text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Networks and Human Behavior
## Tools for computational analysis
### TABLE OF CONTENTS
1. [Introduction](#What-you-need!)
2. [Network Representation](#first)
2.1 [Nodes and Edges](#NE)
2.2 [Graphs](#G)
2.3 [Adjacency Matrix](#A)
3. [Managing Network Data](#Data)
Note: this index and its links don't render properly on github but will work if you open it using jupyter.
***
<a id="What-you-need!"></a>
### Introduction
The empirical work in this course will be organized in a series of Jupyter Notebooks of which this is the first one.
- The notebooks will provide you with enough information to understand the concepts of the class and the tools to perform your own exploratory analysis.
- They are not exhaustive, meaning that there may be alternative methods to do the same tasks.
- They are work in progress, so kindly flag any issues you encounter or any suggestions for improvement.
<br>
<div class="alert alert-block alert-success">
You can vizualize the notebooks on your browser but you will get the most out of the material if you download and run them in Python as you go along.
<br>
<br>
If you have not done so yet, follow the setup instructions.
</div>
***
<a id="first"></a>
### Network Representation
<a id="NE"></a>
#### Nodes and Edges
Networks provide us with a mental model to describe and study interactions of individual entities within a community or group.
The basic elements of the network are thus:
- <u>The individuals</u>: e.g. humans, countries, banks. Abstractly referred to as **nodes** (N).
- <u>The relationships</u>: e.g. friendships, trade treaties, loans. Generically called **edges** (E).
A simple network is thus defined as a set of nodes (think for instance the group of students taking this class) and a description of whether each pair of nodes interacts with each other; we will sometimes talk about this interacting status as the pair being **connected**.
> Naturally, information regarding the same network (that is the same sets of nodes and edges) can be conveyed and stored in different forms.
>
> This is important to remember because it could determine how you handle network data; for instance it could influence which function you use to load a network into a notebook.
To fix some ideas let's consider a group of 5 people: Thomas, Carl, Sara, Mark and Sonia.
<br>
<div class="alert alert-block alert-info"> One of Python's convenient features is that it allows you to build programs based on objects. We will not enter in more detail for the purpose of this class but you will find "pre-programmed" objects such as <b>sets</b> and <b>lists</b> to be very useful.</div>
Let's define a set of our group of students:
```
students = {'Thomas', 'Carl', 'Sara', 'Mark','Sonia'}
# We can see the elements of the set of students by using the generic print() function from python
print(students)
# It's fairly simple to add another name,
# If you have downloaded this notebook and are runnning it as you read, feel free to change YOU for your own name
students.add("YOU")
# Note: there are many other things that can be done with sets. You can search online if you are curious.
print(students)
```
Now, let us fully characterize the network of **friendships** in the class by listing all the pairs of friends:
```
friend_list = [('Sara','Sonia'),
('Sara','Mark'),
('Sonia','Mark'),
('Sonia','Carl'),
('Sonia','Thomas'),
('Mark','Thomas')
]
```
Remark in a few details.
- In this simple network we consider that Sara being friends with Sonia implies that Sonia is friends with Sara. Such networks are known as **undirected** and thus we only need to list the edge between Sara and Sonia once and the ortehr of the elements of the pair is irrelevant.
- The converse type of networks are known as **directed** (simplest example would be Twitter) in which the relationship can be one-sided and thus `('Sara','Sonia')` would be represent a different edge from `('Sonia','Sara')` and both would have to be specified if both indeed appear in the directed network.
- In this classroom network not everyone is connected to everyone else and in particular you as the new student have not yet had the chance to make any connections. This shows why both sets (N and E) are important to fully describe the network. Nodes with no edges (also known as **isolated** nodes) do not appear anywhere on the edgelist.
- Nodes do not have edges with themselves. This is a simple convention that can sometimes be consequential and can be changed in specific contexts.
`students` and `friend_list` fully describe our simple network.
We know all the members of the community and who is friends with whom.
<a id="G"></a>
#### Graphs
Nodes and edges are naturally ameanable to be represented by a graph in which nodes are vertex or points and edges between them show the connections (you can get an idea of where some of the names come from).
Graphs are nice because they provide insightful visualizations but also because many mathematicians and theorists have spent time and effort studying and finding many useful properties some of which we shall cover in this course.
<br>
<div class="alert alert-block alert-info"> Not all objects and functions that we will use on Python are already loaded when you launch the interpreter (or open a notebook). There is a vast number of modules (or packages) that expand your ability to perform computations without having to code the functions from scratch. <br>
To use these functions and objects you have to import them. The best practice is to import them at the begining of your code and to use aliases defined by convention. <br>
Modules are imported by running:
<pre><code>import module_name as module_alias</code></pre>
The word following "as" is known as an alias and it serves as a prefix for when you call the functions of that package. If you import the package without an alias then you can just type the function names without any prefix but this is bad practice because some packages may have functions with the same name or you may have coded functions with already used names. <br>
<br>
In principle, you can choose any alias you want for each package. However, there are some conventions and it is easier if you stick to those.
</div>
In the subsequent notebooks this shall become clear. For now we will just import one package to work with networks (networkx).
In networkx, networks are usually defined as a graph object. Let's play with our example network.
```
import networkx as nx
# Define a graph object G
G=nx.Graph()
# Add nodes
G.add_nodes_from(students)
# Right now, the graph has all the students but no edges
print('# nodes:',G.number_of_nodes())
print(G.nodes())
print('# edges:',G.number_of_edges())
# Let's add the edges from the edge-list
G.add_edges_from(friend_list)
print('# edges:',G.number_of_edges())
# The next line of code allows to visualize plots within the notebook (in the future we will keep it in the preamble)
%matplotlib inline
# Let's take a look at the network
nx.draw_circular(G)
# Don't worry too much about all these extra arguments, it is just to visualize it better.
nx.draw_circular(G, with_labels = True,node_color='crimson',node_size=2500,font_color='white')
```
Now that you are getting so good at networking, you could probably add some edges with the other students.
```
G.add_edge('YOU','Sara')
G.add_edge('YOU','Thomas')
nx.draw_circular(G, with_labels = True,node_color='crimson',node_size=2500,font_color='white')
```
<a id="A"></a>
#### Adjacency Matrix
Another very useful representation for a network is its **adjacency matrix**.
To understand this matrix:
1. Choose an order for the nodes.
2. Consider every possible pair of nodes.
2.1 If the nodes in the pair are connected, input a 1 in the cell of the matrix whose indices correspond to the pair of nodes.
2.2 If they are not connected, input a zero.
For the pre-cooked example in this notebook it should look like this:
| / | Thomas | Mark | Carl | Sonia | Sara | YOU |
| --- | --- | --- | --- | --- | --- | --- |
| Thomas | 0 | 1 | 0 | 1 | 0 | 1 |
| Mark | 1 | 0 | 0 | 1 | 1 | 0 |
| Carl | 0 | 0 | 0 | 1 | 0 | 0 |
| Sonia | 1 | 1 | 1 | 0 | 1 | 0 |
| Sara | 0 | 1 | 0 | 1 | 0 | 1 |
| YOU | 1 | 0 | 0 | 0 | 1 | 0 |
```
# To get the matrix from your networkx Graph simply use nx.adjacency_matrix()
# Note: you can choose the order in which the nodes are indexed into rows and columns.
# Note 2: Python saves the matrix in a "sparse" format, meaning that it only stores the position of the ones
# and ignores the zeros. So to visualize a matrix like the one above we use the .todense() method
# on the matrix to "make it dense", that is to generate the zeros.
print(nx.adjacency_matrix(G,nodelist=['Thomas', 'Mark', 'Carl', 'Sonia', 'Sara', 'YOU']).todense())
```
<br>
<div class="alert alert-block alert-success"> Feel free to <b>play around with the toy network</b> in this notebook. <br>
<ul>
<li>Add/delete nodes. E.g. G.add_node('ME'), G.remove_node('YOU'). </li>
<li>Add/delete edges. E.g. G.add_edge('YOU','Carl'), G.remove_node('Carl','Sonia'). </li>
</ul>
See how this changes affect the edge list, the graph and the adjacency matrix.
<br>
<br>
<b>Make sure you undertand how the different representations work and relate to one another! </b> <br>
If unsure, review the material, ask your classmates and your instructors :)
</div>
***
<a id="Data"></a>
### Managing Network Data
As you go along with the course material (and in your own future research/work) you will come across network datasets that msy contain many more nodes and edges and you will not be creating the network from scratch as we did here.
The network data may be stored as a list of edges, a graph or an adjacency matrix and all of these could be stored in multiple formats (like .csv, .txt, .dta, .xlsx).
In this section we will provide a non-exhaustive list of ways in which you can get your data up an running on Python from an already existing network.
Throughout the course you may encounter some of these and other alternatives (and you may find even more ways to do it online). DO NOT worry about memorizing functions but rather about understanding the concepts as they relate to social and economic networks.
#### Dataframes
One simple way of handling data on Python is with the **Pandas** (pd) module. Pandas is one of the workhorses of data analysis in Python.
Among many other things, it allows you to create tables from data stored in a wide variety of formats.
Remeber that before being able to use it, you will need to load the module:
`import pandas as pd`
In this class the main way in which we will use it is to load/save data.
##### Load Data
We will create a new object, a dataframe, that contains data that was stored in a specific file in your computer.
Let's say that you have a .csv file in a given file: 'PATH_TO_FILE/file_name.csv'
Then way to read and load into a table is simply:
`new_dataframe = pd.read_csv('PATH_TO_FILE/file_name.csv')`
If the file format is not csv you may use other functions (this is not exhaustive):
| Format | function |
| --- | --- |
| excel | `pd.read_excel('PATH_TO_FILE/file_name.xlsx')`|
| stata | `pd.read_stata('PATH_TO_FILE/file_name.dta')`|
| parquet | `pd.read_parquet('PATH_TO_FILE/file_name.parquet')`|
| JSON | `pd.read_json('PATH_TO_FILE/file_name.json')`|
| txt | `pd.read_csv('PATH_TO_FILE/file_name.txt',sep=" ")`|
##### Save Data
When you already have a pandas dataframe (say edgelist_dataframe) then you can save it to different formats using the following fuctions:
| Format | function |
| --- | --- |
| csv | `edgelist_dataframe.to_csv('PATH_TO_FILE/file_name.csv')`|
| excel | `edgelist_dataframe.to_excel('PATH_TO_FILE/file_name.xlsx')`|
| stata | `edgelist_dataframe.to_stata('PATH_TO_FILE/file_name.dta')`|
| parquet | `edgelist_dataframe.to_parquet('PATH_TO_FILE/file_name.parquet')`|
| JSON | `edgelist_dataframe.to_json('PATH_TO_FILE/file_name.json')`|
| txt | `edgelist_dataframe.to_csv('PATH_TO_FILE/file_name.txt',sep=" ")`|
<br>
<div class="alert alert-block alert-info"> Note how in Python some functions are applied by typing the object as one of the arguments in parentheses: <br>
<pre><code>function(OBJECT,other arguments)</code></pre>
while other functions are applied using a '.' after the object: <br>
<pre><code>OBJECT.function(other arguments)</code></pre>
An in-depth explanation of this difference falls outside the scope of this course. For now it should be sufficient to know that the difference exists.
</div>
##### Dataframes $\iff$ Networks
Most commonly, the network dataframes you will be reading will correspond to node/edge lists and adjacency matrices.
- Once you have a pandas dataframe with the edgelist, define a network with:
> `new_graph_object_name = nx.from_pandas_edgelist(new_dataframe)`
- If you want to save the edge list of an existing network as a pandas dataframe then you would run:
> `old_edgelist_data = nx.to_pandas_edgelist(old_graph)`
- If the file contained an adjacency matrix (and therefore the dataframe is a matrix) then you would run:
> `new_graph_object_name = nx.from_pandas_adjacency(new_dataframe)`
- Finally (for now), to save the adjacency matrix of an existing as a pandas dataframe:
> `df = nx.to_pandas_adjacency(old_graph, dtype=int)`
See the example below for our toy example:
```
# since we havent used pandas yet we need to load it
# in the future, follow convention and load modules at the begining of your work
import pandas as pd
# generate pandas dataframe from edgelist
edgelist_dataframe = nx.to_pandas_edgelist(G)
# Take a look a the dataframe. It's a table with two columns, source and target
# these represent the source and target nodes of each connected pair
# since the network is undirected the source/target label is inconsequential here.
print(edgelist_dataframe)
```
<br>
<div class="alert alert-block alert-info">
NOTE: when the edgelist is not from a toy example this dataframe may have thousands of rows.
Instead of trying to print the entire thing you can have a glimpse of the table by running:
<pre><code>print(edgelist_dataframe.head())</code></pre>
</div>
```
# Let's create a pandas dataframe with the adjacency matrix:
adjacency_dataframe = nx.to_pandas_adjacency(G, dtype=int)
print(adjacency_dataframe)
# note how in this case, the node names are preserved in the dataframe
# (as opposed to when we only generated the matrix above)
# Now that we have the pandas dataframes we can create a new network from it:
G_new = nx.from_pandas_adjacency(adjacency_dataframe)
# Check the new network (which should be the same as before/with the corresponding changes you made to ir before)
nx.draw_circular(G_new, with_labels = True,node_color='crimson',node_size=2500,font_color='white')
```
<br><div class="alert alert-block alert-success">
Now that you have seen how to transform our networkx graphs into pandas dataframes and back, convince yourself that you understand how to save (read) such dataframes into (from) files.
<br>
<br>
You will see more examples of this Input/Output behavior in other notebooks as the course progresses.
</div>
| github_jupyter |
# Puma Example
Kevin Walchko
created 7 Nov 2017
---
This is just an example of a more complex serial manipulator.
```
%matplotlib inline
# Let's grab some libraries to help us manipulate symbolic equations
from __future__ import print_function
from __future__ import division
import numpy as np
import sympy
from sympy import symbols, sin, cos, pi, simplify
def makeT(a, alpha, d, theta):
# create a modified DH homogenious matrix
return np.array([
[ cos(theta), -sin(theta), 0, a],
[sin(theta)*cos(alpha), cos(theta)*cos(alpha), -sin(alpha), -d*sin(alpha)],
[sin(theta)*sin(alpha), cos(theta)*sin(alpha), cos(alpha), d*cos(alpha)],
[ 0, 0, 0, 1]
])
def simplifyT(tt):
"""
This goes through each element of a matrix and tries to simplify it.
"""
for i, row in enumerate(tt):
for j, col in enumerate(row):
tt[i,j] = simplify(col)
return tt
```
# Puma
[](dh_pics/puma.png)
<img src="dh_pics/puma.png" width="400px">
Puma robot is an old, but classical serial manipulator. You can see Criag's example in section 3.7, pg 77. Once you have the DH parameters, you can use the above matrix to find the forward kinematics,
```
# craig puma
t1,t2,t3,t4,t5,t6 = symbols('t1 t2 t3 t4 t5 t6')
a2, a3, d3, d4 = symbols('a2 a3 d3 d4')
T1 = makeT(0,0,0,t1)
T2 = makeT(0,-pi/2,0,t2)
T3 = makeT(a2,0,d3,t3)
T4 = makeT(a3,-pi/2,d4,t4)
T5 = makeT(0,pi/2,0,t5)
T6 = makeT(0,-pi/2,0,t6)
ans = np.eye(4)
for T in [T1, T2, T3, T4, T5, T6]:
ans = ans.dot(T)
print(ans)
ans = simplifyT(ans)
print(ans)
print('position x: {}'.format(ans[0,3]))
print('position y: {}'.format(ans[1,3]))
print('position z: {}'.format(ans[2,3]))
```
Looking at the position, this is the same position listed in Craig, eqn 3.14.
Also, **this is the simplified version!!!**. As you get more joints and degrees of freedom, the equations get nastier. You also can run into situations where you end up with singularities (like division by zero) and send your robot into a bad place!
-----------
<a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-sa/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/">Creative Commons Attribution-ShareAlike 4.0 International License</a>.
| github_jupyter |
# Measurement Error Mitigation
```
from qiskit import QuantumCircuit, QuantumRegister, Aer, transpile, assemble
from qiskit_textbook.tools import array_to_latex
```
### Introduction
The effect of noise is to give us outputs that are not quite correct. The effect of noise that occurs throughout a computation will be quite complex in general, as one would have to consider how each gate transforms the effect of each error.
A simpler form of noise is that occurring during final measurement. At this point, the only job remaining in the circuit is to extract a bit string as an output. For an $n$ qubit final measurement, this means extracting one of the $2^n$ possible $n$ bit strings. As a simple model of the noise in this process, we can imagine that the measurement first selects one of these outputs in a perfect and noiseless manner, and then noise subsequently causes this perfect output to be randomly perturbed before it is returned to the user.
Given this model, it is very easy to determine exactly what the effects of measurement errors are. We can simply prepare each of the $2^n$ possible basis states, immediately measure them, and see what probability exists for each outcome.
As an example, we will first create a simple noise model, which randomly flips each bit in an output with probability $p$.
```
from qiskit.providers.aer.noise import NoiseModel
from qiskit.providers.aer.noise.errors import pauli_error, depolarizing_error
def get_noise(p):
error_meas = pauli_error([('X',p), ('I', 1 - p)])
noise_model = NoiseModel()
noise_model.add_all_qubit_quantum_error(error_meas, "measure") # measurement error is applied to measurements
return noise_model
```
Let's start with an instance of this in which each bit is flipped $1\%$ of the time.
```
noise_model = get_noise(0.01)
```
Now we can test out its effects. Specifically, let's define a two qubit circuit and prepare the states $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$. Without noise, these would lead to the definite outputs `'00'`, `'01'`, `'10'` and `'11'`, respectively. Let's see what happens with noise. Here, and in the rest of this section, the number of samples taken for each circuit will be `shots=10000`.
```
qasm_sim = Aer.get_backend('qasm_simulator')
for state in ['00','01','10','11']:
qc = QuantumCircuit(2,2)
if state[0]=='1':
qc.x(1)
if state[1]=='1':
qc.x(0)
qc.measure([0, 1], [0, 1])
t_qc = transpile(qc, qasm_sim)
qobj = assemble(t_qc)
counts = qasm_sim.run(qobj, noise_model=noise_model, shots=10000).result().get_counts()
print(state+' becomes', counts)
```
Here we find that the correct output is certainly the most dominant. Ones that differ on only a single bit (such as `'01'`, `'10'` in the case that the correct output is `'00'` or `'11'`), occur around $1\%$ of the time. Those that differ on two bits occur only a handful of times in 10000 samples, if at all.
So what about if we ran a circuit with this same noise model, and got an result like the following?
```
{'10': 98, '11': 4884, '01': 111, '00': 4907}
```
Here `'01'` and `'10'` occur for around $1\%$ of all samples. We know from our analysis of the basis states that such a result can be expected when these outcomes should in fact never occur, but instead the result should be something that differs from them by only one bit: `'00'` or `'11'`. When we look at the results for those two outcomes, we can see that they occur with roughly equal probability. We can therefore conclude that the initial state was not simply $\left|00\right\rangle$, or $\left|11\right\rangle$, but an equal superposition of the two. If true, this means that the result should have been something along the lines of:
```
{'11': 4977, '00': 5023}
```
Here is a circuit that produces results like this (up to statistical fluctuations).
```
qc = QuantumCircuit(2,2)
qc.h(0)
qc.cx(0,1)
qc.measure([0, 1], [0, 1])
t_qc = transpile(qc, qasm_sim)
qobj = assemble(t_qc)
counts = qasm_sim.run(qobj, noise_model=noise_model, shots=10000).result().get_counts()
print(counts)
```
In this example we first looked at results for each of the definite basis states, and used these results to mitigate the effects of errors for a more general form of state. This is the basic principle behind measurement error mitigation.
### Error mitigation with linear algebra
Now we just need to find a way to perform the mitigation algorithmically rather than manually. We will do this by describing the random process using matrices. For this we need to rewrite our counts dictionaries as column vectors. For example, the dictionary `{'10': 96, '11': 1, '01': 95, '00': 9808}` would be rewritten as
$$
C =
\begin{pmatrix}
9808 \\
95 \\
96 \\
1
\end{pmatrix}.
$$
Here the first element is that for `'00'`, the next is that for `'01'`, and so on.
The information gathered from the basis states $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$ can then be used to define a matrix, which rotates from an ideal set of counts to one affected by measurement noise. This is done by simply taking the counts dictionary for $\left|00\right\rangle$, normalizing it so that all elements sum to one, and then using it as the first column of the matrix. The next column is similarly defined by the counts dictionary obtained for $\left|01\right\rangle$, and so on.
There will be statistical variations each time the circuit for each basis state is run. In the following, we will use the data obtained when this section was written, which was as follows.
```
00 becomes {'10': 96, '11': 1, '01': 95, '00': 9808}
01 becomes {'10': 2, '11': 103, '01': 9788, '00': 107}
10 becomes {'10': 9814, '11': 90, '01': 1, '00': 95}
11 becomes {'10': 87, '11': 9805, '01': 107, '00': 1}
```
This gives us the following matrix.
$$
M =
\begin{pmatrix}
0.9808&0.0107&0.0095&0.0001 \\
0.0095&0.9788&0.0001&0.0107 \\
0.0096&0.0002&0.9814&0.0087 \\
0.0001&0.0103&0.0090&0.9805
\end{pmatrix}
$$
If we now take the vector describing the perfect results for a given state, applying this matrix gives us a good approximation of the results when measurement noise is present.
$$ C_{noisy} = M ~ C_{ideal}$$
.
As an example, let's apply this process for the state $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$,
$$
\begin{pmatrix}
0.9808&0.0107&0.0095&0.0001 \\
0.0095&0.9788&0.0001&0.0107 \\
0.0096&0.0002&0.9814&0.0087 \\
0.0001&0.0103&0.0090&0.9805
\end{pmatrix}
\begin{pmatrix}
5000 \\
0 \\
0 \\
5000
\end{pmatrix}
=
\begin{pmatrix}
4904.5 \\
101 \\
91.5 \\
4903
\end{pmatrix}.
$$
In code, we can express this as follows.
```
import numpy as np
M = [[0.9808,0.0107,0.0095,0.0001],
[0.0095,0.9788,0.0001,0.0107],
[0.0096,0.0002,0.9814,0.0087],
[0.0001,0.0103,0.0090,0.9805]]
Cideal = [[5000],
[0],
[0],
[5000]]
Cnoisy = np.dot(M, Cideal)
array_to_latex(Cnoisy, pretext="\\text{C}_\\text{noisy} = ")
```
Either way, the resulting counts found in $C_{noisy}$, for measuring the $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$ with measurement noise, come out quite close to the actual data we found earlier. So this matrix method is indeed a good way of predicting noisy results given a knowledge of what the results should be.
Unfortunately, this is the exact opposite of what we need. Instead of a way to transform ideal counts data into noisy data, we need a way to transform noisy data into ideal data. In linear algebra, we do this for a matrix $M$ by finding the inverse matrix $M^{-1}$,
$$C_{ideal} = M^{-1} C_{noisy}.$$
```
import scipy.linalg as la
M = [[0.9808,0.0107,0.0095,0.0001],
[0.0095,0.9788,0.0001,0.0107],
[0.0096,0.0002,0.9814,0.0087],
[0.0001,0.0103,0.0090,0.9805]]
Minv = la.inv(M)
array_to_latex(Minv)
```
Applying this inverse to $C_{noisy}$, we can obtain an approximation of the true counts.
```
Cmitigated = np.dot(Minv, Cnoisy)
array_to_latex(Cmitigated, pretext="\\text{C}_\\text{mitigated}=")
```
Of course, counts should be integers, and so these values need to be rounded. This gives us a very nice result.
$$
C_{mitigated} =
\begin{pmatrix}
5000 \\
0 \\
0 \\
5000
\end{pmatrix}
$$
This is exactly the true result we desire. Our mitigation worked extremely well!
### Error mitigation in Qiskit
```
from qiskit.ignis.mitigation.measurement import complete_meas_cal, CompleteMeasFitter
```
The process of measurement error mitigation can also be done using tools from Qiskit. This handles the collection of data for the basis states, the construction of the matrices and the calculation of the inverse. The latter can be done using the pseudo inverse, as we saw above. However, the default is an even more sophisticated method using least squares fitting.
As an example, let's stick with doing error mitigation for a pair of qubits. For this we define a two qubit quantum register, and feed it into the function `complete_meas_cal`.
```
qr = QuantumRegister(2)
meas_calibs, state_labels = complete_meas_cal(qr=qr, circlabel='mcal')
```
This creates a set of circuits to take measurements for each of the four basis states for two qubits: $\left|00\right\rangle$, $\left|01\right\rangle$, $\left|10\right\rangle$ and $\left|11\right\rangle$.
```
for circuit in meas_calibs:
print('Circuit',circuit.name)
print(circuit)
print()
```
Let's now run these circuits without any noise present.
```
# Execute the calibration circuits without noise
t_qc = transpile(meas_calibs, qasm_sim)
qobj = assemble(t_qc, shots=10000)
cal_results = qasm_sim.run(qobj, shots=10000).result()
```
With the results we can construct the calibration matrix, which we have been calling $M$.
```
meas_fitter = CompleteMeasFitter(cal_results, state_labels, circlabel='mcal')
array_to_latex(meas_fitter.cal_matrix)
```
With no noise present, this is simply the identity matrix.
Now let's create a noise model. And to make things interesting, let's have the errors be ten times more likely than before.
```
noise_model = get_noise(0.1)
```
Again we can run the circuits, and look at the calibration matrix, $M$.
```
t_qc = transpile(meas_calibs, qasm_sim)
qobj = assemble(t_qc, shots=10000)
cal_results = qasm_sim.run(qobj, noise_model=noise_model, shots=10000).result()
meas_fitter = CompleteMeasFitter(cal_results, state_labels, circlabel='mcal')
array_to_latex(meas_fitter.cal_matrix)
```
This time we find a more interesting matrix, and one that we cannot use in the approach that we described earlier. Let's see how well we can mitigate for this noise. Again, let's use the Bell state $(\left|00\right\rangle+\left|11\right\rangle)/\sqrt{2}$ for our test.
```
qc = QuantumCircuit(2,2)
qc.h(0)
qc.cx(0,1)
qc.measure([0, 1], [0, 1])
t_qc = transpile(qc, qasm_sim)
qobj = assemble(t_qc, shots=10000)
results = qasm_sim.run(qobj, noise_model=noise_model, shots=10000).result()
noisy_counts = results.get_counts()
print(noisy_counts)
```
In Qiskit we mitigate for the noise by creating a measurement filter object. Then, taking the results from above, we use this to calculate a mitigated set of counts. Qiskit returns this as a dictionary, so that the user doesn't need to use vectors themselves to get the result.
```
# Get the filter object
meas_filter = meas_fitter.filter
# Results with mitigation
mitigated_results = meas_filter.apply(results)
mitigated_counts = mitigated_results.get_counts()
```
To see the results most clearly, let's plot both the noisy and mitigated results.
```
from qiskit.visualization import plot_histogram
noisy_counts = results.get_counts()
plot_histogram([noisy_counts, mitigated_counts], legend=['noisy', 'mitigated'])
```
Here we have taken results for which almost $20\%$ of samples are in the wrong state, and turned it into an exact representation of what the true results should be. However, this example does have just two qubits with a simple noise model. For more qubits, and more complex noise models or data from real devices, the mitigation will have more of a challenge. Perhaps you might find methods that are better than those Qiskit uses!
```
import qiskit
qiskit.__qiskit_version__
```
| github_jupyter |
# Import Libraries
```
# import necessary libraries
import numpy as np
import pandas as pd
import re
import pickle
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import matplotlib.image as mpimg
from sklearn.model_selection import train_test_split
from datetime import date
from datetime import datetime
import torch
from torch import nn
from torch.utils.data import Dataset
from torch.utils.data import DataLoader
from torchvision import transforms
from torch.nn import functional as F
from torch import nn, optim
from rdkit import Chem
from rdkit.Chem import Draw
import selfies as sf
# import chemVAE functions
from chemVAE import main
```
# One-Hot Encode SMILES Data
Dataset is from the Clean Energy Project https://www.worldcommunitygrid.org/research/cep1/overview.do
```
# load dataset
df = pd.read_csv('opv_molecules.csv')
# select SMILEs data
smiles = df['SMILES'].values
selfies = main.smiles2selfies(smiles)
onehot_selfies, idx_to_symbol = main.onehotSELFIES(selfies)
```
# Load One-Hot Encoded Data into Pytorch Dataset
```
# split data in training and testing
X_train, X_test, y_train, y_test = train_test_split(onehot_selfies, onehot_selfies, test_size = 0.40)
X_test, X_val, y_test, y_val = train_test_split(X_test, X_test, test_size = 0.50)
# Pytroch Dataset
train_data = main.SELFIES_Dataset(X_train, y_train, transform = transforms.ToTensor())
test_data = main.SELFIES_Dataset(X_test, y_test, transform = transforms.ToTensor())
val_data = main.SELFIES_Dataset(X_val, y_val, transform = transforms.ToTensor())
```
# Set Model Parameters
```
num_characters, max_seq_len = onehot_selfies[0].shape
params = {'num_characters' : num_characters,
'seq_length' : max_seq_len,
'num_conv_layers' : 3,
'layer1_filters' : 24,
'layer2_filters' : 24,
'layer3_filters' : 24,
'layer4_filters' : 24,
'kernel1_size' : 11,
'kernel2_size' : 11,
'kernel3_size' : 11,
'kernel4_size' : 11,
'lstm_stack_size' : 3,
'lstm_num_neurons' : 396,
'latent_dimensions' : 256,
'batch_size' : 256,
'epochs' : 500,
'learning_rate' : 10**-4}
```
# Train Model
```
# set device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print("Training Model on: " + str(device))
# load data
train_loader = DataLoader(train_data, batch_size = params['batch_size'], shuffle = True)
test_loader = DataLoader(test_data, batch_size = params['batch_size'], shuffle = True)
# initialize model
model = main.VAE(params).to(device)
# set optimizer
optimizer = optim.Adam(model.parameters(), lr = params['learning_rate'])
# set KL annealing
KLD_alpha = np.linspace(0,1, params['epochs'])
## generate unique filenames
date = date.today()
now = datetime.now()
time = now.strftime("%H%M%S")
model_filename = "model_" + str(date)
# train model
epoch = params['epochs']
train_loss = []
test_loss = []
BCE_loss = []
KLD_loss = []
KLD_weight = []
for epoch in range(1, epoch + 1):
alpha = KLD_alpha[epoch-1]
loss, BCE, KLD_wt, KLD = main.train(model, train_loader, optimizer, device, epoch, alpha)
train_loss.append(loss)
BCE_loss.append(BCE)
KLD_loss.append(KLD)
KLD_weight.append(KLD_wt)
test_loss.append(main.test(model, test_loader, optimizer, device, epoch, alpha))
# save model
## save model paramters
output = open(model_filename +'_parameters.pkl', 'wb')
pickle.dump(params, output)
output.close()
print("Saved PyTorch Parameters State to " + model_filename +'_parameters.pkl')
## save model state
torch.save(model.state_dict(), model_filename + '_state.pth')
print("Saved PyTorch Model State to " + model_filename + '_state.pth')
## save model state
torch.save(model, model_filename + '.pth')
print("Saved PyTorch Model State to " + model_filename + '_state.pth')
```
# Deploy Model
```
# Load the model trained in the cell above
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
## load parameters
pkl_file = open(model_filename + '_parameters.pkl', 'rb')
params = pickle.load(pkl_file)
pkl_file.close()
## load model state
model = main.VAE(params).to(device)
model.load_state_dict(torch.load(model_filename + '_state.pth'))
# Load a pretrained model
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
## load parameters
pkl_file = open('pretrained_model_parameters.pkl', 'rb')
params = pickle.load(pkl_file)
pkl_file.close()
## load model state
model = main.VAE(params).to(device)
model.load_state_dict(torch.load('pretrained_model_state.pth'))
# grab random sample from test_data
sample_idx = np.random.randint(0,len(test_data)-1)
img, label = train_data[sample_idx]
# run model
with torch.no_grad():
img = img.to(device)
recon_data, z, mu, logvar = model(img)
recon_data = recon_data[0].cpu()
# grab original smiles
sample = img[0].cpu().numpy()
char_ind = list(np.argmax(sample.squeeze(),axis=0))
string = [idx_to_symbol[i] for i in char_ind]
selfie = ''.join(string)
smiles = sf.decoder(selfie)
# reconstructed smiles
recon_sample = recon_data.numpy()
char_ind = list(np.argmax(recon_sample.squeeze(),axis=0))
string = [idx_to_symbol[i] for i in char_ind]
recon_selfie = ''.join(string)
recon_smiles = sf.decoder(recon_selfie)
# visualize model reconstruction
## draw molecule and reconstruction
m1 = Chem.MolFromSmiles(smiles)
Draw.MolToFile(m1,'original.png')
m2 = Chem.MolFromSmiles(recon_smiles)
Draw.MolToFile(m2,'reconstruct.png')
## visualize molecules in notebokk
figure(figsize=(8, 6), dpi = 100)
plt.subplot(1,2,1)
img = mpimg.imread('original.png')
plt.imshow(img)
plt.axis('off')
plt.title('Original')
plt.subplot(1,2,2)
img = mpimg.imread('reconstruct.png')
plt.imshow(img)
plt.axis('off')
plt.title('Reconstruction')
```
| github_jupyter |
## Breast cancer detection using deep learning
In this notebook we are going to use the [Breast Histopathology Images](https://www.kaggle.com/paultimothymooney/breast-histopathology-images) dataset and the `fastai` library for detecting breast cancer.
**Context**:
Invasive Ductal Carcinoma (IDC) is the most common subtype of all breast cancers. To assign an aggressiveness grade to a whole mount sample, pathologists typically focus on the regions which contain the IDC. As a result, one of the common pre-processing steps for automatic aggressiveness grading is to delineate the exact regions of IDC inside of a whole mount slide.
**About the dataset**:
The original dataset consisted of 162 whole mount slide images of Breast Cancer (BCa) specimens scanned at 40x. From that, 277,524 patches of size 50 x 50 were extracted (198,738 IDC negative and 78,786 IDC positive). Each patch’s file name is of the format: u_xX_yY_classC.png — > example 10253_idx5_x1351_y1101_class0.png . Where u is the patient ID (10253_idx5), X is the x-coordinate of where this patch was cropped from, Y is the y-coordinate of where this patch was cropped from, and C indicates the class where 0 is non-IDC and 1 is IDC.
**Inspiration**:
Breast cancer is the most common form of cancer in women, and invasive ductal carcinoma (IDC) is the most common form of breast cancer. Accurately identifying and categorizing breast cancer subtypes is an important clinical task, and automated methods can be used to save time and reduce error.
**Adrian Rosebrock **of **PyImageSearch** has [this wonderful tutorial](https://www.pyimagesearch.com/2019/02/18/breast-cancer-classification-with-keras-and-deep-learning/) on this same topic as well. Be sure to check that out if you have not. I decided to use the `fastai` library and to see *if I could improve the predictive performance by incorporating modern deep learning practices*.
Let's take a look at the class distribution of the dataset again - >
* 198,738 negative examples (i.e., no breast cancer)
* 78,786 positive examples (i.e., indicating breast cancer was found in the patch)
0 indicates `no IDC` (no breast cancer) while 1 indicates `IDC` (breast cancer)
As we can see, this is a clear example of *class-imbalance*. But we will start simple and do a lot of experimentation for taking major decisions for the model training and tricking.
```
# Get the fastai libraries and other important stuff: https://course.fast.ai/start_colab.html
!curl -s https://course.fast.ai/setup/colab | bash
# Authenticate Colab to use my Google Drive for data storage and retrieval
from google.colab import drive
drive.mount('/content/gdrive', force_remount=True)
root_dir = "/content/gdrive/My Drive/"
base_dir = root_dir + 'BreastCancer'
base_dir
# Change the working directory
%cd /content/gdrive/My\ Drive/BreastCancer
# Verify
!pwd
```
Unzip the zipped folder of data by `!unzip /content/gdrive/My\ Drive/BreastCancer/IDC_regular_ps50_idx5.zip`. It will take time. When the process completes, make sure to move the zipped folder to somwhere else (or remove).
<font color=red>Warning ahead! The unzipping process takes time. So be patient. </font>
```
!unzip /content/gdrive/My\ Drive/BreastCancer/IDC_regular_ps50_idx5.zip
!find /content/gdrive/My\ Drive/BreastCancer -maxdepth 1 -type d | wc -l
```
### Magics and imports
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
from fastai.vision import *
from fastai.metrics import *
import numpy as np
np.random.seed(7)
import torch
torch.cuda.manual_seed_all(7)
import matplotlib.pyplot as ply
plt.style.use('ggplot')
```
### Instantiating the data augmentation object with a number of useful transforms
```
tfms = get_transforms(do_flip=True, flip_vert=True,
max_lighting=0.3, max_warp=0.3, max_rotate=20., max_zoom=0.05)
len(tfms)
```
### Loading the data in mini batches of 128 (48x48)
```
path = '/content/gdrive/My Drive/BreastCancer/'
data = ImageDataBunch.from_folder(path, ds_tfms=tfms, valid_pct=0.2,
size=48, bs=128).normalize(imagenet_stats)
data.show_batch(rows=3, figsize=(8,8))
```
Just to remind you - **0 indicates `no IDC` (no breast cancer) while 1 indicates `IDC` (breast cancer)**.
```
# Training and validation set splits
data.label_list
```
### Distribution of the classes in the new training and validation set
```
from collections import Counter
# Training set
train_counts = Counter(data.train_ds.y)
train_counts.most_common()
```
[(Category 0, 159089), (Category 1, 62931)]
```
# Validation set
valid_counts = Counter(data.valid_ds.y)
valid_counts.most_common()
```
[(Category 0, 39649), (Category 1, 15855)]
### Training a pretrained ResNet50 + Mixed precision policy
Here, we will train the last layer group a pre-trained ResNet50 model (trained on ImageNet) using mixed precision policy and 1cycle policy. We will also tweak the cross-entropy loss function so that it adds weights to the undersampled class effectively.
```
# Initializing the custom class weights and pop it to the GPU
from torch import nn
weights = [0.4, 1]
class_weights=torch.FloatTensor(weights).cuda()
# Begin the training
learn = cnn_learner(data, models.resnet50, metrics=[accuracy]).to_fp16()
learn.loss_func = nn.CrossEntropyLoss(weight=class_weights)
learn.fit_one_cycle(5);
learn.recorder.plot_losses()
```
A mammoth training of **1 hour, 21 minutes and 9 seconds.** The loss surface also seems to be pretty good.
```
# Saving the model
learn.save('stage-1-rn50')
```
### Model's losses, accuracy scores and more
```
# Model's final validation loss and accuracy
learn.validate()
```
`tensor(0.8685)` denotes an accuracy score of **86.85%**.
```
# Model's final training loss and accuracy
learn.validate(learn.data.train_dl)
```
As mentioned in the very beginning, the dataset suffers from the problem of class-imbalance. And for class-imbalanced datasets, we cannot simply go with the accuracy score. Read about [accuracy paradox](https://en.wikipedia.org/wiki/Accuracy_paradox). We will have to consider other metrics like **specificity**, **sensitivity**. Let's start by looking at model's predictions on the validation set.
```
# Looking at model's results
learn.show_results(rows=3)
```
The above figure presents a few IDC(-) samples from the validation set and along with that also shows model's predictions. Let's now see the top losses incurred by the model during the training process.
### Model's top losses and confusion matrix
```
interp = ClassificationInterpretation.from_learner(learn)
losses,idxs = interp.top_losses()
len(data.valid_ds)==len(losses)==len(idxs)
interp.plot_top_losses(9, figsize=(12,10), heatmap=False)
```
We can see that there are some samples which are originally IDC(+) but the model predicts them as IDC(-). **This is a staggering issue**.
We need to be really careful with our false negative here — we don’t want to classify someone as “No cancer” when they are in fact “Cancer positive”.
Our false positive rate is also important — we don’t want to mistakenly classify someone as “Cancer positive” and then subject them to painful, expensive, and invasive treatments when they don’t actually need them.
To get a strong hold of how model is doing on false positives and false negatives, we can plot the confusion matrix.
```
interp.plot_confusion_matrix()
```
There can be improvements in the model's training instructions so that the model minimizes the false predictions.
**What can be done?**
- We are using the pre-trained weights of the ResNet50 model. We can train the other layers of the model to make it a bit more specific
- We have trained our model only for a few epochs. A bit more training will definitely help
- We have not used discriminative learning rates i.e. the way of training models using different learning rates across different layers groups
Before we experiment with all this, let's generate a classification report of the current model's performance.
### Classification report to look at other metrics since there is a class imbalance
```
from sklearn.metrics import classification_report
def return_classification_report(learn):
ground_truth = []
pred_labels = []
for i in range(len(learn.data.valid_ds)):
temp_pred = str(learn.predict(learn.data.valid_ds[i][0])[0])
temp_truth = str(learn.data.valid_ds[i]).split('), ', 1)[1].replace('Category ', '').replace(')', '')
pred_labels.append(temp_pred)
ground_truth.append(temp_truth)
assert len(pred_labels) == len(ground_truth)
return classification_report(ground_truth, pred_labels, target_names=data.classes)
print(return_classification_report(learn))
```
As we can see the model's [recall](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.precision_recall_fscore_support.html#sklearn.metrics.precision_recall_fscore_support) is much better than that of Adrian's `CancerNet` model. This is due to the fact that *we handled the loss function in a custom way*.
Here's a snap of the classification report of `CancerNet` -

**Can we do better?**
We will now start by finding an optimal learning rate for the model.
```
learn.lr_find();
learn.recorder.plot()
```
We now have an idea of what could be a good learning rate for the model. We will now unfreeze the first layer groups of the model and will allow it to fully train. We will train it using discriminative learning rates for another two epochs.
```
learn.unfreeze()
learn.fit_one_cycle(2, max_lr=slice(1e-04, 1e-05))
```
Another **32 minutes and 24 seconds** of training.
```
# Save model
learn.save('stage-2-more-rn50')
# Looking at the classification report
print(return_classification_report(learn))
```
As we can see the recall has improved specifically for the positive classes. Ideally there should be a good balance of specificity and sensitivity.
### Model's architectural summary
```
learn.summary()
# Export the model in pickle format
learn.export('breast-cancer-rn50.pkl')
```
### Conclusion:
We now have a model which is **86.68%** accurate and has got a pretty **improved recall** for both in case of the +ve and the -ve classes.
We still could have trained the network for more. We have trained it for **7 epochs** and it took *approximately* **two hours.** More fine-tuning could have been done. Sophisticated data augmentation and resolution techniques could have been applied. But let's keep them aside for further studies for now :)
| github_jupyter |
# EJERCICIO 4
El conjunto de datos “Iris” ha sido usado como caso de prueba para una gran cantidad de clasificadores y es, quizás, el conjunto de datos más conocido de la literatura específica. Iris es una variedad de planta que se la desea clasificar de acuerdo a su tipo. Se reconocen tres tipos distintos: 'Iris setosa', 'Iris versicolor' e 'Iris virgínica'. El objetivo es lograr clasificar una planta de la variedad Iris a partir del largo y del ancho del pétalo y del largo y del ancho del sépalo.
El conjunto de datos Iris está formado en total por 150 muestras,
siendo 50 de cada uno de los tres tipos de plantas. Cada muestra
está compuesta por el tipo de planta, la longitud y ancho del
pétalo y la longitud y ancho del sépalo. Todos son atributos numéricos continuos.
$$
\begin{array}{|c|c|c|c|c|}
\hline X & Setosa & Versicolor & Virgínica & Inválidas \\
\hline Setosa & 50 & 0 & 0 & 0 \\
\hline Versicolor & 0 & 50 & 0 & 0 \\
\hline Virgínica & 0 & 0 & 50 & 0 \\
\hline
\end{array}
$$
```
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
import mpld3
%matplotlib inline
mpld3.enable_notebook()
from cperceptron import Perceptron
from cbackpropagation import ANN #, Identidad, Sigmoide
import patrones as magia
def progreso(ann, X, T, y=None, n=-1, E=None):
if n % 20 == 0:
print("Pasos: {0} - Error: {1:.32f}".format(n, E))
def progresoPerceptron(perceptron, X, T, n):
y = perceptron.evaluar(X)
incorrectas = (T != y).sum()
print("Pasos: {0}\tIncorrectas: {1}\n".format(n, incorrectas))
iris = np.load('iris.npy')
#Armo Patrones
clases, patronesEnt, patronesTest = magia.generar_patrones(
magia.escalar(iris[:,1:]).round(4),iris[:,:1],80)
X, T = magia.armar_patrones_y_salida_esperada(clases,patronesEnt)
clases, patronesEnt, noImporta = magia.generar_patrones(
magia.escalar(iris[:,1:]),iris[:,:1],100)
Xtest, Ttest = magia.armar_patrones_y_salida_esperada(clases,patronesEnt)
```
a) Entrene perceptrones para que cada uno aprenda a reconocer uno de los distintos tipos de plantas Iris. Informe los parámetros usados para el entrenamiento y el desempeño obtenido. Emplee todos los patrones para el entrenamiento. Muestre la matriz de confusión para la mejor clasificación obtenida luego del entrenamiento, informando los patrones clasificados correcta e incorrectamente.
```
print("Entrenando P1:")
p1 = Perceptron(X.shape[1])
I1 = p1.entrenar_numpy(X, T[:,0], max_pasos=5000, callback=progresoPerceptron, frecuencia_callback=2500)
print("Pasos:{0}".format(I1))
print("\nEntrenando P2:")
p2 = Perceptron(X.shape[1])
I2 = p2.entrenar_numpy(X, T[:,1], max_pasos=5000, callback=progresoPerceptron, frecuencia_callback=2500)
print("Pasos:{0}".format(I2))
print("\nEntrenando P3:")
p3 = Perceptron(X.shape[1])
I3 = p3.entrenar_numpy(X, T[:,2], max_pasos=5000, callback=progresoPerceptron, frecuencia_callback=2500)
print("Pasos:{0}".format(I3))
Y = np.vstack((p1.evaluar(Xtest),p2.evaluar(Xtest),p3.evaluar(Xtest))).T
magia.matriz_de_confusion(Ttest,Y)
```
b) Entrene una red neuronal artificial usando backpropagation como algoritmo de aprendizaje con el fin de lograr la clasificación pedida. Emplee todos los patrones para el entrenamiento. Detalle los parámetros usados para el entrenamiento así como la arquitectura de la red neuronal. Repita más de una vez el procedimiento para confirmar los resultados obtenidos e informe la matriz de confusión para la mejor clasificación obtenida.
```
# Crea la red neuronal
ocultas = 10
entradas = X.shape[1]
salidas = T.shape[1]
ann = ANN(entradas, ocultas, salidas)
ann.reiniciar()
#Entreno
E, n = ann.entrenar_rprop(X, T, min_error=0, max_pasos=100000, callback=progreso, frecuencia_callback=10000)
print("\nRed entrenada en {0} pasos con un error de {1:.32f}".format(n, E))
#Evaluo
Y = (ann.evaluar(Xtest) >= 0.97)
magia.matriz_de_confusion(Ttest,Y)
(ann.evaluar(Xtest)[90])
```
| github_jupyter |
```
import pyspark
from pyspark import SparkContext
sc = SparkContext.getOrCreate();
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # Property used to format output tables better
spark
sc.stop()
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # Property used to format output tables better
spark
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('DataAnalysisOnElonMusk').getOrCreate()
import os
import re
from datetime import date, datetime
import pandas as pd
file_path_name = 'elonmusk.csv'
def open_file(file_path_name):
return pd.read_csv(file_path_name, index_col=[0])
print(open_file(file_path_name).head())
def clean_dataframe(df, columns_to_drop):
df = drop_redundant_columns(df, columns_to_drop)
return df
def transform_dataframe(df):
df = drop_columns_with_constant_values(df)
add_mentions_count(df)
add_weekday(df)
add_reply_to_count(df)
add_photos_count(df)
convert_to_datetime(df)
extract_hour_minute(df)
df = drop_redundant_columns(df, ['photos', 'date', 'mentions', 'reply_to', 'reply_to_count'])
return df
columns_to_drop = ['hashtags', 'cashtags', 'link', 'quote_url', 'urls', 'created_at']
def drop_redundant_columns(df, columns_to_drop):
return df.drop(columns=columns_to_drop, axis=0)
def drop_columns_with_constant_values(df):
return df.drop(columns=list(df.columns[df.nunique() <= 1]))
def add_mentions_count(df):
new_values = []
for i, content in df['mentions'].items():
new_values.append(int(content.count("'") / 2))
df['mentions_count'] = new_values
return df
def add_weekday(df):
weekday = []
for i, content in df['date'].items():
year, month, day = map(int, content.split('-'))
d = date(year, month, day)
weekday.append(d.weekday())
df['weekday'] = weekday
return df
def add_reply_to_count(df):
reply_to_count_values = []
for i, content in df['reply_to'].items():
reply_to_count_values.append((int(content.count("{")) - 1))
df['reply_to_count'] = reply_to_count_values
return df
def add_photos_count(df):
new_values = []
for i, content in df['photos'].items():
new_values.append(int(content.count("https")))
df['photos_count'] = new_values
return df
def convert_to_datetime(df):
df['datetime'] = (df['date'] + " " + df['time']).astype('string')
return df
def extract_hour_minute(df):
year_col = []
month_col = []
hour_col = []
minute_col = []
for i, content in df['datetime'].items():
t1 = datetime.strptime(content, '%Y-%m-%d %H:%M:%S')
year_col.append(t1.year)
month_col.append(t1.month)
hour_col.append(t1.hour)
minute_col.append(t1.minute)
df['year'] = year_col
df['month'] = month_col
df['hour'] = hour_col
df['minute'] = minute_col
return df
df = open_file(file_path_name)
new_df = clean_dataframe(df, columns_to_drop)
new_df = transform_dataframe(new_df)
df['tweet'] = df['tweet'].str.lower()
data = []
for i,j in zip(new_df,new_df.count()):
data.append((i,str(j)))
rdd = spark.sparkContext.parallelize(data)
resultCount = rdd.collect()
print(resultCount)
#Dropping duplicates from previous count data
new_df.drop_duplicates(subset=['tweet'], keep='first', inplace=True)
#print(new_df.shape)
shape = spark.sparkContext.parallelize([new_df.shape]).collect()
print(shape)
data2 = []
for i,j in zip(new_df,new_df.count()):
data2.append((i,str(j)))
rdd = spark.sparkContext.parallelize(data2)
resultCount2 = rdd.collect()
print(resultCount2)
count = new_df['tweet'].str.split().str.len()
count.index = count.index.astype(str) + ' words:'
count.sort_index(inplace=True)
def word_count(df):
words_count = []
for i, content in df['tweet'].items():
new_values =[]
new_values = content.split()
words_count.append(len(new_values))
df['word_count'] = words_count
return df
new_df = word_count(new_df)
print("Total number of words: ", count.sum(), "words")
print("Average number of words per tweet: ", round(count.mean(),2), "words")
print("Max number of words per tweet: ", count.max(), "words")
print("Min number of words per tweet: ", count.min(), "words")
new_df['tweet_length'] = new_df['tweet'].str.len()
print("Total length of a dataset: ", new_df.tweet_length.sum(), "characters")
print("Average length of a tweet: ", round(new_df.tweet_length.mean(),0), "characters")
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 自定义联合算法,第 1 部分:Federated Core 简介
<table class="tfo-notebook-buttons" align="left">
<td> <a target="_blank" href="https://tensorflow.google.cn/federated/tutorials/custom_federated_algorithms_1"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png">在 TensorFlow.org 上查看</a> </td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/federated/tutorials/custom_federated_algorithms_1.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png">在 Google Colab 中运行</a></td>
<td> <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/federated/tutorials/custom_federated_algorithms_1.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png">在 GitHub 上查看源代码</a>
</td>
<td> <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/zh-cn/federated/tutorials/custom_federated_algorithms_1.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png">下载笔记本</a> </td>
</table>
本系列教程包括两个部分,此为第一部分。该系列演示了如何使用 [Federated Core (FC)](../federated_core.md) 在 TensorFlow Federated (TFF) 中实现自定义类型的联合算法。Federated Core 是一组较低级别的接口,这些接口是我们实现[联合学习 (FL)](../federated_learning.md) 层的基础。
第一部分更具概念性;我们将介绍 TFF 中使用的一些关键概念和编程抽象,并在一个非常简单的示例(分布式温度传感器阵列)中演示它们的用法。在[本系列的第二部分](custom_federated_algorithms_2.ipynb)中,我们将使用此处介绍的机制来实现一个联合训练和评估算法的简单版本。我们鼓励您稍后在 <code>tff.learning</code> 中研究联合平均的<a>实现</a>。
在本系列的最后,您应该能够认识到 Federated Core 的应用并不仅限于学习。我们提供的编程抽象非常通用,例如可用于对分布式数据进行分析和其他自定义类型的计算。
尽管本教程可独立使用,但我们建议您先阅读有关[图像分类](federated_learning_for_image_classification.ipynb)和[文本生成](federated_learning_for_text_generation.ipynb)的教程,获得对 TensorFlow Federated 框架和 [Federated Learning](../federated_learning.md) API (`tff.learning`) 更高级和更循序渐进的介绍,它将帮助您在上下文中理解我们在此介绍的概念。
## 预期用途
简而言之,Federated Core (FC) 是一种开发环境,可以紧凑地表达将 TensorFlow 代码与分布式通信算子(比如在[联合平均](https://arxiv.org/abs/1602.05629)中使用的算子)相结合的程序逻辑。它可以在系统中的一组客户端设备上计算分布式总和、平均值和其他类型的分布式聚合,向这些设备广播模型和参数等。
您可能知道 [`tf.contrib.distribute`](https://tensorflow.google.cn/api_docs/python/tf/contrib/distribute),此时自然会问的一个问题可能是:该框架在哪些方面有所不同?毕竟,两种框架都试图使 TensorFlow 进行分布式计算。
其中一种思路是,`tf.contrib.distribute` 的既定目标是*允许用户以最小的更改使用现有模型和训练代码实现分布式训练*,且大部分重点放在如何利用分布式架构来提高现有训练代码的效率。TFF 的 Federated Core 的目标是使研究员和从业者能够明确控制将在系统中使用的分布式通信的具体模式。FC 的重点是提供一种灵活可扩展的语言来表达分布式数据流算法,而不是具体的一组已实现的分布式训练能力。
TFF 的 FC API 的主要目标受众之一是研究员和从业者,他们可能希望尝试新的联合学习算法,并评估微妙的设计选择(这些选择会影响分布式系统中数据流的编排方式)的结果,但又不想被系统实现细节所困扰。FC API 所针对的抽象级别大致对应于伪代码,可以用来描述研究论文中的联合学习算法的机制(系统中存在什么数据以及如何对其进行转换),但又不降低到单个点对点网络消息交换的级别。
TFF 总体上针对的是数据分布的场景(并且出于隐私等原因必须保持这种状态),以及可能无法在某个集中位置收集所有数据的场景。与所有数据都可以在数据中心积累到某个集中位置的场景相比,这意味着机器学习算法的实现需要提高显式控制的程度。
## 准备工作
在深入研究代码之前,请尝试运行以下 “Hello World” 示例,以确保您的环境已正确设置。如果无法正常运行,请参阅[安装](../install.md)指南中的说明。
```
#@test {"skip": true}
!pip install --quiet --upgrade tensorflow-federated-nightly
!pip install --quiet --upgrade nest-asyncio
import nest_asyncio
nest_asyncio.apply()
import collections
import numpy as np
import tensorflow as tf
import tensorflow_federated as tff
@tff.federated_computation
def hello_world():
return 'Hello, World!'
hello_world()
```
## 联合数据
TFF 的一个显著功能是,您可以通过它在*联合数据*上紧凑地表达基于 TensorFlow 的计算。在本教程中,我们将使用*联合数据*这一术语来指代托管在分布式系统中一组设备上的数据项的集合。例如,在移动设备上运行的应用可以收集数据并将其存储在本地,而无需上传到某一集中位置。或者,分布式传感器阵列可以在本地收集并存储温度读数。
像上面示例中这样的联合数据在 TFF 中被视为[一等公民](https://en.wikipedia.org/wiki/First-class_citizen),即,它们可以显示为函数的参数和结果,并具有类型。为了强化这一概念,我们将联合数据集称为*联合值*或*联合类型的值*。
需要理解的一个要点是,我们将所有设备上的数据项的整个集合(例如,来自分布式阵列中所有传感器温度读数的整个集合)建模为单个联合值。
例如,下面是在 TFF 中定义由一组客户端设备托管的*联合浮点*类型的方式。可以将分布式传感器阵列中出现的温度读数的集合建模为此联合类型的值。
```
federated_float_on_clients = tff.type_at_clients(tf.float32)
```
更普遍的是,TFF 中的联合类型是通过指定其*成员组成*(留驻在各个设备上的数据项)的类型 `T` 和托管此类型联合值的设备组 `G`(加上我们会在稍后提及的第三个可选位)来定义的。我们将托管联合值的设备组 `G` 称为该值的*布局*。因此,`tff.CLIENTS` 是布局示例。
```
str(federated_float_on_clients.member)
str(federated_float_on_clients.placement)
```
带有成员组成 `T` 和布局 `G` 的联合类型可以紧凑地表示为 `{T}@G`,如下所示。
```
str(federated_float_on_clients)
```
此简明表示法中的大括号 `{}` 提醒您成员组成(不同设备上的数据项)可能有所不同,例如您所期望的温度传感器读数,因此,客户端会作为整体共同托管 `T` 型项的[多重集](https://en.wikipedia.org/wiki/Multiset),它们共同构成联合值。
需要注意的是,联合值的成员组成通常对程序员不透明,即不应将联合值视为由系统中的设备标识符进行键控的简单 `dict`,这些值应仅由抽象表示各种分布式通信协议(如聚合)的*联合算子*进行集体转换。如果这听起来太过抽象,不要担心,我们稍后将回到这个问题,并用具体示例对其进行演示。
TFF 中的联合类型有两种形式:一种是联合值的成员组成可能不同(如上所示),另一种是联合值的成员组成全部相等。这由 `tff.FederatedType` 构造函数中的第三个可选参数 `all_equal`(默认为 `False`)来控制。
```
federated_float_on_clients.all_equal
```
可以将带有布局 `G`,且其中所有 `T` 型成员组成已知相等的联合类型紧凑地表示为 `T@G`(与 `{T}@G` 相对,也就是说,去掉大括号表示成员组成的多重集由单个项目构成)。
```
str(tff.type_at_clients(tf.float32, all_equal=True))
```
在实际场景中可能会出现的此类联合值的一个示例是超参数(例如学习率、裁剪范数等),该超参数已由服务器广播到参与联合训练的一组设备。
另一个示例是在服务器上预先训练的一组机器学习模型参数,然后将其广播到一组客户端设备,它们可以在这组设备上针对每个用户进行个性化设置。
例如,假设对于一个简单的一维线性回归模型,我们有一对 `float32` 参数 `a` 和 `b`。我们可以构造如下用于 TFF 的(非联合)类型的此类模型。打印的类型字符串中的尖括号 `<>` 是命名或未命名元组的紧凑 TFF 表示法。
```
simple_regression_model_type = (
tff.StructType([('a', tf.float32), ('b', tf.float32)]))
str(simple_regression_model_type)
```
请注意,虽然我们在上文中仅指定了 `dtype`,但也支持非标量类型。在上面的代码中,`tf.float32` 是更通用的 `tff.TensorType(dtype=tf.float32, shape=[])` 的快捷表示法。
将此模型广播到客户端时,生成的联合值类型可以用如下方法表示。
```
str(tff.type_at_clients(
simple_regression_model_type, all_equal=True))
```
根据上面的*联合浮点*的对称性,我们将这种类型称为*联合元组*。一般来说,我们会经常使用术语*联合 XYZ* 来指代联合值,其中成员组成类似 *XYZ*。因此,我们将对*联合元组*、*联合序列*、*联合模型*等进行讨论。
现在,回到 `float32@CLIENTS`,尽管它看起来是在多个设备上复制的,但它实际上是单一的 `float32`,因为所有成员都相同。通常,您可能会想到任意*全等*联合类型(即一种 `T@G` 形式)与非联合类型 `T` 同构,因为这两种情况实际上都只有 `T` 类型的单个(尽管可能是复制的)项。
鉴于 `T` 和 `T@G` 之间的同构性,您可能想知道后一种类型能够起到什么作用(如果有的话)。请继续阅读。
## 布局
### 设计概述
在上一部分中,我们介绍了*布局*的概念(可能会共同托管联合值的系统参与者组),并且我们还演示了将 `tff.CLIENTS` 用作布局的示例规范。
为了解释为什么*放置*的概念如此重要,以至于我们需要将其合并到 TFF 类型系统中,请回想一下本教程开始时提到的有关 TFF 某些预期用途的内容。
尽管在本教程中,您只会看到在模拟环境中本地执行的 TFF 代码,但我们的目标是使 TFF 能够编写可部署在分布式系统中的物理设备组(可能包括运行 Android 的移动或嵌入式设备)上执行的代码。其中,每个设备都将接收单独的一组指令以在本地执行,具体取决于它在系统中所扮演的角色(最终用户设备、集中协调器、多层架构中的中间层等)。重要的是要能够推断出哪些设备子集执行什么代码,以及数据的不同部分可能在哪里进行物理实现。
当处理移动设备上的应用数据时,这一点尤其重要。由于数据私有且可能敏感,我们要能静态验证此类数据永远不会离开设备(并证明对数据进行处理的实际方式)。布局规范是为此目的而设计的一种机制。
TFF 是一种以数据为中心的编程环境,正因为如此,它与一些现有框架不同,这些框架专注于*运算*和这些运算可能*运行*的位置,而 TFF 专注于*数据*、数据*实现*的位置,以及*转换*方式。因此,布局被建模为 TFF 中数据的属性,而不是数据上运算的属性。的确,您将在下一部分中看到,某些 TFF 运算会跨位置,并且可以说是“在网络中”运行,而不是由一台或一组机器执行。
将某个值的类型表示为 `T@G` 或 `{T}@G`(而不仅仅是 `T`)可使数据布局决策明确,并且搭配 TFF 中编写的程序的静态分析,它可以作为为设备端敏感数据提供形式上的隐私保障的基础。
但此时需要注意,虽然我们鼓励 TFF 用户明确托管数据的参与设备*组*(布局),但程序员永远不会处理原始数据或*各个*参与者的身份。
在 TFF 代码的主体内,根据设计,无法枚举构成由 `tff.CLIENTS` 表示的组的设备,也无法探测组中是否存在某个特定设备。在 Federated Core API、基础架构抽象集或我们提供的用于支持模拟的核心运行时基础结构中,没有任何设备或客户端标识的概念。您编写的所有计算逻辑都将表达为在整个客户端组上的运算。
回想一下我们前面提到的联合类型的值与 Python `dict` 的不同,因为它不能简单地枚举其成员组成。将您的 TFF 程序逻辑所操作的值视为与布局(组)所关联,而不是与单个参与者相关联。
Placements *are* designed to be a first-class citizen in TFF as well, and can appear as parameters and results of a `placement` type (to be represented by `tff.PlacementType` in the API). In the future, we plan to provide a variety of operators to transform or combine placements, but this is outside the scope of this tutorial. For now, it suffices to think of `placement` as an opaque primitive built-in type in TFF, similar to how `int` and `bool` are opaque built-in types in Python, with `tff.CLIENTS` being a constant literal of this type, not unlike `1` being a constant literal of type `int`.
### 指定布局
TFF 提供了两种基本的布局文本 `tff.CLIENTS` 和 `tff.SERVER`,使用通过单个集中式*服务器*协调器编排的多种*客户端*设备(移动电话、嵌入式设备、分布式数据库、传感器等),使自然建模为客户端-服务器架构的各种实际场景易于表达。TFF 的设计还支持自定义位置、多客户端组、多层和其他更通用的分布式架构,但对这些内容的讨论不在本教程的范围之内。
TFF 没有规定 `tff.CLIENTS` 或 `tff.SERVER` 实际代表的内容。
In particular, `tff.SERVER` may be a single physical device (a member of a singleton group), but it might just as well be a group of replicas in a fault-tolerant cluster running state machine replication - we do not make any special architectural assumptions. Rather, we use the `all_equal` bit mentioned in the preceding section to express the fact that we're generally dealing with only a single item of data at the server.
同样,在某些应用中,`tff.CLIENTS` 可能代表系统中的所有客户端,在联合学习的上下文中,我们有时将其称为*群体*,但在[联合平均的生产实现](https://arxiv.org/abs/1602.05629)这个示例中,它可能代表*队列*(选择参加某轮训练的客户端的子集)。当部署计算以执行(或者就像模型环境中的 Python 函数那样被调用)时,其中的抽象定义布局将被赋予具体含义。在我们的本地模拟中,客户端组由作为输入提供的联合数据来确定。
## 联合计算
### 声明联合计算
TFF 是支持模块化开发的强类型函数式编程环境。
TFF 中的基本组成单位是*联合计算*,它是可以接受联合值作为输入并返回联合值作为输出的逻辑的一部分。下面的代码定义了一个计算,它计算的是前一个示例中传感器阵列报告的平均温度。
```
@tff.federated_computation(tff.type_at_clients(tf.float32))
def get_average_temperature(sensor_readings):
return tff.federated_mean(sensor_readings)
```
查看上面的代码,此时您可能会问:TensorFlow 中不是已经有用于定义可组合单元的装饰器构造(如 [`tf.function`](https://tensorflow.google.cn/api_docs/python/tf/function))了吗?既然如此,为什么还要引入另一个构造?它们有什么区别?
简短回答是,`tff.federated_computation` 封装容器生成的代码*既不是* TensorFlow,*也不是* Python,它是一种与平台无关的内部*胶水*语言中的分布式系统规范。这听起来确实很神秘,但请牢记这一将联合计算作为分布式系统抽象规范的直观解释。我们稍后将对其进行说明。
首先,我们来思考一下定义。TFF 计算通常会被建模为函数,参数可有可无,但要有明确定义的类型签名。您可以通过查询计算的 `type_signature` 属性来打印计算的类型签名,如下所示。
```
str(get_average_temperature.type_signature)
```
类型签名告诉我们,该计算接受客户端设备上不同传感器读数的集合,并在服务器上返回单个平均值。
在进一步讨论之前,让我们先思考一个问题:此计算的输入和输出*位于不同位置*(在 `CLIENTS` 上和在 `SERVER` 上)。回想一下我们在上一部分所讲的关于 *TFF 运算如何跨位置并在网络中运行*的内容,以及我们刚才讲到的有关联合计算表示分布式系统抽象规范的内容。我们刚刚定义了这样一种计算:一个简单的分布式系统,其中数据在客户端设备上使用,而聚合结果出现在服务器上。
在许多实际场景中,代表顶级任务的计算倾向于接受其输入并在服务器上报告其输出,这表明,计算可能会由在服务器上发起和终止的*查询*所触发。
但是,FC API 并不强制实施此假设,并且我们在内部使用的许多构建块(包括您可能在 API 中见到的许多 `tff.federated_...` 算子)的输入和输出都有不同的布局,因此,通常来说,您不应将联合计算视为*在服务器上运行*或*由服务器执行*的内容。服务器只是联合计算中的一种类型的参与者。在思考此类计算的机制时,最好始终默认使用全局网络范围的视角,而不是使用单个集中协调器的视角。
一般来说,对于输入和输出的类型 `T` 和 `U`,函数类型签名会分别被紧凑地表示为 `(T -> U)`。形式参数的类型(此处为 `sensor_readings`)被指定为装饰器的参数。您无需指定结果的类型,因为它会自动确定。
尽管 TFF 确实提供了有限形式的多态性,但我们强烈建议程序员明确自己使用的数据类型,因为这样可以更轻松地理解、调试和在形式上验证您的代码的属性。在某些情况下,必须明确指定类型(例如,当前无法直接执行多态计算时)。
### 执行联合计算
为了支持开发和调试,TFF 允许您直接调用以此方式定义为 Python 函数的计算,如下所示。对于 `all_equal` 位设置为 `False` 的情况,您可以将其作为 Python 中的普通 `list` 进行馈送,而对于 `all_equal` 位设置为 `True` 的情况,您可以直接馈送(单)成员组成。这也是向您反馈结果的方式。
```
get_average_temperature([68.5, 70.3, 69.8])
```
在模拟模式下运行此类计算时,您将充当具有系统范围视图的外部观察者,您能够在网络中的任何位置提供输入和使用输出,这里正是如此,您提供了客户端值作为输入,并使用了服务器结果。
现在,让我们回到先前关于 `tff.federated_computation` 装饰器用*胶水*语言发出代码的注释。尽管 TFF 计算的逻辑可以用 Python 表达为普通函数(您只需按照上述方法,使用 `tff.federated_computation` 对其进行装饰),而且您可以像此笔记本中的其他 Python 函数一样直接调用它们,但在后台,正如我们前面提到的,TFF 计算实际上*不是* Python。
我们的意思是,当 Python 解释器遇到一个用 `tff.federated_computation` 装饰的函数时,它会对此函数主体中的语句进行一次跟踪(在定义时),然后构造该计算逻辑的[序列化表示](https://github.com/tensorflow/federated/blob/main/tensorflow_federated/proto/v0/computation.proto)以供将来使用(无论是用于执行,还是作为子组件合并到另一个计算中)。
您可以通过添加打印语句来验证这一点,如下所示:
```
@tff.federated_computation(tff.type_at_clients(tf.float32))
def get_average_temperature(sensor_readings):
print ('Getting traced, the argument is "{}".'.format(
type(sensor_readings).__name__))
return tff.federated_mean(sensor_readings)
```
您可以将定义了联合计算的 Python 代码想象成在非 Eager 上下文中构建了 TensorFlow 计算图的 Python 代码(如果您不熟悉 TensorFlow 的非 Eager 用法,请想象您的 Python 代码定义了运算的计算图以稍后执行,但实际上并不立即运行它们)。TensorFlow 中的非 Eager 计算图构建代码是 Python,但用此代码构建的 TensorFlow 计算图与平台无关且可序列化。
同样,TFF 计算用 Python 进行定义,但其主体中的 Python 语句(如我们刚才展示的示例中的 `tff.federated_mean`)会在后台被编译成可移植、与平台无关和可序列化的表示形式。
作为开发者,您无需关注此表示形式的细节,因为您不会直接使用它,但您应该知道它的存在,以及 TFF 计算在本质上非 Eager,且无法捕获任意 Python 状态。TFF 计算主体中包含的 Python 代码会在定义时(即用 `tff.federated_computation` 装饰的 Python 函数在序列化之前被跟踪时)执行。调用时不会再次对其进行跟踪(函数为多态时除外;有关详细信息,请参阅文档页面)。
您可能想知道为什么我们选择引入专用的内部非 Python 表示形式。其中一个原因是,TFF 计算的最终目的是可部署到实际物理环境中,并托管在可能无法使用 Python 的移动或嵌入式设备上。
另一个原因是 TFF 计算表达的是分布式系统的全局行为,而 Python 程序表达的是各个参与者的本地行为。您可以在上面的简单示例中看到这一点,即使用特殊算子 `tff.federated_mean` 接受客户端设备上的数据,但将结果存储在服务器上。
无法将算子 `tff.federated_mean` 轻松建模为 Python 中的普通算子,因为它不在本地执行,如前所述,它表示协调多个系统参与者行为的分布式系统。我们将此类算子称为*联合算子*,以将其与 Python 中的普通(本地)算子进行区分。
因此,TFF 类型系统以及 TFF 语言支持的基本运算集与 Python 中的大不相同,因此必须使用专用表示形式。
### 组成联合计算
如上所述,最好将联合计算及其组成理解为分布式系统的模型,并且可以将联合计算的组成过程想象成从较简单的分布式系统组成较复杂的分布式系统的过程。您可以将 `tff.federated_mean` 算子视为一种具有类型签名 `({T}@CLIENTS -> T@SERVER)` 的内置模板联合计算(实际上,就像您编写的计算一样,该算子的结构也很复杂,我们会在后台把它分解成更简单的算子)。
组成联合计算的过程也是如此。可以在另一个用 `tff.federated_computation` 装饰的 Python 函数主体中调用计算 `get_average_temperature`,这样做会将其嵌入到父级的主体中,这与先前将 `tff.federated_mean` 嵌入到其自身主体中的方式相同。
需要注意的一个重要限制是,用 `tff.federated_computation` 装饰的 Python 函数的主体必须*仅*由联合算子组成(即它们不能直接包含 TensorFlow 运算)。例如,不能直接使用 `tf.nest` 接口添加一对联合值。TensorFlow 代码仅限用 `tff.tf_computation` 装饰的代码块,下一部分将对此进行讨论。只有以这种方式封装,才能在 `tff.federated_computation` 主体中调用封装后的 TensorFlow 代码。
这样区分是出于技术原因(很难欺骗 `tf.add` 之类的算子来使用非张量),以及架构原因。联合计算的语言(即由用 `tff.federated_computation` 装饰的 Python 函数的序列化主体构造的逻辑)被设计用作与平台无关的*胶水*语言。目前,此胶水语言用来从 TensorFlow 代码(限于 `tff.tf_computation` 块)的嵌入部分构建分布式系统。在时间充裕的情况下,我们会预见需要嵌入其他非 TensorFlow 逻辑的各个部分(例如可能表示输入流水线的关系数据库查询),它们全部使用相同的胶水语言(`tff.federated_computation` 块)相互连接。
## TensorFlow 逻辑
### 声明 TensorFlow 计算
TFF 旨在配合 TensorFlow 使用。这样,您将在 TFF 中编写的大部分代码很可能是普通的(即本地执行的) TensorFlow 代码。为了在 TensorFlow 中使用此类代码,如上所述,只需用 `tff.tf_computation` 对其进行装饰。
例如,下面实现了一个函数,该函数接受数字并向其加 `0.5`。
```
@tff.tf_computation(tf.float32)
def add_half(x):
return tf.add(x, 0.5)
```
再次看到此内容,您可能想知道我们为什么应该定义另一个装饰器 `tff.tf_computation`,而不是简单地使用现有机制(如 `tf.function`)。与前一部分不同,我们在这里处理的是一个普通的 TensorFlow 代码块。
这样做有几个原因,虽然对它们的完整处理超出了本教程的范围,但下面这个主要原因值得注意:
- 若要将使用 TensorFlow 代码实现的可重用构建块嵌入到联合计算的主体中,它们需要满足某些属性(例如在定义时进行跟踪和序列化、具有类型签名等)。这通常需要某种形式的装饰器。
一般而言,我们建议尽可能使用 TensorFlow 的原生机制进行组合(如 `tf.function`),因为 TFF 的装饰器与 Eager 函数进行交互的确切方式可能会逐步变化。
现在,回到上面的示例代码段,我们刚才定义的 `add_half` 计算可以像任何其他 TFF 计算一样由 TFF 处理。尤其是,它具有 TFF 类型签名。
```
str(add_half.type_signature)
```
请注意,此类型签名没有布局。TensorFlow 计算无法使用或返回联合类型。
现在,您还可以在其他计算中将 `add_half` 用作构建块。例如,下面是使用 `tff.federated_map` 算子在客户端设备上将 `add_half` 逐点应用到联合浮点的所有成员组成的方法。
```
@tff.federated_computation(tff.type_at_clients(tf.float32))
def add_half_on_clients(x):
return tff.federated_map(add_half, x)
str(add_half_on_clients.type_signature)
```
### 执行 TensorFlow 计算
执行使用 `tff.tf_computation` 定义的计算所遵循的规则与我们为 `tff.federated_computation` 描述的规则相同。可以将它们作为 Python 中的普通可调用对象进行调用,如下所示。
```
add_half_on_clients([1.0, 3.0, 2.0])
```
同样值得注意的是,以这种方式调用 `add_half_on_clients` 计算会模拟分布式过程。数据会在客户端上使用,并在客户端上返回。实际上,此计算会让每个客户端执行一次本地操作。此系统中没有明确提及 `tff.SERVER`(但在实践中,编排此类处理可能会用到)。可以将以这种方式定义的计算理解为在概念上类似于 `MapReduce` 中的 `Map` 阶段。
另外请记住,我们在前一个部分中讲的关于 TFF 计算会在定义时序列化的内容对 `tff.tf_computation` 代码同样适用,`add_half_on_clients` 的 Python 主体会在定义时被跟踪一次。在后续调用中,TFF 将使用其序列化后的表示形式。
用 `tff.federated_computation` 装饰的 Python 方法与用 `tff.tf_computation` 装饰的方法之间的唯一区别是,后者会被序列化为 TensorFlow 计算图(而不允许前者包含直接嵌入其中的 TensorFlow 代码)。
在后台,每个用 `tff.tf_computation` 装饰的方法会暂时停用 Eager Execution,以便捕获计算的结构。虽然 Eager Execution 已在本地停用,但只要您编写的计算逻辑能够正确序列化,欢迎您使用 Eager TensorFlow、AutoGraph、TensorFlow 2.0 构造等。
例如,以下代码将会失败:
```
try:
# Eager mode
constant_10 = tf.constant(10.)
@tff.tf_computation(tf.float32)
def add_ten(x):
return x + constant_10
except Exception as err:
print (err)
```
上述代码失败的原因是,`constant_10` 在计算图外部构造,而该计算图是 `tff.tf_computation` 在序列化过程中在 `add_ten` 的主体内部构造的。
另一方面,您可以对在 `tff.tf_computation` 内部调用时修改当前计算图的 Python 函数进行调用:
```
def get_constant_10():
return tf.constant(10.)
@tff.tf_computation(tf.float32)
def add_ten(x):
return x + get_constant_10()
add_ten(5.0)
```
请注意,TensorFlow 中的序列化机制正在逐步完善,我们期望 TFF 对计算进行序列化方式的细节也将逐步完善。
### 使用 `tf.data.Dataset`
如前所述,`tff.tf_computation` 的独特之处在于,它们允许您使用由您的代码作为形式参数抽象定义的 `tf.data.Dataset`。如果参数需要在 TensorFlow 中表示为数据集,则需要使用 `tff.SequenceType` 构造函数对其进行声明。
例如,类型规范 `tff.SequenceType(tf.float32)` 定义了 TFF 中浮点元素的抽象序列。序列可以包含张量或复杂的嵌套结构(稍后我们将看到相关示例)。`T` 型项的序列的简明表示形式为 `T*`。
```
float32_sequence = tff.SequenceType(tf.float32)
str(float32_sequence)
```
假设在温度传感器示例中,每个传感器包含不只一个温度读数,而是多个。您可以使用下面的代码在 TensorFlow 中定义 TFF 计算,该计算将使用 `tf.data.Dataset.reduce` 算子在单个本地数据集中计算温度的平均值。
```
@tff.tf_computation(tff.SequenceType(tf.float32))
def get_local_temperature_average(local_temperatures):
sum_and_count = (
local_temperatures.reduce((0.0, 0), lambda x, y: (x[0] + y, x[1] + 1)))
return sum_and_count[0] / tf.cast(sum_and_count[1], tf.float32)
str(get_local_temperature_average.type_signature)
```
在用 `tff.tf_computation` 装饰的方法的主体中,TFF 序列类型的形式参数简单地表示为行为类似 `tf.data.Dataset` 的对象(即支持相同的属性和方法,它们目前未作为该类型的子类实现,随着 TensorFlow 中对数据集的支持不断发展,这可能会发生变化)。
您可以轻松验证这一点,如下所示。
```
@tff.tf_computation(tff.SequenceType(tf.int32))
def foo(x):
return x.reduce(np.int32(0), lambda x, y: x + y)
foo([1, 2, 3])
```
请记住,与普通的 `tf.data.Dataset` 不同,这些类似数据集的对象是占位符。它们不包含任何元素,因为它们表示抽象的序列类型参数,在具体上下文中使用时将绑定到具体数据。目前,对抽象定义的占位符数据的支持仍有一定局限,在早期的 TFF 中,您可能会遇到某些限制,但在本教程中不必担心这个问题(有关详细信息,请参阅文档页面)。
当在模拟模式下本地执行接受序列的计算时(如本教程所示),您可以将该序列作为 Python 列表进行馈送,如下所示(还可以用其他方式,例如,在 Eager 模式中作为 `tf.data.Dataset` 进行馈送,但现在我们将简单化处理)。
```
get_local_temperature_average([68.5, 70.3, 69.8])
```
与其他 TFF 类型一样,上面定义的序列可以使用 `tff.StructType` 构造函数定义嵌套结构。例如,下面是一个声明计算的方法,该计算接受 `A`、`B` 的序列对,并返回其乘积的和。我们将跟踪语句包含在计算主体中,以便您能够看到 TFF 类型签名如何转换为数据集的 `output_types` 和 `output_shapes`。
```
@tff.tf_computation(tff.SequenceType(collections.OrderedDict([('A', tf.int32), ('B', tf.int32)])))
def foo(ds):
print('element_structure = {}'.format(ds.element_spec))
return ds.reduce(np.int32(0), lambda total, x: total + x['A'] * x['B'])
str(foo.type_signature)
foo([{'A': 2, 'B': 3}, {'A': 4, 'B': 5}])
```
尽管将 `tf.data.Datasets` 用作形式参数在简单场景(如本教程中的场景)中有效,但对它的支持仍有局限且在不断发展。
## 汇总
现在,让我们再次尝试在联合设置中使用 TensorFlow 计算。假设我们有一组传感器,每个传感器有一个本地温度读数的序列。我们可以通过平均传感器的本地平均值来计算全局平均温度,如下所示。
```
@tff.federated_computation(
tff.type_at_clients(tff.SequenceType(tf.float32)))
def get_global_temperature_average(sensor_readings):
return tff.federated_mean(
tff.federated_map(get_local_temperature_average, sensor_readings))
```
请注意,这并非是来自所有客户端的所有本地温度读数的简单平均,因为这需要根据不同客户端本地维护的读数数量权衡其贡献。我们将其作为练习留给读者来更新上面的代码;`tff.federated_mean` 算子接受权重作为可选的第二个参数(预计为联合浮点)。
还要注意,`get_global_temperature_average` 的输入现在变成了*联合浮点序列*。联合序列是我们通常在联合学习中表示设备端数据的方式,序列元素通常表示数据批次(稍后您将看到相关示例)。
```
str(get_global_temperature_average.type_signature)
```
下面是如何用 Python 在数据样本上本地执行计算的方法。请注意,我们现在提供输入的方式是作为 `list` 的 `list`。外部列表在通过 `tff.CLIENTS` 表示的组中对设备进行迭代,内部列表在每个设备的本地序列中对元素进行迭代。
```
get_global_temperature_average([[68.0, 70.0], [71.0], [68.0, 72.0, 70.0]])
```
本教程的第一部分到此结束。我们鼓励您继续学习[第二部分](custom_federated_algorithms_2.ipynb)。
| github_jupyter |
<h1>Table of Contents<span class="tocSkip"></span></h1>
<div class="toc"><ul class="toc-item"><li><span><a href="#Demonstration-Dimensionless-Learning-for-Keyhole-Dynamics" data-toc-modified-id="Demonstration-Dimensionless-Learning-for-Keyhole-Dynamics-1"><span class="toc-item-num">1 </span>Demonstration Dimensionless Learning for Keyhole Dynamics</a></span></li><li><span><a href="#Import-libraries" data-toc-modified-id="Import-libraries-2"><span class="toc-item-num">2 </span>Import libraries</a></span></li><li><span><a href="#Parametric-space-analysis" data-toc-modified-id="Parametric-space-analysis-3"><span class="toc-item-num">3 </span>Parametric space analysis</a></span><ul class="toc-item"><li><span><a href="#Load-dataset" data-toc-modified-id="Load-dataset-3.1"><span class="toc-item-num">3.1 </span>Load dataset</a></span></li><li><span><a href="#Calculate-dimension-matrix" data-toc-modified-id="Calculate-dimension-matrix-3.2"><span class="toc-item-num">3.2 </span>Calculate dimension matrix</a></span></li><li><span><a href="#Calculate-basis-vectors" data-toc-modified-id="Calculate-basis-vectors-3.3"><span class="toc-item-num">3.3 </span>Calculate basis vectors</a></span></li></ul></li><li><span><a href="#Helper-functions" data-toc-modified-id="Helper-functions-4"><span class="toc-item-num">4 </span>Helper functions</a></span></li><li><span><a href="#Best-representation-learning-discovery" data-toc-modified-id="Best-representation-learning-discovery-5"><span class="toc-item-num">5 </span>Best representation learning discovery</a></span></li><li><span><a href="#Reference" data-toc-modified-id="Reference-6"><span class="toc-item-num">6 </span>Reference</a></span></li></ul></div>
# Demonstration Dimensionless Learning for Keyhole Dynamics
- **Authors**: Xiaoyu Xie, Zhengtao Gan
- **Contact**: xiaoyuxie2020@u.northwestern.edu
- **Date**: Oct. 2021
# Import libraries
```
import numpy as np
import matplotlib.pyplot as plt
from numpy.linalg import matrix_rank
from numpy.linalg import inv
import pandas as pd
import pysindy as ps
import random
from sklearn.linear_model import LinearRegression
from sklearn.metrics import r2_score
from scipy.optimize import minimize
```
# Parametric space analysis
Paraemter list:
$f(\eta P,V_s,r_0, C_p, \alpha, \rho, T_l-T_0)=e$
## Load dataset
```
data = np.loadtxt(
open("../dataset/dataset_keyhole.csv","rb"),
delimiter=',',
skiprows=1,
usecols = (2,3,5,7,8,9,12,14)
)
X = data[:,0:7]
Y = data[:,7]
```
## Calculate dimension matrix
Dimension matrix (input):
\begin{equation}
D_{in}=
\left[
\begin{array}{ccc}
2 & 1 & 1 & 2 &2&-3&0\\
-3 & -1 & 0 & -2 &-1&0&0\\
1& 0 & 0 & 0 &0&1&0\\
0& 0 & 0 & -1 &0&0&1\\
\end{array}
\right]
\end{equation}
Dimension matrix (output):
\begin{equation}
D_{out}=
\left[
\begin{array}{ccc}
1 \\
0 \\
0 \\
0 \\
\end{array}
\right]
\end{equation}
```
D_in = np.mat('2,1,1,2,2,-3,0; -3,-1,0,-2,-1,0,0; 1,0,0,0,0,1,0;0,0,0,-1,0,0,1')
D_out = np.mat('1;0;0;0')
D_in_rank = matrix_rank(D_in)
print(D_in_rank)
```
## Calculate basis vectors
Calculate three basis vectors for equation:
$
D_{in}x=0
$
```
Din1 = D_in[:, 0:4]
Din2 = D_in[:, 4:8]
x2 = np.mat('-1; 0; 0')
x1 = -inv(Din1) * Din2 * x2
basis1_in = np.vstack((x1, x2))
print(f'basis1_in: \n{basis1_in}')
x2 = np.mat('0;-1;0')
x1 = -inv(Din1) * Din2 * x2
basis2_in = np.vstack((x1, x2))
print(f'basis2_in: \n{basis2_in}')
x2 = np.mat('0; 0; -1')
x1 = -inv(Din1) * Din2 * x2
basis3_in = np.vstack((x1, x2))
print(f'basis3_in: \n{basis3_in}')
```
# Helper functions
```
def calc_pi(a):
'''
Calculate pi
Note that the best coef for keyhole is [0.5, 1, 1]
'''
coef_pi = 0.5 * basis1_in + a[0] * basis2_in + a[1] * basis3_in
pi_mat = np.exp(np.log(X).dot(coef_pi))
pi = np.squeeze(np.asarray(pi_mat))
return pi
def calc_y(a, w):
'''
Calculate the prediction y using a polynomial function
'''
pi = calc_pi(a)
y = w[0] + w[1] * pi + w[2] * pi**2 + w[3] * pi**3 + w[4] * pi**4 + w[5] * pi**5
return y
def objective(a, w):
'''
Calculate objective(loss)
'''
return np.square(pi2 - calc_y(a, w)).mean()
def ploter(pi1, pi2, iteration):
fig = plt.figure()
plt.scatter(pi1, pi2)
plt.xlabel('pi1', fontsize=16)
plt.ylabel('pi2', fontsize=16)
plt.title(f'iteration: {iteration}', fontsize=24)
plt.show()
```
# Best representation learning discovery
```
random.seed(3)
niter = 3000
ninital = 1
a = np.zeros(2)
w = np.zeros(6)
global pi2
pi2 = Y / X[:,2]
feature_library = ps.PolynomialLibrary(degree=5)
optimizer =LinearRegression(fit_intercept=False)
for j in range(ninital):
a[0] = 2 * random.random()
a[1] = 2 * random.random()
print(f'Initial a={a}')
info = {}
info['initial'] = a
a_history = np.zeros((niter, 2))
model = ps.SINDy(feature_library=feature_library, optimizer=optimizer)
for i in range(niter):
# update coefficient w for polynomials
pi1 = calc_pi(a)
model.fit(pi1, x_dot=pi2);
coeffi = model.coefficients()
w = coeffi[0]
y_recover = calc_y(a, w)
r2 = r2_score(y_recover, pi2)
# update coefficient a for pi
solution = minimize(objective, a, method='BFGS', tol=1e-3, args=w, options={'maxiter':1})
a = solution.x
y_recover = calc_y(a,w)
r2 = r2_score(y_recover, pi2)
a_history[i,:] = a
if i % 1000 == 0:
# Note that the best coef in keyhole case is [0.5, 1, 1]
# here we only optimize the last two coefficients
print(f'Iteration: {i}, coef: {a}, Objective: {objective(a,w)}')
ploter(pi1, pi2, i)
```
# Reference
- [1] **Xie, X.**, Liu, W. K., & **Gan, Z.** (2021). Data-driven discovery of dimensionless numbers and scaling laws from experimental measurements. ArXiv:2111.03583 [Physics]. http://arxiv.org/abs/2111.03583
- [2] **Gan, Z.**, Kafka, O. L., Parab, N., Zhao, C., Fang, L., Heinonen, O., Sun, T., & Liu, W. K. (2021). Universal scaling laws of keyhole stability and porosity in 3D printing of metals. Nature Communications, 12(1), 2379. https://doi.org/10.1038/s41467-021-22704-0
- [3] Saha, S., **Gan, Z.**, Cheng, L., Gao, J., Kafka, O. L., **Xie, X.**, Li, H., Tajdari, M., Kim, H. A., & Liu, W. K. (2021). Hierarchical Deep Learning Neural Network (HiDeNN): An artificial intelligence (AI) framework for computational science and engineering. Computer Methods in Applied Mechanics and Engineering, 373, 113452. https://doi.org/10.1016/j.cma.2020.113452
| github_jupyter |
# Hatch Template!
## Dandelion Voting
1. Percentage of total tokens that have to vote 'yes' to `something` for it to pass.
```
import param
import panel as pn
import pandas as pd
import hvplot.pandas
import holoviews as hv
pn.extension()
class DandelionVoting(param.Parameterized):
total_tokens = param.Number(1e6, constant=True)
support_required = param.Number(0.5, bounds=(0.5,0.9), step=0.01) # Of those who voted, this percent must be yes to pass
minimum_quorum = param.Number(0.02, bounds=(0,1), step=0.01)
class DandelionVoting(param.Parameterized):
total_tokens = param.Number(1e6, constant=True)
minimum_quorum = param.Number(0.02, bounds=(0,1), step=0.01)
support_required = param.Number(0.5, bounds=(0.5,0.9), step=0.01)
days_to_vote_on_proposal = param.Integer(3 + 8 + 24, bounds=(0,100))
days_to_exit_hatch = param.Integer(8)
# vote_buffer_blocks = param.Integer(8, bounds=(0,None))
# vote_execution_delay_blocks = param.Integer(24, bounds=(0,None))
cost_to_make_a_proposal = param.Number(3, step=1, doc="cost to make a proposal")
maximum_number_proposals_per_month = param.Number(10, bounds=(1, 100))
def view(self):
min_yes_tokens = self.support_required * self.minimum_quorum * self.total_tokens
min_blockers = (1 - self.support_required) * self.minimum_quorum * self.total_tokens
votes = pd.DataFrame.from_dict({'Votes': [min_yes_tokens, min_blockers]}, orient='index', columns=['Minimum Tokens to Pass', 'Minimum Tokens for Quorum'])
vote_plot = votes.hvplot.bar(stacked=True, ylim=(0,self.total_tokens)).opts(color=hv.Cycle(['#0F2EEE', '#0b0a15', '#DEFB48']))
return pn.Row(vote_plot, pn.Column("Minimum Tokens to Meet Quorum: ", int(self.minimum_quorum * self.total_tokens), "Minimum Tokens to Pass a Vote: ", int(min_yes_tokens), "Minimum Tokens to Block a Vote: ", int(min_blockers)))
d = DandelionVoting()
pn.Column(d, d.view)
class Hatch(param.Parameterized):
# CSTK Ratio
total_cstk_tokens = param.Number(700000, constant=True)
hatch_oracle_ratio = param.Number(0.005, constant=True)
@param.depends('hatch_oracle_ratio', 'total_cstk_tokens')
def wxdai_range(self):
return pn.Row(pn.Pane("Cap on wxdai staked: "), self.hatch_oracle_ratio * self.total_cstk_tokens)
# Min and Target Goals
min_goal = param.Number(5, bounds=(1,100), step=10)
max_goal = param.Number(1000, bounds=(100,10000), step=50) # Something to consider -> target goal or max goal
# Hatch params
hatch_period = param.Integer(15, bounds=(5, 30), step=2)
hatch_exchange_rate = param.Number() # This needs to be tested and explained -> See the forum post
hatch_tribute = param.Number(0.05, bounds=(0,1))
h = Hatch()
pn.Pane(h)
import pandas as pd
import panel as pn
import os
import hvplot.pandas
APP_PATH = './'
sheets = [
"Total Impact Hours so far",
"IH Predictions",
"#8 Jan 1",
"#7 Dec 18",
"#6 Dec 4",
"#5 Nov 20",
"#4 Nov 6",
"#3 Oct 23",
"#2 Oct 9",
"#1 Sept 24",
"#0 Sept 7 (historic)",
] + [f"#{i} IH Results" for i in range(9)]
sheets = {i:sheet for i, sheet in enumerate(sheets)}
def read_excel(sheet_name="Total Impact Hours so far", header=1, index_col=0, usecols=None) -> pd.DataFrame:
data = pd.read_excel(
os.path.join(APP_PATH, "data", "TEC Praise Quantification.xlsx"),
sheet_name=sheet_name,
engine='openpyxl',
header=header,
index_col=index_col,
usecols=usecols
).reset_index().dropna(how='any')
return data
## Tests
total_impact_hours = read_excel()
impact_hour_data = read_excel(sheet_name="IH Predictions", header=0, index_col=0, usecols='A:I').drop(index=19)
pn.Row(impact_hour_data.hvplot.table(), total_impact_hours.hvplot.table())
import numpy as np
class ImpactHours(param.Parameterized):
max_ih_rate = param.Number(0.01, bounds=(0,200))
expected_raise_per_ih = param.Number(0.012, bounds=(0,20))
@param.depends('max_ih_rate', 'expected_raise_per_ih')
def impact_hours_rewards(self):
x = np.linspace(h.min_goal, h.max_goal)
R = self.max_ih_rate
m = self.expected_raise_per_ih
H = total_impact_hours['Impact Hours'].sum()
y = [R* (x / (x + m*H)) for x in x]
df = pd.DataFrame([x,y]).T
df.columns = ['x','y']
return df.hvplot(x='x')
i = ImpactHours()
pn.Row(i, i.impact_hours_rewards)
pn.Row(d, h, i)
pn.Row(d.view, i.impact_hours_rewards)
```
# Target/Expected Goals
```
class CommunityParticipation(param.Parameterized)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
path="/content/drive/MyDrive/Research/cifar_entropy/sixth_layer_with_entropy_1235_k01/"
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
#foreground_classes = {'bird', 'cat', 'deer'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck'}
#background_classes = {'plane', 'car', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])#.type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx])#.type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
np.random.seed(i)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=64, kernel_size=3, padding=0)
self.conv4 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, padding=0)
self.conv6 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32,track_running_stats=False)
self.batch_norm2 = nn.BatchNorm2d(64,track_running_stats=False)
self.batch_norm3 = nn.BatchNorm2d(256,track_running_stats=False)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(256,64)
self.fc2 = nn.Linear(64, 32)
self.fc3 = nn.Linear(32, 10)
self.fc4 = nn.Linear(10, 1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,256, 3,3], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
ftr = torch.zeros([batch,9,256,3,3])
y = y.to("cuda")
x = x.to("cuda")
ftr = ftr.to("cuda")
for i in range(9):
out,ftrs = self.helper(z[:,i])
#print(out.shape)
x[:,i] = out
ftr[:,i] = ftrs
log_x = F.log_softmax(x,dim=1) # log_alpha
x = F.softmax(x,dim=1)
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],ftr[:,i])
return x, log_x, y #alpha, avg_data
def helper(self, x):
#x1 = x
#x1 =x
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm3(x))
x = self.conv6(x)
x1 = F.tanh(x)
x = F.relu(x)
x = self.pool(x)
x = x.view(x.size(0), -1)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.dropout2(x)
x = F.relu(self.fc3(x))
x = self.fc4(x)
x = x[:,0]
#print(x.shape)
return x,x1
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=256, out_channels=128, kernel_size=3, padding=1)
self.conv2 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.conv3 = nn.Conv2d(in_channels=128, out_channels=256, kernel_size=3, padding=1)
self.conv4 = nn.Conv2d(in_channels=256, out_channels=256, kernel_size=3, padding=1)
self.conv5 = nn.Conv2d(in_channels=256, out_channels=512, kernel_size=3, padding=1)
self.conv6 = nn.Conv2d(in_channels=512, out_channels=512, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2,padding=1)
self.batch_norm1 = nn.BatchNorm2d(128,track_running_stats=False)
self.batch_norm2 = nn.BatchNorm2d(256,track_running_stats=False)
self.batch_norm3 = nn.BatchNorm2d(512,track_running_stats=False)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.global_average_pooling = nn.AvgPool2d(kernel_size=2)
self.fc1 = nn.Linear(512,128)
# self.fc2 = nn.Linear(128, 64)
# self.fc3 = nn.Linear(64, 10)
self.fc2 = nn.Linear(128, 3)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm3(x))
x = (F.relu(self.conv6(x)))
x = self.pool(x)
#print(x.shape)
x = self.global_average_pooling(x)
x = x.squeeze()
#x = x.view(x.size(0), -1)
#print(x.shape)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
#x = F.relu(self.fc2(x))
#x = self.dropout2(x)
#x = F.relu(self.fc3(x))
x = self.fc2(x)
return x
torch.manual_seed(1235)
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
torch.manual_seed(1235)
classify = Classification().double()
classify = classify.to("cuda")
for params in focus_net.parameters():
params.requires_grad =True
for params in classify.parameters():
params.requires_grad = True
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
np.random.seed(i+30000)
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
# criterion_classify = nn.CrossEntropyLoss()
optimizer_focus = optim.SGD(focus_net.parameters(), lr=0.01, momentum=0.9)
optimizer_classify = optim.SGD(classify.parameters(), lr=0.01, momentum=0.9)
criterion = nn.CrossEntropyLoss()
def my_cross_entropy(x, y,alpha,log_alpha,k):
loss = criterion(x,y)
b = -1.0* alpha * log_alpha
b = torch.mean(torch.sum(b,dim=1))
closs = loss
entropy = b
loss = (1-k)*loss + ((k)*b)
return loss,closs,entropy
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, _ ,avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas,_, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 150
k = 0.01
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
ep_loss=[]
ep_ce=[]
ep_ent =[]
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
running_cross_entropy = 0.0
running_entropy = 0.0
cnt=0
iteration = desired_num // batch
epoch_loss = []
epoch_ce = []
epoch_entropy = []
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas,log_alpha, avg_images = focus_net(inputs)
outputs = classify(avg_images)
# outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
#loss = criterion_classify(outputs, labels)
loss,c_e,entropy = my_cross_entropy(outputs, labels,alphas,log_alpha,k)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
running_cross_entropy += c_e.item()
running_entropy += entropy.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f cross_entropy: %.3f entropy: %.3f' %(epoch + 1, cnt + 1, running_loss / mini,running_cross_entropy/mini,running_entropy/mini))
epoch_loss.append(running_loss/mini)
epoch_ce.append(running_cross_entropy/mini)
epoch_entropy.append(running_entropy/mini)
running_loss = 0.0
running_cross_entropy = 0.0
running_entropy = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if epoch % 5 == 0:
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, _,avg_images = focus_net(inputs)
outputs = classify(avg_images)
#outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
if(np.mean(epoch_loss) <= 0.02):
break;
ep_loss.append(np.mean(epoch_loss))
ep_ce.append(np.mean(epoch_ce))
ep_ent.append(np.mean(epoch_entropy))
print('Finished Training')
torch.save(focus_net.state_dict(),path+"weights_focus_01.pt")
torch.save(classify.state_dict(),path+"weights_classify_01.pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
len(col1)
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
plt.figure(figsize=(6,6))
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig(path+"train_0.png",bbox_inches="tight")
plt.savefig(path+"train_0.pdf",bbox_inches="tight")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend()#loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig(path+"train_0_1.png",bbox_inches="tight")
plt.savefig(path+"train_0_1.pdf",bbox_inches="tight")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
# plt.legend()#loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig(path+"train_0_2.png",bbox_inches="tight")
plt.savefig(path+"train_0_2.pdf",bbox_inches="tight")
plt.show()
df_test
plt.figure(figsize=(6,6))
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend( loc='center left', bbox_to_anchor=(1, 0.5) )
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig(path+"test_0.png",bbox_inches="tight")
plt.savefig(path+"test_0.pdf",bbox_inches="tight")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend()#loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig(path+"test_0_1.png",bbox_inches="tight")
plt.savefig(path+"test_0_1.pdf",bbox_inches="tight")
plt.show()
plt.figure(figsize=(6,6))
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
# plt.legend()#loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig(path+"test_0_2.png",bbox_inches="tight")
plt.savefig(path+"test_0_2.pdf",bbox_inches="tight")
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
focus_net.eval()
classify.eval()
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas,_, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %.3f %%' % ( 100*correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %.3f %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %.3f %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %.3f %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %.3f %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
focus_net.eval()
classify.eval()
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, _,avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %.3f %%' % ( 100.0 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %.3f %%" % (focus_true_pred_true , (100.0 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %.3f %%" % (focus_false_pred_true, (100.0 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %.3f %%" %( focus_true_pred_false , ( 100.0 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %.3f %%" % (focus_false_pred_false, ( 100.0 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
focus_net.eval()
classify.eval()
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas,_, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %.3f %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
focus_net.eval()
classify.eval()
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, _,avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %.3f %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
```
| github_jupyter |
# curve_cifar10.py
```
!python curve_cifar10.py --model 'c6f2_' --test_pth models/cifar10/10152209_c6f2__BCP_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth
```
# drawing curve
- use the last output list to plot the eps-acc curve in the paper.
```
import matplotlib.pyplot as plt
import numpy as np
acc_list = 100-100*np.array([0.34579998254776, 0.3596999943256378, 0.37619999051094055, 0.3919000029563904, 0.40709999203681946, 0.42309999465942383, 0.43879997730255127, 0.45569998025894165, 0.47599998116493225, 0.493399977684021, 0.5144000053405762, 0.5317999720573425, 0.5507000088691711, 0.5703999996185303, 0.5867999792098999, 0.6040999889373779, 0.6189999580383301, 0.6392999887466431, 0.6565999984741211, 0.6746000051498413, 0.689799964427948, 0.7234999537467957, 0.757099986076355, 0.786300003528595, 0.8154000043869019, 0.842799961566925, 0.8675000071525574, 0.8894000053405762, 0.9032999873161316]
)
eps_list = [0,4/255,8/255,12/255,16/255,20/255,24/255,28/255,32/255,36/255,40/255,44/255,48/255,52/255,56/255,60/255,64/255,68/255,72/255,76/255,80/255,88/255,96/255,104/255,112/255,120/255,128/255,136/255,142/255]
import matplotlib.pyplot as plt
plt.figure(figsize=(4,2.7))
plt.vlines(x=36/255, ymin=0, ymax=100, color='gray', alpha=1, linewidth=1)
ax1, =plt.plot(eps_list, acc_list, 'r')
plt.xlim(0,142/255)
plt.ylim(0,70)
plt.legend(handles=[ax1], labels=['BCP'],loc='upper right')
plt.xlabel(r'$\epsilon_{eval}$')
plt.ylabel('Verification Acc')
plt.tight_layout()
plt.show()
!nvidia-smi
!python evaluate_cifar10.py --model 'c6f2' --test_pth models/cifar10/1015_ongoing2_c6f2_BCP_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_pgd_False_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth --cuda_ids 5
!python evaluate_cifar10.py --model 'c6f2' --test_pth log/bcp/1015_c6f2_BCP_bce_False_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_pgd_False_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth
!python curve_cifar10.py --model 'c6f2' --test_pth log/bcp/1015_c6f2_BCP_bce_False_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_pgd_False_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth
!python curve_cifar10.py --model 'c6f2' --test_pth log/bcp/1015_c6f2_BCP_bce_False_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_pgd_False_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth
!python curve_cifar10.py --model 'c6f2' --test_pth log/bcp/1015_c6f2_BCP_bce_False_data_cifar10_epochs_200_epsilon_train_0.1411764705882353_lr_0.001_opt_adam_opt_iter_1_pgd_False_rampup_121_sniter_1_starting_kappa_1.0_train_method_BCP_warmup_10_best.pth
```
| github_jupyter |
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm as tqdm
%matplotlib inline
import torch
import torchvision
import torchvision.transforms as transforms
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import random
from google.colab import drive
drive.mount('/content/drive')
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
type(trainset.targets)
type(trainset.data)
index1 = [np.where(np.array(trainset.targets)==0)[0] , np.where(np.array(trainset.targets)==1)[0], np.where(np.array(trainset.targets)==2)[0] ]
index1 = np.concatenate(index1,axis=0)
len(index1) #15000
#index1
disp = np.array(trainset.targets)
len(trainset.targets)
true_data_count = 15000
epochs= 50
indices = np.random.choice(index1,true_data_count)
_,count = np.unique(disp[indices],return_counts=True)
print(count)
# index = np.where(np.logical_and(np.logical_and(np.array(trainset.targets)!=0, np.array(trainset.targets)!=1), np.array(trainset.targets)!=2))[0] #47335
# len(index)
# values = np.random.choice([0,1,2],size= len(index)) #labeling others as 0,1,2
# print(sum(values ==0),sum(values==1), sum(values==2))
# trainset.data = torch.tensor( trainset.data )
# trainset.targets = torch.tensor(trainset.targets)
# trainset.data = np.concatenate((trainset.data[indices],trainset.data[index]))
# trainset.targets = np.concatenate((np.array(trainset.targets)[indices],values))
x = trainset.data
y = np.array(trainset.targets)
trainset.data = x[indices]
trainset.targets = y[indices]
len(y[indices])
# mnist_trainset.targets[index] = torch.Tensor(values).type(torch.LongTensor)
j =85 # Without Shuffle upto True Training numbers correct , after that corrupted
print(plt.imshow(trainset.data[j]),trainset.targets[j])
trainloader = torch.utils.data.DataLoader(trainset, batch_size=256,shuffle=True, num_workers=2)
testloader = torch.utils.data.DataLoader(testset, batch_size=256,shuffle=False, num_workers=2)
classes = ('zero', 'one','two')
dataiter = iter(trainloader)
images, labels = dataiter.next()
images[:4].shape
# def imshow(img):
# img = img / 2 + 0.5 # unnormalize
# npimg = img.numpy()
# plt.imshow(np.transpose(npimg, (1, 2, 0)))
# plt.show()
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
imshow(torchvision.utils.make_grid(images[:10]))
print('GroundTruth: ', ' '.join('%5s' % classes[labels[j]] for j in range(10)))
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
class Conv_module(nn.Module):
def __init__(self,inp_ch,f,s,k,pad):
super(Conv_module,self).__init__()
self.inp_ch = inp_ch
self.f = f
self.s = s
self.k = k
self.pad = pad
self.conv = nn.Conv2d(self.inp_ch,self.f,k,stride=s,padding=self.pad)
self.bn = nn.BatchNorm2d(self.f)
self.act = nn.ReLU()
def forward(self,x):
x = self.conv(x)
x = self.bn(x)
x = self.act(x)
return x
class inception_module(nn.Module):
def __init__(self,inp_ch,f0,f1):
super(inception_module, self).__init__()
self.inp_ch = inp_ch
self.f0 = f0
self.f1 = f1
self.conv1 = Conv_module(self.inp_ch,self.f0,1,1,pad=0)
self.conv3 = Conv_module(self.inp_ch,self.f1,1,3,pad=1)
#self.conv1 = nn.Conv2d(3,self.f0,1)
#self.conv3 = nn.Conv2d(3,self.f1,3,padding=1)
def forward(self,x):
x1 = self.conv1.forward(x)
x3 = self.conv3.forward(x)
#print(x1.shape,x3.shape)
x = torch.cat((x1,x3),dim=1)
return x
class downsample_module(nn.Module):
def __init__(self,inp_ch,f):
super(downsample_module,self).__init__()
self.inp_ch = inp_ch
self.f = f
self.conv = Conv_module(self.inp_ch,self.f,2,3,pad=0)
self.pool = nn.MaxPool2d(3,stride=2,padding=0)
def forward(self,x):
x1 = self.conv(x)
#print(x1.shape)
x2 = self.pool(x)
#print(x2.shape)
x = torch.cat((x1,x2),dim=1)
return x,x1
class inception_net(nn.Module):
def __init__(self):
super(inception_net,self).__init__()
self.conv1 = Conv_module(3,96,1,3,0)
self.incept1 = inception_module(96,32,32)
self.incept2 = inception_module(64,32,48)
self.downsample1 = downsample_module(80,80)
self.incept3 = inception_module(160,112,48)
self.incept4 = inception_module(160,96,64)
self.incept5 = inception_module(160,80,80)
self.incept6 = inception_module(160,48,96)
self.downsample2 = downsample_module(144,96)
self.incept7 = inception_module(240,176,60)
self.incept8 = inception_module(236,176,60)
self.pool = nn.AvgPool2d(5)
self.linear = nn.Linear(236,10)
def forward(self,x):
x = self.conv1.forward(x)
#act1 = x
x = self.incept1.forward(x)
#act2 = x
x = self.incept2.forward(x)
#act3 = x
x,act4 = self.downsample1.forward(x)
x = self.incept3.forward(x)
#act5 = x
x = self.incept4.forward(x)
#act6 = x
x = self.incept5.forward(x)
#act7 = x
x = self.incept6.forward(x)
#act8 = x
x,act9 = self.downsample2.forward(x)
x = self.incept7.forward(x)
#act10 = x
x = self.incept8.forward(x)
#act11 = x
#print(x.shape)
x = self.pool(x)
#print(x.shape)
x = x.view(-1,1*1*236)
x = self.linear(x)
return x
inc = inception_net()
inc = inc.to("cuda")
criterion_inception = nn.CrossEntropyLoss()
optimizer_inception = optim.SGD(inc.parameters(), lr=0.01, momentum=0.9)
acti = []
loss_curi = []
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_inception.zero_grad()
# forward + backward + optimize
outputs = inc(inputs)
loss = criterion_inception(outputs, labels)
loss.backward()
optimizer_inception.step()
# print statistics
running_loss += loss.item()
mini_batch = 25
if i % mini_batch == mini_batch-1: # print every 50 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / mini_batch))
ep_lossi.append(running_loss/mini_batch) # loss per minibatch
running_loss = 0.0
loss_curi.append(np.mean(ep_lossi)) #loss per epoch
# if (epoch%5 == 0):
# _,actis= inc(inputs)
# acti.append(actis)
print('Finished Training')
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = inc(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %d %%' % (true_data_count, 100 * correct / total))
total,correct
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= inc(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
out = np.concatenate(out,axis=0)
pred = np.concatenate(pred,axis=0)
index = np.logical_or(np.logical_or(out ==1,out==0),out == 2)
print(index.shape)
acc = sum(out[index] == pred[index])/sum(index)
print('Accuracy of the network on the 10000 test images: %d %%' % (
100*acc))
np.unique(out[index],return_counts = True) #== pred[index])
np.unique(pred[index],return_counts = True) #== pred[index])
sum(out[index] == pred[index])
cnt = np.zeros((3,3))
true = out[index]
predict = pred[index]
for i in range(len(true)):
cnt[true[i]][predict[i]] += 1
cnt
# torch.save(inc.state_dict(),"/content/drive/My Drive/Research/CIFAR Random/model_True_"+str(true_data_count)+"_epoch_"+str(epochs)+".pkl")
true_data_count
```
| github_jupyter |
# RSA Decryption
- Ascii plaintext encoded using PKCS1.5
# PKCS1.5
```
RSA Modulo Size: e.g 2048 bits or 256 bytes
+------+------------------------------+------+--------------------+
| 0x02 | RANDOM NONZERO DIGITS | 0x00 | MESSAGE IN ASCII |
+------+------------------------------+------+--------------------+
```
```
# Given
message = "Factoring lets us break RSA."
ct_string = "22096451867410381776306561134883418017410069787892831071731839143676135600120538004282329650473509424343946219751512256465839967942889460764542040581564748988013734864120452325229320176487916666402997509188729971690526083222067771600019329260870009579993724077458967773697817571267229951148662959627934791540"
E = 65537
N_string = "179769313486231590772930519078902473361797697894230657273430081157732675805505620686985379449212982959585501387537164015710139858647833778606925583497541085196591615128057575940752635007475935288710823649949940771895617054361149474865046711015101563940680527540071584560878577663743040086340742855278549092581"
p_string = "13407807929942597099574024998205846127479365820592393377723561443721764030073662768891111614362326998675040546094339320838419523375986027530441562135724301"
q_string = "13407807929942597099574024998205846127479365820592393377723561443721764030073778560980348930557750569660049234002192590823085163940025485114449475265364281"
from os import urandom
from gmpy2 import mpz
from gmpy2 import invert, t_mod, mul, powmod
def decrypt(y, d, N):
return powmod(y, d, N)
def encrypt(x, e, N):
return powmod(x, e, N)
def decrypt_pipeline(c_string, d, N):
m_decimal = decrypt(mpz(c_string), d, N)
m_hex = hex(m_decimal)[2:]
m = m_hex.split('00') #assumes correct format
return bytes.fromhex(m[1]).decode('utf8')
def encrypt_pipeline(message, e, N):
raw_message = bytes(message, 'utf8')
TOTAL_LENGTH = 128
APPENDLENGTH = TOTAL_LENGTH - len(raw_message) - 2
randomhexstring = urandom(APPENDLENGTH).hex()
final_bytes = bytes.fromhex('02' + randomhexstring + '00') + raw_message
final_decimal = mpz(int.from_bytes(final_bytes, 'big'))
return str(encrypt(final_decimal, e, N))
N = mpz(N_string)
p = mpz(p_string)
q = mpz(q_string)
c = mpz(ct_string)
e = mpz(E)
# compute d
phiN = N - p - q + 1
D = invert(e, phiN)
d = mpz(D)
# d * e mod phi(N) = 1
# where phi(N) = N - p - q + 1
assert t_mod(mul(d, e), phiN)
print(decrypt_pipeline(ct_string, d, N))
c = encrypt_pipeline(message, e, N)
m = decrypt_pipeline(c, d, N)
print(m)
```
| github_jupyter |
This notebook shows the construction of simple feed-forward normalizing flows.
```
from lib.toy_data import inf_train_gen
from lib.visualize_flow import plt_flow_2D
import matplotlib.pyplot as plt
import NF
from timeit import default_timer as timer
import torch
%matplotlib inline
```
### Building your first normalizing flow
Let's create our first normalizing flow: an affine (masked) autoregressive flow.
Building such flow is very simple, you should first define the main building blocks of your flow. In this case these are the conditioner (autoregressive) and the normalizer (affine). Once this is done, you have to combine the conditioner and the normalizer into a flow step. You may do that multiple times if you want multiple steps in your flow, here we keep it simple and use only one step.
Once you have a list of normalizing flow steps you may create a normalizing flow by combining the list with a base distribution (often an isotropic Normal).
```
conditioner = NF.AutoregressiveConditioner(2, [50, 50, 50], 2, 8)
normalizer = NF.AffineNormalizer()
flow_steps = [NF.NormalizingFlowStep(conditioner, normalizer)]
flow = NF.FCNormalizingFlow(flow_steps, NF.NormalLogDensity())
opt = torch.optim.Adam(flow.parameters(), 1e-3, weight_decay=1e-5)
```
That's it, congratulations you built your first normalizing flow! ;)
Let's now check how we may train the flow on some data. We will create a function as we will train other flow architectures in the future, fortunately they all may be trained in the same way!
```
def train_and_plot(flow, opt, toy="conditionnal8gaussians", nb_epoch=1000):
time_tot = 0.
for epoch in range(nb_epoch):
loss_tot = 0
start = timer()
# We get some data, (we place ourself in an ideal setting of infinite data).
cur_x, context = inf_train_gen(toy, batch_size=100)
# We compute the log-likelihood as well as the base variable, check NormalizingFlow class for other possibilities
ll, z = flow.compute_ll(torch.tensor(cur_x).float(), torch.tensor(context).float())
# Here we would like to maximize the log-likelihood of our model!
loss = -ll.mean()
opt.zero_grad()
loss.backward()
opt.step()
end = timer()
time_tot += end - start
if epoch % 10 == 0:
print("Approximate time left : {:2f}s - Loss last batch: {:4f}".format(time_tot/(epoch + 1) * (nb_epoch - (epoch + 1)), loss.item()),
end="\r", flush=True)
# Let's check the result!
fig = plt.figure(figsize=(18, 8*6))
def compute_ll(x, i=0):
context = torch.zeros(x.shape[0], 8)
context[:, i] = 1.
return flow.compute_ll(x, context)
for i in range(8):
ax = plt.subplot(8, 3, 1 + 3*i, aspect="equal")
ax.set_title("Learned density")
plt_flow_2D(lambda x: compute_ll(x, i), ax, 100, range_xy=[[-4, 4], [-4, 4]])
ax = plt.subplot(8, 3, 2 + 3*i, aspect="equal")
ax.set_title("Generated samples")
context = torch.zeros(1000, 8)
context[:, i] = 1.
ax.set_xlim(-4.5, 4.5)
ax.set_ylim(-4.5, 4.5)
fake_samples = flow.invert(torch.randn(1000, 2), context).detach().numpy()
ax.scatter(fake_samples[:, 0], fake_samples[:, 1], alpha=1., marker=".")
ax = plt.subplot(8, 3, 3 + 3*i, aspect="equal")
ax.set_xlim(-4.5, 4.5)
ax.set_ylim(-4.5, 4.5)
ax.set_title("Real samples")
samples, _ = inf_train_gen(toy, batch_size=1000)
ax.scatter(samples[:, 0], samples[:, 1], alpha=1., marker=".")
```
And now let us train this one-step affine autoregressive flow!
```
plt.figure()
train_and_plot(flow, opt, nb_epoch=1000)
plt.show()
```
Mmmh not great!
One step does not seem to be enough, let's build a 5-steps affine autoregressive flow. For that you may just reproduce the example above with multiple steps but you can just use one of our factories as follows:
```
conditioner_type = NF.AutoregressiveConditioner
conditioner_args = {"in_size": 2, "hidden": [150, 150, 150], "out_size": 2}
normalizer_type = NF.AffineNormalizer
normalizer_args = {}
nb_flow = 5
flow = NF.buildFCNormalizingFlow(nb_flow, conditioner_type, conditioner_args, normalizer_type, normalizer_args)
opt = torch.optim.Adam(flow.parameters(), 1e-3, weight_decay=1e-5)
```
The flow here is made of 5 steps, each of them being composed of an autoregressive conditioner and an affine normalizer (also denoted a transformer in the flow litterature). The parameters are not shared between flow steps!
```
for i in range(100):
plt.figure()
train_and_plot(flow, opt, nb_epoch=1000)
plt.show()
```
These results are already better but modeling multi-modal distributions with affine normalizing flows is known to be impossible. A possible solution is to replace the affine transformations by monotonic ones, this has been proven to lead to universal density approximators. In the following example we use unconstrained monotonic neural networks to parameterize the monotonic transformation and combine it with an autoregressive conditioner.
```
conditioner = NF.AutoregressiveConditioner(2, [100, 100, 100], 10)
normalizer = NF.MonotonicNormalizer(integrand_net=[50, 50, 50], cond_size=10, nb_steps=50, solver="CCParallel")
flow_steps = [NF.NormalizingFlowStep(conditioner, normalizer)]
flow = NF.FCNormalizingFlow(flow_steps, NormalLogDensity())
opt = torch.optim.Adam(flow.parameters(), 1e-3, weight_decay=1e-5)
for i in range(100):
plt.figure()
train_and_plot(flow, opt, nb_epoch=1000)
plt.show()
```
We have only shown how to use autoregressive conditioners, however you may find other famous conditioners in the library such as coupling conditioners or graphical conditioners.
| github_jupyter |
# Intermediate Linear Algebra - Eigenvalues & Eigenvectors
### Key Equation: $Ax = \lambda b ~~ \text{for} ~~ n \times n $
## Transformations
So what really happens when we multiply the $A$ matrix with a vector $x$
Lets say we have a vector - $x$
$$ x = \begin{bmatrix} -1 \\ 1 \end{bmatrix} $$
What happens when we multiply by a matrix - $A$
$$ A = \begin{bmatrix} 6 & 2 \\ 2 & 6 \end{bmatrix} $$
$$ Ax = \begin{bmatrix} 6 & 2 \\ 2 & 6 \end{bmatrix} \begin{bmatrix} -1 \\ 1 \end{bmatrix} = \begin{bmatrix} -4 \\ 4 \end{bmatrix} $$
$$ Ax = 4Ix $$
$$ Ax = 4x $$
So this particular matrix has just scaled our original vector. It is a scalar transformation. Other matrices can do reflection, rotation and any arbitary transformation in the same 2d space for n = 2.
Lets see what has happened through code.
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
plt.style.use('fivethirtyeight')
plt.rcParams['figure.figsize'] = (10, 6)
def vector_plot (vector):
X,Y,U,V = zip(*vector)
C = [1,1,2,2]
plt.figure()
ax = plt.gca()
ax.quiver(X,Y,U,V,C, angles='xy',scale_units='xy',scale=1)
ax.set_xlim([-6,6])
ax.set_ylim([-6,6])
plt.axhline(0, color='grey', linewidth=1)
plt.axvline(0, color='grey', linewidth=1)
plt.axes().set_aspect('equal')
plt.draw()
A = np.array([[ 6 , 2],
[ 2 , 6]])
x = np.array([[-1],
[1]])
v = A.dot(x)
# All the vectors start at 0, 0
vAX = np.r_[[0,0],A[:,0]]
vAY = np.r_[[0,0],A[:,1]]
vx = np.r_[[0,0],x[:,0]]
vv = np.r_[[0,0],v[:,0]]
vector_plot([vAX, vAY, vx, vv])
```
## Solving Equation $Ax=\lambda x$
### Special Case: $Ax = 0$
So far we have been solving the equation $Ax = b$. Let us just look at special case when $b=0$.
$$ Ax =0 $$
If $A^{-1}$ exists (the matrix is non-singular and invertable), then the solution is trival
$$ A^{-1}Ax =0 $$
$$ x = 0$$
If $A^{-1}$ does not exist, then there may be infinitely many other solutions $x$. And since $A^{-1}$ is a singular matrix then
$$||A|| = 0 $$
### General Case
The second part of linear algebra is solving the equation, for a given $A$ -
$$ Ax = \lambda x$$
Note that both $x$ and $\lambda$ are unknown in this equation. For all solutions of them:
$$ \text{eigenvalues} = \lambda $$
$$ \text{eigenvectors} = x $$
## Calculating Eigenvalues
So let us first solve this for $\lambda$ :
$$ Ax = \lambda Ix $$
$$ (A-\lambda I)x = 0 $$
So for non-trivial solution of $x$, $A$ should be singular:
$$ ||A - \lambda I|| = 0 $$
## For 2 x 2 Matrix
Let us use the sample $A$ vector:
$$ A = \begin{bmatrix}3 & 1\\ 1 & 3\end{bmatrix} $$
So our equation becomes:
$$ \begin{bmatrix}3 & 1\\ 1 & 3\end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}\lambda & 0\\ 0 & \lambda \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} $$
$$ \begin{bmatrix}3 - \lambda & 1\\ 1 & 3 - \lambda \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = 0 $$
So for a singular matrix:
$$ \begin{Vmatrix}3 - \lambda & 1\\ 1 & 3 - \lambda \end{Vmatrix} = 0 $$
$$ (3 - \lambda)^2 - 1 = 0 $$
$$ \lambda^2 - 6\lambda + 8 = 0 $$
$$ (\lambda - 4)(\lambda - 2) = 0 $$
$$ \lambda_1 = 2, \lambda_2 = 4 $$
$$||A|| = \lambda_{1} \lambda_{2} $$
## Calculating Eigenvectors
For $\lambda = 2$,
$$ \begin{bmatrix}3 - \lambda & 1\\ 1 & 3 - \lambda \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}1 & 1\\ 1 & 1 \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = 0 $$
So one simple solution is:
$$ \begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}-1 \\ 1\end{bmatrix} $$
For $\lambda = 4$,
$$ \begin{bmatrix}3 - \lambda & 1\\ 1 & 3 - \lambda \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}-1 & 1\\ 1 & -1 \end{bmatrix} \begin{bmatrix}x \\ y\end{bmatrix} = 0 $$
So one simple solution is:
$$ \begin{bmatrix}x \\ y\end{bmatrix} = \begin{bmatrix}1 \\ 1\end{bmatrix} $$
The eigenvectors are orthonormal to each other in this case.
## Vector Representation (2x2)
A vector representation for this is now:
$$ \begin{bmatrix}3 \\ 1\end{bmatrix} x + \begin{bmatrix}1 \\ 3\end{bmatrix} y = \begin{bmatrix} \lambda \\ 0 \end{bmatrix} x + \begin{bmatrix} 0 \\ \lambda \end{bmatrix} y $$
Now we need to draw these vectors and see the result
```
A = np.array([[ 3 , 1],
[ 1 , 3]])
eigen_val, eigen_vec = np.linalg.eig(A)
eigen_val
eigen_vec
eigen_vec[:,0]
# All the vectors start at 0, 0
vX1 = np.r_[[0,0],A[:,0]]
vY1 = np.r_[[0,0],A[:,1]]
vE1 = np.r_[[0,0],eigen_vec[:,0]] * 2
vE2 = np.r_[[0,0],eigen_vec[:,1]] * 2
vector_plot([vX1, vY1, vE1, vE2])
```
# 3 x 3 Matrix
Let us write it in the form
$$ Ax = \lambda x $$
$$ \begin{bmatrix}1 & 1 & 1 \\ 3 & 8 & 1 \\ 5 & -4 & 3\end{bmatrix}\begin{bmatrix} x \\y \\ z\end{bmatrix}= \lambda \begin{bmatrix} x\\ y \\ x \end{bmatrix} $$
```
f = np.matrix([[1,1,1],
[3,8,1],
[5,-4,3]])
np.linalg.eig(f)
```
## Exercises on Eigenvalues and Eigenvectors
$$ U = \begin{bmatrix}1 & 1 & 0 \\ 0 & 2 & 0 \\ 0 & -1 & 4\end{bmatrix}$$
$$ S = \begin{bmatrix}3 & 1 & 2 \\ 1 & 4 & 5 \\ 2 & 5 & 6 \end{bmatrix}$$
$$ T = \begin{bmatrix}5 & 8 \\ 0 & 5 \end{bmatrix}$$
Write the matrix as np.matrix and find the Eigenvalues and Eigenvectors?
## Symmetric Matrix and Eigen Vectors
$$ V = \begin{bmatrix}1 & 2 & -1 \\ 2 & -1 & -2 \\ -1 & -2 & 1\end{bmatrix}$$
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
import os
os.sys.path.insert(0, '/home/schirrmr/braindecode/code/braindecode/')
```
# Amplitude Perturbation Visualization
In this tutorial, we show how to use perturbations of the input amplitudes to learn something about the trained convolutional networks. For more background, see [Deep learning with convolutional neural networks for EEG decoding and visualization](https://arxiv.org/abs/1703.05051), Section A.5.2.
First we will do some cross-subject decoding, again using the [Physiobank EEG Motor Movement/Imagery Dataset](https://www.physionet.org/physiobank/database/eegmmidb/), this time to decode imagined left hand vs. imagined right hand movement.
<div class="alert alert-warning">
This tutorial might be very slow if you are not using a GPU.
</div>
## Enable logging
```
import logging
import importlib
importlib.reload(logging) # see https://stackoverflow.com/a/21475297/1469195
log = logging.getLogger()
log.setLevel('INFO')
import sys
logging.basicConfig(format='%(asctime)s %(levelname)s : %(message)s',
level=logging.INFO, stream=sys.stdout)
```
## Load data
```
import mne
import numpy as np
from mne.io import concatenate_raws
from braindecode.datautil.signal_target import SignalAndTarget
# First 50 subjects as train
physionet_paths = [ mne.datasets.eegbci.load_data(sub_id,[4,8,12,]) for sub_id in range(1,51)]
physionet_paths = np.concatenate(physionet_paths)
parts = [mne.io.read_raw_edf(path, preload=True,stim_channel='auto')
for path in physionet_paths]
raw = concatenate_raws(parts)
picks = mne.pick_types(raw.info, meg=False, eeg=True, stim=False, eog=False,
exclude='bads')
events = mne.find_events(raw, shortest_event=0, stim_channel='STI 014')
# Read epochs (train will be done only between 1 and 2s)
# Testing will be done with a running classifier
epoched = mne.Epochs(raw, events, dict(hands=2, feet=3), tmin=1, tmax=4.1, proj=False, picks=picks,
baseline=None, preload=True)
# 51-55 as validation subjects
physionet_paths_valid = [mne.datasets.eegbci.load_data(sub_id,[4,8,12,]) for sub_id in range(51,56)]
physionet_paths_valid = np.concatenate(physionet_paths_valid)
parts_valid = [mne.io.read_raw_edf(path, preload=True,stim_channel='auto')
for path in physionet_paths_valid]
raw_valid = concatenate_raws(parts_valid)
picks_valid = mne.pick_types(raw_valid.info, meg=False, eeg=True, stim=False, eog=False,
exclude='bads')
events_valid = mne.find_events(raw_valid, shortest_event=0, stim_channel='STI 014')
# Read epochs (train will be done only between 1 and 2s)
# Testing will be done with a running classifier
epoched_valid = mne.Epochs(raw_valid, events_valid, dict(hands=2, feet=3), tmin=1, tmax=4.1, proj=False, picks=picks_valid,
baseline=None, preload=True)
train_X = (epoched.get_data() * 1e6).astype(np.float32)
train_y = (epoched.events[:,2] - 2).astype(np.int64) #2,3 -> 0,1
valid_X = (epoched_valid.get_data() * 1e6).astype(np.float32)
valid_y = (epoched_valid.events[:,2] - 2).astype(np.int64) #2,3 -> 0,1
train_set = SignalAndTarget(train_X, y=train_y)
valid_set = SignalAndTarget(valid_X, y=valid_y)
```
## Create the model
We use the deep ConvNet from [Deep learning with convolutional neural networks for EEG decoding and visualization](https://arxiv.org/abs/1703.05051) (Section 2.4.2).
```
from braindecode.models.deep4 import Deep4Net
from torch import nn
from braindecode.torch_ext.util import set_random_seeds
from braindecode.models.util import to_dense_prediction_model
# Set if you want to use GPU
# You can also use torch.cuda.is_available() to determine if cuda is available on your machine.
cuda = True
set_random_seeds(seed=20170629, cuda=cuda)
# This will determine how many crops are processed in parallel
input_time_length = 450
# final_conv_length determines the size of the receptive field of the ConvNet
model = Deep4Net(in_chans=64, n_classes=2, input_time_length=input_time_length,
filter_length_3=5, filter_length_4=5,
pool_time_stride=2,
stride_before_pool=True,
final_conv_length=1)
if cuda:
model.cuda()
from braindecode.torch_ext.optimizers import AdamW
import torch.nn.functional as F
optimizer = AdamW(model.parameters(), lr=1*0.01, weight_decay=0.5*0.001) # these are good values for the deep model
model.compile(loss=F.nll_loss, optimizer=optimizer, iterator_seed=1, cropped=True)
```
## Run the training
```
input_time_length = 450
model.fit(train_set.X, train_set.y, epochs=30, batch_size=64, scheduler='cosine',
input_time_length=input_time_length,
validation_data=(valid_set.X, valid_set.y),)
```
## Compute correlation: amplitude perturbation - prediction change
First collect all batches and concatenate them into one array of examples:
```
from braindecode.datautil.iterators import CropsFromTrialsIterator
from braindecode.torch_ext.util import np_to_var
test_input = np_to_var(np.ones((2, 64, input_time_length, 1), dtype=np.float32))
if cuda:
test_input = test_input.cuda()
out = model.network(test_input)
n_preds_per_input = out.cpu().data.numpy().shape[2]
iterator = CropsFromTrialsIterator(batch_size=32,input_time_length=input_time_length,
n_preds_per_input=n_preds_per_input)
train_batches = list(iterator.get_batches(train_set, shuffle=False))
train_X_batches = np.concatenate(list(zip(*train_batches))[0])
```
Next, create a prediction function that wraps the model prediction function and returns the predictions as numpy arrays. We use the predition before the softmax, so we create a new module with all the layers of the old until before the softmax.
```
from torch import nn
from braindecode.torch_ext.util import var_to_np
import torch as th
new_model = nn.Sequential()
for name, module in model.network.named_children():
if name == 'softmax': break
new_model.add_module(name, module)
new_model.eval();
pred_fn = lambda x: var_to_np(th.mean(new_model(np_to_var(x).cuda())[:,:,:,0], dim=2, keepdim=False))
from braindecode.visualization.perturbation import compute_amplitude_prediction_correlations
amp_pred_corrs = compute_amplitude_prediction_correlations(pred_fn, train_X_batches, n_iterations=12,
batch_size=30)
```
## Plot correlations
Pick out one frequency range and mean correlations within that frequency range to make a scalp plot.
Here we use the alpha frequency range.
```
amp_pred_corrs.shape
fs = epoched.info['sfreq']
freqs = np.fft.rfftfreq(train_X_batches.shape[2], d=1.0/fs)
start_freq = 7
stop_freq = 14
i_start = np.searchsorted(freqs,start_freq)
i_stop = np.searchsorted(freqs, stop_freq) + 1
freq_corr = np.mean(amp_pred_corrs[:,i_start:i_stop], axis=1)
```
Now get approximate positions of the channels in the 10-20 system.
```
from braindecode.datasets.sensor_positions import get_channelpos, CHANNEL_10_20_APPROX
ch_names = [s.strip('.') for s in epoched.ch_names]
positions = [get_channelpos(name, CHANNEL_10_20_APPROX) for name in ch_names]
positions = np.array(positions)
```
### Plot with MNE
```
import matplotlib.pyplot as plt
from matplotlib import cm
%matplotlib inline
max_abs_val = np.max(np.abs(freq_corr))
fig, axes = plt.subplots(1, 2)
class_names = ['Left Hand', 'Right Hand']
for i_class in range(2):
ax = axes[i_class]
mne.viz.plot_topomap(freq_corr[:,i_class], positions,
vmin=-max_abs_val, vmax=max_abs_val, contours=0,
cmap=cm.coolwarm, axes=ax, show=False);
ax.set_title(class_names[i_class])
```
### Plot with Braindecode
```
from braindecode.visualization.plot import ax_scalp
fig, axes = plt.subplots(1, 2)
class_names = ['Left Hand', 'Right Hand']
for i_class in range(2):
ax = axes[i_class]
ax_scalp(freq_corr[:,i_class], ch_names, chan_pos_list=CHANNEL_10_20_APPROX, cmap=cm.coolwarm,
vmin=-max_abs_val, vmax=max_abs_val, ax=ax)
ax.set_title(class_names[i_class])
```
From these plots we can see the ConvNet clearly learned to use the lateralized response in the alpha band. Note that the positive correlations for the left hand on the left side do not imply an increase of alpha activity for the left hand in the data, see [Deep learning with convolutional neural networks for EEG decoding and visualization](https://arxiv.org/abs/1703.05051) Result 12 for some notes on interpretability.
## Dataset references
This dataset was created and contributed to PhysioNet by the developers of the [BCI2000](http://www.schalklab.org/research/bci2000) instrumentation system, which they used in making these recordings. The system is described in:
Schalk, G., McFarland, D.J., Hinterberger, T., Birbaumer, N., Wolpaw, J.R. (2004) BCI2000: A General-Purpose Brain-Computer Interface (BCI) System. IEEE TBME 51(6):1034-1043.
[PhysioBank](https://physionet.org/physiobank/) is a large and growing archive of well-characterized digital recordings of physiologic signals and related data for use by the biomedical research community and further described in:
Goldberger AL, Amaral LAN, Glass L, Hausdorff JM, Ivanov PCh, Mark RG, Mietus JE, Moody GB, Peng C-K, Stanley HE. (2000) PhysioBank, PhysioToolkit, and PhysioNet: Components of a New Research Resource for Complex Physiologic Signals. Circulation 101(23):e215-e220.
| github_jupyter |
<p style="margin-top: 3em; margin-bottom: 3em;"><font size="7"><b>Matplotlib & Seaborn: Introduction </b></font></p>
```
%matplotlib inline
```
# Matplotlib
[Matplotlib](http://matplotlib.org/) is a Python package used widely throughout the scientific Python community to produce high quality 2D publication graphics. It transparently supports a wide range of output formats including PNG (and other raster formats), PostScript/EPS, PDF and SVG and has interfaces for all of the major desktop GUI (graphical user interface) toolkits. It is a great package with lots of options.
However, matplotlib is...
> The 800-pound gorilla — and like most 800-pound gorillas, this one should probably be avoided unless you genuinely need its power, e.g., to make a **custom plot** or produce a **publication-ready** graphic.
> (As we’ll see, when it comes to statistical visualization, the preferred tack might be: “do as much as you easily can in your convenience layer of choice [nvdr e.g. directly from Pandas, or with seaborn], and then use matplotlib for the rest.”)
(quote used from [this](https://dansaber.wordpress.com/2016/10/02/a-dramatic-tour-through-pythons-data-visualization-landscape-including-ggplot-and-altair/) blogpost)
And that's we mostly did, just use the `.plot` function of Pandas. So, why do we learn matplotlib? Well, for the *...then use matplotlib for the rest.*; at some point, somehow!
Matplotlib comes with a convenience sub-package called ``pyplot`` which, for consistency with the wider matplotlib community, should always be imported as ``plt``:
```
import numpy as np
import matplotlib.pyplot as plt
```
## - dry stuff - The matplotlib `Figure`, `axes` and `axis`
At the heart of **every** plot is the figure object. The "Figure" object is the top level concept which can be drawn to one of the many output formats, or simply just to screen. Any object which can be drawn in this way is known as an "Artist" in matplotlib.
Lets create our first artist using pyplot, and then show it:
```
fig = plt.figure()
plt.show()
```
On its own, drawing the figure artist is uninteresting and will result in an empty piece of paper (that's why we didn't see anything above).
By far the most useful artist in matplotlib is the "Ax**e**s" artist. The Axes artist represents the "data space" of a typical plot, a rectangular axes (the most common, but not always the case, e.g. polar plots) will have 2 (confusingly named) Ax**i**s artists with tick labels and tick marks.
There is no limit on the number of Axes artists which can exist on a Figure artist. Let's go ahead and create a figure with a single Axes artist, and show it using pyplot:
```
ax = plt.axes()
```
Matplotlib's ``pyplot`` module makes the process of creating graphics easier by allowing us to skip some of the tedious Artist construction. For example, we did not need to manually create the Figure artist with ``plt.figure`` because it was implicit that we needed a figure when we created the Axes artist.
Under the hood matplotlib still had to create a Figure artist, its just we didn't need to capture it into a variable. We can access the created object with the "state" functions found in pyplot called **``gcf``** and **``gca``**.
## - essential stuff - `pyplot` versus Object based
Some example data:
```
x = np.linspace(0, 5, 10)
y = x ** 2
```
Observe the following difference:
**1. pyplot style: plt...** (you will see this a lot for code online!)
```
plt.plot(x, y, '-')
```
**2. creating objects**
```
fig, ax = plt.subplots()
ax.plot(x, y, '-')
```
Although a little bit more code is involved, the advantage is that we now have **full control** of where the plot axes are placed, and we can easily add more than one axis to the figure:
```
fig, ax1 = plt.subplots()
ax.plot(x, y, '-')
ax2 = fig.add_axes([0.2, 0.5, 0.4, 0.3]) # inset axes
ax1.plot(x, y, '-')
ax1.set_ylabel('y')
ax2.set_xlabel('x')
ax2.plot(x, y*2, 'r-')
```
<div class="alert alert-info" style="font-size:18px">
<b>REMEMBER</b>:
<ul>
<li>Use the **object oriented** power of Matplotlib!</li>
<li>Get yourself used to writing `fig, ax = plt.subplots()`</li>
</ul>
</div>
```
fig, ax = plt.subplots()
ax.plot(x, y, '-')
# ...
```
## An small cheat-sheet reference for some common elements
```
x = np.linspace(-1, 0, 100)
fig, ax = plt.subplots()
# Adjust the created axes so that its topmost extent is 0.8 of the figure.
fig.subplots_adjust(top=0.8)
ax.plot(x, x**2, color='0.4', label="power 2")
ax.plot(x, x**3, color='0.8', linestyle='--', label="power 3")
fig.suptitle('Figure title', fontsize=18,
fontweight='bold')
ax.set_title('Axes title', fontsize=16)
ax.set_xlabel('The X axis')
ax.set_ylabel('The Y axis $y=f(x)$', fontsize=16)
ax.set_xlim(-1.0, 1.1)
ax.set_ylim(-0.1, 1.)
ax.text(0.5, 0.2, 'Text centered at (0.5, 0.2)\nin data coordinates.',
horizontalalignment='center', fontsize=14)
ax.text(0.5, 0.5, 'Text centered at (0.5, 0.5)\nin Figure coordinates.',
horizontalalignment='center', fontsize=14,
transform=ax.transAxes, color='grey')
ax.legend(loc='upper right', frameon=True, ncol=2)
```
For more information on legend positioning, check [this post](http://stackoverflow.com/questions/4700614/how-to-put-the-legend-out-of-the-plot) on stackoverflow!
Another nice blogpost about customizing matplotlib figures: http://pbpython.com/effective-matplotlib.html
## I do not like the style...
The power of the object-oriented way of working makes it possible to change everything. However, mostly we just want **quickly a good-looking plot**. Matplotlib provides a number of styles that can be used to quickly change a number of settings:
```
plt.style.available
x = np.linspace(0, 10)
with plt.style.context('seaborn-muted'): # 'ggplot', 'bmh', 'grayscale', 'seaborn-whitegrid'
fig, ax = plt.subplots()
ax.plot(x, np.sin(x) + x + np.random.randn(50))
ax.plot(x, np.sin(x) + 0.5 * x + np.random.randn(50))
ax.plot(x, np.sin(x) + 2 * x + np.random.randn(50))
```
We should not start discussing about colors and styles, just pick **your favorite style**!
## Interaction with Pandas
What we have been doing while plotting with Pandas:
```
import pandas as pd
aqdata = pd.read_csv('data/20000101_20161231-NO2.csv', sep=';', skiprows=[1], na_values=['n/d'],
index_col=0, parse_dates=True)
aqdata = aqdata["2014":].resample('D').mean()
aqdata.plot()
```
### The pandas versus matplotlib
#### Comparison 1: single plot
```
aqdata.plot(figsize=(16, 6)) # shift tab this!
```
Making this with matplotlib...
```
fig, ax = plt.subplots(figsize=(16, 6))
ax.plot(aqdata.index, aqdata["BASCH"],
aqdata.index, aqdata["BONAP"],
aqdata.index, aqdata["PA18"],
aqdata.index, aqdata["VERS"])
ax.legend(["BASCH", "BONAP", "PA18", "VERS"])
```
or...
```
fig, ax = plt.subplots(figsize=(16, 6))
for station in aqdata.columns:
ax.plot(aqdata.index, aqdata[station], label=station)
ax.legend()
```
#### Comparison 2: with subplots
```
axs = aqdata.plot(subplots=True, sharex=True,
figsize=(16, 8), colormap='viridis', # Dark2
fontsize=15)
```
Mimicking this in matplotlib (just as a reference):
```
from matplotlib import cm
import matplotlib.dates as mdates
colors = [cm.viridis(x) for x in np.linspace(0.0, 1.0, len(aqdata.columns))] # list comprehension to set up the colors
fig, axs = plt.subplots(4, 1, figsize=(16, 8))
for ax, col, station in zip(axs, colors, aqdata.columns):
ax.plot(aqdata.index, aqdata[station], label=station, color=col)
ax.legend()
if not ax.is_last_row():
ax.xaxis.set_ticklabels([])
ax.xaxis.set_major_locator(mdates.YearLocator())
else:
ax.xaxis.set_major_locator(mdates.YearLocator())
ax.xaxis.set_major_formatter(mdates.DateFormatter('%Y'))
ax.set_xlabel('Time')
ax.tick_params(labelsize=15)
fig.autofmt_xdate()
```
### Best of both worlds...
```
aqdata.columns
fig, ax = plt.subplots() #prepare a matplotlib figure
aqdata.plot(ax=ax) # use pandas for the plotting
# Provide further adaptations with matplotlib:
ax.set_xlabel("")
ax.tick_params(labelsize=15, pad=8, which='both')
fig.suptitle('Air quality station time series', fontsize=15)
fig, (ax1, ax2) = plt.subplots(2, 1) #provide with matplotlib 2 axis
aqdata[["BASCH", "BONAP"]].plot(ax=ax1) # plot the two timeseries of the same location on the first plot
aqdata["PA18"].plot(ax=ax2) # plot the other station on the second plot
# further adapt with matplotlib
ax1.set_ylabel("BASCH")
ax2.set_ylabel("PA18")
ax2.legend()
```
<div class="alert alert-info">
<b>Remember</b>:
<ul>
<li>You can do anything with matplotlib, but at a cost... [stackoverflow!!](http://stackoverflow.com/questions/tagged/matplotlib)</li>
<li>The preformatting of Pandas provides mostly enough flexibility for quick analysis and draft reporting. It is not for paper-proof figures or customization</li>
</ul>
<br>
</div>
<div class="alert alert-danger">
<b>NOTE</b>:
If you take the time to make you're perfect/spot-on/greatest-ever matplotlib-figure: Make it a **reusable function**! (see tomorrow!)
<ul>
<li>Let your hard work pay off, write your own custom functions!</li>
</ul>
</div>
<div class="alert alert-info" style="font-size:18px">
<b>Remember</b>:
`fig.savefig()` to save your Figure object!
</div>
# Seaborn
```
import seaborn as sns
```
* Built on top of Matplotlib, but providing
1. High level functions
2. Much cleaner default figures
* Works well with Pandas
## First example: `pairplot`
A scatterplot comparing the three stations with a color variation on the months:
```
aqdata["month"] = aqdata.index.month
sns.pairplot(aqdata["2014"].dropna(),
vars=['BASCH', 'BONAP', 'PA18', 'VERS'],
diag_kind='kde', hue="month")
```
## Seaborn works well with Pandas & is built on top of Matplotlib
We will use the Titanic example again:
```
titanic = pd.read_csv('data/titanic.csv')
titanic.head()
```
**Histogram**: Getting the univariaite distribution of the `Age`
```
fig, ax = plt.subplots()
sns.distplot(titanic["Age"].dropna(), ax=ax) # Seaborn does not like Nan values...
sns.rugplot(titanic["Age"].dropna(), color="g", ax=ax) # rugplot provides lines at the individual data point locations
ax.set_ylabel("Frequency")
```
<div class="alert alert-info">
<b>Remember</b>:
Similar to Pandas handling above, we can set up a `figure` and `axes` and add the seaborn output to it; adapt it afterwards
</div>
Compare two variables (**scatter-plot**):
```
g = sns.jointplot(x="Fare", y="Age",
data=titanic,
kind="scatter") #kde, hex
g = sns.jointplot(x="Fare", y="Age",
data=titanic,
kind="scatter") #kde, hex
# Adapt the properties with matplotlib by changing the available axes objects
g.ax_marg_x.set_ylabel("Frequency")
g.ax_joint.set_facecolor('0.1')
g.ax_marg_y.set_xlabel("Frequency")
```
<div class="alert alert-info">
<b>Remember</b>:
Adapting the output of a Seaborn `grid` of different axes can be done as well to adapt it with matplotlib
</div>
Who likes **regressions**?
```
fig, ax = plt.subplots()
sns.regplot(x="Fare", y="Age", data=titanic, ax=ax, lowess=False)
# adding the small lines to indicate individual data points
sns.rugplot(titanic["Fare"].dropna(), axis='x',
color="#6699cc", height=0.02, ax=ax)
sns.rugplot(titanic["Age"].dropna(), axis='y',
color="#6699cc", height=0.02, ax=ax)
```
## Section especially for R `ggplot` lovers
### Regressions with factors/categories: `lmplot`
When you want to take into account a category as well to do regressions, use `lmplot` (which is a special case of `Facetgrid`):
```
sns.lmplot(x="Fare", y="Age", hue="Sex",
data=titanic)
sns.lmplot(x="Fare", y="Age", hue="Sex",
col="Survived", data=titanic)
```
### Other plots with factors/categories: `factorplot`
Another method to create thes **category** based split of columns, colors,... based on specific category columns is the `factorplot`
```
titanic.head()
sns.factorplot(x="Sex",
y="Fare",
col="Pclass",
data=titanic) #kind='strip' # violin,...
sns.factorplot(x="Sex", y="Fare", col="Pclass", row="Embarked",
data=titanic, kind='bar')
g = sns.factorplot(x="Survived", y="Fare", hue="Sex",
col="Embarked", data=titanic,
kind="box", size=4, aspect=.5);
g.fig.set_figwidth(15)
g.fig.set_figheight(6)
```
<div class="alert alert-info">
<b>Remember</b>:
<ul>
<li>`lmplot` and `factorplot` are shortcuts for a more advanced `FacetGrid` functionality</li>
<li>If you want to dig deeper into this `FacetGrid`-based plotting, check the [online manual](http://seaborn.pydata.org/tutorial/axis_grids.html)!</li>
</ul>
</div>
# Need more matplotlib/seaborn inspiration?
For more in-depth material:
* http://www.labri.fr/perso/nrougier/teaching/matplotlib/
* notebooks in matplotlib section: http://nbviewer.jupyter.org/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/Index.ipynb#4.-Visualization-with-Matplotlib
* main reference: [matplotlib homepage](http://matplotlib.org/)
* very nice blogpost about customizing figures with matplotlib: http://pbpython.com/effective-matplotlib.html
<div class="alert alert-info" style="font-size:18px">
<b>Remember</b>(!)
<ul>
<li>[matplotlib Gallery](http://matplotlib.org/gallery.html)</li>
<li>[seaborn gallery ](http://seaborn.pydata.org/examples/index.html)</li>
</ul>
<br>
Important resources to start from!
</div>
# Alternatives for matplotlib
We only use matplotlib (or matplotlib-based plotting) in this workshop, and it is still the main plotting library for many scientists, but it is not the only existing plotting library.
A nice overview of the landscape of visualisation tools in python was recently given by Jake VanderPlas: (or matplotlib-based plotting): https://speakerdeck.com/jakevdp/pythons-visualization-landscape-pycon-2017
Bokeh (http://bokeh.pydata.org/en/latest/): interactive, web-based visualisation
```
from bokeh.io import output_notebook
output_notebook()
from bokeh.plotting import figure, show
from bokeh.sampledata.iris import flowers
colormap = {'setosa': 'red', 'versicolor': 'green', 'virginica': 'blue'}
colors = [colormap[x] for x in flowers['species']]
p = figure(title = "Iris Morphology")
p.xaxis.axis_label = 'Petal Length'
p.yaxis.axis_label = 'Petal Width'
p.circle(flowers["petal_length"], flowers["petal_width"],
color=colors, fill_alpha=0.2, size=10)
show(p)
```
Altair (https://altair-viz.github.io/index.html): declarative statistical visualization library for Python, based on Vega.
```
from altair import Chart, load_dataset
# load built-in dataset as a pandas DataFrame
iris = load_dataset('iris')
Chart(iris).mark_circle().encode(
x='petalLength',
y='petalWidth',
color='species',
)
```
---
# Acknowledgement
> This notebook is partly based on material of © 2016, Joris Van den Bossche and Stijn Van Hoey (<mailto:jorisvandenbossche@gmail.com>, <mailto:stijnvanhoey@gmail.com>, licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/) and partly on material of the Met Office (Copyright (C) 2013 SciTools, GPL licensed): https://github.com/SciTools/courses
| github_jupyter |
# Nipype Quickstart
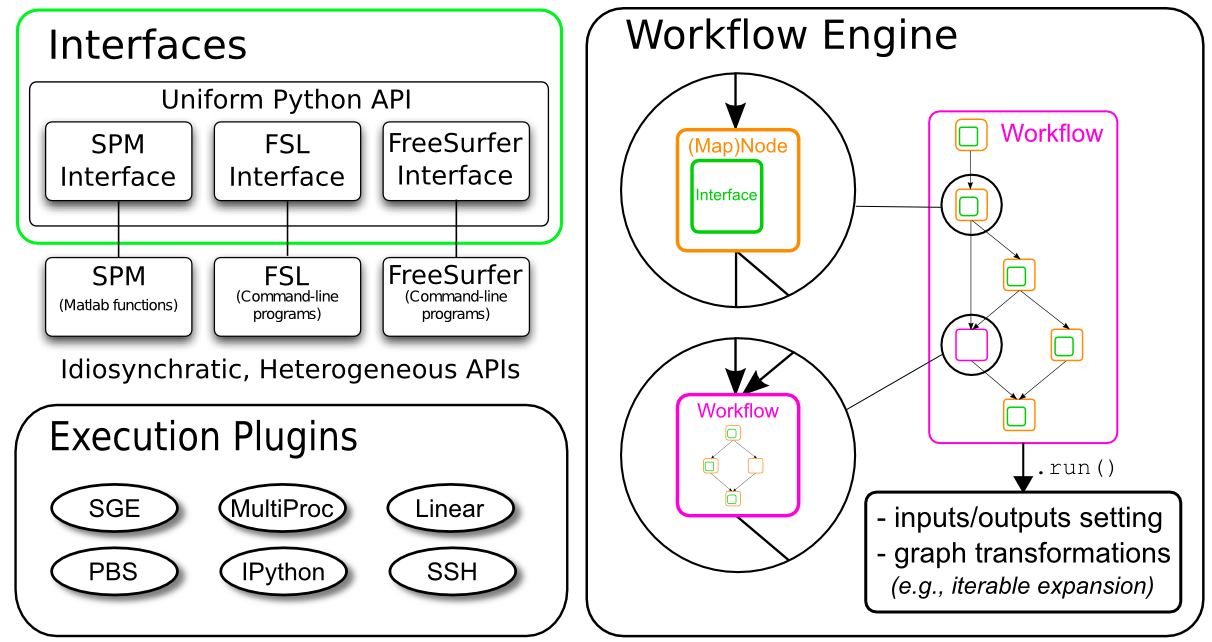
- [Existing documentation](http://nipype.readthedocs.io/en/latest/)
- [Visualizing the evolution of Nipype](https://www.youtube.com/watch?v=cofpD1lhmKU)
- This notebook is taken from [reproducible-imaging repository](https://github.com/ReproNim/reproducible-imaging)
#### Import a few things from nipype and external libraries
```
import os
from os.path import abspath
from nipype import Workflow, Node, MapNode, Function
from nipype.interfaces.fsl import BET, IsotropicSmooth, ApplyMask
from nilearn.plotting import plot_anat
%matplotlib inline
import matplotlib.pyplot as plt
```
## Interfaces
Interfaces are the core pieces of Nipype. The interfaces are python modules that allow you to use various external packages (e.g. FSL, SPM or FreeSurfer), even if they themselves are written in another programming language than python.
**Let's try to use `bet` from FSL:**
```
# will use a T1w from ds000114 dataset
input_file = abspath("/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz")
# we will be typing here
```
If you're lost the code is here:
```
bet = BET()
bet.inputs.in_file = input_file
bet.inputs.out_file = "/output/T1w_nipype_bet.nii.gz"
res = bet.run()
```
let's check the output:
```
res.outputs
```
and we can plot the output file
```
plot_anat('/output/T1w_nipype_bet.nii.gz',
display_mode='ortho', dim=-1, draw_cross=False, annotate=False);
```
you can always check the list of arguments using `help` method
```
BET.help()
```
#### Exercise 1a
Import `IsotropicSmooth` from `nipype.interfaces.fsl` and find out the `FSL` command that is being run. What are the mandatory inputs for this interface?
```
# type your code here
from nipype.interfaces.fsl import IsotropicSmooth
# all this information can be found when we run `help` method.
# note that you can either provide `in_file` and `fwhm` or `in_file` and `sigma`
IsotropicSmooth.help()
```
#### Exercise 1b
Run the `IsotropicSmooth` for `/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz` file with a smoothing kernel 4mm:
```
# type your solution here
smoothing = IsotropicSmooth()
smoothing.inputs.in_file = "/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz"
smoothing.inputs.fwhm = 4
smoothing.inputs.out_file = "/output/T1w_nipype_smooth.nii.gz"
smoothing.run()
# plotting the output
plot_anat('/output/T1w_nipype_smooth.nii.gz',
display_mode='ortho', dim=-1, draw_cross=False, annotate=False);
```
## Nodes and Workflows
Interfaces are the core pieces of Nipype that run the code of your desire. But to streamline your analysis and to execute multiple interfaces in a sensible order, you have to put them in something that we call a Node and create a Workflow.
In Nipype, a node is an object that executes a certain function. This function can be anything from a Nipype interface to a user-specified function or an external script. Each node consists of a name, an interface, and at least one input field and at least one output field.
Once you have multiple nodes you can use `Workflow` to connect with each other and create a directed graph. Nipype workflow will take care of input and output of each interface and arrange the execution of each interface in the most efficient way.
**Let's create the first node using `BET` interface:**
```
# we will be typing here
```
If you're lost the code is here:
```
# Create Node
bet_node = Node(BET(), name='bet')
# Specify node inputs
bet_node.inputs.in_file = input_file
bet_node.inputs.mask = True
# bet node can be also defined this way:
#bet_node = Node(BET(in_file=input_file, mask=True), name='bet_node')
```
#### Exercise 2
Create a `Node` for IsotropicSmooth interface.
```
# Type your solution here:
# smooth_node =
smooth_node = Node(IsotropicSmooth(in_file=input_file, fwhm=4), name="smooth")
```
**We will now create one more Node for our workflow**
```
mask_node = Node(ApplyMask(), name="mask")
```
Let's check the interface:
```
ApplyMask.help()
```
As you can see the interface takes two mandatory inputs: `in_file` and `mask_file`. We want to use the output of `smooth_node` as `in_file` and one of the output of `bet_file` (the `mask_file`) as `mask_file` input.
** Let's initialize a `Workflow`:**
```
# will be writing the code here:
```
if you're lost, the full code is here:
```
# Initiation of a workflow
wf = Workflow(name="smoothflow", base_dir="/output/working_dir")
```
It's very important to specify `base_dir` (as absolute path), because otherwise all the outputs would be saved somewhere in the temporary files.
**let's connect the `bet_node` output to `mask_node` input`**
```
# we will be typing here:
```
if you're lost, the code is here:
```
wf.connect(bet_node, "mask_file", mask_node, "mask_file")
```
#### Exercise 3
Connect `out_file` of `smooth_node` to `in_file` of `mask_node`.
```
# type your code here
wf.connect(smooth_node, "out_file", mask_node, "in_file")
```
**Let's see a graph describing our workflow:**
```
wf.write_graph("workflow_graph.dot")
from IPython.display import Image
Image(filename="/output/working_dir/smoothflow/workflow_graph.png")
```
you can also plot a more detailed graph:
```
wf.write_graph(graph2use='flat')
from IPython.display import Image
Image(filename="/output/working_dir/smoothflow/graph_detailed.png")
```
**and now let's run the workflow**
```
# we will type our code here:
```
if you're lost, the full code is here:
```
# Execute the workflow
res = wf.run()
```
**and let's look at the results**
```
# we can check the output of specific nodes from workflow
list(res.nodes)[0].result.outputs
```
**we can see the fie structure that has been created:**
```
! tree -L 3 /output/working_dir/smoothflow/
```
**and we can plot the results:**
```
import numpy as np
import nibabel as nb
#import matplotlib.pyplot as plt
# Let's create a short helper function to plot 3D NIfTI images
def plot_slice(fname):
# Load the image
img = nb.load(fname)
data = img.get_fdata()
# Cut in the middle of the brain
cut = int(data.shape[-1]/2) + 10
# Plot the data
plt.imshow(np.rot90(data[..., cut]), cmap="gray")
plt.gca().set_axis_off()
f = plt.figure(figsize=(12, 4))
for i, img in enumerate(["/data/ds000114/sub-01/ses-test/anat/sub-01_ses-test_T1w.nii.gz",
"/output/working_dir/smoothflow/smooth/sub-01_ses-test_T1w_smooth.nii.gz",
"/output/working_dir/smoothflow/bet/sub-01_ses-test_T1w_brain_mask.nii.gz",
"/output/working_dir/smoothflow/mask/sub-01_ses-test_T1w_smooth_masked.nii.gz"]):
f.add_subplot(1, 4, i + 1)
plot_slice(img)
```
## Iterables
Some steps in a neuroimaging analysis are repetitive. Running the same preprocessing on multiple subjects or doing statistical inference on multiple files. To prevent the creation of multiple individual scripts, Nipype has as execution plugin for ``Workflow``, called **``iterables``**.
<img src="../static/images/iterables.png" width="240">
Let's assume we have a workflow with two nodes, node (A) does simple skull stripping, and is followed by a node (B) that does isometric smoothing. Now, let's say, that we are curious about the effect of different smoothing kernels. Therefore, we want to run the smoothing node with FWHM set to 2mm, 8mm, and 16mm.
**let's just modify `smooth_node`:**
```
# we will type the code here
```
if you're lost the code is here:
```
smooth_node_it = Node(IsotropicSmooth(in_file=input_file), name="smooth")
smooth_node_it.iterables = ("fwhm", [4, 8, 16])
```
we will define again bet and smooth nodes:
```
bet_node_it = Node(BET(in_file=input_file, mask=True), name='bet_node')
mask_node_it = Node(ApplyMask(), name="mask")
```
** will create a new workflow with a new `base_dir`:**
```
# Initiation of a workflow
wf_it = Workflow(name="smoothflow_it", base_dir="/output/working_dir")
wf_it.connect(bet_node_it, "mask_file", mask_node_it, "mask_file")
wf_it.connect(smooth_node_it, "out_file", mask_node_it, "in_file")
```
**let's run the workflow and check the output**
```
res_it = wf_it.run()
```
**let's see the graph**
```
list(res_it.nodes)
```
We can see the file structure that was created:
```
! tree -L 3 /output/working_dir/smoothflow_it/
```
you have now 7 nodes instead of 3!
### MapNode
If you want to iterate over a list of inputs, but need to feed all iterated outputs afterward as one input (an array) to the next node, you need to use a **``MapNode``**. A ``MapNode`` is quite similar to a normal ``Node``, but it can take a list of inputs and operate over each input separately, ultimately returning a list of outputs.
Imagine that you have a list of items (let's say files) and you want to execute the same node on them (for example some smoothing or masking). Some nodes accept multiple files and do exactly the same thing on them, but some don't (they expect only one file). `MapNode` can solve this problem. Imagine you have the following workflow:
<img src="../static/images/mapnode.png" width="325">
Node `A` outputs a list of files, but node `B` accepts only one file. Additionally, `C` expects a list of files. What you would like is to run `B` for every file in the output of `A` and collect the results as a list and feed it to `C`.
** Let's run a simple numerical example using nipype `Function` interface **
```
def square_func(x):
return x ** 2
square = Function(input_names=["x"], output_names=["f_x"], function=square_func)
```
If I want to know the results only for one `x` we can use `Node`:
```
square_node = Node(square, name="square")
square_node.inputs.x = 2
res = square_node.run()
res.outputs
```
let's try to ask for more values of `x`
```
# NBVAL_SKIP
square_node = Node(square, name="square")
square_node.inputs.x = [2, 4]
res = square_node.run()
res.outputs
```
**It will give an error since `square_func` do not accept list. But we can try `MapNode`:**
```
square_mapnode = MapNode(square, name="square", iterfield=["x"])
square_mapnode.inputs.x = [2, 4]
res = square_mapnode.run()
res.outputs
```
**Notice that `f_x` is a list again!**
| github_jupyter |
# Estrazione dati
## [Scarica zip esercizi](../_static/generated/extraction.zip)
[Naviga file online](https://github.com/DavidLeoni/softpython-it/tree/master/extraction)
## Introduzione
In questo tutorial affronteremo il tema dell'estrazione di dati semi-strutturati, focalizzandoci in particolare sull'HTML. I file in questo formato seguono (o dovrebbero seguire !) le regole dell'XML, quindi guardando dei file HTML possiamo anche imparare qualcosa sul più generico XML.
Scaletta:
0. Impariamo due cose di HTML creando una paginetta web seguendo un semplice tutorial CoderDojoTrento (senza Python).
1. Chiedersi se vale la pena estrarre informazioni dall'HTML
2. Estrazione eventi del Trentino da [visittrentino.info](https://www.visittrentino.info/it/guida/eventi) usando Python e BeautifulSoup 4
3. Per ogni evento, estrarremo nome, data, luogo, tipo, descrizione e li metteremo in una lista di dizionari Python così:
```python
[{'data': '14/12/2017',
'descrizione': 'Al Passo Costalunga sfida tra i migliori specialisti del mondo',
'luogo': 'Passo Costalunga',
'nome': 'Coppa del Mondo di Snowboard',
'tipo': 'Sport, TOP EVENTI SPORT'},
{'data': '18/12/2017',
'descrizione': 'Lunedì 18 dicembre la Coppa Europa fa tappa in Val di Fassa',
'luogo': 'Pozza di Fassa',
'nome': 'Coppa Europa di sci alpino maschile - slalom speciale',
'tipo': 'Sport, TOP EVENTI SPORT'},
....
]
```
4. Infine scriveremo un file CSV `eventi.csv` usando la lista di dizionari generata al punto precedente.
### Che fare
1. Per avere un'idea di cosa sia l'HTML, prova a crearti una paginetta web seguendo il [tutorial 1](http://coderdojotrento.it/web1) di CoderDojoTrento (in questa parte non serve Python, puoi fare tutto online saltando la registrazione su Thimble)
2. Adesso puoi passare ad usare Python e Jupyter come al solito: scarica lo [zip con esercizi e soluzioni](../_static/generated/extraction.zip)
- scompatta lo zip in una cartella, dovresti ottenere qualcosa del genere:
```
extraction
extraction.ipynb
extraction-sol.ipynb
jupman.py
```
<div class="alert alert-warning">
**ATTENZIONE**: Per essere visualizzato correttamente, il file del notebook DEVE essere nella cartella szippata.
</div>
- apri il Jupyter Notebook da quella cartella. Due cose dovrebbero aprirsi, prima una console e poi un browser. Il browser dovrebbe mostrare una lista di file: naviga la lista e apri il notebook `extraction.ipynb`
- Prosegui leggendo il file degli esercizi, ogni tanto al suo interno troverai delle scritte **ESERCIZIO**, che ti chiederanno di scrivere dei comandi Python nelle celle successive.
Scorciatoie da tastiera:
* Per eseguire il codice Python dentro una cella di Jupyter, premi `Control+Invio`
* Per eseguire il codice Python dentro una cella di Jupyter E selezionare la cella seguente, premi `Shift+Invio`
* Per eseguire il codice Python dentro una cella di Jupyter E creare una nuova cella subito dopo, premi `Alt+Invio`
* Se per caso il Notebook sembra inchiodato, prova a selezionare `Kernel -> Restart`
## 1. Guardiamo l'html
Indipendentemente dalla domanda posta al punto precedente, per fini didattici proseguiremo con lo _scraping_ html.
All'interno dello zip degli esercizi, c'è un file chiamato [eventi.html](eventi.html). Dopo aver scompattato lo zip, apri il file nel tuo browser.
<div class="alert alert-warning">
**IMPORTANTE: usa solo il file eventi.html nella cartella del progetto Jupyter!**
Il file [eventi.html](eventi.html) è stato salvato nel 2017 ed è un po' più brutto da vedere della versione online sul sito di visttrentino (mancano immagini, etc). Se sei curioso di provare [quella online](https://www.visittrentino.info/it/guida/eventi), per vedere gli eventi come lista ricordati di cliccare sulla relativa icona:

</div>
**✪ 1.1 ESERCIZIO**: Dopo aver aperto il file nel browser, visualizza l'HTML all'interno (premendo per es `Ctrl+U` in Firefox/Chrome/Safari). Familiarizzati un po' con il file sorgente, cercando all'interno alcuni valori del primo evento , come per esempio il nome `Coppa del Mondo di Snowboard`, la data `14/12/2017`, il luogo `Passo Costalunga`, il tipo `Sport, TOP EVENTI SPORT`, e la descrizione `Al Passo Costalunga sfida tra i migliori specialisti del mondo`.
**NOTA 1**: Per questo esercizio e tutti i seguenti, usa solo il file [eventi.html](eventi.html) indicato, NON usare l'html dalla [versione live](https://www.visittrentino.info/it/guida/eventi) proveniente dalla di visittrentino, che può essere soggetta a cambiamenti !
**NOTA 2**: Evita di usare il browser Internet Explorer, in alternativa cerca di usare uno tra i seguenti (in quest'ordine): Firefox, Chrome, Safari.
Per maggiori informazioni sul formato XML, di cui l'HTML è una incarnazione, puoi consultare il libro Immersione in Python [al capitolo 12](http://gpiancastelli.altervista.org/dip3-it/xml.html),
## 2. Estrazione con BeautifulSoup
Installiamo le librerie `beautifulsoup4` e il parser `lxml`:
- Anaconda:
* `conda install beautifulsoup4`
* `conda install lxml`
- Linux/Mac (`--user` installa nella propria home):
* `python3 -m pip install --user beautifulsoup4`
* `python3 -m pip install --user lxml`
**I parser**
Quanto installato sopra è quanto basta per questo esercizio. In particolare abbiamo installato il parser `lxml`, che è il componente che permette a BeautifulSoup di leggere l'HTML in modo veloce e _lenient_, cioè tollerando possibili errori di formattazione che potrebbero essere presenti nell'HTML (NOTA: la maggior parte dei documenti html nel mondo reale ha problemi di formattazione !). Prendendo pagine a caso da internet, `lxml` potrebbe non essere adatto. Se hai problemi a leggere pagine html, potresti provare a sostituire `lxml` con `html5lib`, o altri parser. Per una lista di possibili parser, vedere la [documentazione di BeautifulSoup](https://www.crummy.com/software/BeautifulSoup/bs4/doc/#installing-a-parser )
### 2.1 Estraiamo i nomi
**✪ ESERCIZIO 2.1.1** Cerca nell'HTML la stringa `Coppa del Mondo di Snowboard`, cercando di capire bene in quali blocchi appare.
**RISPOSTA**:
Compare in 4 blocchi: h2, script Javascript, script semantici `ld+json` e blocchi `h4`, li ricapitoliamo qua sotto:
**RISPOSTA 2.1.1 - blocchi h2** : Compare in blocchi `h2` come `<h2 class="text-secondary">Coppa del Mondo di Snowboard</h2>` qua:
```html
<div class="col col-sm-9 arrange__item">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" class="text-primary list-teaser__link"><span class="icon icon-circle-arrow fz30"></span></a>
<div class="teaser__body">
<span class="text-secondary fz14 text-uppercase strong">Sport, TOP EVENTI SPORT</span>
<h2 class="text-secondary">Coppa del Mondo di Snowboard</h2>
<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>Passo Costalunga</a>
</li>
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>14/12/2017</a>
</li>
</ul>
<p>Al Passo Costalunga sfida tra i migliori specialisti del mondo</p>
</div>
</div>
```
**RISPOSTA 2.1.1 - script Javascript**: Compare in blocchi script Javascript come qua alla riga ` class=\"fz24 text-blue\">Coppa del Mondo di Snowboard<\/` :
```html
<script>
var mapData = [{"id":165375,"lat":46.4021014,"lng":11.6110718,"poiIcon":"\/static\/img\/content\/pois\/poi.png","poiIconActive":"\/static\/img\/content\/pois\/poi-active.png","infoBox":"<div class=\"row\"><div class=\"col-xs-5\"><img src=\"\/website\/var\/tmp\/image-thumbnails\/0\/9500\/thumb__moodboardmap\/snowparks-and-snowboard-brenta-dolomites_1.jpeg\"><\/div><div class=\"col-xs-7\"><h2 class=\"fz24 text-blue\">Coppa del Mondo di Snowboard<\/h2><p class=\"fz14\">14\/12 A Passo Costalunga sfida tra i migliori specialisti del mondo<\/p><a href=\"\/it\/guida\/eventi\/coppa-del-mondo-di-snowboard_e_165375\" class=\"btn btn-primary\">Maggiori info<\/a><\/div><\/div>"},{"id":309611,"lat":46.403707118232,"lng":11.60915851593,"poiIcon":"\/static\/img\/content\/pois\/poi.png","poiIconActive":"\/static\/img\/content\/pois\/poi-active.png","infoBox":"<div class=\"row\"><div class=\"col-xs-5\"><img src=\"\/website\/var\/tmp\/image-thumbnails\/110000\/119958\/thumb__moodboardmap\/foto-3_2.jpeg\"><\/div><div class=\"col-xs-7\"><h2 class=\"fz24 text-blue\">Coppa Europa di sci alpino maschile - slalom speciale<\/h2><p class=\"fz14\">18 dicembre in Val di Fassa<\/p><a href=\"\/it\/guida\/eventi\/coppa-europa-di-sci-alpino-maschile-slalom-speciale_e_309611\" class=\"btn btn-primary\">Maggiori info<\/a><\/div><\/div>"},
....
....
</script>
```
**RISPOSTA 2.1.1 - blocchi script ld+json**: Compare in altri blocchi script semantici jsonld nella parte `"name":"Coppa del Mondo di Snowboard"` come qua. Questi dati sono intesi per le macchine (come il motore di ricerca Google quando indicizza il contenuto delle pagine), quindi sarebbero particolarmente appetibili per i nostri scopi, ma per questo esercizio li ignoreremo:
```
<script type="application/ld+json">
{"@context":"http://schema.org","@type":"Event","name":"Coppa del Mondo di Snowboard","description":"Al Passo Costalunga sfida tra i migliori specialisti del mondo","startDate":"2017-12-14T00:00:00+01:00","endDate":"2017-12-14T00:00:00+01:00","location":{"@type":"Place","name":"Passo Costalunga","address":{"@type":"PostalAddress","addressCountry":"Italy","postalCode":"39056","streetAddress":""}},"image":{"@type":"ImageObject","url":"https://www.visittrentino.info/website/var/tmp/image-thumbnails/0/9500/thumb__contentgallery/snowparks-and-snowboard-brenta-dolomites_1.jpeg","height":396,"width":791},"performer":"Trentino Marketing S.r.l.","url":"https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375","offers":{"@type":"AggregateOffer","lowPrice":"\u20ac0","offerCount":"000","url":"https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375"}}
</script>
```
**Cerchiamo con Python**
Se hai fatto l'esercizio precedente, dovresti aver trovato vari blocchi che contengono `Coppa del Mondo di Snowboard`. Per un elenco e discussione, [guarda la soluzione](extraction-sol.ipynb#2.1-Estraiamo-i-nomi) Per questo esercizio, considereremo principalmente i blocchi `h4`, in cui il nome `Coppa del Mondo di Snowboard` compare in questa riga:
```html
<h4 class="moodboard__item-headline">Coppa del Mondo di Snowboard</h4>
```
vediamo cosa c'è intorno:
```html
<div class="moodboard__item-text text-white">
<div>
<h4 class="moodboard__item-headline">Coppa del Mondo di Snowboard</h4>
<span class="moodboard__item-subline strong fz14 text-uppercase d-b">14/12/2017</span>
<span class="moodboard__item-subline strong fz14 d-b"><span class="icon icon-map-view fz20"></span> Passo Costalunga</span>
</div>
<div class="moodboard__item-text__link text-right">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" role="link"><span class="icon icon-circle-arrow fz30"></span></a>
</div>
</div>
```
Torniamo alla riga:
```html
<h4 class="moodboard__item-headline">Coppa del Mondo di Snowboard</h4>
```
Notiamo che:
* inizia con il tag `<h4>`
* finisce simmetricamente con il tag di chiusura `</h4>`
* il tag di apertura ha un parametro `class="moodboard__item-headline"`, ma al momento non ci interessa
* Il testo che cerchiamo `Coppa del Mondo di Snowboard` è incluso tra i due tag
Per estrarre solo i nomi degli eventi dal documento, possiamo quindi cercare i tag `h4`. Eseguiamo la nostra prima estrazione in Python:
```
# Importa l'oggetto BeatifulSoup dal modulo bs4:
from bs4 import BeautifulSoup
# apriamo il file degli eventi:
# - specifichiamo l'encoding come 'utf-8' ATTENZIONE: MAI DIMENTICARE L'ENCODING !!
# - lo chiamiamo con f, un nome che scegliamo noi
with open("eventi.html", encoding='utf-8') as f:
soup = BeautifulSoup(f, "lxml") # creiamo un oggetto che chiamiamo 'soup', usando il parser lxml
# soup ci permette di chiamare il metodo select, per selezionare per esempio solo tag 'h4':
# il metodo ritorna una lista di tag trovate nel documento. Per ogni tag ritornata, la stampiamo:
for tag in soup.select("h4"):
print(tag)
```
Benissimo, abbiamo filtrato le tag coi nomi degli eventi. Ma noi vogliamo solo i nomi. Tipicamente dalle nostre tag ci interessa estrarre solo il testo. A tal fine. possiamo usare l'attributo `text` delle tag:
```
for tag in soup.select("h4"):
print(tag.text)
```
Casomai volessimo, si può anche estrarre un attributo, per esempio la `class`
```
for tag in soup.select('h4'):
print(tag['class'])
```
#### La lista nomi
Intanto, cerchiamo di mettere tutti i nomi in una lista che chiameremo `nomi`
```
nomi = []
for tag in soup.select("h4"):
nomi.append(tag.text)
nomi
```
#### Una struttura dati per il CSV
Cominciamo a costruirci la struttura dati che modellerà il CSV che vogliamo creare. Un CSV può essere visto come una lista di dizionari. Ogni dizionario rappresenterà un evento. Passo passo, vorremmo arrivare ad avere una forma simile:
```python
[{'data': '14/12/2017',
'descrizione': 'Al Passo Costalunga sfida tra i migliori specialisti del mondo',
'luogo': 'Passo Costalunga',
'nome': 'Coppa del Mondo di Snowboard',
'tipo': 'Sport, TOP EVENTI SPORT'},
{'data': '18/12/2017',
'descrizione': 'Lunedì 18 dicembre la Coppa Europa fa tappa in Val di Fassa',
'luogo': 'Pozza di Fassa',
'nome': 'Coppa Europa di sci alpino maschile - slalom speciale',
'tipo': 'Sport, TOP EVENTI SPORT'},
....
]
```
Inizializziamo la lista delle righe:
```
righe = []
righe
```
Dobbiamo aggiungere dei dizionari vuoti, ma quanti ce ne servono ? Vediamo quanti titoli abbiamo trovato con la funzione `len`:
```
len(nomi)
```
```
for i in range(16):
righe.append({})
righe
```
#### Popoliamo i dizionari con i nomi
Adesso, in ogni dizionario, mettiamo un campo 'nome' usando il valore corrispendente trovato nella lista `nomi`:
```
i = 0
for stringa in nomi:
righe[i]['nome'] = stringa # metti stringa nella riga i-esima, al campo 'nome' del dizionario
i += 1
```
Dovremmo cominciare a vedere i dizionari in `righe` popolati con i nomi. Verifichiamo:
```
righe
```
#### La funzione `aggiungi_campo`
Visto che i campi da aggiungere saranno diversi, definiamoci una funzione per aggiugere comodamente i campi alla variabile `righe`:
```
# 'attributo' potrebbe essere la stringa 'nome'
# 'lista' potrebbe essere la lista dei nomi degli eventi ['Coppa del Mondo di Snowboard', 'Coppa Europa', ...]
def aggiungi_campo(attributo, lista):
i = 0
for stringa in lista:
righe[i][attributo] = stringa
i += 1
return righe # ritorniamo righe per stampare immediatamente il risultato in Jupyter
```
Chiamando `aggiungi_campo` con i nomi, non dovrebbe cambiare nulla perchè andremo semplicemente a riscrivere i campi `nome` dentro i dizionari della lista `righe` :
```
aggiungi_campo('nome', nomi)
```
### 2.2 Estraiamo le date
Bene, è tempo di aggiungere un altro campo, per esempio per la data. Dove trovarlo?
**✪ DOMANDA 2.2.1**: Cerca dentro l'HTML il valore della data della Coppa del Mondo di Snowboard `14/12/2017`. Quanti valori trovi? Intorno alle date, trovi del testo che potrebbero permetterci di filtrare tutte e sole le date ?
**RISPOSTA**: Guardiamo l'HTML intorno alle posizioni `h4`:
```html
<div class="moodboard__item-text text-white">
<div>
<h4 class="moodboard__item-headline">Coppa del Mondo di Snowboard</h4>
<span class="moodboard__item-subline strong fz14 text-uppercase d-b">14/12/2017</span>
<span class="moodboard__item-subline strong fz14 d-b"><span class="icon icon-map-view fz20"></span> Passo Costalunga</span>
</div>
<div class="moodboard__item-text__link text-right">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" role="link"><span class="icon icon-circle-arrow fz30"></span></a>
</div>
</div>
```
Vediamo che le date sono dentro attributi `span`, questi attributi semplicemente indicano una sequenza di testo.
**✪ ESERCIZIO 2.2.2**: Prova a filtrare qui sotto in Python le tag `span`, stampando i risultati. Trovi solo le date ?
```
# scrivi qui il codice
# Le span sono troppe!
#for tag in soup.select("span"):
# print(tag)
```
#### Filtri più selettivi
Noi dobbiamo filtrare solo le tag `span` che contengono l'attributo `class` con un lungo valore criptico:
```
class="moodboard__item-subline strong fz14 text-uppercase d-b"
```
**NOTA**: Non farti spaventare da strani nomi che non conosci. Quando si traffica nei file HTML, si può trovare di tutto e bisogna 'navigare a vista'. Per fortuna, spesso non occorre sapere troppi dettagli tecnici per estrarre testo rilevante.
Per estrarre le nostre date, possiamo sfruttare il fatto che alla funzione `soup.select` possiamo passare non solo tag ma qualunque [espressione di selezione CSS](https://www.mrwebmaster.it/css/selettori-css3_11011.html)
Tra le prime ne troviamo una che ci dice che possiamo scrivere l'attributo e il valore dentro a parentesi quadre dopo il nome della tag, come qua:
```
for tag in soup.select('span[class="moodboard__item-subline strong fz14 text-uppercase d-b"]'):
print(tag.text)
```
Ecco le nostre date! In questo caso particolare, il `"moodboard__item-subline strong fz14 text-uppercase d-b"` è così identificativo che non serve nemmeno specificare `span`:
```
# nota che 'span' è rimosso:
for tag in soup.select('[class="moodboard__item-subline strong fz14 text-uppercase d-b"]'):
print(tag.text)
```
**✪ DOMANDA 2.2.3**: possiamo scrivere direttamente senza nient'altro la stringa `moodboard__item-subline strong fz14 text-uppercase d-b` nella `select` ? Scrivi qui sotto i tuoi esperimenti.
```
# scrivi qui
```
**RISPOSTA**: No, le regole CSS sono abbastanza rigide e non consentono ricerca di testo libero.
Come fatto prima, possiamo crearci una lista per le date:
```
date_eventi = []
for tag in soup.select('[class="moodboard__item-subline strong fz14 text-uppercase d-b"]'):
date_eventi.append(tag.text)
date_eventi
```
Per sincerarci di aver rastrellato tutte le date necessarie, possiamo controllare quante sono:
```
len(date_eventi)
```
Adesso possiamo sfruttare la funzione di prima `aggiungi_campo` per aggiornare `righe`:
```
aggiungi_campo('data', date_eventi)
```
### 2.3 Estraiamo i luoghi
E' tempo di aggiungere il luogo.
**✪✪ ESERCIZIO 2.3.1**: Cerca nell'HTML il luogo `Passo Costalunga` dell'evento Coppa di Snowboard. Con quale criterio possiamo filtrare i luoghi ?
**RISPOSTA**:
Possiamo usare la classe `class="moodboard__item-subline strong fz14 d-b"` nelle righe:
```html
<div class="moodboard__item-text text-white">
<div>
<h4 class="moodboard__item-headline">Coppa del Mondo di Snowboard</h4>
<span class="moodboard__item-subline strong fz14 text-uppercase d-b">14/12/2017</span>
<span class="moodboard__item-subline strong fz14 d-b"><span class="icon icon-map-view fz20"></span> Passo Costalunga</span>
</div>
<div class="moodboard__item-text__link text-right">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" role="link"><span class="icon icon-circle-arrow fz30"></span></a>
</div>
</div>
```
**✪ ESERCIZIO 2.3.2**: Scrivi il codice Python per estrarre i luoghi e metterlo nella lista `luoghi` (usa sempre l'html intorno ai blocchi `h4`)
**SUGGERIMENTO**: Non importa se il testo che cerchi è contenuto in uno span all'interno di un'altro span identificabile
```
# scrivi qui
for tag in soup.select('[class="moodboard__item-subline strong fz14 d-b"]'):
print(tag.text)
luoghi = []
for tag in soup.select('[class="moodboard__item-subline strong fz14 d-b"]'):
luoghi.append(tag.text)
luoghi
```
### 2.3 Sistemiamo i luoghi
**✪ DOMANDA 2.3.3**: Quanti luoghi hai trovato ? Dovrebbero essere 16, ma se ne hai trovato 15 allora manca un dato! Che dato è? Guarda la pagina e cerca se qualche evento non ha un luogo. In che posizione è nella lista ?
**RISPOSTA**: L'evento incriminato è `TrentinoSkiSunrise: sulle piste alla luce dell’alba`, guardando dentro l'html vediamo che manca proprio lo `span` con la classe:
```html
<div class="moodboard__item-text text-white">
<div>
<h4 class="moodboard__item-headline">TrentinoSkiSunrise: sulle piste alla luce dell’alba</h4>
<span class="moodboard__item-subline strong fz14 text-uppercase d-b">06/01/2018 - 07/04/2018</span>
</div>
<div class="moodboard__item-text__link text-right">
<a href="https://www.visittrentino.info/it/guida/eventi/trentinoskisunrise-sulle-piste-alla-luce-dell-alba_e_318133" role="link"><span class="icon icon-circle-arrow fz30"></span></a>
</div>
</div>
```
**✪ ESERCIZIO 2.3.4**: Come rimediare al problema? Ci sono metodo più furbi, ma per stavolta semplicemente possiamo inserire una stringa vuota alla posizione della lista `luoghi` in cui ci dovrebbe essere il campo mancante - e cioè dopo `Val di Fassa`. Per farlo, puoi usare il metodo `insert`. Qua puoi vedere qualche esempio di utilizzo.
**NOTA**: Ricordati che gli indici iniziano da 0 !
```
prova = ['a','b','c','d']
prova.insert(0, 'x') # inserisce 'x' all'inizio in posizione 0
prova
prova = ['a','b','c','d']
prova.insert(2, 'x') # inserisce 'x' a metà
prova
# scrivi qui la soluzione
luoghi.insert(4, '')
luoghi
```
**✪✪ ESERCIZIO 2.3.5**: Prima di procedere, verifica che la lunghezza della lista `len(luoghi)` sia corretta (= 16). Adesso, osserva attentamente le stringhe dei luoghi. Dovresti vedere che iniziano tutte con uno spazio inutile: sistema la lista `luoghi` togliendo gli spazi extra usando una _list comprehension_ (vedere [Capitolo 19.2 Pensare in Python](https://davidleoni.github.io/ThinkPythonItalian/html/thinkpython2020.html#sec227)). Per eliminare gli spazi puoi usare il metodo delle stringhe `.strip()`
**NOTA**: quando crei una _list comprehension_, ne generi una nuova, la lista originale _non_ viene modificata!
```
# scrivi qui
luoghi = [luogo.strip() for luogo in luoghi]
```
### 2.4 Tipo dell'evento
Proseguiamo con il tipo dell'evento. Per esempio, per la Coppa del Mondo di Snowboard, la tipologia dell'evento sarebbe `Sport, TOP EVENTI SPORT`
**✪ ESERCIZIO 2.4.1**: Cerca la stringa `Sport, TOP EVENTI SPORT` nell'HTML. Quante occorrenze ci sono ? E' possibile filtrare il tipo eventi in modo univoco ?
**RISPOSTA**:
Questa volta il tipo eventi appare solo nei blocchi intorno agli `h2`, in particolare notiamo la riga
`<span class="text-secondary fz14 text-uppercase strong">Sport, TOP EVENTI SPORT</span>` :
```html
<div class="col col-sm-9 arrange__item">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" class="text-primary list-teaser__link"><span class="icon icon-circle-arrow fz30"></span></a>
<div class="teaser__body">
<span class="text-secondary fz14 text-uppercase strong">Sport, TOP EVENTI SPORT</span>
<h2 class="text-secondary">Coppa del Mondo di Snowboard</h2>
<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>Passo Costalunga</a>
</li>
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>14/12/2017</a>
</li>
</ul>
<p>Al Passo Costalunga sfida tra i migliori specialisti del mondo</p>
</div>
</div>
```
**✪✪ ESERCIZIO 2.4.2**: Scrivi il codice Python per estrarre la lista degli eventi, e chiamala `tipo_evento`. Per farlo, puoi usare il codice già visto in precedenza, ma questa volta, per creare la lista sforzati di usare una _list comprehension_.
Verifica poi che la lunghezza della stringa sia 16.
**SUGGERIMENTO**: La lista originale da cui prelevare il testo delle le tag sarà creata dalla chiamata a `soup.select`
```
# scrivi qui
for tag in soup.select('[class="text-secondary fz14 text-uppercase strong"]'):
print(tag.text)
tipi_evento = [tag.text for tag in soup.select('[class="text-secondary fz14 text-uppercase strong"]')]
tipi_evento
```
### 2.5 descrizione
Rimane da aggiungere la descrizione, per esempio per la Coppa del Mondo di Snowboard la descrizione sarebbe "Al Passo Costalunga sfida tra i migliori specialisti del mondo".
**✪ ESERCIZIO 2.5.1**: Cerca manualmente nell'HTML la stringa `Al Passo Costalunga sfida tra i migliori specialisti del mondo`. In quante posizioni appare ? In base a quali criteri potremo filtrare i tipi degli eventi ? **NOTA**: In base a quanto sai finora, ti sarà impossibile trovare un selettore che possa filtrare quello che serve, quindi per ora pensa solo a trovare stringhe ricorrenti intorno alle descrizioni
**RISPOSTA**: Vediamo che compare in un blocco html regolare e come già successo prima in un blocco `script`. Concentriamoci sul blocco html. Vediamo che la descrizione sta solo vicino ai blocchi `h2`, in particolare alla linea
`<p>Al Passo Costalunga sfida tra i migliori specialisti del mondo</p>`:
```html
<div class="col col-sm-9 arrange__item">
<a href="https://www.visittrentino.info/it/guida/eventi/coppa-del-mondo-di-snowboard_e_165375" class="text-primary list-teaser__link"><span class="icon icon-circle-arrow fz30"></span></a>
<div class="teaser__body">
<span class="text-secondary fz14 text-uppercase strong">Sport, TOP EVENTI SPORT</span>
<h2 class="text-secondary">Coppa del Mondo di Snowboard</h2>
<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>Passo Costalunga</a>
</li>
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>14/12/2017</a>
</li>
</ul>
<p>Al Passo Costalunga sfida tra i migliori specialisti del mondo</p>
</div>
</div>
```
Stavolta il filtraggio non è così semplice come prima, perchè per `<p>` non c'è nessun comodo attributo come `class`. Possiamo però cercare se qualche attributo prima è più facilmente identificabile. Guardiamo per esempio la tag `<ul>` immediatamente precedente:
```html
<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>Passo Costalunga</a>
</li>
<li>
<a class="fz14 text-uppercase strong text-primary"><span class="icon icon-map-view mr10"></span>14/12/2017</a>
</li>
</ul>
<p>Al Passo Costalunga sfida tra i migliori specialisti del mondo</p>
```
Vediamo che la tag in questione `<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">` contiene una classe, se la cerchiamo nel documento troveremo che tutte queste `ul` con questa classe compaiono sempre e solo prima del `<p>` che vogliamo noi.
#### Trovare la descrizione in Python
Guarda la [soluzione dell'esercizio precedente](#2.5-descrizione). Troverai scritto che la tag `<ul class="list-unstyled list-inline list__teaser__list mb15 mt10">` contiene una classe, se la cerchiamo nel documento scopriremo che tutte queste `ul` con questa classe compaiono sempre e solo prima del `<p>` con le descrizioni che vogliamo noi. Non ci resta quindi che trovare un selettore CSS che
* trovi le tag `ul` con `class=list-unstyled list-inline list__teaser__list mb15 mt10`
* selezioni la tag successiva `p`
Per fare ciò possiamo usare il selettore col simbolo `+`, quindi nella select basterà scrivere:
```
soup.select('ul[class="list-unstyled list-inline list__teaser__list mb15 mt10"] + p')
```
Benissimo, abbiamo ottenuto la lista dei `<p>` che vogliamo. Adesso possiamo mettere il testo interno dei `<p>` dentro una lista che chiameremo `descrizioni`. Possiamo farlo in un colpo solo usando una _list comprehension_:
```
descrizioni = [tag.text for tag in soup.select('ul[class="list-unstyled list-inline list__teaser__list mb15 mt10"] + p')]
descrizioni
```
Come al solito, verifichiamo che la lista delle descrizioni sia di 16 elementi:
```
len(descrizioni)
```
### 2.6 Riempiamo la variabile `righe`
Ricapitolando tutto quanto visto precedentemente, popoliamo `righe` con i dizionari e tutti i campi per assicurarci di avere la struttura dati corretta. In questa riscrittura, usiamo di più le _list comprehnesion_ per avere codice più sintetico:
```
righe = []
for i in range(len(soup.select("h4"))):
righe.append({})
aggiungi_campo('nome', [tag.text for tag in soup.select("h4")])
aggiungi_campo('data', [tag.text for tag in soup.select('[class="moodboard__item-subline strong fz14 text-uppercase d-b"]')])
luoghi = [tag.text.strip() for tag in soup.select('[class="moodboard__item-subline strong fz14 d-b"]')]
luoghi.insert(4, '')
aggiungi_campo('luogo', luoghi)
aggiungi_campo('tipo', [tag.text for tag in soup.select('[class="text-secondary fz14 text-uppercase strong"]')])
aggiungi_campo('descrizione',
[tag.text for tag in soup.select('ul[class="list-unstyled list-inline list__teaser__list mb15 mt10"] + p')])
```
## 3. Scriviamo il CSV
**✪✪ ESERCIZIO 3.1**: Prova a scrivere un file CSV contenente tutti i dati raccolti, con quest'ordine di colonne:
`nome, data, luogo, tipo, descrizione`
Diversamente da quanto fatto finora, in cui abbiamo solo considerato righe come liste, in questo caso dato che abbiamo dizionari ci servirà un oggetto di tipo `DictWriter`. Leggi la [documentazione Python su DictWriter](https://docs.python.org/3/library/csv.html#csv.DictWriter) e copiandone l'esempio prova a scrivere il csv in un file chiamato `eventi.csv`.
**NOTA**: Per testare il tutto, fai attenzione a che nella cartella di questo progetto non sia già presente il file `eventi.csv`, in caso eliminalo a mano.
```
# scrivi qui
import csv
# apriamo il file `eventi.csv`in scrittura :
# 'fouput' è un nome scelto da noi)
# 'w' indica a Python che vogliamo scrivere il file
# se il file esiste già, verrà sovrascritto.
# NOTA: FATE SEMPRE MOLTA ATTENZIONE A NON SOVRASCRIVERE FILE PER SBAGLIO !!!!!
with open('eventi.csv', 'w', encoding='utf-8') as foutput:
# andremo a scrivere dizionari, ma nei dizionari le chiavi sono in ordine casuale:
# per avere una serie di colonne ordinata in modo prevedibile, siamo quindi obbligati
# a fornire una lista di intestazioni:
nomi_campi = ['nome', 'data', 'luogo', 'tipo', 'descrizione']
writer = csv.DictWriter(foutput,fieldnames=nomi_campi) # Ci serve creare un'oggetto 'scrittore' di tipo DictWriter
writer.writeheader()
for diz in righe: # stavolta le righe contengono dizionari
writer.writerow(diz) # chiamiamo l'oggetto scrittore dicendogli di scrivere il dizionario corrente
```
**✪✪✪ ESERCIZIO 3.2**: Abbiamo dei luoghi estratti dall'HTML. Nella [lezione sull'integrazione dati](http://it.softpython.org/data-integration.html), abbiamo imparato a usare Webapi di MapQuest per ottenere delle coordinate geografiche a partire da nomi di luoghi. Riusciresti quindi a:
* usare la libreria `requests` e webapi MapQuest per arricchire prima i dizionari di `righe` con `lat` e `lon`
* aggiungere `lat` e `lon` al CSV generato
* Caricare il CSV in Umap
```
# scrivi qui
```
| github_jupyter |
# Intro to Movie Review Sentiment Analysis

For the movie review sentiment analysis, we will be working on The Rotten Tomatoes movie review dataset from Kaggle.
Here, we'll have to label phrases on a scale of five values: negative, somewhat negative, neutral, somewhat positive, positive based on the sentiment of the movie reviews.
The dataset is comprised of tab-separated files with phrases from the Rotten Tomatoes dataset. Each phrase has a PhraseId. Each sentence has a SentenceId. Phrases that are repeated (such as short/common words) are only included once in the data.
The sentiment labels are:
* 0 - *negative*
* 1 - *somewhat negative*
* 2 - *neutral*
* 3 - *somewhat positive*
* 4 - *positive*
**Any suggestions for improvement or comments are highly appreciated!**
Please upvote(like button) and share this kernel if you like it so that more people can learn from it.
Below is the step by step methodology that we will be following :
- <a href='#1'>1. Initial Look at the Data</a>
- <a href='#1.1'>1.1 Distribution of reviews in each sentiment category</a>
- <a href='#1.2'>1.2 Dropping insignificant columns</a>
- <a href='#1.3'>1.3 Overall Distribution of the length of the reviews under each sentiment class</a>
- <a href='#1.4'>1.4 Creating Word Cloud of negative and positive movie reviews</a>
- <a href='#1.4.1'>1.4.1 Filtering out positive and negative movie reviews</a>
- <a href='#1.4.2'>1.4.2 Word Cloud for negatively classified movie reviews</a>
- <a href='#1.4.3'>1.4.3 Word Cloud for positively classified movie reviews</a>
- <a href='#1.5'>1.5 Term Frequencies of each Sentiment class</a>
- <a href='#1.5.1'>1.5.1 Term Frequency for 'negative' sentiments</a>
- <a href='#1.5.2'>1.5.2 Term Frequency for 'some negative' sentiments</a>
- <a href='#1.5.3'>1.5.3 Term Frequency for 'neutral' sentiments</a>
- <a href='#1.5.4'>1.5.4 Term Frequency for 'some positive' sentiments</a>
- <a href='#1.5.5'>1.5.5 Term Frequency for 'positive' sentiments</a>
- <a href='#1.6'>1.6 Total Term Frequency of all the 5 sentiment classes</a>
- <a href='#1.7'>1.7 Frequency plot of top frequent 500 phrases in movie reviews</a>
- <a href='#1.8'>1.8 Plot of Absolute frequency of phrases against their rank</a>
- <a href='#1.9'>1.9 Movie Reviews Tokens Visualisation</a>
- <a href='#1.9.1'>1.9.1 Plot of top frequently used 50 phrases in negative movie reviews</a>
- <a href='#1.9.2'>1.9.2 Plot of top frequently used 50 phrases in positive movie reviews</a>
- <a href='#2'>2. Traditional Supervised Machine Learning Models</a>
- <a href='#2.1'>2.1 Feature Engineering</a>
- <a href='#2.2'>2.2 Implementation of CountVectorizer & TF-IDF
- <a href='#2.2.1'>2.2.1 CountVectorizer</a>
- <a href='#2.2.2'>2.2.2 How is TF-IDF different from CountVectorizer?</a>
- <a href='#2.2.3'>2.2.3 How exactly does TF-IDF work?</a>
- <a href='#2.2.4'>2.2.4 Understanding the parameters of TfidfVectorizer</a>
- <a href='#2.2.5'>2.2.5 Setting the parametrs of CountVectorizer</a>
- <a href='#2.3'>2.3 Model Training, Prediction and Performance Evaluation</a>
- <a href='#2.3.1'>2.3.1 Logistic Regression model on CountVectorizer</a>
- <a href='#2.3.2'>2.3.2 Logistic Regression model on TF-IDF features</a>
- <a href='#2.3.3'>2.3.3 SGD model on Countvectorizer</a>
- <a href='#2.3.4'>2.3.4 SGD model on TF-IDF</a>
- <a href='#2.3.5'>2.3.5 RandomForest model on TF-IDF</a>
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
```
## <a id='1'>1. Initial Look at the Data</a>
```
df_train = pd.read_csv("../input/train.tsv", sep='\t')
df_train.head()
df_test = pd.read_csv("../input/test.tsv", sep='\t')
df_test.head()
```
## <a id='1.1'>1.1 Distribution of reviews in each sentiment category</a>
Here, the training dataset has dominating neutral phrases from the movie reviews followed by somewhat positive and then somewhat negative.
```
df_train.Sentiment.value_counts()
df_train.info()
```
## <a id='1.2'>1.2 Dropping insignificant columns</a>
```
df_train_1 = df_train.drop(['PhraseId','SentenceId'],axis=1)
df_train_1.head()
```
Let's check the phrase length of each of the movie reviews.
```
df_train_1['phrase_len'] = [len(t) for t in df_train_1.Phrase]
df_train_1.head(4)
```
## <a id='1.3'>1.3 Overall Distribution of the length of the reviews under each sentiment class</a>
```
fig,ax = plt.subplots(figsize=(5,5))
plt.boxplot(df_train_1.phrase_len)
plt.show()
```
From the above box plot, some of the reviews are way more than 100 chracters long.
```
df_train_1[df_train_1.phrase_len > 100].head()
df_train_1[df_train_1.phrase_len > 100].loc[0].Phrase
```
## <a id='1.4'>1.4 Creating Word Cloud of negative and positive movie reviews</a>
### Word Cloud
A word cloud is a graphical representation of frequently used words in a collection of text files. The height of each word in this picture is an indication of frequency of occurrence of the word in the entire text. Such diagrams are very useful when doing text analytics.
It provides a general idea of what kind of words are frequent in the corpus, in a sort of quick and dirty way.
Let's start doing some EDA on text data by Word Cloud.
## <a id='1.4.1'>1.4.1 Filtering out positive and negative movie reviews</a>
```
neg_phrases = df_train_1[df_train_1.Sentiment == 0]
neg_words = []
for t in neg_phrases.Phrase:
neg_words.append(t)
neg_words[:4]
```
**pandas.Series.str.cat ** : Concatenate strings in the Series/Index with given separator. Here we give a space as separator, so, it will concatenate all the strings in each of the index separated by a space.
```
neg_text = pd.Series(neg_words).str.cat(sep=' ')
neg_text[:100]
for t in neg_phrases.Phrase[:300]:
if 'good' in t:
print(t)
```
So, we can very well see, even if the texts contain words like "good", it is a negative sentiment because it indicates that the movie is **NOT** a good movie.
```
pos_phrases = df_train_1[df_train_1.Sentiment == 4] ## 4 is positive sentiment
pos_string = []
for t in pos_phrases.Phrase:
pos_string.append(t)
pos_text = pd.Series(pos_string).str.cat(sep=' ')
pos_text[:100]
```
## <a id='1.4.2'>1.4.2 Word Cloud for negatively classified movie reviews</a>
```
from wordcloud import WordCloud
wordcloud = WordCloud(width=1600, height=800, max_font_size=200).generate(neg_text)
plt.figure(figsize=(12,10))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis("off")
plt.show()
```
Some of the big words can be interpreted quite neutral, such as "movie","film", etc. We can see some of the words in smaller size make sense to be in negative movie reviews like "bad cinema", "annoying", "dull", etc.
However, there are some words like "good" is also present in the negatively classified sentiment about the movie.
Let's go deeper into such words/texts:
## <a id='1.4.3'>1.4.3 Word Cloud for positively classified movie reviews</a>
```
wordcloud = WordCloud(width=1600, height=800, max_font_size=200).generate(pos_text)
plt.figure(figsize=(12,10))
plt.imshow(wordcloud, interpolation='bilinear')
plt.axis('off')
plt.show()
```
Again I see some neutral words in big size, "movie","film", but positive words like "good", "best", "fascinating" also stand out.
## <a id='1.5'>1.5 Term Frequencies of each Sentiment class</a>
We also want to understand how terms are distributed across documents. This helps us to characterize the properties of the algorithms for compressing phrases.
A commonly used model of the distribution of terms in a collection is Zipf's law . It states that, if $t_1$ is the most common term in the collection, $t_2$ is the next most common, and so on, then the collection frequency $cf_i$ of the $i$th most common term is proportional to $1/i$:
$\displaystyle cf_i \propto \frac{1}{i}.$
So if the most frequent term occurs $cf_1$ times, then the second most frequent term has half as many occurrences, the third most frequent term a third as many occurrences, and so on. The intuition is that frequency decreases very rapidly with rank.
The above equation is one of the simplest ways of formalizing such a rapid decrease and it has been found to be a reasonably good model.
We need the Term Frequency data to see what kind of words are used in the movie reviews and how many times have been used.
Let's proceed with CountVectorizer to calculate term frequencies:
```
from sklearn.feature_extraction.text import CountVectorizer
cvector = CountVectorizer(min_df = 0.0, max_df = 1.0, ngram_range=(1,2))
cvector.fit(df_train_1.Phrase)
len(cvector.get_feature_names())
```
It looks like count vectorizer has extracted 94644 words out of the corpus.
Getting term frequency for each class can be obtained with the below code block.
## <a id='1.5.1'>1.5.1 Term Frequency for 'negative' sentiments</a>
```
neg_matrix = cvector.transform(df_train_1[df_train_1.Sentiment == 0].Phrase)
som_neg_matrix = cvector.transform(df_train_1[df_train_1.Sentiment == 1].Phrase)
neu_matrix = cvector.transform(df_train_1[df_train_1.Sentiment == 2].Phrase)
som_pos_matrix = cvector.transform(df_train_1[df_train_1.Sentiment == 3].Phrase)
pos_matrix = cvector.transform(df_train_1[df_train_1.Sentiment == 4].Phrase)
neg_words = neg_matrix.sum(axis=0)
neg_words_freq = [(word, neg_words[0, idx]) for word, idx in cvector.vocabulary_.items()]
neg_tf = pd.DataFrame(list(sorted(neg_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','negative'])
neg_tf.head()
neg_tf_df = neg_tf.set_index('Terms')
neg_tf_df.head()
```
## <a id='1.5.2'>1.5.2 Term Frequency for 'some negative' sentiments</a>
```
som_neg_words = som_neg_matrix.sum(axis=0)
som_neg_words_freq = [(word, som_neg_words[0, idx]) for word, idx in cvector.vocabulary_.items()]
som_neg_tf = pd.DataFrame(list(sorted(som_neg_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','some-negative'])
som_neg_tf_df = som_neg_tf.set_index('Terms')
som_neg_tf_df.head()
```
## <a id='1.5.3'>1.5.3 Term Frequency for 'neutral' sentiments</a>
```
neu_words = neu_matrix.sum(axis=0)
neu_words_freq = [(word, neu_words[0, idx]) for word, idx in cvector.vocabulary_.items()]
neu_words_tf = pd.DataFrame(list(sorted(neu_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','neutral'])
neu_words_tf_df = neu_words_tf.set_index('Terms')
neu_words_tf_df.head()
```
## <a id='1.5.4'>1.5.4 Term Frequency for 'some positive' sentiments</a>
```
som_pos_words = som_pos_matrix.sum(axis=0)
som_pos_words_freq = [(word, som_pos_words[0, idx]) for word, idx in cvector.vocabulary_.items()]
som_pos_words_tf = pd.DataFrame(list(sorted(som_pos_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','some-positive'])
som_pos_words_tf_df = som_pos_words_tf.set_index('Terms')
som_pos_words_tf_df.head()
```
## <a id='1.5.5'>1.5.5 Term Frequency for 'positive' sentiments</a>
```
pos_words = pos_matrix.sum(axis=0)
pos_words_freq = [(word, pos_words[0, idx]) for word, idx in cvector.vocabulary_.items()]
pos_words_tf = pd.DataFrame(list(sorted(pos_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','positive'])
pos_words_tf_df = pos_words_tf.set_index('Terms')
pos_words_tf_df.head()
term_freq_df = pd.concat([neg_tf_df,som_neg_tf_df,neu_words_tf_df,som_pos_words_tf_df,pos_words_tf_df],axis=1)
```
## <a id='1.6'>1.6 Total Term Frequency of all the 5 sentiment classes</a>
```
term_freq_df['total'] = term_freq_df['negative'] + term_freq_df['some-negative'] \
+ term_freq_df['neutral'] + term_freq_df['some-positive'] \
+ term_freq_df['positive']
term_freq_df.sort_values(by='total', ascending=False).head(20)
```
## <a id='1.7'>1.7 Frequency plot of top frequent 500 phrases in movie reviews</a>
**"Given some corpus of natural language utterances, the frequency of any word is inversely proportional to its rank in the frequency table. Thus the most frequent word will occur approximately twice as often as the second most frequent word, three times as often as the third most frequent word, etc."**
In other words, the rth most frequent word has a frequency f(r) that scales according to $${f(r)} \propto \frac{1}{r^\alpha}$$ for $$\alpha \approx {1}$$
Let's see how the movie review tokens and their frequencies look like on a plot.
```
y_pos = np.arange(500)
plt.figure(figsize=(10,8))
s = 1
expected_zipf = [term_freq_df.sort_values(by='total', ascending=False)['total'][0]/(i+1)**s for i in y_pos]
plt.bar(y_pos, term_freq_df.sort_values(by='total', ascending=False)['total'][:500], align='center', alpha=0.5)
plt.plot(y_pos, expected_zipf, color='r', linestyle='--',linewidth=2,alpha=0.5)
plt.ylabel('Frequency')
plt.title('Top 500 phrases in movie reviews')
```
On the X-axis is the rank of the frequency from highest rank from left up to 500th rank to the right. Y-axis is the frequency observed in the corpus.
Another way to plot this is on a log-log graph, with X-axis being log(rank), Y-axis being log(frequency). By plotting on the log-log scale the result will yield roughly linear line on the graph.
## <a id='1.8'>1.8 Plot of Absolute frequency of phrases against their rank</a>
```
from pylab import *
counts = term_freq_df.total
tokens = term_freq_df.index
ranks = arange(1, len(counts)+1)
indices = argsort(-counts)
frequencies = counts[indices]
plt.figure(figsize=(8,6))
plt.ylim(1,10**6)
plt.xlim(1,10**6)
loglog(ranks, frequencies, marker=".")
plt.plot([1,frequencies[0]],[frequencies[0],1],color='r')
title("Zipf plot for phrases tokens")
xlabel("Frequency rank of token")
ylabel("Absolute frequency of token")
grid(True)
for n in list(logspace(-0.5, log10(len(counts)-2), 25).astype(int)):
dummy = text(ranks[n], frequencies[n], " " + tokens[indices[n]],
verticalalignment="bottom",
horizontalalignment="left")
```
We can clearly see that words like "the", "in","it", etc are much higher in frequency but has been ranked less as they don't have any significance regarding the sentiment of the movie review. On the other hand, some words like "downbeat laughably" have been given higher rank as they are very less frequent in the document and seems to be significant related to the sentiment of a movie.
## <a id='1.9'>1.9 Movie Reviews Tokens Visualisation</a>
Next, let's explore about how different the tokens in two different classes(positive, negative).
```
from sklearn.feature_extraction.text import CountVectorizer
cvec = CountVectorizer(stop_words='english',max_features=10000)
cvec.fit(df_train_1.Phrase)
neg_matrix = cvec.transform(df_train_1[df_train_1.Sentiment == 0].Phrase)
som_neg_matrix = cvec.transform(df_train_1[df_train_1.Sentiment == 1].Phrase)
neu_matrix = cvec.transform(df_train_1[df_train_1.Sentiment == 2].Phrase)
som_pos_matrix = cvec.transform(df_train_1[df_train_1.Sentiment == 3].Phrase)
pos_matrix = cvec.transform(df_train_1[df_train_1.Sentiment == 4].Phrase)
neg_words = neg_matrix.sum(axis=0)
neg_words_freq = [(word, neg_words[0, idx]) for word, idx in cvec.vocabulary_.items()]
neg_tf = pd.DataFrame(list(sorted(neg_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','negative'])
neg_tf_df = neg_tf.set_index('Terms')
som_neg_words = som_neg_matrix.sum(axis=0)
som_neg_words_freq = [(word, som_neg_words[0, idx]) for word, idx in cvec.vocabulary_.items()]
som_neg_tf = pd.DataFrame(list(sorted(som_neg_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','some-negative'])
som_neg_tf_df = som_neg_tf.set_index('Terms')
neu_words = neu_matrix.sum(axis=0)
neu_words_freq = [(word, neu_words[0, idx]) for word, idx in cvec.vocabulary_.items()]
neu_words_tf = pd.DataFrame(list(sorted(neu_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','neutral'])
neu_words_tf_df = neu_words_tf.set_index('Terms')
som_pos_words = som_pos_matrix.sum(axis=0)
som_pos_words_freq = [(word, som_pos_words[0, idx]) for word, idx in cvec.vocabulary_.items()]
som_pos_words_tf = pd.DataFrame(list(sorted(som_pos_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','some-positive'])
som_pos_words_tf_df = som_pos_words_tf.set_index('Terms')
pos_words = pos_matrix.sum(axis=0)
pos_words_freq = [(word, pos_words[0, idx]) for word, idx in cvec.vocabulary_.items()]
pos_words_tf = pd.DataFrame(list(sorted(pos_words_freq, key = lambda x: x[1], reverse=True)),columns=['Terms','positive'])
pos_words_tf_df = pos_words_tf.set_index('Terms')
term_freq_df = pd.concat([neg_tf_df,som_neg_tf_df,neu_words_tf_df,som_pos_words_tf_df,pos_words_tf_df],axis=1)
term_freq_df['total'] = term_freq_df['negative'] + term_freq_df['some-negative'] \
+ term_freq_df['neutral'] + term_freq_df['some-positive'] \
+ term_freq_df['positive']
term_freq_df.sort_values(by='total', ascending=False).head(15)
```
## <a id='1.9.1'>1.9.1 Plot of top frequently used 50 phrases in negative movie reviews</a>
```
y_pos = np.arange(50)
plt.figure(figsize=(12,10))
plt.bar(y_pos, term_freq_df.sort_values(by='negative', ascending=False)['negative'][:50], align='center', alpha=0.5)
plt.xticks(y_pos, term_freq_df.sort_values(by='negative', ascending=False)['negative'][:50].index,rotation='vertical')
plt.ylabel('Frequency')
plt.xlabel('Top 50 negative tokens')
plt.title('Top 50 tokens in negative movie reviews')
```
We can see some negative words like "bad", "worst", "dull" are some of the high frequency words. But, there exists few neutral words like "movie", "film", "minutes" dominating the frequency plots.
Let's also take a look at top 50 positive tokens on a bar chart.
## <a id='1.9.2'>1.9.2 Plot of top frequently used 50 phrases in positive movie reviews</a>
```
y_pos = np.arange(50)
plt.figure(figsize=(12,10))
plt.bar(y_pos, term_freq_df.sort_values(by='positive', ascending=False)['positive'][:50], align='center', alpha=0.5)
plt.xticks(y_pos, term_freq_df.sort_values(by='positive', ascending=False)['positive'][:50].index,rotation='vertical')
plt.ylabel('Frequency')
plt.xlabel('Top 50 positive tokens')
plt.title('Top 50 tokens in positive movie reviews')
```
Once again, there are some neutral words like "film", "movie", are quite high up in the rank.
## <a id='2'>2. Traditional Supervised Machine Learning Models</a>
## <a id='2.1'>2.1 Feature Engineering</a>
```
phrase = np.array(df_train_1['Phrase'])
sentiments = np.array(df_train_1['Sentiment'])
# build train and test datasets
from sklearn.model_selection import train_test_split
phrase_train, phrase_test, sentiments_train, sentiments_test = train_test_split(phrase, sentiments, test_size=0.2, random_state=4)
```
Next, we will try to see how different are the tokens in 4 different classes(positive,some positive,neutral, some negative, negative).
## <a id='2.2'>2.2 Implementation of CountVectorizer & TF-IDF
## <a id='2.2.1'>2.2.1 CountVectorizer</a>
As we all know, all machine learning algorithms are good with numbers; we have to extract or convert the text data into numbers without losing much of the information.
One way to do such transformation is Bag-Of-Words (BOW) which gives a number to each word but that is very inefficient.
So, a way to do it is by **CountVectorizer**: it counts the number of words in the document i.e it converts a collection of text documents to a matrix of the counts of occurences of each word in the document.
For Example: If we have a collection of 3 text documents as below, then CountVectorizer converts that into individual counts of occurences of each of the words in the document as below:
```
cv1 = CountVectorizer()
x_traincv = cv1.fit_transform(["Hi How are you How are you doing","Hi what's up","Wow that's awesome"])
x_traincv_df = pd.DataFrame(x_traincv.toarray(),columns=list(cv1.get_feature_names()))
x_traincv_df
```
Now, in case of CountVectorizer, we are just counting the number of words in the document and many times it happens that some words like "are","you","hi",etc are very large in numbers and that would dominate our results in machinelearning algorithm.
## <a id='2.2.2'>2.2.2 How is TF-IDF different from CountVectorizer?</a>
So, TF-IDF (stands for **Term-Frequency-Inverse-Document Frequency**) weights down the common words occuring in almost all the documents and give more importance to the words that appear in a subset of documents.
TF-IDF works by penalising these common words by assigning them lower weights while giving importance to some rare words in a particular document.
## <a id='2.2.3'>2.2.3 How exactly does TF-IDF work?</a>
Consider the below sample table which gives the count of terms(tokens/words) in two documents.

