text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Use a GPU
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/guide/gpu"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/gpu.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/gpu.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/gpu.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
TensorFlow code, and `tf.keras` models will transparently run on a single GPU with no code changes required.
Note: Use `tf.config.experimental.list_physical_devices('GPU')` to confirm that TensorFlow is using the GPU.
The simplest way to run on multiple GPUs, on one or many machines, is using [Distribution Strategies](distributed_training.ipynb).
This guide is for users who have tried these approaches and found that they need fine-grained control of how TensorFlow uses the GPU.
## Setup
Ensure you have the latest TensorFlow gpu release installed.
```
import tensorflow as tf
print("Num GPUs Available: ", len(tf.config.experimental.list_physical_devices('GPU')))
```
## Overview
TensorFlow supports running computations on a variety of types of devices, including CPU and GPU. They are represented with string identifiers for example:
* `"/device:CPU:0"`: The CPU of your machine.
* `"/GPU:0"`: Short-hand notation for the first GPU of your machine that is visible to TensorFlow.
* `"/job:localhost/replica:0/task:0/device:GPU:1"`: Fully qualified name of the second GPU of your machine that is visible to TensorFlow.
If a TensorFlow operation has both CPU and GPU implementations, by default the GPU devices will be given priority when the operation is assigned to a device. For example, `tf.matmul` has both CPU and GPU kernels. On a system with devices `CPU:0` and `GPU:0`, the `GPU:0` device will be selected to run `tf.matmul` unless you explicitly request running it on another device.
## Logging device placement
To find out which devices your operations and tensors are assigned to, put
`tf.debugging.set_log_device_placement(True)` as the first statement of your
program. Enabling device placement logging causes any Tensor allocations or operations to be printed.
```
tf.debugging.set_log_device_placement(True)
# Create some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
```
The above code will print an indication the `MatMul` op was executed on `GPU:0`.
## Manual device placement
If you would like a particular operation to run on a device of your choice
instead of what's automatically selected for you, you can use `with tf.device`
to create a device context, and all the operations within that context will
run on the same designated device.
```
tf.debugging.set_log_device_placement(True)
# Place tensors on the CPU
with tf.device('/CPU:0'):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
```
You will see that now `a` and `b` are assigned to `CPU:0`. Since a device was
not explicitly specified for the `MatMul` operation, the TensorFlow runtime will
choose one based on the operation and available devices (`GPU:0` in this
example) and automatically copy tensors between devices if required.
## Limiting GPU memory growth
By default, TensorFlow maps nearly all of the GPU memory of all GPUs (subject to
[`CUDA_VISIBLE_DEVICES`](https://docs.nvidia.com/cuda/cuda-c-programming-guide/index.html#env-vars)) visible to the process. This is done to more efficiently use the relatively precious GPU memory resources on the devices by reducing memory fragmentation. To limit TensorFlow to a specific set of GPUs we use the `tf.config.experimental.set_visible_devices` method.
```
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only use the first GPU
try:
tf.config.experimental.set_visible_devices(gpus[0], 'GPU')
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPU")
except RuntimeError as e:
# Visible devices must be set before GPUs have been initialized
print(e)
```
In some cases it is desirable for the process to only allocate a subset of the available memory, or to only grow the memory usage as is needed by the process. TensorFlow provides two methods to control this.
The first option is to turn on memory growth by calling `tf.config.experimental.set_memory_growth`, which attempts to allocate only as much GPU memory as needed for the runtime allocations: it starts out allocating very little memory, and as the program gets run and more GPU memory is needed, we extend the GPU memory region allocated to the TensorFlow process. Note we do not release memory, since it can lead to memory fragmentation. To turn on memory growth for a specific GPU, use the following code prior to allocating any tensors or executing any ops.
```
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
try:
# Currently, memory growth needs to be the same across GPUs
for gpu in gpus:
tf.config.experimental.set_memory_growth(gpu, True)
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Memory growth must be set before GPUs have been initialized
print(e)
```
Another way to enable this option is to set the environmental variable `TF_FORCE_GPU_ALLOW_GROWTH` to `true`. This configuration is platform specific.
The second method is to configure a virtual GPU device with `tf.config.experimental.set_virtual_device_configuration` and set a hard limit on the total memory to allocate on the GPU.
```
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Restrict TensorFlow to only allocate 1GB of memory on the first GPU
try:
tf.config.experimental.set_virtual_device_configuration(
gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPUs,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Virtual devices must be set before GPUs have been initialized
print(e)
```
This is useful if you want to truly bound the amount of GPU memory available to the TensorFlow process. This is common practice for local development when the GPU is shared with other applications such as a workstation GUI.
## Using a single GPU on a multi-GPU system
If you have more than one GPU in your system, the GPU with the lowest ID will be
selected by default. If you would like to run on a different GPU, you will need
to specify the preference explicitly:
```
tf.debugging.set_log_device_placement(True)
try:
# Specify an invalid GPU device
with tf.device('/device:GPU:2'):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
except RuntimeError as e:
print(e)
```
If the device you have specified does not exist, you will get a `RuntimeError`: `.../device:GPU:2 unknown device`.
If you would like TensorFlow to automatically choose an existing and supported device to run the operations in case the specified one doesn't exist, you can call `tf.config.set_soft_device_placement(True)`.
```
tf.config.set_soft_device_placement(True)
tf.debugging.set_log_device_placement(True)
# Creates some tensors
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c = tf.matmul(a, b)
print(c)
```
## Using multiple GPUs
Developing for multiple GPUs will allow a model to scale with the additional resources. If developing on a system with a single GPU, we can simulate multiple GPUs with virtual devices. This enables easy testing of multi-GPU setups without requiring additional resources.
```
gpus = tf.config.experimental.list_physical_devices('GPU')
if gpus:
# Create 2 virtual GPUs with 1GB memory each
try:
tf.config.experimental.set_virtual_device_configuration(
gpus[0],
[tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024),
tf.config.experimental.VirtualDeviceConfiguration(memory_limit=1024)])
logical_gpus = tf.config.experimental.list_logical_devices('GPU')
print(len(gpus), "Physical GPU,", len(logical_gpus), "Logical GPUs")
except RuntimeError as e:
# Virtual devices must be set before GPUs have been initialized
print(e)
```
Once we have multiple logical GPUs available to the runtime, we can utilize the multiple GPUs with `tf.distribute.Strategy` or with manual placement.
#### With `tf.distribute.Strategy`
The best practice for using multiple GPUs is to use `tf.distribute.Strategy`.
Here is a simple example:
```
tf.debugging.set_log_device_placement(True)
strategy = tf.distribute.MirroredStrategy()
with strategy.scope():
inputs = tf.keras.layers.Input(shape=(1,))
predictions = tf.keras.layers.Dense(1)(inputs)
model = tf.keras.models.Model(inputs=inputs, outputs=predictions)
model.compile(loss='mse',
optimizer=tf.keras.optimizers.SGD(learning_rate=0.2))
```
This program will run a copy of your model on each GPU, splitting the input data
between them, also known as "[data parallelism](https://en.wikipedia.org/wiki/Data_parallelism)".
For more information about distribution strategies, check out the guide [here](./distributed_training.ipynb).
#### Manual placement
`tf.distribute.Strategy` works under the hood by replicating computation across devices. You can manually implement replication by constructing your model on each GPU. For example:
```
tf.debugging.set_log_device_placement(True)
gpus = tf.config.experimental.list_logical_devices('GPU')
if gpus:
# Replicate your computation on multiple GPUs
c = []
for gpu in gpus:
with tf.device(gpu.name):
a = tf.constant([[1.0, 2.0, 3.0], [4.0, 5.0, 6.0]])
b = tf.constant([[1.0, 2.0], [3.0, 4.0], [5.0, 6.0]])
c.append(tf.matmul(a, b))
with tf.device('/CPU:0'):
matmul_sum = tf.add_n(c)
print(matmul_sum)
```
| github_jupyter |
```
# -*- coding: utf-8 -*-
import re
from tqdm import tqdm
import time
from datetime import datetime
import sqlite3
import sys
import os
import pandas as pd
import unify
def connect(file_path, primary, columns):
con = sqlite3.connect(file_path)
cur = con.cursor()
cols = ", ".join([c + ' Varchar' for c in columns])
cur.execute("create table meta ("+primary+" Varchar PRIMARY KEY, "+cols+" )")
cur.execute("CREATE INDEX log on meta (textid);")
cur.execute("create table plain_texts (id Varchar(128) NOT NULL PRIMARY KEY, text Varchar NOT NULL);")
cur.execute("create table tagged_texts (id Varchar(128) NOT NULL PRIMARY KEY, text Varchar NOT NULL );")
con.commit()
return con, cur
workdir = r'/home/tari/Загрузки/taiga/nplus1'
filename = 'nplus1.db'
file_path = os.path.join(workdir, filename)
metatablepath = os.path.join(workdir,'newmetadata.csv')
tagged = os.path.join(workdir,'texts_tagged')
plain = os.path.join(workdir,'texts')
meta = pd.read_csv(metatablepath, sep='\t', encoding='utf8')
meta = meta.fillna('')
meta.head()
if not os.path.exists(filename):
con, cur = connect(filename, meta.columns[1], [meta.columns[0]]+list(meta.columns[2:]))
else:
con = sqlite3.connect(filename, meta.columns[1], [meta.columns[0]]+list(meta.columns[2:]))
cur = con.cursor()
cur.execute("SELECT * FROM sqlite_master WHERE type='table';")
print(cur.fetchall())
meta.iloc[6].to_dict()
for i in range(len(meta)):
values = meta.iloc[i].to_dict()
values['textid'] = str(values['textid'])
values['textdiff'] = str(values['textdiff'])
columns = ', '.join(values.keys())
#print(list(values.values()))
placeholders = ', '.join('?' * len(values))
sql = 'INSERT INTO meta ({}) VALUES ({})'.format(columns, placeholders)
#print(sql)
cur.execute(sql, list(values.values()))
valuest = {'id': values['textid'], 'text': unify.open_text(os.path.join(plain, str(values['textid'])+".txt"))}
columns = ', '.join(valuest.keys())
placeholders = ', '.join('?' * len(valuest))
sql2 = 'INSERT INTO plain_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql2, list(valuest.values()))
try:
valuest2 = {'id': values['textid'], 'text': unify.open_text(open(os.path.join(tagged, str(values['textid'])+".txt"),'r', encoding='utf8').read())}
columns = ', '.join(valuest2.keys())
placeholders = ', '.join('?' * len(valuest2))
sql3 = 'INSERT INTO tagged_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql3, list(valuest2.values()))
except:
valuest2 = {'id': values['textid'], 'text': ""}
columns = ', '.join(valuest2.keys())
placeholders = ', '.join('?' * len(valuest2))
sql3 = 'INSERT INTO tagged_texts ({}) VALUES ({})'.format(columns, placeholders)
cur.execute(sql3, list(valuest2.values()))
con.commit()
(textid Varchar PRIMARY KEY, segment Varchar, textname Varchar, textregion Varchar, textrubric Varchar, textdiff Varchar, author Varchar, authortexts Varchar, authorreaders Varchar, magazine Varchar, date Varchar, time Varchar, tags Varchar, source Varchar )
```
| github_jupyter |
<small><i>This notebook was put together by [Jake Vanderplas](http://www.vanderplas.com). Source and license info is on [GitHub](https://github.com/jakevdp/sklearn_tutorial/).</i></small>
# Introduction to Scikit-Learn: Machine Learning with Python
This session will cover the basics of Scikit-Learn, a popular package containing a collection of tools for machine learning written in Python. See more at http://scikit-learn.org.
## Outline
**Main Goal:** To introduce the central concepts of machine learning, and how they can be applied in Python using the Scikit-learn Package.
- Definition of machine learning
- Data representation in scikit-learn
- Introduction to the Scikit-learn API
## About Scikit-Learn
[Scikit-Learn](http://github.com/scikit-learn/scikit-learn) is a Python package designed to give access to **well-known** machine learning algorithms within Python code, through a **clean, well-thought-out API**. It has been built by hundreds of contributors from around the world, and is used across industry and academia.
Scikit-Learn is built upon Python's [NumPy (Numerical Python)](http://numpy.org) and [SciPy (Scientific Python)](http://scipy.org) libraries, which enable efficient in-core numerical and scientific computation within Python. As such, scikit-learn is not specifically designed for extremely large datasets, though there is [some work](https://github.com/ogrisel/parallel_ml_tutorial) in this area.
For this short introduction, I'm going to stick to questions of in-core processing of small to medium datasets with Scikit-learn.
## What is Machine Learning?
In this section we will begin to explore the basic principles of machine learning.
Machine Learning is about building programs with **tunable parameters** (typically an
array of floating point values) that are adjusted automatically so as to improve
their behavior by **adapting to previously seen data.**
Machine Learning can be considered a subfield of **Artificial Intelligence** since those
algorithms can be seen as building blocks to make computers learn to behave more
intelligently by somehow **generalizing** rather that just storing and retrieving data items
like a database system would do.
We'll take a look at two very simple machine learning tasks here.
The first is a **classification** task: the figure shows a
collection of two-dimensional data, colored according to two different class
labels. A classification algorithm may be used to draw a dividing boundary
between the two clusters of points:
```
%matplotlib inline
# set seaborn plot defaults.
# This can be safely commented out
import seaborn; seaborn.set()
# Import the example plot from the figures directory
from fig_code import plot_sgd_separator
plot_sgd_separator()
```
This may seem like a trivial task, but it is a simple version of a very important concept.
By drawing this separating line, we have learned a model which can **generalize** to new
data: if you were to drop another point onto the plane which is unlabeled, this algorithm
could now **predict** whether it's a blue or a red point.
If you'd like to see the source code used to generate this, you can either open the
code in the `figures` directory, or you can load the code using the `%load` magic command:
The next simple task we'll look at is a **regression** task: a simple best-fit line
to a set of data:
```
from fig_code import plot_linear_regression
plot_linear_regression()
```
Again, this is an example of fitting a model to data, such that the model can make
generalizations about new data. The model has been **learned** from the training
data, and can be used to predict the result of test data:
here, we might be given an x-value, and the model would
allow us to predict the y value. Again, this might seem like a trivial problem,
but it is a basic example of a type of operation that is fundamental to
machine learning tasks.
## Representation of Data in Scikit-learn
Machine learning is about creating models from data: for that reason, we'll start by
discussing how data can be represented in order to be understood by the computer. Along
with this, we'll build on our matplotlib examples from the previous section and show some
examples of how to visualize data.
Most machine learning algorithms implemented in scikit-learn expect data to be stored in a
**two-dimensional array or matrix**. The arrays can be
either ``numpy`` arrays, or in some cases ``scipy.sparse`` matrices.
The size of the array is expected to be `[n_samples, n_features]`
- **n_samples:** The number of samples: each sample is an item to process (e.g. classify).
A sample can be a document, a picture, a sound, a video, an astronomical object,
a row in database or CSV file,
or whatever you can describe with a fixed set of quantitative traits.
- **n_features:** The number of features or distinct traits that can be used to describe each
item in a quantitative manner. Features are generally real-valued, but may be boolean or
discrete-valued in some cases.
The number of features must be fixed in advance. However it can be very high dimensional
(e.g. millions of features) with most of them being zeros for a given sample. This is a case
where `scipy.sparse` matrices can be useful, in that they are
much more memory-efficient than numpy arrays.

(Figure from the [Python Data Science Handbook](https://github.com/jakevdp/PythonDataScienceHandbook))
## A Simple Example: the Iris Dataset
As an example of a simple dataset, we're going to take a look at the
iris data stored by scikit-learn.
The data consists of measurements of three different species of irises.
There are three species of iris in the dataset, which we can picture here:
```
from IPython.core.display import Image, display
display(Image(filename='images/iris_setosa.jpg'))
print("Iris Setosa\n")
display(Image(filename='images/iris_versicolor.jpg'))
print("Iris Versicolor\n")
display(Image(filename='images/iris_virginica.jpg'))
print("Iris Virginica")
```
### Quick Question:
**If we want to design an algorithm to recognize iris species, what might the data be?**
Remember: we need a 2D array of size `[n_samples x n_features]`.
- What would the `n_samples` refer to?
- What might the `n_features` refer to?
Remember that there must be a **fixed** number of features for each sample, and feature
number ``i`` must be a similar kind of quantity for each sample.
### Loading the Iris Data with Scikit-Learn
Scikit-learn has a very straightforward set of data on these iris species. The data consist of
the following:
- Features in the Iris dataset:
1. sepal length in cm
2. sepal width in cm
3. petal length in cm
4. petal width in cm
- Target classes to predict:
1. Iris Setosa
2. Iris Versicolour
3. Iris Virginica
``scikit-learn`` embeds a copy of the iris CSV file along with a helper function to load it into numpy arrays:
```
from sklearn.datasets import load_iris
iris = load_iris()
iris.keys()
n_samples, n_features = iris.data.shape
print((n_samples, n_features))
print(iris.data[0])
print(iris.data.shape)
print(iris.target.shape)
print(iris.target)
print(iris.target_names)
```
This data is four dimensional, but we can visualize two of the dimensions
at a time using a simple scatter-plot:
```
import numpy as np
import matplotlib.pyplot as plt
x_index = 0
y_index = 1
# this formatter will label the colorbar with the correct target names
formatter = plt.FuncFormatter(lambda i, *args: iris.target_names[int(i)])
plt.scatter(iris.data[:, x_index], iris.data[:, y_index],
c=iris.target, cmap=plt.cm.get_cmap('RdYlBu', 3))
plt.colorbar(ticks=[0, 1, 2], format=formatter)
plt.clim(-0.5, 2.5)
plt.xlabel(iris.feature_names[x_index])
plt.ylabel(iris.feature_names[y_index]);
```
### Quick Exercise:
**Change** `x_index` **and** `y_index` **in the above script
and find a combination of two parameters
which maximally separate the three classes.**
This exercise is a preview of **dimensionality reduction**, which we'll see later.
## Other Available Data
They come in three flavors:
- **Packaged Data:** these small datasets are packaged with the scikit-learn installation,
and can be downloaded using the tools in ``sklearn.datasets.load_*``
- **Downloadable Data:** these larger datasets are available for download, and scikit-learn
includes tools which streamline this process. These tools can be found in
``sklearn.datasets.fetch_*``
- **Generated Data:** there are several datasets which are generated from models based on a
random seed. These are available in the ``sklearn.datasets.make_*``
You can explore the available dataset loaders, fetchers, and generators using IPython's
tab-completion functionality. After importing the ``datasets`` submodule from ``sklearn``,
type
datasets.load_ + TAB
or
datasets.fetch_ + TAB
or
datasets.make_ + TAB
to see a list of available functions.
```
from sklearn import datasets
# Type datasets.fetch_<TAB> or datasets.load_<TAB> in IPython to see all possibilities
# datasets.fetch_
# datasets.load_
```
In the next section, we'll use some of these datasets and take a look at the basic principles of machine learning.
| github_jupyter |
# Plain examples and tests for the Dataset and the Frame classes
If you are looking for a quick example on how to use the `smartdoc15_ch1` package, we recommend you start by looking at the tutorials instead.
## Import
```
from smartdoc15_ch1 import Dataset
```
## Dataset loading
We use a reduced version of the dataset for testing purpose here. It contains only a fraction of the data.
The dataset is made accessible by creating an instance of the `Dataset` class.
```
d = Dataset(data_home="/data/competitions/2015-ICDAR-smartdoc/challenge1/99-computable-version-2017-test",
download_if_missing=False)
```
See `Dataset` documentation for a description of the arguments of the constructor.
## Dataset format
The resulting instance is a modified `list` which makes it easy to access and iterate over elements.
```
d[0]
```
Index-based access is available, as well as iteration.
```
len(d)
```
*(As said previously, we use a reduced version of the dataset here.)*
```
for frame in d[:10]:
print(frame["image_path"])
```
## Dataset content
Dataset elements are `Frame` objects. Each of them describe a frame of the dataset.
`Frame` objects are `dict` objects with few more methods and properties.
```
d0 = d[0]
type(d0)
d0
```
Frames contain everything one may want to know about a particular frame, and every value can be accessed like in a regular `dict` object.
```
d0["bg_name"], d0['model_name'], d0['frame_index']
```
Here, `d0` represents the frame `181` of the capture for `datasheet004` in `background01`.
Please note that `frame_index` values are indexed starting at `1`, as the codecs usually index them in such way. This is the only value which has this indexing, all other values (labels in particular) are indexed starting at `0`.
## Reading images
Of course, the whole point of this structure is to facilitate the iteration over dataset images.
One can read the image associated to a frame using the `Frame.read_image()` method.
```
img0 = d0.read_image()
img0.shape
```
By default, images are loaded in grayscale format, unscaled.
You can ask for the color image if needed.
```
img0_color = d0.read_image(color=True)
img0_color.shape
```
And you can also force the image to be scaled. This reduces the image in both dimensions by the same factor.
```
img0_reduced = d0.read_image(force_scale_factor=0.5)
img0_reduced.shape
```
## Easy-to-use segmentation for each frame
You may need to access the segmentation target for a given frame without browsing the entire segmentation array by index. We offer two formats:
- dict format with a textual indexing of each coordinate
- a list format with coordinates is the same order as the `Dataset.segmentation_targets` array: "tl_x", "tl_y", "bl_x", "bl_y", "br_x", "br_y", "tr_x", "tr_y"
```
d0.segmentation_dict
d0.segmentation_list
```
## Scaled segmentation for frames
You may need the scaled segmentation for a given frame, provided its scaling factor was defined during `Dataset` creation, and not forced with the parameter `force_scale_factor` of `Frame.read_image()`. We provide the scaled equivalents for the previous segmentation accessors.
```
d0.segmentation_dict_scaled
d0.segmentation_list_scaled
```
## Target values
If you process frames in batch, you may want to access the target values for each possible task in a single line.
The following methods enable you to do so, listing the expected values (unscaled in the case of the segmentation) for the segmentation and the classification tasks.
The expected values are returned as a Numpy array for which the rows are sorted as in the `Dataset` object.
```
d.model_classif_targets[:10]
d.modeltype_classif_targets[:10]
d.segmentation_targets[:10]
```
For the segmentation task, the actual physical shape of the document objects is required to compare the returned results and the expected ones.
```
d.model_shapes[:10]
```
For the sake of completeness, background labels can be obtained easily as well.
```
d.background_labels[:10]
```
## Listing label values
Some properties may help you listing the possible values for `background_id`, `model_id` and `modeltype_id`.
```
d.unique_background_ids
d.unique_background_names
d.unique_model_ids
d.unique_model_names
d.unique_modeltype_ids
d.unique_modeltype_names
```
Finally, the underlying Pandas Dataframe is made available directly in case you need more flexibility.
```
d.raw_dataframe[:10]
```
| github_jupyter |
# fastText vs word2vec
# Data
```
import nltk
from smart_open import smart_open
nltk.download('brown')
# Only the brown corpus is needed in case you don't have it.
# Generate brown corpus text file
with smart_open('brown_corp.txt', 'w+') as f:
for word in nltk.corpus.brown.words():
f.write('{word} '.format(word=word))
# Make sure you set FT_HOME to your fastText directory root
FT_HOME = 'fastText/'
# download the text8 corpus (a 100 MB sample of cleaned wikipedia text)
import os.path
if not os.path.isfile('text8'):
!wget -c http://mattmahoney.net/dc/text8.zip
!unzip text8.zip
# download and preprocess the text9 corpus
if not os.path.isfile('text9'):
!wget -c http://mattmahoney.net/dc/enwik9.zip
!unzip enwik9.zip
!perl {FT_HOME}wikifil.pl enwik9 > text9
```
# Train models
For training the models yourself, you'll need to have both [Gensim](https://github.com/RaRe-Technologies/gensim) and [FastText](https://github.com/facebookresearch/fastText) set up on your machine.
```
MODELS_DIR = 'models/'
!mkdir -p {MODELS_DIR}
lr = 0.05
dim = 100
ws = 5
epoch = 5
minCount = 5
neg = 5
loss = 'ns'
t = 1e-4
from gensim.models import Word2Vec, KeyedVectors
from gensim.models.word2vec import Text8Corpus
# Same values as used for fastText training above
params = {
'alpha': lr,
'size': dim,
'window': ws,
'iter': epoch,
'min_count': minCount,
'sample': t,
'sg': 1,
'hs': 0,
'negative': neg
}
def train_models(corpus_file, output_name):
output_file = '{:s}_ft'.format(output_name)
if not os.path.isfile(os.path.join(MODELS_DIR, '{:s}.vec'.format(output_file))):
print('Training fasttext on {:s} corpus..'.format(corpus_file))
%time !{FT_HOME}fasttext skipgram -input {corpus_file} -output {MODELS_DIR+output_file} -lr {lr} -dim {dim} -ws {ws} -epoch {epoch} -minCount {minCount} -neg {neg} -loss {loss} -t {t}
else:
print('\nUsing existing model file {:s}.vec'.format(output_file))
output_file = '{:s}_ft_no_ng'.format(output_name)
if not os.path.isfile(os.path.join(MODELS_DIR, '{:s}.vec'.format(output_file))):
print('\nTraining fasttext on {:s} corpus (without char n-grams)..'.format(corpus_file))
%time !{FT_HOME}fasttext skipgram -input {corpus_file} -output {MODELS_DIR+output_file} -lr {lr} -dim {dim} -ws {ws} -epoch {epoch} -minCount {minCount} -neg {neg} -loss {loss} -t {t} -maxn 0
else:
print('\nUsing existing model file {:s}.vec'.format(output_file))
output_file = '{:s}_gs'.format(output_name)
if not os.path.isfile(os.path.join(MODELS_DIR, '{:s}.vec'.format(output_file))):
print('\nTraining word2vec on {:s} corpus..'.format(corpus_file))
# Text8Corpus class for reading space-separated words file
%time gs_model = Word2Vec(Text8Corpus(corpus_file), **params); gs_model
# Direct local variable lookup doesn't work properly with magic statements (%time)
locals()['gs_model'].wv.save_word2vec_format(os.path.join(MODELS_DIR, '{:s}.vec'.format(output_file)))
print('\nSaved gensim model as {:s}.vec'.format(output_file))
else:
print('\nUsing existing model file {:s}.vec'.format(output_file))
evaluation_data = {}
train_models('brown_corp.txt', 'brown')
train_models(corpus_file='text8', output_name='text8')
train_models(corpus_file='text9', output_name='text9')
```
# Comparisons
```
# download the file questions-words.txt to be used for comparing word embeddings
!wget https://raw.githubusercontent.com/tmikolov/word2vec/master/questions-words.txt
```
Once you have downloaded or trained the models and downloaded `questions-words.txt`, you're ready to run the comparison.
```
import logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
# Training times in seconds
evaluation_data['brown'] = [(18, 54.3, 32.5)]
evaluation_data['text8'] = [(402, 942, 496)]
evaluation_data['text9'] = [(3218, 6589, 3550)]
def print_accuracy(model, questions_file):
print('Evaluating...\n')
acc = model.accuracy(questions_file)
sem_correct = sum((len(acc[i]['correct']) for i in range(5)))
sem_total = sum((len(acc[i]['correct']) + len(acc[i]['incorrect'])) for i in range(5))
sem_acc = 100*float(sem_correct)/sem_total
print('\nSemantic: {:d}/{:d}, Accuracy: {:.2f}%'.format(sem_correct, sem_total, sem_acc))
syn_correct = sum((len(acc[i]['correct']) for i in range(5, len(acc)-1)))
syn_total = sum((len(acc[i]['correct']) + len(acc[i]['incorrect'])) for i in range(5,len(acc)-1))
syn_acc = 100*float(syn_correct)/syn_total
print('Syntactic: {:d}/{:d}, Accuracy: {:.2f}%\n'.format(syn_correct, syn_total, syn_acc))
return (sem_acc, syn_acc)
word_analogies_file = 'questions-words.txt'
accuracies = []
print('\nLoading Gensim embeddings')
brown_gs = KeyedVectors.load_word2vec_format(MODELS_DIR + 'brown_gs.vec')
print('Accuracy for Word2Vec:')
accuracies.append(print_accuracy(brown_gs, word_analogies_file))
print('\nLoading FastText embeddings')
brown_ft = KeyedVectors.load_word2vec_format(MODELS_DIR + 'brown_ft.vec')
print('Accuracy for FastText (with n-grams):')
accuracies.append(print_accuracy(brown_ft, word_analogies_file))
```
The `accuracy` takes an optional parameter `restrict_vocab`, which limits the vocabulary of model considered for fast approximate evaluation (default is 30000).
Word2Vec embeddings seem to be slightly better than fastText embeddings at the semantic tasks, while the fastText embeddings do significantly better on the syntactic analogies. Makes sense, since fastText embeddings are trained for understanding morphological nuances, and most of the syntactic analogies are morphology based.
Let me explain that better.
According to the paper [[1]](https://arxiv.org/abs/1607.04606), embeddings for words are represented by the sum of their n-gram embeddings. This is meant to be useful for morphologically rich languages - so theoretically, the embedding for `apparently` would include information from both character n-grams `apparent` and `ly` (as well as other n-grams), and the n-grams would combine in a simple, linear manner. This is very similar to what most of our syntactic tasks look like.
Example analogy:
`amazing amazingly calm calmly`
This analogy is marked correct if:
`embedding(amazing)` - `embedding(amazingly)` = `embedding(calm)` - `embedding(calmly)`
Both these subtractions would result in a very similar set of remaining ngrams.
No surprise the fastText embeddings do extremely well on this.
Let's do a small test to validate this hypothesis - fastText differs from word2vec only in that it uses char n-gram embeddings as well as the actual word embedding in the scoring function to calculate scores and then likelihoods for each word, given a context word. In case char n-gram embeddings are not present, this reduces (at least theoretically) to the original word2vec model. This can be implemented by setting 0 for the max length of char n-grams for fastText.
```
print('Loading FastText embeddings')
brown_ft_no_ng = KeyedVectors.load_word2vec_format(MODELS_DIR + 'brown_ft_no_ng.vec')
print('Accuracy for FastText (without n-grams):')
accuracies.append(print_accuracy(brown_ft_no_ng, word_analogies_file))
evaluation_data['brown'] += [[acc[0] for acc in accuracies], [acc[1] for acc in accuracies]]
```
A-ha! The results for FastText with no n-grams and Word2Vec look a lot more similar (as they should) - the differences could easily result from differences in implementation between fastText and Gensim, and randomization. Especially telling is that the semantic accuracy for FastText has improved slightly after removing n-grams, while the syntactic accuracy has taken a giant dive. Our hypothesis that the char n-grams result in better performance on syntactic analogies seems fair. It also seems possible that char n-grams hurt semantic accuracy a little. However, the brown corpus is too small to be able to draw any definite conclusions - the accuracies seem to vary significantly over different runs.
Let's try with a larger corpus now - text8 (collection of wiki articles). I'm also curious about the impact on semantic accuracy - for models trained on the brown corpus, the difference in the semantic accuracy and the accuracy values themselves are too small to be conclusive. Hopefully a larger corpus helps, and the text8 corpus likely has a lot more information about capitals, currencies, cities etc, which should be relevant to the semantic tasks.
```
accuracies = []
print('Loading Gensim embeddings')
text8_gs = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text8_gs.vec')
print('Accuracy for word2vec:')
accuracies.append(print_accuracy(text8_gs, word_analogies_file))
print('Loading FastText embeddings (with n-grams)')
text8_ft = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text8_ft.vec')
print('Accuracy for FastText (with n-grams):')
accuracies.append(print_accuracy(text8_ft, word_analogies_file))
print('Loading FastText embeddings')
text8_ft_no_ng = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text8_ft_no_ng.vec')
print('Accuracy for FastText (without n-grams):')
accuracies.append(print_accuracy(text8_ft_no_ng, word_analogies_file))
evaluation_data['text8'] += [[acc[0] for acc in accuracies], [acc[1] for acc in accuracies]]
```
With the text8 corpus, we observe a similar pattern. Semantic accuracy falls by a small but significant amount when n-grams are included in FastText, while FastText with n-grams performs far better on the syntactic analogies. FastText without n-grams are largely similar to Word2Vec.
My hypothesis for semantic accuracy being lower for the FastText-with-ngrams model is that most of the words in the semantic analogies are standalone words and are unrelated to their morphemes (eg: father, mother, France, Paris), hence inclusion of the char n-grams into the scoring function actually makes the embeddings worse.
This trend is observed in the original paper too where the performance of embeddings with n-grams is worse on semantic tasks than both word2vec cbow and skipgram models.
Let's do a quick comparison on an even larger corpus - text9
```
accuracies = []
print('Loading Gensim embeddings')
text9_gs = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text9_gs.vec')
print('Accuracy for word2vec:')
accuracies.append(print_accuracy(text9_gs, word_analogies_file))
print('Loading FastText embeddings (with n-grams)')
text9_ft = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text9_ft.vec')
print('Accuracy for FastText (with n-grams):')
accuracies.append(print_accuracy(text9_ft, word_analogies_file))
print('Loading FastText embeddings')
text9_ft_no_ng = KeyedVectors.load_word2vec_format(MODELS_DIR + 'text9_ft_no_ng.vec')
print('Accuracy for FastText (without n-grams):')
accuracies.append(print_accuracy(text9_ft_no_ng, word_analogies_file))
evaluation_data['text9'] += [[acc[0] for acc in accuracies], [acc[1] for acc in accuracies]]
%matplotlib inline
import matplotlib.pyplot as plt
def plot(ax, data, corpus_name='brown'):
width = 0.25
pos = [(i, i + width, i + 2*width) for i in range(len(data))]
colors = ['#EE3224', '#F78F1E', '#FFC222']
acc_ax = ax.twinx()
# Training time
ax.bar(pos[0],
data[0],
width,
alpha=0.5,
color=colors
)
# Semantic accuracy
acc_ax.bar(pos[1],
data[1],
width,
alpha=0.5,
color=colors
)
# Syntactic accuracy
acc_ax.bar(pos[2],
data[2],
width,
alpha=0.5,
color=colors
)
ax.set_ylabel('Training time (s)')
acc_ax.set_ylabel('Accuracy (%)')
ax.set_title(corpus_name)
acc_ax.set_xticks([p[0] + 1.5 * width for p in pos])
acc_ax.set_xticklabels(['Training Time', 'Semantic Accuracy', 'Syntactic Accuracy'])
# Proxy plots for adding legend correctly
proxies = [ax.bar([0], [0], width=0, color=c, alpha=0.5)[0] for c in colors]
models = ('Gensim', 'FastText', 'FastText (no-ngrams)')
ax.legend((proxies), models, loc='upper left')
ax.set_xlim(pos[0][0]-width, pos[-1][0]+width*4)
ax.set_ylim([0, max(data[0])*1.1] )
acc_ax.set_ylim([0, max(data[1] + data[2])*1.1] )
plt.grid()
# Plotting the bars
fig = plt.figure(figsize=(10,15))
for corpus, subplot in zip(sorted(evaluation_data.keys()), [311, 312, 313]):
ax = fig.add_subplot(subplot)
plot(ax, evaluation_data[corpus], corpus)
plt.show()
```
The results from text9 seem to confirm our hypotheses so far. Briefly summarising the main points -
1. FastText models with n-grams do significantly better on syntactic tasks, because of the syntactic questions being related to morphology of the words
2. Both Gensim word2vec and the fastText model with no n-grams do slightly better on the semantic tasks, presumably because words from the semantic questions are standalone words and unrelated to their char n-grams
3. In general, the performance of the models seems to get closer with the increasing corpus size. However, this might possibly be due to the size of the model staying constant at 100, and a larger model size for large corpora might result in higher performance gains.
4. The semantic accuracy for all models increases significantly with the increase in corpus size.
5. However, the increase in syntactic accuracy from the increase in corpus size for the n-gram FastText model is lower (in both relative and absolute terms). This could possibly indicate that advantages gained by incorporating morphological information could be less significant in case of larger corpus sizes (the corpuses used in the original paper seem to indicate this too)
6. Training times for gensim are slightly lower than the fastText no-ngram model, and significantly lower than the n-gram variant. This is quite impressive considering fastText is implemented in C++ and Gensim in Python (with calls to low-level BLAS routines for much of the heavy lifting). You could read [this post](http://rare-technologies.com/word2vec-in-python-part-two-optimizing/) for more details regarding word2vec optimisation in Gensim. Note that these times include importing any dependencies and serializing the models to disk, and not just the training times.
# Conclusions
These preliminary results seem to indicate fastText embeddings are significantly better than word2vec at encoding syntactic information. This is expected, since most syntactic analogies are morphology based, and the char n-gram approach of fastText takes such information into account. The original word2vec model seems to perform better on semantic tasks, since words in semantic analogies are unrelated to their char n-grams, and the added information from irrelevant char n-grams worsens the embeddings. It'd be interesting to see how transferable these embeddings are for different kinds of tasks by comparing their performance in a downstream supervised task.
| github_jupyter |
```
!pip install statsmodels
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
sales_data = pd.read_csv('sales.csv')
sales_data.head()
plt.figure(figsize=(10,6))
plt.figure(figsize=(10,6))
plt.plot(sales_data['Time'], sales_data['Sales'], label='Sales-Original')
plt.xticks(rotation=60)
plt.legend()
plt.show()
sales_data['3MA']=sales_data['Sales'].rolling(window=3).mean()
sales_data['5MA']=sales_data['Sales'].rolling(window=5).mean()
sales_data.head()
plt.figure(figsize=(10,6))
plt.plot(sales_data['Time'], sales_data['Sales'], label='Sales-Original', color='blue')
plt.plot(sales_data['Time'], sales_data['3MA'], label='3-Moving Average (3MA)', color='green')
plt.plot(sales_data['Time'], sales_data['5MA'], label='5-Moving Average (5MA)', color='red')
plt.xticks(rotation=60)
plt.legend()
plt.show()
import pandas as pd
import statsmodels.api as sm
import matplotlib.pyplot as plt
sales_data = pd.read_csv('sales.csv', index_col='Time')
sales_data.head()
sales_data['simple']=sales_data.Sales.rolling(window=3).mean()
sales_data['boxcar']=sales_data.Sales.rolling(3,win_type='boxcar').mean()
sales_data['triang']=sales_data.Sales.rolling(3,win_type='triang').mean()
sales_data['hamming']=sales_data.Sales.rolling(3,win_type='hamming').mean()
sales_data['blackman']=sales_data.Sales.rolling(3,win_type='blackman').mean()
sales_data.head()
sales_data.plot(kind='line', figsize=(10,6))
import statsmodels.api as sm
import pandas as pd
import statsmodels.tsa.stattools as ts
import numpy as np
def calc_adf(x,y):
result = sm.OLS(x,y).fit()
return ts.adfuller(result.resid)
data = sm.datasets.sunspots.load_pandas().data.values
N = len(data)
t = np.linspace(-2*np.pi, 2*np.pi, N)
sine = np.sin(np.sin(t))
print('Self ADF', calc_adf(sine,sine))
noise = np.random.normal(0,.01,N)
print('ADF sine with noise', calc_adf(sine, sine+noise))
cosine = 100*np.cos(t) + 10
print('ADF sine vs cosine with noise', calc_adf(sine, cosine+noise))
print('Sine vs sunspots', calc_adf(sine, data))
sales_data
first_diffs = sales_data.Sales.values[1:] - sales_data.Sales.values[:-1]
sales_data.Sales.values
sales_data.Sales.values[1:]
sales_data.Sales.values[:-1]
np.concatenate([first_diffs,[0]])
calc_adf(sine,sine)[1]
def calc_ols(x,y):
result = sm.OLS(x,y).fit()
return result
lags = 2
series = [np.random.normal() for _ in range(lags)]
series = [1,2]
coefs = [0.2,0.3]
prev_vals = series[-lags:][::-1]
new_val = np.array(prev_vals)*coefs
prev_vals, new_val
import pandas as pd
import matplotlib.pyplot as plt
from statsmodels.tsa.seasonal import seasonal_decompose
data = pd.read_csv('beer_production.csv')
data.columns = ['date', 'data']
data['date'] = pd.to_datetime(data['date'])
data = data.set_index('date')
decomposed_data = seasonal_decompose(data,model='multiplicative')
import pylab
pylab.rcParams['figure.figsize']=(8,6)
decomposed_data.plot()
plt.show()
import pandas as pd
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
data = sm.datasets.sunspots.load_pandas().data
data
dy = data.SUNACTIVITY - np.mean(data.SUNACTIVITY)
dy_square = np.sum(dy**2)
sun_correlated = np.correlate(dy, dy, mode='full')/dy_square
sun_correlated
result = sun_correlated[int(len(sun_correlated)/2):]
plt.plot(result)
plt.grid(True)
plt.xlabel('Lag')
plt.ylabel('Autocorrelation')
plt.show()
from pandas.plotting import autocorrelation_plot
autocorrelation_plot(data.SUNACTIVITY)
from statsmodels.tsa.ar_model import AR
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
import matplotlib.pyplot as plt
import statsmodels.api as sm
from math import sqrt
data = sm.datasets.sunspots.load_pandas().data
train_ratio = 0.8
train = data[:int(train_ratio*len(data))]
test = data[int(train_ratio*len(data)):]
ar_model = AR(train.SUNACTIVITY)
ar_model = ar_model.fit()
print('Number of Lags:', ar_model.k_ar)
print('Model Coefficients:\n', ar_model.params)
start_point = len(train)
end_point = start_point + len(test) - 1
pred = ar_model.predict(start=start_point, end = end_point, dynamic = False)
mae = mean_absolute_error(test.SUNACTIVITY, pred)
mse = mean_squared_error(test.SUNACTIVITY, pred)
rmse = sqrt(mse)
print('MAE:', mae)
print('MSE:', mse)
print('RMSE:', rmse)
plt.figure(figsize=(10,6))
plt.plot(test.SUNACTIVITY, label='Original-Series')
plt.plot(pred, color='red', label='Predicted Series')
plt.legend()
plt.show()
import statsmodels.api as sm
from statsmodels.tsa.arima_model import ARMA
from sklearn.metrics import mean_absolute_error, mean_squared_error
import matplotlib.pyplot as plt
from math import sqrt
data = sm.datasets.sunspots.load_pandas().data
data
train_ratio = 0.8
train = data[:int(train_ratio*len(data))]
test = data[int(train_ratio*len(data)):]
arma_model = ARMA(train.SUNACTIVITY, order=(10,1))
arma_model = arma_model.fit()
start_point = len(train)
end_point = start_point + len(test) -1
pred = arma_model.predict(start_point, end_point)
mae = mean_absolute_error(test.SUNACTIVITY, pred)
mse = mean_squared_error(test.SUNACTIVITY, pred)
rmse = sqrt(mse)
print('MAE:', mae)
print('MSE:', mse)
print('RMSE:', rmse)
plt.figure(figsize=(10,6))
plt.plot(test.SUNACTIVITY, label='Original-Series')
plt.plot(pred, color='red', label='Predicted Series')
plt.legend()
plt.show()
import numpy as np
import statsmodels.api as sm
from scipy.optimize import leastsq
import matplotlib.pyplot as plt
def model(p,t):
C, p1, f1, phi1, p2, f2, phi2, p3, f3, phi3 = p
return C + p1*np.sin(f1*t + phi1) + p2*np.sin(f2*t + phi2) + p3*np.sin(f3*t+phi3)
def error(p, y, t):
return y - model(p,t)
def fit(y, t):
p0 = [y.mean(), 0, 2*np.pi/11, 0, 0, 2*np.pi/22, 0, 0, 2*np.pi/100, 0]
params = leastsq(error, p0, args=(y,t))[0]
return params
data_loader = sm.datasets.sunspots.load_pandas()
sunspots = data_loader.data['SUNACTIVITY'].values
years = data_loader.data['YEAR'].values
cutoff = int(.9*len(sunspots))
params = fit(sunspots[:cutoff], years[:cutoff])
print('Params', params)
pred = model(params, years[cutoff:])
actual = sunspots[cutoff:]
print('Root mean square error', np.sqrt(np.mean((actual-pred)**2)))
print('Mean absolute error', np.mean(np.abs(actual-pred)))
print('Mean absolute percentage error', 100 * np.mean(np.abs(actual-pred)/actual))
mid = (actual+pred)/2
print('Symmetric mean absolute percentage error', 100 * np.mean(np.abs(actual-pred)/mid))
print('Coefficient of determination', 1 - ((actual-pred)**2).sum()/((actual-actual.mean())**2).sum())
year_range = data_loader.data['YEAR'].values[cutoff:]
plt.plot(year_range, actual, 'o', label='Sunspots')
plt.plot(year_range, pred, 'x',label='Prediction')
plt.grid(True)
plt.xlabel('YEAR')
plt.ylabel('SUNACTIVITY')
plt.legend()
plt.show()
import numpy as np
import statsmodels.api as sm
import matplotlib.pyplot as plt
from scipy.fftpack import rfft, fftshift
data = sm.datasets.sunspots.load_pandas().data
t = np.linspace(-2*np.pi, 2*np.pi, len(data.SUNACTIVITY.values))
mid = np.ptp(data.SUNACTIVITY.values)/2
sine = mid + mid * np.sin(np.sin(t))
sine2 = mid + mid * np.sin(t)
sine_fft = np.abs(fftshift(rfft(sine)))
sine2_fft = np.abs(fftshift(rfft(sine2)))
print('Index of max sine FFT', np.argsort(sine_fft)[-5:])
print('Index of max sine2 FFT', np.argsort(sine2_fft)[-5:])
transformed = np.abs(fftshift(rfft(data.SUNACTIVITY.values)))
print('Indices of max sunspots FFT', np.argsort(transformed)[-5:])
fig, axs = plt.subplots(3, figsize=(12,6), sharex=True)
fig.suptitle('Power Spectrum')
axs[0].plot(data.SUNACTIVITY.values, label='Sunspots')
axs[0].plot(sine, lw=2, label='Sine')
axs[0].plot(sine2, lw=2, label='Sine2')
axs[0].legend()
axs[1].plot(transformed, label='Transformed Sunspots')
axs[1].legend()
axs[2].plot(sine_fft, lw=2, label='Transformed Sine')
axs[2].plot(sine2_fft, lw=2, label='Transformed Sine2')
axs[2].legend()
plt.show()
import numpy as np
import statsmodels.api as sm
from scipy.fftpack import rfft
from scipy.fftpack import fftshift
import matplotlib.pyplot as plt
data = sm.datasets.sunspots.load_pandas().data
transformed = fftshift(rfft(data.SUNACTIVITY.values))
power = transformed **2
phase = np.angle(transformed)
fig, axs = plt.subplots(3, figsize = (12,6), sharex=True)
fig.suptitle('Power Spectrum')
axs[0].plot(data.SUNACTIVITY.values, label='Sunspots')
axs[0].legend(loc='upper right')
axs[1].plot(power, label = 'Power Spectrum')
axs[1].legend()
axs[2].plot(phase, label='Phase Spectrum')
axs[2].legend()
plt.show()
```
| github_jupyter |
# Classificador de Carros usando Tensorflow
Neste notebook iremos implementadar um modelo para classificação de imagens. Classificação é uma das "tarefas" em que podemos utilizar Machine Learning, nesta tarefa o ensino é **supervisionado**, em outras palavras nós vamos ensinar ao modelo através de exemplos com gabarito.
Nosso modelo deverá receber imagens de veículos e não-veículos e identificar a que **classe** (veículo ou não-veículo) estas imagens pertencem.
## Dados
Os dados foram retirados da base de dados da Udacity ([Veículos](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/vehicles.zip) e [Não-veículos](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/non-vehicles.zip)) que contém aproximadamente 9000 imagens de cada classe.
## Modelo
Iremos utilizar uma CNN com __[CONV -> CONV -> POOL -> DROP -> FULLY_CONV -> DROP -> FULLY_CONV]__. As camadas FULLY_CONV são camadas convolucionais que funcionam como camadas Densas. Nós utilizamos camadas FULLY_CONV quando não queremos restringir o tamanho da entrada para as camadas densas. No [artigo original](https://medium.com/@tuennermann/convolutional-neural-networks-to-find-cars-43cbc4fb713), o autor utiliza-se deve artefato para adaptar uma rede treinada inicialmente para classificação como uma rede para detecção.
## Créditos
Essa atividade é baseada no artigo do Medium encontrado [aqui](https://medium.com/@tuennermann/convolutional-neural-networks-to-find-cars-43cbc4fb713) implementada originalmente por [@tuennermann](https://medium.com/@tuennermann/convolutional-neural-networks-to-find-cars-43cbc4fb713).
Obrigada a todos os envolvidos!
## Dependências
```
# Compatibilidade entre Python 2 e Python 3
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
# TensorFlow
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.INFO) # Permitindo visualização de logs
# Bibliotecas auxiliares
import cv2 # maior eficiência ao processar as imagens
import numpy as np # manipular vetores
from glob import glob # carregar arquivos
import matplotlib.pyplot as plt # plotar imagens
from sklearn.model_selection import train_test_split # dividir o banco em treinamento e teste
%matplotlib inline
# IMPORTANTE: essa linha garante que os números gerados aleatoriamente são previsíveis
np.random.seed(0)
print ('Sua versão do TensorFlow:', tf.__version__)
print ('Recomenda-se para esta atividade uma versão >= 1.4.0')
```
## Buscando os Dados
### Baixa e extraia os dados
Para baixar os dados, crie uma pasta chamada *"data"* na pasta em que se encontra este notebook e execute os seguintes comandos dentro da pasta recém-criada.
```bash
curl -O https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/vehicles.zip
curl -O https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/non-vehicles.zip
unzip vehicles.zip
unzip non-vehicles.zip
```
Ou você pode baixar os dados manualmente clicando [neste link](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/vehicles.zip) e [neste link](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/non-vehicles.zip) e extraí-los nesta pasta.
Após extrair as pastas você deverá ver as seguintes subpastas:
```bash
data/
data/vehicles
data/non-vehicles
```
Crie também uma pasta _models_ na mesma pasta desse notebook.
## Visualizando os dados
```
def print_info(x, y):
unique, counts = np.unique(y, return_counts=True)
print('x: {} {} \t y: {} {} \t counts: {}'.format(x.shape, x.dtype, y.shape, y.dtype, dict(zip(unique, counts))))
def get_samples(x, y, class_label, n_samples=10):
mask = np.where(y == class_label)[0][:n_samples]
return x[mask], y[mask]
def plot_batch(img_batch, y_true, y_pred=None, n_cols=10):
plt.figure(figsize=(16,5))
y_pred = y_true if y_pred is None else y_pred
n_rows = img_batch.shape[0] // n_cols + 1
for img, true, pred, sub in zip(img_batch, y_true, y_pred, range(1, len(img_batch)+1)):
plt.subplot(n_rows, n_cols, sub)
plt.imshow(img.astype(np.uint8))
title = "{}:{:.2f}".format("car" if pred > 0 else "non-car", pred)
pred = np.where(pred > 0, 1, -1)
c = 'green' if true == pred else 'red'
plt.title(title, color = c)
plt.axis('off')
plt.tight_layout()
cars = glob('data/vehicles/*/*.png')
non_cars = glob('data/non-vehicles/*/*.png')
y = np.concatenate([np.ones(len(cars)), np.zeros(len(non_cars))-1])
x = []
for file in cars:
x.append(cv2.imread(file))
for file in non_cars:
x.append(cv2.imread(file))
x = np.array(x)
print_info(x, y)
car_samples, car_labels = get_samples(x, y, class_label=1)
noncar_samples, noncar_labels = get_samples(x, y, class_label=-1)
samples = np.vstack((car_samples, noncar_samples))
labels = np.concatenate((car_labels, noncar_labels))
plot_batch(samples, labels)
# Split the dataset into training, validation, and testing
x_train, x_test, y_train, y_test = train_test_split(x.astype(np.float32), y.astype(np.float32), test_size=0.1, stratify=y, random_state=42)
x_train, x_val, y_train, y_val = train_test_split(x_train, y_train, test_size=x_test.shape[0], stratify=y_train, random_state=42)
print_info(x_train, y_train)
print_info(x_val, y_val)
print_info(x_test, y_test)
```
## Implementando o modelo
### Hiper-parâmetros
```
BATCH_SIZE = 128
TRAIN_EPOCHS = 2
```
### Modelo
```
tf.reset_default_graph()
graph = tf.Graph()
with graph.as_default():
tf_dataset = tf.placeholder(dtype=tf.float32, shape=[None, 64, 64, 3], name='tf_dataset')
tf_labels = tf.placeholder(dtype=tf.float32, shape=[None, 1], name='tf_labels')
tf_dropout = tf.placeholder(dtype=tf.bool, name='tf_dropout')
conv1 = tf.layers.conv2d(inputs=tf_dataset,
filters=10,
kernel_size=[3,3],
activation=tf.nn.relu,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
padding='same',
name='conv1')
conv2 = tf.layers.conv2d(inputs=conv1,
filters=10,
kernel_size=[3,3],
activation=tf.nn.relu,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
padding='same',
name='conv2')
pool1 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[8,8], strides=[8,8])
drop1 = tf.layers.dropout(inputs=pool1, rate=0.25, training=tf_dropout)
dens1 = tf.layers.conv2d(inputs=drop1,
filters=128,
kernel_size=[8,8],
activation=tf.nn.relu,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
padding='valid',
name='dense1')
drop2 = tf.layers.dropout(inputs=dens1, rate=0.5, training=tf_dropout)
output = tf.layers.conv2d(inputs=drop2,
filters=1,
kernel_size=[1,1],
activation=tf.nn.tanh,
kernel_initializer=tf.contrib.layers.xavier_initializer(),
padding='valid',
name='dense2')
loss = tf.reduce_mean(tf.squared_difference(tf.squeeze(tf_labels), tf.squeeze(output)))
optimizer = tf.train.AdadeltaOptimizer(learning_rate=1.0, rho=0.95, epsilon=1e-08).minimize(loss)
output = tf.squeeze(output)
tf_pred = tf.where(output > 0, tf.ones_like(output), -1*tf.ones_like(output))
accuracy = tf.reduce_mean(tf.cast(tf.equal(tf.squeeze(tf_labels), tf_pred), tf.float32))*100
```
## Treinando o Modelo
```
steps_by_epoch = x_train.shape[0] // BATCH_SIZE
n_steps = TRAIN_EPOCHS * steps_by_epoch
with tf.Session(graph=graph) as sess:
saver = tf.train.Saver()
sess.run(tf.global_variables_initializer())
for step in range(n_steps+1):
offset = (step*BATCH_SIZE) % (x_train.shape[0] - BATCH_SIZE)
x_batch = x_train[offset:(offset+BATCH_SIZE)]
y_batch = y_train[offset:(offset+BATCH_SIZE)].reshape(-1,1)
x_batch = (x_batch/127.5)-1.0
feed_dict = {tf_dataset:x_batch, tf_labels:y_batch, tf_dropout:True}
_, loss_batch, accuracy_batch = sess.run([optimizer, loss, accuracy], feed_dict=feed_dict)
print('\rstep: {:=3}/{:=3} batch_loss: {:.5f} batch_accuracy: {:.2f}%'.format(step%steps_by_epoch, steps_by_epoch, loss_batch, accuracy_batch), end='')
if step % steps_by_epoch == 0:
feed_dict = {tf_dataset:(x_val/127.5)-1.0, tf_labels:y_val.reshape(-1,1), tf_dropout:False}
loss_val, accuracy_val = sess.run([loss, accuracy], feed_dict=feed_dict)
print('\repoch: {0:=2} batch_loss: {1:.5f} batch_accuracy: {2:.2f}% val_loss: {3:.5f} val_accuracy: {4:.2f}%'.format(step//steps_by_epoch, loss_batch, accuracy_batch, loss_val, accuracy_val))
feed_dict = {tf_dataset:(x_test/127.5)-1.0, tf_labels:y_test.reshape(-1,1), tf_dropout:False}
[accuracy_test] = sess.run([accuracy], feed_dict=feed_dict)
print('Test Accuracy: {:.2f}%'.format(accuracy_test))
saver.save(sess, 'models/cnn_cars')
```
Mesmo com poucas epochs, conseguimos uma __acurácia > 95%__ no banco de teste!
## Restaurando o modelo e Testando
```
car_test, car_labels = get_samples(x_test, y_test, class_label=1)
noncar_test, noncar_labels = get_samples(x_test, y_test, class_label=-1)
test_batch = np.vstack((car_test, noncar_test))
test_labels = np.concatenate((car_labels, noncar_labels))
plot_batch(test_batch, test_labels, test_labels)
# rode o comando abaixo se você quiser ver o nome de todos os nós do nosso grafo
#[n.name for n in tf.get_default_graph().as_graph_def().node]
with tf.Session() as sess:
new_saver = tf.train.import_meta_graph('{}{}.meta'.format('models/', 'cnn_cars'))
new_saver.restore(sess, tf.train.latest_checkpoint('models/'))
graph = tf.get_default_graph()
tf_dataset = graph.get_tensor_by_name("tf_dataset:0")
tf_labels = graph.get_tensor_by_name("tf_labels:0")
tf_dropout = graph.get_tensor_by_name("tf_dropout:0")
tf_dense2 = graph.get_tensor_by_name("dense2/Tanh:0")
feed_dict = {tf_dataset:(test_batch/127.5)-1.0, tf_labels:test_labels.reshape(-1,1), tf_dropout:False}
[output] = sess.run([tf_dense2], feed_dict=feed_dict)
plot_batch(test_batch, test_labels, np.squeeze(output))
```
Os títulos de cada imagem representam:
- __classe predita__: {car, non-car}
- __score de confiança__: [-1, 1], onde -1 = certeza de não ser carro e 1 = certeza de ser carro. Quando o score é próximo de 0, a rede não tem certeza se a imagem é carro ou não-carro.
Os títulos em vermelho representam as imagem __incorretamente__ classificadas pela nossa rede.
## Referências
- [Artigo Original](https://medium.com/@tuennermann/convolutional-neural-networks-to-find-cars-43cbc4fb713)
- [Código Original (in Keras)](https://github.com/HTuennermann/Vehicle-Detection-and-Tracking)
- [Vehicle Dataset](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/vehicles.zip) e [Non-Vehicle Dataset](https://s3.amazonaws.com/udacity-sdc/Vehicle_Tracking/non-vehicles.zip)
- [A quick complete tutorial to save and restore Tensorflow models](http://cv-tricks.com/tensorflow-tutorial/save-restore-tensorflow-models-quick-complete-tutorial/)
| github_jupyter |
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import numpy as np
# from embed import Embedding
# embed_file = 'sgns.sogou.word'
EMBED_DIM = 300
def is_valid(seg):
for w in seg:
if not ('\u4e00' <= w and w <= '\u9fff'):
return False
return True
class Embed:
# def __init__(self, file_path='sgns.sogou.word'):
def __init__(self, file_path='../data/sgns.sogou.word'):
self.idx_seg = ['unk']
self.seg_idx = {'unk': 0}
self.idx_emb = [[0.0 for i in range(EMBED_DIM)]]
with open(file_path, 'r') as f:
for idx, line in enumerate(f.readlines(), start=1):
emb = line.split()[1:]
seg = line.split()[0]
# print(emb, seg)
if is_valid(seg) and (seg not in self.seg_idx):
self.idx_seg.append(seg)
self.seg_idx[seg] = idx
self.idx_emb.append([float(i) for i in emb])
def embed(self, seg):
if seg in self.seg_idx:
return self.seg_idx[seg]
else:
return self.seg_idx['unk']
# s = Embed()
# (s.seg_idx[','])
# (s.seg_idx['的'])
# s.embed(',')
# s.embed('我国')
VOCAB_SIZE = 364182
class SeqRNN(nn.Module):
'''
vocab_size:词向量维度
hidden_size:隐藏单元数量决定输出长度
output_size:输出类别为8,维数为1
'''
def __init__(self, vocab_size=300, hidden_size=10, output_size=8, pretrained_embed=Embed().idx_emb):
super(SeqRNN, self).__init__()
self.embed_dim = vocab_size
self.embed = nn.Embedding(VOCAB_SIZE, self.embed_dim)
self.vocab_size = vocab_size # 这个为词向量的维数300
self.hidden_size = hidden_size # 隐藏单元数
self.output_size = output_size # 最后输出size
self.rnn = nn.RNN(self.vocab_size, self.hidden_size,
batch_first=True, dropout=0.5)
self.linear = nn.Linear(self.hidden_size, self.output_size)
def forward(self, input):
input = self.embed(input)
# print(input)
# print('embeded size:', input.shape)
h0 = torch.zeros(1, 1, self.hidden_size)
# print('h0 size:', h0.shape)
output, hidden = self.rnn(input, h0)
output = output[:, -1, :]
output = self.linear(output)
output = torch.nn.functional.softmax(output, dim=1)
return output
# rnn_model = SeqRNN()
# cnn_model = TextCNN()
import torch
import torch.nn as nn
from torch.nn import functional as F
import numpy as np
import json
from torch.utils.data import Dataset, DataLoader
from scipy.stats import pearsonr
from sklearn.metrics import f1_score
import random
weightFile='./pkl/rnn_weight'
train_file='../data/train_dic.json'
test_file='../data/test_dic.json'
with open(train_file,'r') as f:
train_dic = json.load(f)
with open(test_file,'r') as f:
test_dic=json.load(f)
EPOCH=20
BATCH_SIZE=64
lr=0.001
max_len=len(train_dic['label'])
class trainset(Dataset):
def __init__(self):
self.textdata=torch.LongTensor(train_dic['indexed_text'])
self.labeldata=torch.LongTensor(train_dic['emo'])
def __len__(self):
return len(self.textdata)
def __getitem__(self,index):
return self.textdata[index],self.labeldata[index]
class validset(Dataset):
def __init__(self):
self.textdata=torch.LongTensor(test_dic['indexed_text'])
self.labeldata=torch.LongTensor(test_dic['emo'])
def __len__(self):
return len(self.textdata)
def __getitem__(self,index):
return self.textdata[index],self.labeldata[index]
text = trainset()
textloader = DataLoader(dataset=text,batch_size=BATCH_SIZE,shuffle=True)
from tqdm import tqdm
model = SeqRNN()
optimizer = torch.optim.Adam(model.parameters(), lr=lr)
textloader = DataLoader(dataset=text,batch_size=1,shuffle=True)
cnt = 0
calloss = nn.CrossEntropyLoss()
for epoch in range(2):
aveloss = 0
batchnum = 0
for text, label in tqdm(textloader):
batchnum += 1
optimizer.zero_grad()
out = model(text)
loss = calloss(out, label)
loss.backward()
aveloss += loss.item()
optimizer.step()
aveloss /= batchnum
print('Epoch:', epoch, 'aveloss:', aveloss)
torch.save(model.state_dict(), weightFile+str(epoch)+'.pkl')
# FOR TEST
test = validset()
testloader = DataLoader(dataset=test, batch_size=1, shuffle=False)
testmodel = SeqRNN()
# opt=torch.optim.Adam(testmodel.parameters(),lr=LR)
correct = 0
total = 0
epoch = 8
coef = 0
ground = list()
pred = list()
testmodel.load_state_dict(torch.load(weightFile+str(0)+'.pkl'))
for text, label in tqdm(testloader):
testmodel.eval()
out = testmodel(text)
for ind in range(len(out)):
v0 = test_dic['label'][ind][1:]
ol = []
for i in range(len(out[ind])):
ol.append(float(out[ind][i]))
c = pearsonr(ol, v0)
coef += c[0]
prediction = torch.argmax(out, 1)
ground.append(label)
pred.append(prediction)
correct += (prediction == label).sum().float()
total += len(label)
v = np.array(test_dic['emo'])
print(correct)
print(total)
print('acc:', correct.item()/total)
print(coef)
print('Coef:', coef/total)
# tensor(1217.)
# 2228
# acc: 0.546229802513465
# 717.9179559345431
# Coef: 0.3222252944050912
# F-score: 0.18830698287220027
# F-score: 0.29171621217657023
# F-score: 0.24558080808080807
# F-score: 0.1971957671957672
# F-score: 0.13852813852813853
# 0.2035984339260584
tot = 0
cnt = 0
pred_l = list()
true_l = list()
for i, j in zip(ground, pred):
pred_l.append(i.item())
true_l.append(j.item())
tot += f1_score(i.data,j.data,average='macro')
cnt += 1
print('acc:', tot / cnt)
# epoch 1
# acc: 0.46005385996409337
# epoch 0
# acc: 0.45601436265709155
print('micro f1:', f1_score(pred_l,true_l,average='micro'))
print('macro f1:', f1_score(pred_l,true_l,average='macro'))
print('pearson:', pearsonr(pred_l, true_l))
# epoch 1
# micro f1: 0.46005385996409337
# macro f1: 0.09927234898735894
# pearson: (0.04629288575370093, 0.028885338186686736)
for i, j in zip(ground, pred):
print('F-score:',f1_score(i.data,j.data,average='micro'))
tot += f1_score(i.data,j.data,average='micro')
cnt += 1
print(tot / cnt)
for i, j in zip(ground, pred):
print('F-score:',f1_score(i.data,j.data,average='macro'))
tot += f1_score(i.data,j.data,average='macro')
cnt += 1
print(tot / cnt)
```
| github_jupyter |
# **RNN GRU LSTM in PyTorch**
Here we use the sin wave data as in L18/3
```
import torch
import torch.nn as nn
import numpy as np
import torchvision
import torchvision.datasets as datasets
from torchvision import transforms
import torch.optim as optim
import time
import torch.utils.data
# display routines
%matplotlib inline
from matplotlib import pyplot as plt
from IPython import display
display.set_matplotlib_formats('svg')
embedding = 4 # embedding dimension for autoregressive model
T = 1000 # generate a total of 1000 points
time = torch.arange(0.0,T)
x = torch.sin(0.01 * time) + 0.2 * torch.randn(T)
plt.plot(time.numpy(), x.numpy());
```
### Generating the Regression Dataset
```
features = torch.zeros((T-embedding, embedding))
# Use past features at window size 4 to predict 5th time series data
for i in range(embedding):
features[:,i] = x[i:T-embedding+i]
labels = x[embedding:]
ntrain = 600
train_data = torch.utils.data.TensorDataset(features[:ntrain,:], labels[:ntrain])
test_data = torch.utils.data.TensorDataset(features[ntrain:,:], labels[ntrain:])
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
torch.nn.init.xavier_uniform_(m.weight)
elif classname.find('Linear') != -1:
torch.nn.init.xavier_uniform_(m.weight)
elif classname.find('BatchNorm') != -1:
m.weight.data.normal_(1.0, 0.01)
```
## **Model using LSTM cell**
```
class LSTMcell(nn.Module):
def __init__(self, input_size=1, hidden_size=100, output_size=1,
num_layers = 1, batch_size = 16):
super(LSTMcell, self).__init__()
self.lstm = nn.LSTMCell(input_size = 4, hidden_size = hidden_size)
self.fc = nn.Linear(hidden_size, output_size)
self.hidden_size = hidden_size
self.hidden = (torch.zeros(batch_size, hidden_size), #h_0: (batch, hidden_size)
torch.zeros(batch_size, hidden_size)) # c_0: (batch, hidden_size)
def forward(self, x):
# output h_1, c_1
lstm_out, _ = self.lstm(x)
predictions = self.fc(lstm_out.reshape((-1, self.hidden_size)))
return predictions
loss = nn.MSELoss()
```
### Training
```
# simple optimizer using adam, random shuffle and minibatch size 16
def train_net(net, data, loss, epochs, learningrate):
trainer = optim.Adam(net.parameters(), lr = learningrate)
data_iter = torch.utils.data.DataLoader(data, batch_size = batch_size, shuffle=True)
for epoch in range(1, epochs + 1):
net.train()
for X, y in data_iter:
trainer.zero_grad()
# net.hidden_cell = (torch.zeros(16, 10),
# torch.zeros(16, 10))
# if not (X.shape[0] == 16):
# continue
l = loss(net(X.reshape(-1, 4)), y.reshape(-1,1))
l.backward()
trainer.step()
net.eval()
l = loss(net(data[:][0]), data[:][1].reshape(-1,1))
print('epoch %d, loss: %f' % (epoch, l.mean().item()))
return net
batch_size = 16
net = LSTMcell(batch_size = batch_size)
print(net)
net = train_net(net, train_data, loss, 10, 0.01)
l = loss(net(test_data[:][0]), test_data[:][1].reshape(-1,1))
print('test loss: %f' % l.mean().item())
```
### Results
```
print(features.shape)
estimates = net(features)
plt.plot(time.numpy(), x.numpy(), label='data');
plt.plot(time[embedding:].numpy(), estimates.detach().numpy(), label='estimate');
plt.legend();
```
## Predictions for more than 1 step
```
predictions = torch.zeros_like(estimates)
predictions[:(ntrain-embedding)] = estimates[:(ntrain-embedding)]
for i in range(ntrain-embedding, T-embedding):
predictions[i] = net(predictions[(i-embedding):i].reshape(1,-1)).reshape(1)
plt.plot(time.numpy(), x.numpy(), label='data');
plt.plot(time[embedding:].numpy(), estimates.detach().numpy(), label='estimate');
plt.plot(time[embedding:].numpy(), predictions.detach().numpy(), label='multistep');
plt.legend();
k = 33 # look up to k - embedding steps ahead
features = torch.zeros((T-k, k))
for i in range(embedding):
features[:,i] = x[i:T-k+i]
for i in range(embedding, k):
features[:,i] = net(features[:,(i-embedding):i]).reshape((-1))
for i in (4, 8, 16, 32):
plt.plot(time[i:T-k+i].numpy(), features[:,i].detach().numpy(), label=('step ' + str(i)))
plt.legend();
class LSTMcell(nn.Module):
def __init__(self, input_size=1, hidden_size=100, output_size=1,
num_layers = 1, batch_size = 16):
super(LSTMcell, self).__init__()
self.lstm = nn.LSTMCell(input_size = 4, hidden_size = hidden_size)
self.fc = nn.Linear(hidden_size, output_size)
self.hidden_size = hidden_size
self.hidden = (torch.zeros(batch_size, hidden_size), #h_0: (batch, hidden_size)
torch.zeros(batch_size, hidden_size)) # c_0: (batch, hidden_size)
def forward(self, x):
# output h_1, c_1
lstm_out, _ = self.lstm(x)
predictions = self.fc(lstm_out.reshape((-1, self.hidden_size)))
return predictions
loss = nn.MSELoss()
```
## **Model using LSTM function**
Compared to LSTMcell, LSTM are optimized with cuDNN.
```
class LSTM(nn.Module):
def __init__(self, input_size=1, hidden_size=100, output_size=1,
num_layers = 1, batch_size = 16):
super(LSTM, self).__init__()
self.lstm = nn.LSTM(input_size = 4, hidden_size = hidden_size)
self.fc = nn.Linear(hidden_size, output_size)
self.hidden_size = hidden_size
# h_0: (num_layers * num_directions, batch, hidden_size)
# c_0: (num_layers * num_directions, batch, hidden_size)
self.hidden = (torch.zeros(1, batch_size, hidden_size),
torch.zeros(1, batch_size, hidden_size))
def forward(self, x):
# output h_1, c_1
# print(x.shape)
lstm_out, _ = self.lstm(x, self.hidden)
predictions = self.fc(lstm_out.reshape((-1, self.hidden_size)))
return predictions
loss = nn.MSELoss()
# simple optimizer using adam, random shuffle and minibatch size 16
def train_net(net, data, loss, epochs, learningrate):
trainer = optim.Adam(net.parameters(), lr = learningrate)
data_iter = torch.utils.data.DataLoader(data, batch_size = batch_size, shuffle=True)
for epoch in range(1, epochs + 1):
net.train()
for X, y in data_iter:
trainer.zero_grad()
# net.hidden_cell = (torch.zeros(16, 10),
# torch.zeros(16, 10))
# if not (X.shape[0] == 16):
# continue
X = X.reshape(1, -1, 4)
if not X.shape[1] == batch_size:
continue
l = loss(net(X), y.reshape(-1,1))
l.backward()
trainer.step()
net.eval()
train_l = 0
iters = 0
for X, y in data_iter:
X = X.reshape(1, -1, 4)
if not X.shape[1] == batch_size:
continue
train_l += loss(net(X), y.reshape(-1,1)).mean().item()
iters += 1
print('epoch %d, loss: %f' % (epoch, train_l/iters))
return net
batch_size = 16
net = LSTM(batch_size = batch_size)
print(net)
net = train_net(net, train_data, loss, 10, 0.01)
test_l = 0
iters = 0
test_data_iter = torch.utils.data.DataLoader(test_data, batch_size = batch_size, shuffle=True)
for X, y in test_data_iter:
X = X.reshape(1, -1, 4)
if not X.shape[1] == batch_size:
continue
test_l += loss(net(X), y.reshape(-1,1)).mean().item()
iters += 1
print('test loss: {}'.format(test_l/iters))
```
| github_jupyter |
# Encoder-Decoder model for ENSO-forecasting
The Encoder-Decoder model is enspired by the architecture of autoencoders. Here, the size of the input layer of the neural network is the same size as the output layer. Furthermore, there is a so-called bottleneck layer present in the network which has less neurons than the input/output layers. For an Autoencoder the label and feature are the same. Because of the bottleneck layer, the network is forced to learn a lower dimesional expression of the input
For the Encoder-Decoder model (DEM), a time lag between the feature and the label is introduced. Such that the DEM is a forecast model. Hence, the DEM is forced to learn a lower dimensional expression that explains best the future state of the considered variable field.
<img src="AE_example.png" alt="Drawing" style="width: 400px;"/>
In this tutorial, the considered variable field is again the SST anomaly data from the ERSSTv5 data set.
## Read data
In the following cell, we read the SST anoamly data that was computed in an earlier tutorial.
```
from ninolearn.IO.read_processed import data_reader
#read data
reader = data_reader(startdate='1959-11', enddate='2018-12')
# read SST data and directly make seasonal averages
SSTA = reader.read_netcdf('sst', dataset='ERSSTv5', processed='anom').rolling(time=3).mean()[2:]
# read the ONI for later comparison
oni = reader.read_csv('oni')[2:]
```
## Generate the feature and label arrays
The following cell generates the feature and label arrays. Because `feature` and `label` need to refer to an other object in the backend so that they can be changed without influencing each other.
Moreover, the data is scalled because this helps the DEM to be more precise.
```
from sklearn.preprocessing import StandardScaler
import numpy as np
# make deep copies of the sst data
feature = SSTA.copy(deep=True)
label = SSTA.copy(deep=True)
# reshape the data such that one time step is a 1-D vector
# i.e. the feature_unscaled array is hence 2-D (timestep, number of grid points)
feature_unscaled = feature.values.reshape(feature.shape[0],-1)
label_unscaled = label.values.reshape(label.shape[0],-1)
# scale the data
scaler_f = StandardScaler()
Xorg = scaler_f.fit_transform(feature_unscaled)
scaler_l = StandardScaler()
yorg = scaler_l.fit_transform(label_unscaled)
# replace nans with 0
Xall = np.nan_to_num(Xorg)
yall = np.nan_to_num(yorg)
# shift = 3 is needed to align with the usual way how lead time is defined
shift = 3
# the lead time
lead = 3
y = yall[lead+shift:]
X = Xall[:-lead-shift]
timey = oni.index[lead+shift:]
```
## Split the data set
For the training and testing of machine learning models it is crucial to split the data set into:
1. __Train data set__ which is used to train the weights of the neural network
2. __Validation data set__ which is used to check for overfitting (e.g. when using early stopping) and to optimize the hyperparameters
3. __Test data set__ which is used to to evaluate the trained model.
__NOTE:__ It is important to understand that hyperparamters must be tuned so that the result is best for the Validation data set and __not__ for the test data set. Otherwise you can not rule out the case that the specific hyperparameter setting just works good for the specific test data set but is not generally a good hyperparameter setting.
In the following cell the train and the validation data set are still one data set, because this array will be later splitted into two arrays when th model is fitted.
```
test_indeces = test_indeces = (timey>='2001-01-01') & (timey<='2018-12-01')
train_val_indeces = np.invert(test_indeces)
train_val_X, train_val_y, train_val_timey = X[train_val_indeces,:], y[train_val_indeces,:], timey[train_val_indeces]
testX, testy, testtimey = X[test_indeces,:], y[test_indeces,:], timey[test_indeces]
```
## Fit the model
Let's train the model using a random search. This takes quite some time (for `n_iter=100` it took an hour on my local machine without GPU support). In this training process the train/validation data set will be splitted In this example below this train/validation data is first divided into 5 segments (`n_segments=5`). One segment alwawys serves as the validation data set and the rest as training. Each segment is one time (`n_members_segment=1`) the validation data set. Hence, the ensemble consists in the end out of 5 models.
```
import keras.backend as K
from ninolearn.learn.models.encoderDecoder import EncoderDecoder
K.clear_session()
model = EncoderDecoder()
model.set_parameters(neurons=[(32, 8), (512, 64)], dropout=[0., 0.2], noise=[0., 0.5] , noise_out=[0., 0.5],
l1_hidden=[0.0, 0.001], l2_hidden=[0.0, 0.001], l1_out=[0.0, 0.001], l2_out=[0.0, 0.001], batch_size=100,
lr=[0.0001, 0.01], n_segments=5, n_members_segment=1, patience = 40, epochs=1000, verbose=0)
model.fit_RandomizedSearch(train_val_X, train_val_y, n_iter=10)
```
## Make predictions on the test data set
```
import xarray as xr
# make prediction
pred, pred_all_members = model.predict(testX)
test_label = scaler_l.inverse_transform(testy)
# reshape data into an xarray data array that is 3-D
pred_map = xr.zeros_like(label[lead+shift:,:,:][test_indeces])
pred_map.values = scaler_l.inverse_transform(pred).reshape((testX.shape[0], label.shape[1], label.shape[2]))
test_label_map = xr.zeros_like(label[lead+shift:,:,:][test_indeces])
test_label_map.values = test_label.reshape((testX.shape[0], label.shape[1], label.shape[2]))
# calculate the ONI
pred_oni = pred_map.loc[dict(lat=slice(-5, 5), lon=slice(190, 240))].mean(dim="lat").mean(dim='lon')
```
## Plot prediction for the ONI
```
import matplotlib.pyplot as plt
# Plot ONI Forecasts
plt.subplots(figsize=(8,1.8))
plt.plot(testtimey, pred_oni, c='navy')
plt.plot(testtimey, oni.loc[testtimey[0]: testtimey[-1]], "k")
plt.xlabel('Time [Year]')
plt.ylabel('ONI [K]')
plt.axhspan(-0.5, -6, facecolor='blue', alpha=0.1,zorder=0)
plt.axhspan(0.5, 6, facecolor='red', alpha=0.1,zorder=0)
plt.xlim(testtimey[0], testtimey[-1])
plt.ylim(-3,3)
plt.title(f"Lead time: {lead} month")
plt.grid()
```
## Evaluate the seasonal skill for predicting the ONI
```
from ninolearn.plot.evaluation import plot_correlation
import pandas as pd
# make a plot of the seasonal correaltion
# note: - pd.tseries.offsets.MonthBegin(1) appears to ensure that the correlations are plotted
# agains the correct season
plot_correlation(oni.loc[testtimey[0]: testtimey[-1]], pred_oni, testtimey - pd.tseries.offsets.MonthBegin(1), title="")
```
## Check the correlations on a map
In the following the pearson correlation coefficent between the predicted and the observed value for each grid point are computed and afterwards plotted.
```
# rehshape predictions to a map
corr_map = np.zeros(pred.shape[1])
for j in range(len(corr_map)):
corr_map[j] = np.corrcoef(pred[:,j], test_label[:,j])[0,1]
corr_map = corr_map.reshape((label.shape[1:]))
%matplotlib inline
# Plot correlation map
fig, ax = plt.subplots(figsize=(8,2))
plt.title(f"Lead time: {lead} month")
C=ax.imshow(corr_map, origin='lower', vmin=0, vmax=1)
cb=plt.colorbar(C)
```
## Animate the full test prediction
```
import matplotlib.animation as animation
def animation_ed(true, pred, nino, nino_pred, time):
fig, ax = plt.subplots(3, 1, figsize=(6,7), squeeze=False)
vmin = -3
vmax = 3
true_im = ax[0,0].imshow(true[0], origin='lower', vmin=vmin, vmax=vmax, cmap=plt.cm.bwr)
pred_im = ax[1,0].imshow(pred[0], origin='lower', vmin=vmin, vmax=vmax, cmap=plt.cm.bwr)
title = ax[0,0].set_title('')
ax[2,0].plot(time, nino)
ax[2,0].plot(time, nino_pred)
ax[2,0].set_ylim(-3,3)
ax[2,0].set_xlim(time[0], time[-1])
vline = ax[2,0].plot([time[0], time[0]], [-10,10], color='k')
def update(data):
true_im.set_data(data[0])
pred_im.set_data(data[1])
title_str = np.datetime_as_string(data[0].time.values)[:10]
title.set_text(title_str)
vline[0].set_data([data[2], data[2]],[-10,10])
def data_gen():
k = 0
kmax = len(true)
while k<kmax:
yield true.loc[time[k]], pred.loc[time[k]], time[k]
k+=1
ani = animation.FuncAnimation(fig, update, data_gen, interval=100, repeat=True, save_count=len(true))
plt.close("all")
return ani
from IPython.display import HTML
ani = animation_ed(test_label_map, pred_map,
oni.loc[testtimey[0]:testtimey[-1]], pred_oni,
testtimey)
HTML(ani.to_html5_video())
```
So it looks like that the Encoder-Decoder ensemble has some skill. However, more research is needed to release the full potential of this model.
Maybe you are interested working on this? :D
| github_jupyter |
```
from pyspark.context import SparkContext, SparkConf
from awsglue.dynamicframe import DynamicFrame
import awsglue.transforms as T
# this is just so i can develop with local glue libs
jars = '/Users/joe/aws-glue-libs/jarsv1/*'
sc = SparkContext(conf=SparkConf().setAll([
('spark.executor.extraClassPath', jars),
('spark.driver.extraClassPath', jars)
]))
from awsglue.context import GlueContext
glueContext = GlueContext(sc)
spark = glueContext.spark_session
```
### Read in the data
Use a dynamicframe to deal with null values or variable types
```
data = spark.read.parquet('data/catalog.parquet')
datasource = DynamicFrame.fromDF(data, glueContext, 'datasource')
```
### Grab the Location records
This is the primary source of locations but there might be others in other resources.
```
locations = datasource.filter(
lambda r: r['resourceType'] == 'Location'
)
locations = locations.select_fields(
['identifier','name','type','address','position']
)
locations = T.DropNullFields.apply(locations, "locations")
```
Let's have a look to see where we can find addresses
```
addresses = datasource.filter(
lambda r: r['address']
)
addresses = T.DropNullFields.apply(addresses, "addresses")
df = addresses.toDF()
df.select('resourceType').distinct().show()
```
So we can see there are more locations in **Patient**, **Organization** and **Practitioner**
Let's have a look at the schema:
```
df.select('address').printSchema()
```
So there's a zero or more cardinality and it's either a struct or an array of structs.
We need to normalize this data using the `explode()` method
```
import pyspark.sql.functions as F
df = df.withColumn('address', F.explode(df.address.array))
df.select('address').printSchema()
loc = locations.toDF()
loc = loc.withColumn('address', F.col('address.struct'))
loc.select('address').printSchema()
```
Next we need to join the dataframes, keeping only the relevant fields.
For this we could use an **left outer-join** on the `loc` dataframe.
```
# drop irrelevant fields from struct
def filter_columns(df, root):
cols = df.select(root).columns
fields = filter(lambda x: x in ['postalCode', 'city', 'country', 'state', 'line'], cols)
kept = list(map(lambda x: root[:-1] + x, fields))
return df.select(kept)
other_address = filter_columns(df.select('address'), root='address.*')
loc_address = filter_columns(df.select('address'), root='address.*')
```
We now have the same schema, ready to be combined into one dataframe
```
other_address.printSchema()
loc_address.printSchema()
new_df = loc_address.join(other_address, on=['postalCode','city','country','state','line'], how='left_outer')
new_df.printSchema()
new_df = new_df.drop_duplicates()
new_df.show()
df = new_df
# care_sites = df.na.drop(subset=["type"])
```
TODO:
- [ ] Make identifier based on hash of row
- [ ] Transform to CareSite table based on type
```
df.show()
df.printSchema()
df = df.withColumn('city', F.col('address.struct.city'))\
.withColumn('state', F.col('address.struct.state'))\
.withColumn('zip', F.col('address.struct.postalCode'))\
.withColumn('country', F.col('address.struct.country'))
# df = df.withColumn('exploded', F.explode('address.struct.line'))
df = df.withColumn('address_1', F.col('address.struct.line').getItem(0))
df = df.withColumn('address_2', F.col('address.struct.line').getItem(1))
df = df.withColumnRenamed('id', 'location_id')
df = df.drop(*['address','position','exploded','name','type'])
df.show()
locations = DynamicFrame.fromDF(df, glueContext, 'locations')
```
| github_jupyter |
Copyright (c) Microsoft Corporation. All rights reserved.
Licensed under the MIT License.

# Automated Machine Learning
_**Classification of credit card fraudulent transactions on remote compute **_
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Train](#Train)
1. [Results](#Results)
1. [Test](#Test)
1. [Acknowledgements](#Acknowledgements)
## Introduction
In this example we use the associated credit card dataset to showcase how you can use AutoML for a simple classification problem. The goal is to predict if a credit card transaction is considered a fraudulent charge.
This notebook is using remote compute to train the model.
If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../../../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
In this notebook you will learn how to:
1. Create an experiment using an existing workspace.
2. Configure AutoML using `AutoMLConfig`.
3. Train the model using remote compute.
4. Explore the results.
5. Test the fitted model.
## Setup
As part of the setup you have already created an Azure ML `Workspace` object. For Automated ML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import logging
from matplotlib import pyplot as plt
import pandas as pd
import os
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
```
print("This notebook was created using version 1.36.0 of the Azure ML SDK")
print("You are currently using version", azureml.core.VERSION, "of the Azure ML SDK")
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = 'automl-classification-ccard-remote'
experiment=Experiment(ws, experiment_name)
output = {}
output['Subscription ID'] = ws.subscription_id
output['Workspace'] = ws.name
output['Resource Group'] = ws.resource_group
output['Location'] = ws.location
output['Experiment Name'] = experiment.name
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
```
## Create or Attach existing AmlCompute
A compute target is required to execute the Automated ML run. In this tutorial, you create AmlCompute as your training compute resource.
> Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
#### Creation of AmlCompute takes approximately 5 minutes.
If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster-1"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print('Found existing cluster, use it.')
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(vm_size='STANDARD_DS12_V2',
max_nodes=6)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
# Data
### Load Data
Load the credit card dataset from a csv file containing both training features and labels. The features are inputs to the model, while the training labels represent the expected output of the model. Next, we'll split the data using random_split and extract the training data for the model.
```
data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/creditcard.csv"
dataset = Dataset.Tabular.from_delimited_files(data)
training_data, validation_data = dataset.random_split(percentage=0.8, seed=223)
label_column_name = 'Class'
```
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression|
|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
|**enable_early_stopping**|Stop the run if the metric score is not showing improvement.|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
**_You can find more information about primary metrics_** [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
```
automl_settings = {
"n_cross_validations": 3,
"primary_metric": 'AUC_weighted',
"enable_early_stopping": True,
"max_concurrent_iterations": 2, # This is a limit for testing purpose, please increase it as per cluster size
"experiment_timeout_hours": 0.25, # This is a time limit for testing purposes, remove it for real use cases, this will drastically limit ablity to find the best model possible
"verbosity": logging.INFO,
}
automl_config = AutoMLConfig(task = 'classification',
debug_log = 'automl_errors.log',
compute_target = compute_target,
training_data = training_data,
label_column_name = label_column_name,
**automl_settings
)
```
Call the `submit` method on the experiment object and pass the run configuration. Depending on the data and the number of iterations this can run for a while. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output = False)
# If you need to retrieve a run that already started, use the following code
#from azureml.train.automl.run import AutoMLRun
#remote_run = AutoMLRun(experiment = experiment, run_id = '<replace with your run id>')
```
## Results
#### Widget for Monitoring Runs
The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details
```
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
remote_run.wait_for_completion(show_output=False)
```
#### Explain model
Automated ML models can be explained and visualized using the SDK Explainability library.
## Analyze results
### Retrieve the Best Model
Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
```
best_run, fitted_model = remote_run.get_output()
fitted_model
```
#### Print the properties of the model
The fitted_model is a python object and you can read the different properties of the object.
## Test the fitted model
Now that the model is trained, split the data in the same way the data was split for training (The difference here is the data is being split locally) and then run the test data through the trained model to get the predicted values.
```
# convert the test data to dataframe
X_test_df = validation_data.drop_columns(columns=[label_column_name]).to_pandas_dataframe()
y_test_df = validation_data.keep_columns(columns=[label_column_name], validate=True).to_pandas_dataframe()
# call the predict functions on the model
y_pred = fitted_model.predict(X_test_df)
y_pred
```
### Calculate metrics for the prediction
Now visualize the data on a scatter plot to show what our truth (actual) values are compared to the predicted values
from the trained model that was returned.
```
from sklearn.metrics import confusion_matrix
import numpy as np
import itertools
cf =confusion_matrix(y_test_df.values,y_pred)
plt.imshow(cf,cmap=plt.cm.Blues,interpolation='nearest')
plt.colorbar()
plt.title('Confusion Matrix')
plt.xlabel('Predicted')
plt.ylabel('Actual')
class_labels = ['False','True']
tick_marks = np.arange(len(class_labels))
plt.xticks(tick_marks,class_labels)
plt.yticks([-0.5,0,1,1.5],['','False','True',''])
# plotting text value inside cells
thresh = cf.max() / 2.
for i,j in itertools.product(range(cf.shape[0]),range(cf.shape[1])):
plt.text(j,i,format(cf[i,j],'d'),horizontalalignment='center',color='white' if cf[i,j] >thresh else 'black')
plt.show()
```
## Acknowledgements
This Credit Card fraud Detection dataset is made available under the Open Database License: http://opendatacommons.org/licenses/odbl/1.0/. Any rights in individual contents of the database are licensed under the Database Contents License: http://opendatacommons.org/licenses/dbcl/1.0/ and is available at: https://www.kaggle.com/mlg-ulb/creditcardfraud
The dataset has been collected and analysed during a research collaboration of Worldline and the Machine Learning Group (http://mlg.ulb.ac.be) of ULB (Université Libre de Bruxelles) on big data mining and fraud detection.
More details on current and past projects on related topics are available on https://www.researchgate.net/project/Fraud-detection-5 and the page of the DefeatFraud project
Please cite the following works:
Andrea Dal Pozzolo, Olivier Caelen, Reid A. Johnson and Gianluca Bontempi. Calibrating Probability with Undersampling for Unbalanced Classification. In Symposium on Computational Intelligence and Data Mining (CIDM), IEEE, 2015
Dal Pozzolo, Andrea; Caelen, Olivier; Le Borgne, Yann-Ael; Waterschoot, Serge; Bontempi, Gianluca. Learned lessons in credit card fraud detection from a practitioner perspective, Expert systems with applications,41,10,4915-4928,2014, Pergamon
Dal Pozzolo, Andrea; Boracchi, Giacomo; Caelen, Olivier; Alippi, Cesare; Bontempi, Gianluca. Credit card fraud detection: a realistic modeling and a novel learning strategy, IEEE transactions on neural networks and learning systems,29,8,3784-3797,2018,IEEE
Dal Pozzolo, Andrea Adaptive Machine learning for credit card fraud detection ULB MLG PhD thesis (supervised by G. Bontempi)
Carcillo, Fabrizio; Dal Pozzolo, Andrea; Le Borgne, Yann-Aël; Caelen, Olivier; Mazzer, Yannis; Bontempi, Gianluca. Scarff: a scalable framework for streaming credit card fraud detection with Spark, Information fusion,41, 182-194,2018,Elsevier
Carcillo, Fabrizio; Le Borgne, Yann-Aël; Caelen, Olivier; Bontempi, Gianluca. Streaming active learning strategies for real-life credit card fraud detection: assessment and visualization, International Journal of Data Science and Analytics, 5,4,285-300,2018,Springer International Publishing
Bertrand Lebichot, Yann-Aël Le Borgne, Liyun He, Frederic Oblé, Gianluca Bontempi Deep-Learning Domain Adaptation Techniques for Credit Cards Fraud Detection, INNSBDDL 2019: Recent Advances in Big Data and Deep Learning, pp 78-88, 2019
Fabrizio Carcillo, Yann-Aël Le Borgne, Olivier Caelen, Frederic Oblé, Gianluca Bontempi Combining Unsupervised and Supervised Learning in Credit Card Fraud Detection Information Sciences, 2019
| github_jupyter |
```
from sklearn.model_selection import train_test_split
import pandas as pd
import scripts.classification.classifier as clfs
from scripts.text.skeleton import Skeleton
from scripts.text.textutilities import *
pd.set_option('mode.chained_assignment', None)
categories=['toxic','severe_toxic','obscene','insult','identity_hate','clean']
RANDOM_STATE=20
MODEL_PATH="./word2vec/word2vec.model"
BOOSTING_CLASSIFIER_COUNT=100
skeleton=Skeleton(categories,RANDOM_STATE)
data={}
for key in categories:
data[key]=pd.read_csv('./Data/{}.csv'.format(key))
X_x={}
X_y={}
y_x={}
y_y={}
X = []
y=[]
for key in categories:
X_x[key], y_x[key], X_y[key], y_y[key] = train_test_split(data[key]['comment_text'], data[key][key], test_size=0.2,shuffle=True, random_state=RANDOM_STATE)
X+=list(X_x[key])
y+=list(X_y[key])
print("Created test-train data.")
skeleton.classify([
# clfs.AdaBoostNaiveBayes(n_estimators=2,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=3,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=3,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=3,lr=0.25),
# clfs.AdaBoostNaiveBayes(n_estimators=2,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=5,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=20,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=20,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=100,lr=0.25),
# clfs.AdaBoostNaiveBayes(n_estimators=100,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=100,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=250,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=500,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=500,lr=0.5),
# clfs.AdaBoostNaiveBayes(n_estimators=1000,lr=1),
# clfs.AdaBoostNaiveBayes(n_estimators=1000,lr=0.5),
clfs.AdaBoostDecisionTree(n_estimators=2,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=2,max_tree_depth=1,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=2,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=3,max_tree_depth=2,lr=0.5),
clfs.AdaBoostDecisionTree(n_estimators=3,max_tree_depth=2,lr=0.25),
clfs.AdaBoostDecisionTree(n_estimators=3,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=3,max_tree_depth=2,lr=3),
clfs.AdaBoostDecisionTree(n_estimators=10,max_tree_depth=3,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=15,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=20,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=2,lr=1),
clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=2,lr=2),
clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=2,lr=0.5),
# clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=1,lr=0.1),
# clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=1,lr=0.5),
# clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=3,lr=0.25),
# clfs.AdaBoostSVM(n_estimators=3),
# clfs.AveragingEstimator(),
# clfs.DecisionTree(),
# clfs.NaiveBayes(),
# clfs.SVM()
# clfs.AdaBoostDecisionTree(n_estimators=500,max_tree_depth=2,lr=0.5),
# clfs.AdaBoostDecisionTree(n_estimators=250,max_tree_depth=4,lr=1),
# clfs.AdaBoostDecisionTree(n_estimators=500,max_tree_depth=4,lr=0.1),
# clfs.AdaBoostDecisionTree(n_estimators=500,max_tree_depth=8,lr=0.25),
# clfs.AdaBoostDecisionTree(n_estimators=500,max_tree_depth=8,lr=1),
# clfs.AdaBoostDecisionTree(n_estimators=1000,max_tree_depth=2,lr=0.5),
# clfs.AdaBoostDecisionTree(n_estimators=1000,max_tree_depth=2,lr=0.25)
],X,y,y_x,y_y)
skeleton.save_progress("adaboostdecision_1.txt")
```
| github_jupyter |
# Tutorial Title
A brief introduction to the tutorial that describes:
- The problem that the tutorial addresses
- Who the intended audience is
- The expected experience level of that audience with a concept or tool
- Which environment/language it runs in
If there is another similar tutorial that's more appropriate for another audience, direct the reader there with a linked reference.
## How to Use This Tutorial
A brief explanation of how the reader can use the tutorial. Can the reader copy each code snippet into a Python or other environment? Or can the reader run `<filename>` before or after reading through the explanations to understand how the code works?
You can use this tutorial by *insert method(s) here*.
A bulleted list of the tasks the reader will accomplish and skills he or she will learn. Begin each list item with a noun (Learn, Create, Use, etc.).
You will accomplish the following:
- First task or skill
- Second task or skill
- X task or skill
## Prerequisites
Provide a *complete* list of the software, hardware, knowledge, and skills required to be successful using the tutorial. For each item, link the item to installation instructions, specs, or skill development tools, as appropriate. If good installation instructions aren't available for required software, start the tutorial with instructions for installing it.
To complete this tutorial, you need:
- [MXNet](https://mxnet.incubator.apache.org/install/#overview)
- [Language](https://mxnet.incubator.apache.org/tutorials/)
- [Tool](https://mxnet.incubator.apache.org/api/python/index.html)
- [Familiarity with concept or tool](https://gluon.mxnet.io/)
## The Data
Provide a link to where the data is hosted and explain how to download it. If it requires more than two steps, use a numbered list.
You can download the data used in this tutorial from the [Site Name](http://) site. To download the data:
1. At the `<language>` prompt, type:
`<command>`
2. Second task.
3. Last task.
Briefly describe key aspects of the data. If there are two or more aspects of the data that require involved discussion, use subheads (### `<Concept or Sub-component Name>`). To include a graphic, introduce it with a brief description and use the image linking tool to include it. Store the graphic in GitHub and use the following format: <img width="517" alt="screen shot 2016-05-06 at 10 13 16 pm" src="https://cloud.githubusercontent.com/assets/5545640/15089697/d6f4fca0-13d7-11e6-9331-7f94fcc7b4c6.png">. You do not need to provide a title for your graphics.
The data *add description here. (optional)*
## (Optional) Concept or Component Name
If concepts or components need further introduction, include this section. If there are two or more aspects of the concept or component that require involved discussion, use subheads (### Concept or Sub-component Name).
## Prepare the Data
If appropriate, summarize the tasks required to prepare the data, defining and explaining key concepts.
To prepare the data, *provide explanation here.*
Use a numbered procedure to explain how to prepare the data. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
To prepare the data:
1.
2.
3.
If there are any aspects of data preparation that require elaboration, add it here.
## Create the Model
If appropriate, summarize the tasks required to create the model, defining and explaining key concepts.
To create the model, *provide explanation here.*
Use a numbered procedure to explain how to create the data. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
To create the model:
1.
2.
3.
If there are any aspects of model creation that require elaboration, add it here.
## Fit the Model
If appropriate, summarize the tasks required to fit the model, defining and explaining key concepts.
To fit the model, *provide explanation here.*
Use a numbered procedure to explain how to fit the model. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
To fit the model:
1.
2.
3.
If there are any aspects of model fitting that require elaboration, add it here.
## Evaluate the Model
If appropriate, summarize the tasks required to evaluate the model, defining and explaining key concepts.
To evaluate the model, *provide explanation here.*
Use a numbered procedure to explain how to evaluate the model. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
To evaluate the model:
1.
2.
3.
If there are any aspects of model evaluation that require elaboration, add it here.
## (Optional) Additional Tasks
If appropriate, summarize the tasks required to perform the task, defining and explaining key concepts.
To *perform the task*, *provide explanation here.*
Use a numbered procedure to explain how to perform the task. Add code snippets or blocks that show the code that the user must type or that is used for this task in the Jupyter Notebook. To include code snippets, precede each line of code with four spaces and two tick marks. Always introduce input or output with a description or context or result, followed by a colon.
To *perform the task*:
1.
2.
3.
If there are any aspects of model evaluation that require elaboration, add it here.
## Summary
Briefly describe the end result of the tutorial and how the user can use it or modify it to customize it.
## Next Steps
Provide a bulleted list of other documents, tools, or tutorials that further explain the concepts discussed in this tutorial or build on this tutorial. Start each list item with a brief description of a user task followed by the title of the destination site or topic that is formatted as a link.
- For more information on *topic*, see [Site Name](http://).
- To learn more about using *tool or task*, see [Topic Title](http://).
- To experiment with *service*, *tool*, or *object*, see [Site Name](http://).
- For a more advanced tutorial on *subject*, see [Tutorial Title](http://).
| github_jupyter |
```
import os, os.path
os.chdir('../')
import pandas as pd
import numpy as np
import re
from collections import Counter
from keras.models import Sequential
from keras.layers import Dense
import jieba
import jieba.analyse
```
### Read the Data
```
data = pd.read_csv('data/processed_data.csv')
data.head()
```
### Vectorization
```
text1 = list(data['text1'])
text2 = list(data['text2'])
labels = list(data['label'])
assert len(text1) == len(text2)
texts = text1 + text2
tokens = []
for text in texts:
for sentence in re.findall(r'\w+', text):
for i in range(len(sentence)-1):
word = sentence[i:i+2]
tokens.append(word)
counter_2 = Counter(tokens)
most_common_2 = [word[0] for word in counter_2.most_common(30)]
tokens_3 = []
texts = data['text1']
for text in texts:
for sentence in re.findall(r'\w+', text):
for i in range(len(sentence)-2):
word = sentence[i:i+3]
tokens_3.append(word)
counter_3 = Counter(tokens_3)
most_common_3 = [word[0] for word in counter_3.most_common(30)]
most_common_2
word = most_common_2[0]
word
keywords = most_common_2 + most_common_3
word_vector = np.zeros((len(text1), 2*len(keywords)))
for i, word in enumerate(keywords):
ip = i + len(keywords)
for j in range(len(word_vector)):
if word in text1[j]:
word_vector[j, i] = 1
if word in text2[j]:
word_vector[j, ip] = 1
```
## Key word extraction
```
def get_key_word(sentence,topK):
return jieba.analyse.extract_tags(sentence,topK = topK)
def get_all_key_word(sentences,topK):
res = []
for i in range(len(sentences)):
key_word = get_key_word(sentences[i],topK)
res.extend(key_word)
return res
# add dictionary words
added_words = ['花呗','借呗','怎么','什么','我的']
for i in added_words:
jieba.add_word(i)
counter_key = Counter(get_all_key_word(text1,5)+get_all_key_word(text2,5))
most_common_key = [word[0] for word in counter_key.most_common(120)]
all_key = [word for word in counter_key] #7000+
#### get training data
word_vector = np.zeros((len(text1), 2*len(most_common_key)))
for i, word in enumerate(most_common_key):
ip = i + len(most_common_key)
for j in range(len(word_vector)):
if word in text1[j]:
word_vector[j, i] = 1
if word in text2[j]:
word_vector[j, ip] = 1
```
## Training
```
data = pd.DataFrame(word_vector)
input_dim = word_vector.shape[1]
data_ = np.array(data)
labels = np.array(labels)
model = Sequential()
model.add(Dense(64, input_dim=input_dim, activation='relu'))
model.add(Dense(16, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
model.fit(data_, labels, epochs=15, batch_size=10)
scores = model.evaluate(data_, labels)
print("\n%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
```
| github_jupyter |
# Introduction to `awesimsoss`
**Advanced Webb Exposure Simulator for SOSS**
```
%matplotlib inline
%load_ext autoreload
%autoreload 2
import numpy as np
import astropy.units as q
import astropy.constants as ac
from bokeh.io import output_notebook
from bokeh.plotting import figure, show
import batman
from pkg_resources import resource_filename
from awesimsoss import awesim
output_notebook()
```
## M Dwarf (no planet)
Here is how to generate time series observations of a brown dwarf (or any other isolated star with no transiting planet).
