text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# f-scLVM
In this notebook we illustrate how f-scLVM can be used to identify biological processes driving variability between cells.
First, we load some modules and set some directories; we here use data from 81 differentiating T-cells.
```
import sys
import os
import scipy as SP
import fscLVM
from fscLVM import plotFactors, plotRelevance, saveFA, dumpFA
%pylab inline
#specify where the hdf5 file is
data_dir = '../data/'
out_dir = './results/'
```
f-scLVM expects two input files, a gene expression file and an annotation file. The gene expression file is a text file containing the normalised, log-transformed gene expression matrix, with every row corresponding to a cell. Column names should be gene identifiers matching those in the annotation file. The annotation file is a text file with every row containing the name of a gene set, followed by the gene identifiers annotated to that gene set. We recommend using annotations such as those published in the REACTOME database or the Molecular signature database (MSigDB).
We provide a utility function, `load_txt`, for loading both text files. The loaded data can then be used to initialise the model with the `initFA` function.
```
#Annotation files
#Custom annotations for Th2 term and additional cell cycle term
annoFile = os.path.join(data_dir,'gene_lists_fscLVM.txt') #custom
annoFileMSigDB = os.path.join(data_dir,'h.all.v5.0.symbols.gmt.txt') #MSigDB
#log transformed gene expression (cells x genes)
dataFile = os.path.join(data_dir,'Tcell.csv.zip')
#load data - here we have 2 annotation files, one containng MSigDB annotations,
#the other containing custom gene sets; they can be passed to the load function as a list.
data = fscLVM.load_txt(dataFile, [annoFileMSigDB,annoFile],annoDBs=['MSigDB', 'custom'],
niceTerms=[True,False], dataFile_delimiter=';')
I = data['I']
Y = data['Y']
terms = data['terms']
print(I.shape)
print(Y.shape)
#initialise model
FA = fscLVM.initFA(Y, terms,I,noise='gauss', nHidden=1, nHiddenSparse=0,
pruneGenes=True, minGenes=15)
FA.terms
```
Next, we train the model and print diagnostics.
```
#model training
FA.train()
#print diagnostics
FA.printDiagnostics()
```
We next plot the relevance score of the individual terms. Since 2 of the main drivers of variability are cell cycle and Th2 differenitation, we also generate scatter plot, visualising the cell states of all cell with respect to these factors. Cells are coloured with the clusters identified in Buettner et al 2015.
```
#FA.terms = SP.array(FA.terms)
#fig = plotRelevance(FA)
fscLVM.utils.plotTerms(FA=FA)
#get factors; analogous getters are implemented for relevance and weights (see docs)
plt_terms = ['G2m checkpoint','Th2']
X = FA.getX(terms=plt_terms)
#scatter plot of the top two factors
fig = plotFactors(X=-X, lab=data['lab'], terms=plt_terms,
isCont=False, cols=SP.array(['red','blue']))
```
Next, we can regress out the cell cycle and hidden factors and visualise the residuals using a Bayesian GPLVM. This step requires the `GPy` package which can be installed via `pip`.
```
import GPy #
#Get model residuals
Ycorr = FA.regressOut(terms = ['hidden0', 'G2m checkpoint','Cell.cycle'])
## Model optimization
Ystd = Ycorr-Ycorr.mean(0)
input_dim = 2 # How many latent dimensions to use
kern = GPy.kern.RBF(input_dim) # RBF kernel
m = GPy.models.BayesianGPLVM(Ystd, input_dim=input_dim, kernel=kern, num_inducing=40)
m.optimize('scg', messages=1, max_iters=2000)
```
Visualisation of the residuals. The colors correspond again to the clusters identified in Buettner et al 2015.
```
import pylab as PL
PL.scatter(m.X.mean[:,0], m.X.mean[:,1], 40, data['lab'])
PL.xlabel('Component 1')
PL.ylabel('Component 2')
PL.colorbar()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/vlad-danaila/ml-cancer-detection/blob/master/Cancer_Detection_Ensable_V1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
!pip3 install sklearn matplotlib GPUtil
!pip3 install "pillow<7"
!pip3 install torch==1.3.1+cu92 torchvision==0.4.2+cu92 -f https://download.pytorch.org/whl/torch_stable.html
```
**Download Data**
Mount my google drive, where the dataset is stored.
```
try:
from google.colab import drive
drive.mount('/content/drive')
except Exception as e:
print(e)
```
Unzip the dataset into the folder "dataset".
```
!rm -vrf "dataset"
!mkdir "dataset"
!cp -r "/content/drive/My Drive/Studiu doctorat leziuni cervicale/cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
# !cp -r "cervigram-image-dataset-v2.zip" "dataset/cervigram-image-dataset-v2.zip"
!unzip "dataset/cervigram-image-dataset-v2.zip" -d "dataset"
```
**Constants**
```
# TRAIN_PATH = '/content/dataset/data/train/'
# TEST_PATH = '/content/dataset/data/test/'
TRAIN_PATH = 'dataset/data/train/'
TEST_PATH = 'dataset/data/test/'
CROP_SIZE = 260
IMAGE_SIZE = 224
BATCH_SIZE = 100
# prefix = '/content/drive/My Drive/Studiu doctorat leziuni cervicale/V2/Chekpoints & Notebooks/'
prefix = 'Mobilenetv2 Tuning/'
CHACKPOINT_CROSS_ENTROPY_MODEL = prefix + 'ResNet18 Baseline - Model Checkpoint.tar'
```
**Imports**
```
import torch as t
import torchvision as tv
import numpy as np
import PIL as pil
import matplotlib.pyplot as plt
from torchvision.datasets import ImageFolder
from torch.utils.data import DataLoader
from torch.nn import Linear, BCEWithLogitsLoss
import sklearn as sk
import sklearn.metrics
from os import listdir
import time
import random
import statistics
import math
```
**Deterministic Measurements**
This statements help making the experiments reproductible by fixing the random seeds.
```
SEED = 0
t.manual_seed(SEED)
t.cuda.manual_seed(SEED)
t.cuda.manual_seed_all(SEED)
t.backends.cudnn.deterministic = True
t.backends.cudnn.benchmark = False
np.random.seed(SEED)
random.seed(SEED)
```
**Memory Stats**
```
import GPUtil
def memory_stats():
for gpu in GPUtil.getGPUs():
print("GPU RAM Free: {0:.0f}MB | Used: {1:.0f}MB | Util {2:3.0f}% | Total {3:.0f}MB".format(gpu.memoryFree, gpu.memoryUsed, gpu.memoryUtil*100, gpu.memoryTotal))
memory_stats()
```
**Loading Data**
The dataset is structured in multiple small folders, ecah containing 7 images. The generator iterates through the folders and returns the category and 7 paths: one for each image in the folder.
```
def sortByLastDigits(elem):
chars = [c for c in elem if c.isdigit()]
return 0 if len(chars) == 0 else int(''.join(chars))
def getImagesPaths(root_path):
for class_folder in [root_path + f for f in listdir(root_path)]:
category = int(class_folder[-1])
for case_folder in listdir(class_folder):
case_folder_path = class_folder + '/' + case_folder + '/'
img_files = [case_folder_path + file_name for file_name in listdir(case_folder_path)]
yield category, sorted(img_files, key = sortByLastDigits)
```
We define 3 datasets, which loads 3 kinds of images: natural images, images taken through a green lens and images where the doctor aplied iodine solution (which gives a dark red colour). Each dataset has dynamic and static transformations which could be applied to the data. The static transformations are applied on the initialization of the dataset, while the dynamic ones are applied when loading each batch of data.
```
class SimpleImagesDataset(t.utils.data.Dataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
for i in range(5):
img = pil.Image.open(img_files[i])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
def __getitem__(self, i):
x, y = self.dataset[i]
if self.transforms_x != None:
x = self.transforms_x(x)
if self.transforms_y != None:
y = self.transforms_y(y)
return x, y
def __len__(self):
return len(self.dataset)
class GreenLensImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-2])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
class RedImagesDataset(SimpleImagesDataset):
def __init__(self, root_path, transforms_x_static = None, transforms_x_dynamic = None, transforms_y_static = None, transforms_y_dynamic = None):
self.dataset = []
self.transforms_x = transforms_x_dynamic
self.transforms_y = transforms_y_dynamic
for category, img_files in getImagesPaths(root_path):
# Only the green lens image
img = pil.Image.open(img_files[-1])
if transforms_x_static != None:
img = transforms_x_static(img)
if transforms_y_static != None:
category = transforms_y_static(category)
self.dataset.append((img, category))
```
**Preprocess Data**
Convert pytorch tensor to numpy array.
```
def to_numpy(x):
return x.cpu().detach().numpy()
```
Data transformations for the test and training sets.
```
norm_mean = [0.485, 0.456, 0.406]
norm_std = [0.229, 0.224, 0.225]
transforms_train = tv.transforms.Compose([
tv.transforms.RandomAffine(degrees = 45, translate = None, scale = (1., 2.), shear = 30),
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.RandomHorizontalFlip(),
tv.transforms.ToTensor(),
tv.transforms.Lambda(lambda t: t.cuda()),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
transforms_test = tv.transforms.Compose([
# tv.transforms.CenterCrop(CROP_SIZE),
tv.transforms.Resize(IMAGE_SIZE),
tv.transforms.ToTensor(),
tv.transforms.Normalize(mean=norm_mean, std=norm_std)
])
y_transform = tv.transforms.Lambda(lambda y: t.tensor(y, dtype=t.long, device = 'cuda:0'))
```
Initialize pytorch datasets and loaders for training and test.
```
def create_loaders(dataset_class):
dataset_train = dataset_class(TRAIN_PATH, transforms_x_dynamic = transforms_train, transforms_y_dynamic = y_transform)
dataset_test = dataset_class(TEST_PATH, transforms_x_static = transforms_test,
transforms_x_dynamic = tv.transforms.Lambda(lambda t: t.cuda()), transforms_y_dynamic = y_transform)
loader_train = DataLoader(dataset_train, BATCH_SIZE, shuffle = True, num_workers = 0)
loader_test = DataLoader(dataset_test, BATCH_SIZE, shuffle = False, num_workers = 0)
return loader_train, loader_test, len(dataset_train), len(dataset_test)
loader_train_simple_img, loader_test_simple_img, len_train, len_test = create_loaders(SimpleImagesDataset)
```
**Visualize Data**
Load a few images so that we can see the efects of the data augmentation on the training set.
```
def plot_one_prediction(x, label, pred):
x, label, pred = to_numpy(x), to_numpy(label), to_numpy(pred)
x = np.transpose(x, [1, 2, 0])
if x.shape[-1] == 1:
x = x.squeeze()
x = x * np.array(norm_std) + np.array(norm_mean)
plt.title(label, color = 'green' if label == pred else 'red')
plt.imshow(x)
def plot_predictions(imgs, labels, preds):
fig = plt.figure(figsize = (20, 5))
for i in range(20):
fig.add_subplot(2, 10, i + 1, xticks = [], yticks = [])
plot_one_prediction(imgs[i], labels[i], preds[i])
# x, y = next(iter(loader_train_simple_img))
# for i in range(7):
# plot_predictions(x[i], y, y)
```
**Model**
```
def get_resnet_18():
model = tv.models.resnet18(pretrained = True)
model.fc.out_features = 4
model = model.cuda()
return model
model_simple = t.nn.DataParallel(get_resnet_18())
```
**Train & Evaluate**
Timer utility function. This is used to measure the execution speend.
```
time_start = 0
def timer_start():
global time_start
time_start = time.time()
def timer_end():
return time.time() - time_start
```
This function trains the network and e valuates it at the same time. It outputs the metrics recorder during the training for both train and test. We are measuring accuracy and the loss. The function also saves a checkpoint of the model every time the accuracy is imporved. In the end we will have a checkpoint of the model which gave the best accuracy.
```
def train_eval(optimizer, model, loader_train, loader_test, chekpoint_name, epochs):
metrics = {
'losses_train': [],
'losses_test': [],
'acc_train': [],
'acc_test': [],
'prec_train': [],
'prec_test': [],
'rec_train': [],
'rec_test': [],
'f_score_train': [],
'f_score_test': [],
'mean_class_acc_train': [],
'mean_class_acc_test': []
}
best_mean_acc = 0
loss_weights = t.tensor([1/4] * 4, device='cuda:0')
try:
for epoch in range(epochs):
timer_start()
loss_fn = t.nn.CrossEntropyLoss(weight = loss_weights)
# loss_fn = t.nn.CrossEntropyLoss()
# loss_fn = FocalLoss(gamma = 2)
train_epoch_loss, train_epoch_acc, train_epoch_precision, train_epoch_recall, train_epoch_f_score = 0, 0, 0, 0, 0
test_epoch_loss, test_epoch_acc, test_epoch_precision, test_epoch_recall, test_epoch_f_score = 0, 0, 0, 0, 0
# Train
model.train()
conf_matrix = np.zeros((4, 4))
for x, y in loader_train:
y_pred = model.forward(x)
y_pred = y_pred.narrow(1, 0, 4)
loss = loss_fn(y_pred, y)
loss.backward()
optimizer.step()
# memory_stats()
optimizer.zero_grad()
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_train
train_epoch_loss += (loss.item() * ratio)
train_epoch_acc += (sk.metrics.accuracy_score(y, pred) * ratio)
precision, recall, f_score, _ = sk.metrics.precision_recall_fscore_support(y, pred, average = 'macro')
train_epoch_precision += (precision * ratio)
train_epoch_recall += (recall * ratio)
train_epoch_f_score += (f_score * ratio)
conf_matrix += sk.metrics.confusion_matrix(y, pred, labels = list(range(4)))
class_acc = [conf_matrix[i][i] / sum(conf_matrix[i]) for i in range(len(conf_matrix))]
mean_class_acc = statistics.harmonic_mean(class_acc)
errors = [1 - conf_matrix[i][i] / sum(conf_matrix[i]) for i in range(len(conf_matrix))]
errors_strong = [math.exp(100 * e) for e in errors]
loss_weights = t.tensor([e / sum(errors_strong) for e in errors_strong], device = 'cuda:0')
metrics['losses_train'].append(train_epoch_loss)
metrics['acc_train'].append(train_epoch_acc)
metrics['prec_train'].append(train_epoch_precision)
metrics['rec_train'].append(train_epoch_recall)
metrics['f_score_train'].append(train_epoch_f_score)
metrics['mean_class_acc_train'].append(mean_class_acc)
# Evaluate
model.eval()
with t.no_grad():
conf_matrix_test = np.zeros((4, 4))
for x, y in loader_test:
y_pred = model.forward(x)
y_pred = y_pred.narrow(1, 0, 4)
loss = loss_fn(y_pred, y)
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
ratio = len(y) / len_test
test_epoch_loss += (loss * ratio)
test_epoch_acc += (sk.metrics.accuracy_score(y, pred) * ratio )
precision, recall, f_score, _ = sk.metrics.precision_recall_fscore_support(y, pred, average = 'macro')
test_epoch_precision += (precision * ratio)
test_epoch_recall += (recall * ratio)
test_epoch_f_score += (f_score * ratio)
conf_matrix_test += sk.metrics.confusion_matrix(y, pred, labels = list(range(4)))
class_acc_test = [conf_matrix_test[i][i] / sum(conf_matrix_test[i]) for i in range(len(conf_matrix_test))]
mean_class_acc_test = statistics.harmonic_mean(class_acc_test)
metrics['losses_test'].append(test_epoch_loss)
metrics['acc_test'].append(test_epoch_acc)
metrics['prec_test'].append(test_epoch_precision)
metrics['rec_test'].append(test_epoch_recall)
metrics['f_score_test'].append(test_epoch_f_score)
metrics['mean_class_acc_test'].append(mean_class_acc_test)
if metrics['mean_class_acc_test'][-1] > best_mean_acc:
best_mean_acc = metrics['mean_class_acc_test'][-1]
t.save({'model': model.state_dict()}, 'checkpint {}.tar'.format(chekpoint_name))
print('Epoch {} mean class acc {} acc {} prec {} rec {} f {} minutes {}'.format(
epoch + 1, metrics['mean_class_acc_test'][-1], metrics['acc_test'][-1], metrics['prec_test'][-1], metrics['rec_test'][-1], metrics['f_score_test'][-1], timer_end() / 60))
except KeyboardInterrupt as e:
print(e)
print('Ended training')
return metrics
```
Plot a metric for both train and test.
```
def plot_train_test(train, test, title, y_title):
plt.plot(range(len(train)), train, label = 'train')
plt.plot(range(len(test)), test, label = 'test')
plt.xlabel('Epochs')
plt.ylabel(y_title)
plt.title(title)
plt.legend()
plt.show()
def plot_precision_recall(metrics):
plt.scatter(metrics['prec_train'], metrics['rec_train'], label = 'train')
plt.scatter(metrics['prec_test'], metrics['rec_test'], label = 'test')
plt.legend()
plt.title('Precision-Recall')
plt.xlabel('Precision')
plt.ylabel('Recall')
```
Train a model for several epochs. The steps_learning parameter is a list of tuples. Each tuple specifies the steps and the learning rate.
```
def do_train(model, loader_train, loader_test, checkpoint_name, steps_learning):
t.cuda.empty_cache()
for steps, learn_rate in steps_learning:
metrics = train_eval(t.optim.Adam(model.parameters(), lr = learn_rate, weight_decay = 0), model, loader_train, loader_test, checkpoint_name, steps)
index_max = np.array(metrics['mean_class_acc_test']).argmax()
print('Best mean class accuracy :', metrics['mean_class_acc_test'][index_max])
print('Best test accuracy :', metrics['acc_test'][index_max])
print('Corresponding precision :', metrics['prec_test'][index_max])
print('Corresponding recall :', metrics['rec_test'][index_max])
print('Corresponding f1 score :', metrics['f_score_test'][index_max])
plot_train_test(metrics['mean_class_acc_train'], metrics['mean_class_acc_test'], 'Mean Class Accuracy (lr = {})'.format(learn_rate), 'Mean Class Accuracy')
plot_train_test(metrics['losses_train'], metrics['losses_test'], 'Loss (lr = {})'.format(learn_rate), 'Loss')
plot_train_test(metrics['acc_train'], metrics['acc_test'], 'Accuracy (lr = {})'.format(learn_rate), 'Accuracy')
plot_train_test(metrics['prec_train'], metrics['prec_test'], 'Precision (lr = {})'.format(learn_rate), 'Precision')
plot_train_test(metrics['rec_train'], metrics['rec_test'], 'Recall (lr = {})'.format(learn_rate), 'Recall')
plot_train_test(metrics['f_score_train'], metrics['f_score_test'], 'F1 Score (lr = {})'.format(learn_rate), 'F1 Score')
plot_precision_recall(metrics)
```
Perform actual training
```
do_train(model_simple, loader_train_simple_img, loader_test_simple_img, 'wce resnet 18', [(100, 1e-4)])
checkpoint = t.load('checkpint wce resnet 18.tar')
model_simple.load_state_dict(checkpoint['model'])
def calculate_class_acc_for_test_set(model):
model.eval()
with t.no_grad():
conf_matrix = np.zeros((4, 4))
for x, y in loader_test_simple_img:
y_pred = model.forward(x)
y_pred = y_pred.narrow(1, 0, 4)
y_pred, y = to_numpy(y_pred), to_numpy(y)
pred = y_pred.argmax(axis = 1)
cm = sk.metrics.confusion_matrix(y, pred, labels = list(range(4)))
conf_matrix += cm
print('Confusion matrix:\n', conf_matrix)
class_acc = [conf_matrix[i][i] / sum(conf_matrix[i]) for i in range(len(conf_matrix))]
print('Class acc:\n', class_acc)
return class_acc
def plot_class_acc(class_acc):
plt.bar(list(range(4)), class_acc, align='center', alpha=0.5)
plt.xticks(list(range(4)), list(range(4)))
plt.xlabel('Classes')
plt.ylabel('True Positive Rate')
plt.savefig('AccPerClass.pdf', dpi = 300, format = 'pdf')
plt.show()
def plot_class_acc_comparison(class_acc_1, class_acc_2, title_1, title_2):
width = .3
plt.bar(list(range(4)), class_acc_1, width, alpha=0.5, color = 'green', label = title_1)
plt.bar(np.array(list(range(4))) + width, class_acc_2, width, alpha=0.5, color = 'blue', label = title_2)
plt.xticks(np.array(list(range(4))) + width/2, list(range(4)))
plt.xlabel('Classes')
plt.ylabel('True Positive Rate')
plt.legend()
plt.savefig('ClassAccCompareDenseNet.pdf', dpi = 300, format = 'pdf')
plt.show()
class_acc = calculate_class_acc_for_test_set(model_simple)
plot_class_acc(class_acc)
model_without_wl = t.nn.DataParallel(get_resnet_18().cuda())
checkpoint_without_wl = t.load(CHACKPOINT_CROSS_ENTROPY_MODEL)
model_without_wl.load_state_dict(checkpoint_without_wl['model'])
class_acc_no_wl = calculate_class_acc_for_test_set(model_without_wl)
plot_class_acc(class_acc_no_wl)
plot_class_acc_comparison(class_acc, class_acc_no_wl, 'Weighted Cross Entropy Loss', 'Cross Entropy Loss')
```
| github_jupyter |
```
import pandas as pd
# import numpy as np
import csv
from nltk.tokenize import sent_tokenize
dataset = pd.read_csv("dev_nsp.tsv", delimiter = '\t')
print(dataset)
dataset.context
sentences = []
for c in dataset.context:
sentences.append(c)
print(sentences)
print(sentences[0])
sentences[1]
print(sentences[962])
len(sentences)
test = []
sentence0 = []
sentence1 = []
sentence2 = []
sentence3 = []
# test용 preprocess
for s in sentences:
sentence_list = sent_tokenize(s)
length = len(sentence_list)
index = length//4
if length %4 ==0:
sentence0 = sentence_list[0:index]
sentence1 = sentence_list[index:index*2]
sentence2 = sentence_list[index*2:index*3]
sentence3 = sentence_list[index*3:]
elif length %4 ==1:
sentence0 = sentence_list[0:index]
sentence1 = sentence_list[index:index*2]
sentence2 = sentence_list[index*2:index*3]
sentence3 = sentence_list[index*3:]
elif length %4 ==2:
sentence0 = sentence_list[0:index]
sentence1 = sentence_list[index:index*2]
sentence2 = sentence_list[index*2:index*3+1]
sentence3 = sentence_list[index*3+1:]
elif length %4 ==3:
sentence0 = sentence_list[0:index]
sentence1 = sentence_list[index:index*2+1]
sentence2 = sentence_list[index*2+1:index*3+1]
sentence3 = sentence_list[index*3+1:]
s0 = ''.join(sentence0)
s1 = ''.join(sentence1)
s2 = ''.join(sentence2)
s3 = ''.join(sentence3)
count = sentences.index(s)
items= [s0, s1,s2,s3]
for i, t1 in enumerate(items):
for j, t2 in enumerate(items):
test.append([t1, t2])
len(test)
print(test)
test[0]
test[1]
test_len = len(test) # 963개 데이터를 16개 문장으로 한 리스트
div = test_len//16
print(test_len)
print(div)
# TEXTA_INDEX = 2
# TEXTB_INDEX = 3
with open('test1.tsv', 'wt', -1, "utf-8") as out_file:
tsv_writer = csv.writer(out_file, delimiter='\t')
tsv_writer.writerow([' ', ' ', 'texta', 'textb'])
for i in range(div):
texta = test[i][0]
textb = test[i][1]
tsv_writer.writerow([' ', ' ', texta, textb])
# TEXTA_INDEX = 2
# TEXTB_INDEX = 3
with open('test2.tsv', 'wt', -1, "utf-8") as out_file:
tsv_writer = csv.writer(out_file, delimiter='\t')
tsv_writer.writerow([' ', ' ', 'texta', 'textb'])
for i in range(div):
texta = test[i+div][0]
textb = test[i+div][1]
tsv_writer.writerow([' ', ' ', texta, textb])
```
| github_jupyter |
## We need graphviz for visualization
There's a [tutorial](https://bobswift.atlassian.net/wiki/spaces/GVIZ/pages/20971549/How+to+install+Graphviz+software) to help with the installation, specially the extra stuff needed for Windows or OSX.
```
!pip install graphviz # graphviz for python
import warnings
warnings.filterwarnings("ignore")
%matplotlib inline
import graphviz
import matplotlib.pyplot as plt
import numpy as np
from sklearn import tree
from sklearn import datasets
from sklearn import model_selection
from sklearn.metrics import classification_report
```
# Decision trees
(example from sklearn)
```
iris = datasets.load_iris()
X_train, X_test, y_train, y_test = model_selection.train_test_split(iris.data, iris.target, test_size=0.33, random_state=3)
clf = tree.DecisionTreeClassifier(max_depth=2)
clf = clf.fit(X_train, y_train)
dot_data = tree.export_graphviz(clf, out_file=None,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(dot_data)
graph
predictions = clf.predict(X_train)
print(classification_report(y_train, predictions, target_names=["setosa", "versicolor", "virginica"]))
```
### Increasing the depth...
```
clf = tree.DecisionTreeClassifier()
clf = clf.fit(X_train, y_train)
dot_data = tree.export_graphviz(clf, out_file=None,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(dot_data)
graph
predictions = clf.predict(X_train)
print(classification_report(y_train, predictions, target_names=["setosa", "versicolor", "virginica"]))
```
### And what if we look at the accuracy over the test data?
```
predictions = clf.predict(X_test)
print(classification_report(y_test, predictions, target_names=["setosa", "versicolor", "virginica"]))
```
## Regression Trees
```
boston = datasets.load_boston()
X = boston.data[:, 12] # Only using the LSTAT feature (percentage of lower status of the population)
y = boston.target
# Sort X and y by ascending values of X
sort_idx = X.flatten().argsort()
X = X[sort_idx].reshape(-1, 1)
y = y[sort_idx]
clf = tree.DecisionTreeRegressor(max_depth=3, criterion="mse")
clf = clf.fit(X, y)
```
### What do the leafs return in this case?
```
dot_data = tree.export_graphviz(clf, out_file=None,
filled=True, rounded=True,
special_characters=True)
graph = graphviz.Source(dot_data)
graph
```
### Let's check it out
```
plt.figure(figsize=(16, 8))
plt.scatter(X, y, c='steelblue',
edgecolor='white', s=70)
plt.plot(X, clf.predict(X),
color='black', lw=2)
plt.xlabel('% lower status of the population [LSTAT]')
plt.ylabel('Price in $1000s [MEDV]')
plt.show()
```
| github_jupyter |
```
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
%matplotlib inline
import argparse
import os
import pprint
import shutil
import cv2
from collections import defaultdict
import matplotlib.pyplot as plt
import numpy as np
import pickle
import torch
import torch.nn.parallel
import torch.backends.cudnn as cudnn
import torch.optim
import torch.utils.data
import torch.utils.data.distributed
import torchvision.transforms as transforms
#from tensorboardX import SummaryWriter
import _init_paths
from config import cfg
from config import update_config
from core.loss import JointsMSELoss
from core.function import train
from core.function import validate
from utils.utils import get_optimizer
from utils.utils import save_checkpoint
from utils.utils import create_logger
from utils.utils import get_model_summary
import dataset
import models
def parse_args():
parser = argparse.ArgumentParser(description='Train keypoints network')
# general
parser.add_argument('--cfg',
help='experiment configure file name',
default='experiments/atrw/w48_384x288.yaml',
type=str)
parser.add_argument('opts',
help="Modify config options using the command-line",
default=None,
nargs=argparse.REMAINDER)
# philly
parser.add_argument('--modelDir',
help='model directory',
type=str,
default='')
parser.add_argument('--logDir',
help='log directory',
type=str,
default='')
parser.add_argument('--dataDir',
help='data directory',
type=str,
default='')
parser.add_argument('--prevModelDir',
help='prev Model directory',
type=str,
default='')
args = parser.parse_args()
return args
class Args():
cfg='../experiments/awa/w48_384x288.yaml'
opts=''
modelDir=''
logDir=''
dataDir=''
prevModelDir=''
args=Args()
#args = parse_args()
update_config(cfg, args)
import os
import inspect
# Data loading code
normalize = transforms.Normalize(
mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225]
)
# train_dataset = eval('dataset.'+cfg.DATASET.DATASET)(
# cfg, cfg.DATASET.ROOT, cfg.DATASET.TRAIN_SET, True,
# transforms.Compose([
# transforms.ToTensor(),
# normalize,
# ])
# )
train_dataset = dataset.awa(
cfg, cfg.DATASET.ROOT, cfg.DATASET.TRAIN_SET, True,
transforms.Compose([
transforms.ToTensor(),
normalize,
])
)
print(os.path.abspath(inspect.getfile(dataset.atrw)))
input, target, target_weight, meta = train_dataset.test(0)
#meta['score']
# print(target_weight)
# print(meta['joints_vis'])
with open('/u/snaha/v6/dataset/AWA/Animals_with_Attributes2/quadruped_keypoints/coco_format/dataset.pickle', 'rb') as handle:
dataset = pickle.load(handle)
dataset['annotations'][0]
train_dataset.coco.dataset.keys()
train_dataset.coco.dataset['info']
train_dataset.coco.dataset['annotations'][50]
train_dataset.coco.dataset['images'][0]
train_dataset.coco.dataset['categories']
train_dataset.coco.imgs[4433]
train_dataset.coco.cats
train_dataset.coco.catToImgs[1]
train_dataset.coco.imgToAnns[4433]
train_dataset.coco.anns[2848]
def visualize(b, image):
for animal in b:
for name in b[animal]:
if name == 'bbox':
print(name)
start_point = (int(b[animal]['bbox'][0]+40), int(b[animal]['bbox'][1]+40))
end_point = ( int(b[animal]['bbox'][0]) + int(b[animal]['bbox'][2] - 40), int(b[animal]['bbox'][1]) + int(b[animal]['bbox'][3] - 40) )
color = (255, 0, 0)
thickness = 2
image = cv2.rectangle(image, start_point, end_point, color, thickness)
font = cv2.FONT_HERSHEY_SIMPLEX
org = (int(b[animal]['bbox'][0]), int(b[animal]['bbox'][1]))
fontScale = 1
image = cv2.putText(image, animal, org, font,
fontScale, color, thickness, cv2.LINE_AA)
# cv2.imshow('test', image)
# cv2.waitKey(0);
# cv2.destroyAllWindows();
else:
center_coordinates = (int(b[animal][name][0]), int(b[animal][name][1]))
radious = 1
color = (255, 0, 0)
thickness = 2
image = cv2.circle(image, center_coordinates, radious, color, thickness)
return image
im_root = '/u/snaha/v5/dataset/tiger/images/train'
b = dict()
b['a1'] = dict()
b['a1']['bbox'] = train_dataset.coco.anns[2848]['bbox']
keypoints = np.array(train_dataset.coco.anns[2848]['keypoints']).reshape(15,3)
image_path = os.path.join(im_root, '00'+str(train_dataset.coco.anns[2848]['image_id']) + '.jpg')
print(image_path)
img = cv2.imread(image_path)
print('image shape = ', img.shape)
print('bbox infor = ', b['a1']['bbox'])
for i, key in enumerate(train_dataset.coco.cats[1]['keypoints']):
b['a1'][key] = keypoints[i, :2].tolist()
visimage = visualize(b, img)
plt.imshow(img)
plt.show()
train_dataset.coco.cats[1]['keypoints']
```
| github_jupyter |
```
%%html
<style> table {float:left} </style>
!pip install torch tqdm lazyme nltk gensim
!python -m nltk.downloader punkt
import numpy as np
from tqdm import tqdm
import pandas as pd
from gensim.corpora import Dictionary
import torch
from torch import nn, optim, tensor, autograd
from torch.nn import functional as F
from torch.utils.data import Dataset, DataLoader
try: # Use the default NLTK tokenizer.
from nltk import word_tokenize, sent_tokenize
# Testing whether it works.
# Sometimes it doesn't work on some machines because of setup issues.
word_tokenize(sent_tokenize("This is a foobar sentence. Yes it is.")[0])
except: # Use a naive sentence tokenizer and toktok.
import re
from nltk.tokenize import ToktokTokenizer
# See https://stackoverflow.com/a/25736515/610569
sent_tokenize = lambda x: re.split(r'(?<=[^A-Z].[.?]) +(?=[A-Z])', x)
# Use the toktok tokenizer that requires no dependencies.
toktok = ToktokTokenizer()
word_tokenize = word_tokenize = toktok.tokenize
```
# Classifying Toxic Comments
Lets apply what we learnt in a realistic task and **fight cyber-abuse with NLP**!
From https://www.kaggle.com/c/jigsaw-toxic-comment-classification-challenge/
> *The threat of abuse and harassment online means that many people stop <br>*
> *expressing themselves and give up on seeking different opinions. <br>*
> *Platforms struggle to effectively facilitate conversations, leading many <br>*
> *communities to limit or completely shut down user comments.*
The goal of the task is to build a model to detect different types of of toxicity:
- toxic
- severe toxic
- threats
- obscenity
- insults
- identity-based hate
In this part, you'll be munging the data as how I would be doing it at work.
Your task is to train a feed-forward network on the toxic comments given the skills we have accomplished thus far.
## Digging into the data...
If you're using linux/Mac you can use these bang commands in the notebook:
```
!pip3 install kaggle
!mkdir -p /content/.kaggle/
!echo '{"username":"natgillin","key":"54ae95ab760b52c3307ed4645c6c9b5d"}' > /content/.kaggle/kaggle.json
!chmod 600 /content/.kaggle/kaggle.json
!kaggle competitions download -c jigsaw-toxic-comment-classification-challenge
!unzip /content/.kaggle/competitions/jigsaw-toxic-comment-classification-challenge/*
```
Otherwise, download the data from https://www.kaggle.com/c/jigsaw-toxic-comment-classification-challenge/
```
df_train = pd.read_csv('jigsaw-toxic-comment-classification-challenge/train.csv')
df_train.head()
df_train[df_train['threat'] == 1]['comment_text']
df_train.iloc[3712]['comment_text']
df_train['comment_text_tokenzied'] = df_train['comment_text'].apply(word_tokenize)
# Just in case your Jupyter kernel dies, save the tokenized text =)
# To save your tokenized text you can do this:
import pickle
with open('train_tokenized_text.pkl', 'wb') as fout:
pickle.dump(df_train['comment_text_tokenzied'], fout)
# To load it back:
import pickle
with open('train_tokenized_text.pkl', 'rb') as fin:
df_train['comment_text_tokenzied'] = pickle.load(fin)
```
# How to get a one-hot?
There are many variants of how to get your one-hot embeddings from the individual columns.
This is one way:
```
label_column_names = "toxic severe_toxic obscene threat insult identity_hate".split()
df_train[label_column_names].values
torch.tensor(df_train[label_column_names].values).float()
# Convert one-hot to indices of the column.
print(np.argmax(df_train[label_column_names].values, axis=1))
class ToxicDataset(Dataset):
def __init__(self, texts, labels):
self.texts = texts
self.vocab = Dictionary(texts)
special_tokens = {'<pad>': 0, '<unk>':1}
self.vocab = Dictionary(texts)
self.vocab.patch_with_special_tokens(special_tokens)
self.vocab_size = len(self.vocab)
# Vectorize labels
self.labels = torch.tensor(labels)
# Keep track of how many data points.
self._len = ???
# Find the longest text in the data.
self.max_len = max(len(txt) for txt in texts)
self.num_labels = len(labels[0])
def __getitem__(self, index):
vectorized_sent = self.vectorize(self.texts[index])
# To pad the sentence:
# Pad left = 0; Pad right = max_len - len of sent.
pad_dim = (0, self.max_len - len(vectorized_sent))
padded_vectorized_sent = F.pad(vectorized_sent, pad_dim, 'constant')
return {'x':padded_vectorized_sent,
'y':self.labels[index],
'x_len':len(vectorized_sent)}
def __len__(self):
return self._len
def vectorize(self, tokens):
"""
:param tokens: Tokens that should be vectorized.
:type tokens: list(str)
"""
# See https://radimrehurek.com/gensim/corpora/dictionary.html#gensim.corpora.dictionary.Dictionary.doc2idx
# Lets just cast list of indices into torch tensors directly =)
return torch.tensor(self.vocab.???(tokens))
def unvectorize(self, indices):
"""
:param indices: Converts the indices back to tokens.
:type tokens: list(int)
"""
return [self.vocab[i] for i in indices]
label_column_names = "toxic severe_toxic obscene threat insult identity_hate".split()
toxic_data = ToxicDataset(df_train['comment_text_tokenzied'],
df_train[label_column_names].values)
toxic_data[123]
batch_size = 5
dataloader = DataLoader(???)
class FFNet(nn.Module):
def __init__(self, max_len, num_labels, vocab_size, embedding_size, hidden_dim):
super(FFNet, self).__init__()
self.embeddings = nn.???(num_embeddings=vocab_size,
embedding_dim=embedding_size,
padding_idx=0)
# The no. of inputs to the linear layer is the
# no. of tokens in each input * embedding_size
self.linear1 = nn.???(embedding_size*max_len, hidden_dim)
self.linear2 = nn.???(hidden_dim, num_labels)
def forward(self, inputs):
# We want to flatten the inputs so that we get the matrix of shape.
# batch_size x no. of tokens in each input * embedding_size
batch_size, max_len = inputs.shape
embedded = self.???(inputs).view(batch_size, -1)
hidden = ???
out = ???
return ???
device = 'cuda' if torch.cuda.is_available() else 'cpu'
embedding_size = 100
learning_rate = 0.003
hidden_size = 100
criterion = nn.???()
# Hint: the CBOW model object you've created.
model = FFNet(toxic_data.max_len,
len(label_column_names),
toxic_data.vocab_size,
embedding_size=embedding_size,
hidden_dim=hidden_size)
optimizer = optim.???(???)
#model = nn.DataParallel(model)
losses = []
num_epochs = 100
for _e in range(num_epochs):
epoch_loss = []
for batch in tqdm(dataloader):
x = batch['x'].to(device)
y = batch['y'].to(device)
# Zero gradient.
optimizer.???
# Feed forward.
predictions = ???
loss = ???
# Back propagate the loss
???()
???()
epoch_loss.append(float(loss))
break
print(sum(epoch_loss)/len(epoch_loss))
losses.append(sum(epoch_loss)/len(epoch_loss))
def predict(text):
# Vectorize and Pad.
vectorized_sent = toxic_data.vectorize(word_tokenize(text))
pad_dim = (0, toxic_data.max_len - len(vectorized_sent))
vectorized_sent = F.pad(vectorized_sent, pad_dim, 'constant')
# Forward Propagation.
# Unsqueeze because model is expecting `batch_size` x `sequence_len` shape.
outputs = ???(vectorized_sent.unsqueeze(0))
# To get the boolean output, we check if outputs are > 0.5
return [int(l > 0.5) for l in outputs.squeeze()]
# What happens if you use torch.max instead? =)
##return label_column_names[int(torch.max(outputs, dim=1).indices)]
text = "This is a nice message."
print(label_column_names)
predict(text)
```
| github_jupyter |
# Basic Feature Engineering in Keras
**Learning Objectives**
1. Create an input pipeline using tf.data
2. Engineer features to create categorical, crossed, and numerical feature columns
## Introduction
In this lab, we utilize feature engineering to improve the prediction of housing prices using a Keras Sequential Model.
Each learning objective will correspond to a __#TODO__ in the [student lab notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/feature_engineering/labs/3_keras_basic_feat_eng-lab.ipynb) -- try to complete that notebook first before reviewing this solution notebook.
Start by importing the necessary libraries for this lab.
```
# Run the chown command to change the ownership
!sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
# Install Sklearn
# scikit-learn simple and efficient tools for predictive data analysis
# Built on NumPy, SciPy, and matplotlib
!python3 -m pip install --user sklearn
# Ensure the right version of Tensorflow is installed.
!pip freeze | grep tensorflow==2.1 || pip install tensorflow==2.1
```
**Note:** Please ignore any incompatibility warnings and errors and re-run the cell to view the installed tensorflow version.
`tensorflow==2.1.0` that is the installed version of tensorflow.
```
# You can use any Python source file as a module by executing an import statement in some other Python source file
# The import statement combines two operations; it searches for the named module, then it binds the
# results of that search to a name in the local scope.
import os
import tensorflow.keras
# Use matplotlib for visualizing the model
import matplotlib.pyplot as plt
# Import Pandas data processing libraries
import pandas as pd
import tensorflow as tf
from tensorflow import feature_column as fc
from tensorflow.keras import layers
from sklearn.model_selection import train_test_split
from keras.utils import plot_model
print("TensorFlow version: ",tf.version.VERSION)
```
Many of the Google Machine Learning Courses Programming Exercises use the [California Housing Dataset](https://developers.google.com/machine-learning/crash-course/california-housing-data-description
), which contains data drawn from the 1990 U.S. Census. Our lab dataset has been pre-processed so that there are no missing values.
First, let's download the raw .csv data by copying the data from a cloud storage bucket.
```
if not os.path.isdir("../data"):
os.makedirs("../data")
# Download the raw .csv data by copying the data from a cloud storage bucket.
!gsutil cp gs://cloud-training-demos/feat_eng/housing/housing_pre-proc.csv ../data
# `ls` is a Linux shell command that lists directory contents
# `l` flag list all the files with permissions and details
!ls -l ../data/
```
Now, let's read in the dataset just copied from the cloud storage bucket and create a Pandas dataframe.
```
# `head()` function is used to get the first n rows of dataframe
housing_df = pd.read_csv('../data/housing_pre-proc.csv', error_bad_lines=False)
housing_df.head()
```
We can use .describe() to see some summary statistics for the numeric fields in our dataframe. Note, for example, the count row and corresponding columns. The count shows 20433.000000 for all feature columns. Thus, there are no missing values.
```
# `describe()` is use to get the statistical summary of the DataFrame
housing_df.describe()
```
#### Split the dataset for ML
The dataset we loaded was a single CSV file. We will split this into train, validation, and test sets.
```
# Let's split the dataset into train, validation, and test sets
train, test = train_test_split(housing_df, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
```
Now, we need to output the split files. We will specifically need the test.csv later for testing. You should see the files appear in the home directory.
```
train.to_csv('../data/housing-train.csv', encoding='utf-8', index=False)
val.to_csv('../data/housing-val.csv', encoding='utf-8', index=False)
test.to_csv('../data/housing-test.csv', encoding='utf-8', index=False)
!head ../data/housing*.csv
```
## Lab Task 1: Create an input pipeline using tf.data
Next, we will wrap the dataframes with [tf.data](https://www.tensorflow.org/guide/datasets). This will enable us to use feature columns as a bridge to map from the columns in the Pandas dataframe to features used to train the model.
Here, we create an input pipeline using tf.data. This function is missing two lines. Correct and run the cell.
```
# A utility method to create a tf.data dataset from a Pandas Dataframe
# TODO 1a
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
labels = dataframe.pop('median_house_value')
ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels))
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
return ds
```
Next we initialize the training and validation datasets.
```
batch_size = 32
train_ds = df_to_dataset(train)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
```
Now that we have created the input pipeline, let's call it to see the format of the data it returns. We have used a small batch size to keep the output readable.
```
# TODO 1b
for feature_batch, label_batch in train_ds.take(1):
print('Every feature:', list(feature_batch.keys()))
print('A batch of households:', feature_batch['households'])
print('A batch of ocean_proximity:', feature_batch['ocean_proximity'])
print('A batch of targets:', label_batch)
```
We can see that the dataset returns a dictionary of column names (from the dataframe) that map to column values from rows in the dataframe.
#### Numeric columns
The output of a feature column becomes the input to the model. A numeric is the simplest type of column. It is used to represent real valued features. When using this column, your model will receive the column value from the dataframe unchanged.
In the California housing prices dataset, most columns from the dataframe are numeric. Let's create a variable called **numeric_cols** to hold only the numerical feature columns.
```
# Let's create a variable called `numeric_cols` to hold only the numerical feature columns.
# TODO 1c
numeric_cols = ['longitude', 'latitude', 'housing_median_age', 'total_rooms',
'total_bedrooms', 'population', 'households', 'median_income']
```
#### Scaler function
It is very important for numerical variables to get scaled before they are "fed" into the neural network. Here we use min-max scaling. Here we are creating a function named 'get_scal' which takes a list of numerical features and returns a 'minmax' function, which will be used in tf.feature_column.numeric_column() as normalizer_fn in parameters. 'Minmax' function itself takes a 'numerical' number from a particular feature and return scaled value of that number.
Next, we scale the numerical feature columns that we assigned to the variable "numeric cols".
```
# 'get_scal' function takes a list of numerical features and returns a 'minmax' function
# 'Minmax' function itself takes a 'numerical' number from a particular feature and return scaled value of that number.
# Scalar def get_scal(feature):
# TODO 1d
def get_scal(feature):
def minmax(x):
mini = train[feature].min()
maxi = train[feature].max()
return (x - mini)/(maxi-mini)
return(minmax)
# TODO 1e
feature_columns = []
for header in numeric_cols:
scal_input_fn = get_scal(header)
feature_columns.append(fc.numeric_column(header,
normalizer_fn=scal_input_fn))
```
Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
```
print('Total number of feature coLumns: ', len(feature_columns))
```
### Using the Keras Sequential Model
Next, we will run this cell to compile and fit the Keras Sequential model.
```
# Model create
# `tf.keras.layers.DenseFeatures()` is a layer that produces a dense Tensor based on given feature_columns.
feature_layer = tf.keras.layers.DenseFeatures(feature_columns, dtype='float64')
# `tf.keras.Sequential()` groups a linear stack of layers into a tf.keras.Model.
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
```
Next we show loss as Mean Square Error (MSE). Remember that MSE is the most commonly used regression loss function. MSE is the sum of squared distances between our target variable (e.g. housing median age) and predicted values.
```
# Let's show loss as Mean Square Error (MSE)
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
```
#### Visualize the model loss curve
Next, we will use matplotlib to draw the model's loss curves for training and validation. A line plot is also created showing the mean squared error loss over the training epochs for both the train (blue) and test (orange) sets.
```
# Use matplotlib to draw the model's loss curves for training and validation
def plot_curves(history, metrics):
nrows = 1
ncols = 2
fig = plt.figure(figsize=(10, 5))
for idx, key in enumerate(metrics):
ax = fig.add_subplot(nrows, ncols, idx+1)
plt.plot(history.history[key])
plt.plot(history.history['val_{}'.format(key)])
plt.title('model {}'.format(key))
plt.ylabel(key)
plt.xlabel('epoch')
plt.legend(['train', 'validation'], loc='upper left');
plot_curves(history, ['loss', 'mse'])
```
### Load test data
Next, we read in the test.csv file and validate that there are no null values.
Again, we can use .describe() to see some summary statistics for the numeric fields in our dataframe. The count shows 4087.000000 for all feature columns. Thus, there are no missing values.
```
test_data = pd.read_csv('../data/housing-test.csv')
test_data.describe()
```
Now that we have created an input pipeline using tf.data and compiled a Keras Sequential Model, we now create the input function for the test data and to initialize the test_predict variable.
```
# TODO 1f
def test_input_fn(features, batch_size=256):
"""An input function for prediction."""
# Convert the inputs to a Dataset without labels.
return tf.data.Dataset.from_tensor_slices(dict(features)).batch(batch_size)
test_predict = test_input_fn(dict(test_data))
```
#### Prediction: Linear Regression
Before we begin to feature engineer our feature columns, we should predict the median house value. By predicting the median house value now, we can then compare it with the median house value after feature engineering.
To predict with Keras, you simply call [model.predict()](https://keras.io/models/model/#predict) and pass in the housing features you want to predict the median_house_value for. Note: We are predicting the model locally.
```
# Use the model to do prediction with `model.predict()`
predicted_median_house_value = model.predict(test_predict)
```
Next, we run two predictions in separate cells - one where ocean_proximity=INLAND and one where ocean_proximity= NEAR OCEAN.
```
# Ocean_proximity is INLAND
model.predict({
'longitude': tf.convert_to_tensor([-121.86]),
'latitude': tf.convert_to_tensor([39.78]),
'housing_median_age': tf.convert_to_tensor([12.0]),
'total_rooms': tf.convert_to_tensor([7653.0]),
'total_bedrooms': tf.convert_to_tensor([1578.0]),
'population': tf.convert_to_tensor([3628.0]),
'households': tf.convert_to_tensor([1494.0]),
'median_income': tf.convert_to_tensor([3.0905]),
'ocean_proximity': tf.convert_to_tensor(['INLAND'])
}, steps=1)
# Ocean_proximity is NEAR OCEAN
model.predict({
'longitude': tf.convert_to_tensor([-122.43]),
'latitude': tf.convert_to_tensor([37.63]),
'housing_median_age': tf.convert_to_tensor([34.0]),
'total_rooms': tf.convert_to_tensor([4135.0]),
'total_bedrooms': tf.convert_to_tensor([687.0]),
'population': tf.convert_to_tensor([2154.0]),
'households': tf.convert_to_tensor([742.0]),
'median_income': tf.convert_to_tensor([4.9732]),
'ocean_proximity': tf.convert_to_tensor(['NEAR OCEAN'])
}, steps=1)
```
The arrays returns a predicted value. What do these numbers mean? Let's compare this value to the test set.
Go to the test.csv you read in a few cells up. Locate the first line and find the median_house_value - which should be 249,000 dollars near the ocean. What value did your model predicted for the median_house_value? Was it a solid model performance? Let's see if we can improve this a bit with feature engineering!
## Lab Task 2: Engineer features to create categorical and numerical features
Now we create a cell that indicates which features will be used in the model.
Note: Be sure to bucketize 'housing_median_age' and ensure that 'ocean_proximity' is one-hot encoded. And, don't forget your numeric values!
```
# TODO 2a
numeric_cols = ['longitude', 'latitude', 'housing_median_age', 'total_rooms',
'total_bedrooms', 'population', 'households', 'median_income']
bucketized_cols = ['housing_median_age']
# indicator columns,Categorical features
categorical_cols = ['ocean_proximity']
```
Next, we scale the numerical, bucktized, and categorical feature columns that we assigned to the variables in the preceding cell.
```
# Scalar def get_scal(feature):
def get_scal(feature):
def minmax(x):
mini = train[feature].min()
maxi = train[feature].max()
return (x - mini)/(maxi-mini)
return(minmax)
# All numerical features - scaling
feature_columns = []
for header in numeric_cols:
scal_input_fn = get_scal(header)
feature_columns.append(fc.numeric_column(header,
normalizer_fn=scal_input_fn))
```
### Categorical Feature
In this dataset, 'ocean_proximity' is represented as a string. We cannot feed strings directly to a model. Instead, we must first map them to numeric values. The categorical vocabulary columns provide a way to represent strings as a one-hot vector.
Next, we create a categorical feature using 'ocean_proximity'.
```
# TODO 2b
for feature_name in categorical_cols:
vocabulary = housing_df[feature_name].unique()
categorical_c = fc.categorical_column_with_vocabulary_list(feature_name, vocabulary)
one_hot = fc.indicator_column(categorical_c)
feature_columns.append(one_hot)
```
### Bucketized Feature
Often, you don't want to feed a number directly into the model, but instead split its value into different categories based on numerical ranges. Consider our raw data that represents a homes' age. Instead of representing the house age as a numeric column, we could split the home age into several buckets using a [bucketized column](https://www.tensorflow.org/api_docs/python/tf/feature_column/bucketized_column). Notice the one-hot values below describe which age range each row matches.
Next we create a bucketized column using 'housing_median_age'
```
# TODO 2c
age = fc.numeric_column("housing_median_age")
# Bucketized cols
age_buckets = fc.bucketized_column(age, boundaries=[10, 20, 30, 40, 50, 60, 80, 100])
feature_columns.append(age_buckets)
```
### Feature Cross
Combining features into a single feature, better known as [feature crosses](https://developers.google.com/machine-learning/glossary/#feature_cross), enables a model to learn separate weights for each combination of features.
Next, we create a feature cross of 'housing_median_age' and 'ocean_proximity'.
```
# TODO 2d
vocabulary = housing_df['ocean_proximity'].unique()
ocean_proximity = fc.categorical_column_with_vocabulary_list('ocean_proximity',
vocabulary)
crossed_feature = fc.crossed_column([age_buckets, ocean_proximity],
hash_bucket_size=1000)
crossed_feature = fc.indicator_column(crossed_feature)
feature_columns.append(crossed_feature)
```
Next, we should validate the total number of feature columns. Compare this number to the number of numeric features you input earlier.
```
print('Total number of feature columns: ', len(feature_columns))
```
Next, we will run this cell to compile and fit the Keras Sequential model. This is the same model we ran earlier.
```
# Model create
# `tf.keras.layers.DenseFeatures()` is a layer that produces a dense Tensor based on given feature_columns.
feature_layer = tf.keras.layers.DenseFeatures(feature_columns,
dtype='float64')
# `tf.keras.Sequential()` groups a linear stack of layers into a tf.keras.Model.
model = tf.keras.Sequential([
feature_layer,
layers.Dense(12, input_dim=8, activation='relu'),
layers.Dense(8, activation='relu'),
layers.Dense(1, activation='linear', name='median_house_value')
])
# Model compile
model.compile(optimizer='adam',
loss='mse',
metrics=['mse'])
# Model Fit
history = model.fit(train_ds,
validation_data=val_ds,
epochs=32)
```
Next, we show loss and mean squared error then plot the model.
```
loss, mse = model.evaluate(train_ds)
print("Mean Squared Error", mse)
plot_curves(history, ['loss', 'mse'])
```
Next we create a prediction model. Note: You may use the same values from the previous prediciton.
```
# TODO 2e
# Median_house_value is $249,000, prediction is $234,000 NEAR OCEAN
model.predict({
'longitude': tf.convert_to_tensor([-122.43]),
'latitude': tf.convert_to_tensor([37.63]),
'housing_median_age': tf.convert_to_tensor([34.0]),
'total_rooms': tf.convert_to_tensor([4135.0]),
'total_bedrooms': tf.convert_to_tensor([687.0]),
'population': tf.convert_to_tensor([2154.0]),
'households': tf.convert_to_tensor([742.0]),
'median_income': tf.convert_to_tensor([4.9732]),
'ocean_proximity': tf.convert_to_tensor(['NEAR OCEAN'])
}, steps=1)
```
### Analysis
The array returns a predicted value. Compare this value to the test set you ran earlier. Your predicted value may be a bit better.
Now that you have your "feature engineering template" setup, you can experiment by creating additional features. For example, you can create derived features, such as households per population, and see how they impact the model. You can also experiment with replacing the features you used to create the feature cross.
Copyright 2020 Google Inc.
Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
http://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
| github_jupyter |
```
import numpy as np
from scipy import stats
np.random.seed(2020)
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rcParams['axes.xmargin'] = 0.05
mpl.rcParams['axes.ymargin'] = 0.05
mpl.rcParams['axes.labelsize'] = 24
mpl.rcParams['axes.titlesize'] = 24
mpl.rcParams['legend.fontsize'] = 16
mpl.rcParams['xtick.labelsize'] = 12
mpl.rcParams['ytick.labelsize'] = 12
mpl.rcParams['legend.frameon'] = False
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'sans-serif'
from lio.utils.plot import cov2d, gaussian_mixture, simplex
```
# Data
```
dim_features = 2
num_classes = 3
num_points = 600
# [num_classes]
p_z = np.array([.4, .3, .3])
# [num_classes, dim_features]
mean = np.array([[1.5, 0.0],
[-2.0, 1.0],
[-1.0, -2.0]])
# [num_classes, dim_features]
std = np.array([[1, 1],
[1, 1],
[1, 1]])
# [num_classes]
corr = np.array([0, .5, -.3])
# [num_classes, dim_features, dim_features]
cov = cov2d(std, corr)
x, z = gaussian_mixture(mean, cov, n=num_points, p=p_z)
# [num_points, num_classes]
p_x_z = np.stack([stats.multivariate_normal(mean[i], cov[i]).pdf(x) for i in range(num_classes)], -1)
# [num_points, num_classes]
p_xz = p_x_z * p_z
# [num_points]
p_x = p_xz.sum(1)
# [num_points, num_classes]
p_z_x = p_xz / p_x[:, None]
U = np.array([
[.70, .25, .05],
[.15, .60, .25],
[.05, .15, .80],
])
V = np.array([
[.90, .05, .05],
[.15, .80, .05],
[.00, .10, .90],
])
T = U @ V
# [num_points]
y = np.array([np.random.choice(num_classes, p=T[zi]) for zi in z])
p_y_x = p_z_x @ T
```
# Training
```
import torch
import torch.nn.functional as F
from torch import nn, optim
from torch.utils.data import TensorDataset, DataLoader
torch.manual_seed(2020);
x_tensor = torch.tensor(x).float()
z_tensor = torch.tensor(z).long()
y_tensor = torch.tensor(y).long()
dataset_z = TensorDataset(x_tensor, z_tensor)
dataset_y = TensorDataset(x_tensor, y_tensor)
loader_z = DataLoader(dataset_z, batch_size=64, shuffle=True)
loader_y = DataLoader(dataset_y, batch_size=64, shuffle=True)
def get_model():
return nn.Sequential(
nn.Linear(dim_features, 32),
nn.ReLU(inplace=True),
nn.Linear(32, 32),
nn.ReLU(inplace=True),
nn.Linear(32, num_classes),
)
def train(model, optimizer, loader, epochs=30):
for epoch in range(epochs):
for x_batch, z_batch in loader:
loss = F.cross_entropy(model(x_batch), z_batch)
optimizer.zero_grad()
loss.backward()
optimizer.step()
model = get_model()
optimizer = optim.Adam(model.parameters())
train(model, optimizer, loader_z)
with torch.no_grad():
p_z_x_ = torch.softmax(model(x_tensor), 1).numpy()
z_ = p_z_x_.argmax(1)
model = get_model()
optimizer = optim.Adam(model.parameters())
train(model, optimizer, loader_y)
with torch.no_grad():
p_y_x_ = torch.softmax(model(x_tensor), 1).numpy()
y_ = p_y_x_.argmax(1)
```
# Plot
```
c_z = simplex.Tc[z]
c_y = simplex.Tc[y]
x_range = [0, np.log(3)]
def hist(ax, h, **kwargs):
ax.set_xlim(*x_range)
default = dict(density=True, log=True, color=simplex.Tc)
ax.hist(h, bins=np.linspace(*x_range, 20), **{**default, **kwargs})
def scatter(ax, x, z, c):
for cls, marker in enumerate('ovs'):
ax.scatter(*x[z == cls].T, color=c[z == cls], marker=marker)
def simplex_scatter(ax, p, z, c):
for cls, marker in enumerate('ovs'):
simplex.scatter(ax, p[z == cls], color=c[z == cls], marker=marker)
def entropy(p):
return -(p * np.log(p)).sum(1)
def estimate_transition_matrix(p, q=1):
threshold = np.quantile(p, q, axis=0, interpolation='higher')
return p[np.where(p > threshold, 0, p).argmax(axis=0)]
T_ = estimate_transition_matrix(p_y_x_)
fig, axes = plt.subplots(nrows=2, ncols=4, figsize=(20, 10))
gs = axes[0][0].get_gridspec()
# data
for i, c in zip(range(2), [c_z, c_y]):
ax = axes[i][0]
ax.set_xticks([])
ax.set_yticks([])
scatter(ax, x, z, c)
# empirical
for i, p, c in zip(range(2), [p_z_x, p_y_x], [c_z, c_y]):
ax = axes[i][1]
simplex.init(ax)
simplex.boundary(ax, zorder=0)
simplex_scatter(ax, p, z, c)
ax.axis('equal')
simplex.polygon(ax, T)
# estimated
for i, p, c in zip(range(2), [p_z_x_, p_y_x_], [c_z, c_y]):
ax = axes[i][2]
simplex.init(ax)
simplex.boundary(ax, zorder=0)
simplex_scatter(ax, p, z, c)
ax.axis('equal')
simplex.polygon(ax, T)
simplex.polygon(ax, T_, linestyle='dashed')
# entropy
sub_axes = []
for i, p, q in zip(range(2), [p_z_x, p_y_x], [p_z_x_, p_y_x_]):
axes[i][3].remove()
sub_gs = gs[i, -1].subgridspec(nrows=3, ncols=1, wspace=0, hspace=0)
sub_axes.append(sub_gs.subplots())
for j in range(3):
ax = sub_axes[i][j]
ax.yaxis.tick_right()
hist_params = dict(bins=np.linspace(*x_range, 20), log=True,
histtype='step', linewidth=3)
ax.hist(entropy(p[z == j]), label='Empirical',
color='xkcd:blue', hatch='/' * 3, **hist_params)
ax.hist(entropy(q[z_ == j]), label='Estimated',
color='xkcd:red', hatch='\\' * 3, **hist_params)
sub_axes[0][0].legend(loc='upper right')
sub_axes[0][2].set_xticks([])
# labels
axes[0][0].set_ylabel('Clean')
axes[1][0].set_ylabel('Noisy')
axes[0][0].set_title('Data')
axes[0][1].set_title('Empirical distribution')
axes[0][2].set_title('Estimated distribution')
sub_axes[0][0].set_title('Histogram of entropy')
# annotations
text_params = dict(fontsize=20, ha='center', va='center')
arrowstyle = mpl.patches.ArrowStyle('simple', head_length=1, head_width=1)
arrow_params = dict(coordsA='data', coordsB='data',
facecolor='k', linewidth=1, arrowstyle=arrowstyle)
ax = axes[1][2]
ax.text(0.8, 0.82, 'overconfidence', **text_params)
ax.add_artist(mpl.patches.ConnectionPatch(
xyA=(0.8, 0.78), xyB=(0.45, 0.7), axesA=ax, **arrow_params))
ax.add_artist(mpl.patches.ConnectionPatch(
xyA=(0.8, 0.78), xyB=(0.9, 0.05), axesA=ax, **arrow_params))
sub_axes[1][1].text(0.18, 10, 'overconfidence', **text_params)
sub_axes[1][1].add_artist(mpl.patches.ConnectionPatch(
xyA=(0.42, 10), xyB=(0.6, 5),
axesA=sub_axes[1][1], axesB=sub_axes[1][0], **arrow_params))
sub_axes[1][2].add_artist(mpl.patches.ConnectionPatch(
xyA=(0.42, 10), xyB=(0.5, 5),
axesA=sub_axes[1][1], axesB=sub_axes[1][2], **arrow_params))
fig.tight_layout()
fig.subplots_adjust(wspace=0.05, hspace=0.05)
```
| github_jupyter |
# Multi-Layer Perceptron, MNIST
---
In this notebook, we will train an MLP to classify images from the [MNIST database](http://yann.lecun.com/exdb/mnist/) hand-written digit database.
The process will be broken down into the following steps:
>1. Load and visualize the data
2. Define a neural network
3. Train the model
4. Evaluate the performance of our trained model on a test dataset!
Before we begin, we have to import the necessary libraries for working with data and PyTorch.
```
# import libraries
import torch
import numpy as np
```
---
## Load and Visualize the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a few moments, and you should see your progress as the data is loading. You may also choose to change the `batch_size` if you want to load more data at a time.
This cell will create DataLoaders for each of our datasets.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to torch.FloatTensor
transform = transforms.ToTensor()
# choose the training and test datasets
train_data = datasets.MNIST(root='data', train=True,
download=True, transform=transform)
test_data = datasets.MNIST(root='data', train=False,
download=True, transform=transform)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
```
### Visualize a Batch of Training Data
The first step in a classification task is to take a look at the data, make sure it is loaded in correctly, then make any initial observations about patterns in that data.
```
import matplotlib.pyplot as plt
%matplotlib inline
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
```
### View an Image in More Detail
```
img = np.squeeze(images[1])
fig = plt.figure(figsize = (12,12))
ax = fig.add_subplot(111)
ax.imshow(img, cmap='gray')
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center',
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
The architecture will be responsible for seeing as input a 784-dim Tensor of pixel values for each image, and producing a Tensor of length 10 (our number of classes) that indicates the class scores for an input image. This particular example uses two hidden layers and dropout to avoid overfitting.
```
import torch.nn as nn
import torch.nn.functional as F
# define the NN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# number of hidden nodes in each layer (512)
hidden_1 = 512
hidden_2 = 512
# linear layer (784 -> hidden_1)
self.fc1 = nn.Linear(28 * 28, hidden_1)
# linear layer (n_hidden -> hidden_2)
self.fc2 = nn.Linear(hidden_1, hidden_2)
# linear layer (n_hidden -> 10)
self.fc3 = nn.Linear(hidden_2, 10)
# dropout layer (p=0.2)
# dropout prevents overfitting of data
self.dropout = nn.Dropout(0.2)
def forward(self, x):
# flatten image input
x = x.view(-1, 28 * 28)
# add hidden layer, with relu activation function
x = F.relu(self.fc1(x))
# add dropout layer
x = self.dropout(x)
# add hidden layer, with relu activation function
x = F.relu(self.fc2(x))
# add dropout layer
x = self.dropout(x)
# add output layer
x = self.fc3(x)
return x
# initialize the NN
model = Net()
print(model)
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
It's recommended that you use cross-entropy loss for classification. If you look at the documentation (linked above), you can see that PyTorch's cross entropy function applies a softmax funtion to the output layer *and* then calculates the log loss.
```
# specify loss function (categorical cross-entropy)
criterion = nn.CrossEntropyLoss()
# specify optimizer (stochastic gradient descent) and learning rate = 0.01
optimizer = torch.optim.SGD(model.parameters(), lr=0.01)
```
---
## Train the Network
The steps for training/learning from a batch of data are described in the comments below:
1. Clear the gradients of all optimized variables
2. Forward pass: compute predicted outputs by passing inputs to the model
3. Calculate the loss
4. Backward pass: compute gradient of the loss with respect to model parameters
5. Perform a single optimization step (parameter update)
6. Update average training loss
The following loop trains for 50 epochs; take a look at how the values for the training loss decrease over time. We want it to decrease while also avoiding overfitting the training data.
```
# number of epochs to train the model
n_epochs = 50
# initialize tracker for minimum validation loss
valid_loss_min = np.Inf # set initial "min" to infinity
for epoch in range(n_epochs):
# monitor training loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train() # prep model for training
for data, target in train_loader:
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update running training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval() # prep model for evaluation
for data, target in valid_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update running validation loss
valid_loss += loss.item()*data.size(0)
# print training/validation statistics
# calculate average loss over an epoch
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch+1,
train_loss,
valid_loss
))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
```
model.load_state_dict(torch.load('model.pt'))
```
---
## Test the Trained Network
Finally, we test our best model on previously unseen **test data** and evaluate it's performance. Testing on unseen data is a good way to check that our model generalizes well. It may also be useful to be granular in this analysis and take a look at how this model performs on each class as well as looking at its overall loss and accuracy.
```
# initialize lists to monitor test loss and accuracy
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval() # prep model for evaluation
for data, target in test_loader:
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct = np.squeeze(pred.eq(target.data.view_as(pred)))
# calculate test accuracy for each object class
for i in range(len(target)):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# calculate and print avg test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
str(i), 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Visualize Sample Test Results
This cell displays test images and their labels in this format: `predicted (ground-truth)`. The text will be green for accurately classified examples and red for incorrect predictions.
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds = torch.max(output, 1)
# prep images for display
images = images.numpy()
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, 20/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.squeeze(images[idx]), cmap='gray')
ax.set_title("{} ({})".format(str(preds[idx].item()), str(labels[idx].item())),
color=("green" if preds[idx]==labels[idx] else "red"))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/mashyko/Caffe2_Detectron2/blob/master/Pretrained_Models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
you need to read Model_Quickload.ipynb before run this notebook.
caffe2_tutorials is required to be uploaded in your Google Drive.
# Loading Pre-Trained Models
## Description
In this tutorial, we will use the pre-trained `squeezenet` model from the [ModelZoo](https://github.com/caffe2/caffe2/wiki/Model-Zoo) to classify our own images. As input, we will provide the path (or URL) to an image we want to classify. It will also be helpful to know the [ImageNet object code](https://gist.githubusercontent.com/aaronmarkham/cd3a6b6ac071eca6f7b4a6e40e6038aa/raw/9edb4038a37da6b5a44c3b5bc52e448ff09bfe5b/alexnet_codes) for the image so we can verify our results. The 'object code' is nothing more than the integer label for the class used during training, for example "985" is the code for the class "daisy". Note, although we are using squeezenet here, this tutorial serves as a somewhat universal method for running inference on pretrained models.
If you came from the [Image Pre-Processing Tutorial](https://caffe2.ai/docs/tutorial-image-pre-processing.html), you will see that we are using rescale and crop functions to prep the image, as well as reformatting the image to be CHW, BGR, and finally NCHW. We also correct for the image mean by either using the calculated mean from a provided npy file, or statically removing 128 as a placeholder average.
Hopefully, you will find that loading pre-trained models is simple and syntactically concise. From a high level, these are the three required steps for running inference on a pretrained model:
1. Read the init and predict protobuf (.pb) files of the pretrained model
with open("init_net.pb", "rb") as f:
init_net = f.read()
with open("predict_net.pb", "rb") as f:
predict_net = f.read()
2. Initialize a Predictor in your workspace with the blobs from the protobufs
p = workspace.Predictor(init_net, predict_net)
3. Run the net on some data and get the (softmax) results!
results = p.run({'data': img})
Note, assuming the last layer of the network is a softmax layer, the results come back as a multidimensional array of probabilities with length equal to the number of classes that the model was trained on. The probabilities may be indexed by the object code (integer type), so if you know the object code you can index the results array at that index to view the network's confidence that the input image is of that class.
**Model Download Options**
Although we will use `squeezenet` here, you can check out the [Model Zoo for pre-trained models](https://github.com/caffe2/caffe2/wiki/Model-Zoo) to browse/download a variety of pretrained models, or you can use Caffe2's `caffe2.python.models.download` module to easily acquire pre-trained models from [Github caffe2/models](http://github.com/caffe2/models).
For our purposes, we will use the `models.download` module to download `squeezenet` into the `/caffe2/python/models` folder of our local Caffe2 installation with the following command:
```
python -m caffe2.python.models.download -i squeezenet
```
If the above download worked then you should have a directory named squeezenet in your `/caffe2/python/models` folder that contains `init_net.pb` and `predict_net.pb`. Note, if you do not use the `-i` flag, the model will be downloaded to your CWD, however it will still be a directory named squeezenet containing two protobuf files. Alternatively, if you wish to download all of the models, you can clone the entire repo using:
```
git clone https://github.com/caffe2/models
```
## Code
Before we start, lets take care of the required imports.
```
!git clone --recursive https://github.com/caffe2/tutorials caffe2_tutorials
import os
os.chdir('caffe2_tutorials/')
!pip3 install torch torchvision
```
run the code below to download the pretrained model of squeezenet.
```
!python -m caffe2.python.models.download -i squeezenet
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __future__ import unicode_literals
%matplotlib inline
from caffe2.proto import caffe2_pb2
import numpy as np
import skimage.io
import skimage.transform
from matplotlib import pyplot
import os
from caffe2.python import core, workspace, models
import urllib2
import operator
print("Required modules imported.")
```
### Inputs
Here, we will specify the inputs to be used for this run, including the input image, the model location, the mean file (optional), the required size of the image, and the location of the label mapping file.
```
# Configuration --- Change to your setup and preferences!
# This directory should contain the models downloaded from the model zoo. To run this
# tutorial, make sure there is a 'squeezenet' directory at this location that
# contains both the 'init_net.pb' and 'predict_net.pb'
CAFFE_MODELS = 'caffe2/python/models'
# Some sample images you can try, or use any URL to a regular image.
# IMAGE_LOCATION = "https://upload.wikimedia.org/wikipedia/commons/thumb/f/f8/Whole-Lemon.jpg/1235px-Whole-Lemon.jpg"
# IMAGE_LOCATION = "https://upload.wikimedia.org/wikipedia/commons/7/7b/Orange-Whole-%26-Split.jpg"
# IMAGE_LOCATION = "https://upload.wikimedia.org/wikipedia/commons/a/ac/Pretzel.jpg"
# IMAGE_LOCATION = "https://cdn.pixabay.com/photo/2015/02/10/21/28/flower-631765_1280.jpg"
IMAGE_LOCATION = "images/flower.jpg"
# codes - these help decypher the output and source from a list from ImageNet's object codes
# to provide an result like "tabby cat" or "lemon" depending on what's in the picture
# you submit to the CNN.
codes = "https://gist.githubusercontent.com/aaronmarkham/cd3a6b6ac071eca6f7b4a6e40e6038aa/raw/9edb4038a37da6b5a44c3b5bc52e448ff09bfe5b/alexnet_codes"
print("Config set!")
```
### Image Preprocessing
Now that we have our inputs specified and verified the existance of the input network, we can load the image and pre-processing the image for ingestion into a Caffe2 convolutional neural network! This is a very important step as the trained CNN requires a specifically sized input image whose values are from a particular distribution.
```
# Function to crop the center cropX x cropY pixels from the input image
def crop_center(img,cropx,cropy):
y,x,c = img.shape
startx = x//2-(cropx//2)
starty = y//2-(cropy//2)
return img[starty:starty+cropy,startx:startx+cropx]
# Function to rescale the input image to the desired height and/or width. This function will preserve
# the aspect ratio of the original image while making the image the correct scale so we can retrieve
# a good center crop. This function is best used with center crop to resize any size input images into
# specific sized images that our model can use.
def rescale(img, input_height, input_width):
# Get original aspect ratio
aspect = img.shape[1]/float(img.shape[0])
if(aspect>1):
# landscape orientation - wide image
res = int(aspect * input_height)
imgScaled = skimage.transform.resize(img, (input_width, res))
if(aspect<1):
# portrait orientation - tall image
res = int(input_width/aspect)
imgScaled = skimage.transform.resize(img, (res, input_height))
if(aspect == 1):
imgScaled = skimage.transform.resize(img, (input_width, input_height))
return imgScaled
# Load the image as a 32-bit float
# Note: skimage.io.imread returns a HWC ordered RGB image of some size
INPUT_IMAGE_SIZE = 227
img = skimage.img_as_float(skimage.io.imread(IMAGE_LOCATION)).astype(np.float32)
print("Original Image Shape: " , img.shape)
# Rescale the image to comply with our desired input size. This will not make the image 227x227
# but it will make either the height or width 227 so we can get the ideal center crop.
img = rescale(img, INPUT_IMAGE_SIZE, INPUT_IMAGE_SIZE)
print("Image Shape after rescaling: " , img.shape)
pyplot.figure()
pyplot.imshow(img)
pyplot.title('Rescaled image')
# Crop the center 227x227 pixels of the image so we can feed it to our model
mean=128
img = crop_center(img, INPUT_IMAGE_SIZE, INPUT_IMAGE_SIZE)
print("Image Shape after cropping: " , img.shape)
pyplot.figure()
pyplot.imshow(img)
pyplot.title('Center Cropped')
# switch to CHW (HWC --> CHW)
img = img.swapaxes(1, 2).swapaxes(0, 1)
print("CHW Image Shape: " , img.shape)
pyplot.figure()
for i in range(3):
# For some reason, pyplot subplot follows Matlab's indexing
# convention (starting with 1). Well, we'll just follow it...
pyplot.subplot(1, 3, i+1)
pyplot.imshow(img[i])
pyplot.axis('off')
pyplot.title('RGB channel %d' % (i+1))
# switch to BGR (RGB --> BGR)
img = img[(2, 1, 0), :, :]
# remove mean for better results
img = img * 255 - mean
# add batch size axis which completes the formation of the NCHW shaped input that we want
img = img[np.newaxis, :, :, :].astype(np.float32)
print("NCHW image (ready to be used as input): ", img.shape)
```
### Prepare the CNN and run the net!
Now that the image is ready to be ingested by the CNN, let's open the protobufs, load them into the workspace, and run the net.
```
# when squeezenet is to be used.
from caffe2.python.models import squeezenet as mynet
init_net = mynet.init_net
predict_net = mynet.predict_net
# Initialize the predictor from the input protobufs
p = workspace.Predictor(init_net, predict_net)
# Run the net and return prediction
results = p.run({'data': img})
# Turn it into something we can play with and examine which is in a multi-dimensional array
results = np.asarray(results)
print("results shape: ", results.shape)
# Quick way to get the top-1 prediction result
# Squeeze out the unnecessary axis. This returns a 1-D array of length 1000
preds = np.squeeze(results)
# Get the prediction and the confidence by finding the maximum value and index of maximum value in preds array
curr_pred, curr_conf = max(enumerate(preds), key=operator.itemgetter(1))
print("Prediction: ", curr_pred)
print("Confidence: ", curr_conf)
```
### Process Results
Recall ImageNet is a 1000 class dataset and observe that it is no coincidence that the third axis of results is length 1000. This axis is holding the probability for each category in the pre-trained model. So when you look at the results array at a specific index, the number can be interpreted as the probability that the input belongs to the class corresponding to that index. Now that we have run the predictor and collected the results, we can interpret them by matching them to their corresponding english labels.
```
# the rest of this is digging through the results
results = np.delete(results, 1)
index = 0
highest = 0
arr = np.empty((0,2), dtype=object)
arr[:,0] = int(10)
arr[:,1:] = float(10)
for i, r in enumerate(results):
# imagenet index begins with 1!
i=i+1
arr = np.append(arr, np.array([[i,r]]), axis=0)
if (r > highest):
highest = r
index = i
# top N results
N = 5
topN = sorted(arr, key=lambda x: x[1], reverse=True)[:N]
print("Raw top {} results: {}".format(N,topN))
# Isolate the indexes of the top-N most likely classes
topN_inds = [int(x[0]) for x in topN]
print("Top {} classes in order: {}".format(N,topN_inds))
# Now we can grab the code list and create a class Look Up Table
response = urllib2.urlopen(codes)
class_LUT = []
for line in response:
code, result = line.partition(":")[::2]
code = code.strip()
result = result.replace("'", "")
if code.isdigit():
class_LUT.append(result.split(",")[0][1:])
# For each of the top-N results, associate the integer result with an actual class
for n in topN:
print("Model predicts '{}' with {}% confidence".format(class_LUT[int(n[0])],float("{0:.2f}".format(n[1]*100))))
```
### Feeding Larger Batches
Above is an example of how to feed one image at a time. We can achieve higher throughput if we feed multiple images at a time in a single batch. Recall, the data fed into the classifier is in 'NCHW' order, so to feed multiple images, we will expand the 'N' axis.
```
# List of input images to be fed
images = ["images/cowboy-hat.jpg",
"images/cell-tower.jpg",
"images/Ducreux.jpg",
"images/pretzel.jpg",
"images/orangutan.jpg",
"images/aircraft-carrier.jpg",
"images/cat.jpg"]
# Allocate space for the batch of formatted images
NCHW_batch = np.zeros((len(images),3,227,227))
print ("Batch Shape: ",NCHW_batch.shape)
# For each of the images in the list, format it and place it in the batch
for i,curr_img in enumerate(images):
img = skimage.img_as_float(skimage.io.imread(curr_img)).astype(np.float32)
img = rescale(img, 227, 227)
img = crop_center(img, 227, 227)
img = img.swapaxes(1, 2).swapaxes(0, 1)
img = img[(2, 1, 0), :, :]
img = img * 255 - mean
NCHW_batch[i] = img
print("NCHW image (ready to be used as input): ", NCHW_batch.shape)
# Run the net on the batch
results = p.run([NCHW_batch.astype(np.float32)])
# Turn it into something we can play with and examine which is in a multi-dimensional array
results = np.asarray(results)
# Squeeze out the unnecessary axis
preds = np.squeeze(results)
print("Squeezed Predictions Shape, with batch size {}: {}".format(len(images),preds.shape))
# Describe the results
for i,pred in enumerate(preds):
print("Results for: '{}'".format(images[i]))
# Get the prediction and the confidence by finding the maximum value
# and index of maximum value in preds array
curr_pred, curr_conf = max(enumerate(pred), key=operator.itemgetter(1))
print("\tPrediction: ", curr_pred)
print("\tClass Name: ", class_LUT[int(curr_pred)])
print("\tConfidence: ", curr_conf)
```
以上
| github_jupyter |
### Imports
```
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from sklearn.model_selection import train_test_split
import pandas as pd
import random
```
### Load & Prep Data
The <a href='https://homepages.inf.ed.ac.uk/imurray2/teaching/oranges_and_lemons/'>Fruits Dataset</a> was originally created by Dr. Iain Murray from University of Edinburgh and extended more recently by the University of Michigan. It is a simple multi-class dataset with 4 columns (features) and 4 classes (fruits). The 4 classes are apple, orange, mandarin and lemon. The four features are mass, width, height and color score of the fruit.
The color score feature maps to a color and its intensity in the color spectrum (0 - 1) scale. <br><br>
<table align="left" style="width:50%">
<tr>
<th>Color</th>
<th>Range</th>
</tr>
<tr>
<td>Red</td>
<td>0.85 - 1.00</td>
</tr>
<tr>
<td>Orange</td>
<td>0.75 - 0.85</td>
</tr>
<tr>
<td>Yellow</td>
<td>0.65 - 0.75</td>
</tr>
<tr>
<td>Green</td>
<td>0.45 - 0.65</td>
</tr>
</table>
```
df = pd.read_csv('./DATA/fruits.csv', names=['class', 'mass', 'width', 'height', 'color_score'])
df.head()
df.shape
df['class'].unique().tolist()
X = df[['mass', 'width', 'height', 'color_score']]
y = df['class']
```
#### Encode the classes into numerical values using Sklearn's LabelEncoder
```
label_encoder = LabelEncoder()
label_encoder.fit(['apple', 'orange', 'mandarin', 'lemon'])
y = label_encoder.transform(y)
y
```
#### Split X, y into train and test sets
```
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=123)
X_train.shape
X_test.shape
```
#### Scale feature columns using Sklearn's MinMaxScaler
```
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
X_train[0]
y_train[0]
```
#### Combine Scaled X & y into Train and Test DataFrames
```
X_train = pd.DataFrame(X_train, columns=['mass', 'width', 'height', 'color_score'])
y_train = pd.DataFrame(y_train, columns=['class'])
train_df = pd.concat([y_train, X_train], axis=1)
train_df.head()
X_test = pd.DataFrame(X_test, columns=['mass', 'width', 'height', 'color_score'])
y_test = pd.DataFrame(y_test, columns=['class'])
test_df = pd.concat([y_test, X_test], axis=1)
test_df.head()
```
#### Create a DataFrame for Batch Inference without the Class column
```
batch_test_df = test_df.drop(['class'], axis=1)
batch_test_df.head()
```
#### Write Train & Test Sets to Local Directory
```
train_df.to_csv('./DATA/train/train.csv', header=False, index=False)
test_df.to_csv('./DATA/test/test.csv', header=False, index=False)
batch_test_df.to_csv('./DATA/batch_test/batch_test.csv', header=False, index=False)
```
<b>Write train_df without class label and with header for Model Monitoring Baselining</b>
```
train_df.to_csv('./DATA/train/train_with_header.csv', header=True, index=False)
```
### Let us simulate some artificial data for Model Monitor for our Data Shift experiments later
```
def get_random_val():
val = random.uniform(0, 1)
return round(val, 6)
def generate_row():
row = []
for _ in range(4):
row.append(get_random_val())
return row
def generate_dataset():
rows = []
for _ in range(20):
rows.append(generate_row())
return rows
rows = generate_dataset()
df = pd.DataFrame(rows, columns=['mass', 'width', 'height', 'color_score'])
df.head()
df.to_csv('./DATA/test/model_monitor_test.csv', header=False, index=False)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/lukereichold/3D-MNIST-S4TF/blob/master/3D_MNIST_Classifier_with_S4TF.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## ⚙️ Install Dependencies
```
import TensorFlow
import Foundation
#if canImport(PythonKit)
import PythonKit
#else
import Python
#endif
print(Python.version)
let h5py = Python.import("h5py")
let np = Python.import("numpy")
let os = Python.import("os")
let plt = Python.import("matplotlib.pyplot")
let cm = Python.import("matplotlib.cm")
%include "EnableIPythonDisplay.swift"
IPythonDisplay.shell.enable_matplotlib("inline")
```
## ⬇ Fetch Dataset
3D Point Vectors fetched initially from https://www.kaggle.com/daavoo/3d-mnist/data (29 Mb)
```
os.system("wget https://www.dropbox.com/s/mvre9rojnjp2a36/full_dataset_vectors.h5")
let dataset = h5py.File("full_dataset_vectors.h5", "r")
func floatTensor(from data: PythonObject) -> Tensor<Float> {
return Tensor<Float>(numpy: data.value.astype(np.float32))!
}
func intTensor(from data: PythonObject) -> Tensor<Int32> {
return Tensor<Int32>(numpy: data.value.astype(np.int32))!
}
var trainingFeatures = floatTensor(from: dataset["X_train"])
var trainingLabels = intTensor(from: dataset["y_train"])
var testFeatures = floatTensor(from: dataset["X_test"])
var testLabels = intTensor(from: dataset["y_test"])
```
## 📐 Verify Initial Dataset Shapes
```
assert(trainingFeatures.shape == TensorShape([10_000, 4096]))
assert(trainingLabels.shape == TensorShape([10_000]))
assert(testFeatures.shape == TensorShape([2_000, 4096]))
assert(testLabels.shape == TensorShape([2_000]))
let trainingSize = trainingFeatures.shape[0]
let testSize = testFeatures.shape[0]
```
## 🌈 Give Each Point a Color
Each sample represents an "MNIST digit" in three-dimensional space (x, y, z coordinates), where each of the 4096 possible points has a color.
```
var coloredTrainingSamples = Tensor<Float>(ones: TensorShape([10_000, 4096, 3]))
var coloredTestSamples = Tensor<Float>(ones: TensorShape([2_000, 4096, 3]))
let trainingNp = trainingFeatures.makeNumpyArray()
let testNp = testFeatures.makeNumpyArray()
let scalarMap = cm.ScalarMappable(cmap: "Oranges")
func addColorDimension(scalars: PythonObject) -> Tensor<Float> {
let rgba = scalarMap.to_rgba(scalars).astype(np.float32)
let colorValues = Tensor<Float>(numpy: rgba)!
return colorValues[TensorRange.ellipsis, ..<3]
}
for i in 0 ..< trainingSize {
coloredTrainingSamples[i] = addColorDimension(scalars: trainingNp[i])
}
for i in 0 ..< testSize {
coloredTestSamples[i] = addColorDimension(scalars: testNp[i])
}
```
## 🔲 Expand Points to Three Dimensions
Reshape each sample to be represented as a rank-4 tensor of shape `(16, 16, 16, 3)`.
```
coloredTrainingSamples = coloredTrainingSamples.reshaped(to: TensorShape(trainingSize, 16, 16, 16, 3))
coloredTestSamples = coloredTestSamples.reshaped(to: TensorShape(testSize, 16, 16, 16, 3))
let coloredTrainingSamplesNp = coloredTrainingSamples.makeNumpyArray()
let trainingLabelsNp = trainingLabels.makeNumpyArray()
print(coloredTrainingSamples.shape)
print(coloredTestSamples.shape)
```
## 📚 Defining Our Network
```
let frame_height = 16, frame_width = 16, frame_depth = 16
let RGB_channels = 3
let classCount = 10
let input_shape = (frame_depth, frame_height, frame_width, RGB_channels)
struct Classifier: Layer {
typealias Input = Tensor<Float>
typealias Output = Tensor<Float>
var conv1 = Conv3D<Float>(filterShape: (3, 3, 3, RGB_channels, 8), padding: .same, activation: relu)
var conv2 = Conv3D<Float>(filterShape: (3, 3, 3, 8, 16), padding: .same, activation: relu)
var pool = MaxPool3D<Float>(poolSize: (2, 2, 2), strides: (2, 2, 2))
var conv3 = Conv3D<Float>(filterShape: (3, 3, 3, 16, 32), padding: .same, activation: relu)
var conv4 = Conv3D<Float>(filterShape: (3, 3, 3, 32, 64), padding: .same, activation: relu)
var batchNorm = BatchNorm<Float>(featureCount: 64)
var flatten = Flatten<Float>()
var dense1 = Dense<Float>(inputSize: 4096, outputSize: 4096, activation: relu)
var dense2 = Dense<Float>(inputSize: 4096, outputSize: 1024, activation: relu)
var dropout25 = Dropout<Float>(probability: 0.25)
var dropout50 = Dropout<Float>(probability: 0.5)
var output = Dense<Float>(inputSize: 1024, outputSize: classCount, activation: softmax)
@differentiable
func callAsFunction(_ input: Input) -> Output {
return input
.sequenced(through: conv1, conv2, pool)
.sequenced(through: conv3, conv4, batchNorm, pool)
.sequenced(through: dropout25, flatten, dense1, dropout50)
.sequenced(through: dense2, dropout50, output)
}
}
var model = Classifier()
let dummy = Tensor<Float>(randomNormal: TensorShape(1, 16, 16, 16, 3))
let eval = model(dummy)
assert(eval.shape == TensorShape([1, 10]))
print(eval.shape)
```
## 📈 Training
```
let batchSize = 100
let epochs = 50
let optimizer = Adam(for: model, learningRate: 1e-5, decay: 1e-6)
var trainAccHistory = np.zeros(epochs)
var valAccHistory = np.zeros(epochs)
var trainLossHistory = np.zeros(epochs)
var valLossHistory = np.zeros(epochs)
for epoch in 0 ..< epochs {
Context.local.learningPhase = .training
// Shuffle samples
let shuffledIndices = np.random.permutation(trainingSize)
let shuffledSamples = coloredTrainingSamplesNp[shuffledIndices]
let shuffledLabels = trainingLabelsNp[shuffledIndices]
// Loop over each batch of samples
for batchStart in stride(from: 0, to: trainingSize, by: batchSize) {
let batchRange = batchStart ..< batchStart + batchSize
let labels = Tensor<Int32>(numpy: shuffledLabels[batchRange])!
let samples = Tensor<Float>(numpy: shuffledSamples[batchRange])!
let (_, gradients) = valueWithGradient(at: model) { model -> Tensor<Float> in
let logits = model(samples)
return softmaxCrossEntropy(logits: logits, labels: labels)
}
optimizer.update(&model, along: gradients)
}
// Evaluate model
Context.local.learningPhase = .inference
var correctTrainGuessCount = 0
var totalTrainGuessCount = 0
for batchStart in stride(from: 0, to: trainingSize, by: batchSize) {
let batchRange = batchStart ..< batchStart + batchSize
let labels = Tensor<Int32>(numpy: shuffledLabels[batchRange])!
let samples = Tensor<Float>(numpy: shuffledSamples[batchRange])!
let logits = model(samples)
// accuracy
let correctPredictions = logits.argmax(squeezingAxis: 1) .== labels
correctTrainGuessCount += Int(Tensor<Int32>(correctPredictions).sum().scalarized())
// loss
trainLossHistory[epoch] += PythonObject(softmaxCrossEntropy(logits: logits, labels: labels).scalarized())
}
let trainAcc = Float(correctTrainGuessCount) / Float(trainingSize)
trainAccHistory[epoch] = PythonObject(trainAcc)
var correctValGuessCount = 0
var totalValGuessCount = 0
for batchStart in stride(from: 0, to: testSize, by: batchSize) {
let batchRange = batchStart ..< batchStart + batchSize
let labels = testLabels[batchRange]
let samples = coloredTestSamples[batchRange]
let logits = model(samples)
// accuracy
let correctPredictions = logits.argmax(squeezingAxis: 1) .== labels
correctValGuessCount += Int(Tensor<Int32>(correctPredictions).sum().scalarized())
// loss
valLossHistory[epoch] += PythonObject(softmaxCrossEntropy(logits: logits, labels: labels).scalarized())
}
let valAcc = Float(correctValGuessCount) / Float(testSize)
valAccHistory[epoch] = PythonObject(valAcc)
print("\(epoch + 1) | Training accuracy: \(trainAcc) | Validation accuracy: \(valAcc)")
}
```
## 🕝 Inspecting training history
```
plt.plot(trainAccHistory, label: "Training Accuracy")
plt.plot(valAccHistory, label: "Validation Accuracy")
plt.xlabel("Number of epochs")
plt.legend()
plt.title("Accuracy")
plt.show()
plt.plot(trainLossHistory, label: "Training Set")
plt.plot(valLossHistory, label: "Validation Set")
plt.xlabel("Number of epochs")
plt.legend()
plt.title("Loss")
plt.show()
```
## 🌐 Making Inferences
With our model now trained, we can make inferences to predict the class labels of samples outside the training set.
```
let randomIndex = Int.random(in: 0 ..< testSize)
var randomSample = coloredTestSamples[randomIndex].reshaped(to: TensorShape(1, 16, 16, 16, 3))
let label = testLabels[randomIndex]
print("Predicted: \(model(randomSample).argmax()), Actual: \(label)")
```
| github_jupyter |
# Setup
```
#Save Checkpoints after each round of active learning
store_checkpoint=True
#Mount persistent storage for logs and checkpoints (Drive)
persistent=False
#Load initial model.
'''
Since there is a need to compare all strategies with same initial model,
the base model only needs to be trained once.
True: Will load the model from the model directory configured in section Initial Training
and Parameter Definitions
False: Will train a base model and store it in model directory configured in section Initial Training
and Parameter Definitions
'''
load_model = False
```
**Installations**
```
!pip install apricot-select
!git clone https://github.com/decile-team/distil.git
!git clone https://github.com/circulosmeos/gdown.pl.git
!mv distil asdf
!mv asdf/distil .
```
**Imports, Training Class Definition, Experiment Procedure Definition**
```
import pandas as pd
import numpy as np
import copy
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torch.utils.data import Subset
import torch.nn.functional as F
from torch import nn
from torchvision import transforms
from torchvision import datasets
from PIL import Image
import torch
import torch.optim as optim
from torch.autograd import Variable
import sys
sys.path.append('../')
import matplotlib.pyplot as plt
import time
import math
import random
import os
import pickle
from numpy.linalg import cond
from numpy.linalg import inv
from numpy.linalg import norm
from scipy import sparse as sp
from scipy.linalg import lstsq
from scipy.linalg import solve
from scipy.optimize import nnls
from distil.active_learning_strategies.badge import BADGE
from distil.active_learning_strategies.glister import GLISTER
from distil.active_learning_strategies.margin_sampling import MarginSampling
from distil.active_learning_strategies.entropy_sampling import EntropySampling
from distil.active_learning_strategies.random_sampling import RandomSampling
from distil.active_learning_strategies.fass import FASS
from distil.active_learning_strategies.core_set import CoreSet
from distil.active_learning_strategies.least_confidence import LeastConfidence
from distil.utils.models.resnet import ResNet18
from distil.utils.data_handler import DataHandler_MNIST, DataHandler_CIFAR10, DataHandler_Points, DataHandler_FASHION_MNIST, DataHandler_SVHN
from distil.utils.dataset import get_dataset
from distil.utils.train_helper import data_train
from google.colab import drive
import warnings
warnings.filterwarnings("ignore")
class Checkpoint:
def __init__(self, acc_list=None, indices=None, state_dict=None, experiment_name=None, path=None):
# If a path is supplied, load a checkpoint from there.
if path is not None:
if experiment_name is not None:
self.load_checkpoint(path, experiment_name)
else:
raise ValueError("Checkpoint contains None value for experiment_name")
return
if acc_list is None:
raise ValueError("Checkpoint contains None value for acc_list")
if indices is None:
raise ValueError("Checkpoint contains None value for indices")
if state_dict is None:
raise ValueError("Checkpoint contains None value for state_dict")
if experiment_name is None:
raise ValueError("Checkpoint contains None value for experiment_name")
self.acc_list = acc_list
self.indices = indices
self.state_dict = state_dict
self.experiment_name = experiment_name
def __eq__(self, other):
# Check if the accuracy lists are equal
acc_lists_equal = self.acc_list == other.acc_list
# Check if the indices are equal
indices_equal = self.indices == other.indices
# Check if the experiment names are equal
experiment_names_equal = self.experiment_name == other.experiment_name
return acc_lists_equal and indices_equal and experiment_names_equal
def save_checkpoint(self, path):
# Get current time to use in file timestamp
timestamp = time.time_ns()
# Create the path supplied
os.makedirs(path, exist_ok=True)
# Name saved files using timestamp to add recency information
save_path = os.path.join(path, F"c{timestamp}1")
copy_save_path = os.path.join(path, F"c{timestamp}2")
# Write this checkpoint to the first save location
with open(save_path, 'wb') as save_file:
pickle.dump(self, save_file)
# Write this checkpoint to the second save location
with open(copy_save_path, 'wb') as copy_save_file:
pickle.dump(self, copy_save_file)
def load_checkpoint(self, path, experiment_name):
# Obtain a list of all files present at the path
timestamp_save_no = [f for f in os.listdir(path) if os.path.isfile(os.path.join(path, f))]
# If there are no such files, set values to None and return
if len(timestamp_save_no) == 0:
self.acc_list = None
self.indices = None
self.state_dict = None
return
# Sort the list of strings to get the most recent
timestamp_save_no.sort(reverse=True)
# Read in two files at a time, checking if they are equal to one another.
# If they are equal, then it means that the save operation finished correctly.
# If they are not, then it means that the save operation failed (could not be
# done atomically). Repeat this action until no possible pair can exist.
while len(timestamp_save_no) > 1:
# Pop a most recent checkpoint copy
first_file = timestamp_save_no.pop(0)
# Keep popping until two copies with equal timestamps are present
while True:
second_file = timestamp_save_no.pop(0)
# Timestamps match if the removal of the "1" or "2" results in equal numbers
if (second_file[:-1]) == (first_file[:-1]):
break
else:
first_file = second_file
# If there are no more checkpoints to examine, set to None and return
if len(timestamp_save_no) == 0:
self.acc_list = None
self.indices = None
self.state_dict = None
return
# Form the paths to the files
load_path = os.path.join(path, first_file)
copy_load_path = os.path.join(path, second_file)
# Load the two checkpoints
with open(load_path, 'rb') as load_file:
checkpoint = pickle.load(load_file)
with open(copy_load_path, 'rb') as copy_load_file:
checkpoint_copy = pickle.load(copy_load_file)
# Do not check this experiment if it is not the one we need to restore
if checkpoint.experiment_name != experiment_name:
continue
# Check if they are equal
if checkpoint == checkpoint_copy:
# This checkpoint will suffice. Populate this checkpoint's fields
# with the selected checkpoint's fields.
self.acc_list = checkpoint.acc_list
self.indices = checkpoint.indices
self.state_dict = checkpoint.state_dict
return
# Instantiate None values in acc_list, indices, and model
self.acc_list = None
self.indices = None
self.state_dict = None
def get_saved_values(self):
return (self.acc_list, self.indices, self.state_dict)
def delete_checkpoints(checkpoint_directory, experiment_name):
# Iteratively go through each checkpoint, deleting those whose experiment name matches.
timestamp_save_no = [f for f in os.listdir(checkpoint_directory) if os.path.isfile(os.path.join(checkpoint_directory, f))]
for file in timestamp_save_no:
delete_file = False
# Get file location
file_path = os.path.join(checkpoint_directory, file)
# Unpickle the checkpoint and see if its experiment name matches
with open(file_path, "rb") as load_file:
checkpoint_copy = pickle.load(load_file)
if checkpoint_copy.experiment_name == experiment_name:
delete_file = True
# Delete this file only if the experiment name matched
if delete_file:
os.remove(file_path)
#Logs
def write_logs(logs, save_directory, rd, run):
file_path = save_directory + 'run_'+str(run)+'.txt'
with open(file_path, 'a') as f:
f.write('---------------------\n')
f.write('Round '+str(rd)+'\n')
f.write('---------------------\n')
for key, val in logs.items():
if key == 'Training':
f.write(str(key)+ '\n')
for epoch in val:
f.write(str(epoch)+'\n')
else:
f.write(str(key) + ' - '+ str(val) +'\n')
def train_one(X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, strategy, save_directory, run, checkpoint_directory, experiment_name):
# Define acc initially
acc = np.zeros(n_rounds)
initial_unlabeled_size = X_unlabeled.shape[0]
initial_round = 1
# Define an index map
index_map = np.array([x for x in range(initial_unlabeled_size)])
# Attempt to load a checkpoint. If one exists, then the experiment crashed.
training_checkpoint = Checkpoint(experiment_name=experiment_name, path=checkpoint_directory)
rec_acc, rec_indices, rec_state_dict = training_checkpoint.get_saved_values()
# Check if there are values to recover
if rec_acc is not None:
# Restore the accuracy list
for i in range(len(rec_acc)):
acc[i] = rec_acc[i]
print('Loaded from checkpoint....')
print('Accuracy List:', acc)
# Restore the indices list and shift those unlabeled points to the labeled set.
index_map = np.delete(index_map, rec_indices)
# Record initial size of X_tr
intial_seed_size = X_tr.shape[0]
X_tr = np.concatenate((X_tr, X_unlabeled[rec_indices]), axis=0)
X_unlabeled = np.delete(X_unlabeled, rec_indices, axis = 0)
y_tr = np.concatenate((y_tr, y_unlabeled[rec_indices]), axis = 0)
y_unlabeled = np.delete(y_unlabeled, rec_indices, axis = 0)
# Restore the model
net.load_state_dict(rec_state_dict)
# Fix the initial round
initial_round = (X_tr.shape[0] - initial_seed_size) // budget + 1
# Ensure loaded model is moved to GPU
if torch.cuda.is_available():
net = net.cuda()
strategy.update_model(net)
strategy.update_data(X_tr, y_tr, X_unlabeled)
else:
if torch.cuda.is_available():
net = net.cuda()
acc[0] = dt.get_acc_on_set(X_test, y_test)
print('Initial Testing accuracy:', round(acc[0]*100, 2), flush=True)
logs = {}
logs['Training Points'] = X_tr.shape[0]
logs['Test Accuracy'] = str(round(acc[0]*100, 2))
write_logs(logs, save_directory, 0, run)
#Updating the trained model in strategy class
strategy.update_model(net)
##User Controlled Loop
for rd in range(initial_round, n_rounds):
print('-------------------------------------------------')
print('Round', rd)
print('-------------------------------------------------')
sel_time = time.time()
idx = strategy.select(budget)
sel_time = time.time() - sel_time
print("Selection Time:", sel_time)
#Saving state of model, since labeling new points might take time
# strategy.save_state()
#Adding new points to training set
X_tr = np.concatenate((X_tr, X_unlabeled[idx]), axis=0)
X_unlabeled = np.delete(X_unlabeled, idx, axis = 0)
#Human In Loop, Assuming user adds new labels here
y_tr = np.concatenate((y_tr, y_unlabeled[idx]), axis = 0)
y_unlabeled = np.delete(y_unlabeled, idx, axis = 0)
# Update the index map
index_map = np.delete(index_map, idx, axis = 0)
print('Number of training points -',X_tr.shape[0])
#Reload state and start training
# strategy.load_state()
strategy.update_data(X_tr, y_tr, X_unlabeled)
dt.update_data(X_tr, y_tr)
t1 = time.time()
clf, train_logs = dt.train(None)
t2 = time.time()
acc[rd] = dt.get_acc_on_set(X_test, y_test)
logs = {}
logs['Training Points'] = X_tr.shape[0]
logs['Test Accuracy'] = str(round(acc[rd]*100, 2))
logs['Selection Time'] = str(sel_time)
logs['Trainining Time'] = str(t2 - t1)
logs['Training'] = train_logs
write_logs(logs, save_directory, rd, run)
strategy.update_model(clf)
print('Testing accuracy:', round(acc[rd]*100, 2), flush=True)
# Create a checkpoint
used_indices = np.array([x for x in range(initial_unlabeled_size)])
used_indices = np.delete(used_indices, index_map).tolist()
if store_checkpoint:
round_checkpoint = Checkpoint(acc.tolist(), used_indices, clf.state_dict(), experiment_name=experiment_name)
round_checkpoint.save_checkpoint(checkpoint_directory)
print('Training Completed')
return acc
# Define a function to perform experiments in bulk and return the mean accuracies
def BADGE_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = BADGE(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def random_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = RandomSampling(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def entropy_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = EntropySampling(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def GLISTER_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size'], 'lr': args['lr']}
strategy = GLISTER(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args,valid=False, typeOf='rand', lam=0.1)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def FASS_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = FASS(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def adversarial_bim_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = AdversarialBIM(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def adversarial_deepfool_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = AdversarialDeepFool(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def coreset_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = CoreSet(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def least_confidence_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = LeastConfidence(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def margin_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = MarginSampling(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
# Define a function to perform experiments in bulk and return the mean accuracies
def bald_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, net, n_rounds, budget, args, nclasses, save_directory, checkpoint_directory, experiment_name):
test_acc_list = list()
fig = plt.figure(figsize=(8,6), dpi=160)
x_axis = [np.shape(X_tr)[0] + budget * x for x in range(0, n_rounds)]
for i in range(n_exp):
# Copy data and model to ensure that experiments do not override base versions
X_tr_copy = copy.deepcopy(X_tr)
y_tr_copy = copy.deepcopy(y_tr)
X_unlabeled_copy = copy.deepcopy(X_unlabeled)
y_unlabeled_copy = copy.deepcopy(y_unlabeled)
X_test_copy = copy.deepcopy(X_test)
y_test_copy = copy.deepcopy(y_test)
dt_copy = copy.deepcopy(dt)
clf_copy = copy.deepcopy(net)
#Initializing Strategy Class
strategy_args = {'batch_size' : args['batch_size']}
strategy = BALDDropout(X_tr, y_tr, X_unlabeled, net, handler, nclasses, strategy_args)
test_acc = train_one(X_tr_copy, y_tr_copy, X_test_copy, y_test_copy, X_unlabeled_copy, y_unlabeled_copy, dt_copy, clf_copy, n_rounds, budget, args, nclasses, strategy, save_directory, i, checkpoint_directory, experiment_name)
test_acc_list.append(test_acc)
plt.plot(x_axis, test_acc, label=str(i))
print("EXPERIMENT", i, test_acc)
# Experiment complete; delete all checkpoints related to this experiment
delete_checkpoints(checkpoint_directory, experiment_name)
mean_test_acc = np.zeros(n_rounds)
for test_acc in test_acc_list:
mean_test_acc = mean_test_acc + test_acc
mean_test_acc = mean_test_acc / n_exp
plt.plot(x_axis, mean_test_acc, label="Mean")
plt.xlabel("Labeled Set Size")
plt.ylabel("Test Acc")
plt.legend()
plt.show()
print("MEAN TEST ACC", mean_test_acc)
return mean_test_acc
```
# CIFAR10
**Initial Training and Parameter Definitions**
```
data_set_name = 'CIFAR10'
download_path = '../downloaded_data/'
handler = DataHandler_CIFAR10
net = ResNet18()
# Mount drive containing possible saved model and define file path
if persistent:
drive.mount('/content/drive')
# Retrieve the model from link and save it to the drive
logs_directory = '/content/drive/MyDrive/experiments/cifar10/'
# initial_model = data_set_name
model_directory = "/content/drive/MyDrive/experiments/cifar10/"
os.makedirs(model_directory, exist_ok = True)
model_directory = "/content/drive/MyDrive/experiments/cifar10/base_model.pth"
X, y, X_test, y_test = get_dataset(data_set_name, download_path)
dim = np.shape(X)[1:]
initial_seed_size = 1000
training_size_cap = 25000
print(X.shape, y.shape, X_test.shape, y_test.shape, np.unique(y))
y = y.numpy()
y_test = y_test.numpy()
X_tr = X[:initial_seed_size]
y_tr = y[:initial_seed_size]
X_unlabeled = X[initial_seed_size:]
y_unlabeled = y[initial_seed_size:]
nclasses = 10
budget = 3000
#Initial Training
args = {'n_epoch':300, 'lr':float(0.01), 'batch_size':20, 'max_accuracy':float(0.99), 'num_classes':nclasses, 'islogs':True, 'isreset':True, 'isverbose':True}
# Only train a new model if one does not exist.
if load_model:
net.load_state_dict(torch.load(model_directory))
dt = data_train(X_tr, y_tr, net, handler, args)
clf = net
else:
dt = data_train(X_tr, y_tr, net, handler, args)
clf, train_logs = dt.train(None)
torch.save(clf.state_dict(), model_directory)
# Train on approximately the full dataset given the budget contraints
n_rounds = math.floor(training_size_cap / budget)
n_exp = 1
print("Training for", n_rounds, "rounds with budget", budget, "on unlabeled set size", training_size_cap)
```
**Random Sampling**
```
strat_logs = logs_directory+'random_sampling/'
os.makedirs(strat_logs, exist_ok = True)
checkpoint_directory = '/content/drive/MyDrive/experiments/cifar10/random_sampling/check/'
os.makedirs(checkpoint_directory, exist_ok = True)
mean_test_acc_random = random_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, clf, n_rounds, budget, args, nclasses, strat_logs, checkpoint_directory, "cf_random")
```
**Entropy (Uncertainty) Sampling**
```
strat_logs = logs_directory+'entropy_sampling/'
os.makedirs(strat_logs, exist_ok = True)
checkpoint_directory = '/content/drive/MyDrive/experiments/cifar10/entropy_sampling/check/'
os.makedirs(checkpoint_directory, exist_ok = True)
mean_test_acc_entropy = entropy_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, clf, n_rounds, budget, args, nclasses, strat_logs, checkpoint_directory, "cf_entropy")
```
**BADGE**
```
strat_logs = logs_directory+'badge/'
os.makedirs(strat_logs, exist_ok = True)
checkpoint_directory = '/content/drive/MyDrive/experiments/cifar10/badge/check/'
os.makedirs(checkpoint_directory, exist_ok = True)
mean_test_acc_badge = BADGE_experiment_batch(n_exp, X_tr, y_tr, X_test, y_test, X_unlabeled, y_unlabeled, dt, clf, n_rounds, budget, args, nclasses, strat_logs, checkpoint_directory, "cf_badge")
```
| github_jupyter |
```
import gzip, json
import numpy as np
import matplotlib.pyplot as plt
def combine_results(*jsons):
jsons0 = []
for js in zip(*jsons):
j0 = {}
for j in js:
for k, v in j.items():
j0[k] = v
jsons0.append(j0)
return jsons0
def parse_file(filename):
with gzip.open(filename, "rb") as f:
lines = f.readlines()
return [json.loads(line) for line in lines]
jsons1101_2s = parse_file("../results/r1-mnist-time2-100:200-1101.gz") # Merge's results are invalid in this one
#jsons0100_2s = parse_file("../results/r1-mnist-time2-100:200-0100.gz")
#jsons1101_2s = combine_results(jsons1101_2s, jsons0100_2s)
jsons1100_4s = parse_file("../results/r1-mnist-time4-100:200-1100.gz")
jsons1100_6s = parse_file("../results/r1-mnist-time6-100:200-1100.gz")
jsons1100_8s = parse_file("../results/r1-mnist-time8-100:200-1100.gz")
jsons1100_10s = parse_file("../results/r1-mnist-time10-100:200-1100.gz")
jsons0e00 = parse_file("../results/r1-mnist-100:200-0e00.gz") # external merge (clique size 2, merge level 2, anything more is OOM)
jsons0e00_start40 = parse_file("../results/rob_start40_200-depth8-100:200-0e00.gz") # external merge (clique size 2, merge level 2, anything more is OOM)
kan_deltas = np.array([j["kantchelian_delta"] for j in jsons1101_2s])
kan_times = np.array([j["kantchelian"]["time_p"] for j in jsons1101_2s])
mext_deltas = np.array([j["merge_ext"]["deltas"][-1] for j in jsons0e00])
mext_times = np.array([j["merge_ext"]["times"][-1] for j in jsons0e00])
jsons_dict = {2: jsons1101_2s, 4: jsons1100_4s, 6: jsons1100_6s, 8: jsons1100_8s, 10: jsons1100_10s}
mer_times = {}
mer_deltas = {}
ver1_times = {}
ver1_deltas = {}
ver2_times = {}
ver2_deltas = {}
for seconds, jsons in jsons_dict.items():
mer_times[seconds] = np.array([j["merge_time"] for j in jsons])
mer_deltas[seconds] = np.array([j["merge_deltas"][-1][0] for j in jsons])
ver1_times[seconds] = np.array([j["veritas_time"] for j in jsons])
ver1_deltas[seconds] = np.array([j["veritas_deltas"][-1][0] for j in jsons])
ver2_times[seconds] = np.array([j["veritas_ara_time"] for j in jsons])
ver2_deltas[seconds] = np.array([j["veritas_ara_deltas"][-1][0] for j in jsons])
```
# Bound when given more or less time
```
mer_ts = [t.mean() for t in mer_times.values()]
mer_ys = [np.abs(d-kan_deltas).mean() for d in mer_deltas.values()]
ver1_ts = [t.mean() for t in ver1_times.values()]
ver1_ys = [np.abs(d-kan_deltas).mean() for d in ver1_deltas.values()]
ver2_ts = [t.mean() for t in ver2_times.values()]
ver2_ys = [np.abs(d-kan_deltas).mean() for d in ver2_deltas.values()]
mext_t = mext_times.mean()
mext_y = np.abs(mext_deltas-kan_deltas).mean()
kan_t = kan_times.mean()
fig, ax = plt.subplots(figsize=(16, 6))
ax.set_title("Average absolute deviation from MILP exact robustness delta per time budget (lower better)")
ax.plot(ver1_ts, ver1_ys, "x:", label="Veritas A*")
ax.plot(ver2_ts, ver2_ys, "o:", label="Veritas ARA*")
for i, (x, y) in enumerate(zip(ver2_ts, ver2_ys)):
ax.text(x, y-0.1, f"{M[4,i]:.0f}×", horizontalalignment='right', verticalalignment='top', c="gray")
ax.plot(mer_ts, mer_ys, ".:", label="My Merge")
ax.set_xlabel("Time bugdet [s]")
ax.set_ylabel("Absolute deviation from MILP exact")
l, = ax.plot([mext_t], [mext_y], "v--", lw=1, label="Chen et al.'s Merge")
ax.axhline(y=mext_y, ls=l.get_linestyle(), c=l.get_color(), lw=l.get_linewidth())
l, = ax.plot([kan_t], [0.0], "^--", lw=1, c="gray", label="Kantchelian MILP")
ax.axhline(y=0.0, ls=l.get_linestyle(), c=l.get_color(), lw=l.get_linewidth())
ax.text(kan_t, 0.1, f"{kan_slower_mext:.0f}×", horizontalalignment='right', verticalalignment='bottom', c="gray")
ax.set_xticks(range(0, 31))
ax.legend()
plt.show()
```
# How often better than Chen / as good as Kantchelian MILP
```
mext_ad = np.abs(mext_deltas-kan_deltas)
same_threshold = 0.1
f = 100.0 / len(mext_ad) # as percentage
mer_ts = [t.mean() for t in mer_times.values()]
mer_ys = [f*np.sum(mext_ad - np.abs(d-kan_deltas) > same_threshold) for d in mer_deltas.values()]
ver1_ts = [t.mean() for t in ver1_times.values()]
ver1_ys = [f*np.sum(mext_ad - np.abs(d-kan_deltas) > same_threshold) for d in ver1_deltas.values()]
ver2_ts = [t.mean() for t in ver2_times.values()]
ver2_ys = [f*np.sum(mext_ad - np.abs(d-kan_deltas) > same_threshold) for d in ver2_deltas.values()]
mer_ysb = [f*np.sum(np.abs(d-kan_deltas)-mext_ad > same_threshold) for d in mer_deltas.values()]
ver1_ysb = [f*np.sum(np.abs(d-kan_deltas)-mext_ad > same_threshold) for d in ver1_deltas.values()]
ver2_ysb = [f*np.sum(np.abs(d-kan_deltas)-mext_ad > same_threshold) for d in ver2_deltas.values()]
mer_ys2 = [f*np.sum(same_threshold > np.abs(d-kan_deltas)) for d in mer_deltas.values()]
ver1_ys2 = [f*np.sum(same_threshold > np.abs(d-kan_deltas)) for d in ver1_deltas.values()]
ver2_ys2 = [f*np.sum(same_threshold > np.abs(d-kan_deltas)) for d in ver2_deltas.values()]
mext_y2 = f*np.sum(same_threshold > np.abs(mext_deltas-kan_deltas))
fig, (ax1, ax3, ax2) = plt.subplots(1, 3, figsize=(20, 6))
ax1.set_title("How often better delta than Chen et al.'s Merge \n(higher better)")
ax3.set_title("How often worse delta than Chen et al.'s Merge \n(lower better)")
l, = ax1.plot(ver1_ts, ver1_ys, "x:", label="Veritas A*")
ax3.plot(ver1_ts, ver1_ysb, "x:", c=l.get_color())
l, = ax1.plot(ver2_ts, ver2_ys, "o:", label="Veritas ARA*")
ax3.plot(ver2_ts, ver2_ysb, "x:", c=l.get_color())
for i, (x, y) in enumerate(zip(ver2_ts, ver2_ys)):
ax1.text(x, y+0.1, f"{M[4,i]:.0f}×", horizontalalignment='right', verticalalignment='bottom', c="gray")
l, = ax1.plot(mer_ts, mer_ys, ".:", label="My Merge")
ax3.plot(mer_ts, mer_ysb, "x:", c=l.get_color())
for i, (x, y) in enumerate(zip(mer_ts, mer_ys)):
ax1.text(x, y+0.1, f"{M[5,i]:.0f}×", horizontalalignment='right', verticalalignment='bottom', c="gray")
ax1.set_xlabel("Time bugdet [s]")
ax1.set_ylabel(f"How often better than Chen et al.'s Merge' [%, n={len(mext_ad)}]")
ax3.set_xlabel("Time bugdet [s]")
ax3.set_ylabel(f"How often worse than Chen et al.'s Merge' [%, n={len(mext_ad)}]")
ax1.legend()
ax2.set_title(f"How often optimal (< {same_threshold} difference)\n(higher better)")
ax2.plot(ver1_ts, ver1_ys2, "x:", label="Veritas A*")
ax2.plot(ver2_ts, ver2_ys2, "o:", label="Veritas ARA*")
for i, (x, y) in enumerate(zip(ver1_ts, ver1_ys2)):
ax2.text(x-0.05, y+0.5, f"{M[3,i]:.0f}×", horizontalalignment='right', verticalalignment='bottom', c="gray")
ax2.plot(mer_ts, mer_ys2, ".:", label="My Merge")
l, = ax2.plot([mext_t], [mext_y], "v--", lw=1, label="Chen et al.'s Merge")
ax2.axhline(y=mext_y, ls=l.get_linestyle(), c=l.get_color(), lw=l.get_linewidth())
ax2.set_xlabel("Time bugdet [s]")
ax2.set_ylabel(f"How often (near) optimal [%, n={len(mext_ad)}]")
ax2.legend()
plt.show()
```
# Slower wrt Chen et al.'s Merge / MILP
```
mer_ys = [np.mean(t/mext_times) for t in mer_times.values()]
ver1_ys = [np.mean(t/mext_times) for t in ver1_times.values()]
ver2_ys = [np.mean(t/mext_times) for t in ver2_times.values()]
M = np.zeros((6, 5))
M[0,:] = [np.mean(t) for t in ver1_times.values()]
M[1,:] = [np.mean(t) for t in ver2_times.values()]
M[2,:] = [np.mean(t) for t in mer_times.values()]
M[3,:] = ver1_ys
M[4,:] = ver2_ys
M[5,:] = mer_ys
kan_slower_mext = np.mean(kan_times/mext_times)
fig, ax = plt.subplots(figsize=(8, 6))
ax.set_title("Average absolute deviation from MILP exact robustness delta per time budget")
ax.semilogy(ver1_ts, ver1_ys, "x:", label="Veritas A*")
ax.semilogy(ver2_ts, ver2_ys, "o:", label="Veritas ARA*")
ax.semilogy(mer_ts, mer_ys, ".:", label="My Merge")
ax.set_xlabel("Time bugdet [s]")
ax.set_ylabel("Absolute deviation from MILP exact")
ax.legend()
print("mean Chen et al.'s Merge: ", np.round(np.mean(mext_times), 2), "seconds")
print("Kantchelian MILP: ", np.round(kan_slower_mext, 1), "x")
print()
print("mean times [seconds]")
print(M[0:3,:].round(2))
print("how much slower than Chen et al.'s Merge [times slower than...]")
print(M[3:,:].round(1))
plt.show()
display("num_errors", sum(x["merge_ext"]["exc"] for x in jsons0e00_start40))
display("ad delta", np.mean(list(np.abs(k-x["merge_ext"]["deltas"][-1]) for k, x in zip(kan_deltas, jsons0e00_start40) if not x["merge_ext"]["exc"])))
np.mean(list(x["merge_ext"]["times"][-1] for x in jsons0e00_start40 if not x["merge_ext"]["exc"]))
```
| github_jupyter |
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
%matplotlib inline
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
pickle_file = '../dataset/arbimonTest1.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
image_size = 100
num_labels = 20
num_channels = 1 # grayscale
import numpy as np
def reformat(dataset, labels):
dataset = dataset.reshape(
(-1, image_size, image_size, num_channels)).astype(np.float32)
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
```
### Deep Convolution Neural Network
batch_size = 79
patch_size = 2
depth = 5
num_hidden = 100
### Structure of the Network
Input Layer: The photo
Layer 1: First Convolution Layer
Pooling: RELU
Layer 2: Second Convolution Layer
Pooling: RELU
Layer 5: Reshape
Layer 6: RELU
```
batch_size = 1000
patch_size = 2
depth = 5
num_hidden = 100
graph = tf.Graph()
with graph.as_default():
# Input data.
tf_train_dataset = tf.placeholder(
tf.float32, shape=(batch_size, image_size, image_size, num_channels))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
layer1_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, num_channels, depth], stddev=0.1))
layer1_biases = tf.Variable(tf.zeros([depth]))
layer2_weights = tf.Variable(tf.truncated_normal(
[patch_size, patch_size, depth, depth], stddev=0.1))
layer2_biases = tf.Variable(tf.constant(1.0, shape=[depth]))
layer3_weights = tf.Variable(tf.truncated_normal(
[image_size // 4 * image_size // 4 * depth, num_hidden], stddev=0.1))
layer3_biases = tf.Variable(tf.constant(1.0, shape=[num_hidden]))
layer4_weights = tf.Variable(tf.truncated_normal(
[num_hidden, num_labels], stddev=0.1))
layer4_biases = tf.Variable(tf.constant(1.0, shape=[num_labels]))
# Model.
def model(data):
conv = tf.nn.conv2d(data, layer1_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer1_biases)
conv = tf.nn.conv2d(hidden, layer2_weights, [1, 2, 2, 1], padding='SAME')
hidden = tf.nn.relu(conv + layer2_biases)
shape = hidden.get_shape().as_list()
reshape = tf.reshape(hidden, [shape[0], shape[1] * shape[2] * shape[3]])
hidden = tf.nn.relu(tf.matmul(reshape, layer3_weights) + layer3_biases)
return tf.matmul(hidden, layer4_weights) + layer4_biases
# Training computation.
logits = model(tf_train_dataset)
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits))
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.05).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(model(tf_valid_dataset))
test_prediction = tf.nn.softmax(model(tf_test_dataset))
num_steps = 1001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print('Initialized')
for step in range(num_steps):
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
batch_data = train_dataset[offset:(offset + batch_size), :, :, :]
batch_labels = train_labels[offset:(offset + batch_size), :]
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 50 == 0):
print('Minibatch loss at step %d: %f' % (step, l))
print('Minibatch accuracy: %.1f%%' % accuracy(predictions, batch_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
```
| github_jupyter |
# Home 5: Build a seq2seq model for machine translation.
### Name: [Your-Name?]
### Task: Translate English to [what-language?]
## 0. You will do the following:
1. Read and run my code.
2. Complete the code in Section 1.1 and Section 4.2.
* Translation **English** to **German** is not acceptable!!! Try another pair of languages.
3. **Make improvements.** Directly modify the code in Section 3. Do at least one of the two. By doing both correctly, you will get up to 1 bonus score to the total.
* Bi-LSTM instead of LSTM.
* Attention. (You are allowed to use existing code.)
4. Evaluate the translation using the BLEU score.
* Optional. Up to 1 bonus scores to the total.
5. Convert the notebook to .HTML file.
* The HTML file must contain the code and the output after execution.
6. Put the .HTML file in your Google Drive, Dropbox, or Github repo. (If you submit the file to Google Drive or Dropbox, you must make the file "open-access". The delay caused by "deny of access" may result in late penalty.)
7. Submit the link to the HTML file to Canvas.
### Hint:
To implement ```Bi-LSTM```, you will need the following code to build the encoder. Do NOT use Bi-LSTM for the decoder.
```
from keras.layers import Bidirectional, Concatenate
encoder_bilstm = Bidirectional(LSTM(latent_dim, return_state=True,
dropout=0.5, name='encoder_lstm'))
_, forward_h, forward_c, backward_h, backward_c = encoder_bilstm(encoder_inputs)
state_h = Concatenate()([forward_h, backward_h])
state_c = Concatenate()([forward_c, backward_c])
```
## 1. Data preparation
1. Download data (e.g., "deu-eng.zip") from http://www.manythings.org/anki/
2. Unzip the .ZIP file.
3. Put the .TXT file (e.g., "deu.txt") in the directory "./Data/".
### 1.1. Load and clean text
```
import re
import string
from unicodedata import normalize
import numpy
# load doc into memory
def load_doc(filename):
# open the file as read only
file = open(filename, mode='rt', encoding='utf-8')
# read all text
text = file.read()
# close the file
file.close()
return text
# split a loaded document into sentences
def to_pairs(doc):
lines = doc.strip().split('\n')
pairs = [line.split('\t') for line in lines]
return pairs
def clean_data(lines):
cleaned = list()
# prepare regex for char filtering
re_print = re.compile('[^%s]' % re.escape(string.printable))
# prepare translation table for removing punctuation
table = str.maketrans('', '', string.punctuation)
for pair in lines:
clean_pair = list()
for line in pair:
# normalize unicode characters
line = normalize('NFD', line).encode('ascii', 'ignore')
line = line.decode('UTF-8')
# tokenize on white space
line = line.split()
# convert to lowercase
line = [word.lower() for word in line]
# remove punctuation from each token
line = [word.translate(table) for word in line]
# remove non-printable chars form each token
line = [re_print.sub('', w) for w in line]
# remove tokens with numbers in them
line = [word for word in line if word.isalpha()]
# store as string
clean_pair.append(' '.join(line))
cleaned.append(clean_pair)
return numpy.array(cleaned)
```
#### Fill the following blanks:
```
# e.g., filename = 'Data/deu.txt'
filename = <what is your file name?>
# e.g., n_train = 20000
n_train = <how many sentences are you going to use for training?>
# load dataset
doc = load_doc(filename)
# split into Language1-Language2 pairs
pairs = to_pairs(doc)
# clean sentences
clean_pairs = clean_data(pairs)[0:n_train, :]
for i in range(3000, 3010):
print('[' + clean_pairs[i, 0] + '] => [' + clean_pairs[i, 1] + ']')
input_texts = clean_pairs[:, 0]
target_texts = ['\t' + text + '\n' for text in clean_pairs[:, 1]]
print('Length of input_texts: ' + str(input_texts.shape))
print('Length of target_texts: ' + str(input_texts.shape))
max_encoder_seq_length = max(len(line) for line in input_texts)
max_decoder_seq_length = max(len(line) for line in target_texts)
print('max length of input sentences: %d' % (max_encoder_seq_length))
print('max length of target sentences: %d' % (max_decoder_seq_length))
```
**Remark:** To this end, you have two lists of sentences: input_texts and target_texts
## 2. Text processing
### 2.1. Convert texts to sequences
- Input: A list of $n$ sentences (with max length $t$).