Now, let us define a few terms related to TF-IDF.
**TF (Term Frequency)** :
Denotes the contribution of the word to the document i.e. words relevant to the document should be frequent.
TF = (Number of times term t appears in a document)/(Number of terms in the document)
So, TF(This,Document1) = 1/8
TF(This, Document2)=1/5
**IDF (Inverse Document Frequency)** :
If a word has appeared in all the document, then probably that word is not relevant to a particular document.
But, if it has appeared in a subset of documents then probably the word is of some relevance to the documents it is present in.
IDF = log(N/n), where, N is the number of documents and n is the number of documents a term t has appeared in.
So, IDF(This) = log(2/2) = 0.
IDF(Messi) = log(2/1) = 0.301.
Now, let us compare the TF-IDF for a common word ‘This’ and a word ‘Messi’ which seems to be of relevance to Document 1.
TF-IDF(This,Document1) = (1/8) * (0) = 0
TF-IDF(This, Document2) = (1/5) * (0) = 0
TF-IDF(Messi, Document1) = (4/8) * 0.301 = 0.15
So, for Document1 , TF-IDF method heavily penalises the word ‘This’ but assigns greater weight to ‘Messi’. So, this may be understood as ‘Messi’ is an important word for Document1 from the context of the entire corpus.
## "Rare terms are more informative than frequent terms"
The graphic below attempts to express this intuition. Note that the TF-IDF weight is a relative measurement, so the values in red on the axis are not intended to be taken as absolute weights.

When your corpus (or Structured set of texts) is large, TfIdf is the best option.
Now, let's get back to our problem:
## <a id='2.2.4'>2.2.4 Understanding the parameters of TfidfVectorizer</a>
* min_df : While building the vocabulary, it will ignore terms that have a document frequency strictly lower than the given threshold. In our case, threshold for min_df = 0.0
* max_df : While building the vocabulary, it ignore terms that have a document frequency strictly higher than the given threshold. For us, threshold for max_df = 1.0
* ngram_range : A tuple of lower and upper boundary of the range of n-values for different n-grams to be extracted.

* sublinear_tf : Sublinear tf scaling addresses the problem that 20 occurrences of a word is probably not 20 times more important than 1 occurrence.

** Why is log used when calculating term frequency weight and IDF, inverse document frequency in sublinear_tf transformation?**
Found the answer to this question in Stackoverflow forum which you may find useful.

## <a id='2.2.5'>2.2.5 Setting the parameters of CountVectorizer</a>
**For CountVectorizer**
This time, the stop words will not help much, because of the same high-frequency words, such as "the", "to", will equally frequent in both classes. If these stop words dominate both of the classes, I won't be able to have a meaningful result. So, I decided to remove stop words, and also will limit the max_features to 10,000 with countvectorizer.
```
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
## Build Bag-Of-Words on train phrases
cv = CountVectorizer(stop_words='english',max_features=10000)
cv_train_features = cv.fit_transform(phrase_train)
# build TFIDF features on train reviews
tv = TfidfVectorizer(min_df=0.0, max_df=1.0, ngram_range=(1,2),
sublinear_tf=True)
tv_train_features = tv.fit_transform(phrase_train)
# transform test reviews into features
cv_test_features = cv.transform(phrase_test)
tv_test_features = tv.transform(phrase_test)
print('BOW model:> Train features shape:', cv_train_features.shape, ' Test features shape:', cv_test_features.shape)
print('TFIDF model:> Train features shape:', tv_train_features.shape, ' Test features shape:', tv_test_features.shape)
```
## <a id='2.3'>2.3 Model Training, Prediction and Performance Evaluation</a>
```
####Evaluation metrics
from sklearn import metrics
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from sklearn.base import clone
from sklearn.preprocessing import label_binarize
from scipy import interp
from sklearn.metrics import roc_curve, auc
def get_metrics(true_labels, predicted_labels):
print('Accuracy:', np.round(
metrics.accuracy_score(true_labels,
predicted_labels),
4))
print('Precision:', np.round(
metrics.precision_score(true_labels,
predicted_labels,
average='weighted'),
4))
print('Recall:', np.round(
metrics.recall_score(true_labels,
predicted_labels,
average='weighted'),
4))
print('F1 Score:', np.round(
metrics.f1_score(true_labels,
predicted_labels,
average='weighted'),
4))
def train_predict_model(classifier,
train_features, train_labels,
test_features, test_labels):
# build model
classifier.fit(train_features, train_labels)
# predict using model
predictions = classifier.predict(test_features)
return predictions
def display_confusion_matrix(true_labels, predicted_labels, classes=[1,0]):
total_classes = len(classes)
level_labels = [total_classes*[0], list(range(total_classes))]
cm = metrics.confusion_matrix(y_true=true_labels, y_pred=predicted_labels,
labels=classes)
cm_frame = pd.DataFrame(data=cm,
columns=pd.MultiIndex(levels=[['Predicted:'], classes],
labels=level_labels),
index=pd.MultiIndex(levels=[['Actual:'], classes],
labels=level_labels))
print(cm_frame)
def display_classification_report(true_labels, predicted_labels, classes=[1,0]):
report = metrics.classification_report(y_true=true_labels,
y_pred=predicted_labels,
labels=classes)
print(report)
def display_model_performance_metrics(true_labels, predicted_labels, classes=[1,0]):
print('Model Performance metrics:')
print('-'*30)
get_metrics(true_labels=true_labels, predicted_labels=predicted_labels)
print('\nModel Classification report:')
print('-'*30)
display_classification_report(true_labels=true_labels, predicted_labels=predicted_labels,
classes=classes)
print('\nPrediction Confusion Matrix:')
print('-'*30)
display_confusion_matrix(true_labels=true_labels, predicted_labels=predicted_labels,
classes=classes)
def plot_model_decision_surface(clf, train_features, train_labels,
plot_step=0.02, cmap=plt.cm.RdYlBu,
markers=None, alphas=None, colors=None):
if train_features.shape[1] != 2:
raise ValueError("X_train should have exactly 2 columnns!")
x_min, x_max = train_features[:, 0].min() - plot_step, train_features[:, 0].max() + plot_step
y_min, y_max = train_features[:, 1].min() - plot_step, train_features[:, 1].max() + plot_step
xx, yy = np.meshgrid(np.arange(x_min, x_max, plot_step),
np.arange(y_min, y_max, plot_step))
clf_est = clone(clf)
clf_est.fit(train_features,train_labels)
if hasattr(clf_est, 'predict_proba'):
Z = clf_est.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:,1]
else:
Z = clf_est.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
cs = plt.contourf(xx, yy, Z, cmap=cmap)
le = LabelEncoder()
y_enc = le.fit_transform(train_labels)
n_classes = len(le.classes_)
plot_colors = ''.join(colors) if colors else [None] * n_classes
label_names = le.classes_
markers = markers if markers else [None] * n_classes
alphas = alphas if alphas else [None] * n_classes
for i, color in zip(range(n_classes), plot_colors):
idx = np.where(y_enc == i)
plt.scatter(train_features[idx, 0], train_features[idx, 1], c=color,
label=label_names[i], cmap=cmap, edgecolors='black',
marker=markers[i], alpha=alphas[i])
plt.legend()
plt.show()
def plot_model_roc_curve(clf, features, true_labels, label_encoder=None, class_names=None):
## Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
if hasattr(clf, 'classes_'):
class_labels = clf.classes_
elif label_encoder:
class_labels = label_encoder.classes_
elif class_names:
class_labels = class_names
else:
raise ValueError('Unable to derive prediction classes, please specify class_names!')
n_classes = len(class_labels)
y_test = label_binarize(true_labels, classes=class_labels)
if n_classes == 2:
if hasattr(clf, 'predict_proba'):
prob = clf.predict_proba(features)
y_score = prob[:, prob.shape[1]-1]
elif hasattr(clf, 'decision_function'):
prob = clf.decision_function(features)
y_score = prob[:, prob.shape[1]-1]
else:
raise AttributeError("Estimator doesn't have a probability or confidence scoring system!")
fpr, tpr, _ = roc_curve(y_test, y_score)
roc_auc = auc(fpr, tpr)
plt.plot(fpr, tpr, label='ROC curve (area = {0:0.2f})'
''.format(roc_auc),
linewidth=2.5)
elif n_classes > 2:
if hasattr(clf, 'predict_proba'):
y_score = clf.predict_proba(features)
elif hasattr(clf, 'decision_function'):
y_score = clf.decision_function(features)
else:
raise AttributeError("Estimator doesn't have a probability or confidence scoring system!")
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_test[:, i], y_score[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
## Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_test.ravel(), y_score.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
## Compute macro-average ROC curve and ROC area
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
## Plot ROC curves
plt.figure(figsize=(6, 4))
plt.plot(fpr["micro"], tpr["micro"],
label='micro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["micro"]), linewidth=3)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["macro"]), linewidth=3)
for i, label in enumerate(class_labels):
plt.plot(fpr[i], tpr[i], label='ROC curve of class {0} (area = {1:0.2f})'
''.format(label, roc_auc[i]),
linewidth=2, linestyle=':')
else:
raise ValueError('Number of classes should be atleast 2 or more')
plt.plot([0, 1], [0, 1], 'k--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver Operating Characteristic (ROC) Curve')
plt.legend(loc="lower right")
plt.show()
from sklearn.linear_model import SGDClassifier, LogisticRegression
lr = LogisticRegression(penalty='l2', max_iter=100, C=1)
sgd = SGDClassifier(loss='hinge', n_iter=100)
```
## <a id='2.3.1'>2.3.1 Logistic Regression model on CountVectorizer</a>
```
# Logistic Regression model on BOW features
lr_bow_predictions = train_predict_model(classifier=lr,
train_features=cv_train_features, train_labels=sentiments_train,
test_features=cv_test_features, test_labels=sentiments_test)
display_model_performance_metrics(true_labels=sentiments_test, predicted_labels=lr_bow_predictions,
classes=[0,1,2,3,4])
```
## <a id='2.3.2'>2.3.2 Logistic Regression model on TF-IDF features</a>
```
# Logistic Regression model on TF-IDF features
lr_tfidf_predictions = train_predict_model(classifier=lr,
train_features=tv_train_features, train_labels=sentiments_train,
test_features=tv_test_features, test_labels=sentiments_test)
display_model_performance_metrics(true_labels=sentiments_test, predicted_labels=lr_tfidf_predictions,
classes=[0,1,2,3,4])
```
## <a id='2.3.3'>2.3.3 SGD model on Countvectorizer</a>
```
# SGD model on Countvectorizer
sgd_bow_predictions = train_predict_model(classifier=sgd,
train_features=cv_train_features, train_labels=sentiments_train,
test_features=cv_test_features, test_labels=sentiments_test)
display_model_performance_metrics(true_labels=sentiments_test, predicted_labels=sgd_bow_predictions,
classes=[0,1,2,3,4])
```
## <a id='2.3.4'>2.3.4 SGD model on TF-IDF</a>
```
# SGD model on TF-IDF
sgd_tfidf_predictions = train_predict_model(classifier=sgd,
train_features=tv_train_features, train_labels=sentiments_train,
test_features=tv_test_features, test_labels=sentiments_test)
display_model_performance_metrics(true_labels=sentiments_test, predicted_labels=sgd_tfidf_predictions,
classes=[0,1,2,3,4])
```
## <a id='2.3.5'>2.3.5 RandomForest model on TF-IDF</a>
```
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_jobs=-1)
# RandomForest model on TF-IDF
rfc_tfidf_predictions = train_predict_model(classifier=rfc,
train_features=tv_train_features, train_labels=sentiments_train,
test_features=tv_test_features, test_labels=sentiments_test)
display_model_performance_metrics(true_labels=sentiments_test, predicted_labels=rfc_tfidf_predictions,
classes=[0,1,2,3,4])
```
**Logistic Regression on TF-IDF is outperforming other machine learning algorithms**.
| github_jupyter |
```
# Copyright 2020 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
<img src="http://developer.download.nvidia.com/compute/machine-learning/frameworks/nvidia_logo.png" style="width: 90px; float: right;">
# HugeCTR demo on Movie lens data
## Overview
HugeCTR is a recommender specific framework which is capable of distributed training across multiple GPUs and nodes for Click-Through-Rate (CTR) estimation. It is a component of NVIDIA [Merlin](https://developer.nvidia.com/nvidia-merlin#getstarted), which is a framework accelerating the entire pipeline from data ingestion and training to deploying GPU-accelerated recommender systems.
### Learning objectives
This notebook demonstrates the steps for training a deep learning recommender model (DLRM) on the movie lens 20M [dataset](https://grouplens.org/datasets/movielens/20m/). We will walk you through the process of data preprocessing, train a DLRM model with HugeCTR, then using the movie embedding to answer item similarity queries.
## Content
1. [Pre-requisite](#1)
1. [Data download and preprocessing](#2)
1. [HugeCTR DLRM training](#3)
1. [Answer item similarity with DLRM embedding](#4)
<a id="1"></a>
## 1. Pre-requisite
### 1.1 Docker containers
Follow the steps in [README](README.md) to build and start a HugeCTR development Docker container for the experiments.
### 1.2 Hardware
This notebook requires a Pascal, Volta, Turing, Ampere or newer GPUs, such as P100, V100, T4 or A100.
```
!nvidia-smi
```
<a id="2"></a>
## 2. Data download and preprocessing
We first install a few extra utilities for data preprocessing.
```
!pip3 install torch tqdm
!apt install unzip
```
Next, we download and unzip the movie lens 20M [dataset](https://grouplens.org/datasets/movielens/20m/).
```
%%bash
mkdir -p data
cd data
if [ ! -f "ml-20m.zip" ]; then
echo "Downloading data"
wget http://files.grouplens.org/datasets/movielens/ml-20m.zip
unzip ml-20m.zip
fi
!ls ./data
```
### Movie lens data preprocessing
```
from argparse import ArgumentParser
import pandas as pd
import torch
import tqdm
MIN_RATINGS = 20
USER_COLUMN = 'userId'
ITEM_COLUMN = 'movieId'
```
Since the movie lens data contains only positive examples, let us first define an utility function to generate negative samples.
```
class _TestNegSampler:
def __init__(self, train_ratings, nb_users, nb_items, nb_neg):
self.nb_neg = nb_neg
self.nb_users = nb_users
self.nb_items = nb_items
# compute unique ids for quickly created hash set and fast lookup
ids = (train_ratings[:, 0] * self.nb_items) + train_ratings[:, 1]
self.set = set(ids)
def generate(self, batch_size=128*1024):
users = torch.arange(0, self.nb_users).reshape([1, -1]).repeat([self.nb_neg, 1]).transpose(0, 1).reshape(-1)
items = [-1] * len(users)
random_items = torch.LongTensor(batch_size).random_(0, self.nb_items).tolist()
print('Generating validation negatives...')
for idx, u in enumerate(tqdm.tqdm(users.tolist())):
if not random_items:
random_items = torch.LongTensor(batch_size).random_(0, self.nb_items).tolist()
j = random_items.pop()
while u * self.nb_items + j in self.set:
if not random_items:
random_items = torch.LongTensor(batch_size).random_(0, self.nb_items).tolist()
j = random_items.pop()
items[idx] = j
items = torch.LongTensor(items)
return items
```
Next, we read the data into a Pandas dataframe, and encode userID and itemID with integers.
```
df = pd.read_csv('./data/ml-20m/ratings.csv')
print("Filtering out users with less than {} ratings".format(MIN_RATINGS))
grouped = df.groupby(USER_COLUMN)
df = grouped.filter(lambda x: len(x) >= MIN_RATINGS)
print("Mapping original user and item IDs to new sequential IDs")
df[USER_COLUMN], unique_users = pd.factorize(df[USER_COLUMN])
df[ITEM_COLUMN], unique_items = pd.factorize(df[ITEM_COLUMN])
nb_users = len(unique_users)
nb_items = len(unique_items)
print("Number of users: %d\nNumber of items: %d"%(len(unique_users), len(unique_items)))
# Save the mapping to do the inference later on
import pickle
with open('./mappings.pickle', 'wb') as handle:
pickle.dump({"users": unique_users, "items": unique_items}, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
Next, we split the data into a train and test set, the last movie each user has recently seen will be used for the test set.
```
# Need to sort before popping to get last item
df.sort_values(by='timestamp', inplace=True)
# clean up data
del df['rating'], df['timestamp']
df = df.drop_duplicates() # assuming it keeps order
# now we have filtered and sorted by time data, we can split test data out
grouped_sorted = df.groupby(USER_COLUMN, group_keys=False)
test_data = grouped_sorted.tail(1).sort_values(by=USER_COLUMN)
# need to pop for each group
train_data = grouped_sorted.apply(lambda x: x.iloc[:-1])
train_data['target']=1
test_data['target']=1
train_data.head()
```
Next, we generate the negative samples for training
```
sampler = _TestNegSampler(df.values, nb_users, nb_items, 500) # using 500 negative samples
train_negs = sampler.generate()
train_negs = train_negs.reshape(-1, 500)
sampler = _TestNegSampler(df.values, nb_users, nb_items, 100) # using 100 negative samples
test_negs = sampler.generate()
test_negs = test_negs.reshape(-1, 100)
import numpy as np
# generating negative samples for training
train_data_neg = np.zeros((train_negs.shape[0]*train_negs.shape[1],3), dtype=int)
idx = 0
for i in tqdm.tqdm(range(train_negs.shape[0])):
for j in range(train_negs.shape[1]):
train_data_neg[idx, 0] = i # user ID
train_data_neg[idx, 1] = train_negs[i, j] # negative item ID
idx += 1
# generating negative samples for testing
test_data_neg = np.zeros((test_negs.shape[0]*test_negs.shape[1],3), dtype=int)
idx = 0
for i in tqdm.tqdm(range(test_negs.shape[0])):
for j in range(test_negs.shape[1]):
test_data_neg[idx, 0] = i
test_data_neg[idx, 1] = test_negs[i, j]
idx += 1
train_data_np= np.concatenate([train_data_neg, train_data.values])
np.random.shuffle(train_data_np)
test_data_np= np.concatenate([test_data_neg, test_data.values])
np.random.shuffle(test_data_np)
# HugeCTR expect user ID and item ID to be different, so we use 0 -> nb_users for user IDs and
# nb_users -> nb_users+nb_items for item IDs.
train_data_np[:,1] += nb_users
test_data_np[:,1] += nb_users
np.max(train_data_np[:,1])
```
### Write HugeCTR data files
Next, we will write the data to disk using HugeCTR norm format.
```
from ctypes import c_longlong as ll
from ctypes import c_uint
from ctypes import c_float
from ctypes import c_int
def write_hugeCTR_data(huge_ctr_data, filename='huge_ctr_data.dat'):
print("Writing %d samples"%huge_ctr_data.shape[0])
with open(filename, 'wb') as f:
#write header
f.write(ll(0)) # 0: no error check; 1: check_num
f.write(ll(huge_ctr_data.shape[0])) # the number of samples in this data file
f.write(ll(1)) # dimension of label
f.write(ll(1)) # dimension of dense feature
f.write(ll(2)) # long long slot_num
for _ in range(3): f.write(ll(0)) # reserved for future use
for i in tqdm.tqdm(range(huge_ctr_data.shape[0])):
f.write(c_float(huge_ctr_data[i,2])) # float label[label_dim];
f.write(c_float(0)) # dummy dense feature
f.write(c_int(1)) # slot 1 nnz: user ID
f.write(c_uint(huge_ctr_data[i,0]))
f.write(c_int(1)) # slot 2 nnz: item ID
f.write(c_uint(huge_ctr_data[i,1]))
```
#### Train data
```
!rm -rf ./data/hugeCTR
!mkdir ./data/hugeCTR
for i, data_arr in enumerate(np.array_split(train_data_np,10)):
write_hugeCTR_data(data_arr, filename='./data/hugeCTR/huge_ctr_data_%d.dat'%i)
with open('./data/hugeCTR/filelist.txt', 'wt') as f:
f.write('10\n');
for i in range(10):
f.write('./data/hugeCTR/huge_ctr_data_%d.dat\n'%i)
```
#### Test data
```
for i, data_arr in enumerate(np.array_split(test_data_np,10)):
write_hugeCTR_data(data_arr, filename='./data/hugeCTR/test_huge_ctr_data_%d.dat'%i)
with open('./data/hugeCTR/test_filelist.txt', 'wt') as f:
f.write('10\n');
for i in range(10):
f.write('./data/hugeCTR/test_huge_ctr_data_%d.dat\n'%i)
```
<a id="3"></a>
## 3. HugeCTR DLRM training
In this section, we will train a DLRM network on the augmented movie lens data. First, we write the model config file.
```
%%writefile dlrm_config.json
{
"solver": {
"lr_policy": "fixed",
"display": 1000,
"max_iter":50000,
"gpu": [0],
"batchsize": 65536,
"snapshot": 3000,
"snapshot_prefix": "./hugeCTR_saved_model_DLRM/",
"eval_interval": 3000,
"eval_batches": 1000,
"mixed_precision": 1024,
"eval_metrics": ["AUC:1.0"]
},
"optimizer": {
"type": "SGD",
"global_update": false,
"sgd_hparam": {
"learning_rate": 0.1,
"warmup_steps": 1000,
"decay_start": 10000,
"decay_steps": 40000,
"end_lr": 1e-5
}
},
"layers": [
{
"name": "data",
"type": "Data",
"slot_size_array": [138493 , 26744],
"slot_size_array_orig": [138493 , 26744],
"source": "./data/hugeCTR/filelist.txt",
"eval_source": "./data/hugeCTR/test_filelist.txt",
"check": "None",
"cache_eval_data": true,
"label": {
"top": "label",
"label_dim": 1
},
"dense": {
"top": "dense",
"dense_dim": 1
},
"sparse": [
{
"top": "data1",
"type": "LocalizedSlot",
"max_feature_num_per_sample": 2,
"max_nnz": 1,
"slot_num": 2
}
]
},
{
"name": "sparse_embedding1",
"type": "LocalizedSlotSparseEmbeddingHash",
"bottom": "data1",
"top": "sparse_embedding1",
"sparse_embedding_hparam": {
"slot_size_array": [138493 , 26744],
"embedding_vec_size": 64,
"combiner": 0
}
},
{
"name": "fc1",
"type": "FusedInnerProduct",
"bottom": "dense",
"top": "fc1",
"fc_param": {
"num_output": 64
}
},
{
"name": "fc2",
"type": "FusedInnerProduct",
"bottom": "fc1",
"top": "fc2",
"fc_param": {
"num_output": 128
}
},
{
"name": "fc3",
"type": "FusedInnerProduct",
"bottom": "fc2",
"top": "fc3",
"fc_param": {
"num_output": 64
}
},
{
"name": "interaction1",
"type": "Interaction",
"bottom": ["fc3", "sparse_embedding1"],
"top": "interaction1"
},
{
"name": "fc4",
"type": "FusedInnerProduct",
"bottom": "interaction1",
"top": "fc4",
"fc_param": {
"num_output": 1024
}
},
{
"name": "fc5",
"type": "FusedInnerProduct",
"bottom": "fc4",
"top": "fc5",
"fc_param": {
"num_output": 1024
}
},
{
"name": "fc6",
"type": "FusedInnerProduct",
"bottom": "fc5",
"top": "fc6",
"fc_param": {
"num_output": 512
}
},
{
"name": "fc7",
"type": "FusedInnerProduct",
"bottom": "fc6",
"top": "fc7",
"fc_param": {
"num_output": 256
}
},
{
"name": "fc8",
"type": "InnerProduct",
"bottom": "fc7",
"top": "fc8",
"fc_param": {
"num_output": 1
}
},
{
"name": "loss",
"type": "BinaryCrossEntropyLoss",
"bottom": ["fc8","label"],
"top": "loss"
}
]
}
!rm -rf ./hugeCTR_saved_model_DLRM/
!mkdir ./hugeCTR_saved_model_DLRM/
!CUDA_VISIBLE_DEVICES=0 ../build/bin/huge_ctr --train ./dlrm_config.json
```
<a id="4"></a>
## 4. Answer item similarity with DLRM embedding
In this section, we demonstrate how the output of HugeCTR training can be used to carry out simple inference tasks. Specifically, we will show that the movie embeddings can be used for simple item-to-item similarity queries. Such a simple inference can be used as an efficient candidate generator to generate a small set of cadidates prior to deep learning model re-ranking.
First, we read the embedding tables and extract the movie embeddings.
```
import struct
import pickle
import numpy as np
key_type = 'I32' # {'I64', 'I32'}, default is 'I32'
key_type_map = {"I32": ["I", 4], "I64": ["q", 8]}
embedding_vec_size = 64
HUGE_CTR_VERSION = 2.21 # set HugeCTR version here, 2.2 for v2.2, 2.21 for v2.21
if HUGE_CTR_VERSION <= 2.2:
each_key_size = key_type_map[key_type][1] + key_type_map[key_type][1] + 4 * embedding_vec_size
else:
each_key_size = key_type_map[key_type][1] + 8 + 4 * embedding_vec_size
embedding_table = [{},{}]
with open('./hugeCTR_saved_model_DLRM/0_sparse_9000.model', 'rb') as file:
try:
while True:
buffer = file.read(each_key_size)
if len(buffer) == 0:
break
if HUGE_CTR_VERSION <= 2.2:
key, slot_id = struct.unpack("2" + key_type_map[key_type][0],
buffer[0: 2*key_type_map[key_type][1]])
values = struct.unpack(str(embedding_vec_size) + "f", buffer[2*key_type_map[key_type][1]: ])
else:
key = struct.unpack(key_type_map[key_type][0], buffer[0 : key_type_map[key_type][1]])[0]
slot_id = struct.unpack("Q", buffer[key_type_map[key_type][1] : key_type_map[key_type][1] + 8])[0]
values = struct.unpack(str(embedding_vec_size) + "f", buffer[key_type_map[key_type][1] + 8: ])
if slot_id==0:
embedding_table[slot_id][key] = values
elif slot_id==1:
embedding_table[slot_id][key - 138493] = values
else:
raise(Exception("Slot ID not found - %d"%slot_id))
except BaseException as error:
print(error)
item_embedding = np.zeros((26744, embedding_vec_size), dtype='float')
for i in range(len(embedding_table[1])):
item_embedding[i] = embedding_table[1][i]
len(embedding_table[1])
```
### Answer nearest neighbor queries
```
from scipy.spatial.distance import cdist
def find_similar_movies(nn_movie_id, item_embedding, k=10, metric="euclidean"):
#find the top K similar items according to one of the distance metric: cosine or euclidean
sim = 1-cdist(item_embedding, item_embedding[nn_movie_id].reshape(1, -1), metric=metric)
return sim.squeeze().argsort()[-k:][::-1]
with open('./mappings.pickle', 'rb') as handle:
movies_mapping = pickle.load(handle)["items"]
nn_to_movies = movies_mapping
movies_to_nn = {}
for i in range(len(movies_mapping)):
movies_to_nn[movies_mapping[i]] = i
import pandas as pd
movies = pd.read_csv("./data/ml-20m/movies.csv", index_col="movieId")
for movie_ID in range(1,1000):
try:
print("Query: ", movies.loc[movie_ID]["title"], movies.loc[movie_ID]["genres"])
print("Similar movies: ")
similar_movies = find_similar_movies(movies_to_nn[movie_ID], item_embedding)
for i in similar_movies:
print(nn_to_movies[i], movies.loc[nn_to_movies[i]]["title"], movies.loc[nn_to_movies[i]]["genres"])
print("=================================\n")
except Exception as e:
pass
```
| github_jupyter |
```
# from google.colab import drive
# drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 250
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=12, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=12, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
for i in range(9):
x[:,i] = self.helper(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
return x, y
def helper(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=6, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=6, out_channels=6, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=12, out_channels=20, kernel_size=3, padding=0)
self.fc1 = nn.Linear(1014, 512)
self.fc2 = nn.Linear(512, 64)
# self.fc3 = nn.Linear(512, 64)
# self.fc4 = nn.Linear(64, 10)
self.fc3 = nn.Linear(64,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = (F.relu(self.conv2(x)))
# print(x.shape)
# x = (F.relu(self.conv3(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
# x = F.relu(self.fc3(x))
# x = F.relu(self.fc4(x))
x = self.fc3(x)
return x
classify = Classification().double()
classify = classify.to("cuda")
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.001)#, betas=(0.9, 0.999), eps=1e-08, weight_decay=0, amsgrad=False)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 200
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.005):
break;
if epoch % 5 == 0:
# focus_net.eval()
# classify.eval()
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
# torch.save(focus_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_focus_net.pt")
# torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn_3layers/"+name+"_classify.pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.savefig("train_ftpt.pdf", bbox_inches='tight')
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.savefig("test_ftpt.pdf", bbox_inches='tight')
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
max_alpha =[]
alpha_ftpt=[]
argmax_more_than_half=0
argmax_less_than_half=0
for i, data in enumerate(test_loader):
inputs, labels,fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"),labels.to("cuda")
alphas, avg = focus_net(inputs)
outputs = classify(avg)
mx,_ = torch.max(alphas,1)
max_alpha.append(mx.cpu().detach().numpy())
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if (focus == fore_idx[j] and predicted[j] == labels[j]):
alpha_ftpt.append(alphas[j][focus].item())
max_alpha = np.concatenate(max_alpha,axis=0)
print(max_alpha.shape)
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(max_alpha,bins=50,color ="c")
plt.title("alpha values histogram")
plt.savefig("alpha_hist.pdf")
plt.figure(figsize=(6,6))
_,bins,_ = plt.hist(np.array(alpha_ftpt),bins=50,color ="c")
plt.title("alpha values in ftpt")
plt.savefig("alpha_hist_ftpt.pdf")
```
| github_jupyter |
```
import numpy as np
import tensorflow as tf
from tensorflow import keras
import pandas as pd
import seaborn as sns
from pylab import rcParams
import matplotlib.pyplot as plt
from matplotlib import rc
from sklearn.model_selection import train_test_split
from pandas.plotting import register_matplotlib_converters
from sklearn.metrics import confusion_matrix
import itertools
import matplotlib
import numpy as np
import matplotlib.pyplot as plt
import numpy as np
%matplotlib inline
%matplotlib inline
%config InlineBackend.figure_format='retina'
register_matplotlib_converters()
sns.set(style='whitegrid', palette='muted', font_scale=1.5)
rcParams['figure.figsize'] = 22, 10
RANDOM_SEED = 42
np.random.seed(RANDOM_SEED)
tf.random.set_seed(RANDOM_SEED)
df = pd.read_csv(
"dateindex1.csv",
parse_dates=['datetime'],
index_col="datetime"
)
df['hour'] = df.index.hour
df['day_of_month'] = df.index.day
df['day_of_week'] = df.index.dayofweek
df['month'] = df.index.month
df['year'] = df.index.year
df.head()
y = df[['G']]
X= df.drop(columns=['G'])
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=0)
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
history=regressor.fit(X_train, y_train)
y_pred = regressor.predict(X_test)
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
plt.plot(np.arange(len(y_train), len(y_train) + len(y_test)), y_test, marker='.', label="true")
plt.plot(np.arange(len(y_train), len(y_train) + len(y_test)), y_pred, 'r', label="prediction")
plt.ylabel('Irradiance')
plt.xlabel('Time Step')
plt.legend()
plt.show();
import pickle
with open('final_decision_Tree_model','wb') as f:
pickle.dump(regressor,f)
test_data = np.array([ 2, 1, 5 ,1 ,2005])
print(regressor.predict(test_data.reshape(1,5)))
X_test.head()
y_test.head()
from sklearn.model_selection import train_test_split
x_train, x_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
regressor.fit(x_train, y_train)
regressor.score( X, y)
from sklearn.metrics import confusion_matrix
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
cnf_matrix = confusion_matrix(y_test, y_pred)
plt.figure()
plot_confusion_matrix(cnf_matrix, classes=['Type_A', 'Type_B'], normalize=False,
title='Non Normalized confusion matrix')
```
| github_jupyter |
# Convolutional Neural Net (CNN) Fraud Classifier
Another Neural Net Architecture that can handle series data are __Convolutional Neural Nets (CNN)__. They are mainly used in image recognition and *vision* problems in general. But might have some use in Fraud detection.
With CNN's the network uses convolutional filters, it slides these filters over the data along an axis. For instance in our Transactional data case, the filter would slide along the time-axis. A filter sees a number of elements (here we see two), depending on the size of the filter. This is often named the __kernel size__. The number of elements it moves is named the __stride__, in this case the stride is one. We move the filter/kernel one position at a time.
For time-series we will use a 1d convolution, the kernel_size is a single number and the 'depth' of the filter is the full 'depth/height' of the series, i.e. it spans the full set of features.
At each position the filter outputs one an only one number; the sum of the element wise multiplication of the filter and the portion of the input it spans. $o= \sum_{i=1}^{ks} \sum_{j=1}^{cc} x_{ij} f_{ij}$, where $ks=kernel size$ and $cc=input channel count$ (i.e. features) or $o = \sum x \bigotimes \ f$.
Therefore having one filter slide over the input gives us and array, a 1D - Tensor. The lenght of that tensor is shorter or equal the series length. $Length = \frac {inputlength - kernelsize + 2 * padding} {stride} + 1$.
Intuitively we can think of this as an array that shows the 'reaction' of the filter of a certain input area.

Now, CNN's don't just run one filter over the data. They use multiple filters, the exact amount is defined as '#Filters' or '#Channels'.
Each filter has an output as described above, each of these arrays has the same length and can be concatenated along the second axis. In the example below five filters then give a seconds dimension size of five.
That gives us a 2-D Tensor where the __width__ depends on the formula specified above and the __depth/height__ depends on the number of features/channels specified in the layer. So we end-up with a new block of data that potentially has a different size. Intuitively we can think of that block as showing how *each* filter reacted to a specified input area. Note that the filters are lined up over a 'reduced' time axis, but the aspect/sequence of time is maintained. All elements along the second axis correspond to the same time portion.