We need two components to generate this simulation:
- A flux calibrated stellar spectrum
- A specified number of integrations and groups for the observation
Let's use this 3500K stellar spectrum with a J magnitude of 9.
```
# Get the wavelength and flux of the star with units
star = np.genfromtxt(resource_filename('awesimsoss','files/scaled_spectrum.txt'), unpack=True)
star_wave, star_flux = [star[0][:1000]*q.um, (star[1][:1000]*q.W/q.m**2/q.um).to(q.erg/q.s/q.cm**2/q.AA)]
# Plot it
fig = figure(width=600, height=300, title='3500 K star with Jmag=9')
fig.line(star_wave, star_flux, legend='Input Spectrum')
fig.xaxis.axis_label = 'Wavelength [um]'
fig.yaxis.axis_label = 'Flux Density [erg/s/cm3/A]'
show(fig)
```
Now we can intialize the simulation by passing the number of groups (3) and integrations (10) along with the stellar spectrum to the `TSO` class.
```
# Initialize the simulation with 3 groups and 10 integrations
my_TSO = awesim.TSO(ngrps=2, nints=2, star=[star_wave, star_flux], target='WASP-107')
# Run the simulation (takes ~10 seconds)
my_TSO.simulate()
```
We can view frames of the simulation with the `plot` method like so:
```
# plot the TSO object
my_TSO.plot()
```
## M Dwarf (with planet)
Let's pretend this M dwarf is orbited by WASP107b! Why not? First get the transmission spectrum:
```
# Get the planet data
planet = np.genfromtxt(resource_filename('awesimsoss', '/files/WASP107b_pandexo_input_spectrum.dat'), unpack=True)
planet_wave, planet_trans = [planet[0]*q.um, planet[1]]
# Plot it
fig = figure(width=600, height=300, title='Planetary Transit Spectrum')
fig.line(planet_wave, planet_trans, legend='Input Transmission')
fig.xaxis.axis_label = 'Wavelength [um]'
fig.yaxis.axis_label = 'Transit Depth [ppm]'
show(fig)
# Set the orbital parameters with the Batman package (https://www.cfa.harvard.edu/~lkreidberg/batman/quickstart.html)
params = batman.TransitParams()
params.t0 = 0.001 # Time of inferior conjunction (days)
params.per = 0.03 # Orbital period (days)
params.rp = 0.15 # Planet radius (in units of R*)
params.a = 0.0558*q.AU.to(ac.R_sun)*0.66 # Semi-major axis (in units of R*)
params.inc = 89.8 # Orbital inclination (in degrees)
params.ecc = 0. # Eccentricity
params.w = 90. # Longitude of periastron (in degrees)
params.u = [0.1, 0.1] # Limb darkening coefficients [u1, u2]
params.limb_dark = "quadratic" # Limb darkening model
# Make the transit model and add the stellar params
tmodel = batman.TransitModel(params, my_TSO.time.jd)
tmodel.teff = 3500 # Effective temperature of the host star
tmodel.logg = 5 # log surface gravity of the host star
tmodel.feh = 0 # Metallicity of the host star
# Run the simulation, this time including the planet
my_TSO.simulate(planet=[planet_wave, planet_trans], tmodel=tmodel)
```
## Exporting Results
Create a fits file with your time series observations with ``export`` like so:
```
my_TSO.export('my_soss_simulation.fits')
```
| github_jupyter |
# Data Cleaning and Feature Extraction of ` Phishing URLs `
```
#importing required packages for this module
import pandas as pd
```
# **2.1. Phishing URLs:**
```
#loading the phishing URLs data to dataframe
phishurl = pd.read_csv("3-phishurl.csv")
phishurl.head()
```
# **3.Feature Extraction**
In this step,features are extracted from the URLs dataset.
The extracted features are categorized into
1. Address Bar based Features
2. Domain based Features
3. HTML & Javascript Features
## **3.1.Address Bar Based Features:**
Many features can be extracted that can be consided as address bar base them, below mentioned were considered for this project.
* Domain of URL
* IP Address in URL
* "@"Symbol in URL
* Length of URL
* Depth of URL
* Redirection "//" in URL
* "http/https" in Domain name
* Using URL Shortening Services "TinyURL"
* Prefix or Suffix "-" in Domain
Each of these features are explained and the coded below:
```
#importing required packages for this section
from urllib.parse import urlparse,urlencode
import urllib
import urllib.request
import ipaddress
import re
from bs4 import BeautifulSoup
import whois
from datetime import datetime
import requests
```
# **3.1.1.Domain of the URL**
```
# 1.Domain of the URL(Domain)
def getDomain(url):
domain = urlparse(url).netloc
if re.match(r"^www.",domain):
domain = domain.replace("www.","")
return domain
```
# **3.1.2.IP Address in the URL**
```
# 2.Checks for IP address in URL (Have_IP)
def havingIP(url):
try:
ipaddress.ip_address(url)
ip = 1
except:
ip = 0
return ip
```
# **3.1.3. "@" Symbol in URL**
```
# 3.Checks the presence of @ in URL (Have_At)
def haveAtSign(url):
if "@" in url:
at = 1
else:
at = 0
return at
```
# **3.1.4. Length of URL**
```
# 4.Finding the length of URL and categorizing (URL_Length)
def getLength(url):
if len(url) < 54:
length = 0
else:
length = 1
return length
```
# **3.1.5. Depth of URL**
```
# 5.Gives number of '/' in URL (URL_Depth)
def getDepth(url):
s = urlparse(url).path.split('/')
depth = 0
for j in range(len(s)):
if len(s[j]) != 0:
depth = depth+1
return depth
```
# **3.1.6. Redirection "//" in URL**
```
# 6.Checking for redirection '//' in the url (Redirection)
def redirection(url):
pos = url.rfind('//')
if pos > 6:
if pos > 7:
return 1
else:
return 0
else:
return 0
```
# **3.1.7. "http/https" in Domain name**
```
# 7.Existence of “HTTPS” Token in the Domain Part of the URL (https_Domain)
def httpDomain(url):
domain = urlparse(url).netloc
if 'https' in domain:
return 1
else:
return 0
```
# **3.1.8. Using URL Shortening Services “TinyURL”**
```
#listing shortening services
shortening_services = r"bit\.ly|goo\.gl|shorte\.st|go2l\.ink|x\.co|ow\.ly|t\.co|tinyurl|tr\.im|is\.gd|cli\.gs|" \
r"yfrog\.com|migre\.me|ff\.im|tiny\.cc|url4\.eu|twit\.ac|su\.pr|twurl\.nl|snipurl\.com|" \
r"short\.to|BudURL\.com|ping\.fm|post\.ly|Just\.as|bkite\.com|snipr\.com|fic\.kr|loopt\.us|" \
r"doiop\.com|short\.ie|kl\.am|wp\.me|rubyurl\.com|om\.ly|to\.ly|bit\.do|t\.co|lnkd\.in|db\.tt|" \
r"qr\.ae|adf\.ly|goo\.gl|bitly\.com|cur\.lv|tinyurl\.com|ow\.ly|bit\.ly|ity\.im|q\.gs|is\.gd|" \
r"po\.st|bc\.vc|twitthis\.com|u\.to|j\.mp|buzurl\.com|cutt\.us|u\.bb|yourls\.org|x\.co|" \
r"prettylinkpro\.com|scrnch\.me|filoops\.info|vzturl\.com|qr\.net|1url\.com|tweez\.me|v\.gd|" \
r"tr\.im|link\.zip\.net"
# 8. Checking for Shortening Services in URL (Tiny_URL)
def tinyURL(url):
match=re.search(shortening_services,url)
if match:
return 1
else:
return 0
```
# **3.1.9. Prefix or Suffix "-" in Domain**
```
# 9.Checking for Prefix or Suffix Separated by (-) in the Domain (Prefix/Suffix)
def prefixSuffix(url):
if '-' in urlparse(url).netloc:
return 1 # phishing
else:
return 0 # legitimate
```
## **3.2. Domain Based Features:**
Many features can be extracted that come under this category. Out of them, below mentioned were considered for this project.
* DNS Record
* Website Traffic
*Age of Domain
*End Period of Domain
**3.2.1.DNS Record**
```
# 11.DNS Record availability (DNS_Record)
# obtained in the featureExtraction function itself
```
**3.2.2. Web Traffic**
```
# 12.Web traffic (Web_Traffic)
def web_traffic(url):
try:
#Filling the whitespaces in the URL if any
url = urllib.parse.quote(url)
rank = BeautifulSoup(urllib.request.urlopen("http://data.alexa.com/data?cli=10&dat=s&url=" + url).read(), "xml").find(
"REACH")['RANK']
rank = int(rank)
except TypeError:
return 1
if rank <100000:
return 1
else:
return 0
```
**3.2.3. Age of Domain**
```
# 13.Survival time of domain: The difference between termination time and creation time (Domain_Age)
def domainAge(domain_name):
creation_date = domain_name.creation_date
expiration_date = domain_name.expiration_date
if (isinstance(creation_date,str) or isinstance(expiration_date,str)):
try:
creation_date = datetime.strptime(creation_date,'%Y-%m-%d')
expiration_date = datetime.strptime(expiration_date,"%Y-%m-%d")
except:
return 1
if ((expiration_date is None) or (creation_date is None)):
return 1
elif ((type(expiration_date) is list) or (type(creation_date) is list)):
return 1
else:
ageofdomain = abs((expiration_date - creation_date).days)
if ((ageofdomain/30) < 6):
age = 1
else:
age = 0
return age
```
**3.2.4. End Period of Domain**
```
# 14.End time of domain: The difference between termination time and current time (Domain_End)
def domainEnd(domain_name):
expiration_date = domain_name.expiration_date
if isinstance(expiration_date,str):
try:
expiration_date = datetime.strptime(expiration_date,"%Y-%m-%d")
except:
return 1
if (expiration_date is None):
return 1
elif (type(expiration_date) is list):
return 1
else:
today = datetime.now()
end = abs((expiration_date - today).days)
if ((end/30) < 6):
end = 0
else:
end = 1
return end
```
# **3.3. HTML and JavaScript based Features**
Many features can be extracted that come under this category. Out of them, below mentioned were considered for this project.
* IFrame Redirection
*Status Bar Customization
*Disabling Right Click
*Website Forwarding
### **3.3.1. IFrame Redirection**
```
# 15. IFrame Redirection (iFrame)
def iframe(response):
if response == "":
return 1
else:
if re.findall(r"[<iframe>|<frameBorder>]", response.text):
return 0
else:
return 1
```
### **3.3.2. Status Bar Customization**
```
# 16.Checks the effect of mouse over on status bar (Mouse_Over)
def mouseOver(response):
if response == "" :
return 1
else:
if re.findall("<script>.+onmouseover.+</script>", response.text):
return 1
else:
return 0
```
### **3.3.3. Disabling Right Click**
```
# 17.Checks the status of the right click attribute (Right_Click)
def rightClick(response):
if response == "":
return 1
else:
if re.findall(r"event.button ?== ?2", response.text):
return 0
else:
return 1
```
### **3.3.4. Website Forwarding**
```
# 18.Checks the number of forwardings (Web_Forwards)
def forwarding(response):
if response == "":
return 1
else:
if len(response.history) <= 2:
return 0
else:
return 1
```
## **4. Computing URL Features**
```
#Function to extract features
def featureExtraction(url,label):
features = []
#Address bar based features (10)
features.append(getDomain(url))
features.append(havingIP(url))
features.append(haveAtSign(url))
features.append(getLength(url))
features.append(getDepth(url))
features.append(redirection(url))
features.append(httpDomain(url))
features.append(tinyURL(url))
features.append(prefixSuffix(url))
#Domain based features (4)
dns = 0
try:
domain_name = whois.whois(urlparse(url).netloc)
except:
dns = 1
features.append(dns)
features.append(web_traffic(url))
features.append(1 if dns == 1 else domainAge(domain_name))
features.append(1 if dns == 1 else domainEnd(domain_name))
# HTML & Javascript based features (4)
try:
response = requests.get(url)
except:
response = ""
features.append(iframe(response))
features.append(mouseOver(response))
features.append(rightClick(response))
features.append(forwarding(response))
features.append(label)
return features
```
### **4.2. Reviewing Phishing URLs:**
```
phishurl.shape
```
We will Reviewing URls 2500 by 2500 and storing seperate files and merge them.
```
feature_names = ['Domain', 'Have_IP', 'Have_At', 'URL_Length', 'URL_Depth','Redirection',
'https_Domain', 'TinyURL', 'Prefix/Suffix', 'DNS_Record', 'Web_Traffic',
'Domain_Age', 'Domain_End', 'iFrame', 'Mouse_Over','Right_Click', 'Web_Forwards', 'Label']
label = 1
```
0 - 5000 phishing URLs Feature extraction
```
#Extracting the feautres & storing them in a list
phish_features = []
for i in range(0, 5000):
url = phishurl['url'][i]
print(i)
phish_features.append(featureExtraction(url,label))
#converting the list to dataframe
phishing = pd.DataFrame(phish_features, columns= feature_names)
phishing.head()
# Storing the extracted legitimate URLs fatures to csv file
phishing.to_csv('5-phish_features.csv', index= False)
```
### Contributors - Deshitha Hansajith [Linked In](https://www.linkedin.com/in/deshitha-hansajith/), Nipuni Dilukshika [Linked In](https://www.linkedin.com/in/nipuni-dilukshika-186764197/)
| github_jupyter |
```
# Erasmus+ ICCT project (2018-1-SI01-KA203-047081)
# Toggle cell visibility
from IPython.display import HTML
tag = HTML('''<script>
code_show=true;
function code_toggle() {
if (code_show){
$('div.input').hide()
} else {
$('div.input').show()
}
code_show = !code_show
}
$( document ).ready(code_toggle);
</script>
Toggle cell visibility <a href="javascript:code_toggle()">here</a>.''')
display(tag)
# Hide the code completely
# from IPython.display import HTML
# tag = HTML('''<style>
# div.input {
# display:none;
# }
# </style>''')
# display(tag)
```
## Dinamika hitrosti vozila
Ta interaktivni primer prikazuje popis dinamike hitrosti vozila z uporabe diferencialnih enačb ter pretvorbo le-teh v prostor stanj.
Dinamika hitrosti vozila je pod vplivom dveh glavnih sil: zdrsa in zračnega upora. Podatki za konkreten primer tega sistema so: masa vozila $m = 1000$ kg, maximalni navor motorja $\tau_{\text{max}}=150$ Nm, premer koles $r=25$ cm (poenostavimo, da je prestavno razmerje $\eta$ enako 4:1), koeficient zračnega upora $b = 60$ Ns/m (zračni upor je modeliran kot linearna funkcija hitrosti vozila $v$).
Predpostavimo, da lahko dinamiko motorja, od trenutka pritiska na stopalko za plin do dosega želenega deleža maksimalnega navora $\tau_{\%}$, modeliramo na naslednji način:
$$
\dot{\tau}_{\%}=-\frac{1}{2}\tau_{\%}+\frac{1}{2}t_{r}
$$
Iz zgornjega modela vidimo, da gre za linearni sistem prvega reda s časovno konstanto $T=2$ s in enotskim ojačanjem.
### Diferencialna enačba sistema
Na podlagi drugega Newtonovega zakona ($F=ma$) lahko zapišemo diferencialno enačbo, ki popisuje gibanje vozila:
$$
m\dot{v}= \frac{\tau_{\text{max}}\eta}{r}\tau_{\%}-bv,
$$
kjer dejanski navor, ki ga generira motor dobimo z zmnožkom $\tau_{\%}$ in maksimalnega navora $\tau_{\text{max}}$. Odziv sistema lahko nato opišemo z naslednjim sistemom diferencialnih enačb:
$$
\begin{cases}
m\dot{v}= \frac{\tau_{\text{max}}\eta}{r}\tau_{\%}-bv \\
\dot{\tau}_{\%}=-\frac{1}{2}\tau_{\%}+\frac{1}{2}t_{r}
\end{cases}
$$
### Sistem v prostoru stanj
Zgornji diferencialni enačbi sta prvega reda, tako da je število potrebnih stanj, s katerim lahko popišemo delovanje sistema enako 2. Če definiramo vektor stanj
$x=\begin{bmatrix}x_1&x_2\end{bmatrix}^T=\begin{bmatrix}v&\tau_{\%}\end{bmatrix}^T$ in obravnavamo $t_r$ kot vhod v sistem, lahko zapišemo sistem diferencialnih enačb v prostoru stanj kot:
$$
\dot{x}=\underbrace{\begin{bmatrix}-\frac{b}{m}&\frac{\tau_{\text{max}}\eta}{mr}\\0&-\frac{1}{2}\end{bmatrix}}_{A}x+\underbrace{\begin{bmatrix}0\\\frac{1}{2}\end{bmatrix}}_{B}t_r \\
$$
### Kako upravljati s tem interaktivnim primerom?
Spreminjaj parametre sistema in poizkusi odgovoriti na naslednja vprašanja:
- Ali lahko obravnavani poenostavljen model vozila, ki je predstavljen v tem interaktivnem primeru, doseže neskončno hitrost? Zakaj?
- Ali je maksimalna hitrost vozila odvisna od mase vozia? Zakaj?
- Ali je negativna vrednost $t_r$ relevantna v obranavanem modelu? Zakaj?
```
#Preparatory Cell
%matplotlib notebook
import control
import numpy
from IPython.display import display, Markdown
import ipywidgets as widgets
import matplotlib.pyplot as plt
%matplotlib inline
#print a matrix latex-like
def bmatrix(a):
"""Returns a LaTeX bmatrix - by Damir Arbula (ICCT project)
:a: numpy array
:returns: LaTeX bmatrix as a string
"""
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{bmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{bmatrix}']
return '\n'.join(rv)
# Display formatted matrix:
def vmatrix(a):
if len(a.shape) > 2:
raise ValueError('bmatrix can at most display two dimensions')
lines = str(a).replace('[', '').replace(']', '').splitlines()
rv = [r'\begin{vmatrix}']
rv += [' ' + ' & '.join(l.split()) + r'\\' for l in lines]
rv += [r'\end{vmatrix}']
return '\n'.join(rv)
#matrixWidget is a matrix looking widget built with a VBox of HBox(es) that returns a numPy array as value !
class matrixWidget(widgets.VBox):
def updateM(self,change):
for irow in range(0,self.n):
for icol in range(0,self.m):
self.M_[irow,icol] = self.children[irow].children[icol].value
#print(self.M_[irow,icol])
self.value = self.M_
def dummychangecallback(self,change):
pass
def __init__(self,n,m):
self.n = n
self.m = m
self.M_ = numpy.matrix(numpy.zeros((self.n,self.m)))
self.value = self.M_
widgets.VBox.__init__(self,
children = [
widgets.HBox(children =
[widgets.FloatText(value=0.0, layout=widgets.Layout(width='90px')) for i in range(m)]
)
for j in range(n)
])
#fill in widgets and tell interact to call updateM each time a children changes value
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
self.children[irow].children[icol].observe(self.updateM, names='value')
#value = Unicode('example@example.com', help="The email value.").tag(sync=True)
self.observe(self.updateM, names='value', type= 'All')
def setM(self, newM):
#disable callbacks, change values, and reenable
self.unobserve(self.updateM, names='value', type= 'All')
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].unobserve(self.updateM, names='value')
self.M_ = newM
self.value = self.M_
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].value = self.M_[irow,icol]
for irow in range(0,self.n):
for icol in range(0,self.m):
self.children[irow].children[icol].observe(self.updateM, names='value')
self.observe(self.updateM, names='value', type= 'All')
#self.children[irow].children[icol].observe(self.updateM, names='value')
#overlaod class for state space systems that DO NOT remove "useless" states (what "professor" of automatic control would do this?)
class sss(control.StateSpace):
def __init__(self,*args):
#call base class init constructor
control.StateSpace.__init__(self,*args)
#disable function below in base class
def _remove_useless_states(self):
pass
#define matrixes
C = numpy.matrix([[1,0],[0,1]])
D = numpy.matrix([[0],[0]])
X0 = matrixWidget(2,1)
m = widgets.FloatSlider(
value=1000,
min=400,
max=2000,
step=1,
description='m [kg]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
eta = widgets.FloatSlider(
value=4,
min=0.8,
max=10.0,
step=0.1,
description='$\eta$:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
tau_max = widgets.FloatSlider(
value=150,
min=50,
max=900,
step=1,
description=r'$\tau_{\text{max}}$ [Nm]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
b_air = widgets.FloatSlider(
value=60,
min=0,
max=200,
step=1,
description=r'$b$ [Ns/m]:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
u = widgets.FloatSlider(
value=0.5,
min=0,
max=1,
step=0.01,
description='',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
omega = widgets.FloatSlider(
value=5,
min=0,
max=10.0,
step=0.1,
description='',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='.1f',
)
#create dummy widget
DW = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
#create button widget
START = widgets.Button(
description='Pritisni!',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Pritisni za spremembo začetnih pogojev',
icon='check'
)
def on_start_button_clicked(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW.value> 0 :
DW.value = -1
else:
DW.value = 1
pass
START.on_click(on_start_button_clicked)
#define type of ipout
SELECT = widgets.Dropdown(
options=['impulzna funkcija', 'koračna funkcija', 'sinusoidna funkcija'],
value='koračna funkcija',
description='',
disabled=False
)
def main_callback(X0, m, eta, tau_max, b_air, u, selu, omega, DW):
r = 0.25 # m
a = numpy.matrix([[-b_air/m,tau_max*eta/m/r],[0,-1/2]])
b = numpy.matrix([[0],[1/2]])
eig = numpy.linalg.eig(a)
sys = sss(a,b,C,D)
if min(numpy.real(abs(eig[0]))) != 0:
T = numpy.linspace(0,10/min(numpy.real(abs(eig[0]))),1000)
else:
if max(numpy.real(abs(eig[0]))) != 0:
T = numpy.linspace(0,10/max(numpy.real(abs(eig[0]))),1000)
else:
T = numpy.linspace(0,180,1000)
if selu == 'impulzna funkcija': #selu
U = [0 for t in range(0,len(T))]
U[0] = u
y = control.forced_response(sys,T,U,X0)
if selu == 'koračna funkcija':
U = [u for t in range(0,len(T))]
y = control.forced_response(sys,T,U,X0)
if selu == 'sinusoidna funkcija':
U = u*numpy.sin(omega*T)
y = control.forced_response(sys,T,U,X0)
fig=plt.figure(num=1,figsize=[15, 4])
fig.add_subplot(121)
plt.plot(T,y[1][0])
plt.grid()
plt.xlabel('čas [s]')
plt.ylabel('hitrost [m/s]')
fig.add_subplot(122)
plt.plot(T,y[1][1])
plt.grid()
plt.xlabel('čas [s]')
plt.ylabel(r'$\tau_\%$')
#display(Markdown('The A matrix is: $%s$ and the eigenvalues are: $%s$' % (bmatrix(a),eig[0])))
#create a graphic structure to hold all widgets
alltogether = widgets.VBox([widgets.HBox([widgets.VBox([m,
eta,
tau_max,
b_air]),
widgets.HBox([widgets.VBox([widgets.Label('Izberi vhodno funkcijo:',border=3),
widgets.Label('$t_r$:',border=3),
widgets.Label('omega [rad/s]:',border=3)]),
widgets.VBox([SELECT,u,omega])])]),
widgets.HBox([widgets.Label('Začetni pogoji X0:',border=3),X0,
widgets.Label('Pritisni za spremembo začetnih pogojev:',border=3),START])])
out = widgets.interactive_output(main_callback,{'X0':X0, 'm': m, 'eta': eta, 'tau_max': tau_max, 'b_air': b_air, 'u': u, 'selu': SELECT, 'omega':omega, 'DW':DW})
#out.layout.height = '200px'
display(out,alltogether)
#create dummy widget 2
DW2 = widgets.FloatText(layout=widgets.Layout(width='0px', height='0px'))
DW2.value = -1
#create button widget
START2 = widgets.Button(
description='Pokaži pravilne odgovore',
disabled=False,
button_style='', # 'success', 'info', 'warning', 'danger' or ''
tooltip='Pritisni za preverbo odgovorov',
icon='check',
layout=widgets.Layout(width='200px', height='auto')
)
def on_start_button_clicked2(b):
#This is a workaround to have intreactive_output call the callback:
# force the value of the dummy widget to change
if DW2.value> 0 :
DW2.value = -1
else:
DW2.value = 1
pass
START2.on_click(on_start_button_clicked2)
def main_callback2(DW2):
if DW2 > 0:
display(Markdown(r'''>Odgovori:
>- Da, a le če je koeficient zračnega upora enak 0.
>- Ne, masa avtomobila vpliva le na čas, potreben, da se doseže določeno hitrost; vpliva torej na pospešek in ne na maksimalno hitrost.
>- Da, lahko se uporabi za modeliranje procesa zaviranja vozila.'''))
else:
display(Markdown(''))
#create a graphic structure to hold all widgets
alltogether2 = widgets.VBox([START2])
out2 = widgets.interactive_output(main_callback2,{'DW2':DW2})
#out.layout.height = '300px'
display(out2,alltogether2)
```
| github_jupyter |
```
# default_exp model
#export
from fastai.vision.all import *
import torchaudio
import torchaudio.transforms
#hide
from nbdev.showdoc import *
```
# fastai_resnet_audio model
> ResNet-like 1D CNN model for audio
## Dependencies
- fastai2
- torchaudio
## ResNet-like 1D CNN
This code is inspired by https://www.kaggle.com/readilen/resnet-for-mnist-with-pytorch, https://towardsdatascience.com/understanding-and-visualizing-resnets-442284831be8 and
https://github.com/fastai/fastai2/blob/master/nbs/11_vision.models.xresnet.ipynb
```
#export
def conv1xk(in_channels, out_channels, kernel_size=3, stride=1):
padding = kernel_size // 2
return nn.Conv1d(in_channels, out_channels, kernel_size=kernel_size,
stride=stride, padding=padding, bias=False)
def init_cnn_1d(m):
if getattr(m, 'bias', None) is not None: nn.init.constant_(m.bias, 0)
if isinstance(m, (nn.Conv1d,nn.Linear)): nn.init.kaiming_normal_(m.weight)
for l in m.children(): init_cnn_1d(l)
def splitter(m):
return L(m[0][:6], m[0][6:], m[1]).map(params)
# Residual block
class ResidualBlock(nn.Module):
def __init__(self, in_channels, out_channels, kernel_size=3, stride=1, downsample=None):
super(ResidualBlock, self).__init__()
self.conv1 = conv1xk(in_channels, out_channels, kernel_size, stride)
self.bn1 = nn.BatchNorm1d(out_channels)
self.relu = nn.ReLU(inplace=True)
self.conv2 = conv1xk(out_channels, out_channels, kernel_size)
self.bn2 = nn.BatchNorm1d(out_channels)
self.downsample = downsample
def forward(self, x):
residual = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
if self.downsample:
residual = self.downsample(residual)
out += residual
out = self.relu(out)
return out
class ResNetAudio(nn.Sequential):
def __init__(self, block, layers, in_channels=64, num_classes=10, kernel_size=3, stride=2, dropout=0.2):
in_block = [] # input block
residual_block = [] # residual blocks
header_block = [] # linear head
self.in_channels = in_channels
self.block = block
in_kernel_size = kernel_size * 2 + 1
in_block.append(conv1xk(1, in_channels, in_kernel_size, stride))
in_block.append(nn.BatchNorm1d(in_channels))
in_block.append(nn.ReLU(inplace=True))
in_block.append(nn.MaxPool1d(kernel_size, stride, kernel_size//3))
residual_block = self.make_blocks(layers, in_channels, kernel_size, stride)
header_block.append(nn.AdaptiveAvgPool1d(1))
header_block.append(nn.Flatten())
header_block.append(nn.Dropout(dropout))
header_block.append(nn.Linear(in_channels*2**(len(layers)-1), num_classes))
super().__init__(nn.Sequential(*in_block, *residual_block),nn.Sequential(*header_block))
init_cnn_1d(self)
def make_blocks(self, layers, in_channels, kernel_size, stride):
return [self.make_layer(self.block, in_channels*2**i, l, kernel_size, stride) for i, l in enumerate(layers)]
def make_layer(self, block, out_channels, blocks, kernel_size=3, stride=1):
downsample = None
if (stride != 1) or (self.in_channels != out_channels):
downsample = nn.Sequential(
conv1xk (self.in_channels, out_channels, kernel_size=kernel_size, stride=stride),
nn.BatchNorm1d(out_channels))
layers = []
layers.append(block(self.in_channels, out_channels, kernel_size, stride, downsample))
self.in_channels = out_channels
for i in range(1, blocks):
layers.append(block(out_channels, out_channels, kernel_size))
return nn.Sequential(*layers)
```
## Fine-Tunable model
Call replace_head(learn.model, num_classes=<new_num_classes>) to adapt the model head to a new dataset (new number of classes). Now you can use learn.fine_tune(), learn.unfreeze() and learn.fit_one_cycle() to fine-tune the model.
```
#export
def replace_head(model, num_classes):
model[-1][-1] = nn.Linear(512, num_classes)
apply_init(model[1], nn.init.kaiming_normal_)
```
## Configurations
Configurations for **ResNet18 / ResNet34-like** architectures. Kernel size 34 and stride 4 seem to work quite
well. But there is still room for experimentation and improvement.
```
#export
# resnet 18
resnet1d18 = {
"block": ResidualBlock,
"layers": [2, 2, 2, 2],
"in_channels": 64,
"kernel_size": 15,
"stride": 4,
"num_classes": 10
}
# resnet 34
resnet1d34 = {
"block": ResidualBlock,
"layers": [3, 4, 6, 3],
"in_channels": 64,
"kernel_size": 15,
"stride": 4,
"num_classes": 10
}
```
## Tests
Test architecutre
```
#hide
bs = 8
arch = resnet1d18
model = ResNetAudio(**arch)
inp = torch.randn(bs,1,22050)
out = model(inp)
assert len(out) == bs
assert len(out[0]) == arch['num_classes']
```
Test replace_head
```
#hide
num_classes = 22
replace_head(model, num_classes)
assert getattr(model[-1][-1], 'out_features') == num_classes
```
| github_jupyter |
# Lale: Auto-ML and Types for Scikit-learn
This notebook is an introductory guide to
[Lale](https://github.com/ibm/lale) for scikit-learn users.
[Scikit-learn](https://scikit-learn.org) is a popular, easy-to-use,
and comprehensive data science library for Python. This notebook aims
to show how Lale can make scikit-learn even better in two areas:
auto-ML and type checking. First, if you do not want to manually
select all algorithms or tune all hyperparameters, you can leave it to
Lale to do that for you automatically. Second, when you pass
hyperparameters or datasets to scikit-learn, Lale checks that these
are type-correct. For both auto-ML and type-checking, Lale uses a
single source of truth: machine-readable schemas associated with
scikit-learn compatible transformers and estimators. Rather than
invent a new schema specification language, Lale uses [JSON
Schema](https://json-schema.org/understanding-json-schema/), because
it is popular, widely-supported, and makes it easy to store or send
hyperparameters as JSON objects. Furthermore, by using the same
schemas both for auto-ML and for type-checking, Lale ensures that
auto-ML is consistent with type checking while also reducing the
maintenance burden to a single set of schemas.
Lale is an open-source Python library and you can install it by doing
`pip install lale`. See
[installation](https://github.com/IBM/lale/blob/master/docs/installation.rst)
for further instructions. Lale uses the term *operator* to refer to
what scikit-learn calls machine-learning transformer or estimator.
Lale provides schemas for 144
[operators](https://github.com/IBM/lale/tree/master/lale/lib). Most of
these operators come from scikit-learn itself, but there are also
operators from other frameworks such as XGBoost or PyTorch.
If Lale does not yet support your favorite operator, you can add it
yourself by following this
[guide](https://nbviewer.jupyter.org/github/IBM/lale/blob/master/examples/docs_new_operators.ipynb).
If you do add a new operator, please consider contributing it back to
Lale!
The rest of this notebook first demonstrates auto-ML, then reveals
some of the schemas that make that possible, and finally demonstrates
how to also use the very same schemas for type checking.
## 1. Auto-ML with Lale
Lale serves as an interface for two Auto-ML tasks: hyperparameter tuning
and algorithm selection. Rather than provide new implementations for
these tasks, Lale reuses existing implementations. The next few cells
demonstrate how to use Hyperopt and GridSearchCV from Lale. Lale also
supports additional optimizers, not shown in this notebook. In all
cases, the syntax for specifying the search space is the same.
### 1.1 Hyperparameter Tuning with Lale and Hyperopt
Let's start by looking at hyperparameter tuning, which is an important
subtask of auto-ML. To demonstrate it, we first need a dataset.
Therefore, we load the California Housing dataset and display the
first few rows to get a feeling for the data. Lale can process both
Pandas dataframes and Numpy ndarrays; here we use dataframes.
```
import pandas as pd
import lale.datasets
(train_X, train_y), (test_X, test_y) = lale.datasets.california_housing_df()
pd.concat([train_X.head(), train_y.head()], axis=1)
```
As you can see, the target column is a continuous number, indicating
that this is a regression task. Besides the target, there are eight
feature columns, which are also all continuous numbers. That means
many scikit-learn operators will work out of the box on this data
without needing to preprocess it first. Next, we need to import a few
operators. `PCA` (principal component analysis) is a transformer from
scikit-learn for linear dimensionality reduction.
`DecisionTreeRegressor` is an estimator from scikit-learn that can
predict the target column. `Hyperopt` is a Lale wrapper for
the [hyperopt](http://hyperopt.github.io/hyperopt/) auto-ML library.
And finally, `wrap_imported_operators` augments both `PCA` and `Tree`
with schemas to enable Lale to tune their hyperparameters.
```
from sklearn.decomposition import PCA
from sklearn.tree import DecisionTreeRegressor as Tree
from lale.lib.lale import Hyperopt
lale.wrap_imported_operators()
```
Next, we create a two-step pipeline of `PCA` and `Tree`. Similar to scikit-learn, Lale supports creation of a pipeline using `Pipeline(...)` as well as `make_pipeline`. The only difference compared to a scikit-learn pipeline
is that since we want Lale to tune the hyperparameters for us, we do
not specify them by hand. Specifically, we just write `PCA` instead of
`PCA(...)`, omitting the hyperparameters for `PCA`. Analogously, we
just write `Tree` instead of `Tree(...)`, omitting the hyperparameters
for `Tree`. Rather than binding hyperparameters by hand, we leave them
free to be tuned by hyperopt.
```
pca_tree_planned = lale.operators.Pipeline([('pca', PCA), ('tree', Tree)])
```
We use `auto_configure` on the pipeline and pass `Hyperopt` as an optimizer. This will use the pipeline's search space to find the best pipeline. In this case, the search uses 10 trials. Each
trial draws values for the hyperparameters from the ranges specified
by the JSON schemas associated with the operators in the pipeline.
```
%%time
pca_tree_trained = pca_tree_planned.auto_configure(
train_X, train_y, optimizer=Hyperopt, cv=3, max_evals=10, scoring='r2')
```
By default, Hyperopt uses k-fold cross validation
to evaluate each trial. The end result is the pipeline that
performed best out of all trials. In addition to the cross-val score,
we can also evaluate this best pipeline against the test data. We
simply use the existing R2 score metric from scikit-learn for this
purpose.
```
import sklearn.metrics
predicted = pca_tree_trained.predict(test_X)
print(f'R2 score {sklearn.metrics.r2_score(test_y, predicted):.2f}')
```
### 1.2 Inspecting the Results of Automation
In the previous example, the automation picked hyperparameter values
for PCA and the decision tree. We know the values were valid and we
know how well the pipeline performed with them. But we might also want
to know exactly which values were picked. One way to do that is by
visualizing the pipeline and using tooltips. If you are looking at
this notebook in a viewer that supports tooltips, you can hover the
mouse pointer over either one of the operators to see its
hyperparameters.
```
pca_tree_trained.visualize()
```
Another way to view the results of hyperparameter tuning in Lale is by
pretty-printing the pipeline as Python source code. Calling the
`pretty_print` method with `ipython_display=True` prints the code with
syntax highlighting in a Jupyter notebook. The pretty-printed code
contains the hyperparameters. It also uses the `>>` symbol, which is
just syntactic sugar for calling the `make_pipeline` function.
```
pca_tree_trained.pretty_print(ipython_display=True, show_imports=False)
```
### 1.3 Hyperparameter Tuning with Lale and GridSearchCV
Lale supports multiple auto-ML tools, not just hyperopt. For instance,
you can also use
[GridSearchCV](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html)
from scikit-learn. You could use the exact same `pca_tree_planned`
pipeline for this as we did with the hyperopt tool.
However, to avoid running for a long time, here we simplify the space:
for `PCA`, we bind the `svd_solver` so only the remaining hyperparameters
are being searched, and for `Tree`, we call `freeze_trainable()` to bind
all hyperparameters to their defaults. Lale again uses the schemas
attached to the operators in the pipeline to generate a suitable search grid.
Note that, to be compatible with scikit-learn, `lale.lib.lale.GridSearchCV`
can also take a `param_grid` as an argument if the user chooses to use a
handcrafted grid instead of the one generated automatically.
```
%%time
from lale.lib.lale import GridSearchCV
from lale.operators import make_pipeline
grid_search_planned = lale.operators.Pipeline([
('pca', PCA(svd_solver='auto')), ('tree', Tree().freeze_trainable())])
grid_search_result = grid_search_planned.auto_configure(
train_X, train_y, optimizer=GridSearchCV, cv=3, scoring='r2')
```
Just like we saw earlier with hyperopt, you can use the best pipeline
found for scoring and evaluate the quality of the predictions.
```
predicted = grid_search_result.predict(test_X)
print(f'R2 score {sklearn.metrics.r2_score(test_y, predicted):.2f}')
```
Similarly, to inspect the results of grid search, you have the same
options as demonstrated earlier for hypopt. For instance, you can
pretty-print the best pipeline found by grid search back as Python
source code, and then look at its hyperparameters.
```
grid_search_result.pretty_print(ipython_display=True, show_imports=False)
```
### 1.4 Pipeline Combinators
We already saw that `>>` is syntactic sugar for `make_pipeline`. Lale
refers to `>>` as the *pipe combinator*. Besides `>>`, Lale supports
two additional combinators. Before we introduce them, let's import a
few more things.
```
from lale.lib.lale import NoOp, ConcatFeatures
from sklearn.linear_model import LinearRegression as LinReg
from xgboost import XGBRegressor as XGBoost
lale.wrap_imported_operators()
```
Lale emulates the scikit-learn APIs for composing pipelines using
functions. We already saw `make_pipeline`. Another function in
scikit-learn is `make_union`, which composes multiple sub-pipelines to
run on the same data, then concatenates the features. In other words,
`make_union` produces a horizontal stack of the data transformed by
its sub-pipelines. To support auto-ML, Lale introduces a third
function, `make_choice`, which does not exist in scikit-learn. The
`make_choice` function specifies an algorithmic choice for auto-ML to
resolve. In other words, `make_choice` creates a search space for
automated algorithm selection.
```
dag_with_functions = lale.operators.make_pipeline(
lale.operators.make_union(PCA, NoOp),
lale.operators.make_choice(Tree, LinReg, XGBoost(booster='gbtree')))
dag_with_functions.visualize()
```
The visualization shows `make_union` as multiple sub-pipelines feeding
into `ConcatFeatures`, and it shows `make_choice` using an `|`
combinator. Operators shown in white are already fully trained; in
this case, these operators actually do not have any learnable
coefficients, nor do they have hyperparameters. For each of the three
functions `make_pipeline`, `make_choice`, and `make_union`, Lale also
provides a corresponding combinator. We already saw the pipe
combinator (`>>`) and the choice combinator (`|`). To get the effect
of `make_union`, use the *and combinator* (`&`) with the
`ConcatFeatures` operator. The next example shows the exact same
pipeline as before, but written using combinators instead of
functions.
```
dag_with_combinators = (
(PCA(svd_solver='full') & NoOp)
>> ConcatFeatures
>> (Tree | LinReg | XGBoost(booster='gbtree')))
dag_with_combinators.visualize()
```
### 1.5 Combined Algorithm Selection and Hyperparameter Optimization
Since the `dag_with_functions` specifies an algorithm choice, when we
feed it to a `Hyperopt`, hyperopt will do algorithm selection
for us. And since some of the operators in the dag do not have all
their hyperparameters bound, hyperopt will also tune their free
hyperparameters for us. Note that `booster` for `XGBoost` is fixed to `gbtree` and hence Hyperopt would not tune it.
```
%%time
multi_alg_trained = dag_with_functions.auto_configure(
train_X, train_y, optimizer=Hyperopt, cv=3, max_evals=10, scoring='r2')
```
Visualizing the best estimator reveals what algorithms
hyperopt chose.
```
multi_alg_trained.visualize()
```
Pretty-printing the best estiamtor reveals how hyperopt tuned the
hyperparameters. For instance, we can see that a `randomized` `svd_solver` was chosen for PCA.
```
multi_alg_trained.pretty_print(ipython_display=True, show_imports=False)
```
Of course, the trained pipeline can be used for predictions as usual,
and we can use scikit-learn metrics to evaluate those predictions.
```
predicted = multi_alg_trained.predict(test_X)
print(f'R2 score {sklearn.metrics.r2_score(test_y, predicted):.2f}')
```
## 2. Viewing and Customizing Schemas
This section reveals more of what happens behind the scenes for
auto-ML with Lale. In particular, it shows the JSON Schemas used for
auto-ML, and demonstrates how to customize them if desired.
### 2.1 Looking at Schemas from a Notebook
When writing data science code, I often don't remember all the API
information about what hyperparameters and datasets an operator
expects. Lale attaches this information to the operators and uses it
for auto-ML as demonstrated above. The same information can also be
useful as interactive documentation in a notebook. Most individual
operators in the visualizations shown earlier in this notebook actually
contain a hyperlink to the excellent online documentation of
scikit-learn. We can also retrieve that hyperlink using a method call.
```
print(Tree.documentation_url())
```
Lale's helper function `ipython_display` pretty-prints JSON documents
and JSON schemas in a Jupyter notebook. You can get a quick overview
of the constructor arguments of an operator by calling the
`hyperparam_defaults` method.
```
from lale.pretty_print import ipython_display
ipython_display(Tree.hyperparam_defaults())
```
Hyperparameters can be categorical (meaning they accept a few
discrete values) or continuous (integers or real numbers).
As an example for a categorical hyperparameter, let's look at the
`criterion`. JSON Schema can encode categoricals as an `enum`.
```
ipython_display(Tree.hyperparam_schema('criterion'))
```
As an example for a continuous hyperparameter, let's look at
`max_depth`. The decision tree regressor in scikit-learn accepts
either an integer for that, or `None`, which has its own meaning.
JSON Schema can express these two choices as an `anyOf`, and
encodes the Python `None` as a JSON `null`. Also, while
any positive integer is a valid value, in the context of auto-ML,
Lale specifies a bounded range for the optimizer to search over.
```
ipython_display(Tree.hyperparam_schema('max_depth'))
```
Besides hyperparameter schemas, Lale also provides dataset schemas.
For exampe, NMF, which stands for non-negative matrix factorization,
requires a non-negative matrix as `X`. In JSON Schema, we express this
as an array of arrays of numbers with `minimum: 0`. While NMF also
accepts a second argument `y`, it does not use that argument.
Therefore, Lale gives `y` the empty schema `{}`, which permits any
values.
```
from sklearn.decomposition import NMF
lale.wrap_imported_operators()
ipython_display(NMF.input_schema_fit())
```
### 2.2 Customizing Schemas from a Notebook
While you can use Lale schemas as-is, you can also customize the
schemas to exert more control over the automation. As one example, it is common to tune XGBoost to use a large number for `n_estimators`. However, you might want to
reduce the number of trees in an XGBoost forest to reduce memory
consumption or to improve explainability. As another example, you
might want to hand-pick one of the boosters to reduce the search space
and thus hopefully speed up the search.
```
import lale.schemas as schemas
Grove = XGBoost.customize_schema(
n_estimators=schemas.Int(min=2, max=6),
booster=schemas.Enum(['gbtree']))
```
As this example demonstrates, Lale provides a simple Python API for
writing schemas, which it then converts to JSON Schema internally. The
result of customization is a new copy of the operator that can be used
in the same way as any other operator in Lale. In particular, it can
be part of a pipeline as before.
```
grove_planned = lale.operators.make_pipeline(
lale.operators.make_union(PCA, NoOp),
Grove)
grove_planned.visualize()
```
Given this new planned pipeline, we use hyperopt as before to search
for a good trained pipeline.
```
%%time
grove_trained = grove_planned.auto_configure(
train_X, train_y, optimizer=Hyperopt, cv=3, max_evals=10, scoring='r2')
```
As with all trained Lale pipelines, we can evaluate `grove_trained`
with metrics to see how well it does. Also, we can pretty-print
it back as Python code to double-check whether hyperopt obeyed the
customized schemas for `n_estimators` and `booster`.
```
predicted = grove_trained.predict(test_X)
print(f'R2 score {sklearn.metrics.r2_score(test_y, predicted):.2f}')
grove_trained.pretty_print(ipython_display=True, show_imports=False)
```
## 3. Type-Checking with Lale
The rest of this notebook gives examples for how the same schemas
that serve for auto-ML can also serve for error checking. We will
give comparative examples for error checking in scikit-learn (without
schemas) and in Lale (with schemas). To make it clear which version
of an operator is being used, all of the following examples uses
fully-qualified names (e.g., `sklearn.feature_selection.RFE`). The
fully-qualified names are for presentation purposes only; in typical
usage of either scikit-learn or Lale, these would be simple names
(e.g. just `RFE`).
### 3.1 Hyperparameter Error Example in Scikit-Learn
First, we import a few things.
```
import sys
import sklearn
from sklearn import pipeline, feature_selection, ensemble, tree
```
We use `make_pipeline` to compose a pipeline of two steps: an RFE
transformer and a decision tree regressor. RFE performs recursive
feature elimination, keeping only those features of the input data
that are the most useful for its `estimator` argument. For RFE's
estimator argument, the following code uses a random forest with 10
trees.
```
sklearn_hyperparam_error = sklearn.pipeline.make_pipeline(
sklearn.feature_selection.RFE(
estimator=sklearn.ensemble.RandomForestRegressor(n_estimators=10)),
sklearn.tree.DecisionTreeRegressor(max_depth=-1))
```
The `max_depth` argument for a decision tree cannot be a
negative number. Hence, the above code actually contains a bug: it
sets `max_depth=-1`. Scikit-learn does not check for this mistake from
the `__init__` method, otherwise we would have seen an error message
already. Instead, scikit-learn checks for this mistake during `fit`.
Unfortunately, it takes a few seconds to get the exception, because
scikit-learn first trains the RFE transformer and uses it to transform
the data. Only then does it pass the data to the decision tree.
```
%%time
try:
sklearn_hyperparam_error.fit(train_X, train_y)
except ValueError as e:
message = str(e)
print(message, file=sys.stderr)
```
Fortunately, this error message is pretty clear. Scikit-learn
implements the error check imperatively, using Python if-statements
to raise an exception when hyperparameters are configured wrong.
This notebook is part of Lale's regression test suite and gets run
automatically when changes are pushed to the Lale source code
repository. The assertion in the following cell is a test that the
error-check indeed behaves as expected and documented here.
```
assert message.startswith("max_depth must be greater than zero.")
```
### 3.2 Checking Hyperparameters with Types
Lale performs the same error checks, but using JSON Schema validation
instead of Python if-statements and raise-statements. First, we import
the `jsonschema` validator so we can catch its exceptions.
```
import jsonschema
```
Below is the exact same pipeline as before, but written in Lale
instead of directly in scikit-learn. In both cases, the underlying
implementation is in scikit-learn; Lale only adds thin wrappers to
support type checking and auto-ML.
```
%%time
try:
lale_hyperparam_error = lale.operators.make_pipeline(
lale.lib.sklearn.RFE(
estimator=lale.lib.sklearn.RandomForestRegressor(n_estimators=10)),
lale.lib.sklearn.DecisionTreeRegressor(max_depth=-1))
except jsonschema.ValidationError as e:
message = e.message
print(message, file=sys.stderr)
assert message.startswith("Invalid configuration for DecisionTreeRegressor(max_depth=-1)")
```
Just like in the scikit-learn example, the error message in the Lale
example also pin-points the problem as passing `max_depth=-1` to the
decision tree. It does so in a more stylized way, printing the
relevant JSON schema for this hyperparameter. Lale detects the error
already when the wrong hyperparameter is being passed as an argument,
thus reducing the amount of code you have to look at to find the root
cause. Furthermore, Lale takes only tens of milliseconds to detect
the error, because it does not attempt to train the RFE transformer
first. In this example, that only saves a few seconds, which may not
be significant. But there are situations with larger time savings,
such as when using larger datasets, slower operators, or when auto-ML
tries out many pipelines.
### 3.3 Dataset Error Example in Scikit-Learn
Above, we saw an example for detecting a hyperparameter error in
scikit-learn and in Lale. Next, we look at an analogous example for a
dataset error. Again, let's first look at the experience with
scikit-learn and then the same thing with Lale.
```
from sklearn import decomposition
```
We use scikit-learn to compose a pipeline of two steps: an RFE
transformer as before, this time followed by an NMF transformer.
```
sklearn_dataset_error = sklearn.pipeline.make_pipeline(
sklearn.feature_selection.RFE(
estimator=sklearn.ensemble.RandomForestRegressor(n_estimators=10)),
sklearn.decomposition.NMF())
```
NMF, or non-negative matrix factorization, does not allow any negative
numbers in its input matrix. The California Housing dataset contains
some negative numbers and the RFE does not eliminate those features.
To detect the mistake, scikit-learn must first train the RFE and
transform the data with it, which takes a few seconds. Then, NMF
detects the error and throws an exception.
```
%%time
try:
sklearn_dataset_error.fit(train_X, train_y)
except ValueError as e:
message = str(e)
print(message, file=sys.stderr)
assert message.startswith("Negative values in data passed to NMF (input X)")
```
### 3.4 Types for Dataset Checking
Lale uses types (as expressed using JSON schemas) to check
dataset-related mistakes. Below is the same pipeline as before, using
thin Lale wrappers around scikit-learn operators. We redefine the
pipeline to enable Lale type-checking for it.
```
lale_dataset_error = lale.operators.make_pipeline(
lale.lib.sklearn.RFE(
estimator=lale.lib.sklearn.RandomForestRegressor(n_estimators=10)),
lale.lib.sklearn.NMF())
```
When we call `fit` on the pipeline, before doing the actual training,
Lale checks that the
schema is correct at each step of the pipeline. In other words, it
checks whether the schema of the input data is valid for the first
step of the pipeline, and that the schema of the output from each step
is valid for the next step. By saving the time for training the RFE,
this completes in tens of milliseconds instead of seconds as before.
```
%%time
try:
lale_dataset_error.fit(train_X, train_y)
except ValueError as e:
message = str(e)
print(message, file=sys.stderr)
assert message.startswith('NMF.fit() invalid X: Expected sub to be a subschema of super.')
```
In this example, the schemas for `X` differ: whereas the data is an
array of arrays of unconstrained numbers, NMF expects an array of
arrays of only non-negative numbers.
### 3.5 Hyperparameter Constraint Example in Scikit-Learn
Sometimes, the validity of hyperparameters cannot be checked in
isolation. Instead, the value of one hyperparameter can restrict
which values are valid for another hyperparameter. For example,
scikit-learn imposes a conditional hyperparameter constraint between
the `svd_solver` and `n_components` arguments to PCA.
```
sklearn_constraint_error = sklearn.pipeline.make_pipeline(
sklearn.feature_selection.RFE(
estimator=sklearn.ensemble.RandomForestRegressor(n_estimators=10)),
sklearn.decomposition.PCA(svd_solver='arpack', n_components='mle'))
```
The above notebook cell completed successfully, because scikit-learn
did not yet check for the constraint. To observe the error message
with scikit-learn, we must attempt to fit the pipeline.
```
%%time
message=None
try:
sklearn_constraint_error.fit(train_X, train_y)
except ValueError as e:
message = str(e)
print(message, file=sys.stderr)
assert message.startswith("n_components='mle' cannot be a string with svd_solver='arpack'")
```
Scikit-learn implements constraint-checking as Python code with
if-statements and raise-statements. After a few seconds, we get an
exception, and the error message explains what went wrong.
### 3.6 Types for Constraint Checking
Lale specifies constraints using JSON Schemas. When you configure an
operator with actual hyperparameters, Lale immediately validates them
against their schema including constraints.
```
%%time
try:
lale_constraint_error = lale.operators.make_pipeline(
lale.lib.sklearn.RFE(
estimator=lale.lib.sklearn.RandomForestRegressor(n_estimators=10)),
PCA(svd_solver='arpack', n_components='mle'))
except jsonschema.ValidationError as e:
message = str(e)
print(message, file=sys.stderr)
assert message.startswith("Invalid configuration for pca(svd_solver='arpack', n_components='mle')")
```
Lale reports the error quicker than scikit-learn, taking only tens of
milliseconds instead of multiple seconds. The error message contains
both a natural-language description of the constraint and its formal
representation in JSON Schema. The `'anyOf'` implements an 'or', so
you can read the constraints as
```python
(not (n_components in ['mle'])) or (svd_solver in ['full', 'auto'])
```
By basic Boolean algebra, this is equivalent to an implication
```python
(n_components in ['mle']) implies (svd_solver in ['full', 'auto'])
```
Since the constraint is specified declaratively in the schema, it gets
applied wherever the schema gets used. Specifically, the constraint
gets applied both during auto-ML and during type-checking. In the
context of auto-ML, the constraint prunes the search space: it
eliminates some hyperparameter combinations so that the auto-ML tool
does not have to try them out. We have observed cases where this
pruning makes a big difference in search convergence.
## 4. Conclusion
This notebook showed additions to scikit-learn that simplify auto-ML
as well as error checking. The common foundation for both of these
additions is schemas for operators. For further reading, return to the
Lale github [repository](https://github.com/ibm/lale), where you can
find installation instructions, an FAQ, and links to further
documentation, notebooks, talks, etc.
| github_jupyter |
```
import os
import random
def load_dataset(path_dataset):
"""Load dataset into memory from text file"""
dataset = []
with open(path_dataset) as f:
words, tags = [], []
# Each line of the file corresponds to one word and tag
for line in f:
if line != '\n':
line = line.strip('\n')
if len(line.split()) > 0:
word = line.split()[0]
#tag = line.split()[-1]
else:
word = line.split()[0]
# print(line)
continue
try:
if len(word) > 0:
word = str(word)
words.append(word)
#tags.append(tag)
except Exception as e:
print('An exception was raised, skipping a word: {}'.format(e))
else:
if len(words) > 0:
#assert len(words) == len(tags)
dataset.append((words))
words, tags = [], []
return dataset
def save_dataset(dataset, save_dir,file_name):
# Create directory if it doesn't exist
print('Saving in {}...'.format(save_dir))
if not os.path.exists(save_dir):
os.makedirs(save_dir)
# Export the dataset
with open(os.path.join(save_dir, file_name), 'w') as file_sentences:
for sent in dataset:
for word in sent:
file_sentences.writelines('%s\n' %word)
file_sentences.writelines('\n')
#file_tags.write('{}\n'.format(' '.join(tags)))
file_sentences.writelines('\n')
file_sentences.writelines('\n')
print('- done.')
save_dir = 'submission_Conll'
import glob
filenames = glob.glob('output_data/*')
pred_tags_files = []
for filename in filenames:
pred_tags_file = load_dataset(filename)
pred_tags_files.append(pred_tags_file)
o = "Realizamos en la consulta de Medicina de Familia ecografía cervical clínica observándose adenopatías laterocervicales y submandibulares, fundamentalmente hipoecogénicas con tendencia a la agrupación, afectando de forma bilateral a las cadenas yugulocarotídeas, en general subcentimétricas y con hilio graso conservado, sin necrosis. La mayor mide 2,5 cm de longitud por 1,1 cm de diámetro antero-posterior. En la ecografía abdominal clínica se observa hígado y bazo con tamaño en el límite alto de la normalidad, pero considerando la altura y talla de la paciente podrían considerarse aumentados de tamaño. En ecocardioscopia clínica: Ventrículo izquierdo (VI) hiperdinámico, sin identificarse alteraciones valvulares groseras. Tamaño y función del VI y ventrículo derecho normales. Ausencia de derrame pericárdico."
len(space_remover(o))
```
root_dir = 'data/meddo/test_data/Conll_Format/'
for filename in filenames:
pred_tags_file = load_dataset(filename)
re_name = filename.split('/')[-1]
words_file = load_dataset(root_dir + re_name )
for i in range(len(pred_tags_file)):
for j in range(len(pred_tags_file[i])):
words_file[i][j] = words_file[i][j]+"\t"+pred_tags_file[i][j]
save_dataset(words_file,save_dir,re_name)
```
filenames[0]
sentences_file = glob.glob("./data/meddo/interactive/sentences/*.txt")
def generate_conll(sentences_file, filenames):
with open(sentences_file) as f, open(filenames) as q:
assert sentences_file.split("/")[-1] == filenames.split("/")[-1]
words, tags, comb_str = [], [], []
for line_sentence, line_tags in zip(f,q):
try:
assert len(space_remover(line_sentence)) == len(space_remover(line_tags))
words.append(space_remover(line_sentence))
tags.append(space_remover(line_tags))
except:
token_list = space_remover(line_sentence)
tag_list = space_remover(line_tags)
if len(token_list) <= 0 and len(tag_list) <= 0:
continue
if token_list[-1] == '\n':
token_list = token_list[:-1]
if tag_list[-1] == '\n':
tag_list = tag_list[:-1]
for i in range(len(token_list) - len(tag_list)):
tag_list.append('O')
print(len(tag_list), len(token_list))
assert len(tag_list) == len(token_list)
words.append(token_list)
tags.append(tag_list)
#print("the following file name is wrong"+ str(sentences_file.split("/")[-1]))
for tokens,tag in zip(words,tags):
for token,t in zip(tokens, tag):
if token == '\n' and t == '\n':
comb_str.append("\n")
else:
comb_str.append(token+ "\t" + t)
#print(comb_str)
#exit()
with open("./temp/"+sentences_file.split("/")[-1], 'w') as r:
for item in comb_str:
if item == '\n':
r.write("%s" % item)
else:
r.write("%s\n" % item)
import re
def space_remover(s):
out = re.findall(r'\S+|\n',s)
return out
#generate_conll(sentences_file[1], filenames[1])
#generate_conll(sentences_file[130], filenames[130])
counter = 0
for sentences_f, file_n in zip(sentences_file,filenames):
generate_conll(sentences_f, file_n)
# bert_class = 'mrm8488/bert-base-spanish-wwm-cased-finetuned-spa-squad2-es'
# from transformers import BertTokenizer
# tokenizer = BertTokenizer.from_pretrained(bert_class, do_lower_case=False)
```
| github_jupyter |
# Tracking Information Flow
We have explored how one could generate better inputs that can penetrate deeper into the program in question. While doing so, we have relied on program crashes to tell us that we have succeeded in finding problems in the program. However, that is rather simplistic. What if the behavior of the program is simply incorrect, but does not lead to a crash? Can one do better?
In this chapter, we explore in depth how to track information flows in Python, and how these flows can be used to determine whether a program behaved as expected.
```
from bookutils import YouTubeVideo
YouTubeVideo('MJ0VGzVbhYc')
```
**Prerequisites**
* You should have read the [chapter on coverage](Coverage.ipynb).
* You should have read the [chapter on probabilistic fuzzing](ProbabilisticGrammarFuzzer.ipynb).
We first set up our infrastructure so that we can make use of previously defined functions.
```
import bookutils
from typing import List, Any, Optional, Union
```
## Synopsis
<!-- Automatically generated. Do not edit. -->
To [use the code provided in this chapter](Importing.ipynb), write
```python
>>> from fuzzingbook.InformationFlow import <identifier>
```
and then make use of the following features.
This chapter provides two wrappers to Python _strings_ that allow one to track various properties. These include information on the security properties of the input, and information on originating indexes of the input string.
### Tracking String Taints
`tstr` objects are replacements for Python strings that allows to track and check _taints_ – that is, information on from where a string originated. For instance, one can mark strings that originate from third party input with a taint of "LOW", meaning that they have a low security level. The taint is passed in the constructor of a `tstr` object:
```python
>>> thello = tstr('hello', taint='LOW')
```
A `tstr` object is fully compatible with original Python strings. For instance, we can index it and access substrings:
```python
>>> thello[:4]
'hell'
```
However, the `tstr` object also stores the taint, which can be accessed using the `taint` attribute:
```python
>>> thello.taint
'LOW'
```
The neat thing about taints is that they propagate to all strings derived from the original tainted string.
Indeed, any operation from a `tstr` string that results in a string fragment produces another `tstr` object that includes the original taint. For example:
```python
>>> thello[1:2].taint # type: ignore
'LOW'
```
`tstr` objects duplicate most `str` methods, as indicated in the class diagram:

### Tracking Character Origins
`ostr` objects extend `tstr` objects by not only tracking a taint, but also the originating _indexes_ from the input string, This allows you to exactly track where individual characters came from. Assume you have a long string, which at index 100 contains the password `"joshua1234"`. Then you can save this origin information using an `ostr` as follows:
```python
>>> secret = ostr("joshua1234", origin=100, taint='SECRET')
```
The `origin` attribute of an `ostr` provides access to a list of indexes:
```python
>>> secret.origin
[100, 101, 102, 103, 104, 105, 106, 107, 108, 109]
>>> secret.taint
'SECRET'
```
`ostr` objects are compatible with Python strings, except that string operations return `ostr` objects (together with the saved origin an index information). An index of `-1` indicates that the corresponding character has no origin as supplied to the `ostr()` constructor:
```python
>>> secret_substr = (secret[0:4] + "-" + secret[6:])
>>> secret_substr.taint
'SECRET'
>>> secret_substr.origin
[100, 101, 102, 103, -1, 106, 107, 108, 109]
```
`ostr` objects duplicate most `str` methods, as indicated in the class diagram:

## A Vulnerable Database
Say we want to implement an *in-memory database* service in Python. Here is a rather flimsy attempt. We use the following dataset.
```
INVENTORY = """\
1997,van,Ford,E350
2000,car,Mercury,Cougar
1999,car,Chevy,Venture\
"""
VEHICLES = INVENTORY.split('\n')
```
Our DB is a Python class that parses its arguments and throws `SQLException` which is defined below.
```
class SQLException(Exception):
pass
```
The database is simply a Python `dict` that is exposed only through SQL queries.
```
class DB:
def __init__(self, db={}):
self.db = dict(db)
```
### Representing Tables
The database contains tables, which are created by a method call `create_table()`. Each table data structure is a pair of values. The first one is the meta data containing column names and types. The second value is a list of values in the table.
```
class DB(DB):
def create_table(self, table, defs):
self.db[table] = (defs, [])
```
The table can be retrieved using the name using the `table()` method call.
```
class DB(DB):
def table(self, t_name):
if t_name in self.db:
return self.db[t_name]
raise SQLException('Table (%s) was not found' % repr(t_name))
```
Here is an example of how to use both. We fill a table `inventory` with four columns: `year`, `kind`, `company`, and `model`. Initially, our table is empty.
```
def sample_db():
db = DB()
inventory_def = {'year': int, 'kind': str, 'company': str, 'model': str}
db.create_table('inventory', inventory_def)
return db
```
Using `table()`, we can retrieve the table definition as well as its contents.
```
db = sample_db()
db.table('inventory')
```
We also define `column()` for retrieving the column definition from a table declaration.
```
class DB(DB):
def column(self, table_decl, c_name):
if c_name in table_decl:
return table_decl[c_name]
raise SQLException('Column (%s) was not found' % repr(c_name))
db = sample_db()
decl, rows = db.table('inventory')
db.column(decl, 'year')
```
### Executing SQL Statements
The `sql()` method of `DB` executes SQL statements. It inspects its arguments, and dispatches the query based on the kind of SQL statement to be executed.
```
class DB(DB):
def do_select(self, query):
...
def do_update(self, query):
...
def do_insert(self, query):
...
def do_delete(self, query):
...
def sql(self, query):
methods = [('select ', self.do_select),
('update ', self.do_update),
('insert into ', self.do_insert),
('delete from', self.do_delete)]
for key, method in methods:
if query.startswith(key):
return method(query[len(key):])
raise SQLException('Unknown SQL (%s)' % query)
```
Here's an example of how to use the `DB` class:
```
some_db = DB()
some_db.sql('select year from inventory')
```
However, at this point, the individual methods for handling SQL statements are not yet defined. Let us do this in the next steps.
### Excursion: Implementing SQL Statements
#### Selecting Data
The `do_select()` method handles SQL `select` statements to retrieve data from a table.
```
class DB(DB):
def do_select(self, query):
FROM, WHERE = ' from ', ' where '
table_start = query.find(FROM)
if table_start < 0:
raise SQLException('no table specified')
where_start = query.find(WHERE)
select = query[:table_start]
if where_start >= 0:
t_name = query[table_start + len(FROM):where_start]
where = query[where_start + len(WHERE):]
else:
t_name = query[table_start + len(FROM):]
where = ''
_, table = self.table(t_name)
if where:
selected = self.expression_clause(table, "(%s)" % where)
selected_rows = [hm for i, data, hm in selected if data]
else:
selected_rows = table
rows = self.expression_clause(selected_rows, "(%s)" % select)
return [data for i, data, hm in rows]
```
The `expression_clause()` method is used for two purposes:
1. In the form `select` $x$, $y$, $z$ `from` $t$, it _evaluates_ (and returns) the expressions $x$, $y$, $z$ in the contexts of the selected rows.
2. If a clause `where` $p$ is given, it also evaluates $p$ in the context of the rows and includes the rows in the selection only if $p$ holds.
To evaluate expressions like $x$, $y$, $z$ or $p$, the method `expression_clause()` makes use of the Python `eval()` evaluation function.
```
class DB(DB):
def expression_clause(self, table, statement):
selected = []
for i, hm in enumerate(table):
selected.append((i, self.my_eval(statement, {}, hm), hm))
return selected
```
If `eval()` fails for whatever reason, we raise an exception:
```
class DB(DB):
def my_eval(self, statement, g, l):
try:
return eval(statement, g, l)
except Exception:
raise SQLException('Invalid WHERE (%s)' % repr(statement))
```
**Note:** Using `eval()` here introduces some important security issues, which we will discuss later in this chapter.
Here's how we can use `sql()` to issue a query. Note that the table is yet empty.
```
db = sample_db()
db.sql('select year from inventory')
db = sample_db()
db.sql('select year from inventory where year == 2018')
```
#### Inserting Data
The `do_insert()` method handles SQL `insert` statements.
```
class DB(DB):
def do_insert(self, query):
VALUES = ' values '
table_end = query.find('(')
t_name = query[:table_end].strip()
names_end = query.find(')')
decls, table = self.table(t_name)
names = [i.strip() for i in query[table_end + 1:names_end].split(',')]
# verify columns exist
for k in names:
self.column(decls, k)
values_start = query.find(VALUES)
if values_start < 0:
raise SQLException('Invalid INSERT (%s)' % repr(query))
values = [
i.strip() for i in query[values_start + len(VALUES) + 1:-1].split(',')
]
if len(names) != len(values):
raise SQLException(
'names(%s) != values(%s)' % (repr(names), repr(values)))
# dict lookups happen in C code, so we can't use that
kvs = {}
for k,v in zip(names, values):
for key,kval in decls.items():
if k == key:
kvs[key] = self.convert(kval, v)
table.append(kvs)
```
In SQL, a column can come in any supported data type. To ensure it is stored using the type originally declared, we need the ability to convert the values to specific types which is provided by `convert()`.
```
import ast
class DB(DB):
def convert(self, cast, value):
try:
return cast(ast.literal_eval(value))
except:
raise SQLException('Invalid Conversion %s(%s)' % (cast, value))
```
Here is an example of how to use the SQL `insert` command:
```
db = sample_db()
db.sql('insert into inventory (year, kind, company, model) values (1997, "van", "Ford", "E350")')
db.table('inventory')
```
With the database filled, we can also run more complex queries:
```
db.sql('select year + 1, kind from inventory')
db.sql('select year, kind from inventory where year == 1997')
```
#### Updating Data
Similarly, `do_update()` handles SQL `update` statements.
```
class DB(DB):
def do_update(self, query):
SET, WHERE = ' set ', ' where '
table_end = query.find(SET)
if table_end < 0:
raise SQLException('Invalid UPDATE (%s)' % repr(query))
set_end = table_end + 5
t_name = query[:table_end]
decls, table = self.table(t_name)
names_end = query.find(WHERE)
if names_end >= 0:
names = query[set_end:names_end]
where = query[names_end + len(WHERE):]
else:
names = query[set_end:]
where = ''
sets = [[i.strip() for i in name.split('=')]
for name in names.split(',')]
# verify columns exist
for k, v in sets:
self.column(decls, k)
if where:
selected = self.expression_clause(table, "(%s)" % where)
updated = [hm for i, d, hm in selected if d]
else:
updated = table
for hm in updated:
for k, v in sets:
# we can not do dict lookups because it is implemented in C.
for key, kval in decls.items():
if key == k:
hm[key] = self.convert(kval, v)
return "%d records were updated" % len(updated)
```
Here is an example. Let us first fill the database again with values:
```
db = sample_db()
db.sql('insert into inventory (year, kind, company, model) values (1997, "van", "Ford", "E350")')
db.sql('select year from inventory')
```
Now we can update things:
```
db.sql('update inventory set year = 1998 where year == 1997')
db.sql('select year from inventory')
db.table('inventory')
```
#### Deleting Data
Finally, SQL `delete` statements are handled by `do_delete()`.
```
class DB(DB):
def do_delete(self, query):
WHERE = ' where '
table_end = query.find(WHERE)
if table_end < 0:
raise SQLException('Invalid DELETE (%s)' % query)
t_name = query[:table_end].strip()
_, table = self.table(t_name)
where = query[table_end + len(WHERE):]
selected = self.expression_clause(table, "%s" % where)
deleted = [i for i, d, hm in selected if d]
for i in sorted(deleted, reverse=True):
del table[i]
return "%d records were deleted" % len(deleted)
```
Here is an example. Let us first fill the database again with values:
```
db = sample_db()
db.sql('insert into inventory (year, kind, company, model) values (1997, "van", "Ford", "E350")')
db.sql('select year from inventory')
```
Now we can delete data:
```
db.sql('delete from inventory where company == "Ford"')
```
Our database is now empty:
```
db.sql('select year from inventory')
```
### End of Excursion
Here is how our database can be used.
```
db = DB()
```
We first create a table in our database with the correct data types.
```
inventory_def = {'year': int, 'kind': str, 'company': str, 'model': str}
db.create_table('inventory', inventory_def)
```
Here is a simple convenience function to update the table using our dataset.
```
def update_inventory(sqldb, vehicle):
inventory_def = sqldb.db['inventory'][0]
k, v = zip(*inventory_def.items())
val = [repr(cast(val)) for cast, val in zip(v, vehicle.split(','))]
sqldb.sql('insert into inventory (%s) values (%s)' % (','.join(k),
','.join(val)))
for V in VEHICLES:
update_inventory(db, V)
```
Our database now contains the same dataset as `VEHICLES` under `INVENTORY` table.
```
db.db
```
Here is a sample select statement.
```
db.sql('select year,kind from inventory')
db.sql("select company,model from inventory where kind == 'car'")
```
We can run updates on it.
```
db.sql("update inventory set year = 1998, company = 'Suzuki' where kind == 'van'")
db.db
```
It can even do mathematics on the fly!
```
db.sql('select int(year)+10 from inventory')
```
Adding a new row to our table.
```
db.sql("insert into inventory (year, kind, company, model) values (1, 'charriot', 'Rome', 'Quadriga')")
db.db
```
Which we then delete.
```
db.sql("delete from inventory where year < 1900")
```
### Fuzzing SQL
To verify that everything is OK, let us fuzz. First we define our grammar.
#### Excursion: Defining a SQL grammar
```
import string
from Grammars import START_SYMBOL, Grammar, Expansion, \
is_valid_grammar, extend_grammar
EXPR_GRAMMAR: Grammar = {
"<start>": ["<expr>"],
"<expr>": ["<bexpr>", "<aexpr>", "(<expr>)", "<term>"],
"<bexpr>": [
"<aexpr><lt><aexpr>",
"<aexpr><gt><aexpr>",
"<expr>==<expr>",
"<expr>!=<expr>",
],
"<aexpr>": [
"<aexpr>+<aexpr>", "<aexpr>-<aexpr>", "<aexpr>*<aexpr>",
"<aexpr>/<aexpr>", "<word>(<exprs>)", "<expr>"
],
"<exprs>": ["<expr>,<exprs>", "<expr>"],
"<lt>": ["<"],
"<gt>": [">"],
"<term>": ["<number>", "<word>"],
"<number>": ["<integer>.<integer>", "<integer>", "-<number>"],
"<integer>": ["<digit><integer>", "<digit>"],
"<word>": ["<word><letter>", "<word><digit>", "<letter>"],
"<digit>":
list(string.digits),
"<letter>":
list(string.ascii_letters + '_:.')
}
assert is_valid_grammar(EXPR_GRAMMAR)
PRINTABLE_CHARS: List[str] = [i for i in string.printable
if i not in "<>'\"\t\n\r\x0b\x0c\x00"] + ['<lt>', '<gt>']
INVENTORY_GRAMMAR = extend_grammar(EXPR_GRAMMAR,
{
'<start>': ['<query>'],
'<query>': [
'select <exprs> from <table>',
'select <exprs> from <table> where <bexpr>',
'insert into <table> (<names>) values (<literals>)',
'update <table> set <assignments> where <bexpr>',
'delete from <table> where <bexpr>',
],
'<table>': ['<word>'],
'<names>': ['<column>,<names>', '<column>'],
'<column>': ['<word>'],
'<literals>': ['<literal>', '<literal>,<literals>'],
'<literal>': ['<number>', "'<chars>'"],
'<assignments>': ['<kvp>,<assignments>', '<kvp>'],
'<kvp>': ['<column>=<value>'],
'<value>': ['<word>'],
'<chars>': ['<char>', '<char><chars>'],
'<char>': PRINTABLE_CHARS, # type: ignore
})
assert is_valid_grammar(INVENTORY_GRAMMAR)
```
As can be seen from the source of our database, the functions always check whether the table name is correct. Hence, we modify the grammar to choose our particular table so that it will have a better chance of reaching deeper. We will see in the later sections how this can be done automatically.
```
INVENTORY_GRAMMAR_F = extend_grammar(INVENTORY_GRAMMAR,
{'<table>': ['inventory']})
```
#### End of Excursion
```
from GrammarFuzzer import GrammarFuzzer
gf = GrammarFuzzer(INVENTORY_GRAMMAR_F)
for _ in range(10):
query = gf.fuzz()
print(repr(query))
try:
res = db.sql(query)
print(repr(res))
except SQLException as e:
print("> ", e)
pass
except:
traceback.print_exc()
break
print()
```
Fuzzing does not seem to have triggered any crashes. However, are crashes the only errors that we should be worried about?
### The Evil of Eval
In our database implementation – notably in the `expression_clause()` method -, we have made use of `eval()` to evaluate expressions using the Python interpreter. This allows us to unleash the full power of Python expressions within our SQL statements.
```
db.sql('select year from inventory where year < 2000')
```
In the above query, the clause `year < 2000` is evaluated using `expression_clause()` using Python in the context of each row; hence, `year < 2000` evaluates to either `True` or `False`.
The same holds for the expressions being `select`ed:
```
db.sql('select year - 1900 if year < 2000 else year - 2000 from inventory')
```
This works because `year - 1900 if year < 2000 else year - 2000` is a valid Python expression. (It is not a valid SQL expression, though.)
The problem with the above is that there is _no limitation_ to what the Python expression can do. What if the user tries the following?
```
db.sql('select __import__("os").popen("pwd").read() from inventory')
```
The above statement effectively reads from the users' file system. Instead of `os.popen("pwd").read()`, it could execute arbitrary Python commands – to access data, install software, run a background process. This is where "the full power of Python expressions" turns back on us.
What we want is to allow our _program_ to make full use of its power; yet, the _user_ (or any third party) should not be entrusted to do the same. Hence, we need to differentiate between (trusted) _input from the program_ and (untrusted) _input from the user_.
One method that allows such differentiation is that of *dynamic taint analysis*. The idea is to identify the functions that accept user input as *sources* that *taint* any string that comes in through them, and those functions that perform dangerous operations as *sinks*. Finally we bless certain functions as *taint sanitizers*. The idea is that an input from the source should never reach the sink without undergoing sanitization first. This allows us to use a stronger oracle than simply checking for crashes.
## Tracking String Taints
There are various levels of taint tracking that one can perform. The simplest is to track that a string fragment originated in a specific environment, and has not undergone a taint removal process. For this, we simply need to wrap the original string with an environment identifier (the _taint_) with `tstr`, and produce `tstr` instances on each operation that results in another string fragment. The attribute `taint` holds a label identifying the environment this instance was derived.
### A Class for Tainted Strings
For capturing information flows we need a new string class. The idea is to use the new tainted string class `tstr` as a wrapper on the original `str` class. However, `str` is an *immutable* class. Hence, it does not call its `__init__()` method after being constructed. This means that any subclasses of `str` also will not get the `__init__()` method called. If we want to get our initialization routine called, we need to [hook into `__new__()`](https://docs.python.org/3/reference/datamodel.html#basic-customization) and return an instance of our own class. We combine this with our initialization code in `__init__()`.
```
class tstr(str):
"""Wrapper for strings, saving taint information"""
def __new__(cls, value, *args, **kw):
"""Create a tstr() instance. Used internally."""
return str.__new__(cls, value)
def __init__(self, value: Any, taint: Any = None, **kwargs) -> None:
"""Constructor.
`value` is the string value the `tstr` object is to be constructed from.
`taint` is an (optional) taint to be propagated to derived strings."""
self.taint: Any = taint
class tstr(tstr):
def __repr__(self) -> tstr:
"""Return a representation."""
return tstr(str.__repr__(self), taint=self.taint)
class tstr(tstr):
def __str__(self) -> str:
"""Convert to string"""
return str.__str__(self)
```
For example, if we wrap `"hello"` in `tstr`, then we should be able to access its taint:
```
thello: tstr = tstr('hello', taint='LOW')
thello.taint
repr(thello).taint # type: ignore
```
By default, when we wrap a string, it is tainted. Hence we also need a way to clear the taint in the string. One way is to simply return a `str` instance as above. However, one may sometimes wish to remove the taint from an existing instance. This is accomplished with `clear_taint()`. During `clear_taint()`, we simply set the taint to `None`. This method comes with a pair method `has_taint()` which checks whether a `tstr` instance is currently origined.
```
class tstr(tstr):
def clear_taint(self):
"""Remove taint"""
self.taint = None
return self
def has_taint(self):
"""Check if taint is present"""
return self.taint is not None
```
### String Operators
To propagate the taint, we have to extend string functions, such as operators. We can do so in one single big step, overloading all string methods and operators.
When we create a new string from an existing tainted string, we propagate its taint.
```
class tstr(tstr):
def create(self, s):
return tstr(s, taint=self.taint)
```
The `make_str_wrapper()` function creates a wrapper around an existing string method which attaches the taint to the result of the method:
```
class tstr(tstr):
@staticmethod
def make_str_wrapper(fun):
"""Make `fun` (a `str` method) a method in `tstr`"""
def proxy(self, *args, **kwargs):
res = fun(self, *args, **kwargs)
return self.create(res)
if hasattr(fun, '__doc__'):
# Copy docstring
proxy.__doc__ = fun.__doc__
return proxy
```
We do this for all string methods that return a string:
```
def informationflow_init_1():
for name in ['__format__', '__mod__', '__rmod__', '__getitem__',
'__add__', '__mul__', '__rmul__',
'capitalize', 'casefold', 'center', 'encode',
'expandtabs', 'format', 'format_map', 'join',
'ljust', 'lower', 'lstrip', 'replace',
'rjust', 'rstrip', 'strip', 'swapcase', 'title', 'translate', 'upper']:
fun = getattr(str, name)
setattr(tstr, name, tstr.make_str_wrapper(fun))
informationflow_init_1()
INITIALIZER_LIST = [informationflow_init_1]
def initialize():
for fn in INITIALIZER_LIST:
fn()
```
The one missing operator is `+` with a regular string on the left side and a tainted string on the right side. Python supports a `__radd__()` method which is invoked if the associated object is used on the right side of an addition.
```
class tstr(tstr):
def __radd__(self, value):
"""Return value + self, as a `tstr` object"""
return self.create(value + str(self))
```
With this, we are already done. Let us create a string `thello` with a taint `LOW`.
```
thello = tstr('hello', taint='LOW')
```
Now, any substring will also be tainted:
```
thello[0].taint # type: ignore
thello[1:3].taint # type: ignore
```
String additions will return a `tstr` object with the taint:
```
(tstr('foo', taint='HIGH') + 'bar').taint # type: ignore
```
Our `__radd__()` method ensures this also works if the `tstr` occurs on the right side of a string addition:
```
('foo' + tstr('bar', taint='HIGH')).taint # type: ignore
thello += ', world' # type: ignore
thello.taint # type: ignore
```
Other operators such as multiplication also work:
```
(thello * 5).taint # type: ignore
('hw %s' % thello).taint # type: ignore
(tstr('hello %s', taint='HIGH') % 'world').taint # type: ignore
```
## Tracking Untrusted Input
So, what can one do with tainted strings? We reconsider the `DB` example. We define a "better" `TrustedDB` which only accepts strings tainted as `"TRUSTED"`.
```
class TrustedDB(DB):
def sql(self, s):
assert isinstance(s, tstr), "Need a tainted string"
assert s.taint == 'TRUSTED', "Need a string with trusted taint"
return super().sql(s)
```
Feeding a string with an "unknown" (i.e., non-existing) trust level will cause `TrustedDB` to fail:
```
bdb = TrustedDB(db.db)
from ExpectError import ExpectError
with ExpectError():
bdb.sql("select year from INVENTORY")
```
Additionally any user input would be originally tagged with `"UNTRUSTED"` as taint. If we place an untrusted string into our better calculator, it will also fail:
```
bad_user_input = tstr('__import__("os").popen("ls").read()', taint='UNTRUSTED')
with ExpectError():
bdb.sql(bad_user_input)
```
Hence, somewhere along the computation, we have to turn the "untrusted" inputs into "trusted" strings. This process is called *sanitization*. A simple sanitization function for our purposes could ensure that the input consists only of few allowed characters (not including letters or quotes); if this is the case, then the input gets a new `"TRUSTED"` taint. If not, we turn the string into an (untrusted) empty string; other alternatives would be to raise an error or to escape or delete "untrusted" characters.
```
import re
def sanitize(user_input):
assert isinstance(user_input, tstr)
if re.match(
r'^select +[-a-zA-Z0-9_, ()]+ from +[-a-zA-Z0-9_, ()]+$', user_input):
return tstr(user_input, taint='TRUSTED')
else:
return tstr('', taint='UNTRUSTED')
good_user_input = tstr("select year,model from inventory", taint='UNTRUSTED')
sanitized_input = sanitize(good_user_input)
sanitized_input
sanitized_input.taint
bdb.sql(sanitized_input)
```
Let us now try out our untrusted input:
```
sanitized_input = sanitize(bad_user_input)
sanitized_input
sanitized_input.taint
with ExpectError():
bdb.sql(sanitized_input)
```
In a similar fashion, we can prevent SQL and code injections discussed in [the chapter on Web fuzzing](WebFuzzer.ipynb).
## Taint Aware Fuzzing
We can also use tainting to _direct fuzzing to those grammar rules that are likely to generate dangerous inputs._ The idea here is to identify inputs generated by our fuzzer that lead to untrusted execution. First we define the exception to be thrown when a tainted value reaches a dangerous operation.
```
class Tainted(Exception):
def __init__(self, v):
self.v = v
def __str__(self):
return 'Tainted[%s]' % self.v
```
### TaintedDB
Next, since `my_eval()` is the most dangerous operation in the `DB` class, we define a new class `TaintedDB` that overrides the `my_eval()` to throw an exception whenever an untrusted string reaches this part.
```
class TaintedDB(DB):
def my_eval(self, statement, g, l):
if statement.taint != 'TRUSTED':
raise Tainted(statement)
try:
return eval(statement, g, l)
except:
raise SQLException('Invalid SQL (%s)' % repr(statement))
```
We initialize an instance of `TaintedDB`
```
tdb = TaintedDB()
tdb.db = db.db
```
Then we start fuzzing.
```
import traceback
for _ in range(10):
query = gf.fuzz()
print(repr(query))
try:
res = tdb.sql(tstr(query, taint='UNTRUSTED'))
print(repr(res))
except SQLException as e:
pass
except Tainted as e:
print("> ", e)
except:
traceback.print_exc()
break
print()
```
One can see that `insert`, `update`, `select` and `delete` statements on an existing table lead to taint exceptions. We can now focus on these specific kinds of inputs. However, this is not the only thing we can do. We will see how we can identify specific portions of input that reached tainted execution using character origins in the later sections. But before that, we explore other uses of taints.
## Preventing Privacy Leaks
Using taints, we can also ensure that secret information does not leak out. We can assign a special taint `"SECRET"` to strings whose information must not leak out:
```
secrets = tstr('<Plenty of secret keys>', taint='SECRET')
```
Accessing any substring of `secrets` will propagate the taint:
```
secrets[1:3].taint # type: ignore
```
Consider the _heartbeat_ security leak from [the chapter on Fuzzing](Fuzzer.ipynb), in which a server would accidentally reply not only the user input sent to it, but also secret memory. If the reply consists only of the user input, there is no taint associated with it:
```
user_input = "hello"
reply = user_input
isinstance(reply, tstr)
```
If, however, the reply contains _any_ part of the secret, the reply will be tainted:
```
reply = user_input + secrets[0:5]
reply
reply.taint # type: ignore
```
The output function of our server would now ensure that the data sent back does not contain any secret information:
```
def send_back(s):
assert not isinstance(s, tstr) and not s.taint == 'SECRET' # type: ignore
...
with ExpectError():
send_back(reply)
```
Our `tstr` solution can help to identify information leaks – but it is by no means complete. If we actually take the `heartbeat()` implementation from [the chapter on Fuzzing](Fuzzer.ipynb), we will see that _any_ reply is marked as `SECRET` – even those not even accessing secret memory:
```
from Fuzzer import heartbeat
reply = heartbeat('hello', 5, memory=secrets)
reply.taint # type: ignore
```
Why is this? If we look into the implementation of `heartbeat()`, we will see that it first builds a long string `memory` from the (non-secret) reply and the (secret) memory, before returning the first characters from `memory`.
```python
# Store reply in memory
memory = reply + memory[len(reply):]
```
At this point, the whole memory still is tainted as `SECRET`, _including_ the non-secret part from `reply`.
We may be able to circumvent the issue by tagging the `reply` as `PUBLIC` – but then, this taint would be in conflict with the `SECRET` tag of `memory`. What happens if we compose a string from two differently tainted strings?
```
thilo = tstr("High", taint='HIGH') + tstr("Low", taint='LOW')
```
It turns out that in this case, the `__add__()` method takes precedence over the `__radd__()` method, which means that the right-hand `"Low"` string is treated as a regular (non-tainted) string.
```
thilo
thilo.taint # type: ignore
```
We could set up the `__add__()` and other methods with special handling for conflicting taints. However, the way this conflict should be resolved would be highly _application-dependent_:
* If we use taints to indicate _privacy levels_, `SECRET` privacy should take precedence over `PUBLIC` privacy. Any combination of a `SECRET`-tainted string and a `PUBLIC`-tainted string thus should have a `SECRET` taint.
* If we use taints to indicate _origins_ of information, an `UNTRUSTED` origin should take precedence over a `TRUSTED` origin. Any combination of an `UNTRUSTED`-tainted string and a `TRUSTED`-tainted string thus should have an `UNTRUSTED` taint.
Of course, such conflict resolutions can be implemented. But even so, they will not help us in the `heartbeat()` example differentiating secret from non-secret output data.
## Tracking Individual Characters
Fortunately, there is a better, more generic way to solve the above problems. The key to composition of differently tainted strings is to assign taints not only to strings, but actually to every bit of information – in our case, characters. If every character has a taint on its own, a new composition of characters will simply inherit this very taint _per character_. To this end, we introduce a second bit of information named _origin_.
Distinguishing various untrusted sources may be accomplished by origining each instance as separate instance (called *colors* in dynamic origin research). You will see an instance of this technique in the chapter on [Grammar Mining](GrammarMiner.ipynb).
In this section, we carry *character level* origins. That is, given a fragment that resulted from a portion of the original origined string, one will be able to tell which portion of the input string the fragment was taken from. In essence, each input character index from an origined source gets its own color.
More complex origining such as *bitmap origins* are possible where a single character may result from multiple origined character indexes (such as *checksum* operations on strings). We do not consider these in this chapter.
### A Class for Tracking Character Origins
Let us introduce a class `ostr` which, like `tstr`, carries a taint for each string, and additionally an _origin_ for each character that indicates its source. It is a consecutive number in a particular range (by default, starting with zero) indicating its _position_ within a specific origin.
```
class ostr(str):
"""Wrapper for strings, saving taint and origin information"""
DEFAULT_ORIGIN = 0
def __new__(cls, value, *args, **kw):
"""Create an ostr() instance. Used internally."""
return str.__new__(cls, value)
def __init__(self, value: Any, taint: Any = None,
origin: Optional[Union[int, List[int]]] = None, **kwargs) -> None:
"""Constructor.
`value` is the string value the `ostr` object is to be constructed from.
`taint` is an (optional) taint to be propagated to derived strings.
`origin` (optional) is either
- an integer denoting the index of the first character in `value`, or
- a list of integers denoting the origins of the characters in `value`,
"""
self.taint = taint
if origin is None:
origin = ostr.DEFAULT_ORIGIN
if isinstance(origin, int):
self.origin = list(range(origin, origin + len(self)))
else:
self.origin = origin
assert len(self.origin) == len(self)
```
As with `tstr`, above, we implement methods for conversion into (regular) Python strings:
```
class ostr(ostr):
def create(self, s):
return ostr(s, taint=self.taint, origin=self.origin)
class ostr(ostr):
UNKNOWN_ORIGIN = -1
def __repr__(self):
# handle escaped chars
origin = [ostr.UNKNOWN_ORIGIN]
for s, o in zip(str(self), self.origin):
origin.extend([o] * (len(repr(s)) - 2))
origin.append(ostr.UNKNOWN_ORIGIN)
return ostr(str.__repr__(self), taint=self.taint, origin=origin)
class ostr(ostr):
def __str__(self):
return str.__str__(self)
```
By default, character origins start with `0`:
```
othello = ostr('hello')
assert othello.origin == [0, 1, 2, 3, 4]
```
We can also specify the starting origin as below -- `6..10`
```
tworld = ostr('world', origin=6)
assert tworld.origin == [6, 7, 8, 9, 10]
a = ostr("hello\tworld")
repr(a).origin # type: ignore
```
`str()` returns a `str` instance without origin or taint information:
```
assert type(str(othello)) == str
```
`repr()`, however, keeps the origin information for the original string:
```
repr(othello)
repr(othello).origin # type: ignore
```
Just as with taints, we can clear origins and check whether an origin is present:
```
class ostr(ostr):
def clear_taint(self):
self.taint = None
return self
def has_taint(self):
return self.taint is not None
class ostr(ostr):
def clear_origin(self):
self.origin = [self.UNKNOWN_ORIGIN] * len(self)
return self
def has_origin(self):
return any(origin != self.UNKNOWN_ORIGIN for origin in self.origin)
othello = ostr('Hello')
assert othello.has_origin()
othello.clear_origin()
assert not othello.has_origin()
```
In the remainder of this section, we re-implement various string methods such that they also keep track of origins. If this is too tedious for you, jump right [to the next section](#Checking-Origins) which gives a number of usage examples.
### Excursion: Implementing String Methods
#### Create
We need to create new substrings that are wrapped in `ostr` objects. However, we also want to allow our subclasses to create their own instances. Hence we again provide a `create()` method that produces a new `ostr` instance.
```
class ostr(ostr):
def create(self, res, origin=None):
return ostr(res, taint=self.taint, origin=origin)
othello = ostr('hello', taint='HIGH')
otworld = othello.create('world', origin=6)
otworld.origin
otworld.taint
assert (othello.origin, otworld.origin) == (
[0, 1, 2, 3, 4], [6, 7, 8, 9, 10])
```
#### Index
In Python, indexing is provided through `__getitem__()`. Indexing on positive integers is simple enough. However, it has two additional wrinkles. The first is that, if the index is negative, that many characters are counted from the end of the string which lies just after the last character. That is, the last character has a negative index `-1`
```
class ostr(ostr):
def __getitem__(self, key):
res = super().__getitem__(key)
if isinstance(key, int):
key = len(self) + key if key < 0 else key
return self.create(res, [self.origin[key]])
elif isinstance(key, slice):
return self.create(res, self.origin[key])
else:
assert False
ohello = ostr('hello', taint='HIGH')
assert (ohello[0], ohello[-1]) == ('h', 'o')
ohello[0].taint
```
The other wrinkle is that `__getitem__()` can accept a slice. We discuss this next.
#### Slices
The Python `slice` operator `[n:m]` relies on the object being an `iterator`. Hence, we define the `__iter__()` method, which returns a custom `iterator`.
```
class ostr(ostr):
def __iter__(self):
return ostr_iterator(self)
```
The `__iter__()` method requires a supporting `iterator` object. The `iterator` is used to save the state of the current iteration, which it does by keeping a reference to the original `ostr`, and the current index of iteration `_str_idx`.
```
class ostr_iterator():
def __init__(self, ostr):
self._ostr = ostr
self._str_idx = 0
def __next__(self):
if self._str_idx == len(self._ostr):
raise StopIteration
# calls ostr getitem should be ostr
c = self._ostr[self._str_idx]
assert isinstance(c, ostr)
self._str_idx += 1
return c
```
Bringing all these together:
```
thw = ostr('hello world', taint='HIGH')
thw[0:5]
assert thw[0:5].has_taint()
assert thw[0:5].has_origin()
thw[0:5].taint
thw[0:5].origin
```
#### Splits
```
def make_split_wrapper(fun):
def proxy(self, *args, **kwargs):
lst = fun(self, *args, **kwargs)
return [self.create(elem) for elem in lst]
return proxy
for name in ['split', 'rsplit', 'splitlines']:
fun = getattr(str, name)
setattr(ostr, name, make_split_wrapper(fun))
othello = ostr('hello world', taint='LOW')
othello == 'hello world'
othello.split()[0].taint # type: ignore
```
(Exercise for the reader: handle _partitions_, i.e., splitting a string by substrings)
#### Concatenation
If two origined strings are concatenated together, it may be desirable to transfer the origins from each to the corresponding portion of the resulting string. The concatenation of strings is accomplished by overriding `__add__()`.
```
class ostr(ostr):
def __add__(self, other):
if isinstance(other, ostr):
return self.create(str.__add__(self, other),
(self.origin + other.origin))
else:
return self.create(str.__add__(self, other),
(self.origin + [self.UNKNOWN_ORIGIN for i in other]))
```
```
othello = ostr("hello")
otworld = ostr("world", origin=6)
othw = othello + otworld
assert othw.origin == [0, 1, 2, 3, 4, 6, 7, 8, 9, 10] # type: ignore
```
What if a `ostr` is concatenated with a `str`?
```
space = " "
th_w = othello + space + otworld
assert th_w.origin == [
0,
1,
2,
3,
4,
ostr.UNKNOWN_ORIGIN,
ostr.UNKNOWN_ORIGIN,
6,
7,
8,
9,
10]
```
One wrinkle here is that when adding a `ostr` and a `str`, the user may place the `str` first, in which case, the `__add__()` method will be called on the `str` instance. Not on the `ostr` instance. However, Python provides a solution. If one defines `__radd__()` on the `ostr` instance, that method will be called rather than `str.__add__()`
```
class ostr(ostr):
def __radd__(self, other):
origin = other.origin if isinstance(other, ostr) else [
self.UNKNOWN_ORIGIN for i in other]
return self.create(str.__add__(other, self), (origin + self.origin))
```
We test it out:
```
shello = "hello"
otworld = ostr("world")
thw = shello + otworld
assert thw.origin == [ostr.UNKNOWN_ORIGIN] * len(shello) + [0, 1, 2, 3, 4] # type: ignore
```
These methods: `slicing` and `concatenation` is sufficient to implement other string methods that result in a string, and does not change the character underneath (i.e no case change). Hence, we look at a helper method next.
#### Extract Origin String
Given a specific input index, the method `x()` extracts the corresponding origined portion from a `ostr`. As a convenience it supports `slices` along with `ints`.
```
class ostr(ostr):
class TaintException(Exception):
pass
def x(self, i=0):
"""Extract substring at index/slice `i`"""
if not self.origin:
raise origin.TaintException('Invalid request idx')
if isinstance(i, int):
return [self[p]
for p in [k for k, j in enumerate(self.origin) if j == i]]
elif isinstance(i, slice):
r = range(i.start or 0, i.stop or len(self), i.step or 1)
return [self[p]
for p in [k for k, j in enumerate(self.origin) if j in r]]
thw = ostr('hello world', origin=100)
assert thw.x(101) == ['e']
assert thw.x(slice(101, 105)) == ['e', 'l', 'l', 'o']
```
#### Replace
The `replace()` method replaces a portion of the string with another.
```
class ostr(ostr):
def replace(self, a, b, n=None):
old_origin = self.origin
b_origin = b.origin if isinstance(
b, ostr) else [self.UNKNOWN_ORIGIN] * len(b)
mystr = str(self)
i = 0
while True:
if n and i >= n:
break
idx = mystr.find(a)
if idx == -1:
break
last = idx + len(a)
mystr = mystr.replace(a, b, 1)
partA, partB = old_origin[0:idx], old_origin[last:]
old_origin = partA + b_origin + partB
i += 1
return self.create(mystr, old_origin)
my_str = ostr("aa cde aa")
res = my_str.replace('aa', 'bb')
assert res, res.origin == ('bb', 'cde', 'bb',
[ostr.UNKNOWN_ORIGIN, ostr.UNKNOWN_ORIGIN,
2, 3, 4, 5, 6,
ostr.UNKNOWN_ORIGIN, ostr.UNKNOWN_ORIGIN])
my_str = ostr("aa cde aa")
res = my_str.replace('aa', ostr('bb', origin=100))
assert (
res, res.origin) == (
('bb cde bb'), [
100, 101, 2, 3, 4, 5, 6, 100, 101])
```
#### Split
We essentially have to re-implement split operations, and split by space is slightly different from other splits.
