text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
import os
from pprint import pprint
from solver.darp import Darp
from instance import parser as instance_parser
from solution import parser as solution_parser
import plot.route as route_plot
import json
import logging
# Change logging level to DEBUG to see the construction of the model
format = "%(asctime)s | %(name)s | %(levelname)s | %(message)s"
logging.basicConfig(level=logging.INFO, format=format)
logger = logging.getLogger()
result_folder = "./results/darp_heuristic/"
os.makedirs(result_folder, exist_ok=True)
instance_folder = "./instance/data/darp_cordeau_2006/"
filename = "a2-16"
filepath = os.path.join(instance_folder, filename)
instance = instance_parser.parse_instance_from_filepath(
filepath, instance_parser=instance_parser.PARSER_TYPE_CORDEAU
)
data = instance.get_data()
data["dist_matrix"][0][0]
data["L"]
customers = data["P"]
customers_tw = [(i, data["el"][i]) for i in customers]
pprint(customers_tw)
```
#### Step 1
Index customers in the order of their earliest pick-up times:
```
customers_earliest = sorted(customers_tw, key=lambda x:x[1][0])
pprint(customers_earliest)
from collections import OrderedDict
def get_cost(k, route):
capacity = 0
previous = route[0]
arrival_previous = 0
service_previous = data["d"][previous]
departure_previous = arrival_previous + service_previous
cost = 0
waiting = 0
result = OrderedDict()
result[previous] = dict(
w=0,
b=arrival_previous,
t=0,
q=capacity)
for current in route[1:]:
capacity+= data["q"][current]
if capacity > data["Q"][k]:
return -1, -1
earliest_current = data["el"][current][0]
latest_current = data["el"][current][1]
travel_time = data["dist_matrix"][previous][current]
arrival_current = departure_previous + travel_time
print(f"{previous:>2} -> {travel_time:4.1f} -> {current:>2} ({earliest_current:>5.1f} - {arrival_current:>5.1f} - {latest_current:>6.1f} - cost = {cost:>7.2f} - waiting = {waiting:>7.2f}")
# Update previous
if arrival_current <= latest_current:
cost += travel_time
waiting_at_current = max(earliest_current - arrival_current, 0)
waiting += waiting_at_current
arrival_current = max(departure_previous + travel_time, earliest_current)
ride_delay = 0
if current in data["D"]:
i_delivery_node = data["D"].index(current)
pickup_node = data["P"][i_delivery_node]
departure_at_pickup = result[pickup_node]["b"]
ride_delay = arrival_current - departure_at_pickup
print(pickup_node, current, ride_delay, data["dist_matrix"][pickup_node][current])
if ride_delay > data["L"][pickup_node]:
return -1, -1
result[current] = dict(
w=waiting_at_current,
b=arrival_current,
t=ride_delay,
q=capacity)
previous = current
arrival_previous = arrival_current
service_previous = data["d"][current]
departure_previous = arrival_current + service_previous
else:
return -1, -1
print(f"{previous:>2} -> {travel_time:4.1f} -> {current:>2} ({earliest_current:>5.1f} - {arrival_current:>5.1f} - {latest_current:>6.1f} - cost = {cost:>7.2f} - waiting = {waiting:>7.2f}")
pprint(result)
return cost, waiting
work_schedule = dict()
work_schedule[0] = [0, 12, 6, 28, 22, 4, 11, 27, 20, 3, 19, 13, 29, 9, 8, 25, 24, 2, 18, 1, 17, 0]
work_schedule[1] = [0, 10, 5, 26, 21, 14, 30, 15, 31, 7, 16, 23, 32, 0]
total_cost = 0
total_slack = 0
for k, route in work_schedule.items():
cost, waiting = get_cost(k, route)
total_cost+=cost
total_slack+=waiting
print(k, cost, waiting)
print(total_cost, total_slack)
```
#### Step 2
For each vehicle $j (j = 1, 2, . . . , n)$:
1. Find all the feasible ways in which customer $i$ can be inserted into the work-schedule of vehicle $j$. If it is infeasible to assign customer $i$ to vehicle $j$, examine the next vehicle $j + 1$ (restart *Step 1*); otherwise:
2. Find the insertion of customer i into the work-schedule of vehicle j that results in minimum additional cost (details in Section 5). Call this additional cost COST,.
```
from collections import defaultdict
import math
work_schedule_vehicle_dict = defaultdict(list)
n = len(data["P"])
temp_work_schedule_vehicle_dict = defaultdict(list)
, 'q': 1.0}), (32, {'w': 0.5080375048326005, 'b': 414.0, 't': 30.0, 'q': 0.0}), ('0*', {'w': 0, 'b': 480.0, 't': 0, 'q': 0.0})]})])
# 0 [0, 12, 6, 28, 22, 4, 11, 27, 20, 3, 19, 13, 29, 9, 8, 25, 24, 2, 18, 1, 17, '0*']
# 1 [0, 10, 5, 26, 21, 14, 30, 15, 31, 7, 16, 23, 32, '0*']
def get_cost(route):
o = route[0]
arrival_o = 0
service_o = data["d"][o]
departure_o = arrival_o + service_o
cost = 0
slack = 0
for d in route[1:]:
travel_time_od = data["dist_matrix"][o][d]
earliest_d = data["el"][d][0]
arrival_d = max(departure_o + service_o + travel_time_od, earliest_d)
latest_d = data["el"][d][1]
# Update o
if arrival_d <= latest_d:
cost += travel_time_od
slack += arrival_d - earliest_d
o = d
arrival_o = arrival_d
service_o = data["d"][d]
departure_o = arrival_d + service_o
else:
return -1, -1
return cost, slack
def get_min_insertion(route, pickup, dropoff):
for i in range(len(route)):
temp_route = route.copy()
temp_route.insert(i, pickup)
for j in range(i+1, len(route)):
temp_route.insert(j, dropoff)
cost = get_cost(temp_route)
for k in data["K"]:
work_schedule = work_schedule_vehicle_dict[k].copy()
for pickup_node in customers_earliest:
i_pickup_node = data["N"].index(pickup_node)
delivery_node = data["N"][n + i_pickup_node + 1]
get_min_insertion(temp_work_schedule_vehicle_dict)
data["dist_matrix"][0][0]
print(k)
# ”, EPTi (i = 1, . . . , N), i.e. according to the earliest time at which they are expected to be available for a pick-up. Section 4 shows how EPT, is computed.
# result["instance"] = filename
# solution_obj = solution_parser.parse_solution_dict(result)
s# ol_filepath = f"{os.path.join(result_folder, filename)}.json"
# with open(sol_filepath, "w") as outfile:
# json.dump(result, outfile, indent=4)
# fig, ax = route_plot.plot_vehicle_routes(
# instance,
# solution_obj,
# jointly=True,
# figsize=(10, 10),
# show_arrows=False,
# show_node_labels=False,
# )
# fig_filepath = f"{os.path.join(result_folder, filename)}.pdf"
# fig.savefig(fig_filepath, bbox_inches="tight")
```
| github_jupyter |
# Get Started with TensorFlow
TensorFlow is an open-source machine learning library for research and production. TensorFlow offers APIs for beginners and experts to develop for desktop, mobile, web, and cloud. See the sections below to get started.
## Learn and use ML
The high-level Keras API provides building blocks to create and train deep learning models. Start with these beginner-friendly notebook examples, then read the TensorFlow Keras guide.
### Demo: MNIST (MLP 784-512-512-10)
```
# Let's get started, import the TensorFlow library into your program:
import tensorflow as tf
mnist = tf.keras.datasets.mnist
# Load and prepare the MNIST dataset. Convert the samples from integers to floating-point numbers:
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# Build the tf.keras model by stacking layers. Select an optimizer and loss function used for training:
model = tf.keras.models.Sequential([
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(512, activation=tf.nn.relu),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# Train and evaluate model:
model.fit(x_train, y_train, epochs=5)
scores = model.evaluate(x_test, y_test)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# Save entire model to a HDF5 file
model.save('./models/my_mnist_model.h5')
```
You’ve now trained an image classifier with 97%~98% accuracy on this dataset.
```
print(model.summary())
```
## Train your first neural network: basic classification
This guide trains a neural network model to classify images of clothing, like sneakers and shirts. It's okay if you don't understand all the details, this is a fast-paced overview of a complete TensorFlow program with the details explained as we go.
This guide uses tf.keras, a high-level API to build and train models in TensorFlow.
```
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
```
### Import the Fashion MNIST dataset
This guide uses the [Fashion MNIST](https://github.com/zalandoresearch/fashion-mnist) dataset which contains 70,000 grayscale images in 10 categories. The images show individual articles of clothing at low resolution (28 by 28 pixels), as seen here:

Fashion MNIST is intended as a drop-in replacement for the classic MNIST dataset—often used as the "Hello, World" of machine learning programs for computer vision. The MNIST dataset contains images of handwritten digits (0, 1, 2, etc) in an identical format to the articles of clothing we'll use here.
This guide uses Fashion MNIST for variety, and because it's a slightly more challenging problem than regular MNIST. Both datasets are relatively small and are used to verify that an algorithm works as expected. They're good starting points to test and debug code.
We will use 60,000 images to train the network and 10,000 images to evaluate how accurately the network learned to classify images. You can access the Fashion MNIST directly from TensorFlow, just import and load the data:
```
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
```
Loading the dataset returns four NumPy arrays:
- The train_images and train_labels arrays are the training set—the data the model uses to learn.
- The model is tested against the test set, the test_images, and test_labels arrays.
The images are 28x28 NumPy arrays, with pixel values ranging between 0 and 255. The labels are an array of integers, ranging from 0 to 9. These correspond to the class of clothing the image represents:
Label | Class
----- | -----
0 | T-shirt/top
1 | Trouser
2 | Pullover
3 | Dress
4 | Coat
5 | Sandal
6 | Shirt
7 | Sneaker
8 | Bag
9 | Ankle boot
Each image is mapped to a single label. Since the class names are not included with the dataset, store them here to use later when plotting the images:
```
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
```
### Explore the data
Let's explore the format of the dataset before training the model. The following shows there are 60,000 images in the training set, with each image represented as 28 x 28 pixels:
```
train_images.shape
```
Likewise, there are 60,000 labels in the training set:
```
len(train_labels)
```
Each label is an integer between 0 and 9:
```
train_labels
```
There are 10,000 images in the test set. Again, each image is represented as 28 x 28 pixels:
### Preprocess the data
The data must be preprocessed before training the network. If you inspect the first image in the training set, you will see that the pixel values fall in the range of 0 to 255:
```
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
```
We scale these values to a range of 0 to 1 before feeding to the neural network model. For this, cast the datatype of the image components from an integer to a float, and divide by 255. Here's the function to preprocess the images:
It's important that the training set and the testing set are preprocessed in the same way:
```
train_images = train_images / 255.0
test_images = test_images / 255.0
```
Display the first 25 images from the training set and display the class name below each image. Verify that the data is in the correct format and we're ready to build and train the network.
```
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
```
### Build the model
Building the neural network requires configuring the layers of the model, then compiling the model.
#### Setup the layers
The basic building block of a neural network is the layer. Layers extract representations from the data fed into them. And, hopefully, these representations are more meaningful for the problem at hand.
Most of deep learning consists of chaining together simple layers. Most layers, like `tf.keras.layers.Dense`, have parameters that are learned during training.
```
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
```
The first layer in this network, `tf.keras.layers.Flatten`, transforms the format of the images from a 2d-array (of 28 by 28 pixels), to a 1d-array of 28 * 28 = 784 pixels. Think of this layer as unstacking rows of pixels in the image and lining them up. This layer has no parameters to learn; it only reformats the data.
After the pixels are flattened, the network consists of a sequence of two `tf.keras.layers.Dense` layers. These are densely-connected, or fully-connected, neural layers. The first Dense layer has 128 nodes (or neurons). The second (and last) layer is a 10-node softmax layer—this returns an array of 10 probability scores that sum to 1. Each node contains a score that indicates the probability that the current image belongs to one of the 10 classes.
#### Compile the model
Before the model is ready for training, it needs a few more settings. These are added during the model's compile step:
- **Loss function** —This measures how accurate the model is during training. We want to minimize this function to "steer" the model in the right direction.
- **Optimizer** —This is how the model is updated based on the data it sees and its loss function.
- **Metrics** —Used to monitor the training and testing steps. The following example uses accuracy, the fraction of the images that are correctly classified.
```
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
### Train the model
Training the neural network model requires the following steps:
1. Feed the training data to the model—in this example, the train_images and train_labels arrays.
2. The model learns to associate images and labels.
3. We ask the model to make predictions about a test set—in this example, the test_images array. We verify that the predictions match the labels from the test_labels array.
To start training, call the model.fit method—the model is "fit" to the training data:
```
model.fit(train_images, train_labels, epochs=5)
```
As the model trains, the loss and accuracy metrics are displayed. This model reaches an accuracy of about 0.88~0.90 on the training data. You can evaluate the training results as follows:
```
scores = model.evaluate(train_images, train_labels)
print("Training %s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
```
### Evaluate accuracy
Next, compare how the model performs on the test dataset:
```
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
```
It turns out, the accuracy on the test dataset is a little less than the accuracy on the training dataset. This gap between training accuracy and test accuracy is an example of **overfitting**. Overfitting is when a machine learning model performs worse on new data than on their training data.
### Make predictions
With the model trained, we can use it to make predictions about some images.
```
predictions = model.predict(test_images)
```
Here, the model has predicted the label for each image in the testing set. Let's take a look at the first prediction:
```
predictions[0]
```
A prediction is an array of 10 numbers. These describe the "confidence" of the model that the image corresponds to each of the 10 different articles of clothing. We can see which label has the highest confidence value:
```
np.argmax(predictions[0])
```
So the model is most confident that this image is an ankle boot, or class_names. And we can check the test label to see this is correct:
```
test_labels[0]
print(np.shape(test_labels))
print(np.shape(predictions))
predicted_labels = tf.argmax(predictions, axis = 1)
print(np.shape(predicted_labels))
confusion = tf.confusion_matrix(labels = test_labels, predictions = predicted_labels, num_classes = 10)
print(confusion.eval(session=tf.Session()))
#tf.enable_eager_execution() # tf.enable_eager_execution must be called at program startup.
print(confusion)
```
We can graph this to look at the full set of 10 channels
```
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array[i], true_label[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
```
Let's look at the 0th image, predictions, and prediction array.
```
i = 0
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
i = 12
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(1,2,2)
plot_value_array(i, predictions, test_labels)
```
Let's plot several images with their predictions. Correct prediction labels are blue and incorrect prediction labels are red. The number gives the percent (out of 100) for the predicted label. Note that it can be wrong even when very confident.
```
# Plot the first X test images, their predicted label, and the true label
# Color correct predictions in blue, incorrect predictions in red
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions, test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions, test_labels)
```
Finally, use the trained model to make a prediction about a single image.
```
# Grab an image from the test dataset
img = test_images[0]
print(img.shape)
```
`tf.keras` models are optimized to make predictions on a batch, or collection, of examples at once. So even though we're using a single image, we need to add it to a list:
```
# Add the image to a batch where it's the only member.
img = (np.expand_dims(img,0))
print(img.shape)
```
Now predict the image:
```
predictions_single = model.predict(img)
print(predictions_single)
plot_value_array(0, predictions_single, test_labels)
_ = plt.xticks(range(10), class_names, rotation=45)
```
`model.predict` returns a list of lists, one for each image in the batch of data. Grab the predictions for our (only) image in the batch:
```
np.argmax(predictions_single[0])
```
And, as before, the model predicts a label of 9.
| github_jupyter |
# Modelling techniques in portfolio optimization using second-order cone programming (SOCP) in the NAG Library
# Correct Rendering of this notebook
This notebook makes use of the `latex_envs` Jupyter extension for equations and references. If the LaTeX is not rendering properly in your local installation of Jupyter , it may be because you have not installed this extension. Details at https://jupyter-contrib-nbextensions.readthedocs.io/en/latest/nbextensions/latex_envs/README.html
The notebook is also not rendered well by GitHub so if you are reading it from there, you may prefer the [pdf version instead](./static/portfolio_optimization_using_socp.pdf).
# Installing the NAG library and running this notebook
This notebook depends on the NAG library for Python to run. Please read the instructions in the [Readme.md](https://github.com/numericalalgorithmsgroup/NAGPythonExamples/blob/master/local_optimization/Readme.md#install) file to download, install and obtain a licence for the library.
Instruction on how to run the notebook can be found [here](https://github.com/numericalalgorithmsgroup/NAGPythonExamples/blob/master/local_optimization/Readme.md#jupyter).
# Note for the users of the NAG Library Mark $27.1$ onwards
At Mark $27.1$ of the NAG Library, NAG introduced two new additions to help users easily define a Quadratically Constrained Quadratic Programming (QCQP) problem. All the models in this notebook then can be solved in a much simpler way without the need of a reformulation or any extra effort. It's recommended that the users of the NAG Library Mark $27.1$ or newer should look at the [notebook on QCQP instead](./portfolio_optimization_qcqp.ipynb).
# Introduction
Second-order cone programming (SOCP) is convex optimization which extends linear programming (LP) with second-order (Lorentz or the ice cream) cones. Search region of the solution is the intersection of an affine linear manifold with the Cartesian product of second-order cones. SOCP appears in a broad range of applications from engineering, control theory and quantitative finance to quadratic programming and robust optimization. It has become an important tool for financial optimization due to its powerful nature. Interior point methods (IPM) are the most popular approaches to solve SOCP problems due to their theoretical polynomial complexity and practical performance.
NAG introduces at Mark $27$ an interior point method for large-scale SOCP problem in the standard form:
\begin{equation}\label{SOCP}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & c^Tx\\[0.6ex]
\mbox{subject to} & l_A\leq Ax\leq u_A,\\[0.6ex]
& l_x\leq x\leq u_x,\\[0.6ex]
& x\in{\cal K},
\end{array}
\end{equation}
where $A\in\Re^{m\times n}$, $l_A, u_A\in\Re^m$, $c, l_x, u_x\in\Re^n$ are the problem data, and $\cal K={\cal K}^{n_1}\times\cdots\times{\cal K}^{n_r}\times\Re^{n_l}$ where ${\cal K}^{n_i}$ is either a quadratic cone or a rotated quadratic cone defined as follows:
\begin{itemize}
\item Quadratic cone:
\begin{equation}\label{SOC}
{\cal K}_q^{n_i}:=\left\lbrace x=\left(x_1,\ldots,x_{n_i}\right)\in\Re^{n_i}~:~x_1^2\geq\sum_{j=2}^{n_i} x_j^2, ~x_1\geq0\right\rbrace.
\end{equation}
\item Rotated quadratic cone:
\begin{equation}\label{RSOC}
{\cal K}_r^{n_i}=\left\lbrace x=\left(x_1,x_2,\ldots,x_{n_i}\right)\in\Re^{n_i}~:~2x_1x_2\geq\sum_{j=3}^{n_i} x_j^2,\quad x_1\geq0,\quad x_2\geq0\right\rbrace.
\end{equation}
\end{itemize}
SOCP is widely used in portfolio optimization due to its flexibility and versatility to handle a large variety of problems with different kinds of constraints, which can be transformed into SOCPs that are equivalent to (\ref{SOCP}), see \cite{AG03, LVBL98} for more details. In the rest of this article we demonstrate how to use SOCP solver in the NAG Library for Python to build and solve models with various practical constraints for portfolio optimization.
# Data preparation
We consider daily prices for the 30 stocks in the DJIA from March 2018 to March 2019. In practice, the estimation of the mean return $r$ and covariance $V$ is often a non-trivial task. In this notebook, we estimate those entities using simple sample estimates.
```
# Import necessary libraries
import pickle as pkl
import numpy as np
import matplotlib.pyplot as plt
# Load stock price data from stock_price.plk
# Stock_price: dict = ['close_price': [data], 'date_index': [data]]
stock_price = stock_price = pkl.load(open('./data/stock_price.pkl', 'rb'))
close_price = stock_price['close_price']
date_index = stock_price['date_index']
# Size of data, m: number of observations, n: number of stocks
m = len(date_index)
n = len(close_price)
# Extract stock closing prices to a numpy array
data = np.zeros(shape=(m, n))
i = 0
for stock in close_price:
data[:,i] = close_price[stock]
plt.plot(np.arange(m), data[:,i])
i += 1
# Plot closing prices
plt.xlabel('Time (days)')
plt.ylabel('Closing price ($)')
plt.show()
```
For each stock $i$, we first estimate the $j$th daily relative return as $$relative~return_{i,j} = \frac{closing~price_{i,j+1}-closing~price_{i,j}}{closing~price_{i,j}}.$$
```
# Relative return
rel_rtn = np.zeros(shape=(m-1, n))
for j in range(m-1):
rel_rtn[j,:] = np.divide(data[j+1,:] - data[j,:], data[j,:])
# Plot relative return
for i in range(n):
plt.plot(np.arange(m-1),rel_rtn[:,i])
plt.xlabel('Time (days)')
plt.ylabel('Relative return')
plt.show()
```
Simply take arithmetic mean of each column of relative return to get mean return $r$ for each stock, followed by estimating covariance $V$ using numpy.
```
# Mean return
r = np.zeros(n)
r = rel_rtn.sum(axis=0)
r = r / (m-1)
# Covariance matrix
V = np.cov(rel_rtn.T)
```
# Classic Mean-Variance Model
## Efficient Frontier
One of the major goals of portfolio management is to achieve a certain level of return under a specific risk measurement. Here we demonstrate how to use NAG Library to build efficient frontier by solving classical Markowitz model with long-only constraint (meaning, buy to hold and and short selling is not allowed):
\begin{equation}\label{MV_model}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & -r^Tx+\mu x^TVx\\[0.6ex]
\mbox{subject to} & e^Tx = 1,\\[0.6ex]
& x\geq0,
\end{array}
\end{equation}
where $e\in\Re^n$ is vector of all ones and $\mu$ is a scalar controling trade-off between return and risk. Note one could build the efficient frontier by varying $\mu$ from $0$ to a certain value.
```
# Import the NAG Library
from naginterfaces.base import utils
from naginterfaces.library import opt
from naginterfaces.library import lapackeig
# Import necessary math libraries
import math as mt
import warnings as wn
```
To solve Problem (\ref{MV_model}) via SOCP, we need to transform it to the standard formulation (\ref{SOCP}) and feed the NAG SOCP solver with data $A$, $l_A$, $u_A$, $c$, $l_x$, $u_x$ and $\cal K$. This modelling process is essential to the usage of SOCP. Getting familiar with these reformulation techniques would unleash the maximum power of SOCP.
To manage the data that the solver requires, one could create and maintain a dictionary structure.
```
def model_init(n):
"""
Initialize a dict to store the data that is used to feed NAG socp solver
"""
model = {
# Number of variables
'n': n,
# Number of constraints
'm': 0,
# Coefficient in objective
'c': np.full(n, 0.0, float),
# Bound constraints on variables
'blx': np.full(n, -1.e20, float),
'bux': np.full(n, 1.e20, float),
# Coefficient in linear constraints and their bounds
'linear': {'bl': np.empty(0, float),
'bu': np.empty(0, float),
'irowa': np.empty(0, int),
'icola': np.empty(0, int),
'a': np.empty(0, float)},
# Cone constraint type and variables group
'cone' : {'type': [],
'group': []}
}
return model
```
Once the data in the model has been completed, we could feed the data to the NAG SOCP solver by the following function.
```
def set_nag(model):
"""
Use data in model to feed NAG optimization suite to define problm
"""
# Creat problem handle
handle = opt.handle_init(model['n'])
# Set objective function
opt.handle_set_linobj(handle, model['c'])
# Set box constraints
opt.handle_set_simplebounds(handle, model['blx'], model['bux'])
# Set linear constraints
opt.handle_set_linconstr(handle, model['linear']['bl'], model['linear']['bu'],
model['linear']['irowa'], model['linear']['icola'], model['linear']['a'])
# Set cone constraints
i = 0
while i<len(model['cone']['type']):
opt.handle_set_group(handle,model['cone']['type'][i],
0, model['cone']['group'][i])
i += 1
# Set options
for option in [
'Print Options = NO',
'Print Level = 1',
'Print File = -1',
'SOCP Scaling = A'
]:
opt.handle_opt_set(handle, option)
return handle
```
Now Let's focus on how to get the data in the model ready. In order to add the quadratic objective to the model, we need the following transformation. Note that by introducing variable $t$,
\begin{equation}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & -r^Tx+\mu x^TVx
\end{array}
\end{equation}
is equivalent to
\begin{equation}\label{e_1}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & -r^Tx+t\\
\mbox{subject to} & \mu x^TVx\leq t
\end{array}
\end{equation}
when $V$ is positive semidefinite. Now the objective in (\ref{e_1}) is linear which fits into the standard model (\ref{SOCP}). By factorizing $V=F^TF$, one can rewrite the quadratic inequality in (\ref{e_1}) to
$$\label{e_2}
\mu\|Fx\|^2\leq t,
$$
where $\|\cdot\|$ is the Euclidean norm. Note that by introducing $y=Fx$ and $s=\frac{1}{2\mu}$, (\ref{e_2}) can be rewritten as
$$
\|y\|^2\leq 2st,
$$
which has exactly the same form of cone constraint (\ref{RSOC}). Therefore, the final SOCP formulation of problem (\ref{MV_model}) is
\begin{equation}\label{MV_model_trans}
\begin{array}{ll}
\underset{x\in\Re^n, y\in\Re^n, s\in\Re, t\in\Re}{\mbox{minimize}} & -r^Tx+t\\[0.6ex]
\mbox{subject to} & e^Tx = 1,\\[0.6ex]
& Fx - y = 0,\\
& s=\frac{1}{2\mu},\\
& x\geq0,\\
& (s,t,y)\in{\cal K}^{n+2}_r.
\end{array}
\end{equation}
Factorization of $V$ can be done using the NAG Library as follows.
```
def factorize(V):
"""
For any given positive semidefinite matrix V, factorize it V = F'*F where
F is kxn matrix that is returned
"""
# Size of V
n = V.shape[0]
# Note one could use sparse factorization if V is input as sparse matrix
U, lamda = lapackeig.dsyevd('V', 'L', V)
# Find positive eigenvalues and corresponding eigenvectors
i = 0
k = 0
F = []
while i<len(lamda):
if lamda[i] > 0:
F = np.append(F, mt.sqrt(lamda[i])*U[:,i])
k += 1
i += 1
F = F.reshape((k, n))
return F
```
The following code adds a general quadratic objective
\begin{equation}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & \frac{1}{2}x^TVx + q^Tx
\end{array}
\end{equation}
to the model.
```
def add_qobj(model, F, q=None):
"""
Add quadratic objective defined as: 1/2 * x'Vx + q'x
transformed to second order cone by adding artificial variables
Parameters
----------
model: a dict with structure:
{
'n': int,
'm': int,
'c': float numpy array,
'blx': float numpy array,
'bux': float numpy array,
'linear': {'bl': float numpy array,
'bu': float numpy array,
'irowa': int numpy array,
'icola': int numpy array,
'a': float numpy array},
'cone' : {'type': character list,
'group': nested list of int numpy arrays}
}
F: float 2d numpy array
kxn dense matrix that V = F'*F where k is the rank of V
q: float 1d numpy array
n vector
Returns
-------
model: modified structure of model
Note: imput will not be checked
"""
# Get the dimension of F (kxn)
k, n = F.shape
# Default q
if q is None:
q = np.zeros(n)
# Up-to-date problem size
m_up = model['m']
n_up = model['n']
# Then k + 2 more variables need to be added together with
# k + 1 linear constraints and a rotated cone contraint
# Enlarge the model
model['n'] = model['n'] + k + 2
model['m'] = model['m'] + k + 1
# Initialize c in the objective
# The order of variable is [x, t, y, s]
model['c'][0:n] = q
model['c'] = np.append(model['c'], np.zeros(k+2))
model['c'][n_up] = 1.0
# Enlarge bounds on x, add inf bounds on the new added k+2 variables
model['blx'] = np.append(model['blx'], np.full(k+2, -1.e20, dtype=float))
model['bux'] = np.append(model['bux'], np.full(k+2, 1.e20, dtype=float))
# Enlarge linear constraints
# Get the aparsity pattern of F
row, col = np.nonzero(F)
val = F[row, col]
# Convert to 1-based and move row down by m
row = row + 1 + m_up
col = col + 1
# Add coefficient of y, t and s to the existing linear coefficient A
# The result is
# [A, 0, 0, 0;
# F, 0, -I, 0;
# 0, 0, 0, 1]
row = np.append(row, np.arange(m_up+1, m_up+k+1+1))
col = np.append(col, np.arange(n_up+2, n_up+k+1+1+1))
val = np.append(val, np.append(np.full(k, -1.0, dtype=float), 1.0))
model['linear']['irowa'] = np.append(model['linear']['irowa'], row)
model['linear']['icola'] = np.append(model['linear']['icola'], col)
model['linear']['a'] = np.append(model['linear']['a'], val)
model['linear']['bl'] = np.append(model['linear']['bl'],
np.append(np.zeros(k), 1.0))
model['linear']['bu'] = np.append(model['linear']['bu'],
np.append(np.zeros(k), 1.0))
# Enlarge cone constraints
model['cone']['type'].extend('R')
group = np.zeros(k+2, dtype=int)
group[0] = n_up + 1
group[1] = n_up + 1 + k + 1
group[2:] = np.arange(n_up+2, n_up+k+1+1)
model['cone']['group'].append(group)
return model
```
Note that in the above function, we require the input to be the factor of V instead of V because of two reasons:
\begin{itemize}
\item In some cases, this factorization is already avaliable or easy to compute from the user, for example, when user is using factor-based expected returns, risk and correlations, he should already have $V = B*CF*B^T + \mbox{Diag}(rv)$, then a factorization is a decomposition of a much smaller matrix CF (factor covariance).
\item In many cases, V is not changing during modifications of models, for example, when adding $1/2 * \mu * x^TVx + q^Tx$ with different $\mu$, users do not need to factorize V every time they change $\mu$.
\end{itemize}
Once the objective of (\ref{MV_model}) has been added, we could use the following function to add the long-only constraint$$e^Tx=1~\mbox{and}~x\geq0.$$
```
def add_longonlycon(model, n, b=None):
"""
Add long-only constraint to model
If no b (benchmark) presents, add sum(x) = 1, x >= 0
If b presents, add sum(x) = 0, x + b >= 0
"""
# Up-to-date problem size
m = model['m']
# No of constraints increased by 1
model['m'] = model['m'] + 1
# Bound constraint: x >=0 or x >= -b
if b is not None:
model['blx'][0:n] = -b
else:
model['blx'][0:n] = np.zeros(n)
# Linear constraint: e'x = 1 or e'x = 0
if b is not None:
model['linear']['bl'] = np.append(model['linear']['bl'],
np.full(1, 0.0, dtype=float))
model['linear']['bu'] = np.append(model['linear']['bu'],
np.full(1, 0.0, dtype=float))
else:
model['linear']['bl'] = np.append(model['linear']['bl'],
np.full(1, 1.0, dtype=float))
model['linear']['bu'] = np.append(model['linear']['bu'],
np.full(1, 1.0, dtype=float))
model['linear']['irowa'] = np.append(model['linear']['irowa'],
np.full(n, m+1, dtype=int))
model['linear']['icola'] = np.append(model['linear']['icola'],
np.arange(1, n+1))
model['linear']['a'] = np.append(model['linear']['a'],
np.full(n, 1.0, dtype=float))
return model
```
By using the functions above, we can easily build the efficient frontier as follows.
```
def ef_lo_basic(n, r, V, step=None):
"""
Basic model to build efficient frontier with long-only constraint
by solving:
min -r'*x + mu * x'Vx
s.t. e'x = 1, x >= 0
Parameters
----------
n: number of assets
r: expected return
V: covariance matrix
step: define smoothness of the curve of efficient frontier,
mu would be generated from [0, 2000] with step in default
Output:
-------
risk: a vector of risk sqrt(x'Vx) with repect to certain mu
rtn: a vector of expected return r'x with repect to certain mu
"""
# Set optional argument
if step is None:
step = 2001
# Factorize V for just one time, use the factorization in the rest of the code
# Eigenvalue decomposition on dense V = U*Diag(lamda)*U' to get V = F'*F
F = factorize(V)
risk = np.empty(0, float)
rtn = np.empty(0, float)
for mu in np.linspace(0.0, 2000.0, step):
# Initialize a data structure for build the model
model = model_init(n)
# Quadratic objective function
muf = F * mt.sqrt(2.0*mu)
model = add_qobj(model, muf, -r)
# Add long-only constraint
model = add_longonlycon(model, n)
# Now use model to feed NAG socp solver
handle = set_nag(model)
# Call socp interior point solver
# Mute warnings and do not count results from warnings
wn.simplefilter('error', utils.NagAlgorithmicWarning)
try:
slt = opt.handle_solve_socp_ipm(handle)
# Compute risk and return from the portfolio
risk = np.append(risk, mt.sqrt(slt.x[0:n].dot(V.dot(slt.x[0:n]))))
rtn = np.append(rtn, r.dot(slt.x[0:n]))
except utils.NagAlgorithmicWarning:
pass
# Destroy the handle:
opt.handle_free(handle)
return risk, rtn
# Build efficient frontier and plot the result
ab_risk, ab_rtn = ef_lo_basic(n, r, V, 500)
plt.plot(ab_risk*100.0, ab_rtn*100.0)
plt.ylabel('Total Expected Return (%)')
plt.xlabel('Absolute Risk (%)')
plt.show()
```
## Maximizing the Sharpe ratio
The Sharpe ratio is defined as the ratio of return of portfolio and standard deviation of the portfolio's excess return. It is usually used to measure the efficiency of a portfolio. Find the most efficient portfolio is equivalent to solve the following optimization problem.
\begin{equation}\label{sr_model}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & \frac{\sqrt{x^TVx}}{r^Tx}\\[0.6ex]
\mbox{subject to} & e^Tx = 1,\\[0.6ex]
& x\geq0.
\end{array}
\end{equation}
By replacing $x$ with $\frac{y}{\lambda}, \lambda\gt0$, model (\ref{sr_model}) is equivalent to
\begin{equation}\label{sr_model_eq}
\begin{array}{ll}
\underset{y\in\Re^n, \lambda\in\Re}{\mbox{minimize}} & y^TVy\\[0.6ex]
\mbox{subject to} & e^Ty = \lambda,\\[0.6ex]
& r^Ty=1, \\
& y\geq0, \\
& \lambda\geq0.
\end{array}
\end{equation}
Problem (\ref{sr_model_eq}) is similar to problem (\ref{MV_model}) in the sense that they both have a quadratic objective function and linear constraints. We could reuse most of the functions above since the reformulation is almost the same except for the definition of linear constraints. For that purpose, we need to following function.
```
def add_sr_lincon(model, r, n):
"""
Add linear constraints for Sharpe ratio problem
e'y = lamda, y >= 0, r'y = 1, lamda >= 0
Enlarge model by 1 more variable lamda
Return: model and index of lambda in the final result, need it to
reconstruct the original solution
"""
# Up-to-date problem size
m_up = model['m']
n_up = model['n']
# Add one more var and two more linear constraints
model['n'] = model['n'] + 1
model['m'] = model['m'] + 2
# Enlarge c by one parameter 0.0
model['c'] = np.append(model['c'], 0.0)
# Bounds constraints on y
model['blx'][0:n] = np.zeros(n)
# Set bound constraints on lamda
model['blx'] = np.append(model['blx'], 0.0)
model['bux'] = np.append(model['bux'], 1.e20)
# Add e'y = lamda
row = np.full(n+1, m_up+1, dtype=int)
col = np.append(np.arange(1, n+1), n_up+1)
val = np.append(np.full(n, 1.0, dtype=float), -1.0)
# Add r'y = 1
row = np.append(row, np.full(n, m_up+2, dtype=int))
col = np.append(col, np.arange(1, n+1))
val = np.append(val, r)
# Update model
model['linear']['irowa'] = np.append(model['linear']['irowa'], row)
model['linear']['icola'] = np.append(model['linear']['icola'], col)
model['linear']['a'] = np.append(model['linear']['a'], val)
# Bounds on linear constraints
model['linear']['bl'] = np.append(model['linear']['bl'],
np.append(np.zeros(1), 1.0))
model['linear']['bu'] = np.append(model['linear']['bu'],
np.append(np.zeros(1), 1.0))
return model, n_up
```
Now we can call the NAG SOCP solver as follows.
```
def sr_lo_basic(n, r, V):
"""
Basic model to calculate efficient portfolio that maximize the Sharpe ratio
min y'Vy
s.t. e'y = lamda, y >= 0, r'y = 1, lamda >= 0
Return efficient portfolio y/lamda and corresponding risk and return
"""
# Factorize V for just one time, use the factorization in the rest of the code
# Eigenvalue decomposition on dense V = U*Diag(lamda)*U' to get V = F'*F
F = factorize(V)
# Initialize a data structure for build the model
model = model_init(n)
# Quadratic objective function
muf = F * mt.sqrt(2.0)
model = add_qobj(model, muf)
# Add linear constraints
model, lamda_idx = add_sr_lincon(model, r, n)
# Now use model to feed NAG socp solver
handle = set_nag(model)
# Call socp interior point solver
slt = opt.handle_solve_socp_ipm(handle)
sr_risk = mt.sqrt(slt.x[0:n].dot(V.dot(slt.x[0:n])))/slt.x[lamda_idx]
sr_rtn = r.dot(slt.x[0:n])/slt.x[lamda_idx]
return sr_risk, sr_rtn, slt.x[0:n]/slt.x[lamda_idx]
# Compute the most efficient portfolio and plot result.
sr_risk, sr_rtn, sr_x = sr_lo_basic(n, r, V)
plt.plot(ab_risk*100.0, ab_rtn*100.0, label='Efficient frontier')
plt.plot([sr_risk*100], [sr_rtn*100], 'rs',
label='Portfolio with maximum Sharpe ratio')
plt.plot([sr_risk*100, 0.0], [sr_rtn*100, 0.0], 'r-', label='Capital market line')
plt.axis([min(ab_risk*100), max(ab_risk*100), min(ab_rtn*100), max(ab_rtn*100)])
plt.ylabel('Total Expected Return (%)')
plt.xlabel('Absolute Risk (%)')
plt.legend()
plt.show()
```
# Portfolio optimization with tracking-error constraint
To avoid taking unnecessary risk when beating a benchmark, the investors commonly impose a limit on the volatility of the deviation of the active portfolio from the benchmark, which is also known as tracking-error volatility (TEV) \cite{J03}. The model to build efficient frontier in excess-return space is
\begin{equation}\label{er_tev}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{maximize}} & r^Tx\\
\mbox{subject to} & e^Tx = 0,\\
& x^TVx\leq tev,
\end{array}
\end{equation}
where $tev$ is a limit on the track-error. Roll \cite{R92} noted that problem (\ref{er_tev}) is totally independent of the benchmark and leads to the unpalatable result that the active portfolio has systematically higher risk than the benchmark and is not optimal. Therefore, in this section we solve a more advanced model by taking absolute risk into account as follows.
\begin{equation}\label{tev_model}
\begin{array}{ll}
\underset{x\in\Re^n}{\mbox{minimize}} & -r^Tx+\mu (x+b)^TV(x+b)\\
\mbox{subject to} & e^Tx = 0,\\
& x^TVx\leq tev,\\
& x+b\geq0,
\end{array}
\end{equation}
where $b$ is a benchmark portfolio. In this demonstration, it is generated synthetically. Note here we use the same covariance matrix $V$ for tev and absolute risk measurement for demonstration purpose. In practice one could use different covariance matrices from different markets.
```
# Generate a benchmark portfolio from efficient portfolio that
# maximizes the Sharpe ratio
# Perturb x
b = sr_x + 1.e-1
# Normalize b
b = b/sum(b)
# Compute risk and return at the benchmark
b_risk = mt.sqrt(b.dot(V.dot(b)))
b_rtn = r.dot(b)
```
Note that same as in problem (\ref{MV_model}), the objective function in (\ref{tev_model}) is quadratic, so we can use $add\_qobj()$ to add it to the model. But problem (\ref{tev_model}) has a quadratic constraint, which makes it a quadratically constrained quadratic programming (QCQP). Following a similar procedure to the transformation of constraint in (\ref{e_1}), we can write a function that can be reused repeatedly to add general quadratic constraints.
```
def add_qcon(model, F, q=None, r=None):
"""
Add quadratic contraint defined as: 1/2 * x'Vx + q'x + r <= 0,
which is equivalent to t + q'x + r = 0, 1/2 * x'Vx <= t,
transformed to second order cone by adding artificial variables
Parameters
----------
model: a dict with structure:
{
'n': int,
'm': int,
'c': float numpy array,
'blx': float numpy array,
'bux': float numpy array,
'linear': {'bl': float numpy array,
'bu': float numpy array,
'irowa': int numpy array,
'icola': int numpy array,
'a': float numpy array},
'cone' : {'type': character list,
'group': nested list of int numpy arrays}
}
F: float 2d numpy array
kxn dense matrix that V = F'*F where k is the rank of V
q: float 1d numpy array
n vector
r: float scalar
Returns
-------
model: modified structure of model
Note: imput will not be checked
"""
# Default parameter
if r is None:
r = 0.0
# Get the dimension of F (kxn)
k, n = F.shape
# Up-to-date problem size
m_up = model['m']
n_up = model['n']
# Then k + 2 more variables need to be added together with
# k + 2 linear constraints and a rotated cone contraint
# Enlarge the model
model['n'] = model['n'] + k + 2
model['m'] = model['m'] + k + 2
# All the added auxiliary variables do not take part in obj
# So their coeffients in obj are all zeros.
model['c'] = np.append(model['c'], np.zeros(k+2))
# Enlarge bounds on x, add inf bounds on the new added k+2 variables
model['blx'] = np.append(model['blx'], np.full(k+2, -1.e20, dtype=float))
model['bux'] = np.append(model['bux'], np.full(k+2, 1.e20, dtype=float))
# Enlarge linear constraints
row, col = np.nonzero(F)
val = F[row, col]
# Convert to 1-based and move row down by m_up
# Add Fx = y and s = 1
# [x,t,y,s]
row = row + 1 + m_up
col = col + 1
row = np.append(np.append(row, np.arange(m_up+1, m_up+k+1+1)), m_up+k+1+1)
col = np.append(np.append(col, np.arange(n_up+2, n_up+k+1+1+1)), n_up+1)
val = np.append(np.append(val, np.append(np.full(k, -1.0,
dtype=float), 1.0)), 1.0)
model['linear']['irowa'] = np.append(model['linear']['irowa'], row)
model['linear']['icola'] = np.append(model['linear']['icola'], col)
model['linear']['a'] = np.append(model['linear']['a'], val)
model['linear']['bl'] = np.append(np.append(model['linear']['bl'],
np.append(np.zeros(k), 1.0)), -r)
model['linear']['bu'] = np.append(np.append(model['linear']['bu'],
np.append(np.zeros(k), 1.0)), -r)
# Add t + q'x + r = 0
if q is not None:
model['linear']['irowa'] = np.append(model['linear']['irowa'],
np.full(n, m_up+k+2, dtype=int))
model['linear']['icola'] = np.append(model['linear']['icola'],
np.arange(1, n+1))
model['linear']['a'] = np.append(model['linear']['a'], q)
# Enlarge cone constraints
model['cone']['type'].extend('R')
group = np.zeros(k+2, dtype=int)
group[0] = n_up + 1
group[1] = n_up + 1 + k + 1
group[2:] = np.arange(n_up+2, n_up+k+1+1)
model['cone']['group'].append(group)
return model
```
By using the function above, we can easily build the efficient frotier with TEV constraint as follows.
```
def tev_lo(n, r, V, b, tev, step=None):
"""
TEV contrained portforlio optimization with absolute risk taken into
consideration by solving:
min -r'y + mu*(b+y)'V(b+y)
s.t. sum(y) = 0, y+b >=o, y'Vy <= tev
"""
# Set optional argument
if step is None:
step = 2001
# Factorize V for just one time, use the factorization in the rest of the code
# Eigenvalue decomposition on dense V = U*Diag(lamda)*U' to get V = F'*F
F = factorize(V)
risk = np.empty(0, float)
rtn = np.empty(0, float)
for mu in np.linspace(0.0, 2000.0, step):
# Initialize a data structure for build the model
model = model_init(n)
# Add long-only constraint
model = add_longonlycon(model, n, b)
# Quadratic objective function
muf = F * mt.sqrt(2.0*mu)
mur = 2.0*mu*V.dot(b) - r
model = add_qobj(model, muf, mur)
# Add Quadratic constraint y'Vy <= tev
F_hf = F * mt.sqrt(2.0)
model = add_qcon(model, F_hf, r=-tev)
# Now use model to feed NAG socp solver
handle = set_nag(model)
# Call socp interior point solver
# Mute warnings and do not count results from warnings
wn.simplefilter('error', utils.NagAlgorithmicWarning)
try:
slt = opt.handle_solve_socp_ipm(handle)
# Compute risk and return from the portfolio
risk = np.append(risk, mt.sqrt((slt.x[0:n]+b).dot(V.dot(slt.x[0:n]+b))))
rtn = np.append(rtn, r.dot(slt.x[0:n]+b))
except utils.NagAlgorithmicWarning:
pass
# Destroy the handle:
opt.handle_free(handle)
return risk, rtn
# Set limit on tracking-error
tev = 0.000002
# Solve the model
tev_risk, tev_rtn = tev_lo(n, r, V, b, tev, step=500)
# Plot the result
plt.figure(figsize=(7.5, 5.5))
plt.plot(ab_risk*100.0, ab_rtn*100.0, label='Classic efficient frontier')
plt.plot([sr_risk*100], [sr_rtn*100], 'rs',
label='Portfolio with maximum Sharpe ratio')
plt.plot([sr_risk*100, 0.0], [sr_rtn*100, 0.0], 'r-', label='Capital market line')
plt.plot(b_risk*100, b_rtn*100, 'r*', label='Benchmark portfolio')
plt.plot(tev_risk*100.0, tev_rtn*100.0, 'seagreen',
label='Efficient frontier with tev constraint')
plt.axis([min(ab_risk*100), max(ab_risk*100), min(tev_rtn*100), max(ab_rtn*100)])
plt.ylabel('Total Expected Return (%)')
plt.xlabel('Absolute Risk (%)')
plt.legend()
plt.show()
```
# Conclusion
In this notebook, we demonstrated how to use NAG Library to solve various models in portfolio optimization. One could take some of the functions mentioned above and start to build their own model immediately. It is worth pointing out that the versatility of SOCP is not just limited to the models mentioned here. It covers a lot more problems and constraints. For example, DeMiguel et al. \cite{DGNU09} discussed portfolio optimization with norm constraint, which can be easily transformed into an SOCP problem. We refer readers to the NAG Library documentation \cite{NAGDOC} on SOCP solver and \cite{AG03, LVBL98} for more details.
# References
[<a id="cit-AG03" href="#call-AG03">1</a>] Alizadeh Farid and Goldfarb Donald, ``_Second-order cone programming_'', Mathematical programming, vol. 95, number 1, pp. 3--51, 2003.
[<a id="cit-LVBL98" href="#call-LVBL98">2</a>] Lobo Miguel Sousa, Vandenberghe Lieven, Boyd Stephen <em>et al.</em>, ``_Applications of second-order cone programming_'', Linear algebra and its applications, vol. 284, number 1-3, pp. 193--228, 1998.
[<a id="cit-J03" href="#call-J03">3</a>] Jorion Philippe, ``_Portfolio optimization with tracking-error constraints_'', Financial Analysts Journal, vol. 59, number 5, pp. 70--82, 2003.
[<a id="cit-R92" href="#call-R92">4</a>] Roll Richard, ``_A mean/variance analysis of tracking error_'', The Journal of Portfolio Management, vol. 18, number 4, pp. 13--22, 1992.
[<a id="cit-DGNU09" href="#call-DGNU09">5</a>] DeMiguel Victor, Garlappi Lorenzo, Nogales Francisco J <em>et al.</em>, ``_A generalized approach to portfolio optimization: Improving performance by constraining portfolio norms_'', Management science, vol. 55, number 5, pp. 798--812, 2009.
[<a id="cit-NAGDOC" href="#call-NAGDOC">6</a>] Numerical Algorithms Group, ``_NAG documentation_'', 2019. [online](https://www.nag.com/numeric/fl/nagdoc_latest/html/frontmatter/manconts.html)
| github_jupyter |
```
import pickle as pk
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.patches as mpatches
import os
EDS_files = [
'cora_sampling_method=EDS_K_sparsity=100_results.p',
'cora_sampling_method=EDS_K_sparsity=10_results.p',
'cora_sampling_method=EDS_K_sparsity=5_results.p' ]
Greedy_files = [
'cora_sampling_method=Greedy_K_sparsity=100_label_balance=Greedy_noise=0.01_results.p',
'cora_sampling_method=Greedy_K_sparsity=100_label_balance=Greedy_noise=100_results.p',
'cora_sampling_method=Greedy_K_sparsity=100_label_balance=Greedy_noise=1_results.p',
'cora_sampling_method=Greedy_K_sparsity=10_label_balance=Greedy_noise=0.01_results.p',
'cora_sampling_method=Greedy_K_sparsity=10_label_balance=Greedy_noise=100_results.p',
'cora_sampling_method=Greedy_K_sparsity=10_label_balance=Greedy_noise=1_results.p',
'cora_sampling_method=Greedy_K_sparsity=5_label_balance=Greedy_noise=0.01_results.p',
'cora_sampling_method=Greedy_K_sparsity=5_label_balance=Greedy_noise=100_results.p',
'cora_sampling_method=Greedy_K_sparsity=5_label_balance=Greedy_noise=1_results.p']
max_files = ['cora_sampling_method=MaxDegree_maxdegree_results.p']
random_file = ['cora_sampling_method=Random_random_results.p']
def open_files(files):
file_content = []
for file in files:
try:
with open(file, 'rb') as f:
file_content.append(pk.load(f, encoding='latin1'))
except Exception as e:
print(e)
print("No " + file)
return file_content
eds_results = open_files(EDS_files)
geedy_results = open_files(Greedy_files)
max_results = open_files(max_files)
random_results = open_files(random_file)
def results_to_lines(results):
lines = []
for result in results:
line = result['results']
x = []
y = []
var = []
for point in line:
x.append(point[1])
y.append(point[2])
var.append(point[3])
lines.append((x,y,var,result['info']))
return lines
random_ref_line = results_to_lines(random_results)[0]
def plot(title, save_file,lines, label_name = None):
plt.errorbar(random_ref_line[0],random_ref_line[1],yerr=random_ref_line[2],alpha = 0.7,color = 'r',label="Random sampling",fmt='o-')
for line in lines:
if label_name is not None:
plt.errorbar(line[0],line[1],yerr=line[2],alpha = 0.5,label=label_name+":"+str(line[3][label_name]),fmt='o-')
else:
plt.errorbar(line[0],line[1],yerr=line[2],alpha = 0.5,fmt='o-')
plt.plot(23,0.81,'ko')
plt.xlabel('known labels of training set %')
plt.ylabel('test accuracy')
# plt.title(title)
plt.legend(loc=4)
plt.grid(True)
plt.savefig(os.path.join('../report',save_file), bbox_inches="tight", dpi = 300)
plot("EDS sampling","EDS_sampling_noallfeaturesK_1.jpg",results_to_lines(eds_results)[0:1],'K_sparsity')
plot("EDS sampling","EDS_sampling_noallfeaturesK.jpg",results_to_lines(eds_results)[0:1],'K_sparsity')
plot("EDS sampling","EDS_sampling_noallfeatures.jpg",results_to_lines(eds_results)[0:1],'K_sparsity')
plot("Greedy sampling, K sparsity = 100 ","Greedy_K100_sampling_noallfeatures.jpg",results_to_lines(geedy_results[0:3]),'noise')
plot("Greedy sampling, K sparsity = 10 ","Greedy_K10_sampling_noallfeatures.jpg",results_to_lines(geedy_results[3:6]),'noise')
plot("Greedy sampling, K sparsity = 5 ","Greedy_K5_sampling_noallfeatures.jpg",results_to_lines(geedy_results[6:9]),'noise')
plot("Max degree sampling,","Max_sampling_noallfeatures.jpg",results_to_lines(max_results))
```
| github_jupyter |
```
import pandas as pd
from matplotlib import pyplot as plt
import matplotlib.ticker as ticker
import seaborn as sns
from pandas.plotting import register_matplotlib_converters
register_matplotlib_converters()
%matplotlib inline
plt.style.use('fivethirtyeight')
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['lines.linewidth'] = 1.5
darkgrey = '#3A3A3A'
lightgrey = '#414141'
barblue = plt.rcParams['axes.prop_cycle'].by_key()['color'][0]
plt.rcParams['text.color'] = darkgrey
plt.rcParams['axes.labelcolor'] = darkgrey
plt.rcParams['xtick.color'] = lightgrey
plt.rcParams['ytick.color'] = lightgrey
```
# Your first Monte Carlo Simulation
## Your goal
You want to forecast with a monte carlo simulation based on the gathered data of your team. The data is stored in the raw.csv.
With your simulation you want to answer the question "How many stories can we do in a given time span?"
## How this notebook is structurued
This notebook gives you a structure on how to create a forecast with a monte carlo simulation. Step by step you analyse the data and create the forecast. Each step builds on the previous one. In order to guide and help you each step consists of:
* A small description on what to do in this step
* Code for visualizing the data in this step (optional to use it - but saves time)
* If you get stuck, don't worry. For each step there is a CSV with data needed for this step.
Feel free to follow the structure or find your own way!
## 1. Read and check the raw data
### Goal
This step reads the raw.csv file as pandas.DataFrame and reduces the columns to the data points to calculate throughput (Completed items per day).
Get a feeling about the data:
* What else is in the data set?
* Where could you get this data from in your project?
### Input
raw.csv
### Visualization
Output the data as table (e.g. pd.DataFrame.head())
```
kanban_data = pd.read_csv('raw.csv', usecols=['Done', 'Type'], parse_dates=['Done']).dropna()
kanban_data.head(1)
```
## 2. Read and check the raw data
### Goal
Calculate the throughput (items completed) per day and visualize it over time (e.g. per day or week). Does the data set look valid?
You need the data of througput per day for the next step.
### Visualization
Code to create a simple plot to show datapoints over time is given. X=Date, Y=Throughput
```
# Use the DataFrame kanban_data of the previous step
# Start coding here
# Stuck? Use this to proceed to the next step: throughput_per_week = pd.read_csv('throughput_per_week.csv')
ax = throughput_per_week.plot(
x='Date', y='Throughput', linewidth=2.5, figsize=(14, 3), legend=None)
ax.set_title("Throughput per Week", loc='left', fontdict={
'fontsize': 18, 'fontweight': 'semibold'})
ax.set_xlabel('')
ax.set_ylabel('Items Completed')
ax.axhline(y=0, color=lightgrey, alpha=.5);
```
## 3. Run a Monte Carlo Simulation
### Goal
Run a monte carlo simulation 'how many items can we complete in X days?' with the following steps:
* Define the datapoints you want to use for the simulation (e.g. last 100 days)
* Define the number of days you want to simulate (e.g. 14 days)
* Simulate the number of days at least 10000 times by randomly picking data points for each day
The result is a distribution of how many times a number of completed items has occured in the simulations.
### Visualization
Given: Code to create simple bar plot to visualize the output of the simulation: X=Items Completed, Y=# of occurences of this number of items completed
```
### SIMULATION INPUT ####
LAST_DAYS = 100
SIMULATION_DAYS = 14
SIMULATIONS = 10000
###
# Start coding here, use "throughput per day" of the previous step
# Stuck? Use this to proceed to the next step: distribution = pd.read_csv('distribution.csv')
plt.figure(figsize=(14, 3))
ax = sns.barplot(x='Items', y='Frequency', data=distribution, color=barblue)
ax.set_title(f"Distribution of Monte Carlo Simulation 'How Many' ({SIMULATIONS} Runs)", loc='left',
fontdict={'size': 18, 'weight': 'semibold'})
ax.set_xlabel(f"Total Items Completed in {SIMULATION_DAYS} Days")
ax.set_ylabel('Frequency')
ax.axhline(y=SIMULATIONS*0.001, color=darkgrey, alpha=.5);
```
## 3. Analysis of the Probabilities of Occurence
### Goal
Use the distribution of the simulation to calculate the probability that a number of items is completed.
### Visualization
Given: Code to create simple bar plot to visualize the output of the simulation and highlight the percentiles 95%, 85%, 70%:
* X=Items Completed, Y=Probability to copmlete the # of items
* To highlight the percentiles the samples of the simulation are needed (list of throughput)
```
# Start coding here, use the distribution DataFrame of the previous step.
# Stuck? Use this to proceed to the next step:
#samples = pd.read_csv('samples.csv')
#probability = pd.read_csv('probability.csv')
plt.figure(figsize=(14, 5))
ax = sns.barplot(x='Items', y='Probability', data=probability, color=barblue)
ax.text(x=-1.4, y=118,
s=f"Probabilities of Completing a Scope in {SIMULATION_DAYS} Days", fontsize=18, fontweight='semibold')
ax.text(x=-1.4, y=110,
s=f"Based on a Monte Carlo Simulations ({SIMULATIONS} Runs) with data of last {LAST_DAYS} days", fontsize=16)
ax.set_ylabel('')
ax.set_xlabel('Total Items Completed')
ax.axhline(y=0.5, color=darkgrey, alpha=.5)
ax.axhline(y=70, color=darkgrey, linestyle='--')
ax.axhline(y=85, color=darkgrey, linestyle='--')
ax.axhline(y=95, color=darkgrey, linestyle='--')
label_xpos = distribution['Items'].max()-2
ax.text(y=70, x=label_xpos, s=f'70%% (%d+ Items)' % samples.Items.quantile(0.3),
va='center', ha='center', backgroundcolor='#F0F0F0')
ax.text(y=85, x=label_xpos, s=f'85%% (%d+ Items)' % samples.Items.quantile(0.15),
va='center', ha='center', backgroundcolor='#F0F0F0')
ax.text(y=95, x=label_xpos, s=f'95%% (%d+ Items)' % samples.Items.quantile(0.05),
va='center', ha='center', backgroundcolor='#F0F0F0')
ax.set_yticklabels(labels=['0', '20', '40', '60', '80', '100%']);
```
| github_jupyter |
# Generate Model Interpreter Report with House Price dataset using Contextual AI
This notebook demonstrates how to generate explanations report using complier implemented in the Contextual AI library.
## Motivation
Once the PoC is done (and you know where your data comes from, what it looks like, and what it can predict) comes the ideal next step is to put your model into production and make it useful for the rest of the business.
Does it sound familiar? do you also need to answer the questions below, before promoting your model into production:
1. _How you sure that your model is ready for production?_
2. _How you able to explain the model performance? in business context that non-technical management can understand?_
3. _How you able to compare between newly trained models and existing models is done manually every iteration?_
In Contextual AI project, our simply vision is to:
1. __Speed up data validation__
2. __Simplify model engineering__
3. __Build trust__
For more details, please refer to our [whitepaper](https://sap.sharepoint.com/sites/100454/ML_Apps/Shared%20Documents/Reusable%20Components/Explainability/XAI_Whitepaper.pdf?csf=1&e=phIUNN&cid=771297d7-d488-441a-8a65-dab0305c3f04)
## Steps
1. Create a model to Predict House Price, using the data provide in [house prices dataset](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data)
2. Evaluate the model performance with Contextual AI report
## Credits
1. Pramodh, Manduri <manduri.pramodh@sap.com>
### 1. Performance Model Training
```
import warnings
from pprint import pprint
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
from xgboost import XGBRegressor
```
#### 1.1. Loading Data and XGB-Model
```
data = pd.read_csv('train.csv')
data.dropna(axis=0, subset=['SalePrice'], inplace=True)
y = data.SalePrice
X = data.drop(['SalePrice', 'Id'], axis=1).select_dtypes(exclude=['object'])
train_X, test_X, train_y, test_y = train_test_split(X.values, y.values, test_size=0.25)
my_imputer = SimpleImputer()
train_X = my_imputer.fit_transform(train_X)
test_X = my_imputer.transform(test_X)
my_model = XGBRegressor(n_estimators=1000,
max_depth=5,
learning_rate=0.1,
subsample=0.7,
colsample_bytree=0.8,
colsample_bylevel=0.8,
base_score=train_y.mean(),
random_state=42, seed=42)
hist = my_model.fit(train_X, train_y,
early_stopping_rounds=5,
eval_set=[(test_X, test_y)], eval_metric='rmse',
verbose=100)
```
#### 1.2. Review Best and Worse Predictions
```
test_pred = my_model.predict(test_X)
errors = test_pred - test_y
sorted_errors = np.argsort(abs(errors))
worse_5 = sorted_errors[-5:]
best_5 = sorted_errors[:5]
print(pd.DataFrame({'worse':errors[worse_5]}))
print()
print(pd.DataFrame({'best':errors[best_5]}))
```
#### 1.3. Perform LIME (Local Interpretable Model-Agnostic Explanations)
```
import lime
import lime.lime_tabular
explainer = lime.lime_tabular.LimeTabularExplainer(train_X, feature_names=X.columns, class_names=['SalePrice'], verbose=True, mode='regression')
```
##### Explaining a few worse predictions:
```
type(train_X)
X.columns.tolist()
import pandas as pd
df1 = pd.DataFrame(data =train_X, columns= X.columns.tolist())
#train_y.tolist()
#X.columns.tolist()
X_train = df1
clf = my_model
clf_fn = my_model.predict
y_train = []
feature_names=X.columns.tolist()
target_names_list =['SalePrice']
pprint(target_names_list)
```
### 2. Involve Contextual AI complier
```
import os
import json
import sys
sys.path.append('../../../')
from xai.compiler.base import Configuration, Controller
```
#### 2.1 Specify config file
```
json_config = 'lime-tabular-regressor-model-interpreter.json'
```
#### 2.2 Load and Check config file (before rendering)
```
with open(json_config) as file:
config = json.load(file)
config
pprint(config)
```
#### 2.3 Initial compiler controller with config - withe locals()
```
controller = Controller(config=Configuration(config, locals()))
pprint(controller.config)
```
#### 2.4 Render report
```
controller.render()
```
### Results
```
pprint("report generated : %s/housingpricing-regression-model-interpreter-report.pdf" % os.getcwd())
('report generated : '
'/Users/i062308/Development/Explainable_AI/tutorials/compiler/housingprices/housingpricing-regression-model-interpreter-report.pdf')
```
| github_jupyter |
```
## install dependencies
# !pip install datasets==1.1.2 pytorch_lightning==1.0.3 wandb==0.10.8 transformers==3.4.0
```
## 0. Dependencies
```
# utils
import os
import gc
import tqdm
import torch
import pandas as pd
# data
from transformers import AutoTokenizer
from datasets import load_dataset
from torch.utils.data import random_split, Dataset, DataLoader
# model
from transformers import AutoModel
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
# training and evaluation
import wandb
import pytorch_lightning as pl
from pytorch_lightning.callbacks import EarlyStopping, ProgressBar, ModelCheckpoint
from pytorch_lightning.loggers import WandbLogger
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score, classification_report, roc_auc_score, roc_curve
import seaborn as sns
import matplotlib.pyplot as plt
# device
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# seed
torch.manual_seed(42)
```
## 1. Custom Dataset
```
class NewsDataset(Dataset):
"Custom Dataset class to create the torch dataset"
def __init__(self, root_dir, tokenizer, max_len=128):
"""
root_dir: path where data is residing
tokenizer: tokenizer will be used to tokenize the text
max_len: max_len for text, padding/trimming will be applied to follow this rule
"""
self.tokenizer = tokenizer
self.data = load_dataset("csv", data_files=[os.path.join(root_dir, "Fake.csv"), os.path.join(root_dir, "True.csv")])['train']
self.text = self.data['title']
self.label = self.data['label']
self.max_len = max_len
def __len__(self):
"__len__ function returns the size of the data ="
return len(self.text)
def __getitem__(self, idx):
"""
idx: index of the data to retrieve
returns: A dictionary containing input ids based on tokenizer's vocabulary, attention mask and label tensors
"""
text = self.text[idx]
label = self.label[idx]
input_encoding = self.tokenizer.encode_plus(
text=text,
truncation=True,
max_length=self.max_len,
return_tensors="pt",
return_attention_mask=True,
padding="max_length",
)
return {
"input_ids":input_encoding['input_ids'].squeeze(),
"attention_mask":input_encoding['attention_mask'].squeeze(),
"label":torch.tensor([label], dtype=torch.float)
}
```
## 2. Model
```
class Model(nn.Module):
"""
Fake News Classifier Model
A pretrained model is used as for contextualized embedding and a classifier on top of that.
"""
def __init__(self, model_name, num_classes=2):
"""
model_name: What base model to use from hugginface transformers
num_classes: Number of classes to classify. This is simple binary classification hence 2 classes
"""
super().__init__()
# pretrained transformer model as base
self.base = AutoModel.from_pretrained(pretrained_model_name_or_path=model_name)
# nn classifier on top of base model
self.classfier = nn.Sequential(*[
nn.Linear(in_features=768, out_features=256),
nn.LeakyReLU(),
nn.Linear(in_features=256, out_features=num_classes),
nn.Sigmoid()
])
def forward(self, input_ids, attention_mask=None):
"""
input_ids: input ids tensors for tokens shape = [batch_size, max_len]
attention_mask: attention for input ids, 0 for pad tokens and 1 for non-pad tokens [batch_size, max_len]
returns: logits tensors as output, shape = [batch, num_classes]
"""
outputs = self.base(input_ids=input_ids, attention_mask=attention_mask)
pooler = outputs[1]
# pooler.shape = [batch_size, hidden_size]
logits = self.classfier(pooler)
return logits
```
## 3. PyTorchLightning Data and Trainer Module
#### Data Module
```
class FakeNewsDataModule(pl.LightningDataModule):
"""Lightning Data Module to detach data from model"""
def __init__(self, config):
"""
config: a dicitonary containing data configuration such as batch size, split_size etc
"""
super().__init__()
self.config = config
# prepare and setup the dataset
self.prepare_data()
self.setup()
def prepare_data(self):
"""prepare datset"""
tokenizer = AutoTokenizer.from_pretrained(self.config['model_name'])
self.dataset = NewsDataset(root_dir=self.config['root_dir'], tokenizer=tokenizer, max_len=self.config['max_len'])
def setup(self):
"""make assignments here (val/train/test split)"""
train_size = self.config['train_size']
lengths = [int(len(self.dataset)*train_size), len(self.dataset)-int(len(self.dataset)*train_size)]
self.train_datset, self.test_dataset = random_split(dataset=self.dataset, lengths=lengths)
def train_dataloader(self):
return DataLoader(dataset=self.train_datset, batch_size=self.config['batch_size'], shuffle=True, num_workers=self.config['num_workers'])
def val_dataloader(self):
return DataLoader(dataset=self.test_dataset, batch_size=self.config['batch_size'], shuffle=False, num_workers=self.config['num_workers'])
def test_dataloader(self):
# same as validation data
return DataLoader(dataset=self.test_dataset, batch_size=self.config['batch_size'], shuffle=False, num_workers=self.config['num_workers'])
```
#### Trainer Module
```
class LightningModel(pl.LightningModule):
"""
LightningModel as trainer model
"""
def __init__(self, config):
"""
config: training and other conifguration
"""
super(LightningModel, self).__init__()
self.config = config
self.model = Model(model_name=self.config['model_name'], num_classes=self.config['num_classes'])
def forward(self, input_ids, attention_mask=None):
logits = self.model(input_ids=input_ids, attention_mask=attention_mask)
return logits.squeeze()
def configure_optimizers(self):
return optim.AdamW(params=self.parameters(), lr=self.config['lr'])
def training_step(self, batch, batch_idx):
input_ids, attention_mask, targets = batch['input_ids'], batch['attention_mask'], batch['label'].squeeze()
logits = self(input_ids=input_ids, attention_mask=attention_mask)
loss = F.mse_loss(logits, targets)
pred_labels = logits.cpu() > 0.5 # logits.argmax(dim=1).cpu() for non-sigmoid
acc = accuracy_score(targets.cpu(), pred_labels)
f1 = f1_score(targets.cpu(), pred_labels, average=self.config['average'])
wandb.log({"loss":loss, "accuracy":acc, "f1_score":f1})
return {"loss":loss, "accuracy":acc, "f1_score":f1}
def validation_step(self, batch, batch_idx):
input_ids, attention_mask, targets = batch['input_ids'], batch['attention_mask'], batch['label'].squeeze()
logits = self(input_ids=input_ids, attention_mask=attention_mask)
loss = F.mse_loss(logits, targets)
pred_labels = logits.cpu() > 0.5 # logits.argmax(dim=1).cpu() for non-sigmoid
acc = accuracy_score(targets.cpu(), pred_labels)
f1 = f1_score(targets.cpu(), pred_labels, average=self.config['average'])
precision = precision_score(targets.cpu(), pred_labels, average=self.config['average'])
recall = recall_score(targets.cpu(), pred_labels, average=self.config['average'])
return {"val_loss":loss, "val_accuracy":torch.tensor([acc]), "val_f1":torch.tensor([f1]), "val_precision":torch.tensor([precision]), "val_recall":torch.tensor([recall])}
def validation_epoch_end(self, outputs):
avg_loss = torch.stack([x['val_loss'] for x in outputs]).mean()
avg_acc = torch.stack([x['val_accuracy'] for x in outputs]).mean()
avg_f1 = torch.stack([x['val_f1'] for x in outputs]).mean()
avg_precision = torch.stack([x['val_precision'] for x in outputs]).mean()
avg_recall = torch.stack([x['val_recall'] for x in outputs]).mean()
wandb.log({"val_loss":avg_loss, "val_accuracy":avg_acc, "val_f1":avg_f1, "val_precision":avg_precision, "val_recall":avg_recall})
return {"val_loss":avg_loss, "val_accuracy":avg_acc, "val_f1":avg_f1, "val_precision":avg_precision, "val_recall":avg_recall}
def test_step(self, batch, batch_idx):
input_ids, attention_mask, targets = batch['input_ids'], batch['attention_mask'], batch['label'].squeeze()
logits = self(input_ids=input_ids, attention_mask=attention_mask)
loss = F.mse_loss(logits, targets)
pred_labels = logits.cpu() > 0.5 # logits.argmax(dim=1).cpu() for non-sigmoid
acc = accuracy_score(targets.cpu(), pred_labels)
f1 = f1_score(targets.cpu(), pred_labels, average=self.config['average'])
precision = precision_score(targets.cpu(), pred_labels, average=self.config['average'])
recall = recall_score(targets.cpu(), pred_labels, average=self.config['average'])
return {"test_loss":loss, "test_precision":torch.tensor([precision]), "test_recall":torch.tensor([recall]), "test_accuracy":torch.tensor([acc]), "test_f1":torch.tensor([f1])}
def test_epoch_end(self, outputs):
avg_loss = torch.stack([x['test_loss'] for x in outputs]).mean()
avg_acc = torch.stack([x['test_accuracy'] for x in outputs]).mean()
avg_f1 = torch.stack([x['test_f1'] for x in outputs]).mean()
avg_precision = torch.stack([x['test_precision'] for x in outputs]).mean()
avg_recall = torch.stack([x['test_recall'] for x in outputs]).mean()
return {"test_loss":avg_loss, "test_precision":avg_precision, "test_recall":avg_recall, "test_acc":avg_acc, "test_f1":avg_f1}
```
## 3. Training and Evaluation
##### Preprocessing
- csv file does not have label coloum, adding it to both csv files and save write access wokring/ directory, will read the data from here now
```
root_dir = "../working/Fake-News/"
os.makedirs(root_dir, exist_ok=True)
fake = pd.read_csv(os.path.join('../input/fake-and-real-news-dataset/', "Fake.csv"))
real = pd.read_csv(os.path.join('../input/fake-and-real-news-dataset/', "True.csv"))
fake['label'] = [1]*fake.shape[0]
real['label'] = [0]*real.shape[0]
fake.to_csv(os.path.join(root_dir, "Fake.csv"))
real.to_csv(os.path.join(root_dir, "True.csv"))
config = {
# data
"root_dir":root_dir,
"model_name":"roberta-base",
"num_classes":1,
"max_len":128,
"train_size":0.85,
"batch_size":32,
"num_workers":4,
# training
"average":"macro",
"save_dir":"./",
"project":"fake-news-classification",
"lr":2e-5,
"monitor":"val_accuracy",
"min_delta":0.005,
"patience":2,
"filepath":"./checkpoints/{epoch}-{val_f1:4f}",
"precision":32,
"epochs":5,
}
### Logger, EarlyStopping and Callbacks
logger = WandbLogger(
name=config['model_name'],
save_dir=config["save_dir"],
project=config["project"],
log_model=True,
)
early_stopping = EarlyStopping(
monitor=config["monitor"],
min_delta=config["min_delta"],
patience=config["patience"],
)
checkpoints = ModelCheckpoint(
filepath=config["filepath"],
monitor=config["monitor"],
save_top_k=1
)
trainer = pl.Trainer(
logger=logger,
gpus=[0],
checkpoint_callback=checkpoints,
callbacks=[early_stopping],
default_root_dir="./models/",
max_epochs=config["epochs"],
precision=config["precision"],
automatic_optimization=True
)
dm = FakeNewsDataModule(config=config)
lm = LightningModel(config=config)
trainer.fit(
model=lm,
datamodule=dm
)
trainer.test(
model=lm,
datamodule=dm
)
```
#### Test from Checkpoint
```
test_loader = dm.test_dataloader()
print(os.listdir("../working/checkpoints/"))
l = torch.load(f="./checkpoints/epoch=2-val_f1=0.999528.ckpt")
lm.load_state_dict(l['state_dict'])
trainer.test(
model=lm,
datamodule=dm
)
```
#### Get the pred probs and predicted labels for test set to compute other metrics
```
actual_label = []
pred_label = []
pred_probs = []
for batch in tqdm.tqdm(test_loader):
y_ = lm(input_ids=batch['input_ids'].to(device), attention_mask=batch['attention_mask'].to(device)).detach().cpu()
actual_label += batch['label'].squeeze().tolist()
pred_probs += y_.tolist()
pred_label += y_ > 0.5
del batch
gc.collect()
print(classification_report(y_true=actual_label, y_pred=pred_label, digits=5))
print("ROBERTa with MSE")
print(classification_report(y_true=actual_label, y_pred=pred_label, digits=5))
```
## AUC ROC Curve
```
ns_probs = [0 for _ in range(len(actual_label))]
roc_auc = roc_auc_score(y_true=actual_label, y_score=pred_probs)
ns_auc = roc_auc_score(y_true=actual_label, y_score=ns_probs)
print(f'ROC_AUC_Score = {roc_auc:.4f}')
lr_fpr, lr_tpr, _ = roc_curve(y_true=actual_label, y_score=pred_probs)
ns_fpr, ns_tpr, _ = roc_curve(y_true=actual_label, y_score=ns_probs)
# plot the roc curve for the model
plt.plot(ns_fpr, ns_tpr, linestyle='--', label='No Skill')
plt.plot(lr_fpr, lr_tpr, marker='.', label='RoBERTa')
# axis labels
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
# show the legend
plt.legend()
# show the plot
plt.show()
```
| github_jupyter |
# Lets build a Deep Feed Forward Neural Network
All dependencies for this notebook is listed in the requirements.txt file. One parent above the nbs directory. This list will keep changing as we add to it so be sure to rerun this line after every git pull
```
from google.colab import drive
drive.mount('/content/gdrive')
%cd gdrive/My Drive/cs271p
!ls
!pip install -r requirements.txt
```
Lets declare our imports
```
import numpy as np
import torch
from torch import nn
import math
from pprint import pprint
from tqdm.notebook import tqdm
```
# Kaggle Datasets
### This is an example of how to download datasets directly from kaggle
```
! pip install -q kaggle
from google.colab import files
files.upload()
! mkdir ~/.kaggle
! cp kaggle.json ~/.kaggle/
! chmod 600 ~/.kaggle/kaggle.json
! kaggle datasets list
! kaggle datasets download -d ronitf/heart-disease-uci
class BasicNeuralNetwork(torch.nn.Module):
def __init__(self, in_size=2, out_size=1, hidden_size=30):
super(BasicNeuralNetwork, self).__init__()
# Set the dimensionality of the network
self.input_size = in_size
self.output_size = out_size
self.hidden_size = hidden_size
# Initialize our weights
self._init_weights()
'''
Initialize the weights
'''
def _init_weights(self):
# Create an input tensor of shape (2,3)
self.W_Input = torch.randn(self.input_size, self.hidden_size)
# Create an output tensor of shape (3, 1)
self.W_Output = torch.randn(self.hidden_size, self.output_size)
'''
Create the forward pass
'''
def forward(self, inputs):
# Lets get the element wise dot product
self.z = torch.matmul(inputs, self.W_Input)
# We call the activation
self.state = self._activation(self.z)
# Pass it through the hidden layer
self.z_hidden = torch.matmul(self.state, self.W_Output)
# Finally activate the output
output = self._activation(self.z_hidden)
# Return the output
return output
'''
Backpropagation algorithm implemented
'''
def backward(self, inputs, labels, output):
# What is the error in output
self.loss = labels - output
# What is the delta loss based on the derivative
self.loss_delta = self.loss * self._derivative(output)
# Get the loss for the existing output weight
self.z_loss = torch.matmul(self.loss_delta, torch.t(self.W_Output))
# Compute the delta like before
self.z_loss_delta = self.z_loss * self._derivative(self.state)
# Finally propogate this to our existing weight tensors to update
# the gradient loss
self.W_Input += torch.matmul(torch.t(inputs), self.z_loss_delta)
self.W_Output += torch.matmul(torch.t(self.state), self.loss_delta)
'''
Here we train the network
'''
def train(self, inputs, labels):
# First we do the foward pass
outputs = self.forward(inputs)
# Then we do the backwards pass
self.backward(inputs, labels, outputs)
'''
Here we perform inference
'''
def predict(self, inputs):
pass
'''
Here we save the model
'''
def save(self):
pass
'''
Our non-linear activation function
'''
def _activation(self, s):
# Lets use sigmoid
return 1 / (1 + torch.exp(-s))
'''
Our derivative function used for backpropagation
Usually the sigmoid prime
'''
def _derivative(self, s):
# derivative of sigmoid
return s * (1 - s)
import pandas as pd
df = pd.read_csv('heart.csv')
df.head(20)
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import accuracy_score
sc = MinMaxScaler((-1, 1))
```
Lets split out dataset between inputs and target
```
df.shape
y = df['target']
X = df.drop('target', axis=1)
```
Lets create a test and train split
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25)
```
Here we transform features by scaling each feature to a given range.
This estimator scales and translates each feature individually such that it is in the given range on the training set, e.g. between zero and one.
```
X_train = sc.fit_transform(X_train)
X_train = torch.tensor(X_train, dtype=torch.float)
y_train = torch.tensor((y_train.values,))
y_train = y_train.transpose(0,1)
print(X_train.shape, y_train.shape)
```
Now we instantiate our neural network
```
net = BasicNeuralNetwork(in_size=X_train.shape[1], out_size=1, hidden_size=100)
```
We train our neural network with 1000 epochs (training loops) and we measure the loss
```
for i in tqdm(range(100)):
outputs = net(X_train)
loss = torch.mean((y_train - outputs)**2).detach().item()
tqdm.write("Loss: {}".format(loss))
net.train(X_train, y_train)
with torch.no_grad():
prediction = net(torch.tensor(sc.transform(X_test), dtype=torch.float))
_, preds_y = torch.max(prediction, 1)
accuracy_score(y_test, preds_y)
```
# Excercises
1. Try to initialize the weights with something better. Hint (Xavier Initialization)
2. Add a bias to the forward pass. Recall the affine transform is (inputs . weights) + bias
3. We are missing a learning rate to the backwards pass. See if you can add that in
# How would we rewrite this code using PyTorch built-in methods
PyTorch gives us most of this functionality out of the box. First we can flag all Tensors to use Autograd. You can read more about autograd here: https://pytorch.org/docs/stable/autograd.html
```
X = df.drop('thalach', axis=1)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25)
# Populate the best
X_train = torch.tensor(sc.fit_transform(X_train), dtype=torch.float, requires_grad=True)
X_test = torch.tensor(sc.transform(X_test), dtype=torch.float)
y_train = torch.tensor(y_train.values)
y_test = torch.tensor(y_test.values)
```
This is the first way using the torch.nn.Sequential. In the Sequential model modules will be added to it in the order they are passed in the constructor. This is a quick way to write a small neural network
```
import torch.nn as nn
from collections import OrderedDict
model = torch.nn.Sequential(OrderedDict([
('fc1', nn.Linear(13, 100)),
('relu1', nn.ReLU()),
('fc2', nn.Linear(100, 100)),
('relu2', nn.ReLU()),
('fc3', nn.Linear(100, 2)),
('sigmoid', nn.Sigmoid())
]))
```
What does the architecture of this model look like
```
print(model)
```
Lets register an optimizer and an objective function
```
optimizer = torch.optim.Adam(model.parameters())
criterion = nn.CrossEntropyLoss()
losses = []
for epoch in tqdm(range(1000)):
optimizer.zero_grad()
outputs = model(X_train)
loss = criterion(outputs, y_train)
loss.backward()
optimizer.step()
losses.append(loss.item())
print("Epoch {}, Loss: {}".format(epoch, loss.item()))
prediction = model(X_test)
_, preds_y = torch.max(prediction, 1)
accuracy_score(y_test, preds_y)
import matplotlib.pyplot as plt
plt.plot(losses, label="Loss Curve")
plt.legend()
plt.show()
losses_no_back = []
for epoch in tqdm(range(1000)):
optimizer.zero_grad()
outputs = model(X_train)
loss = criterion(outputs, y_train)
optimizer.step()
losses_no_back.append(loss.item())
print("Epoch {}, Loss: {}".format(epoch, loss.item()))
import matplotlib.pyplot as plt
plt.plot(losses_no_back, label="Loss Curve")
plt.legend()
plt.show()
```
# Is there a better way?
We looked at the torch.nn.Module before. Inherting from this class gives us:
1. More flexibility on how we build our layers
2. Encapsulate our logic into one object
3. Easily swap out optimization functions
4. Easilty swap out cost functions
```
class DeepNeuralNetwork(nn.Module):
def __init__(self, in_size, out_size, hidden_size, layer_depth=4, activation=nn.ReLU):
super(DeepNeuralNetwork, self).__init__()
self.activation = activation()
self.in_size = in_size
self.out_size = out_size
self.fc1 = nn.Linear(in_size, hidden_size)
self.fcn = nn.ModuleDict({})
for l in range(layer_depth):
name = 'fc'+str(1+l)
self.fcn[name] = nn.Linear(hidden_size, hidden_size)
self.out = nn.Linear(hidden_size, out_size)
self.optimizer = torch.optim.Adam(self.parameters())
self.criterion = nn.CrossEntropyLoss()
self.loss_tracker = []
def add_layer(index, layer):
pass
def forward(self, x):
x = self.activation(self.fc1(x))
for k, l in self.fcn.items():
x = self.activation(x)
x = self.out(x)
return x
def train(self, inputs, labels, test_inputs=None, test_labels=None, epochs=10) -> None:
for epoch in tqdm(range(epochs)):
self.optimizer.zero_grad()
outputs = self(X_train)
loss = self.criterion(outputs, y_train)
loss.backward()
self.optimizer.step()
self.loss_tracker.append(loss.item())
acc = self.accuracy(test_inputs, test_labels)
tqdm.write("Epoch {}, Loss: {} Acc: {}".format(epoch, loss.item(), acc))
def accuracy(self, test_inputs, test_labels):
_, preds_y = torch.max(self(test_inputs), 1)
return accuracy_score(test_labels, preds_y)
def show_loss(self):
import matplotlib.pyplot as plt
plt.plot(self.loss_tracker, label="Loss Curve")
plt.legend()
plt.show()
def predict(self, inputs):
"""
Sets the model to evaluation/inference mode, disabling dropout and
gradient calculation.
"""
self.eval()
return self(inputs)
def summary(self):
from torchsummary import summary
summary(self, (1, 1, self.in_size))
```
We create a new instance of the Module we just created
```
model = DeepNeuralNetwork(13, 2, 100)
```
Lets look at what the model summary or architecture looks like.
We will go into much more depth when look at TensorboardX
```
model.summary()
```
Lets go ahead and train the model
```
model.train(X_train, y_train, X_test, y_test, epochs=1000)
```
Lets get the model accuracy
```
model.accuracy(X_test, y_test)
```
What does the loss curve look like
```
model.show_loss()
```
Lets print out the model summary one more time to see if anything has changed
```
model.summary()
```
The model architecture we built above is sufficient for many problems.
But lets look at a SOTA model and see how complex it can become
```
from PIL import Image
from IPython.display import display
img = Image.open('images/inception_v4.jpeg')
display(img)
```
# Excercises
1. Try to implement early stopping
2. Implement a different activation function. What worked, what didn't
3. Implement a different loss function. What worked and what didn't
4. Introduce a different dataset and see if you can use the above model to build an accurate model
# References
https://arxiv.org/pdf/1602.07261.pdf
https://pytorch.org/docs/stable/nn.html
| github_jupyter |
# Experiments with GalSim data
## Generate images with GalSim
The GalSim library is available from https://github.com/GalSim-developers/GalSim.
```
# experiments/galsim_helper.py contains our functions for interacting with GalSim
import galsim_helper
def three_sources_two_overlap(test_case):
test_case.add_star().offset_arcsec(-5, 5)
(test_case.add_galaxy()
.offset_arcsec(2, 5)
.gal_angle_deg(35)
.axis_ratio(0.2)
)
test_case.add_star().offset_arcsec(10, -10)
test_case.include_noise = True
galsim_helper.generate_fits_file("three_sources_two_overlap", [three_sources_two_overlap, ])
```
## Aperture photometry with SExtractor
```
# sep is a Python interface to the code SExtractor libraries.
# See https://sep.readthedocs.io/ for documentation.
import sep
import numpy as np
from astropy.io import fits
import matplotlib.pyplot as plt
from matplotlib import rcParams
%matplotlib inline
rcParams['figure.figsize'] = [10., 8.]
# read image into standard 2-d numpy array
hdul = fits.open("three_sources_two_overlap.fits")
data = hdul[2].data
data = data.byteswap().newbyteorder()
# show the image
m, s = np.mean(data), np.std(data)
plt.imshow(data, interpolation='nearest', cmap='gray', vmin=m-s, vmax=m+s, origin='lower')
plt.colorbar();
# measure a spatially varying background on the image
bkg = sep.Background(data)
# get a "global" mean and noise of the image background:
print(bkg.globalback)
print(bkg.globalrms)
# evaluate background as 2-d array, same size as original image
bkg_image = bkg.back()
# bkg_image = np.array(bkg) # equivalent to above
# show the background
plt.imshow(bkg_image, interpolation='nearest', cmap='gray', origin='lower')
plt.colorbar();
# subtract the background
data_sub = data - bkg
objs = sep.extract(data_sub, 1.5, err=bkg.globalrms)
# how many objects were detected
len(objs)
from matplotlib.patches import Ellipse
# plot background-subtracted image
fig, ax = plt.subplots()
m, s = np.mean(data_sub), np.std(data_sub)
im = ax.imshow(data_sub, interpolation='nearest', cmap='gray',
vmin=m-s, vmax=m+s, origin='lower')
# plot an ellipse for each object
for i in range(len(objs)):
e = Ellipse(xy=(objs['x'][i], objs['y'][i]),
width=6*objs['a'][i],
height=6*objs['b'][i],
angle=objs['theta'][i] * 180. / np.pi)
e.set_facecolor('none')
e.set_edgecolor('red')
ax.add_artist(e)
nelecs_per_nmgy = hdul[2].header["CLIOTA"]
data_sub.sum() / nelecs_per_nmgy
kronrad, krflag = sep.kron_radius(data, objs['x'], objs['y'], objs['a'], objs['b'], objs['theta'], 6.0)
flux, fluxerr, flag = sep.sum_ellipse(data, objs['x'], objs['y'], objs['a'], objs['b'], objs['theta'],
kronrad, subpix=1)
flux_nmgy = flux / nelecs_per_nmgy
fluxerr_nmgy = fluxerr / nelecs_per_nmgy
for i in range(len(objs)):
print("object {:d}: flux = {:f} +/- {:f}".format(i, flux_nmgy[i], fluxerr_nmgy[i]))
kronrad, krflag = sep.kron_radius(data, objs['x'], objs['y'], objs['a'], objs['b'], objs['theta'], 4.5)
flux, fluxerr, flag = sep.sum_ellipse(data, objs['x'], objs['y'], objs['a'], objs['b'], objs['theta'],
kronrad, subpix=1)
flux_nmgy = flux / nelecs_per_nmgy
fluxerr_nmgy = fluxerr / nelecs_per_nmgy
for i in range(len(objs)):
print("object {:d}: flux = {:f} +/- {:f}".format(i, flux_nmgy[i], fluxerr_nmgy[i]))
```
*Celeste.jl* estimates these flux densities much better. The [galsim_julia.ipynb notebook](https://github.com/jeff-regier/Celeste.jl/blob/master/experiments/galsim_julia.ipynb) shows a run of Celeste.jl on the same data.
## Comparision to Hyper Suprime-Cam (HSC) software pipeline
HSC often fails to deblend images with three light sources in a row, including the following one:

"The single biggest failure mode of the deblender occurs when three or more peaks in a blend appear in a straight
line" -- [Bosch, et al. "The Hyper Suprime-Cam software pipeline." (2018)](https://arxiv.org/abs/1705.06766)
So let's use galsim to generate an images with three peaks in a row!
```
def three_sources_in_a_row(test_case):
x = [-11, -1, 12]
test_case.add_galaxy().offset_arcsec(x[0], 0.3 * x[0]).gal_angle_deg(45)
test_case.add_galaxy().offset_arcsec(x[1], 0.3 * x[1]).flux_r_nmgy(3)
test_case.add_star().offset_arcsec(x[2], 0.3 * x[2]).flux_r_nmgy(3)
test_case.include_noise = True
galsim_helper.generate_fits_file("three_sources_in_a_row", [three_sources_in_a_row, ])
hdul = fits.open("three_sources_in_a_row.fits")
data = hdul[2].data
data = data.byteswap().newbyteorder()
# show the image
m, s = np.mean(data), np.std(data)
fig = plt.imshow(data, interpolation='nearest', cmap='gray', vmin=m-s, vmax=m+s, origin='lower')
plt.colorbar();
```
*Celeste.jl* estimates these flux densities much better. The [galsim_julia.ipynb notebook](https://github.com/jeff-regier/Celeste.jl/blob/master/experiments/galsim_julia.ipynb) shows a run of Celeste.jl on the same data.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
import os
import time
import torch
torch.set_default_tensor_type(torch.DoubleTensor)
from spatial_scene_grammars.nodes import *
from spatial_scene_grammars.rules import *
from spatial_scene_grammars.scene_grammar import *
from spatial_scene_grammars.visualization import *
from spatial_scene_grammars_examples.oriented_clusters.grammar import *
from spatial_scene_grammars.parsing import *
from spatial_scene_grammars.sampling import *
import meshcat
import meshcat.geometry as meshcat_geom
if 'vis' not in globals():
vis = meshcat.Visualizer()
base_url = "http://127.0.0.1"
meshcat_url = base_url + ":" + vis.url().split(":")[-1]
print("Meshcat url: ", meshcat_url)
from IPython.display import HTML
HTML("""
<div style="height: 400px; width: 100%; overflow-x: auto; overflow-y: hidden; resize: both">
<iframe src="{url}" style="width: 100%; height: 100%; border: none"></iframe>
</div>
""".format(url=meshcat_url))
# Draw a random sample from the grammar and visualize it.
grammar = SpatialSceneGrammar(
root_node_type = OrientedClusterRoot,
root_node_tf = torch.eye(4)
)
torch.random.manual_seed(5)
tree = grammar.sample_tree(detach=True)
observed_nodes = tree.get_observed_nodes()
print("Sampled scene with %d clusters and %d boxes." %
(len(tree.find_nodes_by_type(OrientedCluster)),
len(tree.find_nodes_by_type(LongBox))))
print("Sampled tree has score %f" % tree.score().item())
draw_scene_tree_contents_meshcat(tree, zmq_url=vis.window.zmq_url)
draw_scene_tree_structure_meshcat(tree, zmq_url=vis.window.zmq_url, alpha=0.5, node_sphere_size=0.01)
# Draw supertree for this grammar
super_tree = grammar.make_super_tree(max_recursion_depth=10)
nx.draw_networkx(super_tree, with_labels=False)
plt.title("Super tree for oriented clusters grammar")
print("Super tree has %d nodes" % len(list(super_tree.nodes)))
# Parse this tree
inference_results = infer_mle_tree_with_mip(
grammar, observed_nodes, verbose=True, max_scene_extent_in_any_dir=5.
)
mip_optimized_tree = get_optimized_tree_from_mip_results(inference_results)
draw_scene_tree_contents_meshcat(mip_optimized_tree, zmq_url=vis.window.zmq_url, prefix="mip")
draw_scene_tree_structure_meshcat(mip_optimized_tree, zmq_url=vis.window.zmq_url, prefix="mip_scene_tree")
for node in mip_optimized_tree:
err = torch.matmul(node.rotation.transpose(0, 1), node.rotation) - torch.eye(3)
print("Avg elementwise deviation from R^T R = I: ", err.abs().mean())
# Do NLP refinement of tree
refinement_results = optimize_scene_tree_with_nlp(grammar, mip_optimized_tree, verbose=False)
refined_tree = refinement_results.refined_tree
draw_scene_tree_contents_meshcat(refined_tree, zmq_url=vis.window.zmq_url, prefix="mip_refined")
draw_scene_tree_structure_meshcat(refined_tree, zmq_url=vis.window.zmq_url, prefix="mip_refined_scene_tree")
try:
mip_optimized_tree.score()
except ValueError as e:
print("MIP optimized tree wasn't happy, as expected: %s" % str(e))
print("Refined tree score: ", refined_tree.score())
print("Original tree score: ", tree.score())
# Now try to do MCMC on the parsed tree
sampled_trees = do_fixed_structure_mcmc(grammar, tree, num_samples=100, verbose=True)
for sample_k, sampled_tree in enumerate(sampled_trees[::5]):
draw_scene_tree_structure_meshcat(sampled_tree, zmq_url=vis.window.zmq_url, prefix="sampled/%d" % sample_k)
```
| github_jupyter |
```
import numpy as np
import tensorflow as tf
from xfmers import utils
from xfmers import ops
from xfmers import layers
def Transformer(vocab_size, dec_layers, ff_units, d_model, num_heads, dropout, max_seq_len=512, causal=False,
weight_sharing=False, efficient_attention=False, shared_qk=False, activation=ops.gelu,
conv_filter=1, conv_padding="same", reversible=False, fused_qkv=False, name="Transformer"):
inputs = tf.keras.Input(shape=(None, ), name="inputs")
padding_mask = layers.PaddingMaskGenerator()(inputs)
embeddings = layers.TokenPosEmbedding(d_vocab=vocab_size, d_model=d_model, pos_length=max_seq_len, scale=1)(inputs)
decoder_block = layers.TransformerStack(layers=dec_layers,
ff_units=ff_units,
d_model=d_model,
num_heads=num_heads,
dropout=dropout,
causal=causal,
activation=activation,
weight_sharing=weight_sharing,
conv_filter=conv_filter,
conv_padding=conv_padding,
reversible=reversible,
fused_qkv=fused_qkv,
name="DecoderBlock")
dec_outputs = decoder_block({"token_inputs": embeddings,
"mask_inputs": padding_mask})
l_dropout = tf.keras.layers.Dropout(rate=dropout)(dec_outputs)
preds = tf.keras.layers.Dense(vocab_size, name="outputs")(l_dropout)
return tf.keras.Model(inputs=inputs, outputs=preds, name=name)
model = Transformer(vocab_size=8192,
dec_layers=12,
ff_units=3072,
d_model=768,
num_heads=12,
dropout=0.1,
max_seq_len=128,
fused_qkv=True,
causal=True)
model.summary()
tlayer = layers.TransformerLayer(ff_units=768*4, d_model=768, num_heads=12, dropout=0.01, causal=True)
_ = tlayer({"token_inputs": np.zeros((1,128,768)),
"mask_inputs": np.zeros((1,128,1))})
tlayer.summary()
for layer in tlayer.layers:
print("name:", layer.name, " - params:", layer.count_params())
tlayer.layers[0].summary()
```
| github_jupyter |
```
import json
from collections import defaultdict, Counter
import numpy as np
import matplotlib.pyplot as plt
from presidio_evaluator import InputSample
from presidio_evaluator.evaluation import ModelError, Evaluator
from presidio_evaluator.models import PresidioAnalyzerWrapper
from presidio_analyzer import AnalyzerEngine
```
# Evaluate Presidio on the I2B2-2014 de-identification dataset
#### Prerequisites:
1. Get access to the data
2. Copy the data to the `/data/i2b2/2014` folder on the top of the repo. You should have three folders:
- `testing-PHI-Gold-fixed`
- `training-PHI-Gold-Set1`
- `training-PHI-Gold-Set2`
3. Run the following cell for creating a list of InputSamples and save them to json:
```
CREATE_DATASET=False #Change to true on the first run
if CREATE_DATASET:
# Data is assumed to be on the data folder (repo root) under i2b2/2014
# train 1
input_path1 = Path("../data/i2b2/2014/training-PHI-Gold-Set1")
output_path1 = Path("../data/i2b2/2014/training-PHI-Gold-Set1.json")
I2B22014Formatter.dataset_to_json(input_path1, output_path1)
# train 2
input_path2 = Path("../data/i2b2/2014/training-PHI-Gold-Set2")
output_path2 = Path("../data/i2b2/2014/training-PHI-Gold-Set2.json")
I2B22014Formatter.dataset_to_json(input_path2, output_path2)
# test
input_path3 = Path("../data/i2b2/2014/testing-PHI-Gold-fixed")
output_path3 = Path("../data/i2b2/2014/testing-PHI-Gold-fixed.json")
I2B22014Formatter.dataset_to_json(input_path3, output_path3)
def read_json_dataset(filepath=None, length=None):
with open(filepath, "r", encoding="utf-8") as f:
dataset = json.load(f)
if length:
dataset = dataset[:length]
input_samples = [InputSample.from_json(row) for row in dataset]
input_samples = [sample for sample in input_samples if len(sample.full_text) < 5120]
return input_samples
dataset = read_json_dataset("../data/i2b2/2014/training-PHI-Gold-Set1.json")
```
Entity types in this dataset and their frequencies:
```
flatten = lambda l: [item for sublist in l for item in sublist]
count_per_entity = Counter([span.entity_type for span in flatten([input_sample.spans for input_sample in dataset])])
count_per_entity
```
Dataset statistics
```
print(f"Number of samples: {len(dataset)}")
print(f"Total number of tokens: {sum([len(sample.tokens) for sample in dataset])}")
print(f"Average number of tokens: {np.mean([len(sample.tokens) for sample in dataset])}")
print(f"Number of spans: {sum(len(sample.spans) for sample in dataset)}")
print("Sentence length")
fig, axs = plt.subplots(2,figsize=(20,10))
lengths = [len(sample.full_text) for sample in dataset]
axs[0].hist(lengths,color="grey")
axs[0].set_title("Number of characters per sample")
axs[0].set(xlabel="Number of characters",ylabel="Number of samples")
tokens = [len(sample.tokens) for sample in dataset]
axs[1].hist(tokens,)
axs[1].set_title("Number of tokens per sample")
axs[1].set(xlabel="Number of characters",ylabel="Number of samples")
```
Translate I2b2 2014 entity types to Presidio's (If available)
```
i2b2_presidio_dict = {
"PATIENT": "PERSON",
"DOCTOR": "PERSON",
"AGE":"AGE", # Not supported in Presidio
"BIOID": "BIOID", # Not supported in Presidio
"COUNTRY": "LOCATION",
"CITY":"LOCATION",
"DATE": "DATE_TIME",
"DEVICE": "DEVICE", # Not supported in Presidio
"EMAIL": "EMAIL_ADDRESS",
"FAX": "US_PHONE_NUMBER",
"HEALTHPLAN": "HEALTHPLAN", # Not supported in Presidio
"HOSPITAL": "ORGANIZATION",
# "IDNUM": "IDNUM", # Not supported in Presidio
"LOCATION-OTHER": "LOCATION",
# "MEDICALRECORD": "MEDICAL_RECORD", # Not supported in Presidio
"ORGANIZATION": "ORGANIZATION",
"PHONE": "PHONE_NUMBER",
"PROFESSION": "PROFESSION", # Not supported in Presidio
"STATE": "LOCATION",
"STREET": "LOCATION",
"URL": "DOMAIN_NAME",
# "USERNAME": "USERNAME", # Not supported in Presidio
"ZIP": "ZIP", # Not supported in Presidio
"O": "O",
}
```
Examine different entity values
```
values_per_entity = defaultdict(set)
for sample in dataset:
for span in sample.spans:
values_per_entity[span.entity_type].add(span.entity_value)
values_per_entity['PROFESSION']
new_dataset = Evaluator.align_entity_types(input_samples=dataset, entities_mapping=i2b2_presidio_dict,
allow_missing_mappings=True)
```
Re-calculate frequency per entity_type
```
count_per_entity_new = Counter([span.entity_type for span in flatten([input_sample.spans for input_sample in new_dataset])])
count_per_entity_new
# Set up analyzer
analyzer = AnalyzerEngine()
# Run evaluation
presidio = PresidioAnalyzerWrapper(analyzer_engine=analyzer,
entities_to_keep=list(count_per_entity_new.keys()))
evaluator = Evaluator(model=presidio)
evaluated = evaluator.evaluate_all(new_dataset)
evaluation_result = evaluator.calculate_score(evaluated)
evaluation_result.print()
```
Analyze wrong predictions
```
errors = evaluation_result.model_errors
```
False positives analysis
```
ModelError.most_common_fp_tokens(errors,n=5)
ModelError.get_fps_dataframe(errors,entity='DATE_TIME')
```
False negatives analysis
```
ModelError.most_common_fn_tokens(errors,n=5)
ModelError.get_fns_dataframe(errors,entity='DATE_TIME')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/EvenSol/NeqSim-Colab/blob/master/notebooks/process/MultiphaseflowMeasurement.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
#@title Multiphase flow measurements
#@markdown This document is part of the module ["Introduction to Gas Processing using NeqSim in Colab"](https://colab.research.google.com/github/EvenSol/NeqSim-Colab/blob/master/notebooks/examples_of_NeqSim_in_Colab.ipynb#scrollTo=_eRtkQnHpL70).
%%capture
!pip install neqsim
import neqsim
from neqsim.thermo.thermoTools import *
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import math
import numpy
import pandas as pd
%matplotlib inline
```
# Multiphase flow measurement
A multiphase flow meter is a device used to measure the individual phase flow rates of constituent phases in a given flow (for example in oil and gas industry) where oil, water and gas mixtures are initially co-mingled together during the oil production processes.
# Litterature
Wikipedia
https://en.wikipedia.org/wiki/Multiphase_flow_meter
HANDBOOK OF MULTIPHASE FLOW METERING:
https://nfogm.no/wp-content/uploads/2014/02/MPFM_Handbook_Revision2_2005_ISBN-82-91341-89-3.pdf
```
#@title Webinar: Advances in Multiphase Metering for Onshore Measurement in Oil & Gas
#@markdown This video gives an intriduction to Oil & Gas facilities Design
from IPython.display import YouTubeVideo
YouTubeVideo('mcxrjJwidl0', width=600, height=400)
```
# Generation of fluid characterization and generationg PVT properites for multi phase meters
In the following section it will be demonstrated how to generate PVT data for a multi phase flow meter. The steps in generation of PVT tables will be:
1. Collection of fluid composition and PVT data from a PVT report
2. Set up fluid composition based on compositional analysis in PVT report, calulate and compare to measured PVT properties from PVT report
3. Selection of parameters to fit, and fit model to PVT data
4. Gerenation of properties for multi phase flow meter
# 1. Collection of fluid composition and PVT data from PVTbreport. Evaluation of data.
We start by optaining a fluid composition and PVT data from a PVT report. The fluid compositon can typically be reported based on a bottom hole sample or from a test separator. The gas and oil from the sample is analysed, and matehamtical recombined into a well stream composition.
In this example we will use a fluid characterization based on a C10+ analysis. A detailed composition is reported up to C6, and the C6, C7, C8 and C9 is defned as oil fraction components with properties calculated based on molar mass nnd density. C10+ is added with molar mass and density of the 10+ fraction.
```
gascondensate = {'ComponentName': ['nitrogen','CO2', 'methane', 'ethane', 'propane','i-butane','n-butane','i-pentane','n-pentane',"C6", "C7", "C8", "C9", "C10"],
'MolarComposition[-]': [0.972, 0.632, 95.111, 2.553, 0.104, 0.121, 0.021, 0.066, 0.02,0.058, 0.107, 0.073, 0.044, 0.118],
'MolarMass[kg/mol]': [None, None, None, None, None, None, None, None, None, 0.08618, 0.096, 0.107, 0.121, 0.202],
'RelativeDensity[-]': [None, None, None, None, None, None, None, None, None, 664.0e-3, 738.0e-3, 765.0e-3, 781.0e-3, 813.30e-3]}
compositionDataFrame = pd.DataFrame(gascondensate)
print(compositionDataFrame.head(20).to_string())
```
## Data from PVT report
Typical data from a PVT report is reported in the following code. In the following example the fluid composition was reported based on test separarator data, and a constand mass expansion test was done (CME). Reservoir pressure and temperature.
```
reservoirTemperature = 80.6 #Celius
reservoirPressure = 320.8 #bara
testSeparatorTemperature = 20.6 # Celcius
testSeparatorPressure = 86.8 # bara
GORseparartorConditions = 58959.0 # Sm3 gas//m3 condesate
GORstandardConditions = 55000.0 # Sm3 gas/Sm3 condesate
# Define a dictionary with PVT data
PVTdata = {'pressure': [555.3,552,518.5,484,449.5,415.3,408.1,401.3,394.3,387.3,380.7,373.8,366.7,360,352.9,346.2,339.3,332.3,329.9,325.6,320,315.1,308.4,301.3,294.5,287.6,280.6,273.8,266.9,259.9,253.1,249.5,246.1,242.7,208.2,173.6,139,104.5,70,46.1],
'relative volume': [0.741,0.7431,0.766,0.7917,0.821,0.8551,0.8631,0.871,0.8795,0.8883,0.8972,0.9066,0.9169,0.9272,0.9385,0.9498,0.9621,0.9751,0.9797,0.9883,1,1.0105,1.0258,1.0428,1.0602,1.0787,1.0989,1.1197,1.1423,1.1668,1.1924,1.2064,1.2202,1.2346,1.4135,1.6806,2.1038,2.8422,4.3612,6.796],
'Zgas': [1.247,1.244,1.204,1.162,1.119,1.077,1.068,1.06,1.051,1.043,1.035,1.027,1.019,1.012,1.004,0.997,0.99,0.982,0.98,0.976,0.97,0.965,0.959,0.952,0.946,0.941,0.935,0.929,0.924,0.919,0.915,0.912,0.91,0.908,0.892,0.884,0.887,0.9,0.926,0.95],
'Density': [0.2671,0.2663,0.2583,0.2499,0.241,0.2314,0.2293,0.2272,0.225,0.2227,0.2206,0.2183,0.2158,0.2134,0.2108,0.2083,0.2057,0.2029,0.202,0.2002,0.1979,0.1958,0.1929,0.1898,0.1866,0.1834,0.1801,0.1767,0.1732,0.1696,0.1659,0.164,0.1622,0.1603,0.14,0.1177,0.0941,0.0696,0.0454,0.0291],
'Bg': [0.0027,0.0027,0.0028,0.0029,0.003,0.0032,0.0032,0.0032,0.0032,0.0033,0.0033,0.0033,0.0034,0.0034,0.0035,0.0035,0.0036,0.0036,0.0036,0.0036,0.0037,0.0037,0.0038,0.0038,0.0039,0.004,0.0041,0.0041,0.0042,0.0043,0.0044,0.0045,0.0045,0.0046,0.0052,0.0062,0.0078,0.0105,0.0161,0.0251],
'gasexpansionfactor': [365.6,364.6,353.7,342.2,330,316.8,313.9,311,308,305,302,298.8,295.5,292.2,288.7,285.2,281.6,277.8,276.5,274.1,270.9,268.1,264.1,259.8,255.5,251.1,246.5,241.9,237.2,232.2,227.2,224.6,222,219.4,191.7,161.2,128.8,95.3,62.1,39.9],
'gasviscosity': [0.0325,0.0324,0.0313,0.0301,0.0288,0.0276,0.0273,0.027,0.0268,0.0265,0.0262,0.0259,0.0257,0.0254,0.0251,0.0248,0.0245,0.0243,0.0242,0.024,0.0237,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0,0]
}
# Convert the PVT data dictionary into DataFrame
CMEdataFrame = pd.DataFrame(PVTdata)
CMEpressures = CMEdataFrame['pressure'].tolist()
CMEtemperature = [80.6+273.15]*len(CMEpressures)
#print and plot the PVT data
print(CMEdataFrame.head(50).to_string())
print("plotting experimental PVT data.....")
CMEdataFrame.plot(kind='scatter',x='relative volume',y='pressure',color='red')
plt.show()
```
# 2. Set up fluid based on fluid composition and properties from PVT report, calculate and compare to measured PVT properties from PVT report
A fluid in neqsim is set up in the following script. We the flash the fluid to calculate the GOR at test separator and standard conditions. Further we will plot the phase envelope of the fluid.
## Setting up the fluid based on the PVT report
In the follwing script we create a neqsim fluid based on composition and data for oil components (molar mass and density) from the PVT report. We will then calcuate the phase envelope based on this fluid characterization.
```
# Running PVTsimulation with default fluid
characterizedFluid = fluid_df(compositionDataFrame, lastIsPlusFraction=True)
print('phase envelope for characterized fluid')
phaseenvelope(characterizedFluid, True)
```
## Compare neqsim calculations to data from PVT report
In the folllowing script we run neqsim simulations to compare how well the PVT data from the PVT report can be represented by the default fluid characterization. NeqSim will by default use SRK-EoS with Peneloux volume correction.
We start by comparing to measured GOR at separator and standard conditions.
```
characterizedFluid.setTemperature(15.0, "C")
characterizedFluid.setPressure(1.0, "atm")
TPflash(characterizedFluid)
#printFLuidCharacterisation(characterizedFluid) #print componentnames, TC, PC, acs, molar mass, density,
printFrame(characterizedFluid)
GORcalcstd = characterizedFluid.getPhase("gas").getNumberOfMolesInPhase()*8.314*288.15/101325 / (characterizedFluid.getPhase("oil").getVolume("m3"))
print("GOR at standard conditions ", GORcalcstd, " Sm3 gas/m3 oil. ", " Deviation from PVT report: ", (GORcalcstd-GORstandardConditions)/GORstandardConditions*100, " %")
characterizedFluid.setTemperature(testSeparatorTemperature, "C")
characterizedFluid.setPressure(testSeparatorPressure, "bara")
TPflash(characterizedFluid)
GORcalc = characterizedFluid.getPhase("gas").getNumberOfMolesInPhase()*8.314*288.15/101325 / (characterizedFluid.getPhase("oil").getVolume("m3"))
print("GOR at test separator conditions: ", GORcalc, " Sm3 gas/m3 oil" , " Deviation from PVT report: ", (GORcalc-GORseparartorConditions)/GORseparartorConditions*100, " %")
#Calculating saturation pressure
#characterizedFluid.setTemperature(reservoirTemperature, "C")
#calcSatPres = saturationpressure(characterizedFluid)
#print("Saturation pressure : ", calcSatPres, " [bara]" , " Deviation from PVT report: ", (calcSatPres-reservoirPressure), " bar")
```
## Comparing to PVT data
In this case CME (constant mass expansion) experiments have been performed, and we will compare to these data.
```
simrelativevolume = []
simliquidrelativevolume = []
Zgas = []
Bgsim=[]
densitysim = []
Yfactor = []
isothermalcompressibility = []
gasviscositysim = []
saturationPressure = None
CME(characterizedFluid,CMEpressures,CMEtemperature,saturationPressure,simrelativevolume, simliquidrelativevolume,Zgas,Yfactor,isothermalcompressibility,densitysim,Bgsim, gasviscositysim)
CMEsimdataFrame = pd.DataFrame(numpy.transpose([CMEpressures, simrelativevolume,Zgas, densitysim, Bgsim, gasviscositysim]), columns=["pressure", "sim relative volume","Zgassim", "densitysim", "Bgsim", "gasviscositysim"])
print(CMEsimdataFrame.head(50).to_string())
print("saturation pressure simulated ", saturationPressure)
pd.concat([CMEdataFrame['pressure'], CMEdataFrame['relative volume'],CMEsimdataFrame['sim relative volume']],axis=1).plot(x='pressure')
pd.concat([CMEdataFrame['pressure'], CMEdataFrame['Zgas'],CMEsimdataFrame['Zgassim']],axis=1).plot(x='pressure')
pd.concat([CMEdataFrame['pressure'], CMEdataFrame['Density']*1e3,CMEsimdataFrame['densitysim']],axis=1).plot(x='pressure')
pd.concat([CMEdataFrame['pressure'], CMEdataFrame['Bg'],CMEsimdataFrame['Bgsim']],axis=1).plot(x='pressure')
pd.concat([CMEdataFrame['pressure'], CMEdataFrame['gasviscosity'],CMEsimdataFrame['gasviscositysim']*1e3],axis=1).plot(x='pressure',kind="line")
matplotlib.pyplot.show()
devanalysisframe = pd.concat([CMEdataFrame['pressure'], (CMEsimdataFrame['sim relative volume']-CMEdataFrame['relative volume'])/CMEdataFrame['relative volume']*100.0],axis=1)
print("Deviation analysis...")
print("Average deviation relative volume: ", devanalysisframe[0].mean(), " %", " Max devation ", devanalysisframe[0].max() , " %")
```
# 3. Selection of parameters to fit, and fit model to PVT data
To be done...
# 4. Gerenation of properties for multi phase flow meter
A typical multi phase flow meter will need calclation of various thermodynamic and physical properties. Such properties have to be updated as the field is produced. The input to the property calculations are a characterized fluid composition from PVT studies.
The following script demonstrates calculation of PVT properties for a multi phase flow meter using a characterized fluid composition.
```
#Creating property tables
pressures = [150.0, 170.0, 180.0, 200.0, 270.0]
temperatures = [30.0, 40.0, 50.0, 60.0, 80.0]
numP = len(pressures)
numT = len(temperatures)
gasViscosity = numpy.zeros((numP, numT))
oilViscosity = numpy.zeros((numP, numT))
gasDensity = numpy.zeros((numP, numT))
oilDensity = numpy.zeros((numP, numT))
GORcalc = numpy.zeros((numP, numT))
GORactual = numpy.zeros((numP, numT))
surfaceTension = numpy.zeros((numP, numT))
gasViscosity[:] = np.NaN
oilViscosity[:] = np.NaN
gasDensity[:] = np.NaN
oilDensity[:] = np.NaN
GORcalc[:] = np.NaN
GORactual[:] = np.NaN
surfaceTension[:] = np.NaN
for i in range(len(temperatures)):
for j in range(len(pressures)):
characterizedFluid.setPressure(pressures[j])
characterizedFluid.setTemperature(temperatures[i]+273.15)
TPflash(characterizedFluid)
characterizedFluid.initProperties()
if(characterizedFluid.hasPhaseType("gas")):
gasViscosity[j][i]=characterizedFluid.getPhase("gas").getViscosity("cP")
gasDensity[j][i]=characterizedFluid.getPhase("gas").getDensity("kg/m3")
if(characterizedFluid.hasPhaseType("oil")):
oilViscosity[j][i]=characterizedFluid.getPhase("oil").getViscosity("cP")
oilDensity[j][i]=characterizedFluid.getPhase("oil").getDensity("kg/m3")
if(characterizedFluid.hasPhaseType("gas") and characterizedFluid.hasPhaseType("oil")):
GORcalc[j][i] = characterizedFluid.getPhase("gas").getNumberOfMolesInPhase()*8.314*288.15/101325 / (characterizedFluid.getPhase("oil").getVolume("m3"))
GORactual[j][i] = (characterizedFluid.getPhase("gas").getVolume("m3"))/ (characterizedFluid.getPhase("oil").getVolume("m3"))
surfaceTension[j][i] = characterizedFluid.getInterfacialTension('gas', 'oil')
gasDensityDataFrame = pd.DataFrame(gasDensity,index=pressures, columns=temperatures)
oilDensityDataFrame = pd.DataFrame(oilDensity,index=pressures, columns=temperatures)
gasviscosityDataFrame = pd.DataFrame(gasViscosity,index=pressures, columns=temperatures)
oilviscosityDataFrame = pd.DataFrame(oilViscosity,index=pressures, columns=temperatures)
GORcalcFrame = pd.DataFrame(GORcalc,index=pressures, columns=temperatures)
GORactualFrame = pd.DataFrame(GORactual,index=pressures, columns=temperatures)
surfaceTensionFrame= pd.DataFrame(surfaceTension,index=pressures, columns=temperatures)
print("gas density [kg/m3]")
print(gasDensityDataFrame.tail())
print("oil density [kg/m3]")
print(oilDensityDataFrame.head())
print("gas viscosity [(mPa.s)]")
print(gasviscosityDataFrame.tail())
print("oil viscosity [(mPa.s)]")
print(oilviscosityDataFrame.head())
#print("GOR (Sm3/m3)")
#print(GORcalcFrame.head())
print("GOR actual (m3/m3)")
print(GORactualFrame.head())
print("Surface Tension (N/m)")
print(surfaceTensionFrame.head())
```
| github_jupyter |
# Student Exam Performance Workshop
Use Python's pandas module to load a dataset containing student demographic information and test scores and find relationships between student attributes and test scores. This workshop will serve as an introduction to pandas and will allow students to practice the following skills:
- Load a csv into a pandas DataFrame and examine summary statistics
- Rename DataFrame column names
- Add columns to a DataFrame
- Change values in DataFrame rows
- Analyze relationships between categorical features and test scores
**Bonus:**
Determine the relationship between the students' lunch classification and average test scores by creating a seaborn boxplot
```
# Import the python modules that we will need to use
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
def load_data(path):
df = pd.read_csv(path)
return df
```
Use the `load_data` function to load the StudentsPerformance.csv file into a pandas dataframe variable called `df`
__Hint__: Keep in mind where the csv file is in relation to this Jupyter Notebook. Do you need to provide an absolute or relative file path?
```
#Write python to call the function above and load the StudentPeformance csv file into a pandas dataframe
#Keep this line so you can see the first five rows of your dataframe once you have loaded it!
df.head(5)
```
__Next step:__ Now that we have loaded our DataFrame, let's look at the summary statistics of our data. We can use the `describe` method to accomplish this:
```
df.describe(include='all')
```
By looking at this breakdown of our dataset, I can make at least the following observations:
1. Our DataFrame consists of eight columns, three of which are student test scores.
2. There are no missing any values in our DataFrame!
3. The data appears to be pretty evenly distributed.
4. The column names are long and difficult to type
## Renaming DataFrame Columns
Let's change our column names so they are easier to work with!
__Hint__: Look into the pandas `columns` attribute to make the change!
```
columns = 'gender', 'race', 'parentDegree', 'lunchStatus', 'courseCompletion', 'mathScore', 'readingScore', 'writingScore'
def renameColumns(df, columns):
df.columns=columns
return df
#Use the above function to rename the DataFrame's column names
df.head(10) #Look at the first ten rows of the DataFrame to ensure the renaming worked!