In order to help build some intuition of how these filters/kernels could help detect Fraud, we'll include a small *purely hypothetical* example.
If we think back to the one-hot encoded vectors showing the 'bin' of the amount. And let's say we have split our amounts in 3 bins, then a small amount might be represented as;
$S_{amount} = \begin{bmatrix}0 \\ 0 \\ 1 \end{bmatrix}$
I.e. there is a 1 in the low range, having a 1 in the high range would mean we have a big amount.
$B_{amount} = \begin{bmatrix}1 \\ 0 \\ 0 \end{bmatrix}$
If we put these in a series, where the columns are the time, then we have a small amount followed by a big amount
$I_{sb} = \begin{bmatrix}0 & 1 \\ 0 & 0 \\ 1 & 0 \end{bmatrix}$.
Then if we were to have a filter that looks like so;
$F= \begin{bmatrix}-1 & 1 \\ 0.3 & 0 \\ 1 & 0 \end{bmatrix}$
Our input $I_{sb}$ filtered by $F$. Would give __2__. $(0 * -1) + (1 * 1) + (0 * 0.3) + (0 * 0) + (1 * 1) + (0 * 0)$. A fairly strong reaction.
Reversing the transactions (big followed by small) $I_{bs} = \begin{bmatrix}1 & 0 \\ 0 & 0 \\ 0 & 1 \end{bmatrix}$. Would give a product of __-1__. A negative reation.
Two big transcations $I_{bb} = \begin{bmatrix}1 & 1 \\ 0 & 0 \\ 0 & 0 \end{bmatrix}$. Would give a product of __0__. No reaction whatsoever.
We can see that these filters react differently to the sequences and thus give the Neural Net the ability to 'detect' patterns in the series. Maybe important to note is the our filter is time-invariant, it can find small payments followed by big ones anywhere in the sequence, whether it's the 9th and 7th or the 5th and 4th... the reaction will be the same.
Clearly it would not be doing this using just the amount, but all of the input features.
Convolutional Nets can __optimize__ the content of the filters to optimally reduce the loss and solve a specific problem. We can see that they theoretically have the ability to come up with a set of filter(s) that can detect certain patterns in the transactional data. The output of the convolutions can then for instance be used to detect various patterns being present in the data.
---
#### Note on the data set
The data set used here is not particularly complex and/or big. It's not really all that challenging to find the fraud. In an ideal world we'd be using more complex data sets to show the real power of Deep Learning. There are a bunch of PCA'ed data sets available, but the PCA obfuscates some of the elements that are useful.
*These examples are meant to show the possibilities, it's not so useful to interpret their performance on this data set*
## Imports
```
import torch
import numpy as np
import pandas as pd
import datetime as dt
import gc
import d373c7.features as ft
import d373c7.engines as en
import d373c7.pytorch as pt
import d373c7.pytorch.models as pm
import d373c7.plot as pl
```
## Set a random seed for Numpy and Torch
> Will make sure we always sample in the same way. Makes it easier to compare results. At some point it should been removed to test the model stability.
```
# Numpy
np.random.seed(42)
# Torch
torch.manual_seed(42)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
```
## Define base feature and read the File
The base features are features found in the input file. They need to be defined after which the file can be read using the `EnginePandasNumpy`. Using the `from_csv` method.
```
# Change this to read from another location
file = '../../../../data/bs140513_032310.csv'
step = ft.FeatureSource('step', ft.FEATURE_TYPE_INT_16)
customer = ft.FeatureSource('customer', ft.FEATURE_TYPE_STRING)
age = ft.FeatureSource('age', ft.FEATURE_TYPE_CATEGORICAL)
gender = ft.FeatureSource('gender', ft.FEATURE_TYPE_CATEGORICAL)
merchant = ft.FeatureSource('merchant', ft.FEATURE_TYPE_CATEGORICAL)
category = ft.FeatureSource('category', ft.FEATURE_TYPE_CATEGORICAL)
amount = ft.FeatureSource('amount', ft.FEATURE_TYPE_FLOAT)
fraud = ft.FeatureSource('fraud', ft.FEATURE_TYPE_INT_8)
base_features = ft.TensorDefinition(
'base',
[
step,
customer,
age,
gender,
merchant,
category,
amount,
fraud
])
amount_binned = ft.FeatureBin('amount_bin', ft.FEATURE_TYPE_INT_16, amount, 30)
def step_to_date(step_count: int):
return dt.datetime(2020, 1, 1) + dt.timedelta(days=int(step_count))
date_time = ft.FeatureExpression('date', ft.FEATURE_TYPE_DATE_TIME, step_to_date, [step])
intermediate_features = ft.TensorDefinition(
'intermediate',
[
customer,
step,
age,
gender,
merchant,
category,
amount_binned,
fraud
])
amount_oh = ft.FeatureOneHot('amount_one_hot', amount_binned)
age_oh = ft.FeatureOneHot('age_one_hot', age)
gender_oh = ft.FeatureOneHot('gender_one_hot', gender)
merchant_oh = ft.FeatureOneHot('merchant_one_hot', merchant)
category_oh = ft.FeatureOneHot('category_one_hot', category)
fraud_label = ft.FeatureLabelBinary('fraud_label', fraud)
learning_features = ft.TensorDefinition(
'learning',
[
customer,
date_time,
age_oh,
gender_oh,
merchant_oh,
category_oh,
amount_oh
])
label = ft.TensorDefinition('label', [fraud_label])
model_features = ft.TensorDefinitionMulti([learning_features, label])
with en.EnginePandasNumpy() as e:
df = e.from_csv(base_features, file, inference=False)
df = e.from_df(intermediate_features, df, inference=False)
ft_df = e.from_df(learning_features, df, inference=False)
lb_df = e.from_df(label, df, inference=False)
lb_np = e.to_numpy_list(label, lb_df)
ser_np = e.to_series_stacked(ft_df, learning_features, key_field=customer, time_field=date_time, length=5)
data_list = en.NumpyList(ser_np.lists + lb_np.lists)
print(data_list.shapes)
print(data_list.dtype_names)
```
## Wrangle the data
Time to split the data. For time series data it is very important to keep the order of the data. Below split will start from the end and work it's way to the front of the data. Doing so the training, validation and test data are nicely colocated in time. You almost *never* want to plain shuffle time based data.
> 1. Split out a test-set of size `test_records`. This is used for model testing.
> 2. Split out a validation-set of size `validation_records`. It will be used to monitor overfitting during training
> 3. All the rest is considered training data.
For time-series we'll perform an additional action.
> 1. The series at the beginning of the data set will all be more or less empty as there is no history, that is not so useful during training, ideally we have records with history and complete series, sometimes named 'mature' series. We'll throw away the first couple of entries.
__Important__; please make sure the data is ordered in ascending fashion on a date(time) field. The split function does not order the data, it assumes the data is in the correct order.
```
test_records = 100000
val_records = 30000
maturation = 30000
train_data, val_data, test_data = data_list.split_time(val_records, test_records)
train_data = train_data[maturation:]
print(f'Training Data shapes {train_data.shapes}')
print(f'Validation Data shapes {val_data.shapes}')
print(f'Test Data shapes {test_data.shapes}')
del data_list, df, ft_df, lb_df, lb_np, ser_np
gc.collect()
print('Done')
```
## Set-up Devices
```
device, cpu = pt.init_devices()
```
## Build the model
We give it the TensorDefinition of Learning features, then ask it to build a __Convolutioanl NN__ having 2 conv layers, the first has 12 output_channels/features, a *kernel_size* of 2 and *stride* of 1, the second layer has 24 output_channels/features, a *kernel_size* of 2 and *stride* of 1. And we ask the model to add 1 *linear* layer of size 8 after that.
```
# Setup Pytorch Datasets for the training and validation
batch_size = 128
train_ds = pt.NumpyListDataSetMulti(model_features, train_data)
val_ds = pt.NumpyListDataSetMulti(model_features, val_data)
train_sampler = pt.ClassSamplerMulti(model_features, train_data).over_sampler()
# Wrap them in a Pytorch Dataloader
train_dl = train_ds.data_loader(cpu, batch_size, num_workers=2, sampler=train_sampler)
val_dl = val_ds.data_loader(cpu, batch_size, num_workers=2)
# Create a Model
m = pm.GeneratedClassifier(model_features, convolutional_layers=[(12, 2, 1), (24, 2, 1)], linear_layers=[8])
print(m)
```
Grapically that model looks like below;
Overall this is very similar to first. the Recurrent example. But here we have a `Convolutional` layer instead of a recurrent layer.
The convolutional layer will reduce the output in 2 layers. First it will be reduced to a (12,4) 2-D tensor. (12 for the output channels and 4 as a result the the formula in the intro to calculate the new length). The second later will further reduce that to a __(24,3) 2-D tensor__. (12 for the output channels and 3 as a result the the formula in the intro to calculate the new length).
the (24,3) 2-D tensor is __flattened__ to a (72) 1-D Tensor, we add the entire last payment to that so end-up with a (179) 1-D Tensor, which follow the path down to linear layers etc...
*(Some layers have been omitted for simplicity)*

# Start Training
### First find a decent Learning Rate.
> Create a trainer and run the find_lr function and plot. This functions iterates over the batches gradually increasing the learning rate from a minimum to a maximum learning rate.
```
t = pt.Trainer(m, device, train_dl, val_dl)
r = t.find_lr(1e-4, 1e-1, 200)
pl.TrainPlot().plot_lr(r)
```
## Start Training and plot the results
> We train for __10 epochs__ and __learning rate 5e-3__. That means we run over the total training data set a total of 10 times/epochs where the model learns, after each epoch we use the trained model and perform a test run on the validation set.
```
t = pt.Trainer(m, device, train_dl, val_dl)
h = t.train_one_cycle(10, 5e-3)
pl.TrainPlot().plot_history(h, fig_size=(10,10))
```
## Test the model on the test data
> Test the model on the test set, it is data that was not seen during training and allows us to validate model results.
```
test_ds = pt.NumpyListDataSetMulti(model_features, test_data)
test_dl = test_ds.data_loader(cpu, 128, num_workers=2)
ts = pt.Tester(m, device, test_dl)
pr = ts.test_plot()
tp = pl.TestPlot()
tp.print_classification_report(pr)
tp.plot_confusion_matrix(pr, fig_size=(6,6))
tp.plot_roc_curve(pr, fig_size=(6,6))
tp.plot_precision_recall_curve(pr, fig_size=(6,6))
```
| github_jupyter |
## ALEJANDRO SANTORUM VARELA - Ejercicios 16-10-2017(Condiciones necesarias y suficientes)
EJERCICIO 1 - Encontrar la condición necesaria y suficiente sobre el entero $n$ para que $1+2+3+...+n$ divida exactamente a $n!$. Una vez encontrada hay que intentar demostrar la equivalencia.
El siguiente programa nos ayudará a visualizar "por donde van los tiros".
```
for k in srange(1,51):
suma = 0
for i in srange(1,k+1):
suma = suma + i
kk = factorial(k)
if kk%suma == 0:
print(k)
```
No es muy difícil ver que para valores de n impares todos cumplen la condición (condición suficiente), pero aún no es necesaria, ya que algunos valores pares lo cumplen también.
Por otro lado, es conocido que $1+2+3+...+n$ es igual a $\frac{n*(n+1)}{2}$ con lo que podemos trabajar más fácilmente.
Si dividimos $n!$ entre $\frac{n*(n+1)}{2}$ y haciendo las simplificaciones oportunas obtenemos: $$\frac{2*(n-1)!}{n+1}$$
Una vez aquí, podemos afirmar que si $n$ es impar, el denominador es par, por lo que se puede simplificar con el dos del numerador y lo restante en el denominador sería menor estricto que $(n-1)!$, por lo que este último sería exactamente divisible por el denominador ya que es uno de sus factores.
No obstante, las cosas no iban a ser tan fáciles. Si $n$ es par, el denominador es impar y ya no se puede simplificar con el 2 del numerador. La única opción que tenemos para que esta división sea exacta es que el denominador ($n+1$) se pueda descomponer en factores de menor valor, los cuales se encontrarían entre los factores de $(n-1)!$ y la división sería exacta. Sin embargo, si $n+1$ no es compuesto, es decir, $n+1$ es un número primo, esta división no es exacta por lo que no se cumpliría la condición.
Para finalizar, en conclusión, la condición necesaria y suficiente sobre $n$ es que este sea un número impar o un número par que no preceda a un primo. Esto se demuestra con todo lo explicado anteriormente.
.
.
EJERCICIO 4 - Demuestra que para $n > 1$ el entero $n^4 + 4$ es siempre compuesto.
Veamos el caso $n=2$: $2^4 +4 = 20 = 2^2 * 5$ compuesto.
Ahora bien, como $n^4 + 4 > 2$ para todo $n>1$, para que $n^4 + 4$ NO sea compuesto, tiene que ser un número impar(hablamos de $n^4 + 4$ no del propio $n$).
-Veamos si $n$ es par:
$n^4 + 4$ ----> Es conocido que todo número par elevado a cualquier potencia es par y que todo número par sumado con otro par da un número par. Con esto podemos afirmar que si $n$ es par, $n^4 + 4$ es par y por lo tanto compuesto, ya que se puede descomponer como mínimo en 2 por otro número.
-Caso en que $n$ es impar: utilizamos inducción y suponemos que $n^4 + 4$ es compuesto (hipótesis de inducción) y lo probamos para el caso n+1:
```
f(n) = (n+1)^4 + 4
f.expand()
```
Entonces $(n+1)^4 + 4 = n^4 + 4 + 4n^3 + 6n^2 + 4n + 1$.
$n^4 + 4$ es un número compuesto por la hipótesis de inducción, por lo que un numero impar.
$4n^3$ es un número par, ya que está multiplicado por 4.
$6n^2$ es un número par, ya que está multiplicado por 6.
$4n$ es un número par, ya que está multiplicado por 4.
5 es trivialmente impar.
Sabiendo que la suma de pares es par, la suma de impares es par, y que la suma de impar+par es impar, podemos agrupar lo anteior.
impar+par+par+par+impar = par
Por lo que $(n+1)^4 + 4$ es par, o lo que es lo mismo, un número compuesto.
Queda demostrado.
.
.
EJERCICIO 3 - La cifra dominante de un entero es la primera por la izquierda que no es nula. Encontrar los dígitos que pueden ser la cifra dominante al mismo tiempo de $2^n$ y $5^n$, con el mismo exponente $n$. Una vez encontrados, hay que intentar demostrar que esos dígitos son los únicos posibles, pero en este caso, no me han sido de ninguna ayuda las pruebas que he hecho con el ordenador.
En primer lugar, vamos a ayudarnos de Sagemath para poder averiguar un posible patrón entre el exponente $n$ y el resultado.
```
for i in srange(1,101):
print("Exponente:")
print(i)
print("Potencia de 2:")
print(2^i)
print("Potencia de 5:")
print(5^i)
print("")
```
Se puede ver que la cifra dominante solo coincide con $n = 5, n = 15, n = 88, n=98$ en los 100 primeros valores de $n$ y en todos ellos el valor dominante es 3. Por lo tanto, nuestra atención se centrará ahora en demostrar que es la única cifra en que pasa esto.
Desafortunadamente, todos mis intentos de demostrar esta propiedad no han tenido éxito. Mantengo que 3 es la única cifra en que pasa esto, pero quedará en conjetura y no en hecho demostrado.
.
.
EJERCICIO 2 - Encontrar la condición necesaria y suficiente para que existan infinitos múltiplos de un entero $n$ que se escriban únicamente con unos en base 10. Una vez encontrada hay que intentar demostrar la equivalencia. ¿Ocurrirá algo similar si queremos que infinitos múltiplos se escriban usando únicamente otro dígito (2, 3, etc.).
Como hecho en los ejercicios anteriores, vamos a ayudarnos de la programación para hacer unos cálculos rápidos y poder conseguir algunas pistas:
```
print(factor(11))
print(factor(111))
print(factor(1111))
print(factor(11111))
print(factor(111111))
print(factor(1111111))
print(factor(11111111))
print(factor(111111111))
print(factor(1111111111))
print(factor(11111111111))
print(factor(111111111111))
print(factor(1111111111111))
print(factor(11111111111111))
print(factor(111111111111111))
print(factor(1111111111111111))
```
Personalmente esto no me dice mucho. La verdad es que no se por donde atacar el problema más.
Quizá lo máximo que puede decir sobre $n$ es que tiene que ser un entero no múltiplo de 2 ni de 5, ya que cualquier número acabado en 1 no puede ser divisible por 2 ni por 5. Pero mantengo que es una información pobre, lejos de demostrar nada.
| github_jupyter |
```
import os
from zenml.repo import Repository
from zenml.datasources import CSVDatasource
from zenml.pipelines import TrainingPipeline
from zenml.steps.evaluator import TFMAEvaluator
from zenml.steps.preprocesser import StandardPreprocesser
from zenml.steps.split import RandomSplit
from zenml.steps.trainer import TFFeedForwardTrainer
from zenml.repo import Repository, ArtifactStore
from zenml.utils.naming_utils import transformed_label_name
from zenml.steps.deployer import GCAIPDeployer
from zenml.steps.deployer import CortexDeployer
from examples.cortex.predictor.tf import TensorFlowPredictor
from zenml.backends.orchestrator import OrchestratorGCPBackend
from zenml.metadata import MySQLMetadataStore
from zenml.backends.processing import ProcessingDataFlowBackend
from zenml.backends.training import SingleGPUTrainingGCAIPBackend
from zenml.backends.processing import ProcessingDataFlowBackend
```
We are going to be creating a ZenML training pipeline and showcasing the modularity of ZenML backends in this example. On a high level, here is what a ZenML training pipeline looks like:
<img src="graphics/architecture.png" width="600" height="600" />
# Set up some variables
```
GCP_BUCKET=os.getenv('GCP_BUCKET')
GCP_PROJECT=os.getenv('GCP_PROJECT')
GCP_REGION=os.getenv('GCP_REGION')
GCP_CLOUD_SQL_INSTANCE_NAME=os.getenv('GCP_CLOUD_SQL_INSTANCE_NAME')
MODEL_NAME=os.getenv('MODEL_NAME')
CORTEX_ENV=os.getenv('CORTEX_ENV')
MYSQL_DB=os.getenv('MYSQL_DB')
MYSQL_USER=os.getenv('MYSQL_USER')
MYSQL_PWD=os.getenv('MYSQL_PWD')
MYSQL_PORT=os.getenv('MYSQL_PORT')
MYSQL_HOST=os.getenv('MYSQL_HOST')
CONNECTION_NAME = f'{GCP_PROJECT}:{GCP_REGION}:{GCP_CLOUD_SQL_INSTANCE_NAME}'
TRAINING_JOB_DIR = os.path.join(GCP_BUCKET, 'gcp_gcaip_training/staging')
repo: Repository = Repository.get_instance()
# Define artifact store in the cloud
cloud_artifact_store = ArtifactStore(os.path.join(GCP_BUCKET, 'all_feature_demo'))
# Define metadata store in the cloud
cloud_metadata_store = MySQLMetadataStore(
host=MYSQL_HOST,
port=int(MYSQL_PORT),
database=MYSQL_DB,
username=MYSQL_USER,
password=MYSQL_PWD,
)
```
# Create first pipeline
```
training_pipeline = TrainingPipeline(name='Experiment 1')
```
#### Add a datasource. This will automatically track and version it.
```
try:
ds = CSVDatasource(name='Pima Indians Diabetes', path='gs://zenml_quickstart/diabetes.csv')
except:
repo: Repository = Repository.get_instance()
ds = repo.get_datasource_by_name('Pima Indians Diabetes')
training_pipeline.add_datasource(ds)
```
#### Add a split step to partition data into train and eval
```
training_pipeline.add_split(RandomSplit(split_map={'train': 0.7, 'eval': 0.2, 'test':0.1}))
```
#### Add a preprocessing step to transform data to be ML-capable
```
training_pipeline.add_preprocesser(
StandardPreprocesser(
features=['times_pregnant', 'pgc', 'dbp', 'tst', 'insulin', 'bmi',
'pedigree', 'age'],
labels=['has_diabetes'],
overwrite={'has_diabetes': {
'transform': [{'method': 'no_transform', 'parameters': {}}]}}
))
```
#### Add a trainer which defines model and training
```
training_pipeline.add_trainer(TFFeedForwardTrainer(
loss='binary_crossentropy',
last_activation='sigmoid',
output_units=1,
metrics=['accuracy'],
epochs=5))
```
#### Add an evaluator to calculate slicing metrics
```
training_pipeline.add_evaluator(
TFMAEvaluator(slices=[['has_diabetes']],
metrics={transformed_label_name('has_diabetes'):
['binary_crossentropy', 'binary_accuracy']}))
```
#### Run and evaluate
```
training_pipeline.run()
training_pipeline.view_statistics(magic=True)
training_pipeline.evaluate(magic=True)
```
#### Inspect datasource
```
datasources = repo.get_datasources()
datasource = datasources[0]
print(datasource)
df = datasource.sample_data()
df.head()
df.shape
df.columns
```
## Skip preprocessing with your next (warm-starting) pipeline
#### Clone first experiment and only change one hyper-parameter
```
training_pipeline_2 = training_pipeline.copy('Experiment 2')
training_pipeline_2.add_trainer(TFFeedForwardTrainer(
loss='binary_crossentropy',
last_activation='sigmoid',
output_units=1,
metrics=['accuracy'],
epochs=20))
training_pipeline_2.run()
training_pipeline_2.evaluate(magic=True)
```
## Post-training
#### Verify theres still only one datasource
```
datasources = repo.get_datasources()
print(f"We have {len(datasources)} datasources")
```
#### Compare pipelines
```
repo.compare_training_runs()
```
# Distribute splitting/preprocessing easily
```
training_pipeline_3 = repo.get_pipeline_by_name('Experiment 1').copy('Experiment 3')
# Define the processing backend
processing_backend = ProcessingDataFlowBackend(
project=GCP_PROJECT,
staging_location=os.path.join(GCP_BUCKET, 'dataflow_processing/staging'),
)
# Run processing step with that backend
training_pipeline_3.add_split(
RandomSplit(split_map={'train': 0.7, 'eval': 0.2, 'test': 0.1}).with_backend(
processing_backend)
)
training_pipeline_3.run(artifact_store=cloud_artifact_store)
```
# Easily train on the cloud
```
training_pipeline_4 = training_pipeline.copy('Experiment 4')
# Add a trainer with a GCAIP backend
training_backend = SingleGPUTrainingGCAIPBackend(
project=GCP_PROJECT,
job_dir=TRAINING_JOB_DIR
)
training_pipeline_4.add_trainer(TFFeedForwardTrainer(
loss='binary_crossentropy',
last_activation='sigmoid',
output_units=1,
metrics=['accuracy'],
epochs=20).with_backend(training_backend))
training_pipeline_4.run(artifact_store=cloud_artifact_store)
```
# Orchestrate pipeline whereever you like
```
training_pipeline_5 = training_pipeline.copy('Experiment 5')
# Define the orchestrator backend
cloud_orchestrator_backend = OrchestratorGCPBackend(
cloudsql_connection_name=CONNECTION_NAME,
project=GCP_PROJECT,
preemptible=True, # reduce costs by using preemptible instances
machine_type='n1-standard-4',
gpu='nvidia-tesla-k80',
gpu_count=1,
)
# Run the pipeline
training_pipeline_5.run(
backend=cloud_orchestrator_backend,
metadata_store=cloud_metadata_store,
artifact_store=cloud_artifact_store,
)
```
# Add a deployer step with different integrations
## Option 1: Deploy to Google Cloud AI Platform
```
training_pipeline_6 = training_pipeline.copy('Experiment 6')
training_pipeline_6.add_deployment(
GCAIPDeployer(
project_id=GCP_PROJECT,
model_name=MODEL_NAME,
)
)
training_pipeline_6.run(artifact_store=cloud_artifact_store)
```
## Option 2: Deploy to Kubernetes via Cortex
```
training_pipeline_7 = training_pipeline.copy('Experiment 7')
# Add cortex deployer
api_config = {
"name": MODEL_NAME,
"kind": "RealtimeAPI",
"predictor": {
"type": "tensorflow",
"models": {"signature_key": "serving_default"}}
}
training_pipeline_7.add_deployment(
CortexDeployer(
env=CORTEX_ENV,
api_config=api_config,
predictor=TensorFlowPredictor,
)
)
training_pipeline_7.run(artifact_store=cloud_artifact_store)
```
| github_jupyter |
```
import pickle
from collections import deque
import gym
import imageio
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
from torch.distributions import Categorical
from running_mean_std import RunningMeanStd
class ActorCriticNet(nn.Module):
def __init__(self, obs_space, action_space):
super().__init__()
h = 64
self.head = nn.Sequential(
nn.Linear(obs_space, h),
nn.Tanh()
)
self.pol = nn.Sequential(
nn.Linear(h, h),
nn.Tanh(),
nn.Linear(h, action_space)
)
self.val = nn.Sequential(
nn.Linear(h, h),
nn.Tanh(),
nn.Linear(h, 1)
)
self.log_softmax = nn.LogSoftmax(dim=-1)
def forward(self, x):
out = self.head(x)
logit = self.pol(out).reshape(out.shape[0], -1)
log_p = self.log_softmax(logit)
v = self.val(out).reshape(out.shape[0], 1)
return log_p, v
def get_action_and_value(obs, old_net):
old_net.eval()
with torch.no_grad():
state = torch.tensor([obs]).to(device).float()
log_p, v = old_net(state)
m = Categorical(log_p.exp())
action = m.sample()
return action.item(), v.item()
```
## Main
```
# set device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# make an environment
# env = gym.make('CartPole-v0')
env = gym.make('CartPole-v1')
# env = gym.make('MountainCar-v0')
# env = gym.make('LunarLander-v2')
SEED = 0
env.seed(SEED)
obs_space = env.observation_space.shape[0]
action_space = env.action_space.n
OBS_NORM = True
n_episodes = 10000
n_eval = env.spec.trials
# global values
total_steps = 0
obses = []
rewards = []
reward_eval = deque(maxlen=n_eval)
# load a model
old_net = ActorCriticNet(obs_space, action_space).to(device)
old_net.load_state_dict(torch.load(
'./saved_models/CartPole-v1_up36_clear_model_ppo_st.pt'))
with open('./saved_models/CartPole-v1_up36_clear_norm_obs.pkl', 'rb') as f:
norm_obs = pickle.load(f)
env.spec.max_episode_steps
env.spec.trials
env.spec.reward_threshold
# play
# frames = []
for i in range(1, n_episodes + 1):
obs = env.reset()
done = False
ep_reward = 0
while not done:
# frames.append(env.render(mode = 'rgb_array'))
env.render()
if OBS_NORM:
obs_norm = np.clip(
(obs - norm_obs.mean) / np.sqrt(norm_obs.var+1e-8),
-10, 10)
action, _ = get_action_and_value(obs_norm, old_net)
else:
action, _ = get_action_and_value(obs, old_net)
_obs, reward, done, _ = env.step(action)
obs = _obs
total_steps += 1
ep_reward += reward
if done:
env.render()
norm_obs.update(_obs)
rewards.append(ep_reward)
reward_eval.append(ep_reward)
print('{:3} Episode in {:5} steps, reward {:.2f}'.format(
i, total_steps, ep_reward))
# frames.append(env.render(mode = 'rgb_array'))
# imageio.mimsave(f'{env.spec.id}.gif', frames,)
if len(reward_eval) >= n_eval:
if np.mean(reward_eval) >= env.spec.reward_threshold:
print('\n{} is sloved! {:3} Episode in {:3} steps'.format(
env.spec.id, i, total_steps))
print(np.mean(reward_eval))
break
env.close()
plt.figure(figsize=(15, 5))
plt.title('reward')
plt.plot(rewards)
plt.show()
[
('CartPole-v0', 412, 1),
('CartPole-v1', 452, 0.05),
('MountainCar-v0', 193, 0.1),
('LunarLander-v2', 260, 0.1)
]
```
| github_jupyter |
## BLU06 - Learning Notebook - Part 2 of 3 - Time Series Pre-processing and Feature Engineering
```
import pandas as pd
from sklearn.linear_model import LinearRegression
from matplotlib import pyplot as plt
from sklearn.ensemble import GradientBoostingRegressor
from joblib import Parallel, delayed
from sklearn.metrics import mean_absolute_error
%matplotlib inline
import numpy as np
plt.rcParams['figure.figsize'] = (16, 4)
from tqdm import tqdm_notebook as tqdm
import warnings
warnings.filterwarnings(action="ignore")
warnings.filterwarnings(action="ignore", module="scipy", message="^internal gelsd")
from utils import * # We've added all the functions from the last BLU to the utils.py
```
In the previous notebook we learned how to formulate time series multi-step forecasting as a regression problem where any ML model can be applied. But the airlines dataset is quite different from the ones you'll find in the wild. In this notebook we'll exlore a dataset that contains the customers that a store had with daily frequency. We'll learn about data pre-processing, feature engineering and play with gradient boosting and compare it with linear regression.
```
store = get_store_data()
store['date'] = pd.to_datetime(store['date'], format='%Y-%m-%d')
store = store.set_index('date')
store = store.sort_index()
```
# Data Pre-processing
Whenever analysing time series data it's always important to verify if it has all the timestamps. Let's look at our data:
```
store.head()
store.isnull().sum()
```
It seems to have daily frequency. To make sure we have no missing days we can resample the data to a daily frequency. This will add all possibly non-existing days as NaNs.
```
store_resampled = store.resample('D').mean()
store_resampled.isnull().sum()
```
Ok! So there were 11 missing days. Let's check which days where these:
```
store_resampled[store_resampled.isnull()['customers']]
```
Interesting! During Christmas and New Year the store should be closed, hence why there was no entry for the number of customers. Since we have no extra information regarding the other days we'll assume the store was also closed. So we have to replace this NaNs be 0s.
```
store_cleaned = store_resampled.fillna(0)
store_cleaned.isnull().sum()
```
# Feature Engineering
In the last BLU we used the simplest feature to predict time series: lags. But there's a lot more you can do! First let's see what a model with lags similar to the one we were using before can do.
```
split_date = '2017-6'
train = store_cleaned.loc[store_cleaned.index < split_date]
test = store_cleaned.loc[store_cleaned.index >= split_date]
predictions = predict_n_periods(series_=train,
n_periods=len(test),
model=LinearRegression(),
num_periods_lagged=30
)
store[1500:].plot(label="original data")
pd.Series(predictions, index=test.index).plot(label="pred")
plt.legend();
mean_absolute_error(test,predictions)
```
### Lags and Diffs
Similar to lags, we can also use diffs of the time series to predict it. It's possible that knowing if the sales are increasing or decreasing can help predict better the next step. We can add this feature by simply doing:
```
store_features = store_cleaned.copy()
store_features['diff'] = store_cleaned.diff()
store_cleaned.head()
store_features.head()
```
### Rolling Window features
Another type of feature that may be relevant are rolling window features. As you may recall, a rolling window refers to a window of days before the current step. We can extract features from that window such as the minimum, mean, maximum and standard deviation. To give information to the model about the recent statistical properties of the time series.
```
store_features['rolling_max'] = store_features['customers'].rolling('7D').max()
store_features['rolling_min'] = store_features['customers'].rolling('7D').min()
store_features['rolling_mean'] = store_features['customers'].rolling('7D').mean()
store_features['rolling_std'] = store_features['customers'].rolling('7D').std()
```
### Datetime features
Datetime features refer to those you can extract simply from the timestamp. For example, giving information regarding which day of the week it is, which month, etc. can go a long way to help the model understand the time series better. For example, in our case we already know that the store is closed in Christmas and the New Year so we can one-hot encode those as a feature to the model.
```
holidays = store_features[((store_features.index.month==12) & (store_features.index.day==25))
|((store_features.index.month==1) & (store_features.index.day==1))].customers
store_features['holidays'] = holidays + 1
store_features['holidays'] = store_features['holidays'].fillna(0)
```
Now let's add the day of the week and month features:
```
store_features['day_of_week'] = store_features.index.weekday
store_features['month'] = store_features.index.month
store_features.head(10)
```
So, regarding day of week you can check pandas [documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DatetimeIndex.dayofweek.html) on what each number means. Basically, Monday is 0 and Sunday is 6. But this brings a good point, is Monday closer to Sunday than to Thursday? "Of course it is closer, what kind of question is that?!" Well, the issue is that the current mapping between day of the week and number is the following:
- Monday: 0
- Thursday: 3
- Sunday: 6
This means that the distance between Monday and Thursday is smaller than Monday and Sunday, which gives wrong intuition to the model. This is particularly important for linear models, as well as kNNm K-means and neural networks. This means that we have to encode these features in a such a way that they convey their cyclical nature to the model.
### Circular Encoding
So how can we encode these features while mantaining their cyclical nature? We can use circular encoding. Basically, the idea is to encode these features using the sine and the cosine. For more information on this, check the optional notebook of this BLU. In practice this is how you add them:
```
store_features['sin_weekday'] = np.sin(2*np.pi*store_features.index.weekday/7)
store_features['cos_weekday'] = np.sin(2*np.pi*store_features.index.weekday/7)
store_features['sin_month'] = np.sin(2*np.pi*store_features.index.month/12)
store_features['cos_month'] = np.sin(2*np.pi*store_features.index.month/12)
```
Now let's drop the previous datetime features that were not encoded
```
store_features = store_features.drop(['day_of_week','month'], axis=1)
```
So our dataframe has a couple more features now:
```
store_features.head()
```
So let's see if any of these actually improve our model, but we have to test all these models in a validation set.
Note: With domain knowledge you can add even more features, such as: is it an holiday? is there a special Event? Even the weather! Feature engineering has a lot to do with creativity and thinking outside of the box.
# Validation
To test which features actually improve model performance we'll add them to the _build_some_features_ function that we started creating on the last BLU:
```
def build_some_features(df_, num_periods_lagged=1, num_periods_diffed=0, weekday=False, month=False, rolling=[], holidays=False):
"""
Builds some features by calculating differences between periods
"""
# make a copy
df_ = df_.copy()
# for a few values, get the lags
for i in range(1, num_periods_lagged+1):
# make a new feature, with the lags in the observed values column
df_['lagged_%s' % str(i)] = df_['customers'].shift(i)
# for a few values, get the diffs
for i in range(1, num_periods_diffed+1):
# make a new feature, with the diffs in the observed values column
df_['diff_%s' % str(i)] = df_['customers'].diff(i)
for stat in rolling:
df_['rolling_%s'%str(stat)] = df_['customers'].rolling('7D').aggregate(stat)
if weekday == True:
df_['sin_weekday'] = np.sin(2*np.pi*df_.index.weekday/7)
df_['cos_weekday'] = np.sin(2*np.pi*df_.index.weekday/7)
if month == True:
df_['sin_month'] = np.sin(2*np.pi*df_.index.month/12)
df_['cos_month'] = np.sin(2*np.pi*df_.index.month/12)
if holidays == True:
holidays = df_[((df_.index.month==12) & (df_.index.day==25)) |((df_.index.month==1) & (df_.index.day==1))].customers
df_['holidays'] = holidays + 1
df_['holidays'] = df_['holidays'].fillna(0)
return df_
```
Now let's create a parameter grid with the combinations we want to test using sklearn's [ParameterGrid](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.ParameterGrid.html). The more combinations the more time it will take to run, so we'll just assume that the rolling statistics and the holidays improve the model. But feel free to add more options to the parameter grid and see if it improves the performance on the validation set.
Note that we're also adding the gradient boosting regressor model, to see if it obtains better results than the vanilla linear regression.
```
from sklearn.model_selection import ParameterGrid
param_grid = {'model': [LinearRegression(), GradientBoostingRegressor()],
'num_periods_lagged':np.arange(1,3),
'num_periods_diffed':np.arange(0,3),
'weekday':[True,False],
'month':[True,False],
'holidays': [True],
'rolling' : [[np.mean,np.min,np.max,np.std]]
}
grid = ParameterGrid(param_grid)
```
Now we can test on a validation set to see which combination of features gives best performance!
```
val_split_date = '2017-3'
test_split_date = '2017-6'
train = store_cleaned.loc[store_cleaned.index < val_split_date]
val = store_cleaned.loc[(val_split_date <= store_cleaned.index) & (store_cleaned.index < test_split_date)]
test = store_cleaned.loc[store_cleaned.index >= test_split_date]
```
The commented cell below implements the for loop that runs through the grid we created and finds the parameter group that minimizes the MAE in the validation set. It is commented because it takes some time. The other cell makes use of the Joblib library to parallelize the code and is considerably faster. In this laptop it takes about 3 minutes.
```
#error_lst = []
#for params in tqdm(grid):
# predictions = predict_n_periods(series_=train,
# n_periods=30,
# model=params['model'],
# num_periods_lagged=params['num_periods_lagged'],
# num_periods_diffed=params['num_periods_diffed'],
# weekday=params['weekday'],
# month=params['month'],
# rolling=[np.mean,np.max,np.min]
# )
#
# error_lst.append(mean_absolute_error(val,predictions))
# pd.Series(error_lst).idxmin()
%%time
# This is another cell that will take a long time to run.
def wrap_model_selection(params):
predictions = predict_n_periods(series_=train,
n_periods=len(val),
model=params['model'],
num_periods_lagged=params['num_periods_lagged'],
num_periods_diffed=params['num_periods_diffed'],
weekday=params['weekday'],
month=params['month'],
rolling=[np.mean,np.max,np.min]
)
return [params,mean_absolute_error(val,predictions)]
res = Parallel(n_jobs=-1)(delayed(wrap_model_selection)(params=params)
for params in tqdm(grid))
df = pd.DataFrame(res, columns=['params','error'])
df.sort_values('error').head()
```
Let's inspect our best model a bit better:
```
df.sort_values('error').iloc[0][0]
```
It seems like the diffed feature and the month didn't really help. On the other hand the weekday feature and the gradient boosting regressor improved the performance.
# Testing
Now we can finally test on the test set with the best parameters and see if it improved since our dataset without feature engineering. But first we need to append the validation set to the training set
```
train = train.append(val)
predictions = predict_n_periods(series_=train,
n_periods=len(test),
model=GradientBoostingRegressor(),
num_periods_lagged=2,
num_periods_diffed=0,
weekday=True,
month=False,
rolling=[np.mean,np.min,np.max,np.std],
holidays=True
)
store[1500:].plot(label="original data")
pd.Series(predictions, index=test.index).plot(label="pred")
plt.legend();
mean_absolute_error(test,predictions)
```
Wow, that's a big improvement from the non-feature-engineered model. Feature engineering works!
But it's interesting that the MAE in the test set is lower than on the val set. This may go a bit against your expectations: Since we did the tunning on the val set we would expect it to perform better there. But that doens't happen. Remember when we talked about stationarity in the last BLU? Well, we'll talk about it in the next notebook.
| github_jupyter |

## 一. Building a Spam Classifier
垃圾邮件分类就是一个 0/1 分类问题,可以用逻辑回归完成,这里不再重复介绍逻辑回归的过程了,我们考虑如何降低分类错误率:
- 尽可能的扩大数据样本:Honypot 做了这样一件事,把自己包装成一个对黑客极具吸引力的机器,来诱使黑客进行攻击,就像蜜罐(honey pot)吸引密封那样,从而记录攻击行为和手段。
- 添加更多特征:例如我们可以增加邮件的发送者邮箱作为特征,可以增加标点符号作为特征(垃圾邮件总会充斥了?,!等吸引眼球的标点)。
- 预处理样本:正如我们在垃圾邮件看到的,道高一尺,魔高一丈,垃圾邮件的制造者也会升级自己的攻击手段,如在单词拼写上做手脚来防止邮件内容被看出问题,例如把 medicine 拼写为 med1cinie 等。因此,我们就要有手段来识别这些错误拼写,从而优化我们输入到逻辑回归中的样本。
假如我们要用机器学习解决一个问题,那么最好的实践方法就是:
1.建立一个简单的机器学习系统,用简单的算法快速实现它。
2.通过画出学习曲线,以及检验误差,来找出我们的算法是否存在高偏差或者高方差的问题,然后再通过假如更多的训练数据、特征变量等等来完善算法。
3.误差分析。例如在构建垃圾邮件分类器,我们检查哪一类型的邮件或者那些特征值总是导致邮件被错误分类,从而去纠正它。当然,误差的度量值也是很重要的,例如我们可以将错误率表示出来,用来判断算法的优劣。
----------------------------------------------------------------------------------------------------------------
## 二. Handling Skewed Data
评估一个模型的好坏,通常使用误差分析可视化,即把预测的准确率(Accuracy)显示出来,其实这样是有缺陷的。这种误差度量又被称为偏斜类(Skewed Classes)问题。
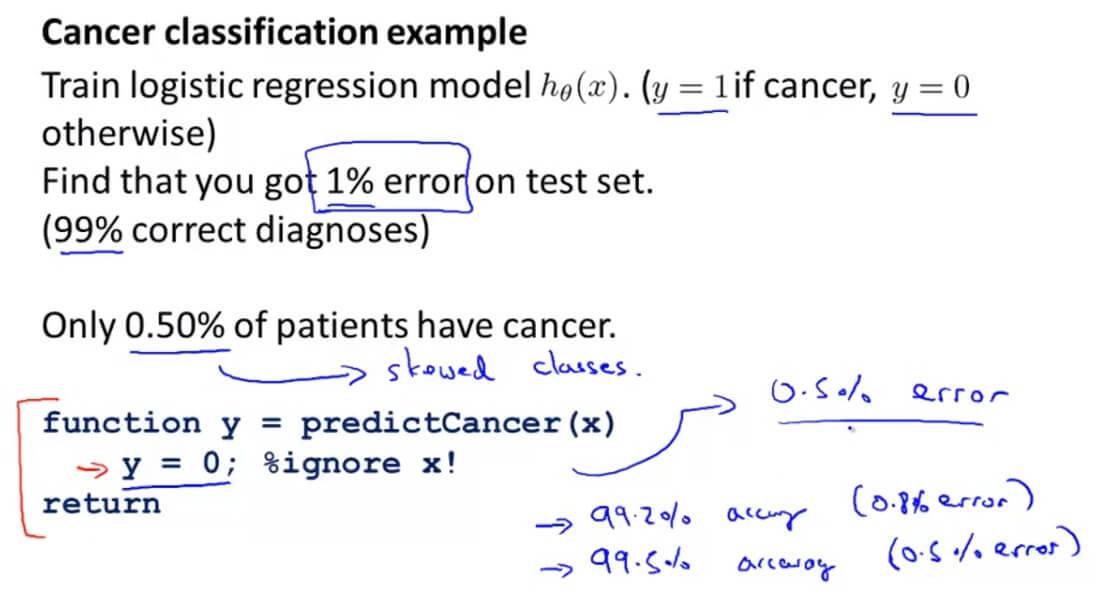

举个例子:假如我们做癌症分析,最后得出该算法只有1%的误差,也就是说准确率达到了99% 。这样看起来99%算是非常高的了,但是我们发现在训练集里面只有0.5%的患者患有癌症,那么这1%的错误率就变得那么准确了。我们再举个极端一点的例子,无论输入是什么,所有预测输出的数据都为0(也就是非癌症),那么我们这里的正确率是99.5%,但是这样的判断标准显然不能体现分类器的性能。
这是因为两者的数据相差非常大,在这里因为癌症的样本非常少,所以导致了预测的结果就会偏向一个极端,我们把这类的情况叫做偏斜类(Skewed Classes)问题。
所以我们需要另一种的评估方法,其中一种评估度量值叫做查准率(Precision)和召回率(Recall)。
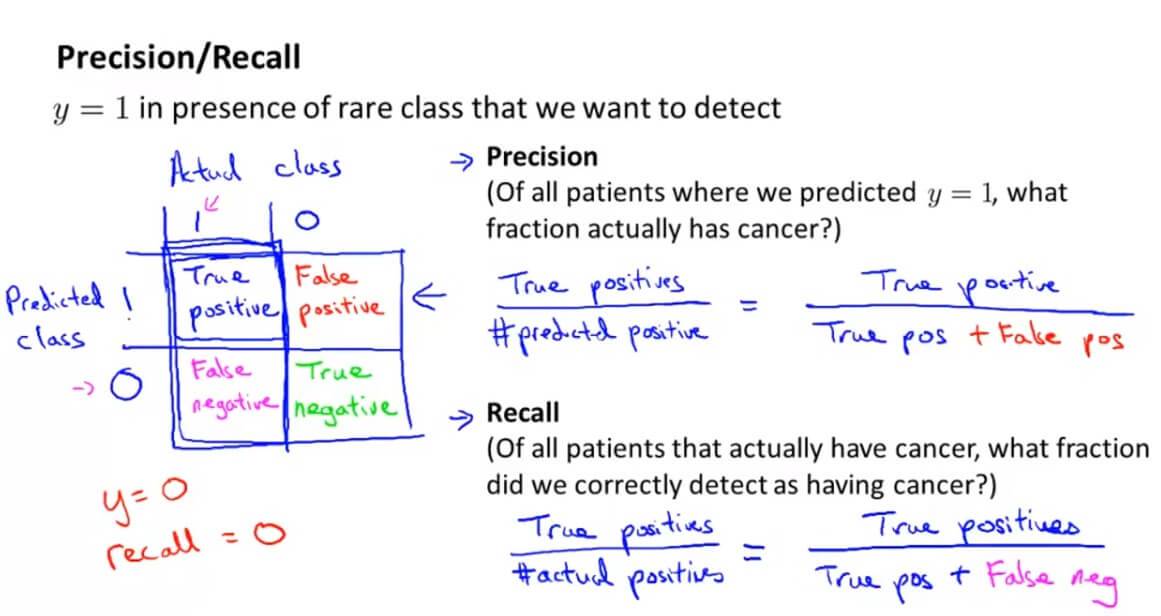
建立一个2 x 2的表格,横坐标为真实值,纵坐标为预测值,表格单元1-4分别代表:预测准确的正样本(True positive)、预测错误的正样本(False positive)、预测错误的负样本(False negative)、预测正确的负样本(True negative)。
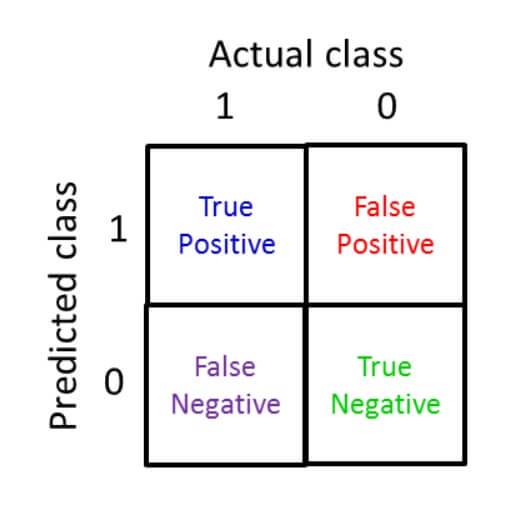
查准率(Precision)= 预测准确的正样本(True positive)/预测的正样本(predicted positive),而其中预测的正样本自然就包括了 预测准确的正样本+ 预测错误的正样本。
$$Precision=\frac{True\;positive}{Predicated\;as\;positive }=\frac{True\;positive}{True\;positive+False\;positive}$$
召回率(Recall)= 预测准确的正样本(True positive)/实际的正样本(actual positive),而其中实际的正样本自然就包括了 预测准确的正样本+ 预测错误的负样本。
$$Recall=\frac{True\;positive}{Actual\;positive}=\frac{True\;positive}{True\;positive+False\;negative}$$
假如像之前的y一直为0,虽然其准确率为99%,但是其召回率是0%。所以这对于评估算法的正确性是非常有帮助的。
那么查准率(Precision)和召回率(Recall)应该如何评估呢?
假如我们选用两者的平均值,这样看起来可行,但是对于之前的极端例子还是不适用。假如我们预测的y一直为1,那么其召回率就100%了,而查准率非常低,但是平均下来还是相对不错。所以我们采用下面一种评估方法,叫做 F 值:
$$F_1\;Score = 2\frac{PR}{P+R}$$
P 指的是 Precision,R 指的是 Recall。
----------------------------------------------------------------------------------------------------------------
## 三. Using Large Data Sets
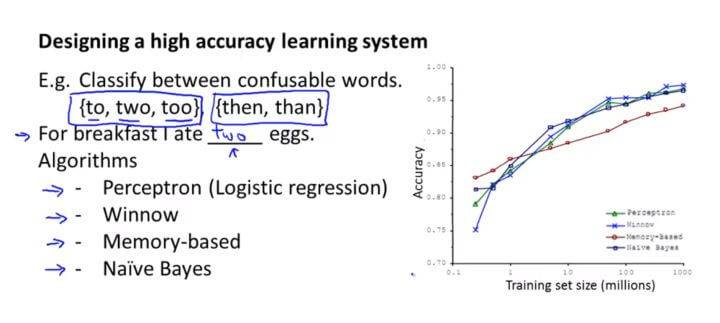
在机器学习领域,流传着这样一句话:
>It's not who has the best algorithm that wins. It's who has the most data.
取得成功的人不是拥有最好算法的人,而是拥有最多数据的人。
这是为什么呢?
首先我们因为有大量的特征量,去训练数据,这样就导致了我们的训练集误差非常小,也就是 $J_{train}(\theta)$ 非常小。然后我们提供了大量的训练数据,这样有利于防止过拟合,可以使得 $J_{train}(\theta)\approx J_{test}(\theta)$ 。这样,我们的假设函数既不会存在高偏差,也不会存在高方差,所以相对而言,大数据训练出来会更加准确。
注意了,这里不仅是由大量的训练数据,而且还要有更多的特征量。因为假如只有一些特征量,例如只有房子的大小,去预测房子的价格,那么就连世界最好的销售员也不能只凭房子大小就能告诉你房子的价格是多少。
什么时候采用大规模的数据集呢,一定要保证模型拥有足够的参数(线索),对于线性回归/逻辑回归来说,就是具备足够多的特征,而对于神经网络来说,就是更多的隐层单元。这样,足够多的特征避免了高偏差(欠拟合)问题,而足够大数据集避免了多特征容易引起的高方差(过拟合)问题。
----------------------------------------------------------------------------------------------------------------
## 四. Machine Learning System Design 测试
### 1. Question 1
You are working on a spam classification system using regularized logistic regression. "Spam" is a positive class (y = 1) and "not spam" is the negative class (y = 0). You have trained your classifier and there are m = 1000 examples in the cross-validation set. The chart of predicted class vs. actual class is:
Actual Class: 1 Actual Class: 0
Predicted Class: 1 85 890
Predicted Class: 0 15 10
For reference:
- Accuracy = (true positives + true negatives) / (total examples)
- Precision = (true positives) / (true positives + false positives)
- Recall = (true positives) / (true positives + false negatives)
- F1 score = (2 * precision * recall) / (precision + recall)
What is the classifier's F1 score (as a value from 0 to 1)?
Enter your answer in the box below. If necessary, provide at least two values after the decimal point.
解答:0.158
代入公式 $2\frac{PR}{P+R}$ 计算即可。
### 2. Question 2
Suppose a massive dataset is available for training a learning algorithm. Training on a lot of data is likely to give good performance when two of the following conditions hold true.
Which are the two?
A. When we are willing to include high order polynomial features of x (such as $x_{1}^{2}$, $x_{2}^{2}$,$x_{1}$,$x_{2}$, etc.).
B. The features x contain sufficient information to predict y accurately. (For example, one way to verify this is if a human expert on the domain can confidently predict y when given only x).
C. We train a learning algorithm with a small number of parameters (that is thus unlikely to overfit).
D. We train a learning algorithm with a large number of parameters (that is able to learn/represent fairly complex functions).
解答:B、D
A. 需要的是足够的特征量而不是高阶。
B. 特征量有足够的信息来准确预测。
C. 少量的特征量显然是不行的。
D. 要有足够多的变量(特征量)。
### 3. Question 3
Suppose you have trained a logistic regression classifier which is outputing hθ(x).
Currently, you predict 1 if $h_{\theta}(x)\geqslant threshold$, and predict 0 if $h_{\theta}(x)<threshold$, where currently the threshold is set to 0.5.
Suppose you decrease the threshold to 0.3. Which of the following are true? Check all that apply.
A. The classifier is likely to have unchanged precision and recall, but higher accuracy.
B. The classifier is likely to now have higher precision.
C. The classifier is likely to now have higher recall.
D. The classifier is likely to have unchanged precision and recall, but lower accuracy.
解答:C
将阈值调低的结果只会导致召回率增大,查准率降低。
### 4. Question 4
Suppose you are working on a spam classifier, where spam emails are positive examples (y=1) and non-spam emails are negative examples (y=0). You have a training set of emails in which 99% of the emails are non-spam and the other 1% is spam. Which of the following statements are true? Check all that apply.
A. If you always predict non-spam (output y=0), your classifier will have 99% accuracy on the training set, but it will do much worse on the cross validation set because it has overfit the training data.
B. If you always predict non-spam (output y=0), your classifier will have 99% accuracy on the training set, and it will likely perform similarly on the cross validation set.
C. A good classifier should have both a high precision and high recall on the cross validation set.
D. If you always predict non-spam (output y=0), your classifier will have an accuracy of 99%.
解答:B、C、D
A. 在交叉验证集因为过拟合的问题会使准确率下降,这不是过拟合的问题,是偏斜类的问题。
B. 假如训练集有99%准确率,那么交叉验证集也有很大可能有99%的准确率,这是正确的,因为数据是随机分布的,训练集的数据分布跟交叉验证集的数据分布相似。
C. 一个好的分类器应该查准率和召回率都比较高,正确。
D. 假如我们都把结果设为全为非垃圾邮件,那么准确率将达到99%,正确。
### 5. Question 5
Which of the following statements are true? Check all that apply.
A. On skewed datasets (e.g., when there are more positive examples than negative examples), accuracy is not a good measure of performance and you should instead use F1 score based on the precision and recall.
B. If your model is underfitting the training set, then obtaining more data is likely to help.
C. After training a logistic regression classifier, you must use 0.5 as your threshold for predicting whether an example is positive or negative.
D. It is a good idea to spend a lot of time collecting a large amount of data before building your first version of a learning algorithm.
E. Using a very large training set makes it unlikely for model to overfit the training data.
解答:A、E
A.利用 F1 score 去衡量准确性,正确。
B.模型不适合训练集,是欠拟合,欠拟合增大数据样本没用。
C.阈值不一定是0.5。
D.在建立第一个学习算法前花大量时间收集数据显然有可能走向浪费时间的不归路。
E.用更多的数据样本可以解决过拟合的现象,正确。
----------------------------------------------------------------------------------------------------------------
> GitHub Repo:[Halfrost-Field](https://github.com/halfrost/Halfrost-Field)
>
> Follow: [halfrost · GitHub](https://github.com/halfrost)
>
> Source: [https://github.com/halfrost/Halfrost-Field/blob/master/contents/Machine\_Learning/Machine\_Learning\_System\_Design.ipynb](https://github.com/halfrost/Halfrost-Field/blob/master/contents/Machine_Learning/Machine_Learning_System_Design.ipynb)
| github_jupyter |
# *grama* Analysis Demo
---
*grama* is a *grammar of model analysis*---a language for describing and analyzing mathematical models. Heavily inspired by [ggplot](https://ggplot2.tidyverse.org/index.html), `py_grama` is a Python package that implements *grama* by providing tools for defining and exploring models.
This notebook illustrates how one can use *grama* to ***analyze a fully-defined model***.
Note that you will need to install `py_grama`, a fork of `dfply`, and dependencies in order to run this notebook. See the [installation instructions](https://github.com/zdelrosario/py_grama) for details.
```
### Setup
import grama as gr
import numpy as np
import pandas as pd
import seaborn as sns
X = gr.Intention()
```
# Quick Tour: Analyzing a model
---
*grama* separates the model *definition* from model *analysis*; once the model is fully defined, only minimal information is necessary for further analysis.
As a quick demonstration, we import a fully-defined model provided with *grama*, and carry out a few analyses.
```
from grama.models import make_cantilever_beam
md_beam = make_cantilever_beam()
md_beam.printpretty()
```
The method `printpretty()` gives us a quick summary of the model; we can see this model has two deterministic variables `w,t` and four random variables `H,V,E,Y`. All of the variables affect the outputs `g_stress, g_displacement`, while only `w,t` affect `c_area`. Since there are random variables, there is a source of *uncertainty* which we must consider when studying this model.
## Studying model behavior with uncertainty
Since the model has sources of randomness (`var_rand`), we must account for this when studying its behavior. We can do so through a Monte Carlo analysis. We make decisions about the deterministic inputs by specifying `df_det`, and the `py_grama` function `gr.ev_monte_carlo` automatically handles the random inputs. Below we fix a nominal value `w = 0.5 * (2 + 4)`, sweep over values for `t`, and account for the randomness via Monte Carlo.
```
## Carry out a Monte Carlo analysis of the random variables
df_beam_mc = \
md_beam >> \
gr.ev_monte_carlo(
n=1e2,
df_det=gr.df_make( # Define deterministic levels
w=0.5*(2 + 4), # Single value
t=np.linspace(2.5, 3, num=10) # Sweep
)
)
```
To help plot the data, we use `gr.tf_gather` to reshape the data, and `seaborn` to quickly visualize results.
```
df_beam_wrangled = \
df_beam_mc >> \
gr.tf_gather("output", "y", ["c_area", "g_stress", "g_disp"])
g = sns.FacetGrid(df_beam_wrangled, col="output", sharey=False)
g.map(sns.lineplot, "t", "y")
```
The mean behavior of the model is shown as a solid line, while the band visualizes the standard deviation of the model output. From this plot, we can see:
- The random variables have no effect on `c_area` (there is no band)
- Comparing `g_stress` and `g_displacement`, the former is more strongly affected by the random inputs, as illustrated by its wider uncertainty band.
While this provides a visual description of how uncertainty affects our outputs, we might be interested in *how* the different random variables affect our outputs.
## Probing random variable effects
One way to quantify the effects of random variables is through *Sobol' indices*, which quantify variable importance by the fraction of output variance "explained" by each random variable. Since distribution information is included in the model, we can carry out a *hybrid-point Monte Carlo* and analyze the results with two calls to `py_grama`.
```
df_sobol = \
md_beam >> \
gr.ev_hybrid(n=1e3, df_det="nom", seed=101) >> \
gr.tf_sobol()
df_sobol
```
The indices should lie between `[0, 1]`, but estimation error can lead to violations. These results suggest that `g_stress` is largely insensitive to `E`, while `g_disp` is insensitive to `Y`. For `g_disp`, the input `V` contributes about twice the variance as variables `H,E`.
To get a *qualitative* sense of how the random variables affect our model, we can perform a set of sweeps over random variable space with a *sinew* design. First, we visualize the design in the six-dimensional full input space.
```
md_beam >> \
gr.ev_sinews(n_density=50, n_sweeps=10, df_det="swp", skip=True) >> \
gr.pt_auto()
```
The `skip` keyword argument allows us to delay evaluating a model; this is useful for inspecting a design before running a potentially expensive calculation. The `pt_auto()` function automatically detects DataFrames generated by `py_grama` functions and constructs an appropriate visualization. This is provided for convenience; you are of course welcome (and encouraged!) to create your own visualizations of the data.
Here we can see the sweeps cross the domain in straight lines at random starting locations. Each of these sweeps gives us a "straight shot" within a single variable. Visualizing the outputs for these sweeps will give us a sense of a single variable's influence, contextualized by the effects of the other variables.
```
df_beam_sweeps = \
md_beam >> \
gr.ev_sinews(n_density=50, n_sweeps=10, df_det="swp")
df_beam_sweeps >> gr.pt_auto()
```
Removing the keyword argument `skip` falls back on the default behavior; the model functions are evaluated at each sample, and `pt_auto()` adjusts to use this new data.
Based on this plot, we can see:
- The output `c_area` is insensitive to all the random variables; it changes only with `t, w`
- As the Sobol' analyis above suggested `g_stress` is insensitive to `E`, and `g_displacement` is insensitive to `Y`
- Visualizing the results shows that inputs `H,E` tend to 'saturate' in their effects on `g_displacement`, while `V` is linear over its domain. This may explain the difference in contributed variance
- Furthermore both `t, w` seem to saturate in their effects on the two limit states---there are diminishing returns on making the beam taller or wider
# Theory: The *grama* language
---
As a language, *grama* has both *objects* and *verbs*.
### Objects
---
*grama* as a language considers two categories of objects:
- **data** (`df`): observations on various quantities, implemented by the Python package `Pandas`
- **models** (`md`): a function and complete description of its inputs, implemented by `py_grama`
For readability, we suggest using prefixes `df_` and `md_` when naming DataFrames and models.
Since data is already well-handled by Pandas, `py_grama` focuses on providing tools to handle models. A `py_grama` model has **functions** and **inputs**: The method `printpretty()` gives a quick summary of the model's inputs and function outputs. Model inputs are organized into:
| | Deterministic | Random |
| ---------- | ---------------------------------------- | ---------- |
| Variables | `model.var_det` | `model.var_rand` |
| Parameters | `model.density.marginals[i].d_param` | (Future*) |
- **Variables** are inputs to the model's functions
+ **Deterministic** variables are chosen by the user; the model above has `w, t`
+ **Random** variables are not controlled; the model above has `H, V, E, Y`
- **Parameters** define random variables
+ **Deterministic** parameters are currently implemented; these are listed under `var_rand` with their associated random variable
+ **Random** parameters* are not yet implemented
The `outputs` section lists the various model outputs. The model above has `c_area, g_stress, g_displacement`.
### Verbs
---
Verbs are used to take action on different *grama* objects. We use verbs to generate data from models, build new models from data, and ultimately make sense of the two.
The following table summarizes the categories of `py_grama` verbs. Verbs take either data (`df`) or a model (`md`), and may return either object type. The prefix of a verb immediately tells one both the input and output types. The short prefix is used to denote the *pipe-enabled version* of a verb.
| Verb Type | Prefix (Short) | In | Out |
| --------- | --------------- | ---- | ----- |
| Evaluate | `eval_` (`ev_`) | `md` | `df` |
| Fit | `fit_` (`ft_`) | `df` | `md` |
| Transform | `tran_` (`tf_`) | `df` | `df` |
| Compose | `comp_` (`cp_`) | `md` | `md` |
### Functional programming (Pipes)
---
`py_grama` provides tools to use functional programming patterns. Short-stem versions of `py_grama` functions are *pipe-enabled*, meaning they can be used in functional programming form with the pipe operator `>>`. These pipe-enabled functions are simply aliases for the base functions, as demonstrated below:
```
df_base = gr.eval_nominal(md_beam, df_det="nom")
df_functional = md_beam >> gr.ev_nominal(df_det="nom")
df_base.equals(df_functional)
```
Functional patterns enable chaining multiple commands, as demonstrated in the Sobol' index code above. In nested form using base functions, this would be:
```python
df_sobol = gr.tran_sobol(gr.eval_hybrid(md_beam, n=1e3, df_det="nom", seed=101))
```
From the code above, it is difficult to see that we first consider `md_beam`, perform a hybrid-point evaluation, then use those data to estimate Sobol' indices. With more chained functions, this only becomes more difficult. One could make the code significantly more readable by introducing intermediate variables:
```python
df_samples = gr.eval_hybrid(md_beam, n=1e3, df_det="nom", seed=101)
df_sobol = gr.tran_sobol(df_samples)
```
Conceptually, using *pipe-enabled* functions allows one to skip assigning intermediate variables, and instead pass results along to the next function. The pipe operator `>>` inserts the results of one function as the first argument of the next function. A pipe-enabled version of the code above would be:
```python
df_sobol = \
md_beam >> \
gr.ev_hybrid(n=1e3, df_det="nom", seed=101) >> \
gr.tf_sobol()
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import warnings
from sklearn.decomposition import PCA
from sklearn.feature_selection import RFE
from sklearn.feature_selection import RFECV
from sklearn.feature_selection import SelectKBest, chi2
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, f1_score
from sklearn.metrics import f1_score,confusion_matrix
from sklearn.model_selection import train_test_split
np.set_printoptions(precision=3)
pd.set_option('display.float_format', lambda x: '%.3f' % x)
warnings.filterwarnings('ignore')
%matplotlib inline
def generate_accuracy_and_heatmap(model, x, y):
ac = accuracy_score(y,model.predict(x))
f_score = f1_score(y,model.predict(x))
print('Accuracy is: ', ac)
print('F1 score is: ', f_score)
print ("\n")
print (pd.crosstab(pd.Series(model.predict(x), name='Predicted'),
pd.Series(y['Outcome'],name='Actual')))
return 1
df = pd.read_csv('diabetes.csv')
df.shape
df.info()
df.Outcome.value_counts()
df.head()
df['BloodPressureSquare'] = np.square(df['BloodPressure'])
df['BloodPressureCube'] = df['BloodPressure']**3
df['BloodPressureSqrt'] = np.sqrt(df['BloodPressure'])
df['GlucoseSquare'] = np.square(df['Glucose'])
df['GlucoseCube'] = df['Glucose']**3
df['GlucoseSqrt'] = np.sqrt(df['Glucose'])
df['GlucoseBloodPressure'] = df['BloodPressure'] * df['Glucose']
df['AgeBMI'] = df['Age'] * df['BMI']
df.head()
categorical_feature_columns = list(set(df.columns) - set(df._get_numeric_data().columns))
categorical_feature_columns
numerical_feature_columns = list(df._get_numeric_data().columns)
numerical_feature_columns
target = 'Outcome'
k = 15 #number of variables for heatmap
cols = df[numerical_feature_columns].corr().nlargest(k, target)[target].index
cm = df[cols].corr()
plt.figure(figsize=(10,6))
sns.heatmap(cm, annot=True, cmap = 'viridis')
X = df.loc[:, df.columns != target]
Y = df.loc[:, df.columns == target]
X.shape
Y.shape
x_train, x_test, y_train, y_test = train_test_split(X, Y,
test_size=0.33,
random_state=8)
clf_lr = LogisticRegression()
lr_baseline_model = clf_lr.fit(x_train,y_train)
generate_accuracy_and_heatmap(lr_baseline_model, x_test, y_test)
```
## Univariate feature selection by SELECTKBEST
```
select_feature = SelectKBest(chi2, k=5).fit(x_train, y_train)
selected_features_df = pd.DataFrame({'Feature':list(x_train.columns),
'Scores':select_feature.scores_})
selected_features_df.sort_values(by='Scores', ascending=False)
x_train_chi = select_feature.transform(x_train)
x_test_chi = select_feature.transform(x_test)
x_train.head(3)
x_train_chi[0:3]
lr_chi_model = clf_lr.fit(x_train_chi,y_train)
generate_accuracy_and_heatmap(lr_chi_model, x_test_chi, y_test)
```
## Recursive Feature Elimination
```
rfe = RFE(estimator=clf_lr, step=1)
rfe = rfe.fit(x_train, y_train)
selected_rfe_features = pd.DataFrame({'Feature':list(x_train.columns),
'Ranking':rfe.ranking_})
selected_rfe_features.sort_values(by='Ranking')
x_train_rfe = rfe.transform(x_train)
x_test_rfe = rfe.transform(x_test)
x_train_rfe[0:3]
lr_rfe_model = clf_lr.fit(x_train_rfe, y_train)
generate_accuracy_and_heatmap(lr_rfe_model, x_test_rfe, y_test)
```
## Recursive feature elimination with cross validation
```
rfecv = RFECV(estimator=clf_lr, step=1, cv=5, scoring='accuracy')
rfecv = rfecv.fit(x_train, y_train)
print('Optimal number of features :', rfecv.n_features_)
print('Best features :', x_train.columns[rfecv.support_])
rfecv.grid_scores_
plt.figure()
plt.xlabel("Number of features selected")
plt.ylabel("Cross validation score of number of selected features")
plt.plot(range(1, len(rfecv.grid_scores_) + 1), rfecv.grid_scores_)
plt.show()
x_train_rfecv = rfecv.transform(x_train)
x_test_rfecv = rfecv.transform(x_test)
lr_rfecv_model = clf_lr.fit(x_train_rfecv, y_train)
generate_accuracy_and_heatmap(lr_rfecv_model, x_test_rfecv, y_test)
```
| github_jupyter |
## Define the Convolutional Neural Network
After you've looked at the data you're working with and, in this case, know the shapes of the images and of the keypoints, you are ready to define a convolutional neural network that can *learn* from this data.
In this notebook and in `models.py`, you will:
1. Define a CNN with images as input and keypoints as output
2. Construct the transformed FaceKeypointsDataset, just as before
3. Train the CNN on the training data, tracking loss
4. See how the trained model performs on test data
5. If necessary, modify the CNN structure and model hyperparameters, so that it performs *well* **\***
**\*** What does *well* mean?
"Well" means that the model's loss decreases during training **and**, when applied to test image data, the model produces keypoints that closely match the true keypoints of each face. And you'll see examples of this later in the notebook.
---
## CNN Architecture
Recall that CNN's are defined by a few types of layers:
* Convolutional layers
* Maxpooling layers
* Fully-connected layers
You are required to use the above layers and encouraged to add multiple convolutional layers and things like dropout layers that may prevent overfitting. You are also encouraged to look at literature on keypoint detection, such as [this paper](https://arxiv.org/pdf/1710.00977.pdf), to help you determine the structure of your network.
### TODO: Define your model in the provided file `models.py` file
This file is mostly empty but contains the expected name and some TODO's for creating your model.
---
## PyTorch Neural Nets
To define a neural network in PyTorch, you define the layers of a model in the function `__init__` and define the feedforward behavior of a network that employs those initialized layers in the function `forward`, which takes in an input image tensor, `x`. The structure of this Net class is shown below and left for you to fill in.
Note: During training, PyTorch will be able to perform backpropagation by keeping track of the network's feedforward behavior and using autograd to calculate the update to the weights in the network.
#### Define the Layers in ` __init__`
As a reminder, a conv/pool layer may be defined like this (in `__init__`):
```
# 1 input image channel (for grayscale images), 32 output channels/feature maps, 3x3 square convolution kernel
self.conv1 = nn.Conv2d(1, 32, 3)
# maxpool that uses a square window of kernel_size=2, stride=2
self.pool = nn.MaxPool2d(2, 2)
```
#### Refer to Layers in `forward`
Then referred to in the `forward` function like this, in which the conv1 layer has a ReLu activation applied to it before maxpooling is applied:
```
x = self.pool(F.relu(self.conv1(x)))
```
Best practice is to place any layers whose weights will change during the training process in `__init__` and refer to them in the `forward` function; any layers or functions that always behave in the same way, such as a pre-defined activation function, should appear *only* in the `forward` function.
#### Why models.py
You are tasked with defining the network in the `models.py` file so that any models you define can be saved and loaded by name in different notebooks in this project directory. For example, by defining a CNN class called `Net` in `models.py`, you can then create that same architecture in this and other notebooks by simply importing the class and instantiating a model:
```
from models import Net
net = Net()
```
```
# load the data if you need to; if you have already loaded the data, you may comment this cell out
# -- DO NOT CHANGE THIS CELL -- #
!mkdir /data
!wget -P /data/ https://s3.amazonaws.com/video.udacity-data.com/topher/2018/May/5aea1b91_train-test-data/train-test-data.zip
!unzip -n /data/train-test-data.zip -d /data
```
<div class="alert alert-info">**Note:** Workspaces automatically close connections after 30 minutes of inactivity (including inactivity while training!). Use the code snippet below to keep your workspace alive during training. (The active_session context manager is imported below.)
</div>
```
from workspace_utils import active_session
with active_session():
train_model(num_epochs)
```
```
# import the usual resources
import matplotlib.pyplot as plt
import numpy as np
# import utilities to keep workspaces alive during model training
from workspace_utils import active_session
# watch for any changes in model.py, if it changes, re-load it automatically
%load_ext autoreload
%autoreload 2
## TODO: Define the Net in models.py
import torch
import torch.nn as nn
import torch.nn.functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
## TODO: Once you've define the network, you can instantiate it
# one example conv layer has been provided for you
from models import Net
net = Net().to(device)
print(net)
```
## Transform the dataset
To prepare for training, create a transformed dataset of images and keypoints.
### TODO: Define a data transform
In PyTorch, a convolutional neural network expects a torch image of a consistent size as input. For efficient training, and so your model's loss does not blow up during training, it is also suggested that you normalize the input images and keypoints. The necessary transforms have been defined in `data_load.py` and you **do not** need to modify these; take a look at this file (you'll see the same transforms that were defined and applied in Notebook 1).
To define the data transform below, use a [composition](http://pytorch.org/tutorials/beginner/data_loading_tutorial.html#compose-transforms) of:
1. Rescaling and/or cropping the data, such that you are left with a square image (the suggested size is 224x224px)
2. Normalizing the images and keypoints; turning each RGB image into a grayscale image with a color range of [0, 1] and transforming the given keypoints into a range of [-1, 1]
3. Turning these images and keypoints into Tensors
These transformations have been defined in `data_load.py`, but it's up to you to call them and create a `data_transform` below. **This transform will be applied to the training data and, later, the test data**. It will change how you go about displaying these images and keypoints, but these steps are essential for efficient training.
As a note, should you want to perform data augmentation (which is optional in this project), and randomly rotate or shift these images, a square image size will be useful; rotating a 224x224 image by 90 degrees will result in the same shape of output.
```
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# the dataset we created in Notebook 1 is copied in the helper file `data_load.py`
from data_load import FacialKeypointsDataset
# the transforms we defined in Notebook 1 are in the helper file `data_load.py`
from data_load import Rescale, RandomCrop, Normalize, ToTensor
## TODO: define the data_transform using transforms.Compose([all tx's, . , .])
# order matters! i.e. rescaling should come before a smaller crop
data_transform = transforms.Compose([Rescale(256),
RandomCrop(224),
Normalize(),
ToTensor()])
# testing that you've defined a transform
assert(data_transform is not None), 'Define a data_transform'
# create the transformed dataset
transformed_dataset = FacialKeypointsDataset(csv_file='./data/training_frames_keypoints.csv',
root_dir='./data/training/',
transform=data_transform)
print('Number of images: ', len(transformed_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
```
## Batching and loading data
Next, having defined the transformed dataset, we can use PyTorch's DataLoader class to load the training data in batches of whatever size as well as to shuffle the data for training the model. You can read more about the parameters of the DataLoader, in [this documentation](http://pytorch.org/docs/master/data.html).
#### Batch size
Decide on a good batch size for training your model. Try both small and large batch sizes and note how the loss decreases as the model trains. Too large a batch size may cause your model to crash and/or run out of memory while training.
**Note for Windows users**: Please change the `num_workers` to 0 or you may face some issues with your DataLoader failing.
```
# load training data in batches
batch_size_training = 48
train_loader = DataLoader(transformed_dataset,
batch_size=batch_size_training,
shuffle=True,
num_workers=4)
```
## Before training
Take a look at how this model performs before it trains. You should see that the keypoints it predicts start off in one spot and don't match the keypoints on a face at all! It's interesting to visualize this behavior so that you can compare it to the model after training and see how the model has improved.
#### Load in the test dataset
The test dataset is one that this model has *not* seen before, meaning it has not trained with these images. We'll load in this test data and before and after training, see how your model performs on this set!
To visualize this test data, we have to go through some un-transformation steps to turn our images into python images from tensors and to turn our keypoints back into a recognizable range.
```
# load in the test data, using the dataset class
# AND apply the data_transform you defined above
# create the test dataset
test_dataset = FacialKeypointsDataset(csv_file='./data/test_frames_keypoints.csv',
root_dir='./data/test/',
transform=data_transform)
print('Number of images: ', len(test_dataset))
# iterate through the transformed dataset and print some stats about the first few samples
for i in range(4):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['keypoints'].size())
# load test data in batches
batch_size_test = 10
test_loader = DataLoader(test_dataset,
batch_size=batch_size_test,
shuffle=True,
num_workers=4)
```
## Apply the model on a test sample
To test the model on a test sample of data, you have to follow these steps:
1. Extract the image and ground truth keypoints from a sample
2. Wrap the image in a Variable, so that the net can process it as input and track how it changes as the image moves through the network.
3. Make sure the image is a FloatTensor, which the model expects.
4. Forward pass the image through the net to get the predicted, output keypoints.
This function test how the network performs on the first batch of test data. It returns the images, the transformed images, the predicted keypoints (produced by the model), and the ground truth keypoints.
```
# test the model on a batch of test images
def net_sample_output():
# iterate through the test dataset
for i, sample in enumerate(test_loader):
# get sample data: images and ground truth keypoints
images = sample['image']
key_pts = sample['keypoints']
# convert images to FloatTensors
images = images.type(torch.FloatTensor)
images = images.to(device)
# forward pass to get net output
output_pts = net(images)
# reshape to batch_size x 68 x 2 pts
output_pts = output_pts.view(output_pts.size()[0], 68, -1)
# break after first image is tested
if i == 0:
return images, output_pts, key_pts
```
#### Debugging tips
If you get a size or dimension error here, make sure that your network outputs the expected number of keypoints! Or if you get a Tensor type error, look into changing the above code that casts the data into float types: `images = images.type(torch.FloatTensor)`.
```
# call the above function
# returns: test images, test predicted keypoints, test ground truth keypoints
test_images, test_outputs, gt_pts = net_sample_output()
# print out the dimensions of the data to see if they make sense
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
```
## Visualize the predicted keypoints
Once we've had the model produce some predicted output keypoints, we can visualize these points in a way that's similar to how we've displayed this data before, only this time, we have to "un-transform" the image/keypoint data to display it.
Note that I've defined a *new* function, `show_all_keypoints` that displays a grayscale image, its predicted keypoints and its ground truth keypoints (if provided).
```
def show_all_keypoints(image, predicted_key_pts, gt_pts=None):
"""Show image with predicted keypoints"""
# image is grayscale
plt.imshow(image, cmap='gray')
plt.scatter(predicted_key_pts[:, 0], predicted_key_pts[:, 1], s=20, marker='.', c='m')
# plot ground truth points as green pts
if gt_pts is not None:
plt.scatter(gt_pts[:, 0], gt_pts[:, 1], s=20, marker='.', c='g')
```
#### Un-transformation
Next, you'll see a helper function. `visualize_output` that takes in a batch of images, predicted keypoints, and ground truth keypoints and displays a set of those images and their true/predicted keypoints.
This function's main role is to take batches of image and keypoint data (the input and output of your CNN), and transform them into numpy images and un-normalized keypoints (x, y) for normal display. The un-transformation process turns keypoints and images into numpy arrays from Tensors *and* it undoes the keypoint normalization done in the Normalize() transform; it's assumed that you applied these transformations when you loaded your test data.
```
# visualize the output
# by default this shows a batch of 10 images
def visualize_output(test_images, test_outputs, gt_pts=None, batch_size=10):
for i in range(batch_size):
plt.figure(figsize=(20,10))
ax = plt.subplot(1, batch_size, i+1)
# un-transform the image data
image = test_images[i].data # get the image from it's Variable wrapper
image = image.cpu().numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
# un-transform the predicted key_pts data
predicted_key_pts = test_outputs[i].data
predicted_key_pts = predicted_key_pts.cpu().numpy()
# undo normalization of keypoints
predicted_key_pts = predicted_key_pts*50.0+100
# plot ground truth points for comparison, if they exist
ground_truth_pts = None
if gt_pts is not None:
ground_truth_pts = gt_pts[i]
ground_truth_pts = ground_truth_pts*50.0+100
# call show_all_keypoints
show_all_keypoints(np.squeeze(image), predicted_key_pts, ground_truth_pts)
plt.axis('off')
plt.show()
# call it
visualize_output(test_images, test_outputs, gt_pts, batch_size_test)
```
## Training
#### Loss function
Training a network to predict keypoints is different than training a network to predict a class; instead of outputting a distribution of classes and using cross entropy loss, you may want to choose a loss function that is suited for regression, which directly compares a predicted value and target value. Read about the various kinds of loss functions (like MSE or L1/SmoothL1 loss) in [this documentation](http://pytorch.org/docs/master/_modules/torch/nn/modules/loss.html).
### TODO: Define the loss and optimization
Next, you'll define how the model will train by deciding on the loss function and optimizer.
---
```
## TODO: Define the loss and optimization
import torch.optim as optim
criterion = nn.SmoothL1Loss()
# optimizer = optim.Adam(net.parameters())
optimizer = optim.Adam(net.parameters(), lr = 0.001)
```
## Training and Initial Observation
Now, you'll train on your batched training data from `train_loader` for a number of epochs.
To quickly observe how your model is training and decide on whether or not you should modify it's structure or hyperparameters, you're encouraged to start off with just one or two epochs at first. As you train, note how your the model's loss behaves over time: does it decrease quickly at first and then slow down? Does it take a while to decrease in the first place? What happens if you change the batch size of your training data or modify your loss function? etc.
Use these initial observations to make changes to your model and decide on the best architecture before you train for many epochs and create a final model.
```
def train_net(n_epochs):
# prepare the net for training
net.train()
for epoch in range(n_epochs): # loop over the dataset multiple times
running_loss = 0.0
# train on batches of data, assumes you already have train_loader
for batch_i, data in enumerate(train_loader):
# get the input images and their corresponding labels
images = data['image']
key_pts = data['keypoints']
# flatten pts
key_pts = key_pts.view(key_pts.size(0), -1)
# convert variables to floats for regression loss
key_pts = key_pts.type(torch.FloatTensor)
images = images.type(torch.FloatTensor)
# transfer data device
key_pts = key_pts.to(device)
images = images.to(device)
# forward pass to get outputs
output_pts = net(images)
# calculate the loss between predicted and target keypoints
loss = criterion(output_pts, key_pts)
# zero the parameter (weight) gradients
optimizer.zero_grad()
# backward pass to calculate the weight gradients
loss.backward()
# update the weights
optimizer.step()
# print loss statistics
running_loss += loss.item()
if (batch_i % 10) == 9: # print every 10 batches
print('Epoch: {}, Batch: {}, Avg. Loss: {}'.format(epoch + 1, batch_i+1, running_loss/10))
running_loss = 0.0
print('Finished Training')
# train your network
n_epochs = 15 # start small, and increase when you've decided on your model structure and hyperparams
# this is a Workspaces-specific context manager to keep the connection
# alive while training your model, not part of pytorch
with active_session():
train_net(n_epochs)
```
## Test data
See how your model performs on previously unseen, test data. We've already loaded and transformed this data, similar to the training data. Next, run your trained model on these images to see what kind of keypoints are produced. You should be able to see if your model is fitting each new face it sees, if the points are distributed randomly, or if the points have actually overfitted the training data and do not generalize.
```
# get a sample of test data again
test_images, test_outputs, gt_pts = net_sample_output()
print(test_images.data.size())
print(test_outputs.data.size())
print(gt_pts.size())
## TODO: visualize your test output
# you can use the same function as before, by un-commenting the line below:
visualize_output(test_images, test_outputs, gt_pts, 5)
```
Once you've found a good model (or two), save your model so you can load it and use it later!
Save your models but please **delete any checkpoints and saved models before you submit your project** otherwise your workspace may be too large to submit.
```
## TODO: change the name to something uniqe for each new model
model_dir = 'saved_models/'
model_name = 'keypoints_model_1.pt'
# after training, save your model parameters in the dir 'saved_models'
# torch.save(net.state_dict(), model_dir+model_name)
```
After you've trained a well-performing model, answer the following questions so that we have some insight into your training and architecture selection process. Answering all questions is required to pass this project.
### Question 1: What optimization and loss functions did you choose and why?
**Answer**:
I chose Smooth L1 Loss as loss function, and Adam as optimizer.
The reason why I chose loss function is that Smooth L1 Loss is not sensitive to outliers than MSE loss function.
Facial keypoints are densely pointed on face in each picture. So that, I assume that outliers appears frequently. Smooth L1 loss is the most expecting loss function among L1 Loss, MSE, and Smooth L1 Loss.
Adam is commonly used in deep neural network recently. Adam is a kind of optimizer which is combination of AdaGrad and Momentum. There are many research which adopts Adam as optimizer, and this optimizer is often evaluated positively compared to Stochastic gradient descent. That's why I chose this optmizer.
### Question 2: What kind of network architecture did you start with and how did it change as you tried different architectures? Did you decide to add more convolutional layers or any layers to avoid overfitting the data?
**Answer**:
I started very simple model: 2 convolution layer and 1 dense (fully-connected) layer without batch normalization and dropout. This model suffers from much under-fitting.
I adopted VGG-16 after confirming that simple model can predict the facial keypoint accurately. I adopted VGG-16 because it is very simple and easy to extend layers. My model has 14 convolution layers with which Batch Normalization layers are connected layer by layer. Besides, dense layers are following convolution layers. There are 2 hidden dense layers, each of which is followed by dropout, and 1 output dense layer.
I aims to avoid overfitting with using batch normalization and dropout. According to best practice shown in the lecture, I applied batch normalization to convolution layers and dropout to dense layers.
### Question 3: How did you decide on the number of epochs and batch_size to train your model?
**Answer**:
I tried to find the number of epochs and batch_size with tial and error approach. I prepare candidate value; 10, 15, and 20 for the number of epochs, and 16, 32, and 48 for `batch_size`.
I paid attention to computational cost for iteration by epoch. If I execute more number of epochs, I assumed, there might be more redundant computation for training. However, less number of epochs prevent my model from being trained accurately. Finally, I set 15 as the number of epochs according to prediction result with test dataset.
Additionally, `batch_size` should be considered carefully. More `batch_size` should require more memory consumption, but might improve the speed of training. Less `batch_size` should costs lots of computation, but might give us better result. As a result I chose 48 as `batch_size` because the prediction result showed that the best `batch_size` was 48. I think that less `batch_size` possibly cause over-fitting, but more `batch_size` cause under-fitting. 48 for `batch_size` was appropriate value which did not cause both over-fitting and under-fitting.
## Feature Visualization
Sometimes, neural networks are thought of as a black box, given some input, they learn to produce some output. CNN's are actually learning to recognize a variety of spatial patterns and you can visualize what each convolutional layer has been trained to recognize by looking at the weights that make up each convolutional kernel and applying those one at a time to a sample image. This technique is called feature visualization and it's useful for understanding the inner workings of a CNN.
In the cell below, you can see how to extract a single filter (by index) from your first convolutional layer. The filter should appear as a grayscale grid.
```
# Get the weights in the first conv layer, "conv1"
# if necessary, change this to reflect the name of your first conv layer
weights1 = net.conv1.weight.data
w = weights1.cpu().numpy()
filter_index = 0
print(w[filter_index][0])
print(w[filter_index][0].shape)
# display the filter weights
plt.imshow(w[filter_index][0], cmap='gray')
```
## Feature maps
Each CNN has at least one convolutional layer that is composed of stacked filters (also known as convolutional kernels). As a CNN trains, it learns what weights to include in it's convolutional kernels and when these kernels are applied to some input image, they produce a set of **feature maps**. So, feature maps are just sets of filtered images; they are the images produced by applying a convolutional kernel to an input image. These maps show us the features that the different layers of the neural network learn to extract. For example, you might imagine a convolutional kernel that detects the vertical edges of a face or another one that detects the corners of eyes. You can see what kind of features each of these kernels detects by applying them to an image. One such example is shown below; from the way it brings out the lines in an the image, you might characterize this as an edge detection filter.
<img src='images/feature_map_ex.png' width=50% height=50%/>
Next, choose a test image and filter it with one of the convolutional kernels in your trained CNN; look at the filtered output to get an idea what that particular kernel detects.
### TODO: Filter an image to see the effect of a convolutional kernel
---
```
import random
import cv2
##TODO: load in and display any image from the transformed test dataset
test_images, test_outputs, gt_pts = net_sample_output()
idx = random.randint(0, 9)
image = test_images[idx].data # get the image from it's Variable wrapper
image = image.cpu().numpy() # convert to numpy array from a Tensor
image = np.transpose(image, (1, 2, 0)) # transpose to go from torch to numpy image
image = np.squeeze(image)
print(image.shape)
## TODO: Using cv's filter2D function,
## apply a specific set of filter weights (like the one displayed above) to the test image
weights1 = net.conv1.weight.data
w = weights1.cpu().numpy()
image_num = len(w)+1
if image_num > 10:
image_num = 10
# display images
fig, axs = plt.subplots(nrows=image_num, figsize=(15, 15))
axs[0].imshow(image, cmap='gray')
for i in range(image_num-1):
w_filter = w[i][0]
filter_image = cv2.filter2D(image, -1, w_filter)
axs[i+1].imshow(filter_image, cmap='gray')
plt.show()
```
### Question 4: Choose one filter from your trained CNN and apply it to a test image; what purpose do you think it plays? What kind of feature do you think it detects?
**Answer**:
I picked up 5 pictures; the first is original image, the others are filtered images.
First, we can notice that the first filtered image (second image of all) showed that edge detection was performed by first convolution layer. It detects both vertical and horizontal lines, but horizontal lines are more clear than vertical.
The second filtered image is very similar to the third filtered image. It might extract brightness of the original image. Darker pixels seems to indicate how bright original pixels are.
The fourth filtered image is very similar to the original image. The filtered image seems blur, so that noise was removed in the last image. This image was input to next layer, and second layer can absorb the image similar to original image.
---
## Moving on!
Now that you've defined and trained your model (and saved the best model), you are ready to move on to the last notebook, which combines a face detector with your saved model to create a facial keypoint detection system that can predict the keypoints on *any* face in an image!
| github_jupyter |
# Project 1: Trading with Momentum
## Instructions
Each problem consists of a function to implement and instructions on how to implement the function. The parts of the function that need to be implemented are marked with a `# TODO` comment. After implementing the function, run the cell to test it against the unit tests we've provided. For each problem, we provide one or more unit tests from our `project_tests` package. These unit tests won't tell you if your answer is correct, but will warn you of any major errors. Your code will be checked for the correct solution when you submit it to Udacity.
## Packages
When you implement the functions, you'll only need to you use the packages you've used in the classroom, like [Pandas](https://pandas.pydata.org/) and [Numpy](http://www.numpy.org/). These packages will be imported for you. We recommend you don't add any import statements, otherwise the grader might not be able to run your code.
The other packages that we're importing is `helper`, `project_helper`, and `project_tests`. These are custom packages built to help you solve the problems. The `helper` and `project_helper` module contains utility functions and graph functions. The `project_tests` contains the unit tests for all the problems.
### Install Packages
```
import sys
!{sys.executable} -m pip install -r requirements.txt
```
### Load Packages
```
import pandas as pd
import numpy as np
import helper
import project_helper
import project_tests
```
## Market Data
The data source we use for most of the projects is the [Wiki End of Day data](https://www.quandl.com/databases/WIKIP) hosted at [Quandl](https://www.quandl.com). This contains data for many stocks, but we'll just be looking at stocks in the S&P 500. We also made things a little easier to run by narrowing our range of time to limit the size of the data.
### Set API Key
Set the `quandl_api_key` variable to your Quandl api key. You can find your Quandl api key [here](https://www.quandl.com/account/api).
```
# TODO: Add your Quandl API Key
quandl_api_key = ''
```
### Download Data
```
import os
snp500_file_path = 'data/tickers_SnP500.txt'
wiki_file_path = 'data/WIKI_PRICES.csv'
start_date, end_date = '2013-07-01', '2017-06-30'
use_columns = ['date', 'ticker', 'adj_close']
if not os.path.exists(wiki_file_path):
with open(snp500_file_path) as f:
tickers = f.read().split()
helper.download_quandl_dataset(quandl_api_key, 'WIKI', 'PRICES', wiki_file_path, use_columns, tickers, start_date, end_date)
else:
print('Data already downloaded')
```
### Load Data
```
df = pd.read_csv(wiki_file_path, parse_dates=['date'], index_col=False)
close = df.reset_index().pivot(index='date', columns='ticker', values='adj_close')
print('Loaded Data')
```
### View Data
Run the cell below to see what the data looks like for `close`.
```
project_helper.print_dataframe(close)
```
### Stock Example
Let's see what a single stock looks like from the closing prices. For this example and future display examples in this project, we'll use Apple's stock (AAPL). If we tried to graph all the stocks, it would be too much information.
```
apple_ticker = 'AAPL'
project_helper.plot_stock(close[apple_ticker], '{} Stock'.format(apple_ticker))
```
## Resample Adjusted Prices
The trading signal you'll develop in this project does not need to be based on daily prices, for instance, you can use month-end prices to perform trading once a month. To do this, you must first resample the daily adjusted closing prices into monthly buckets, and select the last observation of each month.
Implement the `resample_prices` to resample `close_prices` at the sampling frequency of `freq`.
```
def resample_prices(close_prices, freq='M'):
"""
Resample close prices for each ticker at specified frequency.
Parameters
----------
close_prices : DataFrame
Close prices for each ticker and date
freq : str
What frequency to sample at
For valid freq choices, see http://pandas.pydata.org/pandas-docs/stable/timeseries.html#offset-aliases
Returns
-------
prices_resampled : DataFrame
Resampled prices for each ticker and date
"""
# TODO: Implement Function
return None
project_tests.test_resample_prices(resample_prices)
```
### View Data
Let's apply this function to `close` and view the results.
```
monthly_close = resample_prices(close)
project_helper.plot_resampled_prices(
monthly_close.loc[:, apple_ticker],
close.loc[:, apple_ticker],
'{} Stock - Close Vs Monthly Close'.format(apple_ticker))
```
## Compute Log Returns
Compute log returns ($R_t$) from prices ($P_t$) as your primary momentum indicator:
$$R_t = log_e(P_t) - log_e(P_{t-1})$$
Implement the `compute_log_returns` function below, such that it accepts a dataframe (like one returned by `resample_prices`), and produces a similar dataframe of log returns. Use Numpy's [log function](https://docs.scipy.org/doc/numpy/reference/generated/numpy.log.html) to help you calculate the log returns.
```
def compute_log_returns(prices):
"""
Compute log returns for each ticker.
Parameters
----------
prices : DataFrame
Prices for each ticker and date
Returns
-------
log_returns : DataFrame
Log returns for each ticker and date
"""
# TODO: Implement Function
return None
project_tests.test_compute_log_returns(compute_log_returns)
```
### View Data
Using the same data returned from `resample_prices`, we'll generate the log returns.
```
monthly_close_returns = compute_log_returns(monthly_close)
project_helper.plot_returns(
monthly_close_returns.loc[:, apple_ticker],
'Log Returns of {} Stock (Monthly)'.format(apple_ticker))
```
## Shift Returns
Implement the `shift_returns` function to shift the log returns to the previous or future returns in the time series. For example, the parameter `shift_n` is 2 and `returns` is the following:
```
Returns
A B C D
2013-07-08 0.015 0.082 0.096 0.020 ...
2013-07-09 0.037 0.095 0.027 0.063 ...
2013-07-10 0.094 0.001 0.093 0.019 ...
2013-07-11 0.092 0.057 0.069 0.087 ...
... ... ... ... ...
```
the output of the `shift_returns` function would be:
```
Shift Returns
A B C D
2013-07-08 NaN NaN NaN NaN ...
2013-07-09 NaN NaN NaN NaN ...
2013-07-10 0.015 0.082 0.096 0.020 ...
2013-07-11 0.037 0.095 0.027 0.063 ...
... ... ... ... ...
```
Using the same `returns` data as above, the `shift_returns` function should generate the following with `shift_n` as -2:
```
Shift Returns
A B C D
2013-07-08 0.094 0.001 0.093 0.019 ...
2013-07-09 0.092 0.057 0.069 0.087 ...
... ... ... ... ... ...
... ... ... ... ... ...
... NaN NaN NaN NaN ...
... NaN NaN NaN NaN ...
```
_Note: The "..." represents data points we're not showing._
```
def shift_returns(returns, shift_n):
"""
Generate shifted returns
Parameters
----------
returns : DataFrame
Returns for each ticker and date
shift_n : int
Number of periods to move, can be positive or negative
Returns
-------
shifted_returns : DataFrame
Shifted returns for each ticker and date
"""
# TODO: Implement Function
return None
project_tests.test_shift_returns(shift_returns)
```
### View Data
Let's get the previous month's and next month's returns.
```
prev_returns = shift_returns(monthly_close_returns, 1)
lookahead_returns = shift_returns(monthly_close_returns, -1)
project_helper.plot_shifted_returns(
prev_returns.loc[:, apple_ticker],
monthly_close_returns.loc[:, apple_ticker],
'Previous Returns of {} Stock'.format(apple_ticker))
project_helper.plot_shifted_returns(
lookahead_returns.loc[:, apple_ticker],
monthly_close_returns.loc[:, apple_ticker],
'Lookahead Returns of {} Stock'.format(apple_ticker))
```
## Generate Trading Signal
A trading signal is a sequence of trading actions, or results that can be used to take trading actions. A common form is to produce a "long" and "short" portfolio of stocks on each date (e.g. end of each month, or whatever frequency you desire to trade at). This signal can be interpreted as rebalancing your portfolio on each of those dates, entering long ("buy") and short ("sell") positions as indicated.
Here's a strategy that we will try:
> For each month-end observation period, rank the stocks by _previous_ returns, from the highest to the lowest. Select the top performing stocks for the long portfolio, and the bottom performing stocks for the short portfolio.
Implement the `get_top_n` function to get the top performing stock for each month. Get the top performing stocks from `prev_returns` by assigning them a value of 1. For all other stocks, give them a value of 0. For example, using the following `prev_returns`:
```
Previous Returns
A B C D E F G
2013-07-08 0.015 0.082 0.096 0.020 0.075 0.043 0.074
2013-07-09 0.037 0.095 0.027 0.063 0.024 0.086 0.025
... ... ... ... ... ... ... ...
```
The function `get_top_n` with `top_n` set to 3 should return the following:
```
Previous Returns
A B C D E F G
2013-07-08 0 1 1 0 1 0 0
2013-07-09 0 1 0 1 0 1 0
... ... ... ... ... ... ... ...
```
*Note: You may have to use Panda's [`DataFrame.iterrows`](https://pandas.pydata.org/pandas-docs/version/0.21/generated/pandas.DataFrame.iterrows.html) with [`Series.nlargest`](https://pandas.pydata.org/pandas-docs/version/0.21/generated/pandas.Series.nlargest.html) in order to implement the function. This is one of those cases where creating a vecorization solution is too difficult.*
```
def get_top_n(prev_returns, top_n):
"""
Select the top performing stocks
Parameters
----------
prev_returns : DataFrame
Previous shifted returns for each ticker and date
top_n : int
The number of top performing stocks to get
Returns
-------
top_stocks : DataFrame
Top stocks for each ticker and date marked with a 1
"""
# TODO: Implement Function
return None
project_tests.test_get_top_n(get_top_n)
```
### View Data
We want to get the best performing and worst performing stocks. To get the best performing stocks, we'll use the `get_top_n` function. To get the worst performing stocks, we'll also use the `get_top_n` function. However, we pass in `-1*prev_returns` instead of just `prev_returns`. Multiply by negative one will flip all the positive returns to negative and negative returns to positive. Thus, it returns the top worst performing stocks.
```
top_bottom_n = 50
df_long = get_top_n(prev_returns, top_bottom_n)
df_short = get_top_n(-1*prev_returns, top_bottom_n)
project_helper.print_top(df_long, 'Longed Stocks')
project_helper.print_top(df_short, 'Shorted Stocks')
```
## Projected Returns
It's now time to check if your trading signal has the potential to become profitable!
We'll start by computing the net returns this portfolio would return. For simplicity, we'll assume every stock gets an equal dollar amount of investment. This makes it easier to compute a portfolio's returns as the simple arithmetic average of the individual stock returns.
Implement the `portfolio_returns` function to compute the expected portfolio returns. Using `df_long` to indicate which stocks to long and `df_short` to indicate which stocks to short, calculate the returns using `lookahead_returns`. To help with calculation, we've provided you with `n_stocks` as the number of stocks we're investing in a single period.
```
def portfolio_returns(df_long, df_short, lookahead_returns, n_stocks):
"""
Compute expected returns for the portfolio, assuming equal investment in each long/short stock.
Parameters
----------
df_long : DataFrame
Top stocks for each ticker and date marked with a 1
df_short : DataFrame
Bottom stocks for each ticker and date marked with a 1
lookahead_returns : DataFrame
Lookahead returns for each ticker and date
n_stocks: int
The number number of stocks chosen for each month
Returns
-------
portfolio_returns : DataFrame
Expected portfolio returns for each ticker and date
"""
# TODO: Implement Function
return None
project_tests.test_portfolio_returns(portfolio_returns)
```
### View Data
Time to see how the portfolio did.
```
expected_portfolio_returns = portfolio_returns(df_long, df_short, lookahead_returns, 2*top_bottom_n)
project_helper.plot_returns(expected_portfolio_returns.T.sum(), 'Portfolio Returns')
```
## Statistical Tests
### Annualized Rate of Return
```
expected_portfolio_returns_by_date = expected_portfolio_returns.T.sum().dropna()
portfolio_ret_mean = expected_portfolio_returns_by_date.mean()
portfolio_ret_ste = expected_portfolio_returns_by_date.sem()
portfolio_ret_annual_rate = (np.exp(portfolio_ret_mean * 12) - 1) * 100
print("""
Mean: {:.6f}
Standard Error: {:.6f}
Annualized Rate of Return: {:.2f}%
""".format(portfolio_ret_mean, portfolio_ret_ste, portfolio_ret_annual_rate))
```
Annualized rate of return gives you a sense of what the returns are indicating; it would be naive to assume that rate of return will hold over any significant period of time. A better predictor of your signal's general performance is a T-Test.
### T-Test
Our null hypothesis ($H_0$) is that the expected mean return from the signal is zero, and that any positive mean observed here is a matter of chance (within certain expected bounds of deviation). We'll perform a one-sample, one-sided t-test on the observed net returns, to see if we can reject $H_0$.
We'll need to first compute the t-statistic, and then find its corresponding p-value. The p-value will indicate the probability of observing these net returns if the null hypothesis were true. Therefore, a smaller p-value would indicate that the null hypothesis is less likely. In fact, it's good practice to set a desired level of significance or alpha ($\alpha$) _before_ computing the p-value, and then reject the null hypothesis if $p < \alpha$.
For this project, we'll use $\alpha = 0.05$, since it's a common value to use.
Implement the `analyze_alpha` function to perform a t-test on expected portfolio returns by date. We've imported the `scipy.stats` module for you to perform the t-test.
Note: [`scipy.stats.ttest_1samp`](https://docs.scipy.org/doc/scipy-1.0.0/reference/generated/scipy.stats.ttest_1samp.html) performs a two-sided test, so divide the p-value by 2 to get 1-sided p-value
```
from scipy import stats
def analyze_alpha(expected_portfolio_returns_by_date):
"""
Perform a t-test with the null hypothesis being that the expected mean return is zero.
Parameters
----------
expected_portfolio_returns_by_date : Pandas Series
Expected portfolio returns for each date
Returns
-------
t_value
T-statistic from t-test
p_value
Corresponding p-value
"""
# TODO: Implement Function
return None
project_tests.test_analyze_alpha(analyze_alpha)
```
### View Data
Let's see what values we get without portfolio. After you run this, make sure to answer the question below.
```
t_value, p_value = analyze_alpha(expected_portfolio_returns_by_date)
print("""
Alpha analysis:
t-value: {:.3f}
p-value: {:.6f}
""".format(t_value, p_value))
```
### Question: What p-value did you observe? And what does that indicate about your signal?
**A**: I got a p-value of 0.089. Since it's greater than alpha, which is 0.05, we can't throw out the null hypothesis. This means we're not sure if there's a relationship between our signal and our projected returns.
## Submission
Now that you're done with the project, it's time to submit it. Click the submit button in the bottom right. One of our reviewers will give you feedback on your project with a pass or not passed grade. You can continue to the next section while you wait for feedback.
| github_jupyter |
<a href="https://colab.research.google.com/github/nstaudac/DS-Unit-1-Sprint-2-Data-Wrangling-and-Storytelling/blob/master/Nicholas_Staudacher_LS_DS6_224_Sequence_your_narrative.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
_Lambda School Data Science_
# Sequence your narrative
Today we will create a sequence of visualizations inspired by [Hans Rosling's 200 Countries, 200 Years, 4 Minutes](https://www.youtube.com/watch?v=jbkSRLYSojo).
Using this [data from Gapminder](https://github.com/open-numbers/ddf--gapminder--systema_globalis/):
- [Income Per Person (GDP Per Capital, Inflation Adjusted) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--income_per_person_gdppercapita_ppp_inflation_adjusted--by--geo--time.csv)
- [Life Expectancy (in Years) by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--life_expectancy_years--by--geo--time.csv)
- [Population Totals, by Geo & Time](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv)
- [Entities](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv)
- [Concepts](https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--concepts.csv)
Objectives
- sequence multiple visualizations
- combine qualitative anecdotes with quantitative aggregates
Links
- [Hans Rosling’s TED talks](https://www.ted.com/speakers/hans_rosling)
- [Spiralling global temperatures from 1850-2016](https://twitter.com/ed_hawkins/status/729753441459945474)
- "[The Pudding](https://pudding.cool/) explains ideas debated in culture with visual essays."
- [A Data Point Walks Into a Bar](https://lisacharlotterost.github.io/2016/12/27/datapoint-in-bar/): a thoughtful blog post about emotion and empathy in data storytelling
## Make a plan
#### How to present the data?
Variables --> Visual Encodings
- Income --> x
- Lifespan --> y
- Region --> color
- Population --> size
- Year --> animation frame (alternative: small multiple)
- Country --> annotation
Qualitative --> Verbal
- Editorial / contextual explanation --> audio narration (alternative: text)
#### How to structure the data?
| Year | Country | Region | Income | Lifespan | Population |
|------|---------|----------|--------|----------|------------|
| 1818 | USA | Americas | ### | ## | # |
| 1918 | USA | Americas | #### | ### | ## |
| 2018 | USA | Americas | ##### | ### | ### |
| 1818 | China | Asia | # | # | # |
| 1918 | China | Asia | ## | ## | ### |
| 2018 | China | Asia | ### | ### | ##### |
## More imports
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
```
## Load & look at data
```
income = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--income_per_person_gdppercapita_ppp_inflation_adjusted--by--geo--time.csv')
lifespan = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--life_expectancy_years--by--geo--time.csv')
population = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--datapoints--population_total--by--geo--time.csv')
entities = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--entities--geo--country.csv')
concepts = pd.read_csv('https://raw.githubusercontent.com/open-numbers/ddf--gapminder--systema_globalis/master/ddf--concepts.csv')
income.shape, lifespan.shape, population.shape, entities.shape, concepts.shape
income.head()
lifespan.head()
population.head()
pd.options.display.max_columns = 500
entities.head()
concepts.head()
```
## Merge data
https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf
```
# merge income and lifespan csv files
df = pd.merge(income, lifespan)
df.shape
df = pd.merge(df, population)
df.shape
entities_variables = ['country', 'name', 'world_6region']
entities = entities[entities_variables]
print(entities.shape)
entities.head()
entities['world_6region'].value_counts()
merged = pd.merge(df, entities, left_on='geo', right_on='country')
print(merged.shape)
df.head()
merged = merged.drop(['geo', 'country'], axis='columns')
merged.head()
merged = merged.rename(columns = {
'time': 'year',
'income_per_person_gdppercapita_ppp_inflation_adjusted': 'income',
'life_expectancy_years': 'lifespan',
'population_total': 'population',
'name': 'country',
'world_6region': 'region'
})
merged.head()
merged.head()
```
## Explore data
```
merged.dtypes
merged.describe()
merged.describe(exclude='number')
merged.country.unique()
usa = merged[merged.country=="United States"]
usa.head()
usa[usa.year.isin([1818, 1918, 2018])]
china = merged[merged.country=="China"]
china.head()
china[china.year.isin([1818, 1918, 2018])]
```
## Plot visualization
Changed title on graph, adjusted size of markers on graph, set style to 'whitegrid'
```
import seaborn as sns
year_2018= merged[merged['year'] == 2018]
sns.set_style("whitegrid")
sns.relplot(x='income', y='lifespan', hue='region', size='population',
sizes=(20, 500), data=year_2018)
plt.title("Income Per Person GDP Per Capita PPP Inflation Adjusted")
plt.text(x=40000.0, y=85, s="Income vs Lifespan by Country")
plt.show();
```
## Analyze outliers
```
qatar_year_2018 = year_2018[(year_2018.income > 80000) & (year_2018.country == 'Qatar')].sort_values(by='income')
qatar_year_2018
sns.relplot(x='income', y='lifespan', hue='region', size='population',
sizes=(20, 460), data=year_2018);
plt.text(x=qatar_year_2018.income, y=qatar_year_2018.lifespan + 1, s='Qatar')
plt.title("Income Per Person GDP Per Capita PPP Inflation Adjusted")
plt.text(x=30000.0, y=85, s="2018 Qatar Outlier Displayed")
plt.show();
```
## Plot multiple years
```
years = [1818, 1918, 2018]
centuries = merged[merged.year.isin(years)]
sns.relplot(x='income', y='lifespan', hue='region', size='population',
col='year', data=centuries)
plt.xscale('log');
plt.text(x=qatar_year_2018.income-5000, y=qatar_year_2018.lifespan + 1,
s='Qatar');
```
## Point out a story
```
years = [1918, 1938, 1958, 1978, 1998, 2018]
decades = merged[merged.year.isin(years)]
sns.relplot(x='income', y='lifespan', hue='region', size='population',
sizes=(20, 400),
col='year', data=decades);
for year in years:
sns.relplot(x='income', y='lifespan', hue='region', size='population',
sizes=(20, 600), data=merged[merged.year==year])
plt.xscale('log')
plt.xlim((150, 150000))
plt.ylim((0, 90))
plt.title(year)
plt.axhline(y=50, color='grey')
merged[(merged.year==1918) & (merged.lifespan >50)]
merged[(merged.year==2018) & (merged.lifespan <50)]
year = 1883 #@param {type:"slider", min:1800, max:2018, step:1}
sns.relplot(x='income', y='lifespan', hue='region', size='population',
data=merged[merged.year==year])
plt.xscale('log')
plt.xlim((150, 150000))
plt.ylim((20, 90))
plt.title(year);
```
# ASSIGNMENT
Replicate the lesson code
-improve gapminder graph that we made during the lecture
-change the size of the circles
-add a title and subtitle to graph
-(optional: see how graph looks using 4 regions rather than the 6)
-stretch goal -small tick marks/color options
# STRETCH OPTIONS
## 1. Animate!
- [Making animations work in Google Colaboratory](https://medium.com/lambda-school-machine-learning/making-animations-work-in-google-colaboratory-new-home-for-ml-prototyping-c6147186ae75)
- [How to Create Animated Graphs in Python](https://towardsdatascience.com/how-to-create-animated-graphs-in-python-bb619cc2dec1)
- [The Ultimate Day of Chicago Bikeshare](https://chrisluedtke.github.io/divvy-data.html) (Lambda School Data Science student)
## 2. Work on anything related to your portfolio site / project
| github_jupyter |
```
from IPython.display import IFrame
%matplotlib inline
```
# *Hubble and the origins of DESI*
The year 1929 brought us the Oscars, the first car radio and Edwin Hubble's unexpected observation that all galaxies are moving away from us!

Let's take a quick look at some of the galaxies he was looking at, Triangulum and the Large Magellanic Cloud.


In total, Edwin studied the distances of 24 galaxies from us, and their observed `redshifts'. What does that mean?
Maybe you already know that the energy levels of Hydrogen are __quantized__, with electrons habitating a series of shells with __discrete__ energies. When an electron transitions between any two levels, light is emitted with a wavelength neatly given by the "Rydberg" formula:
$$
\frac{1}{\lambda_{\rm vac}} = 1.096 \times 10^{7} \left ( \frac{1}{n^2} - \frac{1}{m^2} \right )
$$
where $n$ and $m$ (any one of $[0, 1, 2, ... \infty]$) label the two energy levels.
```
# First, let's import some useful packages:
import os
import astropy
import pylab as pl
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from scipy import stats
from IPython.display import Image
from tools.wave2rgb import wavelength_to_rgb
from pkg_resources import resource_filename
def Rydberg(n, m):
# Vacuum wavelengths [nanometres]
result = 1.096e-2 * (1. / n / n - 1. / m / m)
return 1. / result
```
Let's workout the wavelengths of light that Hydrogen can emit:
```
waves = []
print('n \t m \t Wavelength [nm]')
for n in np.arange(1, 10, 1):
for m in np.arange(n+1, 10, 1):
wave = Rydberg(n, m)
waves.append(wave)
print('{:d} \t {:d} \t {:.3f}'.format(n, m, wave))
```
Now let's plot the wavelengths and see the color of these lines. If we were to look at a emitting Hydrogen atom, you'd see this:
```
for wave in waves:
# color = [r, g, b]
color = wavelength_to_rgb(wave)
pl.axvline(x=wave, c=color)
pl.xlabel('Vacuum wavelength [nanometers]')
pl.xlim(380., 780.)
```
If the hydrogen exists in a galaxy that is moving, we see the lines Doppler shifted. We will call this the "redshift" of the galaxy, often denote as $z$ (https://en.wikipedia.org/wiki/Redshift). Let's say the galaxy is moving at 1% the speed of light (v = 0.1*c), we can calculate the redshift with the following equation:
$$
1 + z = \sqrt{\frac{1 + v/c}{1 + v/c}}
$$
```
def redshift(v):
# v [speed of light].
result = (1. + v) / (1. - v)
result = np.sqrt(result) - 1.
return result
zz = redshift(0.01)
for restwave in waves:
obswave = (1. + zz) * restwave
color = wavelength_to_rgb(restwave)
pl.axvline(x=restwave, c=color, alpha=0.25)
color = wavelength_to_rgb(obswave)
pl.axvline(x=obswave, c=color)
pl.xlabel('Vacuum wavelength [nanometers]')
pl.xlim(380., 780.)
```
Here you see the original line (faint) and the line shifted if the galaxy with the emitting Hydrogen is moving. https://en.wikipedia.org/wiki/Doppler_effect will tell you all the details.
Hubble knew the lines of Hydrogen, and for many other elements. By reversing above, he was able to calculate the velocity for many galaxies. He found out how far away there were (from how bright some special stars in the galaxy were - https://en.wikipedia.org/wiki/Cepheid_variable) and how fast they were moving (from their redshift, as above):
```
hub = resource_filename('desihigh','dat/hubble.dat')
dat = pd.read_csv(hub, sep='\s+', comment='#', names=['Galaxy name', 'Distance [Mpc]', 'Velocity [km/s]'])
dat
```
Let's plot them.
```
fig = plt.figure(figsize=(10, 7.5))
ax = fig.add_subplot(1, 1, 1)
plt.close()
label_style = {'fontname': 'Georgia', 'fontsize': 16}
ax.plot(dat['Distance [Mpc]'], dat['Velocity [km/s]'], '-', c='k', marker='*', lw=0)
ax.set_xlabel('Distance from us [Megaparsecs]', **label_style)
ax.set_ylabel('Recession velocity [km/s]', **label_style)
plt.tight_layout()
fig
```
Edwin saw a clear trend, but the measurements seemed pretty noisy. Let's figure out our best guess at the true relationship between the two. We'll look at a linear relationship (regression) using the scipy stats package:
```
slope, intercept, r_value, p_value, std_err = stats.linregress(dat['Distance [Mpc]'],dat['Velocity [km/s]'])
print('The gradient to this trend is known as the Hubble constant: {:.3f} [km/s/Mpc]'.format(slope))
```
Let's see what that looks like.
```
distances = np.linspace(-0.5, 2.5, 10)
velocities = slope * distances
ax.plot(distances, velocities, lw=0.25, c='k')
ax.set_xlim(0.0, 2.5)
fig
```
Seems a pretty good fit!
Now it's your turn, can you figure out a good estimate of the error on this measurement of the Hubble costant. How accurately can we predict the recession of a galaxy at a given distance, i.e. how fast or slow could it be moving?
So in conclusion, every galaxy is likely to be moving away from us! We find this to be true of all galaxies - we are not at center or special in any way. Every galaxy is moving away from every other. The fact that the Universe was expanding came as a shock to many in 1929, but an even greater surprise was in store.
# *Dark Energy*
In 1998, the world would change forever. Larry Page and Sergey Brin founded Google, the American Unity node and Russian Zarya module would be brought together to form the [International Space Station](https://en.wikipedia.org/wiki/International_Space_Station), and Lawrence Berkeley Lab's
very own Saul Perlmutter, Brian Schmidt and Adam Reiss irrefutably confirmed the existence of _Dark Energy_. Here's Saul impressing some young Berkeley researchers with these results at the time:

So what was everyone looking at? Let's breakdown the data.
Saul and his team measured the redshift ($z$) and the effective magnitude for several Type Ia Supernovae (https://en.wikipedia.org/wiki/Type_Ia_supernova)
```
perl = resource_filename('desihigh', 'dat/perlmutter.txt')
dat = pd.read_csv(perl, names=['z', 'Effective magnitude'], comment='#', sep='\s+')
toprint = dat[:10]
toprint
```
A plot would show this a lot more clearly:
```
pl.plot(dat['z'], dat['Effective magnitude'], marker='.', lw=0.0)
pl.xlabel('z')
pl.ylabel('Effective magnitude')
```
Saul has good reason to believe (really, he had to tweak them a bit first) that every [type Ia supernovae](https://en.wikipedia.org/wiki/Type_Ia_supernova) shown here was equally bright intrinsically, but those at high redshift appeared relatively faint compared to those at low redshift, as they were simply further away. This explains the trend shown, given that 'effective magnitude' is the awkward way in which astronomers typically express how bright something appears.
The useful thing about this measurement is that how far away a supernovae or galaxy is for a given redshift depends on a few parameters, one of which is how much Dark Energy there might be in the Universe. Almost everyone expected this data to prove there was _no_ _Dark Energy_ when Saul made it, but a few guessed otherwise.
When Hubble discovered the expansion, a natural consequence was that the amount of (rest mass) energy contained within a cubic meter would dilute with time. Dark Energy would be special, as the amount of energy per cubic meter would instead be constant with time and would suggest that spooky effects of [quantum mechanics](https://en.wikipedia.org/wiki/Quantum_mechanics) would be causing the galaxies to separate.
So let's use Saul's data to figure out how much Dark Energy is in the Universe. First, we need a model for the (luminosity) distance of a supernovae at a given redshift, given some amount of Dark Energy. We use $\Omega_\Lambda$ to denote the _fraction_ of all matter that behaves like Dark Energy.
```
from astropy.cosmology import FlatLambdaCDM
def lumdist(z, olambda):
cosmo = FlatLambdaCDM(H0=70, Om0=1. - olambda, Tcmb0=2.725)
return cosmo.luminosity_distance(z)
```
We then need to convert this distance into how astronomers measure brightness:
```
def effmag(z, olambda, MB):
DL = lumdist(z, olambda)
return MB + 5. * np.log10(DL.value)
zs = np.arange(0.01, 0.85, 0.01)
pl.plot(dat['z'], dat['Effective magnitude'], marker='.', lw=0.0)
pl.plot(zs, effmag(zs, 0.0, 6.), c='k', label='No Dark Energy', alpha=0.5)
pl.plot(zs, effmag(zs, 0.5, 6.), c='k', label='Dark Energy!')
pl.xlabel('z')
pl.ylabel('Effective magnitude')
pl.legend(loc=4, frameon=False)
```
Even by eye, the data looks to prefer some Dark Energy. But there's not a huge amount in it. Let's figure out what exactly the data prefers. To do this, we'll assume that minimising the distance between each point and the line is the best measure of how well the theory fits the data (see https://en.wikipedia.org/wiki/Least_squares). Together with the fraction of Dark Energy, we also don't know how bright every supernovae is intrinsically so we'll fit for that simultaneously.
```
from scipy.optimize import minimize
def chi2(x):
olambda = x[0]
MB = x[1]
model = effmag(dat['z'], olambda, MB)
return np.sum((dat['Effective magnitude'] - model)**2.)
res = minimize(chi2, x0=[0.5, 5.0], options={'disp': True})
res.x
zs = np.arange(0.01, 0.85, 0.01)
pl.plot(dat['z'], dat['Effective magnitude'], marker='.', lw=0.0)
pl.plot(zs, effmag(zs, 0.0, 6.), c='k', label='No Dark Energy', alpha=0.5)
pl.plot(zs, effmag(zs, 0.5, 6.), c='k', label='50% Dark Energy!')
pl.plot(zs, effmag(zs, 0.75, 6.), c='c', label='75% Dark Energy!')
pl.xlabel('z')
pl.ylabel('Effective magnitude')
pl.legend(loc=4, frameon=False)
```
So there's something like 75% dark energy in the Universe! As the first people to make this measurement, Saul, together with Brian Schmidt and Adam Reiss, would be awarded the 2011 Nobel Prize for their work.

You can find all the details of his work here: https://arxiv.org/pdf/astro-ph/9812133.pdf. Warning, this is for the pros so don't worry if you don't understand too much!
As honorary principal at DESI High, Saul has a special opening address to all the students, including you!
```
perl = resource_filename('desihigh', 'perlmutter/letter.pdf')
perl = os.path.relpath(perl)
IFrame(perl, width=900, height=1000)
```
The primary motivation for DESI is to repeat similar distant-redshift measurements much more precisely and learn much more about this spooky Dark Energy!
| github_jupyter |
[View in Colaboratory](https://colab.research.google.com/github/ufrpe-eagri-ic/aulas/blob/master/08_Funcoes.ipynb)
# Funções
Funções nos permitem escrever código que podemos usar no futuro. Quando colocamos uma série de instruções em uma função, podemos reutilizá-lo para receber entradas, realizar cálculos ou outras manipulações e retornar saídas, assim como uma função em matemática.
É como se fosse um bloco de código reutilizavel
Como criar uma função:
```
def nome_da_funcao(argumento01, argumento02, ...):
# Código da função
return alguma_coisa
```
O **def** declara que estaremos construindo uma função na qual temos o nome da função e seus argumentos que serão passados para função
O **return** é opcional caso sua função retorne algo, ela pode somente imprimir algo não precisando do return, as funções podem retornar vários valores de uma vez, retornando uma **tupla** ou valores unicos se atribuido para a mesma quantidade de variaveis que retornou
As funções devem ser definidas antes de ser executadas
Usar a função:
```
alguma_coisa = nome_da_funcao(argumento01, argumento02)
```
Nomeclaturas: Não de o nome da função para nome de variaveis
**Exemplo**
Função chamada **maior_numero** que recebe dois numeros, e imprime o maior deles.
```
def maior_numero(num1, num2):
maior = num1
if num2 > num1:
maior = num2
print('O número ', maior, ' é o maior')
```
para usar a função:
```
maior_numero(2, 10)
# usando quantas vezes quiser
maior_numero(10,4)
maior_numero(20, 16)
```
**Exercício**
Escreva uma função que receba uma lista e imprima apenas os números pares
```
```
## Retornando valores
Para retornar um valor, utilizar a palavra **return**
### Exemplo:
Função que retorna True caso o número informado seja ímpar e False caso contrário
```
def eh_impar(x):
if x%2 == 0:
return False
else:
return True
print(eh_impar(4))
print(eh_impar(1))
print(eh_impar(3))
```
# Retornando mais de um valor
```
# Define a função de operações matematicas e retorna duas operações
def soma_multiplica(number1, number2):
return number1+number2, number1*number2
print(soma_multiplica(3, 4))
print(soma_multiplica(2, 5))
```
# Exercício
Escreva uma função em python chamada ‘histograma’que receba como entrada uma string, e retorne um dicionário cujas chaves correspondem à letras da string de entrada, e os valores correspondam à quantidade de vezes que cada letra se repete na string. Por exemplo:
```
>>> h = histograma('brontosaurus')
>>> h
{'a': 1, 'b': 1, 'o': 2, 'n': 1, 's': 2, 'r': 2, 'u': 2, 't': 1}
```
```
```
| github_jupyter |
# Your first neural network
In this project, you'll build your first neural network and use it to predict daily bike rental ridership. We've provided some of the code, but left the implementation of the neural network up to you (for the most part). After you've submitted this project, feel free to explore the data and the model more.
```
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
## Load and prepare the data
A critical step in working with neural networks is preparing the data correctly. Variables on different scales make it difficult for the network to efficiently learn the correct weights. Below, we've written the code to load and prepare the data. You'll learn more about this soon!
```
data_path = 'Bike-Sharing-Dataset/hour.csv'
rides = pd.read_csv(data_path)
rides.head()
```
## Checking out the data
This dataset has the number of riders for each hour of each day from January 1 2011 to December 31 2012. The number of riders is split between casual and registered, summed up in the `cnt` column. You can see the first few rows of the data above.
Below is a plot showing the number of bike riders over the first 10 days or so in the data set. (Some days don't have exactly 24 entries in the data set, so it's not exactly 10 days.) You can see the hourly rentals here. This data is pretty complicated! The weekends have lower over all ridership and there are spikes when people are biking to and from work during the week. Looking at the data above, we also have information about temperature, humidity, and windspeed, all of these likely affecting the number of riders. You'll be trying to capture all this with your model.
```
rides[:24*10].plot(x='dteday', y='cnt')
```
### Dummy variables
Here we have some categorical variables like season, weather, month. To include these in our model, we'll need to make binary dummy variables. This is simple to do with Pandas thanks to `get_dummies()`.
```
dummy_fields = ['season', 'weathersit', 'mnth', 'hr', 'weekday']
for each in dummy_fields:
dummies = pd.get_dummies(rides[each], prefix=each, drop_first=False)
rides = pd.concat([rides, dummies], axis=1)
fields_to_drop = ['instant', 'dteday', 'season', 'weathersit',
'weekday', 'atemp', 'mnth', 'workingday', 'hr']
data = rides.drop(fields_to_drop, axis=1)
data.head()
```
### Scaling target variables
To make training the network easier, we'll standardize each of the continuous variables. That is, we'll shift and scale the variables such that they have zero mean and a standard deviation of 1.
The scaling factors are saved so we can go backwards when we use the network for predictions.
```
quant_features = ['casual', 'registered', 'cnt', 'temp', 'hum', 'windspeed']
# Store scalings in a dictionary so we can convert back later
scaled_features = {}
for each in quant_features:
mean, std = data[each].mean(), data[each].std()
scaled_features[each] = [mean, std]
data.loc[:, each] = (data[each] - mean)/std
data.head()
```
### Splitting the data into training, testing, and validation sets
We'll save the data for the last approximately 21 days to use as a test set after we've trained the network. We'll use this set to make predictions and compare them with the actual number of riders.
```
# Save data for approximately the last 21 days
test_data = data[-21*24:]
# Now remove the test data from the data set
data = data[:-21*24]
print("test: {}, data: {}".format(test_data.shape, data.shape))
# Separate the data into features and targets
target_fields = ['cnt', 'casual', 'registered']
features, targets = data.drop(target_fields, axis=1), data[target_fields]
test_features, test_targets = test_data.drop(target_fields, axis=1), test_data[target_fields]
```
We'll split the data into two sets, one for training and one for validating as the network is being trained. Since this is time series data, we'll train on historical data, then try to predict on future data (the validation set).
```
# Hold out the last 60 days or so of the remaining data as a validation set
train_features, train_targets = features[:-60*24], targets[:-60*24]
val_features, val_targets = features[-60*24:], targets[-60*24:]
print("train_features: {} ({}), train_targets: {} ({})".format(len(train_features), train_features.shape, len(train_targets), train_targets.shape))
```
## Time to build the network
Below you'll build your network. We've built out the structure and the backwards pass. You'll implement the forward pass through the network. You'll also set the hyperparameters: the learning rate, the number of hidden units, and the number of training passes.
<img src="assets/neural_network.png" width=300px>
The network has two layers, a hidden layer and an output layer. The hidden layer will use the sigmoid function for activations. The output layer has only one node and is used for the regression, the output of the node is the same as the input of the node. That is, the activation function is $f(x)=x$. A function that takes the input signal and generates an output signal, but takes into account the threshold, is called an activation function. We work through each layer of our network calculating the outputs for each neuron. All of the outputs from one layer become inputs to the neurons on the next layer. This process is called *forward propagation*.
We use the weights to propagate signals forward from the input to the output layers in a neural network. We use the weights to also propagate error backwards from the output back into the network to update our weights. This is called *backpropagation*.
> **Hint:** You'll need the derivative of the output activation function ($f(x) = x$) for the backpropagation implementation. If you aren't familiar with calculus, this function is equivalent to the equation $y = x$. What is the slope of that equation? That is the derivative of $f(x)$.
Below, you have these tasks:
1. Implement the sigmoid function to use as the activation function. Set `self.activation_function` in `__init__` to your sigmoid function.
2. Implement the forward pass in the `train` method.
3. Implement the backpropagation algorithm in the `train` method, including calculating the output error.
4. Implement the forward pass in the `run` method.
```
# forward . backprop _
# in1 \ . /
# \_ () \ . / () _/
# in2 / > () output . error () err_grad_output < \_
# \_ () / ^ . \ () _/
# in3 / ^ final_out . \
# ^ final_in . ^ \_
# ^ hidden_outputs . err_grad_hidden
# ^ .
# hidden_inputs .
#
# () = node with f(h) = sigmoid activation function
class NeuralNetwork(object):
def __init__(self, input_nodes, hidden_nodes, output_nodes, learning_rate):
# Set number of nodes in input, hidden and output layers.
self.input_nodes = input_nodes
self.hidden_nodes = hidden_nodes
self.output_nodes = output_nodes
# Initialize weights
self.weights_input_to_hidden = np.random.normal(0.0, self.input_nodes**-0.5,
(self.input_nodes, self.hidden_nodes))
self.weights_hidden_to_output = np.random.normal(0.0, self.hidden_nodes**-0.5,
(self.hidden_nodes, self.output_nodes))
self.lr = learning_rate
self.activation_function = lambda x : 1 / (1 + np.exp(-x))
def train(self, features, targets):
''' Train the network on batch of features and targets.
Arguments
---------
features: 2D array, each row is one data record, each column is a feature
targets: 1D array of target values
'''
n_records = features.shape[0]
delta_weights_i_h = np.zeros(self.weights_input_to_hidden.shape)
delta_weights_h_o = np.zeros(self.weights_hidden_to_output.shape)
for X, y in zip(features, targets):
# X: (3,)
### Forward pass ###
hidden_inputs = np.dot(X, self.weights_input_to_hidden)
hidden_outputs = self.activation_function(hidden_inputs)
final_inputs = np.dot(hidden_outputs, self.weights_hidden_to_output)
final_outputs = final_inputs
### Backward pass ###
error = y - final_outputs
# error gradient output layer
err_grad_output = error * 1 # f'(x) = 1
# Weight step (hidden to output)
#err_grad_o: (1,), Who: (2, 1), hidden_outputs: (2,)
delta_weights_h_o += hidden_outputs[:,None] * err_grad_output # (2,1)
# error gradient hidden layer
#err_grad_o: (1,), Who: (2, 1), h_o: (2,)
err_grad_hidden = np.matmul(err_grad_output, self.weights_hidden_to_output.T) * hidden_outputs * (1 - hidden_outputs) # err_grad_output is scalar, (1,2)
# Weight step (input to hidden)
#delta_weights_i_h: (3, 2), X: (3,) (X[:, None](3, 1)), err_grad_hidden: (1, 2)
delta_weights_i_h += X[:,None] * err_grad_hidden
# TODO: Update the weights - Replace these values with your calculations.
self.weights_hidden_to_output += delta_weights_h_o * self.lr / n_records # update hidden-to-output weights with gradient descent step
self.weights_input_to_hidden += delta_weights_i_h * self.lr / n_records # update input-to-hidden weights with gradient descent step
def run(self, features):
''' Run a forward pass through the network with input features
Arguments
---------
features: 1D array of feature values
'''
#### Implement the forward pass here ####
hidden_inputs = np.dot(features, self.weights_input_to_hidden)
hidden_outputs = self.activation_function(hidden_inputs)
final_inputs = np.dot(hidden_outputs, self.weights_hidden_to_output)
final_outputs = final_inputs
return final_outputs
def MSE(y, Y):
return np.mean((y-Y)**2)
```
## Unit tests
Run these unit tests to check the correctness of your network implementation. This will help you be sure your network was implemented correctly before you starting trying to train it. These tests must all be successful to pass the project.
```
import unittest
inputs = np.array([[0.5, -0.2, 0.1]])
targets = np.array([[0.4]])
test_w_i_h = np.array([[0.1, -0.2],
[0.4, 0.5],
[-0.3, 0.2]])
test_w_h_o = np.array([[0.3],
[-0.1]])
class TestMethods(unittest.TestCase):
##########
# Unit tests for data loading
##########
def test_data_path(self):
# Test that file path to dataset has been unaltered
self.assertTrue(data_path.lower() == 'bike-sharing-dataset/hour.csv')
def test_data_loaded(self):
# Test that data frame loaded
self.assertTrue(isinstance(rides, pd.DataFrame))
##########
# Unit tests for network functionality
##########
def test_activation(self):
network = NeuralNetwork(3, 2, 1, 0.5)
# Test that the activation function is a sigmoid
self.assertTrue(np.all(network.activation_function(0.5) == 1/(1+np.exp(-0.5))))
def test_train(self):
# Test that weights are updated correctly on training
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
network.train(inputs, targets)
self.assertTrue(np.allclose(network.weights_hidden_to_output,
np.array([[ 0.37275328],
[-0.03172939]])))
self.assertTrue(np.allclose(network.weights_input_to_hidden,
np.array([[ 0.10562014, -0.20185996],
[0.39775194, 0.50074398],
[-0.29887597, 0.19962801]])))
def test_run(self):
# Test correctness of run method
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
print("test run:", network.run(inputs))
self.assertTrue(np.allclose(network.run(inputs), 0.09998924))
suite = unittest.TestLoader().loadTestsFromModule(TestMethods())
unittest.TextTestRunner().run(suite)
```
## Training the network
Here you'll set the hyperparameters for the network. The strategy here is to find hyperparameters such that the error on the training set is low, but you're not overfitting to the data. If you train the network too long or have too many hidden nodes, it can become overly specific to the training set and will fail to generalize to the validation set. That is, the loss on the validation set will start increasing as the training set loss drops.
You'll also be using a method know as Stochastic Gradient Descent (SGD) to train the network. The idea is that for each training pass, you grab a random sample of the data instead of using the whole data set. You use many more training passes than with normal gradient descent, but each pass is much faster. This ends up training the network more efficiently. You'll learn more about SGD later.
### Choose the number of iterations
This is the number of batches of samples from the training data we'll use to train the network. The more iterations you use, the better the model will fit the data. However, if you use too many iterations, then the model with not generalize well to other data, this is called overfitting. You want to find a number here where the network has a low training loss, and the validation loss is at a minimum. As you start overfitting, you'll see the training loss continue to decrease while the validation loss starts to increase.
### Choose the learning rate
This scales the size of weight updates. If this is too big, the weights tend to explode and the network fails to fit the data. A good choice to start at is 0.1. If the network has problems fitting the data, try reducing the learning rate. Note that the lower the learning rate, the smaller the steps are in the weight updates and the longer it takes for the neural network to converge.
### Choose the number of hidden nodes
The more hidden nodes you have, the more accurate predictions the model will make. Try a few different numbers and see how it affects the performance. You can look at the losses dictionary for a metric of the network performance. If the number of hidden units is too low, then the model won't have enough space to learn and if it is too high there are too many options for the direction that the learning can take. The trick here is to find the right balance in number of hidden units you choose.
```
import sys
### Set the hyperparameters here ###
iterations = 7000
learning_rate = 0.3
hidden_nodes = 9
output_nodes = 1
N_i = train_features.shape[1]
network = NeuralNetwork(N_i, hidden_nodes, output_nodes, learning_rate)
losses = {'train':[], 'validation':[]}
for ii in range(iterations):
# Go through a random batch of 128 records from the training data set
batch = np.random.choice(train_features.index, size=128)
X, y = train_features.ix[batch].values, train_targets.ix[batch]['cnt']
network.train(X, y)
# Printing out the training progress
train_loss = MSE(network.run(train_features).T, train_targets['cnt'].values)
val_loss = MSE(network.run(val_features).T, val_targets['cnt'].values)
sys.stdout.write("\rProgress: {:2.1f}".format(100 * ii/float(iterations)) \
+ "% ... Training loss: " + str(train_loss)[:5] \
+ " ... Validation loss: " + str(val_loss)[:5])
sys.stdout.flush()
losses['train'].append(train_loss)
losses['validation'].append(val_loss)
plt.plot(losses['train'], label='Training loss')
plt.plot(losses['validation'], label='Validation loss')
plt.legend()
_ = plt.ylim()
```
## Check out your predictions
Here, use the test data to view how well your network is modeling the data. If something is completely wrong here, make sure each step in your network is implemented correctly.
```
fig, ax = plt.subplots(figsize=(8,4))
mean, std = scaled_features['cnt']
predictions = network.run(test_features).T*std + mean
ax.plot(predictions[0], label='Prediction')
ax.plot((test_targets['cnt']*std + mean).values, label='Data')
ax.set_xlim(right=len(predictions))
ax.legend()
dates = pd.to_datetime(rides.ix[test_data.index]['dteday'])
dates = dates.apply(lambda d: d.strftime('%b %d'))
ax.set_xticks(np.arange(len(dates))[12::24])
_ = ax.set_xticklabels(dates[12::24], rotation=45)
```
## OPTIONAL: Thinking about your results(this question will not be evaluated in the rubric).
Answer these questions about your results. How well does the model predict the data? Where does it fail? Why does it fail where it does?
* has offset from the minimum values
* does not adapt to changed patterns during holidays
* possibly provide public holiday/typical vacatino periods as dummy variable
| github_jupyter |
# 特徴抽出を改良による極性判定の精度向上確認
scikit-learnを用いて自然言語の極性判定を実装します。また、Bag of Wordsにおける特徴の表現方法によって正解率が変化することを確認します。
今回はテストデータは用いず、10分割交差検証による正解率の平均を確認します。
## データ準備
今回用いるデータセット「MovieReview」の取得です。パッケージを用いてデータセットをダウンロードします。
取得できるデータの種類についての説明と提供元のURLが確認できます。
```
import chazutsu
chazutsu.datasets.MovieReview.polarity().show()
```
ダウンロードされたデータはローカルに展開されると同時に、
シャッフル・訓練データとテストデータの分割されます。
また、pandas形式で読込を行います。
```
r = chazutsu.datasets.MovieReview.polarity().download(force=True, test_size=0.0)
```
ダウンロードされたデータを先頭から5件確認します。
レビュー内容が「review」、極性が「polarity」として「0(ネガティブ)」、「1(ポジティブ)」として格納されています。
```
r.train_data().head(5)
```
## モデル作成
特徴抽出とモデル選択をパイプラインとして作成します。
今回はCountVectorizerとでBag of Words形式に変換します。(TfidfTransformerでtfidf重み付けも行っています)
単語の数え方における「単語まとまりの単位(1単語 or 2単語)」、「数え方の表現(N回 or 出現有無)」を組み合わせて4パターンのモデルを用意し、精度を比較します。
## 1単語・N回モデル
文書の特徴量として単語まとまりの単位は1単語(unigram)、単語の出現回数をN回としてカウントするBoWを用いるモデル。
```
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer
from sklearn.naive_bayes import MultinomialNB
from sklearn.pipeline import Pipeline
def build_pipeline_unigram_multicount():
text_clf = Pipeline([('vect', CountVectorizer(token_pattern=r'[A-Za-z_]+')),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
return text_clf
text_clf_unigram_multicount = build_pipeline_unigram_multicount()
```
## 1単語・出現有無モデル
文書の特徴量として単語まとまりの単位は1単語(unigram)、単語の出現回数を出現有無としてカウントするBoWを用いるモデル。
```
def build_pipeline_unigram_binarycount():
text_clf = Pipeline([('vect', CountVectorizer(binary=True, token_pattern=r'[A-Za-z_]+'),),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
return text_clf
text_clf_unigram_binarycount = build_pipeline_unigram_binarycount()
```
## 2単語・N回モデル
文書の特徴量として単語まとまりの単位は2単語(unigram)、単語の出現回数をN回としてカウントするBoWを用いるモデル。
```
def build_pipeline_bigram_multicount():
text_clf = Pipeline([('vect', CountVectorizer(ngram_range=(2,2), token_pattern=r'[A-Za-z_]+'),),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
return text_clf
text_clf_bigram_multicount = build_pipeline_bigram_multicount()
```
## 2単語・出現有無モデル
文書の特徴量として単語まとまりの単位は2単語(unigram)、単語の出現回数を出現有無としてカウントするBoWを用いるモデル。
```
def build_pipeline_bigram_binarycount():
text_clf = Pipeline([('vect', CountVectorizer(ngram_range=(2,2), binary=True,token_pattern=r'[A-Za-z_]+'),),
('tfidf', TfidfTransformer()),
('clf', MultinomialNB()),
])
return text_clf
text_clf_bigram_binarycount = build_pipeline_bigram_binarycount()
```
# Cross Validationによる正解率確認
作成した4つのモデルの正解率を10分割交差検証で確認します。
## 1単語・N回モデル
```
import pandas as pd
from sklearn.model_selection import cross_val_score
scores = cross_val_score(text_clf_unigram_multicount, r.train_data().review, r.train_data().polarity, cv=10)
scores
print("Accuracy: %0.3f (+/- %0.3f)" % (scores.mean(), scores.std() * 2))
```
## 1単語・出現有無モデル
```
scores = cross_val_score(text_clf_unigram_binarycount, r.train_data().review, r.train_data().polarity, cv=10)
scores
print("Accuracy: %0.3f (+/- %0.3f)" % (scores.mean(), scores.std() * 2))
```
## 2単語・N回モデル
```
scores = cross_val_score(text_clf_bigram_multicount, r.train_data().review, r.train_data().polarity, cv=10)
scores
print("Accuracy: %0.3f (+/- %0.3f)" % (scores.mean(), scores.std() * 2))
```
## 2単語・出現有無モデル
```
scores = cross_val_score(text_clf_bigram_binarycount, r.train_data().review, r.train_data().polarity, cv=10)
scores
print("Accuracy: %0.3f (+/- %0.3f)" % (scores.mean(), scores.std() * 2))
```
| github_jupyter |
# Day 2 - Intcode interpreter
* https://adventofcode.com/2019/day/2
We have a computer again! We've seen this before in 2017 ([day 18](../2017/Day%2018.ipynb), [day 23](../2017/Day%2023.ipynb)), and 2018 ([day 16](../2018/Day%2016.ipynb), [day 19](../2018/Day%2019.ipynb) and [day 21](../2018/Day%2021.ipynb)).
Now we have opcodes with a variable number of operands (called *positions* here); `1` and `2` each have 2 operands and output destination, 99 has none. There are also no registers, all operations take place directly on the memory where our code is also stored, so it can self-modify. Fun!
So we need a CPU with a position counter, memory, and opcode definitions (*instructions*) to call, and the opcodes need access to the memory (to read operand values and write out their result to). Easy peasy.
I'm assuming we'll expand on the instruction set later on, and that we might have instructions with different numbers of operands. So given a function to process the input values and the number of *paramaters* to process, we should be able to produce something readable and reusable.
```
import aocd
data = aocd.get_data(day=2, year=2019)
memory = list(map(int, data.split(',')))
from __future__ import annotations
import operator
from dataclasses import dataclass
from typing import Callable, List, Mapping, Optional
Memory = List[int]
class Halt(Exception):
"""Signal to end the program"""
@classmethod
def halt(cls) -> int: # yes, because Opcode.f callables always produce ints, right?
raise cls
@dataclass
class Instruction:
# the inputs are processed by a function that operates on integers
# returns integers to store in a destination position
f: Callable[..., int]
# An opcode takes N paramaters
paramater_count: int
def __call__(self, memory: Memory, *parameters: int) -> None:
if parameters:
*inputs, output = parameters
memory[output] = self.f(*(memory[addr] for addr in inputs))
else:
# no parameter count, so just call the function directly, no output expected
self.f()
class CPU:
memory: Memory
pos: int
opcodes: Mapping[int, Instruction] = {
1: Instruction(operator.add, 3),
2: Instruction(operator.mul, 3),
99: Instruction(Halt.halt, 0),
}
def reset(self, memory: Memory = None):
if memory is None:
memory = []
self.memory = memory[:]
self.pos: int = 0
def execute(
self, memory: Memory,
noun: Optional[int] = None,
verb: Optional[int] = None
) -> int:
self.reset(memory)
memory = self.memory
if noun is not None:
memory[1] = noun
if verb is not None:
memory[2] = verb
try:
while True:
op = self.opcodes[memory[self.pos]]
paramcount = op.paramater_count
parameters = memory[self.pos + 1 : self.pos + 1 + paramcount]
op(memory, *parameters)
self.pos += 1 + paramcount
except Halt:
return memory[0]
test: Memory = [1, 9, 10, 3, 2, 3, 11, 0, 99, 30, 40, 50]
cpu = CPU()
assert cpu.execute(test) == 3500
print('Part 1:', cpu.execute(memory, 12, 2))
```
## Part 2
Now we need to find the noun and verb that produce a specific programme output. The text suggests we should just brute-force this, so lets try that first and see how long that takes. Given that we'll only have to search through 10000 different inputs, and there are no options to loop, that's not that big a search space anyway.
While the code can self-modify, this is limited to:
- altering what inputs are read
- where to write the output
- replacing a read or write op with another read, write or halt op
so we execute, at most, `len(memory) // 4` instructions, which for my input means there are only 32 steps per execution run, and so we are going to execute, at most, 32.000 instructions. That's pretty cheap:
```
from itertools import product
def bruteforce(target: int, memory: Memory) -> int:
cpu = CPU()
for noun, verb in product(range(100), repeat=2):
result = cpu.execute(memory, noun, verb)
if result == target:
break
return 100 * noun + verb
print('Part 2:', bruteforce(19690720, memory))
```
## Avoiding brute force
Can we just calculate the number? We'd have to disassemble the inputs to see what is going on. Provided the programme never alters its own instructions, we should be able to figure this out.
Lets see if we need to worry about self-modifying code first:
```
# code is expected to alter memory[0], so we don't count that
# as self modifying as the CPU will never return there.
if any(target and target % 4 == 0 for target in memory[3::4]):
print('Code targets opcodes')
elif any(target % 4 and target > (i * 4 + 3) for i, target in enumerate(memory[3::4])):
print('Code targets parameters of later opcodes')
else:
print('Code is not self-modifying')
```
For my puzzle input, the above declares the code to not be self modifying. So all we have is addition and multiplication of memory
already harvested for opcodes and parameter addresses. It's just a big sum!
Note that some operations might write to a destination address that is then never read from, or overwritten by other operations. We could just eliminate those steps, if we could detect those cases.
### What does the sum look like?
We can skip the first operation (`ADD 1 2 3`) because the *next* expression also writes to `3` without using the outcome of the first. That makes sense, because `1` and `2` are our `noun` and `verb` inputs and those can be anywhere in the programme. Or, like I do below, you can just skip the type error that `listobject[string]` throws when trying to use either `'noun'` or `'verb'` as indices.
```
fmemory = memory[:]
fmemory[1:3] = 'noun', 'verb'
for opcode, a, b, target in zip(*([iter(fmemory)] * 4)):
if opcode == 99:
break
try:
fmemory[target] = f"({fmemory[a]}{' +*'[opcode]}{fmemory[b]})"
except TypeError as e:
# the first instruction is to add memory[noun] and memory[verb]
# and store in 3 but the next instruction also stores in 3,
# ignoring the previous result.
assert a == 'noun' and b == 'verb'
formula = fmemory[0]
print(formula)
```
If you were to compile this to a function; Python's AST optimizer will actually replace a lot of the constants; I'm using [Astor](https://github.com/berkerpeksag/astor/) here to simplify roundtripping and pretty printing, so we can see what Python makes of it:
```
import ast
import astor
from textwrap import wrap
simplified = astor.to_source(ast.parse(formula))
print("19690720 =", simplified)
```
This is something we can work with! Clearly this is a simple [linear Diophantine equation](https://en.wikipedia.org/wiki/Diophantine_equation#Linear_Diophantine_equations) that can be solved for either `noun` or `verb`, so let's see if [sympy](https://docs.sympy.org/latest/), the Python symbolic maths solver can do something with this.
We know that both `noun` and `verb` are values in the range `[0, 100)`, so we can use this to see what inputs in that range produce an output in that range:
```
import dis
from IPython.display import display, Markdown
from sympy import diophantine, lambdify, symbols, sympify, Eq, Symbol
from sympy.solvers import solve
# ask Sympy to parse our formula; it'll simplify the formula for us
display(Markdown("### Simplified expression:"))
expr = sympify(formula) - 19690720
display(expr)
# extract the symbols
noun, verb = sorted(expr.free_symbols, key=lambda s: s.name)
display(Markdown("### Solution for the linear diophantine equation"))
# solutions for the two input variables, listed in alphabetical order,
for noun_expr, verb_expr in diophantine(expr):
if isinstance(noun_expr, Symbol):
solution = verb_expr.subs(noun_expr, noun)
arg, result = noun, verb
else:
solution = noun_expr.subs(verb_expr, verb)
arg, result = verb, noun.name
display(Eq(result, solution))
for i in range(100):
other = solution.subs(arg, i)
if 0 <= other < 100:
noun_value = other if result.name == 'noun' else i
verb_value = i if result.name == 'noun' else other
display(Markdown(
f"""### Solution found:
* $noun = {noun_value}$
* $verb = {verb_value}$
* $100noun + verb = {100 * noun_value + verb_value}$
"""))
break
```
Unfortunately, even Sympy's `solveset()` function couldn't help me eliminate the loop over `range(100)`; in principle this should be possible using an `Range(100)` set, but `solveset()` just isn't quite there yet. A [related question on Stack Overflow](https://stackoverflow.com/questions/46013884/get-all-positive-integral-solutions-for-a-linear-equation) appears to confirm that using a loop is the correct method here. I could give Sage a try for this, perhaps.
That said, if you look at the $term1 - term2 \times arg$ solution to the diophantine equation, to me it is clear that `noun` and `verb` are simply the division and modulus, respectively, of $term1$ and $term2$:
```
from sympy import postorder_traversal, Integer
term1, term2 = (abs(int(e)) for e in postorder_traversal(expr) if isinstance(e, Integer))
print(f"divmod({term1}, {term2})")
noun, verb = divmod(term1, term2)
print(f"{noun=}, {verb=}, {100 * noun + verb=}")
```
| github_jupyter |
### Bidirectional LSTM
In this notebook we are going to expand the previous notebook even futher by make use of the Bidirectional LSTM, model that will be able to get reasonable accuracy in translating sequences from english to french.
**Note**: The rest of the notebook will remain the same, when there's a change i will highlight.
### Imports
```
from collections import Counter
import numpy as np
import helper, os, time
from tensorflow import keras
import tensorflow as tf
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tf.__version__
```
### Mounting the Google Drive.
```
from google.colab import drive
drive.mount('/content/drive')
```
### Paths to the files
```
base_path = '/content/drive/MyDrive/NLP Data/seq2seq/fr-en-small'
en_path = 'small_vocab_en.txt'
fr_path = 'small_vocab_fr.txt'
```
### Loading the data.
We have two files that are located at this path `'/content/drive/MyDrive/NLP Data/seq2seq/fr-en-small'` and thes files are:
```
small_vocab_fr.txt
small_vocab_en.txt
```
The following line help us to load the data.
```
eng_sents = open(os.path.join(base_path, en_path), encoding='utf8').read().split('\n')
fre_sents = open(os.path.join(base_path, fr_path), encoding='utf8').read().split('\n')
print("Data Loaded")
eng_sents[1]
```
By looking at the data we can see that the data is already preprocessed, which means we are not going to do that step here.
### Next, Bulding the Vocabulary.
Vocabulary in my definition is just unique words in the curpus. Let's look at the vocabulary size of french and english. But first we need to tokenize each sentence, Inorder for us to do that I'm going to use the `spacy` library which is my favourite when it comes to tokenization of languages.
```
import spacy
spacy.cli.download('fr_core_news_sm')
spacy_fr = spacy.load('fr_core_news_sm')
spacy_en = spacy.load('en_core_web_sm')
def tokenize_fr(sent):
return [tok.text for tok in spacy_fr.tokenizer(sent)]
def tokenize_en(sent):
return [tok.text for tok in spacy_en.tokenizer(sent)]
en_counter = Counter()
fr_counter = Counter()
for sent in eng_sents:
en_counter.update(tokenize_en(sent.lower()))
for sent in fre_sents:
fr_counter.update(tokenize_fr(sent.lower()))
en_vocab_size = len(en_counter)
fr_vocab_size = len(fr_counter)
fr_vocab_size, en_vocab_size
```
Here we have `340` unique words for french in this dataset and `201` unique words for english.
### Preprocessing.
We will convert our text data into sequence of integers so basically we are going to perform the following:
1. Tokenize the words into ids
2. Pad the tokens so that they will have same length.
For this task we are going to use the keras `Tokenizer` class to perform the task, We have been using this for sentiment analyisis so the procedure is the same.
We are going to have two tokenizers for each language.
```
en_tokenizer = Tokenizer(num_words=en_vocab_size, oov_token="<oov>")
en_tokenizer.fit_on_texts(eng_sents)
fr_tokenizer = Tokenizer(num_words=fr_vocab_size, oov_token="<oov>")
fr_tokenizer.fit_on_texts(fre_sents)
en_word_indices = en_tokenizer.word_index
en_word_indices_reversed = dict([
(v, k) for (k, v) in en_word_indices.items()
])
fr_word_indices = fr_tokenizer.word_index
fr_word_indices_reversed = dict([
(v, k) for (k, v) in fr_word_indices.items()
])
```
### Helper functions
We will create some helper function that converts sequences to text and text to sequences for each language. These function will be used for inference later on.
**We have set the out of vocabulary `oov_token|| <"oov">`token to `1` which means the word that does not exist in the vocabulary it's integer representation is 1**
```
def en_seq_to_text(sequences):
return " ".join(en_word_indices_reversed[i] for i in sequences )
def en_seq_to_text(sequences):
return " ".join(fr_word_indices_reversed[i] for i in sequences )
def en_text_to_seq(sent):
words = tokenize_en(sent.lower())
sequences = []
for word in words:
try:
sequences.append(en_word_indices[word])
except:
sequences.append(1)
return sequences
def fr_text_to_seq(sent):
words = tokenize_fr(sent.lower())
sequences = []
for word in words:
try:
sequences.append(fr_word_indices[word])
except:
sequences.append(1)
return sequences
```
### Converting text to sequences
```
en_sequences = en_tokenizer.texts_to_sequences(eng_sents)
fr_sequences = fr_tokenizer.texts_to_sequences(fre_sents)
fr_sequences[0:4]
```
### Padding Sequences.
In our case we are going to assume that the longest sentence has `100` words for both `fr` and `en` languages.
```
max_words = 100
en_tokens_padded = pad_sequences(
en_sequences,
maxlen=max_words,
padding="post",
truncating="post"
)
fr_tokens_padded = pad_sequences(
fr_sequences,
maxlen=max_words,
padding="post",
truncating="post"
)
en_tokens_padded[:2]
```
### Logits to text.
We are going to create 1 more helper function that will help us to take logits or the predictions probabilities and then we convert them to human understandable format.
```
def logits_to_text(logits, tokenizer):
index_to_words = {id: word for word, id
in tokenizer.word_index.items()}
index_to_words[0] = '<pad>'
"""
For every prediction we are going to ignore the pad token
"""
return ' '.join([index_to_words[prediction] for prediction in np.argmax(logits, 1)]).replace("<pad>", "")
```
### Bidirectional LSTM
We are going to create a bidirectional LSTM model with both LSTM layers for the backward and forward layers

```
forward_layer = keras.layers.LSTM(128, dropout=.5,
return_sequences=True,
go_backwards=False
)
backward_layer = keras.layers.LSTM(128, dropout=.5,
return_sequences=True,
go_backwards=True
)
model = keras.Sequential([
keras.layers.Embedding(
en_vocab_size,
128,
input_length=max_words
),
keras.layers.Bidirectional(keras.layers.LSTM(128, return_sequences=False)),
keras.layers.RepeatVector(100),
keras.layers.Bidirectional(forward_layer, backward_layer=backward_layer),
keras.layers.TimeDistributed(keras.layers.Dense(fr_vocab_size, activation='softmax'))
])
model.summary()
tmp_x = en_tokens_padded.reshape(
-1, 100, 1
)
tmp_x.shape
model.compile(
loss='sparse_categorical_crossentropy',
optimizer=keras.optimizers.Adam(),
metrics=['accuracy']
)
model.fit(tmp_x,
fr_tokens_padded,
batch_size=1024,
epochs=30,
validation_split=0.2
)
```
The rest of the notebook will remain the same, I believe we can load the
word pretrained word embeddings in our embedding layer as well, but that will be later on.
### Making some predictions.
Our model is targeting to predict french words, during the predict function we are going to do the following:
1. Get the sequence of the english sentence
2. Pad the english sequences and pass them to the model'
3. Reshape the logits output to the shape of `(max_len, trg_vocabsize(french)`
4. Call the `logits_to_text` function and pass the tokenizer as the `fr_tokenizer`.
5. Get the predictions
```
def predict(sent):
sequences = en_text_to_seq(sent)
padded_tokens = pad_sequences([sequences], maxlen=max_words, padding="post", truncating="post")
logits = model(padded_tokens)
logits = tf.reshape(logits, (100, -1))
return logits_to_text(logits, fr_tokenizer)
predict("your least liked fruit is the grape.")
```
### Making more predictions.
```
from prettytable import PrettyTable
def tabulate_translations(column_names, data, title, max_characters=25):
table = PrettyTable(column_names)
table.title= title
table.align[column_names[0]] = 'l'
table.align[column_names[1]] = 'l'
table.align[column_names[2]] = 'l'
table._max_width = {column_names[0] :max_characters, column_names[1] :max_characters, column_names[2]:max_characters}
for row in data:
table.add_row(row)
print(table)
columns_names = [
"English (real src sentence)", "French (the actual text)", "Translated (translated version)"
]
title = "ENGLISH TO FRENCH TRANSLATOR"
max_characters= 25
total_translations= 10
for i, (eng, fre) in enumerate(zip(eng_sents[:total_translations], fre_sents)):
rows_data = [[eng, fre, predict(eng)]]
if i + 1 != total_translations:
rows_data.append(["-" * max_characters, "-" * max_characters, "-" * max_characters ])
tabulate_translations(columns_names, rows_data, title, max_characters)
```
### Conclusion.
In this notebook we have learnt how to create an `LSTM` bidirectional model. Then what's next.
### Next
In the next notebook we will learn how we can make use of the `Encoder-Decoder` model achitecture. And probably we will get reasonable better accuracy than in this model after just training for a few epochs.
```
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jonkrohn/ML-foundations/blob/master/notebooks/gradient-descent-from-scratch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Gradient Descent from Scratch
This notebook is similar to the [*Batch Regression Gradient* notebook](https://github.com/jonkrohn/ML-foundations/blob/master/notebooks/batch-regression-gradient.ipynb) with the critical exception that we optimize via gradient descent without relying on the built-in PyTorch `SGD()` optimizer.
```
import torch
import matplotlib.pyplot as plt
xs = torch.tensor([0, 1, 2, 3, 4, 5, 6, 7.])
ys = torch.tensor([1.86, 1.31, .62, .33, .09, -.67, -1.23, -1.37])
def regression(my_x, my_m, my_b):
return my_x*my_m + my_b
m = torch.tensor([0.9]).requires_grad_()
b = torch.tensor([0.1]).requires_grad_()
```
**Step 1**: Forward pass
```
yhats = regression(xs, m, b)
yhats
```
**Step 2**: Compare $\hat{y}$ with true $y$ to calculate cost $C$
Mean squared error: $$C = \frac{1}{n} \sum_{i=1}^n (\hat{y_i}-y_i)^2 $$
```
def mse(my_yhat, my_y):
sigma = torch.sum((my_yhat - my_y)**2)
return sigma/len(my_y)
C = mse(yhats, ys)
C
```
**Step 3**: Use autodiff to calculate gradient of $C$ w.r.t. parameters
```
C.backward()
m.grad
```
$\frac{\partial C}{\partial m} = 36.3$ indicates that an increase in $m$ corresponds to a large increase in $C$.
```
b.grad
```
Meanwhile, $\frac{\partial C}{\partial b} = 6.26$ indicates that an increase in $b$ also corresponds to an increase in $C$, though much less so than $m$.
(Using partial derivatives derived in [*Calculus II*](https://github.com/jonkrohn/ML-foundations/blob/master/notebooks/4-calculus-ii.ipynb), we could alternatively calculate these same slopes without automatic numerical computation:)
$$ \frac{\partial C}{\partial m} = \frac{2}{n} \sum (\hat{y}_i - y_i) \cdot x_i $$
```
2*1/len(ys)*torch.sum((yhats - ys)*xs)
```
$$ \frac{\partial C}{\partial b} = \frac{2}{n} \sum (\hat{y}_i - y_i) $$
```
2*1/len(ys)*torch.sum(yhats - ys)
```
The gradient of cost, $\nabla C$, is:
```
gradient = torch.tensor([[b.grad.item(), m.grad.item()]]).T
gradient
```
For convenience, model parameters are often denoted as $\boldsymbol{\theta}$, which, depending on the model, could be, for example, a vector, a matrix, or a collection of tensors of varying dimensions. With our simple linear model, a vector tensor will do:
```
theta = torch.tensor([[b, m]]).T
theta
```
Note the gradient $\nabla C$ could thus alternatively be denoted with respect to $\boldsymbol{\theta}$ as $\nabla_\boldsymbol{\theta} f(\boldsymbol{\theta})$.
(Also, note that we're transposing $\boldsymbol{\theta}$ to make forthcoming tensor operations easier because of the convention in ML to transpose the gradient, $\nabla C$.)
Let's visualize the state of the most pertinent metrics in a single plot:
```
def labeled_regression_plot(my_x, my_y, my_m, my_b, my_C, include_grad=True):
title = 'Cost = {}'.format('%.3g' % my_C.item())
if include_grad:
xlabel = 'm = {}, m grad = {}'.format('%.3g' % my_m.item(), '%.3g' % my_m.grad.item())
ylabel = 'b = {}, b grad = {}'.format('%.3g' % my_b.item(), '%.3g' % my_b.grad.item())
else:
xlabel = 'm = {}'.format('%.3g' % my_m.item())
ylabel = 'b = {}'.format('%.3g' % my_b.item())
fig, ax = plt.subplots()
plt.title(title)
plt.ylabel(ylabel)
plt.xlabel(xlabel)
ax.scatter(my_x, my_y)
x_min, x_max = ax.get_xlim()
y_min, y_max = my_m*x_min + my_b, my_m*x_max + my_b
ax.set_xlim([x_min, x_max])
_ = ax.plot([x_min, x_max], [y_min, y_max], c='C01')
labeled_regression_plot(xs, ys, m, b, C)
```
**Step 4**: Gradient descent
In the first round of training, with $\frac{\partial C}{\partial m} = 36.3$ and $\frac{\partial C}{\partial b} = 6.26$, the lowest hanging fruit with respect to reducing cost $C$ is to decrease the slope of the regression line, $m$. The model would also benefit from a comparatively small decrease in the $y$-intercept of the line, $b$.
To control exactly how much we adjust the model parameters $\boldsymbol{\theta}$, we set a **learning rate**, a hyperparameter of ML models that use gradient descent (that is typically denoted with $\alpha$):
```
lr = 0.01 # Cover rules of thumb
```
We use the learning rate $\alpha$ to scale the gradient, i.e., $\alpha \nabla C$:
```
scaled_gradient = lr * gradient
scaled_gradient
```
We can now use our scaled gradient to adjust our model parameters $\boldsymbol{\theta}$ in directions that will reduce the model cost $C$.
Since, e.g., $\frac{\partial C}{\partial m} = 36.3$ indicates that increasing the slope parameter $m$ corresponds to an increase in cost $C$, we *subtract* the gradient to adjust each individual parameter in a direction that reduces cost: $$ \boldsymbol{\theta}' = \boldsymbol{\theta} - \alpha \nabla C$$
```
new_theta = theta - scaled_gradient
new_theta
```
To see these adjustments even more clearly, you can consider each parameter individually, e.g., $m' = m - \alpha \frac{\partial C}{\partial m}$:
```
m - lr*m.grad
```
...and $b' = b - \alpha \frac{\partial C}{\partial b}$:
```
b - lr*b.grad
```
With our updated parameters $\boldsymbol{\theta}$ now in hand, we can use them to check that they do indeed correspond to a decreased cost $C$:
```
b = new_theta[0]
m = new_theta[1]
C = mse(regression(xs, m, b), ys)
labeled_regression_plot(xs, ys, m, b, C, include_grad=False) # Gradient of C hasn't been recalculated
```
### Rinse and Repeat
To perform another round of gradient descent, we let PyTorch know we'd like to track gradients on the tensors `b` and `m` (as we did at the top of the notebook when we created them the first time):
```
b.requires_grad_()
_ = m.requires_grad_() # "_ =" is to prevent output within Jupyter; it is cosmetic only
epochs = 8
for epoch in range(epochs):
yhats = regression(xs, m, b) # Step 1
C = mse(yhats, ys) # Step 2
C.backward() # Step 3
labeled_regression_plot(xs, ys, m, b, C)
gradient = torch.tensor([[b.grad.item(), m.grad.item()]]).T
theta = torch.tensor([[b, m]]).T
new_theta = theta - lr*gradient # Step 4
b = new_theta[0].requires_grad_()
m = new_theta[1].requires_grad_()
```
(Note that the above plots are identical to those in the [*Batch Regression Gradient* notebook](https://github.com/jonkrohn/ML-foundations/blob/master/notebooks/batch-regression-gradient.ipynb), in which we used the PyTorch `SGD()` method to descend the gradient.)
In later rounds of training, after the model's slope $m$ has become closer to the slope represented by the data, $\frac{\partial C}{\partial b}$ becomes negative, indicating an inverse relationship between $b$ and $C$. Meanwhile, $\frac{\partial C}{\partial m}$ remains positive.
This combination directs gradient descent to simultaneously adjust the $y$-intercept $b$ upwards and the slope $m$ downwards in order to reduce cost $C$ and, ultimately, fit the regression line snugly to the data.
Finally, let's run a thousand more epochs (without plots) to converge on optimal parameters $\boldsymbol{\theta}$:
```
epochs = 992 # accounts for rounds above to match 1000 epochs of regression-in-pytorch.ipynb
for epoch in range(epochs):
yhats = regression(xs, m, b) # Step 1
C = mse(yhats, ys) # Step 2
C.backward() # Step 3
print('Epoch {}, cost {}, m grad {}, b grad {}'.format(epoch, '%.3g' % C.item(), '%.3g' % m.grad.item(), '%.3g' % b.grad.item()))
gradient = torch.tensor([[b.grad.item(), m.grad.item()]]).T
theta = torch.tensor([[b, m]]).T
new_theta = theta - lr*gradient # Step 4
b = new_theta[0].requires_grad_()
m = new_theta[1].requires_grad_()
labeled_regression_plot(xs, ys, m, b, C, include_grad=False)
```
(Note that the above results are identical to those in the [*Regression in PyTorch* notebook](https://github.com/jonkrohn/ML-foundations/blob/master/notebooks/regression-in-pytorch.ipynb), in which we also used the PyTorch `SGD()` method to descend the gradient.)
```
```
| github_jupyter |
# Word2Vec and t-SNE
A question that might come up when working with text is: how do you turn text into numbers?
In the past, common techniques included methods like one-hot vectors, in which we'd have a different number associated with each word, and then turn "on" the value at that index in a vector (making it 1) and setting all the rest to zero.
For instance, if we have the sentence: "I like dogs", we'd have a 3-dimensional one-hot vector (3-dimensional because there are three words), so the word "I" might be `[1,0,0]`, the word "like" might be `[0,1,0]`, and "dogs" would be `[0,0,1]`.
One-hot vectors worked well enough for some tasks but it's not a particularly rich or meaningful representation of text. The indices of these words are arbitrary and don't describe any relationship between them.
[_Word embeddings_](http://arxiv.org/pdf/1301.3781.pdf) provide a meaningful representation of text. Word embeddings, called such because they involve embedding a word in some high-dimensional space, that is, they map a word to some vector, much like one-hot vectors. The difference is that word embeddings are learned for a particular task, so they end up being meaningful representations.
For example, the relationships between words are meaningful (image from the [TensorFlow documentation]((https://www.tensorflow.org/versions/r0.9/tutorials/word2vec/index.html)):
{:width="100%"}
A notable property that emerges is that vector arithmetic is also meaningful. Perhaps the most well-known example of this is:
$$
\text{king} - \text{man} + \text{woman} = \text{queen}
$$
([Chris Olah's piece on word embeddings](Deep Learning, NLP, and Representations) delves more into why this is.)
So the positioning of these words in this space actually tells us something about how these words are used.
This allows us to do things like find the most similar words by looking at the closest words. You can project the resulting embeddings down to 2D so that we can visualize them. We'll use t-SNE ("t-Distributed Stochastic Neighbor Embedding") for this, which is a dimensionality reduction method that works well for visualizing high-dimension data. We'll see that clusters of related words form in a way that a human would probably agree with. We couldn't do this with one-hot vectors - the distances between them are totally arbitrary and their proximity is essentially random.
As mentioned earlier, these word embeddings are trained to help with a particular task, which is learned through a neural network. Two tasks developed for training embeddings is _CBOW_ (continuous bag of words) and _skip-grams_; together these methods of learning word embeddings are called "Word2Vec".
For the CBOW task, we take the context words (the words around the target word) and give the target word. We want to predict whether or not the target word belongs to the context.
The skip-grams is basically the inverse: we take the target word (the "pivot"), then give the context. We want to predict whether or not the context belongs to the word.
They are quite similar but have different properties, e.g. CBOW works better on smaller datasets, where as skip-grams works better for larger ones. In any case, the idea with word embeddings is that they can be trained to help with any task.
We're going to be using the skip-gram task here.
## Corpus
We need a reasonably-sized text corpus to learn from. Here we'll use State of the Union addresses retrieved from [The American Presidency Project](http://www.presidency.ucsb.edu/sou.php). These addresses tend to use similar patterns so we should be able to learn some decent word embeddings. Since the skip-gram task looks at context, texts that use words in a consistent way (i.e. in consistent contexts) we'll be able to learn better.
[The corpus is available here](/guides/data/sotu.tar.gz). The texts were preprocessed a bit (mainly removing URL-encoded characters). The texts provided here are the processed versions (nb: this isn't the complete collection of texts but enough to work with here).
## Skip-grams
Before we go any further, let's get a bit more concrete about what the skip-gram task is.
Let's consider the sentence "I think cats are cool".
The skip-gram task is as follows:
- We take a word, e.g. `'cats'`, which we'll represent as $w_i$. We feed this as input into our neural network.
- We take the word's context, e.g. `['I', 'think', 'are', 'cool']`. We'll represent this as $\{w_{i-2}, w_{i-1}, w_{i+1}, w_{i+2}\}$ and we also feed this into our neural network.
- Then we just want our network to predict (i.e. classify) whether or not $\{w_{i-2}, w_{i-1}, w_{i+1}, w_{i+2}\}$ is the true context of $w_i$.
For this particular example we'd want the network to output 1 (i.e. yes, that is the true context).
If we set $w_i$ to 'frogs', then we'd want the network output 0. In our one sentence corpus, `['I', 'think', 'are', 'cool']` is not the true context for 'frogs'. Sorry frogs 🐸.
## Building the model
We'll use `keras` to build the neural network that we'll use to learn the embeddings.
First we'll import everything:
```
import numpy as np
from keras.models import Sequential
from keras.layers.embeddings import Embedding
from keras.layers import Flatten, Activation, Merge
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import skipgrams, make_sampling_table
```
Then load in our data. We're actually going to define a generator to load our data in on-demand; this way we'll avoid having all our data sitting around in memory when we don't need it.
```
from glob import glob
text_files = glob('../data/sotu/*.txt')
def text_generator():
for path in text_files:
with open(path, 'r') as f:
yield f.read()
len(text_files)
```
Before we go any further, we need to map the words in our corpus to numbers, so that we have a consistent way of referring to them. First we'll fit a tokenizer to the corpus:
```
# our corpus is small enough where we
# don't need to worry about this, but good practice
max_vocab_size = 50000
# `filters` specify what characters to get rid of
tokenizer = Tokenizer(nb_words=max_vocab_size,
filters='!"#$%&()*+,-./:;<=>?@[\\]^_{|}~\t\n\'`“”–')
# fit the tokenizer
tokenizer.fit_on_texts(text_generator())
# we also want to keep track of the actual vocab size
# we'll need this later
# note: we add one because `0` is a reserved index in keras' tokenizer
vocab_size = len(tokenizer.word_index) + 1
```
Now the tokenizer knows what tokens (words) are in our corpus and has mapped them to numbers. The `keras` tokenizer also indexes them in order of frequency (most common first, i.e. index 1 is usually a word like "the"), which will come in handy later.
At this point, let's define the dimensions of our embeddings. It's up to you and your task to choose this number. Like many neural network hyperparameters, you may just need to play around with it.
```
embedding_dim = 256
```
Now let's define the model. When I described the skip-gram task, I mentioned two inputs: the target word (also called the "pivot") and the context. So we're going to build two separate models for each input and then merge them into one.
```
pivot_model = Sequential()
pivot_model.add(Embedding(vocab_size, embedding_dim, input_length=1))
context_model = Sequential()
context_model.add(Embedding(vocab_size, embedding_dim, input_length=1))
# merge the pivot and context models
model = Sequential()
model.add(Merge([pivot_model, context_model], mode='dot', dot_axes=2))
model.add(Flatten())
# the task as we've framed it here is
# just binary classification,
# so we want the output to be in [0,1],
# and we can use binary crossentropy as our loss
model.add(Activation('sigmoid'))
model.compile(optimizer='adam', loss='binary_crossentropy')
```
Finally, we can train the model.
```
n_epochs = 60
# used to sample words (indices)
sampling_table = make_sampling_table(vocab_size)
for i in range(n_epochs):
loss = 0
for seq in tokenizer.texts_to_sequences_generator(text_generator()):
# generate skip-gram training examples
# - `couples` consists of the pivots (i.e. target words) and surrounding contexts
# - `labels` represent if the context is true or not
# - `window_size` determines how far to look between words
# - `negative_samples` specifies the ratio of negative couples
# (i.e. couples where the context is false)
# to generate with respect to the positive couples;
# i.e. `negative_samples=4` means "generate 4 times as many negative samples"
couples, labels = skipgrams(seq, vocab_size, window_size=5, negative_samples=4, sampling_table=sampling_table)
if couples:
pivot, context = zip(*couples)
pivot = np.array(pivot, dtype='int32')
context = np.array(context, dtype='int32')
labels = np.array(labels, dtype='int32')
loss += model.train_on_batch([pivot, context], labels)
print('epoch %d, %0.02f'%(i, loss))
```
With any luck, the model should finish training without a hitch.
Now we can extract the embeddings, which are just the weights of the pivot embedding layer:
```
embeddings = model.get_weights()[0]
```
We also want to set aside the tokenizer's word index for later use (so we can get indices for words) and also create a reverse word index (so we can get words from indices):
```
word_index = tokenizer.word_index
reverse_word_index = {v: k for k, v in word_index.items()}
```
That's it for learning the embeddings. Now we can try using them.
## Getting similar words
Each word embedding is just a mapping of a word to some point in space. So if we want to find words similar to some target word, we literally just need to look at the closest embeddings to that target word's embedding.
An example will make this clearer.
First, let's write a simple function to retrieve an embedding for a word:
```
def get_embedding(word):
idx = word_index[word]
# make it 2d
return embeddings[idx][:,np.newaxis].T
```
Then we can define a function to get a most similar word for an input word:
```
from scipy.spatial.distance import cdist
ignore_n_most_common = 50
def get_closest(word):
embedding = get_embedding(word)
# get the distance from the embedding
# to every other embedding
distances = cdist(embedding, embeddings)[0]
# pair each embedding index and its distance
distances = list(enumerate(distances))
# sort from closest to furthest
distances = sorted(distances, key=lambda d: d[1])
# skip the first one; it's the target word
for idx, dist in distances[1:]:
# ignore the n most common words;
# they can get in the way.
# because the tokenizer organized indices
# from most common to least, we can just do this
if idx > ignore_n_most_common:
return reverse_word_index[idx]
```
Now let's give it a try (you may get different results):
```
print(get_closest('freedom'))
print(get_closest('justice'))
print(get_closest('america'))
print(get_closest('citizens'))
print(get_closest('citizen'))
```
For the most part, we seem to be getting related words!
NB: Here we computed distances to _every_ other embedding, which is far from ideal when dealing with really large vocabularies. `Gensim`'s [`Word2Vec`](https://radimrehurek.com/gensim/models/word2vec.html) class implements a `most_similar` method that uses an approximate, but much faster, method for finding similar words. You can import the embeddings learned here into that class:
```
from gensim.models.doc2vec import Word2Vec
with open('embeddings.dat', 'w') as f:
f.write('{} {}'.format(vocab_size, embedding_dim))
for word, idx in word_index.items():
embedding = ' '.join(str(d) for d in embeddings[idx])
f.write('\n{} {}'.format(word, embedding))
w2v = Word2Vec.load_word2vec_format('embeddings.dat', binary=False)
print(w2v.most_similar(positive=['freedom']))
```
## t-SNE
t-SNE ("t-Distributed Stochastic Neighbor Embedding") is a way of projecting high-dimensional data, e.g. our word embeddings, to a lower-dimension space, e.g. 2D, so we can visualize it.
This will give us a better sense of the quality of our embeddings: we should see clusters of related words.
`scikit-learn` provides a t-SNE implementation that is very easy to use.
```
from sklearn.manifold import TSNE
# `n_components` is the number of dimensions to reduce to
tsne = TSNE(n_components=2)
# apply the dimensionality reduction
# to our embeddings to get our 2d points
points = tsne.fit_transform(embeddings)
```
And now let's plot it out:
```
print(points)
import matplotlib
matplotlib.use('Agg') # for pngs
import matplotlib.pyplot as plt
# plot our results
# make it quite big so we can see everything
fig, ax = plt.subplots(figsize=(40, 20))
# extract x and y values separately
xs = points[:,0]
ys = points[:,1]
# plot the points
# we don't actually care about the point markers,
# just want to automatically set the bounds of the plot
ax.scatter(xs, ys, alpha=0)
# annotate each point with its word
for i, point in enumerate(points):
ax.annotate(reverse_word_index.get(i),
(xs[i], ys[i]),
fontsize=8)
plt.savefig('tsne.png')
```

This looks pretty good! It could certainly be improved upon, with more data or more training, but it's a great start.
## Further Reading
- [Deep Learning, NLP, and Representations](http://colah.github.io/posts/2014-07-NLP-RNNs-Representations/). Chris Olah.
- [On Word Embeddings](http://sebastianruder.com/word-embeddings-1/). Sebastian Ruder.
- Mikolov, T., Chen, K., Corrado, G., & Dean, J. (2013). [Efficient estimation of word representations in vector space](http://arxiv.org/pdf/1301.3781.pdf). arXiv preprint arXiv:1301.3781.
- Mikolov, T., Sutskever, I., Chen, K., Corrado, G. S., & Dean, J. (2013). [Distributed representations of words and phrases and their compositionality](http://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf). In Advances in neural information processing systems (pp. 3111-3119).
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.

# Tutorial: Use automated machine learning to predict taxi fares
In this tutorial, you use automated machine learning in Azure Machine Learning service to create a regression model to predict NYC taxi fare prices. This process accepts training data and configuration settings, and automatically iterates through combinations of different feature normalization/standardization methods, models, and hyperparameter settings to arrive at the best model.
In this tutorial you learn the following tasks:
* Download, transform, and clean data using Azure Open Datasets
* Train an automated machine learning regression model
* Calculate model accuracy
If you don’t have an Azure subscription, create a free account before you begin. Try the [free or paid version](https://aka.ms/AMLFree) of Azure Machine Learning service today.
## Prerequisites
* Complete the [setup tutorial](https://docs.microsoft.com/azure/machine-learning/service/tutorial-1st-experiment-sdk-setup) if you don't already have an Azure Machine Learning service workspace or notebook virtual machine.
* After you complete the setup tutorial, open the **tutorials/regression-automated-ml.ipynb** notebook using the same notebook server.
This tutorial is also available on [GitHub](https://github.com/Azure/MachineLearningNotebooks/tree/master/tutorials) if you wish to run it in your own [local environment](https://docs.microsoft.com/azure/machine-learning/service/how-to-configure-environment#local). Run `pip install azureml-sdk[automl] azureml-opendatasets azureml-widgets` to get the required packages.
## Download and prepare data
Import the necessary packages. The Open Datasets package contains a class representing each data source (`NycTlcGreen` for example) to easily filter date parameters before downloading.
```
from azureml.opendatasets import NycTlcGreen
import pandas as pd
from datetime import datetime
from dateutil.relativedelta import relativedelta
```
Begin by creating a dataframe to hold the taxi data. When working in a non-Spark environment, Open Datasets only allows downloading one month of data at a time with certain classes to avoid `MemoryError` with large datasets. To download taxi data, iteratively fetch one month at a time, and before appending it to `green_taxi_df` randomly sample 2,000 records from each month to avoid bloating the dataframe. Then preview the data.
```
green_taxi_df = pd.DataFrame([])
start = datetime.strptime("1/1/2015","%m/%d/%Y")
end = datetime.strptime("1/31/2015","%m/%d/%Y")
for sample_month in range(12):
temp_df_green = NycTlcGreen(start + relativedelta(months=sample_month), end + relativedelta(months=sample_month)) \
.get_tabular_dataset().to_pandas_dataframe()
green_taxi_df = green_taxi_df.append(temp_df_green.sample(2000))
green_taxi_df.head(10)
```
Now that the initial data is loaded, define a function to create various time-based features from the pickup datetime field. This will create new fields for the month number, day of month, day of week, and hour of day, and will allow the model to factor in time-based seasonality.
Use the `apply()` function on the dataframe to iteratively apply the `build_time_features()` function to each row in the taxi data.
```
def build_time_features(vector):
pickup_datetime = vector[0]
month_num = pickup_datetime.month
day_of_month = pickup_datetime.day
day_of_week = pickup_datetime.weekday()
hour_of_day = pickup_datetime.hour
return pd.Series((month_num, day_of_month, day_of_week, hour_of_day))
green_taxi_df[["month_num", "day_of_month","day_of_week", "hour_of_day"]] = green_taxi_df[["lpepPickupDatetime"]].apply(build_time_features, axis=1)
green_taxi_df.head(10)
```
Remove some of the columns that you won't need for training or additional feature building.
```
columns_to_remove = ["lpepPickupDatetime", "lpepDropoffDatetime", "puLocationId", "doLocationId", "extra", "mtaTax",
"improvementSurcharge", "tollsAmount", "ehailFee", "tripType", "rateCodeID",
"storeAndFwdFlag", "paymentType", "fareAmount", "tipAmount"
]
for col in columns_to_remove:
green_taxi_df.pop(col)
green_taxi_df.head(5)
```
### Cleanse data
Run the `describe()` function on the new dataframe to see summary statistics for each field.
```
green_taxi_df.describe()
```
From the summary statistics, you see that there are several fields that have outliers or values that will reduce model accuracy. First filter the lat/long fields to be within the bounds of the Manhattan area. This will filter out longer taxi trips or trips that are outliers in respect to their relationship with other features.
Additionally filter the `tripDistance` field to be greater than zero but less than 31 miles (the haversine distance between the two lat/long pairs). This eliminates long outlier trips that have inconsistent trip cost.
Lastly, the `totalAmount` field has negative values for the taxi fares, which don't make sense in the context of our model, and the `passengerCount` field has bad data with the minimum values being zero.
Filter out these anomalies using query functions, and then remove the last few columns unnecessary for training.
```
final_df = green_taxi_df.query("pickupLatitude>=40.53 and pickupLatitude<=40.88")
final_df = final_df.query("pickupLongitude>=-74.09 and pickupLongitude<=-73.72")
final_df = final_df.query("tripDistance>=0.25 and tripDistance<31")
final_df = final_df.query("passengerCount>0 and totalAmount>0")
columns_to_remove_for_training = ["pickupLongitude", "pickupLatitude", "dropoffLongitude", "dropoffLatitude"]
for col in columns_to_remove_for_training:
final_df.pop(col)
```
Call `describe()` again on the data to ensure cleansing worked as expected. You now have a prepared and cleansed set of taxi, holiday, and weather data to use for machine learning model training.
```
final_df.describe()
```
## Configure workspace
Create a workspace object from the existing workspace. A [Workspace](https://docs.microsoft.com/python/api/azureml-core/azureml.core.workspace.workspace?view=azure-ml-py) is a class that accepts your Azure subscription and resource information. It also creates a cloud resource to monitor and track your model runs. `Workspace.from_config()` reads the file **config.json** and loads the authentication details into an object named `ws`. `ws` is used throughout the rest of the code in this tutorial.
```
from azureml.core.workspace import Workspace
ws = Workspace.from_config()
```
## Split the data into train and test sets
Split the data into training and test sets by using the `train_test_split` function in the `scikit-learn` library. This function segregates the data into the x (**features**) data set for model training and the y (**values to predict**) data set for testing. The `test_size` parameter determines the percentage of data to allocate to testing. The `random_state` parameter sets a seed to the random generator, so that your train-test splits are deterministic.
```
from sklearn.model_selection import train_test_split
x_train, x_test = train_test_split(final_df, test_size=0.2, random_state=223)
```
The purpose of this step is to have data points to test the finished model that haven't been used to train the model, in order to measure true accuracy.
In other words, a well-trained model should be able to accurately make predictions from data it hasn't already seen. You now have data prepared for auto-training a machine learning model.
## Automatically train a model
To automatically train a model, take the following steps:
1. Define settings for the experiment run. Attach your training data to the configuration, and modify settings that control the training process.
1. Submit the experiment for model tuning. After submitting the experiment, the process iterates through different machine learning algorithms and hyperparameter settings, adhering to your defined constraints. It chooses the best-fit model by optimizing an accuracy metric.
### Define training settings
Define the experiment parameter and model settings for training. View the full list of [settings](https://docs.microsoft.com/azure/machine-learning/service/how-to-configure-auto-train). Submitting the experiment with these default settings will take approximately 20 minutes, but if you want a shorter run time, reduce the `experiment_timeout_hours` parameter.
|Property| Value in this tutorial |Description|
|----|----|---|
|**iteration_timeout_minutes**|2|Time limit in minutes for each iteration. Reduce this value to decrease total runtime.|
|**experiment_timeout_hours**|0.3|Maximum amount of time in hours that all iterations combined can take before the experiment terminates.|
|**enable_early_stopping**|True|Flag to enable early termination if the score is not improving in the short term.|
|**primary_metric**| spearman_correlation | Metric that you want to optimize. The best-fit model will be chosen based on this metric.|
|**featurization**| auto | By using auto, the experiment can preprocess the input data (handling missing data, converting text to numeric, etc.)|
|**verbosity**| logging.INFO | Controls the level of logging.|
|**n_cross_validations**|5|Number of cross-validation splits to perform when validation data is not specified.|
```
import logging
automl_settings = {
"iteration_timeout_minutes": 2,
"experiment_timeout_hours": 0.3,
"enable_early_stopping": True,
"primary_metric": 'spearman_correlation',
"featurization": 'auto',
"verbosity": logging.INFO,
"n_cross_validations": 5
}
```
Use your defined training settings as a `**kwargs` parameter to an `AutoMLConfig` object. Additionally, specify your training data and the type of model, which is `regression` in this case.
```
from azureml.train.automl import AutoMLConfig
automl_config = AutoMLConfig(task='regression',
debug_log='automated_ml_errors.log',
training_data=x_train,
label_column_name="totalAmount",
**automl_settings)
```
Automated machine learning pre-processing steps (feature normalization, handling missing data, converting text to numeric, etc.) become part of the underlying model. When using the model for predictions, the same pre-processing steps applied during training are applied to your input data automatically.
### Train the automatic regression model
Create an experiment object in your workspace. An experiment acts as a container for your individual runs. Pass the defined `automl_config` object to the experiment, and set the output to `True` to view progress during the run.
After starting the experiment, the output shown updates live as the experiment runs. For each iteration, you see the model type, the run duration, and the training accuracy. The field `BEST` tracks the best running training score based on your metric type.
```
from azureml.core.experiment import Experiment
experiment = Experiment(ws, "taxi-experiment")
local_run = experiment.submit(automl_config, show_output=True)
```
## Explore the results
Explore the results of automatic training with a [Jupyter widget](https://docs.microsoft.com/python/api/azureml-widgets/azureml.widgets?view=azure-ml-py). The widget allows you to see a graph and table of all individual run iterations, along with training accuracy metrics and metadata. Additionally, you can filter on different accuracy metrics than your primary metric with the dropdown selector.
```
from azureml.widgets import RunDetails
RunDetails(local_run).show()
```
### Retrieve the best model
Select the best model from your iterations. The `get_output` function returns the best run and the fitted model for the last fit invocation. By using the overloads on `get_output`, you can retrieve the best run and fitted model for any logged metric or a particular iteration.
```
best_run, fitted_model = local_run.get_output()
print(best_run)
print(fitted_model)
```
### Test the best model accuracy
Use the best model to run predictions on the test data set to predict taxi fares. The function `predict` uses the best model and predicts the values of y, **trip cost**, from the `x_test` data set. Print the first 10 predicted cost values from `y_predict`.
```
y_test = x_test.pop("totalAmount")
y_predict = fitted_model.predict(x_test)
print(y_predict[:10])
```
Calculate the `root mean squared error` of the results. Convert the `y_test` dataframe to a list to compare to the predicted values. The function `mean_squared_error` takes two arrays of values and calculates the average squared error between them. Taking the square root of the result gives an error in the same units as the y variable, **cost**. It indicates roughly how far the taxi fare predictions are from the actual fares.
```
from sklearn.metrics import mean_squared_error
from math import sqrt
y_actual = y_test.values.flatten().tolist()
rmse = sqrt(mean_squared_error(y_actual, y_predict))
rmse
```
Run the following code to calculate mean absolute percent error (MAPE) by using the full `y_actual` and `y_predict` data sets. This metric calculates an absolute difference between each predicted and actual value and sums all the differences. Then it expresses that sum as a percent of the total of the actual values.
```
sum_actuals = sum_errors = 0
for actual_val, predict_val in zip(y_actual, y_predict):
abs_error = actual_val - predict_val
if abs_error < 0:
abs_error = abs_error * -1
sum_errors = sum_errors + abs_error
sum_actuals = sum_actuals + actual_val
mean_abs_percent_error = sum_errors / sum_actuals
print("Model MAPE:")
print(mean_abs_percent_error)
print()
print("Model Accuracy:")
print(1 - mean_abs_percent_error)
```
From the two prediction accuracy metrics, you see that the model is fairly good at predicting taxi fares from the data set's features, typically within +- $4.00, and approximately 15% error.
The traditional machine learning model development process is highly resource-intensive, and requires significant domain knowledge and time investment to run and compare the results of dozens of models. Using automated machine learning is a great way to rapidly test many different models for your scenario.
## Clean up resources
Do not complete this section if you plan on running other Azure Machine Learning service tutorials.
### Stop the notebook VM
If you used a cloud notebook server, stop the VM when you are not using it to reduce cost.
1. In your workspace, select **Compute**.
1. Select the **Notebook VMs** tab in the compute page.
1. From the list, select the VM.
1. Select **Stop**.
1. When you're ready to use the server again, select **Start**.
### Delete everything
If you don't plan to use the resources you created, delete them, so you don't incur any charges.
1. In the Azure portal, select **Resource groups** on the far left.
1. From the list, select the resource group you created.
1. Select **Delete resource group**.
1. Enter the resource group name. Then select **Delete**.
You can also keep the resource group but delete a single workspace. Display the workspace properties and select **Delete**.
## Next steps
In this automated machine learning tutorial, you did the following tasks:
> * Configured a workspace and prepared data for an experiment.
> * Trained by using an automated regression model locally with custom parameters.
> * Explored and reviewed training results.
[Deploy your model](https://docs.microsoft.com/azure/machine-learning/service/tutorial-deploy-models-with-aml) with Azure Machine Learning service.
| github_jupyter |
# hana-ml Tutorial - Dataframe
**Author: TI HDA DB HANA Core CN**
The SAP HANA Python Client API for machine learning algorithms (Python Client API for ML) provides a set of client-side Python functions for accessing and querying SAP HANA data, and a set of functions for developing machine learning models.
In this tutorial, we will show you how to use various functions of Dataframe.
A dataframe represents a table (or any SQL statement). Most operations on a dataframe are designed to not bring data back from the database unless explicitly asked for.
## Import necessary libraries and functions
```
from hana_ml import dataframe
from hana_ml.dataframe import ConnectionContext
from hana_ml.algorithms.pal.utility import DataSets, Settings
import pandas as pd
```
## Create a connection to a SAP HANA instance
First, you need to create a connetion to a SAP HANA instance. In the following cell, we use a config file, config/e2edata.ini to control the connection parameters.
In your case, please update the following url, port, user, pwd with your HANA instance information for setting up the connection.
```
# Please replace url, port, user, pwd with your HANA instance information
connection_context = ConnectionContext(url, port, user, pwd)
```
## Obtain a hana-ml dataframe
**1. Create a hana-ml dataframe from pandas DataFrame**
create_dataframe_from_pandas() will create a dataframe from a pandas DataFrame and create a table in SAP HANA.
```
df = dataframe.create_dataframe_from_pandas(connection_context=connection_context,
pandas_df=pd.DataFrame({"ID": [1,2,5],
"ID2": [1,None,5],
"V3": [2,3,4],
"V4": [3,3,3],
"V5": ['a', None, 'b']}),
table_name="#tt_null",
force=True)
print(type(df))
```
**2. Invoke table function**
This function returns a dataframe which represents a specified table in SAP HANA
```
df1 = connection_context.table("#tt_null")
print(type(df1))
```
## Property of dataframe
**SQL select statement**
```
print(df.select_statement)
```
**Connection**
```
df.connection_context
```
## Fetch data in SAP HANA to client
Fetch the first 5 rows of df into client as a <b>Pandas Dataframe</b>
```
pd_df = df.head(5).collect()
print(pd_df)
print(type(pd_df))
```
## Save a dataframe
```
# Creates a table or view holding the current DataFrame's data.
df.save(where="#TT")
print(connection_context.table("#TT").collect())
```
## Simple Operations
In this section, we will show some basic operations of hana-ml dataframe. Please refer more functions in detail in the dataframe documentation.
In hana-ml, we provide a class called DataSets which contains several small public datasets. You could use load_bank_data() to load the bank dataset. We will use the bank dataset in the following examples.
```
# load the dataset and obtain a series of dataframe
bank_df, _, _, _ = DataSets.load_bank_data(connection_context)# the table name in SAP HANA is DBM2_RFULL_TBL
print(bank_df.head(3).collect())
print(type(bank_df))
```
**columns**
```
bank_df.columns
```
**shape**
```
bank_df.shape
```
**get_table_structure**
```
bank_df.get_table_structure()
```
**Add ID**
```
bank_df.add_id(id_col='ID1').head(5).collect()
```
**Add a constant column**
```
bank_df.add_constant(column_name='Constant', value=888).head(5).collect()
```
**Count the number of rows**
```
bank_df.count()
```
**Drop duplicates**
```
df_no_duplicate = bank_df.drop_duplicates()
print(df_no_duplicate.count())# there is no duplicate row in bank dataset
print(df_no_duplicate.select_statement)
```
#### Remove a column
```
df1 = bank_df
print(df1.shape)
df2 = df1.drop(["LABEL"])
print(df2.shape)
print(df2.select_statement)
```
**Filtering Data**
```
print(bank_df.filter('AGE > 60').head(3).collect())
print(bank_df.filter('AGE > 60').select_statement)
```
**Sorting**
```
print(bank_df.filter('AGE>60').sort(['AGE']).head(3).collect())
print(bank_df.filter('AGE>60').sort(['AGE']).select_statement)
```
**Cast**
```
bank_df.cast({"AGE": "BIGINT", "JOB": "NVARCHAR(50)"}).get_table_structure()
```
**Distinct**
```
bank_df.distinct(cols='JOB').collect()
```
**Describing a dataframe**
```
bank_df.describe().collect()
```
**Replace NULL value with a specified value**
```
print(df.collect())
print(df.fillna(value=0).collect())
print(df.collect())
print(df.fillna(value='').collect())
print(df.collect())
print(df.fillna('').fillna(0).collect())
```
**Projection**
```
dsp = bank_df.select("ID", "AGE", "JOB", ('"AGE"*2', "TWICE_AGE"))
print(dsp.head(5).collect())
print(dsp.select_statement)
```
**Simple Joins**
```
df1 = dataframe.create_dataframe_from_pandas(connection_context=connection_context,
pandas_df=pd.DataFrame({"ID": [1,2,3],
"ID2": [1,2,3],
"V1": [2,3,4]}),
table_name="#tt1",
force=True)
df2 = dataframe.create_dataframe_from_pandas(connection_context=connection_context,
pandas_df=pd.DataFrame({"ID": [1,2],
"ID2": [1,2],
"V2": [2,3]}),
table_name="#tt2",
force=True)
df3 = dataframe.create_dataframe_from_pandas(connection_context=connection_context,
pandas_df=pd.DataFrame({"ID": [1,2,5],
"ID2": [1,2,5],
"V3": [2,3,4],
"V4": [3,3,3],
"V5": ['a','a','b']}),
table_name="#tt3",
force=True)
print(df1.collect())
print(df2.collect())
print(df3.collect())
dfs = [df1.set_index("ID"), df2.set_index("ID"), df3.set_index("ID")]
print(dfs[0].join(dfs[1:]).collect())
dfs = [df1.set_index(["ID", "ID2"]), df2.set_index(["ID", "ID2"]), df3.set_index(["ID", "ID2"])]
print(dfs[0].join(dfs[1:]).collect())
print(dfs[0].union([dfs[0], dfs[0]]).collect())
```
**Sort by index**
```
df1.sort_index().collect()
```
**Take min, max, sum, median, mean**
```
df1.min()
df1.select("V1").min()
df1.max()
df1.sum()
df1.median()
df1.mean()
```
**Value counts**
```
df3.value_counts().collect()
```
**Split column**
```
import pandas as pd
split_df = \
dataframe.create_dataframe_from_pandas(connection_context,
pandas_df=pd.DataFrame({"ID": [1,2],
"COL": ['1,2,3', '3,4,4']}),
table_name="#split_test",
force=True)
new_df = split_df.split_column(column="COL", separator=",", new_column_names=["COL1", "COL2", "COL3"])
new_df.collect()
```
**Concat columns**
```
new_df.concat_columns(columns=["COL1", "COL2", "COL3"], separator=",").collect()
```
## Close the connection
```
connection_context.close()
```
## Thank you!
| github_jupyter |
# Lumped Elements Circuits
In this notebook, we construct various network from basic lumped elements (resistor, capacitor, inductor), with the 'classic' and the `Circuit` approach. Generally the `Circuit` approach is more verbose than the 'classic' way for building a circuit. However, as the circuit complexity increases, in particular when components are connected in parallel, the `Circuit` approach is interesting as it increases the readability of the code. Moreover, `Circuit` object can be plotted using its `plot_graph()` method, which is usefull to rapidly control if the circuit is built as expected.
```
import numpy as np
import scipy
import matplotlib.pyplot as plt
import skrf as rf
rf.stylely()
```
## LC Series Circuit
In this section we reproduce a simple equivalent model of a capacitor $C$, as illustrated by the figure below:

```
# reference LC circuit made in Designer
LC_designer = rf.Network('designer_capacitor_30_80MHz_simple.s2p')
# scikit-rf: manually connecting networks
line = rf.media.DefinedGammaZ0(frequency=LC_designer.frequency, z0=50)
LC_manual = line.inductor(24e-9) ** line.capacitor(70e-12)
# scikit-rf: using Circuit builder
port1 = rf.Circuit.Port(frequency=LC_designer.frequency, name='port1', z0=50)
port2 = rf.Circuit.Port(frequency=LC_designer.frequency, name='port2', z0=50)
line = rf.media.DefinedGammaZ0(frequency=LC_designer.frequency, z0=50)
cap = line.capacitor(70e-12, name='cap')
ind = line.inductor(24e-9, name='ind')
connections = [
[(port1, 0), (cap, 0)],
[(cap, 1), (ind, 0)],
[(ind, 1), (port2, 0)]
]
circuit = rf.Circuit(connections)
LC_from_circuit = circuit.network
# testing the equivalence of the results
print(np.allclose(LC_designer.s, LC_manual.s))
print(np.allclose(LC_designer.s, LC_from_circuit.s))
circuit.plot_graph(network_labels=True, edge_labels=True, port_labels=True)
```
## A More Advanced Equivalent Model
In this section we reproduce an equivalent model of a capacitor $C$, as illustrated by the figure below:

```
# Reference results from ANSYS Designer
LCC_designer = rf.Network('designer_capacitor_30_80MHz_adv.s2p')
# scikit-rf: usual way, but this time this is more tedious to deal with connection and port number
freq = LCC_designer.frequency
line = rf.media.DefinedGammaZ0(frequency=freq, z0=50)
elements1 = line.resistor(1e-2) ** line.inductor(24e-9) ** line.capacitor(70e-12)
elements2 = line.resistor(20e6)
T_in = line.tee()
T_out = line.tee()
ntw = rf.connect(T_in, 0, elements1, 0)
ntw = rf.connect(ntw, 2, elements2, 0)
ntw = rf.connect(ntw, 1, T_out, 1)
ntw = rf.innerconnect(ntw, 1, 2)
LCC_manual = ntw ** line.shunt_capacitor(50e-12)
# scikit-rf: using Circuit builder
freq = LCC_designer.frequency
port1 = rf.Circuit.Port(frequency=freq, name='port1', z0=50)
port2 = rf.Circuit.Port(frequency=freq, name='port2', z0=50)
line = rf.media.DefinedGammaZ0(frequency=freq, z0=50)
cap = line.capacitor(70e-12, name='cap')
ind = line.inductor(24e-9, name='ind')
res_series = line.resistor(1e-2, name='res_series')
res_parallel = line.resistor(20e6, name='res_parallel')
cap_shunt = line.capacitor(50e-12, name='cap_shunt')
ground = rf.Circuit.Ground(frequency=freq, name='ground', z0=50)
connections = [
[(port1, 0), (res_series, 0), (res_parallel, 0)],
[(res_series, 1), (cap, 0)],
[(cap, 1), (ind, 0)],
[(ind, 1), (cap_shunt, 0), (res_parallel, 1), (port2, 0)],
[(cap_shunt, 1), (ground, 0)],
]
circuit = rf.Circuit(connections)
LCC_from_circuit = circuit.network
# testing the equivalence of the results
print(np.allclose(LCC_designer.s, LCC_manual.s))
print(np.allclose(LCC_designer.s, LCC_from_circuit.s))
circuit.plot_graph(network_labels=True, edge_labels=True, port_labels=True)
```
## Pass band filter
Below we construct a pass-band filter, from an example given in [Microwaves101](https://www.microwaves101.com/encyclopedias/lumped-element-filter-calculator):