```
class ostr(ostr):
def _split_helper(self, sep, splitted):
result_list = []
last_idx = 0
first_idx = 0
sep_len = len(sep)
for s in splitted:
last_idx = first_idx + len(s)
item = self[first_idx:last_idx]
result_list.append(item)
first_idx = last_idx + sep_len
return result_list
def _split_space(self, splitted):
result_list = []
last_idx = 0
first_idx = 0
sep_len = 0
for s in splitted:
last_idx = first_idx + len(s)
item = self[first_idx:last_idx]
result_list.append(item)
v = str(self[last_idx:])
sep_len = len(v) - len(v.lstrip(' '))
first_idx = last_idx + sep_len
return result_list
def rsplit(self, sep=None, maxsplit=-1):
splitted = super().rsplit(sep, maxsplit)
if not sep:
return self._split_space(splitted)
return self._split_helper(sep, splitted)
def split(self, sep=None, maxsplit=-1):
splitted = super().split(sep, maxsplit)
if not sep:
return self._split_space(splitted)
return self._split_helper(sep, splitted)
my_str = ostr('ab cdef ghij kl')
ab, cdef, ghij, kl = my_str.rsplit(sep=' ')
assert (ab.origin, cdef.origin, ghij.origin,
kl.origin) == ([0, 1], [3, 4, 5, 6], [8, 9, 10, 11], [13, 14])
my_str = ostr('ab cdef ghij kl', origin=list(range(0, 15)))
ab, cdef, ghij, kl = my_str.rsplit(sep=' ')
assert(ab.origin, cdef.origin, kl.origin) == ([0, 1], [3, 4, 5, 6], [13, 14])
my_str = ostr('ab cdef ghij kl', origin=100, taint='HIGH')
ab, cdef, ghij, kl = my_str.rsplit()
assert (ab.origin, cdef.origin, ghij.origin,
kl.origin) == ([100, 101], [105, 106, 107, 108], [110, 111, 112, 113],
[118, 119])
my_str = ostr('ab cdef ghij kl', origin=list(range(0, 20)), taint='HIGH')
ab, cdef, ghij, kl = my_str.split()
assert (ab.origin, cdef.origin, kl.origin) == ([0, 1], [5, 6, 7, 8], [18, 19])
assert ab.taint == 'HIGH'
```
#### Strip
```
class ostr(ostr):
def strip(self, cl=None):
return self.lstrip(cl).rstrip(cl)
def lstrip(self, cl=None):
res = super().lstrip(cl)
i = self.find(res)
return self[i:]
def rstrip(self, cl=None):
res = super().rstrip(cl)
return self[0:len(res)]
my_str1 = ostr(" abc ")
v = my_str1.strip()
assert v, v.origin == ('abc', [2, 3, 4])
my_str1 = ostr(" abc ")
v = my_str1.lstrip()
assert (v, v.origin) == ('abc ', [2, 3, 4, 5, 6])
my_str1 = ostr(" abc ")
v = my_str1.rstrip()
assert (v, v.origin) == (' abc', [0, 1, 2, 3, 4])
```
#### Expand Tabs
```
class ostr(ostr):
def expandtabs(self, n=8):
parts = self.split('\t')
res = super().expandtabs(n)
all_parts = []
for i, p in enumerate(parts):
all_parts.extend(p.origin)
if i < len(parts) - 1:
l = len(all_parts) % n
all_parts.extend([p.origin[-1]] * l)
return self.create(res, all_parts)
my_s = str("ab\tcd")
my_ostr = ostr("ab\tcd")
v1 = my_s.expandtabs(4)
v2 = my_ostr.expandtabs(4)
assert str(v1) == str(v2)
assert (len(v1), repr(v2), v2.origin) == (6, "'ab cd'", [0, 1, 1, 1, 3, 4])
class ostr(ostr):
def join(self, iterable):
mystr = ''
myorigin = []
sep_origin = self.origin
lst = list(iterable)
for i, s in enumerate(lst):
sorigin = s.origin if isinstance(s, ostr) else [
self.UNKNOWN_ORIGIN] * len(s)
myorigin.extend(sorigin)
mystr += str(s)
if i < len(lst) - 1:
myorigin.extend(sep_origin)
mystr += str(self)
res = super().join(iterable)
assert len(res) == len(mystr)
return self.create(res, myorigin)
my_str = ostr("ab cd", origin=100)
(v1, v2), v3 = my_str.split(), 'ef'
assert (v1.origin, v2.origin) == ([100, 101], [103, 104]) # type: ignore
v4 = ostr('').join([v2, v3, v1])
assert (
v4, v4.origin) == (
'cdefab', [
103, 104, ostr.UNKNOWN_ORIGIN, ostr.UNKNOWN_ORIGIN, 100, 101])
my_str = ostr("ab cd", origin=100)
(v1, v2), v3 = my_str.split(), 'ef'
assert (v1.origin, v2.origin) == ([100, 101], [103, 104]) # type: ignore
v4 = ostr(',').join([v2, v3, v1])
assert (v4, v4.origin) == ('cd,ef,ab',
[103, 104, 0, ostr.UNKNOWN_ORIGIN, ostr.UNKNOWN_ORIGIN, 0, 100, 101]) # type: ignore
```
#### Partitions
```
class ostr(ostr):
def partition(self, sep):
partA, sep, partB = super().partition(sep)
return (self.create(partA, self.origin[0:len(partA)]),
self.create(sep,
self.origin[len(partA):len(partA) + len(sep)]),
self.create(partB, self.origin[len(partA) + len(sep):]))
def rpartition(self, sep):
partA, sep, partB = super().rpartition(sep)
return (self.create(partA, self.origin[0:len(partA)]),
self.create(sep,
self.origin[len(partA):len(partA) + len(sep)]),
self.create(partB, self.origin[len(partA) + len(sep):]))
```
#### Justify
```
class ostr(ostr):
def ljust(self, width, fillchar=' '):
res = super().ljust(width, fillchar)
initial = len(res) - len(self)
if isinstance(fillchar, tstr):
t = fillchar.x()
else:
t = self.UNKNOWN_ORIGIN
return self.create(res, [t] * initial + self.origin)
class ostr(ostr):
def rjust(self, width, fillchar=' '):
res = super().rjust(width, fillchar)
final = len(res) - len(self)
if isinstance(fillchar, tstr):
t = fillchar.x()
else:
t = self.UNKNOWN_ORIGIN
return self.create(res, self.origin + [t] * final)
```
#### mod
```
class ostr(ostr):
def __mod__(self, s):
# nothing else implemented for the time being
assert isinstance(s, str)
s_origin = s.origin if isinstance(
s, ostr) else [self.UNKNOWN_ORIGIN] * len(s)
i = self.find('%s')
assert i >= 0
res = super().__mod__(s)
r_origin = self.origin[:]
r_origin[i:i + 2] = s_origin
return self.create(res, origin=r_origin)
class ostr(ostr):
def __rmod__(self, s):
# nothing else implemented for the time being
assert isinstance(s, str)
r_origin = s.origin if isinstance(
s, ostr) else [self.UNKNOWN_ORIGIN] * len(s)
i = s.find('%s')
assert i >= 0
res = super().__rmod__(s)
s_origin = self.origin[:]
r_origin[i:i + 2] = s_origin
return self.create(res, origin=r_origin)
a = ostr('hello %s world', origin=100)
a
(a % 'good').origin
b = 'hello %s world'
c = ostr('bad', origin=10)
(b % c).origin
```
#### String methods that do not change origin
```
class ostr(ostr):
def swapcase(self):
return self.create(str(self).swapcase(), self.origin)
def upper(self):
return self.create(str(self).upper(), self.origin)
def lower(self):
return self.create(str(self).lower(), self.origin)
def capitalize(self):
return self.create(str(self).capitalize(), self.origin)
def title(self):
return self.create(str(self).title(), self.origin)
a = ostr('aa', origin=100).upper()
a, a.origin
```
#### General wrappers
These are not strictly needed for operation, but can be useful for tracing.
```
def make_basic_str_wrapper(fun): # type: ignore
def proxy(*args, **kwargs):
res = fun(*args, **kwargs)
return res
return proxy
import inspect
import types
def informationflow_init_2():
ostr_members = [name for name, fn in inspect.getmembers(ostr, callable)
if isinstance(fn, types.FunctionType) and fn.__qualname__.startswith('ostr')]
for name, fn in inspect.getmembers(str, callable):
if name not in set(['__class__', '__new__', '__str__', '__init__',
'__repr__', '__getattribute__']) | set(ostr_members):
setattr(ostr, name, make_basic_str_wrapper(fn))
informationflow_init_2()
INITIALIZER_LIST.append(informationflow_init_2)
```
#### Methods yet to be translated
These methods generate strings from other strings. However, we do not have the right implementations for any of these. Hence these are marked as dangerous until we can generate the right translations.
```
def make_str_abort_wrapper(fun):
def proxy(*args, **kwargs):
raise ostr.TaintException(
'%s Not implemented in `ostr`' %
fun.__name__)
return proxy
def informationflow_init_3():
for name, fn in inspect.getmembers(str, callable):
# Omitted 'splitlines' as this is needed for formatting output in
# IPython/Jupyter
if name in ['__format__', 'format_map', 'format',
'__mul__', '__rmul__', 'center', 'zfill', 'decode', 'encode']:
setattr(ostr, name, make_str_abort_wrapper(fn))
informationflow_init_3()
INITIALIZER_LIST.append(informationflow_init_3)
```
While generating proxy wrappers for string operations can handle most common cases of transmission of information flow, some of the operations involving strings can not be overridden. For example, consider the following.
### End of Excursion
### Checking Origins
With all this implemented, we now have full-fledged `ostr` strings where we can easily check the origin of each and every character.
To check whether a string originates from another string, we can convert the origin to a set and resort to standard set operations:
```
s = ostr("hello", origin=100)
s[1]
s[1].origin
set(s[1].origin) <= set(s.origin)
t = ostr("world", origin=200)
set(s.origin) <= set(t.origin)
u = s + t + "!"
u.origin
ostr.UNKNOWN_ORIGIN in u.origin
```
### Privacy Leaks Revisited
Let us apply it to see whether we can come up with a satisfactory solution for checking the `heartbeat()` function against information leakage.
```
SECRET_ORIGIN = 1000
```
We define a "secret" that must not leak out:
```
secret = ostr('<again, some super-secret input>', origin=SECRET_ORIGIN)
```
Each and every character in `secret` has an origin starting with `SECRET_ORIGIN`:
```
print(secret.origin)
```
If we now invoke `heartbeat()` with a given string, the origin of the reply should all be `UNKNOWN_ORIGIN` (from the input), and none of the characters should have a `SECRET_ORIGIN`.
```
hello_s = heartbeat('hello', 5, memory=secret)
hello_s
assert isinstance(hello_s, ostr)
print(hello_s.origin)
```
We can verify that the secret did not leak out by formulating appropriate assertions:
```
assert hello_s.origin == [ostr.UNKNOWN_ORIGIN] * len(hello_s)
assert all(origin == ostr.UNKNOWN_ORIGIN for origin in hello_s.origin)
assert not any(origin >= SECRET_ORIGIN for origin in hello_s.origin)
```
All assertions pass, again confirming that no secret leaked out.
Let us now go and exploit `heartbeat()` to reveal its secrets. As `heartbeat()` is unchanged, it is as vulnerable as it was:
```
hello_s = heartbeat('hello', 32, memory=secret)
hello_s
```
Now, however, the reply _does_ contain secret information:
```
assert isinstance(hello_s, ostr)
print(hello_s.origin)
with ExpectError():
assert hello_s.origin == [ostr.UNKNOWN_ORIGIN] * len(hello_s)
with ExpectError():
assert all(origin == ostr.UNKNOWN_ORIGIN for origin in hello_s.origin)
with ExpectError():
assert not any(origin >= SECRET_ORIGIN for origin in hello_s.origin)
```
We can now integrate these assertions into the `heartbeat()` function, causing it to fail before leaking information. Additionally (or alternatively?), we can also rewrite our output functions not to give out any secret information. We will leave these two exercises for the reader.
## Taint-Directed Fuzzing
The previous _Taint Aware Fuzzing_ was a bit unsatisfactory in that we could not focus on the specific parts of the grammar that led to dangerous operations. We fix that with _taint directed fuzzing_ using `TrackingDB`.
The idea here is to track the origins of each character that reaches `eval`. Then, track it back to the grammar nodes that generated it, and increase the probability of using those nodes again.
### TrackingDB
The `TrackingDB` is similar to `TaintedDB`. The difference is that, if we find that the execution has reached the `my_eval`, we simply raise the `Tainted`.
```
class TrackingDB(TaintedDB):
def my_eval(self, statement, g, l):
if statement.origin:
raise Tainted(statement)
try:
return eval(statement, g, l)
except:
raise SQLException('Invalid SQL (%s)' % repr(statement))
```
Next, we need a specially crafted fuzzer that preserves the taints.
### TaintedGrammarFuzzer
We define a `TaintedGrammarFuzzer` class that ensures that the taints propagate to the derivation tree. This is similar to the `GrammarFuzzer` from the [chapter on grammar fuzzers](GrammarFuzzer.ipynb) except that the origins and taints are preserved.
```
import random
from GrammarFuzzer import GrammarFuzzer
from Parser import canonical
class TaintedGrammarFuzzer(GrammarFuzzer):
def __init__(self,
grammar,
start_symbol=START_SYMBOL,
expansion_switch=1,
log=False):
self.tainted_start_symbol = ostr(
start_symbol, origin=[1] * len(start_symbol))
self.expansion_switch = expansion_switch
self.log = log
self.grammar = grammar
self.c_grammar = canonical(grammar)
self.init_tainted_grammar()
def expansion_cost(self, expansion, seen=set()):
symbols = [e for e in expansion if e in self.c_grammar]
if len(symbols) == 0:
return 1
if any(s in seen for s in symbols):
return float('inf')
return sum(self.symbol_cost(s, seen) for s in symbols) + 1
def fuzz_tree(self):
tree = (self.tainted_start_symbol, [])
nt_leaves = [tree]
expansion_trials = 0
while nt_leaves:
idx = random.randint(0, len(nt_leaves) - 1)
key, children = nt_leaves[idx]
expansions = self.ct_grammar[key]
if expansion_trials < self.expansion_switch:
expansion = random.choice(expansions)
else:
costs = [self.expansion_cost(e) for e in expansions]
m = min(costs)
all_min = [i for i, c in enumerate(costs) if c == m]
expansion = expansions[random.choice(all_min)]
new_leaves = [(token, []) for token in expansion]
new_nt_leaves = [e for e in new_leaves if e[0] in self.ct_grammar]
children[:] = new_leaves
nt_leaves[idx:idx + 1] = new_nt_leaves
if self.log:
print("%-40s" % (key + " -> " + str(expansion)))
expansion_trials += 1
return tree
def fuzz(self):
self.derivation_tree = self.fuzz_tree()
return self.tree_to_string(self.derivation_tree)
```
We use a specially prepared tainted grammar for fuzzing. We mark each individual definition, each individual rule, and each individual token with a separate origin (we chose a token boundary of 10 here, after inspecting the grammar). This allows us to track exactly which parts of the grammar were involved in the operations we are interested in.
```
class TaintedGrammarFuzzer(TaintedGrammarFuzzer):
def init_tainted_grammar(self):
key_increment, alt_increment, token_increment = 1000, 100, 10
key_origin = key_increment
self.ct_grammar = {}
for key, val in self.c_grammar.items():
key_origin += key_increment
os = []
for v in val:
ts = []
key_origin += alt_increment
for t in v:
nt = ostr(t, origin=key_origin)
key_origin += token_increment
ts.append(nt)
os.append(ts)
self.ct_grammar[key] = os
# a use tracking grammar
self.ctp_grammar = {}
for key, val in self.ct_grammar.items():
self.ctp_grammar[key] = [(v, dict(use=0)) for v in val]
```
As before, we initialize the `TrackingDB`
```
trdb = TrackingDB(db.db)
```
Finally, we need to ensure that the taints are preserved, when the tree is converted back to a string. For this, we define the `tainted_tree_to_string()`
```
class TaintedGrammarFuzzer(TaintedGrammarFuzzer):
def tree_to_string(self, tree):
symbol, children, *_ = tree
e = ostr('')
if children:
return e.join([self.tree_to_string(c) for c in children])
else:
return e if symbol in self.c_grammar else symbol
```
We define `update_grammar()` that accepts a set of origins that reached the dangerous operations and the derivation tree of the original string used for fuzzing to update the enhanced grammar.
```
class TaintedGrammarFuzzer(TaintedGrammarFuzzer):
def update_grammar(self, origin, dtree):
def update_tree(dtree, origin):
key, children = dtree
if children:
updated_children = [update_tree(c, origin) for c in children]
corigin = set.union(
*[o for (key, children, o) in updated_children])
corigin = corigin.union(set(key.origin))
return (key, children, corigin)
else:
my_origin = set(key.origin).intersection(origin)
return (key, [], my_origin)
key, children, oset = update_tree(dtree, set(origin))
for key, alts in self.ctp_grammar.items():
for alt, o in alts:
alt_origins = set([i for token in alt for i in token.origin])
if alt_origins.intersection(oset):
o['use'] += 1
```
With these, we are now ready to fuzz.
```
def tree_type(tree):
key, children = tree
return (type(key), key, [tree_type(c) for c in children])
tgf = TaintedGrammarFuzzer(INVENTORY_GRAMMAR_F)
x = None
for _ in range(10):
qtree = tgf.fuzz_tree()
query = tgf.tree_to_string(qtree)
assert isinstance(query, ostr)
try:
print(repr(query))
res = trdb.sql(query)
print(repr(res))
except SQLException as e:
print(e)
except Tainted as e:
print(e)
origin = e.args[0].origin
tgf.update_grammar(origin, qtree)
except:
traceback.print_exc()
break
print()
```
We can now inspect our enhanced grammar to see how many times each rule was used.
```
tgf.ctp_grammar
```
From here, the idea is to focus on the rules that reached dangerous operations more often, and increase the probability of the values of that kind.
### The Limits of Taint Tracking
While our framework can detect information leakage, it is by no means perfect. There are several ways in which taints can get lost and information thus may still leak out.
#### Conversions
We only track taints and origins through _strings_ and _characters_. If we convert these to numbers (or other data), the information is lost.
As an example, consider this function, converting individual characters to numbers and back:
```
def strip_all_info(s):
t = ""
for c in s:
t += chr(ord(c))
return t
othello = ostr("Secret")
othello
othello.origin # type: ignore
```
The taints and origins will not propagate through the number conversion:
```
thello_stripped = strip_all_info(thello)
thello_stripped
with ExpectError():
thello_stripped.origin
```
This issue could be addressed by extending numbers with taints and origins, just as we did for strings. At some point, however, this will still break down, because as soon as an internal C function in the Python library is reached, the taint will not propagate into and across the C function. (Unless one starts implementing dynamic taints for these, that is.)
#### Internal C libraries
As we mentioned before, calls to _internal_ C libraries do not propagate taints. For example, while the following preserves the taints,
```
hello = ostr('hello', origin=100)
world = ostr('world', origin=200)
(hello + ' ' + world).origin
```
a call to a `join` that should be equivalent will fail.
```
with ExpectError():
''.join([hello, ' ', world]).origin # type: ignore
```
#### Implicit Information Flow
Even if one could taint all data in a program, there still would be means to break information flow – notably by turning explicit flow into _implicit_ flow, or data flow into _control flow_. Here is an example:
```
def strip_all_info_again(s):
t = ""
for c in s:
if c == 'a':
t += 'a'
elif c == 'b':
t += 'b'
elif c == 'c':
t += 'c'
...
```
With such a function, there is no explicit data flow between the characters in `s` and the characters in `t`; yet, the strings would be identical. This problem frequently occurs in programs that process and manipulate external input.
#### Enforcing Tainting
Both, conversions and implicit information flow are one of several possibilities how taint and origin information get lost. To address the problem, the best solution is to _always assume the worst from untainted strings_:
* As it comes to trust, an untainted string should be treated as _possibly untrusted_, and hence not relied upon unless sanitized.
* As it comes to privacy, an untainted string should be treated as _possibly secret_, and hence not leaked out.
As a consequence, your program should always have two kinds of taints: one for explicitly trusted (or secret) and one for explicitly untrusted (or non-secret). If a taint gets lost along the way, you will may have to restore it from its sources – not unlike the string methods discussed above. The benefit is a trusted application, in which each and every information flow can be checked at runtime, with violations quickly discovered through automated tests.
## Synopsis
This chapter provides two wrappers to Python _strings_ that allow one to track various properties. These include information on the security properties of the input, and information on originating indexes of the input string.
### Tracking String Taints
`tstr` objects are replacements for Python strings that allows to track and check _taints_ – that is, information on from where a string originated. For instance, one can mark strings that originate from third party input with a taint of "LOW", meaning that they have a low security level. The taint is passed in the constructor of a `tstr` object:
```
thello = tstr('hello', taint='LOW')
```
A `tstr` object is fully compatible with original Python strings. For instance, we can index it and access substrings:
```
thello[:4]
```
However, the `tstr` object also stores the taint, which can be accessed using the `taint` attribute:
```
thello.taint
```
The neat thing about taints is that they propagate to all strings derived from the original tainted string.
Indeed, any operation from a `tstr` string that results in a string fragment produces another `tstr` object that includes the original taint. For example:
```
thello[1:2].taint # type: ignore
```
`tstr` objects duplicate most `str` methods, as indicated in the class diagram:
```
# ignore
from ClassDiagram import display_class_hierarchy
display_class_hierarchy(tstr)
```
### Tracking Character Origins
`ostr` objects extend `tstr` objects by not only tracking a taint, but also the originating _indexes_ from the input string, This allows you to exactly track where individual characters came from. Assume you have a long string, which at index 100 contains the password `"joshua1234"`. Then you can save this origin information using an `ostr` as follows:
```
secret = ostr("joshua1234", origin=100, taint='SECRET')
```
The `origin` attribute of an `ostr` provides access to a list of indexes:
```
secret.origin
secret.taint
```
`ostr` objects are compatible with Python strings, except that string operations return `ostr` objects (together with the saved origin an index information). An index of `-1` indicates that the corresponding character has no origin as supplied to the `ostr()` constructor:
```
secret_substr = (secret[0:4] + "-" + secret[6:])
secret_substr.taint
secret_substr.origin
```
`ostr` objects duplicate most `str` methods, as indicated in the class diagram:
```
# ignore
display_class_hierarchy(ostr)
```
## Lessons Learned
* String-based and character-based taints allow to dynamically track the information flow from input to the internals of a system and back to the output.
* Checking taints allows to discover untrusted inputs and information leakage at runtime.
* Data conversions and implicit data flow may strip taint information; the resulting untainted strings should be treated as having the worst possible taint.
* Taints can be used in conjunction with fuzzing to provide a more robust indication of incorrect behavior than to simply rely on program crashes.
## Next Steps
An even better alternative to our taint-directed fuzzing is to make use of _symbolic_ techniques that take the semantics of the program under test into account. The chapter on [flow fuzzing](FlowFuzzer.ipynb) introduces these symbolic techniques for the purpose of exploring information flows; the subsequent chapter on [symbolic fuzzing](SymbolicFuzzer.ipynb) then shows how to make full-fledged use of symbolic execution for covering code. Similarly, [search based fuzzing](SearchBasedFuzzer.ipynb) can often provide a cheaper exploration strategy.
## Background
Taint analysis on Python using a library approach as we implemented in this chapter was discussed by Conti et al. \cite{Conti2010}.
## Exercises
### Exercise 1: Tainted Numbers
Introduce a class `tint` (for tainted integer) that, like `tstr`, has a taint attribute that gets passed on from `tint` to `tint`.
#### Part 1: Creation
Implement the `tint` class such that taints are set:
```python
x = tint(42, taint='SECRET')
assert x.taint == 'SECRET'
```
**Solution.** This is pretty straightforward, as we can apply the same scheme as for `tstr`:
```
class tint(int):
def __new__(cls, value, *args, **kw):
return int.__new__(cls, value)
def __init__(self, value, taint=None, **kwargs):
self.taint = taint
x = tint(42, taint='SECRET')
assert x.taint == 'SECRET'
```
#### Part 2: Arithmetic expressions
Ensure that taints get passed along arithmetic expressions; support addition, subtraction, multiplication, and division operators.
```python
y = x + 1
assert y.taint == 'SECRET'
```
**Solution.** As with `tstr`, we implement a `create()` method and a convenience function to quickly define all arithmetic operations:
```
class tint(tint):
def create(self, n):
# print("New tint from", n)
return tint(n, taint=self.taint)
```
The `make_int_wrapper()` function creates a wrapper around an existing `int` method which attaches the taint to the result of the method:
```
def make_int_wrapper(fun):
def proxy(self, *args, **kwargs):
res = fun(self, *args, **kwargs)
# print(fun, args, kwargs, "=", repr(res))
return self.create(res)
return proxy
```
We do this for all arithmetic operators:
```
for name in ['__add__', '__radd__', '__mul__', '__rmul__', '__sub__',
'__floordiv__', '__truediv__']:
fun = getattr(int, name)
setattr(tint, name, make_int_wrapper(fun))
x = tint(42, taint='SECRET')
y = x + 1
y.taint # type: ignore
```
#### Part 3: Passing taints from integers to strings
Converting a tainted integer into a string (using `repr()`) should yield a tainted string:
```python
x_s = repr(x)
assert x_s.taint == 'SECRET'
```
**Solution.** We define the string conversion functions such that they return a tainted string (`tstr`):
```
class tint(tint):
def __repr__(self) -> tstr:
s = int.__repr__(self)
return tstr(s, taint=self.taint)
class tint(tint):
def __str__(self) -> tstr:
return tstr(int.__str__(self), taint=self.taint)
x = tint(42, taint='SECRET')
x_s = repr(x)
assert isinstance(x_s, tstr)
assert x_s.taint == 'SECRET'
```
#### Part 4: Passing taints from strings to integers
Converting a tainted object (with a `taint` attribute) to an integer should pass that taint:
```python
password = tstr('1234', taint='NOT_EXACTLY_SECRET')
x = tint(password)
assert x == 1234
assert x.taint == 'NOT_EXACTLY_SECRET'
```
**Solution.** This can be done by having the `__init__()` constructor check for a `taint` attibute:
```
class tint(tint):
def __init__(self, value, taint=None, **kwargs):
if taint is not None:
self.taint = taint
else:
self.taint = getattr(value, 'taint', None)
password = tstr('1234', taint='NOT_EXACTLY_SECRET')
x = tint(password)
assert x == 1234
assert x.taint == 'NOT_EXACTLY_SECRET'
```
### Exercise 2: Information Flow Testing
Generate tests that ensure a _maximum_ of information flow, propagating specific taints as much as possible. Implement an appropriate fitness function for [search-based testing](SearchBasedFuzzer.ipynb) and let the search-based fuzzer search for solutions.
**Solution.** This will become a section on its own; as of now, it is an exercise for the reader.
| github_jupyter |
# Let's find boxes!
I will use FindBoxes app
This should works on Mac OS X, Ubuntu 16.04 and Windows 10.
Make sure to install at least:
- dotnet core 2.0
## Imports
```
import cv2
import math
import os
import numpy as np
import statistics
import shutil
import time
import subprocess
from IPython.display import Image
```
## Compile FindBoxes program
We don't need to generate a build.
But, it will be slower if we process many images.
You may want to do this when you deploy your app in production.
```
def copy_dir(src, dest):
try:
shutil.rmtree(dest, ignore_errors=True)
shutil.copytree(src, dest)
except Exception as e:
print('Copy directiory error:' + str(e))
def setup_findboxes():
# Get the code
source_dir = '..' + os.path.sep + '..' + os.path.sep + 'CSharp' + os.path.sep + 'FindBoxes' + os.path.sep
destination_dir = 'findboxesapp'
copy_dir(source_dir, destination_dir)
# Restore package
p = subprocess.Popen('cd ' + destination_dir + " && dotnet restore",
shell=True, stdout=subprocess.PIPE, stderr=subprocess.STDOUT)
for line in p.stdout.readlines():
print(line.decode("utf-8"))
retval = p.wait()
# Build package
p = subprocess.Popen('cd ' + destination_dir + " && dotnet build --configuration Release",
shell=True, stdout=subprocess.PIPE, stderr=subprocess.STDOUT)
for line in p.stdout.readlines():
print(line.decode("utf-8"))
retval = p.wait()
# Temp folder for processing.
temp_dir = 'temp'
if not os.path.exists(temp_dir):
os.makedirs(temp_dir)
# Ensure the program setup.
# We don't have to build the program.
# But, if we do it, it will be faster to process many images.
setup_findboxes()
```
## Call findboxes app
**TODO:** parse results and return it. (Still need some work on c# app.)
```
def findboxes(img_path):
dll_program = 'findboxesapp' + os.path.sep + 'bin' + os.path.sep \
+ 'Release' + os.path.sep + 'netcoreapp2.0' + os.path.sep + 'FindBoxes.dll'
temp_file_output = 'temp/result.json'
p = subprocess.Popen("dotnet " + dll_program + " --input " + img_path + " --output " + 'std',
shell=True, stdout=subprocess.PIPE, stderr=subprocess.STDOUT)
for line in p.stdout.readlines():
print(line.decode("utf-8"))
retval = p.wait()
# Call findBoxes program (python -> dotnet core)
start_time = time.time()
# Pre-processing has already been done with OpenCV with the example image.
# Normally, you would use adaptive threshold, invert pixels (Bitwise not), dilate (Cross 2x2), etc.
findboxes('files/form9.jpg')
end_time = time.time()
duration_ms = int((end_time - start_time) * 1000)
print('Total duration: ' + str(duration_ms) + " ms")
```
| github_jupyter |
```
# Required to access the database
import os
os.environ["DJANGO_ALLOW_ASYNC_UNSAFE"] = "true"
import sys
import numpy
numpy.set_printoptions(threshold=sys.maxsize)
# Data analysis tools
import pandas as pd
import numpy as np
import seaborn as sns
# Models available in our application
from datasets.models import RawFlower, RawUNM, RawDAR
from django.contrib.auth.models import User
from datasets.models import RawNEU
import statsmodels.api as sm
import statsmodels.formula.api as smf
import statsmodels
!pip install lxml
from api import adapters
from api import analysis
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
def crude_reg(df_merged, x_feature, y_feature, adjust_dilution, use_covars):
## adjust dilution
if adjust_dilution == True:
df_merged[x_feature] = df_merged[x_feature] / df_merged['UDR']
if use_covars:
data = df_merged
data.drop(['CohortType'], inplace = True, axis = 1)
else:
data = df_merged[[x_feature,y_feature]]
## problem - if we are using z_score to predict might be an issue
data['intercept'] = 1
#X and Y features TODO: clean up
X = data[[x for x in data.columns if x !=y_feature and x!= 'PIN_Patient']]
#print(X.info())
Y = data[y_feature]
X[x_feature]= np.log(X[x_feature])
if df_merged.shape[0] > 2:
reg = sm.OLS(Y, X).fit()
ret = reg.summary()
else:
ret = 'error'
# model string
fit_string = y_feature + '~'
for x in X.columns:
if x == x_feature:
fit_string += ' + log(' + str(x) +')'
else:
fit_string += ' + ' + str(x)
#htmls = header + ret.tables[0].as_html() + ret.tables[1].as_html()
df = pd.read_html(ret.tables[1].as_html(),header=0,index_col=0)[0]
return df
def crude_logreg(df_merged, x_feature, y_feature, adjust_dilution, use_covars):
## adjust dilution
if adjust_dilution == True:
df_merged[x_feature] = df_merged[x_feature] / df_merged['UDR']
if use_covars:
data = df_merged
data.drop(['CohortType'], inplace = True, axis = 1)
else:
data = df_merged[[x_feature,y_feature]]
## problem - if we are using z_score to predict might be an issue
data['intercept'] = 1
#X and Y features TODO: clean up
X = data[[x for x in data.columns if x !=y_feature and x!= 'PIN_Patient']]
Y = data[y_feature]
X[x_feature]= np.log(X[x_feature])
# fit the model
print('columns going into logreg')
print(X.columns)
if df_merged.shape[0] > 1:
log_reg = sm.Logit(Y, X).fit()
ret = log_reg.summary()
else:
ret = 'error'
# model string
fit_string = y_feature + '~'
for x in X.columns:
if x == x_feature:
fit_string += ' + log(' + str(x) +')'
else:
fit_string += ' + ' + str(x)
df = pd.read_html(ret.tables[1].as_html(),header=0,index_col=0)[0]
return df
def dummy_code(df, covars_cat, contin):
coded_covars = []
orig_shape = df.shape[0]
for var in covars_cat:
df[var] = pd.Categorical(df[var])
dummies_df = pd.get_dummies(df[var], prefix = var, drop_first=True)
coded_covars = coded_covars + [ x for x in dummies_df.columns.tolist()]
df = pd.concat([df, dummies_df], axis = 1)
df.drop([var], inplace = True, axis = 1)
assert df.shape[0] == orig_shape
#print(coded_covars + contin)
return df[coded_covars + contin]
from api import dilutionproc
def printsummary(df):
x = 1
# spearate the data into cat and continuous summary:
# Get the data
## Model 1: Restricted to participants with no fish/seafood consumption.
## Get NEU data with no fish
df_NEU = adapters.neu.get_dataframe_orig()
df_NEU = df_NEU[df_NEU['TimePeriod']==2] # Visit 2
df_NEU_covars = adapters.neu.get_dataframe_covars()
df_NEU = df_NEU_covars.merge(df_NEU, on = ['PIN_Patient','CohortType','TimePeriod']) #Merge the covariates
df_NEU = df_NEU[(df_NEU['fish_pu_v2'] == 0) & (df_NEU['fish'] == 0)] #No fish consumption
## Get DAR data with no fish
df_DAR = adapters.dar.get_dataframe_nofish()
## Get UNM data with no fis
df_UNM = adapters.unm.get_dataframe_orig()
#df_UNM = df_UNM[df_UNM['fish']==0]
df_UNM_covars = adapters.unm.get_dataframe_covars()
df_UNM = df_UNM_covars.merge(df_UNM, on = ['PIN_Patient','CohortType','TimePeriod']) #Merge the covariates
df_NEU = df_NEU.replace(-9,np.nan).replace('-9', np.nan)
#df_ALL = analysis.merge3CohortFrames(df_UNM,df_NEU,df_DAR)
df_ALL = df_NEU
frames_for_adjust = [
('NEU', df_NEU)
]
frames_for_analysis = [
('NEU', df_NEU),
('ALL', df_ALL)
]
for name, df in frames_for_analysis:
print('Data Stats')
print(name)
print(df.shape)
##Run the adjustment
for name, df_coh in frames_for_adjust:
print('Working on ', name)
keep_adj = []
#variables for fitting procedure
x_feature = 'UTAS'
cat_vars = ['babySex','smoking','education','race']
contin_vars = ['PIN_Patient','BMI','UTAS']
# dummy code
df_coh_coded_model = dummy_code(df_coh, cat_vars, contin_vars)
## variables for addjustment procedure
adjust_cat_vars = ['babySex','smoking','education','race']
adjust_contin_vars = ['PIN_Patient','CohortType','BMI', 'ga_collection','birth_year','age']
#add proper variable depending on cohort
if name == 'NEU':
adjust_contin_vars= adjust_contin_vars + ['SPECIFICGRAVITY_V2']
if name == 'UNM':
adjust_contin_vars = adjust_contin_vars + ['cratininemgl']
if name == 'DAR':
adjust_contin_vars = adjust_contin_vars + ['darvar']
## adjustment procedure
if name in ['NEU', 'UNM', 'NEU']:
#dummy code
df_coh_coded_adjust_model = dummy_code(df_coh, adjust_cat_vars, adjust_contin_vars)
d_test = df_coh_coded_adjust_model.dropna()
dil_adj = dilutionproc.predict_dilution(d_test, 'NEU')
fin = df_coh_coded_model.merge(dil_adj[['PIN_Patient','UDR']], on = ['PIN_Patient'])
adjs = dil_adj[['PIN_Patient','UDR']]
adjs.loc[:,'CohortType'] = name
keep_adj.append(adjs)
print('Done')
cohort_adjustmets = pd.concat(keep_adj)
cohort_adjustmets
'''
('UNM', df_UNM),
('DAR', df_DAR),
('NEUUNM', df_NEUUNM),
('NEUDAR', df_NEUDAR),
('UNMDAR', df_UNMDAR),
('UNMDARNEU', df_merged_3),
]
'''
#d_test = df_NEU[['PIN_Patient','CohortType','race', 'education','babySex','BMI', 'ga_collection','birth_year','age','SPECIFICGRAVITY_V2']]
#all_vars = covars + [x_feature]
Y_features_continuous = ['Outcome_weeks','birthWt', 'headCirc', 'birthLen']
Y_features_binary = ['LGA','SGA','Outcome']
outputs_conf = []
outputs_crude = []
for outcome in Y_features_binary + Y_features_continuous:
for name, df_coh in frames_for_analysis:
print('Working on ', name)
#variables for fitting procedure
x_feature = 'UTAS'
cat_vars = ['babySex','smoking','education','race']
contin_vars = ['PIN_Patient','BMI','UTAS'] + [outcome]
# dummy code
df_coh_coded_model = dummy_code(df_coh, cat_vars, contin_vars)
## variables for addjustment procedure
adjust_cat_vars = ['babySex','smoking','education','race']
adjust_contin_vars = ['PIN_Patient','CohortType','BMI', 'ga_collection','birth_year','age']
if name in ['NEU', 'UNM', 'NEU']:
#dummy code
print("go")
fin = df_coh_coded_model.merge(cohort_adjustmets, on = ['PIN_Patient'])
print(fin.columns)
#sdf
if name in ['ALL']:
x = 1
if len(keep_adj) == 1: df_adj_all = pd.concat(keep_adj)
fin = df_coh_coded_model.merge(df_adj_all, on = ['PIN_Patient'])
# run models:
if outcome in Y_features_continuous:
fin = fin.dropna()
output = crude_reg(fin, x_feature, outcome, False, True)
output['y'] = outcome
output['crude'] = False
output['model'] = 'OLS'
outputs_conf.append(output)
output_crude = crude_reg(fin, x_feature, outcome, False, False)
output_crude['y'] = outcome
output_crude['crude'] = True
output_crude['model'] = 'OLS'
outputs_conf.append(output_crude)
if outcome in Y_features_binary:
fin = fin.dropna()
output = crude_logreg(fin, x_feature, outcome, False, True)
output.columns = ['coef', 'std err', 't', 'P>|t|', '[0.025','0.975]']
output['y'] = outcome
output['crude'] = False
output['model'] = 'Logit'
outputs_conf.append(output)
output_crude = crude_logreg(fin, x_feature, outcome, False, False)
output_crude.columns = ['coef', 'std err', 't', 'P>|t|', '[0.025','0.975]']
output_crude['y'] = outcome
output_crude['crude'] = True
output_crude['model'] = 'Logit'
outputs_conf.append(output_crude)
# set output paths for results:
#utput_path_model1_adj = '/usr/src/app/mediafiles/analysisresults/model1adj/'
#utput_path_model1_noadj = '/usr/src/app/mediafiles/analysisresults/model1noadj/'
#ry:
# os.mkdir(output_path_model1_adj)
# os.mkdir(output_path_model1_noadj)
#xcept:
# print('Exists')
# start analysis
pd.concat(outputs_conf)
for name, frame in frames_for_analysis:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continuous:
output = crude_reg(frame, x_feature, y_feature)
#ext_writing(name, frame, x_feature, y_feature, all_vars, output_path_model1_adj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature), 'Linear Regression')
print(output)
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature)
#ext_writing(name, frame, x_feature, y_feature, all_vars, output_path_model1_adj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
print(output)
from api import dilutionproc
d_test = df_NEU[['PIN_Patient','CohortType','race', 'education','babySex','BMI', 'ga_collection','birth_year','age','SPECIFICGRAVITY_V2']]
d_test = d_test.dropna()
dilutionproc.predict_dilution(d_test, 'NEU')
#Model 2: Restricted to participants with arsenic speciation data.
## Get data with fish
df_UNM = adapters.unm.get_dataframe()
df_DAR = adapters.dar.get_dataframe_pred()
## merge data frames
df_UNMDAR = merge2CohortFrames(df_UNM,df_DAR)
frames_for_analysis = [
('UNM', df_UNM),
('DAR', df_DAR),
('UNMDAR', df_UNMDAR)
]
for name, df in frames_for_analysis:
print('Data Stats')
print(name)
print(df.shape)
x_feature = 'UTAS'
covars = 'babySex|BMI|parity|smoking|education'
all_vars = covars.split('|') + [x_feature]
Y_features_continous = ['Outcome_weeks','birthWt', 'headCirc', 'birthLen']
Y_features_binary = ['LGA','SGA','Outcome']
output_path_model2_adj = '/usr/src/app/mediafiles/analysisresults/model2adj/'
output_path_model2_noadj = '/usr/src/app/mediafiles/analysisresults/model2noadj/'
#output_path = '../mediafiles/analysisresults/'
try:
os.mkdir(output_path_model2_adj)
os.mkdir(output_path_model2_noadj)
except:
print('Exists')
for name, frame in frames_for_analysis:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output= crude_reg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model2_adj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model2_adj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
#without adjustment
for name, frame in frames_for_analysis:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output = crude_reg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model2_noadj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model2_noadj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
#Model 3: Restricted to arsenic speciation data with AsB ≤1 µg/L.
x_feature = 'UTAS'
covars = 'babySex|BMI|parity|smoking|education'
all_vars = covars.split('|') + [x_feature]
Y_features_continous = ['Outcome_weeks','birthWt', 'headCirc', 'birthLen']
Y_features_binary = ['LGA','SGA','Outcome']
## Number of Participants
output_path_model3_adj = '/usr/src/app/mediafiles/analysisresults/model3adj/'
output_path_model3_noadj = '/usr/src/app/mediafiles/analysisresults/model3noadj/'
#output_path = '../mediafiles/analysisresults/'
try:
os.mkdir(output_path_model3_adj)
os.mkdir(output_path_model3_noadj)
except:
print('Exists')
# remove the AsB <= 1
df_UNM = df_UNM[df_UNM['UASB'] <= 1]
df_DAR = df_DAR[df_DAR['UASB'] <= 1]
df_UNMDAR_UASB = df_UNMDAR[df_UNMDAR['UASB'] <= 1]
frames_for_analysis3 = [
('UNM', df_UNM),
('DAR', df_DAR),
('UNMDAR', df_UNMDAR)
]
for name, frame in frames_for_analysis3:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output = crude_reg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model3_adj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model3_adj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
#no adj
for name, frame in frames_for_analysis3:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output = crude_reg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model3_noadj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model3_noadj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
#Model 4: Sensitivity analysis
x_feature = 'UTAS'
covars = 'babySex|BMI|parity|smoking|education'
all_vars = covars.split('|') + [x_feature]
Y_features_continous = ['Outcome_weeks','birthWt', 'headCirc', 'birthLen']
Y_features_binary = ['LGA','SGA','Outcome']
## Number of Participants
output_path_model4_adj = '/usr/src/app/mediafiles/analysisresults/model4adj/'
output_path_model4_noadj = '/usr/src/app/mediafiles/analysisresults/model4noadj/'
#output_path = '../mediafiles/analysisresults/'
try:
os.mkdir(output_path_model4_adj)
os.mkdir(output_path_model4_noadj)
except:
print('Exists')
## Get data all
df_NEU = adapters.neu.get_dataframe()
df_UNM = adapters.unm.get_dataframe()
df_DAR = adapters.dar.get_dataframe_pred()
## merge data frames
df_NEUUNM = merge2CohortFrames(df_NEU,df_UNM)
df_NEUDAR = merge2CohortFrames(df_NEU,df_DAR)
df_UNMDAR = merge2CohortFrames(df_UNM,df_DAR)
df_merged_3 = merge3CohortFrames(df_NEU,df_UNM,df_DAR)
frames_for_analysis4 = [
('NEU', df_NEU),
('UNM', df_UNM),
('DAR', df_DAR),
('NEUUNM', df_NEUUNM),
('NEUDAR', df_NEUDAR),
('UNMDAR', df_UNMDAR),
('UNMDARNEU', df_merged_3),
]
for name, frame in frames_for_analysis4:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output = crude_reg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model4_adj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'True', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model4_adj, output, "logistic_reg{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
#no adj
for name, frame in frames_for_analysis3:
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
frame = frame[(frame['UTAS'] > 0) & (~frame['UTAS'].isna())]
print('Min: {} Max: {}'.format(frame['UTAS'].min(), frame['UTAS'].max()))
for y_feature in Y_features_continous:
output = crude_reg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model4_noadj, output, "linear_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Linear Regression')
for y_feature in Y_features_binary:
output = crude_logreg(frame, x_feature, y_feature, covars, 'False', 'csv', True)
text_writing(name, frame, x_feature, y_feature, all_vars, output_path_model4_noadj, output, "logistic_reg_{}_{}_log({}).txt".format(name, y_feature, x_feature),'Logistic Regression')
```
| github_jupyter |
# StyleGAN2 operations
```
# modules for notebook logging
!pip install IPython
!pip install ipywidgets
```
## Training
First, let's prepare the data. Ensure all your images have the same color channels (monochrome, RGB or RGBA).
If you work with patterns or shapes (rather than compostions), you can crop square fragments from bigger images (effectively multiplying their amount). For that, edit source and target paths below; `size` is fragment resolution, `step` is shift between the fragments. This will cut every source image into 512x512px fragments, overlapped with 256px shift by X and Y. Edit `size` and `overlap` according to your dataset.
```
src_dir = 'data/src'
data_dir = 'data/mydata'
size = 512
step = 256
%run src/util/multicrop.py --in_dir $src_dir --out_dir $data_dir --size $size --step $step
```
If you edit the images yourself (e.g. for non-square aspect ratios), ensure their correct size and put to the `data_dir`. For conditional model split the data by subfolders (`mydata/1`, `mydata/2`, ..) and add `--cond` option to the training command below.
```
data_dir = 'data/mydata'
```
Convert directory with images to TFRecords dataset (`mydata-512x512.tfr` file in `data` directory):
```
%run src/training/dataset_tool.py --data $data_dir
```
Now, we can train StyleGAN2 on the prepared dataset:
```
%run src/train.py --data $data_dir
```
> This will run training process, according to the options in `src/train.py` (check and explore those!!). If there was no TFRecords file from the previous step, it will be created at this point. Results (models and samples) are saved under `train` directory, similar to original Nvidia approach. There are two types of models saved: compact (containing only Gs network for inference) as `<dataset>-...pkl` (e.g. `mydata-512-0360.pkl`), and full (containing G/D/Gs networks for further training) as `snapshot-...pkl`.
> By default, the most powerful SG2 config (F) is used; if you face OOM issue, you may resort to `--config E`, requiring less memory (with poorer results, of course). For small datasets (100x images instead of 10000x) one should add `--d_aug` option to use [Differential Augmentation](https://github.com/mit-han-lab/data-efficient-gans) for more effective training.
> The length of the training is defined by `--kimg X` argument (training duration in thousands of images). Reasonable `kimg` value for full training from scratch is 5000-8000, while for finetuning in `--d_aug` mode 1000-2000 may be sufficient.
If the training process was interrupted, we can resume it from the last saved model as following:
*(replace `000-mydata-512-f` with existing training directory)*
```
%run src/train.py --data $data_dir --resume train/000-mydata-512-f
```
NB: In most cases it's much easier to use a "transfer learning" trick, rather than perform full training from the scratch. For that, we use existing well-trained model as a starter, and "finetune" (uptrain) it with our data. This works pretty well, even if our dataset is very different from the original model.
So here is a faster way to train our GAN (presuming we have `ffhq-512.pkl` model already):
```
%run src/train.py --data $data_dir --resume train/ffhq-512.pkl --d_aug --kimg 1000 --finetune
```
## Generation
Let's produce some imagery from the original cat model (download it from [here](https://nvlabs-fi-cdn.nvidia.com/stylegan2/networks/stylegan2-cat-config-f.pkl) and put to `models` directory).
More cool models can be found [here](https://github.com/justinpinkney/awesome-pretrained-stylegan2).
```
from IPython.display import Image, Video
model = 'models/stylegan2-cat-config-f' # without ".pkl" extension
model_pkl = model + '.pkl' # with ".pkl" extension
output = '_out/cats'
frames = '50-10'
```
Generate some animation to test the model:
```
%run src/_genSGAN2.py --model $model_pkl --out_dir $output --frames $frames
Image(filename = output + '/000000.jpg') # show first frame
```
> Here we loaded the model 'as is', and produced 50 frames in its natural resolution, interpolating between random latent `z` space keypoints, with a step of 10 frames between keypoints.
If you have `ffmpeg` installed, you can convert it into video:
```
out_sequence = output + '/%06d.jpg'
out_video = output + '.mp4'
!ffmpeg -y -v warning -i $out_sequence $out_video
Video(out_video)
```
>
Now let's generate custom animation. For that we omit model extension, so the script would load custom network, effectively enabling special features, e.g. arbitrary resolution (set by `--size` argument in `X-Y` format).
`--cubic` option changes linear interpolation to cubic for smoother animation (there is also `--gauss` option for additional smoothing).
```
%run src/_genSGAN2.py --model $model --out_dir $output --frames $frames --size 400-300 --cubic
Image(output+'/000000.jpg')
```
> **Run `ffmpeg` command above after each generation, if you want to check results in motion.**
Adding `--save_lat` option will save all traversed dlatent points (in `w` space) as Numpy array in `*.npy` file (useful for further curating). Set `--seed X` value to produce repeatable results.
Generate more various imagery:
```
%run src/_genSGAN2.py --model $model --out_dir $output --frames $frames --size 768-256 -n 3-1
Image(output+'/000000.jpg')
```
> Here we get animated composition of 3 independent frames, blended together horizontally (like the image in the repo header). Argument `--splitfine X` controls boundary fineness (0 = smoothest/default, higher => thinner).
Instead of frame splitting, we can load external mask from b/w image file (it also could be folder with file sequence):
```
%run src/_genSGAN2.py --model $model --out_dir $output --frames $frames --size 400-300 --latmask _in/mask.jpg
Image(output+'/000000.jpg')
```
>
`--digress X` adds some funky displacements with X strength (by tweaking initial constant layer).
`--trunc X` controls truncation psi parameter (0 = boring, 1+ = weird).
```
%run src/_genSGAN2.py --model $model --out_dir $output --frames $frames --digress 2 --trunc 0.5
Image(output+'/000000.jpg')
```
> Don't forget to check other options of `_genSGAN2.py` by `--help` argument.
### Latent space exploration
For these experiments download [FFHQ model](https://nvlabs-fi-cdn.nvidia.com/stylegan2/networks/stylegan2-ffhq-config-f.pkl) and save to `models`.
```
from IPython.display import Image, Video
model = 'models/stylegan2-ffhq-config-f' # without ".pkl" extension
model_pkl = model + '.pkl' # with ".pkl" extension
```
>
Project external images (aligned face portraits) from `_in/photo` onto StyleGAN2 model dlatent `w` space.
Results (found dlatent points as Numpy arrays in `*.npy` files, and video/still previews) are saved to `_out/proj` directory.
NB: first download [VGG model](https://drive.google.com/uc?id=1N2-m9qszOeVC9Tq77WxsLnuWwOedQiD2) and save it as `models/vgg/vgg16_zhang_perceptual.pkl`
```
%run src/project_latent.py --model $model_pkl --in_dir _in/photo --out_dir _out/proj
```
>
Generate animation between saved dlatent points:
```
dlat = 'dlats'
path_in = '_in/' + dlat
path_out = '_out/ffhq-' + dlat
out_sequence = path_out + '/%06d.jpg'
out_video = path_out + '.mp4'
%run src/_play_dlatents.py --model $model --dlatents $path_in --out_dir $path_out --fstep 10
Image(path_out+'/000000.jpg', width=512, height=512)
!ffmpeg -y -v warning -i $out_sequence $out_video
Video(out_video, width=512, height=512)
```
> This loads saved dlatent points from `_in/dlats` and produces smooth looped animation between them (with interpolation step of 50 frames). `dlats` may be a file or a directory with `*.npy` or `*.npz` files. To select only few frames from a sequence `somename.npy`, create text file with comma-delimited frame numbers and save it as `somename.txt` in the same directory (check given examples for FFHQ model).
Style-blending argument `--style_dlat blonde458.npy` would also load dlatent from `blonde458.npy` and apply it to higher network layers. `--cubic` smoothing and `--digress X` displacements are also applicable here:
```
%run src/_play_dlatents.py --model $model --dlatents $path_in --out_dir $path_out --fstep 10 --style_dlat _in/blonde458.npy --digress 2 --cubic
!ffmpeg -y -v warning -i $out_sequence $out_video
Video(out_video, width=512, height=512)
```
>
Generate animation by moving saved dlatent point `_in/blonde458.npy` along feature direction vectors from `_in/vectors_ffhq` (aging/smiling/etc) one by one: (check preview window!)
```
%run src/_play_vectors.py --model $model_pkl --base_lat _in/blonde458.npy --vector_dir _in/vectors_ffhq --out_dir _out/ffhq_looks
!ffmpeg -y -v warning -i _out/ffhq_looks/%06d.jpg _out/ffhq-vectors.mp4
Video('_out/ffhq-vectors.mp4', width=512, height=512)
```
> Such vectors can be discovered, for example, with the following methods:
> * https://github.com/genforce/sefa
> * https://github.com/harskish/ganspace
> * https://github.com/LoreGoetschalckx/GANalyze
## Tweaking models
NB: No real examples here! Just reference commands, try with your own files.
Strip G/D networks from a full model, leaving only Gs for inference. Resulting file is saved with `-Gs` suffix. It's recommended to add `-r` option to reconstruct the network, saving necessary arguments with it. Useful for foreign downloaded models.
```
%run src/model_convert.py --source snapshot-1024.pkl
```
Add or remove layers (from a trained model) to adjust its resolution for further finetuning. This will produce new model with 512px resolution, populating weights on the layers up to 256px from the source snapshot (the rest will be initialized randomly). It also can decrease resolution (say, make 512 from 1024). Note that this effectively changes the number of layers in the model.
This option works with complete (G/D/Gs) models only, since it's purposed for transfer-learning (the resulting model will contain either partially random weights, or wrong `ToRGB` params).
```
%run src/model_convert.py --source snapshot-256.pkl --res 1024
```
Change aspect ratio of a trained model by cropping or padding layers (keeping their count). Originally from @Aydao. This is experimental function with some voluntary logic, so use with care. This produces working non-square model. In case of basic aspect conversion (like 4x4 => 5x3), complete models (G/D/Gs) will be trainable for further finetuning.
```
%run src/model_convert.py --source snapshot-1024.pkl --res 1280-768
```
Add alpha channel to a trained model for further finetuning:
```
%run src/model_convert.py --source snapshot-1024.pkl --alpha
```
All above (adding/cropping/padding layers + alpha channel) can be done in one shot:
```
%run src/model_convert.py --source snapshot-256.pkl --res 1280-768 --alpha
```
Combine lower layers from one model with higher layers from another. `<res>` is resolution, at which the models are switched (usually 16/32/64); `<level>` is 0 or 1.
For inference (generation) this method works properly only for models from one "family", i.e. uptrained (finetuned) from the same original model. For training may be useful in other cases too (not tested yet).
```
%run src/models_blend.py --pkl1 model1.pkl --pkl2 model2.pkl --res <res> --level <level>
```
Mix few models by stochastic averaging all weights. This would work properly only for models from one "family", i.e. uptrained (finetuned) from the same original model.
```
%run src/models_swa.py --in_dir <models_dir>
```
| github_jupyter |
# RadiusNeighborsRegressor with RobustScaler
This Code template is for the regression analysis using a simple Radius Neighbor Regressor and feature rescaling technique RobustScaler in a pipeline. It implements learning based on the number of neighbors within a fixed radius r of each training point, where r is a floating-point value specified by the user.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import LabelEncoder, RobustScaler
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.neighbors import RadiusNeighborsRegressor
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training .
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Data Rescaling
It scales features using statistics that are robust to outliers.
This method removes the median and scales the data in the range between 1st quartile and 3rd quartile.
[More on RobustScaler module and parameters](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.RobustScaler.html)
### Model
RadiusNeighborsRegressor implements learning based on the neighbors within a fixed radius of the query point, where is a floating-point value specified by the user.
#### Tuning parameters
> **radius**: Range of parameter space to use by default for radius_neighbors queries.
> **algorithm**: Algorithm used to compute the nearest neighbors:
> **leaf_size**: Leaf size passed to BallTree or KDTree.
> **p**: Power parameter for the Minkowski metric.
> **metric**: the distance metric to use for the tree.
> **outlier_label**: label for outlier samples
> **weights**: weight function used in prediction.
For more information refer: [API](https://scikit-learn.org/stable/modules/generated/sklearn.neighbors.RadiusNeighborsRegressor.html#sklearn.neighbors.RadiusNeighborsRegressor)
```
# model initialization and fitting
model = make_pipeline(RobustScaler(),RadiusNeighborsRegressor(radius=2))
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
plt.figure(figsize=(14,10))
plt.plot(range(20),y_test[0:20], color = "green")
plt.plot(range(20),model.predict(x_test[0:20]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Nikhil Shrotri , Github: [Profile](https://github.com/nikhilshrotri)
| github_jupyter |
# Virtual Machine Workload Characterization
## VMware and virtualization
_"We believe that software has the power to unlock new possibilities for people and our planet. Our software forms a digital foundation that powers the apps, services, and experiences transforming the world."_
VMware is an IT company – leader in the server virtualization market. Virtualization is the process of running multiple instances of different computer system in a software layer abstracted from the actual hardware. Most commonly, it refers to running multiple operating systems on a single physical server simultaneously. To the applications running on top of the virtualized machine are not aware that they run on a virtual machine.
## Problem description
As two of the most-mature products, vSphere and vCenter Server provides great deal of customization and flexibility. However, given the complexity of the modern virtualization and the numerous different technologies involved in computing network and storage the customization options for VMware products grew infeasible large for common system administrators to grasp.
At the same time different workloads have different needs. There always is some tradeoff between different functionalities of the system (like throughput and latency or consolidation and redundancy) and there is not one configuration to serve equally well all kind of workloads. Thus, understanding the purpose of the environment is crucial.
And while it is easy to profile and optimize single virtual machine, vSphere stack hosts millions virtual environments. In order to approach their needs proactively we have to find a data driven way to classify them.
</br>
### The Challenge
vSphere stack enables (with the explicit agreement of the owners of the environment) the collection of on demand low level performance telemetry. Based on this data we need to identify groups of virtual machines similar with respect to different properties, such as scale, utilization pattern etc. However, we will diverge from the standard clustering algorithms (although we will use one as supporting tool) and try to achieve this through embeddings.
### But what is an embedding?
Embedding is a representation of high dimensional vector on low dimensional space. Ideally this representation preserves as much information from the original vector by positioning similar inputs close together on the embedding space.
</br>
### The Dataset
The dataset consists of two main data sources related to the performance and the virtual hardware of the virtual machines (VMs).
The **performance telemetry** is organized in a python list, containing multiple python dictionaries. Each dictionary accounts for the data of single VM. The key of the dictionary is the respective ID of the VM, and the value is pandas data frame indexed by the timestamp and containing the actual measurements of each feature.
</br>
Variable Name |Df index| type|unit | Description|
--- | --- |--- | --- |---
ts |yes|timestamp|time|Time stamp (yyyy-mm-dd HH:MM:SS) of the observation|
cpu_run |no|numeric|milliseconds|The time the virtual machine use the CPU|
cpu_ready|no|numeric|milliseconds|The time the virtual machine wants to run a program on the CPU but waited to be scheduled|
mem_active|no|numeric|kiloBytes|The amount of memory actively used by the vm|
mem_activewrite|no|numeric|kiloBytes|Amount of memory actively being written to by the virtual machine.|
net_packetsRx|no|numeric|count|Number of packets received during the interval.|
net_packetsTx|np|numeric|count|Number of packets transmitted during the interval.|
</br>
The **virtual hardware dataset** is a normal rectangular data frame indexed by the id of the VM. It represents “static” features that account basically for the scale of the system.
</br>
Variable Name |Df index| type|unit | Description|
--- | --- |--- | --- |---
id |yes|integer| |Unique identifier of the virtual machine |
memory_mb|no|integer|megabytes|Configured virtual RAM|
num_vcpus|no|integer|count|Number of virtual processor cores|
number_of_nics|no|integer|count|Number of network interface cards|
num_virtual_disks|no|integer|count|Number of the configured hdd|
os_fam|no|categorical|indetity|The operating system of the VM|
</br></br>
## Environment setup
### Before we begin, we should do some groundwork:
* Doing the initial imports
* Mounting Google Drive as our file system
* Setting some path variables and creating working directory
* Download the data from external repo (another location in Google Drive)
* Define some functions that we will reuse many times
```
## SETUP THE ENVIRONMENT
## Import general system modules
from google.colab import drive
from google_drive_downloader import GoogleDriveDownloader
from itertools import product
from itertools import chain
import random
import pickle
import gzip
import sys
import os
## Core modules
import pandas as pd
import numpy as np
## Import plotting libraries
from matplotlib import pyplot as plt
import matplotlib
import seaborn as sns
## ML modules
from sklearn.manifold import TSNE
from sklearn.preprocessing import MinMaxScaler
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
from sklearn.cluster import DBSCAN
import umap
## Plot functions display options
%matplotlib inline
sns.set(rc={'figure.figsize':(11.7,8.27)})
## Mount your google drive as file system
## You should allow this notebook to access your drive
## ultimately receiving temporary token
drive.mount('/content/drive')
## Setup our root dir
gdrive_root = "/content/drive/My Drive/AMLD VMWARE/"
performance_file_name = 'performance_telemetry.gzip'
viritual_hardware_file_name = 'virtual_hardware.csv'
utilities_file = 'workshop_utilities.py'
## Construct the file path of the file we are going to download and use:
performance_data_full_path = gdrive_root + performance_file_name
virtual_hardware_data_full_path = gdrive_root + viritual_hardware_file_name
utilities_file_full_path = gdrive_root + utilities_file
## Check if the default working directory exist and if no, create it:
if not(os.path.exists(gdrive_root)):
os.mkdir(gdrive_root)
print(gdrive_root,' directory has been created!')
## Add the new path to the runtime
## From here we will import some custom functions
sys.path.append(gdrive_root)
## Set the root dir as default
os.chdir(gdrive_root)
print('File paths have been set!')
## This chunk downloads data from our shared storage and makes a "local copy to your personal google drive"
## After the first execution of this chunk the data is cached and in the next run it won’t download the data.
## In order to execute it again, you have to delete the files from the folder (or the entire folder)
## Files location
performance_tm_fileid = '1tQulULMw09XQ5rWKF0fqEzXHRwX2CwBS'
virtual_hw_fileid = "136ZxWcIflPD_yol5Nhkj9LVeYB9-1W6L"
workshop_util_file = '1amC8ZhG7ZQt1fwJoidLC8GF58yXTQrJz'
## Download Data
GoogleDriveDownloader.download_file_from_google_drive(file_id = performance_tm_fileid , dest_path = performance_data_full_path )
GoogleDriveDownloader.download_file_from_google_drive(file_id = virtual_hw_fileid , dest_path = virtual_hardware_data_full_path)
GoogleDriveDownloader.download_file_from_google_drive(file_id = workshop_util_file , dest_path = utilities_file_full_path )
### Print the content of our working folder
#it should contain ['performance_telemetry.gzip', 'virtual_hardware.csv', 'workshop_utilities.py']
print('Dir content:', os.listdir(gdrive_root))
## Import some predefined utility functions
from workshop_utilities import truncate_by_quantile, plot_single_vm, sample_observation, model_params_product, model_train, plot_results, plot_embedding
print('Custom utility functions loaded!')
## Load data in memory for this session:
## Load performance data
with gzip.open(performance_file_name, 'rb') as fconn:
perf_data = pickle.load(fconn)
print('Performance Telemetry successfully loaded !')
## Load virtual hardware
virt_hw = pd.read_csv(viritual_hardware_file_name, sep = ';', index_col = 'id')
print('Virtual Hardware successfully loaded !')
## Define function that applies numeric transformation
def return_transformer(transformer = None, numeric_constant = 1, lambd = 0):
if transformer == None or transformer == 'None':
return(lambda x: x)
elif transformer == 'ln' or (transformer=='boxcox' and lambd == 0):
return(lambda x: np.log(x+numeric_constant))
elif transformer == 'log10':
return(lambda x: np.log10(x+numeric_constant))
elif transformer=='boxcox':
return(lambda x: (np.power(x+numeric_constant, lambd) - 1)/lambd )
### ADD YOUR OWN TRANSFORMATION HERE
### To run all code chunks go to Runtime>Run before (Ctrl + F8)
```
### Env setup end
## First look at our data
```
## Very basic data exploration
## Define some global variables that we will reuse
## N: Number of virtual machines
## D: Number of performance features
## T: Temporal dim: number of time samples
## F: List of features
## VMs: List with all ids of the virtual machines
N = len(perf_data)
VMs = list(perf_data.keys())
T , D = perf_data[VMs[0]].shape
F = list(perf_data[VMs[0]].columns)
print('Dictionary with', N ,' virtual machines!')
print('Each VM has ',T, ' observations in time!')
print('There are',D, 'performance features:', F)
print('\n First rows of the first virtual machine \n')
pd.set_option('display.max_columns', 10)
print(perf_data[1].head())
## D_hw: Number of performance features
## F_hw: List of features
_ , D_hw = virt_hw.shape
F_hw = list(virt_hw.columns)
print('Dictionary with', N ,' virtual machines!')
print('There are',D_hw, 'virtual hardware features:', F_hw, '\n')
print('Missing values:')
print(np.sum(virt_hw.isna()))
print('\nData types:')
print(virt_hw.dtypes)
print('\nData Frame Summary:')
virt_hw.describe()
print(virt_hw.head())
fig, axes = plt.subplots(nrows = 2, ncols=3, sharex = False, sharey = False, figsize = (40, 10))
for i, metric in enumerate(F_hw):
ax = axes.reshape(-1)[i]
data = virt_hw.groupby([metric])[metric].count()
x_ax = [str(i) for i in data.index.values]
ax.bar(x_ax, height = data.values)
ax.set_title(metric)
```
### Inspect single VM series
```
##
DATA = perf_data
VMID = None ## None for random VM
TRANSFORMATION = return_transformer(None)
###
plot_single_vm(__perf_data = DATA
, __vmid = VMID
, transformation = TRANSFORMATION
)
```
### Note on data transformation
Transforming variable means replacing the actual value x with some function of that variable f(x). As a result, we change the distribution and the relationship of this variable in some beneficial way.
Here we will consider two basic techniques:
* Box-Cox(power) transformation - This is a useful data transformation technique used to stabilize variance, make the data more normal distribution-like, improve the validity of measures of association.
* Logarithmic transformation - logarithmic function has many nice properties that are commonly exploited in the analysis of time series. Change of natural log roughly equals percentage change. It also converts multiplicative relationship into additive (log(XY) = log(X) + log(Y)). Natural log transformation is a private case of Box-Cox transformation
* Data truncation - many of the statistics, that we are going to compute are quite sensitive to extreme values and outliers. Thus, it might be worth sacrificing tiny portion of the data in order to limit the effect of extreme values on our estimates. Note that such transformation might obscure the actual distribution and should be applied with care.
```
################
## PARAMETERS ##
################
VMID = None # None or integer between 1 and 707
METRIC = 'mem_activewrite' # If None then random metric is chosen
TRUNCATE_VARIABLE = True #
TRUNC_QRANGE = [0,0.99] # no truncation =[0,1]
TRANSFORMATION = return_transformer(None) ## returns lambda expression. Arguments:None, 'ln', log10, 'boxcox' + lambda
##############
if METRIC == None:
METRIC = F[np.random.randint(0, D)]
VMID, data = sample_observation(perf_data, vmid = VMID)
## Inspect the distribution of the variable:
fig = plt.figure(figsize=(20,10))
gs0 = fig.add_gridspec(2,1)
gs1 = gs0[1].subgridspec(1,2)
ax0 = fig.add_subplot(gs0[0])
ax1 = fig.add_subplot(gs1[0])
ax2 = fig.add_subplot(gs1[1])
ax1.set_title('Untransformed distribution')
ax2.set_title('Transformed distribution')
ax0.set_title('Series in time {feat} for VMID {vmid}:'.format(feat = METRIC, vmid = VMID))
ax0.plot(data.index, data[METRIC], linewidth = 0.9)
ax0_0 = ax0.twinx()
ax0_0.plot(data.index, TRANSFORMATION(data[METRIC]), label = 'transformed', c = 'orange', linewidth = 0.5)
## First plot the original series and then truncate the data
data['truncated'] = data[METRIC]
if TRUNCATE_VARIABLE:
#data = data.apply(lambda x: truncate_by_quantile(x, trunc_range = TRUNC_QRANGE))
data['truncated'] = truncate_by_quantile(data[METRIC], trunc_range = TRUNC_QRANGE)
sns.distplot(data[METRIC].dropna(), kde = False, ax = ax1, bins = 40)
sns.distplot(TRANSFORMATION(data['truncated'].dropna()), kde = False, ax = ax2, bins = 40 )
ax1.set_xlabel(None)
ax2.set_xlabel(None)
plt.legend()
plt.show()
```
### Which transformation?
Among thigs to consider when you transform your variables are what works with the data, does it makes sense and is there a natural interpretation after transforming the data?
</br>
**References:**
[Transformations: an introduction with STATA](http://fmwww.bc.edu/repec/bocode/t/transint.html)
[The logarithm transformation](https://people.duke.edu/~rnau/411log.htm)
[MAKING DATA NORMAL USING BOX-COX POWER TRANSFORMATION](https://www.isixsigma.com/tools-templates/normality/making-data-normal-using-box-cox-power-transformation/)
[Yeo-Johnson Power Transformation](https://www.stat.umn.edu/arc/yjpower.pdf)
</br>
**Some insights regarding the performance telemetry:**
* Our data is strictly non-negative (convenient for power transformations)
* Most of the performance metrics do not exhibit clear trend
* Missing data due do data collection routine
* There are instances with multimodal distribution
## Feature engineering: Basic descriptives
The first stage of our FE is to obtain very basic descriptive statistics. Descriptive statistics are coefficients that summarize some aspect of the empirical distribution of a variable. The most common aspects that are measured by the descriptive statistics are the central tendency (mean, median, mode) and variability (standard deviation, variance, kurtosis and skewness).
<br></br>
**_PROS:_**
* The descriptive are not as sensitive to relatively small amounts of missing data
* Easy and fast to compute and often comes already implemented in the most software packages
* Easy to interpret
**_CONS:_**
* They are not very expressive and can hide a lot of the information
* Some of the basic descriptives are very sensitive to extreme values and need some preprocessing
<br></br>
In order to mitigate the effect of extreme values we will do some statistical tricks, including truncating all values past a threshold and transforming our data on another scale. This way we will improve the statistical properties of our data and make our estimates more robust.
### How will we create our initial features?
1. Apply numeric transformation that will change the scale and the distribution of the initial data
2. Apply multiple function that takes one or more pd.Series as argument and yields single numerical representation of this / these series. Store them in some data structure
3. Flatten this data structure into vector for each virtual machine
Ultimately, we are going to end with tidy rectangular dataset where each row represents single virtual machine and each column represents feature that we have extracted from our original data.
[Pandas Series Methods](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.html)
```
## How can we apply custom functions ?
## Define function that operates on pd Series
def missing_count(series):
return(len(series) - series.count())
def Q25(series):
return(series.quantile(0.25))
def Q50(series):
return(series.quantile(0.50))
def Q75(series):
return(series.quantile(0.75))
## ADD YOUR CUSTOM FUNCTIONS HERE
## Single case example:
## Apply multiple build in aggregation functions with relatively fast implementation
## the available functions are sum, mean, min, max, var, sem, skew, kurt, count
## sem - standard error of the mean
## kurt - kurtosis (4th moment)
## skew - skewness (3rd moment)
## var - variance (2nd moment)
## These functions can be passed as an array to each column of the data frame:
_, data = sample_observation(perf_data)
np.round(data.agg(['mean', 'min', 'max', 'var', 'skew', Q50,missing_count]), 3)
## Correlation matrix
data.corr()
```
#### Creating the features
</br>
By the end of our feature engineering section we will end up with single data frame that combines the information from both descriptive statistics and correlation data.
* Our initial datasets will be called **descriptive_statistics** and **correlation_data**. There is a pandas view that consolidates them into single data frame called **non_temporal_features**
* The features in this data set can be referenced by the column names
* The names of the features, yielded from the descriptive statistics analysis are stored in **descriptive_statistics_columns** variable
* The names of the features, yielded from the correlation matrix are stored in **correlation_features_columns** variable
```
## FEATURE GENERATION CHUNK ##
## FOR EACH VM
## FOR EACH METRIC [cpu_run, cpu_ready ... net_packetsRx]
## 1. TRUNCATE BY QUANTILE
## 2. APPLY TRANSFORMATION
## 3. COMPUTE EACH METRIC IN GENERAL_STATISTICS
## 5. CREATE FEATURE VECTOR WITH COLUMNS {METRIC}_{STATISTIC} [cpu_run_mean, cpu_run_std, .... ]
## 4. COMPUTE CORRELATION
## 5. FOR EACH UNIQUE PAIR OF METRICS CREATE FEATURE VECTOR [corr_cpu_run_cpu_ready, corr_cpu_run_mem_active, ... ]
## 6. COMBINE THE INFORMATION FOR EACH VIRTUAL MACHINE IN TWO DATA FRAMES
GENERAL_STATISTICS = ['mean','std','skew',Q50, Q25, Q75]
TRANSFORMATION = return_transformer('None')
TRUNCATE_VARIABLE = True
TRUNC_QRANGE = [0.001, 0.999]
#######################
## INIT PLACEHOLDERS ##
#######################
descriptive_statistics = []
correlation_data = []
i = 1
for k, data in perf_data.items():
###################
## TRUNCATE DATA ##
###################
if TRUNCATE_VARIABLE:
data = data.apply(lambda x: truncate_by_quantile(x, trunc_range = TRUNC_QRANGE))
## Apply transformation
data = TRANSFORMATION(data)
############################
## Descriptive statistics ##
############################
## Apply aggregation functions
general_descriptives = data.agg(GENERAL_STATISTICS)
## apply modifications to general descriptives:
## 1. converts the data frame to series with index [(summary_statistic,performance_metric)]
general_descriptives = general_descriptives.stack()
## Convert 2d index to 1d by string concat:
general_descriptives.index = general_descriptives.index.map('{0[1]}_{0[0]}'.format)
## Convert the series to data frame and transpose it
general_descriptives = general_descriptives.to_frame().T
## Add the VM indentifier:
general_descriptives['vmid'] = k
########################
## Correlation matrix ##
########################
# Calculate correlation matrix:
correlation_matrix = data.corr()
corr_df_tmp = pd.DataFrame(index=[k])
## Iterate over the elements of the (cross) correlation matrix
## Take the elements below the diagonals
## This way we flatten the correlation matrix into feature vector:
for mrow in range(D):
for mcol in range(D):
if mrow >= mcol:
continue
# Construct new col name:
new_col = 'corr_' + F[mrow] + '_' + F[mcol]
corr_df_tmp[new_col] = correlation_matrix.iloc[mrow,mcol]
if i%100 == 0:
print('Finished iteration', i, 'out of', N)
i += 1
## Finally set vmid as index (on the fly)
## Join correlation data with descriptive statistics:
## as add the record to the data placeholder:
descriptive_statistics.append(general_descriptives.set_index('vmid'))
correlation_data.append(corr_df_tmp)
print('Finished iteration', N, ' out of ', N)
## The loop left us with list of single row data frames both for descriptive statistics and correlation matrix
## Next step is to combine the whole data into one big data frame
descriptive_statistics = pd.concat(descriptive_statistics)
correlation_data = pd.concat(correlation_data)
## We will further process the different families of features
## Thats why we keep their names in separate variables
descriptive_statistics_columns = descriptive_statistics.columns
correlation_features_columns = correlation_data.columns
## Finally merge the final dataset into one
non_temporal_features = descriptive_statistics.merge(correlation_data, left_index = True, right_index= True)
non_temporal_features_index = non_temporal_features.index
## Small trick to ensure alignment between our features and the virtual hardware data
virt_hw = virt_hw.loc[non_temporal_features.index]
```
#### Inspect the features
Visualize the distributions of the features that we’ve created. Please keep in mind the transformation that we’ve applied over the original series.
```
## Inspect the summary statistics of the correlation features
pd.set_option('display.max_columns', 50)
np.round(correlation_data.describe(), 2)
TRANSFORMATION = return_transformer(None)
## Inspect the distribution of cross correlation
g = sns.FacetGrid(pd.melt(TRANSFORMATION(correlation_data)), col = 'variable', col_wrap = 5, aspect = 1.5)
g.map(plt.hist, 'value', bins = 40)
plt.subplots_adjust(top=0.9)
g.fig.suptitle('CORRELATION FEATURES')
```
We can see that the distribution of the correlation is skewed towards the positive correlations . There are several cross-correlation features with bi-modal distributions.
```
## Summary of the descriptive statistics features
pd.set_option('display.max_columns', 50)
print('Min feature value is:', np.min(np.round(descriptive_statistics.describe(), 1).values))
np.round(descriptive_statistics.describe(), 1)
## Inspect the distribution of the descriptive statistics
TRANSFORMATION = return_transformer('ln', numeric_constant=5)
#######
## Transform the data from wide to long format
data = TRANSFORMATION(descriptive_statistics)
data = pd.melt(data)
#data['variable'] = TRANSFORMATION(data['variable'])
## Plot
g = sns.FacetGrid(data, col = 'variable', col_wrap = 6, sharex = False, sharey=False, aspect = 1.5)
g.map(plt.hist, 'value', bins = 40)
plt.subplots_adjust(top=0.9)
g.fig.suptitle('DESCRIPTIVE FEATURES')
```
#### Apply transformation
Each of the following chunks creates a data frame where it applies transformations. We can either choose to leave the data untransformed, to standardize (subtract mean and divide by std) or to normalize data within some numeric range (typically 0, 1 or -1,1).
Typically, is not good idea to apply different scaling to the different features. A rule of the thumb (at least for this workshop) is to have up to one transformation and one scaling.
##### Descriptive statistics transformations
```
############################
##:: DATA PREPROCESSING ::##
##:: DESCRIPTIVES ::##
############################
## Apply transformation
TRANSFORMATION = return_transformer(None)
APPLY_STANDARD_SCALING = False
APPLY_MIN_MAX_SCALING = True
MIN_MAX_SCALER_RANGE = (-1, 1)
##############
## We would like to retain the original features
## Thus we create hard copy of the data and operate on it:
descriptive_statistics_transformed = descriptive_statistics.copy()
## Rescale descriptive statistics data to within MIN MAX SCALER RANGE
## The correlation features are naturally scaled within this range
min_max_scaler_instance = MinMaxScaler(feature_range = MIN_MAX_SCALER_RANGE)
standart_scaler_instance = StandardScaler()
## Apply transformation
descriptive_statistics_transformed[descriptive_statistics_columns] = TRANSFORMATION(descriptive_statistics_transformed[descriptive_statistics_columns])
## APPLY MIN-MAX SCALING TO DESCRIPTIVE STATISTICS DATA
if APPLY_MIN_MAX_SCALING:
descriptive_statistics_transformed[descriptive_statistics_columns] = min_max_scaler_instance.fit_transform(descriptive_statistics_transformed[descriptive_statistics_columns])
## APPLY MIN-MAX SCALING TO DESCRIPTIVE STATISTICS DATA
if APPLY_STANDARD_SCALING:
#modeling_data[STANDARD_SCALING_COLUMNS] = standart_scaler_instance.fit_transform(modeling_data[STANDARD_SCALING_COLUMNS])
descriptive_statistics_transformed[descriptive_statistics_columns] = standart_scaler_instance.fit_transform(descriptive_statistics_transformed[descriptive_statistics_columns])
```
##### Correlation features transformations
```
############################
##:: DATA PREPROCESSING ::##
##:: CORRELATION ::##
############################
## Apply transformation
TRANSFORMATION = return_transformer(None)
APPLY_STANDARD_SCALING = False
APPLY_MIN_MAX_SCALING = False
MIN_MAX_SCALER_RANGE = (-1, 1)
###############
## We would like to retain the original features
## Thus we create hard copy of the data and operate on it:
correlation_data_transformed = correlation_data.copy()
## Rescale descriptive statistics data to within MIN MAX SCALER RANGE
## The correlation features are naturally scaled within this range
min_max_scaler_instance = MinMaxScaler(feature_range = MIN_MAX_SCALER_RANGE)
standart_scaler_instance = StandardScaler()
## Apply transformation
correlation_data_transformed[correlation_features_columns] = TRANSFORMATION(correlation_data_transformed[correlation_features_columns])
## APPLY MIN-MAX SCALING TO DESCRIPTIVE STATISTICS DATA
if APPLY_MIN_MAX_SCALING:
correlation_data_transformed[correlation_features_columns] = min_max_scaler_instance.fit_transform(correlation_data_transformed[correlation_features_columns])
## APPLY MIN-MAX SCALING TO DESCRIPTIVE STATISTICS DATA
if APPLY_STANDARD_SCALING:
#modeling_data[STANDARD_SCALING_COLUMNS] = standart_scaler_instance.fit_transform(modeling_data[STANDARD_SCALING_COLUMNS])
correlation_data_transformed[correlation_features_columns] = standart_scaler_instance.fit_transform(correlation_data_transformed[correlation_features_columns])
```
##### Virtual hardware transformations
```
###############################
###:: DATA PREPROCESSING ::####
###::VIRTUAL HARDWARE DATA::###
###############################
## Apply transformation
TRANSFORMATION = return_transformer(None)
APPLY_STANDARD_SCALING = False
APPLY_MIN_MAX_SCALING = True
MIN_MAX_SCALER_RANGE = (-1, 1)
## Make copy of the original data
virt_hw_transformed = virt_hw.copy()
virt_hw_transformed_columns = virt_hw_transformed.columns
min_max_scaler_instance = MinMaxScaler(feature_range = MIN_MAX_SCALER_RANGE)
## One hot encode categorical
virt_hw_numerics = []
for col in virt_hw_transformed_columns:
col_type = virt_hw_transformed[col].dtypes.name
if col_type == 'object' or col_type == 'category':
dummy_vars = pd.get_dummies(virt_hw_transformed[col], prefix = col)
virt_hw_transformed = pd.concat([virt_hw_transformed,dummy_vars],axis=1)
virt_hw_transformed.drop([col],axis=1, inplace=True)
else:
virt_hw_transformed[col] = TRANSFORMATION(virt_hw_transformed[col])
virt_hw_numerics.append(col)
if APPLY_MIN_MAX_SCALING:
min_max_scaler_instance = MinMaxScaler(feature_range = MIN_MAX_SCALER_RANGE)
virt_hw_transformed[virt_hw_numerics] = min_max_scaler_instance.fit_transform(virt_hw_transformed[virt_hw_numerics])
if APPLY_STANDARD_SCALING:
standart_scaler_instance = StandardScaler()
virt_hw_transformed[virt_hw_numerics] = standart_scaler_instance.fit_transform(virt_hw_transformed[virt_hw_numerics])
```
##### Assemble the dataset for modeling
```
## CREATE FINAL DATASET ##
## CHOOSE WHICH FEATURES TO INCLUDE FOR THE FINAL DATASET
## BY DEFAULT, WE WILL MODEL ALL FEATURES THAT WE’VE CREATED
## HOWEVER, WE CAN MAKE MODELS OUTPUT EASIER TO INTERPRET IF WE USE INFORMATION ONLY FOR ONE RESOURCE TYPE (CPU or NETWORK)
## To include columns that contains cpu_run
## descriptive_statistics_transformed.filter(regex = 'cpu_run')
## To exclude columns containing cpu_run string
## descriptive_statistics_transformed.loc[:,~descriptive_statistics_transformed.columns.str.contains('cpu_run')]
first_stage_data_list = [
descriptive_statistics_transformed
,correlation_data_transformed
,virt_hw_transformed
]
## Init empty data frame:
first_stage_data = pd.DataFrame(index = non_temporal_features_index)
##
for i in first_stage_data_list:
#data = i.copy() ## in the cases where there will be more transformations
data = i
first_stage_data = pd.merge(first_stage_data, data, left_index=True, right_index=True)
print('Final dataset is with shape:', first_stage_data.shape)
### To run all code chunks go to Runtime>Run before (Ctrl + F8)
```
## Principal component analysis
Although PCA can be though as form of embedding by itself, despite all benefits (speed, efficiency, interpretability), it suffers from a major drawback. It is tricky to capture (by default) non-linear relationships.
</br>
**References:**
https://towardsdatascience.com/a-one-stop-shop-for-principal-component-analysis-5582fb7e0a9c
http://setosa.io/ev/principal-component-analysis/
[Making sense of principal component analysis, eigenvectors & eigenvalues](https://stats.stackexchange.com/questions/2691/making-sense-of-principal-component-analysis-eigenvectors-eigenvalues)
```
from sklearn.decomposition import PCA
## Plot correlation matrix
cmap = sns.diverging_palette(220, 20, sep=20, as_cmap=True)
sns.clustermap(first_stage_data.corr(), figsize= (20,20), cmap = cmap)
plt.show()
## One more thing to consider is the correlation among the variables:
pca_instance = PCA().fit(first_stage_data)
## Taking care to preserve the index for later join:
first_stage_data_pca = pd.DataFrame(pca_instance.transform(first_stage_data), index = first_stage_data.index)
fig, ax = plt.subplots(figsize=(20,8),nrows = 1 , ncols = 2)
sns.scatterplot(x = first_stage_data_pca.iloc[:,0], y = first_stage_data_pca.iloc[:,1], ax = ax[0])
ax[0].set_title('Principal Components')
ax[0].set_xlabel("PC 1")
ax[0].set_ylabel("PC 2")
ax[1].bar(x = [i+1 for i in range(pca_instance.n_components_)], height = pca_instance.explained_variance_/ np.sum(pca_instance.explained_variance_))
ax2 = ax[1].twinx()
ax2.plot([i+1 for i in range(pca_instance.n_components_)], np.cumsum(pca_instance.explained_variance_)/ np.sum(pca_instance.explained_variance_), c = 'orange')
ax2.grid([])
ax2.set_ylim(bottom =0)
ax[1].set_title('Explained variance plot:')
ax[1].set_xlabel('Number of principal components')
ax[1].set_ylabel('Explained variance')
ax2.set_ylabel('Cumulative variance')
##
```
The easiest way to reduce the redundancy of our data is to apply PCA.
PCA is beneficial preprocessing step for t-SNE, recommended both by the author and by the ML practitioners.
Number of components to include is a tunable parameter by itself and initially we can adopt the “elbow” approach. Generally we should identify where the marginal explained variance starts to diminish.
## t-Distributed Stochastic Neighbor Embedding (t-SNE)
Intended as a technique for dimensionality reduction well suited for the visualization of high-dimensional datasets
**Materials**
[Official Website With papers and implementations under different languages](https://lvdmaaten.github.io/tsne/)
[How to Use t-SNE Effectively](https://towardsdatascience.com/how-exactly-umap-works-13e3040e1668) an amazing interactive blogpost
[How to tune hyperparameters of tSNE](https://towardsdatascience.com/how-to-tune-hyperparameters-of-tsne-7c0596a18868)
[sklearn implementation](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html)
```
from sklearn.manifold import TSNE
## Simple t-SNE usage:
## Tsne follows the sklearn interface :
# 1. We create instance of the object
# Here we set the parameters that are internal to the algorithm
TSNE_Instance = TSNE(n_components = 2 ## Controls the number of dimebsuibs if the embedded space
,perplexity = 10 ## Perplexity controls how to balance attention between local and global aspects of your data
,early_exaggeration = 12 ## Controls how tight natural clusters are situated
,learning_rate = 100 ## Controls the step during the gradient descent stage
,n_iter = int(1e3) ## Max number of iterations
,random_state = 500 ## Random seed
,verbose = 2 ## controls logging of the algorithm
)
# 2.Fit the object to the data
TSNE_results = TSNE_Instance.fit_transform(first_stage_data)
# The result of fit_transform is ndarray with shape (NSamples, n_components)
plt.figure(figsize=(7,7))
plt.scatter(TSNE_results[:, 0], TSNE_results[:, 1])
plt.xlabel('X')
plt.ylabel('Y')
```
### t-SNE tunable parameters
**n_components** - similar to N components In PCA. It sets the cardinality of your output feature set. sklearn t-sne implementation supports up to 3d plane.
_Practical note:_ algorithm becomes exponentially slow with the increase of this parameter
**perplexity** - Perplexity controls how to balance attention between local and global aspects of your data. Low values yield many small scattered clusters while large values tend to clump up the data. This parameter is very data dependent and it should be explored every time
**PCA_components** - Although not internal for the algorithm in practice t-SNE benefits and in a way complements PCA preprocessing. With the former the linear structure of the data is captured while with the former technique can capture non-linear latent structure.
### Technical note on the utility functions
Since we’ve intended this workshop to be more of a decision making exercise rather that coding one, we are about to introduce several utility functions, defined in order to visualize and automate some of the routines that about to go through.
</br>
#### model_train()
**model_train** : this function will be used to train the embedding with multiple predefined values of the tunable parameters:
* data - pandas data frame or numpy matrix. The data is restricted to numeric arrays without missing observations
* model - string with one of the following values: _"TSNE", "DBSCAN", "UMAP"_
* param_dict - python dictionary with tunable parameters. The name of the parameter must be the key of the dictionary, the value must be **python list** with values.
</br>
param_dict = {
'perplexity':[10,20]
, n_iter:[300,500]
}
* apply_pca - boolean parameter that indicates if the original data will be PCA transformed.
* n_pc - Number of principal components applied as **list of integers** (n_pc = [10, 20]). This parameter is considered as tunable and it is added under the tunable parameters in the output of the function. It is ignored if apply_pca = False
* other_params - non-tunable parameters. Like __param_dict__ the keys should be the name of the parameter. Unlike __param_dict__ the value should not be list
</br>
param_dict = {
'learning_rate':np.float32(1e-3)
}
The result of the function is python list. Each element contains triple object where:
* First element of the triple is the dictionary with tunable parameter values
* The second element is the coordinates of the embedding
* The last element is the non-tunable parameters
This result is fed to several plotting functions in order to produce neat visualizations that will help us interpret the results of our embedding
####plot_results()
This is one of the first plotting function that is designed for exploration of the tuning results. The way it works it that it takes object, generated from model_train() function and creates grid of plots arranged by the values of the parameters that have been used for the embeddings.
Important to note is that this function **plots only the first two dimensions** of the embedding. If the number of dimensions is higher than 2 plot_embedding() can be used to visualize all dimensions.
**Arguments:**
* results - results, generated from model_train()
* leading_param = None - the name of the parameter that should be used to arrange the tuning results row wise.
* color_array - list of numpy arrays that will be used to apply color to the scatterplot matrix. If given, the length of the list should be either 1 (all scatters are colored with the same variable) or equal to the length of the results object (then each object is colored with its respective color array). The second option will be used during exploring the results from DBSCAN clustering.
* force_categorical = False - if the color array should be represented on a discrete color scale. This option is meaningful for integer color arrays with limited number of distinct values
* point_size = 3 - controls the size of the points in the scatterplot
#### plot_embedding()
This is a function that is designed to plot single embedding. It takes **single** object, generated from model_train() and plots all dimensions are scatterplot matrix.
**Arguments**
* result - single element subset from model_train() results.
* fig_scale - plot scaling parameter
* color_var_list- list of numpy arrays that is used for coloring
* force_categorical = False - if the color array should be represented on a discrete color scale. This option is meaningful for integer color arrays with limited number of distinct values
* plot_centers = FALSE - used when we plot DBSCAN clusters. It overlays the cluster centers
* dont_plot_minus_one - used when we plot DBSCAN clusters. Don't plot outlier centers
* point_size = 4 - controls the size of the points in the scatterplot
### Tune T-SNE
```
## EXPERIMENT WITH DIFFERENT TSNE SETTINGS ##
## TSNE PARAMETERS
## https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html
MODEL = 'TSNE'
TUNABLE_PARAMETERS = {'perplexity':[25]}
DEFAULT_PARAMETERS = {'learning_rate':np.int(1e2)}
DATA = first_stage_data
APPLY_PCA = True
N_PC = [30,50, 55]
## Apply tuning:
TSNE_results = model_train( data = DATA
, model = MODEL
, param_dict = TUNABLE_PARAMETERS
, apply_pca = APPLY_PCA
, n_pc = N_PC
, other_params = DEFAULT_PARAMETERS
)
## Inspect the results of the parameters
################
## PARAMETERS ##
################
RESULTS = TSNE_results
LEADING_PARAM = 'n_pc'
COLOR_ARRAY = None
FORCE_CATEGORICAL = False
POINT_SIZE = 5
## APPLY FUNCTION
plot_results(results = RESULTS
,leading_param = LEADING_PARAM
,color_array = COLOR_ARRAY
,force_categorical = FORCE_CATEGORICAL
,point_size = POINT_SIZE
)
## If single combination is being expected
##plot_embedding(RESULTS[0], fig_scale= 1.2,point_size = 5)
```
### Capture "clusters" through DBSCAN :
We will use Density-based spatial clustering of applications with noise (DBSCAN) algorithm to address the different "clusters" in our embedding. This is not a true clustering because t-SNE eliminated the actual structure of the data. However, this pseudo clustering step is useful and can guide us through the profiling of our final clusters
The tunable parameters that we are going to consider are:
**eps:** specifies how close points should be to each other to be considered a part of a cluster. It means that if the distance between two points is lower or equal to this value (**eps**), these points are considered neighbors
**min_samples:** the minimum number of points to form a dense region. For example, if we set the **min_samples** parameter as 5, then we need at least 5 points to form a cluster.
</br>
**Several references:**
[DBSCAN Original paper](http://www2.cs.uh.edu/~ceick/7363/Papers/dbscan.pdf)
[Medium post](https://towardsdatascience.com/machine-learning-clustering-dbscan-determine-the-optimal-value-for-epsilon-eps-python-example-3100091cfbc)
[sklearn implementation](https://scikit-learn.org/stable/modules/generated/sklearn.cluster.DBSCAN.html)
```
from sklearn.cluster import DBSCAN
```
```
## Choose one embedding to operate with
## Zoom plot in :
RESULTS = TSNE_results
MODEL_INDEX = 1
###
plot_embedding(RESULTS[MODEL_INDEX], fig_scale = 2, point_size = 9)
```
#### Tune DBSCAN
```
## Capture the clusters with dbscan
TUNABLE_PARAMETERS = {'eps': [4,5,6]
, 'min_samples':[15,30,45]}
DEFAULT_PARAMETERS = {}
MODEL = 'DBSCAN'
## Apply tuning:
dbscan_clusters = model_train( data = RESULTS[MODEL_INDEX][1]
, model = MODEL
, param_dict = TUNABLE_PARAMETERS
, other_params = DEFAULT_PARAMETERS
)
## Plot the results from DBSCAN clustering
result_for_plotting = [(i[0],RESULTS[MODEL_INDEX][1],i[2]) for i in dbscan_clusters]
dbscan_clusters_results = [i[1] for i in dbscan_clusters]
plot_results(results = result_for_plotting, color_array=dbscan_clusters_results, force_categorical=True, point_size=5)
## Zoom one of the plots:
RESULTS = TSNE_results
CLUSTER_INDEX = 3
plot_embedding(RESULTS[MODEL_INDEX]
, fig_scale = 2.2
, color_var_list = [dbscan_clusters[CLUSTER_INDEX][1]]
, force_categorical=True
, plot_centers=True
, point_size = 10)
```
#### Profile Clusters
```
## COMPARE THE CLUSTERS
## PICK METRICS BY WHICH CLUSTERS WOULD BE COMPARED
DATA = descriptive_statistics
METRICS = ['cpu_run_mean', 'cpu_ready_mean']
### CREATE SUMMARY TABLE
pd.set_option('display.max_rows', None)
pd.options.display.float_format = '{:,.3f}'.format
data = DATA[METRICS].copy()
data['dbscan_clusters'] = dbscan_clusters[CLUSTER_INDEX][1]
display(np.round(pd.melt(data, id_vars = ['dbscan_clusters']).groupby(['variable', 'dbscan_clusters']).describe(), 3))
pd.set_option('display.max_rows', 30)
## PARAMETERS ##
TRANSFORMATION = return_transformer(None)
#DATA = virt_hw.loc[:, virt_hw.columns != 'os_fam'] ## it is categorical variable
DATA = correlation_data
## Plot the distribution as boxplot
clustering_df = pd.DataFrame(index = first_stage_data.index)
clustering_df['dbscan_clusters'] = dbscan_clusters[CLUSTER_INDEX][1]
data = DATA.copy()
data = TRANSFORMATION(data)
data = pd.merge(data,clustering_df, left_index=True, right_index=True)
data = pd.melt(data, id_vars = ['dbscan_clusters'])
ordr = list(sorted(set(dbscan_clusters[CLUSTER_INDEX][1])))
g = sns.FacetGrid(data, col = 'variable', col_wrap = 5, sharex = False, sharey=False, height= 5)
g.map(sns.boxplot, 'dbscan_clusters', 'value').add_legend()
plt.show()
## Parameters:
DATA = correlation_data
COLOR_VAR = 'corr_net_packetsRx_net_packetsTx'
TRANSFORMATION = return_transformer(None)
RESULTS = TSNE_results
###
plot_embedding( RESULTS[MODEL_INDEX]
, fig_scale = 2
, color_var_list = [TRANSFORMATION(DATA[COLOR_VAR].values)]
, point_size = 20
)
## Utility code that helps us compare the embedding generated from pca to the embedding
#pca_result_construct = ({}, first_stage_data_pca.iloc[:,0:5].values,{})
#plot_embedding(pca_result_construct, fig_scale=.8, color_var_list = [TRANSFORMATION(DATA[COLOR_VAR].values)])
## CHECK ORIGINAL SIGNALS ARE DIFFERENT FOR THE DIFFERENT CLUSTERS ##
clusters_to_compare = [1,9]
samples_per_cluster = 5
metric = 'net_packetsRx'
DATA = perf_data
sharey = False
### Fig scaling:
num_cl_to_compare = len(clusters_to_compare)
####
sampled_df = clustering_df[clustering_df['dbscan_clusters'].isin(clusters_to_compare)].groupby('dbscan_clusters').apply(lambda x: x.sample(samples_per_cluster))
fig, axes = plt.subplots(nrows = samples_per_cluster, ncols = num_cl_to_compare, figsize = (15*num_cl_to_compare, 5*samples_per_cluster), sharey=sharey)
ix_i = 0
for i in clusters_to_compare:
_tmp = list(sampled_df.iloc[sampled_df.index.get_level_values(0).isin([i]),:].index.get_level_values(1))
ix_j = 0
for j in _tmp:
axes[ix_j, ix_i].plot(DATA[j][metric])
axes[ix_j, ix_i].set_ylabel('vmid:'+str(j) + '('+str(i)+')')
axes[ix_j, ix_i].yaxis.set_label_position('right')
ix_j += 1
ix_i += 1
```
## Advanced Feature Engineering
### Missing data Imputation
The statistical tests are sensitive or do not tolerate at all missing data.
#### Univariate imputation methods:
* Instance Imputation
* Summary statistics
* Interpolation
* Moving average
```
## Get an intuition about the missing data
## Calculate the % of missing observations:
dta = []
for k, data in perf_data.items():
dta.append(data.isnull().sum().sum() / (T*D))
## Inspect the distribution of missing values
sns.distplot(dta, kde = False, bins = 30)
plt.show()
```
We miss up to 5% of data as the majority of the cases lack only 1-2%
```
## This is a toy array where we can observe the effect of different imputation methods on the data
## These parameters control the steepness of the data
START = 1
STOP = 10
POW = 1
POLY_ORDER = 2
TRANSFORMATION = return_transformer('None')
###
tmp = TRANSFORMATION(pd.Series(np.random.normal(size = 100) + np.linspace(start = START, stop = STOP, num = 100)**POW))
tmp_nan = tmp[43:56].copy()
tmp[45:55] = np.nan
fig = plt.figure(figsize = (18, 6))
ax = fig.add_axes([0,0,1,1])
ax.set_title('Time series imputation methods:')
ax.plot(tmp, label = 'base ts')
base_line = plt.plot(tmp_nan, linestyle = "--", label = 'true values')
ax.plot(tmp.interpolate(method = 'linear')[44:56], label = 'linear interpolation')
## Polynomial
poly_order_label = 'poly '+ str(POLY_ORDER) +' interpolation '
plt.plot(tmp.interpolate(method = 'polynomial', order = POLY_ORDER)[44:56], label = poly_order_label)
tmp_ma_data = tmp.copy()
tmp_ma_data[44:56] = tmp.rolling(12, min_periods = 1).mean()[44:56]
ax.plot(tmp_ma_data[43:57], label = 'Simple MA')
#ax.plot(tmp.ewm(com = 0.5).mean(), label = 'EW MA') MA with Exponential decay
#ax.plot(tmp.rolling(12, min_periods = 1).mean()[44:56], label = 'Simple MA')
plt.legend()
## CREATE NEW DATASET WITH IMPUTED VALUES
IMPUTATION_METHOD = 'interpolation' # mean, median, mode, interpolation, ma
POLY_ORDER = 1 ## The order of the interpolation. 1 = linear
MA_RANGE = 6
TRANSFORMATION = return_transformer(None)
##
OVERWRITE_MA_RANGE = True # IF the max_gap within series > MA_RANGE use MA_RANGE = max_gap.
## Calculate the largest gap
gap_df_list = []
perf_data_no_missing = {}
i = 0
for k, data in perf_data.items():
data_running_copy = data.copy()
data_running_copy = TRANSFORMATION(data_running_copy)
running_dict = {}
for col in F:
max_gap = data[col].isnull().astype(int).groupby(data[col].notnull().astype(int).cumsum()).sum().max()
running_dict[col] = max_gap
## Apply Imputation:
if IMPUTATION_METHOD == 'mean':
data_running_copy[col].fillna(data_running_copy[col].mean(), inplace=True)
elif IMPUTATION_METHOD == 'median':
data_running_copy[col].fillna(data_running_copy[col].median(), inplace=True)
elif IMPUTATION_METHOD == 'mode':
data_running_copy[col].fillna(data_running_copy[col].mode(), inplace=True)
elif IMPUTATION_METHOD == 'interpolation':
data_running_copy[col] = data_running_copy[col].interpolate(method = 'polynomial', order = POLY_ORDER)
elif IMPUTATION_METHOD == 'ma':
## Init the ma parameter
ma_range_running = MA_RANGE
## Subtle hack: If the largest gap > MA_RANGE make MA_RANGE = largest gap for this metric
if MA_RANGE <= max_gap and OVERWRITE_MA_RANGE:
ma_range_running = max_gap +1
data_running_copy[col].fillna(data_running_copy[col].rolling(ma_range_running, min_periods = 1).mean(), inplace=True)
gap_df_list.append(pd.DataFrame(running_dict, index = [k]))
perf_data_no_missing[k] = data_running_copy
i += 1
if i%100 == 0:
print('Iteration', i, 'out of', N )
print('Iteration', i, 'out of', N,'\nDone')
## Zoom to random place in our data in order to see how our gaps have been patched:
MISS_RANGE = 1
DATA_RANGE = np.int(144/4)
## Get random metric
any_missings = 0
while any_missings == 0:
metric_tmp = F[np.random.randint(low = 0, high = D)]
vmid , data = sample_observation(data = perf_data)
any_missings = data[metric_tmp].isnull().sum()
## Get the first and the last data
first_ix = np.min(data[metric_tmp].index)
last_ix = np.max(data[metric_tmp].index)
## Get the indices where the data is missing
missing_data_ix = data[metric_tmp].index[data[metric_tmp].apply(np.isnan)]
## Trick to ensure that we have gap centered between non-missing sequence
miss_date_ix = np.min([dix for dix in missing_data_ix if dix - pd.Timedelta(minutes=5 * DATA_RANGE / 2) > first_ix and dix + pd.Timedelta(minutes=5 * DATA_RANGE / 2) < last_ix])
##
data_ix_reconstruct = [miss_date_ix + pd.Timedelta(minutes = 5*i) for i in range(-DATA_RANGE, DATA_RANGE+1)]
##
tmpdata_orig = data[data.index.isin(data_ix_reconstruct)][metric_tmp]
tmpdf = pd.DataFrame(index = tmpdata_orig.index)
##
miss_lst = []
for i in tmpdata_orig[np.isnan(tmpdata_orig)].index:
# print(i)
miss_lst.append(i)
miss_lst.append(i + pd.Timedelta(minutes=5))
miss_lst.append(i - pd.Timedelta(minutes=5))
miss_lst = list(sorted(set(miss_lst)))
##
fig = plt.figure(figsize = (16,4))
ax = fig.add_axes([0,0,1,1])
ax.plot(data_ix_reconstruct, TRANSFORMATION(data[data.index.isin(data_ix_reconstruct)][metric_tmp]), label = 'original')
ax.plot(miss_lst, perf_data_no_missing[vmid][perf_data_no_missing[vmid].index.isin(miss_lst)][metric_tmp], label = 'imputed', linestyle = "--")
ax.set_title('vmid: '+ str(vmid) + ' ' + metric_tmp)
plt.legend()
plt.show()
#pd.concat(gap_df_list).describe()
## Double check if any variable is still missing
dta_imputed = []
for k, data in perf_data_no_missing.items():
dta_imputed.append(data.isnull().sum().sum() / (T*D))
## Check if there are instances with missing values:
still_missing = [i for i, k in enumerate(dta_imputed) if k > 1e-3]
data = [list(perf_data_no_missing.keys())[i] for i in still_missing]
print('VMs with at least 1 missing observation:' ,len(data))
#
#for i in tmp:
# del perf_data_no_missing[i]
#print(len(perf_data_no_missing.keys()))
```
### Time series specific features
**_PROS_**
More expressive
retain the information encoded in the time domain of the data
**_CONS_**
costly to compute
hard to interpret - thus to profile cluster
requires domain knowledge
not trivial to ensure comparability
some of the tests are very sensitive to data inconsistencies
### Spectral Density
Another set of features that we will consider are related to the presence (or absence) of periodic patterns in the data. One way to check if series show any systematic fluctuations is through spectral decomposition of the data.