```
## Adding Columns to a DataFrame
Great! Next we want to add an `avgScore` column that is an average of the three given test scores (`mathScore`, `readingScore` and `writingScore`). This will allow us to generalize the students' performance and simplify the process of us examining our feature's impact on student performance.
```
#Complete the following line of code to create an avgScore column
df['avgScore'] =
df.head(5)
```
## Analyzing Feature Relationships
Now that our data is looking the way we want, let's examine how some of our features correlate with students' test performances. We will start by looking at the relationship between race and parent degree status on test scores.
__Hint__: Use pandas' `groupby` method to examine these relationships. The documentation for `groupby` can be found here: https://pandas.pydata.org/pandas-docs/version/0.22/generated/pandas.DataFrame.groupby.html
```
df.groupby(['race','parentDegree']).mean()
```
From examining the above output, we can see that across all `race` groups, students with "high school" and "some high school" as their parent degree status (`parentDegree`) had lower test scores.
__Next step__: Since there seems to be a clear distinction between students that have parents with have some college education and those that do not, let's simplify our DataFrame by creating a `degreeBinary` column based on values in the `parentDegree` column. This new column will simply contain either "no_degree" or "has_degree." We can do this by writing a basic function and using pandas' `apply` method:
```
#Complete this function to return the proper strings to denote degree status
def degree_status(edu):
if edu in {'high school', 'some high school'}:
#Fill in your code here!
df['degreeBinary'] = df['parentDegree'].apply(degree_status)
df.head(10)
```
Great job! Now let's continue examining our features to find relationships in our data
__Your turn:__ Use the `groupby` function again examine relationships between other features and student test scores. What can we learn about the relationship between these whether or not the students have completed the course and their test scores? What about the relationship between gender and test scores?
```
##Use groupby to examine the relationship between course completion status and test scores
##Use groupby to examine the relationship between gender and test scores
```
## Bonus: Visualization
Great job making it this far! As a bonus exercise, we will create a simple data visualization. We have examined the relationship between all of our features and student test scores except for one -- student lunch status, which is found in the `lunch` column.
In order to explore this relationship, let's create a `barplot`, with the students'`lunch` status as the x-axis and their average test scores (`avgScore`) as the y-axis.
We will use seaborn, which is a third-party library, to complete this visualization. If you do not already have seaborn installed, `pip install` it now! Follow the seaborn documentation to create the `barplot` in the cell below.
```
import seaborn as sns #import the seaborn module
sns.set(style='whitegrid')
def graph_data(data, xkey='lunchStatus', ykey='avgScore'):
#Fill this in to create the barplot!
graph_data(df)
```
| github_jupyter |
# Sentiment Analysis with an RNN
In this notebook, you'll implement a recurrent neural network that performs sentiment analysis. Using an RNN rather than a feedfoward network is more accurate since we can include information about the *sequence* of words. Here we'll use a dataset of movie reviews, accompanied by labels.
The architecture for this network is shown below.
<img src="assets/network_diagram.png" width=400px>
Here, we'll pass in words to an embedding layer. We need an embedding layer because we have tens of thousands of words, so we'll need a more efficient representation for our input data than one-hot encoded vectors. You should have seen this before from the word2vec lesson. You can actually train up an embedding with word2vec and use it here. But it's good enough to just have an embedding layer and let the network learn the embedding table on it's own.
From the embedding layer, the new representations will be passed to LSTM cells. These will add recurrent connections to the network so we can include information about the sequence of words in the data. Finally, the LSTM cells will go to a sigmoid output layer here. We're using the sigmoid because we're trying to predict if this text has positive or negative sentiment. The output layer will just be a single unit then, with a sigmoid activation function.
We don't care about the sigmoid outputs except for the very last one, we can ignore the rest. We'll calculate the cost from the output of the last step and the training label.
```
import numpy as np
import tensorflow as tf
with open('../sentiment-network/reviews.txt', 'r') as f:
reviews = f.read()
with open('../sentiment-network/labels.txt', 'r') as f:
labels = f.read()
reviews[:2000]
```
## Data preprocessing
The first step when building a neural network model is getting your data into the proper form to feed into the network. Since we're using embedding layers, we'll need to encode each word with an integer. We'll also want to clean it up a bit.
You can see an example of the reviews data above. We'll want to get rid of those periods. Also, you might notice that the reviews are delimited with newlines `\n`. To deal with those, I'm going to split the text into each review using `\n` as the delimiter. Then I can combined all the reviews back together into one big string.
First, let's remove all punctuation. Then get all the text without the newlines and split it into individual words.
```
from string import punctuation
all_text = ''.join([c for c in reviews if c not in punctuation])
reviews = all_text.split('\n')
all_text = ' '.join(reviews)
words = all_text.split()
all_text[:2000]
words[:100]
```
### Encoding the words
The embedding lookup requires that we pass in integers to our network. The easiest way to do this is to create dictionaries that map the words in the vocabulary to integers. Then we can convert each of our reviews into integers so they can be passed into the network.
> **Exercise:** Now you're going to encode the words with integers. Build a dictionary that maps words to integers. Later we're going to pad our input vectors with zeros, so make sure the integers **start at 1, not 0**.
> Also, convert the reviews to integers and store the reviews in a new list called `reviews_ints`.
```
from collections import Counter
counts = Counter(words)
vocab = sorted(counts, key=counts.get, reverse=True)
vocab_to_int = {word: ii for ii, word in enumerate(vocab, 1)}
reviews_ints = []
for each in reviews:
reviews_ints.append([vocab_to_int[word] for word in each.split()])
```
### Encoding the labels
Our labels are "positive" or "negative". To use these labels in our network, we need to convert them to 0 and 1.
> **Exercise:** Convert labels from `positive` and `negative` to 1 and 0, respectively.
```
labels = labels.split('\n')
labels = np.array([1 if each == 'positive' else 0 for each in labels])
review_lens = Counter([len(x) for x in reviews_ints])
print("Zero-length reviews: {}".format(review_lens[0]))
print("Maximum review length: {}".format(max(review_lens)))
```
Okay, a couple issues here. We seem to have one review with zero length. And, the maximum review length is way too many steps for our RNN. Let's truncate to 200 steps. For reviews shorter than 200, we'll pad with 0s. For reviews longer than 200, we can truncate them to the first 200 characters.
> **Exercise:** First, remove the review with zero length from the `reviews_ints` list.
```
non_zero_idx = [ii for ii, review in enumerate(reviews_ints) if len(review) != 0]
len(non_zero_idx)
reviews_ints[-1]
```
Turns out its the final review that has zero length. But that might not always be the case, so let's make it more general.
```
reviews_ints = [reviews_ints[ii] for ii in non_zero_idx]
labels = np.array([labels[ii] for ii in non_zero_idx])
```
> **Exercise:** Now, create an array `features` that contains the data we'll pass to the network. The data should come from `review_ints`, since we want to feed integers to the network. Each row should be 200 elements long. For reviews shorter than 200 words, left pad with 0s. That is, if the review is `['best', 'movie', 'ever']`, `[117, 18, 128]` as integers, the row will look like `[0, 0, 0, ..., 0, 117, 18, 128]`. For reviews longer than 200, use on the first 200 words as the feature vector.
This isn't trivial and there are a bunch of ways to do this. But, if you're going to be building your own deep learning networks, you're going to have to get used to preparing your data.
```
seq_len = 200
features = np.zeros((len(reviews_ints), seq_len), dtype=int)
for i, row in enumerate(reviews_ints):
features[i, -len(row):] = np.array(row)[:seq_len]
features[:10,:100]
```
## Training, Validation, Test
With our data in nice shape, we'll split it into training, validation, and test sets.
> **Exercise:** Create the training, validation, and test sets here. You'll need to create sets for the features and the labels, `train_x` and `train_y` for example. Define a split fraction, `split_frac` as the fraction of data to keep in the training set. Usually this is set to 0.8 or 0.9. The rest of the data will be split in half to create the validation and testing data.
```
split_frac = 0.8
split_idx = int(len(features)*0.8)
train_x, val_x = features[:split_idx], features[split_idx:]
train_y, val_y = labels[:split_idx], labels[split_idx:]
test_idx = int(len(val_x)*0.5)
val_x, test_x = val_x[:test_idx], val_x[test_idx:]
val_y, test_y = val_y[:test_idx], val_y[test_idx:]
print("\t\t\tFeature Shapes:")
print("Train set: \t\t{}".format(train_x.shape),
"\nValidation set: \t{}".format(val_x.shape),
"\nTest set: \t\t{}".format(test_x.shape))
```
With train, validation, and text fractions of 0.8, 0.1, 0.1, the final shapes should look like:
```
Feature Shapes:
Train set: (20000, 200)
Validation set: (2500, 200)
Test set: (2500, 200)
```
## Build the graph
Here, we'll build the graph. First up, defining the hyperparameters.
* `lstm_size`: Number of units in the hidden layers in the LSTM cells. Usually larger is better performance wise. Common values are 128, 256, 512, etc.
* `lstm_layers`: Number of LSTM layers in the network. I'd start with 1, then add more if I'm underfitting.
* `batch_size`: The number of reviews to feed the network in one training pass. Typically this should be set as high as you can go without running out of memory.
* `learning_rate`: Learning rate
```
lstm_size = 256
lstm_layers = 1
batch_size = 500
learning_rate = 0.001
```
For the network itself, we'll be passing in our 200 element long review vectors. Each batch will be `batch_size` vectors. We'll also be using dropout on the LSTM layer, so we'll make a placeholder for the keep probability.
> **Exercise:** Create the `inputs_`, `labels_`, and drop out `keep_prob` placeholders using `tf.placeholder`. `labels_` needs to be two-dimensional to work with some functions later. Since `keep_prob` is a scalar (a 0-dimensional tensor), you shouldn't provide a size to `tf.placeholder`.
```
n_words = len(vocab_to_int) + 1 # Adding 1 because we use 0's for padding, dictionary started at 1
# Create the graph object
graph = tf.Graph()
# Add nodes to the graph
with graph.as_default():
inputs_ = tf.placeholder(tf.int32, [None, None], name='inputs')
labels_ = tf.placeholder(tf.int32, [None, None], name='labels')
keep_prob = tf.placeholder(tf.float32, name='keep_prob')
```
### Embedding
Now we'll add an embedding layer. We need to do this because there are 74000 words in our vocabulary. It is massively inefficient to one-hot encode our classes here. You should remember dealing with this problem from the word2vec lesson. Instead of one-hot encoding, we can have an embedding layer and use that layer as a lookup table. You could train an embedding layer using word2vec, then load it here. But, it's fine to just make a new layer and let the network learn the weights.
> **Exercise:** Create the embedding lookup matrix as a `tf.Variable`. Use that embedding matrix to get the embedded vectors to pass to the LSTM cell with [`tf.nn.embedding_lookup`](https://www.tensorflow.org/api_docs/python/tf/nn/embedding_lookup). This function takes the embedding matrix and an input tensor, such as the review vectors. Then, it'll return another tensor with the embedded vectors. So, if the embedding layer as 200 units, the function will return a tensor with size [batch_size, 200].
```
# Size of the embedding vectors (number of units in the embedding layer)
embed_size = 300
with graph.as_default():
embedding = tf.Variable(tf.random_uniform((n_words, embed_size), -1, 1))
embed = tf.nn.embedding_lookup(embedding, inputs_)
```
### LSTM cell
<img src="assets/network_diagram.png" width=400px>
Next, we'll create our LSTM cells to use in the recurrent network ([TensorFlow documentation](https://www.tensorflow.org/api_docs/python/tf/contrib/rnn)). Here we are just defining what the cells look like. This isn't actually building the graph, just defining the type of cells we want in our graph.
To create a basic LSTM cell for the graph, you'll want to use `tf.contrib.rnn.BasicLSTMCell`. Looking at the function documentation:
```
tf.contrib.rnn.BasicLSTMCell(num_units, forget_bias=1.0, input_size=None, state_is_tuple=True, activation=<function tanh at 0x109f1ef28>)
```
you can see it takes a parameter called `num_units`, the number of units in the cell, called `lstm_size` in this code. So then, you can write something like
```
lstm = tf.contrib.rnn.BasicLSTMCell(num_units)
```
to create an LSTM cell with `num_units`. Next, you can add dropout to the cell with `tf.contrib.rnn.DropoutWrapper`. This just wraps the cell in another cell, but with dropout added to the inputs and/or outputs. It's a really convenient way to make your network better with almost no effort! So you'd do something like
```
drop = tf.contrib.rnn.DropoutWrapper(cell, output_keep_prob=keep_prob)
```
Most of the time, you're network will have better performance with more layers. That's sort of the magic of deep learning, adding more layers allows the network to learn really complex relationships. Again, there is a simple way to create multiple layers of LSTM cells with `tf.contrib.rnn.MultiRNNCell`:
```
cell = tf.contrib.rnn.MultiRNNCell([drop] * lstm_layers)
```
Here, `[drop] * lstm_layers` creates a list of cells (`drop`) that is `lstm_layers` long. The `MultiRNNCell` wrapper builds this into multiple layers of RNN cells, one for each cell in the list.
So the final cell you're using in the network is actually multiple (or just one) LSTM cells with dropout. But it all works the same from an achitectural viewpoint, just a more complicated graph in the cell.
> **Exercise:** Below, use `tf.contrib.rnn.BasicLSTMCell` to create an LSTM cell. Then, add drop out to it with `tf.contrib.rnn.DropoutWrapper`. Finally, create multiple LSTM layers with `tf.contrib.rnn.MultiRNNCell`.
Here is [a tutorial on building RNNs](https://www.tensorflow.org/tutorials/recurrent) that will help you out.
```
with graph.as_default():
# Your basic LSTM cell
lstm = tf.contrib.rnn.BasicLSTMCell(lstm_size)
# Add dropout to the cell
drop = tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
# Stack up multiple LSTM layers, for deep learning
cell = tf.contrib.rnn.MultiRNNCell([drop] * lstm_layers)
# Getting an initial state of all zeros
initial_state = cell.zero_state(batch_size, tf.float32)
```
### RNN forward pass
<img src="assets/network_diagram.png" width=400px>
Now we need to actually run the data through the RNN nodes. You can use [`tf.nn.dynamic_rnn`](https://www.tensorflow.org/api_docs/python/tf/nn/dynamic_rnn) to do this. You'd pass in the RNN cell you created (our multiple layered LSTM `cell` for instance), and the inputs to the network.
```
outputs, final_state = tf.nn.dynamic_rnn(cell, inputs, initial_state=initial_state)
```
Above I created an initial state, `initial_state`, to pass to the RNN. This is the cell state that is passed between the hidden layers in successive time steps. `tf.nn.dynamic_rnn` takes care of most of the work for us. We pass in our cell and the input to the cell, then it does the unrolling and everything else for us. It returns outputs for each time step and the final_state of the hidden layer.
> **Exercise:** Use `tf.nn.dynamic_rnn` to add the forward pass through the RNN. Remember that we're actually passing in vectors from the embedding layer, `embed`.
```
with graph.as_default():
outputs, final_state = tf.nn.dynamic_rnn(cell, embed,
initial_state=initial_state)
```
### Output
We only care about the final output, we'll be using that as our sentiment prediction. So we need to grab the last output with `outputs[:, -1]`, the calculate the cost from that and `labels_`.
```
with graph.as_default():
predictions = tf.contrib.layers.fully_connected(outputs[:, -1], 1, activation_fn=tf.sigmoid)
cost = tf.losses.mean_squared_error(labels_, predictions)
optimizer = tf.train.AdamOptimizer(learning_rate).minimize(cost)
```
### Validation accuracy
Here we can add a few nodes to calculate the accuracy which we'll use in the validation pass.
```
with graph.as_default():
correct_pred = tf.equal(tf.cast(tf.round(predictions), tf.int32), labels_)
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
```
### Batching
This is a simple function for returning batches from our data. First it removes data such that we only have full batches. Then it iterates through the `x` and `y` arrays and returns slices out of those arrays with size `[batch_size]`.
```
def get_batches(x, y, batch_size=100):
n_batches = len(x)//batch_size
x, y = x[:n_batches*batch_size], y[:n_batches*batch_size]
for ii in range(0, len(x), batch_size):
yield x[ii:ii+batch_size], y[ii:ii+batch_size]
```
## Training
Below is the typical training code. If you want to do this yourself, feel free to delete all this code and implement it yourself. Before you run this, make sure the `checkpoints` directory exists.
```
epochs = 10
with graph.as_default():
saver = tf.train.Saver()
with tf.Session(graph=graph) as sess:
sess.run(tf.global_variables_initializer())
iteration = 1
for e in range(epochs):
state = sess.run(initial_state)
for ii, (x, y) in enumerate(get_batches(train_x, train_y, batch_size), 1):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 0.5,
initial_state: state}
loss, state, _ = sess.run([cost, final_state, optimizer], feed_dict=feed)
if iteration%5==0:
print("Epoch: {}/{}".format(e, epochs),
"Iteration: {}".format(iteration),
"Train loss: {:.3f}".format(loss))
if iteration%25==0:
val_acc = []
val_state = sess.run(cell.zero_state(batch_size, tf.float32))
for x, y in get_batches(val_x, val_y, batch_size):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 1,
initial_state: val_state}
batch_acc, val_state = sess.run([accuracy, final_state], feed_dict=feed)
val_acc.append(batch_acc)
print("Val acc: {:.3f}".format(np.mean(val_acc)))
iteration +=1
saver.save(sess, "checkpoints/sentiment.ckpt")
```
## Testing
```
test_acc = []
with tf.Session(graph=graph) as sess:
saver.restore(sess, tf.train.latest_checkpoint('checkpoints'))
test_state = sess.run(cell.zero_state(batch_size, tf.float32))
for ii, (x, y) in enumerate(get_batches(test_x, test_y, batch_size), 1):
feed = {inputs_: x,
labels_: y[:, None],
keep_prob: 1,
initial_state: test_state}
batch_acc, test_state = sess.run([accuracy, final_state], feed_dict=feed)
test_acc.append(batch_acc)
print("Test accuracy: {:.3f}".format(np.mean(test_acc)))
```
| github_jupyter |
# Introduction to Data Science
# Lecture 13: Linear Regression 2
*COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
In this lecture, we'll discuss:
* overfitting, model generalizability, and the bias-variance tradeoff
* cross validation
* using categorical variables for regression
Recommended reading:
* G. James, D. Witten, T. Hastie, and R. Tibshirani, An Introduction to Statistical Learning, Ch. 3 [digitial version available here](http://www-bcf.usc.edu/~gareth/ISL/)
## Review from Lecture 9 (Linear Regression 1)
### Simple Linear Regression (SLR)
**Data**: We have $n$ samples $(x, y)_i$, $i=1,\ldots n$.
**Model**: $y \sim \beta_0 + \beta_1 x$
**Goal**: Find the best values of $\beta_0$ and $\beta_1$, denoted $\hat{\beta}_0$ and $\hat{\beta}_1$, so that the prediction $y = \hat{\beta}_0 + \hat{\beta}_1 x$ "best fits" the data.
<img src="438px-Linear_regression.png" width="40%" alt="https://en.wikipedia.org/wiki/Linear_regression">
**Theorem.**
The parameters that minimize the "residual sum of squares (RSS)",
$RSS = \sum_i (y_i - \beta_0 - \beta_1 x_i)^2$,
are:
$$
\hat{\beta}_1 = \frac{\sum_{i=1}^n (x_i - \overline{x})(y_i - \overline{y}) }{\sum_{i=1}^n (x_i - \overline{x})^2}
\qquad \textrm{and} \qquad
\hat{\beta}_0 = \overline{y} - \hat{\beta}_1 \overline{x}.
$$
where $\overline{x} = \frac{1}{n} \sum_{i=1}^n x_i$ and $\overline{y} = \frac{1}{n} \sum_{i=1}^n y_i$.
### Multilinear regression
**Data**: We have $n$ samples of the form $\big(x_1, x_2 , \ldots, x_m , y \big)_i$, $i=1,\ldots n$.
**Model**: $y \sim \beta_0 + \beta_1 x_1 + \cdots + \beta_m x_m $
### Nonlinear relationships
**Data**: We have $n$ samples $\big(x_1, x_2 , \ldots, x_m , y \big)_i$, $i=1,\ldots n$.
**Model**: $y \sim \beta_0 + \beta_1 f_1(x_1,x_2,\ldots,x_m) + \cdots + \beta_k f_k(x_1,x_2,\ldots,x_m)$
## Regression with python
There are several different python packages that do regression:
1. [statsmodels](http://statsmodels.sourceforge.net/)
+ [scikit-learn](http://scikit-learn.org/)
+ [SciPy](http://www.scipy.org/)
+ ...
Last time, I commented that statsmodels approaches regression from a statistics viewpoint, while scikit-learn approaches from a machine learning viewpoint. I'll say more about this today.
SciPy has some regression tools, but compared to these other two packages, they are relatively limited.
```
# imports and setup
import scipy as sc
import pandas as pd
import statsmodels.formula.api as sm
from sklearn import linear_model
import matplotlib.pyplot as plt
%matplotlib inline
plt.rcParams['figure.figsize'] = (10, 6)
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
```
## Advertisement dataset
Consider the 'Advertising' dataset from
[here](http://www-bcf.usc.edu/~gareth/ISL/data.html).
For 200 different ‘markets’ (think different cities), this dataset consists of the number of sales of a particular product as well as the advertising budget for three different media: TV, radio, and newspaper.
Last time, after trying a variety of linear models, we discovered the following one, which includes a nonlinear relationship between the TV budget and Radio budget:
$$
\text{Sales} = \beta_0 + \beta_1 * \text{TV_budget} + \beta_2*\text{Radio_budget} + \beta_3 * \text{TV_budget} *\text{Radio_budget}.
$$
```
advert = pd.read_csv('Advertising.csv',index_col=0) #load data
ad_NL = sm.ols(formula="Sales ~ TV + Radio + TV*Radio", data=advert).fit()
ad_NL.summary()
```
This model is really excellent:
- $R^2 = 97\%$ of the variability in the data is accounted for by the model.
- The $p$-value for the F-statistic is very small
- The $p$-values for the individual coefficients are small
Interpretation:
- In a particular market, if I spend an additional $1k on TV advertising, what do I expect sales to do?
- Should I spend additional money on TV or Radio advertising?
```
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.scatter(xs=advert['TV'], ys=advert['Radio'], zs=advert['Sales'])
x = sc.linspace(advert['TV'].min(), advert['TV'].max(), 100)
y = sc.linspace(advert['Radio'].min(), advert['Radio'].max(), 100)
X,Y = sc.meshgrid(x,y)
par = dict(ad_NL.params)
Z = par["Intercept"] + par["TV"]*X + par["Radio"]*Y + par["TV:Radio"]*X*Y
surf = ax.plot_surface(X, Y, Z,cmap=cm.Greys, alpha=0.2)
ax.view_init(25,-71)
ax.set_xlabel('TV budget')
ax.set_ylabel('Radio budget')
ax.set_zlabel('Sales')
plt.show()
```
### A word of caution on overfitting
It is tempting to include a lot of terms in the regression, but this is problematic. A useful model will *generalize* beyond the data given to it.
**Questions?**
## Overfitting, underfitting, model generalizability, and the bias–variance tradeoff
In regression, and other prediction problems, we would like to develop a model on a dataset, that would preform well, not only on that dataset, but on similar data that the model hasn't yet seen by the model. If a model satisfies this criterion, we say that it is *generalizable*.
Consider the following data, that has been fit with a linear polynomial model (black) and a high degree polynomial model (blue). For convenience, let me call these the black and blue models, respectively.
<img src="overfitted_data.png" title="https://commons.wikimedia.org/w/index.php?curid=47471056" width="40%">
Let's call the dataset that we train the model on the *training dataset* and the dataset that we test the model on the *testing dataset*. In the above figure, the training dataset are the black points and the testing dataset is not shown, but we imagine it to be similar to the points shown.
Which model is better?
The blue model has 100% accuracy on the training dataset, while the black model has much smaller accuracy. However, the blue model is highly oscillatory and might not generalize well to new data. For example, the model would wildly miss the test point $(3,0)$. We say that the blue model has *overfit* the data. On the other hand, it isn't difficult to see that we could also *underfit* the data. In this case, the model isn't complex enough to have good accuracy on the training dataset.
This phenomena is often described in terms of the *bias-variance tradeoff*. Here, we decompose the error of the model into three terms:
$$
\textrm{Error} =
\textrm{Bias} +
\textrm{Variance} +
\textrm{Irreducible Error}.
$$
- The *bias* of the method is the error caused by the simplifying assumptions built into the method.
+ The *variance* of the method is how much the model will change based on the sampled data.
+ The *irreducible error* is error in the data itself, so no model can capture this error.
There is a tradeoff between the bias and variance of a model.
High-variance methods (e.g., the blue method) are accurate on the training set, but overfit noise in the data, so don't generalized well to new data. High-bias models (e.g., the black method) are too simple to fit the data, but are better at generalizing to new test data.
## Generalizability in practice
Consider the Auto dataset, which contains 9 features (mpg, cylinders, displacement, horsepower, weight, acceleration, year, origin, name) for 397 different used cars. This dataset is available digitally [here](http://www-bcf.usc.edu/~gareth/ISL/).
```
auto = pd.read_csv('Auto.csv') #load data
# one of the horsepowers is '?', so we just remove it and then map the remaining strings to integers
auto = auto[auto.horsepower != '?']
auto['horsepower'] = auto['horsepower'].map(int)
auto
print(auto.describe())
```
Let's consider the relationship between mpg and horsepower.
```
plt.scatter(auto['horsepower'],auto['mpg'],color='black',linewidth=1)
plt.xlabel('horsepower'); plt.ylabel('mpg')
plt.ylim((0,50))
plt.show()
```
We consider the linear model
$$
\text{mpg} = \beta_0 + \beta_1 \text{horsepower} + \beta_2 \text{horsepower}^2 + \cdots + \beta_m \text{horsepower}^m
$$
It might seem that choosing $m$ to be large would be a good thing. After all, a high degree polynomial is more flexible than a small degree polynomial.
```
# fit polynomial models
mr1 = sm.ols(formula="mpg ~ horsepower", data=auto).fit()
par1 = dict(mr1.params)
mr2 = sm.ols(formula="mpg ~ horsepower + I(horsepower ** 2.0)", data=auto).fit()
par2 = dict(mr2.params)
mr3 = sm.ols(formula="mpg ~ horsepower + I(horsepower ** 2.0) + I(horsepower ** 3.0)", data=auto).fit()
par3 = dict(mr3.params)
mr4 = sm.ols(formula="mpg ~ horsepower + I(horsepower ** 2.0) + I(horsepower ** 3.0) + I(horsepower ** 4.0)", data=auto).fit()
par4 = dict(mr4.params)
plt.scatter(auto['horsepower'],auto['mpg'],color='black',label="data")
x = sc.linspace(0,250,1000)
y1 = par1["Intercept"] + par1['horsepower']*x
y2 = par2["Intercept"] + par2['horsepower']*x + par2['I(horsepower ** 2.0)']*x**2
y3 = par3["Intercept"] + par3['horsepower']*x + par3['I(horsepower ** 2.0)']*x**2 + par3['I(horsepower ** 3.0)']*x**3
y4 = par4["Intercept"] + par4['horsepower']*x + par4['I(horsepower ** 2.0)']*x**2 + par4['I(horsepower ** 3.0)']*x**3 + par4['I(horsepower ** 4.0)']*x**4
plt.plot(x,y1,label="degree 1",linewidth=2)
plt.plot(x,y2,label="degree 2",linewidth=2)
plt.plot(x,y3,label="degree 3",linewidth=2)
plt.plot(x,y4,label="degree 4",linewidth=2)
plt.legend()
plt.xlabel('horsepower'); plt.ylabel('mpg')
plt.ylim((0,50))
plt.show()
print('mr1:',mr1.rsquared)
print('mr2:',mr2.rsquared)
print('mr3:',mr3.rsquared)
print('mr4:',mr4.rsquared)
```
As $m$ increases, the $R^2$ value is becoming larger. (You can prove that this is always true if you add more predictors.)
Let's check the $p$-values for the coefficients for the degree 4 fit.
```
mr2.summary()
```
For $m>2$, the $p$-values are very large, so we don't have a strong relationship between the variables.
We could rely on *Occam's razor* to decide between models. Occam's razor can be stated: among many different models that explain the data, the simplest one should be used. Since we don't get much benefit in terms of $R^2$ values by choosing $m>2$, we should use $m=2$.
But there are even better criterion for deciding between models.
## Cross-validation
There is a clever method for developing generalizable models that aren't underfit or overfit, called *cross validation*.
**Cross-validation** is a general method for assessing how the results of a predictive model (regression, classification,...) will *generalize* to an independent data set. In regression, cross-validation is a method for assessing how well the regression model will predict the dependent value for points that weren't used to *train* the model.
The idea of the method is simple:
1. Split the dataset into two groups: the training dataset and the testing dataset.
+ Train a variety of models on the training dataset.
+ Check the accuracy of each model on the testing dataset.
+ By comparing these accuracies, determine which model is best.
In practice, you have to decide how to split the data into groups (i.e. how large the groups should be). You might also want to repeat the experiment so that the assessment doesn't depend on the way in which you split the data into groups. We'll worry about these questions in a later lecture.
As the model becomes more complex ($m$ increases), the accuracy always increases for the training dataset. But, at some point, it starts to overfit the data and the accuracy decreases for the test dataset! Cross validation techniques will allow us to find the sweet-spot for the parameter $m$! (Think: Goldilocks and the Three Bears.)
Let's see this concept for the relationship between mpg and horsepower in the Auto dataset. We'll use the scikit-learn package for the cross validation analysis instead of statsmodels, because it is much easier to do cross validation there.
```
lr = linear_model.LinearRegression() # create a linear regression object
# with scikit-learn, we have to extract values from the pandas dataframe
for m in sc.arange(2,6):
auto['h'+str(m)] = auto['horsepower']**m
X = auto[['horsepower','h2','h3','h4','h5']].values.reshape(auto['horsepower'].shape[0],5)
y = auto['mpg'].values.reshape(auto['mpg'].shape[0],1)
plt.scatter(X[:,0], y, color='black',label='data')
# make data for plotting
xs = sc.linspace(20, 250, num=100)
Xs = sc.zeros([100,5])
Xs[:,0] = xs
for m in sc.arange(1,5):
Xs[:,m] = xs**(m+1)
for m in sc.arange(1,6):
lr.fit(X=X[:,:m], y=y)
plt.plot(xs, lr.predict(X=Xs[:,:m]), linewidth=3, label = "m = " + str(m) )
plt.legend(loc='upper right')
plt.xlabel('horsepower'); plt.ylabel('mpg')
plt.ylim((0,50))
plt.show()
```
### Cross validation using scikit-learn
- In scikit-learn, you can use the [*train_test_split*](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function to split the dataset into a training dataset and a test dataset.
+ The *score* function returns the coefficient of determination, $R^2$, of the prediction.
In the following code, I've split the data in an unusual way - taking the test set to be 90% - to illustrate the point more clearly. Typically, we might make the training set to be 90% of the dataset.
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.9, random_state=1)
print(X_train.shape, y_train.shape)
print(X_test.shape, y_test.shape)
plt.scatter(X_train[:,0], y_train, color='red',label='training data')
plt.scatter(X_test[:,0], y_test, color='black',label='test data')
for m in sc.arange(1,6):
lr.fit(X=X_train[:,:m], y=y_train)
print('m=', m, ', train: ', lr.score(X_train[:,:m], y_train), ' test: ', lr.score(X_test[:,:m], y_test))
plt.plot(xs, lr.predict(X=Xs[:,:m]), linewidth=3, label = "m = " + str(m) )
plt.legend()
plt.xlabel('horsepower'); plt.ylabel('mpg')
plt.ylim((0,50))
plt.show()
```
We observe that as the model complexity increases,
- the accuracy on the training data increases, but
+ the generalizability of the model to the test set decreases.
Our job as data analysts is to find a model that is sufficiently complex to describe the training data, but not so complex that it isn't generalizable to new data.
## Class exercise: analysis of the credit dataset
Next, we'll use [Statsmodels](http://statsmodels.sourceforge.net/) to study a dataset related to credit cards.
We'll use the 'Credit' dataset, available
[here](http://www-bcf.usc.edu/~gareth/ISL/data.html).
This dataset consists of some credit card information for 400 people.
Of course, a *credit card* is a card issued to a person ("cardholder"), typically from a bank, that can be used as a method of payment. The card allows the cardholder to borrow money from the bank to pay for goods and services. Credit cards have a *limit*, the maximum amount you can borrow, which is determined by the bank. The limit is determined from information collected from the cardholder (income, age, ...) and especially (as we will see) the cardholders "credit rating". The *credit rating* is an evaluation of the (1) ability of the cardholder to pay back the borrowed money and (2) the likelihood of the cardholder to defaulting on the borrowed money.
Our focus will be on the use of regression tools to study this dataset. Ideally, we'd like to understand what factors determine *credit ratings* and *credit limits*. We can think about this either from the point of view of (1) a bank who wants to protect their investments by minimizing credit defaults or (2) a person who is trying to increase their credit rating and/or credit limit.
A difficulty we'll encounter is including categorical data in regression models.
```
# Import data from Credit.csv file
credit = pd.read_csv('Credit.csv',index_col=0) #load data
credit
# Summarize and describe data
print(credit.dtypes, '\n')
print(credit['Gender'].value_counts(), '\n')
print(credit['Student'].value_counts(), '\n')
print(credit['Married'].value_counts(), '\n')
print(credit['Ethnicity'].value_counts())
credit.describe()
```
The column names of this data are:
1. Income
+ Limit
+ Rating
+ Cards
+ Age
+ Education
+ Gender (categorial: M,F)
+ Student (categorial: Y,N)
+ Married (categorial: Y,N)
+ Ethnicity (categorial: Caucasian, Asian, African American)
+ Balance
**Question:** What is wrong with the income data? How can it be fixed?
The file 'Credit.csv' is a comma separated file. I assume a period was used instead of a comma to indicate thousands in income so it wouldn't get confused with the separating value? Or maybe this is a dataset from Europe? Or maybe the income is just measured in \$1k units? To change the income data, we can use the Pandas series 'map' function.
```
credit["Income"] = credit["Income"].map(lambda x: 1000*x)
print(credit[:10])
```
We can also look at the covariances in the data. (This is how the variables vary together.) There are two ways to do this:
1. Quantitatively: Compute the correlation matrix. For each pair of variables, $(x_i,y_i)$, we compute
$$
\frac{\sum_i (x_i - \bar x) (y_i - \bar y)}{s_x s_y}
$$
where $\bar x, \bar y$ are sample means and $s_x, s_y$ are sample variances.
+ Visually: Make a scatter matrix of the data
```
credit.corr()
# trick: semi-colon prevents output
pd.plotting.scatter_matrix(credit, figsize=(10, 10), diagonal='kde');
```
**Observations:**
1. Limit and Rating are highly correlated ($99.7\%$)
+ Income strongly correlates with Limit ($79\%$) and Rating ($79\%$)
+ Balance correlates with Limit ($86\%$) and Rating ($86\%$)
+ There are "weird stripes" in some of the data. Why?
+ Categorical information doesn't appear in this plot. Why? How can I visualize the categorical variables?
```
# Plot Categorical variables: Gender, Student, Married, Ethnicity
fig, axes = plt.subplots(nrows=2, ncols=2,figsize=(10,10))
credit["Gender"].value_counts().plot(kind='bar',ax=axes[0,0]);
credit["Student"].value_counts().plot(kind='bar',ax=axes[1,0]);
credit["Married"].value_counts().plot(kind='bar',ax=axes[0,1]);
credit["Ethnicity"].value_counts().plot(kind='bar',ax=axes[1,1]);
```
## A first regression model
**Exercise:** First regress Limit on Rating:
$$
\text{Limit} = \beta_0 + \beta_1 \text{Rating}.
$$
Since credit ratings are primarily used by banks to determine credit limits, we expect that Rating is very predictive for Limit, so this regression should be very good.
Use the 'ols' function from the statsmodels python library.
```
# your code goes here
```
## Predicting Limit without Rating
Since Rating and Limit are almost the same variable, next we'll forget about Rating and just try to predict Limit from the real-valued variables (non-categorical variables): Income, Cards, Age, Education, Balance.
**Exercise:** Develop a multilinear regression model to predict Rating. Interpret the results.
For now, just focus on the real-valued variables (Income, Cards, Age, Education, Balance)
and ignore the categorical variables (Gender, Student, Married, Ethnicity).
```
# your code goes here
```
Which independent variables are good/bad predictors?
**Your observations:**
## Incorporating categorical variables into regression models
We have four categorical variables (Gender, Student, Married, Ethnicity). How can we include them in a regression model?
Let's start with a categorical variable with only 2 categories: Gender (Male, Female).
Idea: Create a "dummy variable" that turns Gender into a real value:
$$
\text{Gender_num}_i = \begin{cases}
1 & \text{if $i$-th person is female} \\
0 & \text{if $i$-th person is male}
\end{cases}.
$$
Then we could try to fit a model of the form
$$
\text{Income} = \beta_0 + \beta_1 \text{Gender_num}.
$$
```
credit["Gender_num"] = credit["Gender"].map({' Male':0, 'Female':1})
credit["Student_num"] = credit["Student"].map({'Yes':1, 'No':0})
credit["Married_num"] = credit["Married"].map({'Yes':1, 'No':0})
credit_model = sm.ols(formula="Income ~ Gender_num", data=credit).fit()
credit_model.summary()
```
Since the $p$-value for the Gender_num coefficient is very large, there is no support for the conclusion that there is a difference in credit card balance between genders.
**Exercise**: Try to find a meaningful relationship in the data including one of the categorical variables (Gender, Student, Married), for example, Balance vs. Student, Credit vs. Married, etc...
```
# your code here
```
## What about a categorical variable with 3 categories?
The Ethnicity variable takes three values: Caucasian, Asian, and African American.
What's wrong with the following?
$$
\text{Ethnicity_num}_i = \begin{cases}
0 & \text{if $i$-th person is Caucasian} \\
1 & \text{if $i$-th person is Asian} \\
2 & \text{if $i$-th person is African American}
\end{cases}.
$$
Hint: Recall Nominal, Ordinal, Interval, Ratio variable types from Lecture 4 (Descriptive Statistics).
We'll need more than one dummy variable:
$$
\text{Asian}_i = \begin{cases}
1 & \text{if $i$-th person is Asian} \\
0 & \text{otherwise}
\end{cases}.
$$
$$
\text{Caucasian}_i = \begin{cases}
1 & \text{if $i$-th person is Caucasian} \\
0 & \text{otherwise}
\end{cases}.
$$
The value with no dummy variable--African American--is called the *baseline*.
We can use the *get_dummies* function to automatically get these values
```
dummy = pd.get_dummies(credit['Ethnicity'])
credit = pd.concat([credit,dummy],axis=1)
credit
```
**Exercise**: Can you find a relationship in the data involving the variable ethnicity?
```
# your code here
```
| github_jupyter |
```
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
```
##### Exercise 7.1
Why do you think a larger random walk task (19 states instead of 5) was used in the examples of this chapter? Would a smaller walk have shifted the advantage to a different value of n? How about the change in left-side outcome from 0 to -1? Would that have made any difference in the best value of n?
A small random walk would truncate large n-step to their total returns since episodes will be shorter (i.e. large n would just result in alpha MC methods). Therefore we should expect the advantage at lower n for smaller random walks.
With values initialized at 0, if the left-most value terminated in 0 reward, we would need longer episodes for an agent to assign the correct values to the states left of center, since episodes that terminate to the left will not cause any updates initially, only the episodes that terminate to the right end with non-zero reward. Thus I would expect the best value of n to increase.
---------
##### Exercise 7.2
Why do you think on-line methods worked better than off-line methods on the example task?
Off-line methods generally take random actions with some small probability $\epsilon$. We would expect at least 1-2 random actions in an environment with a minimum of 10 states to termination, depending on $\epsilon$ (assuming $\epsilon$ is between 10-20%). Therefore, even after finding the optimal action-values, these random actions will attribute erroneous rewards to certain actions, leading to higher RMSEs compared to on-line methods; we also see that larger n is more optimal for off-line methods compared to on-line, presumably because larger n reduces noise from the $\epsilon$ greedy actions.
-----------
##### Exercise 7.3
In the lower part of Figure 7.2, notice that the plot for n=3 is different from the others, dropping to low performance at a much lower value of $\alpha$ than similar methods. In fact, the same was observed for n=5, n=7, and n=9. Can you explain why this might have been so? In fact, we are not sure ourselves.
My hypothesis is that odd values of n have higher RMSE because of the environment. It takes at a minimum, an odd number of steps to reach termination from the starting state. For off-line methods, even after finding the optimal action-values, an agent may still not terminate in an odd number of steps. Therefore my hypothesis is that odd n-step methods are more likely to cause erroneous updates to the $\epsilon$ greedy actions compared to even n-step methods. A quick way to test this, would be to create a random-walk where an agent will terminate at a minimum in an even number of steps, and then to observe the same plots as in Figure 7.2.
----------
#### Exercise 7.4
The parameter $\lambda $ characterizes how fast the exponential weighting in Figure 7.4 falls off, and thus how far into the future the $\lambda $-return algorithm looks in determining its backup. But a rate factor such as $\lambda $ is sometimes an awkward way of characterizing the speed of the decay. For some purposes it is better to specify a time constant, or half-life. What is the equation relating $\lambda $ and the half-life, $\tau$, the time by which the weighting sequence will have fallen to half of its initial value?
The half life occurs when weighting drops in half:
$ \lambda^{n} = 0.5 $,
which occurs at,
$n = -ln(2) / ln(\lambda) = \tau$
-----
Getting (7.3) from the equation above it:
$R_t^\lambda = (1 - \lambda) \sum_{n=1}^\infty \lambda^{n-1} R^{(n)}_t$,
after $T-t-1$, we sum to infinity but with $R^{T-t-1}_t$, which is just the total return $R_t$, so:
$R_t^\lambda = (1 - \lambda) \sum_{n=1}^{T-t-1} \lambda^{n-1} R^{(n)}_t + (1 - \lambda) R_t \sum_{n=T-t-1}^{\infty} \lambda^{n} $
We can remove $\lambda^{T-t-1}$ from the last sum to get $ (1 - \lambda) R_t \lambda^{T-t-1} \sum_{n=0}^\infty \lambda^n = (1 - \lambda) R_t \lambda^{T-t-1} \frac{1}{1 - \lambda}$, so that:
$R_t^\lambda = (1 - \lambda) \sum_{n=1}^{T-t-1} \lambda^{n} R^{(n)}_t + \lambda^{T-t-1} R_t $
----------
##### Exercise 7.5
In order to get TD($\lambda$) to be equivalent to the $\lambda$-return algorithm in the online case, the proposal is that $\delta_t = r_{t+1} + \gamma V_t(s_{t+1}) - V_{t-1}(s_t) $ and the n-step return is $R_t^{(n)} = r_{t+1} + \dots + \gamma^{n-1} r_{t+n} + \gamma^n V_{t+n-1}(s_{t+n}) $. To show that this new TD method is equivalent to the $\lambda$ return, it suffices to show that $\Delta V_t(s_t)$ for the $\lambda$ return is equivalent to the new TD with modified $\delta_t$ and $R_t^{(n)}$.
As such, we expand the $\lambda$ return:
$
\begin{equation}
\begin{split}
\frac{1}{\alpha} \Delta V_t(s_t) =& -V_{t-1}(s_t) + R_t^\lambda\\
=& -V_{t-1}(s_t) + (1 - \lambda) \lambda^0 [r_{t+1} + \gamma V_t(s_{t+1})] + (1-\lambda) \lambda^1 [r_{t+1} + \gamma r_{t+2} + \gamma^2 V_{t+1}(s_{t+2})] + \dots\\
=& -V_{t-1}(s_t) + (\gamma \lambda)^0 [r_{t+1} + \gamma V_t(s_{t+1}) - \gamma \lambda V_t(s_{t+1})] + (\gamma \lambda)^1 [r_{t+2} + \gamma V_{t+1}(s_{t+2}) - \gamma \lambda V_{t+1}(s_{t+2})] + \dots\\
=& (\gamma \lambda)^0 [r_{t+1} + \gamma V_t(s_{t+1}) - V_{t-1}(s_t)] + (\gamma \lambda) [r_{t+2} + \gamma V_{t+1}(s_{t+2}) - V_t(s_t+1)] + \dots\\
=& \sum_{k=t}^\infty (\gamma \lambda)^{k-t} \delta_k
\end{split}
\end{equation}
$
where $\delta_k = r_k + \gamma V_k(s_{k+1}) - V_{k-1}(s_k)$ as defined in the problem. Therefore, for online TD as defined above, the $\lambda$ return is exactly equivalent.
-------------
##### Exercise 7.6
In Example 7.5, suppose from state s the wrong action is taken twice before the right action is taken. If accumulating traces are used, then how big must the trace parameter $\lambda $ be in order for the wrong action to end up with a larger eligibility trace than the right action?
The eligibility trace update is $e_t(s) \leftarrow 1 + \gamma \lambda e_{t-1}(s)$ if $s = s_t$ and $e_t(s) \leftarrow \gamma \lambda e_{t-1}(s)$ if $s \neq s_t$. For two wrong actions, then one right action, $e_t(wrong) = (1 + \gamma \lambda) \gamma \lambda $, and $e_t(right) = 1$. If we want $e_t(wrong) \gt e_t(right)$, we need $(1 + \gamma \lambda) \gamma \lambda \gt 1$, or $\gamma \lambda \gt \frac{1}{2} (\sqrt(5) - 1)$.
-----------
##### Exercise 7.7
```
class LoopyEnvironment(object):
def __init__(self):
self._terminal_state = 5
self._state = 0
self._num_actions = 2
@property
def state(self):
return self._state
@state.setter
def state(self, state):
assert isinstance(state, int)
assert state >= 0 and state <= self._terminal_state
self._state = state
@property
def terminal_state(self):
return self._terminal_state
def reinit_state(self):
self._state = 0
def get_states_list(self):
return range(self._terminal_state + 1)
def get_actions_list(self):
return range(self._num_actions)
def is_terminal_state(self):
return self._state == self._terminal_state
def take_action(self, action):
"""
action int: 0 or 1
if action is 0 = wrong, then don't change the state
if action is 1 = right, then go to the next state
returns int: reward
"""
assert action in [0, 1]
assert self.is_terminal_state() == False
if action == 1:
self._state += 1
if self._state == self._terminal_state:
return 1
return 0
import random
from itertools import product
class SARSA_lambda(object):
def __init__(self, environment):
states = environment.get_states_list()
actions = environment.get_actions_list()
self.environment = environment
self.state_actions = list(product(states, actions))
self.Q = np.random.random([len(states), len(actions)])
self.e = np.zeros([len(states), len(actions)])
def _get_epsilon_greedy_action(self, epsilon, p):
if random.random() <= epsilon:
action = random.randint(0, len(p) - 1)
return action
actions = np.where(p == np.amax(p))[0]
action = np.random.choice(actions)
return action
def learn(self, num_episodes=100, Lambda=.9, gamma=.9, epsilon=.05, alpha=0.05,
replace_trace=False):
"""
Args:
num_episodes (int): Number of episodes to train
Lambda (float): TD(lambda) parameter
(if lambda = 1 we have MC or if lambda = 0 we have 1-step TD)
gamma (float): decay parameter for Bellman equation
epsilon (float): epsilon greedy decisions
alpha (float): determines how big should TD update be
Returns:
list (int): the number of time steps it takes for each episode to terminate
"""
time_steps = []
for n in xrange(num_episodes):
time_idx = 0
self.e = self.e * 0
self.environment.reinit_state()
s = self.environment.state
a = random.randint(0, self.Q.shape[1] - 1)
while not self.environment.is_terminal_state():
r = self.environment.take_action(a)
time_idx += 1
s_prime = self.environment.state
a_prime = self._get_epsilon_greedy_action(epsilon, self.Q[s_prime, :])
delta = r + gamma * self.Q[s_prime, a_prime] - self.Q[s, a]
if replace_trace:
self.e[s, a] = 1
else:
self.e[s, a] = self.e[s, a] + 1
for s, a in self.state_actions:
self.Q[s, a] = self.Q[s, a] + alpha * delta * self.e[s, a]
self.e[s, a] = gamma * Lambda * self.e[s, a]
s = s_prime
a = a_prime
time_steps.append(time_idx)
return time_steps
env = LoopyEnvironment()
s = SARSA_lambda(env)
```
Run both the replace-trace and the SARSA($\lambda$) regular trace methods for X episodes, and repeat N times. Get the average time length over all X episodes for each iteration for each alpha. In the environment in Figure 7.18, it takes at a minimum, 5 time steps to terminate. This is our baseline.
```
def get_results(replace_trace, num_trials, num_episodes):
alphas = np.linspace(.2, 1, num=10)
results = np.array([])
for alpha in alphas:
res = []
for i in xrange(num_trials):
sarsa_lambda = SARSA_lambda(env)
t = sarsa_lambda.learn(num_episodes=num_episodes, alpha=alpha,
replace_trace=replace_trace, gamma=0.9,
epsilon=0.05, Lambda=0.9)
res.append(np.mean(t))
if results.shape[0] == 0:
results = np.array([alpha, np.mean(res)])
else:
results = np.vstack([results, [alpha, np.mean(res)]])
return results
num_trials = 100
num_episodes = 20
replace_trace = get_results(True, num_trials, num_episodes)
regular_trace = get_results(False, num_trials, num_episodes)
plt.plot(replace_trace[:, 0], replace_trace[:, 1], label='replace')
plt.plot(regular_trace[:, 0], regular_trace[:, 1], label='regular')
plt.legend()
plt.title('Exercise 7.7: First %d episodes averaged %d times' %(num_episodes, num_trials))
plt.xlabel('alpha')
plt.ylabel('Time-steps')
```
We see that on average, the replace trace method for $\gamma = 0.9$, $\lambda=0.9$, $\epsilon=0.05$ takes less time to terminate. With lower $\gamma$, the advantage of replace-trace seems to disappear.
-----------
##### Exercise 7.8
sarsa($\lambda$) with replacing traces, has a backup which is equivalent to sarsa($\lambda$) until the first repeated state-action pair. If we use the replace-trace formula in Figure 7.17, the replace-trace backup diagram terminates at the first repeated state-action pair. For the replace-trace formula in Figure 7.16, the backup diagram after the first repeated-state action pair is some hybrid of sarsa($\lambda$) with weights changed only for the repeated state-actions. I'm not sure how to draw that.
-------
##### Exercise 7.9
Write pseudocode for an implementation of TD($\lambda $) that updates only value estimates for states whose traces are greater than some small positive constant.
You can use a hash-map of traces to update, and if the update reduces the value of the trace below some constant, remove the trace from the hash-map. Traces get added to the hash-map as they get visited. If you want to write the pseudo code or real code, feel free to make a pull-request!
-------
##### Exercise 7.10
Prove that the forward and backward views of off-line TD($\lambda $) remain equivalent under their new definitions with variable $\lambda $ given in this section. Follow the example of the proof in Section 7.4.
As given in the book, the backward view is:
$
e_t(s)=\left\{
\begin{array}{ll}
\gamma \lambda_t e_{t-1}(s), & \mbox{ if } s \neq s_t\\
\gamma \lambda_t e_{t-1}(s) + 1, & \mbox{ if } s = s_t
\end{array}
\right.
$
and the forward view is:
$R_t^\lambda = \sum_{k=t+1}^{T-1} R_t^{(k-t)} (1 - \lambda_k) \prod_{i=t+1}^{k-1} \lambda_i + R_t \prod_{i=t+1}^{T-1} \lambda_i$.
The proof is almost identical to 7.4. For the backward view we need to express the eligibility trace nonrecursively:
$e_t(s) = \gamma \lambda_t e_{t-1}(s) + I_{ss_t} = \gamma \lambda_t [\gamma \lambda_{t-1} e_{t-2}(s) + I_{ss_{t-1}}] + I_{ss_t} = \sum_{k=0}^t I_{ss_k}\gamma^{t-k} \prod_{i=k+1}^t \lambda_i$
so that the sum of all updates to a given state is:
$\sum_{t=0}^{T-1}\alpha I_{ss_t} \sum_{k=t}^{T-1} \gamma^{k-t} \prod_{i=t+1}^k \lambda_i \delta_k$
which was obtained by following the same algebra as in 7.9 to 7.12.
The next step is to show that the sum of all updates of the forward view is equivalent to the previous equation above. We start with:
$
\begin{equation}
\begin{split}
\frac{1}{\alpha} \Delta V_t(s_t) =& -V_{t}(s_t) + R_t^\lambda\\
=& -V_t(s_t) + (1 - \lambda_{t+1}) [r_{t+1} + \gamma V_t(s_{t+1})] + (1 - \lambda_{t+2})\lambda_{t+1} [r_{t+1} + \gamma r_{t+2} + \gamma^2 V_t(s_{t+2})] + \dots\\
=& -V_{t}(s_t) + [r_{t+1} + \gamma V_t(s_{t+1}) - \lambda_{t+1} \gamma V_t(s_{t+1})] + \gamma \lambda_{t+1} [r_{t+2} + \gamma V_t(s_{t+2}) - \gamma \lambda_{t+2} V_t(s_{t+2})] + \dots\\
=& [r_{t+1} + \gamma V_t(s_{t+1}) - V_t(s_t)] + (\gamma \lambda_{t+1})[r_{t+2} + \gamma V_t(s_{t+2}) - V_t(s_{t+1})] + (\gamma^2 \lambda_{t+1}\lambda_{t+2}) \delta_{t+3} + \dots\\
\approx& \sum_{k=t}^{T-1} \gamma^{k-t} \delta_k \prod_{i=t+1}^{k} \lambda_i
\end{split}
\end{equation}
$
which is equivalent to the backward case, and becomes an equality for offline updates.
------
** "Eligibility traces are the first line of defense against both long-delayed rewards and non-Markov tasks."**
"In the future it may be possible to vary the trade-off between TD and Monte Carlo methods more finely by using variable $\lambda $, but at present it is not clear how this can be done reliably and usefully."
| github_jupyter |
# Machine learning for yield prediction in Ghana
## Ismail Ougamane, Ismail.Ougamane@gmail.com
## Abstract:
The project can be Summaries into 2 main parts, the first part is for us to show the use of machine learning algorithms for yield prediction, in this part, we have formulate the problematic of the project into 3 research questions, then we have present the use of 2 main keywords in this project yield prediction and machine learning, after that we have present the use of machine learning in yield prediction and in the end of this part, I have answers to the 3 research questions.The second part of this project is a case study for yield prediction in Ghana using machine learning algorithms, the process of the conduct of the project consist of four steps the first step is gathering data, the second step is pre-processing the data, the third step is exploring the data and the last is building and choosing the best machine learning model for yield prediction.
## Introduction:
Although machine learning algorithms are frequently used for yield prediction there is no recent
study available yet to the best of our knowledge that investigates and summarizes the utilization of
machine learning in yield prediction. This study is performed to find answers to the following
questions :
1. Research Question 1: How yield prediction is defined and measured?
2. Research Question 2: What are the factors that control yield prediction, and how these could be included in machine learning?
3. Research Question 3: How machine learning methods could be tailored for modelling yield prediction?
Yield prediction in precision farming is considered of high importance for the improvement of
crop management, since variations in crop yield from year to year impact international trade, food
supply and market prices, this prediction can be useful for policy purposes.
Crop yield measurements or agriculture productivity measurements are important indicators of the
productivity and also provide a basis for assessing whether a landscape is supporting the livelihood
of individuals who farm the land. Kilograms per hectare is commonly used crop yield measurement,
often this standard weighs ( kilograms ) per area measure ( hectare ) crop yield is converted from a
volumetric unit of measurement that is based on a commonly used container, this method of
measuring involve weighing a complete harvest or relying on expert judgement, these two options
are very expensive so we use the following two methods are more economical and provide a
reasonably accurate assessment of crop yield:
• Harvesting: Random sample of the crop in a particular field is cut and weighed. Then the total yield is calculated from the weight multiplied by the total acreage production.
• Framer estimation: We ask farmers for their estimation of the total crop harvested, this value is divided by how much land they planted in order to estimate the yield.
This methods has been to be accurate in determining annual or seasonal crop yield, but is not effective for a continuous crop.
Machine learning gives the computer the ability to learn from experiences without being explicitly programmed , machine learning algorithms can be classified into 3 classes :
• The first class, supervised learning is when the model learns from the labelled data with direct feedback, supervised learning algorithms are used for regression problems like forecasting and for classification problems like image classification.
• The second class of machine learning algorithms is unsupervised learning is when we have no labelled data with no feedback, unsupervised learning algorithms are used for clustering problems like recommended system and dimensionality reduction problem like feature reduction.
• The third class is semi-supervised learning is used when we have a few labelled data and a large amount of unlabelled data, to do that semi supervised learning train with a reward system, providing feedback when the model performs with the best score of labelling the unlabelled data.
Before applying machine learning models, we should check 3 criteria’ The first criterion is to ask if there is a pattern to be found in our data, even that is difficult to prove, and in all times we work
without proving it. The second criterion: we can't pin down the pattern mathematically. The third criterion, we should have data that represent this pattern.
## Machine learning for yield prediction:
Yield prediction is one of the most important topics in precision agriculture, classic yield prediction
models use relationships between various factors like meteorological information and soil
parameters – and the crop yields, the yield prediction models can be categorise into 3 categories:
1. The goal of Statistical analysis is to map crop productivity to the soil proprieties (cation exchange capacity (CEC), pH, organic matter,... .), to soil characteristics (texture, soil types,...) and with climatic information (rainfall, temperature, radiation from the sun,... .), the statistical analysis used linear and non linear regression analysis to achieve the goal, the statistical methods are generally regarded to be unrealistic for particle purposes.
2. Mechanistic Model simulates the process of carbon assimilation using physical environment factors (like pollution or proximity of toxic sites) and other various environmental factors such as climatic information, management practice and soil character issues. In mechanistic modelling we use the relationship between physical environments and crop productivity also the soil conditions are integrated to simulate crop growth and yield prediction, mechanistic models are advantageous in the interpretability of the results, special case of mechanistic model is the chlorophyll meters in yield prediction that correlated directly chlorophyll content of leaves and yield prediction using SPAD chlorophyll meters.
3. Machine learning models based on sensed data , in the past years, many types of sensors and satellite platforms have been used to gather data for yield prediction, the objective of using machine learning models is to match the data with the yield prediction to do that there are 2 methods:
◦ Method 1: Directly correlate the spectral information to crop yield using regression models and vegetation indices (simple ratio, green area above bare soil).
◦ Method 2: Which estimate various crop parameters such as leaf area index and biomass from remotely sensed data.
Computer based image interpretation is seen as the best methods for analysis and interpretation of
remotely sensed images, machine learning algorithms are currently regarded as the key in the
development of image interpretation.
In general remote sensing system are widely used in building decision systems or tools, however
remote sensing based approaches require processing of the enormous amount of data from different
sources using machine learning models, for the error measurement of the models usually depend on
the nature and the goal of the project.
#### Answers to Research Questions:
The answers to research questions are as follows:
Research Question 1: How yield prediction is defined and measured?
Answer: Yield prediction is one of the most important topics in precision agriculture, since the yield prediction can impact the international trade, food supply and market prices. For the crop
yield measurement we use as unit Kilograms per Hectare and to compute the crop yield there are two economical methods:
• Harvesting,
• Farmer estimation.
Research Question 2: What are the factors that control yield prediction, and how these could be included in machine learning?
Answer: The factors that control yield prediction can be categories into 5 categories:
1. Soil proprieties factor: cation exchange caption(CEC), pH, Organic matter, ... .
2. Soil characteristic factor: texture, soil type, ... .
3. Climatic information: rainfall, temperature, radiation from the sun, ... .
4. Physical environment factors: pollution or proximity of toxic sites.
5. Management practice.
All these factors can be included into machine learning models as variables.
Research Question 3: How machine learning methods could be tailored for modelling yield prediction?
Answer: Not only machine learning models can take the factors that control yield prediction as variable, but also can learn from sensed image and satellite images that is seen as the best methods
for analysis and interpretation of the results of yield prediction.
## Machine learning for yield prediction in Ghana:
The purpose of our project is to build a machine learning model for yield prediction in Ghana, the process of the project can be divide into 4 steps:
1. Gather the data,
2. Process the data,
3. Explore the data,
4. Build and choose the model the model.
```
import numpy as np
import pandas as pd
import seaborn as sns
%matplotlib inline
import matplotlib.pyplot as plt
from pandas import read_csv
import seaborn as sns
import glob
import pickle
```
#### Gather the data:
Obtain the data is the core of any machine learning project, in our case we have
used the dataset provide by the “Food and Agriculture Organization of the United Nations
(FAO)”, the dataset contain crop statistics for 173 products in Africa, the Americas, Asia, Europe,
and Oceania(for more details see the cells bellow).
```
data = pd.read_csv('FAOSTAT_data_5-24-2020.csv')
data.info()
data=data[data['Value'].notna()]
data.info()
print(data.shape)
display(data.head())
```
#### Pre-processing data:
After obtaining the data the next immediate thing to do is process the data.
This process is for us to clean and filter the data, to do that we have deal with :
1. Missing values,
2. Categorical data.
```
data.head()
data=data.drop(['Year Code','Flag Description',"Flag"],axis=1)
data.head()
data_new=pd.get_dummies(data)
data_new.head()
```
#### Explore the data:
Before jumping into building the model, we will examine the data, first of all we
will inspect the data and its properties, then we will use the data visualization to help us to identify
significant patterns and trends in our dataset, the scatter-plot below shows a strong positive, non- linear association between the yield estimation and years for items Avocados, banana and beans
whether for the dry beans or for the green beans, and for the other items shows a positive linear
association between yield estimation and years.
```
# Analysis
f,ax = plt.subplots(figsize = (5,5))
sns.heatmap(data.corr(), annot=True)
## this part is optional. I had to do it because the
## plot had been disproportionate.
bottom, top = ax.get_ylim()
ax.set_ylim(bottom + 0.5, top - 0.5)
plt.show()
# Analysis
f,ax = plt.subplots(figsize = (25,20))
sns.scatterplot(data['Year'],data['Value'],legend='full',data=data,hue=data["Item"],s=60)
## this part is optional. I had to do it because the
## plot had been disproportionate.
# bottom, top = ax.get_ylim()
# ax.set_ylim(bottom + 0.5, top - 0.5)
plt.savefig('foo.png')
plt.show()
```
#### Build the model:
Regression models are one of the most powerful models used to find relations
within a dataset, with the key focus being on relationships between the independent variables
(predictors) and a dependent variable (outcome). In this phase, we will apply the following models:
1. Linear Regression,
2. Polynomial Regression,
3. Decision Tree Regression.
To validate our models we split the data set into 2 parties train data (70%) and test data (30%), to
assess the performance of the models we use R2 metrics:
1. Linear Regression score 0.266,
2. Polynomial Regression score 0.748,
3. Decision Tree Regression with Max Depth =2 score 0.525
4. Decision Tree Regression with Max Depth =5 score 0.938
```
from sklearn.model_selection import train_test_split
Y=data_new['Value']
X=data_new.drop(['Value'],axis=1)
print(X.shape,Y.shape)
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=.20, random_state=1)
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import PolynomialFeatures
from sklearn.metrics import r2_score,mean_squared_error
from sklearn.ensemble import RandomForestRegressor
x_train,x_test,y_train,y_test = train_test_split(X,Y, random_state = 0)
lr = LinearRegression().fit(x_train,y_train)
y_train_pred = lr.predict(x_train)
y_test_pred = lr.predict(x_test)
print(lr.score(x_test,y_test))
quad = PolynomialFeatures (degree = 2)
x_quad = quad.fit_transform(X)
X_train,X_test,Y_train,Y_test = train_test_split(x_quad,Y, random_state = 0)
plr = LinearRegression().fit(X_train,Y_train)
Y_train_pred = plr.predict(X_train)
Y_test_pred = plr.predict(X_test)
print(plr.score(X_test,Y_test))
import numpy as np
from sklearn.tree import DecisionTreeRegressor
import matplotlib.pyplot as plt
# Fit regression model
regr_1 = DecisionTreeRegressor(max_depth=2)
regr_2 = DecisionTreeRegressor(max_depth=5)
regr_1.fit(x_train, y_train)
regr_2.fit(x_train, y_train)
# Predict
y_1 = regr_1.predict(x_test)
y_2 = regr_1.predict(x_test)
y_train_pred = regr_1.predict(x_train)
y_test_pred = regr_1.predict(x_test)
print(regr_1.score(x_test,y_test))
y_1 = regr_2.predict(x_test)
y_2 = regr_2.predict(x_test)
y_train_pred = regr_2.predict(x_train)
y_test_pred = regr_2.predict(x_test)
print(regr_2.score(x_test,y_test))
```
## Conlusion:
In important issues for precision agriculture purposes is the accurate yield prediction, machine learning models are an essential approach for achieving practical and effective solutions for this problem.
So in our work we show the use of machine learning models in yield prediction, then for more clarification, we have implemented machine learning algorithms for yield prediction in Ghana.
| github_jupyter |
# Package loading and basic configurations
```
%load_ext autoreload
%autoreload 2
# load dependencies'
import pandas as pd
import geopandas as gpd
from envirocar import TrackAPI, DownloadClient, BboxSelector, ECConfig
# create an initial but optional config and an api client
config = ECConfig()
track_api = TrackAPI(api_client=DownloadClient(config=config))
```
# Querying enviroCar Tracks
The following cell queries tracks from the enviroCar API. It defines a bbox for the area of Münster (Germany) and requests 50 tracks. The result is a GeoDataFrame, which is a geo-extended Pandas dataframe from the GeoPandas library. It contains all information of the track in a flat dataframe format including a specific geometry column.
```
bbox = BboxSelector([
7.601165771484375, # min_x
51.94807412325402, # min_y
7.648200988769531, # max_x
51.97261482608728 # max_y
])
# issue a query
track_df = track_api.get_tracks(bbox=bbox, num_results=50) # requesting 50 tracks inside the bbox
track_df
track_df.plot(figsize=(8, 10))
```
# Inspecting a single Track
```
some_track_id = track_df['track.id'].unique()[5]
some_track = track_df[track_df['track.id'] == some_track_id]
some_track.plot()
ax = some_track['Speed.value'].plot()
ax.set_title("Speed")
ax.set_ylabel(some_track['Speed.unit'][0])
ax
```
## Interactive Map
The following map-based visualization makes use of folium. It allows to visualizate geospatial data based on an interactive leaflet map. Since the data in the GeoDataframe is modelled as a set of Point instead of a LineString, we have to manually create a polyline
```
import folium
lats = list(some_track['geometry'].apply(lambda coord: coord.y))
lngs = list(some_track['geometry'].apply(lambda coord: coord.x))
avg_lat = sum(lats) / len(lats)
avg_lngs = sum(lngs) / len(lngs)
m = folium.Map(location=[avg_lat, avg_lngs], zoom_start=13)
folium.PolyLine([coords for coords in zip(lats, lngs)], color='blue').add_to(m)
m
```
# Example: Visualization with pydeck (deck.gl)
The pydeck library makes use of the basemap tiles from Mapbox. In case you want to visualize the map with basemap tiles, you need to register with MapBox, and configure a specific access token. The service is free until a certain level of traffic is esceeded.
You can either configure it via your terminal (i.e. `export MAPBOX_API_KEY=<mapbox-key-here>`), which pydeck will automatically read, or you can pass it as a variable to the generation of pydeck (i.e. `pdk.Deck(mapbox_key=<mapbox-key-here>, ...)`.
```
import pydeck as pdk
# for pydeck the attributes have to be flat
track_df['lat'] = track_df['geometry'].apply(lambda coord: coord.y)
track_df['lng'] = track_df['geometry'].apply(lambda coord: coord.x)
vis_df = pd.DataFrame(track_df)
vis_df['speed'] = vis_df['Speed.value']
# omit unit columns
vis_df_cols = [col for col in vis_df.columns if col.lower()[len(col)-4:len(col)] != 'unit']
vis_df = vis_df[vis_df_cols]
layer = pdk.Layer(
'ScatterplotLayer',
data=vis_df,
get_position='[lng, lat]',
auto_highlight=True,
get_radius=10, # Radius is given in meters
get_fill_color='[speed < 20 ? 0 : (speed - 20)*8.5, speed < 50 ? 255 : 255 - (speed-50)*8.5, 0, 140]', # Set an RGBA value for fill
pickable=True
)
# Set the viewport location
view_state = pdk.ViewState(
longitude=7.5963592529296875,
latitude=51.96246168188569,
zoom=10,
min_zoom=5,
max_zoom=15,
pitch=40.5,
bearing=-27.36)
r = pdk.Deck(
width=200,
layers=[layer],
initial_view_state=view_state #, mapbox_key=<mapbox-key-here>
)
r.to_html('tracks_muenster.html', iframe_width=900)
```
| github_jupyter |
# Pneumonia Classification on TPU
**Author:** Amy MiHyun Jang<br>
**Date created:** 2020/07/28<br>
**Last modified:** 2020/08/24<br>
**Description:** Medical image classification on TPU.
## Introduction + Set-up
This tutorial will explain how to build an X-ray image classification model
to predict whether an X-ray scan shows presence of pneumonia.
```
import re
import os
import random
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver()
print("Device:", tpu.master())
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.TPUStrategy(tpu)
except:
strategy = tf.distribute.get_strategy()
print("Number of replicas:", strategy.num_replicas_in_sync)
```
We need a Google Cloud link to our data to load the data using a TPU.
Below, we define key configuration parameters we'll use in this example.
To run on TPU, this example must be on Colab with the TPU runtime selected.
```
AUTOTUNE = tf.data.AUTOTUNE
BATCH_SIZE = 25 * strategy.num_replicas_in_sync
IMAGE_SIZE = [180, 180]
CLASS_NAMES = ["NORMAL", "PNEUMONIA"]
```
## Load the data
The Chest X-ray data we are using from
[*Cell*](https://www.cell.com/cell/fulltext/S0092-8674(18)30154-5) divides the data into
training and test files. Let's first load in the training TFRecords.
```
train_images = tf.data.TFRecordDataset(
"gs://download.tensorflow.org/data/ChestXRay2017/train/images.tfrec"
)
train_paths = tf.data.TFRecordDataset(
"gs://download.tensorflow.org/data/ChestXRay2017/train/paths.tfrec"
)
ds = tf.data.Dataset.zip((train_images, train_paths))
```
Let's count how many healthy/normal chest X-rays we have and how many
pneumonia chest X-rays we have:
```
COUNT_NORMAL = len(
[
filename
for filename in train_paths
if "NORMAL" in filename.numpy().decode("utf-8")
]
)
print("Normal images count in training set: " + str(COUNT_NORMAL))
COUNT_PNEUMONIA = len(
[
filename
for filename in train_paths
if "PNEUMONIA" in filename.numpy().decode("utf-8")
]
)
print("Pneumonia images count in training set: " + str(COUNT_PNEUMONIA))
```
Notice that there are way more images that are classified as pneumonia than normal. This
shows that we have an imbalance in our data. We will correct for this imbalance later on
in our notebook.
We want to map each filename to the corresponding (image, label) pair. The following
methods will help us do that.
As we only have two labels, we will encode the label so that `1` or `True` indicates
pneumonia and `0` or `False` indicates normal.
```
def get_label(file_path):
# convert the path to a list of path components
parts = tf.strings.split(file_path, "/")
# The second to last is the class-directory
return parts[-2] == "PNEUMONIA"
def decode_img(img):
# convert the compressed string to a 3D uint8 tensor
img = tf.image.decode_jpeg(img, channels=3)
# resize the image to the desired size.
return tf.image.resize(img, IMAGE_SIZE)
def process_path(image, path):
label = get_label(path)
# load the raw data from the file as a string
img = decode_img(image)
return img, label
ds = ds.map(process_path, num_parallel_calls=AUTOTUNE)
```
Let's split the data into a training and validation datasets.
```
ds = ds.shuffle(10000)
train_ds = ds.take(4200)
val_ds = ds.skip(4200)
```
Let's visualize the shape of an (image, label) pair.
```
for image, label in train_ds.take(1):
print("Image shape: ", image.numpy().shape)
print("Label: ", label.numpy())
```
Load and format the test data as well.
```
test_images = tf.data.TFRecordDataset(
"gs://download.tensorflow.org/data/ChestXRay2017/test/images.tfrec"
)
test_paths = tf.data.TFRecordDataset(
"gs://download.tensorflow.org/data/ChestXRay2017/test/paths.tfrec"
)
test_ds = tf.data.Dataset.zip((test_images, test_paths))
test_ds = test_ds.map(process_path, num_parallel_calls=AUTOTUNE)
test_ds = test_ds.batch(BATCH_SIZE)
```
## Visualize the dataset
First, let's use buffered prefetching so we can yield data from disk without having I/O
become blocking.
Please note that large image datasets should not be cached in memory. We do it here
because the dataset is not very large and we want to train on TPU.
```
def prepare_for_training(ds, cache=True):
# This is a small dataset, only load it once, and keep it in memory.
# use `.cache(filename)` to cache preprocessing work for datasets that don't
# fit in memory.
if cache:
if isinstance(cache, str):
ds = ds.cache(cache)
else:
ds = ds.cache()
ds = ds.batch(BATCH_SIZE)
# `prefetch` lets the dataset fetch batches in the background while the model
# is training.
ds = ds.prefetch(buffer_size=AUTOTUNE)
return ds
```
Call the next batch iteration of the training data.
```
train_ds = prepare_for_training(train_ds)
val_ds = prepare_for_training(val_ds)
image_batch, label_batch = next(iter(train_ds))
```
Define the method to show the images in the batch.
```
def show_batch(image_batch, label_batch):
plt.figure(figsize=(10, 10))
for n in range(25):
ax = plt.subplot(5, 5, n + 1)
plt.imshow(image_batch[n] / 255)
if label_batch[n]:
plt.title("PNEUMONIA")
else:
plt.title("NORMAL")
plt.axis("off")
```
As the method takes in NumPy arrays as its parameters, call the numpy function on the
batches to return the tensor in NumPy array form.
```
show_batch(image_batch.numpy(), label_batch.numpy())
```
## Build the CNN
To make our model more modular and easier to understand, let's define some blocks. As
we're building a convolution neural network, we'll create a convolution block and a dense
layer block.
The architecture for this CNN has been inspired by this
[article](https://towardsdatascience.com/deep-learning-for-detecting-pneumonia-from-x-ray-images-fc9a3d9fdba8).
```
from tensorflow import keras
from tensorflow.keras import layers
def conv_block(filters, inputs):
x = layers.SeparableConv2D(filters, 3, activation="relu", padding="same")(inputs)
x = layers.SeparableConv2D(filters, 3, activation="relu", padding="same")(x)
x = layers.BatchNormalization()(x)
outputs = layers.MaxPool2D()(x)
return outputs
def dense_block(units, dropout_rate, inputs):
x = layers.Dense(units, activation="relu")(inputs)
x = layers.BatchNormalization()(x)
outputs = layers.Dropout(dropout_rate)(x)
return outputs
```
The following method will define the function to build our model for us.
The images originally have values that range from [0, 255]. CNNs work better with smaller
numbers so we will scale this down for our input.
The Dropout layers are important, as they
reduce the likelikhood of the model overfitting. We want to end the model with a `Dense`
layer with one node, as this will be the binary output that determines if an X-ray shows
presence of pneumonia.
```
def build_model():
inputs = keras.Input(shape=(IMAGE_SIZE[0], IMAGE_SIZE[1], 3))
x = layers.Rescaling(1.0 / 255)(inputs)
x = layers.Conv2D(16, 3, activation="relu", padding="same")(x)
x = layers.Conv2D(16, 3, activation="relu", padding="same")(x)
x = layers.MaxPool2D()(x)
x = conv_block(32, x)
x = conv_block(64, x)
x = conv_block(128, x)
x = layers.Dropout(0.2)(x)
x = conv_block(256, x)
x = layers.Dropout(0.2)(x)
x = layers.Flatten()(x)
x = dense_block(512, 0.7, x)
x = dense_block(128, 0.5, x)
x = dense_block(64, 0.3, x)
outputs = layers.Dense(1, activation="sigmoid")(x)
model = keras.Model(inputs=inputs, outputs=outputs)
return model
```
## Correct for data imbalance
We saw earlier in this example that the data was imbalanced, with more images classified
as pneumonia than normal. We will correct for that by using class weighting:
```
initial_bias = np.log([COUNT_PNEUMONIA / COUNT_NORMAL])
print("Initial bias: {:.5f}".format(initial_bias[0]))
TRAIN_IMG_COUNT = COUNT_NORMAL + COUNT_PNEUMONIA
weight_for_0 = (1 / COUNT_NORMAL) * (TRAIN_IMG_COUNT) / 2.0
weight_for_1 = (1 / COUNT_PNEUMONIA) * (TRAIN_IMG_COUNT) / 2.0
class_weight = {0: weight_for_0, 1: weight_for_1}
print("Weight for class 0: {:.2f}".format(weight_for_0))
print("Weight for class 1: {:.2f}".format(weight_for_1))
```
The weight for class `0` (Normal) is a lot higher than the weight for class `1`
(Pneumonia). Because there are less normal images, each normal image will be weighted
more to balance the data as the CNN works best when the training data is balanced.
## Train the model
### Defining callbacks
The checkpoint callback saves the best weights of the model, so next time we want to use
the model, we do not have to spend time training it. The early stopping callback stops
the training process when the model starts becoming stagnant, or even worse, when the
model starts overfitting.
```
checkpoint_cb = tf.keras.callbacks.ModelCheckpoint("xray_model.h5", save_best_only=True)
early_stopping_cb = tf.keras.callbacks.EarlyStopping(
patience=10, restore_best_weights=True
)
```
We also want to tune our learning rate. Too high of a learning rate will cause the model
to diverge. Too small of a learning rate will cause the model to be too slow. We
implement the exponential learning rate scheduling method below.
```
initial_learning_rate = 0.015
lr_schedule = tf.keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate, decay_steps=100000, decay_rate=0.96, staircase=True
)
```
### Fit the model
For our metrics, we want to include precision and recall as they will provide use with a
more informed picture of how good our model is. Accuracy tells us what fraction of the
labels is correct. Since our data is not balanced, accuracy might give a skewed sense of
a good model (i.e. a model that always predicts PNEUMONIA will be 74% accurate but is not
a good model).
Precision is the number of true positives (TP) over the sum of TP and false positives
(FP). It shows what fraction of labeled positives are actually correct.
Recall is the number of TP over the sum of TP and false negatves (FN). It shows what
fraction of actual positives are correct.
Since there are only two possible labels for the image, we will be using the
binary crossentropy loss. When we fit the model, remember to specify the class weights,
which we defined earlier. Because we are using a TPU, training will be quick - less than
2 minutes.
```
with strategy.scope():
model = build_model()
METRICS = [
tf.keras.metrics.BinaryAccuracy(),
tf.keras.metrics.Precision(name="precision"),
tf.keras.metrics.Recall(name="recall"),
]
model.compile(
optimizer=tf.keras.optimizers.Adam(learning_rate=lr_schedule),
loss="binary_crossentropy",
metrics=METRICS,
)
history = model.fit(
train_ds,
epochs=100,
validation_data=val_ds,
class_weight=class_weight,
callbacks=[checkpoint_cb, early_stopping_cb],
)
```
## Visualizing model performance
Let's plot the model accuracy and loss for the training and the validating set. Note that
no random seed is specified for this notebook. For your notebook, there might be slight
variance.
```
fig, ax = plt.subplots(1, 4, figsize=(20, 3))
ax = ax.ravel()
for i, met in enumerate(["precision", "recall", "binary_accuracy", "loss"]):
ax[i].plot(history.history[met])
ax[i].plot(history.history["val_" + met])
ax[i].set_title("Model {}".format(met))
ax[i].set_xlabel("epochs")
ax[i].set_ylabel(met)
ax[i].legend(["train", "val"])
```
We see that the accuracy for our model is around 95%.
## Predict and evaluate results
Let's evaluate the model on our test data!
```
model.evaluate(test_ds, return_dict=True)
```
We see that our accuracy on our test data is lower than the accuracy for our validating
set. This may indicate overfitting.
Our recall is greater than our precision, indicating that almost all pneumonia images are
correctly identified but some normal images are falsely identified. We should aim to
increase our precision.
```
for image, label in test_ds.take(1):
plt.imshow(image[0] / 255.0)
plt.title(CLASS_NAMES[label[0].numpy()])
prediction = model.predict(test_ds.take(1))[0]
scores = [1 - prediction, prediction]
for score, name in zip(scores, CLASS_NAMES):
print("This image is %.2f percent %s" % ((100 * score), name))
```
| github_jupyter |
# Lab Practice Week 4
This notebook does not need to be submitted. This is only for you to gain experience and get some practice.
# Randomness, Histograms, Scatter plot
Today we will review all the manipulation from lectures for numpy arrays and how to use matplotlib to plot graphs.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
## Problem 1: model a biased coin
Reference: Lecture 10, Homework 3.
Consider the following **stochastic process**:
> Consider the following game: we start from the time $t_0 = 0$, at each subsequent $t_i=i$ ($i=1,2,\dots$), we flip a **biased** coin.
> If the coin lands on head, we win $\$ 1 $, otherwise we lose $\$ 1$.
> Suppose $M_i$ denotes our money (in $\$ $) in the wallet at $t_i$, and $M_0 = 0$ (when the money amount is $<0$, it means we owe money to the dealer).
We want to model how $M_i$ evolves after 1000 steps, please finish the following exercises.
### Question 1
* Create a function `coinflip(p)` where $p$ is some real number in $(0,1)$, which returns the float $1.0$ (for Heads) with probability $p$ and the float $-1.0$ with probaility $1-p$ (for Tails).
```
def coinflip(p):
if random.random() < p:
return 1.0
return -1.0
```
### Question 2
Let $p=0.6$, and consider $Y_n$ as follows
$$Y_n = \frac{X_1 + X_2 + ... + X_n}{n},$$
Each $X_i$ represent the flipping result at $i$-th step. If it's heads, $X_i=1$; if it is tails $X_i = -1$. If I play this flipping game $n$ times, the average winning per game is $Y_n$.
Let's design our experiment a little more carefully so that we can get the most information, please finish the following:
* **Sample** $Y_n$ 1000 times for $n=1,2,3,...,200$, that is, create 1000 simulations, each simulation is 200 steps long (200 flippings), record the sampled $Y_1 = X_1$ for all 1000 simulations, i.e., we should have 1000 samples for $Y_1$; $Y_2= (X_1 + X_2)/2$ for all 1000 simulations, etc.
* Use a numpy array `means` of shape `(200,)` to record the mean of the sampled $Y_n$ at each time step.
* Use a numpy array `stdevs` of shape `(200,)` to record the standard deviation of the sampled $Y_n$ at each time step.
### Question 3: Verification of the Law of Large Number and Central Limit Theorem
$p=0.6$ as well as in question 2. In the process of $n\to \infty$, it can be computed that the **true expected valued** of $E(Y_n) = 2p-1$, and the **true standard deviation** is $\dfrac{1- (2p-1)^2}{\sqrt{n}}$ at the $n$-th time step. Use the result from Question 2:
* Use the `plot` function in `matplotlib.pyplot` to plot the sample mean of $Y_n$ for these 200 steps.
* Plot the sample standard deviation of $Y_n$ against the true standard deviation for these 200 steps.
```
# q3
import matplotlib.pyplot as plt
%matplotlib inline
# histogram here
# plt.axis([0, 100, 0, 80])
result_1ktest = simulation_coinflip(0.6)
plt.hist(result_1ktest, bins = 20, edgecolor='k');
# the histogram is concentrated around 60 because p=0.6
```
| github_jupyter |
<a href="https://colab.research.google.com/github/neilpradhan/Deep_reinforcement_learning_for_autonomous_highway_driving_scenario/blob/main/Thesis_Testing_Plots.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
from google.colab import drive
drive.mount('/content/gdrive')
import pandas as pd
import numpy as np
from scipy.stats import norm
import matplotlib.pyplot as plt
import seaborn as sns
from math import exp
import pandas as pd
import io
import pandas as pd
%cd gdrive/My\ Drive/Thesis_training/
%cd train_car_truck/
## dataframe without " "
from collections import Counter
from collections import defaultdict
import pandas as pd
import re
import csv
import numpy as np
def get_dataframe(f):
k = Counter()
d = defaultdict()
cols_1 = ['episode_number','time_stamps_survived', 'survival_status','avg_episode_reward',\
'max_vel' , 'avg_vel', 'epsilon', 'probability_of_survival']
cols_2 = ['action_0','action_1', 'action_2', 'action_3']
df1 = pd.DataFrame(columns=cols_1)
# df2 = pd.DataFrame(columns=cols_2)
# pattern = re.compile()
a_list = []
with open(f, 'r') as v:
lines = v.readlines()
for line in lines:
a = line.split(" ")
# print(a[0:4])
b,c,d,e = [float(x) for x in a[0:4]]
# print(type(b))
match = re.search(r'Counter\(.*?\)',line).group(0)
k = eval(match)
# print(k[0])
# b = line.split(" ",2)
# print(b,c,d,e)
# print(f)
f = line.rsplit(" ")
# print(f)
f = f[::-1]
# print(f[0:5])
# print(f)
g,h,i,j,m = [float(x) for x in f[0:5]]
# print(g)
# print("\n")
# print(h)
# print("\n")
# print(i)
# print("\n")
# print(j)
# break
# break
# with open('numbers.csv', mode='a+') as num_file:
# num_writer = csv.writer(num_file, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
# num_writer.writerow([b,c,d,e,g,h,i,j])
l=[b,c,d,e,k[0],k[1],k[2],k[3],m,j,i,h,g]
a_list.append(l)
# series_row_1 = pd.Series(l)
# series_row_2 = pd.Series(k)
# new_row = {k:v for k,v in zip(cols_1,l)}
# df1 = df1.append(series_row_1, ignore_index = True)
# df2 = df2.append(series_row_2, ignore_index=True)
# return pd.concat([df1,df2],axis=1)
df1 = pd.DataFrame(a_list)
del a_list
return df1
# print(g,h,i,j)
# print(f)
# print(b)
# def line_parsing(line):
# def main():
# if __name__ == "__main__":
# main()
# sen = '0 1 -1 -3.0884098790240953 Counter({2: 1}) 3.0884098790240953 3.0884098790240953 1.0 0.0'
# ep no, tss, win/loss, avg_episodic_reward, counter, max vel, avg vel, epsilon, safety count
## dataframe with " "
# 1. only 4 at the end
# 2. remove additional m or letter
from collections import Counter
from collections import defaultdict
import pandas as pd
import re
import csv
import numpy as np
def get_dataframe(f):
k = Counter()
d = defaultdict()
cols_1 = ['episode_number','time_stamps_survived', 'survival_status','avg_episode_reward',\
'max_vel' , 'avg_vel', 'epsilon', 'probability_of_survival']
cols_2 = ['action_0','action_1', 'action_2', 'action_3']
df1 = pd.DataFrame(columns=cols_1)
# df2 = pd.DataFrame(columns=cols_2)
# pattern = re.compile()
a_list = []
with open(f, 'r') as v:
lines = v.readlines()
for line in lines:
a = line.split(" ")
b,c,d,e = [float(x) for x in a[0:4]]
# print(type(b))
match = re.search(r'Counter\(.*?\)',line).group(0)
k = eval(match)
# print(k[0])
# b = line.split(" ",2)
# print(b,c,d,e)
# print(f)
f = line.rsplit(" ")
# f.pop(7)
f.remove("")
# print(f)
f = f[::-1]
# print(f[0:4])
# print(f)
g,h,i,j = [float(x) for x in f[0:4]]
# print(g)
# print("\n")
# print(h)
# print("\n")
# print(i)
# print("\n")
# print(j)
# break
# break
# with open('numbers.csv', mode='a+') as num_file:
# num_writer = csv.writer(num_file, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
# num_writer.writerow([b,c,d,e,g,h,i,j])
l=[b,c,d,e,k[0],k[1],k[2],k[3],j,i,h,g]
a_list.append(l)
# series_row_1 = pd.Series(l)
# series_row_2 = pd.Series(k)
# new_row = {k:v for k,v in zip(cols_1,l)}
# df1 = df1.append(series_row_1, ignore_index = True)
# df2 = df2.append(series_row_2, ignore_index=True)
# return pd.concat([df1,df2],axis=1)
df1 = pd.DataFrame(a_list)
del a_list
return df1
# print(g,h,i,j)
# print(f)
# print(b)
# def line_parsing(line):
# def main():
# if __name__ == "__main__":
# main()
# sen = '0 1 -1 -3.0884098790240953 Counter({2: 1}) 3.0884098790240953 3.0884098790240953 1.0 0.0'
# ep_no, tss, win/loss, avg_episodic_reward, counter, max vel, avg vel, epsilon, safety count
```
#### POSITION VARIANCE ONLY
```
## dataframe with " "
## REWARD 1
f5 = 'testing_5.txt'
f6 = 'testing_6.txt'
f7 = 'testing_7.txt'
f8 = 'testing_8.txt'
f9 = 'testing_9.txt'
f10 = 'testing_10.txt'
f11 = 'testing_11.txt'
re1_pv1 = get_dataframe(f5) ## 100 points 7 features
re1_pv2 = get_dataframe(f6)
re1_pv3 = get_dataframe(f7)
re1_pv4 = get_dataframe(f8)
re1_pv5 = get_dataframe(f9)
re1_pv6 = get_dataframe(f10)
re1_pv7 = get_dataframe(f11)
xlabels_yo = [0.5,0.75,1.0,1.25, 1.5,1.75,2.0]
# sns.lineplot(data=df1, x=xlabels_yo, y="time_stamps_survived")
# sns.lineplot(data = ax,markers=True, dashes=False)
frames = [re1_pv1,re1_pv2,re1_pv3,re1_pv4,re1_pv5,re1_pv6,re1_pv7]
df1 = pd.concat(frames)
## df1 ready for reward 1 position variance
## REWARD 2
g15 = 'testing_15.txt'
g16 = 'testing_16.txt'
g17 = 'testing_17.txt'
g18 = 'testing_18.txt'
g19 = 'testing_19.txt'
g20 = 'testing_20.txt'
g21 = 'testing_21.txt'
re2_pv1 = get_dataframe(g15) ## 100 points 7 features
re2_pv2 = get_dataframe(g16)
re2_pv3 = get_dataframe(g17)
re2_pv4 = get_dataframe(g18)
re2_pv5 = get_dataframe(g19)
re2_pv6 = get_dataframe(g20)
re2_pv7 = get_dataframe(g21)
xlabels_yo = [0.5,0.75,1.0,1.25, 1.5,1.75,2.0]
# sns.lineplot(data=df1, x=xlabels_yo, y="time_stamps_survived")
# sns.lineplot(data = ax,markers=True, dashes=False)
frames = [re2_pv1,re2_pv2,re2_pv3,re2_pv4,re2_pv5,re2_pv6,re2_pv7]
df2 = pd.concat(frames)
## df1 ready for reward 1 position variance
a = []
for i in range(0,101):
a.append(0.5)
for i in range(101,202):
a.append(0.75) #101 values
for i in range(202,303):
a.append(1.0)
for i in range(303,404):
a.append(1.25)
for i in range(404,505):
a.append(1.50)
for i in range(505,606):
a.append(1.75)
for i in range(606,707):
a.append(2.0)
a = np.asarray(a)
df1[15] = a
df2[15] = a
plt.figure(1)
sns.lineplot(data=df1, x=df1[15], y=df1[1],label = 'gaussian reward', ci = 95)
sns.lineplot(data=df2, x=df2[15], y=df2[1],label = 'exponential rise and fall', ci = 95)
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
# plt.show()
plt.figure(2)
sns.lineplot(data=df1, x=df1[15], y=df1[9]*10,label = 'gaussian reward')
sns.lineplot(data=df2, x=df2[15], y=df2[9]*10,label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average velocity",fontsize=12)
plt.figure(3)
sns.lineplot(data=df1, x=df1[15], y=df1[11],label = 'gaussian reward')
sns.lineplot(data=df2, x=df2[15], y=df2[11],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("safety count",fontsize=12)
plt.figure(4)
sns.lineplot(data=df1, x=df1[15], y=df1[3],label = 'gaussian reward')
sns.lineplot(data=df2, x=df2[15], y=df2[3],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average episode reward",fontsize=12)
df1[1].loc[df1[15] == 0.5].mean()
# df_new.loc[(df_new['T']>2) & (df_new['T']<7), 'G'].mean()
# print(df1[1].loc[df1[15] == 0.5].mean())
# print(df1[1].loc[df1[15] == 0.75].mean())
# print(df1[1].loc[df1[15] == 1.0].mean())
# print(df1[1].loc[df1[15] == 1.25].mean())
# print(df1[1].loc[df1[15] == 1.5].mean())
print("/n")
# print(df2[1].loc[df1[15] == 0.5].mean())
# print(df2[3].loc[df1[15] == 0.5].mean())
# print(df2[11].loc[df1[15] == 0.5].mean())
# print(df2[9].loc[df1[15] == 0.5].mean()*10)
# print(df2[9].loc[df1[15] == 0.75].mean()*10 )
# print(df2[9].loc[df1[15] == 1.0].mean()* 10)
# print(df2[9].loc[df1[15] == 1.25].mean()*10)
# print(df2[9].loc[df1[15] == 1.5].mean()*10)
# print(df2[9].loc[df1[15] == 1.75].mean()*10)
# print(df2[9].loc[df1[15] == 2.0].mean()*10)
# print("/n")
# print(df2[11].loc[df1[15] == 0.5].mean())
# print(df2[11].loc[df1[15] == 0.75].mean())
# print(df2[11].loc[df1[15] == 1.0].mean())
# print(df2[11].loc[df1[15] == 1.25].mean())
# print(df2[11].loc[df1[15] == 1.5].mean())
# print(df2[11].loc[df1[15] == 1.75].mean())
# print(df2[11].loc[df1[15] == 2.0].mean())
# print("/n")
# print(df1[11].loc[df1[15] == 0.5].mean())
# print(df1[11].loc[df1[15] == 0.75].mean())
# print(df1[11].loc[df1[15] == 1.0].mean())
# print(df1[11].loc[df1[15] == 1.25].mean())
# print(df1[11].loc[df1[15] == 1.5].mean())
# print(df1[11].loc[df1[15] == 1.75].mean())
# print(df1[11].loc[df1[15] == 2.0].mean())
# print("/n")
# print(df2[11].loc[df2[15] == 0.5].mean())
# print(df2[11].loc[df2[15] == 0.75].mean())
# print(df2[11].loc[df2[15] == 1.0].mean())
# print(df2[11].loc[df2[15] == 1.25].mean())
# print(df2[11].loc[df2[15] == 1.5].mean())
# print(df2[11].loc[df2[15] == 1.75].mean())
# print(df2[11].loc[df2[15] == 2.0].mean())
print(df2[3].loc[df2[15] == 0.5].mean())
print(df2[3].loc[df2[15] == 0.75].mean())
print(df2[3].loc[df2[15] == 1.0].mean())
print(df2[3].loc[df2[15] == 1.25].mean())
print(df2[3].loc[df2[15] == 1.5].mean())
print(df2[3].loc[df2[15] == 1.75].mean())
print(df2[3].loc[df2[15] == 2.0].mean())
```
###POSITION BIAS AND VARIANCE
```
## dataframe without " "
# reward 1
f1 = 're_1_bias_1.txt'
f2 = 're_1_bias_2.txt'
f3 = 're_1_bias_3.txt'
f4 = 're_1_bias_4.txt'
f5 = 're_1_bias_5.txt'
f6 = 're_1_bias_6.txt'
re1_pbv1 = get_dataframe(f1) ## 100 points 7 features
re1_pbv2 = get_dataframe(f2)
re1_pbv3 = get_dataframe(f3)
re1_pbv4 = get_dataframe(f4)
re1_pbv5 = get_dataframe(f5)
re1_pbv6 = get_dataframe(f6)
frames = [re1_pbv1,re1_pbv2,re1_pbv3,re1_pbv4,re1_pbv5,re1_pbv6]
df3 = pd.concat(frames)
## reward 2
g1 = 're_2_b1.txt'
g2 = 're_2_b2.txt'
g3 = 're_2_b3.txt'
g4 = 're_2_b4.txt'
g5 = 're_2_b5.txt'
g6 = 're_2_bias_6.txt'
re2_pbv1 = get_dataframe(g1) ## 100 points 7 features
re2_pbv2 = get_dataframe(g2)
re2_pbv3 = get_dataframe(g3)
re2_pbv4 = get_dataframe(g4)
re2_pbv5 = get_dataframe(g5)
re2_pbv6 = get_dataframe(g6)
frames = [re2_pbv1,re2_pbv2,re2_pbv3,re2_pbv4,re2_pbv5,re2_pbv6]
df4 = pd.concat(frames)
xlabels_yo = [75,137.5,200,262.5,325,387.5]
a = []
for i in range(0,101):
a.append(xlabels_yo[0])
for i in range(101,202):
a.append(xlabels_yo[1]) #101 values
for i in range(202,303):
a.append(xlabels_yo[2])
for i in range(303,404):
a.append(xlabels_yo[3])
for i in range(404,505):
a.append(xlabels_yo[4])
for i in range(505,606):
a.append(xlabels_yo[5])
a = np.asarray(a)
df3[15] = a
df4[15] = a
plt.figure(1)
sns.lineplot(data=df3, x=df3[15], y=df3[1],label = 'gaussian reward', ci = 95)
sns.lineplot(data=df4, x=df4[15], y=df4[1],label = 'exponential rise and fall', ci = 95)
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
# plt.show()
plt.figure(2)
sns.lineplot(data=df3, x=df3[15], y=df3[9]*10,label = 'gaussian reward')
sns.lineplot(data=df4, x=df4[15], y=df4[9]*10,label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("average velocity",fontsize=12)
plt.figure(3)
sns.lineplot(data=df3, x=df3[15], y=df3[11],label = 'gaussian reward')
sns.lineplot(data=df4, x=df4[15], y=df4[11],label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("safety count",fontsize=12)
plt.figure(4)
sns.lineplot(data=df3, x=df3[15], y=df3[3],label = 'gaussian reward')
sns.lineplot(data=df4, x=df4[15], y=df4[3],label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("average episode reward",fontsize=12)
# df3.head()
# xlabels_yo = [75,137.5,200,262.5,325,387.5]
print(df3[1].loc[df3[15] == 75].mean())
print(df3[1].loc[df3[15] == 137.5].mean())
print(df3[1].loc[df3[15] == 200].mean())
print(df3[1].loc[df3[15] == 262.5].mean())
print(df3[1].loc[df3[15] == 325].mean())
print(df3[1].loc[df3[15] == 387.5].mean())
print("\n")
print(df3[11].loc[df3[15] == 75].mean())
print(df3[11].loc[df3[15] == 137.5].mean())
print(df3[11].loc[df3[15] == 200].mean())
print(df3[11].loc[df3[15] == 262.5].mean())
print(df3[11].loc[df3[15] == 325].mean())
print(df3[11].loc[df3[15] == 387.5].mean())
print("\n")
print(df3[9].loc[df3[15] == 75].mean()*10)
print(df3[9].loc[df3[15] == 137.5].mean()*10)
print(df3[9].loc[df3[15] == 200].mean()*10)
print(df3[9].loc[df3[15] == 262.5].mean()*10)
print(df3[9].loc[df3[15] == 325].mean()*10)
print(df3[9].loc[df3[15] == 387.5].mean()*10)
print("\n")
print(df3[3].loc[df3[15] == 75].mean())
print(df3[3].loc[df3[15] == 137.5].mean())
print(df3[3].loc[df3[15] == 200].mean())
print(df3[3].loc[df3[15] == 262.5].mean())
print(df3[3].loc[df3[15] == 325].mean())
print(df3[3].loc[df3[15] == 387.5].mean())
print(df4[1].loc[df4[15] == 75].mean())
print(df4[1].loc[df4[15] == 137.5].mean())
print(df4[1].loc[df4[15] == 200].mean())
print(df4[1].loc[df4[15] == 262.5].mean())
print(df4[1].loc[df4[15] == 325].mean())
print(df4[1].loc[df4[15] == 387.5].mean())
print("\n")
print(df4[11].loc[df4[15] == 75].mean())
print(df4[11].loc[df4[15] == 137.5].mean())
print(df4[11].loc[df4[15] == 200].mean())
print(df4[11].loc[df4[15] == 262.5].mean())
print(df4[11].loc[df4[15] == 325].mean())
print(df4[11].loc[df4[15] == 387.5].mean())
print("\n")
print(df4[9].loc[df4[15] == 75].mean()*10)
print(df4[9].loc[df4[15] == 137.5].mean()*10)
print(df4[9].loc[df4[15] == 200].mean()*10)
print(df4[9].loc[df4[15] == 262.5].mean()*10)
print(df4[9].loc[df4[15] == 325].mean()*10)
print(df4[9].loc[df4[15] == 387.5].mean()*10)
print("\n")
print(df4[3].loc[df4[15] == 75].mean())
print(df4[3].loc[df4[15] == 137.5].mean())
print(df4[3].loc[df4[15] == 200].mean())
print(df4[3].loc[df4[15] == 262.5].mean())
print(df4[3].loc[df4[15] == 325].mean())
print(df4[3].loc[df4[15] == 387.5].mean())
```
### VELOCITY VARIANCE
```
## dataframe without " "
# reward 1
f1 = 'val_re_1_variance_1.txt'
f2 = 'val_re_1_variance_2.txt'
f3 = 'val_re_1_variance_3.txt'
f4 = 'val_re_1_variance_4.txt'
f5 = 'val_re_1_variance_5.txt'
f6 = 'val_re_1_variance_6.txt'
f7 = 'val_re_1_variance_7.txt'
re1_vv1 = get_dataframe(f1) ## 100 points 7 features
re1_vv2 = get_dataframe(f2)
re1_vv3 = get_dataframe(f3)
re1_vv4 = get_dataframe(f4)
re1_vv5 = get_dataframe(f5)
re1_vv6 = get_dataframe(f6)
re1_vv7 = get_dataframe(f7)
frames = [re1_vv1,re1_vv2,re1_vv3,re1_vv4,re1_vv5,re1_vv6,re1_vv7]
df5 = pd.concat(frames)
## reward 2
g1 = 're_2_vel_error_variance_1.txt'
g2 = 're_2_vel_error_variance_2.txt'
g3 = 're_2_vel_error_variance_3.txt'
g4 = 're_2_vel_error_variance_4.txt'
g5 = 're_2_vel_error_variance_5.txt'
g6 = 're_2_vel_error_variance_6.txt'
g7 = 're_2_vel_error_variance_7.txt'
re2_vv1 = get_dataframe(g1) ## 100 points 7 features
re2_vv2 = get_dataframe(g2)
re2_vv3 = get_dataframe(g3)
re2_vv4 = get_dataframe(g4)
re2_vv5 = get_dataframe(g5)
re2_vv6 = get_dataframe(g6)
re2_vv7 = get_dataframe(g7)
frames = [re2_vv1,re2_vv2,re2_vv3,re2_vv4,re2_vv5,re2_vv6,re2_vv7]
df6 = pd.concat(frames)
xlabels_yo = [0.5,0.75,1.0,1.25, 1.5,1.75,2.0]
a = []
for i in range(0,101):
a.append(0.5)
for i in range(101,202):
a.append(0.75) #101 values
for i in range(202,303):
a.append(1.0)
for i in range(303,404):
a.append(1.25)
for i in range(404,505):
a.append(1.50)
for i in range(505,606):
a.append(1.75)
for i in range(606,707):
a.append(2.0)
a = np.asarray(a)
df5[15] = a
df6[15] = a
plt.figure(1)
sns.lineplot(data=df5, x=df5[15], y=df5[1],label = 'gaussian reward', ci = 95)
sns.lineplot(data=df6, x=df6[15], y=df6[1],label = 'exponential rise and fall', ci = 95)
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
# plt.show()
plt.figure(2)
sns.lineplot(data=df5, x=df5[15], y=df5[9]*10,label = 'gaussian reward')
sns.lineplot(data=df6, x=df6[15], y=df6[9]*10,label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average velocity",fontsize=12)
plt.figure(3)
sns.lineplot(data=df5, x=df5[15], y=df5[11],label = 'gaussian reward')
sns.lineplot(data=df6, x=df6[15], y=df6[11],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("safety count",fontsize=12)
plt.figure(4)
sns.lineplot(data=df5, x=df5[15], y=df5[3],label = 'gaussian reward')
sns.lineplot(data=df6, x=df6[15], y=df6[3],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average episode reward",fontsize=12)
print(df5[3].loc[df5[15] == 0.5].mean())
print(df5[3].loc[df5[15] == 0.75].mean())
print(df5[3].loc[df5[15] == 1.0].mean())
print(df5[3].loc[df5[15] == 1.25].mean())
print(df5[3].loc[df5[15] == 1.5].mean())
print(df5[3].loc[df5[15] == 1.75].mean())
print(df5[3].loc[df5[15] == 2.0].mean())
print("\n")
print(df5[11].loc[df5[15] == 0.5].mean())
print(df5[11].loc[df5[15] == 0.75].mean())
print(df5[11].loc[df5[15] == 1.0].mean())
print(df5[11].loc[df5[15] == 1.25].mean())
print(df5[11].loc[df5[15] == 1.5].mean())
print(df5[11].loc[df5[15] == 1.75].mean())
print(df5[11].loc[df5[15] == 2.0].mean())
print("\n")
print(df5[1].loc[df5[15] == 0.5].mean())
print(df5[1].loc[df5[15] == 0.75].mean())
print(df5[1].loc[df5[15] == 1.0].mean())
print(df5[1].loc[df5[15] == 1.25].mean())
print(df5[1].loc[df5[15] == 1.5].mean())
print(df5[1].loc[df5[15] == 1.75].mean())
print(df5[1].loc[df5[15] == 2.0].mean())
print("\n")
print(df5[9].loc[df5[15] == 0.5].mean()*10)
print(df5[9].loc[df5[15] == 0.75].mean()*10)
print(df5[9].loc[df5[15] == 1.0].mean()*10)
print(df5[9].loc[df5[15] == 1.25].mean()*10)
print(df5[9].loc[df5[15] == 1.5].mean()*10)
print(df5[9].loc[df5[15] == 1.75].mean()*10)
print(df5[9].loc[df5[15] == 2.0].mean()*10)
print(df6[3].loc[df6[15] == 0.5].mean())
print(df6[3].loc[df6[15] == 0.75].mean())
print(df6[3].loc[df6[15] == 1.0].mean())
print(df6[3].loc[df6[15] == 1.25].mean())
print(df6[3].loc[df6[15] == 1.5].mean())
print(df6[3].loc[df6[15] == 1.75].mean())
print(df6[3].loc[df6[15] == 2.0].mean())
print("\n")
print(df6[11].loc[df6[15] == 0.5].mean())
print(df6[11].loc[df6[15] == 0.75].mean())
print(df6[11].loc[df6[15] == 1.0].mean())
print(df6[11].loc[df6[15] == 1.25].mean())
print(df6[11].loc[df6[15] == 1.5].mean())
print(df6[11].loc[df6[15] == 1.75].mean())
print(df6[11].loc[df6[15] == 2.0].mean())
print("\n")
print(df6[1].loc[df6[15] == 0.5].mean())
print(df6[1].loc[df6[15] == 0.75].mean())
print(df6[1].loc[df6[15] == 1.0].mean())
print(df6[1].loc[df6[15] == 1.25].mean())
print(df6[1].loc[df6[15] == 1.5].mean())
print(df6[1].loc[df6[15] == 1.75].mean())
print(df6[1].loc[df6[15] == 2.0].mean())
print("\n")
print(df6[9].loc[df6[15] == 0.5].mean()*10)
print(df6[9].loc[df6[15] == 0.75].mean()*10)
print(df6[9].loc[df6[15] == 1.0].mean()*10)
print(df6[9].loc[df6[15] == 1.25].mean()*10)
print(df6[9].loc[df6[15] == 1.5].mean()*10)
print(df6[9].loc[df6[15] == 1.75].mean()*10)
print(df6[9].loc[df6[15] == 2.0].mean()*10)
```
### VELOCITY BIAS AND VARIANCE
```
#REWARD 1
f1 = 'val_re_1_bias_and_variance_1.txt'
f2 = 'val_re_1_bias_and_variance_2.txt'
f3 = 'val_re_1_bias_and_variance_3.txt'
f4 = 'val_re_1_bias_and_variance_4.txt'
f5 = 'val_re_1_bias_and_variance_5.txt'
f6 = 'val_re_1_bias_and_variance_6.txt'
f7 = 'val_re_1_bias_and_variance_7.txt'
re1_vbv1 = get_dataframe(f1) ## 100 points 7 features
re1_vbv2 = get_dataframe(f2)
re1_vbv3 = get_dataframe(f3)
re1_vbv4 = get_dataframe(f4)
re1_vbv5 = get_dataframe(f5)
re1_vbv6 = get_dataframe(f6)
re1_vbv7 = get_dataframe(f7)
frames = [re1_vbv1,re1_vbv2,re1_vbv3,re1_vbv4,re1_vbv5,re1_vbv6,re1_vbv7]
df7 = pd.concat(frames)
g1 = 'val_re_2_bias_and_variance_1.txt'
g2 = 'val_re_2_bias_and_variance_2.txt'
g3 = 'val_re_2_bias_and_variance_3.txt'
g4 = 'val_re_2_bias_and_variance_4.txt'
g5 = 'val_re_2_bias_and_variance_5.txt'
g6 = 'val_re_2_bias_and_variance_6.txt'
g7 = 'val_re_2_bias_and_variance_7.txt'
re2_vbv1 = get_dataframe(g1) ## 100 points 7 features
re2_vbv2 = get_dataframe(g2)
re2_vbv3 = get_dataframe(g3)
re2_vbv4 = get_dataframe(g4)
re2_vbv5 = get_dataframe(g5)
re2_vbv6 = get_dataframe(g6)
re2_vbv7 = get_dataframe(g7)
frames = [re2_vbv1,re2_vbv2,re2_vbv3,re2_vbv4,re2_vbv5,re2_vbv6,re2_vbv7]
df8 = pd.concat(frames)
# df8
# re1_vbv7
# df8
xlabels_yo = [45,77.5,110,142.5,175,207.5,240]
a = []
for i in range(0,101):
a.append(xlabels_yo[0])
for i in range(101,202):
a.append(xlabels_yo[1]) #101 values
for i in range(202,303):
a.append(xlabels_yo[2])
for i in range(303,404):
a.append(xlabels_yo[3])
for i in range(404,505):
a.append(xlabels_yo[4])
for i in range(505,606):
a.append(xlabels_yo[5])
for i in range(606,707):
a.append(xlabels_yo[6])
a = np.asarray(a)
df7[15] = a
df8[15] = a
plt.figure(1)
sns.lineplot(data=df7, x=df7[15], y=df7[1],label = 'gaussian reward', ci = 95)
sns.lineplot(data=df8, x=df8[15], y=df8[1],label = 'exponential rise and fall', ci = 95)
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
# plt.show()
plt.figure(2)
sns.lineplot(data=df7, x=df7[15], y=df7[9]*10,label = 'gaussian reward')
sns.lineplot(data=df8, x=df8[15], y=df8[9]*10,label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("average velocity",fontsize=12)
plt.figure(3)
sns.lineplot(data=df7, x=df7[15], y=df7[11],label = 'gaussian reward')
sns.lineplot(data=df8, x=df8[15], y=df8[11],label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("safety count",fontsize=12)
plt.figure(4)
sns.lineplot(data=df7, x=df7[15], y=df7[3],label = 'gaussian reward')
sns.lineplot(data=df8, x=df8[15], y=df8[3],label = 'exponential rise and fall reward')
plt.xlabel("max error as sum of bias and variance in pixels",fontsize=12)
plt.ylabel("average episode reward",fontsize=12)
xlabels_yo = [45,77.5,110,142.5,175,207.5,240]
print(df7[9].loc[df7[15] == 45].mean()*10)
print(df7[9].loc[df7[15] == 77.5].mean()*10)
print(df7[9].loc[df7[15] == 110].mean()*10)
print(df7[9].loc[df7[15] == 142.5].mean()*10)
print(df7[9].loc[df7[15] == 175].mean()*10)
print(df7[9].loc[df7[15] == 207.5].mean()*10)
print(df7[9].loc[df7[15] == 240].mean()*10)
print("\n")
print(df7[1].loc[df7[15] == 45].mean())
print(df7[1].loc[df7[15] == 77.5].mean())
print(df7[1].loc[df7[15] == 110].mean())
print(df7[1].loc[df7[15] == 142.5].mean())
print(df7[1].loc[df7[15] == 175].mean())
print(df7[1].loc[df7[15] == 207.5].mean())
print(df7[1].loc[df7[15] == 240].mean())
print("\n")
print(df7[11].loc[df7[15] == 45].mean())
print(df7[11].loc[df7[15] == 77.5].mean())
print(df7[11].loc[df7[15] == 110].mean())
print(df7[11].loc[df7[15] == 142.5].mean())
print(df7[11].loc[df7[15] == 175].mean())
print(df7[11].loc[df7[15] == 207.5].mean())
print(df7[11].loc[df7[15] == 240].mean())
print("\n")
print(df7[3].loc[df7[15] == 45].mean())
print(df7[3].loc[df7[15] == 77.5].mean())
print(df7[3].loc[df7[15] == 110].mean())
print(df7[3].loc[df7[15] == 142.5].mean())
print(df7[3].loc[df7[15] == 175].mean())
print(df7[3].loc[df7[15] == 207.5].mean())
print(df7[3].loc[df7[15] == 240].mean())
print(df8[9].loc[df8[15] == 45].mean()*10)
print(df8[9].loc[df8[15] == 77.5].mean()*10)
print(df8[9].loc[df8[15] == 110].mean()*10)
print(df8[9].loc[df8[15] == 142.5].mean()*10)
print(df8[9].loc[df8[15] == 175].mean()*10)
print(df8[9].loc[df8[15] == 207.5].mean()*10)
print(df8[9].loc[df8[15] == 240].mean()*10)
print("\n")
print(df8[1].loc[df8[15] == 45].mean())
print(df8[1].loc[df8[15] == 77.5].mean())
print(df8[1].loc[df8[15] == 110].mean())
print(df8[1].loc[df8[15] == 142.5].mean())
print(df8[1].loc[df8[15] == 175].mean())
print(df8[1].loc[df8[15] == 207.5].mean())
print(df8[1].loc[df8[15] == 240].mean())
print("\n")
print(df8[11].loc[df8[15] == 45].mean())
print(df8[11].loc[df8[15] == 77.5].mean())
print(df8[11].loc[df8[15] == 110].mean())
print(df8[11].loc[df8[15] == 142.5].mean())
print(df8[11].loc[df8[15] == 175].mean())
print(df8[11].loc[df8[15] == 207.5].mean())
print(df8[11].loc[df8[15] == 240].mean())
print("\n")
print(df8[3].loc[df8[15] == 45].mean())
print(df8[3].loc[df8[15] == 77.5].mean())
print(df8[3].loc[df8[15] == 110].mean())
print(df8[3].loc[df8[15] == 142.5].mean())
print(df8[3].loc[df8[15] == 175].mean())
print(df8[3].loc[df8[15] == 207.5].mean())
print(df8[3].loc[df8[15] == 240].mean())
```
### VARIANCE BOTH POSITION AND VELOCITY
```
## Reward 1
f1 = 'val_re_1_V_and_P_1.txt'
f2 = 'val_re_1_V_and_P_2.txt'
f3 = 'val_re_1_V_and_P_3.txt'
f4 = 'val_re_1_V_and_P_4.txt'
f5 = 'val_re_1_V_and_P_5.txt'
f6 = 'val_re_1_V_and_P_6.txt'
f7 = 'val_re_1_V_and_P_7.txt'
re1_pvv1 = get_dataframe(f1) ## 100 points 7 features
re1_pvv2 = get_dataframe(f2)
re1_pvv3 = get_dataframe(f3)
re1_pvv4 = get_dataframe(f4)
re1_pvv5 = get_dataframe(f5)
re1_pvv6 = get_dataframe(f6)
re1_pvv7 = get_dataframe(f7)
frames = [re1_pvv1,re1_pvv2,re1_pvv3,re1_pvv4,re1_pvv5,re1_pvv6,re1_pvv7]
df9 = pd.concat(frames)
g1 = 'val_re_2_V_and_P_1.txt'
g2 = 'val_re_2_V_and_P_2.txt'
g3 = 'val_re_2_V_and_P_3.txt'
g4 = 'val_re_2_V_and_P_4.txt'
g5 = 'val_re_2_V_and_P_5.txt'
g6 = 'val_re_2_V_and_P_6.txt'
g7 = 'val_re_2_V_and_P_7.txt'
re2_pvv1 = get_dataframe(g1) ## 100 points 7 features
re2_pvv2 = get_dataframe(g2)
re2_pvv3 = get_dataframe(g3)
re2_pvv4 = get_dataframe(g4)
re2_pvv5 = get_dataframe(g5)
re2_pvv6 = get_dataframe(g6)
re2_pvv7 = get_dataframe(g7)
frames = [re2_pvv1,re2_pvv2,re2_pvv3,re2_pvv4,re2_pvv5,re2_pvv6,re2_pvv7]
df10 = pd.concat(frames)
# df10
# df9
xlabels_yo = [0.5,0.75,1.0,1.25, 1.5,1.75,2.0]
a = []
for i in range(0,101):
a.append(xlabels_yo[0])
for i in range(101,202):
a.append(xlabels_yo[1]) #101 values
for i in range(202,303):
a.append(xlabels_yo[2])
for i in range(303,404):
a.append(xlabels_yo[3])
for i in range(404,505):
a.append(xlabels_yo[4])
for i in range(505,606):
a.append(xlabels_yo[5])
for i in range(606,707):
a.append(xlabels_yo[6])
a = np.asarray(a)
df9[15] = a
df10[15] = a
# sns.lineplot(data=df1, x=xlabels_yo, y="time_stamps_survived")
# sns.lineplot(data = ax,markers=True, dashes=False)
plt.figure(1)
sns.lineplot(data=df9, x=df9[15], y=df9[1],label = 'gaussian reward')
sns.lineplot(data=df10, x=df9[15], y=df10[1],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
plt.legend()
plt.figure(2)
sns.lineplot(data=df9, x=df9[15], y=df9[9]*10,label = 'gaussian reward')
sns.lineplot(data=df10, x=df10[15], y=df10[9]*10,label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average velocity",fontsize=12)
plt.figure(3)
sns.lineplot(data=df9, x=df9[15], y=df9[11],label = 'gaussian reward')
sns.lineplot(data=df10, x=df10[15], y=df10[11],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("safety count",fontsize=12)
plt.figure(4)
sns.lineplot(data=df9, x=df9[15], y=df9[3],label = 'gaussian reward')
sns.lineplot(data=df10, x=df10[15], y=df10[3],label = 'exponential rise and fall reward')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("average episode reward",fontsize=12)
print(df9[3].loc[df9[15] == 0.5].mean())
print(df9[3].loc[df9[15] == 0.75].mean())
print(df9[3].loc[df9[15] == 1.0].mean())
print(df9[3].loc[df9[15] == 1.25].mean())
print(df9[3].loc[df9[15] == 1.5].mean())
print(df9[3].loc[df9[15] == 1.75].mean())
print(df9[3].loc[df9[15] == 2.0].mean())
print("\n")
print(df9[11].loc[df9[15] == 0.5].mean())
print(df9[11].loc[df9[15] == 0.75].mean())
print(df9[11].loc[df9[15] == 1.0].mean())
print(df9[11].loc[df9[15] == 1.25].mean())
print(df9[11].loc[df9[15] == 1.5].mean())
print(df9[11].loc[df9[15] == 1.75].mean())
print(df9[11].loc[df9[15] == 2.0].mean())
print("\n")
print(df9[1].loc[df9[15] == 0.5].mean())
print(df9[1].loc[df9[15] == 0.75].mean())
print(df9[1].loc[df9[15] == 1.0].mean())
print(df9[1].loc[df9[15] == 1.25].mean())
print(df9[1].loc[df9[15] == 1.5].mean())
print(df9[1].loc[df9[15] == 1.75].mean())
print(df9[1].loc[df9[15] == 2.0].mean())
print("\n")
print(df9[9].loc[df9[15] == 0.5].mean()*10)
print(df9[9].loc[df9[15] == 0.75].mean()*10)
print(df9[9].loc[df9[15] == 1.0].mean()*10)
print(df9[9].loc[df9[15] == 1.25].mean()*10)
print(df9[9].loc[df9[15] == 1.5].mean()*10)
print(df9[9].loc[df9[15] == 1.75].mean()*10)
print(df9[9].loc[df9[15] == 2.0].mean()*10)
print(df10[3].loc[df10[15] == 0.5].mean())
print(df10[3].loc[df10[15] == 0.75].mean())
print(df10[3].loc[df10[15] == 1.0].mean())
print(df10[3].loc[df10[15] == 1.25].mean())
print(df10[3].loc[df10[15] == 1.5].mean())
print(df10[3].loc[df10[15] == 1.75].mean())
print(df10[3].loc[df10[15] == 2.0].mean())
print("\n")
print(df10[11].loc[df10[15] == 0.5].mean())
print(df10[11].loc[df10[15] == 0.75].mean())
print(df10[11].loc[df10[15] == 1.0].mean())
print(df10[11].loc[df10[15] == 1.25].mean())
print(df10[11].loc[df10[15] == 1.5].mean())
print(df10[11].loc[df10[15] == 1.75].mean())
print(df10[11].loc[df10[15] == 2.0].mean())
print("\n")
print(df10[1].loc[df10[15] == 0.5].mean())
print(df10[1].loc[df10[15] == 0.75].mean())
print(df10[1].loc[df10[15] == 1.0].mean())
print(df10[1].loc[df10[15] == 1.25].mean())
print(df10[1].loc[df10[15] == 1.5].mean())
print(df10[1].loc[df10[15] == 1.75].mean())
print(df10[1].loc[df10[15] == 2.0].mean())
print("\n")
print(df10[9].loc[df10[15] == 0.5].mean()*10)
print(df10[9].loc[df10[15] == 0.75].mean()*10)
print(df10[9].loc[df10[15] == 1.0].mean()*10)
print(df10[9].loc[df10[15] == 1.25].mean()*10)
print(df10[9].loc[df10[15] == 1.5].mean()*10)
print(df10[9].loc[df10[15] == 1.75].mean()*10)
print(df10[9].loc[df10[15] == 2.0].mean()*10)
```
Acceleration : FUEL EFFICIENCY CURVES
```
f1 = 'val_re_2_V_and_P_1.txt'
f2 = 'val_re_2_V_and_P_2.txt'
f3 = 'val_re_2_V_and_P_3.txt'
f4 = 'val_re_2_V_and_P_4.txt'
f5 = 'val_re_2_V_and_P_5.txt'
f6 = 'val_re_2_V_and_P_6.txt'
# f7 = 'val_re_2_V_and_P_7.txt'
re2_pvv1 = get_dataframe(f1) ## 100 points 7 features
re2_pvv2 = get_dataframe(f2)
re2_pvv3 = get_dataframe(f3)
re2_pvv4 = get_dataframe(f4)
re2_pvv5 = get_dataframe(f5)
re2_pvv6 = get_dataframe(f6)
# re2_pvv7 = get_dataframe(f7)
# frames = [re2_pvv1,re2_pvv2,re2_pvv3,re2_pvv4,re2_pvv5,re2_pvv6,re2_pvv7]
frames = [re2_pvv1,re2_pvv2,re2_pvv3,re2_pvv4,re2_pvv5,re2_pvv6]
df_no_fuell = pd.concat(frames)
g1 = 'val_re_2_V_and_P_1_fuel_eff.txt'
g2 = 'val_re_2_V_and_P_2_fuel_eff.txt'
g3 = 'val_re_2_V_and_P_3_fuel_eff.txt'
g4 = 'val_re_2_V_and_P_4_fuel_eff.txt'
g5 = 'val_re_2_V_and_P_5_fuel_eff.txt'
g6 = 'val_re_2_V_and_P_6_fuel_eff.txt'
# g7 = 'val_re_2_V_and_P_7_fuel_eff.txt'
re2_pvv1_fuell = get_dataframe(g1) ## 100 points 7 features
re2_pvv2_fuell = get_dataframe(g2)
re2_pvv3_fuell = get_dataframe(g3)
re2_pvv4_fuell = get_dataframe(g4)
re2_pvv5_fuell = get_dataframe(g5)
re2_pvv6_fuell = get_dataframe(g6)
# re2_pvv7_fuell = get_dataframe(g7)
# frames = [re2_pvv1_fuell,re2_pvv2_fuell,re2_pvv3_fuell,re2_pvv4_fuell,re2_pvv5_fuell,re2_pvv6_fuell,re2_pvv7_fuell]
frames = [re2_pvv1_fuell,re2_pvv2_fuell,re2_pvv3_fuell,re2_pvv4_fuell,re2_pvv5_fuell,re2_pvv6_fuell]
df_fuell = pd.concat(frames)
# df_fuell
# df_no_fuell
# re2_pvv7
# df_no_fuell
xlabels_yo = [0.75,1.0,1.25, 1.5,1.75,2.0]
a = []
for i in range(0,101):
a.append(xlabels_yo[0])
for i in range(101,202):
a.append(xlabels_yo[1]) #101 values
for i in range(202,303):
a.append(xlabels_yo[2])
for i in range(303,404):
a.append(xlabels_yo[3])
for i in range(404,505):
a.append(xlabels_yo[4])
for i in range(505,606):
a.append(xlabels_yo[5])
a = np.asarray(a)
df_no_fuell[15] = a
df_fuell[15] = a
plt.figure(1)
sns.lineplot(data=df_no_fuell, x=df_no_fuell[15], y=df_no_fuell[1],label = 'general policy')
sns.lineplot(data=df_fuell, x=df_fuell[15], y=df_fuell[1],label = 'fuel efficient policy')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("time stamps survived",fontsize=12)
plt.legend()
plt.figure(2)
sns.lineplot(data=df_no_fuell, x=df_no_fuell[15], y=df_no_fuell[12],label = 'general policy')
sns.lineplot(data=df_fuell, x=df_fuell[15], y=df_fuell[12],label = 'fuel efficient policy')
plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
plt.ylabel("fuel score",fontsize=12)
# plt.figure(3)
# sns.lineplot(data=df9, x=df9[15], y=df9[11],label = 'gaussian reward')
# sns.lineplot(data=df10, x=df10[15], y=df10[11],label = 'exponential rise and fall reward')
# plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
# plt.ylabel("safety count",fontsize=12)
# plt.figure(4)
# sns.lineplot(data=df9, x=df9[15], y=df9[3],label = 'gaussian reward')
# sns.lineplot(data=df10, x=df10[15], y=df10[3],label = 'exponential rise and fall reward')
# plt.xlabel("error as % of pixel width size of the vehicle",fontsize=12)
# plt.ylabel("average episode reward",fontsize=12)
```
| github_jupyter |
A double-ended queue, or dequeue, supports adding and removing elements from either end of the queue.
