text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
```
# !nvidia-smi
!pip --quiet install transformers
!pip --quiet install tokenizers
from google.colab import drive
drive.mount('/content/drive')
!cp -r '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Scripts/.' .
COLAB_BASE_PATH = '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/'
MODEL_BASE_PATH = COLAB_BASE_PATH + 'Models/Files/141-roBERTa_base/'
import os
os.mkdir(MODEL_BASE_PATH)
```
## Dependencies
```
import json, warnings, shutil
from tweet_utility_scripts import *
from tweet_utility_preprocess_roberta_scripts import *
from transformers import TFRobertaModel, RobertaConfig
from tokenizers import ByteLevelBPETokenizer
from tensorflow.keras.models import Model
from tensorflow.keras import optimizers, metrics, losses, layers
SEED = 0
seed_everything(SEED)
warnings.filterwarnings("ignore")
```
# Load data
```
database_base_path = COLAB_BASE_PATH + 'Data/aux/'
k_fold = pd.read_csv(database_base_path + '5-fold.csv')
display(k_fold.head())
# Unzip files
!tar -xvf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/aux/fold_1.tar.gz'
!tar -xvf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/aux/fold_2.tar.gz'
!tar -xvf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/aux/fold_3.tar.gz'
# !tar -xvf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/aux/fold_4.tar.gz'
# !tar -xvf '/content/drive/My Drive/Colab Notebooks/Tweet Sentiment Extraction/Data/aux/fold_5.tar.gz'
```
# Model parameters
```
vocab_path = COLAB_BASE_PATH + 'qa-transformers/roberta/roberta-base-vocab.json'
merges_path = COLAB_BASE_PATH + 'qa-transformers/roberta/roberta-base-merges.txt'
base_path = COLAB_BASE_PATH + 'qa-transformers/roberta/'
config = {
"MAX_LEN": 96,
"BATCH_SIZE": 32,
"EPOCHS": 5,
"LEARNING_RATE": 3e-5,
"ES_PATIENCE": 1,
"question_size": 4,
"N_FOLDS": 3,
"base_model_path": base_path + 'roberta-base-tf_model.h5',
"config_path": base_path + 'roberta-base-config.json'
}
with open(MODEL_BASE_PATH + 'config.json', 'w') as json_file:
json.dump(json.loads(json.dumps(config)), json_file)
```
# Tokenizer
```
tokenizer = ByteLevelBPETokenizer(vocab_file=vocab_path, merges_file=merges_path,
lowercase=True, add_prefix_space=True)
```
## Learning rate schedule
```
LR_MIN = 1e-6
LR_MAX = config['LEARNING_RATE']
LR_EXP_DECAY = .5
@tf.function
def lrfn(epoch):
lr = LR_MAX * LR_EXP_DECAY**epoch
if lr < LR_MIN:
lr = LR_MIN
return lr
rng = [i for i in range(config['EPOCHS'])]
y = [lrfn(x) for x in rng]
fig, ax = plt.subplots(figsize=(20, 6))
plt.plot(rng, y)
print("Learning rate schedule: {:.3g} to {:.3g} to {:.3g}".format(y[0], max(y), y[-1]))
```
# Model
```
module_config = RobertaConfig.from_pretrained(config['config_path'], output_hidden_states=True)
def model_fn(MAX_LEN):
input_ids = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='input_ids')
attention_mask = layers.Input(shape=(MAX_LEN,), dtype=tf.int32, name='attention_mask')
base_model = TFRobertaModel.from_pretrained(config['base_model_path'], config=module_config, name="base_model")
_, _, hidden_states = base_model({'input_ids': input_ids, 'attention_mask': attention_mask})
h12 = hidden_states[-1]
h11 = hidden_states[-2]
h10 = hidden_states[-3]
h09 = hidden_states[-4]
x_start_09 = layers.Dropout(.1)(h09)
y_start_09 = layers.Dense(1)(x_start_09)
x_start_10 = layers.Dropout(.1)(h10)
y_start_10 = layers.Dense(1)(x_start_10)
x_start_11 = layers.Dropout(.1)(h11)
y_start_11 = layers.Dense(1)(x_start_11)
x_start_12 = layers.Dropout(.1)(h12)
y_start_12 = layers.Dense(1)(x_start_12)
x_start = layers.Average()([y_start_12, y_start_11, y_start_10, y_start_09])
x_start = layers.Flatten()(x_start)
y_start = layers.Activation('softmax', name='y_start')(x_start)
x_end_09 = layers.Dropout(.1)(h09)
y_end_09 = layers.Dense(1)(x_end_09)
x_end_10 = layers.Dropout(.1)(h10)
y_end_10 = layers.Dense(1)(x_end_10)
x_end_11 = layers.Dropout(.1)(h11)
y_end_11 = layers.Dense(1)(x_end_11)
x_end_12 = layers.Dropout(.1)(h12)
y_end_12 = layers.Dense(1)(x_end_12)
x_end = layers.Average()([y_end_12, y_end_11, y_end_10, y_end_09])
x_end = layers.Flatten()(x_end)
y_end = layers.Activation('softmax', name='y_end')(x_end)
model = Model(inputs=[input_ids, attention_mask], outputs=[y_start, y_end])
return model
```
# Train
```
AUTO = tf.data.experimental.AUTOTUNE
strategy = tf.distribute.get_strategy()
history_list = []
for n_fold in range(config['N_FOLDS']):
n_fold +=1
print('\nFOLD: %d' % (n_fold))
# Load data
base_data_path = 'fold_%d/' % (n_fold)
x_train = np.load(base_data_path + 'x_train.npy')
y_train = np.load(base_data_path + 'y_train.npy')
x_valid = np.load(base_data_path + 'x_valid.npy')
y_valid = np.load(base_data_path + 'y_valid.npy')
step_size = x_train.shape[1] // config['BATCH_SIZE']
valid_step_size = x_valid.shape[1] // config['BATCH_SIZE']
# Build TF datasets
train_dist_ds = strategy.experimental_distribute_dataset(get_training_dataset(x_train, y_train, config['BATCH_SIZE'], AUTO, seed=SEED))
valid_dist_ds = strategy.experimental_distribute_dataset(get_validation_dataset(x_valid, y_valid, config['BATCH_SIZE'], AUTO, repeated=True, seed=SEED))
train_data_iter = iter(train_dist_ds)
valid_data_iter = iter(valid_dist_ds)
# Step functions
@tf.function
def train_step(data_iter):
def train_step_fn(x, y):
with tf.GradientTape() as tape:
probabilities = model(x, training=True)
loss_start = loss_fn_start(y['y_start'], probabilities[0], label_smoothing=0.2)
loss_end = loss_fn_end(y['y_end'], probabilities[1], label_smoothing=0.2)
loss = tf.math.add(loss_start, loss_end)
grads = tape.gradient(loss, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# update metrics
train_loss.update_state(loss)
train_loss_start.update_state(loss_start)
train_loss_end.update_state(loss_end)
for _ in tf.range(step_size):
strategy.experimental_run_v2(train_step_fn, next(data_iter))
@tf.function
def valid_step(data_iter):
def valid_step_fn(x, y):
probabilities = model(x, training=False)
loss_start = loss_fn_start(y['y_start'], probabilities[0])
loss_end = loss_fn_end(y['y_end'], probabilities[1])
loss = tf.math.add(loss_start, loss_end)
# update metrics
valid_loss.update_state(loss)
valid_loss_start.update_state(loss_start)
valid_loss_end.update_state(loss_end)
for _ in tf.range(valid_step_size):
strategy.experimental_run_v2(valid_step_fn, next(data_iter))
# Train model
model_path = 'model_fold_%d.h5' % (n_fold)
model = model_fn(config['MAX_LEN'])
optimizer = optimizers.Adam(learning_rate=lambda: lrfn(tf.cast(optimizer.iterations, tf.float32)//step_size))
loss_fn_start = losses.categorical_crossentropy
loss_fn_end = losses.categorical_crossentropy
train_loss = metrics.Sum()
valid_loss = metrics.Sum()
train_loss_start = metrics.Sum()
valid_loss_start = metrics.Sum()
train_loss_end = metrics.Sum()
valid_loss_end = metrics.Sum()
metrics_dict = {'loss': train_loss, 'loss_start': train_loss_start, 'loss_end': train_loss_end,
'val_loss': valid_loss, 'val_loss_start': valid_loss_start, 'val_loss_end': valid_loss_end}
history = custom_fit(model, metrics_dict, train_step, valid_step, train_data_iter, valid_data_iter,
step_size, valid_step_size, config['BATCH_SIZE'], config['EPOCHS'], config['ES_PATIENCE'],
(MODEL_BASE_PATH + model_path), save_last=False)
history_list.append(history)
model.save_weights(MODEL_BASE_PATH +'last_' + model_path)
model.load_weights(MODEL_BASE_PATH + model_path)
# Make predictions
train_preds = model.predict(get_test_dataset(x_train, config['BATCH_SIZE']))
valid_preds = model.predict(get_test_dataset(x_valid, config['BATCH_SIZE']))
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'start_fold_%d' % (n_fold)] = train_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'train', 'end_fold_%d' % (n_fold)] = train_preds[1].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'start_fold_%d' % (n_fold)] = valid_preds[0].argmax(axis=-1)
k_fold.loc[k_fold['fold_%d' % (n_fold)] == 'validation', 'end_fold_%d' % (n_fold)] = valid_preds[1].argmax(axis=-1)
k_fold['end_fold_%d' % (n_fold)] = k_fold['end_fold_%d' % (n_fold)].astype(int)
k_fold['start_fold_%d' % (n_fold)] = k_fold['start_fold_%d' % (n_fold)].astype(int)
k_fold['end_fold_%d' % (n_fold)].clip(0, k_fold['text_len'], inplace=True)
k_fold['start_fold_%d' % (n_fold)].clip(0, k_fold['end_fold_%d' % (n_fold)], inplace=True)
k_fold['prediction_fold_%d' % (n_fold)] = k_fold.apply(lambda x: decode(x['start_fold_%d' % (n_fold)], x['end_fold_%d' % (n_fold)], x['text'], config['question_size'], tokenizer), axis=1)
k_fold['prediction_fold_%d' % (n_fold)].fillna(k_fold["text"], inplace=True)
k_fold['jaccard_fold_%d' % (n_fold)] = k_fold.apply(lambda x: jaccard(x['selected_text'], x['prediction_fold_%d' % (n_fold)]), axis=1)
```
# Model loss graph
```
sns.set(style="whitegrid")
for n_fold in range(config['N_FOLDS']):
print('Fold: %d' % (n_fold+1))
plot_metrics(history_list[n_fold])
```
# Model evaluation
```
display(evaluate_model_kfold(k_fold, config['N_FOLDS']).style.applymap(color_map))
```
# Visualize predictions
```
display(k_fold[[c for c in k_fold.columns if not (c.startswith('textID') or
c.startswith('text_len') or
c.startswith('selected_text_len') or
c.startswith('text_wordCnt') or
c.startswith('selected_text_wordCnt') or
c.startswith('fold_') or
c.startswith('start_fold_') or
c.startswith('end_fold_'))]].head(15))
```
| github_jupyter |
```
#importing the Figshare dataset
import numpy as np
data=np.load('/content/drive/MyDrive/images5000.npy', allow_pickle="true")
labels=np.load('/content/drive/MyDrive/labels5000.npy', allow_pickle="true")
from google.colab import drive
drive.mount('/content/drive')
#importing other required libraries
from keras import Sequential
import numpy as np
import pandas as pd
from sklearn.utils.multiclass import unique_labels
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import seaborn as sns
import itertools
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from keras import Sequential
from keras.applications import VGG19, VGG16, ResNet50
from keras.preprocessing.image import ImageDataGenerator
from keras.optimizers import SGD,Adam
from keras.callbacks import ReduceLROnPlateau
from keras.layers import Flatten, Dense, BatchNormalization, Activation,Dropout
from keras.utils import to_categorical
import tensorflow as tf
import random
#already preprocessed data (from Matlab) is now divided into X and Y for entering the deep learning model
data=np.asarray(data)
labels=np.asarray(labels)
data = np.delete(data, [1382,2024,2034,2058,2145,2148,2155,2163,2170,2180,2183,2276,2288,2293,2299], axis=0)
labels = np.delete(labels, [1382,2024,2034,2058,2145,2148,2155,2163,2170,2180,2183,2276,2288,2293,2299], axis=0)
from tensorflow.keras.applications.resnet50 import preprocess_input,decode_predictions
for i in range(3049):
data[i]=data[i].flatten()
data[i]=data[i][:261075]
print(i)
data[i]=data[i].reshape((295,295,3))
data[i]=preprocess_input(data[i])
a=tf.keras.utils.to_categorical(labels, num_classes=4)
a=np.delete(a, 0, 1)
d=[]
for i in range(3049):
if i%4!=0:
p=data[i].reshape(295,295,3)
d.append(p)
test_i=[]
for i in range(3049):
if i %4 ==0 :
p=data[i].reshape(295,295,3)
test_i.append(p)
test_c=[]
for i in range(3049):
if i %4 ==0 :
test_c.append(a[i])
test_i=np.asarray(test_i)
test_c=np.asarray(test_c)
np.save("TESTI.npy",test_i)
np.save("TESTC.npy",test_c)
l=[]
for i in range(3049):
if i%4!=0:
l.append(a[i])
# for j in range(2290,3064):
# l.append(a[j])
d=np.asarray(d)
l=np.asarray(l)
l.shape
d.shape
#applying the RESNET 50 model on our preprocessed data
from keras.applications import VGG19, VGG16, ResNet50
from keras.models import Model
from keras.layers import Input
from keras.layers import AveragePooling2D
baseModel = ResNet50(weights="imagenet", include_top=False,input_tensor=Input(shape=(295,295, 3)))
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(7, 7))(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(128, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(3, activation="softmax")(headModel)
model = Model(inputs=baseModel.input, outputs=headModel)
# for layer in baseModel.layers:
# layer.trainable = False
INIT_LR = 1e-4
EPOCHS =10
BS = 32
opt = SGD(lr=1e-4, momentum=0.9)
model.compile(loss="binary_crossentropy", optimizer=opt,
metrics=["accuracy"])
(x_train, x_val, trainY, testY) = train_test_split(d,l,test_size=0.20, random_state=42)
H = model.fit(x_train, trainY, batch_size=11,steps_per_epoch=len(x_train)//11,validation_data=(x_val, testY),validation_steps=len(x_val)//11 ,epochs=20)
model.save("drive/MyDrive/model_75-25_histogram.h5")
#testing and evaluating our model on test data
test_i=np.asarray(test_i)
test_i.shape
test_c=np.asarray(test_c)
import keras
a=keras.models.load_model("drive/MyDrive/model_75-25_histogram.h5")
a.evaluate(test_i,test_c)
test_o=[]
for i in range(3049):
if i %4 ==0 :
test_o.append(labels[i])
k=[]
one=0
two=0
three=0
for i in test_o:
if i==1:
k.append(0)
one=one+1
if i==2:
k.append(1)
two=two+1
if i==3:
k.append(2)
three=three+1
k=np.asarray(k)
# Measuring the validity of our model using some of the metrics
from sklearn.metrics import classification_report
y_pred = a.predict(test_i, batch_size=64, verbose=0)
y_pred_bool = np.argmax(y_pred, axis=1)
print(classification_report(k, y_pred_bool))
```
| github_jupyter |
## Reinforcement Learning Tutorial -3: DDPG
### MD Muhaimin Rahman
contact: sezan92[at]gmail[dot]com
In the last tutorial, I tried to Explain DQN. DQN solves one problem, that is it can deal with continuous state space. But it cannot output continuous action. To solve that problem, here comes DDPG! It means Deep Deterministic Policy Gradient
- [Importing Libraries](#libraries)
- [Algorithm](#algorithm)
- [Model Definition](#model)
- [Replay Buffer](#buffer)
- [Noise Class](#noise)
- [Training](#training)
<a id ="libraries"></a>
### Importing Libraries
```
from __future__ import print_function,division
import gym
import keras
from keras import layers
from keras import backend as K
from collections import deque
from tqdm import tqdm
import random
import numpy as np
import copy
SEED =123
np.random.seed(SEED)
```
Important constants
```
num_episodes = 100
steps_per_episode=500
BATCH_SIZE=256
TAU=0.001
GAMMA=0.95
actor_lr=0.0001
critic_lr=0.001
SHOW= False
from keras.models import Model
```
<a id ="algorithm"></a>
### Algorithm
The actual algorithm was developed by Timothy lilicap et al. The algorithm is an actor-critic based algorithm.. Which means, it has two networks to train- an actor network, which predicts action based on the current state. The other networ- known as Critic network- evaluates the state and action. This is the case for all actor-critic networks. The critic network is updated using Bellman Equation like DQN. The difference is the training of actor network. In DDPG , we train Actor network by trying to get the maximum value of gradient of $Q(s,a)$ for given action $a$ in a state $s$. In a normal machine learning classification and regression algorithm, our target is to get the value with minimum loss. Then we train the network by gradient descent technique using the gradient of Loss .
\begin{equation}
\theta \gets \theta - \alpha \frac{\partial L}{\partial \theta}
\end{equation}
Here, $\theta$ is the weight parameter of the network, and $L$ is loss
But in our case, we have to get the maximize the $Q$ value. So we have to set the weight parameters such that we get the maximum $Q$ value. This technique is known as Gradient Ascent, as it does the exact opposite of Gradient Descent
\begin{equation}
\theta_a \gets \theta_a - \alpha (-\frac{\partial Q(s,a) }{\partial \theta_a})
\end{equation}
The above equation looks like the actual Gradient Descent equation. Only difference is , the minus sign. It makes the equation to minimize the negative value of $Q$ , which in turn maximizes $Q$ value.
So the training is as following
- 1) Define Actor network $actor$ and Critic Network $critic$
- 2) Define Target Actor and Critic Networks - $actor_{target}$ and $critic_{target}$ with exact same weights
- 3) Initialize Replay Buffer
- 4) Get the initial state , $state$
- 5) Get the action $a$ from , $a \gets actor(state)$ + Noise .[Here Noise is given to make the process stochastic and not so deterministic. The paper uses ornstein uhlenbeck noise process , so we will as well]
- 6) Get Next state $state_{next}$ , Reward $r$ , Terminal from agent for given $state$ and $action$
- 7) Add the experience , $state$,$action$,$reward$,$state_{next}$,$terminal$ to replay buffer
- 8) Get sample minibatch from Replay buffer
- 9) Train Critic Network Using Bellman Equation. Like DQN
- 10) Train Actor Network using Gradient Ascent with gradients of $Q$ . $\theta_a \gets \theta_a - \alpha (-\frac{\partial Q(s,a) }{\partial \theta_a}) $
- 11) Update weights of $actor_{target}$ and $critic_{target}$ using the equation $ \theta \gets \tau \theta + (1-\tau)\theta_{target}$
<a id ="model"></a>
### Model Definition
After some trials and errors, I have selected this network. The Actor Network is 3 layer MLP with 320 hidden nodes in each layer. The critic network is also a 3 layer MLP with 640 hidden nodes in each layer.Notice that the return arguments of function ```create_critic_network```.
```
def create_actor_network(state_shape,action_shape):
in1=layers.Input(shape=state_shape,name="state")
l1 =layers.Dense(320,activation="relu")(in1)
l2 =layers.Dense(320,activation="relu")(l1)
l3 =layers.Dense(320,activation="relu")(l2)
action =layers.Dense(action_shape,activation="tanh")(l3)
actor= Model(in1,action)
return actor
def create_critic_network(state_shape,action_shape):
in1 = layers.Input(shape=state_shape,name="state")
in2 = layers.Input(shape=action_shape,name="action")
l1 = layers.concatenate([in1,in2])
l2 = layers.Dense(640,activation="relu")(l1)
l3 = layers.Dense(640,activation="relu")(l2)
l4 = layers.Dense(640,activation="relu")(l3)
value = layers.Dense(1)(l4)
critic = Model(inputs=[in1,in2],outputs=value)
return critic,in1,in2
```
I am chosing ```MountainCarContinuous-v0``` game. Mainly because my GPU is not that good to work on higher dimensional state space
```
env = gym.make("MountainCarContinuous-v0")
state_shape= env.observation_space.sample().shape
action_shape=env.action_space.sample().shape
actor = create_actor_network(state_shape,action_shape[0])
critic,state_tensor,action_tensor = create_critic_network(state_shape,action_shape)
target_actor=create_actor_network(state_shape,action_shape[0])
target_critic,_,_ = create_critic_network(state_shape,action_shape)
target_actor.set_weights(actor.get_weights())
target_critic.set_weights(critic.get_weights())
```
I have chosen ```RMSProp``` optimizer, due to more stability compared to Adam . I found it after trials and errors, no theoritical background on chosing this optimizer
```
actor_optimizer = keras.optimizers.RMSprop(actor_lr)
critic_optimizer = keras.optimizers.RMSprop(critic_lr)
critic.compile(loss="mse",optimizer=critic_optimizer)
```
#### Actor training
I think this is the most critical part of ddpg in keras. The object ```critic``` and ```actor``` has a ```__call__``` method inside it, which will give output tensor if you give input a tensor. So to get the tensor object of ```Q``` we will use this functionality.
```
CriticValues = critic([state_tensor,actor(state_tensor)])
```
Now it is time to get the gradient value of $-\frac{\partial Q(s,a)}{\theta_a}$
```
updates = actor_optimizer.get_updates(
params=actor.trainable_weights,loss=-K.mean(CriticValues))
```
Now we will create a function which will train the actor network.
```
actor_train = K.function(inputs=[state_tensor],outputs=[actor(state_tensor),CriticValues],
updates=updates)
```
<a id ="buffer"></a>
### Replay Buffer
```
memory = deque(maxlen=10000)
state = env.reset()
state = state.reshape(-1,)
for _ in tqdm(range(memory.maxlen)):
action = env.action_space.sample()
next_state,reward,terminal,_=env.step(action)
state=next_state
if terminal:
reward=-100
state= env.reset()
state = state.reshape(-1,)
memory.append((state,action,reward,next_state,terminal))
next_state
```
<a id ="noise"></a>
### Noise class
The use of Noise is to make the process Stochastic and to help the agent explore different actions. The paper used Orstein Uhlenbeck Noise class
```
class OrnsteinUhlenbeckProcess(object):
def __init__(self, theta, mu=0, sigma=1, x0=0, dt=1e-2, n_steps_annealing=10, size=1):
self.theta = theta
self.sigma = sigma
self.n_steps_annealing = n_steps_annealing
self.sigma_step = - self.sigma / float(self.n_steps_annealing)
self.x0 = x0
self.mu = mu
self.dt = dt
self.size = size
def restart(self):
self.x0=copy.copy(self.mu)
def generate(self, step):
#sigma = max(0, self.sigma_step * step + self.sigma)
x = self.x0 + self.theta * (self.mu - self.x0) * self.dt + self.sigma * np.sqrt(self.dt) * np.random.normal(size=self.size)
self.x0 = x
return x
```
<a id ="training"></a>
### Training
```
steps_per_episodes=5000
ou = OrnsteinUhlenbeckProcess(theta=0.35,mu=0.8,sigma=0.4,n_steps_annealing=10)
max_total_reward=0
for episode in range(num_episodes):
state= env.reset()
state = state.reshape(-1,)
total_reward=0
ou.restart()
for step in range(steps_per_episodes):
action= actor.predict(state.reshape(1,-1))+ou.generate(episode)
next_state,reward,done,_ = env.step(action)
total_reward=total_reward+reward
#random minibatch from buffer
batches=random.sample(memory,BATCH_SIZE)
states= np.array([batch[0].reshape((-1,)) for batch in batches])
actions= np.array([batch[1] for batch in batches])
actions=actions.reshape(-1,1)
rewards=np.array([batch[2] for batch in batches])
rewards = rewards.reshape((-1,1))
new_states=np.array([batch[3].reshape((-1,)) for batch in batches])
terminals=np.array([batch[4] for batch in batches])
terminals = terminals.reshape((-1,1))
#training
target_actions = target_actor.predict(new_states)
target_Qs = target_critic.predict([new_states,target_actions])
new_Qs = rewards+GAMMA*target_Qs*terminals
critic.fit([states,actions],new_Qs,verbose=False)
_,critic_values=actor_train(inputs=[states])
target_critic_weights=[TAU*weight+(1-TAU)*target_weight for weight,target_weight in zip(critic.get_weights(),target_critic.get_weights())]
target_actor_weights=[TAU*weight+(1-TAU)*target_weight for weight,target_weight in zip(actor.get_weights(),target_actor.get_weights())]
target_critic.set_weights(target_critic_weights)
target_actor.set_weights(target_actor_weights)
print("Total Reward %f"%total_reward,end="\r")
if SHOW:
env.render()
if done or step==(steps_per_episodes-1):
if total_reward<0:
print("Failed!",end=" ")
reward=-100
elif total_reward>0:
print("Passed!",end=" ")
reward=100
memory.append((state,action,reward,next_state,done))
break
memory.append((state,action,reward,next_state,done))
state=next_state
if total_reward>max_total_reward:
actor.save_weights("MC_DDPG_Weights/Actor_Best_weights episode %d_GAMMA_%f_TAU%f_lr_%f.h5"%(episode,GAMMA,TAU,actor_lr))
critic.save_weights("MC_DDPG_Weights/Critic_Best_weights episode %d_GAMMA_%f_TAU%f_lr_%f.h5"%(episode,GAMMA,TAU,critic_lr))
max_total_reward=total_reward
print("Episode %d Total Reward %f"%(episode,total_reward))
```
### Video
Please watch at 2x speed. I changed some simple mistakes after the video so the rewards are not exactly the same
[](http://www.youtube.com/watch?v=9Fe_n-ovIaA "Keras tutorial DDPG")
| github_jupyter |
# Learning LDA model via Gibbs sampling
# Question 1
<img src="images/Screen Shot 2016-07-29 at 3.14.04 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
# Question 2
<img src="images/Screen Shot 2016-07-29 at 3.14.28 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
# Question 3
<img src="images/Screen Shot 2016-07-29 at 5.11.56 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
# Question 4
<img src="images/Screen Shot 2016-07-29 at 4.53.54 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- https://www.coursera.org/learn/ml-clustering-and-retrieval/discussions/weeks/5/threads/AK3N4kI8EeaXyw5hjmsWew
- https://www.coursera.org/learn/ml-clustering-and-retrieval/discussions/weeks/5/threads/L5yUeFA4EearRRKGx4XuoQ
- Understand notation:
- $n_{i,k}$: count of topics (1's and 2's) in the document after you decrement the target word "manager". If the target word is for topic 1 then you won't need to decrement since the manager = topic 2. So you count the 1's. If the target word "manager" is for topic 2 then you have to decrement the single count of topic 2 which then makes your $n_{i,k}$ = 0.
- $N_i$: count of words in doc i
- V: total count of vocabulary, which is 10
- $m_{manager,k}$: total count of word, "manager" in the corpus assigned to topic k
- $\sum_{w} m_{w,k}$: Sum of count of all words in the corpus assigned to topic k
# Question 5
<img src="images/Screen Shot 2016-07-29 at 4.54.06 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- $\sum_w m_{w, 1}$: total number of words in the corpus of topic 1, which is the sum of all words assigned to topic 1
- 52 + 15 + 9 + 9 + 20 + 17 + 1 = 123
# Question 6
<img src="images/Screen Shot 2016-07-29 at 4.54.09 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- $n_{i, 1}$: # current assignments to topic 1 in doc i, which is how many times topic 1 appears in document i.
- clearly 3 times, for baseball + ticket + owner = 1 + 1 + 1 = 3
# Question 7
<img src="images/Screen Shot 2016-07-29 at 4.54.13 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- $\sum_w m_{w, 1}$: total number of words in the corpus of topic 1, which is the sum of all words assigned to topic 1
- 52 + 15 + 9 + 9 + 20 + 17 + 1 = 123
**Answer**
- When we remove the assignment of manager to topic 2 in document i, we only have 1 assignment of topic 2 which is the word "price"
- $n_{i, 2}$: # current assignments to topic 2 in doc i, which is 1
# Question 8
<img src="images/Screen Shot 2016-07-29 at 4.54.18 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- When we remove the assignment of "manager" to topic 2 in document i -> manager: 0
- The total counts of manager in topic 2 in the corpus:
- number of assignments corpus-wide of word "manager" to topic 2 - number of assignment of word "manager" to topic 2 in document i
- $\large m_{\text{manager,2}} - z_{\text{i,manager}}$ = 37 - 1 = 36
# Question 9
<img src="images/Screen Shot 2016-07-29 at 5.12.06 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
- $\sum_w m_{w, 2}$: total number of words in the corpus of topic 2, which is the sum of all words assigned to topic 2 after we decrement the associated counts
- 2 + 25 + 36 + 32 + 23 + 75 + 19 + 29 = 241
# Question 10
<img src="images/Screen Shot 2016-07-29 at 5.17.14 PM.png">
*Screenshot taken from [Coursera](https://www.coursera.org/learn/ml-clustering-and-retrieval/exam/6ieZu/learning-lda-model-via-gibbs-sampling)*
<!--TEASER_END-->
**Answer**
As discussed in the slides, the unnormalized probability of assigning to topic 1 is
- $p_1 = \frac{n_{i, 1} + \alpha}{N_i - 1 + K \alpha}\frac{m_{\text{manager}, 1} + \gamma}{\sum_w m_{w, 1} + V \gamma}$
where V is the total size of the vocabulary.
Similarly the unnormalized probability of assigning to topic 2 is
- $p_2 = \frac{n_{i, 2} + \alpha}{N_i - 1 + K \alpha}\frac{m_{\text{manager}, 2} + \gamma}{\sum_w m_{w, 2} + V \gamma}$
Using the above equations and the results computed in previous questions, compute the probability of assigning the word “manager” to topic 1.
- Left Formula = (# times topic_1 appears in doc + alpha) / (# words in doc - 1 + K * alpha)
- Right Formula = (# of corpus-wide assignment of 'manager' to topic 1 + gamma) / (Sum of all topic 1 word counts + V * gamma)
- Prob 1 = (Left Formula) * (Right Formula)
Example:
- calculate prob 1
- Left Formula = (3 + 10.0) / (5 - 1 + (2 * 10.0)) = 0.5417
- Right Formula = (20 + 0.1) / (123 + (10 * 0.1)) = 0.162
- Prob 1 = 0.5417 * 0.162 = 0.0877554
- calculate prob 2
- Left Formula = (1 + 10.0) / (5 - 1 + (2 * 10.0)) = 0.4583
- Right Formula = (36 + 0.1) / (241 + (10 * 0.1)) = 0.1492
- Prob 1 = 0.4583 * 0.1492 = 0.06837836
- normalize prob 1
- normalize prob 1 = Prob 1/(Prob 1 + Prob 2) = 0.0877554/(0.0877554 + 0.06837836) = 0.562
```
def calculate_unnorm_prob(n_i, alpha, N_i, K, m_word, gamma, sum_of_m, V):
""" Calculate unnormalized probability of assigning to topic
"""
left_formula = (n_i + alpha)/(N_i - 1 + (K * alpha))
right_formula = (m_word + gamma)/(sum_of_m + (V * gamma))
prob = left_formula * right_formula
return prob
unnorm_prob_1 = calculate_unnorm_prob(3, 10.0, 5, 2, 20, 0.1, 123, 10)
unnorm_prob_2 = calculate_unnorm_prob(1, 10.0, 5, 2, 36, 0.1, 241, 10)
print unnorm_prob_1
print unnorm_prob_2
prob_1 = unnorm_prob_1/(unnorm_prob_1 + unnorm_prob_2)
print prob_1
```
| github_jupyter |
# SciPy - Scientific Computing for Python
SciPy is a framework that is built upon NumPy. It uses NumPy arrays to leverage performance, and then extends NumPy to provide a range of advanced algorithms and functions. We will only cover a brief section of SciPy...there's a lot in it. For more information you can see the SciPy website www.scipy.org.
The different submodules of SciPy include:
* Special functions (`scipy.special`)
* Integration (`scipy.integrate`)
* Optimization (`scipy.optimize`)
* Interpolation (`scipy.interpolate`)
* Fourier Transforms (`scipy.fftpack`)
* Signal Processing (`scipy.signal`)
* Linear Algebra (`scipy.linalg`)
* Sparse Eigenvalue Problems with ARPACK
* Compressed Sparse Graph Routines (`scipy.sparse.csgraph`)
* Spatial data structures and algorithms (`scipy.spatial`)
* Statistics (`scipy.stats`)
* Multidimensional image processing (`scipy.ndimage`)
* File IO (`scipy.io`)
To access the SciPy module, you can import the whole module,
```
from scipy import *
```
or simply import the parts you need:
```
import scipy.linalg as la
```
## Integration
### Quadrature
Many of you are probably familiar with integration from basic calculus:
$\displaystyle \int_a^b f(x) dx$
When you discretize this process and solve an integral numerically, it's called quadrature. SciPy provides several types of quadrature, depending on whether you need to solve single, double, or triple integrals.
```
from scipy.integrate import quad, dblquad, tplquad
# Define a function we want to integrate
def f(x):
return x**2
x0 = 0
x1 = 1
val, abserr = quad(f, x0, x1)
print("integral value =", val, ", absolute error =", abserr)
```
### Ordinary Differential Equations (ODEs)
SciPy's integration package also allows for users to numerically solve ODEs. SciPy provides a function, `odeint`, for solving first order, vector-valued, differential equations:
$\displaystyle \frac{d\mathbf{y}}{dt} = \mathbf{f}(\mathbf{y}, t)$
The `odeint` function takes 3 inputs:
* the function to be evaluated
* initial conditions
* a sequence of time points
In addition to the `odeint` function, SciPy also features a class called `ode` that has more options and finer levels of control. In general, `odeint` is a good starting point for new users.
Here's an example of using `odeint` to solve the predator-prey equations (a simple model that describes the interaction between two species...one the prey, and the other the predator):
$\displaystyle \frac{dx}{dt} = x(a-by)$
$\displaystyle \frac{dy}{dt} = -y(c-dx)$
Let $\displaystyle x$ be the population of rabbits and $\displaystyle y$ be the population of wolves. $\displaystyle a, b, c$ and $\displaystyle d$ are all positive parameters.
```
import numpy as np
from scipy.integrate import odeint
a,b,c,d = 1,1,1,1
# Define our system of ODEs
# Note the vector values, here x is P[0] and y is P[1]
def dP_dt(P, t):
return [P[0]*(a - b*P[1]), -P[1]*(c - d*P[0])]
# Discretize our domain
ts = np.linspace(0, 12, 100)
# Initial conditions
P0 = [1.5, 1.0]
# Call the solver
Ps = odeint(dP_dt, P0, ts)
prey = Ps[:,0]
predators = Ps[:,1]
import matplotlib.pyplot as plt
%matplotlib inline
# Plot the result
plt.plot(ts, prey, "+", label="Rabbits")
plt.plot(ts, predators, "x", label="Wolves")
plt.xlabel("Time")
plt.ylabel("Population")
plt.legend();
```
We can even look at a phase plane plot:
```
ic = np.linspace(1.0, 3.0, 21)
for r in ic:
P0 = [r, 1.0]
Ps = odeint(dP_dt, P0, ts)
plt.plot(Ps[:,0], Ps[:,1], "-")
plt.xlabel("Rabbits")
plt.ylabel("Wolves")
plt.title("Rabbits vs Wolves");
```
## Fourier Transform
SciPy's `fftpack` module provides users with a performant way to compute discrete Fourier transforms (DFT). Underneath the hood, `fftpack` is calling Fortran functions from the Fortran FFTPACK library.
Here's a brief example:
```
from scipy.fftpack import fft
import numpy
x = np.random.randint(10, size=5)
y = fft(x)
y
```
We can also compute the inverse:
```
from scipy.fftpack import ifft
yinv = ifft(y)
yinv
x
```
We won't go into the theory of Fourier transforms, but let us briefly mention what is involved in computing a DFT to highlight SciPy's performance.
Computing Fourier transforms reduces to a simple matrix multiplication; to compute the DFT for a vector $\displaystyle x$ we simply multiply it with the matrix $\displaystyle M$:
$\displaystyle M = e^{-2i\pi kn/N}$
We can use the matrix-vector operations we learned about in NumPy to write a naive implementation for computing DFT:
```
import numpy as np
def DFT(x):
x = np.asarray(x, dtype=float)
N = x.shape[0]
n = np.arange(N)
k = n.reshape((N, 1))
M = np.exp(-2j * np.pi * k * n / N)
return np.dot(M, x)
```
Let's double-check it works by comparing results with `fftpack`:
```
from scipy.fftpack import fft
x = np.random.random(1024)
np.allclose(DFT(x), fft(x))
```
Now let's look at the performance:
```
%timeit DFT(x)
%timeit fft(x)
```
## Linear Algebra
SciPy's `linalg` package greatly expands upon what we saw in NumPy. There are a number of solvers available, including data interpolation modules, eigensolvers, and optimisation routines, but we'll only cover a few here.
Let us consider a system of linear equations, in matrix form:
$A x = b$
where $A$ is a matrix and $x,b$ are vectors.
Such system can be solved with SciPy like:
```
from scipy.linalg import *
A = np.array([[1,2],[3,4]])
b = np.array([3,17])
x = solve(A, b)
x
# check
np.allclose(A @ x, b)
```
SciPy's `solve` function is faster than the built-in `inv` function:
```
A1 = np.random.random((1000,1000))
b1 = np.random.random(1000)
%timeit solve(A1, b1)
%timeit inv(A1) @ b1
```
We can even import LAPACK functions for better performance:
```
import scipy.linalg.lapack as lapack
%timeit lu, piv, x, info = lapack.dgesv(A1, b1)
```
| github_jupyter |
# Print Formatting
In this lecture we will briefly cover the various ways to format your print statements. As you code more and more, you will probably want to have print statements that can take in a variable into a printed string statement.
The most basic example of a print statement is:
### <font color='red'>Python 3 Alert!</font>
Python3 is simpler. We use the format string.
## String formatting
```
print('The first letter is: {}, the second: {}, the third: {}'.format('a','b','c'))
```
Each parameter is indexed. We can manipulate where the argument appears in the string by using its index.
```
print('The first letter is: {2}, the second: {1}, the third: {0}'.format('a','b','c'))
We can also repeat them the argument.
print('The first letter is: {0}, the second: {0}, the third: {0}'.format('a'))
```
## Integers and Floats
There is no need to specify what data type we are passing in.
```
print('The number is {0}, and {0}, and {0}'.format(10))
```
Floats are really just numbers with decimals.
```
print('The number is {0}, and {0}, and {0}'.format(10.00020))
```
## Passing in 'weird' data types aka things we haven't seen yet.
We can also pass in dictionaries, lists, sets, tuples. Don't worry about what these means, they are other types of special data types that are also collections.
```
print('The number is {0}, and {0}, and {0}'.format([20202, 39394]))
print('The number is {0}, and {0}, and {0}'.format({'Tru TV': 'Chris Gethard'}))
print('The number is {0}, and {0}, and {0}'.format({'Tru TV'}))
print('The number is {0}, and {1}, and {0}'.format('A$AP Ferg', 'A$AP Rocky'))
```
## Setting parameters
We can set the arguments to a name and use that name inside the string
```
print('The number is {a}, and {b}, and {a}'.format(a=10.00020, b=20))
```
### <font color='blue'>Python 2 Alert!</font>
Python2 is more complicated to format strings. Python3 improved upon this.
```
print 'This is a string'
```
## Strings
You can use the %s to format strings into your print statements.
```
s = 'STRING'
print 'Place another string with a mod and s: %s' %(s)
```
## Floating Point Numbers aka Floats
Floating point numbers use the format %n1.n2f where the n1 is the total minimum number of digits the string should contain (these may be filled with whitespace if the entire number does not have this many digits. The n2 placeholder stands for how many numbers to show past the decimal point. Lets see some examples:
```
print 'Floating point numbers: %1.2f' %(13.144)
print 'Floating point numbers: %1.0f' %(13.144)
print 'Floating point numbers: %1.5f' %(13.144)
print 'Floating point numbers: %10.2f' %(13.144)
print 'Floating point numbers: %25.2f' %(13.144)
```
## Conversion Format methods.
It should be noted that two methods %s and %r actually convert any python object to a string using two separate methods: str() and repr(). We will learn more about these functions later on in the course, but you should note you can actually pass almost any Python object with these two methods and it will work:
```
print 'Here is a number: %s. Here is a string: %s' %(123.1,'hi')
print 'Here is a number: %r. Here is a string: %r' %(123.1,'hi')
```
## Multiple Formatting
Pass a tuple to the modulo symbol to place multiple formats in your print statements:
```
print 'First: %s, Second: %1.2f, Third: %r' %('hi!',3.14,22)
```
## Using the string .format() method
The best way to format objects into your strings for print statements is using the format method. The syntax is:
'String here {var1} then also {var2}'.format(var1='something1',var2='something2')
Lets see some examples:
```
print 'This is a string with an {p}'.format(p='insert')
# Multiple times:
print 'One: {p}, Two: {p}, Three: {p}'.format(p='Hi!')
# Several Objects:
print 'Object 1: {a}, Object 2: {b}, Object 3: {c}'.format(a=1,b='two',c=12.3)
```
That is the basics of string formatting! Remember that Python 3 uses a print() function, not the print statement!
| github_jupyter |
```
# library of congress
# import libraries
import rdflib, pandas, pathlib, json
import numpy, uuid, xmltodict, pydash
# define graph and namespace
graph = rdflib.Graph()
name_loc = rdflib.Namespace('https://loc.gov/')
name_wb = rdflib.Namespace('http://wikibas.se/ontology')
name_fiaf = rdflib.Namespace('https://www.fiafnet.org/')
# useful functions
def make_claim(s, p, o):
claim_id = name_loc[f"resource/claim/{uuid.uuid4()}"]
graph.add((s, name_wb['#claim'], claim_id))
graph.add((claim_id, p, o))
return claim_id
def make_qual(s, p, o):
qual_id = name_loc[f"resource/qualifier/{uuid.uuid4()}"]
graph.add((s, name_wb['#qualifier'], qual_id))
graph.add((qual_id, p, o))
return qual_id
def reference(claim_id, institute):
ref_id = name_loc[f"resource/reference/{uuid.uuid4()}"]
graph.add((claim_id, name_wb['#reference'], ref_id))
graph.add((ref_id, name_fiaf['ontology/property/contributed_by'], institute))
def single_list(data):
if isinstance(data, list):
return data
else:
return [data]
# define institution
graph.add((name_loc['ontology/item/loc'], rdflib.RDFS.label, rdflib.Literal('Library of Congress', lang='en')))
make_claim(name_loc['ontology/item/loc'], name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/holding_institution'])
make_claim(name_loc['ontology/item/loc'], name_fiaf['ontology/property/located_in'], name_fiaf['ontology/item/usa'])
print(len(graph))
# format data
path = pathlib.Path.home() / 'murnau-data' / 'library_of_congress'
data = list()
for f in [x for x in path.glob('**/*.xml')]:
with open(f, encoding='ISO-8859-1') as xml_data:
element = xmltodict.parse(xml_data.read())
data.append(single_list(pydash.get(element, 'mavis.TitleWork'))[0])
print(len(graph))
# write work
for x in data:
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
make_claim(work, name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/work'])
claim_id = make_claim(work, name_fiaf['ontology/property/external_id'], rdflib.Literal(work_id))
make_qual(claim_id, name_fiaf['ontology/property/institution'], name_loc['ontology/item/loc'])
reference(claim_id, name_loc['ontology/item/loc'])
print(len(graph))
# write title
for x in data:
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
selected_title = ''
for t in ['preferredTitle.Title', 'alternateTitles.Title']:
for y in single_list(pydash.get(x, t)):
if 'Original title' in str(y) or 'German' in str(y):
selected_title = pydash.get(y, '@xl:title')
title_type = name_fiaf['ontology/item/original_title']
if selected_title == '':
selected_title = pydash.get(x, '@xl:title')
title_type = name_fiaf['ontology/item/work_title']
claim_id = make_claim(work, name_fiaf['ontology/property/title'], rdflib.Literal(selected_title[:-1]))
make_qual(claim_id, name_fiaf['ontology/property/title_type'], title_type)
reference(claim_id, name_loc['ontology/item/loc'])
print(len(graph))
# write country
for x in data:
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
country = pydash.get(x, 'countries.WorkCountry.@xl:title')
if country == 'US':
fiaf_country = name_fiaf['ontology/item/usa']
elif country == 'GG':
fiaf_country = name_fiaf['ontology/item/germany']
else:
raise Exception('Unknown country.')
claim = make_claim(work, name_fiaf['ontology/property/production_country'], fiaf_country)
reference(claim, name_loc['ontology/item/loc'])
print(len(graph))
# write agents
def write_credit(work_data, dict_key, agent_type):
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
for a in pydash.get(x, 'roles.Name-Role'):
if 'Person' in pydash.get(a, 'party') and pydash.get(a, 'role.@xl:title') == dict_key:
forename = pydash.get(a, 'party.Person.preferredName.PersonName.firstName')
surname = pydash.get(a, 'party.Person.preferredName.PersonName.name')
contribution = pydash.get(a, 'role.@xl:title')
key = pydash.get(a, 'party.Person.@xl:href').split('/')[-1]
gend = pydash.get(a, 'party.Person.gender.@xl:title')
agent = name_loc[f"resource/agent/{key}"]
claim1 = make_claim(work, name_fiaf['ontology/property/agent'], agent)
make_qual(claim1, name_fiaf['ontology/property/agent_type'], agent_type)
reference(claim1, name_loc['ontology/item/loc'])
make_claim(agent, name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/agent'])
claim2 = make_claim(agent, name_fiaf['ontology/property/external_id'], rdflib.Literal(key))
make_qual(claim2, name_fiaf['ontology/property/institution'], name_loc['ontology/item/loc'])
reference(claim2, name_loc['ontology/item/loc'])
if forename != None:
claim3 = make_claim(agent, name_fiaf['ontology/property/forename'], rdflib.Literal(forename))
reference(claim3, name_loc['ontology/item/loc'])
claim4 = make_claim(agent, name_fiaf['ontology/property/surname'], rdflib.Literal(surname))
reference(claim4, name_loc['ontology/item/loc'])
if gend == 'Male':
claim_id = make_claim(agent, name_fiaf['ontology/property/gender'], name_fiaf['ontology/item/male'])
reference(claim_id, name_loc['ontology/item/loc'])
if gend == 'Female':
claim_id = make_claim(agent, name_fiaf['ontology/property/gender'], name_fiaf['ontology/item/female'])
reference(claim_id, name_loc['ontology/item/loc'])
claim_id = make_claim(agent, name_fiaf['ontology/property/work'], work)
reference(claim_id, name_loc['ontology/item/loc'])
for x in data:
write_credit(x, 'Cast/Actor', name_fiaf['ontology/item/cast'])
write_credit(x, 'Director', name_fiaf['ontology/item/director'])
write_credit(x, 'Producer', name_fiaf['ontology/item/producer'])
write_credit(x, 'Cinematographer/Director of Photography', name_fiaf['ontology/item/cinematographer'])
write_credit(x, 'Scriptwriter', name_fiaf['ontology/item/screenwriter'])
write_credit(x, 'Music Composer', name_fiaf['ontology/item/composer'])
print(len(graph))
# write events
for x in data:
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
date_data = pydash.get(x, 'objectDates.Date-Year')
for y in [a for a in single_list(date_data) if pydash.get(a, 'dateType.@xl:title') == 'Copyright']:
date = pydash.get(y, 'yearFrom')
date += f"-{pydash.get(y, 'monthFrom').zfill(2)}"
date += f"-{pydash.get(y, 'dayFrom').zfill(2)}"
claim_id = make_claim(work, name_fiaf['ontology/property/event'], rdflib.Literal(date))
make_qual(claim_id, name_fiaf['ontology/property/event_type'], name_fiaf['ontology/item/decision_copyright'])
make_qual(claim_id, name_fiaf['ontology/property/country'], name_fiaf['ontology/item/usa'])
reference(claim_id, name_loc['ontology/item/loc'])
print(len(graph))
# write manifestations/items
for x in data:
work_id = x['@xl:href'].split('/')[-1]
work = name_loc[f"resource/work/{work_id}"]
items = list()
for c in [x['components'][y] for y in x['components']]:
c = single_list(c)
for y in c:
items.append(y)
for i in items:
manifestation = name_loc[f"resource/manifestation/{uuid.uuid4()}"]
make_claim(manifestation, name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/manifestation'])
make_claim(manifestation, name_fiaf['ontology/property/manifestation_of'], work)
item_id = pydash.get(i, 'itemId')
item = name_loc[f"resource/item/{item_id}"]
make_claim(item, name_fiaf['ontology/property/instance_of'], name_fiaf['ontology/item/item'])
make_claim(item, name_fiaf['ontology/property/item_of'], manifestation)
claim_id = make_claim(item, name_fiaf['ontology/property/held_at'], name_loc['ontology/item/loc'])
reference(claim_id, name_loc['ontology/item/loc'])
claim_id = make_claim(item, name_fiaf['ontology/property/external_id'], rdflib.Literal(item_id))
make_qual(claim_id, name_fiaf['ontology/property/institution'], name_loc['ontology/item/loc'])
reference(claim_id, name_loc['ontology/item/loc'])
for k, v in {'Safety':name_fiaf['ontology/item/film'], 'Nitrate':name_fiaf['ontology/item/film'],
'Digital':name_fiaf['ontology/item/digital'], 'Video':name_fiaf['ontology/item/video_tape'],
'Tape':name_fiaf['ontology/item/sound_tape'], 'Disc':name_fiaf['ontology/item/disc']}.items():
if pydash.get(i, 'itemType.@xl:title') == k:
claim_id = make_claim(item, name_fiaf['ontology/property/carrier'], v)
reference(claim_id, name_loc['ontology/item/loc'])
for k, v in {'16mm':name_fiaf['ontology/item/16mm'], '35mm':name_fiaf['ontology/item/35mm']}.items():
if pydash.get(i, 'gauge.@xl:title') == k:
claim_id = make_claim(item, name_fiaf['ontology/property/specific_carrier'], v)
reference(claim_id, name_loc['ontology/item/loc'])
for k, v in {'Composite Positive':name_fiaf['ontology/item/print'], 'Duplicate Negative Track':name_fiaf['ontology/item/duplicate_negative'],
'Duplicate Negative Picture':name_fiaf['ontology/item/duplicate_negative'], 'Positive Picture':name_fiaf['ontology/item/duplicate_positive']}.items():
if pydash.get(i, 'techCode.@xl:title') == k:
claim_id = make_claim(item, name_fiaf['ontology/property/element'], v)
reference(claim_id, name_loc['ontology/item/loc'])
for k, v in {'Access':name_fiaf['ontology/item/viewing'], 'Preservation Copy':name_fiaf['ontology/item/master'],
'Access/Browsing copy':name_fiaf['ontology/item/viewing'], 'Limited Access':name_fiaf['ontology/item/restricted'],
'Preservation Material':name_fiaf['ontology/item/master']}.items():
if pydash.get(i, 'categoryMaterial.@xl:title') == k:
claim_id = make_claim(item, name_fiaf['ontology/property/access'], v)
reference(claim_id, name_loc['ontology/item/loc'])
for k, v in {'Safety':name_fiaf['ontology/item/acetate'], 'Nitrate':name_fiaf['ontology/item/nitrate']}.items():
if pydash.get(i, 'itemType.@xl:title') == k:
claim_id = make_claim(item, name_fiaf['ontology/property/base'], v)
reference(claim_id, name_loc['ontology/item/loc'])
make_claim(work, name_fiaf['ontology/property/manifestation'], manifestation)
make_claim(manifestation, name_fiaf['ontology/property/item'], item)
print(len(graph))
graph.serialize(destination=str(pathlib.Path.cwd() / 'library_of_congress.ttl'), format="turtle")
print(len(graph))
```
| github_jupyter |
จาก ep ก่อนที่เราได้เรียนรู้ความสำคัญของ Hyperparameter ของการเทรน Machine Learning ที่ชื่อ [Learning Rate](https://www.bualabs.com/archives/618/learning-rate-deep-learning-how-to-hyperparameter-tuning-ep-1/) ถ้าเรากำหนดค่า Learning น้อยไปก็ทำให้เทรนได้ช้า แต่ถ้ามากเกินไปก็ทำให้ไม่ Converge
แล้วเราจะทราบได้อย่างไร ว่า Learning Rate เท่าไร เป็นค่าที่ดีที่สุดในการเทรน Deep Neural Network ของเรา
เราจะสอนวิธีหา Learning Rate ที่ดีที่สุด ที่ดีที่สุด หมายถึง Learning Rate ที่มากที่สุด ที่ยังไม่ทำให้เกิดการไม่ Converge ด้วยอัลกอริทึมง่าย ๆ ตรงตัว คือ การทดลองเทรน แล้วเพิ่ม Learning Rate ขึ้นไปเรื่อย ๆ แล้วเช็ค Loss จนกว่า Loss จะเพิ่มมากจนผิดปกติ เรียกว่า LR Finder (Learning Rate Finder) โดยใช้ [LR Finder Callback เริ่มที่หัวข้อ 6](#6.-Callbacks)
เมื่อเราได้ข้อมูล ความสัมพันธ์ระหว่าง Learning Rate กับ Loss ของโมเดล Deep Neural Network ของเรามาเรียบร้อยแล้ว เราจะนำมาพล็อตกราฟ เพื่อวิเคราะห์หา Learning Rate ที่ดีที่สุดต่อไป
# 0. Magic
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
```
# 1. Import
```
import torch
from torch import tensor
from torch.nn import *
import torch.nn.functional as F
from torch.utils.data import *
from fastai import datasets
from fastai.metrics import accuracy
import pickle, gzip, math, torch, re
from IPython.core.debugger import set_trace
import matplotlib.pyplot as plt
```
# 2. Data
```
class Dataset(Dataset):
def __init__(self, x, y):
self.x, self.y = x, y
def __len__(self):
return len(self.x)
def __getitem__(self, i):
return self.x[i], self.y[i]
MNIST_URL='http://deeplearning.net/data/mnist/mnist.pkl'
def get_data():
path = datasets.download_data(MNIST_URL, ext='.gz')
with gzip.open(path, 'rb') as f:
((x_train, y_train), (x_valid, y_valid), _) = pickle.load(f, encoding='latin-1')
return map(tensor, (x_train, y_train, x_valid, y_valid))
x_train, y_train, x_valid, y_valid = get_data()
def normalize(x, m, s):
return (x-m)/s
from typing import *
def listify(o):
if o is None: return []
if isinstance(o, list): return o
if isinstance(o, str): return [o]
if isinstance(o, Iterable): return list(o)
return [o]
_camel_re1 = re.compile('(.)([A-Z][a-z]+)')
_camel_re2 = re.compile('([a-z0-9])([A-Z])')
def camel2snake(name):
s1 = re.sub(_camel_re1, r'\1_\2', name)
return re.sub(_camel_re2, r'\1_\2', s1).lower()
train_mean, train_std = x_train.mean(), x_train.std()
x_train = normalize(x_train, train_mean, train_std)
x_valid = normalize(x_valid, train_mean, train_std)
nh, bs = 100, 256
n, m = x_train.shape
c = (y_train.max()+1).numpy()
loss_func = F.cross_entropy
train_ds, valid_ds = Dataset(x_train, y_train), Dataset(x_valid, y_valid)
train_dl, valid_dl = DataLoader(train_ds, bs), DataLoader(valid_ds, bs)
```
# 3. DataBunch
```
class DataBunch():
def __init__(self, train_dl, valid_dl, c=None):
self.train_dl,self.valid_dl,self.c = train_dl,valid_dl,c
@property
def train_ds(self): return self.train_dl.dataset
@property
def valid_ds(self): return self.valid_dl.dataset
data = DataBunch(train_dl, valid_dl, c)
```
# 4. Model
```
lr = 0.03
epoch = 10
def get_model():
# loss function
loss_func = F.cross_entropy
model = Sequential(Linear(m, nh), ReLU(), Linear(nh,c))
return model, loss_func
class Learner():
def __init__(self, model, opt, loss_func, data):
self.model, self.opt, self.loss_func, self.data = model, opt, loss_func, data
```
# 5. Training Loop
Training Loop ที่รองรับ Callback
```
class Runner():
def __init__(self, cbs=None, cb_funcs=None):
cbs = listify(cbs)
for cbf in listify(cb_funcs):
cb = cbf()
setattr(self, cb.name, cb)
cbs.append(cb)
self.stop, self.cbs = False, [TrainEvalCallback()]+cbs
@property
def opt(self): return self.learn.opt
@property
def model(self): return self.learn.model
@property
def loss_func(self): return self.learn.loss_func
@property
def data(self): return self.learn.data
def one_batch(self, xb, yb):
try:
self.xb, self.yb = xb, yb
self('begin_batch')
self.pred = self.model(xb)
self('after_pred')
self.loss = self.loss_func(self.pred, yb)
self('after_loss')
if not self.in_train: return
self.loss.backward()
self('after_backward')
self.opt.step()
self('after_step')
self.opt.zero_grad()
except CancelBatchException: self('after_cancel_batch')
finally: self('after_batch')
def all_batches(self, dl):
self.iters = len(dl)
try:
for xb, yb in dl:
self.one_batch(xb, yb)
except CancelEpochException: self('after_cancel_epoch')
def fit(self, epochs, learn):
self.epochs, self.learn, self.loss = epochs, learn, tensor(0.)
try:
for cb in self.cbs: cb.set_runner(self)
self('begin_fit')
for epoch in range(epochs):
self.epoch = epoch
if not self('begin_epoch'): self.all_batches(self.data.train_dl)
with torch.no_grad():
if not self('begin_validate'): self.all_batches(self.data.valid_dl)
self('after_epoch')
except CancelTrainException: self('after_cancel_train')
finally:
self('after_fit')
self.train = None
def __call__(self, cb_name):
# return True = Cancel, return False = Continue (Default)
res = False
# check if at least one True return True
for cb in sorted(self.cbs, key=lambda x: x._order): res = res or cb(cb_name)
return res
class Callback():
_order = 0
def set_runner(self, run): self.run = run
def __getattr__(self, k): return getattr(self.run, k)
@property
def name(self):
name = re.sub(r'Callback$', '', self.__class__.__name__)
return camel2snake(name or 'callback')
def __call__(self, cb_name):
f = getattr(self, cb_name, None)
if f and f(): return True
return False
class TrainEvalCallback(Callback):
def begin_fit(self):
self.run.n_epochs = 0.
self.run.n_iter = 0
def begin_epoch(self):
self.run.n_epochs = self.epoch
self.model.train()
self.run.in_train=True
def after_batch(self):
if not self.in_train: return
self.run.n_epochs += 1./self.iters
self.run.n_iter += 1
def begin_validate(self):
self.model.eval()
self.run.in_train=False
class CancelTrainException(Exception): pass
class CancelEpochException(Exception): pass
class CancelBatchException(Exception): pass
```
เราจะเพิ่ม Method Recorder.plot สำหรับพล็อตกราฟ ความสัมพันธ์ระหว่าง Learning Rate กับ Loss
และเพิ่ม Parameter ให้กับ plot_lr, plot_loss ดังนี้
* pgid = Parameter Group ID เราแบ่ง Parameter ของโมเดลออกเป็น 3 กลุ่มที่มี Learning Rate แตกต่างกัน จะอธิบายต่อไป ตอนนี้ให้ใช้ -1 หมายถึงสุดท้าย
* skip_last = จำนวน Iteration ที่ข้ามไม่ต้องพล็อตกราฟ นับจากท้ายสุด เนื่องจาก Iteration ท้าย ๆ มักจะ Loss มาก จนทำให้กราฟดูยาก
```
class Recorder(Callback):
def begin_fit(self):
self.lrs = [[] for _ in self.opt.param_groups]
self.losses = []
def after_batch(self):
if not self.in_train: return
for pg, lr in zip(self.opt.param_groups, self.lrs): lr.append(pg['lr'])
self.losses.append(self.loss.detach().cpu())
def plot_lr(self, pgid=-1): plt.plot(self.lrs[pgid])
def plot_loss(self, skip_last=0): plt.plot(self.losses[:len(self.losses)-skip_last])
def plot(self, skip_last=0, pgid=-1):
losses = [o.item() for o in self.losses]
lrs = self.lrs[pgid]
n = len(losses)-skip_last
plt.xscale('log')
plt.plot(lrs[:n], losses[:n])
```
# 6. Callbacks
## 6.1 LR_Find (Learning Rate Finder Callback)
เราสร้าง Callback ที่ในระหว่างการเทรน จะเพิ่มค่า lr (Learning Rate) ในทุก ๆ Batch แบบ Exponential โดยเริ่มที่ min_lr แต่ไม่เกิน max_lr
และหลังจากเทรนจบ Batch ลองเช็คค่า Loss เปรียบเทียบว่ามากกว่า 10 เท่า ของ ค่า Loss ที่น้อยที่สุด ที่เคยได้หรือไม่ ถ้าใช่ก็แปลว่า Learning Rate มากเกินไปแล้ว จบการเทรนได้ แต่ถ้าไม่ใช่ก็เก็บค่า Loss ที่ดีที่สุดไว้ แล้วเทรนต่อ
max_iter = เราจะเทรนกี่ Iteration (Mini-Batch), min_lr = Learning Rate ขั้นต่ำ, max_lr = เพดาน Learning Rate
```
class LR_Find(Callback):
_order = 1
def __init__(self, max_iter=100, min_lr=1e-6, max_lr=10):
self.max_iter, self.min_lr, self.max_lr = max_iter, min_lr, max_lr
self.best_loss = 1e9
def begin_batch(self):
if not self.in_train: return
pos = self.n_iter/self.max_iter
lr = self.min_lr * (self.max_lr/self.min_lr) ** pos
for pg in self.opt.param_groups: pg['lr'] = lr
def after_loss(self):
if self.n_iter>=self.max_iter or self.loss > self.best_loss*10:
raise CancelTrainException()
if self.loss < self.best_loss: self.best_loss = self.loss
```
# 7. Train
ลองเราลองเทรนด้วย Callback 2 ตัว คือ LR_Find เพื่อหา Learning Rate และ และ Recorder เพื่อบันทึกค่า Learning Rate และ Loss ในระหว่างการเทรน
```
model, loss_func = get_model()
opt = torch.optim.SGD(model.parameters(), lr=lr)
learn = Learner(model, opt, loss_func, data)
run = Runner(cb_funcs=[LR_Find, Recorder])
run.fit(5, learn)
```
# 8. Interpret
พล็อตกราฟ Iteration, Learning Rate ดูว่าในแต่ละ Iteration, Learning Rate นั้น เพิ่มแบบ Exponential ในช่วงระหว่าง min_lr, max_lr
```
run.recorder.plot_lr()
```
พล็อตกราฟความสัมพันธ์ระหว่าง Learing Rate, Loss ดูว่า Loss จะลดลงเร็วขึ้นเรื่อย ๆ เมื่อเราเพิ่ม Learning Rate แต่จะลดลงไปถึงค่าหนึ่ง แล้วจะพุ่งขึ้นอย่างรวดเร็ว
```
run.recorder.plot(skip_last=0)
run.recorder.plot(skip_last=2)
```
ให้เราเลือก Learning Rate ที่ต่ำสุดก่อนที่ Loss จะพุ่งขึ้น 1e-1 ย้อนไป 10 เท่า ในที่นี้คือ lr = 1e-2
เมื่อเราลองเทรนด้วย lr = 1e-2 แล้ว เราสามารถลอง +- 10 เท่าได้ เป็น 1e-1, 1e-3 เพื่อเปรียบเทียบได้
# Credit
* https://course.fast.ai/videos/?lesson=10
* http://yann.lecun.com/exdb/mnist/
```
```
| github_jupyter |
# (Effects of Loans Characteristics on Borrower's APR)
## by (Mohammad Aamir)
## Investigation Overview
> In this investigation, I wanted to look at the characteristics of loan that could be used to predict their borrower APR. The main focus was on borrower prosper rating, original loan ammount and Term.
## Dataset Overview
> The prosper loan data set contains 83982 complete loan data for all loans issued through the 2007–2011, it contains 15 variables. Each row contains information on a loan, including loan amount, BorrowerAPR, borrower rate, Term, borrower income, Investors, and more. 14793 data points were removed from the analysis due to inconsistencies or missing information.
```
# import all packages and set plots to be embedded inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sb
%matplotlib inline
# suppress warnings from final output
import warnings
warnings.simplefilter("ignore")
# load in the dataset into a pandas dataframe
loans_df= pd.read_csv('prosperLoanData.csv')
# selecting the interesting features
loans= loans_df.loc[:,('CreditGrade', 'Term', 'BorrowerAPR','BorrowerRate', 'ProsperRating (Alpha)',
'ListingCategory (numeric)', 'BorrowerState', 'EmploymentStatusDuration', 'IsBorrowerHomeowner',
'AmountDelinquent','LoanOriginalAmount', 'DebtToIncomeRatio', 'IncomeVerifiable','StatedMonthlyIncome',
'Investors','AvailableBankcardCredit','MonthlyLoanPayment')]
#Wrangling
#Make a copy of the data frame
loans_clean = loans.copy()
# fill NA with empty strings
loans_clean['ProsperRating (Alpha)'].fillna("", inplace= True)
loans_clean.CreditGrade.fillna("", inplace= True)
# combine credit ratings
loans_clean['Com_ProsperRating']= loans_clean.CreditGrade+ loans_clean['ProsperRating (Alpha)']
# drop rows with no credit ratings
loans_clean= loans_clean.query('Com_ProsperRating != "" & Com_ProsperRating != "NC"')
# drop unnecessary columns
loans_clean.drop(columns= ['ProsperRating (Alpha)', 'CreditGrade'], inplace= True)
# selecting rows only where ProsperRating is not null as this is one of the most important features of the dataset
# and filling in missing values is not possible.
loans_clean = loans_clean[loans_clean['AmountDelinquent'].notnull()]
loans_clean = loans_clean[loans_clean['BorrowerState'].notnull()]
# filling in missing quantitative values as mean of the columns
loans_clean.BorrowerAPR.fillna(loans_clean.BorrowerAPR.mean(), inplace= True)
loans_clean.EmploymentStatusDuration.fillna(loans_clean.EmploymentStatusDuration.mean(), inplace= True)
loans_clean.DebtToIncomeRatio.fillna(loans_clean.DebtToIncomeRatio.mean(), inplace= True)
loans_clean.AvailableBankcardCredit.fillna(loans_clean.AvailableBankcardCredit.mean(), inplace= True)
#Replace listing category number by name
lits_cat_name= {0: 'Not Available', 1 : 'Debt Consolidation', 2 : 'Home Improvement', 3 : 'Business', 4 : 'Personal Loan',\
5 : 'Student Use', 6 : 'Auto', 7 : 'Other', 8 : 'Baby&Adoption', 9 : 'Boat', 10 : 'Cosmetic Procedure',\
11 : 'Engagement Ring', 12 : 'Green Loans', 13 : 'Household Expenses', 14 : 'Large Purchases',\
15 : 'Medical/Dental', 16 : 'Motorcycle', 17 : 'RV', 18 : 'Taxes', 19 : 'Vacation', 20 : 'Wedding Loans'}
loans_clean['ListingCategory (numeric)']= loans_clean['ListingCategory (numeric)'].map(lits_cat_name)
loans_clean= loans_clean.rename(columns={'ListingCategory (numeric)': 'ListingCategory'})
# Convert ProsperRating to ordinal categorical
ordinal_var_dict= {'Com_ProsperRating': ['HR','E','D','C', 'B', 'A', 'AA']}
for var in ordinal_var_dict:
ordered_var= pd.api.types.CategoricalDtype(ordered= True, categories= ordinal_var_dict[var])
loans_clean[var]= loans_clean[var].astype(ordered_var)
#Adjust datatype for all other categorical columns
loans_clean.BorrowerState= loans_clean.BorrowerState.astype('category')
loans_clean.ListingCategory = loans_clean.ListingCategory.astype('category')
# Getting the lowest and highest value for stated monthly income
Q1 = loans_clean['StatedMonthlyIncome'].quantile(0.25)
Q3 = loans_clean['StatedMonthlyIncome'].quantile(0.75)
IQR = Q3 - Q1
fence_low = Q1-1.5*IQR
fence_high = Q3+1.5*IQR
#Removing the outliers
loans_clean = loans_clean.loc[(loans_clean['StatedMonthlyIncome'] > fence_low) & (loans_clean['StatedMonthlyIncome'] < fence_high)]
loans_clean.shape
```
## (Distribution of Borrower APR)
> We see that the the distribution for Borrower APR are normally distrubuted with the peak between 13 and 23 percent in addition we have some increase in the 35 percent.
```
bins= np.arange(0, loans_clean.BorrowerAPR.max()+0.05, 0.01)
plt.figure(figsize=[10, 7])
plt.hist(data= loans_clean, x= 'BorrowerAPR', bins = bins);
plt.xlabel('Borrower APR');
plt.title('Borrower APR Distribution')
```
## (Borrower APR vs. Prosper Rating)
> The borrower APR decreases with the increasingly better rating. Borrowers with the best Prosper ratings have the lowest APR. It means that the Prosper rating has a strong effect on borrower APR.
```
#Violin plot without datapoints in the violin interior
base_color = sb.color_palette()[0]
plt.figure(figsize=[8, 7])
sb.violinplot(data=loans_clean, x='Com_ProsperRating', y='BorrowerAPR', color=base_color, inner=None)
plt.xlabel('Prosper Rating')
plt.ylabel('Borrower APR')
plt.title('Distribution of BorrowerAPR by Prosper rate')
```
## (Borrower APR vs. Loan Original Amount)
> Negative correlation between LoanOriginalAmount and BrowerAPR, that is the more the loan amount, the lower the APR.
```
plt.figure(figsize=[8, 7])
sb.regplot(data = loans_clean, x = 'LoanOriginalAmount', y = 'BorrowerAPR', x_jitter=0.04, scatter_kws={'alpha':.01})
plt.title('Correlation between Loan original amount and Borrower APR');
```
## (Borrower APR by Prosper rating and Term)
> Interestingly we can see that the borrower APR increase with the decrease of term for people with (B, A, AA) raings. But for people with E rating, the APR decrease with the increase of term.
```
fig = plt.figure(figsize = [8,6])
ax = sb.pointplot(data = loans_clean, x = 'Com_ProsperRating', y = 'BorrowerAPR', hue = 'Term',
palette = 'Blues', linestyles = '', dodge = 0.4, ci='sd')
plt.title('Borrower APR across rating and term')
plt.ylabel('Borrower APR')
ax.set_yticklabels([],minor = True);
```
| github_jupyter |
```
import pandas as pd
import numpy as np
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
import re
from sklearn.model_selection import train_test_split
from sklearn.metrics import *
from sklearn.linear_model import LogisticRegression
import warnings
warnings.filterwarnings('ignore')
import os
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from wcbtfidf import Wcbtfidf
df = pd.read_csv('sentiment140_data.csv',names=('target','id','date','flag','username','tweet'))
df.shape
df.head()
# Checking unique ids
print(df['id'].nunique(),df.shape[0])
# Removing duplicate ids
df.drop_duplicates(subset=['id'],keep='first',inplace=True)
print(df['id'].nunique(),df.shape[0])
# Target data distribution
df['target'].value_counts(normalize=True)
# To test our hypothesis let us convert into an imbalance problem with fewer positive samples
# We will take a total of 5 lakh data points with 4.5 lakh belonging to class 4 and 50k to class 0
negative_samples = df[df['target'] == 0].sample(n=50000,random_state=60)
positive_samples = df[df['target'] == 4].sample(n=450000,random_state=60)
final_df = pd.concat([negative_samples,positive_samples]).sample(frac=1,random_state=60) # A sample operation with full data is
# performed to shuffle the data points
final_df['target'] = final_df['target'].map({0:0,4:1})
final_df['target'].value_counts(normalize=True)
def preprocess_text(text):
text = text.lower()
text = re.sub("[^a-z0-9]"," ",text)
text = re.sub("(\s)+"," ",text)
return text
final_df['clean_text'] = final_df['tweet'].apply(preprocess_text)
print(final_df.shape)
final_df = final_df[['clean_text','target']]
print(final_df.shape)
xtrain,xtest,ytrain,ytest = train_test_split(final_df['clean_text'],final_df['target'],test_size=0.25,random_state=60,stratify=final_df['target'])
print(xtrain.shape,ytrain.shape)
print(xtest.shape,ytest.shape)
# Distribution check in train and test
print(ytrain.value_counts(normalize=True))
print(ytest.value_counts(normalize=True))
def check_hypothesis(xtrain,xtest,ytrain,ytest,max_feat,model):
print('Running base version')
tfidf = TfidfVectorizer(max_features=max_feat,stop_words='english')
train_df = pd.DataFrame(tfidf.fit_transform(xtrain).toarray(),columns=tfidf.vocabulary_)
test_df = pd.DataFrame(tfidf.transform(xtest).toarray(),columns=tfidf.vocabulary_)
model.fit(train_df,ytrain)
preds = model.predict(test_df)
print(f'Precision is {precision_score(ytest,preds)}')
print(f'Recall is {recall_score(ytest,preds)}')
print(f'ROC curve is {roc_auc_score(ytest,preds)}')
print(classification_report(ytest,preds))
print('Running my version')
wcbtfidf = Wcbtfidf(max_features=max_feat)
wcbtfidf.fit(xtrain,ytrain)
train_df = wcbtfidf.transform(xtrain)
test_df = wcbtfidf.transform(xtest)
model.fit(train_df,ytrain)
preds = model.predict(test_df)
print(f'Precision is {precision_score(ytest,preds)}')
print(f'Recall is {recall_score(ytest,preds)}')
print(f'ROC curve is {roc_auc_score(ytest,preds)}')
print(classification_report(ytest,preds))
return wcbtfidf,tfidf
model = LogisticRegression()
wcbtfidf_object,tfidf_object = check_hypothesis(xtrain,xtest,ytrain,ytest,300,model)
```
## ANALYSIS
Negative tweets are the minority class. Let us see whether the vocab of wcbtfidf was able to catch words that cater towards the negative class more as compared to tfidf
```
# Length Comparison
tfidf_vocab = tfidf_object.vocabulary_
wcbtfidf_vocab = wcbtfidf_object.combine_vocab
print(len(wcbtfidf_vocab),len(tfidf_vocab))
# Words that are present in tfidf vocab but not in wcbtfidf
print(list(set(tfidf_vocab) - set(wcbtfidf_vocab)))
```
Major words are neutral and rest are positive like **rock,enjoying,loved,wonderful**
```
# Words that are present in wcbtfidf but not in tfidf
print(list(set(wcbtfidf_vocab) - set(tfidf_vocab)))
```
Here as well there are neutral words but rest are towards the negative end like **stupid,sucks,ugh,hurts,shit,headache,poor,least,missing**
| github_jupyter |
```
import opsimsummary as oss
import os
example_dir = os.path.join(oss.__path__[0], 'example_data')
dbName = os.path.join(example_dir, 'enigma_1189_micro.db')
hdfName = os.path.join(example_dir, 'enigma_1189_micro.hdf')
from sqlalchemy import create_engine as create_engine
from opsimsummary import OpSimOutput
```
# From the OpSim Output (sqlite database)
```
opout_hdf = OpSimOutput.fromOpSimHDF(hdfName=hdfName, subset='Combined')
opout = OpSimOutput.fromOpSimDB(dbname=dbName, subset='_all')
opout.proposalTable
opout.propIDDict
opout.summary.propID.unique()
opout.propIDVals('combined', opout.propIDDict, opout.proposalTable)
opout.propIds
opout.subset
opout.summary.head()
opout.summary[['fieldID', 'fieldRA', 'fieldDec', 'filter', 'expMJD', 'ditheredRA', 'ditheredDec']]
print(len(opout.summary))
# Write to hdf file
opout.writeOpSimHDF('/tmp/opsim_small.hdf')
opsim_from_hdf = OpSimOutput.fromOpSimHDF('/tmp/opsim_small.hdf', subset='_all')
from pandas.util.testing import assert_frame_equal
opsim_from_hdf.subset
assert_frame_equal(opsim_from_hdf.summary, opout.summary)
```
#### From OpSimDB but combined (This is WFD and DDF combined)
```
opsim_comb = OpSimOutput.fromOpSimDB(dbname=dbName)
opsim_comb.writeOpSimHDF('tmp/hdfFile.hdf')
```
### From OpSimHDF
```
opsim_all = OpSimOutput.fromOpSimDB(dbname=dbName, subset='_all')
opsim_all.writeOpSimHDF('test_hdf.hdf')
opsim_fromhdf = OpSimOutput.fromOpSimHDF('test_hdf.hdf', subset='_all')
from sqlalchemy import create_engine
engine = create_engine('sqlite:///'+dbName)
import pandas as pd
_ = pd.read_sql_query('SELECT * FROM Summary WHERE propID in (364, 366) LIMIT 5', con=engine)
_
pids = [364, 366]
x = ', '.join(list(str(pid) for pid in pids))
'SELECT * FROM Summary WHERE propID == ({})'.format(x)
opout_comb = oss.OpSimOutput.fromOpSimDB(dbname=dbName, subset='combined')
opout_comb.summary.propID.unique()
import pandas as pd
summarydf = pd.read_hdf('/tmp/opsim_small.hdf', key='Summary')
from pandas.util.testing import assert_frame_equal
assert_frame_equal(summarydf, opout.summary)
engine = create_engine('sqlite:///' + dbName)
import pandas as pd
pids = tuple([366])
pids = [366, 364]
', '.join('{}'.format(x) for x in pids)
pids = [344, 366]
sql_query = "SELECT * FROM Summary WHERE PROPID == "
sql_query += "({})".format(pid) for pid in pids
sql_query += ", ".join(list("({})".format(pid) for pid in pids))
sql_query
pids
df = pd.read_sql_query(sql_query, con=engine)
df = pd.read_sql_query("SELECT * FROM Summary WHERE PROPID in (362, 366)", con=engine)
df.head()
pids = [366]
df.query('propID == @pids')
df.propID.unique()
df = oss.OpSimOutput.fromOpSimDB(dbName, subset='ddf')
df.summary.head()
pids = df.propIDVals('wfd', df.propIDDict, df.proposalTable)
df.summary.query('propID == {}'.format(pids)).propID.unique()
import sqlite3
engine
alldf = oss.summarize_opsim.OpSimOutput.fromOpSimDB(dbName, subset='_all')
len(alldf.summary.drop_duplicates())
len(alldf.summary.reset_index().drop_duplicates(subset='obsHistID'))== len(alldf.summary)
len(alldf.summary)
unique_alldf = oss.summarize_opsim.OpSimOutput.fromOpSimDB(dbName, subset='unique_all')
len(unique_alldf.summary)
unique_alldf.summary.query('propID == [364, 366]').propID.unique()
def myfunc(*args):
if len
return args[0]**2
myfunc(2)
alldf.
alldf.propIDDict
alldf.summary.head()
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.
# Automated Machine Learning: Explain classification model and visualize the explanation
In this example we use the sklearn's [iris dataset](http://scikit-learn.org/stable/modules/generated/sklearn.datasets.load_iris.html) to showcase how you can use the AutoML Classifier for a simple classification problem.
Make sure you have executed the [configuration](../configuration.ipynb) before running this notebook.
In this notebook you would see
1. Creating an Experiment in an existing Workspace
2. Instantiating AutoMLConfig
3. Training the Model using local compute and explain the model
4. Visualization model's feature importance in widget
5. Explore best model's explanation
## Create Experiment
As part of the setup you have already created a <b>Workspace</b>. For AutoML you would need to create an <b>Experiment</b>. An <b>Experiment</b> is a named object in a <b>Workspace</b>, which is used to run experiments.
```
import logging
import os
import random
import pandas as pd
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.train.automl import AutoMLConfig
from azureml.train.automl.run import AutoMLRun
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = 'automl-local-classification'
# project folder
project_folder = './sample_projects/automl-local-classification-model-explanation'
experiment=Experiment(ws, experiment_name)
output = {}
output['SDK version'] = azureml.core.VERSION
output['Subscription ID'] = ws.subscription_id
output['Workspace Name'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Project Directory'] = project_folder
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
pd.DataFrame(data = output, index = ['']).T
```
## Diagnostics
Opt-in diagnostics for better experience, quality, and security of future releases
```
from azureml.telemetry import set_diagnostics_collection
set_diagnostics_collection(send_diagnostics=True)
```
## Load Iris Data Set
```
from sklearn import datasets
iris = datasets.load_iris()
y = iris.target
X = iris.data
features = iris.feature_names
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X,
y,
test_size=0.1,
random_state=100,
stratify=y)
X_train = pd.DataFrame(X_train, columns=features)
X_test = pd.DataFrame(X_test, columns=features)
```
## Instantiate Auto ML Config
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression|
|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
|**max_time_sec**|Time limit in minutes for each iterations|
|**iterations**|Number of iterations. In each iteration Auto ML trains the data with a specific pipeline|
|**X**|(sparse) array-like, shape = [n_samples, n_features]|
|**y**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]<br>Multi-class targets. An indicator matrix turns on multilabel classification. This should be an array of integers. |
|**X_valid**|(sparse) array-like, shape = [n_samples, n_features]|
|**y_valid**|(sparse) array-like, shape = [n_samples, ], [n_samples, n_classes]|
|**model_explainability**|Indicate to explain each trained pipeline or not |
|**path**|Relative path to the project folder. AutoML stores configuration files for the experiment under this folder. You can specify a new empty folder. |
```
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
primary_metric = 'AUC_weighted',
iteration_timeout_minutes = 200,
iterations = 10,
verbosity = logging.INFO,
X = X_train,
y = y_train,
X_valid = X_test,
y_valid = y_test,
model_explainability=True,
path=project_folder)
```
## Training the Model
You can call the submit method on the experiment object and pass the run configuration. For Local runs the execution is synchronous. Depending on the data and number of iterations this can run for while.
You will see the currently running iterations printing to the console.
```
local_run = experiment.submit(automl_config, show_output=True)
```
## Exploring the results
### Widget for monitoring runs
The widget will sit on "loading" until the first iteration completed, then you will see an auto-updating graph and table show up. It refreshed once per minute, so you should see the graph update as child runs complete.
NOTE: The widget displays a link at the bottom. This links to a web-ui to explore the individual run details.
```
from azureml.widgets import RunDetails
RunDetails(local_run).show()
```
### Retrieve the Best Model
Below we select the best pipeline from our iterations. The *get_output* method on automl_classifier returns the best run and the fitted model for the last *fit* invocation. There are overloads on *get_output* that allow you to retrieve the best run and fitted model for *any* logged metric or a particular *iteration*.
```
best_run, fitted_model = local_run.get_output()
print(best_run)
print(fitted_model)
```
### Best Model 's explanation
Retrieve the explanation from the best_run. And explanation information includes:
1. shap_values: The explanation information generated by shap lib
2. expected_values: The expected value of the model applied to set of X_train data.
3. overall_summary: The model level feature importance values sorted in descending order
4. overall_imp: The feature names sorted in the same order as in overall_summary
5. per_class_summary: The class level feature importance values sorted in descending order. Only available for the classification case
6. per_class_imp: The feature names sorted in the same order as in per_class_summary. Only available for the classification case
```
from azureml.train.automl.automlexplainer import retrieve_model_explanation
shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \
retrieve_model_explanation(best_run)
print(overall_summary)
print(overall_imp)
print(per_class_summary)
print(per_class_imp)
```
Beside retrieve the existed model explanation information, explain the model with different train/test data
```
from azureml.train.automl.automlexplainer import explain_model
shap_values, expected_values, overall_summary, overall_imp, per_class_summary, per_class_imp = \
explain_model(fitted_model, X_train, X_test)
print(overall_summary)
print(overall_imp)
```
| github_jupyter |
# 6. Kenel Methods
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from prml.kernel import (
PolynomialKernel,
RBF,
GaussianProcessClassifier,
GaussianProcessRegressor
)
def create_toy_data(func, n=10, std=1., domain=[0., 1.]):
x = np.linspace(domain[0], domain[1], n)
t = func(x) + np.random.normal(scale=std, size=n)
return x, t
def sinusoidal(x):
return np.sin(2 * np.pi * x)
```
## 6.1 Dual Representation
```
x_train, y_train = create_toy_data(sinusoidal, n=10, std=0.1)
x = np.linspace(0, 1, 100)
model = GaussianProcessRegressor(kernel=PolynomialKernel(3, 1.), beta=int(1e10))
model.fit(x_train, y_train)
y = model.predict(x)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", color="blue", label="training")
plt.plot(x, sinusoidal(x), color="g", label="sin$(2\pi x)$")
plt.plot(x, y, color="r", label="gpr")
plt.show()
```
## 6.4 Gaussian Processes
### 6.4.2 Gaussian processes for regression
```
x_train, y_train = create_toy_data(sinusoidal, n=7, std=0.1, domain=[0., 0.7])
x = np.linspace(0, 1, 100)
model = GaussianProcessRegressor(kernel=RBF(np.array([1., 15.])), beta=100)
model.fit(x_train, y_train)
y, y_std = model.predict(x, with_error=True)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", color="blue", label="training")
plt.plot(x, sinusoidal(x), color="g", label="sin$(2\pi x)$")
plt.plot(x, y, color="r", label="gpr")
plt.fill_between(x, y - y_std, y + y_std, alpha=0.5, color="pink", label="std")
plt.show()
```
### 6.4.3 Learning the hyperparameters
```
x_train, y_train = create_toy_data(sinusoidal, n=7, std=0.1, domain=[0., 0.7])
x = np.linspace(0, 1, 100)
plt.figure(figsize=(20, 5))
plt.subplot(1, 2, 1)
model = GaussianProcessRegressor(kernel=RBF(np.array([1., 1.])), beta=100)
model.fit(x_train, y_train)
y, y_std = model.predict(x, with_error=True)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", color="blue", label="training")
plt.plot(x, sinusoidal(x), color="g", label="sin$(2\pi x)$")
plt.plot(x, y, color="r", label="gpr {}".format(model.kernel.params))
plt.fill_between(x, y - y_std, y + y_std, alpha=0.5, color="pink", label="std")
plt.legend()
plt.subplot(1, 2, 2)
model.fit(x_train, y_train, iter_max=100)
y, y_std = model.predict(x, with_error=True)
plt.scatter(x_train, y_train, facecolor="none", edgecolor="b", color="blue", label="training")
plt.plot(x, sinusoidal(x), color="g", label="sin$(2\pi x)$")
plt.plot(x, y, color="r", label="gpr {}".format(np.round(model.kernel.params, 2)))
plt.fill_between(x, y - y_std, y + y_std, alpha=0.5, color="pink", label="std")
plt.legend()
plt.show()
```
### 6.4.4 Automatic relevance determination
```
def create_toy_data_3d(func, n=10, std=1.):
x0 = np.linspace(0, 1, n)
x1 = x0 + np.random.normal(scale=std, size=n)
x2 = np.random.normal(scale=std, size=n)
t = func(x0) + np.random.normal(scale=std, size=n)
return np.vstack((x0, x1, x2)).T, t
x_train, y_train = create_toy_data_3d(sinusoidal, n=20, std=0.1)
x0 = np.linspace(0, 1, 100)
x1 = x0 + np.random.normal(scale=0.1, size=100)
x2 = np.random.normal(scale=0.1, size=100)
x = np.vstack((x0, x1, x2)).T
model = GaussianProcessRegressor(kernel=RBF(np.array([1., 1., 1., 1.])), beta=100)
model.fit(x_train, y_train)
y, y_std = model.predict(x, with_error=True)
plt.figure(figsize=(20, 5))
plt.subplot(1, 2, 1)
plt.scatter(x_train[:, 0], y_train, facecolor="none", edgecolor="b", label="training")
plt.plot(x[:, 0], sinusoidal(x[:, 0]), color="g", label="$\sin(2\pi x)$")
plt.plot(x[:, 0], y, color="r", label="gpr {}".format(model.kernel.params))
plt.fill_between(x[:, 0], y - y_std, y + y_std, color="pink", alpha=0.5, label="gpr std.")
plt.legend()
plt.ylim(-1.5, 1.5)
model.fit(x_train, y_train, iter_max=100, learning_rate=0.001)
y, y_std = model.predict(x, with_error=True)
plt.subplot(1, 2, 2)
plt.scatter(x_train[:, 0], y_train, facecolor="none", edgecolor="b", label="training")
plt.plot(x[:, 0], sinusoidal(x[:, 0]), color="g", label="$\sin(2\pi x)$")
plt.plot(x[:, 0], y, color="r", label="gpr {}".format(np.round(model.kernel.params, 2)))
plt.fill_between(x[:, 0], y - y_std, y + y_std, color="pink", alpha=0.2, label="gpr std.")
plt.legend()
plt.ylim(-1.5, 1.5)
plt.show()
```
### 6.4.5 Gaussian processes for classification
```
def create_toy_data():
x0 = np.random.normal(size=50).reshape(-1, 2)
x1 = np.random.normal(size=50).reshape(-1, 2) + 2.
return np.concatenate([x0, x1]), np.concatenate([np.zeros(25), np.ones(25)]).astype(np.int)[:, None]
x_train, y_train = create_toy_data()
x0, x1 = np.meshgrid(np.linspace(-4, 6, 100), np.linspace(-4, 6, 100))
x = np.array([x0, x1]).reshape(2, -1).T
model = GaussianProcessClassifier(RBF(np.array([1., 7., 7.])))
model.fit(x_train, y_train)
y = model.predict(x)
plt.scatter(x_train[:, 0], x_train[:, 1], c=y_train)
plt.contourf(x0, x1, y.reshape(100, 100), levels=np.linspace(0,1,3), alpha=0.2)
plt.colorbar()
plt.xlim(-4, 6)
plt.ylim(-4, 6)
plt.gca().set_aspect('equal', adjustable='box')
```
| github_jupyter |
#### Part 3 Instructions
#### Create a Travel Itinerary with a Corresponding Map
```
# Import the dependencies.
import pandas as pd
import gmaps
import requests
import os
import time
import json
from datetime import datetime
# Import the API key.
from config import g_key
# Configure gmaps to use your Google API key.
gmaps.configure(api_key=g_key)
# read data from csv and store into dta frame
file_name = os.path.join(".","data","WeatherPy_vacation.csv")
weather_vacation_data_df = pd.read_csv(file_name)
weather_vacation_data_df.head()
info_box_template = """
<dl>
<dt>Hotel Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
<dt>Current Weather</dt><dd>{Current Description} and {Max Temp} °F</dd>
</dl>
"""
# Store the DataFrame Row.
hotel_info = [info_box_template.format(**row) for index, row in weather_vacation_data_df.iterrows()]
# Add a heatmap of temperature for the vacation spots.
locations = weather_vacation_data_df[["Lat", "Lng"]]
# max_temp = hotel_df["Max Temp"]
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
# hotel marker with names
marker_layer = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig.add_layer(marker_layer)
# Call the figure to plot the data.
fig
city_name1 = input("Please enter first city name, you can click on merket to get city name: ")
city_name2 = input("Please enter second city name, you can click on merket to get city name: ")
city_name3 = input("Please enter third city name, you can click on merket to get city name: ")
city_name4 = input("Please enter fourth city name, you can click on merket to get city name: ")
print(f'1:{city_name1}, 2:{city_name2}, 3:{city_name3}, 4:{city_name4}')
# Get infor for each entered city
vacation_stop_1 = weather_vacation_data_df.loc[weather_vacation_data_df["City"] == city_name1]
vacation_stop_2 = weather_vacation_data_df.loc[weather_vacation_data_df["City"] == city_name2]
vacation_stop_3 = weather_vacation_data_df.loc[weather_vacation_data_df["City"] == city_name3]
vacation_stop_4 = weather_vacation_data_df.loc[weather_vacation_data_df["City"] == city_name4]
print(f"1:{vacation_stop_1} \n 2:{vacation_stop_2} \n 3:{vacation_stop_3}\n 4:{vacation_stop_4}")
# Getting Lat and Lg for each city and storing it as tuple
city_1_lat_lng = vacation_stop_1["Lat"].values[0], vacation_stop_1["Lng"].values[0]
city_2_lat_lng = vacation_stop_2["Lat"].values[0], vacation_stop_2["Lng"].values[0]
city_3_lat_lng = vacation_stop_3["Lat"].values[0], vacation_stop_3["Lng"].values[0]
city_4_lat_lng = vacation_stop_4["Lat"].values[0], vacation_stop_4["Lng"].values[0]
print(f'1:{city_1_lat_lng}, 2:{city_2_lat_lng}, 3:{city_3_lat_lng}, 4:{city_4_lat_lng}')
# Get the directions
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
travel_itinerary = gmaps.directions_layer(city_1_lat_lng, city_4_lat_lng,
waypoints=[city_2_lat_lng, city_3_lat_lng],travel_mode='DRIVING')
fig.add_layer(travel_itinerary)
fig
type(vacation_stop_1)
# Create a marker layer map for the four cities
# frames = [df1, df2, df3]
# In [5]: result = pd.concat(frames)
# combine 4 dataframes
frames = [vacation_stop_1, vacation_stop_2, vacation_stop_3, vacation_stop_4]
results_df = pd.concat(frames)
results_df
info_box_template = """
<dl>
<dt>Hotel Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
<dt>Current Weather</dt><dd>{Current Description} and {Max Temp} °F</dd>
</dl>
"""
# Store the DataFrame Row.
itinerary_hotel_info = [info_box_template.format(**row) for index, row in results_df.iterrows()]
# marker layer map
locations = results_df[["Lat", "Lng"]]
# max_temp = hotel_df["Max Temp"]
fig = gmaps.figure(center=(30.0, 31.0), zoom_level=1.5)
# hotel marker with names
marker_layer = gmaps.marker_layer(locations, info_box_content=itinerary_hotel_info)
fig.add_layer(marker_layer)
# Call the figure to plot the data.
fig
```
| github_jupyter |
```
import json
# !wget https://f000.backblazeb2.com/file/malay-dataset/chatbot/blended-skill-talk/blended_skill_talk.json.translate
import re
def cleaning(string):
string = string.replace('\n', ' ')
string = re.sub(r'[ ]+', ' ', string).strip()
return string
with open('blended_skill_talk.json.translate') as fopen:
data = json.load(fopen)
before, after = [], []
for i in data:
if len(i) != 2:
continue
text = i[1]
splitted = text.split('[EENND]')
if len(splitted) != 2:
continue
l, r = splitted
l = cleaning(l)
r = cleaning(r)
before.append(l)
after.append(r)
!wget https://f000.backblazeb2.com/file/malay-dataset/chatbot/convai2/convai2-0.json.translate
!wget https://f000.backblazeb2.com/file/malay-dataset/chatbot/convai2/convai2-100000.json.translate
len(before), len(after)
with open('convai2-0.json.translate') as fopen:
data = json.load(fopen)
for i in data:
if len(i) != 2:
continue
text = i[1]
splitted = text.split('[EENND]')
if len(splitted) == 3:
splitted = splitted[1:]
if len(splitted) != 2:
continue
l, r = splitted
l = cleaning(l)
r = cleaning(r)
before.append(l)
after.append(r)
with open('convai2-100000.json.translate') as fopen:
data = json.load(fopen)
for i in data:
if len(i) != 2:
continue
text = i[1]
splitted = text.split('[EENND]')
if len(splitted) == 3:
splitted = splitted[1:]
if len(splitted) != 2:
continue
l, r = splitted
l = cleaning(l)
r = cleaning(r)
before.append(l)
after.append(r)
len(before), len(after)
list(zip(before[-10:], after[-10:]))
!wget https://f000.backblazeb2.com/file/malay-dataset/wiki-wizard/informations-0.json.translate
!wget https://f000.backblazeb2.com/file/malay-dataset/wiki-wizard/informations-100000.json.translate
!wget https://f000.backblazeb2.com/file/malay-dataset/wiki-wizard/informations-200000.json.translate
from glob import glob
files = glob('informations-*.json.translate')
for file in files:
print(file)
with open(file) as fopen:
data = json.load(fopen)
for i in data:
if len(i) != 2:
continue
splitted = i[1].split('[EENND]')
if len(splitted) != 3:
continue
l, r = splitted[1:]
l = cleaning(l)
r = cleaning(r)
before.append(l)
after.append(r)
list(zip(before[-10:], after[-10:]))
import json
import tensorflow as tf
import itertools
filename = 'chatbot.tsv'
with tf.io.gfile.GFile(filename, 'w') as outfile:
for i in range(len(before)):
outfile.write('%s\t%s\n' % (before[i], after[i]))
import tensorflow as tf
import tensorflow_datasets as tfds
from t5.data import preprocessors as prep
import functools
import t5
import gin
import sentencepiece as spm
from glob import glob
import os
gin.parse_config_file('pretrained_models_base_operative_config.gin')
vocab = 'sp10m.cased.t5.model'
sp = spm.SentencePieceProcessor()
sp.Load(vocab)
def chatbot_dataset(split, shuffle_files = False):
del shuffle_files
ds = tf.data.TextLineDataset(
['chatbot.tsv']
)
ds = ds.map(
functools.partial(
tf.io.decode_csv,
record_defaults = ['', ''],
field_delim = '\t',
use_quote_delim = False,
),
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
ds = ds.map(lambda *ex: dict(zip(['question', 'answer'], ex)))
return ds
def chatbot_preprocessor(ds):
def to_inputs_and_targets(ex):
return {
'inputs': tf.strings.join(['chatbot: ', ex['question']]),
'targets': ex['answer'],
}
return ds.map(
to_inputs_and_targets,
num_parallel_calls = tf.data.experimental.AUTOTUNE,
)
t5.data.TaskRegistry.remove('chatbot_dataset')
t5.data.TaskRegistry.add(
'chatbot_dataset',
dataset_fn = chatbot_dataset,
splits = ['train'],
text_preprocessor = [chatbot_preprocessor],
sentencepiece_model_path = vocab,
metric_fns = [t5.evaluation.metrics.accuracy],
)
from tqdm import tqdm
nq_task = t5.data.TaskRegistry.get("chatbot_dataset")
ds = nq_task.get_dataset(split='qa.tsv', sequence_length={"inputs": 768, "targets": 768})
batch_size, index, part = 50000, 0, 0
fopen = open(f'chatbot-{part}.parse', 'w')
for ex in tqdm(tfds.as_numpy(ds)):
i = sp.DecodeIds(ex['inputs'].tolist())
t = sp.DecodeIds(ex['targets'].tolist())
text = f'{i} [[EENNDD]] {t}\n'
fopen.write(text)
if index == batch_size:
fopen.close()
part += 1
index = 0
fopen = open(f'chatbot-{part}.parse', 'w')
index += 1
fopen.close()
```
| github_jupyter |
```
# Softmax classifier for guessing minesweeper board position and whether it has a mine or not
# Import libraries for simulation
import tensorflow as tf
import numpy as np
import random as r
import datetime as dt
dimensions = (12,12)
mineProbability = 0.16 # Probability that a square contain a mine
# Clears a square on the minesweeper board.
# If it had a mine, return true
# Otherwise if it has no adjacent mines, recursively run on adjacent squares
# Return false
def clearSquare(board,adjacency,row,col):
if board[row,col] == 1:
return True
if adjacency[row,col] >= 0:
return False
n = 0
for r in range(row-1,row+2):
for c in range(col-1,col+2):
if 0 <= r and r < rows and 0 <= c and c < cols:
n += board[r,c]
adjacency[row,col] = n
if n == 0:
for r in range(row-1,row+2):
for c in range(col-1,col+2):
if 0 <= r and r < rows and 0 <= c and c < cols:
clearSquare(board,adjacency,r,c)
return False
# This takes a mine board and gives a mine count with mines removed, and other random squares removed
def boardPartialMineCounts(board):
clearProbability = r.uniform(0.05,0.5)
result = np.full(dimensions,-1)
for index, x in np.ndenumerate(board):
row,col = index
if not(x) and result[row,col] == -1 and r.uniform(0,1) < clearProbability:
clearSquare(board,result,row,col)
return result
# Generates a random training batch of size at most n
def next_training_batch(n):
batch_xs = []
batch_ys = []
boards = []
for _ in range(n):
board = np.random.random(dimensions) < mineProbability
counts = boardPartialMineCounts(board)
validGuesses = np.append(((counts == -1).astype(int) - board).flatten().astype(float),
board.flatten().astype(float))
validGuessesSum = sum(validGuesses)
if validGuessesSum > 0:
# encode counts as one hot
countsOneHot = np.zeros((counts.size,10))
countsOneHot[np.arange(counts.size), counts.flatten() + 1] = 1
batch_xs.append(countsOneHot.flatten())
batch_ys.append(validGuesses / validGuessesSum)
boards.append(board)
return (np.asarray(batch_xs), np.asarray(batch_ys), boards)
# Create the model
rows, cols = dimensions
size = rows*cols
mineCountsOneHot = tf.placeholder(tf.float32, [None, size*10], name="mineCountsOneHot")
#mineCountsOneHot = tf.reshape(tf.one_hot(mineCounts+1,10), [-1, size*10])
W = tf.Variable(tf.random_normal([size*10, size*2], stddev=0.01), name="W")
b = tf.Variable(tf.random_normal([size*2], stddev=0.01), name="b")
y = tf.matmul(mineCountsOneHot, W) + b
validGuessAverages = tf.placeholder(tf.float32, [None, size*2], name="validGuessAverages")
# Loss function
cross_entropy = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(labels=validGuessAverages, logits=y))
# Summaries for tensorboard
with tf.name_scope('W_reshape'):
image_shaped_W = tf.reshape(W, [-1, size*10, size*2, 1])
tf.summary.image('W', image_shaped_W, 1000)
with tf.name_scope('b_reshape'):
image_shaped_b = tf.reshape(b, [-1, rows*2, cols, 1])
tf.summary.image('b', image_shaped_b, 1000)
_ = tf.summary.scalar('loss', cross_entropy)
# Optimiser
train_step = tf.train.AdamOptimizer().minimize(cross_entropy)
# Create session and initialise or restore stuff
savePath = './saves.tf.Mines6/' + str(dimensions) + '/'
saver = tf.train.Saver()
sess = tf.InteractiveSession()
merged = tf.summary.merge_all()
writer = tf.summary.FileWriter('.', sess.graph)
tf.global_variables_initializer().run()
# Restore model?
#saver.restore(sess, savePath + "model-2000")
# Train
for iteration in range(10001):
batch_xs, batch_ys, _ = next_training_batch(1000)
summary, loss, _ = sess.run([merged, cross_entropy, train_step],
feed_dict={mineCountsOneHot: batch_xs, validGuessAverages: batch_ys})
writer.add_summary(summary, iteration)
print('%s: Loss at step %s: %s' % (dt.datetime.now().strftime("%Y-%m-%d %H:%M:%S"), iteration, loss))
if iteration % 100 == 0:
save_path = saver.save(sess, savePath + 'model', global_step=iteration)
print("Model saved in file: %s" % save_path)
# Test trained model on larger batch size
batch_xs, batch_ys, _ = next_training_batch(10000)
print(sess.run(cross_entropy, feed_dict={mineCountsOneHot: batch_xs, validGuessAverages: batch_ys}))
# Run a test
batchSize = 10000
batch_xs, batch_ys, _ = next_training_batch(batchSize)
predictions = sess.run(tf.nn.softmax(y), feed_dict={mineCountsOneHot: batch_xs, validGuessAverages: batch_ys})
bestSquares = [pred.argmax() for pred in predictions]
unfrees = (batch_ys == 0).astype(int)
frees = [unfrees[i][bestSquares[i]] for i in range(batchSize)]
print("Number of errors for batch size of ", batchSize)
print(sum(frees))
# Find boards that we failed on
batchSize = 1000
batch_xs, batch_ys, _ = next_training_batch(batchSize)
predictions = sess.run(tf.nn.softmax(y), feed_dict={mineCountsOneHot: batch_xs, validGuessAverages: batch_ys})
bestSquares = [pred.argmax() for pred in predictions]
unfrees = (batch_ys == 0).astype(int)
guesses = [unfrees[i][bestSquares[i]] for i in range(batchSize)]
for i in range(batchSize):
if guesses[i] == 1:
print(batch_xs[i].reshape(dimensions))
summary = sess.run(tf.summary.image('mine_miss', tf.reshape((batch_xs[i]+1).astype(float),[-1,rows,cols,1]), 100))
writer.add_summary(summary)
#batch_xs = [[-1,1,-1,0,0,0,-1,-1,-1,1,1,1,-1,-1,1,2,2,1,1,-1,2,1,1,-1,-1,2,2,-1,-1,2,-1,1,1,0,-1,-1,2,-1,-1,4,-1,2,-1,1,2,-1,1,0,1,2,-1,3,2,2,1,-1,2,-1,1,0,0,1,1,-1,-1,-1,-1,-1,-1,1,1,-1,-1,0,0,3,-1,4,1,2,-1,1,-1,-1,0,0,0,2,-1,-1,-1,2,-1,1,0,0,0,-1,1,2,-1,2,1,2,2,3,3,2,-1,-1,1,-1,1,-1,0,1,2,-1,-1,-1,1,1,1,-1,1,0,-1,-1,-1,-1,-1,-1,-1,-1,0,-1,-1,-1,-1,-1,1,-1,-1,-1]]
batch_xs0 = [-1] * (size)
batch_xs0[0] = 1
batch_xs0[1] = 1
batch_xs0[cols] = 1
predictions = sess.run(tf.nn.softmax(y), feed_dict={mineCountsOneHot: [batch_xs0]})
bestSquares = [pred.argmax() for pred in predictions]
print(bestSquares[0] // cols, bestSquares[0] % cols)
np.save("./W", sess.run(W))
np.save("./b", sess.run(b))
np.savez("./model", sess.run([W,b]))
```
| github_jupyter |
### Installation
```
# if (!requireNamespace("BiocManager", quietly = TRUE))
# install.packages("BiocManager")
# BiocManager::install("chromVAR", version = "3.8")
# BiocManager::install("motifmatchr", version = "3.8")
# BiocManager::install("BSgenome.Hsapiens.UCSC.hg19", version = "3.8")
# BiocManager::install("JASPAR2016", version = "3.8")
```
### Import packages
```
library(chromVAR)
library(motifmatchr)
library(Matrix)
library(SummarizedExperiment)
library(BiocParallel)
library('JASPAR2016')
library(BSgenome.Hsapiens.UCSC.hg19)
packageVersion("chromVAR")
register(MulticoreParam(10))
```
### Obtain Feature Matrix
```
start_time <- Sys.time()
set.seed(2019)
metadata <- read.table('../../input/metadata.tsv',
header = TRUE,
stringsAsFactors=FALSE,quote="",row.names=1)
peakfile <- "../../input/combined.sorted.merged.bed"
peaks <- getPeaks(peakfile, sort_peaks = TRUE)
```
`width=500` will cause the error: Error in loadFUN(x, seqname, ranges): trying to load regions beyond the boundaries of non-circular sequence "chr17"
```
peaks <- resize(peaks, width = 450, fix = "center")
bamfile <- list.files(path = "../../input/sc-bams_nodup/", pattern = "\\.bam$")
length(bamfile)
cellnames <- sapply(strsplit(bamfile,'.',fixed = TRUE), "[[", 1)
head(cellnames)
sum(cellnames == rownames(metadata))
fragment_counts <- getCounts(paste0("../../input/sc-bams_nodup/",bamfile),
peaks,
paired = TRUE,
by_rg = TRUE,
format = "bam",
colData = data.frame(celltype = cellnames))
fragment_counts
fragment_counts <- addGCBias(fragment_counts, genome = BSgenome.Hsapiens.UCSC.hg19)
head(rowData(fragment_counts))
counts_filtered <- filterPeaks(fragment_counts, non_overlapping = TRUE)
bg <- getBackgroundPeaks(counts_filtered)
# Potentially save the bg object
saveRDS(bg, file = "background_peaks_kmers.rds")
kmer_ix <- matchKmers(6, counts_filtered, genome = BSgenome.Hsapiens.UCSC.hg19)
dev <- computeDeviations(object = counts_filtered, annotations = kmer_ix,
background_peaks = bg)
end_time <- Sys.time()
end_time - start_time
df_zscores = dev@assays[[1]]
dim(df_zscores)
df_zscores[1:5,1:5]
sum(colnames(df_zscores) == rownames(metadata))
saveRDS(df_zscores, file = '../../output/feature_matrices/FM_chromVAR_buenrostro2018_kmers.rds')
```
### Downstream Analysis
```
variability <- computeVariability(dev)
plotVariability(variability, use_plotly = FALSE)
head(variability)
dim(variability)
dev$celltype <- metadata[colnames(dev),]
tsne_results <- deviationsTsne(dev, threshold = 1.5, perplexity = 10)
tsne_plots <- plotDeviationsTsne(dev, tsne_results,
sample_column = "celltype",
shiny = FALSE)
tsne_plots[[1]]
sessionInfo()
save.image(file = 'chromVAR_buenrostro2018_kmers.RData')
```
| github_jupyter |
# Reading and Writing Audio Files with (scikits.)audiolab
[back to overview page](index.ipynb)
http://cournape.github.io/audiolab/
This has support for a lot of formats, but you may have to install it manually.
Works with pcm16, pcm24, pcm32, float32 (both WAV and WAVEX).
Audio data is returned as NumPy array, default `dtype` is `float64`.
This can be changed with the `dtype=` argument, but there is currently (in version 0.11.0) a bug (see <https://github.com/cournape/audiolab/issues/3>)!
Advantages:
* based on [libsndfile](http://www.mega-nerd.com/libsndfile/), can read WAV, OGG, FLAC, ...
* see `audiolab.available_file_formats()` and `audiolab.available_encodings()`
* can read all variants of WAV files (including 24-bit and WAVEX)
* can read parts of audio files
* automatic type conversion and normalization
Disadvantages:
* in most cases, it has to be installed manually (but it's not too hard if you have the necessary dependencies installed)
* there is a bug if `dtype != float64`
* needs NumPy (which normally isn't a problem)
* **doesn't work on Python 3**
Installation:
python2 -m pip install scikits.audiolab
```
import matplotlib.pyplot as plt
import numpy as np
```
## Reading
```
from scikits.audiolab import Sndfile, Format
import contextlib
with contextlib.closing(Sndfile('data/test_wav_pcm16.wav')) as f:
print "sampling rate = {} Hz, length = {} samples, channels = {}".format(f.samplerate, f.nframes, f.channels)
# default dtype: float64
sig = f.read_frames(f.nframes)
print "dtype: {}".format(sig.dtype)
plt.plot(sig);
```
32-bit float and WAVEX are supported:
```
with contextlib.closing(Sndfile('data/test_wavex_float32.wav')) as f:
sig = f.read_frames(f.nframes)
print "dtype: {}".format(sig.dtype)
plt.plot(sig);
```
But, if another type than `float64` is requested, things go wrong!
... at least in version 0.11.0, see https://github.com/cournape/audiolab/issues/3.
```
with contextlib.closing(Sndfile('data/test_wav_pcm16.wav')) as f:
sig = f.read_frames(f.nframes, dtype=np.float32)
plt.plot(sig);
```
Nice format information:
```
Sndfile('data/test_wavex_float32.wav').format
```
## Writing
TODO
## Version
```
from scikits import audiolab
audiolab.__version__
```
<p xmlns:dct="http://purl.org/dc/terms/">
<a rel="license"
href="http://creativecommons.org/publicdomain/zero/1.0/">
<img src="http://i.creativecommons.org/p/zero/1.0/88x31.png" style="border-style: none;" alt="CC0" />
</a>
<br />
To the extent possible under law,
<span rel="dct:publisher" resource="[_:publisher]">the person who associated CC0</span>
with this work has waived all copyright and related or neighboring
rights to this work.
</p>
| github_jupyter |
# Joint Probability, Conditional Probability and Bayes' Rule
```
#Import packages
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
%matplotlib inline
sns.set()
```
## Learning Objectives of Part 1-b
- To understand and be able to simulate joint probabilities and conditional probabilities;
- To understand Bayes' Theorem and its utility.
## Joint Probability & Conditional Probability
### Joint Probability
We have already encountered joint probabilities in the previous notebook, perhaps without knowing it: $P(A,B)$ is the probability two events $A$ and $B$ _both_ occurring.
* For example, getting two heads in a row.
If $A$ and $B$ are independent, then $P(A,B)=P(A)P(B)$ but be warned: this is not always (or often) the case.
One way to think of this is considering "AND" as multiplication: the probability of A **and** B is the probability of A **multiplied** by the probability of B.
#### Hands-On: Joint Probability and Coin Flipping
Verify that $P(A,B)=P(A)P(B)$ in the two fair coin-flip case (A=heads, B=heads) by
- first simulating two coins being flipped together and calculating the proportion of occurences with two heads;
- then simulating one coin flip and calculating the proportion of heads and then doing that again and multiplying the two proportions.
Your two calculations should give "pretty close" results and not the same results due to the (in)accuracy of simulation.
```
# Solution: Calculate P(A,B)
x_0 = np.random.binomial(2, 0.5, 10000)
p_ab = ______________
# Now, plot the histogram of the results
plt.hist(x_0);
print(p_ab)
# Solution: Calculate P(A)P(B)
x_1 = np.random.binomial(1, 0.5, 10000)
x_2 = np.random.binomial(1, 0.5, 10000)
p_a = ___
p_b = ___
p_a*p_b
```
**Note:** In order to use such simulation and _hacker statistics_ approaches to "prove" results such as the above, we're gliding over several coupled and deep technicalities. This is in the interests of the pedagogical nature of this introduction. For the sake of completeness, we'll mention that we're essentially
- Using the proportion in our simulations as a proxy for the probability (which, although Frequentist, is useful to allow you to start getting your hands dirty with probability via simluation).
Having stated this, for ease of instruction, we'll continue to do so when thinking about joint & conditional probabilities of both simulated and real data.
#### Hands-On: Joint probability for birds
What is the probability that two randomly selected birds have beak depths over 10 ?
```
# Import data & store lengths in a pandas series
df_12 = pd.read_csv('../data/finch_beaks_2012.csv')
lengths = df_12['blength']
# Calculate P(A)P(B) of two birds having beak lengths > 10
p_a = ___
p_b = ___
p_a*p_b
```
* Calculate the joint probability using the resampling method, that is, by drawing random samples (with replacement) from the data. First calculate $P(A)P(B)$:
```
# Calculate P(A)P(B) using resampling methods
n_samples = 100000
p_a = sum(___)/n_samples
p_b = sum(___)/n_samples
p_a*p_b
```
Now calculate $P(A,B)$:
```
# Calculate P(A,B) using resampling methods
n_samples = 100000
samples = ___
_ = samples > (10, 10)
p_ab = sum(___)/n_samples
p_ab
```
**Task:** Interpret the results of your simulations.
### Conditional Probability
Now that we have a grasp on joint probabilities, lets consider conditional probabilities, that is, the probability of some $A$, knowing that some other $B$ is true. We use the notation $P(A|B)$ to denote this. For example, you can ask the question "What is the probability of a finch beak having depth $<10$, knowing that the finch of of species 'fortis'?"
#### Example: conditional probability for birds
1. What is the probability of a finch beak having depth > 10 ?
2. What if we know the finch is of species 'fortis'?
3. What if we know the finch is of species 'scandens'?
```
# Q1 Answer
___
# Q2 Answer
df_fortis = df_12.loc[df_12['species'] == 'fortis']
___
# Q3 Answer
df_scandens = df_12.loc[df_12['species'] == 'scandens']
___
```
**Note:** These proportions are definitely different. We can't say much more currently but we'll soon see how to use hypothesis testing to see what else we can say about the differences between the species of finches.
### Joint and conditional probabilities
Conditional and joint probabilites are related by the following:
$$ P(A,B) = P(A|B)P(B)$$
**Homework exercise for the avid learner:** verify the above relationship using simulation/resampling techniques in one of the cases above.

### Hands on example: drug testing
**Question:** Suppose that a test for using a particular drug is 99% sensitive and 99% specific. That is, the test will produce 99% true positive results for drug users and 99% true negative results for non-drug users. Suppose that 0.5% (5 in 1,000) of people are users of the drug. What is the probability that a randomly selected individual with a positive test is a drug user?
**If we can answer this, it will be really cool as it shows how we can move from knowing $P(+|user)$ to $P(user|+)$, a MVP for being able to move from $P(data|model)$ to $P(model|data)$.**
In the spirit of this workshop, it's now time to harness your computational power and the intuition of simulation to solve this drug testing example.
* Before doing so, what do you think the answer to the question _"What is the probability that a randomly selected individual with a positive test is a drug user?"_ is? Write down your guess.
```
# Take 10,000 subjects
n = 100000
# Sample for number of users, non-users
users = ___
non_users = ___
# How many of these users tested +ve ?
u_pos = ___
# How many of these non-users tested +ve ?
non_pos = ___
# how many of those +ve tests were for users?
_____ / (______ + _________)
```
**Discussion**: What you have been able to do here is to solve the following problem: you knew $P(+|user)=0.99$, but you were trying to figure out $P(user|+)$. Is the answer what you expected? If not, why not?
**Key note:** This is related to the serious scientific challenge posed at the beginning here: if you know the underlying parameters/model, you can figure out the distribution and the result, but often we have only the experimental result and we're trying to figure out the most appropriate model and parameters.
It is Bayes' Theorem that lets us move between these.