The general idea is that any real time series (signal) can be approximated through linear combination of sine waves with different frequencies. The amplitude of these sine waves account for the variation, captured through the decomposition.
</br>
**Resources:**
[Relationship between fft and psd](https://dsp.stackexchange.com/questions/24780/power-spectral-density-vs-fft-bin-magnitude)
[Power vs. Variance](https://dsp.stackexchange.com/questions/46385/variance-in-the-time-domain-versus-variance-in-frequency-domain)
[But what is the Fourier Transform? A visual introduction.](https://www.youtube.com/watch?v=spUNpyF58BY)
[Introduction to the Fourier Transform](https://www.youtube.com/watch?v=1JnayXHhjlg)
[Spectral Analysis in R](https://ms.mcmaster.ca/~bolker/eeid/2010/Ecology/Spectral.pdf)
#### Compute spectral density related features
```
## Here we define the workhorse function that will "slide" through the
## spectrogram and extract spectral density volumes within specific intervals
## Arguments:
## df: pandas data frame that contains the time series data
## series_name: Imputed Series
## freq_arr: Time frequencies at which the spectrogram will be captured
## sm_win: Smoothing window +/- Nsamples
## sm_fun: The function that will be used to aggregate the values from the window
## remove_mean: Bool indicating if mean value should be subtracted from the series
## poly_filt: filter polynomial trend
## poly_deg: the degree of the trend (deg 1 = linear trend)
## return_series: If we want to return the frequency/ fft arrays (potentially for plotting)
def calc_PSD_features( df
, series_name
, freq_arr = None
, sm_win = 2
, sm_fun = np.mean
, remove_mean = True
, poly_filt = True
, poly_deg = 1
, return_series = False
):
data = df[series_name]
T_data = len(data)
## Subtract mean:
if remove_mean:
data = data - np.mean(data)
## Remove linear trend:
if poly_filt:
poly_coef = np.polyfit(range(T_data), data, deg = poly_deg)
# This is the poly object that will actually fit the data with the poly_coef
poly_fit = np.poly1d(poly_coef)
## Apply poly filtering
data = data - poly_fit(range(T_data))
## Apply fft:
data_fft = np.fft.fft(data)
## Transform fft to psd
data_fft = np.power(np.abs(data_fft), 2) / T_data
## Include option for returning the PSD arrays for plotting:
if return_series:
return(data_fft)
## Construct the time window at which the data will be aggregated:
## Keep the index in a tuple
filt_win = range(-sm_win , sm_win+1)
per_freq = np.array(range(T_data))
## This creates tuple with freq_value (like 1, 2, 3 etc hours and their position in the array)
time_window_array = [(ix, np.where(per_freq == ix+1) + np.array(filt_win)) for ix in freq_arr]
## Calculate Total Power of the singal / ts
total_power = np.sum(data_fft)
average_power = np.mean(data_fft) ## Under mean = 0 average power = variance
## Calculate the captured power within the given time range
## The column_name corresponds to: {variable_name}_{freq}
data_dict = {}
#ddict_total = '{}_uncaptured'.format(series_name)
#data_dict[ddict_total] = 1
for fr, data_ix in time_window_array:
ddict_key = '{}_pwidth_{}'.format(series_name, fr)
data_dict[ddict_key] = sm_fun(data_fft[data_ix[data_ix > 0]]/average_power)
#data_dict[ddict_total] -= data_dict[ddict_key]
#data_dict[ddict_key] = (sm_fun(data_fft[data_ix[data_ix > 0]])*2)/total_power
## Add Uncaptured variance:
return(data_dict)
```
#### Inspect spectrogram for single vm
```
## Inspect the spectrogram for single VM
DATA = perf_data_no_missing
vmid = 234
metric = 'net_packetsRx'
##
data = calc_PSD_features(DATA[vmid]
,series_name = metric
,return_series = True
)
##
fig, axes = plt.subplots(nrows = 2, figsize = (15,7))
axes[0].plot(DATA[vmid][metric] - np.mean(DATA[vmid][metric].values))
axes[0].set_ylabel('Original Series')
axes[0].yaxis.set_label_position('right')
axes[1].plot(data[:1008])
axes[1].scatter([7,14,21,28,168,336,504, 672], [0,0,0,0,0,0,0,0], c = 'red', s = 5)
axes[1].set_ylabel('Unsmoothed spectrogram')
axes[1].yaxis.set_label_position('right')
plt.show()
## Calculate the psd features for the entire dataset
## Iterate over vms and variables
## and apply function:
########################
## TUNABLE PARAMETERS ##
########################
TRANSFORMATION = 'ln' # one of: None, 'ln', log10, 1/x, -1/x ,'cuberoot'
#FREQ_ARRAY = range(1, 7) ## We will extract the density for each hour
FREQ_ARRAY = [7,14,21,28,168,336,672] ## We will extract the density for each hour
SMOOTHING_WINDOW = 2 ## We will smooth them with +/- SWIN samples
REMOVE_MEAN = True
POLY_FIT = True # We will fit polynomial ...
POLY_DEG = 1 # ... of degree POLY_DEG and subtract it from the original data ultimately eliminating the respective trend
### FREQ_ARRAY INTERPRETATION
# FREQ_ARRAY shows where we expect to observe cycles
# the values in FREQ_ARRAY represents number of cycles per our entire observation period
# thus FREQ_ARRAY = 7 means that we expect to have 7 cycles/ for 1 week timeseries data that we have this corresponds to 1 cycle per day or daily cyclicity
# 14 = half day cyclicity / 12 hours
###########################
## CREATE PSD DATA FRAME ##
###########################
i = 1
psd_features = []
for k, data in perf_data_no_missing.items():
psd_res_ph = {}
transformation = return_transformer(TRANSFORMATION)
data = transformation(data)
for col in data.columns:
psd_running = calc_PSD_features(data, col
, freq_arr = FREQ_ARRAY
, sm_win = SMOOTHING_WINDOW
, remove_mean = REMOVE_MEAN
, poly_filt = POLY_FIT
, poly_deg = POLY_DEG
, sm_fun = np.mean
)
dict.update(psd_res_ph, psd_running)
psd_features.append(pd.DataFrame(psd_res_ph, index = [k]))
if i%100 ==0:
print('Iteration ', i, 'out of', N)
i += 1
print('Done')
## Combine all elements into single pandas data frame :
psd_features = pd.concat(psd_features)
## Combine the PSD features with the ADF features into single data frame
psd_features.describe()
TRANSFORMATION = return_transformer(None)
## inspect the distribution of the features:
## NB: The data has been log transformed
g = sns.FacetGrid(pd.melt(TRANSFORMATION(psd_features)), col = 'variable', col_wrap = 6, sharex = False, sharey=False, aspect = 1.5)
g.map(plt.hist, 'value', bins = 40)
```
## Second wave of modeling:
### Scale PSD features
```
#########################################
##:: psd_features DATA PREPROCESSING ::##
#########################################
## Apply transformation
TRANSFORMATION = return_transformer(None)
APPLY_STANDARD_SCALING = False
APPLY_MIN_MAX_SCALING = True
MIN_MAX_SCALER_RANGE = (-1, 1)
## We would like to retain the features
## Thus, we create hard copy of the data and operate on it:
psd_features_transformed = psd_features.copy()
psd_features_columns = psd_features.columns
psd_features_transformed[psd_features_columns] = TRANSFORMATION(psd_features_transformed[psd_features_columns])
## APPLY MIN-MAX SCALING TO DESCRIPTIVE STATISTICS DATA
if APPLY_STANDARD_SCALING:
standart_scaler_instance = StandardScaler()
psd_features_transformed[psd_features_columns] = standart_scaler_instance.fit_transform(psd_features_transformed[psd_features_columns])
if APPLY_MIN_MAX_SCALING:
psd_features_transformed[psd_features_columns] = min_max_scaler_instance.fit_transform(psd_features_transformed[psd_features_columns])
```
### Construct dataset for modeling
```
## CREATE FINAL DATASET ##
## CHOOSE WHICH FEATURES TO INCLUDE FOR THE FINAL DATASET
## BY DEFAULT, WE WILL MODEL ALL FEATURES THAT WE’VE CREATED
## HOWEVER, WE CAN MAKE MODELS OUTPUT EASIER TO INTERPRET IF WE MODEL INFORMATION ONLY FOR ONE RESOURCE TYPE (CPU or NETWORK)
## To include columns that contains cpu_run
## descriptive_statistics_transformed.filter(regex = 'cpu_run')
## To exclude columns containing cpu_run string
## descriptive_statistics_transformed.loc[:,~descriptive_statistics_transformed.columns.str.contains('cpu_run')]
final_data_list = [
descriptive_statistics_transformed
,correlation_data_transformed
,virt_hw_transformed
,psd_features_transformed
]
## To include columns that contains cpu_urn
## psd_features_transformed.filter(regex = 'cpu_run')
## To exclude columns containing cpu_run string
## psd_features_transformed.loc[:,~psd_features_transformed.columns.str.contains('cpu_run')]
## Init empty data frame:
final_modeling_data = pd.DataFrame(index = list(perf_data.keys()))
for i in final_data_list:
data = i.copy()
final_modeling_data = pd.merge(final_modeling_data, data, left_index=True, right_index=True)
##
print(final_modeling_data.shape)
```
### Inspect linear correlation and PC
```
### Inspect correlation:
DATA = final_modeling_data
cmap = sns.diverging_palette(220, 20, sep=20, as_cmap=True)
fig = plt.figure()
sns.clustermap(DATA.corr(), figsize = (20, 20), cmap = cmap)
plt.show()
### Inspect PCA:
DATA = final_modeling_data
## One more thing to consider is the correlation among the variables:
pca_instance = PCA().fit(DATA)
## Taking care to preserve the index for later join:
pca_data = pd.DataFrame(pca_instance.transform(DATA), index = DATA.index)
fig, ax = plt.subplots(figsize=(20,8),nrows = 1 , ncols = 2)
sns.scatterplot(x = DATA.iloc[:,0], y = pca_data.iloc[:,1], ax = ax[0])
ax[0].set_title('Principal Components')
ax[0].set_xlabel("PC 1")
ax[0].set_ylabel("PC 2")
#tmp_plt = sns.lineplot( x= [i+1 for i in range(pca_instance.n_components_)]
# , y =pca_instance.explained_variance_
# , ax = ax[1]
# #, color = 'blue'
# )
ax[1].bar(x = [i+1 for i in range(pca_instance.n_components_)], height = pca_instance.explained_variance_/np.sum(pca_instance.explained_variance_))
ax2 = ax[1].twinx()
ax2.plot([i+1 for i in range(pca_instance.n_components_)], np.cumsum(pca_instance.explained_variance_)/ np.sum(pca_instance.explained_variance_), c = 'orange')
ax2.grid([])
ax2.set_ylim(bottom =0)
ax[1].set_title('Explained variance plot:')
ax[1].set_xlabel('Number of principal components')
ax[1].set_ylabel('Marginal variance')
ax2.set_ylabel('Cumulative variance')
##
```
## Uniform Manifold Approximation and Projection (UMAP)
**Resources**
[UMAP Python Manual](https://umap-learn.readthedocs.io/en/latest/)
[Understanding UMAP](https://pair-code.github.io/understanding-umap/)
[How Exactly UMAP Works](https://towardsdatascience.com/how-exactly-umap-works-13e3040e1668)
### Basic Umap Usage
```
import umap
## umap usage:
DATA = final_modeling_data
# 1. We create instance of the object
# Here we set the parameters that are internal to the algorithm
UMAP_Instance = umap.UMAP(n_components = 2 ## The dimension of the space to embed into.
, min_dist = 0.5 ## The effective minimum distance between embedded points. Smaller values will result in a more clustered/clumped embedding
, spread = 2.0 ## The effective scale of embedded points. In combination with min_dist this determines how clustered/clumped the embedded points are.
, n_neighbors = 15 ## The size of local neighborhood (in terms of number of neighboring sample points) used for manifold approximation.
, metric = 'euclidean' ## The metric to use to compute distances in high dimensional space. (euclidean, manhattan, cosine, )
,learning_rate = 100 ## Controls the step during the gradient descent stage
,n_epochs = int(1e3) ## selected based on the size of the input dataset (200 for large datasets, 500 for small)
#,random_state = 500 ## Random seed
,verbose = 2 ## controls logging of the algorithm
,a = None ## a and b are another way to control min dist and spread
,b = None ##
)
# 2.Fit the object to the data
result = UMAP_Instance.fit_transform(DATA)
# The result of fit_transform is ndarray with shape (NSamples, n_components)
plt.figure(figsize=(8,8))
plt.scatter(result[:, 0], result[:, 1])
plt.xlabel('X1')
plt.ylabel('X2');
```
### UMAP parameter tuning
</br></br>
**Resources**
[Basic UMAP Parameters](https://umap-learn.readthedocs.io/en/latest/parameters.html)
[Fine-tuning UMAP Visualizations](https://scikit-learn.org/stable/modules/generated/sklearn.manifold.TSNE.html)
```
## EXPERIMENT WITH DIFFERENT UMAP SETTINGS ##
TUNABLE_PARAMETERS = {'n_neighbors':[10,20,30], 'min_dist':[.1,.2,.3], 'spread':[.1,.2,.3]}
DEFAULT_PARAMETERS = {}
APPLY_PCA = False
N_PC = []
MODEL = 'UMAP'
DATA = final_modeling_data
## Apply tuning:
UMAP_results = model_train( data = DATA
, model = MODEL
, param_dict = TUNABLE_PARAMETERS
, apply_pca = APPLY_PCA
, n_pc = N_PC
, other_params = DEFAULT_PARAMETERS
)
## Inspect the results of the parameters
################
## PARAMETERS ##
################
RESULTS = UMAP_results
LEADING_PARAM = 'n_neighbors'
COLOR_ARRAY = None
FORCE_CATEGORICAL = False
POINT_SIZE = 5
## APPLY FUNCTION
plot_results(results = RESULTS
,leading_param = LEADING_PARAM
,color_array = COLOR_ARRAY
,force_categorical = FORCE_CATEGORICAL
,point_size = POINT_SIZE
)
## If single combination is being expected
##plot_embedding(RESULTS[0], fig_scale= 1.2,point_size = 5)
```
### Apply DBSCAN
```
## Choose one embedding to operate with
## Zoom plot in :
RESULTS = UMAP_results
MODEL_INDEX = 2
###
plot_embedding(RESULTS[MODEL_INDEX], fig_scale = 1.5, point_size = 4)
## Capture the clusters with dbscan
TUNABLE_PARAMETERS = {'eps': [.4, .8,1.2,1.4]
, 'min_samples':[10, 26,28]}
DEFAULT_PARAMETERS = {}
MODEL = 'DBSCAN'
## Apply tuning:
dbscan_clusters = model_train( data = RESULTS[MODEL_INDEX][1]
, model = MODEL
, param_dict = TUNABLE_PARAMETERS
, other_params = DEFAULT_PARAMETERS
)
## Plot the results from DBSCAN clustering
result_for_plotting = [(i[0],RESULTS[MODEL_INDEX][1],i[2]) for i in dbscan_clusters]
dbscan_clusters_results = [i[1] for i in dbscan_clusters]
plot_results(results = result_for_plotting, color_array=dbscan_clusters_results, force_categorical=True, point_size=5)
## Zoom one of the plots:
RESULTS = UMAP_results
CLUSTER_INDEX = 9
####
plot_embedding(RESULTS[MODEL_INDEX]
, fig_scale = 2
, color_var_list = [dbscan_clusters[CLUSTER_INDEX][1]]
, force_categorical=True
, plot_centers=True
, point_size = 5)
```
### Profile clusters
```
## PARAMETERS ##
TRANSFORMATION = return_transformer(None)
#DATA = virt_hw.loc[:, virt_hw.columns != 'os_fam'] ## it is categorical variable
DATA = descriptive_statistics_transformed
########
## Plot the distribution as boxplot
clustering_df = pd.DataFrame(index = final_modeling_data.index)
clustering_df['dbscan_clusters'] = dbscan_clusters[CLUSTER_INDEX][1]
data = DATA.copy()
data = TRANSFORMATION(data)
data = pd.merge(data,clustering_df, left_index=True, right_index=True)
data = pd.melt(data, id_vars = ['dbscan_clusters'])
ordr = list(sorted(set(dbscan_clusters[CLUSTER_INDEX][1])))
g = sns.FacetGrid(data, col = 'variable', col_wrap = 5, sharex = False, sharey=False, height= 5)
g.map(sns.boxplot, 'dbscan_clusters', 'value').add_legend()
plt.show()
## Parameters:
DATA = virt_hw# non_temporal_features/ virt_hw_transformed
COLOR_VAR = 'number_of_nics'
TRANSFORMATION = return_transformer(None)
RESULTS = UMAP_results
###
plot_embedding( RESULTS[MODEL_INDEX]
, fig_scale = 2
, color_var_list = [TRANSFORMATION(DATA[COLOR_VAR].values)]
, point_size = 20
, force_categorical = True
)
## Utility code that helps us compare the embedding generated from pca to the embedding
#pca_result_construct = ({}, first_stage_data_pca.iloc[:,0:5].values,{})
#plot_embedding(pca_result_construct, fig_scale=.8, color_var_list = [TRANSFORMATION(DATA[COLOR_VAR].values)])
## PARAMETERS ##
clusters_to_compare = [1,4]
samples_per_cluster = 5
metric = 'net_packetsTx'
DATA = perf_data
### Fig scaling:
num_cl_to_compare = len(clusters_to_compare)
####
sampled_df = clustering_df[clustering_df['dbscan_clusters'].isin(clusters_to_compare)].groupby('dbscan_clusters').apply(lambda x: x.sample(samples_per_cluster))
fig, axes = plt.subplots(nrows = samples_per_cluster, ncols = num_cl_to_compare, figsize = (15*num_cl_to_compare, 5*samples_per_cluster))
ix_i = 0
for i in clusters_to_compare:
_tmp = list(sampled_df.iloc[sampled_df.index.get_level_values(0).isin([i]),:].index.get_level_values(1))
ix_j = 0
for j in _tmp:
axes[ix_j, ix_i].plot(DATA[j][metric])
axes[ix_j, ix_i].set_ylabel('vmid:'+str(j) + '('+str(i)+')')
axes[ix_j, ix_i].yaxis.set_label_position('right')
ix_j += 1
ix_i += 1
```
| github_jupyter |
<font size="+5">#09. Cluster Analysis con k-Means</font>
- Book + Private Lessons [Here ↗](https://sotastica.com/reservar)
- Subscribe to my [Blog ↗](https://blog.pythonassembly.com/)
- Let's keep in touch on [LinkedIn ↗](www.linkedin.com/in/jsulopz) 😄
# Machine Learning Review
- **Supervised Learning:**
- Regression → Predicting a Numerical Variable
- Classification → Predicting a Categorical Variable
- **Unsupervised Learning:**
- Cluster Analysis → Groups based on Explanatory Variables
# Load the Data
> - Simply execute the following lines of code to load the data.
> - This dataset contains **statistics about Car Accidents** (columns)
> - In each one of **USA States** (rows)
https://www.kaggle.com/fivethirtyeight/fivethirtyeight-bad-drivers-dataset/
```
import seaborn as sns
df = sns.load_dataset(name='car_crashes', index_col='abbrev')
df.sample(5)
```
# `KMeans()` Model in Python
## Build the Model
> 1. **Necesity**: Build Model
> 2. **Google**: How do you search for the solution?
> 3. **Solution**: Find the `function()` that makes it happen
## Code Thinking
> Which function computes the Model?
> - `fit()`
>
> How could can you **import the function in Python**?
### Separate Variables for the Model
> Regarding their role:
> 1. **Target Variable `y`**
>
> - [ ] What would you like **to predict**?
>
> Total number of accients? Or Alcohol?
>
> 2. **Explanatory Variable `X`**
>
> - [ ] Which variable will you use **to explain** the target?
### Data Visualization to Analyize Patterns
> - Visualize the 2 variables with a `scatterplot()`
> - And decide *how many `clusters`* you'd like to calculate
### Finally `fit()` the Model
## `predict()` the Cluster for One `USA State`
> **Programming thiking:**
>
> - Which `function()` can we use to make a prediction?
> - How can you answer yourself **without searching in Google**?
## Get the `cluster` for all USA States
> - `model.` + `↹`
> - Create a `dfsel` DataFrame
> - That contains the **columns you used for the model**
> - Add a **new column**
> - That **contains the `cluster` prediction** for every USA State
```
df['cluster'] = ?
```
## Model Visualization
> - You may `hue=` the points with the `cluster` column
## Model Interpretation
> - Do you think the model **makes sense**?
> - Which **variable is the most important** to determine the cluster?
```
%%HTML
<iframe width="560" height="315" src="https://www.youtube.com/embed/4b5d3muPQmA" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
```
# Conclusion
> - You need to `scale` the data
> - Every time the algorithm computes `distances`
> - Between `different variables`
> - Because it's **not the same to increase 1kg of weight than 1m of height**
```
# Draw Weight Height Axes Min Max Scaler 0 - 1
```
# `MinMaxScaler()` the data
> - `scaler.fit_transform()`
# `KMeans()` Model with *Scaled Data*
# Other `Clustering` Models in Python
> - Visit the `sklearn` website [here ↗](https://scikit-learn.org/stable/)
> - **Pick 2 new models** and compute the *Clustering*
## Other Model 1
## Other Model 2
| github_jupyter |
# Portfolio Optimization
-------------------------
## Portfolio Risk-Return optimization based on the Markowitz's Efficient Frontier and CVaR
-------------------------
```
'''
documentation: https://pyportfolioopt.readthedocs.io/en/stable/
'''
# Import libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from pypfopt import EfficientFrontier
from pypfopt import risk_models
from pypfopt import expected_returns
from pypfopt import plotting
%matplotlib inline
# Read in price (Adj Close) data from csv file (Adj Close prices pulled from the yahoo_historical.ipynb)
stock_prices = pd.read_csv("../Resources/multi_stock_prices.csv", parse_dates=True, index_col="Date")
stock_prices.tail()
from pypfopt.risk_models import CovarianceShrinkage
# Calculate expected returns (CAPM based) and covariance (Ledoid Wolf covariance shrinkage)
mu = expected_returns.capm_return(stock_prices)
S = CovarianceShrinkage(stock_prices).ledoit_wolf()
# Print expected returns
mu
# Specify allocation constraints for each sector or individual security to see different portfolio risk-return characteristics.
sector_mapper = {
"ETH-USD": "crypto",
"AGG": "bonds",
"BRK-B": "value",
"ARKK": "growth"
}
sector_lower = {
"value": 0.40, # min 40% equity
"bonds": 0.10, # min 10% bonds
"growth": 0.40, # min 40% growth
"crypto": 0.02 # min 2% crypto
}
sector_upper = {
"crypto": 0.02, # less than 2% crypto
"value": 0.70, # less than 70% value
"growth": 0.40, # less than 40% growth
"bonds": 0.11 # less than 11% bonds
}
# Construct Efficient Frontier optimization model based on the maximum Sharpe ratio (alternatively we can optimize by minimum volatility which is a good option too - by entering w=ef.min_volatility()) and specify gamma to minimize zero weights (higher gamma means less zero weights) and sector allocation constraints
from pypfopt import objective_functions
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.L2_reg, gamma=0.1)
ef.add_sector_constraints(sector_mapper, sector_lower, sector_upper)
w = ef.max_sharpe()
weights = ef.clean_weights()
print(ef.clean_weights())
# Plot asset allocation based on Efficient Frontier model
pd.Series(weights).plot.bar(figsize=(10, 8), ylabel='%', xlabel ='Assets',title='Asset Allocation')
# Evaluate portfolio performance
ef.portfolio_performance(verbose=True)
# Plot Efficient Frontier based on random portfolios and individual assets
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.L2_reg, gamma=0.1)
ef.add_sector_constraints(sector_mapper, sector_lower, sector_upper)
fig, ax = plt.subplots()
plotting.plot_efficient_frontier(ef, ax=ax, show_assets=True)
# Find the tangency portfolio
ef.max_sharpe()
ret_tangent, std_tangent, _ = ef.portfolio_performance()
ax.scatter(std_tangent, ret_tangent, marker="*", s=100, c="r", label="Max Sharpe")
# Generate random portfolios
n_samples = 10000
w = np.random.dirichlet(np.ones(len(mu)), n_samples)
rets = w.dot(mu)
stds = np.sqrt(np.diag(w @ S @ w.T))
sharpes = rets / stds
ax.scatter(stds, rets, marker=".", c=sharpes, cmap="viridis_r")
# Output
ax.set_title("Efficient Frontier with random portfolios")
ax.legend()
plt.tight_layout()
plt.savefig("../Resources/ef_scatter.png", dpi=200)
plt.show()
```
## Portfolio Optimization with CVaR
-------------------------------------
### Conditional Value at Risk (CVaR), also known as the expected shortfall, is a risk assessment measure that quantifies the amount of tail risk an investment portfolio has
```
from pypfopt import EfficientCVaR
# Calculate expected returns (mu) and covariance (risk)
mu = expected_returns.capm_return(stock_prices)
S = CovarianceShrinkage(stock_prices).ledoit_wolf()
# Portfolio optimization based on Efficient Frontier and maximum Sharpe ratio
ef = EfficientFrontier(mu, S)
ef.add_objective(objective_functions.L2_reg, gamma=0.1)
ef.add_sector_constraints(sector_mapper, sector_lower, sector_upper)
ef.max_sharpe()
weights_arr = ef.weights
ef.portfolio_performance(verbose=True)
print(f"weights = {weights_arr}")
# Calculate stock returns from historical stock prices
returns = expected_returns.returns_from_prices(stock_prices).dropna()
returns
# Calculate portfolio returns and plot probability distribution (normal distribution)
portfolio_rets = pd.DataFrame(returns * weights_arr).sum(axis=1)
portfolio_rets.hist(bins=50)
# Calculate 95% CVaR for the max-sharpe portfolio
var = portfolio_rets.quantile(0.05)
cvar = portfolio_rets[portfolio_rets <= var].mean()
print("VaR: {:.2f}%".format(100*var))
print("CVaR: {:.2f}%".format(100*cvar))
print(f"The average loss on the worst 5% of days will be {cvar :.4f}")
# Construct a portfolio with the minimum CVar, print performance and weights
ec = EfficientCVaR(mu, returns)
ec.add_objective(objective_functions.L2_reg, gamma=0.1)
ec.add_sector_constraints(sector_mapper, sector_lower, sector_upper)
ec.min_cvar()
ec.portfolio_performance(verbose=True)
ec.weights
# Another option is to maximize return for the given CvaR (set a % CvaR that is acceptable, however make sure you understand what CvaR means). Print performance and weights
ec = EfficientCVaR(mu, returns)
ec.add_objective(objective_functions.L2_reg, gamma=0.1)
ec.add_sector_constraints(sector_mapper, sector_lower, sector_upper)
ec.efficient_risk(target_cvar=0.05)
ec.portfolio_performance(verbose=True)
ec.weights.round(2)
# Original portfolio performance and weights based on max Sharpe ratio
ef.portfolio_performance(verbose=True)
ef.weights.round(2)
```
| github_jupyter |
```
import spacy
nlp=spacy.load('en_core_web_sm')
def remove_stop_punct(s):
from nltk.corpus import stopwords
stopwords=stopwords.words('english')
doc=nlp(s)
#k=sorted([".","DT","TO","CC","IN"])
#st=" ".join(t.text for t in doc if (t.pos_ not in ["PUNCT"] )and(t.text.lower() not in stopwords))
st=" ".join(t.text for t in doc if (t.pos_ not in ["PUNCT"] ))
return st.strip()
f=open('/home/judson/Desktop/sentenceSeg/POS_IntraSent/decision_POS.txt',"a")
f1=open('/home/judson/Desktop/sentenceSeg/Labelled/decision.txt',"r")
txt=f1.readlines()
for i in txt[374:]:
s=remove_stop_punct(i)
t=nlp(s)
pos=[i.tag_ for i in t]
f.writelines(s+"\t"+",".join(pos)+"\t<DECISION>\n")
f1=open('/home/judson/Desktop/sentenceSeg/Labelled/arguments.txt',"r")
txt=f1.readlines()
import spacy
nlp=spacy.load('en_core_web_sm')
def remove_stop_punct(s):
from nltk.corpus import stopwords
stopwords=stopwords.words('english')
doc=nlp(s)
#k=sorted([".","DT","TO","CC","IN"])
#st=" ".join(t.text for t in doc if (t.pos_ not in ["PUNCT"] )and(t.text.lower() not in stopwords))
st=" ".join(t.text for t in doc if (t.pos_ not in ["PUNCT"] ))
return st.strip()
#print remove_stop_punct(s)
def dict_conv(st):
doc=nlp(st)
root=[]
for i in doc:
if i.dep_=="ROOT":
root.append(i.i)
d={tok.i:[tok.text,tok.dep_,tok.head.text,tok.tag_,[child.i for child in tok.children]] for tok in doc}
return [d,root]
import networkx
def dfs(sd, start, end = "$$"):
h=networkx.Graph(sd)
ptr=[]
for pth in networkx.all_simple_paths(h,start,"$$"):
ptr.append(pth[:-1])
return ptr
def parse_depend(dt,root):
#md= {i:j[3] for i,j in dt.items()}
#print dt
sd=dict()
sd["$$"]=[]
for i,j in dt.items():
if j[4]==[]:
sd[i]=["$$"]
else:
sd[i]=[k for k in j[4]]
#print sd
#m=""
#for i,j in dt.items():
# if j[0]=="ROOT":
# print j[0]
# m=str(i)
# break
#print m
#print sd
ml=[]
for i in root:
ml.extend(dfs(sd,i,"$$"))
#print ml
mpos=[[dt[j][3] for j in i]for i in ml]
mdep=[[dt[j][1] for j in i]for i in ml]
return mdep,mpos
#input nested string list output string
def To_str(nlst):
st="("
for i in nlst:
st=st+"("
st=st+",".join(i)
st=st+"),"
return st[:-1]+")"
f=open('/home/judson/Desktop/sentenceSeg/Dep_IntraSent/Dep_decision',"a")
f1=open('/home/judson/Desktop/sentenceSeg/Labelled/decision.txt',"r")
decision_pickle=[]
txt=f1.readlines()
for i in txt:
s=remove_stop_punct(i)
#print s
#t=nlp(s)
dep,_=parse_depend(*dict_conv(s))
decision_pickle.append(dep)
st=To_str(dep)
#f.writelines(s+"\t"+st+"\t<DECISION>\n")
#print (s+"\t"+st+"\t<ARGUMENTS>\n")
f=open('/home/judson/Desktop/sentenceSeg/Dep_POS_IntraSent/dep_pos_decision.txt',"a")
f1=open('/home/judson/Desktop/sentenceSeg/Labelled/arguments.txt',"r")
txt=f1.readlines()
for i in txt:
s=remove_stop_punct(i)
#print s
#t=nlp(s)
dep,pos=parse_depend(*dict_conv(s))
st=To_str(dep)
st1=To_str(pos)
f.writelines(s+"\t"+st+"\t"+st1+"\t<DECISION>\n")
#find unique nested list
#input ip_list->nested list
#output uniq_list->nested list
def uniq_list(ip_list):
uniq_list=[ip_list[0]]
for i in ip_list:
k=0
for j in uniq_list:
if j==i:
k=0
break
else:
k=1
if k==1:
uniq_list.append(i)
return uniq_list
import pickle
f=open("decision.pickle","wb")
pickle.dump(uniq_list(decision_pickle),f,protocol=2)
def mod_parse_depend(dt,root):
#md= {i:j[3] for i,j in dt.items()}
#print dt
sd=dict()
sd["$$"]=[]
for i,j in dt.items():
if j[4]==[]:
sd[i]=["$$"]
else:
sd[i]=[k for k in j[4]]
ml=[]
for i in root:
ml.extend(dfs(sd,i,"$$"))
#print ml
mpos=[[dt[j][3] for j in i]for i in ml]
mdep=[[dt[j][1] for j in i]for i in ml]
mname=[[dt[j][0] for j in i]for i in ml]
return mdep,mpos,mname
p,d,n=mod_parse_depend(*dict_conv("Apple is looking at buying U.K.startup for $1 billion"))
def mod_file_write(p,d,n):
mn=[[] for _ in range(len(p))]
for i in range(len(p)):
mn[i]=[(p[i][j],n[i][j])for j in range(len(p[i]))]
st="["
for i in mn:
st=st+"["
for j ,k in i:
st=st+"("+j+","+k+"),"
st=st[:-1]+"],"
st=st[:-7]+"]]"
return st+"\n"
f=open("/home/judson/Desktop/sentenceSeg/Labelled1/decision.txt","r")
#f1=open("/home/judson/LabelledDepparsed.txt","a")
id_r=f.readlines()
id_r=id_r[:4]+id_r[5:6]
f1.writelines("Decision:\n")
for i in id_r:
f1.writelines(str(i+":"+mod_file_write(*mod_parse_depend(*dict_conv(i)))))
```
| github_jupyter |
# Bootstrapping (Nonparametric Inference)
In statistics, bootstrapping is any test or metric that relies on random sampling with replacement. Bootstrapping allows assigning measures of accuracy (defined in terms of bias, variance, confidence intervals, prediction error or some other such measure) to sample estimates. This technique allows estimation of the sampling distribution of almost any statistic using random sampling methods. Generally, it falls in the broader class of resampling methods.
Bootstrapping is the practice of estimating properties of an estimator (such as its variance) by measuring those properties when sampling from an approximating distribution. One standard choice for an approximating distribution is the empirical distribution function of the observed data. In the case where a set of observations can be assumed to be from an independent and identically distributed population, this can be implemented by constructing a number of resamples with replacement, of the observed data set (and of equal size to the observed data set).
It may also be used for constructing hypothesis tests. It is often used as an alternative to statistical inference based on the assumption of a parametric model when that assumption is in doubt, or where parametric inference is impossible or requires complicated formulas for the calculation of standard errors.
- Random sampling with replacement*
- Resampling technique to simulate drawing new samples (where repeating experiments is not feasible or possible)
- Typically, the new sample has size *n*, where *n* is the size of the original dataset
```
%matplotlib inline
import pandas as pd
import numpy as np
import scipy.stats
import matplotlib.pyplot as plt
```
## Sample mean, standard error of the mean, and estimating the population mean
```
df = pd.read_csv('../datasets/iris/iris.csv')
x = df['sepal_length'].values
x_mean = np.mean(x)
plt.figure(figsize=(30,15))
plt.hist(x, bins=10)
plt.axvline(x_mean, color='orange', label='sample mean: %.2f' % x_mean)
plt.xlabel('sepal length in cm')
plt.ylabel('count')
plt.legend(loc=1)
plt.show()
```
#### Standard Error (SE)
$$SE_{\bar{x}} = \frac{s}{\sqrt{n}}$$
- the standard error *SE* (or *standard error of the mean*) estimates the standard deviation (*s*) of the sample mean ($\bar{x}$)
- i.e., the *SE* measures the variability when taking different samples from the population
- in other words, the *SE* measures the variability between samples, whereas the sample standard deviation measures the variability within a sample
- we use the standard error to judge how "good" our estimate of the population mean ($\mu$) is
```
se = np.std(x, ddof=1) / np.sqrt(x.shape[0])
print('standard error', se)
scipy.stats.sem(x)
```
#### Bootstrapping and estimating the population mean
```
def bootstrap_means(x, n_bootstrap_samples, seed=None):
rng = np.random.RandomState(seed)
sample_means = np.zeros(shape=n_bootstrap_samples)
for i in range(n_bootstrap_samples):
boot_sample = rng.choice(x, size=x.shape[0], replace=True)
# replicate is a general term for a statistic computed
# from a bootstrap sample
bootstrap_replicate = np.mean(boot_sample)
sample_means[i] = bootstrap_replicate
return sample_means
boot_50 = bootstrap_means(x, n_bootstrap_samples=50, seed=123)
boot_mean = np.mean(boot_50)
plt.figure(figsize=(30,15))
plt.hist(boot_50, bins=10)
plt.axvline(boot_mean, color='orange', label='samples mean: %.2f' % boot_mean)
plt.xlabel('mean sepal length in cm')
plt.ylabel('count')
plt.legend(loc=2)
plt.show()
boot_2500 = bootstrap_means(x, n_bootstrap_samples=2500, seed=123)
boot_mean = np.mean(boot_2500)
plt.figure(figsize=(30,15))
plt.hist(boot_2500, bins=15)
plt.axvline(boot_mean, color='orange', label='samples mean: %.2f' % boot_mean)
plt.xlabel('mean sepal length in cm')
plt.ylabel('count')
plt.legend(loc=2)
plt.show()
```
- note: no matter how the sample is distributed, the sample mean follows a normal distribution
```
np.std(boot_2500, ddof=1)
```
- remember, the standard deviation of the bootstrap replicates (means) estimates the standard error of the mean (which estimates the standard deviation of the population mean)
```
se = np.std(x, ddof=1) / np.sqrt(x.shape[0])
print('standard error', se)
def empirical_cdf(sample):
x = np.sort(sample)
y = np.arange(1, x.shape[0] + 1) / x.shape[0]
return x, y
ecdf_x, ecdf_y = empirical_cdf(boot_2500)
plt.figure(figsize=(30,15))
plt.scatter(ecdf_x, ecdf_y)
plt.xlabel('mean')
plt.ylabel('CDF')
```
## Confidence Intervals
- 95% confidence interval: 95% of the sample means (if we would draw new samples / repeat the experiments) would fall within the confidence interval
#### From bootstrap replicates:
```
boot_2500 = bootstrap_means(x, n_bootstrap_samples=2500, seed=123)
lower, upper = np.percentile(boot_2500, [2.5, 97.5])
print('95%% confidence interval: [%.2f, %.2f]' % (lower, upper))
```
#### From the original data (i.e., from a single sample):
```
def confidence_interval(x, ci=0.95):
x_mean = np.mean(x)
se = np.std(x, ddof=1) / np.sqrt(x.shape[0])
h = se * scipy.stats.t._ppf((1 + ci)/2. , x.shape[0])
return x_mean - h, x_mean + h
lower, upper = confidence_interval(x, ci=0.95)
print('95%% confidence interval: [%.2f, %.2f]' % (lower, upper))
se = np.std(x, ddof=1) / np.sqrt(x.shape[0])
lower, upper = scipy.stats.norm.interval(alpha=0.95,
loc=np.mean(x),
scale=se)
print('95%% confidence interval: [%.2f, %.2f]' % (lower, upper))
```
| github_jupyter |
# Example: CanvasXpress gantt Chart No. 4
This example page demonstrates how to, using the Python package, create a chart that matches the CanvasXpress online example located at:
https://www.canvasxpress.org/examples/gantt-4.html
This example is generated using the reproducible JSON obtained from the above page and the `canvasxpress.util.generator.generate_canvasxpress_code_from_json_file()` function.
Everything required for the chart to render is included in the code below. Simply run the code block.
```
from canvasxpress.canvas import CanvasXpress
from canvasxpress.js.collection import CXEvents
from canvasxpress.render.jupyter import CXNoteBook
cx = CanvasXpress(
render_to="gantt4",
data={
"y": {
"vars": [
"Start",
"End"
],
"smps": [
"S01",
"S02",
"S03",
"S04",
"S05",
"S06",
"S07",
"S08",
"S09",
"S10",
"S11",
"S12"
],
"data": [
[
20140221,
20140521,
20140821,
20141121,
20150221,
20150521,
20140821,
20141121,
20150221,
20150521,
20150821,
20151121
],
[
20140520,
20140820,
20141120,
20150220,
20150520,
20150820,
20141120,
20150220,
20150520,
20150820,
20151120,
20160220
]
]
},
"x": {
"Clinical Trial": [
"CT001",
"CT002",
"CT003",
"CT004",
"CT005",
"CT006",
"CT001",
"CT002",
"CT003",
"CT004",
"CT005",
"CT006"
],
"Indication": [
"Lung",
"Liver",
"Breast",
"Skin",
"Lung",
"Lung",
"Breast",
"Pancreas",
"Stomach",
"Breast",
"Skin",
"Lung"
],
"Completion": [
0.2,
0.6,
1,
0.4,
0.3,
0.4,
0.25,
1,
0.8,
0.9,
0.6,
0.3
],
"Dependencies": [
None,
None,
None,
None,
None,
None,
"S01",
"S02",
"S03",
None,
None,
None
]
}
},
config={
"blockContrast": True,
"colorBy": "Indication",
"ganttCompletion": "Completion",
"ganttDependency": "Dependencies",
"ganttEnd": "End",
"ganttStart": "Start",
"graphType": "Gantt",
"theme": "CanvasXpress"
},
width=613,
height=613,
events=CXEvents(),
after_render=[],
other_init_params={
"version": 35,
"events": False,
"info": False,
"afterRenderInit": False,
"noValidate": True
}
)
display = CXNoteBook(cx)
display.render(output_file="gantt_4.html")
```
| github_jupyter |
# Assignment 2
The main idea of this assignment is to understand **Convolutional Neural Networks** and the basics of **image filtering**. We will implement matrix convolution as well as the convolutional layer from scratch.
**Please note:**
1. Please copy the code from the previous assignment (week 2) into a separate file `Blocks.py`. Make sure it resides in the same folder as this notebook. It should contain the implementation of the building blocks.
2. All functions should be implemented using [**NumPy**](https://docs.scipy.org/doc/).
## Table of contents
* [0. Before you begin](#0.-Before-you-begin)
* [1. Recap](#1.-Recap)
* [2. Matrix Convolution](#2.-Matrix-Convolution) (mandatory)
* [3. Basic Kernels](#3.-Basic-Kernels) (mandatory)
* [4. Convolutional Layer](#4.-Convolutional-Layer)
* [5. MaxPooling Layer](#5.-MaxPooling-Layer) (mandatory)
* [6. Flatten](#6.-Flatten)
* [7. Experiments](#7.-Experiments)
# 0. Before you begin
Run the following block once when you start this notebook.
It imports [NumPy](https://docs.scipy.org/doc/), [Matplotlib](https://matplotlib.org/), AutoMark, and your `Blocks.py` implementations from last week.
We will make use of these imports during the course of this notebook.
```
import numpy as np
import matplotlib.pyplot as plt
# previously implemented blocks for nn
from Blocks import *
import automark as am
#%matplotlib inline
# fill in your studentnumber as your username again
username = '11739371'
# get your progress by running this function
am.get_progress(username)
```
# 1. Recap
During the previous assignment, you implemented the main building blocks of neural networks:
* **Dense Layer**
* Non-linearity with ReLU
* Loss functions
* Optimization
Dense layers can already be very useful. They perform the following mapping on the input matrix $X$ (matrix of objects) using the weights $W$:
$$
H = XW + b
$$
The dense layer enables creation and training of flexible models.
Now, let's take a look at image processing using **Dense Layer**:
1. We have a grayscale image $x$ of size $N \times M$ (width & height)
2. We reshape it into a vector of length $NM$
3. Then we map it with a dense layer
4. And obtain the transformed vector $y$
Each element of $y$ depends on each element of $x$. That's why its also called **fully connected**.
A neural network which only consists of dense layers is therefore called a fully connected network.
When we work with images, we assume that each pixel is correlated with its neighbours and close pixels. Distant pixels are not correlated. Various experiments demonstrate that this assumption is correct.
Dense layers captures these correlations, but they also capture *noisy* correlations to all other pixels.
# 2. Matrix Convolution
There is a way to create a **locally connected** layer which will learn local correlations using a smaller amount of parameters.
This layer is aptly called **Convolutional Layer** and is based on **matrix convolution**
A picture is worth a thousand words which is especially true when learning about convolution:

In image convolution, a **filter**, also called **kernel**, is applied to the source matrix.
Each element from the kernel is multiplied by the corresponding element from the source matrix. The results are summed up and written to the target matrix.
In this example, the output matrix has a smaller size than its source\*. This is because the kernel can not overlap the borders. **Zero padding** can be used to retain the original dimension. It is a simple solution which involves adding a border of zeros to the input.
\* It may seem both matrices have the same size (both are shown with the same number of boxes. In the edges of the right matrix, however, no values are stored. The top-left corner of the right image starts where the $-3$ is placed.
The source matrix $X$ is of size $N \times M$ and the kernel $K$ is of size $(2p+1) \times (2q +1 )$.
We define $X_{ij} = 0$ for $i > N, i < 1$ and $j > M, j < 1$.
In (other) words: If you try to access a pixel which is out of bounds assume that it is zero.
This is called **zero padding**.
Therefore, the convolution of a matrix with a kernel is defined as follows:
$$
Y = X \star K \\
Y_{ij} = \sum\limits_{\alpha=0}^{2p} \sum\limits_{\beta=0}^{2q}
K_{\alpha \beta} X_{i + \alpha - p, j+\beta - q}
$$
This operation's name depends on the field:
* In machine learning: **convolution**
* In mathematics: **cross-correlation**
Finally, its time for you to implement matrix convolution.
You can use the example below this code block to test your implementation.
```
def conv_matrix(matrix, kernel):
"""Perform the convolution of the matrix
with the kernel using zero padding
# Arguments
matrix: input matrix np.array of size `(N, M)`
kernel: kernel of the convolution
np.array of size `(2p + 1, 2q + 1)`
# Output
the result of the convolution
np.array of size `(N, M)`
"""
n, m = matrix.shape
a, b = kernel.shape
p, q = (a - 1) // 2, (b - 1) // 2
# create array of 0
matrix_padded = np.pad(matrix, ((p, p),(q, q)), 'constant')
# insert the image after zero's padding
matrix_padded[p:p + n, q:q + m] = matrix
output = np.zeros_like(matrix)
for j in range(m):
for i in range(n):
output[i, j] = (
kernel * matrix_padded[i:i + 2 * p + 1, j:j + 2 * q + 1]).sum()
return output
```
Let's test the function with the following data:
$$
X = \begin{bmatrix}
1 & 2 & 3 \\
2 & 3 & 4 \\
3 & 4 & 5 \\
\end{bmatrix} \quad
K =
\begin{bmatrix}
1 & 0 & 0 \\
0 & 1 & 0 \\
0 & 0 & 2 \\
\end{bmatrix} \quad
X \star K =
\begin{bmatrix}
7 & 10 & 3 \\
10 & 14 & 6 \\
3 & 6 & 8 \\
\end{bmatrix}
$$
We recreate the example data in Python to perform a local test run.
Don't be confused by [np.eye](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.eye.html). It fills our kernel matrix with ones on the diagonal from top-left to bottom-right.
```
X = np.array([
[1, 2, 3],
[2, 3, 4],
[3, 4, 5]
])
K = np.eye(3)
K[-1, -1] = 2
print(np.zeros(3))
print(K)
```
Run the following code block and compare the result with the example above.
```
print(conv_matrix(X, K))
```
If you feel confident with your solution, check it against the AutoMark-server.
```
am.test_student_function(username, conv_matrix, ['matrix', 'kernel'])
```
# 3. Basic Kernels
Matrix convolution can be used to process an image (think Instagram): blur, shift, detecting edges, and much more.
This [article](http://setosa.io/ev/image-kernels/) (**recommended read**) about image kernels should give you a better understanding of convolutions. It happens to be interactive as well.
In convolutional layers, the kernels are learned by training on the dataset. However, there are predefined kernels, for example used on your Instagram photos. Some examples are:
**Sharpen Kernel:**
$$
\begin{equation*}
\begin{bmatrix}
0 & -1 & 0 \\
-1 & 5 & -1 \\
0 & -1 & 0
\end {bmatrix}
\end{equation*}
$$
**Edge detection filter:**
$$
\begin{equation*}
\begin{bmatrix}
-1 & -1 & -1 \\
-1 & 8 & -1 \\
-1 & -1 & -1
\end {bmatrix}
\end{equation*}
$$
**Box blur of size 3:**
$$ \frac{1}{9}
\begin{equation*}
\begin{bmatrix}
1 & 1 & 1 \\
1 & 1 & 1 \\
1 & 1 & 1
\end {bmatrix}
\end{equation*}
$$
Let's play with convolutions by manipulating an image of a dog.
```
rgb_img = plt.imread('./images/dog.png')
plt.imshow(rgb_img)
```
Coloured images would require a 3-dimensional tensor to represent RGB (red, green, and blue).
Therefore, we will convert it to grayscale. This way it can be processed as a matrix.
```
img = rgb_img.mean(axis=2)
plt.imshow(img, cmap='gray')
```
First of all, let's blur the image with [box blur](https://en.wikipedia.org/wiki/Box_blur). It is just a convolution of a matrix with the kernel of size $N \times N$ of the following form:
$$
\frac{1}{N^2}
\begin{bmatrix}
1 & \dots & 1\\
\vdots & \ddots & \vdots\\
1 & \dots & 1\\
\end{bmatrix}
$$
Every element of this filter is *one* and we divide the sum by the total amount of elements in the blur filter. You could understand it as taking the average of an image region.
**Description:**
Perform the blur of the image.
<u>Arguments:</u>
* `image` - Input matrix - [np.array](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.array.html) of size `(N, M)`
* `box_size` - Size of the blur kernel - `int > 0` the kernel is of size `(box_size, box_size)`
<u>Output:</u>
The result of the blur [np.array](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.array.html) of size `(N, M)`
```
def box_blur(image, box_size):
"""Perform the blur of the image
# Arguments
image: input matrix - np.array of size `(N, M)`
box_size: the size of the blur kernel - int > 0
the kernel is of size `(box_size, box_size)`
# Output
the result of the blur
np.array of size `(N, M)`
"""
kernel = (1 / np.power(box_size, 2)) * np.ones((box_size, box_size))
output = conv_matrix(image, kernel)
return output
```
You can test your solution before submitting it. Running the following code block should yield this result:
$$
\begin{equation*}
\begin{bmatrix}
1 & 2 & 1 \\
2 & 4 & 2 \\
1 & 2 & 1
\end {bmatrix}
\end{equation*}
$$
```
test_image = np.array([
[9, 0, 9],
[0, 0, 0],
[9, 0, 9]
])
print(box_blur(test_image, 3))
```
Submit your solution here:
```
am.test_student_function(username, box_blur, ['image', 'box_size'])
```
Let's apply your blur convolution on our dog:
```
blur_dog = box_blur(img, box_size=5)
plt.imshow(blur_dog, cmap='gray')
```
Now, we will get the vertical and horizontal gradients. To perform it we calculate the convolution of the image with the following kernels:
$$
K_h =
\begin{bmatrix}
-1 & 0 & 1\\
\end{bmatrix} \quad
K_v =
\begin{bmatrix}
1 \\
0 \\
-1\\
\end{bmatrix} \\
X_h = X \star K_h \quad X_v = X \star K_v\\
$$
And then we calculate the amplitude of the gradient:
$$
X_\text{grad} = \sqrt{X_h^2 + X_v^2}
$$
```
dog_h = conv_matrix(blur_dog, np.array([[-1, 0, 1]]))
dog_v = conv_matrix(blur_dog, np.array([[-1, 0, 1]]).T)
dog_grad = np.sqrt(dog_h ** 2 + dog_v ** 2)
plt.imshow(dog_grad, cmap='gray')
```
This yields the edges of our blurred dog. It is not the only way to obtain edges though, there are plenty more:
* [Canny edge detection](https://en.wikipedia.org/wiki/Canny_edge_detector)
* [Sobel operator](https://en.wikipedia.org/wiki/Sobel_operator)
* [Prewitt operator](https://en.wikipedia.org/wiki/Prewitt_operator)
When you convolve an image with a kernel you obtain a map of responses. The more correlated the patch of an image is with the kernel, the higher the response. Let's take a closer look:
```
pattern = np.array([
[0, 1, 0],
[1, 1, 1],
[0, 1, 0]
])
# Create the image
image = np.pad(pattern, [(12, 12), (10, 14)], mode='constant', constant_values=0)
plt.imshow(image, cmap='gray')
plt.title('original image')
plt.show()
# Add some noise
image = 0.5 * image + 0.5 * np.random.random(image.shape)
plt.imshow(image, cmap='gray')
plt.title('noisy image')
plt.show()
# Let's find the cross
response = conv_matrix(image, pattern)
plt.imshow(response, cmap='gray')
plt.title('local response')
plt.show()
```
The brightest pixel highlights where the cross is located. We can find the area where the image is locally close to the kernel. This is especially useful for finding different patterns in images such as: eyes, legs, dogs, cats, etc.
We defined kernels and applied them to images. But we can also learn them by minimizing loss and making the processing as effective as possible. In order to do this, we have to define the **Convolutional Layer** in the next chapter.
# 4. Convolutional Layer
A **Convolutional Layer** works with images. Each image is a 3-dimensional object $N_{\text{channels}} \times H \times W$.
$Channels$ refers to the 3 colors (or 1 for black & white images), $H$ to height, and $W$ to width.
And therefore, the collection of images is 4-dimensional tensor of shape $N_{\text{objects}} \times N_{\text{channels}} \times H \times W$.
For example, 32 RGB images of size $224 \times 224$ are represented as a tensor of shape $32 \times 3 \times 224 \times 224$
A convolutional layer receives an image as its input. Here is how it works:
The layer has `n_in * n_out` kernels. It is a tensor of size `(n_in, n_out, kernel_h, kernel_w)`
It takes a 4-dimensional tensor of size `n_objects, n_in, H, W` as its input.
* `n_objects` is the collection of images.
* Each of them has `n_in` channels.
* The resolution of the images is `(H, W)`
For each of the images the following operation is performed:
* In order to get the 1st output channel, all inputs are convolved with their corresponding kernels.
* Then the results are summed and written to the output channel.
This is our implementation:
```python
for i in range(n_out):
out_channel = 0.0
for j in range(n_in):
kernel_2d = K[i, j] # Retrieve kernel from the collection of kernels
input_channel = input_image[j] # Get one channel of the input iamge
out_channel += conv_matrix(input_channel, kernel_2d) # Perform convolution
output_image.append(out_channel) # Append the calculated channel to the output
```
We implemented the convolutional layer for you. The implementation of `backward` is based on the idea that convolution could be represented as matrix multiplication.
```
class ConvLayer(Layer):
"""
Convolutional Layer. The implementation is based on
the representation of the convolution as matrix multiplication
"""
def __init__(self, n_in, n_out, filter_size):
super(ConvLayer, self).__init__()
self.W = np.random.normal(size=(n_out, n_in, filter_size, filter_size))
self.b = np.zeros(n_out)
def forward(self, x_input):
n_obj, n_in, h, w = x_input.shape
n_out = len(self.W)
self.output = []
for image in x_input:
output_image = []
for i in range(n_out):
out_channel = 0.0
for j in range(n_in):
out_channel += conv_matrix(image[j], self.W[i, j])
output_image.append(out_channel)
self.output.append(np.stack(output_image, 0))
self.output = np.stack(self.output, 0)
return self.output
def backward(self, x_input, grad_output):
N, C, H, W = x_input.shape
F, C, HH, WW = self.W.shape
pad = int((HH - 1) / 2)
self.grad_b = np.sum(grad_output, (0, 2, 3))
# pad input array
x_padded = np.pad(x_input, ((0,0), (0,0), (pad, pad), (pad, pad)), 'constant')
H_padded, W_padded = x_padded.shape[2], x_padded.shape[3]
# naive implementation of im2col
x_cols = None
for i in range(HH, H_padded + 1):
for j in range(WW, W_padded+1):
for n in range(N):
field = x_padded[n, :, i-HH:i, j-WW:j].reshape((1,-1))
if x_cols is None:
x_cols = field
else:
x_cols = np.vstack((x_cols, field))
x_cols = x_cols.T
d_out = grad_output.transpose(1, 2, 3, 0)
dout_cols = d_out.reshape(F, -1)
dw_cols = np.dot(dout_cols, x_cols.T)
self.grad_W = dw_cols.reshape(F, C, HH, WW)
w_cols = self.W.reshape(F, -1)
dx_cols = np.dot(w_cols.T, dout_cols)
dx_padded = np.zeros((N, C, H_padded, W_padded))
idx = 0
for i in range(HH, H_padded + 1):
for j in range(WW, W_padded + 1):
for n in range(N):
dx_padded[n:n+1, :, i-HH:i, j-WW:j] += dx_cols[:, idx].reshape((1, C, HH, WW))
idx += 1
dx = dx_padded[:, :, pad:-pad, pad:-pad]
grad_input = dx
return grad_input
def get_params(self):
return [self.W, self.b]
def get_params_gradients(self):
return [self.grad_W, self.grad_b]
```
This layer transforms images with 3 channels into images with 8 channels by convolving them with kernels of size `(3, 3)`
```
conv_layer = ConvLayer(3, 8, filter_size=3)
am.get_progress(username)
```
# 5. Pooling Layer
The pooling layer **reduces the size of an image**.
In the following figure $2 \times 2$ pooling is applied on the image which effectively reduces the size by half.
If you look closely, pooling operations have no effect on the depth of an image.

There are several types of pooling operations but the most common one is **max pooling**.
During a max pooling operation, the image is split into **windows** (or **filters**) and then the maximum of each window is used as the output.

```
def maxpool_forward(x_input):
"""Perform max pooling operation with 2x2 window
# Arguments
x_input: np.array of size (2 * W, 2 * H)
# Output
output: np.array of size (W, H)
"""
W, H = x_input.shape
output = x_input.reshape(W // 2, 2, H // 2, 2).max(axis=(1, 3))
return output
```
Once again, you can use example data to test your solution:
**Image:**
$$
\begin{equation*}
\begin{bmatrix}
1 & 1 & 2 & 4 \\
5 & 6 & 7 & 8 \\
3 & 2 & 1 & 0 \\
1 & 2 & 3 & 4
\end {bmatrix}
\end{equation*}
$$
**Output:**
$$
\begin{equation*}
\begin{bmatrix}
6 & 8 \\
3 & 4
\end {bmatrix}
\end{equation*}
$$
```
test_image = np.array([
[1, 1, 2, 4],
[5, 6, 7, 8],
[3, 2, 1, 0],
[1, 2, 3, 4]
])
print(maxpool_forward(test_image))
```
Submit your solution once you are confident it is implemented correctly.
```
am.test_student_function(username, maxpool_forward, ['x_input'])
```
We already implemented the gradient calculation.
It is not overly complicated; reading the code should help you to understand the concept.
```
def maxpool_grad_input(x_input, grad_output):
"""Calculate partial derivative of the loss with respect to the input
# Arguments
x_input: np.array of size (2 * W, 2 * H)
grad_output: partial derivative of the loss
with respect to the output
np.array of size (W, H)
# Output
output: partial derivative of the loss
with respect to the input
np.array of size (2 * W, 2 * H)
"""
height, width = x_input.shape
# Create an array of zeros the same size as the input
grad_input = np.zeros(x_input.shape)
# We set 1 if the element is the maximum in its window
for i in range(0, height, 2):
for j in range(0, width, 2):
window = x_input[i:i+2, j:j+2]
i_max, j_max = np.unravel_index(np.argmax(window), (2, 2))
grad_input[i + i_max, j + j_max] = 1
# Overwrite with the corresponding gradient instead of 1
grad_input = grad_input.ravel()
grad_input[grad_input == 1] = grad_output.ravel()
grad_input = grad_input.reshape(x_input.shape)
return grad_input
```
Following up is the full implementation of the **MaxPool Layer**.
```
class MaxPool2x2(Layer):
def forward(self, x_input):
n_obj, n_ch, h, w = x_input.shape
self.output = np.zeros((n_obj, n_ch, h // 2, w // 2))
for i in range(n_obj):
for j in range(n_ch):
self.output[i, j] = maxpool_forward(x_input[i, j])
return self.output
def backward(self, x_input, grad_output):
n_obj, n_ch, _, _ = x_input.shape
grad_input = np.zeros_like(x_input)
for i in range(n_obj):
for j in range(n_ch):
grad_input[i, j] = maxpool_grad_input(x_input[i, j], grad_output[i, j])
return grad_input
```
# 6. Flatten
Convolutional neural networks are better at image processing than fully connected neural networks (dense networks). We will combine convolutional layers, which deal with 4-dimensional tensors, with dense layers, which work with matrices.
In order to bridge the gap between convolutional layers and dense layers we will implement the **Flatten Layer**.
The Flatten layer receives a 4-dimensional tensor of size `(n_obj, n_channels, h, w)` as its input and reshapes it into a 2-dimensional tensor (matrix) of size `(n_obj, n_channels * h * w)`.
The backward pass of this layer is pretty straightforward. Remember that we don't actually change any values; we merely reshape inputs.
**Please implement `flatten_forward` and `flatten_grad_input` functions using [np.reshape](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.reshape.html)**.
```
def flatten_forward(x_input):
"""Perform the reshaping of the tensor of size `(K, L, M, N)`
to the tensor of size `(K, L*M*N)`
# Arguments
x_input: np.array of size `(K, L, M, N)`
# Output
output: np.array of size `(K, L*M*N)`
"""
K, L, M, N = x_input.shape
output = x_input.reshape(K, L * M * N)
return output
```
You can use test data and compare the final shape. It should be `(100, 768)` for the following example.
Please ignore the use of [np.zeros](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.zeros.html) in this case. We are just interested in transforming shapes.
**Be aware:** This test will fail if you do not return an array like object!
```
test_input = np.zeros((100, 3, 16, 16))
print(flatten_forward(test_input).shape)
```
Submit your solution here:
```
am.test_student_function(username, flatten_forward, ['x_input'])
def flatten_grad_input(x_input, grad_output):
"""Calculate partial derivative of the loss with respect to the input
# Arguments
x_input: partial derivative of the loss
with respect to the output
np.array of size `(K, L*M*N)`
# Output
output: partial derivative of the loss
with respect to the input
np.array of size `(K, L, M, N)`
"""
grad_input = grad_output.reshape(x_input.shape)
return grad_input
```
Once again, submit your solution here:
```
am.test_student_function(username, flatten_grad_input, ['x_input', 'grad_output'])
```
This is the, pretty self-explanatory, implemention of the **Flatten Layer**.
```
class FlattenLayer(Layer):
def forward(self, x_input):
self.output = flatten_forward(x_input)
return self.output
def backward(self, x_input, grad_output):
output = flatten_grad_input(x_input, grad_output)
return output
am.get_progress(username)
```
# 7. Experiments
This chapter focuses on conducting several experiments. We will train our neural networks with **mini-batches**. Mini-batches are small portions of our dataset, all mini-batches together should form the original dataset again. With our mini-batches in place we will feed these one-by-one to our neural network.
```
import sys
def iterate_minibatches(x, y, batch_size=16, verbose=True):
assert len(x) == len(y)
indices = np.arange(len(x))
np.random.shuffle(indices)
for i, start_idx in enumerate(range(0, len(x) - batch_size + 1, batch_size)):
if verbose:
print('\rBatch: {}/{}'.format(i + 1, len(x) // batch_size), end='')
sys.stdout.flush()
excerpt = indices[start_idx:start_idx + batch_size]
yield x[excerpt], y[excerpt]
```
Let's import the data. Please [download](http://yann.lecun.com/exdb/mnist/) it first.
If you get an error with loading the data, chances are you'll need to unpack the downloaded files.
```
from dataset_utils import load_mnist
train = list(load_mnist(dataset='training', path='.'))
train
train_images = np.array([im[1] for im in train])
train_targets = np.array([im[0] for im in train])
# We will train a 0 vs. 1 classifier
x_train = train_images[train_targets < 2][:1000]
y_train = train_targets[train_targets < 2][:1000]
y_train = y_train * 2 - 1
y_train = y_train.reshape((-1, 1))
```
You just loaded the MNIST dataset. This dataset consists of gray-scale (so a single channel) images of size `28x28`. These images are represented by the RGB color model. This color model representes a color by 3 integers that range from 0 to 255, or in the case of gray-scale images this is a single integer. This means that each picture in the MNIST dataset is represented by 784 pixels with a value ranging from 0 to 255. This is how a single image looks like:
```
plt.imshow(x_train[0].reshape(28, 28), cmap='gray_r')
```
To make divergence to an optimum easier, we will normalize the images to have values between 0 and 1. Then, by reshaping, we will add the dimensions for the channel which, for simplicity, was removed by the creators of this dataset. As you can see, this doesn't change anything in how the image looks like.
```
x_train = x_train.astype('float32') / 255.0
x_train = x_train.reshape((-1, 1, 28, 28))
plt.imshow(x_train[0].reshape(28, 28), cmap='gray_r')
```
Now we will train a simple convolutional neural network:
```
def get_cnn():
nn = SequentialNN()
nn.add(ConvLayer(1, 2, filter_size=3)) # The output is of size N_obj 2 28 28
nn.add(ReLU()) # The output is of size N_obj 2 28 28
nn.add(MaxPool2x2()) # The output is of size N_obj 2 14 14
nn.add(ConvLayer(2, 4, filter_size=3)) # The output is of size N_obj 4 14 14
nn.add(ReLU()) # The output is of size N_obj 4 14 14
nn.add(MaxPool2x2()) # The output is of size N_obj 4 7 7
nn.add(FlattenLayer()) # The output is of size N_obj 196
nn.add(Dense(4 * 7 * 7, 32))
nn.add(ReLU())
nn.add(Dense(32, 1))
return nn
nn = get_cnn()
loss = Hinge()
optimizer = SGD(nn)
# It will train for about 5 minutes
num_epochs = 5
batch_size = 32
# We will store the results here
history = {'loss': [], 'accuracy': []}
# `num_epochs` represents the number of iterations
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch + 1, num_epochs))
# We perform iteration a one-by-one iteration of the mini-batches
for x_batch, y_batch in iterate_minibatches(x_train, y_train, batch_size):
# Predict the target value
y_pred = nn.forward(x_batch)
# Compute the gradient of the loss
loss_grad = loss.backward(y_pred, y_batch)
# Perform backwards pass
nn.backward(x_batch, loss_grad)
# Update the params
optimizer.update_params()
# Save loss and accuracy values
history['loss'].append(loss.forward(y_pred, y_batch))
prediction_is_correct = (y_pred > 0) == (y_batch > 0)
history['accuracy'].append(np.mean(prediction_is_correct))
print()
# Let's plot the results to get a better insight
plt.figure(figsize=(8, 5))
ax_1 = plt.subplot()
ax_1.plot(history['loss'], c='g', lw=2, label='train loss')
ax_1.set_ylabel('loss', fontsize=16)
ax_1.set_xlabel('#batches', fontsize=16)
ax_2 = plt.twinx(ax_1)
ax_2.plot(history['accuracy'], lw=3, label='train accuracy')
ax_2.set_ylabel('accuracy', fontsize=16)
```
**Things you could try:**
Train the model with a different `batch_size`:
* What would happen with `batch_size=1`?
* What would happen with `batch_size=1000`?
* Does the speed of the computation depend on this parameter? If so, why?
Train the model with a different number of `num_epochs`:
* What would happen with `num_epochs=1`?
* What would happen with `num_epochs=1000`?
* How does it affect computation time, resource strain, and accuracy?
Let's visualize the activations of the intermediate layers:
```
viz_images = x_batch[:2]
_ = nn.forward(viz_images)
activations = {
'conv_1': nn.layers[0].output,
'relu_1': nn.layers[1].output,
'pool_1': nn.layers[2].output,
'conv_2': nn.layers[3].output,
'relu_2': nn.layers[4].output,
'pool_2': nn.layers[5].output,
}
```
### Input Images
```
# Input
f, (ax1, ax2) = plt.subplots(2, 1, figsize=(4, 8))
ax1.imshow(viz_images[0, 0], cmap=plt.cm.gray_r)
ax1.set_xticks([])
ax1.set_yticks([])
ax2.imshow(viz_images[1, 0], cmap=plt.cm.gray_r)
ax2.set_xticks([])
ax2.set_yticks([])
plt.show()
```
### Activations of Conv 1
```
# Conv 1
f, axes = plt.subplots(2, 2, figsize=(8, 8))
for i in range(2):
for j in range(2):
ax = axes[i, j]
ax.imshow(activations['conv_1'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
### Activations of ReLU 1
```
# ReLU 1
f, axes = plt.subplots(2, 2, figsize=(8, 8))
for i in range(2):
for j in range(2):
ax = axes[i, j]
ax.imshow(activations['relu_1'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
### Activations of MaxPooling 1
```
# Max Pooling 1
f, axes = plt.subplots(2, 2, figsize=(8, 8))
for i in range(2):
for j in range(2):
ax = axes[i, j]
ax.imshow(activations['pool_1'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
### Activations of Conv 2
```
# Conv 2
f, axes = plt.subplots(2, 4, figsize=(16, 8))
for i in range(2):
for j in range(4):
ax = axes[i, j]
ax.imshow(activations['conv_2'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
### Activations of ReLU 2
```
# ReLU 2
f, axes = plt.subplots(2, 4, figsize=(16, 8))
for i in range(2):
for j in range(4):
ax = axes[i, j]
ax.imshow(activations['relu_2'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
### Activations of MaxPooling 2
```
# Max Pooling 2
f, axes = plt.subplots(2, 4, figsize=(16, 8))
for i in range(2):
for j in range(4):
ax = axes[i, j]
ax.imshow(activations['pool_2'][i, j], cmap=plt.cm.gray_r)
ax.set_xticks([])
ax.set_yticks([])
ax.set_title('Channel {}'.format(j + 1))
plt.show()
```
As we go deeper and deeper, images become less locally-correlated (the dependance between two neighbours decreases) and more semantically loaded.
Each convoluted pixel stores more and more useful information about the object.
In the end, this will be anaylzed using several **Dense Layers**.
**Things you could try:**
* Change the architecture of the neural network
* Vary the number of kernels
* Vary the size of the kernels
| github_jupyter |
```
import os
os.chdir('..')
```
<img src="flow_0.png">
```
import numpy as np
from flows.flows import Flows
flow = Flows(flow_id=0, categorical_threshold=10)
path = os.path.join('data','flow_0')
files_list = ["train.csv","test.csv"]
dataframe_dict, columns_set = flow.load_data(path, files_list)
columns_set
dataframe_dict, columns_set = flow.encode_categorical_feature(dataframe_dict)
ignore_columns = ['Id', 'SalePrice']
dataframe_dict, columns_set = flow.scale_data(dataframe_dict, ignore_columns)
flow.exploring_data(dataframe_dict, "train")
flow.comparing_statistics(dataframe_dict)
ignore_columns = ["Id", "SalePrice"]
columns = dataframe_dict["train"].columns
train_dataframe = dataframe_dict["train"][[x for x in columns_set["train"]["continuous"] if x not in ignore_columns]]
test_dataframe = dataframe_dict["test"][[x for x in columns_set["train"]["continuous"] if x not in ignore_columns]]
train_target = dataframe_dict["train"]["SalePrice"]
parameters = {
"data": {
"train": {"features": train_dataframe, "target": train_target.to_numpy()},
},
"split": {
"method": "kfold", # "method":"kfold"
"fold_nr": 5, # fold_nr:5 , "split_ratios": 0.3 # "split_ratios":(0.3,0.2)
},
"model": {"type": "Ridge linear regression",
"hyperparameters": {"alpha": "optimize", # alpha:optimize
},
},
"metrics": ["r2_score"],
"predict": {
"test": {"features": test_dataframe}
}
}
model_index_list, save_models_dir, y_test = flow.training(parameters)
parameters_lighgbm = {
"data": {
"train": {"features": train_dataframe, "target": train_target.to_numpy()},
},
"split": {
"method": "kfold", # "method":"kfold"
"fold_nr": 5, # fold_nr:5 , "split_ratios": 0.3 # "split_ratios":(0.3,0.2)
},
"model": {"type": "lightgbm",
"hyperparameters": dict(objective='regression', metric='root_mean_squared_error', num_leaves=5,
boost_from_average=True,
learning_rate=0.05, bagging_fraction=0.99, feature_fraction=0.99, max_depth=-1,
num_rounds=10000, min_data_in_leaf=10, boosting='dart')
},
"metrics": ["mean_squared_error"],
"predict": {
"test": {"features": test_dataframe}
}
}
model_index_list, save_models_dir, y_test = flow.training(parameters_lighgbm)
parameters_xgboost = {
"data": {
"train": {"features": train_dataframe, "target": train_target.to_numpy()},
},
"split": {
"method": "kfold", # "method":"kfold"
"fold_nr": 5, # fold_nr:5 , "split_ratios": 0.3 # "split_ratios":(0.3,0.2)
},
"model": {"type": "xgboost",
"hyperparameters": {'max_depth': 5, 'eta': 1, 'eval_metric': "rmse", "num_round": 100}
},
"metrics": ["r2_score", "mean_squared_error"],
"predict": {
"test": {"features": test_dataframe}
}
}
model_index_list, save_models_dir, ytest = flow.training(parameters_xgboost)
parameters_sklearn = {
"data": {
"train": {"features": train_dataframe, "target": train_target.to_numpy()},
},
"split": {
"method": "split",
"split_ratios": 0.2,
"stratify": False # set to True only for classification tasks
},
"model": {
"type": "sklearn.ensemble.RandomForestRegressor",
"hyperparameters": {
'params_grid':{
'criterion': ["mse", "mae"],
'max_depth': [5, 10, 15, 999],
'min_samples_leaf': [4, 1],
'max_depth': [4, 8, 12],
},
'params_fixed': {
'min_samples_split': 10,
'random_state': 11
},
'params_cv': {
'n_splits': 5,
'shuffle': True,
'random_state': 11
},
'objective': 'regression', # 'classification'
"grid_search_scoring": ['r2', 'neg_mean_squared_error']
},
},
"metrics": ["r2_score", "mean_squared_error"],
"predict": {
"test": {"features": test_dataframe}
}
}
model_index_list, save_models_dir, y_test = flow.training(parameters_sklearn)
parameters_sklearn = {
"data": {
"train": {"features": train_dataframe, "target": train_target.to_numpy()},
},
"split": {
"method": "split",
"split_ratios": 0.2,
"stratify": False # set to True only for classification tasks
},
"model": {
"type": "sklearn.linear_model.ElasticNet",
"hyperparameters": {
'params_grid':{
'alpha': np.logspace(-3,3,7),
'l1_ratio': np.linspace(0, 0., num=4)+0.01
},
'params_fixed': {
'normalize': True,
'max_iter': 2000,
'random_state': 11
},
'params_cv': {
'n_splits': 5,
'shuffle': True,
'random_state': 11
},
'objective': 'regression', # 'classification'
"grid_search_scoring": ['r2', 'neg_mean_squared_error']
},
},
"metrics": ["r2_score", "mean_squared_error"],
"predict": {
"test": {"features": test_dataframe}
}
}
model_index_list, save_models_dir, y_test = flow.training(parameters_sklearn)
```
| github_jupyter |
# Introduction: How H1st.AI enables the Industrial AI Revolution
This tutorial will teach you how H1st AI can help solve the Cold Start problem in domains where labeled data is not available or prohibitively expensive to obtain.
One example of such a domain is cybersecurity, which is increasingly looking forward to adopting ML to detect intrusions. Another domain is predictive maintenance that tries to anticipate industrial machine failures before they happen. In both domains, labels are expensive because fundamentally these occurrences are rare and costly (as compared to NLP where e.g. sentiment are common and labels can be obtained i.g. via crowdsourcing or weak supervision).
Yet this is a fundamental challenge of Industrial AI.
<img src="https://h1st-static.s3.amazonaws.com/batman+h1st.ai.jpg" alt="H1st.AI woke meme" style="float: left; margin-right: 20px; margin-bottom: 20px;" width=320px height=320px>
Jurgen Schmidhuber, one of AI & deep learning's pioneer, [remarked in his 2020s outlook that](http://people.idsia.ch/~juergen/2010s-our-decade-of-deep-learning.html#Sec.%207) in the last decade AI "excelled in virtual worlds, e.g., in video games, board games, and especially on the major WWW platforms", but the main challenge for the next decades is for AI to be "driving industrial processes and machines and robots".
As pioneers in Industrial AI who regularly work with massive global fleets of IoT equipment, Arimo & Panasonic whole-heartedly agrees with this outlook. Importantly, many industrial AI use cases with significant impact have become urgent and demand solutions now that requires a fresh approach. We will work on one such example in this tutorial: detection intrusion in automotive cybersecurity.
We’ll learn that using H1st.AI we can tackle these problems and make it tractable by leveraging human experience and data-driven models in a harmonious way. Especially, we’ll learn how to:
* Perform use-case analysis to decompose problems and adopt different models at the right level of abstractions
* Encode human experience as a model
* Combine human and ML models to work in tandem in a H1st.Graph
Too many tutorials, esp data science ones, start out with some toy applications and the really basic stuff, and then stalls out on the more complex real-world scenario. This one is going to be different.
So, grab a cup of coffee before you continue :)
If you can't wait, go ahead and [star our Github repository](https://github.com/h1st-ai/h1st) and check out the "Quick Start" section. We're open-source!
| github_jupyter |
# Define Concatenation Image Data Loader
```
import os
import torch
from torch.utils.data import Dataset
import numpy as np
from PIL import Image
import albumentations as A
import torchvision.transforms as transforms
from torch.utils.data import DataLoader
import string
import math
augmentation = A.Compose([
A.OneOf([
A.IAAAdditiveGaussianNoise(p=0.9),
A.GaussNoise(p=0.9),
], p=0.9),
A.OneOf([
A.MotionBlur(p=0.9),
A.MedianBlur(blur_limit=3, p=0.9),
A.Blur(blur_limit=4, p=0.9),
], p=0.9),
A.OneOf([
A.CLAHE(clip_limit=2, p=0.9),
A.IAASharpen(p=0.9),
A.IAAEmboss(p=0.9),
A.RandomBrightnessContrast(p=0.95),
], p=0.9),
A.OneOf(
[
A.HueSaturationValue(p=0.9),
A.RandomGamma(p=0.9),
A.IAAPerspective(p=0.05),
], p=0.9,
)
])
def build_transform(shape):
transform = transforms.Compose([
transforms.ToTensor()
])
return transform
class VideoDataset(Dataset):
def __init__(self, folder_list, char_dict,
fixed_frame_num=200, fixed_max_len=6,
image_shape=(100, 100),
aug=augmentation):
self.folders = folder_list
np.random.shuffle(self.folders)
self.fixed_frame_num = fixed_frame_num
self.char_dict = char_dict
self.fixed_max_len = fixed_max_len
self.augmentation = aug
self.image_shape = image_shape
self.transform = build_transform(shape=self.image_shape)
def __len__(self):
return len(self.folders)
def __getitem__(self, index):
image_folder = self.folders[index]
label = image_folder.split("/")[-1].split("_")[-1].strip(" ")
label_digit = [self.char_dict[i] for i in label]
assert len(label_digit) < self.fixed_max_len
label_digit.append(self.char_dict["<eos>"])
rest = self.fixed_max_len - len(label_digit)
if rest:
label_digit += [self.char_dict["<blank>"]] * rest
image_list = [os.path.join(image_folder, i) for i in os.listdir(image_folder) if i.endswith(".jpg")]
image_list = sorted(image_list)
images = []
k_col, k_row = 4, 4
max_frame_num = k_col * k_row
if len(image_list) <= max_frame_num:
image_list += ["pad"] * (max_frame_num - len(image_list))
k_frame_pick_one = math.floor(len(image_list) / (k_col * k_row))
# print('k_frame_pick_one: ', k_frame_pick_one)
for index,i in enumerate(image_list):
if index%k_frame_pick_one == 0:
if i != "pad":
img = Image.open(i).convert("RGB")
if self.augmentation is not None:
img = self.augmentation(image=np.array(img, dtype=np.uint8))["image"]
img = Image.fromarray(img)
else:
img = Image.new("RGB", (self.image_shape[1], self.image_shape[0]))
img = img.resize(self.image_shape)
images.append(img)
x = Image.new('RGB', (self.image_shape[1] * k_row, self.image_shape[0] * k_col))
for i in range(k_col):
for k in range(k_row):
x.paste(images[i * k_col + k], (self.image_shape[1] * k, self.image_shape[0] * i))
x.save('./test.jpg', quality=50)
x = self.transform(x)
y = torch.tensor(label_digit, dtype=torch.long)
return x, y
```
# Build 2DCNN + RNN lipsreading model
```
import torch
import torch.nn.functional as F
class BidirectionalLSTM(torch.nn.Module):
def __init__(self, nIn, nHidden, nOut):
super(BidirectionalLSTM, self).__init__()
self.rnn = torch.nn.LSTM(nIn, nHidden, bidirectional=True, batch_first=True)
self.embedding = torch.nn.Linear(nHidden * 2, nOut)
# self.embedding_1 = torch.nn.Linear(nHidden * 2, nHidden)
# self.embedding_2 = torch.nn.Linear(nHidden, nHidden//2)
# self.embedding_3 = torch.nn.Linear(nHidden//2, nOut)
# self.dropout_1 = torch.nn.Dropout(p=0.1)
# self.dropout_2 = torch.nn.Dropout(p=0.25)
def forward(self, inputs):
recurrent, _ = self.rnn(inputs)
T, b, h = recurrent.size()
t_rec = recurrent.reshape(T * b, h)
# output = self.embedding_1(t_rec) # [T * b, nOut]
# output = self.dropout_1(output)
# output = F.relu(output)
#
# output = self.embedding_2(output)
# # output = self.dropout_2(output)
# output = F.relu(output)
#
# output = self.embedding_3(output)
output = self.embedding(t_rec)
output = output.reshape(T, b, -1)
# output = F.softmax(output, dim=-1)
return output
class VideoModel(torch.nn.Module):
def __init__(self, number_classes=28, max_len=6, image_shape=(60, 60)):
"""
:param number_classes:
our char dictionary is:
0: <blank>
1: a
2: b
3: c
...
26: z
27: <eos>
:param max_len: max_len = 6,
Suppose we said abcde,
the the label should be abcde<eos>
abc -> abc<eos><blank><blank>
number_classes = 28, 26 characters + <eos> + <blank>
"""
super(VideoModel, self).__init__()
self.number_classes = number_classes
self.max_len = max_len
self.conv_block_1 = self. _cnn2d_block_2_conv_layer(3, 32)
self.conv_block_2 = self. _cnn2d_block_2_conv_layer(32, 64)
self.conv_block_3 = self. _cnn2d_block_2_conv_layer(64, 128)
self.conv_block_4 = self. _cnn2d_block_2_conv_layer(128, 256)
self.lstm_decoder = BidirectionalLSTM(nIn=9600,
nHidden=256,
nOut=number_classes)
def _cnn2d_block_2_conv_layer(self, input_size, output_size):
conv2d_block = torch.nn.Sequential(
torch.nn.Conv2d(input_size, output_size, kernel_size=3, padding=1),
torch.nn.ReLU(),
torch.nn.Conv2d(output_size, output_size, kernel_size=3, padding=1),
torch.nn.ReLU(),
torch.nn.MaxPool2d(2,2)
)
return conv2d_block
def _cnn2d_block_1_conv_layer(self, input_size, output_size):
conv2d_block = torch.nn.Sequential(
torch.nn.Conv2d(input_size, output_size, kernel_size=3, padding=1),
torch.nn.ReLU(),
torch.nn.MaxPool2d(2,2)
)
return conv2d_block
def forward(self, x):
# x = x.permute(dims=(0, 2, 3, 4, 1))
x = self.conv_block_1(x)
x = self.conv_block_2(x)
x = self.conv_block_3(x)
x = self.conv_block_4(x)
shape = x.size()
# bs, 256, 3, 3, 14
x = x.view(shape[0], self.max_len, -1) # bs, max_len, rest
x = self.lstm_decoder(x)
return x
```
# Define the dataloader in this task
```
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "0"
from cnn2d_image_generator import VideoDataset
from image_2dcrnn import VideoModel
import torch
from torch.utils.data import DataLoader
import string
import time
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
def make_char_dict():
chars = string.ascii_lowercase
char_dict = {"<blank>": 0}
for idx, c in enumerate(chars):
char_dict[c] = idx + 1
current_len = len(list(char_dict.keys()))
char_dict["<eos>"] = current_len
print(char_dict)
return char_dict
def get_train_test_folders():
test = open("data/eval_lst.txt", "r", encoding="utf-8").readlines()
train = open("data/train_lst.txt", "r", encoding="utf-8").readlines()
train_folders = [os.path.join("data", "data_aligned", i.strip("\n")) for i in train]
test_folders = [os.path.join("data", "data_aligned", i.strip("\n")) for i in test]
print("train videos:{}".format(len(train_folders)))
print("test videos:{}".format(len(test_folders)))
return train_folders, test_folders
image_shape = (60, 60)
char_dict = make_char_dict()
train_folders, test_folders = get_train_test_folders()
train_dataset = VideoDataset(
folder_list=train_folders,
char_dict=char_dict,
fixed_frame_num=200,
fixed_max_len=6,
image_shape=image_shape,
)
batch_size = 10
train_dataloader = DataLoader(
train_dataset, batch_size=batch_size, shuffle=True
)
test_dataset = VideoDataset(
folder_list=test_folders,
char_dict=char_dict,
fixed_frame_num=200,
fixed_max_len=6,
aug=None, # No need to do data augmentation in testing dataset
image_shape=image_shape,
)
test_dataloader = DataLoader(
test_dataset, batch_size=batch_size, shuffle=True
)
```
# Init model
```
model = VideoModel(number_classes=len(list(char_dict.keys())),
max_len=6,
image_shape=image_shape)
model = model.to(device)
print(model)
```
# Set up for training
```
criterion = torch.nn.CrossEntropyLoss().to(device)
optimizer = torch.optim.SGD(model.parameters(), lr=1, momentum=0.9)
steps_per_epoch = len(train_folders) // 10 + 1
epochs = 10
scheduler = torch.optim.lr_scheduler.ReduceLROnPlateau(optimizer,
mode='min',
verbose=True,
factor=0.1,
patience=5,
threshold=0.00001)
```
# Define training process
```
def train_process():
running_loss = 0
num_batches = 0
model.train()
for idx, data in enumerate(train_dataloader):
optimizer.zero_grad()
x, y = data
size = y.size()
x = x.to(device)
y = y.to(device)
x.requires_grad_()
scores = model(x)
scores = scores.view(size[0] * size[1], -1)
y = y.view(size[0] * size[1])
loss = criterion(scores, y)
loss.backward()
torch.nn.utils.clip_grad_norm_(model.parameters(), 5)
optimizer.step()
running_loss += loss.detach().item()
num_batches += 1
print("time:{}, epoch: {} step: {}, avg running loss is {}".format(
time.ctime(), epoch + 1, idx + 1, running_loss / num_batches
))
return running_loss, num_batches
```
# Define validation process
```
def testing_process():
running_loss = 0
num_batches = 0
model.eval()
with torch.no_grad():
for idx, data in enumerate(test_dataloader):
x, y = data
size = y.size()
x = x.to(device)
y = y.to(device)
scores = model(x)
scores = scores.view(size[0] * size[1], -1)
y = y.view(size[0] * size[1])
loss = criterion(scores, y)
running_loss += loss.item()
num_batches += 1
return running_loss, num_batches
```
# Train
```
for epoch in range(epochs):
running_loss, num_batches = train_process()
test_running_loss, test_num_batches = testing_process()
print("*" * 100)
print("epoch: {}, avg training loss:{}, avg validation loss:{}".format(epoch + 1, running_loss / num_batches,
test_running_loss / test_num_batches))
scheduler.step(test_running_loss / test_num_batches)
print("*" * 100)
k_col, k_row = 4, 4
save_name = '2dcrnn_model_'+str(k_col*k_row)+'_epoch_'+str(epochs)+'.pkl'
torch.save(model, save_name)
```
# Load model
```
model = torch.load(save_name)
```
# Test accuracy
```
def compute_val_acc(scores, y):
num = scores.size(0)
prediction = scores.argmax(dim=1)
indicator = (prediction == y)
num_matches = indicator.sum()
return num_matches.float() / num
model.eval()
acc = 0
count = 0
with torch.no_grad():
for idx, data in enumerate(test_dataloader):
x, y = data
size = y.size()
x = x.to(device)
y = y.to(device)
scores = model(x)
scores = scores.view(size[0] * size[1], -1)
y = y.view(size[0] * size[1])
acc += compute_val_acc(scores, y)
count += 1
print("Acc in inference process is {}".format(acc / count))
```
| github_jupyter |
```
from google.colab import drive
drive.mount('/content/drive')
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
from matplotlib import pyplot as plt
import copy
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=True)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
fg1,fg2,fg3 = 0,1,2
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000):
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]].type("torch.DoubleTensor"))
j+=1
else:
image_list.append(foreground_data[fg_idx].type("torch.DoubleTensor"))
label = foreground_label[fg_idx]- fg1 # minus 7 because our fore ground classes are 7,8,9 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
desired_num = 30000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(desired_num):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
batch = 125
msd = MosaicDataset(mosaic_list_of_images, mosaic_label , fore_idx)
train_loader = DataLoader( msd,batch_size= batch ,shuffle=True)
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0)
self.conv4 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv6 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32)
self.batch_norm2 = nn.BatchNorm2d(128)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(128,64)
self.fc2 = nn.Linear(64, 32)
self.fc3 = nn.Linear(32, 10)
self.fc4 = nn.Linear(10, 1)
def forward(self, x):
x = self.conv1(x)
x = F.relu(self.batch_norm1(x))
x = (F.relu(self.conv2(x)))
x = self.pool(x)
x = self.conv3(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv4(x)))
x = self.pool(x)
x = self.dropout1(x)
x = self.conv5(x)
x = F.relu(self.batch_norm2(x))
x = (F.relu(self.conv6(x)))
x = self.pool(x)
x = x.view(x.size(0), -1)
x = self.dropout2(x)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.dropout2(x)
x = F.relu(self.fc3(x))
x = self.fc4(x)
return x
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
# focus_net.load_state_dict( torch.load("/content/drive/My Drive/Research/Cheating_data/Focus_net_weights/focus_net"+"cnn"+".pt"))
```
Changing the last layer of Focus net
```
# focus_net.linear2 = nn.Linear(10,1).double()
# focus_net = focus_net.to("cuda")
for params in focus_net.parameters():
params.requires_grad = False
for params in focus_net.parameters():
print(params)
break;
class Classification(nn.Module):
def __init__(self, focus_net):
super(Classification, self).__init__()
self.focus_net = focus_net
self.conv1 = nn.Conv2d(in_channels=3, out_channels=32, kernel_size=3, padding=0)
self.conv2 = nn.Conv2d(in_channels=32, out_channels=64, kernel_size=3, padding=0)
self.conv3 = nn.Conv2d(in_channels=64, out_channels=128, kernel_size=3, padding=0)
self.conv4 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv5 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=0)
self.conv6 = nn.Conv2d(in_channels=128, out_channels=128, kernel_size=3, padding=1)
self.pool = nn.MaxPool2d(kernel_size=2, stride=2)
self.batch_norm1 = nn.BatchNorm2d(32)
self.batch_norm2 = nn.BatchNorm2d(128)
self.dropout1 = nn.Dropout2d(p=0.05)
self.dropout2 = nn.Dropout2d(p=0.1)
self.fc1 = nn.Linear(128,64)
self.fc2 = nn.Linear(64, 32)
self.fc3 = nn.Linear(32, 10)
self.fc4 = nn.Linear(10, 3)
def forward(self,z): #z batch of list of 9 images
y = torch.zeros([batch,3, 32,32], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
x = x.to("cuda")
y = y.to("cuda")
for i in range(9):
x[:,i] = self.focus_net.forward(z[:,i])[:,0]
x = F.softmax(x,dim=1)
x1 = x[:,0]
torch.mul(x1[:,None,None,None],z[:,0])
for i in range(9):
x1 = x[:,i]
y = y + torch.mul(x1[:,None,None,None],z[:,i])
y1 = self.conv1(y)
y1 = F.relu(self.batch_norm1(y1))
y1 = (F.relu(self.conv2(y1)))
y1 = self.pool(y1)
y1 = self.conv3(y1)
y1 = F.relu(self.batch_norm2(y1))
y1 = (F.relu(self.conv4(y1)))
y1 = self.pool(y1)
y1 = self.dropout1(y1)
y1 = self.conv5(y1)
y1 = F.relu(self.batch_norm2(y1))
y1 = (F.relu(self.conv6(y1)))
y1 = self.pool(y1)
y1 = y1.view(y1.size(0), -1)
y1 = self.dropout2(y1)
y1 = F.relu(self.fc1(y1))
y1 = F.relu(self.fc2(y1))
y1 = self.dropout2(y1)
y1 = F.relu(self.fc3(y1))
y1 = self.fc4(y1)
return y1 , x, y
classify = Classification(focus_net).double()
classify = classify.to("cuda")
test_images =[] #list of mosaic images, each mosaic image is saved as laist of 9 images
fore_idx_test =[] #list of indexes at which foreground image is present in a mosaic image
test_label=[] # label of mosaic image = foreground class present in that mosaic
for i in range(10000):
bg_idx = np.random.randint(0,35000,8)
fg_idx = np.random.randint(0,15000)
fg = np.random.randint(0,9)
fore_idx_test.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
test_images.append(image_list)
test_label.append(label)
test_data = MosaicDataset(test_images,test_label,fore_idx_test)
test_loader = DataLoader( test_data,batch_size= batch ,shuffle=False)
import torch.optim as optim
criterion_classify = nn.CrossEntropyLoss()
optimizer_classify = optim.SGD(classify.parameters(), lr=0.01, momentum=0.9)
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
nos_epochs = 60
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
# zero the parameter gradients
optimizer_classify.zero_grad()
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion_classify(outputs, labels)
loss.backward()
optimizer_classify.step()
running_loss += loss.item()
mini = 60
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.03):
break;
if epoch % 5 == 0:
col1.append(epoch+1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
#************************************************************************
#testing data set
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
print('Finished Training')
for params in focus_net.parameters():
print(params)
break;
name = "4_focus_random_classify_random_train_classify"
print(name)
torch.save(classify.state_dict(),"/content/drive/My Drive/Research/Cheating_data/16_experiments_on_cnn/"+name+".pt")
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
df_train = pd.DataFrame()
df_test = pd.DataFrame()
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
df_train
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.show()
df_test
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.show()
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 30000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
outputs, alphas, avg_images = classify(inputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
```
| github_jupyter |
<table class="ee-notebook-buttons" align="left">
<td><a target="_blank" href="https://github.com/giswqs/earthengine-py-notebooks/tree/master/Datasets/naip_imagery.ipynb"><img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" /> View source on GitHub</a></td>
<td><a target="_blank" href="https://nbviewer.jupyter.org/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/naip_imagery.ipynb"><img width=26px src="https://upload.wikimedia.org/wikipedia/commons/thumb/3/38/Jupyter_logo.svg/883px-Jupyter_logo.svg.png" />Notebook Viewer</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/giswqs/earthengine-py-notebooks/blob/master/Datasets/naip_imagery.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" /> Run in Google Colab</a></td>
</table>
## Install Earth Engine API and geemap
Install the [Earth Engine Python API](https://developers.google.com/earth-engine/python_install) and [geemap](https://github.com/giswqs/geemap). The **geemap** Python package is built upon the [ipyleaflet](https://github.com/jupyter-widgets/ipyleaflet) and [folium](https://github.com/python-visualization/folium) packages and implements several methods for interacting with Earth Engine data layers, such as `Map.addLayer()`, `Map.setCenter()`, and `Map.centerObject()`.
The following script checks if the geemap package has been installed. If not, it will install geemap, which automatically installs its [dependencies](https://github.com/giswqs/geemap#dependencies), including earthengine-api, folium, and ipyleaflet.
**Important note**: A key difference between folium and ipyleaflet is that ipyleaflet is built upon ipywidgets and allows bidirectional communication between the front-end and the backend enabling the use of the map to capture user input, while folium is meant for displaying static data only ([source](https://blog.jupyter.org/interactive-gis-in-jupyter-with-ipyleaflet-52f9657fa7a)). Note that [Google Colab](https://colab.research.google.com/) currently does not support ipyleaflet ([source](https://github.com/googlecolab/colabtools/issues/60#issuecomment-596225619)). Therefore, if you are using geemap with Google Colab, you should use [`import geemap.eefolium`](https://github.com/giswqs/geemap/blob/master/geemap/eefolium.py). If you are using geemap with [binder](https://mybinder.org/) or a local Jupyter notebook server, you can use [`import geemap`](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py), which provides more functionalities for capturing user input (e.g., mouse-clicking and moving).
```
# Installs geemap package
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# Checks whether this notebook is running on Google Colab
try:
import google.colab
import geemap.eefolium as emap
except:
import geemap as emap
# Authenticates and initializes Earth Engine
import ee
try:
ee.Initialize()
except Exception as e:
ee.Authenticate()
ee.Initialize()
```
## Create an interactive map
The default basemap is `Google Satellite`. [Additional basemaps](https://github.com/giswqs/geemap/blob/master/geemap/geemap.py#L13) can be added using the `Map.add_basemap()` function.
```
Map = emap.Map(center=[40,-100], zoom=4)
Map.add_basemap('ROADMAP') # Add Google Map
Map
```
## Add Earth Engine Python script
```
# Add Earth Engine dataset
image = ee.Image('USDA/NAIP/DOQQ/m_4609915_sw_14_1_20100629')
Map.addLayer(image, {'bands': ['N', 'R', 'G']}, 'NAIP')
```
## Display Earth Engine data layers
```
Map.addLayerControl() # This line is not needed for ipyleaflet-based Map.
Map
```
| github_jupyter |
## Create Workshop Notebooks and Users
```
import boto3
import botocore
import json
import time
import os
import base64
import docker
import pandas as pd
import project_path # path to helper methods
from lib import workshop
from botocore.exceptions import ClientError
cfn = boto3.client('cloudformation')
iam = boto3.client('iam')
session = boto3.session.Session()
region = session.region_name
account_id = boto3.client('sts').get_caller_identity().get('Account')
```
### [Create S3 Bucket](https://docs.aws.amazon.com/AmazonS3/latest/gsg/CreatingABucket.html)
We will create an S3 bucket that will be used throughout the workshop for storing our data.
[s3.create_bucket](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/s3.html#S3.Client.create_bucket) boto3 documentation
```
bucket = workshop.create_bucket_name('workshop-')
session.resource('s3').create_bucket(Bucket=bucket, CreateBucketConfiguration={'LocationConstraint': region})
print(bucket)
```
### Create the IAM Users and SageMaker Notebooks
We will use CloudFormation to create the workshop users and SageMaker Notebooks to run the workshop.
```
!cat ../research-env.yml
research_file = 'research-env.yml'
session.resource('s3').Bucket(bucket).Object(os.path.join('cfn', research_file)).upload_file( '../' + research_file)
```
### Execute CloudFormation Stack to generate Users and Environment
Creates a stack as specified in the template. After the call completes successfully, the stack creation starts. You can check the status of the stack via the DescribeStacks API.