- It is represented by a $n\times t$ matrix after the tokenization and zero-padding.
```
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
# encode and pad sequences
def text2sequences(max_len, lines):
tokenizer = Tokenizer(char_level=True, filters='')
tokenizer.fit_on_texts(lines)
seqs = tokenizer.texts_to_sequences(lines)
seqs_pad = pad_sequences(seqs, maxlen=max_len, padding='post')
return seqs_pad, tokenizer.word_index
encoder_input_seq, input_token_index = text2sequences(max_encoder_seq_length,
input_texts)
decoder_input_seq, target_token_index = text2sequences(max_decoder_seq_length,
target_texts)
print('shape of encoder_input_seq: ' + str(encoder_input_seq.shape))
print('shape of input_token_index: ' + str(len(input_token_index)))
print('shape of decoder_input_seq: ' + str(decoder_input_seq.shape))
print('shape of target_token_index: ' + str(len(target_token_index)))
num_encoder_tokens = len(input_token_index) + 1
num_decoder_tokens = len(target_token_index) + 1
print('num_encoder_tokens: ' + str(num_encoder_tokens))
print('num_decoder_tokens: ' + str(num_decoder_tokens))
```
**Remark:** To this end, the input language and target language texts are converted to 2 matrices.
- Their number of rows are both n_train.
- Their number of columns are respective max_encoder_seq_length and max_decoder_seq_length.
The followings print a sentence and its representation as a sequence.
```
target_texts[100]
decoder_input_seq[100, :]
```
## 2.2. One-hot encode
- Input: A list of $n$ sentences (with max length $t$).
- It is represented by a $n\times t$ matrix after the tokenization and zero-padding.
- It is represented by a $n\times t \times v$ tensor ($t$ is the number of unique chars) after the one-hot encoding.
```
from keras.utils import to_categorical
# one hot encode target sequence
def onehot_encode(sequences, max_len, vocab_size):
n = len(sequences)
data = numpy.zeros((n, max_len, vocab_size))
for i in range(n):
data[i, :, :] = to_categorical(sequences[i], num_classes=vocab_size)
return data
encoder_input_data = onehot_encode(encoder_input_seq, max_encoder_seq_length, num_encoder_tokens)
decoder_input_data = onehot_encode(decoder_input_seq, max_decoder_seq_length, num_decoder_tokens)
decoder_target_seq = numpy.zeros(decoder_input_seq.shape)
decoder_target_seq[:, 0:-1] = decoder_input_seq[:, 1:]
decoder_target_data = onehot_encode(decoder_target_seq,
max_decoder_seq_length,
num_decoder_tokens)
print(encoder_input_data.shape)
print(decoder_input_data.shape)
```
## 3. Build the networks (for training)
- Build encoder, decoder, and connect the two modules to get "model".
- Fit the model on the bilingual data to train the parameters in the encoder and decoder.
### 3.1. Encoder network
- Input: one-hot encode of the input language
- Return:
-- output (all the hidden states $h_1, \cdots , h_t$) are always discarded
-- the final hidden state $h_t$
-- the final conveyor belt $c_t$
```
from keras.layers import Input, LSTM
from keras.models import Model
latent_dim = 256
# inputs of the encoder network
encoder_inputs = Input(shape=(None, num_encoder_tokens),
name='encoder_inputs')
# set the LSTM layer
encoder_lstm = LSTM(latent_dim, return_state=True,
dropout=0.5, name='encoder_lstm')
_, state_h, state_c = encoder_lstm(encoder_inputs)
# build the encoder network model
encoder_model = Model(inputs=encoder_inputs,
outputs=[state_h, state_c],
name='encoder')
```
Print a summary and save the encoder network structure to "./encoder.pdf"
```
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(encoder_model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=encoder_model, show_shapes=False,
to_file='encoder.pdf'
)
encoder_model.summary()
```
### 3.2. Decoder network
- Inputs:
-- one-hot encode of the target language
-- The initial hidden state $h_t$
-- The initial conveyor belt $c_t$
- Return:
-- output (all the hidden states) $h_1, \cdots , h_t$
-- the final hidden state $h_t$ (discarded in the training and used in the prediction)
-- the final conveyor belt $c_t$ (discarded in the training and used in the prediction)
```
from keras.layers import Input, LSTM, Dense
from keras.models import Model
# inputs of the decoder network
decoder_input_h = Input(shape=(latent_dim,), name='decoder_input_h')
decoder_input_c = Input(shape=(latent_dim,), name='decoder_input_c')
decoder_input_x = Input(shape=(None, num_decoder_tokens), name='decoder_input_x')
# set the LSTM layer
decoder_lstm = LSTM(latent_dim, return_sequences=True,
return_state=True, dropout=0.5, name='decoder_lstm')
decoder_lstm_outputs, state_h, state_c = decoder_lstm(decoder_input_x,
initial_state=[decoder_input_h, decoder_input_c])
# set the dense layer
decoder_dense = Dense(num_decoder_tokens, activation='softmax', name='decoder_dense')
decoder_outputs = decoder_dense(decoder_lstm_outputs)
# build the decoder network model
decoder_model = Model(inputs=[decoder_input_x, decoder_input_h, decoder_input_c],
outputs=[decoder_outputs, state_h, state_c],
name='decoder')
```
Print a summary and save the encoder network structure to "./decoder.pdf"
```
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(decoder_model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=decoder_model, show_shapes=False,
to_file='decoder.pdf'
)
decoder_model.summary()
```
### 3.3. Connect the encoder and decoder
```
# input layers
encoder_input_x = Input(shape=(None, num_encoder_tokens), name='encoder_input_x')
decoder_input_x = Input(shape=(None, num_decoder_tokens), name='decoder_input_x')
# connect encoder to decoder
encoder_final_states = encoder_model([encoder_input_x])
decoder_lstm_output, _, _ = decoder_lstm(decoder_input_x, initial_state=encoder_final_states)
decoder_pred = decoder_dense(decoder_lstm_output)
model = Model(inputs=[encoder_input_x, decoder_input_x],
outputs=decoder_pred,
name='model_training')
print(state_h)
print(decoder_input_h)
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot, plot_model
SVG(model_to_dot(model, show_shapes=False).create(prog='dot', format='svg'))
plot_model(
model=model, show_shapes=False,
to_file='model_training.pdf'
)
model.summary()
```
### 3.5. Fit the model on the bilingual dataset
- encoder_input_data: one-hot encode of the input language
- decoder_input_data: one-hot encode of the input language
- decoder_target_data: labels (left shift of decoder_input_data)
- tune the hyper-parameters
- stop when the validation loss stop decreasing.
```
print('shape of encoder_input_data' + str(encoder_input_data.shape))
print('shape of decoder_input_data' + str(decoder_input_data.shape))
print('shape of decoder_target_data' + str(decoder_target_data.shape))
model.compile(optimizer='rmsprop', loss='categorical_crossentropy')
model.fit([encoder_input_data, decoder_input_data], # training data
decoder_target_data, # labels (left shift of the target sequences)
batch_size=64, epochs=50, validation_split=0.2)
model.save('seq2seq.h5')
```
## 4. Make predictions
### 4.1. Translate English to XXX
1. Encoder read a sentence (source language) and output its final states, $h_t$ and $c_t$.
2. Take the [star] sign "\t" and the final state $h_t$ and $c_t$ as input and run the decoder.
3. Get the new states and predicted probability distribution.
4. sample a char from the predicted probability distribution
5. take the sampled char and the new states as input and repeat the process (stop if reach the [stop] sign "\n").
```
# Reverse-lookup token index to decode sequences back to something readable.
reverse_input_char_index = dict((i, char) for char, i in input_token_index.items())
reverse_target_char_index = dict((i, char) for char, i in target_token_index.items())
def decode_sequence(input_seq):
states_value = encoder_model.predict(input_seq)
target_seq = numpy.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, target_token_index['\t']] = 1.
stop_condition = False
decoded_sentence = ''
while not stop_condition:
output_tokens, h, c = decoder_model.predict([target_seq] + states_value)
# this line of code is greedy selection
# try to use multinomial sampling instead (with temperature)
sampled_token_index = numpy.argmax(output_tokens[0, -1, :])
sampled_char = reverse_target_char_index[sampled_token_index]
decoded_sentence += sampled_char
if (sampled_char == '\n' or
len(decoded_sentence) > max_decoder_seq_length):
stop_condition = True
target_seq = numpy.zeros((1, 1, num_decoder_tokens))
target_seq[0, 0, sampled_token_index] = 1.
states_value = [h, c]
return decoded_sentence
for seq_index in range(2100, 2120):
# Take one sequence (part of the training set)
# for trying out decoding.
input_seq = encoder_input_data[seq_index: seq_index + 1]
decoded_sentence = decode_sequence(input_seq)
print('-')
print('English: ', input_texts[seq_index])
print('German (true): ', target_texts[seq_index][1:-1])
print('German (pred): ', decoded_sentence[0:-1])
```
### 4.2. Translate an English sentence to the target language
1. Tokenization
2. One-hot encode
3. Translate
```
input_sentence = 'I love you'
input_sequence = <do tokenization...>
input_x = <do one-hot encode...>
translated_sentence = <do translation...>
print('source sentence is: ' + input_sentence)
print('translated sentence is: ' + translated_sentence)
```
## 5. Evaluate the translation using BLEU score
Reference:
- https://machinelearningmastery.com/calculate-bleu-score-for-text-python/
- https://en.wikipedia.org/wiki/BLEU
**Hint:**
- Randomly partition the dataset to training, validation, and test.
- Evaluate the BLEU score using the test set. Report the average.
- A reasonable BLEU score should be 0.1 ~ 0.5.
| github_jupyter |
# Machine Learning Engineer Nanodegree
## Unsupervised Learning
## Project: Creating Customer Segments
Welcome to the third project of the Machine Learning Engineer Nanodegree! In this notebook, some template code has already been provided for you, and it will be your job to implement the additional functionality necessary to successfully complete this project. Sections that begin with **'Implementation'** in the header indicate that the following block of code will require additional functionality which you must provide. Instructions will be provided for each section and the specifics of the implementation are marked in the code block with a `'TODO'` statement. Please be sure to read the instructions carefully!
In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question X'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
>**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
## Getting Started
In this project, you will analyze a dataset containing data on various customers' annual spending amounts (reported in *monetary units*) of diverse product categories for internal structure. One goal of this project is to best describe the variation in the different types of customers that a wholesale distributor interacts with. Doing so would equip the distributor with insight into how to best structure their delivery service to meet the needs of each customer.
The dataset for this project can be found on the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Wholesale+customers). For the purposes of this project, the features `'Channel'` and `'Region'` will be excluded in the analysis — with focus instead on the six product categories recorded for customers.
Run the code block below to load the wholesale customers dataset, along with a few of the necessary Python libraries required for this project. You will know the dataset loaded successfully if the size of the dataset is reported.
```
# Import libraries necessary for this project
import numpy as np
import pandas as pd
from IPython.display import display # Allows the use of display() for DataFrames
# Import supplementary visualizations code visuals.py
import visuals as vs
# Pretty display for notebooks
%matplotlib inline
# Load the wholesale customers dataset
try:
data = pd.read_csv("customers.csv")
data.drop(['Region', 'Channel'], axis = 1, inplace = True)
print "Wholesale customers dataset has {} samples with {} features each.".format(*data.shape)
except:
print "Dataset could not be loaded. Is the dataset missing?"
```
## Data Exploration
In this section, you will begin exploring the data through visualizations and code to understand how each feature is related to the others. You will observe a statistical description of the dataset, consider the relevance of each feature, and select a few sample data points from the dataset which you will track through the course of this project.
Run the code block below to observe a statistical description of the dataset. Note that the dataset is composed of six important product categories: **'Fresh'**, **'Milk'**, **'Grocery'**, **'Frozen'**, **'Detergents_Paper'**, and **'Delicatessen'**. Consider what each category represents in terms of products you could purchase.
```
# Display a description of the dataset
display(data.describe())
```
### Implementation: Selecting Samples
To get a better understanding of the customers and how their data will transform through the analysis, it would be best to select a few sample data points and explore them in more detail. In the code block below, add **three** indices of your choice to the `indices` list which will represent the customers to track. It is suggested to try different sets of samples until you obtain customers that vary significantly from one another.
```
# TODO: Select three indices of your choice you wish to sample from the dataset
indices = [338, 154, 181]
# Create a DataFrame of the chosen samples
samples = pd.DataFrame(data.loc[indices], columns = data.keys()).reset_index(drop = True)
print "Chosen samples of wholesale customers dataset:"
display(samples)
```
### Question 1
Consider the total purchase cost of each product category and the statistical description of the dataset above for your sample customers.
*What kind of establishment (customer) could each of the three samples you've chosen represent?*
**Hint:** Examples of establishments include places like markets, cafes, and retailers, among many others. Avoid using names for establishments, such as saying *"McDonalds"* when describing a sample customer as a restaurant.
**Answer:** I deliberately looked for the records with min(fresh), min(milk) and max(fresh) and it did not dissapoint me to see that they seem to represent vastly different customer segments.
1. The first record is in the top 25% for 'Frozen' goods, top 50% for 'Grocery' and 'Delicatessen'and in the bottom 25% for the last 3 categories. This could be a small grocery store which specializes in frozen goods, but has a grocery and deli section as well. The lack of fresh goods (taken to mean produce), however, seems to suggest otherwise. Though the spending is fairly high, it's not incredibly so (I'm not convinced even a small grocery store only sells ~25,000 m.u. worth of goods in a year). Threfore, it's possible that this could also be a small group of individuals (such as college roommates) who primarily eat frozen foods (eg. frozen pizza, fries).
2. The second record has very low spending all around (WAY below the 25th percentile). This customer is probably an individual, and one that shops at other places.
3. This customer exceeds the 75th percentile in all categories, although they only come close to the max value in one category (Fresh). This is likely a grocery store of some kind, which specializes in selling produce.
### Implementation: Feature Relevance
One interesting thought to consider is if one (or more) of the six product categories is actually relevant for understanding customer purchasing. That is to say, is it possible to determine whether customers purchasing some amount of one category of products will necessarily purchase some proportional amount of another category of products? We can make this determination quite easily by training a supervised regression learner on a subset of the data with one feature removed, and then score how well that model can predict the removed feature.
In the code block below, you will need to implement the following:
- Assign `new_data` a copy of the data by removing a feature of your choice using the `DataFrame.drop` function.
- Use `sklearn.cross_validation.train_test_split` to split the dataset into training and testing sets.
- Use the removed feature as your target label. Set a `test_size` of `0.25` and set a `random_state`.
- Import a decision tree regressor, set a `random_state`, and fit the learner to the training data.
- Report the prediction score of the testing set using the regressor's `score` function.
```
# TODO: Make a copy of the DataFrame, using the 'drop' function to drop the given feature
test_label = 'Grocery'
new_data = data.drop(test_label, axis = 1)
test_feature = data[test_label]
# TODO: Split the data into training and testing sets using the given feature as the target
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(new_data, test_feature, test_size=0.25, random_state=777)
# TODO: Create a decision tree regressor and fit it to the training set
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
regressor.fit(X_train, y_train)
# TODO: Report the score of the prediction using the testing set
score = regressor.score(X_test, y_test)
print score
```
### Question 2
*Which feature did you attempt to predict? What was the reported prediction score? Is this feature is necessary for identifying customers' spending habits?*
**Hint:** The coefficient of determination, `R^2`, is scored between 0 and 1, with 1 being a perfect fit. A negative `R^2` implies the model fails to fit the data.
**Answer:** I attempted to predict 'Grocery'. The reported prediction score ranged between 0.78-0.82 when run multiple times (even with a constant random_state). This feature seems to be pretty good for identifying customer spending habits
### Visualize Feature Distributions
To get a better understanding of the dataset, we can construct a scatter matrix of each of the six product features present in the data. If you found that the feature you attempted to predict above is relevant for identifying a specific customer, then the scatter matrix below may not show any correlation between that feature and the others. Conversely, if you believe that feature is not relevant for identifying a specific customer, the scatter matrix might show a correlation between that feature and another feature in the data. Run the code block below to produce a scatter matrix.
```
# Produce a scatter matrix for each pair of features in the data
pd.scatter_matrix(data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
```
### Question 3
*Are there any pairs of features which exhibit some degree of correlation? Does this confirm or deny your suspicions about the relevance of the feature you attempted to predict? How is the data for those features distributed?*
**Hint:** Is the data normally distributed? Where do most of the data points lie?
**Answer:** Grocery seems to be mostly correlated with 'Milk' and 'Detergents_Paper'. The remaining 3 features are not quite as correlated (In fact, they aren't really correlated with anything else at all). This confirms my suspicion that the feature I chose (Grocery) is relevant. The data, however seems to be highly skewed to the left (a few very large outliers) across all features. This suggests that the company perhaps has a few very large (probably corporate) buyers.
## Data Preprocessing
In this section, you will preprocess the data to create a better representation of customers by performing a scaling on the data and detecting (and optionally removing) outliers. Preprocessing data is often times a critical step in assuring that results you obtain from your analysis are significant and meaningful.
### Implementation: Feature Scaling
If data is not normally distributed, especially if the mean and median vary significantly (indicating a large skew), it is most [often appropriate](http://econbrowser.com/archives/2014/02/use-of-logarithms-in-economics) to apply a non-linear scaling — particularly for financial data. One way to achieve this scaling is by using a [Box-Cox test](http://scipy.github.io/devdocs/generated/scipy.stats.boxcox.html), which calculates the best power transformation of the data that reduces skewness. A simpler approach which can work in most cases would be applying the natural logarithm.
In the code block below, you will need to implement the following:
- Assign a copy of the data to `log_data` after applying logarithmic scaling. Use the `np.log` function for this.
- Assign a copy of the sample data to `log_samples` after applying logarithmic scaling. Again, use `np.log`.
```
# TODO: Scale the data using the natural logarithm
log_data = np.log(data)
# TODO: Scale the sample data using the natural logarithm
log_samples = np.log(samples)
# Produce a scatter matrix for each pair of newly-transformed features
pd.scatter_matrix(log_data, alpha = 0.3, figsize = (14,8), diagonal = 'kde');
```
### Observation
After applying a natural logarithm scaling to the data, the distribution of each feature should appear much more normal. For any pairs of features you may have identified earlier as being correlated, observe here whether that correlation is still present (and whether it is now stronger or weaker than before).
Run the code below to see how the sample data has changed after having the natural logarithm applied to it.
```
# Display the log-transformed sample data
display(log_samples)
```
### Implementation: Outlier Detection
Detecting outliers in the data is extremely important in the data preprocessing step of any analysis. The presence of outliers can often skew results which take into consideration these data points. There are many "rules of thumb" for what constitutes an outlier in a dataset. Here, we will use [Tukey's Method for identfying outliers](http://datapigtechnologies.com/blog/index.php/highlighting-outliers-in-your-data-with-the-tukey-method/): An *outlier step* is calculated as 1.5 times the interquartile range (IQR). A data point with a feature that is beyond an outlier step outside of the IQR for that feature is considered abnormal.
In the code block below, you will need to implement the following:
- Assign the value of the 25th percentile for the given feature to `Q1`. Use `np.percentile` for this.
- Assign the value of the 75th percentile for the given feature to `Q3`. Again, use `np.percentile`.
- Assign the calculation of an outlier step for the given feature to `step`.
- Optionally remove data points from the dataset by adding indices to the `outliers` list.
**NOTE:** If you choose to remove any outliers, ensure that the sample data does not contain any of these points!
Once you have performed this implementation, the dataset will be stored in the variable `good_data`.
```
from collections import defaultdict
outlier_indices = defaultdict(int)
# For each feature find the data points with extreme high or low values
for feature in log_data.keys():
# TODO: Calculate Q1 (25th percentile of the data) for the given feature
Q1 = np.percentile(log_data[feature], 25)
# TODO: Calculate Q3 (75th percentile of the data) for the given feature
Q3 = np.percentile(log_data[feature], 75)
# TODO: Use the interquartile range to calculate an outlier step (1.5 times the interquartile range)
step = (Q3 - Q1) * 1.5
# Display the outliers
print "Data points considered outliers for the feature '{}':".format(feature)
rows = log_data[~((log_data[feature] >= Q1 - step) & (log_data[feature] <= Q3 + step))]
feature_indices = rows.index
display(rows)
# Track all indices that are outliers
for index in feature_indices:
if (index not in indices):
outlier_indices[index] += 1
# OPTIONAL: Select the indices for data points you wish to remove
# If an index appeared 3 times (was an outlier for at least 3 of the categories), drop the row
outliers = []
for index in outlier_indices:
if outlier_indices[index] >= 1:
outliers.append(index)
print(outliers)
print(len(outliers))
# Remove the outliers, if any were specified
good_data = log_data.drop(log_data.index[outliers]).reset_index(drop = True)
display(good_data.describe())
display(log_data.describe())
```
### Question 4
*Are there any data points considered outliers for more than one feature based on the definition above? Should these data points be removed from the dataset? If any data points were added to the `outliers` list to be removed, explain why.*
**Answer:**
There were 42 unique rows that had at least 1 outlier, 5 with 2+, 1 with 3+, and 0 with 4+.
Based on this, I have chosen to remove any data point with an outlier (which works out roughly 10% of the original data). I chose to remove any rows with at least 1 feature that is an outlier. I decided to do this because it seemed to lower the average distance between the mean and the median.
To determine this, I recorded the mean and median of each feature after removing data points with at least 1 feature that is an outlier, at least 2 features that are an outlier, and without removing any data points. Lastly I calculated the difference between the mean and median of each column and averaged the result. The results were:
|Min # of Outlier Features|Average Difference Between Mean and Median|
|---|---|
|None (Base)|0.123|
|1|0.0852|
|2|0.110|
There isn't much improvement when only removing the 5 data points, but there is a much larger improvement when removing all 42 outliers.
Consequently, the first 2 sample points I chose were removed. I have opted to not remove them (and thus, the averages will be slightly different from what I initially calculated, but the overall effect should be the same) by making them an exception and then re-running the code. It is quite surprising that the last sample point wasn't considered an outlier as it had that maximum value in the 'Fresh' category. As a matter of fact, outliers for 'Fresh' were only those that were lower than the 25th percentile - 1.5IQR.
## Feature Transformation
In this section you will use principal component analysis (PCA) to draw conclusions about the underlying structure of the wholesale customer data. Since using PCA on a dataset calculates the dimensions which best maximize variance, we will find which compound combinations of features best describe customers.
### Implementation: PCA
Now that the data has been scaled to a more normal distribution and has had any necessary outliers removed, we can now apply PCA to the `good_data` to discover which dimensions about the data best maximize the variance of features involved. In addition to finding these dimensions, PCA will also report the *explained variance ratio* of each dimension — how much variance within the data is explained by that dimension alone. Note that a component (dimension) from PCA can be considered a new "feature" of the space, however it is a composition of the original features present in the data.
In the code block below, you will need to implement the following:
- Import `sklearn.decomposition.PCA` and assign the results of fitting PCA in six dimensions with `good_data` to `pca`.
- Apply a PCA transformation of `log_samples` using `pca.transform`, and assign the results to `pca_samples`.
```
# TODO: Apply PCA by fitting the good data with the same number of dimensions as features
from sklearn.decomposition import PCA
pca = PCA(n_components=6, random_state=777).fit(good_data)
# TODO: Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Generate PCA results plot
pca_results = vs.pca_results(good_data, pca)
```
### Question 5
*How much variance in the data is explained* ***in total*** *by the first and second principal component? What about the first four principal components? Using the visualization provided above, discuss what the first four dimensions best represent in terms of customer spending.*
**Hint:** A positive increase in a specific dimension corresponds with an *increase* of the *positive-weighted* features and a *decrease* of the *negative-weighted* features. The rate of increase or decrease is based on the indivdual feature weights.
**Answer:** 72.13% of the total variance is explained by the first 2 principal components. 92.95% is explained by the first 4 PCs.
Dimension 1: This dimension suggests that an increase in both 'Fresh' and 'Frozen' results in a moderate decrease in spending on 'Milk' and 'Grocery' and a large decrease in spending in 'Detergents_Paper'
Dimension 2: This dimension suggests that small purchases of 'Detergents_Paper' is correlated with a large decrease in spending on 'Fresh', 'Frozen', and 'Deli' (In fact, it is a decrease in spending in all other categories)
Dimension 3: This dimension suggests that large purchases of 'Frozen' and 'Deli' goods are correlated with a large decrease in spending on 'Fresh'
Dimension 4: This dimension suggests that very large purchases of 'Deli' is correlated with a large decrease in spending on 'Frozen' and a moderate decrease in spending on 'Detergents_Paper'
When comparing with the scatter plots from above, an interesting observation can be made. Previously we determined that 'Grocery', 'Milk' and 'Detergents_Paper' were correlated. In fact, according to the scatter plots, they are all positively correlated (that is, and increase in one results in an increase in the other). The correlation between 'Milk' and 'Detergents_Paper' is a bit weaker but the overall shape is there. However, from the PCA, we can see that asides from dimension 1, 'Grocery' and 'Milk' are negatively correlated with 'Detergents_Paper'. 'Grocery' and 'Milk' are positively correlated in all cases except for the last dimension, which only represents ~2.5% of the total variance and can be considered an edge case.
### Observation
Run the code below to see how the log-transformed sample data has changed after having a PCA transformation applied to it in six dimensions. Observe the numerical value for the first four dimensions of the sample points. Consider if this is consistent with your initial interpretation of the sample points.
```
# Display sample log-data after having a PCA transformation applied
display(log_samples)
display(pd.DataFrame(np.round(pca_samples, 4), columns = pca_results.index.values))
```
### Implementation: Dimensionality Reduction
When using principal component analysis, one of the main goals is to reduce the dimensionality of the data — in effect, reducing the complexity of the problem. Dimensionality reduction comes at a cost: Fewer dimensions used implies less of the total variance in the data is being explained. Because of this, the *cumulative explained variance ratio* is extremely important for knowing how many dimensions are necessary for the problem. Additionally, if a signifiant amount of variance is explained by only two or three dimensions, the reduced data can be visualized afterwards.
In the code block below, you will need to implement the following:
- Assign the results of fitting PCA in two dimensions with `good_data` to `pca`.
- Apply a PCA transformation of `good_data` using `pca.transform`, and assign the results to `reduced_data`.
- Apply a PCA transformation of `log_samples` using `pca.transform`, and assign the results to `pca_samples`.
```
# TODO: Apply PCA by fitting the good data with only two dimensions
pca = PCA(n_components=2, random_state=777).fit(good_data)
# TODO: Transform the good data using the PCA fit above
reduced_data = pca.transform(good_data)
# TODO: Transform log_samples using the PCA fit above
pca_samples = pca.transform(log_samples)
# Create a DataFrame for the reduced data
reduced_data = pd.DataFrame(reduced_data, columns = ['Dimension 1', 'Dimension 2'])
```
### Observation
Run the code below to see how the log-transformed sample data has changed after having a PCA transformation applied to it using only two dimensions. Observe how the values for the first two dimensions remains unchanged when compared to a PCA transformation in six dimensions.
```
# Display sample log-data after applying PCA transformation in two dimensions
display(pd.DataFrame(np.round(pca_samples, 4), columns = ['Dimension 1', 'Dimension 2']))
```
## Visualizing a Biplot
A biplot is a scatterplot where each data point is represented by its scores along the principal components. The axes are the principal components (in this case `Dimension 1` and `Dimension 2`). In addition, the biplot shows the projection of the original features along the components. A biplot can help us interpret the reduced dimensions of the data, and discover relationships between the principal components and original features.
Run the code cell below to produce a biplot of the reduced-dimension data.
```
# Create a biplot
vs.biplot(good_data, reduced_data, pca)
```
### Observation
Once we have the original feature projections (in red), it is easier to interpret the relative position of each data point in the scatterplot. For instance, a point the lower right corner of the figure will likely correspond to a customer that spends a lot on `'Milk'`, `'Grocery'` and `'Detergents_Paper'`, but not so much on the other product categories.
From the biplot, which of the original features are most strongly correlated with the first component? What about those that are associated with the second component? Do these observations agree with the pca_results plot you obtained earlier?
## Clustering
In this section, you will choose to use either a K-Means clustering algorithm or a Gaussian Mixture Model clustering algorithm to identify the various customer segments hidden in the data. You will then recover specific data points from the clusters to understand their significance by transforming them back into their original dimension and scale.
### Question 6
*What are the advantages to using a K-Means clustering algorithm? What are the advantages to using a Gaussian Mixture Model clustering algorithm? Given your observations about the wholesale customer data so far, which of the two algorithms will you use and why?*
**Answer:**
### Implementation: Creating Clusters
Depending on the problem, the number of clusters that you expect to be in the data may already be known. When the number of clusters is not known *a priori*, there is no guarantee that a given number of clusters best segments the data, since it is unclear what structure exists in the data — if any. However, we can quantify the "goodness" of a clustering by calculating each data point's *silhouette coefficient*. The [silhouette coefficient](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.silhouette_score.html) for a data point measures how similar it is to its assigned cluster from -1 (dissimilar) to 1 (similar). Calculating the *mean* silhouette coefficient provides for a simple scoring method of a given clustering.
In the code block below, you will need to implement the following:
- Fit a clustering algorithm to the `reduced_data` and assign it to `clusterer`.
- Predict the cluster for each data point in `reduced_data` using `clusterer.predict` and assign them to `preds`.
- Find the cluster centers using the algorithm's respective attribute and assign them to `centers`.
- Predict the cluster for each sample data point in `pca_samples` and assign them `sample_preds`.
- Import `sklearn.metrics.silhouette_score` and calculate the silhouette score of `reduced_data` against `preds`.
- Assign the silhouette score to `score` and print the result.
```
# TODO: Apply your clustering algorithm of choice to the reduced data
clusterer = None
# TODO: Predict the cluster for each data point
preds = None
# TODO: Find the cluster centers
centers = None
# TODO: Predict the cluster for each transformed sample data point
sample_preds = None
# TODO: Calculate the mean silhouette coefficient for the number of clusters chosen
score = None
```
### Question 7
*Report the silhouette score for several cluster numbers you tried. Of these, which number of clusters has the best silhouette score?*
**Answer:**
### Cluster Visualization
Once you've chosen the optimal number of clusters for your clustering algorithm using the scoring metric above, you can now visualize the results by executing the code block below. Note that, for experimentation purposes, you are welcome to adjust the number of clusters for your clustering algorithm to see various visualizations. The final visualization provided should, however, correspond with the optimal number of clusters.
```
# Display the results of the clustering from implementation
vs.cluster_results(reduced_data, preds, centers, pca_samples)
```
### Implementation: Data Recovery
Each cluster present in the visualization above has a central point. These centers (or means) are not specifically data points from the data, but rather the *averages* of all the data points predicted in the respective clusters. For the problem of creating customer segments, a cluster's center point corresponds to *the average customer of that segment*. Since the data is currently reduced in dimension and scaled by a logarithm, we can recover the representative customer spending from these data points by applying the inverse transformations.
In the code block below, you will need to implement the following:
- Apply the inverse transform to `centers` using `pca.inverse_transform` and assign the new centers to `log_centers`.
- Apply the inverse function of `np.log` to `log_centers` using `np.exp` and assign the true centers to `true_centers`.
```
# TODO: Inverse transform the centers
log_centers = None
# TODO: Exponentiate the centers
true_centers = None
# Display the true centers
segments = ['Segment {}'.format(i) for i in range(0,len(centers))]
true_centers = pd.DataFrame(np.round(true_centers), columns = data.keys())
true_centers.index = segments
display(true_centers)
```
### Question 8
Consider the total purchase cost of each product category for the representative data points above, and reference the statistical description of the dataset at the beginning of this project. *What set of establishments could each of the customer segments represent?*
**Hint:** A customer who is assigned to `'Cluster X'` should best identify with the establishments represented by the feature set of `'Segment X'`.
**Answer:**
### Question 9
*For each sample point, which customer segment from* ***Question 8*** *best represents it? Are the predictions for each sample point consistent with this?*
Run the code block below to find which cluster each sample point is predicted to be.
```
# Display the predictions
for i, pred in enumerate(sample_preds):
print "Sample point", i, "predicted to be in Cluster", pred
```
**Answer:**
## Conclusion
In this final section, you will investigate ways that you can make use of the clustered data. First, you will consider how the different groups of customers, the ***customer segments***, may be affected differently by a specific delivery scheme. Next, you will consider how giving a label to each customer (which *segment* that customer belongs to) can provide for additional features about the customer data. Finally, you will compare the ***customer segments*** to a hidden variable present in the data, to see whether the clustering identified certain relationships.
### Question 10
Companies will often run [A/B tests](https://en.wikipedia.org/wiki/A/B_testing) when making small changes to their products or services to determine whether making that change will affect its customers positively or negatively. The wholesale distributor is considering changing its delivery service from currently 5 days a week to 3 days a week. However, the distributor will only make this change in delivery service for customers that react positively. *How can the wholesale distributor use the customer segments to determine which customers, if any, would react positively to the change in delivery service?*
**Hint:** Can we assume the change affects all customers equally? How can we determine which group of customers it affects the most?
**Answer:**
### Question 11
Additional structure is derived from originally unlabeled data when using clustering techniques. Since each customer has a ***customer segment*** it best identifies with (depending on the clustering algorithm applied), we can consider *'customer segment'* as an **engineered feature** for the data. Assume the wholesale distributor recently acquired ten new customers and each provided estimates for anticipated annual spending of each product category. Knowing these estimates, the wholesale distributor wants to classify each new customer to a ***customer segment*** to determine the most appropriate delivery service.
*How can the wholesale distributor label the new customers using only their estimated product spending and the* ***customer segment*** *data?*
**Hint:** A supervised learner could be used to train on the original customers. What would be the target variable?
**Answer:**
### Visualizing Underlying Distributions
At the beginning of this project, it was discussed that the `'Channel'` and `'Region'` features would be excluded from the dataset so that the customer product categories were emphasized in the analysis. By reintroducing the `'Channel'` feature to the dataset, an interesting structure emerges when considering the same PCA dimensionality reduction applied earlier to the original dataset.
Run the code block below to see how each data point is labeled either `'HoReCa'` (Hotel/Restaurant/Cafe) or `'Retail'` the reduced space. In addition, you will find the sample points are circled in the plot, which will identify their labeling.
```
# Display the clustering results based on 'Channel' data
vs.channel_results(reduced_data, outliers, pca_samples)
```
### Question 12
*How well does the clustering algorithm and number of clusters you've chosen compare to this underlying distribution of Hotel/Restaurant/Cafe customers to Retailer customers? Are there customer segments that would be classified as purely 'Retailers' or 'Hotels/Restaurants/Cafes' by this distribution? Would you consider these classifications as consistent with your previous definition of the customer segments?*
**Answer:**
> **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| github_jupyter |
<h1 align="center"> Tuples </h1>
Tuples are sequences, just like lists. The differences between tuples and lists are, the tuples cannot be changed (immutable) unlike lists (mutable). <br> Tuples use parentheses, whereas lists use square brackets.
# Initialize a Tuple
There are two ways to initialize an empty tuple. You can initialize an empty tuple by having () with no values in them.
```
# Way 1
emptyTuple = ()
```
You can also initialize an empty tuple by using the <b>tuple</b> function.
```
# Way 2
emptyTuple = tuple()
```
A tuple with values can be initialized by making a sequence of values separated by commas.
```
# way 1
z = (3, 7, 4, 2)
# way 2 (tuples can also can be created without parenthesis)
z = 3, 7, 4, 2
```
It is important to keep in mind that if you want to create a tuple containing only one value, you need a trailing comma after your item.
```
# tuple with one value
tup1 = ('Michael',)
# tuple with one value
tup2 = 'Michael',
# This is a string, NOT a tuple.
notTuple = ('Michael')
```
# Accessing Values in Tuples
Each value in a tuple has an assigned index value. It is important to note that python is a zero indexed based language. All this means is that the first value in the tuple is at index 0.
```
# Initialize a tuple
z = (3, 7, 4, 2)
# Access the first item of a tuple at index 0
print(z[0])
```
Python also supports negative indexing. Negative indexing starts from the end of the tuple. It can sometimes be more convenient to use negative indexing to get the last item in a tuple because you don't have to know the length of a tuple to access the last item.
```
# print last item in the tuple
print(z[-1])
```
As a reminder, you could also access the same item using positive indexes (as seen below).
```
print(z[3])
```
# Tuple slices
Slice operations return a new tuple containing the requested items. Slices are good for getting a subset of values in your tuple. For the example code below, it will return a tuple with the items from index 0 up to and not including index 2.
```
# Initialize a tuple
z = (3, 7, 4, 2)
# first index is inclusive (before the :) and last (after the :) is not.
print(z[0:2])
# everything up to but not including index 3
print(z[:3])
```
You can even make slices with negative indexes.
```
print(z[-4:-1])
```
# Tuples are Immutable
Tuples are immutable which means that after initializing a tuple, it is impossible to update individual items in a tuple. As you can see in the code below, you cannot update or change the values of tuple items (this is different from [Python Lists](https://hackernoon.com/python-basics-6-lists-and-list-manipulation-a56be62b1f95) which are mutable).
```
z = (3, 7, 4, 2)
z[1] = "fish"
```
Even though tuples are immutable, it is possible to take portions of existing tuples to create new tuples as the following example demonstrates.
```
# Initialize tuple
tup1 = ('Python', 'SQL')
# Initialize another Tuple
tup2 = ('R',)
# Create new tuple based on existing tuples
new_tuple = tup1 + tup2;
print(new_tuple)
```
# Tuple Methods
Before starting this section, let's first initialize a tuple.
```
# Initialize a tuple
animals = ('lama', 'sheep', 'lama', 48)
```
## index method
The index method returns the first index at which a value occurs.
```
print(animals.index('lama'))
```
## count method
The count method returns the number of times a value occurs in a tuple.
```
print(animals.count('lama'))
```
# Iterate through a Tuple
You can iterate through the items of a tuple by using a for loop.
```
for item in ('lama', 'sheep', 'lama', 48):
print(item)
```
# Tuple Unpacking
Tuples are useful for sequence unpacking.
```
x, y = (7, 10);
print("Value of x is {}, the value of y is {}.".format(x, y))
```
# Enumerate
The enumerate function returns a tuple containing a count for every iteration (from start which defaults to 0) and the values obtained from iterating over a sequence:
```
friends = ('Steve', 'Rachel', 'Michael', 'Monica')
for index, friend in enumerate(friends):
print(index,friend)
```
# Advantages of Tuples over Lists
Lists and tuples are standard Python data types that store values in a sequence. A tuple is <b>immutable</b> whereas a list is <b>mutable</b>. Here are some other advantages of tuples over lists (partially from [Stack Overflow](https://stackoverflow.com/questions/1708510/python-list-vs-tuple-when-to-use-each))
<b>Tuples are faster than lists</b>. If you're defining a constant set of values and all you're ever going to do with it is iterate through it, use a tuple instead of a list. The performance difference can be partially measured using the timeit library which allows you to time your Python code. The code below runs the code for each approach 1 million times and outputs the overall time it took in seconds.
```
import timeit
print('Tuple time: ', timeit.timeit('x=(1,2,3,4,5,6,7,8,9,10,11,12)', number=1000000))
print('List time: ', timeit.timeit('x=[1,2,3,4,5,6,7,8,9,10,11,12]', number=1000000))
```
Some tuples can be used as dictionary keys (specifically, tuples that contain immutable values like strings, numbers, and other tuples). Lists can never be used as dictionary keys, because lists are not immutable (you can learn more about dictionaries [here](https://hackernoon.com/python-basics-10-dictionaries-and-dictionary-methods-4e9efa70f5b9)).
## Tuples can be dictionary keys
```
bigramsTupleDict = {('this', 'is'): 23,
('is', 'a'): 12,
('a', 'sentence'): 2}
print(bigramsTupleDict)
```
## Lists can NOT be dictionary keys
```
bigramsListDict = {['this', 'is']: 23,
['is', 'a']: 12,
['a', 'sentence']: 2}
print(bigramsListDict)
```
# Tuples can be values in a set
```
graphicDesigner = {('this', 'is'),
('is', 'a'),
('a', 'sentence')}
print(graphicDesigner)
```
# Lists can NOT be values in a set
```
graphicDesigner = {['this', 'is'],
['is', 'a'],
['a', 'sentence']}
print(graphicDesigner)
```
### Task: Generating Fibonacci Sequence in Python
Fibonacci sequence is an integer sequence characterized by the fact that every number after the first two is the sum of the two preceding ones. By definition, the first two numbers in the Fibonacci sequence are either 1 and 1 (<b>which is how I like to code it</b>), or 0 and 1, depending on the chosen starting point of the sequence, and each subsequent number is the sum of the previous two.
```
print(1, 1, 2, 3, 5, 8, 13, 21, 34, 55)
```
1. Using looping technique, write a Python program which prints out the first 10 Fibonacci numbers
```
# Note, there are better ways to code this which I will go over in later videos
a,b = 1,1
for i in range(10):
print("Fib(a): ", a, "b is: ", b)
a,b = b,a+b
```
**if this tutorial doesn't cover what you are looking for, please leave a comment on the youtube video and I will try to cover what you are interested in. (Please subscribe if you can!)**
https://www.youtube.com/watch?v=gUHeaQ0qZaw
| github_jupyter |
# Introduction to Deep Learning with PyTorch
In this notebook, you'll get introduced to [PyTorch](http://pytorch.org/), a framework for building and training neural networks. PyTorch in a lot of ways behaves like the arrays you love from Numpy. These Numpy arrays, after all, are just tensors. PyTorch takes these tensors and makes it simple to move them to GPUs for the faster processing needed when training neural networks. It also provides a module that automatically calculates gradients (for backpropagation!) and another module specifically for building neural networks. All together, PyTorch ends up being more coherent with Python and the Numpy/Scipy stack compared to TensorFlow and other frameworks.
## Neural Networks
Deep Learning is based on artificial neural networks which have been around in some form since the late 1950s. The networks are built from individual parts approximating neurons, typically called units or simply "neurons." Each unit has some number of weighted inputs. These weighted inputs are summed together (a linear combination) then passed through an activation function to get the unit's output.
<img src="assets/simple_neuron.png" width=400px>
Mathematically this looks like:
$$
\begin{align}
y &= f(w_1 x_1 + w_2 x_2 + b) \\
y &= f\left(\sum_i w_i x_i \right)
\end{align}
$$
With vectors this is the dot/inner product of two vectors:
$$
h = \begin{bmatrix}
x_1 \, x_2 \cdots x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_1 \\
w_2 \\
\vdots \\
w_n
\end{bmatrix}
$$
### Stack them up!
We can assemble these unit neurons into layers and stacks, into a network of neurons. The output of one layer of neurons becomes the input for the next layer. With multiple input units and output units, we now need to express the weights as a matrix.
<img src='assets/multilayer_diagram_weights.png' width=450px>
We can express this mathematically with matrices again and use matrix multiplication to get linear combinations for each unit in one operation. For example, the hidden layer ($h_1$ and $h_2$ here) can be calculated
$$
\vec{h} = [h_1 \, h_2] =
\begin{bmatrix}
x_1 \, x_2 \cdots \, x_n
\end{bmatrix}
\cdot
\begin{bmatrix}
w_{11} & w_{12} \\
w_{21} &w_{22} \\
\vdots &\vdots \\
w_{n1} &w_{n2}
\end{bmatrix}
$$
The output for this small network is found by treating the hidden layer as inputs for the output unit. The network output is expressed simply
$$
y = f_2 \! \left(\, f_1 \! \left(\vec{x} \, \mathbf{W_1}\right) \mathbf{W_2} \right)
$$
## Tensors
It turns out neural network computations are just a bunch of linear algebra operations on *tensors*, a generalization of matrices. A vector is a 1-dimensional tensor, a matrix is a 2-dimensional tensor, an array with three indices is a 3-dimensional tensor (RGB color images for example). The fundamental data structure for neural networks are tensors and PyTorch (as well as pretty much every other deep learning framework) is built around tensors.
<img src="assets/tensor_examples.svg" width=600px>
With the basics covered, it's time to explore how we can use PyTorch to build a simple neural network.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import numpy as np
import torch
import helper
```
First, let's see how we work with PyTorch tensors. These are the fundamental data structures of neural networks and PyTorch, so it's imporatant to understand how these work.
```
x = torch.rand(3, 2)
x
y = torch.ones(x.size())
y
z = x + y
z
```
In general PyTorch tensors behave similar to Numpy arrays. They are zero indexed and support slicing.
```
z[0]
z[:, 1:]
```
Tensors typically have two forms of methods, one method that returns another tensor and another method that performs the operation in place. That is, the values in memory for that tensor are changed without creating a new tensor. In-place functions are always followed by an underscore, for example `z.add()` and `z.add_()`.
```
# Return a new tensor z + 1
z.add(1)
# z tensor is unchanged
z
# Add 1 and update z tensor in-place
z.add_(1)
# z has been updated
z
```
### Reshaping
Reshaping tensors is a really common operation. First to get the size and shape of a tensor use `.size()`. Then, to reshape a tensor, use `.resize_()`. Notice the underscore, reshaping is an in-place operation.
```
z.size()
z.resize_(2, 3)
z
```
## Numpy to Torch and back
Converting between Numpy arrays and Torch tensors is super simple and useful. To create a tensor from a Numpy array, use `torch.from_numpy()`. To convert a tensor to a Numpy array, use the `.numpy()` method.
```
a = np.random.rand(4,3)
a
b = torch.from_numpy(a)
b
b.numpy()
```
The memory is shared between the Numpy array and Torch tensor, so if you change the values in-place of one object, the other will change as well.
```
# Multiply PyTorch Tensor by 2, in place
b.mul_(2)
# Numpy array matches new values from Tensor
a
```
| github_jupyter |
<a href="https://colab.research.google.com/github/sayakpaul/robustness-vit/blob/master/imagenet_results/imagenet_9/ImageNet_9.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
The weights inside `timm` (the library used here for loading the models) were converted from the official model weights mentioned below.
**Source**:
* BiT: https://tfhub.dev/google/collections/bit/1
* ViT: https://github.com/google-research/vision_transformer/
```
!pip install -q timm
!git clone https://github.com/MadryLab/backgrounds_challenge
%cd backgrounds_challenge
!wget -q https://github.com/MadryLab/backgrounds_challenge/releases/download/data/backgrounds_challenge_data.tar.gz
!tar xf backgrounds_challenge_data.tar.gz
!python challenge_eval.py -h
```
Before running the scripts below, you would need to ajust them to account for the correct `mean` and `std` (both should be [0.5, 0.5, 0.5]). Also, there may be some problems stemming from flashing data and models to the right device. When that happens simply call `.to()` besides the corresponding variable and set the right device.
## BiT
```
!python challenge_eval.py --model resnetv2_101x3_bitm --data-path bg_challenge
!python in9_eval.py --eval-dataset 'original' --model resnetv2_101x3_bitm --data-path bg_challenge
!python in9_eval.py --eval-dataset 'mixed_same' --model resnetv2_101x3_bitm --data-path bg_challenge
!python in9_eval.py --eval-dataset 'mixed_rand' --model resnetv2_101x3_bitm --data-path bg_challenge
```
## ViT
```shell
(base) jupyter@tensorflow24:~/backgrounds_challenge$ python challenge_eval.py --model vit_large_patch16_224 --data-path bg_challenge
==> Preparing dataset ImageNet9..
At image 0 for class 00_dog, used 0.00 since the last print statement.
Up until now, have 0/0 vulnerable foregrounds.
At image 50 for class 00_dog, used 424.84 since the last print statement.
Up until now, have 23/50 vulnerable foregrounds.
At image 100 for class 00_dog, used 348.99 since the last print statement.
Up until now, have 57/100 vulnerable foregrounds.
At image 150 for class 00_dog, used 357.59 since the last print statement.
Up until now, have 87/150 vulnerable foregrounds.
At image 200 for class 00_dog, used 371.92 since the last print statement.
Up until now, have 116/200 vulnerable foregrounds.
At image 250 for class 00_dog, used 383.16 since the last print statement.
Up until now, have 142/250 vulnerable foregrounds.
At image 300 for class 00_dog, used 382.41 since the last print statement.
Up until now, have 169/300 vulnerable foregrounds.
At image 350 for class 00_dog, used 441.94 since the last print statement.
Up until now, have 189/350 vulnerable foregrounds.
At image 400 for class 00_dog, used 385.74 since the last print statement.
Up until now, have 212/400 vulnerable foregrounds.
At image 0 for class 01_bird, used 0.00 since the last print statement.
Up until now, have 238/450 vulnerable foregrounds.
At image 50 for class 01_bird, used 206.51 since the last print statement.
Up until now, have 284/500 vulnerable foregrounds.
At image 100 for class 01_bird, used 107.08 since the last print statement.
Up until now, have 333/550 vulnerable foregrounds.
At image 150 for class 01_bird, used 156.13 since the last print statement.
Up until now, have 380/600 vulnerable foregrounds.
At image 200 for class 01_bird, used 254.28 since the last print statement.
Up until now, have 419/650 vulnerable foregrounds.
At image 250 for class 01_bird, used 198.04 since the last print statement.
Up until now, have 465/700 vulnerable foregrounds.
At image 300 for class 01_bird, used 150.47 since the last print statement.
Up until now, have 513/750 vulnerable foregrounds.
At image 350 for class 01_bird, used 174.16 since the last print statement.
Up until now, have 559/800 vulnerable foregrounds.
At image 400 for class 01_bird, used 173.52 since the last print statement.
Up until now, have 604/850 vulnerable foregrounds.
At image 0 for class 02_wheeled vehicle, used 0.00 since the last print statement.
Up until now, have 651/900 vulnerable foregrounds.
At image 50 for class 02_wheeled vehicle, used 188.95 since the last print statement.
Up until now, have 695/950 vulnerable foregrounds.
At image 100 for class 02_wheeled vehicle, used 217.13 since the last print statement.
Up until now, have 739/1000 vulnerable foregrounds.
At image 150 for class 02_wheeled vehicle, used 295.14 since the last print statement.
Up until now, have 770/1050 vulnerable foregrounds.
At image 200 for class 02_wheeled vehicle, used 175.33 since the last print statement.
Up until now, have 816/1100 vulnerable foregrounds.
At image 250 for class 02_wheeled vehicle, used 207.97 since the last print statement.
Up until now, have 860/1150 vulnerable foregrounds.
At image 300 for class 02_wheeled vehicle, used 153.01 since the last print statement.
Up until now, have 906/1200 vulnerable foregrounds.
At image 350 for class 02_wheeled vehicle, used 129.98 since the last print statement.
Up until now, have 953/1250 vulnerable foregrounds.
At image 400 for class 02_wheeled vehicle, used 197.14 since the last print statement.
Up until now, have 994/1300 vulnerable foregrounds.
At image 0 for class 03_reptile, used 0.00 since the last print statement.
Up until now, have 1043/1350 vulnerable foregrounds.
At image 50 for class 03_reptile, used 249.98 since the last print statement.
Up until now, have 1080/1400 vulnerable foregrounds.
At image 100 for class 03_reptile, used 252.57 since the last print statement.
Up until now, have 1118/1450 vulnerable foregrounds.
At image 150 for class 03_reptile, used 195.30 since the last print statement.
Up until now, have 1159/1500 vulnerable foregrounds.
At image 200 for class 03_reptile, used 186.98 since the last print statement.
Up until now, have 1204/1550 vulnerable foregrounds.
At image 250 for class 03_reptile, used 255.87 since the last print statement.
Up until now, have 1247/1600 vulnerable foregrounds.
At image 300 for class 03_reptile, used 206.11 since the last print statement.
Up until now, have 1289/1650 vulnerable foregrounds.
At image 350 for class 03_reptile, used 190.83 since the last print statement.
Up until now, have 1332/1700 vulnerable foregrounds.
At image 400 for class 03_reptile, used 244.35 since the last print statement.
Up until now, have 1370/1750 vulnerable foregrounds.
At image 0 for class 04_carnivore, used 0.00 since the last print statement.
Up until now, have 1406/1800 vulnerable foregrounds.
At image 50 for class 04_carnivore, used 239.45 since the last print statement.
Up until now, have 1443/1850 vulnerable foregrounds.
At image 100 for class 04_carnivore, used 173.52 since the last print statement.
Up until now, have 1486/1900 vulnerable foregrounds.
At image 150 for class 04_carnivore, used 294.68 since the last print statement.
Up until now, have 1523/1950 vulnerable foregrounds.
At image 200 for class 04_carnivore, used 275.61 since the last print statement.
Up until now, have 1558/2000 vulnerable foregrounds.
At image 250 for class 04_carnivore, used 328.48 since the last print statement.
Up until now, have 1589/2050 vulnerable foregrounds.
At image 300 for class 04_carnivore, used 310.70 since the last print statement.
Up until now, have 1621/2100 vulnerable foregrounds.
At image 350 for class 04_carnivore, used 285.96 since the last print statement.
Up until now, have 1653/2150 vulnerable foregrounds.
At image 400 for class 04_carnivore, used 311.26 since the last print statement.
Up until now, have 1688/2200 vulnerable foregrounds.
At image 0 for class 05_insect, used 0.00 since the last print statement.
Up until now, have 1722/2250 vulnerable foregrounds.
At image 50 for class 05_insect, used 127.48 since the last print statement.
Up until now, have 1769/2300 vulnerable foregrounds.
At image 100 for class 05_insect, used 90.76 since the last print statement.
Up until now, have 1819/2350 vulnerable foregrounds.
At image 150 for class 05_insect, used 115.42 since the last print statement.
Up until now, have 1867/2400 vulnerable foregrounds.
At image 200 for class 05_insect, used 91.27 since the last print statement.
Up until now, have 1917/2450 vulnerable foregrounds.
At image 250 for class 05_insect, used 96.41 since the last print statement.
Up until now, have 1966/2500 vulnerable foregrounds.
At image 300 for class 05_insect, used 109.93 since the last print statement.
Up until now, have 2014/2550 vulnerable foregrounds.
At image 350 for class 05_insect, used 118.48 since the last print statement.
Up until now, have 2063/2600 vulnerable foregrounds.
At image 400 for class 05_insect, used 129.14 since the last print statement.
Up until now, have 2111/2650 vulnerable foregrounds.
At image 150 for class 06_musical instrument, used 123.75 since the last print statement.
Up until now, have 2302/2850 vulnerable foregrounds.
At image 200 for class 06_musical instrument, used 215.24 since the last print statement.
Up until now, have 2343/2900 vulnerable foregrounds.
At image 250 for class 06_musical instrument, used 199.52 since the last print statement.
Up until now, have 2385/2950 vulnerable foregrounds.
At image 300 for class 06_musical instrument, used 152.78 since the last print statement.
Up until now, have 2431/3000 vulnerable foregrounds.
At image 350 for class 06_musical instrument, used 280.11 since the last print statement.
Up until now, have 2469/3050 vulnerable foregrounds.
At image 400 for class 06_musical instrument, used 175.96 since the last print statement.
Up until now, have 2512/3100 vulnerable foregrounds.
At image 0 for class 07_primate, used 0.00 since the last print statement.
Up until now, have 2552/3150 vulnerable foregrounds.
At image 50 for class 07_primate, used 411.45 since the last print statement.
Up until now, have 2576/3200 vulnerable foregrounds.
At image 100 for class 07_primate, used 388.60 since the last print statement.
Up until now, have 2601/3250 vulnerable foregrounds.
At image 150 for class 07_primate, used 319.40 since the last print statement.
Up until now, have 2630/3300 vulnerable foregrounds.
At image 200 for class 07_primate, used 322.67 since the last print statement.
Up until now, have 2664/3350 vulnerable foregrounds.
At image 250 for class 07_primate, used 326.97 since the last print statement.
Up until now, have 2695/3400 vulnerable foregrounds.
At image 300 for class 07_primate, used 355.24 since the last print statement.
Up until now, have 2722/3450 vulnerable foregrounds.
At image 350 for class 07_primate, used 192.03 since the last print statement.
Up until now, have 2765/3500 vulnerable foregrounds.
At image 350 for class 08_fish, used 160.69 since the last print statement.
Up until now, have 3145/3950 vulnerable foregrounds.
At image 400 for class 08_fish, used 113.32 since the last print statement.
Up until now, have 3194/4000 vulnerable foregrounds.
Evaluation complete
Summary: 3239/4050 (79.98%) are vulnerable foregrounds.
```
```
!python in9_eval.py --eval-dataset 'original' --model vit_large_patch16_224 --data-path bg_challenge
!python in9_eval.py --eval-dataset 'mixed_same' --model vit_large_patch16_224 --data-path bg_challenge
!python in9_eval.py --eval-dataset 'mixed_rand' --model vit_large_patch16_224 --data-path bg_challenge
```
| github_jupyter |
ERROR: type should be string, got "https://discourse.julialang.org/t/ode-solvers-why-is-matlab-ode45-uncannily-stable/63052/15\n\n```\nusing DifferentialEquations\nusing Plots\nfunction wk5!(dP, P, params, t)\n\n #=\n\n The 5-Element WK with serial L\n set Ls = 0 for 4-Element WK parallel\n\n Formulation for DifferentialEquations.jl\n\n P: solution vector (pressures p1 and p2)\n params: parameter tuple\n (Rc, Rp, C, Lp, Ls, I, q)\n\n\n I need to find a way to tranfer the function name as well\n for the time being we have to have the function in \"I\"\n\n =#\n\n # Split parameter tuple:\n Rc, Rp, C, Lp, Ls, I, q = params\n\n dP[1] = (\n -Rc / Lp * P[1]\n + (Rc / Lp - 1 / Rp / C) * P[2]\n + Rc * (1 + Ls / Lp) * didt(I, t, q)\n + I(t, q) / C\n )\n\n dP[2] = -1 / Rp / C * P[2] + I(t, q) / C\n\n return\n\nend\n# Generic Input Waveform\n# max volume flow in ml/s\nmax_i = 425\n\n# min volume flow in m^3/s\nmin_i = 0.0\n\nT = 0.9\n\n# Syst. Time in s\nsystTime = 2 / 5 * T\n\n# Dicrotic notch time\ndicrTime = 0.02\n\nq_generic = (max_i, min_i, T, systTime, dicrTime)\n\nfunction I_generic(t, q_generic)\n max_i, min_i, T, systTime, dicrTime = q_generic\n # implicit conditional using boolean multiplicator\n # sine waveform\n (\n (max_i - min_i) * sin(pi / systTime * (t % T))\n * (t % T < (systTime + dicrTime) )\n + min_i\n )\nend\nfunction didt(I, t, q)\n dt = 1e-3;\n didt = (I(t+dt, q) - I(t-dt, q)) / (2 * dt)\n return didt\nend\nplot(range(0, 2; length=2000), t -> didt(I_generic, t, q_generic); label=\"didt\")\n# Initial condition and time span\nP0 = [0.0, 0.0]\ntspan = (0.0, 30.0)\n\n# Set parameters for Windkessel Model\nRc = 0.033\nRp = 0.6\nC = 1.25\n# L for serial model!\nLs = 0.01\n# L for parallel\nLp = 0.02\n\nI = I_generic\nq = q_generic\n\np5 = (Rc, Rp, C, Lp, Ls, I, q)\n\nproblem = ODEProblem(wk5!, P0, tspan, p5)\n\ndtmax = 1e-4\n\n@time solutionTsit = solve(problem); GC.gc()\n@time solutionTsitLowdt = solve(problem, Tsit5(), dtmax=dtmax); GC.gc()\n@time solutionBS3 = solve(problem, BS3()); GC.gc()\n@time solutionBS3Lowdt = solve(problem, BS3(), dtmax=dtmax); GC.gc()\n@time solutionDP5 = solve(problem,DP5()); GC.gc()\n@time solutionDP5Lowdt = solve(problem,DP5(), dtmax=dtmax); GC.gc()\n@time solutionStiff = solve(problem, alg_hints=[:stiff]); GC.gc()\n@time solutionTsit = solve(problem); GC.gc()\n@time solutionTsitLowdt = solve(problem, Tsit5(), dtmax=dtmax); GC.gc()\n@time solutionBS3 = solve(problem, BS3()); GC.gc()\n@time solutionBS3Lowdt = solve(problem, BS3(), dtmax=dtmax); GC.gc()\n@time solutionDP5 = solve(problem,DP5()); GC.gc()\n@time solutionDP5Lowdt = solve(problem,DP5(), dtmax=dtmax); GC.gc()\n@time solutionStiff = solve(problem, alg_hints=[:stiff]); GC.gc()\n@show length(solutionTsit.t)\n@show length(solutionTsitLowdt.t)\n@show length(solutionBS3.t)\n@show length(solutionBS3Lowdt.t)\n@show length(solutionDP5.t)\n@show length(solutionDP5Lowdt.t)\n@show length(solutionStiff.t);\na, b = 0, 2\n\nplot()\n#plot!(t -> solutionTsit(t; idxs=1), a, b; label=\"Tsit\")\nplot!(t -> solutionTsitLowdt(t; idxs=1), a, b; label=\"TsitLowdt\")\n#plot!(t -> solutionBS3(t; idxs=1), a, b; label=\"BS3\", ls=:dash)\nplot!(t -> solutionBS3Lowdt(t; idxs=1), a, b; label=\"BS3Lowdt\", ls=:dash)\n#plot!(t -> solutionDP5(t; idxs=1), a, b; label=\"DP5\", ls=:dashdot)\nplot!(t -> solutionDP5Lowdt(t; idxs=1), a, b; label=\"DP5Lowdt\", ls=:dashdot)\nplot!(t -> solutionStiff(t; idxs=1), a, b; label=\"Stiff\", ls=:dot, lw=1.5)\na, b = 0, 2\n\nplot()\nplot!(t -> solutionTsit(t; idxs=1), a, b; label=\"Tsit\")\n#plot!(t -> solutionTsitLowdt(t; idxs=1), a, b; label=\"TsitLowdt\")\nplot!(t -> solutionBS3(t; idxs=1), a, b; label=\"BS3\", ls=:dash)\n#plot!(t -> solutionBS3Lowdt(t; idxs=1), a, b; label=\"BS3Lowdt\", ls=:dash)\nplot!(t -> solutionDP5(t; idxs=1), a, b; label=\"DP5\", ls=:dashdot)\n#plot!(t -> solutionDP5Lowdt(t; idxs=1), a, b; label=\"DP5Lowdt\", ls=:dashdot)\nplot!(t -> solutionStiff(t; idxs=1), a, b; label=\"Stiff\", ls=:dot, lw=1.5)\na, b = 28, 30\n\nplot()\nplot!(t -> solutionTsit(t; idxs=1), a, b; label=\"Tsit\")\n#plot!(t -> solutionTsitLowdt(t; idxs=1), a, b; label=\"TsitLowdt\")\nplot!(t -> solutionBS3(t; idxs=1), a, b; label=\"BS3\", ls=:dash)\n#plot!(t -> solutionBS3Lowdt(t; idxs=1), a, b; label=\"BS3Lowdt\", ls=:dash)\nplot!(t -> solutionDP5(t; idxs=1), a, b; label=\"DP5\", ls=:dashdot)\n#plot!(t -> solutionDP5Lowdt(t; idxs=1), a, b; label=\"DP5Lowdt\", ls=:dashdot)\nplot!(t -> solutionStiff(t; idxs=1), a, b; label=\"Stiff\", ls=:dot, lw=1.5)\n```\n\n" | github_jupyter |
```
import pandas as pd
import numpy as np
h1_seasons = ['2009',
'2010',
'2009Pan',
'2014',
'2016']
df = pd.read_csv('../raw_data/cases_of_dominant_subtype_by_birth_year.csv')
max_birth_years = pd.read_csv('../raw_data/max_birth_years.csv', index_col='Season')
h1_cases = df[df.Season.isin(h1_seasons)].copy()
h3_cases = df[~df.Season.isin(h1_seasons)].copy()
final_df = pd.DataFrame(columns=['birth_year',
'I_obs_h1',
'I_vac_h1',
'I_obs_h3',
'I_vac_h3',
'season'])
h1_agg = h1_cases.groupby(['Season', 'Birth_year', 'Vaccination_status']).sum().Count
h3_agg = h3_cases.groupby(['Season', 'Birth_year', 'Vaccination_status']).sum().Count
index = 0
for season in set(df.Season):
if season in h1_seasons:
aggdf = h1_agg
subtype = 'h1'
else:
aggdf = h3_agg
subtype = 'h3'
for byear in np.arange(1918, max_birth_years.loc[season, 'max_year'] + 1):
try:
vac_cases = aggdf.loc[(season, byear, 1), ]
except KeyError:
vac_cases = 0
try:
unvac_cases = aggdf.loc[(season, byear, 0), ]
except KeyError:
unvac_cases = 0
final_df.loc[index, 'birth_year'] = byear
final_df.loc[index, 'I_obs_%s'%subtype] = unvac_cases
final_df.loc[index, 'I_vac_%s'%subtype] = vac_cases
final_df.loc[index, 'season'] = season
index += 1
final_df = final_df.sort_values(['season', 'birth_year'])
final_df.to_csv('../data/standard_eligible_observed.csv', index=False)
df = pd.read_csv('../raw_data/cases_of_dominant_subtype_by_age.csv')
h1_cases = df[df.Season.isin(h1_seasons)].copy()
h3_cases = df[~df.Season.isin(h1_seasons)].copy()
final_df = pd.DataFrame(columns=['Age',
'I_obs_H1',
'I_vac_H1',
'I_obs_H3',
'I_vac_H3',
'Season'])
h1_agg = h1_cases.groupby(['Season', 'Age', 'Vaccination_status']).sum().Count
h3_agg = h3_cases.groupby(['Season', 'Age', 'Vaccination_status']).sum().Count
index = 0
for season, seasondf in df.groupby('Season'):
if season in h1_seasons:
aggdf = h1_agg
subtype = 'H1'
else:
aggdf = h3_agg
subtype = 'H3'
for age in set(seasondf.Age):
try:
vac_cases = aggdf.loc[(season, age, 1), ]
except KeyError:
vac_cases = 0
try:
unvac_cases = aggdf.loc[(season, age, 0), ]
except KeyError:
unvac_cases = 0
final_df.loc[index, 'Age'] = age
final_df.loc[index, 'I_obs_%s'%subtype] = unvac_cases
final_df.loc[index, 'I_vac_%s'%subtype] = vac_cases
final_df.loc[index, 'Season'] = season
index += 1
final_df = final_df.sort_values(['Season', 'Age'])
final_df.to_csv('../data/standard_eligible_observed_by_age.csv', index=False)
```
| github_jupyter |
# Convolutional Neural Networks
---
In this notebook, we train a **CNN** to classify images from the CIFAR-10 database.
The images in this database are small color images that fall into one of ten classes; some example images are pictured below.
<img src='notebook_ims/cifar_data.png' width=70% height=70% />
### Test for [CUDA](http://pytorch.org/docs/stable/cuda.html)
Since these are larger (32x32x3) images, it may prove useful to speed up your training time by using a GPU. CUDA is a parallel computing platform and CUDA Tensors are the same as typical Tensors, only they utilize GPU's for computation.
```
import torch
import numpy as np
# check if CUDA is available
train_on_gpu = torch.cuda.is_available()
if not train_on_gpu:
print('CUDA is not available. Training on CPU ...')
else:
print('CUDA is available! Training on GPU ...')
```
---
## Load the [Data](http://pytorch.org/docs/stable/torchvision/datasets.html)
Downloading may take a minute. We load in the training and test data, split the training data into a training and validation set, then create DataLoaders for each of these sets of data.
```
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
# number of subprocesses to use for data loading
num_workers = 0
# how many samples per batch to load
batch_size = 20
# percentage of training set to use as validation
valid_size = 0.2
# convert data to a normalized torch.FloatTensor
transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
##### alternatively with data augmentation
transform_aug = transforms.Compose([
transforms.RandomHorizontalFlip(), # randomly flip and rotate
transforms.RandomRotation(10),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
# choose the training and test datasets
train_data = datasets.CIFAR10('data', train=True,
download=True, transform=transform_aug)
test_data = datasets.CIFAR10('data', train=False,
download=True, transform=transform_test)
# obtain training indices that will be used for validation
num_train = len(train_data)
indices = list(range(num_train))
np.random.shuffle(indices)
split = int(np.floor(valid_size * num_train))
train_idx, valid_idx = indices[split:], indices[:split]
# define samplers for obtaining training and validation batches
train_sampler = SubsetRandomSampler(train_idx)
valid_sampler = SubsetRandomSampler(valid_idx)
# prepare data loaders (combine dataset and sampler)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=train_sampler, num_workers=num_workers)
valid_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size,
sampler=valid_sampler, num_workers=num_workers)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size,
num_workers=num_workers)
# specify the image classes
classes = ['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck']
```
### Visualize a Batch of Training Data
```
import matplotlib.pyplot as plt
%matplotlib inline
# helper function to un-normalize and display an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
plt.imshow(np.transpose(img, (1, 2, 0))) # convert from Tensor image
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
images = images.numpy() # convert images to numpy for display
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
# display 20 images
for idx in np.arange(20):
ax = fig.add_subplot(2, int(20/2), idx+1, xticks=[], yticks=[])
imshow(images[idx])
ax.set_title(classes[labels[idx]])
```
### View an Image in More Detail
Here, we look at the normalized red, green, and blue (RGB) color channels as three separate, grayscale intensity images.
```
rgb_img = np.squeeze(images[3])
channels = ['red channel', 'green channel', 'blue channel']
fig = plt.figure(figsize = (36, 36))
for idx in np.arange(rgb_img.shape[0]):
ax = fig.add_subplot(1, 3, idx + 1)
img = rgb_img[idx]
ax.imshow(img, cmap='gray')
ax.set_title(channels[idx])
width, height = img.shape
thresh = img.max()/2.5
for x in range(width):
for y in range(height):
val = round(img[x][y],2) if img[x][y] !=0 else 0
ax.annotate(str(val), xy=(y,x),
horizontalalignment='center',
verticalalignment='center', size=8,
color='white' if img[x][y]<thresh else 'black')
```
---
## Define the Network [Architecture](http://pytorch.org/docs/stable/nn.html)
This time, you'll define a CNN architecture. Instead of an MLP, which used linear, fully-connected layers, you'll use the following:
* [Convolutional layers](https://pytorch.org/docs/stable/nn.html#conv2d), which can be thought of as stack of filtered images.
* [Maxpooling layers](https://pytorch.org/docs/stable/nn.html#maxpool2d), which reduce the x-y size of an input, keeping only the most _active_ pixels from the previous layer.
* The usual Linear + Dropout layers to avoid overfitting and produce a 10-dim output.
A network with 2 convolutional layers is shown in the image below and in the code, and you've been given starter code with one convolutional and one maxpooling layer.
<img src='notebook_ims/2_layer_conv.png' height=50% width=50% />
#### TODO: Define a model with multiple convolutional layers, and define the feedforward network behavior.
The more convolutional layers you include, the more complex patterns in color and shape a model can detect. It's suggested that your final model include 2 or 3 convolutional layers as well as linear layers + dropout in between to avoid overfitting.
It's good practice to look at existing research and implementations of related models as a starting point for defining your own models. You may find it useful to look at [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py) to help decide on a final structure.
#### Output volume for a convolutional layer
To compute the output size of a given convolutional layer we can perform the following calculation (taken from [Stanford's cs231n course](http://cs231n.github.io/convolutional-networks/#layers)):
> We can compute the spatial size of the output volume as a function of the input volume size (W), the kernel/filter size (F), the stride with which they are applied (S), and the amount of zero padding used (P) on the border. The correct formula for calculating how many neurons define the output_W is given by `(W−F+2P)/S+1`.
For example for a 7x7 input and a 3x3 filter with stride 1 and pad 0 we would get a 5x5 output. With stride 2 we would get a 3x3 output.
```
### see here for the winning architecture:
# http://blog.kaggle.com/2015/01/02/cifar-10-competition-winners-interviews-with-dr-ben-graham-phil-culliton-zygmunt-zajac/
### see here for pytorch tutorial:
# https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py
#################### NOTE this ist version 1, the bigger model
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net1(nn.Module):
def __init__(self):
super(Net1, self).__init__()
# setup
num_classes = 10
drop_p = 0.5
# convolutional layer
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
self.conv3 = nn.Conv2d(32, 64, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# fully connected layer
self.fc1 = nn.Linear(1024, 256, bias=True)
self.fc2 = nn.Linear(256, 64, bias=True)
self.fc3 = nn.Linear(64, num_classes, bias=True)
# dropout
self.dropout = nn.Dropout(drop_p)
def forward(self, x):
# add sequence of convolutional and max pooling layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
# flatten image, keep batch size
x = x.view(x.shape[0], -1)
x = self.dropout(self.fc1(x))
x = self.dropout(self.fc2(x))
x = F.log_softmax(self.fc3(x), dim=1)
return x
##################### NOTE this is version 2, the smaller model
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net2(nn.Module):
def __init__(self):
super(Net2, self).__init__()
# setup
num_classes = 10
drop_p = 0.25
# convolutional layer
self.conv1 = nn.Conv2d(3, 8, 3, padding=1)
self.conv2 = nn.Conv2d(8, 16, 3, padding=1)
self.conv3 = nn.Conv2d(16, 32, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# fully connected layer
self.fc1 = nn.Linear(512, 128, bias=True)
self.fc2 = nn.Linear(128, 64, bias=True)
self.fc3 = nn.Linear(64, num_classes, bias=True)
# dropout
self.dropout = nn.Dropout(drop_p)
def forward(self, x):
# add sequence of convolutional and max pooling layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
# flatten image, keep batch size
x = x.view(x.shape[0], -1)
x = self.dropout(self.fc1(x))
x = self.dropout(self.fc2(x))
x = F.log_softmax(self.fc3(x), dim=1)
return x
##################### NOTE this is the official solution example, it is similar to version 1
# * Conv-layer sind wie bei mir
# * hat einen fc layer weniger als ich
# * hat einen dropout hinter dem letzten conv, das hatte ich nicht
# * hat eine relu hinter dem ersten fc, das hatte ich nicht
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
# convolutional layer
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
self.conv3 = nn.Conv2d(32, 64, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# fully connected layer
self.fc1 = nn.Linear(64 * 4 * 4, 500)
self.fc2 = nn.Linear(500, 10)
# dropout
self.dropout = nn.Dropout(0.25)
def forward(self, x):
# add sequence of convolutional and max pooling layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
# flatten image, keep batch size
x = x.view(-1, 64 * 4 * 4)
x = self.dropout(x)
x = F.relu(self.fc1(x))
x = self.dropout(x)
x = self.fc2(x)
return x
#################### NOTE this is version 3, after seeing the official solution
import torch.nn as nn
import torch.nn.functional as F
# define the CNN architecture
class Net3(nn.Module):
def __init__(self):
super(Net3, self).__init__()
# setup
num_classes = 10
drop_p = 0.5
# convolutional layer
self.conv1 = nn.Conv2d(3, 16, 3, padding=1)
self.conv2 = nn.Conv2d(16, 32, 3, padding=1)
self.conv3 = nn.Conv2d(32, 64, 3, padding=1)
self.conv4 = nn.Conv2d(64, 128, 3, padding=1)
# max pooling layer
self.pool = nn.MaxPool2d(2, 2)
# fully connected layer
self.fc1 = nn.Linear(128 * 2 * 2, 512, bias=True)
self.fc2 = nn.Linear(512, 256, bias=True)
self.fc3 = nn.Linear(256, num_classes, bias=True)
# dropout
self.dropout = nn.Dropout(drop_p)
def forward(self, x):
# add sequence of convolutional and max pooling layers
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = self.pool(F.relu(self.conv3(x)))
x = self.pool(F.relu(self.conv4(x)))
# flatten image, keep batch size
x = x.view(x.shape[0], -1)
x = self.dropout(x)
x = self.dropout(F.relu(self.fc1(x)))
x = self.dropout(F.relu(self.fc2(x)))
x = F.log_softmax(self.fc3(x), dim=1)
return x
# create a complete CNN
#model = Net1() # own bigger model
#model = Net2() # own smaller model
#model = Net() # official solution example
model = Net3() # own final version after seeing the solution example
print(model)
# move tensors to GPU if CUDA is available
if train_on_gpu:
model.cuda()
```
### Specify [Loss Function](http://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizer](http://pytorch.org/docs/stable/optim.html)
Decide on a loss and optimization function that is best suited for this classification task. The linked code examples from above, may be a good starting point; [this PyTorch classification example](https://github.com/pytorch/tutorials/blob/master/beginner_source/blitz/cifar10_tutorial.py) or [this, more complex Keras example](https://github.com/keras-team/keras/blob/master/examples/cifar10_cnn.py). Pay close attention to the value for **learning rate** as this value determines how your model converges to a small error.
#### TODO: Define the loss and optimizer and see how these choices change the loss over time.
```
import torch.optim as optim
# specify loss function
criterion = nn.NLLLoss() # requires SoftMax to be executed prior to the criterion
# NOTE: official solution needs other criterion:
# criterion = nn.CrossEntropyLoss() # includes SoftMax as first step
# specify optimizer
# version 1, dropout = 0.50, mostly with 20 to 30 epochs
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 70% (7029/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 72% (7212/10000)
optimizer = optim.SGD(model.parameters(), lr=0.025) # Test Accuracy (Overall): 71% (7172/10000)
# version 1, dropout = 0.25
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 71% (7185/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 71% (7121/10000)
optimizer = optim.SGD(model.parameters(), lr=0.025) # Test Accuracy (Overall): 71% (7174/10000)
# version 2, dropout = 0.50
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 68% (6894/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 68% (6810/10000)
# version 2, dropout = 0.25
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 67% (6719/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 68% (6802/10000)
# version 3, dropout = 0.50
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 70% (7063/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 74% (7446/10000)
# version 3, dropout = 0.25
optimizer = optim.Adam(model.parameters(), lr=0.001) # Test Accuracy (Overall): 71% (7103/10000)
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 74% (7463/10000)
# version 3, dropout = 0.25, with data augmentation, 35 epochs
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 77% (7702/10000)
# version 3, dropout = 0.25, with data augmentation, 60 epochs
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 79% (7925/10000)
# version 3 + dilation=2 in first 2 conv layers, dropout = 0.25, with data augmentation, 60 epochs
optimizer = optim.SGD(model.parameters(), lr=0.01) # Test Accuracy (Overall): 75% (7595/10000)
```
---
## Train the Network
Remember to look at how the training and validation loss decreases over time; if the validation loss ever increases it indicates possible overfitting.
```
# number of epochs to train the model
n_epochs = 30 # you may increase this number to train a final model
valid_loss_min = np.Inf # track change in validation loss
for epoch in range(1, n_epochs+1):
# keep track of training and validation loss
train_loss = 0.0
valid_loss = 0.0
###################
# train the model #
###################
model.train()
for data, target in train_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# clear the gradients of all optimized variables
optimizer.zero_grad()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# backward pass: compute gradient of the loss with respect to model parameters
loss.backward()
# perform a single optimization step (parameter update)
optimizer.step()
# update training loss
train_loss += loss.item()*data.size(0)
######################
# validate the model #
######################
model.eval()
for data, target in valid_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update average validation loss
valid_loss += loss.item()*data.size(0)
# calculate average losses
train_loss = train_loss/len(train_loader.dataset)
valid_loss = valid_loss/len(valid_loader.dataset)
# print training/validation statistics
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
epoch, train_loss, valid_loss))
# save model if validation loss has decreased
if valid_loss <= valid_loss_min:
print('Validation loss decreased ({:.6f} --> {:.6f}). Saving model ...'.format(
valid_loss_min,
valid_loss))
torch.save(model.state_dict(), 'model_cifar.pt')
valid_loss_min = valid_loss
```
### Load the Model with the Lowest Validation Loss
```
model.load_state_dict(torch.load('model_cifar.pt'))
```
---
## Test the Trained Network
Test your trained model on previously unseen data! A "good" result will be a CNN that gets around 70% (or more, try your best!) accuracy on these test images.
```
# track test loss
test_loss = 0.0
class_correct = list(0. for i in range(10))
class_total = list(0. for i in range(10))
model.eval()
# iterate over test data
for data, target in test_loader:
# move tensors to GPU if CUDA is available
if train_on_gpu:
data, target = data.cuda(), target.cuda()
# forward pass: compute predicted outputs by passing inputs to the model
output = model(data)
# calculate the batch loss
loss = criterion(output, target)
# update test loss
test_loss += loss.item()*data.size(0)
# convert output probabilities to predicted class
_, pred = torch.max(output, 1)
# compare predictions to true label
correct_tensor = pred.eq(target.data.view_as(pred))
correct = np.squeeze(correct_tensor.numpy()) if not train_on_gpu else np.squeeze(correct_tensor.cpu().numpy())
# calculate test accuracy for each object class
for i in range(batch_size):
label = target.data[i]
class_correct[label] += correct[i].item()
class_total[label] += 1
# average test loss
test_loss = test_loss/len(test_loader.dataset)
print('Test Loss: {:.6f}\n'.format(test_loss))
for i in range(10):
if class_total[i] > 0:
print('Test Accuracy of %5s: %2d%% (%2d/%2d)' % (
classes[i], 100 * class_correct[i] / class_total[i],
np.sum(class_correct[i]), np.sum(class_total[i])))
else:
print('Test Accuracy of %5s: N/A (no training examples)' % (classes[i]))
print('\nTest Accuracy (Overall): %2d%% (%2d/%2d)' % (
100. * np.sum(class_correct) / np.sum(class_total),
np.sum(class_correct), np.sum(class_total)))
```
### Question: What are your model's weaknesses and how might they be improved?
Answer:
* Adam Optimizer does not work ... try other learning rates?
* smaller learning rates work
* SGD works fine, but does not go beyond 71 % --> maybe simplify the architecture, use less params
* less params works worse
* best result: more complex model with dropout 0.5, SGD with lr=0.01 --> 72 % test accuracy
### Visualize Sample Test Results
```
# obtain one batch of test images
dataiter = iter(test_loader)
images, labels = dataiter.next()
images.numpy()
# move model inputs to cuda, if GPU available
if train_on_gpu:
images = images.cuda()
# get sample outputs
output = model(images)
# convert output probabilities to predicted class
_, preds_tensor = torch.max(output, 1)
preds = np.squeeze(preds_tensor.numpy()) if not train_on_gpu else np.squeeze(preds_tensor.cpu().numpy())
# plot the images in the batch, along with predicted and true labels
fig = plt.figure(figsize=(25, 4))
for idx in np.arange(20):
ax = fig.add_subplot(2, int(20/2), idx+1, xticks=[], yticks=[])
imshow(images[idx].cpu())
ax.set_title("{} ({})".format(classes[preds[idx]], classes[labels[idx]]),
color=("green" if preds[idx]==labels[idx].item() else "red"))
```
| github_jupyter |
```
# Import libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import re
import sys
from sklearn.cross_validation import cross_val_predict
from sklearn.pipeline import Pipeline
# used for train/test splits and cross validation
from sklearn.cross_validation import train_test_split
from sklearn.grid_search import GridSearchCV
# used to impute mean for data and standardize for computational stability
from sklearn.preprocessing import Imputer
from sklearn.preprocessing import StandardScaler
# logistic regression is our favourite model ever
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LassoCV
# used to calculate AUROC/accuracy
from sklearn import metrics
# used to create confusion matrix
from sklearn.metrics import confusion_matrix
from sklearn.cross_validation import cross_val_score
# gradient boosting - must download package https://github.com/dmlc/xgboost
#import xgboost as xgb
# default colours for prettier plots
col = [[0.9047, 0.1918, 0.1988],
[0.2941, 0.5447, 0.7494],
[0.3718, 0.7176, 0.3612],
[1.0000, 0.5482, 0.1000],
[0.4550, 0.4946, 0.4722],
[0.6859, 0.4035, 0.2412],
[0.9718, 0.5553, 0.7741],
[0.5313, 0.3359, 0.6523]];
marker = ['v','o','d','^','s','o','+']
ls = ['-','-','-','-','-','s','--','--']
%matplotlib inline
from __future__ import print_function
# read data from the got-data.ipynb file
# most of it is scrapped from the game of thrones tv show wiki
df = pd.read_csv('data/got_data.csv', sep=',', index_col=0)
# who do we not have data for?
idxNoData = (df['Season(s)'].isnull())
print('Never appeared in any season:')
print(df[idxNoData].index)
# delete people with no data - they have no scores and never appeared in any season
df = df.loc[~idxNoData]
# print out the data we have
df.sort_values('Total',axis=0,ascending=False).head(n=3).T
# visualize the data a bit
# each histogram shows the points scored across all characters in season 5
# originally sourced from scores.csv
# the scores are broken down into categories
txt_outcome = ['Killing','SexNudity','Insult','Drinking','Injury']
for txt in txt_outcome:
plt.figure(figsize=[6,6])
plt.hist(df[txt],bins=range(120),normed=True)
plt.plot([0,120], 1.0/df.shape[0]*np.ones(2), 'k--',lw=2 )
plt.title(txt,fontsize=16)
plt.xlabel('Points',fontsize=14)
plt.ylabel('Fraction of people',fontsize=14)
plt.show()
# define some useful subfunctions
# given a dataframe/regex phrase, this counts the number of times the regex appears
def count_words(s, phrase):
if s is np.nan:
return 0
else:
return len(re.findall(phrase,s, re.IGNORECASE))
# this calls the above function and adds the data to the given dataframe
def add_data(df_data, column_name, phrase, txt_col=None):
if txt_col is None:
txt_col = ['Background','Season 1','Season 2', 'Season 3', 'Season 4']
df_new = df[txt_col].applymap(lambda x: count_words(x,phrase))
df_data[column_name] = df_new[txt_col].sum(axis=1)
return df_data
# which text columns should we count words from
txt_col = ['Background','Season 1','Season 2', 'Season 3', 'Season 4']
# initialize the dataframe (we'll drop this column later)
idxData = np.ones(df.shape[0], dtype=bool)
df_data = df.loc[idxData, ['Total']]
# start adding data based on counting the frequency of words
# count the number of hyphens as a surrogate as the number of in-show family member
# in the data, the family members appear as "sister - Sansa Stark, father - Eddard Stark," .. etc
df_data = add_data(df_data, 'family_members',' - ', txt_col = ['Family'])
# number of seasons the character appeared in
df_data = add_data(df_data, 'number_of_seasons','[1-5]', txt_col = ['Season(s)'])
# various word "types" across all the data
df_data = add_data(df_data, 'violent_words','(knight|warrior|sword|axe|spear|kill|murder|fight|assassinate)')
df_data = add_data(df_data, 'sexy_words','(sex|naked|love|slept|nude|kiss)')
df_data = add_data(df_data, 'fun_words','(beer|wine|glass|drunk|inebriate)')
df_data = add_data(df_data, 'number_of_words','\w+')
df_data.drop('Total',axis=1,inplace=True) # get rid of the feature we initialized the dataframe with
df_data.head()
```
# logistic regression on target
Since the targets seem to be poorly distributed across the continuous scale (see histograms), we reformulate it from "how many points?" to "are they going to get any points at all?", i.e. a binary prediction task.
```
# decide which target to use
# 'Total','Killing','SexNudity','Injury','Insult','Drinking'
y_name = 'Killing'
# prep data
X = df_data[idxData].values
target = df[y_name].values
X = X[:,1:X.shape[1]]
y = target > 0
# workaround cross_val_predict returning 0s/1s - calling this makes it return probabilities
class proba_logreg(LogisticRegression):
def predict(self, X):
return LogisticRegression.predict_proba(self, X)
# cross-validation performance
mdl = "logreg"
model = LogisticRegression(fit_intercept=True)
estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
strategy="mean",
axis=0)),
("scaler", StandardScaler()),
("lr", model)])
scores = cross_val_score(estimator, X, y, scoring='roc_auc',cv=5)
print('{:10s} {:5g} [{:5g}, {:5g}]'.format("lr", np.mean(scores), np.min(scores), np.max(scores) ))
# plot a roc curve by getting cross-validation predictions
predicted = cross_val_predict(proba_logreg(), X, y, cv=10)
predicted = predicted[:,1]
plt.figure(figsize=[9,9])
ax = plt.gca()
fpr, tpr, thresholds = metrics.roc_curve(y, predicted, pos_label=1)
plt.plot(fpr, tpr, 'bo-',lw=2,markersize=12,
label=mdl + ' ' + '%0.3f' % metrics.auc(fpr, tpr))
plt.xlabel('False positive rate',fontsize=14)
plt.xlabel('True positive rate',fontsize=14)
plt.legend(loc='lower right')
plt.title(y_name,fontsize=16)
plt.show()
# originally we did try linear regressions.. but they're not as good, just look at the scatter plot
y = np.log10(target+1)
# if you need an estimator to handle missing data etc
#estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
# strategy="mean",
# axis=0)),
# ("scaler", StandardScaler()),
# ("lr", LinearRegression(fit_intercept=True))])
predicted = cross_val_predict(LinearRegression(fit_intercept=True), X, y, cv=10)
plt.figure(figsize=[9,9])
ax = plt.gca()
ax.scatter(y, predicted)
ax.plot([y.min(), y.max()], [y.min(), y.max()], 'k--', lw=4)
ax.set_xlabel('Measured')
ax.set_ylabel('Predicted')
plt.show()
plt.figure(figsize=[9,9])
plt.hist(y - predicted)
plt.title('Residuals')
plt.show()
```
| github_jupyter |
<img src='93.jpg' width=600>
```
class Solution:
def restoreIpAddresses(self, s: str):
self.res = []
self.dfs(s, [])
res = ['.'.join(x) for x in self.res]
return res
def dfs(self, s, temp):
if not s and len(temp) == 4 and temp not in self.res:
self.res.append(list(temp))
return
for step in range(1, 4): #字符串截取的长度只能是 1-3 位置
if len(s[step:]) > 9:
continue
sub_str = s[:step]
if sub_str and 0 <= int(sub_str) <= 255 and len(temp) < 4:
if len(sub_str) > 1 and sub_str[0]:
print(sub_str)
continue
temp.append(sub_str)
self.dfs(s[step:], temp)
temp.pop()
class Solution:
def restoreIpAddresses(self, s: str):
if len(s) < 4 or len(s) > 12:
return []
self.res = []
self.dfs(s, 0, '', 0)
return self.res
def dfs(self, s, start, path, count):
if count == 4 and start == len(s):
self.res.append(path[1:])
return
for end in [start+1, start+2, start+3]:
if end <= len(s) and 0 <= int(s[start:end]) <= 255 and str(int(s[start:end])) == s[start:end]:
self.dfs(s, end, path+'.'+s[start:end], count+1)
s_ = "010010"
solution = Solution()
solution.restoreIpAddresses(s_)
a = '1234'
print(a[2:])
class Solution:
def restoreIpAddresses(self, s: str):
if len(s) < 4 or len(s) > 12:
return []
self.res = []
self.dfs(s, 0, '', 0)
return self.res
def dfs(self, s, start, path, count):
if count == 4 and start == len(s):
self.res.append(path[1:])
return
for end in [start+1, start+2, start+3]:
a = end <= len(s)
b = a and 0 <= int(s[start:end]) <= 255 # 防止数字越界
c = a and b and str(int(s[start:end])) == s[start:end] # 防止前导0的出现
if c:
self.dfs(s, end, path + '.' + s[start:end], count+1)
s_ = "101023"
solution = Solution()
solution.restoreIpAddresses(s_)
class Solution:
def restoreIpAddresses(self, s: str):
if len(s) < 4 or len(s) > 12:
return []
self.res = set()
self.dfs(s, '', 0)
return list(self.res)
def dfs(self, s, temp, count):
if not s and count == 4:
print(temp)
self.res.add(temp[1:])
return
for i in range(1, 4):
a = i <= len(s)
sub_str = s[:i]
b = a and 0 <= int(sub_str) <= 255
c = a and b and str(int(sub_str)) == sub_str
if c:
self.dfs(s[i:], temp+'.'+sub_str, count+1)
class Solution:
def restoreIpAddresses(self, s: str):
if len(s) < 4 or len(s) > 12:
return []
self.res = []
self.dfs(s, [], 0)
return ['.'.join(x) for x in self.res]
def dfs(self, s, temp, count):
if not s and count == 4:
self.res.append(list(temp))
return
for i in range(1, 4):
if i <= len(s):
sub_str = s[:i]
if 0 <= int(sub_str) <= 255 and str(int(sub_str)) == sub_str:
temp.append(sub_str)
self.dfs(s[i:], temp, count+1)
temp.pop()
s_ = "101023"
solution = Solution()
solution.restoreIpAddresses(s_)
a = if 2 > 3
a = [1, 2]
a[0:1]
```
| github_jupyter |
# TensorFlow Visual Recognition Sample Application Part 4
## Tranfer Learning: Re-train MobileNet models with custom images
## Define the model metadata
```
import tensorflow as tf
import requests
models = {
"mobilenet": {
"base_url":"https://github.com/DTAIEB/Thoughtful-Data-Science/raw/master/chapter%206/Visual%20Recognition/mobilenet_v1_0.50_224",
"model_file_url": "frozen_graph.pb",
"label_file": "labels.txt",
"output_layer": "MobilenetV1/Predictions/Softmax",
"bottleneck_tensor_name": 'import/MobilenetV1/Predictions/Reshape:0',
"resized_input_tensor_name": 'import/input:0',
"input_width": 224,
"input_height": 224,
"input_depth": 3,
"bottleneck_tensor_size": 1001
}
}
# helper method for reading attributes from the model metadata
def get_model_attribute(model, key, default_value = None):
if key not in model:
if default_value is None:
raise Exception("Require model attribute {} not found".format(key))
return default_value
return model[key]
def ensure_dir_exists(dir_name):
if not os.path.exists(dir_name):
os.makedirs(dir_name)
return dir_name
```
## Helper methods for loading the graph and labels for a given model
```
# Helper method for resolving url relative to the selected model
def get_url(model, path):
return model["base_url"] + "/" + path
# Download the serialized model and create a TensorFlow graph
def load_graph(model):
graph = tf.Graph()
graph_def = tf.GraphDef()
graph_def.ParseFromString(
requests.get( get_url( model, model["model_file_url"] ) ).content
)
with graph.as_default():
tf.import_graph_def(graph_def)
return graph
# Load the labels
def load_labels(model, as_json = False):
labels = [line.rstrip() \
for line in requests.get( get_url( model, model["label_file"] ) ).text.split("\n") \
if line != ""]
if as_json:
return [{"index": item.split(":")[0], "label" : item.split(":")[1]} for item in labels]
return labels
```
## Use BeautifulSoup to scrape the images from a given url
```
from bs4 import BeautifulSoup as BS
import re
# return an array of all the images scraped from an html page
def get_image_urls(url):
# Instantiate a BeautifulSoup parser
soup = BS(requests.get(url).text, "html.parser")
# Local helper method for extracting url
def extract_url(val):
m = re.match(r"url\((.*)\)", val)
val = m.group(1) if m is not None else val
return "http:" + val if val.startswith("//") else val
# List comprehension that look for <img> elements and backgroud-image styles
return [extract_url(imgtag['src']) for imgtag in soup.find_all('img')] + [ \
extract_url(val.strip()) for key,val in \
[tuple(selector.split(":")) for elt in soup.select("[style]") \
for selector in elt["style"].strip(" ;").split(";")] \
if key.strip().lower()=='background-image' \
]
```
## Helper method for downloading an image into a temp file
```
import tempfile
def download_image(url):
response = requests.get(url, stream=True)
if response.status_code == 200:
with tempfile.NamedTemporaryFile(delete=False) as f:
for chunk in response.iter_content(2048):
f.write(chunk)
return f.name
else:
raise Exception("Unable to download image: {}".format(response.status_code))
```
## Decode an image into a tensor
```
# decode a given image into a tensor
def read_tensor_from_image_file(model, file_name):
file_reader = tf.read_file(file_name, "file_reader")
if file_name.endswith(".png"):
image_reader = tf.image.decode_png(file_reader, channels = 3,name='png_reader')
elif file_name.endswith(".gif"):
image_reader = tf.squeeze(tf.image.decode_gif(file_reader,name='gif_reader'))
elif file_name.endswith(".bmp"):
image_reader = tf.image.decode_bmp(file_reader, name='bmp_reader')
else:
image_reader = tf.image.decode_jpeg(file_reader, channels = 3, name='jpeg_reader')
float_caster = tf.cast(image_reader, tf.float32)
dims_expander = tf.expand_dims(float_caster, 0);
# Read some info from the model metadata, providing default values
input_height = get_model_attribute(model, "input_height", 224)
input_width = get_model_attribute(model, "input_width", 224)
input_mean = get_model_attribute(model, "input_mean", 0)
input_std = get_model_attribute(model, "input_std", 255)
resized = tf.image.resize_bilinear(dims_expander, [input_height, input_width])
normalized = tf.divide(tf.subtract(resized, [input_mean]), [input_std])
sess = tf.Session()
result = sess.run(normalized)
return result
```
## Score_image method that run the model and return a dictionary with the generic and custom model if available. Each entry contains the top 5 candidate answers as value
```
import numpy as np
# classify an image given its url
def score_image(graph, model, url):
# Download the image and build a tensor from its data
t = read_tensor_from_image_file(model, download_image(url))
def do_score_image(graph, output_layer, labels):
# Retrieve the tensors corresponding to the input and output layers
input_tensor = graph.get_tensor_by_name("import/" + input_layer + ":0");
output_tensor = graph.get_tensor_by_name( output_layer + ":0");
with tf.Session(graph=graph) as sess:
# Execute the output, overriding the input tensor with the one corresponding
# to the image in the feed_dict argument
sess.run(tf.global_variables_initializer())
results = sess.run(output_tensor, {input_tensor: t})
results = np.squeeze(results)
# select the top 5 candidates and match them to the labels
top_k = results.argsort()[-5:][::-1]
return [(labels[i].split(":")[1], results[i]) for i in top_k]
results = {}
input_layer = get_model_attribute(model, "input_layer", "input")
labels = load_labels(model)
results["mobilenet"] = do_score_image(graph, "import/" + get_model_attribute(model, "output_layer"), labels)
if "custom_graph" in model and "custom_labels" in model:
with open(model["custom_labels"]) as f:
labels = [line.rstrip() for line in f.readlines() if line != ""]
custom_labels = ["{}:{}".format(i, label) for i,label in zip(range(len(labels)), labels)]
results["custom"] = do_score_image(model["custom_graph"], "final_result", custom_labels)
return results
```
## Tooling for acquiring the training data
```
import pandas
wnid_to_urls = pandas.read_csv('/Users/dtaieb/Downloads/fall11_urls.txt', sep='\t', names=["wnid", "url"],
header=0, error_bad_lines=False, warn_bad_lines=False, encoding="ISO-8859-1")
wnid_to_urls['wnid'] = wnid_to_urls['wnid'].apply(lambda x: x.split("_")[0])
wnid_to_urls = wnid_to_urls.dropna()
wnid_to_words = pandas.read_csv('/Users/dtaieb/Downloads/words.txt', sep='\t', names=["wnid", "description"],
header=0, error_bad_lines=False, warn_bad_lines=False, encoding="ISO-8859-1")
wnid_to_words = wnid_to_words.dropna()
def get_url_for_keywords(keywords):
results = {}
for keyword in keywords:
df = wnid_to_words.loc[wnid_to_words['description'] == keyword]
row_list = df['wnid'].values.tolist()
descriptions = df['description'].values.tolist()
if len(row_list) > 0:
results[descriptions[0]] = wnid_to_urls.loc[wnid_to_urls['wnid'] == row_list[0]]["url"].values.tolist()
return results
import pixiedust
display(wnid_to_urls)
from pixiedust.utils.environment import Environment
import os
root_dir = ensure_dir_exists(os.path.join(Environment.pixiedustHome, "imageRecoApp"))
image_dir = root_dir
def download_image_into_dir(url, path):
file_name = url[url.rfind("/")+1:]
if not file_name.endswith(".jpg"):
return
file_name = os.path.join(path, file_name)
if os.path.exists(file_name):
print("Image already downloaded {}".format(url))
else:
print("Downloading image {}...".format(url), end='')
try:
response = requests.get(url, stream=True)
if response.status_code == 200:
with open(file_name, 'wb') as f:
for chunk in response.iter_content(2048):
f.write(chunk)
else:
print("Error: {}".format(response.status_code))
except Exception as e:
print("Error: {}".format(e))
print("done")
image_dict = get_url_for_keywords(["apple", "orange", "pear", "banana"])
for key in image_dict:
path = ensure_dir_exists(os.path.join(image_dir, key))
count = 0
for url in image_dict[key]:
download_image_into_dir(url, path)
count += 1
if count > 500:
break;
```
## Retrain the model using the downloaded images from the previous section
```
import collections
import os.path
import hashlib
from datetime import datetime
from tensorflow.python.framework import graph_util
from tensorflow.python.platform import gfile
from tensorflow.python.util import compat
MAX_NUM_IMAGES_PER_CLASS = 2 ** 27 - 1 # ~134M
final_tensor_name = "final_result"
tmp_root_dir = ensure_dir_exists(os.path.join(root_dir, "tmp"))
bottleneck_dir = ensure_dir_exists(os.path.join(tmp_root_dir, "bottleneck"))
architecture="mobilenet_0.50_224"
train_batch_size = 100
test_batch_size = 1
validation_batch_size = 100
how_many_training_steps = 500
eval_step_interval = 10
output_graph = os.path.join(tmp_root_dir, "output_graph.pb")
output_labels = os.path.join(tmp_root_dir, "output_labels.txt")
def create_image_lists(image_dir, testing_percentage, validation_percentage):
if not gfile.Exists(image_dir):
tf.logging.error("Image directory '" + image_dir + "' not found.")
return None
result = collections.OrderedDict()
sub_dirs = [os.path.join(image_dir,item) for item in gfile.ListDirectory(image_dir)]
sub_dirs = sorted(item for item in sub_dirs if gfile.IsDirectory(item))
for sub_dir in sub_dirs:
extensions = ['jpg', 'jpeg', 'JPG', 'JPEG']
file_list = []
dir_name = os.path.basename(sub_dir)
if dir_name == image_dir:
continue
tf.logging.info("Looking for images in '" + dir_name + "'")
for extension in extensions:
file_glob = os.path.join(image_dir, dir_name, '*.' + extension)
file_list.extend(gfile.Glob(file_glob))
if not file_list:
tf.logging.warning('No files found')
continue
if len(file_list) < 20:
tf.logging.warning('WARNING: Folder has less than 20 images, which may cause issues.')
elif len(file_list) > MAX_NUM_IMAGES_PER_CLASS:
tf.logging.warning(
'WARNING: Folder {} has more than {} images. Some images will '
'never be selected.'.format(dir_name, MAX_NUM_IMAGES_PER_CLASS))
label_name = re.sub(r'[^a-z0-9]+', ' ', dir_name.lower())
training_images = []
testing_images = []
validation_images = []
for file_name in file_list:
base_name = os.path.basename(file_name)
hash_name = re.sub(r'_nohash_.*$', '', file_name)
hash_name_hashed = hashlib.sha1(compat.as_bytes(hash_name)).hexdigest()
percentage_hash = ((int(hash_name_hashed, 16) % (MAX_NUM_IMAGES_PER_CLASS + 1)) * (100.0 / MAX_NUM_IMAGES_PER_CLASS))
if percentage_hash < validation_percentage:
validation_images.append(base_name)
elif percentage_hash < (testing_percentage + validation_percentage):
testing_images.append(base_name)
else:
training_images.append(base_name)
result[label_name] = {
'dir': dir_name,
'training': training_images,
'testing': testing_images,
'validation': validation_images,
}
return result
def add_jpeg_decoding(model):
input_height = get_model_attribute(model, "input_height")
input_width = get_model_attribute(model, "input_width")
input_depth = get_model_attribute(model, "input_depth")
input_mean = get_model_attribute(model, "input_mean", 0)
input_std = get_model_attribute(model, "input_std", 255)
jpeg_data = tf.placeholder(tf.string, name='DecodeJPGInput')
decoded_image = tf.image.decode_jpeg(jpeg_data, channels=input_depth)
decoded_image_as_float = tf.cast(decoded_image, dtype=tf.float32)
decoded_image_4d = tf.expand_dims(decoded_image_as_float, 0)
resize_shape = tf.stack([input_height, input_width])
resize_shape_as_int = tf.cast(resize_shape, dtype=tf.int32)
resized_image = tf.image.resize_bilinear(decoded_image_4d,
resize_shape_as_int)
offset_image = tf.subtract(resized_image, input_mean)
mul_image = tf.multiply(offset_image, 1.0 / input_std)
return jpeg_data, mul_image
def get_bottleneck_path(image_lists, label_name, index, bottleneck_dir,category, architecture):
return get_image_path(image_lists, label_name, index, bottleneck_dir,category) + '_' + architecture + '.txt'
def get_image_path(image_lists, label_name, index, image_dir, category):
label_lists = image_lists[label_name]
category_list = label_lists[category]
if not category_list:
tf.logging.fatal('Label %s has no images in the category %s.',label_name, category)
mod_index = index % len(category_list)
base_name = category_list[mod_index]
sub_dir = label_lists['dir']
full_path = os.path.join(image_dir, sub_dir, base_name)
return full_path
def get_or_create_bottleneck(sess, image_lists, label_name, index, image_dir,
category, bottleneck_dir, jpeg_data_tensor,
decoded_image_tensor, resized_input_tensor,
bottleneck_tensor, architecture):
label_lists = image_lists[label_name]
sub_dir = label_lists['dir']
sub_dir_path = os.path.join(bottleneck_dir, sub_dir)
ensure_dir_exists(sub_dir_path)
bottleneck_path = get_bottleneck_path(image_lists, label_name, index,bottleneck_dir, category, architecture)
if not os.path.exists(bottleneck_path):
try:
create_bottleneck_file(bottleneck_path, image_lists, label_name, index,
image_dir, category, sess, jpeg_data_tensor,
decoded_image_tensor, resized_input_tensor,
bottleneck_tensor)
except:
return None
with open(bottleneck_path, 'r') as bottleneck_file:
bottleneck_string = bottleneck_file.read()
did_hit_error = False
try:
bottleneck_values = [float(x) for x in bottleneck_string.split(',')]
except ValueError:
tf.logging.warning('Invalid float found, recreating bottleneck')
did_hit_error = True
if did_hit_error:
create_bottleneck_file(bottleneck_path, image_lists, label_name, index,
image_dir, category, sess, jpeg_data_tensor,
decoded_image_tensor, resized_input_tensor,
bottleneck_tensor)
with open(bottleneck_path, 'r') as bottleneck_file:
bottleneck_string = bottleneck_file.read()
bottleneck_values = [float(x) for x in bottleneck_string.split(',')]
return bottleneck_values
def cache_bottlenecks(sess, image_lists, image_dir, bottleneck_dir,
jpeg_data_tensor, decoded_image_tensor,
resized_input_tensor, bottleneck_tensor, architecture):
how_many_bottlenecks = 0
ensure_dir_exists(bottleneck_dir)
for label_name, label_lists in image_lists.items():
for category in ['training', 'testing', 'validation']:
category_list = label_lists[category]
for index, unused_base_name in enumerate(category_list):
get_or_create_bottleneck(
sess, image_lists, label_name, index, image_dir, category,
bottleneck_dir, jpeg_data_tensor, decoded_image_tensor,
resized_input_tensor, bottleneck_tensor, architecture)
how_many_bottlenecks += 1
if how_many_bottlenecks % 100 == 0:
tf.logging.info(str(how_many_bottlenecks) + ' bottleneck files created.')
def create_bottleneck_file(bottleneck_path, image_lists, label_name, index,image_dir, category,
sess, jpeg_data_tensor,decoded_image_tensor, resized_input_tensor,bottleneck_tensor):
tf.logging.info('Creating bottleneck at ' + bottleneck_path)
image_path = get_image_path(image_lists, label_name, index,
image_dir, category)
if not gfile.Exists(image_path):
tf.logging.fatal('File does not exist %s', image_path)
image_data = gfile.FastGFile(image_path, 'rb').read()
try:
bottleneck_values = run_bottleneck_on_image(
sess, image_data, jpeg_data_tensor, decoded_image_tensor,resized_input_tensor, bottleneck_tensor
)
except Exception as e:
raise(RuntimeError('Error during processing file {} ({})'.format(image_path, str(e))))
bottleneck_string = ','.join(str(x) for x in bottleneck_values)
with open(bottleneck_path, 'w') as bottleneck_file:
bottleneck_file.write(bottleneck_string)
def run_bottleneck_on_image(sess, image_data, image_data_tensor,decoded_image_tensor,
resized_input_tensor,bottleneck_tensor):
# First decode the JPEG image, resize it, and rescale the pixel values.
resized_input_values = sess.run(decoded_image_tensor,{image_data_tensor: image_data})
# Then run it through the recognition network.
bottleneck_values = sess.run(bottleneck_tensor,{resized_input_tensor: resized_input_values})
bottleneck_values = np.squeeze(bottleneck_values)
return bottleneck_values
def add_final_training_ops(model, class_count, final_tensor_name, bottleneck_tensor,bottleneck_tensor_size):
with tf.name_scope('input'):
bottleneck_input = tf.placeholder_with_default(
bottleneck_tensor,shape=[None, bottleneck_tensor_size],name='BottleneckInputPlaceholder'
)
ground_truth_input = tf.placeholder(tf.float32,[None, class_count],name='GroundTruthInput')
# Organizing the following ops as `final_training_ops` so they're easier to see in TensorBoard
layer_name = 'final_training_ops'
with tf.name_scope(layer_name):
with tf.name_scope('weights'):
initial_value = tf.truncated_normal([bottleneck_tensor_size, class_count], stddev=0.001)
layer_weights = tf.Variable(initial_value, name='final_weights')
with tf.name_scope('biases'):
layer_biases = tf.Variable(tf.zeros([class_count]), name='final_biases')
with tf.name_scope('Wx_plus_b'):
logits = tf.matmul(bottleneck_input, layer_weights) + layer_biases
tf.summary.histogram('pre_activations', logits)
final_tensor = tf.nn.softmax(logits, name=final_tensor_name)
tf.summary.histogram('activations', final_tensor)
with tf.name_scope('cross_entropy'):
cross_entropy = tf.nn.softmax_cross_entropy_with_logits(labels=ground_truth_input, logits=logits)
with tf.name_scope('total'):
cross_entropy_mean = tf.reduce_mean(cross_entropy)
tf.summary.scalar('cross_entropy', cross_entropy_mean)
with tf.name_scope('train'):
learning_rate = get_model_attribute(model, "learning_rate", 0.01)
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
train_step = optimizer.minimize(cross_entropy_mean)
return (train_step, cross_entropy_mean, bottleneck_input, ground_truth_input,final_tensor)
def add_evaluation_step(result_tensor, ground_truth_tensor):
with tf.name_scope('accuracy'):
with tf.name_scope('correct_prediction'):
prediction = tf.argmax(result_tensor, 1)
correct_prediction = tf.equal(prediction, tf.argmax(ground_truth_tensor, 1))
with tf.name_scope('accuracy'):
evaluation_step = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
tf.summary.scalar('accuracy', evaluation_step)
return evaluation_step, prediction
import random
def get_random_cached_bottlenecks(sess, image_lists, how_many, category,
bottleneck_dir, image_dir, jpeg_data_tensor,
decoded_image_tensor, resized_input_tensor,
bottleneck_tensor, architecture):
class_count = len(image_lists.keys())
bottlenecks = []
ground_truths = []
filenames = []
if how_many >= 0:
# Retrieve a random sample of bottlenecks.
for unused_i in range(how_many):
label_index = random.randrange(class_count)
label_name = list(image_lists.keys())[label_index]
image_index = random.randrange(MAX_NUM_IMAGES_PER_CLASS + 1)
image_name = get_image_path(image_lists, label_name, image_index,image_dir, category)
bottleneck = get_or_create_bottleneck(
sess, image_lists, label_name, image_index, image_dir, category,
bottleneck_dir, jpeg_data_tensor, decoded_image_tensor,
resized_input_tensor, bottleneck_tensor, architecture)
if bottleneck is not None:
ground_truth = np.zeros(class_count, dtype=np.float32)
ground_truth[label_index] = 1.0
bottlenecks.append(bottleneck)
ground_truths.append(ground_truth)
filenames.append(image_name)
else:
# Retrieve all bottlenecks.
for label_index, label_name in enumerate(image_lists.keys()):
for image_index, image_name in enumerate(image_lists[label_name][category]):
image_name = get_image_path(image_lists, label_name, image_index,image_dir, category)
bottleneck = get_or_create_bottleneck(
sess, image_lists, label_name, image_index, image_dir, category,
bottleneck_dir, jpeg_data_tensor, decoded_image_tensor,
resized_input_tensor, bottleneck_tensor, architecture)
if bottleneck:
ground_truth = np.zeros(class_count, dtype=np.float32)
ground_truth[label_index] = 1.0
bottlenecks.append(bottleneck)
ground_truths.append(ground_truth)
filenames.append(image_name)
return bottlenecks, ground_truths, filenames
def save_graph_to_file(sess, graph, graph_file_name):
output_graph_def = graph_util.convert_variables_to_constants(sess, graph.as_graph_def(), [final_tensor_name])
with gfile.FastGFile(graph_file_name, 'wb') as f:
f.write(output_graph_def.SerializeToString())
return
model = models['mobilenet']
graph = load_graph(models['mobilenet'])
bottleneck_tensor = graph.get_tensor_by_name(get_model_attribute(model, "bottleneck_tensor_name") )
resized_image_tensor = graph.get_tensor_by_name(get_model_attribute( model, "resized_input_tensor_name") )
testing_percentage = 10
validation_percentage = 10
image_dir = root_dir
image_lists = create_image_lists(image_dir,testing_percentage,validation_percentage)
class_count = len(image_lists.keys())
if class_count == 0:
print('No valid folders of images found')
if class_count == 1:
print('Only one valid folder of images found - multiple classes are needed for classification.')
with tf.Session(graph=graph) as sess:
# Set up the image decoding sub-graph.
jpeg_data_tensor, decoded_image_tensor = add_jpeg_decoding(model)
cache_bottlenecks(sess, image_lists, image_dir,bottleneck_dir, jpeg_data_tensor,
decoded_image_tensor, resized_image_tensor, bottleneck_tensor, architecture)
# Add the new layer that we'll be training.
bottleneck_tensor_size = get_model_attribute(model, "bottleneck_tensor_size")
(train_step, cross_entropy, bottleneck_input, ground_truth_input,final_tensor) = add_final_training_ops(
model, len(image_lists.keys()), final_tensor_name, bottleneck_tensor,bottleneck_tensor_size)
# Create the operations we need to evaluate the accuracy of our new layer.
evaluation_step, prediction = add_evaluation_step(final_tensor, ground_truth_input)
# Merge all the summaries and write them out to the summaries_dir
merged = tf.summary.merge_all()
summaries_dir = tmp_root_dir + "/retrain_logs"
train_writer = tf.summary.FileWriter(summaries_dir + '/train',sess.graph)
validation_writer = tf.summary.FileWriter(summaries_dir + '/validation')
# Set up all our weights to their initial default values.
init = tf.global_variables_initializer()
sess.run(init)
# Run the training for as many cycles as requested on the command line.
for i in range(how_many_training_steps):
# Get a batch of input bottleneck values, either calculated fresh every
# time with distortions applied, or from the cache stored on disk.
(train_bottlenecks,train_ground_truth, _) = get_random_cached_bottlenecks(
sess, image_lists, train_batch_size, 'training',
bottleneck_dir, image_dir, jpeg_data_tensor,
decoded_image_tensor, resized_image_tensor, bottleneck_tensor,
architecture)
# Feed the bottlenecks and ground truth into the graph, and run a training
# step. Capture training summaries for TensorBoard with the `merged` op.
train_summary, _ = sess.run(
[merged, train_step],
feed_dict={bottleneck_input: train_bottlenecks,ground_truth_input: train_ground_truth})
train_writer.add_summary(train_summary, i)
# Every so often, print out how well the graph is training.
is_last_step = (i + 1 == how_many_training_steps)
if (i % eval_step_interval) == 0 or is_last_step:
train_accuracy, cross_entropy_value = sess.run(
[evaluation_step, cross_entropy],
feed_dict={bottleneck_input: train_bottlenecks,ground_truth_input: train_ground_truth})
tf.logging.info('%s: Step %d: Train accuracy = %.1f%%' %(datetime.now(), i, train_accuracy * 100))
tf.logging.info('%s: Step %d: Cross entropy = %f' % (datetime.now(), i, cross_entropy_value))
validation_bottlenecks, validation_ground_truth, _ = (
get_random_cached_bottlenecks(
sess, image_lists, validation_batch_size, 'validation',
bottleneck_dir, image_dir, jpeg_data_tensor,
decoded_image_tensor, resized_image_tensor, bottleneck_tensor,
architecture
)
)
# Run a validation step and capture training summaries for TensorBoard
# with the `merged` op.
validation_summary, validation_accuracy = sess.run(
[merged, evaluation_step],
feed_dict={bottleneck_input: validation_bottlenecks,ground_truth_input: validation_ground_truth})
validation_writer.add_summary(validation_summary, i)
tf.logging.info('%s: Step %d: Validation accuracy = %.1f%% (N=%d)' %
(datetime.now(), i, validation_accuracy * 100,len(validation_bottlenecks)))
# We've completed all our training, so run a final test evaluation on
# some new images we haven't used before.
test_bottlenecks, test_ground_truth, test_filenames = (
get_random_cached_bottlenecks(
sess, image_lists, test_batch_size, 'testing',
bottleneck_dir, image_dir, jpeg_data_tensor,
decoded_image_tensor, resized_image_tensor, bottleneck_tensor,
architecture)
)
test_accuracy, predictions = sess.run(
[evaluation_step, prediction],
feed_dict={bottleneck_input: test_bottlenecks,ground_truth_input: test_ground_truth}
)
tf.logging.info('Final test accuracy = %.1f%% (N=%d)' % (test_accuracy * 100, len(test_bottlenecks)))
# Write out the trained graph and labels with the weights stored as constants.
model["custom_graph"] = graph
model["custom_labels"] = output_labels
save_graph_to_file(sess, graph, output_graph)
with gfile.FastGFile(output_labels, 'w') as f:
f.write('\n'.join(image_lists.keys()) + '\n')
```
## PixieApp with the following screens:
1. First Tab:
- Ask the user for a url to a web page
- Display the images with top 5 candidate classifications
2. Second Tab:
- Display the model Graph Visualization
3. Third Tab:
- Display the label in a PixieDust table
```
from pixiedust.display.app import *
@PixieApp
class ScoreImageApp():
def setup(self):
self.model = self.parent_pixieapp.model
self.graph = self.parent_pixieapp.graph
@route()
def main_screen(self):
return """
<style>
div.outer-wrapper {
display: table;width:100%;height:300px;
}
div.inner-wrapper {
display: table-cell;vertical-align: middle;height: 100%;width: 100%;
}
</style>
<div class="outer-wrapper">
<div class="inner-wrapper">
<div class="col-sm-3"></div>
<div class="input-group col-sm-6">
<input id="url{{prefix}}" type="text" class="form-control"
value="https://www.flickr.com/search/?text=cats"
placeholder="Enter a url that contains images">
<span class="input-group-btn">
<button class="btn btn-default" type="button" pd_options="image_url=$val(url{{prefix}})">Go</button>
</span>
</div>
</div>
</div>
"""
@route(image_url="*")
@templateArgs
def do_process_url(self, image_url):
image_urls = get_image_urls(image_url)
return """
<div>
{%for url in image_urls%}
<div style="float: left; font-size: 9pt; text-align: center; width: 30%; margin-right: 1%; margin-bottom: 0.5em;">
<img src="{{url}}" style="width: 100%">
<div style="display:inline-block" pd_render_onload pd_options="score_url={{url}}"></div>
</div>
{%endfor%}
<p style="clear: both;">
</div>
"""
@route(score_url="*")
@templateArgs
def do_score_url(self, score_url):
scores_dict = score_image(self.graph, self.model, score_url)
return """
{%for model, results in scores_dict.items()%}
<div style="font-weight:bold">{{model}}</div>
<ul style="text-align:left">
{%for label, confidence in results%}
<li><b>{{label}}</b>: {{confidence}}</li>
{%endfor%}
</ul>
{%endfor%}
"""
```
## Visualize the model graph
```
@PixieApp
class TensorGraphApp():
"""Visualize TensorFlow graph."""
def setup(self):
self.graph = self.parent_pixieapp.graph
self.custom_graph = self.parent_pixieapp.model.get("custom_graph", None)
@route()
@templateArgs
def main_screen(self):
strip_def = self.strip_consts(self.graph.as_graph_def())
code = """
<script>
function load() {{
document.getElementById("{id}").pbtxt = {data};
}}
</script>
<link rel="import" href="https://tensorboard.appspot.com/tf-graph-basic.build.html" onload=load()>
<div style="height:600px">
<tf-graph-basic id="{id}"></tf-graph-basic>
</div>
""".format(data=repr(str(strip_def)), id='graph'+ self.getPrefix()).replace('"', '"')
return """
{%if this.custom_graph%}
<div style="margin-top:10px" pd_refresh>
<pd_script>
self.graph = self.custom_graph if self.graph is not self.custom_graph else self.parent_pixieapp.graph
</pd_script>
<span style="font-weight:bold">Select a model to display:</span>
<select>
<option {%if this.graph!=this.custom_graph%}selected{%endif%} value="main">MobileNet</option>
<option {%if this.graph==this.custom_graph%}selected{%endif%} value="custom">Custom</options>
</select>
{%endif%}
<iframe seamless style="width:1200px;height:620px;border:0" srcdoc="{{code}}"></iframe>
"""
def strip_consts(self, graph_def, max_const_size=32):
"""Strip large constant values from graph_def."""
strip_def = tf.GraphDef()
for n0 in graph_def.node:
n = strip_def.node.add()
n.MergeFrom(n0)
if n.op == 'Const':
tensor = n.attr['value'].tensor
size = len(tensor.tensor_content)
if size > max_const_size:
tensor.tensor_content = "<stripped {} bytes>".format(size).encode("UTF-8")
return strip_def
```
## Searchable table for the model categories
```
@PixieApp
class LabelsApp():
def setup(self):
self.labels = self.parent_pixieapp.load_labels(
self.parent_pixieapp.model, as_json=True
)
self.current_labels = self.labels
self.custom_labels = None
if "custom_labels" in self.parent_pixieapp.model:
with open(self.parent_pixieapp.model["custom_labels"]) as f:
content = f.readlines()
labels = [line.rstrip() for line in content if line != ""]
self.custom_labels = \
[{"index": index, "label" : item} for index, item in zip(range(len(labels)), labels)]
@route()
def main_screen(self):
return """
{%if this.custom_labels%}
<div style="margin-top:10px" pd_refresh>
<pd_script>
self.current_labels = self.custom_labels if self.current_labels is not self.custom_labels else self.labels
</pd_script>
<span style="font-weight:bold">Select a model to display:</span>
<select>
<option {%if this.current_labels!=this.labels%}selected{%endif%} value="main">MobileNet</option>
<option {%if this.current_labels==this.custom_labels%}selected{%endif%} value="custom">Custom</options>
</select>
{%endif%}
<div pd_render_onload pd_entity="current_labels">
<pd_options>
{
"table_noschema": "true",
"handlerId": "tableView",
"rowCount": "10000",
"noChartCache": "true"
}
</pd_options>
</div>
"""
```
## Main ImageRecoApp inheriting from TemplateTabbedApp
```
from pixiedust.apps.template import TemplateTabbedApp
@PixieApp
class ImageRecoApp(TemplateTabbedApp):
def setup(self):
self.apps = [
{"title": "Score", "app_class": "ScoreImageApp"},
{"title": "Model", "app_class": "TensorGraphApp"},
{"title": "Labels", "app_class": "LabelsApp"}
]
self.model = models["mobilenet"]
self.graph = self.load_graph(self.model)
app = ImageRecoApp()
app.run()
```
| github_jupyter |
## Title: CI Biodiversity Hotspots (version 2016.1)
### Description
There are currently 36 recognized biodiversity hotspots. These are Earth’s most biologically rich—yet threatened—terrestrial regions. To qualify as a biodiversity hotspot, an area must meet two strict criteria: <br>
- Contain at least 1,500 species of vascular plants found nowhere else on Earth (known as "endemic" species).
- Have lost at least 70 percent of its primary native vegetation. <br>
Many of the biodiversity hotspots exceed the two criteria. For example, both the Sundaland Hotspot in Southeast Asia and the Tropical Andes Hotspot in South America have about 15,000 endemic plant species. The loss of vegetation in some hotspots has reached a startling 95 percent.
### FLINT
This dataset has been pre-processed/checked and is suitable for use in FLINT. Please adhere to individual dataset licence conditions and citations. Processed data can be accessed here: https://datasets.mojaglobal.workers.dev/
### Format
<b>Extent: </b>Global coverage<br>
<b>Format</b>: polygon geoJSON .json<br>
<b>Cordinate system:</b> EPSG:4326 (WGS84)<br>
<b>Temporal Resolution: </b>2016 <br>
<b>Size:</b> 18MB
### Original source
Original Source: https://zenodo.org/record/3261807#.X-p6mtgzZPa Accessed 29/12/2020 <br>
### Licence
This dataset is available under the Creative Commons Attribution-ShareAlike 4.0 International (CC BY-SA 4.0): https://creativecommons.org/licenses/by-sa/4.0/
### Citation
Michael Hoffman, Kellee Koenig, Gill Bunting, Jennifer Costanza, & Williams, Kristen J. (2016). Biodiversity Hotspots (version 2016.1) (Version 2016.1) [Data set]. Zenodo. http://doi.org/10.5281/zenodo.3261807
### Original format
Global coverage, vector, shapefile<br>
Cordinate system EPSG: 4326 (WGS84)
### Metadata
Version 2016.1. 25 April 2016. Added North American Coastal Plains hotspot (Noss, R.F., Platt, W.J., Sorrie, B.A., Weakley, A.S., Means, D.B., Costanza, J., and Peet, R.K. (2015). How global biodiversity hotspots may go unrecognized: lessons from the North American Coastal Plain. Diversity and Distributions, 21, 236–244.) Hotspot boundary modified to remove overlap with Mesoamerica and Madrean Pine-Oak Woodlands hotspots.