```
import collections
d = collections.deque('abcdefg')
print('Deque:', d)
print('Length:', len(d))
print('Left end:', d[0])
print('Right end:', d[-1])
# !! c is not at either end of queue
d.remove('c')
print('remove(c):', d)
```
## Populating
A dequeue can be populated from either end, termed "left" and "right" in the python implementation
```
import collections
# Add to the right
d1 = collections.deque()
d1.extend('abcdefg')
print('extend :', d1)
d1.append('h')
print('append :', d1)
# Add to the left
d2 = collections.deque()
d2.extendleft(range(6))
print('extendleft:', d2)
d2.appendleft(6)
print('appendleft:', d2)
```
### extend, extendleft, append, appendleft
* difference between extend and append is same as list
### insert(x,i)
Insert an element at position i
## Consuming and Modifying
Similarly, the elements of the dequeue can be consumed from both ends or either end.
```
import collections
print('From the right:')
d = collections.deque('abcdefg')
while True:
try:
print(d.pop(), end='')
except IndexError:
break
print()
print('\nFrom the left:')
d = collections.deque(range(6))
while True:
try:
print(d.popleft(), end='')
except IndexError:
break
print()
```
### pop(), popleft()
Remove and return an element from the right side of the deque.
### remove
remove the first occurence of a value|
### clear
Remove all the elements from the deque leaving it with length 0
## Rotating and Reversing
### rotate(n)
rotate the deque n steps to the right. if n is negtive, rotate to he left. Rotating one step to the right is equivalent to: `d.appendleft(d.pop())`
```
import collections
d = collections.deque(range(10))
print('Normal :', d)
d = collections.deque(range(10))
d.rotate(2)
print('Right rotation:', d)
d = collections.deque(range(10))
d.rotate(-2)
print('Left rotation :', d)
```
### reverse()
```
import collections
d = collections.deque(range(10))
print('Normal :', d)
d.reverse()
print('Reverse :', d)
```
## Constraining the Queue Size
A dequeue can be configured with a maximum length so that it never grows beyond that size. When the queue reaches the specified length, existing items are discarded as new items are added. This behavior is useful for finding the last n items in a stream of undetermined length.
```
import collections
import random
# Set the random seed so we see the same output each time
# the script is run.
random.seed(1)
d1 = collections.deque(maxlen=3)
d2 = collections.deque(maxlen=3)
for i in range(5):
n = random.randint(0, 100)
print('n =', n)
d1.append(n)
d2.appendleft(n)
print('D1 append :', d1)
print('D2 append left:', d2)
```
## Other method
### copy
create a shallow copy of the dequeue
### count(x)
count the number of dequeue elements equal to x
| github_jupyter |
### Emotions.
In this notebook we are going to create a pytorch model using torchtext and our custom dataset that identifies emotions of a given sentence.
### Emotions:
````
😞 -> sadness
😨 -> fear
😄 -> joy
😮 -> surprise
😍 -> love
😠 -> anger
````
We are going to use our custom dataset that we will load from my google drive.
### Structure of the data.
We have three files which are:
* test.txt
* train.txt
* val.txt
And each of these file contains lines with a respective lable. The text in these files looks as follows:
```txt
im feeling quite sad and sorry for myself but ill snap out of it soon;sadness
i feel like i am still looking at a blank canvas blank pieces of paper;sadness
i feel like a faithful servant;love
```
We will process these text file to come up with json files which is easy to work with when creating our own dataset using torchtext. The following files will be created
* train.json
* test.json
* validation.json
### Imports
```
import json
import time
from prettytable import PrettyTable
import numpy as np
from matplotlib import pyplot as plt
import torch, os, random
from torch import nn
import torch.nn.functional as F
import pandas as pd
```
### Setting seeds
```
SEED = 42
np.random.seed(SEED)
random.seed(SEED)
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.backends.cudnn.deteministic = True
```
### Mounting my Google Drive
```
from google.colab import drive
drive.mount('/content/drive')
data_path = '/content/drive/MyDrive/NLP Data/emotions-nlp'
os.path.exists(data_path)
```
### Loading files lines
```
with open(os.path.join(data_path, 'test.txt'), 'r') as reader:
test_data = reader.read().splitlines()
with open(os.path.join(data_path, 'val.txt'), 'r') as reader:
valid_data = reader.read().splitlines()
with open(os.path.join(data_path, 'train.txt'), 'r') as reader:
train_data = reader.read().splitlines()
```
### Creating `.json` file from these loaded list.
```
train_data_dicts = []
test_data_dicts = []
valid_data_dicts = []
emotions = ['anger', 'fear', 'joy', 'love', 'sadness', 'surprise' ]
emotions_dict = dict([(v, i) for (i, v) in enumerate(emotions)])
for line in test_data:
text, emotion = line.split(';')
test_data_dicts.append({
'text': text,
"emotion_text": emotion,
"emotion": emotions_dict.get(emotion)
})
for line in train_data:
text, emotion = line.split(';')
train_data_dicts.append({
'text': text,
"emotion_text": emotion,
"emotion": emotions_dict.get(emotion)
})
for line in valid_data:
text, emotion = line.split(';')
valid_data_dicts.append({
'text': text,
"emotion_text": emotion,
"emotion": emotions_dict.get(emotion)
})
test_path = 'test.json'
train_path = 'train.json'
valid_path = 'valid.json'
base_path = '/content/drive/MyDrive/NLP Data/emotions-nlp/json'
if not os.path.exists(base_path):
os.makedirs(base_path)
file_object = open(os.path.join(base_path, train_path), 'w')
for line in train_data_dicts:
file_object.write(json.dumps(line))
file_object.write('\n')
file_object.close()
print("train.json created")
file_object = open(os.path.join(base_path, test_path), 'w')
for line in test_data_dicts:
file_object.write(json.dumps(line))
file_object.write('\n')
file_object.close()
print("test.json created")
file_object = open(os.path.join(base_path, valid_path), 'w')
for line in valid_data_dicts:
file_object.write(json.dumps(line))
file_object.write('\n')
file_object.close()
print("valid.json created")
```
### Checking how many example do we have for each set.
```
def tabulate(column_names, data, title):
table = PrettyTable(column_names)
table.title = title
for row in data:
table.add_row(row)
print(table)
data_rows =["training", len(train_data_dicts) ], ["testing", len(test_data_dicts) ], ["validation", len(valid_data_dicts) ]
title = "EXAMPLES IN EACH SET"
column_data = "SET", "EXAMPLES"
tabulate(column_data,data_rows, title )
```
### Preparing the fields
Now that our `.json` files of all the sets looks as follows:
```json
{"text": "i feel a little mellow today", "emotion_text": "joy", "emotion": 2}
```
We are now ready to create the fields .
```
from torchtext.legacy import data
```
We are going to pass `include_lengths=True` to the text Field because we are using padding padded sequences in this notebook. In the label Field we have to specify the datatype as a LongTensor. This is because when doing multiclass classfication pytorch expects the datatype to be a long tensor.
```
TEXT = data.Field(
tokenize="spacy",
include_lengths = True,
tokenizer_language = 'en_core_web_sm'
)
LABEL = data.LabelField()
```
### Creating Field
```
fields ={
"emotion_text": ("emotion", LABEL),
"text": ("text", TEXT)
}
```
### Now we have to create our datasets.
We are going to use the `TabularDataset` to create sets of data for validation, training and testing sets.
```
train_data, test_data, valid_data = data.TabularDataset.splits(
path=base_path,
train=train_path,
test=test_path,
validation = valid_path,
format=train_path.split('.')[-1],
fields=fields
)
print(vars(train_data.examples[2]))
```
### Loading the pretrained word embeddings.
```
MAX_VOCAB_SIZE = 25_000
TEXT.build_vocab(
train_data,
max_size = MAX_VOCAB_SIZE,
vectors = "glove.6B.100d",
unk_init = torch.Tensor.normal_
)
LABEL.build_vocab(train_data)
```
### Device
```
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
```
### Now lets create iterators.
For this we are going to use my fav ``BucketIterator`` to create iterators for each set.
**Note:** - we have to pass a `sort_key` and `sort_within_batch=True` since we are using packed padded sequences otherwise it wont work.
```
sort_key = lambda x: len(x.text)
BATCH_SIZE = 64
train_iterator, valid_iterator, test_iterator = data.BucketIterator.splits(
(train_data, valid_data, test_data),
device = device,
batch_size = BATCH_SIZE,
sort_key = sort_key,
sort_within_batch=True
)
```
### Creating a model.
```
class EmotionsLSTMRNN(nn.Module):
def __init__(self, vocab_size, embedding_size,
hidden_size, output_size, num_layers,
bidirectional, dropout, pad_index
):
super(EmotionsLSTMRNN, self).__init__()
self.embedding = nn.Embedding(vocab_size,embedding_size,
padding_idx=pad_index)
self.lstm = nn.LSTM(embedding_size, hidden_size = hidden_size,
bidirectional=bidirectional, num_layers=num_layers,
dropout = dropout
)
self.hidden_1 = nn.Linear(hidden_size * 2, out_features=512)
self.hidden_2 = nn.Linear(512, out_features=256)
self.output_layer = nn.Linear(256, out_features=output_size)
self.dropout = nn.Dropout(dropout)
def forward(self, text, text_lengths):
embedded = self.dropout(self.embedding(text))
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, text_lengths.to('cpu'), enforce_sorted=False)
packed_output, (h_0, c_0) = self.lstm(packed_embedded)
output, output_lengths = nn.utils.rnn.pad_packed_sequence(packed_output)
h_0 = self.dropout(torch.cat((h_0[-2,:,:], h_0[-1,:,:]), dim = 1))
out = self.dropout(self.hidden_1(h_0))
out = self.hidden_2(out)
return self.output_layer(out)
```
### Creating the Model instance
```
INPUT_DIM = len(TEXT.vocab) # # 25002
EMBEDDING_DIM = 100
HIDDEN_DIM = 256
OUTPUT_DIM = 6
N_LAYERS = 2
BIDIRECTIONAL = True
DROPOUT = 0.5
PAD_IDX = TEXT.vocab.stoi[TEXT.pad_token] # 0
emotions_model = EmotionsLSTMRNN(INPUT_DIM,
EMBEDDING_DIM,
HIDDEN_DIM,
OUTPUT_DIM,
N_LAYERS,
BIDIRECTIONAL,
DROPOUT,
PAD_IDX).to(device)
emotions_model
```
### Counting parameters of the model.
```
def count_trainable_params(model):
return sum(p.numel() for p in model.parameters()), sum(p.numel() for p in model.parameters() if p.requires_grad)
n_params, trainable_params = count_trainable_params(emotions_model)
print(f"Total number of paramaters: {n_params:,}\nTotal tainable parameters: {trainable_params:,}")
```
### Loading pretrained embeddings
```
pretrained_embeddings = TEXT.vocab.vectors
emotions_model.embedding.weight.data.copy_(pretrained_embeddings)
```
### Zeroiing the `pad` and `unk` indices
```
UNK_IDX = TEXT.vocab.stoi[TEXT.unk_token] or TEXT.vocab.stoi["<unk>"]
emotions_model.embedding.weight.data[UNK_IDX] = torch.zeros(EMBEDDING_DIM)
emotions_model.embedding.weight.data[PAD_IDX] = torch.zeros(EMBEDDING_DIM)
emotions_model.embedding.weight.data
```
### Loss and optimizer.
```
optimizer = torch.optim.Adam(emotions_model.parameters())
criterion = nn.CrossEntropyLoss().to(device)
```
### Accuracy function (`categorical_accuracy`).
```
def categorical_accuracy(preds, y):
top_pred = preds.argmax(1, keepdim = True)
correct = top_pred.eq(y.view_as(top_pred)).sum()
acc = correct.float() / y.shape[0]
return acc
```
### Training and evaluation functions.
```
def train(model, iterator, optimizer, criterion):
epoch_loss = 0
epoch_acc = 0
model.train()
for batch in iterator:
optimizer.zero_grad()
text, text_lengths = batch.text
predictions = model(text, text_lengths).squeeze(1)
loss = criterion(predictions, batch.emotion)
acc = categorical_accuracy(predictions, batch.emotion)
loss.backward()
optimizer.step()
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
def evaluate(model, iterator, criterion):
epoch_loss = 0
epoch_acc = 0
model.eval()
with torch.no_grad():
for batch in iterator:
text, text_lengths = batch.text
predictions = model(text, text_lengths)
loss = criterion(predictions, batch.emotion)
acc = categorical_accuracy(predictions, batch.emotion)
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
```
### Training Loop.
We will create a function that will visualize our training loop `ETA` for each and every epoch.
```
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return "{}:{:>02}:{:>05.2f}".format(h, m, s)
def visualize_training(start, end, train_loss, train_accuracy, val_loss, val_accuracy, title):
data = [
["Training", f'{train_loss:.3f}', f'{train_accuracy:.3f}', f"{hms_string(end - start)}" ],
["Validation", f'{val_loss:.3f}', f'{val_accuracy:.3f}', "" ],
]
table = PrettyTable(["CATEGORY", "LOSS", "ACCURACY", "ETA"])
table.align["CATEGORY"] = 'l'
table.align["LOSS"] = 'r'
table.align["ACCURACY"] = 'r'
table.align["ETA"] = 'r'
table.title = title
for row in data:
table.add_row(row)
print(table)
N_EPOCHS = 10
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
start = time.time()
train_loss, train_acc = train(emotions_model, train_iterator, optimizer, criterion)
valid_loss, valid_acc = evaluate(emotions_model, valid_iterator, criterion)
title = f"EPOCH: {epoch+1:02}/{N_EPOCHS:02} {'saving best model...' if valid_loss < best_valid_loss else 'not saving...'}"
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(emotions_model.state_dict(), 'best-model.pt')
end = time.time()
visualize_training(start, end, train_loss, train_acc, valid_loss, valid_acc, title)
```
### Model Evaluation.
```
emotions_model.load_state_dict(torch.load('best-model.pt'))
test_loss, test_acc = evaluate(emotions_model, test_iterator, criterion)
print(f'Test Loss: {test_loss:.3f} | Test Acc: {test_acc*100:.2f}%')
```
### Model Inference
Making predictions.
```
!pip install emoji
import emoji
emotions_emojis = {
'anger' : ":angry:",
'fear': ":fearful:",
'joy' : ":smile:",
'love' : ":heart_eyes:",
'sadness' : ":disappointed:",
'surprise': ":open_mouth:"
}
import spacy
import en_core_web_sm
nlp = en_core_web_sm.load()
def tabulate(column_names, data, title="EMOTION PREDICTIONS TABLE"):
table = PrettyTable(column_names)
table.align[column_names[0]] = "l"
table.align[column_names[1]] = "l"
for row in data:
table.add_row(row)
print(table)
classes = LABEL.vocab.itos
def predict_emotion(model, sentence, min_len = 5):
model.eval()
with torch.no_grad():
tokenized = [tok.text for tok in nlp.tokenizer(sentence)]
if len(tokenized) < min_len:
tokenized += ['<pad>'] * (min_len - len(tokenized))
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
length = [len(indexed)]
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(1)
length_tensor = torch.LongTensor(length)
probabilities = model(tensor, length_tensor)
prediction = torch.argmax(probabilities, dim=1)
class_name = classes[prediction]
emoji_text = emoji.emojize(emotions_emojis[class_name], language='en', use_aliases=True)
prediction = prediction.item()
table_headers =["KEY", "VALUE"]
table_data = [
["PREDICTED CLASS", prediction],
["PREDICTED CLASS NAME", class_name],
["PREDICTED CLASS EMOJI", emoji_text],
]
tabulate(table_headers, table_data)
```
### Sadness
```
predict_emotion(emotions_model, "im updating my blog because i feel shitty.")
```
### Fear
```
predict_emotion(emotions_model, "i am feeling apprehensive about it but also wildly excited")
```
### Joy
```
predict_emotion(emotions_model, "i feel a little mellow today.")
```
### Surprise
```
predict_emotion(emotions_model, "i feel shocked and sad at the fact that there are so many sick people.")
```
### Love
```
predict_emotion(emotions_model, "i want each of you to feel my gentle embrace.")
```
### Anger.
```
predict_emotion(emotions_model, "i feel like my irritable sensitive combination skin has finally met it s match.")
```
> Next we will clone this repository and use conv nets to perform emotions predictions using the same dataset.
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Image/convolutions.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Image/convolutions.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=Image/convolutions.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Image/convolutions.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The following script checks if the geehydro package has been installed. If not, it will install geehydro, which automatically install its dependencies, including earthengine-api and folium.
```
import subprocess
try:
import geehydro
except ImportError:
print('geehydro package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geehydro'])
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once.
```
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
# Load and display an image.
image = ee.Image('LANDSAT/LC08/C01/T1_TOA/LC08_044034_20140318')
Map.setCenter(-121.9785, 37.8694, 11)
Map.addLayer(image, {'bands': ['B5', 'B4', 'B3'], 'max': 0.5}, 'input image')
# Define a boxcar or low-pass kernel.
# boxcar = ee.Kernel.square({
# 'radius': 7, 'units': 'pixels', 'normalize': True
# })
boxcar = ee.Kernel.square(7, 'pixels', True)
# Smooth the image by convolving with the boxcar kernel.
smooth = image.convolve(boxcar)
Map.addLayer(smooth, {'bands': ['B5', 'B4', 'B3'], 'max': 0.5}, 'smoothed')
# Define a Laplacian, or edge-detection kernel.
laplacian = ee.Kernel.laplacian8(1, False)
# Apply the edge-detection kernel.
edgy = image.convolve(laplacian)
Map.addLayer(edgy,
{'bands': ['B5', 'B4', 'B3'], 'max': 0.5},
'edges')
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
# Tools - matplotlib
*This notebook demonstrates how to use the matplotlib library to plot beautiful graphs.*
## Table of Contents
<p><div class="lev1"><a href="#Plotting-your-first-graph"><span class="toc-item-num">1 </span>Plotting your first graph</a></div><div class="lev1"><a href="#Line-style-and-color"><span class="toc-item-num">2 </span>Line style and color</a></div><div class="lev1"><a href="#Saving-a-figure"><span class="toc-item-num">3 </span>Saving a figure</a></div><div class="lev1"><a href="#Subplots"><span class="toc-item-num">4 </span>Subplots</a></div><div class="lev1"><a href="#Multiple-figures"><span class="toc-item-num">5 </span>Multiple figures</a></div><div class="lev1"><a href="#Pyplot's-state-machine:-implicit-vs-explicit"><span class="toc-item-num">6 </span>Pyplot's state machine: implicit <em>vs</em> explicit</a></div><div class="lev1"><a href="#Pylab-vs-Pyplot-vs-Matplotlib"><span class="toc-item-num">7 </span>Pylab <em>vs</em> Pyplot <em>vs</em> Matplotlib</a></div><div class="lev1"><a href="#Drawing-text"><span class="toc-item-num">8 </span>Drawing text</a></div><div class="lev1"><a href="#Legends"><span class="toc-item-num">9 </span>Legends</a></div><div class="lev1"><a href="#Non-linear-scales"><span class="toc-item-num">10 </span>Non linear scales</a></div><div class="lev1"><a href="#Ticks-and-tickers"><span class="toc-item-num">11 </span>Ticks and tickers</a></div><div class="lev1"><a href="#Polar-projection"><span class="toc-item-num">12 </span>Polar projection</a></div><div class="lev1"><a href="#3D-projection"><span class="toc-item-num">13 </span>3D projection</a></div><div class="lev1"><a href="#Scatter-plot"><span class="toc-item-num">14 </span>Scatter plot</a></div><div class="lev1"><a href="#Lines"><span class="toc-item-num">15 </span>Lines</a></div><div class="lev1"><a href="#Histograms"><span class="toc-item-num">16 </span>Histograms</a></div><div class="lev1"><a href="#Images"><span class="toc-item-num">17 </span>Images</a></div><div class="lev1"><a href="#Animations"><span class="toc-item-num">18 </span>Animations</a></div><div class="lev1"><a href="#Saving-animations-to-video-files"><span class="toc-item-num">19 </span>Saving animations to video files</a></div><div class="lev1"><a href="#What-next?"><span class="toc-item-num">20 </span>What next?</a></div>
## Plotting your first graph
First we need to import the `matplotlib` library.
```
import matplotlib
```
Matplotlib can output graphs using various backend graphics libraries, such as Tk, wxPython, etc. When running python using the command line, the graphs are typically shown in a separate window. In a Jupyter notebook, we can simply output the graphs within the notebook itself by running the `%matplotlib inline` magic command.
```
%matplotlib inline
# matplotlib.use("TKAgg") # use this instead in your program if you want to use Tk as your graphics backend.
```
Now let's plot our first graph! :)
```
import matplotlib.pyplot as plt
plt.plot([1, 2, 4, 9, 5, 3])
plt.show()
```
Yep, it's as simple as calling the `plot` function with some data, and then calling the `show` function!
If the `plot` function is given one array of data, it will use it as the coordinates on the vertical axis, and it will just use each data point's index in the array as the horizontal coordinate.
You can also provide two arrays: one for the horizontal axis `x`, and the second for the vertical axis `y`:
```
plt.plot([-3, -2, 5, 0], [1, 6, 4, 3])
plt.show()
```
The axes automatically match the extent of the data. We would like to give the graph a bit more room, so let's call the `axis` function to change the extent of each axis `[xmin, xmax, ymin, ymax]`.
```
plt.plot([-3, -2, 5, 0], [1, 6, 4, 3])
plt.axis([-4, 6, 0, 7])
plt.show()
```
Now, let's plot a mathematical function. We use NumPy's `linspace` function to create an array `x` containing 500 floats ranging from -2 to 2, then we create a second array `y` computed as the square of `x` (to learn about NumPy, read the [NumPy tutorial](tools_numpy.ipynb)).
```
import numpy as np
x = np.linspace(-2, 2, 500)
y = x**2
plt.plot(x, y)
plt.show()
```
That's a bit dry, let's add a title, and x and y labels, and draw a grid.
```
plt.plot(x, y)
plt.title("Square function")
plt.xlabel("x")
plt.ylabel("y = x**2")
plt.grid(True)
plt.show()
```
## Line style and color
By default, matplotlib draws a line between consecutive points.
```
plt.plot([0, 100, 100, 0, 0, 100, 50, 0, 100], [0, 0, 100, 100, 0, 100, 130, 100, 0])
plt.axis([-10, 110, -10, 140])
plt.show()
```
You can pass a 3rd argument to change the line's style and color.
For example `"g--"` means "green dashed line".
```
plt.plot([0, 100, 100, 0, 0, 100, 50, 0, 100], [0, 0, 100, 100, 0, 100, 130, 100, 0], "g--")
plt.axis([-10, 110, -10, 140])
plt.show()
```
You can plot multiple lines on one graph very simply: just pass `x1, y1, [style1], x2, y2, [style2], ...`
For example:
```
plt.plot([0, 100, 100, 0, 0], [0, 0, 100, 100, 0], "r-", [0, 100, 50, 0, 100], [0, 100, 130, 100, 0], "g--")
plt.axis([-10, 110, -10, 140])
plt.show()
```
Or simply call `plot` multiple times before calling `show`.
```
plt.plot([0, 100, 100, 0, 0], [0, 0, 100, 100, 0], "r-")
plt.plot([0, 100, 50, 0, 100], [0, 100, 130, 100, 0], "g--")
plt.axis([-10, 110, -10, 140])
plt.show()
```
You can also draw simple points instead of lines. Here's an example with green dashes, red dotted line and blue triangles.
Check out [the documentation](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) for the full list of style & color options.
```
x = np.linspace(-1.4, 1.4, 30)
plt.plot(x, x, 'g--', x, x**2, 'r:', x, x**3, 'b^')
plt.show()
```
The plot function returns a list of `Line2D` objects (one for each line). You can set extra attributes on these lines, such as the line width, the dash style or the alpha level. See the full list of attributes in [the documentation](http://matplotlib.org/users/pyplot_tutorial.html#controlling-line-properties).
```
x = np.linspace(-1.4, 1.4, 30)
line1, line2, line3 = plt.plot(x, x, 'g--', x, x**2, 'r:', x, x**3, 'b^')
line1.set_linewidth(3.0)
line1.set_dash_capstyle("round")
line3.set_alpha(0.2)
plt.show()
```
## Saving a figure
Saving a figure to disk is as simple as calling [`savefig`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.savefig) with the name of the file (or a file object). The available image formats depend on the graphics backend you use.
```
x = np.linspace(-1.4, 1.4, 30)
plt.plot(x, x**2)
plt.savefig("my_square_function.png", transparent=True)
```
## Subplots
A matplotlib figure may contain multiple subplots. These subplots are organized in a grid. To create a subplot, just call the `subplot` function, and specify the number of rows and columns in the figure, and the index of the subplot you want to draw on (starting from 1, then left to right, and top to bottom). Note that pyplot keeps track of the currently active subplot (which you can get a reference to by calling `plt.gca()`), so when you call the `plot` function, it draws on the *active* subplot.
```
x = np.linspace(-1.4, 1.4, 30)
plt.subplot(2, 2, 1) # 2 rows, 2 columns, 1st subplot = top left
plt.plot(x, x)
plt.subplot(2, 2, 2) # 2 rows, 2 columns, 2nd subplot = top right
plt.plot(x, x**2)
plt.subplot(2, 2, 3) # 2 rows, 2 columns, 3rd subplot = bottow left
plt.plot(x, x**3)
plt.subplot(2, 2, 4) # 2 rows, 2 columns, 4th subplot = bottom right
plt.plot(x, x**4)
plt.show()
```
* Note that `subplot(223)` is a shorthand for `subplot(2, 2, 3)`.
It is easy to create subplots that span across multiple grid cells like so:
```
plt.subplot(2, 2, 1) # 2 rows, 2 columns, 1st subplot = top left
plt.plot(x, x)
plt.subplot(2, 2, 2) # 2 rows, 2 columns, 2nd subplot = top right
plt.plot(x, x**2)
plt.subplot(2, 1, 2) # 2 rows, *1* column, 2nd subplot = bottom
plt.plot(x, x**3)
plt.show()
```
If you need more complex subplot positionning, you can use `subplot2grid` instead of `subplot`. You specify the number of rows and columns in the grid, then your subplot's position in that grid (top-left = (0,0)), and optionally how many rows and/or columns it spans. For example:
```
plt.subplot2grid((3,3), (0, 0), rowspan=2, colspan=2)
plt.plot(x, x**2)
plt.subplot2grid((3,3), (0, 2))
plt.plot(x, x**3)
plt.subplot2grid((3,3), (1, 2), rowspan=2)
plt.plot(x, x**4)
plt.subplot2grid((3,3), (2, 0), colspan=2)
plt.plot(x, x**5)
plt.show()
```
If you need even more flexibility in subplot positioning, check out the [GridSpec documentation](http://matplotlib.org/users/gridspec.html)
## Multiple figures
It is also possible to draw multiple figures. Each figure may contain one or more subplots. By default, matplotlib creates `figure(1)` automatically. When you switch figure, pyplot keeps track of the currently active figure (which you can get a reference to by calling `plt.gcf()`), and the active subplot of that figure becomes the current subplot.
```
x = np.linspace(-1.4, 1.4, 30)
plt.figure(1)
plt.subplot(211)
plt.plot(x, x**2)
plt.title("Square and Cube")
plt.subplot(212)
plt.plot(x, x**3)
plt.figure(2, figsize=(10, 5))
plt.subplot(121)
plt.plot(x, x**4)
plt.title("y = x**4")
plt.subplot(122)
plt.plot(x, x**5)
plt.title("y = x**5")
plt.figure(1) # back to figure 1, current subplot is 212 (bottom)
plt.plot(x, -x**3, "r:")
plt.show()
```
## Pyplot's state machine: implicit *vs* explicit
So far we have used Pyplot's state machine which keeps track of the currently active subplot. Every time you call the `plot` function, pyplot just draws on the currently active subplot. It also does some more magic, such as automatically creating a figure and a subplot when you call `plot`, if they don't exist yet. This magic is convenient in an interactive environment (such as Jupyter).
But when you are writing a program, *explicit is better than implicit*. Explicit code is usually easier to debug and maintain, and if you don't believe me just read the 2nd rule in the Zen of Python:
```
import this
```
Fortunately, Pyplot allows you to ignore the state machine entirely, so you can write beautifully explicit code. Simply call the `subplots` function and use the figure object and the list of axes objects that are returned. No more magic! For example:
```
x = np.linspace(-2, 2, 200)
fig1, (ax_top, ax_bottom) = plt.subplots(2, 1, sharex=True)
fig1.set_size_inches(10,5)
line1, line2 = ax_top.plot(x, np.sin(3*x**2), "r-", x, np.cos(5*x**2), "b-")
line3, = ax_bottom.plot(x, np.sin(3*x), "r-")
ax_top.grid(True)
fig2, ax = plt.subplots(1, 1)
ax.plot(x, x**2)
plt.show()
```
For consistency, we will continue to use pyplot's state machine in the rest of this tutorial, but we recommend using the object-oriented interface in your programs.
## Pylab *vs* Pyplot *vs* Matplotlib
There is some confusion around the relationship between pylab, pyplot and matplotlib. It's simple: matplotlib is the full library, it contains everything including pylab and pyplot.
Pyplot provides a number of tools to plot graphs, including the state-machine interface to the underlying object-oriented plotting library.
Pylab is a convenience module that imports matplotlib.pyplot and NumPy in a single name space. You will find many examples using pylab, but it is no longer recommended (because *explicit* imports are better than *implicit* ones).
## Drawing text
You can call `text` to add text at any location in the graph. Just specify the horizontal and vertical coordinates and the text, and optionally some extra attributes. Any text in matplotlib may contain TeX equation expressions, see [the documentation](http://matplotlib.org/users/mathtext.html) for more details.
```
x = np.linspace(-1.5, 1.5, 30)
px = 0.8
py = px**2
plt.plot(x, x**2, "b-", px, py, "ro")
plt.text(0, 1.5, "Square function\n$y = x^2$", fontsize=20, color='blue', horizontalalignment="center")
plt.text(px - 0.08, py, "Beautiful point", ha="right", weight="heavy")
plt.text(px, py, "x = %0.2f\ny = %0.2f"%(px, py), rotation=50, color='gray')
plt.show()
```
* Note: `ha` is an alias for `horizontalalignment`
For more text properties, visit [the documentation](http://matplotlib.org/users/text_props.html#text-properties).
It is quite frequent to annotate elements of a graph, such as the beautiful point above. The `annotate` function makes this easy: just indicate the location of the point of interest, and the position of the text, plus optionally some extra attributes for the text and the arrow.
```
plt.plot(x, x**2, px, py, "ro")
plt.annotate("Beautiful point", xy=(px, py), xytext=(px-1.3,py+0.5),
color="green", weight="heavy", fontsize=14,
arrowprops={"facecolor": "lightgreen"})
plt.show()
```
You can also add a bounding box around your text by using the `bbox` attribute:
```
plt.plot(x, x**2, px, py, "ro")
bbox_props = dict(boxstyle="rarrow,pad=0.3", ec="b", lw=2, fc="lightblue")
plt.text(px-0.2, py, "Beautiful point", bbox=bbox_props, ha="right")
bbox_props = dict(boxstyle="round4,pad=1,rounding_size=0.2", ec="black", fc="#EEEEFF", lw=5)
plt.text(0, 1.5, "Square function\n$y = x^2$", fontsize=20, color='black', ha="center", bbox=bbox_props)
plt.show()
```
Just for fun, if you want an [xkcd](http://xkcd.com)-style plot, just draw within a `with plt.xkcd()` section:
```
with plt.xkcd():
plt.plot(x, x**2, px, py, "ro")
bbox_props = dict(boxstyle="rarrow,pad=0.3", ec="b", lw=2, fc="lightblue")
plt.text(px-0.2, py, "Beautiful point", bbox=bbox_props, ha="right")
bbox_props = dict(boxstyle="round4,pad=1,rounding_size=0.2", ec="black", fc="#EEEEFF", lw=5)
plt.text(0, 1.5, "Square function\n$y = x^2$", fontsize=20, color='black', ha="center", bbox=bbox_props)
plt.show()
```
## Legends
The simplest way to add a legend is to set a label on all lines, then just call the `legend` function.
```
x = np.linspace(-1.4, 1.4, 50)
plt.plot(x, x**2, "r--", label="Square function")
plt.plot(x, x**3, "g-", label="Cube function")
plt.legend(loc="best")
plt.grid(True)
plt.show()
```
## Non linear scales
Matplotlib supports non linear scales, such as logarithmic or logit scales.
```
x = np.linspace(0.1, 15, 500)
y = x**3/np.exp(2*x)
plt.figure(1)
plt.plot(x, y)
plt.yscale('linear')
plt.title('linear')
plt.grid(True)
plt.figure(2)
plt.plot(x, y)
plt.yscale('log')
plt.title('log')
plt.grid(True)
plt.figure(3)
plt.plot(x, y)
plt.yscale('logit')
plt.title('logit')
plt.grid(True)
plt.figure(4)
plt.plot(x, y - y.mean())
plt.yscale('symlog', linthreshy=0.05)
plt.title('symlog')
plt.grid(True)
plt.show()
```
## Ticks and tickers
The axes have little marks called "ticks". To be precise, "ticks" are the *locations* of the marks (eg. (-1, 0, 1)), "tick lines" are the small lines drawn at those locations, "tick labels" are the labels drawn next to the tick lines, and "tickers" are objects that are capable of deciding where to place ticks. The default tickers typically do a pretty good job at placing ~5 to 8 ticks at a reasonable distance from one another.
But sometimes you need more control (eg. there are too many tick labels on the logit graph above). Fortunately, matplotlib gives you full control over ticks. You can even activate minor ticks.
```
x = np.linspace(-2, 2, 100)
plt.figure(1, figsize=(15,10))
plt.subplot(131)
plt.plot(x, x**3)
plt.grid(True)
plt.title("Default ticks")
ax = plt.subplot(132)
plt.plot(x, x**3)
ax.xaxis.set_ticks(np.arange(-2, 2, 1))
plt.grid(True)
plt.title("Manual ticks on the x-axis")
ax = plt.subplot(133)
plt.plot(x, x**3)
plt.minorticks_on()
ax.tick_params(axis='x', which='minor', bottom='off')
ax.xaxis.set_ticks([-2, 0, 1, 2])
ax.yaxis.set_ticks(np.arange(-5, 5, 1))
ax.yaxis.set_ticklabels(["min", -4, -3, -2, -1, 0, 1, 2, 3, "max"])
plt.title("Manual ticks and tick labels\n(plus minor ticks) on the y-axis")
plt.grid(True)
plt.show()
```
## Polar projection
Drawing a polar graph is as easy as setting the `projection` attribute to `"polar"` when creating the subplot.
```
radius = 1
theta = np.linspace(0, 2*np.pi*radius, 1000)
plt.subplot(111, projection='polar')
plt.plot(theta, np.sin(5*theta), "g-")
plt.plot(theta, 0.5*np.cos(20*theta), "b-")
plt.show()
```
## 3D projection
Plotting 3D graphs is quite straightforward. You need to import `Axes3D`, which registers the `"3d"` projection. Then create a subplot setting the `projection` to `"3d"`. This returns an `Axes3DSubplot` object, which you can use to call `plot_surface`, giving x, y, and z coordinates, plus optional attributes.
```
from mpl_toolkits.mplot3d import Axes3D
x = np.linspace(-5, 5, 50)
y = np.linspace(-5, 5, 50)
X, Y = np.meshgrid(x, y)
R = np.sqrt(X**2 + Y**2)
Z = np.sin(R)
figure = plt.figure(1, figsize = (12, 4))
subplot3d = plt.subplot(111, projection='3d')
surface = subplot3d.plot_surface(X, Y, Z, rstride=1, cstride=1, cmap=matplotlib.cm.coolwarm, linewidth=0.1)
plt.show()
```
Another way to display this same data is *via* a contour plot.
```
plt.contourf(X, Y, Z, cmap=matplotlib.cm.coolwarm)
plt.colorbar()
plt.show()
```
## Scatter plot
To draw a scatter plot, simply provide the x and y coordinates of the points.
```
from numpy.random import rand
x, y = rand(2, 100)
plt.scatter(x, y)
plt.show()
```
You may also optionally provide the scale of each point.
```
x, y, scale = rand(3, 100)
scale = 500 * scale ** 5
plt.scatter(x, y, s=scale)
plt.show()
```
And as usual there are a number of other attributes you can set, such as the fill and edge colors and the alpha level.
```
for color in ['red', 'green', 'blue']:
n = 100
x, y = rand(2, n)
scale = 500.0 * rand(n) ** 5
plt.scatter(x, y, s=scale, c=color, alpha=0.3, edgecolors='blue')
plt.grid(True)
plt.show()
```
## Lines
You can draw lines simply using the `plot` function, as we have done so far. However, it is often convenient to create a utility function that plots a (seemingly) infinite line across the graph, given a slope and an intercept. You can also use the `hlines` and `vlines` functions that plot horizontal and vertical line segments.
For example:
```
from numpy.random import randn
def plot_line(axis, slope, intercept, **kargs):
xmin, xmax = axis.get_xlim()
plt.plot([xmin, xmax], [xmin*slope+intercept, xmax*slope+intercept], **kargs)
x = randn(1000)
y = 0.5*x + 5 + randn(1000)*2
plt.axis([-2.5, 2.5, -5, 15])
plt.scatter(x, y, alpha=0.2)
plt.plot(1, 0, "ro")
plt.vlines(1, -5, 0, color="red")
plt.hlines(0, -2.5, 1, color="red")
plot_line(axis=plt.gca(), slope=0.5, intercept=5, color="magenta")
plt.grid(True)
plt.show()
```
## Histograms
```
data = [1, 1.1, 1.8, 2, 2.1, 3.2, 3, 3, 3, 3]
plt.subplot(211)
plt.hist(data, bins = 10, rwidth=0.8)
plt.subplot(212)
plt.hist(data, bins = [1, 1.5, 2, 2.5, 3], rwidth=0.95)
plt.xlabel("Value")
plt.ylabel("Frequency")
plt.show()
data1 = np.random.randn(400)
data2 = np.random.randn(500) + 3
data3 = np.random.randn(450) + 6
data4a = np.random.randn(200) + 9
data4b = np.random.randn(100) + 10
plt.hist(data1, bins=5, color='g', alpha=0.75, label='bar hist') # default histtype='bar'
plt.hist(data2, color='b', alpha=0.65, histtype='stepfilled', label='stepfilled hist')
plt.hist(data3, color='r', histtype='step', label='step hist')
plt.hist((data4a, data4b), color=('r','m'), alpha=0.55, histtype='barstacked', label=('barstacked a', 'barstacked b'))
plt.xlabel("Value")
plt.ylabel("Frequency")
plt.legend()
plt.grid(True)
plt.show()
```
## Images
Reading, generating and plotting images in matplotlib is quite straightforward.
To read an image, just import the `matplotlib.image` module, and call its `imread` function, passing it the file name (or file object). This returns the image data, as a NumPy array. Let's try this with the `my_square_function.png` image we saved earlier.
```
import matplotlib.image as mpimg
img = mpimg.imread('my_square_function.png')
print(img.shape, img.dtype)
```
We have loaded a 288x432 image. Each pixel is represented by a 4-element array: red, green, blue, and alpha levels, stored as 32-bit floats between 0 and 1. Now all we need to do is to call `imshow`:
```
plt.imshow(img)
plt.show()
```
Tadaaa! You may want to hide the axes when you are displaying an image:
```
plt.imshow(img)
plt.axis('off')
plt.show()
```
It's just as easy to generate your own image:
```
img = np.arange(100*100).reshape(100, 100)
print(img)
plt.imshow(img)
plt.show()
```
As we did not provide RGB levels, the `imshow` function automatically maps values to a color gradient. By default, the color gradient goes from blue (for low values) to red (for high values), but you can select another color map. For example:
```
plt.imshow(img, cmap="hot")
plt.show()
```
You can also generate an RGB image directly:
```
img = np.empty((20,30,3))
img[:, :10] = [0, 0, 0.6]
img[:, 10:20] = [1, 1, 1]
img[:, 20:] = [0.6, 0, 0]
plt.imshow(img)
plt.show()
```
Since the `img` array is just quite small (20x30), when the `imshow` function displays it, it grows the image to the figure's size. Imagine stretching the original image, leaving blanks between the original pixels. How does imshow fill the blanks? Well, by default, it just colors each blank pixel using the color of the nearest non-blank pixel. This technique can lead to pixelated images. If you prefer, you can use a different interpolation method, such as [bilinear interpolation](https://en.wikipedia.org/wiki/Bilinear_interpolation) to fill the blank pixels. This leads to blurry edges, which many be nicer in some cases:
```
plt.imshow(img, interpolation="bilinear")
plt.show()
```
## Animations
Although matplotlib is mostly used to generate images, it is also capable of displaying animations. First, you need to import `matplotlib.animation`.
```
import matplotlib.animation as animation
```
In the following example, we start by creating data points, then we create an empty plot, we define the update function that will be called at every iteration of the animation, and finally we add an animation to the plot by creating a `FuncAnimation` instance.
The `FuncAnimation` constructor takes a figure, an update function and optional arguments. We specify that we want a 50-frame long animation, with 100ms between each frame. At each iteration, `FuncAnimation` calls our update function and passes it the frame number `num` (from 0 to 49 in our case) followed by the extra arguments that we specified with `fargs`.
Our update function simply sets the line data to be the first `num` data points (so the data gets drawn gradually), and just for fun we also add a small random number to each data point so that the line appears to wiggle.
```
x = np.linspace(-1, 1, 100)
y = np.sin(x**2*25)
data = np.array([x, y])
fig = plt.figure()
line, = plt.plot([], [], "r-") # start with an empty plot
plt.axis([-1.1, 1.1, -1.1, 1.1])
plt.plot([-0.5, 0.5], [0, 0], "b-", [0, 0], [-0.5, 0.5], "b-", 0, 0, "ro")
plt.grid(True)
plt.title("Marvelous animation")
# this function will be called at every iteration
def update_line(num, data, line):
line.set_data(data[..., :num] + np.random.rand(2, num) / 25) # we only plot the first `num` data points.
return line,
line_ani = animation.FuncAnimation(fig, update_line, frames=50, fargs=(data, line), interval=100)
plt.close() # call close() to avoid displaying the static plot
```
Next, let's display the animation. One option is to convert it to HTML5 code (using a `<video>` tag), and render this code using `IPython.display.HTML`:
```
from IPython.display import HTML
HTML(line_ani.to_html5_video())
```
Alternatively, we can display the animation using a nice little HTML/Javascript interactive widget:
```
HTML(line_ani.to_jshtml())
```
You can configure Matplotlib to use this widget by default when rendering animations:
```
matplotlib.rc('animation', html='jshtml')
```
After that, you don't even need to use `IPython.display.HTML` anymore:
```
animation.FuncAnimation(fig, update_line, frames=50, fargs=(data, line), interval=100)
```
**Warning:** if you save the notebook along with its outputs, then the animations will take up a lot of space.
## Saving animations to video files
Matplotlib relies on 3rd-party libraries to write videos such as [FFMPEG](https://www.ffmpeg.org/) or [ImageMagick](https://imagemagick.org/). In this example we will be using FFMPEG so be sure to install it first. To save the animation to the GIF format, you would need ImageMagick.
```
Writer = animation.writers['ffmpeg']
writer = Writer(fps=15, metadata=dict(artist='Me'), bitrate=1800)
line_ani.save('my_wiggly_animation.mp4', writer=writer)
```
## What next?
Now you know all the basics of matplotlib, but there are many more options available. The best way to learn more, is to visit the [gallery](http://matplotlib.org/gallery.html), look at the images, choose a plot that you are interested in, then just copy the code in a Jupyter notebook and play around with it.
| github_jupyter |
# Simulação - Malha de controle com controlador PID
## UNIFEI - Universidade Federal de Itajubá
### Prof. Fernando H. D. Guaracy
### ECA602 - 2º Semestre de 2020 - RTE
```
%matplotlib inline
from ipywidgets import interactive
import ipywidgets as widgets
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
from pylab import rcParams
rcParams['figure.figsize'] = 16, 9
#Configurações da simulação
ti = 0.001 #Passo de simulação
x0 = 0 #Valor inicial
sp = 1 #Setpoint
#Modelo do sistema
def modelo(x, t):
global u
x1,x2,x3 = x
dxdt = [x2,x3,-x1-3*x2-3*x3+u]
return dxdt
```
### Resposta do sistema ao degrau unitário em malha aberta
```
vet_t = []
vet_u = []
vet_y = []
xz = [[x0, 0, 0],[x0, 0, 0]]
#Loop de simulação
for i in range(0,20000,1):
u = 1
#----Solução da EDO-------
xz = odeint(modelo, xz[1][:], [0,ti])
#----Armazenamento de dados----
vet_t.append(i*ti)
vet_u.append(u)
vet_y.append(xz[1][0])
#Gera figuras (estáticas)
plt.plot(vet_t,(np.ones(len(vet_t))),'k')
plt.plot(vet_t, vet_y)
plt.ylabel('Amplitude')
plt.xlabel('Tempo [s]')
plt.legend(['Entrada','Saída'])
plt.ylim(top=1.2)
plt.ylim(bottom=-0.2)
plt.grid(True)
```
### Resposta do sistema a uma referência degrau unitário com o controlador PID em malha fechada
```
#Simulação
def simulacao(Kp,Ki,Kd):
#Parâmetros do controlador
#Inicialização de variáveis
global u, vet_t, vet_u, vet_y
vet_t = []
vet_u = []
vet_y = []
c0 = Kp + ti/2*Ki + Kd/ti
c1 = -Kp + ti/2*Ki - 2*Kd/ti
c2 = Kd/ti
u = 0
u_1 = 0
e = 0
e_1 = 0
e_2 = 0
xz = [[x0, 0, 0],[x0, 0, 0]]
#Loop de simulação
for i in range(0,20000,1):
#----Lei de controle------
e = (sp - xz[1][0])
u = u_1 + c0*e + c1*e_1 + c2*e_2
u_1 = u
e_2 = e_1
e_1 = e
#-Fim da lei de controle----
#----Solução da EDO-------
xz = odeint(modelo, xz[1][:], [0,ti])
#----Armazenamento de dados----
vet_t.append(i*ti)
vet_u.append(u)
vet_y.append(xz[1][0])
#Gera figuras (estáticas)
plt.plot(vet_t,(np.ones(len(vet_t))),'k')
plt.plot(vet_t, vet_y)
plt.legend(['Referência','Saída'])
plt.ylabel('Amplitude')
plt.xlabel('Tempo [s]')
plt.grid(True)
plt.show()
Kp_widget=widgets.FloatText(value=2,step=0.1)
Ki_widget=widgets.FloatText(value=0,step=0.1)
Kd_widget=widgets.FloatText(value=0,step=0.1)
#interact(simulacao,Kp=Kp_widget,Ki=Ki_widget,Kd=Kd_widget,continuous_update=False);
interactive_plot = interactive(simulacao,Kp=Kp_widget,Ki=Ki_widget,Kd=Kd_widget,continuous_update=False)
#output = interactive_plot.children[-1]
#output.layout.height = '450px'
interactive_plot
```
| github_jupyter |
# We award Bitcoins/Ethers: Kaggle with ``Gluon`` and k-fold cross-validation
How have you done so far on the journey of `Deep Learning---the Straight Dope`?
> It's time to get your hands dirty and win awards.
To encourage your study of deep learning, we will [award bitcoins/ethers](https://www.reddit.com/r/MachineLearning/comments/70gaqs/we_award_bitcoinether_for_users_who_use/) to users who complete this Kaggle competition with ``Gluon`` and beat a baseline score on Kaggle.
Let's get started.
## Introduction
In this tutorial, we introduce how to use ``Gluon`` for competition on [Kaggle](https://www.kaggle.com). Specifically, we will take the [house price prediction problem](https://www.kaggle.com/c/house-prices-advanced-regression-techniques) as a case study.
We will provide a basic end-to-end pipeline for completing a common Kaggle challenge. We will demonstrate how to use ``pandas`` to pre-process the real-world data sets, such as:
* Handling categorical data
* Handling missing values
* Standardizing numerical features
Note that this tutorial only provides a very basic pipeline. We expect you to tweak the following code, such as re-designing models and fine-tuning parameters, to obtain a desirable score on Kaggle.
## House Price Prediction Problem on Kaggle
[Kaggle](https://www.kaggle.com) is a popular platform for people who love machine learning. To submit the results, please register a [Kaggle](https://www.kaggle.com) account. Please note that, **Kaggle limits the number of daily submissions to 10**.

We take the [house price prediction problem](https://www.kaggle.com/c/house-prices-advanced-regression-techniques) as an example to demonstrate how to complete a Kaggle competition with ``Gluon``. Please learn details of the problem by clicking the [link to the house price prediction problem](https://www.kaggle.com/c/house-prices-advanced-regression-techniques) before starting.

## Load the data set
There are separate training and testing data sets for this competition. Both data sets describe features of every house, such as type of the street, year of building, and basement conditions. Such features can be numeric, categorical, or even missing (`na`). Only the training data set has the sale price of each house, which shall be predicted based on features of the testing data set.
The data sets can be downloaded via the [link to problem](https://www.kaggle.com/c/house-prices-advanced-regression-techniques). Specifically, you can directly access the [training data set](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/download/train.csv) and the [testing data set](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/download/test.csv) after logging in Kaggle.
We load the data via `pandas`. Please make sure that it is installed (``pip install pandas``).
```
import pandas as pd
import numpy as np
train = pd.read_csv("../data/kaggle/house_pred_train.csv")
test = pd.read_csv("../data/kaggle/house_pred_test.csv")
all_X = pd.concat((train.loc[:, 'MSSubClass':'SaleCondition'],
test.loc[:, 'MSSubClass':'SaleCondition']))
```
We can take a look at a few rows of the training data set.
```
train.head()
```
Here is the shapes of the data sets.
```
train.shape
test.shape
```
## Pre-processing data
We can use ``pandas`` to standardiz the numerical features:
$$x_i = \frac{x_i - \mathbb{E} x_i}{\text{std}(x_i)}$$
```
numeric_feas = all_X.dtypes[all_X.dtypes != "object"].index
all_X[numeric_feas] = all_X[numeric_feas].apply(
lambda x: (x - x.mean()) / (x.std()))
```
Let us transform categorical values to numerical ones.
```
all_X = pd.get_dummies(all_X, dummy_na=True)
```
We can approximate the missing values by the mean values of the current feature.
```
all_X = all_X.fillna(all_X.mean())
```
Let us convert the formats of the data sets.
```
num_train = train.shape[0]
X_train = all_X[:num_train].as_matrix()
X_test = all_X[num_train:].as_matrix()
y_train = train.SalePrice.as_matrix()
```
## Loading data in `NDArray`
To facilitate the interactions with ``Gluon``, we need to load data in the `NDArray` format.
```
from mxnet import ndarray as nd
from mxnet import autograd
from mxnet import gluon
X_train = nd.array(X_train)
y_train = nd.array(y_train)
y_train.reshape((num_train, 1))
X_test = nd.array(X_test)
```
We define the loss function to be the squared loss.
```
square_loss = gluon.loss.L2Loss()
```
Below defines the root mean square loss between the logarithm of the predicted values and the true values used in the competition.
```
def get_rmse_log(net, X_train, y_train):
num_train = X_train.shape[0]
clipped_preds = nd.clip(net(X_train), 1, float('inf'))
return np.sqrt(2 * nd.sum(square_loss(
nd.log(clipped_preds), nd.log(y_train))).asscalar() / num_train)
```
## Define the model
We define a **basic** linear regression model here. This may be modified to achieve better results on Kaggle.
```
def get_net():
net = gluon.nn.Sequential()
with net.name_scope():
net.add(gluon.nn.Dense(1))
net.initialize()
return net
```
We define the training function.
```
%matplotlib inline
import matplotlib as mpl
mpl.rcParams['figure.dpi']= 120
import matplotlib.pyplot as plt
def train(net, X_train, y_train, X_test, y_test, epochs,
verbose_epoch, learning_rate, weight_decay):
train_loss = []
if X_test is not None:
test_loss = []
batch_size = 100
dataset_train = gluon.data.ArrayDataset(X_train, y_train)
data_iter_train = gluon.data.DataLoader(
dataset_train, batch_size,shuffle=True)
trainer = gluon.Trainer(net.collect_params(), 'adam',
{'learning_rate': learning_rate,
'wd': weight_decay})
net.collect_params().initialize(force_reinit=True)
for epoch in range(epochs):
for data, label in data_iter_train:
with autograd.record():
output = net(data)
loss = square_loss(output, label)
loss.backward()
trainer.step(batch_size)
cur_train_loss = get_rmse_log(net, X_train, y_train)
if epoch > verbose_epoch:
print("Epoch %d, train loss: %f" % (epoch, cur_train_loss))
train_loss.append(cur_train_loss)
if X_test is not None:
cur_test_loss = get_rmse_log(net, X_test, y_test)
test_loss.append(cur_test_loss)
plt.plot(train_loss)
plt.legend(['train'])
if X_test is not None:
plt.plot(test_loss)
plt.legend(['train','test'])
plt.show()
if X_test is not None:
return cur_train_loss, cur_test_loss
else:
return cur_train_loss
```
## $K$-Fold Cross-Validation
We described the [overfitting problem](../chapter02_supervised-learning/regularization-scratch.ipynb), where we cannot rely on the training loss to infer the testing loss. In fact, when we fine-tune the parameters, we typically rely on $k$-fold cross-validation.
> In $k$-fold cross-validation, we divide the training data sets into $k$ subsets, where one set is used for the validation and the remaining $k-1$ subsets are used for training.
We care about the average training loss and average testing loss of the $k$ experimental results. Hence, we can define the $k$-fold cross-validation as follows.
```
def k_fold_cross_valid(k, epochs, verbose_epoch, X_train, y_train,
learning_rate, weight_decay):
assert k > 1
fold_size = X_train.shape[0] // k
train_loss_sum = 0.0
test_loss_sum = 0.0
for test_i in range(k):
X_val_test = X_train[test_i * fold_size: (test_i + 1) * fold_size, :]
y_val_test = y_train[test_i * fold_size: (test_i + 1) * fold_size]
val_train_defined = False
for i in range(k):
if i != test_i:
X_cur_fold = X_train[i * fold_size: (i + 1) * fold_size, :]
y_cur_fold = y_train[i * fold_size: (i + 1) * fold_size]
if not val_train_defined:
X_val_train = X_cur_fold
y_val_train = y_cur_fold
val_train_defined = True
else:
X_val_train = nd.concat(X_val_train, X_cur_fold, dim=0)
y_val_train = nd.concat(y_val_train, y_cur_fold, dim=0)
net = get_net()
train_loss, test_loss = train(
net, X_val_train, y_val_train, X_val_test, y_val_test,
epochs, verbose_epoch, learning_rate, weight_decay)
train_loss_sum += train_loss
print("Test loss: %f" % test_loss)
test_loss_sum += test_loss
return train_loss_sum / k, test_loss_sum / k
```
## Train and cross-validate the model
The following parameters can be fine-tuned.
```
k = 5
epochs = 100
verbose_epoch = 95
learning_rate = 5
weight_decay = 0.0
```
Given the above parameters, we can train and cross-validate our model.
```
train_loss, test_loss = k_fold_cross_valid(k, epochs, verbose_epoch, X_train,
y_train, learning_rate, weight_decay)
print("%d-fold validation: Avg train loss: %f, Avg test loss: %f" %
(k, train_loss, test_loss))
```
After fine-tuning, even though the training loss can be very low, but the validation loss for the $k$-fold cross validation can be very high. Thus, when the training loss is very low, we need to observe whether the validation loss is reduced at the same time and watch out for overfitting. We often rely on $k$-fold cross-validation to fine-tune parameters.
## Make predictions and submit results on Kaggle
Let us define the prediction function.
```
def learn(epochs, verbose_epoch, X_train, y_train, test, learning_rate,
weight_decay):
net = get_net()
train(net, X_train, y_train, None, None, epochs, verbose_epoch,
learning_rate, weight_decay)
preds = net(X_test).asnumpy()
test['SalePrice'] = pd.Series(preds.reshape(1, -1)[0])
submission = pd.concat([test['Id'], test['SalePrice']], axis=1)
submission.to_csv('submission.csv', index=False)
```
After fine-tuning parameters, we can predict and submit results on Kaggle.
```
learn(epochs, verbose_epoch, X_train, y_train, test, learning_rate,
weight_decay)
```
After executing the above code, a `submission.csv` file will be generated. It is in the required format by Kaggle. Now, we can submit our predicted sales prices of houses based on the testing data set and compare them with the true prices on Kaggle. You need to log in Kaggle, open the [link to the house prediction problem]((https://www.kaggle.com/c/house-prices-advanced-regression-techniques)), and click the `Submit Predictions` button.

You may click the `Upload Submission File` button to select your results. Then click `Make Submission` at the bottom to view your results.

Just a kind reminder again, **Kaggle limits the number of daily submissions to 10**.
## Homework: [We award Bitcoins/Ethers](https://www.reddit.com/r/MachineLearning/comments/70gaqs/we_award_bitcoinether_for_users_who_use/)
* What loss can you obtain on Kaggle by using this tutorial?
* By re-designing the model, fine-tuning the model, and use the $k$-fold cross-validation results, can you beat [the 0.14765 baseline](https://www.kaggle.com/zubairahmed/randomforestregressor-with-score-of-0-14765/) achieved by Random Forest regressor (a powerful model)?
For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| github_jupyter |
# The Collective Benefit of Wearing Masks
### 1. Masks 50% effective in either direction, worn by 50% of people
We're assuming random contacts between individuals in a population. In this example, we're assuming that masks are 50% effective and that 50% of people wear masks. For now, we're also assuming that masks are equally effective in either direction.
Here are the relative risk and likelihoods for all four disease transmission routes.
(😐 = non-mask wearer, 😷 = mask-wearer)
| Transmission Route | Relative Risk | Likelihood |
| :------------- | :----------: | -----------: |
| **😐 -------------> 😐** | $$1$$ | $$0.25$$ |
| **😷 -‖-----------> 😐** | $$0.5$$ | $$0.25$$ |
| **😐 -----------‖-> 😷** | $$0.5$$ | $$0.25$$ |
| **😷 -‖---------‖-> 😷** | $$0.25$$ | $$0.25$$ |
- **Relative Risk to Mask-Wearers:** $$= \frac{(0.5 + 0.25)} {2} = 0.375$$
- **Relative Risk to Non-Mask Wearers:** $$= \frac{(1 + 0.5)} {2} = 0.75$$
- **Average Relative Risk:** $$= \frac{(0.75 + 0.375)} {2} = 0.5625$$
Non-mask wearers get a benefit, mask wearers get a substantial extra benefit, and the public as a whole gets a benefit that’s somewhere in between these two.
### 2. The General Case
Now let's extend the logic above to any values of mask effectiveness & usage.
Let Eout & Ein be the mask effectiveness on exhalation and inhalation, and let p be the fraction of the population that wears a mask.
Here are the relative risk and likelihoods for all four disease transmission routes:
| Transmission Route | Relative Risk of Infection | Likelihood |
| :------------- | :----------: | -----------: |
| **😐 -------------> 😐** | $$1$$ | $$(1-p)^2$$ |
| **😷 -‖-----------> 😐** | $$1-E_{out}$$ | $$p(1-p)$$ |
| **😐 -----------‖-> 😷** | $$1-E_{in}$$ | $$(1-p)p$$ |
| **😷 -‖---------‖-> 😷** | $$(1-E_{out}) \cdot (1-E_{in})$$ | $$p^2$$ |
In order to calculate the average relative risk for each group, we need to compute a weighted average.
- **Average Relative Risk for Non-Mask Wearer**: $$(1 - E_{out} p)$$
- **Average Relative Risk for Mask-Wearer**: $$(1 - E_{out} p) \cdot (1 - E_{in})$$
- **Average Relative Risk:** ([derivation](https://www.wolframalpha.com/input/?i=FullSimplify%5B%281-p%29%5E2+%2B+%281-x%29*p*%281-p%29+%2B+%281-y%29*p*%281-p%29+%2B+%281-x%29*%281-y%29*p*p%5D)) $$(1 - E_{in} p) \cdot (1 - E_{out} p)$$
- [Interactive graph of above](https://www.desmos.com/calculator/cqsz9lvprf)
If the mask is equally effective in either direction, $$E_{out} = E_{in} = E$$ and so the Average Relative Risk for population becomes $$(1 - E \cdot p)^2$$
So for a 50% effective mask worn by 50% of the population, the relative risk is multiplied by a factor $$=(1 - 0.5*0.5)^2 = 0.5625$$ which is the same value in the simple case above. However we can now calculate this for any E & p. Notice that the factor simply depends on the product of mask effectiveness E & wearership p.
We can interpret this average relative risk as a multiplier on the [effective contact rate](https://en.wikipedia.org/wiki/Transmission_risks_and_rates) due to mask wearing. Since R0 is proportional to the effective contact rate, R0 will vary by the same factor.
$$R_{0}\text{(with masks)} = R_0 \cdot (1 - E_{in} p) \cdot (1 - E_{out} p)$$
```
# https://www.geeksforgeeks.org/3d-surface-plotting-in-python-using-matplotlib/
# Import libraries
from mpl_toolkits import mplot3d
import numpy as np
import matplotlib.pyplot as plt
# Creating dataset
x = np.outer(np.linspace(0, 1, 32), np.ones(32))
y = x.copy().T # transpose
z = 2.5 * (1-x*y)**2
# Creating figyre
fig = plt.figure(figsize =(14, 9))
ax = plt.axes(projection ='3d')
# Creating color map
my_cmap = plt.get_cmap('hot')
# Creating plot
surf = ax.plot_surface(x, y, z,
rstride = 1,
cstride = 1,
alpha = 1,
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='z',
offset = np.min(z),
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='x',
offset =-5,
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='y',
offset = 5,
cmap = my_cmap)
fig.colorbar(surf, ax = ax,
shrink = 0.5,
aspect = 5)
# Adding labels
ax.set_xlabel('Mask Wearership', fontsize = 16)
ax.set_xlim(0, 1)
ax.set_ylabel('Mask Effectiveness', fontsize = 16)
ax.set_ylim(0, 1)
ax.set_zlabel('R0', fontsize = 16)
ax.set_zlim(np.min(z), np.max(z))
ax.set_title('Effect of Masks on R0 (Assuming R0 = 2.5)', fontsize = 20)
# show plot
plt.show()
```
Note that the quadratic dependence leads to a steeper than linear falloff on increasing p, i.e. a small increase in p has a quadratic effect on lowering R0. If 70% of people wore a 70% effective mask, the effect would be to lower R0 by a factor of 4.
### 3. From R0 to Probability of Infection
Assuming a SIR model, we can use the relation $$R_0 = -log(1- P)/P$$ where P is the fraction of the population that will eventually be infected, or equivalently, the probability that an individual will eventually be infected (see [Brienen et al](https://onlinelibrary.wiley.com/doi/full/10.1111/j.1539-6924.2010.01428.x) or page 27 of [this pdf](https://web.stanford.edu/~jhj1/teachingdocs/Jones-Epidemics050308.pdf) for derivation).
This allows us to transform this graph into a graph of infection probability (given a particular R0, mask efficacy E and mask wearership p).
The [solution](https://www.wolframalpha.com/input/?i=Solve%5BR0+%3D+-log%281-+P%29%2FP%2C+P%5D) involves the Lambert W function.
$$P = 1 + \frac{W(R e^{-R})}{R}$$
where $R$ is the value of $R_0$ reduced by mask wearing, i.e.
$$R = R_0 \cdot (1 - E_{in} p) \cdot (1 - E_{out} p)$$
```
from scipy.special import lambertw
# Creating dataset
eps = 10**-6
x = np.outer(np.linspace(0, 1, 32), np.ones(32)) + eps
y = x.copy().T # transpose
R = 2.5 * (1-x*y)**2
z = 1 + np.real(lambertw(-R * np.exp(-R)))/R
# Creating figyre
fig = plt.figure(figsize =(14, 9))
ax = plt.axes(projection ='3d')
# Creating color map
my_cmap = plt.get_cmap('hot')
# Creating plot
surf = ax.plot_surface(x, y, z,
rstride = 1,
cstride = 1,
alpha = 1,
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='z',
offset = 0,
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='x',
offset =-5,
cmap = my_cmap)
cset = ax.contourf(x, y, z,
zdir ='y',
offset = 5,
cmap = my_cmap)
fig.colorbar(surf, ax = ax,
shrink = 0.5,
aspect = 5)
# Adding labels
ax.set_xlabel('Mask Wearership', fontsize = 16)
ax.set_xlim(0, 1)
ax.set_ylabel('Mask Effectiveness', fontsize = 16)
ax.set_ylim(0, 1)
ax.set_zlabel('Probability of Infection', fontsize = 16)
ax.set_zlim(0, 1)
ax.set_title('Effect of Masks on Infection Probability (Assuming R0 = 2.5)', fontsize = 20)
# show plot
plt.show()
```
So the probability of infection has a steeper dependence on mask wearership & mask efficacy, as compared to R0. In particular there is a critical transition when R0 falls below 1. In this region (shown in black above) mask wearing can end an epidemic.
### 4. Non-Random Population Assortment Due To An In-Group Preference
Now let's consider what happens when mask-wearers have a preference for interacting with mask-wearers, and non-mask wearers have a preference for interacting with non-mask wearers.
We can model this with a term epsilon, which represent the bias for a person towards interacting with their own group, and against interacting with the other group.
For simplicity, we'll assume that the in-group preference (and out-group aversion) is the same for mask-wearers and non-mask wearers. This can easily be extended, but our purpose here is to get a basic idea of how the risk changes with in-group preference.
In this case, the relative risk and likelihoods for all four disease transmission routes are modified as follows:
| Transmission Route | Relative Risk of Infection | Likelihood |
| :------------- | :----------: | -----------: |
| **😐 -------------> 😐** | $$1$$ | $$(1-p)^2 + \epsilon$$ |
| **😷 -‖-----------> 😐** | $$1-E_{out}$$ | $$p(1-p) - \epsilon$$ |
| **😐 -----------‖-> 😷** | $$1-E_{in}$$ | $$(1-p)p - \epsilon$$ |
| **😷 -‖---------‖-> 😷** | $$(1-E_{out}) \cdot (1-E_{in})$$ | $$p^2 + \epsilon$$ |
The sum of probabilites of each case still adds to 1. However a positive epsilon biases the system towards in-group preference.
We can now recalculate the average relative risk for each group.
- **Average Relative Risk for Non-Mask Wearer**: $$(1 - E_{out} p) + \frac{\epsilon E_{out}}{p}$$
So the risk for non-mask wearers **increases** due to an in-group preference, which makes intuitive sense, as they are less likely to benefit from mask-wearers.
- **Average Relative Risk for Mask-Wearer**: $$(1 - E_{out} p) \cdot (1 - E_{in}) - \frac{\epsilon E_{out} (1-E_{in})}{p}$$
So the risk for mask-wearers **decreases** due to an in-group preference, which also makes sense, as they are more likely to benefit from other mask-wearers.
The interesting question is what happens in the overall population. Does the benefit to mask wearers due to an in-group preference outweigh the cost to non-mask wearers, or vice-versa?
- **Average Relative Risk:** $$(1 - E_{in} p) \cdot (1 - E_{out} p) + \epsilon E_{in} E_{out}$$
This teaches us that when each group has an in-group preference, the cost to non-mask wearers outweighs the benefit to mask wearers. As a result, the average risk for the population **increases** by an amount that depends on the extent of in-group preference and on the mask effectiveness.
$$R_{0}\text{(with masks and in-group preference)} = R_0 \cdot \big((1 - E_{in} p) \cdot (1 - E_{out} p) + \epsilon E_{out} E_{in}\big)$$
If the in-group preference epsilon goes to zero, we recover the results in section 2.
### 5. Comparison with Existing Literature
This model is similar to the one put forth by [Tian et al](https://arxiv.org/ftp/arxiv/papers/2003/2003.07353.pdf) (Table S3 & Fig S4), also referenced in [Howard et al](https://www.preprints.org/manuscript/202004.0203/v1) (Fig 1), and is consistent with the simulation results of [Eikenberry et al](https://www.sciencedirect.com/science/article/pii/S2468042720300117). [Fisman et al](https://www.sciencedirect.com/science/article/pii/S2468042720300191) also use a similar model of bidirectionally effective masks.
However, other work such as [Ngonghala et al](https://www.sciencedirect.com/science/article/pii/S0025556420300560) (Fig 8), [Stutt et all](https://royalsocietypublishing.org/doi/pdf/10.1098/rspa.2020.0376) (Fig 3 & 4), or [Brienen et al](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC7169241/) (Fig 1) find a linear relationship between R0 and E*p (product of mask effectiveness and wearership) rather than the quadratic relationship shown above. This is likely because they either implicitly or explicitly assume masks work in only one direction. We can recreate this linear relationship by manually setting either Ein or Eout = 0.
| github_jupyter |
# Module 7: APIs and Scraping
In today's module, we will learn about APIs and scraping.
An API is an application programming interface. It provides a structured way to send commands or requests to a piece of software. "API" often refers to a web service API. This is like web site (but designed for applications, rather than humans, to use) that you can send requests to execute commands or query for data. To use them, you can send them a request, and they reply with a response, similar to how a web browser works. The request is sent to an endpoint (a URL) typically with a set of parameters to provide the details of your query or command.
Park. 2019. [How Do APIs Work?](https://tray.io/blog/how-do-apis-work) Tray.io.
"Imagine you’re a customer at a restaurant. The waiter (the API) functions as an intermediary between customers like you (the user) and the kitchen (web server). You tell the waiter your order (API call), and the waiter requests it from the kitchen. Finally, the waiter will provide you with what you ordered." -- Park. 2019.
Tools:
1. [requests](https://docs.python-requests.org/en/master/): a Python packadge for retrieving web resources
2. [beautifulsoup](https://www.crummy.com/software/BeautifulSoup/bs4/doc/): a Python packadge for traversing and extracting elements from a web page HTML and XML files.
Today we are going to work through some examples of using API for requesting government data, and basic webscraping when API is not available:
* [Census Data API](#census)
* [Data Portals API](#local)
* [Web scraping](#scraping)
(Side note: I'm demoing a clickable table of contents here. Give it a look to see how to do this in markdown.)
```
from bs4 import BeautifulSoup
import requests
import pandas as pd
import seaborn as sns
import time
import json
import re
```
<a id='census'></a>
## 1. Census Data API
Traditionally, you may search, access and download census data from the website like [data.census.gov](https://data.census.gov/cedsci/). The U.S. Census Bureau provides [Census Data API](https://www.census.gov/data/developers/guidance/api-user-guide.Overview.html) to allow users to query data directly from Census Bureau servers. The Census Data API is a more efficient and flexible way to query large amount of data for target variables and geographies, compared to traditional webpage interface.
To follow along with this sction, you need a working API key to use the Census API Service. You can register for an API key, free of charge by following these steps:
1. Go to the [Developers site](https://www.census.gov/data/developers.html)
2. Click on the Request a KEY box on the left side of the page.
3. Fill out the pop-up window form.
4. You will receive an email with your key code in the message and a link to register it.
5. Then save your unique key code in a Python script named `keys.py` with information like `census_api_key = 'your-key-code'`
```
# import your KEY inforamtion
# The following code imports only the census_api_key variable from the keys module:
from keys import census_api_key
# which census dataset, in this example, we use ACS 1-year estimate profile dataset
dataset = 'acs/acs1/profile'
# which census variables to retrieve
variables = {'DP05_0033E' : 'tot_pop',
'DP05_0071E' : 'hispanic',
'DP05_0044E' : 'asian',
'DP05_0037E' : 'white',
'DP05_0038E' : 'black'}
# the data year your want to query
year = 2019
# the api key you get from the Census
api_key = census_api_key
```
You need to know the census dataset file well for requesting data via their API service:
- Variables you can query for from ACS: https://api.census.gov/data/2017/acs/acs5/profile/variables.html
https://api.census.gov/data/2017/acs/acs5/subject/variables.html
```
# convert vars to string to send to api
variables_str = ','.join(variables)
variables_str
# in this case, we want to obtain total population for all variables in all US metropolitan and micropolitan statistical areas
url_template = 'https://api.census.gov/data/{year}/{dataset}?' \
'get=NAME,{variable}&for=metropolitan%20statistical%20area/micropolitan%20statistical%20area:*&key={api_key}'
url = url_template.format(year=year, dataset=dataset, variable=variables_str, api_key=api_key)
# take a look at the url to see what data you are expecting
#url
```
You can check the example url for acs data: https://api.census.gov/data/2019/acs/acs1/examples.html, in this case we focus on the metropolitan area example.
```
# request the data by sending a get request
response = requests.get(url, timeout=30)
# parse the json string into a Python dict
json_data = response.json()
```
[JSON](https://en.wikipedia.org/wiki/JSON) is a standardized format the community uses to pass data around. People often use JSON to store and exchange data in websites (they look a lot like looks like a Python dictionary format). But keep in mind, JSON isn’t the only format available for this kind of work, other common ones include XML and YAML.
```
#type(response)
#response.status_code
#print(response.text)
#json_data
# load as dataframe
df = pd.DataFrame(json_data)
df.head()
#df.rename(columns=df.iloc[0])
# set the first row as column name
df = df.rename(columns=df.iloc[0]).drop(df.index[0])
df.head()
# let's rename the dataset to a more intuitive column name
df = df.rename(columns=variables)
df.head()
def get_census_variableprofile_descriptions(dataset, year, variables):
"""Download descriptions of census variables from the API
"""
url_template = 'https://api.census.gov/data/{year}/{dataset}/profile/variables/{variable}.json'
variable_descriptions = {}
for variable in variables:
url = url_template.format(year=year, dataset=dataset, variable=variable)
response = requests.get(url)
data = response.json()
variable_descriptions[variable] = {'concept':data['concept'],
'label':data['label']}
return variable_descriptions
# download and display census descriptions of each variable
variable_descriptions = get_census_variableprofile_descriptions(dataset='acs/acs1',
year=year,
variables=variables)
for v, d in variable_descriptions.items():
print('{}\t{}'.format(variables[v], d['label']))
```
The Census provides very well documented API user guide, check on [Example API queries page](https://www.census.gov/data/developers/guidance/api-user-guide.Example_API_Queries.html), and feel free to experiment and query [available data](https://www.census.gov/data/developers/guidance/api-user-guide.Available_Data.html) on your own.
<a id='local'></a>
## 2. Data Portals
Many governments and agencies now open up their data to the public through a data portal. These often offer APIs to query them for real-time data.
Using the Cambridge Open Data Portal... browse the portal for public datasets: https://data.cambridgema.gov/browse
First we'll look at crime report data in Cambridge: https://data.cambridgema.gov/Public-Safety/Crime-Reports/xuad-73uj
### 2.1. Crime report
```
# define API endpoint
endpoint_url = 'https://data.cambridgema.gov/resource/xuad-73uj.json'
# request the URL and download its response
response = requests.get(endpoint_url)
# parse the json string into a Python dict
data = response.json()
len(data)
```
There are more than 1000 rows in the dataset, but we're limited by the API to only 1000 per request. We have to use pagination to get the rest.
```
# recursive function to keep requesting more rows until there are no more
def request_data(endpoint_url, limit=1000, offset=0, data=[]):
"""Request all data from the API until there are no more
"""
url = endpoint_url + '?$limit={limit}&$offset={offset}'
request_url = url.format(limit=limit, offset=offset)
response = requests.get(request_url)
rows = response.json()
data.extend(rows)
if len(rows) >= limit:
data = request_data(endpoint_url, offset=offset+limit, data=data)
return data
# get all the data from the API, using our recursive function
endpoint_url = 'https://data.cambridgema.gov/resource/xuad-73uj.json'
data = request_data(endpoint_url)
len(data)
# turn the json data into a dataframe
df = pd.DataFrame(data)
df.shape
# what columns are in our data?
df.columns
#count and plot the number of crime by neighborhood
ax = sns.countplot(data=df, y='neighborhood')
```
You can use any web service's API in this same basic way: request the URL with some parameters. Read the API's documentation to know how to use it and what to send. You can also use many web service's through a Python package to make complex services easier to work with. For example, there's a fantastic package called [cenpy](https://cenpy-devs.github.io/cenpy/) that makes downloading and working with US census data super easy.
<a id='scraping'></a>
## 3. Web Scraping
If you need data from a web page that doesn't offer an API, you can scrape it. Note that many web sites prohibit scraping in their terms of use, so proceed respectfully and cautiously. Web scraping means downloading a web page, parsing individual data out of its HTML, and converting those data into a structured dataset.
For straightforward web scraping tasks, you can use the powerful [BeautifulSoup](https://www.crummy.com/software/BeautifulSoup/bs4/doc/) package, but it may not work for some other complex web pages...
In this example, we'll scrape https://en.wikipedia.org/wiki/List_of_National_Basketball_Association_arenas
The first step in scraping is figuring out a way to identify the information you want.
```
url = 'https://en.wikipedia.org/wiki/List_of_National_Basketball_Association_arenas'
response = requests.get(url)
html = response.text
# look at the html string
html[5000:7000]
```
You could manually search for the text and extract them but it would be really cumbersome.
The BeautifulSoup makes this much easier by converting the text into a data structure that is easy to “traverse”
```
# parse the html
soup = BeautifulSoup(html, features='html.parser')
#soup
# use the find_all() function to find all tags of a paticular type
# in this case, we find all the table body, then we find all table row within the table body
rows = soup.find('tbody').findAll('tr')
#rows
rows[1]
```
[HTML Table](https://www.w3schools.com/html/html_tables.asp): `<tbody>` groups the body content in a table. Each table row is defined with a `<tr>` tag. Each table header is defined with a `<th>` tag. Each table data/cell is defined with a `<td>` tag.
```
data = []
for row in rows[1:]:
cells = row.findAll('td')
d = [cell.text.strip('\n') for cell in cells[1:-1]] #The '\n' character is used to find a newline character.
#print(cells[1:-1])
#print(d)
data.append(d)
#data
cols = ['arena', 'city', 'team', 'capacity', 'opened']
df = pd.DataFrame(data=data, columns=cols).dropna()
df
# strip out all the wikipedia notes in square brackets
# The applymap() function is used to apply a function to every elements in a Dataframe
# lambda functions let you write write functions in a quick way
# We can pass lambda function to the applymap() without even naming them, like an anonymous functions.
# the re module provide regular experssion matching in Python
df = df.applymap(lambda x: re.sub(r'\[.\]', '', x))
df
# convert capacity and opened to integer
df['capacity'] = df['capacity'].str.replace(',', '')
df[['capacity', 'opened']] = df[['capacity', 'opened']].astype(int)
df.sort_values('capacity', ascending=False)
```
Web scraping is really hard! You need to understand the website structure well and it takes lots of practice. If you want to use it, read the BeautifulSoup and Selenium documentation carefully, follow examples, and then practice, practice, practice. You'll be an expert before long.
## This week's assignment
Starting this week, you will start thinking about your final project. You are asked to submit a final project proposal following the instruction posted on course website, and also submit at least one dataset you want to use for your final project, from data APIs, public data portals, FTP servers, or directly from an organization etc.
The very first step you need to do is to think about a topic that interests you and explore useful dataset. Below are some datasets for your inspiration.
## Data sources, API, data repository for inspiration:
- US Census and Geographies
- American Community Survey, county business pattern, or Decennial Census data you can download from [data.census.gov](https://data.census.gov/cedsci/); request via [Census Data API](https://www.census.gov/data/developers/guidance/api-user-guide.Overview.html); or via [cenpy](https://cenpy-devs.github.io/cenpy/)
- [IPUMS Census and Survey Data](https://ipums.org/): a great data repository for standardize census microdata across time and space, and other health survey and GIS data from around the world
- Census geospatial data products from [TIGER](https://www.census.gov/cgi-bin/geo/shapefiles/index.php), or from Census API [censusviz package](https://pypi.org/project/censusviz/)
- US economic data:
- [Bureau of Labor Statistics](https://www.bls.gov/developers/) for employment and labor market data
- [Bureau of Economic Analysis](https://www.bea.gov/data) for GDP, income, industrial data etc.
- [Clustering Mapping Project](https://clustermapping.us/region) if you are doing regional economic analysis
- US local open data portals:
- [Cambridge](https://data.cambridgema.gov/browse)
- [Boston](https://data.boston.gov/); Boston Data Portal from [BARI](https://cssh.northeastern.edu/bari/boston-data-portal/)
- [Chicago](https://data.cityofchicago.org/)
- [New York](https://data.cityofnewyork.us/browse?provenance=official&sortBy=most_accessed&utf8=%E2%9C%93)
- [San Francisco](https://datasf.org/opendata/developers/)
- [Los Angeles](https://data.lacity.org/browse)
- U.S. DEPARTMENT OF AGRICULTURE ([USDA](https://www.ers.usda.gov/data-products/)) has great natural resources, environment, food and economic data
- [National Longitudinal Survey of Youth](https://www.nlsinfo.org/content/cohorts) if you are interested in youth development
- [FBI Crime Data](https://crime-data-explorer.fr.cloud.gov/), including an API (can be used for geospatial analyses)
- Global indicator repository for health, sustainability, environment, and urban developemnt indicators around the world:
- [World Bank Development Indicators](https://datahelpdesk.worldbank.org/knowledgebase/topics/125589)
- [WHO Global Health Observatory](https://apps.who.int/gho/data/node.home)
- [UN Habitat Urban Indicators Database](https://data.unhabitat.org/)
- [The Data-Driven EnviroLab](https://datadrivenlab.org/urban/)
- List of APIs from [Data.gov](https://api.data.gov/list-of-apis/)
- Other APIs you may be interested:
- [Twitter API](https://developer.twitter.com/en/docs)
- [google map API](https://developers.google.com/maps/documentation/)
- [National Weather Service API](https://www.weather.gov/documentation/services-web-api)
- [OpenStreetMap](https://www.openstreetmap.org/#map=5/38.007/-95.844) offers geospatial data worldwide, you can request via [Overpass API](https://wiki.openstreetmap.org/wiki/Overpass_API), or via python packadge [OSMnx](https://github.com/gboeing/osmnx)
- [New York Times API](https://developer.nytimes.com/apis)
Also check out some examples notebooks from https://github.com/jupyter/jupyter/wiki/A-gallery-of-interesting-Jupyter-Notebooks#machine-learning-statistics-and-probability and https://anaconda.org/gallery for inspiration to help you move towards final project
| github_jupyter |
# WeatherPy
----
#### Note
* Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
```
# Dependencies and Setup
import requests
import certifi
import json
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import time
from scipy.stats import linregress
from datetime import date
today = date.today()
# Import API key
from api_keys import weather_api_key as api_key
# Incorporated citipy to determine city based on latitude and longitude
from citipy import citipy
# Output File (CSV)
output_data_file = "output_data/cities.csv"
# Range of latitudes and longitudes
lat_range = (-90, 90)
lng_range = (-180, 180)
```
## Generate Cities List
```
# List for holding lat_lngs and cities
lat_lngs = []
cities = []
# Create a set of random lat and lng combinations
lats = np.random.uniform(lat_range[0], lat_range[1], size=1500)
lngs = np.random.uniform(lng_range[0], lng_range[1], size=1500)
lat_lngs = zip(lats, lngs)
# Identify nearest city for each lat, lng combination
for lat_lng in lat_lngs:
city = citipy.nearest_city(lat_lng[0], lat_lng[1]).city_name
# If the city is unique, then add it to a our cities list
if city not in cities:
cities.append(city)
# Print the city count to confirm sufficient count
len(cities)
```
### Perform API Calls
* Perform a weather check on each city using a series of successive API calls.
* Include a print log of each city as it'sbeing processed (with the city number and city name).
```
calls =[]
for city in cities:
base_url = 'https://api.openweathermap.org/'
call = (f'{base_url}data/2.5/weather?q={city}&appid={api_key}')
calls.append(call)
print(calls[0])
df = pd.DataFrame()
for call in calls: #call = https://api.openweathermap.org/data/2.5/weather?q={city}&appid={key}
try:
data = requests.get(call).json()
name = data['name']
data_id = data['id']
data = pd.json_normalize(data)
df = df.append(data)
print(f'City: {name} | City #: {data_id}')
except:
print(f'could not download: {call}')
df.to_csv('output_data/raw_weather_data.csv', index = False, header=True)
print('Dataframe Created.')
```
### Convert Raw Data to DataFrame
* Export the city data into a .csv.
* Display the DataFrame
```
df.head()
```
## Inspect the data and remove the cities where the humidity > 100%.
----
Skip this step if there are no cities that have humidity > 100%.
```
humid = df.loc[df['main.humidity']>100]
print(len(humid))
# Get the indices of cities that have humidity over 100%.
# Make a new DataFrame equal to the city data to drop all humidity outliers by index.
# Passing "inplace=False" will make a copy of the city_data DataFrame, which we call "clean_city_data".
```
## Plotting the Data
* Use proper labeling of the plots using plot titles (including date of analysis) and axes labels.
* Save the plotted figures as .pngs.
## Latitude vs. Temperature Plot
```
lat = 'coord.lat'
temp = 'main.temp'
tem =df.plot.scatter(lat,temp)
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = tem.get_figure()
fig.savefig(f"output_data/Latitude-vs-Temperature-{today}.png")
```
## Latitude vs. Humidity Plot
```
humidity = 'main.humidity'
hum = df.plot.scatter(lat,humidity)
hum.set_xlabel("Latitude")
hum.set_ylabel("Humidity")
hum.set_title('Latitude vs Humidity')
fig = hum.get_figure()
fig.savefig(f"output_data/Latitude-vs-Humidity-{today}.png")
```
## Latitude vs. Cloudiness Plot
```
cloudiness = 'clouds.all'
cl = df.plot.scatter(lat,cloudiness)
cl.set_xlabel("Latitude")
cl.set_ylabel("Cloudiness")
cl.set_title('Latitude vs Cloudiness')
fig = cl.get_figure()
fig.savefig(f"output_data/Latitude-vs-Cloudiness-{today}.png")
```
## Latitude vs. Wind Speed Plot
```
wind_speed = 'wind.speed'
w = df.plot.scatter(lat,wind_speed)
w.set_xlabel("Latitude")
w.set_ylabel("Humidity")
w.set_title('Latitude vs Humidity')
fig = w.get_figure()
fig.savefig(f"output_data/Latitude-vs-Wind-Speed-{today}.png")
```
## Linear Regression
```
n = df.loc[df['coord.lat'] > 0]
s = df.loc[df['coord.lat'] < 0]
```
#### Northern Hemisphere - Max Temp vs. Latitude Linear Regression
```
lat = 'coord.lat'
temp = 'main.temp_max'
(slope, intercept, rvalue, pvalue, stderr) = linregress(n[lat], n[temp])
regress_values = n[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = n.plot.scatter(lat,temp)
plt.plot(n[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = w.get_figure()
fig.savefig(f"output_data/NH-Latitude-vs-Max-Temp-{today}.png")
```
#### Southern Hemisphere - Max Temp vs. Latitude Linear Regression
```
lat = 'coord.lat'
temp = 'main.temp_max'
(slope, intercept, rvalue, pvalue, stderr) = linregress(s[lat], s[temp])
regress_values = s[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = s.plot.scatter(lat,temp)
plt.plot(s[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = w.get_figure()
fig.savefig(f"output_data/SH-Latitude-vs-Max-Temp-{today}.png")
```
#### Northern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
lat = 'coord.lat'
humidity = 'main.humidity'
(slope, intercept, rvalue, pvalue, stderr) = linregress(n[lat], n[humidity])
regress_values = n[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = n.plot.scatter(lat,humidity)
plt.plot(n[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = w.get_figure()
fig.savefig(f"output_data/NH-Latitude-vs-Humidity-{today}.png")
```
#### Southern Hemisphere - Humidity (%) vs. Latitude Linear Regression
```
lat = 'coord.lat'
humidity = 'main.humidity'
(slope, intercept, rvalue, pvalue, stderr) = linregress(s[lat], s[humidity])
regress_values = s[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = s.plot.scatter(lat,humidity)
plt.plot(s[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = w.get_figure()
fig.savefig(f"output_data/SH-Latitude-vs-Humidity-{today}.png")
```
#### Northern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
lat = 'coord.lat'
cloudiness = 'clouds.all'
(slope, intercept, rvalue, pvalue, stderr) = linregress(n[lat], n[cloudiness])
regress_values = n[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = n.plot.scatter(lat,cloudiness)
plt.plot(n[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = w.get_figure()
fig.savefig(f"output_data/NH-Latitude-vs-Cloudiness-{today}.png")
```
#### Southern Hemisphere - Cloudiness (%) vs. Latitude Linear Regression
```
lat = 'coord.lat'
temp = 'main.temp_max'
(slope, intercept, rvalue, pvalue, stderr) = linregress(s[lat], s[cloudiness])
regress_values = s[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = s.plot.scatter(lat,cloudiness)
plt.plot(s[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = tem.get_figure()
fig.savefig(f"output_data/SH-Latitude-vs-Cloudiness-{today}.png")
```
#### Northern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
lat = 'coord.lat'
wind_speed = 'wind.speed'
(slope, intercept, rvalue, pvalue, stderr) = linregress(n[lat], n[wind_speed])
regress_values = n[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = n.plot.scatter(lat,wind_speed)
plt.plot(n[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = tem.get_figure()
fig.savefig(f"output_data/NH-Latitude-vs-Wind-Speed-{today}.png")
```
#### Southern Hemisphere - Wind Speed (mph) vs. Latitude Linear Regression
```
lat = 'coord.lat'
wind_speed = 'wind.speed'
(slope, intercept, rvalue, pvalue, stderr) = linregress(s[lat], s[wind_speed])
regress_values = s[lat] * slope + intercept
line_eq = "y = " + str(round(slope,2)) + "x + " + str(round(intercept,2))
tem = s.plot.scatter(lat,wind_speed)
plt.plot(s[lat],regress_values,"r-")
tem.set_xlabel("Latitude")
tem.set_ylabel("Temperature")
tem.set_title('Latitude vs Temperature')
fig = tem.get_figure()
fig.savefig(f"output_data/SH-Latitude-vs-Wind-Speed-{today}.png")
```
| github_jupyter |
## Analyze whether SNWD varies more from year to year or from place to place.
```
import pandas as pd
import numpy as np
import urllib
import math
import findspark
findspark.init()
from pyspark import SparkContext
#sc.stop()
sc = SparkContext(master="local[3]",pyFiles=['lib/numpy_pack.py','lib/spark_PCA.py','lib/computeStats.py'])
from pyspark import SparkContext
from pyspark.sql import *
sqlContext = SQLContext(sc)
import sys
sys.path.append('./lib')
import numpy as np
from numpy_pack import packArray,unpackArray
from spark_PCA import computeCov
from computeStats import computeOverAllDist, STAT_Descriptions
### Read the data frame from pickle file
data_dir='../../Data/Weather'
file_index='SBBBSBSS'
meas='SNWD'
from pickle import load
#read statistics
filename=data_dir+'/STAT_%s.pickle'%file_index
STAT,STAT_Descriptions = load(open(filename,'rb'))
print('keys from STAT=',STAT.keys())
#!ls -ld $data_dir/*.parquet
#read data
filename=data_dir+'/decon_%s_%s.parquet'%(file_index,meas)
df=sqlContext.read.parquet(filename)
print(df.count())
df.show(2)
print df.columns
#extract longitude and latitude for each station
feature='coeff_1'
sqlContext.registerDataFrameAsTable(df,'weather')
#Features=', '.join(['coeff_1', 'coeff_2', 'coeff_3', 'elevation', 'latitude', 'longitude',\
# 'res_1', 'res_2', 'res_3', 'res_mean', 'year'])
Features='station, year, coeff_1,coeff_2,coeff_3'
Query="SELECT %s FROM weather"%Features
print(Query)
pdf = sqlContext.sql(Query).toPandas()
pdf.head()
year_station_table=pdf.pivot(index='year', columns='station', values='coeff_3')
year_station_table.head(10)
```
### Estimating the effect of the year vs the effect of the station
To estimate the effect of time vs. location on the first eigenvector coefficient we
compute:
* The average row: `mean-by-station`
* The average column: `mean-by-year`
We then compute the RMS before and after subtracting either the row or the column vector.
```
def RMS(Mat):
return np.sqrt(np.nanmean(Mat**2))
mean_by_year=np.nanmean(year_station_table,axis=1)
mean_by_station=np.nanmean(year_station_table,axis=0)
tbl_minus_year = (year_station_table.transpose()-mean_by_year).transpose()
tbl_minus_station = year_station_table-mean_by_station
print 'total RMS = ',RMS(year_station_table)
print 'RMS removing mean-by-station= ',RMS(tbl_minus_station)
print 'RMS removing mean-by-year = ',RMS(tbl_minus_year)
T=year_station_table
print 'initial RMS=',RMS(T)
for i in range(5):
mean_by_year=np.nanmean(T,axis=1)
T=(T.transpose()-mean_by_year).transpose()
print i,'after removing mean by year =',RMS(T)
mean_by_station=np.nanmean(T,axis=0)
T=T-mean_by_station
print i,'after removing mean by stations=',RMS(T)
```
| github_jupyter |
# Comparing TensorFlow (original) and PyTorch models
You can use this small notebook to check the conversion of the model's weights from the TensorFlow model to the PyTorch model. In the following, we compare the weights of the last layer on a simple example (in `input.txt`) but both models returns all the hidden layers so you can check every stage of the model.
To run this notebook, follow these instructions:
- make sure that your Python environment has both TensorFlow and PyTorch installed,
- download the original TensorFlow implementation,
- download a pre-trained TensorFlow model as indicaded in the TensorFlow implementation readme,
- run the script `convert_tf_checkpoint_to_pytorch.py` as indicated in the `README` to convert the pre-trained TensorFlow model to PyTorch.
If needed change the relative paths indicated in this notebook (at the beggining of Sections 1 and 2) to point to the relevent models and code.
```
import os
os.chdir('../')
```
## 1/ TensorFlow code
```
original_tf_inplem_dir = "./tensorflow_code/"
model_dir = "../google_models/uncased_L-12_H-768_A-12/"
vocab_file = model_dir + "vocab.txt"
bert_config_file = model_dir + "bert_config.json"
init_checkpoint = model_dir + "bert_model.ckpt"
input_file = "./samples/input.txt"
max_seq_length = 128
max_predictions_per_seq = 20
masked_lm_positions = [6]
import importlib.util
import sys
import tensorflow as tf
import pytorch_pretrained_bert as ppb
def del_all_flags(FLAGS):
flags_dict = FLAGS._flags()
keys_list = [keys for keys in flags_dict]
for keys in keys_list:
FLAGS.__delattr__(keys)
del_all_flags(tf.flags.FLAGS)
import tensorflow_code.extract_features as ef
del_all_flags(tf.flags.FLAGS)
import tensorflow_code.modeling as tfm
del_all_flags(tf.flags.FLAGS)
import tensorflow_code.tokenization as tft
del_all_flags(tf.flags.FLAGS)
import tensorflow_code.run_pretraining as rp
del_all_flags(tf.flags.FLAGS)
import tensorflow_code.create_pretraining_data as cpp
import re
class InputExample(object):
"""A single instance example."""
def __init__(self, tokens, segment_ids, masked_lm_positions,
masked_lm_labels, is_random_next):
self.tokens = tokens
self.segment_ids = segment_ids
self.masked_lm_positions = masked_lm_positions
self.masked_lm_labels = masked_lm_labels
self.is_random_next = is_random_next
def __repr__(self):
return '\n'.join(k + ":" + str(v) for k, v in self.__dict__.items())
def read_examples(input_file, tokenizer, masked_lm_positions):
"""Read a list of `InputExample`s from an input file."""
examples = []
unique_id = 0
with tf.gfile.GFile(input_file, "r") as reader:
while True:
line = reader.readline()
if not line:
break
line = line.strip()
text_a = None
text_b = None
m = re.match(r"^(.*) \|\|\| (.*)$", line)
if m is None:
text_a = line
else:
text_a = m.group(1)
text_b = m.group(2)
tokens_a = tokenizer.tokenize(text_a)
tokens_b = None
if text_b:
tokens_b = tokenizer.tokenize(text_b)
tokens = tokens_a + tokens_b
masked_lm_labels = []
for m_pos in masked_lm_positions:
masked_lm_labels.append(tokens[m_pos])
tokens[m_pos] = '[MASK]'
examples.append(
InputExample(
tokens = tokens,
segment_ids = [0] * len(tokens_a) + [1] * len(tokens_b),
masked_lm_positions = masked_lm_positions,
masked_lm_labels = masked_lm_labels,
is_random_next = False))
unique_id += 1
return examples
bert_config = tfm.BertConfig.from_json_file(bert_config_file)
tokenizer = ppb.BertTokenizer(
vocab_file=vocab_file, do_lower_case=True)
examples = read_examples(input_file, tokenizer, masked_lm_positions=masked_lm_positions)
print(examples[0])
class InputFeatures(object):
"""A single set of features of data."""
def __init__(self, input_ids, input_mask, segment_ids, masked_lm_positions,
masked_lm_ids, masked_lm_weights, next_sentence_label):
self.input_ids = input_ids
self.input_mask = input_mask
self.segment_ids = segment_ids
self.masked_lm_positions = masked_lm_positions
self.masked_lm_ids = masked_lm_ids
self.masked_lm_weights = masked_lm_weights
self.next_sentence_labels = next_sentence_label
def __repr__(self):
return '\n'.join(k + ":" + str(v) for k, v in self.__dict__.items())
def pretraining_convert_examples_to_features(instances, tokenizer, max_seq_length,
max_predictions_per_seq):
"""Create TF example files from `TrainingInstance`s."""
features = []
for (inst_index, instance) in enumerate(instances):
input_ids = tokenizer.convert_tokens_to_ids(instance.tokens)
input_mask = [1] * len(input_ids)
segment_ids = list(instance.segment_ids)
assert len(input_ids) <= max_seq_length
while len(input_ids) < max_seq_length:
input_ids.append(0)
input_mask.append(0)
segment_ids.append(0)
assert len(input_ids) == max_seq_length
assert len(input_mask) == max_seq_length
assert len(segment_ids) == max_seq_length
masked_lm_positions = list(instance.masked_lm_positions)
masked_lm_ids = tokenizer.convert_tokens_to_ids(instance.masked_lm_labels)
masked_lm_weights = [1.0] * len(masked_lm_ids)
while len(masked_lm_positions) < max_predictions_per_seq:
masked_lm_positions.append(0)
masked_lm_ids.append(0)
masked_lm_weights.append(0.0)
next_sentence_label = 1 if instance.is_random_next else 0
features.append(
InputFeatures(input_ids, input_mask, segment_ids,
masked_lm_positions, masked_lm_ids,
masked_lm_weights, next_sentence_label))
if inst_index < 5:
tf.logging.info("*** Example ***")
tf.logging.info("tokens: %s" % " ".join(
[str(x) for x in instance.tokens]))
tf.logging.info("features: %s" % str(features[-1]))
return features
features = pretraining_convert_examples_to_features(
instances=examples, max_seq_length=max_seq_length,
max_predictions_per_seq=max_predictions_per_seq, tokenizer=tokenizer)
def input_fn_builder(features, seq_length, max_predictions_per_seq, tokenizer):
"""Creates an `input_fn` closure to be passed to TPUEstimator."""
all_input_ids = []
all_input_mask = []
all_segment_ids = []
all_masked_lm_positions = []
all_masked_lm_ids = []
all_masked_lm_weights = []
all_next_sentence_labels = []
for feature in features:
all_input_ids.append(feature.input_ids)
all_input_mask.append(feature.input_mask)
all_segment_ids.append(feature.segment_ids)
all_masked_lm_positions.append(feature.masked_lm_positions)
all_masked_lm_ids.append(feature.masked_lm_ids)
all_masked_lm_weights.append(feature.masked_lm_weights)
all_next_sentence_labels.append(feature.next_sentence_labels)
def input_fn(params):
"""The actual input function."""
batch_size = params["batch_size"]
num_examples = len(features)
# This is for demo purposes and does NOT scale to large data sets. We do
# not use Dataset.from_generator() because that uses tf.py_func which is
# not TPU compatible. The right way to load data is with TFRecordReader.
d = tf.data.Dataset.from_tensor_slices({
"input_ids":
tf.constant(
all_input_ids, shape=[num_examples, seq_length],
dtype=tf.int32),
"input_mask":
tf.constant(
all_input_mask,
shape=[num_examples, seq_length],
dtype=tf.int32),
"segment_ids":
tf.constant(
all_segment_ids,
shape=[num_examples, seq_length],
dtype=tf.int32),
"masked_lm_positions":
tf.constant(
all_masked_lm_positions,
shape=[num_examples, max_predictions_per_seq],
dtype=tf.int32),
"masked_lm_ids":
tf.constant(
all_masked_lm_ids,
shape=[num_examples, max_predictions_per_seq],
dtype=tf.int32),
"masked_lm_weights":
tf.constant(
all_masked_lm_weights,
shape=[num_examples, max_predictions_per_seq],
dtype=tf.float32),
"next_sentence_labels":
tf.constant(
all_next_sentence_labels,
shape=[num_examples, 1],
dtype=tf.int32),
})
d = d.batch(batch_size=batch_size, drop_remainder=False)
return d
return input_fn
def model_fn_builder(bert_config, init_checkpoint, learning_rate,
num_train_steps, num_warmup_steps, use_tpu,
use_one_hot_embeddings):
"""Returns `model_fn` closure for TPUEstimator."""
def model_fn(features, labels, mode, params): # pylint: disable=unused-argument
"""The `model_fn` for TPUEstimator."""
tf.logging.info("*** Features ***")
for name in sorted(features.keys()):
tf.logging.info(" name = %s, shape = %s" % (name, features[name].shape))
input_ids = features["input_ids"]
input_mask = features["input_mask"]
segment_ids = features["segment_ids"]
masked_lm_positions = features["masked_lm_positions"]
masked_lm_ids = features["masked_lm_ids"]
masked_lm_weights = features["masked_lm_weights"]
next_sentence_labels = features["next_sentence_labels"]
is_training = (mode == tf.estimator.ModeKeys.TRAIN)
model = tfm.BertModel(
config=bert_config,
is_training=is_training,
input_ids=input_ids,
input_mask=input_mask,
token_type_ids=segment_ids,
use_one_hot_embeddings=use_one_hot_embeddings)
(masked_lm_loss,
masked_lm_example_loss, masked_lm_log_probs) = rp.get_masked_lm_output(
bert_config, model.get_sequence_output(), model.get_embedding_table(),
masked_lm_positions, masked_lm_ids, masked_lm_weights)
(next_sentence_loss, next_sentence_example_loss,
next_sentence_log_probs) = rp.get_next_sentence_output(
bert_config, model.get_pooled_output(), next_sentence_labels)
total_loss = masked_lm_loss + next_sentence_loss
tvars = tf.trainable_variables()
initialized_variable_names = {}
scaffold_fn = None
if init_checkpoint:
(assignment_map,
initialized_variable_names) = tfm.get_assigment_map_from_checkpoint(
tvars, init_checkpoint)
if use_tpu:
def tpu_scaffold():
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
return tf.train.Scaffold()
scaffold_fn = tpu_scaffold
else:
tf.train.init_from_checkpoint(init_checkpoint, assignment_map)
tf.logging.info("**** Trainable Variables ****")
for var in tvars:
init_string = ""
if var.name in initialized_variable_names:
init_string = ", *INIT_FROM_CKPT*"
tf.logging.info(" name = %s, shape = %s%s", var.name, var.shape,
init_string)
output_spec = None
if mode == tf.estimator.ModeKeys.TRAIN:
masked_lm_positions = features["masked_lm_positions"]
masked_lm_ids = features["masked_lm_ids"]
masked_lm_weights = features["masked_lm_weights"]
next_sentence_labels = features["next_sentence_labels"]
train_op = optimization.create_optimizer(
total_loss, learning_rate, num_train_steps, num_warmup_steps, use_tpu)
output_spec = tf.contrib.tpu.TPUEstimatorSpec(
mode=mode,
loss=total_loss,
train_op=train_op,
scaffold_fn=scaffold_fn)
elif mode == tf.estimator.ModeKeys.EVAL:
masked_lm_positions = features["masked_lm_positions"]
masked_lm_ids = features["masked_lm_ids"]
masked_lm_weights = features["masked_lm_weights"]
next_sentence_labels = features["next_sentence_labels"]
def metric_fn(masked_lm_example_loss, masked_lm_log_probs, masked_lm_ids,
masked_lm_weights, next_sentence_example_loss,
next_sentence_log_probs, next_sentence_labels):
"""Computes the loss and accuracy of the model."""
masked_lm_log_probs = tf.reshape(masked_lm_log_probs,
[-1, masked_lm_log_probs.shape[-1]])
masked_lm_predictions = tf.argmax(
masked_lm_log_probs, axis=-1, output_type=tf.int32)
masked_lm_example_loss = tf.reshape(masked_lm_example_loss, [-1])
masked_lm_ids = tf.reshape(masked_lm_ids, [-1])
masked_lm_weights = tf.reshape(masked_lm_weights, [-1])
masked_lm_accuracy = tf.metrics.accuracy(
labels=masked_lm_ids,
predictions=masked_lm_predictions,
weights=masked_lm_weights)
masked_lm_mean_loss = tf.metrics.mean(
values=masked_lm_example_loss, weights=masked_lm_weights)
next_sentence_log_probs = tf.reshape(
next_sentence_log_probs, [-1, next_sentence_log_probs.shape[-1]])
next_sentence_predictions = tf.argmax(
next_sentence_log_probs, axis=-1, output_type=tf.int32)
next_sentence_labels = tf.reshape(next_sentence_labels, [-1])
next_sentence_accuracy = tf.metrics.accuracy(
labels=next_sentence_labels, predictions=next_sentence_predictions)
next_sentence_mean_loss = tf.metrics.mean(
values=next_sentence_example_loss)
return {
"masked_lm_accuracy": masked_lm_accuracy,
"masked_lm_loss": masked_lm_mean_loss,
"next_sentence_accuracy": next_sentence_accuracy,
"next_sentence_loss": next_sentence_mean_loss,
}
eval_metrics = (metric_fn, [
masked_lm_example_loss, masked_lm_log_probs, masked_lm_ids,
masked_lm_weights, next_sentence_example_loss,
next_sentence_log_probs, next_sentence_labels
])
output_spec = tf.contrib.tpu.TPUEstimatorSpec(
mode=mode,
loss=total_loss,
eval_metrics=eval_metrics,
scaffold_fn=scaffold_fn)
elif mode == tf.estimator.ModeKeys.PREDICT:
masked_lm_log_probs = tf.reshape(masked_lm_log_probs,
[-1, masked_lm_log_probs.shape[-1]])
masked_lm_predictions = tf.argmax(
masked_lm_log_probs, axis=-1, output_type=tf.int32)
next_sentence_log_probs = tf.reshape(
next_sentence_log_probs, [-1, next_sentence_log_probs.shape[-1]])
next_sentence_predictions = tf.argmax(
next_sentence_log_probs, axis=-1, output_type=tf.int32)
masked_lm_predictions = tf.reshape(masked_lm_predictions,
[1, masked_lm_positions.shape[-1]])
next_sentence_predictions = tf.reshape(next_sentence_predictions,
[1, 1])
predictions = {
"masked_lm_predictions": masked_lm_predictions,
"next_sentence_predictions": next_sentence_predictions
}
output_spec = tf.contrib.tpu.TPUEstimatorSpec(
mode=mode, predictions=predictions, scaffold_fn=scaffold_fn)
return output_spec
else:
raise ValueError("Only TRAIN, EVAL and PREDICT modes are supported: %s" % (mode))
return output_spec
return model_fn
is_per_host = tf.contrib.tpu.InputPipelineConfig.PER_HOST_V2
run_config = tf.contrib.tpu.RunConfig(
master=None,
tpu_config=tf.contrib.tpu.TPUConfig(
num_shards=1,
per_host_input_for_training=is_per_host))
model_fn = model_fn_builder(
bert_config=bert_config,
init_checkpoint=init_checkpoint,
learning_rate=0,
num_train_steps=1,
num_warmup_steps=1,
use_tpu=False,
use_one_hot_embeddings=False)
# If TPU is not available, this will fall back to normal Estimator on CPU
# or GPU.
estimator = tf.contrib.tpu.TPUEstimator(
use_tpu=False,
model_fn=model_fn,
config=run_config,
predict_batch_size=1)
input_fn = input_fn_builder(
features=features, seq_length=max_seq_length, max_predictions_per_seq=max_predictions_per_seq,
tokenizer=tokenizer)
tensorflow_all_out = []
for result in estimator.predict(input_fn, yield_single_examples=True):
tensorflow_all_out.append(result)
print(len(tensorflow_all_out))
print(len(tensorflow_all_out[0]))
print(tensorflow_all_out[0].keys())
print("masked_lm_predictions", tensorflow_all_out[0]['masked_lm_predictions'])
print("predicted token", tokenizer.convert_ids_to_tokens(tensorflow_all_out[0]['masked_lm_predictions']))
tensorflow_outputs = tokenizer.convert_ids_to_tokens(tensorflow_all_out[0]['masked_lm_predictions'])[:len(masked_lm_positions)]
print("tensorflow_output:", tensorflow_outputs)
```
## 2/ PyTorch code
```
from examples import extract_features
from examples.extract_features import *
init_checkpoint_pt = "../google_models/uncased_L-12_H-768_A-12/pytorch_model.bin"
device = torch.device("cpu")
model = ppb.BertForPreTraining.from_pretrained('bert-base-uncased')
model.to(device)
all_input_ids = torch.tensor([f.input_ids for f in features], dtype=torch.long)
all_input_mask = torch.tensor([f.input_mask for f in features], dtype=torch.long)
all_segment_ids = torch.tensor([f.segment_ids for f in features], dtype=torch.long)
all_masked_lm_positions = torch.tensor([f.masked_lm_positions for f in features], dtype=torch.long)
eval_data = TensorDataset(all_input_ids, all_input_mask, all_segment_ids, all_masked_lm_positions)
eval_sampler = SequentialSampler(eval_data)
eval_dataloader = DataLoader(eval_data, sampler=eval_sampler, batch_size=1)
model.eval()
import numpy as np
pytorch_all_out = []
for input_ids, input_mask, segment_ids, tensor_masked_lm_positions in eval_dataloader:
print(input_ids)
print(input_mask)
print(segment_ids)
input_ids = input_ids.to(device)
input_mask = input_mask.to(device)
segment_ids = segment_ids.to(device)
prediction_scores, _ = model(input_ids, token_type_ids=segment_ids, attention_mask=input_mask)
prediction_scores = prediction_scores[0, tensor_masked_lm_positions].detach().cpu().numpy()
print(prediction_scores.shape)
masked_lm_predictions = np.argmax(prediction_scores, axis=-1).squeeze().tolist()
print(masked_lm_predictions)
pytorch_all_out.append(masked_lm_predictions)
pytorch_outputs = tokenizer.convert_ids_to_tokens(pytorch_all_out[0])[:len(masked_lm_positions)]
print("pytorch_output:", pytorch_outputs)
print("tensorflow_output:", tensorflow_outputs)
```
| github_jupyter |
Simple single dense layer architecture of neutral network
import dependencies
```
import os
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Embedding,Dense
from tensorflow.keras.layers import SpatialDropout1D,Conv1D,GlobalMaxPooling1D
from tensorflow.keras.layers import Flatten,Dropout
from tensorflow.keras.callbacks import ModelCheckpoint
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import nltk,re
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score, roc_curve
```
Basic text prepocessing function
```
def preprocessing_txt(list_of_sents:pd)->list:
df_sents = list_of_sents
lemmatizer = nltk.WordNetLemmatizer()
stop_words = set(stopwords.words("english"))
new_list_sents = []
for sents in df_sents:
tokens = word_tokenize(sents)
words = " ".join([lemmatizer.lemmatize(word.lower()) for word in tokens if word.isalpha() or word not in stop_words])
new_list_sents.append(words)
return new_list_sents
```
read file and constructed feature engineered data X and proper encoded labels y
```
replace_initial = "someone"
df= pd.read_csv("data-processed.csv")
X_raw = df["DESCRIPTION"]
X_rnm = pd.array([re.sub(r"\{\{[^\{\}]+\}\}\w?", replace_initial, str1) for str1 in X_raw])
X_ = preprocessing_txt(X_rnm)
Y_ = np.array([1 if t == 'Medical emergency' else 0 for t in df["INCIDENT_TYPE_1"]])
```
Here we use sklearn split function to seperate train and test data at 80% and 20%
```
X_train,X_test,Y_train,Y_test = train_test_split(X_,Y_,train_size = 0.80, random_state = 1, shuffle = True)
X_test
```
Here we use tensorflow tokenizer. The unique function we explore here is the oov_token which represent unknown words, in our cases it should provide some better representation since we will be using embedding word vector.