## 2. Bayes' Theorem
$$P(B|A) = \frac{P(A|B)P(B)}{P(A)}$$
As you may have guessed, it is Bayes' Theorem that will allow us to move back and forth between $P(data|model)$ and $P(model|data)$. As we have seen, $P(model|data)$ is usually what we're interested in as data scientists yet $P(data|model)$ is what we can easily compute, either by simulating our model or using analytic equations.
**One of the coolest things:** Bayes Theorem can be proved with a few lines of mathematics. Your instructor will do this on the chalk/white-board now.
### Bayes Theorem solves the above drug testing problem
Bayes Theorem can be used to analytically derive the solution to the 'drug testing' example above as follows.
From Bayes Theorem,
$$P(user|+) = \frac{P(+|user)P(user)}{P(+)}$$
We can expand the denominator here into
$$P(+) = P(+,user) + P(+,non-user) $$
so that
$$ P(+)=P(+|user)P(user) + P(+|non-user)P(non-user)$$
and
$$P(user|+) = \frac{P(+|user)P(user)}{P(+|user)P(user) + P(+|non-user)P(non-user)}$$.
Calculating this explicitly yields
$$P(user|+) = \frac{0.99\times 0.005}{0.99\times 0.005 + 0.01\times 0.995} = 0.332 $$
This means that if an individual tests positive, there is still only a 33.2% chance that they are a user! This is because the number of non-users is so high compared to the number of users.
Coming up: from Bayes Theorem to Bayesian Inference!
| github_jupyter |
<a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/2_transfer_learning_roadmap/4_effect_of_training_epochs/1)%20Understand%20the%20effect%20of%20number%20of%20epochs%20in%20transfer%20learning%20-%20mxnet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Goals
### Understand the role of number of epochs in transfer learning
### Till what point increasing epochs helps in imporving acuracy
### How overtraining can result in overfitting the data
### You will be using skin-cancer mnist to train the classifiers
# Table of Contents
## [0. Install](#0)
## [1. Train a resnet18 network for 5 epochs](#1)
## [2. Re-Train a new experiment for 10 epochs](#2)
## [3. Re-Train a third experiment for 20 epochs](#3)
## [4. Compare the experiments](#4)
<a id='0'></a>
# Install Monk
- git clone https://github.com/Tessellate-Imaging/monk_v1.git
- cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
- (Select the requirements file as per OS and CUDA version)
```
!git clone https://github.com/Tessellate-Imaging/monk_v1.git
# Select the requirements file as per OS and CUDA version
!cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
```
## Dataset Details
- Credits: https://www.kaggle.com/kmader/skin-cancer-mnist-ham10000
- Seven classes
- benign_keratosis_like_lesions
- melanocytic_nevi
- dermatofibroma
- melanoma
- vascular_lesions
- basal_cell_carcinoma
- Bowens_disease
### Download the dataset
```
! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1MRC58-oCdR1agFTWreDFqevjEOIWDnYZ' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1MRC58-oCdR1agFTWreDFqevjEOIWDnYZ" -O skin_cancer_mnist_dataset.zip && rm -rf /tmp/cookies.txt
! unzip -qq skin_cancer_mnist_dataset.zip
```
# Imports
```
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using mxnet-gluon backend
from gluon_prototype import prototype
```
<a id='1'></a>
# Train a resnet18 network for 5 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-5");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Freeze_Base_Network
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Set dataset and select the model
## Quick mode training
- Using Default Function
- dataset_path
- model_name
- freeze_base_network
- num_epochs
## Sample Dataset folder structure
parent_directory
|
|
|------cats
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
|------dogs
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18_v2",
freeze_base_network=True,
num_epochs=5); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 5
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.963
### Final validation loss - 1.062
(You may get a different result)
<a id='2'></a>
# Re-Train a new experiment for 10 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-10");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Epochs-5 (Previously created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-10 (Created Now)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Set dataset and select the model
## Quick mode training
- Using Default Function
- dataset_path
- model_name
- freeze_base_network
- num_epochs
## Sample Dataset folder structure
parent_directory
|
|
|------cats
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
|------dogs
|
|------img1.jpg
|------img2.jpg
|------.... (and so on)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18_v2",
freeze_base_network=True,
num_epochs=10); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 10
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.911
### Final validation loss - 1.017
(You may get a different result)
<a id='3'></a>
# Re-Train a third experiment for 20 epochs
## Creating and managing experiments
- Provide project name
- Provide experiment name
- For a specific data create a single project
- Inside each project multiple experiments can be created
- Every experiment can be have diferent hyper-parameters attached to it
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "Epochs-20");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|
|-----Epochs-5 (Previously created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-10 (Previously Created)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
|
|
|-----Epochs-20 (Created Now)
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
## Modifyable params
- dataset_path: path to data
- model_name: which pretrained model to use
- freeze_base_network: Retrain already trained network or not
- num_epochs: Number of epochs to train for
```
gtf.Default(dataset_path="skin_cancer_mnist_dataset/images",
path_to_csv="skin_cancer_mnist_dataset/train_labels.csv",
model_name="resnet18_v2",
freeze_base_network=True,
num_epochs=20); #Set number of epochs here
#Read the summary generated once you run this cell.
```
## From summary above
Training params
Num Epochs: 20
## Train the classifier
```
#Start Training
gtf.Train();
#Read the training summary generated once you run the cell and training is completed
```
### Final training loss - 0.750
### Final validation loss - 0.829
(You may get a different result)
<a id='4'></a>
# Compare the experiments
```
# Invoke the comparison class
from compare_prototype import compare
```
### Creating and managing comparison experiments
- Provide project name
```
# Create a project
gtf = compare(verbose=1);
gtf.Comparison("Compare-effect-of-num-epochs");
```
### This creates files and directories as per the following structure
workspace
|
|--------comparison
|
|
|-----Compare-effect-of-num-epochs
|
|------stats_best_val_acc.png
|------stats_max_gpu_usage.png
|------stats_training_time.png
|------train_accuracy.png
|------train_loss.png
|------val_accuracy.png
|------val_loss.png
|
|-----comparison.csv (Contains necessary details of all experiments)
### Add the experiments
- First argument - Project name
- Second argument - Experiment name
```
gtf.Add_Experiment("Project", "Epochs-5");
gtf.Add_Experiment("Project", "Epochs-10");
gtf.Add_Experiment("Project", "Epochs-20");
```
### Run Analysis
```
gtf.Generate_Statistics();
```
## Visualize and study comparison metrics
### Training Accuracy Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/train_accuracy.png")
```
### Training Loss Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/train_loss.png")
```
### Validation Accuracy Curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/val_accuracy.png")
```
### Validation loss curves
```
from IPython.display import Image
Image(filename="workspace/comparison/Compare-effect-of-num-epochs/val_loss.png")
```
## Training Accuracies achieved
### With 5 epochs - 67.3%
### With 10 epochs - 68.3%
### With 20 epochs - 73.1%
## Validation accuracies achieved
### With 5 epochs - 63.7%
### With 10 epochs - 63.5%
### With 20 epochs - 72.0%
#### Thing to note - After 15 epochs, accuracies and losses tend to saturate
(You may get a different result)
| github_jupyter |
[](https://colab.research.google.com/github/xxbidiao/plug-and-blend/blob/main/blending_generation_demo_colab.ipynb)
## Introduction
Plug-and-blend generate stories based on both a prompt and one or multiple continuously weighted topics.
Here we show off some capabilities of Plug-and-blend, to illustrate its generation potential and how it can be used in an interactive setting.
## Setup
Let's have code and dataset downloaded and set up.
```
# Downloading the GeDi modifier model.
!wget https://storage.googleapis.com/sfr-gedi-data/gedi_topic.zip
import zipfile
with zipfile.ZipFile('gedi_topic.zip', 'r') as zip_ref:
zip_ref.extractall('./')
```
Then we download the code archive and the base LM.
Here we provide our fine-tuned GPT2-large on ROCStories dataset. You can use a different dataset to fine-tune your own model; as long as its tokenization is compatible with gpt-2, it should work.
```
!gdown --id 1mkNr7unvQKBWayTZPSFlM7XhVMN8iNxA
!unzip plug_and_blend_r1.zip
!gdown --id 1Bhgfp2rZoCfU5tDPxZr5LV36WUfJXNOL
!unzip rocstories_gpt2_large.zip -d baselm
```
Finally, install these dependencies (colab may ask you to restart runtime since we use an older version of `torch`.
```
!pip install transformers==3.5.1
!pip install torch==1.4.0
import nltk
nltk.download('punkt')
```
## Prepare the models
```
from gedi_skill import GediSkill
```
## Load base model
In this notebook, we have prepared all models and their paths set up for you.
However, you can also bring your own base LM.
For this demo, any model that uses the same vocabulary of GPT will work.
If `base_model_path` is not `None`, it is treated as the path to base model.
Otherwise, the original `gpt2-large` model is used instead.
```
base_model_path = "baselm/"
gedi_path = "gedi_topic/"
```
## Demo 1 - Let's generate some sentence!
In this demo, we demonstrate individual-sentence blending generation capability of our blending generation language model.
First, we load in the models (This may take some time):
```
gedi_skill_obj = GediSkill(base_model_path,gedi_path)
# Then we set the parameters needed to generate the sentence.
# Here we first provide a prompt...
sentence = "Welcome to Plug and Blend!"
# Then provide control codes. They can be arbitrary one-token words, so try something else! `Sports`,`Science`,`Business`,`World` works the best, but zero-shot topics are supported too.
topic = {"Science":0.5,"Business":0.5}
# Finally we set the control strength here. A higher value than 1 will result in more steering towards topic specified, but potentially less fluency (Especially when sentence is short and control strength is very high).
# For example, setting it to 2 means 2x control strength.
control_strength = 1
# For ease of demo, scale it to internal strength, so that 1 in control_strength now means 30, the baseline value.
control_strength = control_strength * gedi_skill_obj.disc_weight
# Now just wait for the sentence to be generated.
text = gedi_skill_obj.generate_one_sentence(sentence=sentence, topic=topic,extra_args = {"disc_weight":control_strength})
print(text)
print("OK!")
```
## Demo 2: Run the story generation experience
In this demo, you will provide Control Sketches (described in the paper) so that the planner will generate a 10-sentence story for you.
An agent will interactively ask you (1) a 0-start topic index (e.g. 0 for `Sports` as in the default, `['Sports','Science']` ), (2) a start point of the sketch, which specifies where the effect should emerge, in terms of sentence index (from 0 to 9), (3) an end point of the sketch. See the previous-run log for examples.
(Colab may report out of memory if this demo is performed after demo 1. Feel free to restart the Colab instance.)
For `topics` you can choose from `Sports`,`Science`,`Business`,`World`, or any other word that can be tokenized into one token (extra tokens will be ignored.)
There can be more than 2 topics provided in `topics`. Try having more, and have fun!
```
from story_gen import run_demo
run_demo(
base_language_model_path = base_model_path, # use None to use original GPT2 model.
gedi_path=gedi_path,
topics=['Sports','Science']
)
```
| github_jupyter |
# Summary
Extract SCOPe ids from the training, validation, and test subsets of our training data.
1. Compare the overlap of domains from these three sources:
- `scop95_01Mar17.tgz`, `scop90_01Mar17.tgz`, `scop70_01Mar17.tgz`, `scop40_01Mar17.tgz` from the hh-suite [downloads page](http://wwwuser.gwdg.de/~compbiol/data/hhsuite/databases/hhsuite_dbs/).
- Files downloaded from the SCOPe website.
```bash
rg --text '\0' scop95_a3m.ffdata | wc -l
```
2. Get CATH/Gene3d domains from the PAGNN training set that correspond to different SCOP domains.
----
# Imports
```
import os
import subprocess
import shlex
import io
import re
import itertools
from pathlib import Path
from pprint import pprint
from IPython.display import display
import psutil
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
import pyarrow as pa
import pyarrow.parquet as pq
pd.set_option('display.max_columns', 1000)
%matplotlib inline
```
# Parameters
```
NOTEBOOK_PATH = Path(os.getenv('OUTPUT_DIR', '.')).joinpath('scop_dataset').resolve()
NOTEBOOK_PATH.mkdir(parents=True, exist_ok=True)
NOTEBOOK_PATH
DATA_PATH = Path(os.environ['DATA_DIR']).resolve()
DATA_PATH
DATABIN_PATH = Path(os.environ['DATABIN_DIR']).resolve()
DATABIN_PATH
```
# Spark
```
import pyspark
from pyspark.sql import SparkSession
os.environ['JAVA_HOME']
!ls /home/p/pmkim/strokach/anaconda3
cpu_count = psutil.cpu_count()
spark = (
SparkSession
.builder
.master(f"local[{cpu_count}]")
.appName(NOTEBOOK_PATH.name)
.config('spark.driver.memory', '80G') # Controls both -Xmx and spark.driver.memory
.config('spark.executor.memory', '120g')
.config('spark.memory.storageFraction', 0)
# .config('spark.driver.maxResultSize', '10G')
.getOrCreate()
)
spark
```
# Functions
```
print(re.findall('(-?\d+)-(-?\d+)', '1-10'))
print(re.findall('(-?\d+)-(-?\d+)', '-1-10'))
print(re.findall('(-?\d+)-(-?\d+)', '-1--10'))
print(re.findall('(-?\d+)-(-?\d+)', '1--10'))
print()
def parse_scop_header(header):
columns = [
'scop_id', 'scop_family', 'pdb_chain', 'pdb_domain_range',
'protein_name', 'organism_name',
]
match = re.findall(
'^>(.*) ([a-z]\.\d+\.\d+\.\d+) \((.*):(.*?)\) ([^{}]*) {(.*)}$',
header)
if not match:
raise Exception(f"Could not parser header: {header}")
data = dict(zip(columns, match[0]))
data['pdb_id'] = data['scop_id'][1:5].upper()
domain_range = re.findall('(-?\d+)-(-?\d+)', data['pdb_domain_range'])
if domain_range:
data['pdb_domain_start'], data['pdb_domain_end'] = map(int, domain_range[0])
else:
data['pdb_domain_start'], data['pdb_domain_end'] = None, None
# data['scop_id'] = data['scop_id'] + data['pdb_chain'] + data['pdb_chain_range']
return data
header = ">d1q90l_ f.23.24.1 (L:) PetL subunit of the cytochrome b6f complex {Green alga (Chlamydomonas reinhardtii) [TaxId: 3055]}"
pprint(parse_scop_header(header))
header = ">d2cmyb1 g.2.4.1 (B:7-29) Trypsin inhibitor VhTI {Veronica hederifolia [TaxId: 202477]}"
pprint(parse_scop_header(header))
header = ">d1f93h_ a.34.2.1 (H:) Hepatocyte nuclear factor 1 (HNF-1), N-terminal domain {Mouse (Mus musculus) [TaxId: 10090]}"
pprint(parse_scop_header(header))
header = ">d1ag7a_ g.3.6.1 (A:) Conotoxin {Synthetic, based on Conus geographus, GS}"
pprint(parse_scop_header(header))
def get_scop_domains_from_ffdata(filepath):
assert filepath.suffix == '.ffdata'
proc1 = subprocess.run(
shlex.split(f"head -n1 {filepath}"),
stdout=subprocess.PIPE, stderr=subprocess.PIPE, universal_newlines=True)
if proc1.stderr.strip():
print("proc1 stderr:")
print(proc1.stderr.strip())
proc2 = subprocess.run(
shlex.split(f"rg --text '\\0' {filepath}"),
stdout=subprocess.PIPE, stderr=subprocess.PIPE, universal_newlines=True)
if proc2.stderr.strip():
print("proc2 stderr:")
print(proc2.stderr.strip())
data = []
for i, line in enumerate(itertools.chain(
proc1.stdout.strip().split('\n'), proc2.stdout.strip().split('\n'))):
line = line.replace('\0', '')
if not line:
print(f"Skipping line: {i}")
continue
data.append(parse_scop_header(line))
df = pd.DataFrame(data)
df_ref = pd.read_csv(
filepath.with_suffix('.ffindex'), sep='\t', names=['scop_id', 'start', 'length'])
assert (df['scop_id'].values == df_ref.sort_values('start')['scop_id'].values).all()
return df
df = get_scop_domains_from_ffdata(DATA_PATH / 'hh-suite' / 'scop95_01Mar17' / 'scop95_a3m.ffdata')
df.head()
def get_scop_domains_from_fasta(filepath):
assert filepath.suffix == '.fa'
proc = subprocess.run(
shlex.split(f"rg --text '^>' {filepath}"),
stdout=subprocess.PIPE, stderr=subprocess.PIPE, universal_newlines=True)
if proc.stderr.strip():
print("proc stderr:")
print(proc.stderr.strip())
data = []
for i, line in enumerate(proc.stdout.strip().split('\n')):
if not line:
print(f"Skipping line: {i}")
continue
data.append(parse_scop_header(line))
df = pd.DataFrame(data)
df_ref = pd.read_csv(filepath.with_suffix('.id'), names=['scop_id'])
assert not (set(df['scop_id']) ^ set(df_ref['scop_id']))
return df
df = get_scop_domains_from_fasta(
DATA_PATH / 'scop' / 'v2.07' / 'astral-scopedom-seqres-gd-sel-gs-bib-100-2.07.fa')
df.head()
```
# Databases
```
!ls {DATA_PATH}
```
## `hh-suite`
```
!ls {DATA_PATH}/hh-suite
hhsuite_dfs = {}
for scop_version in ['01Mar17', ]: # '1.75'
for sic in {'01Mar17': [40, 70, 90, 95], '1.75': [70]}[scop_version]:
print(scop_version, sic)
filepath = (
DATA_PATH
.joinpath(f'hh-suite')
.joinpath(f'scop{sic}_{scop_version}')
.joinpath(f'scop{sic}_a3m.ffdata'))
print(filepath, flush=True)
df = get_scop_domains_from_ffdata(filepath)
df['scop_version'] = scop_version
df['seq_identity_cutoff'] = sic
hhsuite_dfs[(scop_version, sic)] = df
hhsuite_dfs.keys()
columns = [
'scop_version', 'seq_identity_cutoff',
'num_domains', 'num_unique_domains', 'scop_domains',
'num_families', 'scop_families'
]
hhsuite_stats = pd.DataFrame(
data=[
(scop_version, seq_identity_cutoff,
len(df), len(set(df['scop_id'])), set(df['scop_id']),
len(set(df['scop_family'])), set(df['scop_family']),
)
for scop_version in ['01Mar17']
for seq_identity_cutoff in [40, 70, 90, 95]
for df in [hhsuite_dfs[(scop_version, seq_identity_cutoff)]]
],
columns=columns,
)
assert (hhsuite_stats['num_domains'] == hhsuite_stats['num_unique_domains']).all()
del hhsuite_stats['num_unique_domains']
hhsuite_stats
schema = None
for (version, sic), df in hhsuite_dfs.items():
filepath = NOTEBOOK_PATH.joinpath(f"scop-hhsuite-{version}-{sic}.parquet")
if schema is None:
table = pa.Table.from_pandas(df, preserve_index=False)
for column_name in ['pdb_domain_start', 'pdb_domain_end']:
idx = table.schema.get_field_index(column_name)
column = table.column(idx).cast(pa.int32())
table = table.remove_column(idx).add_column(idx, column)
schema = table.schema
else:
table = pa.Table.from_pandas(df, schema=schema, preserve_index=False)
pq.write_table(pa.Table.from_pandas(df), filepath, flavor='spark')
```
## `scop`
```
!ls {DATA_PATH}/scop
scop_dfs = {}
for scop_version in ['1.75', '2.07']:
scop_name = 'scop' if float(scop_version) < 2 else 'scope'
print(scop_name, scop_version)
for sic in [10, 20, 25, 30, 35, 40, 50, 70, 90, 95, 100]:
filepath = (
DATA_PATH
.joinpath(f'scop')
.joinpath(f'v{scop_version}')
.joinpath(f'astral-{scop_name}dom-seqres-gd-sel-gs-bib-{sic}-{scop_version}.fa'))
print(filepath, flush=True)
df = get_scop_domains_from_fasta(filepath)
df['scop_version'] = f"{scop_name} v{scop_version}"
df['seq_identity_cutoff'] = sic
scop_dfs[(scop_version, sic)] = df
scop_dfs.keys()
columns = [
'scop_version', 'seq_identity_cutoff',
'num_domains', 'num_unique_domains', 'scop_domains',
'num_families', 'scop_families'
]
scop_stats = pd.DataFrame(
data=[
(scop_version, seq_identity_cutoff,
len(df), len(set(df['scop_id'])), set(df['scop_id']),
len(set(df['scop_family'])), set(df['scop_family']),
)
for scop_version in ['1.75', '2.07']
for seq_identity_cutoff in [10, 20, 25, 30, 35, 40, 50, 70, 90, 95, 100]
for df in [scop_dfs[(scop_version, seq_identity_cutoff)]]
],
columns=columns,
)
assert (scop_stats['num_domains'] == scop_stats['num_unique_domains']).all()
del scop_stats['num_unique_domains']
scop_stats
for (version, sic), df in scop_dfs.items():
filepath = NOTEBOOK_PATH.joinpath(f"scop-astral-{version}-{sic}.parquet")
if schema is None:
table = pa.Table.from_pandas(df, preserve_index=False)
for column_name in ['pdb_domain_start', 'pdb_domain_end']:
idx = table.schema.get_field_index(column_name)
column = table.column(idx).cast(pa.int32())
table = table.remove_column(idx).add_column(idx, column)
schema = table.schema
else:
table = pa.Table.from_pandas(df, schema=schema, preserve_index=False)
pq.write_table(pa.Table.from_pandas(df), filepath, flavor='spark')
```
## Summary
```
# overlap_scop_df
overlap_scop_df = (
scop_stats[scop_stats['scop_version'] == '1.75']
.merge(scop_stats[scop_stats['scop_version'] == '2.07'], on='seq_identity_cutoff', suffixes=('_1', '_2'))
)
overlap_scop_df['num_domains'] = (
overlap_scop_df.apply(lambda r: len(r['scop_domains_1'] & r['scop_domains_2']), axis=1)
)
overlap_scop_df['num_families'] = (
overlap_scop_df.apply(lambda r: len(r['scop_families_1'] & r['scop_families_2']), axis=1)
)
overlap_scop_df['scop_version'] = 'Overlap 1.75 & 2.07'
# overlap_all_df
overlap_all_df = (
scop_stats[scop_stats['scop_version'] == '1.75']
.merge(scop_stats[scop_stats['scop_version'] == '2.07'], on='seq_identity_cutoff', suffixes=('_1', '_2'))
.merge(hhsuite_stats[hhsuite_stats['scop_version'] == '01Mar17'], on='seq_identity_cutoff')
)
overlap_all_df['num_domains'] = (
overlap_all_df.apply(lambda r: len(r['scop_domains_2'] & r['scop_domains']), axis=1)
)
overlap_all_df['num_families'] = (
overlap_all_df.apply(lambda r: len(r['scop_families_2'] & r['scop_families']), axis=1)
)
overlap_all_df['scop_version'] = 'Overlap 2.07 & 01Mar17'
plot_df = pd.concat([
scop_stats[['scop_version', 'seq_identity_cutoff', 'num_domains', 'num_families']],
overlap_scop_df[['scop_version', 'seq_identity_cutoff', 'num_domains', 'num_families']],
hhsuite_stats[['scop_version', 'seq_identity_cutoff', 'num_domains', 'num_families']],
overlap_all_df[['scop_version', 'seq_identity_cutoff', 'num_domains', 'num_families']],
])
with plt.rc_context({'figure.figsize': (12, 6), 'legend.loc': 'upper left', 'savefig.bbox': 'tight'}):
ax = sns.barplot(x="seq_identity_cutoff", y="num_domains", hue="scop_version", data=plot_df)
plt.legend()
plt.savefig(NOTEBOOK_PATH.joinpath('scop_num_domains.png').as_posix(), dpi=300)
with plt.rc_context({'figure.figsize': (12, 6), 'legend.loc': 'upper left', 'savefig.bbox': 'tight'}):
ax = sns.barplot(x="seq_identity_cutoff", y="num_families", hue="scop_version", data=plot_df)
plt.ylim(0, 6100)
plt.legend()
plt.savefig(NOTEBOOK_PATH.joinpath('scop_num_families.png').as_posix(), dpi=300)
```
# Map to training / validation / test datasets
### `scop-hhsuite-01Mar17-40.parquet`
```
ds = spark.sql(f"""\
select *
from parquet.`{NOTEBOOK_PATH}/scop-hhsuite-01Mar17-40.parquet` s
limit 10
""")
df = ds.toPandas()
df.head(2)
hhsuite_dfs.keys()
hhsuite_dfs[('01Mar17', 40)].shape
```
### `adjacency_matrix.parquet`
#### Notes:
- 290,384 domains in in `adjacency_matrix.parquet` map to a PDB chain with 100% sequence identity.
```
ds = spark.sql(f"""\
select *
from parquet.`{DATABIN_PATH}/uniparc_domain/0.1/adjacency_matrix.parquet` am
where am.pc_identity = 100
limit 10
""")
df = ds.toPandas()
df.head(2)
```
### `scop` + `uniparc_domain`
```
ds = spark.sql(f"""\
SELECT
-- scop columns
s.scop_id, s.scop_family, s.pdb_id, s.pdb_chain, s.pdb_domain_start, s.pdb_domain_end,
-- adjacency_matrix columns
am.__index_level_0__ adjacency_matrix_idx, am.uniparc_id, am.database_id cath_id,
am.alignment_length, am.q_start, am.q_end, am.s_start, am.s_end, am.qseq, am.sseq
from parquet.`{NOTEBOOK_PATH}/scop-hhsuite-01Mar17-40.parquet` s
join parquet.`{DATABIN_PATH}/uniparc_domain/0.1/adjacency_matrix.parquet` am ON (
s.pdb_id = am.structure_id and s.pdb_chain = am.chain_id)
where am.pc_identity = 100
and (s.pdb_domain_start is null
or (least(s.pdb_domain_end - s.pdb_domain_start, am.s_end - am.s_start) /
(greatest(s.pdb_domain_end, am.s_end) - least(s.pdb_domain_start, am.s_start))) > 0.9 )
""")
out = (
ds
.write
.format("parquet")
.mode("overwrite")
.save(NOTEBOOK_PATH.joinpath('scop-hhsuite-adjacency_matrix.parquet').as_posix())
)
```
### `scop` + `uniparc`
```
ds = spark.sql(f"""\
select *
from parquet.`{NOTEBOOK_PATH}/scop-hhsuite-01Mar17-40.parquet` s
join parquet.`{DATABIN_PATH}/uniparc_domain/0.1/adjacency_matrix.parquet` am ON (
s.pdb_id = am.structure_id and s.pdb_chain = am.chain_id)
where am.pc_identity = 100
limit 10
""")
df = ds.toPandas()
df.head(2)
```
## Statistics
### Starting
```
df = hhsuite_dfs[('01Mar17', 40)]
display(df.head(2))
print(df.shape)
stating_stats = {
'num_scop_domains': len(set(df['scop_id'])),
'num_scop_families': len(set(df['scop_family'])),
'num_structures': len(set(df['pdb_id'])),
'num_chains': len(set(df[['pdb_id', 'pdb_chain']].apply('-'.join, axis=1))),
}
print(stating_stats)
```
### Mapped
```
df = pq.read_table(NOTEBOOK_PATH.joinpath('scop-hhsuite-adjacency_matrix.parquet').as_posix()).to_pandas()
assert (df['qseq'] == df['sseq']).all()
df = df.drop_duplicates(subset=['qseq'])
display(df.head(2))
print(df.shape)
mapped_stats = {
'num_scop_domains': len(set(df['scop_id'])),
'num_scop_families': len(set(df['scop_family'])),
'num_structures': len(set(df['pdb_id'])),
'num_chains': len(set(df[['pdb_id', 'pdb_chain']].apply('-'.join, axis=1))),
}
print(mapped_stats)
mapped_stats
len(set(df['scop_family']))
len(set(df['cath_id']))
len(set(df['pdb_id']))
.coalesce(100)
df.head(2)
```
| github_jupyter |
$\def\*#1{\mathbf{#1}}$
$\DeclareMathOperator*{\argmax}{arg\,max}$
# Data Types
## Imports
```
import matplotlib as mpl
# pyplot : Provides a MATLAB-like plotting framework
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib notebook
%matplotlib
```
## Data Matrix
The data set is represented by a $n \times d$ **data matrix** :
$$
D =
\begin{pmatrix}
x_{1,1} & x_{1,2} & \cdots & x_{1,d} \\
x_{2,1} & x_{2,2} & \cdots & x_{2,d} \\
\vdots & \vdots & \ddots & \vdots \\
x_{n,1} & x_{n,2} & \cdots & x_{n,d}
\end{pmatrix}
$$
* The *i*-th **row** refers, depending on the application, to an *entity*, *instance*, **record**, *transaction*, *alternative*,...
$$\*x_i = (x_{i1}, x_{i1}, \ldots, x_{id})$$
* The *j*-th **column** refers to an *attribute*, **feature**, *dimension*, *criteria*,...
$$X_j = (x_{1j}, x_{2j}, \ldots, x_{nj})$$
$$
D =
\left(
\begin{array}{c|cccc}
& X_1 & X_2 & \cdots & X_d\\
\hline
\*x_1 & x_{1,1} & x_{1,2} & \cdots & x_{1,d} \\
\*x_2 & x_{2,1} & x_{2,2} & \cdots & x_{2,d} \\
\vdots & \vdots & \vdots & \ddots & \vdots \\
\*x_n & x_{n,1} & x_{n,2} & \cdots & x_{n,d}
\end{array}
\right)
$$
## Iris Data Set
| sepal length (cm) | sepal width (cm) | petal length (cm) | petal width (cm) | Type of iris plant |
| ----------------- | ---------------- | ----------------- | ---------------- | ------------------ |
| 5.1 | 3.5 | 1.4 | 0.2 | Setosa |
| 4.8 | 3.0 | 1.4 | 0.3 | Setosa |
| 6.0 | 3.4 | 4.5 | 1.6 | Versicolor |
| 6.8 | 3.0 | 5.5 | 2.1 | Virginica |
| 6.7 | 3.1 | 5.6 | 2.4 | Virginica |
```
import numpy as np
filename = '../datasets/iris.data'
data = np.loadtxt(filename, delimiter=',', dtype=str)
print(data.nbytes)
print(data)
import numpy as np
filename = '../datasets/iris.data'
data = np.loadtxt(filename, delimiter=',', skiprows=1, usecols=range(4))
print(data.nbytes)
print(data[:5,:])
labels = np.loadtxt(filename, delimiter=',', skiprows=1, usecols=4, dtype=object)
print(labels.nbytes)
print(labels[:5])
import numpy as np
filename = '../datasets/iris.data'
classes = {b'Iris-setosa': 0,
b'Iris-versicolor': 1,
b'Iris-virginica': 2}
classes_converter = {4: lambda c: classes[c]}
data = np.loadtxt(filename, delimiter=',', skiprows=1, converters=classes_converter)
print(data.nbytes)
print(data.dtype)
print(data)
i = 3
xi = data[i]
print(xi)
j = 1
Xj = data[:,j]
print(Xj)
print(data[0:5,:])
Xj = data[0:5,j]
print(Xj)
import pandas
df = pandas.read_csv('../datasets/iris.data')
df.head()
type(df)
df.shape
df.info()
i = 3
xi = df.iloc[i]
print(xi)
j = 1
Xj = df.iloc[:,j]
print(Xj[:5])
print(type(Xj))
df['SepalWidth'][:5]
df['Name'][:5]
```
## Attributes
* Numeric attributes
* Categorical attributes
## Numeric (quantitative) Attributes
* `domain(age)` = $\mathbb{N}$
* `domain(petal length)` = $\mathbb{R}_{>0}$
* **discrete** : finite or countably infinite set of values
* **continuous** : any real value
**Measurement scales**
* **Interval scale** :
* Only addition and substration make sense.
* The *zero point* does not indicate the absence of measurement.
* The `temperature` measured in $^{\circ}C$ is interval-scaled. If two measurements of $20 ^{\circ}C$ and $10 ^{\circ}C$ are compared, what is the right statement ?
* There is a temperature drop of $10 ^{\circ}C$.
* The second measure is twice as cold as the first one.
* **Ratio scale**
* Addition, substraction, and ratio make sense.
* The `Age` attribute is ratio-scaled.
* The `temperature` mesured in *Kelvin* is ratio-scaled.
## Categorical (qualitative) Attributes
* A set of symbols, for example :
* `domain(Education) = {HighSchool, BS, MS, PhD}`
* `domain(Fruits) = {Orange, Apple}`
**Measurement scales**
* **Nominal scale** : values are *unordered*
* **Ordinal scale** : values are *ordered*
## Geometric View
```
fig, ax = plt.subplots()
ax.set_xlabel('Sepal length')
ax.set_ylabel('Sepal width')
X = data[:,0:4]
Y = data[:,4]
ax.scatter(X[:, 0], X[:, 1], c=Y)
fig, axs = plt.subplots(4, 4)
attributes = ['sepal length', 'sepal width', 'petal length', 'petal width']
for i in range(4):
axs[i, 0].set_ylabel(attributes[i])
axs[-1, i].set_xlabel(attributes[i])
for j in range(4):
axs[i, j].scatter(X[:, i], X[:, j], c=Y)
plt.tight_layout(pad=1)
```
### Data binning
```
fig, ax = plt.subplots()
hist = ax.hist([1,1,1,2,2,4,4], bins=3, edgecolor='black', linewidth=1)
fig, ax = plt.subplots()
values = np.array([1, 1, 1, 2, 2, 3, 3, 4, 4, 5, 6, 7])
nbins = 3
size = (max(values) - min(values)) / nbins
print('Size :', size)
ax.hist(values, bins=3, edgecolor='black', linewidth=1)
ax.set_xticks([1, 1 + size, 1 + 2*size, 1 + 3*size])
print(values[values < 1 + size])
print(values[(1 + size <= values) & (values < 1 + 2*size)])
print(values[(1 + 2*size <= values) & (values <= 1 + 3*size)])
fig, ax = plt.subplots()
_, bins, _ = ax.hist(X[:,0], bins=10, edgecolor='black', linewidth=1)
ax.set_xticks(bins)
ax.set_xlabel(attributes[0])
```
## Dependency-oriented data
Relationships between data items :
* **Time-Series** : data generated by continouous measurement over time
* *environmental sensor* : temperature, pressure
* *finantial market analysis*
* **Discrete Sequences**
* *event logs* such as web accesses : Client IP, Web page address
* *strings*
* **Spatial** : non-spatial attributes measured at spatial locations
* *hurricane forecasting* : sea-surface temperature, pressure
* **Spatiotemporal**
* **Network and Graph Data**
* *Web graph*
* *Social networks*
```
rng = pandas.date_range('2017-09-25 08:30:00', periods=30, freq='3s')
temperatures = np.random.randn(len(rng))*2 + 20
ts = pandas.Series(temperatures, index=rng)
print(ts.head())
ts[pandas.Timestamp('2017-09-25 08:30:09')]
```
### Text Data
* A **string** : a discrete sequence of characters
* **Vector-space representation** : words (terms) frequencies (normalized with respect to the document length)
* **Document-term matrix** : $n$ documents $\times$ $d$ terms
```
# import scikit-learn : Machine Learning in Python
# See : http://scikit-learn.org/stable/modules/feature_extraction.html
# and and http://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html
from sklearn.feature_extraction.text import CountVectorizer
# CountVectorizer : Convert a collection of text documents to a matrix of token counts
vectorizer = CountVectorizer()
corpus = ['This is the first document.',
'This is the second second document.',
'And the third one.',
'Is this the first document?']
# Learn the vocabulary dictionary and return term-document matrix.
X = vectorizer.fit_transform(corpus)
print(vectorizer.get_feature_names())
print(type(X))
X.toarray() # dense representation
print(np.sum(X, axis=0))
print(np.sum(X, axis=1))
```
### Graph Data
A graph $G = (V, E)$ with $n$ ***vertices*** and $m$ ***edges*** consists of:
* $V = V(G)$ : a vertex set; $n = |V|$ is the order of $G$
* $E = E(G)$ : a set of pairs of vertices, called edges; $m = |E|$
A ***weighted graph*** is a graph $G = (V, E)$ in which each edge $e \in E(G)$ is given a numerical weight $w(e)$, where $w : E \rightarrow \mathbb{R}$.
```
import networkx as nx
def draw_weighted_graph(g):
pos = nx.spectral_layout(g)
nx.draw_networkx(g, pos)
edge_labels = {edge[0:2]: edge[2]['weight'] for edge in g.edges(data=True)}
nx.draw_networkx_edge_labels(g, pos, edge_labels)
g = nx.Graph()
g.add_nodes_from(['Lille', 'Paris', 'Amiens', 'Arras'])
g.add_edge('Lille', 'Paris', weight=225)
g.add_edge('Lille', 'Amiens', weight=62.7)
g.add_edge('Lille', 'Arras', weight=52.7)
g.add_edge('Paris', 'Amiens', weight=144.4)
g.add_edge('Paris', 'Arras', weight=185.8)
g.add_edge('Amiens', 'Arras', weight=62.6)
draw_weighted_graph(g)
```
| github_jupyter |
# Python tools for data science
(last updated 2020-05-18)
## The PyData Stack
The Python Data Stack comprises a set of packages that makes Python a powerful data science language. These include
+ Numpy: provides arrays and matrix algebra
+ Scipy: provides scientific computing capabilities
+ matplotlib: provides graphing capabilities
These were the original stack that was meant to replace Matlab. However, these were meant to tackle purely numerical data, and the kinds of heterogeneous data we regularly face needed more tools. These were added more recently.
+ Pandas: provides data analytic structures like the data frame, as well as basic descriptive statistical capabilities
+ statsmodels: provides a fairly comprehensive set of statistical functions
+ scikit-learn: provides machine learning capabilities
This is the basic stack of packages we will be using in this workshop. Additionally we will use a few packages that add some functionality to the data science process. These include
+ seaborn: Better statistical graphs
+ plotly: Interactive graphics
+ biopython: Python for bioinformatics
We may also introduce the package `rpy2` which allows one to run R from within Python. This can be useful since many bioinformatic pipelines are already implemented in R.
> The [PyData stack](https://scipy.org) also includes `sympy`, a symbolic mathematics package emulating Maple
## Numpy (numerical and scientific computing)
We start by importing the Numpy package into Python using the alias `np`.
```
import numpy as np
```
Numpy provides both arrays (vectors, matrices, higher dimensional arrays) and vectorized functions which are very fast. Let's see how this works.
```
z = [1,2,3,4,5,6,7,8,9.3,10.6] # This is a list
z_array = np.array(z)
z_array
```
Now, we have already seen functions in Python earlier. In Numpy, there are functions that are optimized for arrays, that can be accessed directly from the array objects. This is an example of *object-oriented programming* in Python, where functions are provided for particular *classes* of objects, and which can be directly accessed from the objects. We will use several such functions over the course of this workshop, but we won't actually talk about how to do this program development here.
> Numpy functions are often very fast, and are *vectorized*, i.e., they are written to work on vectors of numbers rather than single numbers. This is an advantage in data science since we often want to do the same operation to all elements of a column of data, which is essentially a vector
We apply the functions `sum`, `min` (minimum value) and `max` (maximum value) to `z_array`.
```
z_array.sum()
z_array.min()
z_array.max()
```
The versions of these functions in Numpy are optimized for arrays and are quite a bit faster than the corresponding functions available in base Python. When doing data work, these are the preferred functions.
These functions can also be used in the usual function manner:
```
np.max(z_array)
```
Calling `np.max` ensures that we are using the `max` function from numpy, and not the one in base Python.
### Numpy data types
Numpy arrays are homogeneous in type.
```
np.array(['a','b','c'])
np.array([1,2,3,6,8,29])
```
But, what if we provide a heterogeneous list?
```
y = [1,3,'a']
np.array(y)
```
So what's going on here? Upon conversion from a heterogeneous list, numpy converted the numbers into strings. This is necessary since, by definition, numpy arrays can hold data of a single type. When one of the elements is a string, numpy casts all the other entities into strings as well. Think about what would happen if the opposite rule was used. The string 'a' doesn't have a corresponding number, while both numbers 1 and 3 have corresponding string representations, so going from string to numeric would create all sorts of problems.
> The advantage of numpy arrays is that the data is stored in a contiguous section of memory, and you can be very efficient with homogeneous arrays in terms of manipulating them, applying functions, etc. However, `numpy` does provide a "catch-all" `dtype` called `object`, which can be any Python object. This `dtype` essentially is an array of pointers to actual data stored in different parts of the memory. You can get to the actual objects by extracting them. So one could do <a name='object'></a>
```
np.array([1,3,'a'], dtype='object')
```
> which would basically be a valid `numpy` array, but would go back to the actual objects when used, much like a list. We can see this later if we want to transform a heterogeneous `pandas` `DataFrame` into a `numpy` array. It's not particularly useful as is, but it prevents errors from popping up during transformations from `pandas` to `numpy`.
### Generating data in numpy
We had seen earlier how we could generate a sequence of numbers in a list using `range`. In numpy, you can generate a sequence of numbers in an array using `arange` (which actually creates the array rather than provide an iterator like `range`).
```
np.arange(10)
```
You can also generate regularly spaced sequences of numbers between particular values
```
np.linspace(start=0, stop=1, num=11) # or np.linspace(0, 1, 11)
```
You can also do this with real numbers rather than integers.
```
np.linspace(start = 0, stop = 2*np.pi, num = 10)
```
More generally, you can transform lists into `numpy` arrays. We saw this above for vectors. For matrices, you can provide a list of lists. Note the double `[` in front and back.
```
np.array([[1,3,5,6],[4,3,9,7]])
```
You can generate an array of 0's
```
np.zeros(10)
```
This can easily be extended to a two-dimensional array (a matrix), by specifying the dimension of the matrix as a tuple.
```
np.zeros((10,10))
```
You can also generate a matrix of 1s in a similar manner.
```
np.ones((3,4))
```
In matrix algebra, the identity matrix is important. It is a square matrix with 1's on the diagonal and 0's everywhere else.
```
np.eye(4)
```
You can also create numpy vectors directly from lists, as long as lists are made up of atomic elements of the same type. This means a list of numbers or a list of strings. The elements can't be more composite structures, generally. One exception is a list of lists, where all the lists contain the same type of atomic data, which, as we will see, can be used to create a matrix or 2-dimensional array.
```
a = [1,2,3,4,5,6,7,8]
b = ['a','b','c','d','3']
np.array(a)
np.array(b)
```
#### Random numbers
Generating random numbers is quite useful in many areas of data science. All computers don't produce truly random numbers but generate *pseudo-random* sequences. These are completely deterministic sequences defined algorithmically that emulate the properties of random numbers. Since these are deterministic, we can set a *seed* or starting value for the sequence, so that we can exactly reproduce this sequence to help debug our code. To actually see how things behave in simulations we will often run several sequences of random numbers starting at different seed values.
The seed is set by the `RandomState` function within the `random` submodule of numpy. Note that all Python names are case-sensitive.
```
rng = np.random.RandomState(35) # set seed
rng.randint(0, 10, (3,4))
```
We have created a 3x4 matrix of random integers between 0 and 10 (in line with slicing rules, this includes 0 but not 10).
We can also create a random sample of numbers between 0 and 1.
```
rng.random_sample((5,2))
```
We'll see later how to generate random numbers from particular probability distributions.
### Vectors and matrices
Numpy generates arrays, which can be of arbitrary dimension. However the most useful are vectors (1-d arrays) and matrices (2-d arrays).
In these examples, we will generate samples from the Normal (Gaussian) distribution, with mean 0 and variance 1.
```
A = rng.normal(0,1,(4,5))
```
We can compute some characteristics of this matrix's dimensions. The number of rows and columns are given by `shape`.
```
A.shape
```
The total number of elements are given by `size`.
```
A.size
```
If we want to create a matrix of 0's with the same dimensions as `A`, we don't actually have to compute its dimensions. We can use the `zeros_like` function to figure it out.
```
np.zeros_like(A)
```
We can also create vectors by only providing the number of rows to the random sampling function. The number of columns will be assumed to be 1.
```
B = rng.normal(0, 1, (4,))
B
```
#### Extracting elements from arrays
The syntax for extracting elements from arrays is almost exactly the same as for lists, with the same rules for slices.
**Exercise:** State what elements of B are extracted by each of the following statements
```
B[:3]
B[:-1]
B[[0,2,4]]
B[[0,2,5]]
```
For matrices, we have two dimensions, so you can slice by rows, or columns or both.
```
A
```
We can extract the first column by specifying `:` (meaning everything) for the rows, and the index for the column (reminder, Python starts counting at 0)
```
A[:,0]
```
Similarly the 4th row can be extracted by putting the row index, and `:` for the column index.
```
A[3,:]
```
All slicing operations work for rows and columns
```
A[:2,:2]
```
#### Array operations
We can do a variety of vector and matrix operations in `numpy`.
First, all usual arithmetic operations work on arrays, like adding or multiplying an array with a scalar.
```
A = rng.randn(3,5)
A
A + 10
```
We can also add and multiply arrays __element-wise__ as long as they are the same shape.
```
B = rng.randint(0,10, (3,5))
B
A + B
A * B
```
You can also do **matrix multiplication**. Recall what this is.
If you have a matrix $A_{m x n}$ and another matrix $B_{n x p}$, as long as the number of columns of $A$ and rows of $B$ are the same, you can multiply them ($C_{m x p} = A_{m x n}B_{n x p}$), with the (i,j)-th element of C being
$$ c_{ij} = \sum_{k=1}^n a_{ik}b_{kj}, i= 1, \dots, m; j = 1, \dots, p$$
In `numpy` the operant for matrix multiplication is `@`.
In the above examples, `A` and `B` cannot be multiplied since they have incompatible dimensions. However, we can take the *transpose* of `B`, i.e. flip the rows and columns, to make it compatible with `A` for matrix multiplication.
```
A @ np.transpose(B)
np.transpose(A) @ B
```
More generally, you can *reshape* a `numpy` array into a new shape, provided it is compatible with the number of elements in the original array.
```
D = rng.randint(0,5, (4,4))
D
D.reshape(8,2)
D.reshape(1,16)
```
This can also be used to cast a vector into a matrix.
```
e = np.arange(20)
E = e.reshape(5,4)
E
```
> One thing to note in all the reshaping operations above is that the new array takes elements of the old array **by row**. See the examples above to convince yourself of that.
#### Statistical operations on arrays
You can sum all the elements of a matrix using `sum`. You can also sum along rows or along columns by adding an argument to the `sum` function.
```
A = rng.normal(0, 1, (4,2))
A
A.sum()
```
You can sum along rows (i.e., down columns) with the option `axis = 0`
```
A.sum(axis=0)
```
You can sum along columns (i.e., across rows) with `axis = 1`.
```
A.sum(axis=1)
```
> Of course, you can use the usual function calls: `np.sum(A, axis = 1)`
We can also find the minimum and maximum values.
```
A.min(axis = 0)
A.max(axis = 0)
```
We can also find the **position** where the minimum and maximum values occur.
```
A.argmin(axis=0)
A.argmax(axis=0)
```
We can sort arrays and also find the indices which will result in the sorted array. I'll demonstrate this for a vector, where it is more relevant
```
a = rng.randint(0,10, 8)
a
np.sort(a)
np.argsort(a)
a[np.argsort(a)]
```
`np.argsort` can also help you find the 2nd smallest or 3rd largest value in an array, too.
```
ind_2nd_smallest = np.argsort(a)[1]
a[ind_2nd_smallest]
ind_3rd_largest = np.argsort(a)[-3]
a[ind_3rd_largest]
```
You can also sort strings in this way.
```
m = np.array(['Aram','Raymond','Elizabeth','Donald','Harold'])
np.sort(m)
```
If you want to sort arrays **in place**, you can use the `sort` function in a different way.
```
m.sort()
m
```
#### Putting arrays together
We can put arrays together by row or column, provided the corresponding axes have compatible lengths.
```
A = rng.randint(0,5, (3,5))
B = rng.randint(0,5, (3,5))
print('A = ', A)
print('B = ', B)
np.hstack((A,B))
np.vstack((A,B))
```
Note that both `hstack` and `vstack` take a **tuple** of arrays as input.
#### Logical/Boolean operations
You can query a matrix to see which elements meet some criterion. In this example, we'll see which elements are negative.
```
A < 0
```
This is called **masking**, and is useful in many contexts.
We can extract all the negative elements of A using
```
A[A<0]
```
This forms a 1-d array. You can also count the number of elements that meet the criterion
```
np.sum(A<0)
```
Since the entity `A<0` is a matrix as well, we can do row-wise and column-wise operations as well.
### Beware of copies
One has to be a bit careful with copying objects in Python. By default, if you just assign one object to a new name, it does a *shallow copy*, which means that both names point to the same memory. So if you change something in the original, it also changes in the new copy.
```
A[0,:]
A1 = A
A1[0,0] = 4
A[0,0]
```
To actually create a copy that is not linked back to the original, you have to make a *deep copy*, which creates a new space in memory and a new pointer, and copies the original object to the new memory location
```
A1 = A.copy()
A1[0,0] = 6
A[0,0]
```
You can also replace sub-matrices of a matrix with new data, provided that the dimensions are compatible. (Make sure that the sub-matrix we are replacing below truly has 2 rows and 2 columns, which is what `np.eye(2)` will produce)
```
A[:2,:2] = np.eye(2)
A
```
#### Reducing matrix dimensions
Sometimes the output of some operation ends up being a matrix of one column or one row. We can reduce it to become a vector. There are two functions that can do that, `flatten` and `ravel`.
```
A = rng.randint(0,5, (5,1))
A
A.flatten()
A.ravel()
```
So why two functions? I'm not sure, but they do different things behind the scenes. `flatten` creates a **copy**, i.e. a new array disconnected from `A`. `ravel` creates a **view**, so a representation of the original array. If you then changed a value after a `ravel` operation, you would also change it in the original array; if you did this after a `flatten` operation, you would not.
### Broadcasting in Python
Python deals with arrays in an interesting way, in terms of matching up dimensions of arrays for arithmetic operations. There are 3 rules:
1. If two arrays differ in the number of dimensions, the shape of the smaller array is padded with 1s on its _left_ side
2. If the shape doesn't match in any dimension, the array with shape = 1 in that dimension is stretched to match the others' shape
3. If in any dimension the sizes disagree and none of the sizes are 1, then an error is generated
```
A = rng.normal(0,1,(4,5))
B = rng.normal(0,1,5)
A.shape
B.shape
A - B
```
B is 1-d, A is 2-d, so B's shape is made into (1,5) (added to the left). Then it is repeated into 4 rows to make it's shape (4,5), then the operation is performed. This means that we subtract the first element of B from the first column of A, the second element of B from the second column of A, and so on.
You can be explicit about adding dimensions for broadcasting by using `np.newaxis`.
```
B[np.newaxis,:].shape
B[:,np.newaxis].shape
```
#### An example (optional, intermediate/advanced))
This can be very useful, since these operations are faster than for loops. For example:
```
d = rng.random_sample((10,2))
d
```
We want to find the Euclidean distance (the sum of squared differences) between the points defined by the rows. This should result in a 10x10 distance matrix
```
d.shape
d[np.newaxis,:,:]
```
creates a 3-d array with the first dimension being of length 1
```
d[np.newaxis,:,:].shape
d[:, np.newaxis,:]
```
creates a 3-d array with the 2nd dimension being of length 1
```
d[:,np.newaxis,:].shape
```
Now for the trick, using broadcasting of arrays. These two arrays are incompatible without broadcasting, but with broadcasting, the right things get repeated to make things compatible
```
dist_sq = np.sum((d[:,np.newaxis,:] - d[np.newaxis,:,:]) ** 2)
dist_sq.shape
dist_sq
```
Whoops! we wanted a 10x10 matrix, not a scalar.
```
(d[:,np.newaxis,:] - d[np.newaxis,:,:]).shape
```
What we really want is the 10x10 distance matrix.
```
dist_sq = np.sum((d[:,np.newaxis,:] - d[np.newaxis,:,:]) ** 2, axis=2)
```
You can verify what is happening by creating `D = d[:,np.newaxis,:]-d[np.newaxis,:,:]` and then looking at `D[:,:,0]` and `D[:,:,1]`. These are the difference between each combination in the first and second columns of d, respectively. Squaring and summing along the 3rd axis then gives the sum of squared differences.
```
dist_sq
dist_sq.shape
dist_sq.diagonal()
```
### Conclusions moving forward
It's important to understand numpy and arrays, since most data sets we encounter are rectangular. The notations and operations we saw in numpy will translate to data, except for the fact that data is typically heterogeneous, i.e., of different types. The problem with using numpy for modern data analysis is that if you have mixed data types, it will all be coerced to strings, and then you can't actually do any data analysis.
The solution to this issue (which is also present in Matlab) came about with the `pandas` package, which is the main workhorse of data science in Python
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 回帰: 燃費を予測する
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ja/r1/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ja/r1/tutorials/keras/basic_regression.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
Note: これらのドキュメントは私たちTensorFlowコミュニティが翻訳したものです。コミュニティによる 翻訳は**ベストエフォート**であるため、この翻訳が正確であることや[英語の公式ドキュメント](https://www.tensorflow.org/?hl=en)の 最新の状態を反映したものであることを保証することはできません。 この翻訳の品質を向上させるためのご意見をお持ちの方は、GitHubリポジトリ[tensorflow/docs](https://github.com/tensorflow/docs)にプルリクエストをお送りください。 コミュニティによる翻訳やレビューに参加していただける方は、 [docs-ja@tensorflow.org メーリングリスト](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ja)にご連絡ください。
回帰問題では、価格や確率といった連続的な値の出力を予測することが目的となります。これは、分類問題の目的が、(例えば、写真にリンゴが写っているかオレンジが写っているかといった)離散的なラベルを予測することであるのとは対照的です。
このノートブックでは、古典的な[Auto MPG](https://archive.ics.uci.edu/ml/datasets/auto+mpg)データセットを使用し、1970年代後半から1980年台初めの自動車の燃費を予測するモデルを構築します。この目的のため、モデルにはこの時期の多数の自動車の仕様を読み込ませます。仕様には、気筒数、排気量、馬力、重量などが含まれています。
このサンプルでは`tf.keras` APIを使用しています。詳細は[このガイド](https://www.tensorflow.org/r1/guide/keras)を参照してください。
```
# ペアプロットのためseabornを使用します
!pip install seaborn
from __future__ import absolute_import, division, print_function, unicode_literals
import pathlib
import pandas as pd
import seaborn as sns
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
print(tf.__version__)
```
## Auto MPG データセット
このデータセットは[UCI Machine Learning Repository](https://archive.ics.uci.edu/)から入手可能です。
### データの取得
まず、データセットをダウンロードします。
```
dataset_path = keras.utils.get_file("auto-mpg.data", "http://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/auto-mpg.data")
dataset_path
```
pandasを使ってデータをインポートします。
```
column_names = ['MPG','Cylinders','Displacement','Horsepower','Weight',
'Acceleration', 'Model Year', 'Origin']
raw_dataset = pd.read_csv(dataset_path, names=column_names,
na_values = "?", comment='\t',
sep=" ", skipinitialspace=True)
dataset = raw_dataset.copy()
dataset.tail()
```
### データのクレンジング
このデータには、いくつか欠損値があります。
```
dataset.isna().sum()
```
この最初のチュートリアルでは簡単化のためこれらの行を削除します。
```
dataset = dataset.dropna()
```
`"Origin"`の列は数値ではなくカテゴリーです。このため、ワンホットエンコーディングを行います。
```
origin = dataset.pop('Origin')
dataset['USA'] = (origin == 1)*1.0
dataset['Europe'] = (origin == 2)*1.0
dataset['Japan'] = (origin == 3)*1.0
dataset.tail()
```
### データを訓練用セットとテスト用セットに分割
データセットを訓練用セットとテスト用セットに分割しましょう。
テスト用データセットは、作成したモデルの最終評価に使用します。
```
train_dataset = dataset.sample(frac=0.8,random_state=0)
test_dataset = dataset.drop(train_dataset.index)
```
### データの調査
訓練用セットのいくつかの列の組み合わせの同時分布を見てみましょう。
```
sns.pairplot(train_dataset[["MPG", "Cylinders", "Displacement", "Weight"]], diag_kind="kde")
```
全体の統計値も見てみましょう。
```
train_stats = train_dataset.describe()
train_stats.pop("MPG")
train_stats = train_stats.transpose()
train_stats
```
### ラベルと特徴量の分離
ラベル、すなわち目的変数を特徴量から切り離しましょう。このラベルは、モデルに予測させたい数量です。
```
train_labels = train_dataset.pop('MPG')
test_labels = test_dataset.pop('MPG')
```
### データの正規化
上の`train_stats`のブロックをもう一度見て、それぞれの特徴量の範囲がどれほど違っているかに注目してください。
スケールや値の範囲が異なる特徴量を正規化するのは良い習慣です。特徴量の正規化なしでもモデルは収束する**かもしれませんが**、モデルの訓練はより難しくなり、結果として得られたモデルも入力で使われる単位に依存することになります。
注:(正規化に使用する)統計量は意図的に訓練用データセットだけを使って算出していますが、これらはテスト用データセットの正規化にも使うことになります。テスト用のデータセットを、モデルの訓練に使用した分布と同じ分布に射影する必要があるのです。
```
def norm(x):
return (x - train_stats['mean']) / train_stats['std']
normed_train_data = norm(train_dataset)
normed_test_data = norm(test_dataset)
```
この正規化したデータを使ってモデルを訓練することになります。
注意:ここで入力の正規化に使った統計量(平均と標準偏差)は、先程実施したワンホットエンコーディングとともに、モデルに供給する他のどんなデータにも適用する必要があります。テスト用データセットだけでなく、モデルを本番で使用する際の生のデータも同様です。
## モデル
### モデルの構築
それではモデルを構築しましょう。ここでは、2つの全結合の隠れ層と、1つの連続値を返す出力層からなる、`Sequential`モデルを使います。モデルを構築するステップは`build_model`という1つの関数の中に組み込みます。あとから2つ目のモデルを構築するためです。
```
def build_model():
model = keras.Sequential([
layers.Dense(64, activation=tf.nn.relu, input_shape=[len(train_dataset.keys())]),
layers.Dense(64, activation=tf.nn.relu),
layers.Dense(1)
])
optimizer = tf.keras.optimizers.RMSprop(0.001)
model.compile(loss='mean_squared_error',
optimizer=optimizer,
metrics=['mean_absolute_error', 'mean_squared_error'])
return model
model = build_model()
```
### モデルの検証
`.summary`メソッドを使って、モデルの簡単な説明を表示します。
```
model.summary()
```
では、モデルを試してみましょう。訓練用データのうち`10`個のサンプルからなるバッチを取り出し、それを使って`model.predict`メソッドを呼び出します。
```
example_batch = normed_train_data[:10]
example_result = model.predict(example_batch)
example_result
```
うまく動作しているようです。予定通りの型と形状の出力が得られています。
### モデルの訓練
モデルを1000エポック訓練し、訓練と検証の正解率を`history`オブジェクトに記録します。
```
# エポックが終わるごとにドットを一つ出力することで進捗を表示
class PrintDot(keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs):
if epoch % 100 == 0: print('')
print('.', end='')
EPOCHS = 1000
history = model.fit(
normed_train_data, train_labels,
epochs=EPOCHS, validation_split = 0.2, verbose=0,
callbacks=[PrintDot()])
```
`history`オブジェクトに保存された数値を使ってモデルの訓練の様子を可視化します。
```
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
hist.tail()
import matplotlib.pyplot as plt
def plot_history(history):
hist = pd.DataFrame(history.history)
hist['epoch'] = history.epoch
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Abs Error [MPG]')
plt.plot(hist['epoch'], hist['mean_absolute_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_absolute_error'],
label = 'Val Error')
plt.legend()
plt.ylim([0,5])
plt.figure()
plt.xlabel('Epoch')
plt.ylabel('Mean Square Error [$MPG^2$]')
plt.plot(hist['epoch'], hist['mean_squared_error'],
label='Train Error')
plt.plot(hist['epoch'], hist['val_mean_squared_error'],
label = 'Val Error')
plt.legend()
plt.ylim([0,20])
plot_history(history)
```
このグラフを見ると、検証エラーは100エポックを過ぎたあたりで改善が見られなくなり、むしろ悪化しているようです。検証スコアの改善が見られなくなったら自動的に訓練を停止するように、`model.fit`メソッド呼び出しを変更します。ここでは、エポック毎に訓練状態をチェックする*EarlyStopping*コールバックを使用します。設定したエポック数の間に改善が見られない場合、訓練を自動的に停止します。
このコールバックについての詳細は[ここ](https://www.tensorflow.org/versions/master/api_docs/python/tf/keras/callbacks/EarlyStopping)を参照ください。
```
model = build_model()
# The patience parameter is the amount of epochs to check for improvement
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', patience=10)
history = model.fit(normed_train_data, train_labels, epochs=EPOCHS,
validation_split = 0.2, verbose=0, callbacks=[early_stop, PrintDot()])
plot_history(history)
```
検証用データセットでのグラフを見ると、平均誤差は+/- 2 MPG(マイル/ガロン)前後です。これは良い精度でしょうか?その判断はおまかせします。
モデルの訓練に使用していない**テスト用**データセットを使って、モデルがどれくらい汎化できているか見てみましょう。これによって、モデルが実際の現場でどれくらい正確に予測できるかがわかります。
```
loss, mae, mse = model.evaluate(normed_test_data, test_labels, verbose=0)
print("Testing set Mean Abs Error: {:5.2f} MPG".format(mae))
```
### モデルを使った予測
最後に、テストデータを使ってMPG値を予測します。
```
test_predictions = model.predict(normed_test_data).flatten()
plt.scatter(test_labels, test_predictions)
plt.xlabel('True Values [MPG]')
plt.ylabel('Predictions [MPG]')
plt.axis('equal')
plt.axis('square')
plt.xlim([0,plt.xlim()[1]])
plt.ylim([0,plt.ylim()[1]])
_ = plt.plot([-100, 100], [-100, 100])
```
そこそこ良い予測ができているように見えます。誤差の分布を見てみましょう。
```
error = test_predictions - test_labels
plt.hist(error, bins = 25)
plt.xlabel("Prediction Error [MPG]")
_ = plt.ylabel("Count")
```
とても正規分布には見えませんが、サンプル数が非常に小さいからだと考えられます。
## 結論
このノートブックでは、回帰問題を扱うためのテクニックをいくつか紹介しました。
* 平均二乗誤差(MSE: Mean Squared Error)は回帰問題に使われる一般的な損失関数です(分類問題には異なる損失関数が使われます)。
* 同様に、回帰問題に使われる評価指標も分類問題とは異なります。回帰問題の一般的な評価指標は平均絶対誤差(MAE: Mean Absolute Error)です。
* 入力数値特徴量の範囲が異なっている場合、特徴量毎に同じ範囲に正規化するべきです。
* 訓練用データが多くない場合、過学習を避けるために少ない隠れ層を持つ小さいネットワークを使うというのが良い方策の1つです。
* Early Stoppingは過学習を防止するための便利な手法の一つです。
| github_jupyter |
```
from matplotlib import pyplot as plt
from matplotlib.lines import Line2D
import matplotlib
%matplotlib inline
from ovejero import forward_modeling, model_trainer
import os
import corner
# Modifies the paths in the config to agree with the paths being used on the current computer.
def recursive_str_checker(cfg_dict):
for key in cfg_dict:
if isinstance(cfg_dict[key],str):
cfg_dict[key] = cfg_dict[key].replace('/home/swagnercarena/ovejero/',root_path)
if isinstance(cfg_dict[key],dict):
recursive_str_checker(cfg_dict[key])
```
# Comparing Performance to Forward Modeling
__Author:__ Sebastian Wagner-Carena
__Last Run:__ 07/27/2020
__Goals:__ Compare the performance of the BNN model to a forward modeling approach
__Before running this notebook:__ You will have to download and unzip the bnn samples, chains, and datasets that can be found here (TODO). Because we already have the BNN samples, the model weights are not neccesary.
## Comparison of Full and GMM results for 0.1% Dropout
First, we need to load all of our forward modeling and BNN samples for all three of our BNN models. Our forward modeling is done directly through lenstronomy. To get lenstronomy to load the weights for the forward modeling, we will have to sample once. This should take at most a second or two.
```
root_path = os.getcwd()[:-5]
# We will pull our lensing images from an independent test set.
test_set_path = root_path + 'datasets/test/'
# Grab the config paths for our three BNN models
nn1_config_path = root_path + 'configs/nn1_hr.json'
nn2_config_path = root_path + 'configs/nn2_slr.json'
nn3_config_path = root_path + 'configs/nn3_slr.json'
# Load the config for our three models.
nn1_cfg = model_trainer.load_config(nn1_config_path)
nn2_cfg = model_trainer.load_config(nn2_config_path)
nn3_cfg = model_trainer.load_config(nn3_config_path)
recursive_str_checker(nn1_cfg)
recursive_str_checker(nn2_cfg)
recursive_str_checker(nn3_cfg)
# Samples are already generated so we don't need the model weights
lite_class = True
fow_model_nn1 = forward_modeling.ForwardModel(nn1_cfg,lite_class=lite_class,test_set_path=test_set_path)
fow_model_nn2 = forward_modeling.ForwardModel(nn2_cfg,lite_class=lite_class,test_set_path=test_set_path)
fow_model_nn3 = forward_modeling.ForwardModel(nn3_cfg,lite_class=lite_class,test_set_path=test_set_path)
```
We need to select the image we'll use for our forward modeling. So long as you've downloaded the datasets or generated them using the baobab configs provided in this git repo, image index 40 will have the correct meaning. This should print out the same image and information 3 times. Note the random noise is fixed by a set seed to ensure that the forward modeling and the BNN see the exact same image.
```
# Select the image we want to forward model.
image_index = 50
fow_model_nn1.select_image(image_index)
fow_model_nn2.select_image(image_index)
fow_model_nn3.select_image(image_index)
```
Now we need to initialize our three forward modeling samplers. They are all pulling the same weights (since the forward model doesn't care which BNN was used).
```
# Initialize our sampler for the three models
walker_ratio = 50
n_samps = 1
save_path_chains = os.path.join(root_path,'forward_modeling/test_%s.h5'%(
fow_model_nn1.true_values['img_filename'][:-4]))
fow_model_nn1.initialize_sampler(walker_ratio,save_path_chains)
fow_model_nn1.run_sampler(n_samps)
fow_model_nn2.initialize_sampler(walker_ratio,save_path_chains)
fow_model_nn2.run_sampler(n_samps)
fow_model_nn3.initialize_sampler(walker_ratio,save_path_chains)
fow_model_nn3.run_sampler(n_samps)
```
Now we specify the path to the pre-run BNN samples. If you have the weights downloaded, feel free to rerun this on a GPU. Even without a GPU it should only take tens of seconds.
```
num_samples = 10000
burnin = 2000
sample_save_dir_nn1 = os.path.join(root_path,'forward_modeling/nn1_test_%s_samps'%(
fow_model_nn1.true_values['img_filename'][:-4]))
sample_save_dir_nn2 = os.path.join(root_path,'forward_modeling/nn2_test_%s_samps'%(
fow_model_nn1.true_values['img_filename'][:-4]))
sample_save_dir_nn3 = os.path.join(root_path,'forward_modeling/nn3_test_%s_samps'%(
fow_model_nn1.true_values['img_filename'][:-4]))
```
Now we can plot the comparison of the forward modeling posterior and the full / GMM posterior.
```
# Now let's look at the corner plot of the parameters we care about (the lens parameters)
color_map = ['#d95f02','#7570b3','#000000']
truth_color = '#e7298a'
plot_limits = [(0.03,0.09),(-0.03,0.03),(-0.07,-0.04),(-0.1,-0.07),(-0.2,-0.1),(-0.08,0.02),(1.65,1.83),
(-0.05,-0.02)]
save_fig_path = 'figures/fow_model_comp.pdf'
fontsize = 20
matplotlib.rcParams.update({'font.size': 12})
fig = fow_model_nn2.plot_posterior_contours(burnin,num_samples,sample_save_dir=sample_save_dir_nn2,
color_map=color_map,plot_limits=plot_limits,truth_color=truth_color,
save_fig_path=None,show_plot=False,plot_fow_model=False,add_legend=False,
fontsize=fontsize)
fig = fow_model_nn3.plot_posterior_contours(burnin,num_samples,sample_save_dir=sample_save_dir_nn3,
color_map=color_map[1:],plot_limits=plot_limits,truth_color=truth_color,
save_fig_path=None,fig=fig,show_plot=False,add_legend=False,
fontsize=fontsize)
handles = [Line2D([0], [0], color=color_map[0], lw=10),
Line2D([0], [0], color=color_map[1], lw=10),
Line2D([0], [0], color=color_map[2], lw=10)]
fig.legend(handles,[r'Full BNN 0.1% Dropout',r'GMM BNN 0.1% Dropout','Forward Modeling'],loc=(0.525,0.73),
fontsize=20,framealpha=1.0)
fig.savefig(save_fig_path)
```
## Comparison for Diagonal BNN 30% Dropout
We want to generate the same plot as above, but now with the diagaonal BNN model.
```
# Now let's look at the corner plot of the parameters we care about (the lens parameters)
color_map = ['#1b9e77','#000000']
truth_color = '#e7298a'
plot_limits = [(0.00,0.12),(-0.06,0.06),(-0.1,-0.00),(-0.14,-0.04),(-0.3,-0.0),(-0.18,0.12),(1.65,1.90),
(-0.09,0.02)]
save_fig_path = 'figures/fow_model_comp_diag.pdf'
matplotlib.rcParams.update({'font.size': 12})
fig = fow_model_nn1.plot_posterior_contours(burnin,num_samples,sample_save_dir=sample_save_dir_nn1,
color_map=color_map,plot_limits=plot_limits,truth_color=truth_color,
save_fig_path=None,show_plot=False,add_legend=False,fontsize=fontsize)
handles = [Line2D([0], [0], color=color_map[0], lw=10),
Line2D([0], [0], color=color_map[1], lw=10)]
fig.legend(handles,[r'Diagonal BNN 30% Dropout','Forward Modeling'],loc=(0.54,0.73),fontsize=20,framealpha=1.0)
fig.savefig(save_fig_path)
```
| github_jupyter |
```
import matplotlib.pyplot as plt
import numpy as np
plt.style.use('ggplot')
plt.rcParams['font.family'] = 'serif'
plt.rcParams['font.serif'] = 'Ubuntu'
plt.rcParams['font.monospace'] = 'Ubuntu Mono'
plt.rcParams['font.size'] = 12
plt.rcParams['axes.labelsize'] = 11
plt.rcParams['axes.labelweight'] = 'bold'
plt.rcParams['axes.titlesize'] = 12
plt.rcParams['xtick.labelsize'] = 9
plt.rcParams['ytick.labelsize'] = 9
plt.rcParams['legend.fontsize'] = 11
plt.rcParams['figure.titlesize'] = 13
from numpy import genfromtxt
from scipy.stats import multivariate_normal
from sklearn.metrics import f1_score
def read_dataset(filePath,delimiter=','):
return genfromtxt(filePath, delimiter=delimiter)
def feature_normalize(dataset):
mu = np.mean(dataset,axis=0)
sigma = np.std(dataset,axis=0)
return (dataset - mu)/sigma
def estimateGaussian(dataset):
mu = np.mean(dataset, axis=0)
sigma = np.cov(dataset.T)
return mu, sigma
def multivariateGaussian(dataset,mu,sigma):
p = multivariate_normal(mean=mu, cov=sigma)
return p.pdf(dataset)
def selectThresholdByCV(probs,gt):
best_epsilon = 0
best_f1 = 0
f = 0
stepsize = (max(probs) - min(probs)) / 1000;
epsilons = np.arange(min(probs),max(probs),stepsize)
for epsilon in np.nditer(epsilons):
predictions = (probs < epsilon)
f = f1_score(gt, predictions,average='binary')
if f > best_f1:
best_f1 = f
best_epsilon = epsilon
return best_f1, best_epsilon
tr_data = read_dataset('tr_server_data.csv')
cv_data = read_dataset('cv_server_data.csv')
gt_data = read_dataset('gt_server_data.csv')
n_training_samples = tr_data.shape[0]
n_dim = tr_data.shape[1]
print('Number of datapoints in training set: %d' % n_training_samples)
print('Number of dimensions/features: %d' % n_dim)
print(tr_data[1:5,:])
plt.xlabel('Latency (ms)')
plt.ylabel('Throughput (mb/s)')
plt.plot(tr_data[:,0],tr_data[:,1],'bx')
plt.show()
mu, sigma = estimateGaussian(tr_data)
p = multivariateGaussian(tr_data,mu,sigma)
#selecting optimal value of epsilon using cross validation
p_cv = multivariateGaussian(cv_data,mu,sigma)
fscore, ep = selectThresholdByCV(p_cv,gt_data)
print(fscore, ep)
#selecting outlier datapoints
outliers = np.asarray(np.where(p < ep))
plt.figure()
plt.xlabel('Latency (ms)')
plt.ylabel('Throughput (mb/s)')
plt.plot(tr_data[:,0],tr_data[:,1],'bx')
plt.plot(tr_data[outliers,0],tr_data[outliers,1],'ro')
plt.show()
```
## Anomaly Detection Using One-Class SVM
```
from sklearn import svm
# use the same dataset
tr_data = read_dataset('tr_server_data.csv')
clf = svm.OneClassSVM(nu=0.05, kernel="rbf", gamma=0.1)
clf.fit(tr_data)
pred = clf.predict(tr_data)
# inliers are labeled 1, outliers are labeled -1
normal = tr_data[pred == 1]
abnormal = tr_data[pred == -1]
plt.figure()
plt.plot(normal[:,0],normal[:,1],'bx')
plt.plot(abnormal[:,0],abnormal[:,1],'ro')
plt.xlabel('Latency (ms)')
plt.ylabel('Throughput (mb/s)')
plt.show()
```
| github_jupyter |
# Minicurso: Análise e Manipulação de Dados com Python: Parte 2
**Minicurso:** Análise e Manipulação de Dados com Python
**Instrutor:** Humberto da Silva Neto
**Aluno:**
## Tabela de conteúdos:
### [Parte 1:](https://github.com/hsneto/py-pandas-minicourse/blob/master/notebooks/pandas_basic.ipynb)
1. Introdução ao Pandas
2. Correção dos dados
- Identificando e lidando com valores ausentes
- Corrigindo os tipos dos dados
3. Padronização de dados
4. Normalização de dados
5. _Binning_
6. Correlação
### Parte 2:
7. [Preparando os dados](#prep)
8. [Visualização de dados usando Matplotlib](#visualizacao)
- [Gráficos de Linha](#line-plots)
- [Gráficos de Área](#area-plots)
- [Histogramas](#hist-plots)
- [Gráficos de Barras](#bar-plots)
- [Gráficos de Pizza](#pie-plots)
- [Diagrama de Caixa](#box-plots)
- [Gráfico de Dispersão](#scatter-plots)
### Outros:
- [Dicas e observações](#dicas)
- [Referências](#ref)
## Preparando os dados<a name="prep"></a>
### The Dataset: Immigration to Canada from 1980 to 2013<a name="dataset"></a>
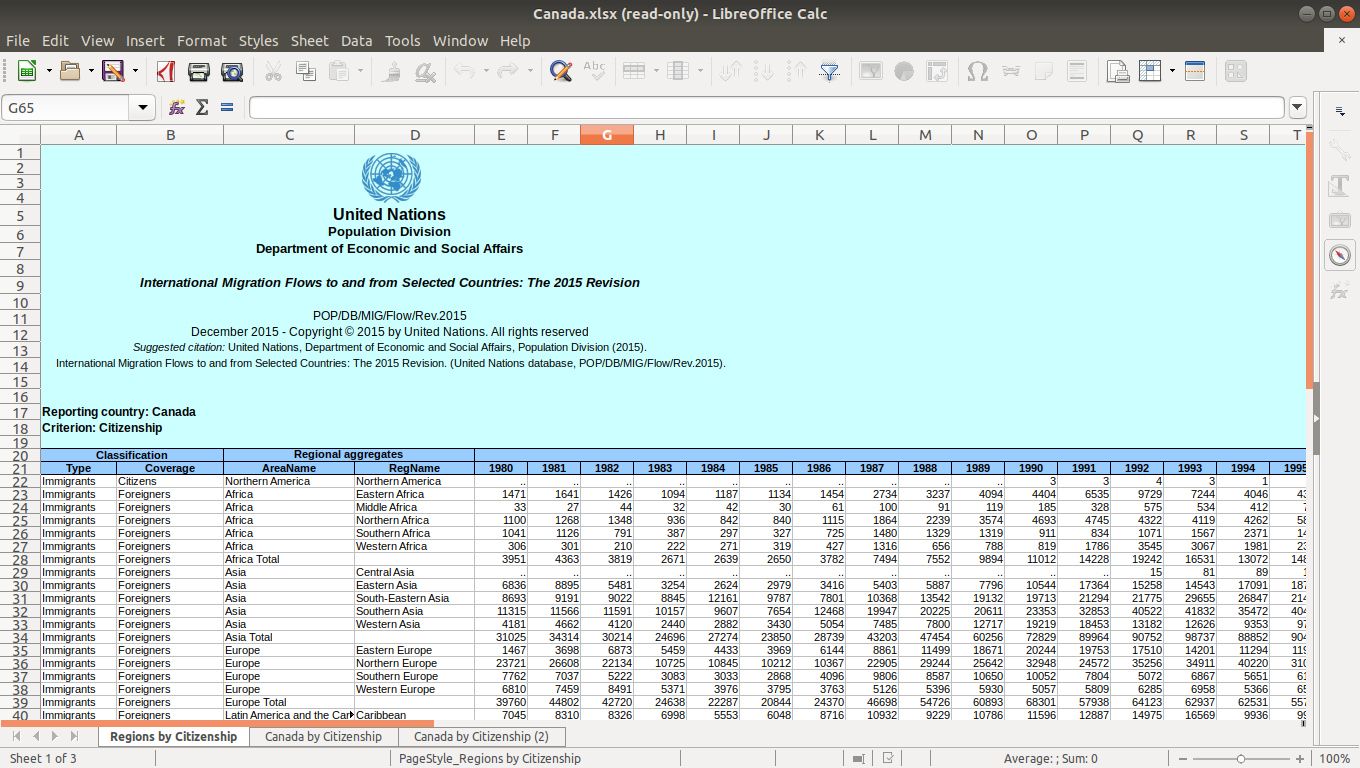
Dataset Source: [International migration flows to and from selected countries - The 2015 revision](http://www.un.org/en/development/desa/population/migration/data/empirical2/migrationflows.shtml).
O conjunto de dados contém dados anuais sobre os fluxos de imigrantes internacionais, conforme registrados pelos países de destino. Os dados apresentam entradas e saídas de acordo com o local de nascimento, cidadania ou local de residência anterior / próxima para estrangeiros e nacionais. A versão atual apresenta dados referentes a 45 países.
Neste laboratório, nos concentraremos nos dados de imigração do Canadá.
```
from __future__ import print_function
import pandas as pd
import numpy as np
# Instale o modulo requerido para para ler arquivos de excel
!pip install xlrd
print("xlrd instalado!")
# Lendo o arquivo .xlsx
xlxs = pd.ExcelFile("https://ibm.box.com/shared/static/lw190pt9zpy5bd1ptyg2aw15awomz9pu.xlsx")
df = pd.read_excel(xlxs, sheet_name="Canada by Citizenship",
skiprows=range(20), skipfooter=2)
df.head(2)
# Verifique as informacoes do dataframe
df.info()
```
### Correção da tabela<a name="clean"></a>
Vamos limpar o conjunto de dados para remover algumas colunas desnecessárias. Podemos usar o método pandas [**`df.drop()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.drop.html)$^{[1]}$ da seguinte maneira:
```python
DataFrame.drop(labels=None, axis=0, index=None, columns=None, level=None, inplace=False, errors='raise')
```
---
$^{[1]}$_**Parameters:**_
_**labels:** single label or list-like_
> _Index or column labels to drop._
_**axis:** {0 or ‘index’, 1 or ‘columns’}, default 0_
> _Whether to drop labels from the index (0 or ‘index’) or columns (1 or ‘columns’)._
_**index, columns:** single label or list-like_
> _Alternative to specifying axis (labels, axis=1 is equivalent to columns=labels)._
_**level:** int or level name, optional_
> _For MultiIndex, level from which the labels will be removed._
_**inplace:** bool, default False_
> _If True, do operation inplace and return None._
_**errors:** {‘ignore’, ‘raise’}, default ‘raise’_
> _If ‘ignore’, suppress error and only existing labels are dropped._
```
############################################################################
# TODO: Remova as colunas: 'AREA','REG','DEV','Type','Coverage' #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Verifique se as colunas foram removidas
df.head(2)
```
Vamos renomear as colunas para que façam sentido. Podemos usar o método [**`df.rename()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.rename.html) passando um dicionário de nomes antigos e novos da seguinte forma:
```python
DataFrame.rename(mapper=None, index=None, columns=None, axis=None, copy=True, inplace=False, level=None)
```
```
############################################################################
# TODO: Altere os nomes das seguintes colunas: #
# - Odname --> Country #
# - AreaName --> Continent #
# - RegName --> Region #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Verifique os nomes das colunas
df.columns
```
Também adicionaremos uma coluna `"Total"` que resume o total de imigrantes por país durante todo o período de 1980 a 2013.
---
Obs.: Esperamos os seguintes valores para as três primeiras linhas
| Country | ... | Total |
|:-----------:|:---:|-------|
| Afghanistan | ... | 58639 |
| Albania | ... | 15699 |
| Algeria | ... | 69439 |
```
############################################################################
# TODO: Crie uma coluna com o nome "Total" que representa a soma do numero #
# de imigrantes de 1980 a 2013 para todos os países. #
# #
# DICA: O comando dataframe.sum() pode ser util. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Verifique a coluna criada
df.head(2)
```
Também é possível detectar valores ausentes do dataframe com os métodos [**`df.isnull()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.isnull.html) e [**`df.isna()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.isna.html). Com isso dito, determine quantos valores ausentes temos em nossa tabela.
```
############################################################################
# TODO: Calcule quantos valores ausentes o dataframe possui #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
Finalmente, vamos ver um breve resumo de cada coluna em nosso dataframe usando o método [**`df.describe()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.describe.html).
```
# Gere estatísticas descritivas que resumem o dataframe
df.describe()
```
### Indexação e Seleção<a name="clean"></a>
**Selecionando colunas:**
Existem duas maneiras de filtrar um nome de coluna:
**Método 1:** Rápido e fácil, mas só funciona se o nome da coluna **NÃO** tiver espaços ou caracteres especiais.
```python
df.column_name
(retorna a série)
```
**Método 2:** mais robusto e pode filtrar em várias colunas.
```python
df ["coluna"]
(retorna a série)
```
```python
df [["coluna 1", "coluna 2"]]
(retorna o dataframe)
```
Obs.: Utilize o comando **`print()`** para exibir os dados extraídos nos próximos exercícios!
```
print("###################################")
print(" pandas.core.series.Series ")
print("###################################")
print("\nMETODO 1:")
############################################################################
# TODO: Utilize o metodo 1 para selecionar a coluna "Country". #
# #
# Verifique o tipo e os dados da Serie encontrada. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 2:")
############################################################################
# TODO: Utilize o metodo 2 para selecionar a coluna "Country". #
# #
# Verifique o tipo e os dados da Serie encontrada. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("####################################")
print(" pandas.core.series.DataFrame ")
print("####################################")
############################################################################
# TODO: Utilize o metodo 2 para selecionar a coluna "Country". #
# #
# Verifique o tipo e os dados do Dataframe encontrado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
############################################################################
# TODO: Utilize o metodo 2 para selecionar as colunas "Country" e "Total". #
# #
# Verifique o tipo e os dados do Dataframe encontrado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
**Selecionando linhas:**
Existem duas$^{[1]}$ maneiras de selecionar uma linha:
**Método 1:** filtra pelos rótulos do índice / coluna
```python
df.loc[label]
```
**Método 2:** filtra pelas posições do índice / coluna
```python
df.iloc[index]
```
---
$^{[1]}$ O método `df.ix[label/index]` está obsoleto!
Antes de prosseguirmos, observe que o índice _default_ do conjunto de dados é um intervalo numérico de 0 a 194. Isso dificulta muito a realização de uma consulta por um país específico. Por exemplo, para procurar dados no Japão, precisamos saber o valor do índice dele.
Outro detalhe, é que os métodos `df.loc` e `df.iloc` recebem o mesmo argumento nesse caso uma vez que o rótulo do índice é também seu index.
Para resolver esse problema, nós iremos definir a coluna **'Country'** como o índice utilizando o método [**`df.set_index()`**](https://pandas.pydata.org/pandas-docs/version/0.22/generated/pandas.DataFrame.set_index.html).
```
############################################################################
# TODO: Altere o indice da tabela usando a coluna "Country". #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Exiba o DataFrame
df.head(2)
############################################################################
# TODO: Remova o nome do index. #
# #
# DICA: O comando dataframe.index.name pode ser util. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Exiba o DataFrame
df.head(2)
print("###################################")
print(" pandas.core.series.Series ")
print("###################################")
print("\nMETODO 1:")
############################################################################
# TODO: Utilize o metodo 1 para selecionar os dados da Alemanha (Germany). #
# #
# Verifique o tipo e os dados da Serie encontrada. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 2:")
############################################################################
# TODO: Utilize o metodo 2 para selecionar os dados da Alemanha (row 67). #
# #
# Verifique o tipo e os dados da Serie encontrada. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("####################################")
print(" pandas.core.series.DataFrame ")
print("####################################")
print("\nMETODO 1:")
############################################################################
# TODO: Utilize o metodo 1 para selecionar os dados da Alemanha (Germany) #
# e do Brasil (Brazil). #
# #
# Verifique o tipo e os dados do DataFrame encontrado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 2:")
############################################################################
# TODO: Utilize o metodo 2 para selecionar os dados da Alemanha (row 67) #
# e do Brasil (row 24). #
# #
# Verifique o tipo e os dados do DataFrame encontrado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 1:")
############################################################################
# TODO: Utilizando o metodo 1, selecione somente a coluna "Total" da #
# Alemanha. #
# #
# OBS.: Imprima o dados dados obtidos. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 2:")
############################################################################
# TODO: Utilizando o metodo 2, selecione somente a coluna "Total" da #
# Alemanha. #
# #
# OBS.: Imprima o dados dados obtidos. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 1:")
############################################################################
# TODO: Utilizando o metodo 1, selecione somente a coluna "Total" da #
# Alemanha e do Brasil. #
# #
# OBS.: Imprima o dados dados obtidos. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
print("\nMETODO 2:")
############################################################################
# TODO: Utilizando o metodo 2, selecione somente a coluna "Total" da #
# Alemanha e do Brasil. #
# #
# OBS.: Imprima o dados dados obtidos. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
Para evitar ambiguidades e manter um padrão, iremos converter todos os nomes do cabeçalho para `str`.
```
def count_type(df):
"""
Imprime o numero de colunas do cabecalho que sao do tipo int ou str.
"""
c_int = 0
c_str = 0
for x in df.columns.values:
if isinstance(x, int):
c_int +=1
elif isinstance(x, str):
c_str +=1
else:
print('A coluna {} e de um tipo desconhecido'.format(x))
print('Numero de classes do tipo int: ', c_int)
print('Numero de classes do tipo str: ', c_str)
print()
# Verifica o numero de colunas do cabecalho que sao do tipo int ou str
print("Antes de transformar: \n")
count_type(df)
############################################################################
# TODO: Converta todos os nomes do cabecalho para o tipo str. #
# #
# DICA: O metodo map pode ser util. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Verifica o numero de colunas do dataframe que sao do tipo int ou str
print("Depois de transformar: \n")
count_type(df)
```
**Filtragem baseada em um critério**
Para filtrar o dataframe com base em uma condição, simplesmente passamos a condição como um vetor booleano.
Por exemplo, vamos filtrar o dataframe para mostrar os dados em países asiáticos (Continent = Asia).
```
############################################################################
# TODO: Crie uma condicao para verificar quais paises pertencem ao #
# continente: Oceania. #
############################################################################
condicao = None
############################################################################
# END OF YOUR CODE #
############################################################################
print(type(condicao), '\n')
print(condicao.head(10))
############################################################################
# TODO: Selecione os dados do dataframe que cumprem a condicao criada. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
## Visualização de dados usando Matplotlib<a name="visualizacao"></a>
A principal biblioteca de plotagem que exploraremos no curso é o Matplotlib. Como mencionado em seu site:
> O Matplotlib é uma biblioteca de plotagem 2D em Python que produz números de qualidade de publicação em uma variedade de formatos impressos e ambientes interativos entre plataformas. O Matplotlib pode ser usado em scripts Python, no shell Python e IPython, Jupyter Notebook, em servidores de aplicativos da Web e em quatro kits de ferramentas de interface gráfica com o usuário.
Se você pretende criar uma visualização impactante com o python, o Matplotlib é uma ferramenta essencial para você ter à sua disposição.
Vamos começar importando o **`Matplotlib`** e o **`Matplotlib.pyplot`** da seguinte maneira:
---
* Opcional: Aplique um estilo no Matplotlib com **`mpl.style.use`**. Para vericar as opções, use:
```python
print(plt.style.available)
```
```
import matplotlib as mpl
import matplotlib.pyplot as plt
%matplotlib inline
# Aplicando o estilo 'seaborn-darkgrid'
mpl.style.use(['seaborn-darkgrid'])
```
### Plotagem em *Pandas*
Felizmente, o Pandas têm uma implementação embutida do Matplotlib que podemos usar. Plotar em *Pandas* é tão simples quanto acrescentar um método [**`.plot()`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.plot.html) a uma série ou dataframe.
Documentação:
- [Plotting with Series](http://pandas.pydata.org/pandas-docs/stable/api.html#plotting)
- [Plotting with Dataframes](http://pandas.pydata.org/pandas-docs/stable/api.html#api-dataframe-plotting)
Desde que convertemos os anos para string, vamos declarar uma variável que nos permita recorrer facilmente a toda a gama de anos:
```
# Isso sera usado para chamar os dados de 1980 a 2014 do dataframe
years = list(map(str, range(1980, 2014)))
```
### Gráficos de linha<a name="line-plots"></a>
**O que é um gráfico de linha e por que usá-lo?**
Um gráfico de linha ou gráfico de linha é um tipo de gráfico que exibe informações como uma série de pontos de dados chamados 'marcadores' conectados por segmentos de linha reta. É um tipo básico de gráfico comum em muitos campos. Use plotagem de linha quando você tiver um conjunto de dados contínuo. Eles são mais adequados para visualizações de dados com base em tendências durante um período de tempo.
**Cuidado:**
O gráfico de linhas é uma ferramenta útil para exibir várias variáveis dependentes contra uma variável independente. No entanto, recomenda-se que não mais do que 5-10 linhas em um único gráfico; mais do que isso e torna-se difícil de interpretar.
**Vamos começar com um estudo de caso:**
Em 2010, o Haiti sofreu um terremoto catastrófico de magnitude 7,0. O terremoto causou devastação generalizada e perda de vidas e cerca de três milhões de pessoas foram afetadas por este desastre natural. Como parte do esforço humanitário do Canadá, o governo do Canadá intensificou seus esforços para aceitar refugiados do Haiti. Podemos visualizar rapidamente esse esforço usando um gráfico de linha:
---
**Questão:** Faça um gráfico de linha da imigração do Haiti usando **`.plot()`**.
[Dica]: Extraia a série de dados para o Haiti.
```
############################################################################
# TODO: Plote um grafico de linha com os dados de imigracao do Haiti entre #
# 1980 e 2014. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
*Pandas* preencheu automaticamente o eixo x com os valores do índice (anos) e o eixo y com os valores da coluna (população).
Vamos rotular os eixos x e y usando **`plt.title()`**, **`plt.ylabel()`** e **`plt.xlabel()`** da seguinte forma:
```
############################################################################
# TODO: Repita o codigo da celula acima. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration from Haiti')
plt.ylabel('Number of immigrants')
plt.xlabel('Years')
# Precisa desta linha para mostrar as atualizações feitas na figura
plt.show()
```
**Questão:** Agora compare os números de imigração entre a China e a India.
```
############################################################################
# TODO: Plote um grafico de linha com os dados de imigracao da China e da #
# India entre 1980 e 2014. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
```
Isso não parece certo ...
Lembre-se de que *Pandas* plota os índices no eixo x e as colunas como linhas individuais no eixo y. Como os dados selecionados são um dataframe com **`country`** como o índice e **`years`** como as colunas, devemos primeiro transpor o dataframe usando o método **`.transpose()`** para trocar as linhas e colunas.
```
############################################################################
# TODO: Repita o processo da celula acima e aplique o metodo .transpose no #
# dataframe selecionado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigrants from China and India')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
```
**Questão:** Compare a tendência dos cinco principais países que mais contribuíram para a imigração para o Canadá.
```
############################################################################
# TODO: Selecione os cinco paises que mais contribuiram para a imigracao #
# no Canada desde 1980. #
# #
# DICA: Tente ordenar o dataframe antes de selecionar os dados. #
# #
# OBS.: Salve esses dados em uma variavel, usaremos eles no futuro! #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
############################################################################
# TODO: Plote um grafico de linha com os dados dos cinco paises que mais #
# contribuiram para a imigracao Canada desde 1980. #
# #
# OBS.: Para melhor visualização, modifique o tamanho da figura plotada. #
# Para tal, utilize o argumento figsize=(x,y) no metodo .plot #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
```
### Gráficos de área <a name="area-plots"></a>
Gráficos de área são empilhados por padrão. E para produzir um gráfico de área empilhada, cada coluna deve ser todos os valores positivos ou negativos (quaisquer valores NaN serão padronizados como 0). Para produzir um gráfico não empilhado, passe **`stacked=False`**.
```
############################################################################
# TODO: Plote um grafico de área com os dados dos cinco paises que mais #
# contribuiram para a imigracao Canada desde 1980. #
# #
# OBS.: Defina o parametro stacked=False #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
```
O gráfico não empilhado tem uma transparência padrão (valor alpha) em 0,5. Podemos modificar esse valor passando o parâmetro **`alpha`**.
```
############################################################################
# TODO: Repita o processo da celula acima, mas dessa vez altere o valor do #
# parametro alpha e verifique o que acontece. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
```
Agora, plote um gráfico empilhado.
```
############################################################################
# TODO: Plote um grafico de área com os dados dos cinco paises que mais #
# contribuiram para a imigracao Canada desde 1980. #
# #
# OBS.: Defina o parametro stacked=True #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration Trend of Top 5 Countries')
plt.ylabel('Number of Immigrants')
plt.xlabel('Years')
plt.show()
```
### Histogramas <a name="hist-plots"></a>
Um histograma é uma maneira de representar a distribuição de frequência do conjunto de dados numéricos. A maneira como funciona é dividir o eixo x em compartimentos, atribuir cada ponto de dados em nosso conjunto de dados a um bloco e, em seguida, contar o número de pontos de dados que foram atribuídos a cada bloco. Portanto, o eixo y é a frequência ou o número de pontos de dados em cada caixa. Observe que podemos alterar o tamanho da caixa e, geralmente, é necessário ajustá-lo para que a distribuição seja exibida corretamente.
**Qustão:** Qual é a distribuição de frequência do número (população) de novos imigrantes dos vários países para o Canadá em 2013?
Antes de prosseguirmos com a criação do gráfico do histograma, vamos primeiro examinar os dados divididos em intervalos. Para fazer isso, nos usaremos o método de histrograma do Numpy para obter os intervalos de bin e as contagens de frequência da seguinte maneira:
```
# Determine o histograma do ano de 2013
count, bin_edges = np.histogram(df['2013'])
print(count)
print(bin_edges)
```
Por padrão, o método **`histrogram`** divide o dataset em 10 blocos. A figura abaixo resume a distribuição de frequência da imigração em 2013 para cada bloco. Podemos ver que em 2013:
* 178 países contribuíram com 0 a 3412,9 imigrantes
* 11 países contribuíram entre 3412,9 e 6825,8 imigrantes
* 1 país contribuiu entre 6285,8 e 10238,7 imigrantes e assim por diante.
<img src = "https://ibm.box.com/shared/static/g54s9q97mrjok0h4272o7g09cyigei0v.jpg" align="center" width=800>
```
############################################################################
# TODO: Plote o histograma referente a imigracao no ano de 2013. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Histogram of Immigration from 195 Countries in 2013')
plt.ylabel('Number of Countries')
plt.xlabel('Number of Immigrants')
plt.show()
```
Na plotagem acima, o eixo x representa a faixa populacional de imigrantes em intervalos de 3412,9. O eixo y representa o número de países que contribuíram para a população mencionada.
Observe que os rótulos do eixo x não correspondem ao tamanho dos intervalos. Isso pode ser corrigido passando-se uma palavra-chave **`xticks`** que contém a lista dos tamanhos desses intervalos.
```
############################################################################
# TODO: Plote o histograma referente a imigracao no ano de 2013, mas desta #
# vez altere o parametro xticks para corrigir o tamanho dos intervalos. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Histogram of Immigration from 195 Countries in 2013')
plt.ylabel('Number of Countries')
plt.xlabel('Number of Immigrants')
plt.show()
```
**Questão:** Qual é a distribuição de imigração para a Dinamarca, Noruega e Suécia para os anos 1980 - 2013?
```
############################################################################
# TODO: Plote o histograma referente a imigracao da Dinamarca (Denmark), #
# Noruega (Norway) e Suécia (Sweden) entre 1980 e 2013. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
```
Ao invés de traçar a distribuição da frequência populacional da população para os 3 países, o *pandas* traçou a distribuição da frequência populacional para os anos.
Isso pode ser facilmente corrigido, primeiro transpondo o conjunto de dados e, em seguida, plotando.
```
############################################################################
# TODO: Corriga o problema da celula acima e plote novamente o histograma. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
```
Vamos fazer algumas modificações para melhorar o impacto e a estética do plot anterior:
* Aumentar o tamanho do bin para 15 passando o parâmetro **`bins`**
* defina a transparência para 60% passando o parâmetro **`alpha`**
* rotule o eixo x passando no parâmetro **`x-label`**
* mudar as cores dos gráficos passando no parâmetro **`color`**
```
# Vamos pegar os valores de x-tick
df_aux = df.loc[['Denmark', 'Norway', 'Sweden'], years].transpose()
count, bin_edges = np.histogram(df_aux, 15)
# Histograma não empilhado
df_aux.plot(kind ='hist',
figsize=(10, 6),
bins=15,
alpha=0.6,
xticks=bin_edges,
color=['coral', 'darkslateblue', 'mediumseagreen'])
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
del df_aux
```
Se não quisermos que os gráficos se sobreponham uns aos outros, podemos empilhá-los usando o parâmetro **stacked=True``**.
```
############################################################################
# TODO: Baseando-se na celula acima, crie um histograma empilhado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Histogram of Immigration from Denmark, Norway, and Sweden from 1980 - 2013')
plt.ylabel('Number of Years')
plt.xlabel('Number of Immigrants')
plt.show()
```
### Gráficos de barras <a name="bar-plots"></a>
Um gráfico de barras é uma maneira de representar dados em que o * comprimento * das barras representa a magnitude / tamanho do recurso / variável. Gráficos de barras geralmente representam variáveis numéricas e categóricas agrupadas em intervalos.
Para criar um gráfico de barras, podemos passar um dos dois argumentos através do parâmetro **`kind`** em **`plot()`**:
* **`kind = bar`** $\to$ cria um gráfico de barra vertical
* **`kind = barh`** $\to$ cria um gráfico de barras horizontal
#### Gráficos de barras verticais
Nos gráficos de barras verticais, o eixo x é usado para rotular e o comprimento das barras no eixo y corresponde à magnitude da variável sendo medida. Gráficos de barras verticais são particularmente úteis na análise de dados de séries temporais. Uma desvantagem é que eles não têm espaço para a rotulagem de texto no pé de cada barra.
** Vamos começar analisando o efeito da crise financeira da Islândia: **
A crise financeira islandesa de 2008 a 2011 foi um importante evento econômico e político na Islândia. Em relação ao tamanho de sua economia, o colapso bancário sistêmico da Islândia foi o maior experimentado por qualquer país na história econômica. A crise levou a uma grave depressão econômica em 2008 - 2011 e uma agitação política significativa.
** Questão: ** Vamos comparar o número de imigrantes islandeses (país = "Iceland") ao Canadá do ano de 1980 a 2013.
[Dica]: Extraia a série de dados para a Islândia.
```
############################################################################
# TODO: Plote um grafico de barras com os dados de imigracao da Islandia #
# (Iceland) entre 1980 e 2014. #
# #
# OBS.: Altere o parametro color e defina uma cor para as barras. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.xlabel('Year')
plt.ylabel('Number of immigrants')
plt.title('Icelandic immigrants to Canada from 1980 to 2013')
plt.show()
```
**Extra:** Adicione o código abaixo antes de **`plt.show()`**
```python
# Anotar seta
plt.annotate('', # s: str. Deixa em branco para nenhum texto
xy=(32, 70), # coloque a ponta da seta no ponto (ano de 2012, pop 70)
xytext=(28, 20), # coloque a base da seta no ponto (ano de 2008, pop 20)
xycoords='data', # usará o sistema de coordenadas do objeto que está sendo anotado
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2))
# Anotar texto
plt.annotate('2008 - 2011 Financial Crisis', # texto a ser exibido
xy=(28,30), # comece o texto no ponto (ano de 2008, pop 30)
rotation=72.5, # Baseado em tentativa e erro para coincidir com a seta
va='bottom', # Deseja que o texto seja alinhado verticalmente na parte inferior
ha='left') # Deseja que o texto seja horizontalmente 'deixado' algned.
```
```
############################################################################
# TODO: Repita o codigo da celula acima #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Anotar seta
plt.annotate('', # s: str. Deixa em branco para nenhum texto
xy=(32, 70), # coloque a ponta da seta no ponto (ano de 2012, pop 70)
xytext=(28, 20), # coloque a base da seta no ponto (ano de 2008, pop 20)
xycoords='data', # usará o sistema de coordenadas do objeto que está sendo anotado
arrowprops=dict(arrowstyle='->', connectionstyle='arc3', color='blue', lw=2))
# Anotar texto
plt.annotate('2008 - 2011 Financial Crisis', # texto a ser exibido
xy=(28,30), # comece o texto no ponto (ano de 2008, pop 30)
rotation=72.5, # Baseado em tentativa e erro para coincidir com a seta
va='bottom', # Deseja que o texto seja alinhado verticalmente na parte inferior
ha='left') # Deseja que o texto seja horizontalmente 'deixado' algned.
plt.xlabel('Year')
plt.ylabel('Number of immigrants')
plt.title('Icelandic immigrants to Canada from 1980 to 2013')
plt.show()
```
#### Gráficos de barras horizontais
Às vezes é mais prático representar os dados horizontalmente, especialmente se você precisar de mais espaço para rotular as barras. Nos gráficos de barras horizontais, o eixo y é usado para rotular e o comprimento das barras no eixo x corresponde à magnitude da variável sendo medida. Como você verá, há mais espaço no eixo y para rotular variáveis categóricas.
**Questão:** Usando o conjunto de dados **df**, crie um gráfico de barras horizontais mostrando o número total de imigrantes para o Canadá dos 15 países que mais imigraram, para o período de 1980 a 2013. Rotule cada país com a contagem total de imigrantes.
```
############################################################################
# TODO: Plote um grafico de barras horizontais com o "Total" dos 15 paises #
# que mais contribuiram para a imigracao no Canada desde 1980. #
# #
# OBS.: Salve esses dados na variavel df_top15 #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# anotar rótulos de valor para cada país
for index, value in enumerate(df_top15):
label = format(int(value), ',')
# coloque o texto no final da barra (subtraindo 47000 de x e 0,1 de y para ajustá-lo dentro da barra)
plt.annotate(label, xy=(value - 47000, index - 0.10), color='white')
plt.xlabel('Number of Immigrants')
plt.title('Top 15 Conuntries Contributing to the Immigration to Canada between 1980 - 2013')
plt.show()
```
### Gráficos de Pizza <a name="pie-plots"></a>
Um `gráfico de pizza` é um gráfico circular que exibe proporções numéricas dividindo um círculo (ou pizza) em fatias proporcionais. Você provavelmente já está familiarizado com gráficos de pizza, pois é amplamente utilizado em negócios e mídia. Podemos criar gráficos de pizza no Matplotlib passando a palavra-chave **`kind = pie`**.
**Questão:** Usando um gráfico de pizza, explore a proporção (porcentagem) de novos imigrantes agrupados por continentes no ano de 2013.
---
**Etapa 1:** Coletar dados.
Vamos usar o método *pandas * [**`dataframe.groupby`**](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.groupby.html) para resumir os dados de imigração por Continente. O processo geral do **`groupby`** envolve os seguintes passos:
1. ** Dividir: ** Dividindo os dados em grupos com base em alguns critérios.
2. ** Aplicar: ** Aplicando uma função para cada grupo de forma independente:
.soma()
.contagem()
.significar()
.std ()
.agregar()
.Aplique()
.etc ..
3. ** Combinar: ** Combinando os resultados em uma estrutura de dados.
<img src = "https://ibm.box.com/shared/static/tkfhxqkehfzpclco8f0eazhie33uxj9j.png" align="center" height=300>
```
############################################################################
# TODO: Agrupe os países por continentes. #
############################################################################
df_continents = None
############################################################################
# END OF YOUR CODE #
############################################################################
# Nota: a saída do método groupby é um objeto groupby.
# não podemos usá-lo até aplicarmos uma função (por exemplo, .sum ())
print(type(df_continents))
############################################################################
# TODO: Agrupe os países por continentes novamente. Dessa vez aplique a #
# funcao .sum() #
############################################################################
df_continents = None
############################################################################
# END OF YOUR CODE #
############################################################################
df_continents.head()
```
**Etapa 2:** plote os dados. Vamos passar a palavra-chave **`kind = "pie"`**, juntamente com os seguintes parâmetros adicionais:
- **`autopct`** - é uma string ou função usada para rotular o valor numérico. O rótulo será colocado dentro da máscara (por exemplo, controlar o número de casas decimais do rótulo). Se for uma string de formato, o rótulo será `fmt% pct`.
- **`startangle`** - gira o início do gráfico de pizza em graus de ângulo no sentido anti-horário a partir do eixo x.
- **`shadow`** - Desenha uma sombra sob a pizza (para dar uma sensação 3D).
```
############################################################################
# TODO: Plote um grafico de pizza agrupando o total de imigracoes desde #
# 1980 por continente. #
# #
# OBS.: Sinta-se livre para alterar os parametros adicionais citados acima.#
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration to Canada by Continent [1980 - 2013]')
plt.axis('equal') # Define o gráfico de pizza para se parecer com um círculo.
plt.show()
```
O visual acima não é muito claro, os números e o texto se sobrepõem em alguns casos. Vamos fazer algumas modificações para melhorar o visual:
* Remova os rótulos de texto no gráfico de pizza passando em "legend" e adicione-o como uma legenda separada usando **`plt.legend()`**.
* Empurre as porcentagens para ficar fora do gráfico de pizza passando o parâmetro **`pctdistance`**.
* Passar em um conjunto personalizado de cores para os continentes, passando no parâmetro **`colors`**.
* Explode o gráfico de pizza para enfatizar os três continentes mais baixos (África, América do Norte e América Latina e Caribe) passando o parâmetro **`explode`**.
```
colors_list = ['gold', 'yellowgreen', 'lightcoral', 'lightskyblue', 'lightgreen', 'pink']
explode_list = [0.1, 0, 0, 0, 0.1, 0.1]
df_continents['Total'].plot(kind='pie',
figsize=(15, 6),
autopct='%1.1f%%',
startangle=90,
shadow=True,
labels=None,
pctdistance=1.12,
colors=colors_list,
explode=explode_list)
# Escale o título em 12% para corresponder à pctdistance
plt.title('Immigration to Canada by Continent [1980 - 2013]', y=1.12)
plt.axis('equal')
plt.legend(labels=df_continents.index, loc='upper left')
plt.show()
```
### Diagrama de caixa <a name="box-plots"></a>
Um diagrama de caixa é uma forma de representar estatisticamente a distribuição dos dados através de cinco dimensões principais:
- ** Minimun: ** O menor número no conjunto de dados.
- ** Primeiro quartil: ** Número do meio entre o "mínimo" e o "mediano".
- ** Segundo quartil (Mediana): ** Número do meio do conjunto de dados (classificado).
- ** Terceiro quartil: ** Número do meio entre "mediano" e "máximo".
- ** Máximo: ** Maior número no conjunto de dados.
<img src = "https://ibm.box.com/shared/static/9nkxsfihu8mgt1go2kfasf61sywlu123.png" width=440, align = "center">
Para fazer uma plotagem de caixa, podemos usar kind = box no método de plotagem chamado em uma série ou dataframe.
**Questão:** Escreva a caixa para os imigrantes japoneses entre 1980 - 2013.
---
**Etapa 1:**
Obtenha o conjunto de dados. Mesmo que estejamos puxando informações para apenas um país, vamos obtê-lo como um dataframe. Isso nos ajudará a chamar o método **`dataframe.describe()`** para visualizar os percentis.
```
############################################################################
# TODO: Plote um diagrama de caixa sobre os dados do Japao (Japan). Antes #
# disso, salve os dados do Japao em uma variavel chamada df_japan #
# #
# OBS.: Selecione os dados do Japao como DataFrame e nao como uma Serie! #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Box plot of Japanese Immigrants from 1980 - 2013')
plt.ylabel('Number of Immigrants')
plt.show()
```
Podemos fazer imediatamente algumas observações importantes do gráfico acima:
1. O número mínimo de imigrantes é de cerca de 200 (min), o número máximo é de cerca de 1300 (max) e o número médio de imigrantes é de cerca de 900 (mediana).
2. 25% dos anos para o período de 1980 a 2013 tiveram uma contagem anual de imigrantes de ~ 500 ou menos (primeiro quartil).
3. 75% dos anos para o período de 1980 a 2013 tiveram uma contagem anual de imigrantes de ~ 1100 ou menos (terceiro quartil).
Podemos ver os números reais chamando o método **`describe()`** no dataframe.
```
df_japan.describe()
```
Um dos principais benefícios dos box plots é comparar a distribuição de vários conjuntos de dados. Em um dos módulos anteriores, observamos que a China e a Índia tinham tendências de imigração muito semelhantes. Vamos analisar mais usando gráficos de caixa.
**Questão:** Compare a distribuição do número de novos imigrantes da Índia e da China para o período de 1980 a 2013.
```
############################################################################
# TODO: Plote um diagrama de caixa sobre os dados do da China e da India #
# para comparar. Confira os dados do diagrama com o metodo .describe #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Box plots of Immigrants from China and India (1980 - 2013)')
plt.xlabel('Number of Immigrants')
plt.show()
```
Podemos observar que, embora ambos os países tenham aproximadamente a mesma população média de imigrantes (~ 20.000), a faixa populacional de imigrantes na China é mais disseminada do que a da Índia. A população máxima da Índia para qualquer ano (36.210) é cerca de 15% menor que a população máxima da China (42.584).
```
# crie uma figura
fig = plt.figure()
# crie as regios do subplot
ax0 = fig.add_subplot(121)
ax1 = fig.add_subplot(122)
############################################################################
# TODO: Com o DataDrame criado com os dados da China e da India, plote um #
# grafico de linhas e um diagrama de caixa lado a lado. #
# #
# OBS.: Utilize o parametro ax para definir a regiao da figura em que o #
# grafico sera plotado. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
ax0.set_title('Box Plots of Immigrants from China and India (1980 - 2013)')
ax0.set_xlabel('Number of Immigrants')
ax0.set_ylabel('Countries')
ax1.set_title ('Line Plots of Immigrants from China and India (1980 - 2013)')
ax1.set_ylabel('Number of Immigrants')
ax1.set_xlabel('Years')
plt.show()
```
**Questão:** Crie uma caixa para visualizar a distribuição dos 15 principais países (com base na imigração total) agrupados nas décadas de 1980, 1990 e 2000.
```
# Criando uma lista de todos os anos nas decada de 80 e 90
years_80s = list(map(str, range(1980, 1990)))
years_90s = list(map(str, range(1990, 2000)))
years_00s = list(map(str, range(2000, 2010)))
############################################################################
# TODO: Selecione os dados dos 15 paises que mais imigraram para o Canada. #
# Agrupe-os pelo numero total de imigracoes em cada decada. Para isso, #
# salve esses valores em tres Series distintas: #
# - df_80s : Serie contendo o total de imigracoes entre 1980 e 1990 #
# - df_90s : Serie contendo o total de imigracoes entre 1990 e 2000 #
# - df_00s : Serie contendo o total de imigracoes entre 2000 e 2010 #
# #
# OBS.: As variaveis criadas acima (years_80s, years_90s, years_00s) podem #
# ser uteis. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
# Mesclar as Series criadas em um novo DataFrame
df_aux = pd.DataFrame({'1980s': df_80s, '1990s': df_90s, '2000s':df_00s})
print(df_aux)
df_aux.describe()
############################################################################
# TODO: Plote um diagrama de caixas do DataFrame criado acima. #
# #
# OBS.: Se necessario, utilize o parametro sym para exibir os outliers. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Immigration from top 15 countries for decades 80s, 90s and 2000s')
plt.show()
del df_aux
```
### Gráfico de dispersão <a name="scatter-plots"></a>
Um gráfico de dispersão (2D) é um método útil para comparar variáveis entre si. Os gráficos de dispersão são semelhantes aos gráficos de linhas, pois ambos mapeiam variáveis independentes e dependentes em um gráfico 2D. Embora os pontos de dados estejam conectados por uma linha em um gráfico de linhas, eles não estão conectados em um gráfico de dispersão. Os dados em um gráfico de dispersão são considerados para expressar uma tendência. Com uma análise mais aprofundada usando ferramentas como a regressão, podemos calcular matematicamente esse relacionamento e usá-lo para prever tendências fora do conjunto de dados.
**Questão:** Usando um gráfico de dispersão, visualize a tendência da imigração total para o Canadá (todos os países combinados) para os anos 1980 - 2013.
**Etapa 1:**
Obtenha o conjunto de dados. Como esperamos usar a relação entre anos e população total, converteremos anos em tipo flutuante.
```
# Podemos usar o método sum() para obter a população total por ano
df_tot = pd.DataFrame(df[years].sum(axis=0))
# Altere os anos para float (util para regressão mais tarde)
df_tot.index = map(float,df_tot.index)
# Redefinir o índice
df_tot.reset_index(inplace = True)
# Renomear colunas
df_tot.columns = ['year', 'total']
# Veja o dataframe final
df_tot.head()
```
**Etapa 2:**
Plotar dados. No `Matplotlib`, podemos criar um gráfico de dispersão ao passar em **`kind = "scatter"`** como argumento de plotagem. Também precisaremos passar as palavras-chave **`x`** e **`y`** para especificar as colunas que vão nos eixos x e y.
```
############################################################################
# TODO: Plote um grafico de dispersao do DataFrame df_tot. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Total Immigration to Canada from 1980 - 2013')
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
plt.show()
```
Observe como o gráfico de dispersão não conecta os pontos de dados juntos. Podemos observar claramente uma tendência ascendente nos dados: à medida que os anos passam, o número total de imigrantes aumenta. Podemos analisar matematicamente essa tendência ascendente usando uma linha de regressão (linha de melhor ajuste).
**Questão:** Faça uma linha linear de melhor ajuste e use-a para prever o número de imigrantes em 2015.
---
**Etapa 1:** obtenha a equação da linha de melhor ajuste. Vamos usar o método **`polyfit()`** do ** Numpy ** passando o seguinte:
- **`x`:** coordenadas x dos dados.
- **`y`:** coordenadas y dos dados.
- **`deg`: ** Grau de ajuste polinomial. 1 = linear, 2 = quadrático e assim por diante
```
x = df_tot.year
y = df_tot.total
fit = np.polyfit(x, y, deg=1)
print(fit)
```
A saída é uma matriz com os coeficientes polinomiais, as maiores potências primeiro. Como estamos plotando uma regressão linear **`y = a*x + b`**, nossa saída tem 2 elementos **`[5.56709228e + 03, -1.09261952e + 07]`** com a inclinação na posição 0 e intercepta na posição 1.
**Etapa 2: **
Plote a linha de regressão no gráfico de dispersão.
```
############################################################################
# TODO: Plote um grafico de dispersao do DataFrame df_tot. #
############################################################################
pass
############################################################################
# END OF YOUR CODE #
############################################################################
plt.title('Total Immigration to Canada from 1980 - 2013')
plt.xlabel('Year')
plt.ylabel('Number of Immigrants')
# Plote a linha de melhor ajuste
plt.plot(x,fit[0]*x + fit[1], color='red') # recall that x is the Years
plt.annotate('y={0:.0f} x + {1:.0f}'.format(fit[0], fit[1]), xy=(2000, 150000))
plt.show()
# Imprima a linha de melhor ajuste
print('No. Immigrants = {0:.0f} * Year + {1:.0f}'.format(fit[0], fit[1]) )
```
Usando a equação da linha de melhor ajuste, podemos estimar o número de imigrantes em 2015:
`` `python
Não. Imigrantes = 5567 * Ano - 10926195
Não. Imigrantes = 5567 * 2015 - 10926195
Não. Imigrantes = 291.310
`` `
Quando comparado com os dados reais do Relatório Anual 2016 da Cidadania e Imigração (CIC) (http://www.cic.gc.ca/english/resources/publications/annual-report-2016/index.asp), que o Canadá aceitou 271.845 imigrantes em 2015. Nosso valor estimado de 291.310 está dentro de 7% do número real, o que é muito bom considerando que nossos dados originais vieram das Nações Unidas (e podem diferir ligeiramente dos dados do CIC).