Version 2016. 4 April 2016. Version 2011 with updated Eastern Afromontane hotspot boundary based on improved elevation data (Eastern Afromontane Outcomes profile, BirdLife International, 2016).
Version 2011. Added Forests of Eastern Australia hotspot (Full set of 35 hotspots: Mittermeier, R. A., Turner, W. R., Larsen, F. W., Brooks, T. M., & Gascon, C. (2011). Global biodiversity conservation: The critical role of hotspots. In F. E. Zachos & J. C. Habel (Eds.), Biodiversity Hotspots (pp. 3–22). Berlin Heidelberg: Springer. New hotspot: Williams, K. J., Ford, A., Rosauer, D. F., Silva, N., Mittermeier, R. A., Bruce, C., … Margules, C. (2011). Forests of East Australia: The 35th biodiversity hotspot. In F. E. Zachos & J. C. Habel (Eds.), Biodiversity Hotspots (pp. 295–310). Berlin Heidelberg: Springer.).
Version 2004. Hotspots Revisited (Mittermeier, R. A., Robles Gil, P., Hoffmann, M., Pilgrim, J., Brooks, T., Mittermeier, C. G., … da Fonseca, G. A. B. (2004). Hotspots Revisited: Earth’s Biologically Richest and Most Endangered Ecoregions (p. 390). Mexico City, Mexico: CEMEX.)
Version 2000. Hotspots (Myers, N., Mittermeier, R. A., Mittermeier, C. G., da Fonseca, G. A. B., & Kent, J. (2000). Biodiversity hotspots for conservation priorities. Nature, 403, 853–858.)
### Notes
Has significant overlapping edges and gaps caused by slight topological error over North American coastal plain, Mesoamerica and Polynesia-Micronesia. Code below finds and fixes these by merging into coincident polygons using the longest edge. Some coastlines are slightly offset from actual coastline (+/- 500m).
Processing time is lengthy. Please note, only gaps smaller than 0.5ha will be fixed as gaps larger than this could be valid. Gaps smaller than 0.5ha could also be valid but are removed in this instance for improved drawing speed.
### Processing
Repair geometry, fix topologial error (remove overlaps), convert to geojson, EPSG:4326 (WGS84), remove/disable Z values. View code below - originally processed in ArcGIS but can be converted to open source QGIS or GDAL (or others).
```
# Import arcpy module
import arcpy
import os
# Input variables
in_folder = r"C:\Users\LennyJenny\Documents\ArcGIS\world\UNFCCC\downloads\CIBiodiversityHotspots"
scr_folder = r"C:\Data\scratch.gdb"
out_folder = r"C:\Data\json"
field = "NAME IS NULL OR NAME = ''"
# Environments
workspace = in_folder
arcpy.env.workspace = workspace
arcpy.env.outputCoordinateSystem = arcpy.SpatialReference(4326)
arcpy.env.outputZFlag = "Disabled"
arcpy.env.overwriteOutput = True
scr = arcpy.CreateFileGDB_management(r"C:\Data", "scratch")
arcpy.env.parallelProcessingFactor = "100%"
# List features to process
featureclasses = arcpy.ListFeatureClasses()
print(featureclasses)
# Repair/check topology and make FLINT ready
for fc in featureclasses:
fcname = os.path.join(os.path.splitext(fc)[0])
outjson = os.path.join(out_folder, fcname)
whereclause = "FID_" +fcname + " =-1 AND AREA_GEO <= 5000 Or AREA_GEO IS NULL"
print(fcname + ' processing...')
fLayer = "project_Layer"
arcpy.management.MakeFeatureLayer(fc, fLayer)
projectIntersect = os.path.join(scr_folder, "projectIntersect")
arcpy.analysis.Intersect(fLayer, projectIntersect, "ONLY_FID")
projectSingle = os.path.join(scr_folder, "projectSingle")
arcpy.management.MultipartToSinglepart(projectIntersect, projectSingle)
dissolveSlither = os.path.join(scr_folder, "dissolveSlither")
arcpy.management.Dissolve(projectSingle, dissolveSlither, None, None,"SINGLE_PART")
# Take action if overlaps
if arcpy.management.GetCount(dissolveSlither)[0] == "0":
print('no overlaps detected...checking for gaps...')
projectUnion = os.path.join(scr_folder, "projectUnion")
arcpy.analysis.Union(fLayer,projectUnion, "ALL", None, "NO_GAPS")
arcpy.management.AddGeometryAttributes(projectUnion, "AREA_GEODESIC", None, "SQUARE_METERS")
uniSelect = os.path.join(scr_folder, "uniSelect")
arcpy.analysis.Select(projectUnion, uniSelect, whereclause)
if arcpy.management.GetCount(uniSelect)[0] == "0":
# Progress report no error
print(fcname, 'No gaps and overlaps. Repairing geometry and conversion to json...')
# Process: Repair Geometry (non-simple geometry)
geomRepair = arcpy.management.RepairGeometry(fLayer, "DELETE_NULL", "OGC")[0]
# Process: Features To JSON
arcpy.conversion.FeaturesToJSON(fLayer, outjson, "NOT_FORMATTED", "NO_Z_VALUES", "NO_M_VALUES", "GEOJSON", "WGS84", "USE_FIELD_NAME")
print(outjson, '.geojson complete')
else:
# Take action if gaps
print('gaps detected')
appendGap = arcpy.management.Append(uniSelect, fLayer, "NO_TEST")
selectGap = arcpy.management.SelectLayerByAttribute(fLayer, "NEW_SELECTION", field)
fixedlyr = os.path.join(scr_folder, "fixedlyr")
arcpy.management.Eliminate(selectGap, fixedlyr, "LENGTH")
# Progress report
print(fcname, 'No overlaps but gaps detected and repaired. Repairing geometry and conversion to json...')
# Process: Repair Geometry (non-simple geometry)
geomRepair = arcpy.management.RepairGeometry(fixedlyr, "DELETE_NULL", "OGC")[0]
# Process: Features To JSON
arcpy.conversion.FeaturesToJSON(fixedlyr, outjson, "NOT_FORMATTED", "NO_Z_VALUES", "NO_M_VALUES", "GEOJSON", "WGS84", "USE_FIELD_NAME")
else:
print('Overlaps detected...')
# Fix overlaps
projectErase = os.path.join(scr_folder, "projectErase")
arcpy.analysis.Erase(fLayer, dissolveSlither, projectErase)
arcpy.management.Append(dissolveSlither, projectErase, "NO_TEST")
selectSlither = arcpy.management.SelectLayerByAttribute(projectErase, "NEW_SELECTION", field)
eliminateSlither = os.path.join(scr_folder, "eliminateSlither")
arcpy.management.Eliminate(selectSlither, eliminateSlither, "LENGTH")
print('Overlaps detected and fixed...checking for gaps...')
projectUnion = os.path.join(scr_folder, "projectUnion")
arcpy.analysis.Union(eliminateSlither, projectUnion, "ALL", None, "NO_GAPS")
arcpy.management.AddGeometryAttributes(projectUnion, "AREA_GEODESIC", None, "SQUARE_METERS")
uniSelect = os.path.join(scr_folder, "uniSelect")
arcpy.analysis.Select(projectUnion, uniSelect, "FID_eliminateSlither = -1 AND AREA_GEO <=5000 OR AREA_GEO IS NULL")
if arcpy.management.GetCount(uniSelect)[0] == "0":
# Progress report no error
print(fcname, ' No gaps detected. Repairing geometry and conversion to json...')
# Process: Repair Geometry (non-simple geometry)
geomRepair = arcpy.management.RepairGeometry(eliminateSlither, "DELETE_NULL", "OGC")[0]
# Process: Features To JSON
arcpy.conversion.FeaturesToJSON(eliminateSlither, outjson, "NOT_FORMATTED", "NO_Z_VALUES", "NO_M_VALUES", "GEOJSON", "WGS84", "USE_FIELD_NAME")
print(outjson, '.geojson complete')
else:
# Take action if gaps
appendGap = arcpy.management.Append(uniSelect, eliminateSlither, "NO_TEST")
selectGap = arcpy.management.SelectLayerByAttribute(eliminateSlither, "NEW_SELECTION", field)
fixedlyr = os.path.join(scr_folder, "fixedlyr")
arcpy.management.Eliminate(selectGap, fixedlyr, "LENGTH")
print('gaps detected and repaired')
# Progress report
print(fcname, 'Gaps and overlaps fixed. Repairing geometry and conversion to json...')
# Process: Repair Geometry (non-simple geometry)
geomRepair = arcpy.management.RepairGeometry(fixedlyr, "DELETE_NULL", "OGC")[0]
# Process: Features To JSON
arcpy.conversion.FeaturesToJSON(fixedlyr, outjson, "NOT_FORMATTED", "NO_Z_VALUES", "NO_M_VALUES", "GEOJSON", "WGS84", "USE_FIELD_NAME")
arcpy.AddMessage("All done!")
print('done')
```
| github_jupyter |
# Reproducing NeurIPS2019 - Subspace Attack
As part of the NeurIPS2019 reproducibility challenge (project 2 of EPFL CS-433 2019) we chose to reproduce the paper [__Subspace Attack: Exploiting Promising Subspaces for Query-Efficient Black-box Attacks__](https://openreview.net/pdf?id=S1g-OVBl8r)
The algorithm is specified in:
<img src="img/algo1.png" style="width:600px;"/>
We need to create the following functions:
- Load random reference model
- Loss function calculation
- Prior gradient calculation wrt dropout/layer ratio
- Attack
The pre-trained models are in (https://drive.google.com/file/d/1aXTmN2AyNLdZ8zOeyLzpVbRHZRZD0fW0/view).
The least demanding target model is the __GDAS__.
__Note!__ we start with 0 droupout ratio.
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import random
import torch
from torchvision import datasets, transforms
import numpy as np
from src.import_models import load_model, MODELS_DATA, MODELS_DIRECTORY
from src.plots import imshow
# Load MNIST dataset
preprocess = transforms.Compose([
transforms.ToTensor(),
])
data = datasets.CIFAR10(root='./data', train=True, download=True, transform=preprocess)
data_loader = torch.utils.data.DataLoader(data, batch_size=1, shuffle=True, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
num_classes = len(classes)
# Load reference models
reference_model_names = ['vgg11_bn', 'vgg13_bn', 'vgg16_bn', 'vgg16_bn', 'vgg19_bn', 'AlexNet_bn']
reference_models = list(map(lambda name: load_model(MODELS_DIRECTORY, MODELS_DATA, name, num_classes), reference_model_names))
# Load victim model
victim_model_name = 'gdas'
victim_model = load_model(MODELS_DIRECTORY, MODELS_DATA, victim_model_name, num_classes)
_ = victim_model.eval()
if torch.cuda.is_available():
reference_models = list(map(lambda model: model.to('cuda'), reference_models))
victim_model = victim_model.to('cuda')
for data, target in data_loader:
if torch.cuda.is_available():
data = data.to('cuda')
target = target.to('cuda')
output = victim_model(data)
label_index = output.max(1, keepdim=True)[1].item()
print('Predicted: ' + classes[label_index])
print('Real: ' + classes[target[0]])
print(torch.nn.functional.softmax(output[0].max(), dim=0).item())
imshow(data[0].cpu())
break
def attack(input_batch, true_label, tau, epsilon, delta, eta_g, eta, victim, references, limit=10000 verbose=False, show_images=False):
# Regulators
regmin = input_batch - epsilon
regmax = input_batch + epsilon
# Initialize the adversarial example
x_adv = input_batch.clone()
y = true_label
# setting a example label
criterion = torch.nn.CrossEntropyLoss()
#Initialize the gradient to be estimated
g = torch.zeros_like(input_batch) # check the dimensions
# move the input and model to GPU for speed if available
if torch.cuda.is_available():
x_adv = x_adv.to('cuda')
y = y.to('cuda')
g = g.to('cuda')
regmin = regmin.to('cuda')
regmax = regmax.to('cuda')
# initialize quiries counter
q_counter = 0
if show_images:
with torch.no_grad():
imshow(x_adv[0].cpu())
success = False
while not success and q_counter < limit:
if q_counter % 50 == 0 and verbose:
# imshow(x_adv[0].cpu())
print(f'Iteration number: {str(q_counter / 2)}')
print(f'{str(q_counter)} queries have been made')
# Load random reference model
random_model = random.randint(0, len(reference_models) - 1)
model = references[random_model]
model.eval()
# calculate the prior gradient - L8
x_adv.requires_grad_(True)
model.zero_grad()
output = model(x_adv)
loss = criterion(output, y)
loss.backward()
u = x_adv.grad
# Calculate delta - L11
with torch.no_grad():
# Calculate g_plus and g_minus - L9-10
g_plus = g + tau * u
g_minus = g - tau * u
g_minus = g_minus / g_minus.norm()
g_plus = g_plus / g_plus.norm()
x_plus = x_adv + delta * g_plus
x_minus = x_adv + delta * g_minus
query_minus = victim(x_minus)
query_plus = victim(x_plus)
q_counter += 2
delta_t = ((criterion(query_plus, y) - criterion(query_minus, y)) / (tau * epsilon)) * u
# Update gradient - L12
g = g + eta_g * delta_t
# Update the adverserial example - L13-15
with torch.no_grad():
x_adv = x_adv + eta * torch.sign(g)
x_adv = torch.max(x_adv, regmin)
x_adv = torch.min(x_adv, regmax)
x_adv = torch.clamp(x_adv, 0, 1)
# Check success
label_minus = query_minus.max(1, keepdim=True)[1].item()
label_plus = query_plus.max(1, keepdim=True)[1].item()
if label_minus != true_label.item() or label_plus != true_label.item():
print('Success! after {} queries'.format(q_counter))
print("True: {}".format(true_label.item()))
print("Label minus: {}".format(label_minus))
print("Label plus: {}".format(label_plus))
if show_images:
imshow(x_adv[0].cpu())
success = True
return q_counter if success else -1
# Hyper-parameters
tau = 0.1
epsilon = 8./255
delta = 1.0
eta_g = 100
eta = 0.1
counter = 0
limit = 10
queries = []
for data, target in data_loader:
print(f'\n-------------\n')
print(f'Target image number {counter}')
queries_counter = attack(data, target, tau, epsilon, delta, eta_g, eta, victim_model, reference_models, verbose=True)
counter += 1
queries.append(queries_counter)
if counter == limit:
break
results = np.array(queries)
failed = results == -1
print(f'Mean number of queries: {results[~failed].mean()}')
print(f'Median number of queries: {np.median(results[~failed])}')
print(f'Number of failed queries: {len(results[failed])}')
```
# Attack implementation
28-11-2019: Dan
```
import torch
from torchvision import datasets, transforms
from torchvision.models import vgg16
from PIL import Image
# Hyper-parameters
tau = 0.1
epsilon = 8./255
delta = 0.01
eta_g = 100
eta = 0.1
# loading true example
import urllib
url, filename = ("https://github.com/pytorch/hub/raw/master/dog.jpg", "dog.jpg")
try: urllib.URLopener().retrieve(url, filename)
except: urllib.request.urlretrieve(url, filename)
input_image = Image.open(filename)
preprocess = transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
# transforms.Normalize(mean=[0.485, 0.456, 0.406],
# std=[0.229, 0.224, 0.225]),
])
input_tensor = preprocess(input_image)
input_batch = input_tensor.unsqueeze(0) # create a mini-batch as expected by the model
# Regulators
regmin = input_batch - epsilon
regmax = input_batch + epsilon
# Initialize the adversarial example
x_adv = input_batch
# setting a example label
true_label = 258
y = torch.tensor([true_label]).long()
criterion = torch.nn.CrossEntropyLoss()
#Initialize the gradient to be estimated
g = torch.zeros_like(input_batch) # check the dimensions
# load model
model = torch.hub.load('pytorch/vision:v0.4.2', 'vgg11', pretrained=True)
model.eval()
# load victem model
victim = torch.hub.load('pytorch/vision:v0.4.2', 'alexnet', pretrained=True)
victim.eval()
# move the input and model to GPU for speed if available
if torch.cuda.is_available():
x_adv = x_adv.to('cuda')
model = model.to('cuda')
victim = victim.to('cuda')
y = y.to('cuda')
g = g.to('cuda')
regmin = regmin.to('cuda')
regmax = regmax.to('cuda')
# initialize quiries counter
q_counter = 0
with torch.no_grad():
imshow(x_adv[0].cpu())
success = False
while not success and q_counter < 1000:
print(q_counter)
# Load model
model = torch.hub.load('pytorch/vision:v0.4.2', 'vgg11', pretrained=True)
model.eval()
if torch.cuda.is_available():
model = model.to('cuda')
# calculate the prior gradient - L8
x_adv.requires_grad_(True)
model.zero_grad()
output = model(x_adv)
loss = criterion(output, y)
loss.backward()
u = x_adv.grad
# Calculate g_plus and g_minus - L9-10
g_plus = g + tau*u
g_minus = g - tau*u
g_minus = g_minus/g_minus.norm()
g_plus = g_plus/g_plus.norm()
# Calculate delta - L11
with torch.no_grad():
x_plus = x_adv + delta*g_plus
x_minus = x_adv + delta*g_minus
query_minus = victim(x_minus)
query_plus = victim(x_plus)
q_counter += 2
print(torch.nn.functional.softmax(query_minus[0].max(), dim=0).item())
print(torch.nn.functional.softmax(query_minus[0].max(), dim=0).item())
delta_t = ((criterion(query_plus, y) - criterion(query_minus, y)) / (tau * epsilon)) * u
# Update gradient - L12
g = g + eta_g * delta_t
# Update the adverserial example - L13-15
with torch.no_grad():
x_adv = x_adv + eta * torch.sign(g)
x_adv = torch.max(x_adv, regmin)
x_adv = torch.min(x_adv, regmax)
x_adv = torch.clamp(x_adv, 0, 1)
imshow(x_adv[0].cpu())
# Check success
label_minus = query_minus.max(1, keepdim=True)[1].item()
label_plus = query_plus.max(1, keepdim=True)[1].item()
if label_minus != true_label or label_plus != true_label :
print('Success! after {} queries'.format(q_counter))
print("True: {}".format(true_label))
print("Label minus: {}".format(label_minus))
print("Label plus: {}".format(label_plus))
success = True
```
| github_jupyter |
# Chapter 1 - Jupyter Notebooks
Welcome to the Python for Linguists course! In this course we will be learning Python primarily within Jupyter notebooks.
The core concept behind Jupyter notebooks is that of the [narrative](https://jupyter.readthedocs.io/en/latest/use-cases/narrative-notebook.html). Based on the concept of a scientists notebook which contains data observations coupled with analytic narrative, the Jupyter Notebook offers a way for writing narrative text interspersed with Python code. Those narratives and code are displayed underneath the hood using fancy HTML and Javascript. You never have to worry about this under the hood stuff. But it is good to know what's going on.
The text you're reading now is written in what's known as [Markdown](https://github.com/adam-p/markdown-here/wiki/Markdown-Cheatsheet). The link there contains a cheatsheet about how to write pretty text in markdown. You can do things like add **bolded text** by wrapping text with double asterix: `**your text here**`. Or you can indicate headings with a hashtag: `# A Primary Heading` or `## A Secondary Heading`.
## Cells
Jupyter notebooks are divided into "cells". These are boxes that you can arbitrarily add and sort according to your needs. This text is currently contained in a cell that is marked for "Markdown" text. You can see this by selecting this cell and looking at the label in the selector above:
<img src="../images/markdown.png">
The cell below, alternatively, contains Python code. That cell is marked as "Code"
```
1+1
```
<img src="../images/code_cell.png">
Make sure that when you're trying to run Python code, the selector at the top of the toolbar says "Code"!
## Beware of the Hidden State
Jupyter notebooks are an interface for writing and experimenting with code. But Jupyter has a weak spot. What you see is not always what you get.
Jupyter runs code in the order that you execute it. And it remembers assignments whether or not they are still there. The image below gives an illustration.
The smaller box on the left represents the hidden state. This is code that you have already executed. Look at the next frame. There we have deleted the variable, but it remains loaded in memory. This can lead to really weird situations and problems.
<img src="../images/jupyter_hidden_state.png">
Try it out below. Run the cell and delete the variable. Run the next cell. See how it still works?
```
tense = 'past perfect'
print(tense)
```
### Fighting back
If your code is behaving strangely, a good first step is to restart the kernel. You can do that most easily by clicking the circle arrow button in toolbar.
<img src="../images/restart.png">
This will reset the state and synchronize it with what you expect.
<img src="../images/hidden_state_solved.png" height="50%" width="50%">
Usually the hidden state of notebooks is no problem. We can write and copy back over the variables we assign with no problems. It's only when we remove or rename things that are subsequently called that we run into problems.
## Ending a Session in Jupyter
When you launch Jupyter, it will also open a box with running text. In Mac this is called the Terminal app. In PC it is called the Anaconda prompt.
When you are done with a Jupyter notebook, the proper way to end the session after closing your browser is to press `control + c`. This will ask you if you want to quit. Press `y`. This will terminate the session.
<img src="../images/terminal.png" height="50%" width="50%">
| github_jupyter |
### Install Sqlite3
```shell
sudo apt-get install sqlite3 sqlite3-doc
# Optional
sudo apt install sqlitebrowser
```
### Create Database
```
# sqlite3_createdb.py
import os
import sqlite3
db_filename="todo.db"
db_is_new = not os.path.exists(db_filename)
conn = sqlite3.connect(db_filename)
if db_is_new:
print("Need to create schema")
else:
print("Database exists,assume schema does,too.")
conn.close()
```
### Initial Database With Schema File
```
# sqlite3_create_schema.py
# create schema with todo_schema.sql
import os
import sqlite3
db_filename = "todo.db"
schema_filename = "todo_schema.sql"
db_is_new = not os.path.exists(db_filename)
with sqlite3.connect(db_filename) as conn:
if db_is_new:
print("Create schema")
with open(schema_filename,'rt') as f:
schema = f.read()
conn.executescript(schema)
print("Inserting initial data")
conn.executescript("""
insert into project (name, description, deadline)
values ('pymotw', 'Python Module of the Week',
'2016-11-01');
insert into task (details, status, deadline, project)
values ('write about select', 'done', '2016-04-25',
'pymotw');
insert into task (details, status, deadline, project)
values ('write about random', 'waiting', '2016-08-22',
'pymotw');
insert into task (details, status, deadline, project)
values ('write about sqlite3', 'active', '2017-07-31',
'pymotw');
"""
)
else:
print("Database exists,assume schema does,too.")
# You can check the initial data with command
!sqlite3 todo.db "select * from task"
```
### Retrieving Data From Database
```
# sqlite3_select_tasks.py
import sqlite3
db_filename = 'todo.db'
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
cursor.execute("""
select id, priority, details, status, deadline from task
where project = 'pymotw'
""")
for row in cursor.fetchall():
task_id, priority, details, status, deadline = row
print('{:2d} [{:d}] {:<25} [{:<8}] ({})'.format(
task_id, priority, details, status, deadline))
# sqlite3_select_variations.py
# fetch one or specify number of items from database
import sqlite3
db_filename = 'todo.db'
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
cursor.execute("""
select name, description, deadline from project
where name = 'pymotw'
""")
name, description, deadline = cursor.fetchone()
print('Project details for {} ({})\n due {}'.format(
description, name, deadline))
cursor.execute("""
select id, priority, details, status, deadline from task
where project = 'pymotw' order by deadline
""")
print('\nNext 5 tasks:')
for row in cursor.fetchmany(5):
task_id, priority, details, status, deadline = row
print('{:2d} [{:d}] {:<25} [{:<8}] ({})'.format(
task_id, priority, details, status, deadline))
```
### Query Metadata
> After cursor.execute() has been called,the cursor should set it's description attribute to hold information about data that will be returned by the fetch methods.
>
> The description value is a sequence of tuples containing the colume name,tye,display size,internal size,precision,scale,and a flag says whether null value are accepted.
```
# sqlite3_cursor_description.py
import sqlite3
db_filename = 'todo.db'
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
cursor.execute("""
select * from task where project = 'pymotw'
""")
print('Task table has these columns:')
for colinfo in cursor.description:
print(colinfo)
```
### Row Object
> By default ,the values returned by the fetch methods as "rows" from the database are tuples.The Caller is responsible for knowing the order of the columns in the query and extracting indirvidual values from the tuple.
>
> When the number of values in query grows,it is usually easier to work with an object and access value using their column names.Then the number and order of the tuple contents can change over time as the query is edited,and code depending on ther query results is less likely to break.
>
> Connection objects have a row_factory property that allows the calling code to control the type of object created to represent each row in the query result set. sqlite3 also includes a Row class intended to be used as a row factory. Column values can be accessed through Row instances by using the column index or name.
```
# sqlite3_row_factory.py
import sqlite3
db_filename = 'todo.db'
with sqlite3.connect(db_filename) as conn:
# Change the row factory to use Row
conn.row_factory = sqlite3.Row
cursor = conn.cursor()
cursor.execute("""
select name, description, deadline from project
where name = 'pymotw'
""")
name, description, deadline = cursor.fetchone()
print('Project details for {} ({})\n due {}'.format(
description, name, deadline))
cursor.execute("""
select id, priority, status, deadline, details from task
where project = 'pymotw' order by deadline
""")
print('\nNext 5 tasks:')
# Access the data with row[column name]
for row in cursor.fetchmany(5):
print('{:2d} [{:d}] {:<25} [{:<8}] ({})'.format(
row['id'], row['priority'], row['details'],
row['status'], row['deadline'],
))
```
### Query With Variables
> SQLite supports two forms for queries with placeholders, positional and named.
```
# sqlite3_argument_positional.py
# A question mark (?) denotes a positional argument,
# passed to execute() as a member of a tuple.
import sqlite3
import sys
db_filename = 'todo.db'
# project_name = sys.argv[1]
project_name = "pymotw"
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
query = """
select id, priority, details, status, deadline from task
where project = ?
"""
cursor.execute(query, (project_name,))
for row in cursor.fetchall():
task_id, priority, details, status, deadline = row
print('{:2d} [{:d}] {:<25} [{:<8}] ({})'.format(
task_id, priority, details, status, deadline))
# sqlite3_argument_named.py
import sqlite3
import sys
db_filename = 'todo.db'
# project_name = sys.argv[1]
project_name = "pymotw"
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
query = """
select id, priority, details, status, deadline from task
where project = :project_name
order by deadline, priority
"""
cursor.execute(query, {'project_name': project_name})
for row in cursor.fetchall():
task_id, priority, details, status, deadline = row
print('{:2d} [{:d}] {:<25} [{:<8}] ({})'.format(
task_id, priority, details, status, deadline))
# sqlite3_argument_update.py
import sqlite3
import sys
db_filename = 'todo.db'
# id = int(sys.argv[1])
# status = sys.argv[2]
id = 2
status = "done"
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
query = "update task set status = :status where id = :id"
cursor.execute(query, {'status': status, 'id': id})
# check updated task
!sqlite3 todo.db "select * from task"
```
### Bulk Loading
> To apply the same SQL instruction to a large set of data ,use executemany().
```
# sqlite3_load_csv.py
import csv
import sqlite3
import sys
db_filename = 'todo.db'
# data_filename = sys.argv[1]
data_filename = "data_filename.csv"
SQL = """
insert into task (details, priority, status, deadline, project)
values (:details, :priority, 'active', :deadline, :project)
"""
with open(data_filename, 'rt') as csv_file:
csv_reader = csv.DictReader(csv_file)
with sqlite3.connect(db_filename) as conn:
cursor = conn.cursor()
cursor.executemany(SQL, csv_reader)
!sqlite3 todo.db "select * from task"
```
### Define New Column Types
> SQLite has native support for integer, floating point, and text columns. Data of these types is converted automatically by sqlite3 from Python’s representation to a value that can be stored in the database, and back again, as needed. Integer values are loaded from the database into int or long variables, depending on the size of the value. Text is saved and retrieved as str, unless the text_factory for the Connection has been changed.
>
> Although SQLite only supports a few data types internally, sqlite3 includes facilities for defining custom types to allow a Python application to store any type of data in a column. Conversion for types beyond those supported by default is enabled in the database connection using the detect_types flag. Use PARSE_DECLTYPES if the column was declared using the desired type when the table was defined.
| github_jupyter |
```
import json
import sys
from collections import defaultdict, Counter
import numpy as np
import re
import matplotlib.pyplot as plt
import pandas as pd
from utils.fix_label import fix_general_label_error
EXPERIMENT_DOMAINS = ["none", "hotel", "train", "restaurant", "attraction", "taxi"]
DOMAIN_INDICES = dict()
for domain in EXPERIMENT_DOMAINS:
DOMAIN_INDICES[domain] = len(DOMAIN_INDICES)
def get_slot_information():
ontology = json.load(open("data/multi-woz/MULTIWOZ2.1/ontology.json", 'r'))
ontology_domains = dict([(k, v) for k, v in ontology.items() if k.split("-")[0] in EXPERIMENT_DOMAINS])
SLOTS = [k.replace(" ","").lower() if ("book" not in k) else k.lower() for k in ontology_domains.keys()]
return SLOTS
ALL_SLOTS = get_slot_information()
def fix_none_typo(value):
if value in ("not men", "not", "not mentioned", "", "not mendtioned", "fun", "art"):
return 'none'
else:
return value
# loading data
filename = 'data/train_dials.json'
with open(filename) as fp:
dialogue_data = json.load(fp)
filename = 'data/dev_dials.json'
dialogue_dev_data = dict()
with open(filename) as fp:
for dialogue in json.load(fp):
dialogue['dialogue'].sort(key=lambda x: int(x['turn_idx']))
dialogue_dev_data[dialogue['dialogue_idx']] = dialogue
max_num_turn = max(max(turn['turn_idx'] for turn in dialogue['dialogue']) for dialogue in dialogue_data)
max_num_turn
# calculate distribution of slots at turns
slot_names = set()
slot_turn_pairs = Counter()
for dialogue in dialogue_data:
for turn in dialogue['dialogue']:
turn_idx = turn['turn_idx']
for node_key, node_value in turn['turn_label']:
if fix_none_typo(node_value) == 'none':
continue
node_key = node_key.replace(' ', '-')
slot_names.add(node_key)
slot_turn_pairs[(node_key, turn_idx)] += 1
slot_names = list(slot_names)
slot_names.sort()
slot_turn_pairs
#slot_names_domain = [name for name in slot_names if name.startswith('restaurant-')]
slot_names_domain = slot_names
data = np.array([
[slot_idx, turn_idx, slot_turn_pairs[(slot_names_domain[slot_idx], turn_idx)]] for slot_idx in range(len(slot_names_domain)) for turn_idx in range(max_num_turn+1)
])
x = data[:, 0]
y = data[:, 1]
s = data[:, 2]
plt.figure(figsize=(17, 5))
#plt.scatter(x, y, s)
plt.xticks(1+np.arange(len(slot_names_domain)), slot_names_domain, rotation='vertical')
violindata = [
[turn_idx for turn_idx in range(max_num_turn+1) for _ in range(slot_turn_pairs[(slot_names_domain[slot_idx], turn_idx)])]
for slot_idx in range(len(slot_names_domain))
]
#violindata
plt.gca().violinplot(violindata)
slot_indices = dict((name, idx) for idx, name in enumerate(slot_names))
slot_distribution = [[] for _ in range(len(slot_names))]
for dialogue in dialogue_data:
for turn in dialogue['dialogue']:
turn_idx = turn['turn_idx']
for node_key, node_value in turn['turn_label']:
if fix_none_typo(node_value) == 'none':
continue
node_key = node_key.replace(' ', '-')
slot_distribution[slot_indices[node_key]].append(turn_idx / len(dialogue['dialogue']))
plt.figure(figsize=(17, 5))
#plt.scatter(x, y, s)
plt.xticks(1+np.arange(len(slot_names_domain)), slot_names_domain, rotation='vertical')
plt.gca().violinplot(slot_distribution)
#test_domain = 'train'
test_domain = None
def remove_none_slots(belief):
for slot_tuple in belief:
domain, slot_name, slot_value = slot_tuple.split('-', maxsplit=2)
if slot_value == 'none':
continue
if test_domain is not None and domain != test_domain:
continue
yield slot_tuple
def get_joint_accuracy(turn):
return float(set(remove_none_slots(turn['turn_belief'])) == set(remove_none_slots(turn['pred_bs_ptr'])))
def print_turn(dialogue_id, model_name, up_to=-1):
dialogue = dialogue_dev_data[dialogue_id]
if up_to < 0:
up_to = len(dialogue['dialogue'])
print()
print(dialogue_id + '/' + str(up_to))
for turn_idx in range(up_to + 1):
if turn_idx > 0:
print('S: ' + dialogue['dialogue'][turn_idx]['system_transcript'])
print('U: ' + dialogue['dialogue'][turn_idx]['transcript'])
pred_data_all[model_name][dialogue_id][str(turn_idx)]['turn_belief'].sort()
pred_data_all[model_name][dialogue_id][str(turn_idx)]['pred_bs_ptr'].sort()
print('Ann:', pred_data_all[model_name][dialogue_id][str(turn_idx)]['turn_belief'])
print('Pred:', pred_data_all[model_name][dialogue_id][str(turn_idx)]['pred_bs_ptr'])
#ALL_MODELS = ['taxi2train-pct0', 'taxi2train-pct0-tr5', 'taxi2train-pct0-tr8', 'taxi2train-pct5', 'baseline21']
#ALL_MODELS = ['baseline21', 'aug5', 'aug6']
ALL_MODELS = ['baseline21', 'baseline21-state-gold', 'baseline21-state-epoch3']
pred_data_all = dict()
for modelname in ALL_MODELS:
with open('./model-' + modelname + '/predictions/' + (test_domain if test_domain else 'full') + '/prediction_TRADE_dev.json') as fp:
pred_data = json.load(fp)
pred_data_all[modelname] = pred_data
#test_domain = 'train'
for modelname in ALL_MODELS:
count = 0
accuracy = 0
for dialogue_id, dialogue in dialogue_dev_data.items():
if test_domain is not None and test_domain not in dialogue['domains']:
continue
pred = pred_data_all[modelname][dialogue_id]
for turn in dialogue['dialogue']:
turn_pred = pred[str(turn['turn_idx'])]
count += 1
accuracy += get_joint_accuracy(turn_pred)
print(modelname, count, accuracy, accuracy/count, sep='\t')
modelname = 'baseline21'
max_num_turn = max(len(dlg) for dlg in pred_data_all[modelname].values())
per_turn_count = np.zeros((max_num_turn+1,), dtype=np.int32)
per_turn_accuracy = np.zeros((max_num_turn+1,), dtype=np.int32)
per_turn_recovery = np.zeros((max_num_turn+1,), dtype=np.int32)
for dialogue_id, dialogue in dialogue_dev_data.items():
#if test_domain not in dialogue['domains']:
# continue
pred = pred_data_all[modelname][dialogue_id]
prev_turn_ok = 0
for turn in dialogue['dialogue']:
turn_idx = turn['turn_idx']
turn_pred = pred[str(turn_idx)]
per_turn_count[turn_idx] += 1
turn_ok = get_joint_accuracy(turn_pred)
per_turn_accuracy[turn_idx] += turn_ok
if turn_ok and not prev_turn_ok:
per_turn_recovery[turn_idx] += 1
prev_turn_ok = turn_ok
data = [
[turn_idx, per_turn_count[turn_idx], per_turn_accuracy[turn_idx],
per_turn_accuracy[turn_idx] / per_turn_count[turn_idx],
per_turn_recovery[turn_idx],
per_turn_recovery[turn_idx] / per_turn_count[turn_idx],
pow(0.96, turn_idx+1)]
for turn_idx in range(max_num_turn)
]
pd.DataFrame(data, columns=['turn_idx', 'count', 'accurate count', 'accuracy', 'recovered count', 'recovered accuracy', 'ceiling'])
per_turn_accuracy_all = dict()
for modelname in ALL_MODELS:
pred_data = pred_data_all[modelname]
max_num_turn = max(len(dlg) for dlg in pred_data.values())
per_turn_count = np.zeros((max_num_turn+1,), dtype=np.int32)
per_turn_accuracy = np.zeros((max_num_turn+1,), dtype=np.int32)
per_turn_accuracy_all[modelname] = per_turn_accuracy
for dialogue_id, dialogue in pred_data.items():
for turn_idx, turn in dialogue.items():
turn_idx = int(turn_idx)
per_turn_count[turn_idx] += 1
per_turn_accuracy[turn_idx] += get_joint_accuracy(turn)
data = [
([turn_idx, per_turn_count[turn_idx]] + [
#100 * ((per_turn_accuracy[turn_idx] / per_turn_count[turn_idx]) - (per_turn_accuracy[turn_idx-1] / per_turn_count[turn_idx-1])
# if turn_idx > 0 else
# (per_turn_accuracy[turn_idx] / per_turn_count[turn_idx]))
100 * ((per_turn_accuracy[turn_idx] / per_turn_count[turn_idx]))
for per_turn_accuracy in per_turn_accuracy_all.values()])
for turn_idx in range(max_num_turn)
]
pd.DataFrame(data, columns=['turn_idx', 'count']+ list(per_turn_accuracy_all.keys()))
per_num_slot_accuracy_all = dict()
max_num_slots = 0
for dialogue in pred_data_all['baseline21'].values():
for turn in dialogue.values():
max_num_slots = max(max_num_slots, len(set(remove_none_slots(turn['turn_belief']))))
max_num_slots += 1
for modelname in ALL_MODELS:
pred_data = pred_data_all[modelname]
per_num_slot_count = np.zeros((max_num_turn+1,), dtype=np.int32)
per_num_slot_accuracy = np.zeros((max_num_turn+1,), dtype=np.int32)
per_num_slot_accuracy_all[modelname] = per_num_slot_accuracy
for dialogue_id, dialogue in pred_data.items():
for turn_idx, turn in dialogue.items():
num_slot = len(set(remove_none_slots(turn['turn_belief'])))
per_num_slot_count[num_slot] += 1
per_num_slot_accuracy[num_slot] += get_joint_accuracy(turn)
data = [
([num_slot, per_num_slot_count[num_slot]] + [
#100 * ((per_turn_accuracy[turn_idx] / per_turn_count[turn_idx]) - (per_turn_accuracy[turn_idx-1] / per_turn_count[turn_idx-1])
# if turn_idx > 0 else
# (per_turn_accuracy[turn_idx] / per_turn_count[turn_idx]))
100 * ((per_num_slot_accuracy[num_slot] / per_num_slot_count[num_slot]))
for per_num_slot_accuracy in per_num_slot_accuracy_all.values()])
for num_slot in range(max_num_slots)
]
pd.DataFrame(data, columns=['# slots', 'count']+ list(per_num_slot_accuracy_all.keys()))
count = 0
accuracy_both = 0
accuracy_neither = 0
accuracy_only_baseline = 0
accuracy_only_improved = 0
max_num_slots = 0
for dialogue in pred_data_all['baseline21'].values():
for turn in dialogue.values():
max_num_slots = max(max_num_slots, len(set(remove_none_slots(turn['turn_belief']))))
max_num_slots += 1
max_num_turn = max(len(dlg) for dlg in pred_data_all['baseline21'].values())
per_num_slot_stats = dict()
for num_slot in range(max_num_slots):
for turn_idx in range(max_num_turn):
per_num_slot_stats[(num_slot, turn_idx)] = {
'count': 0,
'accuracy_both': 0,
'accuracy_neither': 0,
'accuracy_only_baseline': 0,
'accuracy_only_improved': 0,
}
#baseline_model = 'taxi2train-pct0-tr5'
#improved_model = 'taxi2train-pct0-tr7'
baseline_model = 'baseline21'
improved_model = 'aug6'
for dialogue_id, dialogue in dialogue_dev_data.items():
if test_domain is not None and test_domain not in dialogue['domains']:
continue
for turn in dialogue['dialogue']:
turn_idx = str(turn['turn_idx'])
baseline_ok = get_joint_accuracy(pred_data_all[baseline_model][dialogue_id][turn_idx])
improved_ok = get_joint_accuracy(pred_data_all[improved_model][dialogue_id][turn_idx])
num_slot = len(set(remove_none_slots(pred_data_all[baseline_model][dialogue_id][turn_idx]['turn_belief'])))
count += 1
stat_key = (num_slot, turn['turn_idx'])
per_num_slot_stats[stat_key]['count'] += 1
if baseline_ok and improved_ok:
accuracy_both += 1
per_num_slot_stats[stat_key]['accuracy_both'] += 1
elif baseline_ok:
accuracy_only_baseline += 1
per_num_slot_stats[stat_key]['accuracy_only_baseline'] += 1
elif improved_ok:
accuracy_only_improved += 1
per_num_slot_stats[stat_key]['accuracy_only_improved'] += 1
else:
accuracy_neither += 1
per_num_slot_stats[stat_key]['accuracy_neither'] += 1
print('# overall')
print('total =', count)
print('both =', accuracy_both, '(%.1f%%)' % (100* accuracy_both/count,))
print('lost =', accuracy_only_baseline, '(%.1f%%)' % (100* accuracy_only_baseline/count,))
print('gained =', accuracy_only_improved, '(%.1f%%)' % (100* accuracy_only_improved/count,))
print('neither =', accuracy_neither, '(%.1f%%)' % (100* accuracy_neither/count,))
for num_slot in range(max_num_slots):
for turn_idx in range(max_num_turn):
stat_key = (num_slot, turn_idx)
count = per_num_slot_stats[stat_key]['count']
if count == 0:
continue
print()
print('# %d slots, turn %d' % (num_slot, turn_idx))
print('total =', count)
print('both =', per_num_slot_stats[stat_key]['accuracy_both'],
'(%.1f%%)' % (100* per_num_slot_stats[stat_key]['accuracy_both']/count,))
print('lost =', per_num_slot_stats[stat_key]['accuracy_only_baseline'],
'(%.1f%%)' % (100* per_num_slot_stats[stat_key]['accuracy_only_baseline']/count,))
print('gained =', per_num_slot_stats[stat_key]['accuracy_only_improved'],
'(%.1f%%)' % (100* per_num_slot_stats[stat_key]['accuracy_only_improved']/count,))
print('neither =', per_num_slot_stats[stat_key]['accuracy_neither'],
'(%.1f%%)' % (100* per_num_slot_stats[stat_key]['accuracy_neither']/count,))
#baseline_model = 'taxi2train-pct0-tr5'
#improved_model = 'taxi2train-pct0-tr7'
baseline_model = 'baseline21'
improved_model = 'aug6'
count = 0
for dialogue_id, dialogue in dialogue_dev_data.items():
if test_domain is not None and test_domain not in dialogue['domains']:
continue
for turn in dialogue['dialogue']:
turn_idx = str(turn['turn_idx'])
num_slot = len(set(remove_none_slots(pred_data_all[baseline_model][dialogue_id][turn_idx]['turn_belief'])))
if num_slot > 2:
continue
baseline_ok = get_joint_accuracy(pred_data_all[baseline_model][dialogue_id][turn_idx])
improved_ok = get_joint_accuracy(pred_data_all[improved_model][dialogue_id][turn_idx])
if baseline_ok and not improved_ok:
count += 1
print_turn(dialogue_id, improved_model, int(turn_idx))
print(count)
baseline_model = '21notrainfix'
improved_model = 'tr1-taxi2trainfix'
test_domain = 'train'
def has_domain(belief, domain):
return any(slot.split('-', maxsplit=1)[0] == domain for slot in belief)
count = 0
for dialogue_id, dialogue in dialogue_dev_data.items():
if test_domain not in dialogue['domains']:
continue
for turn in dialogue['dialogue']:
turn_idx = turn['turn_idx']
improved_pred = pred_data_all[improved_model][dialogue_id][str(turn_idx)]
if has_domain(improved_pred['pred_bs_ptr'], 'train') and has_domain(improved_pred['pred_bs_ptr'], 'taxi'):
print_turn(dialogue_id, improved_model, turn_idx)
count += 1
if count >= 100:
break
pred_data = pred_data_all['baseline21']
with open('turn1-errors.txt', 'w') as fp:
count = 0
for dialogue_id, dialogue in pred_data.items():
turn = dialogue['1']
num_considered_turns = 2
if not get_joint_accuracy(turn):
turn_objs = dialogue_dev_data[dialogue_id]['dialogue'][ : num_considered_turns]
print(dialogue_id, file=fp)
for turn_obj in turn_objs:
if turn_obj['system_transcript']:
print('S:', turn_obj['system_transcript'], file=fp)
print('U:', turn_obj['transcript'], file=fp)
print('Ann:', json.dumps(list(sorted(turn['turn_belief']))), file=fp)
print('Pred:', json.dumps(list(sorted(turn['pred_bs_ptr']))), file=fp)
print(file=fp)
count += 1
#if count >= :
# break
def get_turn_domains(turn):
for slot_tuple in turn['turn_belief']:
domain, slot_name, slot_value = slot_tuple.split('-')
yield domain
normal_count = 0
normal_accuracy = 0
trans_count = 0
trans_accuracy = 0
delayed_trans_count = 0
delayed_trans_accuracy = 0
end_normal_count = 0
end_normal_accuracy = 0
end_trans_count = 0
end_trans_accuracy = 0
transition_count = 0
for dialogue_id, dialogue in pred_data.items():
prev_turn_domains = set()
delayed_trans_distance = 0
had_transition = False
for turn_idx, turn in dialogue.items():
turn_idx = int(turn_idx)
#turn_id = dialogue_id + '/' + str(turn_idx)
our_turn_domains = set(get_turn_domains(turn))
our_turn_accuracy = get_joint_accuracy(turn)
normal_count += 1
normal_accuracy += our_turn_accuracy
if len(prev_turn_domains) > 0:
is_transition = False
for dom in our_turn_domains:
if dom not in prev_turn_domains:
is_transition = True
break
if is_transition:
trans_count += 1
trans_accuracy += our_turn_accuracy
delayed_trans_distance = 100
had_transition = True
if delayed_trans_distance > 0:
delayed_trans_count += 1
delayed_trans_accuracy += our_turn_accuracy
delayed_trans_distance -= 1
if had_transition:
transition_count += 1
prev_turn_domains = our_turn_domains
if turn_idx == len(dialogue)-1:
end_normal_count += 1
end_normal_accuracy += our_turn_accuracy
if had_transition:
end_trans_count += 1
end_trans_accuracy += our_turn_accuracy
assert trans_count > 0
(normal_accuracy / normal_count, trans_accuracy / trans_count, delayed_trans_accuracy / delayed_trans_count,
end_normal_accuracy / end_normal_count, end_trans_accuracy / end_trans_count, transition_count / normal_count)
def get_slot_names(belief):
for slot_tuple in belief:
domain, slot_name, slot_value = slot_tuple.split('-')
if slot_value == 'none':
continue
yield domain + '-' + slot_name
def get_name_accuracy(turn):
return float(set(get_slot_names(turn['turn_belief'])) == set(get_slot_names(turn['pred_bs_ptr'])))
count = 0
joint_accuracy = 0
name_accuracy = 0
for dialogue_id, dialogue in pred_data.items():
for turn_idx, turn in dialogue.items():
#turn_idx = int(turn_idx)
count += 1
joint_accuracy += get_joint_accuracy(turn)
name_accuracy += get_name_accuracy(turn)
(joint_accuracy / count, name_accuracy / count)
count = 0
insertions = 0
name_errors = 0
max_num_turns = max(len(dialogue) for dialogue in pred_data.values())
count_per_turn = np.zeros((max_num_turns+1,), dtype=np.int32)
name_accuracy_per_turn = np.zeros((max_num_turns+1,), dtype=np.int32)
insertions_per_turn = np.zeros((max_num_turns+1,), dtype=np.int32)
for dialogue_id, dialogue in pred_data.items():
for turn_idx, turn in dialogue.items():
turn_idx = int(turn_idx)
count += 1
count_per_turn[turn_idx] += 1
turn_belief = set(get_slot_names(turn['turn_belief']))
pred_bs_ptr = set(get_slot_names(turn['pred_bs_ptr']))
our_name_accuracy = int(turn_belief == pred_bs_ptr)
name_accuracy_per_turn[turn_idx] += our_name_accuracy
name_errors += 1 - our_name_accuracy
is_insertion = False
for slot_name in pred_bs_ptr:
if slot_name not in turn_belief:
is_insertion = True
break
if is_insertion:
insertions += 1
insertions_per_turn[turn_idx] += 1
(insertions, name_errors, insertions/name_errors, count, insertions / count)
x = np.arange(max_num_turns)
y1 = []
y2 = []
for turn_idx in range(max_num_turns):
print(turn_idx, count_per_turn[turn_idx], insertions_per_turn[turn_idx],
count_per_turn[turn_idx] - name_accuracy_per_turn[turn_idx] - insertions_per_turn[turn_idx],
sep='\t')
insertion_per_turn = insertions_per_turn[turn_idx] / count_per_turn[turn_idx]
name_error_per_turn = 1 - (name_accuracy_per_turn[turn_idx] / count_per_turn[turn_idx])
missing_per_turn = name_error_per_turn - insertion_per_turn
y1.append(insertions_per_turn[turn_idx] / count)
y2.append(missing_per_turn * count_per_turn[turn_idx] / count)
plt.plot(x, y1, label='extra')
plt.plot(x, y2, label='missing')
plt.legend()
```
| github_jupyter |
```
import gc
import numpy as np
import pandas as pd
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
from sklearn.preprocessing import LabelEncoder
ls ../input/
dtypes = {
'MachineIdentifier': 'category',
'ProductName': 'category',
'EngineVersion': 'category',
'AppVersion': 'category',
'AvSigVersion': 'category',
'IsBeta': 'int8',
'RtpStateBitfield': 'float16',
'IsSxsPassiveMode': 'int8',
'DefaultBrowsersIdentifier': 'float16',
'AVProductStatesIdentifier': 'float32',
'AVProductsInstalled': 'float16',
'AVProductsEnabled': 'float16',
'HasTpm': 'int8',
'CountryIdentifier': 'int16',
'CityIdentifier': 'float32',
'OrganizationIdentifier': 'float16',
'GeoNameIdentifier': 'float16',
'LocaleEnglishNameIdentifier': 'int8',
'Platform': 'category',
'Processor': 'category',
'OsVer': 'category',
'OsBuild': 'int16',
'OsSuite': 'int16',
'OsPlatformSubRelease': 'category',
'OsBuildLab': 'category',
'SkuEdition': 'category',
'IsProtected': 'float16',
'AutoSampleOptIn': 'int8',
'PuaMode': 'category',
'SMode': 'float16',
'IeVerIdentifier': 'float16',
'SmartScreen': 'category',
'Firewall': 'float16',
'UacLuaenable': 'float32',
'Census_MDC2FormFactor': 'category',
'Census_DeviceFamily': 'category',
'Census_OEMNameIdentifier': 'float16',
'Census_OEMModelIdentifier': 'float32',
'Census_ProcessorCoreCount': 'float16',
'Census_ProcessorManufacturerIdentifier': 'float16',
'Census_ProcessorModelIdentifier': 'float16',
'Census_ProcessorClass': 'category',
'Census_PrimaryDiskTotalCapacity': 'float32',
'Census_PrimaryDiskTypeName': 'category',
'Census_SystemVolumeTotalCapacity': 'float32',
'Census_HasOpticalDiskDrive': 'int8',
'Census_TotalPhysicalRAM': 'float32',
'Census_ChassisTypeName': 'category',
'Census_InternalPrimaryDiagonalDisplaySizeInInches': 'float16',
'Census_InternalPrimaryDisplayResolutionHorizontal': 'float16',
'Census_InternalPrimaryDisplayResolutionVertical': 'float16',
'Census_PowerPlatformRoleName': 'category',
'Census_InternalBatteryType': 'category',
'Census_InternalBatteryNumberOfCharges': 'float32',
'Census_OSVersion': 'category',
'Census_OSArchitecture': 'category',
'Census_OSBranch': 'category',
'Census_OSBuildNumber': 'int16',
'Census_OSBuildRevision': 'int32',
'Census_OSEdition': 'category',
'Census_OSSkuName': 'category',
'Census_OSInstallTypeName': 'category',
'Census_OSInstallLanguageIdentifier': 'float16',
'Census_OSUILocaleIdentifier': 'int16',
'Census_OSWUAutoUpdateOptionsName': 'category',
'Census_IsPortableOperatingSystem': 'int8',
'Census_GenuineStateName': 'category',
'Census_ActivationChannel': 'category',
'Census_IsFlightingInternal': 'float16',
'Census_IsFlightsDisabled': 'float16',
'Census_FlightRing': 'category',
'Census_ThresholdOptIn': 'float16',
'Census_FirmwareManufacturerIdentifier': 'float16',
'Census_FirmwareVersionIdentifier': 'float32',
'Census_IsSecureBootEnabled': 'int8',
'Census_IsWIMBootEnabled': 'float16',
'Census_IsVirtualDevice': 'float16',
'Census_IsTouchEnabled': 'int8',
'Census_IsPenCapable': 'int8',
'Census_IsAlwaysOnAlwaysConnectedCapable': 'float16',
'Wdft_IsGamer': 'float16',
'Wdft_RegionIdentifier': 'float16'
}
test = pd.read_csv('../input/test.csv',nrows =100,dtype=dtypes)#Remove nrows stuff
dtypes['HasDetections'] = 'int8'
train = pd.read_csv('../input/train.csv',nrows =100, dtype=dtypes)#Remove nrows stuff
train.HasDetections.mean()
train.head()
traintargets = train['HasDetections'].values
del train['HasDetections']
gc.collect()
del train['Census_FirmwareVersionIdentifier']
del test['Census_FirmwareVersionIdentifier']
del train['Census_OEMModelIdentifier']
del test['Census_OEMModelIdentifier']
del train['CityIdentifier']
del test['CityIdentifier']
gc.collect()
true_numerical_columns = [
'Census_ProcessorCoreCount',
'Census_PrimaryDiskTotalCapacity',
'Census_SystemVolumeTotalCapacity',
'Census_TotalPhysicalRAM',
'Census_InternalPrimaryDiagonalDisplaySizeInInches',
'Census_InternalPrimaryDisplayResolutionHorizontal',
'Census_InternalPrimaryDisplayResolutionVertical',
'Census_InternalBatteryNumberOfCharges'
]
for c in true_numerical_columns:
print(c)
if(len(train[c].round(0).unique())>100):
train.loc[~train[c].isnull(),c] = np.log1p(train.loc[~train[c].isnull(),c]).round(0)
test.loc[~test[c].isnull(),c] = np.log1p(test.loc[~test[c].isnull(),c]).round(0)
else:
train.loc[~train[c].isnull(),c] = (train.loc[~train[c].isnull(),c]).round(0)
test.loc[~test[c].isnull(),c] = (test.loc[~test[c].isnull(),c]).round(0)
for c in train.columns[1:]:
print(c)
le = LabelEncoder()
le.fit(list(train.loc[~train[c].isnull(),c].values)+list(test.loc[~test[c].isnull(),c].values))
train['le_'+c] = 0
test['le_'+c] = 0
train.loc[~train[c].isnull(),'le_'+c] = le.transform(train.loc[~train[c].isnull(),c].values)+1
test.loc[~test[c].isnull(),'le_'+c] = le.transform(test.loc[~test[c].isnull(),c].values)+1
train['le_'+c] = train['le_'+c].fillna(0)
test['le_'+c] = test['le_'+c].fillna(0)
del train[c]
del test[c]
gc.collect()
train['HasDetections'] = traintargets
train.shape
test.shape
train.head()
features = train.columns[1:-1]
categories = train.columns[1:-1]
numerics = []
currentcode = len(numerics)
catdict = {}
catcodes = {}
for x in numerics:
catdict[x] = 0
for x in categories:
catdict[x] = 1
noofrows = train.shape[0]
noofcolumns = len(features)
with open("alltrainffm.txt", "w") as text_file:
for n, r in enumerate(range(noofrows)):
if((n%100000)==0):
print('Row',n)
datastring = ""
datarow = train.iloc[r].to_dict()
datastring += str(int(datarow['HasDetections']))
for i, x in enumerate(catdict.keys()):
if(catdict[x]==0):
datastring = datastring + " "+str(i)+":"+ str(i)+":"+ str(datarow[x])
else:
if(x not in catcodes):
catcodes[x] = {}
currentcode +=1
catcodes[x][datarow[x]] = currentcode
elif(datarow[x] not in catcodes[x]):
currentcode +=1
catcodes[x][datarow[x]] = currentcode
code = catcodes[x][datarow[x]]
datastring = datastring + " "+str(i)+":"+ str(int(code))+":1"
datastring += '\n'
text_file.write(datastring)
noofrows = test.shape[0]
noofcolumns = len(features)
with open("alltestffm.txt", "w") as text_file:
for n, r in enumerate(range(noofrows)):
if((n%100000)==0):
print('Row',n)
datastring = ""
datarow = test.iloc[r].to_dict()
datastring += str(0)
for i, x in enumerate(catdict.keys()):
if(catdict[x]==0):
datastring = datastring + " "+str(i)+":"+ str(i)+":"+ str(datarow[x])
else:
if(x not in catcodes):
catcodes[x] = {}
currentcode +=1
catcodes[x][datarow[x]] = currentcode
elif(datarow[x] not in catcodes[x]):
currentcode +=1
catcodes[x][datarow[x]] = currentcode
code = catcodes[x][datarow[x]]
datastring = datastring + " "+str(i)+":"+ str(int(code))+":1"
datastring += '\n'
text_file.write(datastring)
```
| github_jupyter |
# Detecting seasonality <img align="right" src="../Supplementary_data/dea_logo.jpg">
* [**Sign up to the DEA Sandbox**](https://docs.dea.ga.gov.au/setup/sandbox.html) to run this notebook interactively from a browser
* **Compatibility:** Notebook currently compatible with both the `NCI` and `DEA Sandbox` environments
* **Products used:**
[DEA Waterbodies](https://cmi.ga.gov.au/data-products/dea/456/waterboards)
## Description
We often want to analyse a time series to detect long-term trends or events. Time series that span multiple months will have seasonal effects: for example, northern Australia is much wetter in summer due to monsoons. This seasonality may impact our ability to detect the trends or events we are interested in. This notebook provides a few different ways to detect seasonality and check whether deseasonalised data have been correctly deseasonalised. We will look at a seasonal waterbody time series as an example.
***
## Getting started
To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
### Load packages
Load key Python packages and supporting functions for the analysis.
```
%matplotlib inline
import calendar
import matplotlib.pyplot as plt
import matplotlib.cm
import numpy as np
import pandas as pd
import scipy.stats
import statsmodels.tsa.seasonal as sm_seasonal
import statsmodels.tsa.stattools as tsa_stats
import shapely
import sys
sys.path.insert(1, '../Tools/')
from dea_tools.waterbodies import get_waterbody, get_time_series
```
### Analysis parameters
Choose a waterbody to analyse:
```
geohash = "r1tw92u7y" # Lake Tyrrell
```
## Load the waterbody time series
```
ts = get_time_series(geohash)
ts.pc_wet.plot()
```
Then resample the time series to weekly and interpolate. Having a consistent gap between observations makes analysis of time series much easier. In particular, this notebook won't work without an evenly spaced time series.
Here, we interpolate with `pandas` since our time series is in a `pandas.DataFrame`; `xarray` has a similar interpolate method that can interpolate over `xarray.DataArray`.
```
ts = ts.resample("1W").mean().interpolate(method="time").pc_wet
assert not ts.isnull().any()
```
Deseasonalise it for comparison.
```
decomposition = sm_seasonal.seasonal_decompose(ts + 1, model="multiplicative")
ts_deseasonal = decomposition.trend + decomposition.resid - 1
ts_deseasonal = ts_deseasonal[pd.notnull(ts_deseasonal)]
```
## Autocorrelation
The autocorrelation function (ACF) shows how correlated lagged data are. Seasonal data are highly correlated with a lag of one year. Let's compute the ACF for both our waterbody time series and the deseasonalised waterbody time series.
```
acf = tsa_stats.acf(ts, nlags=52 * 3, fft=True)
acf_deseasonal = tsa_stats.acf(ts_deseasonal, nlags=52 * 3, fft=True)
plt.plot(acf, label="seasonal")
plt.plot(acf_deseasonal, label="deseasonalised")
plt.xlabel("Lag (weeks)")
plt.ylabel("ACF")
for i in range(1, 3):
plt.axvline(52 * i, c="grey", linestyle="--")
plt.legend();
```
The seasonal peaks are clearly visible at the 52 and 104 week marks for the seasonal data, while no such peaks can be seen for the deseasonalised data.
## Angular visualisation
A date can be thought of as an angle (the angle of the Earth's position around the sun). In this way we can project the time series into polar coordinates. Non-seasonal data will circle around the origin and be a perfect circle on average; seasonal data will be offset from the origin. Long-term trends may show as spirals.
```
time_angle = 2 * np.pi * ts.index.dayofyear / 365.25
ax = plt.subplot(projection="polar")
ax.plot(time_angle, ts)
ax.set_xticks(np.linspace(0, 2 * np.pi, 12, endpoint=False))
ax.set_xticklabels(list(calendar.month_abbr)[1:]);
```
We can also bin the data by angle to make the circle easier to see.
```
n_bins = 52
binned_mean = scipy.stats.binned_statistic(time_angle, ts, bins=n_bins)
binned_stdev = scipy.stats.binned_statistic(
time_angle, ts, bins=n_bins, statistic="std"
)
ax = plt.subplot(projection="polar")
mean = np.resize(binned_mean.statistic, n_bins + 1)
stdev = np.resize(binned_stdev.statistic, n_bins + 1)
# np.resize is different to ndarray.resize!
ax.plot(binned_mean.bin_edges, mean)
ax.fill_between(
binned_mean.bin_edges, np.clip(mean - stdev, 0, None), mean + stdev, alpha=0.2
)
# Get the centre of the circle so we can plot that too:
proj = ax.transData.transform(np.stack([binned_mean.bin_edges, mean]).T)
polygon = shapely.geometry.Polygon(proj)
reproj = ax.transData.inverted().transform((polygon.centroid.x, polygon.centroid.y))
plt.scatter(*reproj, c="C0")
ax.set_xticks(np.linspace(0, 2 * np.pi, 12, endpoint=False))
ax.set_xticklabels(list(calendar.month_abbr)[1:]);
```
This waterbody is wetter from May to September and drier from November to April. The centre of the circle is clearly offset. Let's compare to the deseasonalised:
```
time_angle_deseasonal = 2 * np.pi * ts_deseasonal.index.dayofyear / 365.25
binned_mean_deseasonal = scipy.stats.binned_statistic(
time_angle_deseasonal, ts_deseasonal, bins=n_bins
)
binned_stdev_deseasonal = scipy.stats.binned_statistic(
time_angle_deseasonal, ts_deseasonal, bins=n_bins, statistic="std"
)
ax = plt.subplot(projection="polar")
ax.plot(binned_mean.bin_edges, mean, label="seasonal")
ax.fill_between(
binned_mean.bin_edges, np.clip(mean - stdev, 0, None), mean + stdev, alpha=0.2
)
plt.scatter(*reproj, c="C0")
mean_deseasonal = np.resize(binned_mean_deseasonal.statistic, n_bins + 1)
stdev_deseasonal = np.resize(binned_stdev_deseasonal.statistic, n_bins + 1)
# np.resize is different to ndarray.resize!
ax.plot(binned_mean_deseasonal.bin_edges, mean_deseasonal, label="deseasonalised")
ax.fill_between(
binned_mean_deseasonal.bin_edges,
np.clip(mean_deseasonal - stdev_deseasonal, 0, None),
mean_deseasonal + stdev_deseasonal,
alpha=0.2,
)
proj_deseasonal = ax.transData.transform(
np.stack([binned_mean_deseasonal.bin_edges, mean_deseasonal]).T
)
polygon_deseasonal = shapely.geometry.Polygon(proj_deseasonal)
reproj_deseasonal = ax.transData.inverted().transform(
(polygon_deseasonal.centroid.x, polygon_deseasonal.centroid.y)
)
plt.scatter(*reproj_deseasonal, c="C1")
ax.set_xticks(np.linspace(0, 2 * np.pi, 12, endpoint=False))
ax.set_xticklabels(list(calendar.month_abbr)[1:])
ax.legend(bbox_to_anchor=(0, 1));
```
The deaseasonalised data are much more circular and the centre is very close to the origin.
We can convert this into a numerical measure to help determine how seasonal the data are (which would be useful for analysis en masse). One way to do this is to compute the Polsby-Popper score, which is the ratio of the area to the squared perimeter, multiplied by $4\pi$. This is a measure of compactness and a circle is a maximally compact shape.
```
pp = 4 * np.pi * polygon.area / polygon.exterior.length ** 2
pp_deseasonal = (
4 * np.pi * polygon_deseasonal.area / polygon_deseasonal.exterior.length ** 2
)
print("Seasonal Polsby-Popper:", round(pp, 2))
print("Deseasonalised Polsby-Popper:", round(pp_deseasonal, 2))
```
The closer we are to 1, the more circular the data.
Circularity alone doesn't tell us that we have non-seasonal data. The circle also needs to be centred. We can measure the distance from the origin:
```
print("Seasonal offset:", reproj[1])
print("Deseasonalised offset:", reproj_deseasonal[1])
```
The closer we are to zero, the less offset from the origin our data are.
## Seasonal subseries plot
A seasonal subseries plot groups data by period (in this case there are 12 months in the year, so the period is 12), and then plots each group. This can help detect both seasonality and change in seasonality over time.
```
plt.figure(figsize=(2 * 6.4, 4.8))
titles = ["seasonal", "deseasonalised"]
monthly_means = [[], []]
for i, ts_ in enumerate([ts, ts_deseasonal]):
plt.subplot(1, 2, i + 1)
colours = matplotlib.cm.rainbow(np.linspace(0, 1, 12))
for month, t in ts_.groupby(ts_.index.month):
plt.plot(
month + np.linspace(0, 1, len(t)),
t.rolling(10).mean(),
c=colours[month - 1],
)
plt.plot([month, month + 1], [t.mean()] * 2, c="k")
plt.plot(
[month + 0.5] * 2,
[t.mean() - t.std(), t.mean() + t.std()],
c="k",
alpha=0.5,
)
monthly_means[i].append(t.mean())
plt.xlabel("Month")
plt.xticks(np.arange(1.5, 13.5), calendar.month_abbr[1:])
plt.ylabel("Percentage wet")
plt.title(titles[i])
```
The seasonal plot again shows very clear maxima during winter and minima during summer. The deseasonalised plot has no such pattern, and the differences between monthly means are within standard error.
We can also aggregate this plot into a single number representing seasonality. There are many ways to do this — any measure of how deviant the means are from a horizontal line would work, for example. One such measure would be the average deviation from the mean:
```
monthly_means = np.array(monthly_means)
mad = abs(monthly_means - monthly_means.mean(axis=1, keepdims=True)).mean(axis=1)
print("Seasonal mean average deviation:", mad[0].round(2))
print("Deseasonalised mean average deviation:", mad[1].round(2))
```
The closer to zero, the less seasonal the data are.
---
## Additional information
**License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
Digital Earth Australia data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
**Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/GeoscienceAustralia/dea-notebooks).
**Last modified:** September 2021
## Tags
Browse all available tags on the DEA User Guide's [Tags Index](https://docs.dea.ga.gov.au/genindex.html)
| github_jupyter |
# Step 1: Import Imporant Libraries and Preparation
```
%matplotlib notebook
!pip install git+https://github.com/am1tyadav/tfutils.git
import tensorflow as tf
import numpy as np
import os
import tfutils
from matplotlib import pyplot as plt
from tensorflow.keras.layers import Dense, Flatten, Conv2D, BatchNormalization
from tensorflow.keras.layers import Conv2DTranspose, Reshape, LeakyReLU
from tensorflow.keras.models import Model, Sequential
from PIL import Image
print('TensoFlow version:', tf.__version__)
```
# Step 2: Importing and Plotting the Data
```
(x_train, y_train), (x_test, y_test) = tfutils.datasets.mnist.load_data(one_hot = False)
x_train = tfutils.datasets.mnist.load_subset([0], x_train, y_train)
x_test = tfutils.datasets.mnist.load_subset([0], x_test, y_test)
x = np.concatenate([x_train, x_test], axis = 0)
tfutils.datasets.mnist.plot_ten_random_examples(plt, x, np.zeros((x.shape[0], 1))).show()
```
# Step 3: Building Discriminator
```
discriminator = Sequential([
Conv2D(64, 3, strides = 2, input_shape = (28, 28, 1)),
LeakyReLU(),
BatchNormalization(),
Conv2D(128, 5, strides = 2),
LeakyReLU(),
BatchNormalization(),
Conv2D(256, 5, strides = 2),
LeakyReLU(),
BatchNormalization(),
Flatten(),
Dense(1, activation = 'sigmoid')
])
optim = tf.keras.optimizers.Adam(lr = 2e-4, beta_1 = 0.5)
discriminator.compile(loss = 'binary_crossentropy', optimizer = optim, metrics = ['accuracy'])
discriminator.summary()
```
# Step 4: Building Generator
```
generator = Sequential([
Dense(256, activation = 'relu', input_shape = (1, )),
Reshape((1, 1, 256)),
Conv2DTranspose(256, 5, activation = 'relu'),
BatchNormalization(),
Conv2DTranspose(128, 5, activation = 'relu'),
BatchNormalization(),
Conv2DTranspose(64, 5, strides = 2, activation = 'relu'),
BatchNormalization(),
Conv2DTranspose(32, 5, activation = 'relu'),
BatchNormalization(),
Conv2DTranspose(1, 4, activation = 'sigmoid')
])
generator.summary()
noise = np.random.randn(1, 1)
generated_image = generator.predict(noise)[0]
plt.figure()
plt.imshow(np.reshape(generated_image, (28, 28)), cmap = 'binary')
plt.show()
```
# Step 5: Building Generative Adversarial Network (GAN)
```
input_layer = tf.keras.layers.Input(shape = (1, ))
generator_output = generator(input_layer)
discriminator_output = discriminator(generator_output)
gan = Model(
input_layer,
discriminator_output
)
discriminator.trainable = False
gan.compile(loss = 'binary_crossentropy', optimizer = optim, metrics = ['accuracy'])
gan.summary()
```
# Step 6: Training the GAN
```
epochs = 25
batch_size = 128
steps_per_epoch = int(2 * x.shape[0] / batch_size)
print('Steps per epoch =', steps_per_epoch)
dp = tfutils.plotting.DynamicPlot(plt, 5, 5, (8, 8))
for e in range(epochs):
dp.start_of_epoch(e)
for step in range(steps_per_epoch):
true_examples = x[int(batch_size / 2) * step : int(batch_size / 2) * (step + 1)]
true_examples = np.reshape(true_examples, (true_examples.shape[0], 28, 28, 1))
noise = np.random.randn(int(batch_size / 2), 1)
generated_examples = generator.predict(noise)
x_batch = np.concatenate([generated_examples, true_examples], axis = 0)
y_batch = np.array([0] * int(batch_size / 2) + [1] * int(batch_size / 2))
indices = np.random.choice(range(batch_size), batch_size, replace = False)
x_batch = x_batch[indices]
y_batch = y_batch[indices]
discriminator.trainable = True
discriminator.train_on_batch(x_batch, y_batch)
discriminator.trainable = False
loss, _ = gan.train_on_batch(noise, np.ones((int(batch_size / 2), 1)))
_, acc = discriminator.evaluate(x_batch, y_batch, verbose = False)
noise = np.random.randn(1, 1)
generated_image = generator.predict(noise)[0]
generated_image = np.reshape(generated_image, (28, 28))
dp.end_of_epoch(generated_image, 'binary', 'DiscAcc:{:.2f}'.format(acc), 'GANLoss:{:.2f}'.format(loss))
```
| github_jupyter |
## Preprocessing
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
url = '/Users/arpanganguli/Documents/Professional/Analysis/ISLR/Datasets/USArrests.csv'
USArrests = pd.read_csv(url, index_col='Unnamed: 0')
USArrests.head()
```
***
## 8.a. Calculating proportion of variance explained (PVE) using PCA through method: pca.explained_variance_ratio_
```
import warnings
warnings.filterwarnings('ignore')
from sklearn.preprocessing import scale
from sklearn.decomposition import PCA
df = pd.DataFrame(scale(USArrests))
df.columns = USArrests.columns
df.index = USArrests.index
df.head()
df.describe().round(4)
pca = PCA(n_components=4)
pca_data = pca.fit_transform(df)
principaldf = pd.DataFrame(data = pca_data, columns = ['PC1', 'PC2', 'PC3', 'PC4'])
principaldf.head()
principaldf.info()
PVAR = principaldf.var()
PVAR
PSUM = np.sum(PVAR)
PSUM
PVE_method = pd.DataFrame([PVAR/PSUM]).T
PVE_method.columns = ['explained variance ratio']
PVE_method.index = principaldf.columns
PVE_method
```
***
## 8.a. Calculating proportion of variance explained (PVE) using PCA through formula: $\frac{\sum_{i=1}^n(\sum_{j=1}^p\phi_{jm}x_{ij})^2}{\sum_{j=1}^p\sum_{i=1}^nx_{ij}^2}$
```
loadings = pca.components_.T
loadings_df = pd.DataFrame(loadings, index=df.columns, columns=principaldf.columns)
loadings_df
# PC1
num = np.sum((np.dot(df, loadings_df.PC1))**2)
denomdf = pd.DataFrame()
for i in range(0, 50):
row_sum = np.sum(df.iloc[i]**2)
denomdf = denomdf.append(pd.DataFrame([row_sum]))
denomdf.columns = ['sums']
denomdf.reset_index(drop=True, inplace=True)
denom = denomdf.sum()
PVE_PC1 = num/denom
PVE_PC1
# PC2
num = np.sum((np.dot(df, loadings_df.PC2))**2)
denomdf = pd.DataFrame()
for i in range(0, 50):
row_sum = np.sum(df.iloc[i]**2)
denomdf = denomdf.append(pd.DataFrame([row_sum]))
denomdf.columns = ['sums']
denomdf.reset_index(drop=True, inplace=True)
denom = denomdf.sum()
PVE_PC2 = num/denom
PVE_PC2
# PC3
num = np.sum((np.dot(df, loadings_df.PC3))**2)
denomdf = pd.DataFrame()
for i in range(0, 50):
row_sum = np.sum(df.iloc[i]**2)
denomdf = denomdf.append(pd.DataFrame([row_sum]))
denomdf.columns = ['sums']
denomdf.reset_index(drop=True, inplace=True)
denom = denomdf.sum()
PVE_PC3 = num/denom
PVE_PC3
# PC4
num = np.sum((np.dot(df, loadings_df.PC4))**2)
denomdf = pd.DataFrame()
for i in range(0, 50):
row_sum = np.sum(df.iloc[i]**2)
denomdf = denomdf.append(pd.DataFrame([row_sum]))
denomdf.columns = ['sums']
denomdf.reset_index(drop=True, inplace=True)
denom = denomdf.sum()
PVE_PC4 = num/denom
PVE_PC4
PVE_formula = pd.DataFrame([PVE_PC1.values, PVE_PC2.values, PVE_PC3.values, PVE_PC4.values])
PVE_formula.columns = ['explained variance ratio']
PVE_formula.index = principaldf.columns
PVE_formula
```
**Therefore, PVE through both method and formula are the same.**
| github_jupyter |
```
# Mount Google Drive
from google.colab import drive # import drive from google colab
ROOT = "/content/drive" # default location for the drive
print(ROOT) # print content of ROOT (Optional)
drive.mount(ROOT) # we mount the google drive at /content/drive
!pip install pennylane
from IPython.display import clear_output
clear_output()
import os
def restart_runtime():
os.kill(os.getpid(), 9)
restart_runtime()
# %matplotlib inline
import matplotlib.pyplot as plt
from mpl_toolkits.axes_grid1 import make_axes_locatable
import pennylane as qml
from pennylane import numpy as np
from pennylane import RX, RY, RZ, CNOT
from pennylane.templates.embeddings import QAOAEmbedding
def feature_encoding_hamiltonian(features, wires):
for idx, w in enumerate(wires):
RX(features[idx], wires=w)
def ising_hamiltonian(weights, wires, l):
# ZZ coupling
CNOT(wires=[wires[1], wires[0]])
RZ(weights[l, 0], wires=wires[0])
CNOT(wires=[wires[1], wires[0]])
# local fields
for idx, w in enumerate(wires):
RY(weights[l, idx + 1], wires=w)
'''
def QAOAEmbedding(features, weights, wires):
repeat = len(weights)
for l in range(repeat):
# apply alternating Hamiltonians
feature_encoding_hamiltonian(features, wires)
ising_hamiltonian(weights, wires, l)
# repeat the feature encoding once more at the end
feature_encoding_hamiltonian(features, wires)
'''
```
# Loading Raw Data
```
import tensorflow as tf
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train_flatten = x_train.reshape(x_train.shape[0], x_train.shape[1]*x_train.shape[2])/255.0
x_test_flatten = x_test.reshape(x_test.shape[0], x_test.shape[1]*x_test.shape[2])/255.0
print(x_train_flatten.shape, y_train.shape)
print(x_test_flatten.shape, y_test.shape)
x_train_0 = x_train_flatten[y_train == 0]
x_train_1 = x_train_flatten[y_train == 1]
x_train_2 = x_train_flatten[y_train == 2]
x_train_3 = x_train_flatten[y_train == 3]
x_train_4 = x_train_flatten[y_train == 4]
x_train_5 = x_train_flatten[y_train == 5]
x_train_6 = x_train_flatten[y_train == 6]
x_train_7 = x_train_flatten[y_train == 7]
x_train_8 = x_train_flatten[y_train == 8]
x_train_9 = x_train_flatten[y_train == 9]
x_train_list = [x_train_0, x_train_1, x_train_2, x_train_3, x_train_4, x_train_5, x_train_6, x_train_7, x_train_8, x_train_9]
print(x_train_0.shape)
print(x_train_1.shape)
print(x_train_2.shape)
print(x_train_3.shape)
print(x_train_4.shape)
print(x_train_5.shape)
print(x_train_6.shape)
print(x_train_7.shape)
print(x_train_8.shape)
print(x_train_9.shape)
x_test_0 = x_test_flatten[y_test == 0]
x_test_1 = x_test_flatten[y_test == 1]
x_test_2 = x_test_flatten[y_test == 2]
x_test_3 = x_test_flatten[y_test == 3]
x_test_4 = x_test_flatten[y_test == 4]
x_test_5 = x_test_flatten[y_test == 5]
x_test_6 = x_test_flatten[y_test == 6]
x_test_7 = x_test_flatten[y_test == 7]
x_test_8 = x_test_flatten[y_test == 8]
x_test_9 = x_test_flatten[y_test == 9]
x_test_list = [x_test_0, x_test_1, x_test_2, x_test_3, x_test_4, x_test_5, x_test_6, x_test_7, x_test_8, x_test_9]
print(x_test_0.shape)
print(x_test_1.shape)
print(x_test_2.shape)
print(x_test_3.shape)
print(x_test_4.shape)
print(x_test_5.shape)
print(x_test_6.shape)
print(x_test_7.shape)
print(x_test_8.shape)
print(x_test_9.shape)
```
# Selecting the dataset
Output: X_train, Y_train, X_test, Y_test
```
X_train = np.concatenate((x_train_list[0][:500, :], x_train_list[1][:500, :]), axis=0)
Y_train = np.zeros((X_train.shape[0],))
Y_train[500:] += 1
X_train.shape, Y_train.shape
X_test = np.concatenate((x_test_list[0][:100, :], x_test_list[1][:100, :]), axis=0)
Y_test = np.zeros((X_test.shape[0],))
Y_test[500:] += 1
X_test.shape, Y_test.shape
```
# Dataset Preprocessing (Standardization + PCA)
## Standardization
```
def normalize(X, use_params=False, params=None):
"""Normalize the given dataset X
Args:
X: ndarray, dataset
Returns:
(Xbar, mean, std): tuple of ndarray, Xbar is the normalized dataset
with mean 0 and standard deviation 1; mean and std are the
mean and standard deviation respectively.
Note:
You will encounter dimensions where the standard deviation is
zero, for those when you do normalization the normalized data
will be NaN. Handle this by setting using `std = 1` for those
dimensions when doing normalization.
"""
if use_params:
mu = params[0]
std_filled = [1]
else:
mu = np.mean(X, axis=0)
std = np.std(X, axis=0)
#std_filled = std.copy()
#std_filled[std==0] = 1.
Xbar = (X - mu)/(std + 1e-8)
return Xbar, mu, std
X_train, mu_train, std_train = normalize(X_train)
X_train.shape, Y_train.shape
X_test = (X_test - mu_train)/(std_train + 1e-8)
X_test.shape, Y_train.shape
```
## PCA
```
from sklearn.decomposition import PCA
from matplotlib import pyplot as plt
num_component = 2
pca = PCA(n_components=num_component, svd_solver='full')
pca.fit(X_train)
np.cumsum(pca.explained_variance_ratio_)
X_train = pca.transform(X_train)
X_test = pca.transform(X_test)
print(X_train.shape, Y_train.shape)
print(X_test.shape, Y_test.shape)
```
# Quantum
```
```
| github_jupyter |
# Create Your Own Chatbot
In this notebook, you can create a series of intents with paraphrase generation and use those in a dialogflow agent
You may optionally mount google drive to use one of its directories as a file system
```
from google.colab import drive
drive.mount('/content/drive')
```
Use this block to declare any variables you will use throughout the notebook
```
# path to the directory you wish to use as a file system
BASE_PATH='/content/drive/MyDrive/VaccineFAQs-BigShot/questions'
# path to the base HTML/XML file from which to extract QAs
HTML_DOC_PATH = BASE_PATH + '/COVID-19-Vaccine-FAQs.html'
# path to a simple QA CSV file which will be the basis for agent generation
QA_CSV_SIMPLE_PATH = BASE_PATH + '/india-qa-simple.csv'
# path to an intermediate CSV file which will contain dialogflow intents
QA_CSV_FORMATTED_PATH = BASE_PATH + '/akuryla-qa-formatted.csv'
```
# Parsing an HTML File for Q/A Pairs (Example)
We'll use lxml to parse Q/A pairs out of our HTML site, and export our parsed data into a simple Q/A CSV
```
from lxml import html, etree
# utility method to print xml elements
def xml_print(elmt):
print(etree.tostring(elmt, pretty_print=True))
# we fetch the content from the html document, stored in drive
html_doc = open(HTML_DOC_PATH, mode='r')
doc = html.parse(html_doc, parser=html.html_parser)
# parse the html document into sets of Q/A pairs
qa_pairs = []
elmt_list=doc.xpath('/html/body/section[4]/div[1]/div/div/div[2]/div[2]/div/div/div/span/div')
for elmt in elmt_list:
elmt_children = elmt.xpath('div')
qa_pairs.append([elmt_children[0].text_content(), elmt_children[1].text_content()])
# utility function to print Q/A pairs
def print_pairs(pairs):
for q, a in qa_pairs:
print("Q:", q)
print("A:", a, "\n")
print_pairs(qa_pairs)
import csv
# write Q/A pairs to csv
def write_pairs_to_csv(qa_pairs, csv_file_path):
with open(csv_file_path, 'w') as csvfile:
filewriter = csv.writer(csvfile)
# write header
filewriter.writerow(['Question', 'Answer'])
# write contents
for entry in qa_pairs:
filewriter.writerow(entry)
# write_pairs_to_csv(qa_pairs, QA_CSV_SIMPLE_PATH)
```
# Generate Intents for Dialogflow Agent
- Parse Q/A pairs from our simple CSV file
- Generate paraphrases for each question
- Output a list of dialogflow intents to our formatted CSV file
- You may manually edit the formatted CSV file after this step to ensure all intents are up to standard
```
import csv
# read entries from csv
def read_entries_from_csv(csv_file_path):
with open(csv_file_path, 'r') as csvfile:
filereader = csv.reader(csvfile)
faqs = []
for row in filereader:
faqs.append({"question": row[0].replace('/', ''), "answer": row[1].replace('/', '')})
return faqs[1:]
faqs = read_entries_from_csv(QA_CSV_SIMPLE_PATH)
faqs
!pip install transformers==2.8.0
import torch
from transformers import T5ForConditionalGeneration,T5Tokenizer
def set_seed(seed):
torch.manual_seed(seed)
if torch.cuda.is_available():
torch.cuda.manual_seed_all(seed)
# set seed to make examples reproducible
set_seed(42)
# load pretrained model for text-to-text conversion
model = T5ForConditionalGeneration.from_pretrained('ramsrigouthamg/t5_paraphraser')
tokenizer = T5Tokenizer.from_pretrained('ramsrigouthamg/t5_paraphraser')
# if GPU is available, set global device to GPU (cuda)
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print ("device ",device)
# move model into current device
model = model.to(device)
def paraphrase_sentence(sentence, max_length=256, num_return_sequences=10):
# generate input encoding
text = "paraphrase: " + sentence + " </s>"
encoding = tokenizer.encode_plus(text, return_tensors="pt")
input_ids, attention_masks = encoding["input_ids"].to(device), encoding["attention_mask"].to(device)
# specify parameters for paraphrasing model and generate tokenized paraphrases
beam_outputs = model.generate(
input_ids=input_ids, attention_mask=attention_masks,
do_sample=True,
max_length=max_length,
top_k=120,
top_p=0.98,
early_stopping=True,
num_return_sequences=num_return_sequences
)
# iterate through results, decode and filter out repeated paraphrases
final_outputs =[]
for beam_output in beam_outputs:
sent = tokenizer.decode(beam_output, skip_special_tokens=True,clean_up_tokenization_spaces=True)
if sent.lower() != sentence.lower() and sent not in final_outputs:
final_outputs.append(sent)
return final_outputs
# generate all paraphrases
for entry in faqs:
entry['paraphrases'] = paraphrase_sentence(entry['question'])
# write faqs in desired format
def write_formatted_faq_to_csv(faqs, csv_file_path):
with open(csv_file_path, 'w') as csvfile:
filewriter = csv.writer(csvfile)
# write header
filewriter.writerow(['IntentID', 'IntentName', 'Query', 'Response'])
# write contents
for idx, entry in enumerate(faqs):
filewriter.writerow([str(idx + 1), entry['question'].replace('/', ''), \
entry['question'].replace('/', ''), entry['answer']])
for paraphrase in entry['paraphrases']:
filewriter.writerow([str(idx + 1), '', paraphrase, ''])
write_formatted_faq_to_csv(faqs, QA_CSV_FORMATTED_PATH)
```
# Generate Dialogflow Agent
- Specify output path
- Read intents from formatted CSV
- specify keywords to filter out unnecessary/incorrectly paraphrased questions
- Add default intents and format output
- Generate an importable Dialogflow agent
```
# specify output folder name
OUTPUT_FOLDER_NAME = 'akuryla_usa_output'
output_path = BASE_PATH + '/' + OUTPUT_FOLDER_NAME
intent_path = output_path + '/intents'
import csv
def read_formatted_csv_to_faq(csv_file_path):
with open(csv_file_path, 'r') as csvfile:
filereader = csv.reader(csvfile)
faqs = []
curr_entry = {}
curr_intent_id = '-1'
for idx, row in enumerate(filereader):
# skip headers
if idx == 0:
continue
# check whether it's a new question
if curr_intent_id != row[0]:
# set current intent id
curr_intent_id = row[0]
# if entry is not empty, add to faqs
if curr_entry:
faqs.append(curr_entry)
# initialize current entry
curr_entry = {'question': row[1], 'answer': row[3], 'paraphrases': []}
else:
# add paraphrase to curr_entry
curr_entry['paraphrases'].append(row[2])
return faqs
faqs = read_formatted_csv_to_faq(QA_CSV_FORMATTED_PATH)
faqs
def add_default_intents(faqs):
faqs.append({
"question": "Default Welcome Intent",
"answer": "Greetings! I am Vaccine chatbot. You can ask me questions about COVID-19 vaccines such as vaccine safety, side effects, immunity and allergies.",
"paraphrases": ["Hi",
"Hello",
"Hi there",
"Hey there",
"Heya",
"Howdy",
"How are you?",
"Just going to say hi"]})
faqs.append({
"question": "Default Fallback Intent",
"answer": "I'm sorry, I don't think I can answer that question. Please try again.",
"paraphrases": []})
faqs.append({
"question": "End Session",
"answer": "",
"paraphrases": ["OK",
"Thank you",
"That's enough",
"Good bye",
"Bye",
"See you",
"Stop",
"No"]})
# you may filter out paraphrases by keyword
def should_filter(phrase, keywords):
return any(elmt.lower() in phrase.lower() for elmt in keywords)
def filter_keywords(faqs, keywords):
for faq in faqs:
faq['paraphrases'] = [x for x in faq['paraphrases'] if not should_filter(x, keywords)]
add_default_intents(faqs)
keywords = ["HIV", "AIDS", "cholera", "cattle", "cow", "colibid", "covirid", \
"covarid", "SVDC-19"]
filter_keywords(faqs, keywords)
faqs
import os
def make_folder_if_absent(output_path, intent_path):
if not os.path.exists(output_path):
os.makedirs(output_path)
if not os.path.exists(intent_path):
os.makedirs(intent_path)
make_folder_if_absent(output_path, intent_path)
import uuid
# there will be one question entity element per paraphrased/original question
def make_question_entity_element(question):
return {
"id": str(uuid.uuid1()),
"data": [
{
"text": question,
"userDefined": False
}
],
"isTemplate": False,
"count": 0,
"lang": "en",
"updated": 0
}
# a question entity is a collection of question entity elements
def make_question_entity(question, paraphrases):
question_entity = []
question_entity.append(make_question_entity_element(question))
for paraphrase in paraphrases:
question_entity.append(make_question_entity_element(paraphrase))
return question_entity
# answer entities typically consist of a single object
def make_answer_entity(intent_name, answer):
return {
"id": str(uuid.uuid1()),
"name": intent_name,
"auto": True,
"contexts": [],
"responses": [
{
"resetContexts": False,
"action": "",
"affectedContexts": [],
"parameters": [],
"messages": [
{
"type": "0",
"title": "",
"textToSpeech": "",
"lang": "en",
"speech": [answer],
"condition": ""
}
],
"speech": []
}
],
"priority": 500000,
"webhookUsed": False,
"webhookForSlotFilling": False,
"fallbackIntent": False,
"events": [],
"conditionalResponses": [],
"condition": "",
"conditionalFollowupEvents": []
}
def format_answer(question, answer):
# only format non-default answers
if question.startswith("Default Welcome Intent") \
or question.startswith("Default Fallback Intent"):
return answer
else:
return "Q: " + question + "\nA: " + answer
for idx, entry in enumerate(faqs):
entry['intent_name'] = f"VaccineFAQ.{entry['question']}"[:56]
entry['question_entity'] = make_question_entity(entry['question'], entry['paraphrases'])
entry['answer_entity'] = make_answer_entity(entry['intent_name'], format_answer(entry['question'], entry['answer']))
import json
agent={
"description": "",
"language": "en",
"shortDescription": "",
"examples": "",
"linkToDocs": "",
"displayName": "Vaccine-Bot-FAQ",
"disableInteractionLogs": False,
"disableStackdriverLogs": True,
"defaultTimezone": "America/New_York",
"isPrivate": False,
"mlMinConfidence": 0.3,
"supportedLanguages": ["en"],
"enableOnePlatformApi": True,
"onePlatformApiVersion": "v2beta1",
"secondaryKey": "9d74e6a3640d4ce3807cf42e2fdcea79",
"analyzeQueryTextSentiment": False,
"enabledKnowledgeBaseNames": [],
"knowledgeServiceConfidenceAdjustment": 0.0,
"dialogBuilderMode": False,
"baseActionPackagesUrl": "",
"enableSpellCorrection": False
}
package={
"version": "1.0.0"
}
def write_to_output_path(faqs):
with open(output_path + "/agent.json", 'w') as outfile:
json.dump(agent, outfile)
with open(output_path + "/package.json", 'w') as outfile:
json.dump(package, outfile)
for entry in faqs:
with open(output_path + "/intents/" + entry["intent_name"] + "_usersays_en.json", 'w') as outfile:
json.dump(entry['question_entity'], outfile)
with open(output_path + "/intents/" + entry["intent_name"] + ".json", 'w') as outfile:
json.dump(entry['answer_entity'], outfile)
write_to_output_path(faqs)
```
| github_jupyter |
# Rate of Returns Over Multiple Periods
## Numpy.cumsum and Numpy.cumprod
You've just leared about active returns and passive returns. Another important concept related to returns is "Cumulative returns" which is defined as the returns over a time period. You can read more about rate of returns [here](https://en.wikipedia.org/wiki/Rate_of_return)!
There are two ways to calcualte cumulative returns, depends on how the returns are calculated. Let's take a look at an example.
```
import numpy as np
import pandas as pd
from datetime import datetime
dates = pd.date_range(datetime.strptime('1/1/2016', '%m/%d/%Y'), periods=12, freq='M')
start_price, stop_price = 0.24, 0.3
abc_close_prices = np.arange(start_price, stop_price, (stop_price - start_price)/len(dates))
abc_close = pd.Series(abc_close_prices, dates)
abc_close
```
Here, we have the historical prices for stock ABC for 2016. We would like to know the yearly cumulative returns for stock ABC in 2016 using time-weighted method, assuming returns are reinvested. How do we do it? Here is the formula:
Assume the returns over n successive periods are:
$ r_1, r_2, r_3, r_4, r_5, ..., r_n $
The cumulative return of stock ABC over period n is the compounded return over period n:
$ (1 + r_1)(1 + r_2)(1 + r_3)(1 + r_4)(1 + r_5)...(1 + r_n) - 1 $
First, let's calculate the returns of stock ABC.
```
returns = abc_close / abc_close.shift(1) - 1
returns
```
The cumulative return equals to the product of the daily returns for the n periods.
That's a very long formula. Is there a better way to calculate this.
The answer is yes, we can use numpy.cumprod().
For example, if we have the following time series: 1, 5, 7, 10 and we want the product of the four numbers. How do we do it? Let's take a look!
```
lst = [1,5,7,10]
np.cumprod(lst)
```
The last element in the list is 350, which is the product of 1, 5, 7, and 10.
OK, let's use numpy.cumprod() to get the cumulative returns for stock ABC
```
(returns + 1).cumprod()[len(returns)-1] - 1
```
The cumulative return for stock ABC in 2016 is 22.91%.
The other way to calculate returns is to use log returns.
The formula of log return is the following:
$ LogReturn = ln(\frac{P_t}{P_t - 1}) $
The cumulative return of stock ABC over period n is the compounded return over period n:
$ \sum_{i=1}^{n} r_i = r_1 + r_2 + r_3 + r_4 + ... + r_n $
Let's see how we can calculate the cumulative return of stock ABC using log returns.
First, let's calculate log returns.
```
log_returns = (np.log(abc_close).shift(-1) - np.log(abc_close)).dropna()
log_returns.head()
```
The cumulative sum equals to the sum of the daily returns for the n periods which is a very long formula.
To calculate cumulative sum, we can simply use numpy.cumsum().
Let's take a look at our simple example of time series 1, 5, 7, 10.
```
lst = [1,5,7,10]
np.cumsum(lst)
```
The last element is 23 which equals to the sum of 1, 5, 7, 10
OK, let's use numpy.cumsum() to get the cumulative returns for stock ABC
```
cum_log_return = log_returns.cumsum()[len(log_returns)-1]
np.exp(cum_log_return) - 1
```
The cumulative return for stock ABC in 2016 is 22.91% using log returns.
## Quiz: Arithmetic Rate of Return
Now, let's use cumprod() and cumsum() to calculate average rate of return.
For consistency, let's assume the rate of return is calculated as $ \frac{P_t}{P_t - 1} - 1 $
### Arithmetic Rate of Return:
$ \frac{1}{n} \sum_{i=1}^{n} r_i = \frac{1}{n}(r_1 + r_2 + r_3 + r_4 + ... + r_n) $
```
import quiz_tests
def calculate_arithmetic_rate_of_return(close):
"""
Compute returns for each ticker and date in close.
Parameters
----------
close : DataFrame
Close prices for each ticker and date
Returns
-------
arithmnetic_returns : Series
arithmnetic_returns at the end of the period for each ticker
"""
# TODO: Implement Function
returns = close / close.shift(1) - 1
arithmetic_returns = returns.cumsum(axis =0).iloc[returns.shape[0]-1]/returns.shape[0]
return arithmetic_returns
quiz_tests.test_calculate_arithmetic_rate_of_return(calculate_arithmetic_rate_of_return)
```
## Quiz Solution
If you're having trouble, you can check out the quiz solution [here](cumsum_and_cumprod_solution.ipynb).
| github_jupyter |
<small><small><i>
All the IPython Notebooks in **[Python Seaborn Module](https://github.com/milaan9/12_Python_Seaborn_Module)** lecture series by **[Dr. Milaan Parmar](https://www.linkedin.com/in/milaanparmar/)** are available @ **[GitHub](https://github.com/milaan9)**
</i></small></small>
<a href="https://colab.research.google.com/github/milaan9/12_Python_Seaborn_Module/blob/main/005_Seaborn_LM_Plot_and_Reg_Plot.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# LM Plot and Reg Plot
Welcome to another lecture on Seaborn! This is going to be the first among a series of plots that we shall be drawing with Seaborn. In this lecture, we shall be covering the concept of plotting **Linear Regression** data analysis, which is a very common method in *Business Intelligence*, and *Data Science* domain in particular. To begin with, we shall at first try to gain *statistical overview* of the concept of *Linear Regression*.
As our intention isn't to dive deeply into each statistical concept, I shall instead pick a curated dataset and show you different ways in which we can visualize whatever we deduced during our analysis. Using Seaborn, there are two important types of figure that we can plot to fulfil our project needs. One is known as **LM Plot** and the other one is **Reg Plot**. Visualy, they have pretty much similar appearance, but do have functional difference that I will highlight in detail for you to understand.
**Linear Regression** is a *statistical concept for predictive analytics*, where the core agenda is to majorly examine three aspects:
- Does a set of predictor variables do a good job in predicting an outcome (dependent) variable?
- Which variables in particular are significant predictors for the outcome variable?
- In what way do they (indicated by the magnitude and sign of the beta estimates) impact the outcome variable? These **Beta Estimates** are just the *standardized coefficients* resulting from a *regression analysis*, that have been standardized so that the variances of dependent and independent variables are 1.
Let us begin by importing the libraries that we might need in our journey and this is something you will find me doing at the start of every lecture so that we don't have to bother about dependancies throughout the lecture.
```
# Importing intrinsic libraries:
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set(style="whitegrid", palette="hsv")
import warnings
warnings.filterwarnings("ignore")
```
Let us now generate some data to play around with using **NumPy** for two imaginary classes of points. Please note that throughout the course I wouldn't be explaining Data generation as that is a component of Data Analysis. With that been said, let us try to plot something here:
```
# Loading Built-in Dataset:
tips = sns.load_dataset("tips")
# Fetching preview of Dataset:
tips.head(10)
```
## Seaborn Lmplots:
Every plot in Seaborn has a set of fixed parameters. For **`sns.lmplot()`**, we have three manadatory parameters and the rest are optional that we may use as per our requirements. These 3 parameters are values for X-axis, values for Y-axis and reference to dataset. These 3 are pre-dominantly visible in almost all of Seaborn plots and in addition, there is an optional parameter which I want you to memorize as it comes in very handy. This is **hue** parameter and it takes in categorical columns and kind of helps us to group our data plot as per *hue* parameter values.
Let me show you how it works:
```
# Basic lmplot visualization:
sns.lmplot(x="total_bill", y="tip", data=tips)
```
Let us now understand what we see on the screen before we jump into adding parameters. This linear line across our plot is the best available fit for the trend of the tip usually customers give with respect to the total bill that gets generated. And the data points that we see at extreme top right which are far away from this line are known as **outliers** in the dataset. You may think of *outliers* as exceptions.
Goal of Data Science is to predict the best fit for understanding the trend in behavior of visiting customers and our algorithm shall be always designed accordingly. You may find this a common scenario while applying Logistic Regression algorithms in Machine Learning. If you very cllosely notice there is this shadow converging at the centre where there is a chunk of our data. This covergent point is actually the statistical mean or in simpler words, the generalized prediction of tip value in this restaurant on a daily basis.
In this case, looking at this plot, we may say that if the total bill is around $20.00, then it shall get a tip of around $3.00. Let us refine this visualization even further by adding more features to the plot, and for this purpose let us try to understand if a Smoker in general *tip* more or *less*:
```
sns.lmplot(x="total_bill", y="tip", hue="smoker", data=tips)
```
Somehow it refects that Smokers that you may see in blue are little more generous but not so consistent with their tipping mechanism as the data points are quite vaguely spread out. So, addition of the 3rd parameter of **hue** helped us visualize this difference in separate color plotting, and has also added a **legend** with *Yes*, *No* to conveniently interpret.
Let us look into other commonly used parameters to customize this plot further:
```
sns.lmplot(x="total_bill", y="tip", hue="smoker", data=tips, markers=["o", "x"], palette="Set1", legend=False)
```
Here, we set data point marker style, altered the coloring and decided to remove the legend which by default is always there. Right now, be it for a smoker or for a non-smoker, the representation is on the same plot so let us get it on separate facets:
```
sns.lmplot(x="total_bill", y="tip", col="smoker", data=tips)
```
There is a lot that you may experiment with by using different optional parameters, but in a shell, basic presentation with mandatory arguments remain the same. Let me show you one more on Tips dataset:
```
sns.lmplot(x="total_bill", y="tip", palette="magma", row="sex", col="time", data=tips, size=3)
```
This plot in 4 separate facets drill deeper into visualizing the data, where we still show the tip being given against total bill but is now also segmented into whether it was Lunch time or not along with dependency on Gender. There shall be multiple occassions where you would like to visualize such a deeper segmentation. Currently we have a small dataset so we still have our hands tied but with real-world dataset exploration, this visualization gets limitless.
Now, I shall show you a generic usage of **`lmplot()`** where we shall generate random data points and then fit a regression line across it. I am showing this implementation just to give an overview of how it generally looks like in production environment and if you're a beginner and not so proficient with Python programming, don't really need to get stressed because with time, you shall gain command over it.
Please note that our focus is just on visualization, so we won't really get into **NumPy** module usage. Let's get started:
```
# Generating Random Data points:
def generatingData():
num_points =1500
category_points =[]
for i in range(num_points):
if np.random.random()>0.5:
x,y = np.random.normal(0.0, 0.9), np.random.normal(0.0, 0.9)
category_points.append([x,y])
else:
x, y = np.random.normal(3.0, 0.5), np.random.normal(1.0, 0.5)
category_points.append([x, y])
df =pd.DataFrame({'x':[v[0] for v in category_points], 'y':
[v[1] for v in category_points]})
sns.lmplot('x', 'y', data=df, fit_reg=True, size=6)
plt.show()
generatingData()
```
Here we see jumbled up data points on the plot with a linearly fitted line passing through, thus reflecting best fit for existing trend as per dataset. In general, I would always recommend to keep the sequence of parameters intact as per **[sns.lmplot()](https://seaborn.pydata.org/generated/seaborn.lmplot.html)** official documentation which looks pretty much like this:
**`seaborn.lmplot(x, y, data, hue=None, col=None, row=None, palette=None, col_wrap=None, size=5, aspect=1, markers='o', sharex=True, sharey=True, hue_order=None, col_order=None, row_order=None, legend=True, legend_out=True, x_estimator=None, x_bins=None, x_ci='ci', scatter=True, fit_reg=True, ci=95, n_boot=1000, units=None, order=1, logistic=False, lowess=False, robust=False, logx=False, x_partial=None, y_partial=None, truncate=False, x_jitter=None, y_jitter=None, scatter_kws=None, line_kws=None)`**
Here the values that we see against few optional parameters are there by default, unless we specifically alter it in our code. Also, we need to always make sure that **`x`** and **`y`** feature values should always be **strings** to maintain tidy data format. If you feel curious to know in depth about **tidy data**, I would suggest reading a research paper by Hadley Wickham that titles **[Journal of Statistical Software](http://vita.had.co.nz/papers/tidy-data.pdf)**. I have attached the link to access it's PDF format in the notebook.
## Seaborn Regplots:
In terms of core functionality, **`reglot()`** is pretty similar to **`lmplot()`** and solves similar purpose of visualizing a linear relationship as determined through Regression. In the simplest invocation, both functions draw a scatterplot of two variables, **`x`** and **`y`**, and then fit the regression model **`y ~ x`**; and plot the resulting regression line and a *95% confidence interval* for that regression. In fact, **`regplot()`** possesses a subset of **`lmplot()`** features.
Important to note is the difference between these two functions in order to choose the correct plot for your usage.
- Very evident difference is the shape of plot that we shall observe shortly.
- Secondly, **[regplot()](https://seaborn.pydata.org/generated/seaborn.regplot.html)** has mandatory input parameter flexibility. This means that **`x`** and **`y`** variables DO NOT necessarily require *strings* as *input*. Unlike **`lmplot()`**, these two parameters shall also accept other formats like simple *NumPy arrays*, *Pandas Series* objects, or as references to variables in a *Pandas DataFrame* object passed to input data.
The parameters for **`regplot()`** as per it's official documentation with all it's parameters look like this:
**`seaborn.regplot(x, y, data=None, x_estimator=None, x_bins=None, x_ci='ci', scatter=True, fit_reg=True, ci=95, n_boot=1000, units=None, order=1, logistic=False, lowess=False, robust=False, logx=False, x_partial=None, y_partial=None, truncate=False, dropna=True, x_jitter=None, y_jitter=None, label=None, color=None, marker='o', scatter_kws=None, line_kws=None, ax=None)`**
There isn't much of a visual difference so let's quickly plot a **`regplot()`** to understand it better. But before I do that, I would like you to make a note of the fact that **`Seaborn regplot()`** or **`lmplot()`** **does not support regression against date data** so if you're dealing with *Time-series algorithms*, please make a careful choice. Also note that **`lmplot()`** that we just finished discussing is just a wrapper around **`regplot()`** and **`facetgrid()`**, that we shall be taking up later in this course.
```
sns.regplot(x="total_bill", y="tip", data=tips, color="g")
```
For a change, let us also try to plot with *NumPy arrays*:
```
import numpy as np
np.random.seed(8) # Initializing RandomState
mean, cov = [6, 8], [(1.5, .7), (.7, 1)] # Mean and Covariance
x, y = np.random.multivariate_normal(mean, cov, 150).T # Generalizing 1-Dimensional Gaussian distribution to higher dimensions.
sns.regplot(x=x, y=y, color="r")
```
With the declaration itself, we know what we have plotted and aesthetically it is something we have already discussed in detail and that brings us to the end of this lecture where we have in length gone through two important aspects of Regression plotting. Third one, i.e. facetgrid is parked in the list and we shall take that as well very soon.
The datasets we have dealt with till now have data points pretty neatly arranged and hence presenting a logistic fit isn't that cumbersome but let us now look at few complex scenarios. The very first one we are going to deal with is to fit a nonparametric regression using a **[lowess smoother](https://en.wikipedia.org/wiki/Local_regression)**.
```
sns.lmplot(x="total_bill", y="tip", data=tips, lowess=True)
```
This is a *computationally intensive* process as it is robust and hence in the backend it doesn't take **`ci`** parameter, i.e. *confidence interval* into consideration. Here the line bends around to get more precise estimate as per the spread of data points, as visible. Let us get another built-in dataset available with Seaborn to have a better view of applied Logistic Regression scenarios:
```
# Loading another Built-in dataset:
anscombe = sns.load_dataset("anscombe")
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'II'"), order=3, ci=None, scatter_kws={"s": 70})
```
This majorly helps to tackle *Outliers* in our dataset to fit a **polynomial regression model** to explore simple kinds of nonlinear trends because the linear relationship is the same but our simple plot wouldn't have been able to trace it. Let me show how it would have looked:
```
sns.lmplot(x="x", y="y", data=anscombe.query("dataset == 'II'"), ci=None, scatter_kws={"s": 70})
```
With all that understanding, the only thing I feel I should get you acquainted with are the commonly used optional parameters:
- Parameters like **`x_jitter`** and **`y_jitter`** are used to add noise to our dataset.
- **`color`** parameter helps you get Matplotlib style color.
- **`dropna`** helps to drop NaN (NULL) values.
- **`x_estimator`** param is useful with discrete variables.
- **`ci`** represents the size of Confidence interval used when plotting a **[central tendency](https://en.wikipedia.org/wiki/Central_tendency)** for discrete values of x.
- **`label`** is used to assign a suitable name to either our *Scatterplot* (in *legends*) or *Regression line*.
Thank You for your patience throughout this visually exhaustive lecture and hope to see you in the next one with **[Scatter Plot and Joint Plot](https://github.com/milaan9/12_Python_Seaborn_Module/blob/main/006_Seaborn_Scatter_Plot_and_Joint_Plot.ipynb)**.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from fastai.vision.all import *
from shopee_utils import *
from train_utils import *
import sklearn.metrics as skm
from tqdm.notebook import tqdm
from fastai.vision.learner import _resnet_split
import h5py
import timm
import debugpy
debugpy.listen(5678)
def efficientnet_b0(pretrained): return timm.create_model('efficientnet_b0', pretrained=pretrained)
def efficientnet_b1(pretrained): return timm.create_model('efficientnet_b1', pretrained=pretrained)
def efficientnet_b2(pretrained): return timm.create_model('efficientnet_b2', pretrained=pretrained)
class conf():
bs = 64
#'arch':resnet34,
arch = efficientnet_b0
arcface_m=.4
arcface_s=30
train_df = add_splits(pd.read_csv(PATH/'train.csv'))
def get_img_file(row):
img =row.image
fn = PATH/'train_images'/img
if not fn.is_file():
fn = PATH/'test_images'/img
return fn
data_block = DataBlock(blocks = (ImageBlock(), CategoryBlock(vocab=train_df.label_group.to_list())),
splitter=ColSplitter(),
#splitter=RandomSplitter(),
get_y=ColReader('label_group'),
get_x=get_img_file,
item_tfms=Resize(460),
batch_tfms=aug_transforms(size=224, min_scale=0.75),
)
dls = data_block.dataloaders(train_df, bs=conf.bs,num_workers=16)
class ArcFaceClassifier(nn.Module):
def __init__(self, in_features, output_classes):
super().__init__()
self.W = nn.Parameter(torch.Tensor(in_features, output_classes))
nn.init.kaiming_uniform_(self.W)
def forward(self, x):
x_norm = F.normalize(x)
W_norm = F.normalize(self.W, dim=0)
return x_norm @ W_norm
class ResnetArcFace(nn.Module):
def __init__(self):
super().__init__()
self.body = create_body(conf.arch, cut=-2)
nf = num_features_model(nn.Sequential(*self.body.children()))
self.after_conv=nn.Sequential(
AdaptiveConcatPool2d(),
Flatten(),
nn.BatchNorm1d(nf*2),
nn.Dropout(.25))
self.classifier = ArcFaceClassifier(nf*2, dls.c)
self.outputEmbs = False
def forward(self, x):
x = self.body(x)
embeddings = self.after_conv(x)
if self.outputEmbs:
return embeddings
return self.classifier(embeddings)
def split_2way(model):
return L(params(model.body),
params(model.classifier))
def modules_params(modules):
return list(itertools.chain(*modules.map(params)))
def split_b0(model):
body =model.body
b0_children = list(body.children())
convs = b0_children[3]
group1 =L(b0_children[:3]) + L(convs[:2])
group2 = L(convs[2:]) + L(b0_children[4:])
group3 = L([model.after_conv,model.classifier])
return [modules_params(g) for g in [group1,group2,group3]]
#opt_func=RMSProp
opt_func=Adam
loss_func=functools.partial(arcface_loss, m=conf.arcface_m, s=conf.arcface_s)
learn = Learner(dls,ResnetArcFace(),splitter=split_b0,
opt_func=opt_func, loss_func=arcface_loss, cbs = F1FromEmbs, metrics=FakeMetric())
learn.fine_tune(8,1e-2, lr_mult=50)
learn.save('b0_788')
learn.load('resnet34_arcface')
```
# VALIDATION
```
model = learn.model.eval().cuda()
model.outputEmbs = True
embs, y = embs_from_model(model, dls.valid)
f1_from_embs(embs,y, True)
```
| github_jupyter |
# "Tableau rocks"
> "And I normally hate stuff like this"
- toc: true
- badges: true
- comments: true
- categories: [Misc]
I’m not the worlds biggest fan of all singing and dancing viz dashboards, but the fact is that there are going to be times when they are useful, and at some point you are probably going to work under important people who *love* visualizations.
I’m very impressed with Tableau. To be honest, when you’ve been around for as long as I have, you get used to be various bits of software like this being market leaders in their little niches because they’re the least dreadful option at what they do. That was depressingly normal for a long time although I’m glad to say it’s getting much rarer.
Still it’s so nice to find something which just works the way it should (and all the more surprising to find that Google has a rival product which is actually *really* bad!).
Below is a mickey-mouse hello world viz which literally takes about 2 mins to produce but it gives you an idea of the kind of thing you can produce and the way people can interact with them.
If you’re new and you’ve never seen Tableau you can go and download the free version [here](https://public.tableau.com/en-us/s/).
<div class='tableauPlaceholder' id='viz1597432657044' style='position: relative'><noscript><a href='#'><img alt=' ' src='https://public.tableau.com/static/images/2K/2KW38M37D/1_rss.png' style='border: none' /></a></noscript><object class='tableauViz' style='display:none;'><param name='host_url' value='https%3A%2F%2Fpublic.tableau.com%2F' /> <param name='embed_code_version' value='3' /> <param name='path' value='shared/2KW38M37D' /> <param name='toolbar' value='yes' /><param name='static_image' value='https://public.tableau.com/static/images/2K/2KW38M37D/1.png' /> <param name='animate_transition' value='yes' /><param name='display_static_image' value='yes' /><param name='display_spinner' value='yes' /><param name='display_overlay' value='yes' /><param name='display_count' value='yes' /><param name='language' value='en' /></object></div> <script type='text/javascript'> var divElement = document.getElementById('viz1597432657044'); var vizElement = divElement.getElementsByTagName('object')[0]; if ( divElement.offsetWidth > 800 ) { vizElement.style.minWidth='420px';vizElement.style.maxWidth='650px';vizElement.style.width='100%';vizElement.style.minHeight='587px';vizElement.style.maxHeight='887px';vizElement.style.height=(divElement.offsetWidth*0.75)+'px';} else if ( divElement.offsetWidth > 500 ) { vizElement.style.minWidth='420px';vizElement.style.maxWidth='650px';vizElement.style.width='100%';vizElement.style.minHeight='587px';vizElement.style.maxHeight='887px';vizElement.style.height=(divElement.offsetWidth*0.75)+'px';} else { vizElement.style.width='100%';vizElement.style.height='727px';} var scriptElement = document.createElement('script'); scriptElement.src = 'https://public.tableau.com/javascripts/api/viz_v1.js'; vizElement.parentNode.insertBefore(scriptElement, vizElement); </script>
| github_jupyter |
```
# Imports
import matplotlib.pyplot as plt
from ptsnet.simulation.sim import PTSNETSimulation
from ptsnet.utils.io import get_example_path
```
## Simulation Settings
Simulation settings can be defined using a dictionary. We show below the default simulation settings.
```
default_settings = {
"time_step" : 0.01, # Simulation time step in [s]
"duration" : 20, # Simulation duration in [s]
"warnings_on" : False,
"skip_compatibility_check" : False, # Dismisses the compatibility check to run a faster initialization
"show_progress" : False, # Shows progress (Warnings should be off)
"save_results" : True, # Saves numerical results in HDF5 format
"profiler_on" : False, # Measures computational times of the simulation
"period" : 0, # Simulation period for EPS
"default_wave_speed" : 1000, # Wave speed value for all pipes in [m/s]
"wave_speed_file_path" : None, # Wave speeds can be defined on a separated file with CSV format
"delimiter" : ',', # Delimiter of text file with wave speed data | pipe,wave_speed_value
"wave_speed_method" : 'optimal' # Method to compute the wave speed adjustment
}
```
## Running a Simulation
The function `PTSNETSimulation` creates a new PTSNET simulation that reads an input file (.inp) located in the path `inpfile` and parses it. Users can use the test cases provided by the library which include:
`["B0", "B0_SURGE", "B1_0", "B3", "B0_0", "B1_1", "B4", "PIPE_IN_SERIES", "TNET3_HAMMER", "B0_1", "B2", "LOOP"]`. Paths to the test cases can be extracted using the `get_example_path(ex_name)` function.
```
# Create a simulation
sim = PTSNETSimulation(
inpfile = get_example_path('TNET3_HAMMER'),
settings = default_settings # If settings are not defined, default settings are loaded automatically
)
print(sim)
```
## Valve Closure
Users can manipulate specific valves of the system via the `define_valve_operation` function. The function allows to manipulate one or many pipes (e.g., `sim.all_pipes`). By default the closure occurs linearly varying the setting of the valve.
```
sim = PTSNETSimulation(inpfile = get_example_path('TNET3_HAMMER'))
sim.define_valve_operation('VALVE-179', initial_setting=1, final_setting=0, start_time=0, end_time=1)
sim.run()
# Plot results
plt.plot(sim['time'], sim['node'].head['JUNCTION-23'], label='JUNCTION-23')
plt.xlabel('Time [s]'); plt.ylabel('Head [m]')
plt.legend()
plt.show()
```
## Pump shut-off
Users can shut-off pumps using the `define_pump_operation`. By default the shut-off occurs linearly varying the setting of the pump.
```
sim = PTSNETSimulation(inpfile = get_example_path('TNET3_HAMMER'))
sim.define_pump_operation('PUMP-172', initial_setting=1, final_setting=0, start_time=0, end_time=2)
sim.run()
# Plot results
plt.plot(sim['time'], sim['node'].head['JUNCTION-34'], label='JUNCTION-34')
plt.xlabel('Time [s]'); plt.ylabel('Head [m]')
plt.legend()
plt.show()
```
## Open Surge Tanks
We generate a transient by closing all the system's valves in 1 s. We add an open surge protection on node N7, with cross-section area, $A_T = 0.1$ $\text{m}^2$
```
sim = PTSNETSimulation(inpfile = get_example_path('TNET3_HAMMER'))
sim.define_pump_operation('PUMP-172', initial_setting=1, final_setting=0, start_time=0, end_time=1)
sim.add_surge_protection('JUNCTION-34', 'open', 0.1)
sim.run()
plt.plot(sim['time'], sim['node'].head['JUNCTION-34'], label='JUNCTION-34')
plt.plot(sim['time'], sim['node'].head['JUNCTION-30'], label='JUNCTION-30')
plt.plot(sim['time'], sim['node'].head['JUNCTION-1'], label='JUNCTION-1')
plt.xlabel('Time [s]')
plt.ylabel('Head [m]')
plt.legend()
plt.show()
```
## Closed Surge Tanks
We generate a transient by closing all the system's valves in 1 s. We add an closed surge protection on node N7, with cross-section area $A_T = 0.1 \text{ m}^2$, height $H_T = 1 \text{ m}$, and initial water level $H_W = 0.2\text{ m}$.
```
sim = PTSNETSimulation(inpfile = get_example_path('TNET3_HAMMER'))
sim.define_pump_operation('PUMP-172', initial_setting=1, final_setting=0, start_time=0, end_time=1)
sim.add_surge_protection('JUNCTION-34', 'closed', 0.1, 1, 0.2)
sim.run()
plt.plot(sim['time'], sim['node'].head['JUNCTION-34'], label='JUNCTION-34')
plt.plot(sim['time'], sim['node'].head['JUNCTION-30'], label='JUNCTION-30')
plt.plot(sim['time'], sim['node'].head['JUNCTION-1'], label='JUNCTION-1')
plt.plot(sim['time'], sim['node'].head['JUNCTION-4'], label='JUNCTION-4')
plt.xlabel('Time [s]')
plt.ylabel('Head [m]')
plt.legend()
plt.show()
```
## Leaks
Leaks can only be modeled as emitters via the .inp file
## Bursts
Users can add bursts defining `start_time` and `end_time` values which define how the burst is going to develop. Users have to define the final loss coefficient associated with the burst.
```
sim = PTSNETSimulation(inpfile = get_example_path('TNET3_HAMMER'), settings={'duration':20})
sim.add_burst('JUNCTION-90', 0.02, 0, 1)
sim.run()
plt.plot(sim['time'], sim['node'].head['JUNCTION-90'], label='JUNCTION-90')
plt.plot(sim['time'], sim['node'].head['JUNCTION-34'], label='JUNCTION-34')
plt.plot(sim['time'], sim['node'].head['JUNCTION-30'], label='JUNCTION-30')
plt.plot(sim['time'], sim['node'].head['JUNCTION-4'], label='JUNCTION-4')
plt.xlabel('Time [s]')
plt.ylabel('Head [m]')
plt.legend()
plt.show()
```
| github_jupyter |
## Предобработка данных и логистическая регрессия для задачи бинарной классификации
## Programming assignment
В задании вам будет предложено ознакомиться с основными техниками предобработки данных, а так же применить их для обучения модели логистической регрессии. Ответ потребуется загрузить в соответствующую форму в виде 6 текстовых файлов.
Для выполнения задания требуется Python версии 2.7 или 3.5, а также актуальные версии библиотек:
- NumPy: 1.10.4 и выше
- Pandas: 0.17.1 и выше
- Scikit-learn: 0.17 и выше
```
import pandas as pd
import numpy as np
import matplotlib
from matplotlib import pyplot as plt
matplotlib.style.use('ggplot')
%matplotlib inline
np.__version__
```
## Описание датасета
Задача: по 38 признакам, связанных с заявкой на грант (область исследований учёных, информация по их академическому бэкграунду, размер гранта, область, в которой он выдаётся) предсказать, будет ли заявка принята. Датасет включает в себя информацию по 6000 заявкам на гранты, которые были поданы в университете Мельбурна в период с 2004 по 2008 год.
Полную версию данных с большим количеством признаков можно найти на https://www.kaggle.com/c/unimelb.
```
data = pd.read_csv('data.csv')
data.shape
```
Выделим из датасета целевую переменную Grant.Status и обозначим её за y
Теперь X обозначает обучающую выборку, y - ответы на ней
```
X = data.drop('Grant.Status', 1)
y = data['Grant.Status']
```
## Теория по логистической регрессии
После осознания того, какую именно задачу требуется решить на этих данных, следующим шагом при реальном анализе был бы подбор подходящего метода. В данном задании выбор метода было произведён за вас, это логистическая регрессия. Кратко напомним вам используемую модель.
Логистическая регрессия предсказывает вероятности принадлежности объекта к каждому классу. Сумма ответов логистической регрессии на одном объекте для всех классов равна единице.
$$ \sum_{k=1}^K \pi_{ik} = 1, \quad \pi_k \equiv P\,(y_i = k \mid x_i, \theta), $$
где:
- $\pi_{ik}$ - вероятность принадлежности объекта $x_i$ из выборки $X$ к классу $k$
- $\theta$ - внутренние параметры алгоритма, которые настраиваются в процессе обучения, в случае логистической регрессии - $w, b$
Из этого свойства модели в случае бинарной классификации требуется вычислить лишь вероятность принадлежности объекта к одному из классов (вторая вычисляется из условия нормировки вероятностей). Эта вероятность вычисляется, используя логистическую функцию:
$$ P\,(y_i = 1 \mid x_i, \theta) = \frac{1}{1 + \exp(-w^T x_i-b)} $$
Параметры $w$ и $b$ находятся, как решения следующей задачи оптимизации (указаны функционалы с L1 и L2 регуляризацией, с которыми вы познакомились в предыдущих заданиях):
L2-regularization:
$$ Q(X, y, \theta) = \frac{1}{2} w^T w + C \sum_{i=1}^l \log ( 1 + \exp(-y_i (w^T x_i + b ) ) ) \longrightarrow \min\limits_{w,b} $$
L1-regularization:
$$ Q(X, y, \theta) = \sum_{d=1}^D |w_d| + C \sum_{i=1}^l \log ( 1 + \exp(-y_i (w^T x_i + b ) ) ) \longrightarrow \min\limits_{w,b} $$
$C$ - это стандартный гиперпараметр модели, который регулирует то, насколько сильно мы позволяем модели подстраиваться под данные.
## Предобработка данных
Из свойств данной модели следует, что:
- все $X$ должны быть числовыми данными (в случае наличия среди них категорий, их требуется некоторым способом преобразовать в вещественные числа)
- среди $X$ не должно быть пропущенных значений (т.е. все пропущенные значения перед применением модели следует каким-то образом заполнить)
Поэтому базовым этапом в предобработке любого датасета для логистической регрессии будет кодирование категориальных признаков, а так же удаление или интерпретация пропущенных значений (при наличии того или другого).
```
data.head()
```
Видно, что в датасете есть как числовые, так и категориальные признаки. Получим списки их названий:
```
numeric_cols = ['RFCD.Percentage.1', 'RFCD.Percentage.2', 'RFCD.Percentage.3',
'RFCD.Percentage.4', 'RFCD.Percentage.5',
'SEO.Percentage.1', 'SEO.Percentage.2', 'SEO.Percentage.3',
'SEO.Percentage.4', 'SEO.Percentage.5',
'Year.of.Birth.1', 'Number.of.Successful.Grant.1', 'Number.of.Unsuccessful.Grant.1']
categorical_cols = list(set(X.columns.values.tolist()) - set(numeric_cols))
```
Также в нём присутствуют пропущенные значения. Очевидны решением будет исключение всех данных, у которых пропущено хотя бы одно значение. Сделаем это:
```
data.dropna().shape
```
Видно, что тогда мы выбросим почти все данные, и такой метод решения в данном случае не сработает.
Пропущенные значения можно так же интерпретировать, для этого существует несколько способов, они различаются для категориальных и вещественных признаков.