```
tokenizer = Tokenizer(num_words=None,
lower =True,oov_token="<oov>")
tokenizer.fit_on_texts(X_train)
unique_words = len(tokenizer.word_index)+1
```
Here we further prepare our text tokens be ready for embedding layers by two techniques:
1. texts to sequences: It encodes word tokens into digits as you can see in word_index below
2. padded text: it is basic fill up the space to match to all same length. here I did not restrict my max words in padding to capture all word count.
```
train_sequences = tokenizer.texts_to_sequences(X_train)
padded_train = pad_sequences(train_sequences,padding = "post")
test_sequences = tokenizer.texts_to_sequences(X_test)
padded_test = pad_sequences(test_sequences,padding = "post")
max_len = len(padded_test[0])
tokenizer.word_index
```
Here we use basic model in tensorflow sequential. the neutral network architecture consist:
1. embedding layer. this layer should always be our default unless we train and import our own specilized embedding matrix.
2. max pooling here is to retain the max activation value. polling is to pick max value on each output dims of the array. at the same time since we are using binary loss so we need make sure last layer will out out 1D
3. Dense layer with 24 output diemnsions and contain most popular activiation function so far relu
4. output layer. since it is binary classfier so we use sigmoid activation function with 1 output
```
model = Sequential([Embedding(input_dim = unique_words, output_dim=128,input_length=max_len),
GlobalMaxPooling1D(),
Dense(128,activation="relu"),
Dropout(0.2),
Dense(128,activation="relu"),
Dropout(0.5),
Dense(1,activation='sigmoid')]
)
model.compile(loss="binary_crossentropy",optimizer = 'adam',metrics = ["accuracy","AUC"])
model.summary()
output = "model_output/dense"
modelcheck = ModelCheckpoint(filepath = output + "weight.{epoch:2d}.hdf5")
if not os.path.exists(output):
os.makedirs(output)
```
we set to train 30 epochs to run and find best weights for the model
```
num_epochs = 40
history = model.fit(padded_train,Y_train,
batch_size = 128,
epochs = num_epochs,
validation_data = (padded_test,Y_test),
verbose = 2,
callbacks = [modelcheck])
model.load_weights("denseweight.22.hdf5")
Y_hat = model.predict_proba(padded_test)
plt.hist(Y_hat)
_= plt.axvline(x=0.5,color = "blue")
plt.show()
pct_auc = roc_auc_score(Y_test,Y_hat) * 100.0
pct_auc
float_y_hat = []
for y in Y_hat:
float_y_hat.append(y[0])
ydf =pd.DataFrame(list(zip(float_y_hat,Y_test)),columns=['y_hat','y'])
word_index = tokenizer.word_index
word_index = {k:(v-1) for k,v in word_index.items()}
index_word ={v:k for k,v in word_index.items()}
print(ydf[(ydf.y == 1) & (ydf.y_hat < 0.5)].head(10))
words = ' '.join(index_word[id] for id in padded_test[2])
words
```
examine the sample sentences for test cases
| github_jupyter |
# Collecting temperature data from an API
## About the data
In this notebook, we will be collecting daily temperature data from the [National Centers for Environmental Information (NCEI) API](https://www.ncdc.noaa.gov/cdo-web/webservices/v2). We will use the Global Historical Climatology Network - Daily (GHCND) dataset; see the documentation [here](https://www1.ncdc.noaa.gov/pub/data/cdo/documentation/GHCND_documentation.pdf).
*Note: The NCEI is part of the National Oceanic and Atmospheric Administration (NOAA) and, as you can see from the URL for the API, this resource was created when the NCEI was called the NCDC. Should the URL for this resource change in the future, you can search for "NCEI weather API" to find the updated one.*
## Using the NCEI API
Request your token [here](https://www.ncdc.noaa.gov/cdo-web/token) and then paste it below.
```
import requests
def make_request(endpoint, payload=None):
"""
Make a request to a specific endpoint on the weather API
passing headers and optional payload.
Parameters:
- endpoint: The endpoint of the API you want to
make a GET request to.
- payload: A dictionary of data to pass along
with the request.
Returns:
A response object.
"""
return requests.get(
f'https://www.ncdc.noaa.gov/cdo-web/api/v2/{endpoint}',
headers={
'token': 'PASTE_TOKEN_HERE'
},
params=payload
)
```
**Note: the API limits us to 5 requests per second and 10,000 requests per day.**
## See which datasets are available
We can make requests to the `datasets` endpoint to see which datasets are available. We also pass in a dictionary for the payload to get datasets that have data after the start date of October 1, 2018.
```
response = make_request('datasets', {'startdate': '2018-10-01'})
```
Status code of `200` means everything is OK. More codes can be found [here](https://en.wikipedia.org/wiki/List_of_HTTP_status_codes).
```
response.status_code
```
Alternatively, we can check the `ok` attribute:
```
response.ok
```
### Get the keys of the result
The result is a JSON payload, which we can access with the `json()` method of our response object. JSON objects can be treated like dictionaries, so we can access the keys just like we would a dictionary:
```
payload = response.json()
payload.keys()
```
The metadata of the response will tell us information about the request and data we got back:
```
payload['metadata']
```
### Figure out what data is in the result
The `results` key contains the data we requested. This is a list of what would be rows in our dataframe. Each entry in the list is a dictionary, so we can look at the keys to get the fields:
```
payload['results'][0].keys()
```
### Parse the result
We don't want all those fields, so we will use a list comphrension to take only the `id` and `name` fields out:
```
[(data['id'], data['name']) for data in payload['results']]
```
## Figure out which data category we want
The `GHCND` data containing daily summaries is what we want. Now we need to make another request to figure out which data categories we want to collect. This is the `datacategories` endpoint. We have to pass the `datasetid` for `GHCND` as the payload so the API knows which dataset we are asking about:
```
# get data category id
response = make_request(
'datacategories', payload={'datasetid': 'GHCND'}
)
response.status_code
```
Since we know the API gives us a `metadata` and a `results` key in each response, we can see what is in the `results` portion of the JSON payload:
```
response.json()['results']
```
## Grab the data type ID for the temperature category
We will be working with temperatures, so we want the `TEMP` data category. Now, we need to find the `datatypes` to collect. For this, we use the `datatypes` endpoint and provide the `datacategoryid` which was `TEMP`. We also specify a limit for the number of `datatypes` to return with the payload. If there are more than this we can make another request later, but for now, we just want to pick a few out:
```
# get data type id
response = make_request(
'datatypes',
payload={
'datacategoryid': 'TEMP',
'limit': 100
}
)
response.status_code
```
We can grab the `id` and `name` fields for each of the entries in the `results` portion of the data. The fields we are interested in are at the bottom:
```
[(datatype['id'], datatype['name']) for datatype in response.json()['results']][-5:] # look at the last 5
```
## Determine which location category we want
Now that we know which `datatypes` we will be collecting, we need to find the location to use. First, we need to figure out the location category. This is obtained from the `locationcategories` endpoint by passing the `datasetid`:
```
# get location category id
response = make_request(
'locationcategories',
payload={'datasetid': 'GHCND'}
)
response.status_code
```
We can use `pprint` to print dictionaries in an easier-to-read format. After doing so, we can see there are 12 different location categories, but we are only interested in `CITY`:
```
import pprint
pprint.pprint(response.json())
```
## Get NYC Location ID
In order to find the location ID for New York, we need to search through all the cities available. Since we can ask the API to return the cities sorted, we can use binary search to find New York quickly without having to make many requests or request lots of data at once. The following function makes the first request to see how big the list is and looks at the first value. From there it decides if it needs to move towards the beginning or end of the list by comparing the item we are looking for to others alphabetically. Each time it makes a request it can rule out half of the remaining data to search.
```
def get_item(name, what, endpoint, start=1, end=None):
"""
Grab the JSON payload for a given field by name using binary search.
Parameters:
- name: The item to look for.
- what: Dictionary specifying what the item in `name` is.
- endpoint: Where to look for the item.
- start: The position to start at. We don't need to touch this, but the
function will manipulate this with recursion.
- end: The last position of the items. Used to find the midpoint, but
like `start` this is not something we need to worry about.
Returns:
Dictionary of the information for the item if found otherwise
an empty dictionary.
"""
# find the midpoint which we use to cut the data in half each time
mid = (start + (end or 1)) // 2
# lowercase the name so this is not case-sensitive
name = name.lower()
# define the payload we will send with each request
payload = {
'datasetid': 'GHCND',
'sortfield': 'name',
'offset': mid, # we will change the offset each time
'limit': 1 # we only want one value back
}
# make our request adding any additional filter parameters from `what`
response = make_request(endpoint, {**payload, **what})
if response.ok:
payload = response.json()
# if response is ok, grab the end index from the response metadata the first time through
end = end or payload['metadata']['resultset']['count']
# grab the lowercase version of the current name
current_name = payload['results'][0]['name'].lower()
# if what we are searching for is in the current name, we have found our item
if name in current_name:
return payload['results'][0] # return the found item
else:
if start >= end:
# if our start index is greater than or equal to our end, we couldn't find it
return {}
elif name < current_name:
# our name comes before the current name in the alphabet, so we search further to the left
return get_item(name, what, endpoint, start, mid - 1)
elif name > current_name:
# our name comes after the current name in the alphabet, so we search further to the right
return get_item(name, what, endpoint, mid + 1, end)
else:
# response wasn't ok, use code to determine why
print(f'Response not OK, status: {response.status_code}')
```
When we use binary search to find New York, we find it in just 8 requests despite it being close to the middle of 1,983 entries:
```
# get NYC id
nyc = get_item('New York', {'locationcategoryid': 'CITY'}, 'locations')
nyc
```
## Get the station ID for Central Park
The most granular data is found at the station level:
```
central_park = get_item('NY City Central Park', {'locationid': nyc['id']}, 'stations')
central_park
```
## Request the temperature data
Finally, we have everything we need to make our request for the New York temperature data. For this, we use the `data` endpoint and provide all the parameters we picked up throughout our exploration of the API:
```
# get NYC daily summaries data
response = make_request(
'data',
{
'datasetid': 'GHCND',
'stationid': central_park['id'],
'locationid': nyc['id'],
'startdate': '2018-10-01',
'enddate': '2018-10-31',
'datatypeid': ['TAVG', 'TMAX', 'TMIN'], # average, max, and min temperature
'units': 'metric',
'limit': 1000
}
)
response.status_code
```
## Create a DataFrame
The Central Park station only has the daily minimum and maximum temperatures.
```
import pandas as pd
df = pd.DataFrame(response.json()['results'])
df.head()
```
We didn't get `TAVG` because the station doesn't measure that:
```
df.datatype.unique()
```
Despite showing up in the data as measuring it... Real-world data is dirty!
```
if get_item(
'NY City Central Park', {'locationid': nyc['id'], 'datatypeid': 'TAVG'}, 'stations'
):
print('Found!')
```
## Using a different station
Let's use LaGuardia airport instead. It contains `TAVG` (average daily temperature):
```
laguardia = get_item(
'LaGuardia', {'locationid': nyc['id']}, 'stations'
)
laguardia
```
We make our request using the LaGuardia airport station this time.
```
# get NYC daily summaries data
response = make_request(
'data',
{
'datasetid': 'GHCND',
'stationid': laguardia['id'],
'locationid': nyc['id'],
'startdate': '2018-10-01',
'enddate': '2018-10-31',
'datatypeid': ['TAVG', 'TMAX', 'TMIN'], # temperature at time of observation, min, and max
'units': 'metric',
'limit': 1000
}
)
response.status_code
```
The request was successful, so let's make a dataframe:
```
df = pd.DataFrame(response.json()['results'])
df.head()
```
We should check that we got what we wanted: 31 entries for TAVG, TMAX, and TMIN (1 per day):
```
df.datatype.value_counts()
```
Write the data to a CSV file for use in other notebooks.
```
df.to_csv('data/nyc_temperatures.csv', index=False)
```
<hr>
<div>
<a href="./1-wide_vs_long.ipynb">
<button>← Previous Notebook</button>
</a>
<a href="./3-cleaning_data.ipynb">
<button style="float: right;">Next Notebook →</button>
</a>
</div>
<hr>
| github_jupyter |
# Soft Sorting
```
import functools
import jax
import jax.numpy as jnp
import numpy as np
import matplotlib.pyplot as plt
import scipy.ndimage
import ott
```
## Sorting operators
Given an array of $n$ numbers, several operators arise around the idea of sorting:
- The `sort` operator reshuffles the values in order, from smallest to largest.
- The `rank` operator associates to each value its rank, when sorting in ascending order.
- The `quantile` operator consider a `level` value between 0 and 1, to return the element of the sorted array indexed at `int(n * level)`, the median for instance if that level is set to $0.5$.
- The `topk` operator is equivalent to the `sort` operator, but only returns the
largest $k$ values, namely the last $k$ values of the sorted vector.
Here are some examples
```
x = jnp.array([1.0, 5.0, 4.0, 8.0, 12.0])
jnp.sort(x)
def rank(x):
return jnp.argsort(jnp.argsort(x))
rank(x)
jnp.quantile(x, 0.5)
```
## Soft operators
Sorting operators are ubiquitous in CS and stats, but have several limitations when used in modern deep learning architectures. For instance, `ranks` is integer valued: if used at some point within a DL pipeline, one won't be able to differentiate through that step because the gradient of these integer values does not exist, or is ill-defined. Indeed, the vector of `ranks` of a slightly perturbed vector $x+\Delta x$ is the same as that for $x$, or switches ranks at some indices when inversions occur. Practially speaking, any loss or intermediary operation based on `ranks` will break backpropagation.
This colab shows how _soft_ counterparts to these operators are defined in OTT. By _soft_, we mean **differentiable**, **approximate** proxies to these original _"hard"_ operators. For instance `soft_sort.ranks` returned by OTT operators won't be integer valued, but instead floating point approximations; `soft_sort.sort` will not contain exactly the `n` values contained in the input array, reordered, but instead `n` combinaisons of thoses values that look very close to them.
**These soft operators trade off accuracy for a more informative Jacobian**. This trade-off is controlled by a non-negative parameter `epsilon`: The *smaller* `epsilon`, the closer to the original ranking and sorting operations; The *bigger*, the more bias yet the more informative gradients. That `epsilon` also correponds to that used in regularized OT (see doc on [sinkhorn](https://ott-jax.readthedocs.io/en/latest/_autosummary/ott.core.sinkhorn.sinkhorn.html#ott.core.sinkhorn.sinkhorn)).
The behavior of these operators is illustrated below.
### Soft sort
```
softsort = jax.jit(ott.tools.soft_sort.sort)
print(softsort(x))
```
As we can see, the values are close to the original ones but not exactly equal. Here, `epsilon` is set by default to `1e-2`. A smaller `epsilon` reduces that gap, whereas a bigger one would tend to squash all returned values to the **average** of the input values.
```
print(softsort(x, epsilon=1e-4))
print(softsort(x, epsilon=1e-1))
```
### Soft topk
The soft operators we propose build on a common idea: formulate sorting operations as optimal transports from an array of $n$ values to a predefined target measure of $m$ points. The user is free to choose $m$, providing great flexibility depending on the use case.
Transporting an input discrete measure of $n$ points towards one of $m$ points results in a $O(nm)$ complexity. The bigger $m$, the more fine grained quantities we recover. For instance, if we wish to get both a fine grained yet differentiable sorted vector, or vector of ranks, one can define a target measure of size $m=n$, leading to a $O(n^2)$ complexity.
On the contrary, if we are only interested in singling out a few important ranks, such as when considering `top k` values, we can simply transport the inputs points onto $k+1$ targets, leading to a smaller complexity in $O(nk)$. When $k \ll n$, the gain in time and memory can be of course substantial.
Here is an example.
```
top5 = jax.jit(functools.partial(ott.tools.soft_sort.sort, topk=5))
# Generates a vector of size 1000
big_x = jax.random.uniform(jax.random.PRNGKey(0), (1000,))
top5(big_x)
```
### Soft Ranks
Similarly, we can compute soft ranks, which do not output integer values, but provide instead a differentiable, float valued, approximation of the vector of ranks.
```
softranks = jax.jit(ott.tools.soft_sort.ranks)
print(softranks(x))
```
### Regularization effect
As mentioned earlier, `epsilon` controls the tradeoff between accuracy and differentiability. Larger `epsilon` tend to merge the _soft_ ranks of values that are close, up to the point where they all collapse to the average rank or agverage value.
```
epsilons = np.logspace(-3, 1,100)
sorted_values = []
ranks = []
for e in epsilons:
sorted_values.append(softsort(x, epsilon=e))
ranks.append(softranks(x, epsilon=e))
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
for values, ax, title in zip(
(sorted_values, ranks), axes, ('sorted values', 'ranks')):
ax.plot(epsilons, np.array(values), color='k', lw=11)
ax.plot(epsilons, np.array(values), lw=7)
ax.set_xlabel('$\epsilon$', fontsize=24)
ax.tick_params(axis='both', which='both', labelsize=18)
ax.set_title(f"Soft {title}", fontsize=24)
ax.set_xscale('log')
```
Note how none of the lines above cross. This is a fundamental property of `soft-sort` operators, proved in [Cuturi et al. 2020](https://arxiv.org/abs/2002.03229): soft sorting and ranking operators are monotonic: the vector of soft-sorted values will remain increasing for any `epsilon`, whereas if an input value $x_i$ has a smaller (hard) rank than $x_j$, its soft-rank, for any value of `epsilon`, will also remain smaller than that for $x_j$.
### Soft quantile
To illustrate further the flexibility provided by setting target measures, one can notice that when a soft quantile is targeted (for instance the soft median), the complexity becomes simply $O(n)$. This is illustrated below to define "soft median" differentiable filter on a noisy image.
```
softquantile = jax.jit(ott.tools.soft_sort.quantile)
softquantile(x, level=0.5)
import io
import requests
url = "https://raw.githubusercontent.com/matplotlib/matplotlib/master/doc/_static/stinkbug.png"
resp = requests.get(url)
image = plt.imread(io.BytesIO(resp.content))
image = image[..., 0]
def salt_and_pepper(im, amount=0.05):
result = np.copy(im)
result = np.reshape(result, (-1,))
num_noises = int(np.ceil(amount * im.size))
indices = np.random.randint(0, im.size, num_noises)
values = np.random.uniform(size=(num_noises,)) > 0.5
result[indices] = values
return np.reshape(result, im.shape)
noisy_image = salt_and_pepper(image, amount=0.1)
softmedian = jax.jit(functools.partial(ott.tools.soft_sort.quantile, level=0.5))
fns = {'original': None, 'median': np.median}
for e in [0.01, 1.0]:
fns[f'soft {e}'] = functools.partial(softmedian, epsilon=e)
fns.update(mean=np.mean)
fig, axes = plt.subplots(1, len(fns), figsize=(len(fns)*6, 4))
for key, ax in zip(fns, axes):
fn = fns[key]
soft_denoised = (
scipy.ndimage.generic_filter(noisy_image, fn, size=(3, 3))
if fn is not None else noisy_image)
ax.imshow(soft_denoised)
ax.set_title(key, fontsize=22)
```
## Learning through a soft ranks operator.
A crucial feature of OTT lies in the ability it provides to **differentiate** seamlessly through any quantities that follow an optimal transport computation, making it very easy for end-users to plug them directly into end-to-end differentiable architectures.
In this tutorial we show how OTT can be used to implement a loss based on soft ranks. That soft 0-1 loss is used here to train a neural network for image classification, as done by [Cuturi et al. 2019](https://papers.nips.cc/paper/2019/hash/d8c24ca8f23c562a5600876ca2a550ce-Abstract.html).
This implementation relies on [FLAX](https://github.com/google/flax) a neural network library for JAX.
### Model
Similarly to [Cuturi et al. 2019](https://papers.nips.cc/paper/2019/hash/d8c24ca8f23c562a5600876ca2a550ce-Abstract.html), we will train a vanilla CNN made of 4 convolutional blocks, in order to classify images from the CIFAR-10 dataset.
```
from typing import Any
import flax
from flax import linen as nn
class ConvBlock(nn.Module):
"""A simple CNN blockl."""
features: int = 32
dtype: Any = jnp.float32
@nn.compact
def __call__(self, x, train: bool = True):
x = nn.Conv(features=self.features, kernel_size=(3, 3))(x)
x = nn.relu(x)
x = nn.Conv(features=self.features, kernel_size=(3, 3))(x)
x = nn.relu(x)
x = nn.max_pool(x, window_shape=(2, 2), strides=(2, 2))
return x
class CNN(nn.Module):
"""A simple CNN model."""
num_classes: int = 10
dtype: Any = jnp.float32
@nn.compact
def __call__(self, x, train: bool = True):
x = ConvBlock(features=32)(x)
x = ConvBlock(features=64)(x)
x = x.reshape((x.shape[0], -1)) # flatten
x = nn.Dense(features=512)(x)
x = nn.relu(x)
x = nn.Dense(features=self.num_classes)(x)
return x
```
### Losses & Metrics
The $0/1$ loss of a classifier on a labeled example is $0$ if the logit of the true class ranks on top (here, would have rank 9, since CIFAR-10 considers 10 classes). Of course the $0/1$ loss is non-differentiable, which is why the cross-entropy loss is used instead.
Here, as in the [paper](https://papers.nips.cc/paper/2019/hash/d8c24ca8f23c562a5600876ca2a550ce-Abstract.html), we consider a differentiable "soft" 0/1 loss by measuring the gap between the soft-rank of the logit of the right answer and the target rank, 9. If that gap is bigger than 0, then we incurr a loss equal to that gap.
```
def cross_entropy_loss(logits, labels):
logits = nn.log_softmax(logits)
return -jnp.sum(labels * logits) / labels.shape[0]
def soft_error_loss(logits, labels):
"""The average distance between the best rank and the rank of the true class."""
ranks_fn = jax.jit(functools.partial(ott.tools.soft_sort.ranks, axis=-1))
soft_ranks = ranks_fn(logits)
return jnp.mean(nn.relu(
labels.shape[-1] - 1 - jnp.sum(labels * soft_ranks, axis=1)))
```
To know more about training a neural network with Flax, please refer to the [Flax Imagenet examples](https://github.com/google/flax/tree/master/examples/imagenet/). After 100 epochs through the CIFAR-10 training examples, we are able to reproduce the results from [Cuturi et al. 2019](https://papers.nips.cc/paper/2019/hash/d8c24ca8f23c562a5600876ca2a550ce-Abstract.html) and see that a soft $0/1$ error loss, building on top of `soft_sort.ranks`, can provide a competitive alternative to the cross entropy loss for classification tasks. As mentioned in that paper, that loss is less prone to overfitting.
```
# Plot results from training
```
| github_jupyter |
<form action="index.ipynb">
<input type="submit" value="Return to Index" style="background-color: green; color: white; width: 150px; height: 35px; float: right"/>
</form>
# Landau Energy - Multidimensional X-data Example
Author(s): Paul Miles | Date Created: July 18, 2019
This example demonstrates using [pymcmcstat](https://github.com/prmiles/pymcmcstat/wiki) with multidimensional input spaces, e.g. functions that depend on two-spatial variables such as $x_1$ and $x_2$. For this particular problem we consider a 6th order Landau energy function, which is used in a wide variety of material science applications.
Consider the 6th order Landau function:
$$ u(q;{\bf P}) = \alpha_{1}(P_1^2+P_2^2+P_3^2) + \alpha_{11}(P_1^2 + P_2^2 + P_3^2)^2 + \alpha_{111}(P_1^6 + P_2^6 + P_3^6) + \alpha_{12}(P_1^2P_2^2 + P_1^2P_3^2 + P_2^2P_3^2) + \alpha_{112}(P_1^4(P_2^2 + P_3^2) + P_2^4(P_1^2 + P_3^2) + P_3^4(P_1^2 + P_2^2)) + \alpha_{123}P_1^2P_2^2P_3^2$$
where $q = [\alpha_{1},\alpha_{11},\alpha_{111},\alpha_{12},\alpha_{112},\alpha_{123}]$. The Landau energy is a function of 3-dimensional polarization space. For the purpose of this example, we consider the case where $P_1 = 0$.
Often times we are interested in using information calculated from Density Functional Theory (DFT) calculations in order to inform our continuum approximations, such as our Landau function. For this example, we will assume we have a set of energy calculations corresponding to different values of $P_2$ and $P_3$ which were found using DFT. For more details regarding this type of research, the reader is referred to:
- Miles, P. R., Leon, L. S., Smith, R. C., Oates, W. S. (2018). Analysis of a Multi-Axial Quantum Informed Ferroelectric Continuum Model: Part 1—Uncertainty Quantification. Journal of Intelligent Material Systems and Structures, 29(13), 2823-2839. https://doi.org/10.1177/1045389X18781023
- Leon, L. S., Smith, R. C., Oates, W. S., Miles, P. R. (2018). Analysis of a Multi-Axial Quantum Informed Ferroelectric Continuum Model: Part 2—Sensitivity Analysis. Journal of Intelligent Material Systems and Structures, 29(13), 2840-2860. https://doi.org/10.1177/1045389X18781024
```
# import required packages
import numpy as np
from pymcmcstat.MCMC import MCMC
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
import seaborn as sns
from pymcmcstat import mcmcplot as mcp
import scipy.io as sio
import pymcmcstat
print(pymcmcstat.__version__)
```
Define the Landau energy function.
```
def landau_energy(q, data):
P = data.xdata[0]
a1, a11, a111, a12, a112, a123 = q
u = a1*((P**2).sum(axis = 1)) + a11*((P**2).sum(axis = 1))**2 + a111*((P**6).sum(axis = 1))
u += a12*(P[:,0]**2*P[:,1]**2 + P[:,0]**2*P[:,2]**2 + P[:,1]**2*P[:,2]**2)
u += a112*(P[:,0]**4*(P[:,1]**2 + P[:,2]**2) + P[:,1]**4*(P[:,0]**2 + P[:,2]**2) + P[:,2]**4*(P[:,0]**2 + P[:,1]**2))
u += a123*P[:,0]**2*P[:,1]**2*P[:,2]**2
return u
```
We can generate a grid in the $P_2-P_3$ space in order to visualize the model response. The model is evaluated at a particular set of parameter values, and the surface plot shows the general behavior of the energy function.
```
# Make data.
P2 = np.linspace(-0.6, 0.6, 100) #np.arange(-0.6, 0.6, 0.25)
P3 = np.linspace(-0.6, 0.6, 100) #np.arange(-0.6, 0.6, 0.25)
P2, P3 = np.meshgrid(P2, P3)
nr, nc = P2.shape
P = np.concatenate((np.zeros([nr*nc,1]), P2.flatten().reshape(nr*nc,1), P3.flatten().reshape(nr*nc,1)), axis = 1)
q = [-389.4, 761.3, 61.46, 414.1, -740.8, 0.]
from pymcmcstat.MCMC import DataStructure
data = DataStructure()
data.add_data_set(
x=P,
y=None)
Z = landau_energy(q, data)
# Plot the surface.
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(P2, P3, Z.reshape([nr,nc]), cmap=cm.coolwarm,
linewidth=0, antialiased=False)
plt.xlabel('$P_2 (C/m^2)$')
plt.ylabel('$P_3 (C/m^2)$')
ax.set_zlabel('$u (MPa)$')
plt.tight_layout()
```
# DFT Data
We define a set of energy results based on a series of DFT calculations which correspond to specific polarization values.
```
pdata = sio.loadmat('data_files/dft_polarization.mat')
polarization = []
for ii, pp in enumerate(pdata['polarization']):
polarization.append(np.zeros([pp[0].shape[0], 3]))
polarization[-1][:,1:] = pp[0]
if ii == 0:
polarization_array = polarization[-1]
else:
polarization_array = np.concatenate((polarization_array, polarization[-1]), axis = 0)
edata = sio.loadmat('data_files/dft_energy.mat')
energy = []
for ii, ee in enumerate(edata['energy']):
energy.append(np.zeros([ee[0].shape[0],1]))
energy[-1][:,:] = ee[0]*1e3 # scale to MPa
if ii == 0:
energy_array = energy[-1]
else:
energy_array = np.concatenate((energy_array, energy[-1]), axis = 0)
dft = dict(energy = energy, polarization = polarization)
from pymcmcstat.MCMC import DataStructure
data = DataStructure()
data.add_data_set(
x=polarization_array,
y=energy_array)
# Plot the surface.
fig = plt.figure()
ax = fig.gca(projection='3d')
for ii, (pp, ee) in enumerate(zip(dft['polarization'], dft['energy'])):
ax.scatter(pp[:,1], pp[:,2], ee)
plt.xlabel('$P_2 (C/m^2)$')
plt.ylabel('$P_3 (C/m^2)$')
ax.set_zlabel('$u (MPa)$')
plt.tight_layout()
```
# MCMC Simulation
First we define our sum-of-squares function
```
def ssfun(q, data):
ud = data.ydata[0]
um = landau_energy(q, data)
ss = ((ud - um.reshape(ud.shape))**2).sum()
return ss
```
## Initialize MCMC Object
We then initialize the MCMC object and define the components:
- data structure
- parameters
- simulation options
- model settings
```
# Initialize MCMC object
mcstat = MCMC()
# setup data structure for dram
mcstat.data = data
# setup model parameters
mcstat.parameters.add_model_parameter(
name = '$\\alpha_{1}$',
theta0 = q[0],
minimum = -1e8,
maximum = 1e8,
)
mcstat.parameters.add_model_parameter(
name = '$\\alpha_{11}$',
theta0 = q[1],
minimum = -1e8,
maximum = 1e8,
)
mcstat.parameters.add_model_parameter(
name='$\\alpha_{111}$',
theta0=q[2],
minimum=-1e8,
maximum=1e8
)
mcstat.parameters.add_model_parameter(
name='$\\alpha_{12}$',
theta0=q[3],
minimum=-1e8,
maximum=1e8
)
mcstat.parameters.add_model_parameter(
name='$\\alpha_{112}$',
theta0=q[4],
minimum=-1e8,
maximum=1e8
)
mcstat.parameters.add_model_parameter(
name='$\\alpha_{123}$',
theta0=q[5],
sample=False, # do not sample this parameter
)
# define simulation options
mcstat.simulation_options.define_simulation_options(
nsimu=int(5e5),
updatesigma=True,
method='dram',
adaptint=100,
verbosity=1,
waitbar=1,
save_to_json=False,
save_to_bin=False,
)
# define model settings
mcstat.model_settings.define_model_settings(
sos_function=ssfun,
N0=1,
N=polarization_array.shape[0],
)
```
## Run MCMC Simulation & Display Stats
```
# Run mcmcrun
mcstat.run_simulation()
results = mcstat.simulation_results.results
# specify burnin period
burnin = int(results['nsimu']/2)
# display chain statistics
chain = results['chain']
s2chain = results['s2chain']
sschain = results['sschain']
names = results['names']
mcstat.chainstats(chain[burnin:,:], results)
print('Acceptance rate: {:6.4}%'.format(100*(1 - results['total_rejected'])))
print('Model Evaluations: {}'.format(results['nsimu'] - results['iacce'][0] + results['nsimu']))
```
## Plot MCMC Diagnostics
```
# generate mcmc plots
mcp.plot_density_panel(chain[burnin:,:], names)
mcp.plot_chain_panel(chain[burnin:,:], names)
f = mcp.plot_pairwise_correlation_panel(chain[burnin:,:], names)
```
# Generate Credible/Prediction Intervals
```
%matplotlib notebook
from pymcmcstat import propagation as up
pdata = []
intervals = []
for ii in range(len(polarization)):
pdata.append(DataStructure())
pdata[ii].add_data_set(polarization[ii], energy[ii])
intervals.append(up.calculate_intervals(chain, results, pdata[ii],
landau_energy,
s2chain=s2chain,
nsample=500, waitbar=True))
```
# Plot 3-D Credible/Prediction Intervals
```
%matplotlib notebook
for ii in range(len(polarization)):
if ii == 0:
fig3, ax3 = up.plot_3d_intervals(intervals[ii], pdata[ii].xdata[0][:, 1::], pdata[ii].ydata[0],
limits=[95],
adddata=True, addprediction=True, addcredible=True,
addlegend=True, data_display=dict(marker='o'))
else:
fig3, ax3 = up.plot_3d_intervals(intervals[ii], pdata[ii].xdata[0][:, 1::], pdata[ii].ydata[0],
limits=[95], fig=fig3,
adddata=True, addprediction=True, addcredible=True,
addlegend=False, data_display=dict(marker='o'))
ax3.set_xlabel('$P_2$')
ax3.set_ylabel('$P_3$')
ax3.set_zlabel('$u$')
tmp = ax3.set_xlim([-0.1, 0.6])
```
# Plot 2-D Credible/Prediction Intervals
```
ii = 0
fig, ax = up.plot_intervals(intervals[ii],
np.linalg.norm(pdata[ii].xdata[0][:, 1::], axis=1),
pdata[ii].ydata[0],
limits=[95],
adddata=True, addprediction=True, addcredible=True,
addlegend=True, data_display=dict(marker='o', mfc='none'),
figsize=(6, 4))
ax.set_title(str('Set {}'.format(ii + 1)))
fig.tight_layout()
```
Let's display them all statically for the sake of brevity.
```
%matplotlib inline
for ii in range(len(polarization)):
fig, ax = up.plot_intervals(intervals[ii],
np.linalg.norm(pdata[ii].xdata[0][:, 1::], axis=1),
pdata[ii].ydata[0],
limits=[95],
adddata=True, addprediction=True, addcredible=True,
addlegend=True, data_display=dict(marker='o', mfc='none'),
figsize=(6, 4))
ax.set_title(str('Set {}'.format(ii + 1)))
fig.tight_layout()
```
| github_jupyter |
# Perceptron Classifier
### Required Packages
```
!pip install imblearn
import warnings
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
from imblearn.over_sampling import RandomOverSampler
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.linear_model import Perceptron
from sklearn.metrics import classification_report,plot_confusion_matrix
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X = df[features]
Y = df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
def EncodeY(df):
if len(df.unique())<=2:
return df
else:
un_EncodedT=np.sort(pd.unique(df), axis=-1, kind='mergesort')
df=LabelEncoder().fit_transform(df)
EncodedT=[xi for xi in range(len(un_EncodedT))]
print("Encoded Target: {} to {}".format(un_EncodedT,EncodedT))
return df
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=EncodeY(NullClearner(Y))
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
#### Distribution Of Target Variable
```
plt.figure(figsize = (10,6))
se.countplot(Y)
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
#### Handling Target Imbalance
The challenge of working with imbalanced datasets is that most machine learning techniques will ignore, and in turn have poor performance on, the minority class, although typically it is performance on the minority class that is most important.
One approach to addressing imbalanced datasets is to oversample the minority class. The simplest approach involves duplicating examples in the minority class.We will perform overspampling using imblearn library.
```
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
```
### Model
the perceptron is an algorithm for supervised learning of binary classifiers.
The algorithm learns the weights for the input signals in order to draw a linear decision boundary.This enables you to distinguish between the two linearly separable classes +1 and -1.
#### Model Tuning Parameters
> **penalty** ->The penalty (aka regularization term) to be used. {‘l2’,’l1’,’elasticnet’}
> **alpha** -> Constant that multiplies the regularization term if regularization is used.
> **l1_ratio** -> The Elastic Net mixing parameter, with 0 <= l1_ratio <= 1. l1_ratio=0 corresponds to L2 penalty, l1_ratio=1 to L1. Only used if penalty='elasticnet'.
> **tol** -> The stopping criterion. If it is not None, the iterations will stop when (loss > previous_loss - tol).
> **early_stopping**-> Whether to use early stopping to terminate training when validation. score is not improving. If set to True, it will automatically set aside a stratified fraction of training data as validation and terminate training when validation score is not improving by at least tol for n_iter_no_change consecutive epochs.
> **validation_fraction** -> The proportion of training data to set aside as validation set for early stopping. Must be between 0 and 1. Only used if early_stopping is True.
> **n_iter_no_change** -> Number of iterations with no improvement to wait before early stopping.
```
# Build Model here
model = Perceptron(random_state=123)
model.fit(x_train, y_train)
```
#### Model Accuracy
score() method return the mean accuracy on the given test data and labels.
In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
#### Confusion Matrix
A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
```
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
```
#### Classification Report
A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
* **where**:
- Precision:- Accuracy of positive predictions.
- Recall:- Fraction of positives that were correctly identified.
- f1-score:- percent of positive predictions were correct
- support:- Support is the number of actual occurrences of the class in the specified dataset.
```
print(classification_report(y_test,model.predict(x_test)))
```
#### Creator: Thilakraj Devadiga , Github: [Profile](https://github.com/Thilakraj1998)
| github_jupyter |
```
import numpy as np
import tensorflow as tf
import keras
import PIL
import pandas as pd
print("Tensorflow version %s" %tf.__version__)
print("Keras version %s" %keras.__version__)
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
from tensorflow.python.client import device_lib
device_lib.list_local_devices()
from keras.models import Sequential, Model
from keras.layers import Dense, Activation, Dropout, Flatten, Convolution2D, MaxPooling2D, ZeroPadding2D
from keras.optimizers import Adam, SGD, RMSprop
from keras.models import model_from_json
from keras_vggface.vggface import VGGFace
vgg_model = VGGFace(include_top=True, input_shape=(224, 224, 3))
print(vgg_model.summary())
for layer in vgg_model.layers:
layer.trainable = False
last_layer = vgg_model.get_layer('fc7/relu').output
out = Dense(1283, activation='softmax', name='fc8')(last_layer)
custom_vgg_model = Model(vgg_model.input, out)
print(custom_vgg_model.summary())
adam = Adam(lr=1e-3, beta_1=0.9, beta_2=0.999)
custom_vgg_model.compile(optimizer=adam, loss='categorical_crossentropy', metrics=['accuracy'])
from keras.preprocessing.image import ImageDataGenerator
from os import listdir
from keras.callbacks import ModelCheckpoint, ReduceLROnPlateau, EarlyStopping
#loading training and validation sets
traindir = '/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/TrainSet'
valdir = '/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/ValidationSet'
batch_size = 64
input_shape = ()
train_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True
)
train_generator = train_datagen.flow_from_directory(
traindir,
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',
shuffle = True
)
val_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True
)
validation_generator = val_datagen.flow_from_directory(
valdir,
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',
shuffle = True
)
num_samples = train_generator.n
num_classes = train_generator.num_classes
#input_shape = train_generator.image_shape
#classnames = [k for k,v in train_generator.class_indices.items()]
#print("Image input %s" %str(input_shape))
#print("Classes: %r" %classnames)
print('Loaded %d training samples from %d classes.' %(num_samples,num_classes))
print('Loaded %d test samples from %d classes.' %(validation_generator.n,validation_generator.num_classes))
callbacks = [EarlyStopping(monitor='val_accuracy', patience=5, restore_best_weights=True, verbose=1),
ModelCheckpoint(filepath='model_blended-accessory.h5', monitor='val_accuracy', save_best_only=True, verbose=1)]
history = custom_vgg_model.fit_generator(
train_generator,
steps_per_epoch = train_generator.samples // batch_size,
callbacks = callbacks,
validation_data = validation_generator,
validation_steps = validation_generator.samples // batch_size,
epochs = 50,
verbose = 1)
custom_vgg_model.load_weights('model_blended-accessory.h5')
loss, acc = custom_vgg_model.evaluate_generator(train_generator,verbose=1)
print('Train loss: %f' %loss)
print('Train accuracy: %f' %acc)
testdir = '/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/TestSet'
test_datagen = ImageDataGenerator(
rescale=1./255,
)
test_generator = test_datagen.flow_from_directory(
testdir,
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',
shuffle = False
)
loss, acc = custom_vgg_model.evaluate_generator(test_generator,verbose=1)
print('Test loss: %f' %loss)
print('Test accuracy: %f' %acc)
# summarize history for accuracy
import matplotlib.pyplot as plt
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
import matplotlib.pyplot as plt
from PIL import Image
img_A = Image.open('/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/PoisonSamples/poison1.jpg')
img_B = Image.open('/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/BackdoorSet/images/backdoor1.jpg')
fig, ax = plt.subplots(1,2)
ax[0].set_title('Poison-Image')
ax[0].imshow(img_A);
ax[1].set_title('Backdoor-Image')
ax[1].imshow(img_B);
from keras.preprocessing import image
import os
from keras.models import load_model
batch_size = 64
backdoor_path = '/home/herson/Desktop/YouTubeFaces/YouTubeFaces/VGG_DB/'
blindset = backdoor_path + 'BackdoorSet/'
blind_datagen = ImageDataGenerator(
rescale = 1. / 255)
blind_generator = blind_datagen.flow_from_directory(
directory=blindset,
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',
shuffle = False
)
filenames = blind_generator.filenames
nb_samples = len(filenames)
prediction = custom_vgg_model.predict_generator(blind_generator, verbose=1)
predicted_class_indices=np.argmax(prediction, axis=1)
labels = (train_generator.class_indices)
labels = dict((v,k) for k,v in labels.items())
predictions = [labels[k] for k in predicted_class_indices]
new_filenames=[]
for f in filenames:
f=f.replace('images/','')
new_filenames.append(f)
results=pd.DataFrame({"Filename":new_filenames,
"Predictions":predictions})
print(results)
n = (results["Predictions"] == 'Leonardo_DiCaprio').value_counts().tolist()[0]
#print(n)
success_rate = (n/57)*100
print('The attack success rate is ' + str(success_rate) + "%")
```
| github_jupyter |
# Compare lithium-ion battery models
We compare three one-dimensional lithium-ion battery models: [the Doyle-Fuller-Newman (DFN) model](./DFN.ipynb), [the single particle model (SPM)](./SPM.ipynb), and [the single particle model with electrolyte (SPMe)](./SPMe.ipynb). Further details on these models can be found in [[4]](#References).
## Key steps:
Comparing models consists of 6 easy steps:
1. Load models and geometry
2. Process parameters
3. Mesh the geometry
4. Discretise models
5. Solve models
6. Plot results
But, as always we first import pybamm and other required modules
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import os
os.chdir(pybamm.__path__[0]+'/..')
import numpy as np
import matplotlib.pyplot as plt
```
## 1. Load models
Since the three models we want to compare are already implemented in PyBaMM, they can be easy loaded using:
```
dfn = pybamm.lithium_ion.DFN()
spme = pybamm.lithium_ion.SPMe()
spm = pybamm.lithium_ion.SPM()
```
To allow us to perform the same operations on each model easily, we create a dictionary of these three models:
```
models = {"DFN": dfn, "SPM": spm, "SPMe": spme}
```
Each model can then be accessed using:
```
models["DFN"]
```
For each model, we must also provide a cell geometry. The geometry is different for different models; for example, the SPM has solves for a single particle in each electrode whereas the DFN solves for many particles. For simplicity, we use the default geometry associated with each model but note that this can be easily changed.
```
geometry = {"DFN": dfn.default_geometry, "SPM": spm.default_geometry, "SPMe": spme.default_geometry}
```
## 2. Process parameters
For simplicity, we use the default parameters values associated with the DFN model, but change the current function to be an input so that we can quickly solve the model with different currents
```
param = dfn.default_parameter_values
param["Current function [A]"] = "[input]"
```
It is simple to change this to a different parameter set if desired.
We then process the parameters in each of the models and geometries using this parameter set:
```
for model_name in models.keys():
param.process_model(models[model_name])
param.process_geometry(geometry[model_name])
```
## 3. Mesh geometry
We use the defaults mesh properties (the types of meshes and number of points to be used) for simplicity to generate a mesh of each model geometry. We store these meshes in a dictionary of similar structure to the geometry and models dictionaries:
```
mesh = {}
for model_name, model in models.items():
mesh[model_name] = pybamm.Mesh(geometry[model_name], model.default_submesh_types, model.default_var_pts)
```
## 4. Discretise model
We now discretise each model using its associated mesh and the default spatial method associated with the model:
```
for model_name, model in models.items():
disc = pybamm.Discretisation(mesh[model_name], model.default_spatial_methods)
disc.process_model(model)
```
## 5. Solve model
We now solve each model using the default solver associated with each model:
```
timer = pybamm.Timer()
solutions = {}
t_eval = np.linspace(0, 3600, 300) # time in seconds
solver = pybamm.CasadiSolver()
for model_name, model in models.items():
timer.reset()
solution = solver.solve(model, t_eval, inputs={"Current function [A]": 1})
print("Solved the {} in {}".format(model.name, timer.time()))
solutions[model_name] = solution
```
## 6. Plot results
To plot results, we extract the variables from the solutions dictionary. Matplotlib can then be used to plot the voltage predictions of each models as follows:
```
for model_name, model in models.items():
time = solutions[model_name]["Time [s]"].entries
voltage = solutions[model_name]["Terminal voltage [V]"].entries
plt.plot(time, voltage, lw=2, label=model.name)
plt.xlabel("Time [s]", fontsize=15)
plt.ylabel("Terminal voltage [V]", fontsize=15)
plt.legend(fontsize=15)
plt.show()
```
Alternatively the inbuilt `QuickPlot` functionality can be employed to compare a set of variables over the discharge. We must first create a list of the solutions
```
list_of_solutions = list(solutions.values())
```
And then employ `QuickPlot`:
```
quick_plot = pybamm.QuickPlot(list_of_solutions)
quick_plot.dynamic_plot();
```
# Changing parameters
Since we have made current an input, it is easy to change it and then perform the calculations again:
```
# update parameter values and solve again
# simulate for shorter time
t_eval = np.linspace(0,800,300)
for model_name, model in models.items():
solutions[model_name] = model.default_solver.solve(model, t_eval, inputs={"Current function [A]": 3})
# Plot
list_of_solutions = list(solutions.values())
quick_plot = pybamm.QuickPlot(list_of_solutions)
quick_plot.dynamic_plot();
```
By increasing the current we observe less agreement between the models, as expected.
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import sys, os
import math
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from Shapes.RaceTrack import RaceTrack
from Shapes.utils import *
from Grid.GridProcessing import Grid
from dynamics.DubinsCar4D2 import DubinsCar4D2
from plot_options import PlotOptions
from solver import HJSolver
name = 'Thruxton'
track = RaceTrack(name)
plt.plot(track.centerline_arr[:, 0],track.centerline_arr[:, 1], 'k--')
plt.plot(track.inside_arr[:, 0], track.inside_arr[:, 1], 'b')
plt.plot(track.outside_arr[:, 0], track.outside_arr[:, 1], 'r')
```
## Usage for Puppy
```
track.raceline_length
idx = 1000
interval = 36
start_pt = track.centerline_arr[idx]
end_pt = track.centerline_arr[idx + interval]
yaw = track.race_yaw[idx]
yaw_min = min(track.race_yaw[idx:idx+interval]) - math.pi/6
yaw_max = max(track.race_yaw[idx:idx+interval]) + math.pi/6
v_min = 0
v_max = 30
pts = []
u_init = idx/track.raceline_length
for pt in [start_pt, end_pt]:
for tck in [track.tck_in, track.tck_out]:
_, p_closest, _ = track._calc_shortest_distance(pt, tck, u_init = u_init)
pts.append(p_closest)
pts = np.array(pts)
pts_local = toLocal(pts, start_pt, yaw)
x_min, x_max, y_min, y_max = fitRectangle(pts_local)
g = Grid(np.array([x_min, y_min-1, v_min, yaw_min]),
np.array([x_max, y_max+1, v_max, yaw_max]), 4, np.array([60, 15, 20, 36]), [3])
# The value function should be calculated in the global coordinate.
basis = np.array([[np.cos(yaw), -np.sin(yaw), start_pt[0]],
[np.sin(yaw), np.cos(yaw), start_pt[1]]])
V0, xx, yy = track.get_init_value(g, u_init = u_init, basis=basis)
fig, ax = plt.subplots()
CS = ax.contour(xx, yy, V0[:, :, 0, 0])
ax.clabel(CS, inline=True, fontsize=10,)
ax.plot(track.centerline_arr[idx:idx+interval, 0],track.centerline_arr[idx:idx+interval, 1], 'k--')
plt.plot(track.inside_arr[idx:idx+interval, 0], track.inside_arr[idx:idx+interval, 1], 'r--')
#plt.plot(track.outside_arr[idx:idx+interval, 0], track.outside_arr[idx:idx+interval, 1], 'r--')
from Grid.GridProcessing import Grid
g = Grid(np.array([0.0, -220.0, 0.0, -math.pi]),
np.array([50.0, -150.0, 4.0, math.pi]),
4, np.array([60, 60, 20, 36]), [3])
name = 'Thruxton'
track = RaceTrack(name)
plt.plot(track.centerline_arr[:, 0],track.centerline_arr[:, 1], 'k--')
plt.plot(track.inside_arr[:, 0], track.inside_arr[:, 1], 'b')
plt.plot(track.outside_arr[:, 0], track.outside_arr[:, 1], 'r')
V0 = track.get_init_value(g, )
plt.xlim(0, 50)
plt.ylim(-220, -150)
point
```
len(track.outside_arr)
| github_jupyter |
<a href="https://colab.research.google.com/github/Theophine/Machine_Learning/blob/master/xgboost_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# import all libraries and dependencies for dataframe
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
import warnings
warnings.filterwarnings("ignore")
# import all libraries and dependencies for data visualization
plt.rcParams['figure.figsize'] = [8,8]
pd.set_option('display.max_columns', 500)
plt.style.use('ggplot')
sns.set(style='darkgrid')
import xgboost as xgb #xgboost package
from sklearn.model_selection import train_test_split #for splitting the data into training and testing dataset
from sklearn.metrics import balanced_accuracy_score, roc_auc_score, make_scorer, confusion_matrix, plot_confusion_matrix #for model validation
from sklearn.model_selection import GridSearchCV #for cross validation
from google.colab import files
uploaded = files.upload()
import pandas as pd
import io
df_churn = pd.read_csv(io.BytesIO(uploaded['Telco-Customer-Churn.csv']))
```
#Access dataframe
```
df_churn.head()
df_churn.shape
```
some data represents the response from people who had left the company. Therefore, we are going to drop these columns
```
for i, v in enumerate(df_churn):
print(i, v)
```
any column with only one value or 85% of only one value must be removed
```
print(df_churn['SeniorCitizen'].unique())
print(df_churn['SeniorCitizen'].value_counts(dropna = False))
```
#data wrangling
Before you begin with data wrangling, make sure that you call the df.dtype attribute, that way you can tell a valid from an invalid value.
```
df_copy = df_churn.copy()
df_copy .dtypes
```
First observation, total charges is supposed to be a float but instead, it has the datatype of object. It is an indication that something is wrong with it.
##access categorical variables to see how many values each have
Because having too many categorical variables would be computationally expensive
```
#print out the unique values of total charges
df_copy ['TotalCharges'].nunique()
#there are a total of 6531 unique values which makes finding the text value difficults
#we have to convert the total charges column to numeric
pd.to_numeric(df_copy ['TotalCharges'])
```
from the above error, it shows that there are blank space in certian cells in the 'Total charges' column. we need to fix this...
```
#how many cells have missing values in the Total charges columns
len(df_copy.loc[df_copy['TotalCharges'] == ' '])
#I will replace those blank values with np.nan and then interpolate to replace the nan values
df_copy.loc[df_copy['TotalCharges'] == ' ', 'TotalCharges'] = np.nan
#Test code
df_copy['TotalCharges'].value_counts(dropna = False)
#convert the totalcharges columns to numeric and then interpolate the nan values
df_copy['TotalCharges'] = pd.to_numeric(df_copy['TotalCharges'])
#Time to interpolate
df_copy['TotalCharges'].interpolate(inplace = True)
#test code
df_copy['TotalCharges'].value_counts(dropna = False)
```
From the above, there are no more nan values
##investigate other columns
```
df_copy['MultipleLines'].value_counts(dropna = False)
df_copy['InternetService'].value_counts(dropna = False)
df_copy['Contract'].value_counts(dropna = False)
df_copy['PaymentMethod'].value_counts(dropna = False)
```
##check for missing values
```
df_copy.isna().sum()
df_copy.shape
```
##check for duplicates
```
df_copy .duplicated(subset= ['customerID']).sum()
```
From the above, there are no duplicate values and thus, the column customerID has to be dropped
```
df_copy .drop(['customerID'], axis = 1, inplace= True)
#test the code
df_copy .shape
df_copy.replace(' ', '_', regex= True, inplace = True)
```
#split your dataset into predictors and target
```
X = df_copy.drop('Churn', axis = 1).copy()
y = df_copy['Churn'].copy()
X.dtypes
#change the data type of gender and seniorcitizen to category
X['gender'] = X['gender'].astype('category')
X['SeniorCitizen'] = X['SeniorCitizen'].astype('category')
```
#OneHot encoding of the object and category column
```
X_encoded = pd.get_dummies(X, drop_first= True)
X_encoded.head(2)
X_encoded.shape
#Inspect the target column
y.value_counts(dropna = False)
```
the data is imbalanced and thus must be balanced
time to split the data into train test split
```
X_train, X_test, y_train, y_test = train_test_split(X_encoded, y, test_size = 0.2, random_state = 42, stratify = y)
```
#Model building stage
##instantiate the xgboost model
```
#instantiate xgboost
xgb_clf = xgb.XGBClassifier(objective = 'binary:logistic', random_state= 42, missing = None)
#Quickly evaluate the performance of the model
xgb_model = xgb_clf.fit(X_train, y_train, eval_set = [(X_test, y_test)], eval_metric= 'auc', early_stopping_rounds= 10, verbose = True)
#print the fitted model to see the parameters it used
xgb_model
```
##Plot confusion matrix
```
#evaluate the model using confusio matrix
plot_confusion_matrix(xgb_model, X_test, y_test, values_format= 'd', display_labels= ['Stayed', 'Left'])
plt.show();
```
##Observation: Remember that we did not tune the hyperparameters and we did not balance the dataset which is why the model performed poorly.
##Tune individual hyperparameters using yellow brick
```
#Testing for 'reg_alpha' (i.e., L1 regularization)
from yellowbrick.model_selection import ValidationCurve
#find the best value for reg_alpha lambda
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz = ValidationCurve(xgb_model, param_name= 'reg_alpha', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, n_jobs = -1, scoring = 'f1_weighted')
viz.fit(X_train, y_train)
viz.poof();
#find the best value for reg_lambda lambda (i.e., L2)
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_lambda = ValidationCurve(xgb_model, param_name='reg_lambda', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_lambda.fit(X_train, y_train)
viz_lambda.poof();
#we check for the learning rate
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_lambda = ValidationCurve(xgb_model, param_name='learning_rate', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_lambda.fit(X_train, y_train)
viz_lambda.poof();
```
inspect each scores and their ranges to see the best combination
```
viz_lambda.param_range[viz_lambda.test_scores_mean_.argmax()]
```
next we check for the pruning parameter (gamma)
```
#we check for the prunning argument i.e., gamma
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_gamma = ValidationCurve(xgb_model, param_name='gamma', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_gamma.fit(X_train, y_train)
viz_gamma.poof();
#we check for the max_depth argument
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_depth = ValidationCurve(xgb_model, param_name='max_depth', param_range= np.arange(0, 30, 1), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_depth.fit(X_train, y_train)
viz_depth.poof();
viz_depth.param_range[viz_depth.test_scores_mean_.argmax()]
viz_depth.test_scores_mean_
#we check for the number of estimators
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_estimators = ValidationCurve(xgb_model, param_name='n_estimators', param_range= np.arange(0, 300, 50), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_estimators.fit(X_train, y_train)
viz_estimators.poof();
viz_estimators.param_range[viz_estimators.test_scores_mean_.argmax()]
#we check for the scale_pos_weight argument i.e., for cases of data imbalance
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_pos_weight = ValidationCurve(xgb_model, param_name='scale_pos_weight', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_pos_weight.fit(X_train, y_train)
viz_pos_weight.poof();
#we check for the min_child_weight argument
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_min_child_weight = ValidationCurve(xgb_model, param_name='min_child_weight', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_min_child_weight.fit(X_train, y_train)
viz_min_child_weight.poof();
viz_min_child_weight.param_range[viz_min_child_weight.test_scores_mean_.argmax()]
viz_min_child_weight.test_scores_mean_
#we check for the subsample argument i.e., percentage of total observation to use for each training/tree building
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_subsample = ValidationCurve(xgb_model, param_name='subsample', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_subsample.fit(X_train, y_train)
viz_subsample.poof();
#we check for the colsample argument i.e., percentage of total features to use for each training/tree building
from sklearn.model_selection import StratifiedKFold
n_splits = StratifiedKFold(n_splits= 6)
fig, ax = plt.subplots(figsize = (10, 6), dpi = 100)
viz_colsample = ValidationCurve(xgb_model, param_name='colsample_bytree', param_range= np.linspace(0, 1, 10), ax = ax, cv = n_splits, scoring= 'f1_weighted', n_jobs= -1)
viz_colsample.fit(X_train, y_train)
viz_colsample.poof();
viz_colsample.param_range[viz_colsample.test_scores_mean_.argmax()]
```
##time to tune our hyperparameters and build a new model
```
from sklearn.feature_selection import f_classif, SelectKBest
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import make_scorer
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline, Pipeline
from sklearn.metrics import f1_score
steps = [('kbest', SelectKBest(f_classif)), ('scaler', StandardScaler()), ('imputer', SimpleImputer()) ,('xgboost', xgb.XGBClassifier(objective='binary:logistic',random_state= 42, n_jobs= -1, missing = None))]
pipeline = Pipeline(steps)
params = [{'kbest__k': np.arange(1, len(X_train.columns), 1), 'xgboost__reg_alpha': [0.7], 'xgboost__max_depth': [2, 3], 'xgboost__learning_rate':[0.11],
'xgboost__n_estimators': [150], 'xgboost__gamma': [0.88, 0.9], 'xgboost__scale_pos_weight':[1],
'xgboost__min_child_weight': [0.7, 0.8], 'xgboost__subsample': [0.3], 'xgboost__colsample_bytree': [0.5]}]
```
#build the grid search
```
gridsearch = GridSearchCV(pipeline, param_grid= params, scoring = make_scorer(f1_score, average = 'weighted'), cv = n_splits, return_train_score= True, n_jobs = -1)
#fit it to the training dataset
grid_model = gridsearch.fit(X_train, y_train)
grid_model.best_score_
grid_model.best_params_
#check for overfitting
grid_cv = grid_model.cv_results_
np.mean(grid_cv['mean_train_score']), np.mean(grid_cv['mean_test_score'])
```
There is no overfit in the dataset
##below we have the predictions and their probabilities
```
y_pred = grid_model.predict(X_test)
y_pred_proba = grid_model.predict_proba(X_test)
```
below I am threshold holding the probabilities to 0.6
#Model evaluation
```
[s[1] > 0.6 for s in y_pred_proba] == y_pred
from sklearn.metrics import roc_curve
from sklearn.preprocessing import LabelEncoder
fpr, tpr, threshold = roc_curve(LabelEncoder().fit_transform(y_test), LabelEncoder().fit_transform([s[1] > 0.6 for s in y_pred_proba]))
plt.plot([0, 1], [0, 1])
plt.plot(fpr, tpr)
plt.title('roc curve')
plt.xlabel('fpr')
plt.ylabel('tpr')
plt.show();
from sklearn.metrics import roc_auc_score
roc_auc_score(LabelEncoder().fit_transform(y_test), LabelEncoder().fit_transform([s[1] > 0.6 for s in y_pred_proba]))
confusion_mat = pd.DataFrame(confusion_matrix(LabelEncoder().fit_transform(y_test), LabelEncoder().fit_transform([s[1] > 0.6 for s in y_pred_proba])))
confusion_mat
sns.heatmap(confusion_mat, annot= True, cmap = 'coolwarm', fmt = '.3g')
plt.title('confusion matrix');
plt.show();
```
##Below we plot the classifcation report
```
from yellowbrick.classifier import ClassificationReport
fig, ax = plt.subplots(figsize = (10,6), dpi =100)
class_rep = ClassificationReport(grid_model, classes= ['Churn', 'Not churn'], ax = ax, support = True)
class_rep.fit(X_train, y_train)
class_rep.score(X_test, y_test)
class_rep.poof();
```
#Evaluating the quality of each xgboosted tree
```
print(grid_model.best_estimator_[3].get_booster().get_dump()[70])
```
#plotting an xgboost tree
```
cnode_params = {'shape':'box',
'style':'filled,rounded',
'fillcolor':'#78bceb'
}
lnode_params = {'shape':'box',
'style':'filled',
'fillcolor':'#e48038'
}
xgb.to_graphviz(grid_model.best_estimator_[3], num_trees= 19, condition_node_params= cnode_params, leaf_node_params= lnode_params, size = "10.10")
```
| github_jupyter |
# Stellargraph Ensembles for node attribute inference
This notebook demonstrates the use of `stellargraph`'s `Ensemble` class for node attribute inference using the Cora and Pubmed-Diabetes citation datasets.
The `Ensemble` class brings ensemble learning to `stellargraph`'s graph neural network models, e.g., `GraphSAGE` and `GCN`, quantifying prediction variance and potentially improving prediction accuracy.
**References**
1. Inductive Representation Learning on Large Graphs. W.L. Hamilton, R. Ying, and J. Leskovec arXiv:1706.02216
[cs.SI], 2017.
2. Semi-Supervised Classification with Graph Convolutional Networks. T. Kipf, M. Welling. ICLR 2017. arXiv:1609.02907
3. Graph Attention Networks. P. Velickovic et al. ICLR 2018
```
import networkx as nx
import pandas as pd
import numpy as np
import itertools
import os
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
import stellargraph as sg
from stellargraph.mapper import GraphSAGENodeGenerator, FullBatchNodeGenerator
from stellargraph.layer import GraphSAGE, GCN, GAT
from stellargraph import globalvar
from stellargraph import Ensemble, BaggingEnsemble
from keras import layers, optimizers, losses, metrics, Model, models
from sklearn import preprocessing, feature_extraction, model_selection
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
def plot_history(history):
def remove_prefix(text, prefix):
return text[text.startswith(prefix) and len(prefix):]
figsize=(6, 4)
c_train = 'b'
c_test = 'g'
metrics = sorted(set([remove_prefix(m, "val_") for m in list(history[0].history.keys())]))
for m in metrics:
# summarize history for metric m
plt.figure(figsize=figsize)
for h in history:
plt.plot(h.history[m], c=c_train)
plt.plot(h.history['val_' + m], c=c_test)
plt.title(m)
plt.ylabel(m)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='best')
```
### Loading the network data
**Downloading the CORA dataset:**
The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/lbc/cora.tgz
The following is the description of the dataset:
> The Cora dataset consists of 2708 scientific publications classified into one of seven classes.
> The citation network consists of 5429 links. Each publication in the dataset is described by a
> 0/1-valued word vector indicating the absence/presence of the corresponding word from the dictionary.
> The dictionary consists of 1433 unique words. The README file in the dataset provides more details.
Download and unzip the cora.tgz file to a location on your computer and set the `data_dir` variable to
point to the location of the dataset (the directory containing "cora.cites" and "cora.content").
**Downloading the PubMed-Diabetes dataset:**
The dataset used in this demo can be downloaded from https://linqs-data.soe.ucsc.edu/public/Pubmed-Diabetes.tgz
The following is the description of the dataset:
>The Pubmed Diabetes dataset consists of 19717 scientific publications from PubMed database pertaining to diabetes classified into one of three classes. The citation network consists of 44338 links. Each publication in the dataset is described by a TF/IDF weighted word vector from a dictionary which consists of 500 unique words.
Download and unzip the Pubmed-Diabetes.tgz file to a location on your computer.
Set the data_dir variable to point to the location of the processed dataset.
First, we select the dataset to use, either Cora or Pubmed-Diabetes
```
use_cora = True # Select the dataset; if False, then Pubmed-Diabetes dataset is used.
if use_cora:
data_dir = os.path.expanduser("~/data/cora")
else:
data_dir = os.path.expanduser("~/data/pubmed/Pubmed-Diabetes/data")
def load_cora(data_dir, largest_cc=False):
g_nx = nx.read_edgelist(path=os.path.expanduser(os.path.join(data_dir, "cora.cites")))
for edge in g_nx.edges(data=True):
edge[2]['label'] = 'cites'
# load the node attribute data
cora_data_location = os.path.expanduser(os.path.join(data_dir, "cora.content"))
node_attr = pd.read_csv(cora_data_location, sep='\t', header=None)
values = { str(row.tolist()[0]): row.tolist()[-1] for _, row in node_attr.iterrows()}
nx.set_node_attributes(g_nx, values, 'subject')
if largest_cc:
# Select the largest connected component. For clarity we ignore isolated
# nodes and subgraphs; having these in the data does not prevent the
# algorithm from running and producing valid results.
g_nx_ccs = (g_nx.subgraph(c).copy() for c in nx.connected_components(g_nx))
g_nx = max(g_nx_ccs, key=len)
print("Largest subgraph statistics: {} nodes, {} edges".format(
g_nx.number_of_nodes(), g_nx.number_of_edges()))
feature_names = ["w_{}".format(ii) for ii in range(1433)]
column_names = feature_names + ["subject"]
node_data = pd.read_csv(os.path.join(data_dir, "cora.content"),
sep="\t", header=None,
names=column_names)
node_data.index = node_data.index.map(str)
node_data = node_data[node_data.index.isin(list(g_nx.nodes()))]
for nid in node_data.index:
g_nx.node[nid][globalvar.TYPE_ATTR_NAME] = "paper" # specify node type
return g_nx, node_data, feature_names
def load_pubmed(data_dir):
edgelist = pd.read_csv(os.path.join(data_dir, 'Pubmed-Diabetes.DIRECTED.cites.tab'),
sep="\t", skiprows=2,
header=None )
edgelist.drop(columns=[0,2], inplace=True)
edgelist.columns = ['source', 'target']
# delete unneccessary prefix
edgelist['source'] = edgelist['source'].map(lambda x: x.lstrip('paper:'))
edgelist['target'] = edgelist['target'].map(lambda x: x.lstrip('paper:'))
edgelist["label"] = "cites" # set the edge type
# Load the graph from the edgelist
g_nx = nx.from_pandas_edgelist(edgelist, edge_attr="label")
# Load the features and subject for each node in the graph
nodes_as_dict = []
with open(os.path.join(os.path.expanduser(data_dir),
"Pubmed-Diabetes.NODE.paper.tab")) as fp:
for line in itertools.islice(fp, 2, None):
line_res = line.split("\t")
pid = line_res[0]
feat_name = ['pid'] + [l.split("=")[0] for l in line_res[1:]][:-1] # delete summary
feat_value = [l.split("=")[1] for l in line_res[1:]][:-1] # delete summary
feat_value = [pid] + [ float(x) for x in feat_value ] # change to numeric from str
row = dict(zip(feat_name, feat_value))
nodes_as_dict.append(row)
# Create a Pandas dataframe holding the node data
node_data = pd.DataFrame(nodes_as_dict)
node_data.fillna(0, inplace=True)
node_data['label'] = node_data['label'].astype(int)
node_data['label'] = node_data['label'].astype(str)
node_data.index = node_data['pid']
node_data.drop(columns=['pid'], inplace=True)
node_data.head()
for nid in node_data.index:
g_nx.node[nid][globalvar.TYPE_ATTR_NAME] = "paper" # specify node type
feature_names = list(node_data.columns)
feature_names.remove("label")
return g_nx, node_data, feature_names
```
Load the graph data.
```
if use_cora:
Gnx, node_data, feature_names = load_cora(data_dir)
else:
Gnx, node_data, feature_names = load_pubmed(data_dir)
```
We aim to train a graph-ML model that will predict the "subject" or "label" attribute on the nodes depending on the selected dataset. These subjects are one of 7 or 3 categories for Cora and PubMed-Diabetes respectively:
```
# Print the class names for the selected dataset
if use_cora:
print(set(node_data["subject"]))
else:
print(set(node_data["label"]))
```
### Splitting the data
For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to do this
```
node_label = "label"
if use_cora:
node_label = "subject"
train_data, test_data = model_selection.train_test_split(node_data,
train_size=0.2, #140
test_size=None,
stratify=node_data[node_label],
random_state=42)
val_data, test_data = model_selection.train_test_split(test_data,
train_size=0.2, #500,
test_size=None,
stratify=test_data[node_label],
random_state=100)
```
### Converting to numeric arrays
For our categorical target, we will use one-hot vectors that will be fed into a soft-max Keras layer during training.
```
target_encoding = feature_extraction.DictVectorizer(sparse=False)
train_targets = target_encoding.fit_transform(train_data[[node_label]].to_dict('records'))
val_targets = target_encoding.transform(val_data[[node_label]].to_dict('records'))
test_targets = target_encoding.transform(test_data[[node_label]].to_dict('records'))
```
We now do the same for the node attributes we want to use to predict the subject. These are the feature vectors that the Keras model will use as input.
```
node_features = node_data[feature_names]
```
### Specify global parameters
Here we specify some parameters that control the type of model we are going to use. For example, we specify the base model type, e.g., GCN, GraphSAGE, etc, and the number of estimators in the ensemble as well as model-specific parameters.
```
model_type = 'graphsage' # Can be either gcn, gat, or graphsage
use_bagging=True # If True, each model in the ensemble is trained on a bootstrapped sample
# of the given training data; otherwise, the same training data are used
# for training each model.
if model_type == "graphsage":
# For GraphSAGE model
batch_size = 50;
num_samples = [10, 10]
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 50 # The number of training epochs
elif model_type == "gcn":
# For GCN model
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 50 # The number of training epochs
elif model_type == "gat":
# For GAT model
layer_sizes = [8, train_targets.shape[1]]
attention_heads = 8
n_estimators = 5 # The number of estimators in the ensemble
n_predictions = 10 # The number of predictions per estimator per query point
epochs = 200 # The number of training epochs
```
## Creating the base graph machine learning model in Keras
Now create a `StellarGraph` object from the `NetworkX` graph and the node features and targets. It is `StellarGraph` objects that we use in this library to perform machine learning tasks on.
```
G = sg.StellarGraph(Gnx, node_features=node_features)
print(G.info())
```
To feed data from the graph to the Keras model we need a generator that feeds data from the graph into the model. The generators are specialized to the model and the learning task.
For training we use only the training nodes returned from our splitter and the target values. The `shuffle=True` argument is given to the `flow` method to improve training for those generators that support shuffling.
```
if model_type == 'graphsage':
generator = GraphSAGENodeGenerator(G, batch_size, num_samples)
train_gen = generator.flow(train_data.index, train_targets, shuffle=True)
elif model_type == 'gcn':
generator = FullBatchNodeGenerator(G, method="gcn")
train_gen = generator.flow(train_data.index, train_targets) # does not support shuffle
elif model_type == 'gat':
generator = FullBatchNodeGenerator(G, method="gat")
train_gen = generator.flow(train_data.index, train_targets) # does not support shuffle
len(train_data.index)
```
Now we can specify our machine learning model, we need a few more parameters for this but the parameters are model-specific.
```
if model_type == 'graphsage':
base_model = GraphSAGE(
layer_sizes=[16, 16],
generator=train_gen,
bias=True,
dropout=0.5,
normalize="l2"
)
x_inp, x_out = base_model.node_model(flatten_output=True)
prediction = layers.Dense(units=train_targets.shape[1], activation="softmax")(x_out)
elif model_type == 'gcn':
base_model = GCN(
layer_sizes=[32, train_targets.shape[1]],
generator = generator,
bias=True,
dropout=0.5,
activations=["elu", "softmax"]
)
x_inp, x_out = base_model.node_model()
prediction = x_out
elif model_type == 'gat':
base_model = GAT(
layer_sizes=layer_sizes,
attn_heads=attention_heads,
generator=generator,
bias=True,
in_dropout=0.5,
attn_dropout=0.5,
activations=["elu", "softmax"],
)
x_inp, prediction = base_model.node_model()
```
Let's have a look at the shape of the output tensor.
```
prediction.shape
```
### Create a Keras model and then an Ensemble
Now let's create the actual Keras model with the graph inputs `x_inp` provided by the `base_model` and outputs being the predictions from the softmax layer.
```
model = Model(inputs=x_inp, outputs=prediction)
```
Next, we create the ensemble model consisting of `n_estimators` models.
We are also going to specify that we want to make `n_predictions` per query point per model. These predictions will differ because of the application of `dropout` and, in the case of ensembling GraphSAGE models, the sampling of node neighbourhoods.
```
if use_bagging:
model = BaggingEnsemble(model, n_estimators=n_estimators, n_predictions=n_predictions)
else:
model = Ensemble(model, n_estimators=n_estimators, n_predictions=n_predictions)
model.compile(
optimizer=optimizers.Adam(lr=0.005),
loss=losses.categorical_crossentropy,
metrics=["acc"],
)
# The model is of type stellargraph.utils.ensemble.Ensemble but has
# a very similar interface to a Keras model
model
```
The ensemble has `n_estimators` models. Let's have a look at the first model's layers.
```
model.layers(0)
```
Train the model, keeping track of its loss and accuracy on the training set, and its performance on the validation set during the training (e.g., for early stopping), and generalization performance of the final model on a held-out test set (we need to create another generator over the test data for this)
```
val_gen = generator.flow(val_data.index, val_targets)
test_gen = generator.flow(test_data.index, test_targets)
```
Note that the amount of time to train the ensemble is linear to `n_estimators`.
Also, we are going to use early stopping by monitoring the accuracy on the validation data and stopping if the accuracy does not increase after 10 training epochs (this is the default grace value specified by the `Ensemble` class but we can set it to a different value by using `model.early_stopping_patience=20` for example.)
```
if use_bagging:
# When using bootstrap samples to train each model in the ensemble, we must specify
# the IDs of the training nodes (train_data) and their corresponding target values
# (train_targets)
history = model.fit_generator(
generator,
train_data = train_data.index,
train_targets = train_targets,
epochs=epochs,
validation_data=val_gen,
verbose=0,
shuffle=False,
bag_size=None,
use_early_stopping=True, # Enable early stopping
early_stopping_monitor="val_acc",
)
else:
history = model.fit_generator(
train_gen,
epochs=epochs,
validation_data=val_gen,
verbose=0,
shuffle=False,
use_early_stopping=True, # Enable early stopping
early_stopping_monitor="val_acc",
)
plot_history(history)
```
Now we have trained the model, let's evaluate it on the test set. Note that the `.evaluate_generator()` method of the `Ensemble` class returns mean and standard deviation of each evaluation metric.
```
test_metrics_mean, test_metrics_std = model.evaluate_generator(test_gen)
print("\nTest Set Metrics of the trained models:")
for name, m, s in zip(model.metrics_names, test_metrics_mean, test_metrics_std):
print("\t{}: {:0.4f}±{:0.4f}".format(name, m, s))
```
### Making predictions with the model
Now let's get the predictions for all nodes, using a new generator for all nodes:
```
all_nodes = node_data.index
all_gen = generator.flow(all_nodes)
all_predictions = model.predict_generator(generator=all_gen)
all_predictions.shape
```
For full-batch methods, the batch dimension is 1 so we will remove any singleton dimensions
```
all_predictions = np.squeeze(all_predictions)
all_predictions.shape
```
These predictions will be the output of the softmax layer, so to get final categories we'll use the `inverse_transform` method of our target attribute specifcation to turn these values back to the original categories
For demonstration, we are going to select one of the nodes in the graph, and plot the ensemble's predictions for that node.
```
selected_query_point = -1
```
The array `all_predictions` has dimensionality $MxKxNxF$ where $M$ is the number of estimators in the ensemble (`n_estimators`); $K$ is the number of predictions per query point per estimator (`n_predictions`); $N$ is the number of query points (`len(all_predictions)`); and $F$ is the output dimensionality of the specified layer determined by the shape of the output layer.
Since we are only interested in the predictions for a single query node, e.g., `selected_query_point`, we are going to slice the array to extract them.
```
# Select the predictions for the point specified by selected_query_point
qp_predictions = all_predictions[:, :, selected_query_point, :]
# The shape should be n_estimators x n_predictions x size_output_layer
qp_predictions.shape
```
Next, to facilitate plotting the predictions using either a density plot or a box plot, we are going to reshape `qp_predictions` to $R\times F$ where $R$ is equal to $M\times K$ as above and $F$ is the output dimensionality of the output layer.
```
qp_predictions = qp_predictions.reshape(np.product(qp_predictions.shape[0:-1]), qp_predictions.shape[-1])
qp_predictions.shape
inv_subject_mapper = {k: v for k, v in enumerate(target_encoding.feature_names_)}
inv_subject_mapper
```
We'd like to assess the ensemble's confidence in its predictions in order to decide if we can trust them or not. Utilising density plots, we can visually inspect the ensemble's distribution of prediction probabilities for a node's label.
This is better demonstrated if the ensemble's base mode is `GraphSAGE` because the predictions of the base model vary most (when compared to GCN and GAT) due to the random sampling of node neighbours during prediction in addition to the inherent stocasticity of the ensemble itself.
If the density plot for the predicted node label is well separated from those of the other labels with little overlap then we can be confident trusting the model's prediction.
```
if model_type not in ['gcn', 'gat']:
fig, ax = plt.subplots(figsize=(12,6))
for i in range(qp_predictions.shape[1]):
sns.kdeplot(data=qp_predictions[:, i].reshape((-1,)), label=inv_subject_mapper[i])
plt.xlabel("Predicted Probability")
plt.title("Density plots of predicted probabilities for each subject")
```
An alternative and possibly more informative view of the distribution of node predictions is a box plot.
```
fig, ax = plt.subplots(figsize=(12,6))
ax.boxplot(x=qp_predictions)
ax.set_xticklabels(target_encoding.feature_names_)
ax.tick_params(axis='x', rotation=45)
if model_type == "graphsage":
y = np.argmax(target_encoding.transform(node_data[[node_label]].to_dict('records')), axis=1)
elif model_type == "gcn" or model_type == "gat":
y = np.argmax(target_encoding.transform(node_data.reindex(G.nodes())[[node_label]].to_dict('records')), axis=1)
plt.title("Correct "+target_encoding.feature_names_[y[selected_query_point]])
plt.ylabel("Predicted Probability")
plt.xlabel("Subject")
```
The above example shows that the ensemble predicts the correct node label with high confidence so we can trust its prediction.
(Note that due to the stochastic nature of training neural network algorithms, the above conclusion may not be valid if you re-run the notebook; however, the general conclusion that the use of ensemble learning can be used to quantify the model's uncertainty about its predictions still holds.)
## Node embeddings
Evaluate node embeddings as activations of the output of one of the graph convolutional or aggregation layers in the ensemble model, and visualise them, coloring nodes by their subject label.
You can find the index of the layer of interest by calling the `Ensemble` class's method `layers`, e.g., `model.layers()`.
```
if model_type == 'graphsage':
# For GraphSAGE, we are going to use the output activations of the second GraphSAGE layer
# as the node embeddings
emb = model.predict_generator(generator=generator,
predict_data=node_data.index,
output_layer=-4) # this selects the output layer
elif model_type == 'gcn' or model_type == 'gat':
# For GCN and GAT, we are going to use the output activations of the first GCN or Graph
# Attention layer as the node embeddings
emb = model.predict_generator(generator=generator,
predict_data=node_data.index,
output_layer=6) # this selects the output layer
```
The array `emb` has dimensionality $MxKxNxF$ (or $MxKx1xNxF$for full batch methods) where $M$ is the number of estimators in the ensemble (`n_estimators`); $K$ is the number of predictions per query point per estimator (`n_predictions`); $N$ is the number of query points (`len(node_data.index)`); and $F$ is the output dimensionality of the specified layer determined by the shape of the readout layer as specified above.
```
emb.shape
emb = np.squeeze(emb)
emb.shape
```
Next we are going to average the predictions over the number of models and the number of predictions per query point.
The dimensionality of the array will then be **NxF** where N is the number of points to predict (equal to the number of nodes in the graph for this example) and F is the dimensionality of the embeddings that depends on the output shape of the readout layer as specified above.
Note that we could have achieved the same by specifying `summarise=True` in the call to the method `predict_generator` above.
```
emb = np.mean(emb, axis=(0,1))
emb.shape
```
Project the embeddings to 2d using either TSNE or PCA transform, and visualise, coloring nodes by their subject label
```
X = emb
if model_type == 'graphsage':
y = np.argmax(target_encoding.transform(node_data[[node_label]].to_dict('records')),
axis=1)
elif model_type == 'gcn' or model_type =='gat':
y = np.argmax(target_encoding.transform(node_data.reindex(G.nodes())[[node_label]].to_dict('records')),
axis=1)
if X.shape[1] > 2:
transform = TSNE # PCA
trans = transform(n_components=2)
emb_transformed = pd.DataFrame(trans.fit_transform(X), index=node_data.index)
emb_transformed['label'] = y
else:
emb_transformed = pd.DataFrame(X, index=node_data.index)
emb_transformed = emb_transformed.rename(columns = {'0':0, '1':1})
emb_transformed['label'] = y
alpha = 0.7
fig, ax = plt.subplots(figsize=(7,7))
ax.scatter(emb_transformed[0], emb_transformed[1], c=emb_transformed['label'].astype("category"),
cmap="jet", alpha=alpha)
ax.set(aspect="equal", xlabel="$X_1$", ylabel="$X_2$")
plt.title('{} visualization of {} embeddings for cora dataset'.format(model_type, transform.__name__))
plt.show()
```
| github_jupyter |
SOP060 - Uninstall kubernetes module
====================================
Steps
-----
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, hyperlinked suggestions, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
first_run = True
rules = None
debug_logging = False
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# apply expert rules (to run follow-on notebooks), based on output
#
if rules is not None:
apply_expert_rules(line_decoded)
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
"""Load a json file from disk and return the contents"""
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
"""Load any 'expert rules' from the metadata of this notebook (.ipynb) that should be applied to the stderr of the running executable"""
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
try:
j = load_json("sop060-uninstall-kubernetes-module.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"expanded_rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["expanded_rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
"""Determine if the stderr line passed in, matches the regular expressions for any of the 'expert rules', if so
inject a 'HINT' to the follow-on SOP/TSG to run"""
global rules
for rule in rules:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
if debug_logging:
print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
if debug_logging:
print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'python': []}
error_hints = {'python': [['Library not loaded: /usr/local/opt/unixodbc', 'SOP012 - Install unixodbc for Mac', '../install/sop012-brew-install-odbc-for-sql-server.ipynb'], ['WARNING: You are using pip version', 'SOP040 - Upgrade pip in ADS Python sandbox', '../install/sop040-upgrade-pip.ipynb']]}
install_hint = {'python': []}
```
### Pip uninstall the kubernetes module
```
import sys
run(f'python -m pip uninstall kubernetes -y')
```
### Pip list installed modules
```
run(f'python -m pip list')
print('Notebook execution complete.')
```
| github_jupyter |
```
class email:
@classmethod
def from_text(cls,name):
if "@" not in name:
raise ValueError("no @")
else:
name, k ,domain = name.partition("@")
#print(name,k,domain)
return cls(name,domain)
def __init__(self,name,domain):
self.name = name
self.domain = domain
def __repr__(self):
return "email is {}@{}".format(self.name,self.domain)
def __str__(self):
return self.__repr__() + " str"
```
Every domain model as an python object
```
a = email.from_text('hello@12@3.com')
```
# Metaclass, \_\_new___, \_\_init__
1. uses metaclass animalbase as to collect class instantiated
2. extention: attach metaclass to animal then inherit animal from dog cats pigeons
3. new - for object creation (can create immut
4. init - to attach properties to obj passed after created from new
```
class AnimalBase(type):
animal_list = {}
cls_list= {}
animal_bse = None
def __new__(cls, name, bases, dct):
AnimalBase.animal_list[name]=dct
AnimalBase.animal_bse=cls
x = super().__new__(cls, name, bases, dct)
AnimalBase.cls_list[name]=x
return x
#def __repr__(self):
# return 'animals'
class cats(metaclass=AnimalBase):
def __new__(cls):
return super(cats, cls).__new__(cls)
def __init__(self,name):
print('meow')
print(name)
return self
pass
class dogs(metaclass=AnimalBase):
def __new__(cls):
return super(dogs, cls).__new__(cls)
def __init__(self,name):
print('woof')
print(name)
return self
pass
class pigeons(metaclass=AnimalBase):
def __new__(cls):
return super(pigeons, cls).__new__(cls)
# if i over write repr of my meta class then type a will change
def __init__(self,name,p):
print('peek')
self.name = "pigeon "+ name
return self
class Animal(metaclass=AnimalBase):
def __new__(cls):
return super(Animal, cls).__new__(cls)
# if i over write repr of my meta class then type a will change
def __init__(self,name,Atype,*a,**kw):
self.Atype = Atype
return self
def animal_factory(name,type,*args, **kwargs):
# POWER OF *ARGS AND *KW
# PIGEONS TAKE 3 ARG SO I DELAY AND PUSH TO OBJ CREATION AND DECIDE PARAM OF PIGEONS
animal_obj=AnimalBase.cls_list[type]
final = animal_obj.__new__(animal_obj).__init__(name,*args,**kwargs)
return final
a_1 = animal_factory("gow",'dogs')
b_1 = animal_factory('new','pigeons','d')
## convert all types to animal
def animal_factory2(name,Atype,*args, **kwargs):
# POWER OF *ARGS AND *KW
# PIGEONS TAKE 3 ARG SO I DELAY AND PUSH TO OBJ CREATION AND DECIDE PARAM OF PIGEONS
animal_obj=AnimalBase.animal_bse
animal_dict=AnimalBase.animal_list
init= animal_dict[Atype]["__init__"]
final = init(animal_obj,name,*args,**kwargs)
return final
a_2 = animal_factory2("gow",'dogs')
b_2 = animal_factory2('new','pigeons','d')
def animal_factory3(name,Atype,*args, **kwargs):
animal_obj=AnimalBase.cls_list["Animal"]
final = animal_obj.__new__(animal_obj).__init__(name, Atype, *args, **kwargs)
return final
a_3 = animal_factory3("gow",'dogs')
b_3 = animal_factory3('new','pigeons','d','d')
a_1,b_1,a_2,b_2
type(a_1),type(b_1),type(a_2),type(a_3)
isinstance(dogs,Animal) , isinstance(a,dogs) , isinstance(a,Animal)
```
| github_jupyter |
```
import numpy as np
import torch
import torch.utils.data
import torch.nn as nn
import torch.optim as optim
import torchdiffeq
from tensorboard_utils import Tensorboard
from tensorboard_utils import tensorboard_event_accumulator
import transformer.Constants as Constants
from transformer.Layers import EncoderLayer, DecoderLayer
from transformer.Modules import ScaledDotProductAttention
from transformer.Models import Decoder, get_attn_key_pad_mask, get_non_pad_mask, get_sinusoid_encoding_table
from transformer.SubLayers import PositionwiseFeedForward
import dataset
import model_process
import checkpoints
from node_transformer import NodeTransformer
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
#%matplotlib notebook
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
print("Torch Version", torch.__version__)
%load_ext autoreload
%autoreload 2
seed = 1
torch.manual_seed(seed)
device = torch.device("cuda")
print("device", device)
data = torch.load("/home/mandubian/datasets/multi30k/multi30k.atok.low.pt")
max_token_seq_len = data['settings'].max_token_seq_len
print(max_token_seq_len)
train_loader, val_loader = dataset.prepare_dataloaders(data, batch_size=128)
```
### Create an experiment with a name and a unique ID
```
exp_name = "transformer_6_layers_multi30k"
unique_id = "2019-06-07_1000"
```
### Create Model
```
model = None
src_vocab_sz = train_loader.dataset.src_vocab_size
print("src_vocab_sz", src_vocab_sz)
tgt_vocab_sz = train_loader.dataset.tgt_vocab_size
print("tgt_vocab_sz", tgt_vocab_sz)
if model:
del model
model = NodeTransformer(
n_src_vocab=max(src_vocab_sz, tgt_vocab_sz),
n_tgt_vocab=max(src_vocab_sz, tgt_vocab_sz),
len_max_seq=max_token_seq_len,
n_layers=6,
#emb_src_tgt_weight_sharing=False,
#d_word_vec=128, d_model=128, d_inner=512,
n_head=8, method='dopri5-ext', rtol=1e-3, atol=1e-3,
has_node_encoder=False, has_node_decoder=False)
model = model.to(device)
```
### Create Tensorboard metrics logger
```
tb = Tensorboard(exp_name, unique_name=unique_id)
```
### Create basic optimizer
```
optimizer = optim.Adam(model.parameters(), lr=1e-4, betas=(0.9, 0.995), eps=1e-9)
```
### Train
```
# Continuous space discretization
timesteps = np.linspace(0., 1, num=6)
timesteps = torch.from_numpy(timesteps).float()
EPOCHS = 50
LOG_INTERVAL = 5
#from torch import autograd
#with autograd.detect_anomaly():
model_process.train(
exp_name, unique_id,
model,
train_loader, val_loader, timesteps,
optimizer, device,
epochs=EPOCHS, tb=tb, log_interval=LOG_INTERVAL,
#start_epoch=0, best_valid_accu=state["acc"]
)
```
### Restore best checkpoint (to restart past training)
```
state = checkpoints.restore_best_checkpoint(
"node_transformer_multi30k", "2019-05-29_1200", "validation", model, optimizer)
print("accuracy", state["acc"])
print("loss", state["loss"])
model = model.to(device)
```
| github_jupyter |
```
#Import required library
import numpy as np
#Manually add the dataset
countries = np.array(['Algeria','Angola','Argentina','Australia','Austria','Bahamas','Bangladesh','Belarus',
'Belgium','Bhutan','Brazil','Bulgaria','Cambodia','Cameroon','Chile','China','Colombia',
'Cyprus','Denmark','El Salvador','Estonia','Ethiopia','Fiji','Finland','France','Georgia',
'Ghana','Grenada','Guinea','Haiti','Honduras','Hungary','India','Indonesia','Ireland',
'Italy','Japan','Kenya', 'South Korea','Liberia','Malaysia','Mexico', 'Morocco','Nepal',
'New Zealand','Norway','Pakistan', 'Peru','Qatar','Russia','Singapore','South Africa',
'Spain','Sweden','Switzerland','Thailand', 'United Arab Emirates','United Kingdom',
'United States','Uruguay','Venezuela','Vietnam','Zimbabwe'])
gdp_per_capita = np.array([2255.225482,629.9553062,11601.63022,25306.82494,27266.40335,19466.99052,588.3691778,
2890.345675,24733.62696,1445.760002,4803.398244,2618.876037,590.4521124,665.7982328,
7122.938458,2639.54156,3362.4656,15378.16704,30860.12808,2579.115607,6525.541272,
229.6769525,2242.689259,27570.4852,23016.84778,1334.646773,402.6953275,6047.200797,
394.1156638,385.5793827,1414.072488,5745.981529,837.7464011,1206.991065,27715.52837,
18937.24998,39578.07441,478.2194906,16684.21278,279.2204061,5345.213415,6288.25324,
1908.304416,274.8728621,14646.42094,40034.85063,672.1547506,3359.517402,36152.66676,
3054.727742,33529.83052,3825.093781,15428.32098,33630.24604,39170.41371,2699.123242,
21058.43643,28272.40661,37691.02733,9581.05659,5671.912202,757.4009286,347.7456605])
#Find and print the name of the country with the highest GDP
max_gdp_per_capita = gdp_per_capita.argmax()
country_name_for_max_gdp = countries[max_gdp_per_capita]
print(country_name_for_max_gdp)
#Find and print the name of the country with the lowest GDP
min_gdp_per_capita = gdp_per_capita.argmin()
country_name_for_min_gdp = countries[min_gdp_per_capita]
print(country_name_for_min_gdp)
#Print out text ('evaluating country') and input value ('country name') iteratively
for country in countries:
print("Evaluating country:- {} ".format(country))
#Print out the entire list of the countries with their GDPs
for i in range(len(countries)):
print("Name of Country: {} and GDP: {}".format(countries[i], gdp_per_capita[i]))
''''Highest GPD value
Lowest GDP value
Mean GDP value
Standardized GDP value
Sum of all the GDPs'''
print("Highest GDP value:- {}".format(max(gdp_per_capita)))
print("Lowest GDP value:- {}".format(min(gdp_per_capita)))
print("Mean GDP value:- {}".format(np.mean(gdp_per_capita)))
print("Standardizes GDP value:- {}".format(np.std(gdp_per_capita)))
print("Sum of all the GDPs value:- {}".format(sum(gdp_per_capita)))
```
| github_jupyter |
# Introduction to C++
## Hello world
```
%%file hello.cpp
#include <iostream>
int main() {
std::cout << "Hello, world!" << std::endl;
}
```
### Compilation
```
%%bash
g++ hello.cpp -o hello.exe
```
### Execution
```
%%bash
./hello.exe
```
## Type conversions
```
%%file type.cpp
#include <iostream>
using std::cout;
using std::string;
using std::stoi;
int main() {
char c = '3'; // A char is an integer type
string s = "3"; // A string is not an integer type
int i = 3;
float f = 3;
double d = 3;
cout << c << "\n";
cout << i << "\n";
cout << f << "\n";
cout << d << "\n";
cout << "c + i is " << c + i << "\n";
cout << "c + i is " << c - '0' + i << "\n";
cout << "s + i is " << stoi(s) + i << "\n"; // Use std::stod to convert to double
}
%%bash
g++ -o type.exe type.cpp -std=c++14
%%bash
./type.exe
```
## Command line inputs
```
%%file command_args.cpp
#include <iostream>
#include <exception>
using std::cout;
using std::stoi;
int main(int argc, char *argv[]) {
for (int i=0; i<argc; i++) {
cout << i << ": " << argv[i];
try {
stoi(argv[i]);
cout << " is an integer\n";
} catch (std::exception& e) {
cout << " is not an integer\n";
}
}
}
%%bash
g++ -o command_args.exe command_args.cpp -std=c++14
%%bash
./command_args.exe 1 2 hello goodbye
```
**Exercise 1**.
Write a C++ program ex01.cpp that takes a single number `n` as input on the command line and then prints the square of that number.
```
%%file ex01.cpp
#include <iostream>
#include <cmath>
using std::cout;
using std::stod;
int main(int argc, char *argv[]) {
if (argc ==2 ) {
double x = stod(argv[1]);
cout << x*x << "\n";
} else {
cout << "Usage: ex01.exe <n>\n";
}
}
%%bash
g++ -o ex01.exe ex01.cpp -std=c++14
%%bash
./ex01.exe 5
```
## Functions
```
%%file func01.cpp
#include <iostream>
double add(double x, double y) {
return x + y;
}
double mult(double x, double y) {
return x * y;
}
int main() {
double a = 3;
double b = 4;
std::cout << add(a, b) << std::endl;
std::cout << mult(a, b) << std::endl;
}
```
### Compilation
```
%%bash
g++ -o func01.exe func01.cpp -std=c++14
```
### Execution
```
%%bash
./func01.exe
```
## Header, implementation and driver files
### Header file(s)
```
%%file func02.hpp
double add(double x, double y);
double mult(double x, double y);
```
### Implementation file(s)
```
%%file func02.cpp
double add(double x, double y) {
return x + y;
}
double mult(double x, double y) {
return x * y;
}
```
### Driver program
```
%%file test_func02.cpp
#include <iostream>
#include "func02.hpp"
int main() {
double a = 3;
double b = 4;
std::cout << add(a, b) << std::endl;
std::cout << mult(a, b) << std::endl;
}
```
### Compilation
```
%%bash
g++ test_func02.cpp func02.cpp -o main02.exe
```
### Execution
```
%%bash
./test_func02.exe
```
## Using `make`
```
%%file Makefile
test_func02.exe: test_func02.o func02.o
g++ -o test_func02.exe test_func02.o func02.o
test_func02.o: test_func02.cpp func02.hpp
g++ -c test_func02.cpp
func02.o: func02.cpp
g++ -c func02.cpp
```
### Compilation
```
%%bash
make
```
### Execution
```
%%bash
./test_func02.exe
```
## A more flexible Makefile
```
%%file Makefile2
CC=g++
CFLAGS=-Wall -std=c++14
test_func02.exe: test_func02.o func02.o
$(CC) $(CFLAGS) -o test_func02.exe test_func02.o func02.o
test_func02.o: test_func02.cpp func02.hpp
$(CC) $(CFLAGS) -c test_func02.cpp
func02.o: func02.cpp
$(CC) $(CFLAGS) -c func02.cpp
```
### Compilation
Note that no re-compilation occurs!
```
%%bash
make -f Makefile2
```
### Execution
```
%%bash
./test_func02.exe
```
## Input and output
```
%%file data.txt
9 6
%%file io.cpp
#include <fstream>
#include "func02.hpp"
int main() {
std::ifstream fin("data.txt");
std::ofstream fout("result.txt");
double a, b;
fin >> a >> b;
fin.close();
fout << add(a, b) << std::endl;
fout << mult(a, b) << std::endl;
fout.close();
}
%%bash
g++ io.cpp -o io.exe func02.cpp
%%bash
./io.exe
! cat result.txt
```
## Arrays
```
%%file array.cpp
#include <iostream>
using std::cout;
using std::endl;
int main() {
int N = 3;
double counts[N];
counts[0] = 1;
counts[1] = 3;
counts[2] = 3;
double avg = (counts[0] + counts[1] + counts[2])/3;
cout << avg << endl;
}
%%bash
g++ -o array.exe array.cpp
%%bash
./array.exe
```
## Loops
```
%%file loop.cpp
#include <iostream>
using std::cout;
using std::endl;
int main()
{
int x[] = {1, 2, 3, 4, 5};
cout << "\nTraditional for loop\n";
for (int i=0; i < sizeof(x)/sizeof(x[0]); i++) {
cout << i << endl;
}
cout << "\nRanged for loop\n\n";
for (auto &i : x) {
cout << i << endl;
}
}
%%bash
g++ -o loop.exe loop.cpp -std=c++14
%%bash
./loop.exe
```
## Function arguments
```
%%file func_arg.cpp
#include <iostream>
using std::cout;
using std::endl;
// Value parameter
void f1(int x) {
x *= 2;
cout << "In f1 : x=" << x << endl;
}
// Reference parameter
void f2(int &x) {
x *= 2;
cout << "In f2 : x=" << x << endl;
}
/* Note
If you want to avoid side effects
but still use references to avoid a copy operation
use a const refernece like this to indicate that x cannot be changed
void f2(const int &x)
*/
/* Note
Raw pointers are prone to error and
generally avoided in modern C++
See unique_ptr and shared_ptr
*/
// Raw pointer parameter
void f3(int *x) {
*x *= 2;
cout << "In f3 : x=" << *x << endl;
}
int main() {
int x = 1;
cout << "Before f1: x=" << x << "\n";
f1(x);
cout << "After f1 : x=" << x << "\n";
cout << "Before f2: x=" << x << "\n";
f2(x);
cout << "After f2 : x=" << x << "\n";
cout << "Before f3: x=" << x << "\n";
f3(&x);
cout << "After f3 : x=" << x << "\n";
}
%%bash
c++ -o func_arg.exe func_arg.cpp --std=c++14
%%bash
./func_arg.exe
```
**Exercise 2**.
Write a C++ program ex02.cpp that uses a function to calculate the 10th Fibonacci number. Here is the Python version to calculate the nth Fibonacci number for comparison.
```
def fib(n):
a, b = 0, 1
for i in range(n):
a, b = b, a + b
return a
fib(10)
%%file ex02.cpp
#include <iostream>
using std::cout;
// In practice, would probably use double rather than int
// as C++ ints will overflow when n is large.
int fib(int n) {
int a = 0;
int b = 1;
int tmp;
for (int i=0; i<n; i++) {
tmp = a;
a = b;
b = tmp + b;
}
return a;
}
int main() {
cout << fib(10) << "\n";
}
%%bash
g++ -o ex02.exe ex02.cpp -std=c++14
%%bash
./ex02.exe
```
## Anonymous functions
```
%%file lambda.cpp
#include <iostream>
using std::cout;
using std::endl;
int main() {
int a = 3, b = 4;
int c = 0;
// Lambda function with no capture
auto add1 = [] (int a, int b) { return a + b; };
// Lambda function with value capture
auto add2 = [c] (int a, int b) { return c * (a + b); };
// Lambda funciton with reference capture
auto add3 = [&c] (int a, int b) { return c * (a + b); };
// Change value of c after function definition
c += 5;
cout << "Lambda function\n";
cout << add1(a, b) << endl;
cout << "Lambda function with value capture\n";
cout << add2(a, b) << endl;
cout << "Lambda function with reference capture\n";
cout << add3(a, b) << endl;
}
%%bash
c++ -o lambda.exe lambda.cpp --std=c++14
%%bash
./lambda.exe
```
## Function pointers
```
%%file func_pointer.cpp
#include <iostream>
#include <vector>
#include <functional>
using std::cout;
using std::endl;
using std::function;
using std::vector;
int main()
{
cout << "\nUsing generalized function pointers\n";
using func = function<double(double, double)>;
auto f1 = [](double x, double y) { return x + y; };
auto f2 = [](double x, double y) { return x * y; };
auto f3 = [](double x, double y) { return x + y*y; };
double x = 3, y = 4;
vector<func> funcs = {f1, f2, f3,};
for (auto& f : funcs) {
cout << f(x, y) << "\n";
}
}
%%bash
g++ -o func_pointer.exe func_pointer.cpp -std=c++14
%%bash
./func_pointer.exe
```
## Generic programming with templates
```
%%file template.cpp
#include <iostream>
template<typename T>
T add(T a, T b) {
return a + b;
}
int main() {
int m =2, n =3;
double u = 2.5, v = 4.5;
std::cout << add(m, n) << std::endl;
std::cout << add(u, v) << std::endl;
}
%%bash
g++ -o template.exe template.cpp
%%bash
./template.exe
```
## Standard template library (STL)
```
%%file stl.cpp
#include <iostream>
#include <vector>
#include <map>
#include <unordered_map>
using std::vector;
using std::map;
using std::unordered_map;
using std::string;
using std::cout;
using std::endl;
struct Point{
int x;
int y;
Point(int x_, int y_) :
x(x_), y(y_) {};
};
int main() {
vector<int> v1 = {1,2,3};
v1.push_back(4);
v1.push_back(5);
cout << "Vecotr<int>" << endl;
for (auto n: v1) {
cout << n << endl;
}
cout << endl;
vector<Point> v2;
v2.push_back(Point(1, 2));
v2.emplace_back(3,4);
cout << "Vector<Point>" << endl;
for (auto p: v2) {
cout << "(" << p.x << ", " << p.y << ")" << endl;
}
cout << endl;
map<string, int> v3 = {{"foo", 1}, {"bar", 2}};
v3["hello"] = 3;
v3.insert({"goodbye", 4});
// Note the a C++ map is ordered
// Note using (traditional) iterators instead of ranged for loop
cout << "Map<string, int>" << endl;
for (auto iter=v3.begin(); iter != v3.end(); iter++) {
cout << iter->first << ": " << iter->second << endl;
}
cout << endl;
unordered_map<string, int> v4 = {{"foo", 1}, {"bar", 2}};
v4["hello"] = 3;
v4.insert({"goodbye", 4});
// Note theunordered_map is similar to Python' dict.'
// Note using (traditional) iterators instead of ranged for loop
cout << "Unordered_map<string, int>" << endl;
for (auto i: v4) {
cout << i.first << ": " << i.second << endl;
}
cout << endl;
}
%%bash
g++ -o stl.exe stl.cpp -std=c++14
%%bash
./stl.exe
```
**Exercise 3**.
Write a C++ program ex03.cpp that implements a map(func, xs) function where xs is an vector of integers and func is any function that takes an integer and returns another integer. Map should return an vector of integers. Test it on a vector [1,2,3,4] and an anonymous function that adds 3 to its input argument.
```
%%file ex03.cpp
#include <iostream>
#include <vector>
#include <functional>
using std::cout;
using std::vector;
using std::function;
using func = function<int(int)>;
vector<int> map(func f, vector<int> xs) {
vector<int> res;
for (auto x: xs) {
res.push_back(f(x));
}
return res;
}
int main() {
vector<int> xs = {1,2,3,4};
vector<int> ys = map([](int x){ return 3+x;}, xs);
for (auto y: ys) {
cout << y << "\n";
}
}
%%bash
g++ -o ex03.exe ex03.cpp -std=c++14
%%bash
./ex03.exe
```
## STL algorithms
```
%%file stl_algorithm.cpp
#include <vector>
#include <iostream>
#include <numeric>
using std::cout;
using std::endl;
using std::vector;
using std::begin;
using std::end;
int main() {
vector<int> v(10);
// iota is somewhat like range
std::iota(v.begin(), v.end(), 1);
for (auto i: v) {
cout << i << " ";
}
cout << endl;
// C++ version of reduce
cout << std::accumulate(begin(v), end(v), 0) << endl;
// Accumulate with lambda
cout << std::accumulate(begin(v), end(v), 1, [](int a, int b){return a * b; }) << endl;
}
%%bash
g++ -o stl_algorithm.exe stl_algorithm.cpp -std=c++14
%%bash
./stl_algorithm.exe
```
## Random numbers
```
%%file random.cpp
#include <iostream>
#include <random>
#include <functional>
using std::cout;
using std::default_random_engine;
using std::uniform_int_distribution;
using std::poisson_distribution;
using std::student_t_distribution;
using std::bind;
// start random number engine with fixed seed
default_random_engine re{12345};
uniform_int_distribution<int> uniform(1,6); // lower and upper bounds
poisson_distribution<int> poisson(30); // rate
student_t_distribution<double> t(10); // degrees of freedom
int main()
{
cout << "\nGenerating random numbers\n";
auto runif = bind (uniform, re);
auto rpois = bind(poisson, re);
auto rt = bind(t, re);
for (int i=0; i<10; i++) {
cout << runif() << ", " << rpois() << ", " << rt() << "\n";
}
}
%%bash
g++ -o random.exe random.cpp -std=c++14
%%bash
./random.exe
```
**Exercise 4**
Generate 100 numbers from $N(100, 15)$ in C++. Write to a plain text file that can be read in Python. Plot a normalized histogram of the numbers.
```
%%file ex04.cpp
#include <iostream>
#include <fstream>
#include <random>
#include <functional>
using std::cout;
using std::endl;
using std::ofstream;
using std::default_random_engine;
using std::normal_distribution;
using std::bind;
// start random number engine with fixed seed
default_random_engine re{12345};
normal_distribution<double> norm(100, 15); // mean and standard deviation
auto rnorm = bind(norm, re);
int main() {
ofstream fout("norm_data.txt");
for (int i=0; i<100; i++) {
fout << rnorm() << "\n";
}
}
%%bash
g++ -o ex04.exe ex04.cpp -std=c++14
%%bash
./ex04.exe
import matplotlib.pyplot as plt
import numpy as np
try:
data = np.loadtxt('norm_data.txt')
plt.hist(data, density=True)
plt.show()
except FileNotFoundError:
pass
```
## Numerics
Using the Eigen library.
```
%%file numeric.cpp
#include <iostream>
#include <fstream>
#include <random>
#include <Eigen/Dense>
#include <functional>
using std::cout;
using std::endl;
using std::ofstream;
using std::default_random_engine;
using std::normal_distribution;
using std::bind;
// start random number engine with fixed seed
default_random_engine re{12345};
normal_distribution<double> norm(5,2); // mean and standard deviation
auto rnorm = bind(norm, re);
int main()
{
using namespace Eigen;
VectorXd x1(6);
x1 << 1, 2, 3, 4, 5, 6;
VectorXd x2 = VectorXd::LinSpaced(6, 1, 2);
VectorXd x3 = VectorXd::Zero(6);
VectorXd x4 = VectorXd::Ones(6);
VectorXd x5 = VectorXd::Constant(6, 3);
VectorXd x6 = VectorXd::Random(6);
double data[] = {6,5,4,3,2,1};
Map<VectorXd> x7(data, 6);
VectorXd x8 = x6 + x7;
MatrixXd A1(3,3);
A1 << 1 ,2, 3,
4, 5, 6,
7, 8, 9;
MatrixXd A2 = MatrixXd::Constant(3, 4, 1);
MatrixXd A3 = MatrixXd::Identity(3, 3);
Map<MatrixXd> A4(data, 3, 2);
MatrixXd A5 = A4.transpose() * A4;
MatrixXd A6 = x7 * x7.transpose();
MatrixXd A7 = A4.array() * A4.array();
MatrixXd A8 = A7.array().log();
MatrixXd A9 = A8.unaryExpr([](double x) { return exp(x); });
MatrixXd A10 = MatrixXd::Zero(3,4).unaryExpr([](double x) { return rnorm(); });
VectorXd x9 = A1.colwise().norm();
VectorXd x10 = A1.rowwise().sum();
MatrixXd A11(x1.size(), 3);
A11 << x1, x2, x3;
MatrixXd A12(3, x1.size());
A12 << x1.transpose(),
x2.transpose(),
x3.transpose();
JacobiSVD<MatrixXd> svd(A10, ComputeThinU | ComputeThinV);
cout << "x1: comman initializer\n" << x1.transpose() << "\n\n";
cout << "x2: linspace\n" << x2.transpose() << "\n\n";
cout << "x3: zeors\n" << x3.transpose() << "\n\n";
cout << "x4: ones\n" << x4.transpose() << "\n\n";
cout << "x5: constant\n" << x5.transpose() << "\n\n";
cout << "x6: rand\n" << x6.transpose() << "\n\n";
cout << "x7: mapping\n" << x7.transpose() << "\n\n";
cout << "x8: element-wise addition\n" << x8.transpose() << "\n\n";
cout << "max of A1\n";
cout << A1.maxCoeff() << "\n\n";
cout << "x9: norm of columns of A1\n" << x9.transpose() << "\n\n";
cout << "x10: sum of rows of A1\n" << x10.transpose() << "\n\n";
cout << "head\n";
cout << x1.head(3).transpose() << "\n\n";
cout << "tail\n";
cout << x1.tail(3).transpose() << "\n\n";
cout << "slice\n";
cout << x1.segment(2, 3).transpose() << "\n\n";
cout << "Reverse\n";
cout << x1.reverse().transpose() << "\n\n";
cout << "Indexing vector\n";
cout << x1(0);
cout << "\n\n";
cout << "A1: comma initilizer\n";
cout << A1 << "\n\n";
cout << "A2: constant\n";
cout << A2 << "\n\n";
cout << "A3: eye\n";
cout << A3 << "\n\n";
cout << "A4: mapping\n";
cout << A4 << "\n\n";
cout << "A5: matrix multiplication\n";
cout << A5 << "\n\n";
cout << "A6: outer product\n";
cout << A6 << "\n\n";
cout << "A7: element-wise multiplication\n";
cout << A7 << "\n\n";
cout << "A8: ufunc log\n";
cout << A8 << "\n\n";
cout << "A9: custom ufucn\n";
cout << A9 << "\n\n";
cout << "A10: custom ufunc for normal deviates\n";
cout << A10 << "\n\n";
cout << "A11: np.c_\n";
cout << A11 << "\n\n";
cout << "A12: np.r_\n";
cout << A12 << "\n\n";
cout << "2x2 block startign at (0,1)\n";
cout << A1.block(0,1,2,2) << "\n\n";
cout << "top 2 rows of A1\n";
cout << A1.topRows(2) << "\n\n";
cout << "bottom 2 rows of A1";
cout << A1.bottomRows(2) << "\n\n";
cout << "leftmost 2 cols of A1";
cout << A1.leftCols(2) << "\n\n";
cout << "rightmost 2 cols of A1";
cout << A1.rightCols(2) << "\n\n";
cout << "Diagonal elements of A1\n";
cout << A1.diagonal() << "\n\n";
A1.diagonal() = A1.diagonal().array().square();
cout << "Transforming diagonal eelemtns of A1\n";
cout << A1 << "\n\n";
cout << "Indexing matrix\n";
cout << A1(0,0) << "\n\n";
cout << "singular values\n";
cout << svd.singularValues() << "\n\n";
cout << "U\n";
cout << svd.matrixU() << "\n\n";
cout << "V\n";
cout << svd.matrixV() << "\n\n";
}
%%bash
g++ -o numeric.exe numeric.cpp -std=c++14 -I/usr/include/eigen3
%%bash
./numeric.exe
```
### Check SVD
```
import numpy as np
A10 = np.array([
[5.17237, 3.73572, 6.29422, 6.55268],
[5.33713, 3.88883, 1.93637, 4.39812],
[8.22086, 6.94502, 6.36617, 6.5961]
])
U, s, Vt = np.linalg.svd(A10, full_matrices=False)
s
U
Vt.T
```
| github_jupyter |
# Refinitiv Data Platform Library for Python
## Delivery - EndPoint - Quantitative Analytics - Financial Contracts
This notebook demonstrates how the Delivery Layer of the library can be used to perform Quantitative Analytics on the Refinitiv Data Platform - using the Delivery Layer's Endpoint interface to retrieve content directly from the Endpoint.
## Set the location of the configuration file
For ease of use, you can set various initialization parameters of the RD Library in the **_refinitiv-data.config.json_** configuration file - as described in the Quick Start -> Sessions example.
### One config file for the tutorials
As these tutorial Notebooks are categorised into sub-folders and to avoid the need for multiple config files, we will use the _RD_LIB_CONFIG_PATH_ environment variable to point to a single instance of the config file in the top-level ***Configuration*** folder.
Before proceeding, please **ensure you have entered your credentials** into the config file in the ***Configuration*** folder.
```
import os
os.environ["RD_LIB_CONFIG_PATH"] = "../../../Configuration"
from refinitiv.data.delivery import endpoint_request
import refinitiv.data as rd
import pandas as pd
import json
```
## Open the default session
To open the default session ensure you have a '*refinitiv-data.config.json*' in the ***Configuration*** directory, populated with your credentials and specified a 'default' session in the config file
```
rd.open_session()
```
## Define the endpoint request
```
request_definition = rd.delivery.endpoint_request.Definition(
url = "/data/quantitative-analytics/beta1/financial-contracts",
method = rd.delivery.endpoint_request.RequestMethod.POST,
body_parameters = {
"fields": [
"InstrumentCode",
"BondType",
"IssueDate",
"EndDate",
"CouponRatePercent",
"Accrued",
"CleanPrice",
"DirtyPrice",
"YieldPercent",
"RedemptionDate",
"ModifiedDuration",
"Duration",
"DV01Bp",
"AverageLife",
"Convexity"
],
"outputs": [
"Headers",
"Data"
],
"universe": [
{
"instrumentType": "Bond",
"instrumentDefinition": {
"instrumentTag": "TreasuryBond_10Y",
"instrumentCode": "US10YT=RR"
}
}
]
}
)
```
## Send a request
```
response = request_definition.get_data()
```
## Display the result
Notice how the response contains the column headings for the data
```
if response.is_success:
headers = [h['name'] for h in response.data.raw['headers']]
df = pd.DataFrame(data=response.data.raw['data'], columns=headers)
display(df)
```
## Close the session
```
rd.close_session()
```
| github_jupyter |
```
import tensorflow as tf
```
# Introduction to input function
Here is a skeleton of basic input function:
```
def my_input_fn():
# Preprocess your data here...
# ...then return 1) a mapping of feature columns to Tensors with
# the corresponding feature data, and 2) a Tensor containing labels
return feature_cols, labels
```
## Function of input function
input function must return
* feature columns
a dict contain key/value paires that map feature column name to `Tensors` containing the corresponding `feature data`.
* labels
a `Tensor` contain your label data
## Converting feature data to tensor
As we said above, **feature column** need **Tensor** that contains feature data. so we need convert our data into **Tensor**
### python array, numpy array, dataframe
if you data is python array, numpy array, dataframe, you can use following data to convert to Tensor.
```
import numpy as np
# numpy input_fn.
my_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": np.array(x_data)},
y=np.array(y_data),
...)
import pandas as pd
# pandas input_fn.
my_input_fn = tf.estimator.inputs.pandas_input_fn(
x=pd.DataFrame({"x": x_data}),
y=pd.Series(y_data),
...)
```
you can check this [official tutorial](https://tensorflow.google.cn/get_started/estimator#construct_a_deep_neural_network_classifier) for a example and run it to get a feeling of it.
these 2 function have other paramter for you to customize the input function. check it out at official documetns:
Here is a list:
* batch_size: int, size of batches to return.
* num_epochs: int, number of epochs to iterate over data. If not None, read attempts that would exceed this value will raise OutOfRangeError.
* shuffle: bool, whether to read the records in random order.
* queue_capacity: int, size of the read queue. If None, it will be set roughly to the size of x.
* num_threads: Integer, number of threads used for reading and enqueueing. In order to have predicted and repeatable order of reading and enqueueing, such as in prediction and evaluation mode, num_threads should be 1.
* target_column: str, name to give the target column y.
## Trick
you can use ```functools.partial``` to wrap your function, so you don't need define separatly function to each type of task(training, validation, prediction)
```
my_input_fn(data_set):
# balalalal
# ...then return 1) a mapping of feature columns to Tensors with
# the corresponding feature data, and 2) a Tensor containing labels
return feature_cols, labels
```
---
```
classifier.train(
input_fn=functools.partial(my_input_fn, data_set=training_set),
steps=2000)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/tirthasheshpatel/Generative-Models/blob/master/VAE.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
%tensorflow_version 1.x
import tensorflow as tf
import keras
import numpy as np
import matplotlib.pyplot as plt
from keras.layers import Input, Dense, Lambda, InputLayer, concatenate, Dropout
from keras.models import Model, Sequential
from keras import backend as K
from keras import metrics
from keras.datasets import mnist
from keras.utils import np_utils
# Start tf session so we can run code.
sess = tf.InteractiveSession()
# Connect keras to the created session.
K.set_session(sess)
def vlb_binomial(x, x_decoded_mean, t_mean, t_log_var):
"""Returns the value of negative Variational Lower Bound
The inputs are tf.Tensor
x: (batch_size x number_of_pixels) matrix with one image per row with zeros and ones
x_decoded_mean: (batch_size x number_of_pixels) mean of the distribution p(x | t), real numbers from 0 to 1
t_mean: (batch_size x latent_dim) mean vector of the (normal) distribution q(t | x)
t_log_var: (batch_size x latent_dim) logarithm of the variance vector of the (normal) distribution q(t | x)
Returns:
A tf.Tensor with one element (averaged across the batch), VLB
"""
vlb = tf.reduce_mean(tf.reduce_sum(x * tf.log( x_decoded_mean+1e-19 ) + (1-x) * tf.log( 1-x_decoded_mean+1e-19 ), axis = 1 ) - 0.5 * tf.reduce_sum( -t_log_var + tf.exp( t_log_var ) + tf.square(t_mean) - 1 , axis = 1 ))
return -vlb
batch_size = 100
original_dim = 784 # Number of pixels in MNIST images.
latent_dim = 3 # d, dimensionality of the latent code t.
intermediate_dim = 128 # Size of the hidden layer.
epochs = 20
x = Input(batch_shape=(batch_size, original_dim))
def create_encoder(input_dim):
# Encoder network.
# We instantiate these layers separately so as to reuse them later
encoder = Sequential(name='encoder')
encoder.add(InputLayer([input_dim]))
encoder.add(Dense(intermediate_dim, activation='relu'))
encoder.add(Dense(2 * latent_dim))
return encoder
encoder = create_encoder(original_dim)
get_t_mean = Lambda(lambda h: h[:, :latent_dim])
get_t_log_var = Lambda(lambda h: h[:, latent_dim:])
h = encoder(x)
t_mean = get_t_mean(h)
t_log_var = get_t_log_var(h)
# Sampling from the distribution
# q(t | x) = N(t_mean, exp(t_log_var))
# with reparametrization trick.
def sampling(args):
"""Returns sample from a distribution N(args[0], diag(args[1]))
The sample should be computed with reparametrization trick.