Como uma nota lateral, podemos observar que a imigração deu um mergulho em torno de 1993-1997. Uma análise mais aprofundada sobre o tema revelou que em 1993 o Canadá interveio Bill C-86, que introduziu revisões no sistema de determinação de refugiados, principalmente restritivo. Outras emendas aos Regulamentos de Imigração cancelaram o patrocínio requerido para "parentes assistidos" e reduziram os pontos atribuídos a eles, tornando mais difícil para os membros da família (exceto família nuclear) imigrar para o Canadá. Estas medidas restritivas tiveram um impacto direto nos números da imigração nos próximos anos.
---
Parabéns, você chegou ao fim do minicurso!
---

---
## Dicas e observações:<a name="dicas"></a>
### Utilizando o Google Colaboratory:
#### 1. Comandos no terminal:
Para utilizar qualquer comando no terminal, comece com um **`!`**. Por exemplo, para mostrar os arquivos do diretório atual utilize
```sh
!ls
```
#### 2. Baixar biliotecas:
Em caso do módulo não estar instalado na máquina, utilize o comando
```python
!pip install nome_do_modulo
```
#### 3. Importando arquivos:
Para importar arquivos da sua máquina para o Colaboratory, o comando abaixo pode ser útil
```pyhon
from google.colab import files
uploaded = files.upload()
```
Para importar de seu drive, o comando abaixo pode ser útil
```python
!apt-get install -y -qq software-properties-common python-software-properties module-init-tools
!add-apt-repository -y ppa:alessandro-strada/ppa 2>&1 > /dev/null
!apt-get update -qq 2>&1 > /dev/null
!apt-get -y install -qq google-drive-ocamlfuse fuse
from google.colab import auth
auth.authenticate_user()
from oauth2client.client import GoogleCredentials
creds = GoogleCredentials.get_application_default()
import getpass
!google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret} < /dev/null 2>&1 | grep URL
vcode = getpass.getpass()
!echo {vcode} | google-drive-ocamlfuse -headless -id={creds.client_id} -secret={creds.client_secret}
!mkdir -p drive
!google-drive-ocamlfuse drive
```
---
## Referências: <a name="ref"></a>
### Livros:
- [Python for Data Analysis, 2nd Edition](http://shop.oreilly.com/product/0636920050896.do)
### Cursos onlines:
- [Cognitive Class (IBM): *Applied Data Science with Python*](https://cognitiveclass.ai/learn/data-science-with-python/)
| github_jupyter |
```
import matplotlib.pyplot as plt
%matplotlib inline
import pandas as pd
import numpy as np
headers = ['episode_num', 'success', 'total_reward', 'total_steps', 'exploration_steps', 'epsilon']
dtypes = {'episode_num': 'int', 'success': 'str', 'total_reward': 'int', 'total_steps': 'int', 'explorations': 'int', 'epsilon': float}
data = pd.read_csv('qtables7_the_one/episodes.txt', header=None, names=headers, dtype=dtypes)
data['success_bool'] = True
data['success_int'] = 0
for i, v in enumerate(data['success'].values):
if v.strip() == 'False':
data.set_value(i, 'success_bool', False)
data.set_value(i, 'success_int', 0)
else:
data.set_value(i, 'success_bool', True)
data.set_value(i, 'success_int', 1)
data.tail()
episode_success = data.iloc[:, [0, 6]].values
episode_reward = data.iloc[:, [0, 2]].values
episode_steps = data.iloc[:, [0, 3]].values
episode_explorations = data.iloc[:, [0, 4]].values
episode_success[:, 1]
np.min(episode_reward[:,1])
# Success
plt.figure(figsize=(15, 3))
b = plt.bar(episode_success[:, 0], episode_success[:, 1], width=1)
for i in range(len(episode_success)):
if episode_explorations[i][1] == 0 and episode_success[i][1]:
b[i].set_color('blue')
b[i].set_linewidth(1)
else:
b[i].set_color('skyblue')
plt.xlabel('Episodes')
plt.ylabel('Success')
plt.title('Episode vs Success')
plt.show()
# Reward
plt.figure(figsize=(15, 5))
b = plt.bar(episode_reward[:, 0], episode_reward[:, 1], width=0.2)
for i in range(len(episode_reward)):
if episode_success[i][1] and episode_explorations[i][1] == 0:
b[i].set_color('b')
b[i].set_linewidth(1)
elif episode_success[i][1] and episode_explorations[i][1] > 0:
b[i].set_color('skyblue')
b[i].set_linewidth(1)
else:
b[i].set_color('lightgray')
b[i].set_linewidth(0.2)
plt.xlabel('Episodes')
plt.ylabel('Total Reward (more the better)')
plt.title('Episode vs Reward')
plt.show()
# Exploration vs Exploitation
plt.figure(figsize=(15, 5))
b = plt.bar(episode_steps[:, 0], episode_steps[:, 1], width=1, color=['lightgray'])
for i in range(len(episode_steps)):
if episode_success[i][1] and episode_explorations[i][1] == 0:
b[i].set_color('b')
b[i].set_linewidth(2)
elif episode_success[i][1] and episode_explorations[i][1] > 0:
b[i].set_color('skyblue')
b[i].set_linewidth(2)
#else:
# b[i].set_color('lightgray')
plt.xlabel('Episodes')
plt.ylabel('Total Actions (lesser the better)')
plt.title('Episode vs Total Action')
plt.show()
```
| github_jupyter |
<table width="100%">
<tr style="border-bottom:solid 2pt #009EE3">
<td style="text-align:left" width="10%">
<a href="FILENAME" download><img src="../../images/icons/download.png"></a>
</td>
<td style="text-align:left" width="10%">
<a href="SOURCE" target="_blank"><img src="../../images/icons/program.png" title="Be creative and test your solutions !"></a>
</td>
<td></td>
<td style="text-align:left" width="5%">
<a href="../MainFiles/biosignalsnotebooks.ipynb"><img src="../../images/icons/home.png"></a>
</td>
<td style="text-align:left" width="5%">
<a href="../MainFiles/contacts.ipynb"><img src="../../images/icons/contacts.png"></a>
</td>
<td style="text-align:left" width="5%">
<a href="https://github.com/biosignalsnotebooks/biosignalsnotebooks" target="_blank"><img src="../../images/icons/github.png"></a>
</td>
<td style="border-left:solid 2pt #009EE3" width="15%">
<img src="../../images/ost_logo.png">
</td>
</tr>
</table>
<link rel="stylesheet" href="../../styles/theme_style.css">
<!--link rel="stylesheet" href="../../styles/header_style.css"-->
<link rel="stylesheet" href="https://cdnjs.cloudflare.com/ajax/libs/font-awesome/4.7.0/css/font-awesome.min.css">
<table width="100%">
<tr>
<td id="image_td" width="15%" class="header_image_color_2"><div id="image_img"
class="header_image_2"></div></td>
<td class="header_text"> Signal Acquisition [OpenSignals] </td>
</tr>
</table>
<div id="flex-container">
<div id="diff_level" class="flex-item">
<strong>Difficulty Level:</strong> <span class="fa fa-star checked"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
<span class="fa fa-star"></span>
</div>
<div id="tag" class="flex-item-tag">
<span id="tag_list">
<table id="tag_list_table">
<tr>
<td class="shield_left">Tags</td>
<td class="shield_right" id="tags">record☁acquire☁opensignals☁real-time</td>
</tr>
</table>
</span>
<!-- [OR] Visit https://img.shields.io in order to create a tag badge-->
</div>
</div>
<a href="https://www.biosignalsplux.com/en/software" target="_blank"><span class="color1"><strong>OpenSignals <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a> is the Plux's software dedicated to acquire, store and process physiological signals acquired with electronic devices developed and commercialized by the company (such as <a href="https://www.biosignalsplux.com/en/" target="_blank"><span class="color2"><strong>biosignalsplux <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a> and <a href="https://bitalino.com/en/" target="_blank"><span class="color4"><strong>bitalino <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></span></a>).
Signal acquisition is a fundamental step that needs to be executed before every processing task. Without signals there are not any information to analyze or knowledge to extract.
Like previously referred, <span class="color1"><strong>OpenSignals</strong></span> ensures that anyone with a Plux's acquisition system can easily acquire physiological data through a desktop or mobile device.
So, in the current <span class="color4"><strong>Jupyter Notebook</strong></span> we will begin our introductory journey through <span class="color1"><strong>OpenSignals</strong></span>, explaining/demonstrating how signals can be acquired in real-time.
<hr>
Before starting an acquisition it is mandatory that your Plux acquisition system (in our case <span class="color1"><strong>biosignalsplux</strong></span>) is paired with the computer.<br>There is available a <span class="color4"><strong>Jupyter Notebook</strong></span> entitled <a href="../Connect/pairing_device.ipynb" target="_blank"><strong>"Pairing a Device at Windows 10 [biosignalsplux]" <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></a> intended to help you in the pairing process between <span class="color1"><strong>biosignalsplux</strong></span> and your computer.
<p class="steps">0 - Execute steps 8 to 10 of <a href="../Connect/pairing_device.ipynb" target="_blank"><strong>"Pairing a Device at Windows 10 [biosignalsplux]" <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></strong></a>, in order to enable your device before starting the acquisition</p>
<p class="steps">1 - Click on the device box. A set of configurable options will appear</p>
<img src="../../images/other/opensignals_dev_click.gif">
<p class="steps">2 - For the current example we will pair an accelerometer (triaxial sensor). Three channels of our <span class="color1"><strong>biosignalsplux</strong></span> device are necessary for doing the acquisition <br>(Channel 1 >> Axis X, Channel 2 >> Axis Y, Channel 3 >> Axis Z)</p>
<img src="../../images/record/biosignalsplux_acc.png" width="50%">
<p class="steps">3 - Activate the used channels and select the respective sensor type </p>
<i>When clicking on our device we can see lots of options and components. Let's focus on the left section of the following image</i>
<p class="steps">3.1 - Activate the used channels</p>
For activating the channels that will be used during the acquisition just click on the "Status Circle" of the respective channel row.
<img src="../../images/record/signal_acquisition_sel_chn.gif">
<p class="steps">3.2 - Select the respective sensor type for each channels (for our accelerometer example, we need to select option "XYZ")</p>
For selecting the desired sensor type you should press on the downside arrow to see a list box containing the available options.
<img src="../../images/record/signal_acquisition_sel_type.gif">
<p class="steps">4 - Specify the desired sampling rate, taking into consideration the signal that you will acquire</p>
For a more deep information about sampling rate choice, there is a <span class="color4"><strong>Jupyter Notebook</strong></span> available in our <a href="../Record/sampling_rate_and_aliasing.ipynb" target="_blank">library <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a>.
While using accelerometer sensor a minimum sampling rate of 100 Hz should be chosen (accordingly to the Nyquist Theorem as the double of the maximum frequency in the sensor bandwidth | <a href="https://www.biosignalsplux.com/en/acc-accelerometer" target="_blank">0-50 Hz <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a>)
<img src="../../images/record/signal_acquisition_sel_sr.gif">
<p class="steps">5 - Select the device resolution</p>
<i>Bigger resolutions ensures a more precise data acquisition but there is the risk of collecting also more <a href="http://www.lionprecision.com/tech-library/technotes/article-0010-sensor-resolution.html" target="_blank">noise <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a> !</i>
<img src="../../images/record/signal_acquisition_sel_res.gif">
<p class="steps">6 - There is only one remaining step left. Just press "Record" button to start acquiring your accelerometer data</p>
<img src="../../images/record/signal_acquisition_rec.png">
The previous steps ensure that all users of Plux's acquisition systems can start their acquisition establishing an important step for beginning the experimental stage of any research.
For concluding, is shown a small video of a real-time accelerometer acquisition made with <span class="color1"><strong>OpenSignals</strong></span> !
<video id="video_1" muted loop src="../../images/record/signal_acquisition_video.mp4" class="video"></video>
```
%%javascript
document.getElementById("video_1").play()
```
<strong><span class="color7">We hope that you have enjoyed this guide. </span><span class="color2">biosignalsnotebooks</span><span class="color4"> is an environment in continuous expansion, so don't stop your journey and learn more with the remaining <a href="../MainFiles/biosignalsnotebooks.ipynb">Notebooks <img src="../../images/icons/link.png" width="10px" height="10px" style="display:inline"></a></span></strong> !
<hr>
<table width="100%">
<tr>
<td style="border-right:solid 3px #009EE3" width="20%">
<img src="../../images/ost_logo.png">
</td>
<td width="40%" style="text-align:left">
<a href="../MainFiles/aux_files/biosignalsnotebooks_presentation.pdf" target="_blank">☌ Project Presentation</a>
<br>
<a href="https://github.com/biosignalsnotebooks/biosignalsnotebooks" target="_blank">☌ GitHub Repository</a>
<br>
<a href="https://pypi.org/project/biosignalsnotebooks/" target="_blank">☌ How to install biosignalsnotebooks Python package ?</a>
<br>
<a href="../MainFiles/signal_samples.ipynb">☌ Signal Library</a>
</td>
<td width="40%" style="text-align:left">
<a href="../MainFiles/biosignalsnotebooks.ipynb">☌ Notebook Categories</a>
<br>
<a href="../MainFiles/by_diff.ipynb">☌ Notebooks by Difficulty</a>
<br>
<a href="../MainFiles/by_signal_type.ipynb">☌ Notebooks by Signal Type</a>
<br>
<a href="../MainFiles/by_tag.ipynb">☌ Notebooks by Tag</a>
</td>
</tr>
</table>
<span class="color6">**Auxiliary Code Segment (should not be replicated by
the user)**</span>
```
from biosignalsnotebooks.__notebook_support__ import css_style_apply
css_style_apply()
%%html
<script>
// AUTORUN ALL CELLS ON NOTEBOOK-LOAD!
require(
['base/js/namespace', 'jquery'],
function(jupyter, $) {
$(jupyter.events).on("kernel_ready.Kernel", function () {
console.log("Auto-running all cells-below...");
jupyter.actions.call('jupyter-notebook:run-all-cells-below');
jupyter.actions.call('jupyter-notebook:save-notebook');
});
}
);
</script>
```
| github_jupyter |
# Linear Elasticity in 3D
## Introduction
This example provides a demonstration of using PyMKS to compute the linear strain field for a two-phase composite material in 3D, and presents a comparison of the computational efficiency of MKS, when compared with the finite element method. The example first provides information on the boundary conditions, used in MKS. Next, delta microstructures are used to calibrate the first-order influence coefficients. The influence coefficients are then used to compute the strain field for a random microstructure. Lastly, the calibrated influence coefficients are scaled up and are used to compute the strain field for a larger microstructure and compared with results computed using finite element analysis.
### Elastostatics Equations and Boundary Conditions
A review of the governing field equations for elastostatics can be found in the [Linear Elasticity in 2D](./elasticity.ipynb) example. The same equations are used in the example with the exception that the second lame parameter (shear modulus) $\mu$ is defined differently in 3D.
$$ \mu = \frac{E}{2(1+\nu)} $$
In general, generating the calibration data for the MKS requires boundary conditions that are both periodic and displaced, which are quite unusual boundary conditions. The ideal boundary conditions are given by:
$$ u(L, y, z) = u(0, y, z) + L\bar{\varepsilon}_{xx} $$
$$ u(0, L, L) = u(0, 0, L) = u(0, L, 0) = u(0, 0, 0) = 0 $$
$$ u(x, 0, z) = u(x, L, z) $$
$$ u(x, y, 0) = u(x, y, L) $$
```
import numpy as np
from sklearn.pipeline import Pipeline
import dask.array as da
from pymks import (
generate_delta,
solve_fe,
plot_microstructures,
PrimitiveTransformer,
LocalizationRegressor,
coeff_to_real
)
#PYTEST_VALIDATE_IGNORE_OUTPUT
%matplotlib inline
%load_ext autoreload
%autoreload 2
```
## Modeling with MKS
### Calibration Data and Delta Microstructures
The first-order MKS influence coefficients are all that is needed to compute a strain field of a random microstructure, as long as the ratio between the elastic moduli (also known as the contrast) is less than 1.5. If this condition is met, we can expect a mean absolute error of 2% or less, when comparing the MKS results with those computed using finite element methods [[1]](#References).
Because we are using distinct phases and the contrast is low enough to only need the first order coefficients, delta microstructures and their strain fields are all that we need to calibrate the first-order influence coefficients [[2]](#References).
The `generate_delta` function can be used to create the two delta microstructures needed to calibrate the first-order influence coefficients for a two phase microstructure. This function uses the Python module [SfePy](http://sfepy.org/doc-devel/index.html) to compute the strain fields using finite element methods.
```
x_delta = generate_delta(n_phases=2, shape=(9, 9, 9)).persist()
plot_microstructures(
x_delta[0, x_delta.shape[1] // 2],
x_delta[1, x_delta.shape[1] // 2],
titles=['[0]', '[1]'],
cmap='gray'
)
```
Using delta microstructures for the calibration of the first-order influence coefficients is essentially the same as using a unit [impulse response](http://en.wikipedia.org/wiki/Impulse_response) to find the kernel of a system in signal processing. Delta microstructures are composed of only two phases. One phase is located only at the center cell of the microstructure, and the rest made up of the other phase.
### Generating Calibration Data
This example models a two-phase microstructure with elastic moduli values of 80 and 120 and Poisson's ratio values of 0.3 and 0.3, respectively. The macroscopic imposed strain is set to 0.02. All of these parameters used in the simulation are used in the `solve_fe` function to calculate the elastic strain.
```
strain_xx = lambda x: solve_fe(
x,
elastic_modulus=(80, 120),
poissons_ratio=(0.3, 0.3),
macro_strain=0.02
)['strain'][...,0]
y_delta = strain_xx(x_delta).persist()
```
Observe the strain field.
```
plot_microstructures(
y_delta[0, x_delta.shape[1] // 2, :, :],
titles=[r'$\mathbf{\varepsilon_{xx}}$']
)
```
### Calibrating First Order Influence Coefficients
Calibrate the influence coefficients by creating a model pipeline using the `PrimitiveTransformer` and the `LocalizationRegressor`.
```
model = Pipeline(steps=[
('discretize', PrimitiveTransformer(n_state=2, min_=0.0, max_=1.0)),
('regressor', LocalizationRegressor())
])
model.fit(x_delta, y_delta);
```
Observe the influence coefficients.
```
to_real = lambda x: coeff_to_real(x.steps[1][1].coeff).real
coeff = to_real(model)
plot_microstructures(
coeff[x_delta.shape[1] // 2, :, :, 0],
coeff[x_delta.shape[1] // 2, :, :, 1],
titles=['Influence coeff [0]', 'Influence coeff [1]']
)
```
The influence coefficients have a Gaussian-like shape.
### Predict of the Strain Field for a Random Microstructure
Use the calibrated `model` to compute the strain field for a random two phase microstructure and compare it with the results from a finite element simulation. The `strain_xx` helper function is used to generate the strain field.
```
# NBVAL_IGNORE_OUTPUT
da.random.seed(99)
x_data = da.random.randint(2, size=(1,) + x_delta.shape[1:]).persist()
%time y_data = strain_xx(x_data).persist()
plot_microstructures(
x_data[0, x_delta.shape[1] // 2, :, :],
titles=['Microstructure']
)
plot_microstructures(
y_data[0, x_delta.shape[1] // 2, :, :],
titles=[r'$\mathbf{\varepsilon_{xx}}$']
)
```
**Note that the calibrated influence coefficients can only be used to reproduce the simulation with the same boundary conditions that they were calibrated with.**
Now to get the strain field from the model, pass the same microstructure to the `predict` method.
```
# NBVAL_IGNORE_OUTPUT
%time y_predict = model.predict(x_data).persist()
```
Finally, compare the results from finite element simulation and the MKS model.
```
plot_microstructures(
y_data[0, x_delta.shape[1] // 2, :, :],
y_predict[0, x_delta.shape[1] // 2, :, :],
titles=[
r'$\mathbf{\varepsilon_{xx}}$ - FE',
r'$\mathbf{\varepsilon_{xx}}$ - MKS'
]
)
```
Observe the difference between the two plots.
```
plot_microstructures(
(y_data - y_predict)[0, x_delta.shape[1] // 2, :, :],
titles=['FE - MKS']
)
```
The MKS model is able to capture the strain field for the random microstructure after being calibrated with delta microstructures.
## Resizing the Coefficeints to use on Larger Microstructures
The influence coefficients that were calibrated on a smaller microstructure can be used to predict the strain field on a larger microstructure though spectral interpolation [[3]](#References), but accuracy of the MKS model drops slightly. To demonstrate how this is done, let's generate a new larger $m$ by $m$ random microstructure and its strain field.
```
new_shape = tuple(np.array(x_delta.shape[1:]) * 3)
x_large = da.random.randint(2, size=(1,) + new_shape).persist()
```
The influence coefficients that have already been calibrated need to be resized to match the shape of the new larger microstructure that we want to compute the strain field for. This can be done by passing the shape of the new larger microstructure into the `coeff_resize` method.
```
model.steps[1][1].coeff_resize(x_large[0].shape);
```
Use the resize coefficients to calculate large strain field. The coefficients can not now be used for the smaller microstructures.
```
# NBVAL_IGNORE_OUTPUT
%time y_large = model.predict(x_large).persist()
plot_microstructures(y_large[0, x_delta.shape[1] // 2], titles=[r'$\mathbf{\varepsilon_{xx}}$'])
```
## References
<a id="ref1"></a>
[1] Binci M., Fullwood D., Kalidindi S.R., *A new spectral framework for establishing localization relationships for elastic behav ior of composites and their calibration to finite-element models*. Acta Materialia, 2008. 56 (10): p. 2272-2282 [doi:10.1016/j.actamat.2008.01.017](http://dx.doi.org/10.1016/j.actamat.2008.01.017).
<a id="ref2"></a>
[2] Landi, G., S.R. Niezgoda, S.R. Kalidindi, *Multi-scale modeling of elastic response of three-dimensional voxel-based microstructure datasets using novel DFT-based knowledge systems*. Acta Materialia, 2009. 58 (7): p. 2716-2725 [doi:10.1016/j.actamat.2010.01.007](http://dx.doi.org/10.1016/j.actamat.2010.01.007).
<a id="ref3"></a>
[3] Marko, K., Kalidindi S.R., Fullwood D., *Computationally efficient database and spectral interpolation for fully plastic Taylor-type crystal plasticity calculations of face-centered cubic polycrystals*. International Journal of Plasticity 24 (2008) 1264–1276 [doi;10.1016/j.ijplas.2007.12.002](http://dx.doi.org/10.1016/j.ijplas.2007.12.002).
| github_jupyter |
# Bond Prices and Yields
Debt securities are often called **fixed-income securities**, because they promise either a fixed stream of income or one determined according to a specified formula.
## Bond Characteristics
**bond**: A security that obligates the issuer to make specified pay-ments to the holder over a period of time.
bond’s **par value, face value**: The payment to the bondholder at the maturity of the bond.
**coupon rate**: A bond’s annual interest payment per dollar of par value.
**zero-coupon bonds**: A bond paying no coupons that sells at a discount and provides only a payment of par value at maturity.
>**e.g.**
>
>- Par value of $\$1,000$
- Coupon rate of $8\%$
- The initial price is $\$997$
- Maturity, $5$ years
- Semi-annual coupon payments
>
>Cash flow of the buyer
>
>
### Treasury Bonds and Notes
$$
\begin{array}{c} \hline
\begin{array}{c:c}
\text{T-Notes} & \text{T-Bonds} \\ \hline
1 \text{ to } 10 \text{ years} & 10 \text{ to } 30 \text{ years} \\ \hline
\end{array} \\
\text{denominations of \$}100 \text{ or \$}1000 \\
\text{semiannual coupon payments} \\ \hline
\end{array}
$$
**Notice**: The number in table with *column* represents the anually coupon, which paid once of six months.
There's also a minimum price increment, or **tick size**.
>**e.g.**
>
>Suppose now the tick size is $\newcommand{\ffrac}{\displaystyle \frac} \ffrac{1} {128}$. And the ask price on the board is $132.9922\%$ of $1000$ dollars.
>
>So when it comes to $\ffrac{126} {128}$, it's $0.984375 < 0.9922$, so there is still space for it to goes up until $\ffrac{127} {128} = 0.9921875 \approx 0.9922$, same value after round to $4$ decimal digits
The other term, **ask yield**, is the yield to maturity (YTM) based on the ask price, which equals to $\text{Ask Price} + \text{Total Coupon}$
### Accured Interest and Quoted Bond Priceds
$\odot$The bond prices that you see quoted in the financial pages are NOT actually the prices that investors pay for the bond. This is because the quoted price does not include the interest that accrues between coupon payment dates.$\Join$
If a bond is purchased between coupon payments, the buyer must pay the seller for accrued interest.
$$\text{Accrued interest} = \frac{\text{Annual coupon payment}} {2} \times \frac{\text{Days since last coupon payment}} {\text{Days separting coupon payments}}$$
The actual payment for the buyer is called **invoice price** which is the sum of **quoted price** and **Accured interest**.
>**e.g.** Suppose that the coupon rate is $8\%$. You buy after $30$ days after last coupon payment, and there are $182$ days in the semiannual coupon period.
>
>The semiannual coupon payment is $40$ dollars. And the seller is entitled to a payment of accrued interest of $\ffrac{30} {182}$ of the semiannual coupon, which is $40 \times \ffrac{30} {182} = 6.59$. If the quoted price of the bond is $\$990$, then the invoice price will be $\$990 + \$6.59 = \$996.59$.
### Corporate Bonds
**Callable bonds**: Bonds that may be repurchased by the issuer at a specified call price during the call period.
- higher coupon rate than noncallable bonds
- higher promised yields to maturity than noncallable bonds
- when the coupon bond are higher than the current interest rate, the firm may call it back
**Convertible bonds**: the buyer has an option to convert bonds into stocks.
- lower coupon rate than nonconvertible bonds
- lower promised yields to maturity than noncallable bonds
**Puttable bonds**: the buyer has an option to retire the bond earlier.
- Retire the bond when the market interest rates are higher than before
**Floating-rate bonds**: coupon rates periodically reset according to some
market rates.
- For example, $\text{next year coupon rate (annully adjusted)} = \text{T-bill rate (at adjustment date)} + 2\%$
### Preferred Stock
- like bonds, preferred stock promises to pay a specified stream of dividends (normally a fixed amount)
- unlike bonds, the failure to pay the promised dividend does not result in corporate bankruptcy
- the claim to the firm’s assets has lower priority than that of bondholders but higher priority than that of common stockholders.
- unlike bonds, payment on divident is not tax-deductible expenses to the firm
- an offsetting tax advantage: When one corporation buys the preferred stock of another corporation, it pays taxes on only $30\%$ of the dividends received. For example: if the firm’s tax bracket is $35\%$, then the **effective tax rate** on preferred dividends is $30\% \times 35\% = 10.5\%$.
- Preferred stock rarely gives its holders full voting privileges in the firm. However, if the preferred dividend is skipped, the preferred stockholders will then be provided some voting power.
- Based on above, most preferred stock is held by corporations.
### International Bonds
Exchange rate risk!
1. Foreign bonds
- Yankee bonds: Dollar-denominated bonds sold in the U.S. by non-U.S. issuers
- Samurai bonds: Yen-denominated bonds sold in Japan by non-Japanese issuers
- Bulldog bonds: Pound-denominated bonds sold in the U.K. by non-U.K. issuers
2. Eurobonds
- Euroyen: Yen-denominated bonds selling outside Japan
- Eurosterling: Pound-denominated bonds selling outside the U.K.
## Bond Pricing
Calculate the present value of all future cash flow at YTM.
**YTM**: universal discount rate for cash flows of any horizons, $r$.
$$\begin{align}
\text{Bond value} &= \text{Present value of coupons} + \text{Present value of par value} \\
&= \sum_{t=1} ^{T} \frac{\text{Coupon}} {(1+r)^t} + \frac{\text{Par}} {(1+r)^T}
\end{align}$$
To better calculate, we define the following
$$\text{Annuity factor}(r,T) = \frac{1} {r} \left[ 1 - \frac{1} {(1+r)^T}\right], \text{PV factor}(r,T) = \frac{1} {(1+r)^T}$$
So that $\text{Bond value} = \text{Coupon} \times \text{Annuity factor}(r,T) + \text{Par Value} \times \text{PV factor}(r,T)$
**The inverse relationship between price and yield**: Bond price will fall as market interest rates rise, which is the central feature of fixed-income securities.
$\odot$Generally, keeping all other factors the same, the longer the maturity of the bond, the greater the sensitivity of its price to fluctuations in the interest rate. $\Join$
>**e.g.** Given $\text{Par} = 100$, $\text{Annual coupon rate} = 10\%$, $\text{YTM} = r = 10\%$, $\text{Maturity} = T = 2$;
>
>If you bought at 1/1/2000, it will pay first coupon at 1/1/2001 and mature at 1/1/2002, so that its price now
>
>$$\sum_{t=1} ^{T} \frac{\text{Coupon}} {(1+r)^t} + \frac{\text{Par}} {(1+r)^T} = \frac{\$10} {(1+0.1)^1} + \frac{\$10 + \$100} {(1+0.1)^2} = \$100$$
>
>And if you bought at 4/1/2000, first we need to calculate the $\text{Accrued Interest}$
>
>$$\frac{1/1/2000 - 4/1/2000} {1/1/2000 - 1/1/ 2001} =\frac{91} {366} = 0.2486$$
>
>Then is the discount, remember to discount to the issue date, which is
>
>$$\sum_{t=1}^{T} \frac{\text{Coupon}} {(1+r)^{t-0.2486}} + \frac{\text{Par}} {(1+r)^{T-0.2486}}= \frac{10} {1.01^{0.751}} + \frac{110} {1.01^{1.751}} = 102.4$$
>
>Or another way:
>
>$$\text{Invoice Price} = \text{Quoted Price} + \text{Accrued Interest} = 99.912 + 2.486 = 102.4$$
## Bond Yields
### Yield to Maturity
**yield to maturity (YTM)**: The discount rate that makes the present value of a bond’s payments equal to its price.
Let $n$ be the number of payments, we solve $r$ using the equation below
$$\text{Bond-Price Today} = \sum_{t=1}^{n}\frac{\text{Coupon}} {\left( 1+r \right)^t} + \frac{\text{Par}} {\left( 1+r \right)^n} $$
So that $\text{YTM} = n * r$, and the effective annual rate is $\left( 1+r \right)^n$
Similar if bond price is during coupon dates.
**current yield**: Annual coupon divided by bond price.
>**e.g.**
>
>- $\text{Par} = \$1000$
- Semiannual coupon payments
- $\text{Annual coupon rate} = 8\%$
- $\text{Maturity} = T = 30$ years
- $\text{Bond price} = \text{PV} = \$1,276.76$
>
>So that $\text{Coupon Rate} = 8\%$
>
>$\text{current yield} = \ffrac {\text{Annual coupon payments}} {\text{Bond price}} = \ffrac{\$ 80} {\$ 1276.76} = 6.27\%$
>
>Using calculator, $\text{YTM} = r = 0.06 $
Normally
- YTM = Coupon Rate →→→ Price = Par (sell at par)
- **Discount Bond**: YTM > Coupon Rate →→→ Price < Par (sell at a discount)
- Coupon Rate < Current Yield < YTM
- **Premium Bond**: YTM < Coupon Rate →→→ Price > Par (sell at a premium)
- Coupon Rate > Current Yield > YTM
And one thing to remember is that both YTM and Coupon Rate are annulized into APR's (Annual Percentage Rate).
### Yield to Call
Similar to Yield to Maturity, now it's mature ahead of the term in a higher call price
>**e.g.**
>
>- $\text{Par} = \$1000$
- Semiannual coupon payments
- $\text{Annual coupon rate} = 8\%$
- $\text{Maturity} = T = 30$ years
- $\text{Bond price} = \text{PV} = \$1150$
- Callable in $10$ years at a call price of $1100$
>
>YTM:```=YIELD(DATE(2000,1,1),DATE(2030,1,1),8%,115,100,2)``` = $6.82\%$
>
>YTC:```=YIELD(DATE(2000,1,1),DATE(2010,1,1),8%,115,110,2)``` = $6.61\%$
## Bond Prices over Time
Even if the interest rate (YTM) is constant over the life of the bond, the bond price still varies over time unless it’s sold at the par (CR=YTM).

## Default Risk and Bond Pricing
**investment grade bond**: A bond rated BBB and above by Standard & Poor's or Baa and above by Moody's.
**speculative grade or junk bonds**: A bond rated BB or lower by Standard & Poor's, Ba or lower by Moody's, or unrated.
Contrary to T-bonds, Corporate bonds have default risk, which affects the bond rating.
### Yield to Maturity and Default Risk
Bondholders are expected to receive only $70\%$ of par when the firm goes bankrupt. And at that time, the expected YTM will be lower than the previously stated YTM.
Default Premium = Yield spreads between corporate and comparable T-bonds
### Credit Default Swaps
CDS is an insurance policy on the default risk of a corporate bond or loan.
In the event of default, CDS buyer may deliver a defaulted bond to the seller in return for the bond's par value.
## The Yield Curve
**Yield curve**: a graph of YTM as a function of term to maturity.
T**erm structure of interest rates**: the relationship between YTM and term to maturity.
Types:
1. Flat Yield Curve
2. Upward-sloping (Rising Yield Curve) (the most common one)
3. Downward-sloping (Inverted Yield Curve)
4. Hump shaped
### The Expectations Theory
**Expectation hypothesis**: The theory that YTM are determined by expectations of future short-term interest rates.
>**e.g.** Returns to two two-year investment strategies
>
>$r_1 = 8\%$, $E(r_2) = 10\%$. what is the fair current YTM for two-year bond?
>
>$$y_2 = \frac{\sqrt{1.08 \times 1.10} - 1} {2} = 8.995\%$$
>
>So that on the yield curve, $y_1 = 8\%$, $y_2 = 8.995\%$
>
>- Upward-sloping yield curve $\Rightarrow$ expect future interest rate $\uparrow$
- More loan and more money in stock market
- Downward-sloping yield curve $\Rightarrow$ expect future interest rate $\downarrow$
**forward rate**: The inferred short-term rate of interest for a future period that makes the expected total return of a long-term bond equal to that of rolling over short-term bonds.
Using the expectations hypothesis to infer the market’s expectation of future short-term rates inversely.
>**e.g.**
>
>Given $y_1=8\%$, $y_2=8.995\%$, since $\left( 1+y_1 \right)\cdot\left( 1+f_2 \right) = \left( 1+y_2 \right)^2$, we have
>
>$$f_2 = \frac{\left( 1+y_2 \right)^2} {\left( 1+y_1 \right)} - 1 = 10\%$$
>
>Given YTM for two-year bond: $6\%$; YTM for three-year bond: $7\%$
>
>What is the forward rate for the third year?
>
>$$f_3 = \frac{\left( 1+y_3 \right)^3} {\left( 1+y_2 \right)^2} - 1 = 9.03\%$$
$$f_n = \frac{\left( 1+y_n \right)^n} {\left( 1+y_{n-1} \right)^{n-1}} - 1$$
### The Liquidity Preference Theory
Long-term bonds are subject to greater interest rate risk than short-term bonds. Therefore investors in long-term bonds might require a risk premium (liquidity premium) to compensate them for this risk.
**liquidity preference theory**: The theory that investors demand a risk premium on long-term bonds.
**liquidity premium**: The extra expected return demanded by investors as compensation for the greater risk of longer-term bonds.
$$\text{Forward Rate}: f_n = E(r_n) + \text{liquidity premium}$$
>**e.g.**
>
>- Without $1\%$ liquidity premium, $y_1 = 8\%$, $f_2 = 8\%$.
- $y_2 = 8\%$, flat yield curve
>
>
>- With $1\%$ liquidity premium, $y_1 = 8\%$, $f_2 = 8\% + 1\% = 9\%$.
- $y_2 = \sqrt{1.08\times 1.09} - 1 = 8.5\% > y_1 = 8\%$, upward-sloping yield curve
So that **in the *presence* of liquidity premium**, even **in the *absence* of any expectation of future increases in interest rates**, still the yield curve will be upward-sloping.
### A Synthesis
A trade-off between Liquidity Preference and Expectations Theories

And for most of time long term t-notes have higher return rate than short term t-bills, meaning that it's commonly upward-sloping.
## Summary
- Debt securities are distinguished by their promise to pay a fixed or specified stream of income to their holders. The coupon bond is a typical debt security.
- Treasury notes and bonds have original maturities greater than one year. They are issued at or near par value, with their prices quoted net of accrued interest.
- Callable bonds should offer higher promised yields to maturity to compensate investors for the fact that they will not realize full capital gains should the interest rate fall and the bonds be called away from them at the stipulated call price. Bonds often are issued with a period of call protection. In addition, discount bonds selling significantly below their call price offer implicit call protection.
- Put bonds give the bondholder rather than the issuer the choice to terminate or extend the life of the bond.
- Convertible bonds may be exchanged, at the bondholder’s discretion, for a specified number of shares of stock. Convertible bondholders “pay” for this option by accepting a lower coupon rate on the security.
- Floating-rate bonds pay a fixed premium over a referenced short-term interest rate. Risk is limited because the rate paid is tied to current market conditions.
- The yield to maturity is the single discount rate that equates the present value of a security’s cash flows to its price. Bond prices and yields are inversely related. For premium bonds, the coupon rate is greater than the current yield, which is greater than the yield to maturity. These inequalities are reversed for discount bonds.
- The yield to maturity often is interpreted as an estimate of the average rate of return to an investor who purchases a bond and holds it until maturity. This interpretation is subject to error, however. Related measures are yield to call, realized compound yield, and expected (versus promised) yield to maturity.
- Treasury bills are U.S. government–issued zero-coupon bonds with original maturities of up to one year. Treasury STRIPS are longer-term default-free zero-coupon bonds. Prices of zero-coupon bonds rise exponentially over time, providing a rate of appreciation equal to the interest rate. The IRS treats this price appreciation as imputed taxable interest income to the investor.
- When bonds are subject to potential default, the stated yield to maturity is the maximum possible yield to maturity that can be realized by the bondholder. In the event of default, however, that promised yield will not be realized. To compensate bond investors for default risk, bonds must offer default premiums, that is, promised yields in excess of those offered by default-free government securities. If the firm remains healthy, its bonds will provide higher returns than government bonds. Otherwise, the returns may be lower.
- Bond safety often is measured using financial ratio analysis. Bond indentures offer safeguards to protect the claims of bondholders. Common indentures specify sinking fund requirements, collateralization, dividend restrictions, and subordination of future debt.
- Credit default swaps provide insurance against the default of a bond or loan. The swap buyer pays an annual premium to the swap seller but collects a payment equal to lost value if the loan later goes into default.
- The term structure of interest rates is the relationship between time to maturity and term to maturity. The yield curve is a graphical depiction of the term structure. The forward rate is the break-even interest rate that would equate the total return on a rollover strategy to that of a longer-term zero-coupon bond.
- The expectations hypothesis holds that forward interest rates are unbiased forecasts of future interest rates. The liquidity preference theory, however, argues that long-term bonds will carry a risk premium known as a liquidity premium. A positive liquidity premium can cause the yield curve to slope upward even if no increase in short rates is anticipated.
## Key Terms
- bond
- callable bonds
- collateral
- convertible bonds
- coupon rate
- credit default swap (CDS)
- current yield
- debenture
- default premium
- discount bonds
- expectations hypothesis
- face value
- floating-rate bonds
- forward rate
- horizon analysis
- indenture
- investment grade bonds
- liquidity preference theory
- liquidity premium
- par value
- premium bonds
- put bond
- realized compound return
- reinvestment rate risk
- sinking fund
- speculative grade or junk bond
- subordination clauses
- term structure of interest rates
- yield curve
- yield to maturity (YTM)
- zero-coupon bond
## Key Formula
- Price of a coupon bond
$$\begin{align}
\text{Bond value} &= \text{Present value of coupons} + \text{Present value of par value} \\
&= \sum_{t=1} ^{T} \frac{\text{Coupon}} {(1+r)^t} + \frac{\text{Par}} {(1+r)^T} \\
&= \text{Coupon} \times \text{Annuity factor}(r,T) + \text{Par Value} \times \text{PV factor}(r,T)
\end{align}$$
- Forward rate of interest
$$1 + f_n = \frac{\left( 1+y_n \right)^n} {\left( 1+y_{n-1} \right)^{n-1}}$$
- Liquidity premium = Forward rate – Expected short rate
## Assignment
1. Sinking funds are commonly viewed as protecting the ___ of the bond.
- ~~Issure~~
- ~~Underwriter~~
- Holder
- ~~Dealer~~
2. A mortgage bond is
- secured by property owned by the firm
- ~~secured by equipment owned by the firm~~
- ~~unsecured~~
- ~~secured by other securities held by the firm~~
3. Floating-rate bonds have a ___ that is adjusted with current market interest rates.
- ~~maturity date~~
- ~~coupon payment date~~
- coupon rate
- ~~dividend yield~~
4. The primary difference between Treasury notes and bonds is
- maturity at issue
- ~~default risk~~
- ~~coupon rate~~
- ~~tax status~~
5. TIPS offer investors inflation protection by ___ by the inflation rate each year.
- ~~increasing only the coupon rate~~
- increasing both the par value and the coupon payment
- ~~increasing only the par value~~
- ~~increasing the promised yield to maturity~~
6. ___ bonds represent a novel way of obtaining insurance from capital markets against specified disasters.
- ~~Asset-backed bonds~~
- ~~TIPS~~
- Catastrophe
- ~~Pay-in-kind~~
7. Everything else equal, the ___ the maturity of a bond and the ___ the coupon, the greater the sensitivity of the bond's price to interest rate changes.
- ~~longer; higher~~
- ~~shorter; higher~~
- longer; lower
- ~~shorter; lower~~
8. A coupon bond that pays interest of $\$60$ annually has a par value of $\$1,000$, matures in $5$ years, and is selling today at an $\$84.52$ discount from par value. The yield to maturity on this bond is
$$8.12\%$$
9. Given zero coupon bonds: A with $1$ year of maturity and YTM: $6\%$; B with $2$ year of maturity and YTM: $7.50\%$, The expected $1\text{-year}$ interest rate $1$ year from now should be about
$$1.075^2 \div 1.06 - 1 \approx 9.0212\%$$
10. **VIP** A $1\%$ decline in yield will have the least effect on the price of a bond with a
- ~~20-year maturity, selling at 80~~
- ~~20-year maturity, selling at 100~~
- 10-year maturity, selling at 100
- ~~10-year maturity, selling at 80~~
11. An investor pays $\$989.40$ for a bond. The bond has an annual coupon rate of $4.8\%$. What is the current yield on this bond?
$$\$48 \div \$989.40 = 4.8514\%$$
12. A bond was purchased at a premium and is now selling at a discount because of a change in market interest rates. If the bond pays a $4\%$ annual coupon, what is the likely impact on the holding-period return if an investor decides to sell now?
- ~~Increased~~
- Decreased
- ~~Stayed the same~~
- ~~The answer cannot be determined from the information given~~
13. You buy a TIPS at issue at par for $\$1,000$. The bond has a $4.0\%$ coupon. Inflation is $3.0\%$, $4.0\%$, and $5.0\%$ over the next $3$ years. The total annual coupon income you will receive in year $3$ is
$$\$44 \times 1.033 \times 1.04 \times 1.05 = \$44.9904$$
14. A coupon bond that pays interest of $\$61$ annually has a par value of $\$1,000$, matures in $5$ years, and is selling today at a $\$75.50$ discount from par value. The current yield on this bond is
$$\$61 \div (\$1000 - \$75.5) = 6.60\%$$
| github_jupyter |
参考:
1. [地址一](https://blog.csdn.net/lh0616/article/details/100067697?depth_1-utm_source=distribute.pc_relevant.none-task&utm_source=distribute.pc_relevant.none-task)
2. [地址二](https://www.cnblogs.com/xleng/p/10948838.html)
3. Gstreamer应用开发手册
**gstreamer 官方网址**
[https://gstreamer.freedesktop.org](https://gstreamer.freedesktop.org/documentation)
# Gstreamer 是什么
GStreamer 是一个创建流媒体应用程序的框架。其基本设计思想来自于俄勒冈(Oregon)研究生学院有关视频管道的创意, 同时也借鉴了 DirectShow 的设计思想。
GStreamer 的程序开发框架使得编写任意类型的流媒体应用程序成为了可能。在编写处理音频、视频或者两者皆有的应用程序时, GStreamer 可以让你的工作变得简单。GStreamer 并不受限于音频和视频处理, 它能够处理任意类型的数据流。管道设计的方法对于实际应用的滤波器几乎没有负荷, 它甚至可
以用来设计出对延时有很高要求的高端音频应用程序。
GStreamer 最显著的用途是在构建一个播放器上。GStreamer 已经支持很多格式的文件了, 包括:MP3、Ogg/Vorbis、MPEG-1/2、AVI、Quicktime、 mod 等等。从这个角度看,GStreamer 更象是一个播放器。但是它主要的优点却是在于: 它的可插入组件能够很方便的接入到任意的管道当中。这个优点使得利用 GStreamer 编写一个万能的可编辑音视频应用程序成为可能。
**GStreamer 框架是基于插件的**, 有些插件中提供了各种各样的多媒体数字信号编解码器,也有些提供了其他的功能。所有的插件都能够被链接到任意的已经定义了的数据流管道中。GStreamer 的管道能够被 GUI 编辑器编辑, 能够以 XML 文件来保存。这样的设计使得管道程序库的消耗变得非常少。
GStreamer 核心库函数是一个处理插件、数据流和媒体操作的框架。GStreamer 核心库还提供了一个API, 这个API是开放给程序员使用的---当程序员需要使用其他的插件来编写他所需要的应用程序的
时候可以使用它。
下图是对基于Gstreamer框架的应用的简单分层:
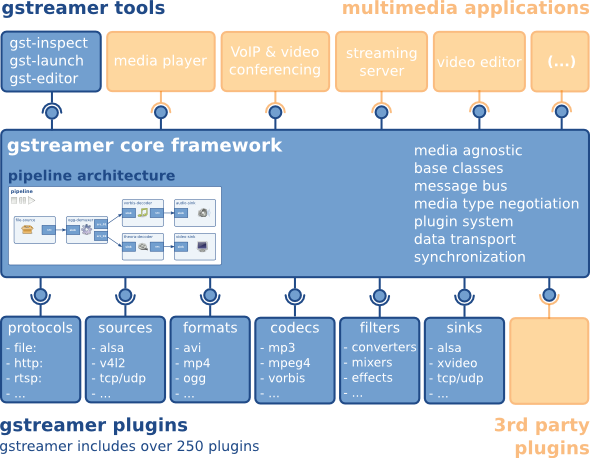
## Media Applications
最上面一层为应用,比如gstreamer自带的一些工具(gst-launch,gst-inspect等),以及基于gstreamer封装的库(gst-player,gst-rtsp-server,gst-editing-services等)根据不同场景实现的应用。
## Core Framework
中间一层为Core Framework,主要提供:
- 上层应用所需接口
- Plugin的框架
- Pipline的框架
- 数据在各个Element间的传输及处理机制
- 多个媒体流(Streaming)间的同步(比如音视频同步)
- 其他各种所需的工具库
## Plugins
最下层为各种插件,实现具体的数据处理及音视频输出,应用不需要关注插件的细节,会由Core Framework层负责插件的加载及管理。主要分类为:
- Protocols:负责各种协议的处理,file,http,rtsp等。
- Sources:负责数据源的处理,alsa,v4l2,tcp/udp等。
- Formats:负责媒体容器的处理,avi,mp4,ogg等。
- Codecs:负责媒体的编解码,mp3,vorbis等。
- Filters:负责媒体流的处理,converters,mixers,effects等。
- Sinks:负责媒体流输出到指定设备或目的地,alsa,xvideo,tcp/udp等。
Gstreamer框架根据各个模块的成熟度以及所使用的开源协议,将core及plugins置于不同的源码包中:
- gstreamer: 包含core framework及core elements。
- gst-plugins-base: gstreamer应用所需的必要插件。
- gst-plugins-good: 高质量的采用LGPL授权的插件。
- gst-plugins-ugly: 高质量,但使用了GPL等其他授权方式的库的插件,比如使用GPL的x264,x265。
- gst-plugins-bad: 质量有待提高的插件,成熟后可以移到good插件列表中。
- gst-libav: 对libav封装,使其能在gstreamer框架中使用。
# Gstreamer 基本概念
## 元件(element)
元件 (element) 是 GStreamer 中最重要的概念。你可以通过创建一系列的元件 (Elements), 并把它们连接起来, 从而让数据流在这个被连接的各个元件(Elements) 之间传输。每个元件 (Elements) 都有一个特殊的函数接口, 对于有些元件 (Elements) 的函数接口它们是用于能够读取文件的数据, 解码文件数据的。而有些元件 (Elements) 的函数接口只是输出相应的数据到具体的设备上 (例如, 声卡设备)。你可以将若干个元件(Elements) 连接在一起, 从而创建一个管道 (`pipeline`) 来完成一个特殊的任务, 例如, 媒体播放或者录音。GStreamer 已经默认安装了很多有用的元件 (Elements), 通过使用这些元件(Elements) 你能够构建一个具有多种功能的应用程序。当然, 如果你需要的话, 你可以自己编写一个新的元件 (Elements)。对于如何编写元件(Elements) 的话题在**GStreamer Plugin Writer's Guide** 中有详细的说明
Elements 根据功能可以分为以下三种:
* Source elements(源元件)
* Sink elements (接收元件)
* Filter elements (过滤元件)
### 源元件 (Source elements)
为管道产生数据,比如从磁盘或者声卡读取数据,源元件不接收数据,仅产生数据。仅有一个源衬垫,形象化的源元件如下图所示

### 过滤元件(Filter elements)
包括过滤器 (filters)、转换器(convertors)、分流器(demuxers)、整流器(muxers) 以及编解码器 (codecs) 等等,同时拥有输入和输出衬垫,形象化过滤元件如下图所示
### 接收元件
是媒体管道的末端,它接收数据但不产生任何数据。仅有一个接收衬垫,形象化接收元件如下图所示

### 元件状态
包括以下几种:
- GST_STATE_NULL: 默认状态
- GST_STATE_READY: 准备状态
- GST_STATE_PAUSED: 暂停状态
- GST_STATE_PLAYING: 运行状态
NULL和READY状态下,element不对数据做任何处理,PLAYING状态对数据进行处理,PAUSE状态介于两者之间,对数据进行preroll。应用程序通过函数调用控制pipeline在不同状态之间进行转换。
element的状态变换不能跳过中间状态,比如不能从READY状态直接变换到PLAYING状态,必须经过中间的PAUSE状态。
**element的状态转换成PAUSE会激活element的pad。首先是source pad被激活,然后是sink pad**。pad被激活后会调用activate函数,有一些pad会启动一个Task。
**PAUSE状态下,pipeline会进行数据的preroll,目的是为后续的PLAYING状态准备好数据,使得PLAYING启动的速度更快。一些element需接收到足够的数据才能完成向PAUSE状态的转变,sink pad只有在接收到第一个数据才能实现向PAUSE的状态转变。**
通常情况下,element的状态转变需要协调一致。
## 衬垫 (Pads)
Pad是一个element的输入/输出接口,分为src pad(生产数据)和sink pad(消费数据)两种。
衬垫 (Pads) 是元件对外的接口。一个衬垫(Pads)可以被看作是一个元件(element)插座或者端口,元件(element)之间的链接就是依靠着衬垫(Pads),**数据流从一个元件的源衬垫 (source pad) 到另一个元件的接收衬垫 (sink pad)。**
衬垫(Pads)有处理特殊数据的能力:一个衬垫(Pads)能够限制数据流类型的通过。链接成功的条件是:**只有在两个衬垫(Pads)允许通过的数据类型一致的时候才被建立**。两个element必须通过pad才能连接起来,pad拥有当前element能处理数据类型的能力(capabilities),会在连接时通过比较src pad和sink pad中所支持的能力,来选择最恰当的数据类型用于传输,如果element不支持,程序会直接退出。在element通过pad连接成功后,数据会从上一个element的src pad传到下一个element的sink pad然后进行处理。
下面的这个比喻可能对你理解衬垫(Pads)有所帮助。一个衬垫(Pads)很象一个物理设备上的插头。例如一个家庭影院系统。一个家庭影院系统由一个功放(amplifier),一个 DVD 机,还有一个无声的视频投影组成。 我们需要连接 DVD 机到功放(amplifier),因为两个设备都有音频插口;我们还需要连接投影机到 DVD 机上,因为 两个设备都有视频处理插口。但我们很难将投影机与功放(amplifier)连接起来,因为他们之间处理的是不同的 插口。GStreamer 衬垫(Pads)的作用跟家庭影院系统中的插口是一样的。
当element支持多种数据处理能力时,我们可以通过Cap来指定数据类型.
例如,下面的命令通过Cap指定了视频的宽高,videotestsrc会根据指定的宽高产生相应数据:
```sh
gst-launch-1.0 videotestsrc ! "video/x-raw,width=1280,height=720" ! autovideosink
```
对于大部分情况,所有的数据流都是在链接好的元素之间流动。数据向元件(element)以外流出可以通过一个或者多个 source 衬垫(Pads),元件(element)接受数据是通过一个或者多个 sink 衬垫(Pads)来完成的。
**Source 元件(element)和 sink 元件(element)分别有且仅有一个 sink 衬垫(Pads)或者 source 衬垫(Pads)。**
Pads 的分类:
- 永久型(always)
- 随机型(sometimes)
- 请求型 (on request)
大部分情况下,所有在 GStreame r 中流经的数据都遵循一个原则:**数据从 element 的一个或多个源衬垫流出,从一个或多个sink 衬垫流入。源和 sink 元件分别只有源和 sink 衬垫。**
**数据在这里代表的是缓冲区(buffers)([GstBuffer对象描述了数据的缓冲区(buffers)的信息](https://gstreamer.freedesktop.org/documentation/gstreamer/gstbuffer.html?gi-language=c#GstBuffer)和事件(events) ([GstEvent 对象描述了数据的事件(events)信息](https://gstreamer.freedesktop.org/documentation/gstreamer/gstevent.html?gi-language=c)。)**
## ❤️箱柜(bin) 和管道(Pipeline)
箱柜 (Bins) 是一个可以装载元件 (element) 的容器。管道 (pipelines) 是箱柜 (Bins) 的一个特殊的子类型, 管道 (pipelines) 可以操作包含在它自身内部的所有元件 (element)。因为箱柜(Bins) 本身又是组件 (element) 的子集, 所以你能够象操作普通组件 (element) 一样的操作一个箱柜 (Bins), 通过这种方法可以降低你的应用程序的复杂度。
**你可以改变一个箱柜(Bins) 的状态来改变箱柜 (Bins) 内部所有组件 (element) 的状态**。箱柜 (Bins) 可以发送总线消息 (bus messages) 给它的子集组件(element)(这些消息包括: 错误消息(error messages), 标签消息(tag messages),EOS 消息(EOS messages))。如果没有bin,我们需要依次操作我们所使用的element。通过bin降低了应用的复杂度。
形象化箱柜如下图所示
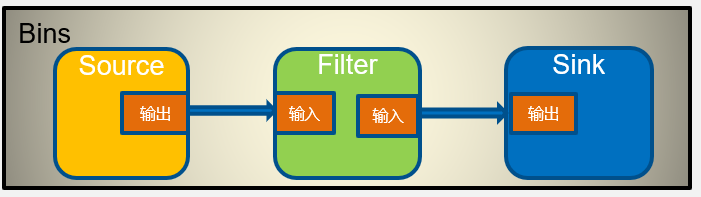
<br>
管道(pipeline)是高级的箱柜(Bins)。**当你设定管道的暂停或者播放状态的时候,数据流将开始流动,并且媒体数据处理也开始处理。一旦开始,管道将在一个单独的线程中运行,直到被停止或者数据流播放完毕。**
## 能力集(Cap)
Caps 描述了能够通过衬垫或当前通过衬垫的数据流格式。一个衬垫能够有多个功能。功能 (Caps) 可以用一个包含一个或多个 GstStructures 的数组来表示。每个 GstStructures 由一个名字字符串 (比如说 "width") 和相应的值 (类型可能为 G_TYPE_INT 或 GST_TYPE_INT_RANGE) 构成。
# Gstreamer数据消息交互
## 数据流
**Gstreamer支持两种类型的数据流,分别是push模式和pull模式**。在push模式下,upstream的element通过调用downstream的sink pads的函数实现数据的传送。在pull模式下,downstream的element通过调用upstream的source pads的函数实现对数据的请求。
**push模式是常用的模式,pull模式一般用于demuxer或者低延迟的音频应用等。**
## 通信对象类型
在pipeline运行的过程中,各个element以及应用之间不可避免的需要进行数据消息的传输,gstreamer提供了bus系统以及多种数据类型(Buffers、Events、Messages,Queries)来达到此目的:
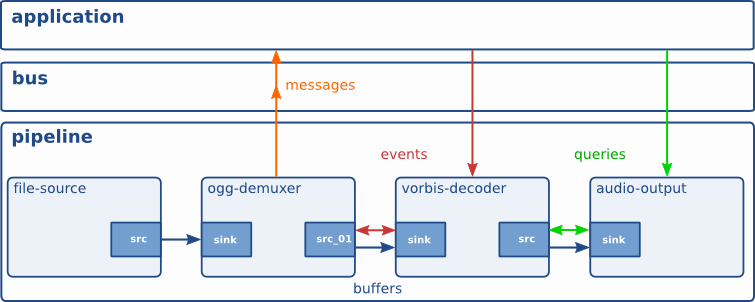
### Bus
Bus是gstreamer内部用于将消息从内部不同的streaming线程,传递到bus线程,再由bus所在线程将消息发送到应用程序。应用程序只需要向bus注册消息处理函数,即可接收到pipline中各element所发出的消息,使用bus后,应用程序就不用关心消息是从哪一个线程发出的,避免了处理多个线程同时发出消息的复杂性。
### Buffers
用于从sources到sinks的媒体数据传输。
缓冲区包含了你创建的管道里的数据流。**通常一个源组件会创建一个新的缓冲区,同时组件还将会把缓冲区的数据传递给下一个组件。**当使用 GStreamer 底层构造来创建一个媒体管道的时候,你不需要自己来处理缓冲区, 组件将会为你处理这些缓冲区。
在pads之间传送的数据封装在Buffer里,一个缓冲区主要由以下几个组成:
- 指向某块内存的指针
- 内存的大小
- 缓冲区的时间戳
- 一些metadata,包括
- Timestamp
- Offset
- Duration
- media type
- 其它
- 一个引用计数,指出了缓冲区所使用的组件数。没有组件可引用的时候,这个引用将用于销毁缓冲区。
在 push 模式下,element通过调用`gst_pad_push()`函数把buffer传送给对应的pad。在 pull 模式下,element通过调用`gst_pad_pull_range()`函数把数据 pull 过来。
element在push buffer之前需要确认对应的element具备处理buffer中的数据类型的能力。在传输之前首先查询对应的element能够处理的格式的种类,并从中选择合适的格式,通过`gst_buffer_set_caps()`函数对buffer进行设置,然后才传送数据。
收到一个buffer后,element要首先对buffer进行检查以确认是否能够处理。
可以调用`gst_buffer_new()`函数创建一个新的buffer,也可以调用`gst_pad_alloc_buffer()`函数申请一个可用的buffer。采用第二种方法接收数据的buffer可以设定接收其它类型的数据,这是通过对buffer的caps进行设定来实现的。
**选择媒体类型并对buffer进行设定的处理过程叫做caps negotianation。**
[GstBuffer对象描述了数据的缓冲区(buffers)的信息](https://gstreamer.freedesktop.org/documentation/gstreamer/gstbuffer.html?gi-language=c#GstBuffer)
### Events
用于element之间或者应用到element之间的信息传递,比如播放时的seek操作是通过event实现的。
事件是一系列控制粒子,随着缓冲区被发送到管道的上游和下游。下游事件通知流状态相同的组件,可能的事件包括中断,flush,流的终止信号等等。**在应用程序与组件之间的交互以及事件与事件之间的交互中, 上游事件被用于改变管道中数据流的状态,如查找。**对于应用程序来说,上游事件非常重要,下游事件则是为了说明获取更加完善的数据概念上的图像。
[GstEvent 对象描述了数据的事件(events)信息](https://gstreamer.freedesktop.org/documentation/gstreamer/gstevent.html?gi-language=c)
### Messages
是由element发出的消息,通过bus,以异步的方式被应用程序处理。通常用于传递errors, tags, state changes, buffering state, redirects等消息。消息处理是线程安全的。由于大部分消息是通过异步方式处理,所以会在应用程序里存在一点延迟,如果要及时的相应消息,需要在streaming线程捕获处理。
### Queries
用于应用程序向gstreamer查询总时间,当前时间,文件大小等信息。
# pipeline是啥?
pipeline是GStreamer设计的核心思想,在说pipeline之前先弄一个meidaplayer的模型,meidaplayer是GStreamer比较重要的应用之一。不管是VLC、mplayer这些开源的多媒体播放器,还是其他商用的。meidaplayer都能抽象成以下的模型:

上面这个图就是个典型的meida播放器的模型:
source: 数据来源,可能是file、http、rtp等.
demuxer: 分流器,负责把容器里的音视频数据剥离出来,然后分别送给audio\video decoder.
decoder: 解码,然后把解完后的数据(yuv、pcm)送给audio\video output输出.
output: 负责将decoder过来的数据呈现出来.
如果把数据想象成流水的话,每个模块的功能虽然不同,但是他们大致抽象的功能都是接收上个模块过来的数据,然后加工把加工后的数据送到下一个模块。
把上述这些模块通过某种方式连接起来,就形成了一个流水线(pipeline),这个流水线就是一个media播放器。
GStreamer 就是把每个模块都看做是一个Element,然后构建了连接和操作这些Element的方法,用户可以通过自己的需求把不同的Elements 排列组合,形成一个又一个的不同的pipeline。
你可以形成具有mediaplayer功能 的pipeline 、VOIP功能的pipeline。
# gstreamer tools
Gstreamer自带了gst-inspect-1.0和gst-launch-1.0等其他命令行工具,我们可以使用这些工具完成常见的处理任务。
## gst-inspect-1.0
查看gstreamer的plugin、element的信息。直接将plugin/element的类型作为参数,会列出其详细信息。如果不跟任何参数,会列出当前系统gstreamer所能查找到的所有插件。
>$ gst-inspect-1.0 playbin
## gst-launch-1.0
用于创建及执行一个Pipline,因此通常使用gst-launch先验证相关功能,然后再编写相应应用。一个pipeline的多个element之间通过 “!" 分隔,同时可以设置element及Cap的属性。例如:
播放音视频
>gst-launch-1.0 playbin file:///home/root/test.mp4
转码
>gst-launch-1.0 filesrc location=/videos/sintel_trailer-480p.ogv ! decodebin name=decode ! \
videoscale ! "video/x-raw,width=320,height=240" ! x264enc ! queue ! \
mp4mux name=mux ! filesink location=320x240.mp4 decode. ! audioconvert ! \
avenc_aac ! queue ! mux.
Streaming
#Server
>gst-launch-1.0 -v videotestsrc ! "video/x-raw,framerate=30/1" ! x264enc key-int-max=30 ! rtph264pay ! udpsink host=127.0.0.1 port=1234
#Client
>gst-launch-1.0 udpsrc port=1234 ! "application/x-rtp, payload=96" ! rtph264depay ! decodebin ! autovideosink sync=false
| github_jupyter |
```
import numpy as np
import pandas as pd
import scipy as sp
import matplotlib.pyplot as plt
import matplotlib as mpl
import seaborn as sns
sns.set()
```
### _csv 파일 가져오기_
```
df = pd.read_csv('~/src/properties_2016.csv')
submission = pd.read_csv('~/src/sample_submission.csv')
submission.tail(5)
train2016 = pd.read_csv('~/src/train_2016.csv')
train2016.tail(5)
```
## *bathfamily_df : bathroomcnt, calculatedbathnbr, threequarterbathnbr, fullbathcnt*
```
bathfamily = ["bathroomcnt", "calculatedbathnbr", "threequarterbathnbr", "fullbathcnt"]
bathfamily_df = pd.DataFrame(columns=bathfamily)
bathfamily_df
for i in bathfamily:
bathfamily_df[i] = df[i]
bathfamily_df.tail()
for i in bathfamily:
bathfamily_df[i].fillna(0, inplace=True)
bathfamily_df.tail()
sns.pairplot(bathfamily_df)
plt.show()
bathroomcnt_result = bathroomcnt_df.sort_values(by='probability', ascending=False)
bathroomcnt_result.reset_index(drop=True, inplace=True)
bathroomcnt_result = bathroomcnt_result.head(10)
bathroomcnt_result
x = bathroomcnt_df['bathroomcnt'].values
y = bathroomcnt_df['probability'].values
plt.figure(figsize = (8, 6))
plt.xticks(range(0, 20))
plt.bar(x, y)
# plt.pie(y, labels=labels, autopct='%d%%')
plt.show()
```
#### _calculatedbathnbr : Number of bathrooms in home including fractional bathroom_
- bathroomcnt와 거의 흡사하다. 그러나 방 갯수 0개인 데이터가 포함되지 않았다.
```
calbathroomcnt_df = df.groupby(['calculatedbathnbr']).size().reset_index(name='counts')
calbathroomcnt_counts = calbathroomcnt_df['counts'].values
calbathroomcnt_df['probability'] = calbathroomcnt_counts / sum(calbathroomcnt_counts)
calbathroomcnt_df
calbathroomcnt_result = calbathroomcnt_df.sort_values(by='probability', ascending=False)
calbathroomcnt_result.reset_index(drop=True, inplace=True)
calbathroomcnt_result = calbathroomcnt_result.head(10)
calbathroomcnt_result
x = calbathroomcnt_df['calculatedbathnbr'].values
y = calbathroomcnt_df['probability'].values
plt.figure(figsize = (8, 6))
plt.xticks(range(0, 20))
plt.bar(x, y)
# plt.pie(y, labels=labels, autopct='%d%%')
plt.show()
y = calbathroomcnt_result['probability'].values
labels = calbathroomcnt_result['calculatedbathnbr'].values
plt.figure(figsize = (8, 6))
plt.pie(y, labels=labels, autopct='%d%%')
plt.show()
```
#### _threequarterbathnbr : Number of 3/4 bathrooms in house (shower + sink + toilet)_
```
# 특정 속성 가져와서 갯수, 비율 칼럼 더하기
threequarterbathnbr_df = df.groupby(['threequarterbathnbr']).size().reset_index(name='counts')
threequarterbathnbr_counts = threequarterbathnbr_df['counts'].values
threequarterbathnbr_df['probability'] = threequarterbathnbr_counts / sum(threequarterbathnbr_counts)
threequarterbathnbr_df
threequarterbathnbr_result = threequarterbathnbr_df.sort_values(by='probability', ascending=False)
threequarterbathnbr_result.reset_index(drop=True, inplace=True)
threequarterbathnbr_result = threequarterbathnbr_result.head(10)
threequarterbathnbr_result
y = threequarterbathnbr_result['probability'].values
labels = threequarterbathnbr_result['threequarterbathnbr'].values
plt.figure(figsize = (8, 6))
plt.pie(y, labels=labels, autopct='%d%%')
plt.show()
```
#### _fullbathcnt : Number of full bathrooms (sink, shower + bathtub, and toilet) present in home_
```
# 특정 속성 가져와서 갯수, 비율 칼럼 더하기
fullbathcnt_df = df.groupby(['fullbathcnt']).size().reset_index(name='counts')
fullbathcnt_counts = fullbathcnt_df['counts'].values
fullbathcnt_df['probability'] = fullbathcnt_counts / sum(fullbathcnt_counts)
fullbathcnt_df
# 비율이 높은 순서대로 정렬하기
fullbathcnt_result = fullbathcnt_df.sort_values(by='probability', ascending=False)
fullbathcnt_result.reset_index(drop=True, inplace=True)
fullbathcnt_result = fullbathcnt_result.head(10)
fullbathcnt_result
# pi 그래프로 시각화
y = fullbathcnt_df['probability'].values
labels = fullbathcnt_df['fullbathcnt'].values
plt.figure(figsize = (8, 6))
plt.pie(y, labels=labels, autopct='%d%%')
plt.show()
```
### _room count_
```
roomcnt_df = df.groupby(['roomcnt']).size().reset_index(name='counts')
roomcnt_df['probability'] = roomcnt_counts / sum(roomcnt_counts) * 100
roomcnt_df
roomcnt_df = df.groupby(['roomcnt']).size().reset_index(name='counts')
roomcnt_df
x = roomcnt_df['roomcnt'].values
y = roomcnt_df['counts'].values
x, y
plt.figure(figsize = (20, 15))
plt.plot(x, y)
plt.xlim(0, 40)
plt.ylim(0, 3000000)
plt.title("room count graph")
plt.show()
```
### _fireplacecnt_
```
fireplacecnt_df = df.groupby(['fireplacecnt']).size().reset_index(name='counts')
fireplacecnt_df
```
## _taxfamily_
```
# 데이터프레임 만들기
taxfamily = ['taxvaluedollarcnt', 'structuretaxvaluedollarcnt',\
'landtaxvaluedollarcnt', 'taxamount']
taxfamily_df = pd.DataFrame(columns=taxfamily)
taxfamily_df
# 데이터프레임에 데이터 채워넣기
for i in taxfamily:
taxfamily_df[i] = df[i]
taxfamily_df.tail()
# NaN value, 0으로 바꾸기
taxfamily_df.fillna(value=0, inplace=True)
taxfamily_df.tail()
# 새로운 속성 structure+land 만들기
taxfamily_df['structure+land'] = taxfamily_df['structuretaxvaluedollarcnt'] + \
taxfamily_df['landtaxvaluedollarcnt']
taxfamily_df.tail()
sns.pairplot(taxfamily_df)
plt.show()
# 스캐터플롯 만들어서 비교하기
for i in taxfamily:
plt.subplot(2, 5, i)
plt.scatter(taxfamily_df[i])
plt.show()
x1 = bathfamily_df['bathroomcnt']
x2 = bathfamily_df['calculatedbathnbr']
x3 = bathfamily_df['threequarterbathnbr']
x4 = bathfamily_df["fullbathcnt"]
plt.subplot(231)
plt.scatter(x1, x2)
plt.show()
plt.subplot(232)
plt.scatter(x1, x3)
plt.show()
plt.subplot(233)
plt.scatter(x1, x4)
plt.show()
plt.subplot(234)
plt.scatter(x2, x3)
plt.show()
plt.subplot(235)
plt.scatter(x2, x4)
plt.show()
plt.subplot(236)
plt.scatter(x3, x4)
plt.show()
```
#### _taxvaluedollarcnt : The total tax assessed value of the parcel_
```
taxvaluedollarcnt_df = df.groupby(['taxvaluedollarcnt']).size().reset_index(name='counts')
taxvaluedollarcnt_df
print(np.mean(taxvaluedollarcnt_df['taxvaluedollarcnt'].values))
print(np.median(taxvaluedollarcnt_df['taxvaluedollarcnt'].values))
x = taxvaluedollarcnt_df['taxvaluedollarcnt'].values
y = taxvaluedollarcnt_df['counts'].values
plt.bar(x, y)
plt.show()
x = taxvaluedollarcnt_df['taxvaluedollarcnt'].values
y = taxvaluedollarcnt_df['counts'].values
plt.subplot(2, 2, 1)
plt.plot(x)
plt.show()
plt.subplot(2, 2, 2)
plt.plot(x)
plt.xlim(638000, 639000)
plt.show()
plt.subplot(2, 2, 3)
plt.plot(x)
plt.xlim(638500, 639000)
plt.show()
plt.subplot(2, 2, 4)
plt.plot(x)
plt.xlim(0, 638500)
plt.show()
```
## _finishedsquare family_
```
list(finishedsquare_df.columns)
finishedsquare_df = pd.DataFrame(columns=['finishedfloor1squarefeet','calculatedfinishedsquarefeet', 'finishedsquarefeet6', 'finishedsquarefeet12',\
'finishedsquarefeet13', 'finishedsquarefeet15',\
'finishedsquarefeet50'])
finishedsquare_df['finishedfloor1squarefeet'] = df['finishedfloor1squarefeet']
finishedsquare_df['calculatedfinishedsquarefeet'] = df['calculatedfinishedsquarefeet']
finishedsquare_df['finishedsquarefeet6'] = df['finishedsquarefeet6']
finishedsquare_df['finishedsquarefeet12'] = df['finishedsquarefeet12']
finishedsquare_df['finishedsquarefeet13'] = df['finishedsquarefeet13']
finishedsquare_df['finishedsquarefeet15'] = df['finishedsquarefeet15']
finishedsquare_df['finishedsquarefeet50'] = df['finishedsquarefeet50']
finishedsquare_df.tail()
print(finishedsquare_df['calculatedfinishedsquarefeet'].notna().sum())
print(finishedsquare_df['finishedsquarefeet6'].notna().sum()+\
finishedsquare_df['finishedsquarefeet12'].notna().sum()+\
finishedsquare_df['finishedsquarefeet13'].notna().sum()+\
finishedsquare_df['finishedsquarefeet15'].notna().sum())
print(finishedsquare_df['finishedfloor1squarefeet'].notna().sum())
print(finishedsquare_df['calculatedfinishedsquarefeet'].notna().sum())
print(finishedsquare_df['finishedsquarefeet6'].notna().sum())
print(finishedsquare_df['finishedsquarefeet12'].notna().sum())
print(finishedsquare_df['finishedsquarefeet13'].notna().sum())
print(finishedsquare_df['finishedsquarefeet15'].notna().sum())
print(finishedsquare_df['finishedsquarefeet50'].notna().sum())
```
### *region family*
```
region = ['regionidcounty', 'regionidcity', 'regionidzip', 'regionidneighborhood']
region, type(region)
region_df = pd.DataFrame(columns=region)
region_df
for i in region:
region_df[i] = df[i]
region_df.tail()
y = region_df['regionidcounty'].fillna(0).values[:1000]
len(y)
x = range(len(y))
plt.plot(x, y)
plt.show()
df['airconditioningtypeid']
airconditioningtype_df = df.groupby('airconditioningtypeid').size().reset_index(name = "Counts")
airconditioningtype_df
```
| github_jupyter |
# 00__preprocess_counts
in this notebook, i merge barcode counts from biological replicates into 1 dataframe. i also filter barcodes such that they have >= 5 counts in the DNA library, and i filter elements such that every element included has >= 3 barcodes represented at this filter.
```
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import math
import matplotlib.pyplot as plt
import numpy as np
import re
import seaborn as sns
import sys
from scipy.stats import spearmanr
# import utils
sys.path.append("../../../utils")
from plotting_utils import *
%matplotlib inline
%config InlineBackend.figure_format = 'svg'
mpl.rcParams['figure.autolayout'] = False
sns.set(**PAPER_PRESET)
fontsize = PAPER_FONTSIZE
np.random.seed(2019)
```
## functions
```
def import_dna(counts_dir, dna_f):
dna_dfs = []
for i in range(len(dna_f)):
f = dna_f[i]
cols = ["barcode", "dna_%s" % (i+1)]
tmp = pd.read_table("%s/%s" % (counts_dir, f), sep="\t")
tmp.columns = cols
dna_dfs.append(tmp)
if len(dna_dfs) > 1:
dna = reduce(lambda x, y: pd.merge(x, y, on = "barcode"), dna_dfs)
else:
dna = dna_dfs[0]
return dna
def import_rna(counts_dir, rna_f, dna):
data = dna.copy()
data_cols = list(dna.columns)
for f in rna_f:
rep = re.findall(r'\d+', f.split("__")[2])[0]
tmp = pd.read_table("%s/%s" % (counts_dir, f), sep="\t")
tmp.columns = ["barcode", "rna_%s" % rep]
data_cols.append("rna_%s" % rep)
data = data.merge(tmp, on="barcode", how="outer")
return data, data_cols
```
## variables
```
counts_dir = "../../../data/02__mpra/01__counts"
barcode_dna_read_threshold = 5
barcode_rna_read_threshold = 0
n_barcodes_per_elem_threshold = 3
```
### DNA files
```
dna_f = ["MPRA__DNA__rep1.BARCODES.txt"]
```
### RNA files
```
HUES64_rna_f = ["MPRA__HUES64__rep1__tfxn1.BARCODES.txt", "MPRA__HUES64__rep1__tfxn2.BARCODES.txt",
"MPRA__HUES64__rep1__tfxn3.BARCODES.txt", "MPRA__HUES64__rep2__tfxn1.BARCODES.txt",
"MPRA__HUES64__rep2__tfxn2.BARCODES.txt", "MPRA__HUES64__rep2__tfxn3.BARCODES.txt",
"MPRA__HUES64__rep3__tfxn1.BARCODES.txt", "MPRA__HUES64__rep3__tfxn2.BARCODES.txt",
"MPRA__HUES64__rep3__tfxn3.BARCODES.txt"]
HUES64_out_f = "HUES64__all_counts.txt"
mESC_rna_f = ["MPRA__mESC__rep1__tfxn1.BARCODES.txt", "MPRA__mESC__rep2__tfxn1.BARCODES.txt",
"MPRA__mESC__rep3__tfxn1.BARCODES.txt"]
mESC_out_f = "mESC__all_counts.txt"
```
### Index file
```
index_f = "../../../data/01__design/02__index/TWIST_pool4_v8_final.txt.gz"
```
## 1. import index
```
index = pd.read_table(index_f, sep="\t")
n_barc = len(index)
n_barc
index_elem = index[["element", "tile_type"]].drop_duplicates()
```
## 2. import dna
```
dna = import_dna(counts_dir, dna_f)
dna.head()
```
## 3. import rna
```
HUES64_data, HUES64_cols = import_rna(counts_dir, HUES64_rna_f, dna)
HUES64_data.head()
mESC_data, mESC_cols = import_rna(counts_dir, mESC_rna_f, dna)
mESC_data.head()
HUES64_data.columns = ["barcode", "dna_1", "HUES64_rep1_tfxn1", "HUES64_rep1_tfxn2", "HUES64_rep1_tfxn3",
"HUES64_rep2_tfxn1", "HUES64_rep2_tfxn2", "HUES64_rep2_tfxn3", "HUES64_rep3_tfxn1",
"HUES64_rep3_tfxn2", "HUES64_rep3_tfxn3"]
mESC_data.columns = ["barcode", "dna_1", "mESC_rep1_tfxn1", "mESC_rep2_tfxn1", "mESC_rep3_tfxn1"]
```
## 4. heatmap showing replicate corrs and clustering
```
# heatmap incl all libraries
tmp = HUES64_data.merge(mESC_data, on=["barcode", "dna_1"])
tmp = tmp.set_index("barcode")
tmp.drop("dna_1", axis=1, inplace=True)
tmp_cols = tmp.columns
tmp[tmp_cols] = np.log10(tmp[tmp_cols] + 1)
tmp_corr = tmp.corr(method="pearson")
cmap = sns.cubehelix_palette(as_cmap=True)
cg = sns.clustermap(tmp_corr, figsize=(5, 5), cmap=cmap, annot=True, vmin=0.7)
_ = plt.setp(cg.ax_heatmap.yaxis.get_majorticklabels(), rotation=0)
plt.suptitle("pearson correlation of replicates\nlog10+1 counts of all barcodes")
#cg.savefig("rep_and_lib_corr_heatmap.pdf", dpi="figure", bbox_inches="tight")
```
## 5. sum technical replicates
```
HUES64_data["HUES64_rep1"] = HUES64_data[["HUES64_rep1_tfxn1", "HUES64_rep1_tfxn2", "HUES64_rep1_tfxn3"]].sum(axis=1)
HUES64_data["HUES64_rep2"] = HUES64_data[["HUES64_rep2_tfxn1", "HUES64_rep2_tfxn2", "HUES64_rep2_tfxn3"]].sum(axis=1)
HUES64_data["HUES64_rep3"] = HUES64_data[["HUES64_rep3_tfxn1", "HUES64_rep3_tfxn2", "HUES64_rep3_tfxn3"]].sum(axis=1)
old_cols = [x for x in HUES64_data.columns if "_tfxn" in x]
HUES64_data.drop(old_cols, axis=1, inplace=True)
HUES64_data.columns = ["barcode", "dna_1", "rep_1", "rep_2", "rep_3"]
HUES64_data.head()
mESC_data.columns = ["barcode", "dna_1", "rep_1", "rep_2", "rep_3"]
mESC_data.head()
HUES64_data[["rep_1", "rep_2", "rep_3"]].sum(axis=0)
mESC_data[["rep_1", "rep_2", "rep_3"]].sum(axis=0)
```
## 6. filter barcodes
```
HUES64_data = HUES64_data.fillna(0)
mESC_data = mESC_data.fillna(0)
HUES64_data_filt = HUES64_data[HUES64_data["dna_1"] >= barcode_dna_read_threshold]
HUES64_data_filt.set_index("barcode", inplace=True)
mESC_data_filt = mESC_data[mESC_data["dna_1"] >= barcode_dna_read_threshold]
mESC_data_filt.set_index("barcode", inplace=True)
HUES64_reps = [x for x in HUES64_data_filt.columns if "rep_" in x]
mESC_reps = [x for x in mESC_data_filt.columns if "rep_" in x]
HUES64_data_filt[HUES64_reps] = HUES64_data_filt[HUES64_data_filt > barcode_rna_read_threshold][HUES64_reps]
HUES64_data_filt.reset_index(inplace=True)
HUES64_data_filt.head()
mESC_data_filt[mESC_reps] = mESC_data_filt[mESC_data_filt > barcode_rna_read_threshold][mESC_reps]
mESC_data_filt.reset_index(inplace=True)
mESC_data_filt.head()
all_names = ["HUES64", "mESC"]
all_dfs = [HUES64_data_filt, mESC_data_filt]
all_cols = [HUES64_data.columns, mESC_data.columns]
print("FILTERING RESULTS:")
for n, df, cs in zip(all_names, all_dfs, all_cols):
index_len = len(index)
dna_barc_len = len(df)
dna_barc_perc = (float(dna_barc_len)/index_len)*100
print("%s: from %s barcodes to %s at DNA level (%s%%)" % (n, index_len, dna_barc_len, dna_barc_perc))
reps = [x for x in cs if "rep_" in x]
for r in reps:
rep = r.split("_")[1]
rna_barc_len = sum(~pd.isnull(df[r]))
rna_barc_perc = (float(rna_barc_len)/index_len)*100
print("\trep %s: %s barcodes at RNA level (%s%%)" % (rep, rna_barc_len, rna_barc_perc))
print("")
```
## 7. filter elements
```
HUES64_data_filt = HUES64_data_filt.merge(index, on="barcode", how="inner")
mESC_data_filt = mESC_data_filt.merge(index, on="barcode", how="inner")
HUES64_barcodes_per_elem = HUES64_data_filt.groupby(["unique_name", "tile_type"])["barcode"].agg("count").reset_index()
HUES64_barcodes_per_elem_neg = HUES64_barcodes_per_elem[HUES64_barcodes_per_elem["tile_type"].isin(["RANDOM", "SCRAMBLED"])]
HUES64_barcodes_per_elem_no_neg = HUES64_barcodes_per_elem[~HUES64_barcodes_per_elem["tile_type"].isin(["RANDOM", "SCRAMBLED"])]
HUES64_barcodes_per_elem_no_neg_filt = HUES64_barcodes_per_elem_no_neg[HUES64_barcodes_per_elem_no_neg["barcode"] >= n_barcodes_per_elem_threshold]
HUES64_total_elems_rep = len(HUES64_barcodes_per_elem_no_neg)
HUES64_total_elems_filt_rep = len(HUES64_barcodes_per_elem_no_neg_filt)
mESC_barcodes_per_elem = mESC_data_filt.groupby(["unique_name", "tile_type"])["barcode"].agg("count").reset_index()
mESC_barcodes_per_elem_neg = mESC_barcodes_per_elem[mESC_barcodes_per_elem["tile_type"].isin(["RANDOM", "SCRAMBLED"])]
mESC_barcodes_per_elem_no_neg = mESC_barcodes_per_elem[~mESC_barcodes_per_elem["tile_type"].isin(["RANDOM", "SCRAMBLED"])]
mESC_barcodes_per_elem_no_neg_filt = mESC_barcodes_per_elem_no_neg[mESC_barcodes_per_elem_no_neg["barcode"] >= n_barcodes_per_elem_threshold]
mESC_total_elems_rep = len(mESC_barcodes_per_elem_no_neg)
mESC_total_elems_filt_rep = len(mESC_barcodes_per_elem_no_neg_filt)
print("ELEMENT FILTERING RESULTS:")
print("HUES64: filtered %s elements to %s represented at >= %s barcodes (%s%%)" % (HUES64_total_elems_rep, HUES64_total_elems_filt_rep,
n_barcodes_per_elem_threshold,
float(HUES64_total_elems_filt_rep)/HUES64_total_elems_rep*100))
print("HUES64: filtered %s elements to %s represented at >= %s barcodes (%s%%)" % (mESC_total_elems_rep, mESC_total_elems_filt_rep,
n_barcodes_per_elem_threshold,
float(mESC_total_elems_filt_rep)/mESC_total_elems_rep*100))
HUES64_good_elems = list(HUES64_barcodes_per_elem_no_neg_filt["unique_name"]) + list(HUES64_barcodes_per_elem_neg["unique_name"])
mESC_good_elems = list(mESC_barcodes_per_elem_no_neg_filt["unique_name"]) + list(mESC_barcodes_per_elem_neg["unique_name"])
HUES64_data_filt = HUES64_data_filt[HUES64_data_filt["unique_name"].isin(HUES64_good_elems)]
mESC_data_filt = mESC_data_filt[mESC_data_filt["unique_name"].isin(mESC_good_elems)]
```
## 8. heatmap comparing barcode counts [biological replicates only]
```
HUES64_cols = ["barcode"]
mESC_cols = ["barcode"]
HUES64_cols.extend(["HUES64_%s" % x for x in HUES64_reps])
mESC_cols.extend(["mESC_%s" % x for x in mESC_reps])
HUES64_cols
HUES64_counts = HUES64_data_filt.copy()
mESC_counts = mESC_data_filt.copy()
mESC_counts.head()
HUES64_counts = HUES64_counts[["barcode", "rep_1", "rep_2", "rep_3"]]
mESC_counts = mESC_counts[["barcode", "rep_1", "rep_2", "rep_3"]]
HUES64_counts.head()
HUES64_counts.columns = HUES64_cols
mESC_counts.columns = mESC_cols
HUES64_cols
all_samples = HUES64_counts.merge(mESC_counts, on="barcode", how="outer")
all_samples.drop("barcode", axis=1, inplace=True)
cols = [x for x in HUES64_cols if x != "barcode"]
all_samples[cols] = np.log10(all_samples[cols]+1)
all_samples_corr = all_samples.corr(method="pearson")
cmap = sns.cubehelix_palette(as_cmap=True)
cg = sns.clustermap(all_samples_corr, figsize=(3,3), cmap=cmap, annot=True)
_ = plt.setp(cg.ax_heatmap.yaxis.get_majorticklabels(), rotation=0)
plt.suptitle("pearson correlation of replicates\nlog10+1 counts of barcodes with >5 DNA counts")
plt.subplots_adjust(top=0.8)
cg.savefig("FigS4.pdf", dpi="figure", transparent=True, bbox_inches="tight")
```
## 10. write final files
```
HUES64_counts = HUES64_data_filt[["barcode", "dna_1", "rep_1", "rep_2", "rep_3"]]
mESC_counts = mESC_data_filt[["barcode", "dna_1", "rep_1", "rep_2", "rep_3"]]
HUES64_counts.to_csv("%s/%s" % (counts_dir, HUES64_out_f), sep="\t", header=True, index=False)
mESC_counts.to_csv("%s/%s" % (counts_dir, mESC_out_f), sep="\t", header=True, index=False)
```
| github_jupyter |
```
#!/usr/bin/env python
# -*- coding: UTF-8
```
# <p style="text-align: center;"> Creating and Refining Dictionaries with Word Embedding Models
<p style="text-align: left;">Charter School Identities Project<br/><br/>Creator: Jaren Haber, PhD Candidate<br/>Institution: Department of Sociology, University of California, Berkeley<br/>Date created: July 20, 2018<br/>Date last modified: September 27, 2019
## Initialize Python
```
# Import key packages
import gensim # for word embedding models
import _pickle as cPickle # Optimized version of pickle
import gc # For managing garbage collector
from collections import Counter # For counting terms across the corpus
# Import functions from other scripts
import sys; sys.path.insert(0, "../data_management/tools/") # To load functions from files in data_management/tools
from textlist_file import write_list, load_list # For saving and loading text lists to/from file
from df_tools import check_df, convert_df, load_filtered_df, replace_df_nulls # For displaying basic DF info, storing DFs for memory efficiency, and loading a filtered DF
from quickpickle import quickpickle_dump, quickpickle_load # For quickly loading & saving pickle files in Python
#import count_dict # For counting word frequencies in corpus (to assess candidate words)
#from count_dict import load_dict, Page, dict_precalc, dict_count, create_cols, count_words, collect_counts, count_master
# FOR CLEANING, TOKENIZING, AND STEMMING THE TEXT
from nltk import word_tokenize, sent_tokenize # widely used text tokenizer
from nltk.stem.porter import PorterStemmer # approximate but effective (and common) method of normalizing words: stems words by implementing a hierarchy of linguistic rules that transform or cut off word endings
# FOR VISUALIZATIONS
import matplotlib
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE # For visualizing word embeddings
from scipy.spatial import distance # To use cosine distances for tSNE metric
# Visualization parameters
%pylab inline
%matplotlib inline
#matplotlib.style.use('white')
import seaborn as sns # To make matplotlib prettier
sns.set(style='white')
#sns.despine()
# Define file paths
home = '/home/jovyan/work/'
wem_path = home + 'wem4themes/data/wem_model_300dims.bin' # path to WEM model
charter_path = home + 'misc_data/charters_2015.pkl' # path to charter school data file
dict_path = home + 'text_analysis/dictionary_methods/dicts/' # path to dictionary files (may not be used here)
# For counting term frequencies, load text corpus:
df = load_filtered_df(charter_path, ["WEBTEXT", "NCESSCH"])
df['WEBTEXT']=df['WEBTEXT'].fillna('') # turn nan to empty iterable for future convenience
```
## Define helper functions
```
def dict_cohere(thisdict, wem_model):
'''Computes the average cosine similarity score of terms within one dictionary with all other terms in that same dictionary,
effectively measuring the coherence of the dictionary.
...question for development: does it make sense to compare the average cosine similarity score between all terms
in thisdict and the average cosine similarity among the total model vocabulary? (Could that be, by definition, 0?)
NOTE: For an unknown reason, calling this function deletes terms from thisdict.
Inputs: List of key terms, word2vec model.
Output: Average cosine similarity score of each word with all other words in the list of key terms.'''
# Initialize average distance variables:
word_avg_dist = 0
word_avg_dists = 0
dict_avg_sim = 0
all_avg_dists = 0
model_avg_dists = 0
# Compute average cosine similarity score of each word with other dict words:
for word in thisdict:
word_avg_dist = (wem_model.distances(word, other_words=thisdict).sum())/len(thisdict) # Total diffs of word with all other words, take average
word_avg_dists += word_avg_dist # Add up each average distance, incrementally
dict_avg_sim = 1 - word_avg_dists/len(thisdict) # Find average cosine similarity score by subtracting avg. distance from 1
# For comparison, compute average cosine similarity score of each word with ALL other words in the model vocabulary:
#for word in thisdict:
# all_avg_dist = (wem_model.distances(word).sum())/len(model.vocab) # Default is to compare each word with all words
# all_avg_dists += all_avg_dist
#model_avg_dist = 1 - all_avg_dists/len(model.vocab) # Find average cosine similarity score by subtracting avg. distance from 1
#print("Average cosine similarities by word for this dictionary: \t" + str(dict_avg_dist))
#print("Compare to avg. cosine similarities by dict words to ALL words:\t" + str(model_avg_dist))
return dict_avg_sim
def expand_dict(thisdict, coredict, maxlen, wem_model):
'''TO DO: Create function to implement random additions to core dictionary,
in contrast to removing least similar word vectors--as focus_dict() does.'''
#code here
thisdict = list(set(thisdict)) # Remove any duplicates
if len(thisdict) != maxlen: # Quality check
print("WARNING: Function produced a dictionary of length " + str(len(thisdict)) + \
", which is not the specified maximum dict length of " + str(maxlen))
return thisdict
def focus_dict(thisdict, coredict, maxlen, wem_model):
'''Focus thisdict by removing least similar word vectors until reaching maxlen.
If any words from coredict get removed, compensate for fact that they will get added back in.
Input: A list of terms, core terms not to remove, desired length, and word2vec model.
Output: The input list focused down to desired length, and still containing all the core terms.'''
core_count = 0 # Counts number of coredict terms that were removed
extend_count = 0 # Counts number of terms removed to offset the coming boost of core terms (that were removed and will be added back in)
while len(thisdict) > maxlen: # Narrow thisdict down to maxlen
badvar = model.doesnt_match(thisdict) # Find least matching term
thisdict.remove(badvar) # Remove that least focal term, to focus dict
if badvar in coredict: # Keep track of number of core terms removed
core_count += 1
while extend_count < core_count: # Remove terms until length = maxlen - number of core terms removed (to offset those core terms that will be added back in later in this script)
badvar = model.doesnt_match(thisdict) # Find least matching term
thisdict.remove(badvar) # Remove that least focal term, to focus dict
extend_count += 1 # Keep track of # non-core terms added
if badvar in coredict: # Keep track of number of core terms removed
core_count += 1
for term in coredict: # Add back in any missing core terms
if term not in thisdict and term in list(model.vocab):
thisdict.append(term)
thisdict = list(set(thisdict)) # Remove any duplicates
if len(thisdict) != maxlen: # Quality check
print("WARNING: Function produced a dictionary of length " + str(len(thisdict)) + \
", which is not the specified maximum dict length of " + str(maxlen))
return thisdict
def display_closestwords(wem_model, word, plotnumber):
'''Use tSNE to display a spatial map (i.e., scatterplot) of words vectors close to given word in Word2Vec model.
Projects each word to a 2D space (a reduction from model size) and plots the position of each word with a label.
Based on this blog:
https://medium.com/@aneesha/using-tsne-to-plot-a-subset-of-similar-words-from-word2vec-bb8eeaea6229)
TO DO: Incorporate bold seed terms, OPTION for getting most similar words
Args:
Word2Vec model
Input word
Number of words to plot
Returns:
Visualization of relationships between given word and its closest word vectors
'''
dimensions = 300
arr = np.empty((0,dimensions), dtype='f')
word_labels = [word]
# get close words
close_words = wem_model.similar_by_word(word, plotnumber)
# add the vector for each of the closest words to the array
arr = np.append(arr, np.array([model[word]]), axis=0)
for wrd_score in close_words:
wrd_vector = model[wrd_score[0]]
word_labels.append(wrd_score[0])
arr = np.append(arr, np.array([wrd_vector]), axis=0)
# find tsne coords for 2 dimensions
tsne = TSNE(n_components=2, random_state=0, metric=distance.cosine)
np.set_printoptions(suppress=True)
Y = tsne.fit_transform(arr)
x_coords = Y[:, 0]
y_coords = Y[:, 1]
sns.set(style='whitegrid')
# display scatter plot
fig, ax = plt.subplots(figsize=(20,20))
ax.scatter(x_coords, y_coords, alpha=1, color='b')
for label, x, y in zip(word_labels, x_coords, y_coords):
plt.annotate(label, xy=(x, y), xytext=(0, 0), textcoords='offset points').set_fontsize(20)
plt.xlim(x_coords.min()+10, x_coords.max()+10)
plt.ylim(y_coords.min()+10, y_coords.max()+10)
plt.show()
```
## Load word embedding model
```
model = gensim.models.KeyedVectors.load_word2vec_format(wem_path, binary=True) # Load word2vec model
# dictionary of words in model (may not work for old gensim)
print(len(model.vocab))
sorted(list(model.vocab)[:200])
```
## Inductive search
### Inspect similar vectors
```
# Repeat inductive process of exploration for a list of seed terms like:
inqseed = ['inquiry-based', 'problem-based', 'discovery-based', 'experiential', 'constructivist']
model.most_similar(inqseed, topn = 20)
# Find the 30 words closest to test word:
test_word = "at-risk"
candidate_sims = model.most_similar(test_word, topn=30)
candidates_list = [pair[0] for pair in candidate_sims] # Convert to list for frequency search below
candidate_sims
model.most_similar("zero_tolerance")
riskseed = ["high-need", "low-income", "high-poverty", "at-risk", "high-risk"]
risk20 = ["high-need", "high-needs",
"low-income", "lower-income", "high-poverty", "impoverished", "economically_challenged",
"underserved", "under-served", "disproportionately",
"at-risk", "high-risk", "under-resourced", "under-represented", "under-performing",
"inner-city", "inner_cities",
"marginalized", "disconnected", "disenfranchised"] #"justice", "afflicting"
model.most_similar(atrisk20, topn = 20)
model.doesnt_match(atrisk20)
```
### Overview of search process
By using the model to look at similar words across terms, create a list of candidate terms for a bigger conceptual dictionary. Manually search all these candidate terms for those that are tightly conceptually related to the seed dictionary. This process blends data-driven search from the model with hand-driven search across the candidate terms.
```
# Find the 15 words closest to each term in seed dictionary:
for term in inqseed:
print("\nT0P 15 WORDS CLOSEST TO " + term)
for line in list(model.most_similar(term, topn=15)):
print(line)
#model.most_similar("problem-based", topn=30)
# Can also look for most similar terms across seed dictionary as a whole, and use it to make new dictionary:
inq50new = []
for line in list(model.most_similar(inqseed, topn=50)):
#print(line[0]) # only print words
inq50new.append(line[0])
inq50new
# By searching through the above candidate terms/phrases, expand from the seed terms into a conceptually tight list like this:
inq50 = ['inquiry-based', 'problem-based', 'discovery-based', 'experiential', 'constructivist', 'hands-on',
'experiential_learning', 'creative', 'problem-solving', 'critical_thinking', 'real_world', 'creativity',
'project-based', 'exploration', 'curiosity', 'investigate', 'real-life', 'student-centered', 'critical_thinkers',
'expeditions', 'child-centered', 'experimentation', 'explorations', 'student-based', 'immersive', 'activity-based',
'open-mindedness', 'student-driven', 'intrinsically', 'reality-based', 'learner-centered', 'interest-based',
'minds-on', 'metacognitive', 'experience-based', 'constructivism', 'project-based', 'student-directed',
'project-oriented', 'inquiry/research', 'project-focused', 'stimulate_critical', 'student-centric', 'active_inquiry',
'inquiry-driven', 'child-directed', 'child-initiated', 'experientially', 'problem-centered', 'project-centered']
# Could add more terms, too:
model.most_similar(inq50, topn=100)
# Can also search for terms that don't match:
model.doesnt_match(inq50)
# Searching for additional terms similar to this list, you can expand even further!
inquiry_fin = [elem.strip('\n') for elem in load_list('data/inquiry.txt')] # Load completed dict of 500 terms
inquiry_fin = list(set(inquiry_fin)) # Remove duplicates
sorted(inquiry_fin) # Show long dictionary resulting from exploring (and hand-cleaning)
# Remove any terms from full dict NOT in current model:
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
print("Removed " + str(word) + " from core dictionary.")
# Repeat for quality:
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
print("Removed " + str(word) + " from core dictionary.")
```
## Visualize vector relationships
### Most similar words
```
# Display 30 closest words to test word
display_closestwords(model, test_word, 30)
```
### Manually defined words
```
# TO DO: Adapt function above to be able to perform this visualization
# Set terms for visualization:
core_words = inqseed
close_words = inq50
# Define model parameters
dimensions = 300
arr = np.empty((0,dimensions), dtype='f')
word_labels = []
# add the vector for each of the closest words to the array
for wrd_label in close_words:
wrd_vector = model[wrd_label]
word_labels.append(wrd_label)
arr = np.append(arr, np.array([wrd_vector]), axis=0)
# find tsne coords for 2 dimensions, using cosine distances as metric
tsne = TSNE(n_components=2, random_state=0, metric=distance.cosine)
np.set_printoptions(suppress=True)
Y = tsne.fit_transform(arr)
x_coords = Y[:, 0]
y_coords = Y[:, 1]
# display scatter plot
fig, ax = plt.subplots(figsize=(20,20))
ax.scatter(x_coords, y_coords, alpha=1, color='b')
sns.set(style='white')
for label, x, y in zip(word_labels, x_coords, y_coords):
if label in core_words:
ax.annotate(label, xy=(x, y), xytext=(0, 0), textcoords='offset points', fontweight='bold').set_fontsize(20)
else:
ax.annotate(label, xy=(x, y), xytext=(0, 0), textcoords='offset points').set_fontsize(20)
ax.set_xlim(x_coords.min()-5, x_coords.max()+50)
ax.set_ylim(y_coords.min()-5, y_coords.max()+5)
ax.set_xlabel('t-SNE dimension 1')
ax.set_ylabel('t-SNE dimension 2')
ax.xaxis.label.set_fontsize(24)
ax.yaxis.label.set_fontsize(24)
for item in (ax.get_xticklabels() + ax.get_yticklabels()):
item.set_fontsize(16)
plt.show()
fig.savefig("data/IBL_core_051519.png", facecolor="white", transparent=True)
```
## Check word frequencies and ranks
### Count frequency of candidate words
Required arguments for `count_master()` function:
- df: DataFrame with text data, each of which is a list of full-text pages (not necessarily preprocessed)
- dict_path: file path to folder containing dictionaries
- dict_names: names of dictionaries on file (list or list of lists)
- file_ext: file extension for dictionary files (probably .txt)
- local_dicts: list of local dictionaries formatted as last of lists of terms--or if singular, just a list of terms
- local_names: names of local dictionaries (list or list of lists)
```
countsdfs = count_master(df, dict_path = dict_path, dict_names = [], file_ext = '.txt',
local_dicts = [candidates_list], local_names = ["candidates"])
```
### Relative ranks of dictionary terms
Word2vec also has built-in method to compare relative ranks of terms, which go from most frequent (highest number) to least frequent. <br/> **Note** that they are NOT real counts/frequencies (see above for that).
```
# Create Counter object: a set containing doubles, where each double is ('term': rank)
counts = Counter({word: vocab.count for (word, vocab) in model.wv.vocab.items()})
counts
# TO DO: 1. Sort entries by rank before displaying; 2. show histogram of term counts for highest 20 and lowest 20 words for each dict
print("RANK FOR SEED DICTIONARY:")
for term in sorted(inqseed):
print(str(term) + ": " + str(counts[term]))
print()
print("RANK FOR CORE DICTIONARY:")
for term in inq30:
print(str(term) + ": " + str(counts[term]))
print()
print("RANK FOR FULL DICTIONARY:")
for term in inquiry_fin:
print(str(term) + ": " + str(counts[term]))
# 50 most frequent words (according to word2vec tracker):
model.wv.index2entity[:50]
```
## Validate dictionaries: Check coherence of terms across sizes
```
for dictionary in [inqseed, inq30, inq50new, inquiry_fin]:
print(str(dict_cohere(dictionary, model)))
# Can also check for words that belong LEAST to a given dictionary:
maindict = inq30
i = 0 # initialize iterator
length = len(maindict) # set length for the for loop
inqtemp = []; inqtemp = maindict # initialize temp list of dict words
similar_words = [] # initialize list of dict words sorted by similarity (least similar words most recently added -> at bottom)
while i < length:
least_similar_word = model.doesnt_match(inqtemp)
similar_words.append(least_similar_word)
inqtemp.remove(least_similar_word)
i += 1 # add to iterator
print("List of words from least to most similar:")
for term in similar_words:
print(term)
# Need to re-define dictionary again at this point:
inq30 = ['discovery-based', 'student-driven_exploration', 'exploration_and_experimentation', 'laboratory-based',
'problem-based', 'prbl', 'learn-by-doing',
'project-based', 'project-centered',
'experiential', 'experiential_approach', 'experientially',
'inquiry-based', 'inquiry-driven', 'student-centered_inquiry-based', 'active_inquiry',
'constructivist', 'constructivism',
'hands-on', 'hand-on', 'hands-on_learning', 'hands-on_and_minds-on', 'hands-on_minds-on', 'hands-on/minds-on',
'socratic', 'socratic_method', 'socratic_dialogue',
'child-centered', 'learner-centered', 'student-centered']
```
### Check integrity of IBL dictionaries of varying lengths
```
# Remove any terms from core dict NOT in current model (these will have to be replaced):
for word in inq30:
if word not in list(model.vocab):
inq30.remove(word)
print("NOT FOUND IN CORE DICT, REMOVING:\t" + word)
count = 0
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
count += 1
#print("NOT FOUND in FULL DICT, REMOVING:\t" + word)
# Loop twice to catch any remaining OOV terms:
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
count += 1
print("Removed " + str(count) + " out-of-vocabulary terms from full dictionary.")
print("Length of original dict: ", len(inq30))
print("Coherence of original dict: ", str(dict_cohere(inq30, model)))
print("Length of extended dict: ", len(inquiry_fin))
print("Coherence of extended dict: ", str(dict_cohere(inquiry_fin, model)))
# Use random adds to create a few dicts of different lengths from max length of 500:
print("TESTING DIFFERENT LENGTHS USING RANDOM ADDITIONS:\n")
# TO DO: Write function expand_dict() above to automate this
inqlist = []; inqlist = inq30 # initialize placeholder for each growing dict
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq30, model)))
print()
# 40 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq30, inquiry_fin) if term in list(model.vocab)], size=10, replace=False))
inq40 = []; inq40 = inqlist
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq40, model)))
print()
# 50 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq40, inquiry_fin) if term in list(model.vocab)], size=10, replace=False))
inq50 = inqlist
#print("50 terms: \n", [term for term in inq50])
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq50, model)))
print()
# 75 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq50, inquiry_fin) if term in list(model.vocab)], size=25, replace=False))
inq75 = inqlist
#print("75 terms: \n", [term for term in inq75])
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq75, model)))
print()
# 100 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq75, inquiry_fin) if term in list(model.vocab)], size=25, replace=False))
inq100 = inqlist
#print("100 terms: \n", [term for term in inq100])
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq100, model)))
print()
# 200 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq100, inquiry_fin) if term in list(model.vocab)], size=100, replace=False))
inq200 = inqlist
#print("200 terms: \n", [term for term in inq200])
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq200, model)))
print()
# 300 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq200, inquiry_fin) if term in list(model.vocab)], size=100, replace=False))
inq300 = inqlist
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq300, model)))
print()
# 400 terms:
inqlist.extend(np.random.choice([term for term in filter(lambda x: x not in inq300, inquiry_fin) if term in list(model.vocab)], size=100, replace=False))
inq400 = inqlist
print(str(len(inqlist)) + "-term dict has this much coherence:", str(dict_cohere(inq400, model)))
print()
print("500-term dict has this much coherence:", str(dict_cohere(inquiry_fin, model)))
# TO DO: Fix this
# Use removal of least similar word vectors to create a few dicts of different lengths from max length of 500:
lens_list = [500, 400, 300, 200, 100, 75, 60, 50, 40, 30] # Define desired dict lengths
# Load full 500-term dicts:
inquiry_fin = [elem.strip('\n') for elem in load_list('data/inquiry.txt')]
# Remove any terms from core dict NOT in current model (these will have to be replaced):
for word in inq30:
if word not in list(model.vocab):
inq30.remove(word)
print("NOT FOUND IN CORE DICT, REMOVING:\t" + word)
print()
count = 0
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
count += 1
#print("NOT FOUND in FULL DICT, REMOVING:\t" + word)
for word in inquiry_fin:
if word not in list(model.vocab):
inquiry_fin.remove(word)
count += 1
print("Removed " + str(count) + " out-of-vocabulary terms from full dictionary.")
print('TESTING DICT LENGTHS BY REMOVING LEAST SIMILAR WORD VECTORS:\n')
for dictlen in lens_list:
inqdict = [],
inqdict = focus_dict(inquiry_fin, inq30, dictlen, model)
print(str(dictlen) + "-term dicts have this much coherence:", str(dict_cohere(inqdict, model)))
print()
```
## Surprising trends in analyzing how dictionary length affects inter-dictionary opposition and within-dictionary coherence (from this and other analyses):
### Greater lengths make for more within-dictionary coherence, but less across-dictionary opposition.
### Trimming word vectors randomly yields dictionaries that are less coherent and more opposing than does trimming those word vectors least similar to the others in the dictionary.
#### Caveat: The above patterns assume (perhaps consequentially, perhaps not) that as we trim dictionaries, we are keeping a small, core list of concepts (here, 30) intact within each dictionary. The size of this core may influence the above dynamics, as may the relative size of the total dictionary compared to the core list.
| github_jupyter |
```
from sys import path as syspath
from os import path as ospath
import pickle
from cbsa import ReactionSystem
import numpy as np
import matplotlib.pyplot as plt
S = [[1 ,-1,1,-2,0],
[-1,-1,0,0,1],
[0 ,0 ,1,0,-1]]
R = [[0,0,1,0,0],
[0,0,0,0,0],
[0,0,0,0,0]]
Y1s = 500
Y2s = 1000
Y3s = 2000
ro1 = 2000
ro2 = 50000
c1X1 = ro1/Y2s
c2 = ro2/(Y1s*Y2s)
c3X2 = (ro1+ro2)/Y1s
c4 = 2*ro1/(Y1s**2)
c5X3 = (ro1+ro2)/Y3s
x = [500,1000,2000]
k = [c1X1,c2,c3X2,c4,c5X3]
cbsa = ReactionSystem(S,R)
cbsa.setup()
cbsa.set_x(x)
cbsa.set_k(k)
total_sim_time = 6.0
cbsa.setup_simulation(use_opencl=False,alpha=0.5,max_dt=0.001)
cbsa.compute_simulation(total_sim_time,batch_steps=1)
cbsa_data = np.array(cbsa.simulation_data)
from matplotlib import rc
from mpl_toolkits import mplot3d
fontsize = 14
rc('text', usetex=True)
plt.style.use("bmh")
plt.rcParams["font.family"] = "serif"
plt.rcParams["xtick.labelsize"] = 12
plt.rcParams["ytick.labelsize"] = 12
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
fig = plt.figure(figsize=(10, 6),constrained_layout=True)
gs = fig.add_gridspec(nrows=2, ncols=4)
ax0 = fig.add_subplot(gs[0:1,0:2])
ax1 = fig.add_subplot(gs[1:2,0:2])
ax2 = fig.add_subplot(gs[0:1,2:3],projection='3d')
ax3 = fig.add_subplot(gs[0:1,3:4])
ax4 = fig.add_subplot(gs[1:2,2:3])
ax5 = fig.add_subplot(gs[1:2,3:4])
ax0.plot(cbsa_data[:,0],cbsa_data[:,1],label="Y1")
ax0.plot(cbsa_data[:,0],cbsa_data[:,2],label="Y2")
ax0.plot(cbsa_data[:,0],cbsa_data[:,3],label="Y3")
ax0.set_xlim(0,6)
#ax0.set_xticklabels([])
ax0.set_ylabel("Number of Molecules",fontsize=fontsize)
#ax0.text(-0.16,0.9,r'\textbf{a)}', transform=ax0.transAxes,fontsize=18)
ax1.plot(cbsa_data[:,0],cbsa_data[:,1],label="Y1")
ax1.plot(cbsa_data[:,0],cbsa_data[:,2],label="Y2")
ax1.plot(cbsa_data[:,0],cbsa_data[:,3],label="Y3")
ax1.set_xlim(0,3)
ax1.set_ylabel("Number of Molecules",fontsize=fontsize)
ax1.set_xlabel("time",fontsize=fontsize)
ax1.legend()
#ax1.text(-0.16,0.9,r'\textbf{b)}', transform=ax1.transAxes,fontsize=18)
ax2.plot3D(cbsa_data[:,1],cbsa_data[:,2],cbsa_data[:,3])
ax2.view_init(elev=30, azim=50)
ax2.set_zlabel("Y3",fontsize=fontsize-2)
ax2.set_ylabel("Y2",fontsize=fontsize-2)
ax2.set_xlabel("Y1",fontsize=fontsize-2)
ax2.set_xticklabels([])
ax2.set_yticklabels([])
ax2.set_zticklabels([])
ax3.plot(cbsa_data[:,1],cbsa_data[:,2])
ax3.set_ylabel("Y2",fontsize=fontsize)
ax3.set_xlabel("Y1",fontsize=fontsize)
ax4.plot(cbsa_data[:,1],cbsa_data[:,3])
ax4.set_ylabel("Y3",fontsize=fontsize)
ax4.set_xlabel("Y1",fontsize=fontsize)
ax5.plot(cbsa_data[:,2],cbsa_data[:,3])
ax5.set_ylabel("Y3",fontsize=fontsize)
ax5.set_xlabel("Y2",fontsize=fontsize)
plt.savefig("oregonator.png",dpi=300, bbox_inches='tight')
plt.show()
```
| github_jupyter |
# Theory Interlude - Tensor and Flow
First of all: Congratulations! You have made it pretty far and built some advanced machine learning systems already. Before we continue, it is time for a little theory interlude. In this chapter, we will peak under the hood of our systems, and see what is going on there. This knowledge will directly help us build even better systems in the next chapters.
## TensorFlow
You might have noticed something peculiar about Keras already. Each time we import it, it gives out a notification about TensorFlow:
```
import keras
```
Keras is a high level library and can be used as a simplified interface to [TensorFlow](https://www.tensorflow.org/). That means, Keras does not do any computations by itself, it is just a simple way to interact with TensorFlow, which is running in the background.

TensorFlow is a clever piece of software library developed by Google. It is very popular for deep learning. In this material we usually try to work with TensorFlow only through Keras, since that is easier than working with TensorFlow directly. However, sometimes we might want to write a bit of TensorFlow code to build more advanced models.
The goal of TensorFlow is to run the computations needed for deep learning as fast as possible. It does so, as the name gives away, by working with tensors in a data flow graph.
## What is a Tensor?
Tensors are arrays of numbers that transform based on specific rules. The simplest kind of tensor is a single number, also called a scalar. Scalars are sometimes referred to as rank zero tensors. The next bigger kind of tensor is a vector, also called rank one tensor. Next higher up the order are matrices, called rank two tensors, cube matrices called rank three tensors and so on.
|Rank|Name|Expresses|
|----|----|--------|
|0|Scalar|Magnitude|
|1|Vector|Magnitude & Direction|
|2|Matrix|Table of numbers|
|3|Cube Matrix|Cube of numbers|
|n|n-dimensional matrix|You get the idea|

Think back to the first week and how we constructed a neural network from scratch. Take a second to think about the numpy matrices and vectors we used. If you can not remember the exact setup, go back to the early chapters and look up the graphics showing our simple neural net.
**Question:** What kind of tensors where used in the neural net built from scratch in week 1?
Take a minute to answer the question without looking at the graphic below

As you can see, all matrices and vectors in our neural network are tensors.
## What about the Flow?
TensorFlow (and every other deep learning library) performs calculations along a 'computational graph'. In a computational graph, operations, such as a matrix multiplication or an activation function are nodes in a network. Tensors get passed along the edges of the graph between the different operations. A forward pass through our simple neural net has the following graph:

The advantage of structuring computations as a graph is that it is easy to run nodes in parallel. Through parallel computation we do not need one very fast machine, we can also achieve fast computation with many slow ones which split up tasks. This is why graphical processing units (GPU's) are so useful for deep learning. They have many small and cores, other than CPU's which only have a few, but fast cores. While a modern CPU might have 4 cores, a modern GPU can have hundreds or even thousands of cores. The entire graph of just a very simple model can look quite complex, but you can see the components of the dense layer: There is a matrix multiplication (matmul), adding of a bias and a relu activation function:

## Derivatives on Computational Graphs
Another advantage of using computational graphs like this is that TensorFlow and other libraries can quickly and automatically calculate derivatives along this graph. As we know from week 1, calculating derivatives is key for training neural networks.