[cfn.create_stack](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/cloudformation.html#CloudFormation.Client.create_stack)
```
user_count = 2
cfn_template = 'https://s3-{0}.amazonaws.com/{1}/cfn/{2}'.format(region, bucket, research_file)
print(cfn_template)
for x in range(user_count):
user_stack_name = 'Workshop-' + str(x + 1)
response = cfn.create_stack(
StackName=user_stack_name,
TemplateURL=cfn_template,
Capabilities = ["CAPABILITY_NAMED_IAM"],
Parameters=[
{
'ParameterKey': 'NotebookInstanceType',
'ParameterValue': 'ml.t2.medium'
},
{
'ParameterKey': 'UserName',
'ParameterValue': 'WorkshopUser' + str(x + 1)
},
{
'ParameterKey': 'IAMUserPassword',
'ParameterValue': 'WorkshopUser#' + str(x + 1)
}
]
)
print(response)
```
## Clean Up
```
for x in range(user_count):
user_stack_name = 'Workshop-' + str(x + 1)
response = cfn.delete_stack(StackName=user_stack_name)
!aws s3 rb s3://$bucket --force
```
| github_jupyter |
```
import numpy as np
import torch
from torch import nn, optim
import torch.nn.functional as F
from torchvision import datasets
import torchvision.transforms as transforms
from torch.utils.data.sampler import SubsetRandomSampler
from sklearn.metrics import accuracy_score
import matplotlib.pyplot as plt
transform = {
"train": transforms.Compose([
transforms.RandomHorizontalFlip(0.5),
transforms.RandomGrayscale(0.1),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))]),
"test": transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])}
batch_size = 100
train_data = datasets.CIFAR10('data3', train=True, download=True, transform=transform["train"])
test_data = datasets.CIFAR10('data3', train=False, download=True, transform=transform["test"])
dev_size = 0.2
idx = list(range(len(train_data)))
np.random.shuffle(idx)
split_size = int(np.floor(dev_size * len(train_data)))
train_idx, dev_idx = idx[split_size:], idx[:split_size]
train_sampler = SubsetRandomSampler(train_idx)
dev_sampler = SubsetRandomSampler(dev_idx)
train_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, sampler=train_sampler)
dev_loader = torch.utils.data.DataLoader(train_data, batch_size=batch_size, sampler=dev_sampler)
test_loader = torch.utils.data.DataLoader(test_data, batch_size=batch_size)
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.conv1 = nn.Conv2d(3, 10, 3, 1, 1)
self.norm1 = nn.BatchNorm2d(10)
self.conv2 = nn.Conv2d(10, 20, 3, 1, 1)
self.norm2 = nn.BatchNorm2d(20)
self.conv3 = nn.Conv2d(20, 40, 3, 1, 1)
self.norm3 = nn.BatchNorm2d(40)
self.pool = nn.MaxPool2d(2, 2)
self.linear1 = nn.Linear(40 * 4 * 4, 100)
self.norm4 = nn.BatchNorm1d(100)
self.linear2 = nn.Linear(100, 10)
self.dropout = nn.Dropout(0.2)
def forward(self, x):
x = self.pool(self.norm1(F.relu(self.conv1(x))))
x = self.pool(self.norm2(F.relu(self.conv2(x))))
x = self.pool(self.norm3(F.relu(self.conv3(x))))
x = x.view(-1, 40 * 4 * 4)
x = self.dropout(x)
x = self.norm4(F.relu(self.linear1(x)))
x = self.dropout(x)
x = F.log_softmax(self.linear2(x), dim=1)
return x
model = CNN()
loss_function = nn.NLLLoss()
optimizer = optim.Adam(model.parameters(), lr=0.001)
epochs = 100
train_losses, dev_losses, train_acc, dev_acc= [], [], [], []
x_axis = []
for e in range(1, epochs+1):
losses = 0
acc = 0
iterations = 0
model.train()
for data, target in train_loader:
iterations += 1
pred = model(data)
loss = loss_function(pred, target)
optimizer.zero_grad()
loss.backward()
optimizer.step()
losses += loss.item()
p = torch.exp(pred)
top_p, top_class = p.topk(1, dim=1)
acc += accuracy_score(target, top_class)
dev_losss = 0
dev_accs = 0
iter_2 = 0
if e%5 == 0 or e == 1:
x_axis.append(e)
with torch.no_grad():
model.eval()
for data_dev, target_dev in dev_loader:
iter_2 += 1
dev_pred = model(data_dev)
dev_loss = loss_function(dev_pred, target_dev)
dev_losss += dev_loss.item()
dev_p = torch.exp(dev_pred)
top_p, dev_top_class = dev_p.topk(1, dim=1)
dev_accs += accuracy_score(target_dev, dev_top_class)
train_losses.append(losses/iterations)
dev_losses.append(dev_losss/iter_2)
train_acc.append(acc/iterations)
dev_acc.append(dev_accs/iter_2)
print("Epoch: {}/{}.. ".format(e, epochs),
"Training Loss: {:.3f}.. ".format(losses/iterations),
"Validation Loss: {:.3f}.. ".format(dev_losss/iter_2),
"Training Accuracy: {:.3f}.. ".format(acc/iterations),
"Validation Accuracy: {:.3f}".format(dev_accs/iter_2))
plt.plot(x_axis,train_losses, label='Training loss')
plt.plot(x_axis, dev_losses, label='Validation loss')
plt.legend(frameon=False)
plt.show()
plt.plot(x_axis, train_acc, label="Training accuracy")
plt.plot(x_axis, dev_acc, label="Validation accuracy")
plt.legend(frameon=False)
plt.show()
model.eval()
iter_3 = 0
acc_test = 0
for data_test, target_test in test_loader:
iter_3 += 1
test_pred = model(data_test)
test_pred = torch.exp(test_pred)
top_p, top_class_test = test_pred.topk(1, dim=1)
acc_test += accuracy_score(target_test, top_class_test)
print(acc_test/iter_3)
```
| github_jupyter |
# Day 4 - brute-force password generation
Because the digits must be the same or increase, the number of combinations is fairly limited. The total number of such 'passwords' is a [*6-simplex polytopic numbers*](https://en.wikipedia.org/wiki/Figurate_number#Triangular_numbers), the value of which can be calculated as
$$\dbinom{10 + 6 - 1}{6} = \dbinom{15}{6} = 5005$$
(where `10` is the number of digits and `6` the number of dimensions). We can easily brute-force this by generating all the possible combinations of increasing digits, using a recursive loop over a digits string (which we shorten for the next recursive call to ensure digits only increase).
The passwords can then be checked to be within the low-high value range, and the number of unique digits needs to be less than 6 for digits to have repeated.
```
from __future__ import annotations
from typing import Callable, Iterable
Checker = Callable[[int], bool]
def produce(n: int = 6, digits: str = '0123456789') -> Iterable[str]:
if n == 0:
yield ''
return
for i, d in enumerate(digits):
for remainder in produce(n - 1, digits[i:]):
yield d + remainder
def password_checker_factory(lo: int, hi: int) -> Checker:
def is_valid(pw: int) -> bool:
return (lo <= pw <= hi) and len(set(str(pw))) < 6
return is_valid
def count_valid(checker: Checker) -> int:
return sum(1 for _ in filter(checker, map(int, produce())))
tests = {
(111111, 111111): 1,
(223450, 223450): 0,
(123789, 123789): 0,
}
for (lo, hi), expected in tests.items():
assert count_valid(password_checker_factory(lo, hi)) == expected
import aocd
lo, hi = map(int, aocd.get_data(day=4, year=2019).strip().split("-"))
print("Part 1:", count_valid(password_checker_factory(lo, hi)))
```
## Part 2
This is just a slightly stricter checker. Instead of the number of unique digits, we need to count consecutive digits and assert there is a group of length 2. This is a job for [`itertools.groupby()`](https://docs.python.org/3/library/itertools.html#itertools.groupby)!
```
from itertools import groupby
def stricter_password_checker_factory(lo: int, hi: int) -> Checker:
def has_2_adjacent(pw: int):
return any(sum(1 for _ in g) == 2 for _, g in groupby(str(pw)))
def is_valid(pw: int):
return (lo <= pw <= hi) and has_2_adjacent(pw)
return is_valid
strict_tests = {112233: True, 123444: False, 111122: True}
for pw, expected in strict_tests.items():
assert stricter_password_checker_factory(pw, pw)(pw) == expected
print("Part 2:", count_valid(stricter_password_checker_factory(lo, hi)))
```
| github_jupyter |
```
import csv
import logging
logging.basicConfig(format='%(message)s')
import requests
from bs4 import BeautifulSoup
import time
import re
import json
import datetime
def scrape_review(eigacom_id):
page_num = 1
data = {
"title" : -1,
"reviews" : []
}
rating_dict = {"val00":0.0, "val05":0.5,"val10":1.0,"val15":1.5,"val20":2.0,"val25":2.5,"val30":3.0,"val35":3.5,"val40":4.0,"val45":4.5,"val50":5.0}
print("START : " + eigacom_id)
url_review='https://eiga.com/movie/' + eigacom_id + '/review/all/'
if url_review is None:
logging.warning("**************************************************")
logging.warning(q + " HAS NO RESULT")
logging.warning("**************************************************")
return None
while(1):
res = requests.get(url_review + str(page_num))
res.encoding = res.apparent_encoding
soup = BeautifulSoup(res.content, "lxml")
if page_num == 1:
title = soup.find('p', attrs={"class":"title-link"}).text
data["title"] = title
if soup.find('div', attrs={"class": "user-review"}) == None: # ページ数の上限を超えたら
print('DONE : ' + eigacom_id )
break
for r in soup.find_all('div', attrs={"class": "user-review"}):
review_title = r.find('h2',attrs={"class": "review-title"})
title = review_title.find('a')
rating_class = review_title.find('span',attrs={"class": "rating-star"}).get('class')[1]
rating = rating_dict[rating_class]
empathy = r.find('div', attrs={"class": "empathy"}).find(('strong')).text
date= r.find('div',attrs={"class": "review-data"}).find('div',attrs={"class": "time"})
main_text = r.find('div',attrs={"class": "txt-block"})
tgl_btn = main_text.find('div',attrs={"class": "toggle-btn"})
if tgl_btn is not None:
tgl_btn.decompose()
item = {
"date" : "",
"rating" : rating,
"empathy" : int(empathy),
"review" : "",
}
review_text = title.text + "\n" + main_text.text.replace("\n", "")
item["review"] = review_text
y, m, d, _ = re.split('[年月日]', date.text)
item["date"] = str(datetime.date(int(y), int(m), int(d)))
data["reviews"].append(item)
page_num += 1
time.sleep(1)
return data
def main():
data_all = {}
movie_id = 1
for year in range(1978, 2020):
print(year)
with open('./eigacom_nomination_id_table/{}.txt'.format(str(year)), 'r') as id_table:
for line in csv.reader(id_table):
if line == "\n":
continue
eigacom_id, *_ = line
print(movie_id)
data = scrape_review(eigacom_id)
if data == None:
movie_id += 1
continue
data_all[str(movie_id)] = data
movie_id += 1
output_file = '../../data/eigacom_review.json'
with open(output_file, 'w') as f:
json.dump(data_all, f, ensure_ascii=False, indent=2)
if __name__ == "__main__":
main()
```
| github_jupyter |
```
import warnings
warnings.filterwarnings('ignore')
import os
import cv2
import keras
from keras.models import Sequential, load_model, Model
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
pre_model = load_model('./saved_models/Inceptionv3_1.h5')
# test_path = 'test/twmoth009748.jpg'
test_path = 'test/twmoth020247.jpg'
img = cv2.resize(cv2.imread(test_path), dsize = (299,299))
img = img.astype(np.float32)
img /= 127.5
img -= 1
img.shape
img = np.expand_dims(img, axis=0)
img.shape
pred = pre_model.predict(img)
pred
img_labels = ['Bombycidae_Bombyx_mandarina_formosana',
'Bombycidae_Ernolatia_moorei',
'Bombycidae_Trilocha_varians',
'Bombycidae_Triuncina_brunnea',
'Brahmaeidae_Brahmaea_wallichii_insulata',
'Callidulidae_Callidula_attenuata',
'Choreutidae_Choreutis_amethystodes',
'Choreutidae_Choreutis_basalis',
'Choreutidae_Choreutis_xanthogramma',
'Choreutidae_Saptha_divitiosa',
'Cossidae_Xyleutes_strix',
'Cossidae_Zeuzera_multistrigata',
'Crambidae_Aethaloessa_calidalis_tiphalis',
'Crambidae_Agrioglypta_itysalis',
'Crambidae_Bocchoris_inspersalis',
'Crambidae_Botyodes_caldusalis',
'Crambidae_Bradina_diagonalis',
'Crambidae_Camptomastix_hisbonalis',
'Crambidae_Cataclysta_angulata',
'Crambidae_Ceratarcha_umbrosa',
'Crambidae_Cirrhochrista_spinuella',
'Crambidae_Cirrhochrista_spissalis',
'Crambidae_Cnaphalocrocis_medinalis',
'Crambidae_Diaphania_indica',
'Crambidae_Eoophyla_conjunctalis',
'Crambidae_Eoophyla_gibbosalis',
'Crambidae_Glyphodes_bivitralis',
'Crambidae_Heortia_vitessoides',
'Crambidae_Maruca_vitrata',
'Crambidae_Nevrina_procopia',
'Crambidae_Paracymoriza_cataclystalis',
'Crambidae_Spoladea_recurvalis',
'Crambidae_Syllepte_taiwanalis',
'Drepanidae_Callidrepana_patrana',
'Drepanidae_Cyclidia_substigmaria',
'Drepanidae_Drapetodes_mitaria',
'Drepanidae_Habrosyne_indica_formosana',
'Drepanidae_Horithyatira_decorata_takamukui',
'Drepanidae_Leucoblepsis_taiwanensis',
'Drepanidae_Leucodrepana_serratilinea',
'Drepanidae_Macrocilix_mysticata_flavotincta',
'Drepanidae_Neoreta_purpureofascia',
'Drepanidae_Nordstromia_semililacina',
'Drepanidae_Oreta_brunnea',
'Drepanidae_Oreta_extensa',
'Drepanidae_Oreta_insignis',
'Drepanidae_Thyatira_batis_formosicola',
'Endromidae_Andraca_theae',
'Erebidae_Aglaomorpha_histrio_formosana',
'Erebidae_Amsactoides_solitaria',
'Erebidae_Anisoneura_salebrosa',
'Erebidae_Areas_galactina_formosana',
'Erebidae_Arna_bipunctapex',
'Erebidae_Aroana_baliensis',
'Erebidae_Artena_dotata',
'Erebidae_Asota_caricae',
'Erebidae_Asota_egens_indica',
'Erebidae_Asota_heliconia_zebrina',
'Erebidae_Asota_plana_lacteata',
'Erebidae_Asura_tricolor',
'Erebidae_Avatha_chinensis',
'Erebidae_Barsine_fuscozonata',
'Erebidae_Barsine_horishanella',
'Erebidae_Barsine_sauteri',
'Erebidae_Bastilla_fulvotaenia',
'Erebidae_Bertula_abjudicalis',
'Erebidae_Bertula_kosemponica',
'Erebidae_Bocana_manifestalis',
'Erebidae_Brunia_antica',
'Erebidae_Calliteara_grotei_horishanella',
'Erebidae_Calliteara_postfusca',
'Erebidae_Chrysaeglia_magnifica',
'Erebidae_Conilepia_nigricosta_paiwan',
'Erebidae_Creatonotos_transiens_vacillans',
'Erebidae_Cyana_hamata_hamata',
'Erebidae_Daddala_lucilla',
'Erebidae_Diduga_flavicostata',
'Erebidae_Dura_alba',
'Erebidae_Ercheia_cyllaria',
'Erebidae_Erebus_ephesperis',
'Erebidae_Erebus_gemmans',
'Erebidae_Eressa_confinis_finitima',
'Erebidae_Eudocima_homaena',
'Erebidae_Eudocima_phalonia',
'Erebidae_Eudocima_salaminia',
'Erebidae_Euproctis_kanshireia',
'Erebidae_Fodina_contigua',
'Erebidae_Garudinia_bimaculata',
'Erebidae_Herminia_vermiculata',
'Erebidae_Hesudra_divisa',
'Erebidae_Hipoepa_fractalis',
'Erebidae_Hydrillodes_lentalis',
'Erebidae_Hydrillodes_nilgirialis',
'Erebidae_Hypospila_bolinoides',
'Erebidae_Ilema_kosemponica',
'Erebidae_Ilema_olivacea',
'Erebidae_Ischyja_ferrifracta',
'Erebidae_Ischyja_manlia',
'Erebidae_Lemyra_alikangensis',
'Erebidae_Lemyra_rhodophilodes',
'Erebidae_Lyclene_alikangiae',
'Erebidae_Lyclene_arcuata',
'Erebidae_Lymantria_concolor_concolor',
'Erebidae_Lymantria_iris',
'Erebidae_Lymantria_mathura_subpallida',
'Erebidae_Lymantria_sinica_sinica',
'Erebidae_Lymantria_xylina',
'Erebidae_Macrobrochis_gigas',
'Erebidae_Metaemene_atrigutta',
'Erebidae_Miltochrista_ziczac',
'Erebidae_Mithuna_arizana',
'Erebidae_Mocis_frugalis',
'Erebidae_Mocis_undata',
'Erebidae_Mosopia_punctilinea',
'Erebidae_Nudaria_ranruna',
'Erebidae_Nyctemera_adversata',
'Erebidae_Nyctemera_carissima_formosana',
'Erebidae_Nyctemera_lacticinia',
'Erebidae_Olene_dudgeoni',
'Erebidae_Olene_mendosa',
'Erebidae_Ommatophora_luminosa',
'Erebidae_Orgyia_postica',
'Erebidae_Oxacme_cretacea',
'Erebidae_Oxyodes_scrobiculata',
'Erebidae_Perina_nuda',
'Erebidae_Pindara_illibata',
'Erebidae_Schistophleps_bipuncta',
'Erebidae_Somena_scintillans',
'Erebidae_Spilarctia_subcarnea',
'Erebidae_Spilarctia_wilemani',
'Erebidae_Sympis_rufibasis',
'Erebidae_Syntomoides_imaon',
'Erebidae_Thyas coronata',
'Erebidae_Thysanoptyx_incurvata',
'Erebidae_Vamuna alboluteora',
'Eupterotidae_Palirisa_cervina_formosana',
'Geometridae_Acolutha pictaria_imbecilla',
'Geometridae_Agathia laetata',
'Geometridae_Alcis admissaria_undularia',
'Geometridae_Alcis taiwanovariegata',
'Geometridae_Amblychia_angeronaria',
'Geometridae_Antitrygodes_divisaria_perturbatus',
'Geometridae_Biston_perclara',
'Geometridae_Borbacha_pardaria',
'Geometridae_Calletaera_obliquata',
'Geometridae_Calletaera_postvittata',
'Geometridae_Catoria_olivescens',
'Geometridae_Catoria_sublavaria',
'Geometridae_Celenna_festivaria_formosensis',
'Geometridae_Chiasmia_emersaria',
'Geometridae_Chorodna_creataria',
'Geometridae_Chorodna_ochreimacula',
'Geometridae_Cleora_fraterna',
'Geometridae_Corymica_arnearia',
'Geometridae_Cusiala_boarmioides',
'Geometridae_Doratoptera_lutea',
'Geometridae_Ectropis_bhurmitra',
'Geometridae_Entomopteryx_rubridisca',
'Geometridae_Epobeidia_lucifera_extranigricans',
'Geometridae_Eumelea _udovicata',
'Geometridae_Gonodontis_pallida',
'Geometridae_Harutalcis_fumigata',
'Geometridae_Herochroma_cristata',
'Geometridae_Hypochrosis_baenzigeri',
'Geometridae_Hyposidra_infixaria',
'Geometridae_Hyposidra_talaca_talaca',
'Geometridae_Krananda semihyalina',
'Geometridae_Krananda_latimarginaria',
'Geometridae_Krananda_oliveomarginata',
'Geometridae_Lassaba_parvalbidaria_parvalbidaria',
'Geometridae_Lophobates_inchoata',
'Geometridae_Lophophelma_taiwana',
'Geometridae_Luxiaria_mitorrhaphes',
'Geometridae_Nothomiza_flavicosta',
'Geometridae_Odontopera_albiguttulata',
'Geometridae_Ophthalmitis_herbidaria',
'Geometridae_Organopoda_carnearia_carnearia',
'Geometridae_Ourapteryx_clara_formosana',
'Geometridae_Pachyodes_subtrita',
'Geometridae_Parapercnia_giraffata',
'Geometridae_Percnia_longitermen',
'Geometridae_Percnia_suffusa',
'Geometridae_Phthonandria_atrilineata_cuneilinearia',
'Geometridae_Pingasa_ruginaria_pacifica',
'Geometridae_Pogonopygia_pavidus_pavidus',
'Geometridae_Problepsis_albidior_matsumurai',
'Geometridae_Psilalcis_breta_rantaizana',
'Geometridae_Psilalcis_pulveraria',
'Geometridae_Racotis_boarmiaria',
'Geometridae_Scopula_propinquaria',
'Geometridae_Tanaoctenia_haliaria',
'Geometridae_Timandra_convectaria',
'Geometridae_Timandra_extremaria',
'Geometridae_Traminda_aventiaria',
'Geometridae_Trotocraspeda_divaricata',
'Geometridae_Xandrames_dholaria',
'Geometridae_Xandrames_latiferaria_curvistriga',
'Geometridae_Xanthorhoe_saturata',
'Geometridae_Xenoplia_trivialis',
'Geometridae_Yashmakia_suffusa',
'Geometridae_Zanclopera_falcata',
'Hyblaeidae_Hyblaea_firmamentum',
'Lasiocampidae_Kunugia_undans_metanastroides',
'Lasiocampidae_Lebeda_nobilis',
'Lasiocampidae_Paralebeda_femorata_mirabilis',
'Lasiocampidae_Trabala_vishnou_guttata',
'Lecithoceridae_Tisis_mesozosta',
'Limacodidae_Cania_heppneri',
'Limacodidae_Miresa_fulgida',
'Limacodidae_Monema_rubriceps',
'Limacodidae_Parasa_consocia',
'Limacodidae_Parasa_shirakii',
'Limacodidae_Phlossa_conjuncta',
'Limacodidae_Susica_sinensis',
'Limacodidae_Thosea_sinensis',
'Noctuidae_Diphtherocome_pulchra',
'Noctuidae_Exsula dentatrix_albomaculata',
'Nolidae_Westermannia_elliptica_elliptica',
'Notodontidae_Benbowia_takamukuanus',
'Notodontidae_Phalera_flavescens_flavescens',
'Notodontidae_Syntypistis_pallidifascia_pallidifascia',
'Pyralidae_Arctioblepsis_rubida',
'Pyralidae_Locastra_muscosalis',
'Saturniidae_Actias_ningpoana_ningtaiwana',
'Saturniidae_Actias_sinensis_subaurea',
'Saturniidae_Antheraea_formosana',
'Saturniidae_Antheraea_yamamai_superba',
'Saturniidae_Attacus_atlas_formosanus',
'Saturniidae_Loepa_formosensis',
'Saturniidae_Samia_wangi',
'Saturniidae_Saturnia_thibeta_okurai',
'Sphingidae_Acherontia_lachesis',
'Sphingidae_Acosmerycoides_harterti',
'Sphingidae_Acosmeryx_castanea',
'Sphingidae_Acosmeryx_naga_naga',
'Sphingidae_Agrius_convolvuli',
'Sphingidae_Callambulyx_tatarinovii_formosana',
'Sphingidae_Cechetra_lineosa',
'Sphingidae_Cechetra_minor',
'Sphingidae_Clanis_bilineata_formosana',
'Sphingidae_Cypoides chinensis',
'Sphingidae_Dolbina_inexacta',
'Sphingidae_Langia_zenzeroides_formosana',
'Sphingidae_Marumba_cristata_bukaiana',
'Sphingidae_Marumba_sperchius_sperchius',
'Sphingidae_Notonagemia_analis_gressitti',
'Sphingidae_Parum_colligata',
'Sphingidae_Pergesa_acteus',
'Sphingidae_Psilogramma_increta',
'Sphingidae_Rhagastis_binoculata',
'Sphingidae_Rhagastis_castor_formosana',
'Sphingidae_Theretra_nessus',
'Thyrididae_Pyrinioides_sinuosus',
'Uraniidae_Acropteris_leptaliata',
'Uraniidae_Dysaethria_cretacea',
'Uraniidae_Dysaethria_flavistriga',
'Zygaenidae_Artona_hainana',
'Zygaenidae_Clelea_formosana',
'Zygaenidae_Erasmia_pulchella_hobsoni',
'Zygaenidae_Eterusia_aedea_formosana',
'Zygaenidae_Gynautocera_rubriscutellata',
'Zygaenidae_Histia_flabellicornis_ultima']
pred_class = np.argmax(pred)
pred_class, img_labels[pred_class]
class_sort = (-pred).argsort()
top5 = class_sort[0][:5]
top5
print(test_path)
ord_nums =['1st', '2nd', '3rd', '4th', '5th']
file_str = test_path.split('/')[1].split('.')[0]
result = file_str+'.txt'
f = open(result, 'w')
for i in range(len(top5)):
ename = img_labels[top5[i]].split('_')
sname = str()
for x in ename[1:]:
sname = sname +' '+x
#name = '科名:'+ename[0]+' '+'屬名:'+' '+ename[1]+' '+'學名:'+sname
name = ' 科名: {} 屬名: {} 學名:{}'.format(ename[0], ename[1], sname)
accu = str(round(pred[0][top5[i]]*100, 2))+'%'
name_str = '{}{}{}\n'.format(ord_nums[i], name, accu)
f.write(name_str)
print (ord_nums[i], name, accu)
#print (name_str)
#print(ord_nums[i],img_labels[top5[i]], str(round(pred[0][top5[i]]*100, 2))+'%')
pred[0][201]
#name = '科名:{} 屬名:{} 學名: {}'.format(ename[0], ename[1], sname)
#u1, u2 = test_path.split('.')
#result = u1+'.txt'
#f = open(result, 'w')
#name_str = '{}{}{}\n'.format(ord_nums[i], name, accu)
#f.write( name_str)
```
| github_jupyter |
```
# Standard imports
import matplotlib.pyplot as plt
from matplotlib.gridspec import GridSpec
import numpy as np
from scipy.stats import linregress
%config InlineBackend.figure_format = 'retina'
seed = np.random.randint(12345)
np.random.seed(seed)
#import pingouin
from scipy.stats import pearsonr
# Useful helper function
r_squared = lambda x,y: linregress(x.ravel().T, y.ravel().T)[2]**2
# def r2_sigma(y, yhat, number_trials=1000, size_drawn_frac=0.25):
# number_data = len(y)
# Rsq_bs = np.zeros(shape=number_trials)
# size_of_drawn_samples = int(size_drawn_frac*number_data)
# for trial in range(number_trials):
# # sample frac (size_drawn_frac) the data with replacement
# ix = np.random.choice(number_data, size=size_of_drawn_samples, replace=True)
# # compute R^2
# Rsq_bs[trial] = np.corrcoef(y[ix], yhat[ix])[0][1]**2
# return np.std(Rsq_bs)/np.sqrt(size_of_drawn_samples)
# Write function to compute bootstrap resampled uncertainties in correlation coefficient.
def my_rsquared(x,y, bootstrap_samples=1000):
"""Returns R^2 and SE thereof based on bootstrap resampling"""
r2 = pearsonr(x,y)[0]**2 # pingouin.corr(x, y).loc['pearson','r']**2
N = len(x)
assert len(x)==len(y), f'len(x)={len(x)} and len(y)={len(y)} are not the same.'
r2s = np.zeros(bootstrap_samples)
for i in range(bootstrap_samples):
ix = np.random.choice(a=bootstrap_samples, size=bootstrap_samples, replace=True)
r2s[i] = pearsonr(x[ix],y[ix])[0]**2 #pingouin.corr(x[ix], y[ix]).loc['pearson','r']**2
dr2 = np.std(r2s)
return r2, dr2
style_file_name = 'fig4.style'
s = """
axes.linewidth: 0.5 # edge linewidth
font.size: 7.0
axes.labelsize: 7.0 # fontsize of the x any y labels
xtick.labelsize: 7.0 # fontsize of the tick labels
ytick.labelsize: 7.0 # fontsize of the tick labels
legend.fontsize: 7.0
legend.borderpad: 0.2 # border whitespace
legend.labelspacing: 0.2 # the vertical space between the legend entries
legend.borderaxespad: 0.2 # the border between the axes and legend edge
legend.framealpha: 1.0
"""
with open(style_file_name, 'w') as f:
f.write(s)
plt.style.use(style_file_name)
plt.rc('font', family='sans-serif')
plt.rc('font', family='sans-serif')
fig_si_npz = np.load('fig4_SI_dict.npz')
y_test_ABeta=fig_si_npz['y_test_ABeta']
yhat_test_ABeta=fig_si_npz['yhat_test_ABeta']
yqs_grid_ABeta=fig_si_npz['yqs_grid_ABeta']
yhat_test_TDP=fig_si_npz['yhat_test_TDP']
y_test_TDP=fig_si_npz['y_test_TDP']
yqs_grid_TDP=fig_si_npz['yqs_grid_TDP']
# Model predictions for wt sequences
phi_wt_ABeta = 1.1025
yhat_wt_ABeta = 0.12027
phi_wt_TDP = -0.01602
yhat_wt_TDP = -0.0022342
fig = plt.figure(figsize=[6.5, 3])
plt.style.use(style_file_name)
gs = fig.add_gridspec(1, 2)
# Define panels
ax_a = fig.add_subplot(gs[0, 0])
ax_b = fig.add_subplot(gs[0, 1])
alpha = 0.2
# Panel A: Scatter plot ABeta
q = [.025,.975]
ylim = [-6, 3]
yticks = [-6, -5, -4, -3, -2, -1, 0, 1, 2, 3]
ax = ax_a
yhat_test = yhat_test_ABeta
xlim = [min(yhat_test), ylim[1]]
yhat_grid = np.linspace(ylim[0], ylim[1], 1000)
yqs_grid = yqs_grid_ABeta
ax.scatter(yhat_test,
y_test_ABeta,
s=2,
alpha=alpha+0.1,
color='C0',
label='test data')
ax.set_xticks(yticks)
ax.set_yticks(yticks)
ax.set_xlabel('model prediction ($\hat{y}$)')
ax.set_ylabel('nucleation score ($y$)', labelpad=10)
ax.set_xlim(ylim)
ax.set_ylim(ylim)
ax.plot(xlim, xlim, linestyle='-', color='C1', linewidth=2, label='$y = \hat{y}$')
ax.plot(yhat_grid, yqs_grid[:,0], linestyle=':', color='C1', linewidth=1, label='95% CI')
ax.plot(yhat_grid, yqs_grid[:,1], linestyle=':', color='C1', linewidth=1)
# draw wt phi
ax.axvline(yhat_wt_ABeta, color='lightgray', zorder=-1, label='WT $\hat{y}$')
leg = ax.legend(loc='lower right')
for lh in leg.legendHandles:
lh.set_alpha(1)
# Compute R^2 and Is
r2, r2_err = my_rsquared(yhat_test, y_test_ABeta)
#ax.text(x=-5.9, y=2.6, s=f'$R^2 =$ {r2:.3f} $\pm$ {r2_err:.3f}',
# uncertainty computed in script in figure 4b from the main text
ax.text(x=-5.9, y=2.6, s=f'$R^2 =$ {r2:.3f} $\pm$ 0.071',
ha='left', va='center');
# Panel B: Scatter plot TDP-43
ylim = [-0.3, 0.4]
yticks = [-0.3, -0.2, -0.1, 0, 0.1, 0.2, 0.3, 0.4]
ax = ax_b
# Select 2000 random point for better visuzlization
ix = np.random.choice(y_test_TDP.shape[0], 2000, replace=False)
yhat_test = yhat_test_TDP
xlim = [min(yhat_test), ylim[1]]
yhat_grid = np.linspace(ylim[0], ylim[1], 1000)
yqs_grid = yqs_grid_TDP
ax.scatter(yhat_test[ix],
y_test_TDP[ix],
s=2,
alpha=alpha,
color='C0',
label='test data')
ax.set_xticks(yticks)
ax.set_yticks(yticks)
ax.set_xlabel('model prediction ($\hat{y}$)')
ax.set_ylabel('toxicity score ($y$)')
ax.set_xlim(ylim)
ax.set_ylim(ylim)
ax.plot(xlim, xlim, linestyle='-', color='C1', linewidth=2, label='$y = \hat{y}$')
ax.plot(yhat_grid, yqs_grid[:,0], linestyle=':', color='C1', linewidth=1, label='95% CI')
ax.plot(yhat_grid, yqs_grid[:,1], linestyle=':', color='C1', linewidth=1)
# draw wt phi
ax.axvline(yhat_wt_TDP, color='lightgray', zorder=-1, label='WT $\hat{y}$')
leg = ax.legend(loc='lower right')
for lh in leg.legendHandles:
lh.set_alpha(1)
# Compute R^2 and I's
r2, r2_err = my_rsquared(yhat_test, y_test_TDP)
yhat_test_TDP = yhat_test
yqs_grid_TDP = yqs_grid
ax.text(x=-0.29, y=0.37,
#s=f'$R^2 =$ {r2:.3f} $\pm$ {r2_err:.3f}',
# uncertainty computed in script for Figure 4d
# from the main text using corrected bootsrap
s=f'$R^2 =$ {r2:.3f} $\pm$ 0.052',
ha='left', va='center');
# Add panel labels
fig.text(0.02, 0.95, 'a', fontsize=11.5, fontweight='bold')
fig.text(0.51, 0.95, 'b', fontsize=11.5, fontweight='bold')
# Clean up and save
fig.tight_layout()
fig.savefig('fig4_supp.png', dpi=400, facecolor='w')
```
| github_jupyter |
# Create your first deep learning neural network
## Introduction
This is the first of our [beginner tutorial series](https://github.com/awslabs/djl/tree/master/jupyter/tutorial) that will take you through creating, training, and running inference on a neural network. In this tutorial, you will learn how to use the built-in `Block` to create your first neural network - a Multilayer Perceptron.
## Neural Network
A neural network is a black box function. Instead of coding this function yourself, you provide many sample input/output pairs for this function. Then, we try to train the network to learn how to match the behavior of the function given only these input/output pairs. A better model with more data can more accurately match the function.
## Multilayer Perceptron
A Multilayer Perceptron (MLP) is one of the simplest deep learning networks. The MLP has an input layer which contains your input data, an output layer which is produced by the network and contains the data the network is supposed to be learning, and some number of hidden layers. The example below contains an input of size 3, a single hidden layer of size 3, and an output of size 2. The number and sizes of the hidden layers are determined through experimentation but more layers enable the network to represent more complicated functions. Between each pair of layers is a linear operation (sometimes called a FullyConnected operation because each number in the input connected to each number in the output by a matrix multiplication). Not pictured, there is also a non-linear activation function after each linear operation. For more information, see [Multilayer Perceptron](https://en.wikipedia.org/wiki/Multilayer_perceptron).

## Step 1: Setup development environment
### Installation
This tutorial requires the installation of the Java Jupyter Kernel. To install the kernel, see the [Jupyter README](https://github.com/awslabs/djl/blob/master/jupyter/README.md).
```
// Add the snapshot repository to get the DJL snapshot artifacts
// %mavenRepo snapshots https://oss.sonatype.org/content/repositories/snapshots/
// Add the maven dependencies
%maven ai.djl:api:0.6.0
%maven org.slf4j:slf4j-api:1.7.26
%maven org.slf4j:slf4j-simple:1.7.26
// See https://github.com/awslabs/djl/blob/master/mxnet/mxnet-engine/README.md
// for more MXNet library selection options
%maven ai.djl.mxnet:mxnet-native-auto:1.7.0-a
import ai.djl.*;
import ai.djl.nn.*;
import ai.djl.nn.core.*;
import ai.djl.training.*;
```
## Step 2: Determine your input and output size
The MLP model uses a one dimensional vector as the input and the output. You should determine the appropriate size of this vector based on your input data and what you will use the output of the model for. In a later tutorial, we will use this model for Mnist image classification.
Our input vector will have size `28x28` because the input images have a height and width of 28 and it takes only a single number to represent each pixel. For a color image, you would need to further multiply this by `3` for the RGB channels.
Our output vector has size `10` because there are `10` possible classes for each image.
```
long inputSize = 28*28;
long outputSize = 10;
```
## Step 3: Create a **SequentialBlock**
### NDArray
The core data type used for working with Deep Learning is the [NDArray](https://javadoc.io/static/ai.djl/api/0.6.0/index.html?ai/djl/ndarray/NDArray.html). An NDArray represents a multidimensional, fixed-size homogeneous array. It has very similar behavior to the Numpy python package with the addition of efficient computing. We also have a helper class, the [NDList](https://javadoc.io/static/ai.djl/api/0.6.0/index.html?ai/djl/ndarray/NDList.html) which is a list of NDArrays which can have different sizes and data types.
### Block API
In DJL, [Blocks](https://javadoc.io/static/ai.djl/api/0.6.0/index.html?ai/djl/nn/Block.html) serve a purpose similar to functions that convert an input `NDList` to an output `NDList`. They can represent single operations, parts of a neural network, and even the whole neural network. What makes blocks special is that they contain a number of parameters that are used in their function and are trained during deep learning. As these parameters are trained, the function represented by the blocks get more and more accurate.
When building these block functions, the easiest way is to use composition. Similar to how functions are built by calling other functions, blocks can be built by combining other blocks. We refer to the containing block as the parent and the sub-blocks as the children.
We provide several helpers to make it easy to build common block composition structures. For the MLP we will use the [SequentialBlock](https://javadoc.io/static/ai.djl/api/0.6.0/index.html?ai/djl/nn/SequentialBlock.html), a container block whose children form a chain of blocks where each child block feeds its output to the next child block in a sequence.
```
SequentialBlock block = new SequentialBlock();
```
## Step 4: Add blocks to SequentialBlock
An MLP is organized into several layers. Each layer is composed of a [Linear Block](https://javadoc.io/static/ai.djl/api/0.6.0/index.html?ai/djl/nn/core/Linear.html) and a non-linear activation function. If we just had two linear blocks in a row, it would be the same as a combined linear block ($f(x) = W_2(W_1x) = (W_2W_1)x = W_{combined}x$). An activation is used to intersperse between the linear blocks to allow them to represent non-linear functions. We will use the popular [ReLU](https://javadoc.io/static/ai.djl/api/0.6.0/ai/djl/nn/Activation.html#reluBlock--) as our activation function.
The first layer and last layers have fixed sizes depending on your desired input and output size. However, you are free to choose the number and sizes of the middle layers in the network. We will create a smaller MLP with two middle layers that gradually decrease the size. Typically, you would experiment with different values to see what works the best on your data set.
```
block.add(Blocks.batchFlattenBlock(inputSize));
block.add(Linear.builder().setOutChannels(128).build());
block.add(Activation::relu);
block.add(Linear.builder().setOutChannels(64).build());
block.add(Activation::relu);
block.add(Linear.builder().setOutChannels(outputSize).build());
block
```
## Summary
Now that you've successfully created your first neural network, you can use this network to train your model.
Next chapter: [Train your first model](train_your_first_model.ipynb)
You can find the complete source code for this tutorial in the [model zoo](https://github.com/awslabs/djl/blob/master/model-zoo/src/main/java/ai/djl/basicmodelzoo/basic/Mlp.java).
| github_jupyter |
# 08: SparkSQL
Originally, SparkSQL was an extension of RDDs with the concept of a `DataFrame` that adds a "schema" for records, defined using Scala _case classes_, tuples, or a built-in schema mechanism. The DataFrame API is inspired by similar `DataFrame` concepts in R and Python libraries. The transformation and action steps written in any of the support languages, as well as SQL queries embedded in strings, are translated to the same, performant query execution model, optimized by a new query engine called *Catalyst*.
The even newer `Dataset` API encapsulates `DataFrame`, but adds more type safety for the data columns. We'll stick with the `DataFrame` here.
> **Tip:** Even if you prefer the Scala collections-like `RDD` API, I recommend using the `DataFrame` API, because the performance is _significantly_ better in most cases, due to internal optimizations.
Furthermore, SparkSQL has convenient support for reading and writing files encoded using [Parquet](http://parquet.io), [ORC](https://orc.apache.org/), JSON, CSV, etc.
Finally, SparkSQL embeds access to a Hive _metastore_, so you can create and delete tables, and run queries against them using SparkSQL.
This example treats the KJV text we've been using as a table with a schema. It runs several SQL queries on the data, then performs the same calculation using the `DataFrame` API.
See the corresponding Spark job [SparkSQL8.scala](https://github.com/deanwampler/spark-scala-tutorial/blob/master/src/main/scala/sparktutorial/SparkSQL8.scala) and the "script" suitable for _spark-shell_, [SparkSQL8-script.scala](https://github.com/deanwampler/spark-scala-tutorial/blob/master/src/main/scala/sparktutorial/SparkSQL8-script.scala), because SQL queries are nice to use interactively!
```
val in = "../data/kjvdat.txt" // '|' separated
val abbrevToNames = "../data/abbrevs-to-names.tsv" // tab separated
```
This time, we won't use the `toText` method we defined before, we'll use a big regex and do pattern matching with it.
```
val lineRE = """^\s*([^|]+)\s*\|\s*([\d]+)\s*\|\s*([\d]+)\s*\|\s*(.*)~?\s*$""".r
```
Let's define a case class `Verse` to represent our records.
```
case class Verse(book: String, chapter: Int, verse: Int, text: String)
```
Now load and parse the data. Note that using `flatMap` effective removes the bad records, for which we return `Nil`, while we return `Seq(verse)` on success.
```
val versesRDD = sc.textFile(in).flatMap {
case lineRE(book, chapter, verse, text) =>
Seq(Verse(book, chapter.toInt, verse.toInt, text))
case line =>
Console.err.println(s"Unexpected line: $line")
Nil // or use Seq.empty[Verse]. It will be eliminated by flattening.
}
```
Now create a `DataFrame` from this `RDD` and create a temporary "view". We're going to reuse this data over and over, so now it's very useful to _cache_ it in memory.
```
val verses = spark.createDataFrame(versesRDD)
verses.createOrReplaceTempView("kjv_bible")
verses.cache()
```
Print the first 20 lines (the default), or pass an integer argument to show a different number
of lines, as here:
```
verses.show(10)
```
Pass an optional `truncate = false` argument for wider output.
```
verses.show(5, truncate = false)
```
Now write some SQL queries, with SQL and with the Dataset/DataFrame API!
```
val godVerses = spark.sql("SELECT * FROM kjv_bible WHERE text LIKE '%God%'")
println("Number of verses that mention God: "+godVerses.count())
godVerses.show(truncate=false)
println("The query plan:")
godVerses.queryExecution // Compare with godVerses.explain(true)
```
Now using the API:
```
val godVersesDF = verses.filter(verses("text").contains("God"))
println("Number of verses that mention God: "+godVersesDF.count())
godVersesDF.show()
```
Use `GROUP BY` and column aliasing
```
val counts = spark.sql("SELECT book, COUNT(*) as count FROM kjv_bible GROUP BY book")
counts.show(100) // print the 1st 100 lines, but there are only 66 books/records...
```
**Exercise**: Update the previous query to sort output by the book names, by the counts. For convenience, I've pasted in the same query (different variable name). How much overhead does this add?
```
val sorted_counts = spark.sql("SELECT book, COUNT(*) as count FROM kjv_bible GROUP BY book")
sorted_counts.show(100) // print the 1st 100 lines, but there are only 66 books/records...
```
Use `coalesce` when you have too many partitions, e.g., a small data set and the default number of partitions (200) is too large.
```
val counts1 = counts.coalesce(1)
val nPartitions = counts.rdd.partitions.size
val nPartitions1 = counts1.rdd.partitions.size
println(s"counts.count (can take a while, # partitions = $nPartitions):")
println(s"result: ${counts.count}")
println(s"counts1.count (usually faster, # partitions = $nPartitions1):")
println(s"result: ${counts1.count}")
```
Now do `GROUP BY` with the DataFrame API.
```
val countsDF = verses.groupBy("book").count()
countsDF.show(20)
countsDF.count
```
**Exercise**: Add sorting by book names and by the counts, using the API. For convenience, I've pasted in the same query (different variable name).
```
val countsDF = verses.groupBy("book").count()
countsDF.show(20)
countsDF.count
```
Aggregations, like in Data Warehousing. We need to import some functions first:
```
import org.apache.spark.sql.functions._ // for min, max, etc.
verses.groupBy("book").agg(
max(verses("chapter")),
max(verses("verse")),
count(verses("*"))
).sort($"count(1)".desc, $"book").show(100)
```
Alternative way of referencing columns in verses.
```
verses.groupBy("book").agg(
max($"chapter"),
max($"verse"),
count($"*")
).sort($"count(1)".desc, $"book").show(100)
```
With just a single column, cube and rollup make less sense, but in a bigger dataset, you could do cubes and rollups, too.
```
verses.cube("book").agg(
max($"chapter"),
max($"verse"),
count($"*")
).sort($"count(1)".desc, $"book").show(100)
verses.rollup("book").agg(
max($"chapter"),
max($"verse"),
count($"*")
).sort($"count(1)".desc, $"book").show(100)
```
Map a field to a method to apply to it, but limited to at most one method per field.
```
verses.rollup("book").agg(Map(
"chapter" -> "max",
"verse" -> "max",
"*" -> "count"
)).sort($"count(1)".desc, $"book").show(100)
```
## Exercises
### Exercise 1: Try joins with the abbreviation data
See the project solution file [SparkSQL8-join-with-abbreviations-script.scala](https://github.com/deanwampler/spark-scala-tutorial/blob/master/src/main/scala/sparktutorial/solns/SparkSQL8-join-with-abbreviations-script.scala).
Here we set up the abbreviations, similar to the RDD Joins examples.
Now load the abbreviations, similar to how we loaded the Bible verses. First we need a case class...
```
case class Abbrev(abbrev: String, name: String)
val abbrevNamesRDD = sc.textFile(abbrevToNames).flatMap { line =>
val ary=line.split("\t")
if (ary.length != 2) {
Console.err.println(s"Unexpected line: $line")
Nil // or use Seq.empty[Abbrev]. It will be eliminated by flattening.
} else {
Seq(Abbrev(ary(0), ary(1)))
}
}
val abbrevNames = spark.createDataFrame(abbrevNamesRDD)
abbrevNames.createOrReplaceTempView("abbrevs_to_names")
```
### Exercise 2: Try other SQL constructs
Both using actual SQL and the API.
| github_jupyter |
# lenstronomy numerics
In this notebook we use the different convolution and super-sampling approximations in lenstronomy. The different approximations are described and demonstrated on the use-case of a single Sersic profile. The example is also been used to compare the computational speed and accuracy of the different approximations. Please note that performance and accuracy depend on the specific use case and an optimal choice is upon the discretion and responsibility of the user.
```
import numpy as np
import os
import time
import astropy.io.fits as pyfits
import matplotlib.pyplot as plt
%matplotlib inline
import lenstronomy.Util.util as util
import lenstronomy.Util.kernel_util as kernel_util
```
## set up the example
We use a single elliptical Sersic profile and a TinyTim PSF as our example.
```
# we define a model consisting of a singe Sersric profile
from lenstronomy.LightModel.light_model import LightModel
light_model_list = ['SERSIC_ELLIPSE']
lightModel = LightModel(light_model_list=light_model_list)
kwargs_light = [{'amp': 100, 'R_sersic': 0.5, 'n_sersic': 2, 'e1': 0, 'e2': 0, 'center_x': 0.03, 'center_y': 0}]
# we define a pixel grid and a higher resolution super sampling factor
supersampling_factor = 5
numPix = 61 # cutout pixel size
deltaPix = 0.05 # pixel size in arcsec (area per pixel = deltaPix**2)
x, y, ra_at_xy_0, dec_at_xy_0, x_at_radec_0, y_at_radec_0, Mpix2coord, Mcoord2pix = util.make_grid_with_coordtransform(
numPix=numPix, deltapix=deltaPix, subgrid_res=1, left_lower=False, inverse=False)
flux = lightModel.surface_brightness(x, y, kwargs_list=kwargs_light)
flux = util.array2image(flux)
flux_max = np.max(flux)
conv_pixels_partial = np.zeros((numPix, numPix), dtype=bool)
conv_pixels_partial[flux >= flux_max/10] = True
print(np.sum(conv_pixels_partial))
# import PSF file
path = os.getcwd()
dirpath, _ = os.path.split(path)
module_path, _ = os.path.split(dirpath)
psf_filename = os.path.join(module_path, 'Data/PSF_TinyTim/psf_example.fits')
kernel = pyfits.getdata(psf_filename)
kernel_size = 91
kernel_super = kernel_util.subgrid_kernel(kernel=kernel, num_iter=10, subgrid_res=supersampling_factor, odd=True)
psf_size_super = kernel_size*supersampling_factor
print(np.shape(kernel_super))
if psf_size_super % 2 == 0:
psf_size_super += 1
kernel_super = kernel_util.cut_psf(psf_data=kernel_super, psf_size=psf_size_super)
print(np.shape(kernel_super))
plt.matshow(np.log(kernel_super), origin='lower')
plt.show()
# make instance of the PixelGrid class
from lenstronomy.Data.pixel_grid import PixelGrid
kwargs_grid = {'nx': numPix, 'ny': numPix, 'transform_pix2angle': Mpix2coord, 'ra_at_xy_0': ra_at_xy_0, 'dec_at_xy_0': dec_at_xy_0}
pixel_grid = PixelGrid(**kwargs_grid)
# make instance of the PSF class
from lenstronomy.Data.psf import PSF
kwargs_psf = {'psf_type': 'PIXEL', 'kernel_point_source': kernel_super, 'point_source_supersampling_factor': supersampling_factor}
psf_class = PSF(**kwargs_psf)
plt.matshow(np.log(psf_class.kernel_point_source), origin='lower')
plt.show()
print(np.shape(psf_class.kernel_point_source))
print(np.shape(psf_class.kernel_point_source_supersampled(supersampling_factor)))
```
## computing convolved image with different numerical settings
Now we define different numerical settings, kwargs_numerics, that are imported into the ImageModel class to perform the image computation.
```
timeit = True # profiling boolean
# high resolution ray-tracing and high resolution convolution, the full calculation
kwargs_numerics_true = {'supersampling_factor': supersampling_factor, # super sampling factor of (partial) high resolution ray-tracing
'compute_mode': 'regular', # 'regular' or 'adaptive'
'supersampling_convolution': True, # bool, if True, performs the supersampled convolution (either on regular or adaptive grid)
'supersampling_kernel_size': None, # size of the higher resolution kernel region (can be smaller than the original kernel). None leads to use the full size
'flux_evaluate_indexes': None, # bool mask, if None, it will evaluate all (sub) pixels
'supersampled_indexes': None, # bool mask of pixels to be computed in supersampled grid (only for adaptive mode)
'compute_indexes': None, # bool mask of pixels to be computed the PSF response (flux being added to). Only used for adaptive mode and can be set =likelihood mask.
'point_source_supersampling_factor': 1, # int, supersampling factor when rendering a point source (not used in this script)
}
# high resolution convolution on a smaller PSF with low resolution convolution on the edges of the PSF and high resolution ray tracing
kwargs_numerics_high_res_narrow = {'supersampling_factor': supersampling_factor,
'compute_mode': 'regular',
'supersampling_convolution': True,
'supersampling_kernel_size': 5,
}
# low resolution convolution based on high resolution ray-tracing grid
kwargs_numerics_low_conv_high_grid = {'supersampling_factor': supersampling_factor,
'compute_mode': 'regular',
'supersampling_convolution': False, # does not matter for supersampling_factor=1
'supersampling_kernel_size': None, # does not matter for supersampling_factor=1
}
# low resolution convolution with a subset of pixels with high resolution ray-tracing
kwargs_numerics_low_conv_high_adaptive = {'supersampling_factor': supersampling_factor,
'compute_mode': 'adaptive',
'supersampling_convolution': False, # does not matter for supersampling_factor=1
'supersampling_kernel_size': None, # does not matter for supersampling_factor=1
'supersampled_indexes': conv_pixels_partial,
}
# low resolution convolution with a subset of pixels with high resolution ray-tracing and high resoluton convolution on smaller kernel size
kwargs_numerics_high_adaptive = {'supersampling_factor': supersampling_factor,
'compute_mode': 'adaptive',
'supersampling_convolution': True, # does not matter for supersampling_factor=1
'supersampling_kernel_size': 5, # does not matter for supersampling_factor=1
'supersampled_indexes': conv_pixels_partial,
}
# low resolution convolution and low resolution ray tracing, the simplest calculation
kwargs_numerics_low_res = {'supersampling_factor': 1,
'compute_mode': 'regular',
'supersampling_convolution': False, # does not matter for supersampling_factor=1
'supersampling_kernel_size': None, # does not matter for supersampling_factor=1
}
from lenstronomy.ImSim.image_model import ImageModel
# without convolution
image_model_true = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_true)
image_unconvolved = image_model_true.image(kwargs_lens_light=kwargs_light, unconvolved=True)
plt.matshow(np.log10(image_unconvolved))
plt.title('log unconvolved')
plt.colorbar()
plt.show()
# True computation
image_true = image_model_true.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_true.image(kwargs_lens_light=kwargs_light)
vmin = np.min(image_true)/20
vmax = np.max(image_true)/20
plt.matshow(np.log10(image_true))
plt.title('log true')
plt.colorbar()
plt.show()
# high_res_narrow
image_model_high_res_narrow = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_high_res_narrow)
image_high_res_narrow = image_model_high_res_narrow.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_high_res_narrow.image(kwargs_lens_light=kwargs_light)
plt.matshow(image_true - image_high_res_narrow, origin='lower', vmin=-vmax, vmax=vmax)
plt.title('true - high res narrow kernle')
plt.colorbar()
plt.show()
# low_conv_high_grid
image_model_low_conv_high_grid = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_low_conv_high_grid)
image_low_conv_high_grid = image_model_low_conv_high_grid.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_low_conv_high_grid.image(kwargs_lens_light=kwargs_light)
plt.matshow(image_true - image_low_conv_high_grid, origin='lower', vmin=-vmax, vmax=vmax)
plt.title('true - low conv high grid')
plt.colorbar()
plt.show()
# low_conv_high_adaptive
image_model_low_conv_high_adaptive = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_low_conv_high_adaptive)
image_low_conv_high_adaptive = image_model_low_conv_high_adaptive.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_low_conv_high_adaptive.image(kwargs_lens_light=kwargs_light)
plt.matshow(image_true - image_low_conv_high_adaptive, origin='lower', vmin=-vmax, vmax=vmax)
plt.title('true - low conv high adaptive')
plt.colorbar()
plt.show()
# high_adaptive
image_model_high_adaptive = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_high_adaptive)
image_high_adaptive = image_model_high_adaptive.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_high_adaptive.image(kwargs_lens_light=kwargs_light)
plt.matshow(image_true - image_high_adaptive, origin='lower', vmin=-vmax, vmax=vmax)
plt.title('true - high adaptive')
plt.colorbar()
plt.show()
# low_res
image_model_low_res = ImageModel(pixel_grid, psf_class, lens_light_model_class=lightModel, kwargs_numerics=kwargs_numerics_low_res)
image_low_res = image_model_low_res.image(kwargs_lens_light=kwargs_light)
if timeit is True:
%timeit image_model_low_res.image(kwargs_lens_light=kwargs_light)
plt.matshow(image_true - image_low_res, origin='lower', vmin=-vmax, vmax=vmax)
plt.title('true - low res')
plt.colorbar()
plt.show()
```
| github_jupyter |
# Exploring MIMIC-IV using Colaboratory and BigQuery
- BigQuery needs to be enabled in CoLaboratory. I followed the instructions [here](https://tech.aaronteoh.com/bigquery-colaboratory-basics/) after creating a Google Cloud project that I named `mimic4-bq`. You will need to modify the code to use the project ID you created.
- It took me a while to get this right and I didn't take good notes, so if anyone else wants to share what they had to do to get BigQuery enabled please share.
# Using `ibis` to connect to MIMIC IV on Google BigQuery
Environments in Google Colaboratory are not persistent. If we use any software that is not part of teh Google Python Colaboratory environment, we must install it during each session.
We are going to be using Ibis, so this must be installed.
```
!pip install ibis-framework[bigquery]
```
### Google has a really nice Pandas DataFrame display that we will enable.
```
%load_ext google.colab.data_table
import ibis
import os
project_id="mimic4-bq"
os.environ["GOOGLE_CLOUD_PROJECT"]=project_id
import seaborn as sns
import pandas as pd
import ipywidgets as ipw
from IPython.display import display, HTML, clear_output
import matplotlib.pyplot as plt
from ipywidgets.widgets.interaction import show_inline_matplotlib_plots
```
The Google display helps with having lots of rows, but not with having lots of columns. This class is a rough attempt to be able to scroll through columns. I've also added on a simple visualization. This needs more work, so be patient with unreadable labels, etc.
```
class PandasBrowser(ipw.VBox):
def __init__(self, df, fixed=None, *args, **kwargs):
self.df = df
if fixed == None:
self.fixed = [self.df.columns[0]]
else:
self.fixed = fixed[:]
self.cols = [c for c in self.df.columns if c not in self.fixed]
self.ncols = len(self.cols)
self.ndisp = max(12-len(self.fixed), 10)
if self.ncols < self.ndisp:
col_max = 0
else:
col_max = self.ncols-self.ndisp
self.start_col = ipw.IntSlider(min=0, max=col_max, value=0, description="Start col")
self.start_col.observe(self.disp_df, "value")
self.out = ipw.Output()
children = kwargs.get("children", [])
self.sub = None
self.graph_type = ipw.Dropdown(options=[None, "describe", "categorical", "numeric"], value=None, description="Plot Type")
self.kind = ipw.Dropdown(options=["count", "swarm", "box", "boxen", "violin", "bar", "point"], value="count")
opts = [None]+list(self.df.columns)
self.xsel = ipw.Dropdown(options=opts, value=opts[1], description="x")
self.ysel = ipw.Dropdown(options=opts, value=None, description="y")
self.hsel = ipw.Dropdown(options=opts, value=None, description="hue")
self.rsel = ipw.Dropdown(options=opts, value=None, description="row var")
self.csel = ipw.Dropdown(options=opts, value=None, description="col var")
self.graph_type.observe(self.disp_plot, "value")
self.kind.observe(self.disp_plot, "value")
self.xsel.observe(self.disp_plot, "value")
self.ysel.observe(self.disp_plot, "value")
self.hsel.observe(self.disp_plot, "value")
self.rsel.observe(self.disp_plot, "value")
self.csel.observe(self.disp_plot, "value")
self.plot_out = ipw.Output()
tmp = ipw.HBox([self.graph_type, self.kind, ipw.VBox([self.xsel, self.ysel]), ipw.VBox([self.hsel, self.rsel, self.csel])])
children= [self.start_col, self.out, tmp, self.plot_out] + children
super(PandasBrowser, self).__init__(children=children)
self.disp_df()
self.disp_plot()
def disp_df(self, *args):
cols = self.fixed + self.cols[self.start_col.value:self.start_col.value+self.ndisp]
#self.sub = self.df.loc[:, cols]
self.out.clear_output()
with self.out:
display(self.df.loc[:, cols])
def disp_plot(self, *args):
self.plot_out.clear_output()
if self.graph_type.value == None:
return
with self.plot_out:
if self.graph_type.value == "describe":
display(self.df.loc[:, cols].describe())
else:
if self.graph_type.value == 'categorical':
g = sns.catplot(data=self.df, kind=self.kind.value,
x=self.xsel.value)
#y=self.ysel.value, row=self.rsel.value, col=self.csel.value)
else:
g = sns.pairplot(data=self.df, hue=self.hsel.value)
g.set_xticklabels(rotation=45)
show_inline_matplotlib_plots()
def disp(self, *args):
self.disp_df(args)
self.disp_plot(args)
```
### Authenticate using `google.colab`
```
from google.colab import auth
auth.authenticate_user()
print('Authenticated')
conn = ibis.bigquery.connect(
project_id="mimic4-bq",
dataset_id='physionet-data.mimic_core')
```
### Once we connect we can list all the databases we have access to
```
dbs = conn.list_databases()
print(dbs)
```
### Since I connected to `mimic_core`, I can list the tables in this database
```
conn.list_tables()
```
https://cloud.google.com/community/tutorials/bigquery-ibis
```
patients = conn.table("patients")
```
### The `schema` method will tell you the data types of each column in the table
```
patients.schema()
```
### And do queries
```
pts = patients.execute(limit=2000)
pv = PandasBrowser(pts)
pv
adm = conn.table("admissions").execute(limit=20000)
PandasBrowser(adm)
```
| github_jupyter |
# Table Extraction Case Study
### AAIC Self Case Study II
### Author: Soumya De
<hr>
## 7. Final Pipeline Method
We will now make use of the compressed tflite model that we have obtained in the previous section and design our final pipeline, that is from getting an image (as a file on the disk) to obtain a csv (also as file on the disk)
```
# importing dependencies
import os
import re
import numpy as np
import csv
import pandas as pd
from tqdm import tqdm
from PIL import Image
import matplotlib.pyplot as plt
import cv2
from time import strftime
import pytesseract
import tensorflow as tf
from tensorflow.keras import Model
# defining helper functions
def load_interpreter(model_path=None):
"""
This function loads a tflite model interpreter
"""
if model_path is None:
model_path = os.path.sep.join(['final_model', 'tablenet_densenet121_lite.tflite'])
interpreter = tf.lite.Interpreter(model_path=model_path)
interpreter.allocate_tensors()
return interpreter
def adjust(new_rows, maxi):
"""
A function to set all with maxi number of columns
for making csv compatible
"""
rows = []
for each_row in new_rows:
if len(each_row) < maxi:
for i in range(maxi - len(each_row)):
each_row.append("-")
rows.append(each_row)
return rows
def text2csv(text):
"""
This funtion transorms a text with newline and spaces to
a csv that treats the spaces in the text as comma and newlines as carriage return
"""
rows = text.split('\n')
new_rows = []
maxi = 0
for each_row in rows:
temp_row = each_row.split()
if maxi < len(temp_row):
maxi = len(temp_row)
new_rows.append(temp_row)
new_rows = adjust(new_rows, maxi)
header = ['column_{}'.format(i) for i in range(maxi)]
tstr = strftime("%Y%m%d-%H%M")
temp_dir = os.path.join('output', 'temporary_files')
if not os.path.exists(temp_dir):
os.makedirs(temp_dir)
temp_file = os.path.join(temp_dir, 'temp_{}.csv'.format(tstr))
with open(temp_file, 'w') as f:
csvwriter = csv.writer(f)
csvwriter.writerow(header)
csvwriter.writerows(new_rows)
return temp_file
def append_offset(name, offset):
"""
This function is used for assigning a name with offset if a file with the same name exists
It takes a filename and a offset and returns a valid equivalent name with offset number
Example :
# assume two variables
name = 'python.py'
offset = '2'
append_offset(name, offset)
# The above invocation will return string as
# 'python_2.py'
"""
fname, extension = name.split('.')
fname = ''.join([fname, '_', offset, '.', extension])
return fname
def render(mask):
mask = tf.argmax(mask, axis=-1)
mask = mask[..., tf.newaxis]
return mask[0]
def visualize(image):
plt.figure(figsize=(15, 15))
title = 'Cropped Table'
plt.title(title)
plt.imshow(tf.keras.preprocessing.image.array_to_img(image))
plt.axis('off')
plt.show()
def final(img_path, output_dir='temp_output', show_table=False):
interpreter = load_interpreter()
image_orig = Image.open(img_path)
original_dim = image_orig.size
image = image_orig.resize((512,512))
np_image = np.asarray(image)/255.0
np_image = np_image.astype(np.float32)
np_image = np.expand_dims(np_image, axis=0)
ip_d = interpreter.get_input_details()[0]
op_d = interpreter.get_output_details()[0]
interpreter.set_tensor(ip_d['index'], np_image)
interpreter.invoke()
tab_mask = interpreter.get_tensor(op_d['index'])
tab_mask = np.squeeze(render(tab_mask).numpy())
tab_mask = Image.fromarray(np.uint8(tab_mask))
tab_mask = tab_mask.resize(original_dim)
tab_mask = np.array(tab_mask)
image_orig = image_orig
x, y, w, h = cv2.boundingRect(tab_mask)
tab = image_orig.crop((x, y, x+w, y+h))
text = pytesseract.image_to_string(tab)
text = text.strip()
text = re.sub("[\r\n]+", "\r\n", text)
csv = text2csv(text)
csv_fname = img_path.split(os.path.sep)[-1].replace('png', 'csv')
dest_dir = os.path.join(output_dir)
if not os.path.exists(dest_dir):
os.makedirs(dest_dir)
dest = os.path.join(dest_dir, csv_fname)
# if file already exists in the temp directory it will save the csv
# by appending some offset to the filename before extension
try:
os.rename(csv, dest)
except:
f_save = 'fail'
i=2
while(f_save=='fail'):
name_off = str(i)
try:
dest = os.path.join(dest_dir, append_offset(csv_fname,name_off))
os.rename(csv, dest)
f_save = 'pass'
except:
i += 1
if show_table:
visualize(tab)
return dest
```
### Inference using the final function
```
img_path = os.path.join('data', 'ICDAR 2017', 'table_images', 'POD_0011.png')
csv_path = final(img_path, show_table=True)
df = pd.read_csv(csv_path)
df
```
| github_jupyter |
```
import DeepGenerator.DeepGenerator as dg
# The library is a generative network built purely in numpy and matplotlib. The library is a sequence to sequence transduction
# library for generating n sequence texts from given corpus. Metrics for accuracy-BLEU has provided a considerable accuracy metric
# for epochs greater than 20000.The library is sequential and includes intermediate tanh activation in the intermediate stages with
# softmax cross entropy loss ,and generalised Adagrad optimizer.
# library facts:
# initialisation:
# import DeepGenerator as dg
# ====================
# creating object:
# deepgen=dg.DeepGenerator()
# Functions:
# 1.attributes for users- learning rate,epochs,local path of data storage(text format),number of hidden layers,kernel size,sequence/step size,count of next words
# 2.data_abstract function- Takes arguements (self,path,choice) -
# path= local path of text file
# choice= 'character_generator' for character generation network
# 'word_generator' for word generator network
# Returns data
# Usage- ouput_data=deepgen.data_preprocess(DeepGenerator.path,DeepGenerator.choice)
# 3.data_preprocess function- Takes arguements (self,path,choice)-
# path= local path of text file
# choice= 'character_generator' for character generation network
# 'word_generator' for word generator network
# Returns data,data_size,vocab_size,char_to_idx,idx_to_char
# Usage- data,data_size,vocab_size,char_to_idx,idx_to_char=deepgen.data_preprocess(DeepGenerator.path,DeepGenerator.choice)
# 4.hyperparameters function-Takes arguements (self,hidden_layers_size,no_hidden_layers,learning_rate,step_size,vocab_size)-
# hidden_layers-kernel size-recommended under 2048
# no_hidden_layers- sequential intermediate layers
# learning_rate- learning_rate (range of 1e-3)
# step_size- sequence length(should be <= vocab_size)
# vocab_size
# Returns hidden_layers,learning_rate,step_size,hid_layer,Wxh,Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by
# Usage- hidden_layers,learning_rate,step_size,hid_layer,Wxh,Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by=deepgen.hyperparamteres(dg.hidden_layers_size,dg.no_hidden_layers,dg.learning_rate,dg.step_size,dg.vocab_size)
# 5. loss_evaluation function- Takes arguements (self,inp,target,h_previous,hidden_layers,hid_layer,Wxh,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by) -
# inp= character to indices encoded dictionary of input text
# target=character to indices encoded dictionary of generated text
# h_previous-value of hidden layer for previous state
# hidden_layers-kernel size
# hid_layer-sequential hidden layers
# ---------- sequential layers---------
# -----weight tensors------
# Wxh- weight tensor of input to first hidden layer
# Wh1- weight tensor of first hidden layer to first layer of sequential network
# Whh_vector-weight tensors of intermediate sequential network
# Whh- weight tensor of last sequential to last hidden layer
# Why-weight tensor of last hidden layer to output layer
# -----bias tensors-------
# bh1-bias of first hidden layer
# bh_vector-bias of intermediate sequential layers
# bhh-bias of end hidden layer
# by-bias at output
# Returns loss,dWxh,dWhh1,dWhh_vector,dWhh,dWhy,dbh1,dbh_vector,dbh,dby,h_state[len(inp)-1],Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by
# Usage loss,dWxh,dWhh1,dWhh_vector,dWhh,dWhy,dbh1,dbh_vector,dbh,dby,h_state[len(inp)-1],Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by=deepgen.loss_evaluation(dg.inp,dg.target,dg.h_previous,dg.hidden_layers,dg.hid_layer,dg.Wxh,dg.Wh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by)
# 6.start_predict function-Takes arguements (self,count,epochs,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by,hid_layer,char_to_idx,idx_to_char,vocab_size,learning_rate,step_size,data,hidden_layers)
# counts-count of sequences to generate
# epochs-epochs
# Whi -weight tensors
# bhi-bias tensors
# hid_layer-no of sequential layers
# char_to_idx-character to index encoder
# idx_to_char-index to character decoder
# vocab_size-vocab_size
# learning_rate-learning_rate
# step_size-sequence length
# hidden_layers-kernel size
# Returns epochs and gradient losses vector
# Usage-epochs,gradient_loss=deepgen.start_predict(dg.count,dg.epochs,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer,dg.char_to_idx,dg.idx_to_char,dg.vocab_size,dg.learning_rate,dg.step_size,dg.data,dg.hidden_layers)
# 7.output_sample function- Takes arguements (self,h1,seed_ix,n,vocab_size,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by,hid_layer)-
# h1-hidden layer previous state
# seed_ix-starting point for generation
# n-count of text to generate
# Whi-weight tensor
# bhi-bias tensor
# hid_layer-no of sequential layers
# Returns ixs- integer vector of maximum probability characters/words
# Usage-ixs=deepgen.output_sample(dg.h1,dg.seed_ix,dg.n,dg.vocab_size,dg.Wh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer)
# 8.plot_loss function -Takes arguements(self,epochs,gradient_loss)-
# epochs-epoch vector
# gradient_loss- gradient loss vector
# Returns void
# Usage-deepgen.plot_loss(dg.epoch,dg.gradient_loss)
#instruction for generation
if __name__=='__main__':
#create instance object
deepgen=dg.DeepGenerator()
#specify hyperparamters
dg.learning_rate=1e-1
dg.step_size=25
dg.no_hidden_layers=24
dg.hidden_layers_size=64
dg.path='C:\\Users\\User\\Desktop\\test2.txt'
dg.choice='word_generator'
dg.epochs=1500
dg.count=100
print("Sequential model for French-De l'embarquement de monseigneur l'archiduc don Fernande, pour venir en Flandre.")
#sequencing model for French#
#data_preprocess
dg.data,dg.data_size,dg.vocab_size,dg.char_to_idx,dg.idx_to_char=deepgen.data_preprocess(dg.path,dg.choice)
#hyperparamters
dg.hidden_layers,dg.learning_rate,dg.step_size,dg.hid_layer,dg.Wxh,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by=deepgen.hyperparamteres(dg.hidden_layers_size,dg.no_hidden_layers,dg.learning_rate,dg.step_size,dg.vocab_size)
#generate text
dg.epoch,dg.gradient_loss=deepgen.start_predict(dg.count,dg.epochs,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer,dg.char_to_idx,dg.idx_to_char,dg.vocab_size,dg.learning_rate,dg.step_size,dg.data,dg.hidden_layers)
#display loss wrt epoch
deepgen.plot_loss(dg.epoch,dg.gradient_loss)
import DeepGenerator.DeepGenerator as dg
# The library is a generative network built purely in numpy and matplotlib. The library is a sequence to sequence transduction
# library for generating n sequence texts from given corpus. Metrics for accuracy-BLEU has provided a considerable accuracy metric
# for epochs greater than 20000.The library is sequential and includes intermediate tanh activation in the intermediate stages with
# softmax cross entropy loss ,and generalised Adagrad optimizer.
# library facts:
# initialisation:
# import DeepGenerator as dg
# ====================
# creating object:
# deepgen=dg.DeepGenerator()
# Functions:
# 1.attributes for users- learning rate,epochs,local path of data storage(text format),number of hidden layers,kernel size,sequence/step size,count of next words
# 2.data_abstract function- Takes arguements (self,path,choice) -
# path= local path of text file
# choice= 'character_generator' for character generation network
# 'word_generator' for word generator network
# Returns data
# Usage- ouput_data=deepgen.data_preprocess(DeepGenerator.path,DeepGenerator.choice)
# 3.data_preprocess function- Takes arguements (self,path,choice)-
# path= local path of text file
# choice= 'character_generator' for character generation network
# 'word_generator' for word generator network
# Returns data,data_size,vocab_size,char_to_idx,idx_to_char
# Usage- data,data_size,vocab_size,char_to_idx,idx_to_char=deepgen.data_preprocess(DeepGenerator.path,DeepGenerator.choice)
# 4.hyperparameters function-Takes arguements (self,hidden_layers_size,no_hidden_layers,learning_rate,step_size,vocab_size)-
# hidden_layers-kernel size-recommended under 2048
# no_hidden_layers- sequential intermediate layers
# learning_rate- learning_rate (range of 1e-3)
# step_size- sequence length(should be <= vocab_size)
# vocab_size
# Returns hidden_layers,learning_rate,step_size,hid_layer,Wxh,Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by
# Usage- hidden_layers,learning_rate,step_size,hid_layer,Wxh,Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by=deepgen.hyperparamteres(dg.hidden_layers_size,dg.no_hidden_layers,dg.learning_rate,dg.step_size,dg.vocab_size)
# 5. loss_evaluation function- Takes arguements (self,inp,target,h_previous,hidden_layers,hid_layer,Wxh,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by) -
# inp= character to indices encoded dictionary of input text
# target=character to indices encoded dictionary of generated text
# h_previous-value of hidden layer for previous state
# hidden_layers-kernel size
# hid_layer-sequential hidden layers
# ---------- sequential layers---------
# -----weight tensors------
# Wxh- weight tensor of input to first hidden layer
# Wh1- weight tensor of first hidden layer to first layer of sequential network
# Whh_vector-weight tensors of intermediate sequential network
# Whh- weight tensor of last sequential to last hidden layer
# Why-weight tensor of last hidden layer to output layer
# -----bias tensors-------
# bh1-bias of first hidden layer
# bh_vector-bias of intermediate sequential layers
# bhh-bias of end hidden layer
# by-bias at output
# Returns loss,dWxh,dWhh1,dWhh_vector,dWhh,dWhy,dbh1,dbh_vector,dbh,dby,h_state[len(inp)-1],Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by
# Usage loss,dWxh,dWhh1,dWhh_vector,dWhh,dWhy,dbh1,dbh_vector,dbh,dby,h_state[len(inp)-1],Whh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by=deepgen.loss_evaluation(dg.inp,dg.target,dg.h_previous,dg.hidden_layers,dg.hid_layer,dg.Wxh,dg.Wh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by)
# 6.start_predict function-Takes arguements (self,count,epochs,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by,hid_layer,char_to_idx,idx_to_char,vocab_size,learning_rate,step_size,data,hidden_layers)
# counts-count of sequences to generate
# epochs-epochs
# Whi -weight tensors
# bhi-bias tensors
# hid_layer-no of sequential layers
# char_to_idx-character to index encoder
# idx_to_char-index to character decoder
# vocab_size-vocab_size
# learning_rate-learning_rate
# step_size-sequence length
# hidden_layers-kernel size
# Returns epochs and gradient losses vector
# Usage-epochs,gradient_loss=deepgen.start_predict(dg.count,dg.epochs,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer,dg.char_to_idx,dg.idx_to_char,dg.vocab_size,dg.learning_rate,dg.step_size,dg.data,dg.hidden_layers)
# 7.output_sample function- Takes arguements (self,h1,seed_ix,n,vocab_size,Wh1,Whh_vector,Whh,Why,bh1,bh_vector,bh,by,hid_layer)-
# h1-hidden layer previous state
# seed_ix-starting point for generation
# n-count of text to generate
# Whi-weight tensor
# bhi-bias tensor
# hid_layer-no of sequential layers
# Returns ixs- integer vector of maximum probability characters/words
# Usage-ixs=deepgen.output_sample(dg.h1,dg.seed_ix,dg.n,dg.vocab_size,dg.Wh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer)
# 8.plot_loss function -Takes arguements(self,epochs,gradient_loss)-
# epochs-epoch vector
# gradient_loss- gradient loss vector
# Returns void
# Usage-deepgen.plot_loss(dg.epoch,dg.gradient_loss)
#instruction for generation
if __name__=='__main__':
#create instance object
deepgen=dg.DeepGenerator()
#specify hyperparamters
dg.learning_rate=1e-1
dg.step_size=25
dg.no_hidden_layers=24
dg.hidden_layers_size=64
dg.path='C:\\Users\\User\\Desktop\\test1.txt'
dg.choice='word_generator'
dg.epochs=1500
dg.count=100
print("Sequence model for English poem - The road not taken- Robert Frost")
#sequencing model for English - The road not taken- Robert Frost#
#data_preprocess
dg.data,dg.data_size,dg.vocab_size,dg.char_to_idx,dg.idx_to_char=deepgen.data_preprocess(dg.path,dg.choice)
#hyperparamters
dg.hidden_layers,dg.learning_rate,dg.step_size,dg.hid_layer,dg.Wxh,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by=deepgen.hyperparamteres(dg.hidden_layers_size,dg.no_hidden_layers,dg.learning_rate,dg.step_size,dg.vocab_size)
#generate text
dg.epoch,dg.gradient_loss=deepgen.start_predict(dg.count,dg.epochs,dg.Whh1,dg.Whh_vector,dg.Whh,dg.Why,dg.bh1,dg.bh_vector,dg.bh,dg.by,dg.hid_layer,dg.char_to_idx,dg.idx_to_char,dg.vocab_size,dg.learning_rate,dg.step_size,dg.data,dg.hidden_layers)
#display loss wrt epoch
deepgen.plot_loss(dg.epoch,dg.gradient_loss)
```
| github_jupyter |
# Maximizing the profit of an oil company
This tutorial includes everything you need to set up the decision optimization engines and build mathematical programming models.
Table of contents:
- [Describe the business problem](#Describe-the-business-problem)
* [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
* [Use decision optimization](#Use-decision-optimization)
- [Step 1: Model the data](#Step-1:-Model-the-data)
* [Step 2: Prepare the data](#Step-2:-Prepare-the-data)
- [Step 3: Set up the prescriptive model](#Step-3:-Set-up-the-prescriptive-model)
* [Define the decision variables](#Define-the-decision-variables)
* [Express the business constraints](#Express-the-business-constraints)
* [Express the objective](#Express-the-objective)
* [Solve with Decision Optimization](#Solve-with-Decision-Optimization)
* [Step 4: Investigate the solution and run an example analysis](#Step-4:-Investigate-the-solution-and-then-run-an-example-analysis)
* [Summary](#Summary)
## Describe the business problem
* An oil company manufactures different types of gasoline and diesel. Each type of gasoline is produced by blending different types of crude oils that must be purchased. The company must decide how much crude oil to buy in order to maximize its profit while respecting processing capacities and quality levels as well as satisfying customer demand.
* Blending problems are a typical industry application of Linear Programming (LP). LP represents real life problems mathematically using an objective function to represent the goal that is to be minimized or maximized, together with a set of linear constraints which define the conditions to be satisfied and the limitations of the real life problem. The function and constraints are expressed in terms of decision variables and the solution, obtained from optimization engines such as IBM® ILOG® CPLEX®, provides the best values for these variables so that the objective function is optimized.
* The oil-blending problem consists of calculating different blends of gasoline according to specific quality criteria.
* Three types of gasoline are manufactured: super, regular, and diesel.
* Each type of gasoline is produced by blending three types of crude oil: crude1, crude2, and crude3.
* The gasoline must satisfy some quality criteria with respect to their lead content and their octane ratings, thus constraining the possible blendings.
* The company must also satisfy its customer demand, which is 3,000 barrels a day of super, 2,000 of regular, and 1,000 of diesel.
* The company can purchase 5,000 barrels of each type of crude oil per day and can process at most 14,000 barrels a day.
* In addition, the company has the option of advertising a gasoline, in which case the demand for this type of gasoline increases by ten barrels for every dollar spent.
* Finally, it costs four dollars to transform a barrel of oil into a barrel of gasoline.
## How decision optimization can help
* Prescriptive analytics technology recommends actions based on desired outcomes, taking into account specific scenarios, resources, and knowledge of past and current events. This insight can help your organization make better decisions and have greater control of business outcomes.
* Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
* Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
<br/>
+ For example:
+ Automate complex decisions and trade-offs to better manage limited resources.
+ Take advantage of a future opportunity or mitigate a future risk.
+ Proactively update recommendations based on changing events.
+ Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
## Use decision optimization
### Step 1: Model the data
* For each type of crude oil, there are capacities of what can be bought, the buying price, the octane level, and the lead level.
* For each type of gasoline or diesel, there is customer demand, selling prices, and octane and lead levels.
* There is a maximum level of production imposed by the factory's limit as well as a fixed production cost.
* There are inventory costs for each type of final product and blending proportions. All of these have actual values in the model.
* The marginal production cost and maximum production are assumed to be identical for all oil types.
Input data comes as *NumPy* arrays with two dimensions. <a href="http://www.numpy.org/" target="_blank" rel="noopener noreferrer">NumPy</a> is the fundamental package for scientific computing with Python.
The first dimension of the *NumPy* array is the number of gasoline types;
and for each gasoline type, you have a *NumPy* array containing capacity, price, octane and lead level, in that order.
```
import numpy as np
gas_names = ["super", "regular", "diesel"]
gas_data = np.array([[3000, 70, 10, 1], [2000, 60, 8, 2], [1000, 50, 6, 1]])
oil_names = ["crude1", "crude2", "crude3"]
oil_data = np.array([[5000, 45, 12, 0.5], [5000, 35, 6, 2], [5000, 25, 8, 3]])
nb_gas = len(gas_names)
nb_oils = len(oil_names)
range_gas = range(nb_gas)
range_oil = range(nb_oils)
print("Number of gasoline types = {0}".format(nb_gas))
print("Number of crude types = {0}".format(nb_oils))
# global data
production_cost = 4
production_max = 14000
# each $1 spent on advertising increases demand by 10.
advert_return = 10
```
### Step 2: Prepare the data
<a href="http://pandas.pydata.org/" target="_blank" rel="noopener noreferrer">Pandas</a> is another Python library that is used to store data. *pandas* contains data structures and data analysis tools for the Python programming language.
```
import pandas as pd
gaspd = pd.DataFrame([(gas_names[i],int(gas_data[i][0]),int(gas_data[i][1]),int(gas_data[i][2]),int(gas_data[i][3]))
for i in range_gas])
oilpd = pd.DataFrame([(oil_names[i],int(oil_data[i][0]),int(oil_data[i][1]),int(oil_data[i][2]),oil_data[i][3])
for i in range_oil])
gaspd.columns = ['name','demand','price','octane','lead']
oilpd.columns= ['name','capacity','price','octane','lead']
```
Use basic HTML and a stylesheet to format the data.
```
CSS = """
body {
margin: 0;
font-family: Helvetica;
}
table.dataframe {
border-collapse: collapse;
border: none;
}
table.dataframe tr {
border: none;
}
table.dataframe td, table.dataframe th {
margin: 0;
border: 1px solid white;
padding-left: 0.25em;
padding-right: 0.25em;
}
table.dataframe th:not(:empty) {
background-color: #fec;
text-align: left;
font-weight: normal;
}
table.dataframe tr:nth-child(2) th:empty {
border-left: none;
border-right: 1px dashed #888;
}
table.dataframe td {
border: 2px solid #ccf;
background-color: #f4f4ff;
}
table.dataframe thead th:first-child {
display: none;
}
table.dataframe tbody th {
display: none;
}
"""
from IPython.core.display import HTML
HTML('<style>{}</style>'.format(CSS))
```
Now display the data that you have just prepared.
```
from IPython.display import display
print("Gas data:")
display(gaspd)
print("Oil data:")
display(oilpd)
```
### Step 4: Set up the prescriptive model
#### Create the DOcplex model
A model is needed to store all the variables and constraints needed to formulate the business problem and submit the problem to the solve service. Use the DOcplex Mathematical Programming (docplex.mp) modeling package.
```
from docplex.mp.model import Model
mdl = Model(name="oil_blending")
```
#### Define the decision variables
For each combination of oil and gas, you must decide the quantity of oil to use to produce a gasoline. A decision variable is needed to represent that amount.
A matrix of continuous variables, indexed by the set of oils and the set of gasolines is thus created.
```
blends = mdl.continuous_var_matrix(keys1=nb_oils, keys2=nb_gas, lb=0)
```
You also need to decide how much should be spent in advertising for each type of gasoline. To do so, you will create a list of continuous variables, indexed by the gasolines.
```
adverts = mdl.continuous_var_list(nb_gas, lb=0)
```
#### Express the business constraints
The business constraints are the following:
* The demand for each gasoline type must be satisfied. The total demand includes the initial demand as stored in the data, plus a variable demand caused by the advertising. This increase is assumed to be proportional to the advertising cost.
- The capacity constraint on each oil type must also be satisfied.
- For each gasoline type, the octane level must be above a minimum level, and the lead level must be below a maximum level.
##### Demand
+ For each gasoline type, the total quantity produced must equal the raw demand plus the demand increase created by the advertising.
```
# gasoline demand is numpy array field #0
mdl.add_constraints(mdl.sum(blends[o, g] for o in range(nb_oils)) == gas_data[g][0] + advert_return * adverts[g]
for g in range(nb_gas))
mdl.print_information()
```
##### Maximum capacity
+ For each type of oil, the total quantity used in all types of gasolines must not exceed the maximum capacity for this oil.
```
mdl.add_constraints(mdl.sum(blends[o,g] for g in range_gas) <= oil_data[o][0]
for o in range_oil)
mdl.print_information()
```
##### Octane and Lead levels
+ For each gasoline type, the octane level must be above a minimum level, and the lead level must be below a maximum level.
```
# minimum octane level
# octane is numpy array field #2
mdl.add_constraints(mdl.sum(blends[o,g]*(oil_data[o][2] - gas_data[g][2]) for o in range_oil) >= 0
for g in range_gas)
# maximum lead level
# lead level is numpy array field #3
mdl.add_constraints(mdl.sum(blends[o,g]*(oil_data[o][3] - gas_data[g][3]) for o in range_oil) <= 0
for g in range_gas)
mdl.print_information()
```
##### Maximum total production
+ The total production must not exceed the maximum (here 14000).
```
# -- maximum global production
mdl.add_constraint(mdl.sum(blends) <= production_max)
mdl.print_information()
```
#### Express the objective
* The objective or goal is to maximize profit, which is made from sales of the final products minus total costs. The costs consist of the purchase cost of the crude oils, production costs, and inventory costs.
- The model maximizes the net revenue, that is the revenue minus oil cost and production cost and advertising cost.
- To define business objective, you can define a few KPIs :
* Total advertising cost
- Total Oil cost
- Total production cost
- Total revenue
```
# KPIs
total_advert_cost = mdl.sum(adverts)
mdl.add_kpi(total_advert_cost, "Total advertising cost")
total_oil_cost = mdl.sum(blends[o,g] * oil_data[o][1] for o in range_oil for g in range_gas)
mdl.add_kpi(total_oil_cost, "Total Oil cost")
total_production_cost = production_cost * mdl.sum(blends)
mdl.add_kpi(total_production_cost, "Total production cost")
total_revenue = mdl.sum(blends[o,g] * gas_data[g][1] for g in range(nb_gas) for o in range(nb_oils))
mdl.add_kpi(total_revenue, "Total revenue")
# finally the objective
mdl.maximize(total_revenue - total_oil_cost - total_production_cost - total_advert_cost)
```
#### Solve with Decision Optimization
Now display the objective and KPI values after the solve by calling the method report() on the model.
```
assert mdl.solve(), "Solve failed"
mdl.report()
```
### Step 4: Investigate the solution and then run an example analysis
#### Displaying the solution
First, get the KPIs values and store them in a *pandas* DataFrame.
```
all_kpis = [(kp.name, kp.compute()) for kp in mdl.iter_kpis()]
kpis_bd = pd.DataFrame(all_kpis, columns=['kpi', 'value'])
blend_values = [ [ blends[o,g].solution_value for g in range_gas] for o in range_oil]
total_gas_prods = [sum(blend_values[o][g] for o in range_oil) for g in range_gas]
prods = list(zip(gas_names, total_gas_prods))
prods_bd = pd.DataFrame(prods)
```
Let's display some KPIs in pie charts using the Python package [*matplotlib*](http://matplotlib.org/).
```
%matplotlib inline
import matplotlib.pyplot as plt
def display_pie(pie_values, pie_labels, colors=None,title=''):
plt.axis("equal")
plt.pie(pie_values, labels=pie_labels, colors=colors, autopct="%1.1f%%")
plt.title(title)
plt.show()
display_pie( [kpnv[1] for kpnv in all_kpis], [kpnv[0] for kpnv in all_kpis],title='KPIs: Revenue - Oil Cost - Production Cost')
```
##### Production
```
display_pie(total_gas_prods, gas_names, colors=["green", "goldenrod", "lightGreen"],title='Gasoline Total Production')
```
You can see that the most produced gasoline type is by far regular.
Now, plot the breakdown of oil blend quantities per gasoline type.
A multiple bar chart diagram is used, displaying all blend values for each couple of oil and gasoline type.
```
sblends = [(gas_names[n], oil_names[o], round(blends[o,n].solution_value)) for n in range_gas for o in range_oil]
blends_bd = pd.DataFrame(sblends)
f, barplot = plt.subplots(1, figsize=(16,5))
bar_width = 0.1
offset = 0.12
rho = 0.7
# position of left-bar boundaries
bar_l = [o for o in range_oil]
mbar_w = 3*bar_width+2*max(0, offset-bar_width)
tick_pos = [b*rho + mbar_w/2.0 for b in bar_l]
colors = ['olive', 'lightgreen', 'cadetblue']
for i in range_oil:
barplot.bar([b*rho + (i*offset) for b in bar_l],
blend_values[i], width=bar_width, color=colors[i], label=oil_names[i])
plt.xticks(tick_pos, gas_names)
barplot.set_xlabel("gasolines")
barplot.set_ylabel("blend")
plt.legend(loc="upper right")
plt.title('Blend Repartition\n')
# Set a buffer around the edge
plt.xlim([0, max(tick_pos)+mbar_w +0.5])
plt.show()
```
Notice the missing bar for (crude2, diesel) which is expected since blend[crude2, diesel] is zero in the solution.
You can check the solution value of blends for *crude2* and *diesel*, remembering that crude2 has offset 1 and diesel has offset 2.
Note how the decision variable is automatically converted to a float here. This would raise an exception if called before submitting a solve, as no solution value would be present.
```
print("* value of blend[crude2, diesel] is %g" % blends[1,2])
```
## Summary
You have learned how to set up and use IBM Decision Optimization CPLEX Modeling for Python to formulate a Mathematical Programming model and solve it with CPLEX.
## References
* <a href="https://rawgit.com/IBMDecisionOptimization/docplex-doc/master/docs/index.html" target="_blank" rel="noopener noreferrer">Decision Optimization CPLEX Modeling for Python documentation</a>
* <a href="https://dataplatform.cloud.ibm.com/docs/content/wsj/getting-started/welcome-main.html" target="_blank" rel="noopener noreferrer">Watson Studio documentation</a>
<hr>
Copyright © 2017-2021. This notebook and its source code are released under the terms of the MIT License.
<div style="background:#F5F7FA; height:110px; padding: 2em; font-size:14px;">
<span style="font-size:18px;color:#152935;">Love this notebook? </span>
<span style="font-size:15px;color:#152935;float:right;margin-right:40px;">Don't have an account yet?</span><br>
<span style="color:#5A6872;">Share it with your colleagues and help them discover the power of Watson Studio!</span>
<span style="border: 1px solid #3d70b2;padding:8px;float:right;margin-right:40px; color:#3d70b2;"><a href="https://ibm.co/wsnotebooks" target="_blank" style="color: #3d70b2;text-decoration: none;">Sign Up</a></span><br>
</div>
| github_jupyter |
# Convolutional Neural Network in TensorFlow
Credits: Forked from [TensorFlow-Examples](https://github.com/aymericdamien/TensorFlow-Examples) by Aymeric Damien
## Setup
Refer to the [setup instructions](http://nbviewer.ipython.org/github/donnemartin/data-science-ipython-notebooks/blob/master/deep-learning/tensor-flow-examples/Setup_TensorFlow.md)
```
import numpy as np
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
# Parameters
learning_rate = 0.001
training_iters = 100000
batch_size = 128
display_step = 20
# Network Parameters
n_input = 784 # MNIST data input (img shape: 28*28)
n_classes = 10 # MNIST total classes (0-9 digits)
dropout = 0.75 # Dropout, probability to keep units
# tf Graph input
x = tf.placeholder(tf.float32, [None, n_input])
y = tf.placeholder(tf.float32, [None, n_classes])
keep_prob = tf.placeholder(tf.float32) #dropout (keep probability)
# Create model
def conv2d(img, w, b):
return tf.nn.relu(tf.nn.bias_add(tf.nn.conv2d(img, w, strides=[1, 1, 1, 1],
padding='SAME'),b))
def max_pool(img, k):
return tf.nn.max_pool(img, ksize=[1, k, k, 1], strides=[1, k, k, 1], padding='SAME')
def conv_net(_X, _weights, _biases, _dropout):
# Reshape input picture
_X = tf.reshape(_X, shape=[-1, 28, 28, 1])
# Convolution Layer
conv1 = conv2d(_X, _weights['wc1'], _biases['bc1'])
# Max Pooling (down-sampling)
conv1 = max_pool(conv1, k=2)
# Apply Dropout
conv1 = tf.nn.dropout(conv1, _dropout)
# Convolution Layer
conv2 = conv2d(conv1, _weights['wc2'], _biases['bc2'])
# Max Pooling (down-sampling)
conv2 = max_pool(conv2, k=2)
# Apply Dropout
conv2 = tf.nn.dropout(conv2, _dropout)
# Fully connected layer
# Reshape conv2 output to fit dense layer input
dense1 = tf.reshape(conv2, [-1, _weights['wd1'].get_shape().as_list()[0]])
# Relu activation
dense1 = tf.nn.relu(tf.add(tf.matmul(dense1, _weights['wd1']), _biases['bd1']))
# Apply Dropout
dense1 = tf.nn.dropout(dense1, _dropout) # Apply Dropout
# Output, class prediction
out = tf.add(tf.matmul(dense1, _weights['out']), _biases['out'])
return out
# Store layers weight & bias
weights = {
# 5x5 conv, 1 input, 32 outputs
'wc1': tf.Variable(tf.random_normal([5, 5, 1, 32])),
# 5x5 conv, 32 inputs, 64 outputs
'wc2': tf.Variable(tf.random_normal([5, 5, 32, 64])),
# fully connected, 7*7*64 inputs, 1024 outputs
'wd1': tf.Variable(tf.random_normal([7*7*64, 1024])),
# 1024 inputs, 10 outputs (class prediction)
'out': tf.Variable(tf.random_normal([1024, n_classes]))
}
biases = {
'bc1': tf.Variable(tf.random_normal([32])),
'bc2': tf.Variable(tf.random_normal([64])),
'bd1': tf.Variable(tf.random_normal([1024])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
# Construct model
pred = conv_net(x, weights, biases, keep_prob)
# Define loss and optimizer
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(pred, y))
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
# Evaluate model
correct_pred = tf.equal(tf.argmax(pred,1), tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# Initializing the variables
init = tf.global_variables_initializer()
# Launch the graph
with tf.Session() as sess:
sess.run(init)
step = 1
# Keep training until reach max iterations
while step * batch_size < training_iters:
batch_xs, batch_ys = mnist.train.next_batch(batch_size)
# Fit training using batch data
sess.run(optimizer, feed_dict={x: batch_xs, y: batch_ys, keep_prob: dropout})
if step % display_step == 0:
# Calculate batch accuracy
acc = sess.run(accuracy, feed_dict={x: batch_xs, y: batch_ys, keep_prob: 1.})
# Calculate batch loss
loss = sess.run(cost, feed_dict={x: batch_xs, y: batch_ys, keep_prob: 1.})
print ("Iter " + str(step*batch_size) + ", Minibatch Loss= " + \
"{:.6f}".format(loss) + ", Training Accuracy= " + "{:.5f}".format(acc))
step += 1
print ("Optimization Finished!")
# Calculate accuracy for 256 mnist test images
print ("Testing Accuracy:", sess.run(accuracy, feed_dict={x: mnist.test.images[:256],
y: mnist.test.labels[:256],
keep_prob: 1.}))
```
| github_jupyter |
##### Copyright 2018 Google LLC.
Licensed under the Apache License, Version 2.0 (the "License");
Licensed under the Apache License, Version 2.0 (the "License");
you may not use this file except in compliance with the License.
You may obtain a copy of the License at
https://www.apache.org/licenses/LICENSE-2.0
Unless required by applicable law or agreed to in writing, software
distributed under the License is distributed on an "AS IS" BASIS,
WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
See the License for the specific language governing permissions and
limitations under the License.
# Training a Simple Neural Network, with PyTorch Data Loading

Let's combine everything we showed in the [quickstart notebook](https://colab.research.google.com/github/google/jax/blob/master/notebooks/quickstart.ipynb) to train a simple neural network. We will first specify and train a simple MLP on MNIST using JAX for the computation. We will use PyTorch's data loading API to load images and labels (because it's pretty great, and the world doesn't need yet another data loading library).
Of course, you can use JAX with any API that is compatible with NumPy to make specifying the model a bit more plug-and-play. Here, just for explanatory purposes, we won't use any neural network libraries or special APIs for builidng our model.
```
from __future__ import print_function, division, absolute_import
import jax.numpy as np
from jax import grad, jit, vmap
from jax import random
```
### Hyperparameters
Let's get a few bookkeeping items out of the way.
```
# A helper function to randomly initialize weights and biases
# for a dense neural network layer
def random_layer_params(m, n, key, scale=1e-2):
w_key, b_key = random.split(key)
return scale * random.normal(w_key, (n, m)), scale * random.normal(b_key, (n,))
# Initialize all layers for a fully-connected neural network with sizes "sizes"
def init_network_params(sizes, key):
keys = random.split(key, len(sizes))
return [random_layer_params(m, n, k) for m, n, k in zip(sizes[:-1], sizes[1:], keys)]
layer_sizes = [784, 512, 512, 10]
param_scale = 0.1
step_size = 0.0001
num_epochs = 10
batch_size = 128
n_targets = 10
params = init_network_params(layer_sizes, random.PRNGKey(0))
```
### Auto-batching predictions
Let us first define our prediction function. Note that we're defining this for a _single_ image example. We're going to use JAX's `vmap` function to automatically handle mini-batches, with no performance penalty.
```
from jax.scipy.special import logsumexp
def relu(x):
return np.maximum(0, x)
def predict(params, image):
# per-example predictions
activations = image
for w, b in params[:-1]:
outputs = np.dot(w, activations) + b
activations = relu(outputs)
final_w, final_b = params[-1]
logits = np.dot(final_w, activations) + final_b
return logits - logsumexp(logits)
```
Let's check that our prediction function only works on single images.
```
# This works on single examples
random_flattened_image = random.normal(random.PRNGKey(1), (28 * 28,))
preds = predict(params, random_flattened_image)
print(preds.shape)
# Doesn't work with a batch
random_flattened_images = random.normal(random.PRNGKey(1), (10, 28 * 28))
try:
preds = predict(params, random_flattened_images)
except TypeError:
print('Invalid shapes!')
# Let's upgrade it to handle batches using `vmap`
# Make a batched version of the `predict` function
batched_predict = vmap(predict, in_axes=(None, 0))
# `batched_predict` has the same call signature as `predict`
batched_preds = batched_predict(params, random_flattened_images)
print(batched_preds.shape)
```
At this point, we have all the ingredients we need to define our neural network and train it. We've built an auto-batched version of `predict`, which we should be able to use in a loss function. We should be able to use `grad` to take the derivative of the loss with respect to the neural network parameters. Last, we should be able to use `jit` to speed up everything.
### Utility and loss functions
```
def one_hot(x, k, dtype=np.float32):
"""Create a one-hot encoding of x of size k."""
return np.array(x[:, None] == np.arange(k), dtype)
def accuracy(params, images, targets):
target_class = np.argmax(targets, axis=1)
predicted_class = np.argmax(batched_predict(params, images), axis=1)
return np.mean(predicted_class == target_class)
def loss(params, images, targets):
preds = batched_predict(params, images)
return -np.sum(preds * targets)
@jit
def update(params, x, y):
grads = grad(loss)(params, x, y)
return [(w - step_size * dw, b - step_size * db)
for (w, b), (dw, db) in zip(params, grads)]
```
### Data Loading with PyTorch
JAX is laser-focused on program transformations and accelerator-backed NumPy, so we don't include data loading or munging in the JAX library. There are already a lot of great data loaders out there, so let's just use them instead of reinventing anything. We'll grab PyTorch's data loader, and make a tiny shim to make it work with NumPy arrays.
```
!pip install torch torchvision
import numpy as onp
from torch.utils import data
from torchvision.datasets import MNIST
def numpy_collate(batch):
if isinstance(batch[0], onp.ndarray):
return onp.stack(batch)
elif isinstance(batch[0], (tuple,list)):
transposed = zip(*batch)
return [numpy_collate(samples) for samples in transposed]
else:
return onp.array(batch)
class NumpyLoader(data.DataLoader):
def __init__(self, dataset, batch_size=1,
shuffle=False, sampler=None,
batch_sampler=None, num_workers=0,
pin_memory=False, drop_last=False,
timeout=0, worker_init_fn=None):
super(self.__class__, self).__init__(dataset,
batch_size=batch_size,
shuffle=shuffle,
sampler=sampler,
batch_sampler=batch_sampler,
num_workers=num_workers,
collate_fn=numpy_collate,
pin_memory=pin_memory,
drop_last=drop_last,
timeout=timeout,
worker_init_fn=worker_init_fn)
class FlattenAndCast(object):
def __call__(self, pic):
return onp.ravel(onp.array(pic, dtype=np.float32))
# Define our dataset, using torch datasets
mnist_dataset = MNIST('/tmp/mnist/', download=True, transform=FlattenAndCast())
training_generator = NumpyLoader(mnist_dataset, batch_size=128, num_workers=0)
# Get the full train dataset (for checking accuracy while training)
train_images = onp.array(mnist_dataset.train_data).reshape(len(mnist_dataset.train_data), -1)
train_labels = one_hot(onp.array(mnist_dataset.train_labels), n_targets)
# Get full test dataset
mnist_dataset_test = MNIST('/tmp/mnist/', download=True, train=False)
test_images = np.array(mnist_dataset_test.test_data.numpy().reshape(len(mnist_dataset_test.test_data), -1), dtype=np.float32)
test_labels = one_hot(onp.array(mnist_dataset_test.test_labels), n_targets)
```
### Training Loop
```
import time
for epoch in range(num_epochs):
start_time = time.time()
for x, y in training_generator:
y = one_hot(y, n_targets)
params = update(params, x, y)
epoch_time = time.time() - start_time
train_acc = accuracy(params, train_images, train_labels)
test_acc = accuracy(params, test_images, test_labels)
print("Epoch {} in {:0.2f} sec".format(epoch, epoch_time))
print("Training set accuracy {}".format(train_acc))
print("Test set accuracy {}".format(test_acc))
```
We've now used the whole of the JAX API: `grad` for derivatives, `jit` for speedups and `vmap` for auto-vectorization.