The inputs are tf.Tensor
args[0]: (batch_size x latent_dim) mean of the desired distribution
args[1]: (batch_size x latent_dim) logarithm of the variance vector of the desired distribution
Returns:
A tf.Tensor of size (batch_size x latent_dim), the samples.
"""
t_mean, t_log_var = args
# YOUR CODE HERE
epsilon = K.random_normal(t_mean.shape)
z = epsilon*K.exp(0.5*t_log_var) + t_mean
return z
t = Lambda(sampling)([t_mean, t_log_var])
def create_decoder(input_dim):
# Decoder network
# We instantiate these layers separately so as to reuse them later
decoder = Sequential(name='decoder')
decoder.add(InputLayer([input_dim]))
decoder.add(Dense(intermediate_dim, activation='relu'))
decoder.add(Dense(original_dim, activation='sigmoid'))
return decoder
decoder = create_decoder(latent_dim)
x_decoded_mean = decoder(t)
loss = vlb_binomial(x, x_decoded_mean, t_mean, t_log_var)
vae = Model(x, x_decoded_mean)
# Keras will provide input (x) and output (x_decoded_mean) to the function that
# should construct loss, but since our function also depends on other
# things (e.g. t_means), it is easier to build the loss in advance and pass
# a function that always returns it.
vae.compile(optimizer=keras.optimizers.RMSprop(lr=0.001), loss=lambda x, y: loss)
# train the VAE on MNIST digits
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# One hot encoding.
y_train = np_utils.to_categorical(y_train)
y_test = np_utils.to_categorical(y_test)
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
x_train = x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test = x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
hist = vae.fit(x=x_train, y=x_train,
shuffle=True,
epochs=epochs,
batch_size=batch_size,
validation_data=(x_test, x_test),
verbose=2)
fig = plt.figure(figsize=(10, 10))
for fid_idx, (data, title) in enumerate(
zip([x_train, x_test], ['Train', 'Validation'])):
n = 10 # figure with 10 x 2 digits
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * 2))
decoded = sess.run(x_decoded_mean, feed_dict={x: data[:batch_size, :]})
for i in range(10):
figure[i * digit_size: (i + 1) * digit_size,
:digit_size] = data[i, :].reshape(digit_size, digit_size)
figure[i * digit_size: (i + 1) * digit_size,
digit_size:] = decoded[i, :].reshape(digit_size, digit_size)
ax = fig.add_subplot(1, 2, fid_idx + 1)
ax.imshow(figure, cmap='Greys_r')
ax.set_title(title)
ax.axis('off')
plt.show()
n_samples = 10
p_samples = tf.random_normal([n_samples, latent_dim], 0.0, 1.0)
# sampled_im_mean is a tf.Tensor of size 10 x 784 with 10 random
# images sampled from the vae model.
sampled_im_mean = decoder(p_samples)
sampled_im_mean_np = sess.run(sampled_im_mean)
# Show the sampled images.
plt.figure()
for i in range(n_samples):
ax = plt.subplot(n_samples // 5 + 1, 5, i + 1)
plt.imshow(sampled_im_mean_np[i, :].reshape(28, 28), cmap='gray')
ax.axis('off')
plt.show()
```
| github_jupyter |
```
# general purpose modules for handling data
import numpy as np
from numpy import array
import pandas as pd
from pandas import ExcelWriter
from pandas import ExcelFile
# for loading telo data column containing individual
# telomere length values
from ast import literal_eval
# custom module for handling telomere length data
import telomere_methods_astros as telo_ma
import importlib
%load_ext autoreload
%autoreload 2
%reload_ext autoreload
```
---
#####
The dataframe containing our telomere length data already contains the means. However, we're interested in analyzing (graphing, stats) the individual telomere length measurements. To do so we must extract the values from the list, while keeping the values associated with their timepont/individual. We'll achieve this by 'exploding' the cell containing individual telomere length measurements as a list, into one row per measurement (5520 rows per sample, 184 telos per 30 metaphases per sample).
As well, we're interested in defining and examining the prevalence of short/long telomeres; this is achieved by finding the values that divide the data into bottom 25% (short telos), middle 50%, and top 25% (long telos). More on that soon.
#####
---
### Reading astronaut dataframe from file
```
astro_df = pd.read_csv('../excel data/All_astronauts_telomere_length_dataframe.csv')
# literal eval enables interpretation of individual telomere length values
# in the list
astro_df['telo data'] = astro_df['telo data'].apply(lambda row: np.array(literal_eval(row)))
astro_df.head(4)
```
### Exploding list containing individual telomere length measurements into rows
this action explodes the list into columns bearing each datapoint.
picture the list, which contains the individual values, expanding to the
right, up to 5520 measurements
importantly, the index #s per row still refer to the timepont/sample
so we can merge these columns back to the original astro_df
then we'll melt the columns into one, resulting in a lot of rows
where each has one individual telomere length measurement
```
explode_telos_raw = astro_df['telo data'].apply(pd.Series)
explode_telos_raw.head(2)
exploded_telos_astro_df = (explode_telos_raw
# merge 5520 columns with original dataframe
.merge(astro_df, right_index = True, left_index = True)
# drop unnecessary columns
.drop(['telo data', 'Q1', 'Q2-3', 'Q4'], axis = 1)
#specify which columns remain constant per indviidual telo
.melt(id_vars = ['astro number', 'astro id', 'timepoint', 'flight status', 'telo means'], value_name = "telo data exploded")
.drop("variable", axis = 1)
.dropna())
exploded_telos_astro_df.shape
```
### Saving exploded individual telos for all astros dataframe to csv for later retrieval
```
copy_exploded_telos_astros_df = exploded_telos_astro_df
copy_exploded_telos_astros_df.to_csv('../excel data/exploded_telos_astros_df.csv', index = False)
```
### Defining short/mid/long telomeres w/ quartiles and counting per timepoint per astro
This function identifies the timepoint per astronaut most distal to spaceflight (L-270 or L-180) and identifies the individual telomere length values which divide the data into the bottom 25% (short telos), mid 50%, and top 25% (long telos). The function then counts how many telos reside in each quartile. Now we have a way to define short/long telomeres.
The function then applies those quartile cutoff values to subsequent datapoints and counts how many telomeres reside within each quartile. In doing so, we count the number of telomeres moving into/out of the quartiles, per timepoint per astronaut, and can quantitatively dicuss # of short/long telos.
```
quartiles_astro_df = telo_ma.make_quartiles_columns(astro_df)
%reload_ext autoreload
# for col in ['Q1', 'Q2-3', 'Q4']:
# quartiles_astro_df[col] = quartiles_astro_df[col].astype('int64')
quartiles_astro_df['Q1'] = quartiles_astro_df['Q1'].astype('int64')
quartiles_astro_df['Q2-3'] = quartiles_astro_df['Q2-3'].astype('int64')
quartiles_astro_df['Q4'] = quartiles_astro_df['Q4'].astype('int64')
quartiles_astro_df[quartiles_astro_df['astro id'] == 2171]
melted_quartiles_astro_df = pd.melt(
quartiles_astro_df,
id_vars=['astro number', 'astro id', 'timepoint', 'flight status', 'telo data', 'telo means'],
# id_vars = [col for col in astro_df.columns if col != 'Q1' and col != 'Q2-3' and col != 'Q4'],
var_name='relative Q',
value_name='Q freq counts')
melted_quartiles_astro_df['Q freq counts'] = melted_quartiles_astro_df['Q freq counts'].astype('int64')
melted_quartiles_astro_df['astro id'] = melted_quartiles_astro_df['astro id'].astype('str')
melted_quartiles_astro_df[melted_quartiles_astro_df['astro id'] == '5163'].head(8)
```
### Saving melted-quartile-count all astros dataframe for later retrieval
```
copy_melted_quartiles_astro_df = melted_quartiles_astro_df
copy_melted_quartiles_astro_df['telo data'] = copy_melted_quartiles_astro_df['telo data'].apply(lambda row: row.tolist())
melted_quartiles_astro_df.to_csv('../excel data/melted_quartiles_astro_df.csv', index = False)
```
| github_jupyter |
# Handling uncertainty with quantile regression
```
%matplotlib inline
```
[Quantile regression](https://www.wikiwand.com/en/Quantile_regression) is useful when you're not so much interested in the accuracy of your model, but rather you want your model to be good at ranking observations correctly. The typical way to perform quantile regression is to use a special loss function, namely the quantile loss. The quantile loss takes a parameter, $\alpha$ (alpha), which indicates which quantile the model should be targeting. In the case of $\alpha = 0.5$, then this is equivalent to asking the model to predict the median value of the target, and not the most likely value which would be the mean.
A nice thing we can do with quantile regression is to produce a prediction interval for each prediction. Indeed, if we predict the lower and upper quantiles of the target then we will be able to obtain a "trust region" in between which the true value is likely to belong. Of course, the likeliness will depend on the chosen quantiles. For a slightly more detailed explanation see [this](https://medium.com/the-artificial-impostor/quantile-regression-part-1-e25bdd8d9d43) blog post.
As an example, let us take the [simple time series model we built in another notebook](building-a-simple-time-series-model.md). Instead of predicting the mean value of the target distribution, we will predict the 5th, 50th, 95th quantiles. This will require training three separate models, so we will encapsulate the model building logic in a function called `make_model`. We also have to slightly adapt the training loop, but not by much. Finally, we will draw the prediction interval along with the predictions from for 50th quantile (i.e. the median) and the true values.
```
import calendar
import math
import matplotlib.pyplot as plt
from river import compose
from river import datasets
from river import linear_model
from river import metrics
from river import optim
from river import preprocessing
from river import stats
from river import time_series
def get_ordinal_date(x):
return {'ordinal_date': x['month'].toordinal()}
def get_month_distances(x):
return {
calendar.month_name[month]: math.exp(-(x['month'].month - month) ** 2)
for month in range(1, 13)
}
def make_model(alpha):
extract_features = compose.TransformerUnion(get_ordinal_date, get_month_distances)
scale = preprocessing.StandardScaler()
learn = linear_model.LinearRegression(
intercept_lr=0,
optimizer=optim.SGD(3),
loss=optim.losses.Quantile(alpha=alpha)
)
model = extract_features | scale | learn
model = time_series.Detrender(regressor=model, window_size=12)
return model
metric = metrics.MAE()
models = {
'lower': make_model(alpha=0.05),
'center': make_model(alpha=0.5),
'upper': make_model(alpha=0.95)
}
dates = []
y_trues = []
y_preds = {
'lower': [],
'center': [],
'upper': []
}
for x, y in datasets.AirlinePassengers():
y_trues.append(y)
dates.append(x['month'])
for name, model in models.items():
y_preds[name].append(model.predict_one(x))
model.learn_one(x, y)
# Update the error metric
metric.update(y, y_preds['center'][-1])
# Plot the results
fig, ax = plt.subplots(figsize=(10, 6))
ax.grid(alpha=0.75)
ax.plot(dates, y_trues, lw=3, color='#2ecc71', alpha=0.8, label='Truth')
ax.plot(dates, y_preds['center'], lw=3, color='#e74c3c', alpha=0.8, label='Prediction')
ax.fill_between(dates, y_preds['lower'], y_preds['upper'], color='#e74c3c', alpha=0.3, label='Prediction interval')
ax.legend()
ax.set_title(metric);
```
An important thing to note is that the prediction interval we obtained should not be confused with a confidence interval. Simply put, a prediction interval represents uncertainty for where the true value lies, whereas a confidence interval encapsulates the uncertainty on the prediction. You can find out more by reading [this](https://stats.stackexchange.com/questions/16493/difference-between-confidence-intervals-and-prediction-intervals) CrossValidated post.
| github_jupyter |
# seaborn.countplot
---
Bar graphs are useful for displaying relationships between categorical data and at least one numerical variable. `seaborn.countplot` is a barplot where the dependent variable is the number of instances of each instance of the independent variable.
dataset: [IMDB 5000 Movie Dataset](https://www.kaggle.com/deepmatrix/imdb-5000-movie-dataset)
```
%matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
plt.rcParams['figure.figsize'] = (20.0, 10.0)
df = pd.read_csv('../../../datasets/movie_metadata.csv')
df.head()
```
For the bar plot, let's look at the number of movies in each category, allowing each movie to be counted more than once.
```
# split each movie's genre list, then form a set from the unwrapped list of all genres
categories = set([s for genre_list in df.genres.unique() for s in genre_list.split("|")])
# one-hot encode each movie's classification
for cat in categories:
df[cat] = df.genres.transform(lambda s: int(cat in s))
# drop other columns
df = df[['director_name','genres','duration'] + list(categories)]
df.head()
# convert from wide to long format and remove null classificaitons
df = pd.melt(df,
id_vars=['duration'],
value_vars = list(categories),
var_name = 'Category',
value_name = 'Count')
df = df.loc[df.Count>0]
# add an indicator whether a movie is short or long, split at 100 minutes runtime
df['islong'] = df.duration.transform(lambda x: int(x > 100))
# sort in descending order
#df = df.loc[df.groupby('Category').transform(sum).sort_values('Count', ascending=False).index]
df.head()
```
Basic plot
```
p = sns.countplot(data=df, x = 'Category')
```
color by a category
```
p = sns.countplot(data=df,
x = 'Category',
hue = 'islong')
```
make plot horizontal
```
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong')
```
Saturation
```
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.5)
```
Various palettes
```
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'deep')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'muted')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'pastel')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'bright')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'dark')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'colorblind')
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = ((50/255, 132/255.0, 191/255.0), (255/255.0, 232/255.0, 0/255.0)))
p = sns.countplot(data=df,
y = 'Category',
hue = 'islong',
saturation=.9,
palette = 'Dark2')
help(sns.color_palette)
help(sns.countplot)
sns.set(rc={"axes.facecolor":"#ccddff",
"axes.grid":False,
'axes.labelsize':30,
'figure.figsize':(20.0, 10.0),
'xtick.labelsize':25,
'ytick.labelsize':20})
p = sns.countplot(data=df, x = 'Category')
plt.text(9,2000, "Color Palettes", fontsize = 95, color='black', fontstyle='italic')
p.get_figure().savefig('../../figures/colors.png')
```
| github_jupyter |
# Create instagram post list
This notebook creates a csv with information on X amount of Flickr images per places in our database. It does so using scripts for the following 3 steps:
1. For each destination, query the top X most interesting images from Flickr.
2. For each author found, query people info to know more about the author.
3. For each place, query the wikivoyage place url for a quick link to more info.
All (intermediate) query results are saved so that we don't need to query again.
In this first run, we query the top 20 images per place. We better have the data, and then we can always do the image generation only for the top 5.
```
%load_ext autoreload
%autoreload 2
data_dir = '../../data/'
TOP_X_IMAGES = 5
```
### Init
```
import pandas as pd
from stairway.apis.flickr.people import get_flickr_people_info, parse_flickr_people_info
from stairway.apis.wikivoyage.page_info import get_wikivoyage_page_info
from stairway.instagram.description import pick_generic_hastags, clean_text, create_draft_description
```
### Read place data
```
# places_file = data_dir + 'wikivoyage/enriched/wikivoyage_destinations.csv'
# df = (
# pd.read_csv(places_file)
# .rename(columns={'id': 'stairway_id'})
# .set_index("stairway_id", drop=False)
# [['stairway_id', 'name', 'country', 'nr_tokens', 'wiki_id']]
# )
# df.shape
```
## 1. Query the api and explode the DF
Implementation moved to `scripts/flickr_image_list.py`.
```
df_images = pd.read_csv(data_dir + 'flickr/flickr_image_list.csv')
df_images.shape
```
## 2. Get Flickr people info
Use https://www.flickr.com/services/api/flickr.people.getInfo.html
First deduplicate the authors from the image list, then retrieve info and join back to avoid querying a single author multiple times.
```
# user = '12962905@N05' #kevinpoh
# user = '61713368@N07' #tiket2
# user = '143466180@N07' # removed user
# output = get_flickr_people_info(user, api_key=FLICKR_KEY)
# output = parse_flickr_people_info(output)
# output
```
Implementation moved to `scripts/flickr_people_list.py`.
```
df_people = pd.read_csv(data_dir + "flickr/flickr_people_list.csv").drop_duplicates()
df_people.shape
```
## 3. Add link to wiki travel for ease of use
Use `wiki_id` and query up to 50 wiki_ids at the same time.
Implementation moved to `scripts/wikivoyage_page_info.py`.
```
# data = get_wikivoyage_page_info([10, 33, 36])
# [v['fullurl'] for k, v in data.items()]
df_wiki_info = pd.read_csv(data_dir + "wikivoyage/clean/wikivoyage_page_info.csv").drop_duplicates()
df_wiki_info.shape
```
## Join all tables together
Now join the people table with the image table
```
df_all = (
df_images
.merge(df_people, left_on='owner', right_on="nsid")
)
df_all.shape
```
Now join the wiki links with the image table
```
df_all = (
df_all
# there is a conflicting 'title' in the image dataset
.merge(df_wiki_info.drop(columns=['title']), left_on='wiki_id', right_on='pageid')
)
df_all.shape
```
## 4. Add a draft description to get started
```
df_all['draft_text'] = df_all.apply(lambda df: create_draft_description(df['name'], df['country'], df['path_alias']), axis=1)
df_all.shape
```
## Dump the final list to file
Last step, is making a nice subselection of the variables and putting them in de right order for the overview.
```
column_rename = {'index': 'image_nr',
'title': 'image_title',
'path_alias': 'owner_tag',
'location': 'owner_location',
'fullurl': 'wiki_url'}
column_order = ['stairway_id', 'index', 'name', 'country', 'article_length', 'title', 'ownername',
'realname', 'path_alias', 'location', 'profileurl', 'image_url', 'fullurl', 'draft_text']
df_insta = (
df_all
.assign(article_length = lambda df: df['length'].astype(int))
.loc[lambda df: df['index'] < TOP_X_IMAGES]
[column_order]
.rename(columns=column_rename)
)
df_insta.shape
df_insta.to_csv(data_dir + 'instagram/instagram_post_list.csv', index=False)
```
Next steps:
1. Import the CSV into Google Spreadsheets
2. Process the images in this list and upload them in Google Drive.
Done.
| github_jupyter |
```
import glob, re
import numpy as np
import pandas as pd
from sklearn import *
from datetime import datetime
from xgboost import XGBRegressor
data = {
'tra': pd.read_csv('../input/air_visit_data.csv'),
'as': pd.read_csv('../input/air_store_info.csv'),
'hs': pd.read_csv('../input/hpg_store_info.csv'),
'ar': pd.read_csv('../input/air_reserve.csv'),
'hr': pd.read_csv('../input/hpg_reserve.csv'),
'id': pd.read_csv('../input/store_id_relation.csv'),
'tes': pd.read_csv('../input/sample_submission.csv'),
'hol': pd.read_csv('../input/date_info.csv').rename(columns={'calendar_date':'visit_date'})
}
data['hr'] = pd.merge(data['hr'], data['id'], how='inner', on=['hpg_store_id'])
for df in ['ar','hr']:
data[df]['visit_datetime'] = pd.to_datetime(data[df]['visit_datetime'])
data[df]['visit_datetime'] = data[df]['visit_datetime'].dt.date
data[df]['reserve_datetime'] = pd.to_datetime(data[df]['reserve_datetime'])
data[df]['reserve_datetime'] = data[df]['reserve_datetime'].dt.date
data[df]['reserve_datetime_diff'] = data[df].apply(lambda r: (r['visit_datetime'] - r['reserve_datetime']).days, axis=1)
tmp1 = data[df].groupby(['air_store_id','visit_datetime'], as_index=False)[['reserve_datetime_diff', 'reserve_visitors']].sum().rename(columns={'visit_datetime':'visit_date', 'reserve_datetime_diff': 'rs1', 'reserve_visitors':'rv1'})
tmp2 = data[df].groupby(['air_store_id','visit_datetime'], as_index=False)[['reserve_datetime_diff', 'reserve_visitors']].mean().rename(columns={'visit_datetime':'visit_date', 'reserve_datetime_diff': 'rs2', 'reserve_visitors':'rv2'})
data[df] = pd.merge(tmp1, tmp2, how='inner', on=['air_store_id','visit_date'])
data['tra']['visit_date'] = pd.to_datetime(data['tra']['visit_date'])
data['tra']['dow'] = data['tra']['visit_date'].dt.dayofweek
data['tra']['year'] = data['tra']['visit_date'].dt.year
data['tra']['month'] = data['tra']['visit_date'].dt.month
data['tra']['visit_date'] = data['tra']['visit_date'].dt.date
data['tes']['visit_date'] = data['tes']['id'].map(lambda x: str(x).split('_')[2])
data['tes']['air_store_id'] = data['tes']['id'].map(lambda x: '_'.join(x.split('_')[:2]))
data['tes']['visit_date'] = pd.to_datetime(data['tes']['visit_date'])
data['tes']['dow'] = data['tes']['visit_date'].dt.dayofweek
data['tes']['year'] = data['tes']['visit_date'].dt.year
data['tes']['month'] = data['tes']['visit_date'].dt.month
data['tes']['visit_date'] = data['tes']['visit_date'].dt.date
unique_stores = data['tes']['air_store_id'].unique()
stores = pd.concat([pd.DataFrame({'air_store_id': unique_stores, 'dow': [i]*len(unique_stores)}) for i in range(7)], axis=0, ignore_index=True).reset_index(drop=True)
#sure it can be compressed...
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].min().rename(columns={'visitors':'min_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].mean().rename(columns={'visitors':'mean_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].median().rename(columns={'visitors':'median_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].max().rename(columns={'visitors':'max_visitors'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
tmp = data['tra'].groupby(['air_store_id','dow'], as_index=False)['visitors'].count().rename(columns={'visitors':'count_observations'})
stores = pd.merge(stores, tmp, how='left', on=['air_store_id','dow'])
stores = pd.merge(stores, data['as'], how='left', on=['air_store_id'])
# NEW FEATURES FROM Georgii Vyshnia
stores['air_genre_name'] = stores['air_genre_name'].map(lambda x: str(str(x).replace('/',' ')))
stores['air_area_name'] = stores['air_area_name'].map(lambda x: str(str(x).replace('-',' ')))
lbl = preprocessing.LabelEncoder()
for i in range(10):
stores['air_genre_name'+str(i)] = lbl.fit_transform(stores['air_genre_name'].map(lambda x: str(str(x).split(' ')[i]) if len(str(x).split(' '))>i else ''))
stores['air_area_name'+str(i)] = lbl.fit_transform(stores['air_area_name'].map(lambda x: str(str(x).split(' ')[i]) if len(str(x).split(' '))>i else ''))
stores['air_genre_name'] = lbl.fit_transform(stores['air_genre_name'])
stores['air_area_name'] = lbl.fit_transform(stores['air_area_name'])
data['hol']['visit_date'] = pd.to_datetime(data['hol']['visit_date'])
data['hol']['day_of_week'] = lbl.fit_transform(data['hol']['day_of_week'])
data['hol']['visit_date'] = data['hol']['visit_date'].dt.date
train = pd.merge(data['tra'], data['hol'], how='left', on=['visit_date'])
test = pd.merge(data['tes'], data['hol'], how='left', on=['visit_date'])
train = pd.merge(train, stores, how='left', on=['air_store_id','dow'])
test = pd.merge(test, stores, how='left', on=['air_store_id','dow'])
for df in ['ar','hr']:
train = pd.merge(train, data[df], how='left', on=['air_store_id','visit_date'])
test = pd.merge(test, data[df], how='left', on=['air_store_id','visit_date'])
train['id'] = train.apply(lambda r: '_'.join([str(r['air_store_id']), str(r['visit_date'])]), axis=1)
train['total_reserv_sum'] = train['rv1_x'] + train['rv1_y']
train['total_reserv_mean'] = (train['rv2_x'] + train['rv2_y']) / 2
train['total_reserv_dt_diff_mean'] = (train['rs2_x'] + train['rs2_y']) / 2
test['total_reserv_sum'] = test['rv1_x'] + test['rv1_y']
test['total_reserv_mean'] = (test['rv2_x'] + test['rv2_y']) / 2
test['total_reserv_dt_diff_mean'] = (test['rs2_x'] + test['rs2_y']) / 2
# NEW FEATURES FROM JMBULL
train['date_int'] = train['visit_date'].apply(lambda x: x.strftime('%Y%m%d')).astype(int)
test['date_int'] = test['visit_date'].apply(lambda x: x.strftime('%Y%m%d')).astype(int)
train['var_max_lat'] = train['latitude'].max() - train['latitude']
train['var_max_long'] = train['longitude'].max() - train['longitude']
test['var_max_lat'] = test['latitude'].max() - test['latitude']
test['var_max_long'] = test['longitude'].max() - test['longitude']
# NEW FEATURES FROM Georgii Vyshnia
train['lon_plus_lat'] = train['longitude'] + train['latitude']
test['lon_plus_lat'] = test['longitude'] + test['latitude']
lbl = preprocessing.LabelEncoder()
train['air_store_id2'] = lbl.fit_transform(train['air_store_id'])
test['air_store_id2'] = lbl.transform(test['air_store_id'])
col = [c for c in train if c not in ['id', 'air_store_id', 'visit_date','visitors']]
train = train.fillna(-1)
test = test.fillna(-1)
def RMSLE(y, pred):
return metrics.mean_squared_error(y, pred)**0.5
#model1 = ensemble.GradientBoostingRegressor(learning_rate=0.2, random_state=3, n_estimators=200, subsample=0.8,
# max_depth =10))
model2 = neighbors.KNeighborsRegressor(n_jobs=-1, n_neighbors=4)
model3 = XGBRegressor(learning_rate=0.2, seed=3, n_estimators=200, subsample=0.8,
colsample_bytree=0.8, max_depth =10)
#model1.fit(train[col], np.log1p(train['visitors'].values))
model2.fit(train[col], np.log1p(train['visitors'].values))
model3.fit(train[col], np.log1p(train['visitors'].values))
#preds1 = model1.predict(train[col])
preds2 = model2.predict(train[col])
preds3 = model3.predict(train[col])
#print('RMSE GradientBoostingRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds1))
print('RMSE KNeighborsRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds2))
print('RMSE XGBRegressor: ', RMSLE(np.log1p(train['visitors'].values), preds3))
from lightgbm import LGBMRegressor
model1 = LGBMRegressor(learning_rate=0.3, num_leaves=1400, max_depth=15, max_bin=300, min_child_weight=5)
model1.fit(train[col], np.log1p(train['visitors'].values))
preds1 = model1.predict(train[col])
print('RMSE LightGBM: ', RMSLE(np.log1p(train['visitors'].values), preds1))
preds1 = model1.predict(test[col])
preds2 = model2.predict(test[col])
preds3 = model3.predict(test[col])
pred = (preds1+preds2+preds3) / 3.0
test['visitors'] = pred
test['visitors'] = np.expm1(test['visitors']).clip(lower=0.)
sub1 = test[['id','visitors']].copy()
print "Hi"
from __future__ import division
# from hklee
# https://www.kaggle.com/zeemeen/weighted-mean-comparisons-lb-0-497-1st/code
dfs = { re.search('/([^/\.]*)\.csv', fn).group(1):
pd.read_csv(fn)for fn in glob.glob('../input/*.csv')}
for k, v in dfs.items(): locals()[k] = v
wkend_holidays = date_info.apply(
(lambda x:(x.day_of_week=='Sunday' or x.day_of_week=='Saturday') and x.holiday_flg==1), axis=1)
date_info.loc[wkend_holidays, 'holiday_flg'] = 0
date_info['weight'] = ((date_info.index + 1) / len(date_info)) ** 5
visit_data = air_visit_data.merge(date_info, left_on='visit_date', right_on='calendar_date', how='left')
visit_data.drop('calendar_date', axis=1, inplace=True)
visit_data['visitors'] = visit_data.visitors.map(pd.np.log1p)
wmean = lambda x:( (x.weight * x.visitors).sum() / x.weight.sum() )
visitors = visit_data.groupby(['air_store_id', 'day_of_week', 'holiday_flg']).apply(wmean).reset_index()
visitors.rename(columns={0:'visitors'}, inplace=True) # cumbersome, should be better ways.
sample_submission['air_store_id'] = sample_submission.id.map(lambda x: '_'.join(x.split('_')[:-1]))
sample_submission['calendar_date'] = sample_submission.id.map(lambda x: x.split('_')[2])
sample_submission.drop('visitors', axis=1, inplace=True)
sample_submission = sample_submission.merge(date_info, on='calendar_date', how='left')
sample_submission = sample_submission.merge(visitors, on=[
'air_store_id', 'day_of_week', 'holiday_flg'], how='left')
missings = sample_submission.visitors.isnull()
sample_submission.loc[missings, 'visitors'] = sample_submission[missings].merge(
visitors[visitors.holiday_flg==0], on=('air_store_id', 'day_of_week'),
how='left')['visitors_y'].values
missings = sample_submission.visitors.isnull()
sample_submission.loc[missings, 'visitors'] = sample_submission[missings].merge(
visitors[['air_store_id', 'visitors']].groupby('air_store_id').mean().reset_index(),
on='air_store_id', how='left')['visitors_y'].values
sample_submission['visitors'] = sample_submission.visitors.map(pd.np.expm1)
sub2 = sample_submission[['id', 'visitors']].copy()
sub_merge = pd.merge(sub1, sub2, on='id', how='inner')
sub_merge['visitors'] = (sub_merge['visitors_x'] + sub_merge['visitors_y']* 1.1)/2
sub_merge['visitors'] = sub_merge['visitors'].astype(int)
sub_merge[['id', 'visitors']].to_csv('submissionINT2.csv', index=False)
```
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 5GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
data = pd.read_csv('../input/eda-data/datasets_11262_15629_50_Startups.csv')
data.head()
data.describe()
!pip install pandas-profiling
import pandas_profiling
profile = data.profile_report(title = 'summary')
profile
data.head()
(data['R&D Spend']==0).sum()
(data['R&D Spend']==0).values.any()
data.dtypes
```
**** #UNIVARIATE ANALYSIS
COLUMNWISE
R&D SPEND ANALYSIS
```
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
data.index
#univariate scatter plot (SCATTER PLOT USING MATPLOTLIB)
plt.scatter(data.index , data['R&D Spend'])
plt.show()
#univariate scatterplot (SCATTER PLOT USING SEABORN)
sns.scatterplot(x = data.index , y = data['R&D Spend'] , hue = data['State'])
#univariate lineplot (LINE PLOT USING MATPLOTLIB)
plt.figure(figsize=(10,10))
plt.title("Line plot of R&D Spend")
plt.xlabel('index' , fontsize = 20)
plt.ylabel('R&D SPEND' , fontsize = 20)
plt.plot(data.index , data['R&D Spend'], markevery = 1,marker = 'o' )
#univariate lineplot (LINE PLOT USING MATPLOTLIB with COLORS WRT TO CLASSES) BY USING GROUP BY
plt.figure(figsize=(10,10))
plt.title("Line plot of R&D Spend")
plt.xlabel('index' , fontsize = 20)
plt.ylabel('R&D SPEND' , fontsize = 20)
for name , group in data.groupby('State'):
plt.plot(group.index , group['R&D Spend'] , label = name , markevery = 1,marker = 'o')
plt.legend()
plt.show()
'R&D Spend'
#univariate lineplot (LINE PLOT USING SEABORN with COLORS WRT TO CLASSES) BY USING GROUP BY
sns.set(rc = {'figure.figsize':(10,10)})
sns.set(font_scale=1.5)
fig = sns.lineplot(x = data.index , y = data['R&D Spend'] , markevery = 1 , marker = 'o' , data = data , hue = data['State'])
fig.set(xlabel = 'index');fig.set(ylabel = 'rd spend')
#comparisons between univariate analysis
#STRIP PLOT USING SEABORN
sns.stripplot(x = data['State'] , y=data['R&D Spend'])
# SWARM PLOT WITH SEARBORN
#The swarm-plot, similar to a strip-plot, provides a visualization technique for
#univariate data to view the spread of values in a continuous variable.
#The only difference between the strip-plot and the swarm-plot is that the
#swarm-plot spreads out the data points of the variable automatically to avoid
#overlap and hence provides a better visual overview of the data.
sns.set(rc = {'figure.figsize': (10,10)})
sns.swarmplot(y = data['R&D Spend'] , x = data['State'])
```
#UNIVARIATE SUMMARY PLOTS
```
#HISTOGRAM
#Histograms are similar to bar charts which display the counts
#or relative frequencies of values falling in different class intervals or ranges.
#A histogram displays the shape and spread of continuous sample data. It also helps us
#understand the skewness and kurtosis of the distribution of the data.
#HISTOGRAM USING MATPLOTLIB
plt.hist(data['R&D Spend'])
#HISTOGRAM USING SEABORN
sns.distplot(data['R&D Spend'] , kde = True , color = 'red' , bins = 10)# KDE IS KERNEL DENSITY : IT DRAWS LINE AROUNF BOUNDARIES OF HISTOGRAM
#DENSITY PLOTS(USING MATPLOTLIB) : SMOOTHER VERSION OF HISTOGRAM
# IT IS USED TO SHOW THE PROBABILITY DENSITY FUNTOIONS OF DATA
plt.figure(figsize=(10,10))
data['R&D Spend'].plot(kind = 'density')
#DENSITY PLOT USING SEABORN
sns.set(rc = {'figure.figsize':(20,20)})
sns.kdeplot(data['R&D Spend'],shade = True , color = 'red' , bw = 0.15)
sns.set(rc = {'figure.figsize':(10,10)})
sns.kdeplot(data['R&D Spend'],shade = True , color = 'red' )
sns.set(rc = {'figure.figsize':(10,10)})
sns.kdeplot(data['R&D Spend'],shade = True ,cbar = True )
#RUG PLOT
#A rug plot is a very simple, but also an ideal legitimate, way of representing a distribution.
#It consists of vertical lines at each data point. Here, the height is arbitrary.
#The density of the distribution can be known by how dense the tick-marks are.
#The connection between the rug plot and histogram is very direct:
# a histogram just creates bins along with the range of the data and then draws a
# bar with height equal to the number of ticks in each bin. In a rug plot, all of
# the data points are plotted on a single axis, one tick mark or line for each one.
# RUG PLOT IN SEABORN , subplot in MATPLOTLIB
fig , ax = plt.subplots()
#sns.rugplot(data['R&D Spend'])
sns.rugplot([1,2,3,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,44,4,4,4,4,4,4,4,4,4,4,5,6,7])
sns.kdeplot([1,2,3,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,4,44,4,4,4,4,4,4,4,4,4,4,5,6,7],shade = True ,cbar = True,bw = 0.0001 )
ax.set_xlim(0,9)
plt.show()
fig , ax = plt.subplots()
#sns.rugplot(data['R&D Spend'])
sns.rugplot(data['R&D Spend'] , color = 'orange')
sns.kdeplot(data['R&D Spend'],shade = True ,color = 'blue' )
#ax.set_xlim(0,9)
plt.show()
#INTEGRATION OF RUG PLOT IN SEABORN HISTOGRAM FUNCTION
sns.distplot(data['R&D Spend'] , rug = True , hist = True , color = 'Blue')
sns.distplot(data['R&D Spend'] , rug =False , hist = True , color = 'Blue')
sns.distplot(data['R&D Spend'] , rug = True , hist = False , color = 'orange')
sns.distplot(data['R&D Spend'] , rug = False , hist = False , color = 'green')
#BOX PLOTS
#A box-plot is a very useful and standardized way of displaying
#the distribution of data based on a five-number summary
#(minimum, first quartile, second quartile(median), third quartile, maximum).
#It helps in understanding these parameters of the distribution of data and is
#extremely helpful in detecting outliers.
```
| github_jupyter |
# Face Recognition
Welcome! In this assignment, you're going to build a face recognition system. Many of the ideas presented here are from [FaceNet](https://arxiv.org/pdf/1503.03832.pdf). In the lecture, you also encountered [DeepFace](https://research.fb.com/wp-content/uploads/2016/11/deepface-closing-the-gap-to-human-level-performance-in-face-verification.pdf).
Face recognition problems commonly fall into one of two categories:
**Face Verification** "Is this the claimed person?" For example, at some airports, you can pass through customs by letting a system scan your passport and then verifying that you (the person carrying the passport) are the correct person. A mobile phone that unlocks using your face is also using face verification. This is a 1:1 matching problem.
**Face Recognition** "Who is this person?" For example, the video lecture showed a [face recognition video](https://www.youtube.com/watch?v=wr4rx0Spihs) of Baidu employees entering the office without needing to otherwise identify themselves. This is a 1:K matching problem.
FaceNet learns a neural network that encodes a face image into a vector of 128 numbers. By comparing two such vectors, you can then determine if two pictures are of the same person.
By the end of this assignment, you'll be able to:
* Differentiate between face recognition and face verification
* Implement one-shot learning to solve a face recognition problem
* Apply the triplet loss function to learn a network's parameters in the context of face recognition
* Explain how to pose face recognition as a binary classification problem
* Map face images into 128-dimensional encodings using a pretrained model
* Perform face verification and face recognition with these encodings
**Channels-last notation**
For this assignment, you'll be using a pre-trained model which represents ConvNet activations using a "channels last" convention, as used during the lecture and in previous programming assignments.
In other words, a batch of images will be of shape $(m, n_H, n_W, n_C)$.
## Table of Contents
- [1 - Packages](#1)
- [2 - Naive Face Verification](#2)
- [3 - Encoding Face Images into a 128-Dimensional Vector](#3)
- [3.1 - Using a ConvNet to Compute Encodings](#3-1)
- [3.2 - The Triplet Loss](#3-2)
- [Exercise 1 - triplet_loss](#ex-1)
- [4 - Loading the Pre-trained Model](#4)
- [5 - Applying the Model](#5)
- [5.1 - Face Verification](#5-1)
- [Exercise 2 - verify](#ex-2)
- [5.2 - Face Recognition](#5-2)
- [Exercise 3 - who_is_it](#ex-3)
- [6 - References](#6)
<a name='1'></a>
## 1 - Packages
Go ahead and run the cell below to import the packages you'll need.
```
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Conv2D, ZeroPadding2D, Activation, Input, concatenate
from tensorflow.keras.models import Model
from tensorflow.keras.layers import BatchNormalization
from tensorflow.keras.layers import MaxPooling2D, AveragePooling2D
from tensorflow.keras.layers import Concatenate
from tensorflow.keras.layers import Lambda, Flatten, Dense
from tensorflow.keras.initializers import glorot_uniform
from tensorflow.keras.layers import Layer
from tensorflow.keras import backend as K
K.set_image_data_format('channels_last')
import os
import numpy as np
from numpy import genfromtxt
import pandas as pd
import tensorflow as tf
import PIL
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
<a name='2'></a>
## 2 - Naive Face Verification
In Face Verification, you're given two images and you have to determine if they are of the same person. The simplest way to do this is to compare the two images pixel-by-pixel. If the distance between the raw images is below a chosen threshold, it may be the same person!
<img src="images/pixel_comparison.png" style="width:380px;height:150px;">
<caption><center> <u> <font color='purple'> <b>Figure 1</b> </u></center></caption>
Of course, this algorithm performs poorly, since the pixel values change dramatically due to variations in lighting, orientation of the person's face, minor changes in head position, and so on.
You'll see that rather than using the raw image, you can learn an encoding, $f(img)$.
By using an encoding for each image, an element-wise comparison produces a more accurate judgement as to whether two pictures are of the same person.
<a name='3'></a>
## 3 - Encoding Face Images into a 128-Dimensional Vector
<a name='3-1'></a>
### 3.1 - Using a ConvNet to Compute Encodings
The FaceNet model takes a lot of data and a long time to train. So following the common practice in applied deep learning, you'll load weights that someone else has already trained. The network architecture follows the Inception model from [Szegedy *et al*..](https://arxiv.org/abs/1409.4842) An Inception network implementation has been provided for you, and you can find it in the file `inception_blocks_v2.py` to get a closer look at how it is implemented.
*Hot tip:* Go to "File->Open..." at the top of this notebook. This opens the file directory that contains the `.py` file).
The key things to be aware of are:
- This network uses 160x160 dimensional RGB images as its input. Specifically, a face image (or batch of $m$ face images) as a tensor of shape $(m, n_H, n_W, n_C) = (m, 160, 160, 3)$
- The input images are originally of shape 96x96, thus, you need to scale them to 160x160. This is done in the `img_to_encoding()` function.
- The output is a matrix of shape $(m, 128)$ that encodes each input face image into a 128-dimensional vector
Run the cell below to create the model for face images!
```
from tensorflow.keras.models import model_from_json
json_file = open('keras-facenet-h5/model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
model = model_from_json(loaded_model_json)
model.load_weights('keras-facenet-h5/model.h5')
```
Now summarize the input and output shapes:
```
print(model.inputs)
print(model.outputs)
```
By using a 128-neuron fully connected layer as its last layer, the model ensures that the output is an encoding vector of size 128. You then use the encodings to compare two face images as follows:
<img src="images/distance_kiank.png\" style="width:680px;height:250px;">
<caption><center> <u> <font color='purple'> <b>Figure 2:</b> <br> </u> <font color='purple'>By computing the distance between two encodings and thresholding, you can determine if the two pictures represent the same person</center></caption>
So, an encoding is a good one if:
- The encodings of two images of the same person are quite similar to each other.
- The encodings of two images of different persons are very different.
The triplet loss function formalizes this, and tries to "push" the encodings of two images of the same person (Anchor and Positive) closer together, while "pulling" the encodings of two images of different persons (Anchor, Negative) further apart.
<img src="images/triplet_comparison.png" style="width:280px;height:150px;"><br>
<caption><center> <u> <font color='purple'> <b>Figure 3: </b> <br> </u> <font color='purple'> In the next section, you'll call the pictures from left to right: Anchor (A), Positive (P), Negative (N)</center></caption>
<a name='3-2'></a>
### 3.2 - The Triplet Loss
**Important Note**: Since you're using a pretrained model, you won't actually need to implement the triplet loss function in this assignment. *However*, the triplet loss is the main ingredient of the face recognition algorithm, and you'll need to know how to use it for training your own FaceNet model, as well as other types of image similarity problems. Therefore, you'll implement it below, for fun and edification. :)
For an image $x$, its encoding is denoted as $f(x)$, where $f$ is the function computed by the neural network.
<img src="images/f_x.png" style="width:380px;height:150px;">
Training will use triplets of images $(A, P, N)$:
- A is an "Anchor" image--a picture of a person.
- P is a "Positive" image--a picture of the same person as the Anchor image.
- N is a "Negative" image--a picture of a different person than the Anchor image.
These triplets are picked from the training dataset. $(A^{(i)}, P^{(i)}, N^{(i)})$ is used here to denote the $i$-th training example.
You'd like to make sure that an image $A^{(i)}$ of an individual is closer to the Positive $P^{(i)}$ than to the Negative image $N^{(i)}$) by at least a margin $\alpha$:
$$
|| f\left(A^{(i)}\right)-f\left(P^{(i)}\right)||_{2}^{2}+\alpha<|| f\left(A^{(i)}\right)-f\left(N^{(i)}\right)||_{2}^{2}
$$
You would thus like to minimize the following "triplet cost":
$$\mathcal{J} = \sum^{m}_{i=1} \large[ \small \underbrace{\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2}_\text{(1)} - \underbrace{\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2}_\text{(2)} + \alpha \large ] \small_+ \tag{3}$$
Here, the notation "$[z]_+$" is used to denote $max(z,0)$.
**Notes**:
- The term (1) is the squared distance between the anchor "A" and the positive "P" for a given triplet; you want this to be small.
- The term (2) is the squared distance between the anchor "A" and the negative "N" for a given triplet, you want this to be relatively large. It has a minus sign preceding it because minimizing the negative of the term is the same as maximizing that term.
- $\alpha$ is called the margin. It's a hyperparameter that you pick manually. You'll use $\alpha = 0.2$.
Most implementations also rescale the encoding vectors to haven L2 norm equal to one (i.e., $\mid \mid f(img)\mid \mid_2$=1); you won't have to worry about that in this assignment.
<a name='ex-1'></a>
### Exercise 1 - triplet_loss
Implement the triplet loss as defined by formula (3). These are the 4 steps:
1. Compute the distance between the encodings of "anchor" and "positive": $\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2$
2. Compute the distance between the encodings of "anchor" and "negative": $\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2$
3. Compute the formula per training example: $ \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2 - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2 + \alpha$
4. Compute the full formula by taking the max with zero and summing over the training examples:$$\mathcal{J} = \sum^{m}_{i=1} \large[ \small \mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2 - \mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2+ \alpha \large ] \small_+ \tag{3}$$
*Hints*:
- Useful functions: `tf.reduce_sum()`, `tf.square()`, `tf.subtract()`, `tf.add()`, `tf.maximum()`.
- For steps 1 and 2, sum over the entries of $\mid \mid f(A^{(i)}) - f(P^{(i)}) \mid \mid_2^2$ and $\mid \mid f(A^{(i)}) - f(N^{(i)}) \mid \mid_2^2$.
- For step 4, you will sum over the training examples.
*Additional Hints*:
- Recall that the square of the L2 norm is the sum of the squared differences: $||x - y||_{2}^{2} = \sum_{i=1}^{N}(x_{i} - y_{i})^{2}$
- Note that the anchor, positive and negative encodings are of shape (*m*,128), where *m* is the number of training examples and 128 is the number of elements used to encode a single example.
- For steps 1 and 2, maintain the number of *m* training examples and sum along the 128 values of each encoding. `tf.reduce_sum` has an axis parameter. This chooses along which axis the sums are applied.
- Note that one way to choose the last axis in a tensor is to use negative indexing (axis=-1).
- In step 4, when summing over training examples, the result will be a single scalar value.
- For `tf.reduce_sum` to sum across all axes, keep the default value axis=None.
```
# UNQ_C1(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: triplet_loss
def triplet_loss(y_true, y_pred, alpha = 0.2):
"""
Implementation of the triplet loss as defined by formula (3)
Arguments:
y_true -- true labels, required when you define a loss in Keras, you don't need it in this function.
y_pred -- python list containing three objects:
anchor -- the encodings for the anchor images, of shape (None, 128)
positive -- the encodings for the positive images, of shape (None, 128)
negative -- the encodings for the negative images, of shape (None, 128)
Returns:
loss -- real number, value of the loss
"""
anchor, positive, negative = y_pred[0], y_pred[1], y_pred[2]
### START CODE HERE
#(≈ 4 lines)
# Step 1: Compute the (encoding) distance between the anchor and the positive
pos_dist = tf.reduce_sum(tf.square(tf.subtract(anchor, positive)), axis = -1)
# Step 2: Compute the (encoding) distance between the anchor and the negative
neg_dist = tf.reduce_sum(tf.square(tf.subtract(anchor, negative)), axis = -1)
# Step 3: subtract the two previous distances and add alpha.
basic_loss = tf.add(tf.subtract(pos_dist, neg_dist), alpha)
# Step 4: Take the maximum of basic_loss and 0.0. Sum over the training examples.
loss = tf.reduce_sum(tf.maximum(basic_loss, 0))
### END CODE HERE
return loss
# BEGIN UNIT TEST
tf.random.set_seed(1)
y_true = (None, None, None) # It is not used
y_pred = (tf.keras.backend.random_normal([3, 128], mean=6, stddev=0.1, seed = 1),
tf.keras.backend.random_normal([3, 128], mean=1, stddev=1, seed = 1),
tf.keras.backend.random_normal([3, 128], mean=3, stddev=4, seed = 1))
loss = triplet_loss(y_true, y_pred)
assert type(loss) == tf.python.framework.ops.EagerTensor, "Use tensorflow functions"
print("loss = " + str(loss))
y_pred_perfect = ([1., 1.], [1., 1.], [1., 1.,])
loss = triplet_loss(y_true, y_pred_perfect, 5)
assert loss == 5, "Wrong value. Did you add the alpha to basic_loss?"
y_pred_perfect = ([1., 1.],[1., 1.], [0., 0.,])
loss = triplet_loss(y_true, y_pred_perfect, 3)
assert loss == 1., "Wrong value. Check that pos_dist = 0 and neg_dist = 2 in this example"
y_pred_perfect = ([1., 1.],[0., 0.], [1., 1.,])
loss = triplet_loss(y_true, y_pred_perfect, 0)
assert loss == 2., "Wrong value. Check that pos_dist = 2 and neg_dist = 0 in this example"
y_pred_perfect = ([0., 0.],[0., 0.], [0., 0.,])
loss = triplet_loss(y_true, y_pred_perfect, -2)
assert loss == 0, "Wrong value. Are you taking the maximum between basic_loss and 0?"
y_pred_perfect = ([[1., 0.], [1., 0.]],[[1., 0.], [1., 0.]], [[0., 1.], [0., 1.]])
loss = triplet_loss(y_true, y_pred_perfect, 3)
assert loss == 2., "Wrong value. Are you applying tf.reduce_sum to get the loss?"
y_pred_perfect = ([[1., 1.], [2., 0.]], [[0., 3.], [1., 1.]], [[1., 0.], [0., 1.,]])
loss = triplet_loss(y_true, y_pred_perfect, 1)
if (loss == 4.):
raise Exception('Perhaps you are not using axis=-1 in reduce_sum?')
assert loss == 5, "Wrong value. Check your implementation"
# END UNIT TEST
```
**Expected Output**:
<table>
<tr>
<td>
<b>loss</b>
</td>
<td>
527.2598
</td>
</tr>
</table>
<a name='4'></a>
## 4 - Loading the Pre-trained Model
FaceNet is trained by minimizing the triplet loss. But since training requires a lot of data and a lot of computation, you won't train it from scratch here. Instead, you'll load a previously trained model in the following cell; which might take a couple of minutes to run.
```
FRmodel = model
```
Here are some examples of distances between the encodings between three individuals:
<img src="images/distance_matrix.png" style="width:380px;height:200px;"><br>
<caption><center> <u> <font color='purple'> <b>Figure 4:</b></u> <br> <font color='purple'> Example of distance outputs between three individuals' encodings</center></caption>
Now use this model to perform face verification and face recognition!
<a name='5'></a>
## 5 - Applying the Model
You're building a system for an office building where the building manager would like to offer facial recognition to allow the employees to enter the building.
You'd like to build a face verification system that gives access to a list of people. To be admitted, each person has to swipe an identification card at the entrance. The face recognition system then verifies that they are who they claim to be.
<a name='5-1'></a>
### 5.1 - Face Verification
Now you'll build a database containing one encoding vector for each person who is allowed to enter the office. To generate the encoding, you'll use `img_to_encoding(image_path, model)`, which runs the forward propagation of the model on the specified image.
Run the following code to build the database (represented as a Python dictionary). This database maps each person's name to a 128-dimensional encoding of their face.
```
#tf.keras.backend.set_image_data_format('channels_last')
def img_to_encoding(image_path, model):
img = tf.keras.preprocessing.image.load_img(image_path, target_size=(160, 160))
img = np.around(np.array(img) / 255.0, decimals=12)
x_train = np.expand_dims(img, axis=0)
embedding = model.predict_on_batch(x_train)
return embedding / np.linalg.norm(embedding, ord=2)
database = {}
database["danielle"] = img_to_encoding("images/danielle.png", FRmodel)
database["younes"] = img_to_encoding("images/younes.jpg", FRmodel)
database["tian"] = img_to_encoding("images/tian.jpg", FRmodel)
database["andrew"] = img_to_encoding("images/andrew.jpg", FRmodel)
database["kian"] = img_to_encoding("images/kian.jpg", FRmodel)
database["dan"] = img_to_encoding("images/dan.jpg", FRmodel)
database["sebastiano"] = img_to_encoding("images/sebastiano.jpg", FRmodel)
database["bertrand"] = img_to_encoding("images/bertrand.jpg", FRmodel)
database["kevin"] = img_to_encoding("images/kevin.jpg", FRmodel)
database["felix"] = img_to_encoding("images/felix.jpg", FRmodel)
database["benoit"] = img_to_encoding("images/benoit.jpg", FRmodel)
database["arnaud"] = img_to_encoding("images/arnaud.jpg", FRmodel)
```
Load the images of Danielle and Kian:
```
danielle = tf.keras.preprocessing.image.load_img("images/danielle.png", target_size=(160, 160))
kian = tf.keras.preprocessing.image.load_img("images/kian.jpg", target_size=(160, 160))
np.around(np.array(kian) / 255.0, decimals=12).shape
kian
np.around(np.array(danielle) / 255.0, decimals=12).shape
danielle
```
Now, when someone shows up at your front door and swipes their ID card (thus giving you their name), you can look up their encoding in the database, and use it to check if the person standing at the front door matches the name on the ID.
<a name='ex-2'></a>
### Exercise 2 - verify
Implement the `verify()` function, which checks if the front-door camera picture (`image_path`) is actually the person called "identity". You will have to go through the following steps:
- Compute the encoding of the image from `image_path`.
- Compute the distance between this encoding and the encoding of the identity image stored in the database.
- Open the door if the distance is less than 0.7, else do not open it.
As presented above, you should use the L2 distance `np.linalg.norm`.
**Note**: In this implementation, compare the L2 distance, not the square of the L2 distance, to the threshold 0.7.
*Hints*:
- `identity` is a string that is also a key in the database dictionary.
- `img_to_encoding` has two parameters: the image_path and model.
```
# UNQ_C2(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: verify
def verify(image_path, identity, database, model):
"""
Function that verifies if the person on the "image_path" image is "identity".
Arguments:
image_path -- path to an image
identity -- string, name of the person you'd like to verify the identity. Has to be an employee who works in the office.
database -- python dictionary mapping names of allowed people's names (strings) to their encodings (vectors).
model -- your Inception model instance in Keras
Returns:
dist -- distance between the image_path and the image of "identity" in the database.
door_open -- True, if the door should open. False otherwise.
"""
### START CODE HERE
# Step 1: Compute the encoding for the image. Use img_to_encoding() see example above. (≈ 1 line)
encoding = img_to_encoding(image_path, model)
# Step 2: Compute distance with identity's image (≈ 1 line)
dist = np.linalg.norm(encoding - database[identity])
# Step 3: Open the door if dist < 0.7, else don't open (≈ 3 lines)
if dist < 0.7:
print("It's " + str(identity) + ", welcome in!")
door_open = True
else:
print("It's not " + str(identity) + ", please go away")
door_open = False
### END CODE HERE
return dist, door_open
```
Younes is trying to enter the office and the camera takes a picture of him ("images/camera_0.jpg"). Let's run your verification algorithm on this picture:
<img src="images/camera_0.jpg\" style="width:100px;height:100px;">
```
# BEGIN UNIT TEST
assert(np.allclose(verify("images/camera_1.jpg", "bertrand", database, FRmodel), (0.54364836, True)))
assert(np.allclose(verify("images/camera_3.jpg", "bertrand", database, FRmodel), (0.38616243, True)))
assert(np.allclose(verify("images/camera_1.jpg", "younes", database, FRmodel), (1.3963861, False)))
assert(np.allclose(verify("images/camera_3.jpg", "younes", database, FRmodel), (1.3872949, False)))
verify("images/camera_0.jpg", "younes", database, FRmodel)
# END UNIT TEST
```
**Expected Output**:
<table>
<tr>
<td>
<b>It's Younes, welcome in!</b>
</td>
<td>
(0.5992946, True)
</td>
</tr>
</table>
Benoit, who does not work in the office, stole Kian's ID card and tried to enter the office. Naughty Benoit! The camera took a picture of Benoit ("images/camera_2.jpg).
<img src="images/camera_2.jpg" style="width:100px;height:100px;">
Run the verification algorithm to check if Benoit can enter.
```
verify("images/camera_2.jpg", "kian", database, FRmodel)
```
**Expected Output**:
<table>
<tr>
<td>
<b>It's not Kian, please go away</b>
</td>
<td>
(1.0259346, False)
</td>
</tr>
</table>
<a name='5-2'></a>
### 5.2 - Face Recognition
Your face verification system is mostly working. But since Kian got his ID card stolen, when he came back to the office the next day he couldn't get in!
To solve this, you'd like to change your face verification system to a face recognition system. This way, no one has to carry an ID card anymore. An authorized person can just walk up to the building, and the door will unlock for them!
You'll implement a face recognition system that takes as input an image, and figures out if it is one of the authorized persons (and if so, who). Unlike the previous face verification system, you will no longer get a person's name as one of the inputs.
<a name='ex-3'></a>
### Exercise 3 - who_is_it
Implement `who_is_it()` with the following steps:
- Compute the target encoding of the image from `image_path`
- Find the encoding from the database that has smallest distance with the target encoding.
- Initialize the `min_dist` variable to a large enough number (100). This helps you keep track of the closest encoding to the input's encoding.
- Loop over the database dictionary's names and encodings. To loop use for (name, db_enc) in `database.items()`.
- Compute the L2 distance between the target "encoding" and the current "encoding" from the database. If this distance is less than the min_dist, then set min_dist to dist, and identity to name.
```
# UNQ_C3(UNIQUE CELL IDENTIFIER, DO NOT EDIT)
# GRADED FUNCTION: who_is_it
def who_is_it(image_path, database, model):
"""
Implements face recognition for the office by finding who is the person on the image_path image.
Arguments:
image_path -- path to an image
database -- database containing image encodings along with the name of the person on the image
model -- your Inception model instance in Keras
Returns:
min_dist -- the minimum distance between image_path encoding and the encodings from the database
identity -- string, the name prediction for the person on image_path
"""
### START CODE HERE
## Step 1: Compute the target "encoding" for the image. Use img_to_encoding() see example above. ## (≈ 1 line)
encoding = img_to_encoding(image_path, model)
## Step 2: Find the closest encoding ##
# Initialize "min_dist" to a large value, say 100 (≈1 line)
min_dist = 100
# Loop over the database dictionary's names and encodings.
for (name, db_enc) in database.items():
# Compute L2 distance between the target "encoding" and the current db_enc from the database. (≈ 1 line)
dist = np.linalg.norm(encoding - database[name])
# If this distance is less than the min_dist, then set min_dist to dist, and identity to name. (≈ 3 lines)
if dist < min_dist:
min_dist = dist
identity = name
### END CODE HERE
if min_dist > 0.7:
print("Not in the database.")
else:
print ("it's " + str(identity) + ", the distance is " + str(min_dist))
return min_dist, identity
```
Younes is at the front door and the camera takes a picture of him ("images/camera_0.jpg"). Let's see if your `who_it_is()` algorithm identifies Younes.
```
# BEGIN UNIT TEST
# Test 1 with Younes pictures
who_is_it("images/camera_0.jpg", database, FRmodel)
# Test 2 with Younes pictures
test1 = who_is_it("images/camera_0.jpg", database, FRmodel)
assert np.isclose(test1[0], 0.5992946)
assert test1[1] == 'younes'
# Test 3 with Younes pictures
test2 = who_is_it("images/younes.jpg", database, FRmodel)
assert np.isclose(test2[0], 0.0)
assert test2[1] == 'younes'
# END UNIT TEST
```
**Expected Output**:
<table>
<tr>
<td>
<b>it's Younes, the distance is 0.5992946</b>
</td>
<td>
(0.5992946, 'younes')
</td>
</tr>
</table>
You can change "camera_0.jpg" (picture of Younes) to "camera_1.jpg" (picture of Bertrand) and see the result.
**Congratulations**!
You've completed this assignment, and your face recognition system is working well! It not only lets in authorized persons, but now people don't need to carry an ID card around anymore!
You've now seen how a state-of-the-art face recognition system works, and can describe the difference between face recognition and face verification. Here's a quick recap of what you've accomplished:
- Posed face recognition as a binary classification problem
- Implemented one-shot learning for a face recognition problem
- Applied the triplet loss function to learn a network's parameters in the context of face recognition
- Mapped face images into 128-dimensional encodings using a pretrained model
- Performed face verification and face recognition with these encodings
Great work!
<font color='blue'>
**What you should remember**:
- Face verification solves an easier 1:1 matching problem; face recognition addresses a harder 1:K matching problem.
- Triplet loss is an effective loss function for training a neural network to learn an encoding of a face image.
- The same encoding can be used for verification and recognition. Measuring distances between two images' encodings allows you to determine whether they are pictures of the same person.
**Ways to improve your facial recognition model**:
Although you won't implement these here, here are some ways to further improve the algorithm:
- Put more images of each person (under different lighting conditions, taken on different days, etc.) into the database. Then, given a new image, compare the new face to multiple pictures of the person. This would increase accuracy.
- Crop the images to contain just the face, and less of the "border" region around the face. This preprocessing removes some of the irrelevant pixels around the face, and also makes the algorithm more robust.
<a name='6'></a>
## 6 - References
1. Florian Schroff, Dmitry Kalenichenko, James Philbin (2015). [FaceNet: A Unified Embedding for Face Recognition and Clustering](https://arxiv.org/pdf/1503.03832.pdf)
2. Yaniv Taigman, Ming Yang, Marc'Aurelio Ranzato, Lior Wolf (2014). [DeepFace: Closing the gap to human-level performance in face verification](https://research.fb.com/wp-content/uploads/2016/11/deepface-closing-the-gap-to-human-level-performance-in-face-verification.pdf)
3. This implementation also took a lot of inspiration from the official FaceNet github repository: https://github.com/davidsandberg/facenet
4. Further inspiration was found here: https://machinelearningmastery.com/how-to-develop-a-face-recognition-system-using-facenet-in-keras-and-an-svm-classifier/
5. And here: https://github.com/nyoki-mtl/keras-facenet/blob/master/notebook/tf_to_keras.ipynb
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
import cv2
from tqdm import tqdm
from ssd_model import DSOD300
from ssd_data import InputGenerator
from ssd_utils import PriorUtil
from ssd_metric import confusion_matrix, plot_confusion_matrix, accuracy, evaluate_results
from utils.model import load_weights
from data_voc import GTUtility
gt_util = GTUtility('./data/VOC2007test/')
from data_coco import GTUtility
gt_util = GTUtility('./data/COCO/', validation=True)
gt_util = gt_util.convert_to_voc()
model = DSOD300(input_shape=(300, 300, 3), num_classes=gt_util.num_classes); confidence_threshold = 0.275
#model = DSOD300(input_shape=(600, 600, 3), num_classes=gt_util.num_classes); confidence_threshold = 0.35
load_weights(model, './checkpoints/201808261035_dsod300fl_voccoco/weights.110.h5')
prior_util = PriorUtil(model)
inputs, data = gt_util.sample_batch(min(gt_util.num_samples, 10000), 0, input_size=model.image_size)
#inputs, data = gt_util.sample_batch(min(gt_util.num_samples, 10000), 0, input_size=model.image_size, preserve_aspect_ratio=True)
#gen = InputGenerator(gt_util, prior_util, min(gt_util.num_samples, 10000), model.image_size, augmentation=True)
#inputs, data = next(gen.generate(encode=False))
preds = model.predict(inputs, batch_size=1, verbose=1)
```
### Grid search
```
steps = np.arange(0.05, 1, 0.05)
fmes_grid = np.zeros((len(steps)))
for i, t in enumerate(steps):
results = [prior_util.decode(p, t) for p in preds]
fmes = evaluate_results(data, results, gt_util, iou_thresh=0.5, max_dets=100, return_fmeasure=True)
fmes_grid[i] = fmes
print('threshold %.2f f-measure %.2f' % (t, fmes))
max_idx = np.argmax(fmes_grid)
print(steps[max_idx], fmes_grid[max_idx])
plt.figure(figsize=[12,6])
plt.plot(steps, fmes_grid)
plt.plot(steps[max_idx], fmes_grid[max_idx], 'or')
plt.xticks(steps)
plt.grid()
plt.xlabel('threshold')
plt.ylabel('f-measure')
plt.show()
confidence_threshold = 0.275 # pascal voc 2007 test
#confidence_threshold = 0.550 # pascal voc 2012 test
#confidence_threshold = 0.125 # ms coco test
```
### Precision-recall curve, mean Average Precision
```
results = [prior_util.decode(p, confidence_threshold=0.01, keep_top_k=400) for p in tqdm(preds)]
evaluate_results(data, results, gt_util, iou_thresh=0.5, max_dets=100)
```
### Confusion matrix of local predictions
```
results = [prior_util.decode(p, confidence_threshold) for p in tqdm(preds)]
encoded_gt = [prior_util.encode(d) for d in tqdm(data)]
y_true_all = []
y_pred_all = []
for i in range(len(data)):
y_true = np.argmax(encoded_gt[i][:,4:], axis=1)
y_pred = np.argmax(preds[i][:,4:], axis=1)
#prior_object_idx = np.where(y_true)[0] # gt prior box contains object
prior_object_idx = np.where(y_true+y_pred)[0] # gt or prediction prior box contains object
y_true_all.extend(y_true[prior_object_idx])
y_pred_all.extend(y_pred[prior_object_idx])
#y_pred_all = [ 17 for i in y_pred_all]
cm = confusion_matrix(y_true_all, y_pred_all, gt_util.num_classes)
plot_confusion_matrix(cm, gt_util.classes, figsize=[12]*2)
accuracy(y_true_all, y_pred_all)
```
### Examples
```
for i in np.random.randint(0, len(inputs), 30):
plt.figure(figsize=[8]*2)
gt_util.plot_input(inputs[i])
#gt_util.plot_gt(data[i])
prior_util.plot_results(results=results[i], classes=gt_util.classes, gt_data=data[i],
confidence_threshold=confidence_threshold)
plt.axis('off')
plt.show()
```
| github_jupyter |
```
import pandas as pd
from pathlib import Path
from tqdm import tqdm
import tensorflow as tf
# The following functions can be used to convert a value to a type compatible
# with tf.Example.
def _bytes_feature(value):
"""Returns a bytes_list from a string / byte."""
if isinstance(value, type(tf.constant(0))):
value = value.numpy() # BytesList won't unpack a string from an EagerTensor.
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[value]))
def _float_feature(value):
"""Returns a float_list from a float / double."""
return tf.train.Feature(float_list=tf.train.FloatList(value=[value]))
def _int64_feature(value):
"""Returns an int64_list from a bool / enum / int / uint."""
return tf.train.Feature(int64_list=tf.train.Int64List(value=[value]))
def serialize(text, label):
feature = {
"text": _bytes_feature(text),
"label": _int64_feature(label)
}
example_proto = tf.train.Example(features=tf.train.Features(feature=feature))
return example_proto.SerializeToString()
def tf_serialize_example(text, label):
tf_string = tf.py_function(
serialize,
(text, label), # pass these args to the above function.
tf.string) # the return type is `tf.string`.
return tf.reshape(tf_string, ())
```
# Zeerak hatespeech
```
df = pd.read_csv("data/hatespeech/NAACL_SRW_2016.csv")
df["racism"]
```
# Supremacist
```
comments = dict()
for p in tqdm(Path("data/hate-speech-dataset/all_files/").glob("*.txt")):
with p.open("r") as f:
t = f.read()
comments[p.stem] = t
df = pd.read_csv("data/hate-speech-dataset/annotations_metadata.csv")
df
df_train = df[:8000]
df_val = df[8000:]
def iter_comments(df):
def iterator():
for i, row in df.iterrows():
comment = comments[row.file_id]
label = int(row.label == "hate")
yield (comment, label)
return iterator
ds_train = tf.data.Dataset.from_generator(iter_comments(df_train), output_types=(tf.string, tf.int32))
ds_val = tf.data.Dataset.from_generator(iter_comments(df_val), output_types=(tf.string, tf.int32))
writer_train = tf.data.experimental.TFRecordWriter("supremacists_train.tfrecord")
writer_train.write(ds_train.map(tf_serialize_example))
writer_val = tf.data.experimental.TFRecordWriter("supremacists_val.tfrecord")
writer_val.write(ds_val.map(tf_serialize_example))
import nlpaug.augmenter.char as nac
import nlpaug.augmenter.word as naw
import nlpaug.augmenter.sentence as nas
import nltk
nltk.download('averaged_perceptron_tagger')
for x, y in ds_val:
text = x.numpy().decode()
break
aug = naw.SynonymAug(aug_src='wordnet')
augmented_text = aug.augment(text)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
aug = naw.AntonymAug()
augmented_text = aug.augment(text,)
print("Original:")
print(text)
print("Augmented Text:")
print(augmented_text)
def augment(x):
x = x.numpy().decode()
x = aug.augment(x)
return x
for x in ds_train.map(
lambda x,y: (tf.py_function(augment, [x], tf.string), y)
):
print(x)
break
```
| github_jupyter |
# Palindrome Program
> For any value (number, string, etc.)
```
# function which return reverse of a string
def reverse(s):
return s[::-1]
def isPalindrome(s):
# Calling reverse function
rev = reverse(s)
# Checking if both string are equal or not
if (s == rev):
return True
return False
# Driver code
s = input("Enter something: ")
ans = isPalindrome(s)
if ans == 1:
print("Yes")
else:
print("No")
```
# Factorial
> Only for integers.
```
def fact(n):
fact = 1
for i in range(2, n+1):
fact *= i
return fact
try:
x = int(input("Enter a number: "))
print(fact(x))
except:
print("Some Error.")
import numpy as np
a = np.array([1, 2, 3]) # Create a rank 1 array
print(type(a)) # Prints "<class 'numpy.ndarray'>"
print(a.shape) # Prints "(3,)"
print(a[0], a[1], a[2]) # Prints "1 2 3"
a[0] = 5 # Change an element of the array
print(a) # Prints "[5, 2, 3]"
b = np.array([[1,2,3],[4,5,6]]) # Create a rank 2 array
print(b.shape) # Prints "(2, 3)"
print(b[0, 0], b[0, 1], b[1, 0]) # Prints "1 2 4"
import numpy as np
a = np.zeros((2,2)) # Create an array of all zeros
print(a) # Prints "[[ 0. 0.]
# [ 0. 0.]]"
b = np.ones((1,2)) # Create an array of all ones
print(b) # Prints "[[ 1. 1.]]"
c = np.full((2,2), 7) # Create a constant array
print(c) # Prints "[[ 7. 7.]
# [ 7. 7.]]"
d = np.eye(2) # Create a 2x2 identity matrix
print(d) # Prints "[[ 1. 0.]
# [ 0. 1.]]"
e = np.random.random((2,2)) # Create an array filled with random values
print(e) # Might print "[[ 0.91940167 0.08143941]
# [ 0.68744134 0.87236687]]"
```
# .csv File Ops
> - Input matrix from user and enter into a .csv file
> - Print contents of .csv file in matrix form
```
def writeCsv(data):
try:
filename = input("Enter filename: ")
fh = open(filename, 'w')
with myFile:
writer = csv.writer(fh)
writer.writerows(data)
print("Writing complete")
except:
print("Some Error.")
import csv
import numpy
def readCsv():
# try:
# filename = input("Enter filename")
filename = '2.csv'
with open(filename, 'r') as csvFile:
reader = csv.reader(csvFile)
lines = np.array(list(reader))
r,c = lines.shape
h = np.zeros(c,)
if lines[0,4] == 'no':
lines[0] = h
lines[0,4] = 'no'
if lines[0,4] == 'yes':
h = lines[0]
for row in range(1,r):
if lines[row, 4] == 'no':
lines[row] = h
if lines[row, 4] == 'yes':
for i in range(len(lines[row])):
if h[i] is not lines[row, i]:
h[i] = str('?')
h = lines[row]
for row in lines:
print(row[0:4])
csvFile.close()
# except:
# print("Some Error.")
readCsv()
```
# Nearest Palindrome
- Input: N (0 <= N <= 10^14)
- Output: The nearest palindrome to the given N. If N is a palindrome, then output N.
- If the difference between the two nearest palindrome is the closest, output is the lower value.
```
# n = input("Enter a num")
# n = '123456678'
n = '1070'
l = len(n)
h1, h2 = n[:l//2], n[(l+1)//2:]
h3 = n[:(l+1)//2]
n1 = str(h1[:-1])+h1[::-1]
n1 = int(n1)
n2 = str(int(h1) - 1)
n2 += h1[::-1][1:]
print(h1,'\t',h2, '\t', n1, '\t', n2)
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%aimport utils_1_1
import pandas as pd
import numpy as np
import altair as alt
from altair_saver import save
import datetime
import dateutil.parser
from os.path import join
from constants_1_1 import SITE_FILE_TYPES
from utils_1_1 import (
get_site_file_paths,
get_site_file_info,
get_site_ids,
get_visualization_subtitle,
get_country_color_map,
)
from theme import apply_theme
from web import for_website
alt.data_transformers.disable_max_rows(); # Allow using rows more than 5000
LOG_LABS = ['alanine aminotransferase (ALT)', 'aspartate aminotransferase (AST)', 'C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer', 'Ferritin', 'lactate dehydrogenase (LDH)']
consistent_loinc = {
"C-reactive protein (CRP) (Normal Sensitivity)": "C-reactive protein (Normal Sensitivity) (mg/dL)",
"creatinine": "Creatinine (mg/dL)",
"Ferritin": "Ferritin (ng/mL)",
"D-dimer": "D-dimer (ng/mL)",
"albumin": "Albumin (g/dL)",
"Fibrinogen": "Fibrinogen (mg/dL)",
"alanine aminotransferase (ALT)": "Alanine aminotransferase (U/L)",
"aspartate aminotransferase (AST)": "Aspartate aminotransferase (U/L)",
"total bilirubin": "Total bilirubin (mg/dL)",
"lactate dehydrogenase (LDH)": "Lactate dehydrogenase (U/L)",
"cardiac troponin": "Cardiac troponin (ng/mL)",
"cardiac troponin (High Sensitivity)": "Cardiac Troponin (High Sensitivity) (ng/mL)",
"cardiac troponin (Normal Sensitivity)": "Cardiac Troponin (Normal Sensitivity) (ng/mL)",
"prothrombin time (PT)": "Prothrombin time (s)",
"white blood cell count (Leukocytes)": "White blood cell count (10*3/uL)",
"lymphocyte count": "Lymphocyte count (10*3/uL)",
"neutrophil count": "Neutrophil count (10*3/uL)",
"procalcitonin": "Procalcitonin (ng/mL)",
}
# Let's remove units
consistent_loinc = {
"C-reactive protein (CRP) (Normal Sensitivity)": "C-reactive protein (Normal Sensitivity)",
"creatinine": "Creatinine",
"Ferritin": "Ferritin",
"D-dimer": "D-dimer",
"albumin": "Albumin",
"Fibrinogen": "Fibrinogen",
"alanine aminotransferase (ALT)": "Alanine aminotransferase",
"aspartate aminotransferase (AST)": "Aspartate aminotransferase",
"total bilirubin": "Total bilirubin",
"lactate dehydrogenase (LDH)": "Lactate dehydrogenase",
"cardiac troponin": "Cardiac troponin",
"cardiac troponin (High Sensitivity)": "Cardiac Troponin (High Sensitivity)",
"cardiac troponin (Normal Sensitivity)": "Cardiac Troponin (Normal Sensitivity)",
"prothrombin time (PT)": "Prothrombin time",
"white blood cell count (Leukocytes)": "White blood cell count",
"lymphocyte count": "Lymphocyte count",
"neutrophil count": "Neutrophil count",
"procalcitonin": "Procalcitonin",
}
df = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_values_standardized.csv"))
df
obs = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_obs_for_Sehi.csv"))
# obs.setting.unique().tolist()
obs
# obs = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_obs.csv"))
# obs
pdf = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals.csv"))
pdf
domain_by_lab = {
'C-reactive protein (CRP) (Normal Sensitivity)': [0.85, 1.15],
'albumin': [0.75, 1.05],
'total bilirubin': [0.95, 1.30],
'creatinine': [0.85, 1.15],
'Ferritin': [0.95, 1.07],
'D-dimer': [0.97, 1.14]
}
domain_by_lab = {
'C-reactive protein (CRP) (Normal Sensitivity)': [0.85, 1.15],
'Ferritin': [0.97, 1.08],
'Fibrinogen': [0.88, 1.05],
'procalcitonin': [0.9, 3],
'D-dimer': [0.96, 1.15],
'creatinine': [0.85, 1.15]
}
def plot(_d, patient_group='all', lab=None, obs=None, pdf=None, i=0, show_patients=False):
d = _d.copy()
o = obs.copy()
p = pdf.copy()
"""
DATA PREPROCESSING...
"""
if lab in LOG_LABS:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d = d.drop(columns=['Unnamed: 0'])
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.Lab == lab]
d17 = d[(1 <= d.days_since_positive) & (d.days_since_positive <= 7)]
d = d[(d.days_since_positive == 0) | (d.days_since_positive == 1) | (d.days_since_positive == 7)]
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d17.days_since_positive = d17.days_since_positive.apply(lambda x: f"Day {x}")
#### PVAL ###############################
p = p[p.setting == patient_group]
p['is_sig'] = False
p.is_sig = p[lab] <= 0.05
p.is_sig = p.is_sig.apply(lambda x: 'p<0.05' if x else 'p>0.05')
p = p[['setting','day', 'is_sig']]
p.day = p.day.apply(lambda x: f"Day {x}")
"""
MERGE
"""
d = pd.merge(d, p, how='left', left_on=['days_since_positive','setting'], right_on = ['day','setting'])
#### OBS ################################
o = o.rename(columns={"p.lwr": "p_lwr", "p.upr": "p_upr"})
o = o[o.lab == lab]
o = o.drop(columns=['Unnamed: 0'])
o.wave = o.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
o = o[o.cohort == 'dayX']
o = o[o.setting == patient_group]
o = o.rename(columns={
'wave': 'Wave'
})
"""
DAY 1-7 AVERAGE
"""
d17['mean'] = d17['mean'] * d17['total_n']
d17 = d17.groupby(['Lab', 'setting', 'Wave']).sum().reset_index()
d17['mean'] = d17['mean'] / d17['total_n']
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
"""
PLOT
"""
titleX=-60
opacity=0.7
"""
LABS
"""
LAB_DROPDOWN = alt.binding_select(options=LABS)
LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": lab if lab != None else LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=4, opacity=opacity, point=False
).encode(
x=alt.X('days_since_positive:N', title=None, axis=alt.Axis(labels=False) if (patient_group == 'all') & (show_patients == False) else alt.Axis()), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False, nice=False, padding=0, domain=domain_by_lab[lab]), title=['Mean Lab Value'] if i == 0 else None, axis=alt.Axis(titleX=titleX)), #domain=domain_by_lab[lab]
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR))
).properties(
width=200,
height=200
)
point_m = alt.Chart(
d
).mark_point(
opacity=opacity, filled=True, strokeWidth=3
).encode(
x=alt.X('days_since_positive:N', title=None), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=['Mean Lab Value'] if i == 0 else None, axis=alt.Axis(titleX=titleX)), # 'All Patients' if patient_group == 'all' else "Ever Severe Patients",
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR)),
stroke=alt.Stroke('is_sig:N', scale=alt.Scale(domain=['p<0.05'], range=['black']), title='Significance'),
size=alt.Size('total_n:Q', title="# of Patients", scale=alt.Scale(domain=[0, 30000], range=[100, 600], zero=False)),
strokeWidth=alt.value(3)
)
line_m = (line_m + point_m)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
"""
Day 1-7 Average
"""
bar_m = alt.Chart(
d17
).mark_bar(
size=32,
stroke='black'
).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value', axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=10, nice=False)),
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR))
).properties(
width=100
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
text = alt.Chart(
d17
).mark_text(size=16, dx=0, dy=-4, color='black', baseline='bottom', align='center', angle=0, fontWeight=500).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value' if i == 0 else None, axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=10, nice=False)),
text=alt.Text('mean:Q', format=".2f")
)
# .transform_filter(
# LAB_SELECTION
# )
bar_m = (bar_m)# + text)
"""
OBSERVATION
"""
LAB_FIELD_NAME = lab.replace(' ', '.').replace('(', '.').replace(')', '.').replace('-', '.') # Used in the original files
o = o.rename(columns={
f"{LAB_FIELD_NAME}": LAB_FIELD_NAME.replace('.', '_')
})
LAB_FIELD_NAME = LAB_FIELD_NAME.replace('.', '_')
obs_line = alt.Chart(
o
).mark_line(
point=True, opacity=0.7, size=4
).encode(
x=alt.X('day:Q', title='Days Since Admission'),
y=alt.Y(f"p:Q", scale=alt.Scale(domain=[0, 1]), axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested"),
color=alt.Color("Wave:N")
)
obs_error = obs_line.mark_errorbar(opacity=0.6).encode(
x=alt.X('day:Q', title='Days Since Admission', scale=alt.Scale(padding=10, nice=False)),
y=alt.Y('ci_95L:Q', axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested" if i == 0 else None),
y2=alt.Y2('ci_95U:Q'),
color=alt.Color("Wave:N"),
strokeWidth=alt.value(1.5)
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
obs_line = alt.layer(obs_line, obs_error).properties(
# title={
# "text": "Percentage of Patients Tested",
# "dx": 80,
# # "fontSize": 16,
# # "color": "gray"
# },
width=200,
height=150
)
plot = (
line_m if show_patients == False else (
alt.vconcat(
# alt.hconcat(line_m, bar_m, spacing=20).resolve_scale(y='shared'),
line_m,
obs_line,
spacing=5
).resolve_scale(x='independent')
)
).properties(
title={
"text": consistent_loinc[lab].replace(' (Normal Sensitivity)', ''),
"anchor": 'middle',
'fontSize': 18,
'dx': (5 if i != 0 else 13) if show_patients == False else (20 if i != 0 else 40)
}
)
return plot
```
# Main Labs
```
SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'Fibrinogen', 'procalcitonin', 'D-dimer', 'creatinine']
show_patients = True
for i, lab in enumerate(SELECTED_LABS):
# DEBUG
# if i == 1:
# break
# new = plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i)
new = alt.vconcat(
plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
# plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')#, color='independent', size='independent', stroke='independent')
if i != 0:
res = alt.hconcat(res, new, spacing=10)
else:
res = new
# DEBUG
#if i == 0:
#break
res = res.properties(
title={
"text": [
f"Mean Standardized Lab Values Of All Patients By Wave"
],
"dx": 80,
# "subtitle": [
# get_visualization_subtitle(data_release='2021-05-06', with_num_sites=False)],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
# res = apply_theme(
# res,
# axis_y_title_font_size=16,
# title_anchor='start',
# legend_orient='right',
# point_size=40
# )
res
# obs
SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'Fibrinogen', 'procalcitonin', 'D-dimer', 'creatinine']
show_patients = True
for i, lab in enumerate(SELECTED_LABS):
# DEBUG
# if i == 1:
# break
# new = plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i)
new = alt.vconcat(
# plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')#, color='independent', size='independent', stroke='independent')
if i != 0:
res2 = alt.hconcat(res2, new, spacing=10)
else:
res2 = new
# DEBUG
#if i == 0:
#break
res2 = res2.properties(
title={
"text": [
f"Mean Standardized Lab Values Of Ever Severe Patients By Wave"
],
"dx": 80,
# "titlePadding": 30,
# "subtitle": [
# get_visualization_subtitle(data_release='2021-05-06', with_num_sites=False)],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
# res = apply_theme(
# res,
# axis_y_title_font_size=16,
# title_anchor='start',
# legend_orient='right'
# )
res = alt.vconcat(res, res2, spacing=50).resolve_scale(size='shared')
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='bottom',
point_size=30
)
res
```
# Different Arrangement (All vs Severe)
```
domain_by_lab = {
'C-reactive protein (CRP) (Normal Sensitivity)': [0.85, 1.15],
'albumin': [0.75, 1.05],
'total bilirubin': [0.95, 1.30],
'creatinine': [0.85, 1.15],
'Ferritin': [0.95, 1.07],
'D-dimer': [0.97, 1.14]
}
domain_by_lab = {
'C-reactive protein (CRP) (Normal Sensitivity)': [0.85, 1.15],
'Ferritin': [0.97, 1.08],
'Fibrinogen': [0.88, 1.05],
'procalcitonin': [0.9, 3],
'D-dimer': [0.96, 1.15],
'creatinine': [0.85, 1.15]
}
def plot(_d, patient_group='all', lab=None, obs=None, pdf=None, i=0, show_patients=False):
d = _d.copy()
o = obs.copy()
p = pdf.copy()
"""
DATA PREPROCESSING...
"""
if lab in LOG_LABS:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d = d.drop(columns=['Unnamed: 0'])
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.Lab == lab]
d17 = d[(1 <= d.days_since_positive) & (d.days_since_positive <= 7)]
d = d[(d.days_since_positive == 0) | (d.days_since_positive == 1) | (d.days_since_positive == 7)]
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d17.days_since_positive = d17.days_since_positive.apply(lambda x: f"Day {x}")
#### PVAL ###############################
p = p[p.setting == patient_group]
p['is_sig'] = False
p.is_sig = p[lab] <= 0.05
p.is_sig = p.is_sig.apply(lambda x: 'p<0.05' if x else 'p>0.05')
p = p[['setting','day', 'is_sig']]
p.day = p.day.apply(lambda x: f"Day {x}")
"""
MERGE
"""
d = pd.merge(d, p, how='left', left_on=['days_since_positive','setting'], right_on = ['day','setting'])
#### OBS ################################
o = o.rename(columns={"p.lwr": "p_lwr", "p.upr": "p_upr"})
o = o[o.lab == lab]
o = o.drop(columns=['Unnamed: 0'])
o.wave = o.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
o = o[o.cohort == 'dayX']
o = o[o.setting == patient_group]
o = o.rename(columns={
'wave': 'Wave'
})
"""
DAY 1-7 AVERAGE
"""
d17['mean'] = d17['mean'] * d17['total_n']
d17 = d17.groupby(['Lab', 'setting', 'Wave']).sum().reset_index()
d17['mean'] = d17['mean'] / d17['total_n']
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
"""
PLOT
"""
titleX=-50
opacity=0.7
"""
LABS
"""
LAB_DROPDOWN = alt.binding_select(options=LABS)
LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": lab if lab != None else LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=4, opacity=opacity, point=False
).encode(
x=alt.X('days_since_positive:N', title=None if patient_group == 'all' else 'Days Since Admission', axis=alt.Axis(labels=False) if (patient_group == 'all') & (show_patients == False) else alt.Axis()), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False, nice=False, padding=0, domain=domain_by_lab[lab]), title=['All Patients' if patient_group == 'all' else 'Ever-Severe Patients'] if i == 0 else None, axis=alt.Axis(titleX=titleX)), #domain=domain_by_lab[lab] Mean Lab Value
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR))
).properties(
width=200,
height=200
)
point_m = alt.Chart(
d
).mark_point(
opacity=opacity, filled=True, strokeWidth=3
).encode(
x=alt.X('days_since_positive:N', title=None if patient_group == 'all' else 'Days Since Admission'), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=['All Patients' if patient_group == 'all' else 'Ever-Severe Patients'] if i == 0 else None), # 'All Patients' if patient_group == 'all' else "Ever Severe Patients", "Mean Lab Value"
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR)),
stroke=alt.Stroke('is_sig:N', scale=alt.Scale(domain=['P<0.05'], range=['black']), title='Significance'),
size=alt.Size('total_n:Q', title="# of Patients", scale=alt.Scale(domain=[0, 30000], range=[100, 600], zero=False)),
strokeWidth=alt.value(3)
)
line_m = (line_m + point_m)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
"""
Day 1-7 Average
"""
bar_m = alt.Chart(
d17
).mark_bar(
size=32,
stroke='black'
).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value', axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=10, nice=False)),
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR))
).properties(
width=100
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
text = alt.Chart(
d17
).mark_text(size=16, dx=0, dy=-4, color='black', baseline='bottom', align='center', angle=0, fontWeight=500).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value' if i == 0 else None, axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=10, nice=False)),
text=alt.Text('mean:Q', format=".2f")
)
# .transform_filter(
# LAB_SELECTION
# )
bar_m = (bar_m)# + text)
"""
OBSERVATION
"""
LAB_FIELD_NAME = lab.replace(' ', '.').replace('(', '.').replace(')', '.').replace('-', '.') # Used in the original files
o = o.rename(columns={
f"{LAB_FIELD_NAME}": LAB_FIELD_NAME.replace('.', '_')
})
LAB_FIELD_NAME = LAB_FIELD_NAME.replace('.', '_')
obs_line = alt.Chart(
o
).mark_line(
point=True, opacity=0.7, size=4
).encode(
x=alt.X('day:Q', title=None if patient_group == 'all' else 'Days Since Admission', axis=alt.Axis(labels=False) if (patient_group == 'all') else alt.Axis()),
y=alt.Y(f"p:Q", scale=alt.Scale(domain=[0, 1]), axis=alt.Axis(format='0.0%', titleX=titleX), title=['All Patients' if patient_group == 'all' else 'Ever-Severe Patients'] if i == 0 else None), # title="% Patients Tested" if i == 0 else None
color=alt.Color("Wave:N")
)
obs_error = obs_line.mark_errorbar(opacity=0.6).encode(
x=alt.X('day:Q', title=None if patient_group == 'all' else 'Days Since Admission', scale=alt.Scale(padding=10, nice=False), axis=alt.Axis(labels=False) if (patient_group == 'all') else alt.Axis()),
y=alt.Y('ci_95L:Q', axis=alt.Axis(format='0.0%', titleX=titleX), title=['All Patients' if patient_group == 'all' else 'Ever-Severe Patients'] if i == 0 else None), # title="% Patients Tested" if i == 0 else None
y2=alt.Y2('ci_95U:Q'),
color=alt.Color("Wave:N", scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR)),
strokeWidth=alt.value(1.5)
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
obs_line = alt.layer(obs_line, obs_error).properties(
# title={
# "text": "Percentage of Patients Tested",
# "dx": 80,
# # "fontSize": 16,
# # "color": "gray"
# },
width=193,
height=150
)
plot = (
line_m if show_patients == False else (
alt.vconcat(
# alt.hconcat(line_m, bar_m, spacing=20).resolve_scale(y='shared'),
# line_m,
obs_line,
# spacing=5
).resolve_scale(x='independent')
)
)
if patient_group == 'all':
plot = plot.properties(
title={
"text": consistent_loinc[lab].replace(' (Normal Sensitivity)', ''),
"anchor": 'middle',
'fontSize': 18,
'dx': (2 if i != 0 else 2) if show_patients == False else (20 if i != 0 else 40)
}
)
return plot
SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'Fibrinogen', 'procalcitonin', 'D-dimer', 'creatinine']
show_patients = False
for i, lab in enumerate(SELECTED_LABS):
new = alt.vconcat(
plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')
if i != 0:
res = alt.hconcat(res, new, spacing=10)
else:
res = new
res = res.properties(
title={
"text": [
f"Mean Standardized Lab Values Of All And Ever-Severe Patients By Wave"
],
"dx": 80
}
)
for i, lab in enumerate(SELECTED_LABS):
new = alt.vconcat(
plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')
if i != 0:
res2 = alt.hconcat(res2, new, spacing=10)
else:
res2 = new
for i, lab in enumerate(SELECTED_LABS):
new = alt.vconcat(
plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=True),
spacing=20
).resolve_scale(y='shared')
if i != 0:
res3 = alt.hconcat(res3, new, spacing=10)
else:
res3 = new
res3 = res3.properties(
title={
"text": [
f"Percentage Of All And Ever-Severe Patients Tested By Wave"
],
"dx": 80
}
)
for i, lab in enumerate(SELECTED_LABS):
new = alt.vconcat(
plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=True),
spacing=20
).resolve_scale(y='shared')
if i != 0:
res4 = alt.hconcat(res4, new, spacing=10)
else:
res4 = new
# res2 = res2.properties(
# title={
# "text": [
# f"Mean Standardized Lab Values Of Ever Severe Patients By Wave"
# ],
# "dx": 80,
# # "titlePadding": 30,
# # "subtitle": [
# # get_visualization_subtitle(data_release='2021-05-06', with_num_sites=False)],
# "subtitleFontSize": 16,
# "subtitleColor": "gray",
# }
# )
# res = apply_theme(
# res,
# axis_y_title_font_size=16,
# title_anchor='start',
# legend_orient='right'
# )
res = alt.vconcat(res, res2, spacing=20).resolve_scale(size='shared')
res2 = alt.vconcat(res3, res4, spacing=20).resolve_scale(size='shared')
res = alt.vconcat(res, res2, spacing=40).resolve_scale(size='independent', stroke='independent', color='independent')
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='bottom',
point_size=30
)
res
```
# All Labs
```
domain_by_lab = {
'C-reactive protein (CRP) (Normal Sensitivity)': [0.85, 1.15],
'Ferritin': [0.97, 1.08],
'Fibrinogen': [0.88, 1.05],
'procalcitonin': [0.9, 2.5],
'D-dimer': [0.96, 1.15],
'creatinine': [0.85, 1.15],
'alanine aminotransferase (ALT)': [0.95, 1.15],
'albumin': [0.75, 1.05],
'aspartate aminotransferase (AST)': [0.96, 1.05],
'lactate dehydrogenase (LDH)': [0.95, 1.05],
'lymphocyte count': [0.85, 1.15],
'neutrophil count': [0.9, 1.55],
'prothrombin time (PT)': [0.95, 1.25],
'total bilirubin': [0.95, 1.25],
'white blood cell count (Leukocytes)': [0.95, 1.5]
}
def plot(_d, patient_group='all', lab=None, obs=None, pdf=None, i=0, show_patients=False):
d = _d.copy()
o = obs.copy()
p = pdf.copy()
"""
DATA PREPROCESSING...
"""
if lab in LOG_LABS:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d = d.drop(columns=['Unnamed: 0'])
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.Lab == lab]
d17 = d[(1 <= d.days_since_positive) & (d.days_since_positive <= 7)]
d = d[(d.days_since_positive == 0) | (d.days_since_positive == 1) | (d.days_since_positive == 7)]
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d17.days_since_positive = d17.days_since_positive.apply(lambda x: f"Day {x}")
#### PVAL ###############################
p = p[p.setting == patient_group]
p['is_sig'] = False
p.is_sig = p[lab] <= 0.05
p.is_sig = p.is_sig.apply(lambda x: 'p<0.05' if x else 'p>0.05')
p = p[['setting','day', 'is_sig']]
p.day = p.day.apply(lambda x: f"Day {x}")
"""
MERGE
"""
d = pd.merge(d, p, how='left', left_on=['days_since_positive','setting'], right_on = ['day','setting'])
#### OBS ################################
o = o.rename(columns={"p.lwr": "p_lwr", "p.upr": "p_upr"})
o = o[o.lab == lab]
o = o.drop(columns=['Unnamed: 0'])
o.wave = o.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
o = o[o.cohort == 'dayX']
o = o[o.setting == patient_group]
o = o.rename(columns={
'wave': 'Wave'
})
"""
DAY 1-7 AVERAGE
"""
d17['mean'] = d17['mean'] * d17['total_n']
d17 = d17.groupby(['Lab', 'setting', 'Wave']).sum().reset_index()
d17['mean'] = d17['mean'] / d17['total_n']
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
"""
PLOT
"""
titleX=-60
opacity=0.7
"""
LABS
"""
LAB_DROPDOWN = alt.binding_select(options=LABS)
LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": lab if lab != None else LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=4, opacity=opacity, point=False
).encode(
x=alt.X('days_since_positive:N', title=None, axis=alt.Axis(labels=False) if (patient_group == 'all') & (show_patients == False) else alt.Axis()), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False, nice=False, padding=0, domain=domain_by_lab[lab]), title=['Mean Lab Value'] if patient_group == 'all' else None, axis=alt.Axis(titleX=titleX)),
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR))
).properties(
width=340,
height=200
)
point_m = alt.Chart(
d
).mark_point(
opacity=opacity, filled=True, strokeWidth=3
).encode(
x=alt.X('days_since_positive:N', title=None), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=['Mean Lab Value'] if patient_group == 'all' else None, axis=alt.Axis(titleX=titleX)), # 'All Patients' if patient_group == 'all' else "Ever Severe Patients",
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR)),
stroke=alt.Stroke('is_sig:N', scale=alt.Scale(domain=['p<0.05'], range=['black']), title='Significance'),
size=alt.Size('total_n:Q', title="# of Patients"),
strokeWidth=alt.value(3)
)
line_m = (line_m + point_m)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
"""
Day 1-7 Average
"""
bar_m = alt.Chart(
d17
).mark_bar(
size=32,
stroke='black'
).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value', axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=1)),
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR))
).properties(
width=200
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
text = alt.Chart(
d17
).mark_text(size=16, dx=0, dy=-4, color='black', baseline='bottom', align='center', angle=0, fontWeight=500).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value' if patient_group == 'all' else None, axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=1)),
text=alt.Text('mean:Q', format=".2f")
)
# .transform_filter(
# LAB_SELECTION
# )
bar_m = (bar_m)# + text)
"""
OBSERVATION
"""
LAB_FIELD_NAME = lab.replace(' ', '.').replace('(', '.').replace(')', '.').replace('-', '.') # Used in the original files
o = o.rename(columns={
f"{LAB_FIELD_NAME}": LAB_FIELD_NAME.replace('.', '_')
})
LAB_FIELD_NAME = LAB_FIELD_NAME.replace('.', '_')
obs_line = alt.Chart(
o
).mark_line(
point=True, opacity=0.7, size=4
).encode(
x=alt.X('day:Q', title='Days Since Positive'),
y=alt.Y(f"p:Q", scale=alt.Scale(domain=[0, 1]), axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested"),
color=alt.Color("Wave:N")
)
obs_error = obs_line.mark_errorbar(opacity=0.3).encode(
x=alt.X('day:Q', title='Days Since Positive', scale=alt.Scale(padding=10, nice=False)),
y=alt.Y('p_upr:Q', axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested" if patient_group == 'all' else None),
y2=alt.Y2('p_lwr:Q'),
color=alt.Color("Wave:N")
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
obs_line = alt.layer(obs_line, obs_error).properties(
# title={
# "text": "Percentage of Patients Tested",
# "dx": 80,
# # "fontSize": 16,
# # "color": "gray"
# },
width=340,
height=200
)
plot = (
line_m if show_patients == False else (
alt.vconcat(
# alt.hconcat(line_m, bar_m, spacing=20).resolve_scale(y='shared'),
line_m,
obs_line,
spacing=5
).resolve_scale(x='independent')
)
).properties(
title={
"text": f"{patient_group.capitalize()} Patients", # == 'all'consistent_loinc[lab].replace(' (Normal Sensitivity)', ''),
"anchor": 'middle',
'fontSize': 16,
'dx': 30 # (5 if i != 0 else 13) if show_patients == False else (20 if i != 0 else 40)
}
)
return plot
SELECTED_LABS = df.Lab.unique().tolist()
show_patients = True
for i, lab in enumerate(SELECTED_LABS):
# DEBUG
# if i == 1:
# break
# new = plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i)
new = alt.hconcat(
plot(df, patient_group='all', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')#, color='independent', size='independent', stroke='independent')
# DEBUG
#if i == 0:
#break
new = new.properties(
title={
"text": [
f"Mean Standardized Lab Values Of All And Severe Patients By Wave"
],
"dx": 80,
"subtitle": [
lab.capitalize()
],
"subtitleFontSize": 18,
"subtitleColor": "gray",
}
)
if i != 0:
res = alt.vconcat(res, new, spacing=30).resolve_scale(y='independent', color='independent', size='independent', stroke='independent')
else:
res = new
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='right',
point_size=40,
title_dy=0
)
res
```
# Standardized Mean Labs
```
def plot(_d, patient_group='all', lab=None, obs=None, pdf=None, i=0, show_patients=False):
d = _d.copy()
o = obs.copy()
p = pdf.copy()
"""
DATA PREPROCESSING...
"""
if lab in LOG_LABS:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d = d.drop(columns=['Unnamed: 0'])
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.Lab == lab]
d17 = d[(1 <= d.days_since_positive) & (d.days_since_positive <= 7)]
d = d[(d.days_since_positive == 0) | (d.days_since_positive == 1) | (d.days_since_positive == 7)]
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d17.days_since_positive = d17.days_since_positive.apply(lambda x: f"Day {x}")
#### PVAL ###############################
p = p[p.setting == patient_group]
p['is_sig'] = False
p.is_sig = p[lab] <= 0.05
p.is_sig = p.is_sig.apply(lambda x: 'p<0.05' if x else 'p>0.05')
p = p[['setting','day', 'is_sig']]
p.day = p.day.apply(lambda x: f"Day {x}")
"""
MERGE
"""
d = pd.merge(d, p, how='left', left_on=['days_since_positive','setting'], right_on = ['day','setting'])
#### OBS ################################
o = o.rename(columns={"p.lwr": "p_lwr", "p.upr": "p_upr"})
o = o[o.lab == lab]
o = o.drop(columns=['Unnamed: 0'])
o.wave = o.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
o = o[o.cohort == 'dayX']
o = o[o.setting == patient_group]
o = o.rename(columns={
'wave': 'Wave'
})
"""
DAY 1-7 AVERAGE
"""
d17['mean'] = d17['mean'] * d17['total_n']
d17 = d17.groupby(['Lab', 'setting', 'Wave']).sum().reset_index()
d17['mean'] = d17['mean'] / d17['total_n']
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
"""
PLOT
"""
titleX=-60
opacity=0.7
"""
LABS
"""
LAB_DROPDOWN = alt.binding_select(options=LABS)
LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": lab if lab != None else LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=4, opacity=opacity, point=False
).encode(
x=alt.X('days_since_positive:N', title=None, axis=alt.Axis(labels=False) if (patient_group == 'all') & (show_patients == False) else alt.Axis()), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False, nice=False, padding=10), title=['Mean Lab Value'] if i == 0 else None, axis=alt.Axis(titleX=titleX)), #domain=domain_by_lab[lab]
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR))
).properties(
width=240,
height=200
)
point_m = alt.Chart(
d
).mark_point(
opacity=opacity, filled=True, strokeWidth=3
).encode(
x=alt.X('days_since_positive:N', title=None), # ''Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=['Mean Lab Value'] if i == 0 else None, axis=alt.Axis(titleX=titleX)), # 'All Patients' if patient_group == 'all' else "Ever Severe Patients",
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR)),
stroke=alt.Stroke('is_sig:N', scale=alt.Scale(domain=['p<0.05'], range=['black']), title='Significance'),
size=alt.Size('total_n:Q', title="# of Patients"),
strokeWidth=alt.value(3)
)
line_m = (line_m + point_m)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
"""
Day 1-7 Average
"""
bar_m = alt.Chart(
d17
).mark_bar(
size=32,
stroke='black'
).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value', axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=1)),
color=alt.Color('Wave:N', scale=alt.Scale(range=WAVE_COLOR))
).properties(
width=100
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
text = alt.Chart(
d17
).mark_text(size=16, dx=0, dy=-4, color='black', baseline='bottom', align='center', angle=0, fontWeight=500).encode(
x=alt.X('Wave:N', title=None), # 'Wave'),
y=alt.Y('mean:Q', title='Day1-7 Mean Lab Value' if i == 0 else None, axis=alt.Axis(ticks=False, labels=False, domain=False, orient='left'), scale=alt.Scale(padding=1)),
text=alt.Text('mean:Q', format=".2f")
)
# .transform_filter(
# LAB_SELECTION
# )
bar_m = (bar_m)# + text)
"""
OBSERVATION
"""
LAB_FIELD_NAME = lab.replace(' ', '.').replace('(', '.').replace(')', '.').replace('-', '.') # Used in the original files
o = o.rename(columns={
f"{LAB_FIELD_NAME}": LAB_FIELD_NAME.replace('.', '_')
})
LAB_FIELD_NAME = LAB_FIELD_NAME.replace('.', '_')
obs_line = alt.Chart(
o
).mark_line(
point=True, opacity=0.7, size=4
).encode(
x=alt.X('day:Q', title='Days Since Positive'),
y=alt.Y(f"p:Q", scale=alt.Scale(domain=[0, 1]), axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested"),
color=alt.Color("Wave:N")
)
obs_error = obs_line.mark_errorbar(opacity=0.3).encode(
x=alt.X('day:Q', title='Days Since Positive', scale=alt.Scale(padding=10, nice=False)),
y=alt.Y('p_upr:Q', axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients Tested" if i == 0 else None),
y2=alt.Y2('p_lwr:Q'),
color=alt.Color("Wave:N")
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
obs_line = alt.layer(obs_line, obs_error).properties(
# title={
# "text": "Percentage of Patients Tested",
# "dx": 80,
# # "fontSize": 16,
# # "color": "gray"
# },
width=240,
height=150
)
plot = (
line_m if show_patients == False else (
alt.vconcat(
# alt.hconcat(line_m, bar_m, spacing=20).resolve_scale(y='shared'),
line_m,
obs_line,
spacing=5
).resolve_scale(x='independent')
)
).properties(
title={
"text": consistent_loinc[lab].replace(' (Normal Sensitivity)', ''),
"anchor": 'middle',
'fontSize': 16,
'dx': (5 if i != 0 else 13) if show_patients == False else (20 if i != 0 else 40)
}
)
return plot
SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'albumin', 'creatinine', 'D-dimer']
show_patients = True
for i, lab in enumerate(SELECTED_LABS):
new = alt.vconcat(
plot(df, patient_group='ever', lab=lab, obs=obs, pdf=pdf, i=i, show_patients=show_patients),
spacing=20
).resolve_scale(y='shared')
if i != 0:
res = alt.hconcat(res, new, spacing=10)
else:
res = new
# DEBUG
if i == 0:
break
res = res.properties(
title={
"text": [
f"Mean Standardized Lab Values Of Ever Severe Patients By Wave"
],
"dx": 80
}
)
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='right'
)
res
cdf = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_values_standardized_bycountry.csv"))
cdf
cobs = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_obs_bycountry_for_Sehi.csv"))
cobs
fr = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_FRANCE.csv"))
ge = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_GERMANY.csv"))
it = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_ITALY.csv"))
sp = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_SPAIN.csv"))
us = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_USA.csv"))
cpdf = fr.append([ge, it, sp, us])
cpdf = cpdf.drop(columns=['Unnamed: 0'])
cpdf
def plot_country(_d, patient_group='all', lab=None, obs=None, pdf=None):
d = _d.copy()
o = obs.copy()
p = pdf.copy()
"""
DATA PREPROCESSING...
"""
if lab in ['C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer']:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d.wave = d.wave.apply(lambda x: { 'early': 'Early', 'late': 'Late' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.country != 'SINGAPORE']
# d = d[d.country != 'GERMANY']
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
d = d[(d.days_since_positive == 0) | (d.days_since_positive == 1) | (d.days_since_positive == 7)]
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d = d[d.Lab == lab]
"""
OBS DATA
"""
# o = o.drop(columns=['Unnamed: 0'])
o = o[o.lab == lab]
o.wave = o.wave.apply(lambda x: { 'early': 'Early', 'late': 'Late' }[x])
o = o[o.cohort == 'dayX']
o = o[o.country != 'GERMANY']
o.country = o.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
o = o[o.setting == patient_group]
o = o.rename(columns={
'wave': 'Wave'
})
"""
PVAL DATA
"""
p = p[p.setting == patient_group]
p.country = p.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
p['is_sig'] = False
p.is_sig = p[lab] <= 0.05
p.is_sig = p.is_sig.apply(lambda x: 'p<0.05' if x else 'p>0.05')
p = p[['setting','day', 'country', 'is_sig']]
p.day = p.day.apply(lambda x: f"Day {x}")
"""
MERGE
"""
d = pd.merge(d, p, how='left', left_on=['days_since_positive','setting', 'country'], right_on = ['day','setting', 'country'])
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
COUNTRIES = ['France', 'Germany', 'Italy', 'Spain', 'USA']
COUNTRY_COLORS = ['#0072B2', '#E6A01B', '#029F73', '#D45E00', '#CB7AA7']
"""
PLOT
"""
titleX=-60
opacity=0.7
"""
LABS
"""
LAB_DROPDOWN = alt.binding_select(options=LABS)
LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=4, opacity=opacity
).encode(
x=alt.X('days_since_positive:N', title=None),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title='Standardized Mean Lab', axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
).properties(
width=200,
height=200
)
point_m = line_m.mark_point(
size=80, opacity=opacity, filled=True
).encode(
x=alt.X('days_since_positive:N', title=None),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title='Standardized Mean Lab', axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS)),
# stroke=alt.Stroke('is_sig:N', scale=alt.Scale(domain=['p<0.05'], range=['black']), title='Significance'),
# strokeWidth=alt.value(3)
)
line_m = (line_m + point_m).facet(
column=alt.Column('Wave:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True))
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
"""
OBSERVATION
"""
LAB_FIELD_NAME = lab.replace(' ', '.').replace('(', '.').replace(')', '.').replace('-', '.') # Used in the original files
o = o.rename(columns={
f"{LAB_FIELD_NAME}": LAB_FIELD_NAME.replace('.', '_')
})
LAB_FIELD_NAME = LAB_FIELD_NAME.replace('.', '_')
obs_line = alt.Chart(
o
).mark_line(
point=True, opacity=0.7, size=4
).encode(
x=alt.X('day:Q', title='Days Since Positive'),
y=alt.Y(f"p:Q", axis=alt.Axis(format='0.0%', titleX=titleX), title="% Patients"),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS)),
column=alt.Column('Wave:N', header=alt.Header(labelOrient="top", title=None, titleOrient="top", labels=False))
).properties(
width=200,
height=100
)
# .properties(
# title={
# "text": "Percentage of Patients Tested",
# "dx": 80,
# # "fontSize": 16,
# # "color": "gray"
# },
# width=350,
# height=100
# )
line_m = alt.vconcat(line_m, obs_line, spacing=20)
plot = (
alt.hconcat(line_m, spacing=20).resolve_scale(y='shared')
).properties(
title={
"text": f"Country-Level Lab Values of {'All' if patient_group == 'all' else 'Ever Severe'} Patients by Wave",
"dx": 80,
"subtitle": [
consistent_loinc[lab], #.title(),
get_visualization_subtitle(data_release='2021-04-27', with_num_sites=False)
],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
return plot
LABS = df.Lab.unique().tolist()
# print(len(LABS))
for i, lab in enumerate(LABS):
new = alt.hconcat(
plot_country(cdf, patient_group='all', lab=lab, obs=cobs, pdf=cpdf),
plot_country(cdf, patient_group='ever', lab=lab, obs=cobs, pdf=cpdf),
spacing=50
).resolve_scale(y='shared', color='independent', size='independent', stroke='independent')
if i != 0:
res = alt.vconcat(res, new, spacing=10)
else:
res = new
# DEBUG
# if i == 0:
# break
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='right',
header_label_font_size=16
)
res
# LABS = df.Lab.unique().tolist()
# print(len(LABS))
for i, lab in enumerate(SELECTED_LABS):
new = alt.hconcat(
plot_country(cdf, patient_group='all', lab=lab, obs=cobs, pdf=cpdf),
plot_country(cdf, patient_group='ever', lab=lab, obs=cobs, pdf=cpdf),
spacing=50
).resolve_scale(y='shared', color='independent', size='independent', stroke='independent')
if i != 0:
res = alt.vconcat(res, new, spacing=10)
else:
res = new
# DEBUG
# if i == 0:
# break
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
legend_orient='right',
header_label_font_size=16
)
res
def day0_overview(_d, lab, _p):
d = _d.copy()
p = _p.copy()
"""
DATA PREPROCESSING...
"""
d.wave = d.wave.apply(lambda x: { 'early': 'Early', 'late': 'Late' }[x])
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.country != 'SINGAPORE']
d = d[d.country != 'GERMANY']
d = d[d.days_since_positive == 0]
d = d[d.Lab == lab]
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
d.days_since_positive = d.days_since_positive.apply(lambda x: f"Day {x}")
d.setting = d.setting.apply(lambda x: {'all': 'All Patients', 'ever': 'Ever Severe Patients', 'never': 'Never Severe Patients'}[x])
if lab in ['C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer']:
d = d[d.scale == 'log']
p = p[p.scale == 'log']
else:
d = d[d.scale == 'original']
p = p[p.scale == 'original']
d = d.drop(columns=['scale', 'Lab', 'days_since_positive'])
d = d.pivot_table(values='mean', index=['country', 'setting'], columns='Wave').reset_index()
d = d[(d.Early.notna() & d.Late.notna())]
d['increase'] = d.Early < d.Late
p.country = p.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
p['sig'] = p[lab].apply(lambda x: 'yes' if x < 0.05 else 'no')
p[lab] = -np.log10(p[lab])
d = pd.merge(d, p, how='left', left_on=['country','setting'], right_on = ['country','setting'])
# print(d)
# print(p[lab])
# print(p)
"""
CONSTANTS
"""
COUNTRIES = ['France', 'Italy', 'Spain', 'USA'] # 'Germany',
COUNTRY_COLORS = ['#0072B2', '#029F73', '#D45E00', '#CB7AA7'] # '#E79F00',
"""
PLOT
"""
titleX=-80
opacity=0.7
"""
LABS
"""
bar = alt.Chart(
d
).mark_bar(
size=6
).encode(
y=alt.Y('Early:Q', scale=alt.Scale(zero=False), title='Mean Lab Value'),
y2=alt.Y2('Late:Q'),
x=alt.X('country:N', title='Country'), # axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
).properties(
width=200,
height=220
)
tr = alt.Chart(
d
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[True])
).mark_point(
shape="triangle-up", filled=True, size=300, yOffset=3, opacity=1
).encode(
y=alt.Y('Late:Q', scale=alt.Scale(zero=False), title='Mean Lab Value'),
x=alt.X('country:N', title='Country'),# axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
)
tl = alt.Chart(
d
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[False])
).mark_point(
shape="triangle-down", filled=True, size=300, yOffset=-3, opacity=1
).encode(
y=alt.Y('Late:Q', scale=alt.Scale(zero=False), title='Mean Lab Value'),
x=alt.X('country:N', title='Country'), # axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
)
baseline = alt.Chart(
pd.DataFrame({'baseline': [1]})
).mark_rule(color='gray').encode(
y=alt.Y('baseline:Q')
)
plot = (bar + tr + tl).facet(
column=alt.Column('setting:N', header=alt.Header(labelOrient="top", title=None, titleOrient="top", labels=True)),
spacing=30
)
# .add_selection(
# LAB_SELECTION
# ).transform_filter(
# LAB_SELECTION
# )
plot = plot.properties(
title={
"text": [
f"Standardized Mean Lab Values by Country from Early to Late Wave"
],
"dx": 50,
"subtitle": [
# 'Relative to Day 0 Mean Value During Early Phase',
consistent_loinc[lab],
#get_visualization_subtitle(data_release='2021-01-25', with_num_sites=False)
],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
"""
p-value triangles
"""
d_notna = d[d[lab].notna()].copy()
p_base = alt.Chart(
d_notna
).mark_point(
size=200, filled=True, opacity=1, shape='triangle-up', strokeWidth=1, xOffset=0
).encode(
x=alt.X('country:N', title='Country'),# axis=alt.Axis(titleX=titleX)),
y=alt.Y(f'{lab}:Q', title="P Value (-log10)", scale=alt.Scale(zero=False)),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS), title='Country'),
stroke=alt.Stroke('sig:N', scale=alt.Scale(domain=['no', 'yes'], range=['white', 'black']), title='p < 0.05?', legend=None)
).properties(
width=200,
height=200
)
p_base_no_y = p_base.encode(
x=alt.X('country:N', title='Country'),
y=alt.Y(f'{lab}:Q', title="P Value (-log10)", scale=alt.Scale(zero=False), axis=alt.Axis(ticks=False, domain=False, title=None, labels=False)),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS)),
stroke=alt.Stroke('sig:N', scale=alt.Scale(domain=['no', 'yes'], range=['white', 'black']), title='p < 0.05?', legend=None)
)
p0_05_all = alt.Chart(
pd.DataFrame({'baseline': [1.30102999566], 'zero': [0]})
).mark_rule(color='firebrick', strokeDash=[3,3]).encode(
y=alt.Y('baseline:Q')
)
p0_05_all_rect = alt.Chart(
pd.DataFrame({'baseline': [1.30102999566]})
).mark_rect(color='transparent', stroke='firebrick', strokeDash=[3,3], strokeWidth=2).encode(
y=alt.Y('baseline:Q'),
y2=alt.value(0)
)
p0_05_text = alt.Chart(
pd.DataFrame({'baseline': [1.30102999566], 'text': 'p=0.05'})
).mark_text(color='firebrick', align='right', baseline='top', y=0, x=195).encode(
# y=alt.Y('baseline:Q'),
y=alt.value(2),
text=alt.value('Statistical Significance: p<0.05')
)
"""
ALL PATIENTS
"""
p_t_all1 = (
#p0_05_all
#+
p0_05_text
+
p_base.transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[True])
).transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['All Patients'])
)
+
p_base.mark_point(
size=200, filled=True, opacity=1, shape='triangle-down', strokeWidth=1, xOffset=0
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[False])
).transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['All Patients'])
)
+
p0_05_all_rect
).properties(
title={
'text': 'All Patients',
'dx': 105,
'fontSize': 16
}
)
"""
EVER SEVERE
"""
p_t_all2 = (
#p0_05_all
#+
p0_05_text
+
p_base_no_y.transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['Ever Severe Patients'])
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[True])
)
+
p_base_no_y.mark_point(
size=200, filled=True, opacity=1, shape='triangle-down', strokeWidth=1, xOffset=0
).transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['Ever Severe Patients'])
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[False])
)
+ p0_05_all_rect
).properties(
title={
'text': 'Ever Severe Patients',
'dx': 30,
'fontSize': 16
}
)
"""
NEVER SEVERE
"""
p_t_all3 = (
#p0_05_all
#+
p0_05_text
+
p_base_no_y.transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['Never Severe Patients'])
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[True])
)
+
p_base_no_y.mark_point(
size=200, filled=True, opacity=1, shape='triangle-down', strokeWidth=1, xOffset=0
).transform_filter(
alt.FieldOneOfPredicate(field='setting', oneOf=['Never Severe Patients'])
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[False])
)
+ p0_05_all_rect
).properties(
title={
'text': 'Never Severe Patients',
'dx': 30,
'fontSize': 16
}
)
"""
COMBINE
"""
p_t = alt.hconcat(p_t_all1, p_t_all2, p_t_all3, spacing=20).resolve_scale(y='shared')
p_t = p_t.properties(
title={
"text": [
f"P-Values at 0 Days Since Positive Across Waves"
],
"dx": 50,
''
"subtitle": [
# 'Relative to Day 0 Mean Value During Early Phase',
consistent_loinc[lab],
#get_visualization_subtitle(data_release='2021-01-25', with_num_sites=False)
],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
return alt.vconcat(plot, p_t, spacing=30).resolve_scale(color='independent', stroke='independent')
fr = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_FRANCE.csv"))
ge = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_GERMANY.csv"))
it = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_ITALY.csv"))
sp = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_SPAIN.csv"))
us = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_pvals_USA.csv"))
pdf = fr.append([ge, it, sp, us])
pdf = pdf.drop(columns=['Unnamed: 0'])
# pdf = pdf[pdf.setting != 'never']
pdf = pdf[pdf.day == 0]
pdf.setting = pdf.setting.apply(lambda x: { 'all': 'All Patients', 'ever': 'Ever Severe Patients', 'never': 'Never Severe Patients'}[x])
pdf
# lab = df.Lab.unique().tolist()[0]
labs = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'D-dimer', 'creatinine', 'albumin']
for l in labs:
plot = day0_overview(cdf, l, pdf)
plot = apply_theme(
plot,
axis_y_title_font_size=16,
title_anchor='start',
# legend_title_orient='left',
legend_orient='right',
header_label_font_size=16
)
plot.display()
# cdf
def plot_country_raw(_d, patient_group='all', lab=None):
d = _d.copy()
"""
DATA PREPROCESSING...