## Summary
- TensorFlow works with Computational Graphs and Tensors
- Operations are 'nodes' in the computational graph
- Tensors get passed along the edges of the computational graph
- TensorFlow can automatically compute derivatives along the computational graph
With this overview of tensors and data flows you are ready to dive deep into advanced models. Let's go!
| github_jupyter |
```
from __future__ import division
import numpy as np
from scipy.stats import norm
from scipy.optimize import curve_fit
from scipy import stats
import random
import tqdm
import pandas as pd
from collections import OrderedDict,Counter
import matplotlib.pyplot as plt
import heapq
import pickle
import math
import seaborn as sns
from matplotlib import pylab
from mpl_toolkits.axes_grid1.inset_locator import InsetPosition
```
### SORN 1 Model Analysis:
#### Connection connection dynamics
#### Histogram of incoming and outgoing connection numbers
#### Population activity/ Fraction of active units/time step
#### Spike rate neurons
#### Spike train plot with firing rate of network
#### Firing rate Smoothness measure
#### Number of active units per timestep
#### Average Correlation coefficient between neurons
#### ISI with Exponential fit
#### ISI-CV
#### Histogram of weight strengths in SORN at 10000th time step and their distribution fit
#### NOTE: Hamming distance, Spike source entropy, Fano factor are in different notebook
### Unpickle simulation matrices for analysis
```
with open('conn_2009_all.pkl','rb') as f:
conn2009_all = pickle.load(f)
with open('conn_2009_NoIP.pkl','rb') as f:
conn2009_NoIP = pickle.load(f)
with open('conn_2009_NoSN.pkl','rb') as f:
conn2009_NoSN = pickle.load(f)
with open('conn_2013_all.pkl','rb') as f:
conn2013_all = pickle.load(f)
with open('conn_2018_NOIP.pkl','rb') as f:
conn2013_NoIP = pickle.load(f)
with open('conn_2013_NoSN.pkl','rb') as f:
conn2013_NoSN = pickle.load(f)
with open('conn_2013_NoiSTDP.pkl','rb') as f:
conn2013_NoiSTDP = pickle.load(f)
# with open('stdp2013_100k.pkl','rb') as f:
# all_matrices1, X,frac_ee_conn6 = pickle.load(f)
# with open('stdp2013_150k.pkl','rb') as f:
# all_matrices1, X,frac_ee_conn7 = pickle.load(f)
# with open('stdp2013_200k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn8 = pickle.load(f)
# with open('stdp2013_290k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn9 = pickle.load(f)
# with open('stdp2013_380k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn10 = pickle.load(f)
# with open('stdp2013_470k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn11 = pickle.load(f)
# with open('stdp2013_500k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn12 = pickle.load(f)
# with open('stdp2013_590k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn13 = pickle.load(f)
# with open('stdp2013_680k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn14 = pickle.load(f)
# with open('stdp2013_770k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn15 = pickle.load(f)
# with open('stdp2013_860k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn16 = pickle.load(f)
# with open('stdp2013_950k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn17 = pickle.load(f)
# with open('stdp2013_1000k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn18 = pickle.load(f)
# with open('stdp2013_1090k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn19 = pickle.load(f)
# with open('stdp2013_1180k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn20 = pickle.load(f)
# with open('stdp2013_1270k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn21 = pickle.load(f)
# with open('stdp2013_1360k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn22 = pickle.load(f)
# with open('stdp2013_1450k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn23 = pickle.load(f)
# with open('stdp2013_1540k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn24 = pickle.load(f)
# with open('stdp2013_1630k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn25 = pickle.load(f)
# with open('stdp2013_1720k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn26 = pickle.load(f)
# with open('stdp2013_1810k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn27 = pickle.load(f)
# with open('stdp2013_1900k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn28 = pickle.load(f)
# with open('stdp2013_1950k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn29 = pickle.load(f)
# with open('stdp2013_2000k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn30 = pickle.load(f)
# with open('stdp2013_2090k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn31 = pickle.load(f)
# with open('stdp2013_2180k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn32 = pickle.load(f)
# with open('stdp2013_2270k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn33 = pickle.load(f)
# with open('stdp2013_2360k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn34 = pickle.load(f)
# with open('stdp2013_2450k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn35 = pickle.load(f)
# with open('stdp2013_2540k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn36 = pickle.load(f)
# with open('stdp2013_2630k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn37 = pickle.load(f)
# with open('stdp2013_2720k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn38 = pickle.load(f)
# with open('stdp2013_2810k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn39 = pickle.load(f)
# with open('stdp2013_2900k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn40 = pickle.load(f)
# with open('stdp2013_3000k.pkl','rb') as f:
# all_matrices, X,frac_ee_conn41 = pickle.load(f)
```
### Number of incoming and outgoing connections
```
with open('stdp2009_50k_all.pkl','rb') as f:
plastic_matrices,X_all,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
weights = plastic_matrices['Wee']
num_incoming_weights = np.sum(np.array(weights) > 0, axis=0)
num_outgoing_weights = np.sum(np.array(weights) > 0, axis=1)
print(np.unique(num_incoming_weights))
print(np.unique(num_outgoing_weights))
f = plt.figure(figsize=(12,5))
ax = f.add_subplot(121)
ax2 = f.add_subplot(122)
ax.set_title('Number of incoming connections')
ax.set_xlabel('Number of connections')
ax.set_ylabel('Count')
ax.hist(num_incoming_weights, bins = 10,histtype = 'step')
ax2.hist(num_outgoing_weights,bins = 10,histtype = 'step')
ax2.set_title('Number of Outgoing connections')
ax2.set_xlabel('Number of connections')
ax2.set_ylabel('Count')
plt.show()
```
#### Guassian fit for histogram of number of incoming connections(Pre- synapses)
```
# Empirical average and variance are computed
avg = np.mean(num_incoming_weights)
var = np.var(num_incoming_weights)
# From hist plot above, it is clear that connection count follow gaussian distribution
pdf_x = np.linspace(np.min(num_incoming_weights),np.max(num_incoming_weights),100)
pdf_y = 1.0/np.sqrt(2*np.pi*var)*np.exp(-0.5*(pdf_x-avg)**2/var)
# Then we plot :
plt.figure()
plt.hist(num_incoming_weights,bins= 6,normed=True)
plt.plot(pdf_x,pdf_y,'k--',label = 'Fit')
plt.title('Gaussian fit')
plt.xlabel('Number of incoming connections')
plt.ylabel('Frequency')
plt.axvline(x=avg, color='r', linestyle='--',label = 'Mean')
plt.legend()
plt.show()
```
### Network connection dynamics: decay and stable phases
```
plt.figure(figsize = (12,8))
plt.plot(conn2009_all,label = 'SORN 2009 with all plasticity')
plt.plot(conn2009_NoIP,label = 'SORN 2009 without IP')
plt.plot(conn2009_NoSN,label = 'SORN 2009 without SN')
plt.plot(conn2013_all,label = 'SORN 2013 with all plasticity')
plt.plot(conn2013_NoIP,label = 'SORN 2013 without IP')
plt.plot(conn2013_NoSN,label = 'SORN 2013 without SN')
plt.plot(conn2013_NoiSTDP,label = 'SORN 2013 without iSTDP')
plt.legend()
plt.xlabel('Time steps')
plt.ylabel('Number of active connections')
plt.savefig('connection_dynamics.png')
plt.show()
```
## Firing rate of entire network
#### Spikes per time step: Should be unimodel as in paper 1
```
## READ ALL THE SIMULATION MATRICES
with open('stdp2009_50k_all.pkl','rb') as f:
plastic_matrices,X_all1,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2009_50k_NoIP.pkl','rb') as f:
plastic_matrices,X_all2,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2009_50k_NoSN.pkl','rb') as f:
plastic_matrices,X_all3,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2013_50k_all.pkl','rb') as f:
plastic_matrices4,X_all4,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2013_50k_NoIP.pkl','rb') as f:
plastic_matrices,X_all5,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2013_50k_NoSN.pkl','rb') as f:
plastic_matrices,X_all6,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
with open('stdp2013_50k_NoiSTDP.pkl','rb') as f:
plastic_matrices,X_all7,Y_all,R_all,frac_pos_active_conn5 = pickle.load(f)
spike_train1 = np.asarray(X_all1)
spike_train2 = np.asarray(X_all2)
spike_train3 = np.asarray(X_all3)
spike_train4 = np.asarray(X_all4)
spike_train5 = np.asarray(X_all5)
spike_train6 = np.asarray(X_all6)
spike_train7 = np.asarray(X_all7)
# X - axis; Neurons
# Y - axis; time_steps
def firing_rate_network(spike_train):
firing_rates = np.count_nonzero(spike_train,1)
# Filter zero entries in firing rate list above
firing_rates = list(filter(lambda a: a != 0, firing_rates))
return firing_rates
fr1 = firing_rate_network(spike_train1[8000:10000])
fr2 = firing_rate_network(spike_train2[8000:10000])
fr3 = firing_rate_network(spike_train3[8000:10000])
fr4 = firing_rate_network(spike_train4[8000:10000])
fr5 = firing_rate_network(spike_train5[8000:10000])
fr6 = firing_rate_network(spike_train6[8000:10000])
fr7 = firing_rate_network(spike_train7[8000:10000])
plt.figure(figsize = (12,8))
# plt.title('Distribution of population activity without inactive timesteps')
plt.xlabel('Spikes/timestep')
plt.ylabel('Count')
plt.hist(fr1,10,histtype = 'stepfilled',label = 'SORN 2009 with all plasticity active')
plt.hist(fr2,10,histtype = 'stepfilled',label = 'SORN 2009 without IP')
plt.hist(fr3,10,histtype = 'stepfilled',label = 'SORN 2009 without SN')
plt.hist(fr4,10,histtype = 'stepfilled',label = 'SORN 2013 with all plasticity active')
plt.hist(fr5,10,histtype = 'stepfilled',label = 'SORN 2013 without IP ')
plt.hist(fr6,10,histtype = 'stepfilled',label = 'SORN 2013 without SN')
plt.hist(fr7,10,histtype = 'stepfilled',label = 'SORN 2013 without iSTDP')
plt.legend()
plt.savefig('Spikespertimestep.png')
plt.show()
### Stacked Firing rates
import seaborn as sns
# plot
sns.set()
plt.figure(figsize = (10,6))
plt.hist([fr1,fr2,fr3,fr4,fr5,fr6,fr7],bins=10, stacked=True)
plt.legend(['SORN 2009 with all plasticity','SORN 2009 without IP','SORN 2009 without SN','SORN 2013 with all plasticity',
'SORN 2013 without IP','SORN 2013 without SN','SORN 2013 without iSTDP'],
ncol=1, loc='upper right')
plt.xlabel('Spikes/timestep')
plt.tight_layout()
plt.savefig('FiringcountStackedHist')
plt.show()
df = pd.DataFrame({'1':pd.Series(fr1),'2':pd.Series(fr2),'3':pd.Series(fr3),'4':pd.Series(fr4),'5':pd.Series(fr5),
'6':pd.Series(fr6),'7':pd.Series(fr7)})
plt.figure(figsize = (10,8))
for col in '1234567':
sns.kdeplot(df[col],shade= True)
plt.legend(['SORN 2009 with all plasticity','SORN 2009 without IP','SORN 2009 without SN','SORN 2013 with all plasticity',
'SORN 2013 without IP','SORN 2013 without SN','SORN 2013 without iSTDP'],
ncol=1, loc='upper right')
plt.xlabel('Spikes/Time step')
plt.ylabel('Density')
plt.savefig('HistFiringcount')
```
## Distribution of Firing rate: Paper 1
```
def firing_rate_neuron(X_all,neuron,time_step):
""" Measure spike rate of given neuron during given time window"""
time_period = len(X_all[:,0])
neuron_spike_train = X_all[:,neuron]
# Split the list(neuron_spike_train) into sublists of lenth time_step
samples_spike_train = [neuron_spike_train[i:i+time_step] for i in range(0, len(neuron_spike_train), time_step)]
spike_rate = 0.
for idx,spike_train in enumerate(samples_spike_train):
spike_rate += list(spike_train).count(1.)
spike_rate = spike_rate*time_step/time_period
# print('Firing rate of neuron %s in %s time steps is %s' %(neuron,time_step,spike_rate/time_step))
return time_period, time_step,spike_rate
_,_,spike_rate=firing_rate_neuron(spike_train1[0:10000],60,10) # Neuron 60; time window size 10 # As per target firing rate
spike_rate
def firing_rate_h(spike_train):
frs = []
for i in range(200):
_,_,fr = firing_rate_neuron(spike_train[0:10000],i,10)
frs.append(fr/10)
return frs
frs1 = firing_rate_h(spike_train1)
frs2 = firing_rate_h(spike_train2)
frs3 = firing_rate_h(spike_train3)
frs4 = firing_rate_h(spike_train4)
frs5 = firing_rate_h(spike_train5)
frs6 = firing_rate_h(spike_train6)
frs7 = firing_rate_h(spike_train7)
plt.figure(figsize=(12,8))
# plt.title('Distribution of firing rates')
plt.xlabel('Rate H')
plt.ylabel('Number of units')
plt.hist(frs1,5,histtype = 'stepfilled',label = 'SORN 2009 with all plasticity')
plt.hist(frs2,5,histtype = 'stepfilled',label = 'SORN 2009 without IP')
plt.hist(frs3,5,histtype = 'stepfilled',label = 'SORN 2009 without SN')
plt.hist(frs4,5,histtype = 'stepfilled',label = 'SORN 2013 with all plasticity')
plt.hist(frs5,5,histtype = 'stepfilled',label = 'SORN 2013 without IP')
plt.hist(frs6,5,histtype = 'stepfilled',label = 'SORN 2013 without SN')
plt.hist(frs7,5,histtype = 'stepfilled',label = 'SORN 2013 without iSTDP')
plt.legend()
plt.savefig('FiringRateH.png')
plt.show()
df = pd.DataFrame({'1':frs1,'2':frs2,'3':frs3,'4':frs4,'5':frs5,'6':frs6,'7':frs7})
import seaborn as sns
# plot
sns.set()
plt.figure(figsize = (10,6))
plt.hist([frs1,frs2,frs3,frs4,frs5,frs6,frs7],bins=10, stacked=True)
plt.legend(['SORN 2009 with all plasticity','SORN 2009 without IP','SORN 2009 without SN','SORN 2013 with all plasticity',
'SORN 2013 without IP','SORN 2013 without SN','SORN 2013 without iSTDP'],
ncol=1, loc='upper right')
plt.xlabel('Rate H')
plt.tight_layout()
plt.savefig('FiringrateStackedHist')
plt.show()
```
### Plot Spike train
```
# Get the indices where spike_train is 1
x,y = np.argwhere(spike_train1[9800:10000].T == 1).T
def _scatter_plot(x,y,firing_rates):
plt.figure(figsize = (8,5))
plt.scatter(y,x, s= 0.1,color = 'black')
# plt.plot(y,x,'|b')
# plt.gca().invert_yaxis()
plt.plot(firing_rates,label = 'Firing rate')
plt.xlabel('Time(ms)')
plt.ylabel('Neuron #')
plt.legend(loc = 'upper right')
plt.savefig('SpikeTrain.png')
plt.show()
def _raster_plot(x,y,firing_rates):
plt.figure(figsize = (11,6))
plt.plot(y,x,'|r')
plt.plot(firing_rates,label = 'Firing rate')
# plt.gca().invert_yaxis()
plt.xlabel('Time(ms)')
plt.ylabel('Neuron #')
plt.legend(loc = 'upper right')
plt.savefig('SpikeTrain.png')
plt.show()
fr = firing_rate_network(spike_train7[9800:10000])
_scatter_plot(x,y,fr)
# # _raster_plot(x,y,fr)
```
## Smoothness firing rate time series
#### Scale dependent method
Smaller values corresponds to smoother series
```
def scale_dependent_smoothness_measure(firing_rates):
"""
Args:
firing_rates - List of number of active neurons per time step
Returns:
sd_diff - Float value signifies the smoothness of the sementic changes in firing rates
"""
diff = np.diff(firing_rates)
sd_diff = np.std(diff)
return sd_diff
```
#### Scale independent measure
Smaller values corresponds to smoother series
```
def scale_independent_smoothness_measure(firing_rates):
"""
Args:
firing_rates - List of number of active neurons per time step
Returns:
coeff_var - Float value signifies the smoothness of the sementic changes in firing rates
"""
diff = np.diff(firing_rates)
mean_diff = np.mean(diff)
sd_diff = np.std(diff)
coeff_var = sd_diff/abs(mean_diff)
return coeff_var
```
#### Using one-lag autocorrelation measure
```
def autocorr(fr,t=2):
return np.corrcoef(np.array([fr[0:len(fr)-t], fr[t:len(fr)]]))
print('smoothness measure using scale_dependent_smoothness_measure',scale_dependent_smoothness_measure(fr[0:200]))
print('smoothness measure using scale_independent_smoothness_measure',scale_independent_smoothness_measure(fr[0:200]))
print('smoothness measure using one-lag auto correlation \n',autocorr(fr[0:200]))
```
## Average Correlation coefficient between neurons
```
def avg_corr_coeff(spike_train):
"""Measure Average Pearson correlation coeffecient between neurons"""
corr_mat = np.corrcoef(np.asarray(spike_train).T)
avg_corr = np.sum(corr_mat,axis = 1) / 200
corr_coeff = avg_corr.sum()/200/2 ; # 2D to 1D and either upper or lower half of correlation matrix.
print('Average correlation coeffecient between neurons in excitatory pool is', corr_coeff)
return corr_mat,corr_coeff
# PLOT THE CORRELATION BETWEEN EXCITATORY NEURONS
def plot_correlation(corr):
""" Plot correlation between neurons"""
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
f, ax = plt.subplots(figsize=(11, 9))
# Custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
return sns.heatmap(corr, mask=mask, cmap=cmap,xticklabels=5, yticklabels=5, vmax=.1, center=0,
square=False, linewidths=0.0, cbar_kws={"shrink": .9})
corr, avg_corr = avg_corr_coeff(spike_train1[0:10000])
plot_correlation(corr)
spike_train = spike_train4.copy()
```
## Exponential fit of Spike time interval
### Exponential fit used first 10000 time steps of network activity
### At 100000 times steps network shows log normal pattern in activity
```
# with open('stdp2009_50k.pkl','rb') as f:
# plastic_matrices,X_all,Y_all,R_all,frac_pos_active_conn = pickle.load(f)
# spike_train = np.asarray(X_all)
def spike_times(x):
""" Get the time instants at which neuron spikes"""
times = np.where(x == 1.)
return times
def spike_time_intervals(spike_times):
""" Generate spike time intervals|spike_times"""
# isi = sorted(np.diff(spike_times)[-1])
isi = np.diff(spike_times)[-1]
return isi
# Define required parameters
y_all,x_all = [],[]
for i in range(200):
spike_time = spike_times(spike_train[0:20000].T[i]) # Locate the spike time of neuron 60 from its spike train
isi = spike_time_intervals(spike_time) # ISI intervals of neuron
bin_size = 19
y, x = np.histogram(sorted(isi),bins = bin_size)
y_all.extend(y)
x_all.extend(x)
len(y_all[1:bin_size])
len(x_all[1:bin_size])
def isi_exponential_fit(x,y,bin_size):
x = [int(i) for i in x]
y = [float(i) for i in y]
def exponential_func(y, a, b, c):
return a*np.exp(-b* np.array(y)) - c
# Curve fit
popt, pcov = curve_fit(exponential_func, x[1:bin_size], y[1:bin_size])
# Plot
plt.plot(x[1:bin_size], exponential_func(x[1:bin_size], *popt),color = 'orange', label = 'Exponential fit')
# plt.scatter(x[1:bin_size],y[1:bin_size], s= 2.0,color = 'black',label = 'ISI')
sns.scatterplot(x[1:bin_size],y[1:bin_size],color = 'green',size = y[1:bin_size],hue = y[1:bin_size])
plt.xlabel('ISI(time step)')
plt.ylabel('Frequency')
plt.legend()
plt.savefig('ISI')
plt.show()
isi_exponential_fit(x,y,bin_size)
df = pd.DataFrame({'x':x[1:19],'y':y[1:19]},columns=['x','y'])
df.size
import seaborn as sns
h = sns.jointplot(x="x", y="y", data=df, kind="kde")
# or set labels via the axes objects
h.ax_joint.set_xlabel('ISI(Time steps)')
h.ax_joint.set_ylabel('Frequency')
plt.tight_layout()
plt.savefig('ISIDensity')
plt.show()
```
### Ditribution of weights using histogram plot
```
weight_strengths = plastic_matrices['Wee']
weights = np.array(weight_strengths.tolist())
weights = weights[weights >= 0.01] # Remove the weight values less than 0.01 # As reported in article SORN 2013
y,x = np.histogram(weights, bins = 100) # Create histogram and the bin size 100
h = sns.scatterplot(x[:-1],y,color = 'orange',size = y,hue = y)
# or set labels via the axes objects
# h.ax_joint.set_xlabel('Connection strength')
# h.ax_joint.set_ylabel('Frequency')
plt.xlabel('Connection strength')
plt.ylabel('Frequency')
plt.tight_layout()
plt.savefig('HistWeights')
plt.show()
import seaborn as sns
cmap = sns.cubehelix_palette(as_cmap=True, dark=0, light=1, reverse=True)
h = sns.jointplot(x="x", y="y",cmap=cmap, data={'x':x[:-1],'y':y}, kind="kde")
# or set labels via the axes objects
h.ax_joint.set_xlabel('Connection strength')
h.ax_joint.set_ylabel('Frequency')
plt.tight_layout()
plt.savefig('WeightsDensity')
plt.show()
```
## Linear and lognormal fit of weight distribution
###### Reference: # http://nbviewer.jupyter.org/url/xweb.geos.ed.ac.uk/~jsteven5/blog/lognormal_distributions.ipynb
```
weight_strengths = plastic_matrices4['Wee']
weights = np.array(weight_strengths.tolist())
weights = weights[weights >= 0.01] # Remove the weight values less than 0.01 # As reported in article SORN 2
y,x = np.histogram(weights, bins = 50) # Create histogram and the bin size 10 which captures a avg firing rate 0.1
# plt.scatter(x[:-1],y,s = 1.0,c ='black')
h = sns.scatterplot(x[:-1],y,color = 'orange',size = y,hue = y,cmap = cmap)
plt.xlabel('Connection strength')
plt.ylabel('Count')
plt.savefig('Connection strenths Sca')
plt.show()
import seaborn as sns
cmap = sns.cubehelix_palette(as_cmap=True, dark=0, light=1, reverse=True)
# h = sns.jointplot(x="x", y="y",cmap=cmap, data={'x':x[:-1],'y':y}, kind="kde")
f, ax = plt.subplots(figsize=(6, 6))
sns.kdeplot(x[:-1], y,cmap=cmap,shade= True, ax=ax)
sns.rugplot(x[:-1], color="g", ax=ax)
sns.rugplot(y, vertical=True, ax=ax);
plt.xlabel('Connection strength')
plt.ylabel('Count')
# or set labels via the axes objects
# h.ax_joint.set_xlabel('Connection strength')
# h.ax_joint.set_ylabel('Frequency')
plt.tight_layout()
plt.savefig('NumberWeightsDensity')
plt.show()
X= weights.copy()
M = float(np.mean(X)) # Geometric mean
s = float(np.std(X)) # Geometric standard deviation
# Lognormal distribution parameters
mu = float(np.mean(np.log(X))) # Mean of log(X)
sigma = float(np.std(np.log(X))) # Standard deviation of log(X)
shape = sigma # Scipy's shape parameter
scale = np.exp(mu) # Scipy's scale parameter
median = np.exp(mu)
mode = np.exp(mu - sigma**2) # Note that mode depends on both M and s
mean = np.exp(mu + (sigma**2/2)) # Note that mean depends on both M and s
x = np.linspace(min(weights),max(weights) , num=100) # values for x-axis
pdf = stats.lognorm.pdf(x, shape, loc=0, scale=scale) # probability distribution
plt.figure(figsize=(12,4.5))
# Figure on linear scale
plt.subplot(121)
plt.plot(x, pdf)
plt.vlines(mode, 0, pdf.max(), linestyle=':', label='Mode')
plt.vlines(mean, 0, stats.lognorm.pdf(mean, shape, loc=0, scale=scale), linestyle='--', color='green', label='Mean')
plt.vlines(median, 0, stats.lognorm.pdf(median, shape, loc=0, scale=scale), color='blue', label='Median')
plt.ylim(ymin=0)
plt.xlabel('Weight')
plt.ylabel('Density')
plt.title('Linear scale')
leg=plt.legend()
# Figure on logarithmic scale
plt.subplot(122)
plt.semilogx(x, pdf)
plt.vlines(mode, 0, pdf.max(), linestyle=':', label='Mode')
plt.vlines(mean, 0, stats.lognorm.pdf(mean, shape, loc=0, scale=scale), linestyle='--', color='green', label='Mean')
plt.vlines(median, 0, stats.lognorm.pdf(median, shape, loc=0, scale=scale), color='blue', label='Median')
plt.ylim(ymin=0)
plt.xlabel('Weight')
plt.title('Logarithmic scale')
plt.savefig('WeightsLogscale55timesteps')
leg=plt.legend()
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
total = pd.read_csv('us.csv', header=0, sep=',')
total.head(10)
total = total.iloc[:224, :]
fig = plt.figure(figsize = (10,6))
plt.bar(total.date,
total.cases,
label = 'cases'
)
plt.plot(total.date,
total.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
ax = plt.gca()
for label in ax.get_xticklabels():
label.set_visible(False)
for label in ax.get_xticklabels()[::20]:
label.set_visible(True)
plt.title('total cases and deaths')
plt.xlabel('date')
plt.ylabel('number of people')
fig.autofmt_xdate(rotation = 45)
plt.legend()
plt.show()
new_increase = pd.DataFrame()
new_increase['date'] = total['date']
new_increase_cases = total['cases'].copy()
new_increase_cases.values[1:] -= new_increase_cases.values[:-1]
new_increase['cases'] = new_increase_cases
new_increase_deaths = total['deaths'].copy()
new_increase_deaths.values[1:] -= new_increase_deaths.values[:-1]
new_increase['deaths'] = new_increase_deaths
new_increase.head(10)
fig = plt.figure(figsize = (10,6))
plt.bar(new_increase.date,
new_increase.cases,
label = 'cases'
)
plt.plot(new_increase.date,
new_increase.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
ax = plt.gca()
for label in ax.get_xticklabels():
label.set_visible(False)
for label in ax.get_xticklabels()[::20]:
label.set_visible(True)
plt.title('daily increased cases and deaths')
plt.xlabel('date')
plt.ylabel('number of people')
fig.autofmt_xdate(rotation = 45)
plt.legend()
plt.show()
# 7-Day Moving Average
average7 = pd.DataFrame()
average7['date'] = new_increase['date']
N = 7
n = np.ones(N)
weights = n/N
new_increase_cases = new_increase['cases'].values
average7_cases = pd.Series(np.convolve(weights, new_increase_cases)[:new_increase_cases.shape[0]])
average7['cases'] = average7_cases
new_increase_deaths = new_increase['deaths'].values
average7_deaths = pd.Series(np.convolve(weights, new_increase_deaths)[:new_increase_deaths.shape[0]])
average7['deaths'] = average7_deaths
average7.head(10)
fig = plt.figure(figsize = (10,6))
plt.bar(average7.date,
average7.cases,
label = 'cases'
)
plt.plot(average7.date,
average7.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
ax = plt.gca()
for label in ax.get_xticklabels():
label.set_visible(False)
for label in ax.get_xticklabels()[::20]:
label.set_visible(True)
plt.title('7 days average of daily new cases and deaths')
plt.xlabel('date')
plt.ylabel('number of people')
fig.autofmt_xdate(rotation = 45)
plt.legend()
plt.show()
# weekly average
average_week = pd.DataFrame()
average_week_date = np.empty([33], dtype='S10')
average_week_date[1:] = new_increase['date'].values[5::7]
average_week_date[0] = new_increase['date'].values[0]
average_week['start_date'] = pd.Series(average_week_date)
new_increase_cases = new_increase['cases'].values
average_week_cases = np.zeros(33)
average_week_cases[0] = np.mean(new_increase_cases[:5])
for i in range(31):
average_week_cases[i+1] = np.mean(new_increase_cases[i*7+5:i*7+12])
average_week_cases[32] = np.mean(new_increase_cases[31*7+5:31*7+7])
average_week['cases'] = pd.Series(average_week_cases)
new_increase_deaths = new_increase['deaths'].values
average_week_deaths = np.zeros(33)
average_week_deaths[0] = np.mean(new_increase_deaths[:5])
for i in range(31):
average_week_deaths[i+1] = np.mean(new_increase_deaths[i*7+5:i*7+12])
average_week_deaths[32] = np.mean(new_increase_deaths[31*7+5:31*7+7])
average_week['deaths'] = pd.Series(average_week_deaths)
average_week.head(10)
fig = plt.figure(figsize = (10,6))
plt.bar(average_week.start_date,
average_week.cases,
label = 'cases'
)
plt.plot(average_week.start_date,
average_week.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
ax = plt.gca()
for label in ax.get_xticklabels():
label.set_visible(False)
for label in ax.get_xticklabels()[::1]:
label.set_visible(True)
plt.title('weekly average of daily new cases and deaths')
plt.xlabel('week_start_date')
plt.ylabel('number of people')
fig.autofmt_xdate(rotation = 45)
plt.legend()
plt.show()
# monthly average
average_month = pd.DataFrame()
average_month_name = np.empty([8], dtype='S10')
average_month_name[0] = 'Jan.'
average_month_name[1] = 'Feb.'
average_month_name[2] = 'Mar.'
average_month_name[3] = 'Apr.'
average_month_name[4] = 'May'
average_month_name[5] = 'June'
average_month_name[6] = 'July'
average_month_name[7] = 'Aug.'
average_month['month'] = pd.Series(average_month_name)
new_increase_cases = new_increase['cases'].values
average_month_cases = np.zeros(8)
average_month_cases[0] = np.mean(new_increase_cases[:11])
average_month_cases[1] = np.mean(new_increase_cases[11:40])
average_month_cases[2] = np.mean(new_increase_cases[40:71])
average_month_cases[3] = np.mean(new_increase_cases[71:101])
average_month_cases[4] = np.mean(new_increase_cases[101:132])
average_month_cases[5] = np.mean(new_increase_cases[132:162])
average_month_cases[6] = np.mean(new_increase_cases[162:193])
average_month_cases[7] = np.mean(new_increase_cases[193:224])
average_month['cases'] = pd.Series(average_month_cases)
new_increase_deaths = new_increase['deaths'].values
average_month_deaths = np.zeros(8)
average_month_deaths[0] = np.sum(new_increase_deaths[:11]) / 31
average_month_deaths[1] = np.mean(new_increase_deaths[11:40])
average_month_deaths[2] = np.mean(new_increase_deaths[40:71])
average_month_deaths[3] = np.mean(new_increase_deaths[71:101])
average_month_deaths[4] = np.mean(new_increase_deaths[101:132])
average_month_deaths[5] = np.mean(new_increase_deaths[132:162])
average_month_deaths[6] = np.mean(new_increase_deaths[162:193])
average_month_deaths[7] = np.mean(new_increase_deaths[193:224])
average_month['deaths'] = pd.Series(average_month_deaths)
average_month.head(10)
fig = plt.figure(figsize = (10,6))
plt.bar(average_month.month,
average_month.cases,
label = 'cases'
)
plt.plot(average_month.month,
average_month.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
ax = plt.gca()
for label in ax.get_xticklabels():
label.set_visible(False)
for label in ax.get_xticklabels()[::1]:
label.set_visible(True)
plt.title('monthly average of daily new cases and deaths')
plt.xlabel('month')
plt.ylabel('number of people')
fig.autofmt_xdate(rotation = 45)
plt.legend()
plt.show()
new_increase.sort_values(by='cases', ascending=False).head(10)[['date', 'cases']]
new_increase.sort_values(by='cases', ascending=True).head(10)[['date', 'cases']]
new_increase.sort_values(by='deaths', ascending=False).head(10)[['date', 'deaths']]
new_increase.sort_values(by='deaths', ascending=True).head(10)[['date', 'deaths']]
most_cases_week = average_week.sort_values(by='cases', ascending=False).head(5)[['start_date', 'cases']]
most_cases_week['cases'] = (most_cases_week['cases'].values * 7).astype(int)
most_cases_week
most_deaths_week = average_week.sort_values(by='deaths', ascending=False).head(5)[['start_date', 'deaths']]
most_deaths_week['deaths'] = (most_deaths_week['deaths'].values * 7).astype(int)
most_deaths_week
fig, axs = plt.subplots(4, 1, constrained_layout=False, figsize = (10,24))
plt.subplots_adjust(hspace = .5)
axs[0].bar(total.date,
total.cases,
label = 'cases'
)
axs[0].plot(total.date,
total.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
for label in axs[0].get_xticklabels():
label.set_visible(False)
for label in axs[0].get_xticklabels()[::7]:
label.set_visible(True)
label.set_rotation(45)
axs[0].set_title('total cases and deaths')
axs[0].set_xlabel('date')
axs[0].set_ylabel('number of people')
axs[0].legend()
axs[1].bar(new_increase.date,
new_increase.cases,
label = 'cases'
)
axs[1].plot(new_increase.date,
new_increase.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
for label in axs[1].get_xticklabels():
label.set_visible(False)
for label in axs[1].get_xticklabels()[::7]:
label.set_visible(True)
label.set_rotation(45)
axs[1].set_title('daily increased cases and deaths')
axs[1].set_xlabel('date')
axs[1].set_ylabel('number of people')
axs[1].legend()
axs[2].bar(average7.date,
average7.cases,
label = 'cases'
)
axs[2].plot(average7.date,
average7.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
for label in axs[2].get_xticklabels():
label.set_visible(False)
for label in axs[2].get_xticklabels()[::7]:
label.set_visible(True)
label.set_rotation(45)
axs[2].set_title('7 days average of daily new cases and deaths')
axs[2].set_xlabel('date')
axs[2].set_ylabel('number of people')
axs[2].legend()
axs[3].bar(average_month.month,
average_month.cases,
label = 'cases'
)
axs[3].plot(average_month.month,
average_month.deaths,
linestyle = '-',
color='red',
label = 'deaths'
)
for label in axs[3].get_xticklabels():
label.set_rotation(45)
axs[3].set_title('monthly average of daily new cases and deaths')
axs[3].set_xlabel('month')
axs[3].set_ylabel('number of people')
axs[3].legend()
plt.show()
```
| github_jupyter |
```
import qiskit
from qiskit import *
from qiskit.tools.visualization import *
from qiskit.tools.monitor import job_monitor
%matplotlib inline
with open('tocken.txt', 'r') as file:
myTocken = file.read().replace('\n', '')
IBMQ.save_account(myTocken,overwrite=True)
IBMQ.load_account()
```
Build a quantum circuit of 2 qubits by building 2 bit quantum register
Build a classical circuit of 2 bits by building 2 bit classical register
classical registers here are essential to store measurements of the corresponding quantum bits
now we can build a circuit of them both
vizualize the circuit, with braket notations
```
circuit.draw()
circuit.draw(output='mpl')
```
Next stage is to build the gates that can make use of this circuit
demonstrate quantum entanglement, Hadamard (H) gate
Apply Hadamard (H) gate on the first Qbit
```
circuit.draw(output='mpl')
```
now, making a 2 qubit operation called 'controlled X' which can map to logical 'if' condition, so it can be the quantum version of 'if this then that'
control of this operation is 1st qubit
target of this operation is 2nd qubit
make a 2 qubit operation, controlled NOT, logical if operation, control is the first qbit, target of the operation is the second qbit
```
circuit.draw(output='mpl')
```
now there is is entanglement
Now, measure the qbits and store them in classical bits
```
circuit.measure(qr, cr)
circuit.draw(output='mpl')
circuit.draw()
```
Now, run the circuit in a simulator
gonna use qasm simulator: quantum assembly language
now call the simulator
```
simulator = Aer.get_backend('qasm_simulator')
```
execute the circuit
```
execute(circuit, backend=simulator,shots=10000)
result = execute(circuit, backend=simulator).result()
circuit.draw(output='mpl')
plot_histogram(result.get_counts(circuit))
```
erros because we are running limited number of shots on simulation instead of infinite number of shots
Load the account to run on IBMQ
```
provider = IBMQ.get_provider('ibm-q')
qcomp = provider.get_backend('Place-your-prefered-quantum-computer')
job = execute(circuit, backend=qcomp)
```
check the job status
```
job_monitor(job)
resultFromQC = job.result()
plot_histogram(resultFromQC.get_counts(circuit))
```
in simulated, only 00, 11 but on real quantum device, we have all
the diff is that simulator simulates perfect, but the real device is siumulating the real quantum errors
01, 10
the simulator simulates a perfect quntum device
the real quantum device takes into consideration 'small quantum errors' which result in getting those 01, 10
| github_jupyter |
<a href='http://www.holoviews.org'><img src="assets/hv+bk.png" alt="HV+BK logos" width="40%;" align="left"/></a>
<div style="float:right;"><h2>11. Deploying Bokeh Apps</h2></div>
In the previous sections we discovered how to use a ``HoloMap`` to build a Jupyter notebook with interactive visualizations that can be exported to a standalone HTML file, as well as how to use ``DynamicMap`` and ``Streams`` to set up dynamic interactivity backed by the Jupyter Python kernel. However, frequently we want to package our visualization or dashboard for wider distribution, backed by Python but run outside of the notebook environment. Bokeh Server provides a flexible and scalable architecture to deploy complex interactive visualizations and dashboards, integrating seamlessly with Bokeh and with HoloViews.
For a detailed background on Bokeh Server see [the Bokeh user guide](http://bokeh.pydata.org/en/latest/docs/user_guide/server.html). In this tutorial we will discover how to deploy the visualizations we have created so far as a standalone Bokeh Server app, and how to flexibly combine HoloViews and ParamBokeh to build complex apps. We will also reuse a lot of what we have learned so far---loading large, tabular datasets, applying Datashader operations to them, and adding linked Streams to our app.
## A simple Bokeh app
The preceding sections of this tutorial focused solely on the Jupyter notebook, but now let's look at a bare Python script that can be deployed using Bokeh Server:
```
with open('./apps/server_app.py', 'r') as f:
print(f.read())
```
Step 1 of this app should be very familiar by now -- declare that we are using Bokeh to render plots, load some taxi dropoff locations, declare a Points object, Datashade them, and set some plot options.
At this point, if we were working with this code in a notebook, we would simply type ``shaded`` and let Jupyter's rich display support take over, rendering the object into a Bokeh plot and displaying it inline. Here, step 2 adds the code necessary to do those steps explicitly:
- get a handle on the Bokeh renderer object using ``hv.renderer``
- create a Bokeh document from ``shaded`` by passing it to the renderer's ``server_doc`` method
- optionally, change some properties of the Bokeh document like the title.
This simple chunk of boilerplate code can be added to turn any HoloViews object into a fully functional, deployable Bokeh app!
## Deploying the app
Assuming that you have a terminal window open with the ``hvtutorial`` environment activated, in the ``notebooks/`` directory, you can launch this app using Bokeh Server:
```
bokeh serve --show apps/server_app.py
```
If you don't already have a favorite way to get a terminal, one way is to [open it from within Jupyter](../terminals/1), then make sure you are in the ``notebooks`` directory, and activate the environment using ``source activate hvtutorial`` (or ``activate tutorial`` on Windows). You can also [open the app script file](../edit/apps/server_app.py) in the inbuilt text editor, or you can use your own preferred editor.
```
# Exercise: Modify the app to display the pickup locations and add a tilesource, then run the app with bokeh serve
# Tip: Refer to the previous notebook
```
## Building an app with custom widgets
The above app script can be built entirely without using Jupyter, though we displayed it here using Jupyter for convenience in the tutorial. Jupyter notebooks are also often helpful when initially developing such apps, allowing you to quickly iterate over visualizations in the notebook, deploying it as a standalone app only once we are happy with it. In this section we will combine everything we have learned so far including declaring of various parameters to control our visualization using a set of widgets.
We begin as usual with a set of imports:
```
import holoviews as hv, geoviews as gv, param, parambokeh, dask.dataframe as dd
from colorcet import cm
from bokeh.models import WMTSTileSource
from holoviews.operation.datashader import datashade
from holoviews.streams import RangeXY
hv.extension('bokeh', logo=False)
```
Next we once again load the Taxi dataset and define a tile source:
```
usecols = ['dropoff_x', 'dropoff_y', 'pickup_x', 'pickup_y', 'dropoff_hour']
df = dd.read_parquet('../data/nyc_taxi_hours.parq/')
df = df[usecols].persist()
url='https://server.arcgisonline.com/ArcGIS/rest/services/World_Imagery/MapServer/tile/{Z}/{Y}/{X}.jpg'
tiles = gv.WMTS(WMTSTileSource(url=url))
tile_options = dict(width=600,height=400,xaxis=None,yaxis=None,bgcolor='black',show_grid=False)
```
Finally we will put together a complete dashboard with a number of parameters controlling our visualization, including controls over the alpha level of the tiles and the colormap as well as the hour of day and whether to plot dropoff or pickup location.
```
class NYCTaxiExplorer(hv.streams.Stream):
alpha = param.Magnitude(default=0.75, doc="Alpha value for the map opacity")
colormap = param.ObjectSelector(default=cm["fire"], objects=[cm[k] for k in cm.keys() if not '_' in k])
hour = param.Range(default=(0, 24), bounds=(0, 24))
location = param.ObjectSelector(default='dropoff', objects=['dropoff', 'pickup'])
def make_view(self, x_range, y_range, **kwargs):
map_tiles = tiles.opts(style=dict(alpha=self.alpha), plot=tile_options)
points = hv.Points(df, kdims=[self.location+'_x', self.location+'_y'], vdims=['dropoff_hour'])
if self.hour != (0, 24): points = points.select(dropoff_hour=self.hour)
taxi_trips = datashade(points, x_sampling=1, y_sampling=1, cmap=self.colormap,
dynamic=False, x_range=x_range, y_range=y_range, width=600, height=400)
return map_tiles * taxi_trips
explorer = NYCTaxiExplorer(name="NYC Taxi Trips")
dmap = hv.DynamicMap(explorer.make_view, streams=[explorer, RangeXY()])
plot = hv.renderer('bokeh').get_plot(dmap)
parambokeh.Widgets(explorer, view_position='right', callback=explorer.event, plots=[plot.state])
```
Now let's open the [text editor](../edit/apps/nyc_taxi/main.py) again and make this edit to a separate app, which we can then launch using Bokeh Server from the [terminal](../terminals/1).
```
# Exercise: Note the differences between the server app and the app defined above
# then add an additional parameter and plot
# Exercise: Click the link below and edit the Jinja2 template to customize the app
```
[Edit the template](../edit/apps/nyc_taxi/templates/index.html)
## Combining HoloViews with bokeh models
Now for a last hurrah let's put everything we have learned to good use and create a bokeh app with it. This time we will go straight to a [Python script containing the app](../edit/apps/player_app.py). If you run the app with ``bokeh serve --show ./apps/player_app.py`` from [your terminal](../terminals/1) you should see something like this:
<img src="./assets/tutorial_app.gif"></img>
This more complex app consists of several components:
1. A datashaded plot of points for the indicated hour of the daty (in the slider widget)
2. A linked ``PointerX`` stream, to compute a cross-section
3. A set of custom Bokeh widgets linked to the hour-of-day stream
We have already covered 1. and 2. so we will focus on 3., which shows how easily we can combine a HoloViews plot with custom Bokeh models. We will not look at the precise widgets in too much detail, instead let's have a quick look at the callback defined for slider widget updates:
```python
def slider_update(attrname, old, new):
stream.event(hour=new)
```
Whenever the slider value changes this will trigger a stream event updating our plots. The second part is how we combine HoloViews objects and Bokeh models into a single layout we can display. Once again we can use the renderer to convert the HoloViews object into something we can display with Bokeh:
```python
renderer = hv.renderer('bokeh')
plot = renderer.get_plot(hvobj, doc=curdoc())
```
The ``plot`` instance here has a ``state`` attribute that represents the actual Bokeh model, which means we can combine it into a Bokeh layout just like any other Bokeh model:
```python
layout = layout([[plot.state], [slider, button]], sizing_mode='fixed')
curdoc().add_root(layout)
```
```
# Advanced Exercise: Add a histogram to the bokeh layout next to the datashaded plot
# Hint: Declare the histogram like this: hv.operation.histogram(aggregated, bin_range=(0, 20))
# then use renderer.get_plot and hist_plot.state and add it to the layout
```
# Onwards
Although the code above is more complex than in previous sections, it's providing a huge range of custom types of interactivity, which if implemented in Bokeh alone would have required far more than a notebook cell of code. Hopefully it is clear that arbitrarily complex collections of visualizations and interactive controls can be built from the components provided by HoloViews, allowing you to make simple analyses very easily and making it practical to make even quite complex apps when needed. The [user guide](http://holoviews.org/user_guide), [gallery](http://holoviews.org/gallery/index.html), and [reference gallery](http://holoviews.org/reference) should have all the information you need to get started with all this power on your own datasets and tasks. Good luck!
| github_jupyter |
# Name
Data preparation using Spark on YARN with Cloud Dataproc
# Label
Cloud Dataproc, GCP, Cloud Storage, Spark, Kubeflow, pipelines, components, YARN
# Summary
A Kubeflow Pipeline component to prepare data by submitting a Spark job on YARN to Cloud Dataproc.
# Details
## Intended use
Use the component to run an Apache Spark job as one preprocessing step in a Kubeflow Pipeline.
## Runtime arguments
Argument | Description | Optional | Data type | Accepted values | Default |
:--- | :---------- | :--- | :------- | :------| :------|
project_id | The ID of the Google Cloud Platform (GCP) project that the cluster belongs to.|No | GCPProjectID | | |
region | The Cloud Dataproc region to handle the request. | No | GCPRegion | | |
cluster_name | The name of the cluster to run the job. | No | String | | |
main_jar_file_uri | The Hadoop Compatible Filesystem (HCFS) URI of the JAR file that contains the main class. | No | GCSPath | | |
main_class | The name of the driver's main class. The JAR file that contains the class must be either in the default CLASSPATH or specified in `spark_job.jarFileUris`.| No | | | |
args | The arguments to pass to the driver. Do not include arguments, such as --conf, that can be set as job properties, since a collision may occur that causes an incorrect job submission.| Yes | | | |
spark_job | The payload of a [SparkJob](https://cloud.google.com/dataproc/docs/reference/rest/v1/SparkJob).| Yes | | | |
job | The payload of a [Dataproc job](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.jobs). | Yes | | | |
wait_interval | The number of seconds to wait between polling the operation. | Yes | | | 30 |
## Output
Name | Description | Type
:--- | :---------- | :---
job_id | The ID of the created job. | String
## Cautions & requirements
To use the component, you must:
* Set up a GCP project by following this [guide](https://cloud.google.com/dataproc/docs/guides/setup-project).
* [Create a new cluster](https://cloud.google.com/dataproc/docs/guides/create-cluster).
* The component can authenticate to GCP. Refer to [Authenticating Pipelines to GCP](https://www.kubeflow.org/docs/gke/authentication-pipelines/) for details.
* Grant the Kubeflow user service account the role `roles/dataproc.editor` on the project.
## Detailed description
This component creates a Spark job from [Dataproc submit job REST API](https://cloud.google.com/dataproc/docs/reference/rest/v1/projects.regions.jobs/submit).
Follow these steps to use the component in a pipeline:
1. Install the Kubeflow Pipeline SDK:
```
%%capture --no-stderr
!pip3 install kfp --upgrade
```
2. Load the component using KFP SDK
```
import kfp.components as comp
dataproc_submit_spark_job_op = comp.load_component_from_url(
'https://raw.githubusercontent.com/kubeflow/pipelines/1.5.0/components/gcp/dataproc/submit_spark_job/component.yaml')
help(dataproc_submit_spark_job_op)
```
### Sample
Note: The following sample code works in an IPython notebook or directly in Python code.
#### Set up a Dataproc cluster
[Create a new Dataproc cluster](https://cloud.google.com/dataproc/docs/guides/create-cluster) (or reuse an existing one) before running the sample code.
#### Prepare a Spark job
Upload your Spark JAR file to a Cloud Storage bucket. In the sample, we use a JAR file that is preinstalled in the main cluster: `file:///usr/lib/spark/examples/jars/spark-examples.jar`.
Here is the [source code of the sample](https://github.com/apache/spark/blob/master/examples/src/main/java/org/apache/spark/examples/JavaSparkPi.java).
To package a self-contained Spark application, follow these [instructions](https://spark.apache.org/docs/latest/quick-start.html#self-contained-applications).
#### Set sample parameters
```
PROJECT_ID = '<Please put your project ID here>'
CLUSTER_NAME = '<Please put your existing cluster name here>'
REGION = 'us-central1'
SPARK_FILE_URI = 'file:///usr/lib/spark/examples/jars/spark-examples.jar'
MAIN_CLASS = 'org.apache.spark.examples.SparkPi'
ARGS = ['1000']
EXPERIMENT_NAME = 'Dataproc - Submit Spark Job'
```
#### Example pipeline that uses the component
```
import kfp.dsl as dsl
import json
@dsl.pipeline(
name='Dataproc submit Spark job pipeline',
description='Dataproc submit Spark job pipeline'
)
def dataproc_submit_spark_job_pipeline(
project_id = PROJECT_ID,
region = REGION,
cluster_name = CLUSTER_NAME,
main_jar_file_uri = '',
main_class = MAIN_CLASS,
args = json.dumps(ARGS),
spark_job=json.dumps({ 'jarFileUris': [ SPARK_FILE_URI ] }),
job='{}',
wait_interval='30'
):
dataproc_submit_spark_job_op(
project_id=project_id,
region=region,
cluster_name=cluster_name,
main_jar_file_uri=main_jar_file_uri,
main_class=main_class,
args=args,
spark_job=spark_job,
job=job,
wait_interval=wait_interval)
```
#### Compile the pipeline
```
pipeline_func = dataproc_submit_spark_job_pipeline
pipeline_filename = pipeline_func.__name__ + '.zip'
import kfp.compiler as compiler
compiler.Compiler().compile(pipeline_func, pipeline_filename)
```
#### Submit the pipeline for execution
```
#Specify pipeline argument values
arguments = {}
#Get or create an experiment and submit a pipeline run
import kfp
client = kfp.Client()
experiment = client.create_experiment(EXPERIMENT_NAME)
#Submit a pipeline run
run_name = pipeline_func.__name__ + ' run'
run_result = client.run_pipeline(experiment.id, run_name, pipeline_filename, arguments)
```
## References
* [Component Python code](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/component_sdk/python/kfp_component/google/dataproc/_submit_spark_job.py)
* [Component Docker file](https://github.com/kubeflow/pipelines/blob/master/components/gcp/container/Dockerfile)
* [Sample notebook](https://github.com/kubeflow/pipelines/blob/master/components/gcp/dataproc/submit_spark_job/sample.ipynb)
* [Dataproc SparkJob](https://cloud.google.com/dataproc/docs/reference/rest/v1/SparkJob)
## License
By deploying or using this software you agree to comply with the [AI Hub Terms of Service](https://aihub.cloud.google.com/u/0/aihub-tos) and the [Google APIs Terms of Service](https://developers.google.com/terms/). To the extent of a direct conflict of terms, the AI Hub Terms of Service will control.
| github_jupyter |
# Image Classification using LeNet CNN
## Fashion MNIST Dataset - Clothing Objects (10 classes)
### t-shirt/top, trouser, pullover, dress, coat, sandal, shirt, sneaker, bag, ankle boot

```
# import tensorflow module. Check API version.
import tensorflow as tf
import numpy as np
print (tf.__version__)
# required for TF to run within docker using GPU (ignore otherwise)
gpu = tf.config.experimental.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(gpu[0], True)
```
## Load the data
```
# grab the Fashion MNIST dataset (may take time the first time)
print("[INFO] downloading Fashion MNIST...")
(trainData, trainLabels), (testData, testLabels) = tf.keras.datasets.fashion_mnist.load_data()
```
## Prepare the data
```
# parameters for Fashion MNIST data set
num_classes = 10
image_width = 28
image_height = 28
image_channels = 1
# define human readable class names
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat/jacket',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# shape the input data using "channels last" ordering
# num_samples x rows x columns x depth
trainData = trainData.reshape(
(trainData.shape[0], image_height, image_width, image_channels))
testData = testData.reshape(
(testData.shape[0], image_height, image_width, image_channels))
# convert to floating point and scale data to the range of [0.0, 1.0]
trainData = trainData.astype("float32") / 255.0
testData = testData.astype("float32") / 255.0
# pad the data to 32X32 for use in LeNet5 network
trainData = np.pad(trainData, ((0,0),(2,2),(2,2),(0,0)), 'constant')
testData = np.pad(testData, ((0,0),(2,2),(2,2),(0,0)), 'constant')
# display data dimentions
print ("trainData:", trainData.shape)
print ("trainLabels:", trainLabels.shape)
print ("testData:", testData.shape)
print ("testLabels:", testLabels.shape)
# parameters for training data set
num_classes = 10
image_width = 32
image_height = 32
image_channels = 1
```
## Define Model

```
# import the necessary packages
from tensorflow.keras import backend
from tensorflow.keras import models
from tensorflow.keras import layers
# define the model as a class
class LeNet:
# INPUT => CONV => TANH => AVG-POOL => CONV => TANH => AVG-POOL => FC => TANH => FC => TANH => FC => SMAX
@staticmethod
def init(numChannels, imgRows, imgCols, numClasses, weightsPath=None):
# if we are using "channels first", update the input shape
if backend.image_data_format() == "channels_first":
inputShape = (numChannels, imgRows, imgCols)
else: # "channels last"
inputShape = (imgRows, imgCols, numChannels)
# initialize the model
model = models.Sequential()
# define the first set of CONV => ACTIVATION => POOL layers
model.add(layers.Conv2D(filters=6, kernel_size=(5, 5), strides=(1, 1),
padding="valid", activation=tf.nn.tanh, input_shape=inputShape))
model.add(layers.AveragePooling2D(pool_size=(2, 2), strides=(2, 2)))
# define the second set of CONV => ACTIVATION => POOL layers
model.add(layers.Conv2D(filters=16, kernel_size=(5, 5), strides=(1, 1),
padding="valid", activation=tf.nn.tanh))
model.add(layers.AveragePooling2D(pool_size=(2, 2), strides=(2, 2)))
# flatten the convolution volume to fully connected layers
model.add(layers.Flatten())
# define the first FC => ACTIVATION layers
model.add(layers.Dense(units=120, activation=tf.nn.tanh))
# define the second FC => ACTIVATION layers
model.add(layers.Dense(units=84, activation=tf.nn.tanh))
# lastly, define the soft-max classifier
model.add(layers.Dense(units=numClasses, activation=tf.nn.softmax))
# if a weights path is supplied (inicating that the model was
# pre-trained), then load the weights
if weightsPath is not None:
model.load_weights(weightsPath)
# return the constructed network architecture
return model
```
## Compile Model
```
# initialize the model
print("[INFO] compiling model...")
model = LeNet.init(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath=None)
# compile the model
model.compile(optimizer=tf.keras.optimizers.SGD(lr=0.01), # Stochastic Gradient Descent
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# print model summary
model.summary()
```
## Train Model
```
# define callback function for training termination criteria
#accuracy_cutoff = 0.99
class myCallback(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
if(logs.get('accuracy') > 0.95):
print("\nReached 95% accuracy so cancelling training!")
self.model.stop_training = True
# initialize training config
batch_size = 128
epochs = 200
# run training
print("[INFO] training...")
history = model.fit(x=trainData, y=trainLabels, validation_data=(testData, testLabels),
batch_size=batch_size, epochs=epochs, verbose=1, callbacks=[myCallback()])
```
## Evaluate Training Performance
### Expected Output
 
```
%matplotlib inline
import matplotlib.pyplot as plt
# retrieve a list of list results on training and test data sets for each training epoch
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
epochs = range(len(acc)) # get number of epochs
# plot training and validation accuracy per epoch
plt.plot(epochs, acc, label='train accuracy')
plt.plot(epochs, val_acc, label='val accuracy')
plt.xlabel('epochs')
plt.ylabel('accuracy')
plt.legend(loc="lower right")
plt.title('Training and validation accuracy')
plt.figure()
# plot training and validation loss per epoch
plt.plot(epochs, loss, label='train loss')
plt.plot(epochs, val_loss, label='val loss')
plt.xlabel('epochs')
plt.ylabel('loss')
plt.legend(loc="upper right")
plt.title('Training and validation loss')
# show the accuracy on the testing set
print("[INFO] evaluating...")
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
model.save_weights("weights/LeNetFashionMNIST.temp.hdf5", overwrite=True)
```
## Evaluate Pre-trained Model
```
# init model and load the model weights
print("[INFO] compiling model...")
model = LeNet.init(numChannels=image_channels,
imgRows=image_height, imgCols=image_width,
numClasses=num_classes,
weightsPath="weights/LeNetFashionMNIST.hdf5")
# compile the model
model.compile(optimizer=tf.keras.optimizers.SGD(lr=0.01), # Stochastic Gradient Descent
loss="sparse_categorical_crossentropy",
metrics=["accuracy"])
# show the accuracy on the testing set
print("[INFO] evaluating...")
batch_size = 128
(loss, accuracy) = model.evaluate(testData, testLabels,
batch_size=batch_size, verbose=1)
print("[INFO] accuracy: {:.2f}%".format(accuracy * 100))
```
## Model Predictions
```
%matplotlib inline
import numpy as np
import cv2
import matplotlib.pyplot as plt
# set up matplotlib fig, and size it to fit 3x4 pics
nrows = 3
ncols = 4
fig = plt.gcf()
fig.set_size_inches(ncols*4, nrows*4)
# randomly select a few testing digits
num_predictions = 12
test_indices = np.random.choice(np.arange(0, len(testLabels)), size=(num_predictions,))
test_images = np.stack(([testData[i] for i in test_indices]))
test_labels = np.stack(([testLabels[i] for i in test_indices]))
# compute predictions
predictions = model.predict(test_images)
for i in range(num_predictions):
# select the most probable class
prediction = np.argmax(predictions[i])
# rescale the test image
image = (test_images[i] * 255).astype("uint8")
# resize the image from a 28 x 28 image to a 96 x 96 image so we can better see it
image = cv2.resize(image, (96, 96), interpolation=cv2.INTER_CUBIC)
# convert grayscale image to RGB color
image = cv2.merge([image] * 3)
# select prediction text color
if prediction == test_labels[i]:
rgb_color = (0, 255, 0) # green for correct predictions
else:
rgb_color = (255, 0, 0) # red for wrong predictions
# show the image and prediction
cv2.putText(image, str(class_names[prediction]), (0, 10),
cv2.FONT_HERSHEY_SIMPLEX, 0.5, rgb_color, 1)
# set up subplot; subplot indices start at 1
sp = plt.subplot(nrows, ncols, i + 1, title="label: %s" % class_names[test_labels[i]])
sp.axis('Off') # don't show axes (or gridlines)
plt.imshow(image)
# show figure matrix
plt.show()
```
| github_jupyter |
```
import os
import sys
module_path = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
if module_path not in sys.path:
sys.path.append(module_path)
import librosa
import museval
import numpy as np
import tensorflow as tf
import IPython.display as ipd
from utils.helper import wav_to_spectrogram_clips, rebuild_audio_from_spectro_clips
from utils.dataset import create_samples
from models.conv_denoising_unet import ConvDenoisingUnet
from models.conv_encoder_denoising_decoder import ConvEncoderDenoisingDecoder
from models.conv_resblock_denoising_unet import ConvResblockDenoisingUnet
from evaluation import evaluate
#from evaluation.evaluate import get_separated_tracks, get_reference_tracks, estimate_and_evaluate
sorted(os.listdir(os.path.join(os.pardir, 'saved_model')))
samples = create_samples('Test')
test_sample = samples[20]
print(test_sample)
model_path = os.path.join(os.pardir, 'saved_model', 'conv_denoising_unet?time=20200223_0347.h5')
model = tf.keras.models.load_model(model_path)
def get_separated_tracks(separator, mix_audio):
# load mix music audio, average the stereo recording to single channel audio track
# convert to spectrogram
sound, sr = librosa.load(mix_audio, sr=44100, mono=True, duration=10)
stft = librosa.stft(sound, n_fft=2048, hop_length=512, win_length=2048)
mag, phase = librosa.magphase(stft)
# chop magnitude of spectrogram into clips, each has 1025 bins, 100 frames
stft_clips = np.empty((0, 1025, 100))
for i in range(mag.shape[1] // 100):
stft_clips = np.concatenate((stft_clips, mag[np.newaxis, :, i * 100: (i + 1) * 100])
# separate components from the mix single channel music audio
separated_sepctrograms = separator.predict(stft_clips)
separated_tracks = list()
# separated_spectrograms contains 4 stem tracks
# the index of spectrograms: 0, 1, 2, 3 -> vocals, bass, drums, other
for i in range(4):
separated_track = np.squeeze(separated_spectrograms[i], axis=-1)
separated_tracks.append(rebuild_audio_from_spectro_clips(separated_track))
return separated_tracks
def get_reference_tracks(sample, track_shape):
reference_tracks = list()
for feat in ['vocals', 'bass', 'drums', 'other']:
track, sr = librosa.load(sample[feat], sr=44100, mono=True, duration=10)
# crop reference track to match separated track shape
track = track[tuple(map(slice, track_shape))]
reference_tracks.append(track)
return reference_tracks
separate_tracks = get_separated_tracks(model, test_sample['mix'])
reference_tracks = get_reference_tracks(test_sample['mix'], separate_tracks[0].shape)
import mir_eval
(sdr, sir, sar, perm) = mir_eval.separation.bss_eval_sources(references, estimates, compute_permutation=False)
```
## wav_to_spectrogram_clips will remove some frames from the original spectrogram
### reconstructon
```
spectrogram_clips = wav_to_spectrogram_clips(test_sample['mix'])
print(spectrogram_clips.shape)
spectrogram = np.concatenate(spectrogram_clips, axis=1)
print(spectrogram_clips.shape)
audio = rebuild_audio_from_spectro_clips(spectrogram_clips)
print('reconstructed audio waveform from wav_to_spectrogram_clips', audio.shape)
```
### original
```
sound, sr = librosa.load(test_sample['mix'], sr=44100, mono=True)
stft = librosa.stft(sound, n_fft=2048, hop_length=512, win_length=2048)
mag, phase = librosa.magphase(stft)
print(mag.shape)
track, sr = librosa.load(test_sample['mix'], sr=44100, mono=True)
print('true size of the original audio waveform', track.shape)
```
| github_jupyter |
```
import keras
from keras.models import Sequential, Model, load_model
from keras.layers import Dense, Dropout, Activation, Flatten, Input, Lambda
from keras.layers import Conv2D, MaxPooling2D, AveragePooling2D, Conv1D, MaxPooling1D, LSTM, ConvLSTM2D, GRU, CuDNNLSTM, CuDNNGRU, BatchNormalization, LocallyConnected2D, Permute, TimeDistributed, Bidirectional
from keras.layers import Concatenate, Reshape, Softmax, Conv2DTranspose, Embedding, Multiply
from keras.callbacks import ModelCheckpoint, EarlyStopping, Callback
from keras import regularizers
from keras import backend as K
from keras.utils.generic_utils import Progbar
from keras.layers.merge import _Merge
import keras.losses
from keras.datasets import mnist
from functools import partial
from collections import defaultdict
import tensorflow as tf
from tensorflow.python.framework import ops
import isolearn.keras as iso
import numpy as np
import tensorflow as tf
import logging
logging.getLogger('tensorflow').setLevel(logging.ERROR)
import os
import pickle
import numpy as np
import scipy.sparse as sp
import scipy.io as spio
import matplotlib.pyplot as plt
from keras.backend.tensorflow_backend import set_session
def contain_tf_gpu_mem_usage() :
config = tf.ConfigProto()
config.gpu_options.allow_growth = True
sess = tf.Session(config=config)
set_session(sess)
contain_tf_gpu_mem_usage()
class EpochVariableCallback(Callback) :
def __init__(self, my_variable, my_func) :
self.my_variable = my_variable
self.my_func = my_func
def on_epoch_begin(self, epoch, logs={}) :
K.set_value(self.my_variable, self.my_func(K.get_value(self.my_variable), epoch))
#Load MNIST data
dataset_name = "mnist_3_vs_5"
img_rows, img_cols = 28, 28
num_classes = 10
batch_size = 32
included_classes = { 3, 5 }
(x_train, y_train), (x_test, y_test) = mnist.load_data()
keep_index_train = []
for i in range(y_train.shape[0]) :
if y_train[i] in included_classes :
keep_index_train.append(i)
keep_index_test = []
for i in range(y_test.shape[0]) :
if y_test[i] in included_classes :
keep_index_test.append(i)
x_train = x_train[keep_index_train]
x_test = x_test[keep_index_test]
y_train = y_train[keep_index_train]
y_test = y_test[keep_index_test]
n_train = int((x_train.shape[0] // batch_size) * batch_size)
n_test = int((x_test.shape[0] // batch_size) * batch_size)
x_train = x_train[:n_train]
x_test = x_test[:n_test]
y_train = y_train[:n_train]
y_test = y_test[:n_test]
x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols, 1)
x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols, 1)
input_shape = (img_rows, img_cols, 1)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
print("x_train.shape = " + str(x_train.shape))
print("n train samples = " + str(x_train.shape[0]))
print("n test samples = " + str(x_test.shape[0]))
y_train = keras.utils.to_categorical(y_train, num_classes)
y_test = keras.utils.to_categorical(y_test, num_classes)
#Binarize images
def _binarize_images(x, val_thresh=0.5) :
x_bin = np.zeros(x.shape)
x_bin[x >= val_thresh] = 1.
return x_bin
x_train = _binarize_images(x_train, val_thresh=0.5)
x_test = _binarize_images(x_test, val_thresh=0.5)
#Visualize background image distribution
pseudo_count = 1.0
x_mean = (np.sum(x_train, axis=(0, 3)) + pseudo_count) / (x_train.shape[0] + pseudo_count)
x_mean_logits = np.log(x_mean / (1. - x_mean))
f = plt.figure(figsize=(4, 4))
plot_ix = 0
plt.imshow(x_mean, cmap="Greys", vmin=0.0, vmax=1.0, aspect='equal')
plt.xticks([], [])
plt.yticks([], [])
plt.tight_layout()
plt.show()
from tensorflow.python.framework import ops
#Stochastic Binarized Neuron helper functions (Tensorflow)
#ST Estimator code adopted from https://r2rt.com/binary-stochastic-neurons-in-tensorflow.html
#See Github https://github.com/spitis/
def bernoulli_sample(x):
g = tf.get_default_graph()
with ops.name_scope("BernoulliSample") as name:
with g.gradient_override_map({"Ceil": "Identity","Sub": "BernoulliSample_ST"}):
return tf.ceil(x - tf.random_uniform(tf.shape(x)), name=name)
@ops.RegisterGradient("BernoulliSample_ST")
def bernoulliSample_ST(op, grad):
return [grad, tf.zeros(tf.shape(op.inputs[1]))]
#Masking and Sampling helper functions
def sample_image_st(x) :
p = tf.sigmoid(x)
return bernoulli_sample(p)
#Generator helper functions
def initialize_templates(generator, background_matrices) :
embedding_backgrounds = []
for k in range(len(background_matrices)) :
embedding_backgrounds.append(background_matrices[k].reshape(1, -1))
embedding_backgrounds = np.concatenate(embedding_backgrounds, axis=0)
generator.get_layer('background_dense').set_weights([embedding_backgrounds])
generator.get_layer('background_dense').trainable = False
#Generator construction function
def build_sampler(batch_size, n_rows, n_cols, n_classes=1, n_samples=1) :
#Initialize Reshape layer
reshape_layer = Reshape((n_rows, n_cols, 1))
#Initialize background matrix
background_dense = Embedding(n_classes, n_rows * n_cols, embeddings_initializer='zeros', name='background_dense')
#Initialize Templating and Masking Lambda layer
background_layer = Lambda(lambda x: x[0] + x[1], name='background_layer')
#Initialize Sigmoid layer
image_layer = Lambda(lambda x: K.sigmoid(x), name='image')
#Initialize Sampling layers
upsampling_layer = Lambda(lambda x: K.tile(x, [n_samples, 1, 1, 1]), name='upsampling_layer')
sampling_layer = Lambda(sample_image_st, name='image_sampler')
permute_layer = Lambda(lambda x: K.permute_dimensions(K.reshape(x, (n_samples, batch_size, n_rows, n_cols, 1)), (1, 0, 2, 3, 4)), name='permute_layer')
def _sampler_func(class_input, raw_logits) :
#Get Template and Mask
background = reshape_layer(background_dense(class_input))
#Add Template and Multiply Mask
image_logits = background_layer([raw_logits, background])
#Compute Image (Sigmoids from logits)
image = image_layer(image_logits)
#Tile each image to sample from and create sample axis
image_logits_upsampled = upsampling_layer(image_logits)
sampled_image = sampling_layer(image_logits_upsampled)
sampled_image = permute_layer(sampled_image)
return image_logits, image, sampled_image
return _sampler_func
#Initialize Encoder and Decoder networks
batch_size = 32
n_rows = 28
n_cols = 28
n_samples = 128
#Load sampler
sampler = build_sampler(batch_size, n_rows, n_cols, n_classes=1, n_samples=n_samples)
#Load Predictor
predictor_path = 'saved_models/mnist_binarized_cnn_10_digits.h5'
predictor = load_model(predictor_path)
predictor.trainable = False
predictor.compile(optimizer=keras.optimizers.SGD(lr=0.1), loss='mean_squared_error')
#Build scrambler model
dummy_class = Input(shape=(1,), name='dummy_class')
input_logits = Input(shape=(n_rows, n_cols, 1), name='input_logits')
image_logits, image, sampled_image = sampler(dummy_class, input_logits)
scrambler_model = Model([input_logits, dummy_class], [image_logits, image, sampled_image])
#Initialize Templates and Masks
initialize_templates(scrambler_model, [x_mean_logits])
scrambler_model.trainable = False
scrambler_model.compile(
optimizer=keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999),
loss='mean_squared_error'
)
scrambler_model.summary()
file_names = [
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_025_kl_divergence_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_03_kl_divergence_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_03_kl_divergence_even_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_example_entropy_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_01_kl_divergence_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_example_entropy_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_005_kl_divergence_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_example_entropy_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_0035_kl_divergence_higher_entropy_penalty_importance_scores_test.npy",
"autoscrambler_dataset_mnist_3_vs_5_inverted_scores_example_entropy_n_samples_32_resnet_5_4_32_3_00_n_epochs_50_target_bits_001_kl_divergence_higher_entropy_penalty_importance_scores_test.npy"
]
model_names =[
"scrambler_inverted",
"scrambler_inverted_higher_entropy_penalty",
"scrambler_inverted_even_higher_entropy_penalty",
"scrambler_inverted",
"scrambler_inverted_higher_entropy_penalty",
"scrambler_inverted_even_higher_entropy_penalty",
"scrambler_inverted_even_even_higher_entropy_penalty"
]
model_importance_scores_test = [np.load(file_name) for file_name in file_names]
feature_quantiles = [0.80, 0.90, 0.95, 0.98]
on_state_logit_val = 50.
dummy_test = np.zeros((x_test.shape[0], 1))
x_test_logits = 2. * x_test - 1.
digit_test = np.argmax(y_test, axis=-1)
y_pred_ref = predictor.predict([x_test], batch_size=32, verbose=True)
model_kl_divergences = []
for model_i in range(len(model_names)) :
print("Benchmarking model '" + str(model_names[model_i]) + "'...")
feature_quantile_kl_divergences = []
for feature_quantile in feature_quantiles :
print("Feature quantile = " + str(feature_quantile))
importance_scores_test = np.abs(model_importance_scores_test[model_i])
quantile_vals = np.quantile(importance_scores_test, axis=(1, 2, 3), q=feature_quantile, keepdims=True)
quantile_vals = np.tile(quantile_vals, (1, importance_scores_test.shape[1], importance_scores_test.shape[2], importance_scores_test.shape[3]))
top_logits_test = np.zeros(importance_scores_test.shape)
top_logits_test[importance_scores_test <= quantile_vals] = on_state_logit_val
print( len(np.nonzero(np.ravel(top_logits_test) > 10.0)[0]) )
top_logits_test = top_logits_test * x_test_logits
_, _, samples_test = scrambler_model.predict([top_logits_test, dummy_test], batch_size=batch_size)
mean_kl_divs = []
for data_ix in range(samples_test.shape[0]) :
if data_ix % 100 == 0 :
print("Processing example " + str(data_ix) + "...")
y_pred_var_samples = predictor.predict([samples_test[data_ix, ...]], batch_size=n_samples)
y_pred_ref_samples = np.tile(np.expand_dims(y_pred_ref[data_ix, :], axis=0), (n_samples, 1))
kl_divs = np.sum(y_pred_ref_samples * np.log(y_pred_ref_samples / y_pred_var_samples), axis=-1)
#kl_divs = (y_pred_ref_samples[:, digit_test[data_ix]] - np.mean(y_pred_ref_samples, axis=-1)) - (y_pred_var_samples[:, digit_test[data_ix]] - np.mean(y_pred_var_samples, axis=-1))
mean_kl_div = np.mean(kl_divs)
mean_kl_divs.append(mean_kl_div)
mean_kl_divs = np.array(mean_kl_divs)
feature_quantile_kl_divergences.append(mean_kl_divs)
model_kl_divergences.append(feature_quantile_kl_divergences)
model_names =[
"scrambler\n(inverted)",
"scrambler\n(inverted,\nhigher entropy)",
"scrambler\n(inverted,\neven higher entropy)",
"scrambler\n(inverted)",
"scrambler\n(inverted,\nhigher entropy)",
"scrambler\n(inverted,\neven higher entropy)",
"scrambler\n(inverted,\nhighest entropy)"
]
def lighten_color(color, amount=0.5):
import matplotlib.colors as mc
import colorsys
try:
c = mc.cnames[color]
except:
c = color
c = colorsys.rgb_to_hls(*mc.to_rgb(c))
return colorsys.hls_to_rgb(c[0], 1 - amount * (1 - c[1]), c[2])
fig = plt.figure(figsize=(13, 6))
benchmark_name = "benchmark_ablation_mnist_scramblers"
save_figs = True
width = 0.2
max_y_val = 4.0
cm = plt.get_cmap('viridis_r')
shades = [0.4, 0.6, 0.8, 1]
quantiles = [0.5, 0.8, 0.9, 0.95]
all_colors = plt.rcParams['axes.prop_cycle'].by_key()['color'] + plt.rcParams['axes.prop_cycle'].by_key()['color']
model_colors = {model_names[i]: all_colors[i] for i in range(len(model_names))}
results = np.zeros((len(quantiles), len(model_names), 1))
for i in range(1, len(feature_quantiles) + 1) :
for j in range(len(model_names)):
kl_div_samples = model_kl_divergences[j][i-1]
for l in range(len(quantiles)):
quantile = quantiles[l]
results[l, j, 0] = np.quantile(kl_div_samples, q=quantile)
xs = range(len(model_names))
xs = [xi + i*width for xi in xs]
for j in range(len(model_names)) :
for l in range(len(quantiles)) :
model_name = model_names[j]
c = model_colors[model_name]
val = results[l, j, 0]
if i == 1 and j == 0 :
lbl = "$%i^{th}$ Perc." % int(100*quantiles[l])
else :
lbl=None
if l == 0 :
plt.bar(xs[j], val, width=width, color=lighten_color(c, shades[l]), edgecolor='k', linewidth=1, label=lbl, zorder=l+1)
else :
prev_val = results[l-1, j].mean(axis=-1)
plt.bar(xs[j],val-prev_val, width=width, bottom = prev_val, color=lighten_color(c, shades[l]), edgecolor='k', linewidth=1, label=lbl, zorder=l+1)
if l == len(quantiles) - 1 :
plt.text(xs[j], val, "Top\n" + str(int(100 - 100 * feature_quantiles[i-1])) + "%", horizontalalignment='center', verticalalignment='bottom', fontdict={ 'family': 'serif', 'color': 'black', 'weight': 'bold', 'size': 10 })
prev_results = results
plt.xticks([i + 2.5*width for i in range(len(model_names))])
all_lbls = [model_names[j].upper() for j in range(len(model_names))]
plt.gca().set_xticklabels(all_lbls, rotation=60)
plt.ylabel("Test Set KL-Divergence")
max_y_val = np.max(results) * 1.1
#plt.ylim([0, max_y_val])
plt.grid(True)
plt.gca().set_axisbelow(True)
plt.gca().grid(color='gray', alpha=0.2)
plt.gca().spines['right'].set_visible(False)
plt.gca().spines['top'].set_visible(False)
plt.gca().yaxis.set_ticks_position('left')
plt.gca().xaxis.set_ticks_position('bottom')
plt.legend(fontsize=12, frameon=True, loc='upper left')
leg = plt.gca().get_legend()
for l in range(len(quantiles)):
leg.legendHandles[l].set_color(lighten_color(all_colors[7], shades[l]))
leg.legendHandles[l].set_edgecolor('k')
plt.tight_layout()
if save_figs :
plt.savefig(benchmark_name + ".png", dpi=300, transparent=True)
plt.savefig(benchmark_name + ".eps")
plt.show()
```
| github_jupyter |
## Loading my Records file
```
!git clone https://github.com/AshishJangra27/Skill-India-AI-ML-Scholarship
```
## Generating Sales and Record file for the first time
```
fd = open('/content/Skill-India-AI-ML-Scholarship/JSON Based Inventory management Assignment/record.json', 'r')
t = fd.read()
fd.close()
fd = open('record.json','w')
fd.write(t)
fd.close()
sales = {}
sales = json.dumps(sales)
fd = open("sales.json", 'w')
fd.write(sales)
fd.close()
```
## Importing Libraries
```
import json
import time
# Loading Inventory and Converting it to Dictionary
fd = open('record.json','r')
t = fd.read()
fd.close()
dct = json.loads(t)
# Loading Sales and Converting it to Dictionary
fd = open("sales.json", 'r')
sl = fd.read()
fd.close()
sales = json.loads(sl)
# Displaying Menu
print("**********************************************")
for i in dct.keys():
print(i, dct[i])
print("******************************************\n")
# Taking Input from User to Enter what he/she wants to purchase
ui_prod = str(input("Enter the product_Id: "))
ui_quant = int(input("Enter the quantity: "))
if (ui_prod in dct.keys()): # Checking If product Exists or not
if (dct[ui_prod]['qn'] >= ui_quant): # If I'm having enough Quantity
print("Name:",dct[ui_prod]['name'])
print("Price:",dct[ui_prod]['pr'])
print("Quantity:", ui_quant)
print("------------------------------")
print("Billing Amount:", dct[ui_prod]['pr'] * ui_quant)
dct[ui_prod]['qn'] = dct[ui_prod]['qn'] - ui_quant # Updating Inventory
sales[str(len(sales)+1)] = {'prod_id' : ui_prod, # Updating Transection
"time_pr" : time.ctime(),
'bill' : dct[ui_prod]['pr'] * ui_quant,
'qn' : ui_quant,
'pr' : dct[ui_prod]['pr'],
'pr_name' : dct[ui_prod]['name']}
else: # If I'm not having enough Quantity
print("Sorry, We are not having that much of quantity.")
print("We're only having",dct[ui_prod]['qn'],"quantity.")
print("Would you like to purchase or not (Y/N)")
ch = str(input("Y/N"))
if (ch == "Y"): # If user wants to purchase the quantitry we're having
print("Name:",dct[ui_prod]['name'])
print("Price:",dct[ui_prod]['pr'])
print("Quantity:", dct[ui_prod]['qn'])
print("------------------------------")
print("Billing Amount:", dct[ui_prod]['pr'] * dct[ui_prod]['qn'])
dct[ui_prod]['qn'] = 0 # Updating Inventory
sales[str(len(sales)+1)] = {'prod_id' : ui_prod, # Updating Transection
"time_pr" : time.ctime(),
'bill' : dct[ui_prod]['pr'] * dct[ui_prod]['qn'],
'qn' : dct[ui_prod]['qn'],
'pr' : dct[ui_prod]['pr'],
'pr_name' : dct[ui_prod]['name']}
else: # If user press anything Except Y
print("Thankyou!")
else: # Product ID Doesn't exists
print("------------------------------")
print("Product doesn't exist!")
print("Please enter a valid product id")
print("------------------------------")
sl = json.dumps(sales) # Updating Sales File
fd = open('sales.json','w')
fd.write(sl)
fd.close()
print("------------------------------")
print("Data Updated in Sales File!")
js = json.dumps(dct) # Updating Inventory File
fd = open('record.json','w')
fd.write(js)
fd.close()
print("Data Updated in Inventory File!")
dct
sales
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import seaborn as sns
import numpy as np
df = pd.read_csv('data.csv')
list(df.columns.values)
# Data cleaning
df.loc[df.AnzahlSexualpartnerAllgemein > 10000, 'AnzahlSexualpartnerAllgemein'] = float('inf')
df.loc[df.AnzahlSexualpartnerGreifswald > 10000, 'AnzahlSexualpartnerGreifswald'] = float('inf')
df.loc[df.AnzahlStechmuecken > 10000, 'AnzahlStechmuecken'] = float('inf')
df.loc[df.Groesse < 20, 'Groesse'] = None
# sorted(df[['LaengeFreundschaft']].values)
df.loc[df.LaengeFreundschaft > 100, ['Alter', 'LaengeFreundschaft']] = None
grouped = df.groupby('Geschlecht')
grouped['EigeneAttraktivitaet'].hist(alpha=0.4, position=0, bins=range(1, 11))
df.loc[(df.Geschlecht == 'Weiblich') & (df.AnzahlStechmuecken < 100), 'AnzahlStechmuecken'].plot.hist()
df.loc[(df.Geschlecht == 'Männlich') & (df.AnzahlStechmuecken < 100), 'AnzahlStechmuecken'].plot.hist()
df.loc[(df.Geschlecht == 'Weiblich') & (df.AnzahlStechmuecken < 100), 'AnzahlStechmuecken'].plot.hist(alpha=0.5)
df.loc[(df.Geschlecht == 'Männlich') & (df.AnzahlStechmuecken < 100), 'AnzahlStechmuecken'].plot.hist(alpha=0.5)
df.loc[df.Geschlecht == 'Männlich', 'EigeneAttraktivitaet'].plot.hist(bins=range(1, 11),alpha=0.5)
# df.loc['EigeneAttraktivitaet'].plot.hist()
df.loc[df.Geschlecht == 'Weiblich', 'EigeneAttraktivitaet'].plot.hist(bins=range(1, 11), alpha=0.5)
categorial = ['Geschlecht',
'Studienfach',
'AG',
'ZungeRollen',
'BeziehungsstatusVorher',
'BeziehungsstatusNachher',
'Tinder',
'GrundSpaetInsBett',
'BesterAbendvortrag',
'Transportmittel',
'Gendern',
'PlutoPlanet',
'ArtikelButter',
'ArtikelNutella',
'NutellaMitButter',
'SchoensterDialekt',
'PfluegenOderNicht',
'BestaussehenderMann',
'BestaussehendeFrau',
'WegZumStipendium',
'Fu\xc3\x9fnoten',
'BeziehungMitStifti',
'InEinemWort']
numeric_cols = ['Alter',
'AnzahlSommerakademien',
'LaengeAnfahrt',
'AnzahlSprachen',
'AnzahlGeschwister',
'AnzahlVornamen',
'Groesse',
'AnzahlStaatsbuergerschaften',
'LaengeFreundschaft',
'AnzahlPokemon',
'MengeKaffee',
'AnzahlSexualpartnerAllgemein',
'AnzahlSexualpartnerGreifswald',
'StundenSport',
'AnzahlBuecher',
'AnzahlFacebookfreunde',
'DurchschnittAbi',
'DurchschnittAktuell',
'QuersummePIN',
'MengeBargeld',
'AnzahlZimmergenossen',
'MengeSchlaf',
'DauerVorbereitungZuhause',
'DauerVorbereitungGreifswald',
'GeldMensakarte',
'AnzahlAbendvortraegeGeschwaenzt',
'AnzahlFremdeBetten',
'ZeitAufWasser',
'AnzahlStechmuecken',
'EigeneAttraktivitaet',]
print("Datenpunkte: %i" % len(df))
for val in numeric_cols:
if val in ['AnzahlSexualpartnerAllgemein', 'AnzahlSexualpartnerGreifswald', 'AnzahlStechmuecken']:
continue
print("{0:>32}: {1:>06.2f} (Range: {2:>4.2f} - {3:>4.2f})\t Mean: {4:>5.2f}".format(
val,
float(df[[val]].median()),
float(df[[val]].min()),
float(df[[val]].max()),
float(df[[val]].mean())))
print("{0:>32}, Group: abs min (AG {1: .0f}) abs max (AG {2: .0f}), mean min (AG {3: .0f}), mean max (AG {4: .0f})".format(
val,
df.groupby('AG')[val].min().idxmin(),
df.groupby('AG')[val].max().idxmax(),
df.groupby('AG')[val].mean().idxmin(),
df.groupby('AG')[val].mean().idxmax()))
print("{0:>32}, Group: median min={1} (AG {2: .0f}), median max={3} (AG {4: .0f}), sum={5} (AG {6: .0f})".format(
val,
df.groupby('AG')[val].median().min(),
df.groupby('AG')[val].median().idxmin(),
df.groupby('AG')[val].median().max(),
df.groupby('AG')[val].median().idxmax(),
df.groupby('AG')[val].sum().max(),
df.groupby('AG')[val].sum().idxmax()))
val_min = int(min(df[val]))
val_max = int(min(10000, max(df[val])))
# df[[val]].plot.hist(bins=np.linspace(minimum, maximum, 10), alpha=0.5, normed=True)
#df.loc[df.Geschlecht == 'Weiblich', val].plot.hist(bins=np.linspace(minimum, maximum, 10), alpha=0.5, normed=True)
# df.loc[df.Geschlecht == 'Männlich', val].plot.hist(bins=np.linspace(minimum, maximum, 10), alpha=0.5, normed=True)
df.groupby('AG')['ZungeRollen'].describe()
# TODO
df.groupby('AG')["StundenSport"].mean()
df.groupby('Geschlecht').describe()
df[df.AG == 5]
df[(df.Geschlecht == "Weiblich") & (df.PlutoPlanet == 'Ist das nicht der Hund von Goofy?')]
gb = df[(df.Geschlecht == "Weiblich")].groupby("PlutoPlanet")
gb.sum()
gb = df.groupby("AG")
gb.sum()[[""]]
gb.sum()
gb.groups
```
| github_jupyter |
# Regularization
Welcome to the second assignment of this week. Deep Learning models have so much flexibility and capacity that **overfitting can be a serious problem**, if the training dataset is not big enough. Sure it does well on the training set, but the learned network **doesn't generalize to new examples** that it has never seen!
**You will learn to:** Use regularization in your deep learning models.
Let's first import the packages you are going to use.
```
# import packages
import numpy as np
import matplotlib.pyplot as plt
from reg_utils import sigmoid, relu, plot_decision_boundary, initialize_parameters, load_2D_dataset, predict_dec
from reg_utils import compute_cost, predict, forward_propagation, backward_propagation, update_parameters
import sklearn
import sklearn.datasets
import scipy.io
from testCases import *
%matplotlib inline
plt.rcParams['figure.figsize'] = (7.0, 4.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
```
**Problem Statement**: You have just been hired as an AI expert by the French Football Corporation. They would like you to recommend positions where France's goal keeper should kick the ball so that the French team's players can then hit it with their head.
<img src="images/field_kiank.png" style="width:600px;height:350px;">
<caption><center> <u> **Figure 1** </u>: **Football field**<br> The goal keeper kicks the ball in the air, the players of each team are fighting to hit the ball with their head </center></caption>
They give you the following 2D dataset from France's past 10 games.
```
train_X, train_Y, test_X, test_Y = load_2D_dataset()
```
Each dot corresponds to a position on the football field where a football player has hit the ball with his/her head after the French goal keeper has shot the ball from the left side of the football field.
- If the dot is blue, it means the French player managed to hit the ball with his/her head
- If the dot is red, it means the other team's player hit the ball with their head
**Your goal**: Use a deep learning model to find the positions on the field where the goalkeeper should kick the ball.
**Analysis of the dataset**: This dataset is a little noisy, but it looks like a diagonal line separating the upper left half (blue) from the lower right half (red) would work well.
You will first try a non-regularized model. Then you'll learn how to regularize it and decide which model you will choose to solve the French Football Corporation's problem.
## 1 - Non-regularized model
You will use the following neural network (already implemented for you below). This model can be used:
- in *regularization mode* -- by setting the `lambd` input to a non-zero value. We use "`lambd`" instead of "`lambda`" because "`lambda`" is a reserved keyword in Python.
- in *dropout mode* -- by setting the `keep_prob` to a value less than one
You will first try the model without any regularization. Then, you will implement:
- *L2 regularization* -- functions: "`compute_cost_with_regularization()`" and "`backward_propagation_with_regularization()`"
- *Dropout* -- functions: "`forward_propagation_with_dropout()`" and "`backward_propagation_with_dropout()`"
In each part, you will run this model with the correct inputs so that it calls the functions you've implemented. Take a look at the code below to familiarize yourself with the model.
```
def model(X, Y, learning_rate = 0.3, num_iterations = 30000, print_cost = True, lambd = 0, keep_prob = 1):
"""
Implements a three-layer neural network: LINEAR->RELU->LINEAR->RELU->LINEAR->SIGMOID.
Arguments:
X -- input data, of shape (input size, number of examples)
Y -- true "label" vector (1 for blue dot / 0 for red dot), of shape (output size, number of examples)
learning_rate -- learning rate of the optimization
num_iterations -- number of iterations of the optimization loop
print_cost -- If True, print the cost every 10000 iterations
lambd -- regularization hyperparameter, scalar
keep_prob - probability of keeping a neuron active during drop-out, scalar.
Returns:
parameters -- parameters learned by the model. They can then be used to predict.
"""
grads = {}
costs = [] # to keep track of the cost
m = X.shape[1] # number of examples
layers_dims = [X.shape[0], 20, 3, 1]
# Initialize parameters dictionary.
parameters = initialize_parameters(layers_dims)
# Loop (gradient descent)
for i in range(0, num_iterations):
# Forward propagation: LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID.
if keep_prob == 1:
a3, cache = forward_propagation(X, parameters)
elif keep_prob < 1:
a3, cache = forward_propagation_with_dropout(X, parameters, keep_prob)
# Cost function
if lambd == 0:
cost = compute_cost(a3, Y)
else:
cost = compute_cost_with_regularization(a3, Y, parameters, lambd)
# Backward propagation.
assert(lambd==0 or keep_prob==1) # it is possible to use both L2 regularization and dropout,
# but this assignment will only explore one at a time
if lambd == 0 and keep_prob == 1:
grads = backward_propagation(X, Y, cache)
elif lambd != 0:
grads = backward_propagation_with_regularization(X, Y, cache, lambd)
elif keep_prob < 1:
grads = backward_propagation_with_dropout(X, Y, cache, keep_prob)
# Update parameters.
parameters = update_parameters(parameters, grads, learning_rate)
# Print the loss every 10000 iterations
if print_cost and i % 10000 == 0:
print("Cost after iteration {}: {}".format(i, cost))
if print_cost and i % 1000 == 0:
costs.append(cost)
# plot the cost
plt.plot(costs)
plt.ylabel('cost')
plt.xlabel('iterations (x1,000)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
return parameters
```
Let's train the model without any regularization, and observe the accuracy on the train/test sets.
```
parameters = model(train_X, train_Y)
print ("On the training set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
The train accuracy is 94.8% while the test accuracy is 91.5%. This is the **baseline model** (you will observe the impact of regularization on this model). Run the following code to plot the decision boundary of your model.
```
plt.title("Model without regularization")
axes = plt.gca()
axes.set_xlim([-0.75,0.40])
axes.set_ylim([-0.75,0.65])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
The non-regularized model is obviously overfitting the training set. It is fitting the noisy points! Lets now look at two techniques to reduce overfitting.
## 2 - L2 Regularization
The standard way to avoid overfitting is called **L2 regularization**. It consists of appropriately modifying your cost function, from:
$$J = -\frac{1}{m} \sum\limits_{i = 1}^{m} \large{(}\small y^{(i)}\log\left(a^{[L](i)}\right) + (1-y^{(i)})\log\left(1- a^{[L](i)}\right) \large{)} \tag{1}$$
To:
$$J_{regularized} = \small \underbrace{-\frac{1}{m} \sum\limits_{i = 1}^{m} \large{(}\small y^{(i)}\log\left(a^{[L](i)}\right) + (1-y^{(i)})\log\left(1- a^{[L](i)}\right) \large{)} }_\text{cross-entropy cost} + \underbrace{\frac{1}{m} \frac{\lambda}{2} \sum\limits_l\sum\limits_k\sum\limits_j W_{k,j}^{[l]2} }_\text{L2 regularization cost} \tag{2}$$
Let's modify your cost and observe the consequences.
**Exercise**: Implement `compute_cost_with_regularization()` which computes the cost given by formula (2). To calculate $\sum\limits_k\sum\limits_j W_{k,j}^{[l]2}$ , use :
```python
np.sum(np.square(Wl))
```
Note that you have to do this for $W^{[1]}$, $W^{[2]}$ and $W^{[3]}$, then sum the three terms and multiply by $ \frac{1}{m} \frac{\lambda}{2} $.
```
# GRADED FUNCTION: compute_cost_with_regularization
def compute_cost_with_regularization(A3, Y, parameters, lambd):
"""
Implement the cost function with L2 regularization. See formula (2) above.
Arguments:
A3 -- post-activation, output of forward propagation, of shape (output size, number of examples)
Y -- "true" labels vector, of shape (output size, number of examples)
parameters -- python dictionary containing parameters of the model
Returns:
cost - value of the regularized loss function (formula (2))
"""
m = Y.shape[1]
W1 = parameters["W1"]
W2 = parameters["W2"]
W3 = parameters["W3"]
cross_entropy_cost = compute_cost(A3, Y) # This gives you the cross-entropy part of the cost
### START CODE HERE ### (approx. 1 line)
L2_regularization_cost = None
### END CODER HERE ###
cost = cross_entropy_cost + L2_regularization_cost
return cost
A3, Y_assess, parameters = compute_cost_with_regularization_test_case()
print("cost = " + str(compute_cost_with_regularization(A3, Y_assess, parameters, lambd = 0.1)))
```
**Expected Output**:
<table>
<tr>
<td>
**cost**
</td>
<td>
1.78648594516
</td>
</tr>
</table>
Of course, because you changed the cost, you have to change backward propagation as well! All the gradients have to be computed with respect to this new cost.
**Exercise**: Implement the changes needed in backward propagation to take into account regularization. The changes only concern dW1, dW2 and dW3. For each, you have to add the regularization term's gradient ($\frac{d}{dW} ( \frac{1}{2}\frac{\lambda}{m} W^2) = \frac{\lambda}{m} W$).
```
# GRADED FUNCTION: backward_propagation_with_regularization
def backward_propagation_with_regularization(X, Y, cache, lambd):
"""
Implements the backward propagation of our baseline model to which we added an L2 regularization.
Arguments:
X -- input dataset, of shape (input size, number of examples)
Y -- "true" labels vector, of shape (output size, number of examples)
cache -- cache output from forward_propagation()
lambd -- regularization hyperparameter, scalar
Returns:
gradients -- A dictionary with the gradients with respect to each parameter, activation and pre-activation variables
"""
m = X.shape[1]
(Z1, A1, W1, b1, Z2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
### START CODE HERE ### (approx. 1 line)
dW3 = 1./m * np.dot(dZ3, A2.T) + None
### END CODE HERE ###
db3 = 1./m * np.sum(dZ3, axis=1, keepdims = True)
dA2 = np.dot(W3.T, dZ3)
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
### START CODE HERE ### (approx. 1 line)
dW2 = 1./m * np.dot(dZ2, A1.T) + None
### END CODE HERE ###
db2 = 1./m * np.sum(dZ2, axis=1, keepdims = True)
dA1 = np.dot(W2.T, dZ2)
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
### START CODE HERE ### (approx. 1 line)
dW1 = 1./m * np.dot(dZ1, X.T) + None
### END CODE HERE ###
db1 = 1./m * np.sum(dZ1, axis=1, keepdims = True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,"dA2": dA2,
"dZ2": dZ2, "dW2": dW2, "db2": db2, "dA1": dA1,
"dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
X_assess, Y_assess, cache = backward_propagation_with_regularization_test_case()
grads = backward_propagation_with_regularization(X_assess, Y_assess, cache, lambd = 0.7)
print ("dW1 = "+ str(grads["dW1"]))
print ("dW2 = "+ str(grads["dW2"]))
print ("dW3 = "+ str(grads["dW3"]))
```
**Expected Output**:
<table>
<tr>
<td>
**dW1**
</td>
<td>
[[-0.25604646 0.12298827 -0.28297129]
[-0.17706303 0.34536094 -0.4410571 ]]
</td>
</tr>
<tr>
<td>
**dW2**
</td>
<td>
[[ 0.79276486 0.85133918]
[-0.0957219 -0.01720463]
[-0.13100772 -0.03750433]]
</td>
</tr>
<tr>
<td>
**dW3**
</td>
<td>
[[-1.77691347 -0.11832879 -0.09397446]]
</td>
</tr>
</table>
Let's now run the model with L2 regularization $(\lambda = 0.7)$. The `model()` function will call:
- `compute_cost_with_regularization` instead of `compute_cost`
- `backward_propagation_with_regularization` instead of `backward_propagation`
```
parameters = model(train_X, train_Y, lambd = 0.7)
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
Congrats, the test set accuracy increased to 93%. You have saved the French football team!
You are not overfitting the training data anymore. Let's plot the decision boundary.
```
plt.title("Model with L2-regularization")
axes = plt.gca()
axes.set_xlim([-0.75,0.40])
axes.set_ylim([-0.75,0.65])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Observations**:
- The value of $\lambda$ is a hyperparameter that you can tune using a dev set.
- L2 regularization makes your decision boundary smoother. If $\lambda$ is too large, it is also possible to "oversmooth", resulting in a model with high bias.
**What is L2-regularization actually doing?**:
L2-regularization relies on the assumption that a model with small weights is simpler than a model with large weights. Thus, by penalizing the square values of the weights in the cost function you drive all the weights to smaller values. It becomes too costly for the cost to have large weights! This leads to a smoother model in which the output changes more slowly as the input changes.
<font color='blue'>
**What you should remember** -- the implications of L2-regularization on:
- The cost computation:
- A regularization term is added to the cost
- The backpropagation function:
- There are extra terms in the gradients with respect to weight matrices
- Weights end up smaller ("weight decay"):
- Weights are pushed to smaller values.
## 3 - Dropout
Finally, **dropout** is a widely used regularization technique that is specific to deep learning.
**It randomly shuts down some neurons in each iteration.** Watch these two videos to see what this means!
<!--
To understand drop-out, consider this conversation with a friend:
- Friend: "Why do you need all these neurons to train your network and classify images?".
- You: "Because each neuron contains a weight and can learn specific features/details/shape of an image. The more neurons I have, the more featurse my model learns!"
- Friend: "I see, but are you sure that your neurons are learning different features and not all the same features?"
- You: "Good point... Neurons in the same layer actually don't talk to each other. It should be definitly possible that they learn the same image features/shapes/forms/details... which would be redundant. There should be a solution."
!-->
<center>
<video width="620" height="440" src="images/dropout1_kiank.mp4" type="video/mp4" controls>
</video>
</center>
<br>
<caption><center> <u> Figure 2 </u>: Drop-out on the second hidden layer. <br> At each iteration, you shut down (= set to zero) each neuron of a layer with probability $1 - keep\_prob$ or keep it with probability $keep\_prob$ (50% here). The dropped neurons don't contribute to the training in both the forward and backward propagations of the iteration. </center></caption>
<center>
<video width="620" height="440" src="images/dropout2_kiank.mp4" type="video/mp4" controls>
</video>
</center>
<caption><center> <u> Figure 3 </u>: Drop-out on the first and third hidden layers. <br> $1^{st}$ layer: we shut down on average 40% of the neurons. $3^{rd}$ layer: we shut down on average 20% of the neurons. </center></caption>
When you shut some neurons down, you actually modify your model. The idea behind drop-out is that at each iteration, you train a different model that uses only a subset of your neurons. With dropout, your neurons thus become less sensitive to the activation of one other specific neuron, because that other neuron might be shut down at any time.
### 3.1 - Forward propagation with dropout
**Exercise**: Implement the forward propagation with dropout. You are using a 3 layer neural network, and will add dropout to the first and second hidden layers. We will not apply dropout to the input layer or output layer.
**Instructions**:
You would like to shut down some neurons in the first and second layers. To do that, you are going to carry out 4 Steps:
1. In lecture, we dicussed creating a variable $d^{[1]}$ with the same shape as $a^{[1]}$ using `np.random.rand()` to randomly get numbers between 0 and 1. Here, you will use a vectorized implementation, so create a random matrix $D^{[1]} = [d^{[1](1)} d^{[1](2)} ... d^{[1](m)}] $ of the same dimension as $A^{[1]}$.
2. Set each entry of $D^{[1]}$ to be 0 with probability (`1-keep_prob`) or 1 with probability (`keep_prob`), by thresholding values in $D^{[1]}$ appropriately. Hint: to set all the entries of a matrix X to 0 (if entry is less than 0.5) or 1 (if entry is more than 0.5) you would do: `X = (X < 0.5)`. Note that 0 and 1 are respectively equivalent to False and True.
3. Set $A^{[1]}$ to $A^{[1]} * D^{[1]}$. (You are shutting down some neurons). You can think of $D^{[1]}$ as a mask, so that when it is multiplied with another matrix, it shuts down some of the values.
4. Divide $A^{[1]}$ by `keep_prob`. By doing this you are assuring that the result of the cost will still have the same expected value as without drop-out. (This technique is also called inverted dropout.)
```
# GRADED FUNCTION: forward_propagation_with_dropout
def forward_propagation_with_dropout(X, parameters, keep_prob = 0.5):
"""
Implements the forward propagation: LINEAR -> RELU + DROPOUT -> LINEAR -> RELU + DROPOUT -> LINEAR -> SIGMOID.
Arguments:
X -- input dataset, of shape (2, number of examples)
parameters -- python dictionary containing your parameters "W1", "b1", "W2", "b2", "W3", "b3":
W1 -- weight matrix of shape (20, 2)
b1 -- bias vector of shape (20, 1)
W2 -- weight matrix of shape (3, 20)
b2 -- bias vector of shape (3, 1)
W3 -- weight matrix of shape (1, 3)
b3 -- bias vector of shape (1, 1)
keep_prob - probability of keeping a neuron active during drop-out, scalar
Returns:
A3 -- last activation value, output of the forward propagation, of shape (1,1)
cache -- tuple, information stored for computing the backward propagation
"""
np.random.seed(1)
# retrieve parameters
W1 = parameters["W1"]
b1 = parameters["b1"]
W2 = parameters["W2"]
b2 = parameters["b2"]
W3 = parameters["W3"]
b3 = parameters["b3"]
# LINEAR -> RELU -> LINEAR -> RELU -> LINEAR -> SIGMOID
Z1 = np.dot(W1, X) + b1
A1 = relu(Z1)
### START CODE HERE ### (approx. 4 lines) # Steps 1-4 below correspond to the Steps 1-4 described above.
D1 = None # Step 1: initialize matrix D1 = np.random.rand(..., ...)
D1 = None # Step 2: convert entries of D1 to 0 or 1 (using keep_prob as the threshold)
A1 = None # Step 3: shut down some neurons of A1
A1 = None # Step 4: scale the value of neurons that haven't been shut down
### END CODE HERE ###
Z2 = np.dot(W2, A1) + b2
A2 = relu(Z2)
### START CODE HERE ### (approx. 4 lines)
D2 = None # Step 1: initialize matrix D2 = np.random.rand(..., ...)
D2 = None # Step 2: convert entries of D2 to 0 or 1 (using keep_prob as the threshold)
A2 = None # Step 3: shut down some neurons of A2
A2 = None # Step 4: scale the value of neurons that haven't been shut down
### END CODE HERE ###
Z3 = np.dot(W3, A2) + b3
A3 = sigmoid(Z3)
cache = (Z1, D1, A1, W1, b1, Z2, D2, A2, W2, b2, Z3, A3, W3, b3)
return A3, cache
X_assess, parameters = forward_propagation_with_dropout_test_case()
A3, cache = forward_propagation_with_dropout(X_assess, parameters, keep_prob = 0.7)
print ("A3 = " + str(A3))
```
**Expected Output**:
<table>
<tr>
<td>
**A3**
</td>
<td>
[[ 0.36974721 0.00305176 0.04565099 0.49683389 0.36974721]]
</td>
</tr>
</table>
### 3.2 - Backward propagation with dropout
**Exercise**: Implement the backward propagation with dropout. As before, you are training a 3 layer network. Add dropout to the first and second hidden layers, using the masks $D^{[1]}$ and $D^{[2]}$ stored in the cache.
**Instruction**:
Backpropagation with dropout is actually quite easy. You will have to carry out 2 Steps:
1. You had previously shut down some neurons during forward propagation, by applying a mask $D^{[1]}$ to `A1`. In backpropagation, you will have to shut down the same neurons, by reapplying the same mask $D^{[1]}$ to `dA1`.
2. During forward propagation, you had divided `A1` by `keep_prob`. In backpropagation, you'll therefore have to divide `dA1` by `keep_prob` again (the calculus interpretation is that if $A^{[1]}$ is scaled by `keep_prob`, then its derivative $dA^{[1]}$ is also scaled by the same `keep_prob`).
```
# GRADED FUNCTION: backward_propagation_with_dropout
def backward_propagation_with_dropout(X, Y, cache, keep_prob):
"""
Implements the backward propagation of our baseline model to which we added dropout.
Arguments:
X -- input dataset, of shape (2, number of examples)
Y -- "true" labels vector, of shape (output size, number of examples)
cache -- cache output from forward_propagation_with_dropout()
keep_prob - probability of keeping a neuron active during drop-out, scalar
Returns:
gradients -- A dictionary with the gradients with respect to each parameter, activation and pre-activation variables
"""
m = X.shape[1]
(Z1, D1, A1, W1, b1, Z2, D2, A2, W2, b2, Z3, A3, W3, b3) = cache
dZ3 = A3 - Y
dW3 = 1./m * np.dot(dZ3, A2.T)
db3 = 1./m * np.sum(dZ3, axis=1, keepdims = True)
dA2 = np.dot(W3.T, dZ3)
### START CODE HERE ### (≈ 2 lines of code)
dA2 = None # Step 1: Apply mask D2 to shut down the same neurons as during the forward propagation
dA2 = None # Step 2: Scale the value of neurons that haven't been shut down
### END CODE HERE ###
dZ2 = np.multiply(dA2, np.int64(A2 > 0))
dW2 = 1./m * np.dot(dZ2, A1.T)
db2 = 1./m * np.sum(dZ2, axis=1, keepdims = True)
dA1 = np.dot(W2.T, dZ2)
### START CODE HERE ### (≈ 2 lines of code)
dA1 = None # Step 1: Apply mask D1 to shut down the same neurons as during the forward propagation
dA1 = None # Step 2: Scale the value of neurons that haven't been shut down
### END CODE HERE ###
dZ1 = np.multiply(dA1, np.int64(A1 > 0))
dW1 = 1./m * np.dot(dZ1, X.T)
db1 = 1./m * np.sum(dZ1, axis=1, keepdims = True)
gradients = {"dZ3": dZ3, "dW3": dW3, "db3": db3,"dA2": dA2,
"dZ2": dZ2, "dW2": dW2, "db2": db2, "dA1": dA1,
"dZ1": dZ1, "dW1": dW1, "db1": db1}
return gradients
X_assess, Y_assess, cache = backward_propagation_with_dropout_test_case()
gradients = backward_propagation_with_dropout(X_assess, Y_assess, cache, keep_prob = 0.8)
print ("dA1 = " + str(gradients["dA1"]))
print ("dA2 = " + str(gradients["dA2"]))
```
**Expected Output**:
<table>
<tr>
<td>
**dA1**
</td>
<td>
[[ 0.36544439 0. -0.00188233 0. -0.17408748]
[ 0.65515713 0. -0.00337459 0. -0. ]]
</td>
</tr>
<tr>
<td>
**dA2**
</td>
<td>
[[ 0.58180856 0. -0.00299679 0. -0.27715731]
[ 0. 0.53159854 -0. 0.53159854 -0.34089673]
[ 0. 0. -0.00292733 0. -0. ]]
</td>
</tr>
</table>
Let's now run the model with dropout (`keep_prob = 0.86`). It means at every iteration you shut down each neurons of layer 1 and 2 with 14% probability. The function `model()` will now call:
- `forward_propagation_with_dropout` instead of `forward_propagation`.
- `backward_propagation_with_dropout` instead of `backward_propagation`.
```
parameters = model(train_X, train_Y, keep_prob = 0.86, learning_rate = 0.3)
print ("On the train set:")
predictions_train = predict(train_X, train_Y, parameters)
print ("On the test set:")
predictions_test = predict(test_X, test_Y, parameters)
```
Dropout works great! The test accuracy has increased again (to 95%)! Your model is not overfitting the training set and does a great job on the test set. The French football team will be forever grateful to you!
Run the code below to plot the decision boundary.
```
plt.title("Model with dropout")
axes = plt.gca()
axes.set_xlim([-0.75,0.40])
axes.set_ylim([-0.75,0.65])
plot_decision_boundary(lambda x: predict_dec(parameters, x.T), train_X, train_Y)
```
**Note**:
- A **common mistake** when using dropout is to use it both in training and testing. You should use dropout (randomly eliminate nodes) only in training.
- Deep learning frameworks like [tensorflow](https://www.tensorflow.org/api_docs/python/tf/nn/dropout), [PaddlePaddle](http://doc.paddlepaddle.org/release_doc/0.9.0/doc/ui/api/trainer_config_helpers/attrs.html), [keras](https://keras.io/layers/core/#dropout) or [caffe](http://caffe.berkeleyvision.org/tutorial/layers/dropout.html) come with a dropout layer implementation. Don't stress - you will soon learn some of these frameworks.
<font color='blue'>
**What you should remember about dropout:**
- Dropout is a regularization technique.
- You only use dropout during training. Don't use dropout (randomly eliminate nodes) during test time.
- Apply dropout both during forward and backward propagation.
- During training time, divide each dropout layer by keep_prob to keep the same expected value for the activations. For example, if keep_prob is 0.5, then we will on average shut down half the nodes, so the output will be scaled by 0.5 since only the remaining half are contributing to the solution. Dividing by 0.5 is equivalent to multiplying by 2. Hence, the output now has the same expected value. You can check that this works even when keep_prob is other values than 0.5.
## 4 - Conclusions
**Here are the results of our three models**:
<table>
<tr>
<td>
**model**
</td>
<td>
**train accuracy**
</td>
<td>
**test accuracy**
</td>
</tr>
<td>
3-layer NN without regularization
</td>
<td>
95%
</td>
<td>
91.5%
</td>
<tr>
<td>
3-layer NN with L2-regularization
</td>
<td>
94%
</td>
<td>
93%
</td>
</tr>
<tr>
<td>
3-layer NN with dropout
</td>
<td>
93%
</td>
<td>
95%
</td>
</tr>
</table>
Note that regularization hurts training set performance! This is because it limits the ability of the network to overfit to the training set. But since it ultimately gives better test accuracy, it is helping your system.
Congratulations for finishing this assignment! And also for revolutionizing French football. :-)
<font color='blue'>
**What we want you to remember from this notebook**:
- Regularization will help you reduce overfitting.
- Regularization will drive your weights to lower values.
- L2 regularization and Dropout are two very effective regularization techniques.
| github_jupyter |
# Using hgvs
This notebook demonstrates major features of the hgvs package.
```
import hgvs
hgvs.__version__
```
## Variant I/O
### Initialize the parser
```
# You only need to do this once per process
import hgvs.parser
hp = hgvsparser = hgvs.parser.Parser()
```
### Parse a simple variant
```
v = hp.parse_hgvs_variant("NC_000007.13:g.21726874G>A")
v
v.ac, v.type
v.posedit
v.posedit.pos
v.posedit.pos.start
```
### Parsing complex variants
```
v = hp.parse_hgvs_variant("NM_003777.3:c.13552_*36del57")
v.posedit.pos.start, v.posedit.pos.end
v.posedit.edit
```
### Formatting variants
All objects may be formatted simply by "stringifying" or printing them using `str`, `print()`, or `"".format()`.
```
str(v)
print(v)
"{v} spans the CDS end".format(v=v)
```
## Projecting variants between sequences
### Set up a dataprovider
Mapping variants requires exon structures, alignments, CDS bounds, and raw sequence. These are provided by a `hgvs.dataprovider` instance. The only dataprovider provided with hgvs uses UTA. You may write your own by subsclassing hgvs.dataproviders.interface.
```
import hgvs.dataproviders.uta
hdp = hgvs.dataproviders.uta.connect()
```
### Initialize mapper classes
The VariantMapper class projects variants between two sequence accessions using alignments from a specified source. In order to use it, you must know that two sequences are aligned. VariantMapper isn't demonstrated here.
AssemblyMapper builds on VariantMapper and handles identifying appropriate sequences. It is configured for a particular genome assembly.
```
import hgvs.variantmapper
#vm = variantmapper = hgvs.variantmapper.VariantMapper(hdp)
am37 = easyvariantmapper = hgvs.variantmapper.AssemblyMapper(hdp, assembly_name='GRCh37')
am38 = easyvariantmapper = hgvs.variantmapper.AssemblyMapper(hdp, assembly_name='GRCh38')
```
### c_to_g
This is the easiest case because there is typically only one alignment between a transcript and the genome. (Exceptions exist for pseudoautosomal regions.)
```
var_c = hp.parse_hgvs_variant("NM_015120.4:c.35G>C")
var_g = am37.c_to_g(var_c)
var_g
am38.c_to_g(var_c)
```
### g_to_c
In order to project a genomic variant onto a transcript, you must tell the AssemblyMapper which transcript to use.
```
am37.relevant_transcripts(var_g)
am37.g_to_c(var_g, "NM_015120.4")
```
### c_to_p
```
var_p = am37.c_to_p(var_c)
str(var_p)
var_p.posedit.uncertain = False
str(var_p)
```
### Projecting in the presence of a genome-transcript gap
As of Oct 2016, 1033 RefSeq transcripts in 433 genes have gapped alignments. These gaps require special handlingin order to maintain the correspondence of positions in an alignment. hgvs uses the precomputed alignments in UTA to correctly project variants in exons containing gapped alignments.
This example demonstrates projecting variants in the presence of a gap in the alignment of NM_015120.4 (ALMS1) with GRCh37 chromosome 2. (The alignment with GRCh38 is similarly gapped.) Specifically, the adjacent genomic positions 73613031 and 73613032 correspond to the non-adjacent CDS positions 35 and 39.
```
NM_015120.4 c 15 > > 58
NM_015120.4 n 126 > CCGGGCGAGCTGGAGGAGGAGGAG > 169
||||||||||| |||||||||| 21=3I20=
NC_000002.11 g 73613021 > CCGGGCGAGCT---GGAGGAGGAG > 73613041
NC_000002.11 g 73613021 < GGCCCGCTCGA---CCTCCTCCTC < 73613041
```
```
str(am37.c_to_g(hp.parse_hgvs_variant("NM_015120.4:c.35G>C")))
str(am37.c_to_g(hp.parse_hgvs_variant("NM_015120.4:c.39G>C")))
```
## Normalizing variants
In hgvs, normalization means shifting variants 3' (as requried by the HGVS nomenclature) as well as rewriting variants. The variant "NM_001166478.1:c.30_31insT" is in a poly-T run (on the transcript). It should be shifted 3' and is better written as dup, as shown below:
```
* NC_000006.11:g.49917127dupA
NC_000006.11 g 49917117 > AGAAAGAAAAATAAAACAAAG > 49917137
NC_000006.11 g 49917117 < TCTTTCTTTTTATTTTGTTTC < 49917137
||||||||||||||||||||| 21=
NM_001166478.1 n 41 < TCTTTCTTTTTATTTTGTTTC < 21 NM_001166478.1:n.35dupT
NM_001166478.1 c 41 < < 21 NM_001166478.1:c.30_31insT
```
```
import hgvs.normalizer
hn = hgvs.normalizer.Normalizer(hdp)
v = hp.parse_hgvs_variant("NM_001166478.1:c.30_31insT")
str(hn.normalize(v))
```
## A more complex normalization example
This example is based on https://github.com/biocommons/hgvs/issues/382/.
```
NC_000001.11 g 27552104 > CTTCACACGCATCCTGACCTTG > 27552125
NC_000001.11 g 27552104 < GAAGTGTGCGTAGGACTGGAAC < 27552125
|||||||||||||||||||||| 22=
NM_001029882.3 n 843 < GAAGTGTGCGTAGGACTGGAAC < 822
NM_001029882.3 c 12 < < -10
^^
NM_001029882.3:c.1_2del
NM_001029882.3:n.832_833delAT
NC_000001.11:g.27552114_27552115delAT
```
```
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.1A>G"))
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.2T>G"))
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.1_2del"))
```
The genomic coordinates for the SNVs at c.1 and c.2 match those for the del at c.1_2. Good!