```
# Reference result calculated from Designer
passband_designer = rf.Network('designer_bandpass_filter_450_550MHz.s2p')
# scikit-rf:
freq = passband_designer.frequency
passband_manual = line.shunt_capacitor(25.406e-12) ** line.shunt_inductor(4.154e-9) ** \
line.capacitor(2.419e-12) ** line.inductor(43.636e-9) ** \
line.shunt_capacitor(25.406e-12) ** line.shunt_inductor(4.154e-9)
# scikit-rf: the filter with the Circuit builder
freq = passband_designer.frequency
line = rf.media.DefinedGammaZ0(frequency=freq)
C1 = line.capacitor(25.406e-12, name='C1')
C2 = line.capacitor(2.419e-12, name='C2')
C3 = line.capacitor(25.406e-12, name='C3')
L1 = line.inductor(4.154e-9, name='L1')
L2 = line.inductor(43.636e-9, name='L2')
L3 = line.inductor(4.154e-9, name='L3')
port1 = rf.Circuit.Port(frequency=freq, name='port1', z0=50)
port2 = rf.Circuit.Port(frequency=freq, name='port2', z0=50)
ground1 = rf.Circuit.Ground(frequency=freq, name='ground1', z0=50)
ground2 = rf.Circuit.Ground(frequency=freq, name='ground2', z0=50)
ground3 = rf.Circuit.Ground(frequency=freq, name='ground3', z0=50)
ground4 = rf.Circuit.Ground(frequency=freq, name='ground4', z0=50)
connections = [
[(port1, 0), (C1, 0), (L1, 0), (C2, 0)],
[(C2, 1), (L2, 0)],
[(L2, 1), (C3, 0), (L3, 0), (port2, 0)],
# grounding must be done on ground ntw having different names
[(C1, 1), (ground1, 0)],
[(C3, 1), (ground2, 0)],
[(L1, 1), (ground3, 0)],
[(L3, 1), (ground4, 0)],
]
circuit = rf.Circuit(connections)
passband_circuit = circuit.network
passband_circuit.name = 'Pass-band circuit'
passband_circuit.plot_s_db(m=0, n=0, lw=2)
passband_circuit.plot_s_db(m=1, n=0, lw=2)
passband_designer.plot_s_db(m=0, n=0, lw=2, ls='-.')
passband_designer.plot_s_db(m=1, n=0, lw=2, ls='-.')
circuit.plot_graph(network_labels=True, port_labels=True, edge_labels=True)
```
| github_jupyter |
# Take a list of all HSW files (edited in notepad++ from google spreadscheet to be "HSW_2021_02_04__18_45_49__04min_59sec__hsamp_64ch_25000sps_a" format and then create a subsequent files named after the date and the extension to be used for processing ( FS03-20210227 justin style).
```
def HSW_text_seperation (animal, text='//10.153.170.3/storage2/fabian/data/raw/'+animal+'/'+animal+'_to_process.txt',base='//10.153.170.3/storage2/fabian/data/raw/'+animal+'/' ):
"""
Take a list of all HSW files (edited in notepad++ from google spreadscheet to be
"HSW_2021_02_04__18_45_49__04min_59sec__hsamp_64ch_25000sps_a" format and then
create a subsequent files named after the date and the extension to be used for
processing ( FS03-20210227 justin style)
"""
last_last='a'
with open(text, 'r') as f: # Open file for read
for line in f: # Read line-by-line
line = line.strip()
print (line)
name = line[4:14]
last = line[-1]
file_name= line [:-2]
print(name)
print (last)
print(file_name)
print(base +animal+'_'+name+'_'+last +'.txt')
justin= animal +"-"+name.replace("_", "")
print(justin)
if last_last== "a":
with open( base +justin+last+'.txt',"a+", encoding = 'utf' ) as file:
file.write("%s\n" %(file_name))
last_last='a'
import os
animal = 'FS04'
text = '//10.153.170.3/storage2/fabian/data/raw/'+animal+'/'+animal+'_to_process.txt'
base = '//10.153.170.3/storage2/fabian/data/raw/'+animal+'/'
```
### Load the large txt file
```
f = open(text, 'r')
line = f.readline()
print (line)
f.close ()
last_last='a'
with open(text, 'r') as f: # Open file for read
for line in f: # Read line-by-line
line = line.strip()
print (line)
name = line[4:14]
last = line[-1]
file_name= line [:-2]
print(name)
print (last)
print(file_name)
print(base +animal+'_'+name+'_'+last +'.txt')
justin= animal +"-"+name.replace("_", "")
print(justin)
if last_last== "a":
with open( base +justin+last+'.txt',"a+", encoding = 'utf' ) as file:
file.write("%s\n" %(file_name))
last_last='a'
Code GRAVEYARD:
import codecs
#read input file
with codecs.open('//10.153.170.3/storage2/fabian/Data/raw/FS03/FS03_2021_02_26_a.txt', 'r', encoding = 'latin-1') as file:
lines = file.read()
#write output file
with codecs.open('//10.153.170.3/storage2/fabian/Data/raw/FS03/FS03_2021_02_26_a_trial.txt', 'w', encoding = 'utf_8_sig') as file:
file.write(lines)
import fnmatch
import os
for file_name in os.listdir('//10.153.170.3/storage2/fabian/data/Tracking_python/RDM_test_98'):
if fnmatch.fnmatch(file_name, 'BPositions_*'):
print(file_name)
for dirpath, dirnames, files in os.walk('.', topdown=False):
print(f'Found directory: {dirpath}')
for file_name in files:
print(file_name)
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
type(trainset.targets)
type(trainset.data)
# trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
# testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
is_fg = [np.where(np.array(trainset.targets)==fg1)[0] , np.where(np.array(trainset.targets)==fg2)[0], np.where(np.array(trainset.targets)==fg3)[0] ]
# print(is_fg)
is_fg = np.concatenate(is_fg,axis=0)
print(is_fg, (is_fg).shape)
trainset.data = trainset.data[is_fg]
trainset.data.shape
trainset.targets = np.array(trainset.targets)[is_fg]
trainset.targets.shape
is_fg = [np.where(np.array(testset.targets)==fg1)[0] , np.where(np.array(testset.targets)==fg2)[0], np.where(np.array(testset.targets)==fg3)[0] ]
# print(is_fg)
is_fg = np.concatenate(is_fg,axis=0)
print(is_fg, (is_fg).shape)
testset.data = testset.data[is_fg]
testset.data.shape
testset.targets = np.array(testset.targets)[is_fg]
testset.targets.shape
np.unique(np.array(testset.targets)) , np.unique(np.array(trainset.targets))
trainloader = torch.utils.data.DataLoader(trainset, batch_size=256,shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=256,shuffle=False)
import torch.nn as nn
import torch.nn.functional as F
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=16, kernel_size=3, padding=0)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(in_channels=16, out_channels=32, kernel_size=3, padding=0)
# self.conv3 = nn.Conv2d(in_channels=32, out_channels=32, kernel_size=3, padding=0)
self.fc1 = nn.Linear(5408, 256)
self.fc2 = nn.Linear(256, 64)
self.fc3 = nn.Linear(64, 10)
self.fc4 = nn.Linear(10,3)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
# x = self.pool(F.relu(self.conv2(x)))
# print(x.shape)
x = (F.relu(self.conv2(x)))
x = x.view(x.size(0), -1)
# print(x.shape)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
what_net = CNN()#.double()
what_net = what_net.to("cuda")
what_net
import torch.optim as optim
criterion_what = nn.CrossEntropyLoss()
optimizer_what = optim.SGD(what_net.parameters(), lr=0.01, momentum=0.9)
acti = []
loss_curi = []
epochs = 1000
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_what.zero_grad()
# forward + backward + optimize
outputs = what_net(inputs)
loss = criterion_what(outputs, labels)
loss.backward()
optimizer_what.step()
# print statistics
running_loss += loss.item()
mini_batch = 50
if i % mini_batch == mini_batch-1: # print every 50 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, i + 1, running_loss / mini_batch))
ep_lossi.append(running_loss/mini_batch) # loss per minibatch
running_loss = 0.0
if(np.mean(ep_lossi) <= 0.005):
break;
loss_curi.append(np.mean(ep_lossi)) #loss per epoch
print('Finished Training')
torch.save(what_net.state_dict(),"/content/drive/My Drive/Research/Cheating_data/Classify_net_weights/classify_net_2layer_cnn_16_32"+".pt")
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = what_net(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the %d train images: %d %%' % (total, 100 * correct / total))
print(total,correct)
correct = 0
total = 0
out = []
pred = []
what_net.eval()
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= what_net(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print(total,correct)
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from numpy.random import permutation
from shutil import unpack_archive
from itertools import permutations
from sklearn.decomposition import PCA
from sklearn.model_selection import train_test_split
from xgboost import XGBClassifier
train_values = pd.read_csv(r'/Users/sabbrielle/Documents/Lambda/train_values.csv', index_col='sequence_id')
train_labels = pd.read_csv(r'/Users/sabbrielle/Documents/Lambda/train_labels.csv', index_col='sequence_id')
test_values = pd.read_csv(r'/Users/sabbrielle/Documents/Lambda/test_values.csv', index_col='sequence_id')
sequence_lengths = train_values.sequence.apply(len)
train_values.iloc[:, 1:].apply(pd.value_counts)
sorted_binary_features = train_values.iloc[:, 1:].mean().sort_values()
lab_ids = pd.DataFrame(train_labels.idxmax(axis=1), columns=['lab_id'])
bases = set(''.join(train_values.sequence.values))
subsequences = [''.join(permutation) for permutation in permutations(bases, r=4)]
def get_ngram_features(data, subsequences):
"""Generates counts for each subsequence.
Args:
data (DataFrame): The data you want to create features from. Must include a "sequence" column.
subsequences (list): A list of subsequences to count.
Returns:
DataFrame: A DataFrame with one column for each subsequence.
"""
features = pd.DataFrame(index=data.index)
for subseq in subsequences:
features[subseq] = data.sequence.str.count(subseq)
return features
ngram_features = get_ngram_features(train_values, subsequences)
all_features = ngram_features.join(train_values.drop('sequence', axis=1))
def top10_accuracy_scorer(estimator, X, y):
"""A custom scorer that evaluates a model on whether the correct label is in
the top 10 most probable predictions.
Args:
estimator (sklearn estimator): The sklearn model that should be evaluated.
X (numpy array): The validation data.
y (numpy array): The ground truth labels.
Returns:
float: Accuracy of the model as defined by the proportion of predictions
in which the correct label was in the top 10. Higher is better.
"""
# predict the probabilities across all possible labels for rows in our training set
probas = estimator.predict_proba(X)
# get the indices for top 10 predictions for each row; these are the last ten in each row
# Note: We use argpartition, which is O(n), vs argsort, which uses the quicksort algorithm
# by default and is O(n^2) in the worst case. We can do this because we only need the top ten
# partitioned, not in sorted order.
# Documentation: https://numpy.org/doc/1.18/reference/generated/numpy.argpartition.html
top10_idx = np.argpartition(probas, -10, axis=1)[:, -10:]
# index into the classes list using the top ten indices to get the class names
top10_preds = estimator.classes_[top10_idx]
# check if y-true is in top 10 for each set of predictions
mask = top10_preds == y.reshape((y.size, 1))
# take the mean
top_10_accuracy = mask.any(axis=1).mean()
return top_10_accuracy
X=all_features
y=lab_ids.values.ravel()
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=42)
pca_whiten = PCA(n_components=3, random_state=42, whiten=True)
X_train_pca_2 = X_train.copy()
X_val_pca_2 = X_val.copy()
princ_comp_train_whitened = pca.fit_transform(X_train_pca_2)
princ_comp_val_whitened = pca.transform(X_val_pca_2)
pca_train_whitened = pd.DataFrame(data = princ_comp_train_whitened,
columns = ['PC1', 'PC2','PC3'],
index=X_train.index)
pca_val_whitened = pd.DataFrame(data = princ_comp_val_whitened,
columns = ['PC1', 'PC2','PC3'],
index=X_val.index)
X_train_w_pca_whitened = pd.concat([X_train, pca_train_whitened], axis=1)
X_val_w_pca_whitened = pd.concat([X_val, pca_val_whitened], axis=1)
model_xgb_4_w_pca = XGBClassifier(random_state=42,
verbosity=1,
n_jobs=-1,
max_delta_step=1)
model_xgb_4_w_pca.fit(X_train_w_pca_whitened, y_train)
print('Training accuracy using XGBoost and 3 (whitened) PCA:', top10_accuracy_scorer(model_xgb_4_w_pca, X_train_w_pca_whitened, y_train))
print('Validation accuracy using XGBoost and 3 (whitened) PCA:', top10_accuracy_scorer(model_xgb_4_w_pca, X_val_w_pca_whitened, y_val))
```
| github_jupyter |
A simple graphical frontend for Libsvm mainly intended for didactic purposes. You can create data points by point and click and visualize the decision region induced by different kernels and parameter settings.
To create positive examples click the left mouse button; to create negative examples click the right button.
If all examples are from the same class, it uses a one-class SVM.
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
This tutorial imports [dump_svmlight_file](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.dump_svmlight_file.html#sklearn.datasets.dump_svmlight_file)
```
from __future__ import division, print_function
print(__doc__)
import matplotlib
matplotlib.use('TkAgg')
from matplotlib.backends.backend_tkagg import FigureCanvasTkAgg
from matplotlib.backends.backend_tkagg import NavigationToolbar2TkAgg
from matplotlib.figure import Figure
from matplotlib.contour import ContourSet
try:
import tkinter as Tk
except ImportError:
# Backward compat for Python 2
import Tkinter as Tk
import sys
import numpy as np
from sklearn import svm
from sklearn.datasets import dump_svmlight_file
from sklearn.externals.six.moves import xrange
```
### Calculations
```
y_min, y_max = -50, 50
x_min, x_max = -50, 50
class Model(object):
"""The Model which hold the data. It implements the
observable in the observer pattern and notifies the
registered observers on change event.
"""
def __init__(self):
self.observers = []
self.surface = None
self.data = []
self.cls = None
self.surface_type = 0
def changed(self, event):
"""Notify the observers. """
for observer in self.observers:
observer.update(event, self)
def add_observer(self, observer):
"""Register an observer. """
self.observers.append(observer)
def set_surface(self, surface):
self.surface = surface
def dump_svmlight_file(self, file):
data = np.array(self.data)
X = data[:, 0:2]
y = data[:, 2]
dump_svmlight_file(X, y, file)
class Controller(object):
def __init__(self, model):
self.model = model
self.kernel = Tk.IntVar()
self.surface_type = Tk.IntVar()
# Whether or not a model has been fitted
self.fitted = False
def fit(self):
print("fit the model")
train = np.array(self.model.data)
X = train[:, 0:2]
y = train[:, 2]
C = float(self.complexity.get())
gamma = float(self.gamma.get())
coef0 = float(self.coef0.get())
degree = int(self.degree.get())
kernel_map = {0: "linear", 1: "rbf", 2: "poly"}
if len(np.unique(y)) == 1:
clf = svm.OneClassSVM(kernel=kernel_map[self.kernel.get()],
gamma=gamma, coef0=coef0, degree=degree)
clf.fit(X)
else:
clf = svm.SVC(kernel=kernel_map[self.kernel.get()], C=C,
gamma=gamma, coef0=coef0, degree=degree)
clf.fit(X, y)
if hasattr(clf, 'score'):
print("Accuracy:", clf.score(X, y) * 100)
X1, X2, Z = self.decision_surface(clf)
self.model.clf = clf
self.model.set_surface((X1, X2, Z))
self.model.surface_type = self.surface_type.get()
self.fitted = True
self.model.changed("surface")
def decision_surface(self, cls):
delta = 1
x = np.arange(x_min, x_max + delta, delta)
y = np.arange(y_min, y_max + delta, delta)
X1, X2 = np.meshgrid(x, y)
Z = cls.decision_function(np.c_[X1.ravel(), X2.ravel()])
Z = Z.reshape(X1.shape)
return X1, X2, Z
def clear_data(self):
self.model.data = []
self.fitted = False
self.model.changed("clear")
def add_example(self, x, y, label):
self.model.data.append((x, y, label))
self.model.changed("example_added")
# update decision surface if already fitted.
self.refit()
def refit(self):
"""Refit the model if already fitted. """
if self.fitted:
self.fit()
class View(object):
"""Test docstring. """
def __init__(self, root, controller):
f = Figure()
ax = f.add_subplot(111)
ax.set_xticks([])
ax.set_yticks([])
ax.set_xlim((x_min, x_max))
ax.set_ylim((y_min, y_max))
canvas = FigureCanvasTkAgg(f, master=root)
canvas.show()
canvas.get_tk_widget().pack(side=Tk.TOP, fill=Tk.BOTH, expand=1)
canvas._tkcanvas.pack(side=Tk.TOP, fill=Tk.BOTH, expand=1)
canvas.mpl_connect('button_press_event', self.onclick)
toolbar = NavigationToolbar2TkAgg(canvas, root)
toolbar.update()
self.controllbar = ControllBar(root, controller)
self.f = f
self.ax = ax
self.canvas = canvas
self.controller = controller
self.contours = []
self.c_labels = None
self.plot_kernels()
def plot_kernels(self):
self.ax.text(-50, -60, "Linear: $u^T v$")
self.ax.text(-20, -60, "RBF: $\exp (-\gamma \| u-v \|^2)$")
self.ax.text(10, -60, "Poly: $(\gamma \, u^T v + r)^d$")
def onclick(self, event):
if event.xdata and event.ydata:
if event.button == 1:
self.controller.add_example(event.xdata, event.ydata, 1)
elif event.button == 3:
self.controller.add_example(event.xdata, event.ydata, -1)
def update_example(self, model, idx):
x, y, l = model.data[idx]
if l == 1:
color = 'w'
elif l == -1:
color = 'k'
self.ax.plot([x], [y], "%so" % color, scalex=0.0, scaley=0.0)
def update(self, event, model):
if event == "examples_loaded":
for i in xrange(len(model.data)):
self.update_example(model, i)
if event == "example_added":
self.update_example(model, -1)
if event == "clear":
self.ax.clear()
self.ax.set_xticks([])
self.ax.set_yticks([])
self.contours = []
self.c_labels = None
self.plot_kernels()
if event == "surface":
self.remove_surface()
self.plot_support_vectors(model.clf.support_vectors_)
self.plot_decision_surface(model.surface, model.surface_type)
self.canvas.draw()
def remove_surface(self):
"""Remove old decision surface."""
if len(self.contours) > 0:
for contour in self.contours:
if isinstance(contour, ContourSet):
for lineset in contour.collections:
lineset.remove()
else:
contour.remove()
self.contours = []
def plot_support_vectors(self, support_vectors):
"""Plot the support vectors by placing circles over the
corresponding data points and adds the circle collection
to the contours list."""
cs = self.ax.scatter(support_vectors[:, 0], support_vectors[:, 1],
s=80, edgecolors="k", facecolors="none")
self.contours.append(cs)
def plot_decision_surface(self, surface, type):
X1, X2, Z = surface
if type == 0:
levels = [-1.0, 0.0, 1.0]
linestyles = ['dashed', 'solid', 'dashed']
colors = 'k'
self.contours.append(self.ax.contour(X1, X2, Z, levels,
colors=colors,
linestyles=linestyles))
elif type == 1:
self.contours.append(self.ax.contourf(X1, X2, Z, 10,
cmap=matplotlib.cm.bone,
origin='lower', alpha=0.85))
self.contours.append(self.ax.contour(X1, X2, Z, [0.0], colors='k',
linestyles=['solid']))
else:
raise ValueError("surface type unknown")
class ControllBar(object):
def __init__(self, root, controller):
fm = Tk.Frame(root)
kernel_group = Tk.Frame(fm)
Tk.Radiobutton(kernel_group, text="Linear", variable=controller.kernel,
value=0, command=controller.refit).pack(anchor=Tk.W)
Tk.Radiobutton(kernel_group, text="RBF", variable=controller.kernel,
value=1, command=controller.refit).pack(anchor=Tk.W)
Tk.Radiobutton(kernel_group, text="Poly", variable=controller.kernel,
value=2, command=controller.refit).pack(anchor=Tk.W)
kernel_group.pack(side=Tk.LEFT)
valbox = Tk.Frame(fm)
controller.complexity = Tk.StringVar()
controller.complexity.set("1.0")
c = Tk.Frame(valbox)
Tk.Label(c, text="C:", anchor="e", width=7).pack(side=Tk.LEFT)
Tk.Entry(c, width=6, textvariable=controller.complexity).pack(
side=Tk.LEFT)
c.pack()
controller.gamma = Tk.StringVar()
controller.gamma.set("0.01")
g = Tk.Frame(valbox)
Tk.Label(g, text="gamma:", anchor="e", width=7).pack(side=Tk.LEFT)
Tk.Entry(g, width=6, textvariable=controller.gamma).pack(side=Tk.LEFT)
g.pack()
controller.degree = Tk.StringVar()
controller.degree.set("3")
d = Tk.Frame(valbox)
Tk.Label(d, text="degree:", anchor="e", width=7).pack(side=Tk.LEFT)
Tk.Entry(d, width=6, textvariable=controller.degree).pack(side=Tk.LEFT)
d.pack()
controller.coef0 = Tk.StringVar()
controller.coef0.set("0")
r = Tk.Frame(valbox)
Tk.Label(r, text="coef0:", anchor="e", width=7).pack(side=Tk.LEFT)
Tk.Entry(r, width=6, textvariable=controller.coef0).pack(side=Tk.LEFT)
r.pack()
valbox.pack(side=Tk.LEFT)
cmap_group = Tk.Frame(fm)
Tk.Radiobutton(cmap_group, text="Hyperplanes",
variable=controller.surface_type, value=0,
command=controller.refit).pack(anchor=Tk.W)
Tk.Radiobutton(cmap_group, text="Surface",
variable=controller.surface_type, value=1,
command=controller.refit).pack(anchor=Tk.W)
cmap_group.pack(side=Tk.LEFT)
train_button = Tk.Button(fm, text='Fit', width=5,
command=controller.fit)
train_button.pack()
fm.pack(side=Tk.LEFT)
Tk.Button(fm, text='Clear', width=5,
command=controller.clear_data).pack(side=Tk.LEFT)
def get_parser():
from optparse import OptionParser
op = OptionParser()
op.add_option("--output",
action="store", type="str", dest="output",
help="Path where to dump data.")
return op
def main(argv):
op = get_parser()
opts, args = op.parse_args(argv[1:])
root = Tk.Tk()
model = Model()
controller = Controller(model)
root.wm_title("Scikit-learn Libsvm GUI")
view = View(root, controller)
model.add_observer(view)
Tk.mainloop()
if opts.output:
model.dump_svmlight_file(opts.output)
if __name__ == "__main__":
main(sys.argv)
```
### License
Author:
Peter Prettenhoer <peter.prettenhofer@gmail.com>
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Libsvm.ipynb', 'scikit-learn/svm-gui/', 'Libsvm GUI| plotly',
' ',
title = 'Libsvm GUI',
name = 'Libsvm GUI',
has_thumbnail='true', thumbnail='thumbnail/scikit-default.jpg',
language='scikit-learn', page_type='example_index',
display_as='real_dataset', order=8,ipynb='~Diksha_Gabha/2662')
```
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fcurriculum-notebooks&branch=master&subPath=Health/CALM/CALM-moving-out-2.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# CALM - Moving Out 2
## Part 2 - Income
📗Now let's take a look at your potential income. First, some definitions:
### Paycheque Definitions
#### Gross Income (pay/earnings)
The amount of income/earnings, for any pay period, before deductions
#### Net income (pay/earnings)
The amount of income/earnings, for any pay period, after deductions (Take home pay)
#### CPP - Canada Pension Plan
2.3% of gross income deducted for insurance in case of unemployment
#### Income Tax
A deduction paid to the Federal and Provincial government for taxes
#### LTD
A deduction for Long Term Disability insurance
#### Union Dues
Fees paid for membership in a union
#### Bonds
An investment in which a business or government pays a set interest rate
#### Advance Earnings
Deducted money that was received in advance of the pay cheque
#### Overtime Earnings
Pay received for working over 8 hours a day or 44 hours a week, whichever is greater
### Calculating Net Income
📗Click on the code cell below, then click the `Run` button on the toolbar to calculate your net income. You may also change some values in the code to see how the results change.
```
#✏️
wagePerHour = 15 # this is your wage in $/hour
hoursPerDay = 8
workdaysPerMonth = 21
grossIncome = wagePerHour * hoursPerDay * workdaysPerMonth
print('Your gross income is $', grossIncome, 'per month.')
incomeTax = .15 + .10 # assume federal income tax is 15% and provincial is 10%
cpp = .0495 # assume Canada Pension Plan is 4.95%
ei = .0188 # assume Employment Insurance is 1.88%
unionDues = .0075 # 0.75% sounds reasonable for union dues
deductions = grossIncome * (incomeTax + cpp + ei + unionDues)
print('$', '{:.2f}'.format(deductions), ' will be taken off your paycheck.')
netIncome = grossIncome - deductions
print('Your net income is $', '{:.2f}'.format(netIncome), 'per month.')
%store netIncome # store that value in memory for later notebooks
```
## Graphing Income
📗
We can also look at how your net income (take-home pay) changes based on your hourly wage. We will use [2019 taxation rates](https://www.canada.ca/en/revenue-agency/services/tax/individuals/frequently-asked-questions-individuals/canadian-income-tax-rates-individuals-current-previous-years.html) as well as [EI](https://www.canada.ca/en/revenue-agency/services/tax/businesses/topics/payroll/payroll-deductions-contributions/employment-insurance-ei/ei-premium-rates-maximums.html) and [CPP](https://www.canada.ca/en/revenue-agency/services/tax/businesses/topics/payroll/payroll-deductions-contributions/canada-pension-plan-cpp/cpp-contribution-rates-maximums-exemptions.html) rates. `Run` the cell below (without editing it) to generate a graph.
</div>
```
#📗
def calculateFederalTax(income):
taxBrackets = [47630, 95259, 147667, 210371]
taxRates = [.15, .205, .26, .29, .33]
taxes = []
for i in range(0, len(taxBrackets)):
taxes.append(taxBrackets[i] * taxRates[i])
if income < taxBrackets[0]:
tax = income * taxRates[0]
elif income < taxBrackets[1]:
tax = taxes[0] + (income - taxBrackets[0]) * taxRates[1]
elif income < taxBrackets[2]:
tax = taxes[1] + (income - taxBrackets[1]) * taxRates[2]
elif income < taxBrackets[3]:
tax = taxes[2] + (income - taxBrackets[2]) * taxRates[3]
else:
tax = taxes[3] + (income - taxBrackets[3]) * taxRates[4]
return tax
def calculateProvincialTax(income):
taxBrackets = [131220, 157464, 209952, 314928] # for Alberta
taxRates = [.1, .12, .13, .14, .15]
taxes = []
for i in range(0, len(taxBrackets)):
taxes.append(taxBrackets[i] * taxRates[i])
if income < taxBrackets[0]:
tax = income * taxRates[0]
elif income < taxBrackets[1]:
tax = taxes[0] + (income - taxBrackets[0]) * taxRates[1]
elif income < taxBrackets[2]:
tax = taxes[1] + (income - taxBrackets[1]) * taxRates[2]
elif income < taxBrackets[3]:
tax = taxes[2] + (income - taxBrackets[2]) * taxRates[3]
else:
tax = taxes[3] + (income - taxBrackets[3]) * taxRates[4]
return tax
def calculateEI(income):
eiMaxInsurableEarnings = 53100
eiRate = 0.0162
if income >= eiMaxInsurableEarnings:
eiPremium = eiMaxInsurableEarnings * eiRate
else:
eiPremium = income * eiRate
return eiPremium
def calculateCPP(income):
cppMaxContributoryEarnings = 53900
cppRate = 0.051
if income >= cppMaxContributoryEarnings:
cppPremium = cppMaxContributoryEarnings * cppRate
else:
cppPremium = income * cppRate
return cppPremium
wages = []
grossIncomes = []
netIncomes = []
for wage in range(15, 150):
wages.append(wage)
grossAnnualIncome = wage * 8 * 240
grossIncomes.append(grossAnnualIncome)
incomeTax = calculateFederalTax(grossAnnualIncome) + calculateProvincialTax(grossAnnualIncome)
eiPremium = calculateEI(grossAnnualIncome)
cppPremium = calculateCPP(grossAnnualIncome)
netIncome = grossAnnualIncome - (incomeTax + eiPremium + cppPremium)
netIncomes.append(netIncome)
import plotly.graph_objects as go
fig = go.Figure()
fig.add_trace(go.Scatter(x=wages, y=grossIncomes, name='Gross Income'))
fig.add_trace(go.Scatter(x=wages, y=netIncomes, name='Net Income'))
fig.update_layout(
title=go.layout.Title(text='Net Income vs Hourly Wage'),
xaxis=go.layout.XAxis(title=go.layout.xaxis.Title(text='Hourly Wage')),
yaxis=go.layout.YAxis(title=go.layout.yaxis.Title(text='Income')))
fig.show()
```
📗The graph shows that the difference between gross income and net income (after deductions) increases as wage increases. For more information about this, you may want to read about [progressive taxation](https://en.wikipedia.org/wiki/Progressive_tax).
You have now completed this section. Proceed to [section 3](./CALM-moving-out-3.ipynb)
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
```
import sys
import itertools
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from math import sqrt
from sklearn.metrics import mean_squared_error
from pandas import datetime
from pandas.tools.plotting import autocorrelation_plot
from statsmodels.tsa.api import ExponentialSmoothing, ARIMA, ARMA, arma_order_select_ic
from statsmodels.tsa.statespace.sarimax import SARIMAX
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.stattools import adfuller
from statsmodels.stats.diagnostic import acorr_ljungbox
from statsmodels.tsa.seasonal import seasonal_decompose
%matplotlib inline
# 设置图片参数
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文
plt.rcParams['axes.unicode_minus'] = False # 用来正常显示负号
plt.rcParams['figure.figsize'] = (15, 8) # 图片大小
plt.rcParams['savefig.dpi'] = 300 # 图片像素
# plt.rcParams['figure.dpi'] = 300 # 分辨率
def error_prop(y_true, y_pred):
return np.sum(np.abs(y_true - y_pred)) / np.sum(y_true)
def parse_date(str):
return datetime.strptime(str, '%Y-%m-%d')
```
## 1 数据预处理
```
# 加载数据
data_path = "../../data/luggage_compartment_door.txt"
df = pd.read_csv(data_path, sep='\t')
# 重新命名列名
df.rename(columns={'物料编码': 'part_id', '物料描述': 'part_name',
'订货数': 'order_num', '缺件数': 'out_of_stock_num',
'受理数': 'delivery_num', '审核日期': 'date', '审核时间': 'time'}, inplace=True)
# 将`part_id`的数据类型设为字符串,方便后面进行重采样
df['part_id'] = df['part_id'].astype('str')
# 重置索引
# df['date'] = pd.to_datetime(df['date'], format="%Y-%m-%d")
df['date'] = df['date'].apply(lambda x: datetime.strptime(x, '%Y-%m-%d'))
df.set_index('date', inplace=True)
# 按照时间排序
df.sort_index(inplace=True)
# 重采样
df_day = df.resample('D').sum()
# 截取2018年的数据
df_2018 = df_day['2018']
fig, ax = plt.subplots()
ax.plot(df_2018.order_num, 'bo-')
ax.set_xlabel('日期(天)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('行李舱门本体总成每天订货量', fontsize=20)
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
```
## 2 主要是星期几订货?
```
df_2018['weekday_name'] = df_2018.index.weekday_name
no_order_day = df_2018[df_2018.order_num == 0.0]
print("2018年没有订货的天数为:%s天" % len(no_order_day))
no_order_counts = no_order_day.weekday_name.value_counts()
no_order_counts / len(no_order_day)
no_order_counts.plot(kind='bar', figsize=(12, 5), fontsize=12)
```
主要是星期六天没有订货,占比 **62.5%**。
```
order_day = df_2018[df_2018.order_num > 0.0]
print("2018年里订货的天数为:%s" % (len(order_day)))
order_counts = order_day.weekday_name.value_counts()
order_counts / len(order_day)
order_counts.plot(kind='bar', figsize=(12, 5), fontsize=12)
```
<u>**2018年里的订货日期比较平均**</u>,除了星期天略少一些。
## 3 较大的订货量主要发生在星期几?
```
big_order_day = df_2018[df_2018.order_num >= 40]
print("2018年订货数非常大的天数有 %s 天" % len(big_order_day))
big_order_counts = big_order_day.weekday_name.value_counts()
big_order_counts / len(big_order_day)
big_order_counts.plot(kind='bar', figsize=(12, 5), fontsize=12)
```
## 建模探索
```
print("2018年到目前为止总的订货量:%s" % order_day.order_num.sum())
print("2018年有订货的日期中平均每天订货数:%s" % order_day.order_num.mean())
print("2018年有订货的日期中订货数的中位数:%s" % order_day.order_num.median())
df_201807_to_201809 = df_2018['2018-07':]
fig, ax = plt.subplots()
ax.plot(df_201807_to_201809.order_num, 'ro-', label='订货数')
# ax.plot(df_201807_to_201809.out_of_stock_num, 'go-', label='缺货数')
# ax.plot(df_201807_to_201809.delivery_num, 'bo-', label='受理数')
ax.set_xlabel('日期(天)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('行李舱门本体总成每天订货量', fontsize=20)
ax.legend(loc='upper left')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
train_day, test_day = df_201807_to_201809[:-7], df_201807_to_201809[-7:]
```
## 2 时间序列检验
### 2.1 自相关图和偏自相关图
```
# 天的自相关图
plot_acf(df_201807_to_201809.order_num.values).show()
plot_pacf(df_201807_to_201809.order_num.values).show()
autocorrelation_plot(df_201807_to_201809.order_num.values)
plt.show()
```
### 2.2 单位根检验
```
adf_result = adfuller(df_201807_to_201809.order_num.values)
output = pd.DataFrame(index=["Test Statistic Value", "p-value", "Lags Used", "Number of Observations Used", "Critical Value (1%)", "Critical Value (5%)", "Critical Value (10%)"], columns=['value'])
output['value']['Test Statistic Value'] = adf_result[0]
output['value']['p-value'] = adf_result[1]
output['value']['Lags Used'] = adf_result[2]
output['value']['Number of Observations Used'] = adf_result[3]
output['value']['Critical Value (1%)'] = adf_result[4]['1%']
output['value']['Critical Value (5%)'] = adf_result[4]['5%']
output['value']['Critical Value (10%)'] = adf_result[4]['10%']
print("单位根检验结果为:")
print(output)
```
### 2.3 纯随机性检验
```
print("序列的纯随机性检验结果为:", end='')
print(acorr_ljungbox(df_201807_to_201809.order_num.values, lags=1)[1][0])
```
## 3 模型拟合
### 3.1 Holt-Winters method
```
train, test = train_day.order_num.values, test_day.order_num.values
model = ExponentialSmoothing(train, seasonal_periods=7, trend='add', seasonal='add')
model_fit = model.fit()
preds = model_fit.forecast(7)
fig, ax = plt.subplots()
# ax.plot(train_day.index, train_day.order_num.values, 'go-', label='Train')
ax.plot(test_day.index, test, 'bo-', label='Test')
ax.plot(test_day.index, preds, 'ro-', label='Holt-Winters method')
ax.set_xlabel('日期(天)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('Holt-Winters方法预测每天订货量', fontsize=20)
ax.legend(loc='best')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
plt.show()
rmse = sqrt(mean_squared_error(test, preds))
print("The RMSE of 'Holt-Winters method' is:", rmse)
print("The error proportion of Holt-Winters is:", error_prop(test, preds))
```
### 3.2 ARIMA
```
train, test = train_day.order_num.values, test_day.order_num.values
# pmax = len(df_day) // 10
# qmax = len(df_day) // 10
pmax = 6
qmax = 6
bic_matrix = []
for p in range(pmax + 1):
temp = []
for q in range(qmax + 1):
try:
temp.append(ARIMA(train, (p, 0, q)).fit().aic)
except:
temp.append(None)
bic_matrix.append(temp)
bic_matrix = pd.DataFrame(bic_matrix).astype('float').dropna(axis=1)
p, q = bic_matrix.stack().idxmin()
print("AIC最小的(p, q)值为:(%s, %s)" % (p, q))
bic_matrix = []
for p in range(pmax + 1):
temp = []
for q in range(qmax + 1):
try:
temp.append(ARIMA(train, (p, 0, q)).fit().aic)
except:
temp.append(None)
bic_matrix.append(temp)
bic_matrix = pd.DataFrame(bic_matrix).astype('float').dropna(axis=1)
p, q = bic_matrix.stack().idxmin()
print("AIC最小的(p, q)值为:(%s, %s)" % (p, q))
arma_order_select_ic(train, max_ar=6, max_ma=6, ic='aic')['aic_min_order']
history = list(train)
preds = list()
for i in range(len(test)):
model = ARMA(history, order=(2, 1))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
preds.append(yhat[0])
obs = test[i]
history.append(obs)
print("expected = %f predicted = %f" % (obs, yhat))
error = sqrt(mean_squared_error(test, preds))
print("Test RMSE: %.3f" % error)
temp = np.array(preds).reshape((len(test), ))
print("The error proportion of ARIMA is:", error_prop(test, temp))
fig, ax = plt.subplots()
ax.plot(test_day.index, test, 'bo-', label='test')
ax.plot(test_day.index, preds, 'ro-', label='pred')
ax.set_xlabel('日期(天)', fontsize=16)
ax.set_ylabel('销量(件)', fontsize=16)
ax.set_title('ARIMA预测行李舱门本体总成每天订货量', fontsize=20)
ax.legend(loc='upper left')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
fig.savefig("../../figs/ARIMA预测图.png")
plt.show()
```
### 3.3 SARIMAX
```
train, test = train_day.order_num.values, test_day.order_num.values
# define the p, d and q parameters to take any value between 0 and 2
p = d = q = range(0, 6)
# generate all different combinations of p, d and q triplets
pdq = list(itertools.product(p, d, q))
# generate all different combinations of seasonal p, q and q triplets
seasonal_pdq = [(x[0], x[1], x[2], 1) for x in pdq]
best_aic = np.inf
best_pdq = None
best_seasonal_pdq = None
tmp_model = None
best_mdl = None
for param in pdq:
for param_seasonal in seasonal_pdq:
try:
tmp_mdl = SARIMAX(train,
order = param,
seasonal_order = param_seasonal,
enforce_stationarity=True,
enforce_invertibility=True)
res = tmp_mdl.fit()
# print("SARIMAX{}x{}12 - AIC:{}".format(param, param_seasonal, results.aic))
if res.aic < best_aic:
best_aic = res.aic
best_pdq = param
best_seasonal_pdq = param_seasonal
best_mdl = tmp_mdl
except:
print("Unexpected error:", sys.exc_info()[0])
continue
print("Best SARIMAX{}x{}12 model - AIC:{}".format(best_pdq, best_seasonal_pdq, best_aic))
history = list(train)
preds = list()
for i in range(len(test)):
model = SARIMAX(train, order=(5, 0, 4), seasonal_order=(0, 1, 1, 7))
model_fit = model.fit(disp=0)
output = model_fit.forecast()
yhat = output[0]
preds.append(yhat)
obs = test[i]
history.append(obs)
print("expected = %f predicted = %f" % (obs, yhat))
error = sqrt(mean_squared_error(test, preds))
print("Test RMSE: %.3f" % error)
fig, ax = plt.subplots()
ax.plot(test_day.index, test, 'bo-', label='test')
ax.plot(test_day.index, preds, 'ro-', label='pred')
ax.set_xlabel('day', fontsize=16)
ax.set_ylabel('sales', fontsize=16)
ax.set_title('ARIMA Rolling Forecast Line Plot', fontsize=20)
ax.legend(loc='upper left')
ax.grid()
# TODO: 改变刻度字体大小不够优雅
plt.xticks(fontsize=12)
plt.yticks(fontsize=12)
fig.savefig("../figs/ARIMA预测图.png")
plt.show()
```
| github_jupyter |
# MNIST With PyTorch Training
## Import Libraries
```
import numpy as np
import torch
import torchvision
import matplotlib.pyplot as plt
from torchvision import datasets, transforms
from torch import nn
from torch import optim
from time import time
import os
from google.colab import drive
```
## Pre-Process Data
Here we download the data using PyTorch data utils and transform the data by using a normalization function. PyTorch provides a data loader abstraction called a `DataLoader` where we can set the batch size, data shuffle per batch loading. Each data loader expecte a Pytorch Dataset. The DataSet abstraction and DataLoader usage can be found [here](https://pytorch.org/tutorials/recipes/recipes/loading_data_recipe.html)
```
# Data transformation function
transform = transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,)),
])
# DataSet
train_data_set = datasets.MNIST('drive/My Drive/mnist/data/', download=True, train=True, transform=transform)
validation_data_set = datasets.MNIST('drive/My Drive/mnist/data/', download=True, train=False, transform=transform)
train_loader = torch.utils.data.DataLoader(train_data_set, batch_size=32, shuffle=True)
validation_loader = torch.utils.data.DataLoader(validation_data_set, batch_size=32, shuffle=True)
```
## Define Network
Here we select the matching input size compared to the network definition.
Here data reshaping or layer reshaping must be done to match input data shape with the network input shape. Also we define a set of hidden unit sizes along with the output layers size. The `output_size` must match with the number of labels associated with the classification problem. The hidden units can be chosesn depending on the problem. `nn.Sequential` is one way to create the network. Here we stack a set of linear layers along with a softmax layer for the classification as the output layer.
```
input_size = 784
hidden_sizes = [128, 128, 64, 64]
output_size = 10
model = nn.Sequential(nn.Linear(input_size, hidden_sizes[0]),
nn.ReLU(),
nn.Linear(hidden_sizes[0], hidden_sizes[1]),
nn.ReLU(),
nn.Linear(hidden_sizes[1], hidden_sizes[2]),
nn.ReLU(),
nn.Linear(hidden_sizes[2], hidden_sizes[3]),
nn.ReLU(),
nn.Linear(hidden_sizes[3], output_size),
nn.LogSoftmax(dim=1))
print(model)
```
## Define Loss Function and Optimizer
Read more about [Loss Functions](https://pytorch.org/docs/stable/nn.html#loss-functions) and [Optimizers](https://pytorch.org/docs/stable/optim.html) supported by PyTorch.
```
criterion = nn.NLLLoss()
optimizer = optim.SGD(model.parameters(), lr=0.003, momentum=0.9)
```
# Model Training
```
epochs = 5
for epoch in range(epochs):
loss_per_epoch = 0
for images, labels in train_loader:
images = images.view(images.shape[0], -1)
# Gradients cleared per batch
optimizer.zero_grad()
# Pass input to the model
output = model(images)
# Calculate loss after training compared to labels
loss = criterion(output, labels)
# backpropagation
loss.backward()
# optimizer step to update the weights
optimizer.step()
loss_per_epoch += loss.item()
average_loss = loss_per_epoch / len(train_loader)
print("Epoch {} - Training loss: {}".format(epoch, average_loss))
```
## Model Evaluation
Similar to training data loader, we use the validation loader to load batch by batch and run the feed-forward network to get the expected prediction and compared to the label associated with the data point.
```
correct_predictions, all_count = 0, 0
# enumerate data from the data validation loader (loads a batch at a time)
for batch_id, (images,labels) in enumerate(validation_loader):
for i in range(len(labels)):
img = images[i].view(1, 784)
# at prediction stage, only feed-forward calculation is required.
with torch.no_grad():
logps = model(img)
# Output layer of the network uses a LogSoftMax layer
# Hence the probability must be calculated with the exponential values.
# The final layer returns an array of probabilities for each label
# Pick the maximum probability and the corresponding index
# The corresponding index is the predicted label
ps = torch.exp(logps)
probab = list(ps.numpy()[0])
pred_label = probab.index(max(probab))
true_label = labels.numpy()[i]
if(true_label == pred_label):
correct_predictions += 1
all_count += 1
print(f"Model Accuracy {(correct_predictions/all_count) * 100} %")
```
### Reference:
1. [Torch NN Sequential](https://pytorch.org/docs/stable/generated/torch.nn.Sequential.html)
2. [Handwritten Digit Recognition Using PyTorch — Intro To Neural Networks](https://towardsdatascience.com/handwritten-digit-mnist-pytorch-977b5338e627)
3. [MNIST Handwritten Digit Recognition in PyTorch](https://nextjournal.com/gkoehler/pytorch-mnist)
| github_jupyter |
# import statements
```
import torch.nn.functional as F
import torch
import numpy as np
from matplotlib import pyplot as plt
%matplotlib inline
from sklearn.metrics import log_loss
import pandas as pd
from torch.nn import BCELoss
#from scipy.optimize import fmin
```
# utils
```
def softmax(x):
"""Compute softmax values for each sets of scores in x."""
m = torch.max(x,dim=1,keepdims=True)
#print(m)
e_x = torch.exp(torch.sub(x,m.values))
return e_x / torch.sum(e_x,dim=1,keepdims=True)
def focus_(a,x):
"""
focus function parametrized by a*x
returns : averaged input for classification function
"""
#print(a*x)
out = softmax(a*x)
#print(out)
out = torch.sum(out*x,dim=1)
return out
def classification_(b,c,x):
"""
classification function parametrized by b*x + c
returns : sigmoid(b*x+c)
"""
out = (b*x) + c
out = 1/(1+torch.exp(-out))
return out
def derv_g(b,c):
"""
derivate of log-loss with respect to b and c using autograd
"""
#print(yhat,y,yhat-y)
#print()
db = b.grad
dc = c.grad
# db = np.dot(xhat,yhat-y)/xhat.shape[0]
# dc = np.sum(yhat-y)/xhat.shape[0]
return db,dc
def derv_f(a):
"""
derivative of log-loss with respect to a using autograd
"""
da = a.grad
# da = np.sum((yhat-y)*b*( ( (x[:,0] - x[:,1])* x[:,0] ) + ( (x[:,1]- x[:,0]) * x[:,1] ) ) * (np.exp((a*x[:,0]+a*x[:,1]))/ (np.exp(a*x[:,0])+ np.exp(a*x[:,1]))**2 )) / xhat.shape[0]
# #print( ( ( (x[:,0] - x[:,1])* x[:,0] ) + ( (x[:,1]- x[:,0]) * x[:,1] ) ) * (np.exp((a*x[:,0]+a*x[:,1]))/ (np.exp(a*x[:,0])+ np.exp(a*x[:,1]))**2 ) )
return da
def gd(w,dw):
"""
updates given parameter in negative direction of gradient
"""
eta = torch.tensor([0.1])
with torch.no_grad():
w = w - torch.dot(eta,dw)
return w
```
# m = 2
```
X = torch.tensor([[-3,-1],[-1,-3],[1,-3],[-3,1]]) # mosaic data m = 2 , d= 1
Y = torch.tensor([0,0,1,1])
def minimize_b_c(x,y,a,b,c,epochs=1000):
# b = 0
# c = 0
#a.requires_grad=False
criterion = BCELoss()
y = y.float()
with torch.no_grad():
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
initial_loss = criterion(yhat,y)
print("x average at 0 epoch", x_average )
print("yhat at 0 epoch",yhat)
print("loss at 0 epoch",criterion(yhat,y).item())
for i in range(epochs):
a.requires_grad = False
b.requires_grad = True
c.requires_grad = True
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
loss = criterion(yhat,y)
b.retain_grad()
c.retain_grad()
loss.backward()
#der_b,der_c = derv_g(b,c)
#print(i,der_b,der_c)
grad_b = b.grad
grad_c = c.grad
b = gd(b,grad_b)
c = gd(c,grad_c)
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
current_loss = criterion(yhat,y)
#print(current_loss<=(initial_loss/2) , current_loss,initial_loss)
if current_loss<= (initial_loss)/2:
break
print(" ")
with torch.no_grad():
x_average = focus_(a,x)
print("x average",x_average)
yhat = classification_(b,c,x_average)
print("Y hat",yhat)
current_loss = criterion(yhat,y)
print("Loss",current_loss.item(),i)
return b,c,current_loss.item()
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
b,c,loss = minimize_b_c(X,Y,a,b,c)
def minimize_a(x,y,a,b,c,epochs=1000):
#b = 0
#c = 0
criterion = BCELoss()
y = y.float()
with torch.no_grad():
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
initial_loss = criterion(yhat,y)
print("x average at 0 epoch", x_average )
print("yhat at 0 epoch",yhat)
print("loss at 0 epoch",criterion(yhat,y).item())
for i in range(epochs):
a.requires_grad =True
b.requires_grad = False
c.requires_grad = False
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
loss = criterion(yhat,y)
a.retain_grad()
loss.backward()
der_a = derv_f(a)
#print(i,der_a)
a = gd(a,der_a)
x_average = focus_(a,x)
yhat = classification_(b,c,x_average)
current_loss = criterion(yhat,y)
if current_loss <= initial_loss/2:
break
print("*"*60)
with torch.no_grad():
x_average = focus_(a,x)
print("x average",x_average)
yhat = classification_(b,c,x_average)
print("Y hat",yhat)
current_loss = log_loss(y,yhat)
print("Loss",current_loss.item(),i)
return a,current_loss.item()
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=False)
c = torch.tensor([0.],requires_grad=False)
a,loss = minimize_a(X,Y,a,b,c)
a = torch.tensor(np.linspace(-1,1,20),requires_grad=True,dtype=torch.float32)
b_list = []
c_list = []
loss_list = []
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
for a1 in a:
#out =focus_(a1,X)
#print(out)
b,c,loss = minimize_b_c(X,Y,a=a1,b=b,c=c)
b_list.append(b.item())
c_list.append(c.item())
#out= classification_(0,0,out)
#print(out)
loss_list.append(loss)
a = np.linspace(-1,1,20)
plt.figure(figsize=(6,5))
plt.plot(a,loss_list,"*-")
plt.grid()
#plt.xticks(a)
plt.xlabel("a")
plt.ylabel("log-loss")
plt.title("loss plot for fix value of a ")
plt.savefig("loss_fixed_a.png")
plt.figure(figsize=(6,5))
plt.plot(a,b_list,"*-")
plt.grid()
#plt.xticks(a)
plt.xlabel("a")
plt.ylabel("b")
plt.title("Minimized value of b for fixed a")
plt.savefig("minimized_b_fixed_a.png")
plt.figure(figsize=(6,5))
plt.plot(a,c_list,"*-")
plt.grid()
plt.xlabel("a")
plt.ylabel("c")
plt.title("Minimized value of c for fixed a")
plt.savefig("minimized_c_fixed_a.png")
loss_ = []
bb,cc= np.meshgrid(np.arange(-21,21,0.2),np.arange(-21,21,0.2))
b_ = bb.reshape(-1,1)
c_ = cc.reshape(-1,1)
a_ = 0
x_average_ = focus_(a_,X)
yhat_ = classification_(b_,c_,x_average_)
#print("Y hat",yhat_)
#Y_ = np.array([list(Y)]*40000)
for i in range(yhat_.shape[0]):
loss_.append(log_loss(Y,yhat_[i]))
loss_ = np.array(loss_)#,axis=0)
plt.figure(figsize=(6,5))
cs = plt.contourf(b_.reshape(bb.shape),c_.reshape(cc.shape),loss_.reshape(bb.shape))
plt.xlabel("b")
plt.ylabel("c")
plt.colorbar(cs)
plt.scatter(0, 0,c="black",s=100)
plt.scatter(15.625194533788827, -1.5129474107626304,c="r",s=100)
plt.title("contour plot for fixed a = "+str(a_) )
plt.savefig("contour_b_c_a_0.png")
minimize_b_c(X,Y,0,0,0)
a = np.linspace(-10,10,5000)
loss = []
for a1 in a:
out =focus_(a1,X)
#print(out)
out = classification_(-10,-10,out)
#print(out)
loss.append(log_loss(Y,out,))
plt.plot(a,loss)
plt.xlabel("a")
plt.ylabel("log-loss")
plt.title("loss plot for fix value of b and c")
plt.savefig("loss_landscape_b_n10_c_n10.png")
```
# Alternate minimization
```
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
criterion = BCELoss()
data = pd.DataFrame(columns=["sno","b_c_fixed","a_fixed","a_value","b_value","c_value","loss"])
Y_ = Y.float()
X_average = focus_(a,X)
Yhat = classification_(b,c,X_average)
initial_loss = criterion(Yhat,Y_)
#print(initial_loss)
k = 0
data.loc[k] = [k,True,True,a.item(),b.item(),c.item(),initial_loss.item()]
k = k+1
j= 1
for i in range(0,40,2):
print("Minimize b and c")
b,c,loss = minimize_b_c(X,Y,a,b,c)
#print(b,c)
data.loc[k] = [j,False,True,a.item(),b.item(),c.item(),loss]
print("*"*60)
print(" ")
print("minimize a")
#print(a,b,c)
a,loss = minimize_a(X,Y,a,b,c)
data.loc[k+1] = [j,True,False,a.item(),b.item(),c.item(),loss]
print(" ")
k = k+2
j = j+1
data
data.to_csv("data_2_m_2_1.csv",index=False)
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
criterion = BCELoss()
data_1 = pd.DataFrame(columns=["sno","b_c_fixed","a_fixed","a_value","b_value","c_value","loss"])
X_average = focus_(a,X)
Yhat = classification_(b,c,X_average)
Y_ = Y.float()
initial_loss = criterion(Yhat,Y_)
k = 0
data_1.loc[k] = [k,True,True,a.item(),b.item(),c.item(),initial_loss.item()]
k = k+1
j= 1
for i in range(0,40,2):
print("minimize a")
a,loss = minimize_a(X,Y,a,b,c)
data_1.loc[k] = [j,True,False,a.item(),b.item(),c.item(),loss]
print("*"*60)
print(" ")
print("Minimize b and c")
b,c,loss = minimize_b_c(X,Y,a,b,c)
data_1.loc[k+1] = [j,False,True,a.item(),b.item(),c.item(),loss]
print(" ")
k = k+2
j = j+1
data_1.to_csv("data_2_m_2_2.csv",index=False)
data_1
```
# m = 9
```
X1 = torch.tensor([[-1,-3,-3,-3,-3,-3,-3,-3,-3],[1,-3,-3,-3,-3,-3,-3,-3,-3],
[-3,-1,-3,-3,-3,-3,-3,-3,-3],[-3,1,-3,-3,-3,-3,-3,-3,-3],
[-3,-3,-1,-3,-3,-3,-3,-3,-3],[-3,-3,1,-3,-3,-3,-3,-3,-3],
[-3,-3,-3,-1,-3,-3,-3,-3,-3],[-3,-3,-3,1,-3,-3,-3,-3,-3],
[-3,-3,-3,-3,-1,-3,-3,-3,-3],[-3,-3,-3,-3,1,-3,-3,-3,-3],
[-3,-3,-3,-3,-3,-1,-3,-3,-3],[-3,-3,-3,-3,-3,1,-3,-3,-3],
[-3,-3,-3,-3,-3,-3,-1,-3,-3],[-3,-3,-3,-3,-3,-3,1,-3,-3],
[-3,-3,-3,-3,-3,-3,-3,-1,-3],[-3,-3,-3,-3,-3,-3,-3,1,-3],
[-3,-3,-3,-3,-3,-3,-3,-3,-1],[-3,-3,-3,-3,-3,-3,-3,-3,1],]) # mosaic data m = 9 , d= 1
Y1 = torch.tensor([0,1,0,1,0,1,0,1,0,1,0,1,0,1,0,1,0,1])
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
criterion = BCELoss()
data = pd.DataFrame(columns=["sno","b_c_fixed","a_fixed","a_value","b_value","c_value","loss"])
Y1_ = Y1.float()
X1_average = focus_(a,X1)
Yhat1 = classification_(b,c,X1_average)
initial_loss = criterion(Yhat1,Y1_)
#print(initial_loss)
k = 0
data.loc[k] = [k,True,True,a.item(),b.item(),c.item(),initial_loss.item()]
k = k+1
j= 1
for i in range(0,40,2):
print("Minimize b and c")
b,c,loss = minimize_b_c(X1,Y1,a,b,c,epochs=20000)
#print(b,c)
data.loc[k] = [j,False,True,a.item(),b.item(),c.item(),loss]
print("*"*60)
print(" ")
print("minimize a")
#print(a,b,c)
a,loss = minimize_a(X1,Y1,a,b,c,epochs=20000)
data.loc[k+1] = [j,True,False,a.item(),b.item(),c.item(),loss]
print(" ")
k = k+2
j = j+1
data
data.to_csv("data_2_m_9_1.csv",index=False)
a = torch.tensor([0.],requires_grad=True)
b = torch.tensor([0.],requires_grad=True)
c = torch.tensor([0.],requires_grad=True)
criterion = BCELoss()
data = pd.DataFrame(columns=["sno","b_c_fixed","a_fixed","a_value","b_value","c_value","loss"])
Y1_ = Y1.float()
X1_average = focus_(a,X1)
Yhat1 = classification_(b,c,X1_average)
initial_loss = criterion(Yhat1,Y1_)
#print(initial_loss)
k = 0
data.loc[k] = [k,True,True,a.item(),b.item(),c.item(),initial_loss.item()]
k = k+1
j= 1
for i in range(0,40,2):
print("minimize a")
a,loss = minimize_a(X1,Y1,a,b,c,epochs=20000)
data.loc[k] = [j,True,False,a.item(),b.item(),c.item(),loss]
print("*"*60)
print(" ")
print("Minimize b and c")
b,c,loss = minimize_b_c(X1,Y1,a,b,c,epochs=20000)
#print(b,c)
data.loc[k+1] = [j,False,True,a.item(),b.item(),c.item(),loss]
print(" ")
k = k+2
j = j+1
data
data.to_csv("data_2_m_9_2.csv",index=False)
```
# m = 50
```
X2 = np.ones((50,50))*3
idx = np.arange(0,50,1)
X2[idx,idx] = -1
X3 = np.ones((50,50))*3
X3[idx,idx] = 1
X3 = np.concatenate((X2,X3),axis=0)
print(X3,X3.shape)
Y3 = np.zeros((100))
Y3[50:] = 1
print(Y3,Y3.shape)
def softmax_(x):
"""Compute softmax values for each sets of scores in x."""
m = torch.max(x,dim=1,keepdims=True)
print(m)
e_x = torch.exp(torch.sub(x,m.values))
return e_x / torch.sum(e_x,dim=1,keepdims=True)
a = torch.tensor([10.],requires_grad=True)
x = torch.tensor([[3.,-1.]])
out = torch.sum(softmax_(a*x) * x,dim=1)
print(out)
out.backward()
a.grad
l = ( torch.exp(a*x[0,0],) + torch.exp(a*x[0,1]) ) **2
#print(l)
f1 = ( ( ( x[0,0] - x[0,1] ) * torch.exp(a*x[0,0] + a*x[0,1]) ) / l ) * x[0,0]
f2 = ( ( ( x[0,1] - x[0,0] ) * torch.exp(a*x[0,0] + a*x[0,1]) ) / l ) * x[0,1]
print(f1.item()+f2.item())
x = np.array([[3,-1]])
a = 10
b = 1
c = 0
y = np.array([1])
xhat = np.sum(softmax(a*x) * x,axis=1)
print(xhat)
yhat = classification_(b,c,xhat)
derv_f(x,xhat,y,yhat,a,b)
```
| github_jupyter |
# Investigate temperature change potential of different aerosol forcing time series
### Figure sizing
AGU’s goal is for the text size of the labeling in each figure to match the text size in the body of the article. This means that for most AGU journals, text and labeling in a figure should be 8 points at the final printing size. Subscript and superscript should be 6 points.
All information within the figure, including symbols, legends, characters, patterns, and shading, must be legible at the final size. Figures should be sized between:
1/4 page figure = 95 mm x 115 mm
Full page = 190 mm x 230 mm
```
import numpy as np
import scipy.stats as st
import pandas as pd
import matplotlib.pyplot as pl
import os
import urllib
import json
import wquantiles
from matplotlib import rc
from matplotlib.ticker import AutoMinorLocator
from matplotlib.lines import Line2D
from scipy.stats import gaussian_kde
from scipy.optimize import root
from scipy.signal import savgol_filter
from scipy.interpolate import interp1d
from tqdm import tqdm_notebook
from scipy.signal import savgol_filter
from netCDF4 import Dataset
import matplotlib.gridspec as gridspec
import random
import h5py
from zipfile import ZipFile
from climateforcing.twolayermodel import TwoLayerModel
# hdf5 utilities
def save_dict_to_hdf5(dic, filename):
"""
....
"""
with h5py.File(filename, 'w') as h5file:
recursively_save_dict_contents_to_group(h5file, '/', dic)
def recursively_save_dict_contents_to_group(h5file, path, dic):
"""
....
"""
for key, item in dic.items():
if isinstance(item, (np.ndarray, np.int64, np.float64, str, bytes)):
h5file[path + key] = item
elif isinstance(item, dict):
recursively_save_dict_contents_to_group(h5file, path + key + '/', item)
else:
raise ValueError('Cannot save %s type'%type(item))
def load_dict_from_hdf5(filename):
"""
....
"""
with h5py.File(filename, 'r') as h5file:
return recursively_load_dict_contents_from_group(h5file, '/')
def recursively_load_dict_contents_from_group(h5file, path):
"""
....
"""
ans = {}
for key, item in h5file[path].items():
if isinstance(item, h5py._hl.dataset.Dataset):
ans[key] = item.value
elif isinstance(item, h5py._hl.group.Group):
ans[key] = recursively_load_dict_contents_from_group(h5file, path + key + '/')
return ans
# get my data
def check_and_download(filepath, url):
"""Checks prescence of a file and downloads if not present.
Inputs
------
filepath : str
filename to download to
url :
url to download from
"""
if not os.path.isfile(filepath):
urllib.request.urlretrieve(url, filepath)
return
pl.rcParams['figure.figsize'] = (12/2.54, 12/2.54)
pl.rcParams['font.size'] = 8
pl.rcParams['font.family'] = 'Arial'
pl.rcParams['xtick.direction'] = 'out'
pl.rcParams['xtick.minor.visible'] = True
pl.rcParams['ytick.minor.visible'] = True
pl.rcParams['ytick.right'] = True
pl.rcParams['xtick.top'] = True
pl.rcParams['figure.dpi'] = 96
```
## von Schuckmann ocean heat uptake
the citation is https://www.earth-syst-sci-data-discuss.net/essd-2019-255/
```
nc = Dataset('../data_input/GCOS_all_heat_content_1960-2018_ZJ_v22062020.nc')
#print(nc.variables)
ohctop = nc.variables['ohc_0-2000m'][:]
ohcbot = nc.variables['ohc_below_2000m'][:]
atmosh = nc.variables['atmospheric_heat_content'][:]
cryosh = nc.variables['energy_cryosphere'][:]
landhc = nc.variables['ground_heat_content'][:]
ohctopu = nc.variables['ohc_0-2000m_uncertainty'][:]
ohcbotu = nc.variables['ohc_below_2000m_uncertainty'][:]
atmoshu = nc.variables['atmospheric_heat_content_uncertainty'][:]
cryoshu = nc.variables['energy_cryosphere_uncertainty'][:]
landhcu = nc.variables['ground_heat_content_uncertainty'][:]
nc.close()
cryosh[-1] = cryosh[-2] # nan for 2017-2018, assume no change
cryoshu[-1] = cryoshu[-2]
OHCobs = (ohctop+ohcbot+atmosh+cryosh+landhc)-(ohctop+ohcbot+atmosh+cryosh+landhc)[11]
print(OHCobs[-1])
#OHCobs_u = np.sqrt(ohctopu**2 + ohcbotu**2 + atmoshu**2 + cryoshu**2 + landhcu**2)
#pl.fill_between(np.arange(1960.5, 2019), OHCobs - 2 * OHCobs_u, OHCobs + 2 * OHCobs_u, alpha=0.3)
#pl.plot(np.arange(1960.5,2019), OHCobs)
```
## Non-aerosol forcing is based on SSP2-4.5
- externally supplied and could be updated when AR6 approved
- this is an update of the RCMIP time series
```
ssp245_allforcing = pd.read_csv('../data_input/ERF_ssp245_1750-2500.csv')
baseline_forcing = ssp245_allforcing[:270].copy()
baseline_forcing.drop(
labels=['total_anthropogenic','total'],
axis='columns',
inplace=True
)
baseline_forcing['total_anthropogenic'] = baseline_forcing[['co2','ch4','n2o','other_wmghg','o3_tropospheric','o3_stratospheric','h2o_stratospheric','contrails','bc_on_snow','land_use','aerosol-radiation_interactions','aerosol-cloud_interactions']].sum(axis=1)
baseline_forcing['total'] = baseline_forcing['total_anthropogenic'] + baseline_forcing['total_natural']
baseline_forcing.set_index('year', inplace=True)
pd.set_option('display.max_rows', 999)
baseline_forcing
# Model-specific ERF datasets
CanESM5_aerforcing = pd.read_csv('../data_output/rfmip_aprp/CanESM5.csv')
E3SM_aerforcing = pd.read_csv('../data_output/rfmip_aprp/E3SM.csv')
GISS_aerforcing = pd.read_csv('../data_output/rfmip_aprp/GISS-E2-1-G.csv')
HadGEM3_aerforcing = pd.read_csv('../data_output/rfmip_aprp/HadGEM3-GC31-LL.csv')
MIROC6_aerforcing = pd.read_csv('../data_output/rfmip_aprp/MIROC6.csv')
GFDL_aerforcing = pd.read_csv('../data_output/rfmip_aprp/GFDL-CM4.csv')
GFDLESM_aerforcing = pd.read_csv('../data_output/rfmip_aprp/GFDL-ESM4.csv')
NorESM2_aerforcing = pd.read_csv('../data_output/rfmip_aprp/NorESM2-LM.csv')
UKESM_aerforcing = pd.read_csv('../data_output/rfmip_aprp/UKESM1-0-LL.csv')
IPSL_aerforcing = pd.read_csv('../data_output/rfmip_aprp/IPSL-CM6A-LR.csv')
MRI_aerforcing = pd.read_csv('../data_output/rfmip_aprp/MRI-ESM2-0.csv')
# Lund et al., 2019, ACP: (details in supplement)
# https://www.atmos-chem-phys.net/19/13827/2019/acp-19-13827-2019-supplement.pdf
# we use SSP2-4.5 for extension
Lund_rfacipoints = [0, -0.03, -0.08, -0.12, -0.13, -0.15, -0.17, -0.21, -0.26, -0.33, -0.38, -0.37,
-0.40, -0.44, -0.40, -0.45, -0.44, -0.44, -0.42, -0.36, -0.35, -0.26, -0.11]
Lund_rfaripoints = [0, 0.002, -0.03, -0.05, -0.06, -0.06, -0.08, -0.10, -0.14, -0.24, -0.26, -0.24,
-0.24, -0.22, -0.19, -0.20, -0.17, -0.17, -0.13, -0.10, -0.13, -0.09, -0.09]
Lund_timepoints = [1750, 1850, 1900, 1910, 1920, 1930, 1940, 1950, 1960, 1970, 1980, 1985, 1990, 1995,
2000, 2005, 2010, 2014, 2015, 2020, 2030, 2050, 2100]
f = interp1d(Lund_timepoints, np.array(Lund_rfacipoints)+np.array(Lund_rfaripoints))
Lund_aerforcing = f(np.arange(1750,2101))
f = interp1d(Lund_timepoints, np.array(Lund_rfaripoints))
Lund_ERFari = f(np.arange(1750,2101))
f = interp1d(Lund_timepoints, np.array(Lund_rfacipoints))
Lund_ERFaci = f(np.arange(1750,2101))
baseline_forcing.index
# Temperature (GMST) observations: Cowtan and Way, accessed 24 November 2020
cw_temp = np.loadtxt('../data_input/CW.txt')
# GSAT/GMST ratio. Use CMIP5 ratio, calculated by me for Rogelj et al. 2019, method originally from Richardson et al. 2016
blratio = np.loadtxt('../data_input/cmip5_data_2019.txt')[5,:]
cowtan = cw_temp[:,1] - np.mean(cw_temp[:51,1])
years = cw_temp[:,0]+0.5
blratio = np.concatenate((np.ones(11), blratio))
Tobs = blratio * cowtan
pl.plot(years, Tobs)
#pl.plot(np.arange(1750,1901), best_land)
print(np.mean(Tobs[:51]))
print(np.mean(Tobs[160:170]))
#print(blratio)
len(Tobs)
```
## Simple experiment with ECS=3.7 and default Geoffroy params
```
def rmse(obs, mod):
return np.sqrt(np.sum((obs-mod)**2)/len(obs))
models = ['CanESM5','E3SM','GFDL-CM4','GFDL-ESM4','GISS-E2-1-G','HadGEM3-GC31-LL','IPSL-CM6A-LR','MIROC6','MRI-ESM2-0','NorESM2-LM','UKESM1-0-LL']
colors = {
'ECLIPSE-constrained' : '0.6',
'CMIP6-constrained' : '0.3',
'CanESM5' : 'red',#'#1e4c24',
'E3SM' : 'darkorange',
'GFDL-ESM4' : 'yellowgreen',
'GFDL-CM4' : 'yellow',#'green',
'GISS-E2-1-G' : 'green',#'#771d7b',
'HadGEM3-GC31-LL': 'turquoise',
'IPSL-CM6A-LR' : 'teal',
'MIROC6' : 'blue',#b85fb7',
'MRI-ESM2-0' : 'blueviolet',
'NorESM2-LM' : 'purple',#'red',
'UKESM1-0-LL' : 'crimson',
'observations' : 'black',
'Oslo-CTM3' : 'pink',
}
ls = {
'CMIP6-constrained' : '-',
'ECLIPSE-constrained' : '-',
'CanESM5' : '-',
'E3SM' : '-',
'GFDL-ESM4' : '-',
'GFDL-CM4' : '-',
'GISS-E2-1-G' : '-',
'HadGEM3-GC31-LL': '-',
'IPSL-CM6A-LR' : '-',
'MIROC6' : '-',
'MRI-ESM2-0' : '-',
'NorESM2-LM' : '-',
'UKESM1-0-LL' : '-',
'Oslo-CTM3' : '-',
'observations' : '-',
}
# load in Geoffroy two layer model parameters (pre-calculated by Glen Harris)
params = pd.read_fwf('../data_input/scmpy2L_calib_n=44_eps=fit_v20200702.txt', sep=' ')
params.set_index('Model', inplace=True)
cmip6_models = list(params.index)
params.rename(columns={"F4x":'q4x', "Lambda":'lamg', "Cmix":'cmix', "Cdeep":'cdeep', "Gamma":'gamma_2l', "Epsilon":'eff'}, inplace=True)
#cmip6_models = list(params['gamma_2l']['model_data']['EBM-epsilon'].keys())
fig, ax = pl.subplots(2,3, figsize=(16/2.54, 12/2.54))
ax[0,0].hist(params['q4x'], bins=np.arange(5,11,0.5), density=True)
f_q4x = st.gaussian_kde(params['q4x'], bw_method='silverman')
ax[0,0].plot(np.linspace(5,11), f_q4x(np.linspace(5,11)), color='k', label='Kernel density')
ax[0,0].set_yticks([])
ax[0,0].set_xlabel('W m$^{-2}$')
ax[0,0].set_title(r'ERF from $F_{4}\times$CO$_2$ ($F_{4\times}$)', fontsize=8)
ax[0,0].set_xlim(5,11)
ax[0,1].hist(-params['lamg'], bins=np.arange(-2,0.2,0.2), density=True)
f_lamg = st.gaussian_kde(-params['lamg'], bw_method='silverman')
ax[0,1].plot(np.linspace(-2.4,0), f_lamg(np.linspace(-2.4,0)), color='k', label='Kernel density')
ax[0,1].set_yticks([])
ax[0,1].set_xlabel('W m$^{-2}$ K$^{-1}$')
ax[0,1].set_title(r'Climate feedback parameter ($\lambda$)', fontsize=8)
ax[0,1].set_xlim(-2.4,0)
ax[0,2].hist(params['eff'], bins=np.arange(0.4,2.2,0.2), density=True)
f_eff = st.gaussian_kde(params['eff'], bw_method='silverman')
ax[0,2].plot(np.linspace(0.4,2.2), f_eff(np.linspace(0.4,2.2)), color='k', label='Kernel density')
ax[0,2].set_yticks([])
ax[0,2].set_title(r'Efficacy of ocean heat uptake ($\epsilon$)', fontsize=8)
ax[0,2].set_xlim(0.4,2.2)
ax[1,0].hist(params['gamma_2l'], bins=np.arange(0.3,1.3,0.1), density=True)
f_gamma_2l = st.gaussian_kde(params['gamma_2l'], bw_method='silverman')
ax[1,0].plot(np.linspace(0.2,1.1), f_gamma_2l(np.linspace(0.2,1.1)), color='k', label='Kernel density')
ax[1,0].set_yticks([])
ax[1,0].set_xlabel('W m$^{-2}$ K$^{-1}$')
ax[1,0].set_title(r'Heat exchange ($\gamma$)', fontsize=8)
ax[1,0].set_xlim(0.2,1.1)
ax[1,1].hist(params['cmix'], bins=np.arange(5,11.5,0.5), density=True)
f_cmix = st.gaussian_kde(params['cmix'], bw_method='silverman')
ax[1,1].plot(np.linspace(5,11.5), f_cmix(np.linspace(5,11.5)), color='k', label='Kernel density')
ax[1,1].set_yticks([])
ax[1,1].set_xlabel('W yr m$^{-2}$ K$^{-1}$')
ax[1,1].set_title(r'Mixed-layer heat capacity (C)', fontsize=8)
ax[1,1].set_xlim(5,11.5)
ax[1,2].hist(params['cdeep'], bins=np.arange(0,420, 20), density=True, label='CMIP6 models')
f_cdeep = st.gaussian_kde(params['cdeep'], bw_method='silverman')
ax[1,2].plot(np.arange(0,400), f_cdeep(np.arange(0,400)), color='k', label='Kernel density')
ax[1,2].set_yticks([])
ax[1,2].set_xlabel('W yr m$^{-2}$ K$^{-1}$')
ax[1,2].set_title(r'Deep ocean heat capacity (C$_0$)', fontsize=8)
ax[1,2].set_xlim(0,400)
ax[1,2].legend()
pl.figtext(0.015,0.775,'Probability density', rotation=90, va='center', ha='center')
pl.figtext(0.015,0.275,'Probability density', rotation=90, va='center', ha='center')
fig.tight_layout(rect=[0.015,0,1,1])
pl.savefig('../figures/figureS4.png', dpi=300)
pl.savefig('../figures/figureS4.pdf')
# construct correlation matrix
pd.set_option('precision', 4)
params.corr()
pd.set_option('precision', 2)
geoff_ecs_data = np.zeros(((len(cmip6_models))))
geoff_ecs_data = params['q4x'].values/params['lamg'].values/2
geoff_df_display = params.copy()
geoff_df_display['ECS'] = geoff_ecs_data
geoff_df_display.sort_index()
samples = 100000
kde = st.gaussian_kde(params.T)
geoff_sample = kde.resample(size=int(samples*1.1), seed=3170812)
# remove unphysical combinations
geoff_sample[:,geoff_sample[0,:] <= 0] = np.nan
geoff_sample[:,geoff_sample[1,:] <= 0.2] = np.nan
geoff_sample[:,geoff_sample[2,:] <= 0] = np.nan
geoff_sample[:,geoff_sample[3,:] <= 0] = np.nan
geoff_sample[:,geoff_sample[4,:] <= 0] = np.nan
geoff_sample[:,geoff_sample[5,:] <= 0] = np.nan
#geoff_sample = geoff_sample[~np.isnan(geoff_sample)]
mask = np.all(np.isnan(geoff_sample), axis=0)
geoff_sample = geoff_sample[:,~mask]
geoff_sample_df=pd.DataFrame(
data=geoff_sample[:,:samples].T, columns=['q4x','lamg','cmix','cdeep','gamma_2l','eff']
)
geoff_sample_df.to_csv('../data_output/geoff_sample.csv', index=False)
geoff_sample_df
# fractional uncertainties on ERF - based on the FAIR code
seed = 36572
zscore = st.norm.ppf(0.95)
# can only use published literature - so revert to FaIR and AR5 uncertainties. Use unmodified Etminan for methane, because RFMIP
# models central estimate is quite close.
unc_ranges = np.array([
0.20, # CO2
0.28, # CH4: updated value from etminan 2016
0.20, # N2O
0.20, # other WMGHGS
0.50, # tropospheric O3
2.00, # stratospheric O3
1.00, # stratospheric WV from CH4
(98-57.4)/57.4, # contrails
0.00, # black carbon on snow (lognormal)
0.75, # land use change
0.50, # volcanic
0.50, # solar (amplitude)
])/(zscore)
scale = st.norm.rvs(size=(samples,12), loc=np.ones((samples,12)), scale=np.ones((samples, 12)) * unc_ranges[None,:], random_state=seed)
scale[:,8] = st.lognorm.rvs(0.5, size=samples, random_state=seed+1)
# contrails slightly asymmetric
scale[scale[:,7]<1,7] = (57.4-19)/(98-57.4)*(scale[scale[:,7]<1,7]-1) + 1
scale_df = pd.DataFrame(
data = scale,
columns = ['co2','ch4','n2o','other_wmghg','o3_tropospheric','o3_stratospheric','h2o_stratospheric','contrails','bc_on_snow','land_use','volcanic','solar']
)
scale_df
pl.hist(scale[:,7])
trend_solar = st.norm.rvs(size=samples, loc=0, scale=0.1/zscore, random_state=138294)
```
## Construct CMIP6 emissions-based forcing
```
# Grab CEDS emissions and unzip (RCMIP should already be here)
check_and_download('../data_input/CEDS_v_2020_09_11_emissions.zip', 'https://zenodo.org/record/4025316/files/CEDS_v_2020_09_11_emissions.zip')
with ZipFile('../data_input/CEDS_v_2020_09_11_emissions.zip', 'r') as zipObj:
zipObj.extractall('../data_input/ceds')
os.remove('../data_input/CEDS_v_2020_09_11_emissions.zip')
emissions = pd.read_csv('../data_input/rcmip/rcmip-emissions-annual-means-v5-1-0.csv')
df_emissions = pd.concat([emissions.loc[(
(emissions.Variable=='Emissions|BC')|
(emissions.Variable=='Emissions|OC')|
(emissions.Variable=='Emissions|Sulfur')|
(emissions.Variable=='Emissions|NOx')|
(emissions.Variable=='Emissions|NH3')|
(emissions.Variable=='Emissions|VOC')|
(emissions.Variable=='Emissions|CO')
) & (emissions.Scenario=='ssp245') & (emissions.Region=='World'), 'Variable'], emissions.loc[(
(emissions.Variable=='Emissions|BC')|
(emissions.Variable=='Emissions|OC')|
(emissions.Variable=='Emissions|Sulfur')|
(emissions.Variable=='Emissions|NOx')|
(emissions.Variable=='Emissions|NH3')|
(emissions.Variable=='Emissions|VOC')|
(emissions.Variable=='Emissions|CO')
) & (emissions.Scenario=='ssp245') & (emissions.Region=='World'), '1750':'2100']], axis=1)#.interpolate(axis=1).T
df_emissions.set_index('Variable', inplace=True)
df_emissions = df_emissions.interpolate(axis=1).T
df_emissions.rename(
columns={
'Emissions|BC': 'BC',
'Emissions|OC': 'OC',
'Emissions|Sulfur': 'SO2',
'Emissions|NOx': 'NOx',
'Emissions|NH3': 'NH3',
'Emissions|VOC': 'VOC',
'Emissions|CO': 'CO'
}, inplace=True
)
# only keep cols we want
emissions = df_emissions[['SO2', 'BC', 'OC', 'NH3', 'NOx', 'VOC', 'CO']]
emissions.index = emissions.index.astype('int')
emissions.index.name='year'
emissions.columns.name=None
emissions_ceds_update = emissions.copy()
emissions_old = pd.read_csv('../data_input/rcmip/rcmip-emissions-annual-means-v5-1-0.csv')
df_emissions = pd.concat([emissions_old.loc[(
(emissions_old.Variable=='Emissions|BC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|OC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|Sulfur|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|NOx|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|NH3|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|VOC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|CO|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|BC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|OC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|Sulfur|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|NOx|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|NH3|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|VOC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|CO|MAGICC AFOLU|Agriculture')
) & (emissions_old.Scenario=='ssp245') & (emissions_old.Region=='World'), 'Variable'], emissions_old.loc[(
(emissions_old.Variable=='Emissions|BC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|OC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|Sulfur|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|NOx|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|NH3|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|VOC|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|CO|MAGICC Fossil and Industrial')|
(emissions_old.Variable=='Emissions|BC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|OC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|Sulfur|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|NOx|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|NH3|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|VOC|MAGICC AFOLU|Agriculture')|
(emissions_old.Variable=='Emissions|CO|MAGICC AFOLU|Agriculture')
) & (emissions_old.Scenario=='ssp245') & (emissions_old.Region=='World'), '1750':'2100']], axis=1)#.interpolate(axis=1).T
df_emissions.set_index('Variable', inplace=True)
df_emissions = df_emissions.interpolate(axis=1).T
for species in ['BC', 'OC', 'Sulfur', 'NOx', 'NH3', 'VOC', 'CO']:
df_emissions[species] = df_emissions['Emissions|{}|MAGICC Fossil and Industrial'.format(species)] + df_emissions['Emissions|{}|MAGICC AFOLU|Agriculture'.format(species)]
df_emissions.rename(columns = {'Sulfur': 'SO2'}, inplace=True)
df_emissions.drop(columns=[
'Emissions|BC|MAGICC Fossil and Industrial',
'Emissions|OC|MAGICC Fossil and Industrial',
'Emissions|Sulfur|MAGICC Fossil and Industrial',
'Emissions|NOx|MAGICC Fossil and Industrial',
'Emissions|NH3|MAGICC Fossil and Industrial',
'Emissions|VOC|MAGICC Fossil and Industrial',
'Emissions|CO|MAGICC Fossil and Industrial',
'Emissions|BC|MAGICC AFOLU|Agriculture',
'Emissions|OC|MAGICC AFOLU|Agriculture',
'Emissions|Sulfur|MAGICC AFOLU|Agriculture',
'Emissions|NOx|MAGICC AFOLU|Agriculture',
'Emissions|NH3|MAGICC AFOLU|Agriculture',
'Emissions|VOC|MAGICC AFOLU|Agriculture',
'Emissions|CO|MAGICC AFOLU|Agriculture',
],
inplace=True
)
df_emissions.index = emissions.index.astype('int')
df_emissions.index.name='year'
df_emissions.columns.name=None
global_total = {}
for species in ['BC', 'OC', 'SO2', 'NH3', 'NOx', 'NMVOC', 'CO']:
df = pd.read_csv('../data_input/ceds/{}_global_CEDS_emissions_by_sector_2020_09_11.csv'.format(species))
global_total[species] = df.sum(axis=0).values[3:].astype(float) / 1000 # yes could get openscm on this
#unit = df.units[0]
#print(unit)
global_total['VOC'] = global_total.pop('NMVOC')
new_ceds = pd.DataFrame(global_total)
new_ceds.index = np.arange(1750,2020)
new_ceds.index = new_ceds.index.astype('int')
new_ceds.index.name='year'
new_ceds.columns.name=None
emissions_ceds_update = new_ceds.loc[1750:2020] + emissions - df_emissions
emissions_ceds_update.drop(index=range(2020,2101), inplace=True)
emissions_ceds_update
#emissions = pd.read_csv('../output_data/historical_slcf_emissions.csv', index_col='year')
emissions = emissions_ceds_update.drop(['CO','VOC','NOx','NH3'], axis=1)
emissions
emissions_old_df = pd.read_csv('../data_input/rcmip/rcmip-emissions-annual-means-v5-1-0.csv')
emissions_old = emissions_old_df.loc[(
(emissions_old_df.Variable=='Emissions|BC')|
(emissions_old_df.Variable=='Emissions|OC')|
(emissions_old_df.Variable=='Emissions|Sulfur')
) & (emissions_old_df.Scenario=='ssp245') & (emissions_old_df.Region=='World'), '1750':'2020'].interpolate(axis=1).T
emissions_old.columns = ['BC','OC','SO2']
fig = pl.figure(figsize=(19/2.54, 9.5/2.54))
pl.plot(emissions.loc[1750:,'SO2'], label='CEDS updated SO$_2$ (Mt SO$_2$)', color='blue')
pl.plot(emissions.loc[1750:,'BC']*10, label='CEDS updated BCx10 (Mt C)', color='black')
pl.plot(emissions.loc[1750:,'OC'], label='CEDS updated OC (Mt C)', color='brown')
pl.plot(np.arange(1750,2020), emissions_old.loc['1750':'2019','SO2'].values, ls='--', color='blue', label='CMIP6 SO$_2$ (Mt SO$_2$)')
pl.plot(np.arange(1750,2020), emissions_old.loc['1750':'2019','BC'].values*10, ls='--', color='black', label='CMIP6 BCx10 (Mt C)')
pl.plot(np.arange(1750,2020), emissions_old.loc['1750':'2019','OC'].values, ls='--', color='brown', label='CMIP6 OC (Mt C)')
pl.xlim(1750,2019)
pl.ylim(0,140)
pl.legend()
pl.ylabel('Mt/yr')
pl.title("Aerosol emissions from CMIP6 and updated CEDS (O'Rourke et al. 2020)")
pl.tight_layout()
pl.savefig('../figures/figureS2.png', dpi=300)
pl.savefig('../figures/figureS2.pdf')
def aerocom(x, bc, oc, so2):
return bc*x[0] + oc*x[1] + so2*x[2]
def ghan(x, beta, n0, n1):
return -beta*np.log(1 + x[0]/n0 + x[1]/n1)
df = pd.read_csv('../data_output/ERFari_samples.csv')
ari_coeffs = df.values
df = pd.read_csv('../data_output/ERFaci_samples.csv')
aci_coeffs = np.exp(df.values)
# Use Ringberg aerosol priors, from script provided to me
def uniform1684(a,b,seed,samples=2000):
interval = (b-a)+((b-a)/(84-16)*32)
lower = a-((b-a)/(84-16)*16)
return st.uniform.rvs(lower, interval, size=samples, random_state=seed)
dtau = uniform1684(0.02,0.04,123,samples=samples)
tau = uniform1684(0.13,0.17,124,samples=samples)
S_tau = uniform1684(-27,-20,125,samples=samples)
RFari_cloudy = uniform1684(-0.1,0.1,126,samples=samples)
dR_dRatm = uniform1684(-0.3,-0.1,127,samples=samples)
dRatm_dtau = uniform1684(17,35,128,samples=samples)
c_tau = uniform1684(0.59,0.71,129,samples=samples)
c_N = uniform1684(0.19,0.29,130,samples=samples)
c_L = uniform1684(0.21,0.29,131,samples=samples)
c_C = uniform1684(0.59,1.07,132,samples=samples)
beta_N_tau = uniform1684(0.3,0.8,133,samples=samples)
beta_L_N = uniform1684(-0.36,-0.011,134,samples=samples)
beta_C_N = uniform1684(0,0.1,135,samples=samples)
S_N = uniform1684(-27,-26,136,samples=samples)
S_L = uniform1684(-56,-54,137,samples=samples)
S_C = uniform1684(-153,-91,138,samples=samples)
rfari = dtau*S_tau*(1-c_tau)+RFari_cloudy
rfari_adj = dtau*dR_dRatm*dRatm_dtau
dlntau = dtau/tau
deltan = dlntau * beta_N_tau
rfaci = dlntau*beta_N_tau*S_N*c_N
erfaci_L = dlntau*beta_N_tau*beta_L_N*S_L*c_L
erfaci_C = dlntau*beta_N_tau*beta_C_N*S_C*c_C
ERFari_scale = rfari + rfari_adj
ERFaci_scale = rfaci + erfaci_L + erfaci_C
fig,ax = pl.subplots(1,2, figsize=(18/2.54,12/2.54))
ax[0].hist(ERFari_scale, bins=np.arange(-1.2,0.05,0.05), density=True);
ax[0].set_title('ERFari prior');
ax[1].hist(ERFaci_scale, bins=np.arange(-4,0.5,0.2), density=True);
ax[1].set_title('ERFaci prior');
# Define our dicts
ERFari = {}
ERFaci = {}
temp = {}
ks = {}
ohc = {}
hflux = {}
ks['temp'] = {}
ks['ohc'] = {}
ks['multi'] = {}
# # load dicts
# ERFari = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/ERFari.h5')
# ERFaci = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/ERFaci.h5')
# temp = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/temp.h5')
# ks = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/knutti_score.h5')
# ohc = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/ohc.h5')
# hflux = load_dict_from_hdf5('/nfs/a65/pmcjs/AR6_tuning/aerosols/hflux.h5')
intvar = np.loadtxt('../data_output/piControl/internal_variability_piControl.txt')
def knutti_score(obs, mod, sigma_D=None):
"""
obs: observations data: array of size (nyears,)
mod: model data: array of size (nyears, nsamples)
"""
samples = mod.shape[1]
rm_d = np.ones(samples) * np.nan
for i in range(samples):
rm_d[i] = rmse(obs, mod[:, i])
if sigma_D==None:
sigma_D = np.nanmin(rm_d)
veracity = np.exp(-rm_d**2/sigma_D**2)
ks_raw = veracity
ks_raw[np.isnan(ks_raw)] = 0
ks = ks_raw/np.sum(ks_raw)
return ks
def weighted_percentile(a, w, q):
if isinstance(q, (list, tuple, np.ndarray)):
result = []
for iq in q:
result.append(wquantiles.quantile(a, w, iq))
else:
result = wquantiles.quantile(a, w, q)
return result
def simple_weight(obs, mod, sigma_D):
veracity = np.exp(-(mod-obs)**2/sigma_D**2)
similarity = 1 # I see no good reason to change this #
ks_raw = veracity/similarity
ks_raw[np.isnan(ks_raw)] = 0
ks = ks_raw/np.sum(ks_raw)
return ks
```
## CEDS emissions
```
ERFari['CMIP6-constrained'] = np.zeros((270,samples))
for i in tqdm_notebook(range(samples)):
ts2010 = np.mean(aerocom([emissions.loc[2005:2015,'BC'], emissions.loc[2005:2015,'OC'], emissions.loc[2005:2015,'SO2']], ari_coeffs[i,1], ari_coeffs[i,2], ari_coeffs[i,0]))
ts1850 = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], ari_coeffs[i,1], ari_coeffs[i,2], ari_coeffs[i,0])
ts1750 = aerocom([emissions.loc[1750,'BC'], emissions.loc[1750,'OC'], emissions.loc[1750,'SO2']], ari_coeffs[i,1], ari_coeffs[i,2], ari_coeffs[i,0])
ERFari['CMIP6-constrained'][:,i] = (
aerocom([emissions['BC'], emissions['OC'], emissions['SO2']], ari_coeffs[i,1], ari_coeffs[i,2], ari_coeffs[i,0])
- ts1750
)/(ts2010-ts1850)*(ERFari_scale[i])
ERFaci['CMIP6-constrained'] = np.zeros((270,samples))
for i in tqdm_notebook(range(samples)):
ts2010 = np.mean(ghan([emissions.loc[2005:2015,'SO2'], emissions.loc[2005:2015,'BC']+emissions.loc[2005:2015,'OC']], 0.97, aci_coeffs[i,0], aci_coeffs[i,1]))
ts1850 = ghan([emissions.loc[1850,'SO2'],emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], 0.97, aci_coeffs[i,0], aci_coeffs[i,1])
ts1750 = ghan([emissions.loc[1750,'SO2'],emissions.loc[1750,'BC']+emissions.loc[1750,'OC']], 0.97, aci_coeffs[i,0], aci_coeffs[i,1])
ERFaci['CMIP6-constrained'][:,i] = (
ghan([emissions['SO2'],emissions['BC']+emissions['OC']], 0.97, aci_coeffs[i,0], aci_coeffs[i,1])
- ts1750
)/(ts2010-ts1850)*(ERFaci_scale[i])
set1 = np.squeeze(np.where(np.logical_and(
1.001*(np.percentile((ERFari['CMIP6-constrained'][-1,:] + ERFaci['CMIP6-constrained'][-1,:]), 50))
< (ERFari['CMIP6-constrained'][-1,:]+ERFaci['CMIP6-constrained'][-1,:]), (ERFari['CMIP6-constrained'][-1,:]+ERFaci['CMIP6-constrained'][-1,:]) <
0.999*(np.percentile((ERFari['CMIP6-constrained'][-1,:] + ERFaci['CMIP6-constrained'][-1,:]), 50)))))
set1
fig, ax = pl.subplots(1,3,figsize=(19/2.54, 9.5/2.54))
ax[0].fill_between(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained'], 5, axis=1), np.percentile(ERFari['CMIP6-constrained'], 95, axis=1), color='0.75', lw=0);
ax[0].fill_between(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained'], 16, axis=1), np.percentile(ERFari['CMIP6-constrained'], 84, axis=1), color='0.5', lw=0);
ax[0].plot(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained'], 50, axis=1), color='k', zorder=10)
ax[0].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,754], color='cyan', label='Parameter set #754')
ax[0].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,1076], color='magenta', label='Parameter set #1076')
ax[0].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,18010], color='lime', label='Parameter set #18010')
ax[0].legend()
ax[0].set_xlim(1800,2020)
ax[0].set_title('ERFari')
ax[0].set_ylabel('W m$^{-2}$')
ax[1].fill_between(np.arange(1750,2020), np.percentile(ERFaci['CMIP6-constrained'], 5, axis=1), np.percentile(ERFaci['CMIP6-constrained'], 95, axis=1), color='0.75', lw=0, label='5-95% range');
ax[1].fill_between(np.arange(1750,2020), np.percentile(ERFaci['CMIP6-constrained'], 16, axis=1), np.percentile(ERFaci['CMIP6-constrained'], 84, axis=1), color='0.5', lw=0, label='16-84% range');
ax[1].plot(np.arange(1750,2020), np.percentile(ERFaci['CMIP6-constrained'], 50, axis=1), color='k', label='median', zorder=10)
ax[1].plot(np.arange(1750,2020), ERFaci['CMIP6-constrained'][:,754], color='cyan')
ax[1].plot(np.arange(1750,2020), ERFaci['CMIP6-constrained'][:,1076], color='magenta')
ax[1].plot(np.arange(1750,2020), ERFaci['CMIP6-constrained'][:,18010], color='lime')
ax[1].legend()
ax[1].set_xlim(1800,2020)
ax[1].set_title('ERFaci')
ax[2].fill_between(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained']+ERFaci['CMIP6-constrained'], 5, axis=1), np.percentile(ERFari['CMIP6-constrained']+ERFaci['CMIP6-constrained'], 95, axis=1), color='0.75', lw=0);
ax[2].fill_between(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained']+ERFaci['CMIP6-constrained'], 16, axis=1), np.percentile(ERFari['CMIP6-constrained']+ERFaci['CMIP6-constrained'], 84, axis=1), color='0.5', lw=0);
ax[2].plot(np.arange(1750,2020), np.percentile(ERFari['CMIP6-constrained']+ERFaci['CMIP6-constrained'], 50, axis=1), color='k', zorder=10)
ax[2].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,754]+ERFaci['CMIP6-constrained'][:,754], color='cyan', label='Ensemble 754')
ax[2].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,1076]+ERFaci['CMIP6-constrained'][:,1076], color='magenta', label='Ensemble 1076')
ax[2].plot(np.arange(1750,2020), ERFari['CMIP6-constrained'][:,18010]+ERFaci['CMIP6-constrained'][:,18010], color='lime', label='Ensemble 18010')
ax[2].set_xlim(1800,2020)
ax[2].set_title('Aerosol ERF')
ax[0].set_ylim(-3,0.2)
ax[1].set_ylim(-3,0.2)
ax[2].set_ylim(-3,0.2)
ax[0].axhline(0, ls=':', lw=0.5, color='k')
ax[1].axhline(0, ls=':', lw=0.5, color='k')
ax[2].axhline(0, ls=':', lw=0.5, color='k')
fig.tight_layout()
pl.savefig('../figures/figure5.png', dpi=300)
pl.savefig('../figures/figure5.pdf')
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[0,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[0], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['CMIP6-constrained'][:270,0]
in_forcing['aerosol-cloud_interactions'] = ERFaci['CMIP6-constrained'][:270,0]
in_forcing['total'] = in_forcing.sum(axis=1)
in_forcing['total']
temp['CMIP6-constrained'] = np.zeros((270, samples))
ohc['CMIP6-constrained'] = np.zeros((270, samples))
hflux['CMIP6-constrained'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['CMIP6-constrained'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['CMIP6-constrained'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['CMIP6-constrained'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['CMIP6-constrained'][:,i] = out.ohc
hflux['CMIP6-constrained'][:,i] = out.hflux
pl.fill_between(np.arange(1750,2020), np.percentile(temp['CMIP6-constrained'], 5, axis=1), np.percentile(temp['CMIP6-constrained'], 95, axis=1))
pl.plot(np.arange(1750,2020), np.median(temp['CMIP6-constrained'], axis=1), color='k')
ks['temp']['CMIP6-constrained'] = knutti_score(Tobs, temp['CMIP6-constrained'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['CMIP6-constrained'] = simple_weight(358, 10*(ohc['CMIP6-constrained'][268,:]-ohc['CMIP6-constrained'][221,:]), sigma_D=37)
ks['multi']['CMIP6-constrained'] = (ks['temp']['CMIP6-constrained']*ks['ohc']['CMIP6-constrained'])/(np.sum(ks['temp']['CMIP6-constrained']*ks['ohc']['CMIP6-constrained']))
print(weighted_percentile(ERFari['CMIP6-constrained'][269,:]+ERFaci['CMIP6-constrained'][269,:], ks['temp']['CMIP6-constrained'][:], [.05,.16,.5,.84,.95]))
print(weighted_percentile(ERFari['CMIP6-constrained'][269,:]+ERFaci['CMIP6-constrained'][269,:], ks['ohc']['CMIP6-constrained'][:], [.05,.16,.5,.84,.95]))
print(weighted_percentile(ERFari['CMIP6-constrained'][269,:]+ERFaci['CMIP6-constrained'][269,:], ks['multi']['CMIP6-constrained'][:], [.05,.16,.5,.84,.95]))
```
## Oslo-CTM3
```
Lund2010ari = Lund_ERFari[255:265].mean()
Lund2010aci = Lund_ERFaci[255:265].mean()
Lund1850ari = Lund_ERFari[100]
Lund1850aci = Lund_ERFaci[100]
ERFari['Oslo-CTM3'] = np.zeros((270,samples))
ERFaci['Oslo-CTM3'] = np.zeros((270,samples))
for i in range(samples):
ERFaci['Oslo-CTM3'][:,i] = Lund_ERFaci[:270]/(Lund2010aci-Lund1850aci)*(ERFaci_scale[i])
ERFari['Oslo-CTM3'][:,i] = Lund_ERFari[:270]/(Lund2010ari-Lund1850aci)*(ERFari_scale[i])
temp['Oslo-CTM3'] = np.zeros((270, samples))
ohc['Oslo-CTM3'] = np.zeros((270, samples))
hflux['Oslo-CTM3'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['Oslo-CTM3'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['Oslo-CTM3'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['Oslo-CTM3'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['Oslo-CTM3'][:,i] = out.ohc
hflux['Oslo-CTM3'][:,i] = out.hflux
ks['temp']['Oslo-CTM3'] = knutti_score(Tobs, temp['Oslo-CTM3'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['Oslo-CTM3'] = simple_weight(358, 10*(ohc['Oslo-CTM3'][268,:]-ohc['Oslo-CTM3'][221,:]), sigma_D=37)
ks['multi']['Oslo-CTM3'] = (ks['temp']['Oslo-CTM3']*ks['ohc']['Oslo-CTM3'])/(np.sum(ks['temp']['Oslo-CTM3']*ks['ohc']['Oslo-CTM3']))
with open("../data_output/cmip6_aerosol_coefficients.json", "r") as read_file:
aero_coeffs = json.load(read_file)
aero_coeffs
```
## GFDL-ESM4
```
GFDLESM2010ari = GFDLESM_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+GFDLESM_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
GFDLESM2010aci = GFDLESM_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+GFDLESM_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['GFDL-ESM4'] = np.zeros((270,samples))
ERFaci['GFDL-ESM4'] = np.zeros((270,samples))
GFDLESMari = np.zeros(270)
GFDLESMaci = np.zeros(270)
GFDLESMari[100:265] = GFDLESM_aerforcing.loc[0:165, 'aprp_ERFariSW'].values+GFDLESM_aerforcing.loc[0:165, 'aprp_ERFariLW'].values
GFDLESMaci[100:265] = GFDLESM_aerforcing.loc[0:165, 'aprp_ERFaciSW'].values+GFDLESM_aerforcing.loc[0:165, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['GFDL-ESM4']['ERFari']['BC'], aero_coeffs['GFDL-ESM4']['ERFari']['OC'], aero_coeffs['GFDL-ESM4']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['GFDL-ESM4']['ERFaci']['beta'], aero_coeffs['GFDL-ESM4']['ERFaci']['n0'], aero_coeffs['GFDL-ESM4']['ERFaci']['n1'])
GFDLESMari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['GFDL-ESM4']['ERFari']['BC'], aero_coeffs['GFDL-ESM4']['ERFari']['OC'], aero_coeffs['GFDL-ESM4']['ERFari']['SO2']) - ts1850ari
GFDLESMaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['GFDL-ESM4']['ERFaci']['beta'], aero_coeffs['GFDL-ESM4']['ERFaci']['n0'], aero_coeffs['GFDL-ESM4']['ERFaci']['n1']) - ts1850aci
GFDLESMari[265:] = aerocom([emissions.loc[2015:2019,'BC'], emissions.loc[2015:2019,'OC'], emissions.loc[2015:2019,'SO2']], aero_coeffs['GFDL-ESM4']['ERFari']['BC'], aero_coeffs['GFDL-ESM4']['ERFari']['OC'], aero_coeffs['GFDL-ESM4']['ERFari']['SO2']) - ts1850ari
GFDLESMaci[265:] = ghan([emissions.loc[2015:2019,'SO2'], emissions.loc[2015:2019,'BC']+emissions.loc[2015:2019,'OC']], aero_coeffs['GFDL-ESM4']['ERFaci']['beta'], aero_coeffs['GFDL-ESM4']['ERFaci']['n0'], aero_coeffs['GFDL-ESM4']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['GFDL-ESM4'][:,i] = (GFDLESMaci-GFDLESMaci[0])/GFDLESM2010aci*(ERFaci_scale[i])
ERFari['GFDL-ESM4'][:,i] = (GFDLESMari-GFDLESMari[0])/GFDLESM2010ari*(ERFari_scale[i])
temp['GFDL-ESM4'] = np.zeros((270, samples))
ohc['GFDL-ESM4'] = np.zeros((270, samples))
hflux['GFDL-ESM4'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['GFDL-ESM4'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['GFDL-ESM4'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['GFDL-ESM4'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['GFDL-ESM4'][:,i] = out.ohc
hflux['GFDL-ESM4'][:,i] = out.hflux
ks['temp']['GFDL-ESM4'] = knutti_score(Tobs, temp['GFDL-ESM4'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['GFDL-ESM4'] = simple_weight(358, 10*(ohc['GFDL-ESM4'][268,:]-ohc['GFDL-ESM4'][221,:]), sigma_D=37)
ks['multi']['GFDL-ESM4'] = (ks['temp']['GFDL-ESM4']*ks['ohc']['GFDL-ESM4'])/(np.sum(ks['temp']['GFDL-ESM4']*ks['ohc']['GFDL-ESM4']))
```
## UKESM1-0-LL
```
UKESM2010ari = UKESM_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+UKESM_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
UKESM2010aci = UKESM_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+UKESM_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['UKESM1-0-LL'] = np.zeros((270,samples))
ERFaci['UKESM1-0-LL'] = np.zeros((270,samples))
UKESMari = np.zeros(270)
UKESMaci = np.zeros(270)
UKESMari[100:265] = UKESM_aerforcing.loc[0:165, 'aprp_ERFariSW'].values+UKESM_aerforcing.loc[0:165, 'aprp_ERFariLW'].values
UKESMaci[100:265] = UKESM_aerforcing.loc[0:165, 'aprp_ERFaciSW'].values+UKESM_aerforcing.loc[0:165, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['UKESM1-0-LL']['ERFari']['BC'], aero_coeffs['UKESM1-0-LL']['ERFari']['OC'], aero_coeffs['UKESM1-0-LL']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['UKESM1-0-LL']['ERFaci']['beta'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n0'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n1'])
UKESMari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['UKESM1-0-LL']['ERFari']['BC'], aero_coeffs['UKESM1-0-LL']['ERFari']['OC'], aero_coeffs['UKESM1-0-LL']['ERFari']['SO2']) - ts1850ari
UKESMaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['UKESM1-0-LL']['ERFaci']['beta'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n0'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n1']) - ts1850aci
UKESMari[265:] = aerocom([emissions.loc[2015:2019,'BC'], emissions.loc[2015:2019,'OC'], emissions.loc[2015:2019,'SO2']], aero_coeffs['UKESM1-0-LL']['ERFari']['BC'], aero_coeffs['UKESM1-0-LL']['ERFari']['OC'], aero_coeffs['UKESM1-0-LL']['ERFari']['SO2']) - ts1850ari
UKESMaci[265:] = ghan([emissions.loc[2015:2019,'SO2'], emissions.loc[2015:2019,'BC']+emissions.loc[2015:2019,'OC']], aero_coeffs['UKESM1-0-LL']['ERFaci']['beta'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n0'], aero_coeffs['UKESM1-0-LL']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['UKESM1-0-LL'][:,i] = (UKESMaci-UKESMaci[0])/UKESM2010aci*(ERFaci_scale[i])
ERFari['UKESM1-0-LL'][:,i] = (UKESMari-UKESMari[0])/UKESM2010ari*(ERFari_scale[i])
temp['UKESM1-0-LL'] = np.zeros((270, samples))
ohc['UKESM1-0-LL'] = np.zeros((270, samples))
hflux['UKESM1-0-LL'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['UKESM1-0-LL'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['UKESM1-0-LL'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['UKESM1-0-LL'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['UKESM1-0-LL'][:,i] = out.ohc
hflux['UKESM1-0-LL'][:,i] = out.hflux
ks['temp']['UKESM1-0-LL'] = knutti_score(Tobs, temp['UKESM1-0-LL'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['UKESM1-0-LL'] = simple_weight(358, 10*(ohc['UKESM1-0-LL'][268,:]-ohc['UKESM1-0-LL'][221,:]), sigma_D=37)
ks['multi']['UKESM1-0-LL'] = (ks['temp']['UKESM1-0-LL']*ks['ohc']['UKESM1-0-LL'])/(np.sum(ks['temp']['UKESM1-0-LL']*ks['ohc']['UKESM1-0-LL']))
```
## CanESM5
```
CanESM2010ari = CanESM5_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+CanESM5_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
CanESM2010aci = CanESM5_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+CanESM5_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['CanESM5'] = np.zeros((270,samples))
ERFaci['CanESM5'] = np.zeros((270,samples))
CanESMari = np.zeros(270)
CanESMaci = np.zeros(270)
CanESMari[100:] = CanESM5_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+CanESM5_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
CanESMaci[100:] = CanESM5_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+CanESM5_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['CanESM5']['ERFari']['BC'], aero_coeffs['CanESM5']['ERFari']['OC'], aero_coeffs['CanESM5']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['CanESM5']['ERFaci']['beta'], aero_coeffs['CanESM5']['ERFaci']['n0'], aero_coeffs['CanESM5']['ERFaci']['n1'])
CanESMari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['CanESM5']['ERFari']['BC'], aero_coeffs['CanESM5']['ERFari']['OC'], aero_coeffs['CanESM5']['ERFari']['SO2']) - ts1850ari
CanESMaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['CanESM5']['ERFaci']['beta'], aero_coeffs['CanESM5']['ERFaci']['n0'], aero_coeffs['CanESM5']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['CanESM5'][:,i] = (CanESMaci-CanESMaci[0])/CanESM2010aci*(ERFaci_scale[i])
ERFari['CanESM5'][:,i] = (CanESMari-CanESMari[0])/CanESM2010ari*(ERFari_scale[i])
temp['CanESM5'] = np.zeros((270, samples))
ohc['CanESM5'] = np.zeros((270, samples))
hflux['CanESM5'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['CanESM5'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['CanESM5'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['CanESM5'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['CanESM5'][:,i] = out.ohc
hflux['CanESM5'][:,i] = out.hflux
ks['temp']['CanESM5'] = knutti_score(Tobs, temp['CanESM5'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['CanESM5'] = simple_weight(358, 10*(ohc['CanESM5'][268,:]-ohc['CanESM5'][221,:]), sigma_D=37)
ks['multi']['CanESM5'] = (ks['temp']['CanESM5']*ks['ohc']['CanESM5'])/(np.sum(ks['temp']['CanESM5']*ks['ohc']['CanESM5']))
```
## E3SM
```
# take 05-14 mean for E3SM
E3SM2010ari = E3SM_aerforcing.loc[135:144,'aprp_ERFariSW'].mean()+E3SM_aerforcing.loc[135:144,'aprp_ERFariLW'].mean()
E3SM2010aci = E3SM_aerforcing.loc[135:144,'aprp_ERFaciSW'].mean()+E3SM_aerforcing.loc[135:144,'aprp_ERFaciLW'].mean()
ERFari['E3SM'] = np.zeros((270,samples))
ERFaci['E3SM'] = np.zeros((270,samples))
E3SMari = np.zeros(270)
E3SMaci = np.zeros(270)
E3SMari[120:265] = E3SM_aerforcing.loc[0:145, 'aprp_ERFariSW'].values+E3SM_aerforcing.loc[0:145, 'aprp_ERFariLW'].values
E3SMaci[120:265] = E3SM_aerforcing.loc[0:145, 'aprp_ERFaciSW'].values+E3SM_aerforcing.loc[0:145, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['E3SM']['ERFari']['BC'], aero_coeffs['E3SM']['ERFari']['OC'], aero_coeffs['E3SM']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['E3SM']['ERFaci']['beta'], aero_coeffs['E3SM']['ERFaci']['n0'], aero_coeffs['E3SM']['ERFaci']['n1'])
E3SMari[:120] = aerocom([emissions.loc[1750:1869,'BC'], emissions.loc[1750:1869,'OC'], emissions.loc[1750:1869,'SO2']], aero_coeffs['E3SM']['ERFari']['BC'], aero_coeffs['E3SM']['ERFari']['OC'], aero_coeffs['E3SM']['ERFari']['SO2']) - ts1850ari
E3SMaci[:120] = ghan([emissions.loc[1750:1869,'SO2'], emissions.loc[1750:1869,'BC']+emissions.loc[1750:1869,'OC']], aero_coeffs['E3SM']['ERFaci']['beta'], aero_coeffs['E3SM']['ERFaci']['n0'], aero_coeffs['E3SM']['ERFaci']['n1']) - ts1850aci
E3SMari[265:] = aerocom([emissions.loc[2015:2019,'BC'], emissions.loc[2015:2019,'OC'], emissions.loc[2015:2019,'SO2']], aero_coeffs['E3SM']['ERFari']['BC'], aero_coeffs['E3SM']['ERFari']['OC'], aero_coeffs['E3SM']['ERFari']['SO2']) - ts1850ari
E3SMaci[265:] = ghan([emissions.loc[2015:2019,'SO2'], emissions.loc[2015:2019,'BC']+emissions.loc[2015:2019,'OC']], aero_coeffs['E3SM']['ERFaci']['beta'], aero_coeffs['E3SM']['ERFaci']['n0'], aero_coeffs['E3SM']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['E3SM'][:,i] = (E3SMaci-E3SMaci[0])/E3SM2010aci*(ERFaci_scale[i])
ERFari['E3SM'][:,i] = (E3SMari-E3SMari[0])/E3SM2010ari*(ERFari_scale[i])
temp['E3SM'] = np.zeros((270, samples))
ohc['E3SM'] = np.zeros((270, samples))
hflux['E3SM'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['E3SM'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['E3SM'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=0.2
)
out = scm.run()
temp['E3SM'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['E3SM'][:,i] = out.ohc
hflux['E3SM'][:,i] = out.hflux
ks['temp']['E3SM'] = knutti_score(Tobs, temp['E3SM'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['E3SM'] = simple_weight(358, 10*(ohc['E3SM'][268,:]-ohc['E3SM'][221,:]), sigma_D=37)
ks['multi']['E3SM'] = (ks['temp']['E3SM']*ks['ohc']['E3SM'])/(np.sum(ks['temp']['E3SM']*ks['ohc']['E3SM']))
```
## GFDL-CM4
```
GFDL2010ari = GFDL_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+GFDL_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
GFDL2010aci = GFDL_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+GFDL_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['GFDL-CM4'] = np.zeros((270,samples))
ERFaci['GFDL-CM4'] = np.zeros((270,samples))
GFDLari = np.zeros(270)
GFDLaci = np.zeros(270)
GFDLari[100:] = GFDL_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+GFDL_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
GFDLaci[100:] = GFDL_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+GFDL_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['GFDL-CM4']['ERFari']['BC'], aero_coeffs['GFDL-CM4']['ERFari']['OC'], aero_coeffs['GFDL-CM4']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['GFDL-CM4']['ERFaci']['beta'], aero_coeffs['GFDL-CM4']['ERFaci']['n0'], aero_coeffs['GFDL-CM4']['ERFaci']['n1'])
GFDLari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['GFDL-CM4']['ERFari']['BC'], aero_coeffs['GFDL-CM4']['ERFari']['OC'], aero_coeffs['GFDL-CM4']['ERFari']['SO2']) - ts1850ari
GFDLaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['GFDL-CM4']['ERFaci']['beta'], aero_coeffs['GFDL-CM4']['ERFaci']['n0'], aero_coeffs['GFDL-CM4']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['GFDL-CM4'][:,i] = (GFDLaci-GFDLaci[0])/GFDL2010aci*(ERFaci_scale[i])
ERFari['GFDL-CM4'][:,i] = (GFDLari-GFDLari[0])/GFDL2010ari*(ERFari_scale[i])
temp['GFDL-CM4'] = np.zeros((270, samples))
ohc['GFDL-CM4'] = np.zeros((270, samples))
hflux['GFDL-CM4'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['GFDL-CM4'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['GFDL-CM4'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['GFDL-CM4'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['GFDL-CM4'][:,i] = out.ohc
hflux['GFDL-CM4'][:,i] = out.hflux
ks['temp']['GFDL-CM4'] = knutti_score(Tobs, temp['GFDL-CM4'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['GFDL-CM4'] = simple_weight(358, 10*(ohc['GFDL-CM4'][268,:]-ohc['GFDL-CM4'][221,:]), sigma_D=37)
ks['multi']['GFDL-CM4'] = (ks['temp']['GFDL-CM4']*ks['ohc']['GFDL-CM4'])/(np.sum(ks['temp']['GFDL-CM4']*ks['ohc']['GFDL-CM4']))
```
## GISS-E2-1-G
```
GISS2010ari = GISS_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+GISS_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
GISS2010aci = GISS_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+GISS_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['GISS-E2-1-G'] = np.zeros((270,samples))
ERFaci['GISS-E2-1-G'] = np.zeros((270,samples))
GISSari = np.zeros(270)
GISSaci = np.zeros(270)
GISSari[100:] = GISS_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+GISS_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
GISSaci[100:] = GISS_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+GISS_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['GISS-E2-1-G']['ERFari']['BC'], aero_coeffs['GISS-E2-1-G']['ERFari']['OC'], aero_coeffs['GISS-E2-1-G']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['GISS-E2-1-G']['ERFaci']['beta'], aero_coeffs['GISS-E2-1-G']['ERFaci']['n0'], aero_coeffs['GISS-E2-1-G']['ERFaci']['n1'])
GISSari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['GISS-E2-1-G']['ERFari']['BC'], aero_coeffs['GISS-E2-1-G']['ERFari']['OC'], aero_coeffs['GISS-E2-1-G']['ERFari']['SO2']) - ts1850ari
GISSaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['GISS-E2-1-G']['ERFaci']['beta'], aero_coeffs['GISS-E2-1-G']['ERFaci']['n0'], aero_coeffs['GISS-E2-1-G']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['GISS-E2-1-G'][:,i] = (GISSaci-GISSaci[0])/GISS2010aci*(ERFaci_scale[i])
ERFari['GISS-E2-1-G'][:,i] = (GISSari-GISSari[0])/GISS2010ari*(ERFari_scale[i])
temp['GISS-E2-1-G'] = np.zeros((270, samples))
ohc['GISS-E2-1-G'] = np.zeros((270, samples))
hflux['GISS-E2-1-G'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['GISS-E2-1-G'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['GISS-E2-1-G'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['GISS-E2-1-G'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['GISS-E2-1-G'][:,i] = out.ohc
hflux['GISS-E2-1-G'][:,i] = out.hflux
ks['temp']['GISS-E2-1-G'] = knutti_score(Tobs, temp['GISS-E2-1-G'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['GISS-E2-1-G'] = simple_weight(358, 10*(ohc['GISS-E2-1-G'][268,:]-ohc['GISS-E2-1-G'][221,:]), sigma_D=37)
ks['multi']['GISS-E2-1-G'] = (ks['temp']['GISS-E2-1-G']*ks['ohc']['GISS-E2-1-G'])/(np.sum(ks['temp']['GISS-E2-1-G']*ks['ohc']['GISS-E2-1-G']))
```
## IPSL-CM6A-LR
```
IPSL2010ari = IPSL_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+IPSL_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
IPSL2010aci = IPSL_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+IPSL_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['IPSL-CM6A-LR'] = np.zeros((270,samples))
ERFaci['IPSL-CM6A-LR'] = np.zeros((270,samples))
IPSLari = np.zeros(270)
IPSLaci = np.zeros(270)
IPSLari[100:] = IPSL_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+IPSL_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
IPSLaci[100:] = IPSL_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+IPSL_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['IPSL-CM6A-LR']['ERFari']['BC'], aero_coeffs['IPSL-CM6A-LR']['ERFari']['OC'], aero_coeffs['IPSL-CM6A-LR']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['beta'], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['n0'], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['n1'])
IPSLari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['IPSL-CM6A-LR']['ERFari']['BC'], aero_coeffs['IPSL-CM6A-LR']['ERFari']['OC'], aero_coeffs['IPSL-CM6A-LR']['ERFari']['SO2']) - ts1850ari
IPSLaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['beta'], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['n0'], aero_coeffs['IPSL-CM6A-LR']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['IPSL-CM6A-LR'][:,i] = (IPSLaci-IPSLaci[0])/IPSL2010aci*(ERFaci_scale[i])
ERFari['IPSL-CM6A-LR'][:,i] = (IPSLari-IPSLari[0])/IPSL2010ari*(ERFari_scale[i])
temp['IPSL-CM6A-LR'] = np.zeros((270, samples))
ohc['IPSL-CM6A-LR'] = np.zeros((270, samples))
hflux['IPSL-CM6A-LR'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['IPSL-CM6A-LR'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['IPSL-CM6A-LR'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['IPSL-CM6A-LR'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['IPSL-CM6A-LR'][:,i] = out.ohc
hflux['IPSL-CM6A-LR'][:,i] = out.hflux
ks['temp']['IPSL-CM6A-LR'] = knutti_score(Tobs, temp['IPSL-CM6A-LR'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['IPSL-CM6A-LR'] = simple_weight(358, 10*(ohc['IPSL-CM6A-LR'][268,:]-ohc['IPSL-CM6A-LR'][221,:]), sigma_D=37)
ks['multi']['IPSL-CM6A-LR'] = (ks['temp']['IPSL-CM6A-LR']*ks['ohc']['IPSL-CM6A-LR'])/(np.sum(ks['temp']['IPSL-CM6A-LR']*ks['ohc']['IPSL-CM6A-LR']))
```
## MIROC6
```
MIROC2010ari = MIROC6_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+MIROC6_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
MIROC2010aci = MIROC6_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+MIROC6_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['MIROC6'] = np.zeros((270,samples))
ERFaci['MIROC6'] = np.zeros((270,samples))
MIROCari = np.zeros(270)
MIROCaci = np.zeros(270)
MIROCari[100:] = MIROC6_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+MIROC6_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
MIROCaci[100:] = MIROC6_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+MIROC6_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['MIROC6']['ERFari']['BC'], aero_coeffs['MIROC6']['ERFari']['OC'], aero_coeffs['MIROC6']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['MIROC6']['ERFaci']['beta'], aero_coeffs['MIROC6']['ERFaci']['n0'], aero_coeffs['MIROC6']['ERFaci']['n1'])
MIROCari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['MIROC6']['ERFari']['BC'], aero_coeffs['MIROC6']['ERFari']['OC'], aero_coeffs['MIROC6']['ERFari']['SO2']) - ts1850ari
MIROCaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['MIROC6']['ERFaci']['beta'], aero_coeffs['MIROC6']['ERFaci']['n0'], aero_coeffs['MIROC6']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['MIROC6'][:,i] = (MIROCaci-MIROCaci[0])/MIROC2010aci*(ERFaci_scale[i])
ERFari['MIROC6'][:,i] = (MIROCari-MIROCari[0])/MIROC2010ari*(ERFari_scale[i])
temp['MIROC6'] = np.zeros((270, samples))
ohc['MIROC6'] = np.zeros((270, samples))
hflux['MIROC6'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['MIROC6'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['MIROC6'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['MIROC6'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['MIROC6'][:,i] = out.ohc
hflux['MIROC6'][:,i] = out.hflux
ks['temp']['MIROC6'] = knutti_score(Tobs, temp['MIROC6'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['MIROC6'] = simple_weight(358, 10*(ohc['MIROC6'][268,:]-ohc['MIROC6'][221,:]), sigma_D=37)
ks['multi']['MIROC6'] = (ks['temp']['MIROC6']*ks['ohc']['MIROC6'])/(np.sum(ks['temp']['MIROC6']*ks['ohc']['MIROC6']))
```
## NorESM2
```
NorESM2010ari = NorESM2_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+NorESM2_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
NorESM2010aci = NorESM2_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+NorESM2_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['NorESM2-LM'] = np.zeros((270,samples))
ERFaci['NorESM2-LM'] = np.zeros((270,samples))
NorESMari = np.zeros(270)
NorESMaci = np.zeros(270)
NorESMari[100:] = NorESM2_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+NorESM2_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
NorESMaci[100:] = NorESM2_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+NorESM2_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['NorESM2-LM']['ERFari']['BC'], aero_coeffs['NorESM2-LM']['ERFari']['OC'], aero_coeffs['NorESM2-LM']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['NorESM2-LM']['ERFaci']['beta'], aero_coeffs['NorESM2-LM']['ERFaci']['n0'], aero_coeffs['NorESM2-LM']['ERFaci']['n1'])
NorESMari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['NorESM2-LM']['ERFari']['BC'], aero_coeffs['NorESM2-LM']['ERFari']['OC'], aero_coeffs['NorESM2-LM']['ERFari']['SO2']) - ts1850ari
NorESMaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['NorESM2-LM']['ERFaci']['beta'], aero_coeffs['NorESM2-LM']['ERFaci']['n0'], aero_coeffs['NorESM2-LM']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['NorESM2-LM'][:,i] = (NorESMaci-NorESMaci[0])/NorESM2010aci*(ERFaci_scale[i])
ERFari['NorESM2-LM'][:,i] = (NorESMari-NorESMari[0])/NorESM2010ari*(ERFari_scale[i])
temp['NorESM2-LM'] = np.zeros((270, samples))
ohc['NorESM2-LM'] = np.zeros((270, samples))
hflux['NorESM2-LM'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['NorESM2-LM'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['NorESM2-LM'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['NorESM2-LM'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['NorESM2-LM'][:,i] = out.ohc
hflux['NorESM2-LM'][:,i] = out.hflux
ks['temp']['NorESM2-LM'] = knutti_score(Tobs, temp['NorESM2-LM'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['NorESM2-LM'] = simple_weight(358, 10*(ohc['NorESM2-LM'][268,:]-ohc['NorESM2-LM'][221,:]), sigma_D=37)
ks['multi']['NorESM2-LM'] = (ks['temp']['NorESM2-LM']*ks['ohc']['NorESM2-LM'])/(np.sum(ks['temp']['NorESM2-LM']*ks['ohc']['NorESM2-LM']))
```
## HadGEM3-GC31-LL
```
HadGEM2010ari = HadGEM3_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+HadGEM3_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
HadGEM2010aci = HadGEM3_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+HadGEM3_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['HadGEM3-GC31-LL'] = np.zeros((270,samples))
ERFaci['HadGEM3-GC31-LL'] = np.zeros((270,samples))
HadGEMari = np.zeros(270)
HadGEMaci = np.zeros(270)
HadGEMari[100:] = HadGEM3_aerforcing.loc[0:169, 'aprp_ERFariSW'].values+HadGEM3_aerforcing.loc[0:169, 'aprp_ERFariLW'].values
HadGEMaci[100:] = HadGEM3_aerforcing.loc[0:169, 'aprp_ERFaciSW'].values+HadGEM3_aerforcing.loc[0:169, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['BC'], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['OC'], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['beta'], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['n0'], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['n1'])
HadGEMari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['BC'], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['OC'], aero_coeffs['HadGEM3-GC31-LL']['ERFari']['SO2']) - ts1850ari
HadGEMaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['beta'], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['n0'], aero_coeffs['HadGEM3-GC31-LL']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['HadGEM3-GC31-LL'][:,i] = (HadGEMaci-HadGEMaci[0])/HadGEM2010aci*(ERFaci_scale[i])
ERFari['HadGEM3-GC31-LL'][:,i] = (HadGEMari-HadGEMari[0])/HadGEM2010ari*(ERFari_scale[i])
temp['HadGEM3-GC31-LL'] = np.zeros((270, samples))
ohc['HadGEM3-GC31-LL'] = np.zeros((270, samples))
hflux['HadGEM3-GC31-LL'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['HadGEM3-GC31-LL'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['HadGEM3-GC31-LL'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['HadGEM3-GC31-LL'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['HadGEM3-GC31-LL'][:,i] = out.ohc
hflux['HadGEM3-GC31-LL'][:,i] = out.hflux
ks['temp']['HadGEM3-GC31-LL'] = knutti_score(Tobs, temp['HadGEM3-GC31-LL'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['HadGEM3-GC31-LL'] = simple_weight(358, 10*(ohc['HadGEM3-GC31-LL'][268,:]-ohc['HadGEM3-GC31-LL'][221,:]), sigma_D=37)
ks['multi']['HadGEM3-GC31-LL'] = (ks['temp']['HadGEM3-GC31-LL']*ks['ohc']['HadGEM3-GC31-LL'])/(np.sum(ks['temp']['HadGEM3-GC31-LL']*ks['ohc']['HadGEM3-GC31-LL']))
```
## MRI-ESM2-0
```
MRI2010ari = MRI_aerforcing.loc[155:165,'aprp_ERFariSW'].mean()+MRI_aerforcing.loc[155:165,'aprp_ERFariLW'].mean()
MRI2010aci = MRI_aerforcing.loc[155:165,'aprp_ERFaciSW'].mean()+MRI_aerforcing.loc[155:165,'aprp_ERFaciLW'].mean()
ERFari['MRI-ESM2-0'] = np.zeros((270,samples))
ERFaci['MRI-ESM2-0'] = np.zeros((270,samples))
MRIari = np.zeros(270)
MRIaci = np.zeros(270)
MRIari[100:265] = MRI_aerforcing.loc[0:165, 'aprp_ERFariSW'].values+MRI_aerforcing.loc[0:165, 'aprp_ERFariLW'].values
MRIaci[100:265] = MRI_aerforcing.loc[0:165, 'aprp_ERFaciSW'].values+MRI_aerforcing.loc[0:165, 'aprp_ERFaciLW'].values
ts1850ari = aerocom([emissions.loc[1850,'BC'], emissions.loc[1850,'OC'], emissions.loc[1850,'SO2']], aero_coeffs['MRI-ESM2-0']['ERFari']['BC'], aero_coeffs['MRI-ESM2-0']['ERFari']['OC'], aero_coeffs['MRI-ESM2-0']['ERFari']['SO2'])
ts1850aci = ghan([emissions.loc[1850,'SO2'], emissions.loc[1850,'BC']+emissions.loc[1850,'OC']], aero_coeffs['MRI-ESM2-0']['ERFaci']['beta'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n0'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n1'])
MRIari[:100] = aerocom([emissions.loc[1750:1849,'BC'], emissions.loc[1750:1849,'OC'], emissions.loc[1750:1849,'SO2']], aero_coeffs['MRI-ESM2-0']['ERFari']['BC'], aero_coeffs['MRI-ESM2-0']['ERFari']['OC'], aero_coeffs['MRI-ESM2-0']['ERFari']['SO2']) - ts1850ari
MRIaci[:100] = ghan([emissions.loc[1750:1849,'SO2'], emissions.loc[1750:1849,'BC']+emissions.loc[1750:1849,'OC']], aero_coeffs['MRI-ESM2-0']['ERFaci']['beta'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n0'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n1']) - ts1850aci
MRIari[265:] = aerocom([emissions.loc[2015:2019,'BC'], emissions.loc[2015:2019,'OC'], emissions.loc[2015:2019,'SO2']], aero_coeffs['MRI-ESM2-0']['ERFari']['BC'], aero_coeffs['MRI-ESM2-0']['ERFari']['OC'], aero_coeffs['MRI-ESM2-0']['ERFari']['SO2']) - ts1850ari
MRIaci[265:] = ghan([emissions.loc[2015:2019,'SO2'], emissions.loc[2015:2019,'BC']+emissions.loc[2015:2019,'OC']], aero_coeffs['MRI-ESM2-0']['ERFaci']['beta'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n0'], aero_coeffs['MRI-ESM2-0']['ERFaci']['n1']) - ts1850aci
for i in range(samples):
ERFaci['MRI-ESM2-0'][:,i] = (MRIaci-MRIaci[0])/MRI2010aci*(ERFaci_scale[i])
ERFari['MRI-ESM2-0'][:,i] = (MRIari-MRIari[0])/MRI2010ari*(ERFari_scale[i])
temp['MRI-ESM2-0'] = np.zeros((270, samples))
ohc['MRI-ESM2-0'] = np.zeros((270, samples))
hflux['MRI-ESM2-0'] = np.zeros((270, samples))
for i in tqdm_notebook(range(samples)):
in_forcing = baseline_forcing.copy()
in_forcing.drop(['aerosol-radiation_interactions', 'aerosol-cloud_interactions','total_anthropogenic','total_natural','total'], axis=1, inplace=True)
in_forcing = in_forcing * scale_df.loc[i,:]
in_forcing['solar'] = in_forcing['solar'] + np.linspace(0, trend_solar[i], 270)
in_forcing['aerosol-radiation_interactions'] = ERFari['MRI-ESM2-0'][:270,i]
in_forcing['aerosol-cloud_interactions'] = ERFaci['MRI-ESM2-0'][:270,i]
in_forcing['total'] = in_forcing.sum(axis=1)
scm = TwoLayerModel(
extforce=in_forcing['total'],
exttime=in_forcing.index,
tbeg=1750,
tend=2020,
q2x=geoff_sample_df.loc[i,'q4x']/2,
lamg=geoff_sample_df.loc[i,'lamg'],
t2x=None,
eff=geoff_sample_df.loc[i,'eff'],
cmix=geoff_sample_df.loc[i,'cmix'],
cdeep=geoff_sample_df.loc[i,'cdeep'],
gamma_2l=geoff_sample_df.loc[i,'gamma_2l'],
outtime=np.arange(1750.5,2020),
dt=1
)
out = scm.run()
temp['MRI-ESM2-0'][:,i] = out.tg - np.mean(out.tg[100:150])
ohc['MRI-ESM2-0'][:,i] = out.ohc
hflux['MRI-ESM2-0'][:,i] = out.hflux
ks['temp']['MRI-ESM2-0'] = knutti_score(Tobs, temp['MRI-ESM2-0'][100:270, :] + intvar[100:270,:samples], sigma_D=0.12)
ks['ohc']['MRI-ESM2-0'] = simple_weight(358, 10*(ohc['MRI-ESM2-0'][268,:]-ohc['MRI-ESM2-0'][221,:]), sigma_D=37)
ks['multi']['MRI-ESM2-0'] = (ks['temp']['MRI-ESM2-0']*ks['ohc']['MRI-ESM2-0'])/(np.sum(ks['temp']['MRI-ESM2-0']*ks['ohc']['MRI-ESM2-0']))
save_dict_to_hdf5(ERFari, '../data_output/results/ERFari.h5')
save_dict_to_hdf5(ERFaci, '../data_output/results/ERFaci.h5')
save_dict_to_hdf5(temp, '../data_output/results/temp.h5')
save_dict_to_hdf5(ks, '../data_output/results/knutti_score.h5')
save_dict_to_hdf5(ohc, '../data_output/results/ohc.h5')
save_dict_to_hdf5(hflux, '../data_output/results/hflux.h5')
```
# After runs, start here
```
expts = ['CMIP6-constrained','CanESM5','E3SM','GFDL-CM4','GFDL-ESM4','GISS-E2-1-G','HadGEM3-GC31-LL','IPSL-CM6A-LR','MIROC6','MRI-ESM2-0','NorESM2-LM','Oslo-CTM3','UKESM1-0-LL']
#expts_all = ['CMIP6-SSP1-1.9','CMIP6-SSP2-4.5','CMIP6-SSP3-7.0','CanESM5','E3SM','GFDL-CM4','GISS-E2-1-G','HadGEM3-GC31-LL','MIROC6','NorESM2-LM','Lund','AR5']
fig, ax = pl.subplots()
for expt in expts:
ax.plot(np.arange(1750.5,2020), 10*np.nansum((ohc[expt]-ohc[expt][221,:])*ks['multi'][expt], axis=1), color=colors[expt])
ax.set_title('Best estimate aerosol forcing')
ax.set_ylabel('Ocean heat uptake relative to 1960, ZJ')
ax.plot(np.arange(1960.5,2019), (OHCobs-OHCobs[11]), color=colors['observations'])
ax.set_xlim(1960,2019)
ax.set_ylim(-50,400)
# Throw TCR into the mix
tcr = geoff_sample_df['q4x'][:samples]/2/(geoff_sample_df[:samples]['lamg'] + geoff_sample_df[:samples]['eff']*geoff_sample_df[:samples]['gamma_2l'])
ecs = geoff_sample_df['q4x'][:samples]/2/(geoff_sample_df[:samples]['lamg'])
pc = {}
for expt in tqdm_notebook(expts):
pc[expt] = {}
for constraint in ['temp', 'ohc', 'multi']:
pc[expt][constraint] = {}
pc[expt][constraint]['ECS'] = {}
pc[expt][constraint]['TCR'] = {}
for metric in ['GSAT','OHC','ERFari','ERFaci','ERFaer']:
pc[expt][constraint][metric] = {}
for perc in ['5','16','50','84','95']:
pc[expt][constraint][metric][perc] = np.zeros(270)
(
pc[expt][constraint]['ECS']['5'],
pc[expt][constraint]['ECS']['16'],
pc[expt][constraint]['ECS']['50'],
pc[expt][constraint]['ECS']['84'],
pc[expt][constraint]['ECS']['95']
) = (
weighted_percentile(ecs, ks[constraint][expt], [.05,.16,.5,.84,.95])
)
(
pc[expt][constraint]['TCR']['5'],
pc[expt][constraint]['TCR']['16'],
pc[expt][constraint]['TCR']['50'],
pc[expt][constraint]['TCR']['84'],
pc[expt][constraint]['TCR']['95']
) = weighted_percentile(tcr, ks[constraint][expt], [.05,.16,.5,.84,.95])
for year in range(270):
(
pc[expt][constraint]['GSAT']['5'][year],
pc[expt][constraint]['GSAT']['16'][year],
pc[expt][constraint]['GSAT']['50'][year],
pc[expt][constraint]['GSAT']['84'][year],
pc[expt][constraint]['GSAT']['95'][year]
) = weighted_percentile(temp[expt][year,:] + intvar[year,:samples], ks[constraint][expt], [.05,.16,.5,.84,.95])
(
pc[expt][constraint]['OHC']['5'][year],
pc[expt][constraint]['OHC']['16'][year],
pc[expt][constraint]['OHC']['50'][year],
pc[expt][constraint]['OHC']['84'][year],
pc[expt][constraint]['OHC']['95'][year]
) = weighted_percentile(ohc[expt][year,:], ks[constraint][expt], [.05,.16,.5,.84,.95])
(
pc[expt][constraint]['ERFari']['5'][year],
pc[expt][constraint]['ERFari']['16'][year],
pc[expt][constraint]['ERFari']['50'][year],
pc[expt][constraint]['ERFari']['84'][year],
pc[expt][constraint]['ERFari']['95'][year]
) = weighted_percentile(ERFari[expt][year,:], ks[constraint][expt], [.05,.16,.5,.84,.95])
(
pc[expt][constraint]['ERFaci']['5'][year],
pc[expt][constraint]['ERFaci']['16'][year],
pc[expt][constraint]['ERFaci']['50'][year],
pc[expt][constraint]['ERFaci']['84'][year],
pc[expt][constraint]['ERFaci']['95'][year]
) = weighted_percentile(ERFaci[expt][year,:], ks[constraint][expt], [.05,.16,.5,.84,.95])
(
pc[expt][constraint]['ERFaer']['5'][year],
pc[expt][constraint]['ERFaer']['16'][year],
pc[expt][constraint]['ERFaer']['50'][year],
pc[expt][constraint]['ERFaer']['84'][year],
pc[expt][constraint]['ERFaer']['95'][year]
) = weighted_percentile(ERFari[expt][year,:]+ERFaci[expt][year,:], ks[constraint][expt], [.05,.16,.5,.84,.95])
save_dict_to_hdf5(pc, '../data_output/results/pc.h5')
```
| github_jupyter |
```
#tag::make_dask_client[]
import dask
from dask.distributed import Client
client = Client() # Here we could specify a cluster, defaults to local mode
#end::make_dask_client[]
#tag::sleepy_task_hello_world[]
import timeit
def slow_task(x):
import time
time.sleep(2) # Do something sciency/business
return x
things = range(10)
very_slow_result = map(slow_task, things)
slowish_result = map(dask.delayed(slow_task), things)
slow_time = timeit.timeit(lambda: list(very_slow_result), number=1)
fast_time = timeit.timeit(lambda: list(dask.compute(*slowish_result)), number=1)
print("In sequence {}, in parallel {}".format(slow_time, fast_time))
#end::sleepy_task_hello_world[]
# Note: if we were on a "real" cluster we'd have to do more magic to install it on all the nodes in the cluster.
!pip install bs4
#tag::mini_crawl_task[]
@dask.delayed
def crawl(url, depth=0, maxdepth=1, maxlinks=4):
links = []
link_futures = []
try:
import requests
from bs4 import BeautifulSoup
f = requests.get(url)
links += [(url, f.text)]
if (depth > maxdepth):
return links # base case
soup = BeautifulSoup(f.text, 'html.parser')
c = 0
for link in soup.find_all('a'):
if "href" in link:
c = c + 1
link_futures += crawl(link["href"], depth=(depth+1), maxdepth=maxdepth)
# Don't branch too much were still in local mode and the web is big
if c > maxlinks:
break
for r in dask.compute(link_futures):
links += r
return links
except requests.exceptions.InvalidSchema:
return [] # Skip non-web links
dask.compute(crawl("http://holdenkarau.com/"))
#end::mini_crawl_task[]
#tag::make_bag_of_crawler[]
import dask.bag as db
githubs = ["https://github.com/scalingpythonml/scalingpythonml", "https://github.com/dask/distributed"]
initial_bag = db.from_delayed(map(crawl, githubs))
#end::make_bag_of_crawler[]
#tag::make_a_bag_of_words[]
words_bag = initial_bag.map(lambda url_contents: url_contents[1].split(" ")).flatten()
#end::make_a_bag_of_words[]
#tag::wc_freq[]
dask.compute(words_bag.frequencies())
#end::wc_freq[]
#tag::wc_func[]
def make_word_tuple(w):
return (w, 1)
def get_word(word_count):
return word_count[0]
def sum_word_counts(wc1, wc2):
return (wc1[0], wc1[1] + wc2[1])
word_count = words_bag.map(make_word_tuple).foldby(get_word, sum_word_counts)
#end::wc_func[]
dask.compute(word_count)
```
| github_jupyter |
#Principal Components Analysis (PCA) of natural images
The idea is that each subimage will be used as a statistical sample. We compute the outer product of each, and the average all samples to get an estimate of the autocovariance matrix.
```
%matplotlib inline
import matplotlib.pyplot as plt # matplotlib provides plot functions similar to MATLAB
import numpy as np
from skimage import color, filter # skimage is an image processing library
imFile = '../data/imgstat/alpine.png'
# imFile = '../data/imgstat/lake.jpg'
im = color.rgb2gray(plt.imread(imFile))
plt.imshow(im, 'gray')
```
##Break a large image into a series of 16x16 subimages
```
# Crop patches from image
patchSize = 16; nDim = patchSize ** 2;
w = im.shape[0]; h = im.shape[1];
nx = im.shape[0] / patchSize; ny = im.shape[1] / patchSize
nSamples = nx * ny
patches = np.zeros((nDim, nSamples)) # column vectors
[X, Y] = np.meshgrid(np.arange(0, w, patchSize), np.arange(0, h, patchSize))
X = X.flatten(); Y = Y.flatten()
for i in range(X.size):
x = X[i]
y = Y[i]
patch = im[x:x+patchSize, y:y+patchSize]
patch = patch - patch.mean()
patches[:,i] = patch.flatten().T
```
##Calculate autocovariance matrix
```
autoCov = np.dot(patches, patches.T) / (nx * ny)
plt.imshow(autoCov)
# io.imshow(patches[0])
```
##Calculate the eigenvectors and eigenvalues of the autocovariance matrix
```
[eigenVals, eigenVecs] = np.linalg.eig(autoCov)
plt.plot(eigenVals, '*')
```
##Display the first 16 eigenvectors as "eigenpictures"
```
for i in range(16):
plt.subplot(4,4,i+1)
plt.imshow(eigenVecs[:,i].reshape((patchSize, patchSize)), 'gray')
plt.axis('off')
```
##Oja's weight normalization rule for Hebbian learning
Oja's rule can be used to iteratively estimate the first principle component
$$ \Delta w_i = \alpha (x_i y - y^2 w_i), \; i= 1,...,n. $$
```
learningRate = 0.01
# Randomly initialize the weight matrix
W = np.random.rand(16, nDim)
nIter = 16
X = patches
plt.figure()
plt.suptitle('Iteration')
for i in range(nIter):
plt.subplot(4,4,i+1)
for j in range(nSamples):
x = X[:,j]
y = np.dot(W, x)
# W: (16, 256), y: (16, 1), x: (256, 1)
# inc = learningRate * (np.outer(y, x.T) - W * np.repeat(y * y,256).reshape(16,256))
inc = learningRate * (np.outer(y, x.T) - W * np.tile((y * y)[:,np.newaxis],(1,256)))
W += inc
plt.imshow(W[0,:].reshape((patchSize, patchSize)), 'gray')
plt.axis('off')
plt.figure()
plt.suptitle('Visualize weight matrix')
for i in range(16):
plt.subplot(4,4,i+1)
plt.imshow(W[i,:].reshape((patchSize, patchSize)), 'gray')
plt.axis('off')
```
| github_jupyter |
# Exercise 1: Uniqueness
## The problem
In [the first tutorial](../tutorials/1_sphere_scatterer_null_field.ipynb), we looked at two formulations for a scattering problem with a Neumann boundary condition.
In this exercise, we will investigate the uniqueness so solutions to the boundary integral formulations for this problem.
## The formulation
In this exercise we will use the null field approach (as in [the first tutorial](../tutorials/1_sphere_scatterer_null_field.ipynb). This uses the following representation formula and boundary integral equation.
### Representation formula
$$
p_\text{total} = \mathcal{D}p_\text{total} + p_\text{inc}
$$
where $\mathcal{D}$ is the double layer potential operator.
### Boundary integral equation
$$
(\mathsf{D}-\tfrac{1}{2}\mathsf{I})p_\text{total} = -p_\text{inc},
$$
where $\mathsf{D}$ is the double layer boundary operator, and $\mathsf{I}$ is the identity operator.
## Finding a resonance
The code below plots the condition number of $\mathsf{D}-\tfrac{1}{2}\mathsf{I}$ for 30 values of $k$ between 2.5 and 3.5. There is a sharp increase in the condition number near 3.2.
Adjust the limits use in `np.linspace` to approximate the value of $k$ for this spike to 4 or 5 decimal places. (For example, you might start be reducing the search to between 3.0 and 3.3, so `np.linspace(3.0, 3.3, 30)`.)
```
%matplotlib inline
import bempp.api
from bempp.api.operators.boundary import helmholtz, sparse
from bempp.api.operators.potential import helmholtz as helmholtz_potential
from bempp.api.linalg import gmres
import numpy as np
from matplotlib import pyplot as plt
grid = bempp.api.shapes.regular_sphere(3)
space = bempp.api.function_space(grid, "DP", 0)
identity = sparse.identity(space, space, space)
x_data = []
y_data = []
for k in np.linspace(2.5, 3.5, 30):
double_layer = helmholtz.double_layer(space, space, space, k)
x_data.append(k)
y_data.append(np.linalg.cond((double_layer - 0.5 * identity).weak_form().to_dense()))
plt.plot(x_data, y_data)
plt.xlabel("Wavenumber ($k$)")
plt.ylabel("Condition number")
plt.show()
```
## The effect on the solution
The code below has been copied from [the first tutorial](../tutorials/1_sphere_scatterer_null_field.ipynb) and the wavenumber has been changed to 3. The solution plot looks like a reasonable soluition.
Change the value of the wavenumber to the resonance that you found above. Does the solution still look reasonable?
```
%matplotlib inline
import bempp.api
from bempp.api.operators.boundary import helmholtz, sparse
from bempp.api.operators.potential import helmholtz as helmholtz_potential
from bempp.api.linalg import gmres
import numpy as np
from matplotlib import pyplot as plt
k = 3.
grid = bempp.api.shapes.regular_sphere(3)
space = bempp.api.function_space(grid, "DP", 0)
identity = sparse.identity(space, space, space)
double_layer = helmholtz.double_layer(space, space, space, k)
@bempp.api.complex_callable
def p_inc_callable(x, n, domain_index, result):
result[0] = np.exp(1j * k * x[0])
p_inc = bempp.api.GridFunction(space, fun=p_inc_callable)
p_total, info = gmres(double_layer - 0.5 * identity, -p_inc, tol=1E-5)
Nx = 200
Ny = 200
xmin, xmax, ymin, ymax = [-3, 3, -3, 3]
plot_grid = np.mgrid[xmin:xmax:Nx * 1j, ymin:ymax:Ny * 1j]
points = np.vstack((plot_grid[0].ravel(),
plot_grid[1].ravel(),
np.zeros(plot_grid[0].size)))
p_inc_evaluated = np.real(np.exp(1j * k * points[0, :]))
p_inc_evaluated = p_inc_evaluated.reshape((Nx, Ny))
double_pot = helmholtz_potential.double_layer(space, points, k)
p_s = np.real(double_pot.evaluate(p_total))
p_s = p_s.reshape((Nx, Ny))
vmax = max(np.abs((p_inc_evaluated + p_s).flat))
fig = plt.figure(figsize=(10, 8))
plt.imshow(np.real((p_inc_evaluated + p_s).T), extent=[-3, 3, -3, 3],
cmap=plt.get_cmap("bwr"), vmin=-vmax, vmax=vmax)
plt.xlabel('x')
plt.ylabel('y')
plt.colorbar()
plt.title("Total wave in the plane z=0")
plt.show()
```
## Obtaining the solution for this wavenumber: the Burton–Miller formulation
The Burton–Miller formulation can be used to obtain solutions to acoustic problems while avoiding spurious resonances.
### Representation formula
$$
p_\text{s} = \mathcal{D}p_\text{total},
$$
where $\mathcal{D}$ is the double layer potential operator.
### Boundary integral equation
$$
\left(\mathsf{D}-\tfrac{1}{2}\mathsf{I}+\frac{1}{\mathsf{i}k}\mathsf{H}\right)p_\text{total}
=
-p_\text{inc} + \frac{1}{\mathsf{i}k}\frac{\partial p_\text{inc}}{\partial \mathbf{n}},
$$
where $\mathsf{D}$ is the double layer boundary operator; $\mathsf{H}$ is the hypersingular boundary operator; $\mathsf{D}$ is the double layer boundary operator; and $\mathsf{I}$ is the identity operator.
### Solving with Bempp
Your task is to adapt and combine the example code in [the first tutorial](../tutorials/1_sphere_scatterer_null_field.ipynb) to solve the problem at the wavenumber you found above using the Burton–Miller formulation.
We can create the hypersingular operator in Bempp by calling `helmholtz.hypersingular`. Complex number can be used in Python by writing (for example) `2 + 1j`, `3j`, or `1j * 3`. In order for the hypersingular operator to be defined, we must use a P1 space. The code needed to create the relevant operators is given below. Your task is to use these to implement the Burton–Miller formulation.
Does the solution you obtain here look more reasonable that the solution above? You might like to adapt the previous example to use a P1 space to be sure that the resonances are still a problem with this alternative space.
Hint: the normal derivative ($\frac{\partial p_\text{inc}}{\partial\mathbf{n}}$) in this case is $\mathrm{i}kn_0\mathrm{e}^{\mathrm{i}kx_0}$, where $\mathbf{n}=(n_0,n_1,n_2)$. If you're not sure how to implement this, have a look at [tutorial 2](../tutorials/2_sphere_scatterer_direct.ipynb).
```
%matplotlib inline
import bempp.api
from bempp.api.operators.boundary import helmholtz, sparse
from bempp.api.operators.potential import helmholtz as helmholtz_potential
from bempp.api.linalg import gmres
import numpy as np
from matplotlib import pyplot as plt
k = 1 # Enter your value here
grid = bempp.api.shapes.regular_sphere(3)
space = bempp.api.function_space(grid, "P", 1)
identity = sparse.identity(space, space, space)
double_layer = helmholtz.double_layer(space, space, space, k)
hypersingular = helmholtz.hypersingular(space, space, space, k)
```
## What next?
After attempting this exercises, you should read [tutorial 2](../tutorials/2_sphere_scatterer_direct.ipynb).
| github_jupyter |
# Your first neural network
In this project, you'll build your first neural network and use it to predict daily bike rental ridership. We've provided some of the code, but left the implementation of the neural network up to you (for the most part). After you've submitted this project, feel free to explore the data and the model more.
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
%config InlineBackend.figure_format = 'retina'
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
```
## Load and prepare the data
A critical step in working with neural networks is preparing the data correctly. Variables on different scales make it difficult for the network to efficiently learn the correct weights. Below, we've written the code to load and prepare the data. You'll learn more about this soon!
```
data_path = 'Bike-Sharing-Dataset/hour.csv'
rides = pd.read_csv(data_path)
rides.head()
```
## Checking out the data
This dataset has the number of riders for each hour of each day from January 1 2011 to December 31 2012. The number of riders is split between casual and registered, summed up in the `cnt` column. You can see the first few rows of the data above.
Below is a plot showing the number of bike riders over the first 10 days or so in the data set. (Some days don't have exactly 24 entries in the data set, so it's not exactly 10 days.) You can see the hourly rentals here. This data is pretty complicated! The weekends have lower over all ridership and there are spikes when people are biking to and from work during the week. Looking at the data above, we also have information about temperature, humidity, and windspeed, all of these likely affecting the number of riders. You'll be trying to capture all this with your model.
```
rides[:24*10].plot(x='dteday', y='cnt')
```
### Dummy variables
Here we have some categorical variables like season, weather, month. To include these in our model, we'll need to make binary dummy variables. This is simple to do with Pandas thanks to `get_dummies()`.
```
dummy_fields = ['season', 'weathersit', 'mnth', 'hr', 'weekday']
for each in dummy_fields:
dummies = pd.get_dummies(rides[each], prefix=each, drop_first=False)
rides = pd.concat([rides, dummies], axis=1)
fields_to_drop = ['instant', 'dteday', 'season', 'weathersit',
'weekday', 'atemp', 'mnth', 'workingday', 'hr']
data = rides.drop(fields_to_drop, axis=1)
data.head()
```
### Scaling target variables
To make training the network easier, we'll standardize each of the continuous variables. That is, we'll shift and scale the variables such that they have zero mean and a standard deviation of 1.
The scaling factors are saved so we can go backwards when we use the network for predictions.
```
quant_features = ['casual', 'registered', 'cnt', 'temp', 'hum', 'windspeed']
# Store scalings in a dictionary so we can convert back later
scaled_features = {}
for each in quant_features:
mean, std = data[each].mean(), data[each].std()
scaled_features[each] = [mean, std]
data.loc[:, each] = (data[each] - mean)/std
```
### Splitting the data into training, testing, and validation sets
We'll save the data for the last approximately 21 days to use as a test set after we've trained the network. We'll use this set to make predictions and compare them with the actual number of riders.
```
# Save data for approximately the last 21 days
test_data = data[-21*24:]
# Now remove the test data from the data set
data = data[:-21*24]
# Separate the data into features and targets
target_fields = ['cnt', 'casual', 'registered']
features, targets = data.drop(target_fields, axis=1), data[target_fields]
test_features, test_targets = test_data.drop(target_fields, axis=1), test_data[target_fields]
```
We'll split the data into two sets, one for training and one for validating as the network is being trained. Since this is time series data, we'll train on historical data, then try to predict on future data (the validation set).
```
# Hold out the last 60 days or so of the remaining data as a validation set
train_features, train_targets = features[:-60*24], targets[:-60*24]
val_features, val_targets = features[-60*24:], targets[-60*24:]
```
## Time to build the network
Below you'll build your network. We've built out the structure. You'll implement both the forward pass and backwards pass through the network. You'll also set the hyperparameters: the learning rate, the number of hidden units, and the number of training passes.
<img src="assets/neural_network.png" width=300px>
The network has two layers, a hidden layer and an output layer. The hidden layer will use the sigmoid function for activations. The output layer has only one node and is used for the regression, the output of the node is the same as the input of the node. That is, the activation function is $f(x)=x$. A function that takes the input signal and generates an output signal, but takes into account the threshold, is called an activation function. We work through each layer of our network calculating the outputs for each neuron. All of the outputs from one layer become inputs to the neurons on the next layer. This process is called *forward propagation*.
We use the weights to propagate signals forward from the input to the output layers in a neural network. We use the weights to also propagate error backwards from the output back into the network to update our weights. This is called *backpropagation*.
> **Hint:** You'll need the derivative of the output activation function ($f(x) = x$) for the backpropagation implementation. If you aren't familiar with calculus, this function is equivalent to the equation $y = x$. What is the slope of that equation? That is the derivative of $f(x)$.
Below, you have these tasks:
1. Implement the sigmoid function to use as the activation function. Set `self.activation_function` in `__init__` to your sigmoid function.
2. Implement the forward pass in the `train` method.
3. Implement the backpropagation algorithm in the `train` method, including calculating the output error.
4. Implement the forward pass in the `run` method.
```
#############
# In the my_answers.py file, fill out the TODO sections as specified
#############
from my_answers import NeuralNetwork
def MSE(y, Y):
return np.mean((y-Y)**2)
```
## Unit tests
Run these unit tests to check the correctness of your network implementation. This will help you be sure your network was implemented correctly befor you starting trying to train it. These tests must all be successful to pass the project.
```
import unittest
inputs = np.array([[0.5, -0.2, 0.1]])
targets = np.array([[0.4]])
test_w_i_h = np.array([[0.1, -0.2],
[0.4, 0.5],
[-0.3, 0.2]])
test_w_h_o = np.array([[0.3],
[-0.1]])
class TestMethods(unittest.TestCase):
##########
# Unit tests for data loading
##########
def test_data_path(self):
# Test that file path to dataset has been unaltered
self.assertTrue(data_path.lower() == 'bike-sharing-dataset/hour.csv')
def test_data_loaded(self):
# Test that data frame loaded
self.assertTrue(isinstance(rides, pd.DataFrame))
##########
# Unit tests for network functionality
##########
def test_activation(self):
network = NeuralNetwork(3, 2, 1, 0.5)
# Test that the activation function is a sigmoid
self.assertTrue(np.all(network.activation_function(0.5) == 1/(1+np.exp(-0.5))))
def test_train(self):
# Test that weights are updated correctly on training
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
network.train(inputs, targets)
self.assertTrue(np.allclose(network.weights_hidden_to_output,
np.array([[ 0.37275328],
[-0.03172939]])))
self.assertTrue(np.allclose(network.weights_input_to_hidden,
np.array([[ 0.10562014, -0.20185996],
[0.39775194, 0.50074398],
[-0.29887597, 0.19962801]])))
def test_run(self):
# Test correctness of run method
network = NeuralNetwork(3, 2, 1, 0.5)
network.weights_input_to_hidden = test_w_i_h.copy()
network.weights_hidden_to_output = test_w_h_o.copy()
self.assertTrue(np.allclose(network.run(inputs), 0.09998924))
suite = unittest.TestLoader().loadTestsFromModule(TestMethods())
unittest.TextTestRunner().run(suite)
```
## Training the network
Here you'll set the hyperparameters for the network. The strategy here is to find hyperparameters such that the error on the training set is low, but you're not overfitting to the data. If you train the network too long or have too many hidden nodes, it can become overly specific to the training set and will fail to generalize to the validation set. That is, the loss on the validation set will start increasing as the training set loss drops.
You'll also be using a method know as Stochastic Gradient Descent (SGD) to train the network. The idea is that for each training pass, you grab a random sample of the data instead of using the whole data set. You use many more training passes than with normal gradient descent, but each pass is much faster. This ends up training the network more efficiently. You'll learn more about SGD later.
### Choose the number of iterations
This is the number of batches of samples from the training data we'll use to train the network. The more iterations you use, the better the model will fit the data. However, this process can have sharply diminishing returns and can waste computational resources if you use too many iterations. You want to find a number here where the network has a low training loss, and the validation loss is at a minimum. The ideal number of iterations would be a level that stops shortly after the validation loss is no longer decreasing.
### Choose the learning rate
This scales the size of weight updates. If this is too big, the weights tend to explode and the network fails to fit the data. Normally a good choice to start at is 0.1; however, if you effectively divide the learning rate by n_records, try starting out with a learning rate of 1. In either case, if the network has problems fitting the data, try reducing the learning rate. Note that the lower the learning rate, the smaller the steps are in the weight updates and the longer it takes for the neural network to converge.
### Choose the number of hidden nodes
In a model where all the weights are optimized, the more hidden nodes you have, the more accurate the predictions of the model will be. (A fully optimized model could have weights of zero, after all.) However, the more hidden nodes you have, the harder it will be to optimize the weights of the model, and the more likely it will be that suboptimal weights will lead to overfitting. With overfitting, the model will memorize the training data instead of learning the true pattern, and won't generalize well to unseen data.
Try a few different numbers and see how it affects the performance. You can look at the losses dictionary for a metric of the network performance. If the number of hidden units is too low, then the model won't have enough space to learn and if it is too high there are too many options for the direction that the learning can take. The trick here is to find the right balance in number of hidden units you choose. You'll generally find that the best number of hidden nodes to use ends up being between the number of input and output nodes.
```
import sys
####################
### Set the hyperparameters in you myanswers.py file ###
####################
from my_answers import iterations, learning_rate, hidden_nodes, output_nodes
N_i = train_features.shape[1]
network = NeuralNetwork(N_i, hidden_nodes, output_nodes, learning_rate)
losses = {'train':[], 'validation':[]}
for ii in range(iterations):
# Go through a random batch of 128 records from the training data set
batch = np.random.choice(train_features.index, size=128)
X, y = train_features.loc[batch].values, train_targets.loc[batch]['cnt']
network.train(X, y)
# Printing out the training progress
train_loss = MSE(network.run(train_features).T, train_targets['cnt'].values)
val_loss = MSE(network.run(val_features).T, val_targets['cnt'].values)
sys.stdout.write("\rProgress: {:2.1f}".format(100 * ii/float(iterations)) \
+ "% ... Training loss: " + str(train_loss)[:5] \
+ " ... Validation loss: " + str(val_loss)[:5])
sys.stdout.flush()
losses['train'].append(train_loss)
losses['validation'].append(val_loss)
plt.plot(losses['train'], label='Training loss')
plt.plot(losses['validation'], label='Validation loss')
plt.legend()
_ = plt.ylim()
```
## Check out your predictions
Here, use the test data to view how well your network is modeling the data. If something is completely wrong here, make sure each step in your network is implemented correctly.
```
fig, ax = plt.subplots(figsize=(8,4))
mean, std = scaled_features['cnt']
predictions = network.run(test_features).T*std + mean
ax.plot(predictions[0], label='Prediction')
ax.plot((test_targets['cnt']*std + mean).values, label='Data')
ax.set_xlim(right=len(predictions))
ax.legend()
dates = pd.to_datetime(rides.loc[test_data.index]['dteday'])
dates = dates.apply(lambda d: d.strftime('%b %d'))
ax.set_xticks(np.arange(len(dates))[12::24])
_ = ax.set_xticklabels(dates[12::24], rotation=45)
```
## OPTIONAL: Thinking about your results(this question will not be evaluated in the rubric).
Answer these questions about your results. How well does the model predict the data? Where does it fail? Why does it fail where it does?
> **Note:** You can edit the text in this cell by double clicking on it. When you want to render the text, press control + enter
#### Your answer below
| github_jupyter |
# [Strings](https://docs.python.org/3/library/stdtypes.html#text-sequence-type-str)
```
my_string = 'Python is my favorite programming language!'
my_string
type(my_string)
len(my_string)
```
### Respecting [PEP8](https://www.python.org/dev/peps/pep-0008/#maximum-line-length) with long strings
```
long_story = ('Lorem ipsum dolor sit amet, consectetur adipiscing elit.'
'Pellentesque eget tincidunt felis. Ut ac vestibulum est.'
'In sed ipsum sit amet sapien scelerisque bibendum. Sed '
'sagittis purus eu diam fermentum pellentesque.')
long_story
```
## `str.replace()`
If you don't know how it works, you can always check the `help`:
```
help(str.replace)
```
This will not modify `my_string` because replace is not done in-place.
```
my_string.replace('a', '?')
print(my_string)
```
You have to store the return value of `replace` instead.
```
my_modified_string = my_string.replace('is', 'will be')
print(my_modified_string)
```
## `str.format()`
```
secret = '{} is cool'.format('Python')
print(secret)
print('My name is {} {}, you can call me {}.'.format('John', 'Doe', 'John'))
# is the same as:
print('My name is {first} {family}, you can call me {first}.'.format(first='John', family='Doe'))
```
## `str.join()`
```
pandas = 'pandas'
numpy = 'numpy'
requests = 'requests'
cool_python_libs = ', '.join([pandas, numpy, requests])
print('Some cool python libraries: {}'.format(cool_python_libs))
```
Alternatives (not as [Pythonic](http://docs.python-guide.org/en/latest/writing/style/#idioms) and [slower](https://waymoot.org/home/python_string/)):
```
cool_python_libs = pandas + ', ' + numpy + ', ' + requests
print('Some cool python libraries: {}'.format(cool_python_libs))
cool_python_libs = pandas
cool_python_libs += ', ' + numpy
cool_python_libs += ', ' + requests
print('Some cool python libraries: {}'.format(cool_python_libs))
```
## `str.upper(), str.lower(), str.title()`
```
mixed_case = 'PyTHoN hackER'
mixed_case.upper()
mixed_case.lower()
mixed_case.title()
```
## `str.strip()`
```
ugly_formatted = ' \n \t Some story to tell '
stripped = ugly_formatted.strip()
print('ugly: {}'.format(ugly_formatted))
print('stripped: {}'.format(ugly_formatted.strip()))
```
## `str.split()`
```
sentence = 'three different words'
words = sentence.split()
print(words)
type(words)
secret_binary_data = '01001,101101,11100000'
binaries = secret_binary_data.split(',')
print(binaries)
```
## Calling multiple methods in a row
```
ugly_mixed_case = ' ThIS LooKs BAd '
pretty = ugly_mixed_case.strip().lower().replace('bad', 'good')
print(pretty)
```
Note that execution order is from left to right. Thus, this won't work:
```
pretty = ugly_mixed_case.replace('bad', 'good').strip().lower()
print(pretty)
```
## [Escape characters](http://python-reference.readthedocs.io/en/latest/docs/str/escapes.html#escape-characters)
```
two_lines = 'First line\nSecond line'
print(two_lines)
indented = '\tThis will be indented'
print(indented)
```
| github_jupyter |
# Evaluation
To be able to make a statement about the performance of a question-answering system, it is important to evalute it. Furthermore, evaluation allows to determine which parts of the system can be improved.
## Start an Elasticsearch server
You can start Elasticsearch on your local machine instance using Docker. If Docker is not readily available in your environment (eg., in Colab notebooks), then you can manually download and execute Elasticsearch from source.
```
# Install the latest release of Haystack in your own environment
#! pip install farm-haystack
# Install the latest master of Haystack and install the version of torch that works with the colab GPUs
!pip install git+https://github.com/deepset-ai/haystack.git
!pip install torch==1.6.0+cu101 -f https://download.pytorch.org/whl/torch_stable.html
# In Colab / No Docker environments: Start Elasticsearch from source
! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.6.2-linux-x86_64.tar.gz -q
! tar -xzf elasticsearch-7.6.2-linux-x86_64.tar.gz
! chown -R daemon:daemon elasticsearch-7.6.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(['elasticsearch-7.6.2/bin/elasticsearch'],
stdout=PIPE, stderr=STDOUT,
preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
! sleep 30
from farm.utils import initialize_device_settings
device, n_gpu = initialize_device_settings(use_cuda=True)
from haystack.preprocessor.utils import fetch_archive_from_http
# Download evaluation data, which is a subset of Natural Questions development set containing 50 documents
doc_dir = "../data/nq"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/nq_dev_subset_v2.json.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# make sure these indices do not collide with existing ones, the indices will be wiped clean before data is inserted
doc_index = "tutorial5_docs"
label_index = "tutorial5_labels"
# Connect to Elasticsearch
from haystack.document_store.elasticsearch import ElasticsearchDocumentStore
# Connect to Elasticsearch
document_store = ElasticsearchDocumentStore(host="localhost", username="", password="", index="document",
create_index=False, embedding_field="emb",
embedding_dim=768, excluded_meta_data=["emb"])
# Add evaluation data to Elasticsearch Document Store
# We first delete the custom tutorial indices to not have duplicate elements
document_store.delete_all_documents(index=doc_index)
document_store.delete_all_documents(index=label_index)
document_store.add_eval_data(filename="../data/nq/nq_dev_subset_v2.json", doc_index=doc_index, label_index=label_index)
```
## Initialize components of QA-System
```
# Initialize Retriever
from haystack.retriever.sparse import ElasticsearchRetriever
retriever = ElasticsearchRetriever(document_store=document_store)
# Alternative: Evaluate DensePassageRetriever
# Note, that DPR works best when you index short passages < 512 tokens as only those tokens will be used for the embedding.
# Here, for nq_dev_subset_v2.json we have avg. num of tokens = 5220(!).
# DPR still outperforms Elastic's BM25 by a small margin here.
# from haystack.retriever.dense import DensePassageRetriever
# retriever = DensePassageRetriever(document_store=document_store,
# query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
# passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
# use_gpu=True,
# embed_title=True,
# max_seq_len=256,
# batch_size=16,
# remove_sep_tok_from_untitled_passages=True)
#document_store.update_embeddings(retriever, index=doc_index)
# Initialize Reader
from haystack.reader.farm import FARMReader
reader = FARMReader("deepset/roberta-base-squad2", top_k_per_candidate=4)
# Initialize Finder which sticks together Reader and Retriever
from haystack.finder import Finder
finder = Finder(reader, retriever)
```
## Evaluation of Retriever
```
## Evaluate Retriever on its own
retriever_eval_results = retriever.eval(top_k=20, label_index=label_index, doc_index=doc_index)
## Retriever Recall is the proportion of questions for which the correct document containing the answer is
## among the correct documents
print("Retriever Recall:", retriever_eval_results["recall"])
## Retriever Mean Avg Precision rewards retrievers that give relevant documents a higher rank
print("Retriever Mean Avg Precision:", retriever_eval_results["map"])
```
## Evaluation of Reader
```
# Evaluate Reader on its own
reader_eval_results = reader.eval(document_store=document_store, device=device, label_index=label_index, doc_index=doc_index)
# Evaluation of Reader can also be done directly on a SQuAD-formatted file without passing the data to Elasticsearch
#reader_eval_results = reader.eval_on_file("../data/nq", "nq_dev_subset_v2.json", device=device)
## Reader Top-N-Accuracy is the proportion of predicted answers that match with their corresponding correct answer
print("Reader Top-N-Accuracy:", reader_eval_results["top_n_accuracy"])
## Reader Exact Match is the proportion of questions where the predicted answer is exactly the same as the correct answer
print("Reader Exact Match:", reader_eval_results["EM"])
## Reader F1-Score is the average overlap between the predicted answers and the correct answers
print("Reader F1-Score:", reader_eval_results["f1"])
```
## Evaluation of Finder
```
# Evaluate combination of Reader and Retriever through Finder
# Evaluate combination of Reader and Retriever through Finder
finder_eval_results = finder.eval(top_k_retriever=1, top_k_reader=10, label_index=label_index, doc_index=doc_index)
finder.print_eval_results(finder_eval_results)
```
| github_jupyter |
# Training Job with Encrypted Static Assets
In the [notebook about creating a training job in VPC mode](https://github.com/aws/amazon-sagemaker-examples/blob/master/sagemaker-fundamentals/create-training-job/create_training_job_vpc.ipynb) you learnt how to create a SageMaker training job with network isolation. Network isolation enables you to protect your data and model from being intercepted by cyber pirates.
Another way you can protect your static assets is to encrypt them before moving them from location A to location B. In this notebook, you will walk through a few techniques on that with the help of AWS Key Management Service [(AWS KMS)](https://docs.aws.amazon.com/kms/latest/developerguide/overview.html).
The following materials are helpful to get you started if you are not familiar with cryptography:
* [AWS KMS Crytography Details](https://docs.aws.amazon.com/kms/latest/cryptographic-details/intro.html)
* [Wikipedia page](https://en.wikipedia.org/wiki/Encryption)
* [Chapter 2 of GNU Privacy Handbook](https://www.gnupg.org/gph/en/manual.html)
You are strongly encouraged to go through the [overview](https://docs.aws.amazon.com/kms/latest/developerguide/overview.html), [concepts](https://docs.aws.amazon.com/kms/latest/developerguide/concepts.html) and [get started](https://docs.aws.amazon.com/kms/latest/developerguide/getting-started.html) sections from the KMS documentations before going through this notebook. This will help you getting familiar with some terminologies we will be using later.
Encryption is a wildly used technology, in addition to the above introductory material, you can find many free lectures online.
## Symmetric Ciphers
We will focus on symmetric ciphers in this notebook. Quote from the GNU Privacy Handbook
> A symmetric cipher is a cipher that uses the same key for both encryption and decryption. Two parties communicating using a symmetric cipher must agree on the key beforehand. Once they agree, the sender encrypts a message using the key, sends it to the receiver, and the receiver decrypts the message using the key. As an example, the German Enigma is a symmetric cipher, and daily keys were distributed as code books. Each day, a sending or receiving radio operator would consult his copy of the code book to find the day's key. Radio traffic for that day was then encrypted and decrypted using the day's key.
## Environment to run this notebook
You can run this notebook on your local machine or EC2 instance as an IAM user or you can run it on SageMaker Notebook Instance as a SageMaker service role. To avoid confusion, we will assume you are running it as an IAM user.
## Permissions
You will need to attach the following permissions to the IAM user
* IAMFullAccess
* AWSKeyManagementServicePowerUser
* AmazonEC2ContainerRegistryFullAccess
## Outline of this notebook
* Generate a symmetric KMS key
* Allow your SageMaker service role to use the KMS key
* Generate a data key from the KMS key
* Encrypt some data with the data key and upload the encrypted data to S3
* Create a SageMaker service role
* Build a training image
* Create a SageMaker training job using the encrypted data
* Verify that data retrieved from S3 is encrypted and SageMaker needs your data key to decrypt
The process of using a data key to encrypt your data instead of using the KMS key directly is called [**envelope encryption**](https://docs.aws.amazon.com/kms/latest/developerguide/concepts.html#enveloping).
Encrypting your data at rest using KMS ensures that only users who have access to call `KMS:Decrypt` on our CMK will be able to view the data, even if they are able to access the data in the s3 bucket or on the disk of a computer
We will discuss the use of data key in detail later. This reduces the changes of many attacks such as man-in-the-middle.