Для вещественных признаков:
- заменить на 0 (данный признак давать вклад в предсказание для данного объекта не будет)
- заменить на среднее (каждый пропущенный признак будет давать такой же вклад, как и среднее значение признака на датасете)
Для категориальных:
- интерпретировать пропущенное значение, как ещё одну категорию (данный способ является самым естественным, так как в случае категорий у нас есть уникальная возможность не потерять информацию о наличии пропущенных значений; обратите внимание, что в случае вещественных признаков данная информация неизбежно теряется)
## Задание 0. Обработка пропущенных значений.
1. Заполните пропущенные вещественные значения в X нулями и средними по столбцам, назовите полученные датафреймы X_real_zeros и X_real_mean соответственно.
2. Все категориальные признаки в X преобразуйте в строки, пропущенные значения требуется также преобразовать в какие-либо строки, которые не являются категориями (например, 'NA'), полученный датафрейм назовите X_cat.
```
# place your code here
X_real_zeros = X[numeric_cols].fillna(0)
X_real_mean = X[numeric_cols].fillna(X.mean())
X_cat = X[categorical_cols].astype(str).fillna('lol') ##не заполняет наны, но код работает энивей
#X_cat = data[categorical_cols].fillna('NA').applymap(lambda s: str(s))` - вот так можно заполнить
# X_cat.shape == X_real_zeros.shape
X_cat.head()
```
## Преобразование категориальных признаков.
В предыдущей ячейке мы разделили наш датасет ещё на две части: в одной присутствуют только вещественные признаки, в другой только категориальные. Это понадобится нам для раздельной последующей обработке этих данных, а так же для сравнения качества работы тех или иных методов.
Для использования модели регрессии требуется преобразовать категориальные признаки в вещественные. Рассмотрим основной способ преоборазования категориальных признаков в вещественные: one-hot encoding. Его идея заключается в том, что мы преобразуем категориальный признак при помощи бинарного кода: каждой категории ставим в соответствие набор из нулей и единиц.
Посмотрим, как данный метод работает на простом наборе данных.
```
from sklearn.linear_model import LogisticRegression as LR
from sklearn.feature_extraction import DictVectorizer as DV
categorial_data = pd.DataFrame({'sex': ['male', 'female', 'male', 'female'],
'nationality': ['American', 'European', 'Asian', 'European']})
print('Исходные данные:\n')
print(categorial_data)
encoder = DV(sparse = False)
encoded_data = encoder.fit_transform(categorial_data.T.to_dict().values())
print('\nЗакодированные данные:\n')
print(encoded_data)
```
Как видно, в первые три колонки оказалась закодированна информация о стране, а во вторые две - о поле. При этом для совпадающих элементов выборки строки будут полностью совпадать. Также из примера видно, что кодирование признаков сильно увеличивает их количество, но полностью сохраняет информацию, в том числе о наличии пропущенных значений (их наличие просто становится одним из бинарных признаков в преобразованных данных).
Теперь применим one-hot encoding к категориальным признакам из исходного датасета. Обратите внимание на общий для всех методов преобработки данных интерфейс. Функция
encoder.fit_transform(X)
позволяет вычислить необходимые параметры преобразования, впоследствии к новым данным можно уже применять функцию
encoder.transform(X)
Очень важно применять одинаковое преобразование как к обучающим, так и тестовым данным, потому что в противном случае вы получите непредсказуемые, и, скорее всего, плохие результаты. В частности, если вы отдельно закодируете обучающую и тестовую выборку, то получите вообще говоря разные коды для одних и тех же признаков, и ваше решение работать не будет.
Также параметры многих преобразований (например, рассмотренное ниже масштабирование) нельзя вычислять одновременно на данных из обучения и теста, потому что иначе подсчитанные на тесте метрики качества будут давать смещённые оценки на качество работы алгоритма. Кодирование категориальных признаков не считает на обучающей выборке никаких параметров, поэтому его можно применять сразу к всему датасету.
```
encoder = DV(sparse = False)
X_cat_oh = encoder.fit_transform(X_cat.T.to_dict().values())
```
Для построения метрики качества по результату обучения требуется разделить исходный датасет на обучающую и тестовую выборки.
Обращаем внимание на заданный параметр для генератора случайных чисел: random_state. Так как результаты на обучении и тесте будут зависеть от того, как именно вы разделите объекты, то предлагается использовать заранее определённое значение для получение результатов, согласованных с ответами в системе проверки заданий.
```
from sklearn.cross_validation import train_test_split
(X_train_real_zeros,
X_test_real_zeros,
y_train, y_test) = train_test_split(X_real_zeros, y,
test_size=0.3,
random_state=0)
(X_train_real_mean,
X_test_real_mean) = train_test_split(X_real_mean,
test_size=0.3,
random_state=0)
(X_train_cat_oh,
X_test_cat_oh) = train_test_split(X_cat_oh,
test_size=0.3,
random_state=0)
```
## Описание классов
Итак, мы получили первые наборы данных, для которых выполнены оба ограничения логистической регрессии на входные данные. Обучим на них регрессию, используя имеющийся в библиотеке sklearn функционал по подбору гиперпараметров модели
optimizer = GridSearchCV(estimator, param_grid)
где:
- estimator - обучающий алгоритм, для которого будет производиться подбор параметров
- param_grid - словарь параметров, ключами которого являются строки-названия, которые передаются алгоритму estimator, а значения - набор параметров для перебора
Данный класс выполняет кросс-валидацию обучающей выборки для каждого набора параметров и находит те, на которых алгоритм работает лучше всего. Этот метод позволяет настраивать гиперпараметры по обучающей выборке, избегая переобучения. Некоторые опциональные параметры вызова данного класса, которые нам понадобятся:
- scoring - функционал качества, максимум которого ищется кросс валидацией, по умолчанию используется функция score() класса esimator
- n_jobs - позволяет ускорить кросс-валидацию, выполняя её параллельно, число определяет количество одновременно запущенных задач
- cv - количество фолдов, на которые разбивается выборка при кросс-валидации
После инициализации класса GridSearchCV, процесс подбора параметров запускается следующим методом:
optimizer.fit(X, y)
На выходе для получения предсказаний можно пользоваться функцией
optimizer.predict(X)
Также можно напрямую получить оптимальный класс estimator и оптимальные параметры, так как они является атрибутами класса GridSearchCV:
- best\_estimator\_ - лучший алгоритм
- best\_params\_ - лучший набор параметров
Класс логистической регрессии выглядит следующим образом:
estimator = LogisticRegression(penalty)
где penalty принимает либо значение 'l2', либо 'l1'. По умолчанию устанавливается значение 'l2', и везде в задании, если об этом не оговорено особо, предполагается использование логистической регрессии с L2-регуляризацией.
## Задание 1. Сравнение способов заполнения вещественных пропущенных значений.
1. Составьте две обучающие выборки из вещественных и категориальных признаков: в одной вещественные признаки, где пропущенные значения заполнены нулями, в другой - средними.
2. Обучите на них логистическую регрессию, подбирая параметры из заданной сетки param_grid по методу кросс-валидации с числом фолдов cv=3. В качестве оптимизируемой функции используйте заданную по умолчанию.
3. Постройте два графика оценок точности +- их стандратного отклонения в зависимости от гиперпараметра и убедитесь, что вы действительно нашли её максимум. Также обратите внимание на большую дисперсию получаемых оценок (уменьшить её можно увеличением числа фолдов cv).
4. Получите две метрики качества AUC ROC на тестовой выборке и сравните их между собой. Какой способ заполнения пропущенных вещественных значений работает лучше? В дальнейшем для выполнения задания в качестве вещественных признаков используйте ту выборку, которая даёт лучшее качество на тесте.
5. Передайте два значения AUC ROC (сначала для выборки, заполненной средними, потом для выборки, заполненной нулями) в функцию write_answer_1 и запустите её. Полученный файл является ответом на 1 задание.
Информация для интересующихся: вообще говоря, не вполне логично оптимизировать на кросс-валидации заданный по умолчанию в классе логистической регрессии функционал accuracy, а измерять на тесте AUC ROC, но это, как и ограничение размера выборки, сделано для ускорения работы процесса кросс-валидации.
```
from sklearn.linear_model import LogisticRegression
from sklearn.grid_search import GridSearchCV
from sklearn.metrics import roc_auc_score
def plot_scores(optimizer):
scores = [[item[0]['C'],
item[1],
(np.sum((item[2]-item[1])**2)/(item[2].size-1))**0.5] for item in optimizer.grid_scores_]
scores = np.array(scores)
plt.semilogx(scores[:,0], scores[:,1])
plt.fill_between(scores[:,0], scores[:,1]-scores[:,2],
scores[:,1]+scores[:,2], alpha=0.3)
plt.show()
def write_answer_1(auc_1, auc_2):
answers = [auc_1, auc_2]
with open("preprocessing_lr_answer1.txt", "w") as fout:
fout.write(" ".join([str(num) for num in answers]))
param_grid = {'C': [0.01, 0.05, 0.1, 0.5, 1, 5, 10]}
cv = 3
# place your code here
clf = LogisticRegression()
gridCVZeros = GridSearchCV(clf, param_grid, scoring = 'accuracy', cv = cv)
gridCVMeans = GridSearchCV(clf, param_grid, scoring = 'accuracy', cv = cv)
X_final_zeros = np.hstack((X_train_real_zeros, X_train_cat_oh))
X_final_means = np.hstack((X_train_real_mean, X_train_cat_oh))
gridCVZeros.fit(X_final_zeros, y_train)
gridCVMeans.fit(X_final_means, y_train)
# print X.shape
# print X_train_real_zeros.shape
# X_train_cat_oh.shape
# plot_scores(gridCVZeros)
pred_zeros = gridCVZeros.predict(np.hstack((X_test_real_zeros, X_test_cat_oh)))
pred_means = gridCVMeans.predict(np.hstack((X_test_real_mean, X_test_cat_oh)))
print roc_auc_score(y_test, pred_means), roc_auc_score(y_test, pred_zeros)
write_answer_1(roc_auc_score(y_test, pred_means), roc_auc_score(y_test, pred_zeros))
```
## Масштабирование вещественных признаков.
Попробуем как-то улучшить качество классификации. Для этого посмотрим на сами данные:
```
from pandas.tools.plotting import scatter_matrix
data_numeric = pd.DataFrame(X_train_real_zeros, columns=numeric_cols)
list_cols = ['Number.of.Successful.Grant.1', 'SEO.Percentage.2', 'Year.of.Birth.1']
scatter_matrix(data_numeric[list_cols], alpha=0.5, figsize=(10, 10))
plt.show()
```
Как видно из графиков, разные признаки очень сильно отличаются друг от друга по модулю значений (обратите внимание на диапазоны значений осей x и y). В случае обычной регрессии это никак не влияет на качество обучаемой модели, т.к. у меньших по модулю признаков будут большие веса, но при использовании регуляризации, которая штрафует модель за большие веса, регрессия, как правило, начинает работать хуже.
В таких случаях всегда рекомендуется делать стандартизацию (масштабирование) признаков, для того чтобы они меньше отличались друг друга по модулю, но при этом не нарушались никакие другие свойства признакового пространства. При этом даже если итоговое качество модели на тесте уменьшается, это повышает её интерпретабельность, потому что новые веса имеют смысл "значимости" данного признака для итоговой классификации.
Стандартизация осуществляется посредством вычета из каждого признака среднего значения и нормировки на выборочное стандартное отклонение:
$$ x^{scaled}_{id} = \dfrac{x_{id} - \mu_d}{\sigma_d}, \quad \mu_d = \frac{1}{N} \sum_{i=1}^l x_{id}, \quad \sigma_d = \sqrt{\frac{1}{N-1} \sum_{i=1}^l (x_{id} - \mu_d)^2} $$
## Задание 1.5. Масштабирование вещественных признаков.
1. По аналогии с вызовом one-hot encoder примените масштабирование вещественных признаков для обучающих и тестовых выборок X_train_real_zeros и X_test_real_zeros, используя класс
StandardScaler
и методы
StandardScaler.fit_transform(...)
StandardScaler.transform(...)
2. Сохраните ответ в переменные X_train_real_scaled и X_test_real_scaled соответственно
```
from sklearn.preprocessing import StandardScaler
# place your code here
scaler = StandardScaler()
X_train_real_scaled = scaler.fit_transform(X_train_real_zeros)
X_test_real_scaled = scaler.transform(X_test_real_zeros)
```
## Сравнение признаковых пространств.
Построим такие же графики для преобразованных данных:
```
data_numeric_scaled = pd.DataFrame(X_train_real_scaled, columns=numeric_cols)
list_cols = ['Number.of.Successful.Grant.1', 'SEO.Percentage.2', 'Year.of.Birth.1']
scatter_matrix(data_numeric_scaled[list_cols], alpha=0.5, figsize=(10, 10))
plt.show()
```
Как видно из графиков, мы не поменяли свойства признакового пространства: гистограммы распределений значений признаков, как и их scatter-plots, выглядят так же, как и до нормировки, но при этом все значения теперь находятся примерно в одном диапазоне, тем самым повышая интерпретабельность результатов, а также лучше сочетаясь с идеологией регуляризации.
## Задание 2. Сравнение качества классификации до и после масштабирования вещественных признаков.
1. Обучите ещё раз регрессию и гиперпараметры на новых признаках, объединив их с закодированными категориальными.
2. Проверьте, был ли найден оптимум accuracy по гиперпараметрам во время кроссвалидации.
3. Получите значение ROC AUC на тестовой выборке, сравните с лучшим результатом, полученными ранее.
4. Запишите полученный ответ в файл при помощи функции write_answer_2.
```
def write_answer_2(auc):
with open("preprocessing_lr_answer2.txt", "w") as fout:
fout.write(str(auc))
# place your code here
gridCVScaled = GridSearchCV(clf, param_grid, scoring = 'accuracy', cv = cv)
X_final_scaled = np.hstack((X_train_real_scaled, X_train_cat_oh))
gridCVScaled.fit(X_final_scaled, y_train)
predScaled = gridCVScaled.predict(np.hstack((X_test_real_scaled, X_test_cat_oh)))
print roc_auc_score(y_test, predScaled)
write_answer_2(roc_auc_score(y_test, predScaled))
# print roc_auc_score(predScaled, y_test)
```
## Балансировка классов.
Алгоритмы классификации могут быть очень чувствительны к несбалансированным классам. Рассмотрим пример с выборками, сэмплированными из двух гауссиан. Их мат. ожидания и матрицы ковариации заданы так, что истинная разделяющая поверхность должна проходить параллельно оси x. Поместим в обучающую выборку 20 объектов, сэмплированных из 1-й гауссианы, и 10 объектов из 2-й. После этого обучим на них линейную регрессию, и построим на графиках объекты и области классификации.
```
np.random.seed(0)
"""Сэмплируем данные из первой гауссианы"""
data_0 = np.random.multivariate_normal([0,0], [[0.5,0],[0,0.5]], size=40)
"""И из второй"""
data_1 = np.random.multivariate_normal([0,1], [[0.5,0],[0,0.5]], size=40)
"""На обучение берём 20 объектов из первого класса и 10 из второго"""
example_data_train = np.vstack([data_0[:20,:], data_1[:10,:]])
example_labels_train = np.concatenate([np.zeros((20)), np.ones((10))])
"""На тест - 20 из первого и 30 из второго"""
example_data_test = np.vstack([data_0[20:,:], data_1[10:,:]])
example_labels_test = np.concatenate([np.zeros((20)), np.ones((30))])
"""Задаём координатную сетку, на которой будем вычислять область классификации"""
xx, yy = np.meshgrid(np.arange(-3, 3, 0.02), np.arange(-3, 3, 0.02))
"""Обучаем регрессию без балансировки по классам"""
optimizer = GridSearchCV(LogisticRegression(), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
"""Строим предсказания регрессии для сетки"""
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
"""Считаем AUC"""
auc_wo_class_weights = roc_auc_score(example_labels_test, optimizer.predict(example_data_test))
plt.title('Without class weights')
plt.show()
print('AUC: %f'%auc_wo_class_weights)
"""Для второй регрессии в LogisticRegression передаём параметр class_weight='balanced'"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced'), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
auc_w_class_weights = roc_auc_score(example_labels_test, optimizer.predict(example_data_test))
plt.title('With class weights')
plt.show()
print('AUC: %f'%auc_w_class_weights)
```
Как видно, во втором случае классификатор находит разделяющую поверхность, которая ближе к истинной, т.е. меньше переобучается. Поэтому на сбалансированность классов в обучающей выборке всегда следует обращать внимание.
Посмотрим, сбалансированны ли классы в нашей обучающей выборке:
```
print(np.sum(y_train==0))
print(np.sum(y_train==1))
```
Видно, что нет.
Исправить ситуацию можно разными способами, мы рассмотрим два:
- давать объектам миноритарного класса больший вес при обучении классификатора (рассмотрен в примере выше)
- досэмплировать объекты миноритарного класса, пока число объектов в обоих классах не сравняется
## Задание 3. Балансировка классов.
1. Обучите логистическую регрессию и гиперпараметры с балансировкой классов, используя веса (параметр class_weight='balanced' регрессии). Убедитесь, что вы нашли максимум accuracy по гиперпараметрам
2. Получите метрику ROC AUC на тестовой выборке.
3. Сбалансируйте выборку, досэмплировав в неё объекты из меньшего класса. Для получения индексов объектов, которые требуется добавить в обучающую выборку, используйте следующую комбинацию вызовов функций:
np.random.seed(0)
indices_to_add = np.random.randint(...)
4. Получите метрику ROC AUC на тестовой выборке, сравните с предыдущим результатом.
5. Внесите ответы в выходной файл при помощи функции write_asnwer_3, передав в неё сначала ROC AUC для балансировки весами, а потом балансировки выборки вручную.
```
X_final_scaled = np.hstack((X_train_real_scaled, X_train_cat_oh))
X_test_final = np.hstack((X_test_real_scaled, X_test_cat_oh))
def write_answer_3(auc_1, auc_2):
answers = [auc_1, auc_2]
with open("preprocessing_lr_answer3.txt", "w") as fout:
fout.write(" ".join([str(num) for num in answers]))
# place your code here
clf = LogisticRegression(class_weight='balanced')
param_grid = {'C': [0.01, 0.05, 0.1, 0.5, 1, 5, 10]}
cv = 3
###захуярим сюда X_real_zeroes
# X_final_scaled = np.hstack((X_train_real_zeros, X_train_cat_oh))
# X_test_final = np.hstack((X_test_real_zeros, X_test_cat_oh))
# X_final_scaled = np.hstack((X_train_real_scaled, X_train_cat_oh))
# X_test_final = np.hstack((X_test_real_scaled, X_test_cat_oh))
gridCV = GridSearchCV(clf, param_grid, scoring = 'accuracy', cv = cv)
gridCV.fit(X_final_scaled, y_train)
auc1 = roc_auc_score(y_test, gridCV.predict(X_test_final))
auc1
# np.random.seed(0)
# ind = y_train[y_train==1].index
# toadd = np.random.randint(len(ind),size=432)
# new_y = y_train[y_train==1].iloc[toadd]
# new_X = X_final_scaled[np.array(y_train == 1)][toadd]
np.random.seed(0)
y_train_0 = np.sum(y_train==0)
y_train_1 = np.sum(y_train==1)
X_man_sampled = X_final_scaled
y_man_sampled = y_train
while y_train_1 < y_train_0:
idx = np.random.randint(y_train_1)
if y_train.iloc[idx] == 1:
new_X = X_final_scaled[idx]
new_y = y_train.iloc[idx]
X_man_sampled = np.vstack((X_man_sampled, new_X))
y_man_sampled = np.hstack((y_man_sampled, new_y))
y_train_1 += 1
X_man_sampled.shape
##Add new lines to X and y:
# X_man_sampled = np.vstack((X_final_scaled, new_X))
# y_train_sampled = np.hstack((y_train, new_y))
# toadd2
# print(np.sum(y_train_sampled==0))
# print(np.sum(y_train_sampled==1))
clf1 = LogisticRegression()
# param_grid = {'C': [0.01, 0.05, 0.1, 0.5, 1, 5, 10]}
# cv = 3
# X_test_final = np.hstack((X_test_real_scaled, X_test_cat_oh))
gridCV_w = GridSearchCV(clf1, param_grid, scoring = 'accuracy', cv = cv)
gridCV_w.fit(X_man_sampled, y_man_sampled)
pred_w2 = gridCV_w.predict(X_test_final)
auc2 = roc_auc_score(y_test, pred_w2)
auc2
write_answer_3(auc1, auc2)
# np.sum(y_train_sampled==1) == np.sum(y_train_sampled==0)
```
## Стратификация выборок.
Рассмотрим ещё раз пример с выборками из нормальных распределений. Посмотрим ещё раз на качество классификаторов, получаемое на тестовых выборках:
```
print('AUC ROC for classifier without weighted classes', auc_wo_class_weights)
print('AUC ROC for classifier with weighted classes: ', auc_w_class_weights)
```
Насколько эти цифры реально отражают качество работы алгоритма, если учесть, что тестовая выборка так же несбалансирована, как обучающая? При этом мы уже знаем, что алгоритм логистический регрессии чувствителен к балансировке классов в обучающей выборке, т.е. в данном случае на тесте он будет давать заведомо заниженные результаты. Метрика классификатора на тесте имела бы гораздо больший смысл, если бы объекты были разделы в выборках поровну: по 20 из каждого класса на обучени и на тесте. Переформируем выборки и подсчитаем новые ошибки:
```
"""Разделим данные по классам поровну между обучающей и тестовой выборками"""
example_data_train = np.vstack([data_0[:20,:], data_1[:20,:]])
example_labels_train = np.concatenate([np.zeros((20)), np.ones((20))])
example_data_test = np.vstack([data_0[20:,:], data_1[20:,:]])
example_labels_test = np.concatenate([np.zeros((20)), np.ones((20))])
"""Обучим классификатор"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced'), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train, example_labels_train)
Z = optimizer.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
auc_stratified = roc_auc_score(example_labels_test, optimizer.predict(example_data_test))
plt.title('With class weights')
plt.show()
print('AUC ROC for stratified samples: ', auc_stratified)
```
Как видно, после данной процедуры ответ классификатора изменился незначительно, а вот качество увеличилось. При этом, в зависимости от того, как вы разбили изначально данные на обучение и тест, после сбалансированного разделения выборок итоговая метрика на тесте может как увеличиться, так и уменьшиться, но доверять ей можно значительно больше, т.к. она построена с учётом специфики работы классификатора. Данный подход является частным случаем т.н. метода стратификации.
## Задание 4. Стратификация выборки.
1. По аналогии с тем, как это было сделано в начале задания, разбейте выборки X_real_zeros и X_cat_oh на обучение и тест, передавая в функцию
train_test_split(...)
дополнительно параметр
stratify=y
Также обязательно передайте в функцию переменную random_state=0.
2. Выполните масштабирование новых вещественных выборок, обучите классификатор и его гиперпараметры при помощи метода кросс-валидации, делая поправку на несбалансированные классы. Убедитесь в том, что нашли оптимум accuracy по гиперпараметрам.
3. Оцените качество классификатора метрике AUC ROC на тестовой выборке.
4. Полученный ответ передайте функции write_answer_4
```
def write_answer_4(auc):
with open("preprocessing_lr_answer4.txt", "w") as fout:
fout.write(str(auc))
# place your code here
(X_train_real_zeros,
X_test_real_zeros,
y_train, y_test) = train_test_split(X_real_zeros, y,
test_size=0.3,
random_state=0, stratify=y)
(X_train_cat_oh,
X_test_cat_oh) = train_test_split(X_cat_oh,
test_size=0.3,
random_state=0, stratify=y)
scaler = StandardScaler()
X_train_strat = np.hstack((scaler.fit_transform(X_train_real_zeros), X_train_cat_oh))
X_test_strat = np.hstack((scaler.transform(X_test_real_zeros), X_test_cat_oh))
gridCVstrat = GridSearchCV(LogisticRegression(class_weight='balanced'), param_grid, cv=cv, n_jobs=-1)
gridCVstrat.fit(X_train_strat, y_train)
auc_strat = roc_auc_score(y_test, gridCVstrat.predict(X_test_strat))
print auc_strat
write_answer_4(auc_strat)
```
Теперь вы разобрались с основными этапами предобработки данных для линейных классификаторов.
Напомним основные этапы:
- обработка пропущенных значений
- обработка категориальных признаков
- стратификация
- балансировка классов
- масштабирование
Данные действия с данными рекомендуется проводить всякий раз, когда вы планируете использовать линейные методы. Рекомендация по выполнению многих из этих пунктов справедлива и для других методов машинного обучения.
## Трансорфмация признаков.
Теперь рассмотрим способы преобразования признаков. Существует достаточно много различных способов трансформации признаков, которые позволяют при помощи линейных методов получать более сложные разделяющие поверхности. Самым базовым является полиномиальное преобразование признаков. Его идея заключается в том, что помимо самих признаков вы дополнительно включаете набор все полиномы степени $p$, которые можно из них построить. Для случая $p=2$ преобразование выглядит следующим образом:
$$ \phi(x_i) = [x_{i,1}^2, ..., x_{i,D}^2, x_{i,1}x_{i,2}, ..., x_{i,D}, x_{i,D-1}, x_{i,1}, ..., x_{i,D}, 1] $$
Рассмотрим принцип работы данных признаков на данных, сэмплированных их гауссиан:
```
from sklearn.preprocessing import PolynomialFeatures
"""Инициализируем класс, который выполняет преобразование"""
transform = PolynomialFeatures(2)
"""Обучаем преобразование на обучающей выборке, применяем его к тестовой"""
example_data_train_poly = transform.fit_transform(example_data_train)
example_data_test_poly = transform.transform(example_data_test)
"""Обращаем внимание на параметр fit_intercept=False"""
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced', fit_intercept=False), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train_poly, example_labels_train)
Z = optimizer.predict(transform.transform(np.c_[xx.ravel(), yy.ravel()])).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
plt.title('With class weights')
plt.show()
```
Видно, что данный метод преобразования данных уже позволяет строить нелинейные разделяющие поверхности, которые могут более тонко подстраиваться под данные и находить более сложные зависимости. Число признаков в новой модели:
```
print(example_data_train_poly.shape)
```
Но при этом одновременно данный метод способствует более сильной способности модели к переобучению из-за быстрого роста числа признаком с увеличением степени $p$. Рассмотрим пример с $p=11$:
```
transform = PolynomialFeatures(11)
example_data_train_poly = transform.fit_transform(example_data_train)
example_data_test_poly = transform.transform(example_data_test)
optimizer = GridSearchCV(LogisticRegression(class_weight='balanced', fit_intercept=False), param_grid, cv=cv, n_jobs=-1)
optimizer.fit(example_data_train_poly, example_labels_train)
Z = optimizer.predict(transform.transform(np.c_[xx.ravel(), yy.ravel()])).reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=plt.cm.Pastel2)
plt.scatter(data_0[:,0], data_0[:,1], color='red')
plt.scatter(data_1[:,0], data_1[:,1], color='blue')
plt.title('Corrected class weights')
plt.show()
```
Количество признаков в данной модели:
```
print(example_data_train_poly.shape)
```
## Задание 5. Трансформация вещественных признаков.
1. Реализуйте по аналогии с примером преобразование вещественных признаков модели при помощи полиномиальных признаков степени 2
2. Постройте логистическую регрессию на новых данных, одновременно подобрав оптимальные гиперпараметры. Обращаем внимание, что в преобразованных признаках уже присутствует столбец, все значения которого равны 1, поэтому обучать дополнительно значение $b$ не нужно, его функцию выполняет один из весов $w$. В связи с этим во избежание линейной зависимости в датасете, в вызов класса логистической регрессии требуется передавать параметр fit_intercept=False. Для обучения используйте стратифицированные выборки, преобразованные признаки требуется заново отмасштабировать.
3. Получите AUC ROC на тесте и сравните данный результат с использованием обычных признаков.
4. Передайте полученный ответ в функцию write_answer_5.
```
def write_answer_5(auc):
with open("preprocessing_lr_answer5.txt", "w") as fout:
fout.write(str(auc))
# place your code here
(X_train_real_zeros,
X_test_real_zeros,
y_train, y_test) = train_test_split(X_real_zeros, y,
test_size=0.3,
random_state=0, stratify=y)
(X_train_cat_oh,
X_test_cat_oh) = train_test_split(X_cat_oh,
test_size=0.3,
random_state=0, stratify=y)
transform = PolynomialFeatures(2)
X_train_poly = transform.fit_transform(X_train_real_zeros)
X_test_poly = transform.transform(X_test_real_zeros)
scaler = StandardScaler()
X_train_strat = np.hstack((scaler.fit_transform(X_train_poly), X_train_cat_oh))
X_test_strat = np.hstack((scaler.transform(X_test_poly), X_test_cat_oh))
"""Обращаем внимание на параметр fit_intercept=False"""
polySearch = GridSearchCV(LogisticRegression(class_weight='balanced', fit_intercept=False), param_grid, cv=cv, n_jobs=-1)
polySearch.fit(X_train_strat, y_train)
auc_p = roc_auc_score(y_test, polySearch.predict(X_test_strat))
write_answer_5(auc_p)
auc_p
```
## Регрессия Lasso.
К логистической регрессии также можно применить L1-регуляризацию (Lasso), вместо регуляризации L2, которая будет приводить к отбору признаков. Вам предлагается применить L1-регуляцию к исходным признакам и проинтерпретировать полученные результаты (применение отбора признаков к полиномиальным так же можно успешно применять, но в нём уже будет отсутствовать компонента интерпретации, т.к. смысловое значение оригинальных признаков известно, а полиномиальных - уже может быть достаточно нетривиально). Для вызова логистической регрессии с L1-регуляризацией достаточно передать параметр penalty='l1' в инициализацию класса.
## Задание 6. Отбор признаков при помощи регрессии Lasso.
1. Обучите регрессию Lasso на стратифицированных отмасштабированных выборках
2. Получите ROC AUC регрессии, сравните его с предыдущими результатами.
3. Найдите номера вещественных признаков, которые имеют нулевые веса в итоговой модели.
4. Передайте их список функции write_answer_6.
```
def write_answer_6(features):
with open("preprocessing_lr_answer6.txt", "w") as fout:
fout.write(" ".join([str(num) for num in features]))
# place your code here
(X_train_real_zeros,
X_test_real_zeros,
y_train, y_test) = train_test_split(X_real_zeros, y,
test_size=0.3,
random_state=0, stratify=y)
(X_train_cat_oh,
X_test_cat_oh) = train_test_split(X_cat_oh,
test_size=0.3,
random_state=0, stratify=y)
scaler = StandardScaler()
X_train_strat = np.hstack((scaler.fit_transform(X_train_real_zeros), X_train_cat_oh))
X_test_strat = np.hstack((scaler.transform(X_test_real_zeros), X_test_cat_oh))
# clf = LogisticRegression(penalty='l1')
lasso = GridSearchCV(LogisticRegression(penalty='l1', class_weight='balanced'), param_grid, cv=cv, scoring = 'accuracy',n_jobs=-1)#
lasso.fit(X_train_strat, y_train)
# clf.fit(X_train_strat, y_train)
# auc_lasso = roc_auc_score(y_test, lasso.predict(X_test_strat))
clf = lasso.best_estimator_
# len(cl.coef_[0])
zero_coef = np.where(clf.coef_[0] == 0)[0]
# print X_train_real_zeros.shape
zero_coef[:3]
# len(clf.coef_[0])
write_answer_6(zero_coef[:3])
```
| github_jupyter |
tgb - 5/21/2019 - The general goal of this notebook is to develop the new Jacobian diagnostics to assess the stability of the constrained/not-constrained networks. The sub-goals are:
1) Developing a Jacobian diagnostics toolbox to normalize the full Jacobian to the right units and analyze the eigenvalues of the dynamical Jacobian
2) Calculate the Jacobian as a function of latitude in our aquaplanet simulation
3) Prepare the Jacobian for the collaboration with Noah Brenowitz
# 1) Jacobian toolbox
## 1.1) Load CBRAIN utilities and a model
```
import tensorflow as tf
#tf.enable_eager_execution()
from cbrain.imports import *
from cbrain.data_generator import *
from cbrain.cam_constants import *
from cbrain.losses import *
from cbrain.utils import limit_mem
from cbrain.layers import *
from cbrain.model_diagnostics import *
import tensorflow.math as tfm
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
import xarray as xr
import numpy as np
from cbrain.model_diagnostics import ModelDiagnostics
from numpy import linalg as LA
import matplotlib.pyplot as plt
# Otherwise tensorflow will use ALL your GPU RAM for no reason
limit_mem()
TRAINDIR = '/local/Tom.Beucler/SPCAM_PHYS/'
DATADIR = '/project/meteo/w2w/A6/S.Rasp/SP-CAM/fluxbypass_aqua/'
PREFIX = '8col009_01_'
%cd /filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM
coor = xr.open_dataset("/project/meteo/w2w/A6/S.Rasp/SP-CAM/fluxbypass_aqua/AndKua_aqua_SPCAM3.0_sp_fbp_f4.cam2.h1.0000-01-01-00000.nc",\
decode_times=False)
lat = coor.lat; lon = coor.lon;
coor.close();
config_fn = '/filer/z-sv-pool12c/t/Tom.Beucler/SPCAM/CBRAIN-CAM/pp_config/8col_rad_tbeucler_local_PostProc.yml'
data_fn = '/local/Tom.Beucler/SPCAM_PHYS/8col009_01_valid.nc'
dict_lay = {'SurRadLayer':SurRadLayer,'MassConsLayer':MassConsLayer,'EntConsLayer':EntConsLayer}
NN = {};
%cd $TRAINDIR/HDF5_DATA
# NNA0.01
path = TRAINDIR+'HDF5_DATA/JNNC.h5'
NN = load_model(path,custom_objects=dict_lay)
md = ModelDiagnostics(NN,config_fn,data_fn)
NN.summary()
```
## 1.2) Calculate Jacobian for specific soundings
tgb - 5/21/2019 - I have no idea why but the tape must be set to persistent=True and the jacobian to experimental_use_pfor=False to be able to calculate the Jacobian in eager execution mode #magic
```
# itime = 1
# X, truth = md.valid_gen[itime]
# inp = tf.convert_to_tensor(np.expand_dims(X[0,:],axis=0))
# with tf.GradientTape(persistent=True) as tape:
# tape.watch(inp)
# pred = NN(inp)
# J = tape.jacobian(pred,inp,experimental_use_pfor=False).numpy()
```
tgb - 5/21/2019 - Renormalize the Jacobian to 1/(Input units) which does not mean much
```
# JAC = J.squeeze()/md.valid_gen.input_transform.div
```
# 1.3) Create corresponding functions
tgb - 5/21/2019 - Following https://github.com/tbeucler/CBRAIN-CAM/blob/master/notebooks/tbeucler_devlog/004_Calculating_Jacobians_of_NN_and_dynamics_of_TQ.ipynb but changed the gradient loop to a single Jacobian calculation and made use of the model diagnostics object
```
def get_TQjacobian(model,inp,sample_index,md):
# model is the neural network model from inp to out
# inp is the input x generator from the generator (object = gen_obj)
# sample_index is the reference number of the sample for x
# md is the model diagnostics object
# x.shape = (#sample,#inputs) so we are evaluating the gradient of
# y(sample_index,:) with respect to x(sample_index,:)
# cf is the conversion factor calculated using CAM constants
cf = np.zeros((1, md.valid_gen.n_inputs))
for index in range (md.valid_gen.n_inputs):
if index<90: cf[0,index]=L_V;
elif index<120: cf[0,index]=C_P;
elif index<150: cf[0,index]=1;
elif index<240: cf[0,index]=L_V*DT;
elif index<270: cf[0,index]=C_P*DT;
elif index<301: cf[0,index]=1;
else: cf[0,index]=DT;
J = np.zeros((md.valid_gen.n_outputs,md.valid_gen.n_inputs))
with tf.GradientTape(persistent=True) as tape:
TFinp = tf.convert_to_tensor(np.expand_dims(inp[0,:],axis=0))
tape.watch(TFinp)
pred = NN(TFinp)
J = tape.jacobian(pred,TFinp,experimental_use_pfor=False)\
.numpy().squeeze()/(cf*md.valid_gen.input_transform.div)
JTQ = np.zeros((60,60))
for i in range (60):
for j in range(60):
if (i<30) and (j<30): JTQ[i,j] = J[i,j] # d(dq/dt)/dq
elif (i>29) and (j<30): JTQ[i,j] = J[90+(i-30),j]-\
J[120+(i-30),j]-J[150+(i-30),j] # d(dTcon/dt)/dq
elif (i<30) and (j>29): JTQ[i,j] = J[i,90+(j-30)] # d(dq/dt)/dT
elif (i>29) and (j>29): JTQ[i,j] = J[90+(i-30),90+(j-30)]-\
J[120+(i-30),90+(j-30)]-J[150+(i-30),90+(j-30)] # d(dTcon/dt)/dT
return JTQ
def get_RADjacobian(model,inp,sample_index,md):
# model is the neural network model from inp to out
# inp is the input x generator from the generator (object = gen_obj)
# sample_index is the reference number of the sample for x
# md is the model diagnostics object
# x.shape = (#sample,#inputs) so we are evaluating the gradient of
# y(sample_index,:) with respect to x(sample_index,:)
# cf is the conversion factor calculated using CAM constants
cf = np.zeros((1, md.valid_gen.n_inputs))
for index in range (md.valid_gen.n_inputs):
if index<90: cf[0,index]=L_V;
elif index<120: cf[0,index]=C_P;
elif index<150: cf[0,index]=1;
elif index<240: cf[0,index]=L_V*DT;
elif index<270: cf[0,index]=C_P*DT;
elif index<301: cf[0,index]=1;
else: cf[0,index]=DT;
J = np.zeros((md.valid_gen.n_outputs,md.valid_gen.n_inputs))
with tf.GradientTape(persistent=True) as tape:
TFinp = tf.convert_to_tensor(np.expand_dims(inp[0,:],axis=0))
tape.watch(TFinp)
pred = NN(TFinp)
J = tape.jacobian(pred,TFinp,experimental_use_pfor=False)\
.numpy().squeeze()/(cf*md.valid_gen.input_transform.div)
JRAD = np.zeros((60,60))
for i in range (60):
for j in range(60):
if (i<30) and (j<30): JRAD[i,j] = J[120+i,j]
elif (i>29) and (j<30): JRAD[i,j] = J[150+(i-30),j]
elif (i<30) and (j>29): JRAD[i,j] = J[120+i,90+(j-30)]
elif (i>29) and (j>29): JRAD[i,j] = J[150+(i-30),90+(j-30)]
return JRAD
```
tgb - 5/21/2019 - Get radiative AND convective Jacobian for an entire batch
```
def get_RADCONjacobian(model,inp,md,ind):
# model is the neural network model from inp to out
# inp is the input x generator from the generator (object = gen_obj)
# sample_index is the reference number of the sample for x
# md is the model diagnostics object
# ind is the indices over which the Jacobian is calculated
# x.shape = (#sample,#inputs) so we are evaluating the gradient of
# y(sample_index,:) with respect to x(sample_index,:)
# cf is the conversion factor calculated using CAM constants
cf = np.zeros((1, md.valid_gen.n_inputs))
for index in range (md.valid_gen.n_inputs):
if index<90: cf[0,index]=L_V;
elif index<120: cf[0,index]=C_P;
elif index<150: cf[0,index]=1;
elif index<240: cf[0,index]=L_V*DT;
elif index<270: cf[0,index]=C_P*DT;
elif index<301: cf[0,index]=1;
else: cf[0,index]=DT;
JCON = np.zeros((60,60,len(ind)))
JRAD = np.zeros((60,60,len(ind)))
J = np.zeros((md.valid_gen.n_outputs,md.valid_gen.n_inputs,len(ind)))
for count,i in enumerate(ind):
print('i=',i,'/',len(ind)-1,end="\r")
with tf.GradientTape(persistent=True) as tape:
TFinp = tf.convert_to_tensor(np.expand_dims(inp[i,:],axis=0))
tape.watch(TFinp)
pred = NN(TFinp)
J[:,:,i] = tape.jacobian(pred,TFinp,experimental_use_pfor=False)\
.numpy().squeeze()/(cf*md.valid_gen.input_transform.div)
for i in range (60):
for j in range(60):
# Convection
if (i<30) and (j<30): JCON[i,j] = J[i,j,:] # d(dq/dt)/dq
elif (i>29) and (j<30): JCON[i,j] = J[90+(i-30),j,:]-\
J[120+(i-30),j,:]-J[150+(i-30),j,:] # d(dTcon/dt)/dq
elif (i<30) and (j>29): JCON[i,j] = J[i,90+(j-30),:] # d(dq/dt)/dT
elif (i>29) and (j>29): JCON[i,j] = J[90+(i-30),90+(j-30),:]-\
J[120+(i-30),90+(j-30),:]-J[150+(i-30),90+(j-30),:] # d(dTcon/dt)/dT
# Radiation
if (i<30) and (j<30): JRAD[i,j,:] = J[120+i,j,:]
elif (i>29) and (j<30): JRAD[i,j,:] = J[150+(i-30),j,:]
elif (i<30) and (j>29): JRAD[i,j,:] = J[120+i,90+(j-30),:]
elif (i>29) and (j>29): JRAD[i,j,:] = J[150+(i-30),90+(j-30),:]
return JCON,JRAD
from tensorflow.python.ops.parallel_for.gradients import batch_jacobian
def get_jacobian(x, model):
sess = tf.keras.backend.get_session()
jac = jacobian(model.output, model.input)
J = sess.run(jac, feed_dict={model.input: x.astype(np.float32)[None]})
return J.squeeze()
def get_batch_jacobian(x, model):
sess = tf.keras.backend.get_session()
jac = batch_jacobian(model.output, model.input)
J = sess.run(jac, feed_dict={model.input: x.astype(np.float32)})
return J.squeeze()
```
tgb - 6/4/2019 - Corrected it to the right units
```
def get_RADCONjacobian(model,md,itime):
# model is the neural network model from inp to out
# inp is the input x generator from the generator (object = gen_obj)
# sample_index is the reference number of the sample for x
# md is the model diagnostics object
# ind is the indices over which the Jacobian is calculated
# x.shape = (#sample,#inputs) so we are evaluating the gradient of
# y(sample_index,:) with respect to x(sample_index,:)
# cf is the conversion factor calculated using CAM constants
X, truth = md.valid_gen[itime]
cf = np.zeros((1, md.valid_gen.n_inputs))
for index in range (md.valid_gen.n_inputs):
if index<90: cf[0,index]=L_V;
elif index<120: cf[0,index]=C_P;
elif index<150: cf[0,index]=1;
elif index<240: cf[0,index]=L_V*DT;
elif index<270: cf[0,index]=C_P*DT;
elif index<301: cf[0,index]=1;
else: cf[0,index]=DT;
cf_oup = np.zeros((1,md.valid_gen.n_outputs))
for index in range (md.valid_gen.n_outputs):
if index<90: cf_oup[0,index]=L_V;
elif index<210: cf_oup[0,index]=C_P;
else: cf_oup[0,index]=1;
JCON = np.zeros((60,60,8192))
JRAD = np.zeros((60,60,8192))
# J = np.zeros((md.valid_gen.n_outputs,md.valid_gen.n_inputs,len(ind)))
# for count,i in enumerate(ind):
# print('i=',i,'/',len(ind)-1,end="\r")
# with tf.GradientTape(persistent=True) as tape:
# TFinp = tf.convert_to_tensor(np.expand_dims(inp[i,:],axis=0))
# tape.watch(TFinp)
# pred = NN(TFinp)
# J[:,:,i] = tape.jacobian(pred,TFinp,experimental_use_pfor=False)\
# .numpy().squeeze()/(cf*md.valid_gen.input_transform.div)
for ind in range(32):
print('ind=',ind,'/',32,end="\r")
sample = np.arange(256*ind,256*(ind+1))
Jtmp = get_batch_jacobian(X[sample,:], model)*\
np.transpose(cf_oup/md.valid_gen.output_transform.scale)\
/(cf*md.valid_gen.input_transform.div)
if ind==0: J = Jtmp
else: J = np.concatenate((J,Jtmp),axis=0)
print(J.shape)
for i in range (60):
for j in range(60):
# Convection
if (i<30) and (j<30): JCON[i,j] = J[:,i,j] # d(dq/dt)/dq
elif (i>29) and (j<30): JCON[i,j] = J[:,90+(i-30),j]-\
J[:,120+(i-30),j]-J[:,150+(i-30),j] # d(dTcon/dt)/dq
elif (i<30) and (j>29): JCON[i,j] = J[:,i,90+(j-30)] # d(dq/dt)/dT
elif (i>29) and (j>29): JCON[i,j] = J[:,90+(i-30),90+(j-30)]-\
J[:,120+(i-30),90+(j-30)]-J[:,150+(i-30),90+(j-30)] # d(dTcon/dt)/dT
# Radiation
if (i<30) and (j<30): JRAD[i,j,:] = J[:,120+i,j]
elif (i>29) and (j<30): JRAD[i,j,:] = J[:,150+(i-30),j]
elif (i<30) and (j>29): JRAD[i,j,:] = J[:,120+i,90+(j-30)]
elif (i>29) and (j>29): JRAD[i,j,:] = J[:,150+(i-30),90+(j-30)]
return JCON,JRAD
X.shape
JRAD2 = np.reshape(JRAD, (60,60,64,128))
JRAD2.shape
JRAD3 = JRAD2.mean(axis=3)
JRAD3.shape
np.mean(np.reshape(JRAD,(60,60,64,128)),axis=3)
import h5py
JCONa = np.zeros((md.nlat,60,60))
JRADa = np.copy(JCONa)
for itime in range(100):
print('itime=',itime)
X, truth = md.valid_gen[itime]
Xgeo = X.values.reshape(md.nlat, md.nlon, 304)
for ilat in range(md.nlat):
print('ilat=',ilat,'/',str(md.nlat-1),end="\r")
Xlat = Xgeo[ilat,:,:]
JCON,JRAD = get_RADCONjacobian(NN,Xlat,md,np.arange(0,md.nlon))
JCONa[ilat,:,:] = JCON.mean(axis=2)/(itime+1)+\
itime/(itime+1)*JCONa[ilat,:,:]
JRADa[ilat,:,:] = JRAD.mean(axis=2)/(itime+1)+\
itime/(itime+1)*JRADa[ilat,:,:]
print('itime=',itime,'Saving the arrays in HDF5 format')
with h5py.File('HDF5_DATA/014_JCONa.h5', 'w') as hf:
hf.create_dataset("name-of-dataset", data=JCONa)
with h5py.File('HDF5_DATA/014_JRADa.h5', 'w') as hf:
hf.create_dataset("name-of-dataset", data=JRADa)
```
tgb - 5/21/2019 - Save in HDF5 format using https://stackoverflow.com/questions/20928136/input-and-output-numpy-arrays-to-h5py
```
# import h5py
# with h5py.File('HDF5_DATA/014_JCON.h5', 'w') as hf:
# hf.create_dataset("name-of-dataset", data=JCON)
# with h5py.File('HDF5_DATA/014_JRAD.h5', 'w') as hf:
# hf.create_dataset("name-of-dataset", data=JRAD)
# JRAD = np.swapaxes(JRAD,1,2)
# JRAD.shape
# JRAD = np.swapaxes(JRAD,0,1)
# JRAD.shape
# with h5py.File('HDF5_DATA/014_JCONa.h5', 'r') as hf:
# JCON = hf['name-of-dataset'][:]
# with h5py.File('HDF5_DATA/014_JRADa.h5', 'r') as hf:
# JRAD = hf['name-of-dataset'][:]
```
tgb - 5/22/2019 - First calculate parameters useful for the particular graph, then graph for each latitude
```
minlev = 200; # Minimum level [hPa]
PS = 1e5; P0 = 1e5;
P = P0*hyai+PS*hybi; # Total pressure [Pa]
minind = np.argmin(abs(P-minlev*1e2));
IND = np.concatenate((np.arange(minind,30),np.arange(30+minind,60)),axis=0)
coor.lat[ilat]
coor.TS[ilat]
ilat = 2
lw = 2.5
siz = 200
plt.figure(num=None, figsize=(20, 2), dpi=80, facecolor='w', edgecolor='k')
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
plt.plot(coor.lat,coor.TS.mean(axis=(0,2)),color='b',linewidth=lw)
plt.xticks(fontsize=20); plt.yticks(fontsize=20)
plt.scatter(coor.lat[ilat],coor.TS.mean(axis=(0,2))[ilat],s=siz,color='b')
plt.xlim((np.min(coor.lat),np.max(coor.lat)));
plt.xlabel(r'$\mathrm{Latitude\ \left(^{\circ}\right)}$', fontsize = 20)
#ax.xaxis.labelpad = 20
plt.ylabel(r'$\mathrm{SST\ \left(K\right)}$', fontsize = 20)
JCON.shape
JCON[ilat,:,:,:].shape
J = JCON[ilat,:,:,:].mean(axis=0)
print(J.shape)
print(J)
def Jacobian_figurelat(Jinp,coor,ilat=32,plot_option='CON',lw=2.5,siz=200,vmin=-3600*0.003, vmax=3600*0.003):
if len(Jinp.shape)>=4: J = Jinp[ilat,:,:,:].mean(axis=0)
#else: J = Jinp[ilat,:,:] tgb - 6/4/2019 - changed shape of Jacobian
else: J = Jinp[:,:,ilat]
# minlev = 200; # Minimum level [hPa]
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
plt.figure(num=None, figsize=(20, 13.5), dpi=80, facecolor='w', edgecolor='k')
fig, ax = plt.subplots(figsize=(10,10))
# Extract the Jacobian from level minlev to the surface (index 30 here)
# PS = 1e5; P0 = 1e5;
# P = P0*hyai+PS*hybi; # Total pressure [Pa]
# minind = np.argmin(abs(P-minlev*1e2));
# IND = np.concatenate((np.arange(minind,30),np.arange(30+minind,60)),axis=0)
J = J[IND,:]; J = J[:,IND];
cax = ax.matshow(24*3600*J, vmin=vmin,vmax=vmax, cmap='bwr')
x = np.linspace(0.,60.,100);
plt.plot(x,J.shape[0]/2*x**0, color='k')
plt.plot(J.shape[0]/2*x**0,x, color='k')
plt.xlim((0,J.shape[0])); plt.ylim((J.shape[0],0))
cbar = fig.colorbar(cax, pad = 0.1)
cbar.ax.tick_params(labelsize=20)
cbar.set_label(r'$\mathrm{Growth\ rate\ \left(1/day\right)}$', rotation=90, fontsize = 20)
plt.xticks(fontsize=20); plt.yticks(fontsize=20)
ax.xaxis.set_label_position('top')
X = plt.xlabel(r'Input level', fontsize = 20)
ax.xaxis.labelpad = 20
Y = plt.ylabel(r'Output level', fontsize = 20)
ax.yaxis.labelpad = 20
# fig.canvas.draw()
labelx = [item.get_text() for item in ax.get_xticklabels()]
# labelx[0] = '$\mathrm{QV_{0hPa}}$'
# labelx[1] = '$\mathrm{QV_{130hPa}}$'
# labelx[2] = '$\mathrm{QV_{650hPa}}$'
# labelx[3] = '$\mathrm{T_{0hPa}}$'
# labelx[4] = '$\mathrm{T_{130hPa}}$'
# labelx[5] = '$\mathrm{T_{650hPa}}$'
# labelx[6] = '$\mathrm{T_{1000hPa}}$'
labelx[0] = '$\mathrm{Q_{200hPa}}$'
labelx[1] = '$\mathrm{Q_{500hPa}}$'
labelx[2] = '$\mathrm{Q_{850hPa}}$'
labelx[3] = '$\mathrm{Q_{967hPa}}$'
labelx[4] = '$\mathrm{T_{350hPa}}$'
labelx[5] = '$\mathrm{T_{730hPa}}$'
labelx[6] = '$\mathrm{T_{925hPa}}$'
labely = [item.get_text() for item in ax.get_yticklabels()]
if plot_option=='RAD':
# labely[0] = '$\mathrm{LW_{0hPa}}$'
# labely[1] = '$\mathrm{LW_{130hPa}}$'
# labely[2] = '$\mathrm{LW_{650hPa}}$'
# labely[3] = '$\mathrm{SW_{0hPa}}$'
# labely[4] = '$\mathrm{SW_{130hPa}}$'
# labely[5] = '$\mathrm{SW_{650hPa}}$'
# labely[6] = '$\mathrm{SW_{1000hPa}}$'
labely[0] = '$\mathrm{LW_{200hPa}}$'
labely[1] = '$\mathrm{LW_{500hPa}}$'
labely[2] = '$\mathrm{LW_{850hPa}}$'
labely[3] = '$\mathrm{LW_{967hPa}}$'
labely[4] = '$\mathrm{SW_{350hPa}}$'
labely[5] = '$\mathrm{SW_{730hPa}}$'
labely[6] = '$\mathrm{SW_{925hPa}}$'
elif plot_option=='CON':
# labely[0] = '$\mathrm{\dot{QV}_{0hPa}}$'
# labely[1] = '$\mathrm{\dot{QV}_{130hPa}}$'
# labely[2] = '$\mathrm{\dot{QV}_{650hPa}}$'
# labely[3] = '$\mathrm{\dot{T}_{0hPa}}$'
# labely[4] = '$\mathrm{\dot{T}_{130hPa}}$'
# labely[5] = '$\mathrm{\dot{T}_{650hPa}}$'
# labely[6] = '$\mathrm{\dot{T}_{1000hPa}}$'
labely[0] = '$\mathrm{\dot{Q}_{200hPa}}$'
labely[1] = '$\mathrm{\dot{Q}_{500hPa}}$'
labely[2] = '$\mathrm{\dot{Q}_{850hPa}}$'
labely[3] = '$\mathrm{\dot{Q}_{967hPa}}$'
labely[4] = '$\mathrm{\dot{T}_{350hPa}}$'
labely[5] = '$\mathrm{\dot{T}_{730hPa}}$'
labely[6] = '$\mathrm{\dot{T}_{925hPa}}$'
ax.set_xticklabels(labelx)
ax.set_yticklabels(labely)
SSTplt = plt.axes([0.1, 0.05, 0.65, .15], facecolor='w')
#plt.figure(num=None, figsize=(20, 2), dpi=80, facecolor='w', edgecolor='k')
#plt.rc('text', usetex=True)
#plt.rc('font', family='serif')
SSTplt.plot(coor.lat,coor.TS.mean(axis=(0,2)),color='b',linewidth=lw)
SSTplt.tick_params(labelsize=15)
SSTplt.scatter(coor.lat[ilat],coor.TS.mean(axis=(0,2))[ilat],s=siz,color='b')
SSTplt.set_xlim((np.min(coor.lat),np.max(coor.lat)));
SSTplt.set_ylim((0.995*np.min(coor.TS.mean(axis=(0,2))),1.005*np.max(coor.TS.mean(axis=(0,2)))));
SSTplt.set_xlabel(r'$\mathrm{Latitude\ \left(^{\circ}\right)}$', fontsize = 15)
#ax.xaxis.labelpad = 20
SSTplt.set_ylabel(r'$\mathrm{SST\ \left(K\right)}$', fontsize = 15)
return fig,ax
Jacobian_figurelat(np.mean(np.reshape(JRAD,(60,60,64,128)),axis=3),\
coor,ilat=35,plot_option='RAD',lw=2.5,siz=200,vmin=-0.2, vmax=0.2)
IND
Jacobian_figurelat(JRAD,coor,ilat=30,plot_option='RAD',lw=2.5,siz=200,vmin=-3600*0.003, vmax=3600*0.003)
for ilat in range(len(coor.lat)):
print('ilat=',ilat,'/',str(len(coor.lat)-1),end="\r")
Jacobian_figurelat(JCON,coor,ilat=ilat,plot_option='CON',lw=2.5,siz=200,vmin=-3600*0.003, vmax=3600*0.003)
plt.savefig('PNG_DATA/'+'CON'+str(ilat))
plt.close('all')
```
tgb - 6/4/2019 - NE runs
```
X, truth = md.valid_gen[itime]
np.floor(8192/32)
ind = 0
X[256*ind:256*(ind+1),:]
Nt = 10
JCON = np.zeros((60,60,8192))
JRAD = np.zeros((60,60,8192))
for it in range(Nt):
print('itime=',it,'/',Nt,end="\r")
JCONa,JRADa = get_RADCONjacobian(NN,md,it)
JCON = JCONa/(it+1)+it/(it+1)*JCON
JRAD = JRADa/(it+1)+it/(it+1)*JRAD
for ilat in range(len(coor.lat)):
print('ilat=',ilat,'/',str(len(coor.lat)-1),end="\r")
Jacobian_figurelat(np.mean(np.reshape(JCON,(60,60,64,128)),axis=3),\
coor,ilat=ilat,plot_option='CON',lw=2.5,siz=200,vmin=-0.2, vmax=0.2)
plt.savefig('PNG_DATA/'+'CON'+str(ilat))
Jacobian_figurelat(np.mean(np.reshape(JRAD,(60,60,64,128)),axis=3),\
coor,ilat=ilat,plot_option='RAD',lw=2.5,siz=200,vmin=-0.2, vmax=0.2)
plt.savefig('PNG_DATA/'+'RAD'+str(ilat))
plt.close('all')
def Jacobian_figure(J,plot_option):
# minlev = 200; # Minimum level [hPa]
plt.rc('text', usetex=True)
plt.rc('font', family='serif')
plt.figure(num=None, figsize=(20, 13.5), dpi=80, facecolor='w', edgecolor='k')
fig, ax = plt.subplots(figsize=(10,10))
# Extract the Jacobian from level minlev to the surface (index 30 here)
# PS = 1e5; P0 = 1e5;
# P = P0*hyai+PS*hybi; # Total pressure [Pa]
# minind = np.argmin(abs(P-minlev*1e2));
# IND = np.concatenate((np.arange(minind,30),np.arange(30+minind,60)),axis=0)
J = J[IND,:]; J = J[:,IND];
cax = ax.matshow(3600*J, vmin=-3600*0.003, vmax=3600*0.003, cmap='bwr')
x = np.linspace(0.,60.,100);
plt.plot(x,J.shape[0]/2*x**0, color='k')
plt.plot(J.shape[0]/2*x**0,x, color='k')
plt.xlim((0,J.shape[0])); plt.ylim((J.shape[0],0))
cbar = fig.colorbar(cax, pad = 0.1)
cbar.ax.tick_params(labelsize=20)
cbar.set_label(r'$\mathrm{Growth\ rate\ \left(1/hour\right)}$', rotation=90, fontsize = 20)
plt.xticks(fontsize=20); plt.yticks(fontsize=20)
ax.xaxis.set_label_position('top')
X = plt.xlabel(r'Input level', fontsize = 20)
ax.xaxis.labelpad = 20
Y = plt.ylabel(r'Output level', fontsize = 20)
ax.yaxis.labelpad = 20
# fig.canvas.draw()
labelx = [item.get_text() for item in ax.get_xticklabels()]
# labelx[0] = '$\mathrm{QV_{0hPa}}$'
# labelx[1] = '$\mathrm{QV_{130hPa}}$'
# labelx[2] = '$\mathrm{QV_{650hPa}}$'
# labelx[3] = '$\mathrm{T_{0hPa}}$'
# labelx[4] = '$\mathrm{T_{130hPa}}$'
# labelx[5] = '$\mathrm{T_{650hPa}}$'
# labelx[6] = '$\mathrm{T_{1000hPa}}$'
labelx[0] = '$\mathrm{Q_{200hPa}}$'
labelx[1] = '$\mathrm{Q_{500hPa}}$'
labelx[2] = '$\mathrm{Q_{850hPa}}$'
labelx[3] = '$\mathrm{Q_{967hPa}}$'
labelx[4] = '$\mathrm{T_{350hPa}}$'
labelx[5] = '$\mathrm{T_{730hPa}}$'
labelx[6] = '$\mathrm{T_{925hPa}}$'
labely = [item.get_text() for item in ax.get_yticklabels()]
if plot_option=='RAD':
# labely[0] = '$\mathrm{LW_{0hPa}}$'
# labely[1] = '$\mathrm{LW_{130hPa}}$'
# labely[2] = '$\mathrm{LW_{650hPa}}$'
# labely[3] = '$\mathrm{SW_{0hPa}}$'
# labely[4] = '$\mathrm{SW_{130hPa}}$'
# labely[5] = '$\mathrm{SW_{650hPa}}$'
# labely[6] = '$\mathrm{SW_{1000hPa}}$'
labely[0] = '$\mathrm{LW_{200hPa}}$'
labely[1] = '$\mathrm{LW_{500hPa}}$'
labely[2] = '$\mathrm{LW_{850hPa}}$'
labely[3] = '$\mathrm{LW_{967hPa}}$'
labely[4] = '$\mathrm{SW_{350hPa}}$'
labely[5] = '$\mathrm{SW_{730hPa}}$'
labely[6] = '$\mathrm{SW_{925hPa}}$'
elif plot_option=='CON':
# labely[0] = '$\mathrm{\dot{QV}_{0hPa}}$'
# labely[1] = '$\mathrm{\dot{QV}_{130hPa}}$'
# labely[2] = '$\mathrm{\dot{QV}_{650hPa}}$'
# labely[3] = '$\mathrm{\dot{T}_{0hPa}}$'
# labely[4] = '$\mathrm{\dot{T}_{130hPa}}$'
# labely[5] = '$\mathrm{\dot{T}_{650hPa}}$'
# labely[6] = '$\mathrm{\dot{T}_{1000hPa}}$'
labely[0] = '$\mathrm{\dot{Q}_{200hPa}}$'
labely[1] = '$\mathrm{\dot{Q}_{500hPa}}$'
labely[2] = '$\mathrm{\dot{Q}_{850hPa}}$'
labely[3] = '$\mathrm{\dot{Q}_{967hPa}}$'
labely[4] = '$\mathrm{\dot{T}_{350hPa}}$'
labely[5] = '$\mathrm{\dot{T}_{730hPa}}$'
labely[6] = '$\mathrm{\dot{T}_{925hPa}}$'
ax.set_xticklabels(labelx)
ax.set_yticklabels(labely)
return fig,ax
```
| github_jupyter |
**Chapter 11 – Training Deep Neural Networks**
_This notebook contains all the sample code and solutions to the exercises in chapter 11._
<table align="left">
<td>
<a href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/11_training_deep_neural_networks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
</td>
<td>
<a target="_blank" href="https://kaggle.com/kernels/welcome?src=https://github.com/ageron/handson-ml2/blob/master/11_training_deep_neural_networks.ipynb"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
</td>
</table>
# Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20 and TensorFlow ≥2.0.
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
# TensorFlow ≥2.0 is required
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
%load_ext tensorboard
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
# Vanishing/Exploding Gradients Problem
```
def logit(z):
return 1 / (1 + np.exp(-z))
z = np.linspace(-5, 5, 200)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [1, 1], 'k--')
plt.plot([0, 0], [-0.2, 1.2], 'k-')
plt.plot([-5, 5], [-3/4, 7/4], 'g--')
plt.plot(z, logit(z), "b-", linewidth=2)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Saturating', xytext=(3.5, 0.7), xy=(5, 1), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Saturating', xytext=(-3.5, 0.3), xy=(-5, 0), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Linear', xytext=(2, 0.2), xy=(0, 0.5), arrowprops=props, fontsize=14, ha="center")
plt.grid(True)
plt.title("Sigmoid activation function", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("sigmoid_saturation_plot")
plt.show()
```
## Xavier and He Initialization
```
[name for name in dir(keras.initializers) if not name.startswith("_")]
keras.layers.Dense(10, activation="relu", kernel_initializer="he_normal")
init = keras.initializers.VarianceScaling(scale=2., mode='fan_avg',
distribution='uniform')
keras.layers.Dense(10, activation="relu", kernel_initializer=init)
```
## Nonsaturating Activation Functions
### Leaky ReLU
```
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
plt.plot(z, leaky_relu(z, 0.05), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([0, 0], [-0.5, 4.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Leak', xytext=(-3.5, 0.5), xy=(-5, -0.2), arrowprops=props, fontsize=14, ha="center")
plt.title("Leaky ReLU activation function", fontsize=14)
plt.axis([-5, 5, -0.5, 4.2])
save_fig("leaky_relu_plot")
plt.show()
[m for m in dir(keras.activations) if not m.startswith("_")]
[m for m in dir(keras.layers) if "relu" in m.lower()]
```
Let's train a neural network on Fashion MNIST using the Leaky ReLU:
```
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.fashion_mnist.load_data()
X_train_full = X_train_full / 255.0
X_test = X_test / 255.0
X_valid, X_train = X_train_full[:5000], X_train_full[5000:]
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(100, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
Now let's try PReLU:
```
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, kernel_initializer="he_normal"),
keras.layers.PReLU(),
keras.layers.Dense(100, kernel_initializer="he_normal"),
keras.layers.PReLU(),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
### ELU
```
def elu(z, alpha=1):
return np.where(z < 0, alpha * (np.exp(z) - 1), z)
plt.plot(z, elu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1, -1], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"ELU activation function ($\alpha=1$)", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("elu_plot")
plt.show()
```
Implementing ELU in TensorFlow is trivial, just specify the activation function when building each layer:
```
keras.layers.Dense(10, activation="elu")
```
### SELU
This activation function was proposed in this [great paper](https://arxiv.org/pdf/1706.02515.pdf) by Günter Klambauer, Thomas Unterthiner and Andreas Mayr, published in June 2017. During training, a neural network composed exclusively of a stack of dense layers using the SELU activation function and LeCun initialization will self-normalize: the output of each layer will tend to preserve the same mean and variance during training, which solves the vanishing/exploding gradients problem. As a result, this activation function outperforms the other activation functions very significantly for such neural nets, so you should really try it out. Unfortunately, the self-normalizing property of the SELU activation function is easily broken: you cannot use ℓ<sub>1</sub> or ℓ<sub>2</sub> regularization, regular dropout, max-norm, skip connections or other non-sequential topologies (so recurrent neural networks won't self-normalize). However, in practice it works quite well with sequential CNNs. If you break self-normalization, SELU will not necessarily outperform other activation functions.
```
from scipy.special import erfc
# alpha and scale to self normalize with mean 0 and standard deviation 1
# (see equation 14 in the paper):
alpha_0_1 = -np.sqrt(2 / np.pi) / (erfc(1/np.sqrt(2)) * np.exp(1/2) - 1)
scale_0_1 = (1 - erfc(1 / np.sqrt(2)) * np.sqrt(np.e)) * np.sqrt(2 * np.pi) * (2 * erfc(np.sqrt(2))*np.e**2 + np.pi*erfc(1/np.sqrt(2))**2*np.e - 2*(2+np.pi)*erfc(1/np.sqrt(2))*np.sqrt(np.e)+np.pi+2)**(-1/2)
def selu(z, scale=scale_0_1, alpha=alpha_0_1):
return scale * elu(z, alpha)
plt.plot(z, selu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1.758, -1.758], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title("SELU activation function", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("selu_plot")
plt.show()
```
By default, the SELU hyperparameters (`scale` and `alpha`) are tuned in such a way that the mean output of each neuron remains close to 0, and the standard deviation remains close to 1 (assuming the inputs are standardized with mean 0 and standard deviation 1 too). Using this activation function, even a 1,000 layer deep neural network preserves roughly mean 0 and standard deviation 1 across all layers, avoiding the exploding/vanishing gradients problem:
```
np.random.seed(42)
Z = np.random.normal(size=(500, 100)) # standardized inputs
for layer in range(1000):
W = np.random.normal(size=(100, 100), scale=np.sqrt(1 / 100)) # LeCun initialization
Z = selu(np.dot(Z, W))
means = np.mean(Z, axis=0).mean()
stds = np.std(Z, axis=0).mean()
if layer % 100 == 0:
print("Layer {}: mean {:.2f}, std deviation {:.2f}".format(layer, means, stds))
```
Using SELU is easy:
```
keras.layers.Dense(10, activation="selu",
kernel_initializer="lecun_normal")
```
Let's create a neural net for Fashion MNIST with 100 hidden layers, using the SELU activation function:
```
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="selu",
kernel_initializer="lecun_normal"))
for layer in range(99):
model.add(keras.layers.Dense(100, activation="selu",
kernel_initializer="lecun_normal"))
model.add(keras.layers.Dense(10, activation="softmax"))
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
```
Now let's train it. Do not forget to scale the inputs to mean 0 and standard deviation 1:
```
pixel_means = X_train.mean(axis=0, keepdims=True)
pixel_stds = X_train.std(axis=0, keepdims=True)
X_train_scaled = (X_train - pixel_means) / pixel_stds
X_valid_scaled = (X_valid - pixel_means) / pixel_stds
X_test_scaled = (X_test - pixel_means) / pixel_stds
history = model.fit(X_train_scaled, y_train, epochs=5,
validation_data=(X_valid_scaled, y_valid))
```
Now look at what happens if we try to use the ReLU activation function instead:
```
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="relu", kernel_initializer="he_normal"))
for layer in range(99):
model.add(keras.layers.Dense(100, activation="relu", kernel_initializer="he_normal"))
model.add(keras.layers.Dense(10, activation="softmax"))
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model.fit(X_train_scaled, y_train, epochs=5,
validation_data=(X_valid_scaled, y_valid))
```
Not great at all, we suffered from the vanishing/exploding gradients problem.
# Batch Normalization
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(100, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(10, activation="softmax")
])
model.summary()
bn1 = model.layers[1]
[(var.name, var.trainable) for var in bn1.variables]
#bn1.updates #deprecated
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
Sometimes applying BN before the activation function works better (there's a debate on this topic). Moreover, the layer before a `BatchNormalization` layer does not need to have bias terms, since the `BatchNormalization` layer some as well, it would be a waste of parameters, so you can set `use_bias=False` when creating those layers:
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(100, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
## Gradient Clipping
All Keras optimizers accept `clipnorm` or `clipvalue` arguments:
```
optimizer = keras.optimizers.SGD(clipvalue=1.0)
optimizer = keras.optimizers.SGD(clipnorm=1.0)
```
## Reusing Pretrained Layers
### Reusing a Keras model
Let's split the fashion MNIST training set in two:
* `X_train_A`: all images of all items except for sandals and shirts (classes 5 and 6).
* `X_train_B`: a much smaller training set of just the first 200 images of sandals or shirts.
The validation set and the test set are also split this way, but without restricting the number of images.
We will train a model on set A (classification task with 8 classes), and try to reuse it to tackle set B (binary classification). We hope to transfer a little bit of knowledge from task A to task B, since classes in set A (sneakers, ankle boots, coats, t-shirts, etc.) are somewhat similar to classes in set B (sandals and shirts). However, since we are using `Dense` layers, only patterns that occur at the same location can be reused (in contrast, convolutional layers will transfer much better, since learned patterns can be detected anywhere on the image, as we will see in the CNN chapter).
```
def split_dataset(X, y):
y_5_or_6 = (y == 5) | (y == 6) # sandals or shirts
y_A = y[~y_5_or_6]
y_A[y_A > 6] -= 2 # class indices 7, 8, 9 should be moved to 5, 6, 7
y_B = (y[y_5_or_6] == 6).astype(np.float32) # binary classification task: is it a shirt (class 6)?
return ((X[~y_5_or_6], y_A),
(X[y_5_or_6], y_B))
(X_train_A, y_train_A), (X_train_B, y_train_B) = split_dataset(X_train, y_train)
(X_valid_A, y_valid_A), (X_valid_B, y_valid_B) = split_dataset(X_valid, y_valid)
(X_test_A, y_test_A), (X_test_B, y_test_B) = split_dataset(X_test, y_test)
X_train_B = X_train_B[:200]
y_train_B = y_train_B[:200]
X_train_A.shape
X_train_B.shape
y_train_A[:30]
y_train_B[:30]
tf.random.set_seed(42)
np.random.seed(42)
model_A = keras.models.Sequential()
model_A.add(keras.layers.Flatten(input_shape=[28, 28]))
for n_hidden in (300, 100, 50, 50, 50):
model_A.add(keras.layers.Dense(n_hidden, activation="selu"))
model_A.add(keras.layers.Dense(8, activation="softmax"))
model_A.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model_A.fit(X_train_A, y_train_A, epochs=20,
validation_data=(X_valid_A, y_valid_A))
model_A.save("my_model_A.h5")
model_B = keras.models.Sequential()
model_B.add(keras.layers.Flatten(input_shape=[28, 28]))
for n_hidden in (300, 100, 50, 50, 50):
model_B.add(keras.layers.Dense(n_hidden, activation="selu"))
model_B.add(keras.layers.Dense(1, activation="sigmoid"))
model_B.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model_B.fit(X_train_B, y_train_B, epochs=20,
validation_data=(X_valid_B, y_valid_B))
model_B.summary()
model_A = keras.models.load_model("my_model_A.h5")
model_B_on_A = keras.models.Sequential(model_A.layers[:-1])
model_B_on_A.add(keras.layers.Dense(1, activation="sigmoid"))
```
Note that `model_B_on_A` and `model_A` actually share layers now, so when we train one, it will update both models. If we want to avoid that, we need to build `model_B_on_A` on top of a *clone* of `model_A`:
```
model_A_clone = keras.models.clone_model(model_A)
model_A_clone.set_weights(model_A.get_weights())
model_B_on_A = keras.models.Sequential(model_A_clone.layers[:-1])
model_B_on_A.add(keras.layers.Dense(1, activation="sigmoid"))
for layer in model_B_on_A.layers[:-1]:
layer.trainable = False
model_B_on_A.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=4,
validation_data=(X_valid_B, y_valid_B))
for layer in model_B_on_A.layers[:-1]:
layer.trainable = True
model_B_on_A.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=16,
validation_data=(X_valid_B, y_valid_B))
```
So, what's the final verdict?
```
model_B.evaluate(X_test_B, y_test_B)
model_B_on_A.evaluate(X_test_B, y_test_B)
```
Great! We got quite a bit of transfer: the error rate dropped by a factor of 4.9!
```
(100 - 97.05) / (100 - 99.40)
```
# Faster Optimizers
## Momentum optimization
```
optimizer = keras.optimizers.SGD(learning_rate=0.001, momentum=0.9)
```
## Nesterov Accelerated Gradient
```
optimizer = keras.optimizers.SGD(learning_rate=0.001, momentum=0.9, nesterov=True)
```
## AdaGrad
```
optimizer = keras.optimizers.Adagrad(learning_rate=0.001)
```
## RMSProp
```
optimizer = keras.optimizers.RMSprop(learning_rate=0.001, rho=0.9)
```
## Adam Optimization
```
optimizer = keras.optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999)
```
## Adamax Optimization
```
optimizer = keras.optimizers.Adamax(learning_rate=0.001, beta_1=0.9, beta_2=0.999)
```
## Nadam Optimization
```
optimizer = keras.optimizers.Nadam(learning_rate=0.001, beta_1=0.9, beta_2=0.999)
```
## Learning Rate Scheduling
### Power Scheduling
```lr = lr0 / (1 + steps / s)**c```
* Keras uses `c=1` and `s = 1 / decay`
```
optimizer = keras.optimizers.SGD(learning_rate=0.01, decay=1e-4)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
import math
learning_rate = 0.01
decay = 1e-4
batch_size = 32
n_steps_per_epoch = math.ceil(len(X_train) / batch_size)
epochs = np.arange(n_epochs)
lrs = learning_rate / (1 + decay * epochs * n_steps_per_epoch)
plt.plot(epochs, lrs, "o-")
plt.axis([0, n_epochs - 1, 0, 0.01])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Power Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
### Exponential Scheduling
```lr = lr0 * 0.1**(epoch / s)```
```
def exponential_decay_fn(epoch):
return 0.01 * 0.1**(epoch / 20)
def exponential_decay(lr0, s):
def exponential_decay_fn(epoch):
return lr0 * 0.1**(epoch / s)
return exponential_decay_fn
exponential_decay_fn = exponential_decay(lr0=0.01, s=20)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 25
lr_scheduler = keras.callbacks.LearningRateScheduler(exponential_decay_fn)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, history.history["lr"], "o-")
plt.axis([0, n_epochs - 1, 0, 0.011])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Exponential Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
The schedule function can take the current learning rate as a second argument:
```
def exponential_decay_fn(epoch, lr):
return lr * 0.1**(1 / 20)
```
If you want to update the learning rate at each iteration rather than at each epoch, you must write your own callback class:
```
K = keras.backend
class ExponentialDecay(keras.callbacks.Callback):
def __init__(self, s=40000):
super().__init__()
self.s = s
def on_batch_begin(self, batch, logs=None):
# Note: the `batch` argument is reset at each epoch
lr = K.get_value(self.model.optimizer.learning_rate)
K.set_value(self.model.optimizer.learning_rate, lr * 0.1**(1 / s))
def on_epoch_end(self, epoch, logs=None):
logs = logs or {}
logs['lr'] = K.get_value(self.model.optimizer.learning_rate)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
lr0 = 0.01
optimizer = keras.optimizers.Nadam(learning_rate=lr0)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
s = 20 * len(X_train) // 32 # number of steps in 20 epochs (batch size = 32)
exp_decay = ExponentialDecay(s)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[exp_decay])
n_steps = n_epochs * len(X_train) // 32
steps = np.arange(n_steps)
lrs = lr0 * 0.1**(steps / s)
plt.plot(steps, lrs, "-", linewidth=2)
plt.axis([0, n_steps - 1, 0, lr0 * 1.1])
plt.xlabel("Batch")
plt.ylabel("Learning Rate")
plt.title("Exponential Scheduling (per batch)", fontsize=14)
plt.grid(True)
plt.show()
```
### Piecewise Constant Scheduling
```
def piecewise_constant_fn(epoch):
if epoch < 5:
return 0.01
elif epoch < 15:
return 0.005
else:
return 0.001
def piecewise_constant(boundaries, values):
boundaries = np.array([0] + boundaries)
values = np.array(values)
def piecewise_constant_fn(epoch):
return values[np.argmax(boundaries > epoch) - 1]
return piecewise_constant_fn
piecewise_constant_fn = piecewise_constant([5, 15], [0.01, 0.005, 0.001])
lr_scheduler = keras.callbacks.LearningRateScheduler(piecewise_constant_fn)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, [piecewise_constant_fn(epoch) for epoch in history.epoch], "o-")
plt.axis([0, n_epochs - 1, 0, 0.011])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Piecewise Constant Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
### Performance Scheduling
```
tf.random.set_seed(42)
np.random.seed(42)
lr_scheduler = keras.callbacks.ReduceLROnPlateau(factor=0.5, patience=5)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
optimizer = keras.optimizers.SGD(learning_rate=0.02, momentum=0.9)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, history.history["lr"], "bo-")
plt.xlabel("Epoch")
plt.ylabel("Learning Rate", color='b')
plt.tick_params('y', colors='b')
plt.gca().set_xlim(0, n_epochs - 1)
plt.grid(True)
ax2 = plt.gca().twinx()
ax2.plot(history.epoch, history.history["val_loss"], "r^-")
ax2.set_ylabel('Validation Loss', color='r')
ax2.tick_params('y', colors='r')
plt.title("Reduce LR on Plateau", fontsize=14)
plt.show()
```
### tf.keras schedulers
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
s = 20 * len(X_train) // 32 # number of steps in 20 epochs (batch size = 32)
learning_rate = keras.optimizers.schedules.ExponentialDecay(0.01, s, 0.1)
optimizer = keras.optimizers.SGD(learning_rate)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
For piecewise constant scheduling, try this:
```
learning_rate = keras.optimizers.schedules.PiecewiseConstantDecay(
boundaries=[5. * n_steps_per_epoch, 15. * n_steps_per_epoch],
values=[0.01, 0.005, 0.001])
```
### 1Cycle scheduling
```
K = keras.backend
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_batch_end(self, batch, logs):
self.rates.append(K.get_value(self.model.optimizer.learning_rate))
self.losses.append(logs["loss"])
K.set_value(self.model.optimizer.learning_rate, self.model.optimizer.learning_rate * self.factor)
def find_learning_rate(model, X, y, epochs=1, batch_size=32, min_rate=10**-5, max_rate=10):
init_weights = model.get_weights()
iterations = math.ceil(len(X) / batch_size) * epochs
factor = np.exp(np.log(max_rate / min_rate) / iterations)
init_lr = K.get_value(model.optimizer.learning_rate)
K.set_value(model.optimizer.learning_rate, min_rate)
exp_lr = ExponentialLearningRate(factor)
history = model.fit(X, y, epochs=epochs, batch_size=batch_size,
callbacks=[exp_lr])
K.set_value(model.optimizer.learning_rate, init_lr)
model.set_weights(init_weights)
return exp_lr.rates, exp_lr.losses
def plot_lr_vs_loss(rates, losses):
plt.plot(rates, losses)
plt.gca().set_xscale('log')
plt.hlines(min(losses), min(rates), max(rates))
plt.axis([min(rates), max(rates), min(losses), (losses[0] + min(losses)) / 2])
plt.xlabel("Learning rate")
plt.ylabel("Loss")
```
**Warning**: In the `on_batch_end()` method, `logs["loss"]` used to contain the batch loss, but in TensorFlow 2.2.0 it was replaced with the mean loss (since the start of the epoch). This explains why the graph below is much smoother than in the book (if you are using TF 2.2 or above). It also means that there is a lag between the moment the batch loss starts exploding and the moment the explosion becomes clear in the graph. So you should choose a slightly smaller learning rate than you would have chosen with the "noisy" graph. Alternatively, you can tweak the `ExponentialLearningRate` callback above so it computes the batch loss (based on the current mean loss and the previous mean loss):
```python
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_epoch_begin(self, epoch, logs=None):
self.prev_loss = 0
def on_batch_end(self, batch, logs=None):
batch_loss = logs["loss"] * (batch + 1) - self.prev_loss * batch
self.prev_loss = logs["loss"]
self.rates.append(K.get_value(self.model.optimizer.learning_rate))
self.losses.append(batch_loss)
K.set_value(self.model.optimizer.learning_rate, self.model.optimizer.learning_rate * self.factor)
```
```
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(learning_rate=1e-3),
metrics=["accuracy"])
batch_size = 128
rates, losses = find_learning_rate(model, X_train_scaled, y_train, epochs=1, batch_size=batch_size)
plot_lr_vs_loss(rates, losses)
class OneCycleScheduler(keras.callbacks.Callback):
def __init__(self, iterations, max_rate, start_rate=None,
last_iterations=None, last_rate=None):
self.iterations = iterations
self.max_rate = max_rate
self.start_rate = start_rate or max_rate / 10
self.last_iterations = last_iterations or iterations // 10 + 1
self.half_iteration = (iterations - self.last_iterations) // 2
self.last_rate = last_rate or self.start_rate / 1000
self.iteration = 0
def _interpolate(self, iter1, iter2, rate1, rate2):
return ((rate2 - rate1) * (self.iteration - iter1)
/ (iter2 - iter1) + rate1)
def on_batch_begin(self, batch, logs):
if self.iteration < self.half_iteration:
rate = self._interpolate(0, self.half_iteration, self.start_rate, self.max_rate)
elif self.iteration < 2 * self.half_iteration:
rate = self._interpolate(self.half_iteration, 2 * self.half_iteration,
self.max_rate, self.start_rate)
else:
rate = self._interpolate(2 * self.half_iteration, self.iterations,
self.start_rate, self.last_rate)
self.iteration += 1
K.set_value(self.model.optimizer.learning_rate, rate)
n_epochs = 25
onecycle = OneCycleScheduler(math.ceil(len(X_train) / batch_size) * n_epochs, max_rate=0.05)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs, batch_size=batch_size,
validation_data=(X_valid_scaled, y_valid),
callbacks=[onecycle])
```
# Avoiding Overfitting Through Regularization
## $\ell_1$ and $\ell_2$ regularization
```
layer = keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01))
# or l1(0.1) for ℓ1 regularization with a factor of 0.1
# or l1_l2(0.1, 0.01) for both ℓ1 and ℓ2 regularization, with factors 0.1 and 0.01 respectively
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01)),
keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01)),
keras.layers.Dense(10, activation="softmax",
kernel_regularizer=keras.regularizers.l2(0.01))
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
from functools import partial
RegularizedDense = partial(keras.layers.Dense,
activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01))
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
RegularizedDense(300),
RegularizedDense(100),
RegularizedDense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
## Dropout
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(300, activation="elu", kernel_initializer="he_normal"),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(100, activation="elu", kernel_initializer="he_normal"),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
## Alpha Dropout
```
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(10, activation="softmax")
])
optimizer = keras.optimizers.SGD(learning_rate=0.01, momentum=0.9, nesterov=True)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 20
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
model.evaluate(X_train_scaled, y_train)
history = model.fit(X_train_scaled, y_train)
```
## MC Dropout
```
tf.random.set_seed(42)
np.random.seed(42)
y_probas = np.stack([model(X_test_scaled, training=True)
for sample in range(100)])
y_proba = y_probas.mean(axis=0)
y_std = y_probas.std(axis=0)
np.round(model.predict(X_test_scaled[:1]), 2)
np.round(y_probas[:, :1], 2)
np.round(y_proba[:1], 2)
y_std = y_probas.std(axis=0)
np.round(y_std[:1], 2)
y_pred = np.argmax(y_proba, axis=1)
accuracy = np.sum(y_pred == y_test) / len(y_test)
accuracy
class MCDropout(keras.layers.Dropout):
def call(self, inputs):
return super().call(inputs, training=True)
class MCAlphaDropout(keras.layers.AlphaDropout):
def call(self, inputs):
return super().call(inputs, training=True)
tf.random.set_seed(42)
np.random.seed(42)
mc_model = keras.models.Sequential([
MCAlphaDropout(layer.rate) if isinstance(layer, keras.layers.AlphaDropout) else layer
for layer in model.layers
])
mc_model.summary()
optimizer = keras.optimizers.SGD(learning_rate=0.01, momentum=0.9, nesterov=True)
mc_model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
mc_model.set_weights(model.get_weights())
```
Now we can use the model with MC Dropout:
```
np.round(np.mean([mc_model.predict(X_test_scaled[:1]) for sample in range(100)], axis=0), 2)
```
## Max norm
```
layer = keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal",
kernel_constraint=keras.constraints.max_norm(1.))
MaxNormDense = partial(keras.layers.Dense,
activation="selu", kernel_initializer="lecun_normal",
kernel_constraint=keras.constraints.max_norm(1.))
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
MaxNormDense(300),
MaxNormDense(100),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
# Exercises
## 1. to 7.
See appendix A.
## 8. Deep Learning on CIFAR10
### a.
*Exercise: Build a DNN with 20 hidden layers of 100 neurons each (that's too many, but it's the point of this exercise). Use He initialization and the ELU activation function.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
activation="elu",
kernel_initializer="he_normal"))
```
### b.
*Exercise: Using Nadam optimization and early stopping, train the network on the CIFAR10 dataset. You can load it with `keras.datasets.cifar10.load_data()`. The dataset is composed of 60,000 32 × 32–pixel color images (50,000 for training, 10,000 for testing) with 10 classes, so you'll need a softmax output layer with 10 neurons. Remember to search for the right learning rate each time you change the model's architecture or hyperparameters.*
Let's add the output layer to the model:
```
model.add(keras.layers.Dense(10, activation="softmax"))
```
Let's use a Nadam optimizer with a learning rate of 5e-5. I tried learning rates 1e-5, 3e-5, 1e-4, 3e-4, 1e-3, 3e-3 and 1e-2, and I compared their learning curves for 10 epochs each (using the TensorBoard callback, below). The learning rates 3e-5 and 1e-4 were pretty good, so I tried 5e-5, which turned out to be slightly better.
```
optimizer = keras.optimizers.Nadam(learning_rate=5e-5)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
```
Let's load the CIFAR10 dataset. We also want to use early stopping, so we need a validation set. Let's use the first 5,000 images of the original training set as the validation set:
```
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.cifar10.load_data()
X_train = X_train_full[5000:]
y_train = y_train_full[5000:]
X_valid = X_train_full[:5000]
y_valid = y_train_full[:5000]
```
Now we can create the callbacks we need and train the model:
```
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_model.h5", save_best_only=True)
run_index = 1 # increment every time you train the model
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
%tensorboard --logdir=./my_cifar10_logs --port=6006
model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_model.h5")
model.evaluate(X_valid, y_valid)
```
The model with the lowest validation loss gets about 47.6% accuracy on the validation set. It took 27 epochs to reach the lowest validation loss, with roughly 8 seconds per epoch on my laptop (without a GPU). Let's see if we can improve performance using Batch Normalization.
### c.
*Exercise: Now try adding Batch Normalization and compare the learning curves: Is it converging faster than before? Does it produce a better model? How does it affect training speed?*
The code below is very similar to the code above, with a few changes:
* I added a BN layer after every Dense layer (before the activation function), except for the output layer. I also added a BN layer before the first hidden layer.
* I changed the learning rate to 5e-4. I experimented with 1e-5, 3e-5, 5e-5, 1e-4, 3e-4, 5e-4, 1e-3 and 3e-3, and I chose the one with the best validation performance after 20 epochs.
* I renamed the run directories to run_bn_* and the model file name to my_cifar10_bn_model.h5.
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
model.add(keras.layers.BatchNormalization())
for _ in range(20):
model.add(keras.layers.Dense(100, kernel_initializer="he_normal"))
model.add(keras.layers.BatchNormalization())
model.add(keras.layers.Activation("elu"))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(learning_rate=5e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_bn_model.h5", save_best_only=True)
run_index = 1 # increment every time you train the model
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_bn_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_bn_model.h5")
model.evaluate(X_valid, y_valid)
```
* *Is the model converging faster than before?* Much faster! The previous model took 27 epochs to reach the lowest validation loss, while the new model achieved that same loss in just 5 epochs and continued to make progress until the 16th epoch. The BN layers stabilized training and allowed us to use a much larger learning rate, so convergence was faster.
* *Does BN produce a better model?* Yes! The final model is also much better, with 54.0% accuracy instead of 47.6%. It's still not a very good model, but at least it's much better than before (a Convolutional Neural Network would do much better, but that's a different topic, see chapter 14).
* *How does BN affect training speed?* Although the model converged much faster, each epoch took about 12s instead of 8s, because of the extra computations required by the BN layers. But overall the training time (wall time) was shortened significantly!
### d.
*Exercise: Try replacing Batch Normalization with SELU, and make the necessary adjustements to ensure the network self-normalizes (i.e., standardize the input features, use LeCun normal initialization, make sure the DNN contains only a sequence of dense layers, etc.).*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(learning_rate=7e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_selu_model.h5", save_best_only=True)
run_index = 1 # increment every time you train the model
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_selu_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
X_means = X_train.mean(axis=0)
X_stds = X_train.std(axis=0)
X_train_scaled = (X_train - X_means) / X_stds
X_valid_scaled = (X_valid - X_means) / X_stds
X_test_scaled = (X_test - X_means) / X_stds
model.fit(X_train_scaled, y_train, epochs=100,
validation_data=(X_valid_scaled, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_selu_model.h5")
model.evaluate(X_valid_scaled, y_valid)
model = keras.models.load_model("my_cifar10_selu_model.h5")
model.evaluate(X_valid_scaled, y_valid)
```
We get 47.9% accuracy, which is not much better than the original model (47.6%), and not as good as the model using batch normalization (54.0%). However, convergence was almost as fast as with the BN model, plus each epoch took only 7 seconds. So it's by far the fastest model to train so far.
### e.
*Exercise: Try regularizing the model with alpha dropout. Then, without retraining your model, see if you can achieve better accuracy using MC Dropout.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(learning_rate=5e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_alpha_dropout_model.h5", save_best_only=True)
run_index = 1 # increment every time you train the model
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_alpha_dropout_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
X_means = X_train.mean(axis=0)
X_stds = X_train.std(axis=0)
X_train_scaled = (X_train - X_means) / X_stds
X_valid_scaled = (X_valid - X_means) / X_stds
X_test_scaled = (X_test - X_means) / X_stds
model.fit(X_train_scaled, y_train, epochs=100,
validation_data=(X_valid_scaled, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_alpha_dropout_model.h5")
model.evaluate(X_valid_scaled, y_valid)
```
The model reaches 48.9% accuracy on the validation set. That's very slightly better than without dropout (47.6%). With an extensive hyperparameter search, it might be possible to do better (I tried dropout rates of 5%, 10%, 20% and 40%, and learning rates 1e-4, 3e-4, 5e-4, and 1e-3), but probably not much better in this case.
Let's use MC Dropout now. We will need the `MCAlphaDropout` class we used earlier, so let's just copy it here for convenience:
```
class MCAlphaDropout(keras.layers.AlphaDropout):
def call(self, inputs):
return super().call(inputs, training=True)
```
Now let's create a new model, identical to the one we just trained (with the same weights), but with `MCAlphaDropout` dropout layers instead of `AlphaDropout` layers:
```
mc_model = keras.models.Sequential([
MCAlphaDropout(layer.rate) if isinstance(layer, keras.layers.AlphaDropout) else layer
for layer in model.layers
])
```
Then let's add a couple utility functions. The first will run the model many times (10 by default) and it will return the mean predicted class probabilities. The second will use these mean probabilities to predict the most likely class for each instance:
```
def mc_dropout_predict_probas(mc_model, X, n_samples=10):
Y_probas = [mc_model.predict(X) for sample in range(n_samples)]
return np.mean(Y_probas, axis=0)
def mc_dropout_predict_classes(mc_model, X, n_samples=10):
Y_probas = mc_dropout_predict_probas(mc_model, X, n_samples)
return np.argmax(Y_probas, axis=1)
```
Now let's make predictions for all the instances in the validation set, and compute the accuracy:
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
y_pred = mc_dropout_predict_classes(mc_model, X_valid_scaled)
accuracy = np.mean(y_pred == y_valid[:, 0])
accuracy
```
We get no accuracy improvement in this case (we're still at 48.9% accuracy).
So the best model we got in this exercise is the Batch Normalization model.
### f.
*Exercise: Retrain your model using 1cycle scheduling and see if it improves training speed and model accuracy.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.SGD(learning_rate=1e-3)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
batch_size = 128
rates, losses = find_learning_rate(model, X_train_scaled, y_train, epochs=1, batch_size=batch_size)
plot_lr_vs_loss(rates, losses)
plt.axis([min(rates), max(rates), min(losses), (losses[0] + min(losses)) / 1.4])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.SGD(learning_rate=1e-2)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
n_epochs = 15
onecycle = OneCycleScheduler(math.ceil(len(X_train_scaled) / batch_size) * n_epochs, max_rate=0.05)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs, batch_size=batch_size,
validation_data=(X_valid_scaled, y_valid),
callbacks=[onecycle])
```
One cycle allowed us to train the model in just 15 epochs, each taking only 2 seconds (thanks to the larger batch size). This is several times faster than the fastest model we trained so far. Moreover, we improved the model's performance (from 47.6% to 52.0%). The batch normalized model reaches a slightly better performance (54%), but it's much slower to train.
| github_jupyter |
# Text Classification Overview
---
## Contents
1. [Overview](#1.-Overview)
2. [How to represent a sentence or tokens?](#2.-How-to-represent-a-sentence-or-tokens?)
```
import numpy as np
np.random.seed(777)
```
---
## 1. Overview
Examples below are **"Text Classification"** problem.
* Sentiment analysis: is this review positive or negative?
* Text categorization: which category does this blog post belong to?
* Intent classification: is this a question about a Chinese restaurant?
We can define "Text Classification" as:
* Input: A natural language sentence / paragraph
* Output: a category to which the input text belongs
- There are a fixed number **C** of categories
---
## 2. How to represent a sentence or tokens?
What is a sentence? A sentence can be viewed as a variable-length sequence of tokens. Each token could be any one from a vocabulary.
It's very similar to talking. For example, "the quick brown fox jumps over the lazy dog" can be viewed as a list "\[the, quick, brown, fox, jumps, over, the, lazy, dog\]" that you speak.
$$X=(x_1, x_2, \cdots, x_t ,\cdots,x_T) \quad where\ x_t \in V$$
So, a sentence $X$ becomes a sequence of tokens, and $V$ is vocabulary set which are total unique tokens in the training data.
The unit of token is not matter, you can try units like words, strings, or even digit bits. Here we take "word" as our token unit.
Since computer machine can't understand "words", so we change words to an integer indices.
After encoded, this sentence can be representated as **a sequence of integer indicies**.
```
sentence = "the quick brown fox jumps over the lazy dog".split()
vocab = {}
for token in sentence:
if vocab.get(token) is None:
vocab[token] = len(vocab)
sentence = list(map(vocab.get, sentence))
sentence
```
The other method to represent a token is called **"one-hot representaion"**. Every token has a length of vocabulary size and only one of the elements is 1 at vocabulary index.
For example: The token "the" has index 0 in our vocabulary. So,
$$the = [1, 0, 0, 0, 0, 0, 0, 0]$$
```
one_hot = np.eye(len(vocab), dtype=np.int)
print('index of "the" in the vocabulary:', vocab['the'])
print('one-hot vector:', one_hot[vocab['the']])
```
However these integer indicies are arbitrary and they can't capture the "meaning" of words.
**What is the "meaning(semantic)" of a word?**
There are some hypotheses in Natural Language Processing. From the paper ["From Frequency to Meaning: Vector Space Models of Semantics"](https://arxiv.org/abs/1003.1141)
>**Statistical semantics hypothesis**: Statistical patterns of human word usage can be
used to figure out what people mean (Weaver, 1955; Furnas et al., 1983). – If units of text
have similar vectors in a text frequency matrix, then they tend to have similar meanings.
(We take this to be a general hypothesis that subsumes the four more specific hypotheses
that follow.)
>
>**Bag of words hypothesis**: The frequencies of words in a document tend to indicate
the relevance of the document to a query (Salton et al., 1975). – If documents and pseudodocuments
(queries) have similar column vectors in a term–document matrix, then they tend to have similar meanings.
>
>**Distributional hypothesis**: Words that occur in similar contexts tend to have similar meanings (Harris, 1954; Firth, 1957; Deerwester et al., 1990). – If words have similar row vectors in a word–context matrix, then they tend to have similar meanings.
>
>**Extended distributional hypothesis**: Patterns that co-occur with similar pairs tend
to have similar meanings (Lin & Pantel, 2001). – If patterns have similar column vectors
in a pair–pattern matrix, then they tend to express similar semantic relations.
>
>**Latent relation hypothesis**: Pairs of words that co-occur in similar patterns tend
to have similar semantic relations (Turney et al., 2003). – If word pairs have similar row
vectors in a pair–pattern matrix, then they tend to have similar semantic relations.
According to these hypotheses, we can know that the meaning of a word can be represented as a vector. Also, Similar vectors have similar meanings which means the distance between two vectors will be closer. **Cosine similarity** is a measure of similarity between two non-zero vectors of an inner product space that measures the cosine of the angle between them. So, we use it to measure the similarity of the meaning between two words.
However, all one hot representations have same distance "0" when using cosine similiarity to measure their distance.
```
def cos_similiarity(x, y):
return (np.dot(x, y) / (np.linalg.norm(x)*np.linalg.norm(y))).round(5)
word1 = one_hot[vocab['fox']] # array([0, 0, 0, 1, 0, 0, 0, 0])
word2 = one_hot[vocab['dog']] # array([0, 0, 0, 0, 0, 0, 0, 1])
print('cosine similiarity between word "fox" and word "dog" is :', cos_similiarity(word1, word2))
```
Then, how should we represent a token so that it reflects its "meaning"? If we want to calculate some similiarity, we need to use **dense vectors** to represent these words. One-hot vectors are **sparse vectors** which contains a lot of zeros.
```
vec1 = np.array([1, 2, 3, 4])
vec2 = np.array([1, 2, 3, 5])
print('cosine similiarity between "vec1" and "vec2" is :', cos_similiarity(vec1, vec2))
```
So, how to slove this? If we assume there is a vector sapce ($\Bbb{R}^{\vert V \vert \times d} $) that can represent these tokens. Then train this vector space in neural network to solve a classification problem. This vector space will capture the token's meaning. This process can be done by a simple matrix multiplication. Let's call word representaion vector space as $W$ which the size is $(\vert V \vert, d)$, since we changed tokens to one hot vectors, by doing matrix multiplication, can get a single row vector of $W$$(w_i)$ and this vector will be the meaning of the word.
$$w_i = t_i \cdot W , \quad where\ t_i = [0, \cdots, \underset{i\text{-}th\ index}{1}, \cdots, 0],\ len(t_i)=\vert V \vert$$
```
d = 5
W = np.random.rand(len(vocab), d).round(3)
print('vector space for all tokens, size of (len(vocab), d)')
print(W)
print()
print('a dense vector representation for word "fox", at column index 3')
print(np.dot(word1, W)) # one-hot token of word "fox" = array([0, 0, 0, 1, 0, 0, 0, 0])
```
In practical, we just get the index of row vector of vector space $W$. At backward time, update the index of previous gradient.
```
idxes = [np.argmax(v) for v in [word1, word2]]
print('Index for word "fox" and "dog" is {}'.format(idxes))
print('Row vector for word "fox" and "dog": ')
print(W[idxes, :])
dout = np.random.rand(2, d).round(3)
dW = np.zeros_like(W)
dW[idxes] = dout
print('Update word "fox" & "dog": ')
print(dW)
```
for same indices, all vector refer to same indices will be summed together.
```
idxes = [np.argmax(v) for v in [word1, word2, word1]]
dout = dout[np.array([0, 1, 0])]
temp = {i: a for i, a in zip(idxes, dout)}
dW = np.zeros_like(W)
for i in idxes:
dW[i] += 1
for i, v in temp.items():
dW[i] = dW[i] * v
print('Update word "fox" & "dog": ')
print(dW)
```
This process is called **Embedding**. In pytorch, use "Embedding" Layer for embedding words to dense vector space.
```
import torch
import torch.nn as nn
import torch.nn.functional as F
# you can set your pretrained vector parameters giving "_weight" args by "torch.Tensor" type
embed_layer = nn.Embedding(len(vocab), d, _weight=torch.FloatTensor(W))
idxes_tensor = torch.LongTensor(idxes)
embeded = embed_layer(idxes_tensor)
print('Embedding Layer parameters: ')
print(embed_layer.weight)
print()
print('Embeded vector for "fox" and "dog": ')
print(embeded)
dout_tensor = torch.tensor(dout, requires_grad=True)
print('Gradient before backward into Embedding Layer: ')
print(dout_tensor)
print()
print('Update word "fox" & "dog": ')
print(embeded.grad_fn(dout_tensor))
```
Very Easy to use, and after training, we can use word vectors in Embedding layer to calculate some similarity of two vectors.
```
wv1 = embed_layer.weight[vocab.get('fox')]
wv2 = embed_layer.weight[vocab.get('dog')]
print('Calculate similarity for "fox" & "dog" {:.4f}'.format(F.cosine_similarity(wv1, wv2, dim=0)))
```
| github_jupyter |
# Create multi-label chips for multi-output regression sampled from the whole GB
Create chips and store proportions of signature types within as an input of multi-output regression problem
```
import geopandas
import tobler
import pyogrio
import pygeos
import numpy
import pandas
import dask_geopandas
import rasterio
from scipy.sparse import coo_matrix
import dask
import dask.bag
from dask.distributed import Client, LocalCluster
%%time
df = geopandas.read_parquet("/home/jovyan/work/chips_gb/chip_bounds_32/").reset_index(drop=True)
df
signatures = pyogrio.read_dataframe(
'/home/jovyan/work/urbangrammar_samba/spatial_signatures/'
'signatures/'
'signatures_combined_levels_simplified.gpkg'
)
bds = df.total_bounds
signatures = signatures.cx[bds[0]:bds[2], bds[1]:bds[3]]
%%time
ids_src, ids_tgt = df.sindex.query_bulk(signatures.geometry, predicate="intersects")
od_matrix = pandas.DataFrame(dict(ids_src=ids_src, ids_tgt=ids_tgt))
%%time
sjoined = df.set_geometry(df.centroid).sjoin(signatures[["signature_type", "geometry"]], how="left", predicate="within")
od_matrix["signature_type"] = sjoined.signature_type.iloc[ids_tgt].values
od_matrix.signature_type.value_counts().plot.bar()
od_matrix.signature_type.value_counts()
cap = 20000
keep = []
counts = {k:0 for k in od_matrix.signature_type.unique()}
for i, t in od_matrix.signature_type.sample(len(od_matrix)).iteritems():
counts[t] += 1
if counts[t] < cap:
keep.append(i)
limited = od_matrix.iloc[keep]
limited.signature_type.value_counts().plot.bar()
limited.shape
%%time
chips = df.geometry.values[limited.ids_tgt.values].data
sig = signatures.geometry.values[limited.ids_src.values].data
# this could be parallelised
r = [
pygeos.clip_by_rect(
g, *chip
) for g, chip in zip(sig, pygeos.bounds(chips))
]
areas = pygeos.area(r)
table = coo_matrix(
(
areas,
(limited.ids_src.values, limited.ids_tgt.values),
),
shape=(signatures.shape[0], df.shape[0]),
dtype=numpy.float32,
)
table = table.tocsr()
%%time
signatures["signature_type"] = pandas.Categorical(signatures["signature_type"])
%%time
categorical = {}
for value in unique:
mask = signatures["signature_type"] == value
categorical[value] = numpy.asarray(
table[mask].sum(axis=0)
)[0]
print(value)
%%time
categorical = pandas.DataFrame(categorical)
%time areas = df.area.values
%time categorical = categorical.div(areas, axis="rows")
(categorical.sum(axis=1) == 0).sum()
geom = df[df.geometry.name].reset_index(drop=True)
ests = geopandas.GeoDataFrame(categorical, geometry=geom, crs=signatures.crs)
limited
ests = ests.iloc[limited.ids_tgt.values]
ests.drop(columns="geometry").sum().plot.bar()
ests.to_parquet("/home/jovyan/work/chips_gb/chip_proportions_32.pq")
ests.centroid.plot(figsize=(20, 20), markersize=.1)
```
Create chips
```
specs = {
'chip_size': 32,
'bands': [1, 2, 3], #RGB
'mosaic_p': (
'/home/jovyan/work/urbangrammar_samba/'
'ghs_composite_s2/GHS-composite-S2.vrt'
),
}
bounds = geopandas.read_parquet("/home/jovyan/work/chips_gb/chip_proportions_32.pq")
centroid = bounds.centroid
bounds['X'] = centroid.x.astype(int)
bounds['Y'] = centroid.y.astype(int)
client = Client(
LocalCluster(n_workers=16, threads_per_worker=1)
)
client
import numpy as np
def bag_of_chips(chip_bbs, specs, npartitions):
'''
Load imagery for `chip_bbs` using a Dask bag
...
Arguments
---------
chip_bbs : GeoDataFrame
Geo-table with bounding boxes of the chips to load
specs : dict
Metadata dict, including, at least:
- `bands`: band index of each band of interest
- `chip_size`: size of each chip size expressed in pixels
- `mosaic_p`: path to the mosaic/file of imagery
npartitions : int
No. of partitions to split `chip_bbs` before sending to
Dask for distributed computation
Returns
-------
chips : ndarray
Numpy tensor of (N, chip_size, chip_size, n_bands) dimension
with imagery data
'''
# Split chip_bbs
thr = np.linspace(0, chip_bbs.shape[0], npartitions+1, dtype=int)
chunks = [
(chip_bbs.iloc[thr[i]:thr[i+1], :], specs) for i in range(len(thr)-1)
]
# Set up the bag
bag = dask.bag.from_sequence(
chunks, npartitions=npartitions
).map(chip_loader)
# Compute
chips = np.concatenate(bag.compute())
return chips
def chip_loader(pars):
'''
Load imagery for `chip_bbs`
...
Arguments (wrapped in `pars`)
-----------------------------
chip_bbs : GeoDataFrame
Geo-table with bounding boxes of the chips to load
specs : dict
Metadata dict, including, at least:
- `bands`: band index of each band of interest
- `chip_size`: size of each chip size expressed in pixels
- `mosaic_p`: path to the mosaic/file of imagery
Returns
-------
chips : ndarray
Numpy tensor of (N, chip_size, chip_size, n_bands) dimension
with imagery data
'''
chip_bbs, specs = pars
b = len(specs['bands'])
s = specs['chip_size']
chips = np.zeros((chip_bbs.shape[0], b, s, s))
with rasterio.open(specs['mosaic_p']) as src:
for i, tup in enumerate(chip_bbs.itertuples()):
img, transform = rasterio.mask.mask(
src, [tup.geometry], crop=True, all_touched=True
)
img = img[:b, :s, :s]
for ban, (l_min, l_max) in enumerate([(350, 1600), (500, 1600), (600, 1800)]):
img[ban][img[ban] > l_max] = l_max
img[ban][img[ban] < l_min] = l_min
a_std = (img[ban] - l_min) / (l_max - l_min)
img[ban] = a_std * 255
chips[i, :, :, :] = img
chips = np.moveaxis(chips, 1, -1)
return chips.astype(rasterio.uint8)
chips = bag_of_chips(bounds, specs, 16)
numpy.save('../../chips_gb/multilabel_chip_32.npy', chips)
```
| github_jupyter |
I've been writing a lot about a [category theory interpretations of data-processing pipelines](http://www.win-vector.com/blog/2019/12/data_algebra-rquery-as-a-category-over-table-descriptions/) and [some of the improvements we feel it is driving](http://www.win-vector.com/blog/2019/12/better-sql-generation-via-the-data_algebra/) in both the [`data_algebra`](https://github.com/WinVector/data_algebra) and in [`rquery`](https://github.com/WinVector/rquery)/[`rqdatatable`](https://github.com/WinVector/rqdatatable).
I think I've found an even better category theory re-formulation of the package, which I will describe here.
In the [earlier formalism](http://www.win-vector.com/blog/2019/12/data_algebra-rquery-as-a-category-over-table-descriptions/) our data transform pipelines were arrows over a category of sets of column names (sets of strings). We will call this the "column equality" version of our category.
These pipelines acted on `Pandas` tables or `SQL` tables, with one table marked as special. Marking one table as special (or using a "pointed set" notation) lets us use a nice compositional notation, without having to appeal to something like operads. Treating one table as the one of interest is fairly compatible with data science, as in data science often when working with many tables one is the primary model-frame and the rest are used to join in additional information.
The above formulation was really working well. But we have found a variation of the `data_algebra` with an even neater formalism.
The `data_algebra` objects have a very nice interpretation as arrows in a category whose objects are set families described by:
* a set of required column names.
* a set of forbidden column names.
We will call this the "set ordered" version of our category.
The arrows `a` and `b` compose as `a >> b` as long as:
* All of the columns required by `b` are produced by `a`.
* None of the columns forbidden by `b` are produced by `a`.
This is still an equality check of domains and co-domains, so as long as we maintain associativity we still have a nice category.
We can illustrate the below.
First we import our modules.
```
import sqlite3
import pandas
from data_algebra.data_ops import *
from data_algebra.arrow import fmt_as_arrow
import data_algebra.SQLite
```
We define our first arrow which is a transform that creates a new column `x` as the sum of the columns `a` and `b`.
```
a = TableDescription(
table_name='table_a',
column_names=['a', 'b']). \
extend({'c': 'a + b'})
a
print(fmt_as_arrow(a))
```
And we define our second arrow, `b`, which renames the column `a` to a new column name `x`.
```
b = TableDescription(table_name='table_b', column_names=['a']). \
rename_columns({'x': 'a'})
b
print(fmt_as_arrow(b))
```
The rules are met, so we can combine these two arrows.
```
ab = a >> b
ab
print(fmt_as_arrow(ab))
```
Notice this produces a new arrow `ab` with appropriate required and forbidden columns. By associativity (one of the primary properties needed to be a category) we get that the arrow `ab` has an action on data frames the same as using the `a` action followed by the `b` action.
Let's illustrate that here.
```
d = pandas.DataFrame({
'a': [1, 2],
'b': [30, 40]
})
d
b.act_on(a.act_on(d))
ab.act_on(d)
```
`.act_on()` copies forward all columns consistent with the transform specification and used at the output. Missing columns are excess columns are checked for at the start of a calculation.
```
excess_frame = pandas.DataFrame({
'a': [1],
'b': [2],
'd': [3],
'x': [4]})
try:
ab.act_on(excess_frame)
except ValueError as ve:
print("caught ValueError: " + str(ve))
```
The `.transform()` method, on the other hand, copies forward only declared columns.
```
ab.transform(excess_frame)
```
Notice in the above that the input `x` did not interfere with the calculation, and `d` was not copied forward. The idea is behavior during composition is very close to behavior during action/application, so we find more issues during composition.
However, `.transform()` does not associate with composition in the set ordered verson of our category, so is not an action of this category. The issue is: we have `b.transform(a.transform(d))` is not equal to `ab.transform(d)` in this category.
```
b.transform(a.transform(d))
```
`.transform()` does associate with the arrows of the stricter [identical column set category we demonstrated earlier](https://github.com/WinVector/data_algebra/blob/master/Examples/Arrow/CDesign.md), so it is an action of this category.
So we have two categories, `.act_on()` is the category compatible action for one, and `.transform()` for the other.
In both cases we still have result-oriented narrowing.
```
c = TableDescription(
table_name='table_c',
column_names=['a', 'b', 'c']). \
extend({'x': 'a + b'}). \
select_columns({'x'})
c
print(fmt_as_arrow(c))
table_c = pandas.DataFrame({
'a': [1, 2],
'b': [30, 40],
'c': [500, 600],
'd': [7000, 8000]
})
table_c
c.act_on(table_c)
c.transform(table_c)
```
`.select_columns()` conditions are propagated back through the calculation.
Another useful operator is `.drop_columns()` which drops columns if they are present, but does not raise an issue if the columns to be removed are already not present. `.drop_columns()` can be used to guarantee forbidden columns are not present. We could use `.act_on()` or `excess_frame` using `.drop_columns()` as follows.
```
tdr = describe_table(excess_frame). \
drop_columns(['x'])
tdr
rab = tdr >> ab
rab
```
The `>>` notation is composing the arrows. `tdr >> ab` is syntactic sugar for `ab.apply_to(tdr)`. Both of these are the arrow composition operations.
```
rab.act_on(excess_frame)
```
Remember, the original `ab` operator rejects `excess_frame`.
```
try:
ab.act_on(excess_frame)
except ValueError as ve:
print("caught ValueError: " + str(ve))
```
We can also adjust the input-specification by composing pipelines with table descriptions.
```
a
bigger = TableDescription(
table_name='bigger',
column_names=['a', 'b', 'x', 'y', 'z'])
bigger
bigger_a = bigger >> a
bigger_a
print(fmt_as_arrow(bigger_a))
```
Notice the new arrow (`bigger_a`) has a wider input specification. Appropriate checking is performed during the composition.
As always, we can also translate any of our operators to `SQL`.
```
db_model = data_algebra.SQLite.SQLiteModel()
print(bigger_a.to_sql(db_model=db_model, pretty=True))
```
The `SQL` translation does not use "`*`", so it is similar to `.transform()` in that it only refers to known columns by name. This means we are safe from extra columns in the source tables. This means if we did derive an action acting on `SQL` or composition over `SQL` it would not associate with the `data_algebra` operator composition (just as `.transform()` did not).
So the column identity is category is a model of our `SQL` translation scheme (needs exact column definitions), and the set ordered category is a model of our in-memory data processing (accepts extra columns). These two systems being slightly different is mitigated by the fact each has an orderly model as a category. So we have tools, in principle, to formally talk about how they relate to each other (again using category theory).
Notice we no longer have to use the arrow-adapter classes (except for formatting), the `data_algebra` itself has been adjusted to a more direct categorical basis.
And that is some of how the `data_algebra` works on our new set-oriented category. In this formulation much less annotation is required from the user, while still allowing very detailed record-keeping. The detailed record-keeping lets us find issues while assembling the pipelines, not later when working with potentially large/slow data.
| github_jupyter |
## Linear Algebra
- Why we need Linear Algebra
1- To solve a system of linear equations
2- Most of machine learning models and all most all deep learning models need it
## Matrix-Vector and Matrix-Matrix Multiplication
- On piece of paper, multiply the following matrix-vector:
<img src="matrix_matrix.png" width="300" height="300">
- On piece of paper, multiply the following matrices:
<img src="matrix_vector.png" width="300" height="300">
## Verify your answer in Python using Numpy
```
A = np.array([[1, 2], [0, 1], [2, 3]])
v = np.array([[2], [6]])
print(A)
print(v)
print(np.dot(A, v))
A = np.array([[1, 2], [0, 1], [2, 3]])
B = np.array([[2, 5], [6, 7]])
print(A)
print(B)
print(np.dot(A, B))
```
## Question: Can we express linear regression in matrix-vector format?
```
import numpy as np
import matplotlib.pyplot as plt
# Running Distance in Mile
x = np.array([3.3,4.4,5.5,6.71,6.93,4.168,9.779,6.182,7.59,2.167,
7.042,10.791,5.313,7.997,5.654,9.27,3.1])
# Water Drinks in Litre
y = np.array([1.7,2.76,2.09,3.19,1.694,1.573,3.366,2.596,2.53,1.221,
2.827,3.465,1.65,2.904,2.42,2.94,1.3])
```
## We can obtain the prediction vector as the following:
```
w1 = 0.25163494
w0 = 0.79880123
y_pred = [w1*i + w0 for i in x]
print(y_pred)
```
## Also, we can define our feature matrix as X and weight vector as w
```
# print(np.ones((len(x), 1)))
# print(np.transpose([x]))
# Concatenate two matrix column-wise
X = np.concatenate((np.transpose([x]), np.ones((len(x), 1))), axis=1)
print(X)
X.shape
w = np.array([w1, w0])
np.dot(X, w)
```
## Another way
```
v_ones = np.ones((1, len(x)))
print(v_ones)
X = np.array([x, np.ones((1, len(x)))])
print(X)
w = np.array([w1, w0])
np.dot(w, X)
```
## Transpose of a Matrix or a Vector
- In linear algebra, the transpose of a matrix is an operator which switches the row and column indices of the matrix by producing another matrix denoted as Aᵀ
<img src="matrix-transpose.jpg" width="300" height="300">
## Transpose the Matrix in Numpy
```
A = np.array([[1, 2, 3], [4, 5, 6], [7, 8, 9]])
print(A)
print(A.T)
```
## Norm of a vector
- We have different norm, here we mean L2-norm, which measures the length of a vector from origin
## Activity: what is the length of the following vector:
- v = [3, 4]
```
from numpy import linalg as LA
v = np.array([3, 4])
LA.norm(v)
```
## Activity: Show that norm of a vector is the sqrt of V*V.T
```
v = np.array([3, 4])
np.sqrt(np.dot(v,v.T))
```
## Activity: The distance between two vector u and v
<img src="norm.png" width="500" height="500">
```
u = np.array([1, 1])
v = np.array([2, 2])
r = np.array([3, 8])
print(LA.norm(u - v))
print(LA.norm(u - r))
print(LA.norm(v - r))
```
## Resources:
- https://docs.scipy.org/doc/numpy/reference/generated/numpy.concatenate.html
- http://matrixmultiplication.xyz
| github_jupyter |
```
import os
if os.getcwd().endswith('visualization'):
os.chdir('..')
import gmaps
import numpy as np
import pandas as pd
from sklearn.externals import joblib
from model import load_clean_data_frame
API_KEY = 'AIzaSyCgyB-8lqWqGhTYSlt2VuJyeuEVotFoYO8'
gmaps.configure(api_key=API_KEY)
model = joblib.load('model/output/xgboost_basic_fs.joblib')
raw_crime_data = load_clean_data_frame()
locations = raw_crime_data[['longitude', 'latitude']].sample(frac=1)
min_longitude = np.min(locations['longitude'])
max_longitude = np.max(locations['longitude'])
min_latitude = np.min(locations['latitude'])
max_latitude = np.max(locations['latitude'])
homicide_data = raw_crime_data.loc[raw_crime_data['type'] == 22.0]
homicide_data = homicide_data[['location', 'iucr', 'hour', 'month', 'type', 'fbi_code']]
homicide_data = homicide_data.sample(frac=1)
homicide_data.head()
# Modeling homicide occuring in an alley
vis_location = 11.0
vis_iucr = 102.0
vis_hour = 2.0
vis_month = 7.0
vis_type = 22.0
vis_fbi_code = 23.0
columns = ['location', 'iucr', 'hour', 'month', 'type', 'fbi_code', 'latitude', 'longitude']
predicted_columns = ['latitude', 'longitude', 'arrest']
predicted_data = pd.DataFrame()
count = 1
for index, row in locations.iterrows():
point = pd.DataFrame([[
vis_location, vis_iucr, vis_hour, vis_month, vis_type, vis_fbi_code,
row['latitude'], row['longitude']
]], columns=columns)
predicted = model.predict(point)
predicted = (model.predict_proba(point)[:, 1] >= 0.2905).astype(float)
predicted_row = pd.DataFrame([[
row['latitude'], row['longitude'], predicted[0]
]], columns=predicted_columns)
predicted_data = predicted_data.append(predicted_row)
count += 1
if count > 3000:
break
homicide_arrests = predicted_data.loc[predicted_data['arrest'] == 1.0]
homicide_no_arrests = predicted_data.loc[predicted_data['arrest'] == 0.0]
print(len(homicide_arrests))
print(len(homicide_no_arrests))
figure_layout = {
'width': '800px',
'height': '800px'
}
fig = gmaps.figure(layout=figure_layout, center=[41.836944, -87.684722], zoom_level=11)
heatmap_layer = gmaps.heatmap_layer(
homicide_no_arrests[['latitude', 'longitude']], weights=homicide_no_arrests['arrest']+1,
max_intensity=1.0, point_radius=10.0, dissipating=True,
opacity=1, gradient=[(255, 0, 0, 0), (255, 0, 0, 1)]
)
fig.add_layer(heatmap_layer)
heatmap_layer = gmaps.heatmap_layer(
homicide_arrests[['latitude', 'longitude']], weights=homicide_arrests['arrest'],
max_intensity=1.0, point_radius=8.0, dissipating=True,
opacity=0.5, gradient=[(0, 0, 255, 0), (0, 0, 255, 1)]
)
fig.add_layer(heatmap_layer)
fig
homicide_truth = raw_crime_data.loc[raw_crime_data['type'] == 22.0]
homicide_truth = homicide_truth.sample(n=3000)
homicide_arrests_truth = homicide_truth.loc[homicide_truth['arrest'] == 1.0]
homicide_arrests_truth = homicide_arrests_truth.loc[homicide_arrests_truth['location'] == 11.0]
homicide_no_arrests_truth = homicide_truth.loc[homicide_truth['arrest'] == 0.0]
homicide_no_arrests_truth = homicide_no_arrests_truth.loc[homicide_no_arrests_truth['location'] == 11.0]
print(len(homicide_arrests_truth))
print(len(homicide_no_arrests_truth))
figure_layout = {
'width': '800px',
'height': '800px'
}
fig = gmaps.figure(layout=figure_layout, center=[41.836944, -87.684722], zoom_level=11)
heatmap_layer = gmaps.heatmap_layer(
homicide_no_arrests_truth[['latitude', 'longitude']], weights=homicide_no_arrests_truth['arrest']+1,
max_intensity=1.0, point_radius=15.0, dissipating=True,
opacity=1, gradient=[(255, 0, 0, 0), (255, 0, 0, 1)]
)
fig.add_layer(heatmap_layer)
heatmap_layer = gmaps.heatmap_layer(
homicide_arrests_truth[['latitude', 'longitude']], weights=homicide_arrests_truth['arrest'],
max_intensity=1.0, point_radius=15.0, dissipating=True,
opacity=0.5, gradient=[(0, 0, 255, 0), (0, 0, 255, 1)]
)
fig.add_layer(heatmap_layer)
fig
homicide_data = raw_crime_data.loc[raw_crime_data['type'] == 12.0]
homicide_data = homicide_data[['iucr', 'type', 'location', 'fbi_code', 'hour', 'property_crime', 'weekday', 'domestic']]
homicide_data = homicide_data.sample(frac=1)
homicide_data.head()
# Modeling assult occuring on a sidewalk.
vis_location = 3.0
vis_iucr = 74.0
vis_hour = 17.0
vis_month = 5.0
vis_type = 12.0
vis_fbi_code = 16.0
columns = ['location', 'iucr', 'hour', 'month', 'type', 'fbi_code', 'latitude', 'longitude']
predicted_columns = ['latitude', 'longitude', 'arrest']
predicted_data = pd.DataFrame()
count = 1
for index, row in locations.iterrows():
point = pd.DataFrame([[
vis_location, vis_iucr, vis_hour, vis_month, vis_type, vis_fbi_code,
row['latitude'], row['longitude']
]], columns=columns)
predicted = (model.predict_proba(point)[:, 1] >= 0.302).astype(float)
predicted_row = pd.DataFrame([[
row['latitude'], row['longitude'], predicted[0]
]], columns=predicted_columns)
predicted_data = predicted_data.append(predicted_row)
count += 1
if count > 3000:
break
assault_arrests = predicted_data.loc[predicted_data['arrest'] == 1.0]
assault_no_arrests = predicted_data.loc[predicted_data['arrest'] == 0.0]
print(len(assault_arrests))
print(len(assault_no_arrests))
figure_layout = {
'width': '800px',
'height': '800px'
}
fig = gmaps.figure(layout=figure_layout, center=[41.836944, -87.684722], zoom_level=11)
heatmap_layer = gmaps.heatmap_layer(
assault_no_arrests[['latitude', 'longitude']], weights=assault_no_arrests['arrest']+1,
max_intensity=1.0, point_radius=8.0, dissipating=True,
opacity=0.5, gradient=[(255, 0, 0, 0), (255, 0, 0, 1)]
)
fig.add_layer(heatmap_layer)
heatmap_layer = gmaps.heatmap_layer(
assault_arrests[['latitude', 'longitude']], weights=assault_arrests['arrest'],
max_intensity=1.0, point_radius=8.0, dissipating=True,
opacity=1, gradient=[(0, 0, 255, 0), (0, 0, 255, 1)]
)
fig.add_layer(heatmap_layer)
fig
assault_truth = raw_crime_data.loc[raw_crime_data['type'] == 12.0]
assault_truth = assault_truth.sample(n=3000)
assault_arrests_truth = assault_truth.loc[assault_truth['arrest'] == 1.0]
assault_arrests_truth = assault_arrests_truth.loc[assault_arrests_truth['location'] == 3.0]
assault_no_arrests_truth = assault_truth.loc[assault_truth['arrest'] == 0.0]
assault_no_arrests_truth = assault_no_arrests_truth.loc[assault_no_arrests_truth['location'] == 3.0]
print(len(assault_arrests_truth))
print(len(assault_no_arrests_truth))
figure_layout = {
'width': '800px',
'height': '800px'
}
fig = gmaps.figure(layout=figure_layout, center=[41.836944, -87.684722], zoom_level=11)
heatmap_layer = gmaps.heatmap_layer(
assault_no_arrests_truth[['latitude', 'longitude']], weights=assault_no_arrests_truth['arrest']+1,
max_intensity=1.0, point_radius=15.0, dissipating=True,
opacity=0.5, gradient=[(255, 0, 0, 0), (255, 0, 0, 1)]
)
fig.add_layer(heatmap_layer)
heatmap_layer = gmaps.heatmap_layer(
assault_arrests_truth[['latitude', 'longitude']], weights=assault_arrests_truth['arrest'],
max_intensity=1.0, point_radius=15.0, dissipating=True,
opacity=1, gradient=[(0, 0, 255, 0), (0, 0, 255, 1)]
)
fig.add_layer(heatmap_layer)
fig
```
| github_jupyter |
# Model to forecast inventory demand based on historical sales data.
```
%matplotlib inline
import numpy as np
import pandas as pd
from scipy import stats
import matplotlib.pyplot as plt
import time
import random
import pickle
import math
import warnings
warnings.filterwarnings("ignore")
```
## Model accuracy is RMSLE
```
def rmsle(y, y_pred):
assert len(y) == len(y_pred)
terms_to_sum = [(math.log(y_pred[i] + 1) - math.log(y[i] + 1)) ** 2.0 for i,pred in enumerate(y_pred)]
return (sum(terms_to_sum) * (1.0/len(y))) ** 0.5
```
## Load Training Data
The size of the training data is quite large (~4 GB). Large datasets require significant amount of memory to process. Instead, we will sample the data randomly for our initial data analysis and visualization.