Now, notice what happens with c.1_3del, c.1_4del, and c.1_5del:
```
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.1_3del"))
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.1_4del"))
am38.c_to_g(hp.parse_hgvs_variant("NM_001029882.3:c.1_5del"))
```
Explanation:
On the transcript, c.1_2delAT deletes AT from …AGGATGCG…, resulting in …AGGGCG…. There's no ambiguity about what sequence was actually deleted.
c.1_3delATG deletes ATG, resulting in …AGGCG…. Note that you could also get this result by deleting GAT. This is an example of an indel that is subject to normalization and hgvs does this.
c.1_4delATGC and 1_5delATGCG have similar behaviors.
Normalization is always 3' with respect to the reference sequence. So, after projecting from a - strand transcript to the genome, normalization will go in the opposite direction to the transcript. It will have roughly the same effect as being 5' shifted on the transcript (but revcomp'd).
For more precise control, see the `normalize` and `replace_reference` options of `AssemblyMapper`.
## Validating variants
`hgvs.validator.Validator` is a composite of two classes, `hgvs.validator.IntrinsicValidator` and `hgvs.validator.ExtrinsicValidator`. Intrinsic validation evaluates a given variant for *internal* consistency, such as requiring that insertions specify adjacent positions. Extrinsic validation evaluates a variant using external data, such as ensuring that the reference nucleotide in the variant matches that implied by the reference sequence and position. Validation returns `True` if successful, and raises an exception otherwise.
```
import hgvs.validator
hv = hgvs.validator.Validator(hdp)
hv.validate(hp.parse_hgvs_variant("NM_001166478.1:c.30_31insT"))
from hgvs.exceptions import HGVSError
try:
hv.validate(hp.parse_hgvs_variant("NM_001166478.1:c.30_32insT"))
except HGVSError as e:
print(e)
```
| github_jupyter |
## Preliminaries
```
# Load libraries
import numpy as np
from keras.datasets import imdb
from keras.preprocessing.text import Tokenizer
from keras import models
from keras import layers
from keras.callbacks import EarlyStopping, ModelCheckpoint
# Set random seed
np.random.seed(0)
```
## Load Movie Review Text Data
```
# Set the number of features we want
number_of_features = 1000
# Load data and target vector from movie review data
(train_data, train_target), (test_data, test_target) = imdb.load_data(num_words=number_of_features)
# Convert movie review data to a one-hot encoded feature matrix
tokenizer = Tokenizer(num_words=number_of_features)
train_features = tokenizer.sequences_to_matrix(train_data, mode='binary')
test_features = tokenizer.sequences_to_matrix(test_data, mode='binary')
```
## Create Neural Network Architecture
```
# Start neural network
network = models.Sequential()
# Add fully connected layer with a ReLU activation function
network.add(layers.Dense(units=16, activation='relu', input_shape=(number_of_features,)))
# Add fully connected layer with a ReLU activation function
network.add(layers.Dense(units=16, activation='relu'))
# Add fully connected layer with a sigmoid activation function
network.add(layers.Dense(units=1, activation='sigmoid'))
```
## Compile Neural Network
```
# Compile neural network
network.compile(loss='binary_crossentropy', # Cross-entropy
optimizer='rmsprop', # Root Mean Square Propagation
metrics=['accuracy']) # Accuracy performance metric
```
## Setup Early Stopping
In Keras, we can implement early stopping as a callback function. Callbacks are functions that can be applied at certain stages of the training process, such as at the end of each epoch. Specifically, in our solution, we included `EarlyStopping(monitor='val_loss', patience=2)` to define that we wanted to monitor the test (validation) loss at each epoch and after the test loss has not improved after two epochs, training is interrupted. However, since we set `patience=2`, we won't get the best model, but the model two epochs after the best model. Therefore, optionally, we can include a second operation, `ModelCheckpoint` which saves the model to a file after every checkpoint (which can be useful in case a multi-day training session is interrupted for some reason. Helpful for us, if we set `save_best_only=True` then `ModelCheckpoint` will only save the best model.
```
# Set callback functions to early stop training and save the best model so far
callbacks = [EarlyStopping(monitor='val_loss', patience=2),
ModelCheckpoint(filepath='best_model.h5', monitor='val_loss', save_best_only=True)]
```
## Train Neural Network
```
# Train neural network
history = network.fit(train_features, # Features
train_target, # Target vector
epochs=20, # Number of epochs
callbacks=callbacks, # Early stopping
verbose=0, # Print description after each epoch
batch_size=100, # Number of observations per batch
validation_data=(test_features, test_target)) # Data for evaluation
```
| github_jupyter |
# 囚徒困境演化博弈分析
姓名:邹子涵
学号:202020085400139
<p style="text-indent:2em"></p>
## 一、囚徒困境
<p style="text-indent:2em">囚徒困境(prisoner's dilemma)是指两个被捕的囚徒之间的一种特殊博弈,说明为什么甚至在合作对双方都有利时,保持合作也是困难的。囚徒困境是博弈论的非零和博弈中具代表性的例子,反映个人最佳选择并非团体最佳选择。虽然困境本身只属模型性质,但现实中的价格竞争、环境保护、人际关系等方面,也会频繁出现类似情况。</p>
### 1.1、理论起源
<p style="text-indent:2em">囚徒困境的故事讲的是,两个嫌疑犯作案后被警察抓住,分别关在不同的屋子里接受审讯。警察知道两人有罪,但缺乏足够的证据。警察告诉每个人:如果两人都抵赖,各判刑一年;如果两人都坦白,各判八年;如果两人中一个坦白而另一个抵赖,坦白的放出去,抵赖的判十年。于是,每个囚徒都面临两种选择:坦白或抵赖。然而,不管同伙选择什么,每个囚徒的最优选择是坦白:如果同伙抵赖、自己坦白的话放出去,抵赖的话判十年,坦白比不坦白好;如果同伙坦白、自己坦白的话判八年,比起抵赖的判十年,坦白还是比抵赖的好。</p>

<p style="text-indent:2em">结果,两个嫌疑犯都选择坦白,各判刑八年。如果两人都抵赖,各判一年,显然这个结果好。囚徒困境所反映出的深刻问题是,人类的个人理性有时能导致集体的非理性--聪明的人类会因自己的聪明而作茧自缚,或者损害集体的利益。</p>
## 二、python实现沙盒模式下囚徒困境演化过程
### 2.1、过程构思
在沙盒模式下,初始化数个拥有不同策略、初始个数相同的人群,是他们在沙盒中随机进行博弈数轮,每轮结束后淘汰一定数量的人数,并在博弈过程中加入随机变异。
### 2.2、代码解读
#### 2.2.1、导包并初始化囚徒选择
将BETRAY(背叛)定义为False
COOPERATE(合作)定义为True
```
import random
from collections import Counter
import matplotlib.pyplot as plt
import sys
import inspect
BETRAY = True
COOPERATE = False
```
工具函数takeSecond(elem),用于获取列表的第二个元素
```
def takeSecond(elem):
return elem[1]
```
#### 2.2.2、定义策略母类
在策略母类中定义了以下几个方法:
+ get_score:得分方法,当自身得分长度大于50时。将得分进行累加
+ __init__:初始化方法,包含三个参数:my_history/theirhistory/score,分别用来记录自身最后的选择、上一博弈对手的选择以及得分情况
+ play:博弈方法
+ reinit_history:重新初始化,重置自身最后的选择和上一博弈对手的选择
```
class Alg:
def get_score(self):
if len(self.score)>50:
self.score = [sum(self.score)]
return self.score
def __init__(self):
self.my_history = []
self.their_history = []
self.score = []
def play(self):
""" @param my_history List of own last choices
@param their_history List of last choices of the other
@return decision {BETRAY, SILENT}
"""
raise NotImplementedError
def __repr__(self):
return "{}".format(type(self).__name__)
def reinit_history(self):
self.my_history = []
self.their_history = []
return self
```
#### 2.2.3、定义分组方法
该方法的目标是将列表ls中的人群按照size大小进行切分分组。
```
def partition(ls, size):
"""
Returns a new list with elements
of which is a list of certain size.
>>> partition([1, 2, 3, 4], 3)
[[1, 2, 3], [4]]
"""
return [ls[i:i + size] for i in range(0, len(ls), size)]
```
#### 2.2.4、定义囚徒困境的收益矩阵
这里通过将囚徒困境的收益矩阵给改为正值,方便计算并比较得分情况,更改后的囚徒困境收益矩阵为下图所示:

```
def score(A_choice, B_choice):
if A_choice == B_choice:
if A_choice == BETRAY:
return (1, 1)
return (3, 3)
if A_choice == BETRAY:
return (5, 0)
return (0, 5)
```
#### 2.2.5、定义博弈过程
A与B博弈,将自己的选择记录在my_history中,将对方的选择记录在their_history中,并将A、B的得分记录在score中。
```
def play(A, B):
A_score = 0
B_score = 0
A_choice = A.play(A.my_history, B.my_history)
B_choice = B.play(B.my_history, A.my_history)
A_score_, B_score_ = score(A_choice, B_choice)
A.my_history.append(A_choice)
B.my_history.append(B_choice)
A.their_history.append(B_choice)
B.their_history.append(A_choice)
A.score.append(A_score_)
B.score.append(B_score_)
return (A_score_, B_score_)
```
#### 2.2.6、定义沙盒模式
在沙盒模式中会将所有人进行两两分组————partition(persons, 2),persons为参与博弈的所有人群对象,rounds为博弈轮数。
在每一轮中,对分好组的两人进行一次博弈
```
# 一期博弈过程,即所有样本随机两两匹配后博弈rounds次
def simulate(persons, rounds, seed=0):
"""
:param persons: 所有参与人对象列表
:param strategy_score: 不同策略得分,初始均为0
:param seed:
:param rounds: 当前期博弈次数
:return: strategy_score
"""
random.shuffle(persons)
# print (persons)
persons_pairs = partition(persons, 2)
# print(partition(persons, 2))
for i in range(rounds):
# tmp = []
for pair in persons_pairs:
play(pair[0], pair[1])
```
#### 2.2.7、沙盒世界更新
在每轮博弈完成后,会对沙盒世界中的人群对象进行更新,包括策略变更、淘汰和生成新的样本(新加入的玩家)
```
def self_reproduction(persons, mutation_rate, out_rate):
"""
:param persons: 博弈后样本列表
:param mutation_rate: 变异率
:param out_rate: 淘汰率
:return: 自我繁殖后样本列表
"""
mutation_count = int(len(persons) * mutation_rate)
out_count = int(len(persons) * out_rate)
for person in persons:
score_list = [sum(person.get_score()) for person in persons]
persons.sort(key=lambda x: sum(x.get_score()), reverse=True)
# print (persons_sorted)
score_sorted = [sum(person.get_score()) for person in persons]
# 获取淘汰样本及变异样本
out_persons = persons[-out_count:]
random.shuffle(out_persons)
mutation_persons = out_persons[:mutation_count]
# 淘汰样本,切分列表
persons = persons[:-out_count]
# 针对淘汰样本中未变异的生成新样本,针对变异样本随机生成新样本后my_history及their_history传递
out2reinit = [eval(person.__repr__() + "()") for person in persons[:out_count - mutation_count]]
for person in out2reinit:
person.my_history = []
person.their_history = []
out2mutation = []
for person in mutation_persons:
algs = Alg.__subclasses__()
new_person = random.choice(algs)()
out2mutation.append(new_person)
# 新旧样本组成persons
print("out2reinit", out2reinit)
print("out2mutation", out2mutation)
print ("persons", persons)
persons = persons + out2reinit + out2mutation
persons = [person.reinit_history() for person in persons]
return persons
```
#### 2.2.8、定义不同的策略
背叛策略,该策略的样本会永远选择背叛
```
class Betray(Alg):
def play(self, my_history, their_history):
return BETRAY
```
合作策略,该策略的样本会永远选择合作
```
class Cooperate(Alg):
def play(self, my_history, their_history):
return COOPERATE
```
报复策略,该策略的样本一开始会选择合作并在后续的选择中倾向于合作,但是如果上一次博弈对手选择了背叛,那么这一次则会选择背叛
```
class Tit4Tat(Alg):
def play(self, my_history, their_history):
if len(their_history) <= 1:
return COOPERATE
if their_history[-1] == BETRAY:
return BETRAY
return COOPERATE
```
混沌随机策略,该策略的样本会以1/3的概率选择背叛,以2/3的概率选择合作
```
class RandomKind(Alg):
def play(self, my_history, their_history):
if random.random() < 0.667:
return COOPERATE
return BETRAY
```
秩序随机策略,该策略的样本会以1/3的概率选择合作,以2/3的概率选择背叛
```
class RandomNasty(Alg):
def play(self, my_history, their_history):
if random.random() < 0.667:
return BETRAY
return COOPERATE
```
理智策略,该策略的样本回一句对手的选择做出改变,如果对手有超过半数的选择选择背叛,那么自己将会选择背叛,否则将选择合作
```
class Dirk(Alg):
def play(self, my_history, their_history):
cnt_betray = len([x for x in their_history if x == BETRAY])
if cnt_betray > 0.5 * len(their_history):
return BETRAY
return COOPERATE
```
秩序规律策略,该策略的样本以三次博弈为循环,每轮循环中前两次博弈选择合作,第三次选择背叛
```
class PerKind(Alg):
def play(self, my_history, their_history):
m = len(their_history) % 3
if m == 0 or m == 1:
return COOPERATE
return BETRAY
```
混沌规律策略,该策略的样本以三次博弈为循环,每轮循环中前两次博弈选择背叛,第三次选择合作
```
class PerNasty(Alg):
def play(self, my_history, their_history):
m = len(their_history) % 3
if m == 0 or m == 1:
return BETRAY
return COOPERATE
```
叛逆策略,该策略的样本会在首次选择背叛,第二次会依据对方的第一次选择做出相反的选择,在往后的选择会依据对手最后两次的选择做出抉择,一旦有背叛行为,将会选择合作
```
class Theresa(Alg):
def play(self, my_history, their_history):
if len(their_history) == 0:
return BETRAY
if len(their_history) == 1:
if their_history[-1] == BETRAY:
return COOPERATE
return BETRAY
if their_history[-1] == BETRAY or their_history[-2] == BETRAY:
return COOPERATE
return BETRAY
```
奇偶策略,该策略的样本会在单数次选择背叛、双数次选择合作
```
class Alternate(Alg):
def play(self, my_history, their_history):
m = len(their_history) % 2
if m == 0:
return BETRAY
return COOPERATE
```
复杂策略,过于复杂。
```
class WinStayLoseShift(Alg):
def play(self, my_history, their_history):
# print ("their_history", their_history)
# print ("my_history", my_history)
if len(their_history) == 0:
return COOPERATE
if len(my_history) == 0:
return COOPERATE
if my_history[-1] == COOPERATE and their_history[-1] == COOPERATE:
return COOPERATE
if my_history[-1] == COOPERATE and their_history[-1] == BETRAY:
return BETRAY
if my_history[-1] == BETRAY and their_history[-1] == COOPERATE:
return BETRAY
return COOPERATE
```
#### 2.2.9、生成策略样本
依据传入的参数counts对每个策略生成counts人数
```
# 生成策略样本
def creat_persons(counts):
""" @param counts 每类策略样本生成个数
@return List[person1,person2,...]
"""
inst = []
for i in range(counts):
examples = Alg.__subclasses__()
inst += [example() for example in examples]
# print(inst)
return inst
```
### 2.3、绘图并分析
这里的超级参数选择如下:演化代数为10代,每代进行20期,每期进行10次重复博弈,每种策略的初始样本数量为50人,每代都会进行策略变异、淘汰样本及更新新样本,淘汰率为0.2,变异概率为0.05。
#### 2.3.1、绘制沙盒最终策略人群饼状图
将最终沙盒中的不同策略人群按照比例绘制饼状图,传入参数为persons_strategy: 各期样本分布列表;i: 绘制期数。
```
def plot_persons_of_strategy(persons_strategy, i):
"""
绘制期末样本分布饼图
:param persons_strategy: 各期样本分布列表
:param i: 绘制期数
:return:
"""
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文标签
labels = persons_strategy[i - 1].keys()
sizes = persons_strategy[i - 1].values()
# explode = (0, 0, 0, 0.1, 0, 0)
plt.pie(sizes, labels=labels, autopct='%1.1f%%', shadow=False, startangle=150)
plt.title("第" + str(i) + "期样本分布(淘汰更新前)")
plt.show()
return
```

#### 2.3.2、输出沙盒最终各策略的得分平均数
传入参数为persons_score:各期策略平均分数;i:绘制期数。
```
def plot_persons_of_score(persons_score, i):
"""
绘制期末策略得分平均数
:param persons_score:各期策略平均分数
:param i:绘制期数
:return:
"""
dict = persons_score[i - 1]
# print(dict)
dict2 = sorted(zip(dict.values(), dict.keys()), reverse=True)
# print (dict2)
plt.show()
return
```
#### 2.3.3、绘制博弈过程不同策略样本的数量
由于每代会进行淘汰和更新,因此每代的策略样本数量都会发生变化,用折线图绘制出策略样本数量的变化
```
def plot_persons_of_counts(persons_strategy):
labels = persons_strategy[0].keys()
dict = {key: [] for key in labels}
for d in persons_strategy:
for key in d:
dict[key].append(d[key])
print (dict)
names = list(range(0, len(persons_strategy)))
x = range(len(names))
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文标签
colorList = ["#ed1299", "#09f9f5", "#246b93", "#cc8e12", "#d561dd", "#c93f00", "#ddd53e",
"#4aef7b", "#e86502", "#9ed84e", "#39ba30", "#6ad157", "#8249aa", "#99db27", "#e07233", "#ff523f",
"#ce2523", "#f7aa5d", "#cebb10", "#03827f", "#931635", "#373bbf", "#a1ce4c", "#ef3bb6", "#d66551",
]
i = 0
for key in dict:
plt.plot(x, dict[key], c=colorList[i], label="$"+key+"$")
i += 1
plt.legend(loc='center left', bbox_to_anchor=(0, 1.05), ncol=5) # 让图例生效
plt.xticks(x, names, rotation=1)
plt.margins(0.01)
plt.subplots_adjust(bottom=0.10)
plt.xlabel('代数') # X轴标签
plt.ylabel("策略样本数") # Y轴标签
plt.show()
return
```

#### 2.3.4、绘制样本选择分布情况
```
def plot_persons_of_choice(persons_choice, i):
plt.rcParams['font.sans-serif'] = ['SimHei'] # 用来正常显示中文标签
labels = persons_choice[i - 1].keys()
sizes = persons_choice[i - 1].values()
# explode = (0, 0, 0, 0.1, 0, 0)
plt.pie(sizes, labels=labels, autopct='%1.1f%%', shadow=False, startangle=150)
plt.title("第" + str(i) + "代样本选择分布")
plt.show()
return
```

#### 2.3.5、启动沙盒博弈过程主函数
```
if __name__ == '__main__':
# Num_of_generation 演化代数
Num_of_generation = 10
# Num_of_periods 每代期数
Num_of_periods = 20
# Play_num_of_per_period 每期重复博弈次数
Play_num_of_per_period = 10
# Num_of_per_strategy 每种策略样本初始数
Num_of_per_strategy = 50
# Out_rate 淘汰率
Out_rate = 0.2
# Mutation_rate 变异率
Mutation_rate = 0.05
persons_strategy = []
persons_score = []
persons_choice = []
# 生成样本
persons = creat_persons(Num_of_per_strategy)
# 开始博弈
for j in range(Num_of_generation):
for i in range(Num_of_periods):
print (j,"代,", i,"期")
simulate(persons, Play_num_of_per_period, seed=0)
# for person in persons:
# print (person.__repr__(), "score", sum(person.get_score()), "\n", person.my_history, "\n", person.their_history)
# break
# 计算期末策略分布
persons_strategy.append(Counter([person.__repr__() for person in persons]))
# 计算期末策略得分均值
strategy_name = Counter([person.__repr__() for person in persons]).keys()
tmp = dict.fromkeys(strategy_name, 0)
for person in persons:
tmp[person.__repr__()] += sum(person.get_score())
for key in tmp:
tmp[key] = tmp[key] / persons_strategy[j][key]
persons_score.append(tmp)
# 计算期末样本选择分布
choice_list = []
for person in persons:
choice_list += person.my_history
persons_choice.append(Counter(choice_list))
persons = self_reproduction(persons, Mutation_rate, Out_rate)
plot_persons_of_choice(persons_choice, Num_of_generation)
plot_persons_of_strategy(persons_strategy, Num_of_generation)
plot_persons_of_counts(persons_strategy)
plot_persons_of_score(persons_score, Num_of_generation)
```
| github_jupyter |
<a href="https://colab.research.google.com/github/rjrahul24/ai-with-python-series/blob/main/05.%20Build%20Concrete%20Time%20Series%20Forecasts/Colab%20Notebook/TimeSeries_Validation_ModelBuilding.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# Importing the basic preprocessing packages
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# The series module from Pandas will help in creating a time series
from pandas import Series,DataFrame
import seaborn as sns
%matplotlib inline
# About the Data Set (Location: https://www.kaggle.com/sumanthvrao/daily-climate-time-series-data)
# To forecast the daily climate of a city in India
time_series = pd.read_csv('https://raw.githubusercontent.com/rjrahul24/ai-with-python-series/main/05.%20Build%20Concrete%20Time%20Series%20Forecasts/Data/DailyDelhiClimateTrain.csv', parse_dates=['date'], index_col='date')
time_series.head()
# Below are a few statistical methods on time series that will help in understanding the data patterns
# Plotting all the individual columns to observe the pattern of data in each column
time_series.plot(subplots=True)
# Calculating the mean, maximum values, and minimum of all individual columns of the dataset
time_series.mean()
time_series.max()
time_series.min()
# The describe() method gives information like count, mean, deviations and quartiles of all columns
time_series.describe()
timeseries_mm = time_series['wind_speed']
timeseries_mm.plot(style='g--')
plt.show()
# Resampling the dataset using the Mean() resample method
timeseries_mm = time_series['wind_speed'].resample("A").mean()
timeseries_mm.plot(style='g--')
plt.show()
# Calculating the rolling mean with a 14-bracket window between time intervals
time_series['wind_speed'].rolling(window=14, center=False).mean().plot(style='-g')
plt.show()
# In the coming sections we will implement time series forecasting on the same dataset
# The series module from Pandas will help in creating a time series
from pandas import Series,DataFrame
%matplotlib inline
# Statsmodel and Adfuller will help in testing the stationarity of the time series
import statsmodels
from statsmodels.tsa.stattools import adfuller
time_series_train = pd.read_csv('https://raw.githubusercontent.com/rjrahul24/ai-with-python-series/main/05.%20Build%20Concrete%20Time%20Series%20Forecasts/Data/DailyDelhiClimateTrain.csv', parse_dates=True)
time_series_train["date"] = pd.to_datetime(time_series_train["date"])
time_series_train.date.freq ="D"
time_series_train.set_index("date", inplace=True)
time_series_train.columns
# Decomposing the time series with Statsmodels Decompose Method
from statsmodels.tsa.seasonal import seasonal_decompose
sd_1 = seasonal_decompose(time_series_train["meantemp"])
sd_2 = seasonal_decompose(time_series_train["humidity"])
sd_3 = seasonal_decompose(time_series_train["wind_speed"])
sd_4 = seasonal_decompose(time_series_train["meanpressure"])
sd_1.plot()
sd_2.plot()
sd_3.plot()
sd_4.plot()
# From the above graph’s observations, it looks like everything other than meanpressure is already stationary
# To re-confirm stationarity, we will run all columns through the ad-fuller test
adfuller(time_series_train["meantemp"])
adfuller(time_series_train["humidity"])
adfuller(time_series_train["wind_speed"])
adfuller(time_series_train["meanpressure"])
# Consolidate the ad-fuller tests to test from static data
temp_var = time_series_train.columns
print('significance level : 0.05')
for var in temp_var:
ad_full = adfuller(time_series_train[var])
print(f'For {var}')
print(f'Test static {ad_full[1]}',end='\n \n')
# With the ad-fuller test, we can now conclude that all data is stationary since static tests are below significance levels. This also rejects the hypothesis that meanpressure was non-static.
# Let us now move towards training and validating the prediction model
from statsmodels.tsa.vector_ar.var_model import VAR
train_model = VAR(time_series_train)
fit_model = train_model.fit(6)
# AIC is lower for lag_order 6. Hence, we can assume the lag_order of 6.
fix_train_test = time_series_train.dropna()
order_lag_a = fit_model.k_ar
X = fix_train_test[:-order_lag_a]
Y = fix_train_test[-order_lag_a:]
# Model Validation
validate_y = X.values[-order_lag_a:]
forcast_val = fit_model.forecast(validate_y,steps=order_lag_a)
train_forecast = DataFrame(forcast_val,index=time_series_train.index[-order_lag_a:],columns=Y.columns)
train_forecast
# Check performance of the predictions’ model
from sklearn.metrics import mean_absolute_error
for i in time_series_train.columns:
print(f'MAE of {i} is {mean_absolute_error(Y[[i]],train_forecast[[i]])}')
# Humidity and Meanpressure are showing higher errors of forecast. We could assume that certain external factors are causing this.
# This model, therefore, forecasts wind speed and mean temperature accurately with less than 5% error
# Let us now implement this on the test data and forecast for the next 6 future periods of time
test_forecast = pd.read_csv('https://raw.githubusercontent.com/rjrahul24/ai-with-python-series/main/05.%20Build%20Concrete%20Time%20Series%20Forecasts/Data/DailyDelhiClimateTrain.csv',parse_dates=['date'], index_col='date')
period_range = pd.date_range('2017-01-05',periods=6)
order_lag_b = fit_model.k_ar
X1,Y1 = test_forecast[1:-order_lag_b],test_forecast[-order_lag_b:]
input_val = Y1.values[-order_lag_b:]
data_forecast = fit_model.forecast(input_val,steps=order_lag_b)
df_forecast = DataFrame(data_forecast,columns=X1.columns,index=period_range)
df_forecast
# Plotting the test data with auto correlation
from statsmodels.graphics.tsaplots import plot_acf
# The next 6 periods of mean temperature (graph 1) and wind_speed (graph 2)
plot_acf(df_forecast["meantemp"])
plot_acf(df_forecast["wind_speed"])
# Import Granger Causality module from the statsmodels package and use the Chi-Squared test metric
from statsmodels.tsa.stattools import grangercausalitytests
test_var = time_series.columns
lag_max = 12
test_type = 'ssr_chi2test'
causal_val = DataFrame(np.zeros((len(test_var),len(test_var))),columns=test_var,index=test_var)
for a in test_var:
for b in test_var:
c = grangercausalitytests ( time_series [ [b,a] ], maxlag = lag_max, verbose = False)
pred_val = [round ( c [ i +1 ] [0] [test_type] [1], 5 ) for i in range (lag_max) ]
min_value = np.min (pred_val)
causal_val.loc[b,a] = min_value
causal_val
pip install pmdarima
```
Let us now use the PMDARIMA Methodology to implement Predictions for the Wind Speed Values within our Time Series
```
from pmdarima.arima import auto_arima
from pmdarima.arima import ADFTest
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.arima_model import ARIMA
adf_test=ADFTest(alpha=0.05)
adf_test.should_diff(time_series_train["wind_speed"])
df = time_series_train["wind_speed"]
df.shape
train=df[:1300]
test=df[-250:]
plt.figure(figsize=(15,10))
plt.plot(train)
plt.plot(test)
plt.autoscale()
# Reducing size of data to run easily on Colab. Please run the entire dataset on a system with very high GPU Performance.
train=df[:100]
test=df[-30:]
model=auto_arima(train,start_p=0,d=1,start_q=0,
max_p=5,max_d=5,max_q=5, start_P=0,
D=1, start_Q=0, max_P=5,max_D=5,
max_Q=5, m=12, seasonal=True,
error_action='warn',trace=True,
supress_warnings=True,stepwise=True,
random_state=20,n_fits=50)
model.summary()
prediction = pd.DataFrame(model.predict(n_periods = 30),index=test.index)
prediction.columns = ['wind_Speed_preds']
prediction
```
| github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork/Template/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
<h1>Convolutional Neural Network with Small Images</h1>
<h3>Objective for this Notebook<h3>
<h5> 1. Learn how to use a Convolutional Neural Network to classify handwritten digits from the MNIST database</h5>
<h5> 2. Learn hot to reshape the images to make them faster to process </h5>
<h2>Table of Contents</h2>
<p>In this lab, we will use a Convolutional Neural Network to classify handwritten digits from the MNIST database. We will reshape the images to make them faster to process </p>
<ul>
<li><a href="https://#Makeup_Data">Get Some Data</a></li>
<li><a href="https://#CNN">Convolutional Neural Network</a></li>
<li><a href="https://#Train">Define Softmax, Criterion function, Optimizer and Train the Model</a></li>
<li><a href="https://#Result">Analyze Results</a></li>
</ul>
<p>Estimated Time Needed: <strong>25 min</strong> 14 min to train model </p>
<hr>
<h2>Preparation</h2>
```
# Import the libraries we need to use in this lab
# Using the following line code to install the torchvision library
# !mamba install -y torchvision
!pip install torchvision==0.9.1 torch==1.8.1
import torch
import torch.nn as nn
import torchvision.transforms as transforms
import torchvision.datasets as dsets
import matplotlib.pylab as plt
import numpy as np
```
Define the function <code>plot_channels</code> to plot out the kernel parameters of each channel
```
# Define the function for plotting the channels
def plot_channels(W):
n_out = W.shape[0]
n_in = W.shape[1]
w_min = W.min().item()
w_max = W.max().item()
fig, axes = plt.subplots(n_out, n_in)
fig.subplots_adjust(hspace=0.1)
out_index = 0
in_index = 0
#plot outputs as rows inputs as columns
for ax in axes.flat:
if in_index > n_in-1:
out_index = out_index + 1
in_index = 0
ax.imshow(W[out_index, in_index, :, :], vmin=w_min, vmax=w_max, cmap='seismic')
ax.set_yticklabels([])
ax.set_xticklabels([])
in_index = in_index + 1
plt.show()
```
Define the function <code>plot_parameters</code> to plot out the kernel parameters of each channel with Multiple outputs .
```
# Define the function for plotting the parameters
def plot_parameters(W, number_rows=1, name="", i=0):
W = W.data[:, i, :, :]
n_filters = W.shape[0]
w_min = W.min().item()
w_max = W.max().item()
fig, axes = plt.subplots(number_rows, n_filters // number_rows)
fig.subplots_adjust(hspace=0.4)
for i, ax in enumerate(axes.flat):
if i < n_filters:
# Set the label for the sub-plot.
ax.set_xlabel("kernel:{0}".format(i + 1))
# Plot the image.
ax.imshow(W[i, :], vmin=w_min, vmax=w_max, cmap='seismic')
ax.set_xticks([])
ax.set_yticks([])
plt.suptitle(name, fontsize=10)
plt.show()
```
Define the function <code>plot_activation</code> to plot out the activations of the Convolutional layers
```
# Define the function for plotting the activations
def plot_activations(A, number_rows=1, name="", i=0):
A = A[0, :, :, :].detach().numpy()
n_activations = A.shape[0]
A_min = A.min().item()
A_max = A.max().item()
fig, axes = plt.subplots(number_rows, n_activations // number_rows)
fig.subplots_adjust(hspace = 0.4)
for i, ax in enumerate(axes.flat):
if i < n_activations:
# Set the label for the sub-plot.
ax.set_xlabel("activation:{0}".format(i + 1))
# Plot the image.
ax.imshow(A[i, :], vmin=A_min, vmax=A_max, cmap='seismic')
ax.set_xticks([])
ax.set_yticks([])
plt.show()
```
Define the function <code>show_data</code> to plot out data samples as images.
```
def show_data(data_sample):
plt.imshow(data_sample[0].numpy().reshape(IMAGE_SIZE, IMAGE_SIZE), cmap='gray')
plt.title('y = '+ str(data_sample[1]))
```
<!--Empty Space for separating topics-->
<h2 id="Makeup_Data">Get the Data</h2>
we create a transform to resize the image and convert it to a tensor .
```
IMAGE_SIZE = 16
composed = transforms.Compose([transforms.Resize((IMAGE_SIZE, IMAGE_SIZE)), transforms.ToTensor()])
```
Load the training dataset by setting the parameters <code>train </code> to <code>True</code>. We use the transform defined above.
```
train_dataset = dsets.MNIST(root='./data', train=True, download=True, transform=composed)
```
Load the testing dataset by setting the parameters train <code>False</code>.
```
# Make the validating
validation_dataset = dsets.MNIST(root='./data', train=False, download=True, transform=composed)
```
We can see the data type is long.
```
# Show the data type for each element in dataset
type(train_dataset[0][1])
```
Each element in the rectangular tensor corresponds to a number representing a pixel intensity as demonstrated by the following image.
<img src="https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0110EN/notebook_images%20/chapter%206/6.2.1imagenet.png" width="550" alt="MNIST data image">
Print out the fourth label
```
# The label for the fourth data element
train_dataset[3][1]
```
Plot the fourth sample
```
# The image for the fourth data element
show_data(train_dataset[3])
```
The fourth sample is a "1".
<!--Empty Space for separating topics-->
<h2 id="CNN">Build a Convolutional Neural Network Class</h2>
Build a Convolutional Network class with two Convolutional layers and one fully connected layer. Pre-determine the size of the final output matrix. The parameters in the constructor are the number of output channels for the first and second layer.
```
class CNN(nn.Module):
# Contructor
def __init__(self, out_1=16, out_2=32):
super(CNN, self).__init__()
self.cnn1 = nn.Conv2d(in_channels=1, out_channels=out_1, kernel_size=5, padding=2)
self.maxpool1=nn.MaxPool2d(kernel_size=2)
self.cnn2 = nn.Conv2d(in_channels=out_1, out_channels=out_2, kernel_size=5, stride=1, padding=2)
self.maxpool2=nn.MaxPool2d(kernel_size=2)
self.fc1 = nn.Linear(out_2 * 4 * 4, 10)
# Prediction
def forward(self, x):
x = self.cnn1(x)
x = torch.relu(x)
x = self.maxpool1(x)
x = self.cnn2(x)
x = torch.relu(x)
x = self.maxpool2(x)
x = x.view(x.size(0), -1)
x = self.fc1(x)
return x
# Outputs in each steps
def activations(self, x):
#outputs activation this is not necessary
z1 = self.cnn1(x)
a1 = torch.relu(z1)
out = self.maxpool1(a1)
z2 = self.cnn2(out)
a2 = torch.relu(z2)
out1 = self.maxpool2(a2)
out = out.view(out.size(0),-1)
return z1, a1, z2, a2, out1,out
```
<h2 id="Train">Define the Convolutional Neural Network Classifier, Criterion function, Optimizer and Train the Model</h2>
There are 16 output channels for the first layer, and 32 output channels for the second layer
```
# Create the model object using CNN class
model = CNN(out_1=16, out_2=32)
```
Plot the model parameters for the kernels before training the kernels. The kernels are initialized randomly.
```
# Plot the parameters
plot_parameters(model.state_dict()['cnn1.weight'], number_rows=4, name="1st layer kernels before training ")
plot_parameters(model.state_dict()['cnn2.weight'], number_rows=4, name='2nd layer kernels before training' )
```
Define the loss function, the optimizer and the dataset loader
```
criterion = nn.CrossEntropyLoss()
learning_rate = 0.1
optimizer = torch.optim.SGD(model.parameters(), lr = learning_rate)
train_loader = torch.utils.data.DataLoader(dataset=train_dataset, batch_size=100)
validation_loader = torch.utils.data.DataLoader(dataset=validation_dataset, batch_size=5000)
```
Train the model and determine validation accuracy technically test accuracy **(This may take a long time)**
```
# Train the model
n_epochs=3
cost_list=[]
accuracy_list=[]
N_test=len(validation_dataset)
COST=0
def train_model(n_epochs):
for epoch in range(n_epochs):
COST=0
for x, y in train_loader:
optimizer.zero_grad()
z = model(x)
loss = criterion(z, y)
loss.backward()
optimizer.step()
COST+=loss.data
cost_list.append(COST)
correct=0
#perform a prediction on the validation data
for x_test, y_test in validation_loader:
z = model(x_test)
_, yhat = torch.max(z.data, 1)
correct += (yhat == y_test).sum().item()
accuracy = correct / N_test
accuracy_list.append(accuracy)
train_model(n_epochs)
```
<!--Empty Space for separating topics-->
<h2 id="Result">Analyze Results</h2>
Plot the loss and accuracy on the validation data:
```
# Plot the loss and accuracy
fig, ax1 = plt.subplots()
color = 'tab:red'
ax1.plot(cost_list, color=color)
ax1.set_xlabel('epoch', color=color)
ax1.set_ylabel('Cost', color=color)
ax1.tick_params(axis='y', color=color)
ax2 = ax1.twinx()
color = 'tab:blue'
ax2.set_ylabel('accuracy', color=color)
ax2.set_xlabel('epoch', color=color)
ax2.plot( accuracy_list, color=color)
ax2.tick_params(axis='y', color=color)
fig.tight_layout()
```
View the results of the parameters for the Convolutional layers
```
# Plot the channels
plot_channels(model.state_dict()['cnn1.weight'])
plot_channels(model.state_dict()['cnn2.weight'])
```
Consider the following sample
```
# Show the second image
show_data(train_dataset[1])
```
Determine the activations
```
# Use the CNN activations class to see the steps
out = model.activations(train_dataset[1][0].view(1, 1, IMAGE_SIZE, IMAGE_SIZE))
```
Plot out the first set of activations
```
# Plot the outputs after the first CNN
plot_activations(out[0], number_rows=4, name="Output after the 1st CNN")
```
The image below is the result after applying the relu activation function
```
# Plot the outputs after the first Relu
plot_activations(out[1], number_rows=4, name="Output after the 1st Relu")
```
The image below is the result of the activation map after the second output layer.
```
# Plot the outputs after the second CNN
plot_activations(out[2], number_rows=32 // 4, name="Output after the 2nd CNN")
```
The image below is the result of the activation map after applying the second relu
```
# Plot the outputs after the second Relu
plot_activations(out[3], number_rows=4, name="Output after the 2nd Relu")
```
We can see the result for the third sample
```
# Show the third image
show_data(train_dataset[2])
# Use the CNN activations class to see the steps
out = model.activations(train_dataset[2][0].view(1, 1, IMAGE_SIZE, IMAGE_SIZE))
# Plot the outputs after the first CNN
plot_activations(out[0], number_rows=4, name="Output after the 1st CNN")
# Plot the outputs after the first Relu
plot_activations(out[1], number_rows=4, name="Output after the 1st Relu")
# Plot the outputs after the second CNN
plot_activations(out[2], number_rows=32 // 4, name="Output after the 2nd CNN")
# Plot the outputs after the second Relu
plot_activations(out[3], number_rows=4, name="Output after the 2nd Relu")
```
Plot the first five mis-classified samples:
```
# Plot the mis-classified samples
count = 0
for x, y in torch.utils.data.DataLoader(dataset=validation_dataset, batch_size=1):
z = model(x)
_, yhat = torch.max(z, 1)
if yhat != y:
show_data((x, y))
plt.show()
print("yhat: ",yhat)
count += 1
if count >= 5:
break
```
<a href="https://dataplatform.cloud.ibm.com/registration/stepone?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01&context=cpdaas&apps=data_science_experience%2Cwatson_machine_learning"><img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-DL0110EN-SkillsNetwork/Template/module%201/images/Watson_Studio.png"/></a>
<!--Empty Space for separating topics-->
<h2>About the Authors:</h2>
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Joseph Santarcangelo</a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
Other contributors: <a href="https://www.linkedin.com/in/michelleccarey/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Michelle Carey</a>, <a href="https://www.linkedin.com/in/jiahui-mavis-zhou-a4537814a?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Mavis Zhou</a>
Thanks to Magnus <a href="http://www.hvass-labs.org/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkDL0110ENSkillsNetwork20647811-2021-01-01">Erik Hvass Pedersen</a> whose tutorials helped me understand convolutional Neural Network
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ----------------------------------------------------------- |
| 2020-09-23 | 2.0 | Srishti | Migrated Lab to Markdown and added to course repo in GitLab |
<hr>
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
from relation_modeling_utils import load_data
train_df = load_data("data/atomic2020_data-feb2021/train.tsv", multi_label=True)
val_df = load_data("data/atomic2020_data-feb2021/dev.tsv", multi_label=True)
test_df = load_data("data/atomic2020_data-feb2021/test.tsv", multi_label=True)
len(train_df), len(val_df), len(test_df)
train_df.head()
from relation_modeling_utils import explode_labels
train_df, test_df = explode_labels(train_df), explode_labels(test_df)
test_df.label_0.value_counts(), test_df.label_1.value_counts(), test_df.label_2.value_counts()
```
## Original data lexical overlap
```
from relation_modeling_utils import create_vocab
train_vocab, val_vocab, test_vocab = create_vocab(train_df), create_vocab(val_df), create_vocab(test_df)
```
### Lexical overlap with stopwords
```
train_test_overlap = set(train_vocab).intersection(set(test_vocab))
len(train_test_overlap) / len(train_vocab), len(train_test_overlap) / len(test_vocab)
train_val_overlap = set(train_vocab).intersection(set(val_vocab))
len(train_val_overlap) / len(train_vocab), len(train_val_overlap) / len(val_vocab)
```
### Lexical overlap without stopwords
```
from spacy.lang.en.stop_words import STOP_WORDS
train_vocab_nostp = create_vocab(train_df, include_stopwords=False)
val_vocab_nostp = create_vocab(val_df, include_stopwords=False)
test_vocab_nostp = create_vocab(test_df, include_stopwords=False)
train_test_overlap_nostp = set(train_vocab_nostp).intersection(set(test_vocab_nostp))
len(train_test_overlap_nostp) / len(train_vocab_nostp), len(train_test_overlap_nostp) / len(test_vocab_nostp)
train_val_overlap_nostp = set(train_vocab_nostp).intersection(set(val_vocab_nostp))
len(train_val_overlap_nostp) / len(train_vocab_nostp), len(train_val_overlap_nostp) / len(val_vocab_nostp)
```
## Create new ATOMIC datasets
```
import pandas as pd
atomic_df = pd.concat([train_df, test_df])
len(atomic_df)
```
### Handle duplicates
```
atomic_df.duplicated(subset=["text"]).sum()
from relation_modeling_utils import explode_labels
atomic_df = explode_labels(atomic_df)
atomic_df.head()
atomic_df.duplicated(subset=["text", "label_0", "label_1", "label_2"]).sum()
exact_dup_df = atomic_df[atomic_df.duplicated(subset=["text", "label_0", "label_1", "label_2"])]
train_org_df = pd.read_csv("data/atomic2020_data-feb2021/train.tsv", sep="\t", header=None, names=["head", "relation", "tail"])
test_org_df = pd.read_csv("data/atomic2020_data-feb2021/test.tsv", sep="\t", header=None, names=["head", "relation", "tail"])
train_org_df.head()
dup_dfs = []
for text in exact_dup_df.text:
train_ex = train_org_df[train_org_df["head"] == text]
test_ex = test_org_df[test_org_df["head"] == text]
merged_df = train_ex.merge(test_ex, left_on=["head", "relation"], right_on=["head", "relation"], how="inner")
if len(merged_df) > 0:
dup_dfs.append(merged_df)
len(dup_dfs)
exact_dup_merged_df = pd.concat(dup_dfs)
exact_dup_merged_df = exact_dup_merged_df.rename(columns={"tail_x": "tail_train", "tail_y": "tail_test"})
exact_dup_merged_df.head()
exact_dup_merged_df.drop_duplicates(subset=["head", "relation"]).to_csv("data/exact_duplicates.tsv", sep="\t")
atomic_df = atomic_df.drop_duplicates(subset=["text", "label_0", "label_1", "label_2"])
duplicated_df = atomic_df[atomic_df.duplicated(subset=["text"])]
len(duplicated_df)
from kogito.core.relation import PHYSICAL_RELATIONS, EVENT_RELATIONS, SOCIAL_RELATIONS
def get_relation_classes(rels):
classes = []
if len(rels & set([str(rel) for rel in PHYSICAL_RELATIONS])) > 0:
classes.append("P")
if len(rels & set([str(rel) for rel in EVENT_RELATIONS])) > 0:
classes.append("E")
if len(rels & set([str(rel) for rel in SOCIAL_RELATIONS])) > 0:
classes.append("S")
return classes
soft_dup_ex = []
for text in duplicated_df.text:
train_ex = train_org_df[train_org_df["head"] == text]
test_ex = test_org_df[test_org_df["head"] == text]
train_rels = set(train_ex.relation)
test_rels = set(test_ex.relation)
if len(train_rels & test_rels) == 0:
soft_dup_ex.append((text, get_relation_classes(train_rels), get_relation_classes(test_rels)))
len(soft_dup_ex)
soft_dup_df = pd.DataFrame(soft_dup_ex, columns=["head", "rel_class_train", "rel_class_test"])
soft_dup_df.head()
soft_dup_df.to_csv("data/soft_duplicates.tsv", sep="\t")
atomic_df[atomic_df.text == "PersonX forgets PersonX's lines"]
train_df[train_df.text == "PersonX forgets PersonX's lines"]
test_df[test_df.text == "PersonX forgets PersonX's lines"]
atomic_df.duplicated(subset=["text", "label_0"]).sum()
all_duplicate_df = atomic_df[atomic_df.duplicated(subset=["text"], keep=False)]
len(all_duplicate_df)
all_duplicate_df.head()
import numpy as np
def group_duplicate_heads(subdf):
label_s = np.logical_or(*[np.array(l) for l in subdf.label]).astype(int).tolist()
label0_s = np.logical_or(*subdf.label_0.to_list()).astype(int)
label1_s = np.logical_or(*subdf.label_1.to_list()).astype(int)
label2_s = np.logical_or(*subdf.label_2.to_list()).astype(int)
return pd.Series({"label": label_s, "label_0": label0_s, "label_1": label1_s, "label_2": label2_s})
handled_dup_df = all_duplicate_df.groupby("text").apply(group_duplicate_heads).reset_index()
handled_dup_df.head()
len(atomic_df)
atomic_df = atomic_df.drop_duplicates(subset=["text"], keep=False)
len(atomic_df)
atomic_df = pd.concat([atomic_df, handled_dup_df])
len(atomic_df)
atomic_df.duplicated(subset=["text"]).sum()
```
### Create docs out of heads
```
import spacy
from tqdm import tqdm
from relation_modeling_utils import IGNORE_WORDS, create_vocab, get_doc
from spacy.lang.en.stop_words import STOP_WORDS
def make_docs(data, vocab, include_stopwords=True):
nlp = spacy.load("en_core_web_sm", exclude=["ner"])
docs = []
for row in tqdm(data.itertuples(), total=len(data)):
doc = get_doc(nlp, row.text)
words = set()
for token in doc:
if token.text not in IGNORE_WORDS and (include_stopwords or token.text not in STOP_WORDS):
words.add(token.lemma_)
doc.user_data['words'] = words
doc.user_data['label'] = row.label
docs.append(doc)
for doc in docs:
freqs = 0
for word in doc.user_data['words']:
freqs += max(vocab.get(word, 0) - 1, 0)
doc.user_data['relative_freq'] = freqs
return docs
atomic_vocab = create_vocab(atomic_df)
atomic_docs = make_docs(atomic_df, atomic_vocab, include_stopwords=False)
sorted(atomic_vocab.items(), key=lambda i: i[1])[-5:]
class1_docs = [doc for doc in atomic_docs if doc.user_data['label'][0] == 1]
class2_docs = [doc for doc in atomic_docs if doc.user_data['label'][1] == 1]
class3_docs = [doc for doc in atomic_docs if doc.user_data['label'][2] == 1]
FREQUENCY_THRESHOLD = 5
class1_freq1_docs = [doc for doc in class1_docs if doc.user_data['relative_freq'] < 1][:500]
class2_freq1_docs = [doc for doc in class2_docs if doc.user_data['relative_freq'] < FREQUENCY_THRESHOLD][:500]
class3_freq1_docs = [doc for doc in class3_docs if doc.user_data['relative_freq'] < FREQUENCY_THRESHOLD][:500]
len(class1_freq1_docs), len(class2_freq1_docs), len(class3_freq1_docs)
test_samples = [doc.text for doc in class1_freq1_docs+class2_freq1_docs+class3_freq1_docs]
len(test_samples)
new_train_data, new_test_data = [], []
for row in atomic_df.itertuples():
if row.text in test_samples:
new_test_data.append((row.text, row.label))
else:
new_train_data.append((row.text, row.label))
new_train_df = pd.DataFrame(new_train_data, columns=["text", "label"])
new_test_df = pd.DataFrame(new_test_data, columns=["text", "label"])
len(new_train_df), len(new_test_df)
from relation_modeling_utils import create_vocab
new_train_vocab, new_test_vocab = create_vocab(new_train_df), create_vocab(new_test_df)
```
### New lexical overlap with stopwords
```
new_train_test_overlap = set(new_train_vocab).intersection(set(new_test_vocab))
len(new_train_test_overlap) / len(new_train_vocab), len(new_train_test_overlap) / len(new_test_vocab)
```
### New lexical overlap without stopwords
```
new_train_vocab_nostp = create_vocab(new_train_df, include_stopwords=False)
new_test_vocab_nostp = create_vocab(new_test_df, include_stopwords=False)
new_train_test_overlap_nostp = set(new_train_vocab_nostp).intersection(set(new_test_vocab_nostp))
len(new_train_test_overlap_nostp) / len(new_train_vocab_nostp), len(new_train_test_overlap_nostp) / len(new_test_vocab_nostp)
```
### Class distributions
```
from relation_modeling_utils import explode_labels
new_train_df, new_test_df = explode_labels(new_train_df), explode_labels(new_test_df)
new_test_df.label_0.value_counts(), new_test_df.label_1.value_counts(), new_test_df.label_2.value_counts()
from relation_modeling_utils import get_class_dist_report
get_class_dist_report(new_test_df)
```
### Vocabulary info
```
new_train_df.to_csv("data/atomic_ood2/n5/train_n5.csv")
new_test_df.to_csv("data/atomic_ood2/n5/test_n5.csv")
from relation_modeling_utils import load_fdata, create_vocab
from spacy.lang.en.stop_words import STOP_WORDS
def get_vocab_info(dataset_type):
train_f = load_fdata(f"data/atomic_ood2/{dataset_type}/train_{dataset_type}.csv")
test_f = load_fdata(f"data/atomic_ood2/{dataset_type}/test_{dataset_type}.csv")
train_f_vocab, test_f_vocab = create_vocab(train_f), create_vocab(test_f)
train_f_nostp, test_f_nostp = create_vocab(train_f,include_stopwords=False), create_vocab(test_f, include_stopwords=False)
return {
'train': len(train_f_vocab), 'test': len(test_f_vocab),
'train_nostp': len(train_f_nostp), 'test_nostp': len(test_f_nostp)
}
get_vocab_info("n1")
get_vocab_info("n3")
get_vocab_info("n5")
```
| github_jupyter |
# Facies Classification Solution By Team_BGC
Cheolkyun Jeong and Ping Zhang From Team_BGC
## Import Header
```
##### import basic function
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
##### import stuff from scikit learn
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.model_selection import KFold, cross_val_score,LeavePGroupsOut, LeaveOneGroupOut, cross_val_predict
from sklearn.metrics import confusion_matrix, make_scorer, f1_score, accuracy_score, recall_score, precision_score
```
## 1. Data Prepocessing
1) Filtered data preparation
After the initial data validation, we figure out the NM_M input is a key differentiator to group non-marine stones (sandstone, c_siltstone, and f_siltstone) and marine stones (marine_silt_shale, mudstone, wakestone, dolomite, packstone, and bafflestone) in the current field. Our team decides to use this classifier aggressively and prepare a filtered dataset which cleans up the outliers.
```
# Input file paths
facies_vector_path = 'facies_vectors.csv'
train_path = 'training_data.csv'
test_path = 'validation_data_nofacies.csv'
# Read training data to dataframe
#training_data = pd.read_csv(train_path)
```
Using Full data to train
```
# 1=sandstone 2=c_siltstone 3=f_siltstone # 4=marine_silt_shale
#5=mudstone 6=wackestone 7=dolomite 8=packstone 9=bafflestone
facies_colors = ['#F4D03F', '#F5B041', '#DC7633','#A569BD',
'#000000', '#000080', '#2E86C1', '#AED6F1', '#196F3D']
feature_names = ['GR', 'ILD_log10', 'DeltaPHI', 'PHIND', 'PE', 'NM_M', 'RELPOS']
facies_labels = ['SS', 'CSiS', 'FSiS', 'SiSh', 'MS',
'WS', 'D','PS', 'BS']
#facies_color_map is a dictionary that maps facies labels
#to their respective colors
training_data = pd.read_csv(facies_vector_path)
X = training_data[feature_names].values
y = training_data['Facies'].values
well = training_data['Well Name'].values
depth = training_data['Depth'].values
facies_color_map = {}
for ind, label in enumerate(facies_labels):
facies_color_map[label] = facies_colors[ind]
def label_facies(row, labels):
return labels[ row['Facies'] -1]
training_data.loc[:,'FaciesLabels'] = training_data.apply(lambda row: label_facies(row, facies_labels), axis=1)
training_data.describe()
# Fitering out some outliers
j = []
for i in range(len(training_data)):
if ((training_data['NM_M'].values[i]==2)and ((training_data['Facies'].values[i]==1)or(training_data['Facies'].values[i]==2)or(training_data['Facies'].values[i]==3))):
j.append(i)
elif((training_data['NM_M'].values[i]==1)and((training_data['Facies'].values[i]!=1)and(training_data['Facies'].values[i]!=2)and(training_data['Facies'].values[i]!=3))):
j.append(i)
training_data_filtered = training_data.drop(training_data.index[j])
print(np.shape(training_data_filtered))
```
Add Missing PE by following AR4 Team
```
#X = training_data_filtered[feature_names].values
# Testing without filtering
X = training_data[feature_names].values
reg = RandomForestRegressor(max_features='sqrt', n_estimators=50)
# DataImpAll = training_data_filtered[feature_names].copy()
DataImpAll = training_data[feature_names].copy()
DataImp = DataImpAll.dropna(axis = 0, inplace=False)
Ximp=DataImp.loc[:, DataImp.columns != 'PE']
Yimp=DataImp.loc[:, 'PE']
reg.fit(Ximp, Yimp)
X[np.array(DataImpAll.PE.isnull()),4] = reg.predict(DataImpAll.loc[DataImpAll.PE.isnull(),:].drop('PE',axis=1,inplace=False))
```
## 2. Feature Selection
Log Plot of Facies
### Filtered Data
```
#count the number of unique entries for each facies, sort them by
#facies number (instead of by number of entries)
#facies_counts_filtered = training_data_filtered['Facies'].value_counts().sort_index()
facies_counts = training_data['Facies'].value_counts().sort_index()
#use facies labels to index each count
#facies_counts_filtered.index = facies_labels
facies_counts.index = facies_labels
#facies_counts_filtered.plot(kind='bar',color=facies_colors,
# title='Distribution of Filtered Training Data by Facies')
facies_counts.plot(kind='bar',color=facies_colors,
title='Distribution of Filtered Training Data by Facies')
#facies_counts_filtered
#training_data_filtered.columns
#facies_counts_filtered
training_data.columns
facies_counts
```
### Filtered facies
```
correct_facies_labels = training_data['Facies'].values
feature_vectors = training_data.drop(['Formation', 'Well Name', 'Depth','Facies','FaciesLabels'], axis=1)
```
Normailization
```
from sklearn import preprocessing
from sklearn.cross_validation import train_test_split
scaler = preprocessing.StandardScaler().fit(X)
scaled_features = scaler.transform(X)
```
## 3. Prediction Model
Accuracy
```
def accuracy(conf):
total_correct = 0.
nb_classes = conf.shape[0]
for i in np.arange(0,nb_classes):
total_correct += conf[i][i]
acc = total_correct/sum(sum(conf))
return acc
adjacent_facies = np.array([[1], [0,2], [1], [4], [3,5], [4,6,7], [5,7], [5,6,8], [6,7]])
def accuracy_adjacent(conf, adjacent_facies):
nb_classes = conf.shape[0]
total_correct = 0.
for i in np.arange(0,nb_classes):
total_correct += conf[i][i]
for j in adjacent_facies[i]:
total_correct += conf[i][j]
return total_correct / sum(sum(conf))
```
Augment Features
```
# HouMath Team algorithm
# Feature windows concatenation function
def augment_features_window(X, N_neig):
# Parameters
N_row = X.shape[0]
N_feat = X.shape[1]
# Zero padding
X = np.vstack((np.zeros((N_neig, N_feat)), X, (np.zeros((N_neig, N_feat)))))
# Loop over windows
X_aug = np.zeros((N_row, N_feat*(2*N_neig+1)))
for r in np.arange(N_row)+N_neig:
this_row = []
for c in np.arange(-N_neig,N_neig+1):
this_row = np.hstack((this_row, X[r+c]))
X_aug[r-N_neig] = this_row
return X_aug
# HouMath Team algorithm
# Feature gradient computation function
def augment_features_gradient(X, depth):
# Compute features gradient
d_diff = np.diff(depth).reshape((-1, 1))
d_diff[d_diff==0] = 0.001
X_diff = np.diff(X, axis=0)
X_grad = X_diff / d_diff
# Compensate for last missing value
X_grad = np.concatenate((X_grad, np.zeros((1, X_grad.shape[1]))))
return X_grad
# HouMath Team algorithm
# Feature augmentation function
def augment_features(X, well, depth, N_neig=1):
# Augment features
X_aug = np.zeros((X.shape[0], X.shape[1]*(N_neig*2+2)))
for w in np.unique(well):
w_idx = np.where(well == w)[0]
X_aug_win = augment_features_window(X[w_idx, :], N_neig)
X_aug_grad = augment_features_gradient(X[w_idx, :], depth[w_idx])
X_aug[w_idx, :] = np.concatenate((X_aug_win, X_aug_grad), axis=1)
# Find padded rows
padded_rows = np.unique(np.where(X_aug[:, 0:7] == np.zeros((1, 7)))[0])
return X_aug, padded_rows
X_aug, padded_rows = augment_features(scaled_features, well, depth)
X_aug.shape
from sklearn.cross_validation import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X_aug, y, test_size=0.3, random_state=16)
X_train_full, X_test_zero, y_train_full, y_test_full = train_test_split(X_aug, y, test_size=0.0, random_state=42)
X_train_full.shape
```
SVM
```
from sklearn.model_selection import KFold, cross_val_score,LeavePGroupsOut, LeaveOneGroupOut, cross_val_predict
from classification_utilities import display_cm, display_adj_cm
clf_filtered = svm.SVC(C=10, gamma=1)
clf_filtered.fit(X_train, y_train)
#predicted_labels_filtered = clf_filtered.predict(X_test_filtered)
predicted_labels = clf_filtered.predict(X_test)
cv_conf_svm = confusion_matrix(y_test, predicted_labels)
print('Optimized facies classification accuracy = %.2f' % accuracy(cv_conf_svm))
print('Optimized adjacent facies classification accuracy = %.2f' % accuracy_adjacent(cv_conf_svm, adjacent_facies))
display_cm(cv_conf_svm, facies_labels,display_metrics=True, hide_zeros=True)
```
## 4. Result Analysis
Prepare test data
```
well_data = pd.read_csv('validation_data_nofacies.csv')
well_data['Well Name'] = well_data['Well Name'].astype('category')
well_features = well_data.drop(['Formation', 'Well Name', 'Depth'], axis=1)
# Prepare test data
well_ts = well_data['Well Name'].values
depth_ts = well_data['Depth'].values
X_ts = well_data[feature_names].values
X_ts = scaler.transform(X_ts)
# Augment features
X_ts, padded_rows = augment_features(X_ts, well_ts, depth_ts)
# Using all data and optimize parameter to train the data
clf_filtered = svm.SVC(C=10, gamma=1)
clf_filtered.fit(X_train_full, y_train_full)
#clf_filtered.fit(X_train_filtered, y_train_filtered)
y_pred = clf_filtered.predict(X_ts)
well_data['Facies'] = y_pred
well_data
well_data.to_csv('predict_result_svm_full_data.csv')
```
## 5. Using Tensorflow
Filtered Data Model
```
X_train.shape
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
# Specify that all features have real-value data
# feature_columns_filtered = [tf.contrib.layers.real_valued_column("", dimension=7)]
feature_columns_filtered = [tf.contrib.layers.real_valued_column("", dimension=28)]
# Build DNN
classifier_filtered = tf.contrib.learn.DNNClassifier(feature_columns=feature_columns_filtered,
hidden_units=[14, 14, 14, 10],
n_classes=10)
classifier_filtered.fit(x=X_train,y=y_train,steps=5000)
y_predict = []
predictions = classifier_filtered.predict(x=X_test)
for i, p in enumerate(predictions):
y_predict.append(p)
#print("Index %s: Prediction - %s, Real - %s" % (i + 1, p, y_test_filtered[i]))
# Evaluate accuracy.
accuracy_score = classifier_filtered.evaluate(x=X_test, y=y_test)["accuracy"]
print('Accuracy: {0:f}'.format(accuracy_score))
cv_conf_dnn = confusion_matrix(y_test, y_predict)
print('Optimized facies classification accuracy = %.2f' % accuracy(cv_conf_dnn))
print('Optimized adjacent facies classification accuracy = %.2f' % accuracy_adjacent(cv_conf_dnn, adjacent_facies))
display_cm(cv_conf_dnn, facies_labels,display_metrics=True, hide_zeros=True)
```
Result from DNN
```
classifier_filtered.fit(x=X_train_full,
y=y_train_full,
steps=10000)
predictions = classifier_filtered.predict(X_ts)
y_predict_filtered = []
for i, p in enumerate(predictions):
y_predict_filtered.append(p)
well_data['Facies'] = y_predict_filtered
well_data
well_data.to_csv('predict_result_dnn_full_data.csv')
```
Find best model by setting different parameters
```
best_score = 0
best_cv = cv_conf_filtered
best_arch = []
tmp_score = 0
tmp_cv = cv_conf_filtered
# try 3 layers and each architecture with 10 iterations training
for i in range (7, 14):
for j in range(14, 21):
for k in range(10, 14):
for l in range(10):
dim_num = 3
arch = [i,j,k]
s_num = 3000
feature_columns_filtered = [tf.contrib.layers.real_valued_column("", dimension=dim_num)]
dnn = tf.contrib.learn.DNNClassifier(feature_columns=feature_columns_filtered,
hidden_units=arch,
n_classes=10)
tmp_score,tmp_cv = dnn_prediction(dim_num, arch, s_num, dnn)
if(tmp_score>best_score):
best_score = tmp_score
best_cv = tmp_cv
best_arch = arch
print('Best facies classification accuracy = %.2f' % accuracy(best_cv))
print('Best adjacent facies classification accuracy = %.2f' % accuracy_adjacent(best_cv, adjacent_facies))
print('Best arch is ', best_arch)
display_cm(cv_conf_dnn, facies_labels,display_metrics=True, hide_zeros=True)
#export_dir_path = 'd:\\'
#classifier_filtered.export(export_dir_path)
```
| github_jupyter |
```
# create bp for 2 ROI into csv - using panda dataframe
import mne
import matplotlib.pyplot as plt
from mne.time_frequency import psd_array_multitaper
from scipy.integrate import simps
import numpy as np
import seaborn as sns
import pandas as pd
precleaned_epochs_path = '/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/'
fmin = 1
fmax = 100
# all subject 63
subjs=['02', '04','07', '11', '12', '14', '16', '18', '19', '21', '22', '26', '28', '30',
'32', '34', '36', '37', '38', '40', '42', '50', '51', '52', '53', '54', '55', '56',
'58', '59','60', '63', '65', '67', '68', '70', '72', '73', '78', '83', '87', '88',
'90', '91', '93', '94', '95', '96','10','25','29','39','57','64','69','80','81','82',
'35','71','79','76','77']
listNovices = ['02', '04', '07', '10', '11', '12', '14', '16', '18', '19', '21', '22', '26',
'28', '29', '30', '32', '34', '35', '36', '37', '38','39', '40', '42', '81', '82',
'83', '87', '88', '90', '91', '93', '94', '95', '96']
listExperts = ['25', '50', '51', '52',
'53', '54', '55', '56', '57', '58', '59', '60', '63', '64', '65', '67','68', '69' ,'70' ,
'71', '72', '73', '76', '77', '78' ,'79', '80']
ROIs = {'frontal':['Fz','F1','F3','AF3','AFz','AF4','F4','F2'],
'occipital':['POz','Oz','O1','O2']}
CondStates={'baseline':{'VD':['111.0','112.0'],'FA':['211.0','212.0'],'OP':['311.0','312.0']},
'safe':{'VD':['121.0','122.0'],'FA':['221.0','222.0'],'OP':['321.0','322.0']},
'threat':{'VD':['131.0','132.0'],'FA':['231.0','232.0'],'OP':['331.0','332.0']}}
wavebands = {'alpha':[8,12],'theta':[3,7],'beta':[13,24],'lowG':[25,40],'highG':[60,90]}
frames_subject = []
for subj in subjs:
precleaned_epochs_fname = precleaned_epochs_path + 'subj0'+subj+'full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
frames_condition = []
for condition in CondStates.keys():
frames_state=[]
for state in CondStates[condition].keys():
# precleaned_epochs_engineering = precleaned_epochs.copy()
precleaned_epochs_selected = precleaned_epochs[CondStates[condition][state]]
frames_ROI=[]
for roi in ROIs.keys():
precleaned_epochs_engineering = precleaned_epochs_selected.copy()
data = precleaned_epochs_engineering.pick_channels(ROIs[roi])._data
frames_waveband = []
for waveband in wavebands.keys():
low,high = wavebands[waveband]
bpAbs_4Epochs=[]
bpRelative_4Epochs=[]
for num_epochs in range(data.shape[0]):
sf = precleaned_epochs_engineering.info['sfreq']
bpAbs_4allchannels = []
bpRelative_4allchannels = []
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = fmin, fmax =fmax,
adaptive=True,normalization='full',verbose=0)
psd = np.log10(psd*10e12)
# average over channels
psd = psd.mean(axis=0)
freq_res = freqs[1] - freqs[0]
bp_total = simps(psd, dx=freq_res)
idx_band = np.logical_and(freqs >= low, freqs <= high)
bp_abs = simps(psd[idx_band], dx=freq_res)
bp_relative = bp_abs/bp_total
bpAbs_4Epochs.append(bp_abs)
bpRelative_4Epochs.append(bp_relative)
bpAbs_mean4Epochs = np.array(bpAbs_4Epochs).mean()
bpRelative_mean4Epochs = np.array(bpRelative_4Epochs).mean()
if subj in listNovices:
group = ['novice']
elif subj in listExperts:
group = ['expert']
data_df_psd = {'subj':[subj],'ROI':[roi],'state':[state],'condition':[condition],
'group':group,'rhythm':[waveband],'AbsBP':[bpAbs_mean4Epochs],
'RelativeBP':[bpRelative_mean4Epochs]}
df_psd_perWaveband = pd.DataFrame(data_df_psd)
frames_waveband.append(df_psd_perWaveband)
df_psd_perROI = pd.concat(frames_waveband,ignore_index=True)
frames_ROI.append(df_psd_perROI)
df_psd_perState = pd.concat(frames_ROI,ignore_index=True)
frames_state.append(df_psd_perState)
df_psd_perCondition = pd.concat(frames_state,ignore_index=True)
frames_condition.append(df_psd_perCondition)
df_psd_perSubject = pd.concat(frames_condition,ignore_index=True)
frames_subject.append(df_psd_perSubject)
df_psd = pd.concat(frames_subject,ignore_index=True)
df_psd.to_csv(path_or_buf='/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/df_bp_log_cluster.csv',index = False)
print('finished')
df_psd.to_csv(path_or_buf='/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/df_bp.csv',index = False)
### get bp matrice for 6 baseline expert/novice VD FA OP -baseline
import pickle
import mne
import matplotlib.pyplot as plt
from mne.time_frequency import psd_array_multitaper
from scipy.integrate import simps
import numpy as np
import seaborn as sns
import pandas as pd
import sys
from mne.channels import find_ch_connectivity
from scipy.stats.distributions import f,t
from mpl_toolkits.axes_grid1 import make_axes_locatable
precleaned_epochs_path = '/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/'
fmin = 1
fmax = 100
def getBpAbsAndRelative4allChannels(epochs,rhythm):
wavebands = {'alpha':[8,12],'theta':[3,7],'beta':[13,24],'lowG':[25,40],'highG':[60,90]}
if rhythm in wavebands.keys():
low,high = wavebands[rhythm]
else:
print('not such rhythm')
bpAbs_4Epochs=[]
bpRelative_4Epochs=[]
data = epochs.get_data(picks=['eeg'])
for num_epochs in range(data.shape[0]):
sf = epochs.info['sfreq']
bpAbs_4allchannels = []
bpRelative_4allchannels = []
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = 1, fmax =100,
adaptive=True,normalization='full',verbose=0)
psd= np.log10(psd*10e12)
freq_res = freqs[1] - freqs[0]
bp_total = simps(psd, dx=freq_res)
idx_band = np.logical_and(freqs >= low, freqs <= high)
bp_abs = simps(psd[:,idx_band], dx=freq_res)
bp_relative = bp_abs/bp_total
bpAbs_4Epochs.append(bp_abs)
bpRelative_4Epochs.append(bp_relative)
bpAbs_mean4Epochs = np.append([bpAbs_4Epochs[0]],bpAbs_4Epochs[1:],axis = 0).mean(axis=0)
bpRelative_mean4Epochs = np.append([bpRelative_4Epochs[0]],bpRelative_4Epochs[1:],axis = 0).mean(axis=0)
return bpAbs_mean4Epochs,bpRelative_mean4Epochs
def getBpAbs4allChannels(epochs,rhythm):
wavebands = {'alpha':[8,12],'theta':[3,7],'beta':[13,24],'lowG':[25,40],'highG':[60,90]}
if rhythm in wavebands.keys():
low,high = wavebands[rhythm]
else:
print('not such rhythm')
bpAbs_4Epochs=[]
data = epochs.get_data(picks=['eeg'])
for num_epochs in range(data.shape[0]):
sf = epochs.info['sfreq']
bpAbs_4allchannels = []
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = 1, fmax =100,
adaptive=True,normalization='full',verbose=0)
psd= np.log10(psd*10e12)
freq_res = freqs[1] - freqs[0]
idx_band = np.logical_and(freqs >= low, freqs <= high)
bp_abs = simps(psd[:,idx_band], dx=freq_res)
bpAbs_4Epochs.append(bp_abs)
bpAbs_mean4Epochs = np.append([bpAbs_4Epochs[0]],bpAbs_4Epochs[1:],axis = 0).mean(axis=0)
return bpAbs_mean4Epochs
# alpha bp clustering test - VD OP
subjs=['02', '04','07', '11', '12', '14', '16', '18', '19', '21', '22', '26', '28', '30',
'32', '34', '36', '37', '38', '40', '42', '50', '51', '52', '53', '54', '55', '56',
'58', '59','60', '63', '65', '67', '68', '70', '72', '73', '78', '83', '87', '88',
'90', '91', '93', '94', '95', '96','10','25','29','39','57','64','69','80','81','82',
'35','71','79','76','77']
# states_codes={'VD':['111.0','112.0','121.0','122.0','131.0','132.0'],
# 'FA':['211.0','212.0','221.0','222.0','231.0','232.0'],
# 'OP':['311.0','312.0','321.0','322.0','331.0','332.0']}
states_codes={'VD':['111.0','112.0'],
'FA':['211.0','212.0'],
'OP':['311.0','312.0']}
listNovices = ['02', '04', '07', '10', '11', '12', '14', '16', '18', '19', '21', '22', '26',
'28', '29', '30', '32', '34', '35', '36', '37', '38','39', '40', '42', '81', '82',
'83', '87', '88', '90', '91', '93', '94', '95', '96']
listExperts = ['25', '50','51', '52',
'53', '54', '55', '56', '57', '58', '59', '60', '63', '64', '65', '67','68', '69' ,'70' ,
'71', '72', '73', '76', '77', '78' ,'79', '80']
group = sys.argv[1]
# group = 'expert'
bpAbs_mean4Epochs_VD4allsubjs = np.array([])
bpAbs_mean4Epochs_FA4allsubjs = np.array([])
bpAbs_mean4Epochs_OP4allsubjs = np.array([])
if group == 'expert':
subjs = listExperts
elif group == 'novice':
subjs = listNovices
else:
print('no such group')
for subj in subjs:
precleaned_epochs_fname = precleaned_epochs_path + 'subj0'+subj+'full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
# precleaned_epochs_VD = precleaned_epochs[states_codes['VD']]
# precleaned_epochs_OP = precleaned_epochs[states_codes['OP']]
# bpAbs_mean4Epochs_VD = getBpAbs4allChannels(precleaned_epochs_VD,'alpha')
# bpAbs_mean4Epochs_OP= getBpAbs4allChannels(precleaned_epochs_OP,'alpha')
precleaned_epochs_VD = precleaned_epochs[states_codes['VD']]
precleaned_epochs_FA = precleaned_epochs[states_codes['FA']]
bpAbs_mean4Epochs_VD = getBpAbs4allChannels(precleaned_epochs_VD,'alpha')
bpAbs_mean4Epochs_FA= getBpAbs4allChannels(precleaned_epochs_FA,'alpha')
# precleaned_epochs_FA = precleaned_epochs[states_codes['FA']]
# precleaned_epochs_OP = precleaned_epochs[states_codes['OP']]
# bpAbs_mean4Epochs_FA = getBpAbs4allChannels(precleaned_epochs_FA,'alpha')
# bpAbs_mean4Epochs_OP= getBpAbs4allChannels(precleaned_epochs_OP,'alpha')
if len(bpAbs_mean4Epochs_VD4allsubjs)==0:
bpAbs_mean4Epochs_VD4allsubjs = bpAbs_mean4Epochs_VD
# bpRelative_mean4Epochs_VD4allsubjs = bpRelative_mean4Epochs_VD
else:
bpAbs_mean4Epochs_VD4allsubjs = np.vstack((bpAbs_mean4Epochs_VD4allsubjs,bpAbs_mean4Epochs_VD))
# bpRelative_mean4Epochs_VD4allsubjs = np.vstack((bpRelative_mean4Epochs_VD4allsubjs,
# bpRelative_mean4Epochs_VD))
# if len(bpAbs_mean4Epochs_OP4allsubjs)==0:
# bpAbs_mean4Epochs_OP4allsubjs = bpAbs_mean4Epochs_OP
# # bpRelative_mean4Epochs_OP4allsubjs = bpRelative_mean4Epochs_OP
# else:
# bpAbs_mean4Epochs_OP4allsubjs = np.vstack((bpAbs_mean4Epochs_OP4allsubjs,bpAbs_mean4Epochs_OP))
# # bpRelative_mean4Epochs_OP4allsubjs = np.vstack((bpRelative_mean4Epochs_OP4allsubjs,
# # bpRelative_mean4Epochs_OP))
if len(bpAbs_mean4Epochs_FA4allsubjs)==0:
bpAbs_mean4Epochs_FA4allsubjs = bpAbs_mean4Epochs_FA
# bpRelative_mean4Epochs_OP4allsubjs = bpRelative_mean4Epochs_OP
else:
bpAbs_mean4Epochs_FA4allsubjs = np.vstack((bpAbs_mean4Epochs_FA4allsubjs,bpAbs_mean4Epochs_FA))
# bpRelative_mean4Epochs_OP4allsubjs = np.vstack((bpRelative_mean4Epochs_OP4allsubjs,
# bpRelative_mean4Epochs_OP))
# bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)]
bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)]
# bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1),
# np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)]
# with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/VDOP_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
# pickle.dump(bpAbs_mean4Epochs2test, fp)
with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/VDFA_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
pickle.dump(bpAbs_mean4Epochs2test, fp)
# with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/FAOP_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
# pickle.dump(bpAbs_mean4Epochs2test, fp)
precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
connectivity, ch_names = find_ch_connectivity(precleaned_epochs.info, ch_type='eeg')
p_threshold = 0.05
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
n_permutations=10000,tail=0,threshold=threshold,
n_jobs=2, buffer_size=None,verbose=True,
connectivity=connectivity)
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
print(clusters)
# good_cluster_inds = np.array(range(len(clusters)))
# precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
# precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
# pos = mne.find_layout(precleaned_epochs.info).pos
# for i_clu, clu_idx in enumerate(good_cluster_inds):
# # unpack cluster information, get unique indices
# time_inds, space_inds = np.squeeze(clusters[clu_idx])
# ch_inds = np.unique(space_inds)
# time_inds = np.unique(time_inds)
# # get topography for bp-mean
# # bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
# # bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
# bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)).mean(axis=0))
# # create spatial mask
# mask = np.zeros((bp_map.shape[0], 1), dtype=bool)
# mask[ch_inds, :] = True
# # initialize figure
# fig, ax_topo = plt.subplots(1, 1, figsize=(10, 3))
# # plot average test statistic and mark significant sensors
# image, _ = mne.viz.plot_topomap(bp_map, pos, mask=mask, axes=ax_topo, cmap='Reds',
# vmin=np.min, vmax=np.max, show=False)
# divider = make_axes_locatable(ax_topo)
# # add axes for colorbar
# ax_colorbar = divider.append_axes('right', size='5%', pad=0.05)
# plt.colorbar(image, cax=ax_colorbar)
# # ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-VD for {}'.format(group))
# # ax_topo.set_xlabel('Averaged baseline alpha bandpower FA-VD for {}'.format(group))
# ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-FA for {}'.format(group))
# mne.viz.tight_layout(fig=fig)
# fig.subplots_adjust(bottom=.05)
# # plt.show()
# # fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-VD_alpha_baseline topoplot.png')
# # fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/FA-VD_alpha_baseline topoplot'+'i_clu'+'.png')
# fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-FA_alpha_baseline topoplot'+'i_clu'+'.png')
with open("/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/VDOP_alpha_baseline_Absnovice.txt", "rb") as fp: # Unpickling
bp = pickle.load(fp)
# bpAbs_mean4Epochs_VD4allsubjs = bp[0]
# bpAbs_mean4Epochs_FA4allsubjs = bp[1]
precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
connectivity, ch_names = find_ch_connectivity(precleaned_epochs.info, ch_type='eeg')
p_threshold = 0.05
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
threshold = -t.ppf(p_threshold/2,bp[0].shape[0]-1)
cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(bp[1]-bp[0],
n_permutations=10000,tail=0,threshold=threshold,
n_jobs=2, buffer_size=None,verbose=True,
connectivity=connectivity)
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
print(clusters)
good_cluster_inds = np.array(range(len(clusters)))
precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
pos = mne.find_layout(precleaned_epochs.info).pos
for i_clu, clu_idx in enumerate(good_cluster_inds):
# unpack cluster information, get unique indices
time_inds, space_inds = np.squeeze(clusters[clu_idx])
ch_inds = np.unique(space_inds)
time_inds = np.unique(time_inds)
# get topography for bp-mean
# bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
# bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
bp_map = np.squeeze((bp[1]-bp[0]).mean(axis=0))
# create spatial mask
mask = np.zeros((bp_map.shape[0], 1), dtype=bool)
mask[ch_inds, :] = True
# initialize figure
fig, ax_topo = plt.subplots(1, 1, figsize=(10, 3))
# plot average test statistic and mark significant sensors
image, _ = mne.viz.plot_topomap(bp_map, pos, mask=mask, axes=ax_topo, cmap='Reds',
vmin=np.min, vmax=np.max, show=False)
divider = make_axes_locatable(ax_topo)
# add axes for colorbar
ax_colorbar = divider.append_axes('right', size='5%', pad=0.05)
plt.colorbar(image, cax=ax_colorbar)
# ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-VD for {}'.format(group))
ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-VD for novice, pv = {}'.format(p_values[i_clu]))
# ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-FA for {}'.format(group))
mne.viz.tight_layout(fig=fig)
fig.subplots_adjust(bottom=.05)
# plt.show()
# fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-VD_alpha_baseline topoplot.png')
fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-VD_alpha_baseline_novice topoplot'+str(i_clu)+'.png')
# fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-FA_alpha_baseline_novice topoplot'+str(i_clu)+'.png')
p_values
from scipy.stats.distributions import f,t
p_threshold = 0.05
threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1),
n_permutations=10000,tail=0,threshold=threshold,
n_jobs=1, buffer_size=None,verbose=True,
connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
clusters[0]
# good_cluster_inds = np.where(p_values < p_threshold)[0]
# good_cluster_inds
clusters[0][1]
from mpl_toolkits.axes_grid1 import make_axes_locatable
good_cluster_inds = np.array([0])
# precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
# precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
pos = mne.find_layout(precleaned_epochs.info).pos
for i_clu, clu_idx in enumerate(good_cluster_inds):
# unpack cluster information, get unique indices
time_inds, space_inds = np.squeeze(clusters[clu_idx])
ch_inds = np.unique(space_inds)
time_inds = np.unique(time_inds)
# get topography for bp-mean
bp_map = np.squeeze(-(np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)).mean(axis=0))
# create spatial mask
mask = np.zeros((f_map.shape[0], 1), dtype=bool)
mask[ch_inds, :] = True
# initialize figure
fig, ax_topo = plt.subplots(1, 1, figsize=(10, 3))
# plot average test statistic and mark significant sensors
image, _ = mne.viz.plot_topomap(bp_map, pos, mask=mask, axes=ax_topo, cmap='Reds',
vmin=np.min, vmax=np.max, show=False)
divider = make_axes_locatable(ax_topo)
# add axes for colorbar
ax_colorbar = divider.append_axes('right', size='5%', pad=0.05)
plt.colorbar(image, cax=ax_colorbar)
np.squeeze((np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)).mean(axis=0)).shape
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
chan_coord=np.array([[-0.3088288, 0.9504773 ]])
mne.viz.plot_topomap(np.array([1.3]),pos=chan_coord,axes=ax)
mne.viz.tight_layout()
from mne.channels import find_ch_connectivity
connectivity, ch_names = find_ch_connectivity(precleaned_epochs.info, ch_type='eeg')
# set cluster threshold
threshold = 2 do not use
# set family-wise p-value
p_accept = 0.5
# threshold=threshold,
cluster_stats = mne.stats.spatio_temporal_cluster_test(bpRelative_mean4Epochs2test, n_permutations=1000,
tail=1,n_jobs=1, buffer_size=None,
connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
good_cluster_inds = np.where(p_values < p_accept)[0]
### get psd matrix ###
## to save at the end
# >>> from tempfile import TemporaryFile
# >>> outfile = TemporaryFile()
# >>> x = np.arange(10)
# >>> np.save(outfile, x)
# >>> _ = outfile.seek(0) # Only needed here to simulate closing & reopening file
# >>> np.load(outfile)
# array([0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
import numpy as np
import mne
from mne.time_frequency import psd_array_multitaper
subjs=['02', '04','07', '11', '12', '14', '16', '18', '19', '21', '22', '26', '28', '30',
'32', '34', '36', '37', '38', '40', '42', '50', '51', '52', '53', '54', '55', '56',
'58', '59','60', '63', '65', '67', '68', '70', '72', '73', '78', '83', '87', '88',
'90', '91', '93', '94', '95', '96','10','25','29','39','57','64','69','80','81','82',
'35','71','79','76','77']
states_codes={'VD':['111.0','112.0'],
'FA':['211.0','212.0'],
'OP':['311.0','312.0']}
# create VD baseline
precleaned_epochs_path = '/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/'
psdList_mean4epochs4allSubjs = []
for subj in subjs:
precleaned_epochs_fname = precleaned_epochs_path + 'subj0'+subj+'full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
precleaned_epochs_OP = precleaned_epochs[states_codes['OP']]
data = precleaned_epochs_OP.get_data(picks=['eeg'])
psd4epochs = []
for num_epochs in range(data.shape[0]):
sf = precleaned_epochs_OP.info['sfreq']
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = 1, fmax =100,
adaptive=True,normalization='full',verbose=0)
psd= np.log10(psd*10e12)
psd4epochs.append(psd)
psd_mean4epochs = np.append([psd4epochs[0]],psd4epochs[1:],axis = 0).mean(axis=0)
psdList_mean4epochs4allSubjs.append(psd_mean4epochs)
psd_final = np.append([psdList_mean4epochs4allSubjs[0]],psdList_mean4epochs4allSubjs[1:],axis = 0)
np.save('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/freq_channels4allSubjsOP.npy', psd_final)
psd_OP = np.load('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/freq_channels4allSubjsOP.npy')
psd_OP.shape
# this script is aimed to add interpolated channel to precleaned_epochs
### important not take into account eletrodes' positions
# import mne
# precleaned_epochs_path = '/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/'
# precleaned_epochs_fname = precleaned_epochs_path + 'subj077full_epo.fif'
# precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
# precleaned_epochs.info['ch_names']
# epochs2E = precleaned_epochs.copy()
# epochs2E.pick_channels(['F1','F3'])
# epochs2E.info['ch_names']=['P9','P10']
# epochs2E.info['chs'][0]['ch_name']='P9'
# epochs2E.info['chs'][1]['ch_name']='P10'
# precleaned_epochs.add_channels([epochs2E])
# precleaned_epochs.info['bads']=['P9','P10']
# precleaned_epochs.interpolate_bads()
# precleaned_epochs.save(precleaned_epochs_fname)
## save list
import pickle
with open("VDOP_alpha_Abs.txt", "wb") as fp: #Pickling
pickle.dump(bpAbs_mean4Epochs2test, fp)
with open("VDOP_alpha_Relative.txt", "wb") as fp: #Pickling
pickle.dump(bpAbs_mean4Epochs2test, fp)
# >>> with open("test.txt", "rb") as fp: # Unpickling
# ... b = pickle.load(fp)
# of no interest psd plot
psd_OP = np.load('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/freq_channels4allSubjsOP.npy')
plt.figure()
average = psd_OP.mean(axis=0)
# average.shape
sem = psd_OP.std(axis = 0)/np.sqrt(psd_OP.shape[0])
plt.semilogx(freqs,average)
plt.fill_between(freqs,average+sem,average-sem,alpha=.4)
plt.title('Average PSD for all subjs for all channels')
plt.legend('OP')
plt.show()
### get bp matrice for 6 baseline expert/novice VD FA OP -baseline local version
import pickle
import mne
import matplotlib.pyplot as plt
from mne.time_frequency import psd_array_multitaper
from scipy.integrate import simps
import numpy as np
import seaborn as sns
import pandas as pd
import sys
from mne.channels import find_ch_connectivity
from scipy.stats.distributions import f,t
from mpl_toolkits.axes_grid1 import make_axes_locatable
precleaned_epochs_path = '/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/'
fmin = 1
fmax = 100
def getBpAbsAndRelative4allChannels(epochs,rhythm):
wavebands = {'alpha':[8,12],'theta':[3,7],'beta':[13,24],'lowG':[25,40],'highG':[60,90]}
if rhythm in wavebands.keys():
low,high = wavebands[rhythm]
else:
print('not such rhythm')
bpAbs_4Epochs=[]
bpRelative_4Epochs=[]
data = epochs.get_data(picks=['eeg'])
for num_epochs in range(data.shape[0]):
sf = epochs.info['sfreq']
bpAbs_4allchannels = []
bpRelative_4allchannels = []
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = 1, fmax =100,
adaptive=True,normalization='full',verbose=0)
psd= np.log10(psd*10e12)
freq_res = freqs[1] - freqs[0]
bp_total = simps(psd, dx=freq_res)
idx_band = np.logical_and(freqs >= low, freqs <= high)
bp_abs = simps(psd[:,idx_band], dx=freq_res)
bp_relative = bp_abs/bp_total
bpAbs_4Epochs.append(bp_abs)
bpRelative_4Epochs.append(bp_relative)
bpAbs_mean4Epochs = np.append([bpAbs_4Epochs[0]],bpAbs_4Epochs[1:],axis = 0).mean(axis=0)
bpRelative_mean4Epochs = np.append([bpRelative_4Epochs[0]],bpRelative_4Epochs[1:],axis = 0).mean(axis=0)
return bpAbs_mean4Epochs,bpRelative_mean4Epochs
def getBpAbs4allChannels(epochs,rhythm):
wavebands = {'alpha':[8,12],'theta':[3,7],'beta':[13,24],'lowG':[25,40],'highG':[60,90]}
if rhythm in wavebands.keys():
low,high = wavebands[rhythm]
else:
print('not such rhythm')
bpAbs_4Epochs=[]
data = epochs.get_data(picks=['eeg'])
for num_epochs in range(data.shape[0]):
sf = epochs.info['sfreq']
bpAbs_4allchannels = []
psd, freqs = psd_array_multitaper(data[num_epochs], sf, fmin = 1, fmax =100,
adaptive=True,normalization='full',verbose=0)
psd= np.log10(psd*10e12)
freq_res = freqs[1] - freqs[0]
idx_band = np.logical_and(freqs >= low, freqs <= high)
bp_abs = simps(psd[:,idx_band], dx=freq_res)
bpAbs_4Epochs.append(bp_abs)
bpAbs_mean4Epochs = np.append([bpAbs_4Epochs[0]],bpAbs_4Epochs[1:],axis = 0).mean(axis=0)
return bpAbs_mean4Epochs
# alpha bp clustering test - VD OP
subjs=['02', '04','07', '11', '12', '14', '16', '18', '19', '21', '22', '26', '28', '30',
'32', '34', '36', '37', '38', '40', '42', '50', '51', '52', '53', '54', '55', '56',
'58', '59','60', '63', '65', '67', '68', '70', '72', '73', '78', '83', '87', '88',
'90', '91', '93', '94', '95', '96','10','25','29','39','57','64','69','80','81','82',
'35','71','79','76','77']
# states_codes={'VD':['111.0','112.0','121.0','122.0','131.0','132.0'],
# 'FA':['211.0','212.0','221.0','222.0','231.0','232.0'],
# 'OP':['311.0','312.0','321.0','322.0','331.0','332.0']}
states_codes={'VD':['111.0','112.0'],
'FA':['211.0','212.0'],
'OP':['311.0','312.0']}
listNovices = ['02', '04', '07', '10', '11', '12', '14', '16', '18', '19', '21', '22', '26',
'28', '29', '30', '32', '34', '35', '36', '37', '38','39', '40', '42', '81', '82',
'83', '87', '88', '90', '91', '93', '94', '95', '96']
listExperts = ['25', '50','51', '52']
# '53', '54', '55', '56', '57', '58', '59', '60', '63', '64', '65', '67','68', '69' ,'70' ,
# '71', '72', '73', '76', '77', '78' ,'79', '80']
# group = sys.argv[1]
group = 'expert'
bpAbs_mean4Epochs_VD4allsubjs = np.array([])
bpAbs_mean4Epochs_FA4allsubjs = np.array([])
bpAbs_mean4Epochs_OP4allsubjs = np.array([])
if group == 'expert':
subjs = listExperts
elif group == 'novice':
subjs = listNovices
else:
print('no such group')
for subj in subjs:
precleaned_epochs_fname = precleaned_epochs_path + 'subj0'+subj+'full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
# precleaned_epochs_VD = precleaned_epochs[states_codes['VD']]
# precleaned_epochs_OP = precleaned_epochs[states_codes['OP']]
# bpAbs_mean4Epochs_VD = getBpAbs4allChannels(precleaned_epochs_VD,'alpha')
# bpAbs_mean4Epochs_OP= getBpAbs4allChannels(precleaned_epochs_OP,'alpha')
# precleaned_epochs_VD = precleaned_epochs[states_codes['VD']]
# precleaned_epochs_FA = precleaned_epochs[states_codes['FA']]
# bpAbs_mean4Epochs_VD = getBpAbs4allChannels(precleaned_epochs_VD,'alpha')
# bpAbs_mean4Epochs_FA= getBpAbs4allChannels(precleaned_epochs_FA,'alpha')
precleaned_epochs_FA = precleaned_epochs[states_codes['FA']]
precleaned_epochs_OP = precleaned_epochs[states_codes['OP']]
bpAbs_mean4Epochs_FA = getBpAbs4allChannels(precleaned_epochs_FA,'alpha')
bpAbs_mean4Epochs_OP= getBpAbs4allChannels(precleaned_epochs_OP,'alpha')
# if len(bpAbs_mean4Epochs_VD4allsubjs)==0:
# bpAbs_mean4Epochs_VD4allsubjs = bpAbs_mean4Epochs_VD
# # bpRelative_mean4Epochs_VD4allsubjs = bpRelative_mean4Epochs_VD
# else:
# bpAbs_mean4Epochs_VD4allsubjs = np.vstack((bpAbs_mean4Epochs_VD4allsubjs,bpAbs_mean4Epochs_VD))
# # bpRelative_mean4Epochs_VD4allsubjs = np.vstack((bpRelative_mean4Epochs_VD4allsubjs,
# # bpRelative_mean4Epochs_VD))
if len(bpAbs_mean4Epochs_OP4allsubjs)==0:
bpAbs_mean4Epochs_OP4allsubjs = bpAbs_mean4Epochs_OP
# bpRelative_mean4Epochs_OP4allsubjs = bpRelative_mean4Epochs_OP
else:
bpAbs_mean4Epochs_OP4allsubjs = np.vstack((bpAbs_mean4Epochs_OP4allsubjs,bpAbs_mean4Epochs_OP))
# bpRelative_mean4Epochs_OP4allsubjs = np.vstack((bpRelative_mean4Epochs_OP4allsubjs,
# bpRelative_mean4Epochs_OP))
if len(bpAbs_mean4Epochs_FA4allsubjs)==0:
bpAbs_mean4Epochs_FA4allsubjs = bpAbs_mean4Epochs_FA
# bpRelative_mean4Epochs_OP4allsubjs = bpRelative_mean4Epochs_OP
else:
bpAbs_mean4Epochs_FA4allsubjs = np.vstack((bpAbs_mean4Epochs_FA4allsubjs,bpAbs_mean4Epochs_FA))
# bpRelative_mean4Epochs_OP4allsubjs = np.vstack((bpRelative_mean4Epochs_OP4allsubjs,
# bpRelative_mean4Epochs_OP))
# bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)]
# bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)]
bpAbs_mean4Epochs2test = [np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1),
np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)]
# with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/VDOP_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
# pickle.dump(bpAbs_mean4Epochs2test, fp)
# with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/VDFA_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
# pickle.dump(bpAbs_mean4Epochs2test, fp)
with open('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/FAOP_alpha_baseline_Abs'+group+'.txt', "wb") as fp: #Pickling
pickle.dump(bpAbs_mean4Epochs2test, fp)
precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
connectivity, ch_names = find_ch_connectivity(precleaned_epochs.info, ch_type='eeg')
p_threshold = 0.05
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
# threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1).shape[0]-1)
# cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1),
# n_permutations=10000,tail=0,threshold=threshold,
# n_jobs=2, buffer_size=None,verbose=True,
# connectivity=connectivity)
threshold = -t.ppf(p_threshold/2,np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1).shape[0]-1)
cluster_stats = mne.stats.spatio_temporal_cluster_1samp_test(np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1),
n_permutations=10000,tail=0,threshold=threshold,
n_jobs=2, buffer_size=None,verbose=True,
connectivity=connectivity)
T_obs, clusters, p_values, _ = cluster_stats
print(clusters)
good_cluster_inds = np.array(range(len(clusters)))
precleaned_epochs_fname = precleaned_epochs_path + 'subj004full_epo.fif'
precleaned_epochs = mne.read_epochs(precleaned_epochs_fname, preload=True)
pos = mne.find_layout(precleaned_epochs.info).pos
for i_clu, clu_idx in enumerate(good_cluster_inds):
# unpack cluster information, get unique indices
time_inds, space_inds = np.squeeze(clusters[clu_idx])
ch_inds = np.unique(space_inds)
time_inds = np.unique(time_inds)
# get topography for bp-mean
# bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
# bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_VD4allsubjs,axis=1)).mean(axis=0))
bp_map = np.squeeze((np.expand_dims(bpAbs_mean4Epochs_OP4allsubjs,axis=1)-np.expand_dims(bpAbs_mean4Epochs_FA4allsubjs,axis=1)).mean(axis=0))
# create spatial mask
mask = np.zeros((bp_map.shape[0], 1), dtype=bool)
mask[ch_inds, :] = True
# initialize figure
fig, ax_topo = plt.subplots(1, 1, figsize=(10, 3))
# plot average test statistic and mark significant sensors
image, _ = mne.viz.plot_topomap(bp_map, pos, mask=mask, axes=ax_topo, cmap='Reds',
vmin=np.min, vmax=np.max, show=False)
divider = make_axes_locatable(ax_topo)
# add axes for colorbar
ax_colorbar = divider.append_axes('right', size='5%', pad=0.05)
plt.colorbar(image, cax=ax_colorbar)
# ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-VD for {}'.format(group))
# ax_topo.set_xlabel('Averaged baseline alpha bandpower FA-VD for {}'.format(group))
ax_topo.set_xlabel('Averaged baseline alpha bandpower OP-FA for {}'.format(group))
mne.viz.tight_layout(fig=fig)
fig.subplots_adjust(bottom=.05)
# plt.show()
# fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-VD_alpha_baseline topoplot.png')
# fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/FA-VD_alpha_baseline topoplot'+'i_clu'+'.png')
fig.savefig('/home/gansheng.tan/process_mne/INSERM_EEG_Enrico_Proc/data_eeglab/full_epochs_data/statistic/OP-FA_alpha_baseline topoplot'+'i_clu'+'.png')
```
| github_jupyter |
# Trading Strategy
<li> Imoport library
```
#pip install pyfolio
%matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn
from sklearn.linear_model import LinearRegression
from scipy import stats
import pylab as pl
import datetime
import math
import pydotplus
from IPython.display import Image
from sklearn.linear_model import LogisticRegression
from sklearn import metrics
import statsmodels.tsa.api as smt
import statsmodels.api as sm
import scipy.stats as scs
from arch import arch_model
#import pyfolio as pf
import warnings
warnings.filterwarnings('ignore')
```
<li> Data preparation
```
def data_pre(stock_file,reddit_file,nytime_file):
## Stock
stock = pd.read_csv(stock_file)#'AMZN.csv'
#daily
stock['log_rt'] = np.log(stock['Adj Close']/stock['Adj Close'].shift(1))
stock['lin_rt'] = stock['Adj Close']-stock['Adj Close'].shift(1)
stock['direction'] = np.where(stock['log_rt']<0,0,1)
# weekly
stock['day']=stock['Date'].apply(lambda x: datetime.datetime.strptime(x,'%Y-%m-%d'))
stock['weekinfo'] =stock['day'].apply(lambda x: x.isocalendar())
stock['week'] = stock['weekinfo'].apply(lambda x: str(x[0])+'-'+str(x[1]))
stock['weekday'] = stock['weekinfo'].apply(lambda x: x[2])
stock_weekly = pd.DataFrame(stock,columns=['week','weekday','log_rt','lin_rt'])
stock_weekly=stock_weekly.groupby(by='week').mean()
stock_weekly['direction'] = np.where(stock_weekly['log_rt']<0,0,1)
stock_weekly['direction'] = np.sign(stock_weekly['log_rt'])
#stock_weekly = pd.DataFrame(stock,columns=['week','direction'])
stock_weekly=stock_weekly.reset_index()
stock_weekly = pd.DataFrame(stock_weekly,columns=['week','direction','lin_rt'])
## Reddit
reddit = pd.read_csv(reddit_file)#'
reddit = pd.DataFrame(reddit,columns=['timestamp','upvote_ratio','score','comms_num','anticipation','sadness',
'joy','negative','trust','positive','surprise','disgust','anger','fear'])
# modified
reddit['modified_positive'] = (reddit['positive']+reddit['joy']+reddit['trust']+reddit['anticipation'])
reddit['modified_negative'] = (reddit['negative']+reddit['disgust']+reddit['anger']+reddit['fear'])
reddit['timestamp']=reddit['timestamp'].apply(lambda x: datetime.datetime.strptime(x,'%Y-%m-%d %H:%M:%S'))
reddit['timestamp']=reddit['timestamp'].apply(lambda x: datetime.datetime.date(x))
# take the average of all article post on same date
reddit=reddit.groupby(by='timestamp').mean()
reddit.sort_index(ascending=True,inplace=True)
reddit=reddit.reset_index()
reddit['Date'] = reddit['timestamp'].apply(lambda x: str(x))
## NY-times
nytimes = pd.read_csv(nytime_file)
nytimes = pd.DataFrame(nytimes,columns=['date','week','anticipation','sadness','joy','negative',
'trust','positive','surprise','disgust','anger','fear'])
# modified
nytimes['modified_positive'] = (nytimes['positive']+nytimes['joy']+nytimes['trust']+nytimes['anticipation'])
nytimes['modified_negative'] = (nytimes['negative']+nytimes['disgust']+nytimes['anger']+nytimes['fear'])
# take the average of all article post on same date
nytimes=nytimes.groupby(by='week').mean()
nytimes.sort_index(ascending=True,inplace=True)
nytimes=nytimes.reset_index()
return stock, stock_weekly, reddit, nytimes
```
<li> Classification function
```
def get_classification(predictions,threshold):
classes = np.ones_like(predictions)
for i in range(len(classes)):
if predictions[i] < threshold:
classes[i] = -1
return classes
```
<li> Machine learning models we contained
```
def ml_modle(x_train,y_train,x_test,y_test,threshold=0.5,SVM_C=1,hid_layer=3,net_pass=10):
#garch
import statsmodels.tsa.api as smt
import statsmodels.api as sm
import scipy.stats as scs
from arch import arch_model
## linear regression model
from sklearn import linear_model
model1 = linear_model.LinearRegression()
model1.fit(x_train,y_train)
linear_pred = model1.predict(x_test)[0]
## random forest
from sklearn.ensemble import RandomForestClassifier
model3 = RandomForestClassifier(n_estimators=10)
model3.fit(x_train,y_train)
rf_pred = model3.predict(x_test)[0]
## SVM
#sigmoid
from sklearn import svm
def sigmoid2(x,deriv=False):
if deriv:
return x*(1-x)
return 1/(1+np.exp(-x))
model5 = svm.SVC(kernel='sigmoid', C=SVM_C,gamma='scale')
model5.fit(x_train,y_train['direction'])
model5.score(x_train,y_train)
SVM_sigmoid_pred = model5.predict(x_test)[0]
#Logistic Regression
logreg = LogisticRegression()
logreg.fit(x_train, y_train)
logistic_pred = [round(value) for value in logreg.predict(x_test)][0]
def net_non_lin(X,y,activation_function=sigmoid2,hidden_layer=1,passes=10):
import time
np.random.seed(1)
syn0 = 2*np.random.random((X.shape[1],hidden_layer)) - 1
syn1 = 2*np.random.random((hidden_layer,1)) - 1
for i in range(passes):
level_0 = X
level_1 = activation_function(np.dot(level_0,syn0))
level_2 = activation_function(np.dot(level_1,syn1))
level_2_error = y - level_2 # error term
level_2_delta = level_2_error*activation_function(level_2,deriv=True)
level_1_error = level_2_delta.dot(syn1.T)
level_1_delta = level_1_error * activation_function(level_1,deriv=True)
syn1 += level_1.T.dot(level_2_delta)
syn0 += level_0.T.dot(level_1_delta)
return syn0,syn1
syn0,syn1 = net_non_lin(x_train,y_train,activation_function=sigmoid2,hidden_layer=hid_layer,passes=net_pass)
level_0 = x_test
level_1 = sigmoid2(np.dot(level_0,syn0))
level_2 = sigmoid2(np.dot(level_1,syn1))
NN_nonlin_pred= level_2[0][0]
return linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred#garch_pred
```
<li> Return Rolling window results for each model
```
def predict(data,emotion,span,threshold=0.5,SVM_C=1,hid_layer=3,net_pass=10):
results = {'linear':[],'Randomforest':[],'SVM_sigmoid':[],'logistic':[],"NN_nonlin":[]}
train_len = int(data.shape[0]*0.7)
a1,a2,a3,a4 = 15,-2,9,10
for i in range(data.shape[0]-train_len):
y_train = data.iloc[i:train_len+i,a3:a4]
y_test = data.iloc[train_len+i:(train_len+i+1),a3:a4]
if emotion == 'all':
x_train = data.iloc[i:train_len+i,a1:a2]
x_test = data.iloc[train_len+i:(train_len+i+1),a1:a2]
else:
x_train = data[['upvote_ratio','score','comms_num','modified_positive','modified_negative']].iloc[i:train_len+i]
x_test = data[['upvote_ratio','score','comms_num','modified_positive','modified_negative']].iloc[train_len+i:(train_len+i+1)]
#print(x_train,y_train)
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = ml_modle(x_train,y_train,x_test,y_test,threshold=0,SVM_C=1)
#,garch_pred
#results['garch'].append(garch_pred)
results['linear']+=list(linear_pred)
results['Randomforest'].append(rf_pred)
#results['SVM_linear'].append(SVM_linear_predictions)
results['SVM_sigmoid'].append(SVM_sigmoid_pred)
results['logistic'].append(logistic_pred)
results['NN_nonlin'].append(NN_nonlin_pred)
return np.array(results['linear']),np.array(results['Randomforest']),np.array(results['SVM_sigmoid']),np.array(results['logistic']),np.array(results['NN_nonlin'])
```
<li> Compute trading results
```
def startegy(data,emotion,span,threshold=0.5,SVM_C=1,hid_layer=4,net_pass=10):
if span == 'daily':
train_len = int(data.shape[0]*0.7)
stock_dtest = data.iloc[(train_len-1):-1,8]
stock=np.array(stock_dtest).reshape(1,len(stock_dtest))
else:
train_len = int(data.shape[0]*0.7)
stock_wtest = data.iloc[(train_len-1):-1,8]
stock=-np.array(stock_wtest).reshape(1,len(stock_wtest))
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = predict(data,emotion,span,threshold=0.5,SVM_C=1)
linear_pred = np.cumsum(stock*get_classification(linear_pred,np.mean(linear_pred)))
rf_pred = np.cumsum(-stock*get_classification(rf_pred,np.mean(rf_pred)))
SVM_sigmoid_pred = np.cumsum(stock*get_classification(SVM_sigmoid_pred,np.mean(SVM_sigmoid_pred)))
logistic_pred =np.cumsum(-stock*get_classification(logistic_pred,np.mean(logistic_pred)))
#NN_nonlin_pred = np.cumsum(stock*get_classification(NN_nonlin_pred,np.mean(NN_nonlin_pred)))
return linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred
```
<li> Evaluate our trading strategies
```
def evaluate(daily,stock_dtest,emotion):
models = startegy(daily,emotion,span='daily',threshold=0.5,SVM_C=1,hid_layer=4,net_pass=100)
model_names = ['raw data','linear','random forest','SVM_sigmoid','logistic']
raw = np.cumsum(stock_dtest)
score={'mean':[np.mean(raw)],
'volatility':[np.std(raw)],
'profit rate':[sum(np.where(raw>=0,1,0))/len(raw)],
'annulized return':[sum(raw)/len(raw)]}
for i in range(4):
score['mean'].append(np.mean(models[i]))
score['volatility'].append(np.std(models[i]))
score['profit rate'].append(sum(np.where(models[i]>=0,1,0))/len(models[i]))
score['annulized return'].append(sum(models[i])/len(models[i]))
#print(score)
score = pd.DataFrame(score,index=model_names)
return score
```
# TEST CASES
# Amazon
```
stock,stock_weekly,reddit,nytimes = data_pre('data/AMZN.csv','data/am_reddit_emotions.csv','data/am_article_emotions.csv')
daily = pd.merge(stock, reddit, how='left', on='Date')
daily=daily.dropna(how='any')
weekly = pd.merge(stock_weekly, nytimes, how='left', on='week')
weekly=weekly.dropna(how='any')
train_len = int(daily.shape[0]*0.7)
x=daily.iloc[train_len:,]['Date']
stock_dtest = daily.iloc[(train_len-1):-1,8]
```
<li> All emotions
```
evaluate(daily,stock_dtest,emotion='all')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='all',span='daily',threshold=0.5,SVM_C=1,hid_layer=4,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'],['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Amazon - Reddit with all emotions')
plt.show()
```
<li> Modified emotions
```
evaluate(daily,stock_dtest,emotion='mod')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='mod',span='daily',threshold=0.5,SVM_C=1,hid_layer=2,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'],['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Amazon - Reddit with modified emotions')
plt.show()
```
# Google
```
stock,stock_weekly,reddit,nytimes = data_pre('data/GOOG.csv','data/gg_reddit_emotions.csv','data/gg_article_emotions.csv')
daily = pd.merge(stock, reddit, how='left', on='Date')
daily=daily.dropna(how='any')
weekly = pd.merge(stock_weekly, nytimes, how='left', on='week')
weekly=weekly.dropna(how='any')
train_len = int(daily.shape[0]*0.7)
x=daily.iloc[train_len:,]['Date']
stock_dtest = daily.iloc[(train_len-1):-1,8]
```
<li> All emotions
```
evaluate(daily,stock_dtest,emotion='all')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='all',span='daily',threshold=0.5,SVM_C=1,hid_layer=4,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'],['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Google - Reddit with all emotions')
plt.show()
```
<li> Modified emotions
```
evaluate(daily,stock_dtest,emotion='mod')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='mod',span='daily',threshold=0.5,SVM_C=1,hid_layer=2,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'],['2019-08-09','2019-09-03','2019-10-01','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Google - Reddit with modified emotions')
plt.show()
```
# Netflix
```
stock,stock_weekly,reddit,nytimes = data_pre('data/NFLX.csv','data/nf_reddit_emotions.csv','data/nf_article_emotions.csv')
daily = pd.merge(stock, reddit, how='left', on='Date')
daily=daily.dropna(how='any')
weekly = pd.merge(stock_weekly, nytimes, how='left', on='week')
weekly=weekly.dropna(how='any')
train_len = int(daily.shape[0]*0.7)
x=daily.iloc[train_len:,]['Date']
stock_dtest = daily.iloc[(train_len-1):-1,8]
```
<li> All emotions
```
evaluate(daily,stock_dtest,emotion='all')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='all',span='daily',threshold=0.5,SVM_C=1,hid_layer=3,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-13','2019-09-13','2019-10-11','2019-11-01','2019-11-26'],['2019-08-13','2019-09-13','2019-10-11','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Netflix - Reddit with all emotions')
plt.show()
```
<li> Modified emotions
```
evaluate(daily,stock_dtest,emotion='mod')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='mod',span='daily',threshold=0.5,SVM_C=1,hid_layer=3,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-13','2019-09-13','2019-10-11','2019-11-01','2019-11-26'],['2019-08-13','2019-09-13','2019-10-11','2019-11-01','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Netflix - Reddit with modified emotions')
plt.show()
```
# Facebook
```
stock,stock_weekly,reddit,nytimes = data_pre('data/FB.csv','data/fb_reddit_emotions.csv','data/fb_article_emotions.csv')
daily = pd.merge(stock, reddit, how='left', on='Date')
daily=daily.dropna(how='any')
weekly = pd.merge(stock_weekly, nytimes, how='left', on='week')
weekly=weekly.dropna(how='any')
train_len = int(daily.shape[0]*0.7)
x=daily.iloc[train_len:,]['Date']
stock_dtest = daily.iloc[(train_len-1):-1,8]
```
<li> All emotions
```
evaluate(daily,stock_dtest,emotion='all')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='all',span='daily',threshold=0.5,SVM_C=1,hid_layer=3,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-20','2019-09-20','2019-10-21','2019-11-11','2019-11-26'],['2019-08-20','2019-09-20','2019-10-21','2019-11-11','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Facebook - Reddit with all emotions')
plt.show()
```
<li> Modified Emotions
```
evaluate(daily,stock_dtest,emotion='mod')
linear_pred,rf_pred,SVM_sigmoid_pred,logistic_pred,NN_nonlin_pred = startegy(daily,emotion='mod',span='daily',threshold=0.5,SVM_C=1,hid_layer=3,net_pass=100)
plt.figure(figsize=(12,8))
from matplotlib.ticker import MultipleLocator
y = np.zeros_like(linear_pred)
plt.plot(x,linear_pred,color='green', label='linear regression')
plt.plot(x, rf_pred, color='orange', label='Random_forest')
plt.plot(x, SVM_sigmoid_pred, color='blue', label='SVM_sigmoid')
plt.plot(x, logistic_pred, color='red', label='logistic regression')
plt.plot(x, np.cumsum(stock_dtest), color='black', label='raw stock data')
plt.plot(x, y, color='yellow', label='0 line')
plt.xlabel("test date")
plt.ylabel("cummulative return($)")
plt.xticks(['2019-08-20','2019-09-20','2019-10-21','2019-11-11','2019-11-26'],['2019-08-20','2019-09-20','2019-10-21','2019-11-11','2019-11-26'])
plt.legend(loc = 'upper left')
plt.title('Facebook - Reddit with modified emotions')
plt.show()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Tessellate-Imaging/monk_v1/blob/master/study_roadmaps/2_transfer_learning_roadmap/3_effect_of_number_of_classes_in_dataset/1)%20Understand%20transfer%20learning%20and%20the%20role%20of%20number%20of%20dataset%20classes%20in%20it%20-%20Mxnet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Goals
### 1. Visualize deep learning network
### 2. Understand how the final layer would change when number of classes in dataset changes
# What do you do with a deep learning model in transfer learning
- These are the steps already done by contributors in pytorch, keras and mxnet
- You take a deep learning architecture, such as resnet, densenet, or even custom network
- Train the architecture on large datasets such as imagenet, coco, etc
- The trained wieghts become your starting point for transfer learning
- The final layer of this pretrained model has number of neurons = number of classes in the large dataset
- In transfer learning
- You take the network and load the pretrained weights on the network
- Then remove the final layer that has the extra(or less) number of neurons
- You add a new layer with number of neurons = number of classes in your custom dataset
- Optionally you can add more layers in between this newly added final layer and the old network
- Now you have two parts in your network
- One that already existed, the pretrained one, the base network
- The new sub-network or a single layer you added
- The hyper-parameter we can see here: Freeze base network
- Freezing base network makes the base network untrainable
- The base network now acts as a feature extractor and only the next half is trained
- If you do not freeze the base network the entire network is trained
(You will take this part in next sessions)
# Table of Contents
## [0. Install](#0)
## [1. Setup Default Params with Cats-Dogs dataset](#1)
## [2. Visualize network](#2)
## [3. Reset Default Params with new dataset - Logo classification](#3)
## [4. Visualize the new network](#4)
<a id='0'></a>
# Install Monk
- git clone https://github.com/Tessellate-Imaging/monk_v1.git
- cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
- (Select the requirements file as per OS and CUDA version)
```
!git clone https://github.com/Tessellate-Imaging/monk_v1.git
# Select the requirements file as per OS and CUDA version
!cd monk_v1/installation/Linux && pip install -r requirements_cu9.txt
```
## Dataset - Sample
- one having 2 classes
- other having 16 classes
```
! wget --load-cookies /tmp/cookies.txt "https://docs.google.com/uc?export=download&confirm=$(wget --save-cookies /tmp/cookies.txt --keep-session-cookies --no-check-certificate 'https://docs.google.com/uc?export=download&id=1jE-ckk0JbrdbJvIBaKMJWkTfbRDR2MaF' -O- | sed -rn 's/.*confirm=([0-9A-Za-z_]+).*/\1\n/p')&id=1jE-ckk0JbrdbJvIBaKMJWkTfbRDR2MaF" -O study_classes.zip && rm -rf /tmp/cookies.txt
! unzip -qq study_classes.zip
```
# Imports
```
# Monk
import os
import sys
sys.path.append("monk_v1/monk/");
#Using mxnet-gluon backend
from gluon_prototype import prototype
```
### Creating and managing experiments
- Provide project name
- Provide experiment name
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "study-num-classes");
```
### This creates files and directories as per the following structure
workspace
|
|--------Project
|
|-----study-num-classes
|
|-----experiment-state.json
|
|-----output
|
|------logs (All training logs and graphs saved here)
|
|------models (all trained models saved here)
<a id='1'></a>
# Setup Default Params with Cats-Dogs dataset
```
gtf.Default(dataset_path="study_classes/dogs_vs_cats",
model_name="resnet18_v1",
num_epochs=5);
```
### From Data summary - Num classes: 2
<a id='2'></a>
# Visualize network
```
gtf.Visualize_With_Netron(data_shape=(3, 224, 224), port=8081);
```
## The final layer
```
from IPython.display import Image
Image(filename='imgs/2_classes_base_gluon.png')
```
<a id='3'></a>
# Reset Default Params with new dataset - Logo classification
```
gtf = prototype(verbose=1);
gtf.Prototype("Project", "study-num-classes");
gtf.Default(dataset_path="study_classes/logos",
model_name="resnet18_v1",
num_epochs=5);
```
### From Data summary - Num classes: 16
<a id='4'></a>
# Visualize network
```
gtf.Visualize_With_Netron(data_shape=(3, 224, 224), port=8082);
```
## The final layer
```
from IPython.display import Image
Image(filename='imgs/16_classes_base_gluon.png')
```
| github_jupyter |
```
import matplotlib.pyplot as plt
```
Here's how to make the coin flip diagram:
```
f = plt.figure(figsize=(5.5, 5.5))
plt.bar([1, 2], [0.5, 0.5])
plt.xticks([1, 2], ['heads', 'tails'])
```
# Distributions
## The Normal Distribution
```
import numpy as np
from scipy.stats import norm
x = np.linspace(-4, 4, 100)
plt.plot(x, norm.pdf(x))
plt.plot(x, norm.cdf(x))
data = norm.rvs(size=10000, random_state=42)
plt.hist(data, bins=30)
x = np.linspace(12, 16, 100)
plt.plot(x, norm.pdf(x, loc=14, scale=0.5))
solar_data = norm.rvs(size=10000, loc=14, scale=0.5, random_state=42)
_ = plt.hist(solar_data, bins=30)
solar_data.mean()
solar_data.std()
import pandas as pd
df = pd.DataFrame(data={'efficiency': solar_data})
df['efficiency'].skew()
df.kurtosis()
import scipy
plt.plot(x, scipy.stats.skewnorm.pdf(x, scale=1.15, loc=13.1, a=5))
plt.plot(x, scipy.stats.lognorm.pdf(x, loc=12, s=0.5))
```
### Fitting distributions to get parameters
```
df = pd.read_csv('data/solar_cell_efficiencies.csv')
df.describe()
df.hist(bins=40)
df['efficiency'].skew()
df['efficiency'].kurt()
scipy.stats.norm.fit(df['efficiency'])
```
### Bernoulli Distribution
```
scipy.stats.bernoulli(p=0.7).rvs()
norm(loc=10).rvs()
```
### Binomial Distribution
```
binom_dist = scipy.stats.binom(p=0.7, n=10)
plt.bar(range(11), binom_dist.pmf(k=range(11)))
binom_dist.pmf(k=range(10))
```
### Exponential and Poisson Distributions
```
from labellines import labelLines
x = np.linspace(0, 5, 100)
plt.plot(x, scipy.stats.expon.pdf(x, scale=1), label='λ=1')
plt.plot(x, scipy.stats.expon.pdf(x, scale=0.25), label='λ=4')
labelLines(plt.gca().get_lines())
plt.bar(range(40), scipy.stats.poisson.pmf(range(40), mu=10))
```
### Weibull distribution
```
x = np.linspace(0, 10, 100)
plt.plot(x, scipy.stats.weibull_min(c=3).pdf(x))
x = range(1, 50)
plt.plot(x, scipy.stats.zipf(a=1.1).pmf(x), marker='.')
plt.xscale('log')
plt.yscale('log')
```
# Sampling
## Central Limit Theorem
```
binom = scipy.stats.binom(p=0.5, n=100)
binom.rvs()
heads = binom.rvs(10000)
import seaborn as sns
sns.histplot(heads, kde=True)
```
## Random sampling
```
df = pd.read_csv('data/solar_cell_efficiencies.csv')
df.sample(100, random_state=42)
df['efficiency'].mean()
import bootstrapped.bootstrap as bs
import bootstrapped.stats_functions as bs_stats
bs.bootstrap(df['efficiency'].values, stat_func=bs_stats.mean)
bs.bootstrap(df['efficiency'].values, stat_func=bs_stats.std)
means = []
for i in range(10000):
sample = np.random.choice(df['efficiency'], 1000, replace=True)
means.append(sample.mean())
sns.histplot(means)
```
| github_jupyter |
# Analysis of Gaussian Mixture Models
This notebook shows examples and usage of the biokit.stats.module.