"""
d['se_top'] = d['mean'] + d['se']
d['se_bottom'] = d['mean'] - d['se']
d.wave = d.wave.apply(lambda x: { 'early': 'First Wave', 'late': 'Second Wave' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.country != 'SINGAPORE']
# d = d[d.country != 'GERMANY']
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
d = d[d.Lab == lab]
# ['C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer']
if lab in ['alanine aminotransferase (ALT)', 'aspartate aminotransferase (AST)', 'C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer', 'Ferritin', 'lactate dehydrogenase (LDH)']:
d = d[d.scale == 'log']
yTitle = 'Mean Lab Values (Log)'
else:
d = d[d.scale == 'original']
yTitle = 'Mean Lab Values'
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
COUNTRIES = ['France', 'Germany', 'Italy', 'Spain', 'USA']
COUNTRY_COLORS = ['#0072B2', '#E6A01B', '#029F73', '#D45E00', '#CB7AA7']
"""
PLOT
"""
titleX=-60
opacity=0.7
width=350
height=250
size=3
"""
LABS
"""
# LAB_DROPDOWN = alt.binding_select(options=LABS)
# LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=size, opacity=opacity, point=True
).encode(
x=alt.X('days_since_positive:N', title='Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=yTitle, axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS), title='Country')
).properties(
width=width,
height=height
)
"""
ERROR BAR
"""
error_m = line_m.mark_errorbar(color='gray', opacity=0.3).encode(
x=alt.X('days_since_positive:N', title='Days Since Positive'),
y=alt.Y('se_top:Q', scale=alt.Scale(zero=False), title=yTitle, axis=alt.Axis(titleX=titleX)),
y2=alt.Y2('se_bottom:Q'),
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
"""
COMBINE
"""
line_m = (line_m + error_m).facet(
column=alt.Column('Wave:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True))
)
plot = (
alt.hconcat(line_m, spacing=20).resolve_scale(y='shared')
).properties(
title={
"text": f"Country-Level Mean Lab Values Of {'All' if patient_group == 'all' else 'Ever Severe'} Patients By Wave",
"dx": 80,
"subtitle": [
consistent_loinc[lab]
# lab, #.title(),
# get_visualization_subtitle(data_release='2021-04-25', with_num_sites=False)
],
"subtitleFontSize": 18,
"subtitleColor": "gray",
}
)
return plot
raw = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_values_bycountry.csv"))
print(raw.Lab.unique().tolist())
# raw = raw[~pd.isnull(raw.mean)]
# !! We are removing the following lab since this does not contain any info
raw = raw[raw.Lab != 'cardiac troponin (Normal Sensitivity)']
labs = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'D-dimer', 'creatinine', 'albumin']
labs = raw.Lab.unique().tolist()
patient_groups = ['all', 'ever']
for li, lab in enumerate(labs):
for i, patient_group in enumerate(patient_groups):
if i == 0:
res = plot_country_raw(raw, patient_group=patient_group, lab=lab)
else:
res = alt.hconcat(
res, plot_country_raw(raw, patient_group=patient_group, lab=lab), spacing=30
).resolve_scale(y='shared', color='shared')
if li == 0:
plot = res
else:
plot = alt.vconcat(
plot, res, spacing=30
).resolve_scale(y='independent', color='independent')
# plot = plot.properties(
# title={
# "text": f"Country-Level Mean Lab Values Of {'All' if patient_group == 'all' else 'Ever Severe'} Patients By Wave",
# "dx": 80
# }
# )
plot = apply_theme(
plot,
axis_y_title_font_size=16,
header_label_font_size=16,
title_anchor='start',
legend_orient='right'
)
plot
def plot_country_raw(_d, patient_group='all', lab=None):
d = _d.copy()
"""
DATA PREPROCESSING...
"""
d['se_top'] = d['mean'] + d['se']
d['se_bottom'] = d['mean'] - d['se']
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.country != 'SINGAPORE']
# d = d[d.country != 'GERMANY']
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
d = d[d.Lab == lab]
# ['C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer']
if lab in ['alanine aminotransferase (ALT)', 'aspartate aminotransferase (AST)', 'C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer', 'Ferritin', 'lactate dehydrogenase (LDH)']:
d = d[d.scale == 'log']
yTitle = 'Mean Lab Values (Log)'
else:
d = d[d.scale == 'original']
yTitle = 'Mean Lab Values'
"""
CONSTANTS
"""
LABS = d.Lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
COUNTRIES = ['France', 'Germany', 'Italy', 'Spain', 'USA']
COUNTRY_COLORS = ['#0072B2', '#E6A01B', '#029F73', '#D45E00', '#CB7AA7']
"""
PLOT
"""
titleX=-60
opacity=0.7
width=350
height=250
size=3
"""
LABS
"""
# LAB_DROPDOWN = alt.binding_select(options=LABS)
# LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=size, opacity=opacity, point=True
).encode(
x=alt.X('days_since_positive:N', title='Days Since Positive'),
y=alt.Y('mean:Q', scale=alt.Scale(zero=False), title=yTitle, axis=alt.Axis(titleX=titleX, tickCount=6)),
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR), title='Wave')
).properties(
width=width,
height=height
)
"""
ERROR BAR
"""
error_m = line_m.mark_errorbar(color='gray', opacity=0.3).encode(
x=alt.X('days_since_positive:N', title='Days Since Positive'),
y=alt.Y('se_top:Q', scale=alt.Scale(zero=False), title=yTitle, axis=alt.Axis(titleX=titleX)),
y2=alt.Y2('se_bottom:Q'),
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
"""
COMBINE
"""
# line_m = (line_m + error_m).facet(
# column=alt.Column('Wave:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True))
# )
line_m = (line_m + error_m).facet(
column=alt.Column('country:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True))
)
plot = (
alt.hconcat(line_m, spacing=20).resolve_scale(y='shared')
).properties(
title={
"text": f"Country-Level Mean Lab Values Of {'All' if patient_group == 'all' else 'Ever Severe'} Patients By Wave",
"dx": 80,
"subtitle": [
consistent_loinc[lab]
# # lab, #.title(),
# # get_visualization_subtitle(data_release='2021-04-25', with_num_sites=False)
],
"subtitleFontSize": 18,
"subtitleColor": "gray",
}
)
return plot
raw = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_values_bycountry.csv"))
# raw = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_values_standardized_bycountry.csv"))
# labs = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'D-dimer', 'creatinine', 'albumin']
labs = raw.Lab.unique().tolist()
# SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'Fibrinogen', 'procalcitonin', 'D-dimer', 'creatinine']
patient_groups = ['all', 'ever']
for li, lab in enumerate(labs):
for i, patient_group in enumerate(patient_groups):
if i == 0:
res = plot_country_raw(raw, patient_group=patient_group, lab=lab)
else:
res = alt.hconcat(
res, plot_country_raw(raw, patient_group=patient_group, lab=lab), spacing=30
).resolve_scale(y='shared', color='independent')
if li == 0:
plot = res
else:
plot = alt.vconcat(
plot, res, spacing=30
).resolve_scale(y='independent', color='independent')
# plot = plot.properties(
# title={
# "text": "Proportion Of Patients Being Tested In Each Country",
# "dx": 80
# }
# )
plot = apply_theme(
plot,
axis_y_title_font_size=16,
header_label_font_size=16,
title_anchor='start',
legend_orient='right',
title_dy=0
)
plot
def plot_country_raw(_d, patient_group='all', lab=None):
d = _d.copy()
"""
DATA PREPROCESSING...
"""
# d['se_top'] = d['mean'] + d['se']
# d['se_bottom'] = d['mean'] - d['se']
d.wave = d.wave.apply(lambda x: { 'early': 'First', 'late': 'Second' }[x])
d = d[d.setting != 'never']
d = d[d.setting == patient_group]
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.cohort == 'dayX']
# d = d[d.country != 'SINGAPORE']
# d = d[d.country != 'GERMANY']
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
d = d[d.lab == lab]
# ['C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer']
yTitle = consistent_loinc[lab].replace('(Normal Sensitivity)', '') # 'Mean Lab Values'
# if lab in ['alanine aminotransferase (ALT)', 'aspartate aminotransferase (AST)', 'C-reactive protein (CRP) (Normal Sensitivity)', 'D-dimer', 'Ferritin', 'lactate dehydrogenase (LDH)']:
# d = d[d.scale == 'log']
# yTitle = 'Mean Lab Values (Log)'
# else:
# d = d[d.scale == 'original']
# yTitle = 'Mean Lab Values'
"""
CONSTANTS
"""
LABS = d.lab.unique().tolist()
WAVE_COLOR = [
'#D45E00', # '#BA4338', # early
'#0072B2', # late
'black'
]
COUNTRY_COLORS = ['#0072B2', '#029F73', '#D45E00', '#CB7AA7']
"""
PLOT
"""
titleX=-60
opacity=0.7
width=260
height=200
size=3
"""
LABS
"""
# LAB_DROPDOWN = alt.binding_select(options=LABS)
# LAB_SELECTION = alt.selection_single(fields=["Lab"], bind=LAB_DROPDOWN, init={"Lab": LABS[0]}, name="Select")
line_m = alt.Chart(
d
).mark_line(
size=size, opacity=opacity, point=True
).encode(
x=alt.X('day:Q', title='Days Since Admission'),
y=alt.Y('p:Q', scale=alt.Scale(zero=True, clamp=True), title=yTitle, axis=alt.Axis(titleX=titleX, format='%')),
color=alt.Color('Wave:N', scale=alt.Scale(domain=['First', 'Second'], range=WAVE_COLOR), title='Wave')
).properties(
width=width,
height=height
)
"""
ERROR BAR
"""
error_m = line_m.mark_errorbar(color='gray', opacity=0.6).encode(
x=alt.X('day:Q', title='Days Since Admission' if lab == 'creatinine' else None, axis=alt.Axis(labels=True if lab == 'creatinine' else False), scale=alt.Scale(padding=10, nice=False)),
y=alt.Y('ci_95U:Q', scale=alt.Scale(zero=True), title=yTitle, axis=alt.Axis(titleX=titleX)),
y2=alt.Y2('ci_95L:Q'),
strokeWidth=alt.value(1.5)
# color=alt.value('gray')
# color=alt.Color('country:N', scale=alt.Scale(range=COUNTRY_COLORS))
)
"""
COMBINE
"""
# line_m = (line_m + error_m).facet(
# column=alt.Column('Wave:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True))
# )
line_m = (line_m + error_m).facet(
column=alt.Column('country:N', header=alt.Header(labelOrient="top", title=None, titleOrient="bottom", labels=True if lab == 'C-reactive protein (CRP) (Normal Sensitivity)' else False))
)
plot = (
alt.hconcat(line_m, spacing=20).resolve_scale(y='shared')
)
# .properties(
# title={
# "text": consistent_loinc[lab], #f"Country-Level Mean Lab Values Of {'All' if patient_group == 'all' else 'Ever Severe'} Patients By Wave",
# "dx": 80,
# # "subtitle": [
# # consistent_loinc[lab]
# # # lab, #.title(),
# # # get_visualization_subtitle(data_release='2021-04-25', with_num_sites=False)
# # ],
# "subtitleFontSize": 18,
# "subtitleColor": "gray",
# }
# )
return plot
obs
obs = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_obs_bycountry_for_Sehi.csv"))
# obs = obs[obs.country != 'GERMANY']
# obs = obs[~((obs.country == 'FRANCE') & (obs.lab == 'procalcitonin'))] # & (obs.wave == 'late'))]
# obs = obs[~((obs.country == 'GERMANY') & (obs.lab == 'creatinine'))] # & (obs.wave == 'late'))]
### Replace values with some problem in the original file
obs_fixed = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "creatinine_germany_procalcitonin_france_lab_obs_bycountry_for_Sehi.csv"))
obs = obs[~((obs.country == 'FRANCE') & (obs.lab == 'procalcitonin'))]
obs = obs[~((obs.country == 'GERMANY') & (obs.lab == 'creatinine'))]
obs = obs.append(obs_fixed)
### Leave a certain sub-panel blank
obs = obs[~((obs.country == 'GERMANY') & (obs.lab == 'creatinine'))]
obs = obs.append({'day': '1', 'setting': 'all', 'p': None, 'lab': 'procalcitonin', 'country': 'FRANCE', 'wave': 'late', 'cohort': 'dayX'}, ignore_index=True)
obs = obs.append({'day': '1', 'setting': 'all', 'p': None, 'lab': 'creatinine', 'country': 'GERMANY', 'wave': 'late', 'cohort': 'dayX'}, ignore_index=True)
SELECTED_LABS = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'Fibrinogen', 'procalcitonin', 'D-dimer', 'creatinine']
patient_groups = ['all']
for li, lab in enumerate(SELECTED_LABS):
for i, patient_group in enumerate(patient_groups):
if i == 0:
res = plot_country_raw(obs, patient_group=patient_group, lab=lab)
else:
res = alt.hconcat(
res, plot_country_raw(obs, patient_group=patient_group, lab=lab), spacing=0
).resolve_scale(y='shared', color='independent')
if li == 0:
plot = res
else:
plot = alt.vconcat(
plot, res, spacing=20
).resolve_scale(y='independent', color='shared')
plot = plot.properties(
title={
"text": "Laboratory Testing Rates Across Hospitalization Days In Each Country",
"dx": 80,
"dy": -10
}
)
plot = apply_theme(
plot,
axis_y_title_font_size=20,
header_label_font_size=20,
title_anchor='start',
legend_orient='bottom',
legend_title_orient='left',
axis_domain_width=0,
title_dy=0,
point_size=100
)
plot
# obs[(obs.country == 'SPAIN') & (obs.day == 14) & (obs.setting == 'all') & (obs.lab == 'C-reactive protein (CRP) (Normal Sensitivity)')]
# obs.country.unique().tolist()
# obs
# obs_fixed[(obs_fixed.country == 'GERMANY') & (obs_fixed.lab == 'creatinine') & (obs_fixed.day > 5)]
cdf[(cdf.country == 'ITALY') & (cdf.Lab == 'Ferritin') & (cdf.days_since_positive == 0)]
obd = pd.read_csv(join("..", "data", "1.1.resurgence", "labs", "lab_obs_bycountry.csv"))
obd
obd = obd[(obd.cohort == 'dayX') & (obd.setting == 'all') & (obd.day == 0)]
obd
def obs(obd, lab):
d = obd.copy()
"""
DATA PREPROCESSING...
"""
d.wave = d.wave.apply(lambda x: { 'early': 'Early', 'late': 'Late' }[x])
d = d.rename(columns={
'wave': 'Wave'
})
d = d[d.country != 'SINGAPORE']
d = d[d.country != 'GERMANY']
d.country = d.country.apply(lambda x: x.capitalize() if x != 'USA' else x)
# "Also, for the lab observations, in germany whenever there’s just a number 0, we’ll count it as unobserved and we should show no arrow for that"
d = d[~((d.country == "Germany") & (d[lab] == 0))]
d = d.pivot_table(values=lab, index=['country', 'setting'], columns='Wave').reset_index()
d = d[(d.Early.notna() & d.Late.notna())]
d['increase'] = d.Early < d.Late
"""
CONSTANTS
"""
COUNTRIES = ['France', 'Italy', 'Spain', 'USA']
COUNTRY_COLORS = ['#0072B2','#029F73', '#D45E00', '#CB7AA7']
"""
PLOT
"""
titleX=-80
opacity=0.7
"""
LABS
"""
bar = alt.Chart(
d
).mark_bar(
size=6
).encode(
x=alt.X('Early:Q', scale=alt.Scale(zero=False), title='Percentage of Patients'),
x2=alt.X2('Late:Q'),
y=alt.Y('country:N', title='Country', axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
).properties(
width=300,
height=320
)
tr = alt.Chart(
d
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[True])
).mark_point(
shape="triangle-right", filled=True, size=300, xOffset=-3, opacity=1
).encode(
x=alt.X('Late:Q', scale=alt.Scale(zero=False), title='Percentage of Patients'),
y=alt.Y('country:N', title='Country', axis=alt.Axis(titleX=titleX)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
)
tl = alt.Chart(
d
).transform_filter(
alt.FieldOneOfPredicate(field='increase', oneOf=[False])
).mark_point(
shape="triangle-left", filled=True, size=300, xOffset=3, opacity=1
).encode(
x=alt.X('Late:Q', scale=alt.Scale(zero=False), title='Percentage of Patients'),
y=alt.Y('country:N', title='Country', axis=alt.Axis(titleX=titleX), scale=alt.Scale(domain=COUNTRIES)),
color=alt.Color('country:N', title='Country', scale=alt.Scale(domain=COUNTRIES, range=COUNTRY_COLORS))
)
baseline = alt.Chart(
pd.DataFrame({'baseline': [1]})
).mark_rule(color='gray').encode(
x=alt.X('baseline:Q')
)
plot = (bar + tr + tl)
plot = plot.properties(
title={
"text": [
f"Patients Tested at Day 0 from Early to Late Wave"
],
"dx": 95,
"subtitle": [
# 'Relative to Day 0 Mean Value During Early Phase',
consistent_loinc[lab],
#get_visualization_subtitle(data_release='2021-01-25', with_num_sites=False)
],
"subtitleFontSize": 16,
"subtitleColor": "gray",
}
)
return plot
labs = ['C-reactive protein (CRP) (Normal Sensitivity)', 'Ferritin', 'D-dimer', 'creatinine', 'albumin']
for lab in labs:
res = obs(obd, lab)
res = apply_theme(
res,
axis_y_title_font_size=16,
title_anchor='start',
# legend_title_orient='left',
legend_orient='right',
header_label_font_size=16
)
res.display()
```
| github_jupyter |
The BWT
-------
Follow along at doi:10.1093/bioinformatics/btp324, _Fast and accurate short read alignment with Burrows–Wheeler transform_ by Heng Li and Richard Durbin.
But note that a couple of their definitions seem to be incorrect. Adjustments will be noted.
For an alphabet of symbols:
$\Sigma = [ A, C, G, T ] $
$\alpha = $ one symbol of alphabet $\Sigma$
${X}$ = source string $\alpha_{0}\alpha_{1}\ldots\alpha_{n-1}$
$\$ =$ end of string
${n}$ = number of symbols in ${X}$
${X}[i] = \alpha_i$
${i} = 0,1,\ldots,{n-1}$
${X}[i,j] = \alpha_i\ldots\alpha_j$ (substring)
${X_i} = X[i,n-1]$ (suffix)
$S(i) = i$ th lexicographically smallest suffix (aka index into X where suffix starts)
$B$ is the BWT string: list of symbols that precede the first symbol in the sorted suffix list
>$B[i] = \$$ when $S(i) = 0$
>$B[i] = X[S(i) - 1]$
$W =$ a potential substring of $X$
Bounds:
>$\underline{R}(W) = min\{k:W $ is the prefix of $X_{S(k)}\}$
>$\overline{R}(W) = max\{k:W $ is the prefix of $X_{S(k)}\}$
For empty string $W = \$$:
>$\underline{R}(W) = 0$
>$\overline{R}(W) = n - 1$
(Note that Li and Durbin define $\underline{R}(W) = 1$ for empty string W to eliminate the need to define $O(\alpha, -1)$, but this leads to off-by-one errors later.)
Set of positions of all occurrences of $W$ in $X$:
>$\{S(k):\underline{R}(W) <= k <= \overline{R}(W)\}$
Is W a substring of X?
> If $\underline{R}(W) > \overline{R}(W)$ then $W$ is not a substring of $X$.
> If $\underline{R}(W) = \overline{R}(W)$ then $W$ matches exactly one BWT entry.
> If $\underline{R}(W) < \overline{R}(W)$ then $W$ matches all BWT entries between (inclusive).
$SA$ interval $= [ \underline{R}(W), \overline{R}(W) ]$
Backward search in $O(|W|)$ time:
>$C(\alpha) =$ # of symbols in $X[0,n-1)$ (exclusive!) that are lexicographically smaller than $\alpha$
>$O(\alpha,i) =$ # of occurrences of $\alpha$ in $B[0,i]$ (inclusive!)
By Spiral's definition:
>$O(\alpha, -1) = 0$
If $W$ is a substring of $X$:
>$\underline{R}(\alpha{W}) = C(\alpha) + O(\alpha,\underline{R}(W)-1) + 1$
>$\overline{R}(\alpha{W}) = C(\alpha) + O(\alpha, \overline{R}(W))$
```
# For string X
X = "ATTGCTAC$"
# Calculate all suffixes
suffixes = sorted([X[i:] for i in range(len(X))])
print "# suffix"
for i, suffix in enumerate(suffixes):
print "{i} {suffix}".format(i=i, suffix=suffix)
# Calculate S
S = []
for suffix in suffixes:
S.append(X.find(suffix))
print S
# C(a) = # of symbols in X[0,n−1) that are lexicographically smaller than a.
# Precalculate the C(a) table. This lets us look up C(a) without knowing B.
Ca = {}
# all unique symbols in X except for $
symbols = ''.join(sorted(list(set(X)))[1:])
for symbol in symbols:
print symbol + ': ' + str([x for x in X[:-1] if x < symbol])
Ca[symbol] = len([x for x in X[:-1] if x < symbol])
print '\n', Ca
# B: X[S(i)-1]
def B(i):
return X[S[i]-1]
# n == |X| == |B| == |S|
n = len(X)
# String representation of B
B_str = ''.join([B(i) for i in range(n)])
print B_str
print n
# O(a,i): number of occurrences of a in B up to index i (inclusive)
def O(a, i):
if i < 0:
return 0 # O(a, -1) == 0
count = 0
for base in B_str[:i+1]:
if base == a:
count += 1
return count
# r underbar: first suffix that matches W (silly linear search)
def r(w):
if not w:
return 0 # r('') == 0
for i, suffix in enumerate(suffixes):
if w == suffix[:len(w)]:
return i
return n
# R overbar: last suffix that matches W (silly linear search)
def R(w):
if not w:
return n - 1 # R('') = n - 1
for i, suffix in enumerate(suffixes[::-1]):
if w == suffix[:len(w)]:
return n - i - 1
return 1
# SA value: compute [i,j] for W
def SA(w):
return [r(w), R(w)]
# Let's find SA values for some substrings
print "# suffix"
for i, suffix in enumerate(suffixes):
print "{i} {suffix}".format(i=i, suffix=suffix)
print "\nB = " + B_str + "\n"
for symbol in symbols:
print symbol + ':', SA(symbol)
queries = [
'GCT', # i == j, exactly one match
'GC', # i == j, exactly one match
'GA', # i > j, not in X
'T', # i < j, more than one match
'', # empty string, full range
]
for q in queries:
print "SA('" + q + "') = " + str(SA(q))
# Calculate bitcounts, saving start entry of each base
from copy import deepcopy
bitcounts = [{'A':0, 'C':0, 'G':0, 'T':0}]
for i, f in enumerate(S):
prev = deepcopy(bitcounts[i])
this = {}
for base in "ACGT":
this[base] = prev[base]
if(f):
base = X[f - 1]
this[base] = this[base] + 1
bitcounts.append(this)
# Drop the placeholder bitcounts[0] row
bitcounts = bitcounts[1:]
print "Bitcounts:\n"
for i, b in enumerate(bitcounts):
print "{i} {b}".format(i=i, b=b)
```
A little bit of bit math
------------------------
The whole point of this exercise is to quickly find the position(s) in $X$ where $W$ is an exact match (if any). One more simplification lets us calculate $O(\alpha, i)$ with bitcount lookups:
>$\underline{R}(\alpha{W}) = C(\alpha) + O(\alpha,\underline{R}(W)-1) + 1$
>$O(\alpha,\underline{R}(W)-1) = \underline{R}(\alpha{W}) - C(\alpha) - 1$
If $\underline{R}(\alpha{W})$ and $C(\alpha)$ are known, then $\underline{R}(W)-1$ becomes a lookup in the bitcount table.
```
# fast_O(a,i): lookup r(W) in the bitcounts table
def fast_O(a, i):
if i < 0:
return 0
return bitcounts[i][a]
# The two methods are equivalent
for a in symbols:
for i in range(n):
if O(a, i) != fast_O(a, i):
raise RuntimeError("O({0},{1}) {2} != {3}".format(a, i, O(a, i), fast_O(a, i)))
print "O(a,i) == fast_O(a,i)"
# r underbar: lower limit of substring W in BWT
# by Spiral's definition, r('$') == 0
r_cache = {'': 0}
def fast_r(w):
# Precache all substrings. We're gonna need them.
for aW in [w[i:] for i in range(len(w))][::-1]:
if(not aW in r_cache):
a = aW[0]
W = aW[1:]
r_cache[aW] = Ca[a] + fast_O(a, fast_r(W) - 1) + 1
return r_cache[w]
# R overbar: upper limit of substring W in BWT
# by definition, $('$') == n - 1
R_cache = {'': n - 1}
def fast_R(w):
for aW in [w[i:] for i in range(len(w))][::-1]:
if(not aW in R_cache):
a = aW[0]
W = aW[1:]
R_cache[aW] = Ca[a] + fast_O(a, fast_R(W))
return R_cache[w]
# SA value: compute [i,j] for W
def fast_SA(w):
return [fast_r(w), fast_R(w)]
print "# suffix"
for i, suffix in enumerate(suffixes):
print "{i} {suffix}".format(i=i, suffix=suffix)
print
for symbol in symbols:
print symbol + ':', SA(symbol), fast_SA(symbol)
print
queries = [
'GCT', # i == j, exactly one match
'GA', # i > j, not in X
'T', # i < j, more than one match
'', # empty string, full range
]
for q in queries:
print "SA('" + q + "') = " + str(SA(q)) + ' ' + str(fast_SA(q))
# Century table: getting back to X
# Keep a position entry every (mod) positions in the original sequence
mod = 3
# Also track the cumulative count of century bits (a la bitcount)
centcount = [0]
print "Century bits:\n"
print "# c o suffix"
for i, s in enumerate(suffixes):
centbit = 0 if S[i]%mod else 1
centcount.append(centcount[-1] + centbit)
print "{i} {m} {o} {s}".format(i=i, m=centbit, o=centcount[i], s=s)
century = []
for i, f in enumerate(S):
if not S[i]%mod:
century.append(f)
print "\nCentury table:\n"
print "# pos"
for i, c in enumerate(century):
print "{i} {c}".format(i=i, c=c)
def w_to_x(W):
(i, j) = fast_SA(W)
reply = []
for k in range(i, j + 1):
e = k # e is the bwt entry under examination
w = W # we will be pushing bases to the front of w
d = 0 # distance from century entry (# of pushes)
# No need to store the full century table: if centcount goes up on the next entry,
# then this is a century entry.
while centcount[e + 1] - centcount[e] == 0:
w = B_str[e] + w
e = fast_r(w)
d += 1
reply.append(century[centcount[e]] + d)
return sorted(reply)
def find_me(W):
print '{}:'.format(W)
for pos in w_to_x(W):
if W != X[pos:pos + len(W)]:
raise RuntimeError("X[{}] {} != {}".format(pos, X[pos:pos + len(W)], W))
print " X[{}] == {}".format(pos, X[pos:pos + len(W)])
print "X = {}\n".format(X)
print 'All symbols:'
for base in symbols:
find_me(base)
print
queries = [
'GCT',
'AAA',
'TGCTAC',
'ATT',
'TGCTA',
'CTA',
'CTTAGGAGAAC',
'AC'
]
print 'Canned lookups:'
for q in queries:
find_me(q)
print
def revcomp(seq):
flip = {
'A': 'T',
'C': 'G',
'G': 'C',
'T': 'A',
'N': 'N',
'a': 't',
'c': 'g',
'g': 'c',
't': 'a',
'n': 'n',
'$': ''
}
reply = ''
for c in seq:
reply += flip[c]
# Got a $? Leave a $.
if c == '$':
reply = '$' + reply
return reply[::-1]
def find_me_twice(W):
print 'Query {}:'.format(W)
for pos in w_to_x(W):
if W != X[pos:pos + len(W)]:
raise RuntimeError("X[{}] {} != {}".format(pos, X[pos:pos + len(W)], W))
print " X[{}] >> {}".format(pos, X[pos:pos + len(W)])
for pos in w_to_x(revcomp(W)):
if revcomp(W) != X[pos:pos + len(W)]:
raise RuntimeError("X[{}] {} != {}".format(pos, revcomp(W), X[pos:pos + len(W)]))
rpos = n - 1 - pos - len(W) # Position in reverse complement(X)
print " R[{}] == X[{}] << {}".format(rpos, pos, revcomp(X)[rpos:rpos + len(W)])
print X
print revcomp(X)
print
find_me_twice('T')
from random import randrange, choice
runs = 10
print '{} random substring lookups:'.format(runs)
for i in range(runs):
r = randrange(0, n - 2)
cache = []
q = 'x'
while q not in cache:
q = X[r:randrange(r + 1, n - 1)]
cache.append(q)
find_me_twice(q)
print '\n{} random 2-mer lookups:'.format(runs)
for i in range(runs):
cache = []
q = 'x'
while q not in cache:
q = choice(symbols) + choice(symbols)
cache.append(q)
find_me_twice(q)
print '\n{} random 3-mer lookups:'.format(runs)
for i in range(runs):
cache = []
q = 'x'
while q not in cache:
q = choice(symbols) + choice(symbols) + choice(symbols)
cache.append(q)
find_me_twice(q)
```
| github_jupyter |

# Semantic Types
Copyright (c) Microsoft Corporation. All rights reserved.<br>
Licensed under the MIT License.
Some string values can be recognized as semantic types. For example, email addresses, US zip codes or IP addresses have specific formats that can be recognized, and then split in specific ways.
When getting a DataProfile you can optionally ask to collect counts of values recognized as semantic types. [`Dataflow.get_profile()`](./data-profile.ipynb) executes the Dataflow, calculates profile information, and returns a newly constructed DataProfile. Semantic type counts can be included in the data profile by calling `get_profile` with the `include_stype_counts` argument set to true.
The `stype_counts` property of the DataProfile will then include entries for columns where some semantic types were recognized for some values.
```
import azureml.dataprep as dprep
dflow = dprep.read_json(path='../data/json.json')
profile = dflow.get_profile(include_stype_counts=True)
print("row count: " + str(profile.row_count))
profile.stype_counts
```
To see all the supported semantic types, you can examine the `SType` enumeration. More types will be added over time.
```
[t.name for t in dprep.SType]
```
You can filter the found semantic types down to just those where all non-empty values matched. The `DataProfile.stype_counts` gives a list of semantic type counts for each column, where at least some matches were found. Those lists are in desecending order of count, so here we consider only the first in each list, as that will be the one with the highest count of values that match.
In this example, the column `inspections.business.postal_code` looks to be a US zip code.
```
stypes_counts = profile.stype_counts
all_match = [
(column, stypes_counts[column][0].stype)
for column in stypes_counts
if profile.row_count - profile.columns[column].empty_count == stypes_counts[column][0].count
]
all_match
```
You can use semantic types to compute new columns. The new columns are the values split up into elements, or canonicalized.
Here we reduce our data down to just the `postal` column so we can better see what a `split_stype` operation can do.
```
dflow_postal = dflow.keep_columns(['inspections.business.postal_code']).rename_columns({'inspections.business.postal_code': 'postal'})
dflow_postal.head(5)
```
With `SType.ZipCode`, values are split into their basic five digit zip code and the plus-four add-on of the Zip+4 format.
```
dflow_split = dflow_postal.split_stype('postal', dprep.SType.ZIPCODE)
dflow_split.head(5)
```
`split_stype` also allows you to specify the fields of the stype to use and the name of the new columns. For example, if you just needed to strip the plus four from our zip codes, you could use this.
```
dflow_no_plus4 = dflow_postal.split_stype('postal', dprep.SType.ZIPCODE, ['zip'], ['zipNoPlus4'])
dflow_no_plus4.head(5)
```
| github_jupyter |
## Learning Objectives:
By the end of this lesson, students will be able to:
- Describe an outlier and how to remove outliers for univariate data
- Visualize the outliers through box-plots
- Describe how to use a Gaussian Mixture Model to fit a data distribution
- Understand correlation and visualize the correlation amongst features
## Outlier Detection
- **Outliers** are extreme values that can skew our dataset, sometimes giving us an incorrect picture of how things actually are in our dataset.
- The hardest part of this is determining which data points are acceptable, and which ones constitute as "outliers".
## Activity: find and remove outliers if our dataset is Normal
When our sample data is close to normal distribution, the samples that are _three standard deviations away from the mean_ can be considered outliers.
**Task**: Write a function that first finds outliers for a normally distributed data, then remove them.
**Hint**: Data samples that are below `mean - 3*std` and above `mean + 3*std` are outliers for Normal distribution
```
import numpy as np
def find_remove_outlier(data_sample):
# calculate summary statistics
data_mean, data_std = np.mean(data), np.std(data)
# define cut-off
cut_off = data_std * 3
lower, upper = data_mean - cut_off, data_mean + cut_off
# identify outliers
outliers = [x for x in data_sample if x < lower or x > upper]
# remove outliers
outliers_removed = [x for x in data_sample if x > lower and x < upper]
return outliers, outliers_removed
```
## Interquartile range (IQR)
We use IQR for finding and removing outliers when the data has any kind of distribution
[John Tukey](https://en.wikipedia.org/wiki/John_Tukey) suggested to _calculate the range between the first quartile (25%) and third quartile (75%) in the data_, which is the IQR.
## Activity: IQR outlier detection and removal
**Task**: write a function to find and remove outliers based on IQR method for this data sample:
**Hint**:
$Q_1$ is the first quartile (25%)
$Q_3$ is the third quartile (75%)
<img src="Images/iqr.png">
`x = [norm.rvs(loc=5 , scale=1 , size=100), -5, 11, 14]`
```
import numpy as np
import scipy.stats
def find_remove_outlier_iqr(data_sample):
# calculate interquartile range
q25, q75 = np.percentile(data_sample, 25), np.percentile(data_sample, 75)
iqr = q75 - q25
print(iqr)
# calculate the outlier cutoff
cut_off = iqr * 1.5
lower, upper = q25 - cut_off, q75 + cut_off
# identify outliers
outliers = [x for x in data_sample if x < lower or x > upper]
# remove outliers
outliers_removed = [x for x in data_sample if x > lower and x < upper]
return outliers
y = np.array([-5, 11, 14])
x = np.concatenate((scipy.stats.norm.rvs(loc=5 , scale=1 , size=100), y))
print(type(x))
print(find_remove_outlier_iqr(x))
print(scipy.stats.iqr(x))
```
## How we can visualy see the outlier?
**Answer**: Box plot use the IQR method to display data and outliers
```
import matplotlib.pyplot as plt
plt.boxplot(x)
plt.show()
```
## Correlation
**Correlation** is used to _test relationships between quantitative variables_
Some examples of data that have a high correlation:
1. Your caloric intake and your weight
1. The amount of time your study and your GPA
**Question**: what is negative correlation?
Correlations are useful because we can find out what relationship variables have, we can make predictions about future behavior.
## Activity: Obtain the correlation among all features of iris dataset
1. Review the iris dataset. What are the features?
1. Eliminate two columns `['Id', 'Species']`
1. Compute the correlation among all features.
1. **Hint**: Use `df.corr()`
1. Plot the correlation by heatmap and corr plot in Seaborn -> `sns.heatmap`, `sns.corrplot`
1. Write a function that computes the correlation (Pearson formula)
1. **Hint**: https://en.wikipedia.org/wiki/Pearson_correlation_coefficient
1. Compare your answer with `scipy.stats.pearsonr` for any given two features
<img src="./Images/iris_vis.jpg">
```
import pandas as pd
import numpy as np
import scipy.stats
import seaborn as sns
import scipy.stats
df = pd.read_csv('Iris.csv')
df = df.drop(columns=['Id', 'Species'])
sns.heatmap(df.corr(), annot=True)
def pearson_corr(x, y):
x_mean = np.mean(x)
y_mean = np.mean(y)
num = [(i - x_mean)*(j - y_mean) for i,j in zip(x,y)]
den_1 = [(i - x_mean)**2 for i in x]
den_2 = [(j - y_mean)**2 for j in y]
correlation_x_y = np.sum(num)/np.sqrt(np.sum(den_1))/np.sqrt(np.sum(den_2))
return correlation_x_y
print(pearson_corr(df['SepalLengthCm'], df['PetalLengthCm']))
print(scipy.stats.pearsonr(df['SepalLengthCm'], df['PetalLengthCm']))
```
## Statistical Analysis
We can approxite the histogram of a dataset with a combination of Gaussian (Normal) distribution functions:
Gaussian Mixture Model (GMM)
Kernel Density Estimation (KDE)
## What is the goal for using GMM?
- We want to appriximate the density (histogram) of any given data sample with a combination of Normal Distribitions
- How many Normal Distributions we need is defined by us (2 or 3 or 4,...)
## Activity : Fit a GMM to a given data sample
Task:
1. Generate the concatination of the random variables as follows:
`x_1 = np.random.normal(-5, 1, 3000)`
`x_2 = np.random.normal(2, 3, 7000) `
`x = np.concatenate((x_1, x_2))`
2. Plot the histogram of `x`
3. Obtain the weights, mean and variances of each Gassuian
You will need to use the following in your solution:
```python
from sklearn import mixture
gmm = mixture.GaussianMixture(n_components=2)
gmm.fit(x.reshape(-1,1))
```
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn import mixture
# Generate data samples and plot its histogram
x_1 = np.random.normal(-5, 1, 3000)
x_2 = np.random.normal(2, 3, 7000)
x = np.concatenate((x_1, x_2))
plt.hist(x, bins=20, density=1)
plt.show()
# Define a GMM model and obtain its parameters
gmm = mixture.GaussianMixture(n_components=2)
gmm.fit(x.reshape(-1,1))
print(gmm.means_)
print(gmm.covariances_)
print(gmm.weights_)
```
## The GMM has learned the probability density function of our data sample
Let's generate the samples from the GMM:
```
z = gmm.sample(10000)
plt.hist(z[0], bins=20, density=1)
plt.show()
```
## Question: Are the samples in z and x the same?
- Not their samples, but we know that the _pdf of x and the pdf of z are the same_
## Kernel Density Estimation (KDE)
**Kernel density estimation (KDE)** is a non-parametric way to estimate the Probability Density Function (PDF) of a data sample. In other words _the goal of KDE is to find PDF for a given data sample._
We can use the below formula to approximate the PDF of the dataset:
$p(x) = \frac{1}{Nh}\sum_{i = 1}^{N} \ K(\frac{x - x_i}{h})$
where $h$ is a bandwidth and $N$ is the number of data points
## Activity: Apply KDE on a given data sample
**Task**: Apply KDE on the previously generated sample data `x`
**Hint**: use
`kde = KernelDensity(kernel='gaussian', bandwidth=0.6)`
```
from sklearn.neighbors import KernelDensity
kde = KernelDensity(kernel='gaussian', bandwidth=0.6)
kde.fit(x.reshape(-1,1))
s = np.linspace(np.min(x), np.max(x))
log_pdf = kde.score_samples(s.reshape(-1,1))
plt.plot(s, np.exp(log_pdf))
```
## The KDE has learned the probability density function of our data sample
Let's generate the samples from the KDE:
```
m = kde.sample(10000)
plt.hist(m, bins=20, density=1)
plt.show()
```
## KDE can learn handwitten digits distribution and generate new digits
http://scikit-learn.org/stable/auto_examples/neighbors/plot_digits_kde_sampling.html
| github_jupyter |
# Using plaidml to understand performance gains for a convolutional architecture
### 1. Import the libraries
```
import plaidml.keras
plaidml.keras.install_backend()
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
from keras import backend as K
```
### 2. Adjusting the model config
```
batch_size = 128
num_classes = 10
epochs = 12
```
### 3. Data Preparation
```
# input image dimensions
img_rows, img_cols = 28, 28
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
if K.image_data_format() == 'channels_first':
x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
else:
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
# convert class vectors to binary class matrices
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
```
### 4. Set the model architecture
```
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=input_shape))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes, activation='softmax'))
```
### 5. Compile the model
```
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
```
### 6. Train the CNN model
#### 6.1 Using `cpu`
```
%%time
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
```
#### 6.2 Using `metal_amd_gpu`
```
# using plaidml backend
# fit the model
%timeit
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
```
### 7. Evaluate the model
```
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
```
| github_jupyter |
# 理解编程语言
简单地说,程序就是人类书写交由机器去执行的一系列指令,而编程语言是书写时遵照的语法规则。
编程很像教孩子,计算机就像一个孩子,一开始什么都不懂,也不会自己思考,但非常听话,你教TA怎样就怎样,一旦学会的就不会忘,可以不走样地做很多次不出错。但是如果教的不对,TA也会不折不扣照做。
对我们来说,和孩子的教育一样,关键在于理解对方的沟通语言,以及理解对方会如何理解你的表达。
## 计算机系统和 CPU
计算机本身并不复杂,大致上就是三部分:
* 负责执行指令的**中央处理器(CPU)**;
* 负责存放 CPU 执行所需要数据的**内存**;
* 各种**外设**,像硬盘、键盘、鼠标、显示设备、网卡等等这些都是。
这里面 CPU 是核心的核心,因为它相当于人的大脑,我们教给计算机的指令实际上就是 CPU 在一条一条地执行,执行过程中会读写内存里的数据,会调用各种外设的接口(通过一种叫做“设备驱动”的软件模块)来完成数据的输入输出(I/O)操作。
CPU 是一种计算机芯片,芯片可以执行的指令是在设计时就定好的, 不同芯片有不同的指令集,固化在硬件里,一般无法改变。因为大小、发热和功耗等问题,芯片的指令集不能面面俱到,而是有所侧重,比如一般计算机里 CPU 芯片比较擅长整数的运算,而 GPU(图形处理芯片,也就是显卡)比较擅长浮点数的运算,计算机图形处理尤其是 3D 图形处理会涉及大量的浮点运算,所以通常交给 GPU 去做,又快又省电;后来人们发现除了 3D 图形处理,人工智能等领域的一些科学计算也多是浮点运算,所以 GPU 也被拿来做这些计算工作。而我们的手机里的芯片,是把 CPU/GPU 还有其他一些做特殊工作的芯片集成在一块硅晶片上,这样可以节省空间,也能节约功耗,这种高度集成的芯片就叫 SoC(*System on Chip*)。
## 汇编和编译器
那么 CPU 执行的代码长啥样呢,我们可以瞄一眼,大致长这样:
```nasm
_add: ## @add
push rbp
mov rbp, rsp
mov dword ptr [rbp - 4], edi
mov dword ptr [rbp - 8], esi
mov esi, dword ptr [rbp - 4]
add esi, dword ptr [rbp - 8]
mov eax, esi
pop rbp
ret
_main: ## @main
push rbp
mov rbp, rsp
sub rsp, 16
mov dword ptr [rbp - 4], 0
mov edi, 27
mov esi, 15
call _add
add rsp, 16
pop rbp
ret
```
这叫汇编指令(*assembly*),和 CPU 实际执行的“机器指令(*machine code*)”几乎一样了,只是机器语言是二进制的,全是 01011001 这样,而汇编将其翻译成了我们稍微看得懂的一些指令和名字。机器可以直接执行的指令其实不多,在 Intel 官方的 [x64 汇编文档](https://software.intel.com/en-us/articles/introduction-to-x64-assembly) 里列出的常用指令码只有 20 多个,上图中的 `mov` `push` `pop` `add` `sub` `call` `ret` 等都是。我们目前并不需要搞懂这些都是干啥的,不过可以简单说一下,`mov` 就是把一个地方保存的数写到另一个地方,`call` 是调用另外一个代码段,`ret` 是返回 `call` 的地方继续执行。
所以其实计算机执行的指令大致就是:
* 把数放到某地方,可能在 CPU 内部也可能在内存里;
* 对某地方的数进行特定运算(通常是加减乘除);
* 跳转到另一个地方执行一段指令;
* 返回原来的位置继续执行。
如此这般。
可以想象,如果要完成复杂的任务,涉及复杂的逻辑流程,用汇编代码写起来可要费好大的劲儿了。但这就是机器的语言,CPU 实际上就只会这种语言,别的它都听不懂,怎么办?
想来想去,是不是有可能我们用某种方式写下我们解决问题的方法,然后让计算机把这些方法翻译成上面那种计算机懂的指令呢?这种翻译的程序肯定不好写,但是一旦写好了以后所有人都可以用更接近人话的方式表达,计算机也能自己翻译给自己并且照做了,所谓辛苦一时享受一世,岂不美哉。程序员的典型思维就是这样:如果有个对人来说很麻烦的事情,就要试试看是不是可以让计算机来做。
这个想法早已被证明完全可行,这样的翻译程序叫做“编译器(*compiler*)”。现在存在于世的编程语言有数百种,每一种都对应一个编译器,它可以读懂你用这语言写的程序,经过词法分析(*tokenizing*)、语法分析(*parsing*)、语义分析(*semantic analysis*)、优化(*optimization*)和代码生成(*code emission*)等环节,最终输出像上面那样机器可以看懂和执行的指令。
编译理论和实现架构已经发展了很多年,已经非常成熟,并在不断优化中。只要愿意学习,任何人都可以设计自己的编程语言并为之实现一个编译器。
顺便,上面的汇编代码是如下的 C 语言代码通过 LLVM/Clang 编译器的处理生成的:
```c
int add(int a, int b) {
return a + b;
}
int main() {
return add(27, 15);
}
```
C 语言被公认是最接近机器语言的“高级编程语言”,因为 C 语言对内存数据的管理方式几乎和机器语言是一样的“手工操作”,需要编程者非常了解 CPU 对内存的管理模式。但尽管如此,C 语言还是高级编程语言,很接近我们人类的语言,基本上一看就懂。
各种各样的高级编程语言,总有一样适合你,这些语言极大地降低了计算机编程的门槛,让几乎所有人都有机会编程,而这一切都是因为有编译器这样的“自动翻译”存在,建立了人类和机器之间畅通的桥梁。
## 解释器和解释运行
前面我们介绍了编程语言编译为机器代码的原理,我们写好的程序可以被编译器翻译为机器代码,然后被机器运行。编译就好像把我们写好的文章整篇读完然后翻译为另一种语言,拿给会那个语言的人看。
还有另外一种运行程序的方法,不需要整篇文章写完,可以一句一句的翻译,写一句翻一句执行一句,做这件事的也是一个程序,叫解释器(*interpreter*)。解释器和编译器的主要区别就在于,它读入源代码是一行一行处理的,读一行执行一行。
解释器的好处是,可以实现一种交互式的编程,启动解释器之后,你说一句解释器就回你一句,有来有回,很亲切,出了问题也可以马上知道。现代人几乎不写信了,打电话和微信是主要的沟通模式,可能也是类似的道理吧。
Python 就是一种解释型语言,在命令行界面运行 `python` 主程序,就会进入一个交互式的界面,输入一行程序立刻可以得到反馈,差不多就是这个样子:
<img src="assets/python-repl.png" width="500">
这就是 Python 的解释器界面,这种输入一行执行一行的界面有个通用的名字叫做 *REPL*(*read–eval–print loop* 的缩写),意思是这个程序可以读取(*read*)你的输入、计算(*evaluate*)、然后打印(*print*)结果,循环往复,直到你退出——在上图的界面中,输入 `exit()` 回车,就可以退出 Python 的 *REPL*。
我们还可以把 Python 程序源代码(*source code*)存进一个文件里,然后让解释器直接运行这个文件。打开命令行界面(我们假定你已经[设置好了你的编程环境](x1-setup.md)),执行下面的操作:
```shell
cd ~/Code
touch hello.py
code hello.py
```
在打开的 VSCode 中输入下面的代码:
```python
print(f'4 + 9 = {4 + 9}')
print('Hello world!')
```
保存,回到命令行界面执行:
```python
python hello.py
```
解释器会读取 `hello.py` 里面的代码,一行一行的执行并输出对应的结果。
> 在有的系统中,`python` 不存在或者指向较老的 Python,可以通过 `python -V` 命令来查看其版本,如果返回 2.x 的话,可以将上述命令改为 `python3 hello.py`。
解释器的实现方式有好几种:
* 分析源代码,直接执行(做某些解释器已经定义好的事情),或者
* 把源代码翻译成某种特定的中间代码,然后交给一个专门运行这种中间代码的程序执行,这种程序通常叫虚拟机(*virtual machine*),或者
* 把源代码编译为机器代码然后交给硬件执行。
无论哪种方式,其实也会有词法分析、句法分析、语义分析、代码优化和生成等过程,和编译器的基本架构是相似的,事实上由于实现上的复杂性,有时候编译和解释之间并没有那么硬的界限。
Python 官方解释器的工作原理是上列的第二种,即生成中间代码——叫做“字节码(*bytecode*)”——和虚拟机的方式;而另外有种 Python 的实现([PyPy](http://pypy.org/))采用的是第三种方式,即预编译为机器码的方式。
在这本书里,绝大部分代码运行用的是 Jupyter Lab,它的目标是把 *REPL* 做的更好用,让我们可以在浏览器中使用,有更好的语法高亮显示,最棒的是,可以把我们交互编程的过程保存下来,与人共享。不过它背后还是 Python 解释器,和命令行界面下运行 `python` 没有本质区别(但用起来的愉快程度和效率还是有很大区别的)。
> Jupyter Lab 通过一个可扩展的架构正在支持越来越多的文档类型和编程语言,以后学习很多别的语言也可以用它。
要使用 Jupyter Lab 需要先用 Python 的包管理工具 `pip` 来安装:
```shell
pip install jupyterlab
```
然后就可以用 `jupyter lab` 来运行 Jupyter Lab,在里面打开 *notebook* 进行交互式编程的尝试了,最好的办法是[使用我们的学习用书](x2-students-book.md)。
## 小结
* 程序是人类书写交由机器去执行的一系列指令,而编程语言是书写时遵照的语法规则;
* 计算机 CPU 只能理解非常基础的指令,人要直接写出机器指令是困难和低效的;
* 编译器和解释器是一些特殊的程序,能够把我们用高级编程语言编写的程序翻译成机器指令,然后交给计算机去执行;
* 可以通过命令行 REPL、命令行加源文件和 Jupyter Lab 这样的可视化环境来运行 Python 程序,背后都是 Python 解释器。
| github_jupyter |
# Data Generation with Flatland
This notebook begins to outline how training datasets will be generated in Flatland. This notebook will evolve into a documentation of how to use the tool through a more formal and simplified API.
### Here's the idea
Some modern protein structure prediction approaches are a little bit complicated to implement. That's why it seems useful to have a simulator that can generate data at least of the same structure that researchers seek to use in such systems. Over time, these simulators can be improved progressively to add some minimal level of realism that should be helpful for initial model debugging. This might include for example modifying simulation parameters to enable a model to train effectively then returning to the more complex form once the simpler problem has been solved. Thus we hope to create a much more smooth path to solving the larger problem than is often followed by those seeking to solve it directly.
Further, even when training larger systems on real data it will be important that system components remain integrated and both the system and its individual components continue to function correctly. Simple toy test cases are often used for this purpose in software test engineering. But in the case of ML software engineering, sometimes it helps if these are slightly realistic. Even further, we are interested in understanding the potential of various additional sources of data to enhance the performance of structure prediction systems.
The simulations performed below involve evolve populations of polymers using a trivially simple fitness metric and in the course of that retain a "genetic history" of the evolved populations. Then, structures for these polymers are simulated using Jax MD. For each "solved" structure we compute a pairwise "residue" distance matrix and a vector of "bond" angles. Lastly, we simulate a compound-protein interaction experiment again using Jax MD.
All of this data is written to Google Cloud Storage in an organized fashion and can be read from there when generating training examples.
### Setup
Ensure the most recent version of Flatland in installed.
```
!pip install git+git://github.com/cayley-group/flatland.git --quiet
import pprint
from flatland import dataset
from flatland import evolution as evo
```
### Customize the dataset
In the interest of keeping things organized, the Flatland datasets are both generated from and readable via the same TensorFlow Datasets object. That includes the configuration used to perform the simulation as well as the name of the Google Cloud Storage bucket to which simulated data is written and from which it is read. Through this project we provide datasets that can be used without the need for users to run simulations. But if you're interested in using different parameters, for example, you'll need to sub-class or fork one of these core objects at least as shown below:
```
class MyFlatlandBase(dataset.FlatlandBase):
def simulation_config(self):
return evo.SimulationConfig(alphabet_size=3,
pop_size=2,
genome_length=10,
mutation_rate=0.15,
num_generations=2,
keep_full_population_history=True,
report_every=10)
def sim_bucket_name(self):
"""The remote bucket in which to store the simulated dataset.
This should be customized to be the name of a GCS bucket for
which you have write permissions.
"""
return "cg-flatland-test"
ds = MyFlatlandBase()
```
## Configuration
Here we'll configure our evolutionary simulations. We'll configure these to be very simple given that this is a demo - polymers of length 10 with elements from an alphabet of size 3. And just 10 population members for only 2 generations.
```
pprint.pprint(dict(ds.simulation_config()._asdict()))
```
## Run simulations
Now we'll evolve our polymer populations using a trivial fitness measure - i.e. how closely the average of the integer encodings of polymer elements come to 1.0! Here we could specify the fitness_fn to be one that simulates the polymers, computes their energy, and simply considered the simulated polymer energy to be a measure of fitness (energies are more negative for more energetically-stable strutures). Or likewise we could simulate the interaction of polymers with a set of compounds and define fitness as the selectivity of systems to be low energy only when including one or more target compounds.
#### Clearly specify the billable project
Our framework requires you clearly specify which project will be billed.
```
#%env FLATLAND_REQUESTER_PAYS_PROJECT_ID="<your project ID here>"
#!gcloud config list
```
#### Evolve the training population
In the future we could scale this up arbitrarily by doing as many of these simulations in parallel as we like and aggregating the result. This is analagous to simulating the independent evolution of polymer families that are evolutionarily distant.
```
ds.simulate_dataset(split="train", shard_id=0)
```
#### Evolve the test population
For now, evolve a single test polymer population. This will be analagous to the problem of inferring the structure of a polymer that is evolutionarily distant from any we have seen before.
In some sense, this kind of generalization may not be necessary - as solvers may do well to just memmorize solutions to the kinds of structures that are known to occur in nature. One way to interpret recent success using evolutionary information for folding is that it does exactly this - cues solvers regarding how to re-use previously-accumulated knowledge about how certain subsequences fold.
The benefit of such a test would be regarding completely novel polymers that are not homologous to anything currently known to occur in nature. Or at least which arise from a anciently-diverged part of the evolutionary tree from the one our model was trained on.
It would be feasible to construct a test set sharing a closer evolutionary history with the training populations by selecting polymers to hold out from these to use in testing - sharing alignments across both.
```
ds.simulate_dataset(split="test", shard_id=0)
```
#### Evolve the validation population
The same for the validation set as for the test set.
```
ds.simulate_dataset(split="validation", shard_id=0)
```
| github_jupyter |
# Generative Adversarial Network (GAN)

Пришло время поговорить о более интересных архитектурах, а именно о GANах или состязательных нейронных сетках. [Впервые GANы были предложены в 2014 году.](https://arxiv.org/abs/1406.2661) Сейчас они очень активно исследуются. GANы состоят из двух нейронных сетей:
* Первая - генератор порождает из некоторого заданного распределения случайные числа и собирает из них объекты, которые идут на вход второй сети.
* Вторая - дискриминатор получает на вход объекты из реальной выборки и объекты, созданные генератором. Она пытается определить какой объект был порождён генератором, а какой является реальным.
Таким образом генератор пытается создавать объекты, которые дискриминатор не сможет отличить от реальных.
```
import tensorflow as tf
print(tf.__version__)
import numpy as np
import time
import matplotlib.pyplot as plt
%matplotlib inline
```
# 1. Данные
Для начала давайте попробуем погонять модели на рукописных цифрах из MNIST как бы скучно это не было.
```
(X, _ ), (_, _) = tf.keras.datasets.mnist.load_data()
X = X/127.5 - 1 # отнормировали данные на отрезок [-1, 1]
X.min(), X.max() # проверили нормировку
X = X[:,:,:,np.newaxis]
X.shape
```
Давайте вытащим несколько рандомных картинок и нарисуем их.
```
cols = 8
rows = 2
fig = plt.figure(figsize=(2 * cols - 1, 2.5 * rows - 1))
for i in range(cols):
for j in range(rows):
random_index = np.random.randint(0, X.shape[0])
ax = fig.add_subplot(rows, cols, i * rows + j + 1)
ax.grid(False)
ax.axis('off')
ax.imshow(np.squeeze(X,-1)[random_index, :], cmap='gray')
plt.show()
```
Соберём для наших данных удобный генератор.
# 2. Дискриминатор
* Дискриминатор - это обычная свёрточная сетка
* Цель этой сетки - отличать сгенерированные изображения от реальных
```
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras import layers as L
```
Должны собрать просто сверточную сетку (discriminator)
1) 16 свертки (3,3) без потери размерности и активацией elu (помним, что это)
2) Пулинг
3) 32 свертки (3,3) без потери размерности и активацией elu (помним, что это)
4) Пулинг
5) слой полносвязный на 128 и выходной
# 3. Генератор
* Генерирует из шума изображения
Будем генерировать новые цифры из шума размера `code_size`.
```
CODE_SIZE = 100
generator = Sequential()
generator.add(L.InputLayer([CODE_SIZE],name='noise'))
generator.add(L.Dense(128*7*7, activation='elu'))
generator.add(L.Reshape((7,7,128)))
generator.add(L.Conv2DTranspose(128, kernel_size=(3,3)))
generator.add(L.LeakyReLU())
generator.add(L.Conv2DTranspose(64, kernel_size=(3,3)))
generator.add(L.LeakyReLU())
generator.add(L.UpSampling2D(size=(2,2)))
generator.add(L.Conv2DTranspose(32,kernel_size=3,activation='elu'))
generator.add(L.Conv2DTranspose(32,kernel_size=3,activation='elu'))
generator.add(L.Conv2DTranspose(32,kernel_size=3,activation='elu'))
generator.add(L.Conv2D(1, kernel_size=3, padding='same'))
# Как проверить то, что мы нигде не ошиблись в размерах?
# Как вообще это подбирать?
print('Выход генератора: ', )
```
Посмотрим на пример, который нам генерирует на выход наша свежая нейронка!
```
noise = tf.random.normal([1, CODE_SIZE])
generated_image = generator(noise)
plt.imshow(generated_image[0, :, :, 0], cmap='gray');
```
Хммм... А что про это всё думает дескриминатор?
```
decision = discriminator(generated_image)
# на выход из дискриминатора мы забираем логарифм!
decision
```
# 4. Функция потерь
Потери для дескриминатора это обычныя кросс-энтропия.
```
cross_entropy = tf.keras.losses.BinaryCrossentropy(from_logits=True)
def discriminator_loss(real_output, fake_output):
real_loss = cross_entropy(tf.ones_like(real_output)+0.05 * np.random.random(real_output.shape),
real_output)
fake_loss = cross_entropy(tf.zeros_like(fake_output)+0.05 * np.random.random(fake_output.shape),
fake_output)
total_loss = real_loss + fake_loss
return total_loss
real_output = discriminator(X[0:10])
fake_output = discriminator(generated_image)
discriminator_loss(real_output, fake_output)
```
Для генератора мы хотим максимизировать ошибку дискриминатора на фэйковых примерах.
```
def generator_loss(fake_output):
return cross_entropy(tf.ones_like(fake_output), fake_output)
generator_loss(fake_output)
```
# 5. Градиентный спуск
Учить пару из сеток будем так:
* Делаем $k$ шагов обучения дискриминатора. Целевая переменная - реальный объект перед нами или порождённый. Веса изменяем стандартно, пытаясь уменьшить кросс-энтропию.
* Делаем $m$ шагов обучения генератора. Веса внутри сетки меняем так, чтобы увеличить логарифм вероятности дискриминатора присвоить сгенерированному объекту лэйбл реального.
* Обучаем итеративно до тех пор, пока дискриминатор больше не сможет найти разницу (либо пока у нас не закончится терпение).
* При обучении может возникнуть огромное количество пробем от взрыва весов до более тонких вещей. Имеет смысл посмотреть на разные трюки, используемые при обучении: https://github.com/soumith/ganhacks
Собираем структуру для обучения.
```
generator_optimizer = tf.keras.optimizers.Adam(1e-4)
discriminator_optimizer = tf.keras.optimizers.Adam(1e-4)
```
Чекпойнты для процесса обучения.
```
import os
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
```
Задаём один шаг процедуры обучения генератора.
```
@tf.function
def train_generator_step(images, noise):
# ищем градиенты
with tf.GradientTape() as gen_tape:
# сгенерировали новое изображение из шума
generated_images = ...
# посчитали прогнозы дискриминатора
real_output = ...
fake_output = ...
# нашли ошибку
gen_loss = ...
# нашли градиенты
grad = ...
# сделали шаг градиентного спуска
generator_optimizer.apply_gradients(zip(grad, generator.trainable_variables))
```
Теперь шаг обучения дискриминатора.
```
@tf.function
def train_discriminator_step(images, noise):
# ищем градиенты
with tf.GradientTape() as disc_tape:
# сгенерировали новое изображение из шума
generated_images = ...
# посчитали прогнозы дискриминатора
real_output = ...
fake_output = ...
# нашли ошибку
disc_loss = ...
# нашли градиенты
grad = ...
# сделали шаг градиентного спуска
discriminator_optimizer.apply_gradients(zip(grad, discriminator.trainable_variables))
```
Мы почти готовы учить нашу сетку. Напишем две простенькие функции для генерации фэйковых и настоящих батчей.
```
# функция, которая генерирует батч с шумом
def sample_noise_batch(bsize):
return tf.random.normal([bsize, CODE_SIZE], dtype=tf.float32)
# функция, которая генерирует батч из реальных данных (для баловства)
def sample_data_batch(bsize):
idxs = np.random.choice(np.arange(X.shape[0]), size=bsize)
return X[idxs]
```
Проверяем отрабатывают ли наши шаги.
```
data_test = sample_data_batch(256)
fake_test = sample_noise_batch(256)
gen_log = discriminator(generator(fake_test))
real_log = discriminator(data_test)
print('Ошибка дескриминатора:', discriminator_loss(real_log, gen_log).numpy())
print('Ошибка генератора:', generator_loss(gen_log).numpy())
# сделали шаг работы генератора
train_generator_step(data_test, fake_test)
gen_log = discriminator(generator(fake_test))
real_log = discriminator(data_test)
print('Ошибка дескриминатора:', discriminator_loss(real_log, gen_log).numpy())
print('Ошибка генератора:', generator_loss(gen_log).numpy())
# сделали шаг работы дискриминатора
train_discriminator_step(data_test, fake_test)
gen_log = discriminator(generator(fake_test))
real_log = discriminator(data_test)
print('Ошибка дескриминатора:', discriminator_loss(real_log, gen_log).numpy())
print('Ошибка генератора:', generator_loss(gen_log).numpy())
```
Как думаете, выглядит адекватно? Мы нигде не ошиблись?
Напишем пару вспомогательных функций для отрисовки картинок.
```
# рисуем изображения
def sample_images(rows, cols, num=0):
images = generator.predict(sample_noise_batch(bsize=rows*cols))
fig = plt.figure(figsize=(2 * cols - 1, 2.5 * rows - 1))
for i in range(cols):
for j in range(rows):
ax = fig.add_subplot(rows, cols, i * rows + j + 1)
ax.grid('off')
ax.axis('off')
ax.imshow(np.squeeze(images[i * rows + j],-1),cmap='gray')
# сохраняем картинку для гифки
if num >0:
plt.savefig('images_gan/image_at_epoch_{:04d}.png'.format(num))
plt.show()
sample_images(2,7)
```
Немного побалуемся с шагами.
```
data_test = sample_data_batch(256)
fake_test = sample_noise_batch(256)
# Генератор
train_generator_step(data_test, fake_test)
gen_log = discriminator(generator(fake_test))
real_log = discriminator(data_test)
print('Ошибка дескриминатора:', discriminator_loss(real_log, gen_log).numpy())
print('Ошибка генератора:', generator_loss(gen_log).numpy())
sample_images(2,7)
data_test = sample_data_batch(256)
fake_test = sample_noise_batch(256)
# Дискриминатор
train_discriminator_step(data_test, fake_test)
gen_log = discriminator(generator(fake_test))
real_log = discriminator(data_test)
print('Ошибка дескриминатора:', discriminator_loss(real_log, gen_log).numpy())
print('Ошибка генератора:', generator_loss(gen_log).numpy())
```
# 6. Обучение
Ну и наконец последний шаг. Тренировка сеток. При обучении нужно соблюдать между сетками баланс. Важно сделать так, чтобы ни одна из них не стала сразу же побеждать. Иначе обучение остановится.
* Чтобы избежать моментального выигрыша дискриминатора, мы добавили в хитрую регуляризацию
* Кроме регуляризации можно пытаться учить модели сбалансированно, делая внутри цикла шаги чуть более умным способом.
```
# Эксперименты с совместным обучением (так цикл работает быстре)
@tf.function
def train_step(images, noise):
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
generated_images = generator(noise)
real_output = discriminator(images)
fake_output = discriminator(generated_images)
gen_loss = generator_loss(fake_output)
disc_loss = discriminator_loss(real_output, fake_output)
gradients_of_generator = gen_tape.gradient(gen_loss, generator.trainable_variables)
gradients_of_discriminator = disc_tape.gradient(disc_loss, discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(gradients_of_generator, generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(gradients_of_discriminator, discriminator.trainable_variables))
from IPython import display
EPOCHS = 300
BSIZE = 256
# время
start = time.time()/60
# вектора для мониторинга сходимости сеток
d_losses = [ ]
g_losses = [ ]
num = 0 # для сохранения картинок
# запускаем цикл обучения
for epoch in range(EPOCHS):
# генерируем батч
X_batch = sample_data_batch(BSIZE)
X_fake = sample_noise_batch(BSIZE)
train_step(X_batch, X_fake)
# делаем N шагов обучения дискриминатора
for i in range(5):
train_discriminator_step(X_batch, X_fake)
# делаем K шагов обучения генератора
for i in range(1):
train_generator_step(X_batch, X_fake)
gen_log = discriminator(generator(X_fake))
real_log = discriminator(X_batch)
d_losses.append(discriminator_loss(real_log, gen_log).numpy())
g_losses.append(generator_loss(gen_log).numpy())
# ну сколько можно ждааать!!!
if epoch % 20==0:
print('Time for epoch {} is {} min'.format(epoch + 1, time.time()/60-start))
print('error D: {}, error G: {}'.format(d_losses[-1], g_losses[-1]))
if epoch % 20==0:
# сохраняем модель и обновляем картинку
# checkpoint.save(file_prefix = checkpoint_prefix)
# можно раскоментировать, если хочется, чтобы картинка обновлялась, а не дополнялас
#display.clear_output(wait=True)
num += 1
sample_images(2,7, num)
# sample_probas(X_batch)
```
Тренируем сетки.
```
# сетка тренировалась много итераций
sample_images(4,8)
# смотрим сошлись ли потери
plt.plot(d_losses, label='Discriminator')
plt.plot(g_losses, label='Generator')
plt.ylabel('loss')
plt.legend();
```
# 7. Интерполяция
Давайте попробуем взять два вектора, сгенерированных из нормального распределения и посмотреть как один из них перетекакет в другой.
```
from scipy.interpolate import interp1d
def show_interp_samples(point1, point2, N_samples_interp):
N_samples_interp_all = N_samples_interp + 2
# линия между двумя точками
line = interp1d([1, N_samples_interp_all], np.vstack([point1, point2]), axis=0)
fig = plt.figure(figsize=(15,4))
for i in range(N_samples_interp_all):
ax = fig.add_subplot(1, 2 + N_samples_interp, i+1)
ax.grid('off')
ax.axis('off')
ax.imshow(generator.predict(line(i + 1).reshape((1, 128)))[0, :, :, 0],cmap='gray')
plt.show()
pass
np.random.seed(seed=42)
# Рандомная точка в пространстве
noise_1 = np.random.normal(0, 1, (1, 128))
# смотрим как она перетекает в симметричкную
show_interp_samples(noise_1, -noise_1, 6)
noise_2 = np.random.normal(0, 1, (1, 128))
show_interp_samples(noise_1, noise_2, 6)
```
А что мы вообще сгенерировали?! Давайте посмотрим на точку из выборки наиболее близкую к получившейся генерации.
```
id_label_sample = 8
img_smp = generator.predict(sample_noise_batch(1))
plt.imshow(img_smp[0,:,:,0], cmap='gray')
img_smp.shape, X.shape
# ищем l1 норму между тем, что сгенерилось и остальным
L1d = np.sum(np.sum(np.abs(X[:,:,:,0] - img_smp[:,:,:,0]), axis=1), axis=1)
idx_l1_sort = L1d.argsort()
idx_l1_sort.shape
idx_l1_sort[:5]
N_closest = 8
fig = plt.figure(figsize=(15,4))
for i in range(N_closest):
ax = fig.add_subplot(1, N_closest, i+1)
ax.grid('off')
ax.axis('off')
ax.imshow(X[idx_l1_sort[i], :, :, 0], cmap='gray')
plt.show()
```
Сохраняю гифку из картинок.
```
import os
import glob
import imageio
def create_animated_gif(files, animated_gif_name, pause=0):
if pause != 0:
# Load the gifs up several times in the array, to slow down the animation
frames = []
for file in files:
count = 0
while count < pause:
frames.append(file)
count+=1
print("Total number of frames in the animation:", len(frames))
files = frames
images = [imageio.imread(file) for file in files]
imageio.mimsave(animated_gif_name, images, duration = 0.005)
pause = 1
animated_gif_name = 'animation_GAN.gif'
image_path = 'images_gan/*.png'
files = glob.glob(image_path)
files = sorted(files, key = lambda w: int(w.split('_')[-1].split('.')[0]))
create_animated_gif(files, animated_gif_name, pause)
```
На этом всё :)

| github_jupyter |
# Part 8 - Introduction to Plans
### Context
> Warning: This is still experimental and may change during June / July 2019
We introduce here an object which is crucial to scale to industrial Federated Learning: the Plan. It reduces dramatically the bandwidth usage, allows asynchronous schemes and give more autonomy to remote devices. The original concept of plan can be found in the paper [Towards Federated Learning at Scale: System Design](https://arxiv.org/pdf/1902.01046.pdf), but it has been adapted to our needs in the PySyft library using PyTorch.
A Plan is intended to store a sequence of torch operations, just like a function, but it allows to send this sequence of operations to remote workers and to keep a reference to it. This way, to compute remotely this sequence of $n$ operations on some remote input referenced through pointers, instead of sending $n$ messages you need now to send a single message with the references of the plan and the pointers. Actually, it's so much like a function that you need a function to build a plan! Hence, for high level users, the notion of plan disappears and is replaced by a magic feature which allow to send to remote workers arbitrary functions containing sequential torch functions.
One thing to notice is that the class of functions that you can transform into plans is currently limited to sequences of hooked torch operations exclusively. This excludes in particular logical structures like `if`, `for` and `while` statements, even if we are working to have workarounds soon. _To be completely precise, you can use these but the logical path you take (first `if` to False and 5 loops in `for` for example) in the first computation of your plan will be the one kept for all the next computations, which we want to avoid in the majority of cases._
Authors:
- Théo Ryffel - Twitter [@theoryffel](https://twitter.com/theoryffel) - GitHub: [@LaRiffle](https://github.com/LaRiffle)
- Bobby Wagner - Twitter [@bobbyawagner](https://twitter.com/bobbyawagner) - GitHub: [@robert-wagner](https://github.com/robert-wagner)
- Marianne Monteiro - Twitter [@hereismari](https://twitter.com/hereismari) - GitHub: [@mari-linhares](https://github.com/mari-linhares)
### Imports and model specifications
First let's make the official imports.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
```
And than those specific to PySyft, with one important note: **the local worker should be a client worker.** *Non client workers can store objects and we need this ability to run a plan.*
```
import syft as sy # import the Pysyft library
hook = sy.TorchHook(torch) # hook PyTorch ie add extra functionalities
# IMPORTANT: Local worker should not be a client worker
hook.local_worker.is_client_worker = False
server = hook.local_worker
```
We define remote workers or _devices_, to be consistent with the notions provided in the reference article.
We provide them with some data.
```
x11 = torch.tensor([-1, 2.]).tag('input_data')
x12 = torch.tensor([1, -2.]).tag('input_data2')
x21 = torch.tensor([-1, 2.]).tag('input_data')
x22 = torch.tensor([1, -2.]).tag('input_data2')
device_1 = sy.VirtualWorker(hook, id="device_1", data=(x11, x12))
device_2 = sy.VirtualWorker(hook, id="device_2", data=(x21, x22))
devices = device_1, device_2
```
### Basic example
Let's define a function that we want to transform into a plan. To do so, it's as simple as adding a decorator above the function definition!
```
@sy.func2plan()
def plan_double_abs(x):
x = x + x
x = torch.abs(x)
return x
```
Let's check, yes we have now a plan!
```
plan_double_abs
```
To use a plan, you need two things: to build the plan (_ie register the sequence of operations present in the function_) and to send it to a worker / device. Fortunately you can do this very easily!
#### Building a plan
To build a plan you just need to call it on some data.
Let's first get a reference to some remote data: a request is sent over the network and a reference pointer is returned.
```
pointer_to_data = device_1.search('input_data')[0]
pointer_to_data
```
If we tell the plan it must be executed remotely on the device`location:device_1`... we'll get an error because the plan was not built yet.
```
plan_double_abs.is_built
# This cell fails
plan_double_abs.send(device_1)
```
To build a plan you just need to call it on some data. When a plan is built all the commands are executed sequentially by the local worker, and are catched by the plan and stored in its `readable_plan` attribute!
```
plan_double_abs(torch.tensor([1., -2.]))
plan_double_abs.is_built
```
If we try to send the plan now it works!
```
# This cell is executed successfully
plan_double_abs.send(device_1)
```
One important thing to remember is that when a plan is built we pre-set ahead of computation the id(s) where the result(s) should be stored. This will allow to send commands asynchronously, to already have a reference to a virtual result and to continue local computations without waiting for the remote result to be computed. One major application is when you require computation of a batch on device_1 and don't want to wait for this computation to end to launch another batch computation on device_2.
#### Running a Plan on a reference pointer
We now feed the plan with a reference pointer to some data. Two things happens: (1) the plan is sent to the device and (2) it is run remotely.
1. This newly plan object is sent to the remote worker in a single communication round.
2. An other command is issued to run this plan remotely, so that the predefined location of the output of the plan now contains the result (remember we pre-set location of result ahead of computation). This also require a single communication round. _That's the place where we could run asynchronously_.
The result is simply a pointer, just like when you call an usual hooked torch function!
```
pointer_to_result = plan_double_abs(pointer_to_data)
print(pointer_to_result)
```
And you can simply ask the value back.
```
pointer_to_result.get()
```
### Towards a concrete example
But what we want to do is to apply Plan to Deep and Federated Learning, right? So let's look to a slightly more complicated example, using neural networks as you might be willing to use them.
Note that we are now transforming a method into a plan, so we use the `@` `sy.meth2plan` decorator instead
```
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(2, 3)
self.fc2 = nn.Linear(3, 2)
@sy.method2plan
def forward(self, x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=0)
net = Net()
```
So the only thing we did is adding the sy.method2plan decorator! And check that `net.forward` is again a plan.
```
net.forward
```
Let's build the plan using some mock data.
```
net(torch.tensor([1., 2.]))
```
Now there is a subtlety: because the plan depend on the net instance, if you send the plan *you also need to send the model*.
> For developers: this is not compulsory as you actually have a reference to the model in the plan, we could call model.send internally.
```
net.send(device_1)
```
Let's retrieve some remote data
```
pointer_to_data = device_1.search('input_data')[0]
```
Then, the syntax is just like normal remote sequential execution, that is, just like local execution. But compared to classic remote execution, there is a single communication round for each execution.
```
pointer_to_result = net(pointer_to_data)
pointer_to_result
```
And we get the result as usual!
```
pointer_to_result.get()
```
Et voilà! We have seen how to dramatically reduce the communication between the local worker (or server) and the remote devices!
### Switch between workers
One major feature that we want to have is to use the same plan for several workers, that we would change depending on the remote batch of data we are considering.
In particular, we don't want to rebuild the plan each time we change of worker. Let's see how we do this, using the previous example with our small network.
```
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.fc1 = nn.Linear(2, 3)
self.fc2 = nn.Linear(3, 2)
@sy.method2plan
def forward(self, x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return F.log_softmax(x, dim=0)
net = Net()
# Build plan
net(torch.tensor([1., 2.]))
```
Here are the main steps we just executed
```
net.send(device_1)
pointer_to_data = device_1.search('input_data')[0]
pointer_to_result = net(pointer_to_data)
pointer_to_result.get()
```
Let's get the model and the network back
```
net.get()
```
And actually the syntax is straight forward: we just send it to another device
```
net.send(device_2)
pointer_to_data = device_2.search('input_data')[0]
pointer_to_result = net(pointer_to_data)
pointer_to_result.get()
```
### Automatically building plans that are functions
For functions (`@sy.func2plan`) we can automatically build the plan with no need to explicitly call it, actually in the moment of creation the plan is already built.
To get this functionality the only thing you need to change when creating a plan is setting an argument to the decorator called `args_shape` which should be a list containing the shapes of each argument.
```
@sy.func2plan(args_shape=[(-1, 1)])
def plan_double_abs(x):
x = x + x
x = torch.abs(x)
return x
plan_double_abs.is_built
```
The `args_shape` parameter is used iternally to create mock tensors with the given shape which are used to build the plan.
```
@sy.func2plan(args_shape=[(1, 2), (-1, 2)])
def plan_sum_abs(x, y):
s = x + y
return torch.abs(s)
plan_sum_abs.is_built
```
At this point, the interest of Plans might not be straightforward, but we will soon reuse all these elements for a larger scale Federated Learning task where we will need more complex interactions between workers.
### Star PySyft on GitHub
The easiest way to help our community is just by starring the repositories! This helps raise awareness of the cool tools we're building.
- [Star PySyft](https://github.com/OpenMined/PySyft)
### Pick our tutorials on GitHub!
We made really nice tutorials to get a better understanding of what Federated and Privacy-Preserving Learning should look like and how we are building the bricks for this to happen.
- [Checkout the PySyft tutorials](https://github.com/OpenMined/PySyft/tree/master/examples/tutorials)
### Join our Slack!
The best way to keep up to date on the latest advancements is to join our community!
- [Join slack.openmined.org](http://slack.openmined.org)
### Join a Code Project!
The best way to contribute to our community is to become a code contributor! If you want to start "one off" mini-projects, you can go to PySyft GitHub Issues page and search for issues marked `Good First Issue`.
- [Good First Issue Tickets](https://github.com/OpenMined/PySyft/issues?q=is%3Aopen+is%3Aissue+label%3A%22good+first+issue%22)
### Donate
If you don't have time to contribute to our codebase, but would still like to lend support, you can also become a Backer on our Open Collective. All donations go toward our web hosting and other community expenses such as hackathons and meetups!
- [Donate through OpenMined's Open Collective Page](https://opencollective.com/openmined)
| github_jupyter |
# Imports
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
#Allows dataset from drive to be utilized
from google.colab import drive
drive.mount("/content/drive", force_remount=True)
#Dataset from location in drive
ds=pd.read_csv(LOCATION_OF_ORIGINAL_DATASET)
```
# Initial Dataset Visualized
These are various ways to analyze the amount of null values in each column of the dataset
```
ds.shape
print(ds)
#ds.head()
ds.loc[:, ds.isnull().any()].head()
pd.set_option('display.max_rows', 149)
#print(ds)
ds.isnull().sum()
#Heatmap of dataset showing the null values
sns.heatmap(ds.isnull(),yticklabels=False,cbar=False,cmap='viridis')
```
# Data Manipulation
## Columns to drop
```
drop_columns = [
'LUNG_RADS_DIAMETER_MM',
'cancer_type',
'eus_performed',
'fna_performed',
'cyto_results',
'operation_performed',
'path_cyst_id',
'path_mucin',
'path_cea',
'other_cysts',
'eus_dx',
'eus_consistency',
'multiples',
'ENTROPY_VOXELS',
'KURTOSIS_VOXELS',
'MEAN_DEVIATION_VOXELS',
'SKEWNESS_VOXELS',
'STD_DEV_VOXELS',
'VARIANCE_VOXELS',
#'ENERGY_VOXELS',
#'MAX_VOXELS',
#'MEAN_VOXELS',
#'MEDIAN_VOXELS',
#'MIN_VOXELS',
#'ROOT_MEAN_SQUARE_VOXELS',
#'SOLID_VOLUME_VOXELS',
#'UNIFORMITY_VOXELS',
#'VOLUME_VOXELS',
'path_duct',
'path_cyst_cat',
'path_dysplastic_margin',
'path_malignancy',
'path_grade_dysplasia',
'cystid',
'character_comment',
'othercyst_comments',
'path_cyst_count',
'path_dx',
'path_num_cysts',
'study_idnumber',
'study_id',
'path_available',
'serum_ca19',
'serum_cea',
'path_size',
'mucin_cyst',
'path_size',
'comments_clinical',
'path_pancreatitis',
'operation_performed_other',
'eus_fluid_other',
'eus_color',
'eus_other',
'amylase_cyst',
'cea_cyst',
'ca19_cyst',
'cyst_location_other',
'max_duct_dil',
'comments_rads',
'cancer_type',
'pancreatitis_dx_year',
'first_scan_reason',
'rad_dx_first_scan',
'rad_dx_last_scan',
'cea_cat',
'ANTPOST_LENGTH_END_MM_X',
'ANTPOST_LENGTH_END_MM_Y',
'ANTPOST_LENGTH_END_MM_Z',
'ANTPOST_LENGTH_START_MM_X',
'ANTPOST_LENGTH_START_MM_Y',
'ANTPOST_LENGTH_START_MM_Z',
'AUTO_CORONAL_LONG_AXIS_END_MM_X',
'AUTO_CORONAL_LONG_AXIS_END_MM_Y',
'AUTO_CORONAL_LONG_AXIS_END_MM_Z',
'AUTO_CORONAL_LONG_AXIS_END_VOXEL',
'V',
'W',
'AUTO_CORONAL_LONG_AXIS_START_MM_',
'Z',
'AA',
'AUTO_CORONAL_LONG_AXIS_START_VOX',
'AC',
'AD',
'AUTO_CORONAL_SHORT_AXIS_END_MM_X',
'AUTO_CORONAL_SHORT_AXIS_END_MM_Y',
'AUTO_CORONAL_SHORT_AXIS_END_MM_Z',
'AUTO_CORONAL_SHORT_AXIS_END_VOXE',
'AI',
'AJ',
'AUTO_CORONAL_SHORT_AXIS_START_MM',
'AM',
'AN',
'AUTO_CORONAL_SHORT_AXIS_START_VO',
'AP',
'AQ',
'AUTO_LARGEST_PLANAR_DIAMETER_END',
'AS',
'AT',
'AU',
'AV',
'AW',
'AUTO_LARGEST_PLANAR_DIAMETER_STA',
'AZ',
'BA',
'BB',
'BC',
'BD',
'AUTO_LARGEST_PLANAR_ORTHO_DIAMET',
'BF',
'BG',
'BH',
'BI',
'BJ',
'BK',
'BL',
'BM',
'BN',
'BO',
'BP',
'BQ',
'AUTO_SAGITTAL_LONG_AXIS_END_MM_X',
'AUTO_SAGITTAL_LONG_AXIS_END_MM_Y',
'AUTO_SAGITTAL_LONG_AXIS_END_MM_Z',
'AUTO_SAGITTAL_LONG_AXIS_END_VOXE',
'BV',
'BW',
'AUTO_SAGITTAL_LONG_AXIS_START_MM',
'BZ',
'CA',
'AUTO_SAGITTAL_LONG_AXIS_START_VO',
'CC',
'CD',
'AUTO_SAGITTAL_SHORT_AXIS_END_MM_',
'CF',
'CG',
'AUTO_SAGITTAL_SHORT_AXIS_END_VOX',
'CI',
'CJ',
'AUTO_SAGITTAL_SHORT_AXIS_START_M',
'CM',
'CN',
'AUTO_SAGITTAL_SHORT_AXIS_START_V',
'CP',
'CQ',
'CENTROID_X_MM',
'CENTROID_Y_MM',
'CENTROID_Z_MM',
'CONFIRMATION_STATUS',
'CORONAL_LONG_AXIS_END_MM_X',
'CORONAL_LONG_AXIS_END_MM_Y',
'CORONAL_LONG_AXIS_END_MM_Z',
'CORONAL_LONG_AXIS_END_VOXELS_X',
'CORONAL_LONG_AXIS_END_VOXELS_Y',
'CORONAL_LONG_AXIS_END_VOXELS_Z',
'CORONAL_LONG_AXIS_START_MM_X',
'CORONAL_LONG_AXIS_START_MM_Y',
'CORONAL_LONG_AXIS_START_MM_Z',
'CORONAL_LONG_AXIS_START_VOXELS_X',
'CORONAL_LONG_AXIS_START_VOXELS_Y',
'CORONAL_LONG_AXIS_START_VOXELS_Z',
'CORONAL_SHORT_AXIS_END_MM_X',
'CORONAL_SHORT_AXIS_END_MM_Y',
'CORONAL_SHORT_AXIS_END_MM_Z',
'CORONAL_SHORT_AXIS_END_VOXELS_X',
'CORONAL_SHORT_AXIS_END_VOXELS_Y',
'CORONAL_SHORT_AXIS_END_VOXELS_Z',
'CORONAL_SHORT_AXIS_START_MM_X',
'CORONAL_SHORT_AXIS_START_MM_Y',
'CORONAL_SHORT_AXIS_START_MM_Z',
'CORONAL_SHORT_AXIS_START_VOXELS_',
'DY',
'DZ',
'CRANIALCAUDAL_LENGTH_END_MM_X',
'CRANIALCAUDAL_LENGTH_END_MM_Y',
'CRANIALCAUDAL_LENGTH_END_MM_Z',
'CRANIALCAUDAL_LENGTH_START_MM_X',
'CRANIALCAUDAL_LENGTH_START_MM_Y',
'CRANIALCAUDAL_LENGTH_START_MM_Z',
'FOOTPRINT_END_MM_X',
'FOOTPRINT_END_MM_Y',
'FOOTPRINT_END_MM_Z',
'FOOTPRINT_END_VOXELS_X',
'FOOTPRINT_END_VOXELS_Y',
'FOOTPRINT_END_VOXELS_Z',
'FOOTPRINT_START_MM_X',
'FOOTPRINT_START_MM_Y',
'FOOTPRINT_START_MM_Z',
'FOOTPRINT_START_VOXELS_X',
'FOOTPRINT_START_VOXELS_Y',
'FOOTPRINT_START_VOXELS_Z',
'FOOTPRINT_X_MM',
'FOOTPRINT_Y_MM',
'FOOTPRINT_Z_MM',
'INIT_DRAG_LONG_END_PATIENT_X',
'INIT_DRAG_LONG_END_PATIENT_Y',
'INIT_DRAG_LONG_END_PATIENT_Z',
'INIT_DRAG_LONG_START_PATIENT_X',
'INIT_DRAG_LONG_START_PATIENT_Y',
'INIT_DRAG_LONG_START_PATIENT_Z',
'L1_AXIS_END_X_MM',
'L1_AXIS_END_Y_MM',
'L1_AXIS_END_Z_MM',
'L1_AXIS_START_X_MM',
'L1_AXIS_START_Y_MM',
'L1_AXIS_START_Z_MM',
'L1_UNIT_AXIS_X_MM',
'L1_UNIT_AXIS_Y_MM',
'L1_UNIT_AXIS_Z_MM',
'L2_AXIS_END_X_MM',
'L2_AXIS_END_Y_MM',
'L2_AXIS_END_Z_MM',
'L2_AXIS_START_X_MM',
'L2_AXIS_START_Y_MM',
'L2_AXIS_START_Z_MM',
'L2_UNIT_AXIS_X_MM',
'L2_UNIT_AXIS_Y_MM',
'L2_UNIT_AXIS_Z_MM',
'L3_AXIS_END_X_MM',
'L3_AXIS_END_Y_MM',
'L3_AXIS_END_Z_MM',
'L3_AXIS_START_X_MM',
'L3_AXIS_START_Y_MM',
'L3_AXIS_START_Z_MM',
'L3_UNIT_AXIS_X_MM',
'L3_UNIT_AXIS_Y_MM',
'L3_UNIT_AXIS_Z_MM',
'LARGEST_PLANAR_DIAMETER_END_MM_X',
'LARGEST_PLANAR_DIAMETER_END_MM_Y',
'LARGEST_PLANAR_DIAMETER_END_MM_Z',
'LARGEST_PLANAR_DIAMETER_END_VOXE',
'HD',
'HE',
'LARGEST_PLANAR_DIAMETER_START_MM',
'HH',
'HI',
'LARGEST_PLANAR_DIAMETER_START_VO',
'HK',
'HL',
'LARGEST_PLANAR_ORTHO_DIAMETER_EN',
'HN',
'HO',
'HP',
'HQ',
'HR',
'LARGEST_PLANAR_ORTHO_DIAMETER_ST',
'HU',
'HV',
'HW',
'HX',
'HY',
'LESION_TYPE',
'LUNG_RADS',
'LUNG_RADS_ISOLATION',
'PERCENT_AIR',
'PERCENT_GGO',
'PERCENT_SOLID',
'PERCENT_SOLID_INCL_AIR',
'SAGITTAL_LONG_AXIS_END_MM_X',
'SAGITTAL_LONG_AXIS_END_MM_Y',
'SAGITTAL_LONG_AXIS_END_MM_Z',
'SAGITTAL_LONG_AXIS_END_VOXELS_X',
'SAGITTAL_LONG_AXIS_END_VOXELS_Y',
'SAGITTAL_LONG_AXIS_END_VOXELS_Z',
'SAGITTAL_LONG_AXIS_START_MM_X',
'SAGITTAL_LONG_AXIS_START_MM_Y',
'SAGITTAL_LONG_AXIS_START_MM_Z',
'SAGITTAL_LONG_AXIS_START_VOXELS_',
'JG',
'JH',
'SAGITTAL_SHORT_AXIS_END_MM_X',
'SAGITTAL_SHORT_AXIS_END_MM_Y',
'SAGITTAL_SHORT_AXIS_END_MM_Z',
'SAGITTAL_SHORT_AXIS_END_VOXELS_X',
'SAGITTAL_SHORT_AXIS_END_VOXELS_Y',
'SAGITTAL_SHORT_AXIS_END_VOXELS_Z',
'SAGITTAL_SHORT_AXIS_START_MM_X',
'SAGITTAL_SHORT_AXIS_START_MM_Y',
'SAGITTAL_SHORT_AXIS_START_MM_Z',
'SAGITTAL_SHORT_AXIS_START_VOXELS',
'JT',
'JU',
'SLICE_INDEX',
'TRANSVERSE_LENGTH_END_MM_X',
'TRANSVERSE_LENGTH_END_MM_Y',
'TRANSVERSE_LENGTH_END_MM_Z',
'TRANSVERSE_LENGTH_START_MM_X',
'TRANSVERSE_LENGTH_START_MM_Y',
'TRANSVERSE_LENGTH_START_MM_Z',
'VOLUMETRIC_LENGTH_END_MM_X',
'VOLUMETRIC_LENGTH_END_MM_Y',
'VOLUMETRIC_LENGTH_END_MM_Z',
'VOLUMETRIC_LENGTH_END_VOXELS_X',
'VOLUMETRIC_LENGTH_END_VOXELS_Y',
'VOLUMETRIC_LENGTH_END_VOXELS_Z',
'VOLUMETRIC_LENGTH_START_MM_X',
'VOLUMETRIC_LENGTH_START_MM_Y',
'VOLUMETRIC_LENGTH_START_MM_Z',
'VOLUMETRIC_LENGTH_START_VOXELS_X',
'VOLUMETRIC_LENGTH_START_VOXELS_Y',
'VOLUMETRIC_LENGTH_START_VOXELS_Z',
'AVG_DENSITY',
'MASS_GRAMS',
'AVG_DENSITY_OF_GGO_REGION',
'AVG_DENSITY_OF_SOLID_REGION',
'EA',
'EB',
'INIT_DRAG_AXIAL_LA_END_MM_X',
'INIT_DRAG_AXIAL_LA_END_MM_Y',
'INIT_DRAG_AXIAL_LA_END_MM_Z',
'INIT_DRAG_AXIAL_LA_START_MM_X',
'INIT_DRAG_AXIAL_LA_START_MM_Y',
'INIT_DRAG_AXIAL_LA_START_MM_Z',
'HM',
'HS',
'HT',
'HZ',
'IC',
'ID',
'IE',
'IF',
'IG',
'MESH_STRUCTURE_MODIFIED',
'JP',
'JQ',
'KC',
'KD',
'SEG_BOUNDING_BOX_END_MM_X',
'SEG_BOUNDING_BOX_END_MM_Y',
'SEG_BOUNDING_BOX_END_MM_Z',
'SEG_BOUNDING_BOX_START_MM_X',
'SEG_BOUNDING_BOX_START_MM_Y',
'SEG_BOUNDING_BOX_START_MM_Z',
'MRNFROMHEALTHMYNE',
'ABS_CHANGE_BL_LA',
'ABS_CHANGE_BL_SA',
'ABS_CHANGE_BL_SLDV',
'ABS_CHANGE_BL_VOL',
'ABS_CHANGE_PR_LA',
'ABS_CHANGE_PR_SA',
'ABS_CHANGE_PR_SLDV',
'ABS_CHANGE_PR_VOL',
'AE',
'AF',
'AL',
'AR',
'AY',
'BE',
'BR',
'BS',
'BT',
'BU',
'BX',
'BY',
'CE',
'CL',
'CO',
'CR',
'CS',
'CV',
'CW',
'CY',
'CZ',
'EK',
'EL',
'DOUBLING_TIME_LA',
'DOUBLING_TIME_LA_DAYS',
'DOUBLING_TIME_SA',
'DOUBLING_TIME_SA_DAYS',
'DOUBLING_TIME_SLDV',
'DOUBLING_TIME_SLDV_DAYS',
'DOUBLING_TIME_VOL',
'DOUBLING_TIME_VOL_DAYS',
'INIT_DRAG_AXIAL_SA_END_MM_X',
'INIT_DRAG_AXIAL_SA_END_MM_Y',
'INIT_DRAG_AXIAL_SA_END_MM_Z',
'INIT_DRAG_AXIAL_SA_START_MM_X',
'INIT_DRAG_AXIAL_SA_START_MM_Y',
'INIT_DRAG_AXIAL_SA_START_MM_Z',
'IJ',
'IK',
'IN',
'IO',
'IQ',
'IR',
'IT',
'IU',
'IV',
'IW',
'IX',
'JA',
'JB',
'JC',
'JD',
'JE',
'PERCENT_CHANGE_BL_LA',
'PERCENT_CHANGE_BL_SA',
'PERCENT_CHANGE_BL_SLDV',
'PERCENT_CHANGE_BL_VOL',
'PERCENT_CHANGE_PR_LA',
'PERCENT_CHANGE_PR_SA',
'PERCENT_CHANGE_PR_SLDV',
'PERCENT_CHANGE_PR_VOL',
'RATE_OF_GROWTH_LA',
'RATE_OF_GROWTH_SA',
'RATE_OF_GROWTH_SLDV',
'RATE_OF_GROWTH_VOL',
'LA',
'LB',
'LN',
'LO',
]
ds.drop(drop_columns,axis=1,inplace=True)
```
## Patients to drop
```
#Rows (Patients) to Drop
ds = ds[ds['mucinous'].notna()] # One patient missing mucinous value
```
## Fill column data
```
#Data to Fill
#ds['']=ds[''].fillna(ds[''].mode()[0])
ds['height']=ds['height'].fillna(ds['height'].mode()[0])
```
# Manipulated Data Visualized
```
#Shape after Dropping
ds.shape
#Heatmap of dataset showing the null values
sns.heatmap(ds.isnull(),yticklabels=False,cbar=False,cmap='viridis')
pd.set_option('display.max_rows', 5)
pd.set_option('display.max_columns', 5)
ds.isnull().sum()
#print(ds)
```
# Export Modified dataset
```
#Export current modified data set
ds.to_csv('original_processed.csv',index=False)
!cp original_processed.csv {DATASET_SAVE_LOCATION}
#Mucinous Dataset
dataframe =pd.read_csv(DATASET_SAVE_LOCATION)
# Drop hgd_malignant row
dataframe.drop('hgd_malignancy',axis=1,inplace=True)
#Export to .csv and save in drive
dataframe.to_csv('mucinous_processed.csv',index=False)
!cp mucinous_processed.csv {DATASET_SAVE_LOCATION}
#hgd dataset
dataframe =pd.read_csv(DATASET_SAVE_LOCATION)
# Find and Delete nonmucinous rows
nonMucinous = dataframe[ dataframe['mucinous'] == 0 ].index
dataframe.drop(nonMucinous, inplace=True)
dataframe.drop('mucinous',axis=1,inplace=True)
#Change missing values in 'hgd_malignancy' to 0
dataframe['hgd_malignancy']=dataframe['hgd_malignancy'].fillna(value=0)
#Export to .csv and save in drive
dataframe.to_csv('hgd_processed.csv',index=False)
!cp hgd_processed.csv {DATASET_SAVE_LOCATION}
```
## Texture only feature set
```
#Mucinous Dataset
dataframe =pd.read_csv(DATASET_SAVE_LOCATION)
# Drop hgd_malignant row
dataframe.drop('hgd_malignancy',axis=1,inplace=True)
#Export to .csv and save in drive
dataframe.to_csv('texture_feature_set_mucinous_processed.csv',index=False)
!cp texture_feature_set_mucinous_processed.csv {DATASET_SAVE_LOCATION}
#hgd dataset
dataframe =pd.read_csv(DATASET_SAVE_LOCATION)
# Find and Delete nonmucinous rows
nonMucinous = dataframe[ dataframe['mucinous'] == 0 ].index
dataframe.drop(nonMucinous, inplace=True)
dataframe.drop('mucinous',axis=1,inplace=True)
#Change missing values in 'hgd_malignancy' to 0
dataframe['hgd_malignancy']=dataframe['hgd_malignancy'].fillna(value=0)
#Export to .csv and save in drive
dataframe.to_csv('texture_feature_set_hgd_processed.csv',index=False)
!cp texture_feature_set_hgd_processed.csv {DATASET_SAVE_LOCATION}
```
## Clinical Only Features
```
#Mucinous Dataset
dataframe = pd.read_csv(DATASET_SAVE_LOCATION)
# Drop hgd_malignant row
dataframe.drop('hgd_malignancy',axis=1,inplace=True)
#Export to .csv and save in drive
dataframe.to_csv('clinical_data_mucinous_processed.csv',index=False)
!cp clinical_data_mucinous_processed.csv {DATASET_SAVE_LOCATION}
#hgd dataset
dataframe =pd.read_csv(DATASET_SAVE_LOCATION)
# Find and Delete nonmucinous rows
nonMucinous = dataframe[ dataframe['mucinous'] == 0 ].index
dataframe.drop(nonMucinous, inplace=True)
dataframe.drop('mucinous',axis=1,inplace=True)
#Change missing values in 'hgd_malignancy' to 0
dataframe['hgd_malignancy']=dataframe['hgd_malignancy'].fillna(value=0)
#Export to .csv and save in drive
dataframe.to_csv('clinical_data_hgd_processed.csv',index=False)
!cp clinical_data_hgd_processed.csv {DATASET_SAVE_LOCATION}
```
| github_jupyter |
```
# !cp /content/drive/MyDrive/TempData/BookingChallenge.zip /content
# !unzip /content/BookingChallenge.zip
```
## ItemPop Model
```
import pandas as pd
train_set = pd.read_csv('train_set.csv').sort_values(by=['utrip_id','checkin'])
print(train_set.shape)
train_set.head()
test_set = pd.read_csv('test_set.csv').sort_values(by=['utrip_id','checkin'])
print(test_set.shape)
test_set.head()
```
Generate Dummy Predictions - use top 4 cities in the trainset as benchmark recommendation
```
topcities = train_set.city_id.value_counts().index[:4]
test_trips = (test_set[['utrip_id']].drop_duplicates()).reset_index().drop('index', axis=1)
cities_prediction = pd.DataFrame([topcities]*test_trips.shape[0]
, columns= ['city_id_1','city_id_2','city_id_3','city_id_4'])
```
Create Submission file according to the format
```
submission = pd.concat([test_trips,cities_prediction], axis =1)
print(submission.shape)
submission.head()
submission.to_csv('submission.csv',index=False)
```
Read submission file and ground truth
```
ground_truth = pd.read_csv('ground_truth.csv',index_col=[0])
submission = pd.read_csv('submission.csv',index_col=[0])
print(ground_truth.shape)
ground_truth.head()
```
Evaluate - use accuracy at 4 to evaluate the prediction
```
def evaluate_accuracy_at_4(submission,ground_truth):
'''checks if the true city is within the four recommended cities'''
data = submission.join(ground_truth, on='utrip_id')
hits = ((data['city_id']==data['city_id_1'])|(data['city_id']==data['city_id_2'])|
(data['city_id']==data['city_id_3'])|(data['city_id']==data['city_id_4']))*1
return hits.mean()
evaluate_accuracy_at_4(submission,ground_truth)
```
## X
```
# # Install RAPIDS
# !git clone https://github.com/rapidsai/rapidsai-csp-utils.git
# !bash rapidsai-csp-utils/colab/rapids-colab.sh stable
# import sys, os
# dist_package_index = sys.path.index('/usr/local/lib/python3.7/dist-packages')
# sys.path = sys.path[:dist_package_index] + ['/usr/local/lib/python3.7/site-packages'] + sys.path[dist_package_index:]
# sys.path
# exec(open('rapidsai-csp-utils/colab/update_modules.py').read(), globals())
import pandas as pd
from sklearn.model_selection import GroupKFold
import cudf
from numba import cuda
train = cudf.read_csv('train_set.csv').sort_values(by=['user_id','checkin'])
test = cudf.read_csv('test_set.csv').sort_values(by=['user_id','checkin'])
print(train.shape, test.shape)
train.head()
test.sample(10)
train['istest'] = 0
test['istest'] = 1
raw = cudf.concat([train,test], sort=False )
raw = raw.sort_values( ['user_id','checkin'], ascending=True )
raw.head()
raw['fold'] = 0
group_kfold = GroupKFold(n_splits=5)
for fold, (train_index, test_index) in enumerate(group_kfold.split(X=raw, y=raw, groups=raw['utrip_id'].to_pandas())):
raw.iloc[test_index,10] = fold
raw['fold'].value_counts()
#This flag tell which row must be part of the submission file.
raw['submission'] = 0
raw.loc[ (raw.city_id==0)&(raw.istest) ,'submission'] = 1
raw.loc[ raw.submission==1 ]
#number of places visited in each trip
aggs = raw.groupby('utrip_id', as_index=False)['user_id'].count().reset_index()
aggs.columns = ['utrip_id', 'N']
raw = raw.merge(aggs, on=['utrip_id'], how='inner')
raw['utrip_id_'], mp = raw['utrip_id'].factorize()
def get_order_in_group(utrip_id_,order):
for i in range(cuda.threadIdx.x, len(utrip_id_), cuda.blockDim.x):
order[i] = i
def add_cumcount(df, sort_col, outputname):
df = df.sort_values(sort_col, ascending=True)
tmp = df[['utrip_id_', 'checkin']].groupby(['utrip_id_']).apply_grouped(
get_order_in_group,incols=['utrip_id_'],
outcols={'order': 'int32'},
tpb=32)
tmp.columns = ['utrip_id_', 'checkin', outputname]
df = df.merge(tmp, how='left', on=['utrip_id_', 'checkin'])
df = df.sort_values(sort_col, ascending=True)
return(df)
raw = add_cumcount(raw, ['utrip_id_','checkin'], 'dcount')
raw['icount'] = raw['N']-raw['dcount']-1
raw.head(20)
```
| github_jupyter |
```
import sqlite3
# Create a SQL connection to our SQLite database
con = sqlite3.connect("db.sqlite3")
cur = con.cursor()
messages = cur.execute('SELECT message FROM comp_493_message')
# The result of a "cursor.execute" can be iterated over by row
for row in cur.execute('SELECT * FROM comp_493_message'):
print(row)
# Be sure to close the connection
print(messages)
import re
list = ["guru99 get", "guru99 give", "guru Selenium"]
for element in list:
z = re.match("(g\w+)\W(g\w+)", element)
if z:
print((z.groups()))
patterns = ['kill', 'hell', 'damn', 'shit']
text = str(messages)
for pattern in patterns:
print('Looking for "%s" in "%s" ->' % (pattern, text), end=' ')
if re.findall(pattern, text):
print('found a match! "%s" in "%s"' % (pattern, text))
con.close()
import sqlite3
import re
# Create a SQL connection to our SQLite database
con = sqlite3.connect("db.sqlite3")
cur = con.cursor()
# Return all results of query
cur.execute('SELECT message, username FROM comp_493_message')
message = cur.fetchall()
print(message)
print('\n')
print(str(message))
# Return first result of query
#cur.execute('SELECT species FROM species WHERE taxa="Bird"')
#cur.fetchone()
text = str(message)
print("\n")
text1 = "'s very sad to see people just coming in there to chat and when they ask of your country and you tell them, they just close the chat window. That is totally rude. If you are not ready to chat with people from certain countries, please suit yourself and don't enter the chat room. It's really unfair to close the chat window whiles the person thinks you are still chatting"
patterns = ['kill', 'hell', 'damn', 'shit', "sad", 'what']
for pattern in patterns:
if re.search(pattern, text):
print('found a match! "%s" in "%s"' % (pattern, text))
print("\n")
#import tkMessageBox
#tkMessageBox.showinfo(title="Threat", message="Hello World!")
# Be sure to close the connection
con.close()
import pandas as pd
import sqlite3
import re
# Read sqlite query results into a pandas DataFrame
con = sqlite3.connect("db.sqlite3")
df = pd.read_sql_query("SELECT * from comp_493_message", con)
message = df.message
# Verify that result of SQL query is stored in the dataframe
print(message)
print('\n')
text = str(message)
print(text)
text = str(messages)
text1 = "fuck you damn shit"
patterns = ['kill', 'hell', 'damn', 'shit']
txt = "The rain in Spain"
x = re.findall(patterns, text1)
print(x)
for pattern in patterns:
if re.search(pattern, text):
print('found a match! "%s" in "%s"' % (pattern, text1))
con.close()
```
| github_jupyter |
```
import os
import pandas as pd
from sklearn.utils import shuffle
from skimage.io import imshow, imread, imsave
data_path = "/home/yanglei/Classfier/dataset"
train_path = os.path.join(data_path, "train")
test_path = os.path.join(data_path, "test")
```
## 1. Train
```
df_train = pd.DataFrame({"image_path": "", "smoking_images": 0, "calling_images": 0, "normal_images": 0}, index = [0])
df_train = df_train.drop(index=0)
for image_path in os.listdir(os.path.join(train_path, "smoking_images")):
image_path = "train/smoking_images/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 1, "calling_images": 0, "normal_images": 0}, ignore_index=True)
for image_path in os.listdir(os.path.join(train_path, "calling_images")):
image_path = "train/calling_images/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 0, "calling_images": 1, "normal_images": 0}, ignore_index=True)
for image_path in os.listdir(os.path.join(train_path, "normal_images")):
image_path = "train/normal_images/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 0, "calling_images": 0, "normal_images": 1}, ignore_index=True)
for image_path in os.listdir(os.path.join(train_path, "smoke")):
image_path = "train/smoke/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 1, "calling_images": 0, "normal_images": 0}, ignore_index=True)
for image_path in os.listdir(os.path.join(train_path, "phone")):
image_path = "train/phone/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 0, "calling_images": 1, "normal_images": 0}, ignore_index=True)
for image_path in os.listdir(os.path.join(train_path, "normal")):
image_path = "train/normal/" + image_path
df_train = df_train.append({"image_path": image_path, "smoking_images": 0, "calling_images": 0, "normal_images": 1}, ignore_index=True)
df_train = shuffle(df_train)
df_train.to_csv(os.path.join(data_path, "train_all_image.csv"), index=0)
```
## 2. Test
```
df_test = pd.DataFrame({"image_path": "", "smoking_images": 0, "calling_images": 0, "normal_images": 0}, index = [0])
df_test = df_test.drop(index=0)
for image_path in os.listdir(test_path):
image_path = "test/" + image_path
df_test = df_test.append({"image_path": image_path, "smoking_images": 0, "calling_images": 0, "normal_images": 0}, ignore_index=True)
df_test
df_test = shuffle(df_test)
df_test.to_csv(os.path.join(data_path, "test.csv"), index=0)
```
## 3. Image Size
```
for image_path in os.listdir(test_path):
image_path = "dataset/test/" + image_path
image = imread(image_path)
print(image.shape)
print(image.shape[1] / image.shape[0])
```
| github_jupyter |
<a href="https://colab.research.google.com/github/bala-codes/Mini_Word_Embeddings/blob/master/codes/Mini_Word_Embedding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import numpy as np
import matplotlib.pylab as plt
import pandas as pd
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
from torch.utils.data import Dataset, DataLoader
import random
from keras.preprocessing.text import one_hot
from keras.preprocessing.sequence import pad_sequences
from sklearn.preprocessing import OneHotEncoder
!pip install torchsummaryX --quiet
from torchsummaryX import summary
```
# Data - Preprocessing - Steps : Visualization
<img src="https://cdn-images-1.medium.com/max/800/1*SANI-0E8qSTHzGHg4HHtNA.png">
<img src="https://cdn-images-1.medium.com/max/1200/0*dEB7_sqWOHhV82EI.png">
# Dataset Source
<img src="https://github.com/bala-codes/Mini_Word_Embeddings/blob/master/codes/Capture.PNG?raw=true">
<img src="https://cdn-images-1.medium.com/max/800/1*pDeswCr7eBhEDtiywMkAqQ.png">
<img src="https://cdn-images-1.medium.com/max/800/1*SANI-0E8qSTHzGHg4HHtNA.png">
<img src="https://cdn-images-1.medium.com/max/1200/1*r_9zfDywyD1TbVpxKENHjw.png">
```
# Sample Document # Recreated from the tom and jerry cartoon
docs = ["cat and mice are buddies",
'mice lives in hole',
'cat lives in house',
'cat chases mice',
'cat catches mice',
'cat eats mice',
'mice runs into hole',
'cat says bad words',
'cat and mice are pals',
'cat and mice are chums',
'mice stores food in hole',
'cat stores food in house',
'mice sleeps in hole',
'cat sleeps in house']
idx_2_word = {}
word_2_idx = {}
temp = []
i = 1
for doc in docs:
for word in doc.split():
if word not in temp:
temp.append(word)
idx_2_word[i] = word
word_2_idx[word] = i
i += 1
print(idx_2_word)
print(word_2_idx)
```
# Words to numbers
```
vocab_size = 25
def one_hot_map(doc):
x = []
for word in doc.split():
x.append(word_2_idx[word])
return x
encoded_docs = [one_hot_map(d) for d in docs]
encoded_docs
```
# Padding
```
max_len = 10
padded_docs = pad_sequences(encoded_docs, maxlen=max_len, padding='post')
padded_docs
```
# Creating dataset tuples for training
```
training_data = np.empty((0,2))
window = 2
for sentence in padded_docs:
sent_len = len(sentence)
for i, word in enumerate(sentence):
w_context = []
if sentence[i] != 0:
w_target = sentence[i]
for j in range(i-window, i + window + 1):
if j != i and j <= sent_len -1 and j >=0 and sentence[j]!=0:
w_context = sentence[j]
training_data = np.append(training_data, [[w_target, w_context]], axis=0)
#training_data.append([w_target, w_context])
print(len(training_data))
print(training_data.shape)
training_data
enc = OneHotEncoder()
enc.fit(np.array(range(30)).reshape(-1,1))
onehot_label_x = enc.transform(training_data[:,0].reshape(-1,1)).toarray()
onehot_label_x
enc = OneHotEncoder()
enc.fit(np.array(range(30)).reshape(-1,1))
onehot_label_y = enc.transform(training_data[:,1].reshape(-1,1)).toarray()
onehot_label_y
print(onehot_label_x[0])
print(onehot_label_y[0])
# From Numpy to Torch
onehot_label_x = torch.from_numpy(onehot_label_x)
onehot_label_y = torch.from_numpy(onehot_label_y)
print(onehot_label_x.shape, onehot_label_y.shape)
```
<img src="https://cdn-images-1.medium.com/max/800/1*rb9i3dT_rH3DB31atto_Ag.png">
```
class WEMB(nn.Module):
def __init__(self, input_size, hidden_size):
super(WEMB, self).__init__()
self.input_size = input_size
self.hidden_size = hidden_size
self.softmax = nn.Softmax(dim=1)
self.l1 = nn.Linear(self.input_size, self.hidden_size, bias=False)
self.l2 = nn.Linear(self.hidden_size, self.input_size, bias=False)
def forward(self, x):
out_bn = self.l1(x) # bn - bottle_neck
out = self.l2(out_bn)
out = self.softmax(out)
return out, out_bn
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
'''
m = nn.Softmax() #nn.Sigmoid()
loss = nn.BCELoss()
input = torch.tensor([0.9,0.0,0.0]) #torch.randn(3, requires_grad=True)
target = torch.tensor([1.0,0.0,0.0])
output = loss(m(input), target)
print(input, m(input), target) tensor([0.9000, 0.0000, 0.0000]) tensor([0.5515, 0.2242, 0.2242]) tensor([1., 0., 0.])
output -> 0.36
'''
'''
m = nn.Sigmoid()
loss = nn.BCELoss()
input = torch.tensor([0.9,0.0,0.0])
target = torch.tensor([1.0,0.0,0.0])
output = loss(m(input), target)
print(input, m(input), target) tensor([0.9000, 0.0000, 0.0000]) tensor([0.7109, 0.5000, 0.5000]) tensor([1., 0., 0.])
output -> 0.5758
'''
input_size = 30
hidden_size = 2
learning_rate = 0.01
num_epochs = 10000
untrained_model = WEMB(input_size, hidden_size).to(device)
model = WEMB(input_size, hidden_size).to(device)
model.train(True)
print(model)
print()
# Loss and optimizer
criterion = nn.BCELoss()
#optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate, momentum=0, dampening=0, weight_decay=0, nesterov=False)
summary(model, torch.ones((1,30)))
loss_val = []
onehot_label_x = onehot_label_x.to(device)
onehot_label_y = onehot_label_y.to(device)
for epoch in range(num_epochs):
for i in range(onehot_label_y.shape[0]):
inputs = onehot_label_x[i].float()
labels = onehot_label_y[i].float()
inputs = inputs.unsqueeze(0)
labels = labels.unsqueeze(0)
# Forward pass
output, wemb = model(inputs)
loss = criterion(output, labels)
# Backward and optimize
optimizer.zero_grad()
loss.backward()
optimizer.step()
loss_val.append(loss.item())
if (epoch+1) % 100 == 0:
print (f'Epoch [{epoch+1}/{num_epochs}], Loss: {loss.item():.4f}')
plt.plot(loss_val)
docs = ['cat and mice are buddies hole lives in house chases catches runs into says bad words pals chums stores sleeps']
encoded_docs = [one_hot_map(d) for d in docs]
test_arr = np.array([[ 1., 2., 3., 4., 5., 8., 6., 7., 9., 10., 11., 13., 14., 15., 16., 17., 18., 19., 20., 22.]])
test = enc.transform(test_arr.reshape(-1,1)).toarray()
output = []
for i in range(test.shape[0]):
_, wemb2 = model(torch.from_numpy(test[i]).unsqueeze(0).float())
wemb2 = wemb2[0].detach().cpu().numpy()
output.append(wemb2)
print(len(output))
print(output)
docs = ['cat', 'and', 'mice', 'are', 'buddies', 'hole', 'lives', 'in', 'house', 'chases', 'catches', 'runs', 'into', 'says', 'bad', 'words', 'pals', 'chums', 'stores', 'sleeps']
for i in range(0, len(docs)):
print("Word - {} - It's Word Embeddings : {:.3} & {:.3} -".format(docs[i], output[i][0], output[i][0]))
xs = []
ys = []
for i in range(len(output)):
xs.append(output[i][0])
ys.append(output[i][1])
print(xs, ys)
docs = ['cat', 'and', 'mice', 'are', 'buddies', 'hole', 'lives', 'in', 'house', 'chases', 'catches', 'runs', 'into', 'says', 'bad', 'words', 'pals', \
'chums', 'stores', 'sleeps']
plt.clf()
plt.figure(figsize=(12,12))
plt.scatter(xs,ys)
label = docs
for i,(x,y) in enumerate(zip(xs,ys)):
plt.annotate(label[i], (x,y), textcoords="offset points", xytext=(0,10), fontsize=20, ha = random.choice(['left', 'right']))
plt.title("Trained Model")
plt.show()
import plotly
import plotly.express as px
import plotly.express as px
fig = px.scatter(x=xs, y=ys, text=docs, size_max=100)
fig.update_traces(textposition= random.choice(['top center', 'bottom center','bottom left']))
fig.update_layout(height=800,title_text='Custom Word Embeddings')
fig.show()
```
# How Untrained Model behaves with our data
```
output = []
for i in range(test.shape[0]):
_, wemb2 = untrained_model(torch.from_numpy(test[i]).unsqueeze(0).float()) # Here I am loading the untrained model
wemb2 = wemb2[0].detach().cpu().numpy()
output.append(wemb2)
print(len(output))
print(output)
xs = []
ys = []
for i in range(len(output)):
xs.append(output[i][0])
ys.append(output[i][1])
print(xs, ys)
docs = ['cat', 'and', 'mice', 'are', 'buddies', 'hole', 'lives', 'in', 'house', 'chases', 'catches', 'runs', 'into', 'says', 'bad', 'words', 'pals', \
'chums', 'stores', 'sleeps']
plt.clf()
plt.figure(figsize=(12,12))
plt.scatter(xs,ys)
label = docs
for i,(x,y) in enumerate(zip(xs,ys)):
plt.annotate(label[i], (x,y), textcoords="offset points", xytext=(0,10), fontsize=20, ha = random.choice(['left', 'right']))
plt.title("Un-Trained Model")
plt.show()
import plotly
import plotly.express as px
import plotly.express as px
fig = px.scatter(x=xs, y=ys, text=docs, size_max=100)
fig.update_traces(textposition= random.choice(['top center', 'bottom center','bottom left']))
fig.update_layout(height=800,title_text='Custom Word Embeddings')
fig.show()
```
| github_jupyter |
# Exploratory data analysis on the San Francisco 311 data
Data can be downloaded from: https://data.sfgov.org/City-Infrastructure/311-Cases/vw6y-z8j6/data
```
%matplotlib inline
%load_ext autoreload
import matplotlib.pyplot as plt
import pandas as pd
pd.options.mode.chained_assignment = None # default='warn'
import numpy as np
## The time window to bucket samples
TIME_RANGE = '24H'
## File path (original data is ~1GB, this is a reduced version with only categories and dates)
#Original file:
#DATAPATH = "SF311_simplified.csv"
#Sample raw data:
DATAPATH = "SF_data/SF-311_simplified.csv"
```
### Read sample of data (original data contains additional columns)
```
raw_sample = pd.read_csv(DATAPATH, nrows=5)
raw_sample.head()
raw = pd.read_csv(DATAPATH).drop(columns='Unnamed: 0')
raw.head(10)
```
#### Initial data prep
```
## Rename columns
raw = raw.rename(columns={'Opened': 'date', 'Category': 'category'})
print(raw.head())
## Turn the raw data into a time series (with date as DatetimeIndex)
from moda.dataprep import raw_to_ts
ts = raw_to_ts(raw,date_format="%m/%d/%Y %H:%M:%S %p")
ts.head()
## Some general stats
print("Dataset length: " + str(len(ts)))
print("Min date: " + str(ts.index.get_level_values('date').min()))
print("Max date: " + str(ts.index.get_level_values('date').max()))
print("Total time: {}".format(ts.index.get_level_values('date').max() - ts.index.get_level_values('date').min()))
print("Dataset contains {} categories.".format(len(ts['category'].unique())))
```
#### Next, we decide on the time interval and aggregate items per time and category
```
from moda.dataprep import ts_to_range
ranged_ts = ts_to_range(ts,time_range=TIME_RANGE)
ranged_ts.head(20)
#I'm using dfply because I like its functional-like syntax. This can also be done with plain pandas.