```
def load_samp_data(filename='train.csv', columns=[], load_pkl=1):
"""
Function returns a dataframe containing the training data sampled randomly.
The data is also stored in a pickle file for later processing.
"""
if load_pkl:
inputfile = open('train_samp_data.pkl', 'rb')
data = pickle.load(inputfile)
inputfile.close()
return data
chunksize= 10 ** 6
datasize = 74180464 #datasize = sum(1 for line in open(filename)) - 1 #number of records in file (excludes header)
samplesize = 10 ** 3 # samples per chunk of data read from the file.
data = pd.DataFrame([],columns=columns)
chunks = pd.read_csv(filename, iterator=True, chunksize=chunksize)
for chunk in chunks:
chunk.columns = columns
data = data.append(chunk.sample(samplesize))
# write data to a pickle file.
outputfile = open('train_samp_data.pkl','wb')
pickle.dump(data,outputfile)
outputfile.close()
return data
load_pkl = 0
columns = ['week_num', 'sales_depot_id', 'sales_chan_id', 'route_id', 'client_id', 'prod_id', 'saleunit_curr_wk', 'saleamt_curr_wk', 'retunit_next_week', 'retamt_next_wk', 'y_pred_demand']
tic = time.time()
train_data_samp = load_samp_data('train.csv', columns, load_pkl)
toc = time.time()
print '*********'
print 'Time to load: ', toc-tic, 'sec'
print
print train_data_samp.describe()
print '*********'
print train_data_samp[['week_num', 'sales_depot_id', 'sales_chan_id', 'route_id', 'client_id', 'prod_id']]
features_train = train_data_samp[['week_num', 'sales_depot_id', 'sales_chan_id', 'route_id', 'client_id', 'prod_id']].values
labels_train_sale = train_data_samp[['saleunit_curr_wk']].values
labels_train_return = train_data_samp[['retunit_next_week']].values
labels_train = train_data_samp[['y_pred_demand']].values
```
## Feature Engineering
```
train_data_samp.groupby(['client_id', 'prod_id']).sum()
```
### Predict sale units $y_{sale}$ and returns $y_{return}$ using two different classifiers. We will use xgboost to fit $y_{sale}$ and $y_{return}$ with the input data.
```
# Utility function to report best scores
def report(grid_scores, n_top=3):
top_scores = sorted(grid_scores, key=itemgetter(1), reverse=True)[:n_top]
for i, score in enumerate(top_scores):
print("Model with rank: {0}".format(i + 1))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
score.mean_validation_score,
np.std(score.cv_validation_scores)))
print("Parameters: {0}".format(score.parameters))
print("")
import warnings
warnings.filterwarnings("ignore")
from sklearn.ensemble import RandomForestClassifier
from sklearn.grid_search import RandomizedSearchCV
from scipy.stats import randint as sp_randint
from operator import itemgetter
clf = RandomForestClassifier(n_estimators=10)
# specify parameters and distributions to sample from
param_dist = {"max_depth": [10],
"max_features": sp_randint(4, 7),
}
# run randomized search
n_iter_search = 10
random_search_sale = RandomizedSearchCV(clf, param_distributions=param_dist,
n_iter=n_iter_search, n_jobs=4, cv=5)
start = time.time()
random_search_sale.fit(features_train, np.ravel(labels_train_sale))
predict = random_search_sale.predict(features_train)
print 'Model Report ********'
print 'Accuracy : ', rmsle(np.ravel(labels_train_sale), predict)
print 'Model Report ********'
print("RandomizedSearchCV took %.2f seconds for %d candidates"
" parameter settings." % ((time.time() - start), n_iter_search))
report(random_search_sale.grid_scores_)
print random_search_sale.best_score_
print random_search_sale.best_estimator_
feat_imp = pd.Series(random_search_sale.best_estimator_.feature_importances_).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
import warnings
warnings.filterwarnings("ignore")
from sklearn.ensemble import RandomForestClassifier
from sklearn.grid_search import RandomizedSearchCV
from scipy.stats import randint as sp_randint
from operator import itemgetter
clf = RandomForestClassifier(n_estimators=15)
# specify parameters and distributions to sample from
param_dist = {"max_depth": [10],
"max_features": sp_randint(3, 5),
}
# run randomized search
n_iter_search = 10
random_search_return = RandomizedSearchCV(clf, param_distributions=param_dist,
n_iter=n_iter_search, n_jobs=4, cv=5)
start = time.time()
random_search_return.fit(features_train, np.ravel(labels_train_return))
predict = random_search_return.predict(features_train)
print 'Model Report ********'
print 'Accuracy : ', rmsle(np.ravel(labels_train_return), predict)
print 'Model Report ********'
print("RandomizedSearchCV took %.2f seconds for %d candidates"
" parameter settings." % ((time.time() - start), n_iter_search))
report(random_search_return.grid_scores_)
print random_search_return.best_score_
print random_search_return.best_estimator_
feat_imp = pd.Series(random_search_return.best_estimator_.feature_importances_).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
predict_sale = random_search_sale.predict(features_train)
predict_return = random_search_return.predict(features_train)
y_pred = [max(0,(predict_sale[i]-predict_return[i])) for i in xrange(len(predict_return))]
plt.scatter(y_pred,np.ravel(labels_train))
print 'Model Report ********'
print 'Accuracy : ', rmsle(y_pred, np.ravel(labels_train))
print 'Model Report ********'
```
### 3. Gradient Boosting
```
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
from matplotlib.pylab import rcParams
rcParams['figure.figsize'] = 12, 4
from sklearn import metrics
def modelfit(alg, Xtrain, ytrain, useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(Xtrain, label=ytrain)
print alg.get_params()['n_estimators']
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round = alg.get_params()['n_estimators'], early_stopping_rounds=early_stopping_rounds)
alg.set_params(n_estimators=cvresult.shape[0])
alg.fit(Xtrain, ytrain, eval_metric='auc')
predict = alg.predict(Xtrain)
return predict
```
## Step 1 Fix learning rate and number of estimators for tuning tree-based parameters
```
xgb1 = XGBClassifier(
learning_rate =0.05,
n_estimators=100,
max_depth=15,
min_child_weight=4,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'reg:linear',
scale_pos_weight=1,
seed=27)
predict = modelfit(xgb1, features_train, np.ravel(labels_train))
#print model report:
print '\nModel Report ********'
print "Accuracy : %.4g" % rmsle(np.ravel(labels_train), predict)
print '\nModel Report ********'
feat_imp = pd.Series(xgb1.booster().get_fscore()).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
```
## Step 2: Tune max_depth and min_child_weight
```
from sklearn.grid_search import GridSearchCV
param_test1 = {
'max_depth':range(3,10,2),
'min_child_weight':range(1,6,2)
}
gsearch1 = GridSearchCV(estimator = XGBClassifier( learning_rate =0.1, n_estimators=100, max_depth=5, min_child_weight=1, gamma=0, subsample=0.8, colsample_bytree=0.8, scale_pos_weight=1, seed=27), param_grid = param_test1, scoring='roc_auc', n_jobs=4,iid=False)
gsearch1.fit(features_train,np.ravel(labels_train))
gsearch1.grid_scores_, gsearch1.best_params_, gsearch1.best_score_
```
## Data Cleaning
There are duplicate client ids in cliente_table, which means one client id may have multiple client name that are very similar. We will cluster them based on a hash function and use a clustering algorithm to evaluate similarity.
```
import re
def hash_eval(s):
hash_base = 4
s = re.sub('[., ]', '', s)
seqlen = len(s)
n = seqlen - 1
h = 0
for c in s:
h += ord(c) * (hash_base ** n)
n -= 1
curhash = h
return curhash
# In the client table, same clients are assigned different client ID. We create a new client table where clients are assigned unique ID.
clientid_hash = dict()
new_client_id = [-1]
for idx, s in enumerate(clientnameid_data.NombreCliente):
t = hash_eval(s)
clientid_hash.setdefault(t, []).append(clientnameid_data.Cliente_ID[idx])
if t in clientid_hash:
a = clientid_hash[t]
new_client_id.append(a[0])
# In the agency table, same agencies (town, state) are assigned different agency ID. We create a new agency table where agencies (town, state) are assigned unique ID.
agencyid_hash = dict()
new_agency_id = [-1]
for idx, s in enumerate(townstate_data.Town+townstate_data.State):
t = hash_eval(s)
agencyid_hash.setdefault(t, []).append(townstate_data.Agencia_ID[idx])
if t in agencyid_hash:
a = agencyid_hash[t]
new_agency_id.append(a[0])
clientnameid_data['New_Cliente_ID'] = new_client_id[1:]
townstate_data['New_Agencia_ID'] = new_agency_id[1:]
print clientnameid_data.head(10)
print '---'
print townstate_data.head()
print '---'
print train_data_samp.head(10)
print train_data_samp.head(10)
print '------'
for idx, cid in enumerate(train_data_samp.client_id):
train_data_samp.client_id.values[idx] = clientnameid_data.New_Cliente_ID[train_data_samp.client_id.values[idx] == clientnameid_data.Cliente_ID.values].values[0]
train_data_samp.sales_depot_id.values[idx] = townstate_data.New_Agencia_ID[train_data_samp.sales_depot_id.values[idx] == townstate_data.Agencia_ID.values].values[0]
print '-----'
print train_data_samp.head()
```
## Load Test Data
```
test_data = pd.read_csv('test.csv')
test_data.columns = ['id', 'week_num', 'sales_depot_id', 'sales_chan_id', 'route_id', 'client id', 'prod_id']
test_labels = pd.read_csv('sample_submission.csv')
test_data = test_data.drop('id', 1)
print test_data.head()
g = sns.PairGrid(data_t)
g.map(plt.scatter)
a = [[1, 2, 3, 4]]
print a
np.array(a)
print np.array(a)
a = np.array(a)
a = sp_randint(10,2)
range(3,10,2)
sp_randint(1, 6)
import subprocess
subprocess.call(['ec2kill'])
from subprocess import call
call(["ec2-terminate-instances", "i-308b33ed "])
# Utility function to report best scores
def report(grid_scores, n_top=3):
top_scores = sorted(grid_scores, key=itemgetter(1), reverse=True)[:n_top]
for i, score in enumerate(top_scores):
print("Model with rank: {0}".format(i + 1))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
score.mean_validation_score,
np.std(score.cv_validation_scores)))
print("Parameters: {0}".format(score.parameters))
print("")
from sklearn.ensemble import RandomForestClassifier
from sklearn.grid_search import RandomizedSearchCV
from scipy.stats import randint as sp_randint
from operator import itemgetter
clf = RandomForestClassifier(n_estimators=30)
# specify parameters and distributions to sample from
param_dist = {"max_depth": [10, None],
"max_features": sp_randint(1, 6),
"min_samples_split": sp_randint(1, 6),
"min_samples_leaf": sp_randint(1, 6),
"bootstrap": [True, False],
"criterion": ["gini", "entropy"]}
# run randomized search
n_iter_search = 20
random_search = RandomizedSearchCV(clf, param_distributions=param_dist,
n_iter=n_iter_search, n_jobs=4, cv=3)
start = time.time()
random_search.fit(features_train, np.ravel(labels_train))
print("RandomizedSearchCV took %.2f seconds for %d candidates"
" parameter settings." % ((time.time() - start), n_iter_search))
report(random_search.grid_scores_)
print random_search.best_score_
print random_search.best_estimator_
```
| github_jupyter |
# Graphics using Seaborn
We previously have covered how to do some basic graphics using `matplotlib`. In this notebook we introduce a package called `seaborn`. `seaborn` builds on top of `matplotlib` by doing 2 things:
1. Gives us access to more types of plots (Note: Every plot created in `seaborn` could be made by `matplotlib`, but you shouldn't have to worry about doing this)
2. Sets better defaults for how the plot looks right away
Before we start, make sure that you have `seaborn` installed. If not, then you can install it by
```
conda install seaborn
```
_This notebook was created by Dave Backus, Chase Coleman, and Spencer Lyon for the NYU Stern course [Data Bootcamp](http://databootcamp.nyuecon.com/)._
```
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import sys
%matplotlib inline
```
As per usual, we begin by listing the versions of each package that is used in this notebook.
```
# check versions (overkill, but why not?)
print('Python version:', sys.version)
print('Pandas version: ', pd.__version__)
print('Matplotlib version: ', mpl.__version__)
print('Seaborn version: ', sns.__version__)
```
## Datasets
There are some classical datasets that get used to demonstrate different types of plots. We will use several of them here.
* tips : This dataset has informaiton on waiter tips. Includes information such as total amount of the bill, tip amount, sex of waiter, what day of the week, which meal, and party size.
* anscombe: This dataset is a contrived example. It has 4 examples which differ drastically when you look at them, but they have the same correlation, regression coefficient, and $R^2$.
* titanic : This dataset has information on each of the passengers who were on the titanic. Includes information such as: sex, age, ticket class, fare paid, whether they were alone, and more.
```
tips = sns.load_dataset("tips")
ansc = sns.load_dataset("anscombe")
tita = sns.load_dataset("titanic")
```
# Better Defaults
Recall that in our [previous notebook](bootcamp_graphics.ipynb) that we used `plt.style.use` to set styles. We will begin by setting the style to `"classic"`; this sets all of our default settings back to `matplotlib`'s default values.
Below we plot open and closing prices on the top axis and the implied returns on the bottom axis.
```
plt.style.use("classic")
fig, ax = plt.subplots(2)
tips[tips["sex"] == "Male"].plot(x="total_bill", y="tip", ax=ax[0], kind="scatter",
color="blue")
tips[tips["sex"] == "Female"].plot(x="total_bill", y="tip", ax=ax[1], kind="scatter",
color="#F52887")
ax[0].set_xlim(0, 60)
ax[1].set_xlim(0, 60)
ax[0].set_ylim(0, 15)
ax[1].set_ylim(0, 15)
ax[0].set_title("Male Tips")
ax[1].set_title("Female Tips")
fig.tight_layout()
# fig.savefig("/home/chase/Desktop/foo.png")
# sns.set() resets default seaborn settings
sns.set()
fig, ax = plt.subplots(2)
tips[tips["sex"] == "Male"].plot(x="total_bill", y="tip", ax=ax[0], kind="scatter",
color="blue")
tips[tips["sex"] == "Female"].plot(x="total_bill", y="tip", ax=ax[1], kind="scatter",
color="#F52887")
ax[0].set_xlim(0, 60)
ax[1].set_xlim(0, 60)
ax[0].set_ylim(0, 15)
ax[1].set_ylim(0, 15)
ax[0].set_title("Male Tips")
ax[1].set_title("Female Tips")
fig.tight_layout()
```
What did you notice about the differences in the settings of the plot?
Which do you like better? We like the second better.
Investigate other styles and create the same plot as above using a style you like. You can choose from the list in the code below.
If you have additional time, visit the [seaborn docs](http://stanford.edu/~mwaskom/software/seaborn/tutorial/aesthetics.html#temporarily-setting-figure-style) and try changing other default settings.
```
plt.style.available
```
We could do the same for a different style (like `ggplot`)
```
plt.style.use("ggplot")
fig, ax = plt.subplots(2)
tips[tips["sex"] == "Male"].plot(x="total_bill", y="tip", ax=ax[0], kind="scatter",
color="blue")
tips[tips["sex"] == "Female"].plot(x="total_bill", y="tip", ax=ax[1], kind="scatter",
color="#F52887")
ax[0].set_xlim(0, 60)
ax[1].set_xlim(0, 60)
ax[0].set_ylim(0, 15)
ax[1].set_ylim(0, 15)
ax[0].set_title("Male Tips")
ax[1].set_title("Female Tips")
fig.tight_layout()
```
**Exercise**: Find a style you like and recreate the plot above using that style.
# The Juicy Stuff
While having `seaborn` set sensible defaults is convenient, it isn't a particularly large innovation. We could choose sensible defaults and set them to be our default. The main benefit of `seaborn` is the types of graphs that it gives you access to -- All of which could be done in `matplotlib`, but, instead of 5 lines of code, it would require possibly hundreds of lines of code. Trust us... This is a good thing.
We don't have time to cover everything that can be done in `seaborn`, but we suggest having a look at the [gallery](http://stanford.edu/~mwaskom/software/seaborn/examples/index.html) of examples.
We will cover:
* `kdeplot`
* `jointplot`
* `violinplot`
* `pairplot`
* ...
```
# Move back to seaborn defaults
sns.set()
```
## kdeplot
What does kde stand for?
kde stands for "kernel density estimation." This is (far far far) beyond the scope of this class, but the basic idea is that this is a smoothed histogram. When we are trying to get information about distributions it sometimes looks nicer than a histogram does.
```
fig, ax = plt.subplots()
ax.hist(tips["tip"], bins=25)
ax.set_title("Histogram of tips")
plt.show()
fig, ax = plt.subplots()
sns.kdeplot(tips["tip"], ax=ax)
ax.hist(tips["tip"], bins=25, alpha=0.25, normed=True, label="tip")
ax.legend()
fig.suptitle("Kernel Density with Histogram")
plt.show()
```
**Exercise**: Create your own kernel density plot using `sns.kdeplot` of `"total_bill"` from the `tips` dataframe
## Jointplot
We now show what `jointplot` does. It draws a scatter plot of two variables and puts their histogram just outside of the scatter plot. This tells you information about not only the joint distribution, but also the marginals.
```
sns.jointplot(x="total_bill", y="tip", data=tips)
```
We can also plot everything as a kernel density estimate -- Notice the main plot is now a contour map.
```
sns.jointplot(x="total_bill", y="tip", data=tips, kind="kde")
```
**Exercise**: Create your own `jointplot`. Feel free to choose your own x and y data (if you can't decide then use `x=size` and `y=tip`). Interpret the output of the plot.
## violinplot
Some of the story of this notebook is that distributions matter and how we can show them. Violin plots are similar to a sideways kernel density and it allows us to look at how distributions matter over some aspect of the data.
```
tita.head()
tips.head()
sns.violinplot(x="class", y="age", data=tita)
```
**Exercise**: We might also want to look at the distribution of prices across ticket classes. Make a violin plot of the prices over the different ticket classes.
## Pairplot
Pair plots show us two things. They show us the histograms of the variables along the diagonal and then the scatter plot of each pair of variables on the off diagonal pictures.
Why might this be useful? It allows us to look get an idea of the correlations across each pair of variables and gives us an idea of their relationships across the variables.
```
sns.pairplot(tips[["tip", "total_bill", "size"]], size=3.5)
```
Below is the same plot, but slightly different. What is different?
```
sns.pairplot(tips[["tip", "total_bill", "size"]], size=3.5, diag_kind="kde")
```
What's different about this plot?
Different colors for each company.
```
tips.head()
sns.pairplot(tips[["tip", "total_bill", "size", "time"]], size=3.5, diag_kind="kde",
hue="time")
```
## swarmplot
Sometimes we simply have too much data. One approach to visualizing all the data is to adjust features like the point size or transparency.
An alternative is to use a swarm plot. This is best understood by example, so let's dive in!
```
fig, ax = plt.subplots()
sns.swarmplot(data=tips, x="day", y="total_bill")
```
## lmplot
We often want to think about running regressions of variables. A statistician named Francis Anscombe came up with four datasets that:
* Same mean for $x$ and $y$
* Same variance for $x$ and $y$
* Same correlation between $x$ and $y$
* Same regression coefficient of $x$ on $y$
Below we show the scatter plot of the datasets to give you an idea of how different they are.
```
fig, ax = plt.subplots(2, 2, figsize=(10, 9))
ansc[ansc["dataset"] == "I"].plot.scatter(x="x", y="y", ax=ax[0, 0])
ansc[ansc["dataset"] == "II"].plot.scatter(x="x", y="y", ax=ax[0, 1])
ansc[ansc["dataset"] == "III"].plot.scatter(x="x", y="y", ax=ax[1, 0])
ansc[ansc["dataset"] == "IV"].plot.scatter(x="x", y="y", ax=ax[1, 1])
ax[0, 0].set_title("Dataset I")
ax[0, 1].set_title("Dataset II")
ax[1, 0].set_title("Dataset III")
ax[1, 1].set_title("Dataset IV")
fig.suptitle("Anscombe's Quartet")
```
`lmplot` plots the data with the regression coefficient through it.
```
sns.lmplot(x="x", y="y", data=ansc, col="dataset", hue="dataset",
col_wrap=2, ci=None)
```
## regplot
`regplot` also shows the regression line through data points
```
sns.regplot(x="x", y="y", data=ansc[ansc["dataset"] == "I"])
```
| github_jupyter |
# Precision-Recall
# Question 1
<img src="images/lec6_quiz01_pic01.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
** Answer **
Recall = 5600 /(5600 + 40) = 0.99
# Question 2
<img src="images/lec6_quiz01_pic02.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
** Answer **
Accuracy = (5600 + 2460)/(5600 + 2460 + 40 + 1900) = 0.8
# Question 3
<img src="images/lec6_quiz01_pic03.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
# Question 4
<img src="images/lec6_quiz01_pic04.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
** Answer **
- Precision: 5600/float(5600 + 1900)= 0.75
- Recall = 5600 /(5600 + 40) = 0.99
# Question 5
<img src="images/lec6_quiz01_pic05.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
# Question 6
<img src="images/lec6_quiz01_pic06.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
**More info: https://www.coursera.org/learn/ml-classification/lecture/IMHs2/trading-off-precision-and-recall**
# Question 7
<img src="images/lec6_quiz01_pic07.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
# Question 8
<img src="images/lec6_quiz01_pic08.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
# Question 9
<img src="images/lec6_quiz01_pic09.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-classification/exam/pGfWZ/precision-recall)*
<!--TEASER_END-->
**Answer**
- Notice that class probability =/= score. In the context of linear classifier, score is the dot product of coefficieints and features.
- Recall that **P(y = +1 | x,w) = sigmoid(score)**. If we want **P(y=+1|x,w)** to be greater than 0.9, how large should the score be?
$\large \frac{1}{1 + e^{-score}} = 0.9$
$=> \large 0.9 + 0.9 e^{-score} = 1$
$=>\large \frac{0.1}{0.9} = e^{-score}$
$=>\large \ln(\frac{0.1}{0.9}) = \ln(e^{-score})$
$=>\large score = 2.20$
| github_jupyter |
# Anomaly Detection Using RNN
Anomaly detection detects data points in data that does not fit well with the rest of data. In this notebook we demonstrate how to do unsupervised anomaly detection using recurrent nueral network (RNN) model.
We used one of the dataset in Numenta Anomaly Benchmark (NAB) ([link](https://github.com/numenta/NAB)) for demo, i.e. NYC taxi passengers dataset, which contains 10320 records, each indicating the total number of taxi passengers in NYC at a corresonponding time spot. We use RNN to learn from 50 previous values, and predict just the 1 next value. The data points whose actual values are distant from predicted values are considered anomalies (distance threshold can be adjusted as needed).
References:
* Unsupervised real-time anomaly detection for streaming data ([link](https://www.sciencedirect.com/science/article/pii/S0925231217309864)).
## Intialization
* import necesary libraries
```
import os
import pandas as pd
import numpy as np
import matplotlib
matplotlib.use('Agg')
%pylab inline
import seaborn
import matplotlib.dates as md
from matplotlib import pyplot as plt
from sklearn import preprocessing
```
* import necessary modules
```
from zoo.pipeline.api.keras.layers import Dense, Dropout, LSTM
from zoo.pipeline.api.keras.models import Sequential
```
## Data Check
* read data
```
try:
dataset_path = os.getenv("ANALYTICS_ZOO_HOME")+"/bin/data/NAB/nyc_taxi/nyc_taxi.csv"
df = pd.read_csv(dataset_path)
except Exception as e:
print("nyc_taxi.csv doesn't exist")
print("you can run $ANALYTICS_ZOO_HOME/bin/data/NAB/nyc_taxi/get_nyc_taxi.sh to download nyc_taxi.csv")
```
* Understand the data.
Each record is in format of (timestamp, value). Timestamps range between 2014-07-01 and 2015-01-31.
```
print(df.info())
# check the timestamp format and frequence
print(df['timestamp'].head(10))
# check the mean of passenger number
print(df['value'].mean())
# change the type of timestamp column for plotting
df['datetime'] = pd.to_datetime(df['timestamp'])
# visualisation of anomaly throughout time (viz 1)
fig, ax = plt.subplots(figsize=(12, 5))
ax.plot(df['datetime'], df['value'], color='blue', linewidth=0.6)
ax.set_title('NYC taxi passengers throughout time')
plt.xlabel('datetime')
plt.xticks(rotation=45)
plt.ylabel('The Number of NYC taxi passengers')
plt.legend(loc='upper left')
plt.show()
```
## Feature engineering
* Extracting some useful features
```
# the hours when people are awake (6:00-00:00)
df['hours'] = df['datetime'].dt.hour
df['awake'] = (((df['hours'] >= 6) & (df['hours'] <= 23)) | (df['hours'] == 0)).astype(int)
# creation of 2 distinct categories that seem useful (sleeping time and awake time)
df['categories'] = df['awake']
a = df.loc[df['categories'] == 0, 'value']
b = df.loc[df['categories'] == 1, 'value']
fig, ax = plt.subplots()
a_heights, a_bins = np.histogram(a)
b_heights, b_bins = np.histogram(b, bins=a_bins)
width = (a_bins[1] - a_bins[0])/6
ax.bar(a_bins[:-1], a_heights*100/a.count(), width=width, facecolor='yellow', label='Sleeping time')
ax.bar(b_bins[:-1]+width, (b_heights*100/b.count()), width=width, facecolor='red', label ='Awake time')
ax.set_title('Histogram of NYC taxi passengers in different categories')
plt.xlabel('The number of NYC taxi passengers')
plt.ylabel('Record counts')
plt.legend()
plt.show()
df['awake'].head(4)
df['timestamp'].head(4)
```
From the above result, we can conclude:
- more people take taxi when they are awake
## Data Preparation
* Standardizing data and spliting them into the train data and the test data
```
#select and standardize data
data_n = df[['value', 'hours', 'awake']]
min_max_scaler = preprocessing.StandardScaler()
np_scaled = min_max_scaler.fit_transform(data_n)
data_n = pd.DataFrame(np_scaled)
#important parameters and train/test size
prediction_time = 1
testdatasize = 1000
unroll_length = 50
testdatacut = testdatasize + unroll_length + 1
#train data
x_train = data_n[0:-prediction_time-testdatacut].as_matrix()
y_train = data_n[prediction_time:-testdatacut ][0].as_matrix()
#test data
x_test = data_n[0-testdatacut:-prediction_time].as_matrix()
y_test = data_n[prediction_time-testdatacut: ][0].as_matrix()
```
* Unroll data: for the data point at index i, create a sequence from i to (i+unroll_length)
for example, if unroll_length=5
[[1]
[2]
[3]
[4]
[5]
[6]
[7]
[8]
[9]
[10]
...
]
will be unrolled to create following sequences
[[ 1, 2, 3, 4, 5]
[ 2, 3, 4, 5, 6]
[ 3, 4, 5, 6, 7]
[ 4, 5, 6, 7, 8]
[ 5, 6, 7, 8, 9]
[ 6, 7, 8, 9, 10]
...
]
```
#unroll: create sequence of 50 previous data points for each data points
def unroll(data,sequence_length=24):
result = []
for index in range(len(data) - sequence_length):
result.append(data[index: index + sequence_length])
return np.asarray(result)
# adapt the datasets for the sequence data shape
x_train = unroll(x_train,unroll_length)
x_test = unroll(x_test,unroll_length)
y_train = y_train[-x_train.shape[0]:]
y_test = y_test[-x_test.shape[0]:]
# see the shape
print("x_train", x_train.shape)
print("y_train", y_train.shape)
print("x_test", x_test.shape)
print("y_test", y_test.shape)
```
## Build Model
* Here we show an example of building a RNN network using Analytics Zoo Keras-Style API.
There are three LSTM layers and one Dense layer.
```
# Build the model
model = Sequential()
model.add(LSTM(
input_shape=(x_train.shape[1], x_train.shape[-1]),
output_dim=20,
return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(
10,
return_sequences=False))
model.add(Dropout(0.2))
model.add(Dense(
output_dim=1))
model.compile(loss='mse', optimizer='rmsprop')
```
## Train the model
```
%%time
# Train the model
print("Training begins.")
model.fit(
x_train,
y_train,
batch_size=1024,
nb_epoch=20)
print("Training completed.")
```
## Prediction
* BigDL models make inferences based on the given data using model.predict(val_rdd) API. A result of RDD is returned. predict_class returns the predicted points.
```
# create the list of difference between prediction and test data
diff=[]
ratio=[]
predictions = model.predict(x_test)
p = predictions.collect()
for u in range(len(y_test)):
pr = p[u][0]
ratio.append((y_test[u]/pr)-1)
diff.append(abs(y_test[u]- pr))
```
## Evaluation
* plot the prediction and the reality
```
# plot the predicted values and actual values (for the test data)
fig, axs = plt.subplots()
axs.plot(p,color='red', label='predicted values')
axs.plot(y_test,color='blue', label='actual values')
axs.set_title('the predicted values and actual values (for the test data)')
plt.xlabel('test data index')
plt.ylabel('number of taxi passengers after min_max_scalar')
plt.legend(loc='upper left')
plt.show()
```
* Set the distance thresold for anomalies. There're many ways to select this threshold. Here we set the expected proportion of anomalies among the entire set. Then we set the threshold as the minimum value of top N distances (here N is the total number of anomalies, i.e. anomaly fraction * total no. of samples)
```
# An estimation of anomly population of the dataset
outliers_fraction = 0.01
# select the most distant prediction/reality data points as anomalies
diff = pd.Series(diff)
number_of_outliers = int(outliers_fraction*len(diff))
threshold = diff.nlargest(number_of_outliers).min()
```
* plot anomalies in the test data throughout time
```
# plot the difference and the threshold (for the test data)
fig, axs = plt.subplots()
axs.plot(diff,color='blue', label='diff')
axs.set_title('the difference between the predicted values and actual values with the threshold line')
plt.hlines(threshold, 0, 1000, color='red', label='threshold')
plt.xlabel('test data index')
plt.ylabel('difference value after min_max_scalar')
plt.legend(loc='upper left')
plt.show()
# data with anomaly label (test data part)
test = (diff >= threshold).astype(int)
# the training data part where we didn't predict anything (overfitting possible): no anomaly
complement = pd.Series(0, index=np.arange(len(data_n)-testdatasize))
last_train_data= (df['datetime'].tolist())[-testdatasize]
# add the data to the main
df['anomaly27'] = complement.append(test, ignore_index='True')
```
* plot anomalies in the test data throughout time
```
# visualisation of anomaly throughout time (viz 1)
fig, ax = plt.subplots(figsize=(12, 5))
a = df.loc[df['anomaly27'] == 1, ['datetime', 'value']] #anomaly
ax.plot(df['datetime'], df['value'], color='blue', label='no anomaly value', linewidth=0.6)
ax.scatter(a['datetime'].tolist(),a['value'], color='red', label='anomalies value')
ax.set_title('the number of nyc taxi value throughout time (with anomalies scattered)')
max_value = df['value'].max()
min_value = df['value'].min()
plt.vlines(last_train_data, min_value, max_value, color='black', linestyles = "dashed", label='test begins')
plt.xlabel('datetime')
plt.xticks(rotation=45)
plt.ylabel('the number of nyc taxi value')
plt.legend(loc='upper left')
plt.show()
```
| github_jupyter |
# nsopy: basic usage examples
Generally, the inputs required are
* a first-order oracle of the problem: for a given $x_k \in \mathbb{X} \subseteq \mathbb{R}^n$, it returns $f(x_k)$ and a valid subgradient $\nabla f(x_k)$,
* the projection function $\Pi_{\mathbb{X}}: \mathbb{R}^n \rightarrow \mathbb{R}^n$.
## Example 1: simple analytical example
Consider the following problem from [1, Sec. 2.1.3]:
$$
\begin{array}{ll}
\min & f(x_1, x_2) = \left\{ \begin{array}{ll}
5(9x_1^2 + 16x_2^2)^{1/2} & \mathrm{if \ } x_1 > \left|x_2\right|, \\
9x_1 + 16\left|x_2\right| & \mathrm{if \ } x_1 \leq \left|x_2\right| \\
\end{array} \right.\\
\mathrm{s.t.} & -3 \leq x_1, x_2 \leq 3.
\end{array}
$$

This problem is interesting because a common gradient algorithm with backtracking initialized anywhere where in the set
$\left\{(x_1,x_2) \left| \ x_1 > \left|x_2\right| > (9/16)^2\left|x_1\right| \right. \right\}$
fails to converge to the optimum (-3,0), by remaining stuck at (0,0), even though it never touches any point where the function is nondifferentiable, see discussion in [1, Sec. 2.1.3]. Here we test our methods on this problem.
We write the oracle and projection as
```
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
%matplotlib notebook
%cd ..
```
**Important Note**: all methods have been devised to solve the maximization of concave functions. To minimize (as in this case), we just need to negate the oracle's returns, i.e., the objective value $f(x_k)$ and the subgradient $\nabla f(x_k)$.
```
def oracle(x):
assert -3 <= x[0] <= 3, 'oracle must be queried within X'
assert -3 <= x[1] <= 3, 'oracle must be queried within X'
# compute function value a subgradient
if x[0] > abs(x[1]):
f_x = 5*(9*x[0]**2 + 16*x[1]**2)**(float(1)/float(2))
diff_f_x = np.array([float(9*5*x[0])/np.sqrt(9*x[0]**2 + 16*x[1]**2),
float(16*5*x[0])/np.sqrt(9*x[0]**2 + 16*x[1]**2)])
else:
f_x = 9*x[0] + 16*abs(x[1])
if x[1] >= 0:
diff_f_x = np.array([9, 16], dtype=float)
else:
diff_f_x = np.array([9, -16], dtype=float)
return 0, -f_x, -diff_f_x # return negation to minimize
def projection_function(x):
# projection on the box is simply saturating the entries
return np.array([min(max(x[0],-3),3), min(max(x[1],-3),3)])
```
We can now solve it by applying one of the several methods available:
**Note**: try to change the method and see for yourself how their trajectories differ!
```
from nsopy import SGMDoubleSimpleAveraging as DSA
from nsopy import SGMTripleAveraging as TA
from nsopy import SubgradientMethod as SG
from nsopy import UniversalPGM as UPGM
from nsopy import UniversalDGM as UDGM
from nsopy import UniversalFGM as UFGM
from nsopy import GenericDualMethodLogger
# method = DSA(oracle, projection_function, dimension=2, gamma=0.5)
# method = TA(oracle, projection_function, dimension=2, variant=2, gamma=0.5)
# method = SG(oracle, projection_function, dimension=2)
method = UPGM(oracle, projection_function, dimension=2, epsilon=10, averaging=True)
# method = UDGM(oracle, projection_function, dimension=2, epsilon=1.0)
# method = UFGM(oracle, projection_function, dimension=2, epsilon=1.0)
method_logger = GenericDualMethodLogger(method)
# start from an different initial point
x_0 = np.array([2.01,2.01])
method.lambda_hat_k = x_0
for iteration in range(100):
method.dual_step()
```
And finally plot the result:
```
box = np.linspace(-3, 3, 31)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
ax.plot_trisurf(np.array([x_1 for x_1 in box for x_2 in box]),
np.array([x_2 for x_1 in box for x_2 in box]),
np.array([-oracle([x_1, x_2])[1] for x_1 in box for x_2 in box]))
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_zlabel('$f(x)$')
plt.plot([x[0] for x in method_logger.lambda_k_iterates],
[x[1] for x in method_logger.lambda_k_iterates],
[-f_x for f_x in method_logger.d_k_iterates], 'r.-')
```
### References
[1] Dimitri Bertsekas, Convex Optimization Algorithms, Athena Scientific Belmont, 2015.
| github_jupyter |
```
# (C) Copyright 1996- ECMWF.
#
# This software is licensed under the terms of the Apache Licence Version 2.0
# which can be obtained at http://www.apache.org/licenses/LICENSE-2.0.
# In applying this licence, ECMWF does not waive the privileges and immunities
# granted to it by virtue of its status as an intergovernmental organisation
# nor does it submit to any jurisdiction.
```
### SOS
This script takes too much time (more than 1 week if run on personal computer). It is recommended to run it in cluster, and preferably in a different session for each lead time, so multiple sessions can run in parallel.
```
import xarray as xr
import pandas as pd
import numpy as np
from itertools import product
from pathlib import Path
import multiprocessing # parallel processing
import tqdm # timing
import sys
Area_used = [48, -10, 27, 41]
P_used = [90, 95, 99] # Thresholds for EPEs [90, 95, 99]
offset_days = 15 # offset days used for getting the EPEs reference climatology of occurrence
bootstraps = 1000
input_dir = ''
output_dir = '/ProcessedData/ForecastsEPEs_Analysis/'
Path(output_dir).mkdir(parents=True, exist_ok=True) # generate subfolder for storing the results
ActualClusters = pd.read_csv(input_dir+'ProcessedData/PatternAllocations_ERA5.csv', index_col=0)
ActualClusters.index = pd.to_datetime(ActualClusters.index)
n_clusters = len(ActualClusters.Label.unique())
AllocatedClusters = pd.read_csv(input_dir+'ProcessedData/ForecastsClusterAllocations.csv').iloc[:, 1:]
AllocatedClusters[['time', 'valid_time']] = AllocatedClusters[['time', 'valid_time']].apply(pd.to_datetime)
# indices for start and end of Summer Half (Summer Half between 16th April - 15th October, inclusive of both dates)
Sorted_Dates = np.array(pd.date_range('20040101', '20041231').strftime('%m%d')) # a leap year for getting all dates
StartSummerHalf = np.where(Sorted_Dates=='0416')[0]
EndSummerHalf = np.where(Sorted_Dates=='1015')[0]
def temp_flagging(valid_dates, temp_subset):
valid_dates = pd.to_datetime(valid_dates)
if temp_subset == 'All':
temporal_flag = ['All']*len(valid_dates)
elif temp_subset == 'HalfYear':
temporal_flag_aux = pd.Series([i[-4:] for i in valid_dates.strftime('%Y%m%d')])
temporal_flag_aux = temporal_flag_aux.map({i: i_c for i_c, i in enumerate(Sorted_Dates)})
temporal_flag_aux = temporal_flag_aux.values
temporal_flag = np.repeat(['WinterHalf'], len(temporal_flag_aux))
temporal_flag[(temporal_flag_aux>=StartSummerHalf) & (temporal_flag_aux<=EndSummerHalf)] = 'SummerHalf'
elif temp_subset == 'Season':
temporal_flag = (valid_dates.month%12 + 3)//3
temporal_flag = temporal_flag.map({1: 'Winter', 2: 'Spring', 3: 'Summer', 4: 'Autumn'})
elif temp_subset == 'Month':
temporal_flag = valid_dates.month.astype(str)
elif temp_subset == 'DayMonth':
temporal_flag = pd.Series([i[-4:] for i in valid_dates.strftime('%Y%m%d')])
temporal_flag = temporal_flag.values
return temporal_flag
for i_tag in ['All', 'HalfYear', 'Season', 'Month', 'DayMonth']:
ActualClusters[i_tag] = temp_flagging(ActualClusters.index, i_tag)
del(i_tag)
# read ERA5 rainfall data
Precipitation = xr.open_dataarray(input_dir+'Data/ERA5/D1_Total_Precipitation.grb', engine='cfgrib')
Precipitation = Precipitation.reset_coords(drop=True)
dates = pd.to_datetime(Precipitation.time.values) # get dates
dates = pd.to_datetime(dates.strftime('%Y%m%d')) # convert to 00:00 hour of day
Precipitation = Precipitation.assign_coords({'time': dates})
Precipitation = Precipitation.sel(longitude=slice(Area_used[1], Area_used[3]),
latitude=slice(Area_used[0], Area_used[2]),
time=slice('1979', '2020')) # keep only full years 1979-2020
precip_dates = Precipitation.time.values # get precipitation dates
Lons_all = Precipitation.longitude.values # get longitudes, since due to memory limititations subsets are needed
del(dates)
# function for getting subset of precipitation data and generating boolean for exceedance of extremes
def exceed_boolean(data):
Quant = data.quantile(np.array(P_used)/100, interpolation='linear', dim='time', keep_attrs=True) # thresholds
Quant = Quant.rename({'quantile': 'percentile'}) # rename coordinate
Quant = Quant.assign_coords({'percentile': P_used}) # assign the dim values based on percentiles
# boolean xarray for identifying if an event is over the threshold
Exceed_xr = [data>Quant.sel(percentile=i_p) for i_p in P_used] # boolean of exceedance per percentile
Exceed_xr = xr.concat(Exceed_xr, dim=pd.Index(P_used, name='percentile')) # concatenate data for all percentiles
return Exceed_xr
# function for calculating "statistical" Brier Score based on conditional probabilities of EPEs at subsets
def statistical_brier_score(cond_probs, weights, dim_used='subset'):
weights = np.array(weights)
brier_score = cond_probs - cond_probs**2 # brier score for climatological probabilities
if len(weights)>1:
brier_score = brier_score.rename({dim_used: 'subsetting'})
brier_score = [brier_score.isel(subsetting=i)*weights[i] for i in range(len(weights))]
brier_score = xr.concat(brier_score, dim='subsetting').sum(dim='subsetting')/weights.sum()
return brier_score.astype('float32')
# function for calculating conditional probabilities and relevant Brier Score for DayMonth temporal subsetting
def DayMonth_EPEs_conditioning(subset_extremes):
nw_crd = ActualClusters.loc[subset_extremes.time.values, 'DayMonth'] # temporal flag to replace coordinate values
exceed_flags = subset_extremes.assign_coords({'time': nw_crd.values}) # rename time based on temporal flag
ConnProb = []
for i_dates_central in Sorted_Dates:
central_loc = np.where(Sorted_Dates==i_dates_central)[0]
dates_check_all = np.linspace(central_loc-offset_days, central_loc+offset_days, 2*offset_days+1)
for loc, i_date_loc in enumerate(dates_check_all):
if i_date_loc >= len(Sorted_Dates):
dates_check_all[loc] = dates_check_all[loc] - len(Sorted_Dates)
dates_check_all = np.take(Sorted_Dates, dates_check_all.astype(int)).flatten()
Kept_dates_locs = [np.where(nw_crd.values==i)[0].tolist() for i in dates_check_all]
Kept_dates_locs = np.array([j for i in Kept_dates_locs for j in i])
i_condprob = exceed_flags.isel(time=Kept_dates_locs).sum('time')/len(Kept_dates_locs)
i_condprob = i_condprob.assign_coords({'temporal': i_dates_central})
ConnProb.append(i_condprob)
ConnProb = xr.concat(ConnProb, dim='temporal')
weights_temp = nw_crd.value_counts() # weights based on occurrence of temporal subsets
weights_temp = weights_temp.reindex(Sorted_Dates).fillna(0) # reorder to same order as the xarray "ConnProb"
BS = statistical_brier_score(ConnProb, weights_temp.values, dim_used='temporal') # statistical Brier Score
BS = BS.assign_coords({'Method': 'DayMonth_Temp'}) # assign new coord with the temporal subsetting info
return (ConnProb.astype('float32'), BS.astype('float32'))
# function for calculating conditional probabilities and relevant Brier Score for specific temporal subsetting
def temporal_conditioning_subset(subset_extremes, subset_type):
nw_crd = ActualClusters.loc[subset_extremes.time.values, subset_type] # temporal flag to replace coordinate values
ConnProb = subset_extremes.assign_coords({'time': nw_crd.values}) # rename time based on temporal flag
ConnProb = ConnProb.groupby('time').sum('time')/ConnProb.groupby('time').count() # get conditional prob
ConnProb = ConnProb.rename({'time': 'temporal'}) # rename coordinate
weights_temp = nw_crd.value_counts() # weights based on occurrence of temporal subsets
weights_temp = weights_temp.reindex(ConnProb.temporal.values) # reorder to same order as the xarray "ConnProb"
BS = statistical_brier_score(ConnProb, weights_temp.values, dim_used='temporal') # statistical Brier Score
BS = BS.assign_coords({'Method': f'{subset_type}_Temp'}) # assign new coord with the temporal subset info
return (ConnProb.astype('float32'), BS.astype('float32'))
# function for calculating conditional probabilities and relevant Brier Score for specific pattern-temporal subsetting
def clusters_EPEs_conditioning(subset_extremes, subset_type):
nw_crd = subset_extremes.time.values # cluster ID
# generate new coordinates values based on the cluster ID and the temporal flag of interest for each instance
nw_crd = ActualClusters.loc[nw_crd, 'Label'].astype(str) + '-' + ActualClusters.loc[nw_crd, subset_type]
DataUsed = subset_extremes.assign_coords({'time': nw_crd.values}) # new coordinate values
DataUsed = DataUsed.groupby('time').sum('time')/DataUsed.groupby('time').count() # conditional prob.
DataUsed = DataUsed.rename({'time': 'cluster'})
weights_cluster = nw_crd.value_counts().reindex(DataUsed.cluster.values) # weights based on occurrence
temporal_splitting = ActualClusters[subset_type].unique() # get all available subsets of temporal flag
ConnProb = []
for i_temp in temporal_splitting:
Subset_used = [i for i in DataUsed.cluster.values if i_temp == i.split('-')[1]] # get all available clusters
Subset_used = DataUsed.sel(cluster=Subset_used) # subset only the available cluster at the temporal subset
Subset_used = Subset_used.assign_coords({'cluster': [int(i[0]) for i in Subset_used.cluster.values]}) # rename
ConnProb.append(Subset_used) # append to final list
ConnProb = xr.concat(ConnProb, dim=pd.Index(temporal_splitting, name='temporal'))
BS = statistical_brier_score(DataUsed, weights_cluster.values, dim_used='cluster') # get Brier Score
BS = BS.assign_coords({'Method': f'{subset_type}_Patt'}) # assign new coord with the temporal subsetting info
return (ConnProb.astype('float32'), BS.astype('float32'))
def connections_stats(input_data):
subset_dates, longs_used = input_data
Exceed_dataset = exceed_boolean(Precipitation.sel(time=subset_dates, longitude=longs_used))
Conn_Clusters, BS_All = [], []
for i_temp in ['All', 'HalfYear']:
i_conn, i_BS = clusters_EPEs_conditioning(subset_extremes=Exceed_dataset, subset_type=i_temp)
Conn_Clusters.append(i_conn)
BS_All.append(i_BS)
Conn_Clusters = xr.concat(Conn_Clusters, dim='temporal')
Conn_Temp, BS_Temp = DayMonth_EPEs_conditioning(subset_extremes=Exceed_dataset)
Conn_Temp2, BS_Temp2 = temporal_conditioning_subset(Exceed_dataset, 'Season')
Conn_Temp = xr.concat([Conn_Temp, Conn_Temp2], dim='temporal')
BS_All.append(BS_Temp)
BS_All.append(BS_Temp2)
BS_All = xr.concat(BS_All, dim='Method')
BS_All.name = 'BS'
BSS_All = 1 - BS_All/BS_All.sel(Method=['DayMonth_Temp', 'Season_Temp']).min('Method')
BSS_All.name = 'BSS'
BS_All = xr.merge([BS_All, BSS_All])
return {'Conn_Clusters': Conn_Clusters, 'Conn_Temp': Conn_Temp, 'BS_All': BS_All}
All_data = connections_stats([Precipitation.time.values, Lons_all])
Connections_Patterns = All_data['Conn_Clusters']
Connections_Patterns.to_netcdf(output_dir+'Connections_Patterns.nc')
Connections_Temporal = All_data['Conn_Temp']
Connections_Temporal.to_netcdf(output_dir+'Connections_Temporal.nc')
BS_All = All_data['BS_All']
BS_All.to_netcdf(output_dir+'BS_ERA5_All.nc')
del(All_data)
# function for generating a 2-d DF with the forecasted cluster allocation for each date (only for specific lead time)
def forecast_subset(lead_time):
Subset_Frcst = AllocatedClusters.query('step==@lead_time and number!=-1') # remove ens. mean data cause high bias
Subset_Frcst = Subset_Frcst.pivot_table(index='valid_time', columns='number', values='Cluster')
Subset_Frcst.index = pd.to_datetime(Subset_Frcst.index)
return Subset_Frcst
def frcst_precip_init_date(init_date_used):
' Get the reforecast data for the selected initialization date '
' There is no need to derive ens mean for precipitation data, cause the mean is very biased and wrong! '
# get the data of the control member (cf)
file_name = input_dir+'Data/Precipitation/cf/Precipitation_cf_'+init_date_used+'.grb'
control_forecast = xr.open_dataarray(file_name, engine='cfgrib')
control_forecast = control_forecast.astype('float32') # float32 for memory efficiency
control_forecast = control_forecast.sel(longitude=slice(Area_used[1], Area_used[3]),
latitude=slice(Area_used[0], Area_used[2]))
control_forecast = control_forecast.assign_coords({'number': 0})
# get the data of the ensemble members (pf)
file_name = input_dir+'Data/Precipitation/pf/Precipitation_pf_'+init_date_used+'.grb'
ensemble_forecast = xr.open_dataarray(file_name, engine='cfgrib')
ensemble_forecast = ensemble_forecast.astype('float32') # float32 for memory efficiency
ensemble_forecast = ensemble_forecast.sel(longitude=slice(Area_used[1], Area_used[3]),
latitude=slice(Area_used[0], Area_used[2]))
final = xr.concat([control_forecast, ensemble_forecast], dim='number') # combine cf and pf data
# Precipitation is a cumulative variable, so for daily values we need differences of next with day of interest
final = xr.concat([final.isel(step=0), final.diff('step')], dim='step')
final = final.assign_coords({'step': final.step.values-np.timedelta64(1, 'D')}) # step is the min possible lag
# slicing the data due to memory limitations: i_lead is defined later on
final = final.sel(step=np.timedelta64(i_lead, 'D'))
return final.reset_coords(drop=True)
def frcst_precip_all(dates):
pool = multiprocessing.Pool() # object for multiprocessing
Data_Pr = list(tqdm.tqdm(pool.imap(frcst_precip_init_date, dates), total=len(dates), position=0, leave=True))
pool.close()
Data_Pr = xr.concat(Data_Pr, dim='time')
return Data_Pr.astype('float32')
# function for getting subset of forecasted precipitation data and generating boolean for exceedance of extremes
def exceed_boolean_frcst(data):
Data = data.stack(all_data=['time', 'number'])
Quant = Data.quantile(np.array(P_used)/100, interpolation='linear', dim='all_data', keep_attrs=True) # thresholds
Quant = Quant.rename({'quantile': 'percentile'}) # rename coordinate
Quant = Quant.assign_coords({'percentile': P_used}) # assign the dim values based on percentiles
# boolean xarray for identifying if an event is over the threshold
Exceed_xr = [data>Quant.sel(percentile=i_p) for i_p in P_used] # boolean of exceedance per percentile
Exceed_xr = xr.concat(Exceed_xr, dim=pd.Index(P_used, name='percentile')) # concatenate data for all percentiles
return Exceed_xr
# function for calculating the cond. probs. for all forecasted dates
def subset_cond_prob_clustering(data_sub, temp_subset, lons_used):
temp_flag = temp_flagging(data_sub.index, temp_subset) # get temporal flags of the forecasted dates
EPEsProb = [Connections_Patterns.sel(cluster=i_cluster, temporal=i_temp, longitude=lons_used).mean('cluster')
for i_cluster, i_temp in list(zip(data_sub.values, temp_flag))]
EPEsProb = xr.concat(EPEsProb, dim=pd.Index(data_sub.index, name='time')) # concat
EPEsProb = EPEsProb.transpose('percentile', ...) # transpose
return EPEsProb.astype('float32')
# function for getting cond. prob. based on temporal subsetting only
def subset_cond_prob_temporal(data_sub, temp_subset, lons_used):
temp_flag = temp_flagging(data_sub.index, temp_subset)
CondProb = [Connections_Temporal.sel(temporal=i_temp, longitude=lons_used) for i_temp in temp_flag]
CondProb = xr.concat(CondProb, dim=pd.Index(data_sub.index, name='time')) # concat
CondProb = CondProb.transpose('percentile', ...) # transpose
return CondProb.reset_coords(drop=True).astype('float32')
# function for calculating the brier score from actual data and not based on statistics as in the other function
def BS_calculation(forecasts, observations):
BS_value = (forecasts - observations)**2
BS_value = BS_value.sum('time')/len(BS_value.time)
return BS_value#.astype('float32')
# Use dates of Cycle 46r1: 11 June 2019 - 30 June 2020
start_date = '20190611'
end_date = '20200630'
initialization_dates = pd.date_range(start_date, end_date)
# keep Mondays (0) and Thursdays (3)
initialization_dates = initialization_dates[(initialization_dates.weekday == 0) | (initialization_dates.weekday == 3)]
initialization_dates = initialization_dates.strftime('%Y%m%d')
del(start_date, end_date)
# function for performing Brier Score analysis
def brier_score_analysis(input_data):
lead_time, dates_subset, lons_used = input_data
Subset_Frcst = forecast_subset(lead_time) # generate dataframe with forecasted cluster allocations
if dates_subset is not None:
Subset_Frcst = Subset_Frcst.loc[dates_subset]
else:
# drop the dates that correspond to the last 5 initiatilization dates, so that the dataset has a
# climatologically correct number of Winter/Spring/Summer/Autumn dates, since the EPEs are based on such data
DropDates = pd.to_datetime(initialization_dates[-5:])+np.timedelta64(lead_time, 'D')
DropDates = [DropDates-pd.DateOffset(years=i) for i in range(1,21)]
DropDates = [j for i in DropDates for j in i]
Subset_Frcst = Subset_Frcst[~Subset_Frcst.index.isin(DropDates)]
Subset_Exceed = exceed_boolean(Precipitation.sel(time=Subset_Frcst.index, longitude=lons_used))
ActualClusters_Subset = ActualClusters.loc[Subset_Frcst.index, ['Label']]
BS_Forecasts = [] # list for appending all Brier Score data (to be converted in DataArray)
# calculate direct EPEs BS based on the forecasted precipitation fields
Frst_Extremes = exceed_boolean_frcst(Precip_Frcst.sel(time=Subset_Frcst.index, longitude=lons_used))
BS_Direct = BS_calculation(Frst_Extremes.mean('number'), Subset_Exceed)
BS_Forecasts.append(BS_Direct.assign_coords({'Method': 'EPEs_Direct'}))
# calculate indirect EPEs BS for forecasted clusters given the CondProb of EPEs based on cluster and halfyear
for i_type in ['HalfYear']: # ['All', 'HalfYear', 'Season']: no need to perform other temporal subsets
Cond_Prob_Frcst = subset_cond_prob_clustering(Subset_Frcst, i_type, lons_used)
Cond_Prob_Frcst = BS_calculation(Cond_Prob_Frcst, Subset_Exceed)
BS_Forecasts.append(Cond_Prob_Frcst.assign_coords({'Method': f'{i_type}_Patt'}))
Cond_Prob_Frcst_Perfect = subset_cond_prob_clustering(ActualClusters_Subset, i_type, lons_used)
Cond_Prob_Frcst_Perfect = BS_calculation(Cond_Prob_Frcst_Perfect, Subset_Exceed)
BS_Forecasts.append(Cond_Prob_Frcst_Perfect.assign_coords({'Method': f'{i_type}_Patt_Perfect'}))
del(Cond_Prob_Frcst)
# calculate precipitation BS for temporal climatological connections (reference scores)
CondProb_Clim = subset_cond_prob_temporal(Subset_Frcst, 'DayMonth', lons_used)
CondProb_Clim = BS_calculation(CondProb_Clim, Subset_Exceed)
BS_Forecasts.append( CondProb_Clim.assign_coords({'Method': 'DayMonth_Temp'}) )
CondProb_Clim = subset_cond_prob_temporal(Subset_Frcst, 'Season', lons_used)
CondProb_Clim = BS_calculation(CondProb_Clim, Subset_Exceed)
BS_Forecasts.append( CondProb_Clim.assign_coords({'Method': 'Season_Temp'}) )
CondProb_Clim = Subset_Exceed.sum('time')/len(Subset_Exceed.time)
CondProb_Clim = statistical_brier_score(CondProb_Clim, [1], dim_used='subset')
BS_Forecasts.append( CondProb_Clim.assign_coords({'Method': 'All_Temp'}) )
del(CondProb_Clim)
BS_Forecasts = xr.concat(BS_Forecasts, dim='Method')
BS_Forecasts.name = 'BS'
BS_Ref_min = BS_Forecasts.sel(Method=['DayMonth_Temp', 'Season_Temp', 'All_Temp']).min('Method')
BSS_Forecasts = 1 - BS_Forecasts/BS_Ref_min
BSS_Forecasts.name = 'BSS'
BS_Forecasts = xr.merge([BS_Forecasts, BSS_Forecasts]).to_array().rename({'variable': 'Var'})
return BS_Forecasts.astype('float32')
# subset data due to memory limitations
Lons_subsets = np.array_split(Lons_all, 2)
# get the index values of the 5th, 95th and median number, when data are ordered
l_m = int(bootstraps*5/100)
l_M = int(bootstraps*95/100)-1
Md = int(bootstraps/2)
# function for performing the full Brier Score analysis for a specific lead time (check all combinations of subsets)
def long_subset_bs_statistics(lons_used, lead_time=0):
BBS_dates = []
for i_ln, i_season in zip([247*3, 252*3, 252*3, 249*3], ['Winter', 'Spring', 'Summer', 'Autumn']):
AllDates = Days_used[temp_flagging(Days_used, 'Season')==i_season]
np.random.seed(10)
BBS_dates_i = np.random.choice(AllDates, i_ln*bootstraps) # generate all bootstrapped values
BBS_dates_i = np.array_split(BBS_dates_i, bootstraps) # split into the number of subsets (samples)
BBS_dates.append(BBS_dates_i)
BBS_dates = np.concatenate(BBS_dates, axis=1)
BBS_dates = list(BBS_dates)+[None] # add also the final bootstrap which is concerning the actual data
del(i_ln, i_season, AllDates, BBS_dates_i)
# generate bootstrapped statistics
pool = multiprocessing.Pool() # object for multiprocessing for bootstrapping
BBS_data = list(product([lead_time], BBS_dates, [lons_used]))
BBS_data = list(tqdm.tqdm(pool.imap(brier_score_analysis, BBS_data), total=len(BBS_data), position=0, leave=True))
pool.close()
BBS_data = xr.concat(BBS_data, dim='bootstrapping_frcst') # concatenate bootstrapping samples
# get the 5th, and 95th value for BS without considering the "Actual" subset
BBS_data_ordered_values = np.sort(BBS_data.isel(bootstrapping_frcst=range(bootstraps)), axis=0) # don't use Actual
P5 = BBS_data[0]*0 + BBS_data_ordered_values[l_m]
P95 = BBS_data[0]*0 + BBS_data_ordered_values[l_M]
# get the P50 after using "Actual" subset as well
BBS_data_ordered_values = np.sort(BBS_data, axis=0)
P50 = BBS_data[0]*0 + BBS_data_ordered_values[Md]
# combine all data to the final xarrays
dim_name = pd.Index(['P5', 'P50', 'Actual', 'P95'], name='bootstraps_frcst')
Final_BS = xr.concat([P5, P50, BBS_data.isel(bootstrapping_frcst=-1).reset_coords(drop=True), P95], dim=dim_name)
# get percentage of methods outperforming Reference (Sign_Ref), and direct prediction of EPEs (Sign_Direct)
Sign_Ref = ( BBS_data.sel(Var='BSS') > 0 ).reset_coords(drop=True)
Sign_Dir = ( BBS_data.sel(Var='BS') < BBS_data.sel(Var='BS', Method='EPEs_Direct') ).reset_coords(drop=True)
Sign_Combo = Sign_Dir & Sign_Ref
Sign_Ref = Sign_Ref.sum('bootstrapping_frcst')
Sign_Ref.name = 'Sign_Ref'
Sign_Dir = Sign_Dir.sum('bootstrapping_frcst')
Sign_Dir.name = 'Sign_Direct'
Sign_Combo = Sign_Combo.sum('bootstrapping_frcst')
Sign_Combo.name = 'Sign_Combo'
Sign_Final = xr.merge([Sign_Ref, Sign_Dir, Sign_Combo])
Sign_Final = Sign_Final/(bootstraps+1)
Final_BS = Final_BS.to_dataset('Var')
Final_BS = xr.merge([Final_BS, Sign_Final])
return Final_BS.astype('float32')
def final_bs_statistics(lead_time=0):
Final = []
for i_lon in Lons_subsets:
Final.append(long_subset_bs_statistics(i_lon, lead_time))
Final = xr.concat(Final, dim='longitude')
Final = Final.assign_coords({'leaddays':lead_time})
return Final
def EconomicValue(input_data):
frcst_data, obs_data, p_t = input_data
HI = ((frcst_data>=p_t).where(obs_data==1)).sum('time')
FA = ((frcst_data>=p_t).where(obs_data==0)).sum('time')
MI = ((frcst_data<p_t).where(obs_data==1)).sum('time')
CR = ((frcst_data<p_t).where(obs_data==0)).sum('time')
HitRate = HI/(HI+MI)
FalseAlarmRate = FA/(FA+CR)
EV = FalseAlarmRate*CostRatio*(1-Extr_Occur)-HitRate*Extr_Occur*(1-CostRatio)+Extr_Occur
EV = (EV_clim - EV)/(EV_clim-Extr_Occur*CostRatio)
EV = EV.assign_coords({'p_thr': p_t})
return EV
def EcVal_analysis(lead_time=0):
Subset_Frcst = forecast_subset(lead_time) # generate dataframe with forecasted pattern allocations
# drop all reforecasts of last 5 initiation dates, so that seasonal frequencies are correct
DropDates = pd.to_datetime(initialization_dates[-5:])+np.timedelta64(lead_time, 'D')
DropDates = [DropDates-pd.DateOffset(years=i) for i in range(1,21)]
DropDates = [j for i in DropDates for j in i]
Subset_Frcst = Subset_Frcst[~Subset_Frcst.index.isin(DropDates)]
# generate boolean with extremes for actual data, direct precip. forecast and indirect based on patterns
Subset_Exceed = exceed_boolean(Precipitation.sel(time=Subset_Frcst.index))
Frst_Extremes = exceed_boolean_frcst(Precip_Frcst.sel(time=Subset_Frcst.index)).mean('number')
Cond_Prob_Frcst = subset_cond_prob_clustering(Subset_Frcst, 'HalfYear', Precipitation.longitude.values)
CondProb_DayMonth = subset_cond_prob_temporal(Subset_Frcst, 'DayMonth', Precipitation.longitude.values)
CondProb_Season = subset_cond_prob_temporal(Subset_Frcst, 'Season', Precipitation.longitude.values)
# generate auxiliary data needed for calculating the Economic Value for different ratios of Cost/Gain measures
global Extr_Occur, CostRatio, EV_clim
Extr_Occur = Subset_Exceed.mean('time')
CostRatio = xr.DataArray(np.linspace(0, 1, 100), dims=['cost_ratio'],
coords={'cost_ratio': np.linspace(0, 1, 100)})
CostRatio = Extr_Occur*0+CostRatio # generate the remaining dimensions of the cost ratio
Extr_Occur = CostRatio*0+Extr_Occur # generate the "cost_ratio" dimension of the extremes percentage occurrences
EV_clim = xr.concat([CostRatio, Extr_Occur], dim='clim').min('clim') # climatological gain for each coordinate
# calculate Economic Value for direct, indirect and climatological forecasting and combine the data
pool = multiprocessing.Pool()
EV_dir = list(product([Frst_Extremes], [Subset_Exceed], np.arange(12)/11))
EV_dir = list(tqdm.tqdm(pool.imap(EconomicValue, EV_dir), total=len(EV_dir), position=0, leave=True))
EV_dir = xr.concat(EV_dir, dim='p_thr').max('p_thr')
pool.close()
thresholds_used = list(np.linspace(0,np.ceil(Cond_Prob_Frcst.max().values*100)/100,100))+[1]
pool = multiprocessing.Pool()
EV_indir = list(product([Cond_Prob_Frcst], [Subset_Exceed], thresholds_used))
EV_indir = list(tqdm.tqdm(pool.imap(EconomicValue, EV_indir), total=len(EV_indir), position=0, leave=True))
EV_indir = xr.concat(EV_indir, dim='p_thr').max('p_thr')
pool.close()
thresholds_used = list(np.linspace(0,np.ceil(CondProb_DayMonth.max().values*100)/100,100))+[1]
pool = multiprocessing.Pool()
EV_DM = list(product([CondProb_DayMonth], [Subset_Exceed], thresholds_used))
EV_DM = list(tqdm.tqdm(pool.imap(EconomicValue, EV_DM), total=len(EV_DM), position=0, leave=True))
EV_DM = xr.concat(EV_DM, dim='p_thr').max('p_thr')
pool.close()
thresholds_used = list(np.linspace(0,np.ceil(CondProb_Season.max().values*100)/100,100))+[1]
pool = multiprocessing.Pool()
EV_Sea = list(product([CondProb_Season], [Subset_Exceed], thresholds_used))
EV_Sea = list(tqdm.tqdm(pool.imap(EconomicValue, EV_Sea), total=len(EV_Sea), position=0, leave=True))
EV_Sea = xr.concat(EV_Sea, dim='p_thr').max('p_thr')
pool.close()
EV_final = xr.concat([EV_dir, EV_indir, EV_DM, EV_Sea],
dim=pd.Index(['Direct', 'Indirect', 'Clim_DayMonth', 'Clim_Seasonal'], name='Method'))
EV_final = EV_final.assign_coords({'leaddays': lead_time})
del Extr_Occur, CostRatio, EV_clim
return EV_final
LeadTimes = AllocatedClusters.step.unique()
BS_EPEs_All = []
EV_EPEs_All = []
for i_lead in tqdm.tqdm(LeadTimes):
# read the forecasted precipitation data
Precip_Frcst = frcst_precip_all(initialization_dates) # get the precip forecasts for the lead time of interest
Days_used = Precip_Frcst.time.values+np.timedelta64(i_lead, 'D') # get the valid dates for the forecasts
Precip_Frcst = Precip_Frcst.assign_coords({'time': Days_used}) # change to the valid time
Days_used = pd.to_datetime(Days_used)
BS_subset = final_bs_statistics(lead_time=i_lead)
BS_EPEs_All.append(BS_subset)
EV_subset = EcVal_analysis(lead_time=i_lead)
EV_EPEs_All.append(EV_subset)
BS_EPEs_All = xr.concat(BS_EPEs_All, dim='leaddays')
BS_EPEs_All.to_netcdf(output_dir+'BS_leaddays.nc')
EV_EPEs_All = xr.concat(EV_EPEs_All, dim='leaddays')
EV_EPEs_All.to_netcdf(output_dir+'EV_leaddays.nc')
```
| github_jupyter |
# Lab 2: Faster and Cheaper Queries with Table Partitions and Clustering
### Learning Objectives
- Create a SQL query to analyze sales from our baseline table
- Analyze the query execution plan for performance optimization opportunities
- Find out how much data was discarded in full table scans
- Analyze the same query against a partitioned table
- Learn how to create partitioned tables with SQL DDL
- Create partitioned tables for the entire dataset
- Run benchmark queries and compare performance to our baseline
## Analyze current architecture for partitions
Let's find the largest table and see the current architecture.
From a previous lab we created the below query:
```
%%bigquery
SELECT
dataset_id,
table_id,
-- Convert bytes to GB.
ROUND(size_bytes/pow(10,9),2) as size_gb,
-- Convert UNIX EPOCH to a timestamp.
TIMESTAMP_MILLIS(creation_time) AS creation_time,
TIMESTAMP_MILLIS(last_modified_time) as last_modified_time,
row_count,
CASE
WHEN type = 1 THEN 'table'
WHEN type = 2 THEN 'view'
ELSE NULL
END AS type
FROM
`dw-workshop.tpcds_2t_baseline.__TABLES__`
ORDER BY size_gb DESC
LIMIT 1
```
Our `store_sales` table is 1,545 GB and 5.7 Billion rows.
### Create a standard sales report
Your manager has asked you to query the existing data warehouse tables and build a report that shows:
- the top 10 sales `ss_net_paid` for all sales on or after `2000-01-01`
- include the name of the product
- include the name and email of the customer and whether they are a preferred customer
- exclude customers with a NULL `ss_customer_sk`
- include the date and time of the order as a formatted timestamp
__Note:__ Develop the query in the [BigQuery console](https://console.cloud.google.com/bigquery/) so you will be able to (1) have the benefit of the query validator as you type and (2) you can view the Execution Plan after your query runs.
```
%%bigquery --verbose
SELECT
PARSE_TIMESTAMP (
"%Y-%m-%d %T %p",
CONCAT(
CAST(d_date AS STRING),
' ',
CAST(t_hour AS STRING),
':',
CAST(t_minute AS STRING),
':',
CAST(t_second AS STRING),
' ',t_am_pm)
,"America/Los_Angeles")
AS timestamp,
s.ss_item_sk,
i.i_product_name,
s.ss_customer_sk,
c.c_first_name,
c.c_last_name,
c.c_email_address,
c.c_preferred_cust_flag,
s.ss_quantity,
s.ss_net_paid
FROM
`dw-workshop.tpcds_2t_baseline.store_sales` AS s
JOIN
`dw-workshop.tpcds_2t_baseline.date_dim` AS d
ON s.ss_sold_date_sk = d.d_date_sk
JOIN
`dw-workshop.tpcds_2t_baseline.time_dim` AS t
ON s.ss_sold_time_sk = t.t_time_sk
JOIN
`dw-workshop.tpcds_2t_baseline.item` AS i
ON s.ss_item_sk = i.i_item_sk
JOIN
`dw-workshop.tpcds_2t_baseline.customer` AS c
ON s.ss_customer_sk = c. c_customer_sk
WHERE d_date >= '2000-01-01' AND ss_customer_sk IS NOT NULL
ORDER BY ss_net_paid DESC
LIMIT 10
```
This simple report took 40+ seconds to execute and processed over 200+ GB of data. Let's see where we can improve.
### Gaining insight from the Query Execution Details (part 1: high level stats)
Learning how BigQuery processes your query under-the-hood is critical to understanding where you can improve performance.
After you executed the previous query, in the BigQuery console click on __Execution details__
<img src="img/bq-exec-details-ui.png" alt="BigQuery Execution Plan" style="border: 2px solid #eee; width: 700px; float: left;"/>
<p style="clear: both; padding: 20px 0;">
Your query plan should be largely similar to ours below. Scan through the execution statistics and answer the questions that follow.
</p>
<img src="img/bq-exec-plan-1.png" alt="BigQuery Execution Plan" style="border: 2px solid #eee; width: 700px; float: left;"/>
### Slot time
As you can see above, your query took `27 seconds` to process 5.7 Billion rows. So what does the `10hr 35min` slot time metric mean?
Recall from our discussion in Lab 1 that inside the BigQuery service are lots of virtual machines that massively process your data and query logic in parallel. These workers, or "slots", work together to process a single query job really quickly.
The BigQuery engine fully-manages the task of taking your query and farming it out to a workers who get the raw data and process the work to be done.
So say we had 30 minutes of slot time or 1800 seconds. If the query took 27 seconds in total to run,
but it was 36000 seconds (or 10 hours) worth of work, how many workers at minimum worked on it?
36000/27 = 1,333
And that's assuming each worker instantly had all the data it needed (no shuffling of data between workers) and was at full capacity for all 27 seconds!
The worker [quota](https://cloud.google.com/bigquery/quotas#queries) for an on-demand query is 2,000 slots at one time so we want to find ways we can optimize and reduce the resources consumed.
### Bytes shuffled
We had `382 GB` of data shuffled. What does that mean?
First let's explore the architecture of BigQuery and how it can process PB+ datasets in seconds. The BigQuery team explains how the managed service is setup in this [detailed blog post](https://cloud.google.com/blog/products/gcp/bigquery-under-the-hood) where the below diagram is sourced:

The actual Google services that the engine uses are
- Dremel (the execution engine)
- Jupiter (the petabit scale Google datacenter network)
- Colossus (distributed clusters of storage)
#### Storage
Our `store_sales` table (1,545 GB and 5.7 Billion rows) isn't stored on just one server. In fact, BigQuery [compresses and stores each column of the data](https://cloud.google.com/blog/products/gcp/inside-capacitor-bigquerys-next-generation-columnar-storage-format) and stores pieces of it across many commodity servers
#### Compute
When it comes time to process a query like `top sales after the year 2000`, BigQuery starts up a fleet of workers to grab and process pieces of data. Since most of the time no single worker has a complete picture of all 5.7 Billon rows, they need to communicate with each other by passing data back and forth. This fast in-memory data `shuffling` or `repartitioning` process can be time and resource intensive.
Below is an example diagram that shows the work-shuffle-work process for each worker:

As you can see in the Execution details, workers have a variety of tasks (waiting for data, reading it, performing computations, and writing data)
### Gaining insight from the Query Execution Details (part 2: repartitioning / shuffling)
Here is the next part of our execution plan after the data is read from disk. Note the longer blue bars indicate more time spent by workers. Generally the average worker (avg) and the slowest worker (max) are aligned unless your dataset is unbalanced and skewed heavily to a few values (hotspots).
Here you can see time spent joining against other datasets and then many repartitions afterward to shuffle the billions of rows across workers for processing. You'll note that the input and output row counts for repartitions for the same as they are purely a shuffling effort across workers.
<img src="img/bq-exec-plan-2.png" alt="BigQuery Execution Plan" style="border: 2px solid #eee; width: 700px; float: left;"/>
#### What triggers repartitions?
The performance-expensive operations in our query were:
1. looking at every record and comparing to see if it was before or after `2000-01-01`
2. sorting large volumes of data by timestamp
3. computing a calculated field
#### How much data was unused?
BigQuery had to scan and compare all records in the dataset to see if it matched our date condition. What percent of records were ultimately thrown away (pre-2000)?
```
%%bigquery
WITH stats AS (
SELECT
COUNT(*) AS all_records,
COUNTIF(d_date >= '2000-01-01') AS after_2000,
COUNTIF(d_date < '2000-01-01') AS before_2000
FROM
`dw-workshop.tpcds_2t_baseline.store_sales` AS s
JOIN
`dw-workshop.tpcds_2t_baseline.date_dim` AS d
ON s.ss_sold_date_sk = d.d_date_sk
)
SELECT
format("%'d",all_records) AS all_records,
format("%'d",after_2000) AS after_2000,
format("%'d",before_2000) AS before_2000,
ROUND(after_2000 / all_records,4) AS percent_dataset_used
FROM stats
```
We only ended up using 60% of our dataset for analysis.
Isn't there a faster way of eliminating the other 40% of records without having to check the date value of each row?
Yes! With date-partitioned tables.
## Reducing data scanned with Partitioned tables
Partitioning automatically buckets groups records of data based on a date or timestamp value which enables fast filtering by date.
We've already created a new table called `dw-workshop.tpcds_2t_flat_part_clust.partitioned_table` which we will show you how to do in the next section.
```
%%bigquery --verbose
SELECT
timestamp,
s.ss_item_sk,
i.i_product_name,
s.ss_customer_sk,
c.c_first_name,
c.c_last_name,
c.c_email_address,
c.c_preferred_cust_flag,
s.ss_quantity,
s.ss_net_paid
FROM
`dw-workshop.tpcds_2t_flat_part_clust.partitioned_table` AS s
/* Date and time tables denormalized as part of the partitioned table
JOIN
`dw-workshop.tpcds_2t_baseline.date_dim` AS d
ON s.ss_sold_date_sk = d.d_date_sk
JOIN
`dw-workshop.tpcds_2t_baseline.time_dim` AS t
ON s.ss_sold_time_sk = t.t_time_sk
*/
JOIN
`dw-workshop.tpcds_2t_baseline.item` AS i
ON s.ss_item_sk = i.i_item_sk
JOIN
`dw-workshop.tpcds_2t_baseline.customer` AS c
ON s.ss_customer_sk = c. c_customer_sk
WHERE DATE(timestamp) >= '2000-01-01' AND ss_customer_sk IS NOT NULL
ORDER BY ss_net_paid DESC
LIMIT 10
```
### Performance comparison
| | Original | Partitioned | Improvement | |
|----------------- |---------- |------------- |-------------------- |--- |
| Query time | 27s | 24.2s | 10% faster | |
| Bytes processed | 290 GB | 144 GB | 50% cheaper | |
| Slot time | 10 hr | 7 hr | 30% more efficient | |
| Bytes Shuffled | 382 GB | 293 GB | 23% more efficient | |


### Creating Partitioned Tables
You can create partitioned tables in a number of ways:
1. Using [SQL DDL](https://cloud.google.com/bigquery/docs/reference/standard-sql/data-definition-language#create_table_statement) From the results of a query
2. When you create a new table schema
For a full list check out the [documentation](https://cloud.google.com/bigquery/docs/creating-column-partitions).
Note that date-partitioned tables (dedicated user-specified date/time column) are different than [ingestion-time partitioned tables](https://cloud.google.com/bigquery/docs/creating-partitioned-tables) which are also available but not covered here.
Below is the example query to create the partitioned table we used earlier (no need to execute it).
```sql
CREATE OR REPLACE TABLE IF NOT EXISTS `dw-workshop.tpcds_2t_flat_part_clust.partitioned_table`
PARTITION BY DATE(timestamp) -- You define the column to partition on (it must be a date or time)
CLUSTER BY ss_net_paid -- Clustering is an added benefit for partitioned tables and explained later
OPTIONS (
require_partition_filter=true -- You can mandate users must provide a WHERE clause when querying
)
AS
SELECT
PARSE_TIMESTAMP (
"%Y-%m-%d %T %p",
CONCAT(
CAST(d_date AS STRING),
' ',
CAST(t_hour AS STRING),
':',
CAST(t_minute AS STRING),
':',
CAST(t_second AS STRING),
' ',t_am_pm)
,"America/Los_Angeles")
AS timestamp,
s.*
FROM
`dw-workshop.tpcds_2t_flat_part_clust.store_sales` AS s
JOIN
`dw-workshop.tpcds_2t_flat_part_clust.date_dim` AS d
ON s.ss_sold_date_sk = d.d_date_sk
JOIN
`dw-workshop.tpcds_2t_flat_part_clust.time_dim` AS t
ON s.ss_sold_time_sk = t.t_time_sk
```
## Converting the TCP-DS tables to Partitioned tables
We can quickly verify that none of the tables in our baseline schema have partitioned columns by using the metadata table `INFORMATION_SCHEMA`
```
%%bigquery
SELECT * FROM
`dw-workshop.tpcds_2t_baseline.INFORMATION_SCHEMA.COLUMNS`
WHERE
is_partitioning_column = 'YES' OR clustering_ordinal_position IS NOT NULL
```
### Create a new dataset to hold our partitioned tables
Let's leave the existing baseline dataset and tables and create a new dataset titled `tpcds_2t_flat_part_clust`
```
%%bash
## Create a BigQuery dataset for tpcds_2t_flat_part_clust if it doesn't exist
datasetexists=$(bq ls -d | grep -w tpcds_2t_flat_part_clust)
if [ -n "$datasetexists" ]; then
echo -e "BigQuery dataset already exists, let's not recreate it."
else
echo "Creating BigQuery dataset titled: tpcds_2t_flat_part_clust"
bq --location=US mk --dataset \
--description 'Partitioned and Clustered' \
$PROJECT:tpcds_2t_flat_part_clust
echo "\nHere are your current datasets:"
bq ls
fi
```
### Create a new empty partitioned table
Let's pick one table to add partitioning and clustering to. It's easiest to add a partitioning column to a data table that has existing date or timestamp columns. Here we will use the `store` table which is a dimensional table for the name and address of each storefront for our business.
#### What to cluster on?
Finding a column to partition on is often the easy part. BigQuery also supports [clustering](https://cloud.google.com/bigquery/docs/clustered-tables) on partitioned tables which can provide performance improvements for commonly filtered or sorted queries. The column(s) you specify are used to colocate related data.
```
%%bigquery
CREATE OR REPLACE TABLE tpcds_2t_flat_part_clust.store(
s_store_sk int64 NOT NULL,
s_store_id string NOT NULL,
s_rec_start_date date ,
s_rec_end_date date ,
s_closed_date_sk int64 ,
s_store_name string ,
s_number_employees int64 ,
s_floor_space int64 ,
s_hours string ,
s_manager string ,
s_market_id int64 ,
s_geography_class string ,
s_market_desc string ,
s_market_manager string ,
s_division_id int64 ,
s_division_name string ,
s_company_id int64 ,
s_company_name string ,
s_street_number string ,
s_street_name string ,
s_street_type string ,
s_suite_number string ,
s_city string ,
s_county string ,
s_state string ,
s_zip string ,
s_country string ,
s_gmt_offset numeric ,
s_tax_precentage numeric )
# TODO: Specify a date field to partition on and a field to cluster on:
PARTITION BY s_rec_start_date
CLUSTER BY s_zip;
SELECT * FROM tpcds_2t_flat_part_clust.store LIMIT 0;
```
Now that you have the empty table, it's time to populate it with data. This can take a while, feel free to cancel the execution and continue with the lab. We'll use the BigQuery Data Transfer Service later to copy over the entire dataset in seconds.
```
%%bigquery
insert into tpcds_2t_flat_part_clust.store(s_store_sk, s_store_id, s_rec_start_date, s_rec_end_date, s_closed_date_sk,
s_store_name, s_number_employees, s_floor_space, s_hours, s_manager, s_market_id, s_geography_class, s_market_desc,
s_market_manager, s_division_id, s_division_name, s_company_id, s_company_name, s_street_number, s_street_name,
s_street_type, s_suite_number, s_city, s_county, s_state, s_zip, s_country, s_gmt_offset, s_tax_precentage)
select s_store_sk, s_store_id, s_rec_start_date, s_rec_end_date, s_closed_date_sk,
s_store_name, s_number_employees, s_floor_space, s_hours, s_manager, s_market_id, s_geography_class, s_market_desc,
s_market_manager, s_division_id, s_division_name, s_company_id, s_company_name, s_street_number, s_street_name,
s_street_type, s_suite_number, s_city, s_county, s_state, s_zip, s_country, s_gmt_offset, s_tax_precentage
from `dw-workshop.tpcds_2t_baseline.store`;
SELECT * FROM tpcds_2t_flat_part_clust.store LIMIT 5;
```
## Compare performance
In the console UI, copy and paste the below queries and run them as one statement.
```sql
SELECT *
FROM `dw-workshop.tpcds_2t_baseline.store`
WHERE s_rec_start_date > '2010-01-01';
SELECT *
FROM `dw-workshop.tpcds_2t_flat_part_clust.store`
WHERE s_rec_start_date > '2010-01-01';
```
The results should look like the below

Why did the first query do an entire table scan of 59 KB (small table) while the second query barely processed any data 117 bytes = .11 KB?
It's because the second query hits a partitioned column and it automatically knows that there is no partition for 2010 data (the dataset goes up to 2003). Unlike the first query, it does this without having to open individual records.
Although this table is quite small, the same benefit applies to any partitioned table no matter how large.
## Running the 99 benchmark queries on partitioned tables
Below are the `CREATE TABLE` statements for the remaining tables in our new `tpcds_2t_flat_part_clust` dataset. Note how you can still use clustering on a table that does not have an existing field to partition on by simply adding an `empty_date` column of type `date`.
Run the below statement to finishing creating the remaining 25 tables for our new partitioned dataset tables.
```
%%bigquery
create table tpcds_2t_flat_part_clust.customer_address(
ca_address_sk int64 NOT NULL,
ca_address_id string NOT NULL,
ca_street_number string ,
ca_street_name string ,
ca_street_type string ,
ca_suite_number string ,
ca_city string ,
ca_county string ,
ca_state string ,
ca_zip string ,
ca_country string ,
ca_gmt_offset numeric ,
ca_location_type string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY ca_address_sk;
create table tpcds_2t_flat_part_clust.customer_demographics(
cd_demo_sk int64 NOT NULL,
cd_gender string ,
cd_marital_status string ,
cd_education_status string ,
cd_purchase_estimate int64 ,
cd_credit_rating string ,
cd_dep_count int64 ,
cd_dep_employed_count int64 ,
cd_dep_college_count int64 ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY cd_demo_sk;
create table tpcds_2t_flat_part_clust.date_dim(
d_date_sk int64 NOT NULL,
d_date_id string NOT NULL,
d_date date ,
d_month_seq int64 ,
d_week_seq int64 ,
d_quarter_seq int64 ,
d_year int64 ,
d_dow int64 ,
d_moy int64 ,
d_dom int64 ,
d_qoy int64 ,
d_fy_year int64 ,
d_fy_quarter_seq int64 ,
d_fy_week_seq int64 ,
d_day_name string ,
d_quarter_name string ,
d_holiday string ,
d_weekend string ,
d_following_holiday string ,
d_first_dom int64 ,
d_last_dom int64 ,
d_same_day_ly int64 ,
d_same_day_lq int64 ,
d_current_day string ,
d_current_week string ,
d_current_month string ,
d_current_quarter string ,
d_current_year string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY d_date_sk;
create table tpcds_2t_flat_part_clust.warehouse(
w_warehouse_sk int64 NOT NULL,
w_warehouse_id string NOT NULL,
w_warehouse_name string ,
w_warehouse_sq_ft int64 ,
w_street_number string ,
w_street_name string ,
w_street_type string ,
w_suite_number string ,
w_city string ,
w_county string ,
w_state string ,
w_zip string ,
w_country string ,
w_gmt_offset numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY w_warehouse_sk;
create table tpcds_2t_flat_part_clust.ship_mode(
sm_ship_mode_sk int64 NOT NULL,
sm_ship_mode_id string NOT NULL,
sm_type string ,
sm_code string ,
sm_carrier string ,
sm_contract string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY sm_carrier;
create table tpcds_2t_flat_part_clust.time_dim(
t_time_sk int64 NOT NULL,
t_time_id string NOT NULL,
t_time int64 ,
t_hour int64 ,
t_minute int64 ,
t_second int64 ,
t_am_pm string ,
t_shift string ,
t_sub_shift string ,
t_meal_time string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY t_time;
create table tpcds_2t_flat_part_clust.reason(
r_reason_sk int64 NOT NULL,
r_reason_id string NOT NULL,
r_reason_desc string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY r_reason_sk;
create table tpcds_2t_flat_part_clust.income_band(
ib_income_band_sk int64 NOT NULL,
ib_lower_bound int64 ,
ib_upper_bound int64 ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY ib_lower_bound;
create table tpcds_2t_flat_part_clust.item(
i_item_sk int64 NOT NULL,
i_item_id string NOT NULL,
i_rec_start_date date ,
i_rec_end_date date ,
i_item_desc string ,
i_current_price numeric ,
i_wholesale_cost numeric ,
i_brand_id int64 ,
i_brand string ,
i_class_id int64 ,
i_class string ,
i_category_id int64 ,
i_category string ,
i_manufact_id int64 ,
i_manufact string ,
i_size string ,
i_formulation string ,
i_color string ,
i_units string ,
i_container string ,
i_manager_id int64 ,
i_product_name string )
PARTITION BY i_rec_start_date
CLUSTER BY i_category;
create table tpcds_2t_flat_part_clust.store(
s_store_sk int64 NOT NULL,
s_store_id string NOT NULL,
s_rec_start_date date ,
s_rec_end_date date ,
s_closed_date_sk int64 ,
s_store_name string ,
s_number_employees int64 ,
s_floor_space int64 ,
s_hours string ,
s_manager string ,
s_market_id int64 ,
s_geography_class string ,
s_market_desc string ,
s_market_manager string ,
s_division_id int64 ,
s_division_name string ,
s_company_id int64 ,
s_company_name string ,
s_street_number string ,
s_street_name string ,
s_street_type string ,
s_suite_number string ,
s_city string ,
s_county string ,
s_state string ,
s_zip string ,
s_country string ,
s_gmt_offset numeric ,
s_tax_precentage numeric )
PARTITION BY s_rec_start_date
CLUSTER BY s_zip;
create table tpcds_2t_flat_part_clust.call_center(
cc_call_center_sk int64 NOT NULL,
cc_call_center_id string NOT NULL,
cc_rec_start_date date ,
cc_rec_end_date date ,
cc_closed_date_sk int64 ,
cc_open_date_sk int64 ,
cc_name string ,
cc_class string ,
cc_employees int64 ,
cc_sq_ft int64 ,
cc_hours string ,
cc_manager string ,
cc_mkt_id int64 ,
cc_mkt_class string ,
cc_mkt_desc string ,
cc_market_manager string ,
cc_division int64 ,
cc_division_name string ,
cc_company int64 ,
cc_company_name string ,
cc_street_number string ,
cc_street_name string ,
cc_street_type string ,
cc_suite_number string ,
cc_city string ,
cc_county string ,
cc_state string ,
cc_zip string ,
cc_country string ,
cc_gmt_offset numeric ,
cc_tax_percentage numeric )
PARTITION BY cc_rec_start_date
CLUSTER BY cc_county;
create table tpcds_2t_flat_part_clust.customer(
c_customer_sk int64 NOT NULL,
c_customer_id string NOT NULL,
c_current_cdemo_sk int64 ,
c_current_hdemo_sk int64 ,
c_current_addr_sk int64 ,
c_first_shipto_date_sk int64 ,
c_first_sales_date_sk int64 ,
c_salutation string ,
c_first_name string ,
c_last_name string ,
c_preferred_cust_flag string ,
c_birth_day int64 ,
c_birth_month int64 ,
c_birth_year int64 ,
c_birth_country string ,
c_login string ,
c_email_address string ,
c_last_review_date_sk int64 ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY c_customer_sk;
create table tpcds_2t_flat_part_clust.web_site(
web_site_sk int64 NOT NULL,
web_site_id string NOT NULL,
web_rec_start_date date ,
web_rec_end_date date ,
web_name string ,
web_open_date_sk int64 ,
web_close_date_sk int64 ,
web_class string ,
web_manager string ,
web_mkt_id int64 ,
web_mkt_class string ,
web_mkt_desc string ,
web_market_manager string ,
web_company_id int64 ,
web_company_name string ,
web_street_number string ,
web_street_name string ,
web_street_type string ,
web_suite_number string ,
web_city string ,
web_county string ,
web_state string ,
web_zip string ,
web_country string ,
web_gmt_offset numeric ,
web_tax_percentage numeric )
PARTITION BY web_rec_start_date
CLUSTER BY web_site_sk;
create table tpcds_2t_flat_part_clust.store_returns(
sr_returned_date_sk int64 ,
sr_return_time_sk int64 ,
sr_item_sk int64 NOT NULL,
sr_customer_sk int64 ,
sr_cdemo_sk int64 ,
sr_hdemo_sk int64 ,
sr_addr_sk int64 ,
sr_store_sk int64 ,
sr_reason_sk int64 ,
sr_ticket_number int64 NOT NULL,
sr_return_quantity int64 ,
sr_return_amt numeric ,
sr_return_tax numeric ,
sr_return_amt_inc_tax numeric ,
sr_fee numeric ,
sr_return_ship_cost numeric ,
sr_refunded_cash numeric ,
sr_reversed_charge numeric ,
sr_store_credit numeric ,
sr_net_loss numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY sr_ticket_number;
create table tpcds_2t_flat_part_clust.household_demographics(
hd_demo_sk int64 NOT NULL,
hd_income_band_sk int64 ,
hd_buy_potential string ,
hd_dep_count int64 ,
hd_vehicle_count int64 ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY hd_buy_potential;
create table tpcds_2t_flat_part_clust.web_page(
wp_web_page_sk int64 NOT NULL,
wp_web_page_id string NOT NULL,
wp_rec_start_date date ,
wp_rec_end_date date ,
wp_creation_date_sk int64 ,
wp_access_date_sk int64 ,
wp_autogen_flag string ,
wp_customer_sk int64 ,
wp_url string ,
wp_type string ,
wp_char_count int64 ,
wp_link_count int64 ,
wp_image_count int64 ,
wp_max_ad_count int64 )
PARTITION BY wp_rec_start_date
CLUSTER BY wp_web_page_sk;
create table tpcds_2t_flat_part_clust.promotion(
p_promo_sk int64 NOT NULL,
p_promo_id string NOT NULL,
p_start_date_sk int64 ,
p_end_date_sk int64 ,
p_item_sk int64 ,
p_cost numeric ,
p_response_target int64 ,
p_promo_name string ,
p_channel_dmail string ,
p_channel_email string ,
p_channel_catalog string ,
p_channel_tv string ,
p_channel_radio string ,
p_channel_press string ,
p_channel_event string ,
p_channel_demo string ,
p_channel_details string ,
p_purpose string ,
p_discount_active string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY p_promo_sk;
create table tpcds_2t_flat_part_clust.catalog_page(
cp_catalog_page_sk int64 NOT NULL,
cp_catalog_page_id string NOT NULL,
cp_start_date_sk int64 ,
cp_end_date_sk int64 ,
cp_department string ,
cp_catalog_number int64 ,
cp_catalog_page_number int64 ,
cp_description string ,
cp_type string ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY cp_catalog_page_sk;
create table tpcds_2t_flat_part_clust.inventory(
inv_date_sk int64 NOT NULL,
inv_item_sk int64 NOT NULL,
inv_warehouse_sk int64 NOT NULL,
inv_quantity_on_hand int64 ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY inv_item_sk;
create table tpcds_2t_flat_part_clust.catalog_returns(
cr_returned_date_sk int64 ,
cr_returned_time_sk int64 ,
cr_item_sk int64 NOT NULL,
cr_refunded_customer_sk int64 ,
cr_refunded_cdemo_sk int64 ,
cr_refunded_hdemo_sk int64 ,
cr_refunded_addr_sk int64 ,
cr_returning_customer_sk int64 ,
cr_returning_cdemo_sk int64 ,
cr_returning_hdemo_sk int64 ,
cr_returning_addr_sk int64 ,
cr_call_center_sk int64 ,
cr_catalog_page_sk int64 ,
cr_ship_mode_sk int64 ,
cr_warehouse_sk int64 ,
cr_reason_sk int64 ,
cr_order_number int64 NOT NULL,
cr_return_quantity int64 ,
cr_return_amount numeric ,
cr_return_tax numeric ,
cr_return_amt_inc_tax numeric ,
cr_fee numeric ,
cr_return_ship_cost numeric ,
cr_refunded_cash numeric ,
cr_reversed_charge numeric ,
cr_store_credit numeric ,
cr_net_loss numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY cr_item_sk;
create table tpcds_2t_flat_part_clust.web_returns(
wr_returned_date_sk int64 ,
wr_returned_time_sk int64 ,
wr_item_sk int64 NOT NULL,
wr_refunded_customer_sk int64 ,
wr_refunded_cdemo_sk int64 ,
wr_refunded_hdemo_sk int64 ,
wr_refunded_addr_sk int64 ,
wr_returning_customer_sk int64 ,
wr_returning_cdemo_sk int64 ,
wr_returning_hdemo_sk int64 ,
wr_returning_addr_sk int64 ,
wr_web_page_sk int64 ,
wr_reason_sk int64 ,
wr_order_number int64 NOT NULL,
wr_return_quantity int64 ,
wr_return_amt numeric ,
wr_return_tax numeric ,
wr_return_amt_inc_tax numeric ,
wr_fee numeric ,
wr_return_ship_cost numeric ,
wr_refunded_cash numeric ,
wr_reversed_charge numeric ,
wr_account_credit numeric ,
wr_net_loss numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY wr_web_page_sk;
create table tpcds_2t_flat_part_clust.web_sales(
ws_sold_date_sk int64 ,
ws_sold_time_sk int64 ,
ws_ship_date_sk int64 ,
ws_item_sk int64 NOT NULL,
ws_bill_customer_sk int64 ,
ws_bill_cdemo_sk int64 ,
ws_bill_hdemo_sk int64 ,
ws_bill_addr_sk int64 ,
ws_ship_customer_sk int64 ,
ws_ship_cdemo_sk int64 ,
ws_ship_hdemo_sk int64 ,
ws_ship_addr_sk int64 ,
ws_web_page_sk int64 ,
ws_web_site_sk int64 ,
ws_ship_mode_sk int64 ,
ws_warehouse_sk int64 ,
ws_promo_sk int64 ,
ws_order_number int64 NOT NULL,
ws_quantity int64 ,
ws_wholesale_cost numeric ,
ws_list_price numeric ,
ws_sales_price numeric ,
ws_ext_discount_amt numeric ,
ws_ext_sales_price numeric ,
ws_ext_wholesale_cost numeric ,
ws_ext_list_price numeric ,
ws_ext_tax numeric ,
ws_coupon_amt numeric ,
ws_ext_ship_cost numeric ,
ws_net_paid numeric ,
ws_net_paid_inc_tax numeric ,
ws_net_paid_inc_ship numeric ,
ws_net_paid_inc_ship_tax numeric ,
ws_net_profit numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY ws_item_sk;
create table tpcds_2t_flat_part_clust.catalog_sales(
cs_sold_date_sk int64 ,
cs_sold_time_sk int64 ,
cs_ship_date_sk int64 ,
cs_bill_customer_sk int64 ,
cs_bill_cdemo_sk int64 ,
cs_bill_hdemo_sk int64 ,
cs_bill_addr_sk int64 ,
cs_ship_customer_sk int64 ,
cs_ship_cdemo_sk int64 ,
cs_ship_hdemo_sk int64 ,
cs_ship_addr_sk int64 ,
cs_call_center_sk int64 ,
cs_catalog_page_sk int64 ,
cs_ship_mode_sk int64 ,
cs_warehouse_sk int64 ,
cs_item_sk int64 NOT NULL,
cs_promo_sk int64 ,
cs_order_number int64 NOT NULL,
cs_quantity int64 ,
cs_wholesale_cost numeric ,
cs_list_price numeric ,
cs_sales_price numeric ,
cs_ext_discount_amt numeric ,
cs_ext_sales_price numeric ,
cs_ext_wholesale_cost numeric ,
cs_ext_list_price numeric ,
cs_ext_tax numeric ,
cs_coupon_amt numeric ,
cs_ext_ship_cost numeric ,
cs_net_paid numeric ,
cs_net_paid_inc_tax numeric ,
cs_net_paid_inc_ship numeric ,
cs_net_paid_inc_ship_tax numeric ,
cs_net_profit numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY cs_item_sk;
create table tpcds_2t_flat_part_clust.store_sales(
ss_sold_date_sk int64 ,
ss_sold_time_sk int64 ,
ss_item_sk int64 NOT NULL,
ss_customer_sk int64 ,
ss_cdemo_sk int64 ,
ss_hdemo_sk int64 ,
ss_addr_sk int64 ,
ss_store_sk int64 ,
ss_promo_sk int64 ,
ss_ticket_number int64 NOT NULL,
ss_quantity int64 ,
ss_wholesale_cost numeric ,
ss_list_price numeric ,
ss_sales_price numeric ,
ss_ext_discount_amt numeric ,
ss_ext_sales_price numeric ,
ss_ext_wholesale_cost numeric ,
ss_ext_list_price numeric ,
ss_ext_tax numeric ,
ss_coupon_amt numeric ,
ss_net_paid numeric ,
ss_net_paid_inc_tax numeric ,
ss_net_profit numeric ,
empty_date date )
PARTITION BY empty_date
CLUSTER BY ss_item_sk
```
### Ingesting data into partitioned tables
We could simply `INSERT INTO` the billions of records from our baseline dataset into our new partitioned dataset but that would take quite a while (a few hours). Instead, we'll simply use the new [BigQuery Dataset Copy API (beta)](https://cloud.google.com/bigquery/docs/copying-datasets) to populate the tables from an already existing solution in our `dw-workshop` project.
### Use the BigQuery Data Transfer Service to copy an existing dataset
1. Enable the [BigQuery Data Transfer Service API](https://console.cloud.google.com/apis/library/bigquerydatatransfer.googleapis.com)
2. Navigate to the [BigQuery console and the existing `dw-workshop` dataset](https://console.cloud.google.com/bigquery?project=dw-workshop&p=dw-workshop&d=tpcds_2t_baseline&page=dataset)
3. Click Copy Dataset

4. In the pop-up, choose your __project name__ and the newly created __dataset name__ from the previous step
__BE SURE TO CHOOSE THE NEW DATASET NAME IN THE DROP DOWN `tpcds_2t_flat_part_clust` OR YOU WILL OVERWRITE YOUR BASELINE TABLES__
5. Click __Copy__
6. Wait for the transfer to complete
### Verify you now have the baseline data in your project
Run the below query and confirm you see data. Note that if you omit the `project-id` ahead of the dataset name in the `FROM` clause, BigQuery will assume your default project.
```
%%bigquery
SELECT COUNT(*) AS store_transaction_count
FROM tpcds_2t_flat_part_clust.store_sales
```
### Running a few benchmark queries with a shell script
```
%%bash
# runs the SQL queries from the TPCDS benchmark
# Pull the current Google Cloud Platform project name
export PROJECT=$(gcloud config list project --format "value(core.project)")
BQ_DATASET="tpcds_2t_flat_part_clust" # let's benchmark our new dataset
QUERY_FILE_PATH="/home/jupyter/$PROJECT/02_add_partition_and_clustering/solution/sql/full_performance_benchmark.sql" #sample_benchmark.sql
IFS=";"
# create perf table to keep track of run times for all 99 queries
printf "\033[32;1m Housekeeping tasks... \033[0m\n\n";
printf "Creating a reporting table perf to track how fast each query runs...";
perf_table_ddl="CREATE TABLE IF NOT EXISTS $BQ_DATASET.perf(performance_test_num int64, query_num int64, elapsed_time_sec int64, ran_on int64)"
bq rm -f $BQ_DATASET.perf
bq query --nouse_legacy_sql $perf_table_ddl
start=$(date +%s)
index=0
for select_stmt in $(<$QUERY_FILE_PATH)
do
# run the test until you hit a line with the string 'END OF BENCHMARK' in the file
if [[ "$select_stmt" == *'END OF BENCHMARK'* ]]; then
break
fi
printf "\n\033[32;1m Let's benchmark this query... \033[0m\n";
printf "$select_stmt";
SECONDS=0;
bq query --use_cache=false --nouse_legacy_sql $select_stmt # critical to turn cache off for this test
duration=$SECONDS
# get current timestamp in milliseconds
ran_on=$(date +%s)
index=$((index+1))
printf "\n\033[32;1m Here's how long it took... \033[0m\n\n";
echo "Query $index ran in $(($duration / 60)) minutes and $(($duration % 60)) seconds."
printf "\n\033[32;1m Writing to our benchmark table... \033[0m\n\n";
insert_stmt="insert into $BQ_DATASET.perf(performance_test_num, query_num, elapsed_time_sec, ran_on) values($start, $index, $duration, $ran_on)"
printf "$insert_stmt"
bq query --nouse_legacy_sql $insert_stmt
done
end=$(date +%s)
printf "Benchmark test complete"
```
## Benchmarking all 99 queries
```
%%bigquery
SELECT * FROM `dw-workshop.tpcds_2t_flat_part_clust.perf` # public table
WHERE
# Let's only pull the results from our most recent test
performance_test_num = (SELECT MAX(performance_test_num) FROM `dw-workshop.tpcds_2t_flat_part_clust.perf`)
ORDER BY ran_on
%%bigquery
SELECT
TIMESTAMP_SECONDS(MAX(performance_test_num)) AS test_date,
MAX(performance_test_num) AS latest_performance_test_num,
COUNT(DISTINCT query_num) AS count_queries_benchmarked,
SUM(elapsed_time_sec) AS total_time_sec,
MIN(elapsed_time_sec) AS fastest_query_time_sec,
MAX(elapsed_time_sec) AS slowest_query_time_sec
FROM
`dw-workshop.tpcds_2t_flat_part_clust.perf`
WHERE
performance_test_num = (SELECT MAX(performance_test_num) FROM `dw-workshop.tpcds_2t_flat_part_clust.perf`)
```
### Results
The total time for the benchmark queries on our newly partitioned dataset is 3680 seconds or 61 minutes. (That's 23% faster than 79 minutes for the baseline)
## Compare vs baseline
Using SQL we can compare our benchmark tests pretty easily.
```
%%bigquery
# TODO write where clause filters to pull latest performance from each table (and debug why they keep getting truncated)
WITH
add_part AS (
SELECT * FROM `dw-workshop.tpcds_2t_flat_part_clust.perf`)
, base AS (
SELECT * FROM `dw-workshop.tpcds_2t_baseline.perf` )
SELECT
base.query_num,
base.elapsed_time_sec AS elapsed_time_sec_base,
add_part.elapsed_time_sec AS elapsed_time_sec_add_part,
add_part.elapsed_time_sec - base.elapsed_time_sec AS delta
FROM base JOIN add_part USING(query_num)
ORDER BY delta
```
__Final Activity:__ Create a Data Studio report or ipynb visualization showing the differences between performance. Which queries saw the most improvement?
| github_jupyter |

---
## 01. Introducción a SQL
Eduard Larrañaga (ealarranaga@unal.edu.co)
---
### Resumen
En este cuaderno se presenta un manejo básico de SQLite3.
---
### SQL
SQL (**S**tructured **Q**uery **L**anguage) es un lenguaje estandarizado para almacenar, manipular y obtener información de bases de datos.
Este lenguaje es particularmente útil para manipular datos estructurados, i.e. datos que incorporan relacinoes entre entidades y variables.
### Conociendo los datos
En esta lección utilizaremos un archivo de datos utilizado en el curso CS50x de la Universidad de Harvard
https://docs.google.com/spreadsheets/d/e/2PACX-1vRfpjV8pF6iNBu5xV-wnzHPXvW69wZcTxqsSnYqHx126N0bPfVhq63UtkG9mqUawB4tXneYh31xJlem/pubhtml
(Este curso esta disponible en https://cs50.harvard.edu/x/2021/).
---
La hoja de calculo y la base de datos contiene información de shows en el sitio Netflix.
---
### SQLite
[SQLite](https://www.sqlite.org/index.html) es una libreria construida en lenguaje C que implementa una versión pequeña y rápida de SQL. Para verificar si ya tiene instalao SQLite en su computador puede utilizar el comando
```
$ sqlite3 --version
```
En caso de no estar instaldo, puede descargarlo de
https://www.sqlite.org/download.html
---
El primer paso para utlizar SQLite es cargar la base de datos utilizando el comando
```
$ sqlite3 shows.db
```
Cuando se carga la información, el promt del sistema se cambiará a `sqlite>`.
La estructura de la base de datos se puede obtener con el comando
```
sqlite> .schema
```
La estructura de una tabla particular dentre de la base de datos se obtiene mediante
```
sqlite> .schema stars
```
---
### Estructura de un 'Llamado' (Query)
Un llamado (query) en SQL consiste de tres partes o bloques: el bloque **SELECT**, el bloque **FROM** y el bloque **WHERE**.
- El bloque SELECT le dice a la base de datos que columnas se desean obtener. El nombre de cada columna o característica se separa con una coma.
- El bloque FROM especifica de cual tabla (o tablas) se quiere extraer la información.
Por ejemplo, si se quiere obtener la columna `year` de la tabla `shows`, se utiliza el comando
```
SELECT year FROM shows;
```
Si se quieren obtener las columnas `title` y `year` de la tabla `shows`, se utiliza
```
SELECT title, year FROM shows;
```
Cuando se quieren seleccionar TODAS las columnas de una tabla, se utiliza
```
SELECT * FROM shows;
```
- El bloque WHERE permite especificar alguna(s) característica(s) para restringir la busqueda. La lista de especificaciones debe estar separada por operadores booleanos.
Suponga que se quiere restringir la busqueda a un show particular definiendo su titulo.
```
SELECT * FROM shows WHERE title='Black Mirror';
```
Ahora intentamos algunos comandos básicos en SQLite.
1. Mostrar la estructura de la base de datos
```
sqlite> .schema
```
2. Mostrar la estructura de una tabla particular en la base de datos
```
sqlite> .schema stars
```
3. Selecciona todas las características de una tabla (en general puede ser mucha información!)
```
SELECT * FROM shows;
```
4. Cuántas entradas se tienen con las características buscadas?
```
SELECT COUNT(*) FROM shows;
```
5. Seleccionar una muestra particular de la lista (e.g. con un titulo específico)
```
SELECT * FROM shows WHERE title = 'Black Mirror';
```
6. Seleccionar los primeros N resultados en una busqueda
```
SELECT title, year FROM shows WHERE year = 2019 LIMIT 5;
```
7. En algunas ocasiones , una busqueda específica puede retornar varios resultados,
```
SELECT * FROM shows WHERE title = 'The Office';
```
8. Seleccionar muestras con una parte de texto,
```
SELECT * FROM shows WHERE title LIKE '%Things%';
```
9. Selecionar muestras mediante una especificación de valores numéricos
```
SELECT * FROM shows WHERE year > 2020;
```
10. Seleccionar una muestra con una característica que no se espcifica completamente
```
SELECT year FROM shows WHERE title LIKE 'Stranger Things';
```
11. Ordenar la selección de acuerdo con una característica
```
SELECT * FROM shows WHERE title LIKE 'Doctor Who%' ORDER BY year;
```
```
SELECT * FROM shows WHERE title LIKE 'Doctor Who%' ORDER BY year DESC;
```
```
SELECT * FROM shows WHERE title LIKE 'Doctor Who%' ORDER BY year DESC LIMIT 10;
```
12. Incluir operadores booleanos en la busqueda
```
SELECT * FROM shows WHERE year > 1990 AND year < 2000 ;
```
```
SELECT * FROM shows WHERE year BETWEEN 1990 AND 2000 ;
```
```
SELECT id FROM shows WHERE title='Stranger Things' AND year = 2016;
```
13. Busqueda involucrando más de una tabla
```
SELECT * FROM genres WHERE show_id = 4574334;
```
Esta busqueda se puede realizar más automáticamente mediante
```
SELECT * FROM genres WHERE show_id = (SELECT id FROM shows WHERE title='Stranger Things' AND year = 2016);
```
14. Para salir de SQLite, se hace
```
.quit
```
---
### Ejercicios
1. Cuantos shows tienen un rating perfecto de 10.0?
2. Cuantos episodios tiene el show 'Black Mirror'?
3. Cuantos shows hay en el genero de Sci-Fi?
4. Cual es el show con mejor rating en el genero de Horror?
5. Cuantos shows del genero Animation hay en la base de datos?
6. Cuales son los 10 shows con peor rating en el genero de animación entre el 2005 y el 2010?
| github_jupyter |
```
import s3fs
import os
import json
import time
import pickle
import requests
import traceback
import time
from datetime import datetime
from sklearn import set_config
import warnings
# Ignore warnings from scikit-learn to make this notebook a bit nicer
warnings.simplefilter('ignore')
warnings.filterwarnings('ignore')
import pandas as pd
from pandas import DataFrame
from pandas import plotting
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import seaborn as sns
import re
from tqdm.autonotebook import tqdm
tqdm.pandas(desc="progress-bar", leave=False)
import string
import unicodedata # might need to pip install unicodedate2 on aws sagemaker
import contractions
from contractions import contractions_dict ## pip installed this
from wordcloud import WordCloud, STOPWORDS #pip install
from textblob import TextBlob
!python -m textblob.download_corpora
import nltk
import nltk.corpus
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
from nltk.tokenize import word_tokenize
from nltk.stem.wordnet import WordNetLemmatizer
from nltk.tokenize import ToktokTokenizer
from nltk.corpus import stopwords
import gensim
from gensim.utils import simple_preprocess
from gensim.parsing.preprocessing import preprocess_string
from gensim.parsing.preprocessing import STOPWORDS
from gensim.models import word2vec
import multiprocessing as mp
import sklearn
from sklearn.utils import resample # Covert too much Rock! to just enough
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.decomposition import TruncatedSVD
%matplotlib inline
sns.set(style='darkgrid',palette='Dark2',rc={'figure.figsize':(9,6),'figure.dpi':90})
# Increase screen size.
#pd.set_option('display.height', 1000)
pd.set_option('display.max_rows', 500)
pd.set_option('display.max_columns', 500)
pd.set_option('display.width', 1000)
%matplotlib inline
sns.set(style='darkgrid',palette='Dark2', rc={'figure.figsize':(9,6), 'figure.dpi':100})
# Set the default figure size for matplotlib
plt.rcParams['figure.figsize'] = (9, 6)
# Visual analysis of model performance
from yellowbrick.classifier import confusion_matrix
from yellowbrick.classifier import classification_report
from yellowbrick.regressor import prediction_error, ResidualsPlot
from yellowbrick.target import ClassBalance
from yellowbrick.target import BalancedBinningReference
from yellowbrick.text import FreqDistVisualizer
from yellowbrick.classifier import ConfusionMatrix, ROCAUC, PrecisionRecallCurve, ClassificationReport
from yellowbrick.classifier import ClassPredictionError
from yellowbrick.model_selection import FeatureImportances
from yellowbrick.model_selection import ValidationCurve
from yellowbrick.contrib.classifier import DecisionViz
#from mlxtend.plotting import plot_decision_regions
#Pipeline toolset
# Used to divide our dataseets into train/test splits
# Data will be randomly shuffled so running this notebook multiple times may lead to different results
from sklearn.model_selection import train_test_split as tts
from sklearn.pipeline import Pipeline
from sklearn.pipeline import FeatureUnion
from sklearn.compose import ColumnTransformer, make_column_selector
from sklearn.preprocessing import Normalizer, RobustScaler, OneHotEncoder, LabelEncoder, OrdinalEncoder, StandardScaler, MinMaxScaler
from sklearn.impute import SimpleImputer
#Model toolset
from sklearn.svm import LinearSVC, NuSVC, SVC
from sklearn.decomposition import TruncatedSVD
from sklearn.ensemble import BaggingClassifier, ExtraTreesClassifier, RandomForestClassifier, AdaBoostClassifier
from sklearn.ensemble import RandomForestRegressor
from sklearn.neighbors import KNeighborsClassifier
from sklearn.linear_model import LogisticRegressionCV, LogisticRegression, SGDClassifier
from sklearn.naive_bayes import MultinomialNB
from sklearn.naive_bayes import GaussianNB
from sklearn.naive_bayes import BernoulliNB
from sklearn.linear_model import Ridge
from sklearn.tree import DecisionTreeClassifier
from sklearn.neural_network import MLPClassifier
#Evaluation toolset
from sklearn.metrics import f1_score
from sklearn.model_selection import StratifiedKFold, cross_val_score
import pickle
from joblib import dump, load
g_df = pd.read_csv('g2_df')
#Drop first, useless column.
g_df.drop(columns=['Unnamed: 0'], axis=1, inplace=True)
g_df.info()
#Decision on what features to include, borne of EDA and visual steering.
df = pd.DataFrame((g_df), columns=['genre','full_word_count','full_character_count',
'med_rock_bool','med_hiphop_bool','med_pop_bool',
'sml_word_count','sml_character_count',
'sml_sent_label','sml_content_affin','sml_vector'])
df.describe(include='all')
df.columns
seed = 99
#Separate majority and minority classes, twice.
majority = df[df.genre=='Rock']
minority = df[df.genre=='Hip Hop']
# Downsample majority class
majority_rock_downsampled = resample(majority,
replace=False, # sample without replacement
n_samples=len(minority), # to match minority class
random_state=seed) # reproducible results
seed = 99
#Separate majority and minority classes, again.
majority = df[df.genre=='Pop']
minority = df[df.genre=='Hip Hop']
# Downsample majority class
majority_pop_downsampled = resample(majority,
replace=False, # sample without replacement
n_samples=len(minority), # to match minority class
random_state=seed) # reproducible results
# Combine minority class with downsampled majority class
dfd = pd.concat([majority_rock_downsampled, majority_pop_downsampled, minority])
# Display new class counts
dfd['genre'].value_counts()
# dfd = 'data frame downsampled'
target_mnb = dfd.genre
features_mnb = dfd[['full_word_count','full_character_count',
'med_rock_bool','med_hiphop_bool','med_pop_bool',
'sml_word_count','sml_character_count',
'sml_sent_label','sml_content_affin','sml_vector']].copy()
X_train_mnb, X_test_mnb, y_train_mnb, y_test_mnb = tts(features_mnb, target_mnb, test_size = 0.2, random_state=123)
print(df.shape); print(X_train_mnb.shape); print(X_test_mnb.shape)
numerical = ['full_word_count','full_character_count','sml_word_count','sml_character_count']
negative_values = ['med_rock_bool','med_hiphop_bool','med_pop_bool','sml_content_affin']
categorical = ['sml_sent_label']
textual = ['sml_vector']
#Had to DROP content_affin, and go MinMax Scaler everywhere else. To avoid a negative in X_train.
%time
ct_mnb = ColumnTransformer(
[('num', MinMaxScaler(feature_range=(0,10)), numerical),
('neg_values', MinMaxScaler(), negative_values),
('sentiment_label', OneHotEncoder(dtype='int', handle_unknown='ignore'), ['sml_sent_label']),
('tfidf', TfidfVectorizer(max_features = 6000, stop_words = 'english', ngram_range=(1,1)), 'sml_vector')], n_jobs=3, verbose=True)
set_config(display='text')
ct_mnb
%%time
# Creating the feature matrix
X_train_mnb = ct_mnb.fit_transform(X_train_mnb)
X_test_mnb = ct_mnb.transform(X_test_mnb)
print(f'Shape of Term Frequency Matrix of train: {X_train_mnb.shape}')
print(f'Shape of Term Frequency Matrix of test: {X_test_mnb.shape}')
%%time
Encoder = LabelEncoder()
y_train_mnb = Encoder.fit_transform(y_train_mnb)
y_test_mnb = Encoder.fit_transform(y_test_mnb)
print(y_train_mnb.shape); print(y_test_mnb.shape)
classes = Encoder.classes_
classes
%%time
# MultinomialNB
mnb = MultinomialNB(alpha=1)
# Training the model
mnb.fit(X_train_mnb, y_train_mnb)
#Predict the Test using MultinominalNB
y_pred_mnb = mnb.predict(X_test_mnb)
print('Accuracy on x_train is',mnb.score(X_train_mnb, y_train_mnb))
print('Accuracy on x_test is',mnb.score(X_test_mnb, y_test_mnb))
from sklearn.model_selection import StratifiedKFold, cross_val_score
scores = cross_val_score(mnb, target_mnb, features_mnb, cv=StratifiedKFold(12))
scores
%time
#save
dump(mnb, 'mnb.joblib')
#load
#mnb = load('mnb.joblib')
%%time
cm_mnb = ConfusionMatrix(mnb, classes=classes, cmap='RdPu')
cm_mnb.fit(X_train_mnb, y_train_mnb)
cm_mnb.score(X_test_mnb, y_test_mnb)
cm_mnb.show()
%%time
cr_mnb = ClassificationReport(mnb, classes=classes, support=True)
cr_mnb.fit(X_train_mnb, y_train_mnb)
cr_mnb.score(X_test_mnb, y_test_mnb)
cr_mnb.show()
%%time
# MultinomialNB
mnb2 = BernoulliNB()
# Training the model
mnb2.fit(X_train_mnb, y_train_mnb)
#Predict the Test using MultinominalNB
y_pred_mnb2 = mnb.predict(X_test_mnb)
print('Accuracy on x_train is',mnb.score(X_train_mnb, y_train_mnb))
print('Accuracy on x_test is',mnb.score(X_test_mnb, y_test_mnb))
%%time
cm_mnb2 = ConfusionMatrix(mnb2, classes=classes, cmap='RdPu')
cm_mnb2.fit(X_train_mnb, y_train_mnb)
cm_mnb2.score(X_test_mnb, y_test_mnb)
cm_mnb2.show()
%%time
cr_mnb2 = ClassificationReport(mnb2, classes=classes, support=True)
cr_mnb2.fit(X_train_mnb, y_train_mnb)
cr_mnb2.score(X_test_mnb, y_test_mnb)
cr_mnb2.show()
%time
from sklearn.model_selection import GridSearchCV
# Set the parameters by cross-validation
tuned_parameters = [{'alpha':[2], 'fit_prior':[False, True]}]
scores = ['precision', 'recall']
for score in scores:
print("# Tuning hyper-parameters for %s" % score)
print()
clf = GridSearchCV(
mnb, tuned_parameters, scoring='%s_macro' % score
)
clf.fit(X_train_mnb, y_train_mnb)
print("Best parameters set found on development set:")
print()
print(clf.best_params_)
print()
print("Grid scores on development set:")
print()
means = clf.cv_results_['mean_test_score']
stds = clf.cv_results_['std_test_score']
for mean, std, params in zip(means, stds, clf.cv_results_['params']):
print("%0.3f (+/-%0.03f) for %r"
% (mean, std * 2, params))
print()
print("Detailed classification report:")
print()
print("The model is trained on the full development set.")
print("The scores are computed on the full evaluation set.")
print()
y_true, y_pred = y_test_mnb, clf.predict(X_test_mnb)
print(classification_report(y_true, y_pred, classes))
print()
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import torch
import torchvision
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from matplotlib import pyplot as plt
%matplotlib inline
from scipy.stats import entropy
from google.colab import drive
drive.mount('/content/drive')
path="/content/drive/MyDrive/Research/alternate_minimisation/type4_data/"
class MosaicDataset1(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list, mosaic_label,fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] , self.fore_idx[idx]
class SyntheticDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, x, y):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.x = x
self.y = y
#self.fore_idx = fore_idx
def __len__(self):
return len(self.y)
def __getitem__(self, idx):
return self.x[idx] , self.y[idx] #, self.fore_idx[idx]
train_data = np.load(path+"train_type4_data.npy",allow_pickle=True)
test_data = np.load(path+"test_type4_data.npy",allow_pickle=True)
data = np.load(path+"type_4_data.npy",allow_pickle=True)
train_mosaic_list_of_images = train_data[0]["mosaic_list"]
train_mosaic_label = train_data[0]["mosaic_label"]
train_fore_idx = train_data[0]["fore_idx"]
test_mosaic_list_of_images = test_data[0]["mosaic_list"]
test_mosaic_label = test_data[0]["mosaic_label"]
test_fore_idx = test_data[0]["fore_idx"]
X = data[0]["X"]
Y = data[0]["Y"]
batch = 250
tr_msd = MosaicDataset1(train_mosaic_list_of_images, train_mosaic_label, train_fore_idx)
train_loader = DataLoader( tr_msd,batch_size= batch ,shuffle=True)
batch = 250
tst_msd = MosaicDataset1(test_mosaic_list_of_images, test_mosaic_label, test_fore_idx)
test_loader = DataLoader( tst_msd,batch_size= batch ,shuffle=True)
dset = SyntheticDataset(X,Y)
dtloader = DataLoader(dset,batch_size =batch,shuffle=True )
```
**Focus Net**
```
class Module1(nn.Module):
def __init__(self):
super(Module1, self).__init__()
self.fc1 = nn.Linear(2, 100)
self.fc2 = nn.Linear(100, 1)
def forward(self, z):
x = torch.zeros([batch,9],dtype=torch.float64)
y = torch.zeros([batch,2], dtype=torch.float64)
x,y = x.to("cuda"),y.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
log_x = F.log_softmax(x,dim=1)
x = F.softmax(x,dim=1) # alphas
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None],z[:,i])
return y , x , log_x
def helper(self,x):
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
```
**Classification Net**
```
class Module2(nn.Module):
def __init__(self):
super(Module2, self).__init__()
self.fc1 = nn.Linear(2, 100)
self.fc2 = nn.Linear(100, 3)
def forward(self,y):
y = F.relu(self.fc1(y))
y = self.fc2(y)
return y
criterion = nn.CrossEntropyLoss()
def my_cross_entropy(x, y,alpha,log_alpha,k):
# log_prob = -1.0 * F.log_softmax(x, 1)
# loss = log_prob.gather(1, y.unsqueeze(1))
# loss = loss.mean()
loss = criterion(x,y)
#alpha = torch.clamp(alpha,min=1e-10)
b = -1.0* alpha * log_alpha
b = torch.mean(torch.sum(b,dim=1))
closs = loss
entropy = b
loss = (1-k)*loss + ((k)*b)
return loss,closs,entropy
```
```
def calculate_attn_loss(dataloader,what,where,criter,k):
what.eval()
where.eval()
r_loss = 0
cc_loss = 0
cc_entropy = 0
alphas = []
lbls = []
pred = []
fidices = []
with torch.no_grad():
for i, data in enumerate(dataloader, 0):
inputs, labels,fidx = data
lbls.append(labels)
fidices.append(fidx)
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
avg,alpha,log_alpha = where(inputs)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
alphas.append(alpha.cpu().numpy())
#ent = np.sum(entropy(alpha.cpu().detach().numpy(), base=2, axis=1))/batch
# mx,_ = torch.max(alpha,1)
# entropy = np.mean(-np.log2(mx.cpu().detach().numpy()))
# print("entropy of batch", entropy)
#loss = (1-k)*criter(outputs, labels) + k*ent
loss,closs,entropy = my_cross_entropy(outputs,labels,alpha,log_alpha,k)
r_loss += loss.item()
cc_loss += closs.item()
cc_entropy += entropy.item()
alphas = np.concatenate(alphas,axis=0)
pred = np.concatenate(pred,axis=0)
lbls = np.concatenate(lbls,axis=0)
fidices = np.concatenate(fidices,axis=0)
#print(alphas.shape,pred.shape,lbls.shape,fidices.shape)
analysis = analyse_data(alphas,lbls,pred,fidices)
return r_loss/i,cc_loss/i,cc_entropy/i,analysis
def analyse_data(alphas,lbls,predicted,f_idx):
'''
analysis data is created here
'''
batch = len(predicted)
amth,alth,ftpt,ffpt,ftpf,ffpf = 0,0,0,0,0,0
for j in range (batch):
focus = np.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
amth +=1
else:
alth +=1
if(focus == f_idx[j] and predicted[j] == lbls[j]):
ftpt += 1
elif(focus != f_idx[j] and predicted[j] == lbls[j]):
ffpt +=1
elif(focus == f_idx[j] and predicted[j] != lbls[j]):
ftpf +=1
elif(focus != f_idx[j] and predicted[j] != lbls[j]):
ffpf +=1
#print(sum(predicted==lbls),ftpt+ffpt)
return [ftpt,ffpt,ftpf,ffpf,amth,alth]
number_runs = 10
full_analysis =[]
FTPT_analysis = pd.DataFrame(columns = ["FTPT","FFPT", "FTPF","FFPF"])
k = 0.001
every_what_epoch = 10
for n in range(number_runs):
print("--"*40)
# instantiate focus and classification Model
torch.manual_seed(n)
where = Module1().double()
torch.manual_seed(n)
what = Module2().double()
where = where.to("cuda")
what = what.to("cuda")
# instantiate optimizer
optimizer_where = optim.Adam(where.parameters(),lr =0.001)
optimizer_what = optim.Adam(what.parameters(), lr=0.001)
#criterion = nn.CrossEntropyLoss()
acti = []
analysis_data = []
loss_curi = []
epochs = 1000
# calculate zeroth epoch loss and FTPT values
running_loss ,_,_,anlys_data= calculate_attn_loss(train_loader,what,where,criterion,k)
loss_curi.append(running_loss)
analysis_data.append(anlys_data)
print('epoch: [%d ] loss: %.3f' %(0,running_loss))
# training starts
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
what.train()
where.train()
if ((epoch) % (every_what_epoch*2) ) <= every_what_epoch-1 :
print(epoch+1,"updating what_net, where_net is freezed")
print("--"*40)
elif ((epoch) % (every_what_epoch*2)) > every_what_epoch-1 :
print(epoch+1,"updating where_net, what_net is freezed")
print("--"*40)
for i, data in enumerate(train_loader, 0):
# get the inputs
inputs, labels,_ = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_where.zero_grad()
optimizer_what.zero_grad()
# forward + backward + optimize
avg, alpha,log_alpha = where(inputs)
outputs = what(avg)
my_loss,_,_ = my_cross_entropy(outputs,labels,alpha,log_alpha,k)
# print statistics
running_loss += my_loss.item()
my_loss.backward()
if ((epoch) % (every_what_epoch*2) ) <= every_what_epoch-1 :
optimizer_what.step()
elif ( (epoch) % (every_what_epoch*2)) > every_what_epoch-1 :
optimizer_where.step()
# optimizer_where.step()
# optimizer_what.step()
#break
running_loss,ccloss,ccentropy,anls_data = calculate_attn_loss(train_loader,what,where,criterion,k)
analysis_data.append(anls_data)
print('epoch: [%d] loss: %.3f celoss: %.3f entropy: %.3f' %(epoch + 1,running_loss,ccloss,ccentropy))
loss_curi.append(running_loss) #loss per epoch
if running_loss<=0.005:
break
print('Finished Training run ' +str(n))
#break
analysis_data = np.array(analysis_data)
FTPT_analysis.loc[n] = analysis_data[-1,:4]/30
full_analysis.append((epoch, analysis_data))
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
images, labels,_ = data
images = images.double()
images, labels = images.to("cuda"), labels.to("cuda")
avg, alpha,log_alpha = where(images)
outputs = what(avg)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 3000 train images: %d %%' % ( 100 * correct / total))
a,b= full_analysis[0]
print(a)
cnt=1
for epoch, analysis_data in full_analysis:
analysis_data = np.array(analysis_data)
# print("="*20+"run ",cnt,"="*20)
plt.figure(figsize=(6,6))
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,0],label="ftpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,1],label="ffpt")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,2],label="ftpf")
plt.plot(np.arange(0,epoch+2,1),analysis_data[:,3],label="ffpf")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title("Training trends for run "+str(cnt))
plt.savefig(path+"what_where/every10/run"+str(cnt)+".png",bbox_inches="tight")
plt.savefig(path+"what_where/every10/run"+str(cnt)+".pdf",bbox_inches="tight")
cnt+=1
np.mean(np.array(FTPT_analysis),axis=0) #array([87.85333333, 5.92 , 0. , 6.22666667])
FTPT_analysis.to_csv(path+"what_where/FTPT_analysis_every10.csv",index=False)
FTPT_analysis
```
| github_jupyter |
# Bayesian optimization with parallel evaluation of an external objection function using Emukit
This tutorial will show you how to leverage Emukit to do Bayesian optimization on an external objective function that we can evaluate multiple times in parallel.