The **mixture** module provides tools to create mixture models (gaussian) and to estimate the parameters of the GMM using either minimization or Expectation Minimization.
We also provide examples to show how to automatically estimate the number of GMMs and limitations of such methods.
```
%pylab inline
matplotlib.rcParams['figure.figsize'] = (12,6)
matplotlib.rcParams['figure.dpi'] = 120
matplotlib.style.use('ggplot')
from biokit.stats import mixture
```
### gaussian mixture model
```
m = mixture.GaussianMixture(mu=[-2, 1], sigma=[0.5,0.5],
mixture=[.2,.8], N=60)
# data is stored in m.data and an histogram can easily be shown:
m.hist()
# In order to figure out the underlying parameters of the distribution
# given in the above sample, we will first use the GaussianMixtureFitting
# class, which minimizes the distribution of the data to a mixture
# of gaussian model. A conveninet class to generate the data is the
# GaussianMixtureModel class, which can be used to plot the final distribution:
gmm = mixture.GaussianMixtureModel(k=2)
X = linspace(-3,3,100)
plot(X, gmm.pdf(X, params=[-1,0.5,0.2,1,0.5,0.8]), 'r')
```
### Gaussian Mixture model Fitting (minimization)
```
# now, let us use the following class to figure out the parameters of the
# gaussian models. We have to provide the number of models defined by
# the k=2 parameter
mf = mixture.GaussianMixtureFitting(m.data, k=2)
#
mf.estimate()
mf.plot()
```
### Expectation Minimization
Another method to estimate the parameters is to use an Expectation
Minimization algorithm
```
em = mixture.EM(m.data, model=gmm)
em.estimate()
em.plot()
em.results.mus
# to compare to numerical values found with the GaussianMixtureFitting
mf.results.mus
```
### How EM and minimization compare in terms of parameter estimation
Is the EM better than MixtureFitting class (scipy minimisation) ?
We can check that by generating N simulated data sets for some parameters
```
res_mf = []; res_em = []
N = 50
from easydev import Progress
replicates = 100
p = Progress(replicates)
mus = [-2,1]
sigmas = [0.5,0.5]
pis= [.2,0.8]
for i in range(0, replicates):
m = mixture.GaussianMixture(mu=mus, sigma=sigmas, mixture=pis, N=N)
mf = mixture.GaussianMixtureFitting(m.data)
mf.estimate(guess=[-.8,0.5,0.5,.8,0.5,0.5])
res_mf.append(mf.results)
em = mixture.EM(m.data, model=gmm)
em.estimate(guess=[-.8, 0.5, 0.5, .8, 0.5, 0.5])
res_em.append(em.results)
#p.animate(i+1)
subplot(2,3,1)
hist([x.mus[0] for x in res_mf], 20, alpha=0.4);
hist([x.mus[0] for x in res_em], 20, color='r', alpha=0.2)
axvline(mus[0], color='r', lw=2)
subplot(2,3,2)
hist([x.sigmas[0] for x in res_mf], 20, alpha=0.4);
hist([x.sigmas[0] for x in res_em], 20, color='r', alpha=0.2)
axvline(sigmas[0], color='r', lw=2)
subplot(2,3,3)
hist([x.pis[0] for x in res_mf], 20, alpha=0.4);
hist([x.pis[0] for x in res_em], 20, color='r', alpha=0.2)
axvline(pis[0], color='r', lw=2)
subplot(2,3,4)
hist([x.mus[1] for x in res_mf], 20, alpha=0.4);
hist([x.mus[1] for x in res_em], 20, color='r', alpha=0.2)
axvline(mus[1], color='r', lw=2)
subplot(2,3,5)
hist([x.sigmas[1] for x in res_mf], 20, alpha=0.4);
hist([x.sigmas[1] for x in res_em], 20, color='r', alpha=0.2)
axvline(sigmas[1], color='r', lw=2)
subplot(2,3,6)
hist([x.pis[1] for x in res_mf], 20, alpha=0.4);
hist([x.pis[1] for x in res_em], 20, color='r', alpha=0.2)
axvline(pis[1], color='r', lw=2)
```
Performace are quite similar on this example, but this may not always be the case !
### Adaptive Estimation of required number of models
```
m = mixture.GaussianMixture(mu=[-1, 1], sigma=[0.5,0.5],
mixture=[.2,.8], N=60)
# if we know that k=2, it is easy to estimate the parameters
res = mf.estimate(k=2)
mf.plot()
```
### What about automatic inference of k ?
```
amf = mixture.AdaptativeMixtureFitting(m.data)
amf.diagnostic(kmin=1, kmax=8)
```
The proposed/optimal k in terms of Aikike is k=4, which is
not correct.
What about using EM ?
```
amf.fitting = mixture.EM(m.data)
amf.run(kmin=1, kmax=8)
amf.diagnostic(k=2)
plot(amf.x[0:8], [amf.all_results[i]['AIC'] for i in amf.x[0:8]], 'or-')
plot(amf.x[0:8], [amf.all_results[i]['AICc'] for i in amf.x[0:8]], 'og-')
```
Now the question is how reliable is this estimation of k on 100 experiments ?
```
bestk_em = []
aics_em = []
bestk_m = []
aics_m = []
replicates = 100
p = Progress(replicates)
for i in range(0, replicates):
m = mixture.GaussianMixture(mu=[-1, 1], sigma=[0.5,0.5],
mixture=[.2,.8], N=1000)
amf = mixture.AdaptativeMixtureFitting(m.data)
amf.verbose = False
amf.fitting = mixture.EM(m.data)
amf.run()
bestk_em.append(amf.best_k)
aics_em.append(amf.fitting.results.AIC)
amf = mixture.AdaptativeMixtureFitting(m.data)
amf.verbose = False
amf.run()
bestk_m.append(amf.best_k)
aics_m.append(amf.fitting.results.AIC)
#p.animate(i+1)
res = hist([bestk_em, bestk_m])
legend(['EM', 'minimization'])
hist([aics_em, aics_m])
```
### Conclusions
When N = 60, the adaptative EM or Minimization methods do not allow the
recovery of k automaticaly. You need more data e.g. ~ 1000 points
| github_jupyter |
# Tutorial 6.2.2.Turbulence Synthesis Inlet for Large Eddy Simulation
Project: Structural Wind Engineering WS20-21
Chair of Structural Analysis @ TUM - R. Wüchner, M. Péntek, A. Kodakkal
Author: Ammar Khallouf
Created on: 24.11.2020
Last Update: 15.12.2020
#### Content:
A random flow generation (RFG) technique is implemented here on the basis of the work by **_Smirnov et al.(2001)_.** and **_Kraichnan (1970)_.**
According to the authors. The method can generate an isotropic divergence-free fluctuating velocity field satisfying the Gaussian's spectral model as well as an inhomogeneous and anisotropic turbulence flow, provided that an anisotropic velocity correlation tensor is given.
The transient flow field is generated as a superposition of harmonic functions with random coefficients.
The approach is used to set inlet boundary conditions to LES models as well as initial boundary conditions in the simulation of turbulent flow around bluff-bodies.
#### Input:
The input of this procedure are the mean speed at the inlet: ${u_{avg}}$ ,turbulent intensities: ${I_u}$ , ${I_v}$ , ${I_w}$
and turbulent length scale: ${L}$
#### References:
[1] _S. S. A. Smirnov and I. Celik, Random flow generation technique for large eddy simulations and particle-dynamics modeling, Journal of Fluids Engineering,123 (2001), pp. 359-371._
[2] _R. Kraichnan, Diffusion by a random velocity field, The Physics of Fluids, 13(1970), pp. 22-31._
```
# Import of some python modules and vector normal random generator.
import numpy as np
import time
import sys
import math
from pylab import *
import random
from IPython.core.display import display, Math
from scipy import signal
from matplotlib import pyplot as plt
```
#### 1. Read Input mean velocity, turbulent intensities and length scale
```
# u_avg -> mean longitudinal (y axis) wind speed, per node, [m/s]:
u_avg = 23.6
#turbulence intensity Iu != Iv != Iw
Iu = 0.1
Iv = 0.1
Iw = 0.1
urms = u_avg*Iu
vrms = u_avg*Iv
wrms = u_avg*Iw
#L -> turbulence length scale [m]
Lu = Lv = Lw = 0.1
Ls = math.sqrt(Lu**2 + Lv**2 + Lw**2)
# function to generate random vectors:
def normalrandomvec(mean,sigma):
x=random.gauss(mean,sigma)
y=random.gauss(mean,sigma)
z=random.gauss(mean,sigma)
return np.array([x,y,z]) # 1x3
```
#### 2. Given an anisotropic velocity correlation tensor $r_{i j}=\overline{\tilde{u}_{i} \tilde{u}_{j}}$ of a turbulent flow field $\tilde{u}_{i}\left(x_{j}, t\right)$.
#### We find an orthogonal transformation tensor $a_{i j}$ that diagonalizes $r_{i j}$:
$$a_{m i} a_{n j} r_{i j}=c_{(n)}^{2} \delta_{m n}
$$
$$a_{m i} a_{n j}=\delta_{i j}
$$
As a result of this step both $a_{i j}$ and $c_{(n)}$ become known functions of space.
Coefficients $c_{n}=\left\{c_{1}, c_{2}, c_{3}\right\}$ play the role of turbulent fluctuating velocities $\left(u^{\prime}, v^{\prime}, w^{\prime}\right)$ in the new coordinate system produced by transformation tensor $a_{i j}$
```
# construct anisotropic velocity correlation tensor:
rij = np.array([[urms**2, 0, 0],[0, vrms**2, 0],[0, 0, wrms**2]])
#find an orthogonal transformation tensor aij that would diagonalize rij:
aij, c2 = np.linalg.eig(rij)
cn = np.sqrt([aij[0], aij[1], aij[2]]) # 1x3
#N -> sampling number for each wavenumber kn
N = 1000
# x -> nodal coordinates
x = np.array([[0.0],[0.0],[0.0]]) # 3x1
timev = np.arange(0,3.,0.0001) # 4000,
```
#### 3. Generate a transient turbulent velocity field in 3D using a modifcation of the Kraichnan method _(Kraichnan,1970)_
$$v_{i}(x, t)=\sqrt{\frac{2}{N}} \sum_{n=1}^{N}\left[p_{i}^{n} \cos \left(\tilde{k}_{j}^{n} \tilde{x}_{j}+\omega_{n} \tilde{t}\right)+q_{i}^{n} \sin \left(\tilde{k}_{j}^{n} \tilde{x}_{j}+\omega_{n} \tilde{t}\right)\right]
$$
#### Where:
$$
\tilde{x}=\frac{x}{l}, \quad \tilde{t}=\frac{t}{\tau}, \quad \tilde{k}_{j}^{n}=k_{j}^{n} \frac{c}{c_{(j)}}, \quad c=\frac{l}{\tau}
$$
$$
p_{i}^{n}=\epsilon_{i j m} \zeta_{j}^{n} k_{m}^{n}, \quad q_{i}^{n}=\epsilon_{i j m} \xi_{j}^{n} k_{m}^{n}
$$
$$
\zeta_{i}^{n}, \xi_{i}^{n}, \omega_{n} \in N(0,1) ; \quad k_{i}^{n} \in N(0,1 / 2)
$$
Where:
$l, \tau$ are the length and time scales of turbulence, $\epsilon_{i j m}$ is the permutation
tensor and $N(M, \sigma)$ is a normal distribution with mean $M$ and standard deviation $\sigma$. $k_{i}^{n}$
and $\omega_{n}$ represent a sample of $n$ wavenumber vectors and frequencies
of the theoretical modeled turbulence spectrum:
$$
E(k)=16\left(\frac{2}{\pi}\right)^{1 / 2} k^{4} \exp \left(-2 k^{2}\right)
$$
```
print ("begin simulation...")
#modified Kraichnan's method
uxt = np.zeros([3,timev.size]) # 3x4000
pni = np.zeros([3,1]) # 3x1
qni = np.zeros([3,1]) # 3x1
knjtil = np.zeros([3,1]) # 3x1
#time-scale of turbulence [sec]
tau = Ls/u_avg
timetil = timev/tau # 4000,
xtil = x/Ls # 3x1
c = Ls/tau
un = np.zeros([3,timev.size]) # 3x4000
# initialize seed:
random.seed()
for n in range(0,N):
omegamn = random.gauss(0,1)
knj = normalrandomvec(0,0.5) # 1x3
Zetan = normalrandomvec(0,1) # 1x3
Xin = normalrandomvec(0,1) # 1x3
pni = np.cross(Zetan.transpose(),knj.transpose()) # 1x3
qni = np.cross(Xin.transpose(),knj.transpose()) # 1x3
knjtil[0,0] = knj[0]*c/cn[0]
knjtil[1,0] = knj[1]*c/cn[1]
knjtil[2,0] = knj[2]*c/cn[2]
un[0,:] = un[0,:] + \
+ pni[0]*cos(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) + \
+ qni[0]*sin(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) \
un[1,:] = un[1,:] + \
+ pni[1]*cos(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) + \
+ qni[1]*sin(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) \
un[2,:] = un[2,:] + \
+ pni[2]*cos(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) + \
+ qni[2]*sin(np.inner(knjtil.T,xtil.T) + omegamn*timetil[:]) \
uxt[0,:] = cn[0]*math.sqrt(2./N)*un[0,:]
uxt[1,:] = cn[1]*math.sqrt(2./N)*un[1,:]
uxt[2,:] = cn[2]*math.sqrt(2./N)*un[2,:]
print ("end simulation...")
```
#### 4. Scale and transform the velocity field $v_{i}$ generated in the previous step to obtain the turbulent flow field $u_{i}$
$$
\begin{array}{c}
w_{i}=c_{i} v_{i} \\
u_{i}=a_{i k} w_{k}
\end{array}
$$
The plots below show the turbulent fluctuations in the velocity field with respect to the average velocity $u_{avg}$
```
um = max(uxt[0,:])
vm = max(uxt[1,:])
wm = max(uxt[2,:])
subplot(311)
plot(timev,uxt[0,:]/u_avg,color="red",linewidth=1)
axis([0, 3, -um/u_avg, um/u_avg])
title('Turbulence Synthesis')
xlabel('time [secs]')
ylabel('$u_x$',fontsize=20)
grid(True)
subplot(312)
plot(timev,uxt[1,:]/u_avg,color="blue",linewidth=1)
axis([0, 3, -vm/u_avg, vm/u_avg])
xlabel('time [secs]')
ylabel('$u_y$',fontsize=20)
grid(True)
subplot(313)
plot(timev,uxt[2,:]/u_avg,color="green",linewidth=1)
axis([0, 3, -wm/u_avg, wm/u_avg])
xlabel('time [secs]')
ylabel('$u_z$', fontsize=20)
grid(True)
```
#### 5. Plot the energy spectrum on a log scale of simulated turbulence against the theoretical turbulence spectrum:
$$
E(k)=16\left(\frac{2}{\pi}\right)^{1 / 2} k^{4} \exp \left(-2 k^{2}\right)
$$
```
# simulated spectrum fluctuations
#fff,pf = signal.welch(uxt)
fff,pf = signal.periodogram(uxt,1/0.0001)
#plt.semilogx(fff,pf[2,:])
# theoretical spectrum fluctuations
kmax = 100
k = np.arange(0.1,kmax,0.5) # 400,
#k = f/u_avg
Ek = 16.*math.sqrt(2./math.pi)*(k/u_avg)**4.*(np.exp(-2.*(k/u_avg)**2.))
fig,ax = subplots()
ax.loglog(fff,pf[0,:],"b",linewidth=1)
ax.loglog(k,Ek,"r",linewidth=1)
ax.axis([1, 1000, 1.e-10, 100])
ax.set_xlabel('$k$ [1/Hz]',fontsize=20)
ax.set_ylabel('$E(k)$', fontsize=20)
ax.set_title('Turbulence Spectra', fontsize=20)
ax.legend(["Simulated Spectrum","Theoretical Spectrum"]);
ax.grid(True)
plt.rc('xtick',labelsize=12)
plt.rc('ytick',labelsize=12)
```
#### 6. Plot the 3D turbulent velocity field compoments as function of simulation time of 3 seconds
```
subplot(311)
plot(timev,u_avg+uxt[0,:],color="red",linewidth=1)
axis([0, 3, u_avg-um, u_avg+um])
title('Turbulence Synthesis')
xlabel('time [secs]')
ylabel('$U_x$ [m/s]',fontsize=12)
grid(True)
subplot(312)
plot(timev,uxt[1,:],color="blue",linewidth=1)
axis([0, 3, -vm, vm])
xlabel('time [secs]')
ylabel('$U_y$ [m/s]',fontsize=12)
grid(True)
subplot(313)
plot(timev,uxt[2,:],color="green",linewidth=1)
axis([0, 3, -wm, wm])
xlabel('time [secs]')
ylabel('$U_z$ [m/s]', fontsize=12)
grid(True)
```
| github_jupyter |
<i>Copyright (c) Microsoft Corporation. All rights reserved.</i>
<i>Licensed under the MIT License.</i>
# LightGBM: A Highly Efficient Gradient Boosting Decision Tree
This notebook will give you an example of how to train a LightGBM model to estimate click-through rates on an e-commerce advertisement. We will train a LightGBM based model on the Criteo dataset.
[LightGBM](https://github.com/Microsoft/LightGBM) is a gradient boosting framework that uses tree-based learning algorithms. It is designed to be distributed and efficient with the following advantages:
* Fast training speed and high efficiency.
* Low memory usage.
* Great accuracy.
* Support of parallel and GPU learning.
* Capable of handling large-scale data.
## Global Settings and Imports
```
import sys
import os
import numpy as np
import lightgbm as lgb
import papermill as pm
import scrapbook as sb
import pandas as pd
import category_encoders as ce
from tempfile import TemporaryDirectory
from sklearn.metrics import roc_auc_score, log_loss
import reco_utils.recommender.lightgbm.lightgbm_utils as lgb_utils
import reco_utils.dataset.criteo as criteo
print("System version: {}".format(sys.version))
print("LightGBM version: {}".format(lgb.__version__))
```
### Parameter Setting
Let's set the main related parameters for LightGBM now. Basically, the task is a binary classification (predicting click or no click), so the objective function is set to binary logloss, and 'AUC' metric, is used as a metric which is less effected by imbalance in the classes of the dataset.
Generally, we can adjust the number of leaves (MAX_LEAF), the minimum number of data in each leaf (MIN_DATA), maximum number of trees (NUM_OF_TREES), the learning rate of trees (TREE_LEARNING_RATE) and EARLY_STOPPING_ROUNDS (to avoid overfitting) in the model to get better performance.
Besides, we can also adjust some other listed parameters to optimize the results. [In this link](https://github.com/Microsoft/LightGBM/blob/master/docs/Parameters.rst), a list of all the parameters is shown. Also, some advice on how to tune these parameters can be found [in this url](https://github.com/Microsoft/LightGBM/blob/master/docs/Parameters-Tuning.rst).
```
MAX_LEAF = 64
MIN_DATA = 20
NUM_OF_TREES = 100
TREE_LEARNING_RATE = 0.15
EARLY_STOPPING_ROUNDS = 20
METRIC = "auc"
SIZE = "sample"
params = {
'task': 'train',
'boosting_type': 'gbdt',
'num_class': 1,
'objective': "binary",
'metric': METRIC,
'num_leaves': MAX_LEAF,
'min_data': MIN_DATA,
'boost_from_average': True,
#set it according to your cpu cores.
'num_threads': 20,
'feature_fraction': 0.8,
'learning_rate': TREE_LEARNING_RATE,
}
```
## Data Preparation
Here we use CSV format as the example data input. Our example data is a sample (about 100 thousand samples) from [Criteo dataset](https://www.kaggle.com/c/criteo-display-ad-challenge). The Criteo dataset is a well-known industry benchmarking dataset for developing CTR prediction models, and it's frequently adopted as evaluation dataset by research papers. The original dataset is too large for a lightweight demo, so we sample a small portion from it as a demo dataset.
Specifically, there are 39 columns of features in Criteo, where 13 columns are numerical features (I1-I13) and the other 26 columns are categorical features (C1-C26).
```
nume_cols = ["I" + str(i) for i in range(1, 14)]
cate_cols = ["C" + str(i) for i in range(1, 27)]
label_col = "Label"
header = [label_col] + nume_cols + cate_cols
with TemporaryDirectory() as tmp:
all_data = criteo.load_pandas_df(size=SIZE, local_cache_path=tmp, header=header)
display(all_data.head())
```
First, we cut three sets (train_data (first 80%), valid_data (middle 10%) and test_data (last 10%)), cut from the original all data. <br>
Notably, considering the Criteo is a kind of time-series streaming data, which is also very common in recommendation scenario, we split the data by its order.
```
# split data to 3 sets
length = len(all_data)
train_data = all_data.loc[:0.8*length-1]
valid_data = all_data.loc[0.8*length:0.9*length-1]
test_data = all_data.loc[0.9*length:]
```
## Basic Usage
### Ordinal Encoding
Considering LightGBM could handle the low-frequency features and missing value by itself, for basic usage, we only encode the string-like categorical features by an ordinal encoder.
```
ord_encoder = ce.ordinal.OrdinalEncoder(cols=cate_cols)
def encode_csv(df, encoder, label_col, typ='fit'):
if typ == 'fit':
df = encoder.fit_transform(df)
else:
df = encoder.transform(df)
y = df[label_col].values
del df[label_col]
return df, y
train_x, train_y = encode_csv(train_data, ord_encoder, label_col)
valid_x, valid_y = encode_csv(valid_data, ord_encoder, label_col, 'transform')
test_x, test_y = encode_csv(test_data, ord_encoder, label_col, 'transform')
print('Train Data Shape: X: {trn_x_shape}; Y: {trn_y_shape}.\nValid Data Shape: X: {vld_x_shape}; Y: {vld_y_shape}.\nTest Data Shape: X: {tst_x_shape}; Y: {tst_y_shape}.\n'
.format(trn_x_shape=train_x.shape,
trn_y_shape=train_y.shape,
vld_x_shape=valid_x.shape,
vld_y_shape=valid_y.shape,
tst_x_shape=test_x.shape,
tst_y_shape=test_y.shape,))
train_x.head()
```
### Create model
When both hyper-parameters and data are ready, we can create a model:
```
lgb_train = lgb.Dataset(train_x, train_y.reshape(-1), params=params, categorical_feature=cate_cols)
lgb_valid = lgb.Dataset(valid_x, valid_y.reshape(-1), reference=lgb_train, categorical_feature=cate_cols)
lgb_test = lgb.Dataset(test_x, test_y.reshape(-1), reference=lgb_train, categorical_feature=cate_cols)
lgb_model = lgb.train(params,
lgb_train,
num_boost_round=NUM_OF_TREES,
early_stopping_rounds=EARLY_STOPPING_ROUNDS,
valid_sets=lgb_valid,
categorical_feature=cate_cols)
```
Now let's see what is the model's performance:
```
test_preds = lgb_model.predict(test_x)
auc = roc_auc_score(np.asarray(test_y.reshape(-1)), np.asarray(test_preds))
logloss = log_loss(np.asarray(test_y.reshape(-1)), np.asarray(test_preds), eps=1e-12)
res_basic = {"auc": auc, "logloss": logloss}
print(res_basic)
sb.glue("res_basic", res_basic)
```
<script type="text/javascript" src="http://cdn.mathjax.org/mathjax/latest/MathJax.js?config=default"></script>
## Optimized Usage
### Label-encoding and Binary-encoding
Next, since LightGBM has a better capability in handling dense numerical features effectively, we try to convert all the categorical features in original data into numerical ones, by label-encoding [3] and binary-encoding [4]. Also due to the sequence property of Criteo, the label-encoding we adopted is executed one-by-one, which means we encode the samples in order, by the information of the previous samples before each sample (sequential label-encoding and sequential count-encoding). Besides, we also filter the low-frequency categorical features and fill the missing values by the mean of corresponding columns for the numerical features. (consulting `lgb_utils.NumEncoder`)
Specifically, in `lgb_utils.NumEncoder`, the main steps are as follows.
* Firstly, we convert the low-frequency categorical features to `"LESS"` and the missing categorical features to `"UNK"`.
* Secondly, we convert the missing numerical features into the mean of corresponding columns.
* Thirdly, the string-like categorical features are ordinal encoded like the example shown in basic usage.
* And then, we target encode the categorical features in the samples order one-by-one. For each sample, we add the label and count information of its former samples into the data and produce new features. Formally, for $i=1,2,...,n$, we add $\frac{\sum\nolimits_{j=1}^{i-1} I(x_j=c) \cdot y}{\sum\nolimits_{j=1}^{i-1} I(x_j=c)}$ as a new label feature for current sample $x_i$, where $c$ is a category to encode in current sample, so $(i-1)$ is the number of former samples, and $I(\cdot)$ is the indicator function that check the former samples contain $c$ (whether $x_j=c$) or not. At the meantime, we also add the count frequency of $c$, which is $\frac{\sum\nolimits_{j=1}^{i-1} I(x_j=c)}{i-1}$, as a new count feature.
* Finally, based on the results of ordinal encoding, we add the binary encoding results as new columns into the data.
Note that the statistics used in the above process only updates when fitting the training set, while maintaining static when transforming the testing set because the label of test data should be considered as unknown.
```
label_col = 'Label'
num_encoder = lgb_utils.NumEncoder(cate_cols, nume_cols, label_col)
train_x, train_y = num_encoder.fit_transform(train_data)
valid_x, valid_y = num_encoder.transform(valid_data)
test_x, test_y = num_encoder.transform(test_data)
del num_encoder
print('Train Data Shape: X: {trn_x_shape}; Y: {trn_y_shape}.\nValid Data Shape: X: {vld_x_shape}; Y: {vld_y_shape}.\nTest Data Shape: X: {tst_x_shape}; Y: {tst_y_shape}.\n'
.format(trn_x_shape=train_x.shape,
trn_y_shape=train_y.shape,
vld_x_shape=valid_x.shape,
vld_y_shape=valid_y.shape,
tst_x_shape=test_x.shape,
tst_y_shape=test_y.shape,))
```
### Training and Evaluation
```
lgb_train = lgb.Dataset(train_x, train_y.reshape(-1), params=params)
lgb_valid = lgb.Dataset(valid_x, valid_y.reshape(-1), reference=lgb_train)
lgb_model = lgb.train(params,
lgb_train,
num_boost_round=NUM_OF_TREES,
early_stopping_rounds=EARLY_STOPPING_ROUNDS,
valid_sets=lgb_valid)
test_preds = lgb_model.predict(test_x)
auc = roc_auc_score(np.asarray(test_y.reshape(-1)), np.asarray(test_preds))
logloss = log_loss(np.asarray(test_y.reshape(-1)), np.asarray(test_preds), eps=1e-12)
res_optim = {"auc": auc, "logloss": logloss}
print(res_optim)
sb.glue("res_optim", res_optim)
```
## Model saving and loading
Now we finish the basic training and testing for LightGBM, next let's try to save and reload the model, and then evaluate it again.
```
with TemporaryDirectory() as tmp:
save_file = os.path.join(tmp, r'finished.model')
lgb_model.save_model(save_file)
loaded_model = lgb.Booster(model_file=save_file)
# eval the performance again
test_preds = loaded_model.predict(test_x)
auc = roc_auc_score(np.asarray(test_y.reshape(-1)), np.asarray(test_preds))
logloss = log_loss(np.asarray(test_y.reshape(-1)), np.asarray(test_preds), eps=1e-12)
print({"auc": auc, "logloss": logloss})
```
## Additional Reading
\[1\] Guolin Ke, Qi Meng, Thomas Finley, Taifeng Wang, Wei Chen, Weidong Ma, Qiwei Ye, and Tie-Yan Liu. 2017. LightGBM: A highly efficient gradient boosting decision tree. In Advances in Neural Information Processing Systems. 3146–3154.<br>
\[2\] The parameters of LightGBM: https://github.com/Microsoft/LightGBM/blob/master/docs/Parameters.rst <br>
\[3\] Anna Veronika Dorogush, Vasily Ershov, and Andrey Gulin. 2018. CatBoost: gradient boosting with categorical features support. arXiv preprint arXiv:1810.11363 (2018).<br>
\[4\] Scikit-learn. 2018. categorical_encoding. https://github.com/scikit-learn-contrib/categorical-encoding<br>
| github_jupyter |
<h1 style = "color:red">Covid-19 Model Analysis</h1>
<h3>Python/DataScience Internship Program</h3>
<hr>
<h3>Batch Code- PD04</h3>
<h3>Name- Nitin Diwakar</h3>
<h3>Guided/Instructed by- Miss Shubhangi Sonker</h3>
#### Importing Modules
```
%matplotlib inline
import numpy as np
# import matplotlib as mpl
import pandas as pd
import seaborn as sns
import os
import matplotlib.pyplot as plt
plt.style.use("ggplot")
import warnings
warnings.filterwarnings("ignore")
```
### Loading Data
```
covid = pd.read_csv("covidreports.csv")
```
### Analysis the data
```
covid.head()
covid.tail()
covid.shape
covid.info()
covid['Age'].value_counts()
covid['Gender'].value_counts()
covid['fever'].value_counts()
covid.info()
covid.describe()
```
### Analysing/ representing the data by Histogram, Heatmap and Countplot
```
covid.hist(bins=20, figsize=(16,9))
plt.show()
corr_matrix = covid.corr()
corr_matrix['Infected'].sort_values(ascending = False)
plt.figure(figsize = (20,20))
sns.heatmap(corr_matrix.corr(), annot=True)
plt.show()
plt.figure(figsize = (12,6))
sns.countplot(data = covid, x = "Infected", hue = "Severity")
```
## Creating a model for analysing the covid case of China
```
China_case = covid[covid["Country"] == "China"]
China_case.head()
China_case.shape
x = China_case['Age']
y = China_case["Infected"]
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(x,y,test_size=0.3, random_state=30)
```
## Training and Testing the data using RandomForestClassifier
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import accuracy_score
rnd_clf = RandomForestClassifier()
rnd_clf.fit(np.array(X_train).reshape(-1,1), np.array(y_train).reshape(-1,1))
rnd_clf.score(np.array(X_train).reshape(-1,1), np.array(y_train).reshape(-1,1))
```
### Training
```
y_rnd_pred = rnd_clf.predict(np.array(X_test).reshape(-1,1))
y_rnd_pred = rnd_clf.predict(np.array(X_train).reshape(-1,1))
from sklearn.metrics import precision_score, recall_score
from sklearn.metrics import accuracy_score, f1_score
print("Accuracy after CV :", accuracy_score(y_train, y_rnd_pred))
print("Pricision after CV:", precision_score(y_train, y_rnd_pred))
print("Recall after CV :", recall_score(y_train, y_rnd_pred))
print("f1_score after CV :", f1_score(y_train, y_rnd_pred))
```
### Testing
```
y_rnd_pred = rnd_clf.predict(np.array(X_test).reshape(-1,1))
print("Accuracy after CV :", accuracy_score(y_test, y_rnd_pred))
print("Pricision after CV:", precision_score(y_test, y_rnd_pred))
print("Recall after CV :", recall_score(y_test, y_rnd_pred))
print("f1_score after CV :", f1_score(y_test, y_rnd_pred))
```
## Testing the model using LogisticRegression
### Test
```
x=covid[['fever','Bodypain','Runny_nose','Difficulty_in_breathing','Nasal_congestion','Sore_throat']].values
y=covid['Infected'].values
X_train,X_test,y_train,y_test=train_test_split(x,y)
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score,f1_score
from sklearn.metrics import precision_score,recall_score
clf=LogisticRegression()
clf.fit(X_train,y_train)
pred=clf.predict(X_test)
print("Accuracy after CV :", round(accuracy_score(pred, y_test)*100,2))
print("Pricision after CV:", round(precision_score(pred, y_test)*100,2))
print("Recall after CV :", round(recall_score(pred, y_test)*100,2))
print("f1_score after CV :", round(f1_score(pred, y_test)*100,2))
```
| github_jupyter |
# This notebook demonstrates some very basic functionality currently present in the library
```
import keras
from importlib import reload
import os
import sys
sys.path.insert(0, "C:/Users/magaxels/AutoML")
import gazer; reload(gazer)
from gazer import GazerMetaLearner
```
## Load som dummy-data
```
from sklearn.datasets import load_digits
X, y = load_digits(return_X_y=True)
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = \
train_test_split(X, y, test_size=0.25, random_state=0)
from sklearn.preprocessing import (MaxAbsScaler,
RobustScaler,
StandardScaler,
MinMaxScaler)
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
X_train.shape, y_train.shape, X_test.shape, y_test.shape
```
## Import selected algorithms using
- *method* = 'select'
- *estimators* = ['adaboost', 'svm', 'neuralnet', 'logreg']
- verbose = 1
- Provides some feedback
```
learner = GazerMetaLearner(method='select',
estimators=['neuralnet', 'adaboost', 'logreg', 'svm'],
verbose=1)
learner.names
```
## Inspect neural network parameters
Note that these have been automatically set for you (~ reasonable defaults)
We shall learn how to change these
```
learner.clf['neuralnet'].network
```
## Perform parameter update using the self.update method
```
learner.update('neuralnet', {'epochs': 100, 'n_hidden': 3, 'input_units': 500})
```
### Note changes in the below dictionary
- We set
- epochs = 100
- n_hidden = 3
- input_units = 500
Note that *input_units* is the number of neurons in each layer
```
learner.clf['neuralnet'].network
```
If the user fails to provide proper input then (providing self.verbose = 1) we provide
the signature to the $__init__()$ method. This helps to determine allowed parameters and their values
```
learner.update('logreg', {'bla': 1})
```
## Train learner
We train all initialized algorithms using **learner.fit**
```
learner.fit(X_train, y_train)
```
Since *verbose* = 1 we get a lot of output. If you wish not to see it, then set *verbose* = 0.
That way, **gazer** stays mute during the training process.
```
learner.verbose = 0
learner.fit(X_train, y_train)
```
See Mom; no output!
## Evalute on test data
We can easily evaluate how well algorithms generalize using **learner.evaluate**
```
learner.evaluate(X_test, y_test, metric='accuracy', get_loss=True)
```
..
..
# End of demo
| github_jupyter |
<img src="../../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# _*Qiskit Aqua: Pricing Fixed-Income Assets*_
The latest version of this notebook is available on https://github.com/Qiskit/qiskit-tutorials.
***
### Contributors
Stefan Woerner<sup>[1]</sup>, Daniel Egger<sup>[1]</sup>, Shaohan Hu<sup>[1]</sup>, Stephen Wood<sup>[1]</sup>, Marco Pistoia<sup>[1]</sup>
### Affliation
- <sup>[1]</sup>IBMQ
### Introduction
We seek to price a fixed-income asset knowing the distributions describing the relevant interest rates. The cash flows $c_t$ of the asset and the dates at which they occur are known. The total value $V$ of the asset is thus the expectation value of:
$$V = \sum_{t=1}^T \frac{c_t}{(1+r_t)^t}$$
Each cash flow is treated as a zero coupon bond with a corresponding interest rate $r_t$ that depends on its maturity. The user must specify the distribution modelling the uncertainty in each $r_t$ (possibly correlated) as well as the number of qubits he wishes to use to sample each distribution. In this example we expand the value of the asset to first order in the interest rates $r_t$. This corresponds to studying the asset in terms of its duration.
<br>
<br>
The approximation of the objective function follows the following paper:<br>
<a href="https://arxiv.org/abs/1806.06893">Quantum Risk Analysis. Woerner, Egger. 2018.</a>
```
import matplotlib.pyplot as plt
%matplotlib inline
import numpy as np
from qiskit import BasicAer
from qiskit_aqua.algorithms import AmplitudeEstimation
from qiskit_aqua.components.random_distributions import MultivariateNormalDistribution
from qiskit_aqua.components.uncertainty_problems import FixedIncomeExpectedValue
```
### Uncertainty Model
We construct a circuit factory to load a multivariate normal random distribution in $d$ dimensions into a quantum state.
The distribution is truncated to a given box $\otimes_{i=1}^d [low_i, high_i]$ and discretized using $2^{n_i}$ grid points, where $n_i$ denotes the number of qubits used for dimension $i = 1,\ldots, d$.
The unitary operator corresponding to the circuit factory implements the following:
$$\big|0\rangle_{n_1}\ldots\big|0\rangle_{n_d} \mapsto \big|\psi\rangle = \sum_{i_1=0}^{2^n_-1}\ldots\sum_{i_d=0}^{2^n_-1} \sqrt{p_{i_1,...,i_d}}\big|i_1\rangle_{n_1}\ldots\big|i_d\rangle_{n_d},$$
where $p_{i_1, ..., i_d}$ denote the probabilities corresponding to the truncated and discretized distribution and where $i_j$ is mapped to the right interval $[low_j, high_j]$ using the affine map:
$$ \{0, \ldots, 2^{n_{j}}-1\} \ni i_j \mapsto \frac{high_j - low_j}{2^{n_j} - 1} * i_j + low_j \in [low_j, high_j].$$
In addition the the uncertainty model, we can also apply an affine map, e.g. resulting from a principal componant analyis. The interest rates used are then given by:
$$ \vec{r} = A * \vec{x} + b,$$
where $\vec{x} \in \otimes_{i=1}^d [low_i, high_i]$ follows the given random distribution.
```
# can be used in case a principal component analysis has been done to derive the uncertainty model, ignored in this example.
A = np.eye(2)
b = np.zeros(2)
# specify the number of qubits that are used to represent the different dimenions of the uncertainty model
num_qubits = [2, 2]
# specify the lower and upper bounds for the different dimension
low = [0, 0]
high = [0.12, 0.24]
mu = [0.12, 0.24]
sigma = 0.01*np.eye(2)
# construct corresponding distribution
u = MultivariateNormalDistribution(num_qubits, low, high, mu, sigma)
# plot contour of probability density function
x = np.linspace(low[0], high[0], 2**num_qubits[0])
y = np.linspace(low[1], high[1], 2**num_qubits[1])
z = u.probabilities.reshape(2**num_qubits[0], 2**num_qubits[1])
plt.contourf(x, y, z)
plt.xticks(x, size=15)
plt.yticks(y, size=15)
plt.grid()
plt.xlabel('$r_1$ (%)', size=15)
plt.ylabel('$r_2$ (%)', size=15)
plt.colorbar()
plt.show()
```
### Cash flow, payoff function, and exact expected value
In the following we define the cash flow per period, the resulting payoff function and evaluate the exact expected value.
For the payoff function we first use a first order approximation and then apply the same approximation technique as for the linear part of the payoff function of the [European Call Option](european_call_option_pricing.ipynb).
```
# specify cash flow
cf = [1.0, 2.0]
periods = range(1, len(cf)+1)
# plot cash flow
plt.bar(periods, cf)
plt.xticks(periods, size=15)
plt.yticks(size=15)
plt.grid()
plt.xlabel('periods', size=15)
plt.ylabel('cashflow ($)', size=15)
plt.show()
# estimate real value
cnt = 0
exact_value = 0.0
for x1 in np.linspace(low[0], high[0], pow(2, num_qubits[0])):
for x2 in np.linspace(low[1], high[1], pow(2, num_qubits[1])):
prob = u.probabilities[cnt]
for t in range(len(cf)):
# evaluate linear approximation of real value w.r.t. interest rates
exact_value += prob * (cf[t]/pow(1 + b[t], t+1) - (t+1)*cf[t]*np.dot(A[:, t], np.asarray([x1, x2]))/pow(1 + b[t], t+2))
cnt += 1
print('Exact value: \t%.4f' % exact_value)
# specify approximation factor
c_approx = 0.125
# get fixed income circuit appfactory
fixed_income = FixedIncomeExpectedValue(u, A, b, cf, c_approx)
# set number of evaluation qubits (samples)
m = 5
# construct amplitude estimation
ae = AmplitudeEstimation(m, fixed_income)
# result = ae.run(quantum_instance=BasicAer.get_backend('qasm_simulator'), shots=100)
result = ae.run(quantum_instance=BasicAer.get_backend('statevector_simulator'))
print('Exact value: \t%.4f' % exact_value)
print('Estimated value:\t%.4f' % result['estimation'])
print('Probability: \t%.4f' % result['max_probability'])
# plot estimated values for "a" (direct result of amplitude estimation, not rescaled yet)
plt.bar(result['values'], result['probabilities'], width=0.5/len(result['probabilities']))
plt.xticks([0, 0.25, 0.5, 0.75, 1], size=15)
plt.yticks([0, 0.25, 0.5, 0.75, 1], size=15)
plt.title('"a" Value', size=15)
plt.ylabel('Probability', size=15)
plt.xlim((0,1))
plt.ylim((0,1))
plt.grid()
plt.show()
# plot estimated values for fixed-income asset (after re-scaling and reversing the c_approx-transformation)
plt.bar(result['mapped_values'], result['probabilities'], width=3/len(result['probabilities']))
plt.plot([exact_value, exact_value], [0,1], 'r--', linewidth=2)
plt.xticks(size=15)
plt.yticks([0, 0.25, 0.5, 0.75, 1], size=15)
plt.title('Estimated Option Price', size=15)
plt.ylabel('Probability', size=15)
plt.ylim((0,1))
plt.grid()
plt.show()
```
| github_jupyter |
# Otimização
## Introdução
- Problemas de otimização são encontrados em diversas situações da Engenharia e indústria
- Muitos algoritmos da inteligência artificial utilizam algum princípio de otimização
- Problemas de otimização são comumente tratados como *problemas de minimização*:
- Busca-se o _mínimo global_ de uma _função objetivo_ (FO) escalar $f(x)$
- Maximizar $f(x)$ equivalente a minimizar $-f(x)$.
### Restrições
- POs são acompanhados de _restrições_:
- Para $g(x) = 0$, $g(x)$ é uma _restrição de igualdade_;
- Para $h(x) \leq 0$, $h(x)$ é uma _restrição de desigualdade_.
### Classificação de problemas de otimização
- _univariado_ (ou _unidimensional_), se $x$ é escalar, i.e. $x \in \mathbb{R}$;
- _multivariado_ (ou _multidimensional_), se $x$ é um vetor, i.e. $x \in \mathbb{R}^n$.
- _linear_: se a FO e as restrições são funções lineares. Neste caso, por razões históricas, diz-se que o problema é de _programação linear_.
- _não-linear_: se a FO e as restrições são funções não-lineares. Neste caso, diz-se que o problema é de _programação não-linear_.
Com respeito às restrições, um PO pode ainda ser:
- _irrestrito_: quando não se assumem limites para os valores de $x$.
- _restrito_: quando limites para os valores de $x$ são impostos.
Aqui trataremos apenas de casos em que $x \in \mathbb{R}$.
### Problemas convexos
- Problemas não-lineares são mais difíceis de resolver do que problemas lineares
- Podem admitir uma ampla variedade de comportamentos.
- Um PO não-linear pode ter tanto _mínimos locais_ quanto _mínimos globais_.
- Encontrar o _mínimo global_ de uma função $f(x)$ não-linear exige técnicas aperfeiçoadas.
- Subclasse de problemas não-lineares que pode ser resolvida eficientemente: _convexos_.
```
import numpy as np
import matplotlib.pyplot as plt
import sympy as sy
sy.init_printing()
```
### Funções convexas
> Uma função convexa definida em um intervalo $[a,b]$ é aquela em que todos os seus valores estão abaixo da reta secante que passa pelos pontos $(a,f(a))$ e $(b,f(b)$. Isto, por sua vez, garante que ela contenha _somente_ um mínimo global.
**Exemplo**: a função $f(x) = 3x^2 - 0.36x - 11.2$ é convexa em $[-2,3]$.
```
# domínio
a,b = -2,3
x = np.linspace(a,b,100)
# função e valores nos extremos
f = lambda x: 5*x**2 - 10.36*x - 11.2
fa,fb = f(a),f(b)
# reta secante
s = fa + (fb - fa)/(b - a)*(x - a)
# ponto de mínimo:
xmin = 10.36/10
# plotagem de funções
plt.figure(figsize=(5,3))
plt.plot(x,f(x))
plt.plot(x,s,color='#ffa500')
# pontos da secante
plt.plot(a,f(a),'o',color='#ffa500')
plt.plot(b,f(b),'o',color='#ffa500')
# ponto de mínimo
plt.plot(xmin,f(xmin),'*r',ms=10);
plt.title('Exemplo de função convexa');
```
**Exemplo**: a função $p(x) = 10x^2\textrm{sen}(6x) - 10.36\exp(x/8) - 11.2$ não é convexa em $[-2,3]$.
```
# função
p = lambda x: 10*x**2*np.sin(6*x) - 10.36*x*np.exp(x/8) - 11.2
# extremos
pa,pb = p(a),p(b)
# secante
t = pa + (pb - pa)/(b - a)*(x - a)
# plotagem de funções
plt.figure(figsize=(5,3))
plt.plot(x,p(x))
plt.plot(x,t,color='#ffa500')
# pontos da secante
plt.plot(a,p(a),'o',color='#ffa500')
plt.plot(b,p(b),'o',color='#ffa500')
# mínimos locais
xloc = [-1.33868618,0.88811853,1.87451904]
for xl in xloc:
plt.plot(xl,p(xl),'or');
# mínimo global
xmin2 = 2.90547127
plt.plot(xmin2,p(xmin2),'*r',ms=10);
plt.title('Exemplo de função não convexa');
```
### Pontos de sela
- Casos particulares onde a derivada de uma FO anula-se mas o ponto não pode ser definido como de mínimo ou máximo.
- Tais situações implicam a existência dos chamados _pontos de sela_.
- Uma função com um único ponto de sela, por exemplo, não admitirá mínimo global nem mínimo local.
- Para testarmos se um ponto crítico é um ponto de sela, devemos verificar o sinal da segunda derivada da função.
- Uma das seguintes situações deve ser obtida em um ponto crítico $x^*$:
- _ponto de mínimo:_ $f''(x^*) > 0$
- _ponto de máximo:_ $f''(x^*) < 0$
- _ponto de sela:_ $f''(x^*) = 0$
**Exemplo:** qualquer função quadrática admite ou um ponto de mínimo ou de máximo. A função $f(x) = x^3$ possui um ponto de sela em $x^* = 0$.
```
x = np.linspace(-1,1)
plt.figure(figsize=(10,3))
plt.subplot(131)
plt.plot(x,x**2 + 1)
plt.plot(0,1,'r*',ms=10)
plt.title('mínimo global')
plt.subplot(132)
plt.plot(x,-x**2 + 1)
plt.plot(0,1,'r*',ms=10)
plt.title('máximo global')
plt.subplot(133)
plt.plot(x,x**3 + 1)
plt.plot(0,1,'r*',ms=10)
plt.title('ponto de sela');
```
## Otimização univariada
- Otimização univariada visa resolver um problema de minimização tomando uma FO que depende apenas de uma variável.
- Matematicamente:
$$\text{Encontre } x^{*} = \min f(x), \, \text{sujeito a} \, g(x) = 0, h(x) \leq 0.$$
- Em geral, $x$ é uma _variável de decisão_, isto é, uma quantidade que pode ser ajustada livremente (ex. comprimentos, áreas, ângulos etc.).
### Técnicas
- As técnicas utilizadas para a resolução de um problema desse tipo são baseadas em:
- métodos analíticos (busca pelos zeros das derivadas);
- métodos computacionais (determinação de raízes por processos iterativos).
- métodos chamados de _root finding_ são estudados em um curso introdutório de Métodos Numéricos.
- Para dar exemplos com problemas aplicados de otimização univariada, usaremos a abordagem analítica por meio de computação simbólica (módulo `sympy`).
### Problema resolvido
Consideremos o seguinte problema: _maximizar a área do retângulo inscrito em uma elipse._
### Resolução
- Área de um retângulo com vértice esquerdo inferior na origem da elipse e com vértice direito superior no ponto $(x,y)$ da elipse que está no primeiro quadrante é dada por $A_r = xy$.
- Logo, a área do retângulo inscrito na elipse será $A = 4xy$.
- A área $A$ pode ser escrita isolando $y$ na equação da elipse:
$$\frac{x^2}{a^2} + \frac{y^2}{b^2} = 1,$$
### Declaração do problema
$$\text{Encontre } x^{*} = \min \,( -A(x) ), \, \text{sujeito a} \, x > 0.$$
- Notemos que maximizar $A(x)$ equivale a minimizar $-A(x)$.
Primeiramente, criamos variáveis simbólicas que representem as variáveis de interesse do problema e a expressão da área total.
```
# cria variáveis simbólicas
x,y,a,b = sy.symbols('x,y,a,b')
# área do retângulo no 1o. quadrante é xy
# logo, área total é 4xy
A = -4*x*y
A
```
Em seguida, resolvemos a equação da elipse para a variável $y$ utilizando a função `sympy.solve`.
```
# resolve equação da elipse para y
sol = sy.solve(x**2/a**2 + y**2/b**2 - 1,y)
sol[0].expand(),sol[1].expand()
```
Duas soluções são possíveis para $y$. Porém, como o nosso ponto de referência sobre a elipse está no primeiro quadrante, tomamos a expressão para $y > 0$ e a substituímos na expressão da área de forma a obter uma expressão univariada $A(x)$.
```
# substitui expressão de y positivo em A para ter -A(x)
A = A.subs({'y':sol[1]})
A.expand()
```
Localizaremos o ponto crítico da função a partir da derivada $A'(x)$. Derivando $A$ em relação a $x$, obtemos:
```
# deriva -A(x) com a,b constantes
dAdx = A.diff(x)
dAdx.expand()
```
Em seguida, buscamos $x^{*}$ tal que $A'(x^{*}) = \frac{dA}{dx}(x^{*}) = 0$.
```
# resolve A'(x*) = 0
sol_x = sy.solve(dAdx,x)
sol_x
```
Duas soluções, são possíveis, porém, podemos verificar qual ponto de crítico, de fato, é o que minimizará $-A(x)$ através da análise da concavidade. Então, calculamos $A''(x)$, para cada ponto crítico.
```
# testa A''(x) para os dois pontos
dAdx2 = dAdx.diff(x)
dAdx2.subs(x,sol_x[0]).simplify(),dAdx2.subs(x,sol_x[1]).simplify()
```
Uma vez que a segunda solução verifica a concavidade positiva, temos que o ponto crítico $x^{*}$ é:
```
# concavidade para cima => ponto de mínimo
xs = sol_x[1]
xs
```
Usando este valor na equação da elipse, obtemos a ordenada correspondente:
```
# resolve para y > 0
ys = sy.solve(xs**2/a**2 + y**2/b**2 - 1,y)[1]
ys
```
Por fim, substituindo $x^{*}$ na expressão da área, temos que $A_{max}$ é:
```
# área máxima
A_max = A.subs(x,xs)
A_max
```
ou, de forma, simplificada,
```
# simplificando
A_max.simplify()
```
### Conclusão
A área do retângulo inscrito na elipse será máxima quando $x = \frac{\sqrt{2}}{2}a$ e $y = \frac{\sqrt{2}}{2}b$. Portanto, $A_{max} = 2ab$, para comprimentos $a$ e $b$ de semi-eixo maior e menor.
## Estudo paramétrico de geometria
- Plotaremos a variação das áreas de retângulos inscritos em uma elipse arbitrária em função do comprimento $x$ da meia-base do retângulo até o limite da meia-base do retângulo de área máxima.
- Adicionalmente, plotaremos a variação do comprimento da diagonal do retângulo.
```
# semi-eixos da elipse
a,b = 60,30
# no. de retângulos inscritos
nx = 40
# base variável do retângulo
X = np.linspace(0,np.sqrt(2)/2*a,nx)
# área da elipse
e = np.pi*a*b
# áreas dos retângulos
R = []; H = []
for x in X:
y = b*np.sqrt(1 - x**2/a**2)
r = 4*x*y
h = np.hypot(2*x,2*y) # diagonal do retângulo
R.append(r)
H.append(h)
# plotagem
fig,ax1 = plt.subplots(figsize=(6,4))
ax1.plot(X,R,'sb',mec='w',alpha=0.8,label='$A_{ret}(x)$')
ax1.plot(X,np.full(X.shape,2*a*b),'--r',alpha=0.8,label='$A_{max}$')
ax1.plot(X,np.full(X.shape,e),'-',alpha=0.8,label='$A_{elip}$')
ax1.legend(fontsize=10)
# labels
plt.xlabel('$x$ [compr. base ret. inscrito]')
plt.ylabel('$A$ [áreas]');
ax2 = ax1.twinx()
ax2.plot(X,H,'og',mec='w',alpha=0.8,label='$h_{ret}(x)$')
ax2.legend(loc=5,ncol=1,fontsize=10)
plt.ylabel('$h$ [compr. diag ret.]');
plt.suptitle('Variação de áreas e diagonais: elipse x retângulo inscrito\n');
plt.title(f'Elipse: $x^2/({a:.1f})^2 + y^2/({b:.1f})^2 = 1$',fontsize=10);
```
| github_jupyter |
<h1> Create TensorFlow wide-and-deep model </h1>
This notebook illustrates:
<ol>
<li> Creating a model using the high-level Estimator API
</ol>
```
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
%%bash
if ! gsutil ls | grep -q gs://${BUCKET}/; then
gsutil mb -l ${REGION} gs://${BUCKET}
fi
%%bash
ls *.csv
```
<h2> Create TensorFlow model using TensorFlow's Estimator API </h2>
<p>
First, write an input_fn to read the data.
```
import shutil
import numpy as np
import tensorflow as tf
print(tf.__version__)
# Determine CSV, label, and key columns
CSV_COLUMNS = 'weight_pounds,is_male,mother_age,plurality,gestation_weeks,key'.split(',')
LABEL_COLUMN = 'weight_pounds'
KEY_COLUMN = 'key'
# Set default values for each CSV column
DEFAULTS = [[0.0], ['null'], [0.0], ['null'], [0.0], ['nokey']]
TRAIN_STEPS = 1000
# Create an input function reading a file using the Dataset API
# Then provide the results to the Estimator API
def read_dataset(filename, mode, batch_size = 512):
def _input_fn():
def decode_csv(value_column):
columns = tf.decode_csv(value_column, record_defaults=DEFAULTS)
features = dict(zip(CSV_COLUMNS, columns))
label = features.pop(LABEL_COLUMN)
return features, label
# Create list of files that match pattern
file_list = tf.gfile.Glob(filename)
# Create dataset from file list
dataset = (tf.data.TextLineDataset(file_list) # Read text file
.map(decode_csv)) # Transform each elem by applying decode_csv fn
if mode == tf.estimator.ModeKeys.TRAIN:
num_epochs = None # indefinitely
dataset = dataset.shuffle(buffer_size=10*batch_size)
else:
num_epochs = 1 # end-of-input after this
dataset = dataset.repeat(num_epochs).batch(batch_size)
return dataset
return _input_fn
```
Next, define the feature columns
```
# Define feature columns
def get_wide_deep():
# Define column types
is_male,mother_age,plurality,gestation_weeks = \
[\
tf.feature_column.categorical_column_with_vocabulary_list('is_male',
['True', 'False', 'Unknown']),
tf.feature_column.numeric_column('mother_age'),
tf.feature_column.categorical_column_with_vocabulary_list('plurality',
['Single(1)', 'Twins(2)', 'Triplets(3)',
'Quadruplets(4)', 'Quintuplets(5)','Multiple(2+)']),
tf.feature_column.numeric_column('gestation_weeks')
]
# Discretize
age_buckets = tf.feature_column.bucketized_column(mother_age,
boundaries=np.arange(15,45,1).tolist())
gestation_buckets = tf.feature_column.bucketized_column(gestation_weeks,
boundaries=np.arange(17,47,1).tolist())
# Sparse columns are wide, have a linear relationship with the output
wide = [is_male,
plurality,
age_buckets,
gestation_buckets]
# Feature cross all the wide columns and embed into a lower dimension
crossed = tf.feature_column.crossed_column(wide, hash_bucket_size=20000)
embed = tf.feature_column.embedding_column(crossed, 3)
# Continuous columns are deep, have a complex relationship with the output
deep = [mother_age,
gestation_weeks,
embed]
return wide, deep
```
To predict with the TensorFlow model, we also need a serving input function. We will want all the inputs from our user.
```
# Create serving input function to be able to serve predictions later using provided inputs
def serving_input_fn():
feature_placeholders = {
'is_male': tf.placeholder(tf.string, [None]),
'mother_age': tf.placeholder(tf.float32, [None]),
'plurality': tf.placeholder(tf.string, [None]),
'gestation_weeks': tf.placeholder(tf.float32, [None])
}
features = {
key: tf.expand_dims(tensor, -1)
for key, tensor in feature_placeholders.items()
}
return tf.estimator.export.ServingInputReceiver(features, feature_placeholders)
# Create estimator to train and evaluate
def train_and_evaluate(output_dir):
wide, deep = get_wide_deep()
EVAL_INTERVAL = 300
run_config = tf.estimator.RunConfig(save_checkpoints_secs = EVAL_INTERVAL,
keep_checkpoint_max = 3)
estimator = tf.estimator.DNNLinearCombinedRegressor(
model_dir = output_dir,
linear_feature_columns = wide,
dnn_feature_columns = deep,
dnn_hidden_units = [64, 32],
config = run_config)
train_spec = tf.estimator.TrainSpec(
input_fn = read_dataset('train.csv', mode = tf.estimator.ModeKeys.TRAIN),
max_steps = TRAIN_STEPS)
exporter = tf.estimator.LatestExporter('exporter', serving_input_fn)
eval_spec = tf.estimator.EvalSpec(
input_fn = read_dataset('eval.csv', mode = tf.estimator.ModeKeys.EVAL),
steps = None,
start_delay_secs = 60, # start evaluating after N seconds
throttle_secs = EVAL_INTERVAL, # evaluate every N seconds
exporters = exporter)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
```
Finally, train!
```
# Run the model
shutil.rmtree('babyweight_trained', ignore_errors = True) # start fresh each time
tf.summary.FileWriterCache.clear() # ensure filewriter cache is clear for TensorBoard events file
train_and_evaluate('babyweight_trained')
```
The exporter directory contains the final model.
<h2> Monitor and experiment with training </h2>
To begin TensorBoard from within AI Platform Notebooks, click the + symbol in the top left corner and select the **Tensorboard** icon to create a new TensorBoard.
In TensorBoard, look at the learned embeddings. Are they getting clustered? How about the weights for the hidden layers? What if you run this longer? What happens if you change the batchsize?
Copyright 2017-2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| github_jupyter |
# RadarCOVID-Report
## Data Extraction
```
import datetime
import json
import logging
import os
import shutil
import tempfile
import textwrap
import uuid
import matplotlib.pyplot as plt
import matplotlib.ticker
import numpy as np
import pandas as pd
import retry
import seaborn as sns
%matplotlib inline
current_working_directory = os.environ.get("PWD")
if current_working_directory:
os.chdir(current_working_directory)
sns.set()
matplotlib.rcParams["figure.figsize"] = (15, 6)
extraction_datetime = datetime.datetime.utcnow()
extraction_date = extraction_datetime.strftime("%Y-%m-%d")
extraction_previous_datetime = extraction_datetime - datetime.timedelta(days=1)
extraction_previous_date = extraction_previous_datetime.strftime("%Y-%m-%d")
extraction_date_with_hour = datetime.datetime.utcnow().strftime("%Y-%m-%d@%H")
current_hour = datetime.datetime.utcnow().hour
are_today_results_partial = current_hour != 23
```
### Constants
```
from Modules.ExposureNotification import exposure_notification_io
spain_region_country_code = "ES"
germany_region_country_code = "DE"
default_backend_identifier = spain_region_country_code
backend_generation_days = 7 * 2
daily_summary_days = 7 * 4 * 3
daily_plot_days = 7 * 4
tek_dumps_load_limit = daily_summary_days + 1
default_shared_diagnosis_generation_to_upload_days = 1
```
### Parameters
```
environment_backend_identifier = os.environ.get("RADARCOVID_REPORT__BACKEND_IDENTIFIER")
if environment_backend_identifier:
report_backend_identifier = environment_backend_identifier
else:
report_backend_identifier = default_backend_identifier
report_backend_identifier
environment_download_only_from_report_backend = \
os.environ.get("RADARCOVID_REPORT__DOWNLOAD_ONLY_FROM_REPORT_BACKEND")
if environment_download_only_from_report_backend:
report_backend_identifiers = [report_backend_identifier]
else:
report_backend_identifiers = None
report_backend_identifiers
environment_shared_diagnosis_generation_to_upload_days = \
os.environ.get("RADARCOVID_REPORT__SHARED_DIAGNOSIS_GENERATION_TO_UPLOAD_DAYS")
if environment_shared_diagnosis_generation_to_upload_days:
shared_diagnosis_generation_to_upload_days = \
int(environment_shared_diagnosis_generation_to_upload_days)
else:
shared_diagnosis_generation_to_upload_days = \
default_shared_diagnosis_generation_to_upload_days
shared_diagnosis_generation_to_upload_days
```
### COVID-19 Cases
```
report_backend_client = \
exposure_notification_io.get_backend_client_with_identifier(
backend_identifier=report_backend_identifier)
@retry.retry(tries=10, delay=10, backoff=1.1, jitter=(0, 10))
def download_cases_dataframe_from_ecdc():
return pd.read_csv(
"https://opendata.ecdc.europa.eu/covid19/casedistribution/csv/data.csv")
confirmed_df_ = download_cases_dataframe_from_ecdc()
confirmed_df = confirmed_df_.copy()
confirmed_df = confirmed_df[["dateRep", "cases", "geoId"]]
confirmed_df.rename(
columns={
"dateRep":"sample_date",
"cases": "new_cases",
"geoId": "country_code",
},
inplace=True)
confirmed_df["sample_date"] = pd.to_datetime(confirmed_df.sample_date, dayfirst=True)
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_days = pd.date_range(
start=confirmed_df.iloc[0].sample_date,
end=extraction_datetime)
source_regions_at_date_df = pd.DataFrame(data=confirmed_days, columns=["sample_date"])
source_regions_at_date_df["source_regions_at_date"] = \
source_regions_at_date_df.sample_date.apply(
lambda x: report_backend_client.source_regions_for_date(date=x))
source_regions_at_date_df.sort_values("sample_date", inplace=True)
source_regions_at_date_df.tail()
confirmed_df = source_regions_at_date_df.merge(confirmed_df, how="left")
confirmed_df = confirmed_df[confirmed_df.apply(
lambda x: x.country_code in x.source_regions_at_date, axis=1)]
confirmed_df.drop(columns=["source_regions_at_date"], inplace=True)
confirmed_df = source_regions_at_date_df.merge(confirmed_df, how="left")
confirmed_df["sample_date"] = confirmed_df.sample_date.dt.strftime("%Y-%m-%d")
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
report_source_regions = list(sorted(confirmed_df.country_code.dropna().unique().tolist()))
if report_backend_identifier in report_source_regions:
report_source_regions = [report_backend_identifier] + \
list(sorted(set(report_source_regions).difference([report_backend_identifier])))
report_source_regions
confirmed_df = confirmed_df.groupby(["sample_date"]).new_cases.sum(min_count=1).reset_index()
confirmed_df.sort_values("sample_date", inplace=True)
confirmed_df.tail()
confirmed_df.rename(columns={"sample_date": "sample_date_string"}, inplace=True)
confirmed_df = confirmed_df[["sample_date_string", "new_cases"]]
confirmed_df.sort_values("sample_date_string", inplace=True)
confirmed_df["covid_cases"] = confirmed_df.new_cases.rolling(7).mean().round()
confirmed_df.fillna(method="ffill", inplace=True)
confirmed_df.tail()
confirmed_df[["new_cases", "covid_cases"]].plot()
```
### Extract API TEKs
```
raw_zip_path_prefix = "Data/TEKs/Raw/"
fail_on_error_backend_identifiers = [report_backend_identifier]
multi_backend_exposure_keys_df = \
exposure_notification_io.download_exposure_keys_from_backends(
backend_identifiers=report_backend_identifiers,
generation_days=backend_generation_days,
fail_on_error_backend_identifiers=fail_on_error_backend_identifiers,
save_raw_zip_path_prefix=raw_zip_path_prefix)
multi_backend_exposure_keys_df["region"] = multi_backend_exposure_keys_df["backend_identifier"]
multi_backend_exposure_keys_df.rename(
columns={
"generation_datetime": "sample_datetime",
"generation_date_string": "sample_date_string",
},
inplace=True)
multi_backend_exposure_keys_df.head()
early_teks_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.rolling_period < 144].copy()
early_teks_df["rolling_period_in_hours"] = early_teks_df.rolling_period / 6
early_teks_df[early_teks_df.sample_date_string != extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
early_teks_df[early_teks_df.sample_date_string == extraction_date] \
.rolling_period_in_hours.hist(bins=list(range(24)))
multi_backend_exposure_keys_df = multi_backend_exposure_keys_df[[
"sample_date_string", "region", "key_data"]]
multi_backend_exposure_keys_df.head()
active_regions = \
multi_backend_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
active_regions
multi_backend_summary_df = multi_backend_exposure_keys_df.groupby(
["sample_date_string", "region"]).key_data.nunique().reset_index() \
.pivot(index="sample_date_string", columns="region") \
.sort_index(ascending=False)
multi_backend_summary_df.rename(
columns={"key_data": "shared_teks_by_generation_date"},
inplace=True)
multi_backend_summary_df.rename_axis("sample_date", inplace=True)
multi_backend_summary_df = multi_backend_summary_df.fillna(0).astype(int)
multi_backend_summary_df = multi_backend_summary_df.head(backend_generation_days)
multi_backend_summary_df.head()
def compute_keys_cross_sharing(x):
teks_x = x.key_data_x.item()
common_teks = set(teks_x).intersection(x.key_data_y.item())
common_teks_fraction = len(common_teks) / len(teks_x)
return pd.Series(dict(
common_teks=common_teks,
common_teks_fraction=common_teks_fraction,
))
multi_backend_exposure_keys_by_region_df = \
multi_backend_exposure_keys_df.groupby("region").key_data.unique().reset_index()
multi_backend_exposure_keys_by_region_df["_merge"] = True
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_df.merge(
multi_backend_exposure_keys_by_region_df, on="_merge")
multi_backend_exposure_keys_by_region_combination_df.drop(
columns=["_merge"], inplace=True)
if multi_backend_exposure_keys_by_region_combination_df.region_x.nunique() > 1:
multi_backend_exposure_keys_by_region_combination_df = \
multi_backend_exposure_keys_by_region_combination_df[
multi_backend_exposure_keys_by_region_combination_df.region_x !=
multi_backend_exposure_keys_by_region_combination_df.region_y]
multi_backend_exposure_keys_cross_sharing_df = \
multi_backend_exposure_keys_by_region_combination_df \
.groupby(["region_x", "region_y"]) \
.apply(compute_keys_cross_sharing) \
.reset_index()
multi_backend_cross_sharing_summary_df = \
multi_backend_exposure_keys_cross_sharing_df.pivot_table(
values=["common_teks_fraction"],
columns="region_x",
index="region_y",
aggfunc=lambda x: x.item())
multi_backend_cross_sharing_summary_df
multi_backend_without_active_region_exposure_keys_df = \
multi_backend_exposure_keys_df[multi_backend_exposure_keys_df.region != report_backend_identifier]
multi_backend_without_active_region = \
multi_backend_without_active_region_exposure_keys_df.groupby("region").key_data.nunique().sort_values().index.unique().tolist()
multi_backend_without_active_region
exposure_keys_summary_df = multi_backend_exposure_keys_df[
multi_backend_exposure_keys_df.region == report_backend_identifier]
exposure_keys_summary_df.drop(columns=["region"], inplace=True)
exposure_keys_summary_df = \
exposure_keys_summary_df.groupby(["sample_date_string"]).key_data.nunique().to_frame()
exposure_keys_summary_df = \
exposure_keys_summary_df.reset_index().set_index("sample_date_string")
exposure_keys_summary_df.sort_index(ascending=False, inplace=True)
exposure_keys_summary_df.rename(columns={"key_data": "shared_teks_by_generation_date"}, inplace=True)
exposure_keys_summary_df.head()
```
### Dump API TEKs
```
tek_list_df = multi_backend_exposure_keys_df[
["sample_date_string", "region", "key_data"]].copy()
tek_list_df["key_data"] = tek_list_df["key_data"].apply(str)
tek_list_df.rename(columns={
"sample_date_string": "sample_date",
"key_data": "tek_list"}, inplace=True)
tek_list_df = tek_list_df.groupby(
["sample_date", "region"]).tek_list.unique().reset_index()
tek_list_df["extraction_date"] = extraction_date
tek_list_df["extraction_date_with_hour"] = extraction_date_with_hour
tek_list_path_prefix = "Data/TEKs/"
tek_list_current_path = tek_list_path_prefix + f"/Current/RadarCOVID-TEKs.json"
tek_list_daily_path = tek_list_path_prefix + f"Daily/RadarCOVID-TEKs-{extraction_date}.json"
tek_list_hourly_path = tek_list_path_prefix + f"Hourly/RadarCOVID-TEKs-{extraction_date_with_hour}.json"
for path in [tek_list_current_path, tek_list_daily_path, tek_list_hourly_path]:
os.makedirs(os.path.dirname(path), exist_ok=True)
tek_list_df.drop(columns=["extraction_date", "extraction_date_with_hour"]).to_json(
tek_list_current_path,
lines=True, orient="records")
tek_list_df.drop(columns=["extraction_date_with_hour"]).to_json(
tek_list_daily_path,
lines=True, orient="records")
tek_list_df.to_json(
tek_list_hourly_path,
lines=True, orient="records")
tek_list_df.head()
```
### Load TEK Dumps
```
import glob
def load_extracted_teks(mode, region=None, limit=None) -> pd.DataFrame:
extracted_teks_df = pd.DataFrame(columns=["region"])
file_paths = list(reversed(sorted(glob.glob(tek_list_path_prefix + mode + "/RadarCOVID-TEKs-*.json"))))
if limit:
file_paths = file_paths[:limit]
for file_path in file_paths:
logging.info(f"Loading TEKs from '{file_path}'...")
iteration_extracted_teks_df = pd.read_json(file_path, lines=True)
extracted_teks_df = extracted_teks_df.append(
iteration_extracted_teks_df, sort=False)
extracted_teks_df["region"] = \
extracted_teks_df.region.fillna(spain_region_country_code).copy()
if region:
extracted_teks_df = \
extracted_teks_df[extracted_teks_df.region == region]
return extracted_teks_df
daily_extracted_teks_df = load_extracted_teks(
mode="Daily",
region=report_backend_identifier,
limit=tek_dumps_load_limit)
daily_extracted_teks_df.head()
exposure_keys_summary_df_ = daily_extracted_teks_df \
.sort_values("extraction_date", ascending=False) \
.groupby("sample_date").tek_list.first() \
.to_frame()
exposure_keys_summary_df_.index.name = "sample_date_string"
exposure_keys_summary_df_["tek_list"] = \
exposure_keys_summary_df_.tek_list.apply(len)
exposure_keys_summary_df_ = exposure_keys_summary_df_ \
.rename(columns={"tek_list": "shared_teks_by_generation_date"}) \
.sort_index(ascending=False)
exposure_keys_summary_df = exposure_keys_summary_df_
exposure_keys_summary_df.head()
```
### Daily New TEKs
```
tek_list_df = daily_extracted_teks_df.groupby("extraction_date").tek_list.apply(
lambda x: set(sum(x, []))).reset_index()
tek_list_df = tek_list_df.set_index("extraction_date").sort_index(ascending=True)
tek_list_df.head()
def compute_teks_by_generation_and_upload_date(date):
day_new_teks_set_df = tek_list_df.copy().diff()
try:
day_new_teks_set = day_new_teks_set_df[
day_new_teks_set_df.index == date].tek_list.item()
except ValueError:
day_new_teks_set = None
if pd.isna(day_new_teks_set):
day_new_teks_set = set()
day_new_teks_df = daily_extracted_teks_df[
daily_extracted_teks_df.extraction_date == date].copy()
day_new_teks_df["shared_teks"] = \
day_new_teks_df.tek_list.apply(lambda x: set(x).intersection(day_new_teks_set))
day_new_teks_df["shared_teks"] = \
day_new_teks_df.shared_teks.apply(len)
day_new_teks_df["upload_date"] = date
day_new_teks_df.rename(columns={"sample_date": "generation_date"}, inplace=True)
day_new_teks_df = day_new_teks_df[
["upload_date", "generation_date", "shared_teks"]]
day_new_teks_df["generation_to_upload_days"] = \
(pd.to_datetime(day_new_teks_df.upload_date) -
pd.to_datetime(day_new_teks_df.generation_date)).dt.days
day_new_teks_df = day_new_teks_df[day_new_teks_df.shared_teks > 0]
return day_new_teks_df
shared_teks_generation_to_upload_df = pd.DataFrame()
for upload_date in daily_extracted_teks_df.extraction_date.unique():
shared_teks_generation_to_upload_df = \
shared_teks_generation_to_upload_df.append(
compute_teks_by_generation_and_upload_date(date=upload_date))
shared_teks_generation_to_upload_df \
.sort_values(["upload_date", "generation_date"], ascending=False, inplace=True)
shared_teks_generation_to_upload_df.tail()
today_new_teks_df = \
shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.upload_date == extraction_date].copy()
today_new_teks_df.tail()
if not today_new_teks_df.empty:
today_new_teks_df.set_index("generation_to_upload_days") \
.sort_index().shared_teks.plot.bar()
generation_to_upload_period_pivot_df = \
shared_teks_generation_to_upload_df[
["upload_date", "generation_to_upload_days", "shared_teks"]] \
.pivot(index="upload_date", columns="generation_to_upload_days") \
.sort_index(ascending=False).fillna(0).astype(int) \
.droplevel(level=0, axis=1)
generation_to_upload_period_pivot_df.head()
new_tek_df = tek_list_df.diff().tek_list.apply(
lambda x: len(x) if not pd.isna(x) else None).to_frame().reset_index()
new_tek_df.rename(columns={
"tek_list": "shared_teks_by_upload_date",
"extraction_date": "sample_date_string",}, inplace=True)
new_tek_df.tail()
shared_teks_uploaded_on_generation_date_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == 0] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_teks_uploaded_on_generation_date",
})
shared_teks_uploaded_on_generation_date_df.head()
estimated_shared_diagnoses_df = shared_teks_generation_to_upload_df[
shared_teks_generation_to_upload_df.generation_to_upload_days == shared_diagnosis_generation_to_upload_days] \
[["upload_date", "shared_teks"]].rename(
columns={
"upload_date": "sample_date_string",
"shared_teks": "shared_diagnoses",
})
estimated_shared_diagnoses_df.head()
```
### Hourly New TEKs
```
hourly_extracted_teks_df = load_extracted_teks(
mode="Hourly", region=report_backend_identifier, limit=25)
hourly_extracted_teks_df.head()
hourly_new_tek_count_df = hourly_extracted_teks_df \
.groupby("extraction_date_with_hour").tek_list. \
apply(lambda x: set(sum(x, []))).reset_index().copy()
hourly_new_tek_count_df = hourly_new_tek_count_df.set_index("extraction_date_with_hour") \
.sort_index(ascending=True)
hourly_new_tek_count_df["new_tek_list"] = hourly_new_tek_count_df.tek_list.diff()
hourly_new_tek_count_df["new_tek_count"] = hourly_new_tek_count_df.new_tek_list.apply(
lambda x: len(x) if not pd.isna(x) else 0)
hourly_new_tek_count_df.rename(columns={
"new_tek_count": "shared_teks_by_upload_date"}, inplace=True)
hourly_new_tek_count_df = hourly_new_tek_count_df.reset_index()[[
"extraction_date_with_hour", "shared_teks_by_upload_date"]]
hourly_new_tek_count_df.head()
hourly_summary_df = hourly_new_tek_count_df.copy()
hourly_summary_df.set_index("extraction_date_with_hour", inplace=True)
hourly_summary_df = hourly_summary_df.fillna(0).astype(int).reset_index()
hourly_summary_df["datetime_utc"] = pd.to_datetime(
hourly_summary_df.extraction_date_with_hour, format="%Y-%m-%d@%H")
hourly_summary_df.set_index("datetime_utc", inplace=True)
hourly_summary_df = hourly_summary_df.tail(-1)
hourly_summary_df.head()
```
### Data Merge
```
result_summary_df = exposure_keys_summary_df.merge(
new_tek_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
shared_teks_uploaded_on_generation_date_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = result_summary_df.merge(
estimated_shared_diagnoses_df, on=["sample_date_string"], how="outer")
result_summary_df.head()
result_summary_df = confirmed_df.tail(daily_summary_days).merge(
result_summary_df, on=["sample_date_string"], how="left")
result_summary_df.head()
result_summary_df["sample_date"] = pd.to_datetime(result_summary_df.sample_date_string)
result_summary_df.set_index("sample_date", inplace=True)
result_summary_df.drop(columns=["sample_date_string"], inplace=True)
result_summary_df.sort_index(ascending=False, inplace=True)
result_summary_df.head()
with pd.option_context("mode.use_inf_as_na", True):
result_summary_df = result_summary_df.fillna(0).astype(int)
result_summary_df["teks_per_shared_diagnosis"] = \
(result_summary_df.shared_teks_by_upload_date / result_summary_df.shared_diagnoses).fillna(0)
result_summary_df["shared_diagnoses_per_covid_case"] = \
(result_summary_df.shared_diagnoses / result_summary_df.covid_cases).fillna(0)
result_summary_df.head(daily_plot_days)
weekly_result_summary_df = result_summary_df \
.sort_index(ascending=True).fillna(0).rolling(7).agg({
"covid_cases": "sum",
"shared_teks_by_generation_date": "sum",
"shared_teks_by_upload_date": "sum",
"shared_diagnoses": "sum"
}).sort_index(ascending=False)
with pd.option_context("mode.use_inf_as_na", True):
weekly_result_summary_df = weekly_result_summary_df.fillna(0).astype(int)
weekly_result_summary_df["teks_per_shared_diagnosis"] = \
(weekly_result_summary_df.shared_teks_by_upload_date / weekly_result_summary_df.shared_diagnoses).fillna(0)
weekly_result_summary_df["shared_diagnoses_per_covid_case"] = \
(weekly_result_summary_df.shared_diagnoses / weekly_result_summary_df.covid_cases).fillna(0)
weekly_result_summary_df.head()
last_7_days_summary = weekly_result_summary_df.to_dict(orient="records")[1]
last_7_days_summary
```
## Report Results
```
display_column_name_mapping = {
"sample_date": "Sample\u00A0Date\u00A0(UTC)",
"datetime_utc": "Timestamp (UTC)",
"upload_date": "Upload Date (UTC)",
"generation_to_upload_days": "Generation to Upload Period in Days",
"region": "Backend",
"region_x": "Backend\u00A0(A)",
"region_y": "Backend\u00A0(B)",
"common_teks": "Common TEKs Shared Between Backends",
"common_teks_fraction": "Fraction of TEKs in Backend (A) Available in Backend (B)",
"covid_cases": "COVID-19 Cases in Source Countries (7-day Rolling Average)",
"shared_teks_by_generation_date": "Shared TEKs by Generation Date",
"shared_teks_by_upload_date": "Shared TEKs by Upload Date",
"shared_diagnoses": "Shared Diagnoses (Estimation)",
"teks_per_shared_diagnosis": "TEKs Uploaded per Shared Diagnosis",
"shared_diagnoses_per_covid_case": "Usage Ratio (Fraction of Cases in Source Countries Which Shared Diagnosis)",
"shared_teks_uploaded_on_generation_date": "Shared TEKs Uploaded on Generation Date",
}
summary_columns = [
"covid_cases",
"shared_teks_by_generation_date",
"shared_teks_by_upload_date",
"shared_teks_uploaded_on_generation_date",
"shared_diagnoses",
"teks_per_shared_diagnosis",
"shared_diagnoses_per_covid_case",
]
```
### Daily Summary Table
```
result_summary_df_ = result_summary_df.copy()
result_summary_df = result_summary_df[summary_columns]
result_summary_with_display_names_df = result_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
result_summary_with_display_names_df
```
### Daily Summary Plots
```
result_plot_summary_df = result_summary_df.head(daily_plot_days)[summary_columns] \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping)
summary_ax_list = result_plot_summary_df.sort_index(ascending=True).plot.bar(
title=f"Daily Summary",
rot=45, subplots=True, figsize=(15, 22), legend=False)
ax_ = summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.95)
ax_.yaxis.set_major_formatter(matplotlib.ticker.PercentFormatter(1.0))
_ = ax_.set_xticklabels(sorted(result_plot_summary_df.index.strftime("%Y-%m-%d").tolist()))
```
### Daily Generation to Upload Period Table
```
display_generation_to_upload_period_pivot_df = \
generation_to_upload_period_pivot_df \
.head(backend_generation_days)
display_generation_to_upload_period_pivot_df \
.head(backend_generation_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping)
fig, generation_to_upload_period_pivot_table_ax = plt.subplots(
figsize=(10, 1 + 0.5 * len(display_generation_to_upload_period_pivot_df)))
generation_to_upload_period_pivot_table_ax.set_title(
"Shared TEKs Generation to Upload Period Table")
sns.heatmap(
data=display_generation_to_upload_period_pivot_df
.rename_axis(columns=display_column_name_mapping)
.rename_axis(index=display_column_name_mapping),
fmt=".0f",
annot=True,
ax=generation_to_upload_period_pivot_table_ax)
generation_to_upload_period_pivot_table_ax.get_figure().tight_layout()
```
### Hourly Summary Plots
```
hourly_summary_ax_list = hourly_summary_df \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.plot.bar(
title=f"Last 24h Summary",
rot=45, subplots=True, legend=False)
ax_ = hourly_summary_ax_list[-1]
ax_.get_figure().tight_layout()
ax_.get_figure().subplots_adjust(top=0.9)
_ = ax_.set_xticklabels(sorted(hourly_summary_df.index.strftime("%Y-%m-%d@%H").tolist()))
```
### Publish Results
```
def get_temporary_image_path() -> str:
return os.path.join(tempfile.gettempdir(), str(uuid.uuid4()) + ".png")
def save_temporary_plot_image(ax):
if isinstance(ax, np.ndarray):
ax = ax[0]
media_path = get_temporary_image_path()
ax.get_figure().savefig(media_path)
return media_path
def save_temporary_dataframe_image(df):
import dataframe_image as dfi
media_path = get_temporary_image_path()
dfi.export(df, media_path)
return media_path
github_repository = os.environ.get("GITHUB_REPOSITORY")
if github_repository is None:
github_repository = "pvieito/Radar-STATS"
github_project_base_url = "https://github.com/" + github_repository
display_formatters = {
display_column_name_mapping["teks_per_shared_diagnosis"]: lambda x: f"{x:.2f}",
display_column_name_mapping["shared_diagnoses_per_covid_case"]: lambda x: f"{x:.2%}",
}
daily_summary_table_html = result_summary_with_display_names_df \
.head(daily_plot_days) \
.rename_axis(index=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.to_html(formatters=display_formatters)
multi_backend_summary_table_html = multi_backend_summary_df \
.head(daily_plot_days) \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(formatters=display_formatters)
def format_multi_backend_cross_sharing_fraction(x):
if pd.isna(x):
return "-"
elif round(x * 100, 1) == 0:
return ""
else:
return f"{x:.1%}"
multi_backend_cross_sharing_summary_table_html = multi_backend_cross_sharing_summary_df \
.rename_axis(columns=display_column_name_mapping) \
.rename(columns=display_column_name_mapping) \
.rename_axis(index=display_column_name_mapping) \
.to_html(
classes="table-center",
formatters=display_formatters,
float_format=format_multi_backend_cross_sharing_fraction)
multi_backend_cross_sharing_summary_table_html = \
multi_backend_cross_sharing_summary_table_html \
.replace("<tr>","<tr style=\"text-align: center;\">")
extraction_date_result_summary_df = \
result_summary_df[result_summary_df.index == extraction_date]
extraction_date_result_hourly_summary_df = \
hourly_summary_df[hourly_summary_df.extraction_date_with_hour == extraction_date_with_hour]
covid_cases = \
extraction_date_result_summary_df.covid_cases.sum()
shared_teks_by_generation_date = \
extraction_date_result_summary_df.shared_teks_by_generation_date.sum()
shared_teks_by_upload_date = \
extraction_date_result_summary_df.shared_teks_by_upload_date.sum()
shared_diagnoses = \
extraction_date_result_summary_df.shared_diagnoses.sum()
teks_per_shared_diagnosis = \
extraction_date_result_summary_df.teks_per_shared_diagnosis.sum()
shared_diagnoses_per_covid_case = \
extraction_date_result_summary_df.shared_diagnoses_per_covid_case.sum()
shared_teks_by_upload_date_last_hour = \
extraction_date_result_hourly_summary_df.shared_teks_by_upload_date.sum().astype(int)
display_source_regions = ", ".join(report_source_regions)
display_brief_source_regions_limit = 2
if len(report_source_regions) <= display_brief_source_regions_limit:
display_brief_source_regions = display_source_regions
else:
prefix_countries = ", ".join(report_source_regions[:display_brief_source_regions_limit])
display_brief_source_regions = f"{len(report_source_regions)} ({prefix_countries}…)"
if len(report_source_regions) == 1:
display_brief_source_regions_warning_prefix_message = ""
else:
display_brief_source_regions_warning_prefix_message = "⚠️ "
summary_plots_image_path = save_temporary_plot_image(
ax=summary_ax_list)
summary_table_image_path = save_temporary_dataframe_image(
df=result_summary_with_display_names_df)
hourly_summary_plots_image_path = save_temporary_plot_image(
ax=hourly_summary_ax_list)
multi_backend_summary_table_image_path = save_temporary_dataframe_image(
df=multi_backend_summary_df)
generation_to_upload_period_pivot_table_image_path = save_temporary_plot_image(
ax=generation_to_upload_period_pivot_table_ax)
```
### Save Results
```
report_resources_path_prefix = "Data/Resources/Current/RadarCOVID-Report-"
result_summary_df.to_csv(
report_resources_path_prefix + "Summary-Table.csv")
result_summary_df.to_html(
report_resources_path_prefix + "Summary-Table.html")
hourly_summary_df.to_csv(
report_resources_path_prefix + "Hourly-Summary-Table.csv")
multi_backend_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Summary-Table.csv")
multi_backend_cross_sharing_summary_df.to_csv(
report_resources_path_prefix + "Multi-Backend-Cross-Sharing-Summary-Table.csv")
generation_to_upload_period_pivot_df.to_csv(
report_resources_path_prefix + "Generation-Upload-Period-Table.csv")
_ = shutil.copyfile(
summary_plots_image_path,
report_resources_path_prefix + "Summary-Plots.png")
_ = shutil.copyfile(
summary_table_image_path,
report_resources_path_prefix + "Summary-Table.png")
_ = shutil.copyfile(
hourly_summary_plots_image_path,
report_resources_path_prefix + "Hourly-Summary-Plots.png")
_ = shutil.copyfile(
multi_backend_summary_table_image_path,
report_resources_path_prefix + "Multi-Backend-Summary-Table.png")
_ = shutil.copyfile(
generation_to_upload_period_pivot_table_image_path,
report_resources_path_prefix + "Generation-Upload-Period-Table.png")
```
### Publish Results as JSON
```
summary_results_api_df = result_summary_df.reset_index()
summary_results_api_df["sample_date_string"] = \
summary_results_api_df["sample_date"].dt.strftime("%Y-%m-%d")
summary_results = dict(
source_regions=report_source_regions,
extraction_datetime=extraction_datetime,
extraction_date=extraction_date,
extraction_date_with_hour=extraction_date_with_hour,
last_hour=dict(
shared_teks_by_upload_date=shared_teks_by_upload_date_last_hour,
shared_diagnoses=0,
),
today=dict(
covid_cases=covid_cases,
shared_teks_by_generation_date=shared_teks_by_generation_date,
shared_teks_by_upload_date=shared_teks_by_upload_date,
shared_diagnoses=shared_diagnoses,
teks_per_shared_diagnosis=teks_per_shared_diagnosis,
shared_diagnoses_per_covid_case=shared_diagnoses_per_covid_case,
),
last_7_days=last_7_days_summary,
daily_results=summary_results_api_df.to_dict(orient="records"))
summary_results = \
json.loads(pd.Series([summary_results]).to_json(orient="records"))[0]
with open(report_resources_path_prefix + "Summary-Results.json", "w") as f:
json.dump(summary_results, f, indent=4)
```
### Publish on README
```
with open("Data/Templates/README.md", "r") as f:
readme_contents = f.read()
readme_contents = readme_contents.format(
extraction_date_with_hour=extraction_date_with_hour,
github_project_base_url=github_project_base_url,
daily_summary_table_html=daily_summary_table_html,
multi_backend_summary_table_html=multi_backend_summary_table_html,
multi_backend_cross_sharing_summary_table_html=multi_backend_cross_sharing_summary_table_html,
display_source_regions=display_source_regions)
with open("README.md", "w") as f:
f.write(readme_contents)
```
### Publish on Twitter
```
enable_share_to_twitter = os.environ.get("RADARCOVID_REPORT__ENABLE_PUBLISH_ON_TWITTER")
github_event_name = os.environ.get("GITHUB_EVENT_NAME")
if enable_share_to_twitter and github_event_name == "schedule" and \
(shared_teks_by_upload_date_last_hour or not are_today_results_partial):
import tweepy
twitter_api_auth_keys = os.environ["RADARCOVID_REPORT__TWITTER_API_AUTH_KEYS"]
twitter_api_auth_keys = twitter_api_auth_keys.split(":")
auth = tweepy.OAuthHandler(twitter_api_auth_keys[0], twitter_api_auth_keys[1])
auth.set_access_token(twitter_api_auth_keys[2], twitter_api_auth_keys[3])
api = tweepy.API(auth)
summary_plots_media = api.media_upload(summary_plots_image_path)
summary_table_media = api.media_upload(summary_table_image_path)
generation_to_upload_period_pivot_table_image_media = api.media_upload(generation_to_upload_period_pivot_table_image_path)
media_ids = [
summary_plots_media.media_id,
summary_table_media.media_id,
generation_to_upload_period_pivot_table_image_media.media_id,
]
if are_today_results_partial:
today_addendum = " (Partial)"
else:
today_addendum = ""
status = textwrap.dedent(f"""
#RadarCOVID – {extraction_date_with_hour}
{display_brief_source_regions_warning_prefix_message}Countries: {display_brief_source_regions}
Today{today_addendum}:
- Uploaded TEKs: {shared_teks_by_upload_date:.0f} ({shared_teks_by_upload_date_last_hour:+d} last hour)
- Shared Diagnoses: ≤{shared_diagnoses:.0f}
- Usage Ratio: ≤{shared_diagnoses_per_covid_case:.2%}
Last 7 Days:
- Shared Diagnoses: ≤{last_7_days_summary["shared_diagnoses"]:.0f}
- Usage Ratio: ≤{last_7_days_summary["shared_diagnoses_per_covid_case"]:.2%}
Info: {github_project_base_url}#documentation
""")
status = status.encode(encoding="utf-8")
api.update_status(status=status, media_ids=media_ids)
```
| github_jupyter |
# Introduction to Hyperopt for Optimizing Neural Networks
## What is Hyperopt?
Hyperopt is a way to search through an hyperparameter space. For example, it can use the [Tree-structured Parzen Estimator (TPE) algorithm](https://pdfs.semanticscholar.org/9f2a/efc3821853e963beda011ed770f740385b77.pdf), which explore intelligently the search space while narrowing down to the estimated best parameters.
It is hence a good method for meta-optimizing a neural network which is itself an optimisation problem: tuning a neural network uses gradient descent methods, and tuning the hyperparameters needs to be done differently since gradient descent can't apply. Therefore, Hyperopt can be useful not only for tuning hyperparameters such as the learning rate, but also to tune more fancy parameters in a flexible way, such as changing the number of layers of certain types, or the number of neurons in a layer, or even the type of layer to use at a certain place in the network given an array of choices, each with nested tunable hyperparameters.
This is an oriented random search, in contrast with a Grid Search where hyperparameters are pre-established with fixed steps increase. [Random Search for Hyper-Parameter Optimization](http://www.jmlr.org/papers/volume13/bergstra12a/bergstra12a) (such as what Hyperopt do) has proven to be an effective search technique. The paper about this technique [sits among the most cited deep learning papers](https://github.com/terryum/awesome-deep-learning-papers#optimization--training-techniques). To sum up, it is more efficient to search randomly through values and to intelligently narrow the search space rather than looping on fixed sets of values for the hyperparameters.
## How to define Hyperopt parameters?
A parameter is defined with a certain uniformrange or else a probability distribution, such as:
- hp.randint(label, upper)
- hp.uniform(label, low, high)
- hp.loguniform(label, low, high)
- hp.normal(label, mu, sigma)
- hp.lognormal(label, mu, sigma)
There is also a few quantized versions of those functions, which rounds the generated values at each step of "q":
- hp.quniform(label, low, high, q)
- hp.qloguniform(label, low, high, q)
- hp.qnormal(label, mu, sigma, q)
- hp.qlognormal(label, mu, sigma, q)
It is also possible to use a "choice" which can lead to hyperparameter nesting:
- hp.choice(label, ["list", "of", "potential", "choices"])
- hp.choice(label, [hp.uniform(sub_label_1, low, high), hp.normal(sub_label_2, mu, sigma), None, 0, 1, "anything"])
Visualisations of the parameters for probability distributions can be found below. Then, more details on choices and parameter nesting will come.
```
# The "%reset" ipython magic command will reset the kernel upon being called.
# (thus flushing loaded variables and previous imports from the current
# notebook session, if there were any)
%reset -f
from hyperopt import pyll, hp
import matplotlib.pyplot as plt
import numpy as np
from scipy.stats.kde import gaussian_kde
# Let's plot the result of sampling from many different probability distributions:
hyperparam_generators = {
'randint': hp.randint('randint', 5),
'uniform': hp.uniform('uniform', -1, 3),
'loguniform': hp.loguniform('loguniform', -0.3, 0.3),
'normal': hp.normal('normal', 1, 2),
'lognormal': hp.lognormal('lognormal', 0, 0.3)
}
n_samples = 5000
for title, space in hyperparam_generators.items():
evaluated = [
pyll.stochastic.sample(space) for _ in range(n_samples)
]
x_domain = np.linspace(min(evaluated), max(evaluated), n_samples)
plt.figure(figsize=(18,6))
hist = gaussian_kde(evaluated, 0.001)
plt.plot(x_domain, hist(x_domain), label="True Histogram")
blurred_hist = gaussian_kde(evaluated, 0.1)
plt.plot(x_domain, blurred_hist(x_domain), label="Smoothed Histogram")
plt.title("Histogram (pdf) for a {} distribution".format(title))
plt.legend()
plt.show()
```
Note on the above charts (especially for the loguniform and uniform distributions): the blurred line averaging the values fades out toward the ends of the signal since it is zero-padded. The line ideally would not fade out by using techniques such as mirror-padding.
## On the loguniform and lognormal distributions
Those are the best distributions for modeling the values a learning rate. That's because we want to observe changes in the learning rate according to changing it with multiplications rather than additions, e.g.: when adjusting the learning rate, we'll want to try to divide it or multiply it by 2 rather than adding and substracting a finite value.
To proove this, let's generate a loguniform distribution for a multiplier of the learning rate, centered at 1.0. Dividing 1 by those values should yield the same distribution.
```
log_hyperparam_generators = {
'loguniform': hp.loguniform('loguniform', -0.3, 0.3),
'lognormal': hp.lognormal('lognormal', 0, 0.3)
}
# For more info about the lognormal distribution, see:
# https://www.wolframalpha.com/input/?i=y%3D2%5E(+(-log4(x))%5E0.5+),+y%3D2%5E(-+(-log4(x))%5E0.5+)+from+0+to+1
# https://www.wolframalpha.com/input/?i=y%3D4%5E-(log2(x)%5E2)+from+0+to+5
n_samples = 5000
for title, space in log_hyperparam_generators.items():
evaluated = [
pyll.stochastic.sample(space) for _ in range(n_samples)
]
inverse_evaluated = [1.0 / y for y in evaluated]
x_domain = np.linspace(min(evaluated), max(evaluated), n_samples)
plt.figure(figsize=(18,6))
hist = gaussian_kde(evaluated, 0.001)
plt.plot(x_domain, hist(x_domain), label="True Histogram")
inverse_hist = gaussian_kde(inverse_evaluated, 0.001)
plt.plot(x_domain, inverse_hist(x_domain), label="1 / True Histogram")
blurred_hist = gaussian_kde(evaluated, 0.1)
plt.plot(x_domain, blurred_hist(x_domain), label="Smoothed Histogram")
blurred_inverse_hist = gaussian_kde(inverse_evaluated, 0.1)
plt.plot(x_domain, blurred_inverse_hist(x_domain), label="1 / Smoothed Histogram")
plt.title("Histogram (pdf) comparing a {} distribution and the distribution of the inverse of all its values".format(title))
plt.legend()
plt.show()
```
## Example - optimizing for finding the minimum of: f(x) = x^2 - x + 1
Let's now define a simple search space and solve for f(x) = x^2 - x + 1, where x is an hyperparameter.
```
%reset -f
from hyperopt import fmin, tpe, hp
import matplotlib.pyplot as plt
def f(x):
return x**2 - x + 1
plt.plot(range(-5, 5), [f(x) for x in range(-5, 5)])
plt.title("Function to optimize: f(x) = x^2 - x + 1")
plt.show()
space = hp.uniform('x', -5, 5)
best = fmin(
fn=f, # "Loss" function to minimize
space=space, # Hyperparameter space
algo=tpe.suggest, # Tree-structured Parzen Estimator (TPE)
max_evals=1000 # Perform 1000 trials
)
print("Found minimum after 1000 trials:")
print(best)
```
## Example with a dict hyperparameter space
Let's solve for minimizing f(x, y) = x^2 + y^2 using a space using a python dict as structure. Later, this will neable us to nest hyperparameters with choices in a clean way.
```
%reset -f
from hyperopt import fmin, tpe, hp
def f(space):
x = space['x']
y = space['y']
return x**2 + y**2
space = {
'x': hp.uniform('x', -5, 5),
'y': hp.uniform('y', -5, 5)
}
best = fmin(
fn=f,
space=space,
algo=tpe.suggest,
max_evals=1000
)
print("Found minimum after 1000 trials:")
print(best)
```
## With choices, Hyperopt hyperspaces can be represented as nested data structures, too
Yet, we have defined spaces as a single parameter. But that is 1D. Normally, spaces contain many parameters. Let's define a more complex one and with one nested hyperparameter choice for an uniform float:
```
%reset -f
from hyperopt import pyll, hp
import pprint
pp = pprint.PrettyPrinter(indent=4, width=100)
# Define a complete space:
space = {
'x': hp.normal('x', 0, 2),
'y': hp.uniform('y', 0, 1),
'use_float_param_or_not': hp.choice('use_float_param_or_not', [
None, hp.uniform('float', 0, 1),
]),
'my_abc_other_params_list': [
hp.normal('a', 0, 2), hp.uniform('b', 0, 3), hp.choice('c', [False, True]),
],
'yet_another_dict_recursive': {
'u': hp.uniform('u', 0, 3),
'v': hp.uniform('v', 0, 3),
'u': hp.uniform('w', -3, 0)
}
}
# Print a few random (stochastic) samples from the space:
for _ in range(10):
pp.pprint(pyll.stochastic.sample(space))
```
## Let's now record the history of every trial
This will require us to import a few more things, and return the results with a dict that has a "status" and "loss" key at least. Let's keep in our return dict the evaluated space too as this may come in handy if we save results to disk.
```
%reset -f
from hyperopt import fmin, tpe, hp, Trials, STATUS_OK, STATUS_FAIL
import pprint
pp = pprint.PrettyPrinter(indent=4)
def f(space):
x = space['x']
y = space['y']
if y > 1:
# Make use of status fail as an example of skipping on error
result = {
"loss": -1,
"status": STATUS_FAIL,
"space": space
}
return result
loss = x**2 + y**2
result = {
"loss": loss,
"status": STATUS_OK,
"space": space
}
return result
space = {
'x': hp.uniform('x', -5, 5),
'y': hp.uniform('y', -5, 5)
}
trials = Trials()
best = fmin(
fn=f,
space=space,
algo=tpe.suggest,
trials=trials,
max_evals=1000
)
print("Found minimum after 1000 trials:")
print(best)
print("")
print("Here are the space and results of the 3 first trials (out of a total of 1000):")
pp.pprint(trials.trials[0])
pp.pprint(trials.trials[1])
pp.pprint(trials.trials[2])
# pp.pprint(trials.trials[...])
# pp.pprint(trials.trials[999])
print("")
print("What interests us most is the 'result' key of each trial (here, we show 7):")
pp.pprint(trials.trials[0]["result"])
pp.pprint(trials.trials[1]["result"])
pp.pprint(trials.trials[2]["result"])
pp.pprint(trials.trials[3]["result"])
pp.pprint(trials.trials[4]["result"])
pp.pprint(trials.trials[5]["result"])
pp.pprint(trials.trials[6]["result"])
# pp.pprint(trials.trials[...]["result"])
# pp.pprint(trials.trials[999]["result"])
```
## Up next: saving results to disk while optimizing for resuming a stopped hyperparameter search
Note that the optimization could be parallelized by using MongoDB and storing the trials' state here. Althought this is a built-in feature of hyperopt, let's keep things simple for our examples here.
Indeed, the TPE algorithm used by the fmin function has state which is stored in the trials and which is useful to narrow the search space dynamically once we have a few trials. It is then interesting to pause and resume a training, and to apply that to a real problem.
This is what's done inside the hyperopt_optimize.py file of the [GitHub repository for this project](https://github.com/Vooban/Hyperopt-Keras-CNN-CIFAR-100). Here, as an example, we optimize a convolutional neural network for solving the CIFAR-100 problem.
| github_jupyter |
\* *[Notice] I wrote thie code while following the examples in [Choi's Tesorflow-101 tutorial](https://github.com/sjchoi86/Tensorflow-101). And, as I know, most of Choi's examples originally come from [Aymeric Damien's](https://github.com/aymericdamien/TensorFlow-Examples/) and [Nathan Lintz's ](https://github.com/nlintz/TensorFlow-Tutorials) tutorials.*
## 1. Linear Regression
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
#%matplotlib inline
```
### Set initial data
My training data : $y = 0.5x + 0.1 + \sigma(0,0.1)$
```
W_ref = 0.5
b_ref = 0.1
nData = 51
noise_mu = 0
noise_std = 0.1
X_train = np.linspace(-2,2,nData)
Y_test = W_ref * X_train + b_ref
Y_train = Y_test + np.random.normal(noise_mu, noise_std, nData)
```
### Plot the data using *matplotlib*
```
plt.figure(1)
plt.plot(X_train, Y_test, 'ro', label='True data')
plt.plot(X_train, Y_train, 'bo', label='Training data')
plt.axis('equal')
plt.legend(loc='lower right')
plt.show()
```
### Write a TF graph
```
X = tf.placeholder(tf.float32, name="input")
Y= tf.placeholder(tf.float32, name="output")
W = tf.Variable(np.random.randn(), name="weight")
b = tf.Variable(np.random.randn(), name="bias")
Y_pred = tf.add(tf.mul(X, W), b)
```
We use a L2 loss function, $loss = -\Sigma (y'-y)^2$
*reduce_mean(X)* returns the mean value for all elements of the tensor *X*
```
loss = tf.reduce_mean(tf.square(Y-Y_pred))
learning_rate = 0.005
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss)
training_epochs = 50 # We will repeat the learning process 2000 times
display_epoch = 5 # We will print the error at every 200 epochs
```
### Run the session
```
sess = tf.Session()
sess.run(tf.initialize_all_variables())
for epoch in range(training_epochs):
for (x,y) in zip(X_train, Y_train):
sess.run(optimizer, feed_dict={X:x, Y:y})
# Print the result
if (epoch+1) % display_epoch == 0:
W_temp = sess.run(W)
b_temp = sess.run(b)
loss_temp = sess.run(loss, feed_dict={X: X_train, Y:Y_train})
print "(epoch {})".format(epoch+1)
print "[W, b / loss] {:05.4f}, {:05.4f} / {:05.4f}".format(W_temp, b_temp, loss_temp)
print " "
# Final results
W_result = sess.run(W)
b_result = sess.run(b)
print "[Final: W, b] {:05.4f}, {:05.4f}".format(W_result, b_result)
print "[Final: W, b] {:05.4f}, {:05.4f}".format(W_ref, b_ref)
plt.figure(2)
plt.plot(X_train, Y_test, 'ro', label='True data')
plt.plot(X_train, Y_train, 'bo', label='Training data')
plt.plot(X_train, W_result*X_train+b_result, 'g-', linewidth=3, label='Regression result')
plt.axis('equal')
plt.legend(loc='lower right')
plt.show()
sess.close()
```
| github_jupyter |
## WEB-SCRAPYING
Web scraping is a computer software technique of extracting information from websites.
This technique mostly focuses on the transformation of unstructured data (HTML format) on the web into structured data (database or spreadsheet).
### PROBLEM STATEMENT
### Crawl popular websites & create a database of Indian movie celebrities with their images and details.
**Libraries used:**
Requests library :https://requests.readthedocs.io/en/master/
- It is a Python module which can be used for fetching URLs
BeautifulSoup library:https://www.crummy.com/software/BeautifulSoup/bs4/doc/
- Beautiful Soup is a Python library for pulling data out of HTML and XML files. It works with your favorite parser to provide idiomatic ways of navigating, searching, and modifying the parse tree.
### IMPORT LIBRARIES
```
""" lxml, re,numpy as np,pandas as pd,os, cv2,from bs4 import BeautifulSoup,from requests import get,
import matplotlib.pyplot as plt
%matplotlib inline"""
os.getcwd()
```
### Make the list of Urls
```
url1 = [""" list of urls"""]
```
### Create a class to extract the details from one website
```
class IMDB(object):
"""docstring for IMDB"""
def __init__(self, url):
super(IMDB, self).__init__()
"""fetch the webpage using the url provided using the requests library"""
"""import the Beautiful soup functions to parse the data returned from the website using lxml parser"""
def articleTitle(self):
"""return the title of the article by searching for class = **header** with h1 tag """
def bodyContent(self):
"""return the body of the article by searching for class **lister-item mode-detail** With div tag """
def IndiancelebData(self):
"""fetch the body content of the page using self.bodyContent function"""
"""initialise the list of features for particular page"""
"""create the celeb_img folder to store folder if required
and change the directory to that particular folder to store image"""
"""searh for the box containing details of each celebrity and append the name,image Url,detail and image"""
CelebData = ["""list of celebrity """]
os.chdir("../")
return CelebData
if __name__ == '__main__':
"""initialise a list for each feature and one new list to store the features"""
for i in range(len(url1)):
"""create an instance for each url in the url list and print the title of the page,
add the information extracted from the particular page onto the list initialised
"""
```
### Create the dataframe
```
"""store the data (CelebName,CelebImageurl,description ,CelebImage) in the dataframe using pandas"""
```
### Remove the duplicates
```
"""print remove the duplicates based on name or even all the data"""
```
### Display the dataset
```
"""print actor's name
actors image url
about actor
image of the actor
"""
```
### Store the dataframe in csv file
```
"""store the dataframe in csv using pandas"""
```
| github_jupyter |
# Preliminaries
Write requirements to file, anytime you run it, in case you have to go back and recover dependencies.
Requirements are hosted for each notebook in the companion github repo, and can be pulled down and installed here if needed. Companion github repo is located at https://github.com/azunre/transfer-learning-for-nlp
```
!ls ../input/jw300entw/jw300.en-tw.en
!pip freeze > kaggle_image_requirements.txt
```
# Fine-tune DistilmBERT on Monolongual Twi Data (multilingual mBERT Tokenizer)
Initialize DistilmBERT tokenizer to DistilmBERT checkpoint
```
from transformers import DistilBertTokenizerFast # this is just a faster version of DistilBertTokenizer, which you could use instead
tokenizer = DistilBertTokenizerFast.from_pretrained("distilbert-base-multilingual-cased") # use pre-trained DistilmBERT tokenizer
```
Having prepared tokenizer, load mBERT checkpoint into a BERT masked language model.
```
from transformers import DistilBertForMaskedLM # use masked language modeling
model = DistilBertForMaskedLM.from_pretrained("distilbert-base-multilingual-cased") # initialize to DistilmBERT checkpoint
print("Number of parameters in DistilmBERT model:")
print(model.num_parameters())
```
Build monolingual Twi dataset with tokenizer using method included with transformers
```
from transformers import LineByLineTextDataset
dataset = LineByLineTextDataset(
tokenizer=tokenizer,
file_path="../input/jw300entw/jw300.en-tw.tw",
block_size=128, # how many lines to read at a time
)
```
We will also need a "data collator". This is a helper method that creates a special object out of a batch of sample data lines (of length block_size). This special object is consummable by PyTorch to neural network training
```
from transformers import DataCollatorForLanguageModeling
data_collator = DataCollatorForLanguageModeling(
tokenizer=tokenizer,
mlm=True, mlm_probability=0.15) # use masked language modeling, and mask words with probability of 0.15
```
Define standard training arguments, and then use them with previously defined dataset and collator to define a "trainer" for one epoch, i.e. across all 600000+ examples.
```
from transformers import Trainer, TrainingArguments
training_args = TrainingArguments(
output_dir="twidistilmbert",
overwrite_output_dir=True,
num_train_epochs=1,
per_gpu_train_batch_size=16,
save_total_limit=1,
)
trainer = Trainer(
model=model,
args=training_args,
data_collator=data_collator,
train_dataset=dataset,
prediction_loss_only=True,
)
```
Train and time.
```
import time
start = time.time()
trainer.train()
end = time.time()
print("Number of seconds for training:")
print((end-start))
trainer.save_model("twidistilmbert") # save model
```
Test model on "fill-in-the-blank" task, by taking a sentence, masking a word and then predicting a completion with pipelines API.
```
# Define fill-in-the-blanks pipeline
from transformers import pipeline
fill_mask = pipeline(
"fill-mask",
model="twidistilmbert",
tokenizer=tokenizer
)
# We modified a sentences as "Eyi de ɔhaw kɛse baa sukuu hɔ." => "Eyi de ɔhaw kɛse baa [MASK] hɔ."
# Predict masked token
print(fill_mask("Eyi de ɔhaw kɛse baa [MASK] hɔ."))
```
| github_jupyter |
# Examples of the options
This is a notebook example to show the abilities of `tutormagic`. You can install the extension using `pip` or `conda`.
```python
pip install tutormagic
```
or
```python
conda install tutormagic
```
```
%load_ext tutormagic
```
# Option *language*: `-l` or `--lang`
Choose the language you want to use for [pythontutor.com](http://pythontutor.com/). Available options are:
* `python2`
* `python3`
* `py3anaconda` (experimental and unsupported)
* `java`
* `javascript`
* `typescript`
* `ruby`
* `c`
* `c++`
If this option is not used the default value is `python3`.
Example:
```
%%tutor -l python3
a = 1
b = 2
print(a + b)
```
# Option *height*: `-h` or `--height`
Change the height of the output area display in pixels.
Example:
```
%%tutor -l python3 -h 100
a = 1
b = 2
print(a + b)
```
# Option *tab*: `-t` or `--tab`
Open pythontutor in a new tab. *height* option will be ignored if *tab* option is used.
Example:
```
%%tutor -l python3 -t
a = 1
b = 2
print(a + b)
```
# Option *secure*: `-s` or `--secure`
Open pythontutor using https in a new tab. *height* and *tab* options will be ignored if *secure* option is used.
Example:
```
%%tutor -l python3 -s
a = 1
b = 2
print(a + b)
```
# Option *link*: `-k` or `--link`
Just display a link to pythontutor with your defined code.
Example:
```
%%tutor -k
a = 1
b = 1
print(a + b)
```
# Option *run*: `-r` or `--run`
Use this option if you also want to run the code in the cell in the notebook.
Example:
```
%%tutor -r
a = 1
b = 1
print("This is run in the notebook as well as in PythonTutor: ", a + b)
print("So, you have access to 'a' and 'b' vars...")
print("the value of a from the previous cell is: ", a)
```
# Option *cumulative*: `--cumulative`
[PythonTutor config](https://github.com/pgbovine/OnlinePythonTutor/blob/master/v3/docs/embedding-HOWTO.md#iframe-embedding-parameters): Set the *cumulative* option to `True`.
Example:
```
%%tutor --cumulative
def func():
return 10
a = func()
print(a)
```
# Option *heapPrimitives*: `--heapPrimitives`
[Render objects on the heap](https://github.com/pgbovine/OnlinePythonTutor/blob/bf157918de5c28cebdc1eb74b77101f639b4e6b5/v3/docs/user-FAQ.md#i-thought-all-objects-in-python-are-conceptually-on-the-heap-why-does-python-tutor-render-primitive-values-eg-numbers-strings-inside-of-stack-frames).
Example:
```
%%tutor --heapPrimitives
a = 1
b = 1
print(a + b)
```
# Option *textReferences*: `--textReferences`
[Use text labels for references](https://github.com/pgbovine/OnlinePythonTutor/blob/bf157918de5c28cebdc1eb74b77101f639b4e6b5/v3/docs/user-FAQ.md#i-thought-all-objects-in-python-are-conceptually-on-the-heap-why-does-python-tutor-render-primitive-values-eg-numbers-strings-inside-of-stack-frames).
Example:
```
%%tutor --textReferences --heapPrimitives
a = 1
b = 1
print(a + b)
```
# Option *curInstr*: `--curInstr`
[PythonTutor config: Start at the defined step](https://github.com/pgbovine/OnlinePythonTutor/blob/master/v3/docs/embedding-HOWTO.md#iframe-embedding-parameters).
Example:
```
%%tutor --curInstr 2
a = 1
b = 1
print(a + b)
```
# Option *verticalStack*: `--verticalStack`
[PythonTutor config: Set visualization to stack atop one another](https://github.com/pgbovine/OnlinePythonTutor/blob/master/v3/docs/embedding-HOWTO.md#iframe-embedding-parameters).