#!pip install dfply
from dfply import *
## Remove categories with less than 1000 items (in more than 10 years) or that existed less than 100 days
min_values = 1000
min_days = 100
categories = ranged_ts.reset_index() >> group_by(X.category) >> \
summarise(value = np.sum(X.value),duration_in_dataset = X.date.max()-X.date.min()) >> \
ungroup() >> \
mask(X.duration_in_dataset.dt.days > min_days) >> \
mask(X.value > min_values) >> \
arrange(X.value,ascending=False)
print("Filtered dataset contains {0} categories,\nafter filtering the small ones that existed less than {1} days or had {2} values of less.".
format(len(categories),min_days,min_values))
categories.head()
```
### Most common categories
```
category_names = categories['category'].values
num_categories = len(categories)
major_category_threshold=11
major_categories = category_names[:major_category_threshold]
minor_categories = category_names[major_category_threshold:]
fig, axes = plt.subplots(nrows=1, ncols=2)
fig.set_figheight(5)
fig.set_figwidth(20)
categories[categories['category'].isin(major_categories)].plot(kind='bar',
x='category',
y='value',
title="Top "+str(major_category_threshold-1)+" common categories on the SF311 dataset",
ax=axes[0])
categories[categories['category'].isin(minor_categories)].plot(kind='bar',
x='category',
y='value',
title=str(major_category_threshold)+"th to "+str(num_categories)+"th most common categories on the SF311 dataset",
ax=axes[1])
plt.savefig("category_values.png",bbox_inches='tight')
```
### Change in requests per category from year to year
```
## Calculate the number of values per category per year
categories_yearly = ranged_ts.reset_index() >> mutate(year = X.date.dt.year) >> group_by(X.category,X.year) >> \
summarise(value_per_year = np.sum(X.value),
duration_in_dataset = X.date.max()-X.date.min()) >>\
ungroup() >> \
mask(X.value_per_year > (min_values/12.0)) >> \
arrange(X.value_per_year,ascending=False)
import seaborn as sns
major_cats_yearly = categories_yearly[categories_yearly['category'].isin(major_categories)]
g = sns.factorplot(x='category', y='value_per_year', hue='year', data=major_cats_yearly, kind='bar', size=4, aspect=4,legend=True)
g.set_xticklabels(rotation=90)
axes = g.axes.flatten()
axes[0].set_title("Yearly number of incidents for the top "+str(major_category_threshold-1)+" categories")
plt.savefig("yearly_values.png",bbox_inches='tight')
minor_cats_yearly = categories_yearly[categories_yearly['category'].isin(minor_categories)]
g = sns.factorplot(x='category', y='value_per_year', hue='year', data=minor_cats_yearly, kind='bar', size=4, aspect=4,legend=True)
g.set_xticklabels(rotation=90)
axes = g.axes.flatten()
axes[0].set_title("Yearly number of incidents for the "+str(major_category_threshold)+"th to "+str(num_categories)+"th categories")
```
### Correlation between categories over time
```
categories_yearly_pivot = categories_yearly.pivot("year", "category", "value_per_year")
categories_yearly_pivot.head()
corr = categories_yearly_pivot.corr()
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
```
### One category inspection
#### Example 1: Noise Reports
```
category = "Noise Report"
ranged_ts.loc[pd.IndexSlice[:, category], :].reset_index().plot(kind='line',x='date',y='value',figsize=(24,6),linewidth=0.7,
title = "Number of incidents per 24 hours for {}".format(category))
```
#### Example 2: Street and Sidewalk Cleaning
```
START = '2015-11-01'
END = '2018-07-01'
category = 'Street and Sidewalk Cleaning'
cleaning = ranged_ts.loc[pd.IndexSlice[:, category], :].reset_index()
cleaning[(cleaning.date > START) & (cleaning.date<=END)].plot(kind='line',x='date',y='value',figsize=(24,6),linewidth=0.7,
title = "Number of incidents per 24 hours for {0} between {1} and {2}".format(category,START,END))
```
As comparison, let's look at the same time series with different time ranges (30 minutes, 1 hour and 24 hours), only on two months of data
```
from moda.dataprep.ts_to_range import ts_to_range
ranged_ts_3H = ts_to_range(ts,time_range='3H',pad_with_zeros=True)
ranged_ts_30min = ts_to_range(ts,time_range='30min',pad_with_zeros=True)
START = '2015-11-01'
END = '2016-01-01'
category = 'Street and Sidewalk Cleaning'
fig, axes = plt.subplots(nrows=3, ncols=1,figsize=(20,12))
cleaning_30min = ranged_ts_30min.loc[pd.IndexSlice[:, category], :].reset_index()
a1=cleaning_30min[(cleaning_30min.date > START) & (cleaning_30min.date<=END)].plot(kind='line',x='date',y='value',linewidth=0.7, ax=axes[0])
cleaning_3H = ranged_ts_3H.loc[pd.IndexSlice[:, category], :].reset_index()
a2=cleaning_3H[(cleaning_3H.date > START) & (cleaning_3H.date<=END)].plot(kind='line',x='date',y='value',linewidth=0.7, ax=axes[1])
cleaning_24H = ranged_ts.loc[pd.IndexSlice[:, category], :].reset_index()
a3=cleaning_24H[(cleaning_24H.date > START) & (cleaning_24H.date<=END)].plot(kind='line',x='date',y='value',linewidth=0.7, ax=axes[2])
```
We can see that there are multiple seasonality factors in this time series. Hourly and weekly patterns are visible on the 30 minute interval time series, and the 3 hours interval time series
## Evaluating different models on the SF 24H data
First, in order to be able to estimate our models, we use [TagAnomaly](https://github.com/Microsoft/TagAnomaly) to tag the points we think are showing trends in the data. Taganomaly can be found here: https://github.com/Microsoft/TagAnomaly
Second, we join the tagged dataset with the time series dataset. Each sample which isn't included in the tagged dataset is assumed to be non-trending (or normal)
```
## Add labeled data
labels24H = pd.read_csv('SF_data/SF_24H_anomalies_only.csv',usecols=['date','category','value'])
labels24H.date = pd.to_datetime(labels24H.date)
labels24H['label'] = 1
labels24H.sort_values(by='date').head()
# Since we have labels only for 2018, we'll filter out previous years.
ts2018 = ranged_ts[ranged_ts.index.get_level_values(0).year == 2018]
ts2018.head()
df24H = pd.merge(ts2018.reset_index(),labels24H,how='left',on=['date','category'])
df24H['label'] = np.where(np.isnan(df24H['value_y']),0,1)
df24H = df24H.set_index([pd.DatetimeIndex(df24H['date']),'category'])
df24H = df24H.drop(columns = ['date','value_y']).rename(columns = {'value_x':'value'})
df24H.head()
df24H.to_csv("SF24H_labeled.csv")
len(df24H)
from moda.evaluators import get_metrics_for_all_categories, get_final_metrics
from moda.dataprep import read_data
from moda.models import TwitterAnomalyTrendinessDetector, MovingAverageSeasonalTrendinessDetector, \
STLTrendinessDetector, AzureAnomalyTrendinessDetector
def run_model(dataset, freq, min_date='01-01-2018', plot=False, model_name='stl', min_value=10,
min_samples_for_category=100):
if model_name == 'twitter':
model = TwitterAnomalyTrendinessDetector(is_multicategory=True, freq=freq, min_value=min_value, threshold=None,
max_anoms=0.49, seasonality_freq=7)
if model_name == 'ma_seasonal':
model = MovingAverageSeasonalTrendinessDetector(is_multicategory=True, freq=freq, min_value=min_value,
anomaly_type='or',
num_of_std=3)
if model_name == 'stl':
model = STLTrendinessDetector(is_multicategory=True, freq=freq, min_value=min_value,
anomaly_type='residual',
num_of_std=3, lo_delta=0)
if model_name == 'azure':
dirname = os.path.dirname(__file__)
filename = os.path.join(dirname, 'config/config.json')
subscription_key = get_azure_subscription_key(filename)
model = AzureAnomalyTrendinessDetector(is_multicategory=True, freq=freq, min_value=min_value,
subscription_key=subscription_key)
# There is no fit/predict here. We take the entire time series and can evaluate anomalies on all of it or just the last window(s)
prediction = model.predict(dataset, verbose=False)
raw_metrics = get_metrics_for_all_categories(dataset[['value']], prediction[['prediction']], dataset[['label']],
window_size_for_metrics=5)
metrics = get_final_metrics(raw_metrics)
print(metrics)
## Plot each category
if plot:
print("Plotting...")
model.plot(labels=dataset['label'],savefig=False)
return prediction
prediction_stl = run_model(df24H,freq='24H',model_name='stl')
def plot_one_category(category_dataset,model_name='stl'):
def ts_subplot(plt, series, label):
plt.plot(series, label=label, linewidth=0.5)
plt.legend(loc='best')
plt.xticks(rotation=90)
plt.subplot(421, )
ts_subplot(plt, category_dataset['value'], label='Original')
if 'residual_anomaly' in category_dataset:
plt.subplot(422)
ts_subplot(plt, category_dataset['residual_anomaly'], label='Residual anomaly')
if 'trend' in category_dataset:
plt.subplot(423)
ts_subplot(plt, category_dataset['trend'], label='Trend')
if 'trend_anomaly' in category_dataset:
plt.subplot(424)
ts_subplot(plt, category_dataset['trend_anomaly'], label='Trend anomaly')
if 'seasonality' in category_dataset:
plt.subplot(425)
ts_subplot(plt, category_dataset['seasonality'], label='Seasonality')
plt.subplot(426)
ts_subplot(plt, category_dataset['prediction'], label='Prediction')
if 'residual' in category_dataset:
plt.subplot(427)
ts_subplot(plt, category_dataset['residual'], label='Residual')
plt.subplot(428)
ts_subplot(plt, category_dataset['label'], label='Labels')
category = category_dataset.category[0]
plt.suptitle("{} results for category {}".format(model_name, category))
graffiti = prediction_stl.loc[pd.IndexSlice[:, 'Graffiti'], :].reset_index(level='category', drop=False)
fig = plt.figure(figsize=(20,8))
plot_one_category(graffiti,model_name='STL')
```
The time series in this case is relatively noisy. The model was more conservative than the labeler in this case.
```
sewer = prediction_stl.loc[pd.IndexSlice[:, 'Sewer Issues'], :].reset_index(level='category', drop=False)
fig = plt.figure(figsize=(20,8))
plot_one_category(sewer,model_name='STL')
```
In this case, we missed the first peak as we didn't have enough historical data to estimate it. Let's compare this result to a different model:
```
prediction_ma = run_model(df24H,freq=TIME_RANGE,model_name='ma_seasonal')
sewer2 = prediction_ma.loc[pd.IndexSlice[:, 'Sewer Issues'], :].reset_index(level='category', drop=False)
fig = plt.figure(figsize=(20,8))
plot_one_category(sewer2,model_name='MA seasonal')
```
This model estimates the trend differently, and found some anomalies on the trend series as well. It too couldn't detect the first peak as it requires some historical data to estimate standard deviation and other statistics.
| github_jupyter |
## Transfer learning
In this notebook, I will use Transfer learning in our problem.
```
from keras import applications
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras import optimizers
from keras.models import Model
from keras.layers import Dropout, Flatten, Dense, GlobalAveragePooling2D
from keras import backend as k
import os
from PIL import Image
import numpy as np
```
I copy $\textbf{10%}$ of the image to a new folder and this will be my new training set.
```
train_data_dir = "../data/sample_new/train"
validation_data_dir = "../data/validation"
nb_train_samples = sum([len(files) for r, d, files in os.walk(train_data_dir)])
nb_validation_samples = sum([len(files) for r, d, files in os.walk(validation_data_dir)])
NUM_CLASSES = 128
```
All the images should be resized. We will first find the smallest resolution in our datasets.
```
smallest_train= None
for dir_path, _, files in os.walk(train_data_dir):
for name in files:
temp_name = os.path.join(dir_path, name)
temp_shape = np.shape(np.array(Image.open(temp_name)))
if smallest_train == None:
smallest_train = temp_shape
elif temp_shape<smallest_train:
smallest_train = temp_shape
else:
continue
smallest_val = None
for dir_path, _, files in os.walk(validation_data_dir):
for name in files:
temp_name = os.path.join(dir_path, name)
temp_shape = np.shape(np.array(Image.open(temp_name)))
if smallest_val == None:
smallest_val = temp_shape
elif temp_shape<smallest_val:
smallest_val = temp_shape
else:
continue
smallest = min(smallest_train, smallest_val)
img_width, img_height = smallest[0], smallest[1]
```
We will use the ResNet50 as model and we will fine-tune the fully connected last layer of pre-trained network.
```
model = applications.resnet50.ResNet50(include_top=False, weights='imagenet', input_tensor=None, input_shape = (img_width, img_height, 3))
```
Let's have a look to some of the parameters:
1. The `weights='imagenet'` parameter will load the final weights after the model trained.
2. The `include_top=False` parameter instantiates the model without its fully connected layers.
Those layers somehow "translates" (I dunno if this the most correct word but you got my point) the convolutional information into the correct classes. Since ImageNet has 1000 different classes and our dataset 128 we don’t need any of the information in these dense layers. Instead, we will train new dense layers, the last one having 128 nodes and a softmax activation.
```
# Freeze the layers which you don't want to train. Here I am freezing all the layers
for layer in model.layers:
layer.trainable = False
# # add a global spatial average pooling layer
x = model.output
x = GlobalAveragePooling2D()(x)
# let's add a fully-connected layer
x = Dense(1024, activation='relu')(x)
# and a logistic layer
predictions = Dense(NUM_CLASSES, activation='softmax')(x)
# create the full network so we can train on it
transfer_model = Model(input=model.input, output=predictions)
INIT_LR = 5e-3 # initial learning rate
BATCH_SIZE = 32
EPOCHS = 1
# prepare model for fitting (loss, optimizer, etc)
transfer_model.compile(
loss='categorical_crossentropy',
optimizer=keras.optimizers.adamax(lr=INIT_LR),
metrics=['accuracy']
)
# scheduler of learning rate (decay with epochs)
def lr_scheduler(epoch):
return INIT_LR * 0.9 ** epoch
# callback for printing of actual learning rate used by optimizer
class LrHistory(keras.callbacks.Callback):
def on_epoch_begin(self, epoch, logs={}):
print("Learning rate:", k.get_value(transfer_model.optimizer.lr))
```
We will rescale our images
```
train_datagen = ImageDataGenerator(rescale = 1./255)
validation_datagen = ImageDataGenerator(rescale = 1./255)
train_generator = train_datagen.flow_from_directory(
train_data_dir,
target_size = (img_width, img_height),
batch_size = BATCH_SIZE,
shuffle = True,
class_mode = "categorical")
validation_generator = validation_datagen.flow_from_directory(
validation_data_dir,
batch_size = BATCH_SIZE,
target_size = (img_width, img_height),
shuffle = True,
class_mode = "categorical")
```
### And we will fine tune it
```
# Train the model
transfer_model.fit_generator(
train_generator,
steps_per_epoch= nb_train_samples//BATCH_SIZE,
epochs = EPOCHS,
validation_data = validation_generator,
validation_steps = nb_validation_samples//BATCH_SIZE,
callbacks=[keras.callbacks.LearningRateScheduler(lr_scheduler),
LrHistory()],
verbose=1)
```
Using the `10%` of the data I achieve an accuracy of `~49%`. However, it took around 5 hours to complete just one epoch.
| github_jupyter |
```
#python packages pd
import numpy as np
import matplotlib.pyplot as plt
#machine learning packages
import tensorflow as tf
import keras
from keras.models import Sequential
from keras.layers import Dense, Embedding, LSTM, SpatialDropout1D, Bidirectional
from keras.layers import CuDNNLSTM
from keras.utils.np_utils import to_categorical
# from keras.callbacks import EarlyStopping
from keras.layers import Dropout
from sklearn.model_selection import train_test_split
import importlib
#custom python scripts
import generator
import utilis
# Check that you are running GPU's
utilis.GPU_checker()
utilis.aws_setup()
%%time
# generators
importlib.reload(generator)
training_generator = generator.Keras_DataGenerator( dataset='train', w_hyp=False)
validation_generator = generator.Keras_DataGenerator(dataset='valid', w_hyp= False)
#Constants
# ARE YOU LOADINNG MODE?
# VOCAB_SIZE = 1254
# INPUT_LENGTH = 1000
# EMBEDDING_DIM = 128
# # model
# def build_model(vocab_size, embedding_dim, input_length):
# model = Sequential()
# model.add(Embedding(vocab_size, embedding_dim, input_length=input_length))
# model.add(SpatialDropout1D(0.2))
# model.add(CuDNNLSTM(128))
# model.add(Dense(41, activation='softmax'))
# return model
# model = build_model(VOCAB_SIZE, EMBEDDING_DIM, INPUT_LENGTH)
# model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# print(model.summary())
## WARNING IF YOU CONTAIN MULTIPLE CORE GPUS
# NOTE unclear if these causes a speed up
# @TANCREDI, I HAVE TREID THIS ON JUST GOALS AND DOES NOT SEEM TO CAUSE A SPEED UP MAY
#CAUSE A SPEED UP IF WE USE HYPOTHESIS
# unclea rif this seepd
# from keras.utils import multi_gpu_model
# model_GPU = multi_gpu_model(model, gpus= 4)
#model_GPU.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
## ARE YOU LOADING A MODEL IF YES RUN TEH FOLLOWING LINES
from keras.models import model_from_json
json_file = open('model.json', 'r')
loaded_model_json = json_file.read()
json_file.close()
loaded_model = model_from_json(loaded_model_json)
# load weights into new model
loaded_model.load_weights("model.h5")
print("Loaded model from disk")
# REMEMEBER TO COMPILE
loaded_model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
#overwriting model
model = loaded_model
print(model.summary())
%%time
n_epochs = 2
history = model.fit_generator(generator=training_generator,
validation_data=validation_generator,
verbose=1,
use_multiprocessing= False,
epochs=n_epochs)
# FOR SAVING MODEL
model_json = model.to_json()
with open("model.json", "w") as json_file:
json_file.write(model_json)
# serialize weights to HDF5
model.save_weights("model.h5")
print("Saved model to disk")
#WARNING_DECIDE_HOW_TO_NAME_LOG
#descriptionofmodel_personwhostartsrun
#e.g. LSTM_128encoder_etc_tanc
LOSS_FILE_NAME = "SIMPLE_LSTM_SMALL_TANK"
#WARNING NUMBER 2 - CURRENTLY EVERYTIME YOU RERUN THE CELLS BELOW THE FILES WITH THOSE NAMES GET WRITTEN OVER
# save history - WARNING FILE NAME
utilis.history_saver_bad(history, LOSS_FILE_NAME)
# read numpy array
# history_toplot = np.genfromtxt("training_logs/"+ LOSS_FILE_NAME +".csv")
# plt.plot(history_toplot)
# plt.title('Loss history')
# plt.show()
%%time
n_epochs = 1
history = loaded_model.fit_generator(generator=training_generator,
validation_data=validation_generator,
verbose=1,
use_multiprocessing= False,
epochs=n_epochs)
```
| github_jupyter |
# Box-Cox Tranformation
### Box-Cox tranformation is a simple generalization of the square root transformation and the log transformation.
## $$\hat{x} = \begin{cases}
\frac{x^{\lambda} - 1}{\lambda} & if \lambda \neq 0 \\
ln(x) & if \lambda = 0\\ \end{cases}$$
### The Box-Cox transformation does only work with positive data. Therefore, sometimes you need to shift the data with adding a fixed constant. The power of $\lambda$ needs to be tuned that the box-cox module in scipy package does this process for you.
```
# Loading Libraries
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import matplotlib.pylab as pylab
import seaborn as sns
from scipy import stats
sns.set_style("ticks")
plot_params = {
"legend.fontsize": "x-large",
"figure.figsize": (10, 6),
"axes.labelsize": "x-large",
"axes.titlesize":"x-large",
"xtick.labelsize":"x-large",
"ytick.labelsize":"x-large",
"axes.linewidth" : 3,
"lines.linewidth" : 2
}
pylab.rcParams.update(plot_params)
x = np.linspace(0.001 , 3., 100)
lambda_list = [0., 0.25, 0.50, 0.75, 1., 1.25, 1.50]
c = ["k", "royalblue", "plum", "aqua", "gold", "tomato", "darkkhaki"]
plt.figure(figsize=(10,6))
for l in lambda_list:
plt.plot(x, stats.boxcox(x, lmbda = l), lw = 3, ls = "--", c = c[lambda_list.index(l)], label = F"$\lambda = {l}$")
plt.legend(loc = 0 , prop={'size': 14})
plt.xlabel("X", fontsize = 20)
plt.ylabel(r"$\hat{X}$", fontsize = 20)
plt.title(r"Box-Cox Transformation for different values of $\lambda$", fontsize = 20)
plt.show()
# using stats module in scipy
from scipy import stats
# non-normally distrubuted data
x = stats.loggamma.rvs(2, size = 1000) + 4
xt, lambda_optimal = stats.boxcox(x)
plt.figure(figsize = (10, 5))
plt.subplot(1,2,1)
plt.hist(x , color = "navy")
plt.title("X")
plt.subplot(1,2,2)
plt.hist(xt , color = "navy")
plt.title("X-transformed with Box-Cox")
plt.show()
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
stats.boxcox_normplot(x, -20,20, plot = ax, N = 200);
ax.axvline(lambda_optimal, lw = 2, color='r', label = F"$\lambda = {lambda_optimal:.4}$");
plt.legend(loc = 0 , prop={'size': 14})
plt.show()
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
res = stats.probplot(x, dist=stats.loggamma, sparams=(5,), plot=ax)
ax.set_title("Probplot for loggamma dist with shape parameter 5", fontsize = 14);
```
| github_jupyter |
### Imports
```
import pandas as pd
import numpy as np
import sys
import os
import math
import matplotlib.pyplot as plt
from terminaltables import AsciiTable
from sklearn.preprocessing import normalize
cmap = plt.get_cmap('viridis')
PATH_TO_DATA = '../data'
```
### Load dataset
```
assert os.path.exists('%s/dataset' % PATH_TO_DATA), \
"Please execute the notebook 'build_feature_set.ipynb' before running this notebook."
dataset = pd.read_pickle('%s/dataset' % PATH_TO_DATA)
```
### Display correlation matrix
```
blog_i = list(dataset.columns).index("blog")
# Normalize the data
dataset_norm = (dataset - dataset.mean()) / (dataset.max() - dataset.min())
correlation_matrix = dataset_norm.corr().values.round(5)
# Plot the correlation matrix
fig = plt.figure(figsize=(14, 12))
ax = fig.add_subplot(111)
cax = ax.matshow(dataset_norm.corr(), cmap=cmap, interpolation='nearest')
# Add title to matrix plot
ax.set_title("Correlation Matrix")
ttl = ax.title
ttl.set_position([.5, 1.1])
# Add feature names to axises
plt.xticks(range(dataset_norm.shape[1]), dataset_norm.columns, rotation=45)
plt.yticks(range(dataset_norm.shape[1]), dataset_norm.columns)
# Add color bar to the right of the matrix
fig.colorbar(cax)
plt.show()
plt.close()
# Extract the blog feature correlations
target_corr = correlation_matrix[:, blog_i]
print("Blog correlation:")
# Show feature table with correlations to blog articles
table_data = [["Num.", "Feature", "Blog Correlation"]]
for i, col in enumerate(dataset.columns):
table_data.append([i, col, target_corr[i]])
print (AsciiTable(table_data).table)
print ("")
# Extract the blog feature correlations
target_corr_sorted = sorted(correlation_matrix[:, blog_i], key=abs, reverse=True)
print ("Ordered by highest absolute correlalation:")
# Show feature table with correlations to blog articles
table_data = [["Num.", "Feature", "Blog Correlation"]]
for corr in target_corr_sorted:
i = np.argmax(target_corr == corr)
col = dataset.columns[i]
table_data.append([i, col, corr])
print (AsciiTable(table_data).table)
print ("")
# Find the features with highest blog correlation
target_highest = np.argsort(target_corr)[::-1]
colors = [cmap(i) for i in np.linspace(0, 1, len(target_highest)-1)][::-1]
x_labels = [dataset.columns[i] for i in target_highest]
plt.figure(figsize=(12, 8))
plt.title("Feature correlation with Blog")
plt.bar(range(len(target_highest)-1), target_corr[target_highest][1:], color=colors)
plt.xticks(range(len(dataset.columns)-1), x_labels[1:], rotation=45)
plt.show()
plt.close()
```
### Compare feature values between distributions
```
feature_names = [col for col in dataset.columns if col != "blog"]
# Extract news and blog articles respectively
news_articles = dataset[dataset['blog'] == 0][feature_names]
blog_articles = dataset[dataset['blog'] == 1][feature_names]
# Get mean values for each feature of each media type
means = list(zip(news_articles.mean(axis=0), blog_articles.mean(axis=0)))
colors = [cmap(0.1), cmap(0.9)]
# Plot grid
n_cols = 4
n_rows = math.ceil(len(feature_names)/n_cols)
fw, fh = n_cols*3, n_rows*2
fig, axes = plt.subplots(n_rows, n_cols, figsize=(fw, fh))
fig.tight_layout()
for i in range(len(feature_names)):
subplot = plt.subplot(n_rows, n_cols, i+1)
subplot.set_title(feature_names[i].capitalize())
subplot.bar(("News", "Blog"), means[i], color=colors)
plt.tight_layout()
plt.show()
plt.close()
```
| github_jupyter |
#### Outline
- for each dataset:
- load dataset;
- for each network:
- load network
- project 1000 test dataset samples
- save to metric dataframe
```
# reload packages
%load_ext autoreload
%autoreload 2
```
### Choose GPU (this may not be needed on your computer)
```
%env CUDA_DEVICE_ORDER=PCI_BUS_ID
%env CUDA_VISIBLE_DEVICES=0
import tensorflow as tf
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
if len(gpu_devices)>0:
tf.config.experimental.set_memory_growth(gpu_devices[0], True)
import numpy as np
import pickle
import pandas as pd
import time
from umap import UMAP
from tfumap.umap import tfUMAP
import tensorflow as tf
from sklearn.decomposition import PCA
from openTSNE import TSNE
from tqdm.autonotebook import tqdm
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
output_dir = MODEL_DIR/'projections'
projection_speeds = pd.DataFrame(columns = ['method_', 'dimensions', 'dataset', 'speed', "nex"])
```
### macosko2015
```
dataset = 'macosko2015'
dims = [50]
```
##### load dataset
```
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
#dataset_address = 'http://file.biolab.si/opentsne/macosko_2015.pkl.gz'
# https://opentsne.readthedocs.io/en/latest/examples/01_simple_usage/01_simple_usage.html
# also see https://github.com/berenslab/rna-seq-tsne/blob/master/umi-datasets.ipynb
import gzip
import pickle
with gzip.open(DATA_DIR / 'macosko_2015.pkl.gz', "rb") as f:
data = pickle.load(f)
x = data["pca_50"]
y = data["CellType1"].astype(str)
print("Data set contains %d samples with %d features" % x.shape)
from sklearn.model_selection import train_test_split
def zero_one_norm(x):
return (x- np.min(x, axis=0))/ (np.max(x, axis=0)-np.min(x, axis=0))
x_norm = zero_one_norm(x)
X_train, X_test, Y_train, Y_test = train_test_split(x_norm, y, test_size=.1, random_state=42)
np.shape(X_train)
n_valid = 10000
X_valid = X_train[-n_valid:]
Y_valid = Y_train[-n_valid:]
X_train = X_train[:-n_valid]
Y_train = Y_train[:-n_valid]
X_train_flat = X_train
from sklearn.preprocessing import OrdinalEncoder
enc = OrdinalEncoder()
Y_train = enc.fit_transform([[i] for i in Y_train]).flatten()
X_train_flat = X_train
X_test_flat = X_test
print(len(X_train), len(X_test), len(X_valid))
```
#### Network
##### 2 dims
```
load_loc = output_dir / dataset / 'network'
embedder = tfUMAP(
direct_embedding=False,
verbose=True,
negative_sample_rate=5,
training_epochs=5,
batch_size = 100,
dims = dims
)
encoder = tf.keras.models.load_model((load_loc / 'encoder').as_posix())
embedder.encoder = encoder
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
encoder(X_test);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['network', 2, dataset, end_time - start_time, len(X_test)]
##### Network CPU
with tf.device('/CPU:0'):
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
encoder(X_test);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['network-cpu', 2, dataset, end_time - start_time, len(X_test)]
```
##### 64 dims
```
load_loc = output_dir / dataset /"64"/ 'network'
embedder = tfUMAP(
direct_embedding=False,
verbose=True,
negative_sample_rate=5,
training_epochs=5,
batch_size = 100,
dims = dims
)
encoder = tf.keras.models.load_model((load_loc / 'encoder').as_posix())
embedder.encoder = encoder
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
encoder(X_test);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['network', 64, dataset, end_time - start_time, len(X_test)]
##### Network CPU
with tf.device('/CPU:0'):
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
encoder(X_test);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['network-cpu', 64, dataset, end_time - start_time, len(X_test)]
```
#### UMAP-learn
##### 2 dims
```
embedder = UMAP(n_components = 2, verbose=True)
z_umap = embedder.fit_transform(X_train_flat)
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
embedder.transform(X_test_flat);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['umap-learn', 2, dataset, end_time - start_time, len(X_test)]
projection_speeds
```
##### 64 dims
```
embedder = UMAP(n_components = 64, verbose=True)
z_umap = embedder.fit_transform(X_train_flat)
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
embedder.transform(X_test_flat);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['umap-learn', 64, dataset, end_time - start_time, len(X_test)]
projection_speeds
```
#### PCA
##### 2 dims
```
pca = PCA(n_components=2)
z = pca.fit_transform(X_train_flat)
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
pca.transform(X_test_flat);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['pca', 2, dataset, end_time - start_time, len(X_test)]
projection_speeds
```
##### 64 dims
```
x_train_flat_padded = np.concatenate([X_train_flat, np.zeros((len(X_train_flat), 14))], axis=1)
X_test_flat_padded = np.concatenate([X_test_flat, np.zeros((len(X_test_flat), 14))], axis=1)
pca = PCA(n_components=64)
z = pca.fit_transform(x_train_flat_padded)
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
pca.transform(X_test_flat_padded);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['pca', 64, dataset, end_time - start_time, len(X_test)]
projection_speeds
```
#### TSNE
##### 2 dims
```
tsne = TSNE(
n_components = 2,
n_jobs=32,
verbose=True
)
embedding_train = tsne.fit(X_train_flat)
n_repeats = 10
times = []
for i in tqdm(range(n_repeats)):
start_time = time.monotonic()
embedding_train.transform(X_test_flat);
end_time = time.monotonic()
print('seconds: ', end_time - start_time)
times.append(end_time - start_time)
projection_speeds.loc[len(projection_speeds)] = ['TSNE', 2, dataset, end_time - start_time, len(X_test)]
projection_speeds
```
### Save
```
save_loc = DATA_DIR / 'projection_speeds' / (dataset + '.pickle')
ensure_dir(save_loc)
projection_speeds.to_pickle(save_loc)
```
| github_jupyter |
Exercise 7 - Advanced Support Vector Machines
===
Support vector machines (SVMs) let us predict categories. In this exercise, we will be using SVM, paying attention to the key steps as we go: formatting data correctly, splitting the data into training and test sets, training an SVM model using the training set, and then evaluating and visualising the SVM model using the test set.
We will be looking at __prions__: misfolded proteins that are associated with several fatal neurodegenerative diseases (kind of like Daleks, if you have seen Doctor Who). Looking at examples of protein mass and weight, we will build a predictive model to detect prions in blood samples.
#### Run the code below to load the required libraries for this exercise.
```
# Load `tidyverse` package
suppressMessages(install.packages("tidyverse"))
suppressMessages(library("tidyverse"))
suppressMessages(install.packages("e1071"))
suppressMessages(library("e1071"))
suppressMessages(install.packages("magrittr"))
suppressMessages(library("magrittr"))
```
Step 1
---
Let's load the required R packages and the prion data for this exercise.
**In the code below, complete the data loading step, by replacing `<prionDataset>` with `prion_data`, and running the code.**
```
###
# REPLACE <prionDataset> WITH prion_data
###
<prionDataset> <- read.csv("Data/PrionData.csv")
###
# Check the structure of `prion_data`
str(prion_data)
head(prion_data)
```
It appears that we have an extra column `X` in `prion_data` that contains the row number. By default, R has labelled the column `X` because the input didn't have a column name (it was blank). This behaviour happens regularly when exporting data sets from a program like Microsoft Excel and then importing them into R.
Let's get rid of the first column from `prion_data`, and then check that it has been successfully removed. We will use the `select` function from the `dplyr` package together with the `-` symbol to "minus" the `X` column from our dataset.
> **N.B. We have used a different assignment symbol `%<>%` from the `magrittr` package in the code below. The `magrittr` assignment symbol `%<>%` is a combination of the `magrittr` pipe symbol `%>%` and the base R assignment symbol `<-`. It takes the variable on the left hand side of the `%<>%` symbol, and updates the value of the variable with the result of the right hand side. So the object on the left hand side acts as both the initial value and the resulting value.**
#### Replace `<removeColumn>` with `-X` to remove the excess column X, then run the code.
```
###
# REPLACE <removeColumn> WITH -X
###
prion_data %<>% select(<removeColumn>)
###
str(prion_data)
head(prion_data)
# Check frequency of `prion_status` in `prion_data`
prion_data %>%
group_by(prion_status) %>%
summarise(n = n()) %>%
mutate(freq = n/sum(n))
```
Excellent, we have successfully removed column `X` from `prion_data`!
Now, looking at the output of `str` and `head`, we can observe that `prion_data` is a `data.frame` that contains 485 observations and 3 variables stored in the following columns:
* `mass` is the first *feature*;
* `weight` is the second *feature*;
* `prion_status` is the *label* (or category).
Of the 485 observations, 375 (77.31%) are non-prions, and 110 (22.68%) are prions.
Step 2
---
Let's graph `prion_data` to better understand the features and labels.
**In the cell below replace:**
**1. `<xData>` with `mass`**
**2. `<yData>` with `weight`**
**3. `<colorData>` with `prion_status`**
** then __run the code__. **
```
prion_data %>%
###
# REPLACE <xData> WITH mass AND <yData> WITH weight AND <colorData> WITH prion_status
###
ggplot(aes(x = <xData> , y = <yData> , colour = <colorData> )) +
###
geom_point() +
ggtitle("Classification plot for prion data") +
# Create labels for x-axis, y-axis, and legend
labs(x = "Mass", y = "Weight", colour = "Prion status") +
# Align title to centre
theme(plot.title = element_text(hjust = 0.5))
```
Step 3
---
To create a SVM model, let's split our data into training and test sets. We'll start by checking the total number of instances in our data set. If we go back to the output from `str(prion_data)` in Step 2, we have 485 observations and 3 variables.
So, let's use 400 examples for our `training` set, and the remainder for our `test` set.
We will use the `slice` function to select the first 400 rows from `prion_data`
#### Replace `<selectData>` with `1:400`, and run the code.
```
###
# REPLACE <selectData> WITH 1:400
###
train_prion <- slice(prion_data, <selectData>)
###
str(train_prion)
# Check percentage of samples that are prions
train_prion %>%
group_by(prion_status) %>%
summarise(n = n()) %>%
mutate(freq = n/sum(n))
# Create test set using the remaining examples
test_prion <- slice(prion_data, 401:n())
str(test_prion)
# Check percentage of samples that are prions
test_prion %>%
group_by(prion_status) %>%
summarise(n = n()) %>%
mutate(freq = n/sum(n))
```
Well done! Let's look at a summary of our training data to get a better idea of what we're dealing with.
#### Replace `<trainDataset>` with `train_prion` and run the code.
```
###
# REPLACE <trainDataset> WITH train_prion
###
summary(<trainDataset>)
###
```
Using the `summary` function, we observe our training data contains 314 non-prions and 86 prions out of a total of 400 observations. This looks right, because the scatter plot we created in Step 2 showed us the majority of observations have 'non-prion' status.
Let's take a look at `test_prion` too, using the `summary` function again.
#### Replace `<testDataset>` with `test_prion` and run the code.
```
###
# REPLACE <testDataset> WITH test_prion
###
summary(<testDataset>)
###
```
Looking good! Alright, now to make a support vector machine.
Step 4
---
Below we will make an SVM similar to the previous exercise. Remember the syntax for SVMs using the `e1071::svm` function:
**`svm_model <- svm(x = x, y = y, data = dataset)`**
where `x` represents the features (a matrix), and `y` represents the labels (factors).
Alternatively, we can use the following syntax for the `svm` function:
**`model <- svm(formula = y ~ x, data = dataset)`**
where `y` represents the labels/categories, and `x` represents the features. Note if you have multiple `x` features in the dataset, you can simply type `.` in the `formula` argument to refer to everything in the data set except the y argument. Let's try out this syntax using the training data as our input.
**In the code below, create an SVM model using the `train_prion` data using the `svm` function with the `formula` argument.**
#### Replace `<dataSelection>` with `prion_status ~ .`, then run the code.
Note: the `prion_status` on the left hand side of the formula selects our labels, and the `.` on the right hand side of the formula selects our features. In this case, the `.` selects all the features in our dataset `train_prion`.
```
###
# REPLACE <dataSelection> WITH prion_status ~ .
###
SVM_Model <- svm(formula = <dataSelection> , data = train_prion)
###
print("Model ready!")
```
Well done! We've made a SVM model using our training set `train_prion`.
Step 5
---
Let's create some custom functions to graph and evaluate SVM models. We will use these functions throughout the remainder of this exercise. You do not need to edit the code block below.
**Run the code below**
```
# Run this box to prepare functions for later use
# Create a custom function named `Graph_SVM` to plot an SVM model
Graph_SVM <- function(model, data_set){
grid <- expand.grid("mass" = seq(min(data_set$mass), max(data_set$mass), length.out = 100),
"weight" = seq(min(data_set$weight), max(data_set$weight), length.out = 100))
preds <- predict(model, grid)
df <- data.frame(grid, preds)
ggplot() +
geom_tile(data = df, aes(x = mass, y = weight, fill = preds)) +
geom_point(data = data_set, aes(x = mass, y = weight, shape = prion_status,
colour = prion_status),
alpha = 0.75) +
scale_colour_manual(values = c("grey10", "grey50")) +
labs(title = paste("SVM model prediction"), x = "Mass", y = "Weight",
fill = "Prediction", shape = "Prion status", colour = "Prion status") +
theme(plot.title = element_text(hjust = 0.5))
}
# Create another custom function named `Evaluate_SVM` to evaluate the SVM model, print results to screen,
# and run the `Graph_SVM` custom function
Evaluate_SVM <- function(model, data_set){
predictions <- predict(model, data_set)
total <- 0
for(i in 1:nrow(data_set)){
if(toString(predictions[i]) == data_set[i, "prion_status"]){
total = total + 1
}
}
# Print results to screen
print("SVM Model Evaluation")
print(paste0("Model name: ", deparse(substitute(model))))
print(paste0("Dataset: ", deparse(substitute(data_set))))
print(paste0("Accuracy: ", total/nrow(data_set)*100, "%"))
print(paste0("Number of samples: ", nrow(data_set)))
# Call our custom function for graphing SVM model
Graph_SVM(model, data_set)
}
print("Custom function ready!")
```
Excellent! Now that we have created the custom function `Evaluate_SVM` (which incorporates the `Graph_SVM` function) let's evaluate our SVM model on the training data.
In the code below, we will change the inputs to the `Evaluate_SVM` function, where the first argument is the SVM model we will evaluate, and the second argument is the dataset we will evaluate the SVM model with.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model` **
** 2. `<dataset>` with `train_prion` **
** Then __run the code__. **
```
###
# REPLACE <svmModel> WITH SVM_Model AND <dataset> WITH train_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
Step 6
---
The SVM has performed reasonably well separating our training data set into two. Now let's take a look at our test set.
In the code below, we will use our custom function `Evaluate_SVM` to evaluate `SVM_Model` on the test set.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model` **
** 2. `<dataset>` with `test_prion` **
** Then __run the code__. **
```
###
# REPLACE <svmModel> WITH SVM_Model AND <dataset> WITH test_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
That's a good result.
Conclusion
---
Well done! We've taken a data set, tidied it, prepared it into training and test sets, created an SVM based on the training set, and evaluated the SVM model using the test set.
You can go back to the course now, or you can try using different kernels with your SVM below.
OPTIONAL: Step 8
---
Want to have a play around with different kernels for your SVM models? It's really easy!
The standard kernel is a radial basis kernel. But there's a few more you can choose from: `linear`, `polynomial`, and `sigmoid`. Let's try them out.
If you want to use a linear kernel, all you need to do is add `kernel = "linear"` to your model. Like this:
`SVM_Model <- svm(formula = y ~ x, data = dataset, kernel = "linear")`
Give it a go with all the different kernels below. The first kernel, `linear`, has been done for you.
**Run the code below**
```
# Run this box to make a linear SVM
# Make a linear SVM model
SVM_Model_Linear <- svm(prion_status ~ . , data = train_prion, kernel = "linear")
print("Model ready")
```
Now we have created the linear SVM model, let's evaluate it on our training and test sets using our custom function we created earlier, `Evaluate_SVM`. Remember the inputs to `Evaluate_SVM` are the SVM model followed by the data you wish to evaluate the model on.
In the code blocks below, we will change the inputs to our `Evaluate_SVM` function to the appropriate variable names to evaluate the linear SVM model on the training and test sets.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model_Linear` **
** 2. `<dataset>` with `train_prion` **
** Then __run the code__. **
```
# Evaluate linear SVM model on training set
###
# REPLACE <svmModel> WITH SVM_Model_Linear AND <dataset> WITH train_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
And now for the test set.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model_Linear` **
** 2. `<dataset>` with `test_prion` **
** Then __run the code__. **
```
# Evaluate linear SVM model on test set
###
# REPLACE <svmModel> WITH SVM_Model_Linear AND <dataset> WITH test_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
You can see the hyperplane is a linear line! Compare the linear SVM model results to the radial SVM model results to see the difference for yourself!
## Now let's try a sigmoid kernel.
** In the cell below replace: **
** 1. `<kernelSelection>` with `"sigmoid"` **
** 2. `<svmModel>` with `SVM_Model_Sigmoid` **
** 3. `<dataset>` with `train_prion` **
** Then __run the code__. **
```
###
# REPLACE <kernelSelection> WITH "sigmoid" (INCLUDING THE QUOTATION MARKS)
###
SVM_Model_Sigmoid <- svm(prion_status ~ . , data = train_prion, kernel = <kernelSelection>)
###
# Evaluate sigmoid SVM model on training set
###
# REPLACE <svmModel> WITH SVM_Model_Sigmoid AND <dataset> WITH train_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
And now for the test set.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model_Sigmoid` **
** 2. `<dataset>` with `test_prion` **
** Then __run the code__. **
```
# Evaluate sigmoid SVM model on test set
###
# REPLACE <svmModel> WITH SVM_Model_Sigmoid AND <dataset> WITH test_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
Perhaps a sigmoid kernel isn't a good idea for this data set....
## Let's try a sigmoid kernel instead.
** In the cell below replace: **
** 1. `<kernelSelection>` with `"polynomial"` **
** 2. `<svmModel>` with `SVM_Model_Sigmoid` **
** 3. `<dataset>` with `train_prion` **
** Then __run the code__. **
```
###
# REPLACE <kernelSelection> WITH "polynomial" (INCLUDING THE QUOTATION MARKS)
###
SVM_Model_Poly <- svm(prion_status ~ . , data = train_prion, kernel = <kernelSelection>)
# Evaluate polynomial SVM model on training set
###
# REPLACE <svmModel> WITH SVM_Model_Poly AND <dataset> WITH train_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
And now for the test set.
** In the cell below replace: **
** 1. `<svmModel>` with `SVM_Model_Poly` **
** 2. `<dataset>` with `test_prion` **
** Then __run the code__. **
```
# Evaluate polynomial SVM model on test set
###
# REPLACE <svmModel> WITH SVM_Model_Poly AND <dataset> WITH test_prion
###
Evaluate_SVM(<svmModel>, <dataset>)
###
```
If we were to carry on analysing prions like this, a polynomial SVM looks like a good choice (based on the performance of the different models on `test_prion`). If the data set was more complicated, we could try different degrees for the polynomial to see which one was the most accurate. This is part of __`tuning`__ a model.
Well done!
| github_jupyter |
<a href="https://colab.research.google.com/github/jwkanggist/EverybodyTensorflow2.0/blob/master/lab15_alexnet_cifar100_tf2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#LAB15: alexnet for cifar100
Alexnet을 구현해보자
- Cifar100 datatset을 위한 실험
```
# preprocessor parts
from __future__ import absolute_import, division, print_function, unicode_literals
# Install TensorFlow
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
from tensorflow.keras.callbacks import TensorBoard
import numpy as np
import pandas as pd
import cv2
import numpy as np
from matplotlib import cm
from matplotlib import gridspec
import matplotlib.pyplot as plt
from datetime import datetime
# for Tensorboard use
LOG_DIR = 'drive/data/tb_logs'
!wget https://bin.equinox.io/c/4VmDzA7iaHb/ngrok-stable-linux-amd64.zip
!unzip ngrok-stable-linux-amd64.zip
import os
if not os.path.exists(LOG_DIR):
os.makedirs(LOG_DIR)
get_ipython().system_raw(
'tensorboard --logdir {} --host 0.0.0.0 --port 6006 &'
.format(LOG_DIR))
get_ipython().system_raw('./ngrok http 6006 &')
!curl -s http://localhost:4040/api/tunnels | python3 -c \
"import sys, json; print(json.load(sys.stdin)['tunnels'][0]['public_url'])"
# dataset loading part
# 데이터 파이프라인 부분
cifar100 = tf.keras.datasets.cifar100
(x_train, y_train), (x_test, y_test) = cifar100.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
print("x_train.shape = %s" % str(x_train.shape))
print("y_train.shape = %s" % str(x_train.shape))
x_train = x_train.reshape([x_train.shape[0],
x_train.shape[1],
x_train.shape[2],3])
x_test = x_test.reshape([x_test.shape[0],
x_test.shape[1],
x_test.shape[2],3])
x_train = np.array(tf.keras.backend.resize_images(x_train, height_factor=2,width_factor=2, data_format='channels_last',interpolation='bilinear'))
x_test = np.array(tf.keras.backend.resize_images(x_test, height_factor=2,width_factor=2, data_format='channels_last',interpolation='bilinear'))
# x_train = cv2.resize(x_train, dsize=(64,64), interpolation=cv2.INTER_CUBIC)
# x_test = cv2.resize(x_test, dsize=(64,64), interpolation=cv2.INTER_CUBIC)
print("x_train.shape = %s" % str(x_train.shape))
print("x_test.shape = %s" % str(x_test.shape))
print("y_train.shape = %s" % str(y_train.shape))
print("y_test.shape = %s" % str(y_test.shape))
# Network Parameters
dropout_rate =0.1
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(filters=24,kernel_size=(11,11),strides=(1,1),activation='relu',padding='same'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D(pool_size=(3,3),strides=(2,2),padding='valid'),
tf.keras.layers.Conv2D(filters=64,kernel_size=(5,5),strides=(1,1),activation='relu',padding='same'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.MaxPool2D(pool_size=(3,3),strides=(2,2),padding='valid'),
tf.keras.layers.Conv2D(filters=96,kernel_size=(3,3),strides=(1,1),activation='relu',padding='same'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(filters=96,kernel_size=(3,3),strides=(1,1),activation='relu',padding='same'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Conv2D(filters=64,kernel_size=(3,3),strides=(1,1),activation='relu',padding='same'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(units=1024,activation='relu'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(units=1024,activation='relu'),
tf.keras.layers.Dropout(dropout_rate),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(units=100,activation='softmax')
])
opt_fn = tf.keras.optimizers.Adam(learning_rate=1e-3,
beta_1=0.9,
beta_2=0.999)
# 'sparse_categorical_crossentropy' is for integer labels
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
tensorboard_callback = TensorBoard(log_dir=LOG_DIR,
histogram_freq=1,
write_graph=True,
write_images=True)
# model training and evaluation part
training_epochs = 20
batch_size = 128
model.fit(x_train, y_train,
epochs=training_epochs,
validation_data=(x_test, y_test),
batch_size=batch_size,
callbacks=[tensorboard_callback])
model.evaluate(x_test, y_test, verbose=2)
# prediction
test_input = x_test[300,:,:,:]
pred_y = model.predict(test_input.reshape([1,64,64,3]))
plt.figure(1)
plt.imshow(test_input.reshape([64,64,3]))
plt.title("input")
print("model prediction = %s"% pred_y.argmax())
```
| github_jupyter |
# BicycleGAN
## 1. Import Libs
```
import torch
import torch.nn as nn
from torch.autograd import Variable
import torch.optim as optim
import torchvision
from torchvision import transforms
from torch.utils.data import DataLoader
from dataloader import Edges2handbags
import util
import os
import argparse
from model import *
CUDA = torch.cuda.is_available()
```
## 2. Settings
```
parser = argparse.ArgumentParser([])
parser.add_argument('--root', required=False, default='./dataset/edges2handbags/edges2handbags')
parser.add_argument('--result_dir', default='result/edges2handbags')
parser.add_argument('--weight_dir', default='weight/edges2handbags')
parser.add_argument('--load_weight', type=bool, default=False)
parser.add_argument('--batch_size', type=int, default=2)
parser.add_argument('--test_size', type=int, default=20)
parser.add_argument('--test_img_num', type=int, default=5)
parser.add_argument('--img_size', type=int, default=128)
parser.add_argument('--num_epoch', type=int, default=50)
parser.add_argument('--save_every', type=int, default=1000)
parser.add_argument('--lr', type=float, default=0.0002)
parser.add_argument('--beta_1', type=float, default=0.5)
parser.add_argument('--beta_2', type=float, default=0.999)
parser.add_argument('--lambda_kl', type=float, default=0.01)
parser.add_argument('--lambda_img', type=int, default=10)
parser.add_argument('--lambda_z', type=float, default=0.5)
parser.add_argument('--z_dim', type=int, default=8)
params = parser.parse_args([])
print(params)
```
## 3. Utils
```
def mse_loss(score, target=1):
dtype = type(score)
if target == 1:
label = util.var(torch.ones(score.size()), requires_grad=False)
elif target == 0:
label = util.var(torch.zeros(score.size()), requires_grad=False)
criterion = nn.MSELoss().cuda()
loss = criterion(score, label)
return loss
def L1_loss(pred, target):
return torch.mean(torch.abs(pred - target))
def lr_decay_rule(epoch, start_decay=100, lr_decay=100):
decay_rate = 1.0 - (max(0, epoch - start_decay) / float(lr_decay))
return decay_rate
```
## 4. Load Dataset
### 4.1 Preprocessing
```
transform = transforms.Compose([
transforms.Scale((params.img_size, params.img_size)),
transforms.ToTensor(),
transforms.Normalize(mean=(0.5, 0.5, 0.5), std=(0.5, 0.5, 0.5))])
```
### 4.2 DataLoader
```
train_dataset = Edges2handbags(params.root, transform=transform, mode='train')
test_dataset = Edges2handbags(params.root, transform=transform, mode='val')
train_data_loader = DataLoader(train_dataset, batch_size=params.batch_size, shuffle=True, num_workers=4, drop_last=True)
test_data_loader = DataLoader(test_dataset, batch_size=params.batch_size, shuffle=True, num_workers=4, drop_last=True)
train_sample = train_dataset.__getitem__(11)
print(train_sample)
print(len(train_sample))
```
## 5. Build Models & Optimizers & Loss Function
### 5.1 Models
```
D_cVAE = Discriminator().cuda()
D_cLR = Discriminator().cuda()
G = Generator(z_dim=params.z_dim).cuda()
E = Encoder(z_dim=params.z_dim).cuda()
print(D_cVAE)
print(D_cLR)
print(G)
print(E)
```
### 5.2 Optimizers
```
optim_D_cVAE = optim.Adam(D_cVAE.parameters(), lr=params.lr, betas=(params.beta_1, params.beta_2))
optim_D_cLR = optim.Adam(D_cLR.parameters(), lr=params.lr, betas=(params.beta_1, params.beta_2))
optim_G = optim.Adam(G.parameters(), lr=params.lr, betas=(params.beta_1, params.beta_2))
optim_E = optim.Adam(E.parameters(), lr=params.lr, betas=(params.beta_1, params.beta_2))
# 20 x 5 x 8
fixed_z = util.var(torch.randn(params.test_size, params.test_img_num, params.z_dim)).cuda()
```
## 6. Load Pretrained Models
```
if params.load_weight is True:
D_cVAE.load_state_dict(torch.load(os.path.join(params.weight_dir, 'D_cVAE.pkl')))
D_cLR.load_state_dict(torch.load(os.path.join(params.weight_dir, 'D_cLR.pkl')))
G.load_state_dict(torch.load(os.path.join(params.weight_dir, 'G.pkl')))
E.load_state_dict(torch.load(os.path.join(params.weight_dir, 'E.pkl')))
D_cVAE.train()
D_cLR.train()
G.train()
E.train()
for epoch in range(1, params.num_epoch + 1):
for iters, (img, ground_truth) in enumerate(train_data_loader): # 1
# img : (2, 3, 128, 128) of domain A / ground_truth : (2, 3, 128, 128) of domain B
img, ground_truth = util.var(img), util.var(ground_truth)
# Seperate data for cVAE_GAN and cLR_GAN
cVAE_data = {'img' : img[0].unsqueeze(dim=0), 'ground_truth' : ground_truth[0].unsqueeze(dim=0)}
cLR_data = {'img' : img[1].unsqueeze(dim=0), 'ground_truth' : ground_truth[1].unsqueeze(dim=0)}
''' ----------------------------- 1. Train D ----------------------------- '''
############# Step 1. D loss in cVAE-GAN #############
# Encoded latent vector
mu, log_variance = E(cVAE_data['ground_truth'])
std = torch.exp(log_variance / 2)
random_z = util.var(torch.randn(1, params.z_dim))
encoded_z = (random_z * std) + mu
# Generate fake image
fake_img_cVAE = G(cVAE_data['img'], encoded_z)
# Get scores and loss
real_d_cVAE_1, real_d_cVAE_2 = D_cVAE(cVAE_data['ground_truth'])
fake_d_cVAE_1, fake_d_cVAE_2 = D_cVAE(fake_img_cVAE)
# mse_loss for LSGAN
D_loss_cVAE_1 = mse_loss(real_d_cVAE_1, 1) + mse_loss(fake_d_cVAE_1, 0)
D_loss_cVAE_2 = mse_loss(real_d_cVAE_2, 1) + mse_loss(fake_d_cVAE_2, 0)
############# Step 2. D loss in cLR-GAN #############
# Random latent vector
random_z = util.var(torch.randn(1, params.z_dim))
# Generate fake image
fake_img_cLR = G(cLR_data['img'], random_z)
# Get scores and loss
real_d_cLR_1, real_d_cLR_2 = D_cLR(cLR_data['ground_truth'])
fake_d_cLR_1, fake_d_cLR_2 = D_cLR(fake_img_cLR)
D_loss_cLR_1 = mse_loss(real_d_cLR_1, 1) + mse_loss(fake_d_cLR_1, 0)
D_loss_cLR_2 = mse_loss(real_d_cLR_2, 1) + mse_loss(fake_d_cLR_2, 0)
D_loss = D_loss_cVAE_1 + D_loss_cLR_1 + D_loss_cVAE_2 + D_loss_cLR_2
# Update
optim_D_cVAE.zero_grad()
optim_D_cLR.zero_grad()
optim_G.zero_grad()
optim_E.zero_grad()
D_loss.backward()
optim_D_cVAE.step()
optim_D_cLR.step()
''' ----------------------------- 2. Train G & E ----------------------------- '''
############# Step 1. GAN loss to fool discriminator (cVAE_GAN and cLR_GAN) #############
# Encoded latent vector
mu, log_variance = E(cVAE_data['ground_truth'])
std = torch.exp(log_variance / 2)
random_z = util.var(torch.randn(1, params.z_dim))
encoded_z = (random_z * std) + mu
# Generate fake image and get adversarial loss
fake_img_cVAE = G(cVAE_data['img'], encoded_z)
fake_d_cVAE_1, fake_d_cVAE_2 = D_cVAE(fake_img_cVAE)
GAN_loss_cVAE_1 = mse_loss(fake_d_cVAE_1, 1)
GAN_loss_cVAE_2 = mse_loss(fake_d_cVAE_2, 1)
# Random latent vector
random_z = util.var(torch.randn(1, params.z_dim))
# Generate fake image and get adversarial loss
fake_img_cLR = G(cLR_data['img'], random_z)
fake_d_cLR_1, fake_d_cLR_2 = D_cLR(fake_img_cLR)
GAN_loss_cLR_1 = mse_loss(fake_d_cLR_1, 1)
GAN_loss_cLR_2 = mse_loss(fake_d_cLR_2, 1)
G_GAN_loss = GAN_loss_cVAE_1 + GAN_loss_cVAE_2 + GAN_loss_cLR_1 + GAN_loss_cLR_2
############# Step 2. KL-divergence with N(0, 1) (cVAE-GAN) #############
KL_div = params.lambda_kl * torch.sum(0.5 * (mu ** 2 + torch.exp(log_variance) - log_variance - 1))
############# Step 3. Reconstruction of ground truth image (|G(A, z) - B|) (cVAE-GAN) #############
img_recon_loss = params.lambda_img * L1_loss(fake_img_cVAE, cVAE_data['ground_truth'])
EG_loss = G_GAN_loss + KL_div + img_recon_loss
optim_D_cVAE.zero_grad()
optim_D_cLR.zero_grad()
optim_G.zero_grad()
optim_E.zero_grad()
EG_loss.backward(retain_graph=True)
optim_E.step()
optim_G.step()
''' ----------------------------- 3. Train ONLY G ----------------------------- '''
############ Step 1. Reconstrution of random latent code (|E(G(A, z)) - z|) (cLR-GAN) ############
# This step should update ONLY G.
mu_, log_variance_ = E(fake_img_cLR)
z_recon_loss = L1_loss(mu_, random_z)
G_alone_loss = params.lambda_z * z_recon_loss
optim_D_cVAE.zero_grad()
optim_D_cLR.zero_grad()
optim_G.zero_grad()
optim_E.zero_grad()
G_alone_loss.backward()
optim_G.step()
log_file = open('log.txt', 'w')
log_file.write(str(epoch))
# Print error and save intermediate result image and weight
if iters % params.save_every == 0:
print('[Epoch : %d / Iters : %d/%d] => D_loss : %f / G_GAN_loss : %f / KL_div : %f / img_recon_loss : %f / z_recon_loss : %f'\
%(epoch, iters, len(train_data_loader),D_loss.data[0], G_GAN_loss.data[0], KL_div.data[0], img_recon_loss.data[0], G_alone_loss.data[0]))
# Save intermediate result image
if os.path.exists(params.result_dir) is False:
os.makedirs(params.result_dir)
result_img = util.make_img(test_data_loader, G, fixed_z, img_num=params.test_img_num, img_size=params.img_size)
img_name = '{epoch}_{iters}.png'.format(epoch=epoch, iters=iters)
img_path = os.path.join(params.result_dir, img_name)
torchvision.utils.save_image(result_img, img_path, nrow=params.test_img_num+1)
# Save intermediate weight
if os.path.exists(params.weight_dir) is False:
os.makedirs(params.weight_dir)
if epoch is None:
d_cVAE_name = 'D_cVAE.pkl'
d_cLR_name = 'D_cLR.pkl'
g_name = 'G.pkl'
e_name = 'E.pkl'
else:
d_cVAE_name = '{epochs}-{name}'.format(epochs=str(epoch), name='D_cVAE.pkl')
d_cLR_name = '{epochs}-{name}'.format(epochs=str(epoch), name='D_cLR.pkl')
g_name = '{epochs}-{name}'.format(epochs=str(epoch), name='G.pkl')
e_name = '{epochs}-{name}'.format(epochs=str(epoch), name='E.pkl')
torch.save(D_cVAE.state_dict(), os.path.join(params.weight_dir, d_cVAE_name))
torch.save(D_cVAE.state_dict(), os.path.join(params.weight_dir, d_cLR_name))
torch.save(G.state_dict(), os.path.join(params.weight_dir, g_name))
torch.save(E.state_dict(), os.path.join(params.weight_dir, e_name))
# Save weight at the end of every epoch
if epoch is None:
d_cVAE_name = 'D_cVAE.pkl'
d_cLR_name = 'D_cLR.pkl'
g_name = 'G.pkl'
e_name = 'E.pkl'
else:
d_cVAE_name = '{epochs}-{name}'.format(epochs=str(epoch), name='D_cVAE.pkl')
d_cLR_name = '{epochs}-{name}'.format(epochs=str(epoch), name='D_cLR.pkl')
g_name = '{epochs}-{name}'.format(epochs=str(epoch), name='G.pkl')
e_name = '{epochs}-{name}'.format(epochs=str(epoch), name='E.pkl')
torch.save(D_cVAE.state_dict(), os.path.join(params.weight_dir, d_cVAE_name))
torch.save(D_cVAE.state_dict(), os.path.join(params.weight_dir, d_cLR_name))
torch.save(G.state_dict(), os.path.join(params.weight_dir, g_name))
torch.save(E.state_dict(), os.path.join(params.weight_dir, e_name))
```
| github_jupyter |
Copyright 2021 Google LLC.
SPDX-License-Identifier: Apache-2.0
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Assessing the veracity of semantic markup for dataset pages
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/google-research/google-research/blob/master/dataset_or_not/dataset_or_not.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/google-research/google-research/tree/master/dataset_or_not/dataset_or_not.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## About this Colab
This is a companion Colab for the paper
[Dataset or Not? A study on the veracity of semantic markup for dataset pages]()\
*Tarfah Alrashed, Dimitris Paparas, Omar Benjelloun, Ying Sheng, and Natasha Noy*
It contains python code for training the two main models from the paper, using the [Veracity of schema.org for datasets (labeled data)](https://www.kaggle.com/googleai/veracity-of-schemaorg-for-datasets-labeled-data) dataset.
## Prerequisites
Before continuing, download and unzip the [Veracity of schema.org for datasets (labeled data)](https://www.kaggle.com/googleai/veracity-of-schemaorg-for-datasets-labeled-data) dataset to your computer.
## Note regarding *prominent terms*
The released dataset and the code in this notebook do not contain the *prominent terms* feature mentioned in the paper. This is because that feature is extracted using proprietary code that cannot be released. The interested reader can replicate this feature extraction using the model proposed in [this paper](https://arxiv.org/abs/1805.01334).
#Install required packages
```
!pip install adanet
!pip install --user --upgrade tensorflow-probability
```
#Import Modules
```
from google.colab import files
import math
import tensorflow.compat.v2 as tf
import adanet
import pandas as pd
import io
from tensorflow.keras.preprocessing import sequence
from tensorflow.keras.preprocessing import text
```
# Upload Dataset
Run the following cell and, when prompted, upload files *testing_set.csv*, *training_set.csv*, and *validation_set.csv* (that you downloaded as part of the prerequisites).
```
uploaded = files.upload()
```
# Load dataset in pandas.DataFrame
```
training_set = pd.read_csv('training_set.csv', keep_default_na=False)
eval_set = pd.read_csv('validation_set.csv', keep_default_na=False)
test_set = pd.read_csv('testing_set.csv', keep_default_na=False)
```
# Select Model
In the next cell you can select which model to train. Remember to run the cell after making a selection. The features each model uses are:
Column Name|Type|Contents|Lightweight Model|Full Model
-----------|----|-|:-:|:--------:
source_url| string |url of a webpage that contains schema.org/Dataset markup| |+
name| string |The name of the dataset| +|+
description| string |Description of the dataset|+|+
has_distribution| bool|True if the dataset contains distribution metadata, false otherwise| |+
has_encoding_or_file_format| bool |True if the dataset contains encoding or file format metadata, false otherwise| |+
provider_or_publisher| string |The name of the provider or publisher of the dataset| |+
author_or_creator| string |The author(s) or creator(s) of the dataset| |+
doi| string|The Digital Object Identifier of the dataset| |+
has_catalog| bool |True if the dataset is included in a data catalog, false otherwise| |+|
has_dateCreated| bool |True if a creation date is provided, false otherwise| |+
has_dateModified| bool |True if a modification date is provided, false otherwise| |+
has_datePublished| bool |True if a publication date is provided, false otherwise| |+
```
SELECTED_MODEL = 'lightweight_model' #@param {type:'string'} ["lightweight_model", "full_model"]
```
# Preprocessing Parameters
Dictionary with the sizes of the feature vocabularies to generate during preprocessing for each of the models
```
P_PARAMS_BY_MODEL = {
'lightweight_model': {
'vocab_size_by_feature': {
'description': 110211,
'name': 18720
},
'MAX_TOKENS': 400
},
'full_model': {
'vocab_size_by_feature': {
'description': 104383,
'name': 17495,
'author_or_creator': 1602,
'doi': 193,
'provider_or_publisher': 773,
'source_url': 17749
},
'MAX_TOKENS': 400
}
}
MODEL_P_PARAMS = P_PARAMS_BY_MODEL[SELECTED_MODEL]
```
# Data Preprocessing
Analyze training dataset and generate tokenizers with custom vocabularies for each text feature
```
tokenizers = {}
for feature_name, vocab_size in MODEL_P_PARAMS['vocab_size_by_feature'].items():
tokenizers[feature_name] = text.Tokenizer(num_words=vocab_size)
tokenizers[feature_name].fit_on_texts(training_set[feature_name])
```
# Hyperparametes
Dictionary with the training hyperparameters for each of the models
```
H_PARAMS_BY_MODEL = {
'lightweight_model': {
'features': ['description', 'name'],
'LEARNING_RATE': 0.00677,
'TRAIN_STEPS': 500,
'SHUFFLE_BUFFER_SIZE': 2048,
'BATCH_SIZE': 128,
'CLIP_NORM': 0.00037,
'HIDDEN_UNITS': [186],
'DROPOUT': 0.28673,
'ACTIVATION_FN': tf.nn.selu,
'MAX_ITERATION_STEPS': 333333,
'DO_BATCH_NORM': True,
'MAX_TRAIN_STEPS': 1000
},
'full_model': {
'features': [
'author_or_creator', 'description', 'doi', 'has_date_created',
'has_date_modified', 'has_date_published', 'has_distribution',
'has_encoding_or_file_format', 'name', 'provider_or_publisher',
'source_url'
],
'LEARNING_RATE': 0.00076,
'TRAIN_STEPS': 500,
'SHUFFLE_BUFFER_SIZE': 2048,
'BATCH_SIZE': 128,
'CLIP_NORM': 0.25035,
'HIDDEN_UNITS': [329, 351, 292],
'DROPOUT': 0.08277,
'ACTIVATION_FN': tf.nn.selu,
'MAX_ITERATION_STEPS': 333333,
'DO_BATCH_NORM': False,
'MAX_TRAIN_STEPS': 1000
}
}
MODEL_H_PARAMS = H_PARAMS_BY_MODEL[SELECTED_MODEL]
```
# Utility functions
Methods used to preprocess and create the input for training the model
```
def tokenize_and_pad(features):
"""Iterates over the features of a labeled sample, tokenizing and padding them.
Args:
features: A dictionary of feature values keyed by feature names. It includes
label as a feature
Returns:
A tuple with the processed features
"""
tokenized_features = list()
for feature in MODEL_H_PARAMS['features']:
# Tokenize text features according to the corresponding vocabulary
if feature in MODEL_P_PARAMS['vocab_size_by_feature']:
# Handle missing features
if not features[feature]:
tokenized = [[MODEL_P_PARAMS['vocab_size_by_feature'][feature]]]
else:
tokenized = tokenizers[feature].texts_to_sequences([features[feature]])
tokenized_features.append([
sequence.pad_sequences(
tokenized,
maxlen=MODEL_P_PARAMS['MAX_TOKENS'],
padding='post',
truncating='post')
])
# Tokenize boolean features into binary values
else:
if features[feature]:
tokenized_features.append([1])
else:
tokenized_features.append([0])
tokenized_features.append(features['label'])
return tuple(tokenized_features)
def generator(dataset):
"""Returns a generator mapping dataset entries to tokenized features-label pairs."""
def _gen():
for entry in dataset.iterrows():
yield tokenize_and_pad(entry[1])
return _gen
def preprocess(*args):
"""Tensorizes its arguments.
Args:
*args: Variable length arguments feature1, ..., featureK, label. Should be
in the same order as in MODEL_H_PARAMS['features']
Returns:
A pair of
1. A dictionary with the features keyed by their names
2. A label
"""
m = {}
for feature, name in zip(args[:-1], MODEL_H_PARAMS['features']):
m[name] = feature
return m, [args[-1]]
def generate_output_types():
"""Returns a vector of output types corresponding to the tuple produced by the generator."""
types = []
# Feature types
types = [tf.int32] * len(MODEL_H_PARAMS['features'])
# Label type
types.append(tf.bool)
return tuple(types)
def input_fn(partition, training, batch_size):
"""Generates an input_fn for the Estimator.
Args:
partition: One of 'train', 'test', and 'eval' for training, testing, and
validation sets respectively
training: If true, then shuffle dataset to add randomness between epochs
batch_size: Number of elements to combine in a single batch
Returns:
The input function
"""
def _input_fn():
if partition == 'train':
dataset = tf.data.Dataset.from_generator(
generator(training_set), generate_output_types())
elif partition == 'test':
dataset = tf.data.Dataset.from_generator(
generator(test_set), generate_output_types())
elif partition == 'eval':
dataset = tf.data.Dataset.from_generator(
generator(eval_set), generate_output_types())
else:
print('Unknown partition')
return
if training:
dataset = dataset.shuffle(MODEL_H_PARAMS['SHUFFLE_BUFFER_SIZE'] *
batch_size).repeat()
dataset = dataset.map(preprocess).batch(batch_size)
iterator = tf.compat.v1.data.make_one_shot_iterator(dataset)
features, labels = iterator.get_next()
return features, labels
return _input_fn
def generate_feature_columns(embed):
"""Creates the feature columns that we will train the model on.
Args:
embed: If true, we embed the columns.
Returns:
A list with the feature columns.
"""
feature_columns = []
for feature in MODEL_H_PARAMS['features']:
if feature in MODEL_P_PARAMS['vocab_size_by_feature']:
# vocab_size + 1 to handle missing features
num_buckets = MODEL_P_PARAMS['vocab_size_by_feature'][feature] + 1
else:
# All none-text features are booleans, so 2 buckets are enough
num_buckets = 2
column = tf.feature_column.categorical_column_with_identity(
key=feature, num_buckets=num_buckets)
if embed:
column = tf.feature_column.embedding_column(
column, dimension=math.ceil(math.log2(num_buckets)))
feature_columns.append(column)
return feature_columns
```
# Build Model
Set up an ensemble estimator combining a Linear estimator and a DNN
```
head = tf.estimator.BinaryClassHead()
adam = lambda: tf.keras.optimizers.Adam(
learning_rate=MODEL_H_PARAMS['LEARNING_RATE'],
clipnorm=MODEL_H_PARAMS['CLIP_NORM'])
estimator = adanet.AutoEnsembleEstimator(
head=head,
candidate_pool={
'linear':
tf.estimator.LinearEstimator(
head=head,
feature_columns=generate_feature_columns(False),
optimizer=adam),
'dnn':
tf.estimator.DNNEstimator(
head=head,
hidden_units=MODEL_H_PARAMS['HIDDEN_UNITS'],
feature_columns=generate_feature_columns(True),
optimizer=adam,
activation_fn=MODEL_H_PARAMS['ACTIVATION_FN'],
dropout=MODEL_H_PARAMS['DROPOUT'],
batch_norm=MODEL_H_PARAMS['DO_BATCH_NORM'])
},
max_iteration_steps=MODEL_H_PARAMS['MAX_ITERATION_STEPS'])
```
# Train Model
For demonstration purposes, we set *max_steps* to a small value so that the
training finishes fast. This is enough to achieve good results. Alternatively, you can remove the *max_steps* argument and let the estimator train to convergence.
```
result = tf.estimator.train_and_evaluate(
estimator,
train_spec=tf.estimator.TrainSpec(
input_fn=input_fn(
'train', training=True, batch_size=MODEL_H_PARAMS['BATCH_SIZE']),
max_steps=MODEL_H_PARAMS['MAX_TRAIN_STEPS']),
eval_spec=tf.estimator.EvalSpec(
input_fn=input_fn(
'eval', training=False, batch_size=MODEL_H_PARAMS['BATCH_SIZE']),
steps=None,
start_delay_secs=1,
throttle_secs=1,
))[0]
```
# Model perfomance on validation set
```
print('AUC:', result['auc'], 'AUC_PR:', result['auc_precision_recall'],
'Recall:', result['recall'], 'Precision:', result['precision'])
```
# Model perfomance on testing set
```
ret = estimator.evaluate(
input_fn('test', training=False, batch_size=MODEL_H_PARAMS['BATCH_SIZE']))
print('AUC:', ret['auc'], 'AUC_PR:', ret['auc_precision_recall'], 'Recall:',
ret['recall'], 'Precision:', ret['precision'])
```
| github_jupyter |
# 常规赛简介
飞桨(PaddlePaddle)以百度多年的深度学习技术研究和业务应用为基础,集深度学习核心框架、基础模型库、端到端开发套件、工具组件和服务平台于一体,是中国首个开源开放、技术领先、功能完备的产业级深度学习平台。更多飞桨资讯,点击此处查看。
飞桨常规赛由百度飞桨于2019年发起,面向全球AI开发者,赛题范围广,涵盖领域多。常规赛旨在通过长期发布的经典比赛项目,为开发者提供学习锻炼机会,助力大家在飞将大赛中获得骄人成绩。
参赛选手需使用飞桨框架,基于特定赛题下的真实行业数据完成并提交任务。常规赛采取月度评比方式,为打破历史最高记录选手和当月有资格参与月度评奖的前10名选手提供飞桨特别礼包奖励。更多惊喜,更多收获,尽在飞桨常规赛。
比赛链接:https://aistudio.baidu.com/aistudio/competition/detail/20/0/introduction
# 导入对应模块
```
import numpy as np
import pandas as pd
import cv2
import matplotlib.pyplot as plt
import os
import random
```
# 数据准备
数据集 简介<br>
train.csv 训练集标签<br>
train_images.zip 训练集全部图片<br>
test_images.zip 测试集全部图片<br>
数据集链接:<br>
常规赛:中文场景文字识别-训练集:[https://aistudio.baidu.com/aistudio/datasetdetail/62842](https://aistudio.baidu.com/aistudio/datasetdetail/62842)<br>
常规赛:中文场景文字识别-测试集:[https://aistudio.baidu.com/aistudio/datasetdetail/62843](http://aistudio.baidu.com/aistudio/datasetdetail/62843)
```
#解压数据
!unzip ./data/data62842/train_images.zip -d ./work/
!unzip ./data/data62843/test_images.zip -d ./work/
!cp ./data/data62842/train_label.csv -d ./work/
#把csv文件转换成txt
%cd /home/aistudio/
train_label_pd=pd.read_csv('./work/train_label.csv',encoding='GBK')
len(train_label_pd)
with open('./work/all.txt','w') as f:
for i in range(len(train_label_pd)):
name= train_label_pd['name'][i]
label = train_label_pd['value'][i]
f.write(name)
f.write('\t')
f.write(label)
f.write('\n')
```
# 划分训练与验证数据
把训练集5%划分为验证集
```
with open('./work/all.txt','r') as f:
lines=f.readlines()
number = len(lines)
print(number)
print(lines[0])
random.shuffle(lines)
print(lines[0])
with open('./work/train_list.txt','w') as f1:
for i in range(int(len(lines)*0.95)):
f1.write(lines[i])
print(int(len(lines)*0.95))
print(len(lines))
with open('./work/evl_list.txt','w') as f2:
for i in range(int(len(lines)*0.95),len(lines)):
f2.write(lines[i])
```
# clone paddleocr
```
# 下载代码
# !git clone https://gitee.com/paddlepaddle/PaddleOCR.git
# 安装运行所需要的whl包
os.chdir('/home/aistudio')
!pip install -U pip
!pip install -r ./PaddleOCR/requirements.txt
```
# 下载推理模型
```
!wget https://paddleocr.bj.bcebos.com/dygraph_v2.0/ch/ch_ppocr_server_v2.0_rec_pre.tar
!tar -xf ch_ppocr_server_v2.0_rec_train.tar
```
修改/PaddleOCR/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml 配置文件开启训练
```
#开启训练
os.chdir('/home/aistudio')
!python ./PaddleOCR/tools/train.py -c ./PaddleOCR/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml
#evl
!python ./PaddleOCR/tools/eval.py -c ./PaddleOCR/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml -o Global.checkpoints=output/rec_chinese_common_v2.0/best_accuracy
```
# 导出模型
```
#导出模型
!python ./PaddleOCR/tools/export_model.py -c ./PaddleOCR/configs/rec/ch_ppocr_v2.0/rec_chinese_common_train_v2.0.yml \
-o Global.pretrained_model="./output/rec_chinese_common_v2.0/best_accuracy" \
Global.save_inference_dir="./my_model"
```
# 预测结果
修改./tools/infer/predict_rec.py文件后台保存result.txt文件
```
#修改./tools/infer/predict_rec.py文件后台保存result.txt文件
import os
os.chdir('/home/aistudio/PaddleOCR/')
!python ./tools/infer/predict_rec.py --image_dir="../work/test_images" --rec_model_dir="/home/aistudio/my_model" --use_gpu=True
```
# 思路介绍
使用paddleocr预训练模型,能有效提升识别效果。
模型下载地址:https://gitee.com/paddlepaddle/PaddleOCR/blob/release/2.4/doc/doc_ch/models_list.md
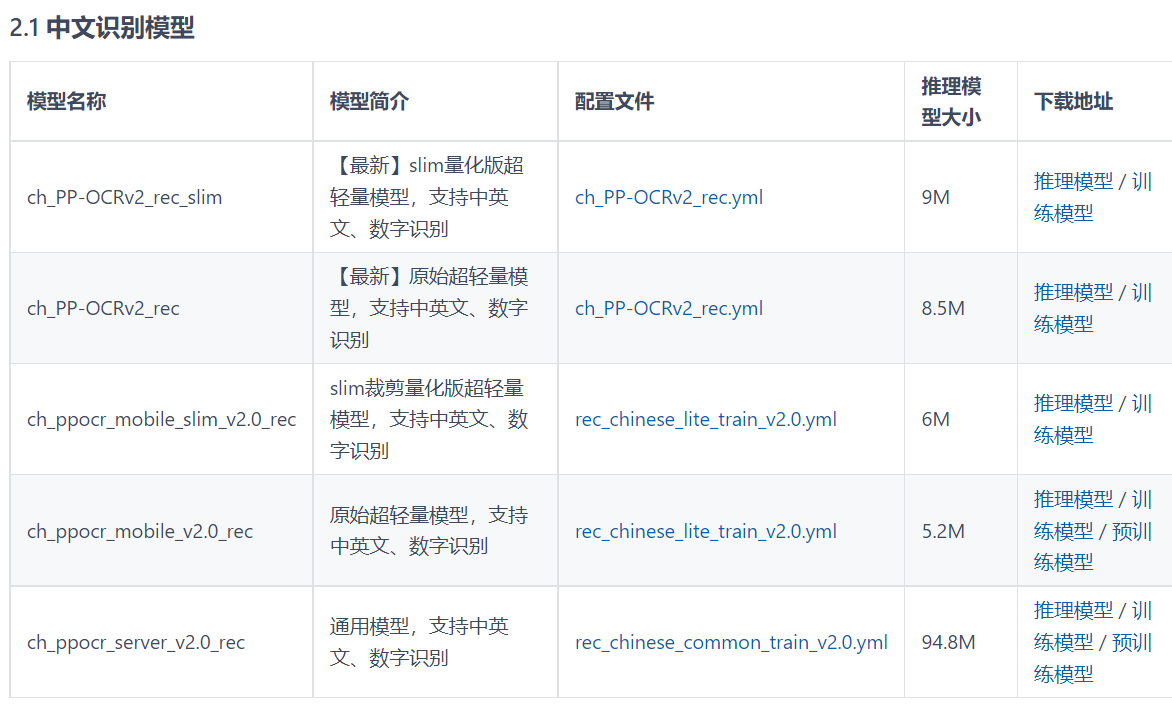
本次使用的是:ch_ppocr_server_v2.0_rec预训练模型<br>
模型使用的是CRNN网络<br>
Backbone:使用的是resnet34<br>
Neck:使用的是rnn<br>
Head:使用的是ctchead<br>
参数设置:
epoch设置的是300轮。<br>
训练的时候发现前面几十轮就差不多是现在这个成绩,跑完了300轮分数还变低了。<br>
learning_rate: 0.001 应该还要设置的再小点。
# 总结
成绩为:74.89251
还有很大的提升空间。
比如,把训练集里面,全角都改为半角,大写转小写,删除空字符等等。
然后再优化下参数等等。
| github_jupyter |
# IQ2 Dataset to Convokit format conversion script
Marianne Aubin Le Quere and Lucas Van Bramer
```
# if needed, change directory to find convokit location
# import required modules and set up environment
import os
# replace file path below with your own local convokit
os.chdir('/Users/marianneaubin/Documents/Classes/CS6742/Cornell-Conversational-Analysis-Toolkit')
import convokit
from convokit import Corpus, User, Utterance
from tqdm import tqdm
import json
# generates all of the users who are listed in the metadata of a specific debate's "speakers" field
# args: debate_id is a key used by the iq2 dataset, e.g. "PerformanceEnhancingDrugs-011508"
# returns: a dictionary in which keys are speakers' full names and values are dictionaries containing metadata
def generate_users(debate_id):
debate = iq2[debate_id]
users = {}
for stance in ["for", "against"]:
for speaker in debate["speakers"][stance]:
meta = {}
meta["stance"] = stance
meta["bio"] = speaker["bio"]
meta["bio_short"] = speaker["bio_short"]
users[speaker["name"]] = meta
mod = debate["speakers"]["moderator"]
modmeta = {}
modmeta["bio"] = mod["bio"]
modmeta["bio_short"] = mod["bio_short"]
modmeta["stance"] = None
users[mod["name"]] = modmeta
users["audience"] = {"bio": None, "bio_short": None, "stance": None}
return users
# generates all of the users in the iq2 dataset
# args: dataset is the python object containing the iq2 dataset parsed from json
# returns: a dictionary in which keys are speakers' full names and values are dictionaries containing metadata
def generate_all_users_convokit(dataset):
all_users = {}
for debate_id in dataset.keys():
res = generate_users(debate_id)
for fullname, usermeta in res.items():
all_users[fullname] = usermeta
print(str(len(all_users.keys())) + " users generated.")
convokit_all_users = {k: User(name = k, meta = v) for k,v in all_users.items()}
return convokit_all_users
# generates all of the convokit utterances from the iq2 dataset
# args: dataset is the python object containing the iq2 dataset parsed from json
# precondition: corpus_users must be populated in the jupyter environment because it is called from and modified here
# returns: convokit corpus representation of iq2 dataset
def generate_utterance_corpus_from_dataset(dataset, v):
utt_id = 0
utterance_corpus = {}
for conversation_id in dataset.keys():
conversation = dataset[conversation_id]
# set root of the conversation to the first utterance id in the conversation
convo_root = utt_id
for turn in conversation["transcript"]:
utterance = {}
utterance["id"] = str(utt_id)
utterance["root"] = str(convo_root)
utterance["timestamp"] = None
meta = {
"nontext": turn["nontext"],
"segment": turn["segment"],
"speakertype": turn["speakertype"],
"debateid": conversation_id
}
utterance["meta"] = meta
# sets replied-to utterance to always be the last utterance
utterance["reply_to"] = utt_id - 1
# text is originally stored as a list of strings; this concatenates them into one string
fulltext = "".join(turn["paragraphs"])
utterance["text"] = fulltext
# "unknown" speakers are generally the audience
utterance["user"] = turn["speaker"] if turn["speakertype"] != "unknown" else "audience"
# in the case that a speaker in the text is not a speaker contained in the debate
# metadata, adds a speaker with the same schema but no metadata to the corpus users
if turn["speaker"] not in corpus_users:
meta = {}
meta["stance"] = None
meta["bio"] = None
meta["bio_short"] = None
corpus_users[turn["speaker"]] = User(name=turn["speaker"], meta=meta)
# adds convokit utterance to corpus object
utterance_corpus[utterance["id"]] = \
Utterance(utterance["id"],
corpus_users[utterance["user"]],
utterance["root"],
utterance["reply_to"],
utterance["timestamp"],
utterance["text"],
meta=utterance["meta"]
)
# increments utterance id
utt_id += 1
# converts utterance dictionary into convokit format
corpus = Corpus(utterances=[utt for _, utt in utterance_corpus.items()], version=v)
return corpus
# replace open location with where the dataset is
!pwd
file = open('../IQ2/iq2_data_release/iq2_data_release.json')
iq2 = json.load(file)
print(str(len(iq2.keys())) + " debates loaded.")
corpus_users = generate_all_users_convokit(iq2)
iq2_corpus = generate_utterance_corpus_from_dataset(iq2, 1)
iq2_corpus.meta["name"] = "IQ2 Debate Corpus"
# this generates conversation metadata based on each debate and updates the corpus conversation instances
for conv_id in iq2_corpus.get_conversation_ids():
conv = iq2_corpus.get_conversation(conv_id)
first_utt = iq2_corpus.get_utterance(conv.get_utterance_ids()[0])
debate = iq2[first_utt.meta["debateid"]]
debate_meta = {}
debate_meta["summary"] = debate["summary"]
debate_meta["title"] = debate["title"]
debate_meta["date"] = debate["date"]
debate_meta["url"] = debate["url"]
debate_meta["results"] = debate["results"]
debate_meta["speakers"] = debate["speakers"]
conv.meta = debate_meta
# this function determines, given a conversation, which is the winner
def determine_winner(conversation):
results = conversation.meta["results"]
fordelta = results["post"]["for"] - results["pre"]["for"]
againstdelta = results["post"]["against"] - results["pre"]["against"]
if(fordelta > againstdelta):
return "for"
elif(againstdelta > fordelta):
return "against"
else:
return "tie"
for conv_id in iq2_corpus.get_conversation_ids():
conv = iq2_corpus.get_conversation(conv_id)
conv.meta["winner"] = determine_winner(conv)
# prints summary stats and dumps the corpus to file
iq2_corpus.print_summary_stats()
iq2_corpus.dump("iq2_corpus")
```
| github_jupyter |
# Writing Your Own Graph Algorithms
The analytical engine in GraphScope derives from [GRAPE](https://dl.acm.org/doi/10.1145/3282488), a graph processing system proposed on SIGMOD-2017. GRAPE differs from prior systems in its ability to parallelize sequential graph algorithms as a whole. In GRAPE, sequential algorithms can be easily **plugged into** with only minor changes and get parallelized to handle large graphs efficiently.
In this tutorial, we will show how to define and run your own algorithm in PIE and Pregel models.
Sounds like fun? Excellent, here we go!
## Writing algorithm in PIE model
GraphScope enables users to write algorithms in the [PIE](https://dl.acm.org/doi/10.1145/3282488) programming model in a pure Python mode, first of all, you should import **graphscope** package and the **pie** decorator.
```
import graphscope
from graphscope.framework.app import AppAssets
from graphscope.analytical.udf.decorators import pie
```
We use the single source shortest path ([SSSP](https://en.wikipedia.org/wiki/Shortest_path_problem)) algorithm as an example. To implement the PIE model, you just need to **fulfill this class**
```
@pie(vd_type="double", md_type="double")
class SSSP_PIE(AppAssets):
@staticmethod
def Init(frag, context):
pass
@staticmethod
def PEval(frag, context):
pass
@staticmethod
def IncEval(frag, context):
pass
```
The **pie** decorator contains two params named `vd_type` and `md_type` , which represent the vertex data type and message type respectively.
You may specify types for your own algorithms, optional values are `int`, `double`, and `string`.
In our **SSSP** case, we compute the shortest distance to the source for all nodes, so we use `double` value for `vd_type` and `md_type` both.
In `Init`, `PEval`, and `IncEval`, it has **frag** and **context** as parameters. You can use these two parameters to access the fragment data and intermediate results. Detail usage please refer to [Cython SDK API](https://graphscope.io/docs/reference/cython_sdk.html).
### Fulfill Init Function
```
@pie(vd_type="double", md_type="double")
class SSSP_PIE(AppAssets):
@staticmethod
def Init(frag, context):
v_label_num = frag.vertex_label_num()
for v_label_id in range(v_label_num):
nodes = frag.nodes(v_label_id)
context.init_value(
nodes, v_label_id, 1000000000.0, PIEAggregateType.kMinAggregate
)
context.register_sync_buffer(v_label_id, MessageStrategy.kSyncOnOuterVertex)
@staticmethod
def PEval(frag, context):
pass
@staticmethod
def IncEval(frag, context):
pass
```
The `Init` function are responsable for 1) setting the initial value for each node; 2) defining the strategy of message passing; and 3) specifing aggregator for handing received message on each rounds.
Note that the algorithm you defined will run on a property graph. So we should get the vertex label first by `v_label_num = frag.vertex_label_num()`, then we can traverse all nodes with the same label
and set the initial value by `nodes = frag.nodes(v_label_id)` and `context.init_value(nodes, v_label_id, 1000000000.0, PIEAggregateType.kMinAggregate)`.
Since we are computing the shorest path between the source node and others nodes. So we use `PIEAggregateType.kMinAggregate` as the aggregator for mesaage aggregation, which means it will
perform `min` operation upon all received messages. Other avaliable aggregators are `kMaxAggregate`, `kSumAggregate`, `kProductAggregate`, and `kOverwriteAggregate`.
At the end of `Init` function, we register the sync buffer for each node with `MessageStrategy.kSyncOnOuterVertex`, which tells the engine how to pass the message.
### Fulfill PEval Function
```
@pie(vd_type="double", md_type="double")
class SSSP_PIE(AppAssets):
@staticmethod
def Init(frag, context):
v_label_num = frag.vertex_label_num()
for v_label_id in range(v_label_num):
nodes = frag.nodes(v_label_id)
context.init_value(
nodes, v_label_id, 1000000000.0, PIEAggregateType.kMinAggregate
)
context.register_sync_buffer(v_label_id, MessageStrategy.kSyncOnOuterVertex)
@staticmethod
def PEval(frag, context):
src = int(context.get_config(b"src"))
graphscope.declare(graphscope.Vertex, source)
native_source = False
v_label_num = frag.vertex_label_num()
for v_label_id in range(v_label_num):
if frag.get_inner_node(v_label_id, src, source):
native_source = True
break
if native_source:
context.set_node_value(source, 0)
else:
return
e_label_num = frag.edge_label_num()
for e_label_id in range(e_label_num):
edges = frag.get_outgoing_edges(source, e_label_id)
for e in edges:
dst = e.neighbor()
distv = e.get_int(2)
if context.get_node_value(dst) > distv:
context.set_node_value(dst, distv)
@staticmethod
def IncEval(frag, context):
pass
```
In `PEval` of **SSSP**, it gets the queried source node by `context.get_config(b"src")`.
`PEval` checks each fragment whether it contains source node by `frag.get_inner_node(v_label_id, src, source)`. Note that the `get_inner_node` method needs a `source` parameter in type `Vertex`, which you can declare by `graphscope.declare(graphscope.Vertex, source)`
If a fragment contains the source node, it will traverse the outgoing edges of the source with `frag.get_outgoing_edges(source, e_label_id)`. For each vertex, it computes the distance from the source, and updates the value if the it less than the initial value.
### Fulfill IncEval Function
```
@pie(vd_type="double", md_type="double")
class SSSP_PIE(AppAssets):
@staticmethod
def Init(frag, context):
v_label_num = frag.vertex_label_num()
for v_label_id in range(v_label_num):
nodes = frag.nodes(v_label_id)
context.init_value(
nodes, v_label_id, 1000000000.0, PIEAggregateType.kMinAggregate
)
context.register_sync_buffer(v_label_id, MessageStrategy.kSyncOnOuterVertex)
@staticmethod
def PEval(frag, context):
src = int(context.get_config(b"src"))
graphscope.declare(graphscope.Vertex, source)
native_source = False
v_label_num = frag.vertex_label_num()
for v_label_id in range(v_label_num):
if frag.get_inner_node(v_label_id, src, source):
native_source = True
break
if native_source:
context.set_node_value(source, 0)
else:
return
e_label_num = frag.edge_label_num()
for e_label_id in range(e_label_num):
edges = frag.get_outgoing_edges(source, e_label_id)
for e in edges:
dst = e.neighbor()
distv = e.get_int(2)
if context.get_node_value(dst) > distv:
context.set_node_value(dst, distv)
@staticmethod
def IncEval(frag, context):
v_label_num = frag.vertex_label_num()
e_label_num = frag.edge_label_num()
for v_label_id in range(v_label_num):
iv = frag.inner_nodes(v_label_id)
for v in iv:
v_dist = context.get_node_value(v)
for e_label_id in range(e_label_num):
es = frag.get_outgoing_edges(v, e_label_id)
for e in es:
u = e.neighbor()
u_dist = v_dist + e.get_int(2)
if context.get_node_value(u) > u_dist:
context.set_node_value(u, u_dist)
```
The only difference between `IncEval` and `PEval` of **SSSP** algorithm is that `IncEval` are invoked
on each fragment, rather than only the fragment with source node. A fragment will repeat the `IncEval` until there is no messages received. When all the fragments are finished computation, the algorithm is terminated.
### Run Your Algorithm on A Graph.
First, let's establish a session and load a graph for testing.
```
from graphscope.framework.loader import Loader
# the location of the property graph for testing
property_dir = '/home/jovyan/datasets/property'
graphscope.set_option(show_log=True)
k8s_volumes = {
"data": {
"type": "hostPath",
"field": {
"path": "/testingdata",
"type": "Directory"
},
"mounts": {
"mountPath": "/home/jovyan/datasets",
"readOnly": True
}
}
}
sess = graphscope.session(k8s_volumes=k8s_volumes)
g = sess.load_from(
edges={
"knows": (
Loader("{}/p2p-31_property_e_0".format(property_dir), header_row=True),
["src_label_id", "dst_label_id", "dist"],
("src_id", "person"),
("dst_id", "person"),
),
},
vertices={
"person": Loader(
"{}/p2p-31_property_v_0".format(property_dir), header_row=True
),
},
generate_eid=False,
)
```
Then initialize your algorithm and query the shorest path from vertex `6` over the graph.
```
sssp = SSSP_PIE()
ctx = sssp(g, src=6)
```
Runing this cell, your algorithm should evaluate successfully. The results are stored in vineyard in the distributed machies. Let's fetch and check the results.
```
r1 = (
ctx.to_dataframe({"node": "v:person.id", "r": "r:person"})
.sort_values(by=["node"])
.to_numpy(dtype=float)
)
r1
```
### Dump and Reload Your Algorithm
You can dump and save your define algorithm for future use.
```
import os
# specify the path you want to dump
dump_path = os.path.expanduser("~/Workspace/sssp_pie.gar")
# dump
SSSP_PIE.to_gar(dump_path)
```
Now, you can find a package named `sssp_pie.gar` in your `~/Workspace`. Reload this algorithm with following code.
```
from graphscope.framework.app import load_app
# specify the path you want to dump
dump_path = os.path.expanduser("~/Workspace/sssp_pie.gar")
sssp2 = load_app("SSSP_PIE", dump_path)
```
### Write Algorithm in Pregel Model
In addition to the sub-graph based PIE model, GraphScope supports vertex-centric Pregel model. To define a Pregel algorithm, you should import **pregel** decorator and fulfil the functions defined on vertex.
```
import graphscope
from graphscope.framework.app import AppAssets
from graphscope.analytical.udf.decorators import pregel
@pregel(vd_type="double", md_type="double")
class SSSP_Pregel(AppAssets):
@staticmethod
def Init(v, context):
pass
@staticmethod
def Compute(messages, v, context):
pass
```
The **pregel** decorator has two parameters named `vd_type` and `md_type`, which represent the vertex data type and message type respectively.
You can specify the types for your algorithm, options are `int`, `double`, and `string`. For **SSSP**, we set both to `double`.
Since Pregel model are defined on vertex, the `Init` and `Compute` functions has a parameter `v` to access the vertex data. See more details in [Cython SDK API](https://graphscope.io/docs/reference/cython_sdk.html).
### Fulfill Init Function¶
```
@pregel(vd_type="double", md_type="double")
class SSSP_Pregel(AppAssets):
@staticmethod
def Init(v, context):
v.set_value(1000000000.0)
@staticmethod
def Compute(messages, v, context):
pass
```
The `Init` function sets the initial value for each node by `v.set_value(1000000000.0)`
### Fulfill Compute function¶
```
@pregel(vd_type="double", md_type="double")
class SSSP_Pregel(AppAssets):
@staticmethod
def Init(v, context):
v.set_value(1000000000.0)
@staticmethod
def Compute(messages, v, context):
src_id = context.get_config(b"src")
cur_dist = v.value()
new_dist = 1000000000.0
if v.id() == src_id:
new_dist = 0
for message in messages:
new_dist = min(message, new_dist)
if new_dist < cur_dist:
v.set_value(new_dist)
for e_label_id in range(context.edge_label_num()):
edges = v.outgoing_edges(e_label_id)
for e in edges:
v.send(e.vertex(), new_dist + e.get_int(2))
v.vote_to_halt()
```
The `Compute` function for **SSSP** computes the new distance for each node by the following steps:
1) Initialize the new value with value 1000000000
2) If the vertex is source node, set its distance to 0.
3) Compute the `min` value of messages received, and set the value if it less than the current value.
Repeat these, until no more new messages(shorter distance) are generated.
### Optional Combiner
Optionally, we can define a combiner to reduce the message communication overhead.
```
@pregel(vd_type="double", md_type="double")
class SSSP_Pregel(AppAssets):
@staticmethod
def Init(v, context):
v.set_value(1000000000.0)
@staticmethod
def Compute(messages, v, context):
src_id = context.get_config(b"src")
cur_dist = v.value()
new_dist = 1000000000.0
if v.id() == src_id:
new_dist = 0
for message in messages:
new_dist = min(message, new_dist)
if new_dist < cur_dist:
v.set_value(new_dist)
for e_label_id in range(context.edge_label_num()):
edges = v.outgoing_edges(e_label_id)
for e in edges:
v.send(e.vertex(), new_dist + e.get_int(2))
v.vote_to_halt()
@staticmethod
def Combine(messages):
ret = 1000000000.0
for m in messages:
ret = min(ret, m)
return ret
```
### Run Your Pregel Algorithm on Graph.
Next, let's run your Pregel algorithm on the graph, and check the results.
```
sssp_pregel = SSSP_Pregel()
ctx = sssp_pregel(g, src=6)
r2 = (
ctx.to_dataframe({"node": "v:person.id", "r": "r:person"})
.sort_values(by=["node"])
.to_numpy(dtype=float)
)
r2
```
It is important to release resources when they are no longer used.
```
sess.close()
```
### Aggregator in Pregel
Pregel aggregators are a mechanism for global communication, monitoring, and counting. Each vertex can provide a value to an aggregator in superstep `S`, the system combines these
values using a reducing operator, and the resulting value is made available to all vertices in superstep `S+1`. GraphScope provides a number of predefined aggregators for Pregel algorithms, such as `min`, `max`, or `sum` operations on data types.
Here is a example for use a builtin aggregator, more details can be found in [Cython SDK API](https://graphscope.io/docs/reference/cython_sdk.html)
```
@pregel(vd_type="double", md_type="double")
class Aggregators_Pregel_Test(AppAssets):
@staticmethod
def Init(v, context):
# int
context.register_aggregator(
b"int_sum_aggregator", PregelAggregatorType.kInt64SumAggregator
)
context.register_aggregator(
b"int_max_aggregator", PregelAggregatorType.kInt64MaxAggregator
)
context.register_aggregator(
b"int_min_aggregator", PregelAggregatorType.kInt64MinAggregator
)
# double
context.register_aggregator(
b"double_product_aggregator", PregelAggregatorType.kDoubleProductAggregator
)
context.register_aggregator(
b"double_overwrite_aggregator",
PregelAggregatorType.kDoubleOverwriteAggregator,
)
# bool
context.register_aggregator(
b"bool_and_aggregator", PregelAggregatorType.kBoolAndAggregator
)
context.register_aggregator(
b"bool_or_aggregator", PregelAggregatorType.kBoolOrAggregator
)
context.register_aggregator(
b"bool_overwrite_aggregator", PregelAggregatorType.kBoolOverwriteAggregator
)
# text
context.register_aggregator(
b"text_append_aggregator", PregelAggregatorType.kTextAppendAggregator
)
@staticmethod
def Compute(messages, v, context):
if context.superstep() == 0:
context.aggregate(b"int_sum_aggregator", 1)
context.aggregate(b"int_max_aggregator", int(v.id()))
context.aggregate(b"int_min_aggregator", int(v.id()))
context.aggregate(b"double_product_aggregator", 1.0)
context.aggregate(b"double_overwrite_aggregator", 1.0)
context.aggregate(b"bool_and_aggregator", True)
context.aggregate(b"bool_or_aggregator", False)
context.aggregate(b"bool_overwrite_aggregator", True)
context.aggregate(b"text_append_aggregator", v.id() + b",")
else:
if v.id() == b"1":
assert context.get_aggregated_value(b"int_sum_aggregator") == 62586
assert context.get_aggregated_value(b"int_max_aggregator") == 62586
assert context.get_aggregated_value(b"int_min_aggregator") == 1
assert context.get_aggregated_value(b"double_product_aggregator") == 1.0
assert (
context.get_aggregated_value(b"double_overwrite_aggregator") == 1.0
)
assert context.get_aggregated_value(b"bool_and_aggregator") == True
assert context.get_aggregated_value(b"bool_or_aggregator") == False
assert (
context.get_aggregated_value(b"bool_overwrite_aggregator") == True
)
context.get_aggregated_value(b"text_append_aggregator")
v.vote_to_halt()
```
| github_jupyter |
# Project 3: Smart Beta Portfolio and Portfolio Optimization
## Instructions
Each problem consists of a function to implement and instructions on how to implement the function. The parts of the function that need to be implemented are marked with a `# TODO` comment. After implementing the function, run the cell to test it against the unit tests we've provided. For each problem, we provide one or more unit tests from our `project_tests` package. These unit tests won't tell you if your answer is correct, but will warn you of any major errors. Your code will be checked for the correct solution when you submit it Udacity.
## Packages
When you implement the functions, you'll only need to use the [Pandas](https://pandas.pydata.org/) and [Numpy](http://www.numpy.org/) packages. Don't import any other packages, otherwise the grader willn't be able to run your code.
The other packages that we're importing is `helper` and `project_tests`. These are custom packages built to help you solve the problems. The `helper` module contains utility functions and graph functions. The `project_tests` contains the unit tests for all the problems.
### Install Packages
```
import sys
!{sys.executable} -m pip install -r requirements.txt
```
### Load Packages
```
import pandas as pd
import numpy as np
import helper
import project_tests
```
## Market Data
The data source we'll be using is the [Wiki End of Day data](https://www.quandl.com/databases/WIKIP) hosted at [Quandl](https://www.quandl.com). This contains data for many stocks, but we'll just be looking at the S&P 500 stocks. We'll also make things a little easier to solve by narrowing our range of time from 2007-06-30 to 2017-09-30.
### Set API Key
Set the `quandl.ApiConfig.api_key ` variable to your Quandl api key. You can find your Quandl api key [here](https://www.quandl.com/account/api).
```
import quandl
# TODO: Add your Quandl API Key
quandl.ApiConfig.api_key = ''
```
### Download Data
```
import os
snp500_file_path = 'data/tickers_SnP500.txt'
wiki_file_path = 'data/WIKI_PRICES.csv'
start_date, end_date = '2013-07-01', '2017-06-30'
use_columns = ['date', 'ticker', 'adj_close', 'adj_volume', 'ex-dividend']
if not os.path.exists(wiki_file_path):
with open(snp500_file_path) as f:
tickers = f.read().split()
print('Downloading data...')
helper.download_quandl_dataset('WIKI', 'PRICES', wiki_file_path, use_columns, tickers, start_date, end_date)
print('Data downloaded')
else:
print('Data already downloaded')
```
### Load Data
```
df = pd.read_csv(wiki_file_path)
```
### Create the Universe
We'll be selecting dollar volume stocks for our stock universe. This universe is similar to large market cap stocks, because they are the highly liquid.
```
percent_top_dollar = 0.2
high_volume_symbols = helper.large_dollar_volume_stocks(df, 'adj_close', 'adj_volume', percent_top_dollar)
df = df[df['ticker'].isin(high_volume_symbols)]
```
### 2-D Matrices
In the previous projects, we used a [multiindex](https://pandas.pydata.org/pandas-docs/stable/advanced.html) to store all the data in a single dataframe. As you work with larger datasets, it come infeasable to store all the data in memory. Starting with this project, we'll be storing all our data as 2-D matrices to match what you'll be expecting in the real world.
```
close = df.reset_index().pivot(index='ticker', columns='date', values='adj_close')
volume = df.reset_index().pivot(index='ticker', columns='date', values='adj_volume')
ex_dividend = df.reset_index().pivot(index='ticker', columns='date', values='ex-dividend')
```
### View Data
To see what one of these 2-d matrices looks like, let's take a look at the closing prices matrix.
```
helper.print_dataframe(close)
```
# Part 1: Smart Beta Portfolio
In Part 1 of this project, you'll build a smart beta portfolio using dividend yield. To see how well it performs, you'll compare this portfolio to an index.
## Index Weights
After building the smart beta portfolio, should compare it to a similar strategy or index.
Implement `generate_dollar_volume_weights` to generate the weights for this index. For each date, generate the weights based on dollar volume traded for that date. For example, assume the following is dollar volume traded data:
| | 10/02/2010 | 10/03/2010 |
|----------|------------|------------|
| **AAPL** | 2 | 2 |
| **BBC** | 5 | 6 |
| **GGL** | 1 | 2 |
| **ZGB** | 6 | 5 |
The weights should be the following:
| | 10/02/2010 | 10/03/2010 |
|----------|------------|------------|
| **AAPL** | 0.142 | 0.133 |
| **BBC** | 0.357 | 0.400 |
| **GGL** | 0.071 | 0.133 |
| **ZGB** | 0.428 | 0.333 |
```
def generate_dollar_volume_weights(close, volume):
"""
Generate dollar volume weights.
Parameters
----------
close : DataFrame
Close price for each ticker and date
volume : str
Volume for each ticker and date
Returns
-------
dollar_volume_weights : DataFrame
The dollar volume weights for each ticker and date
"""
assert close.index.equals(volume.index)
assert close.columns.equals(volume.columns)
#TODO: Implement function
dollar_volume = close * volume
return dollar_volume / dollar_volume.sum()
project_tests.test_generate_dollar_volume_weights(generate_dollar_volume_weights)
```
### View Data
Let's generate the index weights using `generate_dollar_volume_weights` and view them using a heatmap.
```
index_weights = generate_dollar_volume_weights(close, volume)
helper.plot_weights(index_weights, 'Index Weights')
```
## ETF Weights
Now that we have the index weights, it's time to build the weights for the smart beta ETF. Let's build an ETF portfolio that is based on dividends. This is a common factor used to build portfolios. Unlike most portfolios, we'll be using a single factor for simplicity.
Implement `calculate_dividend_weights` to returns the weights for each stock based on its total dividend yield over time. This is similar to generating the weight for the index, but it's dividend data instead.
```
def calculate_dividend_weights(ex_dividend):
"""
Calculate dividend weights.
Parameters
----------
ex_dividend : DataFrame
Ex-dividend for each stock and date
Returns
-------
dividend_weights : DataFrame
Weights for each stock and date
"""
#TODO: Implement function
dividend_cumsum_per_ticker = ex_dividend.T.cumsum().T
return dividend_cumsum_per_ticker/dividend_cumsum_per_ticker.sum()
project_tests.test_calculate_dividend_weights(calculate_dividend_weights)
```
### View Data
Let's generate the ETF weights using `calculate_dividend_weights` and view them using a heatmap.
```
etf_weights = calculate_dividend_weights(ex_dividend)
helper.plot_weights(etf_weights, 'ETF Weights')
```
## Returns
Implement `generate_returns` to generate the returns. Note this isn't log returns. Since we're not dealing with volatility, we don't have to use log returns.
```
def generate_returns(close):
"""
Generate returns for ticker and date.
Parameters
----------
close : DataFrame
Close price for each ticker and date
Returns
-------
returns : Dataframe
The returns for each ticker and date
"""
#TODO: Implement function
return (close.T / close.T.shift(1) -1).T
project_tests.test_generate_returns(generate_returns)
```
### View Data
Let's generate the closing returns using `generate_returns` and view them using a heatmap.
```
returns = generate_returns(close)
helper.plot_returns(returns, 'Close Returns')
```
## Weighted Returns
With the returns of each stock computed, we can use it to compute the returns for for an index or ETF. Implement `generate_weighted_returns` to create weighted returns using returns and weights for an Index or ETF.
```
def generate_weighted_returns(returns, weights):
"""
Generate weighted returns.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
weights : DataFrame
Weights for each ticker and date
Returns
-------
weighted_returns : DataFrame
Weighted returns for each ticker and date
"""
assert returns.index.equals(weights.index)
assert returns.columns.equals(weights.columns)
#TODO: Implement function
return returns * weights
project_tests.test_generate_weighted_returns(generate_weighted_returns)
```
### View Data
Let's generate the etf and index returns using `generate_weighted_returns` and view them using a heatmap.
```
index_weighted_returns = generate_weighted_returns(returns, index_weights)
etf_weighted_returns = generate_weighted_returns(returns, etf_weights)
helper.plot_returns(index_weighted_returns, 'Index Returns')
helper.plot_returns(etf_weighted_returns, 'ETF Returns')
```
## Cumulative Returns
Implement `calculate_cumulative_returns` to calculate the cumulative returns over time.
```
def calculate_cumulative_returns(returns):
"""
Calculate cumulative returns.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
Returns
-------
cumulative_returns : Pandas Series
Cumulative returns for each date
"""
#TODO: Implement function
return (pd.Series([0]).append(returns.sum()) + 1).cumprod().iloc[1:]
project_tests.test_calculate_cumulative_returns(calculate_cumulative_returns)
```
### View Data
Let's generate the etf and index cumulative returns using `calculate_cumulative_returns` and compare the two.
```
index_weighted_cumulative_returns = calculate_cumulative_returns(index_weighted_returns)
etf_weighted_cumulative_returns = calculate_cumulative_returns(etf_weighted_returns)
helper.plot_benchmark_returns(index_weighted_cumulative_returns, etf_weighted_cumulative_returns, 'Smart Beta ETF vs Index')
```
## Tracking Error
In order to check the performance of the smart beta portfolio, we can compare it against the index. Implement `tracking_error` to return the tracking error between the etf and index over time.
```
def tracking_error(index_weighted_cumulative_returns, etf_weighted_cumulative_returns):
"""
Calculate the tracking error.
Parameters
----------
index_weighted_cumulative_returns : Pandas Series
The weighted index Cumulative returns for each date
etf_weighted_cumulative_returns : Pandas Series
The weighted etf Cumulative returns for each date
Returns
-------
tracking_error : Pandas Series
The tracking error for each date
"""
assert index_weighted_cumulative_returns.index.equals(etf_weighted_cumulative_returns.index)
#TODO: Implement function
tracking_error = index_weighted_cumulative_returns - etf_weighted_cumulative_returns
return tracking_error
project_tests.test_tracking_error(tracking_error)
```
### View Data
Let's generate the tracking error using `tracking_error` and graph it over time.
```
smart_beta_tracking_error = tracking_error(index_weighted_cumulative_returns, etf_weighted_cumulative_returns)
helper.plot_tracking_error(smart_beta_tracking_error, 'Smart Beta Tracking Error')
```
# Part 2: Portfolio Optimization
In Part 2, you'll optimize the index you created in part 1. You'll use `cvxopt` to optimize the convex problem of finding the optimal weights for the portfolio. Just like before, we'll compare these results to the index.
## Covariance
Implement `get_covariance` to calculate the covariance of `returns` and `weighted_index_returns`. We'll use this to feed into our convex optimization function. By using covariance, we can prevent the optimizer from going all in on a few stocks.
```
def get_covariance(returns, weighted_index_returns):
"""
Calculate covariance matrices.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
weighted_index_returns : DataFrame
Weighted index returns for each ticker and date
Returns
-------
xtx, xty : (2 dimensional Ndarray, 1 dimensional Ndarray)
"""
assert returns.index.equals(weighted_index_returns.index)
assert returns.columns.equals(weighted_index_returns.columns)
#TODO: Implement function
returns = returns.fillna(0)
weighted_index_returns = weighted_index_returns.sum().fillna(0)
xtx = returns.dot(returns.T)
xty = returns.dot(np.matrix(weighted_index_returns).T)[0]
return xtx.values, xty.values
project_tests.test_get_covariance(get_covariance)
```
### View Data
Let's look the the covariance generated from `get_covariance`.
```
xtx, xty = get_covariance(returns, index_weighted_returns)
xtx = pd.DataFrame(xtx, returns.index, returns.index)
xty = pd.Series(xty, returns.index)
helper.plot_covariance(xty, xtx)
```
## Quadratic Programming
Now that you have the covariance, we can use this to optimize the weights. Implement `solve_qp` to return the optimal `x` in the convex function with the following constraints:
- Sum of all x is 1
- x >= 0
```
import cvxopt
def solve_qp(P, q):
"""
Find the solution for minimize 0.5P*x*x - q*x with the following constraints:
- sum of all x equals to 1
- All x are greater than or equal to 0
Parameters
----------
P : 2 dimensional Ndarray
q : 1 dimensional Ndarray
Returns
-------
x : 1 dimensional Ndarray
The solution for x
"""
assert len(P.shape) == 2
assert len(q.shape) == 1
assert P.shape[0] == P.shape[1] == q.shape[0]
#TODO: Implement function
nn = len(q)
g = cvxopt.spmatrix(-1, range(nn), range(nn))
a = cvxopt.matrix(np.ones(nn), (1,nn))
b = cvxopt.matrix(1.0)
h = cvxopt.matrix(np.zeros(nn))
P = cvxopt.matrix(P)
q = -cvxopt.matrix(q)
# Min cov
# Max return
cvxopt.solvers.options['show_progress'] = False
sol = cvxopt.solvers.qp(P, q, g, h, a, b)
if 'optimal' not in sol['status']:
return np.array([])
return np.array(sol['x']).flatten()
project_tests.test_solve_qp(solve_qp)
```
Run the following cell to generate optimal weights using `solve_qp`.
```
raw_optim_etf_weights = solve_qp(xtx.values, xty.values)
raw_optim_etf_weights_per_date = np.tile(raw_optim_etf_weights, (len(returns.columns), 1))
optim_etf_weights = pd.DataFrame(raw_optim_etf_weights_per_date.T, returns.index, returns.columns)
```
## Optimized Portfolio
With our optimized etf weights built using quadratic programming, let's compare it to the index. Run the next cell to calculate the optimized etf returns and compare the returns to the index returns.
```
optim_etf_returns = generate_weighted_returns(returns, optim_etf_weights)
optim_etf_cumulative_returns = calculate_cumulative_returns(optim_etf_returns)
helper.plot_benchmark_returns(index_weighted_cumulative_returns, optim_etf_cumulative_returns, 'Optimized ETF vs Index')
optim_etf_tracking_error = tracking_error(index_weighted_cumulative_returns, optim_etf_cumulative_returns)
helper.plot_tracking_error(optim_etf_tracking_error, 'Optimized ETF Tracking Error')
```
## Rebalance Portfolio
The optimized etf portfolio used different weights for each day. After calculating in transaction fees, this amount of turnover to the portfolio can reduce the total returns. Let's find the optimal times to rebalance the portfolio instead of doing it every day.
Implement `rebalance_portfolio` to rebalance a portfolio.
```
def rebalance_portfolio(returns, weighted_index_returns, shift_size, chunk_size):
"""
Get weights for each rebalancing of the portfolio.
Parameters
----------
returns : DataFrame
Returns for each ticker and date
weighted_index_returns : DataFrame
Weighted index returns for each ticker and date
shift_size : int
The number of days between each rebalance
chunk_size : int
The number of days to look in the past for rebalancing
Returns
-------
all_rebalance_weights : list of Ndarrays
The etf weights for each point they are rebalanced
"""
assert returns.index.equals(weighted_index_returns.index)
assert returns.columns.equals(weighted_index_returns.columns)
assert shift_size > 0
assert chunk_size >= 0
#TODO: Implement function
date_len = returns.shape[1]
all_rebalance_weights = []
for shift in range(chunk_size, date_len, shift_size):
start_idx = shift - chunk_size
xtx, xty = get_covariance(returns.iloc[:, start_idx:shift], weighted_index_returns.iloc[:, start_idx:shift])
all_rebalance_weights.append(solve_qp(xtx, xty))
return all_rebalance_weights
project_tests.test_rebalance_portfolio(rebalance_portfolio)
```
Run the following cell to rebalance the portfolio using `rebalance_portfolio`.
```
chunk_size = 250
shift_size = 5
all_rebalance_weights = rebalance_portfolio(returns, index_weighted_returns, shift_size, chunk_size)
```
## Portfolio Rebalance Cost
With the portfolio rebalanced, we need to use a metric to measure the cost of rebalancing the portfolio. Implement `get_rebalance_cost` to calculate the rebalance cost.
```
def get_rebalance_cost(all_rebalance_weights, shift_size, rebalance_count):
"""
Get the cost of all the rebalancing.
Parameters
----------
all_rebalance_weights : list of Ndarrays
ETF Returns for each ticker and date
shift_size : int
The number of days between each rebalance
rebalance_count : int
Number of times the portfolio was rebalanced
Returns
-------
rebalancing_cost : float
The cost of all the rebalancing
"""
assert shift_size > 0
assert rebalance_count > 0
#TODO: Implement function
all_rebalance_weights_df = pd.DataFrame(np.array(all_rebalance_weights))
rebalance_total = (all_rebalance_weights_df - all_rebalance_weights_df.shift(-1)).abs().sum().sum()
return (shift_size / rebalance_count) * rebalance_total
project_tests.test_get_rebalance_cost(get_rebalance_cost)
```
Run the following cell to get the rebalance cost from `get_rebalance_cost`.