We used NumPy to specify all of our computation, and borrowed the great data loaders from PyTorch, and ran the whole thing on the GPU.
| github_jupyter |
```
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.layers import Embedding, LSTM, Dense, Dropout, Bidirectional
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
from tensorflow.keras import regularizers
import tensorflow.keras.utils as ku
import numpy as np
tokenizer = Tokenizer()
!wget --no-check-certificate \
https://storage.googleapis.com/laurencemoroney-blog.appspot.com/sonnets.txt \
-O /tmp/sonnets.txt
data = open('/tmp/sonnets.txt').read()
corpus = data.lower().split("\n")
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
# create input sequences using list of tokens
input_sequences = []
for line in corpus:
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
# create predictors and label
predictors, label = input_sequences[:,:-1],input_sequences[:,-1]
label = ku.to_categorical(label, num_classes=total_words)
model = Sequential()
model.add(Embedding(total_words, 100, input_length=max_sequence_len-1))
model.add(Bidirectional(LSTM(150, return_sequences = True)))
model.add(Dropout(0.2))
model.add(LSTM(100))
model.add(Dense(total_words/2, activation='relu', kernel_regularizer=regularizers.l2(0.01)))
model.add(Dense(total_words, activation='softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
print(model.summary())
history = model.fit(predictors, label, epochs=100, verbose=1)
import matplotlib.pyplot as plt
acc = history.history['acc']
loss = history.history['loss']
epochs = range(len(acc))
plt.plot(epochs, acc, 'b', label='Training accuracy')
plt.title('Training accuracy')
plt.figure()
plt.plot(epochs, loss, 'b', label='Training Loss')
plt.title('Training loss')
plt.legend()
plt.show()
seed_text = "Help me Obi Wan Kenobi, you're my only hope"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])[0]
token_list = pad_sequences([token_list], maxlen=max_sequence_len-1, padding='pre')
predicted = model.predict_classes(token_list, verbose=0)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Hub Authors.
Licensed under the Apache License, Version 2.0 (the "License");
```
# Copyright 2019 The TensorFlow Hub Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Bangla Article Classification With TF-Hub
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/hub/tutorials/bangla_article_classifier"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/hub/blob/master/examples/colab/bangla_article_classifier.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/hub/blob/master/examples/colab/bangla_article_classifier.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/hub/examples/colab/bangla_article_classifier.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
Caution: In addition to installing Python packages with pip, this notebook uses
`sudo apt install` to install system packages: `unzip`.
This Colab is a demonstration of using [Tensorflow Hub](https://www.tensorflow.org/hub/) for text classification in non-English/local languages. Here we choose [Bangla](https://en.wikipedia.org/wiki/Bengali_language) as the local language and use pretrained word embeddings to solve a multiclass classification task where we classify Bangla news articles in 5 categories. The pretrained embeddings for Bangla comes from [fastText](https://fasttext.cc/docs/en/crawl-vectors.html) which is a library by Facebook with released pretrained word vectors for 157 languages.
We'll use TF-Hub's pretrained embedding exporter for converting the word embeddings to a text embedding module first and then use the module to train a classifier with [tf.keras](https://www.tensorflow.org/api_docs/python/tf/keras), Tensorflow's high level user friendly API to build deep learning models. Even if we are using fastText embeddings here, it's possible to export any other embeddings pretrained from other tasks and quickly get results with Tensorflow hub.
## Setup
```
%%bash
# https://github.com/pypa/setuptools/issues/1694#issuecomment-466010982
pip install gdown --no-use-pep517
%%bash
sudo apt-get install -y unzip
import os
import tensorflow as tf
import tensorflow_hub as hub
import gdown
import numpy as np
from sklearn.metrics import classification_report
import matplotlib.pyplot as plt
import seaborn as sns
```
# Dataset
We will use [BARD](https://www.researchgate.net/publication/328214545_BARD_Bangla_Article_Classification_Using_a_New_Comprehensive_Dataset) (Bangla Article Dataset) which has around 376,226 articles collected from different Bangla news portals and labelled with 5 categories: economy, state, international, sports, and entertainment. We download the file from Google Drive this ([bit.ly/BARD_DATASET](bit.ly/BARD_DATASET)) link is referring to from [this](https://github.com/tanvirfahim15/BARD-Bangla-Article-Classifier) GitHub repository.
```
gdown.download(
url='https://drive.google.com/uc?id=1Ag0jd21oRwJhVFIBohmX_ogeojVtapLy',
output='bard.zip',
quiet=True
)
%%bash
unzip -qo bard.zip
```
# Export pretrained word vectors to TF-Hub module
TF-Hub provides some useful scripts for converting word embeddings to TF-hub text embedding modules [here](https://github.com/tensorflow/hub/tree/master/examples/text_embeddings_v2). To make the module for Bangla or any other languages, we simply have to download the word embedding `.txt` or `.vec` file to the same directory as `export_v2.py` and run the script.
The exporter reads the embedding vectors and exports it to a Tensorflow [SavedModel](https://www.tensorflow.org/beta/guide/saved_model). A SavedModel contains a complete TensorFlow program including weights and graph. TF-Hub can load the SavedModel as a [module](https://www.tensorflow.org/hub/api_docs/python/hub/Module), which we will use to build the model for text classification. Since we are using `tf.keras` to build the model, we will use [hub.KerasLayer](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer), which provides a wrapper for a TF-Hub module to use as a Keras Layer.
First we will get our word embeddings from fastText and embedding exporter from TF-Hub [repo](https://github.com/tensorflow/hub).
```
%%bash
curl -O https://dl.fbaipublicfiles.com/fasttext/vectors-crawl/cc.bn.300.vec.gz
curl -O https://raw.githubusercontent.com/tensorflow/hub/master/examples/text_embeddings_v2/export_v2.py
gunzip -qf cc.bn.300.vec.gz --k
```
Then, we will run the exporter script on our embedding file. Since fastText embeddings have a header line and are pretty large (around 3.3 GB for Bangla after converting to a module) we ignore the first line and export only the first 100, 000 tokens to the text embedding module.
```
%%bash
python export_v2.py --embedding_file=cc.bn.300.vec --export_path=text_module --num_lines_to_ignore=1 --num_lines_to_use=100000
module_path = "text_module"
embedding_layer = hub.KerasLayer(module_path, trainable=False)
```
The text embedding module takes a batch of sentences in a 1D tensor of strings as input and outputs the embedding vectors of shape (batch_size, embedding_dim) corresponding to the sentences. It preprocesses the input by splitting on spaces. Word embeddings are combined to sentence embeddings with the `sqrtn` combiner(See [here](https://www.tensorflow.org/api_docs/python/tf/nn/embedding_lookup_sparse)). For demonstration we pass a list of Bangla words as input and get the corresponding embedding vectors.
```
embedding_layer(['বাস', 'বসবাস', 'ট্রেন', 'যাত্রী', 'ট্রাক'])
```
# Convert to Tensorflow Dataset
Since the dataset is really large instead of loading the entire dataset in memory we will use a generator to yield samples in run-time in batches using [Tensorflow Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) functions. The dataset is also very imbalanced, so, before using the generator, we will shuffle the dataset.
```
dir_names = ['economy', 'sports', 'entertainment', 'state', 'international']
file_paths = []
labels = []
for i, dir in enumerate(dir_names):
file_names = ["/".join([dir, name]) for name in os.listdir(dir)]
file_paths += file_names
labels += [i] * len(os.listdir(dir))
np.random.seed(42)
permutation = np.random.permutation(len(file_paths))
file_paths = np.array(file_paths)[permutation]
labels = np.array(labels)[permutation]
```
We can check the distribution of labels in the training and validation examples after shuffling.
```
train_frac = 0.8
train_size = int(len(file_paths) * train_frac)
# plot training vs validation distribution
plt.subplot(1, 2, 1)
plt.hist(labels[0:train_size])
plt.title("Train labels")
plt.subplot(1, 2, 2)
plt.hist(labels[train_size:])
plt.title("Validation labels")
plt.tight_layout()
```
To create a [Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) using a generator, we first write a generator function which reads each of the articles from `file_paths` and the labels from the label array, and yields one training example at each step. We pass this generator function to the [`tf.data.Dataset.from_generator`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#from_generator) method and specify the output types. Each training example is a tuple containing an article of `tf.string` data type and one-hot encoded label. We split the dataset with a train-validation split of 80-20 using [`tf.data.Dataset.skip`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#skip) and [`tf.data.Dataset.take`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#take) methods.
```
def load_file(path, label):
return tf.io.read_file(path), label
def make_datasets(train_size):
batch_size = 256
train_files = file_paths[:train_size]
train_labels = labels[:train_size]
train_ds = tf.data.Dataset.from_tensor_slices((train_files, train_labels))
train_ds = train_ds.map(load_file).shuffle(5000)
train_ds = train_ds.batch(batch_size).prefetch(tf.data.AUTOTUNE)
test_files = file_paths[train_size:]
test_labels = labels[train_size:]
test_ds = tf.data.Dataset.from_tensor_slices((test_files, test_labels))
test_ds = test_ds.map(load_file)
test_ds = test_ds.batch(batch_size).prefetch(tf.data.AUTOTUNE)
return train_ds, test_ds
train_data, validation_data = make_datasets(train_size)
```
# Model Training and Evaluation
Since we have already added a wrapper around our module to use it as any other layer in Keras, we can create a small [Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) model which is a linear stack of layers. We can add our text embedding module with `model.add` just like any other layer. We compile the model by specifying the loss and optimizer and train it for 10 epochs. The `tf.keras` API can handle Tensorflow Datasets as input, so we can pass a Dataset instance to the fit method for model training. Since we are using the generator function, `tf.data` will handle generating the samples, batching them and feeding them to the model.
## Model
```
def create_model():
model = tf.keras.Sequential([
tf.keras.layers.Input(shape=[], dtype=tf.string),
embedding_layer,
tf.keras.layers.Dense(64, activation="relu"),
tf.keras.layers.Dense(16, activation="relu"),
tf.keras.layers.Dense(5),
])
model.compile(loss=tf.losses.SparseCategoricalCrossentropy(from_logits=True),
optimizer="adam", metrics=['accuracy'])
return model
model = create_model()
# Create earlystopping callback
early_stopping_callback = tf.keras.callbacks.EarlyStopping(monitor='val_loss', min_delta=0, patience=3)
```
## Training
```
history = model.fit(train_data,
validation_data=validation_data,
epochs=5,
callbacks=[early_stopping_callback])
```
## Evaluation
We can visualize the accuracy and loss curves for training and validation data using the `tf.keras.callbacks.History` object returned by the `tf.keras.Model.fit` method, which contains the loss and accuracy value for each epoch.
```
# Plot training & validation accuracy values
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model accuracy')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Test'], loc='upper left')
plt.show()
```
## Prediction
We can get the predictions for the validation data and check the confusion matrix to see the model's performance for each of the 5 classes. Because `tf.keras.Model.predict` method returns an n-d array for probabilities for each class, they can be converted to class labels using `np.argmax`.
```
y_pred = model.predict(validation_data)
y_pred = np.argmax(y_pred, axis=1)
samples = file_paths[0:3]
for i, sample in enumerate(samples):
f = open(sample)
text = f.read()
print(text[0:100])
print("True Class: ", sample.split("/")[0])
print("Predicted Class: ", dir_names[y_pred[i]])
f.close()
```
## Compare Performance
Now we can take the correct labels for the validation data from `labels` and compare them with our predictions to get a [classification_report](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html).
```
y_true = np.array(labels[train_size:])
print(classification_report(y_true, y_pred, target_names=dir_names))
```
We can also compare our model's performance with the published results obtained in the original [paper](https://www.researchgate.net/publication/328214545_BARD_Bangla_Article_Classification_Using_a_New_Comprehensive_Dataset), which had a 0.96 precision .The original authors described many preprocessing steps performed on the dataset, such as dropping punctuations and digits, removing top 25 most frequest stop words. As we can see in the `classification_report`, we also manage to obtain a 0.96 precision and accuracy after training for only 5 epochs without any preprocessing!
In this example, when we created the Keras layer from our embedding module, we set the parameter`trainable=False`, which means the embedding weights will not be updated during training. Try setting it to `True` to reach around 97% accuracy using this dataset after only 2 epochs.
| github_jupyter |
```
import numpy as np, requests, pandas as pd, zipfile, StringIO
url='http://api.worldbank.org/v2/en/indicator/ny.gdp.pcap.pp.kd?downloadformat=csv'
filename='ny.gdp.pcap.pp.kd_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
gdp=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
gdp.head(2)
url='http://api.worldbank.org/v2/en/indicator/ny.gnp.pcap.pp.kd?downloadformat=csv'
filename='ny.gnp.pcap.pp.kd_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
gnp=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
gnp.head(2)
url='http://api.worldbank.org/v2/en/indicator/sp.dyn.le00.in?downloadformat=csv'
filename='sp.dyn.le00.in_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
le=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
le.head(2)
url='http://api.worldbank.org/v2/en/indicator/se.adt.litr.zs?downloadformat=csv'
filename='se.adt.litr.zs_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
alr=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
alr.head(2)
url='http://api.worldbank.org/v2/en/indicator/se.prm.enrr?downloadformat=csv'
filename='se.prm.enrr_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
ger1=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
ger1.head(2)
url='http://api.worldbank.org/v2/en/indicator/se.sec.enrr?downloadformat=csv'
filename='se.sec.enrr_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
ger2=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
ger2.head(2)
url='http://api.worldbank.org/v2/en/indicator/se.ter.enrr?downloadformat=csv'
filename='se.ter.enrr_Indicator_en_csv_v2.csv'
r = requests.get(url)
z = zipfile.ZipFile(StringIO.StringIO(r.content))
ger3=pd.read_csv(z.open(filename),skiprows=[0,1]).drop('Unnamed: 58',axis=1).drop('Indicator Code',axis=1)
ger3.head(2)
ids=pd.read_csv('http://bl.ocks.org/d/4090846/world-country-names.tsv',sep='\t').set_index(['name'],drop=True)
ids.head()
def country_name_converter(country):
if country=="Venezuela, Bolivarian Republic of": return "Venezuela (Bolivarian Republic of)"
elif country=="Tanzania, United Republic of": return "Tanzania (United Republic of)"
elif country=="Moldova, Republic of": return "Moldova (Republic of)"
elif country=="Micronesia, Federated States of": return "Micronesia (Federated States of)"
elif country=="Macedonia, the former Yugoslav Republic of": return "The former Yugoslav Republic of Macedonia"
elif country=="Korea, Republic of": return "Korea (Republic of)"
elif country=="Korea, Democratic People's Republic of": return "Korea (Democratic People's Rep. of)"
elif country=="Côte d'Ivoire": return "C\xc3\xb4te d'Ivoire"
elif country=="Iran, Islamic Republic of": return "Iran (Islamic Republic of)"
elif country=="Hong Kong": return "Hong Kong, China (SAR)"
elif country=="Palestinian Territory, Occupied": return "Palestine, State of"
elif country=="Congo, the Democratic Republic of the": return "Congo (Democratic Republic of the)"
elif country=="Bolivia, Plurinational State of": return "Bolivia (Plurinational State of)"
else: return country
import re
codes={}
for i in ids.index:
try:
a=[i]
a.append(round(float(re.sub(r'[^\d.]+', '',hdi.loc[country_name_converter(i)]\
[u"Human Development Index (HDI) Value, 2013"])),3))
a.append(round((float(re.sub(r'[^\d.]+', '',hdi.loc[country_name_converter(i)]\
[u"Life expectancy at birth (years), 2013"]))-20)/(85-20),3))
a.append(round((float(re.sub(r'[^\d.]+', '',hdi.loc[country_name_converter(i)]\
[u"Mean years of schooling (years), 2012 a"]))/15+\
float(re.sub(r'[^\d.]+', '',hdi.loc[country_name_converter(i)]\
[u"Expected years of schooling (years), 2012 a"]))/18)/2,3))
a.append(round((np.log(float(re.sub(r'[^\d.]+', '',hdi.loc[country_name_converter(i)]\
[u"Gross national income (GNI) per capita (2011 PPP $), 2013"])))-np.log(100))\
/(np.log(75000)-np.log(100)),3))
a.append(round((a[2]*a[3]*a[4])**(1.0/3.0),3))
codes[repr(ids.loc[i][0])]=a
except: pass
import json
file('hdi2.json','w').write(json.dumps(codes))
```
| github_jupyter |
# 5 - Validation - Global Installations and Waste Literature Comparisons
This journal uses Irena 2016 Installation Data, Irena 2019 Installation Data, and PV ICE installation data ran through PV_ICE tool with Irena Regular Loss Scenario Weibull and Lifetime parameters, and mostly ideal conditions, and compare it to PV ICE baseline. Output is compared to Garvin Heath & Tim Silverman's paper 2020 Mass of Installed PV and Mass of PV Waste.
# Benchmark of Results

Input is from IRENA projections:

Notes on IRENA Data:
- Installation Data < 2010 from D. Jordan (Values too low to digitize properly)
- Installation data >= 2010 from IRENA report (digitized from plot)
Other considerations:
<ul>
<li> Global projected installations from IEA/IRENA (picture below). </li>
<li> No recycling, no reuse, no repair. </li>
<li> 30-year average lifetime with early lifetime failures </li>
<li> Power to Glass conversion: 76 t/MW </li>
</ul>
```
import os
from pathlib import Path
testfolder = str(Path().resolve().parent.parent / 'PV_ICE' / 'TEMP')
# Another option using relative address; for some operative systems you might need '/' instead of '\'
# testfolder = os.path.abspath(r'..\..\PV_DEMICE\TEMP')
print ("Your simulation will be stored in %s" % testfolder)
import PV_ICE
import matplotlib.pyplot as plt
import pandas as pd
plt.rcParams.update({'font.size': 22})
plt.rcParams['figure.figsize'] = (12, 5)
```
## PV ICE
```
r1 = PV_ICE.Simulation(name='Simulation1', path=testfolder)
r1.createScenario(name='PV_ICE_base', file=r'..\baselines\baseline_modules_World.csv')
r1.scenario['PV_ICE_base'].addMaterial('glass', file=r'..\baselines\baseline_material_glass.csv')
r1.createScenario(name='PV_ICE_idealMFG', file=r'..\baselines\baseline_modules_World.csv')
r1.scenario['PV_ICE_idealMFG'].addMaterial('glass', file=r'..\baselines\baseline_material_glass.csv')
r1.scenario['PV_ICE_idealMFG'].data['mod_MFG_eff'] = 100.0
r1.scenario['PV_ICE_idealMFG'].material['glass'].materialdata['mat_MFG_eff'] = 100.0
r1.createScenario(name='Irena_2019', file=r'..\baselines\IRENA\baseline_modules_World_Irena_2019.csv')
r1.scenario['Irena_2019'].addMaterial('glass', file=r'..\baselines\IRENA\baseline_material_glass_Irena.csv')
r1.createScenario(name='A_MassBased', file=r'..\baselines\IRENA\baseline_modules_World_Irena_2019_A_MassBased.csv')
r1.scenario['A_MassBased'].addMaterial('glass', file=r'..\baselines\IRENA\baseline_material_glass_Irena_A_MassBased.csv')
r1.createScenario(name='Irena_2016', file=r'..\baselines\IRENA\baseline_modules_World_Irena_2016.csv')
r1.scenario['Irena_2016'].addMaterial('glass', file=r'..\baselines\IRENA\baseline_material_glass_Irena.csv')
Wambach = True
if Wambach:
r1.createScenario(name='Wambach2020', file=r'C:\Users\sayala\Documents\GitHub\Wambach_Baseline_DonotShare\baseline_modules_World_Wambach2020.csv')
r1.scenario['Wambach2020'].addMaterial('glass', file=r'C:\Users\sayala\Documents\GitHub\Wambach_Baseline_DonotShare\baseline_material_glass_Wambach2020.csv')
'''
r1.scenario['Garvin_2020'].data['mod_Repairing'] = 0
r1.scenario['Garvin_2020'].data['mod_Repowering'] = 0
r1.scenario['Garvin_2020'].data['mod_degradation'] = 0 # Their calculation does not consider degradation of the fleet.
#We're just calculating total waste so everythign goes to landfill
r1.scenario['Garvin_2020'].data['mod_EOL_collection_eff'] = 0
# Setting the shape of the weibull
r1.scenario['Garvin_2020'].data['mod_reliability_t50'] = 45
r1.scenario['Garvin_2020'].data['mod_reliability_t90'] = 50
# Setting Project Lifetime beyond Failures
r1.scenario['Garvin_2020'].data['mod_lifetime'] = 40
'''
pass
r1.scenario['PV_ICE_base'].data.keys()
```
Plot same plot from Garvin's paper from digitized data input
```
fig = plt.figure(figsize=(20,10))
ax1 = plt.subplot(111)
ax1.yaxis.grid()
plt.axvspan(2000, 2018, facecolor='0.9', alpha=0.5)
plt.axvspan(2018, 2050.5, facecolor='yellow', alpha=0.1)
ax1.bar(r1.scenario['Irena_2019'].data['year'], r1.scenario['Irena_2019'].data['new_Installed_Capacity_[MW]']/1000, color='gold', label='IRENA 2019')
plt.legend()
plt.xlabel('Year')
plt.ylabel('Annual Deployments (GW/yr)')
plt.xlim([2000, 2050.5])
plt.ylim([0, 400])
fig = plt.figure(figsize=(20,10))
ax1 = plt.subplot(111)
ax1.yaxis.grid()
plt.axvspan(2000, 2018, facecolor='0.9', alpha=0.5)
plt.axvspan(2018, 2050.5, facecolor='yellow', alpha=0.1)
ax1.bar(r1.scenario['Irena_2016'].data['year'], r1.scenario['Irena_2016'].data['new_Installed_Capacity_[MW]']/1000, color='gold', label='IRENA 2016')
plt.legend()
plt.xlabel('Year')
plt.ylabel('Annual Deployments (GW/yr)')
plt.xlim([2000, 2050.5])
plt.ylim([0, 400])
```
#### Adjusting input parameters to represent the inputs from the IRENA analysis:
```
r1.scenario.keys()
# Selecting only the ones we want with IRENA Assumptions
Irenify = ['Irena_2019', 'A_MassBased', 'Irena_2016', 'Wambach2020']
r1.scenMod_IRENIFY(scens=Irenify)
r1.scenario['PV_ICE_base'].data.keys()
r1.scenario['Irena_2019'].data.keys()
r1.calculateMassFlow()
r1.scenario['PV_ICE_base'].data['WeibullParams'].head(10)
r1.scenario['Irena_2019'].data['WeibullParams'].head(10)
```
## Irena Conversion from Mass to Energy -->
mat_Total_Landfilled is in g, 1 t --> 907.185 kg
1 MW --> 76 t conversion for the Mass in PV service.
Querying some of the values for plotting the flags
```
x2020 = r1.scenario['Irena_2019'].data['year'].iloc[25]
y2020 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[25]*76/1000000
t2020 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[25]/(1E12)
x2030 = r1.scenario['Irena_2019'].data['year'].iloc[35]
y2030 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[35]*76/1000000
t2030 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[35]/(1E12)
x2050 = r1.scenario['Irena_2019'].data['year'].iloc[55]
y2050 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[55]*76/1000000
t2050 = r1.scenario['Irena_2019'].data['Installed_Capacity_[W]'].iloc[55]/(1E12)
print(x2050)
if Wambach:
x2020W = r1.scenario['Wambach2020'].data['year'].iloc[40]
y2020W = r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[40]*76/1000000
t2020W = r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[40]/(1E12)
x2030W = r1.scenario['Wambach2020'].data['year'].iloc[50]
y2030W = r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[50]*76/1000000
t2030W =r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[50]/(1E12)
x2050W = r1.scenario['Wambach2020'].data['year'].iloc[70]
y2050W = r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[70]*76/1000000
t2050W = r1.scenario['Wambach2020'].data['Installed_Capacity_[W]'].iloc[70]/(1E12)
print("Flags calculated for Wambach")
```
Calculating Cumulative Waste isntead of yearly waste
Using glass for proxy of the module; glass is ~76% of the module's mass [REF]
```
cumWaste = r1.scenario['PV_ICE_base'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWaste = (cumWaste*100/76)/1000000 # Converting to tonnes
cumWasteIdeal = r1.scenario['PV_ICE_idealMFG'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWasteIdeal = (cumWasteIdeal*100/76)/1000000 # Converting to tonnes
cumWaste0 = r1.scenario['Irena_2019'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWaste0 = (cumWaste0*100/76)/1000000 # Converting to tonnes
cumWaste1 = r1.scenario['Irena_2016'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWaste1 = (cumWaste1*100/76)/1000000 # Converting to tonnes
cumWaste2 = r1.scenario['A_MassBased'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWaste2 = (cumWaste2*100/76)/1000000 # Converting to tonnes
if Wambach:
cumWaste3 = r1.scenario['Wambach2020'].material['glass'].materialdata['mat_Total_Landfilled'].cumsum()
cumWaste3 = (cumWaste3*100/76)/1000000 # Converting to tonnes
x2020_irena = 2020
y2020_irena = 3.96E+07
t2020_irena = 0.5
x2030_irena = 2030
y2030_irena = 1.24E+08
t2030_irena = 1.6
x2050_irena = 2050
y2050_irena = 3.41E+08
t2050_irena = 4.5
Garvin2020_litCumWaste_X = [2020, 2021.1, 2022.1, 2023.2, 2024.6, 2026.3, 2027.3, 2028.7,
2029.5, 2030.6, 2032.1, 2033.8, 2035.4, 2037.5, 2039.1, 2040.6, 2042, 2044, 2045.5, 2047.3,
2048.9, 2050]
Garvin2020_litCumWaste_Y = [860414.9, 1108341.4, 1383227, 1781800.6, 2295222.2, 3355623.2,
4605006.5, 6319566.5, 7886916, 8951381, 1.15E+07, 1.44E+07, 1.85E+07, 2.31E+07,
2.89E+07, 3.49E+07, 3.96E+07, 4.79E+07, 5.44E+07, 6.57E+07, 7.00E+07, 7.95E+07]
Garvin2020_litMassService_X = [2020, 2021.1, 2022.5, 2024.1, 2025.8, 2027.8, 2030, 2031.8, 2034.5,
2036.9, 2039.5, 2042.6, 2044.9, 2047.4, 2050]
Garvin2020_litMassService_Y = [3.96E+07, 4.79E+07, 5.44E+07, 6.57E+07, 7.95E+07, 1.02E+08,
1.24E+08, 1.45E+08, 1.65E+08, 1.99E+08, 2.19E+08, 2.48E+08, 2.82E+08, 3.10E+08, 3.41E+08]
```
PLOT:
```
fig = plt.figure(figsize=(10,10))
#color = 'C1', 'cornflowerblue'
#Installs
plt.semilogy(r1.scenario['PV_ICE_base'].data.year,r1.scenario['PV_ICE_base'].data['Installed_Capacity_[W]']*76/1000000, color='k', marker='o', label='Mass in Service - PV ICE baseline')
plt.semilogy(Garvin2020_litMassService_X, Garvin2020_litMassService_Y, color='C1', linewidth=5.0, label='Mass in Service - Garvin 2020')
plt.semilogy(r1.scenario['Irena_2016'].data.year,r1.scenario['Irena_2016'].data['Installed_Capacity_[W]']*76/1000000, color='C1', label='Mass in Service - Irena 2016')
plt.semilogy(r1.scenario['Irena_2019'].data.year,r1.scenario['Irena_2019'].data['Installed_Capacity_[W]']*76/1000000, color='C1', marker='o', label='Mass in Service - Irena 2019')
if Wambach:
plt.semilogy(r1.scenario['Wambach2020'].data.year,r1.scenario['Wambach2020'].data['Installed_Capacity_[W]']*76/1000000, color='C1', marker='P', markersize=12, label='Wambach 2020')
# Waste
plt.semilogy(r1.scenario['PV_ICE_base'].data.year,cumWaste, color='k', marker='o', label='Cum Waste - PV ICE baseline')
plt.semilogy(r1.scenario['PV_ICE_idealMFG'].data.year,cumWasteIdeal, color='k', marker='o', label='Cum Waste - PV ICE Ideal')
plt.semilogy(Garvin2020_litCumWaste_X, Garvin2020_litCumWaste_Y, color='cornflowerblue', linewidth=5.0, label='Cum Waste - Garvin 2020')
plt.semilogy(r1.scenario['Irena_2016'].data.year,cumWaste1, color='cornflowerblue', label='Irena 2016 w PV Ice')
plt.semilogy(r1.scenario['Irena_2019'].data.year,cumWaste0, color='cornflowerblue', marker='o', label='Irena 2019 w PV ICE')
plt.semilogy(r1.scenario['A_MassBased'].data.year,cumWaste2, 'k--', alpha=0.4, label='Irena 2019 Approach A & B')
if Wambach:
plt.semilogy(r1.scenario['Wambach2020'].data.year,cumWaste3, color='cornflowerblue', marker='P', markersize=12, label='Wambach 2020')
plt.ylim([1E4, 1E9])
plt.legend()
plt.tick_params(axis='y', which='minor')
plt.xlim([2020,2050])
plt.grid()
plt.ylabel('Mass of PV systems (t)')
plt.xlabel('Years')
offset = (0, 30)
plt.annotate(
'{:.1f} TW'.format(t2050), (x2050, y2050),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030), (x2030, y2030),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2020), (x2020, y2020),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
### IRENA
plt.annotate(
'{:.1f} TW'.format(t2020_irena), (x2020_irena, y2020_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030_irena), (x2030_irena, y2030_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2050_irena), (x2050_irena, y2050_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
# WAMBACH 2020
if Wambach:
plt.annotate(
'{:.1f} TW'.format(t2050W), (x2050W, y2050W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030W), (x2030W, y2030W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2020W), (x2020W, y2020W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.show()
fig = plt.figure(figsize=(10,5))
#color = 'C1', 'cornflowerblue'
Wambach = False
#Installs
plt.semilogy(r1.scenario['PV_ICE_base'].data.year,r1.scenario['PV_ICE_base'].data['Installed_Capacity_[W]']*76/1000000, color='k', marker='o', label='PV ICE baseline')
plt.semilogy(Garvin2020_litMassService_X, Garvin2020_litMassService_Y, color='C1', linewidth=5.0, label='Garvin 2020')
plt.semilogy(r1.scenario['Irena_2019'].data.year,r1.scenario['Irena_2019'].data['Installed_Capacity_[W]']*76/1000000, color='cornflowerblue', marker='o', label='Irena 2019')
plt.semilogy(r1.scenario['Irena_2016'].data.year,r1.scenario['Irena_2016'].data['Installed_Capacity_[W]']*76/1000000, color='cornflowerblue', label='Irena 2016')
if Wambach:
plt.semilogy(r1.scenario['Wambach2020'].data.year,r1.scenario['Wambach2020'].data['Installed_Capacity_[W]']*76/1000000, color='C1', marker='P', markersize=12, label='Wambach 2020')
plt.ylim([1E4, 1E9])
plt.legend()
plt.tick_params(axis='y', which='minor')
plt.xlim([2020,2050])
plt.grid()
plt.ylabel("PV in Service's Mass (t)")
plt.xlabel('Years')
#plt.title('Mass in Service')
offset = (0, 30)
plt.annotate(
'{:.1f} TW'.format(t2050), (x2050, y2050),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030), (x2030, y2030),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2020), (x2020, y2020),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#D3D3D3', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#D3D3D3', ec='none',
relpos=(0.5, 1.5),
)
)
### IRENA
plt.annotate(
'{:.1f} TW'.format(t2020_irena), (x2020_irena, y2020_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030_irena), (x2030_irena, y2030_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2050_irena), (x2050_irena, y2050_irena),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ff7f0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ff7f0e', ec='none',
relpos=(0.5, 1.5),
)
)
# WAMBACH 2020
if Wambach:
plt.annotate(
'{:.1f} TW'.format(t2050W), (x2050W, y2050W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2030W), (x2030W, y2030W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.annotate(
'{:.1f} TW'.format(t2020W), (x2020W, y2020W),
ha='center', va='center',
size=15,
xytext=offset, textcoords='offset points',
bbox=dict(boxstyle='round', fc='#ffdf0e', ec='none'),
arrowprops=dict(arrowstyle='wedge,tail_width=1.',
fc='#ffdf0e', ec='none',
relpos=(0.5, 1.5),
)
)
plt.show()
fig = plt.figure(figsize=(4,5))
#color = 'C1', 'cornflowerblue'
Wambach = False
# Waste
plt.semilogy(r1.scenario['PV_ICE_base'].data.year,cumWaste, color='k', marker='o', label='PV ICE baseline')
plt.semilogy(r1.scenario['PV_ICE_idealMFG'].data.year,cumWasteIdeal, color='k', marker='o', label='PV ICE Ideal')
plt.semilogy(Garvin2020_litCumWaste_X, Garvin2020_litCumWaste_Y, color='C1', linewidth=5.0, label='Garvin 2020')
plt.semilogy(r1.scenario['Irena_2016'].data.year,cumWaste1, color='cornflowerblue', label='Irena 2016')
plt.semilogy(r1.scenario['Irena_2019'].data.year,cumWaste0, color='cornflowerblue', marker='o', label='Irena 2019')
#plt.semilogy(r1.scenario['A_MassBased'].data.year,cumWaste2, 'k--', alpha=0.4, label='Irena 2019 Approach A & B')
if Wambach:
plt.semilogy(r1.scenario['Wambach2020'].data.year,cumWaste3, color='cornflowerblue', marker='P', markersize=12, label='Wambach 2020')
plt.ylim([1E4, 1E9])
#plt.legend()
plt.tick_params(axis='y', which='minor')
plt.xlim([2020,2050])
plt.grid()
plt.ylabel('Cumulative Waste (t)')
plt.xlabel('Years')
plt.title("")
```
| github_jupyter |
```
# learning using a CNN
# CNN for learning!
# learn the states of a double dot
import numpy as np
import tensorflow as tf
import glob
import os
import time
from tensorflow.contrib import learn
from tensorflow.contrib.learn.python.learn.estimators import model_fn as model_fn_lib
tf.logging.set_verbosity(tf.logging.INFO)
# application logic will be added here
def cnn_model_fn(features,labels,mode):
'''Model function for CNN'''
#input layer
input_layer = tf.cast(tf.reshape(features,[-1,30,30,1]),tf.float32)
conv1 = tf.layers.conv2d(inputs=input_layer,
filters=8,
kernel_size=[5,5],
padding="same",
activation=tf.nn.relu)
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2,2],strides=2)
#conv2 = tf.layers.conv2d(inputs=pool1,
# filters=16,
# kernel_size=[5,5],
# padding="same",
# activation=tf.nn.relu)
#pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2,2],strides=2)
flat = tf.contrib.layers.flatten(inputs=pool1)
# dense output layer
out1 = tf.layers.dense(inputs=flat,units=64,activation=tf.nn.relu)
dropout1 = tf.layers.dropout(
inputs=out1, rate=0.4, training=mode == learn.ModeKeys.TRAIN)
out = tf.layers.dense(inputs=dropout1, units=4)
loss = None
train_op = None
# Calculate loss( for both TRAIN AND EVAL modes)
if mode != learn.ModeKeys.INFER:
loss = tf.losses.mean_squared_error(out,labels['prob'])
# Configure the training op (for TRAIN mode)
if mode == learn.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=1e-3,
optimizer=tf.train.AdamOptimizer)
# Generate predictions
predictions= {
"prob" : out,
"states" : tf.argmax(out,axis=1),
}
# Returna ModelFnOps object
return model_fn_lib.ModelFnOps(mode=mode,predictions=predictions,loss=loss, train_op=train_op)
def get_train_inputs():
n_batch = 100
index = np.random.choice(n_train,n_batch,replace=False)
inp = []
oup = []
for i in index:
dat = np.load(files[i])
inp += [dat.item()['current_map']]
oup += [dat.item()['label']]
inp = np.array(inp,dtype=np.float32)
oup = np.array(oup,dtype=np.float32)
x = tf.constant(inp)
y = tf.constant(oup)
labels_dict = {}
labels_dict['prob'] = y
labels_dict['states'] = tf.argmax(y,axis=1)
return x,labels_dict
def get_val_inputs():
inp = []
oup = []
for file in files[n_train:(n_train + 100)]:
dat = np.load(file)
inp += [dat.item()['current_map']]
oup += [dat.item()['label']]
inp = np.array(inp,dtype=np.float32)
oup = np.array(oup,dtype=np.float32)
x = tf.constant(inp)
y = tf.constant(oup)
labels_dict = {}
labels_dict['prob'] = y
labels_dict['states'] = tf.argmax(y,axis=1)
return x,labels_dict
def get_test_inputs():
inp = []
oup = []
for file in files[n_train:]:
dat = np.load(file)
inp += [dat.item()['current_map']]
oup += [dat.item()['label']]
inp = np.array(inp,dtype=np.float32)
oup = np.array(oup,dtype=np.float32)
x = tf.constant(inp)
y = tf.constant(oup)
labels_dict = {}
labels_dict['prob'] = y
labels_dict['states'] = tf.argmax(y,axis=1)
return x,labels_dict
# get the data
data_folder_path = "/Users/ssk4/Downloads/data_subimage/"
files = glob.glob(data_folder_path + "*.npy")
import random
files = files[:]
n_samples = len(files)
train_sample_ratio = 0.8
n_train = int(train_sample_ratio * n_samples)
print("Total number of samples :",n_samples)
print("Training samples :",n_train)
print("Test samples :",n_samples - n_train)
st = time.time()
# create the estimator
dd_classifier = learn.Estimator(model_fn=cnn_model_fn,model_dir="/Users/ssk4/tensorflow_models/cnn_prob_test3/")
n_epochs = 10
steps_per_epoch = 100
for _ in range(n_epochs):
dd_classifier.fit(
input_fn=get_train_inputs,
steps=steps_per_epoch)
train_metrics = {
"Train accuracy" : learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="states",label_key="states"),
}
dd_classifier.evaluate(input_fn=get_train_inputs,metrics=train_metrics,steps=1)
val_metrics = {
"Val accuracy" : learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="states",label_key="states"),
}
dd_classifier.evaluate(input_fn=get_val_inputs,metrics=val_metrics,steps=1)
metrics = {
"accuracy" : learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="states",label_key="states"),
}
dd_classifier.evaluate(input_fn=get_test_inputs,metrics=metrics,steps=1)
print("Training done in",time.time()-st,"seconds.")
def accr_test_input_fn():
inp = []
oup = []
for file in files[n_train:(n_train + 100)]:
dat = np.load(file)
inp += [dat.item()['current_map']]
oup += [dat.item()['label']]
inp = np.array(inp,dtype=np.float32)
oup = np.array(oup,dtype=np.float32)
x = tf.constant(inp)
y = tf.constant(oup)
return x,y
metrics = {
"accuracy" : learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="states"),
}
pred = dd_classifier.predict(input_fn=accr_test_input_fn,as_iterable=False)
metrics = {
"accuracy" : learn.MetricSpec(metric_fn=tf.metrics.accuracy, prediction_key="states",label_key="states"),
}
dd_classifier.evaluate(input_fn=get_train_inputs,metrics=metrics,steps=1)
pred['states']
pred['prob']
out
sess = tf.Session()
x,y = accr_test_input_fn()
st = tf.argmax(y,axis=1)
curr = sess.run(x)
out = sess.run(y)
print(out)
# This notebook will be the auto-tuning routine that will find a single
# dot window using the subimage classifier
import numpy as np
import tensorflow as tf
import glob
import os
import time
import matplotlib.pyplot as plt
%matplotlib inline
tf.logging.set_verbosity(tf.logging.ERROR)
from tensorflow.contrib import learn
from tensorflow.contrib.learn.python.learn.estimators import model_fn as model_fn_lib
# application logic will be added here
def cnn_model_fn(features,labels,mode):
'''Model function for CNN'''
#input layer
input_layer = tf.cast(tf.reshape(features,[-1,30,30,1]),tf.float32)
conv1 = tf.layers.conv2d(inputs=input_layer,
filters=32,
kernel_size=[5,5],
padding="same",
activation=tf.nn.relu)
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2,2],strides=2)
conv2 = tf.layers.conv2d(inputs=pool1,
filters=64,
kernel_size=[5,5],
padding="same",
activation=tf.nn.relu)
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2,2],strides=2)
flat = tf.contrib.layers.flatten(inputs=pool2)
# dense output layer
out1 = tf.layers.dense(inputs=flat,units=1024,activation=tf.nn.relu)
dropout1 = tf.layers.dropout(
inputs=out1, rate=0.4, training=mode == learn.ModeKeys.TRAIN)
out = tf.layers.dense(inputs=dropout1, units=4)
loss = None
train_op = None
# Calculate loss( for both TRAIN AND EVAL modes)
if mode != learn.ModeKeys.INFER:
loss = tf.losses.mean_squared_error(out,labels['prob'])
# Configure the training op (for TRAIN mode)
if mode == learn.ModeKeys.TRAIN:
train_op = tf.contrib.layers.optimize_loss(
loss=loss,
global_step=tf.contrib.framework.get_global_step(),
learning_rate=1e-3,
optimizer=tf.train.AdamOptimizer)
# Generate predictions
predictions= {
"prob" : out,
"states" : tf.argmax(out,axis=1),
}
# Returna ModelFnOps object
return model_fn_lib.ModelFnOps(mode=mode,predictions=predictions,loss=loss, train_op=train_op)
dd_classifier = learn.Estimator(model_fn=cnn_model_fn,model_dir="/Users/ssk4/tensorflow_models/cnn_prob/")
# Testing of the experimental data loading
import numpy as np
import scipy.interpolate
import glob
import matplotlib.pyplot as plt
%matplotlib inline
data_folder_path = "/Users/ssk4/Downloads/exp_data/"
files = glob.glob(data_folder_path + "*.dat")
# Data format is V_LGD I_DC(nA) V_LGS I_AC(nA) t(sec)
# The format of the loaded array is [num_points,5]
dat = np.loadtxt(files[12])
sub_size = 100
grid_x = np.linspace(np.min(dat[:,0]),np.max(dat[:,0]),sub_size)
grid_y = np.linspace(np.min(dat[:,2]),np.max(dat[:,2]),sub_size)
xx,yy = np.meshgrid(grid_x,grid_y)
interpolated_data = scipy.interpolate.griddata((dat[:,0],dat[:,2]),dat[:,1],(xx, yy), method='nearest')
xx, yy = np.meshgrid(grid_x,grid_y)
plt.pcolor(xx,yy,interpolated_data)
plt.xlabel(r'$V_{d1}$',fontsize=16)
plt.ylabel(r'$V_{d2}$',fontsize=16)
cb = plt.colorbar()
cb.set_label('Current (arb. units)',fontsize=16)
sub_size = 30
grid_x = np.linspace(np.min(dat[:,0]),np.max(dat[:,0]),sub_size)
grid_y = np.linspace(np.min(dat[:,2]),np.max(dat[:,2]),sub_size)
xx,yy = np.meshgrid(grid_x,grid_y)
interpolated_data = np.abs(scipy.interpolate.griddata((dat[:,0],dat[:,2]),dat[:,1],(xx, yy), method='nearest'))
interpolated_data = interpolated_data/np.max(interpolated_data)
state = dd_classifier.predict(x=interpolated_data,as_iterable=False)
plt.savefig("/Users/ssk4/Desktop/data1.png",dpi=600)
xbins = np.arange(4)
width = 0.35
fig, ax = plt.subplots()
y = np.abs(state['prob'][0])
y = y/np.sum(y)
color = ["g" if x > 0.3 else "r" for x in y]
ax.bar(xbins,y,color=color)
ax.set_xticks(xbins)
ax.set_ylabel(r'$\alpha$ Probability',fontsize=16)
ax.set_xticklabels(('ShortCircuit','QPC','SD','DD'),fontsize=16)
plt.savefig("/Users/ssk4/Desktop/data1_bar.png",dpi=600)
# Testing of the experimental data loading
import numpy as np
import scipy.interpolate
data_folder_path = "/Users/ssk4/Downloads/exp_data/"
files = glob.glob(data_folder_path + "*.dat")
# Data format is V_LGD I_DC(nA) V_LGS I_AC(nA) t(sec)
# The format of the loaded array is [num_points,5]
index = np.random.randint(len(files))
for i in range(len(files)):
dat = np.loadtxt(files[i])
sub_size = 30
grid_x = np.linspace(np.min(dat[:,0]),np.max(dat[:,0]),sub_size)
grid_y = np.linspace(np.min(dat[:,2]),np.max(dat[:,2]),sub_size)
xx,yy = np.meshgrid(grid_x,grid_y)
interpolated_data = scipy.interpolate.griddata((dat[:,0],dat[:,2]),dat[:,1],(xx, yy), method='nearest')
tf.logging.set_verbosity(tf.logging.ERROR)
#import matplotlib.pyplot as plt
#%matplotlib inline
#plt.pcolor(interpolated_data)
print(i,dd_classifier.predict(x=interpolated_data,as_iterable=False)['states'])
dat = np.loadtxt(files[53])
sub_size = 30
grid_x = np.linspace(np.min(dat[:,0]),np.max(dat[:,0]),sub_size)
grid_y = np.linspace(np.min(dat[:,2]),np.max(dat[:,2]),sub_size)
xx,yy = np.meshgrid(grid_x,grid_y)
interpolated_data = scipy.interpolate.griddata((dat[:,0],dat[:,2]),dat[:,1],(xx, yy), method='nearest')
import matplotlib.pyplot as plt
%matplotlib inline
plt.pcolor(interpolated_data)
interpolated_data
dd_classifier.export_savedmodel('/tmp/dd_classifier',serving_input_fn=)
```
| github_jupyter |
# Implementing the Kalman Model
### Importing required libraries
```
import numpy as np
import pandas as pd
from scipy.optimize import minimize
import matplotlib.pyplot as plt
from math import sqrt
from datetime import datetime
import datetime
from scipy.optimize import minimize
from sklearn.metrics import mean_squared_error
from math import sqrt
# extract data from various Internet sources into a pandas DataFrame
import pandas_datareader as web
!pip install pandas_datareader
start = datetime.datetime(2014, 1, 1)
end = datetime.datetime(2019, 1, 1)
df_amzn = web.DataReader('AMZN', 'yahoo', start, end)
amzn= df_amzn
amzn=amzn.reset_index()
amzn['Date'] = pd.to_datetime(amzn['Date'])
# corresponding csv file is saved in an ouput directory
#df_amzn.to_csv('data/data.csv')
amzn
amzn.columns
amzn.describe()
```
In Stock trading, the **high**, **low** refers to the maximum and the minimum prices in a given time period. **Open** and **Close** are thhighLowOpenClosee prices at which a stock began and ended trading in the same period. **Volume** is the total amount of trading activity. Adjusted values factor in corporate actions such as dividends, stock splits, and new share issuance.
```
def Kalman_Filter(Y):
S = Y.shape[0]
S = S + 1
"Initialize Params:"
Z = param0[0]
T = param0[1]
H = param0[2]
Q = param0[3]
# "Kalman Filter Starts:"
u_predict = np.zeros(S)
u_update = np.zeros(S)
P_predict = np.zeros(S)
P_update = np.zeros(S)
v = np.zeros(S)
F = np.zeros(S)
KF_Dens = np.zeros(S)
for s in range(1,S):
if s == 1:
P_update[s] = 1000
P_predict[s] = T*P_update[1]*np.transpose(T)+Q
else:
F[s]= Z*P_predict[s-1]*np.transpose(Z)+H
v[s] = Y[s-1] - Z*u_predict[s-1]
u_update[s] = u_predict[s-1]+P_predict[s-1]*np.transpose(Z)*(1/F[s])*v[s]
u_predict[s] = T*u_predict[s];
P_update[s] = P_predict[s-1]-P_predict[s-1]*np.transpose(Z)*(1/F[s])*Z*P_predict[s-1]
P_predict[s] = T*P_update[s]*np.transpose(T)+Q
Likelihood = np.sum(KF_Dens[1:-1])
return Likelihood
def Kalman_Smoother(params, Y):
S = Y.shape[0]
S = S + 1
"Initialize Params:"
Z = params[0]
T = params[1]
H = params[2]
Q = params[3]
"Kalman Filter Starts:"
u_predict = np.zeros(S)
u_update = np.zeros(S)
P_predict = np.zeros(S)
P_update = np.zeros(S)
v = np.zeros(S)
F = np.zeros(S)
for s in range(1,S):
if s == 1:
P_update[s] = 1000
P_predict[s] = T*P_update[1]*np.transpose(T)+Q
else:
# "Please fill this part."
F[s]= Z*P_predict[s-1]*np.transpose(Z)+H
v[s] = Y[s-1] - Z*u_predict[s-1]
u_update[s] = u_predict[s-1]+P_predict[s-1]*np.transpose(Z)*(1/F[s])*v[s]
u_predict[s] = T*u_predict[s];
P_update[s] = P_predict[s-1]-P_predict[s-1]*np.transpose(Z)*(1/F[s])*Z*P_predict[s-1]
P_predict[s] = T*P_update[s]*np.transpose(T)+Q
u_smooth = np.zeros(S)
P_smooth = np.zeros(S)
u_smooth[S-1] = u_update[S-1]
P_smooth[S-1] = P_update[S-1]
for t in range(S-1,0,-1):
u_smooth[t-1] = u_update[t] +P_update[t]*np.transpose(T)/P_predict[t]*(u_smooth[t]- T*u_update[t])
P_smooth[t-1] = P_update[t] + (P_update[t]*np.transpose(T)/P_predict[t]*(P_smooth[t]-P_update[t])/P_update[t]*T*P_update[t])
u_smooth = u_smooth[0:-1]
return u_smooth
# amzn = pd.read_csv("AMZN.csv")
# amzn['Typical_Price'] = amzn[['High','Low','Close']].mean(axis=1)
# amzn['lrets'] = (np.log(amzn.Close) - np.log(amzn.Close.shift(1))) * 100.
# amzn.head()
Y = amzn['Open']
T = Y.shape[0]
mu = 1196;
param0 = np.array([0.3, 0.9, 0.8, 1.1])
param_star = minimize(Kalman_Filter, param0, method='BFGS', options={'xtol': 1e-8, 'disp': True})
u = Kalman_Smoother(param_star.x, Y)
timevec = np.linspace(1,T,T)
fig= plt.figure(figsize=(14,6))
plt.plot(timevec, Y,'r-', label='Actual')
plt.plot(timevec, u,'b:', label='Predicted')
plt.legend(loc='upper right')
plt.title("Kalman Filtering")
plt.show()
Y = amzn['Close']
T = Y.shape[0]
mu = 1196;
param0 = np.array([0.3, 0.9, 0.8, 1.1])
param_star = minimize(Kalman_Filter, param0, method='BFGS', options={'xtol': 1e-8, 'disp': True})
u = Kalman_Smoother(param_star.x, Y)
timevec = np.linspace(1,T,T)
fig= plt.figure(figsize=(14,6))
plt.plot(timevec, Y,'r-', label='Actual')
plt.plot(timevec, u,'b:', label='Predicted')
plt.legend(loc='upper right')
plt.title("Kalman Filtering")
plt.show()
results = pd.DataFrame({'Actual': list(Y),
'Predicted' : list(u),
'Date':amzn['Date'],
'Open':amzn['Open'],
'Close':amzn['Close']
})
results.set_index('Date',inplace = True)
results.head(10)
dif = pd.DataFrame({'Actual':list(Y),
'Predicted':list(u)})
```
# Long Short Day trading:
* if predicted > yesterdays close, buy and sell at end of day
* if predicted < yesterdays close, sell and buy at end of day
```
amount = 10000
signal = 0
Amount = []
balance = 0
action = []
portfolio = 0
Portfolio = []
stocks = 0
Stocks = []
for i in range(len(results)):
if results['Predicted'][i] > results['Actual'][i-1]:
action.append('Buy at Open')
stocks = int(amount/results['Open'][i])
balance = int(amount%results['Close'][i])
portfolio = stocks * results ['Open'][i]
print(i,'Buy at Open',round(portfolio,2),stocks,round(balance,2))
action.append('Sell at End')
portfolio = stocks * results['Close'][i]
signal = 0
stocks = 0
amount = balance + portfolio
portfolio = 0
balance = 0
print(i,'Sell at Close',round(amount,2),balance)
Amount.append(amount)
else:
action.append('Sell at Open')
stocks = int(amount/results['Open'][i])
balance = int(amount%results['Close'][i])
portfolio = stocks * results ['Open'][i]
print(i,'Sell at Open',round(portfolio,2),'-',stocks,round(balance,2))
action.append('Buy at Close')
portfolio = stocks * results['Close'][i]
signal = 0
stocks = 0
amount = balance + portfolio
portfolio = 0
balance = 0
print(i,'Buy Back at Close',round(amount,2),balance)
Amount.append(amount)
print('\n')
results['Amount'] = list(Amount)
results['Returns'] = results['Amount'].pct_change()
results.head()
results.tail()
```
#### Sharpe and is used to help investors understand the return of an investment compared to its risk. 12 The ratio is the average return earned in excess of the risk-free rate per unit of volatility or total risk.
```
mean_returns = results['Returns'].mean()
sd = results['Returns'].std()
print(mean_returns,sd)
Market_RF = 0.0464
Sharpe_Ratio = np.sqrt(878)*(mean_returns)/sd
Sharpe_Ratio
```
#### A good model should have an RMSE value less than 180.
#### RMSE Value of Amazon
```
from sklearn.metrics import mean_squared_error
from math import sqrt
rms = sqrt(mean_squared_error(results['Actual'], results['Predicted']))
rms
```
| github_jupyter |
# Generative Adversarial Nets
Training a generative adversarial network to sample from a Gaussian distribution. This is a toy problem, takes < 3 minutes to run on a modest 1.2GHz CPU.
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from scipy.stats import norm
%matplotlib inline
```
Target distribution $p_{data}$
```
mu,sigma=-1,1
xs=np.linspace(-5,5,1000)
plt.plot(xs, norm.pdf(xs,loc=mu,scale=sigma))
#plt.savefig('fig0.png')
TRAIN_ITERS=10000
M=200 # minibatch size
# MLP - used for D_pre, D1, D2, G networks
def mlp(input, output_dim):
# construct learnable parameters within local scope
w1=tf.get_variable("w0", [input.get_shape()[1], 6], initializer=tf.random_normal_initializer())
b1=tf.get_variable("b0", [6], initializer=tf.constant_initializer(0.0))
w2=tf.get_variable("w1", [6, 5], initializer=tf.random_normal_initializer())
b2=tf.get_variable("b1", [5], initializer=tf.constant_initializer(0.0))
w3=tf.get_variable("w2", [5,output_dim], initializer=tf.random_normal_initializer())
b3=tf.get_variable("b2", [output_dim], initializer=tf.constant_initializer(0.0))
# nn operators
fc1=tf.nn.tanh(tf.matmul(input,w1)+b1)
fc2=tf.nn.tanh(tf.matmul(fc1,w2)+b2)
fc3=tf.nn.tanh(tf.matmul(fc2,w3)+b3)
return fc3, [w1,b1,w2,b2,w3,b3]
# re-used for optimizing all networks
def momentum_optimizer(loss,var_list):
batch = tf.Variable(0)
learning_rate = tf.train.exponential_decay(
0.001, # Base learning rate.
batch, # Current index into the dataset.
TRAIN_ITERS // 4, # Decay step - this decays 4 times throughout training process.
0.95, # Decay rate.
staircase=True)
#optimizer=tf.train.GradientDescentOptimizer(learning_rate).minimize(loss,global_step=batch,var_list=var_list)
optimizer=tf.train.MomentumOptimizer(learning_rate,0.6).minimize(loss,global_step=batch,var_list=var_list)
return optimizer
```
# Pre-train Decision Surface
If decider is reasonably accurate to start, we get much faster convergence.
```
with tf.variable_scope("D_pre"):
input_node=tf.placeholder(tf.float32, shape=(M,1))
train_labels=tf.placeholder(tf.float32,shape=(M,1))
D,theta=mlp(input_node,1)
loss=tf.reduce_mean(tf.square(D-train_labels))
optimizer=momentum_optimizer(loss,None)
sess=tf.InteractiveSession()
tf.global_variables_initializer().run()
# plot decision surface
def plot_d0(D,input_node):
f,ax=plt.subplots(1)
# p_data
xs=np.linspace(-5,5,1000)
ax.plot(xs, norm.pdf(xs,loc=mu,scale=sigma), label='p_data')
# decision boundary
r=1000 # resolution (number of points)
xs=np.linspace(-5,5,r)
ds=np.zeros((r,1)) # decision surface
# process multiple points in parallel in a minibatch
for i in range(r/M):
x=np.reshape(xs[M*i:M*(i+1)],(int(M),1))
ds[M*i:M*(i+1)]=sess.run(D,{input_node: x})
ax.plot(xs, ds, label='decision boundary')
ax.set_ylim(0,1.1)
plt.legend()
plot_d0(D,input_node)
plt.title('Initial Decision Boundary')
#plt.savefig('fig1.png')
lh=np.zeros(1000)
for i in range(1000):
#d=np.random.normal(mu,sigma,M)
d=(np.random.random(M)-0.5) * 10.0 # instead of sampling only from gaussian, want the domain to be covered as uniformly as possible
labels=norm.pdf(d,loc=mu,scale=sigma)
lh[i],_=sess.run([loss,optimizer], {input_node: np.reshape(d,(M,1)), train_labels: np.reshape(labels,(M,1))})
# training loss
plt.plot(lh)
plt.title('Training Loss')
plot_d0(D,input_node)
#plt.savefig('fig2.png')
# copy the learned weights over into a tmp array
weightsD=sess.run(theta)
# close the pre-training session
sess.close()
```
# Build Net
Now to build the actual generative adversarial network
```
with tf.variable_scope("G"):
z_node=tf.placeholder(tf.float32, shape=(M,1)) # M uniform01 floats
G,theta_g=mlp(z_node,1) # generate normal transformation of Z
G=tf.multiply(5.0,G) # scale up by 5 to match range
with tf.variable_scope("D") as scope:
# D(x)
x_node=tf.placeholder(tf.float32, shape=(M,1)) # input M normally distributed floats
fc,theta_d=mlp(x_node,1) # output likelihood of being normally distributed
D1=tf.maximum(tf.minimum(fc,.99), 0.01) # clamp as a probability
# make a copy of D that uses the same variables, but takes in G as input
scope.reuse_variables()
fc,theta_d=mlp(G,1)
D2=tf.maximum(tf.minimum(fc,.99), 0.01)
obj_d=tf.reduce_mean(tf.log(D1)+tf.log(1-D2))
obj_g=tf.reduce_mean(tf.log(D2))
# set up optimizer for G,D
opt_d=momentum_optimizer(1-obj_d, theta_d)
opt_g=momentum_optimizer(1-obj_g, theta_g) # maximize log(D(G(z)))
sess=tf.InteractiveSession()
tf.global_variables_initializer().run()
# copy weights from pre-training over to new D network
for i,v in enumerate(theta_d):
sess.run(v.assign(weightsD[i]))
def plot_fig():
# plots pg, pdata, decision boundary
f,ax=plt.subplots(1)
# p_data
xs=np.linspace(-5,5,1000)
ax.plot(xs, norm.pdf(xs,loc=mu,scale=sigma), label='p_data')
# decision boundary
r=5000 # resolution (number of points)
xs=np.linspace(-5,5,r)
ds=np.zeros((r,1)) # decision surface
# process multiple points in parallel in same minibatch
for i in range(r/M):
x=np.reshape(xs[M*i:M*(i+1)],(M,1))
ds[M*i:M*(i+1)]=sess.run(D1,{x_node: x})
ax.plot(xs, ds, label='decision boundary')
# distribution of inverse-mapped points
zs=np.linspace(-5,5,r)
gs=np.zeros((r,1)) # generator function
for i in range(r/M):
z=np.reshape(zs[M*i:M*(i+1)],(M,1))
gs[M*i:M*(i+1)]=sess.run(G,{z_node: z})
histc, edges = np.histogram(gs, bins = 10)
ax.plot(np.linspace(-5,5,10), histc/float(r), label='p_g')
# ylim, legend
ax.set_ylim(0,1.1)
plt.legend()
# initial conditions
plot_fig()
plt.title('Before Training')
#plt.savefig('fig3.png')
# Algorithm 1 of Goodfellow et al 2014
k=1
histd, histg= np.zeros(TRAIN_ITERS), np.zeros(TRAIN_ITERS)
for i in range(TRAIN_ITERS):
for j in range(k):
x= np.random.normal(mu,sigma,M) # sampled m-batch from p_data
x.sort()
z= np.linspace(-5.0,5.0,M)+np.random.random(M)*0.01 # sample m-batch from noise prior
histd[i],_=sess.run([obj_d,opt_d], {x_node: np.reshape(x,(M,1)), z_node: np.reshape(z,(M,1))})
z= np.linspace(-5.0,5.0,M)+np.random.random(M)*0.01 # sample noise prior
histg[i],_=sess.run([obj_g,opt_g], {z_node: np.reshape(z,(M,1))}) # update generator
if i % (TRAIN_ITERS//10) == 0:
print(float(i)/float(TRAIN_ITERS))
plt.plot(range(TRAIN_ITERS),histd, label='obj_d')
plt.plot(range(TRAIN_ITERS), 1-histg, label='obj_g')
plt.legend()
#plt.savefig('fig4.png')
plot_fig()
#plt.savefig('fig5.png')
```
| github_jupyter |
# Installation and Quickstart
This page will explain how to get started using the Summer library in your own project. If you would like to setup Summer as a contributor, or to run the [code examples](http://summerepi.com/examples/index.html), use [these instructions](https://github.com/monash-emu/summer/blob/master/docs/dev-setup.md) instead.
## Prerequisites
This library uses numerical computing packages such as NumPy, SciPy and Numba, which can be difficult to install on Windows and MacOS. As such, we recommend that you use the Anaconda Python distribution to install and run Summer. You can install a minimal Anaconda distribution ("Miniconda") [here](https://docs.conda.io/en/latest/miniconda.html).
In any case, you will need to have Python 3.6+ and Pip (the Python package manager) available.
If you are using Miniconda, then you will need to create an "environment" where you can install Summer and other packages you need for your project. You can create a new environment as follows:
```bash
# Create a new Anaconda environment.
conda create -n myprojectname python=3.6
# Make the new Anaconda environment active in your shell session.
conda activate myprojectname
```
## Installation
You can install summer from PyPI using the Pip package manager
```bash
pip install summerepi
```
Then you can import the library as `summer` and get started building compartmental disease models. You can find a [list of examples](./examples/index.html) and [detailed API documentation](/api/index.html) on this site.
Note the above method installs the latest 'release' version of summer, but that this documentation is based on the current Github master version of summer, which may contain new features or changes to the API.
To install summer directly from Github, use the following command instead
```bash
pip install git+https://github.com/monash-emu/summer.git
```
## Quick Example Model
This is a short example on how summer can be used. See the [list of examples](./examples/index.html) for more.
```
import matplotlib.pyplot as plt
from summer import CompartmentalModel
# Create a model.
model = CompartmentalModel(
times=[1990, 2025],
compartments=["S", "I", "R"],
infectious_compartments=["I"],
timestep=0.1,
)
# Add people to the model.
model.set_initial_population(distribution={"S": 1000, "I": 10})
# Add intercompartmental flows.
model.add_infection_frequency_flow(name="infection", contact_rate=1.2, source="S", dest="I")
model.add_transition_flow(name="recovery", fractional_rate=1/6, source="I", dest="R")
model.add_death_flow(name="infection_death", death_rate=0.5, source="I")
# Spice up the model by importing 500 infected people over the course of 2010.
def get_infected_imports(t, cv=None):
return 500 if 2010 < t <= 2011 else 0
model.add_importation_flow('infected_imports', get_infected_imports, 'I', split_imports=True)
# Run the model
model.run()
# Plot the model results.
fig, ax = plt.subplots(1, 1, figsize=(12, 6), dpi=120)
for i in range(model.outputs.shape[1]):
ax.plot(model.times, model.outputs.T[i])
ax.set_title("SIR Model Outputs")
ax.set_xlabel("Year")
ax.set_ylabel("Compartment size")
ax.legend(["S", "I", "R"])
plt.show()
```
| github_jupyter |
```
import csv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df_anx = pd.read_csv('minimap_hmm.csv')
df_anx = df_anx.replace(r'^s*$', float('NaN'), regex = True)
df_anx = df_anx.replace(r' ', float('NaN'), regex = True)
df_anx = df_anx[df_anx['anger_premeasure'].notna()]
df_anx = df_anx[df_anx['anxiety_premeasure'].notna()]
uids_list = df_anx['uid'].unique()
print(len(uids_list))
```
### Group Assignments
1. High anger, High anxiety
2. Low anger, High anxiety
3. High anger, Low anxiety
4. Low anger, Low anxiety
```
mean_anger = np.mean([float(elem) for elem in df_anx['anger_premeasure'].to_numpy()])
mean_anx = np.mean([float(elem) for elem in df_anx['anxiety_premeasure'].to_numpy()])
uid_to_anger_anx = {}
uid_to_group = {}
for index, row in df_anx.iterrows():
uid = row['uid']
anger = float(row['anger_premeasure'])
anx = float(row['anxiety_premeasure'])
if anger >= mean_anger:
if anx >= mean_anx:
group = 1
else:
group = 3
else:
if anx >= mean_anx:
group = 2
else:
group = 4
group = 1
uid_to_anger_anx[uid] = {'anger': anger, 'anxiety': anx, 'group': group}
uid_to_group[uid] = group
valid_uids = list(uid_to_group.keys())
```
## Compute Metrics
```
robot_X =[84, 84, 84, 84, 84, 84, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 77, 77, 76, 76, 75, 75, 74, 74, 73, 73, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 69, 69, 68, 68, 68, 68, 68, 68, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 70, 70, 71, 71, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 68, 68, 68, 68, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 63, 63, 62, 62, 61, 61, 60, 60, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 61, 61, 62, 62, 62, 62, 62, 62, 62, 62, 63, 63, 63, 63, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 65, 65, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 65, 65, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 63, 63, 62, 62, 62, 62, 61, 61, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 59, 59, 59, 59, 59, 59, 60, 60, 61, 61, 62, 62, 63, 63, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 69, 69, 70, 70, 71, 71, 72, 72, 73, 73, 74, 74, 75, 75, 76, 76, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 79, 79, 80, 80, 80, 80, 80, 80, 80, 80, 81, 81, 81, 81, 82, 82, 82, 82, 82, 82, 82, 82, 82, 82, 83, 83, 82, 82, 82, 82, 81, 81, 81, 81, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 80, 80, 80, 80, 80, 80, 79, 79, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 78, 78, 79, 79, 80, 80, 81, 81, 82, 82, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 77, 77, 77, 77, 77, 77, 77, 77, 76, 76, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 74, 74, 73, 73, 72, 72, 71, 71, 70, 70, 69, 69, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 63, 63, 62, 62, 61, 61, 60, 60, 59, 59, 58, 58, 57, 57, 56, 56, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 56, 56, 57, 57, 58, 58, 58, 58, 59, 59, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 58, 58, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 58, 58, 57, 57, 56, 56, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 56, 56, 57, 57, 58, 58, 59, 59, 60, 60, 61, 61, 62, 62, 63, 63, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 69, 69, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 71, 71, 72, 72, 73, 73, 74, 74, 75, 75, 76, 76, 77, 77, 78, 78, 78, 78, 78, 78, 78, 78, 79, 79, 80, 80, 81, 81, 82, 82, 83, 83, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 85, 85, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 88, 88, 88, 88, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 88, 88, 87, 87, 86, 86, 86, 86, 86, 86, 86, 86, 85, 85, 84, 84, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 77, 77, 77, 77, 76, 76, 76, 76, 75, 75, 75, 75, 74, 74, 74, 74, 73, 73, 72, 72, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 72, 72, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 70, 70, 69, 69, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 69, 69]
robot_Y =[38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 39, 39, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 42, 42, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 45, 45, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 47, 47, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 45, 45, 44, 44, 44, 44, 44, 44, 45, 45, 45, 45, 45, 45, 45, 45, 45, 45, 46, 46, 45, 45, 45, 45, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 39, 39, 38, 38, 37, 37, 36, 36, 35, 35, 34, 34, 33, 33, 32, 32, 31, 31, 30, 30, 29, 29, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 30, 30, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 30, 30, 31, 31, 32, 32, 33, 33, 34, 34, 34, 34, 34, 34, 34, 34, 34, 34, 35, 35, 34, 34, 34, 34, 34, 34, 35, 35, 36, 36, 37, 37, 37, 37, 37, 37, 36, 36, 35, 35, 34, 34, 33, 33, 32, 32, 31, 31, 30, 30, 29, 29, 28, 28, 27, 27, 26, 26, 25, 25, 24, 24, 23, 23, 22, 22, 21, 21, 20, 20, 19, 19, 18, 18, 17, 17, 16, 16, 15, 15, 14, 14, 13, 13, 12, 12, 11, 11, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 3, 3, 3, 3, 3, 3, 4, 4, 5, 5, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 5, 5, 5, 5, 5, 5, 5, 5, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 9, 9, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 1, 1, 1, 1, 1, 1, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 3, 3, 3, 3, 3, 3, 4, 4, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 11, 11, 12, 12, 12, 12, 12, 12, 12, 12, 12, 12, 13, 13, 12, 12, 12, 12, 12, 12, 12, 12, 12, 12, 11, 11, 10, 10, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 8, 8, 8, 8, 7, 7, 7, 7, 6, 6, 6, 6, 5, 5, 5, 5, 5, 5, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 11, 11, 12, 12, 13, 13, 13, 13, 14, 14, 15, 15, 16, 16, 16, 16, 16, 16, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 16, 16, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 18, 18, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 18, 18, 19, 19, 20, 20, 21, 21, 22, 22, 23, 23, 24, 24, 25, 25, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 27, 27, 26, 26, 26, 26, 26, 26, 27, 27, 27, 27, 27, 27, 27, 27, 27, 27, 28, 28, 27, 27, 27, 27, 27, 27, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 29, 29, 28, 28, 28, 28, 27, 27, 26, 26, 25, 25, 24, 24, 23, 23, 22, 22, 21, 21, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20]
assert len(robot_X) == len(robot_Y)
# if (minutes ==4 && seconds < 45 && seconds >= 41) {
# otherNumSteps += 0;
# }
# else if (minutes ==4 && seconds < 26 && seconds >= 24) {
# otherNumSteps += 0;
# }
# else if (minutes ==3 && seconds < 46 && seconds >= 43) {
# otherNumSteps += 0;
# }
# else if (minutes ==2 && seconds < 50 && seconds >= 48) {
# otherNumSteps += 0;
# }
# else if (minutes ==1 && seconds < 32 && seconds >= 30) {
# otherNumSteps += 0;
# }
plt.scatter(robot_X, robot_Y)
```
# Process Robot Trajectory
```
N = len(robot_X)
print(N)
difference = int(N/10)
print(difference)
min_to_robot_traj = {(5,0):[]}
counter = 0
for minute in [4,3,2, 1, 0]:
for second in [30, 0]:
counter += 1
keyname = (minute, second)
t_robot_x = robot_X[difference*(counter-1): difference*(counter)]
t_robot_y = robot_Y[difference*(counter-1): difference*(counter)]
traj_robot = [(t_robot_x[i], t_robot_y[i]) for i in range(len(t_robot_x))]
min_to_robot_traj[keyname] = traj_robot
min_to_robot_traj
```
# Get Human Data
```
df = pd.read_csv('minimap_study_3_data.csv')
i = 40
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['time_spent'], ascending=False)
def compute_minute_to_traj(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_traj = {}
minute_to_msgs = {}
minute_to_victims = {}
curr_min = 5
curr_sec = 0
where_start = np.where(times == 'start')[0]
if len(where_start) == 0:
minute_to_traj[(curr_min, curr_sec)] = []
else:
# print("where_start = ", where_start)
start_idx = where_start[0]
t = str(times[start_idx])
traj_t = str(traj[start_idx])
msgs_t = str(sent_msgs[start_idx])
vic_t = str(victims[start_idx])
if traj_t == 'nan':
curr_traj = []
else:
curr_traj = [eval(x) for x in traj_t.split(';')]
if msgs_t == 'nan':
curr_msgs = []
else:
curr_msgs = msgs_t
minute_to_traj[(curr_min, curr_sec)] = curr_traj
# minute_to_msgs[(curr_min, curr_sec)] = minute_to_msgs
# minute_to_victims[(curr_min, curr_sec)] = []
if curr_sec == 0:
curr_min -= 1
curr_sec = 30
curr_traj = []
prev_min = 5
prev_seconds = 60
for i in range(len(traj)):
t = str(times[i])
traj_t = str(traj[i])
msgs_t = sent_msgs[i]
vic_t = victims[i]
if ':' not in t:
continue
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
if traj_t != 'nan':
curr_t_traj = [eval(x) for x in traj_t.split(';')]
# print("INPUTS TO ROUND")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
if t_min == curr_min:
if t_sec == curr_sec:
curr_traj.extend(curr_t_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
if curr_sec == 0:
curr_min -= 1
curr_sec = 30
else:
curr_sec = 0
curr_traj = []
elif t_sec > curr_sec:
curr_traj.extend(curr_t_traj)
elif t_sec < curr_sec and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:45-2:30
diff_in_next_section = abs(curr_sec - t_sec) #2:30-2
# 2- 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
elif t_sec == 0 and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:45 - 2:30
diff_in_next_section = abs(curr_sec - t_sec) # 2:30- 2
# 2 - 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:15 - 2:00
diff_in_next_section = abs(curr_sec - t_sec) # 2:00- 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
curr_sec = 0
curr_traj = []
elif t_min == curr_min-1:
if t_sec > curr_sec and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - curr_sec)
diff_in_next_section = abs(curr_sec - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec ==30:
diff_in_past_section = abs(prev_seconds - 30) #2:40-2:30
diff_in_mid_section = 30 # 2:30-2
diff_in_next_section = 30 # 2-1:30
#1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:next_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - 0) #2:04 -2
diff_in_mid_section = 30 # 2 - 1:30
diff_in_next_section = 30 # 1:30 - 1
#1-0:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:next_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 0 and curr_sec ==30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_section = 30 #3:30 - 3
# diff_in_next_section = abs(30 - t_sec) # 3 - 2:30
# 2:30 - 2
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section))
# next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
# next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
# curr_traj = next_section_traj
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
# curr_sec = 0
curr_traj = []
elif t_sec == 30 and curr_sec == 0:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3
diff_in_mid_section = 30 #3 - 2:30
# diff_in_next_section = abs(30 - t_sec) # 2:30 - 2
# 2 - 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section))
# next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
# next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
# curr_traj = next_section_traj
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
# curr_min -= 1
# curr_sec = 30
curr_traj = []
elif t_sec > curr_sec and curr_sec == 30:
# 2 sections off
diff_in_past_section = abs(prev_seconds - curr_sec)
diff_in_mid_section = 30
diff_in_next_section = abs(60 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:mid_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
# print("past_section_traj = ", past_section_traj)
# print("mid_section_traj = ", mid_section_traj)
# print("next_section_traj = ", next_section_traj)
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
elif t_sec == 0 and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - 0)
diff_in_mid_section = 30
diff_in_next_section = abs(60 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
next_section_traj = []
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
elif t_sec < curr_sec and curr_sec == 30:
# 3 sections off, 2:30 --> 1:14, 2:40-2:30, 2:30-2, 2-1:30, 1:30-1:14
# print("PROBLEMMM")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
diff_in_past_section = abs(prev_seconds - 30)
diff_in_mid_1_section = 30
diff_in_mid_2_section = 30
diff_in_next_section = abs(30 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
# elif t_sec <= curr_sec and curr_sec == 0:
# # 3 sections off, 2:30 --> 1:14, 2:40-2:30, 2:30-2, 2-1:30, 1:30-1:14
# print("PROBLEMMM 2")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
elif t_min == curr_min-2:
### 4:30 --> 2:30, 4:00 --> 2:00, 4:00 --> 2:30
# if t_sec > curr_sec and curr_sec == 0:
# print("MAJOR PROBLEM 2: ", t)
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
if t_sec == 30 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3
diff_in_mid_1_section = 30 # 3-2:30
diff_in_mid_2_section = 30 # 2:30- 2
diff_in_next_section = abs(30 - t_sec) #2 - 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_1_section = 30 # 3:30-3
diff_in_mid_2_section = 30 # 3- 2:30
diff_in_next_section = abs(30 - t_sec) #2:30 - 2
# 2-1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec == 30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_1_section = 30 # 3:30-3
diff_in_mid_2_section = 30 # 3- 2:30
diff_in_mid_3_section = 30 # 2:30 - 2
diff_in_next_section = abs(30 - t_sec) #2 - 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_3_idx = mid_section_2_idx + 1+ int(diff_in_mid_3_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
next_section_idx = mid_section_3_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
mid_section_3_traj = curr_t_traj[mid_section_2_idx+1:mid_section_3_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_3_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 0:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3:00
diff_in_mid_1_section = 30 # 3:0-2:30
diff_in_mid_2_section = 30 # 2:30- 2
diff_in_mid_3_section = 30 # 2 - 1:30
diff_in_next_section = abs(30 - t_sec) #1:30 - 1
# 1 - 0:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_3_idx = mid_section_2_idx + 1+ int(diff_in_mid_3_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
next_section_idx = mid_section_3_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
mid_section_3_traj = curr_t_traj[mid_section_2_idx+1:mid_section_3_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_3_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
else:
# print("MAJOR PROBLEM: ", t)
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
to_include = False
prev_min = t_min
prev_seconds = t_sec
stop_indices = np.where(times == 'stop')[0]
if len(stop_indices) > 0:
stop_idx = stop_indices[0]
t = str(times[start_idx])
traj_t = str(traj[start_idx])
msgs_t = str(sent_msgs[start_idx])
vic_t = str(victims[start_idx])
if traj_t != 'nan':
curr_traj.extend([eval(x) for x in traj_t.split(';')])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
return minute_to_traj, to_include
def compute_minute_to_msgs(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_msgs = {}
curr_min = 5
curr_sec = 0
# minute_to_msgs[(curr_min, curr_sec)] = []
# minute_to_victims[(curr_min, curr_sec)] = []
prev_time = (5,0)
for i in range(len(traj)):
t = str(times[i])
msgs_t = str(sent_msgs[i])
if ':' in t:
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
prev_time = (t_min, t_sec)
if msgs_t == 'nan':
continue
prev_t_min = prev_time[0]
prev_t_sec = prev_time[1]
window_sec = 0
window_min = prev_t_min
if prev_t_sec > 30:
window_sec = 30
# print((prev_t_min, prev_t_sec), 't = ', (window_min, window_sec))
# print("msgs_t: ", msgs_t)
# print()
keyname = (window_min, window_sec)
if keyname not in minute_to_msgs:
minute_to_msgs[keyname] = []
minute_to_msgs[keyname].append(msgs_t)
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
return minute_to_msgs
def compute_minute_to_victims(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_victims = {}
curr_min = 5
curr_sec = 0
# minute_to_msgs[(curr_min, curr_sec)] = []
minute_to_victims[(curr_min, curr_sec)] = []
prev_time = (5,0)
for i in range(len(traj)):
t = str(times[i])
victims_t = str(victims[i])
if ':' in t:
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
prev_time = (t_min, t_sec)
if victims_t in ['nan', 'door']:
continue
prev_t_min = prev_time[0]
prev_t_sec = prev_time[1]
window_sec = 0
window_min = prev_t_min
if prev_t_sec > 30:
window_sec = 30
# print((prev_t_min, prev_t_sec), 't = ', (window_min, window_sec))
# print("victims_t: ", victims_t)
# print()
keyname = (window_min, window_sec)
if keyname not in minute_to_victims:
minute_to_victims[keyname] = []
minute_to_victims[keyname].append(victims_t)
return minute_to_victims
```
## Get Data Dictionaries
```
uid_to_minute_victims = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['created_at'], ascending=False)
minute_to_victims = compute_minute_to_victims(df_traj)
# if to_include:
uid_to_minute_victims[p_uid] = minute_to_victims
uid_to_minute_msgs = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['created_at'], ascending=False)
minute_to_msgs = compute_minute_to_msgs(df_traj)
# if to_include:
uid_to_minute_msgs[p_uid] = minute_to_msgs
uid_to_minute_traj = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['time_spent'], ascending=False)
minute_to_traj, to_include = compute_minute_to_traj(df_traj)
if to_include:
uid_to_minute_traj[p_uid] = minute_to_traj
```
## Process Data Into Metrics
```
def l2(x1, x2):
return np.sqrt((x1[0] - x2[0])**2 + (x1[1] - x2[1])**2)
def compute_effort(traj_list, robot_list):
eff = 0
for i in range(len(traj_list)-1):
eff += l2(traj_list[i], traj_list[i+1])
eff_robot = 0
for i in range(len(robot_list)-1):
eff_robot += l2(robot_list[i], robot_list[i+1])
eff = eff + eff_robot
return eff
uid_min_to_data = {}
for puid in uid_to_minute_traj:
uid_min_to_data[puid] = {}
human_traj = uid_to_minute_traj[puid]
robot_traj = min_to_robot_traj
human_msgs = uid_to_minute_msgs[puid]
human_victims = uid_to_minute_victims[puid]
counter = 0
for minute in [4,3,2, 1, 0]:
for second in [30, 0]:
keyname = (minute, second)
uid_min_to_data[puid][counter] = {}
if keyname not in human_traj:
curr_human_traj = []
else:
curr_human_traj = human_traj[keyname]
curr_robot_traj = robot_traj[keyname]
effort = compute_effort(curr_human_traj, curr_robot_traj)
uid_min_to_data[puid][counter]['effort'] = effort
if keyname not in human_msgs:
num_msgs = 0
else:
num_msgs = len(human_msgs[keyname])
uid_min_to_data[puid][counter]['msgs'] = num_msgs
if keyname not in human_victims:
num_victims = 0
else:
num_victims = len(human_victims[keyname])
uid_min_to_data[puid][counter]['victims'] = num_victims
counter += 1
uid_min_to_data
```
# Split Into Groups
```
data_by_group = {1:{}, 2:{}, 3:{}, 4:{}}
for p_uid in uid_min_to_data:
# print(p_uid)
# print(uid_min_to_data[puid])
group_no = uid_to_group[p_uid]
data_by_group[group_no][p_uid] = uid_min_to_data[p_uid]
# break
```
# Binarize Data
```
group_to_means = {1:{}, 2:{}, 3:{}, 4:{}}
for group in data_by_group:
group_data = data_by_group[group]
all_effort = {i:[] for i in range(10)}
all_victims = {i:[] for i in range(10)}
all_msgs = {i:[] for i in range(10)}
for p_uid in group_data:
for i in range(10):
all_effort[i].append(group_data[p_uid][i]['effort'])
all_msgs[i].append(group_data[p_uid][i]['msgs'])
all_victims[i].append(group_data[p_uid][i]['victims'])
# print(all_victims)
final_effort = {i:np.mean(all_effort[i]) for i in range(10)}
final_victims = {i:np.mean(all_victims[i]) for i in range(10)}
final_msgs = {i:np.mean(all_msgs[i]) for i in range(10)}
group_to_means[group]['effort'] = final_effort
group_to_means[group]['victims'] = final_victims
group_to_means[group]['msgs'] = final_msgs
group_to_binary_data = {}
for group in data_by_group:
group_data = data_by_group[group]
mean_effort = group_to_means[group]['effort']
mean_victims = group_to_means[group]['victims']
mean_msgs = group_to_means[group]['msgs']
new_group_data = {}
for p_uid in group_data:
p_uid_data = []
for i in range(10):
effort_binary = 0 if group_data[p_uid][i]['effort'] < mean_effort[i] else 1
msgs_binary = 0 if group_data[p_uid][i]['msgs'] < mean_msgs[i] else 1
victims_binary = 0 if group_data[p_uid][i]['victims'] < mean_victims[i] else 1
p_uid_data.append((effort_binary, msgs_binary, victims_binary))
new_group_data[p_uid] = p_uid_data
group_to_binary_data[group] = new_group_data
group_to_binary_state_data = {}
group_to_state_list = {}
for group in data_by_group:
group_data = data_by_group[group]
group_to_state_list[group] = []
mean_effort = group_to_means[group]['effort']
mean_victims = group_to_means[group]['victims']
mean_msgs = group_to_means[group]['msgs']
new_group_data = {}
for p_uid in group_data:
p_uid_data = []
for i in range(9):
effort_binary = 0 if group_data[p_uid][i]['effort'] < mean_effort[i] else 1
msgs_binary = 0 if group_data[p_uid][i]['msgs'] < mean_msgs[i] else 1
victims_binary = 0 if group_data[p_uid][i]['victims'] < mean_victims[i] else 1
effort_binary_next = 0 if group_data[p_uid][i+1]['effort'] < mean_effort[i+1] else 1
msgs_binary_next = 0 if group_data[p_uid][i+1]['msgs'] < mean_msgs[i+1] else 1
victims_binary_next = 0 if group_data[p_uid][i+1]['victims'] < mean_victims[i+1] else 1
state_vector = (effort_binary_next, msgs_binary_next, victims_binary_next, effort_binary, msgs_binary, victims_binary)
p_uid_data.append(state_vector)
if state_vector not in group_to_state_list[group]:
group_to_state_list[group].append(state_vector)
new_group_data[p_uid] = p_uid_data
group_to_binary_state_data[group] = new_group_data
group_to_state_mapping = {}
for group in group_to_state_list:
group_to_state_mapping[group]= {}
state_id_to_state = dict(enumerate(group_to_state_list[group]))
state_to_state_id = {v: k for k, v in state_id_to_state.items()}
group_to_state_mapping[group]['id_to_vec'] = state_id_to_state
group_to_state_mapping[group]['vec_to_id'] = state_to_state_id
```
# Generate States
```
group_to_state_id_data_w_pid = {}
group_to_state_id_data = {}
for group in group_to_binary_state_data:
group_data = group_to_binary_state_data[group]
state_to_state_id = group_to_state_mapping[group]['vec_to_id']
all_data = []
all_data_w_pid = {}
for p_uid in group_data:
state_data = [state_to_state_id[elem] for elem in group_data[p_uid]]
all_data.append(state_data)
all_data_w_pid[p_uid] = state_data
group_to_state_id_data[group] = all_data
group_to_state_id_data_w_pid[group] = all_data_w_pid
group_to_state_id_data
import pickle
with open('minimap_group_to_state_data_AGG.pickle', 'wb') as handle:
pickle.dump(group_to_state_id_data, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_state_data_w_pid_AGG.pickle', 'wb') as handle:
pickle.dump(group_to_state_id_data_w_pid, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_binary_state_data_AGG.pickle', 'wb') as handle:
pickle.dump(group_to_binary_state_data, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_state_mapping_AGG.pickle', 'wb') as handle:
pickle.dump(group_to_state_mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
```
| github_jupyter |
```
import sys,os
sys.path.append('.')
sys.path.append('/home/lev/slot_attention/')
import torch
from typing import Optional
import pytorch_lightning.loggers as pl_loggers
from pytorch_lightning import Trainer
from pytorch_lightning.callbacks import LearningRateMonitor
from torchvision import transforms
from slot_attention.data import CLEVRDataModule, ESC50DataModule, SyntaticDataModule
from slot_attention.method import SlotAttentionMethod
from slot_attention.model import SlotAttentionModel
from slot_attention.params import SlotAttentionParams
from slot_attention.utils import ImageLogCallback
from slot_attention.utils import rescale, normalize_audio
import hydra
import wandb
from omegaconf import DictConfig, OmegaConf
import logging
import matplotlib.pyplot as plt
import seaborn as sns
import os
os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"]="4"
def get_dataset(cfg:DictConfig):
if "_dataset" in cfg.dataset.name:
return SyntaticDataModule(cfg)
elif cfg.dataset.name == 'esc50':
clevr_transforms = transforms.Compose(
[
transforms.Lambda(normalize_audio)
])
return ESC50DataModule(cfg=cfg, clevr_transforms=clevr_transforms)
elif cfg.dataset.name == 'clever':
clevr_transforms = transforms.Compose(
[
transforms.ToTensor(),
transforms.Lambda(rescale), # rescale between -1 and 1
transforms.Resize(tuple(cfg.model.resolution)),
]
)
clevr_datamodule = CLEVRDataModule(
data_root=cfg.dataset.data_root,
max_n_objects=cfg.model.num_slots - 1,
train_batch_size=cfg.dataset.train_batch_size,
val_batch_size=cfg.dataset.val_batch_size,
clevr_transforms=clevr_transforms, # change also this moment))
num_train_images=cfg.dataset.num_train_images,
num_val_images=cfg.dataset.num_val_images,
num_workers=cfg.dataset.num_workers,
)
return clevr_datamodule
else:
print('Choose the dataset')
def main(cfg):
assert cfg.model.num_slots > 1, "Must have at least 2 slots."
if cfg.additional.is_verbose:
print(f"INFO: limiting the dataset to only images with `num_slots - 1` ({cfg.model.num_slots - 1}) objects.")
if cfg.dataset.num_train_images:
print(f"INFO: restricting the train dataset size to `num_train_images`: {cfg.dataset.num_train_images}")
if cfg.dataset.num_val_images:
print(f"INFO: restricting the validation dataset size to `num_val_images`: {cfg.dataset.num_val_images}")
clevr_datamodule = get_dataset(cfg)
print(f"Training set size (images must have {cfg.model.num_slots - 1} objects):", len(clevr_datamodule.train_dataset))
model = SlotAttentionModel(
cfg=cfg
)
method = SlotAttentionMethod(model=model, datamodule=clevr_datamodule, hparams=cfg)
return method, clevr_datamodule
# from hydra import compose, initialize
# from omegaconf import OmegaConf
# with initialize(config_path="../buffer/configs_benchmark", job_name="test_app"):
# cfg = compose(config_name="default", overrides=[]) #clever_benchmark
# method, clevr_datamodule = main(cfg)
# print(OmegaConf.to_yaml(cfg))
checkpoint_path="/home/lev/slot_attention/outputs/2021-12-01/clever_model_100epoch/slot-attention-search/1ci43d71/checkpoints/epoch=99-step=27299.ckpt"
data = torch.load(checkpoint_path)
method.load_state_dict(data['state_dict'])
method.eval()
print('1')
for batch in clevr_datamodule.val_dataset:
batch = batch.unsqueeze(0)
break
recon_combined, recons, masks, slots = method(batch)
from slot_attention.utils import to_rgb_from_tensor
from torchvision import utils as vutils
# combine images in a nice way so we can display all outputs in one grid, output rescaled to be between 0 and 1
out = to_rgb_from_tensor(
torch.cat(
[
batch.unsqueeze(1), # original images
recon_combined.unsqueeze(1), # reconstructions
recons * masks + (1 - masks), # each slot
],
dim=1,
)
)
batch_size, num_slots, C, H, W = recons.shape
images = vutils.make_grid(
out.view(batch_size * out.shape[1], C, H, W).cpu(), normalize=False, nrow=out.shape[1],
)
ch, h, w = images.shape
plt.imshow(images.reshape(h, w, ch))
```
### Trained model
```
from collections import defaultdict
for batch in clevr_datamodule.val_dataloader():
#batch = batch.unsqueeze(0)
enc_out = method.model.encoder(batch)
for num_iter in range(1,100,5):
method.model.slot_attention.num_iterations = num_iter
a = method.model.slot_attention(enc_out)
d[num_iter].append(torch.mean(torch.cdist(a,a)).detach().cpu().numpy())
stat = defaultdict(list)
for key in d.keys():
stat[key] = (np.mean(d[key]), np.std(d[key]))
plt.plot([stat[key][0] for key in stat.keys()])
stat
```
### Random initialization
```
# random weights
method, clevr_datamodule = main(cfg)
method.to(torch.device('cuda'))
d = defaultdict(list)
for batch in clevr_datamodule.val_dataloader():
#batch = batch.unsqueeze(0)
enc_out = method.model.encoder(batch.to(torch.device('cuda')))
for num_iter in [1,2,3,5,10,15,25,30,100]:
method.model.slot_attention.num_iterations = num_iter
a = method.model.slot_attention(enc_out)
d[num_iter].append(torch.mean(torch.cdist(a,a)).detach().cpu().numpy())
stat = defaultdict(list)
for key in d.keys():
stat[key] = (np.mean(d[key]), np.std(d[key]))
plt.plot([stat[key][0] for key in stat.keys()])
stat
```
### Pretrained encoder + Slot random weights
```
# random weights
method1, clevr_datamodule = main(cfg)
method1.to(torch.device('cuda')).eval()
# pretrained encoder
method2, clevr_datamodule = main(cfg)
checkpoint_path="/home/lev/slot_attention/outputs/2021-12-01/clever_model_100epoch/slot-attention-search/1ci43d71/checkpoints/epoch=99-step=27299.ckpt"
data = torch.load(checkpoint_path)
method2.load_state_dict(data['state_dict'])
method2.to(torch.device('cuda')).eval()
print('1')
method.to(torch.device('cuda'))
d = defaultdict(list)
for batch in clevr_datamodule.val_dataloader():
#batch = batch.unsqueeze(0)
enc_out = method2.model.encoder(batch.to(torch.device('cuda')))
for num_iter in [1,2,3,5,10,15,25,30,100]:
method1.model.slot_attention.num_iterations = num_iter
a = method1.model.slot_attention(enc_out)
d[num_iter].append(torch.mean(torch.cdist(a,a)).detach().cpu().numpy())
stat = defaultdict(list)
for key in d.keys():
stat[key] = (np.mean(d[key]), np.std(d[key]))
plt.plot([stat[key][0] for key in stat.keys()])
```
### Slot retrained + Encoder random weights
```
# random weights
method1, clevr_datamodule = main(cfg)
method1.to(torch.device('cuda')).eval()
# pretrained encoder
method2, clevr_datamodule = main(cfg)
checkpoint_path="/home/lev/slot_attention/outputs/2021-12-01/clever_model_100epoch/slot-attention-search/1ci43d71/checkpoints/epoch=99-step=27299.ckpt"
data = torch.load(checkpoint_path)
method2.load_state_dict(data['state_dict'])
method2.to(torch.device('cuda')).eval()
print('1')
method.to(torch.device('cuda'))
d = defaultdict(list)
for batch in clevr_datamodule.val_dataloader():
#batch = batch.unsqueeze(0)
enc_out = method1.model.encoder(batch.to(torch.device('cuda')))
for num_iter in [1,2,3,5,10,15,25,30,100]:
method2.model.slot_attention.num_iterations = num_iter
a = method2.model.slot_attention(enc_out)
d[num_iter].append(torch.mean(torch.cdist(a,a)).detach().cpu().numpy())
stat = defaultdict(list)
for key in d.keys():
stat[key] = (np.mean(d[key]), np.std(d[key]))
plt.plot([stat[key][0] for key in stat.keys()])
from hydra import compose, initialize
from omegaconf import OmegaConf
with initialize(config_path="../slot_attention/configs", job_name="test_app"):
cfg = compose(config_name="default", overrides=[])
method, clevr_datamodule = main(cfg)
checkpoint_path="/home/lev/slot_attention/outputs/2021-12-09/20-00-00/implicit_experiments/2619mscn/checkpoints/epoch=99-step=27298.ckpt"
data = torch.load(checkpoint_path)
method.load_state_dict(data['state_dict'])
method.to(torch.device('cuda')).eval()
print('1')
#
from slot_attention.utils import to_rgb_from_tensor
from torchvision import utils as vutils
for i, batch in enumerate(clevr_datamodule.val_dataset):
if i == 2:
batch = batch.unsqueeze(0).to(torch.device('cuda'))
break
recon_combined, recons, masks, slots = method(batch)
# combine images in a nice way so we can display all outputs in one grid, output rescaled to be between 0 and 1
# combine images in a nice way so we can display all outputs in one grid, output rescaled to be between 0 and 1
out = to_rgb_from_tensor(
torch.cat(
[
batch.unsqueeze(1), # original images
recon_combined.unsqueeze(1), # reconstructions
recons * masks + (1 - masks), # each slot
],
dim=1,
)
)
batch_size, num_slots, C, H, W = recons.shape
images = vutils.make_grid(
out.view(batch_size * out.shape[1], C, H, W).cpu(), normalize=False, nrow=out.shape[1],
)
ch, h, w = images.shape
plt.imshow(images.reshape(h, w, ch))
data = masks.squeeze(0).cpu().detach().numpy()
for idx in range(len(data)):
plt.imshow(data[idx,0], cmap='hot')
ax = sns.heatmap(data[idx,0])
plt.show()
batch.shape
plt.imshow(data[idx,0])
for batch in clevr_datamodule.val_dataset:
batch = batch.unsqueeze(0).to(torch.device('cuda'))
break
recon_combined, recons, masks, slots = method(batch)
from slot_attention.utils import to_rgb_from_tensor
from torchvision import utils as vutils
# combine images in a nice way so we can display all outputs in one grid, output rescaled to be between 0 and 1
out = to_rgb_from_tensor(
torch.cat(
[
batch.unsqueeze(1), # original images
recon_combined.unsqueeze(1), # reconstructions
recons * masks + (1 - masks), # each slot
],
dim=1,
)
)
batch_size, num_slots, C, H, W = recons.shape
images = vutils.make_grid(
out.view(batch_size * out.shape[1], C, H, W).cpu(), normalize=False, nrow=out.shape[1],
)
ch, h, w = images.shape
plt.imshow(images.reshape(h, w, ch))
import seaborn as sns
data = 1 - masks.squeeze(0).cpu().detach().numpy()
for idx in range(len(data)):
plt.imshow(data[idx,0], cmap='hot')
ax = sns.heatmap(data[idx,0])
plt.show()
```
| github_jupyter |
```
import numpy as np
import random
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
import torch.optim.lr_scheduler as sched
import torch.utils.data as data
import util
from args import get_train_args
from collections import OrderedDict
from json import dumps
from models import BiDAF
#from tensorboardX import SummaryWriter
from torch.utils.tensorboard import SummaryWriter
from tqdm import tqdm
from ujson import load as json_load
from util import collate_fn, SQuAD
from argparse import Namespace
argDict = {'train_record_file': './data/train.npz',
'dev_record_file': './data/dev.npz',
'test_record_file': './data/test.npz',
'word_emb_file': './data/word_emb.json',
'char_emb_file': './data/char_emb.json',
'train_eval_file': './data/train_eval.json',
'dev_eval_file': './data/dev_eval.json',
'test_eval_file': './data/test_eval.json',
'name': 'devNonPCE',
'max_ans_len': 15,
'num_workers': 4,
'save_dir': './save/',
'batch_size': 16,
'use_squad_v2': True,
'hidden_size': 100,
'num_visuals': 10,
'load_path': None,
'rnn_type': 'LSTM',
'char_embeddings': False,
'eval_steps': 5000,
'lr': 0.5, 'l2_wd': 0,
'num_epochs': 3,
'drop_prob': 0.2,
'metric_name': 'F1',
'max_checkpoints': 5,
'max_grad_norm': 5.0,
'seed': 224,
'ema_decay': 0.999,
'char_out_channels': 5,
'char_kernel_size': 100,
'maximize_metric': True,
'gpu_ids': []}
args = Namespace(**argDict)
# Set up logging and devices
args.save_dir = util.get_save_dir(args.save_dir, args.name, training=True)
log = util.get_logger(args.save_dir, args.name)
tbx = SummaryWriter(args.save_dir)
#if args.device_cpu:
# device = 'cpu',
# args.gpu_ids = []
#else:
device, args.gpu_ids = util.get_available_devices()
log.info(f'Args: {dumps(vars(args), indent=4, sort_keys=True)}')
args.batch_size *= max(1, len(args.gpu_ids))
torch.cuda.is_available()
print(args)
# Set random seed
log.info(f'Using random seed {args.seed}...')
random.seed(args.seed)
np.random.seed(args.seed)
torch.manual_seed(args.seed)
torch.cuda.manual_seed_all(args.seed)
# Get embeddings
log.info('Loading embeddings...')
word_vectors = util.torch_from_json(args.word_emb_file)
char_vectors = util.torch_from_json(args.char_emb_file)
# Get model
log.info('Building model...')
model = BiDAF(word_vectors = word_vectors,
char_vectors = char_vectors,
hidden_size=args.hidden_size,
rnn_type=args.rnn_type,
drop_prob=args.drop_prob)
#if args.device_cpu:
# args.gpu_ids = []
# device = 'cpu'
#else:
#model = nn.DataParallel(model, args.gpu_ids)
if args.load_path:
log.info(f'Loading checkpoint from {args.load_path}...')
model, step = util.load_model(model, args.load_path, args.gpu_ids)
else:
step = 0
model = model.to(device)
model.train()
ema = util.EMA(model, args.ema_decay)
print(model)
device
# Get saver
saver = util.CheckpointSaver(args.save_dir,
max_checkpoints=args.max_checkpoints,
metric_name=args.metric_name,
maximize_metric=args.maximize_metric,
log=log)
# Get optimizer and scheduler
optimizer = optim.Adadelta(model.parameters(), args.lr,
weight_decay=args.l2_wd)
scheduler = sched.LambdaLR(optimizer, lambda s: 1.) # Constant LR
# Get data loader
log.info('Building dataset...')
train_dataset = SQuAD(args.train_record_file, args.use_squad_v2)
train_loader = data.DataLoader(train_dataset,
batch_size=args.batch_size,
shuffle=True,
num_workers=args.num_workers,
collate_fn=collate_fn)
dev_dataset = SQuAD(args.dev_record_file, args.use_squad_v2)
dev_loader = data.DataLoader(dev_dataset,
batch_size=args.batch_size,
shuffle=False,
num_workers=args.num_workers,
collate_fn=collate_fn)
print(train_loader.__dir__())
print(train_loader.batch_sampler)
# get a training input
cw_idxs, cc_idxs, qw_idxs, qc_idxs, y1, y2, ids = next(iter(train_loader))
cw_idxs=cw_idxs.to(device)
cc_idxs=cc_idxs.to(device)
qw_idxs=qw_idxs.to(device)
qc_idxs=qc_idxs.to(device)
print('cw_idxs',cw_idxs.shape,cw_idxs.device)
print('cc_idxs',cc_idxs.shape)
print('qw_idxs',qw_idxs.shape)
print('qc_idxs',qc_idxs.shape)
tbx.add_graph(model,[cw_idxs, cc_idxs, qw_idxs, qc_idxs])
# Train
log.info('Training...')
steps_till_eval = args.eval_steps
epoch = step // len(train_dataset)
while epoch != args.num_epochs:
epoch += 1
log.info(f'Starting epoch {epoch}...')
with torch.enable_grad(), \
tqdm(total=len(train_loader.dataset)) as progress_bar:
for cw_idxs, cc_idxs, qw_idxs, qc_idxs, y1, y2, ids in train_loader:
# Setup for forward
cw_idxs = cw_idxs.to(device)
qw_idxs = qw_idxs.to(device)
cc_idxs = cc_idxs.to(device)
qc_idxs = qc_idxs.to(device)
batch_size = cw_idxs.size(0)
optimizer.zero_grad()
# Forward
log_p1, log_p2 = model(cw_idxs,cc_idxs, qw_idxs, qc_idxs)
y1, y2 = y1.to(device), y2.to(device)
loss = F.nll_loss(log_p1, y1) + F.nll_loss(log_p2, y2)
loss_val = loss.item()
# Backward
loss.backward()
nn.utils.clip_grad_norm_(model.parameters(), args.max_grad_norm)
optimizer.step()
scheduler.step() # removed parm step // batch_size per scheduler 1.8 release notes
ema(model, step // batch_size)
# Log info
step += batch_size
progress_bar.update(batch_size)
progress_bar.set_postfix(epoch=epoch,
NLL=loss_val)
tbx.add_scalar('train/NLL', loss_val, step)
tbx.add_scalar('train/LR',
optimizer.param_groups[0]['lr'],
step)
steps_till_eval -= batch_size
if steps_till_eval <= 0:
steps_till_eval = args.eval_steps
# Evaluate and save checkpoint
log.info(f'Evaluating at step {step}...')
ema.assign(model)
results, pred_dict = evaluate(model, dev_loader, device,
args.dev_eval_file,
args.max_ans_len,
args.use_squad_v2)
saver.save(step, model, results[args.metric_name], device)
ema.resume(model)
# Log to console
results_str = ', '.join(f'{k}: {v:05.2f}' for k, v in results.items())
log.info(f'Dev {results_str}')
# Log to TensorBoard
log.info('Visualizing in TensorBoard...')
for k, v in results.items():
tbx.add_scalar(f'dev/{k}', v, step)
util.visualize(tbx,
pred_dict=pred_dict,
eval_path=args.dev_eval_file,
step=step,
split='dev',
num_visuals=args.num_visuals)
hparms = args.__dict__.copy()
hparms['gpu_ids']=str(args.gpu_ids)
metrics = dict([ ('met/'+k,v) for (k,v) in results.items()])
tbx.add_hparams(hparms,metrics)
```
| github_jupyter |
# Import Packages
[LaTeX Editor](https://latex.codecogs.com/eqneditor/editor.php)
```
# Run cell ini dulu!!!
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
def showGraph(result, range):
x = np.arange(result-range, result+range, 0.01)
y = f(x)
y1 = 0
plt.clf()
plt.plot(x, y)
plt.plot(x, 0/x)
plt.grid()
plt.show()
a = 2.023624234234
print(round(a, 3))
```
# Bisection Method
$x_{mid} = \frac{x_{low}+x_{up}}{2}$
$|\epsilon_{a}| = |\frac{x_{mid}^{new}-x_{mid}^{old}}{x_{mid}^{new}}| \cdot 100\%$
```
table = {
'xlow': [],
'xup': [],
'xmid': [],
'rae': [],
'f(xlow)': [],
'f(xmid)': []
}
# ====== Ganti f(x) ======
def f(x):
return x**2 + 4*x - 12
def bisection(xlow, xup, xmidold=9999, rae=9999, iter=0):
xmid = (xlow + xup)/2
rae = abs((xmid-xmidold)/xmid)
f_xlow = f(xlow)
f_xmid = f(xmid)
table['xlow'].append(xlow)
table['xup'].append(xup)
table['xmid'].append(xmid)
if iter > 0:
table['rae'].append(rae)
else:
table['rae'].append(999)
table['f(xlow)'].append(f_xlow)
table['f(xmid)'].append(f_xmid)
if iter > 100 or rae < es:
return xmid
if f_xlow * f_xmid < 0:
return bisection(xlow, xmid, xmid, rae, iter+1)
elif f_xlow * f_xmid > 0:
return bisection(xmid, xup, xmid, rae, iter+1)
return xmid
# ====== Ganti Titik Awal ======
xlow = 1
xup = 8
es = 0.005 # 0.5%
result = bisection(xlow, xup)
print(result, "\n")
# showGraph(result, 5)
# showGraph(result, 1)
print(pd.DataFrame(table).round(4))
```
# False Position Method
$x_{mid} = x_{up} - \frac{f(x_{up})(x_{low}-x_{up})}{f(x_{low})-f(x_{up})}$
$|\epsilon_{a}| = |\frac{x_{mid}^{new}-x_{mid}^{old}}{x_{mid}^{new}}| \cdot 100\%$
```
table = {
'xlow': [],
'xup': [],
'xmid': [],
'rae': [],
'f(xlow)': [],
'f(xmid)': []
}
# ====== Ganti f(x) ======
def f(x):
return x**2 + 4*x - 12
def falsePosition(xlow, xup, xmidold=9999, rae=9999, iter=0):
xmid = xup - (f(xup)*(xlow-xup))/(f(xlow)-f(xup))
rae = abs((xmid-xmidold)/xmid)
f_xlow = f(xlow)
f_xmid = f(xmid)
table['xlow'].append(xlow)
table['xup'].append(xup)
table['xmid'].append(xmid)
if iter > 0:
table['rae'].append(rae)
else:
table['rae'].append(999)
table['f(xlow)'].append(f_xlow)
table['f(xmid)'].append(f_xmid)
if iter > 100 or rae < es:
return xmid
if f_xlow * f_xmid < 0:
return falsePosition(xlow, xmid, xmid, rae, iter+1)
elif f_xlow * f_xmid > 0:
return falsePosition(xmid, xup, xmid, rae, iter+1)
return xmid
# ====== Ganti Titik Awal ======
xlow = 1
xup = 8
es = 0.005
result = falsePosition(xlow, xup)
print(result, "\n")
# showGraph(result, 5)
# showGraph(result, 1)
print(pd.DataFrame(table).round(4))
```
# Fixed Point Iteration
```
table = {
'xi': [],
'f(xi)': []
}
# ====== ganti f(x) ======
def f(x):
return 2*(x**3) - 11.7*(x**2) + 17.7*x - 5
# ====== ganti g(x)
def g(x):
return ((11.7*(x**2) - 17.7*x + 5)/2)**(1/3)
def fixedPointIteration(xi, iter=0):
f_xi = f(xi)
table['xi'].append(xi)
table['f(xi)'].append(f_xi)
xii = g(xi)
if iter < 100 and abs(f_xi) >= es:
return fixedPointIteration(xii, iter+1)
return xi
x0 = 3
result = fixedPointIteration(x0)
print(result, "\n")
print(pd.DataFrame(table))
```
# Newton Raphson Method
$x_{i+1} = x_{i} - \frac{f(x_{i})}{f'(x_{i})}$
$|\epsilon_{a}| = |\frac{x_{i+1}-x_{i}}{x_{i+1}}| \cdot 100\% $
```
table = {
'xi': [],
'xi+1': [],
'rae': [],
'f(xi)': [],
"f'(xi)": []
}
# ====== Ganti f(x) ======
def f(x):
return 9.34-21.97*x+16.3*(x**2)-3.704*(x**3)
# ====== Ganti f'(x) ======
def fa(x):
return -21.97 + 32.6*x - 3*3.704*(x**2)
def newtonRaphson(xi, rae=9999, iter=0):
xii = xi - f(xi)/fa(xi)
rae = abs((xii-xi)/xii)
table['xi'].append(xi)
table['xi+1'].append(xii)
table['rae'].append(rae)
table['f(xi)'].append(f(xi))
table["f'(xi)"].append(fa(xi))
if iter < 100 and rae >= es:
return newtonRaphson(xii, rae, iter+1)
return xii
# ====== Ganti Titik Awal ======
x0 = 0.5
es = 0.005
result = newtonRaphson(x0)
print(result, "\n")
# showGraph(result, 10)
# showGraph(result, 1)
print(pd.DataFrame(table).round(4))
```
# Secant Method
$x_{i+1} = x_{i} - \frac{f(x_{i})(x_{i-1}-x_{i})}{f(x_{i-1})-f(x_{i})}$
$|\epsilon_{a}| = |\frac{x_{i+1}-x_{i}}{x_{i+1}}| \cdot 100\% $
```
table = {
'x': [],
'xi': [],
'xi+1': [],
'rae': [],
"f(xi)": [],
"f(xi-1)": [],
"xi-1 - xi": [],
}
# ====== Ganti f(x) ======
def f(x):
return 9.34-21.97*x+16.3*(x**2)-3.704*(x**3)
def secant(x, xi, rae=9999, iter=0):
xii = xi - (f(xi)*(x-xi))/(f(x)-f(xi))
rae = abs((xii-xi)/xii)
table['x'].append(x)
table['xi'].append(xi)
table['xi+1'].append(xii)
table['rae'].append(rae)
table['f(xi-1)'].append(f(x))
table['f(xi)'].append(f(xi))
table['xi-1 - xi'].append(x - xi)
if iter < 100 and rae >= es:
return secant(xi, xii, rae, iter+1)
return xii
# ====== Ganti Titik Awal ======
x99 = 1.05
x0 = 0.5
es = 0.005
result = secant(x99, x0)
print(result, "\n")
# showGraph(result, 10)
# showGraph(result, 1)
print(pd.DataFrame(table).round(4))
```
# Modified Secant Method
```
table = {
'xi': [],
'xi+1': [],
'rae': [],
'dxi': [],
'xi + dxi': [],
'f(xi)': [],
'f(xi + dxi)': [],
}
# ====== Ganti f(x) ======
def f(x):
return 9.34-21.97*x+16.3*(x**2)-3.704*(x**3)
def modifiedSecant(xi, d, rae=9999, iter=0):
# print(f(xi+d*xi))
xii = xi - (d*xi*f(xi))/(f(xi+d*xi)-f(xi))
rae = abs((xii-xi)/xii)
table['xi'].append(xi)
table['xi+1'].append(xii)
table['rae'].append(rae)
table['dxi'].append(d*xi)
table['xi + dxi'].append(xi+d*xi)
table['f(xi)'].append(f(xi))
table['f(xi + dxi)'].append(f(xi + d*xi))
if iter < 100 and rae >= es:
return modifiedSecant(xii, d, rae, iter+1)
return xii
# ====== Ganti Titik Awal ======
x0 = 0.8
d = 0.01
es = 0.00005
result = modifiedSecant(x0, d)
print(result, "\n")
# showGraph(result, 10)
# showGraph(result, 1)
print(pd.DataFrame(table).round(4))
```
| github_jupyter |
# Deriving a vegetation index from PlanetScope imagery
Researchers often use a vegetation index called NDVI to measure the "greeenness" or density of vegetation across a landscape. In addition to monitoring vegetation health, NDVI (Normalized Difference Vegetation Index) can be used to track climate change, agricultural production, desertification, and land cover change. Developed by NASA scientist Compton Tucker in 1977, NDVI is derived from satellite imagery and compares reflected near-infrared light to reflected visible red light. It can be expressed as following equation:

In general, healthy and/or dense vegetation reflects a lot of near-infrared light and not as much red visible light. Conversely, when vegetation is sparse or not-so-healthy, its near-infrared reflectance decreases and its red light reflectance increases. You can read more about how NDVI is used to study cyclical, seasonal, and long-term changes to the Earth's physical characteristics from [NASA](https://earthobservatory.nasa.gov/Features/MeasuringVegetation/measuring_vegetation_1.php) and [USGS](https://phenology.cr.usgs.gov/ndvi_foundation.php) researchers.