```
# set ups
import boto3
import datetime
import json
import pprint
pp = pprint.PrettyPrinter(indent=1)
kms = boto3.client("kms")
# Some helper functions
def current_time():
ct = datetime.datetime.now()
return str(ct.now()).replace(":", "-").replace(" ", "-")[:19]
def account_id():
return boto3.client("sts").get_caller_identity()["Account"]
```
### Generate a symmetric KMS key
You will use [kms:CreateKey](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/kms.html#KMS.Client.create_key) API to generate a **symmetric key** used for **encryption** and **decryption**. You need to use a IAM policy to define who has access (and with what level of access) to the key.
If you create the key from AWS console, then by following the default steps you will end up the following key policy:
```
root_arn = f"arn:aws:iam::{account_id()}:root"
user_arn = boto3.client("sts").get_caller_identity()["Arn"]
key_policy = {
"Id": "key-consolepolicy-3",
"Version": "2012-10-17",
"Statement": [
{
"Sid": "Enable IAM User Permissions",
"Effect": "Allow",
"Principal": {"AWS": root_arn}, # enable root user to perform all actions
"Action": "kms:*",
"Resource": "*",
},
{
"Sid": "Allow access for Key Administrators",
"Effect": "Allow",
"Principal": {
"AWS": [user_arn] # give myself admin permission to this key
# you can add more admin users by appending this list
},
"Action": [
"kms:Create*",
"kms:Describe*",
"kms:Enable*",
"kms:List*",
"kms:Put*",
"kms:Update*",
"kms:Revoke*",
"kms:Disable*",
"kms:Get*",
"kms:Delete*",
"kms:TagResource",
"kms:UntagResource",
"kms:ScheduleKeyDeletion",
"kms:CancelKeyDeletion",
],
"Resource": "*",
},
{
"Sid": "Allow use of the key",
"Effect": "Allow",
"Principal": {
"AWS": [user_arn] # allow myself to use the key
# you can add more users / roles to this list
# for example you can add SageMaker service role
# here. But we will allow SageMaker service role
# to use this key via grant (see below)
},
"Action": [
"kms:Encrypt",
"kms:Decrypt",
"kms:ReEncrypt*",
"kms:GenerateDataKey*",
"kms:DescribeKey",
],
"Resource": "*",
},
{
"Sid": "Allow attachment of persistent resources",
"Effect": "Allow",
"Principal": {
"AWS": [user_arn] # allow myself to create grant for this key
# see ref below to understand the diff
# between user and grant
# https://docs.aws.amazon.com/kms/latest/developerguide/concepts.html#grant
},
"Action": ["kms:CreateGrant", "kms:ListGrants", "kms:RevokeGrant"],
"Resource": "*",
"Condition": {"Bool": {"kms:GrantIsForAWSResource": "true"}},
},
],
}
key_policy = json.dumps(key_policy)
```
You can either create a new key or use an existing key. If you want to use an existing key, set `create_new_key` variable to `False` and replace `None` in line 23 by your key id. Note that in order to run this notebook, the key policy of your existing key should grant you AT LEAST the same level of access as the above key policy.
```
# create a key with the above key policy
create_new_key = False
if create_new_key:
ck_res = kms.create_key(
Policy=key_policy,
Description="a symmetric key to demonstrate KMS",
KeyUsage="ENCRYPT_DECRYPT", # use this key to encrypt and decrypt
Origin="AWS_KMS", # created via AWS KMS
CustomerMasterKeySpec="SYMMETRIC_DEFAULT", # symmetric key
)
pp.pprint(ck_res)
kms_key = kms_key ## replace `kms_key` here with ck_res["KeyMetadata"]["KeyId"]
print("The id of the key: ")
print(kms_key)
else:
print("Supply an existing KMS key by setting kms_key variable to your key id")
# replace None by your CMK key id
kms_key = None
if kms_key is None:
raise ValueError("Supply a valid KMS key id or create a new one")
kms_key = ck_res["KeyMetadata"]["KeyId"]
print("The id of the key: ")
```
You can use this KMS key to encrypt your data directly. It is not a good practice in production. But it is good to know what you can do.
```
my_secret_message = "1729 is the smallest number expressible \
as the sum of two cubes in two different ways".encode(
"utf-8"
)
# 1729 = 1^3 + 12^3 = 9^3 + 10^3 (Srinivasa Ramanujan)
# make the above secret a ciphertext
enc_res = kms.encrypt(KeyId=kms_key, Plaintext=my_secret_message)
pp.pprint(enc_res)
# decrypt your secret message
dec_res = kms.decrypt(KeyId=kms_key, CiphertextBlob=enc_res["CiphertextBlob"])
print("Decrpyted message:")
print(dec_res["Plaintext"].decode())
```
One thing to notice is encryption and decryption should happen at **bytes** level. If you want to encrypt a python object (list, numpy array, pandas data frame, pytorch model or a string) then the first step is to serialize it into bytes. One easy way to do it is to use `pickle.dumps` method.
## Client-side encryption with data key
Now let's pretend you are a data engineer and you need to move a chuck of data from location A to location B. Location A is the machine you are using now to run this notebook, location B is an S3 bucket that your data scientist buddy will be using later to create a training job. You want to ensure that while data is on its way from location A to location B, it is not intercepted and stolen by cyber-attacker in the middle.
One solution is genereate a data key `DK` from the KMS key and use `DK` to encrypt your data at location A (client side) and save the encrypted to S3 bucket.
You will get a different data key each time you request it from the KMS key and the plaintext data key is intended to be **short-lived** and you should only save the **encrypted** data key for later use.
Use [kms:GenerateDataKey](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/kms.html#KMS.Client.generate_data_key) to generate a data key.
```
key_length = 32 # 32 bytes
data_key_res = kms.generate_data_key(
KeyId=kms_key,
NumberOfBytes=key_length # your data key is will be 32x8=256-bit long
# takes 2^256 number of guesses to crack your data key
)
pp.pprint(data_key_res)
plaintext, ciphertext = data_key_res["Plaintext"], data_key_res["CiphertextBlob"]
assert len(plaintext) == key_length
```
The ciphertext above is the encrypted data key. Of course it is encrypted by the KMS key. And the ciphtertext is what you should keep for long term. There is nothing preventing you from encrypting your plaintext data key with a different KMS key. You just need to remember which KMS you used to encrypt it.
```
assert kms.decrypt(KeyId=kms_key, CiphertextBlob=ciphertext)["Plaintext"] == plaintext
```
Note that the plaintext data key is a byte-like object. It is not a string and in fact it cannot be decoded to a python string.
```
try:
plaintext.decode("utf-8")
except Exception as e:
print(e)
```
We will be using the AWS Encryption SDK library for client-side encryption. For more information on the AWS Encryption library, see [AWS Encryption SDK library](https://docs.aws.amazon.com/encryption-sdk/latest/developer-guide/introduction.html).
If you do not have this library, then you will need to install it as follows:
```
pip install -q 'aws-encryption-sdk'
import aws_encryption_sdk
from aws_encryption_sdk.identifiers import CommitmentPolicy
client = aws_encryption_sdk.EncryptionSDKClient(
commitment_policy=CommitmentPolicy.REQUIRE_ENCRYPT_REQUIRE_DECRYPT
)
kms_key_provider = aws_encryption_sdk.StrictAwsKmsMasterKeyProvider(
key_ids=[kms_key] ## Add your KMS key here
)
def encrypt(data, kms_key_provider):
"""Encrypt a chunk of bytes on client-side
data: a chunk of bytes
kms_key_provider: kms key
"""
ciphertext, encryptor_header = client.encrypt(
source=my_plaintext,
key_provider=kms_key_provider
)
return ciphertext
def decrypt(data, kms_key_provider):
"""Encrypt a chunk of bytes on client-side
data: a chunk of bytes that is encrypted.
kms_key_provider: kms key
"""
decrypted_plaintext, decryptor_header = client.decrypt(
source=data,
key_provider=kms_key_provider
)
return decrypted_plaintext
import pickle
# encrypt
data = [i for i in range(1729)]
encrypted_data = encrypt(pickle.dumps(data), plaintext) # python object -> bytes
```
Once you finished encryption, you should delete the plaintext data key as soon as possible.
```
del plaintext
# decrypt
b = decrypt(encrypted_data, ciphertext)
data_ = pickle.loads(b) # bytes -> python object
for x, y in zip(data, data_):
assert x == y
```
## Save encrypted objects on S3
Now you understand how encryption at client-side works. It should be straightforward to you how to save encrypted data on an S3 bucket.
```
# create a bucket to be shared by SageMaker later
def create_bucket():
"""Create an S3 bucket that is intended to be used for short term"""
bucket = f"sagemaker-{current_time()}"
region_name = boto3.Session().region_name
create_bucket_config = {}
if region_name != "us-east-1":
# us-east-1 is the default region for S3 bucket
# specify LocationConstraint if your VPC is not
# in us-east-1
create_bucket_config["LocationConstraint"] = region_name
boto3.client("s3").create_bucket(Bucket=bucket, CreateBucketConfiguration=create_bucket_config)
return bucket
bucket = create_bucket()
# put your encrypted data on the S3 bucket
s3 = boto3.client("s3")
input_prefix = "data" # will be used later as S3Prefix when calling CreateTrainingJob
put_obj_res = s3.put_object(
Bucket=bucket, Key=input_prefix + "/" + "a_chunk_of_secrets", Body=encrypted_data
)
pp.pprint(put_obj_res)
```
## Create a SageMaker training job with encrypted data
Now you understand how to move your data from location $A$ to location $B$ encrypted. Let's see how this workflow can be merged into a SageMaker training job. What you want to achieve is, the static assets (model and data) need to be encrypted before you traffic them in the Internet.
Let $M$ denote the customer KMS key hosted on KMS, $D$ the plaintext data key and $C$ the ciphertext data key.
Suppose your training data is in an S3 bucket encrypted by the data key $D$. In order to use the training data, the SageMaker training job needs to be able to decrypt it. Of course you **would not** want to move $D$ (plaintext) around in the Internet and hand it to a SageMaker training job. Instead you will hand the encrypted data key (ciphertext) $C$ to the SageMaker training job.
The SageMaker training job will do the following things with $C$
- Decrypt it to plaintext using the KMS key $M$ and get $D$
- Download the encrypted data from the S3 bucket and decrypt the data with $D$
- Train the model and encrypt the model with $D$
- Send the encrypted model to an S3 bucket
Of course, you could use a different data key to encrypt the model.
### How SageMaker uses your KMS key $M$
Remember a managed service like SageMaker *assumes* an IAM role (service role) in your account and it procures the resources in your AWS account based on the permission of the service role.
When you created $M$, key policy said that the IAM user (you) and the root user of your account are the only entities entitled to use $M$. So does SageMaker use $M$ then?
There are two ways to achieve this:
Suppose your SageMaker service role is called `example-role`.
1. Update the key policy to allow `example-role` to use $M$
2. Allow `example-role` to use $M$ via a **grant**
Quote from the [KMS docs](https://docs.aws.amazon.com/kms/latest/developerguide/concepts.html#grant)
>A grant is a policy instrument that allows AWS principals to use AWS KMS key in cryptographic operations. It also can let them view a CMK (DescribeKey) and create and manage grants. When authorizing access to a CMK, grants are considered along with key policies and IAM policies. Grants are often used for temporary permissions because you can create one, use its permissions, and delete it without changing your key policies or IAM policies. Because grants can be very specific, and are easy to create and revoke, they are often used to provide temporary permissions or more granular permissions.
We will the grant approach this tutorial as it involves less activities on your key policy. In a prodcution environment, you should think of an activity on your key policy as *a big deal*.
First, get some helper functions for creating a SageMaker service role.
```
%%bash
cp ../execution-role/iam_helpers.py .
# set up service role for SageMaker
from iam_helpers import create_execution_role
iam = boto3.client("iam")
role_name = "example-role"
role_arn = create_execution_role(role_name=role_name)["Role"]["Arn"]
iam.attach_role_policy(
RoleName=role_name, PolicyArn="arn:aws:iam::aws:policy/AmazonSageMakerFullAccess"
)
```
Now, you will verify that `example-role` cannot use your master key $M$ at this point. The cell below is expected to raise an exception.
```
# create a boto3 session with example-role
import time
print(role_arn)
def create_session(role_arn):
"""Create a boto3 session with an IAM role"""
now = str(time.time()).split(".")[0]
obj = boto3.client("sts").assume_role(RoleArn=role_arn, RoleSessionName=now)
cred = obj["Credentials"]
sess = boto3.session.Session(
aws_access_key_id=cred["AccessKeyId"],
aws_secret_access_key=cred["SecretAccessKey"],
aws_session_token=cred["SessionToken"],
)
return sess
sess = create_session(role_arn)
try:
sess.client("kms").encrypt(KeyId=kms_key, Plaintext="it will not go through".encode("utf-8"))
except Exception as e:
print(e)
del sess
grant_res = kms.create_grant(
KeyId=kms_key,
GranteePrincipal=role_arn,
Operations=["Decrypt", "Encrypt"], # allow example-role to use M to encrypt and decrypt
)
pp.pprint(grant_res)
# Verify example-role has can use M
sess = create_session(role_arn)
enc_res = sess.client("kms").encrypt(
KeyId=kms_key, Plaintext="it will go through this time".encode("utf-8")
)
pp.pprint(enc_res)
del sess
# put C to the bucket
s3.put_object(Bucket=bucket, Key="dont_look", Body=ciphertext)
```
### Build a training container
You will build a training image here like in [the notebook on basics of `CreateTrainingJob`](https://github.com/hsl89/amazon-sagemaker-examples/blob/sagemaker-fundamentals/sagemaker-fundamentals/create-training-job/create_training_job.ipynb)
```
# View the Dockerfile
!cat container_kms/Dockerfile
# View the entrypoint script
!pygmentize container_kms/train.py
```
You will need to build your AWS credentials into the container, because you will need to decrypt your ciphertext data key within the container.
```
cred = boto3.Session().get_credentials()
access_key, secret_key = cred.access_key, cred.secret_key
region_name = boto3.Session().region_name
%%bash -s "$access_key" "$secret_key" "$region_name"
# build the image
cd container_kms/
# tag it as example-image:latest
docker build -t example-image:latest . --build-arg ACCESS_KEY=$1 \
--build-arg SECRET_KEY=$2 --build-arg REGION_NAME=$3
```
## Test your container locally
You programmed the entrypoint `container_kms/train.py` so that it gets to know the id of the master key as well as the S3 object key for the data key ciphertext via hyperparameters in `/opt/ml/input/config/hyperparameters.json`. That means you will need to [inject those hyperparamters to the container](https://github.com/aws/amazon-sagemaker-examples/blob/master/sagemaker-fundamentals/create-training-job/create_training_job_hyperparameter_injection.ipynb)
You can checkout the [notebook on basics of create a training job](https://github.com/aws/amazon-sagemaker-examples/blob/master/sagemaker-fundamentals/create-training-job/create_training_job.ipynb)(section Test your container) for more details.
To recap, you will mount `container_kms/local_test/ml` (OS) to `/opt/ml`(container) as a docker volume and exchange training information with the container there.
Look at what hyperparameters we used in `container_kms/train.py`. The hyperparameters are:
```
hyperparameters = {
"ciphertext_s3_key": "dont_look",
"kms_key_id": kms_key,
"train_channel": "train",
"train_file": "a_chunk_of_secrets",
"key_bucket": bucket,
}
pp.pprint(hyperparameters)
```
The hyperparameters are made available to the training container at `/opt/ml/input/config/hyperparameter.json`, so you will write the hyperparameters to `container_kms/local_test/ml/input/config/hyperparameters.json` for local testing.
```
import json
with open("container_kms/local_test/ml/input/config/hyperparameters.json", "w") as f:
json.dump(hyperparameters, f)
!cat container_kms/local_test/ml/input/config/hyperparameters.json
```
Also, you need to have `container_kms/local_test/ml/input/train/a_chunk_of_secrets` available
```
import os
with open(
os.path.join(
"container_kms", "local_test", "ml", "input", "data", "train", "a_chunk_of_secrets"
),
"wb",
) as f:
f.write(encrypted_data)
!ls -R container_kms/local_test/ml
!python container_kms/local_test/test_container.py
# create a repo in ECR called example-image
ecr = boto3.client("ecr")
try:
# The repository might already exist
# in your ECR
cr_res = ecr.create_repository(repositoryName="example-image")
pp.pprint(cr_res)
except Exception as e:
print(e)
%%bash
account=$(aws sts get-caller-identity --query Account | sed -e 's/^"//' -e 's/"$//')
region=$(aws configure get region)
ecr_account=${account}.dkr.ecr.${region}.amazonaws.com
# Give docker your ECR login password
aws ecr get-login-password --region $region | docker login --username AWS --password-stdin $ecr_account
# Fullname of the repo
fullname=$ecr_account/example-image:latest
#echo $fullname
# Tag the image with the fullname
docker tag example-image:latest $fullname
# Push to ECR
docker push $fullname
```
Now you have all the ingredients for a SageMaker training job.
```
# configure a training job
sm_cli = boto3.client("sagemaker")
# input
data_path = "s3://" + bucket + "/" + input_prefix
# location that SageMaker saves the model artifacts
output_prefix = "output"
output_path = "s3://" + bucket + "/" + output_prefix
# ECR URI of your image
region = boto3.Session().region_name
account = account_id()
image_uri = "{}.dkr.ecr.{}.amazonaws.com/example-image:latest".format(account, region)
algorithm_specification = {
"TrainingImage": image_uri,
"TrainingInputMode": "File",
}
input_data_config = [
{
"ChannelName": "train",
"DataSource": {
"S3DataSource": {
"S3DataType": "S3Prefix",
"S3Uri": data_path,
"S3DataDistributionType": "FullyReplicated",
}
},
}
]
output_data_config = {"S3OutputPath": output_path}
resource_config = {"InstanceType": "ml.m5.large", "InstanceCount": 1, "VolumeSizeInGB": 10}
stopping_condition = {
"MaxRuntimeInSeconds": 120,
}
enable_network_isolation = False
# some helper functions to monitor the training job
import time
def monitor_training_job_status(training_job_name, log_freq=30):
"""Print out training job status every $log_freq seconds"""
stopped = False
while not stopped:
tj_state = sm_cli.describe_training_job(TrainingJobName=training_job_name)
if tj_state["TrainingJobStatus"] in ["Completed", "Stopped", "Failed"]:
stopped = True
else:
print("Training in progress")
time.sleep(log_freq)
if tj_state["TrainingJobStatus"] == "Failed":
print("Training job failed ")
print("Failed Reason: {}".format(tj_state["FailureReason"]))
else:
print("Training job completed")
return
def print_logs(training_job_name):
"""Print out stdout in the container from CloudWatch"""
logs = boto3.client("logs")
log_res = logs.describe_log_streams(
logGroupName="/aws/sagemaker/TrainingJobs", logStreamNamePrefix=training_job_name
)
for log_stream in log_res["logStreams"]:
# get one log event
log_event = logs.get_log_events(
logGroupName="/aws/sagemaker/TrainingJobs", logStreamName=log_stream["logStreamName"]
)
# print out messages from the log event
for ev in log_event["events"]:
for k, v in ev.items():
if k == "message":
print(v)
return
# name training job
training_job_name = "example-training-job-{}".format(current_time())
ct_res = sm_cli.create_training_job(
TrainingJobName=training_job_name,
AlgorithmSpecification=algorithm_specification,
RoleArn=role_arn,
HyperParameters=hyperparameters, # use the same hyperparameters for local testing
InputDataConfig=input_data_config,
OutputDataConfig=output_data_config,
ResourceConfig=resource_config,
StoppingCondition=stopping_condition,
EnableNetworkIsolation=enable_network_isolation,
EnableManagedSpotTraining=False,
)
monitor_training_job_status(training_job_name)
print_logs(training_job_name)
```
## Review && Discussion
In this notebook, you went through a typical workflow for creating a SageMaker training job with client-side encryption. You have
* Generated a data key from a KMS key
* Used the data key to encrypt your dataset before putting it to a SageMaker-accessible S3 bucket
* Created a SageMaker training job and passed the encrypted data key to the training job
* Decrypted the data key within the training container and used the decrypted data key to decrypted training data from the S3 bucket
* Encrypted your trained model at the end of the training job
You might have a few questions lingering at this point:
* Is pickle my best option to serialize and deserialize python objects?
* A: Pickle might not be your best option in your production env (thanks to @seebees). Checkout the following content:
* https://nedbatchelder.com/blog/202006/pickles_nine_flaws.html
* https://lwn.net/Articles/595352/
* https://intoli.com/blog/dangerous-pickles/
* Client-side encryption seems complicated, can I have a push-button solution?
* A: There are many technical details to take care of when doing client-side encryption. The purpose of this tutorial is to show you the basic concepts involved in client-side encryption. [AWS Encryption SDK](https://docs.aws.amazon.com/encryption-sdk/latest/developer-guide/introduction.html) is a library for client-side encryption and it can take care much of nuts and bolts for you. You are highly encouraged to explore it.
* I do not like to build my AWS credentials into an image.
* A: A much better way to access your credentials when running a container service like SageMaker training job is to use [`ContainerProvider`](https://github.com/boto/botocore/blob/develop/botocore/credentials.py#L1819). This class can be used for a wider range of problems, so we will discuss it in an standalone notebook.
## Clean up
You cannot delete the KMS key with one stroke. In this tutorial we created key for sake of running this tutorial, but in a production environment, deleting a key is a BIG deal, because once you deleled the KMS key, all data encryted under that key becomes unavailable to you. That's why you need to exercise extreme caution when deleting a key. Checkout the [section on deleting a KMS key](https://docs.aws.amazon.com/kms/latest/developerguide/deleting-keys.html) for more detail.
You can schedule a key deletion in $X$ days by calling [kms:ScheduleKeyDeletion](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/kms.html#KMS.Client.schedule_key_deletion).
Once this API is called, you key status will be **Pending Deletion**. This is to remind you that you need to sort out all your data encrypted under this key in $X$ days and encrypt them using a different key.
During those $X$ days, if you changed your mind and decided not to delete the key, you can call [kms:CancelKeyDeletion](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/kms.html#KMS.Client.cancel_key_deletion) API.
```
def schedule_key_deletion(key_id, waiting_period):
"""Delete a key in $waiting_period days
Args:
key_id: id of the key to be deleted
waiting_period: number of days to wait before key deletion
"""
dk_res = kms.schedule_key_deletion(KeyId=key_id, PendingWindowInDays=waiting_period)
pp.pprint(dk_res)
return
# call schedule_key_deletion if you want to delete the key
# schedule_key_deletion(key_id=kms_key, waiting_period=7)
def delete_force(bucket_name):
"""Helper function to delete a bucket"""
objs = s3.list_objects_v2(Bucket=bucket_name)["Contents"]
for obj in objs:
s3.delete_object(Bucket=bucket_name, Key=obj["Key"])
return s3.delete_bucket(Bucket=bucket_name)
def delete_ecr_repo(repo_name):
"""Helper function to delete an ECR repo"""
ecr.delete_repository(repositoryName=repo_name, force=True)
return
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/ImageCollection/overview.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/ImageCollection/overview.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://mybinder.org/v2/gh/giswqs/earthengine-py-notebooks/master?filepath=ImageCollection/overview.ipynb"><img width=58px src="https://mybinder.org/static/images/logo_social.png" />Run in binder</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/ImageCollection/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geehydro](https://github.com/giswqs/geehydro). The **geehydro** Python package builds on the [folium](https://github.com/python-visualization/folium) package and implements several methods for displaying Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, `Map.centerObject()`, and `Map.setOptions()`.
The following script checks if the geehydro package has been installed. If not, it will install geehydro, which automatically install its dependencies, including earthengine-api and folium.
```
import subprocess
try:
import geehydro
except ImportError:
print('geehydro package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geehydro'])
```
Import libraries
```
import ee
import folium
import geehydro
```
Authenticate and initialize Earth Engine API. You only need to authenticate the Earth Engine API once.
```
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
This step creates an interactive map using [folium](https://github.com/python-visualization/folium). The default basemap is the OpenStreetMap. Additional basemaps can be added using the `Map.setOptions()` function.
The optional basemaps can be `ROADMAP`, `SATELLITE`, `HYBRID`, `TERRAIN`, or `ESRI`.
```
Map = folium.Map(location=[40, -100], zoom_start=4)
Map.setOptions('HYBRID')
```
## Add Earth Engine Python script
```
# Create arbitrary constant images.
constant1 = ee.Image(1)
constant2 = ee.Image(2)
# Create a collection by giving a list to the constructor.
collectionFromConstructor = ee.ImageCollection([constant1, constant2])
print('collectionFromConstructor: ', collectionFromConstructor.getInfo())
# Create a collection with fromImages().
collectionFromImages = ee.ImageCollection.fromImages(
[ee.Image(3), ee.Image(4)])
print('collectionFromImages: ', collectionFromImages.getInfo())
# Merge two collections.
mergedCollection = collectionFromConstructor.merge(collectionFromImages)
print('mergedCollection: ', mergedCollection.getInfo())
# # Create a toy FeatureCollection
# features = ee.FeatureCollection(
# [ee.Feature({}, {'foo': 1}), ee.Feature({}, {'foo': 2})])
# # Create an ImageCollection from the FeatureCollection
# # by mapping a function over the FeatureCollection.
# images = features.map(function(feature) {
# return ee.Image(ee.Number(feature.get('foo')))
# })
# # Print the resultant collection.
# print('Image collection: ', images)
```
## Display Earth Engine data layers
```
Map.setControlVisibility(layerControl=True, fullscreenControl=True, latLngPopup=True)
Map
```
| github_jupyter |
# Exercise 5.03: Logistic Regression - Multi Class Classifier
```
import struct
import numpy as np
import gzip
import urllib.request
import matplotlib.pyplot as plt
from array import array
from sklearn.linear_model import LogisticRegression
```
Load the MNIST data into memory
```
with gzip.open('../Datasets/train-images-idx3-ubyte.gz', 'rb') as f:
magic, size, rows, cols = struct.unpack(">IIII", f.read(16))
img = np.array(array("B", f.read())).reshape((size, rows, cols))
with gzip.open('../Datasets/train-labels-idx1-ubyte.gz', 'rb') as f:
magic, size = struct.unpack(">II", f.read(8))
labels = np.array(array("B", f.read()))
with gzip.open('../Datasets/t10k-images-idx3-ubyte.gz', 'rb') as f:
magic, size, rows, cols = struct.unpack(">IIII", f.read(16))
img_test = np.array(array("B", f.read())).reshape((size, rows, cols))
with gzip.open('../Datasets/t10k-labels-idx1-ubyte.gz', 'rb') as f:
magic, size = struct.unpack(">II", f.read(8))
labels_test = np.array(array("B", f.read()))
```
Visualise a sample of the data
```
for i in range(10):
plt.subplot(2, 5, i + 1)
plt.imshow(img[i], cmap='gray');
plt.title(f'{labels[i]}');
plt.axis('off')
```
## Construct a Logistic Model to Classify Digits 0 - 9
In this model as we are predicting classes 0 - 9 we will require images from all available data. However given the extremely large dataset we will need to sample only a small amount of the original MNIST set due to limited system requirements and anticipated training time. We will select 2000 samples at random:
```
np.random.seed(0) # Give consistent random numbers
selection = np.random.choice(len(img), 5000)
selected_images = img[selection]
selected_labels = labels[selection]
```
In order to provide the image information to the Logistic model we must first flatten the data out so that each image is 1 x 784 pixels in shape.
```
selected_images = selected_images.reshape((-1, rows * cols))
selected_images.shape
```
Try applying normalisation by uncommenting the lines the next cell. **What effect does it have on performance?**
```
#selected_images = selected_images / 255.0
#img_test = img_test / 255.0
```
Let's construct the model, use the sklearn LogisticRegression API and call the fit function.
```
model = LogisticRegression(solver='lbfgs', multi_class='multinomial', max_iter=500, tol=0.1)
model.fit(X=selected_images, y=selected_labels)
```
Determine the score against the training set
```
model.score(X=selected_images, y=selected_labels)
```
Display the first two predictions for the Logistic model against the training data
```
model.predict(selected_images)[:2]
plt.subplot(1, 2, 1)
plt.imshow(selected_images[0].reshape((28, 28)), cmap='gray');
plt.axis('off');
plt.subplot(1, 2, 2)
plt.imshow(selected_images[1].reshape((28, 28)), cmap='gray');
plt.axis('off');
```
Examine the corresponding predicted probabilities for the first two training samples
```
model.predict_proba(selected_images)[0]
```
Compare the performance against the test set
```
model.score(X=img_test.reshape((-1, rows * cols)), y=labels_test)
```
| github_jupyter |
Linear models assume that the independent variables are normally distributed. In this recipe, we will learn how to assess normal distributions of variables.
```
import pandas as pd
import numpy as np
# for plotting
import matplotlib.pyplot as plt
import seaborn as sns
# for the Q-Q plots
import scipy.stats as stats
# the dataset for the demo
from sklearn.datasets import load_boston
# load the the Boston House price data
# this is how we load the boston dataset from sklearn
boston_dataset = load_boston()
# create a dataframe with the independent variables
boston = pd.DataFrame(boston_dataset.data,
columns=boston_dataset.feature_names)
boston.head()
# this is the information about the boston house prince dataset
# get familiar with the variables before continuing with
# the notebook
# the aim is to predict the "Median value of the houses"
# MEDV column of this dataset
# and we have variables with characteristics about
# the homes and the neighborhoods
print(boston_dataset.DESCR)
# I will create a dataframe with the variable x that
# follows a normal distribution
# this will provide the expected plots
# i.e., how the plots should look like if the
# assumption is met
np.random.seed(29) # for reproducibility
n = 200 # in the book, we pass 200 within brackets directly, without defining n
x = np.random.randn(n)
data = pd.DataFrame([x]).T
data.columns = ['x']
data.head()
```
Normality can be assessed by histograms:
```
# histogram of the simulated independent variable x
# which we know follows a Gaussian distribution
sns.distplot(data['x'], bins=30)
# histogram of the variable RM from the boston
# house price dataset from sklearn
# RM is the average number of rooms per dwelling
sns.distplot(boston['RM'], bins=30)
# histogram of the variable LSTAT
# (% lower status of the population)
sns.distplot(boston['LSTAT'], bins=30)
```
Normality can be also assessed by Q-Q plots. In a Q-Q plot we plot the quantiles of the variable in the y-axis and the expected quantiles of the normal distribution in the x-axis. If the variable follows a normal distribution, the dots in the Q-Q plot should fall in a 45 degree diagonal line as indicated below.
```
# let's plot the Q-Q plot for the simualted data.
# the dots should adjust to the 45 degree line
stats.probplot(data['x'], dist="norm", plot=plt)
plt.show()
# let's do the same for RM
stats.probplot(boston['RM'], dist="norm", plot=plt)
plt.show()
```
Most of the observations of RM fall on the 45 degree line, which suggests that the distribution is approximately Gaussian, with some deviation towards the larger and smaller values of the variable.
```
# just for comparison, let's go ahead and plot CRIM
stats.probplot(boston['CRIM'], dist="norm", plot=plt)
plt.show()
```
CRIM does not follow a Gaussian distribution as most of its observations deviate from the 45 degree line in the Q-Q plot.
| github_jupyter |
### Reconstruction with a custom network.
This notebook extends the last notebook to simultaneously train a decoder network, which translates from embedding back into dataspace. It also shows you how to use validation data for the reconstruction network during training.
### load data
```
from tensorflow.keras.datasets import mnist
(train_images, Y_train), (test_images, Y_test) = mnist.load_data()
train_images = train_images.reshape((train_images.shape[0], -1))/255.
test_images = test_images.reshape((test_images.shape[0], -1))/255.
```
### define the encoder network
```
import tensorflow as tf
dims = (28,28, 1)
n_components = 2
encoder = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=dims),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation="relu", padding="same"
),
tf.keras.layers.Conv2D(
filters=128, kernel_size=3, strides=(2, 2), activation="relu", padding="same"
),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=n_components),
])
encoder.summary()
decoder = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=(n_components)),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=512, activation="relu"),
tf.keras.layers.Dense(units=7 * 7 * 128, activation="relu"),
tf.keras.layers.Reshape(target_shape=(7, 7, 128)),
tf.keras.layers.UpSampling2D((2)),
tf.keras.layers.Conv2D(
filters=64, kernel_size=3, padding="same", activation="relu"
),
tf.keras.layers.UpSampling2D((2)),
tf.keras.layers.Conv2D(
filters=32, kernel_size=3, padding="same", activation="relu"
),
])
decoder.summary()
```
### create parametric umap model
```
from umap.parametric_umap import ParametricUMAP
embedder = ParametricUMAP(
encoder=encoder,
decoder=decoder,
dims=dims,
n_components=n_components,
n_training_epochs=5,
parametric_reconstruction= True,
reconstruction_validation=test_images,
verbose=True,
)
embedding = embedder.fit_transform(train_images)
```
### plot reconstructions
```
test_images_recon = embedder.inverse_transform(embedder.transform(test_images))
import numpy as np
nex = 10
fig, axs = plt.subplots(ncols=10, nrows=2, figsize=(nex, 2))
for i in range(nex):
axs[0, i].matshow(np.squeeze(test_images[i].reshape(28, 28, 1)), cmap=plt.cm.Greys)
axs[1, i].matshow(
tf.nn.sigmoid(np.squeeze(test_images_recon[i].reshape(28, 28, 1))),
cmap=plt.cm.Greys,
)
for ax in axs.flatten():
ax.axis("off")
```
### plot results
```
embedding = embedder.embedding_
import matplotlib.pyplot as plt
fig, ax = plt.subplots( figsize=(8, 8))
sc = ax.scatter(
embedding[:, 0],
embedding[:, 1],
c=Y_train.astype(int),
cmap="tab10",
s=0.1,
alpha=0.5,
rasterized=True,
)
ax.axis('equal')
ax.set_title("UMAP in Tensorflow embedding", fontsize=20)
plt.colorbar(sc, ax=ax);
```
### plotting loss
```
embedder._history.keys()
fig, axs = plt.subplots(ncols=2, figsize=(10,5))
ax = axs[0]
ax.plot(embedder._history['loss'])
ax.set_ylabel('Cross Entropy')
ax.set_xlabel('Epoch')
ax = axs[1]
ax.plot(embedder._history['reconstruction_loss'], label='train')
ax.plot(embedder._history['val_reconstruction_loss'], label='valid')
ax.legend()
ax.set_ylabel('Cross Entropy')
ax.set_xlabel('Epoch')
```
| github_jupyter |
# Job Listings
```
# Dependencies & Setup
import pandas as pd
import numpy as np
import requests
import json
from os.path import exists
import simplejson as json
# Retrieve Google API Key from config.py
from config_3 import gkey
# File to Load
ba_file = "data/bay_area_job_listings.csv"
# Read Scraped Data (CSV File) & Store Into Pandas DataFrame
ba_job_listings_df = pd.read_csv(ba_file, encoding="ISO-8859-1")
# Drop BA NaN's
revised_ba_job_listings_df = ba_job_listings_df.dropna()
revised_ba_job_listings_df.head()
# Reorganize WC File Column Names
organized_wc_job_listings_df = cleaned_wc_job_listings_df.rename(columns={"company":"Company Name",
"job_title":"Job Title",
"location":"Location"})
# Extract Only Job Titles with "Data" as String
new_organized_wc_job_listings_df = organized_wc_job_listings_df[organized_wc_job_listings_df["Job Title"].
str.contains("Data", case=True)]
new_organized_wc_job_listings_df.head()
print(len(new_organized_wc_job_listings_df))
# Extract Unique Locations
new_organized_wc_job_listings_df["company_address"] = new_organized_wc_job_listings_df["Company Name"] + ", " + new_organized_wc_job_listings_df["Location"]
unique_locations = new_organized_wc_job_listings_df["company_address"].unique().tolist()
print(len(unique_locations))
# Reorganize BA File Column Names
organized_ba_job_listings_df = revised_ba_job_listings_df.rename(columns={"company":"Company Name",
"job_title":"Job Title",
"location":"Location"})
organized_ba_job_listings_df.head()
# Extract Only Company Names to Pass to Google Maps API to Gather GeoCoordinates
company = organized_ba_job_listings_df[["Company Name"]]
company.head()
# What are the GeoCoordinates (Latitude/Longitude) of the Companies?
company_list = list(company["Company Name"])
# Build URL using the Google Maps API
base_url = "https://maps.googleapis.com/maps/api/geocode/json"
new_json = []
for target_company in company_list:
params = {"address": target_company + ", San Francisco CA", "key": gkey}
# Run Request
response = requests.get(base_url, params=params)
# Extract lat/lng
companies_geo = response.json()
lat = companies_geo["results"][0]["geometry"]["location"]["lat"]
lng = companies_geo["results"][0]["geometry"]["location"]["lng"]
new_json.append({"company": target_company,"lat": lat,"lng": lng})
# print(f"{target_company}, {lat}, {lng}")
print(new_json)
# What are the GeoCoordinates (Latitude/Longitude) of the Companies?
company_list = list(company["Company Name"])
# Build URL using the Google Maps API
base_url = "https://maps.googleapis.com/maps/api/geocode/json"
new_json = []
counter = 1
for location in company_list:
params = {"address": location, "key": gkey}
# Run Request
response = requests.get(base_url, params=params)
try:
# Extract lat/lng
companies_geo = response.json()
# print(companies_geo)
lat = companies_geo["results"][0]["geometry"]["location"]["lat"]
lng = companies_geo["results"][0]["geometry"]["location"]["lng"]
new_json.append({"company": location,"lat": lat,"lng": lng})
print(counter)
counter += 1
except IndexError:
print(location)
# What are the geocoordinates (latitude/longitude) of the Company Names?
company_list = list(company["Company Name"])
# Build URL using the Google Maps API
base_url = "https://maps.googleapis.com/maps/api/geocode/json"
new_json = []
for target_company in company_list:
# print(target_company)
params = {"address": target_company + ", San Francisco CA", "key": gkey}
# print(params)
# print("The Geocoordinates of LinkedIn Company Names")
# Run Request
response = requests.get(base_url, params=params)
# print(response.url)
# Extract lat/lng
companies_geo = response.json()
lat = companies_geo["results"][0]["geometry"]["location"]["lat"]
lng = companies_geo["results"][0]["geometry"]["location"]["lng"]
new_json.append({"company":target_company,"lat":lat,"lng":lng})
# print(f"{target_company}, {lat}, {lng}")
print(new_json)
print(job_list_json)
# Convert JSON into GeoJSON
geojson = {
"type": "FeatureCollection",
"features": [
{
"type": "Feature",
"company": d["company"],
"geometry" : {
"type": "Point",
"coordinates": [d["lat"], d["lng"]],
},
} for d in job_list_json]
}
print(job_list_geojson)
job_listing_coordinates = pd.DataFrame(new_json)
job_listing_coordinates
updated_job_listings = job_listings.merge(job_listing_coordinates, how="left", left_on="company_address", right_on="company")
updated_job_listings
# Drop NaN's
updated_job_listings_no_missing = updated_job_listings.dropna()
updated_job_listings_no_missing.head()
updated_job_listings[["company","lat","lng"]].to_dict()
json_job_listings = updated_job_listings[["company","lat","lng"]].to_json(orient="records")
json_job_listings
with open('data.json', 'w') as outfile:
outfile.write(json_job_listings)
```
| github_jupyter |
# EDA
> Exploratory Data Analysis
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "end_to_end_project"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
# Ignore useless warnings (see SciPy issue #5998)
import warnings
warnings.filterwarnings(action="ignore", message="^internal gelsd")
```
### Preview of Dataset
Available at https://raw.githubusercontent.com/ageron/handson-ml2/master/datasets/housing/housing.tgz
```
import os
import tarfile
import urllib
DOWNLOAD_ROOT = "https://raw.githubusercontent.com/ageron/handson-ml2/master/"
HOUSING_PATH = os.path.join("datasets", "housing")
HOUSING_URL = DOWNLOAD_ROOT + "datasets/housing/housing.tgz"
def fetch_housing_data(housing_url=HOUSING_URL, housing_path=HOUSING_PATH):
if not os.path.isdir(housing_path):
os.makedirs(housing_path)
tgz_path = os.path.join(housing_path, "housing.tgz")
urllib.request.urlretrieve(housing_url, tgz_path)
housing_tgz = tarfile.open(tgz_path)
housing_tgz.extractall(path=housing_path)
housing_tgz.close()
fetch_housing_data()
import pandas as pd
def load_housing_data(housing_path=HOUSING_PATH):
csv_path = os.path.join(housing_path, "housing.csv")
return pd.read_csv(csv_path)
```
### Preview of First 10 Rows
```
housing = load_housing_data()
print(f'Full dataset has {housing.shape[0]} rows, {housing.shape[0]} columns')
housing.head(10)
from pandas_profiling import ProfileReport
profile = ProfileReport(housing, title='Pandas Profiling Report', minimal=True)
profile.to_file(output_file="your_report.html")
housing.describe()
housing["ocean_proximity"].value_counts()
%matplotlib inline
import matplotlib.pyplot as plt
housing.hist(bins=50, figsize=(20,15))
save_fig("attribute_histogram_plots")
plt.show()
# to make this notebook's output identical at every run
np.random.seed(42)
import numpy as np
# For illustration only. Sklearn has train_test_split()
def split_train_test(data, test_ratio):
shuffled_indices = np.random.permutation(len(data))
test_set_size = int(len(data) * test_ratio)
test_indices = shuffled_indices[:test_set_size]
train_indices = shuffled_indices[test_set_size:]
return data.iloc[train_indices], data.iloc[test_indices]
train_set, test_set = split_train_test(housing, 0.2)
from zlib import crc32
def test_set_check(identifier, test_ratio):
return crc32(np.int64(identifier)) & 0xffffffff < test_ratio * 2**32
def split_train_test_by_id(data, test_ratio, id_column):
ids = data[id_column]
in_test_set = ids.apply(lambda id_: test_set_check(id_, test_ratio))
return data.loc[~in_test_set], data.loc[in_test_set]
```
The implementation of `test_set_check()` above works fine in both Python 2 and Python 3. In earlier releases, the following implementation was proposed, which supported any hash function, but was much slower and did not support Python 2:
```
import hashlib
def test_set_check(identifier, test_ratio, hash=hashlib.md5):
return hash(np.int64(identifier)).digest()[-1] < 256 * test_ratio
```
If you want an implementation that supports any hash function and is compatible with both Python 2 and Python 3, here is one:
```
def test_set_check(identifier, test_ratio, hash=hashlib.md5):
return bytearray(hash(np.int64(identifier)).digest())[-1] < 256 * test_ratio
housing_with_id = housing.reset_index() # adds an `index` column
train_set, test_set = split_train_test_by_id(housing_with_id, 0.2, "index")
housing_with_id["id"] = housing["longitude"] * 1000 + housing["latitude"]
train_set, test_set = split_train_test_by_id(housing_with_id, 0.2, "id")
```
### Median Income
```
housing["median_income"].hist()
housing["income_cat"] = pd.cut(housing["median_income"],
bins=[0., 1.5, 3.0, 4.5, 6., np.inf],
labels=[1, 2, 3, 4, 5])
```
# Geographic Distribution
```
import matplotlib.image as mpimg
california_img=mpimg.imread(os.path.join(images_path, filename))
ax = housing.plot(kind="scatter", x="longitude", y="latitude", figsize=(10,7),
s=housing['population']/100, label="Population",
c="median_house_value", cmap=plt.get_cmap("jet"),
colorbar=False, alpha=0.4,
)
plt.imshow(california_img, extent=[-124.55, -113.80, 32.45, 42.05], alpha=0.5,
cmap=plt.get_cmap("jet"))
plt.ylabel("Latitude", fontsize=14)
plt.xlabel("Longitude", fontsize=14)
prices = housing["median_house_value"]
tick_values = np.linspace(prices.min(), prices.max(), 11)
cbar = plt.colorbar()
cbar.ax.set_yticklabels(["$%dk"%(round(v/1000)) for v in tick_values], fontsize=14)
cbar.set_label('Median House Value', fontsize=16)
plt.legend(fontsize=16)
#save_fig("california_housing_prices_plot")
plt.show()
```
## Correlation with `median_house_value`
```
corr_matrix["median_house_value"].sort_values(ascending=False)
```
## Scatterplot Matrix
```
# from pandas.tools.plotting import scatter_matrix # For older versions of Pandas
from pandas.plotting import scatter_matrix
attributes = ["median_house_value", "median_income", "total_rooms",
"housing_median_age"]
scatter_matrix(housing[attributes], figsize=(12, 8))
#save_fig("scatter_matrix_plot")
plt.show()
```
#### Median House Value vs. Median Income
```
housing.plot(kind="scatter", x="median_income", y="median_house_value",
alpha=0.1)
plt.axis([0, 16, 0, 550000])
#save_fig("income_vs_house_value_scatterplot")
plt.show()
housing["rooms_per_household"] = housing["total_rooms"]/housing["households"]
housing["bedrooms_per_room"] = housing["total_bedrooms"]/housing["total_rooms"]
housing["population_per_household"]=housing["population"]/housing["households"]
```
#### Scatterplot Rooms Per Household vs. Median House Value
```
housing.plot(kind="scatter", x="rooms_per_household", y="median_house_value",
alpha=0.2)
plt.axis([0, 5, 0, 520000])
plt.show()
```
| github_jupyter |
### This post is to help provide an intuitive understanding of a (dumb) 2-Layered Neural Network.
* In this following post, we will start off with building a 1-Layer Neural Net, and how it maps to Logistic Regression. We will test our functions/ model with multiple datasets.
* Later, we will improve on our initial model and include a single hidden layer, and see how our performance increases in classification task. We will also tune/ tweak the number of hidden nodes in our hidden layer, and try to gain an intuition about how Neural Nets could be used efficiently over traditional classification models.
* In the next post, once we understand the basic building blocks of non-linear functions, along with a generalized forward and backward propagation steps, we will construct our very own Deep Neural Network package, with Model() and Layer().
* My 2 cents: Don't worry about optimization yet; details about activation functions, mini-batch gradient descent, RMSprop, Adams Optimization and other techniques are used to fine-tune model performance, which will be covered in a different post.
```
#importing necessary libraries
import numpy as np
import matplotlib.pyplot as plt
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
```
### Activation Functions
Let's load the activation functions we will use in this classification task. We use Sigmoid function to map our input to [0,1] in binary classification task.
* The final output layer uses the Sigmoid function (remember that the activation function of Logistic regression is based on sigmoid fucntion).
* Tanh activation function is similary to Sigmoid function, but y-values go below 0. (i.e., [-1,1]), derivates are steeper when compared to Sigmoid function.
* Let's take an array from -3 to +3 with step size of 0.05. After 'activating' the input with sigmoid, tanh and relu, we see that tanh is a transformation of the sigmoid activation, whereas relu gives maximum of 0 and the value itself.
```
# Sigmoid function -> maps input to [0,1]
def sigmoid(X):
return (1/(1+np.exp(-X)))
# Tanh fucntions -> maps input to [-1,1]
def tanh(X):
return np.tanh(X)
# Rectifier which maps to [0,max(x)]
def relu(X):
return np.maximum(0,X)
# input vector to activation functions
x = np.arange(-3,3,0.05)
# activation steps
y_sigmoid = sigmoid(x)
y_tanh = tanh(x)
y_relu = relu(x)
# 2 plots
plt.figure(figsize=(14,6))
plt.subplot(121), plt.scatter(x, y_sigmoid, label='sigmoid'), plt.scatter(x, y_tanh, label='tanh')
plt.grid(), plt.legend(loc='upper left'), plt.title("Sigmoid vs Tanh")
plt.xlabel('x'), plt.ylabel('activated x')
plt.subplot(122)
plt.scatter(x, y_sigmoid, label='sigmoid'), plt.scatter(x, y_tanh, label='tanh')
plt.scatter(x, y_relu, label='relu'), plt.grid()
plt.legend(loc='upper left'), plt.title("Sigmoid vs Tanh vs Relu"), plt.xlabel('x'), plt.ylabel('activated x')
plt.show()
```
* From the above graphs, we understand why Sigmoid is used for our final Layer (or Logistic Regression) in case of Binary classification; to predict either 0 or 1. Now, lets load our dataset and explore it further.
## 1-Layer Neural Net (Logistic regression)
Let's load a simple dataset with two classes, and see how a basic 1-Layer Neural Network, is the same as a Logistic Regression Task, and how activation functions are used to activate linear combinations of input vectors and parameters.
```
# take features into X and targets into y
from sklearn.datasets import make_moons
# Dataset, set sample size
X, y = make_moons(n_samples=800, noise=0.5)
# Splitting into train test
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y.ravel(), test_size=0.5, random_state=27)
# working with transposes for ease of matrix manipulation
X_train, X_test, y_train, y_test = X_train.T, X_test.T, y_train.T, y_test.T
# shapes
X.shape, y.shape
# Viz of input data
plt.scatter(X[:,0], X[:, 1], c=y)
```
* Now, let us consider the input feature matrix X-transpose, with shape (2, 800), represented as the zeroth layer/ input layer of our Neural Network. As we have two features (x1, x2) for our input data, we would have 2-weights (w1, w2), along with the term b, as the parameters of our model.
### Propagation Step:
* The function initialize_weights returns (w, b), where w is the weight vector of shape (2,1) and b is zero. It uses numpy random.rand function to generate values with seed set to 27.
* We form a linear combination of (x1, x2), by using randomly initialized weights (w1, w2, b), to form Z.
* We then activate Z, by using Sigmoid function, which returns a probability of belonging to class 1, from [0,1].
* Then, we calculate the cost/ loss with the parameters (w1, w2, b). However, we need to update our parameters such that the Log loss is minimized, read <a href="https://adivarma27.github.io/LogisticRegressionCost/#">this article</a> for detailed explanation. Hence, we need to update the parameter vectors w and b, in such a way that the Log loss is minimized. We need to perform gradient descent, and find the slope of Loss function, with respect to each of the parameters.
* Below is a flow diagram of the 1-Layer NN.
Note: In the following image below, w1 and w2 are vectorized into w for matrix multiplication.
<img src="img1.jpeg" width="700">
### Back-Propagation Step:
* As this is a Single Layer Neural Net, there is just one back-propagation step involved. Each of the input parameter vector (w and b, in this case), decreases its value by a learning rate '$\alpha$' times the derivative of loss function with respect to itself. Hence, the learning_rate in itself is a hyperparameter and needs to be initialized properly, for efficient training process.
* Below is the math behind finding derivatives of cost function with respect to parameters, which will aid with update parameter step.
<img src="img2.jpeg" width="800">
<img src="img3.jpeg" width="800">
* In the back-propagation step, we need to update only our weight parameters (w, b); however to calculate derivative of cost function with respect to parameter, we need to propagate through the Linear combination (Z), as well the activated output (A).
* As this Neural-Net has a single layer, the back-propagation is straight forward. As the number of hidden layers increase, we can generalize the forward & backward propagation step.
* Once the output is computed, the predict function takes in the y vector which contains probabilities of belonging to class 0 or 1 and assigns class based on 0.5 threshold.
```
# function to initialize weights
def initialize_weights(X, seed=27):
np.random.seed(27)
w = np.random.rand(X.shape[0],1)
b = 0
return w, b
# function to predict class on 0.5 threshold
def predict(y):
y_list = []
y = y[0]
for i in range(0,len(y)):
if y[i] > 0.5:
y_list.append(1)
else:
y_list.append(0)
return y_list
# See post https://adivarma27.github.io/LogisticRegressionCost/# for detailed explanation
def train(X, y, w, b, max_iterations=1000, learning_rate=0.01):
m = X.shape[1]
learning_rate = learning_rate
max_iterations = max_iterations
cost_list, train_acc_list, test_acc_list, w1_list, w2_list, b_list = [], [], [], [], [], []
# iteration
for iteration in range(max_iterations):
# linear-combination, activation step
Z = np.dot(w.T, X) + b
A = sigmoid(Z)
# compute cost
cost = -(1/m)*np.sum(y*np.log(A) + (1-y)*np.log(1-A))
cost_list.append(cost)
# back-prop
# derivative with respect to cost function
dw = (1/m)*np.dot(X, (A-y).T)
db = (1/m)*np.sum(A-y)
# parameter update step
w = w - learning_rate*dw
b = b - learning_rate*db
w1_list.append(w[0])
w2_list.append(w[1])
b_list.append(b)
train_acc_list.append(accuracy_score(predict(A), y.squeeze()))
test_acc_list.append(PredictClass(X_test, w, b))
return cost_list, train_acc_list, test_acc_list, w1_list, w2_list, b_list
# Forward prop class prediction
def PredictClass(X, w, b):
y_pred_list = []
A = sigmoid(np.dot(w.T, X) + b)
ypred = predict(A)
y_pred_list.append(ypred)
return y_pred_list
# Function to rpedict test data
def PredTestData(test_acc_list):
y_pred_list = []
for i in range(0,len(test_acc_list)):
y_pred_list.append(accuracy_score(test_acc_list[i][0], y_test))
return y_pred_list
```
### For each iteration:
Propagation Steps:
1. compute linear combination of weights and input vector
2. activate input vector and store in vector A
3. compute cost with the current weights
Back-Prop Steps:
4. compute derivates/ slope of cost function with respect to corresponding weights
5. Update weights
Prediction:
6. Predict class
```
# hyperparameters alpha, max_iterations
learning_rate = 0.05
max_iterations = 1000
# Initialize weights
w, b = initialize_weights(X_train, seed=27)
# train over X_train, y_train
cost_list, train_acc_list, test_acc_list, w1_list, w2_list, b_list = train(X_train, y_train, w, b)
# testing data prediction task
test_pred = PredTestData(test_acc_list)
# subplots
plt.figure(figsize=(14,6)), plt.subplot(121)
plt.scatter(np.arange(0, max_iterations), cost_list), plt.xlabel('# of Iterations'), plt.ylabel('Cost')
plt.title('Scatter plot of Decreasing Cost over number of iterations'), plt.grid()
plt.subplot(122), plt.scatter(np.arange(0,max_iterations), train_acc_list, label='Training Accuracy')
plt.scatter(np.arange(0,max_iterations), test_pred, label='Testing Accuracy')
plt.legend()
plt.xlabel('# of Iterations'), plt.ylabel('Accuracy')
plt.title('Scatter plot of Training Accuracy over number of iterations'), plt.grid()
plt.show()
```
#### As we see from the above graphs, as the number of iterations reaches 1000, we observe saturating Training & Testing Accuracy, and we achieve a testing accuracy of ~ 77 %. In the above 1-Layer Neural Net (Logistic Regression), we do a decent job, where the input features are just linear combinations along with weights w1, w2, b.
#### Below, we plot the weights w1, w2 and see how they converge by our gradient descent step, with x-axis as w1, w2 values and y-axis as cost function. Initially, the gradient step is higher; in later iterations the step size decreases.
```
plt.scatter(w1_list, cost_list, label='w1'), plt.scatter(w2_list, cost_list, label='w2'),
plt.legend(), plt.xlabel('w1, w2 values'), plt.ylabel('Cost Function')
```
## 2-Layer Neural Net (1-Hidden Layer, n-Hidden Units)
* In the above exercise, we saw how Logistic Regression, is a **simple Feed-forward neural network**. Now, lets include a hidden layer in our Neural Net, and select number of hidden units as a hyperparameter in our model.
* We can also choose between different activation functions (relu or tanh) functions for activating our hidden layer, after which we will activate the final layer with Sigmoid function, to obtain probability of belonging to class 1. This is where we understand the advantages of Neural Nets over Logistic Regression model.
* Let us choose a complex dataset to understand the power of hidden layers/ hidden units in our Neural net model.
```
# Complex dataset
# source: https://github.com/rvarun7777/Deep_Learning/blob/master/Neural%20Networks%20and%20Deep%20Learning/Week%203/Planar%20data%20classification%20with%20one%20hidden%20layer/planar_utils.py
# function to load our dataset
def load_planar_dataset():
np.random.seed(1)
m = 400 # number of examples
N = int(m/2) # number of points per class
D = 2 # dimensionality
X = np.zeros((m,D)) # data matrix where each row is a single example
Y = np.zeros((m,1), dtype='uint8') # labels vector (0 for red, 1 for blue)
a = 4 # maximum ray of the flower
for j in range(2):
ix = range(N*j,N*(j+1))
t = np.linspace(j*3.12,(j+1)*3.12,N) + np.random.randn(N)*0.2 # theta
r = a*np.sin(4*t) + np.random.randn(N)*0.2 # radius
X[ix] = np.c_[r*np.sin(t), r*np.cos(t)]
Y[ix] = j
X = X.T
Y = Y.T
return X, Y
# take features into X and targets into y
X, y = load_planar_dataset()
# shapes
X.shape, y.shape
```
* We obtained around 75-80 % testing accuracy on previous dataset by using 1 Layer NN. We observe that the same model has only 50 % accuracy on the new dataset. (Model has just w1, w2 & b to capture complex decision boundary)
```
plt.scatter(X[0,:], X[1,:], c=y.ravel())
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegressionCV
X_train, X_test, y_train, y_test = train_test_split(X.T, y.T, test_size=0.5)
clf = LogisticRegressionCV()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(accuracy_score(y_test, y_pred)*100, '% accuracy')
```
## Why is 2-Layer NN better than 1-Layer NN/ Any traditional Machine Learning Model ?
* From the above, we see that traditional single layer NN/ Logistic Regression gives only around 50 % accuracy. Clearly, there are hidden patterns which the model is unable to capture. The model needs more number of parameters to be able to capture the pattern in our dataset, to form/ contruct non-linear complex boundaries.
* The **power** of hidden layers lies in the fact that these complex decision boundaries can be captured; and generally speaking, more the number of hidden units in a layer, better we can train the data. Previously, we had only w1, w2 & b to tune to find the best decision boundary. As the number of layers and number of nodes increase, we have many more parameters to optimize. The weights which are randomly initialized, start 'self-adjusting' or 'self-correcting', where the combinations of different weights in different nodes and layers, try to minimize log-loss. After multiple iterations, the layers start recognizing patterns in the data and start self-correcting. Finally, we are left with many more number of parameters/ weight vectors, which could overfit to the training data. Essentially, each parameter (on each layer), tries to find the derivative of cost/ loss function with respect to itself, to update the parameter value.
* Since we have one hidden layer and 4 hidden units, our parameters are now vectors W1, W2, B1, B2, where W1, B1 correspond to the input layer (input layer to hidden layer), and W2, B2 (weights corresponding to hidden layer to output). W1 = [w1, w2] (input layer), W2 = [w1, w2, w3, w4] (4 hidden nodes).
<img src="img4.jpeg" width="800">
* Below are the functions **initialize_weights_2layers(X, n_hidden_units)** and **trainHidden(params, X, max_iterations)** to initialize weights vector based on number of hidden features in second layer.
```
# function to initialize parameters
def initialize_weights_2layers(X, n_hidden_units = 10, seed=27):
np.random.seed(seed=seed)
params = {}
W1 = np.random.randn(n_hidden_units, X.shape[0])*0.01
B1 = np.zeros(shape=(n_hidden_units, 1))
W2 = np.random.randn(1,n_hidden_units)*0.01
B2 = np.zeros(shape=(1,1))
params['W1'], params['W2'], params['B1'], params['B2'] = W1, W2, B1, B2
return params
params = initialize_weights_2layers(X, n_hidden_units=2)
W1, W2, B1, B2 = params['W1'], params['W2'], params['B1'], params['B2']
params
# function to train 2 Layered NN
def trainHidden(params, X, max_iterations=8000):
W1, W2, B1, B2 = params['W1'], params['W2'], params['B1'], params['B2']
m = X.shape[1]
learning_rate = 1
acc = []
# iteratiing
for iteration in range(max_iterations):
# linear combination and first activation
Z1 = np.dot(W1, X) + B1
A1 = tanh(Z1)
# linear combination and second activation
Z2 = np.dot(W2, A1) + B2
A2 = sigmoid(Z2)
acc.append(predict(A2))
# calculating cost
cost = -(1/m)*np.sum(np.multiply(np.log(A2), y) + np.multiply((1 - y), np.log(1 - A2)))
# backprop step
dz2 = A2 - y
dw2 = (1/m)*np.dot(dz2,A1.T)
db2 = (1/m)*np.sum(dz2,axis=1,keepdims=True)
dz1 = np.multiply(np.dot(W2.T,dz2),1-np.power(A1, 2))
dw1 = (1/m)*np.dot(dz1,X.T)
db1 = (1/m)*np.sum(dz1,axis=1,keepdims=True)
# parameter update step
W1 = W1 - learning_rate*dw1
B1 = B1 - learning_rate*db1
W2 = W2 - learning_rate*dw2
B2 = B2 - learning_rate*db2
return acc
# function to train over 'n' hidden units
def trainhiddenUnits(n_hidden_units):
params = initialize_weights_2layers(X, n_hidden_units)
acc = trainHidden(params, X=X, max_iterations=5000)
accuracy = []
for i in range(0,len(acc)):
accuracy.append(accuracy_score(acc[i], y.squeeze()))
return accuracy
```
Below are the accuracies after including one hidden layer (with various number of hidden units). We observe that by using 3-hidden units, the model is able to capture the complexity and able to predict pretty well. (~ 40% increase in accuracy when compared to 1 Layered Neural Net).
```
# list of accuracies for n hidden units
accuracies_ = []
for n_hidden_units in range(1,5):
accuracies_.append(trainhiddenUnits(n_hidden_units))
# Plots for various number of hidden units
plt.figure(figsize=(16,8))
plt.scatter(np.arange(0,5000), accuracies_[0], label='1 hidden unit')
plt.scatter(np.arange(0,5000), accuracies_[1], label='2 hidden units')
plt.scatter(np.arange(0,5000), accuracies_[2], label='3 hidden units')
plt.scatter(np.arange(0,5000), accuracies_[3], label='4 hidden units')
plt.xlabel('# of iterations')
plt.ylabel('Accuracy')
plt.legend()
plt.grid()
```
### Why does predictive power saturate at 3 Hidden Units ? Why not 100 units if it performs better ?
* As you see below, once we reach the decision boundaries which can form separation between the two classes, predictive power saturates, beyond which it overfits to the data.
* For the same data, 1 Layered NN performed poorly (~45 % testing accuracy), as it was not able to understand this pattern, which looks simple to the human eye.
```
plt.figure(figsize=(12,8))
plt.scatter(X[0,:], X[1,:], c=y.ravel())
plt.plot([-4.5, 4.5], [-0.8, 0.8], 'k-', lw=2)
plt.plot([-4.5, 4.5], [-3.1, 3.1], 'k-', lw=2)
plt.plot([-4.5, 4.5], [3.25, -3.2], 'k-', lw=2)
```
* Now, lets try another dataset, where decision boundary can be visually understood. First, we use Logistic Regression to see if the model can understand the obvious pattern of different classes.
```
from sklearn.datasets.samples_generator import make_circles
X, y = make_circles(n_samples=400,factor=0.5,noise = 0.1)
plt.figure(figsize=(12,8))
plt.scatter(X[:, 0], X[:, 1], c=y, s=50, cmap = 'winter')
plt.title('Nonlinear Data')
X, y = X.T, y.T
```
* We observe that the accuracy is only around 50 %. Hence, just W and b vectors are not enough to model the non-linearity that exists in the data.
```
X_train, X_test, y_train, y_test = train_test_split(X.T, y.T, test_size=0.5)
clf = LogisticRegressionCV()
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(accuracy_score(y_test, y_pred)*100, '% accuracy')
```
### Hence, we traing the data using 2-Layered Neural Net, over various number of hidden layers, and find that 3 hidden units can predict with 100 % accuracy.
```
# list of accuracies for n hidden units
accuracies_ = []
for n_hidden_units in range(1,4):
accuracies_.append(trainhiddenUnits(n_hidden_units))
# Plots for various number of hidden units
plt.figure(figsize=(16,8))
plt.scatter(np.arange(0,5000), accuracies_[0], label='1 hidden unit')
plt.scatter(np.arange(0,5000), accuracies_[1], label='2 hidden units')
plt.scatter(np.arange(0,5000), accuracies_[2], label='3 hidden units')
plt.xlabel('# of iterations')
plt.ylabel('Accuracy')
plt.legend()
plt.grid()
```
#### Deep Layered Neural Nets can now outperform us at classification tasks. We are yet to tune all the hyper-parameters, use optimization techniques to help parameters converge faster, and also look at Convoluted/ Recurrent layers to improve performance.
| github_jupyter |
# Jupyter notebook test file
Welcome to the first jupyter notebook. This is a test file to verify your installation of python.
If you have never heard about or used jupyter notebooks before, don't worry! Jupyter notebooks provide an easy way to combine formatted text (such as this paragraph) with python code (such as the box below). The great thing is that you can also execute the python code live on this page. Try it out by clicking on the `Run` button in the toolbar above.
On the first click, the blue box around the headline probably jumped to the next paragraph, but nothing else happened, right? This is how jupyter notebooks operate: the formatted text blocks and the code blocks are organised in _cells_ – each of which can be executed. Now, executing a cell with only text doesn't do anything of course ... But with every click on `Run`, you run the active (blue) cell and jump to the next one. If this paragraph is currently marked in blue, click on `Run` twice, to execute both this cell with formatted text, and the cell with python code below.
```
import sys
sys.version
```
Great, you've executed your first bit of python code! Python is an extremely _modular_ language, meaning that it is organised in chunks of code called _modules_. These modules provide all sorts of functionalities, usually dedicated to specific tasks. For example, the module collection [numpy](https://www.numpy.org/) provides functionality for numerical computations. It is one of the most widely used collections in scientific programming. By the way, collections of modules are called `packages`.
The above two commands import one module called `sys`, which provides some functions to learn about the python installation itself. The command `sys.version` for example tells you the version of python that you are running on. Hopefully, the output says something like:
```
Python version: 3.8.8 (some more info ...)
```
which is the python version that we would like to use for these tutorials.
### Installation of basic packages
Click on the `Run` button again to jump to the cell of python code below and execute it.
```
import numpy as np
np.__version__
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.__version__
```
As you might have guessed, the two `import` lines above load two more modules: the numpy package for scientific programming, and [matplotlib](https://matplotlib.org), a package for visualising data. Two more gimmicks:
- the `import` command can also creates aliases for modules/packages with the `as` keyword. This can be useful if the names of modules/packages are very long. In the above example, anything within `matplotlib` can be accessed with the alias `mpl`.
- You can also import only _parts_ of a package. Many packages have functionalities bundled in their own small modules. In the above example, the `matplotlib.pyplot` module is imported separately from `matplotlib` and is given the alias `plt`.
If your python installation was done correctly, you should get version numbers like these:
```
Numpy version: 1.19.5
Matplotlib version: 3.3.4
```
The patch version (which is the number after the second dot) might be different, but the major and minor version number should be identical.
Usually, this should be enough to verify the installation. But let's run the following piece of code, which uses these two packages to create some random data, perform a polynomial regression, and plot the result:
```
import seaborn as sns
# Generate 50 random x values in the interval [0, 2].
# For each of these random values, generate a y value
# following the formula y = 3 + 4x. To make our data
# more interesting, we add some random noise.
m = 50
X = 2 * np.random.rand(m)
y = 3 + 5 * X + 2 * np.random.randn(m)
# Add some labels to the axes before plotting.
plt.xlabel("$x$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
# Now, import the seaborn package, another library for
# data visualisation (built on top of matplotlib). Use
# seaborn to perform a linear regression and plot it.
sns.regplot(x=X, y=y)
```
Hopefully this worked, too. By the way, you can always click on the code and start to modify it! For example, try to change the value of `m` in the above cell, and then run the cell again. Does the plot look different?
Another thing: you can always jump back in forth within a notebook by using your up/down arrow keys or simply clicking on the cells. Sometimes it is useful to reset the jupyter notebook and start from scratch. This will remove all generated output and only leave the formatted text and code cells. To do this, click on the `Kernel` drop-down menu and click on `Restart & Clear Output`. The kernel restart means that python will also "forget" about anything you did: variables you assigned, modules you imported etc.
### Machine-learning libraries
Before we finish, let's make sure our machine-learning packages have the correct version as well. Execute the following code cell to import [Scikit-Learn](https://scikit-learn.org/) and [tensorflow](https://www.tensorflow.org/) and check their versions:
```
import sklearn as skl
skl.__version__
import tensorflow as tf
tf.__version__
```
If everything is set up correctly, the versions should be:
```
scikit-learn version: 0.24.2
tensorflow version: 2.4.1
```
Hurray, the setup was successful!
| github_jupyter |
<a href="https://colab.research.google.com/github/The-Geology-Guy/sample_determination/blob/main/SampleDetermination.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Null Hypothesis Significance Testing (NHST)
##### _March, 2019_
PAJER, Luke. luke.pajer@gmail.com
---------------------
```
import numpy as np
import pandas as pd
import scipy.stats
from IPython.display import display, Markdown, Latex
pd.set_option('display.max_colwidth', 0)
```
#### Determine Alpha Function
```
def alpha_cv(a):
alpha_dict = {"0.10": 1.282, "0.05": 1.645, "0.025": 1.960, "0.01": 2.326, "0.005": 2.576, "0.001": 3.090, "0.0005": 3.291}
return alpha_dict.get(a, "")
```
#### Determine Rejection of Null Hypothesis Function
```
def rejection(z_cv, z_alpha, rr):
if (rr == 'upper'):
test = (z_cv >= z_alpha)
elif (rr == 'lower'):
test = (z_cv <= (-1 * z_alpha))
elif (rr == 'two'):
test = ((z_cv >= z_alpha) & (z_cv <= z_alpha))
return test
```
#### Explain the Rejection Function
```
def rej_explain(z, a, alpha, r, rr):
if ((r == True) & (rr == 'upper')):
explain = (str(np.round(z, 3)) + " $\geq$ " + str(a) +
r": $\text{Reject}$ $H_{0}$ because the Test statistic $(z)$ falls in the Rejection Region ($z$ $\geq$ $z_{\alpha}$ = " +
str(alpha) + ").")
elif ((r == True) & (rr == 'lower')):
explain = (str(np.round(z, 3)) + " $\leq$ -" + str(a) +
r": $\text{Reject}$ $H_{0}$ because the Test statistic $(z)$ falls in the Rejection Region ($z$ $\leq$ $-z_{\alpha}$ = " +
str(alpha) + ").")
elif ((r == True) & (rr == 'two')):
explain = ""
elif ((r == False) & (rr == 'upper')):
explain = (str(np.round(z, 3)) + " $\leq$ " + str(a) +
r": $\text{Do not reject}$ $H_{0}$ because the Test statistic $(z)$ does not fall in the Rejection Region ($z$ $\geq$ $z_{\alpha}$ = " +
str(alpha) + ").")
elif ((r == False) & (rr == 'lower')):
explain = (str(np.round(z, 3)) + " $\geq$ -" + str(a) +
r": $\text{Do not reject}$ $H_{0}$ because the Test statistic $(z)$ does not fall in the Rejection Region ($z$ $\leq$ $-z_{\alpha}$ = " +
str(alpha) + ").")
elif ((r == False) & (rr == 'two')):
explain = ""
return display(Markdown(explain))
```
#### Compute the Test Statistic Value
```
def ztest(p_hat, p_0, nt, alpha, reject_region):
valid_head = ((nt * p_0) >= 10)
valid_tail = ((nt * (1 - p_0)) >= 10)
if (valid_head & valid_tail):
zt = (p_hat - p_0)/(np.sqrt((p_0 *(1 - p_0))/nt))
alpha_1 = alpha_cv(alpha)
answer = rejection(zt, alpha_1, reject_region)
return rej_explain(zt, alpha_1, alpha, answer, reject_region)
else:
return display(Markdown("Does not meet the test procedures requirements"))
```
#### Solve for Sample Size needed
_Take a look into the beta error function, does it need to be subtracted by one?_
```
def sampleDeter(p0, p1, za, ns):
za = alpha_cv(za)
phi_z = (p0 - p1 + za * np.sqrt(abs(p0 * (1 - p0))/ns)) / np.sqrt(abs((p1 * (1 - p1)))/ns)
beta_err = scipy.stats.norm.cdf(phi_z)
n = (((za * np.sqrt(p0 * (1 - p0))) + (za * np.sqrt(p1 * (1 - p1)))) / (p1 - p0)) ** 2
retest = p0 + (za * np.sqrt((p0 * (1-p0))/ns))
return pd.DataFrame.from_dict({r'$\beta$': [np.round(beta_err, 3)],
r"$n$": [int(np.round(n, 0))],
r"$z$": [np.round(phi_z, 3)],
r"$p$": [np.round(p0, 3)],
r"$p'$": [np.round(p1, 3)],
r"$c$": [np.round(retest, 3)]
})
```
### Run the NHST for each Type I Error $(\alpha)$
```
def to_md(p0, p1, n, tail):
alpha_list = ["0.10", "0.05", "0.025", "0.01", "0.005", "0.001", "0.0005"]
i = 0
while i < len(alpha_list):
alpha_a = alpha_list[i]
display(Markdown("--------------------"))
display(Markdown(r"$\text{For Type I Error } (\alpha)$ = " + alpha_a))
ztest(p1, p0, n, str(alpha_a), tail)
test = sampleDeter(p0, p1, str(alpha_a), n)
beta_, n_, z_, p0_, p1_, c_ = test.iloc[0,0], test.iloc[0,1], test.iloc[0,2], test.iloc[0,3], test.iloc[0,4], test.iloc[0,5]
display(Markdown(r"If the experiment collects data on $n =$ " + str(n_) + r" users, and $\text{Rejects}$ $H_{0}: p = $ "
+ str(float(round(p0, 3))) + " if the proportion of sampled users who favor the change is greater than " + str(c_) +
", then there will be a " + str(int(round(float(alpha_a), 2) * 100)) +
r"% chance of committing a Type I error $(\alpha)$ and a " +
str(int(round(beta_, 2) * 100)) + r"% chance of committing a Type II error $(\beta)$ if the population proportion were actually " +
str(p1_) + "."))
display(Markdown(r"$\hat{p} =$ " + str(float(round(p0, 3))) + r", $p =$ " + str(float(round(p1, 3))) + r", $c =$ " + str(c_)))
display(Markdown(r"$n =$ " + str(n) + r", $n_{c} =$ " + str(n_)))
display(Markdown(r"$\alpha =$ " + str(alpha_a) + r", $\beta =$ " + str(beta_)))
i +=1
```
#### Enter values for the variables
`p0` is the original or unmodified data to test
`p1` is the modified data to test
`n` is the total number of trials for the p1
`upper` is for predicting an increase
```
p0, p1, n, tail, = 0.0301, 0.0487, 256836, "upper"
display(Markdown("### Transaction Analysis"))
to_md(p0, p1, n, tail)
```
| github_jupyter |
Copyright 2020 Vasile Rus, Andrew M. Olney and made available under [CC BY-SA](https://creativecommons.org/licenses/by-sa/4.0) for text and [Apache-2.0](http://www.apache.org/licenses/LICENSE-2.0) for code.
# Decision Trees
So far we have talked about the KNN classifier, which uses *closeness to known points* to classify, and logistic regression, which uses *probability* to classify.
Decision trees are a new idea that uses a *tree* to classify.
Building a decision tree means, as the name implies, generating a tree in which each internal node of the tree is a decision point, usually deciding which of that node's children to consider next based on the values of the predictor or feature associated with the current node.
The goal is to reach a leaf node which corresponds to a prediction, for instance, the category of the object/instance described by the predictors/features.
## What you will learn
In this notebook, you will learn about decision trees, an original data
science paradigm to approach primarily classification tasks, and how it can
be used to infer from labeled/annotated data decision tree based
classifiers. We will study the following:
- The basics of decision trees
- Details about how decision trees are built
- Criteria to evaluate features/predictors
- Interpreting paths from the root to leaf nodes
- Visualization of decision trees
- Evaluation of performance for decision trees
## When to use decision trees
Decision trees are useful when you have a categorical response/outcome
variable and there are multiple features/predictors that can be used to
predict the correct value of the outcome variable. The ultimate goal is to
build automatically a decision tree to predict the
correct value of the outcome variable for a new instance described by the
set of predictors/features. It can also be used as a feature selection tool
to determine which features/predictors are most discriminatory with respect
to the outcome variable. Decision trees have the major of being
interpretable over many other classification paradigms.
## Example
An example of such a tree is given below, just for illustration purposes.
It uses two decision nodes (the circles/blue nodes): `Rainy`, which has two values Yes and No, and `Fit`, which has two values as well: Yes - meaning being fit to play - and No - not being fit to play for whatever reason, medical or not.
The leaf nodes at the bottom correspond to the outcomes: Play or No-Play.
Using this tree, if it is NOT `Rainy` and `Fit` is Yes, the decision in the corresponding leaf node is Play.
That is, a path from the root/top node to a leaf node illustrates a decision path resulting in the decision shown in the leaf node.
Each path in decision trees can be expressed as an IF-THEN, for instance: IF (Rainy == No) AND (Fit == Yes) THEN Play.
While this example uses nominal variables that are binary (yes/no), the same approach also works for other variable types.
For example, ratio variables can be given a decision threshold such that one path is followed if the value is above that threshold vs. below.
Continuing the example below, if we replaced `Rainy` with `Cloud cover`, then we could set a threshold of 50%, so that if `Cloud cover` was above 50%, then we would follow the arrow to No-Play and otherwise we'd follow the arrow to `Fit`.

It's one thing to create a tree by hand - the magic is learning a tree like this from the data.
As a simple example, we consider the artificially created dataset below:
| Rainy | Fit | Play |
|------------|------------|-------------|
| 0 | 0 | 0 |
| 0 | 1 | 1 |
| 1 | 1 | 0 |
| 1 | 0 | 0 |
<!-- NOTE: trying to simplify
In this dataset, there are four instances. Each instances has two predictors, say $predictor_1$ and $predictor_2$, which are the first values in each instance. $Predictor_1$ could indicate whether it is rainy (1) or not (0) whereas $predictor_2$ may indicate whether a person feels fit (1) to play, say, tennis or not (0). The last value in each instance is the class: 0 (no play) or 1 (play).
The Python code below illustrates how to build a decision tree based on this simple data. -->
Let's build a decision tree using this simple data:
- Create variable `X` and set it to a list containing
- a list containing
- 0
- 0
- a list containing
- 0
- 1
- a list containing
- 1
- 1
- a list containing
- 1
- 0
- Create a variable `Y` and set it to a list containing
- 0
- 1
- 0
- 0
As you can see, we've recreated the table above using these lists.
Normally we work with larger dataframes, so you may not have thought of it this way, but ultimately all our data is just lists of numbers like this.
```
X = [[0, 0], [0, 1], [1, 1], [1, 0]]
Y = [0, 1, 0, 0]
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="_z/H/tTnOYS1Lla2~;*0">X</variable><variable id="o8#tp%!,_Okj2aVwX(DL">Y</variable></variables><block type="variables_set" id="Iwb#y?nx(@|hrx`dPD21" x="225" y="122"><field name="VAR" id="_z/H/tTnOYS1Lla2~;*0">X</field><value name="VALUE"><block type="lists_create_with" id="M(r*hCGi-1+FP3$9I/jI"><mutation items="4"></mutation><value name="ADD0"><block type="lists_create_with" id="!mtX-`LPTj2ek-kix54P"><mutation items="2"></mutation><value name="ADD0"><block type="math_number" id="w@E:*IEeZ_vb.xt/W.1`"><field name="NUM">0</field></block></value><value name="ADD1"><block type="math_number" id="m1Q];BPvl{pmtXT^T9*u"><field name="NUM">0</field></block></value></block></value><value name="ADD1"><block type="lists_create_with" id="/G|.+K|h9-p~!!LHIkw}"><mutation items="2"></mutation><value name="ADD0"><block type="math_number" id="Q=zSR?(Ysp[S{hh_0}t@"><field name="NUM">0</field></block></value><value name="ADD1"><block type="math_number" id="KT4OlB?BCwAy~6VyG0oh"><field name="NUM">1</field></block></value></block></value><value name="ADD2"><block type="lists_create_with" id="^_N6We|_:gx(xQiFu]aW"><mutation items="2"></mutation><value name="ADD0"><block type="math_number" id="O+U8]nd%wt}c-:C#Ho1D"><field name="NUM">1</field></block></value><value name="ADD1"><block type="math_number" id=":*9[$?!#gwbPt69dmLo|"><field name="NUM">1</field></block></value></block></value><value name="ADD3"><block type="lists_create_with" id="edXG56R~rZt/}bb9.FNe"><mutation items="2"></mutation><value name="ADD0"><block type="math_number" id="#2(OH@:RIQG-221lz|Nc"><field name="NUM">1</field></block></value><value name="ADD1"><block type="math_number" id="tgFCw4(dD*iA^?h`{xP-"><field name="NUM">0</field></block></value></block></value></block></value><next><block type="variables_set" id="Hb9H-%nIxKc,UMwINy57"><field name="VAR" id="o8#tp%!,_Okj2aVwX(DL">Y</field><value name="VALUE"><block type="lists_create_with" id="qk~BgitfP@aj/2oa+`W("><mutation items="4"></mutation><value name="ADD0"><block type="math_number" id="IV{G{Q+%sJS#Nv)XO4;I"><field name="NUM">0</field></block></value><value name="ADD1"><block type="math_number" id="DinOCv.!$Vk[!{j5WaW;"><field name="NUM">1</field></block></value><value name="ADD2"><block type="math_number" id="DEjz[VWhV:(ZleV{V(Y1"><field name="NUM">0</field></block></value><value name="ADD3"><block type="math_number" id="2EzEW`TvJ(5/i@?`.Kh;"><field name="NUM">0</field></block></value></block></value></block></next></block></xml>
```
Next the imports for creating the decision tree model and graphing it:
- `import sklearn.tree as tree`
- `import graphviz as graphviz`
```
import sklearn.tree as tree
import graphviz as graphviz
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="tkeU~z;dQ]^6[?rbF?)9">tree</variable><variable id="rrXqQ:9R(~X0,}F[1UD]">graphviz</variable></variables><block type="importAs" id="g[,/=2yYgO?4Bv[OrVf;" x="16" y="10"><field name="libraryName">sklearn.tree</field><field name="libraryAlias" id="tkeU~z;dQ]^6[?rbF?)9">tree</field><next><block type="importAs" id="5j3UPoKQcO/bU0u!@P06"><field name="libraryName">graphviz</field><field name="libraryAlias" id="rrXqQ:9R(~X0,}F[1UD]">graphviz</field></block></next></block></xml>
```
Create the model:
- Create `decisionTreeExample`
- Set it to `with tree create DecisionTreeClassifier using`
```
decisionTreeExample = tree.DecisionTreeClassifier()
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</variable><variable id="tkeU~z;dQ]^6[?rbF?)9">tree</variable></variables><block type="variables_set" id="}:yCB*e+[r5M{rIc($M]" x="84" y="299"><field name="VAR" id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</field><value name="VALUE"><block type="varCreateObject" id="bcO!l)K~/m6oq}xWIulv"><field name="VAR" id="tkeU~z;dQ]^6[?rbF?)9">tree</field><field name="MEMBER">DecisionTreeClassifier</field><data>tree:DecisionTreeClassifier</data></block></value></block></xml>
```
And fit the model
- `with decisionTreeExample do fit using` a list containing
- `X`
- `Y`
```
decisionTreeExample.fit(X, Y)
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</variable><variable id="_z/H/tTnOYS1Lla2~;*0">X</variable><variable id="o8#tp%!,_Okj2aVwX(DL">Y</variable></variables><block type="varDoMethod" id=",S.=/_X:;`m5Ahy=!xmE" x="40" y="235"><field name="VAR" id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</field><field name="MEMBER">fit</field><data>decisionTreeExample:fit</data><value name="INPUT"><block type="lists_create_with" id="}F0@!:1xqW/}(Dj%-V78"><mutation items="2"></mutation><value name="ADD0"><block type="variables_get" id="2:V:wGJK%j=I8yZEc^o}"><field name="VAR" id="_z/H/tTnOYS1Lla2~;*0">X</field></block></value><value name="ADD1"><block type="variables_get" id="BG#`Opts05WCR={8qe{k"><field name="VAR" id="o8#tp%!,_Okj2aVwX(DL">Y</field></block></value></block></value></block></xml>
```
Let's visualize the tree and interpret how the algorithm fit the tree to the data.
This is a little complicated to set up, but the results are worth it:
- Create `dot_data` and set to `with tree do export_graphviz using` a list containing
- `decisionTreeExample`
- freestyle `out_file=None`
- freestyle `feature_names=['Rainy','Fit']`
- freestyle `class_names=['NoPlay','Play']`
- freestyle `filled=True`
- freestyle `rounded=True`
- freestyle `special_characters=True`
- `with graphviz create Source using` a list containing `dot_data`
```
dot_data = tree.export_graphviz(decisionTreeExample, out_file=None, feature_names=['Rainy','Fit'], class_names=['NoPlay','Play'], filled=True, rounded=True, special_characters=True)
graphviz.Source(dot_data)
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="F.ej=Hud7vT9uozVaF5C">dot_data</variable><variable id="rrXqQ:9R(~X0,}F[1UD]">graphviz</variable><variable id="tkeU~z;dQ]^6[?rbF?)9">tree</variable><variable id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</variable></variables><block type="variables_set" id=":6*zm74#9$!O3?QVUukE" x="209" y="280"><field name="VAR" id="F.ej=Hud7vT9uozVaF5C">dot_data</field><value name="VALUE"><block type="varDoMethod" id="jsRyzb,9t~G^G@G@SMkS"><field name="VAR" id="tkeU~z;dQ]^6[?rbF?)9">tree</field><field name="MEMBER">export_graphviz</field><data>dot_data:</data><value name="INPUT"><block type="lists_create_with" id="r2_rY*ef1v:96w)e_B6K"><mutation items="7"></mutation><value name="ADD0"><block type="variables_get" id="x$]b$%X#}qjO@0H/!O|N"><field name="VAR" id="@[.K6R7D##dBf3mu(%2`">decisionTreeExample</field></block></value><value name="ADD1"><block type="dummyOutputCodeBlock" id="K!=:ty3DMPs#b^ziH=5p"><field name="CODE">out_file=None</field></block></value><value name="ADD2"><block type="dummyOutputCodeBlock" id="by]EM[Y$kMG~MfF0{q^,"><field name="CODE">feature_names=['Rainy','Fit']</field></block></value><value name="ADD3"><block type="dummyOutputCodeBlock" id="2hz~E.{8:W9-wWb*9Ut~"><field name="CODE">class_names=['NoPlay','Play']</field></block></value><value name="ADD4"><block type="dummyOutputCodeBlock" id="^IJ3b3Eg7K9k^6Of_4i%"><field name="CODE">filled=True</field></block></value><value name="ADD5"><block type="dummyOutputCodeBlock" id="VH!NxdBRgItjnN]Zs^Uh"><field name="CODE">rounded=True</field></block></value><value name="ADD6"><block type="dummyOutputCodeBlock" id="f*|7fCy,K/0{_WzfribG"><field name="CODE">special_characters=True</field></block></value></block></value></block></value></block><block type="varCreateObject" id="-a(nDg:Sgtf)Ey5vB4(x" x="187" y="513"><field name="VAR" id="rrXqQ:9R(~X0,}F[1UD]">graphviz</field><field name="MEMBER">Source</field><data>graphviz:Source</data><value name="INPUT"><block type="variables_get" id="!M}@xBlLyRkl~Us)Smh%"><field name="VAR" id="F.ej=Hud7vT9uozVaF5C">dot_data</field></block></value></block></xml>
```
Let's pause for a moment and look at the tree.
The first node *splits* on whether `Rainy` is ≤ 0.05 (i.e is 0 or 1).
If it isn't (i.e. is 1), then we go to the right and `NoPlay`.
If it is, we go to the left and test whether `Fit` is ≤ 0.05 (i.e is 0 or 1).
If it isn't (i.e. is 1), we go to the right and `Play`.
If it is, we go to the left and `NoPlay`.
Underneath the split rule for each node are several properties with an = :
- `samples=` describes how many datapoints fall under that node. So at the top, we have 4 because all samples fall under that node. When we get to the bottom nodes, or **leaves** (which are all fully shaded), `samples` shows how many datapoints fell into each leaf, and those numbers *sum* to 4.
- `value=` describes how many of each class (`NoPlay`, `Play`) fall under that node. Notice the leaf nodes only contain one class or the other - this is important as we'll discuss later.
- `class=` is the class that will be returned for datapoints in the node, but it only has meaning for the leaves.
## Theoretical Background
Building a decision tree means deciding which predictor/feature should correspond to which node of the tree and which predictions should correspond to which terminal nodes or leaf nodes of the tree.
The predictions in the leaf nodes in our case are class label.
Decision trees are commonly used for classification.
The tree is constructed so that predictors that better split the instances according to some criterion such as information gain (IG; to be discussed shortly) are placed higher in the decision tree.
The net effect of such a strategy is to minimize the depth of the tree, i.e., to make the tree as flat as possible.
The general algorithm to infer a particular Decision Tree (i.e., a particular configuration of internal and leaf nodes arranged in a tree) for a given dataset is the following:
1. Select the best predictor based on some criterion such as Information Gain (discussed shortly)
2. Split the data set into subsets based on the values of the chosen predictor in Step 1.
3. Repeat the above process for the subsets until one of the following conditions is met:
- all the instances in the subset belong to the same class.
- there are no more predictors/features left.
- no more instances.
The most widely used predictor selection criteria are:
- Entropy
- Information Gain
- Gain Ratio
- Gini Index
- Chi-square
- Reduction in Variance
### Entropy and Information Gain
One of the most widely used criteria to guide the construction of Decision Trees is $Information Gain (IG)$ which relies on another fundamental concept called $Entropy (E)$.
We will define first entropy E and then IG.
*Entropy*
Entropy is a measure of uncertainty or impurity of a set of items.
It can also be regarded as a measure of the diversity of a set of items.
For instance, in the left urn (Urn A) below there are only red balls.
The impurity of this set of balls is 0 - the purity is perfect as all the balls are of the same color (red) or we can say they all belong to class 'red'.
The urn on the right (Urn B) has two types of balls - red and blue - and therefore is less pure or more diverse.
When drawing balls from Urn A we are certain that the ball is red.
When we draw balls from urn B we are less certain whether the ball will be red as we may also draw a blue ball.

In order to measure this degree of uncertainty, the following formula has been proposed which accounts for the distribution of the possible values/labels/classes/categories C in a set of items S:
$$ E (S) = \sum \limits _{j=1} ^{C} P(c_j)log_2 P(c_j) $$
In general, the more balanced the distribution of categories in a set of items the higher the entropy. You can see the entropy for a set of items with two categories.
<!-- NOTE: on this image, the y-axis is H(X); should we make E(S) to match the equations? -->

As you can see, the entropy is maximum when the two categories have an equal probability 50-50, e.g., 50% chance of drawing a red ball versus 50% chance of drawing a blue ball.
For instance, a set of 30 red balls and 30 blue balls is perfectly balanced and therefore has maximum entropy as opposed to a set with 50 red balls and 10 blue balls in which case the chances/probability of drawing a red balls are much higher.
When we have a set S of items (X,Y) where Y describes possible outcomes and X are the predictors/features/attributes then we can compute the entropy of a partition of S according to the values $v$ of a particular predictor $x$ as in the following:
$$ E (S, x) = \sum \limits _{v \in x} P(x=v) E(S_v) $$
where $x$ is a predictor from the set of predictors $X$ and $E(S,x)$ is the entropy of the split of the set S into subsets ${S_v}$ according to the values of $x$.
As it can be seen, the entropy $E(S,x)$ of such as split is computed as the weighted average of the entropies of each of the subsets $S_v$ corresponding to each of the values $v$ of predictor $x$.
The weight corresponds to the probability of that value $P(x=v)$ and is computed as $\frac {|S_v|} {|S|}$.
*Information Gain*
Information Gain is a selection criterion for predictors and measures how well a predictor can separate the dataset with respect to outcome categories Y.
Predictors that separate the dataset into more pure subsets are preferred as they are more informative, i.e., they will lead to more flat (less deep), simpler trees which should generalize better to new instances.
Intuitively, you can see in the image below two splits of a set S according to one predictor ($P_i$, bottom left split in the figure) or another ($P_j$, bottom right split).