```
unconstrained_costs = get_rebalance_cost(all_rebalance_weights, shift_size, returns.shape[1])
print(unconstrained_costs)
# IGNORE THIS CODE
# THIS CODE IS TEST CODE FOR BUILDING PROJECT
# THIS WILL BE REMOVED BEFORE FINAL PROJECT
# Error checking while refactoring
assert np.isclose(optim_etf_weights, np.load('check_data/po_weights.npy'), equal_nan=True).all()
assert np.isclose(optim_etf_tracking_error, np.load('check_data/po_tracking_error.npy'), equal_nan=True).all()
assert np.isclose(smart_beta_tracking_error, np.load('check_data/sb_tracking_error.npy'), equal_nan=True).all()
# Error checking while refactoring
assert np.isclose(unconstrained_costs, 0.10739965758876144), unconstrained_costs
```
## Submission
Now that you're done with the project, it's time to submit it. Click the submit button in the bottom right. One of our reviewers will give you feedback on your project with a pass or not passed grade. You can continue to the next section while you wait for feedback.
| github_jupyter |
**Outline of Steps**
+ Initialization
+ Download COCO detection data from http://cocodataset.org/#download
+ http://images.cocodataset.org/zips/train2014.zip <= train images
+ http://images.cocodataset.org/zips/val2014.zip <= validation images
+ http://images.cocodataset.org/annotations/annotations_trainval2014.zip <= train and validation annotations
+ Run this script to convert annotations in COCO format to VOC format
+ https://gist.github.com/chicham/6ed3842d0d2014987186#file-coco2pascal-py
+ Download pre-trained weights from https://pjreddie.com/darknet/yolo/
+ https://pjreddie.com/media/files/yolo.weights
+ Specify the directory of train annotations (train_annot_folder) and train images (train_image_folder)
+ Specify the directory of validation annotations (valid_annot_folder) and validation images (valid_image_folder)
+ Specity the path of pre-trained weights by setting variable *wt_path*
+ Construct equivalent network in Keras
+ Network arch from https://github.com/pjreddie/darknet/blob/master/cfg/yolo-voc.cfg
+ Load the pretrained weights
+ Perform training
+ Perform detection on an image with newly trained weights
+ Perform detection on an video with newly trained weights
# Initialization
```
from keras.models import Sequential, Model
from keras.layers import Reshape, Activation, Conv2D, Input, MaxPooling2D, BatchNormalization, Flatten, Dense, Lambda
from keras.layers.advanced_activations import LeakyReLU
from keras.callbacks import EarlyStopping, ModelCheckpoint, TensorBoard
from keras.optimizers import SGD, Adam, RMSprop
from keras.layers.merge import concatenate
import matplotlib.pyplot as plt
import keras.backend as K
import tensorflow as tf
import imgaug as ia
from tqdm import tqdm
from imgaug import augmenters as iaa
import numpy as np
import pickle
import os, cv2
from preprocessing import parse_annotation, BatchGenerator
from utils import WeightReader, decode_netout, draw_boxes, normalize
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = ""
%matplotlib inline
LABELS = ['person']
IMAGE_H, IMAGE_W = 416, 416
GRID_H, GRID_W = 13 , 13
BOX = 5
CLASS = len(LABELS)
CLASS_WEIGHTS = np.ones(CLASS, dtype='float32')
OBJ_THRESHOLD = 0.3#0.5
NMS_THRESHOLD = 0.3#0.45
ANCHORS = [0.57273, 0.677385, 1.87446, 2.06253, 3.33843, 5.47434, 7.88282, 3.52778, 9.77052, 9.16828]
NO_OBJECT_SCALE = 1.0
OBJECT_SCALE = 5.0
COORD_SCALE = 1.0
CLASS_SCALE = 1.0
BATCH_SIZE = 16
WARM_UP_BATCHES = 3
TRUE_BOX_BUFFER = 50
wt_path = 'yolo.weights'
train_image_folder = '/home/peng/data/coco/images/train2014/'
train_annot_folder = '/home/peng/data/coco/annotations/train2014ann/'
valid_image_folder = '/home/peng/data/coco/images/val2014/'
valid_annot_folder = '/home/peng/data/coco/annotations/val2014ann/'
```
# Construct the network
```
# the function to implement the orgnization layer (thanks to github.com/allanzelener/YAD2K)
def space_to_depth_x2(x):
return tf.space_to_depth(x, block_size=2)
input_image = Input(shape=(IMAGE_H, IMAGE_W, 3))
true_boxes = Input(shape=(1, 1, 1, TRUE_BOX_BUFFER , 4))
# Layer 1
x = Conv2D(32, (3,3), strides=(1,1), padding='same', name='conv_1', use_bias=False)(input_image)
x = BatchNormalization(name='norm_1')(x)
x = LeakyReLU(alpha=0.1)(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
# Layer 2
x = Conv2D(64, (3,3), strides=(1,1), padding='same', name='conv_2', use_bias=False)(x)
x = BatchNormalization(name='norm_2')(x)
x = LeakyReLU(alpha=0.1)(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
# Layer 3
x = Conv2D(128, (3,3), strides=(1,1), padding='same', name='conv_3', use_bias=False)(x)
x = BatchNormalization(name='norm_3')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 4
x = Conv2D(64, (1,1), strides=(1,1), padding='same', name='conv_4', use_bias=False)(x)
x = BatchNormalization(name='norm_4')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 5
x = Conv2D(128, (3,3), strides=(1,1), padding='same', name='conv_5', use_bias=False)(x)
x = BatchNormalization(name='norm_5')(x)
x = LeakyReLU(alpha=0.1)(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
# Layer 6
x = Conv2D(256, (3,3), strides=(1,1), padding='same', name='conv_6', use_bias=False)(x)
x = BatchNormalization(name='norm_6')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 7
x = Conv2D(128, (1,1), strides=(1,1), padding='same', name='conv_7', use_bias=False)(x)
x = BatchNormalization(name='norm_7')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 8
x = Conv2D(256, (3,3), strides=(1,1), padding='same', name='conv_8', use_bias=False)(x)
x = BatchNormalization(name='norm_8')(x)
x = LeakyReLU(alpha=0.1)(x)
x = MaxPooling2D(pool_size=(2, 2))(x)
# Layer 9
x = Conv2D(512, (3,3), strides=(1,1), padding='same', name='conv_9', use_bias=False)(x)
x = BatchNormalization(name='norm_9')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 10
x = Conv2D(256, (1,1), strides=(1,1), padding='same', name='conv_10', use_bias=False)(x)
x = BatchNormalization(name='norm_10')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 11
x = Conv2D(512, (3,3), strides=(1,1), padding='same', name='conv_11', use_bias=False)(x)
x = BatchNormalization(name='norm_11')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 12
x = Conv2D(256, (1,1), strides=(1,1), padding='same', name='conv_12', use_bias=False)(x)
x = BatchNormalization(name='norm_12')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 13
x = Conv2D(512, (3,3), strides=(1,1), padding='same', name='conv_13', use_bias=False)(x)
x = BatchNormalization(name='norm_13')(x)
x = LeakyReLU(alpha=0.1)(x)
skip_connection = x
x = MaxPooling2D(pool_size=(2, 2))(x)
# Layer 14
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_14', use_bias=False)(x)
x = BatchNormalization(name='norm_14')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 15
x = Conv2D(512, (1,1), strides=(1,1), padding='same', name='conv_15', use_bias=False)(x)
x = BatchNormalization(name='norm_15')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 16
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_16', use_bias=False)(x)
x = BatchNormalization(name='norm_16')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 17
x = Conv2D(512, (1,1), strides=(1,1), padding='same', name='conv_17', use_bias=False)(x)
x = BatchNormalization(name='norm_17')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 18
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_18', use_bias=False)(x)
x = BatchNormalization(name='norm_18')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 19
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_19', use_bias=False)(x)
x = BatchNormalization(name='norm_19')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 20
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_20', use_bias=False)(x)
x = BatchNormalization(name='norm_20')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 21
skip_connection = Conv2D(64, (1,1), strides=(1,1), padding='same', name='conv_21', use_bias=False)(skip_connection)
skip_connection = BatchNormalization(name='norm_21')(skip_connection)
skip_connection = LeakyReLU(alpha=0.1)(skip_connection)
skip_connection = Lambda(space_to_depth_x2)(skip_connection)
x = concatenate([skip_connection, x])
# Layer 22
x = Conv2D(1024, (3,3), strides=(1,1), padding='same', name='conv_22', use_bias=False)(x)
x = BatchNormalization(name='norm_22')(x)
x = LeakyReLU(alpha=0.1)(x)
# Layer 23
x = Conv2D(BOX * (4 + 1 + CLASS), (1,1), strides=(1,1), padding='same', name='conv_23')(x)
output = Reshape((GRID_H, GRID_W, BOX, 4 + 1 + CLASS))(x)
# small hack to allow true_boxes to be registered when Keras build the model
# for more information: https://github.com/fchollet/keras/issues/2790
output = Lambda(lambda args: args[0])([output, true_boxes])
model = Model([input_image, true_boxes], output)
model.summary()
```
# Load pretrained weights
**Load the weights originally provided by YOLO**
```
weight_reader = WeightReader(wt_path)
weight_reader.reset()
nb_conv = 23
for i in range(1, nb_conv+1):
conv_layer = model.get_layer('conv_' + str(i))
if i < nb_conv:
norm_layer = model.get_layer('norm_' + str(i))
size = np.prod(norm_layer.get_weights()[0].shape)
beta = weight_reader.read_bytes(size)
gamma = weight_reader.read_bytes(size)
mean = weight_reader.read_bytes(size)
var = weight_reader.read_bytes(size)
weights = norm_layer.set_weights([gamma, beta, mean, var])
if len(conv_layer.get_weights()) > 1:
bias = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[1].shape))
kernel = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[0].shape))
kernel = kernel.reshape(list(reversed(conv_layer.get_weights()[0].shape)))
kernel = kernel.transpose([2,3,1,0])
conv_layer.set_weights([kernel, bias])
else:
kernel = weight_reader.read_bytes(np.prod(conv_layer.get_weights()[0].shape))
kernel = kernel.reshape(list(reversed(conv_layer.get_weights()[0].shape)))
kernel = kernel.transpose([2,3,1,0])
conv_layer.set_weights([kernel])
```
**Randomize weights of the last layer**
```
layer = model.layers[-4] # the last convolutional layer
weights = layer.get_weights()
new_kernel = np.random.normal(size=weights[0].shape)/(GRID_H*GRID_W)
new_bias = np.random.normal(size=weights[1].shape)/(GRID_H*GRID_W)
layer.set_weights([new_kernel, new_bias])
```
# Perform training
**Loss function**
$$\begin{multline}
\lambda_\textbf{coord}
\sum_{i = 0}^{S^2}
\sum_{j = 0}^{B}
L_{ij}^{\text{obj}}
\left[
\left(
x_i - \hat{x}_i
\right)^2 +
\left(
y_i - \hat{y}_i
\right)^2
\right]
\\
+ \lambda_\textbf{coord}
\sum_{i = 0}^{S^2}
\sum_{j = 0}^{B}
L_{ij}^{\text{obj}}
\left[
\left(
\sqrt{w_i} - \sqrt{\hat{w}_i}
\right)^2 +
\left(
\sqrt{h_i} - \sqrt{\hat{h}_i}
\right)^2
\right]
\\
+ \sum_{i = 0}^{S^2}
\sum_{j = 0}^{B}
L_{ij}^{\text{obj}}
\left(
C_i - \hat{C}_i
\right)^2
\\
+ \lambda_\textrm{noobj}
\sum_{i = 0}^{S^2}
\sum_{j = 0}^{B}
L_{ij}^{\text{noobj}}
\left(
C_i - \hat{C}_i
\right)^2
\\
+ \sum_{i = 0}^{S^2}
L_i^{\text{obj}}
\sum_{c \in \textrm{classes}}
\left(
p_i(c) - \hat{p}_i(c)
\right)^2
\end{multline}$$
```
def custom_loss(y_true, y_pred):
mask_shape = tf.shape(y_true)[:4]
cell_x = tf.to_float(tf.reshape(tf.tile(tf.range(GRID_W), [GRID_H]), (1, GRID_H, GRID_W, 1, 1)))
cell_y = tf.transpose(cell_x, (0,2,1,3,4))
cell_grid = tf.tile(tf.concat([cell_x,cell_y], -1), [BATCH_SIZE, 1, 1, 5, 1])
coord_mask = tf.zeros(mask_shape)
conf_mask = tf.zeros(mask_shape)
class_mask = tf.zeros(mask_shape)
seen = tf.Variable(0.)
total_recall = tf.Variable(0.)
"""
Adjust prediction
"""
### adjust x and y
pred_box_xy = tf.sigmoid(y_pred[..., :2]) + cell_grid
### adjust w and h
pred_box_wh = tf.exp(y_pred[..., 2:4]) * np.reshape(ANCHORS, [1,1,1,BOX,2])
### adjust confidence
pred_box_conf = tf.sigmoid(y_pred[..., 4])
### adjust class probabilities
pred_box_class = y_pred[..., 5:]
"""
Adjust ground truth
"""
### adjust x and y
true_box_xy = y_true[..., 0:2] # relative position to the containing cell
### adjust w and h
true_box_wh = y_true[..., 2:4] # number of cells accross, horizontally and vertically
### adjust confidence
true_wh_half = true_box_wh / 2.
true_mins = true_box_xy - true_wh_half
true_maxes = true_box_xy + true_wh_half
pred_wh_half = pred_box_wh / 2.
pred_mins = pred_box_xy - pred_wh_half
pred_maxes = pred_box_xy + pred_wh_half
intersect_mins = tf.maximum(pred_mins, true_mins)
intersect_maxes = tf.minimum(pred_maxes, true_maxes)
intersect_wh = tf.maximum(intersect_maxes - intersect_mins, 0.)
intersect_areas = intersect_wh[..., 0] * intersect_wh[..., 1]
true_areas = true_box_wh[..., 0] * true_box_wh[..., 1]
pred_areas = pred_box_wh[..., 0] * pred_box_wh[..., 1]
union_areas = pred_areas + true_areas - intersect_areas
iou_scores = tf.truediv(intersect_areas, union_areas)
true_box_conf = iou_scores * y_true[..., 4]
### adjust class probabilities
true_box_class = tf.argmax(y_true[..., 5:], -1)
"""
Determine the masks
"""
### coordinate mask: simply the position of the ground truth boxes (the predictors)
coord_mask = tf.expand_dims(y_true[..., 4], axis=-1) * COORD_SCALE
### confidence mask: penelize predictors + penalize boxes with low IOU
# penalize the confidence of the boxes, which have IOU with some ground truth box < 0.6
true_xy = true_boxes[..., 0:2]
true_wh = true_boxes[..., 2:4]
true_wh_half = true_wh / 2.
true_mins = true_xy - true_wh_half
true_maxes = true_xy + true_wh_half
pred_xy = tf.expand_dims(pred_box_xy, 4)
pred_wh = tf.expand_dims(pred_box_wh, 4)
pred_wh_half = pred_wh / 2.
pred_mins = pred_xy - pred_wh_half
pred_maxes = pred_xy + pred_wh_half
intersect_mins = tf.maximum(pred_mins, true_mins)
intersect_maxes = tf.minimum(pred_maxes, true_maxes)
intersect_wh = tf.maximum(intersect_maxes - intersect_mins, 0.)
intersect_areas = intersect_wh[..., 0] * intersect_wh[..., 1]
true_areas = true_wh[..., 0] * true_wh[..., 1]
pred_areas = pred_wh[..., 0] * pred_wh[..., 1]
union_areas = pred_areas + true_areas - intersect_areas
iou_scores = tf.truediv(intersect_areas, union_areas)
best_ious = tf.reduce_max(iou_scores, axis=4)
conf_mask = conf_mask + tf.to_float(best_ious < 0.6) * (1 - y_true[..., 4]) * NO_OBJECT_SCALE
# penalize the confidence of the boxes, which are reponsible for corresponding ground truth box
conf_mask = conf_mask + y_true[..., 4] * OBJECT_SCALE
### class mask: simply the position of the ground truth boxes (the predictors)
class_mask = y_true[..., 4] * tf.gather(CLASS_WEIGHTS, true_box_class) * CLASS_SCALE
"""
Warm-up training
"""
no_boxes_mask = tf.to_float(coord_mask < COORD_SCALE/2.)
seen = tf.assign_add(seen, 1.)
true_box_xy, true_box_wh, coord_mask = tf.cond(tf.less(seen, WARM_UP_BATCHES),
lambda: [true_box_xy + (0.5 + cell_grid) * no_boxes_mask,
true_box_wh + tf.ones_like(true_box_wh) * np.reshape(ANCHORS, [1,1,1,BOX,2]) * no_boxes_mask,
tf.ones_like(coord_mask)],
lambda: [true_box_xy,
true_box_wh,
coord_mask])
"""
Finalize the loss
"""
nb_coord_box = tf.reduce_sum(tf.to_float(coord_mask > 0.0))
nb_conf_box = tf.reduce_sum(tf.to_float(conf_mask > 0.0))
nb_class_box = tf.reduce_sum(tf.to_float(class_mask > 0.0))
loss_xy = tf.reduce_sum(tf.square(true_box_xy-pred_box_xy) * coord_mask) / (nb_coord_box + 1e-6) / 2.
loss_wh = tf.reduce_sum(tf.square(true_box_wh-pred_box_wh) * coord_mask) / (nb_coord_box + 1e-6) / 2.
loss_conf = tf.reduce_sum(tf.square(true_box_conf-pred_box_conf) * conf_mask) / (nb_conf_box + 1e-6) / 2.
loss_class = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=true_box_class, logits=pred_box_class)
loss_class = tf.reduce_sum(loss_class * class_mask) / (nb_class_box + 1e-6)
loss = loss_xy + loss_wh + loss_conf + loss_class
nb_true_box = tf.reduce_sum(y_true[..., 4])
nb_pred_box = tf.reduce_sum(tf.to_float(true_box_conf > 0.5) * tf.to_float(pred_box_conf > 0.3))
"""
Debugging code
"""
current_recall = nb_pred_box/(nb_true_box + 1e-6)
total_recall = tf.assign_add(total_recall, current_recall)
loss = tf.Print(loss, [tf.zeros((1))], message='Dummy Line \t', summarize=1000)
loss = tf.Print(loss, [loss_xy], message='Loss XY \t', summarize=1000)
loss = tf.Print(loss, [loss_wh], message='Loss WH \t', summarize=1000)
loss = tf.Print(loss, [loss_conf], message='Loss Conf \t', summarize=1000)
loss = tf.Print(loss, [loss_class], message='Loss Class \t', summarize=1000)
loss = tf.Print(loss, [loss], message='Total Loss \t', summarize=1000)
loss = tf.Print(loss, [current_recall], message='Current Recall \t', summarize=1000)
loss = tf.Print(loss, [total_recall/seen], message='Average Recall \t', summarize=1000)
return loss
```
**Parse the annotations to construct train generator and validation generator**
```
generator_config = {
'IMAGE_H' : IMAGE_H,
'IMAGE_W' : IMAGE_W,
'GRID_H' : GRID_H,
'GRID_W' : GRID_W,
'BOX' : BOX,
'LABELS' : LABELS,
'CLASS' : len(LABELS),
'ANCHORS' : ANCHORS,
'BATCH_SIZE' : BATCH_SIZE,
'TRUE_BOX_BUFFER' : 50,
}
train_imgs, seen_train_labels = parse_annotation(train_annot_folder, train_image_folder, labels=LABELS)
### write parsed annotations to pickle for fast retrieval next time
#with open('train_imgs', 'wb') as fp:
# pickle.dump(train_imgs, fp)
### read saved pickle of parsed annotations
#with open ('train_imgs', 'rb') as fp:
# train_imgs = pickle.load(fp)
train_batch = BatchGenerator(train_imgs, generator_config, norm=normalize)
valid_imgs, seen_valid_labels = parse_annotation(valid_annot_folder, valid_image_folder, labels=LABELS)
### write parsed annotations to pickle for fast retrieval next time
#with open('valid_imgs', 'wb') as fp:
# pickle.dump(valid_imgs, fp)
### read saved pickle of parsed annotations
#with open ('valid_imgs', 'rb') as fp:
# valid_imgs = pickle.load(fp)
valid_batch = BatchGenerator(valid_imgs, generator_config, norm=normalize, jitter=False)
```
**Setup a few callbacks and start the training**
```
early_stop = EarlyStopping(monitor='val_loss',
min_delta=0.001,
patience=3,
mode='min',
verbose=1)
checkpoint = ModelCheckpoint('weights_coco.h5',
monitor='val_loss',
verbose=1,
save_best_only=True,
mode='min',
period=1)
tb_counter = len([log for log in os.listdir(os.path.expanduser('~/logs/')) if 'coco_' in log]) + 1
tensorboard = TensorBoard(log_dir=os.path.expanduser('~/logs/') + 'coco_' + '_' + str(tb_counter),
histogram_freq=0,
write_graph=True,
write_images=False)
optimizer = Adam(lr=0.5e-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
#optimizer = SGD(lr=1e-4, decay=0.0005, momentum=0.9)
#optimizer = RMSprop(lr=1e-4, rho=0.9, epsilon=1e-08, decay=0.0)
model.compile(loss=custom_loss, optimizer=optimizer)
model.fit_generator(generator = train_batch,
steps_per_epoch = len(train_batch),
epochs = 100,
verbose = 1,
validation_data = valid_batch,
validation_steps = len(valid_batch),
callbacks = [early_stop, checkpoint, tensorboard],
max_queue_size = 3)
```
# Perform detection on image
```
model.load_weights("weights_coco.h5")
dummy_array = np.zeros((1,1,1,1,TRUE_BOX_BUFFER,4))
image = cv2.imread('images/giraffe.jpg')
plt.figure(figsize=(10,10))
input_image = cv2.resize(image, (416, 416))
input_image = input_image / 255.
input_image = input_image[:,:,::-1]
input_image = np.expand_dims(input_image, 0)
netout = model.predict([input_image, dummy_array])
boxes = decode_netout(netout[0],
obj_threshold=OBJ_THRESHOLD,
nms_threshold=NMS_THRESHOLD,
anchors=ANCHORS,
nb_class=CLASS)
image = draw_boxes(image, boxes, labels=LABELS)
plt.imshow(image[:,:,::-1]); plt.show()
```
# Perform detection on video
```
model.load_weights("weights_coco.h5")
dummy_array = np.zeros((1,1,1,1,TRUE_BOX_BUFFER,4))
video_inp = '../basic-yolo-keras/images/phnom_penh.mp4'
video_out = '../basic-yolo-keras/images/phnom_penh_bbox.mp4'
video_reader = cv2.VideoCapture(video_inp)
nb_frames = int(video_reader.get(cv2.CAP_PROP_FRAME_COUNT))
frame_h = int(video_reader.get(cv2.CAP_PROP_FRAME_HEIGHT))
frame_w = int(video_reader.get(cv2.CAP_PROP_FRAME_WIDTH))
video_writer = cv2.VideoWriter(video_out,
cv2.VideoWriter_fourcc(*'XVID'),
50.0,
(frame_w, frame_h))
for i in tqdm(range(nb_frames)):
ret, image = video_reader.read()
input_image = cv2.resize(image, (416, 416))
input_image = input_image / 255.
input_image = input_image[:,:,::-1]
input_image = np.expand_dims(input_image, 0)
netout = model.predict([input_image, dummy_array])
boxes = decode_netout(netout[0],
obj_threshold=0.3,
nms_threshold=NMS_THRESHOLD,
anchors=ANCHORS,
nb_class=CLASS)
image = draw_boxes(image, boxes, labels=LABELS)
video_writer.write(np.uint8(image))
video_reader.release()
video_writer.release()
```
| github_jupyter |
# Reading JWST ASDF-in-FITS with `astrowidgets`
This is a proof-of-concept using `astrowidgets` to read in JWST ASDF-in-FITS data. As it is using the dev versions of several different packages, this notebook is not guaranteed to work as-is in the future. As Ginga is primarily an image viewer, we will not concern ourselves with spectrocopic data models in this notebook.
Relevant specs used in testing (more or less):
* Python 3.7.3
* `aggdraw` 1.3.11
* `asdf` 2.4.0.dev
* `astropy` 4.0.dev
* `astrowidgets` 0.1.0.dev
* `ginga` 3.0.dev (https://github.com/ejeschke/ginga/pull/781 and https://github.com/ejeschke/ginga/pull/764)
* `gwcs` 0.12.dev
* `ipyevents` 0.6.2
* `ipywidgets` 7.5.0
* `jsonschema` 2.6.0
* `jupyter` 1.0.0
* `jupyter_client` 5.3.1
* `jupyter_console` 6.0.0
* `jupyter_core` 4.5.0
* `jwst` 0.13.8a0.dev
* `notebook` 6.0.0
* `numpy` 1.16.4
* `opencv` 3.4.2
* `scipy` 1.3.0
* `stginga` 1.1.dev308 (https://github.com/spacetelescope/stginga/pull/177 and https://github.com/spacetelescope/stginga/pull/179)
```
import warnings
from astropy.io import fits
from astrowidgets import ImageWidget
from ginga.misc.log import get_logger
from ginga.util import wcsmod
```
We need to ask Ginga to explicitly use its `astropy_ape14` WCS interface. This is unnecessary if every user sets it in their `~/.ginga/general.cfg` but that is not always guaranteed.
```
wcsmod.use('astropy_ape14')
```
This notebook assumes that you have downloaded a JWST data file of interest into the working directory. Example file can be found at https://stsci.box.com/s/hwrc5reqygmmv2rl3yvvz90l7ryjac6h (requires permission for access).
```
filename = 'jw1069001001_01203_00002_nrca1_level2_cal.fits'
```
The following cell should show a multi-extension FITS with an ASDF-in-FITS extension. For example:
Filename: jw1069001001_01203_00002_nrca1_level2_cal.fits
No. Name Ver Type Cards Dimensions Format
0 PRIMARY 1 PrimaryHDU 232 ()
1 SCI 1 ImageHDU 53 (2048, 2048) float32
2 ERR 1 ImageHDU 10 (2048, 2048) float32
3 DQ 1 ImageHDU 11 (2048, 2048) int32 (rescales to uint32)
4 AREA 1 ImageHDU 9 (2048, 2048) float32
5 VAR_POISSON 1 ImageHDU 9 (2048, 2048) float32
6 VAR_RNOISE 1 ImageHDU 9 (2048, 2048) float32
7 ASDF 1 BinTableHDU 11 1R x 1C [14057B]
```
with fits.open(filename) as pf:
pf.info()
```
Then, we customize our image widget by subclassing `ImageWidget` and adding a method to load the file. In the future, after https://github.com/astropy/astrowidgets/pull/78 is merged, this subclassing will be unnecessary and we will be able to use `ImageWidget` directly.
```
class JWSTImageWidget(ImageWidget):
def load_file(self, filename):
from ginga.util.io import io_asdf
# Even if jwst package is not called directly, it is used
# to load the correct GWCS data models behind the scenes.
image = io_asdf.load_file(filename)
self._viewer.set_image(image)
```
We define a Ginga logger to go with our image widget. This logger prints out error messages to screen directly.
```
logger = get_logger('my viewer', log_stderr=True, log_file=None, level=40)
```
We create the widget instance. This would be the thing that you interface with for widget magic.
```
w = JWSTImageWidget(logger=logger)
```
We load our JWST data file into our widget instance.
```
with warnings.catch_warnings():
warnings.simplefilter('ignore') # Ignore validation warning
w.load_file(filename)
```
This would display the widget. When you mouse over the pixels, you would see coordinates information (both pixels and sky) change. See https://astrowidgets.readthedocs.io/en/latest/ for documentation on `astrowidgets`.
```
w
```
Let's change the colormap, the stretch, and the cuts.
```
w.set_colormap('viridis_r')
w.stretch = 'log'
w.cuts = 'histogram'
```
Now, we mark some stars of interest by running the following cell and then click on the widget above. You will see a marker appear where you click each time. We will also customize how the marker would show in the viewer.
```
marker_params = {'type': 'circle', 'color': 'red', 'radius': 10,
'linewidth': 2}
w.start_marking(marker_name='demo', marker=marker_params)
print(w.is_marking)
```
When we are done, we run the following cell to stop marking.
```
w.stop_marking()
print(w.is_marking)
```
We can see the points we manually selected above.
```
tab = w.get_markers(marker_name='all')
tab.pprint_all()
```
Optional: Delete all the markers to start over marking objects again above.
```
w.reset_markers()
```
<img style="float: right;" src="https://raw.githubusercontent.com/spacetelescope/notebooks/master/assets/stsci_pri_combo_mark_horizonal_white_bkgd.png" alt="Space Telescope Logo" width="200px"/>
| github_jupyter |
# PRMT 2241: What is the average message size for GP2GP messages including / excluding attachments?
In order for the Practice Migration team to estimate the potential transfer speed of a Migration Data Pipeline and provide a basis to the total potential total size of a bulk data set.
Two questions:
What is the average message size for GP2GP messages including attachments?
What is the average message size for GP2GP messages Excluding attachments? <- we won't be able to answer this as we only have attachment size data and not core extract size data
### Requirements
In order to replicate this notebook, perform the following steps:
1. Log into Splunk and run the following query for:
- 01/06/2021 00:00:00:00 to 01/07/2021 00:00:00, export the result as a csv named `6-2021-attachment-metadata.csv` and gzip it.
```
index="spine2vfmmonitor" logReference=MPS0208
| table _time, attachmentID, conversationID, FromSystem, ToSystem, attachmentType, Compressed, ContentType, LargeAttachment, Length, OriginalBase64, internalID
```
2. Run the following Splunk query for the same time range. Export the results as a csv named `6-2021-gp2gp-messages.csv` and gzip it.
```
index="spine2vfmmonitor" service="gp2gp" logReference="MPS0053d"
| table _time, conversationID, internalID, interactionID
```
```
import pandas as pd
import numpy as np
import datetime
attachment_metadata_file = "s3://prm-gp2gp-data-sandbox-dev/PRMT-2240-tpp-attachment-limit/6-2021-attachment-metadata.csv.gz"
attachments = pd.read_csv(attachment_metadata_file, parse_dates=["_time"], na_values=["Unknown"], dtype={"Length": pd.Int64Dtype()})
gp2gp_messages_file = "s3://prm-gp2gp-data-sandbox-dev/PRMT-2240-tpp-attachment-limit/6-2021-gp2gp-messages.csv.gz"
gp2gp_messages = pd.read_csv(gp2gp_messages_file, parse_dates=["_time"])
```
## Deduplicate attachment data
```
ehr_request_completed_messages = gp2gp_messages[gp2gp_messages["interactionID"] == "urn:nhs:names:services:gp2gp/RCMR_IN030000UK06"]
unique_ehr_request_completed_messages = ehr_request_completed_messages.sort_values(by="_time").drop_duplicates(subset=["conversationID"], keep="last")
unique_ehr_request_completed_messages.shape
ehr_attachments = pd.merge(attachments, unique_ehr_request_completed_messages[["internalID", "interactionID"]], on="internalID", how="inner")
```
## Find total average size of attachments for a transfer
```
attachments_grouped_by_conversation = ehr_attachments.groupby(by="conversationID").agg({"Length": "sum"})
attachments_grouped_by_conversation["Length in Mb"] = attachments_grouped_by_conversation["Length"].fillna(0)/(1024**2)
attachments_grouped_by_conversation["Length in Mb"].describe()
attachments_grouped_by_conversation.boxplot(column="Length in Mb")
attachments_grouped_by_conversation.hist(column="Length in Mb", bins=40)
```
| github_jupyter |
CER040 - Install signed Management Proxy certificate
====================================================
This notebook installs into the Big Data Cluster the certificate signed
using:
- [CER030 - Sign Management Proxy certificate with generated
CA](../cert-management/cer030-sign-service-proxy-generated-cert.ipynb)
Steps
-----
### Parameters
```
app_name = "mgmtproxy"
scaledset_name = "mgmtproxy"
container_name = "service-proxy"
prefix_keyfile_name = "service-proxy"
common_name = "mgmtproxy-svc"
test_cert_store_root = "/var/opt/secrets/test-certificates"
```
### Common functions
Define helper functions used in this notebook.
```
# Define `run` function for transient fault handling, suggestions on error, and scrolling updates on Windows
import sys
import os
import re
import json
import platform
import shlex
import shutil
import datetime
from subprocess import Popen, PIPE
from IPython.display import Markdown
retry_hints = {} # Output in stderr known to be transient, therefore automatically retry
error_hints = {} # Output in stderr where a known SOP/TSG exists which will be HINTed for further help
install_hint = {} # The SOP to help install the executable if it cannot be found
first_run = True
rules = None
debug_logging = False
def run(cmd, return_output=False, no_output=False, retry_count=0):
"""Run shell command, stream stdout, print stderr and optionally return output
NOTES:
1. Commands that need this kind of ' quoting on Windows e.g.:
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='data-pool')].metadata.name}
Need to actually pass in as '"':
kubectl get nodes -o jsonpath={.items[?(@.metadata.annotations.pv-candidate=='"'data-pool'"')].metadata.name}
The ' quote approach, although correct when pasting into Windows cmd, will hang at the line:
`iter(p.stdout.readline, b'')`
The shlex.split call does the right thing for each platform, just use the '"' pattern for a '
"""
MAX_RETRIES = 5
output = ""
retry = False
global first_run
global rules
if first_run:
first_run = False
rules = load_rules()
# When running `azdata sql query` on Windows, replace any \n in """ strings, with " ", otherwise we see:
#
# ('HY090', '[HY090] [Microsoft][ODBC Driver Manager] Invalid string or buffer length (0) (SQLExecDirectW)')
#
if platform.system() == "Windows" and cmd.startswith("azdata sql query"):
cmd = cmd.replace("\n", " ")
# shlex.split is required on bash and for Windows paths with spaces
#
cmd_actual = shlex.split(cmd)
# Store this (i.e. kubectl, python etc.) to support binary context aware error_hints and retries
#
user_provided_exe_name = cmd_actual[0].lower()
# When running python, use the python in the ADS sandbox ({sys.executable})
#
if cmd.startswith("python "):
cmd_actual[0] = cmd_actual[0].replace("python", sys.executable)
# On Mac, when ADS is not launched from terminal, LC_ALL may not be set, which causes pip installs to fail
# with:
#
# UnicodeDecodeError: 'ascii' codec can't decode byte 0xc5 in position 4969: ordinal not in range(128)
#
# Setting it to a default value of "en_US.UTF-8" enables pip install to complete
#
if platform.system() == "Darwin" and "LC_ALL" not in os.environ:
os.environ["LC_ALL"] = "en_US.UTF-8"
# When running `kubectl`, if AZDATA_OPENSHIFT is set, use `oc`
#
if cmd.startswith("kubectl ") and "AZDATA_OPENSHIFT" in os.environ:
cmd_actual[0] = cmd_actual[0].replace("kubectl", "oc")
# To aid supportabilty, determine which binary file will actually be executed on the machine
#
which_binary = None
# Special case for CURL on Windows. The version of CURL in Windows System32 does not work to
# get JWT tokens, it returns "(56) Failure when receiving data from the peer". If another instance
# of CURL exists on the machine use that one. (Unfortunately the curl.exe in System32 is almost
# always the first curl.exe in the path, and it can't be uninstalled from System32, so here we
# look for the 2nd installation of CURL in the path)
if platform.system() == "Windows" and cmd.startswith("curl "):
path = os.getenv('PATH')
for p in path.split(os.path.pathsep):
p = os.path.join(p, "curl.exe")
if os.path.exists(p) and os.access(p, os.X_OK):
if p.lower().find("system32") == -1:
cmd_actual[0] = p
which_binary = p
break
# Find the path based location (shutil.which) of the executable that will be run (and display it to aid supportability), this
# seems to be required for .msi installs of azdata.cmd/az.cmd. (otherwise Popen returns FileNotFound)
#
# NOTE: Bash needs cmd to be the list of the space separated values hence shlex.split.
#
if which_binary == None:
which_binary = shutil.which(cmd_actual[0])
if which_binary == None:
if user_provided_exe_name in install_hint and install_hint[user_provided_exe_name] is not None:
display(Markdown(f'HINT: Use [{install_hint[user_provided_exe_name][0]}]({install_hint[user_provided_exe_name][1]}) to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)")
else:
cmd_actual[0] = which_binary
start_time = datetime.datetime.now().replace(microsecond=0)
print(f"START: {cmd} @ {start_time} ({datetime.datetime.utcnow().replace(microsecond=0)} UTC)")
print(f" using: {which_binary} ({platform.system()} {platform.release()} on {platform.machine()})")
print(f" cwd: {os.getcwd()}")
# Command-line tools such as CURL and AZDATA HDFS commands output
# scrolling progress bars, which causes Jupyter to hang forever, to
# workaround this, use no_output=True
#
# Work around a infinite hang when a notebook generates a non-zero return code, break out, and do not wait
#
wait = True
try:
if no_output:
p = Popen(cmd_actual)
else:
p = Popen(cmd_actual, stdout=PIPE, stderr=PIPE, bufsize=1)
with p.stdout:
for line in iter(p.stdout.readline, b''):
line = line.decode()
if return_output:
output = output + line
else:
if cmd.startswith("azdata notebook run"): # Hyperlink the .ipynb file
regex = re.compile(' "(.*)"\: "(.*)"')
match = regex.match(line)
if match:
if match.group(1).find("HTML") != -1:
display(Markdown(f' - "{match.group(1)}": "{match.group(2)}"'))
else:
display(Markdown(f' - "{match.group(1)}": "[{match.group(2)}]({match.group(2)})"'))
wait = False
break # otherwise infinite hang, have not worked out why yet.
else:
print(line, end='')
if rules is not None:
apply_expert_rules(line)
if wait:
p.wait()
except FileNotFoundError as e:
if install_hint is not None:
display(Markdown(f'HINT: Use {install_hint} to resolve this issue.'))
raise FileNotFoundError(f"Executable '{cmd_actual[0]}' not found in path (where/which)") from e
exit_code_workaround = 0 # WORKAROUND: azdata hangs on exception from notebook on p.wait()
if not no_output:
for line in iter(p.stderr.readline, b''):
try:
line_decoded = line.decode()
except UnicodeDecodeError:
# NOTE: Sometimes we get characters back that cannot be decoded(), e.g.
#
# \xa0
#
# For example see this in the response from `az group create`:
#
# ERROR: Get Token request returned http error: 400 and server
# response: {"error":"invalid_grant",# "error_description":"AADSTS700082:
# The refresh token has expired due to inactivity.\xa0The token was
# issued on 2018-10-25T23:35:11.9832872Z
#
# which generates the exception:
#
# UnicodeDecodeError: 'utf-8' codec can't decode byte 0xa0 in position 179: invalid start byte
#
print("WARNING: Unable to decode stderr line, printing raw bytes:")
print(line)
line_decoded = ""
pass
else:
# azdata emits a single empty line to stderr when doing an hdfs cp, don't
# print this empty "ERR:" as it confuses.
#
if line_decoded == "":
continue
print(f"STDERR: {line_decoded}", end='')
if line_decoded.startswith("An exception has occurred") or line_decoded.startswith("ERROR: An error occurred while executing the following cell"):
exit_code_workaround = 1
# inject HINTs to next TSG/SOP based on output in stderr
#
if user_provided_exe_name in error_hints:
for error_hint in error_hints[user_provided_exe_name]:
if line_decoded.find(error_hint[0]) != -1:
display(Markdown(f'HINT: Use [{error_hint[1]}]({error_hint[2]}) to resolve this issue.'))
# apply expert rules (to run follow-on notebooks), based on output
#
if rules is not None:
apply_expert_rules(line_decoded)
# Verify if a transient error, if so automatically retry (recursive)
#
if user_provided_exe_name in retry_hints:
for retry_hint in retry_hints[user_provided_exe_name]:
if line_decoded.find(retry_hint) != -1:
if retry_count < MAX_RETRIES:
print(f"RETRY: {retry_count} (due to: {retry_hint})")
retry_count = retry_count + 1
output = run(cmd, return_output=return_output, retry_count=retry_count)
if return_output:
return output
else:
return
elapsed = datetime.datetime.now().replace(microsecond=0) - start_time
# WORKAROUND: We avoid infinite hang above in the `azdata notebook run` failure case, by inferring success (from stdout output), so
# don't wait here, if success known above
#
if wait:
if p.returncode != 0:
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(p.returncode)}.\n')
else:
if exit_code_workaround !=0 :
raise SystemExit(f'Shell command:\n\n\t{cmd} ({elapsed}s elapsed)\n\nreturned non-zero exit code: {str(exit_code_workaround)}.\n')
print(f'\nSUCCESS: {elapsed}s elapsed.\n')
if return_output:
return output
def load_json(filename):
"""Load a json file from disk and return the contents"""
with open(filename, encoding="utf8") as json_file:
return json.load(json_file)
def load_rules():
"""Load any 'expert rules' from the metadata of this notebook (.ipynb) that should be applied to the stderr of the running executable"""
try:
# Load this notebook as json to get access to the expert rules in the notebook metadata.
#
j = load_json("cer040-install-service-proxy-cert.ipynb")
except:
pass # If the user has renamed the book, we can't load ourself. NOTE: Is there a way in Jupyter, to know your own filename?
else:
if "metadata" in j and \
"azdata" in j["metadata"] and \
"expert" in j["metadata"]["azdata"] and \
"rules" in j["metadata"]["azdata"]["expert"]:
rules = j["metadata"]["azdata"]["expert"]["rules"]
rules.sort() # Sort rules, so they run in priority order (the [0] element). Lowest value first.
# print (f"EXPERT: There are {len(rules)} rules to evaluate.")
return rules
def apply_expert_rules(line):
"""Determine if the stderr line passed in, matches the regular expressions for any of the 'expert rules', if so
inject a 'HINT' to the follow-on SOP/TSG to run"""
global rules
for rule in rules:
# rules that have 9 elements are the injected (output) rules (the ones we want). Rules
# with only 8 elements are the source (input) rules, which are not expanded (i.e. TSG029,
# not ../repair/tsg029-nb-name.ipynb)
if len(rule) == 9:
notebook = rule[1]
cell_type = rule[2]
output_type = rule[3] # i.e. stream or error
output_type_name = rule[4] # i.e. ename or name
output_type_value = rule[5] # i.e. SystemExit or stdout
details_name = rule[6] # i.e. evalue or text
expression = rule[7].replace("\\*", "*") # Something escaped *, and put a \ in front of it!
if debug_logging:
print(f"EXPERT: If rule '{expression}' satisfied', run '{notebook}'.")
if re.match(expression, line, re.DOTALL):
if debug_logging:
print("EXPERT: MATCH: name = value: '{0}' = '{1}' matched expression '{2}', therefore HINT '{4}'".format(output_type_name, output_type_value, expression, notebook))
match_found = True
display(Markdown(f'HINT: Use [{notebook}]({notebook}) to resolve this issue.'))
print('Common functions defined successfully.')
# Hints for binary (transient fault) retry, (known) error and install guide
#
retry_hints = {'kubectl': ['A connection attempt failed because the connected party did not properly respond after a period of time, or established connection failed because connected host has failed to respond']}
error_hints = {'kubectl': [['no such host', 'TSG010 - Get configuration contexts', '../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb'], ['no such host', 'TSG011 - Restart sparkhistory server', '../repair/tsg011-restart-sparkhistory-server.ipynb'], ['No connection could be made because the target machine actively refused it', 'TSG056 - Kubectl fails with No connection could be made because the target machine actively refused it', '../repair/tsg056-kubectl-no-connection-could-be-made.ipynb']]}
install_hint = {'kubectl': ['SOP036 - Install kubectl command line interface', '../install/sop036-install-kubectl.ipynb']}
```
### Get the Kubernetes namespace for the big data cluster
Get the namespace of the Big Data Cluster use the kubectl command line
interface .
**NOTE:**
If there is more than one Big Data Cluster in the target Kubernetes
cluster, then either:
- set \[0\] to the correct value for the big data cluster.
- set the environment variable AZDATA\_NAMESPACE, before starting
Azure Data Studio.
```
# Place Kubernetes namespace name for BDC into 'namespace' variable
if "AZDATA_NAMESPACE" in os.environ:
namespace = os.environ["AZDATA_NAMESPACE"]
else:
try:
namespace = run(f'kubectl get namespace --selector=MSSQL_CLUSTER -o jsonpath={{.items[0].metadata.name}}', return_output=True)
except:
from IPython.display import Markdown
print(f"ERROR: Unable to find a Kubernetes namespace with label 'MSSQL_CLUSTER'. SQL Server Big Data Cluster Kubernetes namespaces contain the label 'MSSQL_CLUSTER'.")
display(Markdown(f'HINT: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'HINT: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print(f'The SQL Server Big Data Cluster Kubernetes namespace is: {namespace}')
```
### Create a temporary directory to stage files
```
# Create a temporary directory to hold configuration files
import tempfile
temp_dir = tempfile.mkdtemp()
print(f"Temporary directory created: {temp_dir}")
```
### Helper function to save configuration files to disk
```
# Define helper function 'save_file' to save configuration files to the temporary directory created above
import os
import io
def save_file(filename, contents):
with io.open(os.path.join(temp_dir, filename), "w", encoding='utf8', newline='\n') as text_file:
text_file.write(contents)
print("File saved: " + os.path.join(temp_dir, filename))
print("Function `save_file` defined successfully.")
```
### Get name of the ‘Running’ `controller` `pod`
```
# Place the name of the 'Running' controller pod in variable `controller`
controller = run(f'kubectl get pod --selector=app=controller -n {namespace} -o jsonpath={{.items[0].metadata.name}} --field-selector=status.phase=Running', return_output=True)
print(f"Controller pod name: {controller}")
```
### Get the name of the `management proxy` `pod`
```
# Place the name of the mgmtproxy pod in variable `pod`
pod = run(f'kubectl get pod --selector=app=mgmtproxy -n {namespace} -o jsonpath={{.items[0].metadata.name}}', return_output=True)
print(f"Management proxy pod name: {pod}")
```
### Copy certifcate files from `controller` to local machine
```
import os
cwd = os.getcwd()
os.chdir(temp_dir) # Use chdir to workaround kubectl bug on Windows, which incorrectly processes 'c:\' on kubectl cp cmd line
run(f'kubectl cp {controller}:{test_cert_store_root}/{app_name}/{prefix_keyfile_name}-certificate.pem {prefix_keyfile_name}-certificate.pem -c controller -n {namespace}')
run(f'kubectl cp {controller}:{test_cert_store_root}/{app_name}/{prefix_keyfile_name}-privatekey.pem {prefix_keyfile_name}-privatekey.pem -c controller -n {namespace}')
os.chdir(cwd)
```
### Copy certifcate files from local machine to `controldb`
```
import os
cwd = os.getcwd()
os.chdir(temp_dir) # Workaround kubectl bug on Windows, can't put c:\ on kubectl cp cmd line
run(f'kubectl cp {prefix_keyfile_name}-certificate.pem controldb-0:/var/opt/mssql/{prefix_keyfile_name}-certificate.pem -c mssql-server -n {namespace}')
run(f'kubectl cp {prefix_keyfile_name}-privatekey.pem controldb-0:/var/opt/mssql/{prefix_keyfile_name}-privatekey.pem -c mssql-server -n {namespace}')
os.chdir(cwd)
```
### Get the `controller-db-rw-secret` secret
Get the controller SQL symmetric key password for decryption.
```
import base64
controller_db_rw_secret = run(f'kubectl get secret/controller-db-rw-secret -n {namespace} -o jsonpath={{.data.encryptionPassword}}', return_output=True)
controller_db_rw_secret = base64.b64decode(controller_db_rw_secret).decode('utf-8')
print("controller_db_rw_secret retrieved")
```
### Update the files table with the certificates through opened SQL connection
```
import os
sql = f"""
OPEN SYMMETRIC KEY ControllerDbSymmetricKey DECRYPTION BY PASSWORD = '{controller_db_rw_secret}'
DECLARE @FileData VARBINARY(MAX), @Key uniqueidentifier;
SELECT @Key = KEY_GUID('ControllerDbSymmetricKey');
SELECT TOP 1 @FileData = doc.BulkColumn FROM OPENROWSET(BULK N'/var/opt/mssql/{prefix_keyfile_name}-certificate.pem', SINGLE_BLOB) AS doc;
EXEC [dbo].[sp_set_file_data_encrypted] @FilePath = '/config/scaledsets/{scaledset_name}/containers/{container_name}/files/{prefix_keyfile_name}-certificate.pem',
@Data = @FileData,
@KeyGuid = @Key,
@Version = '0';
SELECT TOP 1 @FileData = doc.BulkColumn FROM OPENROWSET(BULK N'/var/opt/mssql/{prefix_keyfile_name}-privatekey.pem', SINGLE_BLOB) AS doc;
EXEC [dbo].[sp_set_file_data_encrypted] @FilePath = '/config/scaledsets/{scaledset_name}/containers/{container_name}/files/{prefix_keyfile_name}-privatekey.pem',
@Data = @FileData,
@KeyGuid = @Key,
@Version = '0';
"""
save_file("insert_certificates.sql", sql)
cwd = os.getcwd()
os.chdir(temp_dir) # Workaround kubectl bug on Windows, can't put c:\ on kubectl cp cmd line
run(f'kubectl cp insert_certificates.sql controldb-0:/var/opt/mssql/insert_certificates.sql -c mssql-server -n {namespace}')
run(f"""kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c "SQLCMDPASSWORD=`cat /var/run/secrets/credentials/mssql-sa-password/password` /opt/mssql-tools/bin/sqlcmd -b -U sa -d controller -i /var/opt/mssql/insert_certificates.sql" """)
# Clean up
run(f"""kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c "rm /var/opt/mssql/insert_certificates.sql" """)
run(f"""kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c "rm /var/opt/mssql/{prefix_keyfile_name}-certificate.pem" """)
run(f"""kubectl exec controldb-0 -c mssql-server -n {namespace} -- bash -c "rm /var/opt/mssql/{prefix_keyfile_name}-privatekey.pem" """)
os.chdir(cwd)
```
### Clear out the controller\_db\_rw\_secret variable
```
controller_db_rw_secret= ""
```
### Clean up certificate staging area
Remove the certificate files generated on disk (they have now been
placed in the controller database).
```
cmd = f"rm -r {test_cert_store_root}/{app_name}"
run(f'kubectl exec {controller} -c controller -n {namespace} -- bash -c "{cmd}"')
```
### Restart Pod
```
run(f'kubectl delete pod {pod} -n {namespace}')
```
### Clean up temporary directory for staging configuration files
```
# Delete the temporary directory used to hold configuration files
import shutil
shutil.rmtree(temp_dir)
print(f'Temporary directory deleted: {temp_dir}')
print('Notebook execution complete.')
```
Related
-------
- [CER041 - Install signed Knox
certificate](../cert-management/cer041-install-knox-cert.ipynb)
- [CER030 - Sign Management Proxy certificate with generated
CA](../cert-management/cer030-sign-service-proxy-generated-cert.ipynb)
- [CER020 - Create Management Proxy
certificate](../cert-management/cer020-create-management-service-proxy-cert.ipynb)
| github_jupyter |
```
#default_exp callback.training_utils
```
# Training Utility Callbacks
> Very basic Callbacks to enhance the training experience including CUDA support
```
#export
# Contains code used/modified by fastai_minima author from fastai
# Copyright 2019 the fast.ai team.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language
#hide
from nbdev.showdoc import *
from fastcore.test import *
#export
from fastai_minima.callback.core import Callback
from fastai_minima.learner import Learner
from fastai_minima.utils import defaults, noop, default_device, to_device
from fastprogress.fastprogress import progress_bar,master_bar
from fastcore.basics import patch, ifnone
from contextlib import contextmanager
#export
class ProgressCallback(Callback):
"A `Callback` to handle the display of progress bars"
order,_stateattrs = 60,('mbar','pbar')
def before_fit(self):
"Setup the master bar over the epochs"
assert hasattr(self.learn, 'recorder')
if self.create_mbar: self.mbar = master_bar(list(range(self.n_epoch)))
if self.learn.logger != noop:
self.old_logger,self.learn.logger = self.logger,self._write_stats
self._write_stats(self.recorder.metric_names)
else: self.old_logger = noop
def before_epoch(self):
"Update the master bar"
if getattr(self, 'mbar', False): self.mbar.update(self.epoch)
def before_train(self):
"Launch a progress bar over the training dataloader"
self._launch_pbar()
def before_validate(self):
"Launch a progress bar over the validation dataloader"
self._launch_pbar()
def after_train(self):
"Close the progress bar over the training dataloader"
self.pbar.on_iter_end()
def after_validate(self):
"Close the progress bar over the validation dataloader"
self.pbar.on_iter_end()
def after_batch(self):
"Update the current progress bar"
self.pbar.update(self.iter+1)
if hasattr(self, 'smooth_loss'): self.pbar.comment = f'{self.smooth_loss:.4f}'
def _launch_pbar(self):
self.pbar = progress_bar(self.dl, parent=getattr(self, 'mbar', None), leave=False)
self.pbar.update(0)
def after_fit(self):
"Close the master bar"
if getattr(self, 'mbar', False):
self.mbar.on_iter_end()
delattr(self, 'mbar')
if hasattr(self, 'old_logger'): self.learn.logger = self.old_logger
def _write_stats(self, log):
if getattr(self, 'mbar', False): self.mbar.write([f'{l:.6f}' if isinstance(l, float) else str(l) for l in log], table=True)
if not hasattr(defaults, 'callbacks'): defaults.callbacks = [TrainEvalCallback, Recorder, ProgressCallback]
elif ProgressCallback not in defaults.callbacks: defaults.callbacks.append(ProgressCallback)
#hide
import torch
from torch.utils.data import TensorDataset, DataLoader
from fastai_minima.learner import DataLoaders
from torch import nn
def synth_dbunch(a=2, b=3, bs=16, n_train=10, n_valid=2):
"A simple dataset where `x` is random and `y = a*x + b` plus some noise."
def get_data(n):
x = torch.randn(int(bs*n))
return TensorDataset(x, a*x + b + 0.1*torch.randn(int(bs*n)))
train_ds = get_data(n_train)
valid_ds = get_data(n_valid)
train_dl = DataLoader(train_ds, batch_size=bs, shuffle=True, num_workers=0)
valid_dl = DataLoader(valid_ds, batch_size=bs, num_workers=0)
return DataLoaders(train_dl, valid_dl)
def synth_learner(n_train=10, n_valid=2, lr=defaults.lr, **kwargs):
data = synth_dbunch(n_train=n_train,n_valid=n_valid)
return Learner(data, RegModel(), loss_func=nn.MSELoss(), lr=lr, **kwargs)
class RegModel(nn.Module):
"A r"
def __init__(self):
super().__init__()
self.a,self.b = nn.Parameter(torch.randn(1)),nn.Parameter(torch.randn(1))
def forward(self, x): return x*self.a + self.b
learn = synth_learner()
learn.fit(5)
#export
@patch
@contextmanager
def no_bar(self:Learner):
"Context manager that deactivates the use of progress bars"
has_progress = hasattr(self, 'progress')
if has_progress: self.remove_cb(self.progress)
try: yield self
finally:
if has_progress: self.add_cb(ProgressCallback())
learn = synth_learner()
with learn.no_bar(): learn.fit(5)
#hide
#Check validate works without any training
import torch.nn.functional as F
def tst_metric(out, targ): return F.mse_loss(out, targ)
learn = synth_learner(metrics=tst_metric)
preds,targs = learn.validate()
show_doc(ProgressCallback.before_fit)
show_doc(ProgressCallback.before_epoch)
show_doc(ProgressCallback.before_train)
show_doc(ProgressCallback.before_validate)
show_doc(ProgressCallback.after_batch)
show_doc(ProgressCallback.after_train)
show_doc(ProgressCallback.after_validate)
show_doc(ProgressCallback.after_fit)
#export
class CollectDataCallback(Callback):
"Collect all batches, along with `pred` and `loss`, into `self.data`. Mainly for testing"
def before_fit(self): self.data = L()
def after_batch(self):
self.data.append(self.learn.to_detach((self.xb,self.yb,self.pred,self.loss)))
# export
class CudaCallback(Callback):
"Move data to CUDA device"
def __init__(self, device=None): self.device = ifnone(device, default_device())
def before_batch(self): self.learn.xb,self.learn.yb = to_device(self.xb),to_device(self.yb)
def before_fit(self): self.model.to(self.device)
```
| github_jupyter |
This notebook takes the associated primary contigs and haplotigs contigs and performs specific alignments for those
This notebook was only designed for the purpose of analyzing the Pst-104E genome. No gurantees it works in any other situtation. It will have spelling errors due to the lack of autocorrection.
This is it uses the FLACON unzip h contigs of each p contig and aligns them with the following settings
> ##For Assembletics e.g.
/home/benjamin/anaconda3/bin//nucmer -maxmatch -l 100 -c 500 Pst_104E_v12_pcontig_193.fa Pst_104E_v12_pcontig_193_h_ctgs.fa -prefix Pst_104E_v12_pcontig_193_php
with
##Assemblytics delta_file <output_prefix unique_anchor_length maximum_feature_length path_to_R_scripts Assemblytics Pst_104E_v12_pcontig_166_php.delta Pst_104E_v12_pcontig_166_php_8kbp_50kp 8000 50000 /home/benjamin/genome_assembly/PST79/FALCON/p_assemblies/v9_1/downstream_analysis_2017/scripts/Assemblytics
For Mummerplot
```
import os
from Bio import SeqIO
import shutil
import subprocess
#Define the PATH for variable parameters
BASE_AA_PATH = '/home/benjamin/genome_assembly/PST79/FALCON/p_assemblies/v9_1/Pst_104E_v12'
BASE_A_PATH = '/home/benjamin/genome_assembly/PST79/FALCON/p_assemblies/v9_1/032017_assembly'
OUT_PATH_NUCMER = os.path.join(BASE_AA_PATH, 'nucmer_analysis')
OUT_PATH_ASSEMBLETICS = os.path.join(BASE_AA_PATH, 'Assembletics')
MUMMER_PATH_PREFIX = '/home/benjamin/anaconda3/bin/'
ASSEMBLETICS_PATH = '/home/benjamin/genome_assembly/PST79/FALCON/p_assemblies/v9_1/downstream_analysis_2017/scripts/Assemblytics'
py2_env = 'py27' #python 2 environment in conda
if not os.path.isdir(OUT_PATH_NUCMER):
os.mkdir(OUT_PATH_NUCMER)
if not os.path.isdir(OUT_PATH_ASSEMBLETICS):
os.mkdir(OUT_PATH_ASSEMBLETICS)
#Define your p and h genome and move it into the allele analysis folder
genome_prefix = 'Pst_104E_v12'
p_genome = 'Pst_104E_v12_p_ctg'
h_genome = 'Pst_104E_v12_h_ctg'
genome_file_suffix = '.genome_file'
for x in (x + '.fa' for x in [p_genome, h_genome]):
shutil.copy2(BASE_A_PATH+'/'+x, OUT_PATH_NUCMER)
shutil.copy2(BASE_A_PATH+'/'+x, OUT_PATH_ASSEMBLETICS)
#define the scripts to generate
bash_script_q= genome_prefix+"_ph_ctg_qmapping.sh"
bash_script_g=genome_prefix+"_ph_ctg_gmapping.sh"
bash_script_nucmer_assemlytics = genome_prefix +"_ncumer_assembleticsmapping.sh"
bash_script_assemlytics = genome_prefix + '_assembletics.sh'
outfq = open(os.path.join(OUT_PATH_NUCMER, bash_script_q), 'w')
outfq.write('#!/bin/bash\n')
outfg = open(os.path.join(OUT_PATH_NUCMER,bash_script_g), 'w')
outfg.write('#!/bin/bash\n')#parsing out P and corresponding h contigs and writing a short ncumer script that aligns them against each other
outfna = open(os.path.join(OUT_PATH_ASSEMBLETICS,bash_script_nucmer_assemlytics), 'w')
outfna.write('#!/bin/bash\n')
for pseq_record in SeqIO.parse(OUT_PATH_ASSEMBLETICS+'/'+genome_prefix+'_p_ctg.fa', 'fasta'):
p_acontigs = []
p_contig = pseq_record.id.split("_")[0]+"_"+pseq_record.id.split("_")[1]
suffix = genome_prefix+"_"+p_contig+"_php"
p_file = genome_prefix+"_"+p_contig+'.fa'
SeqIO.write(pseq_record, OUT_PATH_ASSEMBLETICS+'/'+ p_file, 'fasta')
SeqIO.write(pseq_record,OUT_PATH_NUCMER+'/'+ p_file, 'fasta')
for aseq_record in SeqIO.parse(OUT_PATH_ASSEMBLETICS+'/'+genome_prefix+'_h_ctg.fa', 'fasta'):
if aseq_record.id.split("_")[1] == pseq_record.id.split("_")[1]:
p_acontigs.append(aseq_record)
a_file = genome_prefix +"_"+pseq_record.id.split("_")[0]+"_"+pseq_record.id.split("_")[1]+'_h_ctgs.fa'
#if we have alternative contigs safe those too
if p_acontigs != []:
SeqIO.write(p_acontigs, OUT_PATH_ASSEMBLETICS+'/'+ a_file, 'fasta')
SeqIO.write(p_acontigs, OUT_PATH_NUCMER+'/'+ a_file, 'fasta')
outfq.write(MUMMER_PATH_PREFIX +'/nucmer '+p_file+' '+a_file+" > "+'out.delta\n')
outfq.write(MUMMER_PATH_PREFIX +'/delta-filter -q '+'out.delta'+" > "+suffix+"_qfiltered.delta\n")
outfq.write(MUMMER_PATH_PREFIX +'/show-coords -T '+suffix+"_qfiltered.delta > "+suffix+".qcoords\n")
outfq.write(MUMMER_PATH_PREFIX +'/mummerplot -p '+suffix+'_qfiltered --png '+suffix+"_qfiltered.delta\n")
outfq.write(MUMMER_PATH_PREFIX +'/mummerplot -c -p '+suffix+'_qfiltered_cov --png '+suffix+"_qfiltered.delta\n")
#for g_file bash script
outfg.write(MUMMER_PATH_PREFIX +'/nucmer '+p_file+' '+a_file+" > "+'out.delta\n')
outfg.write(MUMMER_PATH_PREFIX +'/delta-filter -g '+'out.delta'+" > "+suffix+"_gfiltered.delta\n")
outfg.write(MUMMER_PATH_PREFIX +'/show-coords -T '+suffix+"_gfiltered.delta > "+suffix+".gcoords\n")
outfg.write(MUMMER_PATH_PREFIX +'/mummerplot -p '+suffix+'_gfiltered --png '+suffix+"_gfiltered.delta\n")
outfg.write(MUMMER_PATH_PREFIX +'/mummerplot -c -p '+suffix+'_gfiltered_cov --png '+suffix+"_gfiltered.delta\n")
#for nucmer assemblytics out
outfna.write(MUMMER_PATH_PREFIX +'/nucmer -maxmatch -l 100 -c 500 '+p_file+' '+a_file+' -prefix ' + suffix +'\n')
outfna.close()
outfq.close()
outfg.close()
#run the scripts and check if they are errors
bash_script_q_stderr = subprocess.check_output('bash %s' %os.path.join(OUT_PATH_NUCMER, bash_script_q), shell=True, stderr=subprocess.STDOUT)
bash_script_g_stderr = subprocess.check_output('bash %s' %os.path.join(OUT_PATH_NUCMER, bash_script_g), shell=True, stdeorr=subprocess.STDOUT)
bash_script_assembletics_stderr = subprocess.check_output('bash %s' %os.path.join(OUT_PATH_ASSEMBLETICS, bash_script_nucmer_assemlytics), shell=True, stderr=subprocess.STDOUT)
bash_script_assembletics_stderr = subprocess.check_output('bash %s' %os.path.join(OUT_PATH_ASSEMBLETICS, bash_script_nucmer_assemlytics), shell=True, stderr=subprocess.STDOUT)
#write the Assemblytics script
outfnarun = open(os.path.join(OUT_PATH_ASSEMBLETICS, bash_script_assemlytics), 'w')
outfnarun.write('#!/bin/bash\n')
delta_files = [x for x in os.listdir(OUT_PATH_ASSEMBLETICS) if x.endswith('delta')]
outfnarun.write('export PATH="%s":$PATH\n'% ASSEMBLETICS_PATH)
outfnarun.write('#Assemblytics delta_file output_prefix unique_anchor_length maximum_feature_length path_to_R_scripts\n')
outfnarun.write('source activate %s\n' %py2_env)
for delta in delta_files:
folder_name = delta[:-6] + '_8kbp'
output_prefix = delta[:-6] + '_8kbp_50kp'
outfnarun.write("mkdir %s\ncp %s %s\ncd %s\n" % (folder_name, delta, folder_name, folder_name))
outfnarun.write("Assemblytics %s %s 8000 50000 %s\n" % (delta, output_prefix, ASSEMBLETICS_PATH) )
output_prefix = delta[:-6] + '_8kbp_10kp'
outfnarun.write("Assemblytics %s %s 8000 10000 %s\ncd ..\n" % (delta, output_prefix, ASSEMBLETICS_PATH) )
outfnarun.write('source deactivate\n')
outfnarun.close()
bash_script_assemlytics_stderr = subprocess.check_output('bash %s' %os.path.join(OUT_PATH_ASSEMBLETICS, bash_script_assemlytics), shell=True, stderr=subprocess.STDOUT)
```
| github_jupyter |
# ERA5 Data
Data download was requested from
https://cds.climate.copernicus.eu/cdsapp#!/dataset/reanalysis-era5-single-levels?tab=form
*10m u-component of wind (m s-1)*: This parameter is the eastward component of the 10m wind. It is the horizontal speed of air moving towards the east, at a height of ten metres above the surface of the Earth, in metres per second. Care should be taken when comparing this parameter with observations, because wind observations vary on small space and time scales and are affected by the local terrain, vegetation and buildings that are represented only on average in the ECMWF Integrated Forecasting System (IFS). This parameter can be combined with the V component of 10m wind to give the speed and direction of the horizontal 10m wind.
*10m v-component of wind (m s-1)*: This parameter is the northward component of the 10m wind. It is the horizontal speed of air moving towards the north, at a height of ten metres above the surface of the Earth, in metres per second. Care should be taken when comparing this parameter with observations, because wind observations vary on small space and time scales and are affected by the local terrain, vegetation and buildings that are represented only on average in the ECMWF Integrated Forecasting System (IFS). This parameter can be combined with the U component of 10m wind to give the speed and direction of the horizontal 10m wind.
*Evaporation (m of water equivalent)*: This parameter is the accumulated amount of water that has evaporated from the Earth's surface, including a simplified representation of transpiration (from vegetation), into vapour in the air above. This parameter is accumulated over a particular time period which depends on the data extracted. The ECMWF Integrated Forecasting System (IFS) convention is that downward fluxes are positive. Therefore, negative values indicate evaporation and positive values indicate condensation.
*Sea surface temperature (K)*: This parameter is the temperature of sea water near the surface. This parameter is taken from various providers, who process the observational data in different ways. Each provider uses data from several different observational sources. For example, satellites measure sea surface temperature (SST) in a layer a few microns thick in the uppermost mm of the ocean, drifting buoys measure SST at a depth of about 0.2-1.5m, whereas ships sample sea water down to about 10m, while the vessel is underway. Deeper measurements are not affected by changes that occur during a day, due to the rising and setting of the Sun (diurnal variations). Sometimes this parameter is taken from a forecast made by coupling the NEMO ocean model to the ECMWF Integrated Forecasting System (IFS). In this case, the SST is the average temperature of the uppermost metre of the ocean and does exhibit diurnal variations. This parameter has units of kelvin (K). Temperature measured in kelvin can be converted to degrees Celsius (°C) by subtracting 273.15.
*Surface latent heat flux (J m-2)*: This parameter is the transfer of latent heat (resulting from water phase changes, such as evaporation or condensation) between the Earth's surface and the atmosphere through the effects of turbulent air motion. Evaporation from the Earth's surface represents a transfer of energy from the surface to the atmosphere. This parameter is accumulated over a particular time period which depends on the data extracted. The units are joules per square metre (J m-2 ). To convert to watts per square metre (W m-2 ), the accumulated values should be divided by the accumulation period expressed in seconds. The ECMWF convention for vertical fluxes is positive downwards.
*Surface net solar radiation (J m-2)*: This parameter is the amount of solar radiation (also known as shortwave radiation) that reaches a horizontal plane at the surface of the Earth (both direct and diffuse) minus the amount reflected by the Earth's surface (which is governed by the albedo). Radiation from the Sun (solar, or shortwave, radiation) is partly reflected back to space by clouds and particles in the atmosphere (aerosols) and some of it is absorbed. The remainder is incident on the Earth's surface, where some of it is reflected. This parameter is accumulated over a particular time period which depends on the data extracted. The units are joules per square metre (J m-2 ). To convert to watts per square metre (W m-2 ), the accumulated values should be divided by the accumulation period expressed in seconds. The ECMWF convention for vertical fluxes is positive downwards.
*Surface net thermal radiation (J m-2)*: Thermal radiation (also known as longwave or terrestrial radiation) refers to radiation emitted by the atmosphere, clouds and the surface of the Earth. This parameter is the difference between downward and upward thermal radiation at the surface of the Earth. It is the amount of radiation passing through a horizontal plane. The atmosphere and clouds emit thermal radiation in all directions, some of which reaches the surface as downward thermal radiation. The upward thermal radiation at the surface consists of thermal radiation emitted by the surface plus the fraction of downwards thermal radiation reflected upward by the surface. This parameter is accumulated over a particular time period which depends on the data extracted. The units are joules per square metre (J m-2 ). To convert to watts per square metre (W m-2 ), the accumulated values should be divided by the accumulation period expressed in seconds. The ECMWF convention for vertical fluxes is positive downwards.
*Surface sensible heat flux (J m-2)*: This parameter is the transfer of heat between the Earth's surface and the atmosphere through the effects of turbulent air motion (but excluding any heat transfer resulting from condensation or evaporation). The magnitude of the sensible heat flux is governed by the difference in temperature between the surface and the overlying atmosphere, wind speed and the surface roughness. For example, cold air overlying a warm surface would produce a sensible heat flux from the land (or ocean) into the atmosphere. This parameter is accumulated over a particular time period which depends on the data extracted. The units are joules per square metre (J m-2 ). To convert to watts per square metre (W m-2 ), the accumulated values should be divided by the accumulation period expressed in seconds. The ECMWF convention for vertical fluxes is positive downwards.
*Total precipitation (m)*: This parameter is the accumulated liquid and frozen water, comprising rain and snow, that falls to the Earth's surface. It is the sum of large-scale precipitation and convective precipitation. Large-scale precipitation is generated by the cloud scheme in the ECMWF Integrated Forecasting System (IFS). The cloud scheme represents the formation and dissipation of clouds and large-scale precipitation due to changes in atmospheric quantities (such as pressure, temperature and moisture) predicted directly by the IFS at spatial scales of the grid box or larger. Convective precipitation is generated by the convection scheme in the IFS, which represents convection at spatial scales smaller than the grid box. This parameter does not include fog, dew or the precipitation that evaporates in the atmosphere before it lands at the surface of the Earth. This parameter is the total amount of water accumulated over a particular time period which depends on the data extracted. The units of this parameter are depth in metres of water equivalent. It is the depth the water would have if it were spread evenly over the grid box. Care should be taken when comparing model parameters with observations, because observations are often local to a particular point in space and time, rather than representing averages over a model grid box.
```
import cdsapi
c = cdsapi.Client()
c.retrieve(
'reanalysis-era5-single-levels',
{
'product_type': 'reanalysis',
'format': 'netcdf',
'variable': [
'10m_u_component_of_wind', '10m_v_component_of_wind',
],
'year': '2019',
'month': ['01','02',
'03', '04', '05',
'06', '07', '08',
'09', '10', '11',
'12',
],
'day': [
'01', '02', '03',
'04', '05', '06',
'07', '08', '09',
'10', '11', '12',
'13', '14', '15',
'16', '17', '18',
'19', '20', '21',
'22', '23', '24',
'25', '26', '27',
'28', '29', '30',
'31',
],
'time': [
'00:00', '01:00', '02:00',
'03:00', '04:00', '05:00',
'06:00', '07:00', '08:00',
'09:00', '10:00', '11:00',
'12:00', '13:00', '14:00',
'15:00', '16:00', '17:00',
'18:00', '19:00', '20:00',
'21:00', '22:00', '23:00',
],
'area': '-45/25/-62/45'
},
'download_winds.nc')
c = cdsapi.Client()
c.retrieve(
'reanalysis-era5-single-levels',
{
'product_type': 'reanalysis',
'format': 'netcdf',
'variable': [
'sea_surface_temperature',
],
'year': '2019',
'month': ['01','02',
'03', '04', '05',
'06', '07', '08',
'09', '10', '11',
'12',
],
'day': [
'01', '02', '03',
'04', '05', '06',
'07', '08', '09',
'10', '11', '12',
'13', '14', '15',
'16', '17', '18',
'19', '20', '21',
'22', '23', '24',
'25', '26', '27',
'28', '29', '30',
'31',
],
'time': [
'00:00', '01:00', '02:00',
'03:00', '04:00', '05:00',
'06:00', '07:00', '08:00',
'09:00', '10:00', '11:00',
'12:00', '13:00', '14:00',
'15:00', '16:00', '17:00',
'18:00', '19:00', '20:00',
'21:00', '22:00', '23:00',
],
'area': '-45/25/-62/45'
},
'download_SST.nc')
c = cdsapi.Client()
c.retrieve(
'reanalysis-era5-single-levels',
{
'product_type': 'reanalysis',
'format': 'netcdf',
'variable': [
'evaporation', 'surface_latent_heat_flux', 'surface_net_solar_radiation',
'surface_net_thermal_radiation', 'surface_sensible_heat_flux', 'total_precipitation',
],
'year': '2019',
'month': ['01','02',
'03', '04', '05',
'06', '07', '08',
'09', '10', '11',
'12',
],
'day': [
'01', '02', '03',
'04', '05', '06',
'07', '08', '09',
'10', '11', '12',
'13', '14', '15',
'16', '17', '18',
'19', '20', '21',
'22', '23', '24',
'25', '26', '27',
'28', '29', '30',
'31',
],
'time': [
'00:00', '01:00', '02:00',
'03:00', '04:00', '05:00',
'06:00', '07:00', '08:00',
'09:00', '10:00', '11:00',
'12:00', '13:00', '14:00',
'15:00', '16:00', '17:00',
'18:00', '19:00', '20:00',
'21:00', '22:00', '23:00',
],
'area': '-45/25/-62/45'
},
'download_surf_flux.nc')
```
| github_jupyter |
# AgavePy
<img style="display:inline;vertical-align:center" src="python-logo-generic.svg" width="150px">
<span style="font-size:28pt">+</span>
<img style="display:inline;vertical-align:bottom" src="logo-white-lg.png" width="160px">
These are instructions and a basic example of how to run `agavepy`.
## Setting up
Fill the following variables to the values for your system. The default ones work against a sandbox.
```
USERNAME = 'jdoe'
PASSWORD = 'abcde'
```
This is the IP of a machine with `ssh` access, to work as an storage system:
```
STORAGE_IP = '191.236.150.54'
STORAGE_USERNAME = 'jstubbs'
# home directory in STORAGE_IP
HOME_DIR = '/home/jstubbs'
# credentials to access STORAGE_IP
STORAGE_PASSWORD = 'your password here'
```
Here are the parameters needed for an execution system. In this example we point it to the same IP as the storage system.
```
EXECUTION_IP = STORAGE_IP
EXECUTION_USERNAME = STORAGE_USERNAME
EXECUTION_PASSWORD = STORAGE_PASSWORD
```
## Creating the agave object
Import the `agavepy` module:
```
import agavepy.agave as a
import json
```
Instantiate with your username and password.
`api_server` points to an agave server. In the current container, `dev.agave.tacc.utexas.edu` is defined in `/etc/hosts` pointing to an Agave sandbox.
```
agave = a.Agave(api_server='https://dev.agave.tacc.utexas.edu',
username=USERNAME,
password=PASSWORD,
verify=False)
```
Usually there are already clients defined. If you created them with `agavepy`, you already have the credentials to use those clients in a file `~/.agavepy`, which is used automatically. Otherwise, you can create new ones.
List them with:
```
agave.clients.list()
```
You can create a new client, in which case `agavepy` gets automatically connected to it:
```
agave.clients.delete(clientName='myclient') # delete old client first (if any)
agave.clients.create(body={'clientName': 'myclient'})
```
The new client should appear now in the list. Let's check it:
```
[client for client in agave.clients.list() if client['name'] == 'myclient']
```
## Using the API
### Systems
At this point, the object `agave` is connected to the API using a client. As a test, let's list the systems associated to this client:
```
agave.systems.list()
```
This is the JSON object necessary to register a new storage system:
```
storage = {
"id": "my.test.system",
"name": "Agavepy test storage system",
"status": "UP",
"type": "STORAGE",
"description": "Storage system for agavepy test",
"site": "agaveapi.co",
"storage":{
"host": STORAGE_IP,
"port": 22,
"protocol": "SFTP",
"rootDir": "/",
"homeDir": HOME_DIR,
"auth":{
"username": STORAGE_USERNAME,
"type": "PASSWORD",
"password": STORAGE_PASSWORD
}
}
}
```
The actuall call to the Agave API is:
```
agave.systems.add(body=storage)
```
Let's check that the storage system is up and that we have access by listing files:
```
[f.name for f in agave.files.list(systemId='my.test.system', filePath=HOME_DIR)]
```
And let's be bold and add a file.
We create the file locally first:
```
import tempfile
temp = tempfile.mktemp()
with open(temp, 'w') as f:
f.write('Hello world!')
```
and we upload it with:
```
response = agave.files.importData(
systemId='my.test.system',
filePath=HOME_DIR,
fileName='hello.txt',
fileToUpload=open(temp))
response
```
Note the field `response['uuid']`, which is an identifier that can be used to refer to the file.
Let's try downloading the file:
```
agave.files.download(systemId='my.test.system', filePath='hello.txt').text
```
Now we register an execution system. This is the JSON object:
```
execution = {
"id": "my.execution.system",
"name": "Agavepy execution test system",
"status": "UP",
"type": "EXECUTION",
"description": "Execution system for agavepy test",
"site": "agaveapi.co",
"executionType": "CLI",
"scratchDir": HOME_DIR,
"workDir": HOME_DIR,
"queues": [
{
"name": "debug",
"maxJobs": 100,
"maxUserJobs": 10,
"maxNodes": 128,
"maxMemoryPerNode": "2GB",
"maxProcessorsPerNode": 128,
"maxRequestedTime": "24:00:00",
"customDirectives": "",
"default": True
}
],
"login":{
"host": EXECUTION_IP,
"port": 22,
"protocol": "SSH",
"scratchDir": HOME_DIR,
"workDir": HOME_DIR,
"auth":{
"username": EXECUTION_USERNAME,
"type":"PASSWORD",
"password": EXECUTION_PASSWORD
}
},
"storage":{
"host": STORAGE_IP,
"port": 22,
"protocol": "SFTP",
"rootDir": "/",
"homeDir": HOME_DIR,
"auth":{
"username": STORAGE_USERNAME,
"type":"PASSWORD",
"password": STORAGE_PASSWORD
}
},
"maxSystemJobs": 100,
"maxSystemJobsPerUser": 10,
"scheduler": "FORK",
"environment": "",
"startupScript": "./bashrc"
}
```
The execution system is registered in the same way as the storage:
```
agave.systems.add(body=execution)
```
### Apps
Let's create a simple app that counts words. The Python functions is:
```
%%writefile wc.py
import sys
def word_count(filename):
return len(open(filename).read().split())
if __name__ == '__main__':
with open('out.txt', 'w') as out:
out.write(str(word_count(sys.argv[1])))
```
We test it by creating a temporary file and passing its name:
```
temp = tempfile.mktemp()
with open(temp, 'w') as f:
f.write('Lorem ipsum dolor sit amet, consectetur adipiscing elit.')
import wc
print(wc.word_count(temp))
```
Let's wrap the Python function in a shell script:
```
%%writefile wc.sh
python2 wc.py ${temp}
```
And let's test it:
```
!chmod +x wc.sh # make shell script executable
!temp=$temp ./wc.sh # call wc.sh with env variable 'temp'
!cat out.txt # display output created by wc.sh
```
Here we upload the files `wc.py` and `wc.sh` to the storage system:
```
for filename in ['wc.py', 'wc.sh']:
agave.files.importData(
systemId='my.test.system',
filePath=HOME_DIR,
fileName=filename,
fileToUpload=open(filename))
```
Now we register an app with name `my.wc` using the previously registered storage and execution systems. The app takes an input with id `temp`, matching the name of the input used in `wc.sh` above.
The JSON object is:
```
app = {
"defaultNodes": 1,
"defaultMemoryPerNode": "2GB",
"datePublished": "",
"defaultQueue": "debug",
"author": "agavepy",
"shortDescription": "Word count app",
"label": "word-count",
"defaultProcessorsPerNode": 1,
"version": "0.0.1",
"defaultMaxRunTime": "01:00:00",
"available": True,
"inputs": [
{
"semantics": {
"fileTypes": [
"text-0"
],
"minCardinality": 1,
"ontology": [
"http://sswapmeet.sswap.info/util/TextDocument"
],
"maxCardinality": 1
},
"id": "temp",
"value": {
"default": "",
"required": True,
"enquote": False,
"visible": True,
"validator": "",
"order": 0
},
"details": {
"repeatArgument": False,
"showArgument": False,
"description": "",
"label": "A text file."
}
}
],
"tags": [
"containers",
"docker"
],
"outputs": [],
"longDescription": "",
"executionSystem": "my.execution.system",
"testPath": HOME_DIR,
"deploymentPath": HOME_DIR,
"templatePath": "wc.sh",
"deploymentSystem": "my.test.system",
"name": "my.wc",
"checkpointable": True,
"executionType": "CLI",
"publiclyAvailable": "false",
"parallelism": "SERIAL",
"helpURI": "https://example.com"
}
```
Registering the app is similar to registering the systems:
```
agave.apps.add(body=app)
```
We should see our new app in the list of apps:
```
[app.name for app in agave.apps.list()]
```
We can access all the information of the app by its name together with its version:
```
agave.apps.get(appId='my.wc-0.0.1')
```
### Jobs
Submit a job to exercise the app. We'll set as input the file `hello.txt` that we uploaded before:
```
job = {
"name": "My Word Count",
"appId": "my.wc-0.0.1",
"inputs": {
"temp": ["agave://my.test.system//{HOME_DIR}/hello.txt".format(HOME_DIR=HOME_DIR)]
},
"archive": True,
"notifications": [
{
"url": "http://requestb.in/p9fm7ap9?job_id=${JOB_ID}&status=${JOB_STATUS}",
"event": "*",
"persistent": True
}
]
}
my_job = agave.jobs.submit(body=job)
my_job
```
We can retrieve the status of the job with:
```
agave.jobs.getStatus(jobId=my_job['id'])
```
Let's wait until the job is finished:
```
import time
while agave.jobs.getStatus(jobId=my_job['id'])['status'] != 'FINISHED':
time.sleep(1)
print 'Done!'
```
Finally, we can retrieve the result that was left in the file `out.txt`:
```
agave.files.download(systemId='my.test.system', filePath=my_job['archivePath']+'/out.txt').text
```
That's precisely the word count of "`Hello World!`"!!
| github_jupyter |
# MobileNetV2で欅坂46とけやき坂46のメンバーの顔認識
```
import keras
from keras.applications.mobilenetv2 import MobileNetV2
from matplotlib import pyplot as plt
from sklearn.model_selection import train_test_split
from utils.utils import load_data
%matplotlib inline
```
### モデルと学習の設定
EPOCHを2000
テストとバリデーションのデータサイズはともに0.1(=全体の10%)を指定した。
```
# モデルの設定
NUMBER_OF_MEMBERS = 41 # 漢字とひらがな合わせたメンバー数
CLASSES = NUMBER_OF_MEMBERS + 1 # one hot表現は0から始まるため
LOG_DIR = './logs/1.2_0.8/' # LossとAccuracyのログ
# 学習の設定
EPOCHS = 2000
TEST_SIZE = 0.1
VALIDATION_SPLIT = 0.1
```
### データの読み込み
```
X, Y = load_data('/home/ishiyama/notebooks/keyakizaka_member_detection/image/mobilenet/')
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size=TEST_SIZE, shuffle=True)
```
### モデル構築
```
model = MobileNetV2(alpha=1.2, depth_multiplier=0.8, include_top=True, weights=None, classes=CLASSES)
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
callbacks = keras.callbacks.TensorBoard(log_dir=LOG_DIR)
```
### 学習させる
```
fit_result = model.fit(
x=X_train,
y=Y_train,
epochs=EPOCHS,
validation_split=VALIDATION_SPLIT,
verbose=2,
callbacks=[callbacks]
)
```
### LossとAccuracyのグラフを表示する
(参考)[MNISTでハイパーパラメータをいじってloss/accuracyグラフを見てみる](https://qiita.com/hiroyuki827/items/213146d551a6e2227810)
```
fig, (axL, axR) = plt.subplots(ncols=2, figsize=(16,5))
# loss
def plot_history_loss(fit):
# Plot the loss in the history
axL.plot(fit.history['loss'],label="loss for training")
axL.plot(fit.history['val_loss'],label="loss for validation")
axL.set_title('model loss')
axL.set_xlabel('epoch')
axL.set_ylabel('loss')
axL.legend(loc='upper right')
# acc
def plot_history_acc(fit):
# Plot the loss in the history
axR.plot(fit.history['acc'],label="accuracy for training")
axR.plot(fit.history['val_acc'],label="accuracy for validation")
axR.set_title('model accuracy')
axR.set_xlabel('epoch')
axR.set_ylabel('accuracy')
axR.legend(loc='upper right')
plot_history_loss(fit_result)
plot_history_acc(fit_result)
plt.show()
plt.close()
```
周期的に精度が下がる原因を調査する。
### テストデータで精度を確認する
```
test_result = model.evaluate(
x=X_test,
y=Y_test
)
print('loss for test:', test_result[0])
print('accuracy for test:', test_result[1])
```
### 今回学習したモデルを保存する
```
model.save('keyakizaka_member_detection_mobilenetv2.h5')
```
## 混同行列を作成する
```
from matplotlib import pyplot as plt
import pandas as pd
import seaborn as sns
%matplotlib inline
plt.rcParams['figure.figsize'] = (12, 10)
predicted = model.predict(X_test)
predicted_member_id = predicted.argmax(axis=1)
true_member_id = Y_test.argmax(axis=1)
predicted_result = pd.DataFrame(
data={
'y_true': true_member_id,
'y_pred': predicted_member_id,
'cnt': 1
}
)
confusion_matrix = pd.pivot_table(
data=predicted_result,
index=['y_pred'],
columns=['y_true'],
values='cnt',
aggfunc='count',
fill_value=0
)
sns.heatmap(confusion_matrix)
```
| github_jupyter |
```
from joblib import dump, load
import numpy as np
import cv2
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
from sklearn import metrics
import matplotlib.pyplot as plt
#set the directory for custom scripts
import sys
sys.path.append('/Users/macbook/Box/git_hub/Insight_Project_clean/scripts/')
#import custom scripts
import sql_con
from sql_con import df_from_query
import hsv_shift as hsv
#import the clustered ds swatches
hsv_knn_chroma = load('/Users/macbook/Box/git_hub/Insight_Project_clean/models/ds_h_chroma.joblib')
hsv_knn_neutral = load('/Users/macbook/Box/git_hub/Insight_Project_clean/models/ds_h_neutrals.joblib')
#custom function imports image and converts to hsv and 1-D pixel array
pixels = hsv.import_convert_pixelize('/Users/macbook/Box/insight_project_data/test_image/tucan2.jpg')
pixels
```
# convert the image to pixels and seperate out the neutrals from the chroma
```
#custom function takes the pixel array and splits it into chroma and neutrals and returns two dataframes
#takes longer with larger images
shifted_colors, shifted_neutrals = hsv.shift_h_split(pixels, .25, .25)
shifted_colors
```
# cluster the colors using k-means and return the values
```
X_pixels = shifted_colors[['h']]
from sklearn.cluster import KMeans
kmeans = KMeans(n_clusters=8, random_state=42, algorithm = 'full')
kmeans.fit(X_pixels)
image2show = kmeans.cluster_centers_[kmeans.labels_]
kmeans_df = pd.DataFrame(image2show, columns=['h'])
kmeans_df['label'] = kmeans.labels_
```
## Take the clustered colors and match them to the pigment KNN model
```
X = kmeans_df[['h']]
predict_colors = hsv_knn_chroma.predict(X)
colors2 = np.array(np.unique(predict_colors, return_counts=True)).T
colors2df = pd.DataFrame(colors2, columns = ['name', 'count'])
names = colors2df.sort_values(by=['count'], ascending = False)
names
```
## cluster the neutrals
```
X_pixels_n = shifted_neutrals[['h']]
from sklearn.cluster import KMeans
kmeans_n = KMeans(n_clusters=2, random_state=42, algorithm = 'full')
kmeans_n.fit(X_pixels_n)
image2show_n = kmeans_n.cluster_centers_[kmeans_n.labels_]
kmeans_df_n = pd.DataFrame(image2show_n, columns=['h'])
kmeans_df_n['label'] = kmeans_n.labels_
kmeans_df_n
```
# Cluster the neutrals to Knn model
```
X_n = kmeans_df_n[['h']]
predict_neutrals = hsv_knn_neutral.predict(X_n)
neutrals = np.array(np.unique(predict_neutrals, return_counts=True)).T
neutrals_df = pd.DataFrame(neutrals, columns = ['name', 'count'])
names_n = neutrals_df.sort_values(by=['count'], ascending = False)
names_n
```
| github_jupyter |
# 2. Linear Combination, Linear Transformation, Dot Product
As we progress in our understanding of the math surrounding machine learning, AI, and DS, there will be a host of linear algebra concepts that we are forced to reckon with. From PCA and it's utilization of eigenvalues and eigenvectors, to neural networks reliance on linear combinations and matrix multiplication, the list goes on and on. Having a very solid grasp on linear algebra is crucial to realizing _how_ and _why_ these algorithms work.
This notebook in particular is going to focus on the connection between the following:
* **Linear Combinations**
* **Linear Transformations**
* **The Dot Product**
* **Functions**
These concepts are incredibly prevelant and linked to each other in beautiful ways, however, this link is generally missing in the way linear algebra is taught-particularly when studying machine learning. Before moving, on I recommend reviewing my notebook concerning vectors.
# 1. Linear Combination
If you go to wikipedia, you can find the following definition regarding a **linear combination**:
> A linear combination is an expression constructed from a set of terms by multiplying each term by a constant and adding the results. For example, a linear combination of $x$ and $y$ would be any expression of the form $ax + by$, where a and b are constants.
Now, this can be defined slightly more formally in regards to vectors as follows:
If $v_1,...,v_n$ is a set of vectors, and $a_1,...,a_n$ is a set of scalars, then their linear combination would take the form:<br>
$$a_1\vec{v_1} + a_2\vec{v_2}+...+a_n\vec{v_n}$$
Where, it should be noted that all $\vec{v}$'s are vectors. Hence, it can be expanded as:<br>
<br>
$$a_1\begin{bmatrix}
v_1^1 \\
v_1^2 \\
. \\
. \\
v_1^m
\end{bmatrix}
+
a_2\begin{bmatrix}
v_2^1 \\
v_2^2 \\
. \\
. \\
v_2^m
\end{bmatrix}
+
a_n\begin{bmatrix}
v_n^1 \\
v_n^2 \\
. \\
. \\
v_n^m
\end{bmatrix}
$$
where for generality we have defined $\vec{v}$ to be an $m$ dimensional vector. Notice that the final result is a single $m$ dimensional vector. So, for instance, in a simple case, we could have:<br>
<br>
$$a_1\begin{bmatrix}
v_1^1
\end{bmatrix}
+
a_2\begin{bmatrix}
v_2^1
\end{bmatrix}
+
a_n\begin{bmatrix}
v_n^1
\end{bmatrix}
$$
<br>
$$a_1\begin{bmatrix}
v_1
\end{bmatrix}
+
a_2\begin{bmatrix}
v_2
\end{bmatrix}
+
a_n\begin{bmatrix}
v_n
\end{bmatrix}
$$<br>
$$a_1v_1+a_2v_2+a_nv_n$$
And end up with a 1 dimensional vector, often just viewed as a scalar.
Now, this definition is good to have in mind, however we can make it a bit more concrete by expanding visually. For instance, if you have a pair of numbers that is meant to describe, a vector, such as:
$$\begin{bmatrix}
3 \\
-2
\end{bmatrix}
$$
<img src="https://drive.google.com/uc?id=1dzL3c1wCklOE78qBiqf0QIbt70eQtsOD" width="300">
We can think of each coordinate as a scalar (how does it stretch or squish vectors?). In linear algebra, there are two very important vectors, commonly known as $\hat{i}$ and $\hat{j}$:
<img src="https://drive.google.com/uc?id=1M8_Uhcddg2bqET8yfrvsuzW6KqezsGgH" width="300">
Now, we can think of the coordinates of our vector as stretching $\hat{i}$ and $\hat{j}$:
<img src="https://drive.google.com/uc?id=1tAeBsXNFUdC_hV9z3RqahavR0KIhDGPt" width="300">
In this sense, the vector that these coordinates describe is the sum of two scaled vectors:
$$(3)\hat{i} + (-2)\hat{j}$$
Note that $\hat{i}$ and $\hat{j}$ have a special name; they are refered to as the _basis vectors_ of the _xy_ coordinate system. This means that when you think about vector coordinates as scalars, the basis vectors are what those coordinates are actually scaling.
Now, this brings us to our first definiton:
> **Linear Combination:** Any time you are scaling two vectors and then adding them together, you have a linear combination. For example: <br>
$$(3)\hat{i} + (-2)\hat{j}$$<br>
Or, more generally:
$$a\vec{v} + b \vec{w}$$<br>
Where, above both $a$ and $b$ are scalars.
This can be seen visually:
<img src="https://drive.google.com/uc?id=1Uzf5dAXN5sUOXM3VM7QtgyvD3yDXptBp" width="300">
And we can see that as we scale $\vec{v}$ and $\vec{w}$ we can create many different linear combinations:
<img src="https://drive.google.com/uc?id=1-ZS40KYmjMyWAVC1gm4GP2utTr2LyBYs" width="400">
This brings up another definition, _span_.
> **Span**: The set of all possible vectors that you can reach with a linear combination of a given pair of vectors is known as the _span_ of those two vectors.
So, the span of most 2-d vectors is all of space, however, if they line up then they span is a specific line. When two vectors do happen to line up we can say that they are _linearly dependent_, and one can be expressed in terms of the other. On the other hand, if they do line up, they are said to be _linearly independent_.
---
# 2. Linear Transformations and Matrices
Linear transformations are absolutely fundamental in order to understand matrix vector multiplication (well, unless you want to rely on memorization). To start, let's just parse the term "Linear Transformation".
Transformation is essentially just another way of saying _function_. This is where the first bit of confusion can arise though if you are being particularly thoughtful about the process's-what exactly is a function? It is helpful to define it before moving forward.
**Function**<br>
Generally, in mathematics we view a function as a process that take in an input and returns an output (this coincides nicely with the computer science view as well). It can be viewed as:
$$x \rightarrow f(x) \rightarrow y$$
Or, expanded as:
$$x_1, x_2, ... , x_n \rightarrow f(x_1, x_2, ... , x_n) \rightarrow y$$
This is how it is _generally_ encountered, where anywhere from one to several inputs are taken in, and a single output is produced. However, let's define a function more rigorously:
> A function is a _process_ or a relation that associates each element $x$ of a set $X$, the _domain_ of the function, to a single element $y$ of another set $Y$ (possibly the same set), the codomain of the function.
The important point to recognize from the above definition is that, while it is common for a function to map elements from a set $X$ to a different set $Y$, the two sets can be _same_. Hence, although it is not encountered quite as often in ML, a function can map $X \rightarrow X$.
Now, back to our term transformation; it is something that takes in inputs, and spits out an output for each one. In the context of linear algebra, we like to think about transformations that take in some vector, and spit out another vector:
$$\begin{bmatrix}
5 \\
7
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
2 \\
-3
\end{bmatrix}
$$
This is where we can see an example of a function that does not map to a different space necessarily, but potentially to itself. In other words, generally if we have a function that takes in two inputs, we end up with one output:
$$f(x,y) = z$$
However, we can clearly see here that we take in two inputs (coordinates of the vector) and end up with two outputs (coordinates of the transformed vector).
So, why use the word transformation instead of function if they essentially mean the same thing? Well, it is to be suggestive of _movement_! The way to think about functions of vectors is to use movement. If a transformation takes some input vector to some output vector, we image that input vector moving over to the output vector:
<img src="https://drive.google.com/uc?id=1JxUVZYtLwQh9eTK6RrRjGhfCP3Z_i5mh" width="400">
And in order to think about the transformation as a whole, we can think about _every possible input vector_ moving over to its corresponding _output vector_.
**Key Point**<br>
> We are now seeing that our general interpretation of a function can be expanded (via its full definition), to not simply taking in one or several inputs and producing a single output, but taking in one to several inputs, and producing one to several outputs. This expansion is key to recognizing the relationship between functions, linear transformations, and dot products.
Now let's pose the following question: If we were given the coordinates of a vector, and we then were trying to determine the coordinates of where that vector landed after being linearly transformed, how would we represent this?
$$\begin{bmatrix}
x_{in} \\
y_{in}
\end{bmatrix}
\rightarrow
????
\rightarrow
\begin{bmatrix}
x_{out} \\
y_{out}
\end{bmatrix}
$$
Well, it turns out that you actually only need to record where the two basis vectors land and everything else will follow from that! For example, let's consider the vector:
$$ \vec{v} =
\begin{bmatrix}
-1 \\
2
\end{bmatrix}$$
<img src="https://drive.google.com/uc?id=18WMVsEdWeDd6nt7XCxMkKIU_khRG73jo" width="250">
Where $\vec{v}$ can also be written as:
$$\vec{v} = -1 \hat{i} + 2 \hat{j}$$
If we then perform some transformation and watch where all of the vectors go, the property (of linear transformations) that all gridlines remain parallel and evenly spaced has a really important consequence!
<img src="https://drive.google.com/uc?id=1KJPU9eq8wAw1QBiM7rCZSOG2lj5Fwd9P" width="400">
The place where $\vec{v}$ lands will be -1 times the vector where $\hat{i}$ landed, and 2 times the vector where $\hat{j}$ landed.
$$\text{Transformed } \vec{v} = -1 (\text{Transformed }\hat{i}) + 2 (\text{Transformed }\hat{j})$$
In other words, it started off as a certain _linear combination_ of $\hat{i}$ and $\hat{j}$, and it ended up as that _same linear combination_ of where those two vectors landed! This means that we can determine where $\vec{v}$ must go, based only on where the two basis vectors land!
Because we have a copy of our original gridlines in the background, we can see where the vectors landed:
<img src="https://drive.google.com/uc?id=120YMveKLwGeS492LiMf0_RGJL9FThxyk" width="500">
We see that our transformed $\hat{i}$ and $\hat{j}$ landed at:
$$ \text{Transformed }\hat{i} =
\begin{bmatrix}
1 \\
-2
\end{bmatrix} \hspace{1cm} \hspace{1cm}
\text{Transformed }\hat{j} =
\begin{bmatrix}
3 \\
0
\end{bmatrix}
$$
Meaning that our transformed $\vec{v}$ is:
$$\text{Transformed } \vec{v} = -1 \begin{bmatrix}
1 \\
-2
\end{bmatrix} + 2 \begin{bmatrix}
3 \\
0
\end{bmatrix}$$
$$\text{Transformed } \vec{v} = \begin{bmatrix}
5 \\
2
\end{bmatrix}$$
Now, the cool thing about this is that we have just discovered a way to determine where _any_ transformed vector will land, only by knowing where $\hat{i}$ and $\hat{j}$ land, without needing to watch the transformation itself! We can write a vector with more general coordinates:
$$\begin{bmatrix}
x \\
y
\end{bmatrix}$$
And it will land on $x$ times the vector where $\hat{i}$ lands, and $y$ times the vector where $\hat{j}$ lands:
$$ \begin{bmatrix}
x \\
y
\end{bmatrix} \rightarrow x \begin{bmatrix}
1 \\
-2
\end{bmatrix} + y \begin{bmatrix}
3 \\
0
\end{bmatrix} =
\begin{bmatrix}
1x + 3y\\
-2x + 0y
\end{bmatrix}
$$
We can now get to a very key point:
This is all to say that a 2-dimensional linear transformation is completely described by _just four numbers_: the 2 coordinates for where $\hat{i}$ lands, and the 2 coordinates for where $\hat{j}$ lands. It is common to package these coordinates in a 2x2 grid of numbers, commonly referred to as a 2x2 _**matrix**_:<br>
<br>
$$\begin{bmatrix}
1 & 3\\
-2 & 0
\end{bmatrix}$$<br>
Here you can interprete the columns as where $\hat{i}$ lands and where $\hat{j}$ lands!
So, for just one more example, let's say we are given the 2x2 matrix representing a linear transformation:
$$
\begin{bmatrix}
3 & 2\\
-2 & 1
\end{bmatrix}$$
And we are then given a random vector:
$$\begin{bmatrix}
5\\
7
\end{bmatrix}$$
If we want to determine where that linear transformation takes that vector, we can take the coordinates of the vector, multiply them by the corresponding columns of the matrix, and then add together what you get:
$$
5\begin{bmatrix}
3 \\
-2
\end{bmatrix} +
7\begin{bmatrix}
2 \\
1
\end{bmatrix}$$
## 2.1 The General Case
Let's now try and generalize this as much as possible. Assume our 2x2 matrix (the typographical representation of our linear transformation of the vector space-in other words, it is a convenient way to package the information needed to describe a linear transformation), is:
$$
\begin{bmatrix}
a & b\\
c & d
\end{bmatrix}$$
If we apply this transformation to some vector:
$$ \begin{bmatrix}
x \\
y
\end{bmatrix}$$
What do we end up with? Well it will be:
$$
x\begin{bmatrix}
a \\
c
\end{bmatrix} +
y\begin{bmatrix}
b \\
d
\end{bmatrix} =
\begin{bmatrix}
ax + by\\
cx + dy
\end{bmatrix}$$
Now, what we just did-which has been intuitive and clear (or I should at least hope)-is frequently taught _with no intuition whatsoever_. It is referred to as _**matrix vector multiplication**_, and would written as follows:
$$\begin{bmatrix}
a & b\\
c & d
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix} =
x\begin{bmatrix}
a \\
c
\end{bmatrix} +
y\begin{bmatrix}
b \\
d
\end{bmatrix} =
\begin{bmatrix}
ax + by\\
cx + dy
\end{bmatrix}$$
Where our matrix (linear transformation, a form of a _function_) is on the left, and the vector is on the right. This, when taught without the necessary background is incredibly confusing and leaves us to simply utilize rote memorization of symbol manipulation processes. When we view this transformation as a type of function that takes in a vector as input and yields an output:
$$\begin{bmatrix}
5 \\
7
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
2 \\
-3
\end{bmatrix}
$$
It begins to become far more intuitive _why the order matters_, and _why the top number in the vector is multiplied by the first column of the matrix_, and subsequently _why the bottom number in the vector is multiplied by the second column in the matrix_. It is simply due to:
> 1. The way that vectors are, by convention, packaged and represented.
2. The way that matrices are, by convention, packaged and represented.
In order for the process to perform isomorphically to the geometric interpretation, we must follow the conventions that have been laid out. However, if we _only new the conventions_, while we did not have the underlying _intuition_, then there would be very little _meaning_ in what was going on.
## 2.2 Matrix Multiplication
We won't dig into it here, but as a quick note I wanted to touch on matrix multiplication. If we had the following two matrices:
$$\begin{bmatrix}
a & b\\
c & d
\end{bmatrix}$$
$$\begin{bmatrix}
e & f\\
g & h
\end{bmatrix}$$
And we wanted to the result of applying one, then the other, to a given vector:
$$\begin{bmatrix}
a & b\\
c & d
\end{bmatrix}
\begin{bmatrix}
e & f\\
g & h
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix}$$
We can first thinking about this a a _**composition of functions**_. What this would look like from a standard function perspective is:
$$f(g(x))$$
Where we first run $x$ through function $g$, and then the result is run through function $f$. In our case, where the function is a matrix (linear transformation), we can say that the vector is first _transformed_ via the matrix:
$$
\begin{bmatrix}
e & f\\
g & h
\end{bmatrix}$$
Which would look like:
$$
\begin{bmatrix}
e & f\\
g & h
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix} =
\begin{bmatrix}
ex + fy\\
gx + hy
\end{bmatrix}
$$
And that resulting 2-d vector is then transformed via the second _linear transformation_:
$$
\begin{bmatrix}
a & b\\
c & d
\end{bmatrix}
\begin{bmatrix}
ex + fy\\
gx + hy
\end{bmatrix} =
\begin{bmatrix}
a(ex + fy) + b(gx + hy)\\
c(ex + fy) + d(gx + hy)
\end{bmatrix}
$$
Resulting in a final 2-d vector! We can now easily understand what the following property of matrices exists:
$$M_1M_2 \neq M_2M_1$$
We know that because a matrix represents a _transformation_ or _function_, that switching the order changes the order in which they are applied. In a more familiar case, that would mean:
$$f(g(x)) \rightarrow g(f(x))$$
We know that the above is not true, and it follows that it is not true for matrices either!
---
# 3. Nonsquare Matrices $\rightarrow$ Transformations between dimensions
Up until this point we have been dealing with transformations from 2-d vectors to other 2-d vectors, by the means of a 2x2 matrix. Dealing with non-square matrices and having an underlying intuition is crucial, especially when dealing with machine learning (specifically deep neural networks). More importantly, it should help bring our generalization around to include some of the functions we are more familiar with.
As we go through this, recall what was mentioned at the start of this notebook. There are functions that are dependent on several inputs, such as:
$$f(x,y) = z$$
Or:
$$f(x_1,x_2, ...,x_n) = z$$
Now, what is interesting about functions of that sort is that we start with _multiple dimensions_, and the result of our function process is a _one dimensional output_. Keep that in mind as we move forward.
Okay, so we can start by saying that it is perfectly reasonable to talk about transformations between dimensions, such as:
$$\begin{bmatrix}
2 \\
7
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
1 \\
8 \\
2
\end{bmatrix}
$$
Above, we have a 2-d input, and a 3-d output. In order to determine the transformation $L$, we just look at the coordinates of where the basis vectors land! For instance, it may be (left column is $\hat{i}$, right column is $\hat{j}$):
$$L =
\begin{bmatrix}
2 & 0\\
-1 & 1 \\
-2 & 1
\end{bmatrix}$$
Now, the matrix above which encodes our transformation has 3 rows and 2 columns, making it a 3x2 matrix. We can intuitively say that a 3x2 matrix has a geometric interpretation of mapping 2 dimensions to 3 dimensions. This is because the 2 columns mean that the input space has 2 basis vectors, and the 3 rows indicate that the landing spots for each of those 3 basis vectors is described with 3 separate coordinates.
Likewise, if we see a 2x3 matrix (2 rows, 3 columns):
$$\begin{bmatrix}
3 & 1 & 4\\
1 & 5 & 9
\end{bmatrix}$$
We know that the 3 columns indicate that we are starting in a space that has 3 basis vectors, and the 2 rows indicate that the landing spot for each of those 3 basis vectors is described with only 2 coordinates.
Now, we can complete the bridge from linear transformations to our standard functions. We have come across functions before of the form:
$$f(x,y) = z$$
Where we take a 2 dimensional input, and produce a single 1 dimensional output. Well, linear transformations are capable of the same thing! Below, we take a 2-d input, and produce a 1-d output:
$$\begin{bmatrix}
2 \\
7
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
1.8
\end{bmatrix}
$$
1-dimensional space is just the number line, so a transformation such as the one above essentially just takes in 2-d vectors, and spits out a single number. A transformation such as this is encoded as a 1x2 matrix, where each column has a single entry:
$$L =
\begin{bmatrix}
1 & 2\\
\end{bmatrix}$$
The 2 columns represent where each of the basis vectors land, and each column requires just one number; the number where that basis vector landed on. This is a very interesting concept, with ties to the dot product.
---
# 4. The Dot Product
In general, the dot product would be introduced as follows: we have 2 vector of the same length, and we take there dot product by multiplying the corresponding rows and adding the results:
$$\begin{bmatrix}
2 \\
7 \\
1
\end{bmatrix} \cdot
\begin{bmatrix}
1 \\
8 \\
2
\end{bmatrix}
$$
$$(2*1) + (7*8) + (1*2) = 60$$
### 4.1 Geometric Interpretation
Now, this computation has a very nice geometric intepretation. Let's say we have two vectors, $\vec{v}$ and $\vec{w}$:
$$
\vec{v} =
\begin{bmatrix}
4 \\
1
\end{bmatrix}
\hspace{1cm}
\vec{w} =
\begin{bmatrix}
2 \\
-1
\end{bmatrix}
$$
<img src="https://drive.google.com/uc?id=1ain7m5PeTnwX-Wu5NepUCxNGxRwJR5xA" width="250">
To think about the dot product between $\vec{v}$ and $\vec{w}$, think about project $\vec{w}$ onto the line that passes through the origin and the tip of $\vec{v}$:
<img src="https://drive.google.com/uc?id=14wuptrxqsBeGVWFM_lu6JmC2Lw7LFcYr" width="250">
Multiplying the length of the projected $\vec{w}$ by the length of $\vec{v}$, we end up with $\vec{v} \cdot \vec{w}$:
$$
\vec{v} \cdot \vec{w} =
(\text{Length of projected } \vec{w})(\text{Length of }\vec{v})=
\begin{bmatrix}
4 \\
1
\end{bmatrix}
\cdot
\begin{bmatrix}
2 \\
-1
\end{bmatrix}
$$
Note: if the direction of the projection of $\vec{w}$ is pointing in the opposite direction of $\vec{v}$, that dot product should be _negative_.
So, when two vectors are generally pointing in the same direction their dot product is positive, when they are perpendicular their dot product is 0, and if they point in generally the opposite direction, their dot product is negative.
Note: Additionally I should mention that _order doesn't matter_. We will not going into the derivation here.
**Why are these two views connected?**<br>
You may be sitting there at this point wondering:
> "Why does this numerical process of matching coordinates, multiplying pairs, and adding them together, have anything to do with projection?"
Well, in order to give a fully satisfactory answer here, we are going to need to unearth something a little deeper here; _**duality**_. However, before we can get into that, we first need to talk about linear transformations from _multiple dimensions_ to _one dimension_.
### 4.2 Linear Transformations from Multiple to One Dimension
These are functions that take in a 2-d vector and spit out some number (a 1-d output):
$$\begin{bmatrix}
2 \\
7
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
1.8
\end{bmatrix}
$$
_Linear transformations_ are much more restrictive that your run of the mill function with a 2-d input with a 1-d output. For example, a _non-linear_ transformation (function) may look like:
$$f\Big( \begin{bmatrix}
x \\
y
\end{bmatrix} \Big) =
x^2 + y^2
$$
Now, recall from section 2 that we whenever we are performing a transformation, in order to determine the matrix that represents that transformation (remember: this operates isomorphically to a linear function), we just follow $\hat{i}$ and $\hat{j}$:
<img src="https://drive.google.com/uc?id=1oCORXDrbx5tgH1a0pfAej3AYfewTg5h7" width="250">
<img src="https://drive.google.com/uc?id=18vI0-5JaHen5X47qBV0NMeWRyhz4YiVD" width="250">
Only this time, each one of our basis vectors just lands on a number! When we record where they land as the columns of a matrix ($\hat{i}$ lands on 2 and $\hat{j}$ lands on 1), each of those columns just has a single number:
$$\text{Transformation Matrix:}\begin{bmatrix}
2 & 1
\end{bmatrix}
$$
This is a 1x2 matrix. Let's go through an example of what it means to apply one of these transformations to a vector. We will start by defining our vector, $\vec{v}$, to be:
$$\vec{v} =
\begin{bmatrix}
4 \\
3
\end{bmatrix}$$
<img src="https://drive.google.com/uc?id=1K4-RnkmUDUrbjbz3EBEVwYhug8s_htlZ" width="250">
Let's say that we have a linear transformation that takes $\hat{i}$ to 1, and $\hat{j}$ to -2:
<img src="https://drive.google.com/uc?id=1eNrwCAmXJhfbL4vVUI3nsn8s98Yn5N6g" width="250">
<img src="https://drive.google.com/uc?id=1VFoh9kgT4ZGcfao919SxSorLCRrieyLh" width="250">
Recall that we can write our original vector $\vec{v}$ as _linear combination_ of 4 times $\hat{i}$ plus 3 times $\hat{j}$:
$$(4)\hat{i} + (3)\hat{j}$$
A consequence of linearity is that when we perform a linear transformation on $\vec{v}$, the transformed output can be written as:
$$\text{Transformed } \vec{v} = 4 (\text{Transformed }\hat{i}) + 3 (\text{Transformed }\hat{j})$$
$$4(1) + 3(-2) = -2$$
When we perform this calculation purely numerically, it is matrix vector multiplication:
$$\begin{bmatrix}
1 & -2
\end{bmatrix}
\begin{bmatrix}
4 \\
3
\end{bmatrix} =
4*1 + 3*-2 = -2
$$
Where again, $\begin{bmatrix}1 & -2\end{bmatrix}$ is our transform, and $\begin{bmatrix}4 \\3\end{bmatrix}$ is our vector. Now, this numerical operation of multiplying a 1x2 matrix by a 2x1 vector feels _just like taking the dot product of two vectors_. The 1x2 matrix looks just like a vector that we tipped on its side. In fact, we can say that there is a nice association between 1x2 matrices and 2-d vectors:
$$\text{1 x 2 Matrices} \longleftrightarrow \text{2-d vectors}$$
At this point it is worth noting again that a matrix is a representation of a linear transformation, or function. What is often confusing in linear algebra is when matrices and vectors begin to look _the same_. For instance, we could have a matrice that is 2x1, looking identical to a vector. That means that it is expect 1 input dimension, and will result in with 2 output dimensions. It is important to remain aware of what typographical objects are meant to represent matrices and vectors as we continue.
Back to where we were, the above association suggests something very cool form the geometric view:
> _There is some kind of connections between linear transformations that take vectors to numbers, and vectors themselves_.
### 4.3 Example
We can discover more surrounding this connection with the following example. Let's take a copy of the number line and place it diagonaly in space somehow, with the number 0 sitting at the origin:
<img src="https://drive.google.com/uc?id=1X9gtfjtXSvtWCdvTX2x3xDhFil3usihH" width="300">
Now, think of the 2-dimensional unit vector whose tip sits where the number 1 on the number line is.
<img src="https://drive.google.com/uc?id=1hFUoKsnlH6xXUx3ZR2vNNbnmBQI4Q8JR" width="300">
We will call that vector $\hat{u}$. This vector plays a large role in what is about to happen. If we project 2 vectors straight onto this number line:
<img src="https://drive.google.com/uc?id=1PMBbMedbuwGtfmZGZv3bwONGVdJYq5ua" width="400">
<img src="https://drive.google.com/uc?id=1E9vfzNvDZsVbZeFjN1FDE3HL6ni5qKBM" width="400">
<img src="https://drive.google.com/uc?id=1Q-_lUtkLPB3O_BTYA3uMJ0mppYt-CKva" width="400">
In effect, _**we have just defined a function that takes 2-d vectors to numbers**_. What is more, this function is actually linear, since it passes the visual test _any line of evenly spaced dots must remain evenly spaced once landing on number line._
To be clear, even though the number line was embeded in 2-d space, the outputs are _numbers_, not _2-d vectors_. Again, we should be thinking of a function that takes in two coordinates, and outputs a single coordinate:
$$\begin{bmatrix}
x \\
y
\end{bmatrix} \rightarrow
L(\vec{v})
\rightarrow
\begin{bmatrix}
z
\end{bmatrix}
$$
But, the vector $\hat{u}$ is a two dimensional vector living in the input space. It is just situated in such a way that overlaps with the embedding of the number line.
With this projection, we just defined a linear transformation from 2-d vectors, to numbers. This means that we will be able to find a 1x2 matrix that describes this transformation:
$$\begin{bmatrix}
\text{Where } \hat{i} \text{ lands} & \text{Where } \hat{j} \text{ lands}
\end{bmatrix}$$
To find this 1x2 matrix, we can zoom in on our diagonal number line setup and think about where $\hat{i}$ and $\hat{j}$ each land, since those landing spots are going to be the columns of the matrix:
<img src="https://drive.google.com/uc?id=1cQMF-LalYiPdkJLEdV_DgD3wLT7DDjjP" width="230">
<img src="https://drive.google.com/uc?id=1fZRb-9RU5aVlcZ2P05sBGVj_AAoVT6Kn" width="230">
We can actually reason through this with a piece of symmetry; since $\hat{i}$ and $\hat{u}$ are both unit vectors, projecting $\hat{i}$ onto the line passing through $\hat{u}$, looks completely symmetric to projecting $\hat{u}$ onto the x-axis.
<img src="https://drive.google.com/uc?id=160NpTPVDm5ydgQnBYcVOlADlwfM55xni" width="230">
So, when we ask what number does $\hat{i}$ land on when it gets projected, the answer will be the same as what number $\hat{u}$ lands on when it gets projected onto the x-axis. But, projecting $\hat{u}$ onto the x-axis just means taking the x coordinate of $\hat{u}$:
<img src="https://drive.google.com/uc?id=11zjaLPVuWoa5WIFLGmL19Cgs4_-GjFif" width="230">
We can use nearly identical reasoning when determining where $\hat{j}$ lands!
<img src="https://drive.google.com/uc?id=1_LWImNskhCi7MUqqfkjgLDior1IgCAOh" width="230">
So, the entries of a 1x2 transformation matrix describing the projection transformation are going to be the coordinates of $\hat{u}$:
$$\begin{bmatrix}
u_x & u_y
\end{bmatrix}$$
And computing that transformation for arbitrary vectors in space:
<img src="https://drive.google.com/uc?id=1lCP3loW0YfY7oL8ihr6Qjogy9HCmOwby" width="300">
Requires multiplying that matrix by those vectors:
$$
\begin{bmatrix}
u_x & u_y
\end{bmatrix}
\begin{bmatrix}
x \\
y
\end{bmatrix} =
u_x * x + u_y * y
$$
This is computationally identical to taking the dot product with $\hat{u}$!
$$
\begin{bmatrix}
u_x \\
u_y
\end{bmatrix}
\cdot
\begin{bmatrix}
x \\
y
\end{bmatrix} =
u_x * x + u_y * y
$$
**Key Point**:<br>
> This is why _taking the dot product with a unit vector_ can be interpreted as _projecting a vector onto the span of that unit vector and taking the length_.
Let's take a moment to think about what just happened here; we started with a linear transformation from 2-d space to the number line, which was _not_ defined in terms of numerical vectors or numerical dot products. Rather, it was defined by projecting space onto a diagonal copy of the number line. But, _because the translation was linear_, it was necessarily _described by some 1x2 matrix_. And since multiplying some 1x2 matrix by a 2-d vector is the same as turning that matrix on its side and taking the dot product, this transformation was, _inescapably_ related to some 2-d vector.
The lesson here is that any time you have one of these linear transformations, whose output space is the number line, no matter how it was defined, there is going to be some unique vector $\vec{v}$ corresponding to that transformation, in the sense that applying that transformation is the same as taking a dot product with that vector.
### 4.4 Duality
What just happened above in math is an example of _**duality**_. Loosely speaking, it is defined as:
> _**Duality: A natural-but-surprising correspondence between two types of mathematical things.**_
For the linear algebra case we just saw, we would say the dual of a vector is the linear transformation that it encodes. And the dual of a linear transformation from some space to 1-dimension, is a certain vector in that space.
### 4.5 Summary
In summation:
> The dot product is a very useful geometric tool for understanding projections, and for testing whether or not vectors generally point in the same direction.
The above statement is probably the most important thing to remember about the dot product. But at a deeper level:
> Dotting two vectors together is a way to translate one of them into the world of transformations.
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.