## Overview
By the end of the tutorial, you will be able to:
1. Generate batches $\{X_t | t \in 1..\}$ of objective function evaluation locations $\{x_i | x_i \in X_t\}$
2. Evaluate the objective function at these suggested locations in parallel $f(x_i)$
3. Use `asyncio` to implement the concurrency structure supporting this parallel evaluation
This tutorial requires basic familiarity with Bayesian optimization and concurrency. If you've never run Bayesian optimization using Emukit before, please refer to the [introductory tutorial](Emukit-tutorial-intro.ipynb) for more information. The concurrency used here is not particularly complicated, so you should be able to follow just fine without much more than an understanding of the [active object design pattern](https://en.wikipedia.org/wiki/Active_object).
The overview must start with the general imports and plots configuration
The overview section must finish with a Navigation that links to the main sections of the notebook
```
### General imports
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import colors as mcolors
### --- Figure config
colors = dict(mcolors.BASE_COLORS, **mcolors.CSS4_COLORS)
LEGEND_SIZE = 15
TITLE_SIZE = 25
AXIS_SIZE = 15
FIG_SIZE = (12,8)
```
### Navigation
1. [Define Objective Function](#1.-Define-objective-function)
2. [Setup BO & Run BO](#2.-Run-BO-using-parallel-evaluation-of-batched-suggestions)
3. [Conclusions](#3.-Conclusions)
## 1. Define objective function
```
# Specific imports that are used in a section should be loaded at the beginning of that section.
# It is ok if an import is repeated multiple times over the notebook
import time
import asyncio
import GPy
import emukit
import numpy as np
from math import pi
from emukit.test_functions.branin import (
branin_function as _branin_function,
)
### Define the cost and objective functions
_branin, _ps = _branin_function()
async def a_cost(x: np.ndarray):
# Cost function, defined arbitrarily
t = max(x.sum()/10, 0.1)
await asyncio.sleep(t)
async def a_objective(x: np.ndarray):
# Objective function
r = _branin(x)
await a_cost(x)
return r
async def demo_async_obj():
'''This function demonstrates a simple usage of the async objective function'''
# Configure
_x = [7.5, 12.5]
d = len(_x)
x = np.array(_x).reshape((1, d))
assert _ps.check_points_in_domain(x).all(), ("You configured a point outside the objective"
f"function's domain: {x} is outside {_ps.get_bounds()}")
# Execute
print(f"Input: x={x}")
t0 = time.perf_counter()
r = await a_objective(x)
t1 = time.perf_counter()
print(f"Output: result={r}")
print(f"Time elapsed: {t1-t0} sec")
await demo_async_obj()
```
## 2. Run BO using parallel evaluation of batched suggestions
```
from GPy.models import GPRegression
from emukit.model_wrappers import GPyModelWrapper
from emukit.core.initial_designs.latin_design import LatinDesign
from emukit.core import ParameterSpace, ContinuousParameter
from emukit.core.loop import UserFunctionWrapper, UserFunctionResult
from emukit.core.loop.stopping_conditions import FixedIterationsStoppingCondition
from emukit.core.optimization import GradientAcquisitionOptimizer
from emukit.bayesian_optimization.loops import BayesianOptimizationLoop
from emukit.bayesian_optimization.acquisitions import NegativeLowerConfidenceBound
import warnings
warnings.filterwarnings('ignore') # to quell the numerical errors in hyperparameter fitting
# Plotting stuff (from constrained optimization tutorial)
x1b, x2b = _ps.get_bounds()
plot_granularity = 50
x_1 = np.linspace(x1b[0], x1b[1], plot_granularity)
x_2 = np.linspace(x2b[0], x2b[1], plot_granularity)
x_1_grid, x_2_grid = np.meshgrid(x_1, x_2)
x_all = np.stack([x_1_grid.flatten(), x_2_grid.flatten()], axis=1)
y_all = _branin(x_all)
y_reshape = np.reshape(y_all, x_1_grid.shape)
x_best = np.array([(-pi,12.275), (pi,2.275), (9.425,2.475)])
def plot_progress(loop_state, batch_size: int):
plt.figure(figsize=FIG_SIZE)
plt.contourf(x_1, x_2, y_reshape)
plt.plot(loop_state.X[:-batch_size, 0], loop_state.X[:-batch_size, 1], linestyle='', marker='.', markersize=16, color='b')
plt.plot(loop_state.X[-batch_size:, 0], loop_state.X[-batch_size:, 1], linestyle='', marker='.', markersize=16, color='r')
plt.plot(x_best[:,0], x_best[:,1], linestyle='', marker='x', markersize=18, color='g')
plt.legend(['Previously evaluated points', 'Last evaluation', 'True best'])
plt.show()
async def async_run_bo():
# Configure
max_iter = 50
n_init = 6
batch_size = 6
beta = 0.1 # tradeoff parameter for NCLB acq. opt.
update_interval = 1 # how many results before running hyperparam. opt.
# Build Bayesian optimization components
space = _ps
design = LatinDesign(space)
X_init = design.get_samples(n_init)
input_coroutines = [a_objective(x.reshape((1,space.dimensionality))) for x in X_init]
_Y_init = await asyncio.gather(*input_coroutines, return_exceptions=True)
Y_init = np.concatenate(_Y_init)
model_gpy = GPRegression(X_init, Y_init)
model_gpy.optimize()
model_emukit = GPyModelWrapper(model_gpy)
acquisition_function = NegativeLowerConfidenceBound(model=model_emukit, beta=beta)
acquisition_optimizer = GradientAcquisitionOptimizer(space=space)
bo_loop = BayesianOptimizationLoop(
model = model_emukit,
space = space,
acquisition = acquisition_function,
acquisition_optimizer = acquisition_optimizer,
update_interval = update_interval,
batch_size = batch_size,
)
# Run BO loop
results = None
n = bo_loop.model.X.shape[0]
while n < max_iter:
print(f"Optimizing: n={n}")
# TODO use a different acquisition function because currently X_batch is 5 identical sugg.
# ^ only on occasion, apparently
X_batch = bo_loop.get_next_points(results)
coroutines = [a_objective(x.reshape((1, space.dimensionality))) for x in X_batch]
# TODO update model as soon as any result is available
# ^ as-is, only updates and makes new suggestions when all results come in
# TODO make suggestions cost-aware
_results = await asyncio.gather(*coroutines, return_exceptions=True)
Y_batch = np.concatenate(_results)
results = list(map(UserFunctionResult, X_batch, Y_batch))
n = n + len(results)
plot_progress(bo_loop.loop_state, batch_size)
final_result = bo_loop.get_results()
true_best = 0.397887
# rel_err = (final_result.minimum_value - true_best)/true_best
print(
"############################################################\n"
f"Minimum found at location: {final_result.minimum_location}\n"
f"\twith score: {final_result.minimum_value}\n"
f"True minima at:\n{x_best}\n"
f"\twith score: {true_best}\n"
# f"Relative error (%): {rel_err*100:.2f}\n"
"\tsource: https://www.sfu.ca/~ssurjano/branin.html\n"
"############################################################"
)
await async_run_bo()
```
## 3. Conclusions
1. I generated batches of suggestions using the `bo_loop.get_next_points()` function having configured the `bo_loop` with `batch_size` > 1
2. I evaluated these suggestions in parallel using `_Y_init = await asyncio.gather(*input_coroutines, return_exceptions=True)`
3. The `asyncio` structure is bare-bones:
1. The coroutines are prepared by mapping the async external objective function over the inputs
2. The coroutines are executed using `asyncio.gather`
| github_jupyter |
# Dataset cleansing
```
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
```
## 1. Load the (standardized) Data
```
patients = pd.read_csv("../../Data/standardized_patients.csv",index_col=0, header=0)
patients.dropna(inplace=True)
patients.head()
patients = patients.sample(frac=1).reset_index(drop=True)
```
## 2. Create the two dataset from the t-SNE analysis
### 2.1 Women in menopause condition
```
women_menopause = patients[patients['menopause']>1]
women_menopause.drop(columns=['sex','menopause'], axis=1, inplace=True)
women_menopause.head()
```
#### Create the X and y
```
women_X = women_menopause.copy()
women_X.drop("Class", axis=1, inplace=True)
print(women_X.shape)
women_y = women_menopause[['Class']].copy()
print(women_y.shape)
```
#### Split the data into Train, Test and Validation
The ratio in the splitting, in percentage, is 60/20/20
```
women_X_train_and_val, women_X_test, women_y_train_and_val, women_y_test = train_test_split(women_X, women_y, test_size=0.2, random_state=42)
women_X_train, women_X_val, women_y_train, women_y_val = train_test_split(women_X_train_and_val, women_y_train_and_val, test_size=0.25, random_state=42)
women_y_val[women_y_val['Class']==1].shape
women_y_train[women_y_train['Class']==1].shape
women_y_test[women_y_test['Class']==1].shape
```
#### Save to Files
```
patients.to_csv("../../Data/women_menopause/standardized_patients.csv")
women_X_train.to_csv("../../Data/women_menopause/X_train_total.csv")
women_y_train.to_csv("../../Data/women_menopause/y_train_total.csv")
women_X_test.to_csv("../../Data/women_menopause/X_test.csv")
women_y_test.to_csv("../../Data/women_menopause/y_test.csv")
women_X_val.to_csv("../../Data/women_menopause/X_val.csv")
women_y_val.to_csv("../../Data/women_menopause/y_val.csv")
```
### 2.2 Other patients
```
other_patients = patients[patients['menopause']<1]
other_patients.drop(columns=['menopause','HRT'], axis=1, inplace=True)
other_patients.head()
```
#### Create the X and y
```
other_patients_X = other_patients.copy()
other_patients_X.drop("Class", axis=1, inplace=True)
print(other_patients.shape)
other_patients_y = other_patients[['Class']].copy()
print(other_patients_y.shape)
```
#### Split the data into Train, Test and Validation
The ratio in the splitting, in percentage, is 60/20/20
```
other_patients_X_train_and_val, other_patients_X_test, other_patients_y_train_and_val, other_patients_y_test = train_test_split(other_patients_X, other_patients_y, test_size=0.2, random_state=42)
other_patients_X_train, other_patients_X_val, other_patients_y_train, other_patients_y_val = train_test_split(other_patients_X_train_and_val, other_patients_y_train_and_val, test_size=0.25, random_state=42)
other_patients_y_val[other_patients_y_val['Class']==1].shape
other_patients_y_train[other_patients_y_train['Class']==1].shape
other_patients_y_test[other_patients_y_test['Class']==1].shape
```
#### Save to Files
```
patients.to_csv("../../Data/other_patients/standardized_patients.csv")
other_patients_X_train.to_csv("../../Data/other_patients/X_train_total.csv")
other_patients_y_train.to_csv("../../Data/other_patients/y_train_total.csv")
other_patients_X_test.to_csv("../../Data/other_patients/X_test.csv")
other_patients_y_test.to_csv("../../Data/other_patients/y_test.csv")
other_patients_X_val.to_csv("../../Data/other_patients/X_val.csv")
other_patients_y_val.to_csv("../../Data/other_patients/y_val.csv")
```
| github_jupyter |
```
#Libs
import pandas as pd
import os
import sqlite3
```
# **Exportação de Dados**
```
#Abri o arquivo de dados
fileIO = 'C:\\Users\\User\\Desktop\\desafio\\DesafioETL01\\resources\\META.xlsx'
#Transforma o arquivo em Dataframe pandas
df = pd.read_excel(fileIO)
#Apresenta a tabela
df
```
# **Transformação de Dados**
```
#Retira as colunas com dados faltantes
df = df.dropna(axis=1, how='all')
#Apresenta os dados
df
#Retira colunas desnecessárias
df = df.drop(columns=['Erro?', 0])
#Apresenta a tabela
df
#Retira as linhas com dados faltantes
df = df.dropna(axis=0, how='all')
#Apresenta os dados
df
#Verifica se existe(m) valor(es) nulo(s)
print('\n', df.isnull().values.any())
df.info()
#Faz uma cópia dos dados
data = df.copy()
#Apresenta os dados
data
```
# **Carregamento de Dados**
```
#Verifica se já existe um banco de dados
try:
os.remove('resources/db/bancolocal.db')
except OSError:
pass
#Conexão com o banco de dados
con = sqlite3.connect('resources/db/bancolocal.db')
#Criando um cursor
inst = con.cursor()
#Verifica se tabela já existe
SQL = '''
DROP TABLE IF EXISTS meta
'''
#Executa a instrução SQL no cursor
inst.execute(SQL)
#Grava a informação no db
con.commit()
#Linguagem SQL (Criação da Tabela meta)
SQL = '''
CREATE TABLE meta (
anometa smallint,
mesmeta smallint,
vendedorcodigo varchar(255),
vendedornome varchar(255),
valormeta double precision
);
'''
#Executa a instrução SQL no cursor
inst.execute(SQL)
#Definido informações estáticas
anometa = data.iloc[0,1]
#transforma as colunas em dataframe
colunas = pd.DataFrame(data.columns)
#Captura somente os meses
for i in range(2,14,1):
mesmeta = colunas.iloc[i,0]
#captura cada instância
for j in range(1,18,1):
vendedorcodigo = data.iloc[j,0]
vendedornome = data.iloc[j,1]
valormeta = data.iloc[j,i]
#Lista de dados
info = [anometa, mesmeta, vendedorcodigo, vendedornome, valormeta]
#Alimenta os dados na tabela
inst.execute("INSERT INTO meta VALUES(?,?,?,?,?)", info)
#Grava a informação no db
con.commit()
#Mostra os valores alimentados na tabela
inst.execute('SELECT * FROM meta')
itemAll = inst.fetchall()
for item in itemAll:
print(item)
SQL2 = '''
select
vendedorcodigo,
vendedornome,
sum(case when mesmeta = 'Janeiro' then valormeta end) as Janeiro,
sum(case when mesmeta = 'Fevereiro' then valormeta end) as Fevereiro,
sum(case when mesmeta = 'Março' then valormeta end) as Março,
sum(case when mesmeta = 'Abril' then valormeta end) as Abril,
sum(case when mesmeta = 'Maio' then valormeta end) as Maio,
sum(case when mesmeta = 'Junho' then valormeta end) as Junho,
sum(case when mesmeta = 'Julho' then valormeta end) as Julho,
sum(case when mesmeta = 'Agosto' then valormeta end) as Agosto,
sum(case when mesmeta = 'Setembro' then valormeta end) as Setembro,
sum(case when mesmeta = 'Outubro' then valormeta end) as Outubro,
sum(case when mesmeta = 'Novembro' then valormeta end) as Novembro,
sum(case when mesmeta = 'Dezembro' then valormeta end) as Dezembro
from meta
group by vendedorcodigo, vendedornome;
'''
data.to_csv("resources\meta.csv", index=False)
```
| github_jupyter |
```
import cobra
import pandas as pd
from util.manipulation import load_latest_model
import numpy as np
import matplotlib.pyplot as plt
```
### Tutorial 1: General manipulations
This tutorial covers the general manipulations that can performed on the *i*JL208 model. The following basic manipulations are executed:
1- Load the latest version of the model and display the number of reactions, genes and metabolites
2- Get the optimal solution and growth rate prediction
3- Get and display a predicted flux state, the number of used reactions
4- Formulate a single gene essentiality prediction
5- Using parsimonious FBA (pFBA) for results more similar to expression data
```
#Use the util function to load the latest model from the model_versions folder
model = load_latest_model()
#Display the number of reactions, genes and metabolites in the model
print(f"The {model.name} model contains {len(model.reactions)} reactions, {len(model.genes)} genes and {len(model.metabolites)} metabolites")
for g in model.reactions.PDH.genes:
print(g.id, g.notes)
for r in model.genes.Mfl041.reactions:
print('#########')
print(r.id, r.name)
print(r.reaction)
print(r.metabolites)
l = []
for g in model.genes:
if len(g.reactions) > 1:
l.append(g)
print(len(l))
for g in model.genes:
print(g.id, g.reactions)
for r in g.reactions:
print(r)
print(f"les metabolites associes a la reaction {r.metabolites}")
break
gene_associated_reactions = []
for r in model.reactions:
if len(r.genes) > 0:
gene_associated_reactions.append(r)
print(len(gene_associated_reactions))
model.reactions.PDH.gene_reaction_rule = 'Mfl039 and Mfl040 and Mfl041 and Mfl042'
model.reactions.PDH.gene_reaction_rule
cobra.io.save_json_model(model,'iJL208_PDH.json')
model.solver
#Get the predicted growth rate and validate that the model solves correctly
solution = model.optimize()
if solution.status == 'optimal':
print(f"The {model.name} has an optimal solution and the predicted growth rate is: {solution.objective_value}")
else:
raise ValueError(f"The model does not have an optimal solution, ensure the ")
solution.fluxes['PDH']
solution.fluxes['EX_sucr_e']
solution.fluxes.idxmax()
model.reactions.ENO.reaction
model.reactions.ENO.genes
model.metabolites.pep_c.name
#Plot the predicted flux state
predicted_fluxes = pd.DataFrame({'Predicted fluxes':solution.fluxes.to_list()},index=solution.fluxes.index.to_list())
predicted_fluxes.plot(kind='hist')
#Get the number of used/unused reactions, those are the reactions for which no flux is predicted
used_reactions = predicted_fluxes[predicted_fluxes['Predicted fluxes']!=0]
unused_reactions = predicted_fluxes[predicted_fluxes['Predicted fluxes']==0]
print(f"The model has {len(used_reactions)} used reactions and {len(unused_reactions)} unused reactions")
#Formulate a single gene essentiality prediction
from cobra.flux_analysis import single_gene_deletion
single_gene_essentiality_prediction = single_gene_deletion(model)
single_gene_essentiality_prediction
#Get the list of essential gene
#Define a threshold for essential as 50% of the original growth rate
essentiality_threshold = solution.objective_value * 0.5
#Essential genes have a predicted growth rate inferior or equal to the threshold or NaN
essential_genes = [list(g)[0] for g in single_gene_essentiality_prediction[(single_gene_essentiality_prediction['growth']<=essentiality_threshold)\
|(single_gene_essentiality_prediction['growth'].isna())].index]
#Non essential genes have a predicted growth rate higher than threshold
non_essential_genes = [list(g)[0] for g in single_gene_essentiality_prediction[single_gene_essentiality_prediction['growth']> essentiality_threshold].index]
#Display the number of essential and non essential genes
print(f"The model has {len(essential_genes)} essential genes and {len(non_essential_genes)} non essentials")
#Optimize the model using pFBA
from cobra.flux_analysis import pfba
#Generate the pFBA solution
pfba_solution = pfba(model)
#This solution contains the same elements as a regular FBA solution
#--> objective_value, fluxes, shadow_prices, reduced_costs and status
#Now compare the two flux predictions
#Plot the predicted flux state
pfba_predicted_fluxes = pd.DataFrame({'pFBA predicted fluxes':pfba_solution.fluxes.to_list()},index=pfba_solution.fluxes.index.to_list())
#Compare the two flux distributions
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
x = predicted_fluxes['Predicted fluxes'].tolist()
y = pfba_predicted_fluxes['pFBA predicted fluxes'].tolist()
ax.scatter(x=x,y=y)
#Display on log scale
ax.set_yscale('log')
ax.set_xscale('log')
ax.set_xlim(1e-3,1e2)
ax.set_ylim(1e-3,1e2)
ax.set_xlabel('FBA')
ax.set_ylabel('pFBA')
#Get the flux predictions difference
all_fluxes = pd.concat([predicted_fluxes,pfba_predicted_fluxes],axis=1,sort=True)
all_fluxes['difference'] = all_fluxes['pFBA predicted fluxes'] - all_fluxes['Predicted fluxes']
#Display
all_fluxes[all_fluxes['difference'] > 1e-9]
```
#### End of the first tutorial
In this tutorial, only static observations of the model were performed and slighlty compared to one another. In Tutorial 2, we will apply modifications to the model, generate and compare new predictions resulting from these modifications.
| github_jupyter |
```
# Only to use if Jupyter Notebook
# %load_ext google.datalab.kernel
from __future__ import division
import os, hashlib, math
import numpy as np
import pandas as pd
import sklearn.metrics as metrics
import seaborn as sns
import matplotlib.pyplot as plt
import google.datalab.contrib.mlworkbench.commands
import google.datalab.ml as ml
```
This Notebook shows you how to perform the basic steps in order to build, train and deploy a model on Google Cloud Platform using ML Toolbox
1. Collect data
2. Organize data
3. Design the model
4. Train and generate the model
5. Deploy the model
Note that we will build, train and deploy our model on this machine only as our dataset is small enough and a model created locally can still be deployed on Google ML Engine.
```
WORKING_FOLDER = 'priority'
CSV_FILE = "issues.csv"
!rm -rf $WORKING_FOLDER
!mkdir $WORKING_FOLDER
```
# 1 of 7 - Collect data
In a use case like this, data can be collected in different ways usually through dump of your database or export from your CRM into CSV.
The data that we have collected is available on Google Cloud Storage as gs://solutions-public-assets/smartenup-helpdesk/ml/data.csv (public dataset).
Note that in some cases, data might be saved in BigQuery (which is both a storage and queryuing engine) which is perfectly fine and would actually facilitate the filtering of the data specially with big datasets.
```
# Copy data from the cloud to this instance
!gsutil cp gs://solutions-public-assets/smartenup-helpdesk/ml/issues.csv $CSV_FILE
# Read data from csv into a Panda dataframe
df_data = pd.read_csv(CSV_FILE, dtype=str)
print '%d rows' % len(df_data)
df_data.head()
```
We need to make sure that we do not have any duplicate. If that was the case some identical rows could be found in the training set and validation set for example which would have an impact on the model training (remember, validation and training set can't overlap)
```
df_data = df_data.drop_duplicates(df_data.columns.difference(['ticketid']))
print '%d rows' % len(df_data)
```
# 2 of 7 - Organize data
## Filter data
We keep only the columns that we are interested and discard some others:
- ownerid because we won't know its value when doing a prediction
- priority as it is a value that we will predict in another Notebook
- statisfaction as it is not a value that we know when doing a prediction. It might also be a value we woud like to predict later
```
def transform_data(df):
# Lists the column names that we want to keep from our dataframe.
interesting_columns = ['ticketid', 'seniority', 'experience', 'category', 'type', 'impact', 'priority']
# Filters the dataframe to keep only the relevant data and return the dataframe.
df = df[interesting_columns]
return df
df_data = transform_data(df_data)
# Displays the new dataframe.
print '%d rows' % len(df_data)
df_data.head()
```
## Create datasets
Here we create training and test datasets on a 80/20 basis. To keep consistency for every load of data we use a column that follows these two requirements:
- Is a unique identifer for each row
- Will not be used as a training input
ticketid is a good candidate
```
def is_test_set(identifier, test_ratio, hash):
h = int(hash(identifier.encode('ascii')).hexdigest()[-7:], 16)
return (h/0xFFFFFFF) < test_ratio
def create_datasets(df, id_column, test_ratio=0.2, hash=hashlib.md5):
ids = df[id_column]
ids_test_set = ids.apply(lambda x: is_test_set(x, test_ratio, hash))
return df.loc[~ids_test_set], df.loc[ids_test_set]
df_train, df_eval = create_datasets(df_data, 'ticketid')
# Set paths for CSV datasets
training_data_path = './{}/train.csv'.format(WORKING_FOLDER)
test_data_path = './{}/eval.csv'.format(WORKING_FOLDER)
# Write Panda Dataframes to CSV files
df_train.to_csv(training_data_path, header=False, index=False)
df_eval.to_csv(test_data_path, header=False, index=False)
```
## Explore
One of the most important part of Machine Learning is to explore the data before building a model. A few things will help improving your model quality such as
- Normalization
- Look at feature correlation
- Feature crossing
- ...
Because the main goal of this Notebook and solution is to show how to build and deploy a model for serverless enrichment, we won't spend too much time here (also because the provided dataset is fake) but keep in mind that this is not a part that should be ignored in a real world example.
One important thing in Classification though is to have a balanced set of Labels.
```
print "Training set:\n{}".format(df_train.priority.value_counts())
print "Eval set:\n{}".format(df_eval.priority.value_counts())
```
# 3 of 7: Analyse
ML Workbench comes with a pre-buit function that analyzes training data and generate stats, such as min/max/mean for numeric values, vocabulary for text columns. Note that if cloud is set to True, the function leverages BigQuery making the switch from small dataset to big data seamless.
```
!rm -rf ./priority/analysis
%%ml dataset create
format: csv
train: ./priority/train.csv
eval: ./priority/eval.csv
name: issues_data_priority
schema:
- name: ticketid
type: STRING
- name: seniority
type: FLOAT
- name: experience
type: STRING
- name: category
type: STRING
- name: type
type: STRING
- name: impact
type: STRING
- name: priority
type: STRING
%%ml analyze
output: ./priority/analysis
data: $issues_data_priority
features:
ticketid:
transform: key
seniority:
transform: identity
experience:
transform: one_hot
category:
transform: one_hot
type:
transform: one_hot
impact:
transform: one_hot
priority:
transform: target
```
# 4 of 7: Transform
This section is optional but can be an important step when dealing with big data. While the Analysis phase provides enough details for the training step to append, the transform phase creates tfRecord files which is required for Tensorflow processing. Doing it now make sure that the training step can start from the preprocessed data and does not have to do this for each row for every pass of the data which is not recommended when handling text or image data.
```
!rm -rf ./priority/transform
%%ml transform
output: ./priority/transform
analysis: ./priority/analysis
data: $issues_data_priority
%%ml dataset create
format: transformed
name: issues_data_priority_transformed
train: ./priority/transform/train-*
eval: ./priority/transform/eval-*
```
# 5 of 7 Train
This steps leverages Tensorflow canned models in the background without you having to write any code.
```
!rm -rf ./priority/train
%%ml train
output: ./priority/train
analysis: ./priority/analysis
data: $issues_data_priority_transformed
model_args:
model_args:
model: dnn_classification
max-steps: 5000
hidden-layer-size1: 256
hidden-layer-size2: 128
train-batch-size: 8
eval-batch-size: 100
learning-rate: 0.001
tensorboard_pid = ml.TensorBoard.start('./{}/train'.format(WORKING_FOLDER))
ml.TensorBoard.stop(tensorboard_pid)
```
# 6 of 7
In this section, we will test our model to see how well it performs. For demo purposes, we are reusing the evaluation dataset. Consider using a 3rd separated dataset for production cases.
```
!rm -rf ./priority/evalme
%%ml batch_predict
model: ./priority/train/evaluation_model/
output: ./priority/evalme
format: csv
data:
csv: ./priority/eval.csv
!head -n 2 ./priority/evalme/predict_results_eval.csv
ml.ConfusionMatrix.from_csv(
input_csv='./{}/evalme/predict_results_eval.csv'.format(WORKING_FOLDER),
schema_file='./{}/evalme/predict_results_schema.json'.format(WORKING_FOLDER)
).plot()
```
We see on this confusion matrix that the predictions are not too bad, the good ones being on the diagonale. Results might improve by doing extra prepration steps such as:
- Feature crossing
- Correlation analysis
- Normalization
# 7 of 7 - Deploy the Model
```
model_name = 'mdl_helpdesk_priority'
model_version = 'v1'
storage_bucket = 'gs://' + google.datalab.Context.default().project_id + '-datalab-workspace/'
storage_region = 'us-central1'
# Check that we have the model files created by the training
!ls -R train/model
# Create a model
!gcloud ml-engine models create {model_name} --regions {storage_region}
# Create a staging bucket required to write staging files
# When creating a model from local files
staging_bucket = 'gs://' + google.datalab.Context.default().project_id + '-dtlb-staging-resolution'
!gsutil mb -c regional -l {storage_region} {staging_bucket}
# Create our version of the model.
!gcloud ml-engine versions create {model_version} --model {model_name} --origin train/model --staging-bucket {staging_bucket}
```
That version that is deployed is the one that you will be able to update automatically in a production environment if you need to update your model daily/weekly/monthly for example
| github_jupyter |
# Information about the system & environment
```
from datetime import datetime
import os
import platform
import psutil
import sh
import sys
```
Note that `psutil` is not in Python's standard library.
## Operating system
Using the `platform` module, it is easy to obtain information on the platform the Python interpreter is running on.
Information about the machine name:
```
platform.node()
```
The architecture and hardware:
```
platform.processor()
sys.byteorder
os.cpu_count()
os.sched_getaffinity(0)
```
The operating system:
```
platform.system()
platform.release()
platform.version()
platform.linux_distribution()
platform.platform()
```
## Numerics
The properties of floating point numbers can be obtained easily from the `sys.floatinfo` object.
The largest floating point value that can be represented, the smallest positive non-zero value:
```
print(sys.float_info.max, sys.float_info.min)
```
The number of significant digits of a floating point value:
```
sys.float_info.dig
```
## Processes
Detailed information is available on the processes running on the system.
```
for process in psutil.process_iter():
if 'bash' in process.name():
cpu_times = process.cpu_times()
thread_str = f'threads: {process.num_threads()}'
cpu_str = f'user: {cpu_times.user}, sys: {cpu_times.system}'
print(f'{process.pid}: {process.name()} ({thread_str}, {cpu_str})')
```
CPU times are cumulative over the process' life time.
```
for process in psutil.process_iter():
if process.cpu_times().user > 2.5:
print(f'{process.name()}: {process.cpu_times().user}')
```
It is easy to kill processes, so you might want to be careful.
```
sleep = sh.sleep(120, _bg=True)
for process in psutil.process_iter():
if 'sleep' in process.name():
print(process)
name = 'sleep'
killed_pids = []
for process in psutil.process_iter():
if name == process.name():
print(f'killing {name}...')
killed_pids.append(process.pid)
process.kill()
print('killed: ', ', '.join(map(str, killed_pids)))
```
## Users
You can retrieve information on the users on the system as well.
```
for user in psutil.users():
started = datetime.strftime(datetime.fromtimestamp(user.started), '%Y-%m-%d %H:%M:%S')
print(f'{user.name}: {started}')
```
## Performance
The `psutil` module makes quite some interesting statistics related to system performance available. This can be useful when writing monitoring tools.
The cumulative times for user, nice, system and so on are readily available.
```
psutil.cpu_times()
```
Memory usage can be queried.
```
psutil.virtual_memory()
```
Disk I/O measures such as the total IOP and read/write sizes in byte are easy to obain.
```
psutil.disk_io_counters()
```
Network I/O can similarly be monitored.
```
psutil.net_io_counters()
```
Disk usage for all partitions can be queried.
```
for partition in psutil.disk_partitions():
mountpoint = partition.mountpoint
if 'snap' not in mountpoint:
print(f'{mountpoint}: {psutil.disk_usage(mountpoint).percent:.1f}')
```
| github_jupyter |
# What are `LightCurve` objects?
`LightCurve` objects are data objects which encapsulate the brightness of a star over time. They provide a series of common operations, for example folding, binning, plotting, etc. There are a range of subclasses of `LightCurve` objects specific to telescopes, including `KeplerLightCurve` for Kepler and K2 data and `TessLightCurve` for TESS data.
Although *lightkurve* was designed with Kepler, K2 and TESS in mind, these objects can be used for a range of astronomy data.
You can create a `LightCurve` object from a `TargetPixelFile` object using Simple Aperture Photometry (see our tutorial for more information on Target Pixel Files [here](http://lightkurve.keplerscience.org/tutorials/1.02-target-pixel-files.html). Aperture Photometry is the simple act of summing up the values of all the pixels in a pre-defined aperture, as a function of time. By carefully choosing the shape of the aperture mask, you can avoid nearby contaminants or improve the strength of the specific signal you are trying to measure relative to the background.
To demonstrate, lets create a `KeplerLightCurve` from a `KeplerTargetPixelFile`.
```
from lightkurve import search_targetpixelfile
# First we open a Target Pixel File from MAST, this one is already cached from our previous tutorial!
tpf = search_targetpixelfile('KIC 6922244', quarter=4).download()
# Then we convert the target pixel file into a light curve using the pipeline-defined aperture mask.
lc = tpf.to_lightcurve(aperture_mask=tpf.pipeline_mask)
```
We've built a new `KeplerLightCurve` object called `lc`. Note in this case we've passed an **aperture_mask** to the `to_lightcurve` method. The default is to use the *Kepler* pipeline aperture. (You can pass your own aperture, which is a boolean `numpy` array.) By summing all the pixels in the aperture we have created a Simple Aperture Photometry (SAP) lightcurve.
`KeplerLightCurve` has many useful functions that you can use. As with Target Pixel Files you can access the meta data very simply:
```
lc.meta['mission']
lc.meta['quarter']
```
And you still have access to time and flux attributes. In a light curve, there is only one flux point for every time stamp:
```
lc.time
lc.flux
```
You can also check the "CDPP" noise metric of the lightcurve using the built in method:
```
lc.estimate_cdpp()
```
Now we can use the built in `plot` function on the `KeplerLightCurve` object to plot the time series. You can pass `plot` any keywords you would normally pass to `matplotlib.pyplot.plot`.
```
%matplotlib inline
lc.plot();
```
There are a set of useful functions in `LightCurve` objects which you can use to work with the data. These include:
* `flatten()`: Remove long term trends using a [Savitzky–Golay filter](https://en.wikipedia.org/wiki/Savitzky%E2%80%93Golay_filter)
* `remove_outliers()`: Remove outliers using simple sigma clipping
* `remove_nans()`: Remove infinite or NaN values (these can occur during thruster firings)
* `fold()`: Fold the data at a particular period
* `bin()`: Reduce the time resolution of the array, taking the average value in each bin.
We can use these simply on a light curve object
```
flat_lc = lc.flatten(window_length=401)
flat_lc.plot();
folded_lc = flat_lc.fold(period=3.5225)
folded_lc.plot();
binned_lc = folded_lc.bin(time_bin_size=0.01)
binned_lc.plot();
```
Or we can do these all in a single (long) line!
```
lc.remove_nans().flatten(window_length=401).fold(period=3.5225).bin(time_bin_size=0.01).plot();
```
| github_jupyter |
```
from __future__ import print_function
import os
import sys
import numpy as np
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.utils import to_categorical
from keras.layers import Dense, Input, GlobalMaxPooling1D
from keras.layers import Conv1D, MaxPooling1D, Embedding
from keras.models import Model
BASE_DIR = ''
GLOVE_DIR = os.path.join(BASE_DIR, 'glove.6B')
TEXT_DATA_DIR = os.path.join(BASE_DIR, '20_newsgroup')
MAX_SEQUENCE_LENGTH = 1000
MAX_NUM_WORDS = 20000
EMBEDDING_DIM = 100
VALIDATION_SPLIT = 0.2
print('Indexing word vectors.')
embeddings_index = {}
with open(os.path.join(GLOVE_DIR, 'glove.6B.100d.txt')) as f:
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
print('Found %s word vectors.' % len(embeddings_index))
# second, prepare text samples and their labels
print('Processing text dataset')
texts = [] # list of text samples
labels_index = {} # dictionary mapping label name to numeric id
labels = [] # list of label ids
for name in sorted(os.listdir(TEXT_DATA_DIR)):
path = os.path.join(TEXT_DATA_DIR, name)
if os.path.isdir(path):
label_id = len(labels_index)
labels_index[name] = label_id
for fname in sorted(os.listdir(path)):
if fname.isdigit():
fpath = os.path.join(path, fname)
args = {} if sys.version_info < (3,) else {'encoding': 'latin-1'}
with open(fpath, **args) as f:
t = f.read()
i = t.find('\n\n') # skip header
if 0 < i:
t = t[i:]
texts.append(t)
labels.append(label_id)
print('Found %s texts.' % len(texts))
okenizer = Tokenizer(num_words=MAX_NUM_WORDS)
tokenizer.fit_on_texts(texts)
sequences = tokenizer.texts_to_sequences(texts)
word_index = tokenizer.word_index
print('Found %s unique tokens.' % len(word_index))
data = pad_sequences(sequences, maxlen=MAX_SEQUENCE_LENGTH)
labels = to_categorical(np.asarray(labels))
print('Shape of data tensor:', data.shape)
print('Shape of label tensor:', labels.shape)
# split the data into a training set and a validation set
indices = np.arange(data.shape[0])
np.random.shuffle(indices)
data = data[indices]
labels = labels[indices]
num_validation_samples = int(VALIDATION_SPLIT * data.shape[0])
x_train = data[:-num_validation_samples]
y_train = labels[:-num_validation_samples]
x_val = data[-num_validation_samples:]
y_val = labels[-num_validation_samples:]
num_words = min(MAX_NUM_WORDS, len(word_index) + 1)
embedding_matrix = np.zeros((num_words, EMBEDDING_DIM))
for word, i in word_index.items():
if i >= MAX_NUM_WORDS:
continue
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
# words not found in embedding index will be all-zeros.
embedding_matrix[i] = embedding_vector
# load pre-trained word embeddings into an Embedding layer
# note that we set trainable = False so as to keep the embeddings fixed
embedding_layer = Embedding(num_words,
EMBEDDING_DIM,
weights=[embedding_matrix],
input_length=MAX_SEQUENCE_LENGTH,
trainable=False)
print('Training model.')
sequence_input = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
embedded_sequences = embedding_layer(sequence_input)
x = Conv1D(128, 5, activation='relu')(embedded_sequences)
x = MaxPooling1D(5)(x)
x = Conv1D(128, 5, activation='relu')(x)
x = MaxPooling1D(5)(x)
x = Conv1D(128, 5, activation='relu')(x)
x = GlobalMaxPooling1D()(x)
x = Dense(128, activation='relu')(x)
preds = Dense(len(labels_index), activation='softmax')(x)
model = Model(sequence_input, preds)
model.compile(loss='categorical_crossentropy',
optimizer='rmsprop',
metrics=['acc'])
model.fit(x_train, y_train,
batch_size=128,
epochs=10,
validation_data=(x_val, y_val))
```
| github_jupyter |
###Set up working directory
```
cd ~/Desktop/SSUsearch/
mkdir -p ./workdir
#check seqfile files to process in data directory (make sure you still remember the data directory)
!ls ./data/test/data
```
#README
## This part of pipeline search for the SSU rRNA gene fragments, classify them, and extract reads aligned specific region. It is also heavy lifting part of the whole pipeline (more cpu will help).
## This part works with one seqfile a time. You just need to change the "Seqfile" and maybe other parameters in the two cells bellow.
## To run commands, click "Cell" then "Run All". After it finishes, you will see "\*** pipeline runs successsfully :)" at bottom of this pape.
##If your computer has many processors, there are two ways to make use of the resource:
1. Set "Cpu" higher number.
2. make more copies of this notebook (click "File" then "Make a copy" in menu bar), so you can run the step on multiple files at the same time.
(Again we assume the "Seqfile" is quality trimmed.)
###Here we will process one file at a time; set the "Seqfile" variable to the seqfile name to be be processed
###First part of seqfile basename (separated by ".") will be the label of this sample, so named it properly.
e.g. for "/usr/local/notebooks/data/test/data/1c.fa", "1c" will the label of this sample.
```
Seqfile='./data/test/data/2c.fa'
```
###Other parameters to set
```
Cpu='1' # number of maxixum threads for search and alignment
Hmm='./data/SSUsearch_db/Hmm.ssu.hmm' # hmm model for ssu
Gene='ssu'
Script_dir='./scripts'
Gene_model_org='./data/SSUsearch_db/Gene_model_org.16s_ecoli_J01695.fasta'
Ali_template='./data/SSUsearch_db/Ali_template.silva_ssu.fasta'
Start='577' #pick regions for de novo clustering
End='727'
Len_cutoff='100' # min length for reads picked for the region
Gene_tax='./data/SSUsearch_db/Gene_tax.silva_taxa_family.tax' # silva 108 ref
Gene_db='./data/SSUsearch_db/Gene_db.silva_108_rep_set.fasta'
Gene_tax_cc='./data/SSUsearch_db/Gene_tax_cc.greengene_97_otus.tax' # greengene 2012.10 ref for copy correction
Gene_db_cc='./data/SSUsearch_db/Gene_db_cc.greengene_97_otus.fasta'
# first part of file basename will the label of this sample
import os
Filename=os.path.basename(Seqfile)
Tag=Filename.split('.')[0]
import os
New_path = '{}:{}'.format('~/Desktop/SSUsearch/external_tools/bin/', os.environ['PATH'])
Hmm=os.path.abspath(Hmm)
Seqfile=os.path.abspath(Seqfile)
Script_dir=os.path.abspath(Script_dir)
Gene_model_org=os.path.abspath(Gene_model_org)
Ali_template=os.path.abspath(Ali_template)
Gene_tax=os.path.abspath(Gene_tax)
Gene_db=os.path.abspath(Gene_db)
Gene_tax_cc=os.path.abspath(Gene_tax_cc)
Gene_db_cc=os.path.abspath(Gene_db_cc)
os.environ.update(
{'PATH':New_path,
'Cpu':Cpu,
'Hmm':os.path.abspath(Hmm),
'Gene':Gene,
'Seqfile':os.path.abspath(Seqfile),
'Filename':Filename,
'Tag':Tag,
'Script_dir':os.path.abspath(Script_dir),
'Gene_model_org':os.path.abspath(Gene_model_org),
'Ali_template':os.path.abspath(Ali_template),
'Start':Start,
'End':End,
'Len_cutoff':Len_cutoff,
'Gene_tax':os.path.abspath(Gene_tax),
'Gene_db':os.path.abspath(Gene_db),
'Gene_tax_cc':os.path.abspath(Gene_tax_cc),
'Gene_db_cc':os.path.abspath(Gene_db_cc)})
!echo "*** make sure: parameters are right"
!echo "Seqfile: $Seqfile\nCpu: $Cpu\nFilename: $Filename\nTag: $Tag"
cd workdir
mkdir -p $Tag.ssu.out
### start hmmsearch
%%bash
echo "*** hmmsearch starting"
time hmmsearch --incE 10 --incdomE 10 --cpu $Cpu \
--domtblout $Tag.ssu.out/$Tag.qc.$Gene.hmmdomtblout \
-o /dev/null -A $Tag.ssu.out/$Tag.qc.$Gene.sto \
$Hmm $Seqfile
echo "*** hmmsearch finished"
!python $Script_dir/get-seq-from-hmmout.py \
$Tag.ssu.out/$Tag.qc.$Gene.hmmdomtblout \
$Tag.ssu.out/$Tag.qc.$Gene.sto \
$Tag.ssu.out/$Tag.qc.$Gene
```
### Pass hits to mothur aligner
```
%%bash
echo "*** Starting mothur align"
cat $Gene_model_org $Tag.ssu.out/$Tag.qc.$Gene > $Tag.ssu.out/$Tag.qc.$Gene.RFadded
# mothur does not allow tab between its flags, thus no indents here
time mothur "#align.seqs(candidate=$Tag.ssu.out/$Tag.qc.$Gene.RFadded, template=$Ali_template, threshold=0.5, flip=t, processors=$Cpu)"
rm -f mothur.*.logfile
```
### Get aligned seqs that have > 50% matched to references
```
!python $Script_dir/mothur-align-report-parser-cutoff.py \
$Tag.ssu.out/$Tag.qc.$Gene.align.report \
$Tag.ssu.out/$Tag.qc.$Gene.align \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter \
0.5
!python $Script_dir/remove-gap.py $Tag.ssu.out/$Tag.qc.$Gene.align.filter $Tag.ssu.out/$Tag.qc.$Gene.align.filter.fa
```
### Search is done here (the computational intensive part). Hooray!
- \$Tag.ssu.out/\$Tag.qc.\$Gene.align.filter:
aligned SSU rRNA gene fragments
- \$Tag.ssu.out/\$Tag.qc.\$Gene.align.filter.fa:
unaligned SSU rRNA gene fragments
### Extract the reads mapped 150bp region in V4 (577-727 in *E.coli* SSU rRNA gene position) for unsupervised clustering
```
!python $Script_dir/region-cut.py $Tag.ssu.out/$Tag.qc.$Gene.align.filter $Start $End $Len_cutoff
!mv $Tag.ssu.out/$Tag.qc.$Gene.align.filter."$Start"to"$End".cut.lenscreen $Tag.ssu.out/$Tag.forclust
```
### Classify SSU rRNA gene seqs using SILVA
```
%%bash
rm -f $Tag.ssu.out/$Tag.qc.$Gene.align.filter.silva_taxa_family*.taxonomy
mothur "#classify.seqs(fasta=$Tag.ssu.out/$Tag.qc.$Gene.align.filter.fa, template=$Gene_db, taxonomy=$Gene_tax, cutoff=50, processors=$Cpu)"
mv $Tag.ssu.out/$Tag.qc.$Gene.align.filter.silva_taxa_family*.taxonomy \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.silva.taxonomy
!python $Script_dir/count-taxon.py \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.silva.taxonomy \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.silva.taxonomy.count
!rm -f mothur.*.logfile
```
### Classify SSU rRNA gene seqs with Greengene for copy correction later
```
%%bash
rm -f $Tag.ssu.out/$Tag.qc.$Gene.align.filter.greengene_97_otus*.taxonomy
mothur "#classify.seqs(fasta=$Tag.ssu.out/$Tag.qc.$Gene.align.filter.fa, template=$Gene_db_cc, taxonomy=$Gene_tax_cc, cutoff=50, processors=$Cpu)"
mv $Tag.ssu.out/$Tag.qc.$Gene.align.filter.greengene_97_otus*.taxonomy \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.gg.taxonomy
!python $Script_dir/count-taxon.py \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.gg.taxonomy \
$Tag.ssu.out/$Tag.qc.$Gene.align.filter.wang.gg.taxonomy.count
!rm -f mothur.*.logfile
# check the output directory
!ls $Tag.ssu.out
```
### This part of pipeline (working with one sequence file) finishes here. Next we will combine samples for community analysis (see unsupervised analysis).
Following are files useful for community analysis:
* 1c.577to727: aligned fasta file of seqs mapped to target region for de novo clustering
* 1c.qc.ssu.align.filter: aligned fasta file of all SSU rRNA gene fragments
* 1c.qc.ssu.align.filter.wang.gg.taxonomy: Greengene taxonomy (for copy correction)
* 1c.qc.ssu.align.filter.wang.silva.taxonomy: SILVA taxonomy
```
!echo "*** pipeline runs successsfully :)"
```
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os
import pickle
import numpy as np
import pandas as pd
import settings as conf
# genes_associations_dir = os.path.join(conf.PREPROCESSED_BASED_DIR, 'gene_associations')
# smultixcan_gene_association_dirs = os.path.join(genes_associations_dir, 'mashr')
output_dir = os.path.join(conf.DELIVERABLES_DIR, 'roc_validation', 'classifier_tables')
os.makedirs(output_dir, exist_ok=True)
RCP_CUTOFF = 0.10
```
# Load gene mappings
```
with open(os.path.join(conf.GENES_METADATA_DIR, 'genes_mapping_simplified-0.pkl'), 'rb') as f:
genes_mapping_0 = pickle.load(f)
with open(os.path.join(conf.GENES_METADATA_DIR, 'genes_mapping_simplified-1.pkl'), 'rb') as f:
genes_mapping_1 = pickle.load(f)
```
# Load S-MultiXcan results
```
smultixcan_genes_associations_filename = os.path.join(conf.GENE_ASSOC_DIR, 'smultixcan-mashr-zscores.pkl.xz')
display(smultixcan_genes_associations_filename)
smultixcan_genes_associations = pd.read_pickle(smultixcan_genes_associations_filename)
smultixcan_genes_associations.shape
smultixcan_genes_associations.head(5)
```
# Load fastENLOC results
```
fastenloc_genes_associations_filename = os.path.join(conf.GENE_ASSOC_DIR, 'fastenloc-torus-rcp.pkl.xz')
display(fastenloc_genes_associations_filename)
fastenloc_genes_associations = pd.read_pickle(fastenloc_genes_associations_filename)
fastenloc_genes_associations.shape
fastenloc_genes_associations.head(5)
```
# Genes in common between S-MultiXcan and fastENLOC
```
common_genes = fastenloc_genes_associations.index.intersection(smultixcan_genes_associations.index)
display(common_genes)
```
# Load OMIM silver standard
```
omim_silver_standard = pd.read_csv(os.path.join(conf.DATA_DIR, 'omim_silver_standard.tsv'), sep='\t')
omim_silver_standard = omim_silver_standard.dropna(subset=['ensembl_gene_id', 'trait', 'pheno_mim'])
display(omim_silver_standard.shape)
display(omim_silver_standard.head())
```
# Read gwas2gene results
These results were generated with scripts in `scripts/extras/gwas2gene`.
```
from glob import glob
import rpy2.robjects as robjects
from rpy2.robjects import pandas2ri
pandas2ri.activate()
readRDS = robjects.r['readRDS']
f_files = glob(os.path.join(conf.OMIM_SILVER_STANDARD_GWAS_TO_GENE_DIR, '*.rds'))
display(len(f_files))
if len(f_files) != len(omim_silver_standard['trait'].unique()):
print(f'WARNING: some files are not there. {len(omim_silver_standard["trait"].unique())} expected, {len(f_files)} found.')
gwas2genes_results = {}
for f in f_files:
f_base = os.path.basename(f)
f_code, _ = os.path.splitext(f_base)
#print(f_base)
rds_contents = readRDS(f)
if len(rds_contents[1]) > 0:
f_gene_list = list(rds_contents[1][0].iter_labels())
else:
print(f'{f_code}: empty')
f_gene_list = []
gwas2genes_results[f_code] = common_genes.intersection(set(f_gene_list))
gwas2gene_all_genes = []
for k in gwas2genes_results.keys():
gwas2gene_all_genes.extend(gwas2genes_results[k])
display(len(gwas2gene_all_genes))
gwas2gene_all_genes = set(gwas2gene_all_genes)
display(len(gwas2gene_all_genes))
```
# Create list of UKB-OMIM traits
```
omim_silver_standard.head()
```
# Create PrediXcan classifier table
```
_tmp = omim_silver_standard[['trait', 'ensembl_gene_id']]
ukb_traits_common = _tmp['trait'].unique()
omim_true_classes = _tmp[['trait', 'ensembl_gene_id']].drop_duplicates()
omim_true_classes = omim_true_classes.assign(omim_value=1)
omim_true_classes = omim_true_classes.set_index(['trait', 'ensembl_gene_id'])
len(ukb_traits_common)
omim_true_classes.shape
omim_true_classes.head()
len(ukb_traits_common)
from entity import Trait
index_tuples = []
for t in ukb_traits_common:
t_code = Trait(full_code=t).code
if t_code not in gwas2genes_results:
continue
for g in gwas2genes_results[t_code]:
index_tuples.append((t, g))
len(index_tuples)
index_tuples[:5]
classifier_index = pd.MultiIndex.from_tuples(
index_tuples,
names=['ukb_efo', 'gene']
)
len(gwas2gene_all_genes)
classifier_index.shape
predixcan_classifier_df = pd.DataFrame(index=classifier_index, columns=['score', 'predicted_class', 'true_class'])
predixcan_classifier_df = predixcan_classifier_df.sort_index()
predixcan_classifier_df.shape
predixcan_classifier_df['true_class'] = 0
predixcan_classifier_df.head()
true_classes = omim_true_classes.squeeze()
display(true_classes.shape)
display(true_classes.head())
predixcan_classifier_df.loc[predixcan_classifier_df.index.intersection(true_classes.index), 'true_class'] = 1
assert predixcan_classifier_df['true_class'].isna().sum() == 0
# some testing
predixcan_classifier_df.loc[('M41-Diagnoses_main_ICD10_M41_Scoliosis',)].head()
true_classes.loc[('M41-Diagnoses_main_ICD10_M41_Scoliosis', 'ENSG00000112234')]
'ENSG00000090263' not in true_classes.loc['M41-Diagnoses_main_ICD10_M41_Scoliosis'].index
assert predixcan_classifier_df.loc[('M41-Diagnoses_main_ICD10_M41_Scoliosis', 'ENSG00000112234'), 'true_class'] == 1.0
assert predixcan_classifier_df.loc[('M41-Diagnoses_main_ICD10_M41_Scoliosis', 'ENSG00000090263'), 'true_class'] == 0.0
len(gwas2gene_all_genes)
# score
df_score = pd.Series(index=classifier_index, dtype=float)
for trait in ukb_traits_common:
trait_code = Trait(full_code=trait).code
if trait_code not in gwas2genes_results:
print(trait_code)
continue
trait_genes = gwas2genes_results[trait_code]
if len(trait_genes) == 0:
print(f'Empty: {trait}')
continue
smultixcan_zscores = smultixcan_genes_associations.loc[trait_genes, trait]
fastenloc_rcps = fastenloc_genes_associations.loc[trait_genes, trait].fillna(0.0)
smultixcan_zscores[(fastenloc_rcps < RCP_CUTOFF) & (~np.isnan(smultixcan_zscores))] = 0
scores = smultixcan_zscores
df_score.loc[trait] = scores.values
# for S-MultiXcan dropna
df_score = df_score.dropna().sort_index()
assert df_score.isna().sum().sum() == 0
df_score.head()
# some testing
_gene, _trait = ('ENSG00000090263', 'M41-Diagnoses_main_ICD10_M41_Scoliosis')
display(smultixcan_genes_associations.loc[_gene, _trait])
display(fastenloc_genes_associations.loc[_gene, _trait])
assert 0.0 == df_score.loc[_trait, _gene]
_gene, _trait = ('ENSG00000070061', 'O14-Diagnoses_main_ICD10_O14_Gestational_pregnancyinduced_hypertension_with_significant_proteinuria')
display(smultixcan_genes_associations.loc[_gene, _trait])
display(fastenloc_genes_associations.loc[_gene, _trait])
assert 0.0 == df_score.loc[_trait, _gene]
_gene, _trait = ('ENSG00000004534', '1200-Sleeplessness_insomnia')
display(smultixcan_genes_associations.loc[_gene, _trait])
display(fastenloc_genes_associations.loc[_gene, _trait])
assert smultixcan_genes_associations.loc[_gene, _trait] == df_score.loc[_trait, _gene]
df_score.shape
df_score.head()
df_score.describe()
predixcan_classifier_df = predixcan_classifier_df.assign(score=df_score)
# assert not predixcan_classifier_df['score'].isna().any()
from scipy import stats
_n_genes = len(gwas2gene_all_genes)
display(_n_genes)
_n_ukb_traits = len(omim_silver_standard['trait'].unique())
display(_n_ukb_traits)
display(_n_genes * _n_ukb_traits)
PVALUE_THRESHOLD = (0.05 / (_n_genes * _n_ukb_traits))
display(PVALUE_THRESHOLD)
ZSCORE_THRESHOLD = np.abs(stats.norm.ppf(PVALUE_THRESHOLD / 2))
display(ZSCORE_THRESHOLD)
def _assign_predicted_class(x):
# trait, gene = x.name
# smultixcan_zscore = smultixcan_genes_associations.loc[gene, trait]
if x > ZSCORE_THRESHOLD:
return 1
else:
return 0
predixcan_classifier_df = predixcan_classifier_df.assign(predicted_class=predixcan_classifier_df['score'].apply(_assign_predicted_class))
predixcan_classifier_df.shape
predixcan_classifier_df.head()
predixcan_classifier_df.loc['M41-Diagnoses_main_ICD10_M41_Scoliosis'].sort_values('true_class', ascending=False).head()
```
## Select genes per trait
```
#selected_predixcan_classifier_df = predixcan_classifier_df.loc[predixcan_classifier_df.index.intersection(trait_genes_to_keep)]
selected_predixcan_classifier_df = predixcan_classifier_df
# some testing
selected_predixcan_classifier_df.shape
selected_predixcan_classifier_df.head()
selected_predixcan_classifier_df.sort_values('predicted_class', ascending=False).head()
_tmp = selected_predixcan_classifier_df.sort_values(['true_class', 'ukb_efo'], ascending=False)
display(_tmp.shape)
display(_tmp[_tmp['true_class'] > 0].shape)
display(_tmp[_tmp['true_class'] > 0].head())
```
### Test classes
```
selected_predixcan_classifier_df.index.get_level_values('ukb_efo').unique().shape
selected_predixcan_classifier_df.index.get_level_values('gene').unique().shape
_pheno = 'N20-Diagnoses_main_ICD10_N20_Calculus_of_kidney_and_ureter'
_clinvar_asthma_genes = omim_silver_standard[omim_silver_standard['trait'] == _pheno]['ensembl_gene_id'].unique()
display(_clinvar_asthma_genes)
display(_clinvar_asthma_genes.shape)
_tmp = selected_predixcan_classifier_df.loc[_pheno]
_tmp.loc[_tmp.index.intersection(_clinvar_asthma_genes)]
_predixcan_asthma_genes = selected_predixcan_classifier_df.loc[_pheno]
_predixcan_asthma_genes.head()
selected_predixcan_classifier_df.shape
selected_predixcan_classifier_df['predicted_class'].value_counts()
selected_predixcan_classifier_df['true_class'].value_counts()
selected_predixcan_classifier_df.sort_values(['true_class'], ascending=[False])
```
# Save classifier table
```
# remove nans
selected_predixcan_classifier_df = selected_predixcan_classifier_df.dropna()
assert selected_predixcan_classifier_df.index.is_unique
selected_predixcan_classifier_df.head()
selected_predixcan_classifier_df.shape
# this variant is not used, do not save
# selected_predixcan_classifier_df.to_csv(
# os.path.join(output_dir, 'combined-classifier_data.tsv.gz'),
# sep='\t', index=False
# )
```
| github_jupyter |
```
# ~4/1 5465, ~3/1 11074 조선일보는 기사 연속해서 볼 때 로그인해야해서 시험해보지 않음
## 입력값(이것만 입력할 것) ##
num = [0, 5466] # 수행할 구간, 조선일보은 range(0, 5465+1) 설정해야함, 내가 처음에 range(0,50)까지했으면 다음에는 range(50,이후숫자)
loot = 'C:/Users/###/###/###/' # 저장할 위치, 파일이 산만해지니 프로젝트가아닌폴더에서 관리할 것 # loot = './/' 현재위치에 저장하는 변수
# 파일이름
name = 'text' + '_' + str(num[0]) + '_' + str(num[1]) # 파일이름 Ex. text_0_50
import sqlite3
from selenium import webdriver
from selenium.webdriver.remote.webelement import WebElement
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.common.by import By
from selenium.webdriver.support import expected_conditions as EC
from bs4 import BeautifulSoup
import requests
import pandas as pd
import csv
import time
import re
df = pd.read_excel('.//NewsResult_20200901-20210430 (7).xlsx', names=['identical', 'date', 'press', 'name', 'title', 'c1', 'c2', 'c3', 'a1', 'a2', 'a3', 'person', 'place', 'institute', 'keyword', 'topkeyword', 'body', 'url', 'tf'])
# 전처리
df = df[df.tf != '예외']
df = df[df.tf != '중복']
df = df[df.tf != '중복, 예외']
# df = df[~df.title.str.contains('경향포토')]
# df = df[~df.title.str.contains('인터랙티브')]
# df = df[~df.place.str.contains('korea', na=False)]
# df = df[~df.place.str.contains('la', na=False)]
# df = df[~df.place.str.contains('LA', na=False)]
df = df.reset_index()
df = df.drop(columns=['index'], axis=1)
len(df)
# df.to_excel('.//pre_press_7.xls', header=None, index=False)
# ~4/1 5465, ~3/1 11074
df = df.iloc[0:5465] # 4/30 ~ 4/1 1달간 총 5465개 *50개(3분)
# 셀레니움 창 오픈, 탭에 애드블락은 설치후 엑스해도 됨
options = webdriver.ChromeOptions()
# options.add_extension(r'..//extension_3_11_1_0.crx')
options.add_argument('headless')
browser = webdriver.Chrome('..//chromedriver.exe', options=options)
from selenium.common.exceptions import NoSuchElementException
def check_exists_by_xpath(xpath):
try:
browser.find_element_by_xpath(xpath)
except NoSuchElementException:
return False
return True
# section.article-body > p.article-body__content
texts = []
for i in range(num[0], num[1]):
try:
url = df.url[i]
browser.get(url)
WebDriverWait(browser, 3).until(EC.presence_of_element_located((By.XPATH, "//p[@class='article-body__content']")))
if check_exists_by_xpath("//p[@class='article-body__content']") == True:
html = browser.page_source
soup = BeautifulSoup(html, 'html.parser')
contents = soup.select('p.content_text')
text = ''
for content in contents:
text = text + content.text.strip()
texts.append(text)
else:
texts.append('None')
continue
except:
texts.append('None')
continue
len(texts)
text = pd.DataFrame(data=texts) # texts를 text로 pandas로 저장
spot = loot + name + '.xls'
text.to_excel(spot, header=None,index=False)
# 검증, 통합 설명과 수행방법(주석제거)
# 분할된 text를 순서대로 합치는 법[2개(파일이름1, 파일이름2)를 불러울 경우]
# 파일이름 = pd.read_excel('C:/Develops/newspapers/press0/파일이름.xls', header=None) # 엑셀파일불러오기
# 파일이름 = 파일이름.values.tolist() # 리스트로 환원
# 파일이름2 = pd.read_excel('C:/Develops/newspapers/press0/파일이름2.xls', header=None)
# 파일이름2 = 파일이름2.values.tolist()
# co_text = 파일이름 + 파일이름2 # 리스트 순서대로 붙이기
# 제목과 내용 일치 여부 판단
# len(co_text)
# i = 400 # co_text에 있는 숫자 구간에서 임의의 숫자 넣어서 제목과 내용이 일치하는지 대략적으로 검수
# df.title[i] # 번호에 해당하는 제목
# co_text[i] # 번호에 해당하는 내용, co가 아니라 바로 결과를 확인할 때는 texts[i]
# df에 text 넣어서 저장하기 # 모든 기사내용을 스크래핑해서 co_text로 합쳤으면 excel로 저장
# df['total_text'] = co_text
# final_spot = loot + 'press_0.xls'
# df.to_excel(final_spot, header=None,index=False)
# 끝
```
| github_jupyter |
# Introduction
Here we process the data from the electron microscopy 3D dataset, the first lines automatically download it if you have Kaggle setup otherwise you can download the dataset [here](https://www.kaggle.com/kmader/electron-microscopy-3d-segmentation/data)
```
import os
if not os.path.exists('input'):
!kaggle datasets download -d kmader/electron-microscopy-3d-segmentation -wp emdata
!mkdir input
!mv emdata/* input
%matplotlib inline
from skimage.io import imread # for reading images
import matplotlib.pyplot as plt # for showing plots
from skimage.measure import label # for labeling regions
from skimage.measure import regionprops # for shape analysis
import numpy as np # for matrix operations and array support
from skimage.color import label2rgb # for making overlay plots
import matplotlib.patches as mpatches # for showing rectangles and annotations
```
# Connected Component Labeling
scikit-image has basic support for [connected component labeling](http://scikit-image.org/docs/dev/api/skimage.measure.html#skimage.measure.label) and we can do some small demos with the label function and small test images, before moving onto bigger datasets.
## Neighborhood
In the course we use the term neighborhood and here we use the term ```connectivity``` for the same idea.
- For a 2D image a connectivity = 1 is just the 4-neighborhood (or pixels that share an edge, connectivity = 2 is then 8-neighborhood (or pixels that share a vertex).
```
# simple test image diagonal
test_img=np.eye(4)
print('Input Image')
print(test_img)
test_label_4=label(test_img,connectivity=1)
print('Labels with 4-neighborhood')
print(test_label_4)
test_label_8=label(test_img,connectivity=2)
print('Labels with 8-neighborhood')
print(test_label_8)
```
## 3D Neighborhood
For a 3D image a connectivity = 1 is just the 6-neighborhood (or voxels that share an face, connectivity = 2 is then voxels that share an edge and 3 is voxels that share a vertex
```
test_img=np.array([1 if x in [0,13,26] else 0 for x in range(27)]).reshape((3,3,3))
print('Input Image')
print(test_img)
test_label_1=label(test_img,connectivity=1)
print('Labels with Face-sharing')
print(test_label_1)
test_label_2=label(test_img,connectivity=2)
print('Labels with Edge-Sharing')
print(test_label_2)
test_label_3=label(test_img,connectivity=3)
print('Labels with Vertex-Sharing')
print(test_label_3)
import os, numpy as np
def imread_or_invent(in_path):
np.random.seed(2018)
if os.path.exists(in_path):
return imread(in_path)
else:
print('Getting creative...')
fake_shape = (10, 50, 75)
if 'groundtruth' in in_path:
return (np.random.uniform(0, 1, size = fake_shape)>0.99).astype(int)
else:
return np.random.uniform(0, 1, size = fake_shape)
em_image_vol = imread_or_invent('input/training.tif')
em_thresh_vol = imread_or_invent('input/training_groundtruth.tif')
print("Data Loaded, Dimensions", em_image_vol.shape,'->',em_thresh_vol.shape)
```
# 2D Analysis
Here we work with a single 2D slice to get started and take it randomly from the middle
```
em_idx = np.random.permutation(range(em_image_vol.shape[0]))[0]
em_slice = em_image_vol[em_idx]
em_thresh = em_thresh_vol[em_idx]
print("Slice Loaded, Dimensions", em_slice.shape)
# show the slice and threshold
fig, (ax1, ax2, ax3) = plt.subplots(1,3, figsize = (9, 4))
ax1.imshow(em_slice, cmap = 'gray')
ax1.axis('off')
ax1.set_title('Image')
ax2.imshow(em_thresh, cmap = 'gray')
ax2.axis('off')
ax2.set_title('Segmentation')
# here we mark the threshold on the original image
ax3.imshow(label2rgb(em_thresh,em_slice, bg_label=0))
ax3.axis('off')
ax3.set_title('Overlayed');
# make connected component labels
em_label = label(em_thresh)
print(em_label.max(), 'number of labels')
# show the segmentation, labels and overlay
fig, (ax1, ax2, ax3) = plt.subplots(1,3, figsize = (9, 4))
ax1.imshow(em_thresh, cmap = 'gray')
ax1.axis('off')
ax1.set_title('Segmentation')
ax2.imshow(em_label, cmap = plt.cm.gist_earth)
ax2.axis('off')
ax2.set_title('Labeling')
# here we mark the threshold on the original image
ax3.imshow(label2rgb(em_label,em_slice, bg_label=0))
ax3.axis('off')
ax3.set_title('Overlayed');
```
# Shape Analysis
For shape analysis we use the regionprops function which calculates the area, perimeter, and other features for a shape. The analysis creates a list of these with one for each label in the original image.
```
shape_analysis_list = regionprops(em_label)
first_region = shape_analysis_list[0]
print('List of region properties for',len(shape_analysis_list), 'regions')
print('Features Calculated:',', '.join([f for f in dir(first_region) if not f.startswith('_')]))
fig, ax = plt.subplots(figsize=(10, 6))
ax.imshow(label2rgb(em_label,em_slice, bg_label=0))
for region in shape_analysis_list:
# draw rectangle using the bounding box
minr, minc, maxr, maxc = region.bbox
rect = mpatches.Rectangle((minc, minr), maxc - minc, maxr - minr,
fill=False, edgecolor='red', linewidth=2)
ax.add_patch(rect)
ax.set_axis_off()
plt.tight_layout()
```
## Anisotropy
We can calculate anisotropy as we did in the course by using the largest and shortest lengths, called here as ```major_axis_length``` and ```minor_axis_length``` respectively
- Try using different formulas for anisotropy to see how it changes what is shown
$$ Aiso1 = \frac{\text{Longest Side}}{\text{Shortest Side}} - 1 $$
$$ Aiso2 = \frac{\text{Longest Side}-\text{Shortest Side}}{\text{Longest Side}} $$
$$ Aiso3 = \frac{\text{Longest Side}}{\text{Average Side Length}} - 1 $$
$$ Aiso4 = \frac{\text{Longest Side}-\text{Shortest Side}}{\text{Average Side Length}} $$
```
fig, ax = plt.subplots(figsize=(10, 6))
ax.imshow(label2rgb(em_label,em_slice, bg_label=0))
for region in shape_analysis_list:
x1=region.major_axis_length
x2=region.minor_axis_length
anisotropy = (x1-x2)/np.clip(x1+x2, 0.1, 9999)
# for anisotropic shapes use red for the others use blue
print('Label:',region.label,'Anisotropy %2.2f' % anisotropy)
if anisotropy>0.1:
minr, minc, maxr, maxc = region.bbox
rect = mpatches.Rectangle((minc, minr), maxc - minc, maxr - minr,
fill=False, edgecolor='red', linewidth=2)
ax.add_patch(rect)
else:
minr, minc, maxr, maxc = region.bbox
rect = mpatches.Rectangle((minc, minr), maxc - minc, maxr - minr,
fill=False, edgecolor='green', linewidth=2)
ax.add_patch(rect)
ax.set_axis_off()
plt.tight_layout()
```
# Tasks
- Perform the analysis in 3D
- Find the largest and smallest structures
- Find the structures with the highest and lowest 3D anisotropy
| github_jupyter |
Licensed to the Apache Software Foundation (ASF) under one or more contributor license agreements; and to You under the Apache License, Version 2.0.
# SINGA Core Classes
<img src="http://singa.apache.org/en/_static/images/singav1-sw.png" width="500px"/>
# Device
A device instance represents a hardware device with multiple execution units, e.g.,
* A GPU which has multile cuda streams
* A CPU which has multiple threads
All data structures (variables) are allocated on a device instance. Consequently, all operations are executed on the resident device.
## Create a device instance
```
from singa import device
default_dev = device.get_default_device()
gpu = device.create_cuda_gpu() # the first gpu device
gpu
```
** NOTE: currently we can only call the creating function once due to the cnmem restriction.**
```
gpu = device.create_cuda_gpu_on(1) # use the gpu device with the specified GPU ID
gpu_list1 = device.create_cuda_gpus(2) # the first two gpu devices
gpu_list2 = device.create_cuda_gpus([0,2]) # create the gpu instances on the given GPU IDs
opencl_gpu = device.create_opencl_device() # valid if SINGA is compiled with USE_OPENCL=ON
device.get_num_gpus()
device.get_gpu_ids()
```
# Tensor
A tensor instance represents a multi-dimensional array allocated on a device instance.
It provides linear algbra operations, like +, -, *, /, dot, pow ,etc
NOTE: class memeber functions are inplace; global functions are out-of-place.
### Create tensor instances
```
from singa import tensor
import numpy as np
a = tensor.Tensor((2, 3))
a.shape
a.device
gb = tensor.Tensor((2, 3), gpu)
gb.device
```
### Initialize tensor values
```
a.set_value(1.2)
gb.gaussian(0, 0.1)
```
### To and from numpy
```
tensor.to_numpy(a)
tensor.to_numpy(gb)
c = tensor.from_numpy(np.array([1,2], dtype=np.float32))
c.shape
c.copy_from_numpy(np.array([3,4], dtype=np.float32))
tensor.to_numpy(c)
```
### Move tensor between devices
```
gc = c.clone()
gc.to_device(gpu)
gc.device
b = gb.clone()
b.to_host() # the same as b.to_device(default_dev)
b.device
```
### Operations
**NOTE: tensors should be initialized if the operation would read the tensor values**
#### Summary
```
gb.l1()
a.l2()
e = tensor.Tensor((2, 3))
e.is_empty()
gb.size()
gb.memsize()
# note we can only support matrix multiplication for tranposed tensors;
# other operations on transposed tensor would result in errors
c.is_transpose()
et=e.T()
et.is_transpose()
et.shape
et.ndim()
```
#### Member functions (in-place)
These functions would change the content of the tensor
```
a += b
tensor.to_numpy(a)
a -= b
tensor.to_numpy(a)
a *= 2
tensor.to_numpy(a)
a /= 3
tensor.to_numpy(a)
d = tensor.Tensor((3,))
d.uniform(-1,1)
tensor.to_numpy(d)
a.add_row(d)
tensor.to_numpy(a)
```
#### Global functions (out of place)
These functions would not change the memory of the tensor, instead they return a new tensor
**Unary functions**
```
h = tensor.sign(d)
tensor.to_numpy(h)
tensor.to_numpy(d)
h = tensor.abs(d)
tensor.to_numpy(h)
h = tensor.relu(d)
tensor.to_numpy(h)
g = tensor.sum(a, 0)
g.shape
g = tensor.sum(a, 1)
g.shape
tensor.bernoulli(0.5, g)
tensor.to_numpy(g)
g.gaussian(0, 0.2)
tensor.gaussian(0, 0.2, g)
tensor.to_numpy(g)
```
#### Binary functions
```
f = a + b
tensor.to_numpy(f)
g = a < b
tensor.to_numpy(g)
tensor.add_column(2, c, 1, f) # f = 2 *c + 1* f
tensor.to_numpy(f)
```
#### BLAS
BLAS function may change the memory of input tensor
```
tensor.axpy(2, a, f) # f = 2a + f
tensor.to_numpy(b)
f = tensor.mult(a, b.T())
tensor.to_numpy(f)
tensor.mult(a, b.T(), f, 2, 1) # f = 2a*b.T() + 1f
tensor.to_numpy(f)
```
## Next: [SINGA model classes](./model.ipynb)
| github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork/labs/Module%204/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Hierarchical Clustering
Estimated time needed: **25** minutes
## Objectives
After completing this lab you will be able to:
* Use scikit-learn to do Hierarchical clustering
* Create dendograms to visualize the clustering
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="https://#hierarchical_agglomerative">Hierarchical Clustering - Agglomerative</a></li>
<ol>
<li><a href="https://#generating_data">Generating Random Data</a></li>
<li><a href="https://#agglomerative_clustering">Agglomerative Clustering</a></li>
<li><a href="https://#dendrogram">Dendrogram Associated for the Agglomerative Hierarchical Clustering</a></li>
</ol>
<li><a href="https://#clustering_vehicle_dataset">Clustering on the Vehicle Dataset</a></li>
<ol>
<li><a href="https://#data_cleaning">Data Cleaning</a></li>
<li><a href="https://#clustering_using_scipy">Clustering Using Scipy</a></li>
<li><a href="https://#clustering_using_skl">Clustering using scikit-learn</a></li>
</ol>
</ol>
</div>
<br>
<hr>
<h1 id="hierarchical_agglomerative">Hierarchical Clustering - Agglomerative</h1>
We will be looking at a clustering technique, which is <b>Agglomerative Hierarchical Clustering</b>. Remember that agglomerative is the bottom up approach. <br> <br>
In this lab, we will be looking at Agglomerative clustering, which is more popular than Divisive clustering. <br> <br>
We will also be using Complete Linkage as the Linkage Criteria. <br> <b> <i> NOTE: You can also try using Average Linkage wherever Complete Linkage would be used to see the difference! </i> </b>
```
import numpy as np
import pandas as pd
from scipy import ndimage
from scipy.cluster import hierarchy
from scipy.spatial import distance_matrix
from matplotlib import pyplot as plt
from sklearn import manifold, datasets
from sklearn.cluster import AgglomerativeClustering
from sklearn.datasets._samples_generator import make_blobs
%matplotlib inline
```
<hr>
<h3 id="generating_data">Generating Random Data</h3>
We will be generating a set of data using the <b>make_blobs</b> class. <br> <br>
Input these parameters into make_blobs:
<ul>
<li> <b>n_samples</b>: The total number of points equally divided among clusters. </li>
<ul> <li> Choose a number from 10-1500 </li> </ul>
<li> <b>centers</b>: The number of centers to generate, or the fixed center locations. </li>
<ul> <li> Choose arrays of x,y coordinates for generating the centers. Have 1-10 centers (ex. centers=[[1,1], [2,5]]) </li> </ul>
<li> <b>cluster_std</b>: The standard deviation of the clusters. The larger the number, the further apart the clusters</li>
<ul> <li> Choose a number between 0.5-1.5 </li> </ul>
</ul> <br>
Save the result to <b>X1</b> and <b>y1</b>.
```
X1, y1 = make_blobs(n_samples=50, centers=[[4,4], [-2, -1], [1, 1], [10,4]], cluster_std=0.9)
```
Plot the scatter plot of the randomly generated data.
```
plt.scatter(X1[:, 0], X1[:, 1], marker='o');
```
<hr>
<h3 id="agglomerative_clustering">Agglomerative Clustering</h3>
We will start by clustering the random data points we just created.
The <b> Agglomerative Clustering </b> class will require two inputs:
<ul>
<li> <b>n_clusters</b>: The number of clusters to form as well as the number of centroids to generate. </li>
<ul> <li> Value will be: 4 </li> </ul>
<li> <b>linkage</b>: Which linkage criterion to use. The linkage criterion determines which distance to use between sets of observation. The algorithm will merge the pairs of cluster that minimize this criterion. </li>
<ul>
<li> Value will be: 'complete' </li>
<li> <b>Note</b>: It is recommended you try everything with 'average' as well </li>
</ul>
</ul> <br>
Save the result to a variable called <b> agglom </b>.
```
agglom = AgglomerativeClustering(n_clusters = 4, linkage = 'average')
```
Fit the model with <b> X2 </b> and <b> y2 </b> from the generated data above.
```
agglom.fit(X1,y1)
```
Run the following code to show the clustering! <br>
Remember to read the code and comments to gain more understanding on how the plotting works.
```
# Create a figure of size 6 inches by 4 inches.
plt.figure(figsize=(6,4))
# These two lines of code are used to scale the data points down,
# Or else the data points will be scattered very far apart.
# Create a minimum and maximum range of X1.
x_min, x_max = np.min(X1, axis=0), np.max(X1, axis=0)
# Get the average distance for X1.
X1 = (X1 - x_min) / (x_max - x_min)
# This loop displays all of the datapoints.
for i in range(X1.shape[0]):
# Replace the data points with their respective cluster value
# (ex. 0) and is color coded with a colormap (plt.cm.spectral)
plt.text(X1[i, 0], X1[i, 1], str(y1[i]),
color=plt.cm.nipy_spectral(agglom.labels_[i] / 10.),
fontdict={'weight': 'bold', 'size': 9})
# Remove the x ticks, y ticks, x and y axis
plt.xticks([])
plt.yticks([])
#plt.axis('off')
# Display the plot of the original data before clustering
plt.scatter(X1[:, 0], X1[:, 1], marker='.')
# Display the plot
plt.show()
```
<h3 id="dendrogram">Dendrogram Associated for the Agglomerative Hierarchical Clustering</h3>
Remember that a <b>distance matrix</b> contains the <b> distance from each point to every other point of a dataset </b>.
Use the function <b> distance_matrix, </b> which requires <b>two inputs</b>. Use the Feature Matrix, <b> X1 </b> as both inputs and save the distance matrix to a variable called <b> dist_matrix </b> <br> <br>
Remember that the distance values are symmetric, with a diagonal of 0's. This is one way of making sure your matrix is correct. <br> (print out dist_matrix to make sure it's correct)
```
dist_matrix = distance_matrix(X1,X1)
print(dist_matrix)
```
Using the <b> linkage </b> class from hierarchy, pass in the parameters:
<ul>
<li> The distance matrix </li>
<li> 'complete' for complete linkage </li>
</ul> <br>
Save the result to a variable called <b> Z </b>.
```
Z = hierarchy.linkage(dist_matrix, 'complete')
```
A Hierarchical clustering is typically visualized as a dendrogram as shown in the following cell. Each merge is represented by a horizontal line. The y-coordinate of the horizontal line is the similarity of the two clusters that were merged, where cities are viewed as singleton clusters.
By moving up from the bottom layer to the top node, a dendrogram allows us to reconstruct the history of merges that resulted in the depicted clustering.
Next, we will save the dendrogram to a variable called <b>dendro</b>. In doing this, the dendrogram will also be displayed.
Using the <b> dendrogram </b> class from hierarchy, pass in the parameter:
<ul> <li> Z </li> </ul>
```
dendro = hierarchy.dendrogram(Z)
```
## Practice
We used **complete** linkage for our case, change it to **average** linkage to see how the dendogram changes.
```
Z = hierarchy.linkage(dist_matrix, 'average')
dendro = hierarchy.dendrogram(Z)
```
<details><summary>Click here for the solution</summary>
```python
Z = hierarchy.linkage(dist_matrix, 'average')
dendro = hierarchy.dendrogram(Z)
```
</details>
<hr>
<h1 id="clustering_vehicle_dataset">Clustering on Vehicle dataset</h1>
Imagine that an automobile manufacturer has developed prototypes for a new vehicle. Before introducing the new model into its range, the manufacturer wants to determine which existing vehicles on the market are most like the prototypes--that is, how vehicles can be grouped, which group is the most similar with the model, and therefore which models they will be competing against.
Our objective here, is to use clustering methods, to find the most distinctive clusters of vehicles. It will summarize the existing vehicles and help manufacturers to make decision about the supply of new models.
### Download data
To download the data, we will use **`!wget`** to download it from IBM Object Storage.\
**Did you know?** When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
```
!wget -O cars_clus.csv https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork/labs/Module%204/data/cars_clus.csv
```
## Read data
Let's read dataset to see what features the manufacturer has collected about the existing models.
```
filename = 'cars_clus.csv'
#Read csv
pdf = pd.read_csv(filename)
print ("Shape of dataset: ", pdf.shape)
pdf.head(5)
```
The feature sets include price in thousands (price), engine size (engine_s), horsepower (horsepow), wheelbase (wheelbas), width (width), length (length), curb weight (curb_wgt), fuel capacity (fuel_cap) and fuel efficiency (mpg).
<h2 id="data_cleaning">Data Cleaning</h2>
Let's clean the dataset by dropping the rows that have null value:
```
print ("Shape of dataset before cleaning: ", pdf.size)
pdf[[ 'sales', 'resale', 'type', 'price', 'engine_s',
'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap',
'mpg', 'lnsales']] = pdf[['sales', 'resale', 'type', 'price', 'engine_s',
'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap',
'mpg', 'lnsales']].apply(pd.to_numeric, errors='coerce')
pdf = pdf.dropna()
pdf = pdf.reset_index(drop=True)
print ("Shape of dataset after cleaning: ", pdf.size)
pdf.head(5)
```
### Feature selection
Let's select our feature set:
```
featureset = pdf[['engine_s', 'horsepow', 'wheelbas', 'width', 'length', 'curb_wgt', 'fuel_cap', 'mpg']]
```
### Normalization
Now we can normalize the feature set. **MinMaxScaler** transforms features by scaling each feature to a given range. It is by default (0, 1). That is, this estimator scales and translates each feature individually such that it is between zero and one.
```
from sklearn.preprocessing import MinMaxScaler
x = featureset.values #returns a numpy array
min_max_scaler = MinMaxScaler()
feature_mtx = min_max_scaler.fit_transform(x)
feature_mtx [0:5]
```
<h2 id="clustering_using_scipy">Clustering using Scipy</h2>
In this part we use Scipy package to cluster the dataset.
First, we calculate the distance matrix.
```
import scipy
leng = feature_mtx.shape[0]
D = scipy.zeros([leng,leng])
for i in range(leng):
for j in range(leng):
D[i,j] = scipy.spatial.distance.euclidean(feature_mtx[i], feature_mtx[j])
D
```
In agglomerative clustering, at each iteration, the algorithm must update the distance matrix to reflect the distance of the newly formed cluster with the remaining clusters in the forest.
The following methods are supported in Scipy for calculating the distance between the newly formed cluster and each:
\- single
\- complete
\- average
\- weighted
\- centroid
We use **complete** for our case, but feel free to change it to see how the results change.
```
import pylab
import scipy.cluster.hierarchy
Z = hierarchy.linkage(D, 'complete')
```
Essentially, Hierarchical clustering does not require a pre-specified number of clusters. However, in some applications we want a partition of disjoint clusters just as in flat clustering.
So you can use a cutting line:
```
from scipy.cluster.hierarchy import fcluster
max_d = 3
clusters = fcluster(Z, max_d, criterion='distance')
clusters
```
Also, you can determine the number of clusters directly:
```
from scipy.cluster.hierarchy import fcluster
k = 5
clusters = fcluster(Z, k, criterion='maxclust')
clusters
```
Now, plot the dendrogram:
```
fig = pylab.figure(figsize=(18,50))
def llf(id):
return '[%s %s %s]' % (pdf['manufact'][id], pdf['model'][id], int(float(pdf['type'][id])) )
dendro = hierarchy.dendrogram(Z, leaf_label_func=llf, leaf_rotation=0, leaf_font_size =12, orientation = 'right')
```
<h2 id="clustering_using_skl">Clustering using scikit-learn</h2>
Let's redo it again, but this time using the scikit-learn package:
```
from sklearn.metrics.pairwise import euclidean_distances
dist_matrix = euclidean_distances(feature_mtx,feature_mtx)
print(dist_matrix)
Z_using_dist_matrix = hierarchy.linkage(dist_matrix, 'complete')
fig = pylab.figure(figsize=(18,50))
def llf(id):
return '[%s %s %s]' % (pdf['manufact'][id], pdf['model'][id], int(float(pdf['type'][id])) )
dendro = hierarchy.dendrogram(Z_using_dist_matrix, leaf_label_func=llf, leaf_rotation=0, leaf_font_size =12, orientation = 'right')
```
Now, we can use the 'AgglomerativeClustering' function from scikit-learn library to cluster the dataset. The AgglomerativeClustering performs a hierarchical clustering using a bottom up approach. The linkage criteria determines the metric used for the merge strategy:
* Ward minimizes the sum of squared differences within all clusters. It is a variance-minimizing approach and in this sense is similar to the k-means objective function but tackled with an agglomerative hierarchical approach.
* Maximum or complete linkage minimizes the maximum distance between observations of pairs of clusters.
* Average linkage minimizes the average of the distances between all observations of pairs of clusters.
```
agglom = AgglomerativeClustering(n_clusters = 6, linkage = 'complete')
agglom.fit(dist_matrix)
agglom.labels_
```
We can add a new field to our dataframe to show the cluster of each row:
```
pdf['cluster_'] = agglom.labels_
pdf.head()
import matplotlib.cm as cm
n_clusters = max(agglom.labels_)+1
colors = cm.rainbow(np.linspace(0, 1, n_clusters))
cluster_labels = list(range(0, n_clusters))
# Create a figure of size 6 inches by 4 inches.
plt.figure(figsize=(16,14))
for color, label in zip(colors, cluster_labels):
subset = pdf[pdf.cluster_ == label]
for i in subset.index:
plt.text(subset.horsepow[i], subset.mpg[i],str(subset['model'][i]), rotation=25)
plt.scatter(subset.horsepow, subset.mpg, s= subset.price*10, c=color, label='cluster'+str(label),alpha=0.5)
# plt.scatter(subset.horsepow, subset.mpg)
plt.legend()
plt.title('Clusters')
plt.xlabel('horsepow')
plt.ylabel('mpg')
```
As you can see, we are seeing the distribution of each cluster using the scatter plot, but it is not very clear where is the centroid of each cluster. Moreover, there are 2 types of vehicles in our dataset, "truck" (value of 1 in the type column) and "car" (value of 0 in the type column). So, we use them to distinguish the classes, and summarize the cluster. First we count the number of cases in each group:
```
pdf.groupby(['cluster_','type'])['cluster_'].count()
```
Now we can look at the characteristics of each cluster:
```
agg_cars = pdf.groupby(['cluster_','type'])['horsepow','engine_s','mpg','price'].mean()
agg_cars
```
It is obvious that we have 3 main clusters with the majority of vehicles in those.
**Cars**:
* Cluster 1: with almost high mpg, and low in horsepower.
* Cluster 2: with good mpg and horsepower, but higher price than average.
* Cluster 3: with low mpg, high horsepower, highest price.
**Trucks**:
* Cluster 1: with almost highest mpg among trucks, and lowest in horsepower and price.
* Cluster 2: with almost low mpg and medium horsepower, but higher price than average.
* Cluster 3: with good mpg and horsepower, low price.
Please notice that we did not use **type** and **price** of cars in the clustering process, but Hierarchical clustering could forge the clusters and discriminate them with quite a high accuracy.
```
plt.figure(figsize=(16,10))
for color, label in zip(colors, cluster_labels):
subset = agg_cars.loc[(label,),]
for i in subset.index:
plt.text(subset.loc[i][0]+5, subset.loc[i][2], 'type='+str(int(i)) + ', price='+str(int(subset.loc[i][3]))+'k')
plt.scatter(subset.horsepow, subset.mpg, s=subset.price*20, c=color, label='cluster'+str(label))
plt.legend()
plt.title('Clusters')
plt.xlabel('horsepow')
plt.ylabel('mpg')
```
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="https://www.ibm.com/analytics/spss-statistics-software?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://www.ibm.com/cloud/watson-studio?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01">Watson Studio</a>
### Thank you for completing this lab!
## Author
Saeed Aghabozorgi
### Other Contributors
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01" target="_blank">Joseph Santarcangelo</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | --------------------------------------------------- |
| 2021-01-11 | 2.2 | Lakshmi | Changed distance matrix in agglomerative clustering |
| 2020-11-03 | 2.1 | Lakshmi | Updated URL |
| 2020-08-27 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
# Foundations of Computational Economics #10
by Fedor Iskhakov, ANU
<img src="_static/img/dag3logo.png" style="width:256px;">
## Two simple algorithms: parity and max
<img src="_static/img/lab.png" style="width:64px;">
<img src="_static/img/youtube.png" style="width:65px;">
[https://youtu.be/fKFZZc77if0](https://youtu.be/fKFZZc77if0)
Description: Parity of a number, bitwise operations in Python. Finding maximum in an array.
### Divisibility by number base
Whether a decimal number is divisible by 10 can be easily seen from its last digit.
Similarly, whether a binary number is divisible by 2 can be easily seen from its last digit.
**If last digit of a number is 0, it is divisible by its base!**
#### Parity of a number algorithm
1. Convert the number to binary
1. Check if last digit is zero
- All integers already have a clear binary representation
*This algorithm only applies to integers*
### Bitwise operations in Python
- bitwise AND **&**
- bitwise OR **|**
- bitwise XOR **^**
- bitwise NOT **~** (including sign bit!)
- right shift **>>**
- left shift **<<** (without overflow!)
#### Bitwise AND, OR and XOR
<img src="_static/img/bitwise.png" style="width:600px;">
```
# bitwise logic
a,b = 3,5 # 3=0011, 5=0101
print(' a = {0:d} ({0:04b})\n b = {1:d} ({1:04b})'.format(a,b))
print('a&b = {0:d} ({0:04b})'.format(a&b))
# print('a|b = {0:d} ({0:04b})'.format(a|b))
# print('a^b = {0:d} ({0:04b})'.format(a^b))
```
#### Bit shifts in Python
<img src="_static/img/bitshift.png" style="width:600px;">
#### Replacing arithmetic operations with bit operations
Is it possible?
Which operations can be done in this *geeky* way?
```
# bit shifts
a = 0b11100011
b = a >> 1
print(' a = {0:4d} ({0:016b})\n b = {1:4d} ({1:016b})\n'.format(a,b))
b = a << 2
print(' a = {0:4d} ({0:016b})\n b = {1:4d} ({1:016b})\n'.format(a,b))
# arythmetic operations with bit shifts
a = 0b11100011
print(' a = {0:4d} ({0:016b})'.format(a))
for i in range(1,10):
x = 2**i
d = a//x
s = a>>i
print('a//%d = %d, a>>%d = %d' % (x,d,i,s))
```
### Parity algorithm
Run a single bitwise AND operation to
compare against **0b0000001** which is simply 1 in decimal
Complexity is constant because only one bit must be checked!
*However, when running AND are all bits checked?*
```
# parity check
def parity (n,verbose=False):
'''Returns 1 if passed integer number is odd
'''
pass
# check parity of various numbers
for n in [2,4,7,32,543,671,780]:
print('n = {0:5d} ({0:08b}), parity={}'.format(n,parity(n)))
def parity (n,verbose=False):
'''Returns 1 if passed integer number is odd
'''
if not isinstance(n, int): raise TypeError('Only integers in parity()')
if verbose: print('n = {:08b}'.format(n)) # print binary form of the number
return n & 1 # bitwise and operation returns the value of last bit
```
### Finding max/min in a list
- In the worst case, there is no way to avoid checking *all elements*
- Complexity is linear in the number of elements on the list
```
def maximum_from_list (vars):
'''Returns the maximum from a list of values
'''
pass
# find maximum in some random lists
import numpy as np
for i in range(5):
list = np.random.uniform(low=0.0, high=100.0, size=10)
m = maximum_from_list(list)
print('Maximum in {} is {:.2f}'.format(list,m))
def maximum_from_list (vars):
'''Returns the maximum from a list of values
'''
m=float('-inf') # init with the worst value
for v in vars:
if v > m: m = v
return m
```
### Further learning resources
- Formatting strings
[https://www.digitalocean.com/community/tutorials/how-to-use-string-formatters-in-python-3](https://www.digitalocean.com/community/tutorials/how-to-use-string-formatters-in-python-3)
- Bitwise operations post on Geeksforgeeks
[https://www.geeksforgeeks.org/python-bitwise-operators/](https://www.geeksforgeeks.org/python-bitwise-operators/)
| github_jupyter |
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version Check
Note: The static image export API is available in version <b>3.2.0.+</b><br>
```
import plotly
plotly.__version__
```
### Static Image Export
New in version 3.2.0. It's now possible to programmatically export figures as high quality static images while fully offline.
#### Install Dependencies
Static image generation requires the [orca](https://github.com/plotly/orca) commandline utility and the [psutil](https://github.com/giampaolo/psutil) Python library. There are 3 general approach to installing these dependencies.
##### conda
Using the [conda](https://conda.io/docs/) package manager, you can install these dependencies in a single command:
```
$ conda install -c plotly plotly-orca psutil
```
**Note:** Even if you don't want to use conda to manage your Python dependencies, it is still useful as a cross platform tool for managing native libraries and command-line utilities (e.g. git, wget, graphviz, boost, gcc, nodejs, cairo, etc.). For this use-case, start with [Miniconda](https://conda.io/miniconda.html) (~60MB) and tell the installer to add itself to your system `PATH`. Then run `conda install plotly-orca` and the orca executable will be available system wide.
##### npm + pip
You can use the [npm](https://www.npmjs.com/get-npm) package manager to install `orca` (and its `electron` dependency), and then use pip to install `psutil`:
```
$ npm install -g electron@1.8.4 orca
$ pip install psutil
```
##### Standalone Binaries + pip
If you are unable to install conda or npm, you can install orca as a precompiled binary for your operating system. Follow the instructions in the orca [README](https://github.com/plotly/orca) to install orca and add it to your system `PATH`. Then use pip to install `psutil`.
```
$ pip install psutil
```
### Create a Figure
Now let's create a simple scatter plot with 100 random points of variying color and size.
```
from plotly.offline import iplot, init_notebook_mode
import plotly.graph_objs as go
import plotly.io as pio
import os
import numpy as np
```
We'll configure the notebook for use in [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode
```
init_notebook_mode(connected=True)
N = 100
x = np.random.rand(N)
y = np.random.rand(N)
colors = np.random.rand(N)
sz = np.random.rand(N)*30
fig = go.Figure()
fig.add_scatter(x=x,
y=y,
mode='markers',
marker={'size': sz,
'color': colors,
'opacity': 0.6,
'colorscale': 'Viridis'
})
iplot(fig)
```
### Write Image File
The `plotly.io.write_image` function is used to write an image to a file or file-like python object.
Let's first create an output directory to store our images
```
if not os.path.exists('images'):
os.mkdir('images')
```
If you are running this notebook live, click to [open the output directory](./images) so you can examine the images as they're written.
#### Raster Formats: PNG, JPEG, and WebP
Orca can output figures to several raster image formats including **PNG**, ...
```
pio.write_image(fig, 'images/fig1.png')
```
**JPEG**, ...
```
pio.write_image(fig, 'images/fig1.jpeg')
```
and **WebP**
```
pio.write_image(fig, 'images/fig1.webp')
```
#### Vector Formats: SVG and PDF...
Orca can also output figures in several vector formats including **SVG**, ...
```
pio.write_image(fig, 'images/fig1.svg')
```
**PDF**, ...
```
pio.write_image(fig, 'images/fig1.pdf')
```
and **EPS** (requires the poppler library)
```
pio.write_image(fig, 'images/fig1.eps')
```
**Note:** It is important to note that any figures containing WebGL traces (i.e. of type `scattergl`, `heatmapgl`, `contourgl`, `scatter3d`, `surface`, `mesh3d`, `scatterpolargl`, `cone`, `streamtube`, `splom`, or `parcoords`) that are exported in a vector format will include encapsulated rasters, instead of vectors, for some parts of the image.
### Get Image as Bytes
The `plotly.io.to_image` function is used to return an image as a bytes object.
Let convert the figure to a **PNG** bytes object...
```
img_bytes = pio.to_image(fig, format='png')
```
and then display the first 20 bytes.
```
img_bytes[:20]
```
#### Display Bytes as Image Using `IPython.display.Image`
A bytes object representing a PNG image can be displayed directly in the notebook using the `IPython.display.Image` class. This also works in the [Qt Console for Jupyter](https://qtconsole.readthedocs.io/en/stable/)!
```
from IPython.display import Image
Image(img_bytes)
```
### Change Image Dimensions and Scale
In addition to the image format, the `to_image` and `write_image` functions provide arguments to specify the image `width` and `height` in logical pixels. They also provide a `scale` parameter that can be used to increase (`scale` > 1) or decrease (`scale` < 1) the physical resolution of the resulting image.
```
img_bytes = pio.to_image(fig, format='png', width=600, height=350, scale=2)
Image(img_bytes)
```
### Summary
In summary, to export high-quality static images from plotly.py all you need to do is install orca and psutil and then use the `plotly.io.write_image` and `plotly.io.to_image` functions.
If you want to know more about how the orca integration works, or if you need to troubleshoot an issue, please check out the [Orca Management](../orca-management/) section.
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'static-image-export.ipynb', 'python/static-image-export/', 'Static Image Export | plotly',
'Plotly allows you to save static images of your plots. Save the image to your local computer, or embed it inside your Jupyter notebooks as a static image.',
title = 'Static Image Export | plotly',
name = 'Static Image Export',
thumbnail='thumbnail/static-image-export.png',
language='python',
uses_plotly_offline=True,
page_type='example_index', has_thumbnail='true', display_as='file_settings', order=1,
ipynb='~notebook_demo/252')
```
| github_jupyter |
# 列表
有了前一章关于 *iterable* 和 *iterator* 的知识基础,这一章开始正式学习**数据容器**(*data container*)。
顾名思义,数据容器就是可以装其他数据的数据,Python 有四种基本的数据容器:**列表**(*list*)、**元组**(*tuple*)、**字典**(*dictionary*)、**集合**(*set*),这些容器有共性也有差异,它们都是 *iterable*,所以对这些容器要迭代遍历里面的元素是非常简单的,方法也几乎一样。我们会首先重点介绍**列表**(*list*),然后介绍另外三个,共性的部分基本都会在这一章介绍,后面就重点讲另外三个各自的特色了。
## 列表简介
Python 的列表(*list*),很多编程语言里称为数组(*array*),是最常用的的数据结构之一。有序性是列表的最核心特征,也就是说:
1. 列表里的元素排列是有序的,每个元素有唯一的序号;
2. 第一个元素的序号为 0,第二个序号是 1,依次递增;
3. 可以使用序号来读写指定位置的元素。
Python 的列表中可以是任何类型的数据,一个列表里甚至可以有不同类型的数据,不过一般情况下同一个列表里的数据类型应该是相同的,所以我们可以有整数列表、浮点数列表、字符串列表等等,也可以有列表的列表(列表里的每个元素是一个列表),还可以有我们自定义对象组成的列表。
## 创建列表
Python 提供了若干种方法来创建列表,最基本的就是创建一个空的列表:
```
lst = []
```
要给列表一些初始值的话,就用方括号括起来就行了:
```
lst = [3, 1, 2]
```
列表拥有类型 `list`,这是一个内置的类,有很多定义好的方法给我们使用:
```
type(lst)
```
Python 还提供了一个函数 `list()` 来将其他数据类型转换为列表,可以被转换的数据类型包括:
* **string**:之前我们就说过字符串其实就是字符列表,`list()` 会把字符串转换为其字符组成的数组;
* **tuple**:后面我们会介绍的一种数据容器,和列表其实很像,区别在于 *tuple* 一旦建立其值不可以再更改(*list* 则可以);
* **set**:后面我们会介绍的一种数据容器,里面没有重复元素,`list()` 会把其中所有元素取出来组成一个列表,顺序是不可预期的(因为 *set* 中的元素是没有顺序概念的);
* **dict**:后面我们会介绍的一种数据容器,里面每个元素都是一个 *key-value* 对,就是一个值(value)和它的名字(key),`list()` 会把其中所有的 keys 取出来组成一个列表,顺序是不可预期的(因为 *dictionary* 中的元素是没有顺序概念的);
事实上任何“**可迭代**(*iterable*)”的对象都可以被转换为 *list*,这也是为啥我们要先学 [iterable](p2-7-iterable-iterator.ipynb) 的原因之一。
下面是 `list()` 的例子。
```
# 不带参数调用 list() 返回空列表
print(list())
# 输入字符串,list() 返回字符组成的列表
print(list("aeiou"))
# 输入 tuple(和列表的区别在于不是用方括号而是圆括号),list() 返回元素及排序严格一样的列表
print(list(('a', 'e', 'i', 'o', 'u')))
# 输入 set(用花括号括起来的一组元素),返回其元素组成的列表,顺序不可预测
print(list({'a', 'e', 'i', 'o', 'u'}))
# 输入 dictionary(用花括号括起来的一组 key-value 对,每个元素是 “key: value” 的格式)
# list() 会返回其所有 key 组成的列表,顺序不可预测
print(list({'a': 1, 'e': 2, 'i': 3, 'o':4, 'u': 5}))
```
`list()` 可以把任何 *iterable object*——包括我们要讲的四种数据容器,还有任何迭代器和生成器——转换为列表,比如下面这个来自[上一章](p2-7-iterable-iterator.ipynb)中的例子:
```
from itertools import islice
def fib():
prev, curr = 0, 1
while True:
yield curr
prev, curr = curr, prev + curr
f = fib()
list(islice(f, 0, 10))
```
Python 本身也内置了不少生成器帮我们快速生成列表,比如我们已经熟悉的[等差数列](https://zh.wikipedia.org/wiki/%E7%AD%89%E5%B7%AE%E6%95%B0%E5%88%97)生成器 `range()`,输出的是 *iterable*,所以也可以很方便的变成列表:
```
# range(n) 生成 0 开始但小于 6(不包含)的整数列表:
print(list(range(6)))
# range(n) 生成 3 开始但小于 10(不包含)的整数列表:
print(list(range(3, 10)))
# range(n) 生成 3 开始但小于 10(不包含)、相邻两数相差 2 的整数列表:
print(list(range(3, 10, 2)))
# range(n) 生成 3 开始但小于 20(不包含)、相邻两数相差 4 的整数列表:
print(list(range(3, 20, 4)))
```
## 访问列表元素和列表长度
创建列表之后就可以用列表中元素的序号来访问元素,这个序号一般称为“下标(*index*)”,语法如下:
```
lst = [3, 1, 2]
print(lst[0], lst[1], lst[2])
# Python 允许使用负的序号,-1 表示从最后开始倒数第一个元素,-2 则是倒数第二个,依此类推
print(lst[-1], lst[-2])
# 使用列表序号不仅可以读取指定数据也可以写入,而且可以写入不同类型的数据(虽然一般不推荐这么做)
lst[2] = 'foo'
print(lst)
# len() 函数返回列表的长度,也就是列表中元素的个数
print(len(lst))
```
列表的下标从 0 开始,最大的下标应该是 `len(lst)-1`,如果访问超出列表下标的元素(无论读写)都会产生 `IndexError` 运行时异常,这叫“下标越界(*out of range*)”,这也是最容易出现的程序错误之一。
## 列表切片
我们可以非常简单地从已有列表中切出一段子列表出来,在列表下标中使用 `[m:n]`(m 和 n 都是列表合法整数下标)即返回列表从 `m` 到 `n-1` 的这一段组成的子列表,这个机制一般称为“切片(*slicing*)”。
注意这个语法和上面用下标 `[n]` 访问列表元素很接近,但是含义完全不一样,`[n]` 返回的是列表里的一个元素,类型是该元素的类型;而 `[m:n]` 返回的是一个列表,类型是列表,哪怕里面只有一个元素也如此。
```
nums = list(range(7)) # => [0, 1, 2, 3, 4, 5, 6]
# 切出下标为 2 到 4 的子列表,由于列表的下标是从 0 开始的,所以这个相当于取出了列表第 3 到第 5 的元素
print(nums[2:5])
# 切出下标为 3 及以后的子列表
print(nums[3:])
# 切出下标为 3 以前的子列表
print(nums[:3])
# 从头到尾,其实就是整个列表了,不过切片返回的总是一个新的列表而不是原先的列表,所以这会建立一个原列表的副本
print(nums[:])
# 同样的,负数下标也可以,:-1 就是倒数第一个之前(不包含倒数第一个)的所有元素
print(nums[:-1])
```
上面使用切片的方式会生成原来列表一个片段组成的新列表对象,原列表不会变化。
我们也可以把列表切片放在赋值语句的左边,给列表的某一个片段赋值,这样会直接替换掉原列表中指定的那个片段,比如下面的例子,就是用右边的 `[9, 12]` 替换掉原列表中的第 3 和第 4 个元素:
```
# 如果切片出现在赋值语句左边,
nums[2:4] = [9, 12]
print(nums)
```
上面这个代码其实有不少坑,我们稍微解释下。首先,左边片段的元素个数和右边列表的元素个数不必一致,比如原列表片段有两个元素,右边可以是 0 个 1 个 2 个 3 个以至任何多个元素,这个赋值是吧原片段整体替换为右边的列表,所以可能会改变原列表的元素个数,看下面两个例子:
```
# 右边的列表元素更多,那么都会塞进去,原列表总元素数会变多
nums[2:4] = [10, 11, 12, 13]
print(nums)
# 右边的列表元素较少,原列表总元素数会变少
nums[2:4] = [2]
print(nums)
```
另一个坑是,上面这样的赋值,和单个列表元素的赋值不一样。单个元素赋值时指定的下标是不能越界的,比如上面这个列表目前是 8 个元素,最大下标为 7,`nums[8] = 20` 这样的赋值会抛出 `IndexError: list assignment index out of range` 的下标越界异常;
但切片的赋值是可以的,如果切片范围超过了最大下标,就会把右边的列表扩充到原列表的最后,比如下面的例子,`[8:]` 似乎已越界,但此语句成立,会把 `[7, 8]` 扩充到原列表最后:
```
nums[8:] = [7, 8]
print(nums)
```
这段代码里,不仅 `[8:]` 可以,写 `[100:]` 效果也一样,都是紧跟原列表最后添加上去,不过这种做法并不易于理解,我们要在列表最后增加元素,最好用下面一节介绍的方法。
## 增加、删除元素和其他列表操作
Python 的 `list` 类里面有很多方法可以让我们对已建立的列表进行增删元素等操作。
```
fruits = ['orange', 'apple', 'pear', 'banana']
```
`append(x)` 方法把 x 加入到列表的最后;如上节所述,下面的代码等价于 `fruits[4:] = ['coconut']`,但是要清晰很多:
```
fruits.append('coconut')
print(fruits)
```
`append()` 有个批量版本叫 `extend(x)`,也是把 x 扩展到原列表最后,但这里 x 不是一个元素,而是一组元素,任何 *iterable* 都可以:
```
fruits.extend(['apple', 'banana'])
print(fruits)
```
`insert(i, x)` 把 x 插入到列表下标 i 的元素之前(成为新的下标 i 的元素),比如 `i = 0` 就会插入到原列表最前面:
```
fruits.insert(0, 'kiwi')
print(fruits)
```
`pop()` 删除原列表最后一个元素并返回该元素,原列表长度减一:
```
fruit = fruits.pop()
print(fruit, 'popped')
print(fruits)
```
`remove(x)` 删除列表中第一个出现的值等于 x 的元素,如果整个列表里都没有等于 x 的元素,会扔出一个 `ValueError` 运行时异常
```
fruits.remove('apple')
print(fruits)
```
方法 `reverse()` 将整个列表中的元素排列顺序倒转,注意这个方法是直接修改原列表的:
```
fruits.reverse()
print(fruits)
# clear() 删除列表中的所有元素,还原为一个空列表 []
fruits.clear()
print(fruits)
```
列表里每个元素都有下标,我们可以把上面的操作分成两类:
* 一类是在列表尾部进行的操作,不影响绝大部分元素的下标,包括 `append`、`extend`、`pop` 这几个;
* 另一类是其他的,它们会在列表的中间增加或者删除元素,那么在操作点之后的所有元素下标都要改变。
一般来说前一类的性能会很好,后面一类要做的处理更多,性能会差一些,尤其是在很大很大的列表中会更明显。
## 查找和排序
```
fruits = ['kiwi', 'orange', 'apple', 'pear', 'banana', 'coconut', 'apple', 'banana']
```
`index(x)` 查找列表中值为 x 的第一个元素并返回其下标,如果没找到会扔出一个 `ValueError` 运行时异常:
```
fruits.index('apple')
```
`index(x)` 还可以指定查找范围,用起始和结束下标表示,包括起始下标但不包括结束下标;如果不指定结束下标,则查找到列表尾:
```
fruits.index('apple', 4)
```
`count(x)` 返回列表中出现值为 x 的元素个数,如果没有找到则返回 0:
```
fruits.count('apple')
fruits.count('grape')
```
列表还有个重要的操作,就是根据列表里元素的值来排序,`sort()` 方法就是用来做排序的,最简单的例子什么参数也不用,`sort()` 会自动按照缺省逻辑从小到大排序,所谓缺省逻辑就是用 Python 内置的 `<` `>` 操作符的逻辑,对字符串来说是字母顺序,对数字来说是大小顺序:
```
# sort() 将整个列表排序
fruits.sort()
print(fruits)
```
`sort()` 允许定制这个大小判断的逻辑,输入一些参数来定制排序的规则,实现非常个性化的效果。在介绍这些定制参数之前我们先来看看排序这个事情的本质。
排序,最终的结果是列表里的元素按照**一定的顺序**排列,那么怎么定义**一定的顺序**呢?要么从小到大排(正序,下面的解说如果没有特别说明,都以正序为例),要么从大到小排(逆序),所以我们做排序之前必须先定义元素间“大小规则”,即随便拿两个元素出来,能分出大小,只有这样才能进行有意义的排序。
我们以后会看到越来越多的例子,要排序的东西是各种各样的,其大小关系并不都是那么直观和显而易见的。比如要对一组人排序,怎么定义两个人之间谁大谁小呢?年龄?身高?体重?工作年限?职位?这就需要我们根据需要去选择和定义。
定义好大小比较的规则之后,计算机可以按照一定的**算法**来进行排序的操作。排序算法是非常好的编程入门练习,有很多经典的排序算法,这些算法本质都一样:**比较和移动**。
以正序排序为例:选取两个元素 `a` 和 `b`,按照我们实现约定好的大小规则作比较,如果 `a <= b` 那么 `a` 应该在前面,`b` 在后面(有少数场景下 `a == b` 的情况需要特别处理,我们暂时不考虑这种情况);反之如果 `a > b`,`a` 应该在后面,`b` 应该在前面;然后对元素做相应的移动。
各种算法的差别,在于选取元素和移动元素的策略,随着策略的不同,进行比较和移动的次数会有巨大差异,那么运行的性能(占用的内存和消耗的时间)都会有明显的差别。有兴趣的话可以看看各种排序算法可视化的演示,比如 [这个](http://sorting.at/)、[这个](https://www.cs.usfca.edu/~galles/visualization/ComparisonSort.html)、或者 [这个](https://www.toptal.com/developers/sorting-algorithms),进而可以找本讲算法的入门教材来看看这些算法的实现,以及怎么评估它们在不同情况下的“**时间复杂度**”。
上面说的可以有空时候去研究,回到我们目前的问题,关键是理解一点:排序的结果由我们定义的“大小规则”决定。比如我们不想按照字母顺序,而想按照每个词包含字母的多少来排序,可以这么写:
```
fruits.sort(key=len)
print(fruits)
```
我们给 `sort()` 方法传入了一个参数名为 `key` 的参数,这个参数是一个**函数**,我们在前面提到过,**函数也是数据**,这里是一个例子:函数也可以作为参数传递。传进来的函数是内置的 `len()` 函数,这么做的效果是:排序的大小规则由这个 `len()` 函数的返回值大小来决定:任取列表中两个元素 `a` `b`,如果 `len(a) < len(b)` 那么 `a` 就比 `b` “小”(这个小是我们特别定义的,不是常规概念的“小”);反之如果 `len(a) > len(b)` 那么 `a` 就比 `b` 大。
`sort()` 方法还可以传一个布尔型参数 `reverse` 进去,告诉它要不要逆序排列(大的在前小的在后),`reverse` 缺省值是 `False`,即不用逆序,就正序排列,如果传入 `reverse=True`,`sort()` 方法会按逆序排列:
```
fruits.sort(key=len, reverse=True)
print(fruits)
```
继续说 `sort()` 的这个 `key` 参数,我们不仅可以传给它一个内置函数(如 `len()`),还可以传自定义的函数,比如我们想按照词的最后一个字母的字母表顺序排序,我们可以这么写:
```
def last_letter(s):
return s[-1]
fruits.sort(key=last_letter)
print(fruits)
```
这个 `last_letter` 函数太简单了,而且就用于排序,我们可以简单地将其用 `lambda` 写成一个**匿名函数**:
```
fruits.sort(key=lambda s:s[-1])
print(fruits)
```
效果完全一样,如果你不记得 `lambda` 可以回去重新看看前面的[相关章节](p2-5-functional-1.ipynb)。
## 列表的遍历
遍历(*traverse*)是指不重不漏地访问数据容器中的每个元素各一次,由于列表是一种有序容器,遍历通常暗含着要按照列表顺序访问的要求。而“访问”的含义是取得元素并做相应的处理。
Python 提供两种对列表进行遍历的方式,一种是我们前面已经熟悉的 `for...in` 循环,实际上对任何 *iterable* 都适用;另外一种称为 *list comprehension*,在某些情况下更加简洁和清晰。先来看看 `for...in` 循环。
```
animals = ['cat', 'dog', 'monkey']
for animal in animals:
print(animal)
```
这个 `for...in` 还有另一种写法,可以带着列表的下标做操作,需要用到 Python 内置函数 `enumerate()`:
```
enumerate(animals).__next__()
# 另一种写法允许我们连带取得每个元素的下标
for index, animal in enumerate(animals):
print(f'#{index}: {animal}')
```
`enumerate()` 函数接受一个 *iterable* 对象,然后会调用其迭代器的 `__next__()` 方法,每次调用会把一个序号和 `__next__()` 返回结果绑在一起,形成一个元素,然后用这些绑着序号的元素生成一个新的迭代器返回,比如 `enumerate(animals)` 会生成下面这个序列的迭代器:
`(0, 'cat'), (1, 'dog'), (2, 'monkey')`
于是上面的 `for...in` 循环每次取出一个 `(0, 'cat')` 这样的元素,然后把里面两个值分别赋给循环变量 `index` 和 `animal`。
小括号括起来的 `(0, 'cat')` 这种数据叫**元组**(*tuple*),我们会在下一章介绍。
### List Comprehension
下面我们来看看 *list comprehension*,这实际上是一种列表变换,就是把一个列表变成另外一个列表,变换后列表的元素是原列表元素按照我们指定算法操作得来的。比如我们有一个人名的列表,现在希望把里面所有的人名都变成全小写,如果采用循环的方式我们会这么写:
```
names = ['Neo', 'Trinity', 'Morpheus', 'Smith']
lowercased = []
for name in names:
lowercased.append(name.lower())
print(lowercased)
```
运用 *list comprehension* 的写法可以更加简洁:
```
lowercased = [name.lower() for name in names]
print(lowercased)
```
上面这个简单的例子可以看出,*list comprehension* 是从一个列表的元素出发构造另一个列表的方法,其格式为:
`lst2 = [expr_of_x for x in lst1]`
含义是“对列表 `lst1` 里的每个元素 `x` 计算表达式 `expr_of_x` 的值并将结果依序组成列表 `lst2`”。
有的时候我们并不需要对所有原来列表里的元素都进行变换,所以上面这个格式还可以带个条件尾巴,变成这样:
`lst2 = [expr_of_x for x in lst1 if cond]`
含义和上面一样但多了个条件:只对 `cond` 表达式值为 `True` 的那些元素进行处理,其他那些就直接忽略了。或者更加通用的:
`lst2 = [expr_of_x1 if cond else expr_of_x2 for x in lst1 ]`
含义是:对 `cond` 表达式值为 `True` 的那些元素 `x` 计算表达式 `expr_of_x1` 的值,否则计算表达式 `expr_of_x2` 的值并放入新列表。下面有个例子:
```
lowercased_good = [name.lower() if name != 'Smith' else name.upper() for name in names]
print(lowercased_good)
```
*List comprehension* 是源自函数式抽象的一种强大而优雅的工具,以后我们会看到更多的应用实例。
## 容器与函数参数
前面我们[提到过](p2-1-function-def.ipynb),全局变量和局部变量是彼此隔离的,即使名字一样也是完全不同的东西,就像下面的代码,函数体内的 n 和外面的 n 其实完全是两个变量,函数体内对 n 执行加一,但外面的全局变量 n 不会改变:
```
def inc(n):
n += 1
n = 0
inc(n)
print(n)
```
当我们用一个数据容器做函数的输入参数时,情况会变得复杂起来。我们来看下面的例子:
```
def reassign(l):
l = [0, 1, 2, 3]
def append(l):
l.append(3)
def modify(l):
l[1] = 100
l = [0, 1, 2]
reassign(l)
print(l)
append(l)
print(l)
modify(l)
print(l)
```
`reassign` `append` `modify` 三个函数都接受一个列表输入,我们发现 `reassign` 没有改变全局变量 `l`——这很好理解,因为函数体内给局部变量 `l` 重新赋值,影响的是局部变量 `l`,全局变量 `l` 是另一个东西,自然不受影响,和前面我们学过的知识点完全吻合。
但是 `append` `modify` 确实改变了全局变量 `l` 的内容,分别给它增加了一个元素,以及修改了一个元素的值,这又是怎么回事呢?
原来,当数据容器(其实对任何对象都如此,数据容器也是对象)作为函数输入参数时,传递的是一种“**引用**(*reference*)”,你可以理解为是对同一个对象的不同名字。按照这个思路我们来详细分析下上面的三个函数。
**reassign(l)**
`l = [0, 1, 2]` 创建了一个全局变量 `l`,并且赋值,这时候内存里有了 `[0, 1, 2]` 这样的一个列表对象,全局变量 `l` 是指向它的一个名字。
当我们调用 `reassign(l)` 时首先吧小括号里的参数值求出来,就是内存里的那个 `[0, 1, 2]` 对象,匹配到函数 `reassign()` 的唯一参数 `l`,这个 `l` 是函数 `reassign` 内的局部变量(所有函数参数都是局部变量),于是局部变量 `l` 和全局变量 `l` 目前都指向内存里同一个对象。
然后函数内执行 `l = [0, 1, 2, 3]`,这一赋值语句实际上在内存中创建了另外一个列表对象 `[0, 1, 2, 3]`,并让局部变量 `l` 指向它,注意这时候全局变量没有动,还是指向原先的那个 `[0, 1, 2]` 对象,所以第一个 `print(l)` 语句输出的还是 `[0, 1, 2]`(全局变量 `l` 指向的对象)。
这个过程和前面输入参数为整数的情形是完全一样的,只要理解了全局变量 `l` 和局部变量 `l` 是两个变量就可以理解。
**append(l)**
前面都一样,函数调用发生时,局部变量 `l` 和全局变量 `l` 都指向内存里同一个 `[0, 1, 2]` 对象。
然后函数内执行 `l.append(3)`,并没有新的对象创建,这个 `append()` 就是针对上述 `[0, 1, 2]` 对象操作的,于是内存中的这个对象在尾部增加了一个元素,变成了 `[0, 1, 2, 3]`。注意,全局变量 `l` 也是指向这个对象的。
所以第二个 `print(l)` 语句输出了变化之后的对象,变成了 `[0, 1, 2, 3]`。
**modify()**
同 `append(l)` 理,不再赘述。
所以,在[前面关于函数和变量作用域的规则](p2-1-function-def.ipynb)基础之上,我们要再补充下面的规则,才算完整:
* 当数据容器等对象作为函数变量是,传递的是指向内存中对象的 *reference*,对这个对象直接进行的操作会改变对象的内容。这样的操作包括修改其元素或属性值,调用其(会改变自身内容)的方法等。
注意有些对象方法本来就不会改变自身内容,而是生成一个新对象返回的,不在此列,比如列表的切片、*comprehension* 这些。
## 小结
*List* 是我们学习的第一个数据容器(可能也会是最多用到的),我们花了不少的时间来介绍了列表的特性:有序、*iterable*,进而学习了列表的各种操作:
* 创建列表的方法,尤其是用 `list()` 可以把任何 *iterable* 转换为列表;
* 用 `len()` 可以获得列表的长度,用 `lst[n]` 可以访问下标 `n` 的元素,注意下标是从 `0` 开始的;
* 用 `lst[n1:n2]` 获得列表下标 `n1` 到 `n2-1` 的**切片**(*slice*);需要格外留意给切片赋值的语法和效果;
* 了解列表增加和删除元素等操作,注意在列表尾部增加删除元素开销较小;
* 了解在列表中查找元素和对列表做排序的方法,注意函数对象和匿名函数在 `sort()` 方法中的应用;
* 熟悉列表遍历的两种方法,尤其是新的 *list comprehensions* 方法;注意匿名函数在 *list comprehensions* 的应用;
* 当容器或者其他对象做函数参数时,传递的是对象引用,在函数内对这个对象的操作可能影响其他指向该对象的变量。
| github_jupyter |
# Open spectrum from root-files
Author:
J. Angevaare // <j.angevaare@nikhef.nl> // 2020-05-25
Until now we have only been dealing with small files that make it easy to see what is going on. Perhaps we want at some point to get more data from the stoomboot computing cluster or the appended root-file as in this folder. This notebook will show how to and we make an exemplary coincidence plot for Ti-44 using much more data than in the previous tutorials.