The partition on the right is better as it leads to purer subsets, i.e., the resulting subsets have more balls of the same color.
Information Gain for a predictor $x$ and items set S is defined as:
$$ IG (S,x) = E(S) - E(S,x) $$
which can be expanded based on the previous formulas as follows:
$$ IG (S,x) = E(S) - E(S,x) = \sum \limits _{j=1} ^{C} P(c_j)log_2 P(c_j) - \sum \limits _{v \in x} P(x=v) E(S_v) = \sum \limits _{j=1} ^{C} P(c_j)log_2 P(c_j) - \sum \limits _{v \in x} \frac {|S_v|} {|S|} E(S_v) $$
From the definition, IG can be viewed as a measure quantifying the reduction in entropy of the original dataset versus the entropy of the subsets resulting from splitting the original dataset into subsets based on the values of the predictor $x$.
*ID3 Algorithm*
The ID3 (Iterative Dichotomizer) algorithm selects a decision tree using a top-down greedy search strategy through the space of possible decision trees, i.e., the set or space of all possible trees that can be arranged by assigning the predictors to nodes in all possible combinations.
The greedy strategy in ID3 can be summarized as below:
- Give the set of unselected predictors, select the predictor with the highest Information Gain (IG)
- Divide the dataset, i.e., the set of instances, into subsets based on the values of the selected attribute
- Repeat the above process for each of the subsets until the subset is empty or the subset is homogeneous (contains instances belonging to on class), or no more predictors are available.
We will illustrate the ID3 algorithm on the Iris data set next.
## Iris Example
We will now exemplify how to build a decision tree for the Iris dataset.
The goal is to classify each instance in the dataset using the predictors corresponding to each such instance.
| Variable | Type | Description |
|:-------------|:---------|:-----------------------|
| SepalLength | Ratio | the sepal length (cm) |
| SepalWidth | Ratio | the sepal width (cm) |
| PetalLength | Ratio | the petal length (cm) |
| PetalWidth | Ratio | the petal width (cm) |
| Species | Nominal | the flower species |
<div style="text-align:center;font-size: smaller">
<b>Source:</b> This dataset was taken from the <a href="https://archive.ics.uci.edu/ml/datasets/iris">UCI Machine Learning Repository library
</a></div>
<br>
## Load data
Import `pandas` to work with dataframes:
- `import pandas as pd`
```
import pandas as pd
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="/%IFN5#t2uAm}E`8:KV:">pd</variable></variables><block type="importAs" id="_@5-r*j4E`}d?=DXLinf" x="16" y="10"><field name="libraryName">pandas</field><field name="libraryAlias" id="/%IFN5#t2uAm}E`8:KV:">pd</field></block></xml>
```
Load the dataframe with the `iris` dataset:
- Create variable `iris` and set it to `with pd do read_csv using "datasets/iris.csv"`
- `iris` (to display)
```
iris = pd.read_csv('datasets/iris.csv')
iris
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="@dNx?X?Zd[|UZ.PI?OJ2">iris</variable><variable id="/%IFN5#t2uAm}E`8:KV:">pd</variable></variables><block type="variables_set" id="gM*jw`FfIR3)8=g0iEB7" x="11" y="186"><field name="VAR" id="@dNx?X?Zd[|UZ.PI?OJ2">iris</field><value name="VALUE"><block type="varDoMethod" id="ny0sjvqTnn2B]K2za7Li"><field name="VAR" id="/%IFN5#t2uAm}E`8:KV:">pd</field><field name="MEMBER">read_csv</field><data>pd:read_csv</data><value name="INPUT"><block type="text" id="dfrpI5b@DHr+DQ:|@vpv"><field name="TEXT">datasets/iris.csv</field></block></value></block></value></block><block type="variables_get" id="dn{+Q#DO%lN;G_tFGJ#B" x="8" y="304"><field name="VAR" id="@dNx?X?Zd[|UZ.PI?OJ2">iris</field></block></xml>
```
## Model
### Prepare data
Separate the predictor variables (`X`) from the class label (`Y`):
- Set `X` to `with iris do drop using` a list containing
- freestyle `columns=['Species']`
- Set `Y` to `iris[` a list containing `'Species'` `]`
```
X = iris.drop(columns=['Species'])
Y = iris[['Species']]
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="_z/H/tTnOYS1Lla2~;*0">X</variable><variable id="o8#tp%!,_Okj2aVwX(DL">Y</variable><variable id="@dNx?X?Zd[|UZ.PI?OJ2">iris</variable></variables><block type="variables_set" id="M9g^9:hbZQ0Cb9jKq9V6" x="14" y="218"><field name="VAR" id="_z/H/tTnOYS1Lla2~;*0">X</field><value name="VALUE"><block type="varDoMethod" id="HF9HKI:lNqY;?[|G]m=P"><field name="VAR" id="@dNx?X?Zd[|UZ.PI?OJ2">iris</field><field name="MEMBER">drop</field><data>iris:drop</data><value name="INPUT"><block type="lists_create_with" id="Mm^0W8}3,2|YO6m+rQ1Z"><mutation items="1"></mutation><value name="ADD0"><block type="dummyOutputCodeBlock" id="WkdEs~{tWz~8u4:XkYHO"><field name="CODE">columns=['Species']</field></block></value></block></value></block></value><next><block type="variables_set" id=":eCC|=CT/c:_fzcI.h5%"><field name="VAR" id="o8#tp%!,_Okj2aVwX(DL">Y</field><value name="VALUE"><block type="indexer" id="i*OWk(?KoBxj{6v-G8RH"><field name="VAR" id="@dNx?X?Zd[|UZ.PI?OJ2">iris</field><value name="INDEX"><block type="lists_create_with" id="H3L8XKx]~iB1,]H3b2H("><mutation items="1"></mutation><value name="ADD0"><block type="text" id="64q(iYfgFHZZK4?N%T@y"><field name="TEXT">Species</field></block></value></block></value></block></value></block></next></block></xml>
```
Split the data into train/test sets:
- `import sklearn.model_selection as model_selection`
```
import sklearn.model_selection as model_selection
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="-q~R*yx.VGQ@%!0Q+!gh">model_selection</variable></variables><block type="importAs" id="E{QJ$O@lc8u.;OTWdVgf" x="-89" y="168"><field name="libraryName">sklearn.model_selection</field><field name="libraryAlias" id="-q~R*yx.VGQ@%!0Q+!gh">model_selection</field></block></xml>
```
And do the actual split:
- Create `splits` and set to `with model_selection do train_test_split using` a list containing:
- `X` (the features in an array)
- `Y` (the labels in an array)
- freestyle `random_state=1` (this will make your random split the same as mine)
```
splits = model_selection.train_test_split(X, Y, random_state=1)
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="{Yt*9=2tTC$JepE9BcRz">splits</variable><variable id="-q~R*yx.VGQ@%!0Q+!gh">model_selection</variable><variable id="_z/H/tTnOYS1Lla2~;*0">X</variable><variable id="o8#tp%!,_Okj2aVwX(DL">Y</variable></variables><block type="variables_set" id="HpD!.HSC`PPRg]21i*7c" x="-102" y="134"><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field><value name="VALUE"><block type="varDoMethod" id="fUetl3$Ak=SI~T6T5!6c"><field name="VAR" id="-q~R*yx.VGQ@%!0Q+!gh">model_selection</field><field name="MEMBER">train_test_split</field><data>model_selection:train_test_split</data><value name="INPUT"><block type="lists_create_with" id="uUY%r2L/7acQcS/KbQvx"><mutation items="3"></mutation><value name="ADD0"><block type="variables_get" id="jN^;NIVaV%22*W_^|!uK"><field name="VAR" id="_z/H/tTnOYS1Lla2~;*0">X</field></block></value><value name="ADD1"><block type="variables_get" id="OOZSj;YDm{T6q%{~5?B1"><field name="VAR" id="o8#tp%!,_Okj2aVwX(DL">Y</field></block></value><value name="ADD2"><block type="dummyOutputCodeBlock" id="Gg*^%q4qIJwc*of,V|rb"><field name="CODE">random_state=1</field></block></value></block></value></block></value></block></xml>
```
### Fit model
First create the model:
- Create variable `decisionTree` and set to `with tree create DecisionTreeClassifier using`
```
decisionTree = tree.DecisionTreeClassifier()
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="fYtGc,k9LOv^%_9|[cfE">decisionTree</variable><variable id="tkeU~z;dQ]^6[?rbF?)9">tree</variable></variables><block type="variables_set" id="0{MIimjRn(`+`5M$0_U7" x="-90" y="132"><field name="VAR" id="fYtGc,k9LOv^%_9|[cfE">decisionTree</field><value name="VALUE"><block type="varCreateObject" id="?`dwaKnOBjnrd}`doxhF"><field name="VAR" id="tkeU~z;dQ]^6[?rbF?)9">tree</field><field name="MEMBER">DecisionTreeClassifier</field><data>tree:DecisionTreeClassifier</data></block></value></block></xml>
```
Fit it, and get predictions:
- `with decisionTree do fit` a list containing:
- `in list splits get # 1` (this is Xtrain)
- `in list splits get # 3` (this is Ytrain)
- Create variable `predictions` and set to `with decisionTree do predict using` a list containing:
- `in list splits get # 2` (this is Xtest)
```
decisionTree.fit(splits[0], splits[2])
predictions = decisionTree.predict(splits[1])
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="fYtGc,k9LOv^%_9|[cfE">decisionTree</variable><variable id=".hU+?FvKPIm~!dS$d-nj">predictions</variable><variable id="{Yt*9=2tTC$JepE9BcRz">splits</variable></variables><block type="varDoMethod" id="-@fm,911cJiCu@jkys%R" x="-94" y="192"><field name="VAR" id="fYtGc,k9LOv^%_9|[cfE">decisionTree</field><field name="MEMBER">fit</field><data>decisionTree:fit</data><value name="INPUT"><block type="lists_create_with" id="qj[~!aNmN}=v;xQ.=$%L"><mutation items="2"></mutation><value name="ADD0"><block type="lists_getIndex" id="W~.ZU([-)(,-i3=bTsB|"><mutation statement="false" at="true"></mutation><field name="MODE">GET</field><field name="WHERE">FROM_START</field><value name="VALUE"><block type="variables_get" id="GIJn[jpD_~wa}#7IO!1K"><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field></block></value><value name="AT"><block type="math_number" id="Rh[;nrMLN$[cFFsZC~T|"><field name="NUM">1</field></block></value></block></value><value name="ADD1"><block type="lists_getIndex" id="YkCD57xF*SSN3[`_op{u"><mutation statement="false" at="true"></mutation><field name="MODE">GET</field><field name="WHERE">FROM_START</field><value name="VALUE"><block type="variables_get" id="M@?wn}aoHxJYan0=AE3$"><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field></block></value><value name="AT"><block type="math_number" id="jNUa2BdXPG?H8|.br*@e"><field name="NUM">3</field></block></value></block></value></block></value></block><block type="variables_set" id="W2#bH~%cZ$JYTYX@t`OJ" x="-89" y="291"><field name="VAR" id=".hU+?FvKPIm~!dS$d-nj">predictions</field><value name="VALUE"><block type="varDoMethod" id="uF}tgd}f*m!CoCE6Vnfc"><field name="VAR" id="fYtGc,k9LOv^%_9|[cfE">decisionTree</field><field name="MEMBER">predict</field><data>decisionTree:predict</data><value name="INPUT"><block type="lists_create_with" id="g4~yox0A5a]I|VUV:ff9"><mutation items="1"></mutation><value name="ADD0"><block type="lists_getIndex" id="D[QG~pcCO~@%*GRtEXGC"><mutation statement="false" at="true"></mutation><field name="MODE">GET</field><field name="WHERE">FROM_START</field><value name="VALUE"><block type="variables_get" id="k]*6kE_B2NOcRC4T2DMa"><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field></block></value><value name="AT"><block type="math_number" id="nwBf+AnSmiv.DN_wSPy("><field name="NUM">2</field></block></value></block></value></block></value></block></value></block></xml>
```
### Measure performance
Import the evaluation metrics:
- `import sklearn.metrics as metrics`
```
import sklearn.metrics as metrics
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="jpz]0=,hLYv~rN^#0dFO">metrics</variable></variables><block type="importAs" id="CEkPf]fPOl]|@Gl1lho[" x="-101" y="139"><field name="libraryName">sklearn.metrics</field><field name="libraryAlias" id="jpz]0=,hLYv~rN^#0dFO">metrics</field></block></xml>
```
Get the accuracy:
- `with metrics do accuracy_score using` a list containing
- `in list splits get # 4` (this is `Ytest`)
- `predictions`
```
print(metrics.accuracy_score(splits[3], predictions))
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="jpz]0=,hLYv~rN^#0dFO">metrics</variable><variable id=".hU+?FvKPIm~!dS$d-nj">predictions</variable><variable id="{Yt*9=2tTC$JepE9BcRz">splits</variable></variables><block type="text_print" id="u8M]eo-vRBhX!L/a@4+J" x="-102" y="208"><value name="TEXT"><shadow type="text" id=")gVsrXo+bq|?9m0cIU/Y"><field name="TEXT">abc</field></shadow><block type="varDoMethod" id="0B;?~qgT4?Xe8I|O=P)%"><field name="VAR" id="jpz]0=,hLYv~rN^#0dFO">metrics</field><field name="MEMBER">accuracy_score</field><data>metrics:</data><value name="INPUT"><block type="lists_create_with" id="qe7YbU#WU/F|Iuks0C:c"><mutation items="2"></mutation><value name="ADD0"><block type="lists_getIndex" id="13J^-{z`P=,aau/j:VF~"><mutation statement="false" at="true"></mutation><field name="MODE">GET</field><field name="WHERE">FROM_START</field><value name="VALUE"><block type="variables_get" id="m^Z6y-8@L:s!oS)`%iJW"><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field></block></value><value name="AT"><block type="math_number" id="u}VxLP~*Dbl0g4[AO,bL"><field name="NUM">4</field></block></value></block></value><value name="ADD1"><block type="variables_get" id="|3h,TPit9wg+a;bR[r$P"><field name="VAR" id=".hU+?FvKPIm~!dS$d-nj">predictions</field></block></value></block></value></block></value></block></xml>
```
And get the recall and precision:
- `print with metrics do classification_report using` a list containing
- `in list splits get # 4` (this is `Ytest`)
- `predictions`
```
print(metrics.classification_report(splits[3], predictions))
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="jpz]0=,hLYv~rN^#0dFO">metrics</variable><variable id=".hU+?FvKPIm~!dS$d-nj">predictions</variable><variable id="{Yt*9=2tTC$JepE9BcRz">splits</variable></variables><block type="text_print" id="jFo8knaDn;JRZ=Fj7Tzo" x="57" y="316"><value name="TEXT"><shadow type="text" id="QWuxwm12CzEZ-r2sdwCR"><field name="TEXT">abc</field></shadow><block type="varDoMethod" id="(y`#h=AgNH6X+ahL$2L$"><field name="VAR" id="jpz]0=,hLYv~rN^#0dFO">metrics</field><field name="MEMBER">classification_report</field><data>metrics:classification_report</data><value name="INPUT"><block type="lists_create_with" id="A/#8KhfiGd[(@p,uaFFl"><mutation items="2"></mutation><value name="ADD0"><block type="lists_getIndex" id="*N[l.zzXc*IdHJ-Ur0D)"><mutation statement="false" at="true"></mutation><field name="MODE">GET</field><field name="WHERE">FROM_START</field><value name="VALUE"><block type="variables_get" id="fQYj?Tz[y/t|%9~{2Pf["><field name="VAR" id="{Yt*9=2tTC$JepE9BcRz">splits</field></block></value><value name="AT"><block type="math_number" id="V(5k43YOsHPep%LEEd9Q"><field name="NUM">4</field></block></value></block></value><value name="ADD1"><block type="variables_get" id="mX(20KX!C)d_,/ddeysr"><field name="VAR" id=".hU+?FvKPIm~!dS$d-nj">predictions</field></block></value></block></value></block></value></block></xml>
```
As we can see, both the accuracy and the average precision, recall, and f1 are all very good.
## Display the Tree
Copy the blocks above and the necessary changes:
- Create `dot_data` and set to `with tree do export_graphviz using` a list containing
- `decisionTree`
- freestyle `out_file=None`
- freestyle `feature_names=["SepalLength","SepalWidth","PetalLength","PetalWidth"]`
- freestyle `class_names=['setosa','versicolor','virginica']`
- freestyle `filled=True`
- freestyle `rounded=True`
- freestyle `special_characters=True`
- `with graphviz create Source using` a list containing `dot_data`
**To get the feature names, we just used the columns from the data frame displayed above
To get the class names, we copied the names in the classification report above.
It is important that both of these lists be in the right order.**
```
dot_data = tree.export_graphviz(decisionTree, out_file=None, feature_names=["SepalLength","SepalWidth","PetalLength","PetalWidth"], class_names=['setosa','versicolor','virginica'], filled=True, rounded=True, special_characters=True)
graphviz.Source(dot_data)
#<xml xmlns="https://developers.google.com/blockly/xml"><variables><variable id="F.ej=Hud7vT9uozVaF5C">dot_data</variable><variable id="rrXqQ:9R(~X0,}F[1UD]">graphviz</variable><variable id="tkeU~z;dQ]^6[?rbF?)9">tree</variable><variable id="fYtGc,k9LOv^%_9|[cfE">decisionTree</variable></variables><block type="variables_set" id=":6*zm74#9$!O3?QVUukE" x="209" y="280"><field name="VAR" id="F.ej=Hud7vT9uozVaF5C">dot_data</field><value name="VALUE"><block type="varDoMethod" id="jsRyzb,9t~G^G@G@SMkS"><field name="VAR" id="tkeU~z;dQ]^6[?rbF?)9">tree</field><field name="MEMBER">export_graphviz</field><data>dot_data:</data><value name="INPUT"><block type="lists_create_with" id="r2_rY*ef1v:96w)e_B6K"><mutation items="7"></mutation><value name="ADD0"><block type="variables_get" id="x$]b$%X#}qjO@0H/!O|N"><field name="VAR" id="fYtGc,k9LOv^%_9|[cfE">decisionTree</field></block></value><value name="ADD1"><block type="dummyOutputCodeBlock" id="K!=:ty3DMPs#b^ziH=5p"><field name="CODE">out_file=None</field></block></value><value name="ADD2"><block type="dummyOutputCodeBlock" id="by]EM[Y$kMG~MfF0{q^,"><field name="CODE">feature_names=["SepalLength","SepalWidth","PetalLength","PetalWidth"]</field></block></value><value name="ADD3"><block type="dummyOutputCodeBlock" id="2hz~E.{8:W9-wWb*9Ut~"><field name="CODE">class_names=['setosa','versicolor','virginica']</field></block></value><value name="ADD4"><block type="dummyOutputCodeBlock" id="^IJ3b3Eg7K9k^6Of_4i%"><field name="CODE">filled=True</field></block></value><value name="ADD5"><block type="dummyOutputCodeBlock" id="VH!NxdBRgItjnN]Zs^Uh"><field name="CODE">rounded=True</field></block></value><value name="ADD6"><block type="dummyOutputCodeBlock" id="f*|7fCy,K/0{_WzfribG"><field name="CODE">special_characters=True</field></block></value></block></value></block></value></block><block type="varCreateObject" id="-a(nDg:Sgtf)Ey5vB4(x" x="187" y="513"><field name="VAR" id="rrXqQ:9R(~X0,}F[1UD]">graphviz</field><field name="MEMBER">Source</field><data>graphviz:Source</data><value name="INPUT"><block type="variables_get" id="!M}@xBlLyRkl~Us)Smh%"><field name="VAR" id="F.ej=Hud7vT9uozVaF5C">dot_data</field></block></value></block></xml>
```
The learning algorithm decided that `PetalWidth` was the most important variable (or feature).
If the value of `PetalWidth` is less than .8, the iris will be classified as `setosa`.
Notice that all 37 `setosa` are classified by this one decision!
For `PetalWidth` ≥ .8 but &le 1.65 AND has `PetalLength` ≤ 4.95 most iris are `versicolor` (32 of 34).
We can see some complexity here with `versicolor` that don't quite fit as nicely lower in the tree.
Everything else, more or less, is `virginica`.
| github_jupyter |
# Experiment #1 - Baseline Model vs. Baseline ML Models
## Overview
The purpose of this experiment is to establish a baseline for a domain-driven model and to compare it to more sophisticated machine learning models using baseline features. Without collaboration with stakeholders or additional predictors such as customer tenure, lifetime value, historical response rate, etc., our baseline model will simply follow our intuited rule:
> Include the customer if they responded positively to the previous marketing campaign. If there were not enough respondents to use up the entire budget then randomly sample the training set for additional instances to ensure that the maximum number of customers are included.
To estimate the performance of machine learning models, we will train the following models with three different hyperparameter configurations, selecting the best configuration and averaging the scores of the best models:
* Naive Bayes
* Decision tree
* Logististic regression
* Neural network
* SVM
Scores will be based on how well a classifier can prioritize 6.6% of the population since that is all our budget allows for when deploying a model to classify all 100,000 customers.
```
%load_ext autoreload
%autoreload 2
from utils import code
import pandas as pd
import numpy as np
import scikitplot as skplt
# Model evaluation
from sklearn.metrics import make_scorer, roc_auc_score, classification_report, confusion_matrix
from sklearn.model_selection import train_test_split, cross_val_predict
from support.evaluation import plot_learning_curve, evaluate_model
# Support
from support.pipeline import get_pipeline, get_feature_names_for_pipeline
from support.model import Model, build_tuned_model
from support.datasets import get_data
from support.experiments import experiment_1, get_scorer
from support import parameters as params
# Algos
from sklearn.naive_bayes import BernoulliNB
from sklearn.dummy import DummyClassifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
from sklearn.neural_network import MLPClassifier
%matplotlib inline
```
## Data
```
X, y = get_data('../data/train.csv')
X.head()
n_instances = len(X)
p_instances = y.sum() / len(y)
p_targeted = .066
n_targeted = int(n_instances*p_targeted)
print('Number of instances: {:,}'.format(n_instances))
print('Number of conversions {:,}'.format(y.sum()))
print('Conversion rate: {:.2f}%'.format(p_instances*100.))
print('6.6% of the population {:,}'.format(n_targeted))
print('Expected number of conversions targetting {:,} @ {:.2f}%: {:,}'.format(n_targeted, p_instances*100., int(p_instances * n_targeted)))
```
Create a validation set and train the model then score on the test set. The performance will be biased since we are using less training data.
```
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=.2, stratify=y, random_state=1)
n_targeted_test = int(len(X_test) * p_targeted)
```
## Baseline Model
As stated above, our baseline model is simply the rule "predict the customer will convert if they converted when contacted as part of the previous campaign". Whether the customer converted during the previous campaign is encoded in the attribute `poutcome` which can take on three values: success, failure, or nonexistent. We are assuming that 'nonexistent' means that the customer was not part of the previous campaign (see the exploratory notebook - Exploratory Questions).
```
# Setup costs and benefits
avg_revenue = params.AVG_REVENUE
avg_cost = params.AVG_COST
# Get all of the instances where the previous campaign was a success
X_test_success = X_test[X_test.poutcome == 'success']
# Calcuate how many more instances we need
n_rest = n_targeted_test - len(X_test_success)
# Randomly choose from the remaining instances
rest = X_test[~(X_test.index.isin(X_test_success.index))].sample(n=n_rest, random_state=1)
# Combine the targeted and random groups
baseline_targets = pd.concat([X_test_success, rest], axis=0)
baseline_ys = y_test.loc[baseline_targets.index]
baseline_outcomes = baseline_ys.apply(lambda x: avg_cost if x == 0 else avg_cost + avg_revenue)
assert(len(baseline_targets) == n_targeted_test)
# Create the random targets
random_targets = X_test.sample(n=n_targeted_test)
random_ys = y.loc[random_targets.index]
random_outcomes = random_ys.apply(lambda x: avg_cost if x == 0 else avg_cost + avg_revenue)
# Compute profit
random_profit = sum(random_outcomes)
baseline_profit = sum(baseline_outcomes)
print('Number of customers targeted: {:,}/{:,}\n'.format(len(baseline_targets), len(X_test)))
print('Conversion rate under random policy: {:.1f}%'.format(random_ys.sum() / len(random_ys)*100.))
print('Expected profit under random policy: ${:,}\n'.format(random_profit))
print('Conversion rate under baseline policy: {:.1}%'.format(baseline_ys.sum() / len(baseline_ys)*100.))
print('Expected profit under baseline policy: ${:,}'.format(baseline_profit))
print('Lift over random policy: {:.1f} or ${:,}'.format(baseline_profit / random_profit, baseline_profit - random_profit))
```
## ML Models
Optimize each model we're interested in evaluating then choose the best one and estimate financial impact.
```
scorer = get_scorer()
```
Apply feature computations
```
pipeline = experiment_1.get_pipeline()
ps = pipeline.fit_transform(X).shape
print('Instances: {:,}, Features: {}'.format(ps[0], ps[1]))
```
Configure and run the different algos
```
results = []
# Naive Bayes
param_grid = [{
'nb__alpha': [0, 0.01, 0.1, 1],
'nb__fit_prior': [True, False]
}]
# We need to use the categorical pipeline since BernoulliNB only handles
# categorical features
nb_pipeline = experiment_1.get_categorical_pipeline()
result = evaluate_model(X, y, 'nb', BernoulliNB(), param_grid, scorer, nb_pipeline)
results.append(result)
# Decision tree
param_grid = [{
'dt__criterion': ['gini', 'entropy'],
'dt__max_depth': [1, 50, 100],
'dt__min_samples_leaf': [5, 10]
}]
result = evaluate_model(X, y, 'dt', DecisionTreeClassifier(), param_grid, pipeline)
results.append(result)
# Logistic regression
param_grid = [{
'lr__C': [.001, .01, 1.],
'lr__penalty': ['l1', 'l2']
}]
result = evaluate_model(X, y, 'lr', LogisticRegression(), param_grid, pipeline)
results.append(result)
# NN
param_grid = [{
'nn__hidden_layer_sizes': [(10), (100), (200)],
# 'nn__activation': ['logistic', 'tanh', 'relu'],
# 'nn__solver': ['lbfgs']
}]
result = evaluate_model(X, y, 'nn', MLPClassifier(), param_grid, pipeline)
results.append(result)
# SVM
param_grid = [{
'svm__C': [.001, .01, 1.],
'svm__gamma': [.1, 1., 2.],
'svm__kernel': ['rbf']
}]
result = evaluate_model(X, y, 'svm', SVC(), param_grid, pipeline)
results.append(result)
pd.DataFrame.from_dict(list(map(lambda x: { 'model': x[1], 'mean': x[2], 'std': x[3] }, results)))[[
'model', 'mean', 'std'
]].sort_values('mean', ascending=False).style.bar()
```
## Financial Impact of the Best Model
So now that we have evaluated models and estimated their generalization accuracy, we will select the best model and take a closer look at its performance and what we can expect in terms of profitability for targeting 6.6% of the population.
```
model_result = list(filter(lambda x: x[1] == 'nb', results))[0]
model = model_result[0]
model.model
print('Best model performance mean:', model_result[2])
print('Best model performance std:', model_result[3])
plot_learning_curve(model.model, 'Naive Bayes Learning Curves', model.pipeline.fit_transform(X), y, cv=5, scoring=scorer);
```
The model may be suffering from high bias - we could improve the performance - but we don't have to be concerned with variance (our model will generalize well).
Run 5-fold cross validation using the optimized model and print the metrics
```
preds = cross_val_predict(model.get_model_pipeline(), X, y, cv=5)
print(classification_report(y, preds))
```
While our model isn't very precise (we are correct in predicting a conversion 20% of the time, 80% of the time we get a false positive), it does a very good job (compared to random guessing) at finding customers that are likely to convert.
```
model.pipeline.fit(X_train)
model.model.fit(model.pipeline.transform(X_train), y_train)
probs = model.model.predict_proba(model.pipeline.transform(X_test))
preds = model.model.predict(model.pipeline.transform(X_test))
_ = skplt.metrics.cumulative_gain_curve(y_test, probs);
```
If we wanted to reduce the number of false positives, we could choose a probability threshold that corresponds to a lower (FPR, TPR) point on the ROC curve but since our profit margin is so high (assuming that our cost and revenue estimatios are correct), then we can afford the misclassifications so long as the classifier does a good job of selecting customers who end up converting.
Evaluate financial performance
```
# Create a dataframe of probabilities and actual / predicted outcomes
probs_df = pd.DataFrame(np.hstack([probs, y_test.values.reshape(-1,1), preds.reshape(-1,1)]), columns=['p_no', 'p_yes', 'actual', 'predicted'])
# Sort customers by the probability that they will convert
model_targets = probs_df.sort_values('p_yes', ascending=False)
# Take the top 6.6%
model_targets = model_targets.head(n_targeted_test)
# Calculate financial outcomes
model_outcomes = model_targets.actual.apply(lambda x: avg_cost if x == 0 else avg_cost + avg_revenue)
```
Model performance for top 6.6%
```
print(classification_report(model_targets.actual, model_targets.predicted))
# Calculate profit
model_profit = sum(model_outcomes)
print('Number of customers targeted: {:,}/{:,}'.format(len(model_targets), len(X_test)))
print('Conversion rate of model policy: {:.2f}%'.format(model_targets.actual.sum() / len(model_outcomes)*100.))
print('Expected profit of model policy: ${:,}'.format(model_profit))
print('Lift over random: {:.1f} or ${:,}'.format(model_profit / random_profit, model_profit - random_profit))
print('Lift over baseline: {:.1f} or ${:,}'.format(model_profit / baseline_profit, model_profit - baseline_profit))
```
Save the model
```
model.save('../models/experiment-1-model.pkl')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Edward-TL/amazon_scraper/blob/master/Backend_load.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
import requests
url = 'http://34.95.224.171/api'
json = { 'query' : 'mutation { getProducts { name } }' }
# r = requests.post(url=url, json=json)
# print (r.text)
from google.colab import drive
drive.mount('/content/drive')
#For the system
import os
import progressbar
from tabulate import tabulate
#Manage of time
from datetime import datetime, timedelta
from pytz import timezone
import time
import re
#Manage of files
import pandas as pd
import csv
#scrap
from bs4 import BeautifulSoup
from openpyxl.workbook import Workbook
import requests
%cd 'drive/My Drive/Colab Notebooks/4SS/4SS_db/testing/'
!ls
%cd 'Products/mx/parquet'
!ls
df = pd.read_parquet('mx-master_db_amazon-devices.parquet')
df.drop(columns=['Image ID (product page)', 'Product Description', ' Product Bullets', 'Principal Review - Title', 'Principal Review - Body', 'Principal Review - Stars'], inplace=True)
df
countries = ['mx', 'br']
#On the file
product_name = df['Product Names'].to_list()
date = df['time'].to_list()
rank = df['Rank'].to_list()
stars = df['Stars'].to_list()
reviews = df['Reviews'].to_list()
price = df['Price_std_or_min'].to_list()
max_price = df['Max_prices'].to_list()
#images ID to URL
image_id = df['Image ID'].to_list()
image_url = [f'https://images-na.ssl-images-amazon.com/images/I/{img_id}.jpg' for img_id in image_id]
#Static and quasi-static properties
store = 'Amazon'
#Country
country = countries[0]
country = country
#Product ID to URL
product_id = df['Product ID'].to_list()
product_url = [f'https://www.amazon.com.{country}/dp/{pid}' for pid in product_id]
#Constant
parquet_len = len('.parquet')
#variables
#Main Name Extracter
file_name_len = len(df_name)-parquet_len
file_name = df_name[13:file_name_len]
#Dictionary
category = file_name
string_1 = '''mutation {
releaseProduct (input: {'''
string_3 = '''}) {
_id
name
stars
}
}'''
query_json = string_1 + query_string + string_3
for n in product_name:
query_string = f'''
name : "{product_name[n]}",
imageURL: "{image_url[n]}",
link: "{product_url[n]}",
cathegory: "{category}",
country: "{country}",
store: "Amazon",
Time: "{date[n]}",
Rank: {int(rank[n])},
Price : {price[n]},
'''
url = 'http://34.95.224.171/api'
json = { 'query' : query_json}
r = requests.post(url=url, json=json)
print(query_json)
url = 'http://34.95.224.171/api'
json = { 'query' : query_json}
r = requests.post(url=url, json=json)
print (r.text)
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
import matplotlib as mpl
import pandas as pd
import sys
import datetime
from pathlib import Path
sys.path.append("../")
sys.path.append("../../")
sys.path.append("./analysis")
sys.path.append("./analysis/technical-analysis_python")
mpl.use('tkagg') # issues with Big Sur
import matplotlib.pyplot as plt
from strategy.macd_crossover import macdCrossover
from strategy.relative_strength_index import rsi
from strategy.stochastic_oscillator import stc_oscillator
from strategy.parabolic_stop_and_reverse import ParabolicSAR
from strategy.williams_R import williamsR
from backtest import Backtest
from evaluate import PortfolioReturn, SharpeRatio, MaxDrawdown, CAGR
import warnings
warnings.filterwarnings('ignore')
directory_name = "hkex_ticks_day"
pathlist = Path(directory_name).rglob('*.csv')
# traverse over files in directory
for path in pathlist:
# load data
df = pd.read_csv(path, header=0, index_col='Date', parse_dates=True)
# select time range
df = df.loc[pd.Timestamp('2017-01-01'):pd.Timestamp('2019-01-01')]
path_str = str(path)
ticker = path_str[len(directory_name)+6:len(directory_name)+6+4]
# MACD
"""
macd_cross = macdCrossover(df)
macd_fig = macd_cross.plot_MACD()
mpl.pyplot.close()
signals = macd_cross.gen_signals()
signal_fig = macd_cross.plot_signals(signals)
mpl.pyplot.close()
signal_fig = macd_cross.plot_signals_MACD()
mpl.pyplot.close()
"""
# RSI
"""
rsi = rsi(df)
rsi_fig = rsi.plot_RSI()
mpl.pyplot.close()
signals = rsi.gen_signals()
signal_fig = rsi.plot_signals(signals)
mpl.pyplot.close()
"""
# Stochastic Oscillator
stc = stc_oscillator(df)
stc_fig = stc.plot_KD()
signals = stc.gen_signals()
signal_fig = stc.plot_signals(signals)
# Parabolic SAR
psar = ParabolicSAR(df)
psar_fig = psar.plot_PSAR()
signals = psar.gen_signals()
signal_fig = psar.plot_signals(signals)
# William's R
wr = williamsR(df)
wr_fig = wr.plot_wr()
signals = wr.gen_signals()
signal_fig = wr.plot_signals(signals)
# Chaikin Oscillator
co = co(df)
co_fig = co.plot_CO()
signals = co.gen_signals()
signal_fig = co.plot_signals(signals)
# Volume Rate of Change
vroc = volume_roc(df)
vroc_fig = vroc.plot_VROC()
signals = vroc.gen_signals()
signal_fig = vroc.plot_signals(signals)
# Backtesting
portfolio, backtest_fig = Backtest(ticker, signals, df)
mpl.pyplot.close()
if (len(portfolio) != 0):
print("Final total value: {value:.4f} ".format(value = portfolio['total'][-1]))
print("Total return: {value:.4f}%".format(value = (portfolio['total'][-1] - portfolio['total'][0])/portfolio['total'][-1]*100))
# for analysis
print("No. of trade: {value}".format(value = len(signals[signals.positions == 1])))
filename = "test.txt"
with open(filename,'a') as outfile:
port_ret = (portfolio['total'][-1] - portfolio['total'][0])/portfolio['total'][-1]*100
num_trade = len(signals[signals.positions == 1])
start_date = pd.Timestamp('2017-01-01').strftime("%Y-%m-%d")
end_date = pd.Timestamp('2019-06-01').strftime("%Y-%m-%d")
outfile.write("{},{},{},{},{}".format(ticker, start_date, end_date, port_ret, num_trade))
outfile.write('\n')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/DaraSamii/DataDays2021/blob/main/train_mix_model.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Dara R Samii Login
```
from google.colab import drive
drive.mount("/content/drive",force_remount=True)
%cd /content/drive/MyDrive/DataDays2021/
pwd = %pwd
pwd
```
**-------------------------------------------------------------------------------------------------------------------------------**
# installing packages
```
!pip install "dask[complete]"
!pip install -Uqq fastai
!pip install parsivar
```
# Imports
```
import dask.dataframe as dd
import pandas as pd
import os
from fastai.text.all import *
from fastai.tabular.all import *
import pickle
from torch.utils.data import Dataset
from tqdm import tqdm
import torch as T
from fastai.data.core import DataLoaders
import torch.nn as nn
from torch.utils.data.dataloader import DataLoader
from helper import utils
```
# declaring paths
```
data_folder = os.path.join(pwd,"data")
final_clicked = os.path.join(data_folder,"final","final_clicked.csv")
final_products = os.path.join(data_folder,"final","final_products.csv")
mix_model_path = os.path.join(data_folder,"models","mix_model")
category_classifier_path = os.path.join(data_folder,"models","category_classifier")
cdf = dd.read_csv(final_clicked)
cdf.head()
```
## loading vocab
```
#if vocab exits:
vocab = pickle.load(open(os.path.join(category_classifier_path,"vocab"),'rb'))
vocab
```
## nlp trasnforms pipeline
```
class nlp_pipeline:
def __init__(self, vocab,):
self.vocab = vocab
self.tok = SpacyTokenizer(lang='fa')
self.num = Numericalize(vocab=self.vocab)
def encode(self,x):
x = utils._normalize_text(x)
x = tokenize1(x, self.tok)
x = self.num.encodes(x)
return x
def decode(self,x):
x = self.num.decodes(x)
x = " ".join(x)
return x
drop_cols =["Unnamed: 0",
"product_showed",
"DAY(datetime)",
"HOUR(datetime)",
"IS_WEEKEND(datetime)",
"MINUTE(datetime)",
"MONTH(datetime)",
"WEEKDAY(datetime)",
"YEAR(datetime)",
"_id"]
cont_cols = ["rank",
"NUM_WORDS(raw_query)",
"page",
"products.sellers_count",
"products.availabilty_ratio",
"products.mean_all_price",
"products.max_all_price",
"products.min_all_price",
"products.std_all_price",
"products.skew_all_price",
"products.mean_available_price",
"products.max_available_price",
"products.min_available_price",
"products.std_available_price",
"products.skew_available_price",
"products.COUNT(clicked_merged)",
"products.PERCENT_TRUE(clicked_merged.is_clicked)",
"products.NUM_WORDS(product_name_normalized)"]
cat_cols = ['products.DAY(first_added_date)',
'products.DAY(last_added_date)',
'products.HOUR(first_added_date)',
'products.HOUR(last_added_date)',
'products.IS_WEEKEND(first_added_date)',
'products.IS_WEEKEND(last_added_date)',
'products.MINUTE(first_added_date)',
'products.MINUTE(last_added_date)',
'products.MONTH(first_added_date)',
'products.MONTH(last_added_date)',
'products.WEEKDAY(first_added_date)',
'products.WEEKDAY(last_added_date)',
'products.YEAR(first_added_date)',
'products.YEAR(last_added_date)']
text_col = ['raw_query',
'products.category_name',
'products.product_name_normalized',]
target = 'is_clicked'
df = cdf.drop(drop_cols,axis=1).compute()
df
class MixDataSet(Dataset):
def __init__(self, df, cat_col_names, cont_col_names, query_col_name, category_col_name, product_col_name, vocab,target_col_name=None, test=False,normalize=True,):
self.df = df.reset_index(drop=True).copy()
self.cat_col_names = cat_col_names
self.cont_col_names = cont_col_names
self.query_col_name = query_col_name
self.category_col_name = category_col_name
self.product_col_name = product_col_name
self.target_col_name = target_col_name
self.test = test
self.normalize = normalize
self.vocab = vocab
self.nlp_pipeline = nlp_pipeline(vocab=self.vocab)
if self.test == False:
self.target_col_name = target_col_name
def __len__(self):
return self.df.shape[0]
def __getitem__(self, i):
ndf = self.df.iloc[i]
ndf = ndf.fillna(0.0,inplace=False)
if self.normalize == True:
if type(ndf) == pd.core.series.Series:
for col in ndf.index:
if "price" in col:
ndf[col] = np.log10(ndf[col]/1000 + 1)
elif "NUM" in col or "COUNT" in col:
ndf[col] = np.log10(ndf[col] + 1)
elif type(ndf) == pd.core.frame.DataFrame:
for col in ndf.columns:
if "price" in col:
ndf[col] = np.log10(ndf[col]/1000 + 1)
elif "NUM" in col or "COUNT" in col:
ndf[col] = np.log10(ndf[col] + 1)
for col in self.cat_col_names:
if "YEAR" in col:
ndf[col] = ndf[col] - 2017
cat = ndf[self.cat_col_names].values
cont = ndf[self.cont_col_names].values
query = self.nlp_pipeline.encode(ndf[self.query_col_name])
category = self.nlp_pipeline.encode(ndf[self.category_col_name])
product = self.nlp_pipeline.encode(ndf[self.product_col_name])
if self.test == False:
target = ndf[self.target_col_name]
return (T.tensor(cat.astype(np.int32)),T.tensor(cont.astype(np.float32)),query,category,product), T.tensor(target)
else:
return (T.tensor(cat.astype(np.int32)),T.tensor(cont.astype(np.float32)),query,category,product)
stratified_df = pd.concat([df[df["is_clicked"] == True].reset_index(drop=True)[0:10000],
df[df["is_clicked"] == False].reset_index(drop=True)[0:10000]]).reset_index(drop=True).sample(frac=1)
stratified_df.groupby(by="is_clicked").count()
b = MixDataSet(stratified_df,cat_cols,cont_cols,"raw_query","products.category_name","products.product_name_normalized",target_col_name="is_clicked",vocab=vocab[0])
b[0:10]
len(b)
stratified_df["is_valid"] = False
stratified_df["is_valid"] = stratified_df["is_valid"].apply(lambda x: True if random.random() < 0.08 else False)
def my_collate(batch):
b = list(zip(*batch))
x,y = b
x1,x2,x3,x4,x5 = list(zip(*x))
return (T.stack(x1),T.stack(x2), nn.utils.rnn.pad_sequence(x3).T, nn.utils.rnn.pad_sequence(x4).T, nn.utils.rnn.pad_sequence(x5).T), T.stack(y).to(T.long)
train_ds = MixDataSet(stratified_df[stratified_df["is_valid"]==False],
cat_cols,
cont_cols,
"raw_query",
"products.category_name",
"products.product_name_normalized",
target_col_name="is_clicked",
vocab=vocab[0])
valid_ds = MixDataSet(stratified_df[stratified_df["is_valid"]==True],
cat_cols,
cont_cols,
"raw_query",
"products.category_name",
"products.product_name_normalized",
target_col_name="is_clicked",
vocab = vocab[0])
len(train_ds), len(valid_ds)
train_dl = DataLoader(dataset=train_ds, batch_size=150, shuffle=True, collate_fn=my_collate)
valid_dl = DataLoader(dataset=valid_ds, batch_size=150, shuffle=True, collate_fn=my_collate)
next(iter(train_dl))
next(iter(valid_dl))
class MixModel(nn.Module):
def __init__(self, embed_sz, n_cont, layers, awd_config, vocab_sz, lin_ftrs,joint_layers):
super(MixModel, self).__init__()
self.embed_sz = embed_sz
self.n_cont = n_cont
self.layers = layers
self.awd_config = awd_config
self.joint_layers = joint_layers
self.tab_model = TabularModel(emb_szs=self.embed_sz, n_cont=n_cont, out_sz=2,layers=layers,)
self.tab_model.layers = self.tab_model.layers[:-1]
self.awd1 = get_text_classifier(AWD_LSTM,vocab_sz=vocab_sz, n_class=2, config=awd_config,lin_ftrs=lin_ftrs)
self.awd1[-1].layers = self.awd1[-1].layers[:-1]
self.awd2 = get_text_classifier(AWD_LSTM,vocab_sz=vocab_sz, n_class=2, config=awd_config,lin_ftrs=lin_ftrs)
self.awd2[-1].layers = self.awd2[-1].layers[:-1]
self.awd3 = get_text_classifier(AWD_LSTM,vocab_sz=vocab_sz, n_class=2, config=awd_config,lin_ftrs=lin_ftrs)
self.awd3[-1].layers = self.awd3[-1].layers[:-1]
self.joint_layers = [lin_ftrs[-1]*3 + layers[-1]] + joint_layers + [2]
linBins = []
for i in range(0,len(self.joint_layers)-1):
linBins.append(LinBnDrop(self.joint_layers[i],self.joint_layers[i+1]))
self.LinBins = nn.Sequential(*linBins)
self.Softmax = nn.Softmax(dim=1)
def forward(self, x):
xtab = self.tab_model(x[0],x[1])
xnlp1 = self.awd1(x[2])[0]
xnlp2 = self.awd2(x[3])[0]
xnlp3 = self.awd3(x[4])[0]
X = T.cat([xtab, xnlp1, xnlp2, xnlp3],dim=1)
X = self.LinBins(X)
return self.Softmax(X)
def reset(self):
self.awd1.reset()
self.awd2.reset()
self.awd3.reset()
for col in cat_cols:
print(col,"-->", df[col].nunique())
len(cont_cols)
emb_sz = [
(32, 20),
(32, 20),
(24, 20),
(24, 20),
(3, 10),
(3, 10),
(61, 30),
(61, 30),
(13, 10),
(13, 10),
(8, 10),
(8, 10),
(6, 20),
(6, 20)
]
awd_conf = {'bidir': True,
'emb_sz': 1000,
'embed_p': 0.05,
'hidden_p': 0.3,
'input_p': 0.4,
'n_hid': 1000,
'n_layers': 3,
'output_p': 0.4,
'pad_token': 1,
'weight_p': 0.5}
mix_model = MixModel(embed_sz = emb_sz, n_cont= len(cont_cols), layers=[200], awd_config=awd_conf, vocab_sz=len(vocab[0]), lin_ftrs=[500],joint_layers=[500])
state_dict = T.load(open(os.path.join(mix_model_path,"models","mix_model.pth"),'rb'))
mix_model.load_state_dict(state_dict)
mix_model
sum(p.numel() for p in mix_model.parameters())
a = next(iter(train_dl))
tab = mix_model.tab_model(a[0][0],a[0][1])
nlp1 = mix_model.awd1(a[0][2])
nlp2 = mix_model.awd2(a[0][3])
nlp3 = mix_model.awd3(a[0][4])
print(tab.shape, nlp1[0].shape, nlp2[0].shape, nlp3[0].shape)
mix_model(a[0])
fast_mix_dl = DataLoaders(train_dl,valid_dl)
loss_func = nn.CrossEntropyLoss()
learn = Learner(fast_mix_dl,
mix_model,
loss_func = loss_func,
path = mix_model_path,
metrics=[accuracy,error_rate,Recall(),Precision(),F1Score()]).to_fp16()
grp = ShowGraphCallback
svm = SaveModelCallback(at_end=False,every_epoch=False,reset_on_fit=False,monitor='f1_score',fname="mix_model",)
esc = EarlyStoppingCallback(patience=3)
rlr = ReduceLROnPlateau(monitor="valid_loss",patience=2,factor=10,)
learn.add_cbs([grp,svm,esc,rlr,ModelResetter])
learn.cbs
learn.lr_find()
learn.fine_tune(10,1e-04,10)
stratified_df = pd.concat([df[df["is_clicked"] == True].reset_index(drop=True)[0:30000],
df[df["is_clicked"] == False].reset_index(drop=True)[0:30000]]).reset_index(drop=True).sample(frac=1)
stratified_df["is_valid"] = False
stratified_df["is_valid"] = stratified_df["is_valid"].apply(lambda x: True if random.random() < 0.08 else False)
train_ds1 = MixDataSet(stratified_df[stratified_df["is_valid"]==False],
cat_cols,
cont_cols,
"raw_query",
"products.category_name",
"products.product_name_normalized",
target_col_name="is_clicked",
vocab=vocab[0])
valid_ds1 = MixDataSet(stratified_df[stratified_df["is_valid"]==True],
cat_cols,
cont_cols,
"raw_query",
"products.category_name",
"products.product_name_normalized",
target_col_name="is_clicked",
vocab = vocab[0])
train_dl1 = DataLoader(dataset=train_ds1, batch_size=200, shuffle=True, collate_fn=my_collate)
valid_dl1 = DataLoader(dataset=valid_ds1, batch_size=200, shuffle=True, collate_fn=my_collate)
fast_mix_dl1 = DataLoaders(train_dl1,valid_dl1)
loss_func = nn.CrossEntropyLoss()
learn = Learner(fast_mix_dl1,
mix_model,
loss_func = loss_func,
path = mix_model_path,
metrics=[accuracy,error_rate,Recall(),Precision(),F1Score()]).to_fp16()
grp = ShowGraphCallback
svm = SaveModelCallback(at_end=False,every_epoch=False,reset_on_fit=False,monitor='f1_score',fname="mix_model",)
esc = EarlyStoppingCallback(patience=3)
rlr = ReduceLROnPlateau(monitor="valid_loss",patience=2,factor=10,)
learn.add_cbs([grp,svm,esc,rlr,ModelResetter])
learn.cbs
learn.fine_tune(10,1e-04,10)
```
| github_jupyter |
```
import os
import glob
import sklearn
import numpy as np
import cv2 as cv
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
from matplotlib.colors import ListedColormap
from ana_classification import preprocess
sklearn.show_versions()
```
### Data loading and processing
```
files = glob.glob('data/original/*/*')
print(len(files))
files[:10]
negatives_files = [f for f in files if 'NEG' in f]
positives_files = [f for f in files if 'NEG' not in f]
len(negatives_files), len(positives_files)
%%time
neg_imgs = np.array([preprocess(cv.imread(f), normalize=False, equalize=False) for f in negatives_files])
pos_imgs = np.array([preprocess(cv.imread(f), normalize=False, equalize=False) for f in positives_files])
neg_imgs.shape, pos_imgs.shape
pos_means = np.mean(pos_imgs.reshape(pos_imgs.shape[0], -1), axis=1)
neg_means = np.mean(neg_imgs.reshape(neg_imgs.shape[0], -1), axis=1)
pos_stds = np.std(pos_imgs.reshape(pos_imgs.shape[0], -1), axis=1)
neg_stds = np.std(neg_imgs.reshape(neg_imgs.shape[0], -1), axis=1)
means = np.concatenate([neg_means, pos_means])
stds = np.concatenate([neg_stds, pos_stds])
x = np.column_stack([means, stds])
y = np.array([0] * len(neg_imgs) + [1] * len(pos_imgs))
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=1)
```
### Data visualization
```
f, a = plt.subplots(1, 2)
f.set_size_inches(15, 6)
for ax in a:
ax.scatter(neg_means, neg_stds, c='blue', s=3, label='negative samples')
ax.scatter(pos_means, pos_stds, c='red', s=3, label='positive samples')
ax.xaxis.set_label('mean pixel intentity')
ax.yaxis.set_label('pixel intensity std')
ax.legend()
a[1].set_xlim(-1, 15)
a[1].set_ylim(0, 10)
plt.show()
```
### Data saving
```
np.savez('neg-classifier-data.npz', x_train=x_train, y_train=y_train)
```
### Data loading
```
data = np.load('neg-classifier-data.npz')
x_train = data['x_train']
y_train = data['y_train']
```
### Classifier
```
classifier = SVC(C=0.9, kernel='poly', degree=2, gamma=0.35)
classifier.fit(x_train, y_train)
```
### Decision boundary
```
h = 0.05
cmap_light = ListedColormap(['#AAAAFF', '#FFAAAA'])
cmap_bold = ListedColormap(['#0000FF', '#FF0000'])
x_min, x_max = -1, 15
y_min, y_max = 0, 10
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
Z = classifier.predict(np.c_[xx.ravel(), yy.ravel()])
Z = Z.reshape(xx.shape)
plt.pcolormesh(xx, yy, Z, cmap=cmap_light)
plt.scatter(x[:, 0], x[:, 1], c=y, cmap=cmap_bold, s=4)
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.title('classifier decision boundary')
plt.xlabel('mean')
plt.ylabel('std')
plt.show()
```
### Results
```
train_preds = classifier.predict(x_train)
test_preds = classifier.predict(x_test)
train_acc = np.mean(train_preds == y_train)
test_acc = np.mean(test_preds == y_test)
print(f'train accuracy: {train_acc}')
print(f'test accuracy: {test_acc}')
```
| github_jupyter |
# 1D Heat Equation
Heat transfer in 1D is governed by the following PDE (See [here](https://ocw.mit.edu/courses/mathematics/18-303-linear-partial-differential-equations-fall-2006/lecture-notes/heateqni.pdf) for more info):
$\frac{\partial u}{\partial t} = \kappa \frac{\partial^2 u}{\partial x^2}$
$\kappa = K_0/c\rho$
$\hat{x}=x/L_*$
$\hat{t}=t/T_*$
$\hat{u}(\hat{x},\hat{t})=u(x,t)/U_*$
$\frac{\partial \hat{u}}{\partial \hat{t}} = \frac{T_* \kappa}{L^2_*} \frac{\partial^2 \hat{u}}{\partial \hat{x}^2}$
$T_* = L^2_*/ \kappa$
$\frac{\partial \hat{u}}{\partial \hat{t}} = \frac{\partial^2 \hat{u}}{\partial \hat{x}^2}$
Suppose the initial temperature distibution in a 1D rod is constant i.e. $f(x) = u_0$
$u(x,t) = \sum ^\inf _{n=1} B_n sin(n\pi x)e^{-n^2 \pi^2 t}$
where
$B_n = 2u_0 \int_0^1 sin(n \pi x) dx$
$u(x,t) = \frac{4u_0}{\pi} \sum_{n=1}^{\inf} \frac{sin((2n-1)\pi x)}{(2n-1)} e^{-(2n-1)^2\pi^2 t}$
$u(x,t) = \frac{4u_0}{\pi}\left( sin(\pi x)e^{-\pi^2 t} + \frac{sin(3\pi x)}{3}e^{-9\pi ^2 t} + ... \right)$
```
import numpy as np
np.random.seed(10)
np.set_printoptions(5)
import matplotlib.pyplot as plt
def approx_sol(u0, x, t, N):
out = 0
for i in range(N):
n = i+1
out += np.sin((2*n-1)*np.pi*x) * np.exp(-(2*n-1)**2 * np.pi**2 * t) / (2*n-1)
return (4*u0/np.pi)*out
# NBVAL_IGNORE_OUTPUT
u0 = 1.0
x = np.linspace(0, 1, 101)
plt.figure()
for t in [1/np.pi**2]:
sol = approx_sol(u0, x, t, N=10000)
plt.plot(x, sol)
print(f"Solution maximum: {sol.max():.5f}")
plt.show()
# NBVAL_IGNORE_OUTPUT
t = np.linspace(0, 1, 101)
plt.figure()
for x in [0.01, 0.25, 0.5, 0.75]:
plt.plot(t, approx_sol(u0, x, t, N=10000))
plt.show()
# Copper
K0 = 400
rho = 8960
Cp = 385
kappa = K0/(rho*Cp)
print(f"Thermal diffusivity (kappa): {kappa:.5e}")
l = 1.0
def approx_sol_dim(u0, x, t, N, kappa, l):
out = 0
for i in range(N):
n = i+1
out += np.sin((2*n-1)*np.pi*x/l) * np.exp(-(2*n-1)**2 * np.pi**2 * t * kappa/l**2) / (2*n-1)
return (4*u0/np.pi)*out
# NBVAL_IGNORE_OUTPUT
x = np.linspace(0, l, 101)
plt.figure()
t_diffusion = (1/np.pi**2)/(kappa/l**2)
print(f"Diffusion time: {t_diffusion/60:.5f} mins")
for t in np.linspace(0.5, 1.5, 3)*t_diffusion:
sol = approx_sol_dim(u0, x, t, 10000, kappa, l)
plt.plot(x, sol)
print(f"Solution maximum: {sol.max():.5f}")
plt.show()
import openpnm as op
ws = op.Workspace()
spacing = 1e-2
net = op.network.Cubic(shape=[101, 1, 1], spacing=spacing)
# translate to origin
net['pore.coords'] -= np.array([spacing, spacing, spacing])/2
l = net['pore.coords'][:, 0].max() - net['pore.coords'][:, 0].min()
print(f"Length: {l:.5f}")
geo = op.geometry.GenericGeometry(network=net, pores=net.Ps, throats=net.Ts)
geo['pore.diameter'] = spacing
geo['throat.diameter'] = spacing
geo['throat.length'] = spacing
geo['throat.area'] = spacing**2
geo['pore.area'] = spacing**2
geo['pore.volume'] = spacing**3
geo['throat.volume'] = 0.0
phase = op.phases.GenericPhase(network=net)
phase['pore.conductivity'] = kappa
phys = op.physics.GenericPhysics(network=net, geometry=geo, phase=phase)
c = 1.0 # mol/m^3
phys['throat.conductance'] = c*kappa*geo['throat.area']/geo['throat.length']
alg = op.algorithms.FourierConduction(network=net)
alg.setup(phase=phase, conductance='throat.conductance')
alg.set_value_BC(pores=[0], values=1.0)
alg.set_value_BC(pores=[-1], values=0.0)
alg.run()
K_eff = alg.calc_effective_conductivity(domain_length=l, domain_area=spacing**2)[0]
print(f"Effective conductivity: {K_eff:.5e}, kappa: {kappa:.5e}, do they match? {np.allclose(K_eff, kappa)}")
# NBVAL_IGNORE_OUTPUT
alg = op.algorithms.TransientReactiveTransport(network=net)
alg.setup(phase=phase, conductance='throat.conductance', quantity='pore.temperature',
t_initial=0, t_final=880, t_step=87, t_output=10,
t_tolerance=1e-12, t_precision=12, rxn_tolerance=1e-12, t_scheme='implicit')
alg.set_IC(values=u0)
alg.set_value_BC(pores=[0], values=0.0)
alg.set_value_BC(pores=[-1], values=0.0)
alg.run()
res = alg.results()
times = list(res.keys())
times.sort()
plt.figure()
for time in times:
plt.plot(alg[time])
print(f"Diffusion time: {t_diffusion:.5f} sec")
'pore.temperature@870' in times
# NBVAL_IGNORE_OUTPUT
plt.figure()
x = net['pore.coords'][:, 0]
a = alg['pore.temperature@870']
b = approx_sol_dim(u0, x, 870.0, 10000, kappa, l)
plt.plot(a)
plt.plot(b, '--')
plt.show()
print(f"Maximum error: {np.max(a-b):.5f}")
def approx_sol_inhom(u1, x, t, N):
out = 0
for i in range(N):
n = i+1
out += ((-1)**n)/n * np.sin(n*np.pi*x) * np.exp(-n**2 * np.pi**2 * t)
return u1*x + (2*u1/np.pi)*out
# NBVAL_IGNORE_OUTPUT
u1 = 1.0
x = np.linspace(0, 1, 101)
plt.figure()
for t in np.array([0.01, 0.1, 0.5, 1.0, 1.5, 3])/np.pi**2:
sol = approx_sol_inhom(u1, x, t, N=10000)
plt.plot(x, sol)
plt.show()
def approx_sol_inhom_dim(u1, x, t, N, kappa, l):
out = 0
for i in range(N):
n = i+1
out += ((-1)**n)/n * np.sin(n*np.pi*x/l) * np.exp(-n**2 * np.pi**2 * t * kappa/l**2)
return u1*x/l + (2*u1/np.pi)*out
# NBVAL_IGNORE_OUTPUT
alg = op.algorithms.TransientReactiveTransport(network=net)
alg.setup(phase=phase, conductance='throat.conductance', quantity='pore.temperature',
t_initial=0, t_final=870, t_step=87, t_output=10,
t_tolerance=1e-12, t_precision=12, rxn_tolerance=1e-12, t_scheme='implicit')
alg.set_IC(values=0.0)
alg.set_value_BC(pores=[0], values=0.0)
alg.set_value_BC(pores=[-1], values=u1)
alg.run()
res = alg.results()
times = list(res.keys())
times.sort()
plt.figure()
for time in times:
plt.plot(alg[time])
# NBVAL_IGNORE_OUTPUT
plt.figure()
x = net['pore.coords'][:, 0]
a = alg['pore.temperature@870']
b = approx_sol_inhom_dim(u0, x, 870.0, 10000, kappa, l)
plt.plot(a)
plt.plot(b, '--')
plt.show()
print(np.max(a-b))
```
Now consider a source term $sin(\pi x)$
$u_{t} = u_{xx} + sin(\pi x)$
subject to B.Cs
$u(0) = u(1) = 0$
```
def approx_sol_source(x, t):
return (np.sin(np.pi*x)/(np.pi**2))*(1-np.exp(-np.pi**2 * t))
def approx_sol_source_dim(x, t, kappa, l):
return (np.sin(np.pi*x/l)/(np.pi**2))*(1-np.exp(-np.pi**2 * t * kappa/l**2))
# NBVAL_IGNORE_OUTPUT
x = np.linspace(0, 1, 101)
plt.figure()
for t in np.array([0.01, 0.1, 0.5, 1.0, 1.5, 3, 100])/np.pi**2:
sol = approx_sol_source(x, t)
plt.plot(x, sol)
plt.show()
# NBVAL_IGNORE_OUTPUT
Qf = 2000*net['pore.volume']/(Cp*rho)
Q = Qf*np.sin(np.pi*x)
phys['pore.source.S1'] = 0.0
phys['pore.source.S2'] = Q
phys['pore.source.rate'] = Q
alg = op.algorithms.TransientReactiveTransport(network=net)
alg.setup(phase=phase, conductance='throat.conductance', quantity='pore.temperature',
t_initial=0, t_final=10000, t_step=100, t_output=100,
t_tolerance=1e-7, t_precision=12, rxn_tolerance=1e-7, t_scheme='implicit')
alg.set_IC(values=0.0)
alg.set_value_BC(pores=[0], values=0.0)
alg.set_value_BC(pores=[-1], values=0.0)
alg.set_source(propname='pore.source', pores=net.pores()[1:-1])
alg.run()
res = alg.results()
times = list(res.keys())
times.sort()
plt.figure()
for time in times:
plt.plot(alg[time])
print(f"Maximum temperature: {alg['pore.temperature@4500'].max():.5f}")
# NBVAL_IGNORE_OUTPUT
plt.figure()
x = net['pore.coords'][:, 0]
a = alg['pore.temperature@4500']
b = approx_sol_source_dim(x, 4500.0, kappa, l)
plt.plot(a)
plt.plot(b, '--')
plt.show()
print(f"Maximum error: {np.max(a-b):.5f}")
def crazy_source(x, t, N):
# https://www.math.upenn.edu/~deturck/m241/inhomogeneous.pdf
out = 0
for n in range(N):
a = 4*((2*n+1)**2 * np.pi**2 - 2)*np.exp(-(2*n+1)**2 * np.pi**2 * t)/((2*n+1)**3 * np.pi**3 * ((2*n+1)**2 *np.pi**2 - 1))
b = 4*np.exp(-t)/((2*n+1)*np.pi*((2*n+1)**2*np.pi**2-1))
c = np.sin((2*n+1)*np.pi*x)
out += (a+b)*c
return out
# NBVAL_IGNORE_OUTPUT
x = np.linspace(0, 1, 101)
plt.figure()
times = np.array([0.01, 0.1, 0.5, 1.0, 1.5, 3])/np.pi**2
for t in times:
sol = crazy_source(x, t, 10000)
plt.plot(x, sol)
plt.show()
# NBVAL_IGNORE_OUTPUT
plt.figure()
x = np.linspace(0, 2, 101)
plt.plot(1-np.abs(x-1))
plt.show()
# NBVAL_IGNORE_OUTPUT
plt.figure()
plt.plot(2*x-x**2)
plt.show()
def qn(n, x):
return (8/(n**2 * np.pi**2))*np.sin(n*np.pi*x/2)
def an(n, x):
return (4/(n**2 * np.pi**2))*qn(n, x)
def cn(n):
return (16/(n**3 * np.pi**3))*(1-np.cos(n*np.pi))
def bn(n, x):
return cn(n) - an(n, x)
def approx_sol_sawtooth(x, t, N):
#https://faculty.uca.edu/darrigo/Students/M4315/Fall%202005/sep-var.pdf
out = 0
for i in range(N):
n = i+1
np2 = (n*np.pi/2)
e = np.exp(-((np2**2) * t))
out += ( an(n, x) + bn(n, x) * e ) * np.sin(np2*x)
return out
# NBVAL_IGNORE_OUTPUT
for t in [0.0, 0.01, 0.1, 0.5, 1.0, 2.0, 3.0, 100.0]:
sol = approx_sol_sawtooth(x, t, 10000)
plt.plot(x, sol)
plt.show()
```
| github_jupyter |
# House Building with worker skills
This tutorial includes everything you need to set up decision optimization engines, build constraint programming models.
When you finish this tutorial, you'll have a foundational knowledge of _Prescriptive Analytics_.
>It requires either an [installation of CPLEX Optimizers](http://ibmdecisionoptimization.github.io/docplex-doc/getting_started.html) or it can be run on [IBM Cloud Pak for Data as a Service](https://www.ibm.com/products/cloud-pak-for-data/as-a-service/) (Sign up for a [free IBM Cloud account](https://dataplatform.cloud.ibm.com/registration/stepone?context=wdp&apps=all>)
and you can start using `IBM Cloud Pak for Data as a Service` right away).
>
> CPLEX is available on <i>IBM Cloud Pack for Data</i> and <i>IBM Cloud Pak for Data as a Service</i>:
> - <i>IBM Cloud Pak for Data as a Service</i>: Depends on the runtime used:
> - <i>Python 3.x</i> runtime: Community edition
> - <i>Python 3.x + DO</i> runtime: full edition
> - <i>Cloud Pack for Data</i>: Community edition is installed by default. Please install `DO` addon in `Watson Studio Premium` for the full edition
Table of contents:
- [Describe the business problem](#Describe-the-business-problem)
* [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
* [Use decision optimization](#Use-decision-optimization)
* [Step 1: Download the library](#Step-1:-Download-the-library)
* [Step 2: Set up the engines](#Step-2:-Set-up-the-prescriptive-engine)
- [Step 3: Model the Data](#Step-3:-Model-the-data)
- [Step 4: Set up the prescriptive model](#Step-4:-Set-up-the-prescriptive-model)
* [Define the decision variables](#Define-the-decision-variables)
* [Express the business constraints](#Express-the-business-constraints)
* [Express the objective](#Express-the-objective)
* [Solve with Decision Optimization solve service](#Solve-with-Decision-Optimization-solve-service)
* [Step 5: Investigate the solution and run an example analysis](#Step-5:-Investigate-the-solution-and-then-run-an-example-analysis)
* [Summary](#Summary)
****
### Describe the business problem
* This is a problem of building five houses in different locations; the masonry, roofing, painting, etc. must be scheduled. Some tasks must necessarily take place before others and these requirements are expressed through precedence constraints.
* There are three workers, and each worker has a given skill level for each task. Each task requires one worker; the worker assigned must have a non-null skill level for the task. A worker can be assigned to only one task at a time.
* Each house has a deadline.
* The objective is to maximize the skill levels of the workers assigned to the tasks.
*****
## How decision optimization can help
* Prescriptive analytics technology recommends actions based on desired outcomes, taking into account specific scenarios, resources, and knowledge of past and current events. This insight can help your organization make better decisions and have greater control of business outcomes.
* Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
* Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
<br/>
+ For example:
+ Automate complex decisions and trade-offs to better manage limited resources.
+ Take advantage of a future opportunity or mitigate a future risk.
+ Proactively update recommendations based on changing events.
+ Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
<h2>About Detailed Scheduling concepts</h2>
<p>
<ul>
<li> Scheduling consists of assigning starting and completion times to a set of activities while satisfying different types of constraints (resource availability, precedence relationships, … ) and optimizing some criteria (minimizing tardiness, …)
<!-- <img src = "./house_building_utils/activity.png" > -->
<img src = "https://github.com/IBMDecisionOptimization/docplex-examples/blob/master/examples/cp/jupyter/house_building_utils/activity.PNG?raw=true " >
<li> Time is considered as a continuous dimension: domain of possible start/completion times for an activity is potentially very large
<li>Beside start and completion times of activities, other types of decision variables are often involved in real industrial scheduling problems (resource allocation, optional activities …)
</ul>
## Use decision optimization
### Step 1: Download the library
Run the following code to install Decision Optimization CPLEX Modeling library. The *DOcplex* library contains the two modeling packages, Mathematical Programming and Constraint Programming, referred to earlier.
```
import sys
try:
import docplex.cp
except:
if hasattr(sys, 'real_prefix'):
#we are in a virtual env.
!pip install docplex
else:
!pip install --user docplex
```
Note that the more global package <i>docplex</i> contains another subpackage <i>docplex.mp</i> that is dedicated to Mathematical Programming, another branch of optimization.
### Step 2: Set up the prescriptive engine
For display of the solution, ensure last version of matplotlib is available:
```
try:
import matplotlib
if matplotlib.__version__ < "1.4.3":
!pip install --upgrade matplotlib
except:
!pip install --user matplotlib
```
Now, we need to import all required modeling functions that are provided by the <i>docplex.cp</i> package:
```
from docplex.cp.model import CpoModel
from sys import stdout
from collections import namedtuple
```
### Step 3: Model the data
Planning contains the number of houses and the max amount of periods (<i>days</i>) for our schedule
```
NB_HOUSES = 5
MAX_AMOUNT_OF_PERIODS = 318
HOUSES = range(1, NB_HOUSES + 1)
```
All tasks must start and end between 0 and the max amount of periods
```
period_domain = (0, MAX_AMOUNT_OF_PERIODS)
```
For each task type in the house building project, the following table shows the duration of the task in days along with the tasks that must be finished before the task can start. A worker can only work on one task at a time; each task, once started, may not be interrupted.
<p>
| *Task* | *Duration* | *Preceding tasks* |
|---|---|---|
| masonry | 35 | |
| carpentry | 15 | masonry |
| plumbing | 40 | masonry |
| ceiling | 15 | masonry |
| roofing | 5 | carpentry |
| painting | 10 | ceiling |
| windows | 5 | roofing |
| facade | 10 | roofing, plumbing |
| garden | 5 | roofing, plumbing |
| moving | 5 | windows, facade, garden, painting |
##### Tasks' durations
```
Task = (namedtuple("Task", ["name", "duration"]))
TASKS = {Task("masonry", 35),
Task("carpentry", 15),
Task("plumbing", 40),
Task("ceiling", 15),
Task("roofing", 5),
Task("painting", 10),
Task("windows", 5),
Task("facade", 10),
Task("garden", 5),
Task("moving", 5),
}
```
##### The tasks precedences
```
TaskPrecedence = (namedtuple("TaskPrecedence", ["beforeTask", "afterTask"]))
TASK_PRECEDENCES = {TaskPrecedence("masonry", "carpentry"),
TaskPrecedence("masonry", "plumbing"),
TaskPrecedence("masonry", "ceiling"),
TaskPrecedence("carpentry", "roofing"),
TaskPrecedence("ceiling", "painting"),
TaskPrecedence("roofing", "windows"),
TaskPrecedence("roofing", "facade"),
TaskPrecedence("plumbing", "facade"),
TaskPrecedence("roofing", "garden"),
TaskPrecedence("plumbing", "garden"),
TaskPrecedence("windows", "moving"),
TaskPrecedence("facade", "moving"),
TaskPrecedence("garden", "moving"),
TaskPrecedence("painting", "moving"),
}
```
There are three workers with varying skill levels in regard to the ten tasks. If a worker has a skill level of zero for a task, he may not be assigned to the task.
<p>
| *Task* | *Joe* | *Jack* | *Jim* |
|---|---|---|---|
|masonry |9 | 5 | 0|
|carpentry |7 | 0 | 5|
|plumbing |0 | 7 | 0|
|ceiling |5 | 8 | 0|
|roofing |6 | 7 | 0|
|painting |0 | 9 | 6|
|windows |8 | 0 | 5|
|façade |5 | 5 | 0|
|garden |5 | 5 | 9|
|moving |6 | 0 | 8|
##### Workers Names
```
WORKERS = {"Joe", "Jack", "Jim"}
```
##### Workers Name and level for each of there skill
```
Skill = (namedtuple("Skill", ["worker", "task", "level"]))
SKILLS = {Skill("Joe", "masonry", 9),
Skill("Joe", "carpentry", 7),
Skill("Joe", "ceiling", 5),
Skill("Joe", "roofing", 6),
Skill("Joe", "windows", 8),
Skill("Joe", "facade", 5),
Skill("Joe", "garden", 5),
Skill("Joe", "moving", 6),
Skill("Jack", "masonry", 5),
Skill("Jack", "plumbing", 7),
Skill("Jack", "ceiling", 8),
Skill("Jack", "roofing", 7),
Skill("Jack", "painting", 9),
Skill("Jack", "facade", 5),
Skill("Jack", "garden", 5),
Skill("Jim", "carpentry", 5),
Skill("Jim", "painting", 6),
Skill("Jim", "windows", 5),
Skill("Jim", "garden", 9),
Skill("Jim", "moving", 8)
}
```
##### Utility functions
find_tasks: returns the task it refers to in the TASKS vector
```
def find_tasks(name):
return next(t for t in TASKS if t.name == name)
```
find_skills: returns the skill it refers to in the SKILLS vector
```
def find_skills(worker, task):
return next(s for s in SKILLS if (s.worker == worker) and (s.task == task))
```
find_max_level_skill: returns the tuple "skill" where the level is themaximum for a given task
```
def find_max_level_skill(task):
st = [s for s in SKILLS if s.task == task]
return next(sk for sk in st if sk.level == max([s.level for s in st]))
```
### Step 4: Set up the prescriptive model
<h3>Create the model container</h3>
<p>
The model is represented by a Python object that is filled with the different model elements (variables, constraints, objective function, etc). The first thing to do is then to create such an object:
```
mdl = CpoModel(name="HouseBuilding")
```
#### Define the decision variables
<h5><i><font color=blue>Concept: interval variable</font></i></h5>
<p>
<ul>
<li> What for?<br>
<blockquote> Modeling an interval of time during which a particular property holds <br>
(an activity executes, a resource is idle, a tank must be non-empty, …)</blockquote>
<li> Example:<br>
<blockquote><code><font color=green>interval_var(start=(0,1000), end=(0,1000), size=(10,20))</font></code>
</blockquote>
<!-- <img src = "./house_building_utils/intervalVar.png" > -->
<img src = "https://github.com/IBMDecisionOptimization/docplex-examples/blob/master/examples/cp/jupyter/house_building_utils/intervalVar.PNG?raw=true" >
<li>Properties:
<ul>
<li>The **value** of an interval variable is an integer interval [start,end)
<li>**Domain** of possible values: [0,10), [1,11), [2,12),...[990,1000), [0,11),[1,12),...
<li>Domain of interval variables is represented **compactly** in CP Optimizer (a few bounds: smin, smax, emin, emax, szmin, szmax)
</ul>
</ul>
For each house, an interval variable is created for each task.<br>
This interval must start and end inside the period_domain and its duration is set as the value stated in TASKS definition.
```
tasks = {} # dict of interval variable for each house and task
for house in HOUSES:
for task in TASKS:
tasks[(house, task)] = mdl.interval_var(start=period_domain,
end=period_domain,
size=task.duration,
name="house {} task {}".format(house, task))
```
<h5><i><font color=blue>Concept: optional interval variable</font></i></h5>
<p>
<ul>
<li>Interval variables can be defined as being **optional** that is, it is part of the decisions of the problem to decide whether the interval will be **present** or **absent** in the solution<br><br>
<li> What for?<br>
<blockquote> Modeling optional activities, alternative execution modes for activities, and … most of the discrete decisions in a schedule</blockquote>
<li> Example:<br>
<blockquote><code><font color=green>interval_var(</font><font color=red>optional=True</font><font color=green>, start=(0,1000), end=(0,1000), size=(10,20))</font></code>
</blockquote>
<li>Properties:
<ul>
<li>An optional interval variable has an additional possible value in its domain (absence value)
<li>**Optionality** is a powerful property that you must learn to leverage in your models
</ul>
</ul>
For each house, an __optional__ interval variable is created for each skill.<br>
Skill being a tuple (worker, task, level), this means that for each house, an __optional__ interval variable is created for each couple worker-task such that the skill level of this worker for this task is > 0.<p>
The "**set_optional()**" specifier allows a choice between different variables, thus between different couples house-skill.
This means that the engine decides if the interval will be present or absent in the solution.
```
wtasks = {} # dict of interval variable for each house and skill
for house in HOUSES:
for skill in SKILLS:
iv = mdl.interval_var(name='H' + str(house) + '-' + skill.task + '(' + skill.worker + ')')
iv.set_optional()
wtasks[(house, skill)] = iv
```
#### Express the business constraints
<h5>Temporal constraints</h5>
<h5><i><font color=blue>Concept: precedence constraint</font></i></h5>
<p>
<ul>
<li> What for?<br>
<ul>
<li>Modeling temporal constraints between interval variables
<li>Modeling constant or variable minimal delays
</ul>
<li>Properties
<blockquote>Semantic of the constraints handles optionality (as for all constraints in CP Optimizer).<br>
Example of endBeforeStart:<br>
<code><font color=green>end_before_start(a,b,z)</font></code><br>
present(a) <font color=red>AND</font> present(b) ⇒ end(a)+z ⩽ start(b)
</blockquote>
<ul>
The tasks in the model have precedence constraints that are added to the model.
```
for h in HOUSES:
for p in TASK_PRECEDENCES:
mdl.add(mdl.end_before_start(tasks[(h, find_tasks(p.beforeTask))], tasks[(h, find_tasks(p.afterTask))]))
```
<h5>Alternative workers</h5>
<h5><i><font color=blue>Concept: alternative constraint</font></i></h5>
<p>
<ul>
<li> What for?<br>
<ul>
<li>Modeling alternative resource/modes/recipes
<li>In general modeling a discrete selection in the schedule
</ul>
<li> Example:<br>
<blockquote><code><font color=green>alternative(a,[b1,...,bn])</font></code>
</blockquote>
<!-- <img src = "./house_building_utils/alternative.png" > -->
<img src = "https://github.com/IBMDecisionOptimization/docplex-examples/blob/master/examples/cp/jupyter/house_building_utils/alternative.PNG?raw=true" >
<li>Remark: Master interval variable **a** can of course be optional
</ul>
To constrain the solution so that exactly one of the interval variables wtasks associated with a given task of a given house is to be present in the solution, an "**alternative**" constraint is used.
```
for h in HOUSES:
for t in TASKS:
mdl.add(mdl.alternative(tasks[(h, t)], [wtasks[(h, s)] for s in SKILLS if (s.task == t.name)], 1))
```
<h5>No overlap constraint</h5>
<h5><i><font color=blue>Concept: No-overlap constraint</font></i></h5>
<p>
<ul>
<li> Constraint noOverlap schedules a group of interval variables in such a way that they do not overlap in time.
<li> Absent interval variables are ignored.
<li>It is possible to constrain minimum delays between intervals using transition matrix.
<li>It is possible to constraint the first, last in the sequence or next or preceding interval
</ul>
<!-- <img src = "./house_building_utils/noOverlap.png" > -->
<img src = "https://github.com/IBMDecisionOptimization/docplex-examples/blob/master/examples/cp/jupyter/house_building_utils/noOverlap.PNG?raw=true" >
To add the constraints that a given worker can be assigned only one task at a given moment in time, a **noOverlap** constraint is used.
```
for w in WORKERS:
mdl.add(mdl.no_overlap([wtasks[(h, s)] for h in HOUSES for s in SKILLS if s.worker == w]))
```
#### Express the objective
The presence of an interval variable in wtasks in the solution must be accounted for in the objective. Thus for each of these possible tasks, the cost is incremented by the product of the skill level and the expression representing the presence of the interval variable in the solution.<p>
The objective of this problem is to maximize the skill level used for all the tasks.
```
obj = mdl.sum([s.level * mdl.presence_of(wtasks[(h, s)]) for s in SKILLS for h in HOUSES])
mdl.add(mdl.maximize(obj))
```
#### Solve the model
The model is now completely defined. It is time to solve it !
```
# Solve the model
print("\nSolving model....")
msol = mdl.solve(TimeLimit=10)
```
### Step 5: Investigate the solution and then run an example analysis
```
print("Solve status: " + msol.get_solve_status())
if msol.is_solution():
stdout.write("Solve time: " + str(msol.get_solve_time()) + "\n")
# Sort tasks in increasing begin order
ltasks = []
for hs in HOUSES:
for tsk in TASKS:
(beg, end, dur) = msol[tasks[(hs, tsk)]]
ltasks.append((hs, tsk, beg, end, dur))
ltasks = sorted(ltasks, key = lambda x : x[2])
# Print solution
print("\nList of tasks in increasing start order:")
for tsk in ltasks:
print("From " + str(tsk[2]) + " to " + str(tsk[3]) + ", " + tsk[1].name + " in house " + str(tsk[0]))
else:
stdout.write("No solution found\n")
```
#### Import graphical tools
*You can set __POP\_UP\_GRAPHIC=True__ if you prefer a pop up graphic window instead of an inline one.*
```
POP_UP_GRAPHIC=False
import docplex.cp.utils_visu as visu
import matplotlib.pyplot as plt
if not POP_UP_GRAPHIC:
%matplotlib inline
#Change the plot size
from pylab import rcParams
rcParams['figure.figsize'] = 15, 3
```
#### Draw solution
#### Useful functions
With the aim to facilitate the display of tasks names, we keep only the n first characters.
```
def compact_name(name,n): return name[:n]
if msol and visu.is_visu_enabled():
workers_colors = {}
workers_colors["Joe"] = 'lightblue'
workers_colors["Jack"] = 'violet'
workers_colors["Jim"] = 'lightgreen'
visu.timeline('Solution per houses', 0, MAX_AMOUNT_OF_PERIODS)
for h in HOUSES:
visu.sequence(name="house " + str(h))
for s in SKILLS:
wt = msol.get_var_solution(wtasks[(h,s)])
if wt.is_present():
color = workers_colors[s.worker]
wtname = compact_name(s.task,2)
visu.interval(wt, color, wtname)
visu.show()
```
The purpose of this function is to compact the names of the different tasks with the aim of making the graphical display readable. </p>
For example "H3-garden" becomes "G3"
```
def compact_house_task(name):
loc, task = name[1:].split('-', 1)
return task[0].upper() + loc
```
Green-like color when task is using the most skilled worker
Red-like color when task does not use the most skilled worker
```
if msol and visu.is_visu_enabled():
visu.timeline('Solution per workers', 0, MAX_AMOUNT_OF_PERIODS)
for w in WORKERS:
visu.sequence(name=w)
for h in HOUSES:
for s in SKILLS:
if s.worker == w:
wt = msol.get_var_solution(wtasks[(h,s)])
if wt.is_present():
ml = find_max_level_skill(s.task).level
if s.level == ml:
color = 'lightgreen'
else:
color = 'salmon'
wtname = compact_house_task(wt.get_name())
visu.interval(wt, color, wtname)
visu.show()
```
<h4>Going further with Constraint Programming</h4>
The last available installable package is available on Pypi here: https://pypi.python.org/pypi/docplex
A complete set of modeling examples can be downloaded here: https://github.com/IBMDecisionOptimization/docplex-examples
## Summary
You learned how to set up and use the IBM Decision Optimization CPLEX Modeling for Python to build and solve a Constraint Programming model.
#### References
* [CPLEX Modeling for Python documentation](https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html)
* [IBM Decision Optimization](https://www.ibm.com/analytics/decision-optimization)
* Need help with DOcplex or to report a bug? Please go [here](https://stackoverflow.com/questions/tagged/docplex)
* Contact us at dofeedback@wwpdl.vnet.ibm.com
Copyright © 2017, 2021 IBM. IPLA licensed Sample Materials.
| github_jupyter |
# Aplicando Python para análisis de precios: simulación de escenarios futuros de precios
<img style="float: right; margin: 0px 0px 15px 15px;" src="https://upload.wikimedia.org/wikipedia/commons/d/d7/Philippine-stock-market-board.jpg" width="400px" height="125px" />
> En la clase anterior vimos como importar datos de activos de la base de datos de Yahoo Finance usando el paquete pandas-datareader. En esta clase, veremos como pronosticar escenarios de evolución de precios, suponiendo que los rendimientos diarios se distribuyen normalmente. Como esta evolución de precios es aleatoria, utilizaremos la simulación montecarlo (hacer muchas simulaciones de escenarios de evolución de precios) para obtener probabilidades de que los precios de cierre estén encima de un valor umbral y tomar decisiones con base en estas probabilidades.
**Referencias:**
- http://pandas.pydata.org/
- http://www.learndatasci.com/python-finance-part-yahoo-finance-api-pandas-matplotlib/
## 1. Recordemos como descargar datos...
Antes que nada, para poder hacer simular escenarios de predicción de precios, vamos a recordar lo que hicimos en la clase pasada de descargar los datos de Yahoo Finance, utilizando el paquete `data` de la librería `pandas_datareader`.
Esta vez, utilizaremos los datos de precios de cierre ajustados de activos de la compañía Apple en el año 2016 para nuestra aplicación.
```
# Importamos librerías
import pandas as pd
pd.core.common.is_list_like = pd.api.types.is_list_like
import pandas_datareader.data as web
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
# Función para descargar precios de cierre ajustados de varios activos a la vez:
def get_closes(tickers, start_date=None, end_date=None, freq='d'):
# Fecha inicio por defecto (start_date='2010-01-01') y fecha fin por defecto (end_date=today)
# Frecuencia de muestreo por defecto (freq='d')
# Importamos paquetes necesarios
import pandas as pd
pd.core.common.is_list_like = pd.api.types.is_list_like
import pandas_datareader.data as web
# Creamos DataFrame vacío de precios, con el índice de las fechas
closes = pd.DataFrame(columns = tickers, index=web.YahooDailyReader(symbols=tickers[0], start=start_date, end=end_date, interval=freq).read().index)
# Agregamos cada uno de los precios con YahooDailyReader
for ticker in tickers:
df = web.YahooDailyReader(symbols=ticker, start=start_date, end=end_date, interval=freq).read()
closes[ticker]=df['Adj Close']
closes.index_name = 'Date'
closes = closes.sort_index()
return closes
# Descargamos datos...
# Instrumento: Apple
name = ['AAPL']
# Fechas de interés (inicio y fin): 2010-2016
start, end = '2010-01-01', '2016-12-31'
# Función DataReader
closes = get_closes(tickers=name, start_date=start, end_date=end, freq='d')
closes
# Graficamos
closes.plot(figsize=(10,6));
```
## 2. Simulación de rendimientos diarios
Recordemos que los precios diarios de cierre ajustados no son un proceso estocástico estacionario, pero los rendimientos diarios si lo son. Por tanto calculamos los rendimientos a partir de los precios de cierre, obtenemos sus propiedades estadísticas muestrales y proyectamos los rendimientos. Luego, obtenemos la proyección de los precios.
Para una sucesión de precios $\{S_t\}_{t=0}^{n}$, el rendimiento simple $R_t$ se define como el el cambio porcentual
$$
R_t=\frac{S_t-S_{t-1}}{S_{t-1}}\approx \ln\left(\frac{S_t}{S_{t-1}}\right)=r_t.
$$
para $t=1,\ldots,n$.
Para el ejemplo en curso, ¿cómo calcular esto?
Además, supusimos que los rendimientos diarios eran una variable aleatoria con distribución normal (que se caracteriza con su media y varianza). Por tanto obtenemos la media y desviación estandar muestrales. Hagamos una función que retorne lo anterior.
```
# Calcular rendimientos diarios y graficarlos
ret = closes.pct_change().dropna()
ret.plot(figsize=(10,6));
```
Entonces, suponemos que la diferencia logaritmica de los precios (rendimientos diarios) tiene una distribución normal.
¿Cómo se caracteriza una [distribución normal](https://es.wikipedia.org/wiki/Distribuci%C3%B3n_normal)?
```
# Calculamos media y desviación estándar
mu, std = ret.mean().AAPL, ret.std().AAPL
mu, std
```
Habiendo caracterizado los rendimientos diarios como una variable aleatoria normal con la media y la varianza muestral obtenida de los datos del 2016, podemos generar números aleatorios con estas características para simular el comportamiento de los precios de cierre de las acciones en el 2017 (hay un supuesto de que las cosas no cambiarán fundamentalmente).
Sin embargo, cada simulación que hagamos nos conducirá a distintos resultados (los precios siguen evolucionando aleatoriamente). Entonces, lo que haremos es simular varios escenarios para así ver alguna tendencia y tomar decisiones.
Hagamos una una función que simule varios escenarios de rendimientos diarios rendimientos diarios y que devuelva un dataframe con esta simulación.
```
# Ayuda en la función np.random.randn
help(np.random.randn)
# Función que simula varios escenarios de rendimientos diarios
def ret_sim(mu, sigma, ndays, nscen, start_date):
dates = pd.date_range(start=start_date,periods=ndays)
return pd.DataFrame(data = sigma*np.random.randn(ndays, nscen)+mu, index = dates)
# Simulamos 100 escenarios para todo el 2017
simret = ret_sim(mu, std, 252, 10000, '2017-01-01')
# Mostrar
#simret
```
## 3. Proyección de precios de cierre
Por tanto, para calcular los precios, tenemos:
$$\begin{align}
p_i&=p_{i-1}(R_i+1)\\
p_{i+1}&=p_i(R_{i+1}+1)=p_{i-1}(R_i+1)(R_{i+1}+1)\\
&\vdots\\
p_{i+k}&=p_{i-1}(R_i+1)\cdots(R_{i+k}+1).
\end{align}$$
Si hacemos $i=0$ en la última ecuación, tenemos que $p_{k}=p_{-1}\exp(r_0+\cdots+r_{k})$, donde $p_{-1}$ es el último precio reportado en el 2016.
Con los rendimientos, calculamos los precios de cierre...
```
# Obtenemos los precios
simcloses = closes.iloc[-1].AAPL*((1+simret).cumprod())
#simcloses
```
Concatenamos y graficamos...
```
# Concatenar y graficar
#simcloses_ex = pd.concat([closes.iloc[-200:], simcloses])
#simcloses_ex.plot(figsize=(10,6), legend=False);
closes.iloc[-1].AAPL*1.1
```
## 4. Probabilidad Precio-Umbral
Ya que tenemos muchos escenarios de precios proyectados, podemos ver varias cosas. Por ejemplo, ¿cuál es la probabilidad de que el precio de cierre sobrepase algún valor umbral en algún momento?
```
# Umbral de 110% del ultimo precio
K = closes.iloc[-1].AAPL*1.1
# Fechas
dates = simcloses.index
# DataFrame de Strikes
Strike = pd.DataFrame(index = dates, columns=['K'], data = K*np.ones(len(dates)))
# Concatenar y graficar
#simcloses_ex_K = pd.concat([simcloses_ex.T, Strike.T]).T
#simcloses_ex_K.plot(figsize=(10,6), legend=False);
# Comparar cada escenario en cada fecha
TF = simcloses>K
# Sumamos para cada fecha y dividimos entre el número de escenarios
prob = pd.DataFrame(TF.sum(axis=1)/100)
# Gráfico de probabilidad
prob.plot(figsize=(10,6), legend=False);
```
___
Entonces, ya aprendimos a bajar datos con pandas-datareader. En específico, a partir de los precios de cierre ajustados obtuvimos los rendimientos diarios.
Suponiendo que los rendimientos diarios son un proceso estocástico estacionario de distribución normal, pudimos caracaterizarlo y proyectar varios escenarios de evolución de los precios (montecarlo).
Con estas proyecciones pudimos calcular probabilidades de sobrepasar cierto precio umbral: toma de decisiones.
<script>
$(document).ready(function(){
$('div.prompt').hide();
$('div.back-to-top').hide();
$('nav#menubar').hide();
$('.breadcrumb').hide();
$('.hidden-print').hide();
});
</script>
<footer id="attribution" style="float:right; color:#808080; background:#fff;">
Created with Jupyter by Esteban Jiménez Rodríguez.
</footer>
| github_jupyter |
# freud.density.LocalDensity
The `freud.density` module is intended to compute a variety of quantities that relate spatial distributions of particles with other particles.
In this notebook, we demonstrate `freud`'s local density calculation, which can be used to characterize the particle distributions in some systems.
In this example, we consider a toy example of calculating the particle density in the vicinity of a set of other points.
This can be visualized as, for example, billiard balls on a table with certain regions of the table being stickier than others.
In practice, this method could be used for analyzing, *e.g*, binary systems to determine how densely one species packs close to the surface of the other.
```
import freud
import matplotlib.pyplot as plt
import numpy as np
from matplotlib import patches
# Define some helper plotting functions.
def add_patches(ax, points, radius=1, fill=False, color="#1f77b4", ls="solid", lw=None):
"""Add set of points as patches with radius to the provided axis"""
for pt in points:
p = patches.Circle(
pt,
fill=fill,
linestyle=ls,
radius=radius,
facecolor=color,
edgecolor=color,
lw=lw,
)
ax.add_patch(p)
def plot_lattice(box, points, radius=1, ls="solid", lw=None):
"""Helper function for plotting points on a lattice."""
fig, ax = plt.subplots(1, 1, figsize=(9, 9))
box.plot(ax=ax)
add_patches(ax, points, radius, ls=ls, lw=lw)
return fig, ax
```
Let us consider a set of regions on a square lattice.
```
area = 2
radius = np.sqrt(area / np.pi)
spot_area = area * 100
spot_radius = np.sqrt(spot_area / np.pi)
num = 6
scale = num * 4
uc = freud.data.UnitCell(freud.Box.square(1), [[0.5, 0.5, 0]])
box, spot_centers = uc.generate_system(num, scale=scale)
fig, ax = plot_lattice(box, spot_centers, spot_radius, ls="dashed", lw=2.5)
plt.tick_params(axis="both", which="both", labelsize=14)
plt.show()
```
Now let's add a set of points to this box.
Points are added by drawing from a normal distribution centered at each of the regions above.
For demonstration, we will assume that each region has some relative "attractiveness," which is represented by the covariance in the normal distributions used to draw points.
Specifically, as we go up and to the right, the covariance increases proportional to the distance from the lower right corner of the box.
```
points = []
fractional_distances_to_corner = np.linalg.norm(
box.make_fractional(spot_centers), axis=-1
)
cov_basis = 20 * fractional_distances_to_corner
for i, p in enumerate(spot_centers):
np.random.seed(i)
cov = cov_basis[i] * np.diag([1, 1, 0])
points.append(np.random.multivariate_normal(p, cov, size=(50,)))
points = box.wrap(np.concatenate(points))
fig, ax = plot_lattice(box, spot_centers, spot_radius, ls="dashed", lw=2.5)
plt.tick_params(axis="both", which="both", labelsize=14)
add_patches(ax, points, radius, True, "k", lw=None)
plt.show()
```
We see that the density decreases as we move up and to the right.
In order to compute the actual densities, we can leverage the `LocalDensity` class.
The class allows you to specify a set of query points around which the number of other points is computed.
These other points can, but need not be, distinct from the query points.
In our case, we want to use the blue regions as our query points with the small black dots as our data points.
When we construct the `LocalDensity` class, there are two arguments.
The first is the radius from the query points within which particles should be included in the query point's counter.
The second is the circumsphere diameter of the **data points**, not the query points.
This distinction is critical for getting appropriate density values, since these values are used to actually check cutoffs and calculate the density.
```
density = freud.density.LocalDensity(spot_radius, radius)
density.compute(system=(box, points), query_points=spot_centers);
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
for i, data in enumerate([density.num_neighbors, density.density]):
poly = np.poly1d(np.polyfit(cov_basis, data, 1))
axes[i].tick_params(axis="both", which="both", labelsize=14)
axes[i].scatter(cov_basis, data)
x = np.linspace(*axes[i].get_xlim(), 30)
axes[i].plot(x, poly(x), label="Best fit")
axes[i].set_xlabel("Covariance", fontsize=16)
axes[0].set_ylabel("Number of neighbors", fontsize=16)
axes[1].set_ylabel("Density", fontsize=16)
plt.show()
```
As expected, we see that increasing the variance in the number of points centered at a particular query point decreases the total density at that point.
The trend is noisy since we are randomly sampling possible positions, but the general behavior is clear.
| github_jupyter |
```
%reload_ext autoreload
%autoreload 2
%matplotlib inline
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID";
os.environ["CUDA_VISIBLE_DEVICES"]="0"
import numpy as np
import tensorflow as tf
print("Tensorflow version " + tf.__version__)
```
# Using *ktrain* to Facilitate a Normal TensorFlow Workflow
This example notebook simply illustrates how *ktrain* can be used in a **minimally-invasive** way within
a normal TensorFlow workflow. In this notebook, we will store our datasets in the form of `tf.Datasets` and build our own `tf.Keras` model following the example of TensorFlow's [Keras MNIST TPU.ipynb](https://colab.research.google.com/github/tensorflow/tpu/blob/master/tools/colab/keras_mnist_tpu.ipynb#scrollTo=cCpkS9C_H7Tl). We will then simply use **ktrain** as a lightweight wrapper for our model and data to estimate a learning rate, train the model, inspect the model, and make predictions.
## Detect Hardware: CPU vs. GPU vs. TPU
```
# Detect hardware
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection
except ValueError:
tpu = None
gpus = tf.config.experimental.list_logical_devices("GPU")
# Select appropriate distribution strategy
if tpu:
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu, steps_per_run=128) # Going back and forth between TPU and host is expensive. Better to run 128 batches on the TPU before reporting back.
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
elif len(gpus) > 1:
strategy = tf.distribute.MirroredStrategy([gpu.name for gpu in gpus])
print('Running on multiple GPUs ', [gpu.name for gpu in gpus])
elif len(gpus) == 1:
strategy = tf.distribute.get_strategy() # default strategy that works on CPU and single GPU
print('Running on single GPU ', gpus[0].name)
else:
strategy = tf.distribute.get_strategy() # default strategy that works on CPU and single GPU
print('Running on CPU')
print("Number of accelerators: ", strategy.num_replicas_in_sync)
```
## Prepare Training and Validation Data as `tf.Datasets`
Download the dataset files from [LeCun's website](http://yann.lecun.com/exdb/mnist/).
```
BATCH_SIZE = 64 * strategy.num_replicas_in_sync # Gobal batch size.
training_images_file = 'data/mnist_lecun/train-images-idx3-ubyte'
training_labels_file = 'data/mnist_lecun/train-labels-idx1-ubyte'
validation_images_file = 'data/mnist_lecun/t10k-images-idx3-ubyte'
validation_labels_file = 'data/mnist_lecun/t10k-labels-idx1-ubyte'
```
Note that, if training using a TPU, these should be set as follows:
```python
training_images_file = 'gs://mnist-public/train-images-idx3-ubyte'
training_labels_file = 'gs://mnist-public/train-labels-idx1-ubyte'
validation_images_file = 'gs://mnist-public/t10k-images-idx3-ubyte'
validation_labels_file = 'gs://mnist-public/t10k-labels-idx1-ubyte'
```
You may need to authenticate:
```python
IS_COLAB_BACKEND = 'COLAB_GPU' in os.environ # this is always set on Colab, the value is 0 or 1 depending on GPU presence
if IS_COLAB_BACKEND:
from google.colab import auth
# Authenticates the Colab machine and also the TPU using your
# credentials so that they can access your private GCS buckets.
auth.authenticate_user()
```
```
def read_label(tf_bytestring):
label = tf.io.decode_raw(tf_bytestring, tf.uint8)
label = tf.reshape(label, [])
label = tf.one_hot(label, 10)
return label
def read_image(tf_bytestring):
image = tf.io.decode_raw(tf_bytestring, tf.uint8)
image = tf.cast(image, tf.float32)/255.0
image = tf.reshape(image, [28*28])
return image
def load_dataset(image_file, label_file):
imagedataset = tf.data.FixedLengthRecordDataset(image_file, 28*28, header_bytes=16)
imagedataset = imagedataset.map(read_image, num_parallel_calls=16)
labelsdataset = tf.data.FixedLengthRecordDataset(label_file, 1, header_bytes=8)
labelsdataset = labelsdataset.map(read_label, num_parallel_calls=16)
dataset = tf.data.Dataset.zip((imagedataset, labelsdataset))
return dataset
def get_training_dataset(image_file, label_file, batch_size):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM
dataset = dataset.shuffle(5000, reshuffle_each_iteration=True)
dataset = dataset.repeat() # Mandatory for Keras for now
dataset = dataset.batch(batch_size, drop_remainder=True) # drop_remainder is important on TPU, batch size must be fixed
dataset = dataset.prefetch(-1) # fetch next batches while training on the current one (-1: autotune prefetch buffer size)
return dataset
def get_validation_dataset(image_file, label_file):
dataset = load_dataset(image_file, label_file)
dataset = dataset.cache() # this small dataset can be entirely cached in RAM
dataset = dataset.batch(10000, drop_remainder=True) # 10000 items in eval dataset, all in one batch
dataset = dataset.repeat() # Mandatory for Keras for now
return dataset
def load_label_dataset(label_file):
labelsdataset = tf.data.FixedLengthRecordDataset(label_file, 1, header_bytes=8)
labelsdataset = labelsdataset.map(read_label, num_parallel_calls=16)
return labelsdataset
# instantiate the datasets
training_dataset = get_training_dataset(training_images_file, training_labels_file, BATCH_SIZE)
validation_dataset = get_validation_dataset(validation_images_file, validation_labels_file)
# exract ground truth labels
training_labels = np.vstack(list(load_label_dataset(training_labels_file).as_numpy_iterator()))
validation_labels = np.vstack(list(load_label_dataset(validation_labels_file).as_numpy_iterator()))
```
## Build a Model
```
# This model trains to 99.4% accuracy in 10 epochs (with a batch size of 64)
def make_model():
model = tf.keras.Sequential(
[
tf.keras.layers.Reshape(input_shape=(28*28,), target_shape=(28, 28, 1), name="image"),
tf.keras.layers.Conv2D(filters=12, kernel_size=3, padding='same', use_bias=False), # no bias necessary before batch norm
tf.keras.layers.BatchNormalization(scale=False, center=True), # no batch norm scaling necessary before "relu"
tf.keras.layers.Activation('relu'), # activation after batch norm
tf.keras.layers.Conv2D(filters=24, kernel_size=6, padding='same', use_bias=False, strides=2),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Conv2D(filters=32, kernel_size=6, padding='same', use_bias=False, strides=2),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(200, use_bias=False),
tf.keras.layers.BatchNormalization(scale=False, center=True),
tf.keras.layers.Activation('relu'),
tf.keras.layers.Dropout(0.4), # Dropout on dense layer only
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam', # learning rate will be set by LearningRateScheduler
loss='categorical_crossentropy',
metrics=['accuracy'])
return model
with strategy.scope():
model = make_model()
# set up learning rate decay [FROM ORIGINAL EXAMPLE BUT NOT USED]
# NOT NEEDED: we will use ktrain to find LR and decay learning rate during training
LEARNING_RATE = 0.01
LEARNING_RATE_EXP_DECAY = 0.6 if strategy.num_replicas_in_sync == 1 else 0.7
lr_decay = tf.keras.callbacks.LearningRateScheduler(
lambda epoch: LEARNING_RATE * LEARNING_RATE_EXP_DECAY**epoch,
verbose=True)
```
## Use *ktrain* With Our Model and Data
### Wrap tf.Datasets in a `ktrain.TFDataset` wrapper and create `Learner`
```
import ktrain
trn = ktrain.TFDataset(training_dataset, n=training_labels.shape[0], y=training_labels)
val = ktrain.TFDataset(validation_dataset, n=validation_labels.shape[0], y=validation_labels)
learner = ktrain.get_learner(model, train_data=trn, val_data=val)
```
### Find Learning Rate
```
learner.lr_find(show_plot=True)
```
## Train the Model Using a Cosine Annealing LR Schedule
```
learner.fit(5e-3, 1, cycle_len=10, checkpoint_folder='/tmp/mymodel')
# cosine annealed LR schedule
learner.plot('lr')
# training vs. validation loss
learner.plot('loss')
```
### Inspect Model
#### Evaluate as Normal
```
learner.model.evaluate(validation_dataset, steps=1)
```
#### Validation Metrics
```
learner.validate(class_names=list(map(str, range(10))))
```
#### View Top Losses
```
learner.view_top_losses(n=1)
```
### Making Predictions
```
preds = learner.predict(val)
preds = np.argmax(preds, axis=1)
actual = learner.ground_truth(val)
actual = np.argmax(actual, axis=1)
import pandas as pd
df = pd.DataFrame(zip(preds, actual), columns=['Predicted', 'Actual'])
df.head()
```
## Save Model and Reload Model
```
learner.save_model('/tmp/my_tf_model')
learner.load_model('/tmp/my_tf_model')
learner.model.evaluate(validation_dataset, steps=1)
```
| github_jupyter |
# Design of experiments
This notebook is about the statistical considerations related to fMRI experimental design. Make sure you do the other lab (`glm_part2_inference.ipynb`) first!
Experimental designs for fMRI studies come in different flavors, depending on what hypotheses you have about the phenomenon of interest and how you manipulate this. Apart from the different types of experimental designs (e.g., subtractive, factorial, parametric), there are a couple of general recommendations w.r.t. experimental design that can optimize the chance of finding positive results, which will be discussed in this notebook as well. These recommendations have to do with the specific ordering and timing of the events in your experiment. For example, suppose you show images of cats (condition: "C") and dogs (condition: "D") to subjects in the scanner, and you're interested if the brain responds differently to images of dogs compared to images of cats. What ordering ("CCCCDDDD" or "CDCDCDCD" or "CCDCDDCD"?) and timing (how long should I wait to present another stimulus?) of the stimuli will yield the best (here: highest) effect possible, and why?
**What you'll learn**: after this lab, you'll ...
- understand what 'design variance' is and how it relates to 'efficiency'
- understand the effect of design variance *t*-values
- know how to calculate design variance in Python
**Estimated time needed to complete**: 4-6 hours<br>
```
# First some imports
import numpy as np
import matplotlib.pyplot as plt
from nilearn.glm.first_level.hemodynamic_models import glover_hrf
from scipy.stats import pearsonr
```
## Types of designs: factorial and parametric designs
Last week, we discussed contrasts at length and how to use contrast-vectors to specify simple hypotheses (e.g., happy faces > sad faces). The contrast-vectors from last week's lab were examples of either simple contrasts-against-baseline ($H_{0}: \beta = 0$) or examples of *subtractive designs* (also called categorical designs; e.g., $H_{0}: \beta_{1} - \beta_{2} = 0$). There are, however, more types of designs possible, like *factorial* and *parametric* designs. In this first section, we'll discuss these two designs shortly and have you implement GLMs to test hypotheses forwarded by these designs.
### Factorial designs
Factorial designs are designs in which each event (e.g., stimulus) may be represented by a combination of different conditions. For example, you could show images of squares and circles (condition 1: shape) which may be either green or red (condition 2: color). See the image below for a visualization of these conditions and the associated contrasts.

Before we'll explain this figure in more detail, let's generate some data. We'll assume that the TR is 1 second and that all onsets are always syncronized with the TR (so there won't be onsets at, e.g., 10.295 seconds). This way, we can ignore the downsampling issue. We'll convolve the data with an HRF already and plot the design matrix below:
```
from niedu.utils.nii import simulate_signal
exp_length = 160
TR = 1
xmax = exp_length // TR
y, X = simulate_signal(
onsets=np.array(
[0, 40, 80, 120, # red squares
10, 50, 90, 130, # red circles
20, 60, 100, 140, # green squares
30, 70, 110, 150] # green circles
),
conditions=['rq'] * 4 + ['rc'] * 4 + ['gs'] * 4 + ['gc'] * 4,
TR=TR,
duration=exp_length,
icept=0,
params_canon=[1, 2, 3, 4],
rnd_seed=42,
plot=False
)
X = X[:, :5]
plt.figure(figsize=(15, 5))
plt.plot(X[:, 1], c='r', ls='--')
plt.plot(X[:, 2], c='r')
plt.plot(X[:, 3], c='g', ls='--')
plt.plot(X[:, 4], c='g')
plt.xlim(0, xmax)
plt.ylim(-0.3, 1.2)
plt.xlabel('Time (seconds/TRs)', fontsize=20)
plt.ylabel('Activity (A.U.)', fontsize=20)
plt.legend(['red squares', 'red circles', 'green squares', 'green circles'], frameon=False)
plt.grid()
plt.show()
```
We have also the time series of a (hypothetical) fMRI voxel, loaded and plotted below:
```
plt.figure(figsize=(15, 5))
plt.plot(y)
plt.xlim(0, xmax)
plt.axhline(0, ls='--', c='k', lw=0.75)
plt.xlabel('Time (seconds/TRs)', fontsize=20)
plt.ylabel('Activity (A.U.)', fontsize=20)
plt.grid()
plt.show()
```
<div class='alert alert-warning'>
<b>ToDo</b> (2 points): Time to refresh your memory on how to implement the GLM! Run linear regression with the design specified above (i.e., the <tt>X</tt> variable). Store the resulting parameters (i.e., the "betas") in a new variable named <tt>betas_todo</tt>. Check whether the design already includes an intercept!
</div>
```
from numpy.linalg import inv
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo'''
from niedu.tests.nii.week_3 import test_glm_refresher
test_glm_refresher(X, y, betas_todo)
```
Alright, now, from the figure above, you can see there are many different contrast possible! First of all, we can test for *main effects*: these are effects of a single condition, collapsing over the other(s). For example, testing whether red stimuli lead to different activity levels than green stimuli (regardless of shape) would be a test of a main effect. Technically, main effects within factorial designs are tested with F-tests, which are undirectional tests, which mean that they test for *any* difference between conditions (e.g., *either* that red > green *or* green > red). However, this rarely happens in cognitive neuroscience, as most hypotheses are directional (e.g., red > green), so we'll focus on those type of hypothese in factorial designs here.
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Define a contrast-vector below (which should be a numpy array with 5 values) with the name <tt>cvec_red_green</tt> that would test the hypothesis that red stimuli evoke more activity than green stimuli (regardless of shape).
</div>
```
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo '''
from niedu.tests.nii.week_3 import test_red_larger_than_green
test_red_larger_than_green(cvec_red_green)
```
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Another hypothesis that you could have is that circles evoke more activity than squares (regardless of color). Define a contrast-vector below (which should be a numpy array with 5 values) with the name <tt>cvec_circle_square</tt> that would test this hypothesis.
</div>
```
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo '''
from niedu.tests.nii.week_3 import test_circles_larger_than_squares
test_circles_larger_than_squares(cvec_circle_square)
```
Alright, these (directional) main effects should be familiar as they don't differ very much from those that you saw last week. However, factorial designs are unique in that they, additionally, can test for *interactions* between conditions. Again, technically, (undirectional) F-tests should be used, but again, these are rarely used in cognitive neuroscience.
So, let's define a directional interaction effect. Suppose that, for some reason, I believe that red stimuli evoke more activity than green stimuli, but more so for circles than for squares. In other words:
\begin{align}
(\hat{\beta}_{\mathrm{red, circle}} - \hat{\beta}_{\mathrm{green,circle}}) > (\hat{\beta}_{\mathrm{red, square}} - \hat{\beta}_{\mathrm{green,square}})
\end{align}
It turns out, there is a very nice trick to figure out the corresponding contrast for this interaction: you can simply (elementwise) multiply the contrast vector for "red > green" and the contrast vector for "circle > squares"!
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Define a contrast vector below, named <tt>cvec_interaction</tt>, that tests the hypothesis that red stimuli evoke more activity than green stimuli, but more so for circles than for squares (i.e., the one from the example above).
</div>
```
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo '''
from niedu.tests.nii.week_3 import test_interaction
test_interaction(cvec_interaction)
```
Let's practice working with interactions once more.
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Define a contrast vector below, named <tt>cvec_interaction2</tt>, that tests the hypothesis that squares evoke more activity than circles, but less so for green stimuli than for red stimuli.
</div>
```
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo '''
from niedu.tests.nii.week_3 import test_interaction2
test_interaction2(cvec_interaction2)
```
### Parametric designs
So far, we have discussed only designs with conditions that are categorical, such as "male vs. female faces" and "circles vs. squares". The independent variables in your experimental design, however, do not *have* to be categorical! They can be continuous or ordinal, meaning that a particular variable might have different values (or "weights") across trials. Designs involving continuously varying properties are often called *parametric designs* or *parametric modulation*.
In parametric designs, we assume that our design affects the voxel response in two ways:
1. An "unmodulated" response (a response to the stimulus/task *independent* of parametric value);
2. A parametric modulation of the response
To make this more tangible, let's consider an example. Suppose that we have fMRI data from a reward-study. On every trial in this experiment, trials would start with a word "guess" on the screen for 1 second. Then, participants had to guess a number between 1 and 10 (which they indicated using an MRI-compatible button box). Before the experiment started, participants were told that the closer they were to the "correct" number (which was predetermined by the experimenter for every trial), the larger the reward they would get: 1 euro when their guess was correct and 10 cents less for every number that they were off (e.g., when the "correct" number was 7 and they would guess 5, then they'd receive 80 eurocents). After the participant's response and a inter-stimulus interval of 4 seconds, participants would see the amount they won on the screen.

One hypothesis you might be interested in is whether there are voxels/brain regions which response is modulated by the reward magnitude (e.g., higher activity for larger rewards, or vice versa). Before we go on, let's create some (hypothetical) experimental data. Suppose that the experiment lasted 10 minutes and contained 30 trials with varying reward magnitude, and fMRI was acquired with a TR of 1 second and onsets of the reward presentations were synchronized with the TR (again, while this is not very realistic, this obviates the need for up/downsampling).
```
np.random.seed(42)
exp_time = 60 * 10 # i.e., 14 minutes in seconds
n_trials = 30
reward_onsets = np.arange(4, exp_time, exp_time / n_trials).astype(int)
print("Number of trials: %i" % reward_onsets.size)
reward_magnitudes = np.random.randint(1, 11, size=reward_onsets.size) / 10
plt.figure(figsize=(15, 5))
plt.plot(reward_magnitudes, marker='o')
plt.xlim(-1, reward_magnitudes.size)
plt.ylim(0, 1.1)
plt.xlabel('Trial number, 0-%i (NOT TIME)' % n_trials, fontsize=20)
plt.ylabel('Reward (in euro)', fontsize=20)
plt.grid()
plt.show()
```
Now, in non-parametric designs, we would create regressors with zeros everywhere and ones at the onset of stimuli (or whatever we think will impact the fMRI data). However, in parametric designs, we create two regressors for every parametric modulation: one for the unmodulated response and one for the modulated response.
Let's start with the unmodulated response. This predictor is created like we did before: convolving a stick predictor with an HRF:
```
hrf = glover_hrf(tr=1, oversampling=1)
x_unmod = np.zeros(exp_time)
x_unmod[reward_onsets] = 1
x_unmod = np.convolve(x_unmod, hrf)[:exp_time]
plt.figure(figsize=(15, 5))
plt.plot(x_unmod)
plt.xlim(0, exp_time)
plt.xlabel('Time (sec./vols)', fontsize=20)
plt.ylabel('Activation (A.U.)', fontsize=20)
plt.title('Unmodulated regressor', fontsize=25)
plt.grid()
plt.show()
```
Now, the parametrically modulated regressor is created as follows: instead of created an initial array with zeros and *ones* at indices corresponding to the reward onset, we use the (mean-subtracted) *reward magnitude*. It is important to subtract the mean from the parametric modulation values, because this will "decorrelate" the modulated regressor from the unmodulated regressor (such that the modulated regressor explains only variance that is due to modulation of the response, not the common response towards the stimulus/task). In other words, subtracting the mean from the parametric regressor *orthogonalises* the parametric regressor with respect to the unmodulated regressor.
Then, the predictor is again convolved with the HRF to create the final modulated predictor.
<div class='alert alert-success'>
<b>Tip</b>: Check out <a href="https://journals.plos.org/plosone/article?id=10.1371/journal.pone.0126255">this excellent paper</a> by Mumford and colleagues (2015), which discusses orthogonalization in fMRI designs and when it is (in)appropriate. <a href="https://www.youtube.com/watch?v=2W7Rso-4Hqg">This video</a> also nicely explains orthogonalization in the context of parametric modulation analyses.
</div>
Let's try to create a modulated regression in a ToDo!
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Subtract the mean from the parametric modulation values (<tt>reward_magnitudes</tt>) and save this in a new variable named <tt>reward_magnitudes_ms</tt> (note: no for-loop necessary!). Now, create a new zeros-filled predictor, and set the values corresponding to the reward onsets to the mean-subtract reward magnitudes. Then, convolve the predictor with the HRF (use the variable <tt>hrf</tt> defined earlier). Make sure to trim off the excess values. Store the result in a variable named <tt>x_mod</tt>. Also plot the modulated regressor.
</div>
```
''' Implement your ToDo here. '''
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo. '''
from niedu.tests.nii.week_3 import test_parametric_modulation
test_parametric_modulation(reward_onsets, reward_magnitudes, exp_time, hrf, x_mod)
```
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Now, stack an intercept, the unmodulated regressor, and the modulated regressor in a single design matrix (with three columns; you might have to create a singleton axis with <tt>np.newaxis</tt>!). Make sure the order of the columns is as follows: intercept, unmodulated regressor, modulated regressor. Then, run linear regression with this design matrix on the variable <tt>y_reward_signal</tt> below. Save the parameters in a variable named <tt>betas_reward</tt>.
</div>
```
# Implement your ToDo here
y_reward_signal = np.load('y_reward_signal.npy')
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the ToDo above. '''
np.testing.assert_array_almost_equal(betas_reward, np.array([0, 1.05, 2.19]), decimal=2)
print("Well done!")
```
<div class='alert alert-info'>
<b>ToThink</b> (1 point): Interpret the direction of the effect of the unmodulated and modulated predictors. How does this voxel respond to the reward events?
</div>
YOUR ANSWER HERE
## Design variance/efficiency
Alright, hopefully you now know how to design contrasts for factorial and parameteric designs! This section has a slightly different focus, namely the mathematics behind design variance and efficiency! Remember that we wouldn't tell you what "design variance" was last week? Well, this week we're going to discuss and explain it *extensively*! Before we delve into this topic, let's first recap the (conceptual) formula for the *t*-value from last week (the relevance of this will become clear shortly).
Last week, you learned about the GLM and how to apply it to fMRI data to find out how much influence each predictor in your design has on the signal of a voxel. Crucially, you learned that you shouldn't look at raw beta-parameters to infer the effect of predictors, but that you should look at *normalized beta-parameters* — the **_t_-value**. Remember the formula for the *t*-value for a given contrast ($c$)?
\begin{align}
t_{\mathbf{c}\hat{\beta}} = \frac{\mathrm{effect}}{\sqrt{\mathrm{noise \cdot design\ variance}}} = \frac{\mathbf{c}\hat{\beta}}{\sqrt{\frac{SSE}{\mathrm{DF}} \cdot \mathrm{design\ variance}}}
\end{align}
The formula for the *t*-value embodies the concept that the statistics you (should) care about, *t*-values, depend both on the **effect** (sometimes confusingly called the "signal"; $\hat{\beta}$), the **noise** ($\hat{\sigma}^{2} = \frac{SSE}{\mathrm{DF}}$), and the **"design variance"**.
So, to find optimal (i.e. largest) *t*-values, we should try to optimize both the effect of our predictors (i.e. the betas), try to minimize the errors ($\hat{\sigma}^2$), and try to minimize the design variance of our model. In this lab, we'll shortly discuss the "effect" component ($\hat{\beta}$) and thereafter we'll discuss in detail the "design variance" part. We won't discuss the "noise" part, as this will be the topic of next week (preprocessing).
### Optimizing "effects"
#### Psychological factors
As discussed above, the "effect" part of the conceptual formula for the t-statistic refers to the $\beta$-parameter in the statistical formula. It may sound weird to try to "optimize" your effect, because there is no way to magically acquire a better/stronger effect from your data, right? (Well, apart from using a better/stronger MRI-scanner.) Actually, don't forget that the effect you're measuring is coming from the brain of a *human* beings (your subjects)! There are real and important psychological influences that affect the strength of your signal, and thus influence eventually the size of your $\beta$-parameter.
So, what are these psychological influences? Well, think about inattention/boredom, anxiety, sleepiness (don't underestimate how many subjects fall asleep in the scanner!), and subjects not understanding your task. As an extreme example: suppose you're showing your subject some visual stimuli in order to measure the effect of some visual property (e.g., object color) in the visual cortex. Imagine that your subject finds the task so boring that he/she falls asleep; the $\beta$-parameters in this scenario are going to be *much* lower than when the subject wouldn't have fallen asleep, of course! Sure, this is an extreme (but not uncommon!) example, but it shows the potential influence of psychological factors on the "effect" you're measuring in your data!
In short, when designing an experiment, you want to continually ask yourself: "Are subjects really doing/thinking the way I want them to?", and consequently: "Am I really measuring what I think I'm measuring?"
(The effect of psychological aspects on the measured effect is thoroughly explained in the video [Psychological principles in experimental design](https://www.youtube.com/watch?v=lwy2k8YQ-cM) from Tor Wager, which you can also find on Canvas.)
#### Design factors
Apart from taking psychological factors into account when designing your experiment, there are also design-technical factors that influence the (potential) strength of your signal: using blocked designs. We will, however, discuss this topic in a later section, because you need a better understanding of another part of the conceptual *t*-value formula first: design variance.
### Optimizing design variance
So, last week we talked quite a bit about this mysterious term "design variance" and we promised to discuss it the next week. That's exactly what we're going to do now. As we shortly explained last week, *design variance is the part of the standard error caused by the design-matrix ($X$)*. Importantly, design variance is closely related to the *efficiency* of the design matrix ($X$), i.e., efficiency is the inverse of design variance:
\begin{align}
\mathrm{efficiency} = \frac{1}{\mathrm{design\ variance}}
\end{align}
This term, efficiency, will be important in the rest of this notebook.
As these terms are inversely related, high design variance means low efficiency (which we don't want) and low design variance means high efficiency (which we want). Phrased differently, high design variance means that your design-matrix is (relatively) *inefficient* for our goal to measure significant effects (i.e., high *t*-values). But, as you might have noticed, this definition is kind of circular. What causes low design variance (high efficiency), or: what constitutes an efficient design?
Basicially, two factors contribute to an efficient design:
1. The predictors in your design should have **high variance** (i.e. they should vary a lot relative to their mean)
2. The predictors should **not** have **high covariance** (i.e. they should not correlate between each other a lot)
In general, for any **contrast between two $\beta$-parameters corresponding to predictor $j$ and $k$**, we can define their design variance as follows\*:
\begin{align}
\mathrm{design\ variance}_{j,k} = \frac{1}{\mathrm{var}[X_{j}] + \mathrm{var}[X_{k}] - 2\cdot \mathrm{cov}[X_{j}, X_{k}]}
\end{align}
As such, efficiency for this contrast would the inverse:
\begin{align}
\mathrm{efficiency}_{j,k} = \mathrm{var}[X_{j}] + \mathrm{var}[X_{k}] - 2\cdot \mathrm{cov}[X_{j}, X_{k}]
\end{align}
As you can see, design variance thus depends on the variance of the predictors *and* the covariance between predictors. Note that this formulation only applies to contrasts involving more than one parameter. For **contrasts against baseline, in which only one parameter is tested (e.g. predictor $j$)**, there is only one variance term (the other variance term and the covariance term are dropped out):
\begin{align}
\mathrm{design\ variance}_{j} = \frac{1}{\mathrm{var}[X_{j}]}
\end{align}
It is of course kind of annoying to have two different definitions (and computations) of design variance, which depend on whether you want to test a parameter against baseline or against another parameter. Therefore, people usually use the vectorized computation (i.e. using matrix multiplication), which allows you to define the formula for design variance *for any contrast-vector $\mathbf{c}$*:
\begin{align}
\mathrm{design\ variance} = \frac{1}{\mathrm{var}[X_{j}] + \mathrm{var}[X_{k}] - 2\cdot \mathrm{cov}[X_{j}, X_{k}]} = \mathbf{c}(\mathbf{X}^T\mathbf{X})^{-1}\mathbf{c}^{T}
\end{align}
While this notation, $\mathbf{c}(\mathbf{X}^{T}\mathbf{X})^{-1}\mathbf{c}$, may seem quite different than the above definitions using the $\mathrm{var}$ and $\mathrm{cov}$ terms, it is mathematically doing the same thing. The term $(X^{T}X)^{-1}$ represents (the inverse of) the variance-covariance matrix of the design ($X$) and **c** (contrast vector) is used only to "extract" the relevant variances and covariance for the particular contrast out of the entire covariance matrix of $X$.
While appearing more complex, the advantage of the vectorized definition, however, is that it works for both contrasts against baseline (e.g. `[0, 0, 1]`) and contrasts between parameters (e.g. `[0, 1, -1]`). Now, if we plug in this mathematical definition of design variance in the formula of the standard error of a given contrast, we get:
\begin{align}
\mathrm{SE}_{\mathbf{c}\hat{\beta}} = \sqrt{\mathrm{noise} \cdot \mathrm{design\ variance}} = \sqrt{\hat{\sigma}^{2}\mathbf{c}(\mathbf{X}^{T}\mathbf{X})^{-1}\mathbf{c}^{T}}
\end{align}
Now, we can write out the entire formula for the *t*-statistic:
\begin{align}
t_{\mathbf{c}\hat{\beta}} = \frac{\mathbf{c}\hat{\beta}}{\sqrt{\hat{\sigma}^{2}\mathbf{c}(X'X)^{-1}\mathbf{c}'}} = \frac{\mathrm{effect}}{\sqrt{\mathrm{noise} \cdot \mathrm{design\ variance}}}
\end{align}
---
\* Actually, design variance does not depend on the "variance" and "covariance", but on the sums-of-squares of each predictor $j$ ($\mathrm{SS}_{X_{j}}$) and sums-of-squares cross-products ($\mathrm{SS}_{X_{j}, X_{k}}$), respectively. These are just the variance and covariance terms, but without dividing by $N - 1$! We used the terms variance and covariance here because they are more intuitive.
### Summary: effects, noise, and design variance
Alright, that's a lot of math. Sorry about that. But the above formula nicely illustrates that, to obtain large effects (i.e. *t*-values), you need three things:
1. A large response/effect (i.e. $\beta$)
2. An efficient design or, in other words, low design variance (i.e. high variance, low covariance: $\frac{1}{c(X^{T}X)^{-1}c'}$)
3. Low noise/unexplained variance (i.e. low $\mathrm{SSE}\ /\ \mathrm{DF}$)
This week, we'll discuss how to optimize (2): the efficiency from the design. Next week, we'll discuss how to minimize (3): noise (unexplained variance).
If you remember these three components and how they conceptually relate to the effect we want to measure (*t*-values), you understand the most important aspect of experimental design in fMRI! In the rest of the tutorial, we're going to show you **why** you want high variance and low covariance in your design ($X$) and **how** to achieve this by designing your experiment in a specific way.
<div class='alert alert-warning'>
<b>ToDo</b> (0 points)
In the previous section, you've seen a lot of math and definitions of (statistical) concepts. Especially the part on the inverses (e.g. efficiency is the inverse of design-variance, and vice versa).
It is important to understand how all the concepts (signal/beta, noise/SSE, design variance, efficiency) relate to each other and to the thing we're after: strong effects (high *t*-values)!
Therefore, we captured the *conceptual* formula in a function below named <tt>conceptual_tvalue_calculator</tt>, which takes three inputs — signal, noise, and design variance — and outputs the effect (*t*-value) and design efficiency.
In the cell below, we call the function with some particular values for the three input-arguments (<tt>SIGNAL</tt>, <tt>NOISE</tt>, <tt>DESIGN_VARIANCE</tt>). For this (ungraded) ToDo, try to change these input parameters and try to understand how changing the inputs changes the outputs!
</div>
```
def conceptual_tvalue_calculator(signal, noise, design_variance):
""" Calculate the effect (t-value) from the signal, noise, and design variance components.
Parameters
----------
signal : int/float
noise : int/float
design_variance : int/float
Returns
-------
effect : float
efficiency : float
"""
efficiency = 1 / design_variance
effect = signal / (noise * design_variance)
return effect, efficiency
# Change the capitalized variables to see what effect it has on the t-value and efficiency
SIGNAL = 0.5
NOISE = 2.8
DESIGN_VARIANCE = 0.02
effect, efficiency = conceptual_tvalue_calculator(signal=SIGNAL, noise=NOISE, design_variance=DESIGN_VARIANCE)
print("Effect ('t-value'): %.3f" % effect)
print("Efficiency: %.3f" % efficiency)
```
<div class='alert alert-info'>
<b>ToThink</b> (1 point): Researchers do not need to acquire (fMRI) data ($\mathbf{y}$) to calculate the efficiency of their design ($\mathbf{X}$). Why?
</div>
YOUR ANSWER HERE
### How to calculate design variance and efficiency in Python
As discussed in the previous section, the formula for design variance (and efficiency) is often expressed using linear algebra notation:
\begin{align}
\mathrm{design\ variance} = \mathbf{c}(\mathbf{X}^{T}\mathbf{X})^{-1}\mathbf{c}^{T}
\end{align}
You have seen the $(\mathbf{X}^{T}\mathbf{X})$ earlier when we discussed the solution for finding the least squares solution. Now, design variance is calculated by pre and postmultiplying this term with the (transpose of the) contrast vector (denoted with `c` and `c.T`). As such, so the full design variance calculation can be implemented in python as follows:
```python
design_var = c @ inv(X.T @ X) @ c.T
```
Given that efficiency is the inverse of design variance:
\begin{align}
\mathrm{efficiency} = \frac{1}{\mathbf{c}(\mathbf{X}^{T}\mathbf{X})^{-1}\mathbf{c}^{T}}
\end{align}
... we can calculate efficiency as:
```python
efficiency = 1.0 / c @ inv(X.T @ X) @ c.T
```
You'll have to implement this yourself in a later ToDo! But first, let's go into more detail *why* high variance and low covariance are important to get large effects!
## The effect of predictor variance on design variance/efficiency
As explained in the previous section, design variance depends on (1) predictor variance and (2) predictor covariance. In this section, we'll focus on predictor variance. In the next section, we'll focus on predictor covariance.
As you probably know, *variance* is a statistical property of a random variable that describes the average squared deviation from the variable's mean. Formally, for any variable $x$ with mean $\bar{x}$ and length $N$, its sample variance is defined as:
\begin{align}
\mathbf{var}[x] = \frac{1}{N - 1}\sum_{i=1}^{N}(x - \bar{x})^{2}
\end{align}
So, the more values of a variable deviate from its mean on average, the more variance it has.
To demonstrate the effect of predictor variance on design variance/efficiency, we will focus (for simplicity) on non-time series designs that have just a single condition and thus a single predictor (apart from the intercept). In these examples, we'll focus on why high variance is important.
### An example of the effect of (high) design variance
To start, we want to show you — conceptually — why it is important to have a lot of variance in your predictors for a low standard error of your beta, and thus high t-values. We're going to show you an example of 'regular' linear regression (so no time-series signal, but the example holds for MRI data).
Suppose we want to investigate the effect of someone's IQ on their income ($\mathbf{X} = IQ$, $\mathbf{y} = income$). We've gathered some data, which we'll as usual represent as an independent variable ($\mathbf{X}$) and a dependent variable ($\mathbf{y}$). We'll also run the regression analysis and calculate the beta-parameters, MSE and *t*-value (corresponding to the IQ-parameter "against baseline").
```
from niedu.utils.nii import calculate_stats_for_iq_income_dataset
# Load the data
iq_income_data = np.load('iq_variance_example.npz')
X_lowvar = iq_income_data['X_lv']
y_lowvar = iq_income_data['y_lv']
print("Shape X: %s" % (X_lowvar.shape,))
print("Shape y: %s" % (y_lowvar.shape,))
beta_lv, mse_lv, tval_lv = calculate_stats_for_iq_income_dataset(iq_income_data, which='lowvar')
plt.figure(figsize=(7, 7))
plt.title("Relation between IQ (X) and income (y)", fontsize=15)
plt.scatter(X_lowvar[:, 1], y_lowvar, c='tab:blue')
plt.ylabel('Income (x 1000 euro)', fontsize=20)
plt.xlabel('IQ', fontsize=20)
plt.plot(
(116, 128),
(116 * beta_lv[1] + beta_lv[0], 128 * beta_lv[1] + beta_lv[0]), c='tab:orange', lw=3
)
plt.text(118, 66, r'$\hat{\beta}_{IQ} = %.3f$' % beta_lv[1], fontsize=15)
plt.text(118, 65, 'MSE = %.3f' % mse_lv, fontsize=15)
plt.grid()
plt.xlim(116, 128)
plt.show()
```
This is pretty awesome data! On average, our prediction is on average less than 1 point off (i.e., $\mathrm{MSE} < 1$)! But you might also have noticed that the *range* of values for $X$ (i.e., IQ) is quite limited: we only measured people with IQs between about 118 and 127. This is quite a narrow range — in other words: little variance — knowing that IQ varies according to a normal distribution with mean 100 and standard deviation 15. In other words, we have a pretty good model, but it is only based on a specific range of the IQ-variable.
Think about it this way: this model captures the relationship between IQ and income, but only for relatively high-intelligence people. Sure, you can extrapolate to IQ-values like 80 and 90, but this extrapolation is quite uncertain because you've never even measured someone with that IQ-value!
So, for comparison, let's a similar dataset with IQ and income, but this time with a much larger range of the IQ-variable. We'll plot the two datasets (the low-variance and high-variance data) next to each other.
```
X_highvar = iq_income_data['X_hv']
y_highvar = iq_income_data['y_hv']
x_lim = (65, 130)
y_lim = (0, 80)
plt.figure(figsize=(15, 7))
plt.subplot(1, 2, 1)
plt.title("Low-variance data (zoomed out)", fontsize=20)
plt.scatter(X_lowvar[:, 1], y_lowvar, c='tab:blue')
plt.ylabel('Income (x 1000 euro)', fontsize=20)
plt.xlabel('IQ', fontsize=20)
plt.xlim(x_lim)
plt.ylim(y_lim)
plt.plot(x_lim, (x_lim[0] * beta_lv[1] + beta_lv[0], x_lim[1] * beta_lv[1] + beta_lv[0]), c='tab:orange', lw=3)
plt.text(70, 70, r'$\hat{\beta}_{IQ} = %.3f$' % beta_lv[1], fontsize=20)
plt.text(70, 65, 'MSE = %.3f' % mse_lv, fontsize=20)
plt.grid()
# Now, do the same calculations for the highvar data
beta_hv, mse_hv, tval_hv = calculate_stats_for_iq_income_dataset(iq_income_data, which='highvar')
plt.subplot(1, 2, 2)
plt.title("High-variance data", fontsize=20)
plt.scatter(X_highvar[:, 1], y_highvar, c='tab:blue')
plt.xlim(x_lim)
plt.ylim(y_lim)
plt.xlabel('IQ', fontsize=20)
plt.plot(x_lim, (x_lim[0] * beta_hv[1] + beta_hv[0], x_lim[1] * beta_hv[1] + beta_hv[0]), c='tab:orange', lw=3)
plt.text(70, 70, r'$\hat{\beta}_{IQ} = %.3f$' % beta_hv[1], fontsize=20)
plt.text(70, 65, 'MSE = %.3f' % mse_hv, fontsize=20)
plt.tight_layout()
plt.grid()
plt.show()
```
As you can see from the plots of the two datasets side-by-side, both the low-variance (left) and the high-variance plot (right) capture approximately the model: for each increase in an IQ-point, people earn about 1000 (low-variance model) / 959 (high-variance model) euro extra (these are reflected by the beta-parameters!).
But, you also see that the MSE for the high-variance model is *much* higher, which is also evident from the residuals (distance of the red points from the blue line).
```
mse_ratio = mse_hv / mse_lv
print("The MSE of the high-variance data is %.3f times larger than the low-variance data!" % mse_ratio)
```
Given these statistics, you might guess that the t-value of the IQ-parameter in the high-variance model would be way lower than the same parameter in the low-variance model, right? Well, let's check it out:
```
# We calcalated the t-values earlier with the calculate_stats_for_iq_income_dataset function
print("T-value low-variance model: %.3f" % tval_lv)
print("T-value high-variance model: %.3f" % tval_hv)
```
You probably by now understand what's the culprit: the design-variance! Given that the effect ($\hat{\beta}_{IQ}$) is about the same for the two models and the MSE is higher for the high-variance model, the logical conclusion is that *the design-variance of the high-variance model must be waaaaay lower*.
<div class='alert alert-warning'>
<b>ToDo</b> (2 points): Use the two design-matrices (<tt>X_highvar</tt> and <tt>X_lowvar</tt>) to calculate the design-variances of both the low-variance and the high-variance dataset for the "contrast against baseline", i.e., $H_{0}: \beta_{IQ} = 0$ and $H_{a}: \beta_{IQ} \neq 0$. Then, divide the design-variance of the low-variance dataset by the high-variance dataset and store this in the variable <tt>desvar_ratio</tt> (this indicates how much higher the design-variance of the low-variance dataset is compared to the high-variance dataset). Make sure to use an appropriate contrast-vector!
</div>
```
# Implement your ToDo here
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo. '''
from niedu.tests.nii.week_3 import test_lowvar_vs_highvar_iq_design
test_lowvar_vs_highvar_iq_design(X_lowvar, X_highvar, desvar_ratio)
```
<div class='alert alert-warning'>
<b>ToDo</b> (1 point): Design efficiency (and design-variance) is a metric without a clear unit of measurement; herefore, efficiency (and design variance) should always be interpreted in relative terms. To show this, we are going to look at the weight-height example from last week, in which we used weight as a predictor for height. Now, we're going to rescale the predictor ('weight') such that it represents weight in <em>grams</em> instead of *kilos* (as was originally the case).
Calculate efficiency for both the weight-in-kilos data (<tt>X_kilos</tt>) and the weight-in-grams data (<tt>X_grams</tt>). Store the efficiency for the weight-in-kilos data in a variable named <tt>efficiency_kilos</tt> and the efficiency for the weight-in-grams data in a variable named <tt>efficiency_grams</tt>.
</div>
```
with np.load('weight_height_data.npz') as data:
X_kilos = data['X']
X_grams = X_kilos * 1000
# We'll stack an intercept for you!
intercept = np.ones((X_kilos.size, 1))
X_kilos = np.hstack((intercept, X_kilos))
X_grams = np.hstack((intercept, X_grams))
# Start your ToDo here
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo. '''
from niedu.tests.nii.week_3 import test_design_variance_scaling
test_design_variance_scaling(X_kilos, X_grams, efficiency_kilos, efficiency_grams)
```
<div class='alert alert-info'>
<b>ToThink</b> (1 point): If you've done the above ToDo correctly, you should see that (everything else being equal) the design with weight in grams is a 1,000,000 times more efficient than the design with weight in kilos.
Why is the efficiency a million times higher and not 1000 times higher (as the scale-difference would suggest)?
</div>
YOUR ANSWER HERE
## The effect of predictor covariance on design variance/efficiency
### Multicollinearity
In the previous section, we discussed the influence of predictor variance - $\mathrm{var}[\mathbf{X}_{j}]$ - on the design-variance term, showing that high variance leads to (relatively) low design variance (and thus high efficiency).
We know, however, that design variance *also* depends on the *covariance* between predictors - $\mathrm{cov}[\mathbf{X}_{j}, \mathbf{X}_{k}]$. This "covariance between predictors" is also known as **multicollinearity**.
Specifically, the **higher** the covariance (multicollinearity), the **lower** the design efficiency (the worse our design is). Conceptually, you can think of high covariance between predictors as causing *uncertainty* of the estimation of your beta-estimates: if your predictors are correlated, the GLM "doesn't know" what (de)activation it should assign to which predictor. This uncertainty due to correlated predictors is reflected in a (relatively) higher design variance term.
Anyway, let's look at some (simulated) data. This time (unlike the variance-example), we're going to look at fMRI timeseries data. We'll simulate a design with two predictors, we'll calculate the correlation between the two predictors, and the efficiency of the design for the difference contrast between the predictors (`c = [0, 1, -1]`).
```
def simulate_two_predictors(N=360, shift=30, TR=2):
''' Simulates two predictors with evenly spaced trials,
shifted a given number of time-points. '''
offset = 20
stop = 300
space = 60
pred1 = np.zeros(N)
pred1[offset:stop:space] = 1
pred2 = np.zeros(N)
pred2[(offset + shift):stop:space] = 1
hrf = glover_hrf(tr=1, oversampling=1)
hrf /= hrf.max()
pred1 = np.convolve(pred1, hrf)[:N:int(TR)]
pred2 = np.convolve(pred2, hrf)[:N:int(TR)]
X = np.hstack((np.ones((int(N / 2), 1)), pred1[:, np.newaxis], pred2[:, np.newaxis]))
return X
# We set the "shift" (distance between predictor 1 and 2 to 30 seconds)
X = simulate_two_predictors(N=350, shift=30, TR=2)
cvec = np.array([0, 1, -1])
corr = pearsonr(X[:, 1], X[:, 2])[0]
eff = 1.0 / cvec.dot(np.linalg.inv(X.T.dot(X))).dot(cvec.T)
plt.figure(figsize=(20, 5))
plt.plot(X[:, 1])
plt.plot(X[:, 2])
plt.text(150, 0.8, 'Corr predictors: %.3f' % corr, fontsize=14)
plt.text(150, 0.7, 'Efficiency: %.3f' % eff, fontsize=14)
plt.xlim(0, 175)
plt.legend(['Predictor 1', 'Predictor 2'], loc='lower right')
plt.xlabel("Time (volumes)")
plt.ylabel("Activation (A.U.)")
plt.title("Almost no collinearity", fontsize=20)
plt.grid()
plt.show()
print("Variance predictor 1: %.3f" % np.var(X[:, 1], ddof=1))
print("Variance predictor 2: %.3f" % np.var(X[:, 2], ddof=1))
```
As you can see, the predictors are almost perfectly uncorrelated - $\mathrm{corr}(\mathbf{X}_{1}, \mathbf{X}_{2}) \approx 0$ - which corresponds to a design efficiency of 4.539. Remember, the absolute value of efficiency is not interpretable, but we can interpret it *relative to other designs*. As such, we can investigate how a design with more correlated predictors will change in terms of efficiency.
To do so, we can simply "shift" the second predictor (blue line) to the left (i.e., the stimuli of predictor 2 follow the stimuli of predictor 1 more quickly). Let's check out what happens to the efficiency if we induce corelation this way:
```
# We set shift to 4 seconds (instead of 30 like before)
X2 = simulate_two_predictors(N=350, shift=2, TR=2)
corr2 = pearsonr(X2[:, 1], X2[:, 2])[0]
eff2 = 1.0 / cvec.dot(np.linalg.inv(X2.T.dot(X2))).dot(cvec.T)
plt.figure(figsize=(20, 5))
plt.plot(X2[:, 1])
plt.plot(X2[:, 2])
plt.text(150, 0.8, 'Corr predictors: %.3f' % corr2, fontsize=14)
plt.text(150, 0.7, 'Efficiency: %.3f' % eff2, fontsize=14)
plt.xlim(0, 175)
plt.legend(['Predictor 1', 'Predictor 2'], loc='lower right', frameon=False)
plt.xlabel("Time (volumes)", fontsize=20)
plt.ylabel("Activation (A.U.)", fontsize=20)
plt.title("Quite a bit of collinearity", fontsize=25)
plt.grid()
plt.show()
print("Variance predictor 1: %.3f" % np.var(X2[:, 1], ddof=1))
print("Variance predictor 2: %.3f" % np.var(X2[:, 2], ddof=1))
# Let's calculate the reduction in efficiency
reduction_eff = ((eff - eff2) / eff) * 100
print("Efficiency is reduced with %.1f%% when increasing the correlation to %.3f" % (reduction_eff, corr2))
```
As you can see, increasing correlation between predictors has the effect of reducing efficiency, even if the predictor variance stays the same! Like we discussed earlier, this is because correlation between predictors reflects *ambiguity* about the "source" of an effect.
To get a better intuition of this ambiguity, suppose that for the above design (with the correlated predictors), we observe the following signal (we just simulate the signal as the linear sum of the predictors + noise; sort of a "reverse linear regression"):
```
np.random.seed(42)
# Here we simulate a signal based on the predictors + noise
some_noise = + np.random.normal(0, 0.3, X2.shape[0])
sim_signal = X2[:, 1] * 2 + X2[:, 2] * 2 + some_noise
plt.figure(figsize=(15, 5))
plt.plot(X2[:, 1])
plt.plot(X2[:, 2])
plt.plot(sim_signal)
plt.xlim(0, 175)
plt.legend(['Predictor 1', 'Predictor 2', 'Signal'], loc='upper right', frameon=False)
plt.xlabel("Time (volumes)", fontsize=20)
plt.ylabel("Activation (A.U.)", fontsize=20)
plt.title("Simulated data + multicollinear predictors", fontsize=25)
plt.grid()
plt.show()
```
Now, if we calculate the beta-parameters of both predictors, we see that they both are given approximately equal "importance" (i.e., their beta-parameters are about equally high):
```
betas = inv(X2.T @ X2) @ X2.T @ sim_signal
print("Betas (w/o intercept): %r" % betas[1:])
```
However, it is unclear to the GLM whether the peaks in the signal (green line) are caused by predictor 1 or predictor 2! While the betas themselves are not affected on average (i.e., there is no *bias*), this "uncertainty" (or "ambiguity") is reflected in the GLM through a relatively higher design variance term, that will subsequently lead to (relatively) lower *t*-values!
<div class='alert alert-info'>
<b>ToThink</b> (1 point): Suppose that due to a mistake in your experimental paradigm, you actually present the two classes of stimuli (reflecting predictor 1 and predictor 2 in the above example) at the same time (blue and orange predictors completely overlap). As it turns out, you cannot (reliably) calculate the design variance for such a design. Explain concisely why this is the case.
</div>
YOUR ANSWER HERE
### Evaluating multiple contrasts
Thus far, we only evaluated the efficiency for a *single* contrast, like one particular predictor against baseline, e.g. `contrast_vec = np.array([0, 1, 0])`. Often, though, you might be interested in *more than one contrast*. For example, you might be interested in the contrast of predictor "A" against baseline, predictor "B" against baseline, and the difference between predictor "A" and "B"\*. We can simply extend our formula for efficiency to allow more than one contrast. For $K$ contrasts, efficiency is defined as:
\begin{align}
\mathrm{efficiency} = \frac{K}{\sum_{k=1}^{K} \mathbf{c}_{k}(\mathbf{X}^{T}\mathbf{X})^{-1}\mathbf{c}_{k}^{T}}
\end{align}
This specific calculation of efficiency is also referred to as "A optimality". From the formula, you can see that the overall efficiency for multiple contrasts is basically (but not precisely) the "average" of the efficiencies for the individual contrasts.
Let's practice the Python-implementation of overall efficiency for multiple contrasts in a short assignment (graded with hidden tests).
---
\* Note that evaluating different contrasts separately is not the same as doing an F-test (like we discussed in week 2)!
<div class='alert alert-warning'>
<b>ToDo</b> (1 point):
With the data from the correlation-simulation (i.e., the variable `X`), calculate the efficiency for the set of the following contrasts:
- predictor 1 against baseline
- predictor 2 against baseline
- prediction 1 - predictor 2
You have to define the contrasts yourself.
Store the overall efficiency in a variable named <tt>overall_eff</tt>. Hint: you probably need a for loop (or a list comprehension). Hint 2: don't forget the intercept in your contrast-definitions (it's the first column).
</div>
```
# Implement your ToDo here using X
X = simulate_two_predictors(N=350, shift=30, TR=2)
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo. '''
from niedu.tests.nii.week_3 import test_overall_eff
test_overall_eff(X, overall_eff)
```
## How should we design our experiment to maximize efficiency?
Alright, thus far we talked about **why** you want high predictor variance and low predictor covariance for optimal estimation of effects (i.e., t-values). But this leaves us with the question: **how** should we design our experiment such that it has high variance and low covariance?
The answer is (as is often the case): it depends.
### Estimation vs. detection
The specific design of your experiment (how the stimuli are ordered and their timing) mainly depends on the *type of question* you're investigating with your fMRI experiment. These types of questions are usually divided into two categories within (univariate) fMRI studies:
**1) You want to know whether different conditions activate voxel activity differently.**
We've learned how to do this in week 2: essentially, you want to estimate just a beta-parameter (reflecting activation/deactivation) of your stimulus-regressors. This is a question about HRF-*amplitude* only. The far majority of fMRI research falls in this category (and it's the type we'll focus on in this course). It is often said that this type of research focuses on **detection** of a signal's response. For this question, designs are often based on canonical HRF-based convolution (or based on a basis set).
**2) You want investigate how different conditions influence voxel activity not only by investigating the "amplitude" parameter of the HRF, but also parameters relating to other properties of the shape of the HRF (like width, lag, strenght of undershoot, etc.).**
A small proportion of fMRI studies have this goal. Examples are studies that investigate [how the shape of the HRF changes with age](http://www.sciencedirect.com/science/article/pii/S1053811907010877) or that investigate [differences in HRF shape across clinical populations](http://www.sciencedirect.com/science/article/pii/S1053811907001371). We won't focus on this type of research in this course, but you should know that you can also investigate other parameters of the HRF in neuroimaging research! It if often said that this type of research focuses on **estimation** of the (shape of the) signal's response. Often, "finite impulse reponse" (FIR) models are used for these types of studies (which you might have seen in the videos).
The far majority of the fMRI studies focus on questions about **detection**, i.e., based on analysis of the "amplitude" of the HRF. This is why we won't discuss the estimation approach (and associated models, like the FIR-based GLM) in the rest of this lab and the course in general.
Now, given that we aim for detection, how should we design our experiment? Well, there are two "main" types of designs: event-related designs and blocked designs, which are discussed in the next section.
### Event-related vs. blocked designs
As you've probably read in the book or seen in the videos, event-related and blocked designs differ in the *ordering* of the stimuli. Basically, event-related designs are designs in which the stimuli from different conditions are ordered randomly, while blocked designs are designs in which the stimuli of the same condition are grouped together in "blocks". Below, we visualized an example of each design side-by-side:
```
np.random.seed(2)
N = 180
dg_hrf = glover_hrf(tr=1, oversampling=1)
blocked_pred1_onsets = list(range(10, 30)) + list(range(90, 110))
blocked_pred2_onsets = list(range(50, 70)) + list(range(130, 150))
N_stim = len(blocked_pred1_onsets)
blocked_pred1, blocked_pred2 = np.zeros(N), np.zeros(N)
blocked_pred1[blocked_pred1_onsets] = 1
blocked_pred2[blocked_pred2_onsets] = 1
icept = np.ones((N, 1))
X_blocked = np.hstack((
icept,
np.convolve(blocked_pred1, dg_hrf)[:N, np.newaxis],
np.convolve(blocked_pred2, dg_hrf)[:N, np.newaxis]
))
plt.figure(figsize=(20, 10))
plt.subplot(2, 2, 1)
plt.title("Event onsets (BLOCKED)", fontsize=20)
plt.xlim(0, N)
plt.axhline(0, c='tab:blue')
plt.grid()
for onset in blocked_pred1_onsets:
plt.plot((onset, onset), (0, 1), 'k-', c='tab:blue')
for onset in blocked_pred2_onsets:
plt.plot((onset, onset), (0, 1), 'k-', c='tab:orange')
plt.subplot(2, 2, 3)
plt.xlim(0, N)
plt.title("Convolved predictors (BLOCKED)", fontsize=20)
plt.ylim(-1, 2)
plt.plot(X_blocked[:, 1], c='tab:blue')
plt.plot(X_blocked[:, 2], c='tab:orange')
plt.grid()
plt.xlabel("Time (volumes)", fontsize=15)
er_stims = np.arange(N)
er_pred1_onsets = np.random.choice(er_stims, N_stim, replace=False)
er_stims_new = np.array([o for o in er_stims if o not in er_pred1_onsets])
er_pred2_onsets = np.random.choice(er_stims_new, N_stim, replace=False)
er_pred1, er_pred2 = np.zeros(N), np.zeros(N)
er_pred1[er_pred1_onsets] = 1
er_pred2[er_pred2_onsets] = 1
plt.subplot(2, 2, 2)
plt.xlim(0, N)
plt.title("Event onsets (EVENT-RELATED)", fontsize=20)
plt.axhline(0, c='tab:blue')
plt.grid()
for onset in er_pred1_onsets:
plt.plot((onset, onset), (0, 1), 'k-', c='tab:blue')
for onset in er_pred2_onsets:
plt.plot((onset, onset), (0, 1), 'k-', c='tab:orange')
X_er = np.hstack((
icept,
np.convolve(er_pred1, dg_hrf)[:N, np.newaxis],
np.convolve(er_pred2, dg_hrf)[:N, np.newaxis]
))
plt.subplot(2, 2, 4)
plt.title("Convolved predictors (EVENT-RELATED)", fontsize=20)
plt.ylim(-1, 2)
plt.plot(X_er[:, 1], c='tab:blue')
plt.plot(X_er[:, 2], c='tab:orange')
plt.axhline(0, ls='--', c='k')
plt.xlim(0, N)
plt.grid()
plt.xlabel("Time (volumes)", fontsize=15)
plt.tight_layout()
plt.show()
```
As you can see in the plot above, a blocked design groups trials of the same condition together in blocks, while the event-related design is completely random in the sequence of trials. Note that designs can of course also be a "mixture" between blocked and event-related (e.g., largely random with some "blocks" in between).
So, if we're interested in detection (i.e., the amplitude of the response), what should we choose?
Well, the answer is simple: **blocked designs**.
This is because blocked designs simply (almost always) have lower design variance because of:
- lower covariance ("correlation")
- higher variance ("spread")
Let's check this for the designs from the plot. First, we'll look at the predictor covariance, but because predictor correlation is often more interpretable (correlation = standardized covariance), we'll calculate that instead:
```
corr_blocked = pearsonr(X_blocked[:, 1], X_blocked[:, 2])
corr_er = pearsonr(X_er[:, 1], X_er[:, 2])
print("Correlation blocked: %.3f. Correlation event-related: %.3f" % (corr_blocked[0], corr_er[0]))
```
<div class='alert alert-warning'>
<b>ToDo</b> (1 point):
We've seen that predictor correlation is lower in blocked designs than in event-related designs. But what about predictor variance? Calculate predictor variance for predictor 1 (column 2) and predictor 2 (column 3) for both the blocked design (<tt>X_blocked</tt>) and the event-related design (<tt>X_er</tt>).
Remember: (sample) variance is the summed squared deviation of values from a variable's mean divided by the number of observations minus 1, or formally:
\begin{align}
\mathbf{var}[x] = \frac{1}{N - 1}\sum_{i=1}^{N}(x - \bar{x})^{2}
\end{align}
Store the variance of the four predictors (2 predictions $\times$ 2 designs) in the following variables:
- <tt>blocked_pred1_var</tt>
- <tt>blocked_pred2_var</tt>
- <tt>er_pred1_var</tt>
- <tt>er_pred2_var</tt>
Note: do **not** use the numpy function <tt>np.var</tt> for this ToDo (also because it's going to give you the wrong answer).
</div>
```
# Implement your ToDo here
# YOUR CODE HERE
raise NotImplementedError()
''' Tests the above ToDo. '''
from niedu.tests.nii.week_3 import test_variance_computation
test_variance_computation(X_blocked, X_er, blocked_pred1_var, blocked_pred2_var, er_pred1_var, er_pred2_var)
```
<div class='alert alert-info'>
<b>ToThink</b> (1 point): One property/characteristic of the BOLD-response is especially important in generating high predictor variance, which becomes especially clear in blocked designs. Which property is this? Write your answer in the cell below.
</div>
YOUR ANSWER HERE
<div class='alert alert-warning'>
<b>ToDo/ToThink</b> (2 points)
As you've seen in the previous ToDo, blocked designs have larger predictor variance and (everything else being equal) are more efficient. In fact, up to a certain point, the larger the blocks of trials, the more efficient the design. This may, at first sight, reflect idea in most psychological research that more trials (events) lead to more power.
In fMRI designs, however, this is *not* the case, because at a certain point, longer blocks yield a *less* efficient design. Below, we define a function that simulates a single predictor for a blocked-design with a variable amount of trials in it (for a fixed experiment duration of 500 seconds and a TR of 1). So, for example, if we call the function with <tt>trials=10</tt>, it will create a design and predictor with a block of 10 consecutive stimuli (all lasting a second). We'll also plot the predictor after simulating the data.
Now, suppose I would like to evaluate the contrast of that single predictor against baseline. In the text-cell below, argue why adding more trials does not necessarily mean a more efficient design, assuming some fixed length of the experiment.
Hint: set the number of trials very high (e.g. <tt>N_TRIALS = 500</tt>) and see what happens with the predictor.<br>
</div>
```
def simulate_single_predictor(trials, time_exp=500):
if trials > time_exp:
raise ValueError("Cannot have more trials than timepoints!")
pred = np.zeros(time_exp)
onsets = np.arange(trials)
pred[onsets] = 1
dg = glover_hrf(tr=1, oversampling=1)
pred_conv = np.convolve(pred, dg)[:time_exp, np.newaxis]
X = np.hstack((np.ones((time_exp, 1)), pred_conv))
return X
# We'll call the function above here:
contrast = np.array([0, 1])
# you can change this variable to investigate the effect of increasing/decreasing the amount of trials
N_TRIALS = 1
X = simulate_single_predictor(trials=N_TRIALS)
# ... and plot the predictor
plt.figure(figsize=(15, 5))
plt.plot(X[:, 1])
plt.xlim(0, 500)
plt.title("Simulated design/predictor with %i trials" % N_TRIALS, fontsize=25)
plt.xlabel("Time (seconds/volumes)", fontsize=20)
plt.ylabel("Activation (A.U.)", fontsize=20)
plt.grid()
plt.show()
```
YOUR ANSWER HERE
### The "paradox" of efficiency
So, we've discussed blocked and event-related designs and we've come to the conclusion that blocked designs are simply more efficient than event-related designs. "So, we should always use blocked designs?", you may ask.
Well, no.
We've discussed the mathematics behind design variance, efficiency, and t-value in detail, but we shouldn't forget that ultimately **we're measuring the data from a living human beings in the MRI-scanner**, who tend to get bored, fall asleep, and otherwise not pay attention if the task they're doing is monotonous, predictable or simply uninteresting!
Blocked designs, however, are (usually) exactly this: designs that are experienced as predictable, monotonous, and (relatively) boring! Like we said earlier, the effects we're going to measure depend on three things - effects, noise, and design efficiency - and psychological factors may strongly influence the "effect" part and thus affect the statistics we're interested in (i.e., t-values). In addition to psychological factors like boredom and inattention, blocked designs may also lead to unwanted effects like habituation (attenuation of the BOLD-response after repeated stimulation), which violate the assumption of the BOLD-response as being 'linear time-invariant' (LTI). In other words, the BOLD-response may stop 'behaving' like we assume it behaves when we use blocked designs.
This is, essentially, the paradox of designing fMRI experiments: the most efficient designs are also the designs that (potentially) lead to the lowest signal or otherwise unintended effects (due to boredom, predictability, habituation, etc.).
So, what do we do in practice? Usually, we use (semi-random) event-related designs. We lose some efficiency by using event-related designs instead of blocked designs, but we reduce the chance of psychological factors and other effects that reduce the measured signal or mess with the assumption of linear time-invariance.
Given that we're going to use some event-related (i.e., "random") design, let's investigate how we can optimize this type of design.
### Improving design efficiency for event-related designs using jittering
Usually, events in an experimental design are separated by short periods without any event; this is called the "inter-stimulus interval" (ISI; also called stimulus onset asynchrony, SOA) - the time between successive stimuli. For example, the experiment from the image below has an ISI of 8 seconds:
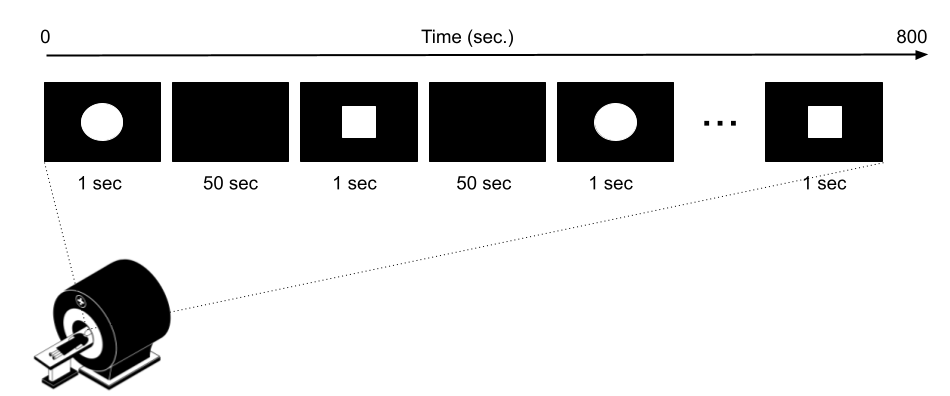
Let's simulate some event-onsets and predictors for this experiment. We have two predictors (circles and squares). The stimuli ('events') take 1 second and the ISI is 8 seconds (like the figure above). Suppose we're interested in both contrasts against baseline (circles against baseline; squares against baseline).
Now, let's simulate one design (with a fixed ISI), calculate efficiency and plot it:
```
def simulate_data_fixed_ISI(N=420):
dg_hrf = glover_hrf(tr=1, oversampling=1)
# Create indices in regularly spaced intervals (9 seconds, i.e. 1 sec stim + 8 ISI)
stim_onsets = np.arange(10, N - 15, 9)
stimcodes = np.repeat([1, 2], stim_onsets.size / 2) # create codes for two conditions
np.random.shuffle(stimcodes) # random shuffle
stim = np.zeros((N, 1))
c = np.array([[0, 1, 0], [0, 0, 1]])
# Fill stim array with codes at onsets
for i, stim_onset in enumerate(stim_onsets):
stim[stim_onset] = 1 if stimcodes[i] == 1 else 2
stims_A = (stim == 1).astype(int)
stims_B = (stim == 2).astype(int)
reg_A = np.convolve(stims_A.squeeze(), dg_hrf)[:N]
reg_B = np.convolve(stims_B.squeeze(), dg_hrf)[:N]
X = np.hstack((np.ones((reg_B.size, 1)), reg_A[:, np.newaxis], reg_B[:, np.newaxis]))
dvars = [(c[i, :].dot(np.linalg.inv(X.T.dot(X))).dot(c[i, :].T))
for i in range(c.shape[0])]
eff = c.shape[0] / np.sum(dvars)
return X, eff
X, eff = simulate_data_fixed_ISI()
plt.figure(figsize=(15, 5))
plt.title('Fixed ISI of 8 seconds (Efficiency = %.3f)' % eff, fontsize=20)
plt.plot(X[:, 1])
plt.plot(X[:, 2])
plt.legend(['Predictor A', 'Predictor B'])
plt.ylabel('Amplitude (a.u.)')
plt.xlabel('Time (TR)')
plt.xlim(0, N)
plt.grid()
plt.show()
```
Often, though, researchers do not use a *fixed* ISI, but they vary the ISI from trial to trial. This process is called "jittering". Usually, the ISIs are drawn randomly from a known distribution (e.g., truncated exponential or normal distribution). Compared to using fixed ISIs, jittering may yield more efficient designs by reducing covariance and increasing predictor variance. Let's simulate another dataset, but this time with a variable ISI between 2-6 seconds (which is on average 4 seconds, but variable across trials):
```
def simulate_data_jittered_ISI(N=420):
dg_hrf = glover_hrf(tr=1, oversampling=1)
stim_onsets = np.arange(10, N - 15, 9)
stimcodes = np.repeat([1, 2], stim_onsets.size / 2)
np.random.shuffle(stimcodes)
# Here, we pick some *deviations* from the standard ISI (i.e., 8),
# so possible ISIs are (8 - 2, 8 - 1, 8 - 0, 8 + 1, 8 + 2)
ISIs = np.repeat([-2, -1, 0, 1, 2], repeats=11)
np.random.shuffle(ISIs)
stim = np.zeros((N, 1))
c = np.array([[0, 1, 0], [0, 0, 1]])
for i, stim_onset in enumerate(stim_onsets):
# We subtract the stim-onset with -2, -1, 0, 1, or 2 (from ISIs)
# to simulate jittering
stim[stim_onset - ISIs[i]] = 1 if stimcodes[i] == 1 else 2
stims_A = (stim == 1).astype(int)
stims_B = (stim == 2).astype(int)
reg_A = np.convolve(stims_A.squeeze(), dg_hrf)[:N]
reg_B = np.convolve(stims_B.squeeze(), dg_hrf)[:N]
X = np.hstack((np.ones((reg_B.size, 1)), reg_A[:, np.newaxis], reg_B[:, np.newaxis]))
# Loop over the two contrasts
dvars = [(c[i, :].dot(np.linalg.inv(X.T.dot(X))).dot(c[i, :].T))
for i in range(c.shape[0])]
eff = c.shape[0] / np.sum(dvars)
return X, eff
plt.figure(figsize=(15, 5))
X, eff = simulate_data_jittered_ISI()
plt.title('Variable (jittered) ISI of 2-6 seconds (Efficiency = %.3f)' % eff, fontsize=20)
plt.plot(X[:, 1])
plt.plot(X[:, 2])
plt.legend(['Predictor A', 'Predictor B'])
plt.ylabel('Amplitude (a.u.)')
plt.xlabel('Time (TR)')
plt.xlim(0, N)
plt.grid()
plt.show()
```
As you can see in the plot above, jittering improved design efficiency quite a bit! It is important to realize that jittering does not *always* improve design efficiency, but by "injecting" randomness (by selecting semi-random ISIs) it allows for *a larger variety* of designs, which also include designs that happen to be more efficient than the fixed-ISI designs.
You'll follow up on this idea in the next ToDo.
<div class='alert alert-warning'>
<b>ToDo</b> (3 points)
In the previous two examples (fixed-ISI and jittered ISI examples), we saw that the fixed-ISI design was less efficient than the jittered ISI design. In general, jittering increases the number of different designs you can simulate relative to fixed-ISI designs. A great way to visualize this is to simply run the simulation of fixed-ISI and jittered-ISI designs a number of times and plot the resulting efficiencies in two separate histograms. You should see that the histogram of efficiencies from fixed-ISI designs is quite a bit narrower than the histogram of efficiencies from jittered-ISI designs (but you might also also see that some jittered-ISI designs are *less* efficient that the average fixed-ISI design).
So, in this ToDo you will have to call the two simulation-functions (<tt>simulate_data_jittered_ISI</tt> and <tt>simulate_data_fixed_ISI</tt>) each 1000 times (use <tt>N=420</tt>, the default value) and keep track of the efficiency from both. Then, plot in *a single plot*, the histogram (using <tt>plt.hist</tt>) of the fixed-ISI efficiencies and the histogram of the jittered-ISI efficiencies. To plot two different histograms in a single plot, just call <tt>plt.hist</tt> twice (*before* calling <tt>plt.show</tt>); it will plot both histograms in the same plot.
Also, add a legend (showing which histogram refers to which efficiencies), give a sensible label to the x-axis and y-axis (if you don't know what the axes of a histogram refer to, look it up!), and give the plot a descriptive title.
So, in summary, you have to do the following:
- run the two functions 1000 times each
- with the resulting efficiency values (1000 for the fixed-ISI simulation function, and 1000 for the jittered-ISI simulation function), plot a histogram of the fixed-ISI efficiencies and a histogram of the jittered-ISI efficiencies in a single plot
- add a legend, labels for the axes, and a title
Hint: the functions output two things (the design and the efficiency); you only need the efficiency (second output)
This assignment is manually graded (no test-cell).
</div>
```
# implement your ToDo here
iterations = 1000
# YOUR CODE HERE
raise NotImplementedError()
```
### Summary: how to optimize your design for efficiency
So, in this section we discussed how to structure your experiment such that it yields a (relatively) high design efficiency, which will optimize our chance to find significant effects. How you do this depends on whether you aim for estimation (what is the shape of the HRF?) or for detection (what is the amplitude of the response?). Usually, we aim for detection; in that case, designs can be roughly grouped in two types: blocked designs an event-related designs. Purely statistically speaking, blocked designs are (almost always) more efficient, because they generally have lower covariance and higher variance than event-related designs. However, due to psychological factors and potential violations of the linear time-invariance of the BOLD-response, we often opt for event-related designs in the end. For event-related designs, we can increase our chance of finding a relatively efficient design by jittering our ISIs.
<div class='alert alert-success'>
<b>Tip!</b>
Before handing in your notebooks, we recommend restarting your kernel (<em>Kernel</em> → <em>Restart & Clear Ouput</em>) and running all your cells again (manually, or by <em>Cell</em> → <em>Run all</em>). By running all your cells one by one (from "top" to "bottom" of the notebook), you may spot potential errors that are caused by accidentally overwriting your variables or running your cells out of order (e.g., defining the variable 'x' in cell 28 which you then use in cell 15).
</div>
| github_jupyter |
# Time Series with Python
© Francesco Mosconi, 2016
## Regression
- detrending
- lagged variables
- train-test split
- validation
```
import pandas as pd
import numpy as np
%matplotlib inline
import matplotlib.pyplot as plt
from pylab import rcParams
rcParams['figure.figsize'] = 11, 7
```
## Regression
```
df = pd.read_csv('retail_sales.csv')
df.head()
df = df.set_index(pd.to_datetime(df['Period'])).drop('Period', axis=1)
df.plot(legend=False)
plt.title('Retail Sales US 1992-2016')
```
## Rescale values
```
df /= 1e3
df.plot(legend=False)
plt.title('Retail Sales US 1992-2016')
```
## Julian date
Using Julian dates turns dates to a real number, making it easier to build a regression.
We also remove the date of the first day, so that dates will start from zero. Notice that Julian dates are measured in days.
```
df['Julian'] = df.index.to_julian_date() - df.index.to_julian_date().min()
df.head()
```
## The most important thing: split Past and Future
```
cutoff = pd.Timestamp('2013-01-01')
train = df.loc[:cutoff].copy()
test = df.loc[cutoff:].copy()
train['Value'].plot()
test['Value'].plot()
```
## Simplest trend: linear
```
from sklearn.linear_model import LinearRegression
lr = LinearRegression()
X_train = train[['Julian']]
X_test = test[['Julian']]
y_train = train['Value']
y_test = test['Value']
lr.fit(X_train, y_train)
lr.score(X_test, y_test)
train['linear_prediction'] = lr.predict(X_train)
test['linear_prediction'] = lr.predict(X_test)
df['Value'].plot()
train['linear_prediction'].plot()
test['linear_prediction'].plot()
```
## Exercise:
Play around with the cutoff date.
- How does the regression score change?
- What happens if you move the cutoff date to before the 2008 crisis?
## Predicting the stationary component
```
df['linear_prediction'] = lr.predict(df[['Julian']])
df['value_minus_linear'] = df['Value'] - df['linear_prediction']
df['value_minus_linear'].plot()
```
## Delayed variables
```
def add_shifts(tdf):
df = tdf.copy()
for i in xrange(1,24):
df['shift_'+str(i)] = df['value_minus_linear'].shift(i).fillna(0)
return df
add_shifts(df).head()
cutoff = pd.Timestamp('2013-01-01')
train = df.loc[:cutoff].copy()
test = df.loc[cutoff:].copy()
train['value_minus_linear'].plot()
test['value_minus_linear'].plot()
train = add_shifts(train)
test = add_shifts(test)
model = LinearRegression()
features = ['Julian'] + list(train.loc[:, 'shift_1':].columns)
print features
X_train = train[features]
X_test = test[features]
y_train = train['value_minus_linear']
y_test = test['value_minus_linear']
model.fit(X_train, y_train)
plt.plot(y_train.values)
plt.plot(model.predict(X_train))
plt.plot(y_test.values)
plt.plot(model.predict(X_test))
```
first year predictions are bad, why?
```
coefs = model.coef_[np.abs(model.coef_) > 0.]
cols = X_train.columns[np.abs(model.coef_) > 0.]
plt.figure(figsize=(10,7))
s = pd.Series(coefs, index=cols).sort_values()
s.plot(kind='bar', fontsize=18)
plt.title('Non-null feature coefficients', fontsize=20)
model.score(X_test, y_test)
```
## Exercise
Scikit Learn offers many other regression models. Try experimenting with any of the following models and see if you can improve your test score.
```
from sklearn.ensemble import RandomForestRegressor
from sklearn.linear_model import ARDRegression, BayesianRidge, ElasticNet, Hinge, Huber
from sklearn.linear_model import Lars, Lasso, LassoLars, ModifiedHuber
from sklearn.linear_model import MultiTaskElasticNet, MultiTaskLasso, OrthogonalMatchingPursuit
from sklearn.linear_model import PassiveAggressiveRegressor, RANSACRegressor, RandomizedLasso
from sklearn.linear_model import Ridge, SGDRegressor, TheilSenRegressor
```
## Question:
You have successfully implemented a model that is able to predict the future. Can you use it to detect anomalies? How? Could you have predicted a rare event like the 2008 crisis?
## Next steps:
- [Autoregressive models](https://en.wikipedia.org/wiki/Autoregressive_integrated_moving_average)
- [Bayesian Time series modeling](http://multithreaded.stitchfix.com/blog/2016/04/21/forget-arima/)
- [Recurrent neural network models](http://www.wildml.com/2015/09/recurrent-neural-networks-tutorial-part-1-introduction-to-rnns/)
| github_jupyter |
# Example Groups
I'm using this notebook to implement example groups using the JSON format defined here.
Many of these groups were obtained from [Group Explorer](https://nathancarter.github.io/group-explorer/index.html): <i>"Visualization software for the abstract algebra classroom".</i>
```
import json
import os
import itertools as it
import algebras as alg
```
## Path Definitions
```
# Path to this repo
aa_path = os.path.join(os.getenv('PYPROJ'), 'abstract_algebra')
# Path to a directory containing Algebra definitions in JSON
alg_dir = os.path.join(aa_path, "Algebras")
```
## Table of Contents
* [S3 -- Symmetric Group on 3 Letters](#S3)
* [Symmetric Group, S3X (not same as S3, above)](#S3X)
* [Z2 (Cyclic Group of Order 2) with Direct Products: Z2_x_Z2 and Z2_x_Z2_x_Z2](#cyclicgroups)
* [Z4 -- Cyclic Group of Order 4](#Z4)
* [Dihedral Group of Order 6](#dihedralgroup6)
* [A4 -- Alternating Group on 4 Letters](#A4)
* [D4 -- Dihedral Group on Four Vertices](#D4)
* [V4 -- Klein-4 Group](#V4)
* [Algebra from Pinter's book, top of page 29](#pinter29)
* [Tesseract Group](#tesseract)
## S3 -- Symmetric Group on 3 Letters<a class="anchor" id="S3"></a>
See this [definition at GitHub](https://github.com/nathancarter/group-explorer/blob/master/groups/S_3.group). "Another name for this group is "Dihedral group on 3 vertices."
```
s3_path = os.path.join(alg_dir, "s3_symmetric_group_on_3_letters.json")
!cat {s3_path}
s3 = alg.Group(s3_path)
s3.pprint()
print(f"\nAbelian? {s3.is_abelian()}")
```
## Symmetric Group, S3X (not same as S3, above)<a class="anchor" id="S3X"></a>
This is the [Symmetric group, S3, as specified at Groupprops](https://groupprops.subwiki.org/wiki/Symmetric_group:S3).
```
s3x = alg.Group('S3X',
'Another version of the symmetric group on 3 letters',
['()', '(1,2)', '(2,3)', '(1,3)', '(1,2,3)', '(1,3,2)'],
[[0, 1, 2, 3, 4, 5],
[1, 0, 4, 5, 2, 3],
[2, 5, 0, 4, 3, 1],
[3, 4, 5, 0, 1, 2],
[4, 3, 1, 2, 5, 0],
[5, 2, 3, 1, 0, 4]]
)
s3x.pprint()
```
## Z2 (Cyclic Group of Order 2) with Direct Products: Z2_x_Z2 and Z2_x_Z2_x_Z2<a class="anchor" id="cyclicgroups"></a>
```
z2 = alg.generate_cyclic_group(2)
z2.pprint()
print(f"\nAbelian? {s3.is_abelian()}")
z2xz2 = z2 * z2
z2xz2.pprint()
print(f"\nAbelian? {s3.is_abelian()}")
z2xz2xz2 = z2 * z2 * z2
z2xz2xz2.pprint()
print(f"\nAbelian? {s3.is_abelian()}")
z2.proper_subgroups()
z2xz2.proper_subgroups()
z2xz2xz2_subgroups = z2xz2xz2.proper_subgroups()
len(z2xz2xz2_subgroups)
[g.order for g in z2xz2xz2_subgroups]
subs = z2xz2xz2_subgroups
for sub in subs:
sub.pprint()
```
## Z4 -- Cyclic Group of Order 4<a class="anchor" id="Z4"></a>
See this [definition at GitHub](https://github.com/nathancarter/group-explorer/blob/master/groups/Z_4.group).
```
z4_json = os.path.join(alg_dir, "z4_cyclic_group_of_order_4.json")
!cat {z4_json}
z4 = alg.Group(z4_json)
z4.pprint()
```
## Dihedral Group of Order 6<a class="anchor" id="dihedralgroup6"></a>
[See Wikipedia](https://en.wikipedia.org/wiki/Dihedral_group_of_order_6)
```
d3_path = os.path.join(alg_dir, "d3_dihedral_group_of_order_6.json")
!cat {d3_path}
d3 = alg.Group(d3_path)
d3.pprint()
subs = d3.proper_subgroups()
for sub in subs:
sub.pprint()
```
## D4 -- Dihedral Group on Four Vertices<a class="anchor" id="D4"></a>
```
d4_path = os.path.join(alg_dir, "d4_dihedral_group_on_4_vertices.json")
!cat {d4_path}
d4 = alg.Group(d4_path)
d4.pprint()
subs = d4.proper_subgroups()
for sub in subs:
sub.pprint()
```
## A4 -- Alternating Group on 4 Letters<a class="anchor" id="A4"></a>
```
a4_path = os.path.join(alg_dir, "a4_alternating_group_on_4_letters.json")
!cat {a4_path}
a4 = alg.Group(a4_path)
a4.pprint()
subs = a4.proper_subgroups()
for sub in subs:
sub.pprint()
```
## Klein-4 Group<a class="anchor" id="V4"></a>
See this [definition at GitHub](https://github.com/nathancarter/group-explorer/blob/master/groups/V_4.group).
```
v4_json = os.path.join(alg_dir, "v4_klein_4_group.json")
!cat {v4_json}
v4 = alg.Group(v4_json)
v4.pprint()
```
## Algebra from Pinter's book, top of page 29<a class="anchor" id="pinter29"></a>
```
pinter_p29_path = os.path.join(alg_dir, "Pinter_page_29.json")
pinter_p29 = alg.Group(pinter_p29_path)
pinter_p29.pprint()
```
## Tesseract group<a class="anchor" id="tesseract"></a>
This group has 384 elements and was converted to JSON from [this definition](https://github.com/nathancarter/group-explorer/blob/master/groups/Tesseract.group).
```
tesseract_path = os.path.join(alg_dir, "tesseract.json")
tesseract = alg.Group(tesseract_path)
print(tesseract)
# %time tesseract.is_abelian() # False
# %time tesseract.is_associative() # True (WARNING: LONG RUNNING TIME)
tesseract.order
```
Here are the first n elements:
```
n = 25
tesseract.elements[:n]
```
## Whatever
```
whatever = alg.Group("Whatever",
"Experimenting",
['e', 'a', 'b', 'c'],
[[0, 1, 2, 3],
[1, 0, 3, 2],
[2, 3, 0, 1],
[3, 2, 1, 0]])
whatever.pprint(True)
foo = [z2xz2, z4, v4]
[whatever.isomorphic(x) for x in foo]
whatever
v4
z2xz2
z4
whatever2 = whatever.deepcopy()
whatever2.about(use_table_names=True)
whatever2.set_elements(['E', 'A', 'B', 'C'])
whatever2.about(use_table_names=True)
```
| github_jupyter |
This notebook is part of the $\omega radlib$ documentation: https://docs.wradlib.org.
Copyright (c) $\omega radlib$ developers.
Distributed under the MIT License. See LICENSE.txt for more info.
# Adjusting radar-base rainfall estimates by rain gauge observations
## Background
There are various ways to correct specific errors and artifacts in radar-based quantitative precipitation estimates (*radar QPE*). Alternatively, you might want to correct your radar QPE regardless of the error source - by using ground truth, or, more specifically, rain gauge observations. Basically, you define the error of your radar QPE at a rain gauge location by the discrepancy between rain gauge observation (considered as "the truth") and radar QPE at that very location. Whether you consider this "discrepancy" as an *additive* or *multiplicative* error is somehow arbitrary - typically, it's a *mix* of both. If you quantify this error at various locations (i.e. rain gauges), you can go ahead and construct correction fields for your radar QPE. You might compute a single correction factor for your entire radar domain (which would e.g. make sense in case of hardware miscalibration), or you might want to compute a spatially variable correction field. This typically implies to interpolate the error in space.
$\omega radlib$ provides different error models and different spatial interpolation methods to address the adjustment problem. For details, please refer to $\omega radlib's$ [library reference](https://docs.wradlib.org/en/latest/adjust.html).
```
import wradlib.adjust as adjust
import wradlib.verify as verify
import wradlib.util as util
import numpy as np
import matplotlib.pyplot as pl
try:
get_ipython().magic("matplotlib inline")
except:
pl.ion()
```
## Example for the 1-dimensional case
Looking at the 1-D (instead of 2-D) case is more illustrative.
### Create synthetic data
First, we **create synthetic data**:
- true rainfall,
- point observations of the truth,
- radar observations of the truth.
The latter is disturbed by some kind of error, e.g. a combination between systemtic and random error.
```
# gage and radar coordinates
obs_coords = np.array([5, 10, 15, 20, 30, 45, 65, 70, 77, 90])
radar_coords = np.arange(0, 101)
# true rainfall
np.random.seed(1319622840)
truth = np.abs(1.5 + np.sin(0.075 * radar_coords)) + np.random.uniform(
-0.1, 0.1, len(radar_coords))
# radar error
erroradd = 0.7 * np.sin(0.2 * radar_coords + 10.)
errormult = 0.75 + 0.015 * radar_coords
noise = np.random.uniform(-0.05, 0.05, len(radar_coords))
# radar observation
radar = errormult * truth + erroradd + noise
# gage observations are assumed to be perfect
obs = truth[obs_coords]
# add a missing value to observations (just for testing)
obs[1] = np.nan
```
### Apply different adjustment methods
- additive error, spatially variable (`AdjustAdd`)
- multiplicative error, spatially variable (`AdjustMultiply`)
- mixed error, spatially variable (`AdjustMixed`)
- multiplicative error, spatially uniform (`AdjustMFB`)
```
# number of neighbours to be used
nnear_raws = 3
# adjust the radar observation by additive model
add_adjuster = adjust.AdjustAdd(obs_coords, radar_coords,
nnear_raws=nnear_raws)
add_adjusted = add_adjuster(obs, radar)
# adjust the radar observation by multiplicative model
mult_adjuster = adjust.AdjustMultiply(obs_coords, radar_coords,
nnear_raws=nnear_raws)
mult_adjusted = mult_adjuster(obs, radar)
# adjust the radar observation by AdjustMixed
mixed_adjuster = adjust.AdjustMixed(obs_coords, radar_coords,
nnear_raws=nnear_raws)
mixed_adjusted = mixed_adjuster(obs, radar)
# adjust the radar observation by MFB
mfb_adjuster = adjust.AdjustMFB(obs_coords, radar_coords,
nnear_raws=nnear_raws,
mfb_args = dict(method="median"))
mfb_adjusted = mfb_adjuster(obs, radar)
```
### Plot adjustment results
```
# Enlarge all label fonts
font = {'size' : 15}
pl.rc('font', **font)
pl.figure(figsize=(10,5))
pl.plot(radar_coords, radar, 'k-', linewidth=2., linestyle="dashed", label="Unadjusted radar", )
pl.plot(radar_coords, truth, 'k-', linewidth=2., label="True rainfall", )
pl.plot(obs_coords, obs, 'o', markersize=10.0, markerfacecolor="grey", label="Gage observation")
pl.plot(radar_coords, add_adjusted, '-', color="red", label="Additive adjustment")
pl.plot(radar_coords, mult_adjusted, '-', color="green", label="Multiplicative adjustment")
pl.plot(radar_coords, mfb_adjusted, '-', color="orange", label="Mean Field Bias adjustment")
pl.plot(radar_coords, mixed_adjusted,'-', color="blue", label="Mixed (mult./add.) adjustment")
pl.xlabel("Distance (km)")
pl.ylabel("Rainfall intensity (mm/h)")
leg = pl.legend(prop={'size': 10})
```
### Verification
We use the `verify` module to compare the errors of different adjustment approaches.
*Here, we compare the adjustment to the "truth". In practice, we would carry out a cross validation.*
```
# Verification for this example
rawerror = verify.ErrorMetrics(truth, radar)
mfberror = verify.ErrorMetrics(truth, mfb_adjusted)
adderror = verify.ErrorMetrics(truth, add_adjusted)
multerror = verify.ErrorMetrics(truth, mult_adjusted)
mixerror = verify.ErrorMetrics(truth, mixed_adjusted)
# Helper function for scatter plot
def scatterplot(x, y, title=""):
"""Quick and dirty helper function to produce scatter plots
"""
pl.scatter(x, y)
pl.plot([0, 1.2 * maxval], [0, 1.2 * maxval], '-', color='grey')
pl.xlabel("True rainfall (mm)")
pl.ylabel("Estimated rainfall (mm)")
pl.xlim(0, maxval + 0.1 * maxval)
pl.ylim(0, maxval + 0.1 * maxval)
pl.title(title)
# Verification reports
maxval = 4.
# Enlarge all label fonts
font = {'size' : 10}
pl.rc('font', **font)
fig = pl.figure(figsize=(14, 8))
ax = fig.add_subplot(231, aspect=1.)
scatterplot(rawerror.obs, rawerror.est, title="Unadjusted radar")
ax.text(0.2, maxval, "Nash=%.1f" % rawerror.nash(), fontsize=12)
ax = fig.add_subplot(232, aspect=1.)
scatterplot(adderror.obs, adderror.est, title="Additive adjustment")
ax.text(0.2, maxval, "Nash=%.1f" % adderror.nash(), fontsize=12)
ax = fig.add_subplot(233, aspect=1.)
scatterplot(multerror.obs, multerror.est, title="Multiplicative adjustment")
ax.text(0.2, maxval, "Nash=%.1f" % multerror.nash(), fontsize=12)
ax = fig.add_subplot(234, aspect=1.)
scatterplot(mixerror.obs, mixerror.est, title="Mixed (mult./add.) adjustment")
ax.text(0.2, maxval, "Nash=%.1f" % mixerror.nash(), fontsize=12)
ax = fig.add_subplot(235, aspect=1.)
scatterplot(mfberror.obs, mfberror.est, title="Mean Field Bias adjustment")
ax.text(0.2, maxval, "Nash=%.1f" % mfberror.nash(), fontsize=12)
pl.tight_layout()
```
## Example for the 2-dimensional case
For the 2-D case, we follow the same approach as before:
- create synthetic data: truth, rain gauge observations, radar-based rainfall estimates
- apply adjustment methods
- verification
The way these synthetic data are created is totally arbitrary - it's just to show how the methods are applied.
### Create 2-D synthetic data
```
# grid axes
xgrid = np.arange(0, 10)
ygrid = np.arange(20, 30)
# number of observations
num_obs = 10
# create grid
gridshape = len(xgrid), len(ygrid)
grid_coords = util.gridaspoints(ygrid, xgrid)
# Synthetic true rainfall
truth = np.abs(10. * np.sin(0.1 * grid_coords).sum(axis=1))
# Creating radar data by perturbing truth with multiplicative and
# additive error
# YOU CAN EXPERIMENT WITH THE ERROR STRUCTURE
np.random.seed(1319622840)
radar = 0.6 * truth + 1. * np.random.uniform(low=-1., high=1,
size=len(truth))
radar[radar < 0.] = 0.
# indices for creating obs from raw (random placement of gauges)
obs_ix = np.random.uniform(low=0, high=len(grid_coords),
size=num_obs).astype('i4')
# creating obs_coordinates
obs_coords = grid_coords[obs_ix]
# creating gauge observations from truth
obs = truth[obs_ix]
```
### Apply different adjustment methods
```
# Mean Field Bias Adjustment
mfbadjuster = adjust.AdjustMFB(obs_coords, grid_coords)
mfbadjusted = mfbadjuster(obs, radar)
# Additive Error Model
addadjuster = adjust.AdjustAdd(obs_coords, grid_coords)
addadjusted = addadjuster(obs, radar)
# Multiplicative Error Model
multadjuster = adjust.AdjustMultiply(obs_coords, grid_coords)
multadjusted = multadjuster(obs, radar)
```
### Plot 2-D adjustment results
```
# Helper functions for grid plots
def gridplot(data, title):
"""Quick and dirty helper function to produce a grid plot
"""
xplot = np.append(xgrid, xgrid[-1] + 1.) - 0.5
yplot = np.append(ygrid, ygrid[-1] + 1.) - 0.5
grd = ax.pcolormesh(xplot, yplot, data.reshape(gridshape), vmin=0,
vmax=maxval)
ax.scatter(obs_coords[:, 0], obs_coords[:, 1], c=obs.ravel(),
marker='s', s=50, vmin=0, vmax=maxval)
#pl.colorbar(grd, shrink=0.5)
pl.title(title)
# Maximum value (used for normalisation of colorscales)
maxval = np.max(np.concatenate((truth, radar, obs, addadjusted)).ravel())
# open figure
fig = pl.figure(figsize=(10, 6))
# True rainfall
ax = fig.add_subplot(231, aspect='equal')
gridplot(truth, 'True rainfall')
# Unadjusted radar rainfall
ax = fig.add_subplot(232, aspect='equal')
gridplot(radar, 'Radar rainfall')
# Adjusted radar rainfall (MFB)
ax = fig.add_subplot(234, aspect='equal')
gridplot(mfbadjusted, 'Adjusted (MFB)')
# Adjusted radar rainfall (additive)
ax = fig.add_subplot(235, aspect='equal')
gridplot(addadjusted, 'Adjusted (Add.)')
# Adjusted radar rainfall (multiplicative)
ax = fig.add_subplot(236, aspect='equal')
gridplot(multadjusted, 'Adjusted (Mult.)')
pl.tight_layout()
# Open figure
fig = pl.figure(figsize=(6, 6))
# Scatter plot radar vs. observations
ax = fig.add_subplot(221, aspect='equal')
scatterplot(truth, radar, 'Radar vs. Truth (red: Gauges)')
pl.plot(obs, radar[obs_ix], linestyle="None", marker="o", color="red")
# Adjusted (MFB) vs. radar (for control purposes)
ax = fig.add_subplot(222, aspect='equal')
scatterplot(truth, mfbadjusted, 'Adjusted (MFB) vs. Truth')
# Adjusted (Add) vs. radar (for control purposes)
ax = fig.add_subplot(223, aspect='equal')
scatterplot(truth, addadjusted, 'Adjusted (Add.) vs. Truth')
# Adjusted (Mult.) vs. radar (for control purposes)
ax = fig.add_subplot(224, aspect='equal')
scatterplot(truth, multadjusted, 'Adjusted (Mult.) vs. Truth')
pl.tight_layout()
```
| github_jupyter |
<p><b>Report #6, 27-05-2020</p></b>
```
import pandas as pd
pd.set_option('display.max_columns', None)
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn import preprocessing
import numpy as np
from sklearn.metrics import confusion_matrix
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
import itertools
import sklearn.metrics as metrics
from sklearn.model_selection import cross_val_score, cross_val_predict
from sklearn import metrics
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
from sklearn.model_selection import cross_val_predict
from sklearn.metrics import confusion_matrix
import matplotlib.pyplot as plt
# %matplotlib notebook
# %%html
# pd.set_option('display.max_colwidth', -1)
# pd.set_option('display.max_columns', None)
```
<p><b>1- Lets have look at the dataset:</p></b>
<P> - text fields before prepocessing: {essay0, essay1, essay2, essay7, essay8, essay9}</p>
<P> - text field after preprocessing: {clean_text}</p>
<p> In the preprocessing phase all the essays are merged for each user and useless information is omitted from the text.</p>
<P> - target field: {isced}</p>
<P> - shape of dataset: (49924, 20)</p>
<p> It means that the dataset consists of 49924 rows(user) and 20 columns(features for each user) after removing all the null values.</p>
```
okcupid_df = pd.read_csv('../../data/processed/preprocessed_cupid.csv')
okcupid_df.dropna(subset=['isced', 'text', 'clean_text'], inplace=True)
print('A small sample of OkCupid dataset:')
okcupid_df.head(3)
```
<p><b>2- Using LIWC(Linguistic Inquiry and Word Count)</p></b>
<p> - LIWC is a transparent text analysis program that counts words in psychologically meaningful categories.</p>
<p> - Table 1 provides a list of the default LIWC2015 dictionary categories, scales, sample scale words, and relevant scale word counts.</p>
<img src="images/img1.jpg">
<img src="images/img2.jpg">
<img src="images/img3.jpg">
<a href="https://repositories.lib.utexas.edu/bitstream/handle/2152/31333/LIWC2015_LanguageManual.pdf">Link to the manual</a>
<p>- The LIWC package relies on internal default dictionaries that define which words should be counted in which target category. we are using LIWC2015 v1.6.</p>
<p>- LIWC receives our dataset in the csv format and goes through each rows of dataset word by word. Each word would be compared with the dictionary file(LIWC2015) and the associated categories would be distinguished. After going through all the words in the text , LIWC would calculate the percentage of each LIWC category. So, for example, we might discover that 2.34% of all the words in a given text are impersonal pronouns and 3.33% are auxiliary verbs.
</p>
```
liwc_df = pd.read_csv('../../data/liwc-data/liwic_cupid_v0.csv', dtype=object, low_memory = False)
# Some preprations on data
# Rename first three columns
liwc_df.rename(columns={'A':'pre_index', 'B':'text', 'C':'isced'}, inplace=True)
# skip first row (header) and first two columns
liwc_df = liwc_df.iloc[1:, 2:]
# print(liwc_df.columns)
# change type to float
liwc_df.replace(',','.',inplace=True, regex=True)
liwc_df = liwc_df.astype(float)
liwc_df['isced'].mask(liwc_df['isced'].isin([3.0, 5.0, 1.0]) , 0.0, inplace=True)
liwc_df['isced'].mask(liwc_df['isced'].isin([6.0, 7.0, 8.0]) , 1.0, inplace=True)
data = liwc_df.iloc[:, 1:]
target = liwc_df.iloc[:, 0].astype(float)
print('A small sample of dataset with LIWC outputs:')
liwc_df.head(10)
```
<p> - LIWC is applied to the 'text' column in the original dataset before preprocessing.</p>
<p> - shape of LIWC out put is (49943, 94)</p>
<p> It means that there are 49943 rows in the dataset and for each row we have extracted 94 features from the text</p>
<p> - List of all the columns in LIWC output:</p>
<p> ['isced', 'WC', 'Analytic', 'Clout', 'Authentic', 'Tone', 'WPS',
'Sixltr', 'Dic', 'function', 'pronoun', 'ppron', 'i', 'we', 'you',
'shehe', 'they', 'ipron', 'article', 'prep', 'auxverb', 'adverb',
'conj', 'negate', 'verb', 'adj', 'compare', 'interrog', 'number',
'quant', 'affect', 'posemo', 'negemo', 'anx', 'anger', 'sad', 'social',
'family', 'friend', 'female', 'male', 'cogproc', 'insight', 'cause',
'discrep', 'tentat', 'certain', 'differ', 'percept', 'see', 'hear',
'feel', 'bio', 'body', 'health', 'sexual', 'ingest', 'drives',
'affiliation', 'achieve', 'power', 'reward', 'risk', 'focuspast',
'focuspresent', 'focusfuture', 'relativ', 'motion', 'space', 'time',
'work', 'leisure', 'home', 'money', 'relig', 'death', 'informal',
'swear', 'netspeak', 'assent', 'nonflu', 'filler', 'AllPunc', 'Period',
'Comma', 'Colon', 'SemiC', 'QMark', 'Exclam', 'Dash', 'Quote',
'Apostro', 'Parenth', 'OtherP']</p>
<p> - isced : </p>
<p> 0.0 -> primary educated</p>
<p> 1.0 -> high educated</p>
<p> - Most LIWC2015 output variables are expressed as percentage of total words.
There are six exceptions: word count (WC; raw word count), (WPS; mean words per sentence), and four summary variables: Analytic, Clout, Authentic, and Tone. </p>
<p> - LIWC2015 converts all the text to lower case before processing.</p>
<p> - LIWC2015 can only count words that are in its dictionaries. Misspellings, colloquialisms, foreign words, and abbreviations are usually not in the internal dictionaries.</p>
<p> - The Words per sentence (WPS) category is based on the number of times that end-of-sentence markers are detected. These include all periods (.), question marks, and exclamation points. </p>
<p> - Many types of “netspeak” that is used as shorthand interpersonal communication (e.g., “lol”, “4ever”) are captured by the LIWC2015 dictionary.</p>
### The sumary statistics of quantitative variables
```
data.describe()
```
<p><b> 3- Training Logistic regression based on LIWC output without including text.<p><b>
### Grid search settings for Logistic regression
<img src="images/img5.jpg">
```
def plot_confusion_matrix(cm, classes, normalize=False, title='Confusion matrix', cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0])
, range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# Train dataset
X_train,X_test, y_train, y_test = train_test_split(data, target, test_size = 0.25, random_state=0)
X_train, X_val, y_train, y_val = train_test_split(X_train, y_train, test_size = 0.25)
scaler = preprocessing.StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
# X_train_scaled.mean(axis=0)
# X_train_scaled.std(axis=0)
LogisticRegr = LogisticRegression(random_state=0, max_iter=10000, solver='lbfgs', penalty='l2', class_weight='balanced')
LogisticRegr.fit(X_train_scaled, y_train)
X_val_scaler = scaler.transform(X_val)
predictions = LogisticRegr.predict(X_val_scaler)
print("Final Accuracy for Logistic: %s"% accuracy_score(y_val, predictions))
cm = confusion_matrix(y_val,predictions)
plt.figure()
plot_confusion_matrix(cm, classes=[0,1], normalize=False,
title='Confusion Matrix')
print(classification_report(y_val, predictions))
# calculate the fpr and tpr for all thresholds of the classification
probs = LogisticRegr.predict_proba(X_val_scaler)
preds = probs[:,1]
fpr, tpr, threshold = metrics.roc_curve(y_val, preds)
roc_auc = metrics.auc(fpr, tpr)
import matplotlib.pyplot as plt
plt.title('Receiver Operating Characteristic(ROC curve)')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
```
<b>Logistic regression with Cross Validation:</b>
```
# Logistic regression with Cross Validation
print('# Logistic regression with 5-Fold Cross Validation:')
LogisticRegr_cross = LogisticRegression(random_state=0, max_iter=10000, solver='lbfgs', penalty='l2', class_weight='balanced')
LogisticRegr_cross.fit(X_train_scaled, y_train)
data_scaler = scaler.transform(data)
scores = cross_val_score(LogisticRegr_cross, data_scaler, target, cv=5)
print(scores)
print(scores.mean())
y_pred = cross_val_predict(LogisticRegr_cross, X_val_scaler, y_val, cv=5)
conf_mat = confusion_matrix(y_val, y_pred)
print(conf_mat)
```
<p><b> 5- Logistic regression feature coefficient values</b></p>
```
# Most Important features
import matplotlib.pyplot as plt
importance = LogisticRegr.coef_[0]
# summarize feature importance
for i,v in enumerate(importance):
print('Feature: %0d, Score: %.5f' % (i,v))
# plot feature importance
plt.bar([x for x in range(len(importance))], importance)
plt.show()
col_no_pos =[]
col_no_neg =[]
for i,v in enumerate(importance):
if v>0.3:
col_no_pos. append(i)
print('Feature: %0d, Score: %.5f' % (i,v))
elif v< -0.3:
col_no_neg. append(i)
# print('Feature: %0d, Score: %.5f' % (i,v))
print("positive:",col_no_pos)
print("negetive",col_no_neg)
```
<b> features with |coefficient| >0.2: </b>
```
cols_name = data.columns
# liwc_df['isced'] = liwc_df['isced'].astype(float)
# liwc_df['isced'].mask(liwc_df['isced'].isin([3, 5.0, 1.0]) , 0 , inplace=True) # 0 for primary education
# liwc_df['isced'].mask(liwc_df['isced'].isin([6.0, 7.0, 8.0]) , 1 , inplace=True) # 1 for high education
cols_no_pos = [1, 2, 6, 8, 9, 30, 55, 62, 65, 84, 90]
cols_no_neg = [10, 11, 16, 18, 29, 67, 75]
imp_cols_pos = []
imp_cols_neg =[]
for i in cols_no_pos:
imp_cols_pos.append( cols_name[i])
for i in cols_no_neg:
imp_cols_neg.append(cols_name[i])
print('list of features that are more involved in predicting class 0:', imp_cols_neg)
print('list of features that are more involved in predicting class 1:', imp_cols_pos)
```
| Abbrev | Category |Examples|
|------|------|-----|
|ppron|Personal pronouns|I, them, her|
|i|1st pers singular|I, me, mine|
|ipron|Impersonal pronouns|it, it’s, those|
| prep | Prepositions|to, with, above|
|affect|Affective processes|happy, cried|
|space|space|down, in, thin|
|informal|Informal language|
| Abbrev | Category |Examples|
|------|------|-----|
|Analytic|Analytical thinking|
|Clout|Clout|
|Sixltr|Words > 6 letters|
|function|Total function words|it, to, no, very|
|posemo|Positive emotion|love, nice, sweet|
|pronoun|Total pronouns|I, them, itself|
|ingest|Ingestion|dish, eat, pizza|
|focuspast|Past focus|ago, did, talked|
|relativ|Relativity|area, bend, exit|
|Apostro||
|Colon||
<P><b>6- training logistic regression considering both clean_text and LIWC features:</p></b>
```
from sklearn.preprocessing import FunctionTransformer
from sklearn.pipeline import FeatureUnion
from sklearn.pipeline import Pipeline
from sklearn.feature_extraction.text import CountVectorizer
df_merge = pd.read_csv('D:\projects\okcupid\data\liwc-data/liwic_cupid.csv', dtype=object, low_memory = False)
df_merge.rename(columns={'A':'pre_index', 'B':'text', 'C':'isced', 'D':'clean_text'}, inplace=True)
df_merge = df_merge.iloc[1:, :]
df_merge = df_merge.iloc[ : , 2: ]
df_merge.replace(',','.',inplace=True, regex=True)
df_merge.iloc[:, 2: ] = df_merge.iloc[ : , 2: ].astype(float)
df_merge['clean_text'] = df_merge['clean_text'].astype(str)
df_merge['isced'] = df_merge['isced'].astype(float)
df_merge['isced'].mask(df_merge['isced'].isin([3, 5.0, 1.0]) , 0 , inplace=True) # 0 for primary education
df_merge['isced'].mask(df_merge['isced'].isin([6.0, 7.0, 8.0]) , 1 , inplace=True) # 1 for high education
# df['isced'] = df['isced'].astype(int)
df_merge.head(2)
data = df_merge.iloc[:, 1:]
target = df_merge.iloc[:, 0].astype(float)
# vectorization
X_t, X_test, y_t, y_test = train_test_split(data, target, train_size=0.75, stratify=target, test_size=0.25, random_state = 0)
X_train, X_val, y_train, y_val = train_test_split(X_t, y_t, train_size=0.75, stratify=y_t, test_size=0.25, random_state = 0)
cols = X_train.columns
cols = cols[1:]
cols
get_text_data = FunctionTransformer(lambda x: x['clean_text'], validate=False)
get_numeric_data = FunctionTransformer(lambda x: x[cols], validate=False)
# merge vectorized text data and scaled numeric data
process_and_join_features = Pipeline([
('features', FeatureUnion([
('numeric_features', Pipeline([
('selector', get_numeric_data),
('scaler', preprocessing.StandardScaler())
])),
('text_features', Pipeline([
('selector', get_text_data),
('vec', CountVectorizer(binary=False, ngram_range=(1, 2), lowercase=True))
]))
])),
('clf', LogisticRegression(random_state=0,max_iter=5000, solver='sag', penalty='l2', class_weight='balanced'))
])
#
process_and_join_features.fit(X_train, y_train)
predictions = process_and_join_features.predict(X_val)
print("Final Accuracy for Logistic: %s"% accuracy_score(y_val, predictions))
cm = confusion_matrix(y_val,predictions)
# plt.figure()
# plot_confusion_matrix(cm, classes=[0,1], normalize=False,
# title='Confusion Matrix')
print(classification_report(y_val, predictions))
plt.figure()
plot_confusion_matrix(cm, classes=[0,1], normalize=False, title='Confusion Matrix')
# calculate the fpr and tpr for all thresholds of the classification
probs = process_and_join_features.predict_proba(X_val)
preds = probs[:,1]
fpr, tpr, threshold = metrics.roc_curve(y_val, preds)
roc_auc = metrics.auc(fpr, tpr)
import matplotlib.pyplot as plt
plt.title('Receiver Operating Characteristic(ROC curve)')
plt.plot(fpr, tpr, 'b', label = 'AUC = %0.2f' % roc_auc)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
plt.show()
```
| github_jupyter |
# Advanced Object Oriented Programming
This notebook is created by Eda AYDIN through by Udemy, DATAI Team.
## Classes
```
# While writing the class name, you should use capital letter for the first letter.
class Employee(object): # object: Object is created to use inside Employee (We don't write this, because Python creates it automatically.)
pass # If we do not write anything in the class, but we do not want to see the Syntax Error while running the whole, we use pass.
# attribute : age, address, name
# behavior : pass, run
employee1 = Employee()
print(employee1) # We created object for Employee class
```
## Attributes
```
class FootballPlayers:
football_club = "Barcelona"
age = 30
f1 = FootballPlayers()
print(f1) # print the object we created
print(f1.age)
print(f1.football_club)
f1.football_club = "Real Madrid"
print(f1.football_club)
```
It may not always be correct to change it in any way, normally in code written by someone else. In this section, I created a class in this way to explain what the Attribute expression is and how it works.
## Methods
```
class Square:
def __init__(self, edge):
self.edge = edge
def calculateCircumference(self):
return self.edge * 4
def calculateArea(self):
return self.edge ** 2
square = Square(int(input("Enter the number:")))
print("Circumference of square : {} and Area of square: {}".format(square.calculateCircumference(),square.calculateArea()))
```
## Methods and Functions
```
class Employee2:
def __init__(self,age,salary):
self.age = age
self.salary = salary
def ageSalaryRatio(self): # This is a method. This takes objects not variable.
return self.age / self.salary
def ageSalaryRatio2(age,salary): # This is a function. This function does not depend on any class. So, this takes just variable, not object.
return age / salary
age = int(input())
salary = int(input())
employee = Employee2(age,salary)
employee.ageSalaryRatio()
ageSalaryRatio2(age,salary)
import math
class Area:
def __init__(self,radius,a,b):
self.radius = radius
self.a = a
self.b = b
self.pi = math.pi
def circleArea(self):
print("Radius is : {}".format(self.radius))
return self.pi * self.radius **2
def squareArea(self):
print("Edge is : {}".format(self.a))
return self.a ** 2
def rectangularArea(self):
print("Edges are : {} and {}".format(self.a, self.b))
return self.a * self.b
square = Area(0,int(input()),0)
square.squareArea()
rectangular = Area(0, int(input()), int(input()))
rectangular.rectangularArea()
radius = Area(int(input()),0,0)
radius.circleArea()
```
## Constructor / Initializer
```
class Animal:
def __init__(self,breed,age):
self.breed = breed
self.age = age
def getAge(self):
return self.age
def getBreed(self):
return self.breed
animal1 = Animal(input(),int(input())) #If you use just input(), the Python understand that you write a string. Remember in each coding part.
animal1.getAge()
animal1.getBreed()
```
## Calculator Project
```
class Calculator:
"""Calculator"""
def __init__(self,x, y,operation):
"""Initialize values"""
self.x = x
self.y = y
self.operation = operation
def printResult(self,result):
print("You entered: {} and {} numbers, and {} mathematical operation. Your result is {} {} {} = {}".format(
self.x,self.y,self.operation,self.x,self.operation,self.y,result))
def determineOperation(self):
"""Get the user 2 integer input with 1 string operation sign and calculate the result according to operation sign.
If user enter the wrong operation sign, the code will ask it again."""
if self.operation == "+": self.printResult(self.x + self.y)
elif self.operation == "-": self.printResult(self.x - self.y)
elif self.operation == "*": self.printResult(self.x * self.y)
elif self.operation == "/": self.printResult(self.x / self.y)
elif self.operation == "%": self.printResult(self.x % self.y)
elif self.operation == "**": self.printResult(self.x ** self.y)
elif self.operation == "//": self.printResult(self.x // self.y)
else:
print("You entered {} operation.This operation is not define in the mathematics. Try again.".format(self.operation))
time.sleep(1)
input2 = Calculator(int(input("Enter the first number: ")),
int(input("Enter the second number: ")),
input("Enter the operation sign: "))
return input2.determineOperation()
input1 = Calculator(int(input("Enter the first number: ")),
int(input("Enter the second number: ")),
input("Enter the operation sign: "))
input1.determineOperation()
```
## Encapsulation
Restricting external access to variables defined in any class is called **encapsulation**.
```
import time
import pandas as pd
class BankAccount:
def __init__(self,name, money, address):
self.name = name # global variable
self.__money = money # private variable
self.address = address # global variable
def getMoney(self):
return self.__money
def setMoney(self,money):
self.__money = money
# private
def __increase(self):
self.__money = self.__money + 500
if __name__ == '__main__':
n = int(input("Enter the number as the number of people your want to add: "))
for i in range(1,n+1):
globals()['person{}'.format(i)] = BankAccount(input("Enter the name:"),
int(input("Enter the money:")),
input("Enter the address:"))
person1.getMoney()
person2.getMoney()
print("get method:",person1.getMoney())
person1.setMoney(500)
print("after set method:",person1.getMoney())
person1.__increase()
print("after raise:",person1.getMoney())
```
Since the __increase() method is a private method, it cannot be accessed by any means from the outside.
## Inheritance
Creating a new class using the variables or methods of a previously created class is called **inheritance**.
```
# Parent class
class Animal:
def __init__(self):
print("Animal is created.")
def toString(self):
print("This is animal.")
def walk(self):
print("Animal can walk.")
# Child class
class Monkey(Animal):
def __init__(self):
super().__init__() #get the variables from Animal. I can use init of (animal) class
print("Monkey is created")
def toString(self):
print("This is monkey.")
def climb(self):
print("Monkey can climb.")
m1 = Monkey()
m1.toString()
m1.walk()
m1.climb()
class Bird(Animal):
def __init__(self):
super().__init__()
print("Bird is created")
def toString(self):
print("This is bird.")
def fly(self):
print("Monkey can fly.")
b1 = Bird()
b1.toString()
b1.walk()
b1.fly()
b1.climb() # This climb method is ony a method that belongs to the Monkey class.
```
### Inheritance Project
```
class Website:
def __init__(self,name, surname):
self.name = name
self.surname = surname
def loginInfo(self):
return self.name, self.surname
class Website1(Website):
def __init__(self, name, surname, ids, email):
Website.__init__(self,name, surname)
self.ids = ids
self.email = email
def LoginId(self):
print("Name: {} Surname: {}, Id: {}".format(self.name, self.surname, self.ids))
def loginEmail(self):
print("Name: {} Surname: {}, Email: {}".format(self.name, self.surname, self.email))
if __name__ == '__main__':
n = int(input("Enter the number as the number of people your want to add: "))
for i in range(1,n+1):
globals()['person{}'.format(i)] = Website(input("Enter the person{} name:".format(i)),
input("Enter the person{} surname:".format(i)))
for i in range(1,n+1):
globals()['person{}'.format(i)] = Website1(globals()['person{}'.format(i)].name,
globals()["person{}".format(i)].surname,
input("Enter the id of person{}:".format(i)),
input("Enter the email of person{}:".format(i)))
person1.LoginId()
person1.loginEmail()
person2.LoginId()
person2.loginEmail()
```
## Abstract Classes
- The abstract classes can't create an instance.
- All methods defined as Abstract must be used in other classes defined as inheritance over this class. In other words, Abstract methods must be used by subclasses.
```
from abc import ABC, abstractmethod
class Animal2(ABC): # super class
@abstractmethod
def walk(self): pass
@abstractmethod
def run(self): pass
a = Animal2()
```
I was expecting to get this error because I created an abstract class.
```
class Bird2(Animal2):
def __init__(self):
print("Bird")
b2 = Bird2()
class Bird3(Animal2):
def __init__(self):
print("Bird")
def walk(self):
print("walk")
def run(self):
print("run")
b3 = Bird3() # I didn't get any error while creating an instance where I used both walk and run methods defined as abstract in the Bird3 class inherited by Animal2.
```
## Overriding
- If I have two different methods with the same name in both the parent class and the subclass, then the subclass executes its own function using the overriding property.
```
class Animal3: #parent
def toString(self):
print("Animal")
class Monkey3(Animal3): #subclass
def toString(self):
print("Monkey")
a3 = Animal3()
a3.toString()
m1 = Monkey3()
m1.toString() # Monkey3 calls overriding method
class Rectangle():
def __init__(self,length,breadth):
self.length = length
self.breadth = breadth
def getArea(self):
print(self.length*self.breadth," is area of rectangle")
class Square(Rectangle):
def __init__(self,side):
self.side = side
Rectangle.__init__(self,side,side)
def getArea(self):
print(self.side*self.side," is area of square")
s = Square(4)
r = Rectangle(2,4)
s.getArea()
r.getArea()
```
## Polymorphism
- Using methods transferred from superclass via subclass inherit in a different way is called Polymorphism.
- Overriding is the sample of this situation.
```
class FinancialTransaction:
def __init__(self,name, surname,id, money,profession):
self.name = name
self.surname = surname
self.money = money
self.id = id
self.profession = profession
def getMoney(self):
return self.money
def getID(self):
return self.id
class NonEngineer(FinancialTransaction):
def __init__(self,name, surname,id, money,profession):
FinancialTransaction.__init__(self,name, surname,id, money,profession)
self.raisemoney = 0
def getMoney(self):
return self.money
def raiseMoney(self):
raise_rate = 0.6
return self.money + self.money * raise_rate
class ComputerEngineer(FinancialTransaction):
def __init__(self,name, surname,id, money,profession):
FinancialTransaction.__init__(self,name, surname,id, money,profession)
self.raisemoney = 0
def getMoney(self):
return self.money
def raiseMoney(self):
raise_rate = 0.3
return self.money + self.money * raise_rate
class DataScientist(FinancialTransaction):
def __init__(self,name, surname,id, money,profession):
FinancialTransaction.__init__(self,name, surname,id, money,profession)
def getMoney(self):
return self.money
def raiseMoney(self):
raise_rate = 0.6
return self.money + self.money * raise_rate
if __name__ == '__main__':
n = int(input("Enter the number as the number of people your want to add: "))
for i in range(1,n+1):
globals()['person{}'.format(i)] = FinancialTransaction(input("Enter the person{} name:".format(i)),
input("Enter the person{} surname:".format(i)),
int(input("Enter the person{} id:".format(i))),
int(input("Enter the person{} money:".format(i))),
int(input("Enter the person{} profession: (Non-engineer: 0 - Computer Engineer: 1, Data Scientist: 2)".format(i))))
for j in range(1, n+1):
if str(globals()['person{}'.format(i)].profession) == "0":
globals()['person_neng{}'.format(j)] = NonEngineer(globals()['person{}'.format(i)].name,
globals()['person{}'.format(i)].surname,
globals()['person{}'.format(i)].id,
globals()['person{}'.format(i)].money,
globals()['person{}'.format(i)].profession)
elif str(globals()['person{}'.format(i)].profession) == "1":
globals()['personcs{}'.format(j)] = ComputerEngineer(globals()['person{}'.format(i)].name,
globals()['person{}'.format(i)].surname,
globals()['person{}'.format(i)].id,
globals()['person{}'.format(i)].money,
globals()['person{}'.format(i)].profession)
elif str(globals()['person{}'.format(i)].profession) == "2":
globals()['personds{}'.format(j)] = DataScientist(globals()['person{}'.format(i)].name,
globals()['person{}'.format(i)].surname,
globals()['person{}'.format(i)].id,
globals()['person{}'.format(i)].money,
globals()['person{}'.format(i)].profession)
for i in range(1,n+1):
if str(globals()['person{}'.format(i)].profession) == "0":
print("Previous salary of non-engineer with {} id number: {} Salary after raise: {}".format(globals()["person_neng{}".format(i)].id,
globals()["person_neng{}".format(i)].money,
globals()["person_neng{}".format(i)].raiseMoney()))
for i in range(1,n+1):
if str(globals()['person{}'.format(i)].profession) == "1":
print("Previous salary of computer engineer with {} id number: {} Salary after raise: {}".format(globals()["personcs{}".format(i)].id,
globals()["personcs{}".format(i)].money,
globals()["personcs{}".format(i)].raiseMoney()))
for i in range(1,n+1):
if str(globals()['person{}'.format(i)].profession) == "2":
print("Previous salary of data scientist with {} id number: {} Salary after raise: {}".format(globals()["personds{}".format(i)].id,
globals()["personds{}".format(i)].money,
globals()["personds{}".format(i)].raiseMoney()))
```
| github_jupyter |
```
import os
import folium
print(folium.__version__)
```
**Note** : The examples presented below are the copy of the ones presented on https://github.com/bbecquet/Leaflet.PolylineOffset
## Basic Demo
- The dashed line is the "model", with no offset applied.
- The Red line is with a -5px offset,
- The Green line is with a 10px offset.
The three are distinct Polyline objects but uses the same coordinate array
```
from folium import plugins
m = folium.Map(location=[58.0, -11.0], zoom_start=4, tiles="cartodbpositron")
coords = [
[58.44773, -28.65234],
[53, -23.33496],
[53, -14.32617],
[58.1707, -10.37109],
[59, -13],
[57, -15],
[57, -18],
[60, -18],
[63, -5],
[59, -7],
[58, -3],
[56, -3],
[60, -4],
]
plugins.PolyLineOffset(
coords, weight=2, dash_array="5,10", color="black", opacity=1
).add_to(m)
plugins.PolyLineOffset(coords, color="#f00", opacity=1, offset=-5).add_to(m)
plugins.PolyLineOffset(coords, color="#080", opacity=1, offset=10).add_to(m)
m.save(os.path.join('results', "PolyLineOffset_simple.html"))
m
```
## Bus Lines
A more complex demo.
Offsets are computed automatically depending on the number of bus lines using the same segment.
Other non-offset polylines are used to achieve the white and black outline effect.
```
m = folium.Map(location=[48.868, 2.365], zoom_start=15)
geojson = {
"type": "FeatureCollection",
"features": [
{
"type": "Feature",
"properties": {"lines": [0, 1]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.357919216156006, 48.87621773324153],
[2.357339859008789, 48.874834693731664],
[2.362983226776123, 48.86855408432749],
[2.362382411956787, 48.86796126699168],
[2.3633265495300293, 48.86735432768131],
],
},
},
{
"type": "Feature",
"properties": {"lines": [2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.351503372192383, 48.86443950493823],
[2.361609935760498, 48.866775611250205],
[2.3633265495300293, 48.86735432768131],
],
},
},
{
"type": "Feature",
"properties": {"lines": [1, 2]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.369627058506012, 48.86619159489603],
[2.3724031448364253, 48.8626397112042],
[2.3728322982788086, 48.8616233285001],
[2.372767925262451, 48.86080456075567],
],
},
},
{
"type": "Feature",
"properties": {"lines": [0]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3647427558898926, 48.86653565369396],
[2.3647642135620117, 48.86630981023694],
[2.3666739463806152, 48.86314789481612],
[2.3673176765441895, 48.86066339254944],
],
},
},
{
"type": "Feature",
"properties": {"lines": [0, 1, 2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3633265495300293, 48.86735432768131],
[2.3647427558898926, 48.86653565369396],
],
},
},
{
"type": "Feature",
"properties": {"lines": [1, 2, 3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.3647427558898926, 48.86653565369396],
[2.3650002479553223, 48.86660622956524],
[2.365509867668152, 48.866987337550164],
[2.369627058506012, 48.86619159489603],
],
},
},
{
"type": "Feature",
"properties": {"lines": [3]},
"geometry": {
"type": "LineString",
"coordinates": [
[2.369627058506012, 48.86619159489603],
[2.372349500656128, 48.865702850895744],
],
},
},
],
}
# manage overlays in groups to ease superposition order
outlines = folium.FeatureGroup("outlines")
line_bg = folium.FeatureGroup("lineBg")
bus_lines = folium.FeatureGroup("busLines")
bus_stops = folium.FeatureGroup("busStops")
line_weight = 6
line_colors = ["red", "#08f", "#0c0", "#f80"]
stops = []
for line_segment in geojson["features"]:
# Get every bus line coordinates
segment_coords = [[x[1], x[0]] for x in line_segment["geometry"]["coordinates"]]
# Get bus stops coordinates
stops.append(segment_coords[0])
stops.append(segment_coords[-1])
# Get number of bus lines sharing the same coordinates
lines_on_segment = line_segment["properties"]["lines"]
# Width of segment proportional to the number of bus lines
segment_width = len(lines_on_segment) * (line_weight + 1)
# For the white and black outline effect
folium.PolyLine(
segment_coords, color="#000", weight=segment_width + 5, opacity=1
).add_to(outlines)
folium.PolyLine(
segment_coords, color="#fff", weight=segment_width + 3, opacity=1
).add_to(line_bg)
# Draw parallel bus lines with different color and offset
for j, line_number in enumerate(lines_on_segment):
plugins.PolyLineOffset(
segment_coords,
color=line_colors[line_number],
weight=line_weight,
opacity=1,
offset=j * (line_weight + 1) - (segment_width / 2) + ((line_weight + 1) / 2),
).add_to(bus_lines)
# Draw bus stops
for stop in stops:
folium.CircleMarker(
stop,
color="#000",
fill_color="#ccc",
fill_opacity=1,
radius=10,
weight=4,
opacity=1,
).add_to(bus_stops)
outlines.add_to(m)
line_bg.add_to(m)
bus_lines.add_to(m)
bus_stops.add_to(m)
m.save(os.path.join('results', "PolyLineOffset_bus.html"))
m
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import json
import torch
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
from gluonts.dataset.repository.datasets import get_dataset
from gluonts.dataset.util import to_pandas
from gluonts.evaluation import Evaluator
from gluonts.evaluation.backtest import make_evaluation_predictions
from pts.model.deepar import DeepAREstimator
from pts.dataset.repository.datasets import dataset_recipes
from pts.modules import ZeroInflatedNegativeBinomialOutput
from pts import Trainer
dataset = get_dataset("pts_m5", regenerate=False)
entry = next(iter(dataset.train))
train_series = to_pandas(entry)
train_series.plot()
plt.grid(which="both")
plt.legend(["train series"], loc="upper left")
plt.title(entry['item_id'])
plt.show()
entry = next(iter(dataset.test))
test_series = to_pandas(entry)
test_series.plot()
plt.axvline(train_series.index[-1], color='r') # end of train dataset
plt.grid(which="both")
plt.legend(["test series", "end of train series"], loc="upper left")
plt.title(entry['item_id'])
plt.show()
print(f"Recommended prediction horizon: {dataset.metadata.prediction_length}")
print(f"Frequency of the time series: {dataset.metadata.freq}")
estimator = DeepAREstimator(
distr_output=ZeroInflatedNegativeBinomialOutput(),
cell_type='GRU',
input_size=62,
num_cells=64,
num_layers=3,
dropout_rate=0.1,
use_feat_dynamic_real=True,
use_feat_static_cat=True,
cardinality=[int(cat_feat_info.cardinality) for cat_feat_info in dataset.metadata.feat_static_cat],
embedding_dimension = [4, 4, 4, 4, 16],
prediction_length=dataset.metadata.prediction_length,
context_length=dataset.metadata.prediction_length*2,
freq=dataset.metadata.freq,
scaling=True,
trainer=Trainer(device=device,
epochs=25,
learning_rate=1e-3,
num_batches_per_epoch=120,
batch_size=256,
)
)
predictor = estimator.train(dataset.train, num_workers=8, shuffle_buffer_length=512)
forecast_it, ts_it = make_evaluation_predictions(
dataset=dataset.test, # test dataset
predictor=predictor, # predictor
num_samples=100, # number of sample paths we want for evaluation
)
forecasts = list(forecast_it)
tss = list(ts_it)
evaluator = Evaluator()
agg_metrics, item_metrics = evaluator(iter(tss), iter(forecasts), num_series=len(dataset.test))
print(json.dumps(agg_metrics, indent=4))
item_metrics.plot(x='MSIS', y='MASE', kind='scatter')
plt.grid(which="both")
plt.show()
```
| github_jupyter |
# TensorFlow によるモデルの学習とマルチモデルエンドポイントでのホスティング
こちらは、TensorFlow を SageMaker 上で学習し、ひとつの推論エンドポイントに複数のモデルをデプロイする [マルチモデルエンドポイント (MME)](https://docs.aws.amazon.com/ja_jp/sagemaker/latest/dg/multi-model-endpoints.html#multi-model-endpoint-instance) の機能を使ってモデルをホスティングするノートブックです。このノートブックでは、SageMaker の pre-build Tensorflow コンテナを使う方法と、OSS の [Multi-Model Server (MMS)](https://github.com/awslabs/multi-model-server) を使う方法の 2通りをご紹介します。状況に合わせて適切な方法をご選択ください。2021年5月現在、マルチモデルエンドポイントは GPU インスタンスには対応していません。
[SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk) は、SageMaker学習インスタンスへのスクリプトの転送を処理します。学習インスタンスでは、SageMakerのネイティブTensorFlowサポートが学習関連の環境変数を設定し、学習スクリプトを実行します。このチュートリアルでは、SageMaker Python SDKを使用して学習ジョブを起動し、学習されたモデルを展開します。
TensorFlow Training についての詳細についてはこちらの[ドキュメント](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/using_tf.html#train-a-model-with-tensorflow)にアクセスしてください
このノートブックでは、マルチモデルエンドポイントを実現するための 2通りの方法をご紹介しますが、それぞれの基本的な違いは以下の通りです。
- SageMaker の pre-build Tensorflow コンテナを使う方法
- Tensorflow 2.2.2 かそれより新しいバージョンを使用する場合、かつ、推論データの前処理、後処理が不要な場合。<br>基本的にこちらの方法がシンプルでおすすめ。
- Multi-Model Server を使う方法
- マルチモデルエンドポイントがサポートされていないバージョンの Tensorflow を使う場合、または、推論データの前処理、後処理が必要な場合。
**※本ノートブックは TensorFlow バージョン 2 以上で動作します。**
---
## コンテンツ
モデルを学習する部分は通常の SageMaker 学習ジョブで行います。学習ジョブによって作成された学習済みモデルを使ってマルチモデルエンドポイントを作成します。
1. [環境のセットアップ](#1.-環境のセットアップ)
1. [学習データの準備](#2.-学習データの準備)
1. [分散学習用のスクリプトを作成する](#3.-分散学習用のスクリプトを作成する)
1. [TensorFlow Estimator を利用して学習ジョブを作成する](#4.-TensorFlow-Estimatorを利用して学習ジョブを作成する)
1. [Pre-build Tensorflow コンテナを使ってマルチモデルエンドポイントを作成する](#5.-Pre-build-Tensorflow-コンテナを使ってマルチモデルエンドポイントを作成する)
1. [Multi-Model Server を使ってマルチモデルエンドポイントを作成する](#6.-Multi-Model-Server-を使ってマルチモデルエンドポイントを作成する)
1. [エンドポイントを削除する](#7.エンドポイントを削除する)
---
# 1. 環境のセットアップ
まずは環境のセットアップを行いましょう。
```
import os, sagemaker, urllib
import matplotlib.pyplot as plt
import numpy as np
import boto3
from sagemaker import get_execution_role
sagemaker_session = sagemaker.Session()
sm_client = boto3.client(service_name='sagemaker')
runtime_sm_client = boto3.client(service_name='sagemaker-runtime')
role = get_execution_role()
region = sagemaker_session.boto_session.region_name
account_id = boto3.client('sts').get_caller_identity().get('Account')
s3_output = sagemaker_session.default_bucket()
s3_prefix = 'tensorflow-mme'
tag = ':latest'
print(f'Current SageMaker Python SDK Version = {sagemaker.__version__}')
```
注) このノートブックでは SageMaker SDK が 2.19.0 以上で動作します。上記の出力結果がそれ以前のバージョンになった際は、下記のセルの#を削除(コメントアウトを解除)して実行、Jupyterカーネルを再起動し、再度上記のセルを実行し、バージョンがアップデートされたことを確認してください。カーネルが再起動されない場合は、SageMaker SDK バージョン更新が反映されません。
```
# !pip install -U --quiet "sagemaker>=2.19.0"
```
# 2. 学習データの準備
MNISTデータセットは、パブリックS3バケット ``sagemaker-sample-data-<REGION>`` の下のプレフィックス ``tensorflow/mnist`` の下にロードされています。 このプレフィックスの下には4つの ``.npy`` ファイルがあります:
* ``train_data.npy``
* ``eval_data.npy``
* ``train_labels.npy``
* ``eval_labels.npy``
学習データが保存されている s3 の URI を変数に格納しておきます。
```
training_data_uri = f's3://sagemaker-sample-data-{region}/tensorflow/mnist/'
print(training_data_uri)
!aws s3 ls {training_data_uri}
```
# 3. 分散学習用のスクリプトを作成する
このチュートリアルの学習スクリプトは、TensorFlowの公式の [CNN MNISTの例](https://www.tensorflow.org/tutorials/images/cnn?hl=ja) をベースに作成されました。 SageMaker から渡された `` model_dir`` パラメーターを処理するように変更しています。 これは、分散学習時のデータ共有、チェックポイント、モデルの永続保存などに使用できるS3パスです。 また、学習関連の変数を扱うために、引数をパースする関数も追加しました。
学習ジョブの最後に、学習済みモデルを環境変数 ``SM_MODEL_DIR`` に保存されているパスにエクスポートするステップを追加しました。このパスは常に ``/opt/ml/model`` をポイントします。 SageMaker は、学習の終了時にこのフォルダー内のすべてのモデル成果物をS3にアップロードするため、これは重要です。
スクリプト全体は次のとおりです。
```
!pygmentize 'mnist.py'
```
# 4. TensorFlow Estimatorを利用して学習ジョブを作成する
`sagemaker.tensorflow.TensorFlow` estimator は、スクリプトモード対応の TensorFlow コンテナの指定、学習・推論スクリプトの S3 へのアップロード、および SageMaker 学習ジョブの作成を行います。ここでいくつかの重要なパラメーターを呼び出しましょう。
* `py_version`は` 'py3'`に設定されています。レガシーモードは Python 2 のみをサポートしているため、この学習スクリプトはスクリプトモードを使用していることを示しています。Python2は間もなく廃止されますが、 `py_version` を設定することでPython 2でスクリプトモードを使用できます。`'py2'`と` script_mode`を `True`にします。
* `distributions` は、分散学習設定を構成するために使用されます。インスタンスのクラスターまたは複数の GPU をまたいで分散学習を行う場合にのみ必要です。ここでは、分散学習スキーマとしてパラメーターサーバーを使用しています。 SageMaker 学習ジョブは同種のクラスターで実行されます。 SageMaker セットアップでパラメーターサーバーのパフォーマンスを向上させるために、クラスター内のすべてのインスタンスでパラメーターサーバーを実行するため、起動するパラメーターサーバーの数を指定する必要はありません。スクリプトモードは、[Horovod](https://github.com/horovod/horovod) による分散学習もサポートしています。 `distributions` の設定方法に関する詳細なドキュメントは[こちら](https://github.com/aws/sagemaker-python-sdk/tree/master/src/sagemaker/tensorflow#distributed-training) をご参照ください。
* 実際にモデル開発をする際はコード(ここでは `mnist.py` )にバグが混入していないか確認しながら実行することになりますが、学習インスタンスを利用すると、インスタンスの起動に時間がかかるため、学習開始コマンドを打ち込んでから 10 分後に気づいてやり直し、となってしまうことがあります。そのオーバヘッドを防止するために、ローカルモードでの学習が Sagemaker ではサポートされています。``instance_type=local``を指定するだけで、ノートブックインスタンスで学習(=インスタンスの立ち上げ時間なしで)を試すことができます。よくやるやり方としてはコードの確認用途のため、 epoch の数やデータを減らして動くかどうかの確認を行うことが多いです。
また、Spot Instanceを用いて実行する場合は、下記のコードを `Estimator` の `train_instance_type` の次の行に追加しましょう。
```python
max_run = 5000, # 学習は最大で5000秒までにする設定
use_spot_instances = 'True',
max_wait = 7200 # 学習完了を待つ最大時間
```
```
from sagemaker.tensorflow import TensorFlow
mnist_estimator = TensorFlow(entry_point='mnist.py',
role=role,
instance_count=2,
# instance_type='local',
instance_type='ml.p3.2xlarge',
framework_version='2.2.2',
py_version='py37',
distribution={'parameter_server': {'enabled': True}},
hyperparameters={
"epochs": 4,
'batch-size':16
}
# max_run = 5000, # 学習は最大で5000秒までにする設定
# use_spot_instances = 'True',
# max_wait = 7200 # 学習完了を待つ最大時間
)
```
## ``fit`` による学習ジョブの実行
学習ジョブを開始するには、`estimator.fit(training_data_uri)` を呼び出します。
ここでは、S3 ロケーションが入力として使用されます。 `fit` は、`training` という名前のデフォルトチャネルを作成します。これは、このS3ロケーションを指します。学習スクリプトでは、 `SM_CHANNEL_TRAINING` に保存されている場所から学習データにアクセスできます。 `fit`は、他のいくつかのタイプの入力も受け入れます。詳細については、APIドキュメント[こちら](https://sagemaker.readthedocs.io/en/stable/estimators.html#sagemaker.estimator.EstimatorBase.fit) を参照してください。
学習が開始されると、TensorFlow コンテナは mnist.py を実行し、スクリプトの引数として estimator から`hyperparameters` と `model_dir` を渡します。この例では、estimator 内で定義していないハイパーパラメーターは渡されず、 `model_dir` のデフォルトは `s3://<DEFAULT_BUCKET>/<TRAINING_JOB_NAME>` であるため、スクリプトの実行は次のようになります。
```bash
python mnist.py --model_dir s3://<DEFAULT_BUCKET>/<TRAINING_JOB_NAME>
```
学習が完了すると、学習ジョブは保存されたモデルを TensorFlow serving にアップロードします。
```
mnist_estimator.fit(training_data_uri)
```
マルチモデルエンドポイントは、指定された S3 パスにデプロイしたい全てのモデルを保存します。このノートブックでは、先ほど実行した学習ジョブの学習済みモデルが保存されているパスをマルチモデルエンドポイント用のパスとして使用します。
以下のセルでは、学習済みモデルが保存されたパスを取得しています。
```
import os
dirname = os.path.dirname(mnist_estimator.model_data)
dirname
```
複数のモデルを学習させるのは時間がかかるので、このノートブックでは先ほど学習したモデルを複製してデプロイします。
以下のセルでは、S3 に保存されている model.tar.gz を model2.tar.gz から model7.tar.gz までの 6 回複製しています。
```
!aws s3 cp $dirname/model.tar.gz $dirname/model2.tar.gz
!aws s3 cp $dirname/model.tar.gz $dirname/model3.tar.gz
!aws s3 cp $dirname/model.tar.gz $dirname/model4.tar.gz
!aws s3 cp $dirname/model.tar.gz $dirname/model5.tar.gz
!aws s3 cp $dirname/model.tar.gz $dirname/model6.tar.gz
!aws s3 cp $dirname/model.tar.gz $dirname/model7.tar.gz
```
あとで推論を実行する際に使用する入力データを準備しておきます。
```
!aws --region {region} s3 cp s3://sagemaker-sample-data-{region}/tensorflow/mnist/eval_data.npy eval_data.npy
!aws --region {region} s3 cp s3://sagemaker-sample-data-{region}/tensorflow/mnist/eval_labels.npy eval_labels.npy
eval_data = np.load('eval_data.npy').reshape(-1,28,28,1)
eval_labels = np.load('eval_labels.npy')
```
データセットから 50 枚のみ抜き出して `test_data` とします。
```
k = 1000 # choose your favorite number from 0 to 9950
test_data = eval_data[k:k+50]
test_data
for i in range(5):
for j in range(10):
plt.subplot(5, 10, 10* i + j+1)
plt.imshow(test_data[10 * i + j, :].reshape(28, 28), cmap='gray')
plt.title(10* i + j+1)
plt.tick_params(labelbottom=False, labelleft = False)
plt.subplots_adjust(wspace=0.2, hspace=1)
plt.show()
```
# 5. Pre-build Tensorflow コンテナを使ってマルチモデルエンドポイントを作成する
まずは、SageMaker が用意している pre-build Tensorflow コンテナを使ってマルチモデルエンドポイントを作成してみましょう。Tensorflow 2.2.2 を含む新しいバージョンがマルチモデルエンドポイントに対応しています。Pre-build コンテナの一覧は [こちら](https://github.com/aws/deep-learning-containers/blob/master/available_images.md) から参照可能です。古いバージョンや、SageMaker が用意していないバージョンの Tensorflow を使いたい場合は、[6. Multi-Model Server を使ってマルチモデルエンドポイントを作成する](#6.-Multi-Model-Server-を使ってマルチモデルエンドポイントを作成する) の方法を使用してください。
まず TensorFlowModel を作成し、それを引数として MultiDataModel を作成します。
```
from sagemaker.tensorflow.serving import TensorFlowModel
model = TensorFlowModel(role=role,
image_uri= '763104351884.dkr.ecr.us-east-1.amazonaws.com/tensorflow-inference:2.2.2-cpu-py37-ubuntu18.04',
model_data=mnist_estimator.model_data)
```
MultiDataModel の引数 `model_data_prefix` には、デプロイしたいモデルたちが保存されている S3 パスを指定します。
```
from sagemaker.multidatamodel import MultiDataModel
from time import gmtime, strftime
endpoint_name = 'tensorflow-mnist-mme-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
model_name = endpoint_name
mme = MultiDataModel(name=model_name,
model_data_prefix=dirname + '/',
model=model,# passing our model - passes container image needed for the endpoint
sagemaker_session=sagemaker_session)
```
MultiDataModel の deploy() を使って推論エンドポイントを起動します。このエンドポイントがマルチモデルエンドポイントとなります。エンドポイントの起動には 10 分ほどかかります。セルの下にしばらく - が表示されたのち、最後に ! が表示されたらエンドポイントの起動完了です。
```
predictor = mme.deploy(initial_instance_count=1,
instance_type='ml.m5.xlarge',
endpoint_name=endpoint_name)
```
エンドポイントにデプロイされているモデルの一覧を見てみましょう。先ほど複製した合計 7 つのモデルが表示されます。
```
list(mme.list_models())
```
それでは、起動完了した推論エンドポイントに推論リクエストを投げて推論を実行しましょう。`instances` というラベルで入力データを指定して predict() を実行します。合わせて、`TargetModel` に使用したいモデル名を指定します。
```
%%time
payload = {'instances': test_data.reshape(-1,28,28,1)}
predictions = predictor.predict(data=payload, initial_args={'TargetModel': 'model.tar.gz'})
```
推論結果を確認します。0.96 ほどの精度で MNIST 画像分類ができていることがわかります。
```
count_true = 0
for i in range(0, 50):
prediction = np.argmax(predictions['predictions'][i])
label = eval_labels[i+k]
if prediction == label:
count_true += 1
print(' [{}]: prediction is {}, label is {}, matched: {}'.format(i+1, prediction, label, prediction == label))
print('Accuracy: ', (count_true/50.0))
```
次に、`model2.tar.gz` を使って推論を実行します。
```
%%time
predictions = predictor.predict(data=payload, initial_args={'TargetModel': 'model2.tar.gz'})
```
もう一度、`model2.tar.gz` を使って推論を実行します。先ほどの初回実行時と推論時間がどれくらい変わったでしょうか。
```
%%time
predictions = predictor.predict(data=payload, initial_args={'TargetModel': 'model2.tar.gz'})
```
さらに、`model3.tar.gz` を使って推論を実行します。初回実行時は推論結果が返ってくるまでに 3秒程度かかりますが、2回目以降の呼び出しでは 100 ms ほどになっていたのではないでしょうか。これは、初回は S3 からモデルをダウンロードする必要がありますが、2回目以降はモデルがメモリにキャッシュされるためです。メモリに乗り切らないほど多数、もしくはサイズの大きいのモデルをデプロイした場合、メモリからは追い出されますが推論エンドポイントにアタッチされたストレージにモデルは保存されるため、初回推論時ほどの時間はかかりません。
```
%%time
predictions = predictor.predict(data=payload, initial_args={'TargetModel': 'model3.tar.gz'})
```
## 新しいモデルのアップロード
新しくモデルを追加してみましょう。モデルの追加のためにエンドポイントの設定などを変更する必要はありません。デプロイ済みのモデルが保存されている S3 パスに新しいモデルをアップロードします。
```
!aws s3 cp $dirname/model.tar.gz $dirname/model8.tar.gz
```
アップロードしたモデルがマルチモデルエンドポイントの参照先に反映されているか確認します。
```
list(mme.list_models())
```
新しいモデルを使って推論を実行します。
```
%%time
payload = {'instances': test_data.reshape(-1,28,28,1)}
predictions = predictor.predict(data=payload, initial_args={'TargetModel': 'model8.tar.gz'})
```
# 6. Multi-Model Server を使ってマルチモデルエンドポイントを作成する
前処理、後処理を定義したい、MME が対応していないバージョンの Tensorflow を使いたい場合、OSS の [Multi-Model Server](https://github.com/awslabs/multi-model-server) を使って MME を実現することが可能です。
まずは、推論で使用するコンテナをビルドする準備をします。
```
!mkdir -p docker/inference
%%writefile docker/inference/Dockerfile
# FROM ubuntu:18.04
FROM tensorflow/tensorflow:2.2.2-py3
# Set a docker label to advertise multi-model support on the container
LABEL com.amazonaws.sagemaker.capabilities.multi-models=true
# Set a docker label to enable container to use SAGEMAKER_BIND_TO_PORT environment variable if present
LABEL com.amazonaws.sagemaker.capabilities.accept-bind-to-port=true
# Install necessary dependencies for MMS and SageMaker Inference Toolkit
RUN apt-get update && \
apt-get -y install --no-install-recommends \
build-essential \
ca-certificates \
openjdk-8-jdk-headless \
python3-dev \
curl \
vim \
&& rm -rf /var/lib/apt/lists/* \
&& curl -O https://bootstrap.pypa.io/get-pip.py \
&& python3 get-pip.py
RUN update-alternatives --install /usr/bin/python python /usr/bin/python3 1
RUN update-alternatives --install /usr/local/bin/pip pip /usr/local/bin/pip3 1
# Install MXNet, MMS, and SageMaker Inference Toolkit to set up MMS
RUN pip3 --no-cache-dir install \
multi-model-server \
sagemaker-inference \
retrying
# Copy entrypoint script to the image
COPY dockerd-entrypoint.py /usr/local/bin/dockerd-entrypoint.py
RUN chmod +x /usr/local/bin/dockerd-entrypoint.py
RUN mkdir -p /home/model-server/
# Copy the default custom service file to handle incoming data and inference requests
COPY model_handler.py /home/model-server/model_handler.py
# Define an entrypoint script for the docker image
ENTRYPOINT ["python", "/usr/local/bin/dockerd-entrypoint.py"]
# Define command to be passed to the entrypoint
CMD ["serve"]
%%writefile docker/inference/dockerd-entrypoint.py
import subprocess
import sys
import shlex
import os
from retrying import retry
from subprocess import CalledProcessError
from sagemaker_inference import model_server
def _retry_if_error(exception):
return isinstance(exception, CalledProcessError or OSError)
@retry(stop_max_delay=1000 * 50,
retry_on_exception=_retry_if_error)
def _start_mms():
# by default the number of workers per model is 1, but we can configure it through the
# environment variable below if desired.
# os.environ['SAGEMAKER_MODEL_SERVER_WORKERS'] = '2'
model_server.start_model_server(handler_service='/home/model-server/model_handler.py:handle')
def main():
if sys.argv[1] == 'serve':
_start_mms()
else:
subprocess.check_call(shlex.split(' '.join(sys.argv[1:])))
# prevent docker exit
subprocess.call(['tail', '-f', '/dev/null'])
main()
```
推論リクエストを処理する部分を model_handler.py で定義します。`initialize()` で学習済みモデルをロードし、`preprocess()` でデータの前処理、`inference()` で推論実行、`postprocess()` で推論結果の後処理をします。
```
%%writefile docker/inference/model_handler.py
from collections import namedtuple
import glob
import json
import logging
import os
import re
import numpy as np
import tensorflow as tf
import os,json,argparse
from tensorflow.keras.layers import *
from tensorflow.keras.models import *
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.losses import categorical_crossentropy
class ModelHandler(object):
"""
A sample Model handler implementation.
"""
def __init__(self):
self.initialized = False
self.model = None
self.shapes = None
def initialize(self, context):
"""
Initialize model. This will be called during model loading time
:param context: Initial context contains model server system properties.
:return:
"""
self.initialized = True
properties = context.system_properties
# Contains the url parameter passed to the load request
model_dir = properties.get("model_dir")
# Load model
try:
self.model = tf.keras.models.load_model(os.path.join(model_dir, '000000001'))
except (RuntimeError) as memerr:
if re.search('Failed to allocate (.*) Memory', str(memerr), re.IGNORECASE):
logging.error("Memory allocation exception: {}".format(memerr))
raise MemoryError
raise
def preprocess(self, request):
"""
Transform raw input into model input data.
:param request: list of raw requests
:return: list of preprocessed model input data
"""
# Take the input data and pre-process it make it inference ready
payload = request[0]['body']
data = np.frombuffer(payload, dtype=np.float32).reshape(-1,28,28,1)
return data
def inference(self, model_input):
"""
Internal inference methods
:param model_input: transformed model input data list
:return: list of inference output in NDArray
"""
prediction = self.model.predict(model_input)
return prediction
def postprocess(self, inference_output):
"""
Return predict result in as list.
:param inference_output: list of inference output
:return: list of predict results
"""
print('======inference=======')
return [str(inference_output.tolist())]
def handle(self, data, context):
"""
Call preprocess, inference and post-process functions
:param data: input data
:param context: mms context
"""
model_input = self.preprocess(data)
model_out = self.inference(model_input)
return self.postprocess(model_out)
_service = ModelHandler()
def handle(data, context):
if not _service.initialized:
_service.initialize(context)
if data is None:
return None
return _service.handle(data, context)
```
作成したファイルを使って Docker イメージをビルドし、Amazon ECR に push します。
```
ecr_repository_inference = 'tensorflow-mme'
uri_suffix = 'amazonaws.com'
inference_repository_uri = '{}.dkr.ecr.{}.{}/{}'.format(account_id, region, uri_suffix, ecr_repository_inference + tag)
# Create ECR repository and push docker image
!docker build -t $ecr_repository_inference docker/inference
!$(aws ecr get-login --region $region --registry-ids $account_id --no-include-email)
!aws ecr create-repository --repository-name $ecr_repository_inference
!docker tag {ecr_repository_inference + tag} $inference_repository_uri
!docker push $inference_repository_uri
```
push したコンテナイメージを使って `create_model` を実行します。コンテナイメージを指定する際に、デプロイしたいモデルたちが保存されている S3 パスも指定します。
```
from time import gmtime, strftime
model_name = 'tf-MultiModelModel-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
container = {
'Image': inference_repository_uri,
'ModelDataUrl': dirname + '/',
'Mode': 'MultiModel'
}
create_model_response = sm_client.create_model(
ModelName = model_name,
ExecutionRoleArn = role,
Containers = [container])
print("Model Arn: " + create_model_response['ModelArn'])
```
`create_model` で作成したモデルを使って `create_endpoint_config` を実行してエンドポイント設定を作成します。
```
endpoint_config_name = 'tf-MultiModelEndpointConfig-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print('Endpoint config name: ' + endpoint_config_name)
create_endpoint_config_response = sm_client.create_endpoint_config(
EndpointConfigName = endpoint_config_name,
ProductionVariants=[{
'InstanceType': 'ml.m5.xlarge',
'InitialInstanceCount': 2,
'InitialVariantWeight': 1,
'ModelName': model_name,
'VariantName': 'AllTraffic'}])
print("Endpoint config Arn: " + create_endpoint_config_response['EndpointConfigArn'])
```
作成したエンドポイント設定を使って推論エンドポイントを起動します。このエンドポイントがマルチモデルエンドポイントとなります。エンドポイントの起動には 10 分ほどかかります。
```
import time
endpoint_name_mms = 'tensorflow-mnist-mme-mms-' + strftime("%Y-%m-%d-%H-%M-%S", gmtime())
print('Endpoint name: ' + endpoint_name_mms)
create_endpoint_response = sm_client.create_endpoint(
EndpointName=endpoint_name_mms,
EndpointConfigName=endpoint_config_name)
print('Endpoint Arn: ' + create_endpoint_response['EndpointArn'])
resp = sm_client.describe_endpoint(EndpointName=endpoint_name_mms)
status = resp['EndpointStatus']
print("Endpoint Status: " + status)
print('Waiting for {} endpoint to be in service...'.format(endpoint_name_mms))
waiter = sm_client.get_waiter('endpoint_in_service')
waiter.wait(EndpointName=endpoint_name_mms)
```
推論エンドポイントの起動が完了したら、`invoke_endpoint()` を使って推論を実行します。まずは `model.tar.gz` を使って推論を実行します。
```
%%time
import json
predictions = runtime_sm_client.invoke_endpoint(
EndpointName=endpoint_name_mms,
ContentType='application/x-npy',
TargetModel='model.tar.gz', # this is the rest of the S3 path where the model artifacts are located
Body=test_data.reshape(-1,28,28,1).tobytes())
```
推論エンドポイントからは str として結果が返ってくるので、それを numpy に変換します。
```
pred_str = predictions['Body'].read().decode('utf-8')
pred = np.array(json.loads(pred_str))
```
推論結果をラベルデータと比較します。問題なく推論できていそうです。
```
count_true = 0
for i in range(0, 50):
prediction = np.argmax(pred[i])
label = eval_labels[i+k]
if prediction == label:
count_true += 1
print(' [{}]: prediction is {}, label is {}, matched: {}'.format(i+1, prediction, label, prediction == label))
print('Accuracy: ', (count_true/50.0))
```
もう一度、`model.tar.gz` で推論を実行してみます。推論にかかる時間はどれくらい変わったでしょうか?
```
%%time
import json
predictions = runtime_sm_client.invoke_endpoint(
EndpointName=endpoint_name_mms,
ContentType='application/x-npy',
TargetModel='model.tar.gz', # this is the rest of the S3 path where the model artifacts are located
Body=test_data.reshape(-1,28,28,1).tobytes())
```
今度は `model2.tar.gz` を実行してみます。すぐ上のセルに表示された実行時間と比べてどうでしょうか。
初回実行時は推論結果が返ってくるまでに 6 秒程度かかりますが、2回目以降の呼び出しでは 100 ms ほどになっていたのではないでしょうか。これは、初回は S3 からモデルをダウンロードする必要がありますが、2回目以降はモデルがメモリにキャッシュされるためです。メモリに乗り切らないほど多数、もしくはサイズの大きいのモデルをデプロイした場合、メモリからは追い出されますが推論エンドポイントにアタッチされたストレージにモデルは保存されるため、初回推論時ほどの時間はかかりません。
```
%%time
import json
predictions = runtime_sm_client.invoke_endpoint(
EndpointName=endpoint_name_mms,
ContentType='application/x-npy',
TargetModel='model2.tar.gz', # this is the rest of the S3 path where the model artifacts are located
Body=test_data.reshape(-1,28,28,1).tobytes())
```
## 新しいモデルのアップロード
新しくモデルを追加してみましょう。モデルの追加のためにエンドポイントの設定などを変更する必要はありません。デプロイ済みのモデルが保存されている S3 パスに新しいモデルをアップロードします。
```
!aws s3 cp $dirname/model.tar.gz $dirname/model9.tar.gz
```
アップロードしたモデルを使って推論を実行します。
```
%%time
import json
predictions = runtime_sm_client.invoke_endpoint(
EndpointName=endpoint_name_mms,
ContentType='application/x-npy',
TargetModel='model9.tar.gz', # this is the rest of the S3 path where the model artifacts are located
Body=test_data.reshape(-1,28,28,1).tobytes())
```
# 7.エンドポイントを削除する
推論用エンドポイントは停止されるまで課金が発生します。そのため。不要になったエンドポイントはすぐに削除することをおすすめします。以下のコードでエンドポイントが削除されます。AWS コンソールの左側のメニューから「エンドポイント」をクリックし、停止したいエンドポイントを選択して削除することも可能です。
こちらは、pre-build コンテナを使用して作成したエンドポイントを削除するコードです。
```
predictor.delete_endpoint()
```
こちらは、カスタムコンテナと Multi-Model Server を使って作成したエンドポイントを削除するコードです。
```
response = sm_client.delete_endpoint(
EndpointName=endpoint_name_mms
)
```
| github_jupyter |
# Chapter 4 - Boolean Expressions and Conditions
*This notebook uses code snippets and explanations from [this course](https://github.com/kadarakos/python-course/blob/master/Chapter%202%20-%20Conditions.ipynb).*
So far, we have learned how to use Python as a basic calculator and how to store information in variables. Now we will set the first steps to an actual useful program. A lot of programming has to do with executing code if a particular condition holds. This enables a program to act upon its inputs. For example: an app on your phone might give a warning if the battery level is lower than 5%. This means that the program needs to check if the variable `battery_level` is lower than the value of 5. We can do these checks using so called Boolean expressions. These Boolean expressions are the main element in probably one of the most used things in Python: *if statements*.
#### At the end of this topic, you will be able to:
* work with and understand *boolean expressions*
* work with and understand *if statements*
* understand what *indentation* is
* understand what *nesting* is
#### If you want to learn more about these topics, you might find the following links useful:
* Documentation: [Built-in Types (boolean expressions)](https://docs.python.org/3.5/library/stdtypes.html#)
* Video: [Python Booleans](https://www.youtube.com/watch?v=9OK32jb_TdI)
* Video: [Conditionals](https://www.youtube.com/watch?v=mQrci1kAwh4)
* Explanation: [if elif else](http://www.programiz.com/python-programming/if-elif-else)
If you have **questions** about this chapter, please contact us(cltl.python.course@gmail.com)**.
## 1. Boolean expressions
A **Boolean expression** (or simply: boolean) is an expression that results in the type `bool` in Python. Possible values are either **`True`** or **`False`**. Boolean expressions are the building blocks of programming. Any expression that results in `True` or `False` can be considered a Boolean expression.
So far you've mainly seen:
```
print(type('this is a string'))
print(type(101))
print(type(0.8))
```
Now we're introducing:
```
print(type(False))
print(type(True))
```
### 1.1 Comparison operators
Here is a list of **[comparison operators](https://docs.python.org/3/library/stdtypes.html#comparisons)** used in Boolean expressions:
(You have already used these operators in the previous chapters, but we are treating them in more detail here.)
| Operator | Meaning | `True` | `False` |
|-----------|--------|--------|--------|
| `==` | equal | `2 == 2` | `2 == 3` |
| `!=` | not equal | `3 != 2` | `2 != 2` |
| `<` | less than | `2 < 13` | `2 < 2` |
| `<=` | less than or equal to | `2 <= 2` | `3 <= 2` |
| `>` | greater than | `13 > 2` | `2 > 13` |
| `>=` | greather than or equal to | `3 >= 3` | `2 >= 3` |
Remember that the single = is reserved for assignment! Boolean expressions look at variables but never change them.
```
print(2 < 5)
print(2 <= 5)
print(3 > 7)
print(3 >= 7)
print(3 == 3)
print("school" == "school")
print("Python" != "SPSS")
```
The relevant 'logical operators' that we used here are: <, <=, >,>=,==,!=. In Python-speak, we say that such a logical expression gets 'evaluated' when you run the code. The outcome of such an evaluation is a 'binary value' or a so-called 'boolean' that can take only two possible values: `True` or `False`. You can assign such a boolean to a variable:
```
greater = 5 > 2
print(greater, type(greater))
greater = 5 < 2
print(greater, type(greater))
```
Let's look at some examples. Try to guess the output based on the information about the operators in the table above. Hence, will the expression result in `True` or `False` in the following examples?
```
print(5 == 5)
print(5 == 4)
print(10 < 20)
print(10 < 8)
print(10 < 10)
print(10 <= 10)
print(20 >= 21)
print(20 == 20)
print(1 == '1')
print(1 != 2)
boolean_expression = 5 == 4
print(boolean_expression)
```
### 1.2 Membership operators
Python also has so-called **[membership operators](https://docs.python.org/3.5/reference/expressions.html#not-in)**:
| Operator | function | `True` | `False` |
|-----------|--------|--------|--------|
| `in` | left object is a member of right object | `"c" in "cat"` | `"f" in "cat"` |
| `not in` |left object is NOT a member of right object | `"f" not in "cat"` | `"c" not in "cat"` |
We have already seen the operator **`in`** being used for checking whether a string (single or multiple characters) is a substring of another one:
```
print("fun" in "function")
print("pie" in "python")
```
We can only use membership operators with *iterables* (i.e. python objects that can be split up into smaller components - e.g. characters of a string). The following will therefore not work, because an integer is not iterable:
```
print(5 in 10)
```
However, we can use membership operators with other types of 'containers', such as *lists*. We will discuss lists in much more detail later on, but they represent ordered sequences of objects like strings, integers or a combination. We can use *in* and *not in* to check whether an object is a member of a list:
```
letters = ['a','b','c','d']
numbers = [1,2,3,4,5]
mixed = [1,2,3,'a','b','c']
print('a' in letters)
print('g' not in letters)
print('d' in mixed)
print(1 in numbers)
print(3 not in mixed)
print('a' not in 'hello world')
```
### 1.3 And, or, and not
Finally, boolean operations are often performed using the [**boolean operators `and`, `or` and `not`**](https://docs.python.org/3.5/library/stdtypes.html#boolean-operations-and-or-not). Given two boolean expressions, **bool1** and **bool2**, this is how they work:
| operation | function | `True` | `False` |
|-----------|--------|----------|---------|
| **bool1** `and` **bool2** | `True` if both **bool1** and **bool2** are `True`, otherwise `False` | (`5 == 5 and 3 < 5`) | (`5 == 5 and 3 > 5`) |
| **bool1** `or` **bool2** | `True` when at least one of the boolean expressions is `True`, otherwise `False` | (`5 == 5 or 3 > 5`) | (`5 != 5 or 3 > 5`) |
| `not` **bool1** | `True` if **bool1** is `False`, otherwise `False` | (`not 5 != 5`) | (`not 5 == 5`) |
Here are some examples of **`and`**:
```
letters = ['a','b','c','d']
numbers = [1,2,3,4,5]
print('a' in letters and 2 in numbers)
print("z" in letters and 3 in numbers)
print("f" in letters and 0 in numbers)
```
Here are some examples of **`or`**:
```
letters = ['a','b','c','d']
numbers = [1,2,3,4,5]
print('f' in letters or 2 in numbers)
print('a' in letters or 2 in numbers)
print('f' in letters or 10 in numbers)
```
Here are some example of **`not`**:
```
a_string = "hello"
letters = ['a','b','c','d']
numbers = [1,2,3,4,5]
print(not a_string.endswith("o"))
print(not a_string.startswith("o"))
print(not 'x' in letters)
print(not 4 == 4)
print(not (4 == 4 and "4" == 4))
```
Note that for some of these, there are alternative ways of writing them. For example, `'x not in y'` and `'not x in y'` are identical, and so are `'not x == y'` and `'x != y'`. For now, it does not really matter which one you use. If you want to read more about it, have a look [here](https://stackoverflow.com/questions/8738388/order-of-syntax-for-using-not-and-in-keywords) and [here](https://stackoverflow.com/questions/31026754/python-if-not-vs-if/31026976).
```
print(not 'x' in letters)
print('x' not in letters)
print(not 4 == 4)
print(4 != 4)
```
### 1.4 EXTRA: `all()` and `any()`
Take a look at the following example. Do you think it is clear?
```
print("test" != "testing" and 1 == 1 and 2 == 2 or 20 in [1, 20, 3, 4,5])
```
Not really, right? Luckily, Python has another trick to deal with this type of examples: [**`all` and `any`**]((https://docs.python.org/3/library/functions.html#all). Given a list of boolean expressions, this is how they work:
| operation | function |
|-----------|--------|
| `all` | True if all boolean expressions are True, otherwise False |
| `any` | True if at least one boolean expression is True, otherwise False |
If you don't completely understand `all()` and `any()`, don't worry, you will not necessarily need them right now. They are just a nice alternative to make your code more readable and you may appreciate that in the future.
Here are some examples of **`all()`**:
```
letters = ['a','b','c','d']
numbers = [1,2,3,4,5]
list_bools1 = ['a' in letters,
2 in numbers]
print(list_bools1)
boolean_expression1 = all(list_bools1)
print(boolean_expression1)
list_bools2 = ['a' in letters,
20 in numbers]
print(list_bools2)
boolean_expression2 = all(list_bools2)
print(boolean_expression2)
```
Here are some examples of **`any()`**:
```
list_bools3 = ['f' in letters,
200 in numbers]
print(list_bools3)
boolean_expression3 = any(list_bools3)
print(boolean_expression3)
list_bools4 = ['a' in letters,
20 in numbers,
2 in numbers]
print(list_bools4)
boolean_expression4 = any(list_bools4)
print(boolean_expression4)
```
## 2. Conditions: `if` statements
You might wonder why we took quite some time explaining boolean expresisons. One of the reasons is that they are the main element in probably one of the most used things in Python: **`if` statements**. The following picture explains what happens in an `if` statement in Python.

Let's look at an example (modify the value of *number* to understand what is happening here):
```
number = 2 # try changing this value to 6
if number <= 5:
print(number)
```
You can use as many `if` statements as you like:
```
number = 5
if number == 5:
print("number equals 5")
if number > 4:
print("number is greater than 4")
if number >= 5:
print("number is greater than or equals 5")
if number < 6:
print("number is less than 6")
if number <= 5:
print("number is less than or equals 5")
if number != 6 :
print("number does not equal 6")
```
### 2.1 Two-way decisions
But what if we want to have options for two different scenarios? We could just use a bunch of `if` statements. However, Python has a more efficient way. Apart from `if` we also have the **`else` statement** for two-way decisions (modify the value of `number` to understand what is happening here):
```
number = 10 # try changing this value to 2
if number <= 5:
print(number)
else:
print('number is higher than 5')
```
Now Python always runs one of the two pieces of code. It's like arriving at a fork in the road and choosing one path to follow.
### 2.2 Multi-way decisions
But of course we don't have to stop there. If you have multiple options, you can use the **`elif` statement**. For every `if` block, you can have one `if` statement, multiple `elif` statements and one `else` statement. So now we know the entire **`if-elif-else` construct**:

```
age = 21
if age < 12:
print("You're still a child!")
elif age < 18:
print("You are a teenager!")
elif age < 30:
print("You're pretty young!")
else:
print("Wow, you're old!")
```
First the `if` statement will be evaluated. Only if that statement turns out to be `False` the computer will proceed to evaluate the `elif` statement. If the `elif` statements in turn would prove to be `False`, the machine will proceed and execute the lines of code associated with the `else` statement. You can think of this coding structure as a decision tree! Remember: if somewhere along the tree, your machine comes across a logical expression which is `True`, it won't bother anymore to evaluate the remaining options! Note that the statements are evaluated in order of occurence.
Can you identify the difference between the code above and the code below? (Try changing `age`)
```
age = 21
if age < 12:
print("You're still a child!")
if age < 18:
print("You are a teenager!")
if age < 30:
print("You're pretty young!")
else:
print("Wow, you're old!")
```
**Remember:**
- `if-if`: your code wil check all the `if` statements
- `if-elif`: if one condition results to `True`, it will not check the other conditions
Unless you *need* to check all conditions, using `if-elif` is usually preferred because it's *more efficient*.
## 3. Indentation
Let's take another look at the example from above (we've added line numbers):
```python
1. if number <= 5:
2. print(number)
3. else:
4. print('number is higher than 5')
```
You might have noticed that line 2 starts with 4 spaces. This is on purpose! The indentation lets Python know when it needs to execute the piece of code. When the boolean expression in line 1 is `True`, Python executes the code from the next line that starts four spaces or one tab (an indent) to the right. This is called **indentation**. All statements with the same distance to the right belong to the same 'block' of code.
Unlike other languages, Python does not make use of curly braces to mark the start and end of pieces of code, like `if` statements. The only delimiter is a colon (:) and the indentation of the code. Both four spaces and tabs can be used for indentation. This indentation must be used consistently throughout your code. The most popular way to indent is four spaces (see [stackoverflow](http://stackoverflow.com/questions/120926/why-does-python-pep-8-strongly-recommend-spaces-over-tabs-for-indentation)). For now, you do not have to worry about this, since a tab is automatically converted to four spaces in notebooks.
Take a look at the code below. We see that the indented block is not executed, but the unindented lines of code are. Now go ahead and change the value of the `person` variable. The conversation should be a bit longer now!
```
person = "John"
print("hello!")
if person == "Alice":
print("how are you today?") #this is indented
print("do you want to join me for lunch?") #this is indented
elif person == "Lisa":
print("let's talk some other time!") #this is indented
print("goodbye!")
```
### 3.1 Nesting
We have seen that all statements with the same distance to the right belong to the same block of code, i.e. the statements within a block line up vertically. The block ends at a line less indented or the end of the file.
Blocks can contain blocks as welll; this way, we get a nested block structure. The block that has to be more deeply **nested** is simply indented further to the right:

There may be a situation when you want to check for another condition after a condition resolves to `True`. In such a situation, you can use the nested `if` construct. As you can see if you run the code below, the second `if` statement is only executed if the first `if` statement returns `True`. Try changing the value of x to see what the code does.
```
x = float(input("Enter a number: "))
if x >= 0:
if x == 0:
print("Zero")
else:
print("Positive number")
else:
print("Negative number")
```
## Exercises
### Exercise 1:
It's important to practice a lot with boolean expressions. Here is a list of them, which orginate from [learnpythonthehardway](http://learnpythonthehardway.org/book/ex28.html). Try to guess the output.
```
print(True and True)
print(False and True)
print(1 == 1 and 2 == 1)
print("test" == "test")
print(1 == 1 or 2 != 1)
print(True and 1 == 1)
print(False and 0 != 0)
print(True or 1 == 1)
print("test" == "testing")
print(1 != 0 and 2 == 1)
print("test" != "testing")
print("test" == 1)
print(not (True and False))
print(not (1 == 1 and 0 != 1))
print(not (10 == 1 or 1000 == 1000))
print(not (1 != 10 or 3 == 4))
print(not ("testing" == "testing" and "Zed" == "Cool Guy"))
print(1 == 1 and (not ("testing" == 1 or 1 == 0)))
print("chunky" == "bacon" and (not (3 == 4 or 3 == 3)))
print(3 == 3 and (not ("testing" == "testing" or "Python" == "Fun")))
print("test" != "testing" and 1 == 1 and 2 == 2 and 20 in [1, 20, 3, 4,5])
```
### Exercise 2:
Write a small program that defines a variable `weight`. If the weight is > 50 pounds, print "There is a $25 charge for luggage that heavy." If it is not, print: "Thank you for your business." If the weight is exactly 50, print: "Pfiew! The weight is just right!". Change the value of weight a couple of times to check whether your code works. Make use of the logical operators and the if-elif-else construct!
```
# insert your code here
```
### Exercise 3:
What's wrong in the following code? Correct the mistake.
```
my_string = "hello"
if my_string == "hello":
print("world")
```
Why is the last line in the following code red? Correct the mistake.
```
my_string = "hello"
if my_string == "hello":
print("world")
```
What's wrong in the following code? Correct the mistake.
```
my_string = "hello"
if my_string == "hello"
print("world")
```
What's wrong in the following code? Correct the mistake.
```
my_string = "hello"
if my_string = "hello":
print("world")
```
### Exercise 4:
Can you rewrite the code below without nesting? Hint: use the `if-elif-else` construct.
```
x = float(input("Enter a number: "))
if x >= 0:
if x == 0:
print("Zero")
else:
print("Positive number")
else:
print("Negative number")
```
### Exercise 5:
A friend wants your advice on how much oranges he should buy. Write a program that will give the advice to buy 24 oranges if the price is lower than 1.50 EUR per kg, 12 oranges if the price is between 1.50 EUR and 3 EUR, and only 1 orange if the price is higher than 3 EUR. But also tell him that he should only buy them if the oranges are fresh; otherwise, he should not get any. Use nesting and the if-elif-else construct.
```
orange_quality = "fresh"
orange_price = 1.75
# your code here
```
| github_jupyter |
# TopK benchmark
The notebook compares [onnxruntime](https://github.com/microsoft/onnxruntime) and [mlprodict](http://www.xavierdupre.fr/app/mlprodict/helpsphinx/index.html) implementation of operator [TopK](https://github.com/onnx/onnx/blob/master/docs/Operators.md#TopK).
```
from jyquickhelper import add_notebook_menu
add_notebook_menu()
%matplotlib inline
```
## Plots
We measure two runtimes by computing a ratio between their time execution through the following kind of graphs.
```
import matplotlib.pyplot as plt
from mlprodict.plotting.plotting import plot_benchmark_metrics
def plot_metric(metric, ax=None, xlabel="N", ylabel="k", middle=1.,
transpose=False, shrink=1.0, title=None):
ax, cbar = plot_benchmark_metrics(metric, ax=ax, xlabel=xlabel, ylabel=ylabel,
middle=middle, transpose=transpose,
cbar_kw={'shrink': shrink})
if title is not None:
ax.set_title(title)
return ax
data = {(1, 1): 0.1, (10, 1): 1, (1, 10): 2,
(10, 10): 100, (100, 1): 100, (100, 10): 1000}
fig, ax = plt.subplots(1, 2, figsize=(10, 4))
plot_metric(data, ax[0], shrink=0.6)
plot_metric(data, ax[1], transpose=True);
```
## TopK in ONNX
The following lines creates an ONNX graph using one TopK ONNX node. The outcome is the ONNX graph converted into json.
```
import numpy
from numpy.random import randn
from skl2onnx.algebra.onnx_ops import OnnxTopK_11
X32 = randn(100000, 100).astype(numpy.float32)
node = OnnxTopK_11('X', numpy.array([5], dtype=numpy.int64),
output_names=['dist', 'ind'])
model_onnx = node.to_onnx([('X', X32)], target_opset=12,
# shape inference does not seem to work in onnxruntime
# so we speccify the output shape
outputs=[('dist', X32[:1, :5]),
('ind', X32[:1, :5].astype(numpy.int64))])
model_onnx
from mlprodict.onnxrt import OnnxInference
oinf = OnnxInference(model_onnx, runtime="python")
res = oinf.run({'X': X32})
dist, ind = res['dist'], res['ind']
dist[:2], ind[:2]
from mlprodict.tools import get_ir_version_from_onnx
model_onnx.ir_version = get_ir_version_from_onnx()
from onnxruntime import InferenceSession
sess = InferenceSession(model_onnx.SerializeToString())
dist, ind = sess.run(None, {'X': X32})
dist[:2], ind[:2]
```
Let's compare two implementations.
```
import numpy
import sklearn
from mlprodict.onnxrt.validate.validate_benchmark import benchmark_fct
def benchmark(X, fct1, fct2, N, repeat=10, number=10):
def ti(n):
if n <= 1:
return 50
if n <= 1000:
return 2
if n <= 10000:
return 0.51
return 0.11
# to warm up the engine
time_kwargs = {n: dict(repeat=10, number=10) for n in N[:2]}
benchmark_fct(fct1, X, time_kwargs=time_kwargs, skip_long_test=False)
benchmark_fct(fct2, X, time_kwargs=time_kwargs, skip_long_test=False)
# real measure
time_kwargs = {n: dict(repeat=int(repeat * ti(n)),
number=int(number * ti(n))) for n in N}
res1 = benchmark_fct(fct1, X, time_kwargs=time_kwargs, skip_long_test=False)
res2 = benchmark_fct(fct2, X, time_kwargs=time_kwargs, skip_long_test=False)
res = {}
for r in sorted(res1):
r1 = res1[r]
r2 = res2[r]
ratio = r2['total'] / r1['total']
res[r] = ratio
return res
N = [1, 10, 100, 1000, 10000, 100000]
benchmark(X32, lambda x: sess.run(None, {'X': x}),
lambda x: oinf.run({'X': x}), N=N)
```
The implementation in [mlprodict](https://github.com/sdpython/mlprodict/blob/master/mlprodict/onnxrt/ops_cpu/_op_onnx_numpy.cpp#L246) is faster when the number of rows grows. It is faster for 1 rows, for many rows, the implementation uses openmp to parallelize.
## C++ implementation vs numpy
*scikit-learn* uses numpy to compute the top *k* elements.
```
from mlprodict.onnxrt.ops_cpu.op_topk import (
topk_sorted_implementation, topk_sorted_implementation_cpp)
benchmark(X32, lambda x: topk_sorted_implementation(x, 5, 1, 0),
lambda x: topk_sorted_implementation_cpp(x, 5, 1, 0), N=N)
```
It seems to be faster too. Let's profile.
```
from pyquickhelper.pycode.profiling import profile
from IPython.display import HTML
xr = randn(1000000, 100)
HTML(profile(lambda: topk_sorted_implementation(xr, 5, 1, 0),
pyinst_format='html')[1])
```
## Parallelisation
We need to disable the parallelisation to really compare both implementation.
```
from tqdm import tqdm
def benchmark_test(X, fct1, fct2, N, K, repeat=10, number=10):
res = {}
for k in tqdm(K):
f1 = lambda x, k=k: fct1(x, k=k)
f2 = lambda x, k=k: fct2(x, k=k)
r = benchmark(X32, f1, f2, N=N, repeat=repeat, number=number)
for n, v in r.items():
res[n, k] = v
return res
K = [1, 2, 5, 10, 15]
N = [1, 2, 3, 10, 100, 1000, 10000, 100000]
bench_para = benchmark_test(
X32, (lambda x, k: topk_sorted_implementation_cpp(x, k=k, axis=1, largest=0, th_para=100000000)),
(lambda x, k: topk_sorted_implementation_cpp(x, k=k, axis=1, largest=0, th_para=1)),
N=N, K=K)
plot_metric(bench_para, transpose=False, title="TopK and parallelisation\n"
"< 1 means parallelisation is faster", shrink=0.75);
```
This is somehow expected. First column is closed to 1 as it is the same code being compared. Next columns are red, meaning the parallelisation does not help, the parallelisation helps for bigger N, as least more than 100.
## Parallellisation with ONNX
We replicate the same experiment with an ONNX graph.
```
k_ = numpy.array([3], dtype=numpy.int64)
node = OnnxTopK_11('X', 'k',
output_names=['dist', 'ind'])
model_onnx = node.to_onnx([('X', X32), ('k', k_)], target_opset=12,
# shape inference does not seem to work in onnxruntime
# so we speccify the output shape
outputs=[('dist', X32[:1, :5]),
('ind', X32[:1, :5].astype(numpy.int64))])
oinf_no_para = OnnxInference(model_onnx, runtime="python")
res = oinf_no_para.run({'X': X32, 'k': k_})
dist, ind = res['dist'], res['ind']
dist[:2], ind[:2]
oinf_para = OnnxInference(model_onnx, runtime="python")
oinf_no_para.sequence_[0].ops_.th_para = 100000000
oinf_para.sequence_[0].ops_.th_para = 1
bench_onnx_para = benchmark_test(
X32, (lambda x, k: oinf_no_para.run({'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
(lambda x, k: oinf_para.run({'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
N=N, K=K)
plot_metric(bench_onnx_para, transpose=False, title="TopK and parallelisation with ONNX\n"
"< 1 means parallelisation is faster", shrink=0.75);
```
Pretty much the same results.
## onnxruntime vs mlprodict (no parallelisation)
```
model_onnx.ir_version = get_ir_version_from_onnx()
sess = InferenceSession(model_onnx.SerializeToString())
bench_ort = benchmark_test(
X32, (lambda x, k: sess.run(None, {'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
(lambda x, k: oinf_no_para.run({'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
N=N, K=K)
plot_metric(bench_ort, transpose=False, title="TopK, onnxruntime vs mlprodict\n"
"< 1 means mlprodict is faster\nno parallelisation", shrink=0.75);
```
It seems the implement of operator TopK in onnxruntime 1.1.1 can be improved.
```
from onnxruntime import __version__ as ort_version
from mlprodict import __version__ as mlp_version
ort_version, mlp_version
```
And with parallelisation above 50 rows.
```
oinf_para.sequence_[0].ops_.th_para = 50
bench_ort_para = benchmark_test(
X32, (lambda x, k: sess.run(None, {'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
(lambda x, k: oinf_para.run({'X': x, 'k': numpy.array([k], dtype=numpy.int64)})),
N=N, K=K)
plot_metric(bench_ort_para, transpose=False, title="TopK, onnxruntime vs mlprodict\n"
"< 1 means mlprodict is faster\nparallelisation above 50 rows", shrink=0.75);
```
Interesting...
## Comparison with onnxruntime
```
from skl2onnx.algebra.onnx_ops import OnnxTopK
from skl2onnx.common.data_types import FloatTensorType
from mlprodict.onnx_conv import to_onnx
from onnxruntime import InferenceSession
X = numpy.array([
[0, 1, 2, 3],
[4, 5, 6, 7],
[8, 9, 10, 11],
], dtype=numpy.float32)
K = numpy.array([3], dtype=numpy.int64)
node = OnnxTopK('X', K, output_names=['values', 'indices'],
op_version=12)
onx = node.to_onnx([('X', FloatTensorType())])
py_topk = OnnxInference(onx, runtime="python_compiled")
ort_topk = InferenceSession(onx.SerializeToString())
py_topk.run({'X': X})
ort_topk.run(None, {'X': X})
%timeit py_topk.run({'X': X})
%timeit ort_topk.run(None, {'X': X})
X = numpy.random.randn(10000, 100).astype(numpy.float32)
%timeit py_topk.run({'X': X})
%timeit ort_topk.run(None, {'X': X})
```
| github_jupyter |
# Data Wrangling
This notebook presents different data wrangling techniques used commonly
```
# import required libraries
import random
import datetime
import numpy as np
import pandas as pd
from random import randrange
from sklearn import preprocessing
pd.options.mode.chained_assignment = None
```
## Utilities
```
def _random_date(start,date_count):
"""This function generates a random date based on params
Args:
start (date object): the base date
date_count (int): number of dates to be generated
Returns:
list of random dates
"""
current = start
while date_count > 0:
curr = current + datetime.timedelta(days=randrange(42))
yield curr
date_count-=1
def generate_sample_data(row_count=100):
"""This function generates a random transaction dataset
Args:
row_count (int): number of rows for the dataframe
Returns:
a pandas dataframe
"""
# sentinels
startDate = datetime.datetime(2016, 1, 1,13)
serial_number_sentinel = 1000
user_id_sentinel = 5001
product_id_sentinel = 101
price_sentinel = 2000
# base list of attributes
data_dict = {
'Serial No': np.arange(row_count)+serial_number_sentinel,
'Date': np.random.permutation(pd.to_datetime([x.strftime("%d-%m-%Y")
for x in _random_date(startDate,
row_count)]).date
),
'User ID': np.random.permutation(np.random.randint(0,
row_count,
size=int(row_count/10)) + user_id_sentinel).tolist()*10,
'Product ID': np.random.permutation(np.random.randint(0,
row_count,
size=int(row_count/10))+ product_id_sentinel).tolist()*10 ,
'Quantity Purchased': np.random.permutation(np.random.randint(1,
42,
size=row_count)),
'Price': np.round(np.abs(np.random.randn(row_count)+1)*price_sentinel,
decimals=2),
'User Type':np.random.permutation([chr(random.randrange(97, 97 + 3 + 1))
for i in range(row_count)])
}
# introduce missing values
for index in range(int(np.sqrt(row_count))):
data_dict['Price'][np.argmax(data_dict['Price'] == random.choice(data_dict['Price']))] = np.nan
data_dict['User Type'][np.argmax(data_dict['User Type'] == random.choice(data_dict['User Type']))] = np.nan
data_dict['Date'][np.argmax(data_dict['Date'] == random.choice(data_dict['Date']))] = np.nan
data_dict['Product ID'][np.argmax(data_dict['Product ID'] == random.choice(data_dict['Product ID']))] = 0
data_dict['Serial No'][np.argmax(data_dict['Serial No'] == random.choice(data_dict['Serial No']))] = -1
data_dict['User ID'][np.argmax(data_dict['User ID'] == random.choice(data_dict['User ID']))] = -101
# create data frame
df = pd.DataFrame(data_dict)
return df
def describe_dataframe(df=pd.DataFrame()):
"""This function generates descriptive stats of a dataframe
Args:
df (dataframe): the dataframe to be analyzed
Returns:
None
"""
print("\n\n")
print("*"*30)
print("About the Data")
print("*"*30)
print("Number of rows::",df.shape[0])
print("Number of columns::",df.shape[1])
print("\n")
print("Column Names::",df.columns.values.tolist())
print("\n")
print("Column Data Types::\n",df.dtypes)
print("\n")
print("Columns with Missing Values::",df.columns[df.isnull().any()].tolist())
print("\n")
print("Number of rows with Missing Values::",len(pd.isnull(df).any(1).nonzero()[0].tolist()))
print("\n")
print("Sample Indices with missing data::",pd.isnull(df).any(1).nonzero()[0].tolist()[0:5])
print("\n")
print("General Stats::")
print(df.info())
print("\n")
print("Summary Stats::")
print(df.describe())
print("\n")
print("Dataframe Sample Rows::")
display(df.head(5))
def cleanup_column_names(df,rename_dict={},do_inplace=True):
"""This function renames columns of a pandas dataframe
It converts column names to snake case if rename_dict is not passed.
Args:
rename_dict (dict): keys represent old column names and values point to
newer ones
do_inplace (bool): flag to update existing dataframe or return a new one
Returns:
pandas dataframe if do_inplace is set to False, None otherwise
"""
if not rename_dict:
return df.rename(columns={col: col.lower().replace(' ','_')
for col in df.columns.values.tolist()},
inplace=do_inplace)
else:
return df.rename(columns=rename_dict,inplace=do_inplace)
def expand_user_type(u_type):
"""This function maps user types to user classes
Args:
u_type (str): user type value
Returns:
(str) user_class value
"""
if u_type in ['a','b']:
return 'new'
elif u_type == 'c':
return 'existing'
elif u_type == 'd':
return 'loyal_existing'
else:
return 'error'
```
## Generate a Sample Dataset
```
df = generate_sample_data(row_count=1000)
```
### Describe the Dataset
```
describe_dataframe(df)
```
### Rename Columns
```
print("Dataframe columns:\n{}".format(df.columns.tolist()))
cleanup_column_names(df)
print("Dataframe columns:\n{}".format(df.columns.tolist()))
```
### Sort Rows on defined attributes
```
display(df.sort_values(['serial_no', 'price'],
ascending=[True, False]).head())
```
### Rearrange Columns in a Dataframe
```
display(df[['serial_no','date','user_id','user_type',
'product_id','quantity_purchased','price']].head())
```
### Filtering Columns
Using Column Index
```
# print 10 values from column at index 3
print(df.iloc[:,3].values[0:10])
```
Using Column Name
```
# print 10 values of quantity purchased
print(df.quantity_purchased.values[0:10])
```
Using Column Datatype
```
# print 10 values of columns with data type float
print(df.select_dtypes(include=['float64']).values[:10,0])
```
### Filtering Rows
Select specific rows
```
display(df.iloc[[10,501,20]])
```
Exclude Specific Row indices
```
display(df.drop([0,24,51], axis=0).head())
```
Conditional Filtering
```
display(df[df.quantity_purchased>25].head())
```
Offset from top of the dataframe
```
display(df[100:].head())
```
Offset from bottom of the dataframe
```
display(df[-10:].head())
```
### TypeCasting/Data Type Conversion
```
df['date'] = pd.to_datetime(df.date)
# compare dtypes of the original df with this one
print(df.dtypes)
```
### Apply/Map Usage
Map : Create a derived attribute using map
```
df['user_class'] = df['user_type'].map(expand_user_type)
display(df.tail())
```
Apply: Using apply to get attribute ranges
```
display(df.select_dtypes(include=[np.number]).apply(lambda x:
x.max()- x.min()))
```
Applymap: Extract week from date
```
df['purchase_week'] = df[['date']].applymap(lambda dt:dt.week
if not pd.isnull(dt.week)
else 0)
display(df.head())
```
### Missing Values
Drop Rows with missing dates
```
df_dropped = df.dropna(subset=['date'])
display(df_dropped.head())
```
Fill Missing Price values with mean price
```
df_dropped['price'].fillna(value=np.round(df.price.mean(),decimals=2),
inplace=True)
```
Fill Missing user_type values with value from previous row (forward fill)
```
df_dropped['user_type'].fillna(method='ffill',inplace=True)
```
Fill Missing user_type values with value from next row (backward fill)
```
df_dropped['user_type'].fillna(method='bfill',inplace=True)
```
### Duplicates
Drop Duplicate serial_no rows
```
# sample duplicates
display(df_dropped[df_dropped.duplicated(subset=['serial_no'])].head())
print("Shape of df={}".format(df_dropped.shape))
df_dropped.drop_duplicates(subset=['serial_no'],inplace=True)
# updated dataframe
display(df_dropped.head())
print("Shape of df={}".format(df_dropped.shape))
```
Remove rows which have less than 3 attributes with non-missing data
```
display(df.dropna(thresh=3).head())
print("Shape of df={}".format(df.dropna(thresh=3).shape))
```
### Encode Categoricals
One Hot Encoding using get_dummies()
```
display(pd.get_dummies(df,columns=['user_type']).head())
```
Label Mapping
```
type_map={'a':0,'b':1,'c':2,'d':3,np.NAN:-1}
df['encoded_user_type'] = df.user_type.map(type_map)
display((df.tail()))
```
### Random Sampling data from DataFrame
```
display(df.sample(frac=0.2, replace=True, random_state=42).head())
```
### Normalizing Numeric Values
Normalize price values using **Min-Max Scaler**
```
df_normalized = df.dropna().copy()
min_max_scaler = preprocessing.MinMaxScaler()
np_scaled = min_max_scaler.fit_transform(df_normalized['price'].values.reshape(-1,1))
df_normalized['price'] = np_scaled.reshape(-1,1)
display(df_normalized.head())
```
Normalize quantity purchased values using **Robust Scaler**
```
df_normalized = df.dropna().copy()
robust_scaler = preprocessing.RobustScaler()
rs_scaled = robust_scaler.fit_transform(df_normalized['quantity_purchased'].values.reshape(-1,1))
df_normalized['quantity_purchased'] = rs_scaled.reshape(-1,1)
display(df_normalized.head())
```
### Data Summarization
Condition based aggregation
```
print("Mean price of items purchased by user_type=a :: {}".format(df['price'][df['user_type']=='a'].mean()))
```
Condtion based counts
```
print(df['purchase_week'].value_counts())
```
### Group By
Group By certain attributes
```
print(df.groupby(['user_class'])['quantity_purchased'].sum())
```
Group By with different aggregate functions
```
display(df.groupby(['user_class'])['quantity_purchased'].agg([np.sum,
np.mean,
np.count_nonzero]))
```
Group by specific aggregate functions for each attribute
```
display(df.groupby(['user_class','user_type']).agg({'price':np.mean,
'quantity_purchased':np.max}))
```
Group by with multiple agg for each attribute
```
display(df.groupby(['user_class','user_type']).agg({'price':{
'total_price':np.sum,
'mean_price':np.mean,
'variance_price':np.std,
'count':np.count_nonzero},
'quantity_purchased':np.sum}))
```
### Pivot Tables
```
display(df.pivot_table(index='date', columns='user_type',
values='price',aggfunc=np.mean))
```
### Stack a Dataframe
```
print(df.stack())
```
| github_jupyter |
```
import itertools
import string
import collections
import random
F = range(0,7)
# letters = ["".join(x) for x in itertools.combinations(string.letters, 2)]
# letters = ["".join(x) for x in itertools.combinations(string.letters, 2)]
letters = [('\U{0:08x}'.format(x)).decode('unicode-escape') for x in itertools.chain(range(0x1f600,0x1F623), range(0x1F624, 0x1f650), range(0x250, 0x2b0), range(0x2c80, 0x2cf4), range(0x2460, 0x2500))]
for l in letters:
print l,
print "..."
def reciprocal(x):
return reciprocal.table[x]
reciprocal.table = { i : j for i,j in itertools.product(F,F) if (i * j) % len(F) == 1 }
print reciprocal.table
if (len(reciprocal.table) < (len(F) - 1)):
print "There are elements with no reciprocal..."
def multiply(k, x):
return tuple([k * i % len(F) for i in x])
def lmod(x):
return multiply(1, x)
def add(x, y):
return tuple([(i + j) % len(F) for i,j in zip(x,y)])
def get_representative_in_P2(x):
x = lmod(x)
if (max(x) == 0):
return tuple([0,0,0])
if x[2] != 0:
k = reciprocal(x[2])
elif x[1] != 0:
k = reciprocal(x[1])
else:
k = reciprocal(x[0])
return multiply(k, x)
print get_representative_in_P2([1,2,3])
print get_representative_in_P2([1,2,0])
print get_representative_in_P2([1,0,0])
def span(x, y):
points = set()
for m, n in itertools.product(F,F):
if m == n == 0:
continue
p = get_representative_in_P2(add(multiply(m, x), multiply(n, y)))
points.add(p)
return frozenset(points)
def validate(L):
expected_size = len(F) ** 2 + len(F) + 1
if len(L) != expected_size:
print "Unexpected length of L: expected: {}, actual {}".format(expected_size, len(L))
return False
for x in L:
expected_size = len(F) + 1
if len(x) != expected_size:
print "Unexpected length of {}: expected: {}, actual {}".format(x, expected_size, len(x))
return False
for x, y in itertools.product(L, L):
if x == y:
continue
if len(x & y) != 1:
print "Unexpected intersection of {} and {}: expected: {}, actual {}".format(x, y, 1, len(x & y))
return False
return True
def get_P2_points():
points = set()
l2p = {}
p2l = {}
i = 0
for x,y,z in itertools.product(F,F,F):
if x == y == z == 0:
continue
p = get_representative_in_P2((x,y,z))
if p not in points:
points.add(p)
l2p[letters[i]] = p
p2l[p] = letters[i]
i += 1
return points, l2p, p2l
P2, l2p, p2l = get_P2_points()
print len(P2)
print len(F) ** 2 + len(F) + 1
for k, v in l2p.iteritems():
print k, '-->', v
s = span([1, 2, 1], [0, 1, 2])
# for p in s:
# print p2l[p], "-->", p
def get_P2_lines():
lines = set()
P2, l2p, p2l = get_P2_points()
for x, y in itertools.combinations_with_replacement(P2, 2):
s = span(x, y)
# linear dependent
if (0,0,0) in s:
continue
lines.add(s)
return lines
L = get_P2_lines()
print "Number of lines {}".format(len(L))
print "Validating each two lines have 1 point in common: {}".format(validate(L))
print "The lines in P2(F):\n"
L_as_letters = [" ".join(([p2l[p] for p in line])) for line in L]
for i, line in enumerate(L_as_letters):
s = u"{})\t {}\n".format(i, line)
print s
print random.choice(L_as_letters)
print random.choice(L_as_letters)
```
| github_jupyter |

```
Spam Filtering using SVM
```
# Load Dataset
**Check the link for the dataset from Kaggle**
https://www.kaggle.com/uciml/sms-spam-collection-dataset
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
data = pd.read_csv('spamham.csv')
data.head()
data.shape
print("We have",data.shape[0],"observations")
dis = data['Category'].value_counts()
print("We have",dis[0], 'normal mails')
print("We have",dis[1], 'spam mails')
```
This is our distribution in the total data
```
plt.pie(x = dis.values, explode = (0.1,0), labels = dis.index, autopct='%1.1f%%')
plt.show()
```
# Label Spam as 1 and Ham as 0
### Using loc
```
data.loc[data['Category'] == 'spam']
data.loc[data['Category'] == 'spam']['Category']
data.loc[data['Category'] == 'spam', 'Category'] = 1
data.loc[data['Category'] == 'ham', 'Category'] = 0
data.head()
```
### Use get_dummies
```
pd.get_dummies(data['Category'])
```
Select whether you need the ham as 1 or spam as 1 and insert that column into the data and drop the categorical category already present in the dataframe
# Build the classic X and y
## X -> Feature y -> Label
```
X = data['Message']
y = data['Category']
print(X,'\n\n',y)
```
# Train - Test Split
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size = 0.8)
print("Entire data :", data.shape)
print("X_train shape : ", X_train.shape)
print("y_train shape : ", y_train.shape)
print("X_test shape : ", X_test.shape)
print("y_test shape : ", y_test.shape)
1115/5572
```
0.2 % is maintained as we have given
# Feature Extraction
```
from sklearn.feature_extraction.text import TfidfVectorizer
```
## Why use Tfidf
Classic approach is to use a bad of words
Why we chose Tfidf
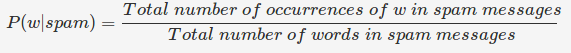

For example, there are two messages in the dataset.
‘hello world’ and ‘hello foo bar’.
1.TF(‘hello’) is 2.
Since hello appears once in the message and has hello in both of the message 2/1
2.IDF(‘hello’) is log(2/2).
If a word occurs a lot, it means that the word gives less information.
say irrelevant words like mail recieves a high impact so that it will be a spam, as a result of using tfidf we will have rare things like offer have high impact
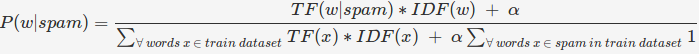
min_df = When building the vocabulary ignore terms that have a document
frequency strictly lower than the given threshold. This value is also
called cut-off in the literature.
Stop words are words like a,an,are,etc.
```
extractor = TfidfVectorizer(min_df = 1, stop_words = 'english', lowercase=True )
X_train_features = extractor.fit_transform(X_train)
X_test_features = extractor.transform(X_test)
# Convert object type to int
y_train = y_train.astype(int)
y_test = y_test.astype(int)
```
# Build the model
**SVM model**

SVM builds a classifier by searching for a separating hyperplane (optimal hyperplane) which is optimal and maximises the margin that separates the categories (in our case spam and ham). Thus, SVM has the advantage of robustness in general and effectiveness when the number of dimensions is greater than the number of samples.
```
from sklearn.svm import LinearSVC
model = LinearSVC()
model.fit(X_train_features, y_train)
```
# Evaluate the model
```
from sklearn.metrics import accuracy_score
train_pred = model.predict(X_train_features)
print("Training accuracy : ",accuracy_score(y_train, train_pred))
test_pred = model.predict(X_test_features)
print("Test accuracy : ",accuracy_score(y_test, test_pred))
```
# Checking
```
mail = ["WINNER!! As a valued network customer you have been selected to receivea £900 prize reward! To claim call 09061701461. Claim code KL341. Valid 12 hours only."]
processed = extractor.transform(mail)
model.predict(processed)
```
Your model may not give best output because it hasn't seen a lot of data.
You can implement this model using a large dataset and you will be able to build a better model
# Success
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
```
Add some physical examples to motivate root finding. Just using algebraic examples up to this point.
#Solving Algebraic Equations: Root-finding Methods
Solving a linear equation (e.g. $y=mx+b$) is one of the first topic one encounters in any sort of introductory algebra course. This of course can be done by hand quite easily. One of course then moves onto more sophisticated equations involving terms like $x^2,x^3,\ldots$. For example, the equation $$x^2+10x+9=0$$ can be solved using the familiar quadratic formula $x=\frac{-b\pm\sqrt{b^2-4ac}}{2a}$ where $a=1$, $b=10$, and $c=9$. Applying the quadratic formula, we see that $x=-1,-9$. When we plot the function $f(x)=x^2+10x+9$, we can see that $f=0$ at $x=-1$ and $x=-9$.
```
x=np.linspace(-15,5,1000)
x_zeros=[-1,-9]
fig = plt.figure(figsize=(10,10))
ax = fig.gca()
ax.plot(x,0*x,'--k',linewidth=2.0)
ax.plot(x,x**2+10*x+9,linewidth=2.0)
ax.plot(x_zeros,[0,0],'ro',markersize=10)
ax.set_xlabel(r'$x$',fontsize=22)
ax.set_ylabel(r'$f(x)$',fontsize=22)
ax.set_title(r'$f(x)=x^2+10x+9$',fontsize=22)
ax.set_ylim([-20,20])
plt.show()
```
These types of equations, often referred to as polynomials, can also be solved through factorization. For example, the equation $$x^3+6x^2+11x+6=0$$ can be rewritten as $$(x+1)(x+2)(x+3)=0$$ such that $x=-1,-2,-3$ are all roots of the equation. Again, let's plot the function and the roots to check our answer.
```
x=np.linspace(-4,0,1000)
x_zeros=[-1,-2,-3]
fig = plt.figure(figsize=(10,10))
ax = fig.gca()
ax.plot(x,0*x,'--k',linewidth=2.0)
ax.plot(x,x**3+6*x**2+11*x+6,linewidth=2.0)
ax.plot(x_zeros,[0,0,0],'ro',markersize=10)
ax.set_xlabel(r'$x$',fontsize=22)
ax.set_ylabel(r'$f(x)$',fontsize=22)
ax.set_title(r'$f(x)=x^3+6x^2+11x+6$',fontsize=22)
ax.set_ylim([-2,2])
plt.show()
```
Note also that finding the value of $x$ where $f(x)=0$ is the just as easy as finding the value of $x$ where $f(x)=a$, where $a$,a constant, is some value of $f$. This is because $f(x)=a$ can be rewritten as $f(x)-a=0$. Thus, we can rewrite $f(x)=a$ as a root-finding problem $g(x)=0$ where $g(x)=f(x)-a$.
So far, we've only considered the case of finding roots of polynomials. In these cases, we could always find the roots using either the quadratic formula (when our leading term was $x^2$) or a factorization method when possible. But what if our function is more complicated, say $f(x)=2x^2\sin(x)+\ln(x)$, or better yet, what if our function is unknown altogether? Often, we resort to **root finding algorithms** where we make a first guess about the root of our function and then repeatedly refine our guess until we have a reasonably accurate estimate of the root.
##Bisection Method
One of the most popular root-finding methods is known as the **bisection method**. In order to apply the bisection method, there are two important requirements:
1. the function $f$ must be _monotonically increasing (or decreasing)_ between $x=a$ and $x=b$,
2. between $f(x=a)$ and $f(x=b)$, $f(x)=0$ for some value of $x=x_0$ such that $a<x_0<b$.
Let's consider what both of these conditions actually mean. Consider a function $f$ that is _monotonically increasing (or decreasing)_ from $x=a$ to $x=b$. This is just a fancy way of saying that as we move from $a$ to $b$, $f$ **only** increases (or **only** decreases). For example, the above function ($f(x)=x^3+6x^2+11x+6$) is neither monotonically increasing or decreasing on the interval shown, $x\in[-4,0]$. However, from $x=-4$ to $x=-2-\sqrt{3}/3$, the function increases, from $x=-2-\sqrt{3}/3$ to $x=-2+\sqrt{3}/3$ the function decreases, and then from $x=-2+\sqrt{3}/3$ to $x=0$, the function increases. Thus, when applying our bisection method, we must use it on each of these intervals individually. Lets consider the middle interval, $x\in[-2-\sqrt{3}/3,-2+\sqrt{3}/3]$.
```
xzoom = np.linspace(-2.0-np.sqrt(3.0)/3.0,-2.0+np.sqrt(3.0)/3.0,1000)
def cubic_func(x):
return x**3+6*x**2+11*x+6
fig = plt.figure(figsize=(10,10))
ax = fig.gca()
ax.plot(xzoom,0*xzoom,'--k',linewidth=2.0)
ax.plot(xzoom,cubic_func(xzoom),linewidth=2.0)
ax.set_xlabel(r'$x$',fontsize=22)
ax.set_ylabel(r'$f(x)$',fontsize=22)
ax.set_title(r'$f(x)=x^3+6x^2+11x+6$',fontsize=22)
ax.set_ylim([-.5,.5])
plt.show()
```
Let's call $a=-2.3$ and $b=-1.5$. Looking at the above figure, we can see that between $f(a)$ and $f(b)$, $f=0$ (i.e. the blue curve intersects the black dotted line). Thus, we have also satisfied the second condition for applying the bisection method to find the roots of our function. Now that we've shown that conditions for applying the bisection method are satisfied, how exactly does the method work?
1. Input interval endpoints $a,b$ for a function $f$ which satisfies the above conditions.
2. Calculate first guess of root $c=a+(b-a)/2$ and evaluate $f(c)$.
3. Calculate $f(a)\times f(c)$,
* If $f(a)\times f(c)>0$ (i.e. $c$ is on the same side of the root as $a$), set $a=c$ and update $c$.
* If $f(a)\times f(c)<0$ (i.e. $c$ is on the same side of the root as $b$), set $b=c$ and update $c$.
4. Evalute $f(c)$.
5. Repeat steps 3 and 4 for a finite number of steps or until $f(c)<\epsilon$, where $\epsilon$ is some specified error.
Now let's write a function that performs the bisection method which accepts the bounds on our interval $a,b$ and our function $f$. Note that we can write our bisection method function in such a way that it is not specific to the kind of function to which we apply it. This will allow us to reuse this piece of code for several different functions.
```
def bisection(a,b,f,options):
"""Bisection method for interval (a,b) as applied to a monotonically increasing (or decreasing) function f on (a,b)"""
#First guess for root value
c = a + (b - a)/2
#Evaluate function at first guess
f_guess = f(c)
#Initialize the counter
count = 0
#Initialize lists of endpoints and roots
a_list = [a]
b_list = [b]
c_list = [c]
#Begin iteration
while(count<options['max_count'] and np.fabs(f_guess)>options['tol']):
#Output iteration and endpoints
print "Iteration %d, Endpoints (%f,%f)"%(count,a,b)
#Check relative sign
if f_guess*f(a)>0:
#advance left endpoint
a=c
elif f_guess*f(a)<0:
#advance right endpoint
b=c
else:
#case where f(c)==0
print "f(c) == 0 exactly. Breaking out of loop."
break
#update root value guess
c = a + (b-a)/2
#evaluate function at current guess
f_guess = f(c)
#save a,b,c values
a_list.append(a)
b_list.append(b)
c_list.append(c)
#update the counter
count = count + 1
#print final zero approximation
print "Value of the function at approximate zero is f(c=%f)=%f"%(c,f_guess)
#return lists of a,b,c
return a_list,b_list,c_list
```
Now that we have an algorithm for our bisection method, let's apply it to the above function for our chosen endpoints $a=-2.3$ and $b=-1.5$.
```
opt = {'max_count':100,'tol':1.0e-10}
alist,blist,clist = bisection(-2.5,-1.9,cubic_func,opt)
```
Thus, we see that our bisection method performs very well for our cubic function on the given interval. This provides a nice check of our method as we can analytically verify (by hand) that our root on this interval is $x=-2$. Now let's apply this method over the two remaining intervals to find our other zeros. For the leftmost interval, we'll choose $a=-5,b=-2-\sqrt{3}/3$ and for the rightmost interval, we'll choose $a=-2+\sqrt{3}/3,b=4$.
```
opt['tol'] = 1.0e-6
aleft,bleft,cleft=bisection(-5.0,-2.0-np.sqrt(3.0)/3.0,cubic_func,opt)
aright,bright,cright=bisection(-2.0+np.sqrt(3.0)/3.0,4.0,cubic_func,opt)
```
Our bisection method is able to calculate the zeros of our cubic function in a reasonable number of iterations. Now that we've shown that our bisection algorithm works, let's apply it to a more useful case: an unknown function where we can't use a simple method like the quadratic formula or factorization to find the zeros.
```
%run make_unknown_function
unknowns = np.loadtxt('example_data/root_finding_unknown_function.txt')
x_unknown = unknowns[0,:]
y_unknown = unknowns[1,:]
```
Now let's plot our unknown function to determine where it is monotonically increasing/decreasing. We'll make smart choices about our intervals by "bracketing" our function at $(0.5,2.9)$ and $(3.5,6)$. Note that we are free to make any choice when it comes to our intervals provided that on $(a,b)$, $f$ is either monotonically increasing or decreasing.
```
a1 = 0.5
b1 = 2.9
a2 = 3.5
b2 = 6.0
fig = plt.figure(figsize=(10,10))
ax = fig.gca()
ax.plot(x_unknown,y_unknown,linewidth=2.0)
ax.plot(x_unknown,0*x_unknown,'--k',linewidth=2.0)
ax.plot([a1,a2],[y_unknown[np.where(x_unknown<a1)[0][-1]],y_unknown[np.where(x_unknown<a2)[0][-1]]],'ko',marker=r'$[$',markersize=25)
ax.plot([b1,b2],[y_unknown[np.where(x_unknown<b1)[0][-1]],y_unknown[np.where(x_unknown<b2)[0][-1]]],'ko',marker=r'$]$',markersize=25)
ax.set_xlabel(r'$x$',fontsize=22)
ax.set_ylabel(r'$f(x)$',fontsize=22)
ax.set_title(r'Unknown Function',fontsize=22)
ax.set_xlim([x_unknown[0],x_unknown[-1]])
plt.show()
```
Now, we can apply our bisection method to this new unknown function to try to find the roots of this equation. Note that here we cannot use our quadratic method or factorization; the form of the function has been defined when we ran the script `make_unknown_function.py`. It is only important to know the form of the function so that we can feed it to the bisection method; suffice it to say that is complicated.
```
a1list,b1list,c1list = bisection(a1,b1,sine_integral_func,opt)
a2list,b2list,c2list = bisection(a2,b2,sine_integral_func,opt)
```
Now, let's plot the zeros on top of our plot of the unknown function to get a sense of how good our guess is.
```
fig = plt.figure(figsize=(10,10))
ax = fig.gca()
ax.plot(x_unknown,y_unknown,linewidth=2.0)
ax.plot(x_unknown,0*x_unknown,'--k',linewidth=2.0)
ax.plot([a1,a2],[y_unknown[np.where(x_unknown<a1)[0][-1]],y_unknown[np.where(x_unknown<a2)[0][-1]]],'ko',marker=r'$[$',markersize=25)
ax.plot([b1,b2],[y_unknown[np.where(x_unknown<b1)[0][-1]],y_unknown[np.where(x_unknown<b2)[0][-1]]],'ko',marker=r'$]$',markersize=25)
ax.plot([c1list[-1],c2list[-1]],[sine_integral_func(c1list[-1]),sine_integral_func(c2list[-1])],'ro',markersize=10)
ax.set_xlabel(r'$x$',fontsize=22)
ax.set_ylabel(r'$f(x)$',fontsize=22)
ax.set_title(r'Unknown Function',fontsize=22)
ax.set_xlim([x_unknown[0],x_unknown[-1]])
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/zerotodeeplearning/ztdl-masterclasses/blob/master/notebooks/Convolutional_Neural_Networks.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## Learn with us: www.zerotodeeplearning.com
Copyright © 2021: Zero to Deep Learning ® Catalit LLC.
```
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Convolutional Neural Networks
This notebook is best run using a GPU backend
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
from tensorflow.keras.datasets import fashion_mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout, Activation
from tensorflow.keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Flatten
(X_train, y_train), (X_test, y_test) = fashion_mnist.load_data()
X_train = X_train.reshape(-1, 28, 28, 1).astype('float32') / 255.0
X_test = X_test.reshape(-1, 28, 28, 1).astype('float32') / 255.0
X_train.shape
model = Sequential([
Conv2D(32, (3, 3), input_shape=(28, 28, 1), activation='relu'),
Flatten(),
Dense(10, activation='softmax')
])
model.compile(loss='sparse_categorical_crossentropy',
optimizer='rmsprop',
metrics=['accuracy'])
h = model.fit(X_train, y_train, batch_size=128, epochs=3, validation_split=0.1)
pd.DataFrame(h.history).plot()
```
### Exercise 1: A better CNN
The CNN we just implemented is very basic. Improve its design by adding layers before and after the `Flatten` layer. It is up to you what the model will be. Here are some things you may consider:
- how many convolutional layers to use?
- what filter size?
- how many filters in each layer?
- what activation function?
- pooling? what type?
- how many fully-connected layers after Flatten?
- dropout or batch normalization?
- what batch size for training?
Once you've defined your new model, compile it and train it on the straining data for 5 epochs. Can you get the accuracy above 90% ?
Your code should look like:
```python
model = Sequential([
# YOUR CODE HERE
# ...
])
model.compile(# YOUR CODE HERE)
model.fit(# YOUR CODE HERE)
```
### Exercise 2: Modularize the network
You may have noticed that, as a network becomes deeper, some parts can be naturally grouped to be considered small sub-networks or modules within the larger architecture. Can we leverage this fact and build a network made of modules?
Define 2 functions to achieve that.
#### Function 1
The first function should return a block with the following components:
- Conv2D with variable number of filters and `'same'` padding.
- Conv2D with variable number of filters
- Optional BatchNormalization
- MaxPooling2D
- Dropout
Use the functional API to do this, and write a function with the following signature:
```python
def cnn_block(inputs, n_filters, activation, dropout, bn):
x = ....(inputs)
....
return x
```
#### Function 2
The second function should return a compiled model, it should use the first function to include CNN blocks and it should have the following signature:
```python
def convolutional_model(n_blocks=2):
inputs = Input(shape=(28, 28, 1))
x = inputs
for i in range(n_blocks):
x = cnn_block(x, 32, 'relu', 0.4, True)
# YOUR CODE HERE
# ..
outputs = Dense(10)(x)
model = Model(# YOUR CODE HERE)
model.compile(
# YOUR CODE HERE
)
return model
```
Test your functions by creating a model and training it for 1 epoch
```
from tensorflow.keras.layers import Input, BatchNormalization
from tensorflow.keras.models import Model
from tensorflow.keras.losses import SparseCategoricalCrossentropy
from tensorflow.keras.optimizers import SGD, Adam, RMSprop, Adamax
```
### Exercise 3: Hyperparameter search with Tensorboard
Extend your search for the best model using what you've learned in the Hyperparameter tuning class.
Take the `convolutional_model` function from the previous exercise and modify it to accept hyperparameters.
You could define a dictionary to hold your ranges:
```python
hp_ranges = {
'n_blocks': # YOUR CODE HERE
'n_conv_filters_1': # YOUR CODE HERE
# YOUR CODE HERE
# ...
}
```
and then change the signature of your function to look like this:
```python
def convolutional_model(hparams):
inputs = Input(shape=(28, 28, 1))
x = inputs
for i in range(hparams['n_blocks']):
x = cnn_block(x,
hparams[f'n_conv_filters_{i+1}'],
...
# YOUR CODE HERE
# ...
```
Finally, sample the parameter space and see if you can identify important and unimportant parameters for this particular dataset. The `train_test_model` and search loop are provided for you.
```
from tensorflow.keras.callbacks import TensorBoard
from tensorboard.plugins.hparams import api as hp
from sklearn.model_selection import ParameterGrid, ParameterSampler
from scipy.stats.distributions import expon, uniform, randint
logdir = 'logs/hparam_tuning/'
# YOUR CODE HERE
def train_test_model(hparams, run):
model = convolutional_model(hparams)
model.fit(X_train, y_train, epochs=5,
verbose=2,
batch_size=hparams['batch_size'],
validation_split=0.1,
callbacks=[TensorBoard(logdir + run_name)]
)
model.fit(X_train, y_train, epochs=1,
verbose=2,
batch_size=hparams['batch_size'],
validation_split=0.1,
callbacks=[hp.KerasCallback(logdir+run, hparams)]
)
session_num = 0
for hparams in ParameterSampler(hp_ranges, n_iter=50, random_state=0):
print(hparams)
run_name = "run-%d" % session_num
print('--- Starting trial: %s' % run_name)
train_test_model(hparams, run_name)
session_num += 1
%load_ext tensorboard
%tensorboard --logdir logs
```
| github_jupyter |
### Dataset Reading
```
import pandas as pd
data = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Train.xlsx')
pd.set_option('display.max_colwidth',150)
data.head()
data.shape
print(data.dtypes)
```
### Making of "label" Variable
```
label = data['label']
label.head()
```
### Checking Dataset Balancing
```
print(label.value_counts())
import matplotlib.pyplot as plt
label.value_counts().plot(kind='bar', color='blue')
```
### Convering label into "0" or "1"
```
import numpy as np
classes_list = ["fake","real"]
label_index = data['label'].apply(classes_list.index)
final_label = np.asarray(label_index)
print(final_label[:10])
from keras.utils.np_utils import to_categorical
label_twoDimension = to_categorical(final_label, num_classes=2)
print(label_twoDimension[:10])
```
### Making of "text" Variable
```
text = data['tweet']
text.head(10)
```
### Dataset Pre-processing
```
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
clean_text = text.apply(lambda x:text_clean(x))
clean_text.head(10)
```
### Removing stopwords
```
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
clean_text_ns=clean_text.apply(lambda x: stop_words_removal(x))
print(clean_text_ns.head(10))
```
### Lemmatization
```
"""# Lemmatization
import nltk
nltk.download('wordnet')
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
def word_lemmatizer(text):
lem_text = "".join([lemmatizer.lemmatize(i) for i in text])
return lem_text"""
"""clean_text_lem = clean_text_ns.apply(lambda x : word_lemmatizer(x))"""
"""print(clean_text_lem.head(10))"""
```
### Stemming
```
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
clean_text_stem = clean_text_ns.apply(lambda x : word_stemmer(x))
print(clean_text_stem.head(10))
# final_text = [x for x in clean_text_lem if len(x) > 3]
#print(final_text)
```
### Tokenization using "keras"
```
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_all = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_all.fit_on_texts(clean_text_stem)
```
### Making Vocab for words
```
vocabulary_all = len(tok_all.word_counts)
print(vocabulary_all)
l = tok_all.word_index
print(l)
```
### encoding or sequencing
```
encoded_clean_text_stem = tok_all.texts_to_sequences(clean_text_stem)
print(clean_text_stem[1])
print(encoded_clean_text_stem[1])
```
### Pre-padding
```
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_stem = sequence.pad_sequences(encoded_clean_text_stem, maxlen=max_length, padding='pre')
```
# Test Data Pre-processing
# Data test Reading
```
data_t = pd.read_excel('drive/My Drive/Constraint_Competition_Dataset/Constraint_Covid-19_English_Val.xlsx')
pd.set_option('display.max_colwidth',150)
data_t.head()
data_t.shape
print(data_t.dtypes)
```
# Making of "label" Variable
```
label_t = data_t['label']
label_t.head()
```
# Checking Dataset Balancing
```
print(label_t.value_counts())
import matplotlib.pyplot as plt
label_t.value_counts().plot(kind='bar', color='red')
```
# Convering label into "0" or "1"
```
import numpy as np
classes_list_t = ["fake","real"]
label_t_index = data_t['label'].apply(classes_list_t.index)
final_label_t = np.asarray(label_t_index)
print(final_label_t[:10])
from keras.utils.np_utils import to_categorical
label_twoDimension_t = to_categorical(final_label_t, num_classes=2)
print(label_twoDimension_t[:10])
```
# Making of "text" Variable
```
text_t = data_t['tweet']
text_t.head(10)
```
# **Dataset Pre-processing**
1. Remove unwanted words
2. Stopwords removal
3. Stemming
4. Tokenization
5. Encoding or Sequencing
6. Pre-padding
### 1. Removing Unwanted Words
```
import re
def text_clean(text):
''' Pre process and convert texts to a list of words '''
text=text.lower()
# Clean the text
text = re.sub(r"[^A-Za-z0-9^,!.\/'+-=]", " ", text)
text = re.sub(r"what's", "what is ", text)
text = re.sub(r"I'm", "I am ", text)
text = re.sub(r"\'s", " ", text)
text = re.sub(r"\'ve", " have ", text)
text = re.sub(r"can't", "cannot ", text)
text = re.sub(r"wouldn't", "would not ", text)
text = re.sub(r"shouldn't", "should not ", text)
text = re.sub(r"shouldn", "should not ", text)
text = re.sub(r"didn", "did not ", text)
text = re.sub(r"n't", " not ", text)
text = re.sub(r"i'm", "i am ", text)
text = re.sub(r"\'re", " are ", text)
text = re.sub(r"\'d", " would ", text)
text = re.sub(r"\'ll", " will ", text)
text = re.sub('https?://\S+|www\.\S+', "", text)
text = re.sub(r",", " ", text)
text = re.sub(r"\.", " ", text)
text = re.sub(r"!", " ! ", text)
text = re.sub(r"\/", " ", text)
text = re.sub(r"\^", " ^ ", text)
text = re.sub(r"\+", " + ", text)
text = re.sub(r"\-", " - ", text)
text = re.sub(r"\=", " = ", text)
text = re.sub(r"'", " ", text)
text = re.sub(r"(\d+)(k)", r"\g<1>000", text)
text = re.sub(r":", " : ", text)
text = re.sub(r" e g ", " eg ", text)
text = re.sub(r" b g ", " bg ", text)
text = re.sub(r" u s ", " american ", text)
text = re.sub(r"\0s", "0", text)
text = re.sub(r" 9 11 ", "911", text)
text = re.sub(r"e - mail", "email", text)
text = re.sub(r"j k", "jk", text)
text = re.sub(r"\s{2,}", " ", text)
text = re.sub(r"[0-9]", "", text)
# text = re.sub(r"rt", " ", text)
return text
clean_text_t = text_t.apply(lambda x:text_clean(x))
clean_text_t.head(10)
```
### 2. Removing Stopwords
```
import nltk
from nltk.corpus import stopwords
nltk.download('stopwords')
def stop_words_removal(text1):
text1=[w for w in text1.split(" ") if w not in stopwords.words('english')]
return " ".join(text1)
clean_text_t_ns=clean_text_t.apply(lambda x: stop_words_removal(x))
print(clean_text_t_ns.head(10))
```
### 3. Stemming
```
# Stemming
from nltk.stem import PorterStemmer
stemmer = PorterStemmer()
def word_stemmer(text):
stem_text = "".join([stemmer.stem(i) for i in text])
return stem_text
clean_text_t_stem = clean_text_t_ns.apply(lambda x : word_stemmer(x))
print(clean_text_t_stem.head(10))
```
### 4. Tokenization
```
import keras
import tensorflow
from keras.preprocessing.text import Tokenizer
tok_test = Tokenizer(filters='!"#$%&\'()*+,-./:;<=>?@[\\]^_`{|}~', lower=True, char_level = False)
tok_test.fit_on_texts(clean_text_t_stem)
vocabulary_all_test = len(tok_test.word_counts)
print(vocabulary_all_test)
test_list = tok_test.word_index
print(test_list)
```
### 5. Encoding or Sequencing
```
encoded_clean_text_t_stem = tok_all.texts_to_sequences(clean_text_t_stem)
print(clean_text_t_stem[0])
print(encoded_clean_text_t_stem[0])
```
### 6. Pre-padding
```
from keras.preprocessing import sequence
max_length = 100
padded_clean_text_t_stem = sequence.pad_sequences(encoded_clean_text_t_stem, maxlen=max_length, padding='pre')
```
# fastText Embedding
```
# fastText Embedding link - https://fasttext.cc/docs/en/crawl-vectors.html
'''import os
import numpy as np
embeddings_index = {}
f = open('drive/My Drive/ML Internship IIIT Dharwad/Copy of cc.en.300.vec',encoding='utf-8',errors='ignore')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Loaded %s word vectors.' % len(embeddings_index))'''
'''embedding_matrix = np.zeros((vocabulary_all+1, 300))
for word, i in tok_all.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector'''
```
## **Word2Vec Embedding**
```
import gensim
from gensim import corpora
from gensim.models import Word2Vec
sen = []
for line in clean_text:
words = line.split()
sen.append(words)
allwords = []
for l in sen:
allwords += l
print (len(allwords))
print (len(set(allwords)))
# model = Word2Vec(sen, size=300,window=5,min_count=5, negative=20)
model = Word2Vec(sen, size=300, min_count=1)
len(model.wv[sen[0][0]])
filename = 'embedding_word2vec.txt'
model.wv.save_word2vec_format(filename,binary=False)
import os
embeddings_index = {}
f = open('embedding_word2vec.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
embeddings_index[word] = coefs
f.close()
print('Loaded %s word vectors.' % len(embeddings_index))
embedding_matrix = np.zeros((vocabulary_all+1, 300))
for word, i in tok_all.word_index.items():
embedding_vector = embeddings_index.get(word)
if embedding_vector is not None:
embedding_matrix[i] = embedding_vector
sim = model.wv.most_similar(positive=[sen[0][11]])
print (sen[0][11])
```
# **CNN Model**
```
from keras.preprocessing import sequence
from keras.preprocessing import text
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation
from keras.layers import Embedding, LSTM
from keras.layers import Conv1D, Flatten
from keras.preprocessing import text
from keras.models import Sequential,Model
from keras.layers import Dense ,Activation,MaxPool1D,Conv1D,Flatten,Dropout,Activation,Dropout,Input,Lambda,concatenate
from keras.utils import np_utils
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
from nltk.stem.porter import PorterStemmer
import nltk
import csv
import pandas as pd
from keras.preprocessing import text as keras_text, sequence as keras_seq
Embedding_Layer = Embedding(vocabulary_all+1, 300, weights=[embedding_matrix], input_length=max_length, trainable=False)
CNN2_model=Sequential([Embedding_Layer,
Conv1D(128,5,activation="relu",padding='same'),
Dropout(0.2),
MaxPool1D(2),
Conv1D(32,3,activation="relu",padding='same'),
Dropout(0.2),
MaxPool1D(2),
Flatten(),
Dense(64,activation="relu"),
Dense(2,activation="sigmoid")
])
CNN2_model.summary()
from keras.optimizers import Adam
CNN2_model.compile(loss = "binary_crossentropy", optimizer=Adam(lr=0.00003), metrics=["accuracy"])
from keras.utils.vis_utils import plot_model
plot_model(CNN2_model, to_file='CNN2_model.png', show_shapes=True, show_layer_names=True)
from keras.callbacks import EarlyStopping, ReduceLROnPlateau,ModelCheckpoint
earlystopper = EarlyStopping(patience=8, verbose=1)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.9,
patience=2, min_lr=0.00001, verbose=1)
```
### **Model Fitting or Training**
```
hist = CNN2_model.fit(padded_clean_text_stem,label_twoDimension,validation_data=(padded_clean_text_t_stem,label_twoDimension_t),epochs=150,batch_size=32,callbacks=[earlystopper, reduce_lr])
```
# log loss
```
CNN2_model_predictions = CNN2_model.predict(padded_clean_text_t_stem)
from sklearn.metrics import log_loss
log_loss_test= log_loss(label_twoDimension_t,CNN2_model_predictions)
log_loss_test
```
# Classification Report
```
predictions = np.zeros_like(CNN2_model_predictions)
predictions[np.arange(len(CNN2_model_predictions)), CNN2_model_predictions.argmax(1)] = 1
predictionInteger=(np.argmax(predictions, axis=1))
predictionInteger
'''pred_label = np.array(predictionInteger)
df = pd.DataFrame(data=pred_label , columns=["task1"])
print(df)'''
# df.to_csv("submission_EN_A.csv", index=False)
from sklearn.metrics import classification_report
print(classification_report(label_twoDimension_t,predictions))
```
# Epoch v/s Loss Plot
```
from matplotlib import pyplot as plt
plt.plot(hist.history["loss"],color = 'red', label = 'train_loss')
plt.plot(hist.history["val_loss"],color = 'blue', label = 'val_loss')
plt.title('Loss Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.savefig('CNN2_HASOC_Eng_lossPlot.pdf',dpi=1000)
from google.colab import files
files.download('CNN2_HASOC_Eng_lossPlot.pdf')
```
# Epoch v/s Accuracy Plot
```
plt.plot(hist.history["accuracy"],color = 'red', label = 'train_accuracy')
plt.plot(hist.history["val_accuracy"],color = 'blue', label = 'val_accuracy')
plt.title('Accuracy Visualisation')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend()
plt.savefig('CNN2_HASOC_Eng_accuracyPlot.pdf',dpi=1000)
files.download('CNN2_HASOC_Eng_accuracyPlot.pdf')
```
# Area under Curve-ROC
```
pred_train = CNN2_model.predict(padded_clean_text_stem)
pred_test = CNN2_model.predict(padded_clean_text_t_stem)
import numpy as np
import matplotlib.pyplot as plt
from itertools import cycle
from sklearn import svm, datasets
from sklearn.metrics import roc_curve, auc
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import label_binarize
from sklearn.multiclass import OneVsRestClassifier
from scipy import interp
def plot_AUC_ROC(y_true, y_pred):
n_classes = 2 #change this value according to class value
# Compute ROC curve and ROC area for each class
fpr = dict()
tpr = dict()
roc_auc = dict()
for i in range(n_classes):
fpr[i], tpr[i], _ = roc_curve(y_true[:, i], y_pred[:, i])
roc_auc[i] = auc(fpr[i], tpr[i])
# Compute micro-average ROC curve and ROC area
fpr["micro"], tpr["micro"], _ = roc_curve(y_true.ravel(), y_pred.ravel())
roc_auc["micro"] = auc(fpr["micro"], tpr["micro"])
############################################################################################
lw = 2
# Compute macro-average ROC curve and ROC area
# First aggregate all false positive rates
all_fpr = np.unique(np.concatenate([fpr[i] for i in range(n_classes)]))
# Then interpolate all ROC curves at this points
mean_tpr = np.zeros_like(all_fpr)
for i in range(n_classes):
mean_tpr += interp(all_fpr, fpr[i], tpr[i])
# Finally average it and compute AUC
mean_tpr /= n_classes
fpr["macro"] = all_fpr
tpr["macro"] = mean_tpr
roc_auc["macro"] = auc(fpr["macro"], tpr["macro"])
# Plot all ROC curves
plt.figure()
plt.plot(fpr["micro"], tpr["micro"],
label='micro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["micro"]),
color='deeppink', linestyle=':', linewidth=4)
plt.plot(fpr["macro"], tpr["macro"],
label='macro-average ROC curve (area = {0:0.2f})'
''.format(roc_auc["macro"]),
color='navy', linestyle=':', linewidth=4)
colors = cycle(['aqua', 'darkorange'])
#classes_list1 = ["DE","NE","DK"]
classes_list1 = ["Non-duplicate","Duplicate"]
for i, color,c in zip(range(n_classes), colors,classes_list1):
plt.plot(fpr[i], tpr[i], color=color, lw=lw,
label='{0} (AUC = {1:0.2f})'
''.format(c, roc_auc[i]))
plt.plot([0, 1], [0, 1], 'k--', lw=lw)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic curve')
plt.legend(loc="lower right")
#plt.show()
plt.savefig('CNN2_HASOC_Eng_Area_RocPlot.pdf',dpi=1000)
files.download('CNN2_HASOC_Eng_Area_RocPlot.pdf')
plot_AUC_ROC(label_twoDimension_t,pred_test)
```
| github_jupyter |
```
if 0 :
%matplotlib inline
else :
%matplotlib notebook
```
# Import libraries
```
import sys
import os
module_path = os.path.abspath('.') +"\\_scripts"
print(module_path)
if module_path not in sys.path:
sys.path.append(module_path)
from _00_Import_packages_git3 import *
from platform import python_version
print(python_version())
import numpy
print(numpy.__version__)
import scipy
print(scipy.__version__)
import pandas
print(pandas.__version__)
from numpy import array
import pandas as pd
from sos_trades_core.execution_engine.execution_engine import ExecutionEngine
from numpy.testing import assert_array_equal, assert_array_almost_equal # @UnresolvedImport
import os
from gemseo.core.mdo_scenario import MDOScenario
```
# Description of the model
DESC_IN = {'x': {'type': 'array', 'visibility': SoSDiscipline.SHARED_VISIBILITY, 'namespace': 'ns_OptimSellar'},
'y_2': {'type': 'array', 'visibility': SoSDiscipline.SHARED_VISIBILITY, 'namespace': 'ns_OptimSellar'},
'z': {'type': 'array', 'visibility': SoSDiscipline.SHARED_VISIBILITY, 'namespace': 'ns_OptimSellar'}}
DESC_OUT = {'y_1': {'type': 'array',
'visibility': SoSDiscipline.SHARED_VISIBILITY, 'namespace': 'ns_OptimSellar'}}
mod_path = 'SoSTrade.dfmodel.d_dfmodel_sos.DfModel'
ns_test = 'MyStudy'
ns_dict = {'ns_OptimSellar': ns_test}
disc_name = 'Sellar1'
Model :
$$y_1 = z_0 ^ 2 + x + z_1 - 0.2 * y_2$$
# Set the discipline : here Sellar1
```
# Choice of the discipline (File that provides the wrapper and associated wrapper)
mod_path = 'sos_trades_core.sos_wrapping.test_discs.sellar.Sellar1'
#from sos_trades_core.sos_wrapping.test_discs.sellar import Sellar1
from sos_trades_core.execution_engine.execution_engine import ExecutionEngine
ns_test = 'MyStudy'
ee = ExecutionEngine(ns_test)
ns_dict = {'ns_OptimSellar': ns_test} # conform to shared variable x
ee.ns_manager.add_ns_def(ns_dict)
disc_name = 'Sellar1'
disc1_builder = ee.factory.get_builder_from_module(disc_name, mod_path)
ee.factory.set_builders_to_coupling_builder(disc1_builder)
ee.configure()
```
# Display discipline structure
```
my_disc1 = ee.root_process.sos_disciplines[0]
DESC_IN = my_disc1.DESC_IN
DESC_OUT = my_disc1.DESC_OUT
DESC_IN_df = pd.DataFrame.from_dict(DESC_IN,orient='index')
DESC_OUT_df = pd.DataFrame.from_dict(DESC_OUT,orient='index')
DESC_IN_df
DESC_OUT_df
```
# Display study structure
```
ee.display_treeview_nodes()
```
# Provide inputs
```
import numpy as np
priv_in_values = {
'MyStudy.z': np.array([1.,0.]),
'MyStudy.x': 0.,
'MyStudy.y_2': 1.
}
priv_in_values
ee.dm.set_values_from_dict(priv_in_values)
ee.root_process.update_from_dm()
```
# Execute
```
print(ee.execute())
```
# Display Results
```
my_disc1.get_data_io_dict('out')['y_1']['value']
my_disc1.compute_sos_jacobian()
my_disc1.jac
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.