**In this guide, you'll perform a basic NDVI calculation on PlanetScope imagery using just a few lines of Python. Here are the steps:**
1. Download a PlanetScope image
2. Extract data from the red and near-infrared bands
3. Normalize the data
4. Perform the NDVI calculation
5. Save the NDVI image
6. Apply a color scheme and visualize NDVI on a map
7. Generate a histogram to view NDVI values
### Requirements
- Python 2.7 or 3+
- [Planet's Python Client](https://www.planet.com/docs/api-quickstart-examples/cli/)
- [rasterio](https://github.com/mapbox/rasterio)
- [numpy](http://www.numpy.org/)
- [matplotlib](https://matplotlib.org/)
- [Planet API Key](https://www.planet.com/account/#/), stored as environment variable `$PL_API_KEY`.
- [Planet 4-Band Imagery](https://www.planet.com/docs/imagery-quickstart/) with the following specifications: `item-type`: `PSOrthoTile`, `REOrthoTile`, or `PSScene4Band`; `asset-type`: `analytic`, or `basic_analytic`
## Step 1. Download a PlanetScope image
First, you're going to download a [4-band PlanetScope satellite image](https://www.planet.com/docs/spec-sheets/sat-imagery/#ps-imagery-product) of agricultural land in California's Central Valley, captured in late August 2016 (`item-id`: `20160831_180302_0e26`). You can do this using [Planet's Python client](https://www.planet.com/docs/api-quickstart-examples/cli/) to interact with our Data API, or by browsing [Planet Explorer](https://www.planet.com/products/explorer/), filtering for 4 Band PlanetScope scene (`PSScene4Band`) or Planetscope ortho tile (`PSOrthoTile`), and downloading an `analytic` asset.
Before you download the full image, you can [preview a thumbnail](https://www.planet.com/docs/reference/data-api/previews/) of the image via Planet's Data API. (The thumbnails are 256x256 by default, and can be scaled up to 512x512 by passing in a `width` parameter.)
```
from IPython.display import Image
Image(url="https://api.planet.com/data/v1/item-types/PSScene4Band/items/20160831_180302_0e26/thumb?width=512")
```
Next, you'll use [Planet's Python client](https://planetlabs.github.io/planet-client-python/index.html) to download the image. *Note: when you run this command, you'll get a stream of messages in your Jupyter notebook as the Python client polls the Data API to determine if the image is [activated and ready to download](https://www.planet.com/docs/api-quickstart-examples/step-2-download/#activate).*
```
!planet data download --item-type PSScene4Band --dest data --asset-type analytic,analytic_xml --string-in id 20160831_180302_0e26
```
**Congratulations!** You now have two files in your `data` directory: `20160831_180302_0e26_3B_AnalyticMS.tif` and `20160831_180302_0e26_3B_AnalyticMS_metadata.xml`. The first file is a GeoTIFF, the image you requested with spatial reference data embedded. The second file is a metadata file for that image that includes the data you'll need to calculate the NDVI.
## Step 2. Extract the data from the red and near-infrared bands
In this step, you'll use [Rasterio](https://github.com/mapbox/rasterio), a Python library for reading and writing geospatial raster datasets, to open the raster image you downloaded (the .tif file). Then you'll extract the data from the red and near-infrared bands and load the band data into arrays that you can manipulate using Python's [NumPy](http://www.numpy.org/) libary. *Note: in PlanetScope 4-band images, the band order is BGRN: (1) Blue, (2) Green, (3) Red, (4) Near-infrared.*
```
import rasterio
import numpy as np
filename = "data/20160831_180302_0e26_3B_AnalyticMS.tif"
# Load red and NIR bands - note all PlanetScope 4-band images have band order BGRN
with rasterio.open(filename) as src:
band_red = src.read(3)
with rasterio.open(filename) as src:
band_nir = src.read(4)
```
## Step 3. Normalize the band data
Before you can calculate NDVI, you must normalize the values in the arrays for each band using the [Top of Atmosphere (TOA) reflectance coefficients](https://landsat.usgs.gov/using-usgs-landsat-8-product) stored in metadata file you downloaded (the .xml file).
```
from xml.dom import minidom
xmldoc = minidom.parse("data/20160831_180302_0e26_3B_AnalyticMS_metadata.xml")
nodes = xmldoc.getElementsByTagName("ps:bandSpecificMetadata")
# XML parser refers to bands by numbers 1-4
coeffs = {}
for node in nodes:
bn = node.getElementsByTagName("ps:bandNumber")[0].firstChild.data
if bn in ['1', '2', '3', '4']:
i = int(bn)
value = node.getElementsByTagName("ps:reflectanceCoefficient")[0].firstChild.data
coeffs[i] = float(value)
# Multiply the Digital Number (DN) values in each band by the TOA reflectance coefficients
band_red = band_red * coeffs[3]
band_nir = band_nir * coeffs[4]
```
## Step 4. Perform the NDVI calculation
Next, you're going to calculate NDVI through subtraction and division of the normalized values stored in the NumPy arrays. This calculation will give you NDVI values that range from -1 to 1. Values closer to 1 indicate a greater density of vegetation or higher level of "greenness."
```
# Allow division by zero
np.seterr(divide='ignore', invalid='ignore')
# Calculate NDVI. This is the equation at the top of this guide expressed in code
ndvi = (band_nir.astype(float) - band_red.astype(float)) / (band_nir + band_red)
# check range NDVI values, excluding NaN
np.nanmin(ndvi), np.nanmax(ndvi)
```
## Step 5. Save the NDVI image
Next, you're going to save the calculated NDVI values to a new image file, making sure the new image file has the same geospatial metadata as the original GeoTIFF we downloaded.
```
# Set spatial characteristics of the output object to mirror the input
kwargs = src.meta
kwargs.update(
dtype=rasterio.float32,
count = 1)
# Write band calculations to a new raster file
with rasterio.open('output/ndvi.tif', 'w', **kwargs) as dst:
dst.write_band(1, ndvi.astype(rasterio.float32))
```
## Step 6. Apply a color scheme to visualize the NDVI values on the image
In the last two steps, you'll use [Matplotlib](https://matplotlib.org/) to visualize the NDVI values you calculated for the PlanetScope scene. First you'll view a map of the NDVI values; then you'll generate a histogram of NDVI values.
```
import matplotlib.pyplot as plt
import matplotlib.colors as colors
"""
The NDVI values will range from -1 to 1. You want to use a diverging color scheme to visualize the data,
and you want to center the colorbar at a defined midpoint. The class below allows you to normalize the colorbar.
"""
class MidpointNormalize(colors.Normalize):
"""
Normalise the colorbar so that diverging bars work there way either side from a prescribed midpoint value)
e.g. im=ax1.imshow(array, norm=MidpointNormalize(midpoint=0.,vmin=-100, vmax=100))
Credit: Joe Kington, http://chris35wills.github.io/matplotlib_diverging_colorbar/
"""
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
self.midpoint = midpoint
colors.Normalize.__init__(self, vmin, vmax, clip)
def __call__(self, value, clip=None):
# I'm ignoring masked values and all kinds of edge cases to make a
# simple example...
x, y = [self.vmin, self.midpoint, self.vmax], [0, 0.5, 1]
return np.ma.masked_array(np.interp(value, x, y), np.isnan(value))
# Set min/max values from NDVI range for image (excluding NAN)
# set midpoint according to how NDVI is interpreted: https://earthobservatory.nasa.gov/Features/MeasuringVegetation/
min=np.nanmin(ndvi)
max=np.nanmax(ndvi)
mid=0.1
fig = plt.figure(figsize=(20,10))
ax = fig.add_subplot(111)
# diverging color scheme chosen from https://matplotlib.org/users/colormaps.html
cmap = plt.cm.RdYlGn
cax = ax.imshow(ndvi, cmap=cmap, clim=(min, max), norm=MidpointNormalize(midpoint=mid,vmin=min, vmax=max))
ax.axis('off')
ax.set_title('Normalized Difference Vegetation Index', fontsize=18, fontweight='bold')
cbar = fig.colorbar(cax, orientation='horizontal', shrink=0.65)
fig.savefig("output/ndvi-fig.png", dpi=200, bbox_inches='tight', pad_inches=0.7)
plt.show()
```
## 7. Generate a histogram of NDVI values
```
fig2 = plt.figure(figsize=(10,10))
ax = fig2.add_subplot(111)
plt.title("NDVI Histogram", fontsize=18, fontweight='bold')
plt.xlabel("NDVI values", fontsize=14)
plt.ylabel("# pixels", fontsize=14)
x = ndvi[~np.isnan(ndvi)]
numBins = 20
ax.hist(x,numBins,color='green',alpha=0.8)
fig2.savefig("output/ndvi-histogram.png", dpi=200, bbox_inches='tight', pad_inches=0.7)
plt.show()
```
| github_jupyter |
```
%%configure -f
{"conf":{"spark.driver.extraClassPath":"/usr/lib/hadoop-lzo/lib/*:/usr/lib/hadoop/hadoop-aws.jar:/usr/share/aws/aws-java-sdk/*:/usr/share/aws/emr/goodies/lib/emr-spark-goodies.jar:/usr/share/aws/emr/security/conf:/usr/share/aws/emr/security/lib/*:/usr/share/aws/hmclient/lib/aws-glue-datacatalog-spark-client.jar:/usr/share/java/Hive-JSON-Serde/hive-openx-serde.jar:/usr/share/aws/sagemaker-spark-sdk/lib/sagemaker-spark-sdk.jar:/usr/share/aws/emr/s3select/lib/emr-s3-select-spark-connector.jar:/docker/usr/lib/hadoop-lzo/lib/*:/docker/usr/lib/hadoop/hadoop-aws.jar:/docker/usr/share/aws/aws-java-sdk/*:/docker/usr/share/aws/emr/goodies/lib/emr-spark-goodies.jar:/docker/usr/share/aws/emr/security/conf:/docker/usr/share/aws/emr/security/lib/*:/docker/usr/share/aws/hmclient/lib/aws-glue-datacatalog-spark-client.jar:/docker/usr/share/java/Hive-JSON-Serde/hive-openx-serde.jar:/docker/usr/share/aws/sagemaker-spark-sdk/lib/sagemaker-spark-sdk.jar:/docker/usr/share/aws/emr/s3select/lib/emr-s3-select-spark-connector.jar,file:///opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar",
"spark.executor.extraClassPath":"/usr/lib/hadoop-lzo/lib/*:/usr/lib/hadoop/hadoop-aws.jar:/usr/share/aws/aws-java-sdk/*:/usr/share/aws/emr/goodies/lib/emr-spark-goodies.jar:/usr/share/aws/emr/security/conf:/usr/share/aws/emr/security/lib/*:/usr/share/aws/hmclient/lib/aws-glue-datacatalog-spark-client.jar:/usr/share/java/Hive-JSON-Serde/hive-openx-serde.jar:/usr/share/aws/sagemaker-spark-sdk/lib/sagemaker-spark-sdk.jar:/usr/share/aws/emr/s3select/lib/emr-s3-select-spark-connector.jar:/docker/usr/lib/hadoop-lzo/lib/*:/docker/usr/lib/hadoop/hadoop-aws.jar:/docker/usr/share/aws/aws-java-sdk/*:/docker/usr/share/aws/emr/goodies/lib/emr-spark-goodies.jar:/docker/usr/share/aws/emr/security/conf:/docker/usr/share/aws/emr/security/lib/*:/docker/usr/share/aws/hmclient/lib/aws-glue-datacatalog-spark-client.jar:/docker/usr/share/java/Hive-JSON-Serde/hive-openx-serde.jar:/docker/usr/share/aws/sagemaker-spark-sdk/lib/sagemaker-spark-sdk.jar:/docker/usr/share/aws/emr/s3select/lib/emr-s3-select-spark-connector.jar,file:///opt/benchmark-tools/spark-sql-perf/target/scala-2.12/spark-sql-perf_2.12-0.5.1-SNAPSHOT.jar"}}
val scaleFactor = "1" // data scale 1GB
val iterations = 1 // how many times to run the whole set of queries.
val format = "parquet" // support parquer or orc
val storage = "s3" // support hdfs or s3
var bucket_name = "aws-emr-resources-348941870272-us-east-2" // when storage is "s3", this value will be use.
val partitionTables = true // create partition tables
val query_filter = Seq() // Seq() == all queries
//val query_filter = Seq("q1-v2.4", "q2-v2.4") // run subset of queries
val randomizeQueries = false // run queries in a random order. Recommended for parallel runs.
if (storage == "hdfs"){
bucket_name = "/user/livy" // scala notebook only has the write permission of "hdfs:///user/livy" directory
}
// detailed results will be written as JSON to this location.
var resultLocation = s"${storage}://${bucket_name}/results/tpcds_${format}/${scaleFactor}/"
var databaseName = s"tpcds_${format}_scale_${scaleFactor}_db"
val use_arrow = false // when you want to use gazella_plugin to run TPC-DS, you need to set it true.
if (use_arrow){
val data_path= s"${storage}://${bucket_name}/datagen/tpcds_${format}/${scaleFactor}"
resultLocation = s"${storage}://${bucket_name}/results/tpcds_arrow/${scaleFactor}/"
databaseName = s"tpcds_arrow_scale_${scaleFactor}_db"
val tables = Seq("call_center", "catalog_page", "catalog_returns", "catalog_sales", "customer", "customer_address", "customer_demographics", "date_dim", "household_demographics", "income_band", "inventory", "item", "promotion", "reason", "ship_mode", "store", "store_returns", "store_sales", "time_dim", "warehouse", "web_page", "web_returns", "web_sales", "web_site")
sql(s"DROP database $databaseName CASCADE")
if (spark.catalog.databaseExists(s"$databaseName")) {
println(s"$databaseName has exists!")
}else{
spark.sql(s"create database if not exists $databaseName").show
spark.sql(s"use $databaseName").show
for (table <- tables) {
if (spark.catalog.tableExists(s"$table")){
println(s"$table has exists!")
}else{
spark.catalog.createTable(s"$table", s"$data_path/$table", "arrow")
}
}
if (partitionTables) {
for (table <- tables) {
try{
spark.sql(s"ALTER TABLE $table RECOVER PARTITIONS").show
}catch{
case e: Exception => println(e)
}
}
}
}
}
val timeout = 60 // timeout in hours
// COMMAND ----------
// Spark configuration
spark.conf.set("spark.sql.broadcastTimeout", "10000") // good idea for Q14, Q88.
// ... + any other configuration tuning
// COMMAND ----------
sql(s"use $databaseName")
import com.databricks.spark.sql.perf.tpcds.TPCDS
val tpcds = new TPCDS (sqlContext = spark.sqlContext)
def queries = {
val filtered_queries = query_filter match {
case Seq() => tpcds.tpcds2_4Queries
case _ => tpcds.tpcds2_4Queries.filter(q => query_filter.contains(q.name))
}
if (randomizeQueries) scala.util.Random.shuffle(filtered_queries) else filtered_queries
}
val experiment = tpcds.runExperiment(
queries,
iterations = iterations,
resultLocation = resultLocation,
tags = Map("runtype" -> "benchmark", "database" -> databaseName, "scale_factor" -> scaleFactor))
println(experiment.toString)
experiment.waitForFinish(timeout*60*60)
```
| github_jupyter |
```
# Copyright 2020 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# 1.1. BigQuery Storage & Spark DataFrames - Python
### Create Dataproc Cluster with Jupyter
This notebook is designed to be run on Google Cloud Dataproc.
Follow the links below for instructions on how to create a Dataproc Cluster with the Juypter component installed.
* [Tutorial - Install and run a Jupyter notebook on a Dataproc cluster](https://cloud.google.com/dataproc/docs/tutorials/jupyter-notebook)
* [Blog post - Apache Spark and Jupyter Notebooks made easy with Dataproc component gateway](https://medium.com/google-cloud/apache-spark-and-jupyter-notebooks-made-easy-with-dataproc-component-gateway-fa91d48d6a5a)
### Python 3 Kernel
Use a Python 3 kernel (not PySpark) to allow you to configure the SparkSession in the notebook and include the [spark-bigquery-connector](https://github.com/GoogleCloudDataproc/spark-bigquery-connector) required to use the [BigQuery Storage API](https://cloud.google.com/bigquery/docs/reference/storage).
### Scala Version
Check what version of Scala you are running so you can include the correct spark-bigquery-connector jar
```
!scala -version
```
### Create Spark Session
Include the correct version of the spark-bigquery-connector jar
Scala version 2.11 - `'gs://spark-lib/bigquery/spark-bigquery-latest.jar'`.
Scala version 2.12 - `'gs://spark-lib/bigquery/spark-bigquery-latest_2.12.jar'`.
```
from pyspark.sql import SparkSession
spark = SparkSession.builder \
.appName('1.1. BigQuery Storage & Spark DataFrames - Python')\
.config('spark.jars', 'gs://spark-lib/bigquery/spark-bigquery-latest.jar') \
.getOrCreate()
```
### Enable repl.eagerEval
This will output the results of DataFrames in each step without the new need to show `df.show()` and also improves the formatting of the output
```
spark.conf.set("spark.sql.repl.eagerEval.enabled",True)
```
### Read BigQuery table into Spark DataFrame
Use `filter()` to query data from a partitioned table.
```
table = "bigquery-public-data.wikipedia.pageviews_2020"
df_wiki_pageviews = spark.read \
.format("bigquery") \
.option("table", table) \
.option("filter", "datehour >= '2020-03-01' AND datehour < '2020-03-02'") \
.load()
df_wiki_pageviews.printSchema()
```
Select required columns and apply a filter using `where()` which is an alias for `filter()` then cache the table
```
df_wiki_en = df_wiki_pageviews \
.select("title", "wiki", "views") \
.where("views > 1000 AND wiki in ('en', 'en.m')") \
.cache()
df_wiki_en
```
Group by title and order by page views to see the top pages
```
import pyspark.sql.functions as F
df_wiki_en_totals = df_wiki_en \
.groupBy("title") \
.agg(F.sum('views').alias('total_views'))
df_wiki_en_totals.orderBy('total_views', ascending=False)
```
### Write Spark Dataframe to BigQuery table
Write the Spark Dataframe to BigQuery table using BigQuery Storage connector. This will also create the table if it does not exist. The GCS bucket and BigQuery dataset must already exist.
If the GCS bucket and BigQuery dataset do not exist they will need to be created before running `df.write`
- [Instructions here for creating a GCS bucket](https://cloud.google.com/storage/docs/creating-buckets)
- [Instructions here for creating a BigQuery Dataset](https://cloud.google.com/bigquery/docs/datasets)
```
# Update to your GCS bucket
gcs_bucket = 'dataproc-bucket-name'
# Update to your BigQuery dataset name you created
bq_dataset = 'dataset_name'
# Enter BigQuery table name you want to create or overwite.
# If the table does not exist it will be created when you run the write function
bq_table = 'wiki_total_pageviews'
df_wiki_en_totals.write \
.format("bigquery") \
.option("table","{}.{}".format(bq_dataset, bq_table)) \
.option("temporaryGcsBucket", gcs_bucket) \
.mode('overwrite') \
.save()
```
### Use BigQuery magic to query table
Use the [BigQuery magic](https://googleapis.dev/python/bigquery/latest/magics.html) to check if the data was created successfully in BigQuery. This will run the SQL query in BigQuery and the return the results
```
%%bigquery
SELECT title, total_views
FROM dataset_name.wiki_total_pageviews
ORDER BY total_views DESC
LIMIT 10
```
| github_jupyter |
<a href="https://colab.research.google.com/github/diascarolina/project-icu-prediction/blob/main/notebooks/01_data_cleaning_and_analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# COVID-19 - Clinical Data to Assess Diagnosis
<p align="center">
<img width="700" src="https://i.imgur.com/wxaTMWn.png">
</p>
# Table of Contents
1. [What **problem** do we have here? What can we do to **help**?](#intro)
2. [How was this notebook **organized**?](#orga)
3. [Where did we get this **data**? What kind of **information** do we have in it?](#data)
4. [Libraries & Configurations](#libs)
5. [Data Checking: is everything in its right place?](#check)
6. [Missing Values & Data Preparation](#prep)
7. [Data Analysis](#eda)
7.1. [Patient Demographic Information](#pdi)
7.2. [Patient Previous Grouped Diseases](#ppgd)
7.3. [Vital Signs](#vitals)
7.4. [Blood Results](#blood)
8. [Conclusions](#conc)
[Part 2 - Machine Learning](#partwo)
<a name="intro"></a>
# 1 What **problem** do we have here? What can we do to **help**?
The COVID-19 pandemic. Unfortunately we are all aware of it by now. We all know the sad and high number of deaths and the hopeful increasing number of vaccinated people (in some countries).
Seeing as the number of people infected by the virus keeps growing, we need more and more ways of better allocating these patients so as not to overwhelm our healthcare systems.
In this context, brazilian hospital Sírio-Libânes [published a dataset at Kaggle](https://www.kaggle.com/S%C3%ADrio-Libanes/covid19) urging people to take action in predicting the need for ICU beds.
The tasks at hand are:
**Task 01:** Predict admission to the ICU of confirmed COVID-19 cases.
Based on the data available, is it feasible to predict which patients will need intensive care unit support?
The aim is to provide tertiary and quarternary hospitals with the most accurate answer, so ICU resources can be arranged or patient transfer can be scheduled.
**Task 02:** Predict NOT admission to the ICU of confirmed COVID-19 cases.
Based on the subsample of widely available data, is it feasible to predict which patients will need intensive care unit support?
The aim is to provide local and temporary hospitals a good enough answer, so frontline physicians can safely discharge and remotely follow up with these patients.
<a name="orga"></a>
# 2 How was this notebook organized?
In this **Part 1** we'll begin by **checking** and **cleaning** our data. For this, we'll follow these steps:

After that, we'll **analyse** the data as the following image:

<a name="data"></a>
# 3 Where did we get this **data**? What kind of **information** do we have in it?
The data was taken directly from the [Kaggle problem](https://www.kaggle.com/S%C3%ADrio-Libanes/covid19) and uploaded to [Github](https://github.com/diascarolina/data-science-bootcamp/blob/main/data/Kaggle_Sirio_Libanes_ICU_Prediction.xlsx?raw=true) for easy access.
**From Kaggle:**
> The dataset contains **anonymized data** from Hospital Sírio-Libanês, São Paulo and Brasília. All data were anonymized following the best international practices and recommendations.
> Data has been cleaned and **scaled** by column according to MinMaxScaler to fit between -1 and 1.
_Disclaimer._ It is highly recommended that we apply a scaler to the data only **after** splitting between train and test data (_source:_ [Data Normalization Before or After Splitting a Data Set?](https://www.baeldung.com/cs/data-normalization-before-after-splitting-set)). But since our data is already scaled, we won't go in too much detail in this aspect.
**From our dataset we have:**
1. Patient demographic information (03 columns)
2. Patient previous grouped diseases (09 columns)
3. Blood results (36 columns)
4. Vital signs (06 columns)
In total there are **54 features**, expanded when pertinent to the _mean, median, max, min, diff and relative diff_, where
- _diff = max - min_
- _relative diff = diff/median_
Furthermore, **our target variable is the ICU** column, in which we have **0** (zero) if that patient did not go to the ICU and **1** (one) if that corresponding patient did go to the ICU.
We also have a column called **Window**. What does this variable mean?
Again, from Kaggle:
> We were careful to include real life cenarios of window of events and available data. Data was obtain and grouped as follows:
- patient
- patient encounter
- aggregated by windows in chronological order
Window | Description
:---: | :---:
**0-2** | From 0 to 2 hours of the admission
**2-4** | From 2 to 4 hours of the admission
**4-6** | From 4 to 6 hours of the admission
**6-12** | From 6 to 12 hours of the admission
**Above-12** | Above 12 hours from admission
> Beware **NOT to use the data when the target variable is present**, as it is unknown the order of the event (maybe the target event happened before the results were obtained). They were kept there so we can grow this dataset in other outcomes latter on.
**Examples obtained from Kaggle description:**
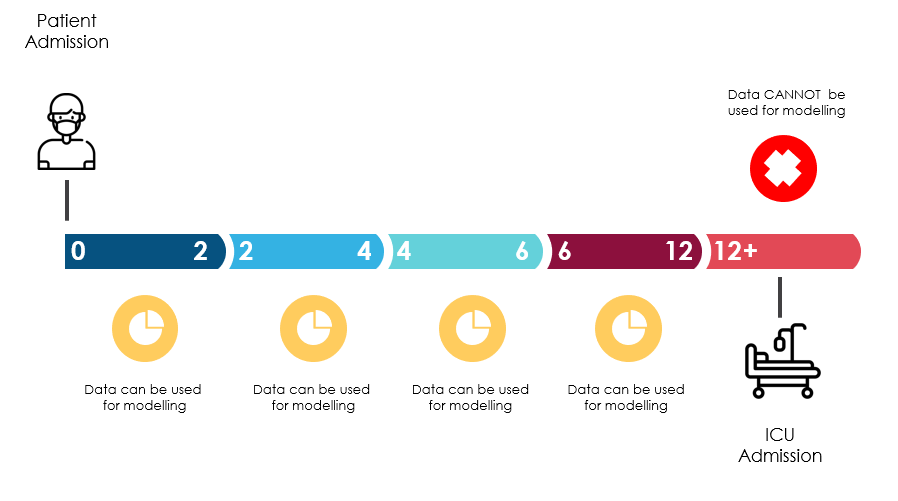
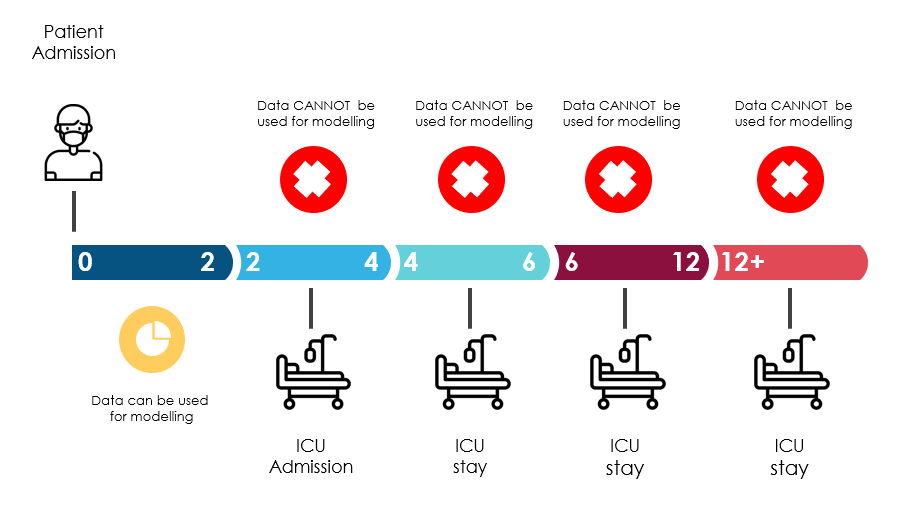
<a name="libs"></a>
# 4 Libraries & Configurations
Remember that this notebook is only used for data cleaning and analysis, so we'll only use libraries related to this task.
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# colors definition
GRAY1, GRAY2, GRAY3 = '#231F20', '#414040', '#555655'
GRAY4, GRAY5, GRAY6 = '#646369', '#76787B', '#828282'
GRAY7, GRAY8, GRAY9 = '#929497', '#A6A6A5', '#BFBEBE'
BLUE1, BLUE2, BLUE3, BLUE4 = '#174A7E', '#4A81BF', '#94B2D7', '#94AFC5'
RED1, RED2 = '#C3514E', '#E6BAB7'
GREEN1, GREEN2 = '#0C8040', '#9ABB59'
ORANGE1 = '#F79747'
# charts configs
plt.rcParams['font.family'] = 'Arial'
plt.rcParams['mathtext.fontset'] = 'custom'
plt.rcParams['mathtext.bf'] = 'Arial:bold'
plt.rcParams['mathtext.it'] = 'Arial:italic'
```
<a name="check"></a>
# 5 Data Checking: is everything in its right place?
```
data_url = 'https://github.com/diascarolina/data-science-bootcamp/blob/main/data/Kaggle_Sirio_Libanes_ICU_Prediction.xlsx?raw=true'
raw_data = pd.read_excel(data_url)
raw_data.head(15)
```
Let's check this data.
```
def print_data_shape(dataset):
'''
Prints the number of rows and columns in a pandas dataframe.
Input:
dataset -> a pandas dataframe
'''
print(f'Number of rows in the dataset: {dataset.shape[0]}')
print(f'Number of columns in the dataset: {dataset.shape[1]}')
print_data_shape(raw_data)
```
We have 1925 rows in our dataset. But does this mean that each row represents a patient? Let's check the column ```PATIENT_VISIT_IDENTIFIER```.
```
num_unique = len(raw_data['PATIENT_VISIT_IDENTIFIER'].unique())
print(f'Number of unique values in the column PATIENT_VISIT_IDENTIFIER: {num_unique}')
```
With the cell above, we see that we have 385 unique values in the ```PATIENT_VISIT_IDENTIFIER``` columns. This is exactly the number of rows, 1925, divided by 5. This means we only have 385 patients in this dataset, each one represented by 5 rows. Each of these 5 rows for a patient correspond with the ```WINDOW``` column.
```
raw_data['WINDOW'].unique()
```
We can see that the ```WINDOW``` column is divided into the following categories:
- 0-2
- 2-4
- 4-6
- 6-12
- ABOVE_12
Each of these categories represent a time window with how many hours did it take for the patient to go the ICU. For example, if a patient went to the ICU between 2 and 6 of being admited, then we would have the following data:
**WINDOW** | **ICU**
:---:|:---:
0-2 | 0
2-4 | 1
4-6 | 1
6-12 | 1
ABOVE_12 | 1
**Obs.** All of this information is also present in the [Kaggle problem page](https://www.kaggle.com/S%C3%ADrio-Libanes/covid19).
<a name="prep"></a>
# 6 Missing Values & Data Preparation
Just from taking a quick look at the data we can see we have some missing values, specially on the columns where we have continuous values, ranging from the column ```ALBUMIN_MEDIAN``` to ```OXYGEN_SATURATION_DIFF_REL```. These represent the measurements from the patient and said measurements are not taken constantly, as explained bellow (also found at the [Kaggle problem page](https://www.kaggle.com/S%C3%ADrio-Libanes/covid19)).
> **Problem:** One of the major challenges of working with health care data is that the sampling rate varies across different type of measurements. For instance, vital signs are sampled more frequently (usually hourly) than blood labs (usually daily).
> **Tips & Tricks:** It is reasonable to assume that a patient who does not have a measurement recorded in a time window is clinically stable, potentially presenting vital signs and blood labs similar to neighboring windows. Therefore, one may fill the missing values using the next or previous entry. Attention to multicollinearity and zero variance issues in this data when choosing your algorithm.
With this in mind, we'll use this approach of filling the missing values using the next or the previous entry. For this, we'll create a function that fills the missing values for the continuous variables in the dataset. These variables are already aggregated for us in the columns ```[13:-2]```.
The methods used will be the "forward fill"and the "backfill".
But first let's check how many missing values we have in the whole dataset.
```
def check_missing_values(dataset):
'''
Function that prints the number of missing values in a dataframe, together
with its percentage against the total number of values in the dataframe.
Input:
dataset -> a pandas dataframe
Output:
A print statement.
'''
number_of_null_values = dataset.isnull().values.sum()
# this is just the number of rows times the number of columns
total_values_in_dataset = dataset.shape[0] * dataset.shape[1]
percentage = number_of_null_values / total_values_in_dataset * 100
print(f'We have {number_of_null_values} null values in this dataset.')
print(f'This represents {round(percentage, 2)}% of the total number of values in the dataset')
check_missing_values(raw_data)
```
We could try and visualize graphically the missing values and its distribution in the dataset, using a library like [missingno](https://github.com/ResidentMario/missingno). But since we have so many columns to visualize, it would not be a very good visualization.
Bellow we define the function to fill the values in the continous variables using the previous or the next observation in the dataset.
```
def fill_dataset(data):
'''
Function that fills the missing values in the dataset with the methods 'bfill'
and 'ffill'. This substitution is only applied in the continuous variables.
Input:
data -> a pandas dataframe
Output:
filled_data -> a pandas dataframe without missing values in the continuous variables
'''
# select the columns with the continuous variables
columns_continuous_features = data.iloc[:, 13:-2].columns
# group the dataset per patient and we substitute the missing values
# using the 'bfill' and 'ffill' methods for each patient
continuous_features = data.groupby('PATIENT_VISIT_IDENTIFIER',
as_index = False)[columns_continuous_features].fillna(method = 'bfill').fillna(method = 'ffill')
categorical_features = data.iloc[:, :13]
output_data = data.iloc[:, -2:]
filled_data = pd.concat([categorical_features, continuous_features, output_data],
ignore_index = True,
axis = 1)
filled_data.columns = data.columns
return filled_data
data = raw_data.copy()
filled_data = fill_dataset(data)
filled_data.head(10)
```
So now, by just taking a look at the data above, we see that the missing values in the continuous variables were filled.
Let's check if we have any other null values in the other columns.
```
check_missing_values(filled_data)
```
We can see we still have some missing values, but significantly less than before. We can simply go ahead and remove these ones, since they represent only a smal percentage of the dataset and are probably from errors in the data.
We can also go ahead and remove the observations in which the admitted patients went to the ICU at window 0-2. This is also a recommendation from the hospital itself, regarding machine learning model and the usefulness of the data:
**The earlier, the better!**
> **Problem:** Early identification of those patients who will develop an adverse course of illness (and need intensive care) is a key for an appropriate treatment (saving lives) and to managing beds and resources.
> **Tips & Tricks:** _Whereas a predictive model using all time windows will probably yield a greater accuracy, a nice model using only the first (0-2) is likely to be more clinically relevant._
We also have:
> [...] patients that went to the ICU on 0-2 window have no available data for predicting, therefore should also be excluded from the sample.
So let's do that.
```
# check the number of 1's in the 0-2 window
pd.crosstab(filled_data['WINDOW'], filled_data['ICU'])
```
As seen above, we have **32 observations in which the patient went to the ICU at the 0-2 window**. So we'll discard these 32 patients from our dataset.
```
# let's get only the patients which did not went to the ICU in the first two hours of admission
to_be_removed = filled_data.query("WINDOW=='0-2' and ICU==1")['PATIENT_VISIT_IDENTIFIER'].values
filled_data = filled_data.query("PATIENT_VISIT_IDENTIFIER not in @to_be_removed")
# drop the remaining missing values
filled_data = filled_data.dropna()
filled_data.head()
check_missing_values(filled_data)
print_data_shape(filled_data)
```
Now, we'll group the rows by patient to obtain a dataset where each row represents only one patient, instead of 5 rows per patient.
```
def prepare_window(rows):
'''
Clean the "WINDOW" row in the dataframe.
'''
if(np.any(rows['ICU'])):
rows.loc[rows['WINDOW'] == '0-2', 'ICU'] = 1
# return the WINDOW column for the first two hours
return rows.loc[rows['WINDOW'] == '0-2']
# group the data by patient and apply the function above
clean_data = filled_data.groupby('PATIENT_VISIT_IDENTIFIER').apply(prepare_window)
# change the AGE_PERCENTIL column to a 'category' type
clean_data.AGE_PERCENTIL = clean_data.AGE_PERCENTIL.astype('category').cat.codes
clean_data.head()
```
We can go ahead and drop the column ```WINDOW``` since we now have only one value in it: ```0-2```. We'll also drop the ```PATIENT_VISIT_IDENTIFIER``` since we don't need this information in order to predict the ICU admission.
```
clean_data = clean_data.drop(['WINDOW', 'PATIENT_VISIT_IDENTIFIER'], axis = 1)
check_missing_values(clean_data)
print_data_shape(clean_data)
```
Comparing to the previous dataframe, we now reduced our information from 1760 rows to only 352!
With this new and clean dataset, we'll take a closer look at the variables in order to reduce the large number of columns and explore these features. Furthermore, seeing as we have many features that were created from a single variable, we have to address them and check their correlation against each other. This helps us in reducing the risk of overfitting our machine learning model.
For the variables pertaining to the blood results and vital signs, they were expanded by the mean, median, max, min, diff and relative diff. Where ```diff = max - min``` and ```relative diff = diff/median```.
```
# let's check our data again
clean_data.head()
```
To check the highly correlated columns, we first declare a function that checks the correlation between each variable in the whole dataset and then it removes the ones that pass a certain threshold.
```
def remove_corr_var(data, threshold = 0.95):
'''
Given a dataset, this function calculates the correlation between the
variable and return a new dataset with only the variable with a correlation
below a certain threshold.
Input:
data -> a pandas dataframe
threshold -> a number (usually between 0 and 1) used as a cutoff for the
correlation, default = 0.95
'''
matrix_corr = data.iloc[:,4:-2].corr().abs()
matrix_upper = matrix_corr.where(np.triu(np.ones(matrix_corr.shape), k = 1).astype(np.bool))
to_remove = [coluna for coluna in matrix_upper.columns if any(matrix_upper[coluna] > threshold)]
return data.drop(to_remove, axis = 1)
final_data = remove_corr_var(clean_data)
final_data.head()
print_data_shape(final_data)
```
So now we have a dataset with 352 rows, one for each patient, and 99 columns. From these 99 columns, 98 are variables and 1 is the target, the ICU column.
```
# uncomment the following line to download the final data
#final_data.to_csv('final_data.csv', index = False)
#from google.colab import files
#files.download('final_data.csv')
```
<a name="eda"></a>
# 7 Data Analysis
Now we'll take a closer look at some features to check their distribution and whether or not they are balanced in order to be used in a machine learning model. If they are highly unbalanced, it could create a problem for the modelling part.
```
# remembering our dataframe
final_data.head()
```
We have 03 patient demographic information variables: ```AGE_ABOVE65```, ```AGE_PERCENTIL``` and ```GENDER```. Let's take a look at them together with the ```ICU``` column, our target variable. For reference, we can also take a look at the other continuous variables.
<a name="pdi"></a>
## 7.1 Patient Demographic Information
Let's check graphically how many patiets are in each category of demographic information.
```
color = [GREEN2, RED1]
ax = final_data['ICU'].value_counts().plot(kind = 'bar',
color = color,
figsize = (10, 6),
width = 0.9)
ax.tick_params(color = 'darkgrey', bottom = 'off')
ax.tick_params(axis = 'both',
which = 'both',
bottom = 'off',
top = 'off',
labelbottom = 'off',
right = 'off',
left = 'off',
labelleft = 'off')
sns.despine(left = True, bottom = True)
ax.set_xticklabels(['Patient did not\n go to the ICU', 'Patient went\n to the ICU'],
fontsize = 18, color = GRAY4)
ax.get_yaxis().set_visible(False)
plt.xticks(rotation = 0)
plt.title('ICU Admission', fontsize = 25, loc = 'left', color = GRAY2)
perc_not_icu = round(len(final_data[final_data['ICU'] != 1]) / len(final_data) * 100, 1)
perc_icu = round(len(final_data[final_data['ICU'] == 1]) / len(final_data) * 100, 1)
labels = [str(len(final_data[final_data['ICU'] == 0])) + ' patients\n(' + str(perc_not_icu) + ')%',
str(len(final_data[final_data['ICU'] == 1])) + ' patients\n(' + str(perc_icu) + ')%']
rects = ax.patches
for rect, label in zip(rects, labels):
height = rect.get_height()
ax.text(rect.get_x() + rect.get_width() / 2,
height-30,
label,
ha = 'center',
va = 'bottom',
color = 'white',
fontsize = 22,
fontweight = 'bold')
plt.show()
```
We don't have a hard threshold for when to consider our data unbalanced, but [some sources](https://developers.google.com/machine-learning/data-prep/construct/sampling-splitting/imbalanced-data) indicate that we can begin to consider our data **unbalanced** when the difference between the classes is **above 20%**. Here in our case, the difference is only **7.4%**, so we'll consider our **target variable balanced**.
```
color = [GREEN2, RED1]
fig, axes = plt.subplots(1, 2, sharey = True, figsize = (16, 7))
final_data[final_data.GENDER == 0].ICU.value_counts().sort_values(ascending = True).plot(kind = 'bar',
ax = axes[0],
color = color,
width = 0.9)
axes[0].set_title('Gender 0: Man', loc = 'left', fontsize = 19, color = GRAY2)
final_data[final_data.GENDER == 1].ICU.value_counts().plot(kind = 'bar',
ax = axes[1],
color = color,
width = 0.9)
axes[1].set_title('Gender 1: Woman', loc = 'left', fontsize = 19, color = GRAY2)
sns.despine(left = True, bottom = True)
for i in [0, 1]:
axes[i].tick_params(color = 'darkgrey', bottom = 'off')
axes[i].tick_params(axis = 'both',
which = 'both',
bottom = 'off',
top = 'off',
labelbottom = 'off',
right = 'off',
left = 'off',
labelleft = 'off')
axes[i].set_xticklabels(['Patient did not\n go to the ICU', 'Patient went\n to the ICU'],
fontsize = 18, color = GRAY4, rotation = 0)
axes[i].get_yaxis().set_visible(False)
plt.suptitle('ICU Admission by Gender', fontsize = 24, color = GRAY1, y = 1)
# Code for chart annotation
perc_not_icu_0 = round(final_data[final_data.GENDER == 0].ICU.value_counts()[0] / len(final_data[final_data.GENDER == 0]) * 100, 1)
perc_icu_0 = round(final_data[final_data.GENDER == 0].ICU.value_counts()[1] / len(final_data[final_data.GENDER == 0]) * 100, 1)
labels = [str(final_data[final_data.GENDER == 0].ICU.value_counts()[0]) + ' patients\n(' + str(perc_not_icu_0) + ')%',
str(final_data[final_data.GENDER == 0].ICU.value_counts()[1]) + ' patients\n(' + str(perc_icu_0) + ')%']
rects = axes[0].patches
for rect, label in zip(rects, labels):
height = rect.get_height()
axes[0].text(rect.get_x() + rect.get_width() / 2,
height-15,
label,
ha = 'center',
va = 'bottom',
color = 'white',
fontsize = 18,
fontweight = 'bold')
perc_not_icu_1 = round(final_data[final_data.GENDER == 1].ICU.value_counts()[0] / len(final_data[final_data.GENDER == 1]) * 100, 1)
perc_icu_1 = round(final_data[final_data.GENDER == 1].ICU.value_counts()[1] / len(final_data[final_data.GENDER == 1]) * 100, 1)
labels = [str(final_data[final_data.GENDER == 1].ICU.value_counts()[0]) + ' patients\n(' + str(perc_not_icu_1) + ')%',
str(final_data[final_data.GENDER == 1].ICU.value_counts()[1]) + ' patients\n(' + str(perc_icu_1) + ')%']
rects = axes[1].patches
for rect, label in zip(rects, labels):
height = rect.get_height()
axes[1].text(rect.get_x() + rect.get_width() / 2,
height-15,
label,
ha = 'center',
va = 'bottom',
color = 'white',
fontsize = 18,
fontweight = 'bold')
plt.show()
```
Here we can immediately see some interesting facts about how many man go to the ICU in contrast to the women.
This phenomenon has already been well documented in this study: [Male sex identified by global COVID-19 meta-analysis as a risk factor for death and ITU admission](nature.com/articles/s41467-020-19741-6)
So gender really plays an important role in the ICU prediction we'll be making.
```
color = [GREEN2, RED1]
fig, axes = plt.subplots(1, 2, sharey = True, figsize = (16, 7))
final_data[final_data.AGE_ABOVE65 == 0].ICU.value_counts().plot(kind = 'bar',
ax = axes[0],
color = color,
width = 0.9)
axes[0].set_title('Patient is Younger than 65 Years', loc = 'left', fontsize = 19, color = GRAY2)
final_data[final_data.AGE_ABOVE65 == 1].ICU.value_counts().sort_values(ascending = True).plot(kind = 'bar',
ax = axes[1],
color = color,
width = 0.9)
axes[1].set_title('Patient is Older than 65 Years', loc = 'left', fontsize = 19, color = GRAY2)
sns.despine(left = True, bottom = True)
for i in [0, 1]:
axes[i].tick_params(color = 'darkgrey', bottom = 'off')
axes[i].tick_params(axis = 'both',
which = 'both',
bottom = 'off',
top = 'off',
labelbottom = 'off',
right = 'off',
left = 'off',
labelleft = 'off')
axes[i].set_xticklabels(['Patient did not\n go to the ICU', 'Patient went\n to the ICU'],
fontsize = 18, color = GRAY4, rotation = 0)
axes[i].get_yaxis().set_visible(False)
plt.suptitle('ICU Admission by Age', fontsize = 24, color = GRAY1, y = 1)
# Code for chart annotation
per_not_icu_0 = round(final_data[final_data.AGE_ABOVE65 == 0].ICU.value_counts()[0] / len(final_data[final_data.AGE_ABOVE65 == 0]) * 100, 1)
per_icu_0 = round(final_data[final_data.AGE_ABOVE65 == 0].ICU.value_counts()[1] / len(final_data[final_data.AGE_ABOVE65 == 0]) * 100, 1)
labels = [str(final_data[final_data.AGE_ABOVE65 == 0].ICU.value_counts()[0]) + ' patients\n(' + str(per_not_icu_0) + ')%',
str(final_data[final_data.AGE_ABOVE65 == 0].ICU.value_counts()[1]) + ' patients\n(' + str(per_icu_0) + ')%']
rects = axes[0].patches
for rect, label in zip(rects, labels):
height = rect.get_height()
axes[0].text(rect.get_x() + rect.get_width() / 2,
height-18,
label,
ha = 'center',
va = 'bottom',
color = 'white',
fontsize = 18,
fontweight = 'bold')
per_not_icu_1 = round(final_data[final_data.AGE_ABOVE65 == 1].ICU.value_counts()[0] / len(final_data[final_data.AGE_ABOVE65 == 1]) * 100, 1)
per_icu_1 = round(final_data[final_data.AGE_ABOVE65 == 1].ICU.value_counts()[1] / len(final_data[final_data.AGE_ABOVE65 == 1]) * 100, 1)
labels = [str(final_data[final_data.AGE_ABOVE65 == 1].ICU.value_counts()[0]) + ' patients\n(' + str(per_not_icu_1) + ')%',
str(final_data[final_data.AGE_ABOVE65 == 1].ICU.value_counts()[1]) + ' patients\n(' + str(per_icu_1) + ')%']
rects = axes[1].patches
for rect, label in zip(rects, labels):
height = rect.get_height()
axes[1].text(rect.get_x() + rect.get_width() / 2,
height-18,
label,
ha = 'center',
va = 'bottom',
color = 'white',
fontsize = 18,
fontweight = 'bold')
plt.show()
```
It's also noticeable how age affects ICU admission. Let's see a more detailed breakdown below.
```
label_names = ['10th', '20th', '30th', '40th', '50th', '60th', '70th', '80th', '90th', '>90th']
label_codes = [0, 1, 2, 3, 4, 5, 6, 7, 8, 9]
not_icu = []
icu = []
for code in label_codes:
not_icu.append(final_data[final_data.AGE_PERCENTIL == code].ICU.value_counts()[0])
icu.append(final_data[final_data.AGE_PERCENTIL == code].ICU.value_counts()[1])
x = np.arange(len(label_names))
width = 0.35
fig, ax = plt.subplots(figsize = (15, 8))
rects1 = ax.bar(x - width/2, not_icu, width, label = 'No ICU', color = GREEN2)
rects2 = ax.bar(x + width/2, icu, width, label = 'ICU', color = RED1)
ax.set_ylabel('Number of Patients', fontsize = 18)
ax.set_xlabel('Age Percentil', fontsize = 18, color = GRAY4)
ax.set_title('ICU Admission by Age Group', loc = 'left', fontsize = 24, color = GRAY2)
ax.set_xticks(x)
ax.set_xticklabels(label_names, fontsize = 18, color = GRAY4, rotation = 0)
ax.yaxis.label.set_color(GRAY4)
ax.tick_params(axis = 'both', colors = GRAY4)
ax.tick_params(axis = 'y', labelsize = 18)
ax.set_xticklabels(label_names)
plt.legend(fontsize = 18, frameon = False)
sns.despine()
fig.tight_layout()
plt.show()
```
Here, it is imediate to see that people above 70 years of age are highly more susceptible in needing intensive care than younger people.
This also has been largely studied and we all know by know how age is a risk factor for Covid-19.
_Source:_ [The impact of frailty on survival in elderly intensive care patients with COVID-19](https://ccforum.biomedcentral.com/articles/10.1186/s13054-021-03551-3)
<a name="ppgd"></a>
## 7.2 Patient Previous Grouped Diseases
```
diseases = {
'DISEASE GROUPING 1': {'not_icu': [], 'icu': []},
'DISEASE GROUPING 2': {'not_icu': [], 'icu': []},
'DISEASE GROUPING 3': {'not_icu': [], 'icu': []},
'DISEASE GROUPING 4': {'not_icu': [], 'icu': []},
'DISEASE GROUPING 5': {'not_icu': [], 'icu': []},
'DISEASE GROUPING 6': {'not_icu': [], 'icu': []},
'HTN': {'not_icu': [], 'icu': []},
'IMMUNOCOMPROMISED': {'not_icu': [], 'icu': []},
'OTHER': {'not_icu': [], 'icu': []}
}
color = [GREEN2, RED1]
label_names = ['Patient doesn\'t have\nGroup Disease', 'Patient has\nGroup Disease']
for disease in diseases.keys():
for num in [0, 1]:
diseases[disease]['not_icu'].append(final_data[final_data[disease] == num].ICU.value_counts()[0])
diseases[disease]['icu'].append(final_data[final_data[disease] == num].ICU.value_counts()[1])
x = np.arange(len(label_names))
width = 0.40
ax = [0, 1, 2, 3, 4, 5, 6, 7, 8]
fig, ((ax[0], ax[1], ax[2]), (ax[3], ax[4], ax[5]), (ax[6],ax[7], ax[8])) = plt.subplots(3, 3, sharey = True, figsize=(16, 10))
for i in [0, 1, 2, 3, 4, 5, 6, 7, 8]:
disease = list(diseases.keys())[i]
ax[i].bar(x - width/2, diseases[disease]['not_icu'], width, label = 'No ICU', color = GREEN2)
ax[i].bar(x + width/2, diseases[disease]['icu'], width, label = 'ICU', color = RED1)
#ax[i].set_ylabel('Number of Patients', fontsize = 16)
ax[i].set_title(disease.title(), loc = 'left', fontsize = 18, color = GRAY2)
ax[i].set_xticks(x)
ax[i].set_xticklabels(label_names, fontsize = 12, color = GRAY4, rotation = 0)
ax[i].yaxis.label.set_color(GRAY4)
ax[i].tick_params(axis = 'both', colors = GRAY4)
ax[i].tick_params(axis = 'y', labelsize = 14)
ax[i].set_xticklabels(label_names)
ax[2].legend(fontsize = 16, frameon = False, loc = 'upper right', bbox_to_anchor = (1.3, 1.0))
sns.despine()
plt.suptitle('ICU Admission by Disease Grouping', color = GRAY1, fontsize = 22, y = 1.05)
fig.tight_layout()
plt.show()
```
Here we can see how some disease groups directly impact in the ICU need for a patient. In almost all cases, patients with previous diseases are more prone to need intensive care for Covid-19 than those without previous diseases.
<a name="vitals"></a>
## 7.3 Vital Signs
We have different measurement that are expanded for various metrics. Here we'll only use the mean of the vital signs columns.
```
columns = ['BLOODPRESSURE_DIASTOLIC_MEAN', 'BLOODPRESSURE_SISTOLIC_MEAN',
'HEART_RATE_MEAN', 'OXYGEN_SATURATION_MEAN',
'RESPIRATORY_RATE_MEAN', 'TEMPERATURE_MEAN', 'ICU']
ax = [0, 1, 2, 3, 4, 5]
fig, ((ax[0], ax[1], ax[2]), (ax[3], ax[4], ax[5])) = plt.subplots(nrows = 2,
ncols = 3,
sharex = True,
figsize = (16, 10))
for i in [0, 1, 2, 3, 4, 5]:
column = columns[i]
temp = final_data.reset_index()[[column, 'ICU']]
mdf = pd.melt(temp, id_vars = ['ICU'], var_name = [column[:-5]])
sns.boxplot(x = 'value', y = column[:-5], hue = 'ICU', data = mdf, palette = color, ax = ax[i])
ax[i].set_yticklabels([])
ax[i].set_ylabel(column[:-5], fontsize = 15)
if i != 2:
ax[i].legend([],[], frameon = False)
for j in ax[i].get_xticklabels():
j.set_fontsize(13)
j.set_color(GRAY1)
sns.despine()
plt.suptitle('ICU Admission by Vital Signs Results', fontsize = 20, color = GRAY1)
plt.show()
```
**What information can we gather from this charts?**
First of all, the data, as seen before, has been scaled to fit between -1 and 1 in order to anonymize it, so we cannot be sure what exactly are these values. But we can make some assumptions from this data as is.
First, ```BLOODPRESSURE_DIASTOLIC``` in patients that needed to go to the ICU is a bit lower than in those patients that didn't go to the ICU. But the ```BLOODPRESSURE_SISTOLIC``` is just a bit above in those patients in the ICU, as oposed to those out of the ICU.
The ```HEART_RATE``` is a bit lower on ICU patients, but with outliers in both groups.
One of the most relevant features, ```OXYGEN_SATURATION``` has many outliers in the group that needed the ICU. A low oxygen saturation is a cause for concern in respiratory and COVID-19 patients. The ```RESPIRATORY_RATE``` is also higher in patients with admission to the ICU, as seen in the chart above.
_Source:_ [Oxygenation and Ventilation](https://www.covid19treatmentguidelines.nih.gov/management/critical-care/oxygenation-and-ventilation/)
The average ```TEMPERATURE``` is also higher in those patients that needed to go to the ICU, which could represent fever as a symptom.
_Source:_ [Symptoms of COVID-19](https://www.cdc.gov/coronavirus/2019-ncov/symptoms-testing/symptoms.html)
<a name="blood"></a>
## 7.4 Blood Results
Here we will use the median of the relevant columns.
```
# all other columns represent the blood results
columns_to_drop = ['BLOODPRESSURE_DIASTOLIC_MEAN', 'BLOODPRESSURE_SISTOLIC_MEAN',
'HEART_RATE_MEAN', 'OXYGEN_SATURATION_MEAN',
'RESPIRATORY_RATE_MEAN', 'TEMPERATURE_MEAN', 'GENDER',
'AGE_ABOVE65', 'AGE_PERCENTIL', 'DISEASE GROUPING 1', 'DISEASE GROUPING 2',
'DISEASE GROUPING 3', 'DISEASE GROUPING 4', 'DISEASE GROUPING 5',
'DISEASE GROUPING 6', 'HTN', 'IMMUNOCOMPROMISED', 'OTHER']
# we only want the columns that end with '_MEDIAN'
blood_results = final_data.drop(columns_to_drop, axis = 1).columns[final_data.drop(columns_to_drop, axis = 1).columns.str.endswith('_MEDIAN')]
blood_results
```
From the above columns let's pick some relevant ones so as not to plot them all and overwhelm our charts.
```
blood_results = ['CALCIUM_MEDIAN', 'HEMATOCRITE_MEDIAN', 'LACTATE_MEDIAN',
'LEUKOCYTES_MEDIAN', 'LINFOCITOS_MEDIAN', 'PC02_ARTERIAL_MEDIAN',
'PC02_VENOUS_MEDIAN', 'PCR_MEDIAN', 'PH_ARTERIAL_MEDIAN',
'PH_VENOUS_MEDIAN', 'PLATELETS_MEDIAN', 'POTASSIUM_MEDIAN',
'SAT02_ARTERIAL_MEDIAN', 'SAT02_VENOUS_MEDIAN', 'UREA_MEDIAN']
columns = list(blood_results)
columns.append('ICU')
ax = list(range(0, 15))
fig, ((ax[0], ax[1], ax[2], ax[3], ax[4]),
(ax[5], ax[6], ax[7], ax[8], ax[9]),
(ax[10], ax[11], ax[12], ax[13], ax[14])) = plt.subplots(ncols = 5, nrows = 3, sharex = True, figsize = (20, 10))
for i in range(0, 15):
column = columns[i]
temp = final_data.reset_index()[[column, 'ICU']]
mdf = pd.melt(temp, id_vars = ['ICU'], var_name = [column[:-7]])
sns.boxplot(x = 'value', y = column[:-7], hue = 'ICU', data = mdf, palette = color, ax = ax[i])
ax[i].set_yticklabels([])
ax[i].set_ylabel(column[:-7], fontsize = 15)
if i != 4:
ax[i].legend([],[], frameon = False)
for j in ax[i].get_xticklabels():
j.set_fontsize(13)
j.set_color(GRAY1)
sns.despine()
plt.suptitle('ICU Admission by Blood Results', fontsize = 25, color = GRAY1, y = 0.95)
plt.show()
```
We can see that there is a variation in the results from people who did not go to the ICU against those who had to be admitted to the ICU.
We have lower ```HEMATOCRITE```, ```LINFOCITOS``` and ```PLATELETS``` in people who went to the ICU. And higher ```LEUKOCYTES```, ```PCR```, ```POTASSIUM``` and ```UREA``` for the patients who went to the ICU.
<a name="conc"></a>
# 8 Conclusions
From the cleaning part, we see that some data preparation was need in order to get an accurate sense of the data and to obtain the right kind information for analysis.
From the data analysis part, we can clearly see that many variables correlate with the need for a patient to go to the ICU. Age, gender, vital signs and blood results can be (and will be) very indicative and useful for our future machine learning modelling in predicting ICU admission.
<a name="partwo"></a>
# Part 2 - Machine Learning
- Click [here to access part 2 in Google Colab](https://colab.research.google.com/drive/1bLzbafHgRTXe2T4fFutOzRASrsSSs2or?usp=sharing). In this next part we'll begin the ICU prediction for our problem.
- [Project Repository: ICU Prediction](https://github.com/diascarolina/project-icu-prediction)
| github_jupyter |
```
import numpy as np
import tensorflow as tf
import scipy.sparse as sp
import sys
import pickle as pkl
import networkx as nx
conv1d = tf.layers.conv1d
#注意力层
def attn_head(seq, out_sz, bias_mat, activation, in_drop=0.0, coef_drop=0.0, residual=False):
with tf.name_scope('my_attn'):
if in_drop != 0.0:
#防止过拟合,保留seq中1.0 - in_drop个数,保留的数并变为1/1.0 - in_drop
seq = tf.nn.dropout(seq, 1.0 - in_drop)
#将原始节点特征 seq 进行变换得到了 seq_fts。这里,作者使用卷积核大小为 1 的 1D 卷积模拟投影变换,
# 投影变换后的维度为 out_sz。注意,这里投影矩阵 W是所有节点共享,所以 1D 卷积中的多个卷积核也是共享的。
#seq_fts 的大小为 [num_graph, num_node, out_sz]
print('【seq】',seq)
#(1, 2708, 1433)
seq_fts = tf.layers.conv1d(seq, out_sz, 1, use_bias=False)
print('【seq_fts】',seq_fts)
#(1, 2708, 8)
# simplest self-attention possible
# f_1 和 f_2 维度均为 [num_graph, num_node, 1]
#(1, 2708, 1)
f_1 = tf.layers.conv1d(seq_fts, 1, 1) #节点投影
f_2 = tf.layers.conv1d(seq_fts, 1, 1) #邻居投影
#将 f_2 转置之后与 f_1 叠加,通过广播得到的大小为 [num_graph, num_node, num_node] 的 logits
logits = f_1 + tf.transpose(f_2, [0, 2, 1])#注意力矩阵
#(1, 2708, 2708)
print('【logits】',logits)
#+biase_mat是为了对非邻居节点mask,归一化的注意力矩阵
#邻接矩阵的作用,把和中心节点没有链接的注意力因子mask掉
coefs = tf.nn.softmax(tf.nn.leaky_relu(logits) + bias_mat)
if coef_drop != 0.0:
coefs = tf.nn.dropout(coefs, 1.0 - coef_drop)
if in_drop != 0.0:
seq_fts = tf.nn.dropout(seq_fts, 1.0 - in_drop)
#将 mask 之后的注意力矩阵 coefs 与变换后的特征矩阵 seq_fts 相乘,
# 即可得到更新后的节点表示 vals。
#(1, 2708, 2708)*(1,2708,8)=(1, 2708, 8)
vals = tf.matmul(coefs, seq_fts)
ret = tf.contrib.layers.bias_add(vals)
# residual connection
if residual:
if seq.shape[-1] != ret.shape[-1]:
ret = ret + conv1d(seq, ret.shape[-1], 1) # activation
else:
ret = ret + seq
return activation(ret) # activation
#BaseGAttN
class BaseGAttN:
def loss(logits, labels, nb_classes, class_weights):
sample_wts = tf.reduce_sum(tf.multiply(tf.one_hot(labels, nb_classes), class_weights), axis=-1)
xentropy = tf.multiply(tf.nn.sparse_softmax_cross_entropy_with_logits(
labels=labels, logits=logits), sample_wts)
return tf.reduce_mean(xentropy, name='xentropy_mean')
def training(loss, lr, l2_coef):
# weight decay
vars = tf.trainable_variables()
print(vars.shape)
lossL2 = tf.add_n([tf.nn.l2_loss(v) for v in vars if v.name not
in ['bias', 'gamma', 'b', 'g', 'beta']]) * l2_coef
# optimizer
opt = tf.train.AdamOptimizer(learning_rate=lr)
# training op
train_op = opt.minimize(loss+lossL2)
return train_op
def preshape(logits, labels, nb_classes):
new_sh_lab = [-1]
new_sh_log = [-1, nb_classes]
log_resh = tf.reshape(logits, new_sh_log)
lab_resh = tf.reshape(labels, new_sh_lab)
return log_resh, lab_resh
def confmat(logits, labels):
preds = tf.argmax(logits, axis=1)
return tf.confusion_matrix(labels, preds)
##########################
# Adapted from tkipf/gcn #
##########################
def masked_softmax_cross_entropy(logits, labels, mask):
"""Softmax cross-entropy loss with masking."""
loss = tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=labels)
mask = tf.cast(mask, dtype=tf.float32)
mask /= tf.reduce_mean(mask)
loss *= mask
return tf.reduce_mean(loss)
def masked_sigmoid_cross_entropy(logits, labels, mask):
"""Softmax cross-entropy loss with masking."""
labels = tf.cast(labels, dtype=tf.float32)
loss = tf.nn.sigmoid_cross_entropy_with_logits(logits=logits, labels=labels)
loss=tf.reduce_mean(loss,axis=1)
mask = tf.cast(mask, dtype=tf.float32)
mask /= tf.reduce_mean(mask)
loss *= mask
return tf.reduce_mean(loss)
def masked_accuracy(logits, labels, mask):
"""Accuracy with masking."""
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(labels, 1))
accuracy_all = tf.cast(correct_prediction, tf.float32)
mask = tf.cast(mask, dtype=tf.float32)
mask /= tf.reduce_mean(mask)
accuracy_all *= mask
return tf.reduce_mean(accuracy_all)
def micro_f1(logits, labels, mask):
"""Accuracy with masking."""
predicted = tf.round(tf.nn.sigmoid(logits))
# Use integers to avoid any nasty FP behaviour
predicted = tf.cast(predicted, dtype=tf.int32)
labels = tf.cast(labels, dtype=tf.int32)
mask = tf.cast(mask, dtype=tf.int32)
# expand the mask so that broadcasting works ([nb_nodes, 1])
mask = tf.expand_dims(mask, -1)
# Count true positives, true negatives, false positives and false negatives.
tp = tf.count_nonzero(predicted * labels * mask)
tn = tf.count_nonzero((predicted - 1) * (labels - 1) * mask)
fp = tf.count_nonzero(predicted * (labels - 1) * mask)
fn = tf.count_nonzero((predicted - 1) * labels * mask)
# Calculate accuracy, precision, recall and F1 score.
precision = tp / (tp + fp)
recall = tp / (tp + fn)
fmeasure = (2 * precision * recall) / (precision + recall)
fmeasure = tf.cast(fmeasure, tf.float32)
return fmeasure
#from utils import layers
#from models.base_gattn import BaseGAttN
class GAT(BaseGAttN):
def inference(inputs, nb_classes, nb_nodes, training, attn_drop, ffd_drop,
bias_mat, hid_units, n_heads, activation=tf.nn.elu, residual=False):
attns = []
#将多头输出连接在一起concat,有n个输入层
for _ in range(n_heads[0]):
attns.append(attn_head(inputs, bias_mat=bias_mat,
out_sz=hid_units[0], activation=activation,
in_drop=ffd_drop, coef_drop=attn_drop, residual=False))
#(1, 2708, 64)
h_1 = tf.concat(attns, axis=-1)
print('【hid_units】',len(hid_units))
for i in range(1, len(hid_units)):
h_old = h_1
attns = []
for _ in range(n_heads[i]):
print('【hid_units】',hid_units)
attns.append(attn_head(h_1, bias_mat=bias_mat,
out_sz=hid_units[i], activation=activation,
in_drop=ffd_drop, coef_drop=attn_drop, residual=residual))
h_1 = tf.concat(attns, axis=-1)
out = []
print('【h_1】',h_1)
for i in range(n_heads[-1]):
out.append(attn_head(h_1, bias_mat=bias_mat,
out_sz=nb_classes, activation=lambda x: x,
in_drop=ffd_drop, coef_drop=attn_drop, residual=False))
logits = tf.add_n(out) / n_heads[-1]
return logits
def load_data(dataset_str): # {'pubmed', 'citeseer', 'cora'}
"""Load data."""
names = ['x', 'y', 'tx', 'ty', 'allx', 'ally', 'graph']
objects = []
for i in range(len(names)):
with open("data/ind.{}.{}".format(dataset_str, names[i]), 'rb') as f:
if sys.version_info > (3, 0):
objects.append(pkl.load(f, encoding='latin1'))
else:
objects.append(pkl.load(f))
x, y, tx, ty, allx, ally, graph = tuple(objects)
test_idx_reorder = parse_index_file("data/ind.{}.test.index".format(dataset_str))
test_idx_range = np.sort(test_idx_reorder)
if dataset_str == 'citeseer':
# Fix citeseer dataset (there are some isolated nodes in the graph)
# Find isolated nodes, add them as zero-vecs into the right position
test_idx_range_full = range(min(test_idx_reorder), max(test_idx_reorder)+1)
tx_extended = sp.lil_matrix((len(test_idx_range_full), x.shape[1]))
tx_extended[test_idx_range-min(test_idx_range), :] = tx
tx = tx_extended
ty_extended = np.zeros((len(test_idx_range_full), y.shape[1]))
ty_extended[test_idx_range-min(test_idx_range), :] = ty
ty = ty_extended
features = sp.vstack((allx, tx)).tolil()
features[test_idx_reorder, :] = features[test_idx_range, :]
adj = nx.adjacency_matrix(nx.from_dict_of_lists(graph))
labels = np.vstack((ally, ty))
labels[test_idx_reorder, :] = labels[test_idx_range, :]
idx_test = test_idx_range.tolist()
idx_train = range(len(y))
idx_val = range(len(y), len(y)+500)
train_mask = sample_mask(idx_train, labels.shape[0])
val_mask = sample_mask(idx_val, labels.shape[0])
test_mask = sample_mask(idx_test, labels.shape[0])
y_train = np.zeros(labels.shape)
y_val = np.zeros(labels.shape)
y_test = np.zeros(labels.shape)
y_train[train_mask, :] = labels[train_mask, :]
y_val[val_mask, :] = labels[val_mask, :]
y_test[test_mask, :] = labels[test_mask, :]
print(adj.shape)
print(features.shape)
return adj, features, y_train, y_val, y_test, train_mask, val_mask, test_mask
def parse_index_file(filename):
"""Parse index file."""
index = []
for line in open(filename):
index.append(int(line.strip()))
return index
def load_random_data(size):
adj = sp.random(size, size, density=0.002) # density similar to cora
features = sp.random(size, 1000, density=0.015)
int_labels = np.random.randint(7, size=(size))
labels = np.zeros((size, 7)) # Nx7
labels[np.arange(size), int_labels] = 1
train_mask = np.zeros((size,)).astype(bool)
train_mask[np.arange(size)[0:int(size/2)]] = 1
val_mask = np.zeros((size,)).astype(bool)
val_mask[np.arange(size)[int(size/2):]] = 1
test_mask = np.zeros((size,)).astype(bool)
test_mask[np.arange(size)[int(size/2):]] = 1
y_train = np.zeros(labels.shape)
y_val = np.zeros(labels.shape)
y_test = np.zeros(labels.shape)
y_train[train_mask, :] = labels[train_mask, :]
y_val[val_mask, :] = labels[val_mask, :]
y_test[test_mask, :] = labels[test_mask, :]
# sparse NxN, sparse NxF, norm NxC, ..., norm Nx1, ...
return adj, features, y_train, y_val, y_test, train_mask, val_mask, test_mask
def sparse_to_tuple(sparse_mx):
"""Convert sparse matrix to tuple representation."""
def to_tuple(mx):
if not sp.isspmatrix_coo(mx):
mx = mx.tocoo()
coords = np.vstack((mx.row, mx.col)).transpose()
values = mx.data
shape = mx.shape
return coords, values, shape
if isinstance(sparse_mx, list):
for i in range(len(sparse_mx)):
sparse_mx[i] = to_tuple(sparse_mx[i])
else:
sparse_mx = to_tuple(sparse_mx)
return sparse_mx
def sample_mask(idx, l):
"""Create mask."""
mask = np.zeros(l)
mask[idx] = 1
return np.array(mask, dtype=np.bool)
def standardize_data(f, train_mask):
"""Standardize feature matrix and convert to tuple representation"""
# standardize data
f = f.todense()
mu = f[train_mask == True, :].mean(axis=0)
sigma = f[train_mask == True, :].std(axis=0)
f = f[:, np.squeeze(np.array(sigma > 0))]
mu = f[train_mask == True, :].mean(axis=0)
sigma = f[train_mask == True, :].std(axis=0)
f = (f - mu) / sigma
return f
def preprocess_features(features):
"""Row-normalize feature matrix and convert to tuple representation"""
rowsum = np.array(features.sum(1))
r_inv = np.power(rowsum, -1).flatten()
r_inv[np.isinf(r_inv)] = 0.
r_mat_inv = sp.diags(r_inv)
features = r_mat_inv.dot(features)
return features.todense(), sparse_to_tuple(features)
def adj_to_bias(adj, sizes, nhood=1):
nb_graphs = adj.shape[0]
mt = np.empty(adj.shape)
for g in range(nb_graphs):
mt[g] = np.eye(adj.shape[1])
for _ in range(nhood):
mt[g] = np.matmul(mt[g], (adj[g] + np.eye(adj.shape[1])))
for i in range(sizes[g]):
for j in range(sizes[g]):
if mt[g][i][j] > 0.0:
mt[g][i][j] = 1.0
return -1e9 * (1.0 - mt)
print('Dataset: ' + dataset)
print('----- Opt. hyperparams -----')
print('lr: ' + str(lr))
print('l2_coef: ' + str(l2_coef))
print('----- Archi. hyperparams -----')
print('nb. layers: ' + str(len(hid_units)))
print('nb. units per layer: ' + str(hid_units))
print('nb. attention heads: ' + str(n_heads))
print('residual: ' + str(residual))
print('nonlinearity: ' + str(nonlinearity))
print('model: ' + str(model))
adj, features, y_train, y_val, y_test, train_mask, val_mask, test_mask = load_data(dataset)
features, spars = preprocess_features(features)
nb_nodes = features.shape[0]
ft_size = features.shape[1]
nb_classes = y_train.shape[1]
adj = adj.todense()
features = features[np.newaxis]
adj = adj[np.newaxis]
y_train = y_train[np.newaxis]
y_val = y_val[np.newaxis]
y_test = y_test[np.newaxis]
train_mask = train_mask[np.newaxis]
val_mask = val_mask[np.newaxis]
test_mask = test_mask[np.newaxis]
biases = adj_to_bias(adj, [nb_nodes], nhood=1)
with tf.Graph().as_default():
with tf.name_scope('input'):
ftr_in = tf.placeholder(dtype=tf.float32, shape=(batch_size, nb_nodes, ft_size))
bias_in = tf.placeholder(dtype=tf.float32, shape=(batch_size, nb_nodes, nb_nodes))
lbl_in = tf.placeholder(dtype=tf.int32, shape=(batch_size, nb_nodes, nb_classes))
msk_in = tf.placeholder(dtype=tf.int32, shape=(batch_size, nb_nodes))
attn_drop = tf.placeholder(dtype=tf.float32, shape=())
ffd_drop = tf.placeholder(dtype=tf.float32, shape=())
is_train = tf.placeholder(dtype=tf.bool, shape=())
logits = model.inference(ftr_in, nb_classes, nb_nodes, is_train,
attn_drop, ffd_drop,
bias_mat=bias_in,
hid_units=hid_units, n_heads=n_heads,
residual=residual, activation=nonlinearity)
log_resh = tf.reshape(logits, [-1, nb_classes])
lab_resh = tf.reshape(lbl_in, [-1, nb_classes])
msk_resh = tf.reshape(msk_in, [-1])
loss = model.masked_softmax_cross_entropy(log_resh, lab_resh, msk_resh)
accuracy = model.masked_accuracy(log_resh, lab_resh, msk_resh)
train_op = model.training(loss, lr, l2_coef)
saver = tf.train.Saver()
init_op = tf.group(tf.global_variables_initializer(), tf.local_variables_initializer())
vlss_mn = np.inf
vacc_mx = 0.0
curr_step = 0
with tf.Session() as sess:
sess.run(init_op)
train_loss_avg = 0
train_acc_avg = 0
val_loss_avg = 0
val_acc_avg = 0
for epoch in range(nb_epochs):
tr_step = 0
tr_size = features.shape[0]
while tr_step * batch_size < tr_size:
_, loss_value_tr, acc_tr = sess.run([train_op, loss, accuracy],
feed_dict={
ftr_in: features[tr_step*batch_size:(tr_step+1)*batch_size],
bias_in: biases[tr_step*batch_size:(tr_step+1)*batch_size],
lbl_in: y_train[tr_step*batch_size:(tr_step+1)*batch_size],
msk_in: train_mask[tr_step*batch_size:(tr_step+1)*batch_size],
is_train: True,
attn_drop: 0.6, ffd_drop: 0.6})
train_loss_avg += loss_value_tr
train_acc_avg += acc_tr
tr_step += 1
vl_step = 0
vl_size = features.shape[0]
while vl_step * batch_size < vl_size:
loss_value_vl, acc_vl = sess.run([loss, accuracy],
feed_dict={
ftr_in: features[vl_step*batch_size:(vl_step+1)*batch_size],
bias_in: biases[vl_step*batch_size:(vl_step+1)*batch_size],
lbl_in: y_val[vl_step*batch_size:(vl_step+1)*batch_size],
msk_in: val_mask[vl_step*batch_size:(vl_step+1)*batch_size],
is_train: False,
attn_drop: 0.0, ffd_drop: 0.0})
val_loss_avg += loss_value_vl
val_acc_avg += acc_vl
vl_step += 1
print('Training: loss = %.5f, acc = %.5f | Val: loss = %.5f, acc = %.5f' %
(train_loss_avg/tr_step, train_acc_avg/tr_step,
val_loss_avg/vl_step, val_acc_avg/vl_step))
if val_acc_avg/vl_step >= vacc_mx or val_loss_avg/vl_step <= vlss_mn:
if val_acc_avg/vl_step >= vacc_mx and val_loss_avg/vl_step <= vlss_mn:
vacc_early_model = val_acc_avg/vl_step
vlss_early_model = val_loss_avg/vl_step
#saver.save(sess, checkpt_file)
vacc_mx = np.max((val_acc_avg/vl_step, vacc_mx))
vlss_mn = np.min((val_loss_avg/vl_step, vlss_mn))
curr_step = 0
else:
curr_step += 1
if curr_step == patience:
print('Early stop! Min loss: ', vlss_mn, ', Max accuracy: ', vacc_mx)
print('Early stop model validation loss: ', vlss_early_model, ', accuracy: ', vacc_early_model)
break
train_loss_avg = 0
train_acc_avg = 0
val_loss_avg = 0
val_acc_avg = 0
#saver.restore(sess, checkpt_file)
ts_size = features.shape[0]
ts_step = 0
ts_loss = 0.0
ts_acc = 0.0
while ts_step * batch_size < ts_size:
loss_value_ts, acc_ts = sess.run([loss, accuracy],
feed_dict={
ftr_in: features[ts_step*batch_size:(ts_step+1)*batch_size],
bias_in: biases[ts_step*batch_size:(ts_step+1)*batch_size],
lbl_in: y_test[ts_step*batch_size:(ts_step+1)*batch_size],
msk_in: test_mask[ts_step*batch_size:(ts_step+1)*batch_size],
is_train: False,
attn_drop: 0.0, ffd_drop: 0.0})
ts_loss += loss_value_ts
ts_acc += acc_ts
ts_step += 1
print('Test loss:', ts_loss/ts_step, '; Test accuracy:', ts_acc/ts_step)
sess.close()
```
| github_jupyter |
# Using DALI in PyTorch Lightning
### Overview
This example shows how to use DALI in PyTorch Lightning.
Let us grab [a toy example](https://pytorch-lightning.readthedocs.io/en/latest/introduction_guide.html) showcasing a classification network and see how DALI can accelerate it.
The DALI_EXTRA_PATH environment variable should point to a [DALI extra](https://github.com/NVIDIA/DALI_extra) copy. Please make sure that the proper release tag, the one associated with your DALI version, is checked out.
```
import torch
from torch.nn import functional as F
from torch import nn
from pytorch_lightning.core.lightning import LightningModule
from pytorch_lightning import Trainer
from torch.optim import Adam
from torchvision.datasets import MNIST
from torchvision import datasets, transforms
from torch.utils.data import DataLoader
import os
BATCH_SIZE = 64
# workaround for https://github.com/pytorch/vision/issues/1938 - error 403 when downloading mnist dataset
import urllib
opener = urllib.request.build_opener()
opener.addheaders = [('User-agent', 'Mozilla/5.0')]
urllib.request.install_opener(opener)
```
We will start by implement a training class that uses the native data loader
```
class LitMNIST(LightningModule):
def __init__(self):
super().__init__()
# mnist images are (1, 28, 28) (channels, width, height)
self.layer_1 = torch.nn.Linear(28 * 28, 128)
self.layer_2 = torch.nn.Linear(128, 256)
self.layer_3 = torch.nn.Linear(256, 10)
def forward(self, x):
batch_size, channels, width, height = x.size()
# (b, 1, 28, 28) -> (b, 1*28*28)
x = x.view(batch_size, -1)
x = self.layer_1(x)
x = F.relu(x)
x = self.layer_2(x)
x = F.relu(x)
x = self.layer_3(x)
x = F.log_softmax(x, dim=1)
return x
def process_batch(self, batch):
return batch
def training_step(self, batch, batch_idx):
x, y = self.process_batch(batch)
logits = self(x)
loss = F.nll_loss(logits, y)
return loss
def cross_entropy_loss(self, logits, labels):
return F.nll_loss(logits, labels)
def configure_optimizers(self):
return Adam(self.parameters(), lr=1e-3)
def prepare_data(self):# transforms for images
transform=transforms.Compose([transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))])
self.mnist_train = MNIST(os.getcwd(), train=True, download=True, transform=transform)
def train_dataloader(self):
return DataLoader(self.mnist_train, batch_size=64, num_workers=8, pin_memory=True)
```
And see how it works
```
model = LitMNIST()
trainer = Trainer(gpus=1, distributed_backend="ddp", max_epochs=5)
trainer.fit(model)
```
The next step is to define a DALI pipeline that will be used for loading and pre-processing data.
```
import nvidia.dali as dali
from nvidia.dali import pipeline_def
import nvidia.dali.fn as fn
import nvidia.dali.types as types
from nvidia.dali.plugin.pytorch import DALIClassificationIterator, LastBatchPolicy
# Path to MNIST dataset
data_path = os.path.join(os.environ['DALI_EXTRA_PATH'], 'db/MNIST/training/')
@pipeline_def
def GetMnistPipeline(device, shard_id=0, num_shards=1):
jpegs, labels = fn.readers.caffe2(path=data_path, shard_id=shard_id, num_shards=num_shards, random_shuffle=True, name="Reader")
images = fn.decoders.image(jpegs,
device='mixed' if device == 'gpu' else 'cpu',
output_type=types.GRAY)
images = fn.crop_mirror_normalize(images,
dtype=types.FLOAT,
std=[0.3081 * 255],
mean=[0.1307 * 255],
output_layout="CHW")
if device == "gpu":
labels = labels.gpu()
# PyTorch expects labels as INT64
labels = fn.cast(labels, dtype=types.INT64)
return images, labels
```
Now we are ready to modify the training class to use the DALI pipeline we have just defined. Because we want to integrate with PyTorch, we wrap our pipeline with a PyTorch DALI iterator, that can replace the native data loader with some minor changes in the code. The DALI iterator returns a list of dictionaries, where each element in the list corresponds to a pipeline instance, and the entries in the dictionary map to the outputs of the pipeline. For more information, check the documentation of DALIGenericIterator.
```
class DALILitMNIST(LitMNIST):
def __init__(self):
super().__init__()
def prepare_data(self):
device_id = self.local_rank
shard_id = self.global_rank
num_shards = self.trainer.world_size
mnist_pipeline = GetMnistPipeline(batch_size=BATCH_SIZE, device='gpu', device_id=device_id, shard_id=shard_id,
num_shards=num_shards, num_threads=8)
self.train_loader = DALIClassificationIterator(mnist_pipeline, reader_name="Reader",
last_batch_policy=LastBatchPolicy.PARTIAL, auto_reset=True)
def train_dataloader(self):
return self.train_loader
def process_batch(self, batch):
x = batch[0]["data"]
y = batch[0]["label"].squeeze(-1)
return (x, y)
```
We can now run the training
```
# Even if previous Trainer finished his work it still keeps the GPU booked, force it to release the device.
if 'PL_TRAINER_GPUS' in os.environ:
os.environ.pop('PL_TRAINER_GPUS')
model = DALILitMNIST()
trainer = Trainer(gpus=1, distributed_backend="ddp", max_epochs=5)
trainer.fit(model)
```
For even better integration, we can provide a custom DALI iterator wrapper so that no extra processing is required inside `LitMNIST.process_batch`. Also, PyTorch can learn the size of the dataset this way.
```
class BetterDALILitMNIST(LitMNIST):
def __init__(self):
super().__init__()
def prepare_data(self):
device_id = self.local_rank
shard_id = self.global_rank
num_shards = self.trainer.world_size
mnist_pipeline = GetMnistPipeline(batch_size=BATCH_SIZE, device='gpu', device_id=device_id, shard_id=shard_id, num_shards=num_shards, num_threads=8)
class LightningWrapper(DALIClassificationIterator):
def __init__(self, *kargs, **kvargs):
super().__init__(*kargs, **kvargs)
def __next__(self):
out = super().__next__()
# DDP is used so only one pipeline per process
# also we need to transform dict returned by DALIClassificationIterator to iterable
# and squeeze the lables
out = out[0]
return [out[k] if k != "label" else torch.squeeze(out[k]) for k in self.output_map]
self.train_loader = LightningWrapper(mnist_pipeline, reader_name="Reader", last_batch_policy=LastBatchPolicy.PARTIAL, auto_reset=True)
def train_dataloader(self):
return self.train_loader
```
Let us run the training one more time
```
# Even if previous Trainer finished his work it still keeps the GPU booked, force it to release the device.
if 'PL_TRAINER_GPUS' in os.environ:
os.environ.pop('PL_TRAINER_GPUS')
model = BetterDALILitMNIST()
trainer = Trainer(gpus=1, distributed_backend="ddp", max_epochs=5)
trainer.fit(model)
```
| github_jupyter |
[Previous Notebook](Part_2.ipynb)
     
     
     
     
[Home Page](../Start_Here.ipynb)
# CNN Primer and Keras 101 - Continued
This notebook covers introduction to Convolutional Neural Networks, and it's terminologies.
**Contents of the this Notebook:**
- [Convolution Neural Networks ( CNNs )](#Convolution-Neural-Networks-(-CNNs-))
- [Why CNNs are good in Image related tasks? ](#Why-CNNs-are-good-in-Image-related-tasks?)
- [Implementing Image Classification using CNN's](#Implementing-Image-Classification-using-CNN's)
- [Conclusion](#Conclusion-:)
**By the end of this notebook you will:**
- Understand how a Convolution Neural Network works
- Write your own CNN Classifier and train it.
## Convolution Neural Networks ( CNNs )
Convolution Neural Networks are widely used in the field of Image Classification, Object Detection, and Face Recognition because they are very effective in reducing the number of parameters without losing on the quality of models.
Let's now understand what makes up a CNN Architecture and how it works :
Here is an example of a CNN Architecture for a Classification task :

*Source: https://fr.mathworks.com/solutions/deep-learning/convolutional-neural-network.html*
Each input image will pass it through a series of convolution layers with filters (Kernels), pooling, fully connected layers (FC) and apply Softmax function to classify an object with probabilistic values between 0 and 1.
Let us discuss in brief about the following in detail :
- Convolution Layer
- Strides and Padding
- Pooling Layer
- Fully Connected Layer
#### Convolution Layer :
Convolution layer is the first layer to learn features from the input by preserving the relationships between neighbouring pixels. The Kernel Size is a Hyper-parameter and can be altered according to the complexity of the problem.
Now that we've discussed Kernels. Let's see how a Kernel operates on the layer.

*Source: https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53*
We have seen how the convolution operation works, and now let us now see how convolution operation is carried out with multiple layers.

*Source: https://towardsdatascience.com/a-comprehensive-introduction-to-different-types-of-convolutions-in-deep-learning-669281e58215*
Let us define the terms :
- Hin : Height dimension of the layer
- Win : Width dimension of the layer
- Din : Depth of the layer
- h : height of the kernel
- w : width of the kernal
- Dout : Number of kernels acting on the Layer
Note : Din for the Layer and Kernel needs to be the same.
Here the Din and Dout is also called as the number of channels of the layer. We can notice from the first image that typically the number of channels keeps increasing over the layers while the height and width keep decreasing. This is done so that the filters learn the features from the previous layers, they can also be called as feature channels.
#### Strides and Padding
Stride is the number of pixels shifts over the input matrix during convolution. When the stride is 1, then we move the filters to 1 pixel at a time. When the stride is 2, then we move the filters to 2 pixels at a time and so on.
Sometimes filter do not fit perfectly on the input image. So, we have two options:
- Pad the picture with zeros (zero-padding) so that it fits
- Drop the part of the image where the filter did not fit. This is called valid padding which keeps only the valid part of the image.
#### Pooling Layer :
Pooling layers section would reduce the number of parameters when the images are too large. Spatial pooling also called subsampling or downsampling, which reduces the dimensionality of each map but retains important information. Spatial pooling can be of different types:
- Max Pooling :
- Max pooling is one of the common pooling used, and it takes the largest element from the rectified feature map.
- Average Pooling
- Taking the average of the elements is called Average pooling.
- Sum Pooling
- Sum of all elements in the feature map call is called as sum pooling.

*Source: https://www.programmersought.com/article/47163598855/*
#### Fully Connected Layer :
We will then flatten the output from the convolutions layers and feed into it a _Fully Connected layer_ to generate a prediction. The fully connected layer is an ANN Model whose inputs are the features of the Inputs obtained from the Convolutions Layers.
These Fully Connected Layers are then trained along with the _kernels_ during the training process.
We will also be comparing later between CNN's and ANN's during our example to benchmark their results on Image Classification tasks.
### Transposed Convolution :
When we apply our Convolution operation over an image, we find that the number of channels increase while the height and width of the image decreases, now in some cases, for different applications we will need to up-sample our images, _Transposed convolution_ helps to up sample the images from these layers.
Here is an animation to Tranposed convolution:
<table><tr>
<td> <img src="images/convtranspose.gif" alt="Drawing" style="width: 540px;"/></td>
<td> <img src="images/convtranspose_conv.gif" alt="Drawing" style="width: 500px;"/> </td>
</tr></table>
*Source https://towardsdatascience.com/a-comprehensive-introduction-to-different-types-of-convolutions-in-deep-learning-669281e58215*
Tranposed Convolution can also be visualised as Convolution of a Layer with 2x2 padding as displayed in the right gif.
## Why CNNs are good in Image related tasks?
1970, **David Marr** wrote a book called [vision](https://mitpress.mit.edu/books/vision).It was a breakthrough in understating of how the brain does vision; he stated that vision task is performed in a hierarchal manner. You start simple and get complex. For example, you start with as simple as identifying edge, colours and then build upon them to detect the object and then classify them and so on.
The architecture of CNNs is designed as such to emulate the human brain's technique to deal with images. As convolutions are mainly used for extracting high-level features from the images such as edges/other patterns, these algorithms try to emulate our understanding of the vision. Certain filters do operations such as blurring the image, sharpening the image and then performing pooling operations on each of these filters to extract information from an image. As stated earlier, our understanding of vision consists that vision is a hierarchal process, and our brain deals with vision in a similar fashion. CNN also deals with understanding and classifying images similarly, thereby making them the appropriate choice for these kinds of tasks.
# Implementing Image Classification using CNN's
We will the following the same steps for Data Pre-processing as mentioned in the previous Notebook :
```
# Import Necessary Libraries
from __future__ import absolute_import, division, print_function, unicode_literals
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
# Let's Import the Dataset
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
#Print Array Size of Training Set
print("Size of Training Images :"+str(train_images.shape))
#Print Array Size of Label
print("Size of Training Labels :"+str(train_labels.shape))
#Print Array Size of Test Set
print("Size of Test Images :"+str(test_images.shape))
#Print Array Size of Label
print("Size of Test Labels :"+str(test_labels.shape))
#Let's See how our Outputs Look like
print("Training Set Labels :"+str(train_labels))
#Data in the Test Set
print("Test Set Labels :"+str(test_labels))
train_images = train_images / 255.0
test_images = test_images / 255.0
```
## Further Data pre-processing :
You may have noticed by now that the Training Set is of Shape `(60000,28,28)`.
In CNN's, we need to feed the data in the form of a 4D Array as follows :
`( Num_Images, X-dims, Y-dims, # of Channels of Image )`
So, as our image is grayscale, we will reshape it to `(60000,28,28,1)` before passing it to our Architecture.
```
# Reshape input data from (28, 28) to (28, 28, 1)
w, h = 28, 28
train_images = train_images.reshape(train_images.shape[0], w, h, 1)
test_images = test_images.reshape(test_images.shape[0], w, h, 1)
```
## Defining Convolution Layers
Let us see how to define a Convolution Layer, MaxPooling Layer and Dropout
#### Convolution Layer
We will be using the following API to define the Convolution Layer.
```tf.keras.layers.Conv2D(filters, kernel_size, padding='valid', activation=None, input_shape)```
Let us define the parameters in brief :
- Filters: The dimensionality of the output space (i.e. the number of output filters in the convolution).