Below we:
- open a root file using uproot
- show a calibrated spectrum
- show a Ti-44 coincidence plot
## Open the data
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
import numba
import uproot3
import tqdm
import os
try:
import RP2021
except ModuleNotFoundError:
# This module is not installed correctly, let's hack it in
import sys
import os
path = os.path.join(os.path.abspath('.'), '..')
sys.path.append(path)
import RP2021
# Let's look at the first root file in the list above
path = '../data/mx_n_20200104_1055_000055.root'
file = uproot3.open(path)
tree = file['T;2']
data = tree.pandas.df()
# Let's have a look at how the data looks like
data[['channel', 'integral', 'height', 'time', 'istestpulse', 'error', 'baseline', 'rms', 'ratio', 'humid', 'sec']]
```
## plot a spectrum of a source, we are starting with Co60!
```
mask = (
(data['channel'] == 4 ) | (data['channel'] == 5 )
& (data['istestpulse'] == 0)
& (data['error'] == 0)
)
co60 = data[mask]
co60
for ch in np.unique(co60['channel']):
plt.title('Sectrum of ${}^{60}$Co'f', ch{ch}')
selection = (co60['channel']==ch)
plt.hist(co60[selection]['integral'].values, bins = 300, range=[0,3000])
plt.yscale('log')
plt.ylim(10,plt.ylim()[-1])
plt.axvline(1173.2, c = 'r', linestyle = '--')
plt.axvline(1332.5, c = 'b', linestyle = '--')
plt.xlabel('E [keV]')
plt.ylabel('Counts / 10 keV')
plt.show()
```
## Save small selection
For further use cases we want to save a copy where we have somewhat less data such that we can use this to do small tests with.
This is actually where the CSV file that was used in the previous tutorials was created. You can see that we have capped the max number of events in this file to 100000.
```
co60[['channel', 'integral','time']][:100000].to_csv('../data/Co60_sample.csv',index=False)
```
# Ti44
Below, we'll do the same thing as we have done earlier for Co60 in the 3rd session. Now we'll do the same for Ti44 but with much higher statistics!
Can you again explain all of the spectrum we will see below? Good luck!
```
# Let's cut out some data where we are not interested in anyway
mask = ((data['istestpulse'] == 0) & (data['error'] == 0) )
data = data[mask]
print(f'Pay attention, we are going to go through quite a lot of data!\n'
f'The data we are using now is a staggering {len(data)} events (that is ~{int(len(data)/1e6)} million!)')
%%time
matched_ti44 = RP2021.easy_coincidence_matching(data, source='Ti44', check_time_order = False)
%%time
matched_ti44 = RP2021.easy_coincidence_matching(data, source='Ti44', check_time_order = False)
matched_ti44
plt.figure(figsize=(10,7))
plt.hist2d(
matched_ti44['e_ch2'],
matched_ti44['e_ch3'],
bins = 200,
norm=LogNorm(),
range=[[0,2000],[0,2000]]);
plt.plot([511,0], [0,511], linestyle = '--', linewidth = 2, c = 'cyan')
plt.plot([0,511], [511,0], linestyle = '--', linewidth = 2, c = 'lightgreen')
plt.plot([0,511+511], [511+511,0], linestyle = '--', linewidth = 2, c = 'green')
plt.gca().set_aspect(1)
plt.colorbar(label='Counts/bin')
plt.xlabel('Energy ch2 [keV]')
plt.ylabel('Energy ch3 [keV]')
plt.title('Coincidence ${}^{44}$Ti');
plt.axvline(511, c = 'r', linestyle = '--')
plt.axhline(511, c = 'r', linestyle = '--')
plt.axvline(511*2, c = 'b', linestyle = '--')
plt.axhline(511*2, c = 'b', linestyle = '--')
plt.axvline(1157, c = 'purple', linestyle = '--', label = '1157 keV')
plt.axhline(1157, c = 'purple', linestyle = '--', label = '1157 keV')
selected_data = RP2021.select_peak(matched_ti44, 'e_ch2', energy = 511, energy_range = 50)
plt.hist(selected_data['e_ch2'], bins = 200, range=[0,2000], label = 'Selected peak channel 2')
plt.hist(selected_data['e_ch3'], bins = 200, range=[0,2000], label = 'Coincident in channel 3')
plt.hist(matched_ti44['e_ch3'], bins = 200, range=[0,2000], label = 'All data 3', alpha = 0.3)
plt.axvline(511, c = 'r', linestyle = '--', label = '511 keV')
plt.axvline(1157, c = 'green', linestyle = '--', label = '1157 keV')
plt.axvline(1157+511, c = 'purple', linestyle = '--', label = f'{1157+511} keV')
plt.yscale('log')
plt.legend(loc=(1,0))
plt.xlabel('Energy [keV]')
plt.ylabel('Counts / 10 keV')
plt.ylim(ymin=50);
```
### Downloading more data.
You may want to use more data at some point, you can download additional data via:
- https://surfdrive.surf.nl/files/index.php/s/6CBbzKGvCrttLkp
## Generate huge dataframes (high statistics)
### WARNING:
unproper handing may result in high RAM usage and performance loss
I assume you have downloaded **and extracted** the zip files (see above) and the data is stored under ``/data/...``
## Ti44
```
save_dir = f'../data/ti44'
# Great we have the data where we expected it!
!ls $save_dir
# Let's make one giant array of pandas dataframes. We will combine them in a second
combined = []
for f in tqdm.tqdm(os.listdir(save_dir)):
path = os.path.join(save_dir, f)
combined.append(pd.read_csv(path))
%%time
combined_ti44 = pd.concat(combined)
combined_ti44
```
Wow! That is 116 **million** events, that is a lot! Things will take longer to compute.
```
%%time
# Notice that we do have to check the time ordering as the csv-files are loaded in a random order
matched_ti44 = RP2021.easy_coincidence_matching(combined_ti44, source='Ti44', check_time_order = True)
%%time
plt.figure(figsize=(10,7))
plt.hist2d(
matched_ti44['e_ch2'],
matched_ti44['e_ch3'],
bins = 200,
norm=LogNorm(),
range=[[0,2000],[0,2000]]);
plt.plot([511,0], [0,511], linestyle = '--', linewidth = 2, c = 'cyan')
plt.plot([0,511], [511,0], linestyle = '--', linewidth = 2, c = 'lightgreen')
plt.plot([0,511+511], [511+511,0], linestyle = '--', linewidth = 2, c = 'green')
plt.gca().set_aspect(1)
plt.colorbar(label='Counts/bin')
plt.xlabel('Energy ch2 [keV]')
plt.ylabel('Energy ch3 [keV]')
plt.title('Coincidence ${}^{44}$Ti');
plt.axvline(511, c = 'r', linestyle = '--')
plt.axhline(511, c = 'r', linestyle = '--')
plt.axvline(511*2, c = 'b', linestyle = '--')
plt.axhline(511*2, c = 'b', linestyle = '--')
plt.axvline(1157, c = 'purple', linestyle = '--', label = '1157 keV')
plt.axhline(1157, c = 'purple', linestyle = '--', label = '1157 keV')
```
### Co60
we can do the same trick again
```
save_dir = f'../data/co60'
# Great we have the data where we expected it!
!ls $save_dir
# Let's make one giant array of pandas dataframes. We will combine them in a second
combined = []
for f in tqdm.tqdm(os.listdir(save_dir)):
path = os.path.join(save_dir, f)
combined.append(pd.read_csv(path))
%%time
combined_co60 = pd.concat(combined)
combined_co60
%%time
# Notice that we do have to check the time ordering as the csv-files are loaded in a random order
matched_co60 = RP2021.easy_coincidence_matching(combined_co60, source='Co60', check_time_order = True)
plt.figure(figsize=(10,7))
plt.hist2d(
matched_co60['e_ch4'],
matched_co60['e_ch5'],
bins = 200,
norm=LogNorm(),
range=[[0,3000],[0,3000]]);
plt.plot([1332.5,0], [0,1332.5], linestyle = '--', linewidth = 2, c = 'cyan')
plt.plot([0,1173.2], [1173.2,0], linestyle = '--', linewidth = 2, c = 'lightgreen')
plt.plot([0,1173.2+1332.5], [1173.2+1332.5,0], linestyle = '--', linewidth = 2, c = 'green')
plt.gca().set_aspect(1)
plt.colorbar(label='Counts/bin')
plt.xlabel('Energy ch4 [keV]')
plt.ylabel('Energy ch5 [keV]')
plt.title('Coincidence ${}^{60}$Co');
plt.axvline(1173.2, c = 'r', linestyle = '--')
plt.axhline(1173.2, c = 'r', linestyle = '--')
plt.axvline(1332.5, c = 'b', linestyle = '--')
plt.axhline(1332.5, c = 'b', linestyle = '--')
```
| github_jupyter |
# QDA + Pseudo Labeling + Gaussian Mixture = LB 0.975
The dataset for Kaggle competition "Instant Gratification" appears to be 512 datasets concatenated where each sub dataset is believed to be created by Sklearn's `make_classification`. EDA suggests the following parameters:
X, y = make_classification(n_samples=1024, n_features=255, n_informative=33+x,
n_redundant=0, n_repeated=0, n_classes=2, n_clusters_per_class=3,
weights=None, flip_y=0.05, class_sep=1.0, hypercube=True, shift=0.0,
scale=1.0, shuffle=True, random_state=None) # where 0<=x<=14
The important parameters to note are `n_clusters_per_class=3` and `n_informative=33+x`. This means that the data resides in `33+x` dimensional space within 6 hyper-ellipsoids. Each hyper-ellipsoid is a multivariate Gaussian distribution therefore the best classifiers to use are QDA, Pseudo Labeling, and Gaussian Mixture. (See appendix for EDA showing 3 clusters per class).
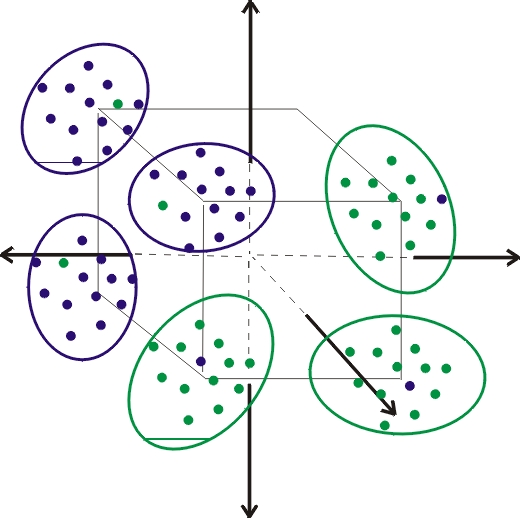
# Better than Perfect Classifier!
If Kaggle generated the data with the above function call to `make_classification` then
Many participants will submit a perfect classifier since the data is easy to separate using QDA and GM. Therefore to win this Kaggle competition, we must submit a better than perfect classifier! The code presented in this kernel has randomness added (and marked in the code below) which allows this classifier to score as much as LB 0.00050 better than a perfect classifier.
If the data is made with Sklearn's `make_classification`, perfect classification is classifying everything correctly except the 2.5% randomly flipped labels. No model can consistently predict the flipped labels correct. However, if you add randomness to your model sometimes it will do better than perfect (and get some flipped targets correct) and sometimes it will do worse. Below is a scatter plot of this kernel's performance. The dotted line is a perfect classifier and each dot is one attempt of this kernel to classify a synthetic dataset (that is similar to this comp's data).
The 200 dots represent 10 attempts made on each of 20 different randomly created synthetic datasets. The black dotted lines were determined by modifying `make_classification` to output the AUC of perfect classification. (See Appendix 3 for more info).
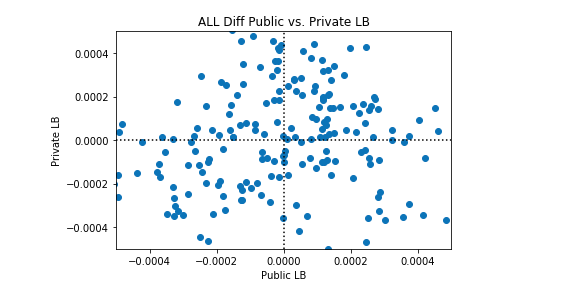
Besides indicating perfect classification on average, this scatter plot also shows that there is no correlation between this kernel's public LB and private LB performance. Therefore for our final submission, we will not choose our highest public LB submission. We will run this kernel 30 times and submit two versions chosen at random regardless of their public LB performance. Then we cross our fingers and hope that those two runs score better than perfect private test dataset classification :P
```
import numpy as np
np.random.seed(42)
x = np.random.choice(np.arange(30),2)
print('We will submit versions',x[0],'and',x[1])
np.random.seed(None)
```
## UPDATE: How Lucky Were We?
Hip hip hooray! `np.random.seed(42)` chose well. After the competition ended, I made the plot below showing this kernel's 30 public and private LB scores. The two randomly chosen final submissions are versions 6 and 19 colored green below. You can see that 19 has the best private LB score :-)
The highest **public** LB is 0.97481 achieved by version 25. The lowest public LB is 0.97439 by version 7. The highest **private** LB is 0.97588 by version 19. The lowest private LB is 0.97543 by version 23. The black dotted lines represent this kernel's public and private prediction averages. (For more info about this plot and the above plot, see Appendix 3).
```
import matplotlib.pyplot as plt, numpy as np
pu = np.array([68,53,70,67,54,54,39,68,60,65,46,62,62,55,54,
60,59,55,43,52,63,68,51,75,81,56,68,55,60,48])
pr = np.array([58,54,68,54,48,61,59,49,60,57,70,54,53,69,72,
56,64,44,88,63,74,70,43,48,77,65,51,70,51,48])
plt.scatter(0.974+pu/1e5,0.975+pr/1e5)
plt.scatter(0.974+pu[5]/1e5,0.975+pr[5]/1e5,color='green',s=100)
plt.scatter(0.974+pu[18]/1e5,0.975+pr[18]/1e5,color='green',s=100)
mpu = 0.974 + np.mean(pu)/1e5
mpr = 0.975 + np.mean(pr)/1e5
plt.plot([mpu,mpu],[mpr-0.0005,mpr+0.0005],':k')
plt.plot([mpu-0.0005,mpu+0.0005],[mpr,mpr],':k')
plt.xlabel('Public LB'); plt.xlim((mpu-0.0005,mpu+0.0005))
plt.ylabel('Private LB'); plt.ylim((mpr-0.0005,mpr+0.0005))
plt.title("Public and private LB scores from 30 runs of this kernel.\n \
The green dots were the two randomly chosen submissions")
plt.show()
```
# Load Libraries and Data
```
# IMPORT LIBRARIES
import numpy as np, pandas as pd, os
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
from sklearn.mixture import GaussianMixture
from sklearn.model_selection import StratifiedKFold
from sklearn.feature_selection import VarianceThreshold
from sklearn.metrics import roc_auc_score
import matplotlib.pyplot as plt
from sklearn.datasets import make_classification
# LOAD TRAIN AND TEST
df_train = pd.read_csv('../input/train.csv')
df_test = pd.read_csv('../input/test.csv')
df_train.head()
```
# Identify Useful Features
Sklearn's `make_classification` leaks which features are useful by increasing their variance.
```
# IDENTIFY USEFUL FEATURES PER MAGIC SUB-DATASET
useful = np.zeros((256,512))
for i in range(512):
partial = df_train[ df_train['wheezy-copper-turtle-magic']==i ]
useful[:,i] = np.std(partial.iloc[:,1:-1], axis=0)
useful = useful > 1.5
useful = np.sum( useful, axis=0 )
```
# Model and Predict
Our model is as follows. First we use QDA plus Pseudo Labeling to predict `test.csv` with CV 0.970 accuracy as previously done [here][1]. Next we will use these predictions (pseudo labels) to find the 6 ellipses. We separately find the 3 ellipses of the target=1 data and 3 ellipses of the target=0 data using Sklearn GaussianMixture. Then we label each point with 0, 1, 2, 3, 4, 5 representing which ellipse it belongs to. Finally we train QDA on these 6 ellipses and use QDA to make our final predictions with `Pr(target=1) = Pr(in ellipse 3) + Pr(in ellipse 4) + Pr(in ellipse 5)`. (See appendix 2 for advanced techniques).
For validation, we didn't use typical k-folds CV. Instead we created synthetic data and optimized our technique on synthetic data. This has proven to be more reliable than CV. Also it allows our model to use all 1024 rows of sub datasets when building models. Our model has demonstrated that on average it can perfectly classify Sklearn's `make_classification` data. However many other participants can too. So randomness is added which allows us to do better than perfect sometimes. Then two random versions' output were submitted to Kaggle and hopefully those are the high scoring ones! :P
When the code below is run locally, it generates synthetic data and calculates validation AUC. When the code is submitted to Kaggle, it uses real data and predicts `test.csv`.
[1]: https://www.kaggle.com/cdeotte/pseudo-labeling-qda-0-969
```
# RUN LOCALLY AND VALIDATE
models = 512
RunLocally = True
# RUN SUBMITTED TO KAGGLE
if len(df_test)>512*300:
repeat = 1
models = 512 * repeat
RunLocally = False
# INITIALIZE
all_preds = np.zeros(len(df_test))
all_y_pu = np.array([])
all_y_pr = np.array([])
all_preds_pu = np.array([])
all_preds_pr = np.array([])
# MODEL AND PREDICT
for k in range(models):
# IF RUN LOCALLY AND VALIDATE
# THEN USE SYNTHETIC DATA
if RunLocally:
obs = 512
X, y = make_classification(n_samples=1024, n_features=useful[k%512],
n_informative=useful[k%512], n_redundant=0, n_repeated=0,
n_classes=2, n_clusters_per_class=3, weights=None, flip_y=0.05,
class_sep=1.0, hypercube=True, shift=0.0, scale=1.0, shuffle=True,
random_state=None)
# IF RUN SUBMITTED TO KAGGLE
# THEN USE REAL DATA
else:
df_train2 = df_train[df_train['wheezy-copper-turtle-magic']==k%512]
df_test2 = df_test[df_test['wheezy-copper-turtle-magic']==k%512]
sel = VarianceThreshold(1.5).fit(df_train2.iloc[:,1:-1])
df_train3 = sel.transform(df_train2.iloc[:,1:-1])
df_test3 = sel.transform(df_test2.iloc[:,1:])
obs = df_train3.shape[0]
X = np.concatenate((df_train3,df_test3),axis=0)
y = np.concatenate((df_train2['target'].values,np.zeros(len(df_test2))))
# TRAIN AND TEST DATA
train = X[:obs,:]
train_y = y[:obs]
test = X[obs:,:]
test_y = y[obs:]
comb = X
# FIRST MODEL : QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.5,0.5])
clf.fit(train,train_y)
test_pred = clf.predict_proba(test)[:,1]
# SECOND MODEL : PSEUDO LABEL + QDA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred)) #randomness
clf = QuadraticDiscriminantAnalysis(priors = [0.5, 0.5])
clf.fit(comb, np.concatenate((train_y,test_pred)) )
test_pred = clf.predict_proba(test)[:,1]
# THIRD MODEL : PSEUDO LABEL + GAUSSIAN MIXTURE
test_pred = test_pred > np.random.uniform(0,1,len(test_pred)) #randomness
all_y = np.concatenate((train_y,test_pred))
least = 0; ct = 1; thx=150
while least<thx:
# STOPPING CRITERIA
if ct>=10: thx -= 10
else: thx = 150
# FIND CLUSTERS
clusters = np.zeros((len(comb),6))
# FIND THREE TARGET=1 CLUSTERS
train4 = comb[ all_y==1, :]
clf = GaussianMixture(n_components=3).fit(train4) #randomness
clusters[ all_y==1, 3:] = clf.predict_proba(train4)
# FIND THREE TARGET=0 CLUSTERS
train4 = comb[ all_y==0, :]
clf = GaussianMixture(n_components=3).fit(train4) #randomness
clusters[ all_y==0, :3] = clf.predict_proba(train4)
# ADJUST CLUSTERS (EXPLAINED IN KERNEL COMMENTS)
for j in range(5): clusters[:,j+1] += clusters[:,j]
rand = np.random.uniform(0,1,clusters.shape[0])
for j in range(6): clusters[:,j] = clusters[:,j]>rand #randomness
clusters2 = 6 - np.sum(clusters,axis=1)
# IF IMBALANCED TRY AGAIN
least = pd.Series(clusters2).value_counts().min(); ct += 1
# FOURTH MODEL : GAUSSIAN MIXTURE + QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.167, 0.167, 0.167, 0.167, 0.167, 0.167])
clf.fit(comb,clusters2)
pds = clf.predict_proba(test)
test_pred = pds[:,3]+pds[:,4]+pds[:,5]
# IF RUN LOCALLY, STORE TARGETS AND PREDS
if RunLocally:
all_y_pu = np.append(all_y_pu, test_y[:256])
all_y_pr = np.append(all_y_pr, test_y[256:])
all_preds_pu = np.append(all_preds_pu, test_pred[:256])
all_preds_pr = np.append(all_preds_pr, test_pred[256:])
# IF RUN SUBMIT TO KAGGLE, PREDICT TEST.CSV
else:
all_preds[df_test2.index] += test_pred / repeat
# PRINT PROGRESS
if ((k+1)%64==0)|(k==0): print('modeled and predicted',k+1,'magic sub datasets')
# IF RUN LOCALLY, COMPUTE AND PRINT VALIDATION AUCS
if RunLocally:
all_y_pu_pr = np.concatenate((all_y_pu,all_y_pr))
all_preds_pu_pr = np.concatenate((all_preds_pu,all_preds_pr))
auc1 = roc_auc_score(all_y_pu_pr, all_preds_pu_pr)
auc2 = roc_auc_score(all_y_pu, all_preds_pu)
auc3 = roc_auc_score(all_y_pr, all_preds_pr)
print()
print('Validation AUC =',np.round(auc1,5))
print('Approx Public LB =',np.round(auc2,5))
print('Approx Private LB =',np.round(auc3,5))
```
# Submit to Kaggle
Alright, let's cross our fingers and hope that our submission has a high private LB !!
```
sub = pd.read_csv('../input/sample_submission.csv')
sub['target'] = all_preds
sub.to_csv('submission.csv',index=False)
plt.hist( test_pred ,bins=100)
plt.title('Model 512 test predictions')
plt.show()
```
# Appendix 1 - EDA revealing n_clusters_per_class = 3
We believe the data was made from Sklearn's `make_classification`. An important question is what parameter for `n_clusters_per_class` did Kaggle use?
According to Sklearn's documentation [here][1]:
> This initially creates clusters of points normally distributed (std=1) about vertices of an n-informative-dimensional hypercube with sides of length 2 * class_sep and assigns an equal number of clusters to each class.
In three dimensions that means, that the clusters will be centered at one of these 8 locations: (-1, -1, -1), (-1, -1, 1), (-1, 1, -1), (-1, 1, 1), (1, -1, -1), (1, -1, 1), (1, 1, -1), (1, 1, 1) where you replace all 1's by `class_sep`. If you create 1024 rows of data and have 2 clusters per class, then for `target=1`, you may have 256 points centered at (-1, 1, -1) and 256 points centered at (1, 1, -1). Then for `target=0`, you may have 256 points centered at (1, 1, 1) and 256 points centered at (-1, -1, 1).
Using EDA, we can determine the number of clusters per class of the real data. Sklearn's `make_classification` generates data (ellipses) at hypercube corners. Therefore if there is only `n_clusters_per_class=1` then the center of each ellipse (target=1 and target=0) of data will have all coordinates 1's and -1's, for example (1,1,-1,1,-1,1,...). So if we plot a histogram of all the variables' means (center coordinates), we will see a bump at 1 and -1. (By variable we mean within each sub dataset. That's 768 rows of train and public test. We don't mean all 262144 rows of original train columns).
If `n_clusters_per_class=2`, then within one sub dataset there will be 2 ellipses for target=1. For example, there may be one ellipse centered at (-1,1,...) and one at (1,1,...) and the first coordinates of 1 and -1 will average to 0 when we compute that variable's mean. Therefore, if `clusters=2`, we will see histogram bumps at -1, 0, and 1. If `n_clusters_per_class=3`, we will see 4 bumps. Etc, etc. We can confirm this with synthetic data. We will use `n_samples=768` because we only have training and public test data to compare with which only has 768 rows per `wheezy-magic` sub dataset.
Afterward, we will plot a histogram of the real data's variable means (within sub datasets) and see which `n_clusters_per_class` it matches. Alternatively, we can build a model and assume that `n_clusters_per_class` equals 1, 2, 3, or 4. Then we check which `n_clusters_per_class` has the greatest CV. Both of these methods determine that `n_clusters_per_class=3`.
[1]: https://scikit-learn.org/stable/modules/generated/sklearn.datasets.make_classification.html
```
for clusters in range(4):
centers = np.array([])
for k in range(512):
X, y = make_classification(n_samples=768, n_features=useful[k],
n_informative=useful[k], n_redundant=0, n_repeated=0,
n_classes=2, n_clusters_per_class=clusters+1, weights=None,
flip_y=0.05, class_sep=1.0, hypercube=True, shift=0.0, scale=1.0,
shuffle=True, random_state=None)
centers = np.append(centers,np.mean(X[ np.argwhere(y==0).flatten() ,:],axis=0))
centers = np.append(centers,np.mean(X[ np.argwhere(y==1).flatten() ,:],axis=0))
plt.hist(centers,bins=100)
plt.title('Variable means if clusters='+str(clusters+1))
plt.show()
```
## Now let's plot the real data
First we will use QDA to create pseudo labels for the public test data. Then we will plot a histogram of the variable means (data centers' coordinates) of target=0 and target=1 using all training and pseudo labeled public test data combined (768 rows per sub dataset). The plot below shows that Kaggle used `n_clusters_per_class=3`.
```
centers = np.array([])
for k in range(512):
# REAL DATA
df_train2 = df_train[df_train['wheezy-copper-turtle-magic']==k]
df_test2 = df_test[df_test['wheezy-copper-turtle-magic']==k]
sel = VarianceThreshold(1.5).fit(df_train2.iloc[:,1:-1])
df_train3 = sel.transform(df_train2.iloc[:,1:-1])
df_test3 = sel.transform(df_test2.iloc[:,1:])
obs = df_train3.shape[0]
X = np.concatenate((df_train3,df_test3),axis=0)
y = np.concatenate((df_train2['target'].values,np.zeros(len(df_test2))))
# TRAIN AND TEST DATA
train = X[:obs,:]
train_y = y[:obs]
test = X[obs:,:]
test_y = y[obs:]
comb = X
# FIRST MODEL : QDA
clf = QuadraticDiscriminantAnalysis(priors = [0.5,0.5])
clf.fit(train,train_y)
test_pred = clf.predict_proba(test)[:,1]
# SECOND MODEL : PSEUDO LABEL + QDA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred))
clf = QuadraticDiscriminantAnalysis(priors = [0.5, 0.5])
clf.fit(comb, np.concatenate((train_y,test_pred)) )
test_pred = clf.predict_proba(test)[:,1]
# PSEUDO LABEL TEST DATA
test_pred = test_pred > np.random.uniform(0,1,len(test_pred))
y[obs:] = test_pred
# COLLECT CENTER COORDINATES
centers = np.append(centers,np.mean(X[ np.argwhere(y==0).flatten() ,:],axis=0))
centers = np.append(centers,np.mean(X[ np.argwhere(y==1).flatten() ,:],axis=0))
# PLOT CENTER COORDINATES
plt.hist(centers,bins=100)
plt.title('Real Data Variable Means (match clusters=3)')
plt.show()
```
# Appendix 2 - Advanced Techniques
Since the code above already classifies Sklearn's `make_classification` on average prefectly, there is no need to improve it. However, below are some ideas that could be used if improvement was possible and/or neccessary.
1. After building model 4's classifier, you could use it to classify the training data. Then all training data with `abs(oof - true)>0.9` are erroneous training data with their labels flippped. Next correct those training labels and run the entire kernel a second time.
2. Since each cluster is centered at a hypercube corner, you can modify Sklearn's Quadratic Discriminant Analysis code by adding `meang[ np.argwhere(meang>=0) ] = 1.0` and `meang[ np.argwhere(meang<0) ] = -1.0`. This moves the centers of all clusters to hypercube corners.
3. Computer accuracy cannot distinguish between predictions that are close to 1. Using an example with 6 digit accuracy, the numbers 1.000001 and 1.000002 are the same because both become 1.00000. To help improve AUC, you can add the following code to this kernel. `temp = np.log(pds[:,0]+pds[:,1]+pds[:,2])`; `temp[ np.isinf(temp) ] = -1e6`; `test_pred -= temp`. This improves AUC by differentiating between predictions close to 1. Note that this isn't a problem for predictions close to 0 because the numbers 0.0000001 and 0.0000002 are 1.0e-7 and 2.0e-7 and the computer can already differentiate them.
4. After making predictions for `test.csv`, you can use them as pseudo labels and run the entire kernel a second time. Then use those labels and run the kernel a third time. Each iteration can give a slight boost.
5. You can run this kernel multiple times and take an average. Or use k-folds. This removes this code's variance (randomness) and achieves close to prefection everytime but it also removes the possibility of scoring LB 0.00050 more or less than perfection.
6. We can also remove this code's variance (randomness) by modifying Sklearn's code for Quadratic Discriminant Analysis and Gaussian Mixture. Each of these models can only accept training labels that are either 0 or 1. By adding a few lines of code, we can allow these models to accept continuous probabilities and use them as weights. This would allow us to remove the randomization line `test_pred = test_pred > np.random.uniform(0,1,len(test_pred))`. Instead we can leave pseudo labels as probabilities between 0 and 1 and still call `QuadraticDiscriminantAnalysis.fit(test_data,test_pred)`.
Using combinations of these additional advanced techniques, this kernel was able to score LB 0.97489 on this competition's public leaderboard. But validation showed that these techniques didn't make the basic perfect classifier any more perfect. Therefore for final submission, the basic classifier was used.
# Appendix 3 - Final Submission Strategy
By applying our model to synthetic data, we can learn how it performs on a simulated public and private leaderboard. We observe that this kernel achieves perfection on average (if Kaggle used `make_classification` with the parameters we suspect). Sklearn's code for `make_classification` includes
# Randomly replace labels
if flip_y >= 0.0:
flip_mask = generator.rand(n_samples) < flip_y
y[flip_mask] = generator.randint(n_classes, size=flip_mask.sum())
Before variable `y` gets rewritten, you can store it by adding `y_orig = y.copy()`. Next update the shuffle line to `X, y, y_orig = util_shuffle(X, y, y_orig, random_state=generator)`. Then change the last line of `make_classification` to `return X, y, y_orig`. By doing this, we can compute the AUC of a perfect classifier with `prefect = roc_auc_score(y, y_orig)`.
Now we can make hundreds of synthetic datasets that are the similar to this competition's data and apply this kernel to see how well it does compared with a perfect classifier. For each synthetic dataset, we will run this kernel 10 times. This will show us patterns and help us decide how to choose our two final submissions.
We observe that sometimes this kenel does better than perfect and sometimes it does worse than perfect. It's interesting to note that there is no correlation between its performance on public LB versus private LB. In the example plots below, perfect classification is represented by the black dotted line.
If we take the average of this kernel's performance over many sythnetic datasets, it achieves perfect classification. Therefore there is no reason to improve this kernel. We can never expect to perform better than perfect classification on average. The only change we could consider is altering this kernel's standard deviation from perfection. We could either try to achieve perfection every kernel run, or (leave it as is and) randomly try to exceed perfection by a desired amount on some kernel runs.
## Synthetic Dataset 1
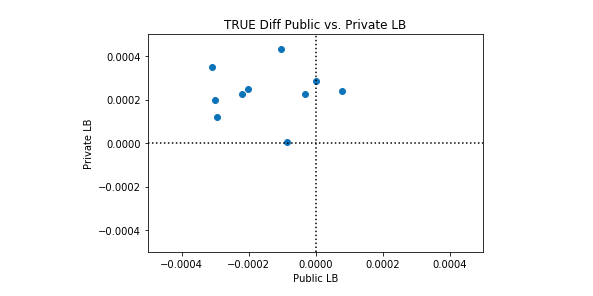
## Synthetic Dataset 2
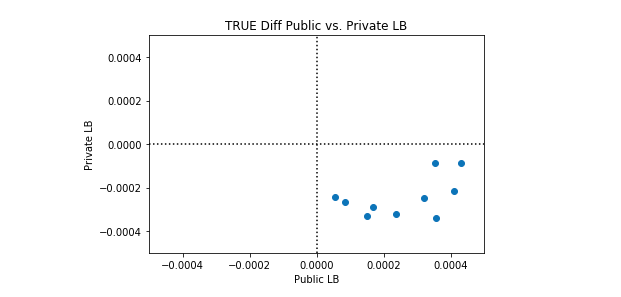
## Synthetic Dataset 3
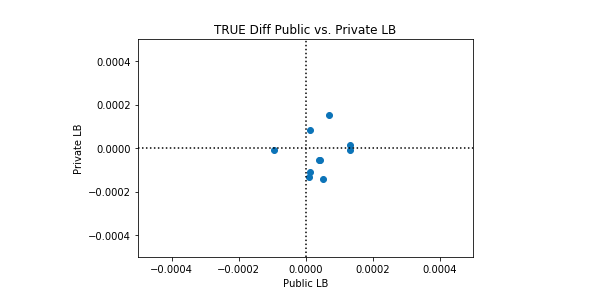
## Synthetic Dataset 4
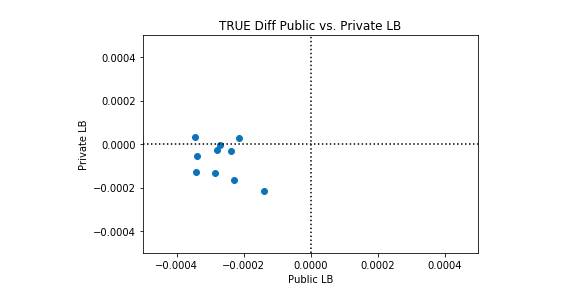
## Synthetic Dataset 5
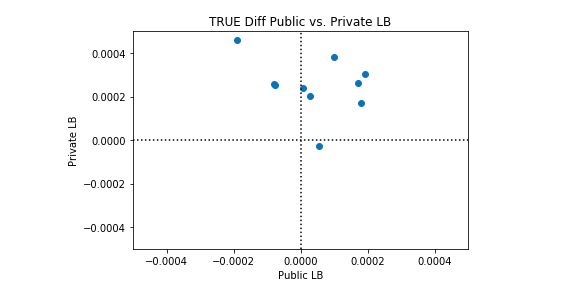
## Many Synthetic Datasets Together

| github_jupyter |
**The Sparks Foundation Internship Program**
**Data Science & Business Analytics Internship**
**Technical TASK 1 :- Prediction using Supervised ML**
In this task, we will predict the score of student based upon the number of hours they studied. This is a simple linear regression task as it involves just two variables.
Author: NOMAN SAEED SOOMRO
**Step 1: Importing Required Libraries**
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
```
**Step 2 : Reading Data from online source**
```
url="https://raw.githubusercontent.com/AdiPersonalWorks/Random/master/student_scores%20-%20student_scores.csv"
data=pd.read_csv(url)
data.head()
data.isnull().sum()
```
**Step 3: Plotting the distribution of scores**
```
data.plot(x='Hours', y='Scores', style='+')
plt.title('Hours vs Percentage')
plt.xlabel('Hours Studied')
plt.ylabel('Percentage Score')
plt.show()
```
**Step 4 : Preparing The Data and splitting the data**
The next step is to divide the data into "attributes"/"X" (inputs) and "labels"/"Y" (outputs).Splitting the data into training data-set and test data-set.
```
X= data.iloc[:, :-1].values
Y= data.iloc[:, 1].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,Y,test_size=0.2, random_state=0)
```
**Step 5 : Model Training**
```
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
print("Model Trained")
# Plotting the regression line
line = regressor.coef_*X+regressor.intercept_
# Plotting for the test data
plt.scatter(X, y)
plt.plot(X, line);
plt.show()
print(X_test) # Testing data - In Hours
y_pred = regressor.predict(X_test) # Predicting the scores
```
**Step 6 : Comparing Actual vs Predicted**
```
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
df
```
**Step 7:
Predicting the output on the New Data.**
```
### Testing your own data.
hours = 9.25
test = np.array([hours])
test = test.reshape(-1,1)
own_pred = regressor.predict(test)
print ("No. of Hours = {}".format(hours))
print ("Predicted Score = {}".format(own_pred[0]))
```
**Step 8 : Evaluating the model**
The final step is to evaluate the performance of algorithm. This step is particularly important to compare how well different algorithms perform on a particular dataset. For simplicity here, we have chosen the mean square error. There are many such metrics.
```
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root mean squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
```
| github_jupyter |
```
values = [10,8,12,11,7,10,8,9,12,11,10]
import numpy as np
np.mean(values)
values = [10,8,12,11,7,10,8,9,12,11,10]
import numpy as np
np.median(values)
values = [10,8,12,11,7,10,8,9,12,11,10]
import statistics
statistics.mode(values)
values = [10,8,12,11,7,10,8,9,12,11,10]
import numpy as np
np.std(values, ddof=1)
values = [10,8,12,11,7,10,8,9,12,11,10]
import numpy as np
np.var(values, ddof=1)
values = [10,8,12,11,7,10,8,9,12,11,10]
import scipy.stats
scipy.stats.iqr(values)
values_x = [10,8,12,11,7,10,8,9,12,11,10]
values_y = [12,9,11,11,8,11,9,10,14,10,9]
import numpy as np
np.corrcoef(values_x,values_y)
import pandas as pd
data_batch = pd.DataFrame({
'temperature': [10, 11, 10, 11, 12, 11, 10, 9, 10, 11, 12, 11, 9, 12, 11],
'pH': [5, 5.5, 6, 5, 4.5, 5, 4.5, 5, 4.5, 5, 4, 4.5, 5, 4.5, 6]
})
data_batch
def super_simple_alert(datapoint):
if datapoint[‘temperature’] < 10:
print(‘this is a real time alert. Temp too low’)
if datapoint[‘pH’] > 5.5:
print(‘this is a real time alert. pH too high’)
data_iterable = data_batch.iterrows()
for i,new_datapoint in data_iterable:
print(new_datapoint.to_json())
super_simple_alert(new_datapoint)
import numpy as np
def super_simple_alert(hist_datapoints):
print(hist_datapoints)
if np.mean(hist_datapoints['temperature']) < 10:
print('this is a real time alert. temp too low')
if np.mean(hist_datapoints['pH']) > 5.5:
print('this is a real time alert. pH too high')
data_iterable = data_batch.iterrows()
# create historization for window
hist_temp = []
hist_ph = []
for i,new_datapoint in data_iterable:
hist_temp.append([new_datapoint['temperature']])
hist_ph.append([new_datapoint['pH']])
hist_datapoint = {
'temperature': hist_temp[-3:],
'pH': hist_ph[-3:]
}
super_simple_alert(hist_datapoint)
import numpy as np
def super_simple_alert(hist_datapoints):
print(hist_datapoints)
if np.std(hist_datapoints['temperature']) > 1:
print('this is a real time alert. temp variations too high')
if np.std(hist_datapoints['pH']) > 1:
print('this is a real time alert. pH variations too high')
data_iterable = data_batch.iterrows()
# create historization for window
hist_temp = []
hist_ph = []
for i,new_datapoint in data_iterable:
hist_temp.append([new_datapoint['temperature']])
hist_ph.append([new_datapoint['pH']])
hist_datapoint = {
'temperature': hist_temp[-3:],
'pH': hist_ph[-3:]
}
super_simple_alert(hist_datapoint)
```
| github_jupyter |
# matplotlib
- matplotlib는 Python에서 차트나 플롯을 시각화하는 패키지
- [홈페이지](http://matplotlib.org/gallery.html)
- matplotlib 내부에 pylab 서브패키지는 matlab 수치해석 소프트웨어 시각화 명령을 그대로 사용!
- ipython에선 %matplotlib inline을 통해 바로 그림을 볼 수 있음
```
import matplotlib as mpl
import matplotlib.pylab as plt
import numpy as np
import pandas as pd
import matplotlib.ticker as tkr
%matplotlib inline
# plt의 내부 메소드
dir(plt)
# 라인플롯
plt.plot([1,4,5,10])
plt.plot([1, 4, 9, 16], c="b", lw=5, ls="--", marker="o", ms=15, mec="g", mew=5, mfc="r")
plt.xlim(-0.2, 3.2)
plt.ylim(-1, 18)
plt.show()
# 여러개의 그래프 표현
t = np.arange(0., 5., 0.2)
plt.plot(t, t, 'r--', t, 0.5*t**2, 'bs:', t, 0.2*t**3, 'g^-')
plt.show()
# matplotlib 1.5까진 hold(True)옵션을 줬어야했으나, matplotlib 2.0부터는 모든 플롯 명령에 hold(True)가 자동 적용
plt.plot([1, 4, 9, 16], c="b", lw=5, ls="--", marker="o", ms=15, mec="g", mew=5, mfc="r")
# plt.hold(True) # <- 1,5 버전에서는 이 코드가 필요하다.
plt.plot([9, 16, 4, 1], c="k", lw=3, ls=":", marker="s", ms=10, mec="m", mew=5, mfc="c")
# plt.hold(False) # <- 1,5 버전에서는 이 코드가 필요하다.
plt.show()
```
## 범례는 legend를 사용
```
X = np.linspace(-np.pi, np.pi, 256)
C, S = np.cos(X), np.sin(X)
plt.plot(X, C, label="cosine")
plt.plot(X, S, label="sine")
plt.legend(loc=2)
plt.show()
```
## 부가 설명은 plt.annotate를 사용
```
Signature: plt.annotate(*args, **kwargs)
Docstring:
Annotate the point ``xy`` with text ``s``.
Additional kwargs are passed to `~matplotlib.text.Text`.
Parameters
----------
s : str
The text of the annotation
xy : iterable
Length 2 sequence specifying the *(x,y)* point to annotate
xytext : iterable, optional
Length 2 sequence specifying the *(x,y)* to place the text
at. If None, defaults to ``xy``.
xycoords : str, Artist, Transform, callable or tuple, optional
The coordinate system that ``xy`` is given in.
For a `str` the allowed values are:
================= ===============================================
Property Description
================= ===============================================
'figure points' points from the lower left of the figure
'figure pixels' pixels from the lower left of the figure
'figure fraction' fraction of figure from lower left
'axes points' points from lower left corner of axes
'axes pixels' pixels from lower left corner of axes
'axes fraction' fraction of axes from lower left
'data' use the coordinate system of the object being
annotated (default)
'polar' *(theta,r)* if not native 'data' coordinates
================= ===============================================
If a `~matplotlib.artist.Artist` object is passed in the units are
fraction if it's bounding box.
If a `~matplotlib.transforms.Transform` object is passed
in use that to transform ``xy`` to screen coordinates
If a callable it must take a
`~matplotlib.backend_bases.RendererBase` object as input
and return a `~matplotlib.transforms.Transform` or
`~matplotlib.transforms.Bbox` object
If a `tuple` must be length 2 tuple of str, `Artist`,
`Transform` or callable objects. The first transform is
used for the *x* coordinate and the second for *y*.
See :ref:`plotting-guide-annotation` for more details.
Defaults to ``'data'``
textcoords : str, `Artist`, `Transform`, callable or tuple, optional
The coordinate system that ``xytext`` is given, which
may be different than the coordinate system used for
``xy``.
All ``xycoords`` values are valid as well as the following
strings:
================= =========================================
Property Description
================= =========================================
'offset points' offset (in points) from the *xy* value
'offset pixels' offset (in pixels) from the *xy* value
================= =========================================
defaults to the input of ``xycoords``
arrowprops : dict, optional
If not None, properties used to draw a
`~matplotlib.patches.FancyArrowPatch` arrow between ``xy`` and
``xytext``.
If `arrowprops` does not contain the key ``'arrowstyle'`` the
allowed keys are:
========== ======================================================
Key Description
========== ======================================================
width the width of the arrow in points
headwidth the width of the base of the arrow head in points
headlength the length of the arrow head in points
shrink fraction of total length to 'shrink' from both ends
? any key to :class:`matplotlib.patches.FancyArrowPatch`
========== ======================================================
If the `arrowprops` contains the key ``'arrowstyle'`` the
above keys are forbidden. The allowed values of
``'arrowstyle'`` are:
============ =============================================
Name Attrs
============ =============================================
``'-'`` None
``'->'`` head_length=0.4,head_width=0.2
``'-['`` widthB=1.0,lengthB=0.2,angleB=None
``'|-|'`` widthA=1.0,widthB=1.0
``'-|>'`` head_length=0.4,head_width=0.2
``'<-'`` head_length=0.4,head_width=0.2
``'<->'`` head_length=0.4,head_width=0.2
``'<|-'`` head_length=0.4,head_width=0.2
``'<|-|>'`` head_length=0.4,head_width=0.2
``'fancy'`` head_length=0.4,head_width=0.4,tail_width=0.4
``'simple'`` head_length=0.5,head_width=0.5,tail_width=0.2
``'wedge'`` tail_width=0.3,shrink_factor=0.5
============ =============================================
Valid keys for `~matplotlib.patches.FancyArrowPatch` are:
=============== ==================================================
Key Description
=============== ==================================================
arrowstyle the arrow style
connectionstyle the connection style
relpos default is (0.5, 0.5)
patchA default is bounding box of the text
patchB default is None
shrinkA default is 2 points
shrinkB default is 2 points
mutation_scale default is text size (in points)
mutation_aspect default is 1.
? any key for :class:`matplotlib.patches.PathPatch`
=============== ==================================================
Defaults to None
annotation_clip : bool, optional
Controls the visibility of the annotation when it goes
outside the axes area.
If `True`, the annotation will only be drawn when the
``xy`` is inside the axes. If `False`, the annotation will
always be drawn regardless of its position.
The default is `None`, which behave as `True` only if
*xycoords* is "data".
Returns
-------
Annotation
plt.plot(X, S, label="sine")
plt.scatter([0], [0], color="r", linewidth=10)
plt.annotate(r'$(0,0)$', xy=(0, 0), xycoords='data', xytext=(-50, 50),
textcoords='offset points', fontsize=16,
arrowprops=dict(arrowstyle="->", linewidth=3, color="g"))
plt.show()
```
# Figure의 이해
<img src='https://datascienceschool.net/upfiles/4e20efe6352e4f4fac65c26cb660f522.png' height="400" width="400">
figure : 도화지라고 생각하면 편함
plt.figure(figsize=(10,2)) 또는 plt.gcf()을 사용해 생성
```
plt.figure(figsize=(10,5))
plt.plot(np.random.randn(100))
plt.show()
# gcf로 figure를 2개 생성해서 그림
f1=plt.gcf()
plt.plot(np.random.randn(100))
f2=plt.gcf()
plt.plot(np.random.randn(100))
```
# Axes, Subplot
- 하나의 윈도우(Figure)에 여러개의 플롯을 배열 형태로 보여야 하는 경우-!
- subplot 명령으로 명시적으로 Axes 객체를 얻어야 함
- plt.subplot(a,b,c) : a,b는 행렬의 모양을 뜻하고 c는 axb개중 몇번째인지 나타냄
- plt.subplot(211) 같이 줄여서 작성 가능
```
x1 = np.linspace(0.0, 5.0)
x2 = np.linspace(0.0, 2.0)
y1 = np.cos(2 * np.pi * x1) * np.exp(-x1)
y2 = np.cos(2 * np.pi * x2)
ax1 = plt.subplot(2, 1, 1)
plt.plot(x1, y1, 'yo-')
plt.title('A tale of 2 subplots')
plt.ylabel('Damped oscillation')
print(ax1)
ax2 = plt.subplot(2, 1, 2)
plt.plot(x2, y2, 'r.-')
plt.xlabel('time (s)')
plt.ylabel('Undamped')
print(ax2)
plt.show()
```
# pandas와 연계
```
dates = pd.date_range('20160620', periods=6)
df = pd.DataFrame(np.random.randn(6,4), index=dates, columns=list('ABCD'))
df
fig, axes = plt.subplots(nrows=2, ncols=2)
df1.plot(ax=axes[0,0])
df2.plot(ax=axes[0,1])
# subplot을 True로 주고 layout 설정-! (이 경우 column name이 기준이 됩니다 )
df.plot(subplots=True, layout=(2,2), figsize=[10,5])
plt.show()
```
### subplot을 False로 주는 경우
```
df.plot(subplots=False, figsize=[10,5])
plt.show()
```
# ticks 설정
- plt.xticks(fontsize=14, rotation=90)
- matplotlib.text.Text 클래스에 있는 것들은 모두 사용가능
locs, labels = xticks()
# plot y축 reverse
- plt.gca().invert_yaxis()
# 글자크기 조절
- mpl.rcParams['font.size'] = 20
- mpl.rcParams['xtick.labelsize'] = 20
- plt.title('platform', fontsize=25)
# layout 조절
- plt.tight_layout()
# colobar
- cb = plt.colorbar()
# y축 쉼표 추가하는 방법
- ax.yaxis.set_major_formatter(
tkr.FuncFormatter(lambda y, p: format(int(y), ',')))
```
data = pd.DataFrame(np.random.randint(1500, size=(100, 2)), columns=['meas', 'modeled'])
# data = pd.read_csv('data.csv')
x = data['meas']
y = data['modeled']
xmin = 0
xmax = 1500
ymin = 0
ymax = 1500
fig, ax = plt.subplots()
plt.hexbin(x, y, cmap=plt.cm.gnuplot2_r)
plt.axis([xmin, xmax, ymin, ymax])
plt.xlabel("Measured baseflow, in cfs")
plt.ylabel("Simulated baseflow, in cfs")
cb = plt.colorbar()
cb.set_label('count')
p2, = plt.plot([0,1500],[0,1500], c='g')
l2 = plt.legend([p2], ["1:1 Line"], loc=2)
ax.yaxis.set_major_formatter(
tkr.FuncFormatter(lambda y, p: format(int(y), ',')))
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/yohanesnuwara/mem/blob/master/01_loaddata.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Load Data
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
```
Access Google Drive to get data
```
from google.colab import drive
drive.mount('/content/drive')
import sys
sys.path.append('/content/drive/My Drive/Colab Notebooks')
```
46 LAS files from Norne Field.
```
import glob
import os
file_path = "/content/drive/My Drive/Colab Notebooks/well_logs"
read_files = glob.glob(os.path.join(file_path, "*.las"))
read_files
```
Get the name of the well from the file.
```
well_names = []
for files in read_files:
files = os.path.splitext(os.path.basename(files))[0]
well_names.append(files)
well_names
```
If the well name too long, shorten it by renaming it.
```
well_names = np.array(well_names)
wellnames = np.array(['B1BH', 'E2AH', 'B1AHT2', 'K1HT2',
'B1H', 'B2H', 'B3H', 'B4AH',
'B4BH', 'B4CH', 'B4DHT2', 'B4H',
'C1H', 'C2H', 'C3H', 'C4AH',
'C4H', 'D1AH', 'D1BH', 'D1CH',
'D1H', 'D2HT2', 'D3AH', 'D3BY1HT2',
'D3BY2H', 'D3H', 'D4AH', 'D4AHT2',
'D4H', 'E1H', 'E2H', 'E3AH',
'E3AHT2', 'E3BH', 'E3CHT2','E3H',
'E4AH', 'E4AHT2', 'E4H','E4HT2',
'F1H', 'F2H', 'F3H',
'F4H','K1H', 'K3H'])
```
Import `lasio` library to import LAS data
```
!pip install lasio
import lasio
```
Read LAS file (if command: `Header section Parameter regexp=~P was not found.`, it's OK)
```
lases = []
for files in read_files:
las = lasio.read(files)
lases.append(las)
```
## Well catalogue
Input the name of the well you want to view in the `find` and check what data is present.
```
wellnames
# type the numbers from 0 to 45 in the wellames[...]
find = wellnames[0]
# print the name of the searched well
print("Well name:", find)
id_ = np.int64(np.where(wellnames==find)).item()
print("Data available:", lases[id_].keys())
# check more details
print("Detail about data:")
print(lases[id_].curves)
# peek or overviewing the well log
# which data you want to see
data_view = 'RHOB'
print("Data", data_view, "of well:", find)
plt.figure(figsize=(5,10))
plt.plot((lases[id_][data_view]), (lases[id_]['DEPTH']))
plt.xlabel(data_view); plt.ylabel("Depth (m)")
plt.grid(True)
plt.gca().invert_yaxis()
```
| github_jupyter |
# ML Pipeline Preparation
### In this notebook work we will work on creating ML Pipeline
- Import Necessary Python Modules
- Load data from sqlite database created earlier
- **Build , Train and Evaluate** your model
```
################### Import Python Modules ###########################
#To Handle datasets
import pandas as pd
import numpy as np
#To handle Databases
from sqlalchemy import create_engine
import re
import pickle
import string
import sys
#To Handle text data using Natural Language ToolKit
import nltk
nltk.download(['punkt', 'wordnet','stopwords'])
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem import WordNetLemmatizer
from nltk.stem.porter import PorterStemmer
#Sklearn Libraries for Ml Models
from sklearn.pipeline import Pipeline, FeatureUnion
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.multioutput import MultiOutputClassifier
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
from sklearn.metrics import classification_report
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
```
## 1.Load Dataset from sqlite database
- Use `read_sql_table` to read data from DisasterResponse database
```
engine= create_engine('sqlite:///DisasterResponse.db')
#
df = pd.read_sql_table('DS_messages',engine)
X=df['message']
y=df[df.columns[4:]]
print(X.head(),y.head())
```
## 2.Function to tokenize text
```
def tokenize(text):
"""
INPUT - text - messages column from the table
Returns tokenized text
1. Remove Punctuation and normalize text
2. Tokenize text and remove stop words
3.Use stemmer and Lemmatizer to Reduce words to its root form
"""
# Remove Punctuations and normalize text by converting text into lower case
text = re.sub(r"[^a-zA-Z0-9]", " ", text.lower())
# Tokenize text and remove stop words
tokens = word_tokenize(text)
stop_words = stopwords.words("english")
words = [w for w in tokens if w not in stop_words]
#Reduce words to its stem/Root form
stemmer = PorterStemmer()
stemmed = [stemmer.stem(w) for w in words]
#Lemmatizer - Reduce words to its root form
lemmatizer = WordNetLemmatizer()
lemm = [lemmatizer.lemmatize(w) for w in stemmed]
return lemm
```
## 3.Build Model
Here we have X as messages columns which is the input to the model and y as the 36 categories which results as output classification to our model. As we have multi features to classify we can make use of [MultiOutputClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.multioutput.MultiOutputClassifier.html)
```
#create pipeline
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(RandomForestClassifier()))
])
```
## 4.Train Model
```
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
pipeline.fit(X_train, y_train)
```
## 5.Predict the model
prints [classification_report](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html?highlight=classification_report#sklearn.metrics.classification_report) from sklearn library which returns Precision,Recall and f1-score of your model
```
# Predict on test set
y_pred = pipeline.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories, zero_division="warn"))
```
**Use KNN Classifier**
```
#Use KNN Classifier
#create pipeline
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(KNeighborsClassifier()))
])
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
pipeline.fit(X_train, y_train)
# Predict on test set
y_pred = pipeline.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories, zero_division="warn"))
```
## 6.Improve Model
**Use GridSearchCV**
```
parameters = {
'vect__max_df': (0.5, 0.75, 1.0),
'vect__ngram_range': ((1, 1), (1,2)),
'vect__max_features': (None, 5000,10000),
'tfidf__use_idf': (True, False)
}
cv = GridSearchCV(estimator=pipeline, param_grid=parameters, cv=3, verbose=3)
cv.fit(X_train, y_train)
# Predict on test set
y_pred = pipeline.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories, zero_division="warn"))
```
## Save your model
```
import joblib
joblib.dump(cv, 'DS_model.pkl')
```
with the help of this notebook we can build ML Pipeline in `train_classifie.py` Script
## AdaBoostClassifier
```
from sklearn.neighbors import KNeighborsClassifier
#create pipeline
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
#('clf', MultiOutputClassifier(KNeighborsClassifier()))
#('clf', MultiOutputClassifier(GradientBoostingClassifier(n_estimators=100, learning_rate=1.0,max_depth=1, random_state=0)))
('clf', MultiOutputClassifier(AdaBoostClassifier(n_estimators=200, random_state=0)))
])
# Parameters for GridSearchCV
param_grid = {'vect__max_df': (0.75,1.0)}
# 'vect__ngram_range': ((1, 1), (1,2)),
# 'vect__max_features': (None, 5000),
# 'tfidf__use_idf': (True, False)
cv = GridSearchCV(pipeline, param_grid)
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
cv.fit(X_train, y_train)
# Predict on test set
y_pred = cv.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories))
```
## RandomForestClassifier
```
#create pipeline
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
#('clf', MultiOutputClassifier(KNeighborsClassifier()))
#('clf', MultiOutputClassifier(GradientBoostingClassifier(n_estimators=100, learning_rate=1.0,max_depth=1, random_state=0)))
#('clf', MultiOutputClassifier(AdaBoostClassifier(n_estimators=200, random_state=0)))
('clf', MultiOutputClassifier(RandomForestClassifier()))
])
# Parameters for GridSearchCV
param_grid = {'vect__max_df': (0.75,1.0)}
# 'vect__ngram_range': ((1, 1), (1,2)),
# 'vect__max_features': (None, 5000),
# 'tfidf__use_idf': (True, False)
cv = GridSearchCV(pipeline, param_grid)
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
cv.fit(X_train, y_train)
# Predict on test set
y_pred = cv.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories))
```
## SDGC Classifier
```
from sklearn.linear_model import SGDClassifier
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(SGDClassifier()))
])
# Parameters for GridSearchCV
param_grid = {'vect__max_df': (0.75,1.0)}
# 'vect__ngram_range': ((1, 1), (1,2)),
# 'vect__max_features': (None, 5000),
# 'tfidf__use_idf': (True, False)
cv = GridSearchCV(pipeline, param_grid)
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
cv.fit(X_train, y_train)
# Predict on test set
y_pred = cv.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories))
```
## GradientBoostingClassifier
```
#create pipeline
from sklearn.ensemble import GradientBoostingClassifier
pipeline = Pipeline([
('vect', CountVectorizer(tokenizer=tokenize)),
('tfidf', TfidfTransformer()),
('clf', MultiOutputClassifier(GradientBoostingClassifier(n_estimators=100, learning_rate=1.0,max_depth=1, random_state=0)))
])
# Parameters for GridSearchCV
param_grid = {'vect__max_df': (0.75,1.0)}
# 'vect__ngram_range': ((1, 1), (1,2)),
# 'vect__max_features': (None, 5000),
# 'tfidf__use_idf': (True, False)
cv = GridSearchCV(pipeline, param_grid)
#Split train and test data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.4, random_state=42)
# Fit pipeline
cv.fit(X_train, y_train)
# Predict on test set
y_pred = cv.predict(X_test)
categories = y.columns.tolist()
# Test model on test set
print(classification_report(y_test, y_pred, target_names=categories))
```
| github_jupyter |
# EDR Signatures
Author: Louis Richard\
A routine to compute various parameters used to identify electron diffusion regions for the four MMS spacecraft.
Quantities calculated so far are:
- sqrt(Q) : Based on Swisdak, GRL ,2016. Values around 0.1 indicate electron agyrotropies. Computed based on the off-diagonal terms in the pressure tensor for Pe_perp1 = Pe_perp2.
- Dng: Based on Aunai et al., 2013; Computed based on the off-diagonal terms in the pressure tensor for Pe_perp1 = Pe_perp2. Similar to sqrt(Q) but with different normalization. Calculated but not plotted.
- AG^(1/3): Based on Che et al., POP, 2018. Constructed from determinant of field-aligned rotation of the electron pressure tensor (Pe_perp1 = Pe_perp2).
- A phi_e/2 = abs(Perp1-Perp2)/(Perp1+Perp2): This is a measure of electron agyrotropy. Values of O(1) are expected for EDRs. We transform the pressure tensor into field-aligned coordinates such that the difference in Pe_perp1 and Pe_perp2 is maximal. This corresponds to P23 being zero. (Note that this definition of agyrotropy neglects the off-diagonal pressure terms P12 and P13, therefore it doesn't capture all agyrotropies.)
- A n_e = T_parallel/T_perp: Values much larger than 1 are expected. Large T_parallel/T_perp are a feature of the ion diffusion region. For MP reconnection ion diffusion regions have A n_e ~ 3 based on MMS observations. Scudder says A n_e ~ 7 at IDR-EDR boundary, but this is extremely large for MP reconnection.
- Mperp e: electron Mach number: bulk velocity divided by the electron thermal speed perpendicular to B. Values of O(1) are expected in EDRs (Scudder et al., 2012, 2015).
- J.E': J.E > 0 is expected in the electron diffusion region, corresponding to dissipation of field energy. J is calculated on each spacecraft using the particle moments (Zenitani et al., PRL, 2011).
- epsilon_e: Energy gain per cyclotron period. Values of O(1) are expected in EDRs (Scudder et al., 2012, 2015).
- delta_e: Relative strength of the electric and magnetic force in the bulk electron rest frame. N. B. Very sensitive to electron moments and electric field. Check version of these quantities (Scudder et al., 2012, 2015).
Notes:
kappa_e (not yet included) is taken to be the largest value of epsilon_e and delta_e at any given point. Requires electron distributions with version number v2.0.0 or higher. Calculations of agyrotropy measures (1)--(3) become unreliable at low densities n_e <~ 2 cm^-3, when the raw particle counts are low. Agyrotropies are removed for n_e < 1 cm^-3
```
import tqdm
import numpy as np
import matplotlib.pyplot as plt
from astropy import constants
from pyrfu.mms import get_data, rotate_tensor
from pyrfu.plot import make_labels, pl_tx
from pyrfu.pyrf import (resample, norm, cross, dot, trace, calc_sqrtq,
calc_dng, calc_ag, calc_agyro)
```
## Time interval selection
```
tint = ["2015-12-14T01:17:38.000", "2015-12-14T01:17:41.000"]
```
## Load Data
### Load fields
```
b_mms = [get_data("b_dmpa_fgm_srvy_l2", tint, i) for i in range(1, 5)]
e_mms = [get_data("e_dsl_edp_brst_l2", tint, i) for i in range(1, 5)]
```
### Load particles moments
```
n_mms_e = [get_data("ne_fpi_brst_l2", tint, i) for i in range(1, 5)]
v_mms_e = [get_data("ve_dbcs_fpi_brst_l2", tint, i) for i in range(1, 5)]
v_mms_i = [get_data("vi_dbcs_fpi_brst_l2", tint, i) for i in range(1, 5)]
t_mms_e = [get_data("te_dbcs_fpi_brst_l2", tint, i) for i in range(1, 5)]
p_mms_e = [get_data("pe_dbcs_fpi_brst_l2", tint, i) for i in range(1, 5)]
```
### Resample to DES sampling frequency
```
e_mms = [resample(e_xyz, n_e) for e_xyz, n_e in zip(e_mms, n_mms_e)]
b_mms = [resample(b_xyz, n_e) for b_xyz, n_e in zip(b_mms, n_mms_e)]
v_mms_i = [resample(v_xyz_i, n_e) for v_xyz_i, n_e in zip(v_mms_i, n_mms_e)]
```
## Rotate pressure and temperature tensors
```
p_mms_e_pp = [rotate_tensor(p_xyz, "fac", b_xyz, "pp") for p_xyz, b_xyz in zip(p_mms_e, b_mms)]
p_mms_e_qq = [rotate_tensor(p_xyz, "fac", b_xyz, "qq") for p_xyz, b_xyz in zip(p_mms_e, b_mms)]
t_mms_e_fac = [rotate_tensor(t_xyz, "fac", b_xyz) for t_xyz, b_xyz in zip(t_mms_e, b_mms)]
```
## Compute tests for EDR
### Compute Q and Dng from Pepp
```
sqrtq_mms = [calc_sqrtq(p_pp) for p_pp in p_mms_e_pp]
dng_mms = [calc_dng(p_pp) for p_pp in p_mms_e_pp]
```
### Compute agyrotropy measure AG1/3
```
ag_mms = [calc_ag(p_pp) for p_pp in p_mms_e_pp]
ag_cr_mms = [ag ** (1 / 3) for ag in ag_mms]
```
### Compute agyrotropy Aphi from Peqq
```
agyro_mms = [calc_agyro(p_qq) for p_qq in p_mms_e_qq]
```
### Simple fix to remove spurious points
```
for sqrtq, dng, agyro, ag_cr in zip(sqrtq_mms, dng_mms, agyro_mms, ag_cr_mms):
for coeff in [sqrtq, dng, agyro, ag_cr]:
coeff_data = coeff.data.copy()
for ii in tqdm.tqdm(np.arange(len(coeff_data) - 1)):
if coeff[ii] > 2 * coeff[ii - 1] and coeff[ii] > 2 * coeff[ii + 1]:
coeff_data[ii] = np.nan
coeff.data = coeff_data
```
### Remove all points corresponding to densities below 1cm^-3
```
for n_e, sqrtq, dng, agyro, ag_cr in zip(n_mms_e, sqrtq_mms, dng_mms, agyro_mms, ag_cr_mms):
sqrtq.data[n_e.data < 1] = np.nan
dng.data[n_e.data < 1] = np.nan
agyro.data[n_e.data < 1] = np.nan
ag_cr.data[n_e.data < 1] = np.nan
```
### Compute temperature ratio An
```
t_rat_mms = [p_pp[:, 0, 0] / p_pp[:, 1, 1] for p_pp in p_mms_e_pp]
```
### Compute electron Mach number
```
qe, me = [constants.e.value, constants.m_e.value]
v_mms_e_mag = [norm(v_xyz_e) for v_xyz_e in v_mms_e]
v_mms_e_per = [np.sqrt((t_fac_e[:, 1, 1] + t_fac_e[:, 2, 2]) * qe / me) for t_fac_e in t_mms_e_fac]
m_mms_e = [1e3 * v_e_mag / v_e_perp for v_e_mag, v_e_perp in zip(v_mms_e_mag, v_mms_e_per)]
```
### Compute current density and J.E
```
# Current density in nA m^-2
j_mms_moms = [1e18 * qe * n_e * (v_xyz_i - v_xyz_e) for n_e, v_xyz_i, v_xyz_e in zip(n_mms_e, v_mms_i, v_mms_e)]
vexb_mms = [e_xyz + 1e-3 * cross(v_xyz_e, b_xyz) for e_xyz, v_xyz_e, b_xyz in zip(e_mms, v_mms_e, b_mms)]
# J (nA/m^2), E (mV/m), E.J (nW/m^3)
edotj_mms = [1e-3 * dot(vexb_xyz, j_xyz) for vexb_xyz, j_xyz in zip(vexb_mms, j_mms_moms)]
```
### Calculate epsilon and delta parameters
```
w_mms_ce = [1e-9 * qe * norm(b_xyz) /me for b_xyz in b_mms]
edotve_mms = [dot(e_xyz, v_xyz_e) for e_xyz, v_xyz_e in zip(e_mms, v_mms_e)]
eps_mms_e = [np.abs(6 * np.pi * edotve_xyz /(w_ce * trace(t_fac_e))) for edotve_xyz, w_ce, t_fac_e in zip(edotve_mms, w_mms_ce, t_mms_e_fac)]
delta_mms_e = [1e-3 * norm(vexb_xyz) / (v_xyz_e_per * norm(b_xyz) * 1e-9) for vexb_xyz, v_xyz_e_per, b_xyz in zip(vexb_mms, v_mms_e_per, b_mms)]
```
## Plot figure
```
legend_options = dict(ncol=4, frameon=True, loc="upper right")
%matplotlib notebook
f, axs = plt.subplots(9, sharex="all", figsize=(6.5, 11))
f.subplots_adjust(bottom=.1, top=.95, left=.15, right=.85, hspace=0)
pl_tx(axs[0], b_mms, 2)
axs[0].set_ylabel("$B_{z}$ [nT]")
labels = ["MMS{:d}".format(ic) for ic in range(1, 5)]
axs[0].legend(labels, **legend_options)
pl_tx(axs[1], sqrtq_mms, 0)
axs[1].set_ylabel("$\sqrt{Q}$")
pl_tx(axs[2], ag_cr_mms, 0)
axs[2].set_ylabel("$AG^{1/3}$")
pl_tx(axs[3], agyro_mms, 0)
axs[3].set_ylabel("$A\Phi_e / 2$")
pl_tx(axs[4], t_rat_mms, 0)
axs[4].set_ylabel("$T_{e||}/T_{e \perp}$")
pl_tx(axs[5], m_mms_e, 0)
axs[5].set_ylabel("$M_{e \perp}$")
pl_tx(axs[6], edotj_mms, 0)
axs[6].set_ylabel("$E'.J$ [nW m$^{-3}$]")
pl_tx(axs[7], eps_mms_e, 0)
axs[7].set_ylabel("$\epsilon_{e}$")
pl_tx(axs[8], delta_mms_e, 0)
axs[8].set_ylabel("$\delta_{e}$")
make_labels(axs, [0.025, 0.83])
axs[-1].set_xlim(tint)
f.align_ylabels(axs)
```
| github_jupyter |
# Evaluation Dynamic Interpolation (DYMOST):
This notebook presents the evaluation of the SSH reconstructions based on the Dynamic Interpolation method ([Ballarotta et al., 2020](https://journals.ametsoc.org/view/journals/atot/37/9/jtechD200030.xml)) and performed for the **"2021a_SSH_mapping_OSE" ocean data challenge**.
```
import os
import sys
sys.path.append('..')
import logging
import pandas as pd
from src.mod_inout import *
from src.mod_interp import *
from src.mod_stats import *
from src.mod_spectral import *
from src.mod_plot import *
logger = logging.getLogger()
logger.setLevel(logging.INFO)
```
### Study Area & Ouput Parameters
```
# study area
lon_min = 295.
lon_max = 305.
lat_min = 33.
lat_max = 43.
is_circle = False
time_min = '2017-01-01'
time_max = '2017-12-31'
# Outputs
bin_lat_step = 1.
bin_lon_step = 1.
bin_time_step = '1D'
output_directory = '../results'
if not os.path.exists(output_directory):
os.mkdir(output_directory)
output_filename = f'{output_directory}/stat_OSE_DYMOST_{time_min}_{time_max}_{lon_min}_{lon_max}_{lat_min}_{lat_max}.nc'
output_filename_timeseries = f'{output_directory}/stat_timeseries_OSE_DYMOST_{time_min}_{time_max}_{lon_min}_{lon_max}_{lat_min}_{lat_max}.nc'
# Spectral parameter
# C2 parameter
delta_t = 0.9434 # s
velocity = 6.77 # km/s
delta_x = velocity * delta_t
lenght_scale = 1000 # km
output_filename_spectrum = f'{output_directory}/psd_OSE_DYMOST_{time_min}_{time_max}_{lon_min}_{lon_max}_{lat_min}_{lat_max}.nc'
```
### Open your AVISO+ session: fill the ```<AVISO_LOGIN>``` and ```<AVISO_PWD>``` items below
```
my_aviso_session = rq.Session()
my_aviso_session.auth = ("<AVISO_LOGIN>", "<AVISO_PWD>")
url_alongtrack = 'https://tds.aviso.altimetry.fr/thredds/dodsC/2021a-SSH-mapping-OSE-along-track-data'
url_map = 'https://tds.aviso.altimetry.fr/thredds/dodsC/2021a-SSH-mapping-OSE-grid-data'
```
### Read L3 datasets
```
# independent along-track
alontrack_independent_dataset = f'{url_alongtrack}/dt_gulfstream_c2_phy_l3_20161201-20180131_285-315_23-53.nc'
# Read along-track
ds_alongtrack = read_l3_dataset_from_aviso(alontrack_independent_dataset,
my_aviso_session,
lon_min=lon_min,
lon_max=lon_max,
lat_min=lat_min,
lat_max=lat_max,
time_min=time_min,
time_max=time_max)
ds_alongtrack
```
### Read L4 dataset and interpolate onto along-track positions
```
# series of maps to evaluate
gridded_dataset = [f'{url_map}/OSE_ssh_mapping_DYMOST.nc', my_aviso_session]
# Interpolate maps onto alongtrack dataset
time_alongtrack, lat_alongtrack, lon_alongtrack, ssh_alongtrack, ssh_map_interp = interp_on_alongtrack(gridded_dataset,
ds_alongtrack,
lon_min=lon_min,
lon_max=lon_max,
lat_min=lat_min,
lat_max=lat_max,
time_min=time_min,
time_max=time_max,
is_circle=is_circle)
```
### Compute statistical score
```
leaderboard_nrmse, leaderboard_nrmse_std = compute_stats(time_alongtrack,
lat_alongtrack,
lon_alongtrack,
ssh_alongtrack,
ssh_map_interp,
bin_lon_step,
bin_lat_step,
bin_time_step,
output_filename,
output_filename_timeseries)
plot_spatial_statistics(output_filename)
plot_temporal_statistics(output_filename_timeseries)
```
### Compute spectral scores
```
compute_spectral_scores(time_alongtrack,
lat_alongtrack,
lon_alongtrack,
ssh_alongtrack,
ssh_map_interp,
lenght_scale,
delta_x,
delta_t,
output_filename_spectrum)
leaderboard_psds_score = plot_psd_score(output_filename_spectrum)
# Print leaderboard
data = [['DYMOST',
leaderboard_nrmse,
leaderboard_nrmse_std,
int(leaderboard_psds_score),
'Dynamic mapping',
'example_eval_dymost.ipynb']]
Leaderboard = pd.DataFrame(data,
columns=['Method',
"µ(RMSE) ",
"σ(RMSE)",
'λx (km)',
'Notes',
'Reference'])
print("Summary of the leaderboard metrics:")
Leaderboard
print(Leaderboard.to_markdown())
```
| github_jupyter |
```
#@title
from google.colab import drive
drive.mount('/content/drive')
#@title
!cp -r '/content/drive/My Drive/Colab Notebooks/Melanoma/Scripts/.' .
COLAB_BASE_PATH = '/content/drive/My Drive/Colab Notebooks/Melanoma/'
MODEL_NAME = '81-efficientnetb6'
MODEL_BASE_PATH = f'{COLAB_BASE_PATH}Models/Files/{MODEL_NAME}/'
SUBMISSION_BASE_PATH = f'{COLAB_BASE_PATH}Submissions/'
SUBMISSION_PATH = SUBMISSION_BASE_PATH + f'{MODEL_NAME}.csv'
SUBMISSION_LAST_PATH = SUBMISSION_BASE_PATH + f'{MODEL_NAME}_last.csv'
SUBMISSION_BLEND_PATH = SUBMISSION_BASE_PATH + f'{MODEL_NAME}_blend.csv'
import os
os.mkdir(MODEL_BASE_PATH)
```
## Dependencies
```
#@title
!pip install --quiet efficientnet
# !pip install --quiet image-classifiers
#@title
import warnings, json, re, glob, math
from scripts_step_lr_schedulers import *
from melanoma_utility_scripts import *
from sklearn.model_selection import KFold
import tensorflow.keras.layers as L
import tensorflow.keras.backend as K
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint
from tensorflow.keras import optimizers, layers, metrics, losses, Model
import tensorflow_addons as tfa
import efficientnet.tfkeras as efn
# from classification_models.tfkeras import Classifiers
SEED = 42
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
## TPU configuration
```
#@title
strategy, tpu = set_up_strategy()
REPLICAS = strategy.num_replicas_in_sync
print("REPLICAS: ", REPLICAS)
AUTO = tf.data.experimental.AUTOTUNE
```
# Model parameters
```
#@title
config = {
"HEIGHT": 384,
"WIDTH": 384,
"CHANNELS": 3,
"BATCH_SIZE": 256,
"EPOCHS": 12,
"LEARNING_RATE": 0.00000125 * REPLICAS * 256,
"ES_PATIENCE": 5,
"N_FOLDS": 5,
"N_USED_FOLDS": 5,
"TTA_STEPS": 10,
"BASE_MODEL": 'EfficientNetB6',
"BASE_MODEL_WEIGHTS": 'imagenet',
"DATASET_PATH": 'melanoma-384x384'
}
with open(MODEL_BASE_PATH + 'config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
config
```
# Load data
```
database_base_path = COLAB_BASE_PATH + 'Data/'
k_fold = pd.read_csv(database_base_path + 'train.csv')
test = pd.read_csv(database_base_path + 'test.csv')
print('Train samples: %d' % len(k_fold))
display(k_fold.head())
print(f'Test samples: {len(test)}')
display(test.head())
GCS_PATH = 'gs://kds-3c0c447892078c6cb0cb069958214c9fedc65fe9efc33682fef72a3e'
TRAINING_FILENAMES = np.sort(tf.io.gfile.glob(GCS_PATH + '/train*.tfrec'))
TEST_FILENAMES = np.sort(tf.io.gfile.glob(GCS_PATH + '/test*.tfrec'))
```
# Augmentations
```
#@title
def data_augment(image, label):
p_spatial = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_spatial2 = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_rotation = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_crop = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_pixel = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
p_cutout = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
if p_spatial >= .2:
if p_spatial >= .6: # Flips
image['input_image'] = data_augment_spatial(image['input_image'])
else: # Rotate
image['input_image'] = data_augment_rotate(image['input_image'])
if p_crop >= .4: # Crops
image['input_image'] = data_augment_crop(image['input_image'])
if p_spatial2 >= .4:
if p_spatial2 >= .75: # Shift
image['input_image'] = data_augment_shift(image['input_image'])
else: # Shear
image['input_image'] = data_augment_shear(image['input_image'])
if p_pixel >= .3: # Pixel-level transforms
if p_pixel >= .8:
image['input_image'] = data_augment_hue(image['input_image'])
elif p_pixel >= .6:
image['input_image'] = data_augment_saturation(image['input_image'])
elif p_pixel >= .4:
image['input_image'] = data_augment_contrast(image['input_image'])
else:
image['input_image'] = data_augment_brightness(image['input_image'])
if p_rotation >= .4: # Rotation
image['input_image'] = data_augment_rotation(image['input_image'])
if p_cutout >= .4: # Cutout
image['input_image'] = data_augment_cutout(image['input_image'])
return image, label
def data_augment_rotation(image, max_angle=45.):
image = transform_rotation(image, config['HEIGHT'], max_angle)
return image
def data_augment_shift(image, h_shift=50., w_shift=50.):
image = transform_shift(image, config['HEIGHT'], h_shift, w_shift)
return image
def data_augment_shear(image, shear=25.):
image = transform_shear(image, config['HEIGHT'], shear)
return image
def data_augment_hue(image, max_delta=.02):
image = tf.image.random_hue(image, max_delta)
return image
def data_augment_saturation(image, lower=.8, upper=1.2):
image = tf.image.random_saturation(image, lower, upper)
return image
def data_augment_contrast(image, lower=.8, upper=1.2):
image = tf.image.random_contrast(image, lower, upper)
return image
def data_augment_brightness(image, max_delta=.1):
image = tf.image.random_brightness(image, max_delta)
return image
def data_augment_spatial(image):
p_spatial = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
image = tf.image.random_flip_left_right(image)
image = tf.image.random_flip_up_down(image)
if p_spatial > .75:
image = tf.image.transpose(image)
return image
def data_augment_rotate(image):
p_rotate = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
if p_rotate > .66:
image = tf.image.rot90(image, k=3) # rotate 270º
elif p_rotate > .33:
image = tf.image.rot90(image, k=2) # rotate 180º
else:
image = tf.image.rot90(image, k=1) # rotate 90º
return image
def data_augment_crop(image):
p_crop = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
if p_crop > .8:
image = tf.image.random_crop(image, size=[int(config['HEIGHT']*.7), int(config['WIDTH']*.7), config['CHANNELS']])
elif p_crop > .6:
image = tf.image.random_crop(image, size=[int(config['HEIGHT']*.8), int(config['WIDTH']*.8), config['CHANNELS']])
elif p_crop > .4:
image = tf.image.random_crop(image, size=[int(config['HEIGHT']*.9), int(config['WIDTH']*.9), config['CHANNELS']])
elif p_crop > .2:
image = tf.image.central_crop(image, central_fraction=.8)
else:
image = tf.image.central_crop(image, central_fraction=.7)
image = tf.image.resize(image, size=[config['HEIGHT'], config['WIDTH']])
return image
def data_augment_cutout(image, min_mask_size=(int(config['HEIGHT'] * .05), int(config['HEIGHT'] * .05)),
max_mask_size=(int(config['HEIGHT'] * .25), int(config['HEIGHT'] * .25))):
p_cutout = tf.random.uniform([1], minval=0, maxval=1, dtype='float32')
if p_cutout > .9: # 3 cut outs
image = random_cutout(image, config['HEIGHT'], config['WIDTH'],
min_mask_size=min_mask_size, max_mask_size=max_mask_size, k=3)
elif p_cutout > .75: # 2 cut outs
image = random_cutout(image, config['HEIGHT'], config['WIDTH'],
min_mask_size=min_mask_size, max_mask_size=max_mask_size, k=2)
else: # 1 cut out
image = random_cutout(image, config['HEIGHT'], config['WIDTH'],
min_mask_size=min_mask_size, max_mask_size=max_mask_size, k=1)
return image
```
## Auxiliary functions
```
#@title
# Datasets utility functions
def read_tfrecord(example, labeled=False, return_names=False, height=config['HEIGHT'], width=config['WIDTH'], channels=config['CHANNELS']):
if labeled:
example = tf.io.parse_single_example(example, LABELED_TFREC_FORMAT)
label = tf.cast(example['target'], tf.float32)
else:
example = tf.io.parse_single_example(example, UNLABELED_TFREC_FORMAT)
if return_names:
image_name = example['image_name']
image = decode_image(example['image'], height, width, channels)
# meta features
data = {}
data['patient_id'] = tf.cast(example['patient_id'], tf.int32)
data['sex'] = tf.cast(example['sex'], tf.int32)
data['age_approx'] = tf.cast(example['age_approx'], tf.int32)
data['anatom_site_general_challenge'] = tf.cast(tf.one_hot(example['anatom_site_general_challenge'], 7), tf.int32)
if labeled:
if return_names:
return {'input_image': image, 'input_meta': data}, label, image_name
else:
return {'input_image': image, 'input_meta': data}, label
else:
if return_names:
return {'input_image': image, 'input_meta': data}, image_name
else:
return {'input_image': image, 'input_meta': data}
def load_dataset(filenames, labeled=False, return_names=False, ordered=False, buffer_size=-1):
ignore_order = tf.data.Options()
if not ordered:
ignore_order.experimental_deterministic = False # disable order, increase speed
dataset = tf.data.TFRecordDataset(filenames, num_parallel_reads=buffer_size)
dataset = dataset.with_options(ignore_order) # uses data as soon as it streams in, rather than in its original order
dataset = dataset.map(lambda example: read_tfrecord(example, labeled, return_names), num_parallel_calls=buffer_size)
return dataset
def get_dataset(filenames, labeled=True, return_names=False, ordered=False, repeated=False, augment=False,
batch_size=32, buffer_size=-1):
dataset = load_dataset(filenames, labeled, return_names, ordered, buffer_size)
if augment:
dataset = dataset.map(data_augment, num_parallel_calls=buffer_size)
if repeated:
dataset = dataset.repeat()
if not ordered:
dataset = dataset.shuffle(2048)
dataset = dataset.batch(batch_size, drop_remainder=False)
dataset = dataset.prefetch(buffer_size)
return dataset
```
## Learning rate scheduler
```
#@title
lr_min = 1e-6
lr_start = 5e-6
lr_max = config['LEARNING_RATE']
steps_per_epoch = 24519 // config['BATCH_SIZE']
total_steps = config['EPOCHS'] * steps_per_epoch
warmup_steps = steps_per_epoch * 5
hold_max_steps = 0
step_decay = .8
step_size = steps_per_epoch * 1
rng = [i for i in range(0, total_steps, 32)]
y = [step_schedule_with_warmup(tf.cast(x, tf.float32), step_size=step_size,
warmup_steps=warmup_steps, hold_max_steps=hold_max_steps,
lr_start=lr_start, lr_max=lr_max, step_decay=step_decay) for x in rng]
sns.set(style="whitegrid")
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
#@title
# Initial bias
pos = len(k_fold[k_fold['target'] == 1])
neg = len(k_fold[k_fold['target'] == 0])
initial_bias = np.log([pos/neg])
print('Bias')
print(pos)
print(neg)
print(initial_bias)
# class weights
total = len(k_fold)
weight_for_0 = (1 / neg)*(total)/2.0
weight_for_1 = (1 / pos)*(total)/2.0
class_weight = {0: weight_for_0, 1: weight_for_1}
print('Class weight')
print(class_weight)
def model_fn(input_shape):
input_image = L.Input(shape=input_shape, name='input_image')
base_model = efn.EfficientNetB6(weights=config['BASE_MODEL_WEIGHTS'],
include_top=False)
x = base_model(input_image)
x = L.GlobalAveragePooling2D()(x)
output = L.Dense(1, activation='sigmoid', name='output',
bias_initializer=tf.keras.initializers.Constant(initial_bias))(x)
model = Model(inputs=input_image, outputs=output)
return model
```
# Training
```
NUM_TEST_IMAGES = count_data_items(TEST_FILENAMES)
NUM_TRAIN_IMAGES = count_data_items(TRAINING_FILENAMES)
# Evaluation
eval_dataset = get_dataset(TRAINING_FILENAMES, labeled=False, return_names=True, ordered=True, repeated=False, augment=False,
batch_size=1024, buffer_size=AUTO)
image_names = next(iter(eval_dataset.unbatch().map(lambda data, image_name: image_name).batch(NUM_TRAIN_IMAGES))).numpy().astype('U')
image_data = eval_dataset.map(lambda data, image_name: data)
# Test
test_preds = np.zeros((NUM_TEST_IMAGES, 1)); test_preds_last = np.zeros((NUM_TEST_IMAGES, 1))
test_dataset = get_dataset(TEST_FILENAMES, labeled=False, return_names=True, ordered=True, repeated=False, augment=True,
batch_size=1024, buffer_size=AUTO)
image_names_test = next(iter(test_dataset.unbatch().map(lambda data, image_name: image_name).batch(NUM_TEST_IMAGES))).numpy().astype('U')
test_image_data = test_dataset.map(lambda data, image_name: data)
# Resample dataframe
k_fold = k_fold[k_fold['image_name'].isin(image_names)]
k_fold_best = k_fold.copy()
history_list = []
kfold = KFold(config['N_FOLDS'], shuffle=True, random_state=SEED)
for n_fold, (trn_idx, val_idx) in enumerate(kfold.split(TRAINING_FILENAMES)):
if n_fold < config['N_USED_FOLDS']:
n_fold +=1
print('\nFOLD: %d' % (n_fold))
tf.tpu.experimental.initialize_tpu_system(tpu)
K.clear_session()
### Data
train_filenames = np.array(TRAINING_FILENAMES)[trn_idx]
valid_filenames = np.array(TRAINING_FILENAMES)[val_idx]
steps_per_epoch = count_data_items(train_filenames) // config['BATCH_SIZE']
# Train model
model_path = f'model_fold_{n_fold}.h5'
checkpoint = ModelCheckpoint((MODEL_BASE_PATH + model_path), monitor='val_loss', mode='min',
save_best_only=True, save_weights_only=True)
with strategy.scope():
model = model_fn((config['HEIGHT'], config['WIDTH'], config['CHANNELS']))
lr = lambda: step_schedule_with_warmup(tf.cast(optimizer.iterations, tf.float32),
step_size=step_size, warmup_steps=warmup_steps,
hold_max_steps=hold_max_steps, lr_start=lr_start,
lr_max=lr_max, step_decay=step_decay)
optimizer = optimizers.Adam(learning_rate=lr)
model.compile(optimizer, loss=losses.BinaryCrossentropy(label_smoothing=0.05),
metrics=[metrics.AUC()])
history = model.fit(get_dataset(train_filenames, labeled=True, return_names=False, ordered=False, repeated=True,
augment=True, batch_size=config['BATCH_SIZE'], buffer_size=AUTO),
validation_data=get_dataset(valid_filenames, labeled=True, return_names=False, ordered=True,
repeated=False, augment=False, batch_size=config['BATCH_SIZE'], buffer_size=AUTO),
epochs=config['EPOCHS'],
steps_per_epoch=steps_per_epoch,
callbacks=[checkpoint],
verbose=2).history
history_list.append(history)
# Save last epoch weights
model.save_weights((MODEL_BASE_PATH + 'last_' + model_path))
# Get validation IDs
valid_dataset = get_dataset(valid_filenames, labeled=False, return_names=True, ordered=True, repeated=False,
augment=False, batch_size=config['BATCH_SIZE'], buffer_size=AUTO)
valid_image_names = next(iter(valid_dataset.unbatch().map(lambda data, image_name: image_name).batch(count_data_items(valid_filenames)))).numpy().astype('U')
k_fold[f'fold_{n_fold}'] = k_fold.apply(lambda x: 'validation' if x['image_name'] in valid_image_names else 'train', axis=1)
k_fold_best[f'fold_{n_fold}'] = k_fold_best.apply(lambda x: 'validation' if x['image_name'] in valid_image_names else 'train', axis=1)
##### Last model #####
print(f'Last model evaluation...')
preds = model.predict(image_data)
name_preds_eval = dict(zip(image_names, preds.reshape(len(preds))))
k_fold[f'pred_fold_{n_fold}'] = k_fold.apply(lambda x: name_preds_eval[x['image_name']], axis=1)
print(f'Last model inference (TTA {config["TTA_STEPS"]} steps)...')
for step in range(config['TTA_STEPS']):
test_preds_last += model.predict(test_image_data)
##### Best model #####
print(f'Best model evaluation...')
model.load_weights(MODEL_BASE_PATH + model_path)
preds = model.predict(image_data)
name_preds_eval = dict(zip(image_names, preds.reshape(len(preds))))
k_fold_best[f'pred_fold_{n_fold}'] = k_fold_best.apply(lambda x: name_preds_eval[x['image_name']], axis=1)
print(f'Best model inference (TTA {config["TTA_STEPS"]} steps)...')
for step in range(config['TTA_STEPS']):
test_preds += model.predict(test_image_data)
# normalize preds
test_preds /= (config['N_USED_FOLDS'] * config['TTA_STEPS'])
test_preds_last /= (config['N_USED_FOLDS'] * config['TTA_STEPS'])
name_preds = dict(zip(image_names_test, test_preds.reshape(NUM_TEST_IMAGES)))
name_preds_last = dict(zip(image_names_test, test_preds_last.reshape(NUM_TEST_IMAGES)))
test['target'] = test.apply(lambda x: name_preds[x['image_name']], axis=1)
test['target_last'] = test.apply(lambda x: name_preds_last[x['image_name']], axis=1)
```
## Model loss graph
```
#@title
for n_fold in range(config['N_USED_FOLDS']):
print(f'Fold: {n_fold + 1}')
plot_metrics(history_list[n_fold])
```
## Model loss graph aggregated
```
#@title
# plot_metrics_agg(history_list, config['N_USED_FOLDS'])
```
# Model evaluation (best)
```
#@title
display(evaluate_model(k_fold_best, config['N_USED_FOLDS']).style.applymap(color_map))
display(evaluate_model_Subset(k_fold_best, config['N_USED_FOLDS']).style.applymap(color_map))
```
# Model evaluation (last)
```
#@title
display(evaluate_model(k_fold, config['N_USED_FOLDS']).style.applymap(color_map))
```
# Confusion matrix
```
#@title
for n_fold in range(config['N_USED_FOLDS']):
n_fold += 1
pred_col = f'pred_fold_{n_fold}'
train_set = k_fold_best[k_fold_best[f'fold_{n_fold}'] == 'train']
valid_set = k_fold_best[k_fold_best[f'fold_{n_fold}'] == 'validation']
print(f'Fold: {n_fold}')
plot_confusion_matrix(train_set['target'], np.round(train_set[pred_col]),
valid_set['target'], np.round(valid_set[pred_col]))
```
# Visualize predictions
```
#@title
k_fold['pred'] = 0
k_fold_best['pred'] = 0
for n_fold in range(config['N_USED_FOLDS']):
k_fold['pred'] += k_fold[f'pred_fold_{n_fold+1}'] / config['N_USED_FOLDS']
k_fold_best['pred'] += k_fold_best[f'pred_fold_{n_fold+1}'] / config['N_USED_FOLDS']
k_fold['pred_best'] = k_fold_best['pred']
print('Label/prediction distribution')
print(f"Train positive labels: {len(k_fold[k_fold['target'] > .5])}")
print(f"Train positive predictions: {len(k_fold[k_fold['pred'] > .5])}")
print(f"Train positive correct predictions: {len(k_fold[(k_fold['target'] > .5) & (k_fold['pred'] > .5)])}")
print('Top 5 samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred', 'pred_best'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].head(5))
print('Top 5 positive samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred', 'pred_best'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].query('target == 1').head(5))
print('Top 5 predicted positive samples')
display(k_fold[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'diagnosis',
'target', 'pred', 'pred_best'] + [c for c in k_fold.columns if (c.startswith('pred_fold'))]].query('pred > .5').head(5))
display(k_fold.describe())
fig, (ax1, ax2) = plt.subplots(1, 2, sharex=True, figsize=(20, 6))
ax1.hist(k_fold['pred'], bins=100)
ax2.hist(k_fold['pred_best'], bins=100)
ax1.set_title('Last')
ax2.set_title('Best')
plt.show()
```
# Visualize test predictions
```
#@title
print(f"Test predictions {len(test[test['target'] > .5])}|{len(test[test['target'] <= .5])}")
print(f"Test predictions (last) {len(test[test['target_last'] > .5])}|{len(test[test['target_last'] <= .5])}")
print('Top 10 samples')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last']
+ [c for c in test.columns if (c.startswith('pred_fold'))]].head(10))
print('Top 10 positive samples')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last']
+ [c for c in test.columns if (c.startswith('pred_fold'))]].query('target > .5').head(10))
print('Top 10 positive samples (last)')
display(test[['image_name', 'sex', 'age_approx','anatom_site_general_challenge', 'target', 'target_last']
+ [c for c in test.columns if (c.startswith('pred_fold'))]].query('target_last > .5').head(10))
fig = plt.subplots(figsize=(20, 5))
plt.hist(test['target'], bins=100)
plt.show()
```
# Test set predictions
```
#@title
submission = pd.read_csv(database_base_path + 'sample_submission.csv')
submission['target'] = test['target']
submission['target_last'] = test['target_last']
submission['target_blend'] = (test['target'] * .5) + (test['target_last'] * .5)
display(submission.head(10))
display(submission.describe())
### BEST ###
submission[['image_name', 'target']].to_csv(SUBMISSION_PATH, index=False)
### LAST ###
submission_last = submission[['image_name', 'target_last']]
submission_last.columns = ['image_name', 'target']
submission_last.to_csv(SUBMISSION_LAST_PATH, index=False)
### BLEND ###
submission_blend = submission[['image_name', 'target_blend']]
submission_blend.columns = ['image_name', 'target']
submission_blend.to_csv(SUBMISSION_BLEND_PATH, index=False)
```
| github_jupyter |
<center>
<img src="../../img/ods_stickers.jpg">
## Open Machine Learning Course
Authors: [Maria Sumarokova](https://www.linkedin.com/in/mariya-sumarokova-230b4054/), senior data scientist/analyst at Veon, and [Yury Kashnitsky](https://www.linkedin.com/in/festline/), data scientist at Mail.Ru Group. Translated and edited by Gleb Filatov, Aleksey Kiselev, [Anastasia Manokhina](https://www.linkedin.com/in/anastasiamanokhina/), [Egor Polusmak](https://www.linkedin.com/in/egor-polusmak/), and [Yuanyuan Pao](https://www.linkedin.com/in/yuanyuanpao/). All content is distributed under the [Creative Commons CC BY-NC-SA 4.0](https://creativecommons.org/licenses/by-nc-sa/4.0/) license.
# <center> Assignment #3 (demo)
## <center> Decision trees with a toy task and the UCI Adult dataset
Please fill in the answers in the [web-form](https://docs.google.com/forms/d/1wfWYYoqXTkZNOPy1wpewACXaj2MZjBdLOL58htGWYBA/edit).
Let's start by loading all necessary libraries:
```
%matplotlib inline
from matplotlib import pyplot as plt
plt.rcParams['figure.figsize'] = (10, 8)
import seaborn as sns
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder
import collections
from sklearn.model_selection import GridSearchCV
from sklearn import preprocessing
from sklearn.tree import DecisionTreeClassifier, export_graphviz
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
from ipywidgets import Image
from io import StringIO
import pydotplus #pip install pydotplus
```
### Part 1. Toy dataset "Will They? Won't They?"
Your goal is to figure out how decision trees work by walking through a toy problem. While a single decision tree does not yield outstanding results, other performant algorithms like gradient boosting and random forests are based on the same idea. That is why knowing how decision trees work might be useful.
We'll go through a toy example of binary classification - Person A is deciding whether they will go on a second date with Person B. It will depend on their looks, eloquence, alcohol consumption (only for example), and how much money was spent on the first date.
#### Creating the dataset
```
# Create dataframe with dummy variables
def create_df(dic, feature_list):
out = pd.DataFrame(dic)
out = pd.concat([out, pd.get_dummies(out[feature_list])], axis = 1)
out.drop(feature_list, axis = 1, inplace = True)
return out
# Some feature values are present in train and absent in test and vice-versa.
def intersect_features(train, test):
common_feat = list( set(train.keys()) & set(test.keys()))
return train[common_feat], test[common_feat]
features = ['Looks', 'Alcoholic_beverage','Eloquence','Money_spent']
```
#### Training data
```
df_train = {}
df_train['Looks'] = ['handsome', 'handsome', 'handsome', 'repulsive',
'repulsive', 'repulsive', 'handsome']
df_train['Alcoholic_beverage'] = ['yes', 'yes', 'no', 'no', 'yes', 'yes', 'yes']
df_train['Eloquence'] = ['high', 'low', 'average', 'average', 'low',
'high', 'average']
df_train['Money_spent'] = ['lots', 'little', 'lots', 'little', 'lots',
'lots', 'lots']
df_train['Will_go'] = LabelEncoder().fit_transform(['+', '-', '+', '-', '-', '+', '+'])
df_train = create_df(df_train, features)
df_train
```
#### Test data
```
df_test = {}
df_test['Looks'] = ['handsome', 'handsome', 'repulsive']
df_test['Alcoholic_beverage'] = ['no', 'yes', 'yes']
df_test['Eloquence'] = ['average', 'high', 'average']
df_test['Money_spent'] = ['lots', 'little', 'lots']
df_test = create_df(df_test, features)
df_test
# Some feature values are present in train and absent in test and vice-versa.
y = df_train['Will_go']
df_train, df_test = intersect_features(train=df_train, test=df_test)
df_train
df_test
```
#### Draw a decision tree (by hand or in any graphics editor) for this dataset. Optionally you can also implement tree construction and draw it here.
1\. What is the entropy $S_0$ of the initial system? By system states, we mean values of the binary feature "Will_go" - 0 or 1 - two states in total.
```
n = len(y)
counts = y.value_counts()
p1 = counts[1] / n
p0 = counts[0] / n
-1 * (p1 * np.log2(p1) + p0 * np.log2(p0))
```
2\. Let's split the data by the feature "Looks_handsome". What is the entropy $S_1$ of the left group - the one with "Looks_handsome". What is the entropy $S_2$ in the opposite group? What is the information gain (IG) if we consider such a split?
```
# you code here
```
#### Train a decision tree using sklearn on the training data. You may choose any depth for the tree.
```
tree = DecisionTreeClassifier()
tree.fit(df_train, y)
```
#### Additional: display the resulting tree using graphviz. You can use pydot or [web-service](https://www.coolutils.com/ru/online/DOT-to-PNG) dot2png.
```
#tree.predict(df_test)
tree.predict_proba(df_test)
dot_tree = export_graphviz(tree)
graph = pydotplus.graphviz.graph_from_dot_file("tree.dot")
graph.write_png('tree.png')
```
### Part 2. Functions for calculating entropy and information gain.
Consider the following warm-up example: we have 9 blue balls and 11 yellow balls. Let ball have label **1** if it is blue, **0** otherwise.
```
balls = [1 for i in range(9)] + [0 for i in range(11)]
```
<img src = '../../img/decision_tree3.png'>
Next split the balls into two groups:
<img src = '../../img/decision_tree4.png'>
```
# two groups
balls_left = [1 for i in range(8)] + [0 for i in range(5)] # 8 blue and 5 yellow
balls_right = [1 for i in range(1)] + [0 for i in range(6)] # 1 blue and 6 yellow
```
#### Implement a function to calculate the Shannon Entropy
```
def entropy(a_list):
seen = []
entropy = 0
for el in a_list:
if el not in seen:
seen.append(el)
p = a_list.count(el) / len(a_list)
entropy += -p * np.log2(p)
return entropy
```
Tests
```
print(entropy(balls)) # 9 blue и 11 yellow
print(entropy(balls_left)) # 8 blue и 5 yellow
print(entropy(balls_right)) # 1 blue и 6 yellow
print(entropy([1,2,3,4,5,6])) # entropy of a fair 6-sided die
```
3\. What is the entropy of the state given by the list **balls_left**?
```
print(entropy(balls_left))
```
4\. What is the entropy of a fair dice? (where we look at a dice as a system with 6 equally probable states)?
```
# information gain calculation
def information_gain(root, left, right):
''' root - initial data, left and right - two partitions of initial data'''
# you code here
pass
```
5\. What is the information gain from splitting the initial dataset into **balls_left** and **balls_right** ?
```
def best_feature_to_split(X, y):
'''Outputs information gain when splitting on best feature'''
# you code here
pass
```
#### Optional:
- Implement a decision tree building algorithm by calling **best_feature_to_split** recursively
- Plot the resulting tree
### Part 3. The "Adult" dataset
#### Dataset description:
[Dataset](http://archive.ics.uci.edu/ml/machine-learning-databases/adult) UCI Adult (no need to download it, we have a copy in the course repository): classify people using demographical data - whether they earn more than \$50,000 per year or not.
Feature descriptions:
- **Age** – continuous feature
- **Workclass** – continuous feature
- **fnlwgt** – final weight of object, continuous feature
- **Education** – categorical feature
- **Education_Num** – number of years of education, continuous feature
- **Martial_Status** – categorical feature
- **Occupation** – categorical feature
- **Relationship** – categorical feature
- **Race** – categorical feature
- **Sex** – categorical feature
- **Capital_Gain** – continuous feature
- **Capital_Loss** – continuous feature
- **Hours_per_week** – continuous feature
- **Country** – categorical feature
**Target** – earnings level, categorical (binary) feature.
#### Reading train and test data
```
data_train = pd.read_csv('../../data/adult_train.csv', sep=';')
data_train.tail()
data_test = pd.read_csv('../../data/adult_test.csv', sep=';')
data_test.tail()
# necessary to remove rows with incorrect labels in test dataset
data_test = data_test[(data_test['Target'] == ' >50K.') | (data_test['Target']==' <=50K.')]
# encode target variable as integer
data_train.loc[data_train['Target']==' <=50K', 'Target'] = 0
data_train.loc[data_train['Target']==' >50K', 'Target'] = 1
data_test.loc[data_test['Target']==' <=50K.', 'Target'] = 0
data_test.loc[data_test['Target']==' >50K.', 'Target'] = 1
```
#### Primary data analysis
```
data_test.describe(include='all').T
data_train['Target'].value_counts()
fig = plt.figure(figsize=(25, 15))
cols = 5
rows = np.ceil(float(data_train.shape[1]) / cols)
for i, column in enumerate(data_train.columns):
ax = fig.add_subplot(rows, cols, i + 1)
ax.set_title(column)
if data_train.dtypes[column] == np.object:
data_train[column].value_counts().plot(kind="bar", axes=ax)
else:
data_train[column].hist(axes=ax)
plt.xticks(rotation="vertical")
plt.subplots_adjust(hspace=0.7, wspace=0.2)
```
#### Checking data types
```
data_train.dtypes
data_test.dtypes
```
As we see, in the test data, age is treated as type **object**. We need to fix this.
```
data_test['Age'] = data_test['Age'].astype(int)
```
Also we'll cast all **float** features to **int** type to keep types consistent between our train and test data.
```
data_test['fnlwgt'] = data_test['fnlwgt'].astype(int)
data_test['Education_Num'] = data_test['Education_Num'].astype(int)
data_test['Capital_Gain'] = data_test['Capital_Gain'].astype(int)
data_test['Capital_Loss'] = data_test['Capital_Loss'].astype(int)
data_test['Hours_per_week'] = data_test['Hours_per_week'].astype(int)
```
#### Fill in missing data for continuous features with their median values, for categorical features with their mode.
```
# choose categorical and continuous features from data
categorical_columns = [c for c in data_train.columns
if data_train[c].dtype.name == 'object']
numerical_columns = [c for c in data_train.columns
if data_train[c].dtype.name != 'object']
print('categorical_columns:', categorical_columns)
print('numerical_columns:', numerical_columns)
# fill missing data
for c in categorical_columns:
data_train[c].fillna(data_train[c].mode(), inplace=True)
data_test[c].fillna(data_train[c].mode(), inplace=True)
for c in numerical_columns:
data_train[c].fillna(data_train[c].median(), inplace=True)
data_test[c].fillna(data_train[c].median(), inplace=True)
```
We'll dummy code some categorical features: **Workclass**, **Education**, **Martial_Status**, **Occupation**, **Relationship**, **Race**, **Sex**, **Country**. It can be done via pandas method **get_dummies**
```
data_train = pd.concat([data_train[numerical_columns],
pd.get_dummies(data_train[categorical_columns])], axis=1)
data_test = pd.concat([data_test[numerical_columns],
pd.get_dummies(data_test[categorical_columns])], axis=1)
set(data_train.columns) - set(data_test.columns)
data_train.shape, data_test.shape
```
#### There is no Holland in the test data. Create new zero-valued feature.
```
data_test['Country_ Holand-Netherlands'] = 0
set(data_train.columns) - set(data_test.columns)
data_train.head(2)
data_test.head(2)
X_train = data_train.drop(['Target'], axis=1)
y_train = data_train['Target']
X_test = data_test.drop(['Target'], axis=1)
y_test = data_test['Target']
```
### 3.1 Decision tree without parameter tuning
Train a decision tree **(DecisionTreeClassifier)** with a maximum depth of 3, and evaluate the accuracy metric on the test data. Use parameter **random_state = 17** for results reproducibility.
```
# you code here
# tree =
# tree.fit
```
Make a prediction with the trained model on the test data.
```
# you code here
# tree_predictions = tree.predict
# you code here
# accuracy_score
```
6\. What is the test set accuracy of a decision tree with maximum tree depth of 3 and **random_state = 17**?
### 3.2 Decision tree with parameter tuning
Train a decision tree **(DecisionTreeClassifier, random_state = 17).** Find the optimal maximum depth using 5-fold cross-validation **(GridSearchCV)**.
```
tree_params = {'max_depth': range(2,11)}
locally_best_tree = GridSearchCV # you code here
locally_best_tree.fit; # you code here
```
Train a decision tree with maximum depth of 9 (it is the best **max_depth** in my case), and compute the test set accuracy. Use parameter **random_state = 17** for reproducibility.
```
# you code here
# tuned_tree =
# tuned_tree.fit
# tuned_tree_predictions = tuned_tree.predict
# accuracy_score
```
7\. What is the test set accuracy of a decision tree with maximum depth of 9 and **random_state = 17**?
### 3.3 (Optional) Random forest without parameter tuning
Let's take a sneak peek of upcoming lectures and try to use a random forest for our task. For now, you can imagine a random forest as a bunch of decision trees, trained on slightly different subsets of the training data.
Train a random forest **(RandomForestClassifier)**. Set the number of trees to 100 and use **random_state = 17**.
```
# you code here
# rf =
# rf.fit # you code here
```
Make predictions for the test data and assess accuracy.
```
# you code here
```
### 3.4 (Optional) Random forest with parameter tuning
Train a random forest **(RandomForestClassifier)**. Tune the maximum depth and maximum number of features for each tree using **GridSearchCV**.
```
# forest_params = {'max_depth': range(10, 21),
# 'max_features': range(5, 105, 20)}
# locally_best_forest = GridSearchCV # you code here
# locally_best_forest.fit # you code here
```
Make predictions for the test data and assess accuracy.
```
# you code here
```
| github_jupyter |
##### Copyright 2021 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# SavedModel Migration
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/migrate/saved_model">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/migrate/saved_model.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/migrate/saved_model.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/migrate/saved_model.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
Once you have migrated your model from graphs and sessions to `tf.function`, `tf.Module`, and `tf.keras.Model`, it is time to migrate the saving and loading code. This is specifically a guide about migrating from TF1 to TF2, [see here a more general guide about the TF2 SavedModel API](../../guide/saved_model.ipynb).
This is a quick overview of the API changes from TF1 to TF2.
||TF1|Migration to TF2|
| ----| ----|---|
|**Saving**|`tf.compat.v1.saved_model.Builder`<br>`tf.compat.v1.saved_model.simple_save`|`tf.saved_model.save`<br>`tf.keras.models.save_model`
|**Loading**|`tf.compat.v1.saved_model.load`|`tf.saved_model.load`
|**Signatures**: a set of input <br>and output tensors that <br>can be used to run the<br> model|These are generated using the `*.signature_def` utils<br>(e.g. `tf.compat.v1.saved_model.predict_signature_def`)|Write a `tf.function`, and export it using the<br> `signatures` argument in <br>`tf.saved_model.save`.
**Classify and regress**:<br>special types of signatures|Generated with `classification_signature_def`, <br> `regresssion_signature_def`, and certain Estimator exports.|These two signatures types have been removed <br>from TF2.If the serving library requires these <br>method names, use the [`MethodNameUpdater`](https://www.tensorflow.org/api_docs/python/tf/compat/v1/saved_model/signature_def_utils/MethodNameUpdater).
For a more in-depth explanation of the mapping see the *Changes from TF1 to TF2* section
Below are code examples of writing saving and loading code in TF1 and TF2.
## Code Sample Set up
The code examples below show how to export and load the same dummy TensorFlow model (`add_two`) to SavedModel using the TF1 and TF2 APIs.
Run the below code to set up the imports and utils functions
```
import tensorflow as tf
import tensorflow.compat.v1 as tf1
import shutil
def remove_dir(path):
try:
shutil.rmtree(path)
except:
pass
def add_two(input):
return input + 2
```
## Building TF1 SavedModels
The TF1 APIs, `tf.compat.v1.saved_model.Builder`, `tf.compat.v1.saved_model.simple_save`, and `tf.estimator.Estimator.export_saved_model` export the TensorFlow graph and session.
### Builder
```
remove_dir("saved-model-builder")
with tf.Graph().as_default() as g:
with tf1.Session() as sess:
input = tf1.placeholder(tf.float32, shape=[])
output = add_two(input)
print("add two output: ", sess.run(output, {input: 3.}))
# Save with SavedModelBuilder
builder = tf1.saved_model.Builder('saved-model-builder')
sig_def = tf1.saved_model.predict_signature_def(
inputs={'input': input},
outputs={'output': output})
builder.add_meta_graph_and_variables(
sess, tags=["serve"], signature_def_map={
tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY: sig_def
})
builder.save()
!saved_model_cli run --dir simple-save --tag_set serve \
--signature_def serving_default --input_exprs input=10
```
### Simple Save
```
remove_dir("simple-save")
with tf.Graph().as_default() as g:
with tf1.Session() as sess:
input = tf1.placeholder(tf.float32, shape=[])
output = add_two(input)
print("add two output: ", sess.run(output, {input: 3.}))
tf1.saved_model.simple_save(
sess, 'simple-save',
inputs={'input': input},
outputs={'output': output})
!saved_model_cli run --dir simple-save --tag_set serve \
--signature_def serving_default --input_exprs input=10
```
### Estimator export
In the definition of the Estimator `model_fn`, you can define signatures in your model by returning `export_outputs` in the `EstimatorSpec`. There are different types of outputs:
* `tf.estimator.export.ClassificationOutput`
* `tf.estimator.export.RegressionOutput`
* `tf.estimator.export.PredictOutput`
These will produce `classify`, `regress`, and `predict` signature types.
When the estimator is exported with `tf.estimator.Estimator.export_saved_model`, these signatures will be saved with the model.
```
def model_fn(features, labels, mode):
output = add_two(features['input'])
step = tf1.train.get_global_step()
return tf.estimator.EstimatorSpec(
mode,
predictions=output,
train_op=step.assign_add(1),
loss=tf.constant(0.),
export_outputs={
tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY: \
tf.estimator.export.PredictOutput({'output': output})})
est = tf.estimator.Estimator(model_fn, 'estimator-checkpoints')
# Train for one step to create a checkpoint
def train_fn():
return tf.data.Dataset.from_tensors({'input': 3.})
est.train(train_fn, steps=1)
# This util function `build_raw_serving...` takes in raw tensor features
# and builds an "input serving receiver function", which creates placeholder
# inputs to the model.
serving_input_fn = tf.estimator.export.build_raw_serving_input_receiver_fn(
{'input': tf.constant(3.)}) # Pass in a dummy input batch
estimator_path = est.export_saved_model('exported-estimator', serving_input_fn)
# Estimator's export_saved_model creates a timestamped directory. Move this
# to a set path so it can be inspected with saved_model_cli in the cell below:
!rm -rf estimator-model
import shutil
shutil.move(estimator_path, 'estimator-model')
!saved_model_cli run --dir estimator-model --tag_set serve \
--signature_def serving_default --input_exprs input=[10]
```
## Building TF2 SavedModels
### TF2 Saving
To export your model in TF2, you must define a `tf.Module` or `tf.keras.Model` to hold all of your model's variables and functions. Then, call `tf.saved_model.save` to create a SavedModel.
```
class MyModel(tf.Module):
@tf.function
def __call__(self, input):
return add_two(input)
model = MyModel()
@tf.function
def serving_default(input):
return {'output': model(input)}
signature_function = serving_default.get_concrete_function(
tf.TensorSpec(shape=[], dtype=tf.float32))
tf.saved_model.save(
model, 'tf2-save', signatures={
tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY: signature_function})
!saved_model_cli run --dir tf2-save --tag_set serve \
--signature_def serving_default --input_exprs input=10
```
### Keras SavedModel (TF2)
The Keras `tf` format exports a SavedModel from a `tf.keras.Model`. SavedModels exported with Keras can be reloaded using any of the loading APIs, in addition to the Keras loading function.
```
inp = tf.keras.Input(3)
out = add_two(inp)
model = tf.keras.Model(inputs=inp, outputs=out)
@tf.function(input_signature=[tf.TensorSpec(shape=[], dtype=tf.float32)])
def serving_default(input):
return {'output': model(input)}
model.save('keras-model', save_format='tf', signatures={
tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY: serving_default})
!saved_model_cli run --dir keras-model --tag_set serve \
--signature_def serving_default --input_exprs input=10
```
## Loading SavedModels
SavedModels saved with any of the above APIs can be loaded with either TF1 or TF2.
TF1 SavedModel can generally be used for inference when loaded into TF2, but training (generating gradients) is only possible if the SavedModel contains *resource variables*. You can check the dtype of the variables. If the variable dtype contains "_ref" then it is a reference variable.
TF2 SavedModel can be loaded and executed from TF1 as long as the SavedModel is saved with signatures.
The sections below contain code samples showing how to load the SavedModels saved in the previous sections, and call the exported signature.
### TF1 Loading
TF1 imports the SavedModel directly in to the current graph and session. You can call `session.run` on the tensor input and output names.
```
def load_tf1(path, input):
print('Loading from', path)
with tf.Graph().as_default() as g:
with tf1.Session() as sess:
meta_graph = tf1.saved_model.load(sess, ["serve"], path)
sig_def = meta_graph.signature_def[tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY]
input_name = sig_def.inputs['input'].name
output_name = sig_def.outputs['output'].name
print(' Output with input', input, ': ',
sess.run(output_name, feed_dict={input_name: input}))
load_tf1('saved-model-builder', 5.)
load_tf1('simple-save', 5.)
load_tf1('estimator-model', [5.]) # Estimator's input must be batched.
load_tf1('tf2-save', 5.)
load_tf1('keras-model', 5.)
```
### TF2 Loading
In TF2, objects are loaded into a Python object that stores the variables and functions. This is compatible with models saved from TF1. See `tf.saved_model.load` for details.
```
def load_tf2(path, input):
print('Loading from', path)
loaded = tf.saved_model.load(path)
out = loaded.signatures[tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY](
tf.constant(input))['output']
print(' Output with input', input, ': ', out)
load_tf2('saved-model-builder', 5.)
load_tf2('simple-save', 5.)
load_tf2('estimator-model', [5.]) # Estimator's input must be batched.
load_tf2('tf2-save', 5.)
load_tf2('keras-model', 5.)
```
Models saved with the TF2 API can also access tf.functions and variables that are attached to the model (instead of those exported as signatures). Example:
```
loaded = tf.saved_model.load('tf2-save')
print('restored __call__:', loaded.__call__)
print('output with input 5.', loaded(5))
```
### Keras loading
The Keras loading API allows you to reload a saved model back into a Keras Model object. Note that this only allows you to load SavedModels saved with Keras (`model.save` or `tf.keras.models.save_model`). Models saved with `tf.saved_model.save` can not be loaded.
Load the previously saved Keras model:
```
loaded_model = tf.keras.models.load_model('keras-model')
loaded_model.predict_on_batch(tf.constant([1, 3, 4]))
```
## Changes from TF1 to TF2
This section lists out key terms from TF1, their TF2 equivalents, and what has changed.
### SavedModel
SavedModel is a format that stores a TensorFlow model. It contains signatures which are used by serving platforms to run the model.
The file format itself has not changed significantly, so SavedModels can be loaded and served using either TF1 and TF2 APIs.
**Differences between TF1 and TF2**
The *serving* and *inference* use cases have not been updated, aside from API changes. The improvement in TF2 is the ability to *reuse* and *compose models* loaded from SavedModel.
In TF2, the program is represented by objects like `tf.Variable`, `tf.Module`, or Keras models and layers. There are no more global variables that have values stored in a session, and the graph now exists in different `tf.functions`. Consequently, during export, SavedModel saves each component and function graphs separately.
When you write a TensorFlow program with the TF2 Python API, you must build an object to manage the variables, functions, and other resources. Generally, this is accomplished by using the Keras API, but you can also build the object by creating or subclassing `tf.Module`.
Keras models and `tf.Module` automatically track variables and functions attached to them. SavedModel saves these connections between modules, variables and functions so that they can be restored when loading.
### Signatures
Signatures are the endpoints of a SavedModel -- they tell the user how to run the model and what inputs are needed.
In TF1, signatures are created by listing the input and output tensors. In TF2, signatures are generated by passing in *concrete functions*.
To read more about TensorFlow functions, [see this guide](../guide/intro_to_graphs). In short, a concrete function is generated from a `tf.function`:
```
# Option 1: specify an input signature
@tf.function(input_signature=[...])
def fn(...):
...
return outputs
tf.saved_model.save(model, path, signatures={
'name': fn
})
```
```
# Option 2: call get_concrete_function
@tf.function
def fn(...):
...
return outputs
tf.saved_model.save(model, path, signatures={
'name': fn.get_concrete_function(...)
})
```
### `session.run`
In TF1, you could call `session.run` with the imported graph as long as you already know the tensor names. This allows you to retrieve the restored variable values, or run parts of the model that were not exported in the signatures.
In TF2, you can directly access the variable (e.g. `loaded.dense_layer.kernel`), or call `tf.function`s attached the model object (e.g. `loaded.__call__`).
Unlike TF1, there is no way to extract parts of a function and access intermediate values. You *must* export all of the needed functionality in the saved object.
## Serving Migration notes
SavedModel was originally created to work with [TensorFlow Serving](https://www.tensorflow.org/tfx/guide/serving). This platform offers different types of prediction requests: classify, regress, and predict.
The TF1 API allows you to create these types of signatures with the utils:
* `tf.compat.v1.saved_model.classification_signature_def`
* `tf.compat.v1.saved_model.regression_signature_def`
* `tf.compat.v1.saved_model.predict_signature_def`
Classify and regress restrict the inputs and outputs, so the inputs must be a `tf.Example`, and the outputs must be `classes` or `scores` or `prediction`. Meanwhile, the predict signature has no restrictions.
SavedModels exported with the **TF2** API are compatible with TensorFlow serving, but will only contain `predict` signatures. The `classify` and `regress` signatures have ben removed.
If you require the use of the `classify` and `regress` signatures, you may modify the exported SavedModel using `tf.compat.v1.saved_model.signature_def_utils.MethodNameUpdater`.
## Other Saving Migration guides
If you are using TF Hub, then here are some guides that you may fine useful:
* https://www.tensorflow.org/hub/model_compatibility
* https://www.tensorflow.org/hub/migration_tf2
| github_jupyter |
##### Copyright 2021 Google LLC.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<a href="https://colab.research.google.com/github/google-research/vision_transformer/blob/master/vit_jax_augreg.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## How to train your ViT? Data, Augmentation, and Regularization in Vision Transformers
Model repository published with the paper
[**How to train your ViT? Data, Augmentation, and Regularization in Vision
Transformers**](https://arxiv.org/abs/TODO)
This Colab shows how to
[find checkpoints](#scrollTo=F4SLGDtFxlsC)
in the repository, how to
[select and load a model](#scrollTo=wh_SLkQtQ6K4)
form the repository and use it for inference
([also with PyTorch](#scrollTo=1nMyWmDycpAo)),
and how to
[fine-tune on a dataset](#scrollTo=iAruT3YOxqB6).
For more details, please refer to the repository:
https://github.com/google-research/vision_transformer/
Note that this Colab directly uses the unmodified code from the repository. If
you want to modify the modules and persist your changes, you can do all that
using free GPUs and TPUs without leaving the Colab environment - see
https://colab.research.google.com/github/google-research/vision_transformer/blob/master/vit_jax.ipynb
### Imports
```
# Fetch vision_transformer repository.
![ -d vision_transformer ] || git clone --depth=1 https://github.com/google-research/vision_transformer
# Install dependencies.
!pip install -qr vision_transformer/vit_jax/requirements.txt
# Import files from repository.
import sys
if './vision_transformer' not in sys.path:
sys.path.append('./vision_transformer')
%load_ext autoreload
%autoreload 2
from vit_jax import checkpoint
from vit_jax import models
from vit_jax import train
from vit_jax.configs import augreg as augreg_config
from vit_jax.configs import models as models_config
# Connect to TPUs if runtime type is of type TPU.
import os
if 'google.colab' in str(get_ipython()) and 'COLAB_TPU_ADDR' in os.environ:
import jax
import jax.tools.colab_tpu
jax.tools.colab_tpu.setup_tpu()
print('Connected to TPU.')
else:
# Otherwise print information about GPU.
!nvidia-smi
# Some more imports used in this Colab.
import glob
import os
import random
import shutil
import time
from absl import logging
import pandas as pd
import seaborn as sns
import tensorflow as tf
import tensorflow_datasets as tfds
from matplotlib import pyplot as plt
pd.options.display.max_colwidth = None
logging.set_verbosity(logging.INFO) # Shows logs during training.
```
### Explore checkpoints
This section contains shows how to use the `index.csv` table for model
selection.
See
[`vit_jax.checkpoint.get_augreg_df()`](https://github.com/google-research/vision_transformer/blob/ed1491238f5ff6099cca81087c575a215281ed14/vit_jax/checkpoint.py#L181-L228)
for a detailed description of the individual columns
```
# Load master table from Cloud.
with tf.io.gfile.GFile('gs://vit_models/augreg/index.csv') as f:
df = pd.read_csv(f)
# This is a pretty large table with lots of columns:
print(f'loaded {len(df):,} rows')
df.columns
# Number of distinct checkpoints
len(tf.io.gfile.glob('gs://vit_models/augreg/*.npz'))
# Any column prefixed with "adapt_" pertains to the fine-tuned checkpoints.
# Any column without that prefix pertains to the pre-trained checkpoints.
len(set(df.filename)), len(set(df.adapt_filename))
# Upstream AugReg parameters (section 3.3):
(
df.groupby(['ds', 'name', 'wd', 'do', 'sd', 'aug']).filename
.count().unstack().unstack().unstack()
.dropna(1, 'all').astype(int)
.iloc[:7] # Just show beginning of a long table.