Example:
```
%%tutor --verticalStack
a = 1
b = 1
print(a + b)
```
# Option lang *py3anaconda*: `--lang py3anaconda`
This option allows you to import more modules like `numpy`. It is experimental and unsupported (check pythontutor web page for more information).
Example:
```
%%tutor --lang py3anaconda
import numpy as np
arr = np.arange(10)
print(arr * 10)
```
| github_jupyter |
# MEGI001-2101033 Introduction to Earth System Data
## Task 5.3 - Data Handling Analysis
Created on: Jan 24, 2019 by Ralph Florent <r.florent@jacobs-university.de>
## T 5.3 : Spectral analysis
Please perform the operations above (in a new notebook, properly renamed) for the labotory spectrum of Montmorillonite, i.e. ```montmorillonite-1292F35-RELAB.txt```.
Please document any isssue/trouble
```
# -*- coding: utf-8 -*-
"""
Created on Jan 24 2019
@author: Angelo Rossi, Ralph Florent
"""
# Import relevant libraries
import matplotlib
from matplotlib import pyplot as plt
%matplotlib inline
import numpy as np
import pandas as pd # we'll use pandas to load data
from scipy.interpolate import interp1d
# We'll be looking at phylosiclicates (clays), as they have a fairly complex spectra with many features:
# In the data_samples folder we have two spectra for Montmorillonite
PATH = '../assets/data/spectral_data/'
specSlPhyMontm = pd.read_csv(PATH + 'FRTC596_537-220-5x5_Al-phyllo_montm.txt', skipinitialspace=True, sep=' ',
skiprows=3, header=None)
relabMontm = pd.read_csv(PATH + 'montmorillonite-1292F35-RELAB.txt', skipinitialspace=True, sep=' ',
skiprows=3, header=None)
# also we have two spectra for Nontronite
specFePhyNontr = pd.read_csv(PATH + 'FRTC596_537-220-5x5_Fe-phyllo_nontr.txt', skipinitialspace=True, sep=' ',
skiprows=3, header=None)
relabNontr = pd.read_csv(PATH + 'nontronite-CBJB26-RELAB.txt', skipinitialspace=True, sep=' ',
skiprows=3, header=None)
specFePhyNontr[:5] # take a look at what our data looks like
# rename columns using dictionary
specFePhyNontr.rename(columns={0:'wavelength',1:'specFePhyNontr'})[:5]
# This worked, lets fix all dataframes now:
specSlPhyMontm = specSlPhyMontm.rename(columns={0:'wavelength',1:'specSlPhyMontm'})
relabMontm = relabMontm.rename(columns={0:'wavelength',1:'relabMontm'})
specFePhyNontr = specFePhyNontr.rename(columns={0:'wavelength',1:'specFePhyNontr'})
relabNontr = relabNontr.rename(columns={0:'wavelength',1:'relabNontr'})
#lets check:
relabNontr[:5]
#lets plot:
ax1 = specSlPhyMontm.plot.line(0, figsize=(15,5), title="Montmorillonite")
relabMontm.plot.line(0, ax=ax1) # ax=ax plots all on the same axis
ax2 = specFePhyNontr.plot.line(0, figsize=(15,5), title="Nontronite")
relabNontr.plot.line(0, ax=ax2)
ax3 = relabMontm.plot.line(0, figsize=(15,5), title="Montmorillonite vs Nontronite")
relabNontr.plot.line(0, ax=ax3)
plt.show()
```
## Continuum removal
When processing reflective spectrometry, we normally want to look at absorption features.
Remember that this is a plot of reflectivity against wavelength, the higher absorption means lower reflection,
so what we want to find are location of "troughs", ie local minima.
The first step we could do towards solving this non-trivial problem would be to remove continuum, or the shape of the spectra "at large" then the troughs would become more aparent and easier to identify.
For this example we shall use the *relabNontr* spectra, as it is the smoothest and would be easiest to show the process in an ideal case.
In this part of the tutorial we will show how not only how to apply code, but also how to write it, thus demonstrating the entire workflow of ad-hoc data analysis.
We begin by creating simple fake datasets to test our code while we're writing it, gradually increasing complexity
while develping code, and lastly applying it to real data.
```
#lets put this into a function so we can reuse it later:
def removeCont(pSample):
pSampleLineX=[pSample[0][0],pSample[0][-1]]
pSampleLineY=[pSample[1][0],pSample[1][-1]]
pSampleLine=[pSampleLineX,pSampleLineY]
finterp = interp1d(pSampleLine[0],pSampleLine[1])#create interploation function
return pSample[1]-finterp(pSample[0])
# Lets try this on a bigger fake dataset:
pivot = lambda sample: [[a[0] for a in sample],[a[1] for a in sample]]
sample = np.asarray([list(x) for x in zip(range(32),[x[0] for x in np.random.random((32,1)).tolist()])])
pSample=pivot(sample)
plt.plot(pSample[0],pSample[1], '-',pSample[0],removeCont(pSample),'-')
plt.legend(['original', 'continuum removed'], loc='best')
plt.show()
#we can now easily find a maximum value above the continuum for points other than first and last:
max(removeCont(pSample)[1:-1])
#furthermore, we can ask numpy for an index of max:
maxIndex = np.argmax(removeCont(pSample)[1:-1]) +1
maxIndex
#Armed with this information we can repeat the previous step for a subset:
plt.plot(pSample[0][:maxIndex],pSample[1][:maxIndex], '-',pSample[0][:maxIndex],
removeCont([pSample[0][:maxIndex],pSample[1][:maxIndex]]),'-')
plt.legend(['original', 'continuum removed'], loc='best')
plt.show()
# From henceforth we could do this recursively - that is to use the output of a function as an input to itself:
def getMaxima(pSample):
def getMaximaInner(innerSample):
contRem=removeCont(innerSample)
#print(contRem)
maxIndex=np.argmax(contRem)
#print(maxIndex)
maxVal=contRem[maxIndex]
maxLoc=innerSample[0][maxIndex]
if len(contRem)>2 and maxVal>contRem[0] and maxVal>contRem[-1]: # check that the maximum is more than edges
maxLocArray.append(maxLoc)
#print(maxLoc)
subsetLeft=[innerSample[0][:maxIndex+1],innerSample[1][:maxIndex+1]]
#print(subsetLeft[0])
subsetRight=[innerSample[0][maxIndex:],innerSample[1][maxIndex:]]
#print(subsetRight[0])
getMaximaInner(subsetLeft)
getMaximaInner(subsetRight)
maxLocArray=[] #initialize array to store a list of points on a convex hull
getMaximaInner(pSample)
maxLocArray.sort()
return [pSample[0][0]]+maxLocArray+[pSample[0][-1]]
#maxList=getMaxima([pSample[0][:5],pSample[1][:5]])
maxList=getMaxima(pSample)
print(maxList)
hull=[maxList,[x[1] for x in sample if x[0] in maxList]]
plt.plot(pSample[0],pSample[1], '-',hull[0],hull[1],'-')
plt.show()
#Now lets try it with a real dataset:
sample = np.asarray(relabMontm)
pSample = pivot(sample)
maxList = getMaxima(pSample)
print(maxList)
hull = [maxList,[x[1] for x in sample if x[0] in maxList]]
plt.plot(pSample[0],pSample[1], '-',hull[0],hull[1],'-')
# You may notice that the formation of a convex hull is distorted
# by a long row of zeros at the end of the data sample
# Lets remove all zeros:
cleanSample=[value for value in sample if value[1]>0]
pSample=pivot(cleanSample)
maxList=getMaxima(pSample)
#print(maxList)
pHull=[maxList,[x[1] for x in sample if x[0] in maxList]]
plt.plot(pSample[0],pSample[1], '-',pHull[0],pHull[1],'-')
plt.legend(['spectrum', 'convex hull'])
plt.show()
# Next we can subtact the convex hull from our data, in a manner similar to how we subtracted continuum ealier
def removeHull(pSample,pHull):
finterp = interp1d(pHull[0],pHull[1])#create interploation function
return pSample[1]-finterp(pSample[0])
hullRemoved = removeHull(pSample,pHull)
plt.plot(pSample[0],pSample[1],'-',pSample[0],hullRemoved, '-')
plt.legend(['spectrum', 'convex hull removed'])
plt.show()
# we can easily find indices for these values:
splitInd=[pSample[0].index(x) for x in pHull[0]]
print(splitInd)
#then we can split the array along the indices using list comprehension:
splitSample=[[pSample[0][splitInd[i]:splitInd[i+1]],hullRemoved[splitInd[i]:splitInd[i+1]]]
for i in range(len(splitInd)-1) if splitInd[i+1]-splitInd[i]>2]
for s in splitSample:
plt.plot(s[0],s[1],'-')
plt.show()
# Finding local minima is then straightforward:
listMinimaX=[x[0][np.argmin(x[1])] for x in np.asarray(splitSample)]
print(listMinimaX)
# we can use list comprehension again to get corresponding Y-values
listMinimaY=[pSample[1][pSample[0].index(x)] for x in listMinimaX]
print(listMinimaY)
#And we can combine the two and plot the minima on a graph:
splitSample=[[pSample[0][splitInd[i]:splitInd[i+1]],pSample[1][splitInd[i]:splitInd[i+1]]]
for i in range(len(splitInd)-1) if splitInd[i+1]-splitInd[i]>2]
for s in splitSample:
plt.plot(s[0],s[1],'-')
plt.plot(listMinimaX,listMinimaY,'x',color='black')
plt.show()
# We've now identified some deep absorption bands, a some shallow that are probably noise.
# We can filter out the shallow ones by appling a threshold:
# First get the band depths with hull removed:
listMinimaYhullRemoved=[hullRemoved[pSample[0].index(x)] for x in listMinimaX]
print(listMinimaYhullRemoved)
# Now apply a threshold:
threshold=0.05
listMinimaSigX=[q[0] for q in list(zip(listMinimaX,listMinimaYhullRemoved)) if q[1]<-threshold]
listMinimaSigYhullRemoved=[q[1] for q in list(zip(listMinimaX,listMinimaYhullRemoved)) if q[1]<-threshold]
listMinimaSigY=[pSample[1][pSample[0].index(x)] for x in listMinimaSigX]
#then we can split the array along the indices using list comprehension:
splitSample=[[pSample[0][splitInd[i]:splitInd[i+1]],hullRemoved[splitInd[i]:splitInd[i+1]]]
for i in range(len(splitInd)-1) if splitInd[i+1]-splitInd[i]>2]
plt.figure(figsize=(15,5)) #make larger figure
for s in splitSample:
plt.plot(s[0],s[1],'-')
for xc in listMinimaSigX:
plt.axvline(x=xc,color='black')
plt.show()
# Finally, lets see where do these band depths plot on original spectra:
plt.figure(figsize=(15,5))
splitSample=[[pSample[0][splitInd[i]:splitInd[i+1]],pSample[1][splitInd[i]:splitInd[i+1]]]
for i in range(len(splitInd)-1) if splitInd[i+1]-splitInd[i]>2]
for s in splitSample:
plt.plot(s[0],s[1],'-')
plt.plot(listMinimaSigX,listMinimaSigY,'o',color='black')
plt.show()
print("adsorption band center wavelenghts are:")
for item in listMinimaSigX:
print(item, "micrometers")
```
| github_jupyter |
```
import json
import time
fd=open("records.json","r")
txt=fd.read()
fd.close()
products=json.loads(txt)
fd=open("sales.json","r")
txt=fd.read()
fd.close()
sales=json.loads(txt)
```
# Purchasing The Products
```
print("*******************MENU*******************")
print("product_id product_name",end="\n\n")
for i in products.keys():
print(i," ",products[i]["Name"])
```
## Getting Product Id and quantity
```
user_id=input("Enter the product id :")
quantity=int(input("Enter The Quantity :"))
product_id=products.keys()
if (user_id not in product_id):
print("Entered wrong Product Id ")
flag=0
if(products[user_id]["Quantity"]<quantity):
print("The available quantity is :" ,products[user_id]["Quantity"])
ans=int(input("Do you want to take these quantity(1-yes/0-No)"))
if(ans):
print("The bill is processed!!")
quantity=products[user_id]["Quantity"]
else:
flag=1
print("choose any other product")
products[user_id]["Quantity"]-=quantity
```
## Printing The Bill
```
if(flag==0):
print("*************Bill***********",end=" ")
print("Time :",time.ctime(),"**********************")
print("Product :",products[user_id]["Name"])
print("Quanity :",quantity)
print("Price :",products[user_id]["Price"])
print("Billing Amount :",products[user_id]["Price"]*quantity)
print("***********************************************************************************")
```
## Making Transaction Details
```
trans_id=len(sales)+1
trans_details=products[user_id].copy()
trans_details['time']=time.ctime()
trans_details['Quantity']=quantity
sales[trans_id]=trans_details
```
# Adding the products to the Inventory
```
print("To Add Product")
product_id=products.keys()
new_id=input("Enter the id of the product")
if new_id in product_id:
qun=int(input("Enter the product Quantity"))
products[new_id]["Quantity"]+=qun
print("Qunatity Added")
else:
nname=input("Enter the name of the product")
nprice=int(input("enter the price"))
nquantity=int(input("enter the quantity"))
nbrand=input("Enter the brand of the product")
medate=input("Enter the expiry date of the product")
new_product["Name"]=nname
new_product["Price"]=nprice
new_product["Quantity"]=nquantity
new_product["ExpiryDate"]=medate
new_product["Brand"]=nbrand
products[new_id]=new_product
fd=open("records.json","w")
jformat=json.dumps(products)
fd.write(jformat)
fd.close()
fd=open("sales.json","w")
jformat=json.dumps(sales)
fd.write(jformat)
fd.close()
```
| github_jupyter |
<a href="https://colab.research.google.com/github/Nirzu97/pyprobml/blob/logreg_jax/notebooks/logreg_jax.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Logistic regression <a class="anchor" id="logreg"></a>
In this notebook, we illustrate how to perform logistic regression on some small datasets. We will compare binary logistic regression as implemented by sklearn with our own implementation, for which we use a batch optimizer from scipy. We code the gradients by hand. We also show how to use the JAX autodiff package (see [JAX AD colab](https://github.com/probml/pyprobml/tree/master/book1/supplements/autodiff_jax.ipynb)).
```
# Standard Python libraries
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import time
import numpy as np
import glob
import matplotlib.pyplot as plt
import PIL
import imageio
from IPython import display
import sklearn
import seaborn as sns;
sns.set(style="ticks", color_codes=True)
# https://github.com/google/jax
import jax
import jax.numpy as jnp
from jax.scipy.special import logsumexp
from jax import grad, hessian, jacfwd, jacrev, jit, vmap
from jax.experimental import optimizers
print("jax version {}".format(jax.__version__))
# First we create a dataset.
import sklearn.datasets
from sklearn.model_selection import train_test_split
iris = sklearn.datasets.load_iris()
X = iris["data"]
y = (iris["target"] == 2).astype(np.int) # 1 if Iris-Virginica, else 0'
N, D = X.shape # 150, 4
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.33, random_state=42)
# Now let's find the MLE using sklearn. We will use this as the "gold standard"
from sklearn.linear_model import LogisticRegression
# We set C to a large number to turn off regularization.
# We don't fit the bias term to simplify the comparison below.
log_reg = LogisticRegression(solver="lbfgs", C=1e5, fit_intercept=False)
log_reg.fit(X_train, y_train)
w_mle_sklearn = jnp.ravel(log_reg.coef_)
print(w_mle_sklearn)
# First we define the model, and check it gives the same output as sklearn.
def sigmoid(x): return 0.5 * (jnp.tanh(x / 2.) + 1)
def predict_logit(weights, inputs):
return jnp.dot(inputs, weights) # Already vectorized
def predict_prob(weights, inputs):
return sigmoid(predict_logit(weights, inputs))
ptest_sklearn = log_reg.predict_proba(X_test)[:,1]
print(jnp.round(ptest_sklearn, 3))
ptest_us = predict_prob(w_mle_sklearn, X_test)
print(jnp.round(ptest_us, 3))
assert jnp.allclose(ptest_sklearn, ptest_us, atol=1e-2)
# Next we define the objective and check it gives the same output as sklearn.
from sklearn.metrics import log_loss
from jax.scipy.special import logsumexp
#from scipy.misc import logsumexp
def NLL_unstable(weights, batch):
inputs, targets = batch
p1 = predict_prob(weights, inputs)
logprobs = jnp.log(p1) * targets + jnp.log(1 - p1) * (1 - targets)
N = inputs.shape[0]
return -jnp.sum(logprobs)/N
def NLL(weights, batch):
# Use log-sum-exp trick
inputs, targets = batch
# p1 = 1/(1+exp(-logit)), p0 = 1/(1+exp(+logit))
logits = predict_logit(weights, inputs).reshape((-1,1))
N = logits.shape[0]
logits_plus = jnp.hstack([jnp.zeros((N,1)), logits]) # e^0=1
logits_minus = jnp.hstack([jnp.zeros((N,1)), -logits])
logp1 = -logsumexp(logits_minus, axis=1)
logp0 = -logsumexp(logits_plus, axis=1)
logprobs = logp1 * targets + logp0 * (1-targets)
return -jnp.sum(logprobs)/N
# We can use a small amount of L2 regularization, for numerical stability
def PNLL(weights, batch, l2_penalty=1e-5):
nll = NLL(weights, batch)
l2_norm = jnp.sum(jnp.power(weights, 2)) # squared L2 norm
return nll + l2_penalty*l2_norm
# We evaluate the training loss at the MLE, where the parameter values are "extreme".
nll_train = log_loss(y_train, predict_prob(w_mle_sklearn, X_train))
nll_train2 = NLL(w_mle_sklearn, (X_train, y_train))
nll_train3 = NLL_unstable(w_mle_sklearn, (X_train, y_train))
print(nll_train)
print(nll_train2)
print(nll_train3)
# Next we check the gradients compared to the manual formulas.
# For simplicity, we initially just do this for a single random example.
np.random.seed(42)
D = 5
w = np.random.randn(D)
x = np.random.randn(D)
y = 0
#d/da sigmoid(a) = s(a) * (1-s(a))
deriv_sigmoid = lambda a: sigmoid(a) * (1-sigmoid(a))
deriv_sigmoid_jax = grad(sigmoid)
a = 1.5 # a random logit
assert jnp.isclose(deriv_sigmoid(a), deriv_sigmoid_jax(a))
# mu(w)=sigmoid(w'x), d/dw mu(w) = mu * (1-mu) .* x
def mu(w): return sigmoid(jnp.dot(w,x))
def deriv_mu(w): return mu(w) * (1-mu(w)) * x
deriv_mu_jax = grad(mu)
assert jnp.allclose(deriv_mu(w), deriv_mu_jax(w))
# NLL(w) = -[y*log(mu) + (1-y)*log(1-mu)]
# d/dw NLL(w) = (mu-y)*x
def nll(w): return -(y*jnp.log(mu(w)) + (1-y)*jnp.log(1-mu(w)))
def deriv_nll(w): return (mu(w)-y)*x
deriv_nll_jax = grad(nll)
assert jnp.allclose(deriv_nll(w), deriv_nll_jax(w))
# Now let's check the gradients on the batch version of our data.
N = X_train.shape[0]
mu = predict_prob(w_mle_sklearn, X_train)
g1 = grad(NLL)(w_mle_sklearn, (X_train, y_train))
g2 = jnp.sum(jnp.dot(jnp.diag(mu - y_train), X_train), axis=0)/N
print(g1)
print(g2)
assert jnp.allclose(g1, g2, atol=1e-2)
H1 = hessian(NLL)(w_mle_sklearn, (X_train, y_train))
S = jnp.diag(mu * (1-mu))
H2 = jnp.dot(jnp.dot(X_train.T, S), X_train)/N
print(H1)
print(H2)
assert jnp.allclose(H1, H2, atol=1e-2)
# Finally, use BFGS batch optimizer to compute MLE, and compare to sklearn
import scipy.optimize
def training_loss(w):
return NLL(w, (X_train, y_train))
def training_grad(w):
return grad(training_loss)(w)
np.random.seed(43)
N, D = X_train.shape
w_init = np.random.randn(D)
w_mle_scipy = scipy.optimize.minimize(training_loss, w_init, jac=training_grad, method='BFGS').x
print("parameters from sklearn {}".format(w_mle_sklearn))
print("parameters from scipy-bfgs {}".format(w_mle_scipy))
assert jnp.allclose(w_mle_sklearn, w_mle_scipy, atol=1e-1)
prob_scipy = predict_prob(w_mle_scipy, X_test)
prob_sklearn = predict_prob(w_mle_sklearn, X_test)
print(jnp.round(prob_scipy, 3))
print(jnp.round(prob_sklearn, 3))
assert jnp.allclose(prob_scipy, prob_sklearn, atol=1e-2)
```
| github_jupyter |
# Matplotlib
Matplotlib is the standard library in Python used for plotting. It can be really powerful and offers a lot of customization, but with that can come frustration and an initial learning curve. Plotting something requires interacting different levels. Often you can use high level coommands like plot this array but sometimes it is necessary to interact at a low-level or use very specific commands like change the color of this pixel. Matplotlib does a great job of allowing you to interact at a high level most of the time to easily make plots, while leaving the freedom to make changes at a lower level when necessary.
Today, we are going to focus on learning to use the built in functions within matplotlib to make a variety of plot types. Matplotlib has a number of usefule modules, but we will focus on the most commonly used **pyplot**. Pyplot is a collection of functions which allow matplotlib to be run like MATLAB. This is sufficient for most plot types you will need to make.
First, we import the pyplot module. We alias this as **plt** to make calling all of the plotting functions easer (less typing)! **plt** is an extremely common alias for matplotlib and most code you see will use this.
```
%matplotlib notebook
import matplotlib.pyplot as plt
import numpy as np
```
The %matplotlib notebook command just tells Jupyter to automatically display the plots we generate. This is not needed when making plots outside of a Jupyter notebook.
We'll start by making a simple plot and taking a closer look at each part.
```
x = np.linspace(0,10) #array with values from 0 to 10
plt.plot(x, x, label='Line') #Plot a line
plt.plot(x, x**2, label='x Squared') # Plot x^2
plt.title('This is the Title')
plt.xlabel('X')
plt.ylabel('Y')
plt.legend()
```
## Parts of the Figure
**Figure**
The entire plot is called a figure. It is an object that keeps track of all parts of the figure including the axes and any number of special artists (ie: title, legend, labels, etc. The figure also keeps track of the canvas, but for now we will ignore this. When using matplotlib any interaction with the canvas should happen under the hood, making it virtually invisible.
**Axes**
The axes part of a figure contains all of the information for both axis of a 2D plot (ie: axis labels, tick mark spacing and numbering, the title, etc). You can have multiple axis on a figure, as you will see below, but each axes can only be on one figure. The attributes of the axes can be controlled with a set of methods, such as set_xlabel(), set_ylabel(), set_title(), and many more.
**Axis**
The axis is the object containing the information for each axis on your plot. Two or more of these are combined with some extra information to make the axes. The axis can set graph limits, set tickmarks and labels and change the visual properties of an individual axis (for example making thicker tick marks, or moving the tick marks inside the plot). Many of the changes you want to make to an axis can be accomplished using the methods within **axes**, however there is a greater level of control over specific details here, when the need arises.
**Artist**
Basically everything you can see on the figure is an artist (even the Figure, Axes, and Axis objects). Anything you add to a plot is also an artist, for example adding a text box or the legend. Artists are usually tied to an axes object and cannot be shared by multiple axes at once.
## Subplots
The ability to use subplots within pyplot is extremely useful. It allows you to place multiple axes options on a single figure. For example:
```
x = np.linspace(0,10) #array with values from 0 to 10
fig, ax = plt.subplots(2,1, figsize=(10,5))
ax[0].plot(x, x) #Plot a line
ax[1].plot(x, x**2) # Plot x^2
fig.suptitle('This is the Main Title')
ax[0].set_title('X vs X')
ax[1].set_title('X Squared')
ax[1].set_xlabel('X')
ax[0].set_ylabel('Y')
ax[1].set_ylabel('Y')
```
Here, we have two **axes**. In the plt.subplots(2,1) function we specify that we want the figure to have 2 axis vertically and only one horizontally. Changing this to plt.subplots(1,2) would render the plots next to one another.
We also use a slightly different coding style here (with subplots) than in the first example. To set all of the axis labes and titles we use the methods for each **axes**. The top plot is axis 1, accessed using `ax[0]` and the bottom with `ax[1]`.
| github_jupyter |
# Lecture 5 - Halfway Review
*Monday, June 15th 2020*
*Rahul Dani*
In this lecture, we will recap all the material we have learned so far and try to work on some fun tasks!
"Everyone should learn how to code, it teaches you how to think" - Steve Jobs
## Activity 1 : Mad Libs
```
noun_1
noun_2
plural_noun_1
game
plural_noun_2
verb_ending_ING_1
verb_ending_ING_2
plural_noun_3
verb_ending_ING_3
noun_3
plant
body_part
place
verb_ending_ING_4
```
Make variables names with the words I have given above and take user input for those words with "Enter an adjective: " for adjectives, etc. Also come up with creative answers to each of the words. I might ask you to share some funny ones. Please choose appropriate words!
<!-- Sentence : 'A vacation is when you take a trip to some ' + adjective_1 + ' place with your '+adjective_2+ 'family. Usually you go to some place that is near a/an ' +noun_1+ ' or up on a/an '+noun_2+'. A good vacation place is one where you can ride '+plural_noun_1+ ' or play '+game+ ' or go hunting for '+plural_noun_2+'. I like to spend my time '+verb_ending_ING_1+' or '+verb_ending_ING_2+'. When parents go on a vacation, they spend their time eating three '+plural_noun_3+' a day, and fathers play golf, and mothers sit around '+verb_ending_ING_3+'. Last summer, my little brother fell in a/an '+noun_3+' and got poison '+plant+' all over his '+body_part+'. My family is going to go to (the) '+place+', and I will practice '+verb_ending_ING_4+'.' -->
<!-- Answer:
```
print('Welcome to the Mad Libs Game!\n')
adjective_1 = input('Enter an adjective: ')
adjective_2 = input('Enter an adjective: ')
noun_1 = input('Enter a noun: ')
noun_2 = input('Enter a noun: ')
plural_noun_1 = input('Enter a plural noun: ')
game = input('Enter a game: ')
plural_noun_2 = input('Enter a plural noun: ')
verb_ending_ING_1 = input('Enter a verb ending with ING: ')
verb_ending_ING_2 = input('Enter a verb ending with ING: ')
plural_noun_3 = input('Enter a plural noun: ')
verb_ending_ING_3 = input('Enter a verb ending with ING: ')
noun_3 = input('Enter a noun: ')
plant = input('Enter a plant: ')
body_part = input('Enter a body part: ')
place = input('Enter a place: ')
verb_ending_ING_4 = input('Enter a verb ending with ING: ')
print('\nA vacation is when you take a trip to some ' + adjective_1 + ' place with your '+adjective_2+ ' \nfamily. Usually you go to some place that is near a/an ' +noun_1+ ' or up on a/an '+noun_2+'. A good \nvacation place is one where you can ride '+plural_noun_1+ ' or play '+game+ ' or go hunting for '+plural_noun_2+'. I like to spend \nmy time '+verb_ending_ING_1+' or '+verb_ending_ING_2+'. When parents go on a vacation, they spend their time eating \nthree '+plural_noun_3+' a day, and fathers play golf, and mothers sit around '+verb_ending_ING_3+'. Last summer, my \nlittle brother fell in a/an '+noun_3+' and got poison '+plant+' all over his '+body_part+'. My family is going to go \nto (the) '+place+', and I will practice '+verb_ending_ING_4+'.' )
``` -->
```
print('Welcome to the Mad Libs Game!\n')
adjective_1 = input('Enter an adjective: ')
adjective_2 = input('Enter an adjective: ')
noun_1 = input('Enter a noun: ')
noun_2 = input('Enter a noun: ')
plural_noun_1 = input('Enter a plural noun: ')
game = input('Enter a game: ')
plural_noun_2 = input('Enter a plural noun: ')
verb_ending_ING_1 = input('Enter a verb ending with ING: ')
verb_ending_ING_2 = input('Enter a verb ending with ING: ')
plural_noun_3 = input('Enter a plural noun: ')
verb_ending_ING_3 = input('Enter a verb ending with ING: ')
noun_3 = input('Enter a noun: ')
plant = input('Enter a plant: ')
body_part = input('Enter a body part: ')
place = input('Enter a place: ')
verb_ending_ING_4 = input('Enter a verb ending with ING: ')
print('\nA vacation is when you take a trip to some ' + adjective_1 + ' place with your '+adjective_2+ ' \nfamily. Usually you go to some place that is near a/an ' +noun_1+ ' or up on a/an '+noun_2+'. A good \nvacation place is one where you can ride '+plural_noun_1+ ' or play '+game+ ' or go hunting for '+plural_noun_2+'. I like to spend \nmy time '+verb_ending_ING_1+' or '+verb_ending_ING_2+'. When parents go on a vacation, they spend their time eating \nthree '+plural_noun_3+' a day, and fathers play golf, and mothers sit around '+verb_ending_ING_3+'. Last summer, my \nlittle brother fell in a/an '+noun_3+' and got poison '+plant+' all over his '+body_part+'. My family is going to go \nto (the) '+place+', and I will practice '+verb_ending_ING_4+'.' )
```
## Activity 2 : Guessing game
In this exercise you will make a game that lets the user guess a random number between 1 and 100.
For example:
Let the random number be 75.
If the user guesses 60, say "Your guess was too low"
If the user guesses 80, say "Your guess was too high"
Let the user keep entering numbers and only end the program when the user guesses correctly.
Use this to generate a random number between 1 and 100:
import random
number = random.randint(1,100)
<!-- Answer:
```
import random
number = random.randint(1, 100)
guess = -1
attempts = 0
while(guess != number):
guess = int(input('What is your guess? : '))
if (guess < number):
print('Your guess was too low!')
attempts += 1
elif (guess > number):
print('Your guess was too high!')
attempts += 1
else:
attempts += 1
print('Yay you got it!')
print('You took ' + str(score) + ' attempts to get the answer!')
``` -->
```
import random
number = random.randint(1, 100)
guess = -1
attempts = 0
while(guess != number):
guess = int(input('What is your guess? : '))
if (guess < number):
print('Your guess was too low!')
attempts += 1
elif (guess > number):
print('Your guess was too high!')
attempts += 1
else:
attempts += 1
print('Yay you got it!')
print('You took ' + str(attempts) + ' attempts to get the answer!')
```
## Activity 3 : Adding numbers game (Gauss' Formula)
Can you add all the numbers from 1 to 100? Seems like a challenging task, so make a program that can add all the numbers from 1 to 100. Ex: 1+2+3...+98+99+100. Hint: Use a for loop with the range() command.
<!-- Answer:
```
sum = 0
for num in range(101):
sum = sum + num
print(sum)
``` -->
```
sum = 0
for num in range(1,101):
sum = sum + num
print(sum)
```
Now, can you do this task without using a for loop?
<!-- Answer:
```
sum = (100*(101))/2
sum = int(sum)
print(sum)
``` -->
```
sum = int((100*(101))/2)
print(sum)
```
Gauss Formula method:

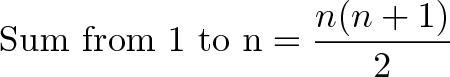
You may be asking: **Why does this matter?**
It matters because we are excuting the same code in less time. We are optimizing our solution!
Let's try adding all the numbers between 1 and 1 million and check which method is faster.
<!-- Solution 1:
```
import time
start_time = time.time()
sum = 0
for num in range(1000001):
sum = sum + num
print(sum)
print("--- %s seconds ---" % (time.time() - start_time))
```
Solution 2:
```
import time
start_time = time.time()
sum = (1000000*(1000001))//2
print(sum)
print("--- %s seconds ---" % (time.time() - start_time))
``` -->
```
import time
start_time = time.time()
sum = 0
for num in range(100000001):
sum = sum + num
print(sum)
print("--- %s seconds ---" % (time.time() - start_time))
import time
start_time = time.time()
sum = (100000000*(100000001))//2
print(sum)
print("--- %s seconds ---" % (time.time() - start_time))
```
## Activity 4: Odd/Even Number Checker
Take a user input number, if the number is even then print 'This number is even', otherwise print 'This number is odd'.
<!-- Answer:
```
num = input('Enter your number here: ')
num = int(num)
if(num % 2 == 0):
print('This number is even!')
else:
print('This number is odd!')
``` -->
```
import random
number = random.randint(1, 100)
print(number)
if (number%2 == 0):
print('even')
else:
print('odd')
```
Practice Problems from previous lectures to work on after class:
Lecture 1 : https://colab.research.google.com/drive/1RlMF5WD6YvUf7sbGs0XkpYyWvOLyT30b?usp=sharing
Lecture 2 : https://colab.research.google.com/drive/14B7NaXdTWmfFhb6wRldo8nTdPMHNX7fZ?usp=sharing
Lecture 3 : https://colab.research.google.com/drive/1WLEpSIk2eDn9YORxIuBJKAIBtvQ0E50Q?usp=sharing
Lecture 4 : https://colab.research.google.com/drive/1eZ3Xzojdu8QYzLD3AfkImMgSf9xUMsHD?usp=sharing
| github_jupyter |
<a href="https://colab.research.google.com/github/aayushkubb/Deep_Learning_Tutorial/blob/master/CIFAR_with_Tensorflow_KERAS_CNN_and_DNN(ALL%20MODELS).ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
### Import the Libraries
```
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import tensorflow as tf
sns.set()
```
### Load the Dataset and Prepare testing and training data
```
from keras.datasets import cifar10
import keras
from keras.models import Sequential
from keras.layers import Dense,Conv2D,Flatten, MaxPool2D
(x_train,y_train),(x_test,y_test)=cifar10.load_data()
print(x_train.shape)
print(x_test.shape)
print('Training data shape : ', x_train.shape, y_train.shape)
print('Testing data shape : ', x_test.shape, y_test.shape)
# Find the unique numbers from the train labels
classes = np.unique(y_train)
nClasses = len(classes)
print('Total number of outputs : ', nClasses)
print('Output classes : ', classes)
plt.figure(figsize=[4,2])
# Display the first image in training data
plt.subplot(121)
plt.imshow(x_train[0,:,:], cmap='gray')
plt.title("Ground Truth : {}".format(y_train[0]))
# Display the first image in testing data
plt.subplot(122)
plt.imshow(x_test[0,:,:], cmap='gray')
plt.title("Ground Truth : {}".format(y_test[0]))
```
## Display images for each category
```
for category in np.unique(y_train):
counter=2
print(category)
for idx, label in enumerate(y_train):
if counter <1:
break
if label==category:
counter-= 1
plt.imshow(x_train[idx])
plt.show()
print("*"*50)
```
# Approach -1 - Using simple DNN or FC
```
# Load the data
(x_train,y_train),(x_test,y_test)=cifar10.load_data()
# train set / data
x_train = x_train.reshape(-1, 32*32*3)
x_train = x_train.astype('float32') / 255
# test set / data
x_test = x_test.reshape(-1, 32*32*3)
x_test = x_test.astype('float32') / 255
# train set / target
y_train = tf.keras.utils.to_categorical(y_train , num_classes=10)
# test set / target
y_test = tf.keras.utils.to_categorical(y_test , num_classes=10)
x_train.shape,x_test.shape
#Models
model = Sequential()
model.add(Dense(800, input_dim=3072, activation="relu"))
model.add(Dense(10, activation="softmax"))
# For using custom metrics
# https://keras.io/api/metrics/#custom-metrics
model.compile(loss="categorical_crossentropy", optimizer="SGD", metrics=["accuracy",tf.keras.metrics.AUC()])
history = model.fit(x_train, y_train, validation_data=(x_test,y_test),
batch_size=200,
epochs=5,
verbose=1)
# https://keras.io/api/models/model_training_apis/
model.summary()
history.history
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['loss'],'r',linewidth=3.0)
plt.plot(history.history['val_loss'],'b',linewidth=3.0)
plt.legend(['Training loss', 'Validation Loss'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('Loss',fontsize=16)
plt.title('Loss Curves',fontsize=16)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['accuracy'],'r',linewidth=3.0)
plt.plot(history.history['val_accuracy'],'b',linewidth=3.0)
plt.legend(['Training accuracy', 'Validation accuracy'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('accuracy',fontsize=16)
plt.title('Accuracy Curves',fontsize=16)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['auc'],'r',linewidth=3.0)
plt.plot(history.history['val_auc'],'b',linewidth=3.0)
plt.legend(['Training AUC', 'Validation AUC'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('AUC',fontsize=16)
plt.title('AUC Curves',fontsize=16)
```
# Approach -2 : Using CNN
## Change the Datatype to Float and Normalize it
```
# Load the data
(x_train,y_train),(x_test,y_test)=cifar10.load_data()
#Normalize
x_train=x_train.astype('float64')/255
x_test=x_test.astype('float64')/255
print(y_train.shape)
print(y_test.shape)
```
## Do the one hot Encoding of the Dependent Variable using Keras.utils
```
from keras.utils import np_utils
from tensorflow.keras.utils import to_categorical
num_classes= len(np.unique(y_train))
y_train= to_categorical(y_train,num_classes)
y_test=to_categorical(y_test,num_classes)
```
### Divide the Training Dataset into validation, Training and Test.
```
(y_train, y_valid) = y_train[5000:], y_train[:5000]
(x_train, x_valid) = x_train[5000:], x_train[:5000]
print(y_train.shape)
print(x_train.shape)
print(y_valid.shape)
print(x_valid.shape)
```
#
## Build a CNN Architecture with KERAS
#### CNN using just the CONV layers
```
model=Sequential()
model.add(Conv2D(filters=9,kernel_size=3,activation='relu',input_shape=(32, 32, 3)))
model.add(Conv2D(filters=6,kernel_size=3,activation='relu'))
model.add(Conv2D(filters=3,kernel_size=3,activation='relu'))
model.summary()
model.add(Flatten())
model.add(Dense(units=256,activation='relu'))
model.add(Dense(units=10,activation='softmax'))
model.summary()
#Compile & Train the model
model.compile(loss="categorical_crossentropy", optimizer="SGD", metrics=["accuracy"])
history = model.fit(x_train, y_train, validation_data=(x_valid,y_valid),
batch_size=200,
epochs=5,
verbose=1)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['accuracy'],'r',linewidth=3.0)
plt.plot(history.history['val_accuracy'],'b',linewidth=3.0)
plt.legend(['Training accuracy', 'Validation accuracy'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('accuracy',fontsize=16)
plt.title('Accuracy Curves',fontsize=16)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['loss'],'r',linewidth=3.0)
plt.plot(history.history['val_loss'],'b',linewidth=3.0)
plt.legend(['Training loss', 'Validation Loss'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('Loss',fontsize=16)
plt.title('Loss Curves',fontsize=16)
```
#### CNN using CONV+ POOLING Layers
```
model=Sequential()
model.add(Conv2D(filters=16,kernel_size=3,activation='relu',input_shape=(32, 32, 3)))
model.add(MaxPool2D(pool_size=2))
model.add(Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(MaxPool2D(pool_size=2))
model.add(Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(MaxPool2D(pool_size=2))
model.add(Flatten())
model.add(Dense(units=512,activation='relu'))
model.add(Dense(units=512,activation='relu'))
model.add(Dense(units=10,activation='softmax'))
model.summary()
#Compile & Train the model
model.compile(loss="categorical_crossentropy", optimizer="SGD", metrics=["accuracy"])
history = model.fit(x_train, y_train, validation_data=(x_valid,y_valid),
batch_size=200,
epochs=5,
verbose=1)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['loss'],'r',linewidth=3.0)
plt.plot(history.history['val_loss'],'b',linewidth=3.0)
plt.legend(['Training loss', 'Validation Loss'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('Loss',fontsize=16)
plt.title('Loss Curves',fontsize=16)
import matplotlib.pyplot as plt
plt.figure(figsize=[8,6])
plt.plot(history.history['accuracy'],'r',linewidth=3.0)
plt.plot(history.history['val_accuracy'],'b',linewidth=3.0)
plt.legend(['Training accuracy', 'Validation accuracy'],fontsize=18)
plt.xlabel('Epochs ',fontsize=16)
plt.ylabel('accuracy',fontsize=16)
plt.title('Accuracy Curves',fontsize=16)
```
## Build a CNN Architecture with Tensorflow
```
import tensorflow as tf
del model
model= tf.keras.Sequential()
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu',input_shape=(32,32,3)))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Flatten())
model.add(Dense(10, activation='softmax'))
model.compile(optimizer='adam',loss="categorical_crossentropy",metrics=['accuracy'])
model.summary()
x_train.shape,y_train.shape
answers=model.fit(x_train,y_train,
validation_data=(x_valid,y_valid),
shuffle=True,epochs=2,verbose=2,batch_size=200)
```
### Stack an ANN on top of CNN
```
model= tf.keras.Sequential()
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu',input_shape=(32,32,3)))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Conv2D(filters=16,kernel_size=3,activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Flatten())
model.summary()
### Stack an ANN on top of CNN
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=512,activation='relu',kernel_initializer='uniform'))
model.add(tf.keras.layers.Dense(units=10,activation='softmax',kernel_initializer='uniform'))
model.compile(optimizer='adam',loss="categorical_crossentropy",metrics=['accuracy'])
model.summary()
```
### Fit the model
```
answers=model.fit(x_train,y_train,validation_data=(x_valid,y_valid),shuffle=True,epochs=2,verbose=2,batch_size=500)
```
### Evaluate the Results
```
results=model.evaluate(x_test,y_test)
answers.history.keys()
plt.plot(answers.history['val_loss'],label='Validation Loss');
plt.plot(answers.history['loss'],label='Training Loss');
plt.legend();
plt.plot(answers.history['val_accuracy'],label='Validation Accuracy');
plt.plot(answers.history['accuracy'],label='Training Accuracy');
plt.legend;
```
# Approch -3 CNN
```
from keras.layers import Conv2D, Flatten
simple_cnn_model = Sequential()
simple_cnn_model.add(Conv2D(32, (3,3), input_shape=(32,32,3), activation='relu'))
simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
simple_cnn_model.add(Flatten())
simple_cnn_model.add(Dense(10, activation='softmax'))
simple_cnn_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
simple_cnn_model_history = simple_cnn_model.fit(x_train,y_train,validation_data=(x_valid,y_valid), batch_size=100, epochs=8)
```
# CNN - Experiment - 4
```
from keras.layers import Conv2D, Flatten
simple_cnn_model = Sequential()
simple_cnn_model.add(Conv2D(32, (3,3), input_shape=(32,32,3), activation='relu'))
simple_cnn_model.add(MaxPool2D(pool_size=2))
simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
simple_cnn_model.add(MaxPool2D(pool_size=2))
simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
simple_cnn_model.add(Flatten())
simple_cnn_model.add(Dense(10, activation='softmax'))
simple_cnn_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
simple_cnn_model_history = simple_cnn_model.fit(x_train,y_train,validation_data=(x_valid,y_valid), batch_size=100, epochs=8)
```
# Alternative Approach
https://www.kaggle.com/aayushkubba/cifar-10-analysis-with-a-neural-network
## Data Augmentation
To get a more generalised model
```
from keras.preprocessing.image import ImageDataGenerator
datagen = ImageDataGenerator(
rotation_range=20,
width_shift_range=0.2,
height_shift_range=0.2,
horizontal_flip=True, vertical_flip=True,
validation_split=0.2)
type(datagen)
from keras.layers import Conv2D, Flatten
augmented_simple_cnn_model = Sequential()
augmented_simple_cnn_model.add(Conv2D(32, (3,3), input_shape=(32,32,3), activation='relu'))
augmented_simple_cnn_model.add(MaxPool2D(pool_size=2))
augmented_simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
augmented_simple_cnn_model.add(MaxPool2D(pool_size=2))
augmented_simple_cnn_model.add(Conv2D(32, (3,3), activation='relu'))
augmented_simple_cnn_model.add(Flatten())
augmented_simple_cnn_model.add(Dense(10, activation='softmax'))
augmented_simple_cnn_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
augmented_simple_cnn_model.summary()
augmented_simple_cnn_model_history=augmented_simple_cnn_model.fit_generator(datagen.flow(x_train,y_train,batch_size=256),
validation_data=(x_valid,y_valid),
epochs=8)
plt.plot(augmented_simple_cnn_model_history.history['val_loss'],label='Validation Loss');
plt.plot(augmented_simple_cnn_model_history.history['loss'],label='Training Loss');
plt.legend();
plt.plot(augmented_simple_cnn_model_history.history['val_accuracy'],label='Validation Accuracy');
plt.plot(augmented_simple_cnn_model_history.history['accuracy'],label='Training Accuracy');
plt.legend;
```
# Transfer Learning
## RESNET
```
from keras.models import Model, load_model
from tensorflow.keras.applications.resnet50 import preprocess_input
from keras.layers import GlobalAveragePooling2D, Dropout
# RESNET as the base model
base_model=tf.keras.applications.ResNet50(weights='imagenet',include_top=False)
# We will only use resnet for feature extarction, freeze all the layers that we dont need
for layer in base_model.layers:
layer.trainable=False
# Take the output output of the base layers
base_model_output=base_model.output
# Add our layers
## Pooling
x=GlobalAveragePooling2D()(base_model_output)
## FC
x=Dense(2056,activation='relu')(x)
## Dropout
x=Dropout(0.5)(x)
## Output layer
x=Dense(10,activation='softmax')(x)
resnet_model=Model(inputs=base_model.input,outputs=x)
resnet_model.summary()
resnet_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
```
For reference
Without freezing
Epoch 1/8
176/176 [==============================] - 50s 238ms/step - loss: 1.4530 - accuracy: 0.5499 - val_loss: 4.2816 - val_accuracy: 0.1038
Epoch 2/8
```
resnet_model_history=resnet_model.fit(x_train,y_train,batch_size=256,
validation_data=(x_valid,y_valid),epochs=4,shuffle=True)
```
# VGGNET
```
from keras.models import Model, load_model
from tensorflow.keras.applications.resnet50 import preprocess_input
from keras.layers import GlobalAveragePooling2D, Dropout, BatchNormalization
# RESNET as the base model
base_model=tf.keras.applications.VGG16(weights='imagenet',include_top=False)
# We will only use resnet for feature extarction, freeze all the layers that we dont need
for layer in base_model.layers:
layer.trainable=False
# Take the output output of the base layers
base_model_output=base_model.output
# Add our layers
## Pooling
x=GlobalAveragePooling2D()(base_model_output)
## FC
x=Dense(2056,activation='relu')(x)
## BAtch Normalisation - For Speed
x= BatchNormalization()(x)
## Dropout
x=Dropout(0.5)(x)
## Output layer
x=Dense(10,activation='softmax')(x)
vgg_model=Model(inputs=base_model.input,outputs=x)
vgg_model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
vgg_model_history=vgg_model.fit(x_train,y_train,batch_size=256,
validation_data=(x_valid,y_valid),epochs=40,shuffle=True)
```
| github_jupyter |
```
from sklearn.ensemble import RandomForestRegressor
from sklearn.datasets import make_regression
import pandas as pd
import numpy as np
from sklearn import preprocessing
from sklearn.model_selection import train_test_split
import numpy as np, tensorflow as tf
from sklearn.preprocessing import OneHotEncoder
import os
import csv
import gc
from sklearn.metrics import mean_squared_error
import math
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import ConstantKernel, RBF
from sklearn.gaussian_process.kernels import RationalQuadratic
from sklearn.model_selection import GridSearchCV
from sklearn.model_selection import RepeatedKFold
from sklearn import linear_model
from xgboost.sklearn import XGBRegressor
from sklearn.decomposition import PCA
import copy
import pyflux as pf
import datetime
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
PRICED_BITCOIN_FILE_PATH = "C:/Users/wang.yuhao/Documents/ChainNet/data/original_data/pricedBitcoin2009-2018.csv"
DAILY_OCCURRENCE_FILE_PATH = "C:/Users/wang.yuhao/Documents/ChainNet/data/original_data/dailyOccmatrices/"
betti0_input_path = "C:/Users/wang.yuhao/Documents/ChainNet/data/original_data/betti_0(100).csv"
betti1_input_path = "C:/Users/wang.yuhao/Documents/ChainNet/data/original_data/betti_1(100).csv"
DAILY_FILTERED_OCCURRENCE_FILE_PATH = "C:/Users/wang.yuhao/Documents/ChainNet/data/original_data/filteredDailyOccMatrices/"
ROW = -1
COLUMN = -1
TEST_SPLIT = 0.01
ALL_YEAR_INPUT_ALLOWED = False
YEAR = 2017
START_YEAR = 2017
END_YEAR = 2018
SLIDING_BATCH_SIZE = 7
# Baseline
from sklearn.metrics import mean_squared_error
from sklearn import metrics
import matplotlib.pyplot as plt
def exclude_days(train, test):
row, column = train.shape
train_days = np.asarray(train[:, -1]).reshape(-1, 1)
x_train = train[:, 0:column - 1]
test_days = np.asarray(test[:, -1]).reshape(-1, 1)
x_test = test[:, 0:column - 1]
return x_train, x_test, train_days, test_days
def merge_data(occurrence_data, daily_occurrence_normalized_matrix, aggregation_of_previous_days_allowed):
if(aggregation_of_previous_days_allowed):
if(occurrence_data.size==0):
occurrence_data = daily_occurrence_normalized_matrix
else:
occurrence_data = np.add(occurrence_data, daily_occurrence_normalized_matrix)
else:
if(occurrence_data.size == 0):
occurrence_data = daily_occurrence_normalized_matrix
else:
occurrence_data = np.concatenate((occurrence_data, daily_occurrence_normalized_matrix), axis=0)
#print("merge_data shape: {} occurrence_data: {} ".format(occurrence_data.shape, occurrence_data))
return occurrence_data
def get_normalized_matrix_from_file(day, year, totaltx):
daily_occurrence_matrix_path_name = DAILY_OCCURRENCE_FILE_PATH + "occ" + str(year) + '{:03}'.format(day) + '.csv'
daily_occurence_matrix = pd.read_csv(daily_occurrence_matrix_path_name, sep=",", header=None).values
return np.asarray(daily_occurence_matrix).reshape(1, daily_occurence_matrix.size)/totaltx
def fl_get_normalized_matrix_from_file(day, year, totaltx, n_components):
daily_occurence_matrix = np.asarray([],dtype=np.float32)
for filter_number in range(0, 50, 10):
daily_occurrence_matrix_path_name = DAILY_FILTERED_OCCURRENCE_FILE_PATH + "occ" + str(year) + '{:03}'.format(day) + "_" + str(filter_number) +'.csv'
daily_occurence_matrix_read = pd.read_csv(daily_occurrence_matrix_path_name, sep=",", header=None).values
if(daily_occurence_matrix.size == 0):
daily_occurence_matrix = daily_occurence_matrix_read
else:
daily_occurence_matrix = np.concatenate((daily_occurence_matrix, daily_occurence_matrix_read), axis = 1)
#print("daily_occurence_matrix: ", daily_occurence_matrix, daily_occurence_matrix.shape)
#return np.asarray(daily_occurence_matrix).reshape(1, daily_occurence_matrix.size)/totaltx
return np.asarray(daily_occurence_matrix).reshape(1, daily_occurence_matrix.size)
def get_daily_occurrence_matrices(priced_bitcoin, current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
#print("priced_bitcoin: ", priced_bitcoin, priced_bitcoin.shape)
#print("current_row: ", current_row, current_row.shape)
previous_price_data = np.array([], dtype=np.float32)
occurrence_data = np.array([], dtype=np.float32)
for index, row in priced_bitcoin.iterrows():
if not ((row.values == current_row.values).all()):
previous_price_data = np.append(previous_price_data, row['price'])
previous_price_data = np.append(previous_price_data, row['totaltx'])
#print("previous_price_data: ", previous_price_data,row['day'], row['year'], row['totaltx'])
#print("occurrence_data: ", occurrence_data)
if(is_price_of_previous_days_allowed):
#print("previous_price_data: ", np.asarray(previous_price_data).reshape(1, -1), np.asarray(previous_price_data).reshape(1, -1).shape)
occurrence_data = np.asarray(previous_price_data).reshape(1, -1)
occurrence_input = np.concatenate((occurrence_data, np.asarray(current_row['price']).reshape(1,1)), axis=1)
#print("current_row: ", current_row, current_row.shape)
#print(" price occurrence_input: ", np.asarray(current_row['price']).reshape(1,1), (np.asarray(current_row['price']).reshape(1,1)).shape)
#print("concatenate with price occurrence_input: ", occurrence_input, occurrence_input.shape)
occurrence_input = np.concatenate((occurrence_input, np.asarray(current_row['day']).reshape(1,1)), axis=1)
#print(" price occurrence_input: ", np.asarray(current_row['day']).reshape(1,1), (np.asarray(current_row['day']).reshape(1,1)).shape)
#print("concatenate with day occurrence_input: ", occurrence_input, occurrence_input.shape)
return occurrence_input
def betti_get_daily_occurrence_matrices(priced_bitcoin, current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
previous_price_data = np.array([], dtype=np.float32)
occurrence_data = np.array([], dtype=np.float32)
for index, row in priced_bitcoin.iterrows():
if not ((row.values == current_row.values).all()):
previous_price_data = np.append(previous_price_data, row['price'])
previous_price_data = np.append(previous_price_data, row['totaltx'])
betti0_50 = read_betti(betti0_input_path, row['day'])
occurrence_data = np.append(occurrence_data, np.asarray(betti0_50).reshape(1,-1))
betti1_50 = read_betti(betti1_input_path, row['day'])
occurrence_data = np.append(occurrence_data, np.asarray(betti1_50).reshape(1,-1))
if (is_price_of_previous_days_allowed):
occurrence_data = np.concatenate((np.asarray(previous_price_data).reshape(1,-1),occurrence_data.reshape(1,-1)), axis=1)
occurrence_input = np.concatenate((occurrence_data.reshape(1,-1), np.asarray(current_row['price']).reshape(1,1)), axis=1)
occurrence_input = np.concatenate((occurrence_input, np.asarray(current_row['day']).reshape(1,1)), axis=1)
#print("occurrence_input: ",occurrence_input, occurrence_input.shape)
return occurrence_input
def betti_der_get_daily_occurrence_matrices(priced_bitcoin, current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
previous_price_data = np.array([], dtype=np.float32)
occurrence_data = np.array([], dtype=np.float32)
for index, row in priced_bitcoin.iterrows():
if not ((row.values == current_row.values).all()):
previous_price_data = np.append(previous_price_data, row['price'])
previous_price_data = np.append(previous_price_data, row['totaltx'])
betti0_50 = read_betti(betti0_input_path, row['day'])
occurrence_data = np.append(occurrence_data, np.asarray(betti0_50).reshape(1,-1))
betti1_50 = read_betti(betti1_input_path, row['day'])
occurrence_data = np.append(occurrence_data, np.asarray(betti1_50).reshape(1,-1))
betti0_50_diff1 = betti0_50.diff(1).dropna()
occurrence_data = np.concatenate((occurrence_data.reshape(1,-1), np.asarray(betti0_50_diff1).reshape(1,-1)), axis=1)
betti1_50_diff1 = betti1_50.diff(1).dropna()
occurrence_data = np.concatenate((occurrence_data, np.asarray(betti1_50_diff1).reshape(1,-1)), axis=1)
#print("previous_price_data:",previous_price_data, previous_price_data.shape)
if (is_price_of_previous_days_allowed):
occurrence_data = np.concatenate((np.asarray(previous_price_data).reshape(1,-1),occurrence_data.reshape(1,-1)), axis=1)
occurrence_input = np.concatenate((occurrence_data.reshape(1,-1), np.asarray(current_row['price']).reshape(1,1)), axis=1)
occurrence_input = np.concatenate((occurrence_input, np.asarray(current_row['day']).reshape(1,1)), axis=1)
#print("occurrence_input: ",occurrence_input, occurrence_input.shape)
return occurrence_input
def fl_get_daily_occurrence_matrices(priced_bitcoin, current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
previous_price_data = np.array([], dtype=np.float32)
occurrence_data = np.array([], dtype=np.float32)
for index, row in priced_bitcoin.iterrows():
if not ((row.values == current_row.values).all()):
previous_price_data = np.append(previous_price_data, row['price'])
previous_price_data = np.append(previous_price_data, row['totaltx'])
daily_occurrence_normalized_matrix = fl_get_normalized_matrix_from_file(row['day'], row['year'], row['totaltx'], 20)
occurrence_data = merge_data(occurrence_data, daily_occurrence_normalized_matrix, aggregation_of_previous_days_allowed)
#print("occurrence_data: ",occurrence_data, occurrence_data.shape)
if (is_price_of_previous_days_allowed):
occurrence_data = np.concatenate((np.asarray(previous_price_data).reshape(1,-1),occurrence_data.reshape(1,-1)), axis=1)
occurrence_input = np.concatenate((occurrence_data.reshape(1,-1), np.asarray(current_row['price']).reshape(1,1)), axis=1)
occurrence_input = np.concatenate((occurrence_input, np.asarray(current_row['day']).reshape(1,1)), axis=1)
#print("occurrence_input: ",occurrence_input, occurrence_input.shape)
return occurrence_input
def read_betti(file_path, day):
day = day - 1
betti = pd.read_csv(file_path, index_col=0)
try:
betti_50 = betti.iloc[day, 0:50]
except:
print("day:", day)
return betti_50
def rf_mode(train_input, train_target, test_input, test_target):
param = {
'n_estimators':400
}
rf_regression = RandomForestRegressor(**param)
rf_regression.fit(train_input, train_target.ravel() )
rf_predicted = rf_regression.predict(test_input)
return rf_predicted
def gp_mode(train_input, train_target, test_input, test_target):
param = {
'kernel': RationalQuadratic(alpha=0.01, length_scale=1),
'n_restarts_optimizer': 2
}
gpr = GaussianProcessRegressor(**param)
gpr.fit(train_input,train_target.ravel())
gp_predicted = gpr.predict(test_input)
return gp_predicted
def enet_mode(train_input, train_target, test_input, test_target):
param = {
'alpha': 10,
'l1_ratio': 1,
}
elastic = linear_model.ElasticNet(**param)
elastic.fit(train_input,train_target.ravel())
enet_predicted = elastic.predict(test_input)
return enet_predicted
def xgbt_mode(train_input, train_target, test_input, test_target):
param = {
'n_estimators':1000,
'learning_rate': 0.01,
'objective': 'reg:squarederror',
}
xgbt = XGBRegressor(**param)
xgbt.fit(train_input,train_target.ravel())
xgbt_predicted = xgbt.predict(test_input)
return xgbt_predicted
def arimax_initialize_setting(dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
data = preprocess_data(dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
train = data[0:100, :]
test = data[100:100+prediction_horizon, :]
x_train, x_test, train_days, test_days = exclude_days(train, test)
row, column = x_train.shape
train_target = np.asarray(x_train[:, -1]).reshape(-1)
train_input = x_train[:, 0:column - 1]
test_target = x_test[: , -1]
test_input = x_test[ : , 0:column - 1]
return train_input, train_target, test_input, test_target, train_days, test_days
def arimax_base_rmse_mode(train_input, train_target, test_input, test_target):
train_input_diff_arr = np.array([])
train_columns_name = []
train_input_column = int(train_input.shape[1])
for i in range(train_input_column):
if(i%2==0):
train_columns_name.append('price_' + str(i))
else:
train_columns_name.append('totaltx_' + str(i))
train_input_diff = np.diff(train_input[:,i] )
if i == 0:
train_input_diff_arr = train_input_diff
else:
train_input_diff_arr = np.dstack((train_input_diff_arr, train_input_diff))
columns_name = copy.deepcopy(train_columns_name)
columns_name.append('current_price')
train_target_diff = np.diff(train_target )
train_input_diff_arr = np.dstack((train_input_diff_arr, train_target_diff))
train_input_diff_arr = pd.DataFrame(train_input_diff_arr[0], columns = columns_name)
model = pf.ARIMAX(data=train_input_diff_arr,formula="current_price~totaltx_5",ar=1,ma=2,integ=0)
model_1 = model.fit("MLE")
model_1.summary()
test_input_pd = pd.DataFrame(test_input, columns = train_columns_name)
test_target_pd = pd.DataFrame(test_target, columns = ['current_price'])
test_input_target = pd.concat([test_input_pd, test_target_pd], axis=1)
pred = model.predict(h=test_input_target.shape[0],
oos_data=test_input_target,
intervals=True, )
arimax_base_rmse = mean_squared_error([test_input_target.iloc[0, 6]],[(train_target[99])+pred.current_price[99]])
print("arimax_base_rmse:",arimax_base_rmse)
return arimax_base_rmse
def run_print_model(train_input, train_target, test_input, test_target, train_days, test_days):
rf_prediction = rf_mode(train_input, train_target, test_input, test_target)
xgbt_prediction = xgbt_mode(train_input, train_target, test_input, test_target)
gp_prediction = gp_mode(train_input, train_target, test_input, test_target)
enet_prediction = enet_mode(train_input, train_target, test_input, test_target)
return rf_prediction, xgbt_prediction, gp_prediction, enet_prediction
#print_results(predicted, test_target, original_log_return, predicted_log_return, cost, test_days, rmse)
#return rf_base_rmse
def filter_data(priced_bitcoin, window_size):
end_day_of_previous_year = max(priced_bitcoin[priced_bitcoin['year'] == START_YEAR-1]["day"].values)
start_index_of_previous_year = end_day_of_previous_year - window_size
previous_year_batch = priced_bitcoin[(priced_bitcoin['year'] == START_YEAR-1) & (priced_bitcoin['day'] > start_index_of_previous_year)]
input_batch = priced_bitcoin[(priced_bitcoin['year'] >= START_YEAR) & (priced_bitcoin['year'] <= END_YEAR)]
filtered_data = previous_year_batch.append(input_batch)
filtered_data.insert(0, 'index', range(0, len(filtered_data)))
filtered_data = filtered_data.reset_index(drop=True)
return filtered_data
def preprocess_data(dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
priced_bitcoin = pd.read_csv(PRICED_BITCOIN_FILE_PATH, sep=",")
if(ALL_YEAR_INPUT_ALLOWED):
pass
else:
#priced_bitcoin = filter_data(priced_bitcoin, window_size)
priced_bitcoin = priced_bitcoin[priced_bitcoin['year']==YEAR].reset_index(drop=True)
#print("priced_bitcoin:",priced_bitcoin)
daily_occurrence_input = np.array([],dtype=np.float32)
temp = np.array([], dtype=np.float32)
for current_index, current_row in priced_bitcoin.iterrows():
if(current_index<(window_size+prediction_horizon-1)):
pass
else:
start_index = current_index - (window_size + prediction_horizon) + 1
end_index = current_index - prediction_horizon
if(dataset_model=="base"):
temp = get_daily_occurrence_matrices(priced_bitcoin[start_index:end_index+1], current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
elif(dataset_model=="betti"):
temp = betti_get_daily_occurrence_matrices(priced_bitcoin[start_index:end_index+1], current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
elif(dataset_model=="fl"):
temp = fl_get_daily_occurrence_matrices(priced_bitcoin[start_index:end_index+1], current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
elif(dataset_model=="betti_der"):
temp = betti_der_get_daily_occurrence_matrices(priced_bitcoin[start_index:end_index+1], current_row, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
else:
sys.exit("Dataset model support only baseline, betti, fl and betti_der!")
if(daily_occurrence_input.size == 0):
daily_occurrence_input = temp
else:
daily_occurrence_input = np.concatenate((daily_occurrence_input, temp), axis=0)
return daily_occurrence_input
def initialize_setting( features, price, day, test_start, dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed):
train_target = price[test_start : test_start + 100]
train_days = day[test_start : test_start + 100]
pca_features = features[test_start : test_start + 100+1, :]
train_input = pca_features[0 : 100, :]
#x_test = x_test.reshape(-1,1)
test_target = price[test_start + 100]
test_days = day[test_start + 100]
#print(pca_features, pca_features.shape)
test_input = pca_features[ 100, :].reshape(1, -1)
#print(pca_features, pca_features.shape)
#print("***"*20)
#print(train_input,train_input.shape, train_target, test_input,test_input.shape, test_target, train_days, test_days)
return train_input, train_target, test_input, test_target, train_days, test_days
def split_process(data,dataset_model,window_size):
baseline_features = data[:, 0:window_size*2]
fl_features = data[: , window_size*2:-2]
price = data[:, -2]
day = data[:,-1]
return baseline_features, fl_features, price, day
parameter_dict = {#0: dict({'is_price_of_previous_days_allowed':True, 'aggregation_of_previous_days_allowed':True})}
1: dict({'is_price_of_previous_days_allowed':True, 'aggregation_of_previous_days_allowed':False})}
for step in parameter_dict:
t = datetime.datetime.now()
dir_name = t.strftime('%m_%d___%H_%M')
drive_path = "drive/MyDrive/Colab Notebooks/ChainNet/processed_data/"+dir_name
if not os.path.exists(dir_name):
os.makedirs(drive_path)
print("drive_path: ", drive_path)
result_path = drive_path + "/"
names = locals()
gc.collect()
evalParameter = parameter_dict.get(step)
is_price_of_previous_days_allowed = evalParameter.get('is_price_of_previous_days_allowed')
aggregation_of_previous_days_allowed = evalParameter.get('aggregation_of_previous_days_allowed')
print("IS_PRICE_OF_PREVIOUS_DAYS_ALLOWED: ", is_price_of_previous_days_allowed)
print("AGGREGATION_OF_PREVIOUS_DAYS_ALLOWED: ", aggregation_of_previous_days_allowed)
window_size_array = [3, 5, 7]
horizon_size_array = [1, 2, 5, 7, 10, 15, 20, 25, 30]
dataset_model_array = ["base","betti", "betti_der","fl"]
for dataset_model in dataset_model_array:
print('dataset_model: ', dataset_model)
for window_size in window_size_array:
print('WINDOW_SIZE: ', window_size)
for prediction_horizon in horizon_size_array:
data = preprocess_data(dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
baseline_features, fl_features, price, day = split_process(data, dataset_model, window_size)
if dataset_model == "base":
#print("features: ", baseline_features, fl_features, price, day)
features = baseline_features
#print("pca features: ", features, features.shape)
else:
#print("features: ", baseline_features, fl_features, price, day)
pca = PCA(n_components = 20)
pca.fit(fl_features)
fl_features = pca.transform(fl_features)
features = np.concatenate((baseline_features, fl_features), axis=1)
#print("pca features: ", features, features.shape)
print("PREDICTION_HORIZON: ", prediction_horizon)
for test_start in range(0, 100):
#print("test_start: ", test_start)
train_input, train_target, test_input, test_target, train_days, test_days = initialize_setting( features, price, day, test_start, dataset_model, window_size, prediction_horizon, is_price_of_previous_days_allowed, aggregation_of_previous_days_allowed)
rf_prediction, xgbt_prediction, gp_prediction, enet_prediction = run_print_model(train_input, train_target, test_input, test_target, train_days, test_days)
prediction = pd.DataFrame({'rf_' + dataset_model + '_prediction_'+str(window_size): [rf_prediction], 'xgbt_' + dataset_model + '_prediction_'+str(window_size): [xgbt_prediction], 'gp_' + dataset_model + '_prediction_'+str(window_size): [gp_prediction], 'enet_' + dataset_model + '_prediction_'+str(window_size): [enet_prediction]})
test_target_df = pd.DataFrame({'test_target': [test_target]})
#print("prediction: ",prediction)
if(test_start==0):
prediction_total = prediction
test_target_total = test_target_df
else:
prediction_total = [prediction_total, prediction]
test_target_total = [test_target_total, test_target_df]
prediction_total = pd.concat(prediction_total)
test_target_total = pd.concat(test_target_total)
print("+++"*10)
print("prediction_total: ",prediction_total)
print("test_target_total: ",test_target_total)
rmse = ((((prediction_total.sub(test_target_total.values))**2).mean())**0.5).to_frame().T
print("rmse: ",rmse)
#rmse.to_csv(result_path + "rmse_" + dataset_model + "_"+ str(window_size) +"_"+ str(prediction_horizon) + ".csv", index=True)
if(prediction_horizon==1):
rmse_total = rmse
else:
rmse_total = [rmse_total, rmse]
rmse_total = pd.concat(rmse_total)
if(window_size==3):
names['rmse_' + dataset_model + '_total'] = rmse_total
else:
names['rmse_' + dataset_model + '_total'] = pd.concat([names.get('rmse_' + dataset_model + '_total') , rmse_total], axis=1)
names['rmse_' + dataset_model + '_total'].index = pd.Series(horizon_size_array)
names.get('rmse_' + dataset_model + '_total').to_csv(result_path + "rmse_" + dataset_model + "_total.csv", index=True)
print('rmse_{}_total = {}'.format(dataset_model, names.get('rmse_' + dataset_model + '_total')))
betti_gain = 100 * (1 -rmse_betti_total.div(rmse_base_total.values))
fl_gain = 100 * (1 -rmse_fl_total.div(rmse_base_total.values))
betti_der_gain = 100 * (1 -rmse_betti_der_total.div(rmse_base_total.values))
for i in range(12):
perf = pd.concat([betti_gain.iloc[:,i],betti_der_gain.iloc[:,i], fl_gain.iloc[:, i]], axis=1).plot.bar()
modelnames = ["rf","xgbt","gp","enet"]
windows = [3, 5, 7]
filename = result_path + modelnames[int(i%4)] +"_window_" + str(windows[int(i/4)])
perf.figure.savefig(filename)
horizon_size_array_str = [str(x) for x in horizon_size_array]
rmse_base_total.index = pd.Series(horizon_size_array_str)
base_rmse_window_3_line = rmse_base_total.iloc[:,0:4].plot()
base_rmse_window_3_line.figure.savefig(result_path + "base_rmse_window_3_line")
base_rmse_window_5_line = rmse_base_total.iloc[:,4:8].plot()
base_rmse_window_5_line.figure.savefig(result_path + "base_rmse_window_5_line")
base_rmse_window_7_line = rmse_base_total.iloc[:,8:12].plot()
base_rmse_window_7_line.figure.savefig(result_path + "base_rmse_window_7_line")
horizon_size_array_str = [str(x) for x in horizon_size_array]
rmse_base_total.index = pd.Series(horizon_size_array_str)
base_rmse_window_3_line = rmse_base_total.iloc[:,0:4].plot()
base_rmse_window_3_line.figure.savefig(result_path + "base_rmse_window_3_line")
base_rmse_window_5_line = rmse_base_total.iloc[:,4:8].plot()
base_rmse_window_5_line.figure.savefig(result_path + "base_rmse_window_5_line")
base_rmse_window_7_line = rmse_base_total.iloc[:,8:12].plot()
base_rmse_window_7_line.figure.savefig(result_path + "base_rmse_window_7_line")
pd.Series(horizon_size_array).map(str)
horizon_size_array_str
rmse_base_total.index
base_rmse_window_5_line = rmse_base_total.iloc[:,4:8].plot()
base_rmse_window_5_line.figure.savefig(result_path + "base_rmse_window_5_line")
base_rmse_window_7_line = rmse_base_total.iloc[:,8:12].plot()
base_rmse_window_7_line.figure.savefig(result_path + "base_rmse_window_7_line")
rmse_base_total
rmse_base_total.iloc[:,0:4].plot()
```
| github_jupyter |
# Chapter 3: Word2vec - Learning Word Embeddings
** Warning **:This code can consume a significant amount of memory. If you have significantly large RAM memory (>4GB) you do not need to worry. Otherwise please reduce the `batch_size` or `embedding_size` parameter to allow the model to fit in memory.
```
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
%matplotlib inline
import collections
import math
import numpy as np
import os
import random
import tensorflow as tf
import bz2
from matplotlib import pylab
from six.moves import range
from six.moves.urllib.request import urlretrieve
from sklearn.manifold import TSNE
from sklearn.cluster import KMeans
import nltk # standard preprocessing
import operator # sorting items in dictionary by value
#nltk.download() #tokenizers/punkt/PY3/english.pickle
from math import ceil
import csv
```
## Dataset
This code downloads a [dataset](http://www.evanjones.ca/software/wikipedia2text.html) consisting of several Wikipedia articles totaling up to roughly 61 megabytes. Additionally the code makes sure the file has the correct size after downloading it.
```
url = 'http://www.evanjones.ca/software/'
def maybe_download(filename, expected_bytes):
"""Download a file if not present, and make sure it's the right size."""
if not os.path.exists(filename):
print('Downloading file...')
filename, _ = urlretrieve(url + filename, filename)
statinfo = os.stat(filename)
if statinfo.st_size == expected_bytes:
print('Found and verified %s' % filename)
else:
print(statinfo.st_size)
raise Exception(
'Failed to verify ' + filename + '. Can you get to it with a browser?')
return filename
filename = maybe_download('wikipedia2text-extracted.txt.bz2', 18377035)
```
## Read Data without Preprocessing
Reads data as it is to a string and tokenize it using spaces and returns a list of words
```
def read_data(filename):
"""Extract the first file enclosed in a zip file as a list of words"""
with bz2.BZ2File(filename) as f:
data = []
file_string = f.read().decode('utf-8')
file_string = nltk.word_tokenize(file_string)
data.extend(file_string)
return data
words = read_data(filename)
print('Data size %d' % len(words))
print('Example words (start): ',words[:10])
print('Example words (end): ',words[-10:])
```
## Read Data with Preprocessing with NLTK
Reads data as it is to a string, convert to lower-case and tokenize it using the nltk library. This code reads data in 1MB portions as processing the full text at once slows down the task and returns a list of words. You will have to download the necessary tokenizer.
```
def read_data(filename):
"""
Extract the first file enclosed in a zip file as a list of words
and pre-processes it using the nltk python library
"""
with bz2.BZ2File(filename) as f:
data = []
file_size = os.stat(filename).st_size
chunk_size = 1024 * 1024 # reading 1 MB at a time as the dataset is moderately large
print('Reading data...')
for i in range(ceil(file_size//chunk_size)+1):
bytes_to_read = min(chunk_size,file_size-(i*chunk_size))
file_string = f.read(bytes_to_read).decode('utf-8')
file_string = file_string.lower()
# tokenizes a string to words residing in a list
file_string = nltk.word_tokenize(file_string)
data.extend(file_string)
return data
words = read_data(filename)
print('Data size %d' % len(words))
print('Example words (start): ',words[:10])
print('Example words (end): ',words[-10:])
```
## Building the Dictionaries
Builds the following. To understand each of these elements, let us also assume the text "I like to go to school"
* `dictionary`: maps a string word to an ID (e.g. {I:0, like:1, to:2, go:3, school:4})
* `reverse_dictionary`: maps an ID to a string word (e.g. {0:I, 1:like, 2:to, 3:go, 4:school}
* `count`: List of list of (word, frequency) elements (e.g. [(I,1),(like,1),(to,2),(go,1),(school,1)]
* `data` : Contain the string of text we read, where string words are replaced with word IDs (e.g. [0, 1, 2, 3, 2, 4])
It also introduces an additional special token `UNK` to denote rare words to are too rare to make use of.
```
# we restrict our vocabulary size to 50000
vocabulary_size = 50000
def build_dataset(words):
count = [['UNK', -1]]
# Gets only the vocabulary_size most common words as the vocabulary
# All the other words will be replaced with UNK token
count.extend(collections.Counter(words).most_common(vocabulary_size - 1))
dictionary = dict()
# Create an ID for each word by giving the current length of the dictionary
# And adding that item to the dictionary
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
# Traverse through all the text we have and produce a list
# where each element corresponds to the ID of the word found at that index
for word in words:
# If word is in the dictionary use the word ID,
# else use the ID of the special token "UNK"
if word in dictionary:
index = dictionary[word]
else:
index = 0 # dictionary['UNK']
unk_count = unk_count + 1
data.append(index)
# update the count variable with the number of UNK occurences
count[0][1] = unk_count
reverse_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
# Make sure the dictionary is of size of the vocabulary
assert len(dictionary) == vocabulary_size
return data, count, dictionary, reverse_dictionary
data, count, dictionary, reverse_dictionary = build_dataset(words)
print('Most common words (+UNK)', count[:5])
print('Sample data', data[:10])
del words # Hint to reduce memory.
```
## Generating Batches of Data for Skip-Gram
Generates a batch or target words (`batch`) and a batch of corresponding context words (`labels`). It reads `2*window_size+1` words at a time (called a `span`) and create `2*window_size` datapoints in a single span. The function continue in this manner until `batch_size` datapoints are created. Everytime we reach the end of the word sequence, we start from beginning.
```
data_index = 0
def generate_batch_skip_gram(batch_size, window_size):
# data_index is updated by 1 everytime we read a data point
global data_index
# two numpy arras to hold target words (batch)
# and context words (labels)
batch = np.ndarray(shape=(batch_size), dtype=np.int32)
labels = np.ndarray(shape=(batch_size, 1), dtype=np.int32)
# span defines the total window size, where
# data we consider at an instance looks as follows.
# [ skip_window target skip_window ]
span = 2 * window_size + 1
# The buffer holds the data contained within the span
buffer = collections.deque(maxlen=span)
# Fill the buffer and update the data_index
for _ in range(span):
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
# This is the number of context words we sample for a single target word
num_samples = 2*window_size
# We break the batch reading into two for loops
# The inner for loop fills in the batch and labels with
# num_samples data points using data contained withing the span
# The outper for loop repeat this for batch_size//num_samples times
# to produce a full batch
for i in range(batch_size // num_samples):
k=0
# avoid the target word itself as a prediction
# fill in batch and label numpy arrays
for j in list(range(window_size))+list(range(window_size+1,2*window_size+1)):
batch[i * num_samples + k] = buffer[window_size]
labels[i * num_samples + k, 0] = buffer[j]
k += 1
# Everytime we read num_samples data points,
# we have created the maximum number of datapoints possible
# withing a single span, so we need to move the span by 1
# to create a fresh new span
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
return batch, labels
print('data:', [reverse_dictionary[di] for di in data[:8]])
for window_size in [1, 2]:
data_index = 0
batch, labels = generate_batch_skip_gram(batch_size=8, window_size=window_size)
print('\nwith window_size = %d:' %window_size)
print(' batch:', [reverse_dictionary[bi] for bi in batch])
print(' labels:', [reverse_dictionary[li] for li in labels.reshape(8)])
```
## Skip-Gram Algorithm
### Defining Hyperparameters
Here we define several hyperparameters including `batch_size` (amount of samples in a single batch) `embedding_size` (size of embedding vectors) `window_size` (context window size).
```
batch_size = 128 # Data points in a single batch
embedding_size = 128 # Dimension of the embedding vector.
window_size = 4 # How many words to consider left and right.
# We pick a random validation set to sample nearest neighbors
valid_size = 16 # Random set of words to evaluate similarity on.
# We sample valid datapoints randomly from a large window without always being deterministic
valid_window = 50
# When selecting valid examples, we select some of the most frequent words as well as
# some moderately rare words as well
valid_examples = np.array(random.sample(range(valid_window), valid_size))
valid_examples = np.append(valid_examples,random.sample(range(1000, 1000+valid_window), valid_size),axis=0)
num_sampled = 32 # Number of negative examples to sample.
```
### Defining Inputs and Outputs
Here we define placeholders for feeding in training inputs and outputs (each of size `batch_size`) and a constant tensor to contain validation examples.
```
tf.reset_default_graph()
# Training input data (target word IDs).
train_dataset = tf.placeholder(tf.int32, shape=[batch_size])
# Training input label data (context word IDs)
train_labels = tf.placeholder(tf.int32, shape=[batch_size, 1])
# Validation input data, we don't need a placeholder
# as we have already defined the IDs of the words selected
# as validation data
valid_dataset = tf.constant(valid_examples, dtype=tf.int32)
```
### Defining Model Parameters and Other Variables
We now define several TensorFlow variables such as an embedding layer (`embeddings`) and neural network parameters (`softmax_weights` and `softmax_biases`)
```
# Variables
# Embedding layer, contains the word embeddings
embeddings = tf.Variable(tf.random_uniform([vocabulary_size, embedding_size], -1.0, 1.0))
# Softmax Weights and Biases
softmax_weights = tf.Variable(
tf.truncated_normal([vocabulary_size, embedding_size],
stddev=0.5 / math.sqrt(embedding_size))
)
softmax_biases = tf.Variable(tf.random_uniform([vocabulary_size],0.0,0.01))
```
### Defining the Model Computations
We first defing a lookup function to fetch the corresponding embedding vectors for a set of given inputs. With that, we define negative sampling loss function `tf.nn.sampled_softmax_loss` which takes in the embedding vectors and previously defined neural network parameters.
```
# Model.
# Look up embeddings for a batch of inputs.
embed = tf.nn.embedding_lookup(embeddings, train_dataset)
# Compute the softmax loss, using a sample of the negative labels each time.
loss = tf.reduce_mean(
tf.nn.sampled_softmax_loss(
weights=softmax_weights, biases=softmax_biases, inputs=embed,
labels=train_labels, num_sampled=num_sampled, num_classes=vocabulary_size)
)
```
### Calculating Word Similarities
We calculate the similarity between two given words in terms of the cosine distance. To do this efficiently we use matrix operations to do so, as shown below.
```
# Compute the similarity between minibatch examples and all embeddings.
# We use the cosine distance:
norm = tf.sqrt(tf.reduce_sum(tf.square(embeddings), 1, keepdims=True))
normalized_embeddings = embeddings / norm
valid_embeddings = tf.nn.embedding_lookup(
normalized_embeddings, valid_dataset)
similarity = tf.matmul(valid_embeddings, tf.transpose(normalized_embeddings))
```
### Model Parameter Optimizer
We then define a constant learning rate and an optimizer which uses the Adagrad method. Feel free to experiment with other optimizers listed [here](https://www.tensorflow.org/api_guides/python/train).
```
# Optimizer.
optimizer = tf.train.AdagradOptimizer(1.0).minimize(loss)
```
## Running the Skip-Gram Algorithm
```
num_steps = 100001
skip_losses = []
# ConfigProto is a way of providing various configuration settings
# required to execute the graph
with tf.Session(config=tf.ConfigProto(allow_soft_placement=True)) as session:
# Initialize the variables in the graph
tf.global_variables_initializer().run()
print('Initialized')
average_loss = 0
# Train the Word2vec model for num_step iterations
for step in range(num_steps):
# Generate a single batch of data
batch_data, batch_labels = generate_batch_skip_gram(
batch_size, window_size)
# Populate the feed_dict and run the optimizer (minimize loss)
# and compute the loss
feed_dict = {train_dataset : batch_data, train_labels : batch_labels}
_, l = session.run([optimizer, loss], feed_dict=feed_dict)
# Update the average loss variable
average_loss += l
if (step+1) % 2000 == 0:
if step > 0:
average_loss = average_loss / 2000
skip_losses.append(average_loss)
# The average loss is an estimate of the loss over the last 2000 batches.
print('Average loss at step %d: %f' % (step+1, average_loss))
average_loss = 0
# Evaluating validation set word similarities
if (step+1) % 10000 == 0:
sim = similarity.eval()
# Here we compute the top_k closest words for a given validation word
# in terms of the cosine distance
# We do this for all the words in the validation set
# Note: This is an expensive step
for i in range(valid_size):
valid_word = reverse_dictionary[valid_examples[i]]
top_k = 8 # number of nearest neighbors
nearest = (-sim[i, :]).argsort()[1:top_k+1]
log = 'Nearest to %s:' % valid_word
for k in range(top_k):
close_word = reverse_dictionary[nearest[k]]
log = '%s %s,' % (log, close_word)
print(log)
skip_gram_final_embeddings = normalized_embeddings.eval()
# We will save the word vectors learned and the loss over time
# as this information is required later for comparisons
np.save('skip_embeddings',skip_gram_final_embeddings)
with open('skip_losses.csv', 'wt') as f:
writer = csv.writer(f, delimiter=',')
writer.writerow(skip_losses)
```
## Visulizing the Learnings of the Skip-Gram Algorithm
### Finding Only the Words Clustered Together Instead of Sparsely Distributed Words
```
def find_clustered_embeddings(embeddings,distance_threshold,sample_threshold):
'''
Find only the closely clustered embeddings.
This gets rid of more sparsly distributed word embeddings and make the visualization clearer
This is useful for t-SNE visualization
distance_threshold: maximum distance between two points to qualify as neighbors
sample_threshold: number of neighbors required to be considered a cluster
'''
# calculate cosine similarity
cosine_sim = np.dot(embeddings,np.transpose(embeddings))
norm = np.dot(np.sum(embeddings**2,axis=1).reshape(-1,1),np.sum(np.transpose(embeddings)**2,axis=0).reshape(1,-1))
assert cosine_sim.shape == norm.shape
cosine_sim /= norm
# make all the diagonal entries zero otherwise this will be picked as highest
np.fill_diagonal(cosine_sim, -1.0)
argmax_cos_sim = np.argmax(cosine_sim, axis=1)
mod_cos_sim = cosine_sim
# find the maximums in a loop to count if there are more than n items above threshold
for _ in range(sample_threshold-1):
argmax_cos_sim = np.argmax(cosine_sim, axis=1)
mod_cos_sim[np.arange(mod_cos_sim.shape[0]),argmax_cos_sim] = -1
max_cosine_sim = np.max(mod_cos_sim,axis=1)
return np.where(max_cosine_sim>distance_threshold)[0]
```
### Computing the t-SNE Visualization of Word Embeddings Using Scikit-Learn
```
num_points = 1000 # we will use a large sample space to build the T-SNE manifold and then prune it using cosine similarity
tsne = TSNE(perplexity=30, n_components=2, init='pca', n_iter=5000)
print('Fitting embeddings to T-SNE. This can take some time ...')
# get the T-SNE manifold
selected_embeddings = skip_gram_final_embeddings[:num_points, :]
two_d_embeddings = tsne.fit_transform(selected_embeddings)
print('Pruning the T-SNE embeddings')
# prune the embeddings by getting ones only more than n-many sample above the similarity threshold
# this unclutters the visualization
selected_ids = find_clustered_embeddings(selected_embeddings,.25,10)
two_d_embeddings = two_d_embeddings[selected_ids,:]
print('Out of ',num_points,' samples, ', selected_ids.shape[0],' samples were selected by pruning')
```
### Plotting the t-SNE Results with Matplotlib
```
def plot(embeddings, labels):
n_clusters = 20 # number of clusters
# automatically build a discrete set of colors, each for cluster
label_colors = [pylab.cm.spectral(float(i) /n_clusters) for i in range(n_clusters)]
assert embeddings.shape[0] >= len(labels), 'More labels than embeddings'
# Define K-Means
kmeans = KMeans(n_clusters=n_clusters, init='k-means++', random_state=0).fit(embeddings)
kmeans_labels = kmeans.labels_
pylab.figure(figsize=(15,15)) # in inches
# plot all the embeddings and their corresponding words
for i, (label,klabel) in enumerate(zip(labels,kmeans_labels)):
x, y = embeddings[i,:]
pylab.scatter(x, y, c=label_colors[klabel])
pylab.annotate(label, xy=(x, y), xytext=(5, 2), textcoords='offset points',
ha='right', va='bottom',fontsize=10)
# use for saving the figure if needed
#pylab.savefig('word_embeddings.png')
pylab.show()
words = [reverse_dictionary[i] for i in selected_ids]
plot(two_d_embeddings, words)
```
# CBOW Algorithm
## Changing the data generation process
We need to define a new data generator for CBOW. Shape of the new input array is (batch_size, context_window*2). That is, a batch in CBOW captures all the words in the context of a given word.
```
data_index = 0
def generate_batch_cbow(batch_size, window_size):
# window_size is the amount of words we're looking at from each side of a given word
# creates a single batch
# data_index is updated by 1 everytime we read a set of data point
global data_index
# span defines the total window size, where
# data we consider at an instance looks as follows.
# [ skip_window target skip_window ]
# e.g if skip_window = 2 then span = 5
span = 2 * window_size + 1 # [ skip_window target skip_window ]
# two numpy arras to hold target words (batch)
# and context words (labels)
# Note that batch has span-1=2*window_size columns
batch = np.ndarray(shape=(batch_size,span-1), dtype=np.int32)
labels = np.ndarray(shape=(batch_size, 1), dtype=np.int32)
# The buffer holds the data contained within the span
buffer = collections.deque(maxlen=span)
# Fill the buffer and update the data_index
for _ in range(span):
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
# Here we do the batch reading
# We iterate through each batch index
# For each batch index, we iterate through span elements
# to fill in the columns of batch array
for i in range(batch_size):
target = window_size # target label at the center of the buffer
target_to_avoid = [ window_size ] # we only need to know the words around a given word, not the word itself
# add selected target to avoid_list for next time
col_idx = 0
for j in range(span):
# ignore the target word when creating the batch
if j==span//2:
continue
batch[i,col_idx] = buffer[j]
col_idx += 1
labels[i, 0] = buffer[target]
# Everytime we read a data point,
# we need to move the span by 1
# to create a fresh new span
buffer.append(data[data_index])
data_index = (data_index + 1) % len(data)
return batch, labels
for window_size in [1,2]:
data_index = 0
batch, labels = generate_batch_cbow(batch_size=8, window_size=window_size)
print('\nwith window_size = %d:' % (window_size))
print(' batch:', [[reverse_dictionary[bii] for bii in bi] for bi in batch])
print(' labels:', [reverse_dictionary[li] for li in labels.reshape(8)])
```
### Defining Hyperparameters
Here we define several hyperparameters including `batch_size` (amount of samples in a single batch) `embedding_size` (size of embedding vectors) `window_size` (context window size).
```
batch_size = 128 # Data points in a single batch
embedding_size = 128 # Dimension of the embedding vector.
# How many words to consider left and right.
# Skip gram by design does not require to have all the context words in a given step
# However, for CBOW that's a requirement, so we limit the window size
window_size = 2
# We pick a random validation set to sample nearest neighbors
valid_size = 16 # Random set of words to evaluate similarity on.
# We sample valid datapoints randomly from a large window without always being deterministic
valid_window = 50
# When selecting valid examples, we select some of the most frequent words as well as
# some moderately rare words as well
valid_examples = np.array(random.sample(range(valid_window), valid_size))
valid_examples = np.append(valid_examples,random.sample(range(1000, 1000+valid_window), valid_size),axis=0)
num_sampled = 32 # Number of negative examples to sample.
```
### Defining Inputs and Outputs
Here we define placeholders for feeding in training inputs and outputs (each of size `batch_size`) and a constant tensor to contain validation examples.
```
tf.reset_default_graph()
# Training input data (target word IDs). Note that it has 2*window_size columns
train_dataset = tf.placeholder(tf.int32, shape=[batch_size,2*window_size])
# Training input label data (context word IDs)
train_labels = tf.placeholder(tf.int32, shape=[batch_size, 1])
# Validation input data, we don't need a placeholder
# as we have already defined the IDs of the words selected
# as validation data
valid_dataset = tf.constant(valid_examples, dtype=tf.int32)
```
### Defining Model Parameters and Other Variables
We now define several TensorFlow variables such as an embedding layer (`embeddings`) and neural network parameters (`softmax_weights` and `softmax_biases`)
```
# Variables.
# Embedding layer, contains the word embeddings
embeddings = tf.Variable(tf.random_uniform([vocabulary_size, embedding_size], -1.0, 1.0,dtype=tf.float32))
# Softmax Weights and Biases
softmax_weights = tf.Variable(tf.truncated_normal([vocabulary_size, embedding_size],
stddev=0.5 / math.sqrt(embedding_size),dtype=tf.float32))
softmax_biases = tf.Variable(tf.random_uniform([vocabulary_size],0.0,0.01))
```
### Defining the Model Computations
We first defing a lookup function to fetch the corresponding embedding vectors for a set of given inputs. Concretely, we define 2$\times$`window_size` embedding lookups. We then concatenate all these looked up embedding vectors to form a matrix of size `[batch_size, embedding_size, 2*window_size]`. Thereafter, we average these embedding lookups to produce an average embeddings of size `[batch_size, embedding_size]`. With that, we define negative sampling loss function `tf.nn.sampled_softmax_loss` which takes in the embedding vectors and previously defined neural network parameters.
```
# Model.
# Look up embeddings for a batch of inputs.
# Here we do embedding lookups for each column in the input placeholder
# and then average them to produce an embedding_size word vector
stacked_embedings = None
print('Defining %d embedding lookups representing each word in the context'%(2*window_size))
for i in range(2*window_size):
embedding_i = tf.nn.embedding_lookup(embeddings, train_dataset[:,i])
x_size,y_size = embedding_i.get_shape().as_list()
if stacked_embedings is None:
stacked_embedings = tf.reshape(embedding_i,[x_size,y_size,1])
else:
stacked_embedings = tf.concat(axis=2,values=[stacked_embedings,tf.reshape(embedding_i,[x_size,y_size,1])])
assert stacked_embedings.get_shape().as_list()[2]==2*window_size
print("Stacked embedding size: %s"%stacked_embedings.get_shape().as_list())
mean_embeddings = tf.reduce_mean(stacked_embedings,2,keepdims=False)
print("Reduced mean embedding size: %s"%mean_embeddings.get_shape().as_list())
# Compute the softmax loss, using a sample of the negative labels each time.
# inputs are embeddings of the train words
# with this loss we optimize weights, biases, embeddings
loss = tf.reduce_mean(
tf.nn.sampled_softmax_loss(weights=softmax_weights, biases=softmax_biases, inputs=mean_embeddings,
labels=train_labels, num_sampled=num_sampled, num_classes=vocabulary_size))
```
### Model Parameter Optimizer
We then define a learning rate as a constant and an optimizer which uses the Adagrad method. Feel free to experiment with other optimizers listed [here](https://www.tensorflow.org/api_guides/python/train).
```
# Optimizer.
optimizer = tf.train.AdagradOptimizer(1.0).minimize(loss)
```
### Calculating Word Similarities
We calculate the similarity between two given words in terms of the cosine distance. To do this efficiently we use matrix operations to do so, as shown below.
```
# Compute the similarity between minibatch examples and all embeddings.
# We use the cosine distance:
norm = tf.sqrt(tf.reduce_sum(tf.square(embeddings), 1, keepdims=True))
normalized_embeddings = embeddings / norm
valid_embeddings = tf.nn.embedding_lookup(normalized_embeddings, valid_dataset)
similarity = tf.matmul(valid_embeddings, tf.transpose(normalized_embeddings))
```
## Running the CBOW Algorithm
```
num_steps = 100001
cbow_losses = []
# ConfigProto is a way of providing various configuration settings
# required to execute the graph
with tf.Session(config=tf.ConfigProto(allow_soft_placement=True)) as session:
# Initialize the variables in the graph
tf.global_variables_initializer().run()
print('Initialized')
average_loss = 0
# Train the Word2vec model for num_step iterations
for step in range(num_steps):
# Generate a single batch of data
batch_data, batch_labels = generate_batch_cbow(batch_size, window_size)
# Populate the feed_dict and run the optimizer (minimize loss)
# and compute the loss
feed_dict = {train_dataset : batch_data, train_labels : batch_labels}
_, l = session.run([optimizer, loss], feed_dict=feed_dict)
# Update the average loss variable
average_loss += l
if (step+1) % 2000 == 0:
if step > 0:
average_loss = average_loss / 2000
# The average loss is an estimate of the loss over the last 2000 batches.
cbow_losses.append(average_loss)
print('Average loss at step %d: %f' % (step+1, average_loss))
average_loss = 0
# Evaluating validation set word similarities
if (step+1) % 10000 == 0:
sim = similarity.eval()
# Here we compute the top_k closest words for a given validation word
# in terms of the cosine distance
# We do this for all the words in the validation set
# Note: This is an expensive step
for i in range(valid_size):
valid_word = reverse_dictionary[valid_examples[i]]
top_k = 8 # number of nearest neighbors
nearest = (-sim[i, :]).argsort()[1:top_k+1]
log = 'Nearest to %s:' % valid_word
for k in range(top_k):
close_word = reverse_dictionary[nearest[k]]
log = '%s %s,' % (log, close_word)
print(log)
cbow_final_embeddings = normalized_embeddings.eval()
np.save('cbow_embeddings',cbow_final_embeddings)
with open('cbow_losses.csv', 'wt') as f:
writer = csv.writer(f, delimiter=',')
writer.writerow(cbow_losses)
```
## Final Remarks
I'm grateful to the Google's free deep learning course at Udacity, as some of the helper functions were adopted from code files provided in their [Github repo](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/examples/udacity/5_word2vec.ipynb)
| github_jupyter |
# Why You Should Hedge Beta and Sector Exposures (Part I)
by Jonathan Larkin and Maxwell Margenot
Part of the Quantopian Lecture Series:
* [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
* [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
Notebook released under the Creative Commons Attribution 4.0 License.
---
Whenever we have a trading strategy of any sort, we need to be considering the impact of systematic risk. There needs to be some risk involved in a strategy in order for there to be a return above the risk-free rate, but systematic risk poisons the well, so to speak. By its nature, systematic risk provides a commonality between the many securities in the market that cannot be diversified away. As such, we need to construct a hedge to get rid of it.
```
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
from sklearn.covariance import LedoitWolf
import seaborn as sns
import statsmodels.api as sm
```
# The Fundamental Law of Asset Management
The primary driver of the value of any strategy is whether or not it provides a compelling risk-adjusted return, i.e., the Sharpe Ratio. As expressed in [The Foundation of Algo Success](https://blog.quantopian.com/the-foundation-of-algo-success/) and "The Fundamental Law of Active Management", by Richard Grinold, Sharpe Ratio can be decomposed into two components, skill and breadth, as:
$$IR = IC \sqrt{BR}$$
Technically, this is the definition of the Information Ratio (IR), but for our purposes it is equivalent to the Sharpe Ratio. The IR is the ratio of the excess return of a portfolio over its benchmark per unit active risk, i.e., the excess return of a long-only portfolio less its benchmark per unit tracking error. In the time of Grinold’s publication, however, long/short investing was a rarity. Today, in the world of hedge funds and long/short investing, there is no benchmark. We seek absolute returns so, in this case, the IR is equivalent to the Sharpe ratio.
In this equation, skill is measured by IC (Information Coefficient), calculated with [Alphalens](https://github.com/quantopian/alphalens). The IC is essentially the Spearman rank correlation, used to correlate your prediction and its realization. Breadth is measured as the number of **independent** bets in the period. The takeaway from this "law" is that, with any strategy, we need to:
1. Bet well (high IC),
2. Bet often (high number of bets), *and*
3. **Make independent bets**
If the bets are completely independent, then breadth is the total number of bets we have made for every individual asset, the number of assets times the number of periods. If the bets are not independent then the **effective breadth** can be much much less than the number of assets. Let's see precisely what beta exposure and sector exposure do to **effective breadth**.
<div class="alert alert-warning">
<b>TL;DR:</b> Beta exposure and sector exposure lead to a significant increase in correlation among bets. Portfolios with beta and sector bets have very low effective breadth. In order to have high Sharpe then, these portfolios must have very high IC. It is easier to increase effective breadth by hedging beta and sector exposure than it is to increase your IC.
</div>
# Forecasts and Bet Correlation
We define a bet as the forecast of the *residual* of a security return. This forecast can be implicit -- i.e., we buy a stock and thus implicity we forecast that the stock will go up. What though do we mean by *residual*? Without any fancy math, this simply means the return **less a hedge**. Let's work through three examples. We use the Ledoit-Wolf covariance estimator to assess our covariance in all cases. For more information on why we use Ledoit-Wolf instead of typical sample covariance, check out [Estimating Covariance Matrices](https://www.quantopian.com/lectures/estimating-covariance-matrices).
### Example 1: No Hedge!
If we go long on a set of securities, but do not hold any short positions, there is no hedge! So the *residual* is the stock return itself.
$$r_{resid,i} = r_i$$
Let's see what the correlation of our bets are in this case.
```
tickers = ['WFC', 'JPM', 'USB', 'XOM', 'BHI', 'SLB'] # The securities we want to go long on
historical_prices = get_pricing(tickers, start_date='2015-01-01',end_date='2017-02-22') # Obtain prices
rets = historical_prices['close_price'].pct_change().fillna(0) # Calculate returns
lw_cov = LedoitWolf().fit(rets).covariance_ # Calculate Ledoit-Wolf estimator
def extract_corr_from_cov(cov_matrix):
# Linear algebra result:
# https://math.stackexchange.com/questions/186959/correlation-matrix-from-covariance-matrix
d = np.linalg.inv(np.diag(np.sqrt(np.diag(cov_matrix))))
corr = d.dot(cov_matrix).dot(d)
return corr
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
# Plot prices
left = historical_prices['close_price'].plot(ax=ax1)
# Plot covariance as a heat map
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=tickers, yticklabels=tickers)
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print 'Average pairwise correlation: %.4f' % average_corr
```
The result here is that we have six bets and they are all very highly correlated.
### Example 2: Beta Hedge
In this case, we will assume that each bet is hedged against the market (SPY). In this case, the residual is calculated as:
$$ r_{resid,i} = r_i - \beta_i r_i $$
where $\beta_i$ is the beta to the market of security $i$ calculated with the [CAPM](https://www.quantopian.com/lectures/the-capital-asset-pricing-model-and-arbitrage-pricing-theory) and $r_i$ is the return of security $i$.
```
tickers = ['WFC', 'JPM', 'USB', 'SPY', 'XOM', 'BHI', 'SLB' ] # The securities we want to go long on plus SPY
historical_prices = get_pricing(tickers, start_date='2015-01-01',end_date='2017-02-22') # Obtain prices
rets = historical_prices['close_price'].pct_change().fillna(0) # Calculate returns
market = rets[symbols(['SPY'])]
stock_rets = rets.drop(symbols(['SPY']), axis=1)
residuals = stock_rets.copy()*0
for stock in stock_rets.columns:
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
residuals[stock] = results.resid
lw_cov = LedoitWolf().fit(residuals).covariance_ # Calculate Ledoit-Wolf Estimator
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
left = (1+residuals).cumprod().plot(ax=ax1)
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=tickers, yticklabels=tickers)
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print 'Average pairwise correlation: %.4f' % average_corr
```
The beta hedge has brought down the average correlation significanty. Theoretically, this should improve our breadth. It is obvious that we are left with two highly correlated clusters however. Let's see what happens when we hedge the sector risk.
### Example 3: Sector Hedge
The sector return and the market return are themselves highly correlated. As such, you cannot do a multivariate regression due to multicollinearity, a classic [violation of regression assumptions](https://www.quantopian.com/lectures/violations-of-regression-models). To hedge against both the market and a given security's sector, you first estimate the market beta residuals and then calculate the sector beta on *those* residuals.
$$
r_{resid,i} = r_i - \beta_i r_i \\
r_{resid_{SECTOR},i}= r_{resid,i} - \beta_{SECTOR,i}r_{resid,i}
$$
Here, $r_{resid, i}$ is the residual between the security return and a market beta hedge and $r_{resid_{SECTOR}, i}$ is the residual between *that* residual and a hedge of that residual against the relevant sector.
```
tickers = ['WFC', 'JPM', 'USB', 'XLF', 'SPY', 'XOM', 'BHI', 'SLB', 'XLE']
historical_prices = get_pricing(tickers, start_date='2015-01-01',end_date='2017-02-22')
rets = historical_prices['close_price'].pct_change().fillna(0)
# Get market hedge ticker
mkt = symbols(['SPY'])
# Get sector hedge tickers
sector_1_hedge = symbols(['XLF'])
sector_2_hedge = symbols(['XLE'])
# Identify securities for each sector
sector_1_stocks = symbols(['WFC', 'JPM', 'USB'])
sector_2_stocks = symbols(['XOM', 'BHI', 'SLB'])
market_rets = rets[mkt]
sector_1_rets = rets[sector_1_hedge]
sector_2_rets = rets[sector_2_hedge]
stock_rets = rets.drop(symbols(['XLF', 'SPY', 'XLE']), axis=1)
residuals_market = stock_rets.copy()*0
residuals = stock_rets.copy()*0
# Calculate market beta of sector 1 benchmark
model = sm.OLS(sector_1_rets.values, market.values)
results = model.fit()
sector_1_excess = results.resid
# Calculate market beta of sector 2 benchmark
model = sm.OLS(sector_2_rets.values, market.values)
results = model.fit()
sector_2_excess = results.resid
for stock in sector_1_stocks:
# Calculate market betas for sector 1 stocks
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
# Calculate residual of security + market hedge
residuals_market[stock] = results.resid
# Calculate sector beta for previous residuals
model = sm.OLS(residuals_market[stock], sector_1_excess)
results = model.fit()
# Get final residual
residuals[stock] = results.resid
for stock in sector_2_stocks:
# Calculate market betas for sector 2 stocks
model = sm.OLS(stock_rets[stock], market.values)
results = model.fit()
# Calculate residual of security + market hedge
residuals_market[stock] = results.resid
# Calculate sector beta for previous residuals
model = sm.OLS(residuals_market[stock], sector_2_excess)
results = model.fit()
# Get final residual
residuals[stock] = results.resid
# Get covariance of residuals
lw_cov = LedoitWolf().fit(residuals).covariance_
fig, (ax1, ax2) = plt.subplots(ncols=2)
fig.tight_layout()
corr = extract_corr_from_cov(lw_cov)
left = (1+residuals).cumprod().plot(ax=ax1)
right = sns.heatmap(corr, ax=ax2, fmt='d', vmin=-1, vmax=1, xticklabels=tickers, yticklabels=tickers)
average_corr = np.mean(corr[np.triu_indices_from(corr, k=1)])
print 'Average pairwise correlation: %.4f' % average_corr
```
There we go! The sector hedge brought down the correlation between our bets to close to zero.
## Calculating Effective Breadth
This section is based on "How to calculate breadth: An evolution of the fundamental law of active portfolio management", by David Buckle; Vol. 4, 6, 393-405, 2003, _Journal of Asset Management_. Buckle derives the "semi-generalised fundamental law of active management" under several weak assumptions. The key result of this paper (for us) is a closed-form calculaiton of effective breadth as a function of the correlation between bets. Buckle shows that breadth, $BR$, can be modeled as
$$BR = \frac{N}{1 + \rho(N -1)}$$
where N is the number of stocks in the portfolio and $\rho$ is the assumed single correlation of the expected variation around the forecast.
```
def buckle_BR_const(N, rho):
return N/(1 + rho*(N - 1))
corr = np.linspace(start=0, stop=1.0, num=500)
plt.plot(corr, buckle_BR_const(6, corr))
plt.title('Effective Breadth as a function of Forecast Correlation (6 Stocks)')
plt.ylabel('Effective Breadth (Number of Bets)')
plt.xlabel('Forecast Correlation');
```
Here we see that in the case of the long-only portfolio, where the average correlation is 0.56, we are *effectively making only approximately 2 bets*. When we hedge beta, with a resulting average correlation of 0.22, things get a little better, *three effective bets*. When we add the sector hedge, we get close to zero correlation, and in this case the number of bets equals the number of assets, 6.
**More independent bets with the same IC leads to higher Sharpe ratio.**
## Using this in Practice
Trading costs money due to market impact and commissions. As such, the post hoc implementation of a hedge is almost always suboptimal. In that case, you are trading purely to hedge risk. It is preferable to think about your sector and market exposure *throughout the model development process*. Sector and market risk is naturally hedged in a pairs-style strategy; in a cross-sectional strategy, consider de-meaning the alpha vector by the sector average; with an event-driven strategy, consider adding additional alphas so you can find offsetting bets in the same sector. As a last resort, hedge with a well chosen sector ETF.
*This presentation is for informational purposes only and does not constitute an offer to sell, a solicitation to buy, or a recommendation for any security; nor does it constitute an offer to provide investment advisory or other services by Quantopian, Inc. ("Quantopian"). Nothing contained herein constitutes investment advice or offers any opinion with respect to the suitability of any security, and any views expressed herein should not be taken as advice to buy, sell, or hold any security or as an endorsement of any security or company. In preparing the information contained herein, Quantopian, Inc. has not taken into account the investment needs, objectives, and financial circumstances of any particular investor. Any views expressed and data illustrated herein were prepared based upon information, believed to be reliable, available to Quantopian, Inc. at the time of publication. Quantopian makes no guarantees as to their accuracy or completeness. All information is subject to change and may quickly become unreliable for various reasons, including changes in market conditions or economic circumstances.*
| github_jupyter |
<figure>
<IMG SRC="https://raw.githubusercontent.com/mbakker7/exploratory_computing_with_python/master/tudelft_logo.png" WIDTH=250 ALIGN="right">
</figure>
# Exploratory Computing with Python
*Developed by Mark Bakker*
## Notebook 11: Distribution of the mean, hypothesis tests, and the central limit theorem
In this notebook we first investigate the distribution of the mean of a dataset, we simulate several hypothesis tests, and finish with exploring the central limit theorem.
```
import numpy as np
import matplotlib.pyplot as plt
import numpy.random as rnd
%matplotlib inline
```
Consider a dataset of 100 points. The data are drawn from a normal distribution with mean 4 and standard deviation 2. As we noticed before, the sample mean of the 100 data points almost always differs from 4. And every time we generate a new set of 100 points, the mean will be somewhat different.
```
for i in range(5):
a = 2 * rnd.standard_normal(100) + 4
print('mean a:', np.mean(a))
```
In fact, the mean of the dataset itself can be considered as a random variable with a distribution of its own.
### Sample standard deviation
The sample standard deviation $s_n$ of a dataset of $n$ values is defined as
$s_n = \sqrt{ \frac{1}{n-1} \sum_{i=1}^n (x_i - \overline{x}_n)^2 }$
and can be computed with the `std` function of the `numpy` package. By default, the `std` function devides the sum by $n$ rather than by $n-1$. To divide by $n-1$, as we want for an unbiased estimate of the standard deviation, specify the keyword argument `ddof=1` in the `np.std` function.
### Exercise 1. <a name="back1"></a>Histogram of the means of datasets with 100 values
Generate 1000 datasets each with 100 values drawn from a normal distribution with mean 4 and standard deviation 2; use a seed of 22. Compute the mean of each dataset and store them in an array of length 1000. Compute the mean of the means and the standard deviation of the means, and print them to the screen. Draw a boxplot of the means. In a separate figure, draw a histogram of the means. Make sure the vertical axis of the boxplot and the horizontal axis of the histogram extend from 3 to 5.
<a href="#ex1answer">Answers to Exercise 1</a>
### Exercise 2. <a name="back2"></a>Histogram of the means of datasets with 1000 values
Repeat exercise 1 but now generate 1000 datasets each with 1000 values (rather than 100 values) drawn from the same normal distribution with mean 4 and standard deviation 2, and again with a seed of 22. Make sure the vertical axis of the boxplot and the horizontal axis of the histogram extend from 3 to 5, so that the graphs can be compared to the graphs you created in the previous exercise. Is the spread of the mean much smaller now as compared to the datasets consisting of only 100 values?
<a href="#ex2answer">Answers to Exercise 2</a>
### Sample standard deviation of the sample mean
The histogram of the means looks like the bell-shaped curve of a Normal distribution, but you may recall that it is actually a Student's $t$-distribution, also simply called a $t$-distribution. A $t$-distribution arises when estimating the mean of a normally distributed variable in situations where the sample size is relatively small and the standard deviation is unknown (as it pretty much always is in practice) and needs to be estimated from the data.
The sample mean of a dataset of $n$ values is commonly written as $\overline{x}_n$, while the sample standard deviation is written as $s_n$ (as defined above). Here, we are computing the sample standard deviation of the sample means, which we write as $\hat{s}_n$ for a dataset of size $n$. Theoretically, the value of the sample standard deviation of the sample mean $\hat{s}_n$ is related to the sample standard deviation as (see [here](http://en.wikipedia.org/wiki/Standard_deviation#Standard_deviation_of_the_mean))
$\hat{s}_n = s_n / \sqrt{n}$
### Percentiles of $t$-distribution
You may recall that the 90% interval around the mean for a Normally distributed variable runs from $\mu-1.64\sigma$ to $\mu+1.64\sigma$. In other words, 5% of the data is expected to lie below $\mu-1.64\sigma$ and 5% of the data is expected to lie above $\mu+1.64\sigma$. What now if you forgot it is $1.64\sigma$ to the left and right of the mean? Or what if you want to know the value for some other percentile. You may look that up in a table in a Statistics book (or on the web), or use the percent point function `ppf`, which is part of any statistical distribution function defined in the `scipy.stats` package. The `ppf` function is the inverse of the cumulative distribution function. For example, `ppf(0.05)` returns the value of the data such that the cumulative distribution function is equal to 0.05 at the returned value. To find the 5% and 95% values, type (recall that by default the `norm` distribution has mean zero and standard deviation 1; you can specify different values with the `loc` and `scale` keyword arguments, respectively).
```
from scipy.stats import norm
xvalue_05 = norm.ppf(0.05)
xvalue_95 = norm.ppf(0.95)
print('5% limit:', xvalue_05)
print('95% limit:', xvalue_95)
print('check if it works for 5%:', norm.cdf(xvalue_05))
print('check if it works for 95%:', norm.cdf(xvalue_95))
# Next, specify a mean and standard deviation
xvalue_05_musig = norm.ppf(0.05, loc=20, scale=10) # mu = 20, sigma = 10
print('5% limit with mu=20, sig=10:', xvalue_05_musig)
print('check:', norm.cdf(xvalue_05_musig, loc=20, scale=10))
```
A similar function exists for the $t$ distribution. The $t$-distribution takes one additional argument: the number of degrees of freedom, which is equal to the number of data points minus 1. For example, consider a sample with 40 data points, a sample mean of 20, and a sample standard deviation of the mean of 2, then the 5 and 95 percentiles are
```
from scipy.stats import t
xvalue_05 = t.ppf(0.05, 39, loc=20, scale=2)
xvalue_95 = t.ppf(0.95, 39, loc=20, scale=2)
print('5% limit: ',xvalue_05)
print('95% limit: ',xvalue_95)
print('check if it works for 5%:', t.cdf(xvalue_05, 39, loc=20, scale=2))
print('check if it works for 95%:', t.cdf(xvalue_95, 39, loc=20, scale=2))
```
### Exercise 3. <a name="back3"></a>Count the number of means outside 95 percentile
Go back to Exercise 1. Generate 1000 datasets each with 100 values drawn from a normal distribution with mean 4 and standard deviation 2. For each dataset, evaluate whether the sample mean is within the 95 percentile of the $t$-distribution around the true mean of 4 (the standard deviation of the sample mean is different every time, of course). Count how many times the sample mean is outside the 95 percentile around the true mean of the $t$ distribution. If the theory is correct, it should, of course, be the case for about 5% of the datasets. Try five different seeds and report the percentage of means in the dataset that is outside the 95 percentile around the true mean.
<a href="#ex3answer">Answers to Exercise 3</a>
### Exercise 4. <a name="back4"></a>$t$ test on dataset of 20 values
Generate 20 datapoints from a Normal distribution with mean 39 and standard deviation 4. Use a seed of 2. Compute and report the sample mean and sample standard deviation of the dataset and the sample standard deviation of the sample mean.
If you computed it correctly, the mean of the 20 data points generated above is 38.16. Somebody now claims that the 20 datapoints are taken from a distribution with a mean of 40. You are asked to decide wether the true underlying mean could indeed be 40. In statistical terms, you are asked to perform a Hypothesis test, testing the null hypothesis that the mean is 40 against the alternative hypothesis that the mean is not 40 at significance level 5%. Hence, you are asked to do a two-sided $t$-test. All you can do in Hypothesis testing it trying to reject the null hypothesis, so let's try that. Most statistics books give a cookbook recipe for performing a $t$-test. Here we will visualize the $t$-test. We reject the null hypothesis if the sample mean is outside the 95% interval around the mean of the corresponding $t$-distribution. If the mean is inside the 95% interval we can only conclude that there is not enough evidence to reject the null hypothesis. Draw the probability density function of a $t$-distribution with mean 40 and standard deviation equal to the sample standard deviation of the sample mean you computed above. Draw red vertical lines indicating the left and right limits of the 95% interval around the mean. Draw a heavy black vertical line at the position of the sample mean you computed above. Decide whether you can reject the null hypothesis that the mean is 40 and add that as a title to the figure.
<a href="#ex4answer">Answers to Exercise 4</a>
### Exercise 5. <a name="back5"></a>Hypothesis tests on Wooden beam data
Load the data set of experiments on wooden beams stored in the file `douglas_data.csv`. First, consider the first 20 measurements of the bending strength. Compute the sample mean and the standard deviation of the sample mean. The manufacturer claims that the mean bending strength is only 50 Pa. Perform a $t$-test (significance level 5%) with null hypothesis that the mean is indeed 50 Pa and alternative hypothesis that the mean is not 50 Pa using the approach applied in Exercise 4.
Repeat the $t$-test above but now with all the measurements of the bending strength. Do you reach the same conclusion?