)
# Downstream parameters (table 4)
(
df.groupby(['adapt_resolution', 'adapt_ds', 'adapt_lr', 'adapt_steps']).filename
.count().astype(str).unstack().unstack()
.dropna(1, 'all').fillna('')
)
# Let's first select the "best checkpoint" for every model. We show in the
# paper (section 4.5) that one can get a good performance by simply choosing the
# best model by final pre-train validation accuracy ("final-val" column).
# Pre-training with imagenet21k 300 epochs (ds=="i21k") gives the best
# performance in almost all cases (figure 6, table 5).
best_filenames = set(
df.query('ds=="i21k"')
.groupby('name')
.apply(lambda df: df.sort_values('final_val').iloc[-1])
.filename
)
# Select all finetunes from these models.
best_df = df.loc[df.filename.apply(lambda filename: filename in best_filenames)]
# Note: 9 * 68 == 612
len(best_filenames), len(best_df)
best_df.columns
# Note that this dataframe contains the models from the "i21k_300" column of
# table 3:
best_df.query('adapt_ds=="imagenet2012"').groupby('name').apply(
lambda df: df.sort_values('adapt_final_val').iloc[-1]
)[[
# Columns from upstream
'name', 'ds', 'filename',
# Columns from downstream
'adapt_resolution', 'infer_samples_per_sec','adapt_ds', 'adapt_final_test', 'adapt_filename',
]].sort_values('infer_samples_per_sec')
# Visualize the 2 (resolution) * 9 (models) * 8 (lr, steps) finetunings for a
# single dataset (Pets37).
# Note how larger models get better scores up to B/16 @384 even on this tiny
# dataset, if pre-trained sufficiently.
sns.relplot(
data=best_df.query('adapt_ds=="oxford_iiit_pet"'),
x='infer_samples_per_sec',
y='adapt_final_val',
hue='name',
style='adapt_resolution'
)
plt.gca().set_xscale('log');
# More details for a single pre-trained checkpoint.
best_df.query('name=="R26+S/32" and adapt_ds=="oxford_iiit_pet"')[[
col for col in best_df.columns if col.startswith('adapt_')
]].sort_values('adapt_final_val')
```
### Load a checkpoint
```
# Select a value from "adapt_filename" above that is a fine-tuned checkpoint.
filename = 'R26_S_32-i21k-300ep-lr_0.001-aug_light1-wd_0.1-do_0.0-sd_0.0--oxford_iiit_pet-steps_0k-lr_0.003-res_384'
tfds_name = filename.split('--')[1].split('-')[0]
model_config = models_config.AUGREG_CONFIGS[filename.split('-')[0]]
resolution = int(filename.split('_')[-1])
path = f'gs://vit_models/augreg/{filename}.npz'
print(f'{tf.io.gfile.stat(path).length / 1024 / 1024:.1f} MiB - {path}')
# Fetch dataset that the checkpoint was finetuned on.
# (Note that automatic download does not work with imagenet2012)
ds, ds_info = tfds.load(tfds_name, with_info=True)
ds_info
# Get model instance - no weights are initialized yet.
model = models.VisionTransformer(
num_classes=ds_info.features['label'].num_classes, **model_config)
# Load a checkpoint from cloud - for large checkpoints this can take a while...
params = checkpoint.load(path)
# Get a single example from dataset for inference.
d = next(iter(ds['test']))
def pp(img, sz):
"""Simple image preprocessing."""
img = tf.cast(img, float) / 255.0
img = tf.image.resize(img, [sz, sz])
return img
plt.imshow(pp(d['image'], resolution));
# Inferance on batch with single example.
logits, = model.apply({'params': params}, [pp(d['image'], resolution)], train=False)
# Plot logits (you can use tf.nn.softmax() to show probabilities instead).
plt.figure(figsize=(10, 4))
plt.bar(list(map(ds_info.features['label'].int2str, range(len(logits)))), logits)
plt.xticks(rotation=90);
```
#### Using `timm`
If you know PyTorch, you're probably already familiar with `timm`.
If not yet - it's your lucky day! Please check out their docs here:
https://rwightman.github.io/pytorch-image-models/
```
# Checkpoints can also be loaded directly into timm...
!pip install timm
import timm
import torch
# For available model names, see here:
# https://github.com/rwightman/pytorch-image-models/blob/master/timm/models/vision_transformer.py
# https://github.com/rwightman/pytorch-image-models/blob/master/timm/models/vision_transformer_hybrid.py
timm_model = timm.create_model(
'vit_small_r26_s32_384', num_classes=ds_info.features['label'].num_classes)
# Non-default checkpoints need to be loaded from local files.
if not tf.io.gfile.exists(f'{filename}.npz'):
tf.io.gfile.copy(f'gs://vit_models/augreg/{filename}.npz', f'{filename}.npz')
timm.models.load_checkpoint(timm_model, f'{filename}.npz')
def pp_torch(img, sz):
"""Simple image preprocessing for PyTorch."""
img = pp(img, sz)
img = img.numpy().transpose([2, 0, 1]) # PyTorch expects NCHW format.
return torch.tensor(img[None])
with torch.no_grad():
logits, = timm_model(pp_torch(d['image'], resolution)).detach().numpy()
# Same results as above (since we loaded the same checkpoint).
plt.figure(figsize=(10, 4))
plt.bar(list(map(ds_info.features['label'].int2str, range(len(logits)))), logits)
plt.xticks(rotation=90);
```
### Fine-tune
You want to be connected to a TPU or GPU runtime for fine-tuning.
Note that here we're just calling into the code. For more details see the
annotated Colab
https://colab.sandbox.google.com/github/google-research/vision_transformer/blob/linen/vit_jax.ipynb
Also note that Colab GPUs and TPUs are not very powerful. To run this code on
more powerful machines, see:
https://github.com/google-research/vision_transformer/#running-on-cloud
In particular, note that due to the Colab "TPU Node" setup, transfering data to
the TPUs is realtively slow (for example the smallest `R+Ti/16` model trains
faster on a single GPU than on 8 TPUs...)
#### TensorBoard
```
# Launch tensorboard before training - maybe click "reload" during training.
%load_ext tensorboard
%tensorboard --logdir=./workdirs
```
#### From tfds
```
# Create a new temporary workdir.
workdir = f'./workdirs/{int(time.time())}'
workdir
# Get config for specified model.
# Note that we can specify simply the model name (in which case the recommended
# checkpoint for that model is taken), or it can be specified by its full
# name.
config = augreg_config.get_config('R_Ti_16')
# A very small tfds dataset that only has a "train" split. We use this single
# split both for training & evaluation by splitting it further into 90%/10%.
config.dataset = 'tf_flowers'
config.pp.train = 'train[:90%]'
config.pp.test = 'train[90%:]'
# tf_flowers only has 3670 images - so the 10% evaluation split will contain
# 360 images. We specify batch_eval=120 so we evaluate on all but 7 of those
# images (remainder is dropped).
config.batch_eval = 120
# Some more parameters that you will often want to set manually.
# For example for VTAB we used steps={500, 2500} and lr={.001, .003, .01, .03}
config.base_lr = 0.01
config.shuffle_buffer = 1000
config.total_steps = 100
config.warmup_steps = 10
config.accum_steps = 0 # Not needed with R+Ti/16 model.
config.pp['crop'] = 224
# Call main training loop. See repository and above Colab for details.
state = train.train_and_evaluate(config, workdir)
```
#### From JPG files
The codebase supports training directly form JPG files on the local filesystem
instead of `tfds` datasets. Note that the throughput is somewhat reduced, but
that only is noticeable for very small models.
The main advantage of `tfds` datasets is that they are versioned and available
globally.
```
base = '.' # Store data on VM (ephemeral).
# Uncomment below lines if you want to download & persist files in your Google
# Drive instead. Note that Colab VMs are reset (i.e. files are deleted) after
# some time of inactivity. Storing data to Google Drive guarantees that it is
# still available next time you connect from a new VM.
# Note that this is significantly slower than reading from the VMs locally
# attached file system!
# from google.colab import drive
# drive.mount('/gdrive')
# base = '/gdrive/My Drive/vision_transformer_images'
# Download some dataset & unzip.
! rm -rf '$base/flower_photos'; mkdir -p '$base'
! (cd '$base' && curl https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz | tar xz)
# Since the default file format of above "tf_flowers" dataset is
# flower_photos/{class_name}/{filename}.jpg
# we first need to split it into a "train" (90%) and a "test" (10%) set:
# flower_photos/train/{class_name}/{filename}.jpg
# flower_photos/test/{class_name}/{filename}.jpg
def split(base_dir, test_ratio=0.1):
paths = glob.glob(f'{base_dir}/*/*.jpg')
random.shuffle(paths)
counts = dict(test=0, train=0)
for i, path in enumerate(paths):
split = 'test' if i < test_ratio * len(paths) else 'train'
*_, class_name, basename = path.split('/')
dst = f'{base_dir}/{split}/{class_name}/{basename}'
if not os.path.isdir(os.path.dirname(dst)):
os.makedirs(os.path.dirname(dst))
shutil.move(path, dst)
counts[split] += 1
print(f'Moved {counts["train"]:,} train and {counts["test"]:,} test images.')
split(f'{base}/flower_photos')
# Create a new temporary workdir.
workdir = f'./workdirs/{int(time.time())}'
workdir
# Read data from directory containing files.
# (See cell above for more config settings)
config.dataset = f'{base}/flower_photos'
# And fine-tune on images provided
opt = train.train_and_evaluate(config, workdir)
```
| github_jupyter |
# The instruction for data acquired from neuPrint
The instructions below was based on the description from [neuPrintExplorer](https://neuprint.janelia.org/help/cypherexamples). For more information, please visit the original website. A technical neuPrint paper exists [here](https://www.biorxiv.org/content/10.1101/2020.01.16.909465v1.full).
## Detailed information about the data
```
import pandas as pd
import matplotlib.pyplot as plt
df = pd.read_csv('Neuprint_connections.csv')
df.head
Id_pre = list(df['bodyId_pre'])
Id_post = list(df['bodyId_post'])
R_l = list(df['roi'])
wt = list(df['weight'])
t_pre = list(df['type_pre'])
ins_pre = list(df['instance_pre'])
t_post = list(df['type_post'])
ins_post = list(df['instance_post'])
```
### bodyId
bodyId is the unique identifier of a body in neuPrint. A body is a segmentation piece with at least 1 synapse. They are treated as nodes in this graph.
### nodes and edges
As shown above, the total number of edges in this graph is 8034510.
```
print('The number of source nodes is', len(pd.value_counts(Id_pre)))
print('The number of target nodes is', len(pd.value_counts(Id_post)))
print('The total number of nodes is', len(pd.value_counts(Id_pre + Id_post)))
```
### ROI
There are 64 regions of interest (ROI), the name of the regions and the numbers of connections in each ROI are listed below.
```
print(pd.value_counts(R_l))
```
### weight
Weight indicates the number of synapese in a connection between two neurons.
- It ranges from 1 to 1409 in this graph. The total counts for the connections with different weight are listed below.
- The first column shows the value of the weight, and the sencond column shows the number of connections with corresponding weight value.
```
print(pd.value_counts(wt))
```
### type
Type contains the neuron type of the body.
- By default cells get a type of the form NPXXX, where NP is a acronym for the neuropil with the largest overlap, such as CL for clamp, and XXX is numeric id
- If a cell is clearly recognised as a previously published type, then the systematic name will instead be replaced by that published name
- The types and the counts are listed in the blocks below. The first block shows the types of presynaptic neurons, and the second block shows the types of postsynaptic neurons. The first column in each blocks shows the name of the cell types, and the sencond column shows the number of of the corresponding type.
```
print(pd.value_counts(t_pre))
print(pd.value_counts(t_post))
```
### instance
Instance indicates a name that indicates a more specific instance of a neuron type.
- The instance and the counts are listed in the blocks below. The first block shows the instance of presynaptic neurons, and the second block shows the instance of postsynaptic neurons. The first column in each blocks shows the name of the instance, and the sencond column shows the number of of the corresponding instance.
```
print(pd.value_counts(ins_pre))
print(pd.value_counts(ins_post))
```
## More information about the node attributes
:Segment (:Neuron) nodes
#### bodyId: a unique number for each distinct segment
#### pre: Number of pre-synaptic sites on the segment
#### post: Number of post-synaptic sites on the segment
#### type: Cell type name for given neuron (if provided)
#### instance: String identifier for a neuron (if provided)
#### size: Number of voxels in the body
#### roiInfo: JSON string showing the pre and post breakdown for each ROI the neuron intersects.
#### roi: This property only exists for the ROIs that intersect this segment
#### tatus: Reconstruction status for a neuron. By convention, we broadly consider proofread neurons as being “Traced”.
#### cropped: Since datasets often involve a portion of a larger brain, cropped indicates that a significant portion of a neuron is cut-off by the dataset extents. By convention, all “Traced” neurons should be explicitly noted whether they are cropped or not.
_quote from [here](https://www.biorxiv.org/content/10.1101/2020.01.16.909465v1.full)_
| github_jupyter |
# Tutorial 1. Introduction to Statistical Quantities in Wind Engineering
## Part 1: Basic quantities
### Description: Wind data (measured or simulated) in wind engineering is usually recorded as a time series. Typical quantities measured are velocity (certain components) at a reference height or pressure measured at locations of interest along the structure. Evaluating the statistical quantities of these time series is a crucial task. In this tutorial a time series is generated and analyzed. Various statistical quantities, which are introduced during the lecture, are calculated for a generated signal. Some additional exercises are proposed for individual studies.
#### Students are advised to complete the proposed excercises
Project : Structural Wind Engineering WS19-20
Chair of Structural Analysis @ TUM - R. Wüchner, M. Péntek
Author : anoop.kodakkal@tum.de, mate.pentek@tum.de
Created on: 30.11.2015
Last update: 27.09.2019
Reference: G. Coles, Stuart. (2001). An introduction to statistical modeling of extreme values. Springer. 10.1007/978-1-4471-3675-0.
##### Contents:
1. Generating a time series as a superposition of constant, cosine and random signals
2. Introduction of some common statistical tools in python
3. Interquartile range and box plots
4. Probability Distribution Function (PDF)
5. Fast Fourier Transform (FFT)
```
# import python modules
import numpy as np
import scipy
from matplotlib import pyplot as plt
# import own modules
import custom_utilities as c_utils
from ipywidgets import interactive
```
#### Creating the time instances as an array
The start time, end time and the number of time steps are specified here for generating the time series.
```
# start time
start_time = 0.0
# end time
end_time = 10.0
# steps
n_steps = 10000
# time step
delta_time = end_time / (n_steps-1)
# time series
# generate grid size vector (array) 1D
time_series = np.arange(start_time, end_time + delta_time, delta_time)
```
#### Generating signals in time domain (from herein referred to as a certain series (of values)).
##### Three signals are created.
1. A Harmonic (cosine) signal with given amplitude and frequency
2. A constant signal with given amplitude
3. A random signal with specified distribution and given properties
###### 1. Cosine signal with given amplitude and frequency
```
# frequency of the cosine
cos_freq = 10
# amplitude of the cosine
cos_ampl = 1
# series of the cosine
cos_series = cos_ampl * np.cos(2*np.pi * cos_freq * time_series)
```
###### Let us look at the plot to see how the signal looks like
```
def plot_cosine_signal ( amplitude = 1, frequency = 10):
cos_series = amplitude * np.cos(2*np.pi * frequency * time_series)
fig = plt.figure(num=1, figsize=(15, 4))
ax = plt.axes()
ax.plot(time_series, cos_series)
ax.set_ylabel('Amplitude')
ax.set_xlabel('Time [s]')
ax.set_title('1. Cosine signal')
ax.grid(True)
plt.show()
cos_plot = interactive(plot_cosine_signal, amplitude = (0.0,50.0),frequency = (0.0,20.0))
cos_plot
```
### Exercise 1: Try different frequencies
Try different frequencies for the harmonic function.
###### 2. Constant signal with given amplitude
```
# amplitude of the constant
const_ampl = 10
# series of the constant
const_series = const_ampl * np.ones(len(time_series))
```
###### Let us look at the plot to see how the signals look like
```
plt.figure(num=2, figsize=(15, 4))
plt.plot(time_series, const_series)
plt.ylabel('Amplitude')
plt.xlabel('Time [s]')
plt.title('2. Constant signal')
plt.grid(True)
```
###### 3. Random signal with specified distribution and given properties
```
# random signal
# assuming normal distribution
# with given mean m = 0 and standard deviation std = 0.25
rand_m = 0.0
rand_std = 0.25
# series of the random
rand_series = np.random.normal(rand_m, rand_std, len(time_series))
```
###### Let us look at the plot to see how the signal looks like
```
plt.figure(num=3, figsize=(15, 4))
plt.plot(time_series, rand_series)
plt.ylabel('Amplitude')
plt.xlabel('Time [s]')
plt.title('3. Random signal')
plt.grid(True)
```
### Exercise 2 : Different distributions and parameters for random signal
Instead of the [normal](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.random.normal.html) distribution for the random signal try [lognormal](https://docs.scipy.org/doc/numpy-1.14.0/reference/generated/numpy.random.lognormal.html), [beta](https://docs.scipy.org/doc/numpy-1.15.1/reference/generated/numpy.random.beta.html), [standard normal](https://docs.scipy.org/doc/numpy-1.15.1/reference/generated/numpy.random.randn.html) and [uniform](https://docs.scipy.org/doc/numpy-1.15.0/reference/generated/numpy.random.uniform.html) distribution
```
#rand_series = np.random.lognormal(0, 0.25, len(time_series))
#rand_series = np.random.beta(1, 0.25, len(time_series))
#rand_series = np.random.rand(len(time_series))
#rand_series = np.random.uniform(0,1,len(time_series))
```
#### 4. Generic signal - for example a superposition of the above ones
A general signal (here) is represented as a superposition of the above three - constant, cosine and random signals
###### Superposed signal
The above three signals are superposed with corresponding weights
```
const_coeff = 1
cos_coeff = 0.25
rand_coeff = 0.25
superposed_series = const_coeff * const_series + cos_coeff * cos_series + rand_coeff * rand_series
```
###### Let us look at the plot to see how the signal look like
```
# coefs -> weighting factors for the respective series of signals
def plot_superposed_signal(const_coeff = 1,cos_coeff = 0.25,rand_coeff = 0.25):
superposed_series = const_coeff * const_series + cos_coeff * cos_series + rand_coeff * rand_series
fig = plt.figure(num=4, figsize=(15, 4))
ax = plt.axes()
ax.plot(time_series, superposed_series)
ax.set_ylabel('Amplitude')
ax.set_xlabel('Time [s]')
ax.set_title('4. Superposed signal')
ax.grid(True)
plt.show()
```
###### Let us look at the plot to see how the signal look like
```
mean_plot=interactive(plot_superposed_signal, const_coeff = (0.0,10.0),cos_coeff = (0.0,5.0),rand_coeff = (0.0,2.0))
mean_plot
```
### Exercise 3: Different weights for superposition
Try different weights for the superposition. What do you observe in the plots?
Try different frequencies for the cosine function and observe the difference in the superposed signal.
## Check Point 1: Discussion
#### Discuss among groups the observations and outcomes from exercise 1-3.
## 1.1 Statistical tools and quantities used to evaluate the signal
##### The following statistical quantities are computed for the given signal.
1. Mean (Arithmetic)
2. Root Mean Square (RMS)
3. Median
4. Standard deviation
5. Skewness
Recall from the lecture the definitions of these quantities.
These quantites can be computed using the inbuilt functions of numpy
[mean (arithmetic)](https://docs.scipy.org/doc/numpy/reference/generated/numpy.mean.html),
[median](https://docs.scipy.org/doc/numpy/reference/generated/numpy.median.html),
[standard deviation](https://docs.scipy.org/doc/numpy/reference/generated/numpy.std.html#numpy.std)
and
[skewness](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.stats.skew.html)
###### 1. Cosine signal with given amplitude and frequency
```
# computing statistical quantitites (scalar values) and "converting" to an array for later plotting
cos_series_m = np.mean(cos_series) * np.ones(len(time_series))
cos_series_std = np.std(cos_series) * np.ones(len(time_series))
cos_series_rms = np.sqrt(np.mean(np.square(cos_series))) * np.ones(len(time_series))
# printing statistical quantitites (scalar values) to the console
print('Mean: ', np.mean(cos_series))
print('STD: ', np.std(cos_series))
print('RMS: ', np.sqrt(np.mean(np.square(cos_series))))
print('Median: ', np.median(cos_series))
print('Skewness: ',(np.mean(cos_series) - np.median(cos_series))/np.std(cos_series))
```
###### 2. Constant signal with given amplitude
```
const_series_m = np.mean(const_series) * np.ones(len(time_series))
const_series_std = np.std(const_series) * np.ones(len(time_series))
const_series_rms = np.sqrt(np.mean(np.square(const_series))) * np.ones(len(time_series))
print('Mean: ', np.mean(const_series))
print('STD: ', np.std(const_series))
print('RMS: ', np.sqrt(np.mean(np.square(const_series))))
print('Median: ', np.median(const_series))
print('Skewness: ', (np.mean(const_series) - np.median(const_series))/np.std(const_series))
```
###### 3. Random signal with specified distribution and given properties
```
rand_series_m = np.mean(rand_series) * np.ones(len(time_series))
rand_series_std = np.std(rand_series) * np.ones(len(time_series))
rand_series_rms = np.sqrt(np.mean(np.square(rand_series))) * np.ones(len(time_series))
print('Mean: ', np.mean(rand_series))
print('STD: ', np.std(rand_series))
print('RMS: ', np.sqrt(np.mean(np.square(rand_series))))
print('Median: ', np.median(rand_series))
print('Skewness: ', (np.mean(rand_series) - np.median(rand_series))/np.std(rand_series))
```
#### Superposed signal
```
superposed_series_m = np.mean(superposed_series) * np.ones(len(time_series))
superposed_series_std = np.std(superposed_series) * np.ones(len(time_series))
superposed_series_rms = np.sqrt(np.mean(np.square(superposed_series))) * np.ones(len(time_series))
print('Mean: ', np.mean(superposed_series))
print('STD: ', np.std(superposed_series))
print('RMS: ', np.sqrt(np.mean(np.square(superposed_series))))
print('Median: ', np.median(superposed_series))
print('Skewness: ', (np.mean(superposed_series) - np.median(superposed_series))/np.std(superposed_series))
```
What do the mean, median, mode, RMS, standard deviation and skewness represent?
### Histogram of the signals
The variation of each signal with time and their histograms are plotted.
```
# const
plt.figure(num=5, figsize=(15, 4))
plt.suptitle('Constant signal')
plt.subplot(1, 2, 1)
plt.plot(time_series, const_series,
time_series, const_series_m,
time_series, const_series_m - const_series_std,
time_series, const_series_m + const_series_std,
time_series, const_series_rms)
plt.ylabel('Amplitude')
plt.title('Time series')
plt.grid(True)
bins = 100
plt.subplot(1, 2, 2)
plt.hist(const_series, bins)
plt.title('Histogram of ' + str(n_steps) +' values')
plt.ylabel('Frequency of occurrence.')
plt.grid(True)
# cos
plt.figure(num=6, figsize=(15, 4))
plt.suptitle('Cosine signal')
plt.subplot(1, 2, 1)
plt.plot(time_series, cos_series)
plt.plot(time_series, cos_series_m, label = 'Mean')
plt.plot(time_series, cos_series_m - cos_series_std, label = 'Mean - STD')
plt.plot(time_series, cos_series_m + cos_series_std,label = 'Mean + STD')
plt.plot(time_series, cos_series_rms, label = 'RMS')
plt.ylabel('Amplitude')
plt.legend()
plt.grid(True)
plt.subplot(1, 2, 2)
plt.hist(cos_series, bins)
plt.ylabel('Frequency of occurrence.')
plt.grid(True)
# rand
plt.figure(num=7, figsize=(15, 4))
plt.suptitle('Random signal')
plt.subplot(1, 2, 1)
plt.plot(time_series, rand_series,
time_series, rand_series_m,
time_series, rand_series_m - rand_series_std,
time_series, rand_series_m + rand_series_std,
time_series, rand_series_rms)
plt.ylabel('Amplitude')
plt.grid(True)
plt.subplot(1, 2, 2)
plt.hist(rand_series, bins)
plt.ylabel('Frequency of occurrence.')
plt.grid(True)
# superposed
plt.figure(num=8, figsize=(15, 4))
plt.suptitle('Superposed signal')
plt.subplot(1, 2, 1)
plt.plot(time_series, superposed_series,
time_series, superposed_series_m,
time_series, superposed_series_m - superposed_series_std,
time_series, superposed_series_m + superposed_series_std,
time_series, superposed_series_rms)
plt.ylabel('Amplitude')
plt.xlabel('Time [s]')
plt.grid(True)
plt.subplot(1, 2, 2)
plt.hist(superposed_series, bins)
plt.ylabel('Frequency of occurrence.')
plt.xlabel('Amplitude')
plt.grid(True)
```
### Interquartile range and percentile
The [interquartile range (IQR)](https://en.wikipedia.org/wiki/Interquartile_range) , also called the midspread or middle 50%, or technically H-spread, is a measure of statistical dispersion. This is computed as the difference between 75th and 25th percentiles, or between upper and lower quartiles. In statistics of extreme values the interquartile range is also considered along with standard deviation as a measure of the dispersion.
The [percentile](https://en.wikipedia.org/wiki/Percentile) is a measure used in statistics indicating the value below which a given percentage of observations in a group of observations fall. These quantites can be computed using the inbuilt functions of numpy
[interquartile range (IQR)](https://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.iqr.html)
[percentile](https://docs.scipy.org/doc/numpy/reference/generated/numpy.percentile.html)
```
iqr = scipy.stats.iqr(superposed_series)
q75, q25 = np.percentile(superposed_series, [75 ,25])
print('Interquartile range = ',iqr, 'Interquantile range computed = ', q75-q25)
```
The [boxplots](https://en.wikipedia.org/wiki/Box_plot) can be obtained from the interquartile range to identify possible outliers. The box indicate the middle quartile and the lines extending indicating the variability outside the lower and upper quartiles. The in built python function [boxplots](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.boxplot.html) can be used for plotting.
```
# coefs -> weighting factors for the respective series of signals
def boxplot_superposed_signal(const_coeff = 1,cos_coeff = 0.25,rand_coeff = 0.25):
superposed_series = const_coeff * const_series + cos_coeff * cos_series + rand_coeff * rand_series
fig = plt.figure(num=9, figsize=(6, 8))
ax = plt.axes()
ax.boxplot(superposed_series)
ax.grid(True)
plt.show()
```
###### Let us look at the plot to see how the signal look like
```
box_plot=interactive(boxplot_superposed_signal, const_coeff = (0.0,10.0),cos_coeff = (0.0,5.0),rand_coeff = (0.0,10))
box_plot
```
### Probability Distribution Function (PDF) and Cumulative Distribution Function (CDF)
The PDF and CDF of the signals are derived and are plotted later. Recall from the lecture the definitions of PDF, CDF of a continuous random variables.
##### Tip: Have a look at the get_pdf function in the "custom_utilities.py" for details
```
# const
[const_pdf_x, const_pdf_y] = c_utils.get_pdf(const_series,'Constant')
# the 'Constant' is used for obtaining pdf of a constant signal.
# check the implimentation for details
# cos
[cos_pdf_x, cos_pdf_y] = c_utils.get_pdf(cos_series)
# rand
[rand_pdf_x, rand_pdf_y] = c_utils.get_pdf(rand_series)
# superposed
[superposed_pdf_x, superposed_pdf_y] = c_utils.get_pdf(superposed_series)
```
### Converting to Frequency domain - Fast Fourier Transform (FFT)
FFT computes the frequency contents of the given signal. Recall from the lecture the basic definitions and procedure for FFT.
##### Tip: Have a look at the get_fft function in the "custom_utilities.py" for details
```
# sampling frequency the same in this case for all time series
sampling_freq = 1/delta_time
# const
[const_freq_half, const_series_fft] = c_utils.get_fft(const_series, sampling_freq)
# cos
[cos_freq_half, cos_series_fft] = c_utils.get_fft(cos_series, sampling_freq)
# rand
[rand_freq_half, rand_series_fft] = c_utils.get_fft(rand_series, sampling_freq)
# superposed
[superposed_freq_half, superposed_series_fft] = c_utils.get_fft(superposed_series, sampling_freq)
# pdf, cdf and frequency domain
plt.rcParams["figure.figsize"] = (15,4)
# const
plt.figure(num=10)
plt.suptitle('Constant signal')
plt.subplot(1,3,1)
plt.plot(const_pdf_x, const_pdf_y)
plt.xlabel(' ')
plt.ylabel('PDF(Amplitude)')
plt.title('PDF')
plt.grid(True)
const_ecdf = c_utils.get_ecdf(const_pdf_x, const_pdf_y)
plt.subplot(1,3,2)
plt.plot(const_pdf_x, const_ecdf)
plt.ylabel('CDF(Amplitude)')
plt.title('Empirical CDF')
plt.grid(True)
plt.subplot(1,3,3)
plt.plot(const_freq_half, const_series_fft)
plt.xlim([1, 25])
plt.ylabel('|Amplitude|')
plt.title('Frequency domain using FFT')
plt.grid(True)
plt.show()
# cos
plt.figure(num=11)
plt.suptitle('Cosine signal')
plt.subplot(1,3,1)
plt.plot(cos_pdf_x, cos_pdf_y)
plt.xlabel(' ')
plt.ylabel('PDF(Amplitude)')
plt.grid(True)
cos_ecdf = c_utils.get_ecdf(cos_pdf_x, cos_pdf_y)
plt.subplot(1,3,2)
plt.plot(cos_pdf_x, cos_ecdf)
plt.ylabel('CDF(Amplitude)')
plt.grid(True)
plt.subplot(1,3,3)
plt.plot(cos_freq_half, cos_series_fft)
plt.xlim([1, 25])
plt.ylabel('|Amplitude|')
plt.grid(True)
plt.show()
# rand
plt.figure(num=12)
plt.suptitle('Random signal')
plt.subplot(1,3,1)
plt.plot(rand_pdf_x, rand_pdf_y)
plt.xlabel(' ')
plt.ylabel('PDF(Amplitude)')
plt.grid(True)
rand_ecdf = c_utils.get_ecdf(rand_pdf_x, rand_pdf_y)
plt.subplot(1,3,2)
plt.plot(rand_pdf_x, rand_ecdf)
plt.ylabel('CDF(Amplitude)')
plt.grid(True)
plt.subplot(1,3,3)
plt.plot(rand_freq_half, rand_series_fft)
plt.xlim([1, 25])
plt.ylabel('|Amplitude|')
plt.grid(True)
plt.show()
# superposed
plt.figure(num=13)
plt.suptitle('Superposed signal')
plt.subplot(1,3,1)
plt.plot(superposed_pdf_x, superposed_pdf_y)
plt.xlabel(' ')
plt.ylabel('PDF(Amplitude)')
plt.xlabel('Amplitude')
plt.grid(True)
superposed_ecdf = c_utils.get_ecdf(superposed_pdf_x, superposed_pdf_y)
plt.subplot(1,3,2)
plt.plot(superposed_pdf_x, superposed_ecdf)
plt.ylabel('CDF(Amplitude)')
plt.xlabel('Amplitude')
plt.grid(True)
plt.subplot(1,3,3)
plt.plot(superposed_freq_half, superposed_series_fft)
plt.ylim([0, 0.4])
plt.xlim([1, 25])
plt.xlabel('Frequency [Hz]')
plt.ylabel('|Amplitude|')
plt.grid(True)
plt.show()
```
PDF follows the normalized hystograms. Observe the predominant frequency in the superimposed signal.
### Excercise 4: Try two or more harmonic function
Try two or more cosine functions and superimpose them. What difference do you observe?
What do you observe in the FFT plots?
## Check Point 2: Discussion
#### Discuss among groups the uses of various statistical quantities and their significance.
| github_jupyter |
```
%matplotlib inline
```
# `scikit-learn` - Machine Learning in Python
[scikit-learn](http://scikit-learn.org) is a simple and efficient tool for data mining and data analysis. It is built on [NumPy](www.numpy.org), [SciPy](https://www.scipy.org/), and [matplotlib](https://matplotlib.org/). The following examples show some of `scikit-learn`'s power. For a complete list, go to the official homepage under [examples](http://scikit-learn.org/stable/auto_examples/index.html) or [tutorials](http://scikit-learn.org/stable/tutorial/index.html).
## Blind source separation using FastICA
This example of estimating sources from noisy data is adapted from [`plot_ica_blind_source_separation`](http://scikit-learn.org/stable/auto_examples/decomposition/plot_ica_blind_source_separation.html).
```
import numpy as np
import matplotlib.pyplot as plt
from scipy import signal
from sklearn.decomposition import FastICA, PCA
# Generate sample data
n_samples = 2000
time = np.linspace(0, 8, n_samples)
s1 = np.sin(2 * time) # Signal 1: sinusoidal signal
s2 = np.sign(np.sin(3 * time)) # Signal 2: square signal
s3 = signal.sawtooth(2 * np.pi * time) # Signal 3: saw tooth signal
S = np.c_[s1, s2, s3]
S += 0.2 * np.random.normal(size=S.shape) # Add noise
S /= S.std(axis=0) # Standardize data
# Mix data
A = np.array([[1, 1, 1], [0.5, 2, 1.0], [1.5, 1.0, 2.0]]) # Mixing matrix
X = np.dot(S, A.T) # Generate observations
# Compute ICA
ica = FastICA(n_components=3)
S_ = ica.fit_transform(X) # Reconstruct signals
A_ = ica.mixing_ # Get estimated mixing matrix
# For comparison, compute PCA
pca = PCA(n_components=3)
H = pca.fit_transform(X) # Reconstruct signals based on orthogonal components
# Plot results
plt.figure(figsize=(12, 4))
models = [X, S, S_, H]
names = ['Observations (mixed signal)', 'True Sources',
'ICA recovered signals', 'PCA recovered signals']
colors = ['red', 'steelblue', 'orange']
for ii, (model, name) in enumerate(zip(models, names), 1):
plt.subplot(2, 2, ii)
plt.title(name)
for sig, color in zip(model.T, colors):
plt.plot(sig, color=color)
plt.subplots_adjust(0.09, 0.04, 0.94, 0.94, 0.26, 0.46)
plt.show()
```
# Anomaly detection with Local Outlier Factor (LOF)
This example presents the Local Outlier Factor (LOF) estimator. The LOF algorithm is an unsupervised outlier detection method which computes the local density deviation of a given data point with respect to its neighbors. It considers as outlier samples that have a substantially lower density than their neighbors. This example is adapted from [`plot_lof`](http://scikit-learn.org/stable/auto_examples/neighbors/plot_lof.html).
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn.neighbors import LocalOutlierFactor
# Generate train data
X = 0.3 * np.random.randn(100, 2)
# Generate some abnormal novel observations
X_outliers = np.random.uniform(low=-4, high=4, size=(20, 2))
X = np.r_[X + 2, X - 2, X_outliers]
# fit the model
clf = LocalOutlierFactor(n_neighbors=20)
y_pred = clf.fit_predict(X)
y_pred_outliers = y_pred[200:]
# Plot the level sets of the decision function
xx, yy = np.meshgrid(np.linspace(-5, 5, 50), np.linspace(-5, 5, 50))
Z = clf._decision_function(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.title("Local Outlier Factor (LOF)")
plt.contourf(xx, yy, Z, cmap=plt.cm.Blues_r)
a = plt.scatter(X[:200, 0], X[:200, 1], c='white', edgecolor='k', s=20)
b = plt.scatter(X[200:, 0], X[200:, 1], c='red', edgecolor='k', s=20)
plt.axis('tight')
plt.xlim((-5, 5))
plt.ylim((-5, 5))
plt.legend([a, b], ["normal observations", "abnormal observations"], loc="upper left")
plt.show()
```
# SVM: Maximum margin separating hyperplane
Plot the maximum margin separating hyperplane within a two-class separable dataset using a Support Vector Machine classifier with a linear kernel. This example is adapted from [`plot_separating_hyperplane`](http://scikit-learn.org/stable/auto_examples/svm/plot_separating_hyperplane.html).
```
import numpy as np
import matplotlib.pyplot as plt
from sklearn import svm
from sklearn.datasets import make_blobs
# we create 40 separable points
X, y = make_blobs(n_samples=40, centers=2, random_state=6)
# fit the model, don't regularize for illustration purposes
clf = svm.SVC(kernel='linear', C=1000)
clf.fit(X, y)
plt.scatter(X[:, 0], X[:, 1], c=y, s=30, cmap=plt.cm.Paired)
# plot the decision function
ax = plt.gca()
xlim = ax.get_xlim()
ylim = ax.get_ylim()
# create grid to evaluate model
xx = np.linspace(xlim[0], xlim[1], 30)
yy = np.linspace(ylim[0], ylim[1], 30)
YY, XX = np.meshgrid(yy, xx)
xy = np.vstack([XX.ravel(), YY.ravel()]).T
Z = clf.decision_function(xy).reshape(XX.shape)
# plot decision boundary and margins
ax.contour(XX, YY, Z, colors='k', levels=[-1, 0, 1], alpha=0.5,
linestyles=['--', '-', '--'])
# plot support vectors
ax.scatter(clf.support_vectors_[:, 0], clf.support_vectors_[:, 1], s=100,
linewidth=1, facecolors='none')
plt.show()
```
# `Scikit-Image` - Image processing in python
[scikit-image](http://scikit-image.org/) is a collection of algorithms for image processing and is based on [scikit-learn](http://scikit-learn.org). The following examples show some of `scikit-image`'s power. For a complete list, go to the official homepage under [examples](http://scikit-image.org/docs/stable/auto_examples/).
## Sliding window histogram
Histogram matching can be used for object detection in images. This example extracts a single coin from the `skimage.data.coins` image and uses histogram matching to attempt to locate it within the original image. This example is adapted from [`plot_windowed_histogram`](http://scikit-image.org/docs/stable/auto_examples/features_detection/plot_windowed_histogram.html).
```
from __future__ import division
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
from skimage import data, transform
from skimage.util import img_as_ubyte
from skimage.morphology import disk
from skimage.filters import rank
def windowed_histogram_similarity(image, selem, reference_hist, n_bins):
# Compute normalized windowed histogram feature vector for each pixel
px_histograms = rank.windowed_histogram(image, selem, n_bins=n_bins)
# Reshape coin histogram to (1,1,N) for broadcast when we want to use it in
# arithmetic operations with the windowed histograms from the image
reference_hist = reference_hist.reshape((1, 1) + reference_hist.shape)
# Compute Chi squared distance metric: sum((X-Y)^2 / (X+Y));
# a measure of distance between histograms
X = px_histograms
Y = reference_hist
num = (X - Y) ** 2
denom = X + Y
denom[denom == 0] = np.infty
frac = num / denom
chi_sqr = 0.5 * np.sum(frac, axis=2)
# Generate a similarity measure. It needs to be low when distance is high
# and high when distance is low; taking the reciprocal will do this.
# Chi squared will always be >= 0, add small value to prevent divide by 0.
similarity = 1 / (chi_sqr + 1.0e-4)
return similarity
# Load the `skimage.data.coins` image
img = img_as_ubyte(data.coins())
# Quantize to 16 levels of greyscale; this way the output image will have a
# 16-dimensional feature vector per pixel
quantized_img = img // 16
# Select the coin from the 4th column, second row.
# Co-ordinate ordering: [x1,y1,x2,y2]
coin_coords = [184, 100, 228, 148] # 44 x 44 region
coin = quantized_img[coin_coords[1]:coin_coords[3],
coin_coords[0]:coin_coords[2]]
# Compute coin histogram and normalize
coin_hist, _ = np.histogram(coin.flatten(), bins=16, range=(0, 16))
coin_hist = coin_hist.astype(float) / np.sum(coin_hist)
# Compute a disk shaped mask that will define the shape of our sliding window
# Example coin is ~44px across, so make a disk 61px wide (2 * rad + 1) to be
# big enough for other coins too.
selem = disk(30)
# Compute the similarity across the complete image
similarity = windowed_histogram_similarity(quantized_img, selem, coin_hist,
coin_hist.shape[0])
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(12, 4))
axes[0].imshow(quantized_img, cmap='gray')
axes[0].set_title('Quantized image')
axes[0].axis('off')
axes[1].imshow(coin, cmap='gray')
axes[1].set_title('Coin from 2nd row, 4th column')
axes[1].axis('off')
axes[2].imshow(img, cmap='gray')
axes[2].imshow(similarity, cmap='hot', alpha=0.5)
axes[2].set_title('Original image with overlaid similarity')
axes[2].axis('off')
plt.tight_layout()
plt.show()
```
## Local Thresholding
If the image background is relatively uniform, then you can use a global threshold value as presented above. However, if there is large variation in the background intensity, adaptive thresholding (a.k.a. local or dynamic thresholding) may produce better results. This example is adapted from [`plot_thresholding`](http://scikit-image.org/docs/dev/auto_examples/xx_applications/plot_thresholding.html#local-thresholding).
```
from skimage.filters import threshold_otsu, threshold_local
image = data.page()
global_thresh = threshold_otsu(image)
binary_global = image > global_thresh
block_size = 35
adaptive_thresh = threshold_local(image, block_size, offset=10)
binary_adaptive = image > adaptive_thresh
fig, axes = plt.subplots(ncols=3, figsize=(16, 6))
ax = axes.ravel()
plt.gray()
ax[0].imshow(image)
ax[0].set_title('Original')
ax[1].imshow(binary_global)
ax[1].set_title('Global thresholding')
ax[2].imshow(binary_adaptive)
ax[2].set_title('Adaptive thresholding')
for a in ax:
a.axis('off')
plt.show()
```
## Finding local maxima
The peak_local_max function returns the coordinates of local peaks (maxima) in an image. A maximum filter is used for finding local maxima. This operation dilates the original image and merges neighboring local maxima closer than the size of the dilation. Locations, where the original image is equal to the dilated image, are returned as local maxima. This example is adapted from [`plot_peak_local_max`](http://scikit-image.org/docs/stable/auto_examples/segmentation/plot_peak_local_max.html).
```
from scipy import ndimage as ndi
import matplotlib.pyplot as plt
from skimage.feature import peak_local_max
from skimage import data, img_as_float
im = img_as_float(data.coins())
# image_max is the dilation of im with a 20*20 structuring element
# It is used within peak_local_max function
image_max = ndi.maximum_filter(im, size=20, mode='constant')
# Comparison between image_max and im to find the coordinates of local maxima
coordinates = peak_local_max(im, min_distance=20)
# display results
fig, axes = plt.subplots(1, 3, figsize=(12, 5), sharex=True, sharey=True,
subplot_kw={'adjustable': 'box'})
ax = axes.ravel()
ax[0].imshow(im, cmap=plt.cm.gray)
ax[0].axis('off')
ax[0].set_title('Original')
ax[1].imshow(image_max, cmap=plt.cm.gray)
ax[1].axis('off')
ax[1].set_title('Maximum filter')
ax[2].imshow(im, cmap=plt.cm.gray)
ax[2].autoscale(False)
ax[2].plot(coordinates[:, 1], coordinates[:, 0], 'r.')
ax[2].axis('off')
ax[2].set_title('Peak local max')
fig.tight_layout()
plt.show()
```
## Label image region
This example shows how to segment an image with image labeling. The following steps are applied:
1. Thresholding with automatic Otsu method
2. Close small holes with binary closing
3. Remove artifacts touching image border
4. Measure image regions to filter small objects
This example is adapted from [`plot_label`](http://scikit-image.org/docs/stable/auto_examples/segmentation/plot_label.html).
```
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from skimage import data
from skimage.filters import threshold_otsu
from skimage.segmentation import clear_border
from skimage.measure import label, regionprops
from skimage.morphology import closing, square
from skimage.color import label2rgb
image = data.coins()[50:-50, 50:-50]
# apply threshold
thresh = threshold_otsu(image)
bw = closing(image > thresh, square(3))
# remove artifacts connected to image border
cleared = clear_border(bw)
# label image regions
label_image = label(cleared)
image_label_overlay = label2rgb(label_image, image=image)
fig, ax = plt.subplots(figsize=(10, 6))
ax.imshow(image_label_overlay)
for region in regionprops(label_image):
# take regions with large enough areas
if region.area >= 100:
# draw rectangle around segmented coins
minr, minc, maxr, maxc = region.bbox
rect = mpatches.Rectangle((minc, minr), maxc - minc, maxr - minr,
fill=False, edgecolor='red', linewidth=2)
ax.add_patch(rect)
ax.set_axis_off()
plt.tight_layout()
plt.show()
```
| github_jupyter |
# An Introduction to SageMaker Neural Topic Model
***Unsupervised representation learning and topic extraction using Neural Topic Model***
1. [Introduction](#Introduction)
1. [Data Preparation](#Data-Preparation)
1. [Model Training](#Model-Training)
1. [Model Hosting and Inference](#Model-Hosting-and-Inference)
1. [Model Exploration](#Model-Exploration)
---
# Introduction
Amazon SageMaker Neural Topic Model (NTM) is an unsupervised learning algorithm that attempts to describe a set of observations as a mixture of distinct categories. NTM is most commonly used to discover a user-specified number of topics shared by documents within a text corpus. Here each observation is a document, the features are the presence (or occurrence count) of each word, and the categories are the topics. Since the method is unsupervised, the topics are not specified upfront and are not guaranteed to align with how a human may naturally categorize documents. The topics are learned as a probability distribution over the words that occur in each document. Each document, in turn, is described as a mixture of topics.
In this notebook, we will use the Amazon SageMaker NTM algorithm to train a model on the [20NewsGroups](https://archive.ics.uci.edu/ml/datasets/Twenty+Newsgroups) data set. This data set has been widely used as a topic modeling benchmark.
The main goals of this notebook are as follows:
1. learn how to obtain and store data for use in Amazon SageMaker,
2. create an AWS SageMaker training job on a data set to produce an NTM model,
3. use the model to perform inference with an Amazon SageMaker endpoint.
4. explore trained model and visualized learned topics
If you would like to know more please check out the [SageMaker Neural Topic Model Documentation](https://docs.aws.amazon.com/sagemaker/latest/dg/ntm.html).
### A Brief Overview of SageMaker NTM
Topic models are a classical example of probablistic graphical models that involve challenging posterior inference problems. We implement topic modeling under a neural-network based variational inference framework. The difficult inference problem is framed as an optimization problem solved by scalable methods such as stochastic gradient descent. Compared to conventional inference schemes, the neural-network implementation allows for scalable model training as well as low-latency inference. Furthermore, the flexibility of the neural inference framework allows us to more quickly add new functionalities and serve a wider range of customer use cases.
The high-level diagram of SageMaker NTM is shown below:
<img src="ntm_diagram.png" width="600">
$$
\begin{equation}
\begin{split}
\textrm{Encoder} \ q(z\vert x): & \\
& \pi = f_X^{MLP}(x), \quad \mu(x) = l_1(\pi), \quad \log \sigma(x) = l_2(\pi)\\
& h(x, \epsilon) = \mu + \sigma \epsilon, \ \textrm{where} \ \epsilon \sim \mathcal{N}(0,I),\quad z=g(h), \ \textrm{where} \ h \sim \mathcal{N}(\mu, \sigma^2 I) \\
\textrm{Decoder} \ p(x\vert z): & \\
& y(z) = \textrm{softmax}(Wz+b), \quad \log p(x\vert z) = \sum x \odot \log(y(z))
\end{split}
\end{equation}
$$
where $l_1$ and $l_2$ are linear transformations with bias.
### Beyond Text Data
In principle, topic models can be applied to types of data beyond text documents. For example, topic modeling has been applied on network traffice data to discover [peer-to-peer applications usage patterns](http://isis.poly.edu/~baris/papers/GangsOfInternet-CNS13.pdf). We would be glad to hear about your novel use cases and are happy to help provide additional information. Please feel free to post questions or feedback at our [GitHub repository](https://github.com/awslabs/amazon-sagemaker-examples) or in the [Amazon SageMaker](https://forums.aws.amazon.com/forum.jspa?forumID=285) section of AWS Developer Forum.
# Install Python packages
```
import sys
!{sys.executable} -m pip install "scikit_learn==0.20.0" "nltk==3.4.4"
```
---
# Data Preparation
The 20Newsgroups data set is a collection of approximately 20,000 newsgroup documents, partitioned (nearly) evenly across 20 different newsgroups. This collection has become a popular data set for experiments in text applications of machine learning techniques, such as text classification and text clustering. Here, we will see what topics we can learn from this set of documents with NTM. The data setis available at the UCI Machine Learning Repository at this [location](https://archive.ics.uci.edu/ml/datasets/Twenty+Newsgroups). Please aware of the following requirements about ackonwledge, copyright and availability, cited from the [data set description page](https://archive.ics.uci.edu/ml/machine-learning-databases/20newsgroups-mld/20newsgroups.data.html).
> **Acknowledgements, Copyright Information, and Availability**
>You may use this material free of charge for any educational purpose, provided attribution is given in any lectures or publications that make use of this material.
## Fetching Data Set
First let's define the folder to hold the data and clean the content in it which might be from previous experiments.
```
import os
import shutil
data_dir = "20_newsgroups_bulk"
if os.path.exists(data_dir): # cleanup existing data folder
shutil.rmtree(data_dir)
```
Now we can download the data. We download the [`20 newsgroups dataset`](http://qwone.com/~jason/20Newsgroups/). The `20 newsgroups dataset` consists of 20000 messages taken from 20 Usenet newsgroups.
```
!aws s3 cp s3://sagemaker-sample-files/datasets/text/20_newsgroups/20_newsgroups_bulk.tar.gz .
!tar xzf 20_newsgroups_bulk.tar.gz
!ls 20_newsgroups_bulk
file_list = [os.path.join(data_dir, f) for f in os.listdir(data_dir)]
print("Number of files:", len(file_list))
import pandas as pd
documents_count = 0
for file in file_list:
df = pd.read_csv(file, header=None, names=["text"])
documents_count = documents_count + df.shape[0]
print("Number of documents:", documents_count)
```
This following function will remove the header, footer and quotes (of earlier messages in each text).
```
def strip_newsgroup_item(item):
item = strip_newsgroup_header(item)
item = strip_newsgroup_quoting(item)
item = strip_newsgroup_footer(item)
return item
from sklearn.datasets.twenty_newsgroups import (
strip_newsgroup_header,
strip_newsgroup_quoting,
strip_newsgroup_footer,
)
data = []
for file in file_list:
print(f"Processing {file}")
label = file.split("/")[1]
df = pd.read_csv(file, header=None, names=["text"])
df["text"] = df["text"].apply(strip_newsgroup_item)
data.extend(df["text"].tolist())
```
As we can see below, the entries in the data set are just plain text paragraphs. We will need to process them into a suitable data format.
```
data[10:13]
```
---
## From Plain Text to Bag-of-Words (BOW)
The input documents to the algorithm, both in training and inference, need to be vectors of integers representing word counts. This is so-called bag-of-words (BOW) representation. To convert plain text to BOW, we need to first "tokenize" our documents, i.e identify words and assign an integer id to each of them.
$$
\text{"cat"} \mapsto 0, \; \text{"dog"} \mapsto 1 \; \text{"bird"} \mapsto 2, \ldots
$$
Then, we count the occcurence of each of the tokens in each document and form BOW vectors as illustrated in the following example:
$$
w = \text{"cat bird bird bird cat"} \quad \longmapsto \quad w = [2, 0, 3, 0, \ldots, 0]
$$
Also, note that many real-world applications have large vocabulary sizes. It may be necessary to represent the input documents in sparse format. Finally, the use of stemming and lemmatization in data preprocessing provides several benefits. Doing so can improve training and inference compute time since it reduces the effective vocabulary size. More importantly, though, it can improve the quality of learned topic-word probability matrices and inferred topic mixtures. For example, the words *"parliament"*, *"parliaments"*, *"parliamentary"*, *"parliament's"*, and *"parliamentarians"* are all essentially the same word, *"parliament"*, but with different conjugations. For the purposes of detecting topics, such as a *"politics"* or *governments"* topic, the inclusion of all five does not add much additional value as they all essentially describe the same feature.
In this example, we will use a simple lemmatizer from [`nltk`](https://www.nltk.org/) package and use `CountVectorizer` in `scikit-learn` to perform the token counting. For more details please refer to their documentation respectively. Alternatively, [`spaCy`](https://spacy.io/) also offers easy-to-use tokenization and lemmatization functions.
---
In the following cell, we use a tokenizer and a lemmatizer from `nltk`. In the list comprehension, we implement a simple rule: only consider words that are longer than 2 characters, start with a letter and match the `token_pattern`.
```
import nltk
nltk.download("punkt")
nltk.download("wordnet")
from nltk import word_tokenize
from nltk.stem import WordNetLemmatizer
import re
token_pattern = re.compile(r"(?u)\b\w\w+\b")
class LemmaTokenizer(object):
def __init__(self):
self.wnl = WordNetLemmatizer()
def __call__(self, doc):
return [
self.wnl.lemmatize(t)
for t in word_tokenize(doc)
if len(t) >= 2 and re.match("[a-z].*", t) and re.match(token_pattern, t)
]
```
With the tokenizer defined we perform token counting next while limiting the vocabulary size to `vocab_size`. Use a maximum document frequency of 95% of documents (`max_df=0.95`) and a minimum document frequency of 2 documents (`min_df=2`).
```
import time
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer
vocab_size = 2000
print("Tokenizing and counting, this may take a few minutes...")
start_time = time.time()
vectorizer = CountVectorizer(
input="content",
analyzer="word",
stop_words="english",
tokenizer=LemmaTokenizer(),
max_features=vocab_size,
max_df=0.95,
min_df=2,
)
vectors = vectorizer.fit_transform(data)
vocab_list = vectorizer.get_feature_names()
print("vocab size:", len(vocab_list))
# random shuffle
idx = np.arange(vectors.shape[0])
np.random.shuffle(idx)
vectors = vectors[idx]
print("Done. Time elapsed: {:.2f}s".format(time.time() - start_time))
```
Optionally, we may consider removing very short documents, the following cell removes documents shorter than 25 words. This certainly depends on the application, but there are also some general justifications. It is hard to imagine very short documents express more than one topic. Topic modeling tries to model each document as a mixture of multiple topics, thus it may not be the best choice for modeling short documents.
```
threshold = 25
vectors = vectors[
np.array(vectors.sum(axis=1) > threshold).reshape(
-1,
)
]
print("removed short docs (<{} words)".format(threshold))
print(vectors.shape)
```
The output from `CountVectorizer` are sparse matrices with their elements being integers.
```
print(type(vectors), vectors.dtype)
print(vectors[0])
```
Because all the parameters (weights and biases) in the NTM model are `np.float32` type we'd need the input data to also be in `np.float32`. It is better to do this type-casting upfront rather than repeatedly casting during mini-batch training.
```
import scipy.sparse as sparse
vectors = sparse.csr_matrix(vectors, dtype=np.float32)
print(type(vectors), vectors.dtype)
```
As a common practice in modeling training, we should have a training set, a validation set, and a test set. The training set is the set of data the model is actually being trained on. But what we really care about is not the model's performance on training set but its performance on future, unseen data. Therefore, during training, we periodically calculate scores (or losses) on the validation set to validate the performance of the model on unseen data. By assessing the model's ability to generalize we can stop the training at the optimal point via early stopping to avoid over-training.
Note that when we only have a training set and no validation set, the NTM model will rely on scores on the training set to perform early stopping, which could result in over-training. Therefore, we recommend always supply a validation set to the model.
Here we use 80% of the data set as the training set and the rest for validation set and test set. We will use the validation set in training and use the test set for demonstrating model inference.
```
n_train = int(0.8 * vectors.shape[0])
# split train and test
train_vectors = vectors[:n_train, :]
test_vectors = vectors[n_train:, :]
# further split test set into validation set (val_vectors) and test set (test_vectors)
n_test = test_vectors.shape[0]
val_vectors = test_vectors[: n_test // 2, :]
test_vectors = test_vectors[n_test // 2 :, :]
print(train_vectors.shape, test_vectors.shape, val_vectors.shape)
```
---
## Store Data on S3
The NTM algorithm, as well as other first-party SageMaker algorithms, accepts data in [RecordIO](https://mxnet.apache.org/api/python/io/io.html#module-mxnet.recordio) [Protobuf](https://developers.google.com/protocol-buffers/) format. The SageMaker Python API provides helper functions for easily converting your data into this format. Below we convert the from numpy/scipy data and upload it to an Amazon S3 destination for the model to access it during training.
### Setup AWS Credentials
We first need to specify data locations and access roles. In particular, we need the following data:
- The S3 `bucket` and `prefix` that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
- The IAM `role` is used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
```
import os
import sagemaker
from sagemaker import get_execution_role
role = get_execution_role()
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = "20newsgroups"
train_prefix = os.path.join(prefix, "train")
val_prefix = os.path.join(prefix, "val")
output_prefix = os.path.join(prefix, "output")
s3_train_data = os.path.join("s3://", bucket, train_prefix)
s3_val_data = os.path.join("s3://", bucket, val_prefix)
output_path = os.path.join("s3://", bucket, output_prefix)
print("Training set location", s3_train_data)
print("Validation set location", s3_val_data)
print("Trained model will be saved at", output_path)
```
Here we define a helper function to convert the data to RecordIO Protobuf format and upload it to S3. In addition, we will have the option to split the data into several parts specified by `n_parts`.
The algorithm inherently supports multiple files in the training folder ("channel"), which could be very helpful for large data set. In addition, when we use distributed training with multiple workers (compute instances), having multiple files allows us to distribute different portions of the training data to different workers conveniently.
Inside this helper function we use `write_spmatrix_to_sparse_tensor` function provided by [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) to convert scipy sparse matrix into RecordIO Protobuf format.
```
# update sagemake package, in order to use write_spmatrix_to_sparse_tensor in the next cell
# !pip install -U sagemaker
def split_convert_upload(sparray, bucket, prefix, fname_template="data_part{}.pbr", n_parts=2):
import io
import boto3
import sagemaker.amazon.common as smac
chunk_size = sparray.shape[0] // n_parts
for i in range(n_parts):
# Calculate start and end indices
start = i * chunk_size
end = (i + 1) * chunk_size
if i + 1 == n_parts:
end = sparray.shape[0]
# Convert to record protobuf
buf = io.BytesIO()
smac.write_spmatrix_to_sparse_tensor(array=sparray[start:end], file=buf, labels=None)
buf.seek(0)
# Upload to s3 location specified by bucket and prefix
fname = os.path.join(prefix, fname_template.format(i))
boto3.resource("s3").Bucket(bucket).Object(fname).upload_fileobj(buf)
print("Uploaded data to s3://{}".format(os.path.join(bucket, fname)))
split_convert_upload(
train_vectors, bucket=bucket, prefix=train_prefix, fname_template="train_part{}.pbr", n_parts=8
)
split_convert_upload(
val_vectors, bucket=bucket, prefix=val_prefix, fname_template="val_part{}.pbr", n_parts=1
)
```
---
# Model Training
We have created the training and validation data sets and uploaded them to S3. Next, we configure a SageMaker training job to use the NTM algorithm on the data we prepared
SageMaker uses Amazon Elastic Container Registry (ECR) docker container to host the NTM training image. The following ECR containers are currently available for SageMaker NTM training in different regions. For the latest Docker container registry please refer to [Amazon SageMaker: Common Parameters](https://docs.aws.amazon.com/sagemaker/latest/dg/sagemaker-algo-docker-registry-paths.html).
```
import boto3
from sagemaker.image_uris import retrieve
container = retrieve("ntm", boto3.Session().region_name)
```
The code in the cell below automatically chooses an algorithm container based on the current region. In the API call to `sagemaker.estimator.Estimator` we also specify the type and count of instances for the training job. Because the 20NewsGroups data set is relatively small, we have chosen a CPU only instance (`ml.c4.xlarge`), but do feel free to change to [other instance types](https://aws.amazon.com/sagemaker/pricing/instance-types/). NTM fully takes advantage of GPU hardware and in general trains roughly an order of magnitude faster on a GPU than on a CPU. Multi-GPU or multi-instance training further improves training speed roughly linearly if communication overhead is low compared to compute time.
```
import sagemaker
sess = sagemaker.Session()
ntm = sagemaker.estimator.Estimator(
container,
role,
instance_count=2,
instance_type="ml.c4.xlarge",
output_path=output_path,
sagemaker_session=sess,
)
```
## Hyperparameters
Here we highlight a few hyperparameters. For information about the full list of available hyperparameters, please refer to [NTM Hyperparameters](https://docs.aws.amazon.com/sagemaker/latest/dg/ntm_hyperparameters.html).
- **feature_dim** - the "feature dimension", it should be set to the vocabulary size
- **num_topics** - the number of topics to extract
- **mini_batch_size** - this is the batch size for each worker instance. Note that in multi-GPU instances, this number will be further divided by the number of GPUs. Therefore, for example, if we plan to train on an 8-GPU machine (such as `ml.p2.8xlarge`) and wish each GPU to have 1024 training examples per batch, `mini_batch_size` should be set to 8196.
- **epochs** - the maximal number of epochs to train for, training may stop early
- **num_patience_epochs** and **tolerance** controls the early stopping behavior. Roughly speaking, the algorithm will stop training if within the last `num_patience_epochs` epochs there have not been improvements on validation loss. Improvements smaller than `tolerance` will be considered non-improvement.
- **optimizer** and **learning_rate** - by default we use `adadelta` optimizer and `learning_rate` does not need to be set. For other optimizers, the choice of an appropriate learning rate may require experimentation.
```
num_topics = 20
ntm.set_hyperparameters(
num_topics=num_topics,
feature_dim=vocab_size,
mini_batch_size=128,
epochs=100,
num_patience_epochs=5,
tolerance=0.001,
)
```
Next, we need to specify how the training data and validation data will be distributed to the workers during training. There are two modes for data channels:
- `FullyReplicated`: all data files will be copied to all workers
- `ShardedByS3Key`: data files will be sharded to different workers, i.e. each worker will receive a different portion of the full data set.
At the time of writing, by default, the Python SDK will use `FullyReplicated` mode for all data channels. This is desirable for validation (test) channel but not suitable for training channel. The reason is that when we use multiple workers we would like to go through the full data set by each of them going through a different portion of the data set, so as to provide different gradients within epochs. Using `FullyReplicated` mode on training data not only results in slower training time per epoch (nearly 1.5X in this example), but also defeats the purpose of distributed training. To set the training data channel correctly we specify `distribution` to be `ShardedByS3Key` for the training data channel as follows.
```
from sagemaker.inputs import TrainingInput
s3_train = TrainingInput(s3_train_data, distribution="ShardedByS3Key")
```
Now we are ready to train. The following cell takes a few minutes to run. The command below will first provision the required hardware. You will see a series of dots indicating the progress of the hardware provisioning process. Once the resources are allocated, training logs will be displayed. With multiple workers, the log color and the ID following `INFO` identifies logs emitted by different workers.
```
ntm.fit({"train": s3_train, "validation": s3_val_data})
```
If you see the message
> `===== Job Complete =====`
at the bottom of the output logs then that means training successfully completed and the output NTM model was stored in the specified output path. You can also view information about and the status of a training job using the AWS SageMaker console. Just click on the "Jobs" tab and select training job matching the training job name, below:
```
print("Training job name: {}".format(ntm.latest_training_job.job_name))
```
# Model Hosting and Inference
A trained NTM model does nothing on its own. We now want to use the model we computed to perform inference on data. For this example, that means predicting the topic mixture representing a given document.
We create an inference endpoint using the SageMaker Python SDK `deploy()` function from the job we defined above. We specify the instance type where inference is computed as well as an initial number of instances to spin up.
```
from sagemaker.serializers import CSVSerializer
from sagemaker.deserializers import JSONDeserializer
ntm_predictor = ntm.deploy(
initial_instance_count=1,
instance_type="ml.m4.xlarge",
serializer=CSVSerializer(),
deserializer=JSONDeserializer(),
)
```
Congratulations! You now have a functioning SageMaker NTM inference endpoint. You can confirm the endpoint configuration and status by navigating to the "Endpoints" tab in the AWS SageMaker console and selecting the endpoint matching the endpoint name, below:
```
print("Endpoint name: {}".format(ntm_predictor.endpoint_name))
```
Let's pass 5 examples from the test set to the inference endpoint
```
test_data = np.array(test_vectors.todense())
results = ntm_predictor.predict(test_data[:5])
print(results)
```
We can see the output format of SageMaker NTM inference endpoint is a Python dictionary with the following format.
```
{
'predictions': [
{'topic_weights': [ ... ] },
{'topic_weights': [ ... ] },
{'topic_weights': [ ... ] },
...
]
}
```
We extract the topic weights, themselves, corresponding to each of the input documents.
```
predictions = np.array([prediction["topic_weights"] for prediction in results["predictions"]])
print(predictions)
```
---
### Inference with RecordIO Protobuf
The inference endpoint also supports JSON-formatted and RecordIO Protobuf, see [Common Data Formats—Inference](https://docs.aws.amazon.com/sagemaker/latest/dg/cdf-inference.html) for more information.
At the time of writing SageMaker Python SDK does not yet have a RecordIO Protobuf serializer, but it is fairly straightforward to create one as follows.
```
def recordio_protobuf_serializer(spmatrix):
import io
import sagemaker.amazon.common as smac
buf = io.BytesIO()
smac.write_spmatrix_to_sparse_tensor(array=spmatrix, file=buf, labels=None)
buf.seek(0)
return buf
```
If you decide to compare these results to the known topic weights generated above keep in mind that SageMaker NTM discovers topics in no particular order. That is, the approximate topic mixtures computed above may be (approximate) permutations of the known topic mixtures corresponding to the same documents.
---
Now we can take a look at how the 20 topics are assigned to the 5 test documents with a bar plot.
```
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
fs = 12
df = pd.DataFrame(predictions.T)
df.plot(kind="bar", figsize=(16, 4), fontsize=fs)
plt.ylabel("Topic assignment", fontsize=fs + 2)
plt.xlabel("Topic ID", fontsize=fs + 2)
```
## Stop / Close the Endpoint
Finally, we should delete the endpoint before we close the notebook.
To restart the endpoint you can follow the code above using the same `endpoint_name` we created or you can navigate to the "Endpoints" tab in the SageMaker console, select the endpoint with the name stored in the variable `endpoint_name`, and select "Delete" from the "Actions" dropdown menu.
```
sagemaker.Session().delete_endpoint(ntm_predictor.endpoint_name)
```
# Model Exploration
***Note: The following section is meant as a deeper dive into exploring the trained models. The demonstrated functionalities may not be fully supported or guaranteed. For example, the parameter names may change without notice.***
The trained model artifact is a compressed package of MXNet models from the two workers. To explore the model, we first need to install mxnet.
```
# If you use conda_mxnet_p36 kernel, mxnet is already installed, otherwise, uncomment the following line to install.
# !pip install mxnet
import mxnet as mx
```
Here we download unpack the artifact
```
model_path = os.path.join(output_prefix, ntm._current_job_name, "output/model.tar.gz")
model_path
boto3.resource("s3").Bucket(bucket).download_file(model_path, "downloaded_model.tar.gz")
!pwd
import tarfile
tarfile.open("downloaded_model.tar.gz").extractall()
import zipfile
with zipfile.ZipFile("model_algo-1", "r") as zip_ref:
zip_ref.extractall("./")
```
We can load the model parameters and extract the weight matrix $W$ in the decoder as follows
```
model = mx.ndarray.load("params")
W = model["arg:projection_weight"]
```
Matrix $W$ corresponds to the $W$ in the NTM digram at the beginning of this notebook. Each column of $W$ corresponds to a learned topic. The elements in the columns of $W$ corresponds to the pseudo-probability of a word within a topic. We can visualize each topic as a word cloud with the size of each word be proportional to the pseudo-probability of the words appearing under each topic.
```
import sys
!{sys.executable} -m pip install wordcloud
import wordcloud as wc
import matplotlib.pyplot as plt
word_to_id = dict()
for i, v in enumerate(vocab_list):
word_to_id[v] = i
limit = 24
n_col = 4
counter = 0
plt.figure(figsize=(20, 16))
for ind in range(num_topics):
if counter >= limit:
break
title_str = "Topic{}".format(ind)
# pvals = mx.nd.softmax(W[:, ind]).asnumpy()
pvals = mx.nd.softmax(mx.nd.array(W[:, ind])).asnumpy()
word_freq = dict()
for k in word_to_id.keys():
i = word_to_id[k]
word_freq[k] = pvals[i]
wordcloud = wc.WordCloud(background_color="white").fit_words(word_freq)
plt.subplot(limit // n_col, n_col, counter + 1)
plt.imshow(wordcloud, interpolation="bilinear")
plt.axis("off")
plt.title(title_str)
# plt.close()
counter += 1
```
| github_jupyter |
```
%load_ext watermark
%watermark -v -p numpy,sklearn,scipy,matplotlib,tensorflow
```
**11장 – 심층 신경망 훈련**
_이 노트북은 11장에 있는 모든 샘플 코드를 가지고 있습니다._
# 설정
파이썬 2와 3을 모두 지원합니다. 공통 모듈을 임포트하고 맷플롯립 그림이 노트북 안에 포함되도록 설정하고 생성한 그림을 저장하기 위한 함수를 준비합니다:
```
# 파이썬 2와 파이썬 3 지원
from __future__ import division, print_function, unicode_literals
# 공통
import numpy as np
import os
# 일관된 출력을 위해 유사난수 초기화
def reset_graph(seed=42):
tf.reset_default_graph()
tf.set_random_seed(seed)
np.random.seed(seed)
# 맷플롯립 설정
%matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# 한글출력
plt.rcParams['font.family'] = 'NanumBarunGothic'
plt.rcParams['axes.unicode_minus'] = False
# 그림을 저장할 폴더
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
def save_fig(fig_id, tight_layout=True):
path = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id + ".png")
if tight_layout:
plt.tight_layout()
plt.savefig(path, format='png', dpi=300)
```
# 그래디언트 소실/폭주 문제
```
def logit(z):
return 1 / (1 + np.exp(-z))
z = np.linspace(-5, 5, 200)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [1, 1], 'k--')
plt.plot([0, 0], [-0.2, 1.2], 'k-')
plt.plot([-5, 5], [-3/4, 7/4], 'g--')
plt.plot(z, logit(z), "b-", linewidth=2)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('수렴', xytext=(3.5, 0.7), xy=(5, 1), arrowprops=props, fontsize=14, ha="center")
plt.annotate('수렴', xytext=(-3.5, 0.3), xy=(-5, 0), arrowprops=props, fontsize=14, ha="center")
plt.annotate('선형', xytext=(2, 0.2), xy=(0, 0.5), arrowprops=props, fontsize=14, ha="center")
plt.grid(True)
plt.title("로지스틱 활성화 함수", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("sigmoid_saturation_plot")
plt.show()
```
## Xavier와 He 초기화
```
import tensorflow as tf
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
he_init = tf.variance_scaling_initializer()
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
kernel_initializer=he_init, name="hidden1")
```
## 수렴하지 않는 활성화 함수
### Leaky ReLU
```
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
plt.plot(z, leaky_relu(z, 0.05), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([0, 0], [-0.5, 4.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('통과', xytext=(-3.5, 0.5), xy=(-5, -0.2), arrowprops=props, fontsize=14, ha="center")
plt.title("Leaky ReLU 활성화 함수", fontsize=14)
plt.axis([-5, 5, -0.5, 4.2])
save_fig("leaky_relu_plot")
plt.show()
```
텐서플로에서 Leaky ReLU 구현하기:
_텐서플로 1.4에서 tf.nn.leaky_\__relu(z, alpha) 함수가 추가되었습니다._
```
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
def leaky_relu(z, name=None):
return tf.maximum(0.01 * z, z, name=name)
hidden1 = tf.layers.dense(X, n_hidden1, activation=leaky_relu, name="hidden1")
```
Leaky ReLU를 사용하여 신경망을 훈련시켜 보죠. 먼저 그래프를 정의합니다:
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=leaky_relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=leaky_relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
```
데이터를 로드합니다:
주의: `tf.examples.tutorials.mnist`은 삭제될 예정이므로 대신 `tf.keras.datasets.mnist`를 사용하겠습니다.
```
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train = X_train.astype(np.float32).reshape(-1, 28*28) / 255.0
X_test = X_test.astype(np.float32).reshape(-1, 28*28) / 255.0
y_train = y_train.astype(np.int32)
y_test = y_test.astype(np.int32)
X_valid, X_train = X_train[:5000], X_train[5000:]
y_valid, y_train = y_train[:5000], y_train[5000:]
def shuffle_batch(X, y, batch_size):
rnd_idx = np.random.permutation(len(X))
n_batches = len(X) // batch_size
for batch_idx in np.array_split(rnd_idx, n_batches):
X_batch, y_batch = X[batch_idx], y[batch_idx]
yield X_batch, y_batch
# from tensorflow.examples.tutorials.mnist import input_data
# mnist = input_data.read_data_sets("/tmp/data/")
n_epochs = 40
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
if epoch % 5 == 0:
acc_batch = accuracy.eval(feed_dict={X: X_batch, y: y_batch})
acc_valid = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "배치 데이터 정확도:", acc_batch, "검증 세트 정확도:", acc_valid)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
### ELU
```
def elu(z, alpha=1):
return np.where(z < 0, alpha * (np.exp(z) - 1), z)
plt.plot(z, elu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1, -1], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"ELU 활성화 함수 ($\alpha=1$)", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("elu_plot")
plt.show()
```
텐서플로에서 ELU를 구현하는 것은 간단합니다. 층을 구성할 때 활성화 함수에 지정하기만 하면 됩니다:
```
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.elu, name="hidden1")
```
### SELU
이 활성화 함수는 Günter Klambauer, Thomas Unterthiner, Andreas Mayr가 2017년에 쓴 [논문](https://arxiv.org/pdf/1706.02515.pdf)에서 소개되었습니다(나중에 책에 추가하겠습니다). 훈련할 때 SELU 활성화 함수를 사용한 완전 연결 신경망은 스스로 정규화를 합니다. 각 층의 출력은 훈련하는 동안 같은 평균과 분산을 유지하려는 경향이 있어 그래디언트 소실과 폭주 문제를 해결합니다. 이 활성화 함수는 심층 신경망에서 다른 활성화 함수보다 뛰어난 성능을 내므로 꼭 이 함수를 시도해봐야 합니다.
```
def selu(z,
scale=1.0507009873554804934193349852946,
alpha=1.6732632423543772848170429916717):
return scale * elu(z, alpha)
plt.plot(z, selu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1.758, -1.758], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"SELU 활성화 함수", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("selu_plot")
plt.show()
```
기본적으로 SELU 하이퍼파라미터(`scale`과 `alpha`)는 평균이 0, 표준 편차가 1에 가깝게 유지되도록 조정합니다(입력도 평균이 0, 표준 편차가 1로 표준화되었다고 가정합니다). 이 활성화 함수를 사용하면 100층으로 된 심층 신경망도 그래디언트 소실/폭주 문제없이 모든 층에서 대략 평균이 0이고 표준 편차가 1을 유지합니다:
```
np.random.seed(42)
Z = np.random.normal(size=(500, 100))
for layer in range(100):
W = np.random.normal(size=(100, 100), scale=np.sqrt(1/100))
Z = selu(np.dot(Z, W))
means = np.mean(Z, axis=1)
stds = np.std(Z, axis=1)
if layer % 10 == 0:
print("층 {}: {:.2f} < 평균 < {:.2f}, {:.2f} < 표준 편차 < {:.2f}".format(
layer, means.min(), means.max(), stds.min(), stds.max()))
```
텐서플로 1.4 버전에 `tf.nn.selu()` 함수가 추가되었습니다. 이전 버전을 사용할 때는 다음 구현을 사용합니다:
```
def selu(z,
scale=1.0507009873554804934193349852946,
alpha=1.6732632423543772848170429916717):
return scale * tf.where(z >= 0.0, z, alpha * tf.nn.elu(z))
```
하지만 SELU 활성화 함수는 일반적인 드롭아웃과 함께 사용할 수 없습니다(드롭아웃은 SELU 활성화 함수의 자동 정규화 기능을 없애버립니다). 다행히 같은 논문에 실린 알파 드롭아웃(Alpha Dropout)을 사용할 수 있습니다. 텐서플로 1.4에 `tf.contrib.nn.alpha_dropout()`이 추가되었습니다(Linz 대학교 생물정보학 연구소(Institute of Bioinformatics)의 Johannes Kepler가 만든 [구현](https://github.com/bioinf-jku/SNNs/blob/master/selu.py)을 확인해 보세요).
SELU 활성화 함수를 사용한 신경망을 만들어 MNIST 문제를 풀어 보겠습니다:
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=selu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=selu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 40
batch_size = 50
```
이제 훈련할 차례입니다. 입력을 평균 0, 표준 편차 1로 스케일 조정해야 합니다:
```
means = X_train.mean(axis=0, keepdims=True)
stds = X_train.std(axis=0, keepdims=True) + 1e-10
X_val_scaled = (X_valid - means) / stds
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
X_batch_scaled = (X_batch - means) / stds
sess.run(training_op, feed_dict={X: X_batch_scaled, y: y_batch})
if epoch % 5 == 0:
acc_batch = accuracy.eval(feed_dict={X: X_batch_scaled, y: y_batch})
acc_valid = accuracy.eval(feed_dict={X: X_val_scaled, y: y_valid})
print(epoch, "배치 데이터 정확도:", acc_batch, "검증 세트 정확도:", acc_valid)
save_path = saver.save(sess, "./my_model_final_selu.ckpt")
```
# 배치 정규화
각 은닉층의 활성화 함수 전에 배치 정규화를 추가하기 위해 ELU 활성화 함수를 배치 정규화 층 이후에 수동으로 적용하겠습니다.
노트: `tf.layers.dense()` 함수가 (책에서 사용하는) `tf.contrib.layers.arg_scope()`와 호환되지 않기 때문에 대신 파이썬의 `functools.partial()` 함수를 사용합니다. 이를 사용해 `tf.layers.dense()`에 필요한 매개변수가 자동으로 설정되도록 `my_dense_layer()`를 만듭니다(그렇지 않으면 `my_dense_layer()`를 호출할 때마다 덮어씌여질 것입니다). 다른 코드는 이전과 비슷합니다.
```
reset_graph()
import tensorflow as tf
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 100
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
training = tf.placeholder_with_default(False, shape=(), name='training')
hidden1 = tf.layers.dense(X, n_hidden1, name="hidden1")
bn1 = tf.layers.batch_normalization(hidden1, training=training, momentum=0.9)
bn1_act = tf.nn.elu(bn1)
hidden2 = tf.layers.dense(bn1_act, n_hidden2, name="hidden2")
bn2 = tf.layers.batch_normalization(hidden2, training=training, momentum=0.9)
bn2_act = tf.nn.elu(bn2)
logits_before_bn = tf.layers.dense(bn2_act, n_outputs, name="outputs")
logits = tf.layers.batch_normalization(logits_before_bn, training=training,
momentum=0.9)
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
training = tf.placeholder_with_default(False, shape=(), name='training')
```
같은 매개변수를 계속 반복해서 쓰지 않도록 파이썬의 `partial()` 함수를 사용합니다:
```
from functools import partial
my_batch_norm_layer = partial(tf.layers.batch_normalization,
training=training, momentum=0.9)
hidden1 = tf.layers.dense(X, n_hidden1, name="hidden1")
bn1 = my_batch_norm_layer(hidden1)
bn1_act = tf.nn.elu(bn1)
hidden2 = tf.layers.dense(bn1_act, n_hidden2, name="hidden2")
bn2 = my_batch_norm_layer(hidden2)
bn2_act = tf.nn.elu(bn2)
logits_before_bn = tf.layers.dense(bn2_act, n_outputs, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
```
각 층에 ELU 활성화 함수와 배치 정규화를 사용하여 MNIST를 위한 신경망을 만듭니다:
```
reset_graph()
batch_norm_momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
training = tf.placeholder_with_default(False, shape=(), name='training')
with tf.name_scope("dnn"):
he_init = tf.variance_scaling_initializer()
my_batch_norm_layer = partial(
tf.layers.batch_normalization,
training=training,
momentum=batch_norm_momentum)
my_dense_layer = partial(
tf.layers.dense,
kernel_initializer=he_init)
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
bn1 = tf.nn.elu(my_batch_norm_layer(hidden1))
hidden2 = my_dense_layer(bn1, n_hidden2, name="hidden2")
bn2 = tf.nn.elu(my_batch_norm_layer(hidden2))
logits_before_bn = my_dense_layer(bn2, n_outputs, name="outputs")
logits = my_batch_norm_layer(logits_before_bn)
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
```
노트: 배치 정규화를 위해 별도의 업데이트 연산을 실행해 주어야 합니다(`sess.run([training_op, extra_update_ops],...`).
```
n_epochs = 20
batch_size = 200
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run([training_op, extra_update_ops],
feed_dict={training: True, X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
어!? MNIST 정확도가 좋지 않네요. 물론 훈련을 더 오래하면 정확도가 높아지겠지만 이런 얕은 신경망에서는 배치 정규화와 ELU가 큰 효과를 내지 못합니다. 대부분 심층 신경망에서 빛을 발합니다.
업데이트 연산에 의존하는 훈련 연산을 만들 수도 있습니다:
```python
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
extra_update_ops = tf.get_collection(tf.GraphKeys.UPDATE_OPS)
with tf.control_dependencies(extra_update_ops):
training_op = optimizer.minimize(loss)
```
이렇게 하면 훈련할 때 `training_op`만 평가하면 텐서플로가 업데이트 연산도 자동으로 실행할 것입니다:
```python
sess.run(training_op, feed_dict={training: True, X: X_batch, y: y_batch})
```
한가지 더, 훈련될 변수 개수가 전체 전역 변수 개수보다 적습니다. 이동 평균을 위한 변수는 훈련되는 변수가 아니기 때문입니다. 미리 학습한 신경망을 재사용할 경우(아래 참조) 이런 훈련되지 않는 변수를 놓쳐서는 안됩니다.
```
[v.name for v in tf.trainable_variables()]
[v.name for v in tf.global_variables()]
```
## 그래디언트 클리핑
MNIST를 위한 간단한 신경망을 만들고 그래디언트 클리핑을 적용해 보겠습니다. 시작 부분은 이전과 동일합니다(학습한 모델을 재사용하는 예를 만들기 위해 몇 개의 층을 더 추가했습니다. 아래 참조):
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_hidden3 = 50
n_hidden4 = 50
n_hidden5 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3")
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4")
hidden5 = tf.layers.dense(hidden4, n_hidden5, activation=tf.nn.relu, name="hidden5")
logits = tf.layers.dense(hidden5, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
learning_rate = 0.01
```
이제 그래디언트 클리핑을 적용합니다. 먼저 그래디언트를 구한 다음 `clip_by_value()` 함수를 사용해 클리핑하고 적용합니다:
```
threshold = 1.0
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var)
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
```
나머지는 이전과 동일합니다:
```
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
## 학습된 모델 재사용하기
## 텐서플로 모델 재사용하기
먼저 그래프 구조를 로드해야 합니다. `import_meta_graph()` 함수가 그래프 연산들을 로드하여 기본 그래프에 적재하고 모델의 상태를 복원할 수 있도록 `Saver` 객체를 반환합니다. 기본적으로 `Saver` 객체는 `.meta` 확장자를 가진 파일에 그래프 구조를 저장하므로 이 파일을 로드해야 합니다:
```
reset_graph()
saver = tf.train.import_meta_graph("./my_model_final.ckpt.meta")
```
다음으로 훈련해야 할 모든 연산을 가져와야 합니다. 그래프 구조를 모를 때는 모든 연산을 출력해 볼 수 있습니다:
```
for op in tf.get_default_graph().get_operations():
print(op.name)
```
웁스, 연산이 엄청 많네요! 텐서보드로 그래프를 시각화해보는 것이 더 좋을 것 같습니다. 다음 코드는 주피터에서 그래프를 그려줍니다(만약 브라우저에서 보이지 않는다면 `FileWriter`로 그래프를 저장한 다음 텐서보드에서 열어 보세요):
```
from tensorflow_graph_in_jupyter import show_graph
show_graph(tf.get_default_graph())
```
필요한 연산을 찾았다면 그래프의 `get_operation_by_name()`이나 `get_tensor_by_name()` 메서드를 사용하여 추출할 수 있습니다:
```
X = tf.get_default_graph().get_tensor_by_name("X:0")
y = tf.get_default_graph().get_tensor_by_name("y:0")
accuracy = tf.get_default_graph().get_tensor_by_name("eval/accuracy:0")
training_op = tf.get_default_graph().get_operation_by_name("GradientDescent")
```
원본 모델을 만들 때 다른 사람이 재사용하기 쉽게 연산에 명확한 이름을 부여하고 문서화를 하는 것이 좋습니다. 또 다른 방법은 처리해야 할 중요한 연산들을 모두 모아 놓은 컬렉션을 만드는 것입니다:
```
for op in (X, y, accuracy, training_op):
tf.add_to_collection("my_important_ops", op)
```
이렇게 하면 모델을 재사용할 때 다음과 같이 간단하게 쓸 수 있습니다:
```
X, y, accuracy, training_op = tf.get_collection("my_important_ops")
```
이제 세션을 시작하고 모델을 복원하여 준비된 훈련 데이터로 훈련을 계속할 수 있습니다:
```
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
# 모델 훈련 계속하기...
```
실제로 테스트를 해보죠!
```
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
```
또 다른 방법으로 원본 그래프를 만든 파이썬 코드에 접근할 수 있다면 `import_meta_graph()`를 대신 사용할 수 있습니다:
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_hidden3 = 50
n_hidden4 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3")
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4")
hidden5 = tf.layers.dense(hidden4, n_hidden5, activation=tf.nn.relu, name="hidden5")
logits = tf.layers.dense(hidden5, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
threshold = 1.0
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
grads_and_vars = optimizer.compute_gradients(loss)
capped_gvs = [(tf.clip_by_value(grad, -threshold, threshold), var)
for grad, var in grads_and_vars]
training_op = optimizer.apply_gradients(capped_gvs)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
```
그 다음 훈련을 계속할 수 있습니다:
```
with tf.Session() as sess:
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
```
일반적으로 하위층만 재사용할 것입니다. `import_meta_graph()`를 사용하면 전체 그래프를 로드하지만 필요하지 않은 부분은 무시하면 됩니다. 이 예에서는 학습된 3번째 층 위에 4번째 은닉층을 새로 추가합니다(원래 4번째 층은 무시됩니다). 새로운 출력층도 추가하고 이 출력으로 손실을 계산하고 이를 최소화하기 위한 새로운 옵티마이저를 만듭니다. 전체 그래프(원본 그래프 전체와 새로운 연산)를 저장할 새로운 `Saver` 객체와 새로운 모든 변수를 초기화할 초기화 연산도 필요합니다:
```
reset_graph()
n_hidden4 = 20 # 새 층
n_outputs = 10 # 새 층
saver = tf.train.import_meta_graph("./my_model_final.ckpt.meta")
X = tf.get_default_graph().get_tensor_by_name("X:0")
y = tf.get_default_graph().get_tensor_by_name("y:0")
hidden3 = tf.get_default_graph().get_tensor_by_name("dnn/hidden4/Relu:0")
new_hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="new_hidden4")
new_logits = tf.layers.dense(new_hidden4, n_outputs, name="new_outputs")
with tf.name_scope("new_loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=new_logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("new_eval"):
correct = tf.nn.in_top_k(new_logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("new_train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
new_saver = tf.train.Saver()
```
새로운 모델을 훈련시킵니다:
```
with tf.Session() as sess:
init.run()
saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = new_saver.save(sess, "./my_new_model_final.ckpt")
```
원본 모델을 만든 파이썬 코드에 접근할 수 있다면 필요한 부분만 재사용하고 나머지는 버릴 수 있습니다:
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1") # 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2") # 재사용
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3") # 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
```
그러나 이전에 학습된 모델을 복원하기 위해 (복원할 변수 리스트를 전달합니다. 그렇지 않으면 그래프와 맞지 않는다고 에러를 낼 것입니다) `Saver` 객체를 하나 만들고 훈련이 끝난 후 새로운 모델을 저장하기 위해 또 다른 `Saver` 객체를 만들어야 합니다:
```
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs): # 책에는 없음
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size): # 책에는 없음
sess.run(training_op, feed_dict={X: X_batch, y: y_batch}) # 책에는 없음
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_new_model_final.ckpt")
```
## 다른 프레임워크의 모델 재사용하기
이 예에서는 재사용하려는 각 변수에 대해 변수 초기화 할당 연산을 찾고, 초기화 될 값에 해당하는 두 번째 입력 핸들을 구합니다. 초기화가 실행될 때 여기에 `feed_dict` 매개변수를 사용하여 초깃값 대신 원하는 값을 주입합니다:
```
reset_graph()
n_inputs = 2
n_hidden1 = 3
original_w = [[1., 2., 3.], [4., 5., 6.]] # 다른 프레임워크로부터 가중치를 로드
original_b = [7., 8., 9.] # 다른 프레임워크로부터 편향을 로드
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
# [...] 모델의 나머지 부분을 구성
# hidden1 변수의 할당 노드에 대한 핸들을 구합니다
graph = tf.get_default_graph()
assign_kernel = graph.get_operation_by_name("hidden1/kernel/Assign")
assign_bias = graph.get_operation_by_name("hidden1/bias/Assign")
init_kernel = assign_kernel.inputs[1]
init_bias = assign_bias.inputs[1]
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init, feed_dict={init_kernel: original_w, init_bias: original_b})
# [...] 새 작업에 모델을 훈련시킵니다
print(hidden1.eval(feed_dict={X: [[10.0, 11.0]]})) # 책에는 없음
```
또 다른 방법은 전용 할당 노드와 플레이스홀더를 만든는 것입니다. 이 방법은 더 번거롭고 효율적이지 않지만 하려는 방식이 잘 드러나는 방법입니다:
```
reset_graph()
n_inputs = 2
n_hidden1 = 3
original_w = [[1., 2., 3.], [4., 5., 6.]] # 다른 프레임워크로부터 가중치를 로드
original_b = [7., 8., 9.] # 다른 프레임워크로부터 편향을 로드
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
# [...] 모델의 나머지를 구성
# hidden1 변수의 할당 노드에 대한 핸들을 구합니다
with tf.variable_scope("", default_name="", reuse=True): # 루트 범위
hidden1_weights = tf.get_variable("hidden1/kernel")
hidden1_biases = tf.get_variable("hidden1/bias")
# 전용 플레이스홀더와 할당 노드를 만듭니다
original_weights = tf.placeholder(tf.float32, shape=(n_inputs, n_hidden1))
original_biases = tf.placeholder(tf.float32, shape=n_hidden1)
assign_hidden1_weights = tf.assign(hidden1_weights, original_weights)
assign_hidden1_biases = tf.assign(hidden1_biases, original_biases)
init = tf.global_variables_initializer()
with tf.Session() as sess:
sess.run(init)
sess.run(assign_hidden1_weights, feed_dict={original_weights: original_w})
sess.run(assign_hidden1_biases, feed_dict={original_biases: original_b})
# [...] 새 작업에 모델을 훈련시킵니다
print(hidden1.eval(feed_dict={X: [[10.0, 11.0]]}))
```
`get_collection()`에 `scope`를 지정하여 변수의 핸들을 가져올 수도 있습니다:
```
tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES, scope="hidden1")
```
또는 그래프의 `get_tensor_by_name()` 메서드를 사용할 수 있습니다:
```
tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
tf.get_default_graph().get_tensor_by_name("hidden1/bias:0")
```
### 하위층 동결하기
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1") # 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2") # 재사용
hidden3 = tf.layers.dense(hidden2, n_hidden3, activation=tf.nn.relu, name="hidden3") # 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu, name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"): # 책에는 없음
optimizer = tf.train.GradientDescentOptimizer(learning_rate) # 책에는 없음
train_vars = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES,
scope="hidden[34]|outputs")
training_op = optimizer.minimize(loss, var_list=train_vars)
init = tf.global_variables_initializer()
new_saver = tf.train.Saver()
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
name="hidden1") # 동결층 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
name="hidden2") # 동결층 재사용
hidden2_stop = tf.stop_gradient(hidden2)
hidden3 = tf.layers.dense(hidden2_stop, n_hidden3, activation=tf.nn.relu,
name="hidden3") # 동결하지 않고 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu,
name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
```
훈련하는 코드는 이전과 완전히 동일합니다:
```
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_new_model_final.ckpt")
```
### 동결층 캐싱하기
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300 # 재사용
n_hidden2 = 50 # 재사용
n_hidden3 = 50 # 재사용
n_hidden4 = 20 # 새로 만듦!
n_outputs = 10 # 새로 만듦!
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
name="hidden1") # 동결층 재사용
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
name="hidden2") # 동결층 재사용 & 캐싱
hidden2_stop = tf.stop_gradient(hidden2)
hidden3 = tf.layers.dense(hidden2_stop, n_hidden3, activation=tf.nn.relu,
name="hidden3") # 동결하지 않고 재사용
hidden4 = tf.layers.dense(hidden3, n_hidden4, activation=tf.nn.relu,
name="hidden4") # 새로 만듦!
logits = tf.layers.dense(hidden4, n_outputs, name="outputs") # 새로 만듦!
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
reuse_vars = tf.get_collection(tf.GraphKeys.GLOBAL_VARIABLES,
scope="hidden[123]") # 정규 표현식
restore_saver = tf.train.Saver(reuse_vars) # 1-3층 복원
init = tf.global_variables_initializer()
saver = tf.train.Saver()
import numpy as np
n_batches = len(X_train) // batch_size
with tf.Session() as sess:
init.run()
restore_saver.restore(sess, "./my_model_final.ckpt")
h2_cache = sess.run(hidden2, feed_dict={X: X_train})
h2_cache_valid = sess.run(hidden2, feed_dict={X: X_valid}) # 책에는 없음
for epoch in range(n_epochs):
shuffled_idx = np.random.permutation(len(X_train))
hidden2_batches = np.array_split(h2_cache[shuffled_idx], n_batches)
y_batches = np.array_split(y_train[shuffled_idx], n_batches)
for hidden2_batch, y_batch in zip(hidden2_batches, y_batches):
sess.run(training_op, feed_dict={hidden2:hidden2_batch, y:y_batch})
accuracy_val = accuracy.eval(feed_dict={hidden2: h2_cache_valid, # 책에는 없음
y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_new_model_final.ckpt")
```
# 고속 옵티마이저
## 모멘텀 옵티마이저
```
optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate,
momentum=0.9)
```
## 네스테로프 가속 경사
```
optimizer = tf.train.MomentumOptimizer(learning_rate=learning_rate,
momentum=0.9, use_nesterov=True)
```
## AdaGrad
```
optimizer = tf.train.AdagradOptimizer(learning_rate=learning_rate)
```
## RMSProp
```
optimizer = tf.train.RMSPropOptimizer(learning_rate=learning_rate,
momentum=0.9, decay=0.9, epsilon=1e-10)
```
## Adam 최적화
```
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
```
## 학습률 스케줄링
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
with tf.name_scope("train"): # 책에는 없음
initial_learning_rate = 0.1
decay_steps = 10000
decay_rate = 1/10
global_step = tf.Variable(0, trainable=False, name="global_step")
learning_rate = tf.train.exponential_decay(initial_learning_rate, global_step,
decay_steps, decay_rate)
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum=0.9)
training_op = optimizer.minimize(loss, global_step=global_step)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 5
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
# 규제로 과대적합 피하기
## $\ell_1$과 $\ell_2$ 규제
$\ell_1$ 규제를 직접 구현해 보죠. 먼저 평상시처럼 모델을 만듭니다(간단하게 하기 위해 은닉층을 하나만 두겠습니다):
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
logits = tf.layers.dense(hidden1, n_outputs, name="outputs")
```
그다음, 층의 가중치에 대한 핸들을 얻어 크로스 엔트로피 손실에 $\ell_1$ 손실(즉, 가중치의 절댓값)을 더해 전체 손실을 계산합니다:
```
W1 = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
W2 = tf.get_default_graph().get_tensor_by_name("outputs/kernel:0")
scale = 0.001 # l1 규제 하이퍼파라미터
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y,
logits=logits)
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy")
reg_losses = tf.reduce_sum(tf.abs(W1)) + tf.reduce_sum(tf.abs(W2))
loss = tf.add(base_loss, scale * reg_losses, name="loss")
```
나머지는 이전과 동일합니다:
```
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
다른 방법으로는 `tf.layers.dense()` 함수에 규제 함수를 전달할 수 있습니다. 이 함수는 규제 손실을 계산하기 위한 연산을 만들고 규제 손실 컬렉션에 이 연산을 추가합니다. 모델 선언부는 이전과 동일합니다:
```
reset_graph()
n_inputs = 28 * 28 # MNIST
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
```
그다음, 동일한 매개변수를 매번 반복하지 않으려고 파이썬의 `partial()` 함수를 사용합니다. `kernel_regularizer` 매개변수를 지정해야 합니다:
```
scale = 0.001
my_dense_layer = partial(
tf.layers.dense, activation=tf.nn.relu,
kernel_regularizer=tf.contrib.layers.l1_regularizer(scale))
with tf.name_scope("dnn"):
hidden1 = my_dense_layer(X, n_hidden1, name="hidden1")
hidden2 = my_dense_layer(hidden1, n_hidden2, name="hidden2")
logits = my_dense_layer(hidden2, n_outputs, activation=None,
name="outputs")
```
기본 손실에 규제 손실을 추가합니다:
```
with tf.name_scope("loss"): # 책에는 없음
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits( # 책에는 없음
labels=y, logits=logits) # 책에는 없음
base_loss = tf.reduce_mean(xentropy, name="avg_xentropy") # 책에는 없음
reg_losses = tf.get_collection(tf.GraphKeys.REGULARIZATION_LOSSES)
loss = tf.add_n([base_loss] + reg_losses, name="loss")
```
나머지는 평상시와 동일합니다:
```
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32), name="accuracy")
learning_rate = 0.01
with tf.name_scope("train"):
optimizer = tf.train.GradientDescentOptimizer(learning_rate)
training_op = optimizer.minimize(loss)
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 200
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
## 드롭아웃
```
reset_graph()
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
training = tf.placeholder_with_default(False, shape=(), name='training')
dropout_rate = 0.5 # == 1 - keep_prob
X_drop = tf.layers.dropout(X, dropout_rate, training=training)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X_drop, n_hidden1, activation=tf.nn.relu,
name="hidden1")
hidden1_drop = tf.layers.dropout(hidden1, dropout_rate, training=training)
hidden2 = tf.layers.dense(hidden1_drop, n_hidden2, activation=tf.nn.relu,
name="hidden2")
hidden2_drop = tf.layers.dropout(hidden2, dropout_rate, training=training)
logits = tf.layers.dense(hidden2_drop, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum=0.9)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
n_epochs = 20
batch_size = 50
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={training: True, X: X_batch, y: y_batch})
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid})
print(epoch, "검증 세트 정확도:", accuracy_val)
save_path = saver.save(sess, "./my_model_final.ckpt")
```
## 맥스 노름
2개의 은닉층을 가진 간단한 MNIST 신경망을 만들어 보겠습니다:
```
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
```
다음으로 첫 번째 은닉층의 가중치에 대한 핸들을 얻고 `clip_by_norm()` 함수를 사용해 가중치를 클리핑하는 연산을 만듭니다. 그런 다음 클리핑된 가중치를 가중치 변수에 할당하는 연산을 만듭니다:
```
threshold = 1.0
weights = tf.get_default_graph().get_tensor_by_name("hidden1/kernel:0")
clipped_weights = tf.clip_by_norm(weights, clip_norm=threshold, axes=1)
clip_weights = tf.assign(weights, clipped_weights)
```
두 번째 층에 대해서도 동일하게 할 수 있습니다:
```
weights2 = tf.get_default_graph().get_tensor_by_name("hidden2/kernel:0")
clipped_weights2 = tf.clip_by_norm(weights2, clip_norm=threshold, axes=1)
clip_weights2 = tf.assign(weights2, clipped_weights2)
```
초기와 연산과 `Saver` 객체를 만듭니다:
```
init = tf.global_variables_initializer()
saver = tf.train.Saver()
```
이제 모델을 훈련시킵니다. 이전과 매우 동일한데 `training_op`을 실행한 후에 `clip_weights`와 `clip_weights2` 연산을 실행하는 것만 다릅니다:
```
n_epochs = 20
batch_size = 50
with tf.Session() as sess: # 책에는 없음
init.run() # 책에는 없음
for epoch in range(n_epochs): # 책에는 없음
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size): # 책에는 없음
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
clip_weights.eval()
clip_weights2.eval() # 책에는 없음
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_model_final.ckpt") # 책에는 없음
```
위 구현은 이해하기 쉽고 잘 작동하지만 조금 번거롭습니다. 더 나은 방법은 `max_norm_regularizer()` 함수를 만드는 것입니다:
```
def max_norm_regularizer(threshold, axes=1, name="max_norm",
collection="max_norm"):
def max_norm(weights):
clipped = tf.clip_by_norm(weights, clip_norm=threshold, axes=axes)
clip_weights = tf.assign(weights, clipped, name=name)
tf.add_to_collection(collection, clip_weights)
return None # 규제 손실을 위한 항이 없습니다
return max_norm
```
그런 다음 (필요한 임계값을 지정해서) 맥스 노름 규제 매개변수에 넘길 함수를 만들기 위해 이 함수를 호출합니다. 은닉층을 만들 때 이 규제 함수를 `kernel_regularizer` 매개변수를 통해 전달할 수 있습니다:
```
reset_graph()
n_inputs = 28 * 28
n_hidden1 = 300
n_hidden2 = 50
n_outputs = 10
learning_rate = 0.01
momentum = 0.9
X = tf.placeholder(tf.float32, shape=(None, n_inputs), name="X")
y = tf.placeholder(tf.int32, shape=(None), name="y")
max_norm_reg = max_norm_regularizer(threshold=1.0)
with tf.name_scope("dnn"):
hidden1 = tf.layers.dense(X, n_hidden1, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden1")
hidden2 = tf.layers.dense(hidden1, n_hidden2, activation=tf.nn.relu,
kernel_regularizer=max_norm_reg, name="hidden2")
logits = tf.layers.dense(hidden2, n_outputs, name="outputs")
with tf.name_scope("loss"):
xentropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=logits)
loss = tf.reduce_mean(xentropy, name="loss")
with tf.name_scope("train"):
optimizer = tf.train.MomentumOptimizer(learning_rate, momentum)
training_op = optimizer.minimize(loss)
with tf.name_scope("eval"):
correct = tf.nn.in_top_k(logits, y, 1)
accuracy = tf.reduce_mean(tf.cast(correct, tf.float32))
init = tf.global_variables_initializer()
saver = tf.train.Saver()
```
훈련 연산이 실행된 후에 가중치 클리핑 연산을 실행하는 것을 제외하면 이전과 동일합니다:
```
n_epochs = 20
batch_size = 50
clip_all_weights = tf.get_collection("max_norm")
with tf.Session() as sess:
init.run()
for epoch in range(n_epochs):
for X_batch, y_batch in shuffle_batch(X_train, y_train, batch_size):
sess.run(training_op, feed_dict={X: X_batch, y: y_batch})
sess.run(clip_all_weights)
accuracy_val = accuracy.eval(feed_dict={X: X_valid, y: y_valid}) # 책에는 없음
print(epoch, "검증 세트 정확도:", accuracy_val) # 책에는 없음
save_path = saver.save(sess, "./my_model_final.ckpt") # 책에는 없음
```
# 연습문제 해답
11장의 연습문제는 [11_deep_learning_exercise](11_deep_learning_exercises.ipynb) 노트북에 있습니다.
| github_jupyter |
# A1: core module notebook
# Introduction
This notebook includes a description of the 'core' python module in the JBEI Quantitative Metabolic Modeling (QMM) library. A description and demonstration of the diffent classes can be found below.
# Setup
First, we need ot set the path and environment variable properly:
```
%matplotlib inline
import sys, os
pythonPath = "/scratch/david.ando/quantmodel/code/core"
if pythonPath not in sys.path:
sys.path.append('/scratch/david.ando/quantmodel/code/core')
os.environ["QUANTMODELPATH"] = '/scratch/david.ando/quantmodel'
```
Importing the required modules for the demo:
```
from IPython.display import Image
import core, FluxModels
import os
```
# Classes description
## Metabolite related classes
### metbolite class
The *metabolite* class is used to store all information related to a metabolite. For example the following instantation:
```
ala = core.Metabolite('ala-L', ncarbons=3, source=True, feed='100% 1-C', destination=False, formula='C3H7NO2')
```
creates a metabolite with nbame 'ala-L', 3 carbon atoms, which is the source of labeling, is labeled in the first carbon,
is not a destination (measured) metabolite and with a composition formula equal to 'C3H7NO2'
the **generateEMU** function creates the corresponding Elementary Metabolite Unit (EMU):
```
ala.generateEMU([2])
```
In this case the EMU contains the first and last carbon in alanine. The input ([2]) specifies which carbons to exclude:
```
ala.generateEMU([2,3])
```
### reactant and product classes
*Reactant* and *product* are classes derived from metabolite and the only difference is that they represent metabolites in the context of a reaction. Hence, the stoichiometry of the metabolite and the labeling pattern in that reaction are included:
```
R_ala = core.Reactant(ala, 1, 'abc')
```
Notice that the stoichiometry information (1, meaning in the reaction only 1 molecule participates in the reaction) and the labeling data ('abc', one part of the labeling pattern, see below) only make sense in the context of a reaction, so they are not included in the metabolite class.
Both classes are derived from metabolites, so they inherit their methods:
```
R_ala.generateEMU([2,3])
```
## Reaction related classes
### reaction class
The *reaction* class produces a reaction instance:
```
# Create reactant metabolites
coa_c = core.Metabolite('coa_c')
nad_c = core.Metabolite('nad_c')
pyr_c = core.Metabolite('pyr_c')
# Convert into reactants
Rcoa_c = core.Reactant(coa_c, 1.0)
Rnad_c = core.Reactant(nad_c, 1.0)
Rpyr_c = core.Reactant(pyr_c, 1.0)
# Create product metabolites
accoa_c = core.Metabolite('accoa_c')
co2_c = core.Metabolite('co2_c')
nadh_c = core.Metabolite('nadh_c')
# Convert into products
Raccoa_c = core.Product(accoa_c, 1.0)
Rco2_c = core.Product(co2_c, 1.0)
Rnadh_c = core.Product(nadh_c, 1.0)
# Create reaction
PDH = core.Reaction('PDH',reactants=[Rcoa_c,Rnad_c,Rpyr_c] , products=[Raccoa_c,Rco2_c,Rnadh_c]
,subsystem='S_GlycolysisGluconeogenesis')
```
Reactions can also initialized from a string:
```
PDH2 = core.Reaction.from_string('PDH : coa_c + nad_c + pyr_c --> accoa_c + co2_c + nadh_c ')
```
The *reaction* class contains some useful functions such as:
**stoichLine** to obtain the stoichiometric line for the reaction:
```
print PDH.stoichLine()
print PDH2.stoichLine()
```
**getReactDict** produces a dictionary of reactants:
```
PDH.getReactDict()
```
**getProdDict** produces a dictionary of products:
```
PDH.getProdDict()
```
## Elementary Metabolite Unit (EMU) related classes
Elementary Metabolite Units (or EMUs) of a compound are the molecule parts (moieties) comprising any distinct subset of the compound’s atoms (Antoniewicz MR, Kelleher JK, Stephanopoulos G: Elementary metabolite units (EMU): a novel framework for modeling isotopic distributions. Metab Eng 2007, 9:68-86.). For example, cit$_{123}$ represents the first 3 carbon atoms in the citrate molecule.
### EMU class
The EMU class provides a class to hold and manipulate EMUs:
```
cit321= core.EMU('cit_3_2_1')
```
The method **findnCarbons** produces the number of carbons in the EMU:
```
print cit321.findnCarbons()
```
The method **getMetName** produces the name of the corresponding metabolite:
```
print cit321.getMetName()
str(cit321.getMetName()) == 'cit'
```
The method **getIndices** produces the indices:
```
print cit321.getIndices()
```
**getSortedName** sorts the indices in the EMU name:
```
print cit321.getSortedName()
```
**getEmuInSBML** produces the name of the EMU in SBML format:
```
print cit321.getEmuInSBML()
```
## Transitions related classes
Transitions contain the information on how carbon (or other) atoms are passed in each reaction. Atom transitions describe, for example, the fate of each carbon in a reaction, whereas EMU transitions describe this information by using EMUs, as described below.
### AtomTransition class
Atom transitions represent the fate of each carbon in a reaction (Wiechert W. (2001) 13C metabolic flux analysis. Metabolic engineering 3: 195-206). For example, in:
AKGDH akg --> succoa + co2 abcde : bcde + a
akg gets split into succoa and co2, with the first 4 carbons going to succoa and the remaining carbon going to co2.
```
AT = core.AtomTransition('AKGDH akg --> succoa + co2 abcde : bcde + a')
print AT
```
The method **findEMUtransition** provides for a given input EMU (e.g. succoa_1_2_3_4), which EMU it comes from in the form of a EMU transition:
```
emu1 = core.EMU('co2_1')
print AT.findEMUtransition(emu1)
emu2 = core.EMU('succoa_1_2_3_4')
print AT.findEMUtransition(emu2)
```
This is done through the method **findEMUs**, which finds the emus from which the input emanates in the given atom transition:
```
print emu2.name
print AT.findEMUs(emu2)
for emus in AT.findEMUs(emu2):
for emu_ in emus:
print emu_.name
```
which in turn, uses the method **getOriginDictionary** which provides for a given input EMU the originating metabolite and the correspondance in indices:
```
AT.getOriginDictionary(emu2)
```
### EMUTransition class
Class for EMU transitions that contain information on how different EMUs transform intto each other. For example:
TA1_b, TAC3_c_1_2_3 + g3p_c_1_2_3 --> f6p_c_1_2_3_4_5_6
indicating that TAC3_c_1_2_3 and g3p_c_1_2_3 combine to produce f6p_c_1_2_3_4_5_6 in reaction TA1_b (backward reaction of TA1), or:
SSALy, (0.5) sucsal_c_4 --> (0.5) succ_c_4
which indicates that the fourth atom of sucsal_c becomes the fourth atom of succ_c. The (0.5) contribution coefficient indicates that reaction SSALy contains a symmetric molecule and two labeling correspondences are equally likely. Hence this transition only contributes half the flux to the final labeling.
```
emuTrans = core.EMUTransition('TA1_b, TAC3_c_1_2_3 + g3p_c_1_2_3 --> f6p_c_1_2_3_4_5_6')
print emuTrans
str(emuTrans) == 'TA1_b, TAC3_c_1_2_3 + g3p_c_1_2_3 --> f6p_c_1_2_3_4_5_6'
```
## Ranged number class
The *rangedNumber* class describes floating point numbers for which a confidence interval is available. For example, fluxes obtained through 2S-$^{13}$C MFA are described through the flux that best fits the data and the highest and lowest values that are found to be compatible with labeling data (see equations 16-23 in Garcia Martin *et al* 2015). However, this class has been abstracted out so it can be used with other ranged intervals. Ranged numbers can used as follows:
```
number = core.rangedNumber(0.3,0.6,0.9) # 0.3 lowest, 0.6 best fit, 0.9 highest
```
Ranged numbers can be printed:
```
print number
```
and added, substracted, multiplied and divided following the standard error propagation rules(https://en.wikipedia.org/wiki/Propagation_of_uncertainty):
```
A = core.rangedNumber(0.3,0.6,0.9)
B = core.rangedNumber(0.1,0.15,0.18)
print A+B
print A-B
print 2*A
print B/3
```
## Flux class
The flux class describes fluxes attached to a reaction. For example, if the net flux is described by the ranged number A and the exchange flux by the ranged number B, the corresponding flux would be:
```
netFlux = A
exchangeFlux = B
flux1 = core.flux(net_exc_tup=(netFlux,exchangeFlux))
print flux1
```
Fluxes can easily multiplied:
```
print 3*flux1
```
| github_jupyter |
# IPython: beyond plain Python
When executing code in IPython, all valid Python syntax works as-is, but IPython provides a number of features designed to make the interactive experience more fluid and efficient.
## First things first: running code, getting help
In the notebook, to run a cell of code, hit `Shift-Enter`. This executes the cell and puts the cursor in the next cell below, or makes a new one if you are at the end. Alternately, you can use:
- `Alt-Enter` to force the creation of a new cell unconditionally (useful when inserting new content in the middle of an existing notebook).
- `Control-Enter` executes the cell and keeps the cursor in the same cell, useful for quick experimentation of snippets that you don't need to keep permanently.
```
print("Hi")
```
Getting help:
```
?
```
Typing `object_name?` will print all sorts of details about any object, including docstrings, function definition lines (for call arguments) and constructor details for classes.
```
import collections
collections.namedtuple?
collections.Counter??
*int*?
```
An IPython quick reference card:
```
%quickref
```
## Tab completion
Tab completion, especially for attributes, is a convenient way to explore the structure of any object you’re dealing with. Simply type `object_name.<TAB>` to view the object’s attributes. Besides Python objects and keywords, tab completion also works on file and directory names.
```
collections.
```
## The interactive workflow: input, output, history
```
2+10
_+10
```
You can suppress the storage and rendering of output if you append `;` to the last cell (this comes in handy when plotting with matplotlib, for example):
```
10+20;
_
```
The output is stored in `_N` and `Out[N]` variables:
```
_10 == Out[10]
```
And the last three have shorthands for convenience:
```
from __future__ import print_function
print('last output:', _)
print('next one :', __)
print('and next :', ___)
In[11]
_i
_ii
print('last input:', _i)
print('next one :', _ii)
print('and next :', _iii)
%history -n 1-5
```
**Exercise**
Write the last 10 lines of history to a file named `log.py`.
## Accessing the underlying operating system
```
!pwd
files = !ls
print("My current directory's files:")
print(files)
!echo $files
!echo {files[0].upper()}
```
Note that all this is available even in multiline blocks:
```
import os
for i,f in enumerate(files):
if f.endswith('ipynb'):
!echo {"%02d" % i} - "{os.path.splitext(f)[0]}"
else:
print('--')
```
## Beyond Python: magic functions
The IPyhton 'magic' functions are a set of commands, invoked by prepending one or two `%` signs to their name, that live in a namespace separate from your normal Python variables and provide a more command-like interface. They take flags with `--` and arguments without quotes, parentheses or commas. The motivation behind this system is two-fold:
- To provide an orthogonal namespace for controlling IPython itself and exposing other system-oriented functionality.
- To expose a calling mode that requires minimal verbosity and typing while working interactively. Thus the inspiration taken from the classic Unix shell style for commands.
```
%magic
```
Line vs cell magics:
```
%timeit list(range(1000))
%%timeit
list(range(10))
list(range(100))
```
Line magics can be used even inside code blocks:
```
for i in range(1, 5):
size = i*100
print('size:', size, end=' ')
%timeit list(range(size))
```
Magics can do anything they want with their input, so it doesn't have to be valid Python:
```
%%bash
echo "My shell is:" $SHELL
echo "My disk usage is:"
df -h
```
Another interesting cell magic: create any file you want locally from the notebook:
```
%%writefile test.txt
This is a test file!
It can contain anything I want...
And more...
!cat test.txt
```
Let's see what other magics are currently defined in the system:
```
%lsmagic
```
## Running normal Python code: execution and errors
Not only can you input normal Python code, you can even paste straight from a Python or IPython shell session:
```
>>> # Fibonacci series:
... # the sum of two elements defines the next
... a, b = 0, 1
>>> while b < 10:
... print(b)
... a, b = b, a+b
In [1]: for i in range(10):
...: print(i, end=' ')
...:
```
And when your code produces errors, you can control how they are displayed with the `%xmode` magic:
```
%%writefile mod.py
def f(x):
return 1.0/(x-1)
def g(y):
return f(y+1)
```
Now let's call the function `g` with an argument that would produce an error:
```
import mod
mod.g(0)
%xmode plain
mod.g(0)
%xmode verbose
mod.g(0)
```
The default `%xmode` is "context", which shows additional context but not all local variables. Let's restore that one for the rest of our session.
```
%xmode context
```
## Running code in other languages with special `%%` magics
```
%%perl
@months = ("July", "August", "September");
print $months[0];
%%ruby
name = "world"
puts "Hello #{name.capitalize}!"
```
## Raw Input in the notebook
Since 1.0 the IPython notebook web application support `raw_input` which for example allow us to invoke the `%debug` magic in the notebook:
```
mod.g(0)
%debug
```
Don't forget to exit your debugging session. Raw input can of course be used to ask for user input:
```
enjoy = input('Are you enjoying this tutorial? ')
print('enjoy is:', enjoy)
```
## Plotting in the notebook
This magic configures matplotlib to render its figures inline:
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 2*np.pi, 300)
y = np.sin(x**2)
plt.plot(x, y)
plt.title("A little chirp")
fig = plt.gcf() # let's keep the figure object around for later...
```
## The IPython kernel/client model
```
%connect_info
```
We can automatically connect a Qt Console to the currently running kernel with the `%qtconsole` magic, or by typing `ipython console --existing <kernel-UUID>` in any terminal:
```
%qtconsole
```
| github_jupyter |
# Cryptarithms: Send More Money
The July, 1924, issue of the famous British magazine *The Strand* included a word puzzle by Henry E. Dudeney in his regular contribution "Perplexities". The puzzle is to assign a unique digit to each letter appearing in the equation
S E N D
+ M O R E
= M O N E Y
such that the arithmetic equation is satisfied, and the leading digit for M is non-zero. There are [many more examples](http://cryptarithms.awardspace.us/puzzles.html) of these puzzles, but this is perhaps the most well-known.
This notebook demonstrates a solution to this puzzle using Pyomo disjuctions and the `gecode` solver, a constraint solving package written in C++. This same puzzle is used in the `gecode` documentation, so this notebook may provide a useful contrast between Pyomo modeling and use of a native C++ API.
```
# install Pyomo and solvers for Google Colab
import sys
if "google.colab" in sys.modules:
!wget -N -q https://raw.githubusercontent.com/jckantor/MO-book/main/tools/install_on_colab.py
%run install_on_colab.py
```
## Modeling and Solution
There are several possible approaches to modeling this puzzle in Pyomo.
[One approach](https://stackoverflow.com/questions/67456379/pyomo-model-constraint-programming-for-sendmore-money-task) would be to using a matrix of binary variables $x_{a,d}$ indexed by letter $a$ and digit $d$ such that $x_{a,d} = 1$ designates the corresponding assignment. The problem constraints can then be implemented by summing the binary variables along the two axes. The arithmetic constraint becomes a more challenging.
[Another approach](https://www.gecode.org/doc/6.0.1/MPG.pdf) is to use Pyomo integer variables indexed by letters, then setup an linear expression to represent the puzzle. If we use the notation $n_a$ to represent the digit assigned to letter $a$, the algebraic constraint becomes
$$
\begin{align*}
1000 n_s + 100 n_e + 10 n_n + n_d \\
+ 1000 n_m + 100 n_o + 10 n_r + n_e \\
= 10000 n_m + 1000 n_o + 100 n_n + 10 n_e + n_y
\end{align*}
$$
The requirement that no two letters be assigned the same digit can be represented as a \ disjunction. Letting $n_a$ and $n_b$ denote the integers assigned to letters $a$ and $b$, the disjuction becomes
$$
\begin{align*}
\begin{bmatrix}n_a \lt n_b\end{bmatrix}
\ \veebar\ &
\begin{bmatrix}n_b \lt n_a\end{bmatrix}
& \forall a \lt b
\end{align*}$$
```
import pyomo.environ as pyo
import pyomo.gdp as gdp
m = pyo.ConcreteModel()
m.LETTERS = pyo.Set(initialize=['S', 'E', 'N', 'D', 'M', 'O', 'R', 'Y'])
m.PAIRS = pyo.Set(initialize=m.LETTERS * m.LETTERS, filter = lambda m, a, b: a < b)
m.n = pyo.Var(m.LETTERS, domain=pyo.Integers, bounds=(0, 9))
@m.Constraint()
def message(m):
return 1000*m.n['S'] + 100*m.n['E'] + 10*m.n['N'] + m.n['D'] \
+ 1000*m.n['M'] + 100*m.n['O'] + 10*m.n['R'] + m.n['E'] \
== 10000*m.n['M'] + 1000*m.n['O'] + 100*m.n['N'] + 10*m.n['E'] + m.n['Y']
# leading digit must be non-zero
@m.Constraint()
def leading_digit_nonzero(m):
return m.n['M'] >= 1
# assign a different number to each letter
@m.Disjunction(m.PAIRS)
def unique_assignment(m, a, b):
return [m.n[a] >= m.n[b] + 1, m.n[b] >= m.n[a] + 1]
# assign a "dummy" objective to avoid solver errors
@m.Objective()
def dummy_objective(m):
return m.n['M']
pyo.TransformationFactory('gdp.bigm').apply_to(m)
solver = pyo.SolverFactory('cbc')
solver.solve(m)
def letters2num(s):
return ' '.join(map(lambda s: f"{int(m.n[s]())}", list(s)))
print(" ", letters2num('SEND'))
print(" + ", letters2num('MORE'))
print(" ----------")
print("= ", letters2num('MONEY'))
```
## Suggested exercises
1. Pyomo includes a logic-based solver `GDPopt` for generalized disjunctive programming problems. Implement and test `GDPopt` using combinations of solution strategies and MIP solvers. Compare the performance of `GDPopt` to the constraint solver `gecode`.
2. There are [many more examples](http://cryptarithms.awardspace.us/puzzles.html) this puzzles. Refactor this code and create a function that can be used to solve generic puzzles of this type.
| github_jupyter |
#### Assignment 11
**Sai Gauthami Kuravi**
<ol>
<li>Construct, train and test a neural network to predict high utilization in the next one year using recurrent neural networks. </li>
<li>You should use tensorflow library to do so</li>
<li>Please submit python notebook and html version of the output showing results. </li>
</ol>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from tensorflow import keras
from tensorflow import feature_column
from tensorflow.keras import layers
from sklearn.metrics import accuracy_score,roc_auc_score
import os
os.getcwd()
os.chdir(r'C:\Users\skura\Desktop')
df = pd.read_csv('exlix_binary_claims_w_dates.csv')
back_window = 20 # this is number of claims
shift = 1 # this is number of claims
# forward window defined in terms of days
forward_window = 365
#
# split in training/testing by patient
#
from sklearn.model_selection import train_test_split
pts = df['patient_id'].unique()
trp, tsp = train_test_split(pts)
tr = df[df['patient_id'].isin(trp)]
ts = df[df['patient_id'].isin(tsp)]
x_tr = [] # inputs
y_tr = [] # outputs
for p in trp[:10000]:
pt = tr[tr['patient_id'] == p]
for t in range(back_window, len(pt)):
# input variables
x_tr.append(pt[t-back_window:t].iloc[:,2:])
# output variable
t_days = pt.iloc[t].claim_days_cum
y_tr.append(len(pt[(pt['claim_days_cum'] > t_days) & (pt['claim_days_cum'] <= t_days + 365) ]))
#stop looping if past year 2
if pt.iloc[t].claim_days_cum > 730:
break
x_ts = [] # inputs
y_ts = [] # outputs
for p in tsp[:1000]:
pt = ts[ts['patient_id'] == p]
for t in range(back_window, len(pt)):
# input variables
x_ts.append(pt[t-back_window:t].iloc[:,2:])
# output variable
t_days = pt.iloc[t].claim_days_cum
y_ts.append(len(pt[(pt['claim_days_cum'] > t_days) & (pt['claim_days_cum'] <= t_days + 365) ]))
#stop looping if past year 2
if pt.iloc[t].claim_days_cum > 730:
break
y_ts = (np.array(y_ts) >= 100).astype('int')
y_ts
x_tr_vals = np.array([xx.values for xx in x_tr]).astype('float32')
x_ts_vals = np.array([xx.values for xx in x_ts]).astype('float32')
y_tr = (np.array(y_tr) >= 100).astype('int')
import tensorflow as tf
elix_lstm_model = tf.keras.models.Sequential([
tf.keras.layers.LSTM(29, input_shape=(20,29)),
tf.keras.layers.Dense(1, activation='sigmoid')
])
elix_lstm_model.compile(optimizer='adam', loss='binary_crossentropy')
elix_lstm_model.fit(x_tr_vals, y_tr, epochs=40,
steps_per_epoch=None)
x_ts_vals.shape
x_tr_vals.shape
probs = elix_lstm_model.predict_proba(x_ts_vals)
probs
results = elix_lstm_model.evaluate(x_ts_vals, y_ts, batch_size=128)
results
```
| github_jupyter |
```
import xgboost as xgb
import pandas as pd
from sklearn.metrics import f1_score
train_features = pd.read_csv('dataset1_train_features.csv')
test_features = pd.read_csv('dataset1_test_features.csv')
train_types = []
for row in train_features['Type']:
if row == 'Class':
train_types.append(1)
else:
train_types.append(0)
train_features['Type_encode'] = train_types
test_types = []
for row in test_features['Type']:
if row == 'Class':
test_types.append(1)
else:
test_types.append(0)
test_features['Type_encode'] = test_types
X_train = train_features.loc[:, 'Ngram1_Entity':'Type_encode']
y_train = train_features['Match']
X_test = test_features.loc[:, 'Ngram1_Entity':'Type_encode']
y_test = test_features['Match']
df_train = train_features.loc[:, 'Ngram1_Entity':'Type_encode']
df_train['Match'] = train_features['Match']
df_test = test_features.loc[:, 'Ngram1_Entity':'Type_encode']
df_test['Match'] = test_features['Match']
X_train = X_train.fillna(value=0)
X_test = X_test.fillna(value=0)
min_child_weight_s = [1, 5, 10]
gamma_s = [0.5, 1, 1.5, 2, 5]
subsample_s = [0.6, 0.8, 1.0]
colsample_bytree_s = [0.6, 0.8, 1.0]
max_depth_s = [3, 4, 5]
def f1_eval(y_pred, dtrain):
y_true = dtrain.get_label()
err = 1-f1_score(y_true, np.round(y_pred))
return 'f1_err', err
dtrain = xgb.DMatrix(X_train, label=y_train)
dtest = xgb.DMatrix(X_test, label=y_test)
data = []
for min_child_weight in min_child_weight_s:
for gamma in gamma_s:
for subsample in subsample_s:
for colsample_bytree in colsample_bytree_s:
for max_depth in max_depth_s:
param = {'silent': 0,
'objective': 'binary:logistic',
'min_child_weight': min_child_weight,
'gamma': gamma,
'subsample': subsample,
'colsample_bytree': colsample_bytree,
'max_depth': max_depth
}
param['nthread'] = 4
param['eval_metric'] = f1_eval
evallist = [(dtest, 'eval'), (dtrain, 'train')]
plst = param.items()
num_round = 10
bst = xgb.train(plst, dtrain, num_round, evallist, verbose_eval=False)
y_pred = bst.predict(dtest)
f1 = f1_score(y_test, y_pred)
print(min_child_weight, gamma, subsample, colsample_bytree, max_depth)
data.append((min_child_weight, gamma, subsample, colsample_bytree, max_depth, f1))
dataset = pd.DataFrame(data, columns=['min_child_weight', 'gamma', 'subsample',
'colsample_bytree', 'max_depth', 'f1'])
dataset.to_csv('dataset1_xgboost.csv', index=False)
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.