<a href="#ex5answer">Answers to Exercise 5</a>
### Central limit theorem
So far we looked at the distribution of the sample mean of a dataset while we knew that the data was taken from a normal distribution (except for the wooden beam data, but that looked very much like a Normal distribution). Such a sample mean has a Student $t$-distribtion, which approaches the Normal distribution when the dataset is large. Actually, 100 datapoints is already enough to approach the Normal distribution fairly closely. You may check this by comparing, for example, the percent point function `ppf` of a Normal distribution with a $t$-distribution with 99 degrees of freedom, or by simply plotting the pdf of both distributions:
```
print('95 percentile Standard Normal: ',norm.ppf(0.95))
print('95 percentile t-dist with n=99: ',t.ppf(0.95,99))
x = np.linspace(-4,4,100)
y1 = norm.pdf(x)
y2 = t.pdf(x,99)
plt.plot(x,y1,'b',label='Normal')
plt.plot(x,y2,'r',label='t-dist')
plt.legend();
```
The Central limit theorem now states that the distribution of the sample mean approaches the Normal distribution in the limit even if the dataset is drawn from an entirely different distribution! We are going to test this theorem by drawing numbers from a Gamma distribution. The Gamma distribution is a skewed distribution and takes a shape parameter $k$ and a scale parameter $\theta$, and is defined for $x>0$. Details on the Gamma distribution can be found, for example [here](http://en.wikipedia.org/wiki/Gamma_distribution). Let's set the shape parameter equal to 2 and the scale parameter equal to 1 (which happens to be the default). When the scale parameter is equal to 1, the mean is equal to the shape parameter. The pdf of the Gamma distribution for these values is shown below. The mean is indicated with the red vertical line.
```
from scipy.stats import gamma
x = np.linspace(1e-6, 10, 100)
y = gamma.pdf(x, 2, scale=1)
plt.plot(x, y)
plt.axvline(2, color='r');
```
Random numbers may be drawn from any distribution in the `scipy.stats` package with the `rvs` function. Here, we draw 1000 numbers and add the histogram to the previous figure
```
x = np.linspace(1e-6, 10, 100)
y = gamma.pdf(x, 2)
plt.plot(x, y)
plt.axvline(2, color='r')
data = gamma.rvs(2, size=1000)
plt.hist(data, bins=20, normed=True);
```
### Exercise 6. <a name="back6"></a>Explore Central Limit Theorem for Gamma Distribution
Generate $N$ datasets of 20 numbers randomly drawn from a Gamma distribution with shape parameter equal to 2 and scale equal to 1. Draw a histogram of the means of the $N$ datasets using 20 bins. On the same graph, draw the pdf of the Normal distribution using the mean of means and sample standard deviation of the means; choose the limits of the $x$-axis between 0 and 4. Make 3 graphs, for $N=100,1000,10000$ and notice that the distribution starts to approach a Normal distribution. Add a title to each graph stating the number of datasets.
<a href="#ex6answer">Answers to Exercise 6</a>
### Answers to the exercises
<a name="ex1answer">Answers to Exercise 1</a>
```
rnd.seed(22)
mean_of_data = np.mean(2 * rnd.standard_normal((1000, 100)) + 4, 1)
print('The mean of the means is:', np.mean(mean_of_data))
print('The standard deviation of the means is:', np.std(mean_of_data, ddof=1))
plt.figure()
plt.boxplot(mean_of_data)
plt.ylim(3, 5)
plt.figure()
plt.hist(mean_of_data, normed=True)
plt.xlim(3,5)
```
<a href="#back1">Back to Exercise 1</a>
<a name="ex2answer">Answers to Exercise 2</a>
```
rnd.seed(22)
mean_of_data = np.mean(2 * rnd.standard_normal((1000, 1000)) + 4, 1)
print('The mean of the means is:', np.mean(mean_of_data))
print('The standard deviation of the means is:', np.std(mean_of_data, ddof=1))
plt.figure()
plt.boxplot(mean_of_data)
plt.ylim(3,5)
plt.figure()
plt.hist(mean_of_data)
plt.xlim(3, 5)
```
<a href="#back2">Back to Exercise 2</a>
<a name="ex3answer">Answers to Exercise 3</a>
```
from scipy.stats import t
for s in [22, 32, 42, 52, 62]:
rnd.seed(s)
data = 2.0 * rnd.standard_normal((1000, 100)) + 4.0
mean = np.mean(data, 1)
sighat = np.std(data, axis=1, ddof=1) / np.sqrt(100)
count = 0
for i in range(1000):
low = t.ppf(0.025, 99, loc=4, scale=sighat[i])
high = t.ppf(0.975, 99, loc=4, scale=sighat[i])
if (mean[i] < low) or (mean[i] > high): count += 1
print('percentage of datasets where sample mean is outside 95 percentile:', count * 100 / 1000)
```
<a href="#back3">Back to Exercise 3</a>
<a name="ex4answer">Answers to Exercise 4</a>
```
rnd.seed(2)
data = 4 * rnd.standard_normal(20) + 39
mu = np.mean(data)
sig = np.std(data, ddof=1)
sighat = np.std(data, ddof=1) / np.sqrt(20)
print('mean of the data:', mu)
print('std of the data:', sig)
print('std of the mean:', sighat)
x = np.linspace(37, 43, 100)
y = t.pdf(x, 19, loc=40, scale=sighat)
plt.plot(x, y)
perc025 = t.ppf(0.025, 19, loc=40, scale=sighat)
perc975 = t.ppf(0.975, 19, loc=40, scale=sighat)
plt.axvline(perc025, color='r')
plt.axvline(perc975, color='r')
plt.axvline(mu, color='k', lw=5)
plt.title('H0 cannot be rejected');
```
<a href="#back4">Back to Exercise 4</a>
<a name="ex5answer">Answers to Exercise 5</a>
```
from pandas import read_csv
w = read_csv('douglas_data.csv', skiprows=[1], skipinitialspace=True)
mu20 = np.mean(w.bstrength[:20])
sig20 = np.std(w.bstrength[:20], ddof=1) / np.sqrt(20)
print('sample mean, standard deviation of sample mean: ', mu20, sig20)
x = np.linspace(30,70,100)
y = t.pdf(x, 19, loc=50, scale=sig20)
plt.plot(x,y)
perc025 = t.ppf(0.025, 19, loc=50, scale=sig20)
perc975 = t.ppf(0.975, 19, loc=50, scale=sig20)
plt.axvline(perc025, color='r')
plt.axvline(perc975, color='r')
plt.axvline(mu20, color='k', lw=4)
plt.title('H0 is rejected: mean is not 50 Pa');
from pandas import read_csv
w = read_csv('douglas_data.csv', skiprows=[1], skipinitialspace=True)
N = len(w.bstrength)
mu = np.mean(w.bstrength)
sig = np.std(w.bstrength, ddof=1) / np.sqrt(N)
print('sample mean, standard deviation of sample mean: ', mu, sig)
x = np.linspace(30, 70, 100)
y = t.pdf(x, N - 1, loc=50, scale=sig)
plt.plot(x, y)
perc025 = t.ppf(0.025, N - 1, loc=50, scale=sig)
perc975 = t.ppf(0.975, N - 1, loc=50, scale=sig)
plt.axvline(perc025, color='r')
plt.axvline(perc975, color='r')
plt.axvline(mu, color='k', lw=4)
plt.title('Not enough evidence to reject H0: mean may very well be 50');
```
<a href="#back5">Back to Exercise 5</a>
<a name="ex6answer">Answers to Exercise 6</a>
```
from scipy.stats import norm, gamma
for N in [100, 1000, 10000]:
data = gamma.rvs(2, size=(N, 20))
mean_of_data = np.mean(data, 1)
mu = np.mean(mean_of_data)
sig = np.std(mean_of_data, ddof=1)
plt.figure()
plt.hist(mean_of_data, bins=20, normed=True)
x = np.linspace(0, 4, 100)
y = norm.pdf(x, loc=mu, scale=sig)
plt.plot(x, y, 'r')
plt.title('N=' + str(N))
```
<a href="#back6">Back to Exercise 6</a>
| github_jupyter |
```
from os import path, listdir
import re
from types import SimpleNamespace as simplenamespace
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
%matplotlib inline
!ls logs
LOG_DIR = path.join(path.abspath("."), "logs")
n_points = [1000, 5000, 10_000, 100_000, 250_000, 500_000, 750_000, 1_000_000]
def make_regex_func(pattern, fname):
def _wrapper(n_points):
matches = [re.findall(pattern, line) for line in
open(path.join(LOG_DIR, f"{fname}_{n_points}.log"))]
matches = list(filter(len, matches))
matches = list(map(lambda x: float(x[0]), matches))
return np.array(matches)
return _wrapper
parse_opentsne = make_regex_func(r"openTSNE: Full (\d+\.\d+)", "openTSNEapprox")
parse_opentsne_nn = make_regex_func(r"openTSNE: NN search (\d+\.\d+)", "openTSNEapprox")
parse_opentsne_optimization = make_regex_func(r"openTSNE: Optimization (\d+\.\d+)", "openTSNEapprox")
opentsne = simplenamespace(full=parse_opentsne, nn=parse_opentsne_nn, optim=parse_opentsne_optimization)
print(opentsne.full(100000))
parse_opentsne8 = make_regex_func(r"openTSNE: Full (\d+\.\d+)", "openTSNEapprox8core")
parse_opentsne_nn8 = make_regex_func(r"openTSNE: NN search (\d+\.\d+)", "openTSNEapprox8core")
parse_opentsne_optimization8 = make_regex_func(r"openTSNE: Optimization (\d+\.\d+)", "openTSNEapprox8core")
opentsne8 = simplenamespace(full=parse_opentsne8, nn=parse_opentsne_nn8, optim=parse_opentsne_optimization8)
print(opentsne8.full(100000))
parse_fitsne = make_regex_func(r"FIt-SNE: (\d+\.\d+)", "FItSNE")
parse_fitsne_nn = make_regex_func(r"100\% (\d+\.\d+)", "FItSNE")
def parse_fitsne_optimization(n_points):
full_times = parse_fitsne(n_points)
nn_times = parse_fitsne_nn(n_points)
return full_times - nn_times
fitsne = simplenamespace(full=parse_fitsne, nn=parse_fitsne_nn, optim=parse_fitsne_optimization)
print(fitsne.full(100000))
parse_fitsne8 = make_regex_func(r"FIt-SNE: (\d+\.\d+)", "FItSNE8core")
parse_fitsne_nn8 = make_regex_func(r"100\% (\d+\.\d+)", "FItSNE8core")
def parse_fitsne_optimization8(n_points):
full_times = parse_fitsne8(n_points)
nn_times = parse_fitsne_nn8(n_points)
return full_times - nn_times
fitsne8 = simplenamespace(full=parse_fitsne8, nn=parse_fitsne_nn8, optim=parse_fitsne_optimization8)
print(fitsne8.full(1000))
parse_multicore = make_regex_func(r"Multicore t-SNE: (\d+\.\d+)", "MulticoreTSNE")
parse_multicore_nn = make_regex_func(r"Done in (\d+\.\d+) seconds", "MulticoreTSNE")
parse_multicore_optimization = make_regex_func(r"Fitting performed in (\d+\.\d+) seconds", "MulticoreTSNE")
multicore = simplenamespace(full=parse_multicore, nn=parse_multicore_nn, optim=parse_multicore_optimization)
print(multicore.full(100000))
parse_multicore8 = make_regex_func(r"Multicore t-SNE: (\d+\.\d+)", "MulticoreTSNE8core")
parse_multicore_nn8 = make_regex_func(r"Done in (\d+\.\d+) seconds", "MulticoreTSNE8core")
parse_multicore_optimization8 = make_regex_func(r"Fitting performed in (\d+\.\d+) seconds", "MulticoreTSNE8core")
multicore8 = simplenamespace(full=parse_multicore8, nn=parse_multicore_nn8, optim=parse_multicore_optimization8)
print(multicore8.full(1000))
parse_sklearn = make_regex_func(r"scikit-learn t-SNE: (\d+\.\d+)", "sklearn")
parse_sklearn_nn = make_regex_func(r"neighbors for .* samples in (\d+\.\d+)s", "sklearn")
def parse_sklearn_optimization(n_points):
full_times = parse_sklearn(n_points)
nn_times = parse_sklearn_nn(n_points)
return full_times - nn_times
sklearn = simplenamespace(full=parse_sklearn, nn=parse_sklearn_nn, optim=parse_sklearn_optimization)
print(sklearn.full(10000))
parse_umap = make_regex_func(r"UMAP: (\d+\.\d+)", "UMAP")
umap = simplenamespace(full=parse_umap, nn=lambda *a, **kw: ..., optim=lambda *a, **kw: ...)
print(umap.full(10000))
import warnings
df = pd.DataFrame(columns=["time", "nn_time", "optim_time", "n_samples", "method"])
for method_name, method in [("openTSNE (1 core)", opentsne),
("openTSNE (8 cores)", opentsne8),
("MulticoreTSNE (1 core)", multicore),
("MulticoreTSNE (8 cores)", multicore8),
("FIt-SNE (1 core)", fitsne),
("FIt-SNE (8 cores)", fitsne8),
("scikit-learn (1 core)", sklearn),
("UMAP (1 core)", umap)]:
for n in n_points:
try:
tmp_df = pd.DataFrame({
"time": method.full(n),
#"nn_time": method.nn(n),
#"optim_time": method.optim(n),
})
tmp_df["n_samples"] = n
tmp_df["method"] = method_name
df = df.append(tmp_df, ignore_index=True)
except FileNotFoundError as e:
warnings.warn(str(e))
df.head()
df.groupby(["method", "n_samples"]).count()
import seaborn as sns
#sns.set("notebook", "whitegrid")
df["time_min"] = df["time"] / 60
df.head()
fig, ax = plt.subplots(figsize=(8, 6))
#g = sns.lineplot(x="n_samples", y="time_min", hue="method", data=df, ax=ax)
sns.despine(offset=20)
ax.set_title("t-SNE implementation benchmarks", loc="Left", fontdict={
"fontsize": "13"
})
ax.set_xlabel("$n$ samples")
ax.set_ylabel("Time (in minutes)")
ax.grid(color="0.9", linestyle="--", linewidth=1)
# Lines
d = df.groupby(["method", "n_samples"]).mean().reset_index()
d_std = df.groupby(["method", "n_samples"]).std().reset_index()
# openTSNE
which = "openTSNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#4C72B0", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#4C72B0")
which = "openTSNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#4C72B0", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#4C72B0")
# FIt-SNE
which = "FIt-SNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#DD8452", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#DD8452")
which = "FIt-SNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#DD8452", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#DD8452")
# MulticoreTSNE
which = "MulticoreTSNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#55A868", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#55A868")
which = "MulticoreTSNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#55A868", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#55A868")
# sklearn
which = "scikit-learn (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#C44E52", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#C44E52")
ax.set_xlim(0, 1_000_000)
ax.set_ylim(0, 130)
ax.set_yticks(range(0, 130, 15))
ax.get_xaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(
lambda x, p: format(int(x), ',').replace(",", ".")))
handles, labels = ax.get_legend_handles_labels()
ax.legend(frameon=False, loc='upper right')
plt.savefig("benchmarks.png", dpi=300, transparent=True)
fig, ax = plt.subplots(figsize=(8, 6))
#g = sns.lineplot(x="n_samples", y="time_min", hue="method", data=df, ax=ax)
sns.despine(offset=20)
ax.set_title("t-SNE implementation benchmarks", loc="Left", fontdict={
"fontsize": "13"
})
ax.set_xlabel("$n$ samples")
ax.set_ylabel("Time (in minutes)")
ax.grid(color="0.9", linestyle="--", linewidth=1)
# Lines
d = df.groupby(["method", "n_samples"]).mean().reset_index()
d_std = df.groupby(["method", "n_samples"]).std().reset_index()
# openTSNE
which = "openTSNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#4C72B0", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#4C72B0")
which = "openTSNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#4C72B0", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#4C72B0")
# FIt-SNE
which = "FIt-SNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#DD8452", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#DD8452")
which = "FIt-SNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#DD8452", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#DD8452")
# MulticoreTSNE
which = "MulticoreTSNE (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#55A868", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#55A868")
which = "MulticoreTSNE (8 cores)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#55A868", linestyle="dashed", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#55A868")
# sklearn
#which = "scikit-learn (1 core)"
#tmp = d[d["method"] == which]
#tmp1 = d_std[d_std["method"] == which]
#ax.plot(tmp["n_samples"], tmp["time_min"], c="#C44E52", label=which)
#ax.fill_between(tmp1["n_samples"],
# tmp["time_min"] + tmp1["time_min"],
# tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#C44E52")
# UMAP
which = "UMAP (1 core)"
tmp = d[d["method"] == which]
tmp1 = d_std[d_std["method"] == which]
ax.plot(tmp["n_samples"], tmp["time_min"], c="#C44E52", label=which)
ax.fill_between(tmp1["n_samples"],
tmp["time_min"] + tmp1["time_min"],
tmp["time_min"] - tmp1["time_min"], alpha=0.25, color="#C44E52")
ax.set_xlim(0, 1_000_000)
ax.set_ylim(0, 130)
ax.set_yticks(range(0, 130, 15))
ax.get_xaxis().set_major_formatter(matplotlib.ticker.FuncFormatter(
lambda x, p: format(int(x), ',').replace(",", ".")))
handles, labels = ax.get_legend_handles_labels()
ax.legend(frameon=False, loc='upper right')
#plt.savefig("benchmarks.png", dpi=300, transparent=True)
```
| github_jupyter |
```
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
path = "data/kc_house_data.csv"
df = pd.read_csv(path, parse_dates=["date"])
df.head(3)
def get_features(data):
corr = data.corr()['price']
for lst in corr:
if lst > 0.4:
print(lst)
else:
pass
dd = get_features(df)
dd
import numpy as np
corr = df.corr()["price"]
corr[np.argsort(corr, axis=0)[::-1]]
not_include = ['id', 'zipcode', 'long', 'condition']
features = list(df.columns)
rms = [1,2,4,5,6,7,8]
def remove(lst):
x = lst
for i in x:
if i > 6:
print(i)
else:
pass
remove(rms)
# we can use the important features to train our model
X = df[['sqft_above', 'sqft_living', 'grade', 'sqft_living15', 'bathrooms', 'view', 'sqft_basement', 'bedrooms', 'lat', 'waterfront', 'floors']]
# assign price column in our target output
y = df['price']
# our X only holds variables that are related to 'price'
X.head()
# Splitting our data in subsets
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=0)
# using 'RobustScaler' makes our data scaled in a robust way... depends on your target values
#from sklearn.preprocessing import RobustScaler
#scaler = RobustScaler()
#robust_scaled_df = scaler.fit_transform(X_train)
#robust_scaled_df = pd.DataFrame(robust_scaled_df)
#robust_scaled_df.shape
#robust_scaled_Test = scaler.fit_transform(X_test)
#robust_scaled_Test = pd.DataFrame(robust_scaled_Test)
#robust_scaled_Test.shape
X_train.head()
# Random forest are an ensemble learning method for classification, regression and other tasks.
# Random decision forests correct for decision trees' habit of overfitting to their training set.
# if
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(n_estimators=20, oob_score=True, random_state= 0)
rf.fit(X_train, y_train)
from sklearn.metrics import accuracy_score
predicted = rf.predict(X_test)
print('score estimate: ', round(rf.oob_score_, 3))
print('Mean accuracy score: ', round(rf.score(X_test, y_test), 3))
# we got score of 81% on our testing data, it is really good, but we can do better. If you notice that, we are only using
#... one parameter tuning in our model which is "n_estimators", next we'll use gridsearch to find best parameter for this model
# Setting parameters. Remember each model in sklearn has different parameter tuning, for example, logistic regression has two
#... well used parameter, 'penalty' for regularization which it has two options 'l1' or 'l2' and the second parameter
#... in logistic regression is 'C' parameter.
# But here, we are using 'randomForestRegression' which has few parameters we need to choose.
n_estimators = [20, 30, 60]
criterion = ['mse', 'mae']
max_features = ['auto','sqrt']
max_depth = [4,6]
n_jobs = [-1]
# Create hyperparameter options
hyperparameters = dict(n_estimators=n_estimators,
criterion=criterion,
max_features=max_features,
max_depth=max_depth,
n_jobs=n_jobs)
# Create grid search using 10-fold cross validation
from sklearn.model_selection import GridSearchCV
clf = GridSearchCV(rf, hyperparameters, cv=5, verbose=5)
# Fit grid search - NOTE: it will take time to train the data, start cv=5
best_model = clf.fit(X_train, y_train)
# View best hyperparameters - gridsearch
print('Best n_estimators:', best_model.best_estimator_.get_params()['n_estimators'])
print('Best criterion:', best_model.best_estimator_.get_params()['criterion'])
print('Best max_features:', best_model.best_estimator_.get_params()['max_features'])
print('Best max_depth:', best_model.best_estimator_.get_params()['max_depth'])
# Predict target values
best_model.predict(X_test)
# View the accuracy score
print('Best score for data1:', best_model.best_score_)
from sklearn.model_selection import RandomizedSearchCV
random = RandomizedSearchCV(rf, hyperparameters, random_state=1, n_iter=60, cv=5, verbose=0, n_jobs=-1)
compare_random = random.fit(X_train, y_train)
# View best hyperparameters - randomizedsearch
print('Best n_estimators:', compare_random.best_estimator_.get_params()['n_estimators'])
print('Best criterion:', compare_random.best_estimator_.get_params()['criterion'])
print('Best max_features:', compare_random.best_estimator_.get_params()['max_features'])
print('Best max_depth:', compare_random.best_estimator_.get_params()['max_depth'])
print('Best score for randomized search:', compare_random.best_score_)
# Note - if you not have xgboost installed in your anaconda, it won't work. Plus its expensive to run
#from xgboost.sklearn import XGBClassifier
#from xgboost.sklearn import XGBRegressor
#xclas = XGBClassifier() # and for classifier
#xclas.fit(X_train, y_train)
#xclas.predict(X_test)
# normalizing using standardscaler: NOTE to scale our data, because our output in not binary (0, 1), if we're to normalize our
#... data, it would heavly effect our score... some data are better to normalize and some are not...
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X_train)
X_train_scaled = pd.DataFrame(scaler.transform(X_train), index=X_train.index.values, columns=X_train.columns.values)
X_test_scaled = pd.DataFrame(scaler.transform(X_test), index=X_test.index.values, columns=X_test.columns.values)
from sklearn.decomposition import PCA
# Note - this cell takes more time to execute, just uncomment 'sns' to see the output
pca = PCA()
pca.fit(X_train)
cpts = pd.DataFrame(pca.transform(X_train))
x_axis = np.arange(1, pca.n_components_+1)
pca_scaled = PCA()
pca_scaled.fit(X_train_scaled)
cpts_scaled = pd.DataFrame(pca.transform(X_train_scaled))
# matplotlib boilerplate goes here
#sns.pairplot(cpts_scaled)
rf2 = RandomForestRegressor(n_estimators=500, oob_score=True, random_state=0)
rf2.fit(X_train, y_train)
from sklearn.metrics import r2_score
from scipy.stats import spearmanr, pearsonr
predicted_train = rf2.predict(X_train)
predicted_test = rf2.predict(X_test)
test_score = r2_score(y_test, predicted_test)
# finds corr between two vairble that are related
# FIND: http://statisticslectures.com/topics/spearman/
spearman = spearmanr(y_test, predicted_test)
#Pearsonr measures the strength of the linear relationship between two variables. Pearson�s r is always between -1 and 1.
pearson = pearsonr(y_test, predicted_test)
print('Out-of-bag R-2 score estimate: ', round(rf2.oob_score_, 3))
print('Test data R-2 score: ', round(test_score, 3))
print('Test data Spearman correlation: ', round(spearman[0], 3))
print('Test data Pearson correlation: ',round(pearson[0], 3))
```
| github_jupyter |
# CT-LTI: Figure 3
Figure for qualitative evaluation of reached states between NODEC and OC.
Furthermore, please make sure that the required data folder is available at the paths used by the script.
You may generate the required data by running the python script
```nodec_experiments/ct_lti/gen_parameters.py```.
Please also make sure that a trainingproceedure has produced results in the corresponding paths used below.
Running ```nodec_experiments/ct_lti/single_sample/train.ipynb``` with default paths is expected to generate at the requiered location.
As neural network intialization is stochastic, please make sure that appropriate seeds are used or expect some variance to paper results.
```
# %load_ext autoreload
# %autoreload 2
import os
os.sys.path.append('../../../')
import torch
from torchdiffeq import odeint
import numpy as np
import pandas as pd
import networkx as nx
import plotly
from copy import deepcopy
import scipy
from plotly import graph_objects as go
import plotly.express as px
from tqdm.auto import tqdm
from nnc.helpers.plot_helper import square_lattice_heatmap, trendplot
from nnc.helpers.torch_utils.file_helpers import read_tensor_from_collection, \
save_tensor_to_collection
from nnc.helpers.plot_helper import ColorRegistry, base_layout
from plotly.subplots import make_subplots
```
## Load required Data
This script requires ```train.ipynb``` to have run first or to download the precomputed results in the ```data``` folder.
```
# Load the results
results_data_folder = '../../../../data/results/ct_lti/single_sample/'
experiment_data_folder = '../../../../data/parameters/ct_lti/'
graph='lattice'
n_interactions = ['50', '500', '5000']
collection_file = 'epochs.zip'
evaluation_files = dict(
oc_50 = results_data_folder + 'oc_sample_ninter_50/',
oc_500 = results_data_folder + 'oc_sample_ninter_500/',
oc_5000 = results_data_folder + 'oc_sample_ninter_5000/',
nodec_50 = results_data_folder + 'eval_nn_sample_ninter_50/',
nodec_500 = results_data_folder + 'eval_nn_sample_ninter_500/',
nodec_5000 = results_data_folder + 'eval_nn_sample_ninter_5000/',
)
all_files = dict(
train_file = results_data_folder + 'nn_sample_train/',
)
all_files.update(evaluation_files)
# Load graph and initial-target states
graph='lattice'
graph_folder = experiment_data_folder+graph+'/'
device='cpu'
target_states = torch.load(graph_folder+'target_states.pt').to(device)
initial_states = torch.load(experiment_data_folder+'init_states.pt').to(device)
current_sample_id = 24
x0 = initial_states[current_sample_id].unsqueeze(0)
xstar = target_states[current_sample_id].unsqueeze(0)
T = 0.5
```
## Load initial and target states
```
fig_x0 = square_lattice_heatmap(x0, color_scale=
plotly.colors.sequential.Agsunset)
fig_x0.layout.paper_bgcolor = 'rgba(0,0,0,0)'
fig_xstar = square_lattice_heatmap(xstar, color_scale=
plotly.colors.sequential.Agsunset)
fig_xstar.layout.paper_bgcolor = 'rgba(0,0,0,0)'
```
## Load reached states for all baselines and different interactions
```
all_figs = dict()
for file, path in all_files.items():
metadata = pd.read_csv(path+'epoch_metadata.csv')
epoch = metadata['epoch'].iloc[-1]
reached_state = read_tensor_from_collection(path + 'epochs', 'reached_state/ep_'+str(epoch)+'.pt')
fig = square_lattice_heatmap(reached_state, color_scale=
plotly.colors.sequential.Agsunset)
all_figs[file] = fig
fig.layout.paper_bgcolor = 'rgba(0,0,0,0)'
## Plot the figure
fig = make_subplots(2,4,
row_heights=2*[80],
column_width = 4*[80],
horizontal_spacing = 0.05,
vertical_spacing=0.1,
subplot_titles=['$x_0$', 'NODEC 0.01', 'NODEC 0.001', 'NODEC 0.0001',
'$x^*$', 'OC 0.01', 'OC 0.001', 'OC 0.0001',
]
)
fig.add_trace(fig_x0.data[0], 1, 1)
fig.add_trace(fig_xstar.data[0], 2, 1)
fig.add_trace(all_figs['nodec_5000'].data[0], 1, 2)
fig.add_trace(all_figs['nodec_500'].data[0], 1, 3)
fig.add_trace(all_figs['nodec_50'].data[0], 1, 4)
fig.add_trace(all_figs['oc_5000'].data[0], 2, 2)
fig.add_trace(all_figs['oc_500'].data[0], 2, 3)
fig.add_trace(all_figs['oc_50'].data[0], 2, 4)
fig.update_xaxes(visible = False)
fig.update_yaxes(visible = False)
fig.layout.margin = dict(t=30,b=30,l=30,r=30)
fig.update_traces(showscale=False)
fig
```
| github_jupyter |
# Using crack submodels in PyBaMM
In this notebook we show how to use the crack submodel with battery DFN or SPM models. To see all of the models and submodels available in PyBaMM, please take a look at the documentation [here](https://pybamm.readthedocs.io/en/latest/source/models/index.html).
```
%pip install pybamm -q # install PyBaMM if it is not installed
import pybamm
import os
import numpy as np
import matplotlib.pyplot as plt
os.chdir(pybamm.__path__[0]+'/..')
```
Then we load the DFN, SPMe or SPM, by choosing one and commenting the others.
When you load a model in PyBaMM it builds by default. Building the model sets all of the model variables and sets up any variables which are coupled between different submodels: this is the process which couples the submodels together and allows one submodel to access variables from another.
```
model = pybamm.lithium_ion.DFN(
options = {
"particle": "Fickian diffusion",
"particle mechanics": "swelling and cracking", # other options are "none", "swelling only"
}
)
```
Load the parameter set Ai2020 which contains mechanical parameters. Other sets may not contain mechanical parameters should add them manually.
```
param = pybamm.ParameterValues("Ai2020")
## It can update the speed of crack propagation using the commands below:
# param.update({"Negative electrode Cracking rate":3.9e-20*10})
```
We can get the default parameters for the model and update them with the parameters required by the cracking model. Eventually, we would like these to be added to their own chemistry (you might need to adjust the path to the parameters file to your system).
Now the model can be processed and solved in the usual way, and we still have access to model defaults such as the default geometry and default spatial methods
```
sim = pybamm.Simulation(
model,
parameter_values=param,
solver=pybamm.CasadiSolver(dt_max=600),
)
solution = sim.solve(t_eval=[0, 3600], inputs={"C-rate": 1})
# plot
quick_plot = pybamm.QuickPlot(solution)
quick_plot.dynamic_plot()
```
Plot the results as required.
```
# extract voltage
stress_t_n_surf = solution["Negative particle surface tangential stress"]
x = solution["x [m]"].entries[0:19, 0]
c_s_n = solution['Negative particle concentration']
r_n = solution["r_n [m]"].entries[:, 0, 0]
# plot
def plot_concentrations(t):
f, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4 ,figsize=(20,4))
ax1.plot(x, stress_t_n_surf(t=t,x=x))
ax1.set_xlabel(r'$x_n$ [m]')
ax1.set_ylabel('$\sigma_t/E_n$')
plot_c_n, = ax2.plot(r_n, c_s_n(r=r_n,t=t,x=x[0])) # can evaluate at arbitrary x (single representative particle)
ax2.set_ylabel('Negative particle concentration')
ax2.set_xlabel(r'$r_n$ [m]')
ax2.set_ylim(0, 1)
ax2.set_title('Close to current collector')
ax2.grid()
plot_c_n, = ax3.plot(r_n, c_s_n(r=r_n,t=t,x=x[10])) # can evaluate at arbitrary x (single representative particle)
ax3.set_ylabel('Negative particle concentration')
ax3.set_xlabel(r'$r_n$ [m]')
ax3.set_ylim(0, 1)
ax3.set_title('In the middle')
ax3.grid()
plot_c_n, = ax4.plot(r_n, c_s_n(r=r_n,t=t,x=x[-1])) # can evaluate at arbitrary x (single representative particle)
ax4.set_ylabel('Negative particle concentration')
ax4.set_xlabel(r'$r_n$ [m]')
ax4.set_ylim(0, 1)
ax4.set_title('Close to separator')
ax4.grid()
plt.show()
import ipywidgets as widgets
widgets.interact(plot_concentrations, t=widgets.FloatSlider(min=0,max=3600,step=10,value=0));
```
Plot results using the default functions
```
label = ["Crack model"]
output_variables = [
"Negative particle crack length",
"Positive particle crack length",
"X-averaged negative particle crack length",
"X-averaged positive particle crack length"
]
quick_plot = pybamm.QuickPlot(solution, output_variables, label,variable_limits='tight')
quick_plot.dynamic_plot();
```
## References
The relevant papers for this notebook are:
```
pybamm.print_citations()
```
| github_jupyter |
```
import pnad
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pnad import Gender, Race
```
# Accessing data
First, select the year we will use to make the rest of the analysis.
Our main entry point is the `pnad.load_person` function, that load specific columns from a PNAD survey of an specific year. We will select some basic variables.
```
YEAR = 2012
data = pnad.load_person(YEAR, ['income', 'age', 'education_years', 'race', 'gender', 'weight'])
```
Pandas makes it very easy to filter and transform the dataframe. We will select only the adult entries (people with age >= 18) and make some further analysis.
We use the `groupby` function of data frames to collect basic statistics about our sample. We can, for instance, check how income and education differs in different categories.
```
adults = data[data.age >= 18]
adults.groupby('race').mean()
adults.groupby('gender').mean()
```
Those results cannot be taken as the real sample mean since we do not consider the statistical weight of each entry. The "weight" column indicates the number of people each entry represents in the survey. Summing over it gives an estimate of the total population.
```
population_mi = data.weight.sum() / 1e6
population_mi
```
## Basic statistics
The values shown on the tables above are similar to the correct ones, but now we will account for the weights.
```
col = 'income'
by = ['gender'] # ['race', 'gender']
a, b = 70, 75
df = data[(data.age > a) | (data.age < b)]
df.income.fillna(0, inplace=True)
df = data
df['weighted'] = df[col] * df.weight
groups = df[[*by, 'weight', 'weighted']].groupby(by).sum()
income = groups.weighted / groups.weight
income / income.max()
```
# Age distribution
```
df = data[data.race != 'UNKNOWN']
df = df[['gender', 'age', 'weight']]
MAX_AGE = 120
females = df[['age', 'weight']][df.gender == 'FEMALE']
males = df[['age', 'weight']][df.gender == 'MALE']
female_data = females.groupby('age').sum().weight.values.copy()
male_data = males.groupby('age').sum().weight.values.copy()
def smooth(arr, by=5):
arr.resize(MAX_AGE, refcheck=False)
arr = arr.reshape((MAX_AGE // by, by))
return arr.sum(1)
female_data = smooth(female_data)
male_data = smooth(male_data)
plt.barh(np.arange(2, MAX_AGE, 5), female_data / 1e6, height=4.0, label='female')
plt.barh(np.arange(2, MAX_AGE, 5), -male_data / 1e6, height=4.0, label='male')
plt.plot([0, 0], [0, MAX_AGE], 'r--')
plt.legend()
ax = plt.gca()
ax.set_aspect(1.5e6 / max(female_data.max(), male_data.max()))
plt.show()
import pnad
import pandas as pd
import matplotlib.pyplot as plt
year_born = 1982
df = pd.DataFrame()
for year in pnad.years():
if year in (1976, 1977, 1978, 1979, 1981, 1983, 1984, 1985, 1986, 1988, 2015):
continue
print(year)
data = pnad.load_person(year, ['race', 'weight', 'age'])
data = data[data.age == year - year_born]
df[year - year_born] = (data[['race', 'weight']].groupby('race').sum() / data.weight.sum()).weight
#df[year - year_born] = data.groupby('race').count().weight
df.T.plot()
plt.show()
df.T.WHITE.plot()
df.T.iloc[5:].plot()
plt.show()
df.T
df = pnad.load_person(1982, ['age', 'race', 'gender', 'income', 'weight'])
df = df[df.income >= 2 * df.income.mean()]
df = data[data.race != 'UNKNOWN'].groupby(['age', 'race']).sum()
df[df.index.names] = df.index.to_frame()
x = df.pivot('age', 'race', 'weight').fillna(0).iloc[:90]
x = x / x.sum(1).values[:, None]
x.plot()
x.plot.scatter('BROWN', 'WHITE')
x
np.corrcoef(x.BLACK, x.WHITE + x.BROWN)
x.corr()
data
from pnad import Race
s = Race.categorical([1, 2, 4, 8, 16, 16, 16, 2, 2, 2, 2])
s == 'ASIAN'
s.cat.
import sklearn.encode
df = data[['income', 'education_years', 'gender']].copy()
df['gender'] = (df.pop('gender') == 'MALE') + 0.0
from sklearn.decomposition import PCA
plt.scatter(*PCA(2).fit_transform(df.fillna(0).values).T)
plt.show()
from sklearn.naive_bayes import GaussianNB
nb = GaussianNB()
X = df.fillna(0).values
y = (data.race == 'WHITE') | (data.race == 'ASIAN')
nb.fit(X, y)
nb.score(X, y)
from sklearn.ensemble import RandomForestClassifier
rf = ExtraTreesClassifier()
rf.fit(X, y)
rf.score(X, y)
rf.feature_importances_
from sklearn.tree import DecisionTreeClassifier
dt = DecisionTreeClassifier(max_depth=3)
dt.fit(X, y)
dt.score(X, y)
dt.decision_path(X)
tree.plot_tree(dt)
path = '/home/chips/usa_00001.csv.gz'
pd.read_csv(path)
d = _
d.describe()
df = d[['YEAR', 'RACE', 'AGE', 'HHWT']]
df = df[df.YEAR == 2002]
df.HHWT.sum()
dd = df[['HHWT', 'RACE', 'AGE']].groupby(['AGE', 'RACE']).sum()
dd[['AGE', 'RACE']] = dd.index.to_frame()
d1 = dd.pivot('AGE', 'RACE', 'HHWT')
d1 /= d1.sum(1).values[:, None]
d1.plot()
```
| github_jupyter |
```
class GoppaCode:
def __init__(self, n, m, g):
t = g.degree()
F2 = GF(2)
F_2m = g.base_ring()
Z = F_2m.gen()
PR_F_2m = g.parent()
X = PR_F_2m.gen()
factor_list = list(factor(2 ^ m - 1))
final_factor_list = []
for i in range(len(factor_list)):
for j in range(factor_list[i][1]):
final_factor_list.append(factor_list[i][0])
while 1:
primitive_root = F_2m.random_element()
if primitive_root == 0:
continue
for i in range(len(final_factor_list)):
for j in itertools.combinations(final_factor_list, i):
exponent = 1
for _ in range(len(j)):
exponent *= j[_]
if primitive_root ^ exponent == 1:
output = False
break
else:
output = True
continue
if not output:
break
if output:
break
codelocators = []
for i in range(2 ^ m - 1):
codelocators.append(primitive_root ^ (i + 1))
codelocators.append(F_2m(0))
h = PR_F_2m(1)
gamma = []
for a_i in codelocators:
gamma.append((h * ((X - a_i).inverse_mod(g))).mod(g))
H_check_poly = matrix(F_2m, t, n)
for i in range(n):
coeffs = list(gamma[i])
for j in range(t):
if j < len(coeffs):
H_check_poly[j, i] = coeffs[j]
else:
H_check_poly[j, i] = F_2m(0)
H_Goppa = matrix(F2, m * H_check_poly.nrows(), H_check_poly.ncols())
for i in range(H_check_poly.nrows()):
for j in range(H_check_poly.ncols()):
be = bin(H_check_poly[i, j].integer_representation())[2:]
be = be[::-1]
be = be + '0' * (m - len(be))
be = list(be)
H_Goppa[m * i:m * (i + 1), j] = vector(map(int, be))
G_Goppa = H_Goppa.transpose().kernel().basis_matrix()
G_Goppa_poly = H_check_poly.transpose().kernel().basis_matrix()
SyndromeCalculator = matrix(PR_F_2m, 1, len(codelocators))
for i in range(len(codelocators)):
SyndromeCalculator[0, i] = (X - codelocators[i]).inverse_mod(g)
self._n = n
self._m = m
self._g = g
self._t = t
self._codelocators = codelocators
self._SyndromeCalculator = SyndromeCalculator
self._H_Goppa = H_Goppa
self._H_gRS = H_check_poly
self._G_Goppa = G_Goppa
def _split(self, p):
Phi = p.parent()
p0 = Phi([sqrt(c) for c in p.list()[0::2]])
p1 = Phi([sqrt(c) for c in p.list()[1::2]])
return p0, p1
def _g_inverse(self, p):
(d, u, v) = xgcd(p, self.goppa_polynomial())
return u.mod(self.goppa_polynomial())
def _norm(self, a, b):
X = self.goppa_polynomial().parent().gen()
return 2 ^ ((a ^ 2 + X * b ^ 2).degree())
def _lattice_basis_reduce(self, s):
g = self.goppa_polynomial()
t = g.degree()
a = [0]
b = [0]
(q, r) = g.quo_rem(s)
(a[0], b[0]) = simplify((g - q * s, 0 - q))
if self._norm(a[0], b[0]) > 2 ^ t:
a.append(0)
b.append(0)
(q, r) = s.quo_rem(a[0])
(a[1], b[1]) = (r, 1 - q * b[0])
if a[1] == 0:
return s, 1
else:
return a[0], b[0]
i = 1
while self._norm(a[i], b[i]) > 2 ^ t:
a.append(0)
b.append(0)
(q, r) = a[i - 1].quo_rem(a[i])
(a[i + 1], b[i + 1]) = (r, b[i - 1] - q * b[i])
i += 1
return (a[i], b[i])
def SyndromeDecode(self, syndrome_poly, mode='Patterson'):
g = self.goppa_polynomial()
X = g.parent().gen()
error = matrix(GF(2), 1, self.parity_check_matrix().ncols())
if mode == 'Patterson':
(g0, g1) = self._split(g)
sqrt_X = g0 * self._g_inverse(g1)
T = syndrome_poly.inverse_mod(g)
(T0, T1) = self._split(T - X)
R = (T0 + sqrt_X * T1).mod(g)
(alpha, beta) = self._lattice_basis_reduce(R)
sigma = (alpha * alpha) + (beta * beta) * X
if (X ^ (2 ^ self._m)).mod(sigma) != X:
print("sigma: Decodability Test Failed")
return error # return a zero vector
for i in range(len(self._codelocators)):
if sigma(self._codelocators[i]) == 0:
error[0, i] = 1
return error
def generator_matrix(self):
return self._G_Goppa
def goppa_polynomial(self):
return self._g
def parity_check_matrix(self):
return self._H_Goppa
from math import floor
import itertools
def GetGoppaPolynomial(polynomial_ring, polynomial_degree):
while 1:
irr_poly = polynomial_ring.random_element(polynomial_degree)
irr_poly_list = irr_poly.list()
irr_poly_list[-1] = 1
irr_poly = polynomial_ring(irr_poly_list)
if irr_poly.degree() != polynomial_degree:
continue
elif irr_poly.is_irreducible():
break
else:
continue
return irr_poly
class Niederreiter:
def __init__(self):
m = 4
n = 2 ** m
t = 2
F_2m = GF(n, 'Z', modulus='random')
PR_F_2m = PolynomialRing(F_2m, 'X')
Z = F_2m.gen()
X = PR_F_2m.gen()
irr_poly = GetGoppaPolynomial(PR_F_2m, t)
goppa_code = GoppaCode(n, m, irr_poly)
k = goppa_code.generator_matrix().nrows()
# Random binary non singulary matrix -> S
S = matrix(GF(2), n - k, [random() < 0.5 for _ in range((n - k) ^ 2)])
while rank(S) < n - k:
S[floor((n - k) * random()), floor((n - k) * random())] += 1
# parity check matrix for code -> H
H = goppa_code.parity_check_matrix()
# Random permutation matrix -> P
rng = range(n)
P = matrix(GF(2), n)
for i in range(n):
p = floor(len(rng) * random())
P[i, rng[p]] = 1
rng = [*rng[:p], *rng[p + 1:]]
self._m_GoppaCode = goppa_code
self._g = irr_poly
self._t = self._g.degree()
self._S = S
self.H = H
self._P = P
self._PublicKey = S * H * P
def encrypt(self, message):
# verify length of message
assert (message.ncols() == self._PublicKey.ncols()), "Message is not of the correct length"
code_word = self._PublicKey*(message.transpose())
return code_word.transpose()
def decrypt(self, received_word):
# verify length of received word
received_word = received_word.transpose()
assert (received_word.nrows() == self._PublicKey.nrows()), "Received word is not of the correct row length"
assert (received_word.ncols() == 1), "Received word is not of the correct column length"
message = ~(self._S)*received_word
# Syndrome decoding
t = self._t
m = message.nrows()/t
g = self._m_GoppaCode.goppa_polynomial()
F2 = GF(2)
F_2m = g.base_ring()
Z = F_2m.gen()
PR_F_2m = g.parent()
X = PR_F_2m.gen()
syndrome_poly = 0
for i in range(t):
tmp = []
for j in range(m):
tmp.append(message[i*m+j,0])
syndrome_poly += F_2m(tmp[::1])*X^i
message = self._m_GoppaCode.SyndromeDecode(syndrome_poly)
message = message*self._P
return message
# function to create a random message to encrypt
def GetRandomMessageWithWeight(message_length, message_weight):
message = matrix(GF(2), 1, message_length)
rng = range(message_length)
for i in range(message_weight):
p = floor(len(rng)*random())
message[0,rng[p]] = 1
rng=[*rng[:p],*rng[p+1:]]
return message
crypto = Niederreiter()
message = GetRandomMessageWithWeight(crypto._PublicKey.ncols(),crypto._g.degree())
encrypted_message = crypto.encrypt(message)
decrypted_message = crypto.decrypt(encrypted_message)
print('random message:', message.str())
print('encrypted message:', encrypted_message.str())
print('decrpted message:', decrypted_message.str())
print('decryption is: ', message==decrypted_message)
```
| github_jupyter |
# Hybrid Monte Carlo
## Products And Exposures
In this notebook we demonstrate the setup of products and the calculation of exposures. In practice products are specified by actual dates (not model times). We use QuantLib to handle date arithmetics (calenders, day counts, etc.) in product specifications.
This notebook is structured along the following sections:
1. Setting up a cash flow leg in QuantLib
2. Constructing and inspecting a swap product
3. Calculating exposures with MC and AMC
We use a couple of standard packages for calculation and analysis
```
import sys
sys.path.append('../') # make python find our modules
import numpy as np
import pandas as pd
import plotly.express as px
```
The following auxilliary method lists the relevant members of an object. We use it to inspect the objects created.
```
def members(obj):
return [f for f in dir(obj) if not f.startswith('_')]
```
### Setting up a cash flow leg in QuantLib
For general details on QuantLib see e.g. https://www.quantlib.org/
Here we demonstrate the features required to set up simple cash flow legs.
QuantLib has a global evaluation date. This is the date from which time periods for term structures and models are calculated. We set evaluation date at inception and keep it fixed during the run of the session.
```
import QuantLib as ql
today = ql.Date(5,ql.October,2020)
ql.Settings.instance().evaluationDate = today
```
We want to set up a fixed leg and a standard floating leg. For multi-curve modelling we need to take into account the tenor basis. Roughly speaking, tenor basis is the difference between projection curve and discount curve. Consequently, to model tenor basis we need a projection curve and a discount curve.
```
discYtsH = ql.YieldTermStructureHandle(
ql.FlatForward(today,0.01,ql.Actual365Fixed()))
projYtsH = ql.YieldTermStructureHandle(
ql.FlatForward(today,0.02,ql.Actual365Fixed()))
```
In this example we set discount curve flat at *1%* continuous componded zero rate and projection curve flat at *2%* continuous componded zero rate. A *ql.FlatForward()* object is equivalent to our *YieldCurve()* object. QuantLib's *ql.YieldTermStructureHandle()* is just a double-indirection for the underlying curve.
Euribor/Libor forward rates are modelled in QunatLib via indizes. We set up an *Euribor* index.
```
index = ql.Euribor6M(projYtsH)
```
Cash flow dates are specified via *Schedule* objects. For details on how to construct QuantLib schedules see https://github.com/lballabio/QuantLib-SWIG/blob/master/SWIG/scheduler.i.
```
# we set start in the future to avoid the need of index fixings
startDate = ql.Date(12,ql.October,2020)
endDate = ql.Date(12,ql.October,2030)
calendar = ql.TARGET()
fixedTenor = ql.Period('1y')
floatTenor = ql.Period('6m')
fixedSchedule = ql.MakeSchedule(startDate,endDate,tenor=fixedTenor,calendar=calendar)
floatSchedule = ql.MakeSchedule(startDate,endDate,tenor=floatTenor,calendar=calendar)
```
A schedule behaves essentially like a list of dates.
```
display(list(fixedSchedule))
```
Now we can setup a fixed leg. For details on how to setup cash flow legs with QuantLib see https://github.com/lballabio/QuantLib-SWIG/blob/master/SWIG/cashflows.i.
```
couponDayCount = ql.Thirty360()
notional = 1.0
fixedRate = 0.02
fixedLeg = ql.FixedRateLeg(fixedSchedule,couponDayCount,[notional],[fixedRate])
```
Similarly, we can setup a floating rate leg.
```
floatingLeg = ql.IborLeg([notional],floatSchedule,index)
```
Cash flow legs behave like lists of cash flows.
```
display([ cf.amount() for cf in fixedLeg ])
```
We can calculate present value of the legs using the discount curve.
```
display('FixedLeg npv: %.4f' % ql.CashFlows_npv(fixedLeg,discYtsH,True))
display('FloatingLeg npv: %.4f' % ql.CashFlows_npv(floatingLeg,discYtsH,True))
```
### Constructing and inspecting a swap product
A *Product* object represents a financial instrument with one or several cash flows. Each cash flow is represented as a Monte Carlo payoff $V_i(T_i)$ that is paid at $T_i$.
The *Product* objects implement a function *cashflows(obsTime)*. This function calculates payoffs with observation time $t$ that calculate (or estimate)
$$
V(t) = B(t) \mathbb{E} \left[
\sum_{T_i>t} \frac{V_i(T_i)}{B(T_i)} \, | \, \cal{F}_t
\right]
= \sum_{T_i>t} \mathbb{E} \left[
\frac{V_i(T_i)}{B(T_i)} \, | \, \cal{F}_t
\right].
$$
If the payoffs $V_i$ are simple enough such that $\mathbb{E} \left[ V_i(T_i) / B(T_i) \, | \, \cal{F}_t \right]$ can be calculated in closed form then we use this analytic expression in the cash flow method. This is typically the case for linear products and Vanilla options.
For complex payoffs without analytical expression for $\mathbb{E} \left[ V_i(T_i) / B(T_i) \, | \, \cal{F}_t \right]$ we use AMC to estimate the conditional expectation.
We implement a *Swap* product that uses the discounted cash flows and forward Libor rates for the analytical payoff expessions.
A *Swap* product is represented by a list of fixed or Ibor legs. Moreover, we need to specify whether we receive (+1) or pay (-1) a leg. The calculation of tenor basis also requires the discount curve.
```
from hybmc.products.Swap import Swap
swap = Swap([fixedLeg,floatingLeg],[1.0,-1.0],discYtsH)
```
We check the cash flow calculation at $t=9.0$, i.e. approximately 1y bevor swap maturity.
```
cfs = swap.cashFlows(9.0)
display([str(cf) for cf in cfs])
```
We make the following observations:
- We have two remaining fixed leg payments. The first fixed leg payment is paid at $T=9.02$ and
the second fixed leg payment is paid at $T=10.03$.
- Similarly, we have three remaining floating rate payments.
- Payoffs are discounted to time $t=9.0$, see *P_None(9.00,.)*.
- Payoffs are observed at $t=9.0$, see *@ 9.00*.
Also note that the first Libor cash flow has fixing time $8.52$ compared to $9.0$ for the other cash flows. This is correct, because at observation time $t=9.0$ the the Libor rate is alsready fixed but the coupon is not yet paid.
For the other Libor cash flows the actual Libor fixing is in the future (later than $t=9.0$). However, we can calculate (using $T_i$-forward measure)
$$
B(t) \cdot \mathbb{E} \left[ V_i(T_i) / B(T_i) \, | \, \cal{F}_t \right]
=
P(t,T_i) \cdot \mathbb{E}^{T_i} \left[ V_i(T_i) \, | \, \cal{F}_t \right].
$$
And for a Libor rate $\mathbb{E}^{T_i} \left[ L_i(T_i) \, | \, \cal{F}_t \right]$ becomes the forward Libor rate with observation time equal to $t$.
If we calculate cash flows at $t=0$ then we get the full list of product cash flows.
```
cfs = swap.cashFlows(0.0)
display([str(cf) for cf in cfs])
```
We can use a deterministic model to calculate these payoffs.
```
from hybmc.models.DeterministicModel import DcfModel
path = DcfModel(discYtsH).path()
amounts = np.array([ cf.discountedAt(path) for cf in cfs ])
display(amounts)
```
We can double-check the valuation against QuantLib's valuation of the fixed and float leg.
```
amountsQl = np.array(
[cf.amount() * discYtsH.discount(cf.date()) for cf in fixedLeg] +
[-cf.amount() * discYtsH.discount(cf.date()) for cf in floatingLeg] )
display(amountsQl)
```
This looks good except the 12th floating rate cash flow (with index 11):
```
display(str(cfs[21]))
display(cfs[21].discountedAt(path))
cf = list(floatingLeg)[11]
display(-cf.amount() * discYtsH.discount(cf.date()))
```
That is an interesting case. Differences are probably due to some slight date mismatch in accrual period versus fixing period.
### Calculating exposures with MC and AMC
Once the cash flow method is implemented for a product we can call it for a range of observation times. This gives a *time line* of payoffs.
```
timeline = swap.timeLine([0.0, 3.0, 10.0])
for t in timeline:
print('ObsTime: %.2f' % t)
for p in timeline[t]:
print(p)
```
For exposure valuation we need to set up a model and a MC simulation.
```
from hybmc.models.HullWhiteModel import HullWhiteModel
from hybmc.simulations.McSimulation import McSimulation
model = HullWhiteModel(discYtsH,0.03,np.array([10.0]),np.array([0.0050]))
mcsim = McSimulation(model,np.linspace(0.0,10.0,41),2**10,314159265359,True)
```
Now we can calculate scenarios for the time line.
```
times = np.linspace(0.0,10.0,41)
scens = swap.scenarios(times,mcsim)
```
For exposure simulation we are interested in the $\mathbb{E}[V(t)^+]$.
```
epeDcf = np.average(np.maximum(scens,0.0),axis=0)
```
We plot the exposure profile.
```
dfDcf = pd.DataFrame([ times, epeDcf ]).T
dfDcf.columns = ['times', 'epeDcf']
fig = px.line(dfDcf,x='times',y='epeDcf')
fig.show()
```
Alternatively (and as a proof of concept), we can also setup a swap time line using American Monte Carlo.
```
from hybmc.products.Swap import AmcSwap
mcsim_training = McSimulation(model,np.linspace(0.0,10.0,41),2**10,2718281828,True)
swap = AmcSwap([fixedLeg,floatingLeg],[1.0,-1.0],mcsim_training,2,discYtsH)
cfs = swap.cashFlows(9.0)
display([str(cf) for cf in cfs])
```
Here we see a single payoff per observation time. That payoff is a AMC regression payoff that references the actual swap payoffs at future pay times.
As regresssion variable we use a *co-terminal Libor rate*. That is a bit unusual, but it does the job.
Similarly, as with the analytic approach we calculate scenarion and expected (positive) exposures.
```
scens = swap.scenarios(times,mcsim)
epeAmc = np.average(np.maximum(scens,0.0),axis=0)
dfAmc = pd.DataFrame([ times, epeAmc ]).T
dfAmc.columns = ['times', 'epeAmc']
fig = px.line(dfAmc,x='times',y='epeAmc')
fig.show()
```
Finally, we compare profiles from analytic and AMC method.
```
dfDcf.columns = ['times', 'epe']
dfDcf['type'] = 'Dcf'
dfAmc.columns = ['times', 'epe']
dfAmc['type'] = 'Amc'
df = pd.concat([dfDcf,dfAmc],axis=0)
fig = px.line(df,x='times',y='epe', color='type')
fig.show()
```
| github_jupyter |
## Arithmetic, Function Application, Mapping with pandas
### Working with pandas
*Curtis Miller*
Here we will see several examples of concepts discussed in the slides.
### `Series` Arithmetic
Let's first suit up.
```
import pandas as pd
from pandas import Series, DataFrame
import numpy as np
srs1 = Series([1, 9, -4, 3, 3])
srs2 = Series([2, 3, 4, 5, 10], index=[0, 1, 2, 3, 5])
print(srs1)
print(srs2)
```
Notice that the indices do not line up, even though the `Series` are of the same length.
Predict the outcomes:
```
srs1 + srs2
srs1 * srs2
srs1 ** srs2
# Boolean arithmetic is different
srs1 > srs2
srs1 <= srs2 # Opposite of above
srs1 > Series([1, 2, 3, 4, 5], index = [4, 3, 2, 1, 0])
np.sqrt(srs2)
np.abs(srs1)
type(np.abs(srs1))
# Define a cusom ufunc: notice the decorator notation?
@np.vectorize
def trunc(x):
return x if x > 0 else 0
trunc(np.array([-1, 5, 4, -3, 0]))
trunc(srs1)
type(trunc(srs1))
```
### `Series` Methods and Function Application
Having seen basic arithmetic with Series, let's look at useful Series methods.
```
# Mean of a series
srs1.mean()
srs1.std()
srs1.max()
srs1.argmax() # Returns the index where the maximum is
srs1.cumsum()
srs1.abs() # An alternative to the abs function in NumPy
```
Now let's look at function application and mapping.
```
srs1.apply(lambda x: x if x > 2 else 2)
srs3 = Series(['alpha', 'beta', 'gamma', 'delta'], index = ['a', 'b', 'c', 'd'])
print(srs3)
obj = {"alpha": 1, "beta": 2, "gamma": -1, "delta": -3}
srs3.map(obj)
srs4 = Series(obj)
print(srs4)
srs3.map(srs4)
srs1.map(lambda x: x if x > 2 else 2) # Works like apply
```
### `DataFrame`s
Many of the tricks that work with `Series` work with `DataFrame`s, but with some more complication.
```
df = DataFrame(np.arange(15).reshape(5, 3), columns=["AAA", "BBB", "CCC"])
print(df)
# Should get 0's, and CCC gets NaN because no match
df - df.loc[:,["AAA", "BBB"]]
df.mean()
df.std()
# This is known as standardization
(df - df.mean())/df.std()
```
Let's now look at vectorization
```
np.sqrt(df)
# trunc is a custom ufunc: does not give a DataFrame
trunc(df)
# Mixed data
df2 = DataFrame({"AAA": [1, 2, 3, 4], "BBB": [0, -9, 9, 3], "CCC": ["Bob", "Terry", "Matt", "Simon"]})
print(df2)
# Produces an error
np.sqrt(df2)
# Let's select JUST numeric data
# The select_dtypes() method selects columns based on their dtype
# np.number indicates numeric dtypes
# Here we select columns only with numeric data
df2.select_dtypes([np.number])
np.sqrt(df2.select_dtypes([np.number]))
```
A brief look at function application. Here we work with a function that computes the geometric mean, which is defined as:
$$\text{geometric mean} = \left(\prod_{i = 1}^n x_i\right)^{\frac{1}{n}}$$
```
# Define a function for the geometric mean
def geomean(srs):
return srs.prod() ** (1 / len(srs)) # prod method is product of all elements of srs
# Demo
geomean(Series([2, 3, 4]))
df.apply(geomean)
df.apply(geomean, axis='columns')
# Apply a truncation function to each element of df
df.applymap(lambda x: x if x > 3 else 3)
```
| github_jupyter |
# Generative Models
- Train on data from certain probability distribution
- To be able to generate new data within this probability distribution is must learn the features and characteristics that make up this distribution
- “What I cannot create, I do not understand.” — Richard Feynman
<img src="https://openai.com/assets/research/generative-models/gen-c994c9370597f62edbce64af321e7186c41e8fcf4d7503ea876f8a6bdf901135.svg">
<img src="https://openai.com/assets/research/generative-models/gencnn-afe135ff8d2725325a22455a488562b0e1cb7ac6a3f60b3cecb373fd043eb202.svg">
<img src="https://openai.com/assets/research/generative-models/learning-gan-ffc4c09e6079283f334b2485ae663a6587d937a45ebc1d8aeac23a67889a3cf5.gif">
# Extracting Features from the Generative Models
### Technique 1: Examine the effect of each dimention of the code
For example, in the images of 3D faces below we vary one continuous dimension of the code, keeping all others fixed. It's clear from the five provided examples (along each row) that the resulting dimensions in the code capture interpretable dimensions, and that the model has perhaps understood that there are camera angles, facial variations, etc., without having been told that these features exist and are important
### Elevation
<img src="https://openai.com/assets/research/generative-models/infogan-2-069a9ff24c4194a444ba286980a5f2693446c1d8f42c2dc240da05fe48e0378d.jpg">
### Lighting
<img src="https://openai.com/assets/research/generative-models/infogan-3-20e68c4ad01bd22874596ec9b799f76865e89db6768d412c6d9f6e26e37e6823.jpg">
### Technique 2: Decompose the "though vector" in a learned feature basis (<a href="http://gabgoh.github.io/ThoughtVectors/">Goh</a>)
<img style="float:left" src="http://gabgoh.github.io/ThoughtVectors/c2c.svg">
<img style="float:left" src="http://gabgoh.github.io/ThoughtVectors/c2r.svg">
<img style="float:left" src="http://gabgoh.github.io/ThoughtVectors/r2c.svg">
<img style="float:left" src="http://gabgoh.github.io/ThoughtVectors/r2r.svg">
### Technique 3: Use a deconvolution on each layer of the encoder to examine the features being learned

# My Current Proposal
James' astrocyte data is significantly simpler than the majority of data these techniques were designed to operate on. These may be overkill. I propose we start simple and add complexity as it is needed.
### Spatio-temporal Features
These are often extracted using two approaches.
1. They use recurrent networks to "remember" past activity. The effect this has on the data/network/features is quite unclear. In addition most techniques only use a delay of 1 meaning the impact of different "features" of the data at different times may not be captured best.
2. Few people have used 3D neural networks. Instead of passing a sinlge image through a 2D convolutional network, they pass 3D volumes through a 3D convolutional network. This has been shown to provide superior predicition/classification on videos. The features extracted are more "isolated"

### Short-term goal
I would like to explore the features of a 3D convolutional auto-encoder. I will start simple with only 1 hidden layer. I can examine the features using the 3 techniques outlined above. With my current understanding of the auto-encoders and our data I believe 1, maybe two layers should be sufficient to extract the features necessary. This will be because we are using 3D convolution. Depending on the results, I may use a variational auto-encoder to see if superior results are found.
1. Implement 3D convolution auto-encoder (Keras or TensorFlow)
2. Implement the 3 tools for extracting and visualizing the features extracted (Software packages available for 1)
| github_jupyter |
# ORE Jupyter Dashboard
This dashboard contains several modules to run ORE functionality and visualize results.
It assumes a standard installation of Anaconda Python 3, see https://www.continuum.io/downloads.
Some plots require more dependencies, see below.
## Launch ORE
Kick off a process in ORE as specified in Input/ore.xml
```
from OREAnalytics import *
print ("Loading parameters...")
params = Parameters()
print (" params is of type", type(params))
params.fromFile("Input/ore.xml")
print (" setup/asofdate = " + params.get("setup","asofDate"))
print ("Building OREApp...")
ore = OREApp(params)
print (" ore is of type", type(ore))
print ("Running ORE process...");
ore.buildMarket()
print ("ORE process finished");
```
## Query ORE App Members
Retrieve the market object stored in OREApp and initialized in the previous step when running the ORE process.
Then query the market object for some members (a discount and a forward curve) and evaluate discount factors and zero rates at some point in time in the future.
```
print ("Retrieve market object...");
market = ore.getMarket()
print (" retrieved market is of type", type(market))
asof = market.asofDate();
print (" retrieved market's asof date is", asof)
ccy = "EUR"
index = "EUR-EURIBOR-6M"
print ("Get term structures for ccy ", ccy, "and index", index);
discountCurve = market.discountCurve(ccy)
print (" discount curve is of type", type(discountCurve))
iborIndex = market.iborIndex(index)
print (" ibor index is of type", type(iborIndex))
forwardCurve = iborIndex.forwardingTermStructure()
print (" forward curve is of type", type(forwardCurve))
date = asof + 10*Years;
zeroRateDc = Actual365Fixed()
discount = discountCurve.discount(date)
zero = discountCurve.zeroRate(date, zeroRateDc, Continuous)
fwdDiscount = forwardCurve.discount(date)
fwdZero = forwardCurve.zeroRate(date, zeroRateDc, Continuous)
print (" 10y discount factor (discount curve) is", discount)
print (" 10y discout factor (forward curve) is", fwdDiscount)
print (" 10y zero rate (discount curve) is", zero)
print (" 10y zero rate (forward curve) is", fwdZero)
dc = ActualActual()
# date grid
dates = []
times = []
zeros1 = []
zeros2 = []
date = asof
previousDate = asof
for i in range (1,10*53):
date = date + Period(1, Weeks);
time = dc.yearFraction(asof, date)
dates.append(date)
times.append(time)
zero1 = discountCurve.forwardRate(previousDate, date, zeroRateDc, Continuous).rate()
zero2 = forwardCurve.forwardRate(previousDate, date, zeroRateDc, Continuous).rate()
zeros1.append(zero1)
zeros2.append(zero2)
previousDate = date
#print (date, time, zero1, zero2)
#print(times)
```
## Curve Plot
Plot the discount and forward curves above using Bloomberg's bqplot, see https://github.com/bloomberg/bqplot
```
import numpy as np
import bqplot as bq
from traitlets import link
from IPython.display import display
xs = bq.LinearScale()
ys = bq.LinearScale()
x = times
y = np.vstack((zeros1,zeros2))
line = bq.Lines(x=x, y=y, scales={'x': xs, 'y': ys}, display_legend=True, labels=['Discount Curve','Forward Curve'])
xax = bq.Axis(scale=xs, label='years', grid_lines='solid')
yax = bq.Axis(scale=ys, orientation='vertical', grid_lines='solid')
fig = bq.Figure(marks=[line], axes=[xax, yax], animation_duration=1000, legend_location='top-left')
display(fig)
```
| github_jupyter |
# Example1: Principal Component Analysis (PCA)
Principal Component Analysis (PCA) is a classic statistical technique for dimensionality reduction. It defines a mapping between the $d$-dimensional data-representation of a point $\boldsymbol{x}$ and its $k$-dimensional latent representation, $\boldsymbol{z}$. The latent representation is known as the scores, and the affine transformation is performed using the loading matrix $\boldsymbol{\beta}$, which has dimensions $k\times d$.
A simplified probabilistic view of PCA (Tipping & Bishop, 1999) is given below, which provides a pseudo-code a description of the generative model of a probabilistic PCA model.
<img src="https://raw.githubusercontent.com/PGM-Lab/ProbModelingDNNs/master/img/pca_pseudocode.png" alt="PCA-pseudocode" style="width: 700px;"/>
The present notebook experimentally illustrates the behavior of Probabilistic PCA as a feature reduction method on (a reduced version of) MNIST dataset. The model is implemented using the InferPy.
## Setting up the system
First, we will install and import the required packages as follows.
```
!pip install tensorflow==1.14.0
!pip install tensorflow-probability==0.7.0
!pip install keras
!pip install matplotlib
!pip install inferpy==1.2.0
!pip install setuptools
!pip install pandas
import tensorflow as tf
from tensorflow_probability import edward2 as ed
import numpy as np
import matplotlib.pyplot as plt
import inferpy as inf
from inferpy.data import mnist
import pandas as pd
import sys
import ssl
ssl._create_default_https_context = ssl._create_unverified_context
import warnings
warnings.filterwarnings("ignore")
# Global configuration
num_epochs = 3000
tf.reset_default_graph()
tf.set_random_seed(1234)
verbose = True
learning_rate = 0.01
```
## Data
For illustrating purposes, the MNIST dataset containg handwritten digits will be used. In particular we might obtain the data from the ``inferpy`` package (and hence from keras):
```
# number of observations (dataset size)
N = 1000
# digits considered
DIG = [0,1,2]
# load the data
(x_train, y_train), _ = mnist.load_data(num_instances=N, digits=DIG)
# plot the digits
mnist.plot_digits(x_train, grid=[5,5])
```
## Model definition
The implementation for the generative model for a PCA model (Algorithm 1) is defined below. The input parameters are: `k` is the latent dimension, `d` is the data-dimension and `N`the number of samples or data instances.
```
# Model constants
k, d = 2, np.shape(x_train)[-1]
@inf.probmodel
def pca(k,d):
w = inf.Normal(loc=tf.zeros([k,d]),
scale=1, name="w") # shape = [k,d]
w0 = inf.Normal(loc=tf.zeros([d]),
scale=1, name="w0") # shape = [d]
with inf.datamodel():
z = inf.Normal(tf.zeros([k]),1, name="z") # shape = [N,k]
x = inf.Normal( z @ w + w0, 1, name="x") # shape = [N,d]
print(pca)
```
This is a latent variable model (LVM) where the latent representation $\boldsymbol{z}$ is known as the scores, and the affine transformation is performed using the loading matrix $\boldsymbol{\beta}$.
## Inference
Variational inference is a deterministic technique that finds a tractable approximation to an intractable (posterior) distribution.
We will use $q$ to denote the approximation, and use $p$ to signify the true distribution (like $p(\boldsymbol{\beta},\boldsymbol{z}|\boldsymbol{x})$ in the example above).
More specifically, ${\cal Q}$ will denote a set of possible approximations $q$.
In practice, we define a generative model for sampling from $q(\boldsymbol{\beta},\boldsymbol{z} | \boldsymbol{\lambda}, \boldsymbol{\phi})$, where $\boldsymbol{\lambda}, \boldsymbol{\phi}$ are the variational parameters to optimise.
```
@inf.probmodel
def Q(k,d):
qw_loc = inf.Parameter(tf.zeros([k,d]), name="qw_loc")
qw_scale = tf.math.softplus(inf.Parameter(tf.ones([k,d]), name="qw_scale"))
qw = inf.Normal(qw_loc, qw_scale, name="w")
qw0_loc = inf.Parameter(tf.ones([d]), name="qw0_loc")
qw0_scale = tf.math.softplus(inf.Parameter(tf.ones([d]), name="qw0_scale"))
qw0 = inf.Normal(qw0_loc, qw0_scale, name="w0")
with inf.datamodel():
qz_loc = inf.Parameter(np.zeros([k]), name="qz_loc")
qz_scale = tf.math.softplus(inf.Parameter(tf.ones([k]), name="qz_scale"))
qz = inf.Normal(qz_loc, qz_scale, name="z")
print(Q)
```
Variational methods adjusts the parameters by maximizing the ELBO (Evidence LOwer Bound) denoted $\cal{L}$ and expressed as
$\cal{L}(\boldsymbol{\lambda},\boldsymbol{\phi}) = \mathbb{E}_q [\ln p(\boldsymbol{x}, \boldsymbol{z}, \boldsymbol{\beta})] - \mathbb{E}_q [\ln q(\boldsymbol{\beta},\boldsymbol{z}|\boldsymbol{\lambda},\boldsymbol{\phi})]$
In InferPy, this is transparent to the user: it is only required to create the instances of the P and Q models, the optimizer and inference method objects.
```
# create an instance of the P model and the Q model
m = pca(k,d)
q = Q(k,d)
# load the data
(x_train, y_train), _ = mnist.load_data(num_instances=N, digits=DIG)
optimizer = tf.train.AdamOptimizer(learning_rate)
VI = inf.inference.VI(q, optimizer=optimizer, epochs=2000)
```
Finally, the ELBO function is maximized.
```
m.fit({"x": x_train}, VI)
```
After the inference, we can plot the hidden representation:
```
post = {"z": m.posterior("z", data={"x": x_train}).sample()}
markers = ["x", "+", "o"]
colors = [plt.get_cmap("gist_rainbow")(0.05),
plt.get_cmap("gnuplot2")(0.08),
plt.get_cmap("gist_rainbow")(0.33)]
transp = [0.9, 0.9, 0.5]
fig = plt.figure()
for c in range(0, len(DIG)):
col = colors[c]
plt.scatter(post["z"][y_train == DIG[c], 0], post["z"][y_train == DIG[c], 1], color=col,
label=DIG[c], marker=markers[c], alpha=transp[c], s=60)
plt.legend()
```
## Test
For testing our model, we will generate samples of $\boldsymbol{x}$ given the infered posterior distributions.
```
x_gen = m.posterior_predictive('x', data=post).sample()
# plot the digits
mnist.plot_digits(x_gen, grid=[5,5])
```
| github_jupyter |
```
import json
import numpy as np
import tensorflow as tf
import collections
from sklearn.cross_validation import train_test_split
with open('ctexts.json','r') as fopen:
ctexts = json.loads(fopen.read())[:100]
with open('headlines.json','r') as fopen:
headlines = json.loads(fopen.read())[:100]
def build_dataset(words, n_words):
count = [['GO', 0], ['PAD', 1], ['EOS', 2], ['UNK', 3]]
count.extend(collections.Counter(words).most_common(n_words))
dictionary = dict()
for word, _ in count:
dictionary[word] = len(dictionary)
data = list()
unk_count = 0
for word in words:
index = dictionary.get(word, 0)
if index == 0:
unk_count += 1
data.append(index)
count[0][1] = unk_count
reversed_dictionary = dict(zip(dictionary.values(), dictionary.keys()))
return data, count, dictionary, reversed_dictionary
concat_from = ' '.join(ctexts).split()
vocabulary_size_from = len(list(set(concat_from)))
data_from, count_from, dictionary_from, rev_dictionary_from = build_dataset(concat_from, vocabulary_size_from)
print('vocab from size: %d'%(vocabulary_size_from))
print('Most common words', count_from[4:10])
print('Sample data', data_from[:10], [rev_dictionary_from[i] for i in data_from[:10]])
concat_to = ' '.join(headlines).split()
vocabulary_size_to = len(list(set(concat_to)))
data_to, count_to, dictionary_to, rev_dictionary_to = build_dataset(concat_to, vocabulary_size_to)
print('vocab to size: %d'%(vocabulary_size_to))
print('Most common words', count_to[4:10])
print('Sample data', data_to[:10], [rev_dictionary_to[i] for i in data_to[:10]])
for i in range(len(headlines)):
headlines[i] = headlines[i] + ' EOS'
headlines[0]
GO = dictionary_from['GO']
PAD = dictionary_from['PAD']
EOS = dictionary_from['EOS']
UNK = dictionary_from['UNK']
def str_idx(corpus, dic):
X = []
for i in corpus:
ints = []
for k in i.split():
try:
ints.append(dic[k])
except Exception as e:
print(e)
ints.append(UNK)
X.append(ints)
return X
X = str_idx(ctexts, dictionary_from)
Y = str_idx(headlines, dictionary_to)
train_X, test_X, train_Y, test_Y = train_test_split(X, Y, test_size = 0.05)
class Summarization:
def __init__(self, size_layer, num_layers, embedded_size,
from_dict_size, to_dict_size, batch_size,
dropout = 0.5, beam_width = 15):
def lstm_cell(reuse=False):
return tf.nn.rnn_cell.LSTMCell(size_layer, reuse=reuse)
self.X = tf.placeholder(tf.int32, [None, None])
self.Y = tf.placeholder(tf.int32, [None, None])
self.X_seq_len = tf.placeholder(tf.int32, [None])
self.Y_seq_len = tf.placeholder(tf.int32, [None])
# encoder
encoder_embeddings = tf.Variable(tf.random_uniform([from_dict_size, embedded_size], -1, 1))
encoder_embedded = tf.nn.embedding_lookup(encoder_embeddings, self.X)
encoder_cells = tf.nn.rnn_cell.MultiRNNCell([lstm_cell() for _ in range(num_layers)])
encoder_dropout = tf.contrib.rnn.DropoutWrapper(encoder_cells, output_keep_prob = 0.5)
self.encoder_out, self.encoder_state = tf.nn.dynamic_rnn(cell = encoder_dropout,
inputs = encoder_embedded,
sequence_length = self.X_seq_len,
dtype = tf.float32)
self.encoder_state = tuple(self.encoder_state[-1] for _ in range(num_layers))
main = tf.strided_slice(self.Y, [0, 0], [batch_size, -1], [1, 1])
decoder_input = tf.concat([tf.fill([batch_size, 1], GO), main], 1)
# decoder
decoder_embeddings = tf.Variable(tf.random_uniform([to_dict_size, embedded_size], -1, 1))
decoder_cells = tf.nn.rnn_cell.MultiRNNCell([lstm_cell() for _ in range(num_layers)])
dense_layer = tf.layers.Dense(to_dict_size)
training_helper = tf.contrib.seq2seq.ScheduledEmbeddingTrainingHelper(
inputs = tf.nn.embedding_lookup(decoder_embeddings, decoder_input),
sequence_length = self.Y_seq_len,
embedding = decoder_embeddings,
sampling_probability = 0.5,
time_major = False)
training_decoder = tf.contrib.seq2seq.BasicDecoder(
cell = decoder_cells,
helper = training_helper,
initial_state = self.encoder_state,
output_layer = dense_layer)
training_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = training_decoder,
impute_finished = True,
maximum_iterations = tf.reduce_max(self.Y_seq_len))
predicting_decoder = tf.contrib.seq2seq.BeamSearchDecoder(
cell = decoder_cells,
embedding = decoder_embeddings,
start_tokens = tf.tile(tf.constant([GO], dtype=tf.int32), [batch_size]),
end_token = EOS,
initial_state = tf.contrib.seq2seq.tile_batch(self.encoder_state, beam_width),
beam_width = beam_width,
output_layer = dense_layer,
length_penalty_weight = 0.0)
predicting_decoder_output, _, _ = tf.contrib.seq2seq.dynamic_decode(
decoder = predicting_decoder,
impute_finished = False,
maximum_iterations = tf.reduce_max(self.X_seq_len)//3)
self.training_logits = training_decoder_output.rnn_output
self.predicting_ids = predicting_decoder_output.predicted_ids[:, :, 0]
masks = tf.sequence_mask(self.Y_seq_len, tf.reduce_max(self.Y_seq_len), dtype=tf.float32)
self.cost = tf.contrib.seq2seq.sequence_loss(logits = self.training_logits,
targets = self.Y,
weights = masks)
self.optimizer = tf.train.AdamOptimizer().minimize(self.cost)
size_layer = 512
num_layers = 2
embedded_size = 128
batch_size = 16
epoch = 20
tf.reset_default_graph()
sess = tf.InteractiveSession()
model = Summarization(size_layer, num_layers, embedded_size, len(dictionary_from),
len(dictionary_to), batch_size)
sess.run(tf.global_variables_initializer())
def pad_sentence_batch(sentence_batch, pad_int, maxlen=500):
padded_seqs = []
seq_lens = []
max_sentence_len = min(max([len(sentence) for sentence in sentence_batch]),maxlen)
for sentence in sentence_batch:
sentence = sentence[:maxlen]
padded_seqs.append(sentence + [pad_int] * (max_sentence_len - len(sentence)))
seq_lens.append(len(sentence))
return padded_seqs, seq_lens
def check_accuracy(logits, Y):
acc = 0
for i in range(logits.shape[0]):
internal_acc = 0
count = 0
for k in range(len(Y[i])):
try:
if Y[i][k] == logits[i][k]:
internal_acc += 1
count += 1
if Y[i][k] == EOS:
break
except:
break
acc += (internal_acc / count)
return acc / logits.shape[0]
for i in range(epoch):
total_loss, total_accuracy = 0, 0
for k in range(0, (len(train_X) // batch_size) * batch_size, batch_size):
batch_x, seq_x = pad_sentence_batch(train_X[k: k+batch_size], PAD)
batch_y, seq_y = pad_sentence_batch(train_Y[k: k+batch_size], PAD)
predicted, loss, _ = sess.run([model.predicting_ids, model.cost, model.optimizer],
feed_dict={model.X:batch_x,
model.Y:batch_y,
model.X_seq_len:seq_x,
model.Y_seq_len:seq_y})
total_loss += loss
total_accuracy += check_accuracy(predicted,batch_y)
rand = np.random.randint(0, len(train_X)-batch_size)
batch_x, seq_x = pad_sentence_batch(train_X[rand:rand+batch_size], PAD)
batch_y, seq_y = pad_sentence_batch(train_Y[rand:rand+batch_size], PAD)
predicted, test_loss = sess.run([model.predicting_ids,model.cost],
feed_dict={model.X:batch_x,
model.Y:batch_y,
model.X_seq_len:seq_x,
model.Y_seq_len:seq_y})
print('epoch %d, train loss %f, valid loss %f'%(i+1,loss,test_loss))
print('expected output:',' '.join([rev_dictionary_to[n] for n in batch_y[0] if n not in [-1,0,1,2,3]]))
print('predicted output:',' '.join([rev_dictionary_to[n] for n in predicted[0] if n not in [-1,0,1,2,3]]))
total_loss /= (len(train_X) // batch_size)
total_accuracy /= (len(train_X) // batch_size)
print('epoch: %d, avg loss: %f, avg accuracy: %f'%(i+1, total_loss, total_accuracy),'\n')
```
| github_jupyter |
Let us go through a full worked example. Here we have a Tipsy SPH dataset. By general
inspection, we see that there are stars present in the dataset, since
there are fields with field type: `Stars` in the `ds.field_list`. Let's look
at the `derived_field_list` for all of the `Stars` fields.
```
import yt
import numpy as np
ds = yt.load("TipsyGalaxy/galaxy.00300")
for field in ds.derived_field_list:
if field[0] == 'Stars':
print (field)
```
We will filter these into young stars and old stars by masking on the ('Stars', 'creation_time') field.
In order to do this, we first make a function which applies our desired cut. This function must accept two arguments: `pfilter` and `data`. The first argument is a `ParticleFilter` object that contains metadata about the filter its self. The second argument is a yt data container.
Let's call "young" stars only those stars with ages less 5 million years. Since Tipsy assigns a very large `creation_time` for stars in the initial conditions, we need to also exclude stars with negative ages.
Conversely, let's define "old" stars as those stars formed dynamically in the simulation with ages greater than 5 Myr. We also include stars with negative ages, since these stars were included in the simulation initial conditions.
We make use of `pfilter.filtered_type` so that the filter definition will use the same particle type as the one specified in the call to `add_particle_filter` below. This makes the filter definition usable for arbitrary particle types. Since we're only filtering the `"Stars"` particle type in this example, we could have also replaced `pfilter.filtered_type` with `"Stars"` and gotten the same result.
```
def young_stars(pfilter, data):
age = data.ds.current_time - data[pfilter.filtered_type, "creation_time"]
filter = np.logical_and(age.in_units('Myr') <= 5, age >= 0)
return filter
def old_stars(pfilter, data):
age = data.ds.current_time - data[pfilter.filtered_type, "creation_time"]
filter = np.logical_or(age.in_units('Myr') >= 5, age < 0)
return filter
```
Now we define these as particle filters within the yt universe with the
`add_particle_filter()` function.
```
yt.add_particle_filter("young_stars", function=young_stars, filtered_type='Stars', requires=["creation_time"])
yt.add_particle_filter("old_stars", function=old_stars, filtered_type='Stars', requires=["creation_time"])
```
Let us now apply these filters specifically to our dataset.
Let's double check that it worked by looking at the derived_field_list for any new fields created by our filter.
```
ds.add_particle_filter('young_stars')
ds.add_particle_filter('old_stars')
for field in ds.derived_field_list:
if "young_stars" in field or "young_stars" in field[1]:
print (field)
```
We see all of the new `young_stars` fields as well as the 4 deposit fields. These deposit fields are `mesh` fields generated by depositing particle fields on the grid. Let's generate a couple of projections of where the young and old stars reside in this simulation by accessing some of these new fields.
```
p = yt.ProjectionPlot(ds, 'z', [('deposit', 'young_stars_cic'), ('deposit', 'old_stars_cic')], width=(40, 'kpc'), center='m')
p.set_figure_size(5)
p.show()
```
We see that young stars are concentrated in regions of active star formation, while old stars are more spatially extended.
| github_jupyter |
# Introduction to Basic Functionality of NTM
_**Finding Topics in Synthetic Document Data with the Neural Topic Model**_
---
---
# Contents
***
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Data](#Data)
1. [Train](#Train)
1. [Host](#Host)
1. [Extensions](#Extensions)
# Introduction
***
Amazon SageMaker NTM (Neural Topic Model) is an unsupervised learning algorithm that attempts to describe a set of observations as a mixture of distinct categories. NTM is most commonly used to discover a user-specified number of topics shared by documents within a text corpus. Here each observation is a document, the features are the presence (or occurrence count) of each word, and the categories are the topics. Since the method is unsupervised, the topics are not specified up front, and are not guaranteed to align with how a human may naturally categorize documents. The topics are learned as a probability distribution over the words that occur in each document. Each document, in turn, is described as a mixture of topics.
In this notebook we will use the Amazon SageMaker NTM algorithm to train a model on some example synthetic data. We will then use this model to classify (perform inference on) the data. The main goals of this notebook are to,
* learn how to obtain and store data for use in Amazon SageMaker,
* create an AWS SageMaker training job on a data set to produce a NTM model,
* use the model to perform inference with an Amazon SageMaker endpoint.
# Setup
***
_This notebook was created and tested on an ml.m4xlarge notebook instance._
Let's start by specifying:
- The S3 bucket and prefix that you want to use for training and model data. This should be within the same region as the Notebook Instance, training, and hosting.
- The IAM role arn used to give training and hosting access to your data. See the documentation for how to create these. Note, if more than one role is required for notebook instances, training, and/or hosting, please replace the boto regexp with a the appropriate full IAM role arn string(s).
```
# Define IAM role
import sagemaker
import boto3
import re
from sagemaker import get_execution_role
sess = sagemaker.Session()
bucket=sess.default_bucket()
role = get_execution_role()
```
Next we'll import the libraries we'll need throughout the remainder of the notebook.
```
import numpy as np
from generate_example_data import generate_griffiths_data, plot_topic_data
import io
import os
import time
import json
import sys
import pandas as pd
import matplotlib.pyplot as plt
from IPython.display import display
import scipy
import sagemaker.amazon.common as smac
from sagemaker.predictor import csv_serializer, json_deserializer
```
# Data
***
We generate some example synthetic document data. For the purposes of this notebook we will omit the details of this process. All we need to know is that each piece of data, commonly called a "document", is a vector of integers representing "word counts" within the document. In this particular example there are a total of 25 words in the "vocabulary".
```
# generate the sample data
num_documents = 5000
num_topics = 5
vocabulary_size = 25
known_alpha, known_beta, documents, topic_mixtures = generate_griffiths_data(
num_documents=num_documents, num_topics=num_topics, vocabulary_size=vocabulary_size)
# separate the generated data into training and tests subsets
num_documents_training = int(0.8*num_documents)
num_documents_test = num_documents - num_documents_training
documents_training = documents[:num_documents_training]
documents_test = documents[num_documents_training:]
topic_mixtures_training = topic_mixtures[:num_documents_training]
topic_mixtures_test = topic_mixtures[num_documents_training:]
data_training = (documents_training, np.zeros(num_documents_training))
data_test = (documents_test, np.zeros(num_documents_test))
```
## Inspect Example Data
*What does the example data actually look like?* Below we print an example document as well as its corresponding *known* topic mixture. Later, when we perform inference on the training data set we will compare the inferred topic mixture to this known one.
As we can see, each document is a vector of word counts from the 25-word vocabulary
```
print('First training document = {}'.format(documents[0]))
print('\nVocabulary size = {}'.format(vocabulary_size))
np.set_printoptions(precision=4, suppress=True)
print('Known topic mixture of first training document = {}'.format(topic_mixtures_training[0]))
print('\nNumber of topics = {}'.format(num_topics))
```
Because we are visual creatures, let's try plotting the documents. In the below plots, each pixel of a document represents a word. The greyscale intensity is a measure of how frequently that word occurs. Below we plot the first tes documents of the training set reshaped into 5x5 pixel grids.
```
%matplotlib inline
fig = plot_topic_data(documents_training[:10], nrows=2, ncols=5, cmap='gray_r', with_colorbar=False)
fig.suptitle('Example Documents')
fig.set_dpi(160)
```
## Store Data on S3
A SageMaker training job needs access to training data stored in an S3 bucket. Although training can accept data of various formats recordIO wrapped protobuf is most performant.
_Note, since NTM is an unsupervised learning algorithm, we simple put 0 in for all label values._
```
buf = io.BytesIO()
smac.write_numpy_to_dense_tensor(buf, data_training[0].astype('float32'))
buf.seek(0)
key = 'ntm.data'
boto3.resource('s3').Bucket(bucket).Object(os.path.join(prefix, 'train', key)).upload_fileobj(buf)
s3_train_data = 's3://{}/{}/train/{}'.format(bucket, prefix, key)
```
# Training
***
Once the data is preprocessed and available in a recommended format the next step is to train our model on the data. There are number of parameters required by the NTM algorithm to configure the model and define the computational environment in which training will take place. The first of these is to point to a container image which holds the algorithms training and hosting code.
```
from sagemaker.amazon.amazon_estimator import get_image_uri
container = get_image_uri(boto3.Session().region_name, 'ntm')
```
An NTM model uses the following hyperparameters:
* **`num_topics`** - The number of topics or categories in the NTM model. This has been pre-defined in our synthetic data to be 5.
* **`feature_dim`** - The size of the *"vocabulary"*, in topic modeling parlance. In this case, this has been set to 25 by `generate_griffiths_data()`.
In addition to these NTM model hyperparameters, we provide additional parameters defining things like the EC2 instance type on which training will run, the S3 bucket containing the data, and the AWS access role.
```
sess = sagemaker.Session()
ntm = sagemaker.estimator.Estimator(container,
role,
train_instance_count=1,
train_instance_type='ml.c4.xlarge',
output_path='s3://{}/{}/output'.format(bucket, prefix),
sagemaker_session=sess)
ntm.set_hyperparameters(num_topics=num_topics,
feature_dim=vocabulary_size)
ntm.fit({'train': s3_train_data})
```
# Inference
***
A trained model does nothing on its own. We now want to use the model to perform inference. For this example, that means predicting the topic mixture representing a given document.
This is simplified by the deploy function provided by the Amazon SageMaker Python SDK.
```
ntm_predictor = ntm.deploy(initial_instance_count=1,
instance_type='ml.m4.xlarge')
```
## Perform Inference
With this real-time endpoint at our fingertips we can finally perform inference on our training and test data. We should first discuss the meaning of the SageMaker NTM inference output.
For each document we wish to compute its corresponding `topic_weights`. Each set of topic weights is a probability distribution over the number of topics, which is 5 in this example. Of the 5 topics discovered during NTM training each element of the topic weights is the proportion to which the input document is represented by the corresponding topic.
For example, if the topic weights of an input document $\mathbf{w}$ is,
$$\theta = \left[ 0.3, 0.2, 0, 0.5, 0 \right]$$
then $\mathbf{w}$ is 30% generated from Topic #1, 20% from Topic #2, and 50% from Topic #4. Below, we compute the topic mixtures for the first ten traning documents.
First, we setup our serializes and deserializers which allow us to convert NumPy arrays to CSV strings which we can pass into our HTTP POST request to our hosted endpoint.
```
ntm_predictor.content_type = 'text/csv'
ntm_predictor.serializer = csv_serializer
ntm_predictor.deserializer = json_deserializer
```
Now, let's check results for a small sample of records.
```
results = ntm_predictor.predict(documents_training[:10])
print(results)
```
We can see the output format of SageMaker NTM inference endpoint is a Python dictionary with the following format.
```
{
'predictions': [
{'topic_weights': [ ... ] },
{'topic_weights': [ ... ] },
{'topic_weights': [ ... ] },
...
]
}
```
We extract the topic weights, themselves, corresponding to each of the input documents.
```
predictions = np.array([prediction['topic_weights'] for prediction in results['predictions']])
print(predictions)
```
If you decide to compare these results to the known topic weights generated above keep in mind that SageMaker NTM discovers topics in no particular order. That is, the approximate topic mixtures computed above may be (approximate) permutations of the known topic mixtures corresponding to the same documents.
```
print(topic_mixtures_training[0]) # known topic mixture
print(predictions[0]) # computed topic mixture
```
With that said, let's look at how our learned topic weights map to known topic mixtures for the entire training set. Because NTM inherently creates a soft clustering (meaning that documents can sometimes belong partially to multiple topics), we'll evaluate correlation of topic weights. This gives us a more relevant picture than just selecting the single topic for each document that happens to have the highest probability.
To do this, we'll first need to generate predictions for all of our training data. Because our endpoint has a ~6MB per POST request limit, let's break the training data up into mini-batches and loop over them, creating a full dataset of predictions.
```
def predict_batches(data, rows=1000):
split_array = np.array_split(data, int(data.shape[0] / float(rows) + 1))
predictions = []
for array in split_array:
results = ntm_predictor.predict(array)
predictions += [r['topic_weights'] for r in results['predictions']]
return np.array(predictions)
predictions = predict_batches(documents_training)
```
Now we'll look at how the actual and predicted topics correlate.
```
data = pd.DataFrame(np.concatenate([topic_mixtures_training, predictions], axis=1),
columns=['actual_{}'.format(i) for i in range(5)] + ['predictions_{}'.format(i) for i in range(5)])
display(data.corr())
pd.plotting.scatter_matrix(pd.DataFrame(np.concatenate([topic_mixtures_training, predictions], axis=1)), figsize=(12, 12))
plt.show()
```
As we can see:
- The upper left quadrant of 5 * 5 cells illustrates that the data are synthetic as the correlations are all slightly negative, but too perfectly triangular to occur naturally.
- The upper right quadrant, which tells us about our model fit, shows some similarities, with many correlations having very near triangular shape, and negative correlations of a similar magnitude.
- Notice, actual topic #2 maps to predicted topic #2. Similarly actual topic #3 maps to predicted topic #3, and #4 to #4. However, there's a slight bit of uncertainty in topics #0 and #1. Actual topic #0 appears to map to predicted topic #1, but actual topic #1 also correlates most highly with predicted topic #1. This is not unexpected given that we're working with manufactured data and unsupervised algorithms. The important part is that NTM is picking up aggregate structure well and with increased tuning of hyperparameters may fit the data even more closely.
_Note, specific results may differ due to randomized steps in the data generation and algorithm, but the general story should remain unchanged._
## Stop / Close the Endpoint
Finally, we should delete the endpoint before we close the notebook.
To restart the endpoint you can follow the code above using the same `endpoint_name` we created or you can navigate to the "Endpoints" tab in the SageMaker console, select the endpoint with the name stored in the variable `endpoint_name`, and select "Delete" from the "Actions" dropdown menu.
```
sagemaker.Session().delete_endpoint(ntm_predictor.endpoint)
```
# Extensions
***
This notebook was a basic introduction to the NTM . It was applied on a synthetic dataset merely to show how the algorithm functions and represents data. Obvious extensions would be to train the algorithm utilizing real data. We skipped the important step of qualitatively evaluating the outputs of NTM. Because it is an unsupervised model, we want our topics to make sense. There is a great deal of subjectivity involved in this, and whether or not NTM is more suitable than another topic modeling algorithm like Amazon SageMaker LDA will depend on your use case.
| github_jupyter |
```
#np.pad(np.array([[1,2,3,4]]).reshape(-1,1), [[5, 0], [0, 0]],'constant')
'''import librosa
import numpy as np
energy = librosa.feature.rmse(np.array([1,0,0,4],dtype=float), frame_length=2,hop_length=1,center=True)
print(energy)
frames = np.nonzero(energy > 0)
print(librosa.core.frames_to_samples(frames,hop_length=1))
indices = librosa.core.frames_to_samples(frames,hop_length=1)[1]'''
'''import re
FILE_PATTERN = r'p([0-9]+)_([0-9]+)\.wav'
id_reg_exp = re.compile(FILE_PATTERN)
ids = id_reg_exp.findall('./VCTK-Corpus/wav48/p257/p257_390.wav')
print(ids[0][0])'''
"""Training script for the WaveNet network on the VCTK corpus.
This script trains a network with the WaveNet using data from the VCTK corpus,
which can be freely downloaded at the following site (~10 GB):
http://homepages.inf.ed.ac.uk/jyamagis/page3/page58/page58.html
"""
from __future__ import print_function
import argparse
from datetime import datetime
import json
import os
import sys
import time
import tensorflow as tf
from tensorflow.python.client import timeline
from wavenet import WaveNetModel, AudioReader, optimizer_factory
BATCH_SIZE = 1
DATA_DIRECTORY = './VCTK-Corpus'
LOGDIR_ROOT = './logdir'
CHECKPOINT_EVERY = 50
NUM_STEPS = int(1e5)
LEARNING_RATE = 1e-3
WAVENET_PARAMS = './wavenet_params.json'
STARTED_DATESTRING = "{0:%Y-%m-%dT%H-%M-%S}".format(datetime.now())
SAMPLE_SIZE = 100000
L2_REGULARIZATION_STRENGTH = 0
#SILENCE_THRESHOLD = 0.3
SILENCE_THRESHOLD = 0
EPSILON = 0.001
MOMENTUM = 0.9
MAX_TO_KEEP = 5
METADATA = False
def get_arguments():
def _str_to_bool(s):
"""Convert string to bool (in argparse context)."""
if s.lower() not in ['true', 'false']:
raise ValueError('Argument needs to be a '
'boolean, got {}'.format(s))
return {'true': True, 'false': False}[s.lower()]
parser = argparse.ArgumentParser(description='WaveNet example network')
parser.add_argument('--batch_size', type=int, default=BATCH_SIZE,
help='How many wav files to process at once. Default: ' + str(BATCH_SIZE) + '.')
parser.add_argument('--data_dir', type=str, default=DATA_DIRECTORY,
help='The directory containing the VCTK corpus.')
parser.add_argument('--store_metadata', type=bool, default=METADATA,
help='Whether to store advanced debugging information '
'(execution time, memory consumption) for use with '
'TensorBoard. Default: ' + str(METADATA) + '.')
parser.add_argument('--logdir', type=str, default=None,
help='Directory in which to store the logging '
'information for TensorBoard. '
'If the model already exists, it will restore '
'the state and will continue training. '
'Cannot use with --logdir_root and --restore_from.')
parser.add_argument('--logdir_root', type=str, default=None,
help='Root directory to place the logging '
'output and generated model. These are stored '
'under the dated subdirectory of --logdir_root. '
'Cannot use with --logdir.')
parser.add_argument('--restore_from', type=str, default=None,
help='Directory in which to restore the model from. '
'This creates the new model under the dated directory '
'in --logdir_root. '
'Cannot use with --logdir.')
parser.add_argument('--checkpoint_every', type=int,
default=CHECKPOINT_EVERY,
help='How many steps to save each checkpoint after. Default: ' + str(CHECKPOINT_EVERY) + '.')
parser.add_argument('--num_steps', type=int, default=NUM_STEPS,
help='Number of training steps. Default: ' + str(NUM_STEPS) + '.')
parser.add_argument('--learning_rate', type=float, default=LEARNING_RATE,
help='Learning rate for training. Default: ' + str(LEARNING_RATE) + '.')
parser.add_argument('--wavenet_params', type=str, default=WAVENET_PARAMS,
help='JSON file with the network parameters. Default: ' + WAVENET_PARAMS + '.')
parser.add_argument('--sample_size', type=int, default=SAMPLE_SIZE,
help='Concatenate and cut audio samples to this many '
'samples. Default: ' + str(SAMPLE_SIZE) + '.')
parser.add_argument('--l2_regularization_strength', type=float,
default=L2_REGULARIZATION_STRENGTH,
help='Coefficient in the L2 regularization. '
'Default: False')
parser.add_argument('--silence_threshold', type=float,
default=SILENCE_THRESHOLD,
help='Volume threshold below which to trim the start '
'and the end from the training set samples. Default: ' + str(SILENCE_THRESHOLD) + '.')
parser.add_argument('--optimizer', type=str, default='adam',
choices=optimizer_factory.keys(),
help='Select the optimizer specified by this option. Default: adam.')
parser.add_argument('--momentum', type=float,
default=MOMENTUM, help='Specify the momentum to be '
'used by sgd or rmsprop optimizer. Ignored by the '
'adam optimizer. Default: ' + str(MOMENTUM) + '.')
parser.add_argument('--histograms', type=_str_to_bool, default=False,
help='Whether to store histogram summaries. Default: False')
parser.add_argument('--gc_channels', type=int, default=None,
help='Number of global condition channels. Default: None. Expecting: Int')
parser.add_argument('--max_checkpoints', type=int, default=MAX_TO_KEEP,
help='Maximum amount of checkpoints that will be kept alive. Default: '
+ str(MAX_TO_KEEP) + '.')
return parser.parse_args([])
def save(saver, sess, logdir, step):
model_name = 'model.ckpt'
checkpoint_path = os.path.join(logdir, model_name)
print('Storing checkpoint to {} ...'.format(logdir), end="")
sys.stdout.flush()
if not os.path.exists(logdir):
os.makedirs(logdir)
saver.save(sess, checkpoint_path, global_step=step)
print(' Done.')
def load(saver, sess, logdir):
print("Trying to restore saved checkpoints from {} ...".format(logdir),
end="")
ckpt = tf.train.get_checkpoint_state(logdir)
if ckpt:
print(" Checkpoint found: {}".format(ckpt.model_checkpoint_path))
global_step = int(ckpt.model_checkpoint_path
.split('/')[-1]
.split('-')[-1])
print(" Global step was: {}".format(global_step))
print(" Restoring...", end="")
saver.restore(sess, ckpt.model_checkpoint_path)
print(" Done.")
return global_step
else:
print(" No checkpoint found.")
return None
def get_default_logdir(logdir_root):
logdir = os.path.join(logdir_root, 'train', STARTED_DATESTRING)
return logdir
def validate_directories(args):
"""Validate and arrange directory related arguments."""
# Validation
if args.logdir and args.logdir_root:
raise ValueError("--logdir and --logdir_root cannot be "
"specified at the same time.")
if args.logdir and args.restore_from:
raise ValueError(
"--logdir and --restore_from cannot be specified at the same "
"time. This is to keep your previous model from unexpected "
"overwrites.\n"
"Use --logdir_root to specify the root of the directory which "
"will be automatically created with current date and time, or use "
"only --logdir to just continue the training from the last "
"checkpoint.")
# Arrangement
logdir_root = args.logdir_root
if logdir_root is None:
logdir_root = LOGDIR_ROOT
logdir = args.logdir
if logdir is None:
logdir = get_default_logdir(logdir_root)
print('Using default logdir: {}'.format(logdir))
restore_from = args.restore_from
if restore_from is None:
# args.logdir and args.restore_from are exclusive,
# so it is guaranteed the logdir here is newly created.
restore_from = logdir
return {
'logdir': logdir,
'logdir_root': args.logdir_root,
'restore_from': restore_from
}
args = get_arguments()
try:
directories = validate_directories(args)
except ValueError as e:
print("Some arguments are wrong:")
print(str(e))
logdir = directories['logdir']
restore_from = directories['restore_from']
# Even if we restored the model, we will treat it as new training
# if the trained model is written into an arbitrary location.
is_overwritten_training = logdir != restore_from
with open(args.wavenet_params, 'r') as f:
wavenet_params = json.load(f)
# Create coordinator.
coord = tf.train.Coordinator()
# Load raw waveform from VCTK corpus.
with tf.name_scope('create_inputs'):
# Allow silence trimming to be skipped by specifying a threshold near
# zero.
silence_threshold = args.silence_threshold if args.silence_threshold > \
EPSILON else None
gc_enabled = args.gc_channels is not None
reader = AudioReader(
args.data_dir,
coord,
sample_rate=wavenet_params['sample_rate'], #"sample_rate": 16000,
gc_enabled=gc_enabled,
receptive_field=WaveNetModel.calculate_receptive_field(wavenet_params["filter_width"],
wavenet_params["dilations"],
wavenet_params["scalar_input"],
wavenet_params["initial_filter_width"]),
sample_size=args.sample_size, #SAMPLE_SIZE = 100000
silence_threshold=silence_threshold)
audio_batch = reader.dequeue(args.batch_size) #BATCH_SIZE = 1
if gc_enabled:
gc_id_batch = reader.dequeue_gc(args.batch_size)
else:
gc_id_batch = None
# Create network.
net = WaveNetModel(
batch_size=args.batch_size,
dilations=wavenet_params["dilations"],
filter_width=wavenet_params["filter_width"],
residual_channels=wavenet_params["residual_channels"],
dilation_channels=wavenet_params["dilation_channels"],
skip_channels=wavenet_params["skip_channels"],
quantization_channels=wavenet_params["quantization_channels"],
use_biases=wavenet_params["use_biases"],
scalar_input=wavenet_params["scalar_input"],
initial_filter_width=wavenet_params["initial_filter_width"],
histograms=args.histograms,
global_condition_channels=args.gc_channels,
global_condition_cardinality=reader.gc_category_cardinality)
if args.l2_regularization_strength == 0:
args.l2_regularization_strength = None
#print('audio_batch',audio_batch)
loss = net.loss(input_batch=audio_batch,
global_condition_batch=gc_id_batch,
l2_regularization_strength=args.l2_regularization_strength)
optimizer = optimizer_factory[args.optimizer](
learning_rate=args.learning_rate,
momentum=args.momentum)
trainable = tf.trainable_variables()
optim = optimizer.minimize(loss, var_list=trainable)
# Set up logging for TensorBoard.
writer = tf.summary.FileWriter(logdir)
writer.add_graph(tf.get_default_graph())
run_metadata = tf.RunMetadata()
summaries = tf.summary.merge_all()
# Set up session
sess = tf.Session(config=tf.ConfigProto(log_device_placement=False))
init = tf.global_variables_initializer()
sess.run(init)
# Saver for storing checkpoints of the model.
saver = tf.train.Saver(var_list=tf.trainable_variables(), max_to_keep=args.max_checkpoints)
try:
saved_global_step = load(saver, sess, restore_from)
if is_overwritten_training or saved_global_step is None:
# The first training step will be saved_global_step + 1,
# therefore we put -1 here for new or overwritten trainings.
saved_global_step = -1
except:
print("Something went wrong while restoring checkpoint. "
"We will terminate training to avoid accidentally overwriting "
"the previous model.")
raise
threads = tf.train.start_queue_runners(sess=sess, coord=coord)
reader.start_threads(sess)
step = None
last_saved_step = saved_global_step
try:
for step in range(saved_global_step + 1, args.num_steps):
start_time = time.time()
if args.store_metadata and step % 50 == 0:
# Slow run that stores extra information for debugging.
print('Storing metadata')
run_options = tf.RunOptions(
trace_level=tf.RunOptions.FULL_TRACE)
summary, loss_value, _ = sess.run(
[summaries, loss, optim],
options=run_options,
run_metadata=run_metadata)
writer.add_summary(summary, step)
writer.add_run_metadata(run_metadata,
'step_{:04d}'.format(step))
tl = timeline.Timeline(run_metadata.step_stats)
timeline_path = os.path.join(logdir, 'timeline.trace')
with open(timeline_path, 'w') as f:
f.write(tl.generate_chrome_trace_format(show_memory=True))
else:
summary, loss_value, _ = sess.run([summaries, loss, optim])
writer.add_summary(summary, step)
duration = time.time() - start_time
print('step {:d} - loss = {:.3f}, ({:.3f} sec/step)'
.format(step, loss_value, duration))
if step % args.checkpoint_every == 0:
save(saver, sess, logdir, step)
last_saved_step = step
except KeyboardInterrupt:
# Introduce a line break after ^C is displayed so save message
# is on its own line.
print()
finally:
if step > last_saved_step:
save(saver, sess, logdir, step)
coord.request_stop()
coord.join(threads)
```
| github_jupyter |
```
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.manifold import TSNE
import warnings
warnings.filterwarnings('ignore')
plt.rcParams['figure.figsize'] = [10, 5]
def calStability(raw, task_order, method, normalise=False, raw_acc=False, _offline=False):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
target = rows[rows["method"]==m]
_m = target[target["task_index"]==1][["accuracy", "no_of_test", "no_of_correct_prediction"]]
if m=="offline":
if _offline:
if raw_acc:
val = list(_m["accuracy"])
else:
val = float((_m["accuracy"]).mean())
values[m].append(val)
continue
if normalise:
_ideal = offline[offline["task_index"]==1]["accuracy"]
val = float((_m["accuracy"]/float(_ideal)).sum()/len(_m))
values[m].append(val)
elif raw_acc:
val = (_m["accuracy"])
values[m].append(list(val))
else:
val = float((_m["accuracy"]).sum()/len(_m))
values[m].append(val)
return values
def calPlasticity(raw, task_order, method, normalise=False, raw_acc=False, _offline=False):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
_sum = 0.0
if raw_acc:
_sum = []
target = rows[rows["method"]==m]
train_session = target["train_session"].unique()
if m=="offline":
if _offline:
if raw_acc:
val = list(target[target["train_session"]==target["task_index"]]["accuracy"])
else:
val = target[target["train_session"]==target["task_index"]]["accuracy"].mean()
values[m].append(val)
continue
target["train_session"] = pd.to_numeric(target["train_session"], errors='coerce')
_m = target[target["train_session"]==(target["task_index"])][["accuracy", "no_of_test", "no_of_correct_prediction"]]
if normalise:
_ideal = offline["accuracy"]
val = _m["accuracy"].div(_ideal.values, axis=0).sum()/len(_m)
values[m].append(val)
elif raw_acc:
val = (_m["accuracy"])
values[m].append(list(val))
else:
val = float((_m["accuracy"]).sum()/len(_m))
values[m].append(val)
return values
def calOverallAcc(raw, task_order, method, normalise=False, raw_acc=False):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
if m=="offline":
continue
_sum = 0.0
if raw_acc:
_sum = []
target = rows[rows["method"]==m]
task_index = target["task_index"].unique()
train_session = target["train_session"].unique()
_m = target[target["train_session"]==str(len(task_index))]
for t in task_index:
t = int(t)
_m1 = _m[_m["task_index"]==t]["accuracy"]
assert len(_m1)==1
if normalise:
_ideal = offline[offline["task_index"]==t]["accuracy"]
_sum += float(_m1)/float(_ideal)
elif raw_acc:
_sum.append(float(_m1))
else:
_sum += float(_m1)
if raw_acc:
values[m].append(_sum)
continue
if len(train_session)==0:
values[m].append(np.nan)
else:
val = _sum/len(train_session)
values[m].append(val)
return values
# Overall accuracy at each step
def calOverallAccEach(raw, task_order, method, normalise=False, raw_acc=False, _offline=False):
values = {k:[] for k in method}
for t in task_order:
rows = raw[raw["task_order"]==t]
offline = rows[rows["method"]=="offline"]
for m in method:
if m=="offline" and (not _offline):
continue
_sum = 0.0
if raw_acc:
_sum = []
target = rows[rows["method"]==m]
task_index = target["task_index"].unique()
train_session = target["train_session"].unique()
target["train_session"] = pd.to_numeric(target["train_session"], errors='coerce')
val = []
for s in train_session:
s = int(s)
_m = target[target["train_session"]==(s)]
_sum = _m["accuracy"].sum()/len(_m)
val.append(_sum)
values[m].append(val)
return values
all_values = {}
for d in ["CASAS"]:
dataset = d
folder = "../../Results/"+dataset+"/exp_no_of_hidden/"
_folder = "../../Results/"+dataset+"/exp_offline_acc/"
raw = pd.read_csv(folder+"results.txt")
_raw = pd.read_csv(_folder+"results.txt")
raw.columns = [c.strip() for c in raw.columns]
_raw.columns = [c.strip() for c in raw.columns]
cmd = raw["cmd"].unique()
task_order = raw["task_order"].unique()
method = raw["method"].unique()
stability = []
plasticity = []
overallAcc = []
for c in cmd:
target = raw[raw["cmd"]==c]
m = calStability(target, task_order, method, raw_acc=True)
stability.append(m)
m = calPlasticity(target, task_order, method, raw_acc=True)
plasticity.append(m)
m = calOverallAccEach(target, task_order, method, raw_acc=True)
overallAcc.append(m)
print(d, "DONE")
all_values[d] = (stability, plasticity, overallAcc)
```
# Continual learning
```
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from scipy import stats
def plotline(values, label, x=[], models=None, legends=None):
plt.rcParams['figure.figsize'] = [10, 5]
plt.rcParams.update({'font.size': 20})
m = []
merr = []
if models is None:
models = ["mp-gan", "mp-wgan", "sg-cgan", "sg-cwgan"]
if legends is None:
legends = models
numbers = []
for model in models:
nTaskOrder = len(values[model])
nTask = len(values[model][0])
_pivot = [[] for i in range(nTask)]
for i in range(nTaskOrder):
for j in range(nTask):
_pivot[j].append(values[model][i][j])
avg = [(np.nanmean(v), stats.sem(v, nan_policy="omit")) for v in _pivot]
numbers.append(avg)
fig, ax = plt.subplots()
patterns = [ "-s" , "-o" , "-x" , "-D" , "-+" , "-*", "-2" ]
for i, model in enumerate(models):
mval = [v for (v, _) in numbers[i]]
merr = [e for (_, e) in numbers[i]]
print(x, model, mval)
ax.errorbar(x, mval, yerr=merr, fmt=patterns[i])
mx = [v+e for (v, e) in numbers[i]]
mn = [v-e for (v, e) in numbers[i]]
# ax.fill_between(x, mn, mx, alpha=0.2)
ax.set_ylim(ymin=-0.1, ymax=1.1)
ax.set_title(label)
ax.set_xticks(x)
ax.set_xticklabels(x)
ax.legend(legends, prop={'size': 20}, loc=3, bbox_to_anchor=(1, 0.4))
# ax.legend(prop={'size': 20}, bbox_to_anchor=(1.05, 1), loc=0, borderaxespad=0.)
fig.tight_layout()
plt.show()
# select the best model (best GAN and best hidden units) => "sg-cgan" with highest number of neuron
def selectModel(db, model, cmd):
(stability, plasticity, overallAcc) = all_values[db]
models = [model, "exact", "none"]
legends = ["HAR-GAN", "Exact Reply", "None"]
x = [1,2,3,4,5]
if db=="HouseA":
x = [1,2,3]
# stability[cmd]["batch"] = []
# plasticity[cmd]["batch"] = []
# overallAcc[cmd]["batch"] = []
# for i in range(len(stability[cmd]["offline"])):
# v = stability[cmd]["offline"][i]
# stability[cmd]["batch"].append([v[-1] for i in v])
# v = plasticity[cmd]["offline"][i]
# plasticity[cmd]["batch"].append([v[-1] for i in v])
# v = overallAcc[cmd]["offline"][i]
# overallAcc[cmd]["batch"].append([v[-1] for i in v])
plotline(stability[cmd], "Stability of the model", x=x, models=models, legends=legends)
plotline(plasticity[cmd], "Plasticity of the model", x=x, models=models, legends=legends)
plotline(overallAcc[cmd], "Overall performance", x=x, models=models, legends=legends)
selectModel("CASAS", "sg-cgan", 3)
task_orders = [
["R2_dinner", "R2_lunch", "R2_sleeppppp", "R1_sleep", "R2_toilet", "R1_work_dining_room_table", "R1_work_computer", "R2_watch_TV", "R1_toilet", "R2_work_computer"],
["R2_sleeppppp", "R2_toilet", "R2_lunch", "R2_dinner", "R2_watch_TV", "R1_work_dining_room_table", "R2_work_computer", "R1_work_computer", "R1_toilet", "R1_sleep"],
["R1_toilet", "R2_lunch", "R1_work_computer", "R2_dinner", "R2_work_computer", "R1_work_dining_room_table", "R2_toilet", "R2_watch_TV", "R2_sleeppppp", "R1_sleep"],
["R1_toilet", "R2_lunch", "R2_sleeppppp", "R1_work_computer", "R2_dinner", "R2_watch_TV", "R2_toilet", "R1_sleep", "R1_work_dining_room_table", "R2_work_computer"],
["R2_dinner", "R1_work_dining_room_table", "R2_watch_TV", "R1_work_computer", "R2_work_computer", "R1_toilet", "R1_sleep", "R2_sleeppppp", "R2_lunch", "R2_toilet"],
["R2_watch_TV", "R1_work_computer", "R2_dinner", "R1_toilet", "R2_toilet", "R2_work_computer", "R2_sleeppppp", "R2_lunch", "R1_work_dining_room_table", "R1_sleep"],
["R2_dinner", "R2_sleeppppp", "R2_watch_TV", "R1_sleep", "R2_toilet", "R1_work_dining_room_table", "R1_work_computer", "R2_lunch", "R1_toilet", "R2_work_computer"],
["R1_work_computer", "R2_work_computer", "R1_sleep", "R2_toilet", "R2_lunch", "R2_dinner", "R1_toilet", "R2_sleeppppp", "R2_watch_TV", "R1_work_dining_room_table"],
["R1_sleep", "R1_work_dining_room_table", "R1_work_computer", "R1_toilet", "R2_sleeppppp", "R2_toilet", "R2_watch_TV", "R2_dinner", "R2_work_computer", "R2_lunch"],
["R1_work_dining_room_table", "R1_toilet", "R2_sleeppppp", "R2_lunch", "R2_work_computer", "R2_dinner", "R1_sleep", "R2_toilet", "R2_watch_TV", "R1_work_computer"],
]
(stability, plasticity, overallAcc) = all_values["CASAS"]
values = plasticity[3]["sg-cgan"]
print()
for i in range(len(values)):
for j in range(len(values[0])):
print(task_orders[i][j*2][3:9], task_orders[i][j*2+1][3:9], end='')
if values[i][j] > 0.4:
# print(" ", end=' ')
print(" -".format(values[i][j]), end=' ')
else:
print("{0:.4f}".format(values[i][j]), end=' ')
# print("*", end=' ')
print("")
# print ("======")
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.