- Kernel_size: An integer or tuple/list of 2 integers, specifying the height and width of the 2D convolution window. Can be a single integer to specify the same value for all spatial dimensions.
- Padding: one of "valid" or "same" (case-insensitive).
- Activation: Activation function to use (see activations). If you don't specify anything, no activation is applied (ie. "linear" activation: a(x) = x).
Refer here for the Full Documentation -> [Convolutional Layers](https://keras.io/layers/convolutional/)
#### Pooling Layer
`tf.keras.layers.MaxPooling2D(pool_size=2)`
- Pool size : Size of the max pooling windows.
Keras Documentation -> [Pooling Layers](https://keras.io/layers/pooling/)
#### Dropout
Dropout is an approach to regularization in neural networks which helps reducing interdependent learning amongst the neurons.
Simply put, dropout refers to ignoring units (i.e. neurons) during the training phase of certain set of neurons which is chosen at random. By “ignoring”, we mean these units are not considered during a particular forward or backward pass.
It is defined by the following function :
`tf.keras.layers.Dropout(0.3)`
- Parameter : float between 0 and 1. Fraction of the input units to drop.
Keras Documentation -> [Dropout](https://keras.io/layers/core/#dropout)
## Defining our Model and Training
Now that we are aware of the code for building a CNN , Let us now build a 5 Layer Model :
- Input Layer : ( 28 , 28 ,1 )
- Size of the Input Image
- Convolution layers :
- First Layer : Kernel Size ( 2x2 ) and we obtain 64 layers from it.
- Pooling of Size ( 2 x 2) making the layer to be ( 14 x 14 x 64 )
- Second Layer : Kernel Size ( 2 x 2 ) and obtaining 32 layers.
- Pooling of Size ( 2 x 2 ) making the layer to be ( 7 x 7 x 32 )
- Fully Connected Layers :
- Flatten the Convolution layers to nodes of 1567 = ( 7 * 7 * 32 )
- Dense Layer of 256
- Output Layer :
- Densely Connected Layer with 10 classes with `softmax` activation

Here , Let us now define our Model in Keras
```
from tensorflow.keras import backend as K
import tensorflow as tf
K.clear_session()
model = tf.keras.Sequential()
# Must define the input shape in the first layer of the neural network
model.add(tf.keras.layers.Conv2D(filters=64, kernel_size=2, padding='same', activation='relu', input_shape=(28,28,1)))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
# model.add(tf.keras.layers.Dropout(0.3))
#Second Convolution Layer
model.add(tf.keras.layers.Conv2D(filters=32, kernel_size=2, padding='same', activation='relu'))
model.add(tf.keras.layers.MaxPooling2D(pool_size=2))
model.add(tf.keras.layers.Dropout(0.3))
#Fully Connected Layer
model.add(tf.keras.layers.Flatten())
model.add(tf.keras.layers.Dense(256, activation='relu'))
model.add(tf.keras.layers.Dropout(0.5))
model.add(tf.keras.layers.Dense(10, activation='softmax'))
# Take a look at the model summary
model.summary()
```
### Compile the model
Before the model is ready for training, it needs a few more settings. These are added during the model's *compile* step:
* *Loss function* —This measures how accurate the model is during training. You want to minimize this function to "steer" the model in the right direction.
* *Optimizer* —This is how the model is updated based on the data it sees and its loss function.
* *Metrics* —Used to monitor the training and testing steps. The following example uses *accuracy*, the fraction of the images that are correctly classified.
```
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
```
## Train the model
Training the neural network model requires the following steps:
1. Feed the training data to the model. In this example, the training data is in the `train_images` and `train_labels` arrays.
2. The model learns to associate images and labels.
3. You ask the model to make predictions about a test set—in this example, the `test_images` array. Verify that the predictions match the labels from the `test_labels` array.
To start training, call the `model.fit` method—so called because it "fits" the model to the training data:
```
model.fit(train_images, train_labels,batch_size=32 ,epochs=5)
#Evaluating the Model using the Test Set
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('\nTest accuracy:', test_acc)
```
## Making Predictions :
```
# Making Predictions from the test_images
predictions = model.predict(test_images)
# Reshape input data from (28, 28) to (28, 28, 1)
w, h = 28, 28
train_images = train_images.reshape(train_images.shape[0], w, h)
test_images = test_images.reshape(test_images.shape[0], w, h)
# Helper Functions to Plot Images
def plot_image(i, predictions_array, true_label, img):
predictions_array,true_label, img = predictions_array, true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'blue'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
def plot_value_array(i, predictions_array, true_label):
predictions_array, true_label = predictions_array, true_label[i]
plt.grid(False)
plt.xticks(range(10))
plt.yticks([])
thisplot = plt.bar(range(10), predictions_array, color="#777777")
plt.ylim([0, 1])
predicted_label = np.argmax(predictions_array)
thisplot[predicted_label].set_color('red')
thisplot[true_label].set_color('blue')
# Plot the first X test images, their predicted labels, and the true labels.
# Color correct predictions in blue and incorrect predictions in red.
num_rows = 5
num_cols = 3
num_images = num_rows*num_cols
plt.figure(figsize=(2*2*num_cols, 2*num_rows))
for i in range(num_images):
plt.subplot(num_rows, 2*num_cols, 2*i+1)
plot_image(i, predictions[i], test_labels, test_images)
plt.subplot(num_rows, 2*num_cols, 2*i+2)
plot_value_array(i, predictions[i], test_labels)
plt.tight_layout()
plt.show()
```
### Conclusion :
Running both our models for 5 Epochs here is a table comparing them :
| Model | Train Accuracy | Train Loss | Test Accuracy | Test Loss |
|----------|-----------------|-------------|---------------|-----------|
| Fully connected Neural Networks -After 5 Epochs | 0.8923 | 0.2935 | 0.8731 | 0.2432|
| Convolution networks - After 5 Epochs | 0.8860| 0.3094 | 0.9048 | 0.1954 |
Congrats on coming this far, now that you are introduced to Machine Learning and Deep Learning, you can get started on the Domain Specific Problem accessible through the Home Page.
## Exercise
Play with different hyper-parameters ( Epoch, depth of layers , kernel size ... ) to bring down loss further
## Important:
<mark>Shutdown the kernel before clicking on “Next Notebook” to free up the GPU memory.</mark>
## Acknowledgements :
[Transposed Convolutions explained](https://medium.com/apache-mxnet/transposed-convolutions-explained-with-ms-excel-52d13030c7e8)
[Why are CNNs used more for computer vision tasks than other tasks?](https://www.quora.com/Why-are-CNNs-used-more-for-computer-vision-tasks-than-other-tasks)
[Comprehensive introduction to Convolution](https://towardsdatascience.com/a-comprehensive-introduction-to-different-types-of-convolutions-in-deep-learning-669281e58215)
## Licensing
This material is released by OpenACC-Standard.org, in collaboration with NVIDIA Corporation, under the Creative Commons Attribution 4.0 International (CC BY 4.0).
[Previous Notebook](Part_2.ipynb)
     
     
     
     
[Home Page](../Start_Here.ipynb)
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
data = np.genfromtxt('quasar_train.csv', delimiter=',')
lambd = data[0, :]
m = lambd.shape[0]
train_set = data[1:, :]
test_set = np.genfromtxt('quasar_test.csv', delimiter=',')[1:, :]
num_train = train_set.shape[0]
num_test = test_set.shape[0]
lambd_add_bias = np.vstack(
[np.ones(lambd.shape), lambd]).T
first_sample_x = train_set[0, :]
smooth_train_set = np.load('smooth_train_set.npy')
smooth_test_set = np.load('smooth_test_set.npy')
def weight_matrix(index, tau):
return np.diag(np.exp(-np.square(lambd - lambd[index]) / (2 * tau ** 2)))
def local_weight_LR(sample):
y_hat = np.zeros((m,))
tau = 5
for i in range(m):
Wi = weight_matrix(i, tau)
theta_i = np.linalg.inv(lambd_add_bias.T.dot(Wi).dot(lambd_add_bias)) \
.dot(np.dot(lambd_add_bias.T.dot(Wi), sample))
y_hat[i] = theta_i[0] + lambd[i] * theta_i[1]
return y_hat
def distance_matrix(dataset):
num = dataset.shape[0]
matrix = np.zeros((num, num_train))
for i in range(num):
matrix[i, :] = np.linalg.norm(smooth_train_set - dataset[i], axis=1)
return matrix / np.amax(matrix, axis=1, keepdims=True)
def neighb(k=3, dataset='train'):
if dataset == 'train':
matrix = distance_matrix(smooth_train_set)
k_index = np.argpartition(matrix, range(1, k+1), axis=1)[:, 1:(k + 1)]
ker = 1. - np.partition(matrix, range(1, k+1), axis=1)[:, 1:(k + 1)]
else:
matrix = distance_matrix(smooth_test_set)
k_index = np.argpartition(matrix, k, axis=1)[:, :k]
ker = 1. - np.partition(matrix, k, axis=1)[:, :k]
return k_index, ker
right_trains = smooth_train_set[:, 150:]
left_trains = smooth_train_set[:, :50]
right_tests = smooth_test_set[:, 150:]
left_tests = smooth_test_set[:, :50]
k_neighb_index, ker = neighb()
f_left_estimates = np.zeros_like(left_trains)
for i in range(num_train):
f_left_estimates[i] = np.sum(ker[i][:, np.newaxis] * left_trains[k_neighb_index[i]], axis=0) / np.sum(ker[i])
error_train = np.sum((f_left_estimates - left_trains) ** 2)
print(error_train / num_train)
k_test_index, ker_test = neighb(dataset='test')
f_left_estimates_test = np.zeros_like(left_tests)
for i in range(num_test):
f_left_estimates_test[i] = np.sum(ker[i][:, np.newaxis] * left_trains[k_neighb_index[i]], axis=0) / np.sum(ker[i])
error_train = np.sum((f_left_estimates_test - left_tests) ** 2)
print(error_train / num_test)
estimates_stack = np.hstack([f_left_estimates_test, smooth_test_set[:, 50:]])
plt.plot(lambd, smooth_test_set[0])
plt.plot(range(1150, 1200), f_left_estimates_test[0])
index = np.partition(a, [1,2,3], axis=1)[:,1:4]
index
np.maximum(1 - a, 0)
neighb(3)[:10, :4]
mt = distance_matrix(smooth_train_set)
mt[1, 1]
```
| github_jupyter |
# Training to label with BERT and Cleanlab
In this notebook, we'll use the labels that we have generated and checked with [Cleanlab](https://github.com/cgnorthcutt/cleanlab) to build a classifier using BERT embeddings to label the remaining data. We'll check our results with Cleanlab and adjust as necessary.
```
%load_ext autoreload
%autoreload 2
%matplotlib inline
import warnings
warnings.simplefilter(action='ignore', category=FutureWarning)
import matplotlib.pyplot as plt
from tqdm import tqdm
from random import sample
from sklearn.model_selection import train_test_split
import numpy as np
import cleanlab
from sklearn.linear_model import LogisticRegression
from tensorflow.keras import models, layers
import pandas as pd
import torch
from transformers import BertTokenizer, BertModel
import os
import sys
import inspect
from pathlib import Path
currentdir = Path.cwd()
parentdir = os.path.dirname(currentdir)
sys.path.insert(0,parentdir)
from mlyoucanuse.embeddings import get_embeddings_index
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
occs_df = pd.read_csv('occupations.wikidata.all.gnews.labeled.final.csv', sep='\t')
occs_df.head()
pos_eg = occs_df.query('label==1')['occupation'].tolist()
neg_eg = occs_df.query('label==0')['occupation'].tolist()
print(f"Positive examples: {len(pos_eg):,}; Negative examples {len(neg_eg):,}")
tokenizer = BertTokenizer.from_pretrained('bert-large-cased')
model = BertModel.from_pretrained('bert-large-cased')
model.eval()
model.to(device)
device
```
## Here's an example of how we'll use BERT to provide embeddings for the text of an occupation label
```
input_ids = torch.tensor(tokenizer.encode("The quick brown fox jumped over the lazy dog.")).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
last_hidden_states.detach()
print(last_hidden_states.shape)
text_embedding = last_hidden_states[0][0]
print(text_embedding.shape)
```
# Generate X; BERT embeddings for each Occupation we labeled
```
X =[]
with torch.no_grad():
for name in tqdm(pos_eg, total=len(pos_eg)):
input_ids = torch.tensor(tokenizer.encode(name)).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
X.append(last_hidden_states[0][0].detach().cpu().numpy())
for name in tqdm(neg_eg, total=len(neg_eg)):
input_ids = torch.tensor(tokenizer.encode(name)).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
X.append(last_hidden_states[0][0].detach().cpu().numpy())
X = np.array(X)
X.shape
unlabeled = occs_df.query("label==-1")['occupation'].tolist()
unlabeled_X = []
with torch.no_grad():
for name in tqdm(unlabeled, total=len(unlabeled)):
input_ids = torch.tensor(tokenizer.encode(name)).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
unlabeled_X.append(last_hidden_states[0][0].detach().cpu().numpy())
unlabeled_X = np.array(unlabeled_X)
unlabeled_X.shape
```
## Split our data into train, validation and test sets
```
y = np.concatenate([np.ones(len(pos_eg), dtype=np.int64), np.zeros(len(neg_eg), dtype=np.int64)])
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=.8, random_state=12)
X_test, X_validation, y_test, y_validation = train_test_split(X_test, y_test, train_size=.5, random_state=12)
X.shape, y.shape
```
## Build a simple model
```
model = models.Sequential([
layers.Dense(512, input_shape=(1024,), activation='relu'),
layers.Dropout(.2),
layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(X_train,
y_train,
epochs=20,
batch_size=128,
validation_data=(X_validation, y_validation),
verbose=0)
```
## Evaluate and adjust as necessary
```
res = model.evaluate(X_train, y_train)
print(f"Train results: loss {res[0]:.3f} acc: {res[1]:.3f}")
res = model.evaluate(X_validation, y_validation)
print(f"Validation results: loss {res[0]:.3f} acc: {res[1]:.3f}")
res = model.evaluate(X_test, y_test)
print(f"Test results: loss {res[0]:.3f} acc: {res[1]:.3f}")
history_dict = history.history
loss_values = history_dict['loss']
val_loss_values = history_dict['val_loss']
epochs = range(1, len(loss_values) + 1)
plt.plot(epochs, loss_values, 'bo', label='Training loss')
plt.plot(epochs, val_loss_values, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.show()
```
## Train final model
```
final_train_x = np.vstack((X_train, X_test))
final_train_y = np.hstack((y_train, y_test))
model = models.Sequential([
layers.Dense(512, input_shape=(1024,), activation='relu'),
layers.Dropout(0.2),
layers.Dense(1, activation='sigmoid')
])
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
history = model.fit(final_train_x,
final_train_y,
epochs=7,
batch_size=128,
validation_data=(X_validation, y_validation),
verbose=0)
```
## Make Predictions
```
predictions = {}
for idx, word in tqdm(enumerate(unlabeled)):
res = model.predict(unlabeled_X[idx].reshape(1, -1))
predictions[word] = res[0][0]
```
## Update our dataset with the BERT predictions
```
for occ in tqdm(predictions, total=len(predictions)):
if occ in occs_df['occupation'].tolist():
the_row = occs_df[occs_df['occupation'] == occ]
the_index = the_row.index[0]
occs_df.loc[the_index, 'label'] = int(round(predictions[occ]))
occs_df.loc[the_index, 'labeled_by'] = 'bert'
```
## Use Cleanlab to find potentially problematic labels
* Get out-of-sample predicted probabilities using cross-validation
* Compute confident joint
* Find label errors
## Step 1: Get out-of-sample predicted probabilities using cross-validation
```
labeled_y = [int(round(val)) for key,val in predictions.items() ]
all_X = np.vstack((X, unlabeled_X))
all_y = np.hstack((y, labeled_y))
# using a simple, non-optimized logistic regression classifier; the cross-validation will expose label weaknesses
psx = cleanlab.latent_estimation.estimate_cv_predicted_probabilities(
all_X, all_y, clf=LogisticRegression(max_iter=1000, multi_class='auto', solver='lbfgs'))
```
## Step 2: Compute confident joint
```
def compute_confident_joint(psx, y, verbose=False):
# Verify inputs
psx = np.asarray(psx)
# Find the number of unique classes if K is not given
K = len(np.unique(y))
# Estimate the probability thresholds for confident counting
# You can specify these thresholds yourself if you want
# as you may want to optimize them using a validation set.
# By default (and provably so) they are set to the average class prob.
thresholds = [np.mean(psx[:,k][y == k]) for k in range(K)] # P(s^=k|s=k)
thresholds = np.asarray(thresholds)
# Compute confident joint
confident_joint = np.zeros((K, K), dtype = int)
for i, row in enumerate(psx):
y_label = y[i]
# Find out how many classes each example is confidently labeled as
confident_bins = row >= thresholds - 1e-6
num_confident_bins = sum(confident_bins)
# If more than one conf class, inc the count of the max prob class
if num_confident_bins == 1:
confident_joint[y_label][np.argmax(confident_bins)] += 1
elif num_confident_bins > 1:
confident_joint[y_label][np.argmax(row)] += 1
# Normalize confident joint (use cleanlab, trust me on this)
confident_joint = cleanlab.latent_estimation.calibrate_confident_joint(
confident_joint, y)
if verbose:
cleanlab.util.print_joint_matrix(confident_joint)
return confident_joint
confident_joint = compute_confident_joint(psx, all_y)
```
## Step 3: Find label errors
```
def find_label_errors(confident_joint, y, verbose=False):
# We arbitrarily choose at least 5 examples left in every class.
# Regardless of whether some of them might be label errors.
MIN_NUM_PER_CLASS = 5
# Leave at least MIN_NUM_PER_CLASS examples per class.
# NOTE prune_count_matrix is transposed (relative to confident_joint)
prune_count_matrix = cleanlab.pruning.keep_at_least_n_per_class(
prune_count_matrix=confident_joint.T,
n=MIN_NUM_PER_CLASS,
)
K = len(np.unique(y)) # number of unique classes
y_counts = np.bincount(y)
noise_masks_per_class = []
# For each row in the transposed confident joint
for k in range(K):
noise_mask = np.zeros(len(psx), dtype=bool)
psx_k = psx[:, k]
if y_counts[k] > MIN_NUM_PER_CLASS: # Don't prune if not MIN_NUM_PER_CLASS
for j in range(K): # noisy label index (k is the true label index)
if k != j: # Only prune for noise rates, not diagonal entries
num2prune = prune_count_matrix[k][j]
if num2prune > 0:
# num2prune'th largest p(classk) - p(class j)
# for x with noisy label j
margin = psx_k - psx[:, j]
y_filter = y == j
threshold = -np.partition(
-margin[y_filter], num2prune - 1
)[num2prune - 1]
noise_mask = noise_mask | (y_filter & (margin >= threshold))
noise_masks_per_class.append(noise_mask)
else:
noise_masks_per_class.append(np.zeros(len(s), dtype=bool))
# Boolean label error mask
label_errors_bool = np.stack(noise_masks_per_class).any(axis=0)
# Remove label errors if given label == model prediction
for i, pred_label in enumerate(psx.argmax(axis=1)):
# np.all let's this work for multi_label and single label
if label_errors_bool[i] and np.all(pred_label == y[i]):
label_errors_bool[i] = False
# Convert boolean mask to an ordered list of indices for label errors
label_errors_idx = np.arange(len(y))[label_errors_bool]
# self confidence is the holdout probability that an example
# belongs to its given class label
self_confidence = np.array(
[np.mean(psx[i][y[i]]) for i in label_errors_idx]
)
margin = self_confidence - psx[label_errors_bool].max(axis=1)
label_errors_idx = label_errors_idx[np.argsort(margin)]
label_errors_idx.sort()
if verbose:
print('Indices of label errors found by confident learning:')
print('Note label errors are sorted by likelihood of being an error')
print('but here we just sort them by index for comparison with above.')
print(np.array(label_errors_idx))
return label_errors_idx
label_errors_idx = find_label_errors(confident_joint, all_y)
```
## Print out questionable labels and their existing scores for manual review
```
idx_name = {idx : name for idx, name in enumerate(pos_eg + neg_eg + unlabeled)}
# we'll sort the bad labels for easier review
questionable_good = []
questionable_bad = []
for bad_idx in label_errors_idx:
true_name = idx_name[bad_idx]
the_idx = occs_df.index[ occs_df['occupation'] == true_name].tolist()[0]
if int(occs_df.iloc[the_idx]['label']) ==0:
questionable_bad.append((occs_df.iloc[the_idx]['occupation'], occs_df.iloc[the_idx]['label']))
if int(occs_df.iloc[the_idx]['label']) ==1:
questionable_good.append((occs_df.iloc[the_idx]['occupation'], occs_df.iloc[the_idx]['label']))
for name, label in questionable_good + questionable_bad:
print('"{}", {}'.format(name, label))
```
## Update the dataframe with manually adjusted labels
```
bad_bert_occs = ["shamakhi dancers", "circus arts", "yaksha", "Protestant theology", "architectural photography", "lumières", "consumer protection", "Jerome Mayo Greenberg", "role-playing game", "media literacy", "Pensioner Guards", "social anthropology", "Aviation historians", "short story", "communication medium", "bleachery", "video on demand", "armed struggle", "traditional song", "Rugby league match officials", "zootechnics", "special effects", "plastic surgery", "male prostitution", "deported French resistance", "china painting", "Perm Governorate", "Aalto University", "A Few Good Men", "beer pouring", "transport planning", "Chinese calligraphy", "local government", "natural philosophy", "petroleum-gas industry", "Transactions of the Royal Society of Tropical Medicine and Hygiene", "local history", "fackskolelärare", "digital photography", "social engagement", "Principle of Aberdeen University", "local authority", "computational science", "animal control service", "skolförestånderska", "Academy of Finland", "sigillography", "voice-over", "musical instrument", "Salinas de Ibargoiti", "Saint Petersburg State University", "Tobolsk Governorate", "General Francos's opposition", "Erivan Governorate", "staff and line", "Người dẫn chương trình truyền hình (chương trình TV)", "製本職人", "29 January 28", "Brigade RISTA - Electronic Warfare Intelligence", "Bomberos Voluntarios (Guatemala)", "London South Bank University", "supreme court", "Gubkin University", "High Priests of Amun", "community service", "Postgraduate Work", "Old lesbians: Gendered histories and persistent challenges.", "Canvas print", "theatrical makeup", "list of type designers", "Nazi plunder", "Domestikos", "Catholicos of All Armenians", "conservation movement", "arts administration", "Medical Specimens", "political analysis", "Shadow Theatre", "capo dei capi", "Letrados", "La Révolution prolétarienne", "Ña Catita", "Sofer", "skogsägare", "akademifogde", "organizational leadership", "Carabinieri", "furniture construction", "capitano del popolo", "Željko Ražnatović", "土木技師", "space exploration", "major police MVD", "dollmaking"]
bert_good_occs = ["Ambassadeur", "Chief Executive", "Named Professor", "folk hero", "cinephile", "peasant", "District Chief Executive", "scholar of the bible as literature", "Captain of the guard", "marxist", "Werkmeister", "Franciscan", "public figure", "land surveyor in Poland", "Member of the Congress of Deputies of Spain", "Jägermeister", "galley slave", "Member of Parliament in the Parliament of England", "Young Guard", "madam", "pietist", "Nazi", "patriarch", "Judge Advocate General of the Armed Forces", "competition judge", "diving instructor", "King of Jerusalem", "Tournament director", "Deputy Commander, ROK/US Combined Forces Command", "proselyte", "principal of Uppsala University", "headmaster in France", "reiki master", "Spiker", "combined track and field event athlete", "mayor of Albacete", "medical assistant", "research assistant", "Universal Esperanto Association committee member", "groom", "personality", "celebrity", "centenarian", "Naturaliste", "superhero", "sinecure", "General Officer Commanding", "revenger", "Candidate of Biology Sciences", "human", "Freiherr", "tsarina", "member of the Senate of France", "erudite", "delegate", "passenger", "youth sports minister", "vegetarian", "wanker", "streamer", "mythological king", "internet celebrity", "Polymath", "charcoal burner", "Director-General of the National Heritage Board", "General of the Infantry", "streaker", "pacemaker", "person of short stature", "Iceman", "vase painter", "mother", "patostreamer", "schoolbook publisher", "registrar", "supercentenarian", "fado singer", "Tattooed Lady", "maltster", "cure doctor", "Jedi", "elder", "building contractor", "court Jew"]
print(f"Dataframe length before update {len(occs_df):,}")
for occ in tqdm(bert_good_occs, total=len(bert_good_occs)):
if occ in occs_df['occupation'].tolist():
the_row = occs_df[occs_df['occupation'] == occ ]
the_index = the_row.index[0]
occs_df.loc[the_index, 'label'] = 1
occs_df.loc[the_index, 'labeled_by'] = 'cleanlab'
for occ in tqdm(bad_bert_occs, total=len(bad_bert_occs)):
if occ in occs_df['occupation'].tolist():
the_row = occs_df[occs_df['occupation'] == occ ]
the_index = the_row.index[0]
occs_df.loc[the_index, 'label'] = 0
occs_df.loc[the_index, 'labeled_by'] = 'cleanlab'
print(f"Dataframe length after update {len(occs_df):,}")
```
## Cleanlab, 2nd pass
```
def print_label_names(idx_label_map, label_errors_idx):
label_error_names = [idx_label_map.get(tmp) for tmp in label_errors_idx]
# We'll sort the labels alphabetically and by label value for easier manual review
label_error_names.sort()
pos_labels = []
neg_labels = []
for name in label_error_names:
key_name = name.replace('_', ' ')
val = int(occs_df[occs_df['occupation'] == key_name].label )
if val ==1:
pos_labels.append((key_name, val))
else:
neg_labels.append((key_name, val))
for key_name, val in pos_labels + neg_labels:
print('"{}", {}'.format(key_name, val))
good_occs = occs_df.query("in_google_news ==1 and label ==1")['occupation'].tolist()
bad_occs = occs_df.query("in_google_news ==1 and label ==0")['occupation'].tolist()
y = np.concatenate([np.ones(len(good_occs), dtype=np.int32), np.zeros(len(bad_occs), dtype=np.int32)])
X =[]
model = BertModel.from_pretrained('bert-large-cased')
model.eval()
model.to(device)
with torch.no_grad():
for name in tqdm(good_occs, total=len(good_occs)):
input_ids = torch.tensor(tokenizer.encode(name)).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
X.append(last_hidden_states[0][0].detach().cpu().numpy())
for name in tqdm(bad_occs, total=len(bad_occs)):
input_ids = torch.tensor(tokenizer.encode(name)).unsqueeze(0).to(device) # Batch size 1
outputs = model(input_ids)
last_hidden_states = outputs[0] # The last hidden-state is the first element of the output tuple
X.append(last_hidden_states[0][0].detach().cpu().numpy())
X = np.array(X)
X.shape
all_words = good_occs + bad_occs
# We'll save the mapping of index to words for decoding
idx_label_map = {}
for idx, name in enumerate(all_words):
idx_label_map[idx]= name
psx = cleanlab.latent_estimation.estimate_cv_predicted_probabilities(
X, y, clf=LogisticRegression(max_iter=1000, multi_class='auto', solver='lbfgs'))
confident_joint = compute_confident_joint(psx, y)
label_errors_idx = find_label_errors(confident_joint, y)
print_label_names(idx_label_map, label_errors_idx)
print_label_names(idx_label_map, label_errors_idx)
bad_occs_manual=[ "Freedom Fighters", "Met Éireann", "autopilot", "counterintelligence", "pastry shop", ]
good_occs_manual = [ "Faujdar", "MV Explorer", "Naik", "Raja", "Rinpoche", "Zamindar", "daimyo", "father", ]
print(f"Dataframe length before update {len(occs_df):,}")
for occ in tqdm(good_occs_manual, total=len(good_occs_manual)):
if occ in occs_df['occupation'].tolist():
the_row = occs_df[occs_df['occupation'] == occ ]
the_index = the_row.index[0]
occs_df.loc[the_index, 'label'] = 1
occs_df.loc[the_index, 'labeled_by'] = 'cleanlab'
for occ in tqdm(bad_occs_manual, total=len(bad_occs_manual)):
if occ in occs_df['occupation'].tolist():
the_row = occs_df[occs_df['occupation'] == occ ]
the_index = the_row.index[0]
occs_df.loc[the_index, 'label'] = 0
occs_df.loc[the_index, 'labeled_by'] = 'cleanlab'
print(f"Dataframe length after update {len(occs_df):,}")
occs_df.to_csv('occupations.wikidata.all.labeled.csv', index=False, sep='\t')
occs_df.head()
```
## The dataset is now labeled and verified. Next we will use it to generate negative examples in preparing the Employers dataset
| github_jupyter |
# Finding nemo a bug journey
> follow up from having locally installed xla
- toc:true
- branch: master
- badges: true
- comments: true
- categories: [xla, fastai]
# What happened at first
The major part of the last months we were trying to solve one bug that we didn't know how to solve until this days, some time before fastai version 2 release having a POC and passing the hackathon we participated we left a little the project, but when we came back to it, there were some extrange things happening, some models where not training and apparently others where training, we didn't understand what happened, in that time we thought we introduced an error in our code some way.
## The journey
So, these last few days since I had locally installed XLA and been able to run things, I planned to take a new round to find our bug and it was a perfect opportunity to test what having locally installed `pytorch`+`xla`+`fatai_xla_etensions` could do.
So I passed a lot of assumptions to finally find the solution.
0. So after having locally installed XLA, one of the first things I wanted to test is if locally I could reproduce the error, and I could!
1. The first one was that the error was in our code, but having locally XLA allowed me to test the exact same code changing the device to either: CPU, CUDA or XLA. So I ended up having 2 data loaders and 2 trains on the same python file. Also one main point was that I wrapped a Adam but from pytorch with `OptimWrapper` and it trained correctly so I was more suspicious of differences between the optimizer from fastai and the native to pytorch because there is one know difference about `__getstate__` that is also a requirement for TPU Pods.
2. In the past we have also thought that it was a freeze unfreeze problem, but it was also discarded, so this time I was checking the optimizer, but could not find why the params were not training even when looking under the lens.
3. But after more testings and so on, I see that the second example started to train correctly while the first on the file not, and it was that with all the fresh runs, so I thought it was a problem with the learner, but could not find a "real problem" so I returned back to the optimizer and all, but this time I have a new "tool" I learned, counting the trainable parameters so, the trainable parameters their gradients are updated when you call `backward`, so I started to count there and for the first example they where **always zero** while for the second run since start, they have a number. So the next task was to find why on the first the trainable parameters are always zero and the second not.
But I still didn't get why one model was training and the other one not!
I passed `_BaseOptimizer`, `Optimizer`, `Learner` and others and still could not find the problem, so I decided to compare models and found the problem! I updated the example I found in pytorch forums https://discuss.pytorch.org/t/two-models-with-same-weights-different-results/8918/7 the original one at first run did break because it compared tensors not on same device, so it threw error, I modified it so that it prints them nicely instead of being caught in that error.
```python
def compare_models(model_1, model_2):
models_differ = 0
for key_item_1, key_item_2 in zip(model_1.state_dict().items(), model_2.state_dict().items()):
if key_item_1[1].device == key_item_2[1].device and torch.equal(key_item_1[1], key_item_2[1]):
pass
else:
models_differ += 1
if (key_item_1[0] == key_item_2[0]):
_device = f'device {key_item_1[1].device}, {key_item_2[1].device}' if key_item_1[1].device != key_item_2[1].device else ''
print(f'Mismatch {_device} found at', key_item_1[0])
else:
raise Exception
if models_differ == 0:
print('Models match perfectly! :)')
```
And that was the solution to the problem, I focused on seeing why the models parameters were on different devices. At the end I have something like (remember I don't need to patch the optimizer because I have all installed locally).
```python
def create_opt(self):
print('trainable count before', len(self.all_params(with_grad=True)))
self.opt = self.opt_func(self.splitter(self.model), lr=self.lr)
print('trainable count after', len(self.all_params(with_grad=True)))
if not self.wd_bn_bias:
for p in self._bn_bias_state(True ): p['do_wd'] = False
if self.train_bn:
for p in self._bn_bias_state(False): p['force_train'] = True
```
and
```python
print('trainable count before backward', len(self.all_params(with_grad=True)))
self('before_backward')
print('trainable count before backward', len(self.all_params(with_grad=True)))
self._backward()
self('after_backward')
```
So at the end I see that even that the model is moved later to the device, the first time when `splitter=trainable_params` in `self.opt = self.opt_func(self.splitter(self.model), lr=self.lr)` inside `create_opt` it is not there, so the parameters where stuck on CPU while the the data and the model is later moved to the XLA device.
This does not affects GPUs, but thinking about it could mean also something about the pickable behaviour of xla tensors, specially the optimizer, but that is an history for another time, right now, we have again a simple lib that works for `Single-device TPUs` where you need to modify zero code from fastai.
## Conclusion
So the model need to be in TPU/XLA device before their parameters are taked by splitter on `Optimizer` initialization, I guess we assumed some things in between then and now. At the end it was not exactly an error but was. But sure it was difficult to track, now knowing what it is is solved and we can continue forward.
I hope to add in the next release a `show_lowering_ops` (or similar) to print the counters if you have hit some of those and it is easy to print in a model that runs with this activated. The [MNIST demo](https://github.com/butchland/fastai_xla_extensions/blob/master/samples/MNIST_TPU_demo.ipynb) should be working again don't forget to peek at [fastai_xla_extensions](https://github.com/butchland/fastai_xla_extensions/).
EXTRA NOTE: But there was error because XLA model on CPU not trained when updating backward from data operations on XLA device? well, now I think that XLA worked on TPU with model and data copied to TPU on first time but somehow our model got stuck on CPU so not trained, it became another model separate from the execution happening on TPU (I can think of pickable things, but that is unknown at the moment)
| github_jupyter |
# WORK Token Distribution
A cadCAD sketch for the Workbench WORK token distribution.
## Check cadCAD
This cell doesn't do anything, but it does ensure that you know what version of cadCAD you're running. That way if/when you encounter technical difficulties you can tell the community which version of cadCAD you're running. Might save you hours of pain if all you need to do is upgrade to the latest version.
```
%pip show cadCAD
```
## Import Stuff
These are the libraries you'll need (cadCAD stuff) and that could be useful (python stuff) for your cadCAD model.
```
# python stuff
import numpy as np
import random as random
# cadCAD stuff
from cadCAD.configuration.utils import config_sim
from cadCAD.configuration import Experiment
from cadCAD import configs
from cadCAD.engine import ExecutionMode, ExecutionContext
from cadCAD.engine import Executor
```
## Initial Params
These are the parameters that control the behavior of a system. For example, if you were modeling a Web3 protocol these would be the parameters within a system that token holders could vote to modify. All of the parameters of a protocol that can be modified should be accessible here. That way it's easy to modify and run similations to explore how parameter changes might affect the network.
```
# the initial state of the cadCAD model
# in a stock and flow model you can think of this as the stocks
genesis_states = {
'tokens': 1,
'weekly_token_minting': 100,
'community_treasury': 0,
'contributors': {
'token_balance': 0
}
}
# The parameters to run the model
# Often these are found towards the end of the file near the cadCAD engine
# but we're putting them here so that you can easily configure and run
# the notebook without having to scroll back and forth
sim_config_dict = {
# timesteps: weekly
'T': range(52),
# parallel runs
'N': 3,
# more advanced stuff
#'M': {}
}
```
## Policy Functions
Policy functions are like flows in stock and flow diagrams. They modify the inputs to state update functions.
We start policy functions with p_ so that they're easier to keep track of when we put them into state update blocks.
```
# Mint tokens on a weekly basis (each timestep)
def p_mint_tokens(params, step, sH, s):
minted_tokens = s['weekly_token_minting']
return ({'minted_tokens': minted_tokens})
# Do work to earn tokens
def p_do_work(params, step, sH, s):
# some contributions create more value than others
impact = random.randint(1, 10)
return ({'impact': impact})
```
## State Update Functions
These functions take in inputs (state variables and policies) and modify the state.
We start policy functions with p_ so that they're easier to keep track of when we put them into state update blocks.
```
# Update token count based on weekly minting
def s_update_token_count(params, step, sH, s, _input):
y = 'tokens'
x = s[y]
x += _input['minted_tokens']
return (y, x)
# Put newly minted tokens into the community treasury
def s_update_community_treasury(params, step, sH, s, _input):
y = 'community_treasury'
x = s[y]
x += _input['minted_tokens']
return (y, x)
# Recognize and reward community contributors
def s_reward_contributions(params, step, sH, s, _input):
y = 'contributors'
x = s[y]
x['token_balance'] += _input['impact']
return (y, x)
```
## State Update Block
If you're approaching cadCAD from Web3 you can think of the state of the model as something that evolves in blocks. Each block has a set of actions that updates the state. Those transactions then get batched into blocks to be processed together. In cadCAD blocks are called "`partial_state_update_blocks`." As you can see below, this is an array that is very similar to a "block" in a blockchain in that it represents a set of actions to update the state. That state is then updated across many timesteps. This is similar to how the state of a blockchain is updated over many timesteps as new blocks are added.
```
partial_state_update_blocks = [
{
# mint tokens
'policies': {
'minted_tokens': p_mint_tokens
},
# update token count and community treasury
'variables': {
'tokens': s_update_token_count,
'community_treasury': s_update_community_treasury
}
},
{
# do work
'policies': {
'impact': p_do_work
},
# recognize and reward contributors
'variables': {
'contributors': s_reward_contributions
}
}
]
```
## Running the cadCAD Engine
```
# imported some addition utilities to help with configuration set-up
exp = Experiment()
c = config_sim(sim_config_dict)
# The configurations above are then packaged into a `Configuration` object
del configs[:]
# dict containing variable names and initial values
exp.append_configs(initial_state=genesis_states,
# dict containing state update functions
partial_state_update_blocks=partial_state_update_blocks,
# preprocessed dictionaries containing simulation parameters
sim_configs=c)
%%capture
exec_mode = ExecutionMode()
local_mode_ctx = ExecutionContext(exec_mode.local_mode)
# pass the configuration object inside an array
simulation = Executor(exec_context=local_mode_ctx, configs=configs)
# the `execute()` method returns a tuple; its first elements contains the raw results
raw_system_events, tensor_field, sessions = simulation.execute()
```
## Data Visualization
This is often half the battle. Not only do need to design and build a cadCAD model, but you need to understand how it's working and be able to effectively communicate that to other people. A picture says a thousand words, thus enter data viz. Getting good at using python data viz libraries is probably the highest leverage thing you can do after you learn the cadCAD basics.
```
%matplotlib inline
import pandas as pd
simulation_result = pd.DataFrame(raw_system_events)
simulation_result.set_index(['subset', 'run', 'timestep', 'substep'])
```
simulation_result.plot('timestep', ['tokens', 'community_treasury'], grid=True,
colormap = 'gist_rainbow',
xticks=list(simulation_result['timestep'].drop_duplicates()),
yticks=list(range(1+(simulation_result['tokens']+simulation_result['community_treasury']).max())))
| github_jupyter |
```
import pandas as pd
test_data = pd.read_csv('test/test.csv')
length_data = pd.read_csv('train/length.csv')
# Convert to 0-based
test_data.species_id = test_data.species_id - 1
length_data.species_id = length_data.species_id - 1
import configparser
config = configparser.ConfigParser()
config.read('train_bigelow.ini')
species_names = config.get('Data', 'Species').split(',')
# Convert to species names for easier plotting
def get_name(row):
return species_names[row.species_id]
test_names = test_data.apply(get_name, axis=1)
length_names = length_data.apply(get_name, axis=1)
test_names=test_names.sort_values()
length_names=length_names.sort_values()
def check_for_missing(df):
for name in species_names:
if len(df.loc[df == name]) == 0:
print(f"Missing {name}")
import plotly.express as px
check_for_missing(test_names)
px.histogram(x=test_names, title='Test Data (hold out)')
check_for_missing(length_names)
px.histogram(x=length_names, title='Train (total of train) data')
```
# Retinanet Working
```
retinanet_cols=['img', 'x1','y1','x2','y2', 'species_name']
annotations=pd.read_csv('openem_work/retinanet/annotations.csv', header=None, names=retinanet_cols)
validation=pd.read_csv('openem_work/retinanet/validation.csv', header=None, names=retinanet_cols)
# Check for leakage:
if len(annotations.loc[annotations.img.isin(validation.img)]) != 0:
print("Leakage between train/val")
else:
print("No Leakage between train/val")
unique_train=annotations.img.unique()
unique_val=validation.img.unique()
print
print(f"Unique Training images {len(unique_train)}")
print(f"Unique Training images {len(unique_val)}")
sorted_names=annotations.species_name.sort_values()
check_for_missing(annotations.species_name)
px.histogram(x=sorted_names, title='Train into retinanet')
sorted_names=validation.species_name.sort_values()
check_for_missing(validation.species_name)
px.histogram(x=sorted_names, title='Retinanet Validation')
```
# Annotation Previews
This only shows one box per image, even if there are multiple present as it iterates over annotations.csv each is 1 box per row.
```
import asyncio
class Timer:
def __init__(self, timeout, callback):
self._timeout = timeout
self._callback = callback
self._task = asyncio.ensure_future(self._job())
async def _job(self):
await asyncio.sleep(self._timeout)
self._callback()
def cancel(self):
self._task.cancel()
def debounce(wait):
""" Decorator that will postpone a function's
execution until after `wait` seconds
have elapsed since the last time it was invoked. """
def decorator(fn):
timer = None
def debounced(*args, **kwargs):
nonlocal timer
def call_it():
fn(*args, **kwargs)
if timer is not None:
timer.cancel()
timer = Timer(wait, call_it)
return debounced
return decorator
import ipywidgets as widgets
from IPython.display import clear_output
import matplotlib.pyplot as plt
import os
import numpy as np
w = widgets.IntSlider(min=0, max=len(annotations))
@debounce(0.2)
def value_changed(change):
clear_output()
display(w)
annotation=annotations.iloc[change['new']]
print(annotation)
img_path=os.path.relpath(annotation['img'], '/data')
img = plt.imread(img_path)
fig,axes=plt.subplots(1,1)
axes.set_title(annotation['species_name'])
axes.imshow(img)
x=np.array([annotation.x1, annotation.x2])
y=np.array([annotation.y1, annotation.y2])
axes.plot(x,y, color='red')
plt.show()
display(w)
w.observe(value_changed, 'value')
```
| github_jupyter |
# Second Exploratory Notebook
## Used for Data Exploration of Full Listings Data
```
import pandas as pd
import numpy as np
import nltk
import sklearn
import string, re
import urllib
import seaborn as sbn
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.preprocessing import OneHotEncoder,StandardScaler,LabelEncoder
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.ensemble import RandomForestRegressor
from sklearn.decomposition import PCA
from nltk.corpus import stopwords
data = pd.read_csv('../../Data/2019/listings122019long.csv')
data.head()
data1 = data.drop(columns=['listing_url','scrape_id','last_scraped','summary','space','description','experiences_offered',
'neighborhood_overview','notes','transit','access','interaction','house_rules',
'thumbnail_url','medium_url','picture_url','xl_picture_url','host_id','host_url',
'host_name','host_since', 'host_location','host_about','host_response_time','host_response_rate',
'host_acceptance_rate', 'host_thumbnail_url','host_picture_url', 'host_neighbourhood',
'host_listings_count','host_total_listings_count', 'host_verifications','host_has_profile_pic',
'host_identity_verified','street', 'city','state','zipcode','market','country_code',
'country', 'is_location_exact','bed_type','amenities','square_feet','weekly_price',
'monthly_price','security_deposit','guests_included','extra_people','maximum_nights',
'minimum_minimum_nights','maximum_minimum_nights', 'minimum_maximum_nights', 'maximum_maximum_nights',
'minimum_nights_avg_ntm', 'maximum_nights_avg_ntm', 'calendar_updated', 'has_availability',
'availability_30', 'availability_60', 'availability_90', 'availability_365', 'calendar_last_scraped','first_review',
'last_review','review_scores_accuracy', 'review_scores_cleanliness', 'review_scores_checkin',
'review_scores_communication', 'review_scores_location','requires_license', 'license',
'jurisdiction_names', 'instant_bookable', 'is_business_travel_ready', 'cancellation_policy',
'require_guest_profile_picture', 'require_guest_phone_verification', 'calculated_host_listings_count',
'calculated_host_listings_count_entire_homes', 'calculated_host_listings_count_private_rooms',
'calculated_host_listings_count_shared_rooms', 'neighbourhood','smart_location','id','property_type'
])
data1.describe()
data1.columns
data1.head()
```
# Change Price to Numerical and Get rid of NaNs in data
```
data1['price'] = data1['price'].str.extract('(\d+)', expand=False).astype('float')
data1['cleaning_fee'] = data1['cleaning_fee'].str.extract('(\d+)', expand=False).astype('float')
data1['host_is_superhost'] = (data1['host_is_superhost'] == 't').astype('int')
data1['reviews_per_month'] = data1['reviews_per_month'].fillna(0)
data1['bathrooms'] = data1['bathrooms'].fillna(data1['bathrooms'].mean())
data1['bedrooms'] = data1['bedrooms'].fillna(data1['bedrooms'].mean())
data1['beds'] = data1['beds'].fillna(data1['beds'].mean())
data1['cleaning_fee'] = data1['cleaning_fee'].fillna(data1['cleaning_fee'].mean())
data1['review_scores_rating'] = data1['review_scores_rating'].fillna(0)
data1['longitude'] = data1['longitude'].round(decimals=4)
data1['latitude'] = data1['latitude'].round(decimals=4)
data1 = data1[data1.minimum_nights<31]
```
# Vectorize Categorical Variables
```
ohe_ng = OneHotEncoder(sparse=False)
ohe_n = OneHotEncoder(sparse=False)
ohe_r = OneHotEncoder(sparse=False)
ohe_mn = OneHotEncoder(sparse=False)
neigh_group = ohe_ng.fit_transform(data1[['neighbourhood_group_cleansed']])
neigh_group_cat = ohe_ng.categories_
neigh = ohe_n.fit_transform(data1[['neighbourhood_cleansed']])
neigh_cat = ohe_n.categories_
room = ohe_r.fit_transform(data1[['room_type']])
room_cat = ohe_r.categories_
# prop = ohe.fit_transform(data1[['room_type']])
# prop_cat = ohe.categories_
minimum = ohe_mn.fit_transform(data1[['minimum_nights']])
minimum_cat = ohe_mn.categories_
list(neigh_group_cat)
# ohe_ng.transform([['Northgate']])
len(list(neigh_cat[0]))
# np.array([['Ballard']])
# import pickle
# pickle.dump(ohe_ng, open('ohe_ng.sav', 'wb'))
# pickle.dump(ohe_n, open('ohe_n.sav', 'wb'))
# pickle.dump(ohe_r, open('ohe_r.sav', 'wb'))
# pickle.dump(ohe_mn, open('ohe_mn.sav', 'wb'))
minimum_cat = minimum_cat[0].astype('str')
minimum_cat = list(minimum_cat)
new_minimum_cat = [(lambda x : f'minimum_nights: {x}')(x) for x in range(len(minimum_cat))]
def rename(name_of_columns,pre_addition):
new_list = []
for x in name_of_columns:
for x in x:
new_list.append(pre_addition+ '' + x)
return new_list
new_neigh_group_cat = rename(neigh_group_cat,'neighbourhood_group: ')
new_neigh_cat = rename(neigh_cat,'neighbourhood: ')
new_room_cat = rename(room_cat,'room_type: ')
# new_prop_cat = rename(prop_cat, 'property_type: ')
# Create categories for neighborhood_group, neighborhood and room_type
neigh_group_df = pd.DataFrame(data=neigh_group,columns=new_neigh_group_cat)
neigh_df = pd.DataFrame(data=neigh,columns=new_neigh_cat)
room_type_df = pd.DataFrame(data=room,columns = new_room_cat)
# property_df = pd.DataFrame(data=prop, columns= new_prop_cat)
minimum_df = pd.DataFrame(data=minimum,columns=new_minimum_cat)
```
# NLP for Name Category
```
stopwords_list = stopwords.words('english') + list(string.punctuation)
vectorizer = TfidfVectorizer(strip_accents='unicode',stop_words=stopwords_list,min_df=60,max_df = 800, ngram_range=(1,3))
# get rid of na in name column
data1.fillna({'name':''}, inplace=True)
tf_idf = vectorizer.fit_transform(data['name'])
nlp_name = pd.DataFrame(tf_idf.toarray(), columns=vectorizer.get_feature_names())
nlp_name[13:16]
# import pickle
# pickle.dump(tf_idf, open('nlp.sav', 'wb'))
```
# Reconnect DataFrames / Drop Duplicates
```
clean_data = pd.concat([data1,neigh_group_df,neigh_df,room_type_df,nlp_name,minimum_df],axis=1)
clean_data = clean_data.drop(columns=['name','neighbourhood_cleansed','minimum_nights',
'neighbourhood_group_cleansed','room_type'])
```
# Remove New/ Unsuccessful Properties
```
clean_data.describe()
clean_data = clean_data[clean_data.review_scores_rating>60]
clean_data = clean_data[clean_data.price>20]
clean_data = clean_data[clean_data.price<800]
clean_data = clean_data.dropna()
```
# Visualize Data
```
import seaborn as sbn
import statsmodels
import statsmodels.api as sm
import matplotlib.pyplot as plt
import scipy.stats as stats
fig, axes = plt.subplots(1,3, figsize=(21,6))
sbn.distplot((clean_data['price']), ax=axes[0])
axes[0].set_xlabel('price')
sm.qqplot(np.log1p(clean_data['price']), stats.norm, fit=True, line='45', ax=axes[1])
sbn.scatterplot(x= clean_data['latitude'], y=clean_data['longitude'],hue=clean_data['price'],ax=axes[2]);
correlations = data1.corr()
f, ax = plt.subplots(figsize = (12, 9))
mask = np.zeros_like(correlations, dtype = np.bool)
mask[np.triu_indices_from(mask)] = True
heatmap_one = sbn.heatmap(correlations, cmap ='Reds', mask = mask)
print(heatmap_one)
# fig = heatmap_one.get_figure()
# fig.savefig("heatmap.png")
```
# Train Test Split
```
ss = StandardScaler()
X = clean_data.drop(columns=['price','number_of_reviews_ltm','review_scores_rating',
'review_scores_value','reviews_per_month'])
Xss = ss.fit_transform(X)
y = clean_data['price']
Xtrain,Xtest,ytrain,ytest = train_test_split(Xss,y,test_size = .05,random_state=11)
X.columns
```
# Random Forest
```
rfr = RandomForestRegressor(n_estimators=1000,min_samples_split=5,min_samples_leaf=3,random_state=11)
rfr.fit(Xtrain,ytrain)
rfr.score(Xtrain,ytrain)
yprebootstrapdtrain = rfr.predict(Xtrain)
ypredtest = rfr.predict(Xtest)
from sklearn.metrics import r2_score, explained_variance_score,mean_absolute_error,mean_squared_error
print(r2_score(ytrain,ypredtrain))
print(r2_score(ytest,ypredtest))
# import pickle
# pickle.dump(rfr, open('rf.sav', 'wb'))
sorted(list(zip(rfr.feature_importances_,X.columns)),reverse=True)[0:15]
list(zip(ytest,ypredtest))[0:10]
abs(sum(ytest)-sum(ypredtest))/len(ytest)
y
from sklearn import tree
fig, axes = plt.subplots(nrows = 1,ncols = 1,figsize = (4,4), dpi=800)
tree.plot_tree(rfr.estimators_[0],
feature_names = X.columns,
filled = True);
fig.savefig('rf_individualtree.png')
```
# Neural Net
```
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Dense
from tensorflow.python.keras.layers import Dropout
from tensorflow.python.keras.wrappers.scikit_learn import KerasRegressor
from sklearn.model_selection import KFold
model = Sequential()
model.add(Dense(len(X.columns), input_dim=len(X.columns), kernel_initializer='normal', activation='relu'))
model.add(Dense(200,activation='relu'))
model.add(Dense(1, activation='linear'))
model.summary()
model.compile(loss='mse', optimizer='adam', metrics=['mse','mae'])
history = model.fit(Xtrain, ytrain, epochs=100, batch_size=3, verbose=1, validation_split=0.2)
nnpreds = []
for x in model.predict(Xtest):
for x in x:
nnpreds.append(x)
abs(sum(ytest)-sum(nnpreds))/len(ytest)
print(explained_variance_score(ytest,ypredtest))
# print(explained_variance_score(ytest,nnpreds))
print(mean_absolute_error(ytest,ypredtest))
# print(mean_absolute_error(ytest,nnpreds))
print(r2_score(ytest,ypredtest))
print(r2_score(ytest,nnpreds))
fig, axes = plt.subplots(1,3, figsize=(21,6))
sbn.distplot(list(abs(ytest-ypredtest)/ytest), ax=axes[0])
axes[0].set_xlabel('percentage difference')
sm.qqplot(np.log1p(clean_data['price']), stats.norm, fit=True, line='45', ax=axes[1])
sbn.scatterplot(x= clean_data['latitude'], y=clean_data['longitude'],hue=clean_data['price'],ax=axes[2]);
len(ytest)
list(abs(ytest-ypredtest)/ytest)
sum(abs(ytest-nnpreds)/ytest)/len(ytest)
list(zip(abs(ytest-nnpreds),abs(ytest-ypredtest)))
list(zip(nnpreds,ytest,ypredtest))
list(data1.columns)
list(X.columns)
```
| github_jupyter |
# Recurrent Neural Networks I
Classical neural networks, including convolutional ones, suffer from two severe limitations:
+ They only accept a fixed-sized vector as input and produce a fixed-sized vector as output.
+ They do not consider the sequential nature of some data (language, video frames, time series, etc.)
Recurrent neural networks overcome these limitations by allowing to operate over sequences of vectors (in the input, in the output, or both).
## Vanilla Recurrent Neural Network
<img src="images/vanilla.png" alt="" style="width: 400px;"/>
## Unrolling in time of a RNN
By unrolling we mean that we write out the network for the complete sequence.
$$ s_t = \mbox{tanh }(Ux_t + W s_{t-1}) $$
$$ y_t = \mbox{softmax }(V s_t) $$
<img src="images/unrolling.png" alt="" style="width: 600px;"/>
<img src="images/TanhReal.gif" alt="" style="width: 200px;"/>
## Vanilla Recurrent Neural Network (minibatch version)
<img src="images/minibatch.png" alt="" style="width: 400px;"/>
+ We can think of the **hidden state** $s_t$ as a memory of the network that captures information about the previous steps.
+ The RNN **shares the parameters** $U,V,W$ across all time steps.
+ It is not necessary to have outputs $y_t$ at each time step.
<img src="images/kar.png" alt="" style="width: 600px;"/>
Source: http://karpathy.github.io/2015/05/21/rnn-effectiveness/
RNN have shown success in:
+ Language modeling and generation.
+ Machine Translation.
+ Speech Recognition.
+ Image Description.
+ Question Answering.
+ Etc.
## RNN Computation
```python
class RNN:
#...
def step(self,x):
self.h = np.tanh(np.dot(self.W_hh, self.h) +
np.dot(self.W_xh, self.x))
y = np.dot(self.W_hy, self.h)
return y
#...
```
We can go deep by stacking RNNs:
```python
y1 = rnn1.step(x)
y2 = rnn2.step(y1)
```
Training a RNN is similar to training a traditional NN, but some modifications. The main reason is that parameters are shared by all time steps: in order to compute the gradient at $t=4$, we need to propagate 3 steps and sum up the gradients. This is called **Backpropagation through time (BPTT)**.
The inputs of a recurrent network are always vectors, but we can process sequences of symbols/words by representing these symbols by numerical vectors.
Let's suppose we are classifying a series of words: $x_1, ..., x_{t-1}, x_t, x_{t+1}, ... x_{T}$ are the word vectors corresponding to a corpus with T symbols. Then, the relationship to compute the hidden layer output features at each time-step $t$ is $h_t = \sigma(W^{(hh)} h_{t-1} + W^{(hx)} x_{t})$, where:
+ $x_{t} \in \mathbb{R}^{d}$ is input word vector at time $t$.
+ $W^{hx} \in \mathbb{R}^{D_h \times d}$ is the weights matrix used to condition the input word vector, $x_t$.
+ $W^{hh} \in \mathbb{R}^{D_h \times D_h}$ is the weights matrix used to condition the output of the previous time-step, $h_{t-1}$.
+ $h_{t-1} \in \mathbb{R}^{D_h}$ is the output of the non-linear function at the previous time-step, $t-1$.
+ $h_0 \in \mathbb{R}^{D_h}$ is an initialization vector for the hidden layer at time-step $t = 0$.
+ $\sigma ()$ is the non-linearity function (normally, ``tanh``).
+ $\hat{y}_t = softmax (W^{(S)}h_t)$ is the output probability distribution over the vocabulary at each time-step $t$. Essentially, $\hat{y}_t$ is the next predicted word given the document context score so far (i.e. $h_{t-1}$) and the last observed word vector $x^{(t)}$. Here, $W^{(S)} \in \mathbb{R}^{|V| \times D_h}$ and $\hat{y} \in \mathbb{R}^{|V|}$ where $|V|$ is the vocabulary.
The loss function used in RNNs is often the cross entropy error:
$$
L^{(t)}(W) = - \sum_{j=1}^{|V|} y_{t,j} \times log (\hat{y}_{t,j})
$$
The cross entropy error over a corpus of size $T$ is:
$$
L = \dfrac{1}{T} \sum_{t=1}^{T} L^{(t)}(W) = - \dfrac{1}{T} \sum_{t=1}^{T} \sum_{j=1}^{|V|} y_{t,j} \times log (\hat{y}_{t,j})
$$
In the case of classifying a series of symbols/words, the *perplexity* measure can be used to assess the goodness of our model. It is basically 2 to the power of the negative log probability of the cross entropy error function:
$$
Perplexity = 2^{L}
$$
Perplexity is a measure of confusion where lower values imply more confidence in predicting the next word in the sequence (compared to the ground truth outcome).
## RNN Training
Recurrent neural networks propagate weight matrices from one time-step to the next. Recall the goal of a RNN implementation is to enable propagating context information through faraway time-steps. When these propagation results in a long series of matrix multiplications, weights can vanish or explode.
Once the gradient value grows extremely large, it causes an overflow (i.e. ``NaN``) which is easily detectable at runtime; this issue is called the *Gradient Explosion Problem*.
When the gradient value goes to zero, however, it can go undetected while drastically reducing the learning quality of the model for far-away words in the corpus; this issue is called the *Vanishing Gradient Problem*.
### Gradient Clipping
To solve the problem of exploding gradients, Thomas Mikolov first introduced a simple heuristic solution that *clips* gradients to a small number whenever they explode. That is, whenever they reach a certain threshold, they are set back to a small number.
<img src="images/exploding.png" alt="" style="width: 400px;"/>
### Better initialization
To solve the problem of vanishing gradients, instead of initializing $W^{hh}$ randomly, starting off from **random orthogocal matrices** works better, i.e., a square matrix $W$ for which $W^T W=I$.
There are two properties of orthogonal matrices that are useful for training deep neural networks:
+ they are norm-preserving, i.e., $ ||Wx||^2=||x||^2$, and
+ their columns (and rows) are all orthonormal to one another.
At least at the start of training, the first of these should help to keep the norm of the input constant throughout the network, which can help with the problem of exploding/vanishing gradients.
Similarly, an intuitive understanding of the second is that having orthonormal weight vectors encourages the weights to learn different input features.
You can obtain a random $n \times n$ orthogonal matrix $W$, (uniformly distributed) by performing a QR factorization of an $n \times n$ matrix with elements i.i.d. Gaussian random variables of mean $0$ and variance $1$. Here is an example:
```
import numpy as np
from scipy.linalg import qr
n = 3
H = np.random.randn(n, n)
print(H)
print('\n')
Q, R = qr(H)
print (Q.dot(Q.T))
```
### Steeper Gates
We can make the "gates steeper" so they change more repidly from 0 to 1 and the model is learnt quicker.
<img src="images/steeper.png" alt="" style="width: 600px;"/>
### Gated Units
The most important types of gated RNNs are:
+ Long Short Term Memories (LSTM). It was introduced by S.Hochreiter and J.Schmidhuber in 1997 and is widely used. LSTM is very good in the long run due to its high complexity.
+ Gated Recurrent Units (GRU). It was recently introduced by K.Cho. It is simpler than LSTM, fasters and optimizes quicker.
#### LSTM
The key idea of LSTMs is the cell state $C$, the horizontal line running through the top of the diagram.
The cell state is kind of like a conveyor belt. It runs straight down the entire chain, with only some minor linear interactions. It’s very easy for information to just flow along it unchanged.
<img src="images/lstm.png" alt="Source: http://colah.github.io/posts/2015-08-Understanding-LSTMs/" style="width: 600px;"/>
LSTM has the ability to remove or add information to the cell state, carefully regulated by structures called gates.
Gates are a way to optionally let information through. They are composed out of a *sigmoid* neural net layer and a pointwise multiplication operation.
Let us see how a LSTM uses $h_{t-1}, C_{t-1}$ and $x_{t}$ to generate the next hidden states $C_t, h_{t}$:
$$ f_t = \sigma(W_f \cdot [h_{t-1}, x_t]) \mbox{ (Forget gate)} $$
$$ i_t = \sigma(W_i \cdot [h_{t-1}, x_t]) \mbox{ (Input gate)} $$
$$ \tilde C_t = \operatorname{tanh}(W_C \cdot [h_{t-1}, x_t]) $$
$$ C_t = f_t * C_{t-1} + i_t * \tilde C_t \mbox{ (Update gate)} $$
$$ o_t = \sigma(W_o \cdot [h_{t-1}, x_t]) $$
$$ h_t = o_t * \operatorname{tanh}(C_t) \mbox{ (Output gate)} $$
There are other variants of LSTM (f.e. LSTM with peephole connections of Gers & Schmidhuber (2000))
#### GRU
The transition from hidden state $h_{t-1}$ to $h_{t}$ in vanilla RNN is defined by using an affine transformation and a point-wise nonlinearity.
What motivates the use of gated units? Although RNNs can theoretically capture long-term dependencies, they are very hard to actually train to do this. Gated recurrent units are designed in a manner to have more persistent memory thereby making it easier for RNNs to capture long-term dependencies.
<img src="images/gru.png" alt="Source: http://colah.github.io/posts/2015-08-Understanding-LSTMs/" style="width: 300px;"/>
Let us see how a GRU uses $h_{t-1}$ and $x_{t}$ to generate the next hidden state $h_{t}$.
$$ z_{t} = \sigma(W_z \cdot [x_{t}, h_{t-1}]) \mbox{ (Update gate)}$$
$$ r_{t} = \sigma(W_r \cdot [x_{t}, h_{t-1}]) \mbox{ (Reset gate)}$$
$$ \tilde{h}_{t} = \operatorname{tanh}(r_{t} \cdot [x_{t}, r_t \circ h_{t-1}] ) \mbox{ (New memory)}$$
$$ h_{t} = (1 - z_{t}) \circ \tilde{h}_{t-1} + z_{t} \circ h_{t} \mbox{ (Hidden state)}$$
It combines the forget and input gates into a single “update gate.” It also merges the cell state and hidden state, and makes some other changes. The resulting model is simpler than standard LSTM models.
### RNN in Keras
Whenever you train or test your LSTM/GRU, you first have to build your input matrix $X$ of shape ``nb_samples``, ``timesteps``, ``input_dim`` where your batch size divides ``nb_samples``.
For instance, if ``nb_samples=1024`` and ``batch_size=64``, it means that your model will receive blocks of 64 samples, compute each output (whatever the number of timesteps is for every sample), average the gradients and propagate it to update the parameters vector.
> By default, **Keras shuffles (permutes) the samples in $X$** and the dependencies between $X_i$ and $X_{i+1}$ are lost.
With the stateful model, all the states are propagated to the next batch. It means that the state of the sample located at index $i$, $X_i$, will be used in the computation of the sample $X_{i+bs}$ in the next batch, where $bs$ is the batch size (no shuffling).
> Keras requires the batch size in ``stateful`` mode and ``shuffle=False``.
```
'''Example script showing how to use stateful RNNs
to model long sequences efficiently.
'''
from __future__ import print_function
import numpy as np
import matplotlib.pyplot as plt
from keras.models import Sequential
from keras.layers.core import Dense
from keras.layers.recurrent import LSTM, GRU
# since we are using stateful rnn tsteps can be set to 1
tsteps = 1
batch_size = 25
# number of elements ahead that are used to make the prediction
lahead = 1
def gen_cosine_amp(amp=100, period=25, x0=0, xn=50000, step=1, k=0.0001):
"""Generates an absolute cosine time series with the amplitude
exponentially decreasing
Arguments:
amp: amplitude of the cosine function
period: period of the cosine function
x0: initial x of the time series
xn: final x of the time series
step: step of the time series discretization
k: exponential rate
"""
cos = np.zeros(((xn - x0) * step, 1, 1))
for i in range(len(cos)):
idx = x0 + i * step
cos[i, 0, 0] = amp * np.cos(idx / (2 * np.pi * period))
cos[i, 0, 0] = cos[i, 0, 0] * np.exp(-k * idx)
return cos
print('Generating Data')
cos = gen_cosine_amp()
print('Input shape:', cos.shape)
expected_output = np.zeros((len(cos), 1))
for i in range(len(cos) - lahead):
expected_output[i, 0] = np.mean(cos[i + 1:i + lahead + 1])
print('Output shape')
print(expected_output.shape)
print("Sample: ",cos[0], expected_output[0])
plt.subplot(2, 1, 1)
plt.plot(expected_output)
plt.title('Expected')
plt.show()
epochs = 15
print('Creating Model')
model = Sequential()
model.add(LSTM(50,
batch_input_shape=(batch_size, tsteps, 1),
return_sequences=True,
stateful=True))
model.add(LSTM(50,
batch_input_shape=(batch_size, tsteps, 1),
return_sequences=False,
stateful=True))
model.add(Dense(1))
model.compile(loss='mse', optimizer='rmsprop')
print('Training')
for i in range(epochs):
print('Epoch', i, '/', epochs)
model.fit(cos,
expected_output,
batch_size=batch_size,
verbose=1,
nb_epoch=1,
shuffle=False)
model.reset_states()
print('Predicting')
predicted_output = model.predict(cos, batch_size=batch_size)
print('Ploting Results')
plt.subplot(2, 1, 1)
plt.plot(predicted_output)
plt.title('Predicted')
plt.show()
```
| github_jupyter |
```
import pandas as pd
from constants import *
```
# 2011
------------
<br>
### DATASETS
BLOGS: 120,000 articles
BOOKS: 1,000 books
NEWS: 29,000 articles
PUBMED: 77,000 articles
<br>
### TOPIC MODELING pipeline:
- token -> lemma -> stop -> filter out token w/ count < 10 in vocabulary
- BOW for each document
- 100 topic models
- average PMI score for each topic (Newman 2010b) -> filter out topics w/ score < 0.4
- filter out topics w/ count(token: token default nominal in Wikipedia) < 5
<br>
### RESULTS
| **topics** | **dataset** |
| ------------- |:-------------:|
| 45 | BLOGS |
| 38 | BOOKS |
| 60 | NEWS |
| 85 | PUBMED |
| *228* | *sum* |
=> 6000 labels (~27 per topic)
<br>
#### annotation task:
per topic:
- 10 top topic terms
- 10 label suggestions
- 4 answer options
10 annotations (at least) per label candidate
-> filtered malitious annotators
<br>
45,533 label ratings => ~4,500 successfull HITs (Amazon Mechanical Turk)
```
data_2011 = '../data/topiclabel/2011'
f_topics2011 = join(data_2011, 'topics.csv')
f_annotation2011 = join(data_2011, 'topiclabels.csv')
readme = join(data_2011, 'README.txt')
topics2011 = pd.read_csv(f_topics2011)
annotation2011 = pd.read_csv(f_annotation2011)
print(annotation2011.iloc[:, 2:].count().sum())
examples = ['2008 summer olympics', 'gothic architecture', 'israeli–palestinian conflict', 'immune system']
mask11 = annotation2011.label.isin(examples)
annotation2011['mean'] = annotation2011.loc[mask11, 'rate0':].mean(axis=1)
annotation2011.loc[mask11]
annotation2011[annotation2011.duplicated('label', keep=False)].sort_values('label')
print(topics2011.shape[0])
topics2011.iloc[[7, 11, 89, 105]]
```
# 2016
------
<br>
### DATASETS and annotation
- same datasets and topics as 2011
- very similar annotation method
- annotation quality controlled by 2011 dataset mean ratings
<br>
- 19 top candidates from unsupervised relevance ranking
- 10 annotations per label candidate (pre-filtering)
- 6.4 annotations per label candidate on average (post-filtering)
<br>
#### annotation task:
per topic:
- 10 top topic terms
- 10 label suggestions
- 4 answer options
<br>
27,788 label ratings => ~2,800 successfull HITs (CrowdFlower)
<br>
compute mean of ratings and rank accordingly
```
data_2016 = '../data/topiclabel/2016'
f_topics2016 = join(data_2016, 'topics.csv')
f_annotation2016 = join(data_2016, 'annotated_dataset.csv')
topics2016 = pd.read_csv(f_topics2016)
annotation2016 = pd.read_csv(f_annotation2016, sep='\t')
print(annotation2016.iloc[:, 2:].count().sum())
mask16 = annotation2016.label.isin(examples)
annotation2016['mean'] = annotation2016.loc[mask16, 'annotator1':].mean(axis=1)
annotation2016.loc[mask16]
annotation2016[annotation2016.duplicated('label', keep=False)].sort_values('label')
print(topics2016.shape[0])
topics2016.iloc[[35, 72, 141, 34]]
```
| github_jupyter |
Copyright (c) Microsoft Corporation.
Licensed under the MIT License.
# FWI in Azure project
## Set-up AzureML resources
This project ports devito (https://github.com/opesci/devito) into Azure and runs tutorial notebooks at:
https://nbviewer.jupyter.org/github/opesci/devito/blob/master/examples/seismic/tutorials/
In this notebook we setup AzureML resources. This notebook should be run once and will enable all subsequent notebooks.
<a id='user_input_requiring_steps'></a>
User input requiring steps:
- [Fill in and save sensitive information](#dot_env_description)
- [Azure login](#Azure_login) (may be required first time the notebook is run)
- [Set __create_ACR_FLAG__ to true to trigger ACR creation and to save of ACR login info](#set_create_ACR_flag)
- [Azure CLI login ](#Azure_cli_login) (may be required once to create an [ACR](https://azure.microsoft.com/en-us/services/container-registry/))
```
# Allow multiple displays per cell
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
```
## Azure Machine Learning and Pipeline SDK-specific imports
```
import sys, os
import shutil
import urllib
import azureml.core
from azureml.core import Workspace, Experiment
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
import platform, dotenv
import pathlib
import subprocess
print("Azure ML SDK Version: ", azureml.core.VERSION)
platform.platform()
os.getcwd()
```
#### 1. Create utilities file
##### 1.1 Define utilities file (project_utils.py) path
Utilities file created here has code for Azure resources access authorization, project configuration settings like directories and file names in __project_consts__ class.
```
utils_file_name = 'project_utils'
auxiliary_files_dir = os.path.join(*(['.', 'src']))
utils_path_name = os.path.join(os.getcwd(), auxiliary_files_dir)
utils_full_name = os.path.join(utils_path_name, os.path.join(*([utils_file_name+'.py'])))
os.makedirs(utils_path_name, exist_ok=True)
def ls_l(a_dir):
return ([f for f in os.listdir(a_dir) if os.path.isfile(os.path.join(a_dir, f))])
```
##### 1.2. Edit/create project_utils.py file
```
%%writefile $utils_full_name
from azureml.core.authentication import ServicePrincipalAuthentication
from azureml.core.authentication import AzureCliAuthentication
from azureml.core.authentication import InteractiveLoginAuthentication
from azureml.core.authentication import AuthenticationException
import dotenv, logging, pathlib, os
# credit Mathew Salvaris
def get_auth(env_path):
"""Tries to get authorization info by first trying to get Service Principal info, then CLI, then interactive.
"""
logger = logging.getLogger(__name__)
crt_sp_pwd = os.environ.get("SP_PASSWORD", None)
if crt_sp_pwd:
logger.debug("Trying to create Workspace with Service Principal")
aml_sp_password = crt_sp_pwd
aml_sp_tennant_id = dotenv.get_key(env_path, 'SP_TENANT_ID')
aml_sp_username = dotenv.get_key(env_path, 'SP_APPLICATION_ID')
auth = ServicePrincipalAuthentication(
tenant_id=aml_sp_tennant_id,
username=aml_sp_username,
password=aml_sp_password,
)
else:
logger.debug("Trying to create Workspace with CLI Authentication")
try:
auth = AzureCliAuthentication()
auth.get_authentication_header()
except AuthenticationException:
logger.debug("Trying to create Workspace with Interactive login")
auth = InteractiveLoginAuthentication()
return auth
def set_dotenv_info(dotenv_file_path, env_dict):
"""Use dict loop to set multiple keys in dotenv file.
Minimal file error management.
"""
logger = logging.getLogger(__name__)
if bool(env_dict):
dotenv_file = pathlib.Path(dotenv_file_path)
if not dotenv_file.is_file():
logger.debug('dotenv file not found, will create "{}" using the sensitive info you provided.'.format(dotenv_file_path))
dotenv_file.touch()
else:
logger.debug('dotenv file "{}" found, will (over)write it with current sensitive info you provided.'.format(dotenv_file_path))
for crt_key, crt_val in env_dict.items():
dotenv.set_key(dotenv_file_path, crt_key, crt_val)
else:
logger.debug(\
'Trying to save empty env_dict variable into {}, please set your sensitive info in a dictionary.'\
.format(dotenv_file_path))
class project_consts(object):
"""Keep project's file names and directory structure in one place.
Minimal setattr error management.
"""
AML_WORKSPACE_CONFIG_DIR = ['.', '..', 'not_shared']
AML_EXPERIMENT_DIR = ['.', '..', 'temp']
AML_WORKSPACE_CONFIG_FILE_NAME = 'aml_ws_config.json'
DOTENV_FILE_PATH = AML_WORKSPACE_CONFIG_DIR + ['general.env']
DOCKER_DOTENV_FILE_PATH = AML_WORKSPACE_CONFIG_DIR + ['dockerhub.env']
def __setattr__(self, *_):
raise TypeError
if __name__=="__main__":
"""Basic function/class tests.
"""
import sys, os
prj_consts = project_consts()
logger = logging.getLogger(__name__)
logging.basicConfig(level=logging.DEBUG) # Logging Levels: DEBUG 10, NOTSET 0
logger.debug('AML ws file = {}'.format(os.path.join(*([os.path.join(*(prj_consts.AML_WORKSPACE_CONFIG_DIR)),
prj_consts.AML_WORKSPACE_CONFIG_FILE_NAME]))))
crt_dotenv_file_path = os.path.join(*(prj_consts.DOTENV_FILE_PATH))
set_dotenv_info(crt_dotenv_file_path, {})
```
##### 1.3. Import utilities functions defined above
```
def add_path_to_sys_path(path_to_append):
if not (any(path_to_append in paths for paths in sys.path)):
sys.path.append(path_to_append)
paths_to_append = [os.path.join(os.getcwd(), auxiliary_files_dir)]
[add_path_to_sys_path(crt_path) for crt_path in paths_to_append]
import project_utils
prj_consts = project_utils.project_consts()
```
#### 2. Set-up the AML SDK infrastructure
* Create Azure resource group (rsg), workspaces,
* save sensitive info using [python-dotenv](https://github.com/theskumar/python-dotenv)
Notebook repeateability notes:
* The notebook tries to find and use an existing Azure resource group (rsg) defined by __crt_resource_group__. It creates a new one if needed.
<a id='set_create_ACR_flag'></a>
##### Create [ACR]() first time this notebook is run.
Either docker hub or ACR can be used to store the experimentation image. To create the ACR, set:
```
create_ACR_FLAG=True
```
It will create an ACR by running severral steps described below in section 2.7. __Create an [ACR]__
[Back](#user_input_requiring_steps) to summary of user input requiring steps.
```
create_ACR_FLAG = False #True False
sensitive_info = {}
```
<a id='dot_env_description'></a>
##### 2.1. Input here sensitive and configuration information
[dotenv](https://github.com/theskumar/python-dotenv) is used to hide sensitive info, like Azure subscription name/ID. The serialized info needs to be manually input once.
* REQUIRED ACTION for the 2 cells below: uncomment them, add the required info in first cell below, run both cells one.
The sensitive information will be packed in __sensitive_info__ dictionary variable, which that will then be saved in a following cell in an .env file (__dotenv_file_path__) that should likely be git ignored.
* OPTIONAL STEP: After running once the two cells below to save __sensitive_info__ dictionary variable with your custom info, you can comment them and leave the __sensitive_info__ variable defined above as an empty python dictionary.
__Notes__:
* An empty __sensitive_info__ dictionary is ignored by the __set_dotenv_info__ function defined above in project_utils.py .
* The saved .env file will be used thereafter in each cell that starts with %dotenv.
* The saved .env file contains user specific information and it shoulld __not__ be version-controlled in git.
* If you would like to [use service principal authentication](https://github.com/Azure/MachineLearningNotebooks/blob/master/how-to-use-azureml/manage-azureml-service/authentication-in-azureml/authentication-in-azure-ml.ipynb) make sure you provide the optional values as well (see get_auth function definition in project_utils.py file created above for details).
[Back](#user_input_requiring_steps) to summary of user input requiring steps.
```
# subscription_id = ""
# resource_group = "ghiordanfwirsg01"
# workspace_name = "ghiordanfwiws"
# workspace_region = "eastus2"
# gpu_cluster_name = "gpuclstfwi02"
# gpucluster_admin_user_name = ""
# gpucluster_admin_user_password = ""
# experimentation_docker_image_name = "fwi01_azureml"
# experimentation_docker_image_tag = "sdk.v1.0.60"
# docker_container_mount_point = os.getcwd() # use project directory or a subdirectory
# docker_login = "georgedockeraccount"
# docker_pwd = ""
# acr_name="fwi01acr"
# sensitive_info = {
# 'SUBSCRIPTION_ID':subscription_id,
# 'RESOURCE_GROUP':resource_group,
# 'WORKSPACE_NAME':workspace_name,
# 'WORKSPACE_REGION':workspace_region,
# 'GPU_CLUSTER_NAME':gpu_cluster_name,
# 'GPU_CLUSTER_ADMIN_USER_NAME':gpucluster_admin_user_name,
# 'GPU_CLUSTER_ADMIN_USER_PASSWORD':gpucluster_admin_user_password,
# 'EXPERIMENTATION_DOCKER_IMAGE_NAME':experimentation_docker_image_name,
# 'EXPERIMENTATION_DOCKER_IMAGE_TAG':experimentation_docker_image_tag,
# 'DOCKER_CONTAINER_MOUNT_POINT':docker_container_mount_point,
# 'DOCKER_LOGIN':docker_login,
# 'DOCKER_PWD':docker_pwd,
# 'ACR_NAME':acr_name
# }
```
##### 2.2. Save sensitive info
An empty __sensitive_info__ variable will be ingored.
A non-empty __sensitive_info__ variable will overwrite info in an existing .env file.
```
%load_ext dotenv
dotenv_file_path = os.path.join(*(prj_consts.DOTENV_FILE_PATH))
os.makedirs(os.path.join(*(prj_consts.DOTENV_FILE_PATH[:-1])), exist_ok=True)
pathlib.Path(dotenv_file_path).touch()
# # show .env file path
# !pwd
dotenv_file_path
#save your sensitive info
project_utils.set_dotenv_info(dotenv_file_path, sensitive_info)
```
##### 2.3. Use (load) saved sensitive info
THis is how sensitive info will be retrieved in other notebooks
```
%dotenv $dotenv_file_path
subscription_id = os.getenv('SUBSCRIPTION_ID')
# # print a bit of subscription ID, to show dotenv file was found and loaded
# subscription_id[:2]
crt_resource_group = os.getenv('RESOURCE_GROUP')
crt_workspace_name = os.getenv('WORKSPACE_NAME')
crt_workspace_region = os.getenv('WORKSPACE_REGION')
```
##### 2.4. Access your workspace
* In AML SDK we can get a ws in two ways:
- via Workspace(subscription_id = ...)
- via Workspace.from_config(path=some_file_path).
For demo purposes, both ways are shown in this notebook.
* At first notebook run:
- the AML workspace ws is typically not found, so a new ws object is created and persisted on disk.
- If the ws has been created other ways (e.g. via Azure portal), it may be persisted on disk by calling ws1.write_config(...).
```
workspace_config_dir = os.path.join(*(prj_consts.AML_WORKSPACE_CONFIG_DIR))
workspace_config_file = prj_consts.AML_WORKSPACE_CONFIG_FILE_NAME
# # print debug info if needed
# workspace_config_dir
# ls_l(os.path.join(os.getcwd(), os.path.join(*([workspace_config_dir]))))
```
<a id='Azure_login'></a>
###### Login into Azure may be required here
[Back](#user_input_requiring_steps) to summary of user input requiring steps.
```
try:
ws1 = Workspace(
subscription_id = subscription_id,
resource_group = crt_resource_group,
workspace_name = crt_workspace_name,
auth=project_utils.get_auth(dotenv_file_path))
print("Workspace configuration loading succeeded. ")
ws1.write_config(path=os.path.join(os.getcwd(), os.path.join(*([workspace_config_dir]))),
file_name=workspace_config_file)
del ws1 # ws will be (re)created later using from_config() function
except Exception as e :
print('Exception msg: {}'.format(str(e )))
print("Workspace not accessible. Will create a new workspace below")
workspace_region = crt_workspace_region
# Create the workspace using the specified parameters
ws2 = Workspace.create(name = crt_workspace_name,
subscription_id = subscription_id,
resource_group = crt_resource_group,
location = workspace_region,
create_resource_group = True,
exist_ok = False)
ws2.get_details()
# persist the subscription id, resource group name, and workspace name in aml_config/config.json.
ws2.write_config(path=os.path.join(os.getcwd(), os.path.join(*([workspace_config_dir]))),
file_name=workspace_config_file)
#Delete ws2 and use ws = Workspace.from_config() as shwon below to recover the ws, rather than rely on what we get from one time creation
del ws2
```
##### 2.5. Demo access to created workspace
From now on, even in other notebooks, the provisioned AML workspace will be accesible using Workspace.from_config() as shown below:
```
# path arg is:
# - a file path which explictly lists aml_config subdir for function from_config()
# - a dir path with a silently added <<aml_config>> subdir for function write_config().
ws = Workspace.from_config(path=os.path.join(os.getcwd(),
os.path.join(*([workspace_config_dir, '.azureml', workspace_config_file]))))
# # print debug info if needed
# print(ws.name, ws.resource_group, ws.location, ws.subscription_id[0], sep = '\n')
```
##### 2.6. Create compute cluster used in following notebooks
```
gpu_cluster_name = os.getenv('GPU_CLUSTER_NAME')
gpu_cluster_name
max_nodes_value = 3
try:
gpu_cluster = ComputeTarget(workspace=ws, name=gpu_cluster_name)
print("Found existing gpu cluster")
except ComputeTargetException:
print("Could not find gpu cluster, please create one")
# # Specify the configuration for the new cluster, add admin_user_ssh_key='ssh-rsa ... ghiordan@microsoft.com' if needed
# compute_config = AmlCompute.provisioning_configuration(vm_size="Standard_NC12",
# min_nodes=0,
# max_nodes=max_nodes_value,
# admin_username=os.getenv('GPU_CLUSTER_ADMIN_USER_NAME'),
# admin_user_password=os.getenv('GPU_CLUSTER_ADMIN_USER_NAME'))
# # Create the cluster with the specified name and configuration
# gpu_cluster = ComputeTarget.create(ws, gpu_cluster_name, compute_config)
# # Wait for the cluster to complete, show the output log
# gpu_cluster.wait_for_completion(show_output=True)
```
##### 2.7. Create an [ACR](https://docs.microsoft.com/en-us/azure/container-registry/) if you have not done so using the [portal](https://docs.microsoft.com/en-us/azure/container-registry/container-registry-get-started-portal)
- Follow the 4 ACR steps described below.
- Uncomment cells' lines as needed to login and see commands responses while you set the right subscription and then create the ACR.
- You need [Azure CLI](https://docs.microsoft.com/en-us/cli/azure/install-azure-cli) to run the commands below.
<a id='Azure_cli_login'></a>
##### ACR Step 1. Select ACR subscription (az cli login into Azure may be required here)
[Back](#user_input_requiring_steps) to summary of user input requiring steps.
```
!az --version
# !az login
# ! az account set --subscription $subscription_id
if create_ACR_FLAG:
!az login
response01 = ! az account list --all --refresh -o table
response02 = ! az account set --subscription $subscription_id
response03 = ! az account list -o table
response04 = ! $cli_command
response01
response02
response03
response04
```
##### ACR Step 2. Create the ACR
```
%dotenv $dotenv_file_path
acr_name = os.getenv('ACR_NAME')
cli_command='az acr create --resource-group '+ crt_resource_group +' --name ' + acr_name + ' --sku Basic'
cli_command
response = !$cli_command
response[-14:]
```
##### ACR Step 3. Also enable password and login via __ [--admin-enabled true](https://docs.microsoft.com/en-us/azure/container-registry/container-registry-authentication) __ and then use the az cli or portal to set up the credentials
```
# per https://docs.microsoft.com/en-us/azure/container-registry/container-registry-authentication
cli_command='az acr update -n '+acr_name+' --admin-enabled true'
cli_command
response = !$cli_command
# response
```
##### ACR Step 4. Save the ACR password and login
```
cli_command = 'az acr credential show -n '+acr_name
acr_username = subprocess.Popen(cli_command+' --query username',shell=True,stdout=subprocess.PIPE, stderr=subprocess.PIPE).\
communicate()[0].decode("utf-8").split()[0].strip('\"')
acr_password = subprocess.Popen(cli_command+' --query passwords[0].value',shell=True,stdout=subprocess.PIPE, stderr=subprocess.PIPE).\
communicate()[0].decode("utf-8").split()[0].strip('\"')
response = dotenv.set_key(dotenv_file_path, 'ACR_PASSWORD', acr_password)
response = dotenv.set_key(dotenv_file_path, 'ACR_USERNAME', acr_username)
%reload_ext dotenv
%dotenv -o $dotenv_file_path
# print acr password and login info saved in dotenv file
if create_ACR_FLAG:
os.getenv('ACR_PASSWORD')
os.getenv('ACR_USERNAME')
```
print('Finished running 000_Setup_GeophysicsTutorial_FWI_Azure_devito!')
| github_jupyter |
# Alzheimer's Disease Capstone Project
## Machine Learning and Predictive Modeling
The two primary questions that this analysis is trying to answer are:
1. Which biomarkers are correlated with a change in diagnosis to Alzheimer's Disease?
2. Which biomarkers might be able to predict a final diagnosis of Alzhiemer's Disease at an initial visit?
```
# load packages
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.multiclass import OneVsRestClassifier
from sklearn import linear_model
from sklearn.naive_bayes import MultinomialNB
from sklearn.neighbors import KNeighborsClassifier
from sklearn.model_selection import train_test_split, cross_val_score, GridSearchCV
from sklearn.decomposition import PCA
from sklearn.metrics import roc_curve, roc_auc_score, confusion_matrix, classification_report
# load custom modules
from adnidatawrangling import wrangle_adni
import eda, ml
# set default plotting
plt.style.use('ggplot')
```
## 1. Which Biomarkers Are Correlated with a Progression Towards Alzheimer's Disease?
- Steps
- Prepare data for analysis
- Import/wrangle/clean
- Extract the last exam and basline data
- Calculate the change for each variables (deltas)
- Extract the features and standardize the data
- Separate the target and feature data
```
# import data, clean, and extract data
adni_comp, clin_data, scan_data = wrangle_adni()
# extract final exam data: only the last exam for each patient
final_exam = eda.get_final_exam(adni_comp)
# calculate the change in variables over the course of the study
eda.calc_deltas(final_exam)
# extract and scale the deltas data for ML analysis
feature_names, Xd, yd = ml.get_delta_scaled(final_exam)
# examine the structure of the data
print(feature_names.shape)
print(Xd.shape, yd.shape)
# split the data into test and train sets
Xd_train, Xd_test, yd_train, yd_test = train_test_split(Xd, yd, test_size=0.3,
random_state=21, stratify=yd)
```
#### First Model k-Nearest Neighbors Classification
- The first model will try k-NN to predict the groups
- First, the best k will be chosen
- Graphic approach
- GridSearchCV approach
```
# plot the training and test accuracy with varying k values
ml.plot_best_k(Xd_train, Xd_test, yd_train, yd_test, kmax=50)
```
- This approach suggests 17 nearest neighbors is a good number.
```
# find best k using GridSearchCV
param_grid = {'n_neighbors': np.arange(1, 50)}
knn = KNeighborsClassifier()
knn_cv = GridSearchCV(knn, param_grid, cv=5)
knn_cv.fit(Xd_train, yd_train)
# print the best value(s) for each hyperparameter and the mean of the best validation score
print(knn_cv.best_params_, knn_cv.best_score_)
```
- GridSearchCV returned 39 as the best k. Because the first approach gave 17, that will be the first k used in the model.
#### k-Nearest Neighbors Model in Action
```
# create the model
knn = KNeighborsClassifier(n_neighbors=17)
# fit the model
knn.fit(Xd_train, yd_train)
# generate predictions
y_pred = knn.predict(Xd_test)
# print the confusion matrix
print(confusion_matrix(yd_test, y_pred))
# print the accuracy
print('Training Accuracy: {}'.format(knn.score(Xd_train, yd_train)))
print('Testing Accuracy: {}'.format(knn.score(Xd_test, yd_test)))
print(classification_report(yd_test, y_pred))
# old code for reference
# map the diagnosis group and assign to dx_group
nc_idx = final_exam[final_exam.DX == final_exam.DX_bl2].index
cn_mci_idx = final_exam[(final_exam.DX == 'MCI') & (final_exam.DX_bl2 == 'CN')].index
mci_ad_idx = final_exam[(final_exam.DX == 'AD') & (final_exam.DX_bl2 == 'MCI')].index
cn_ad_idx = final_exam[(final_exam.DX == 'AD') & (final_exam.DX_bl2 == 'CN')].index
labels = pd.concat([pd.DataFrame({'dx_group': 'No Change'}, index=nc_idx),
pd.DataFrame({'dx_group': 'CN to MCI'}, index=cn_mci_idx),
pd.DataFrame({'dx_group': 'MCI to AD'}, index=mci_ad_idx),
pd.DataFrame({'dx_group': 'CN to AD'}, index=cn_ad_idx)
]).sort_index()
# add to the dataframe and ensure every row has a label
deltas_df = final_exam.loc[labels.index]
deltas_df.loc[:,'dx_group'] = labels.dx_group
type(deltas_df)
pd.get_dummies(deltas_df, drop_first=True, columns=['PTGENDER'])
# extract the features for change in diagnosis
X_delta_male = deltas_df[deltas_df.PTGENDER == 'Male'].reindex(columns=['CDRSB_delta', 'ADAS11_delta',
'ADAS13_delta', 'MMSE_delta',
'RAVLT_delta', 'Hippocampus_delta',
'Ventricles_delta', 'WholeBrain_delta',
'Entorhinal_delta', 'MidTemp_delta'])
X_delta_female = deltas_df[deltas_df.PTGENDER == 'Female'].reindex(columns=['CDRSB_delta', 'ADAS11_delta',
'ADAS13_delta', 'MMSE_delta',
'RAVLT_delta', 'Hippocampus_delta',
'Ventricles_delta', 'WholeBrain_delta',
'Entorhinal_delta', 'MidTemp_delta'])
male_scaler = StandardScaler()
female_scaler = StandardScaler()
Xd_male = male_scaler.fit_transform(X_delta_male)
Xd_female = female_scaler.fit_transform(X_delta_female)
# extract the labels
yd_male = np.array(deltas_df[deltas_df.PTGENDER == 'Male'].dx_group)
yd_female = np.array(deltas_df[deltas_df.PTGENDER == 'Female'].dx_group)
print(X_delta_male.shape, yd_male.shape)
print(X_delta_female.shape, yd_female.shape)
```
| github_jupyter |
# Análise, Limpeza e Engenharia de Dados
```
##Bibliotecas úteis
import pandas as pd ## Para manipulação dos conjuntos de dados
import numpy as np ## Para
import chardet ## Biblioteca para lidar com a codificação de caracteres
import matplotlib.pyplot as plt
import seaborn as sns
# para escala
from mlxtend.preprocessing import minmax_scaling
# Para transformação Box-Cox
from scipy import stats
from sklearn.preprocessing import OneHotEncoder, LabelEncoder, MinMaxScaler
%matplotlib inline
```
## Abrir o conjunto de dados
```
weather_df = pd.read_csv('bases/summary_of_weather.csv', index_col=0)
```
## Visualizar o conjunto de dados
```
weather_df.head()
```
#### Tem alguns dados faltando...
## Limpar o conjunto de dados
--Lidar com dados faltantes, como: __NaN__ ou __None__
--Remover colunas sujas
### -- Lidar com dados faltantes
#### Verificando o número de dados faltantes por coluna
```
weather_df.isnull().sum()
weather_df.isnull().sum().plot(kind='bar')
```
#### Verificando a porcentagem de dados faltantes
```
## Vamos checar a quantidade de linhas e colunas
weather_df.shape
## Agora, o número total de células no dataset
num_total_celulas_weather_df = np.product(weather_df.shape)
num_total_celulas_weather_df
## E o numero de células com números faltantes
num_celulas_faltantes = weather_df.isnull().sum().sum()
num_celulas_faltantes
print('Porcentagem de dados faltantes:', 100 * num_celulas_faltantes / num_total_celulas_weather_df)
```
### Remover dados faltantes
Vamos começar removendo todas as __linhas__ que contém algum valor faltante
```
weather_df.dropna()
```
#### Eita! Não sobrou nenhuma linha!
Vamos tentar remover todas as __colunas__ que têm algum valor faltante:
```
weather_df_sem_nan = weather_df.dropna(axis=1)
weather_df_sem_nan.head()
```
Vamos ver a quantidade de dados perdidos:
```
print('Quantidade de colunas no dataset original:', weather_df.shape[1])
print('Quantidade de colunas sem valores faltantes:', weather_df_sem_nan.shape[1])
```
Sobraram 9 colunas!
###### Conseguimos remover dados e continuar com uma parcela do dataset!
##### Mas quanto?
```
## Agora, o número total de células no dataset
num_total_celulas_weather_df = np.product(weather_df.shape)
print('num_total_celulas_weather_df', num_total_celulas_weather_df)
## E o numero de células com números faltantes
num_celulas_weather_df_sem_nan = np.product(weather_df_sem_nan.shape)
print('num_celulas_weather_df_sem_nan', num_celulas_weather_df_sem_nan)
print('Porcentagem de dados resultantes:', 100 * num_celulas_weather_df_sem_nan / num_total_celulas_weather_df)
```
#### Perdemos MUITO dado!
###### Vamos olhar mais atentamente para o conjunto de dados.
###### Quais dados não existem e quais não foram coletados?
-- Se o dado não existe, não faz sentido tentar adivinhar qual valor deveria estar ali
-- Se o dado não foi coletado, vale a pena tentar colocar algo na célula desfalcada
```
#Mostrando 5 amostras do dataset
weather_df.sample(5)
### Escolha uma coluna e analise seus dados faltosos
```
#### Exemplo: coluna 'Snowfall'
```
### Ver quantos e quais dados não-nulos
pd.value_counts(weather_df['Snowfall'])
weather_df.sort_values(by='Date', inplace=True)
weather_df.head()
weather_df['Snowfall'] = weather_df['Snowfall'].fillna(method='bfill', axis=0).fillna(0)
weather_df.isnull().sum()
pd.value_counts(weather_df['Snowfall'])
```
###### Tratar os valores que não são numéricos
```
weather_df.loc[:, 'Snowfall'].replace(0.0, 0, inplace=True)
weather_df.loc[:, 'Snowfall'].replace('0', 0, inplace=True)
weather_df.loc[:, 'Snowfall'].replace('#VALUE!', method='bfill', inplace=True)
pd.value_counts(weather_df['Snowfall'])
```
#### -- Remover colunas com todos os valores nulos
```
weather_df.dropna(how='all', axis=1, inplace=True)
weather_df.isnull().sum()
```
#### -- Aplicar fillna para todo o dataset
```
weather_df = weather_df.fillna(method='ffill', axis=0).fillna(0)
```
### -- Coluna Precip
```
pd.value_counts(weather_df['Precip'])
```
###### Limpar dados não-numéricos da coluna
```
weather_df.loc[:,'Precip'].replace('T',0, inplace=True)
weather_df.sample(5)
```
###### Mudar tipo da coluna
```
weather_df.Precip = weather_df.Precip.astype(float)
```
## Engenharia de Dados
##### Tratamento de Datas
```
### Sua vez! Trate a coluna Date
### Sua vez! monte uma coluna nova com o mês
### Sua vez! monte uma coluna nova com o valor da precipitação do mês anterior
```
## Escala e Padronização
### Escala
```
weather_df_MaxTemp = weather_df.MaxTemp
# Colocar em escala de 0 a 1
scaled_data = minmax_scaling(weather_df_MaxTemp, columns = [0])
# plotar os dados originais e em escala
fig, ax=plt.subplots(1,2)
sns.distplot(weather_df_MaxTemp, ax=ax[0])
ax[0].set_title("Dados Originais")
sns.distplot(scaled_data, ax=ax[1])
ax[1].set_title("Dados em Escala")
# Sua vez!
# Aplique a função scale para a coluna "MinTemp"
## Sua vez!
## Identifique as colunas de dados numéricos utilizando a biblioteca pandas e numpy
## Sua vez! Aplique a função scale para todas as colunas de dados numéricos
numeric_columns = list(weather_df.select_dtypes(include=[np.number]).columns)
numeric_columns.remove('STA')
numeric_columns.remove('YR')
numeric_columns.remove('MO')
numeric_columns.remove('DA')
scaled_columns = minmax_scaling(weather_df[numeric_columns], numeric_columns)
scaled_columns.head()
```
### Normalização
```
scaled_columns['MaxTemp'].hist()
# normalize the exponential data with boxcox
scaled_maxtemp = scaled_columns['MaxTemp']
normalized_data = stats.boxcox(scaled_maxtemp[scaled_maxtemp>0])
# plot both together to compare
fig, ax=plt.subplots(1,2)
sns.distplot(scaled_maxtemp, ax=ax[0])
ax[0].set_title("Dados Originais")
sns.distplot(normalized_data[0], ax=ax[1])
ax[1].set_title("Dados Normalizados")
# Sua vez!
# Normalize a coluna "MinTemp"
## Sua vez! Normalize todas as colunas de dados numéricos
```
###### Sem usar biblioteca:
```
# normalize the exponential data with boxcox
scaled_maxtemp = scaled_columns['MaxTemp']
normalized_data = ( scaled_maxtemp - scaled_maxtemp.mean() ) / scaled_maxtemp.std()
# plot both together to compare
fig, ax=plt.subplots(1,2)
sns.distplot(scaled_maxtemp, ax=ax[0])
ax[0].set_title("Dados Originais")
sns.distplot(normalized_data, ax=ax[1])
ax[1].set_title("Dados Normalizados")
```
## One Hot Encoder
#### Via sklearn
```
weather_df['Month'] = pd.to_datetime(weather_df['Date'], format="%Y-%m-%d").dt.strftime("%b")
one_hot_encoder = OneHotEncoder(categories='auto')
ohe_month = one_hot_encoder.fit_transform(weather_df['Month'].values.reshape(-1,1)).toarray()
ohe_month_df = pd.DataFrame(ohe_month, columns=['month_' + str(i) for i in range(ohe_month.shape[1])])
ohe_month_df.head()
weather_df_month_ohe = pd.concat([weather_df, ohe_month_df],axis=1)
weather_df_month_ohe.head()
```
###### Via pandas
```
ohe_month = pd.get_dummies(weather_df['Month'], dummy_na=True)
weather_df_month_ohe = pd.concat([weather_df, ohe_month],axis=1)
weather_df_month_ohe.head()
# Faça OHE para a coluna YR. Escolha um método de sua preferência.
```
| github_jupyter |
Due to the few points in each dimension and the straight line that linear regression uses to follow these points as well as it can, noise on the observations will cause great variance as shown in the first plot. Every line’s slope can vary quite a bit for each prediction due to the noise induced in the observations.
Ridge regression is basically minimizing a penalised version of the least-squared function. The penalising shrinks the value of the regression coefficients. Despite the few data points in each dimension, the slope of the prediction is much more stable and the variance in the line itself is greatly reduced, in comparison to that of the standard linear regression
#### New to Plotly?
Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
<br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
<br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
### Version
```
import sklearn
sklearn.__version__
```
### Imports
```
import plotly.plotly as py
import plotly.graph_objs as go
from plotly import tools
import numpy as np
from sklearn import linear_model
```
### Calculations
```
X_train = np.c_[.5, 1].T
y_train = [.5, 1]
X_test = np.c_[0, 2].T
np.random.seed(0)
classifiers = dict(ols=linear_model.LinearRegression(),
ridge=linear_model.Ridge(alpha=.1))
```
### Plot Results
```
fig = tools.make_subplots(rows=1, cols=2,
print_grid=False,
subplot_titles=('ols', 'ridge'))
def data_to_plotly(x):
k = []
for i in range(0, len(X_test)):
k.append(x[i][0])
return k
fignum = 1
for name, clf in classifiers.items():
for _ in range(6):
this_X = .1 * np.random.normal(size=(2, 1)) + X_train
clf.fit(this_X, y_train)
p1 = go.Scatter(x=data_to_plotly(X_test),
y=clf.predict(X_test),
mode='lines', showlegend=False,
line=dict(color='gray', width=1))
p2 = go.Scatter(x=data_to_plotly(this_X),
y=y_train, showlegend=False,
mode='markers',
marker=dict(color='gray')
)
fig.append_trace(p1, 1, fignum)
fig.append_trace(p2, 1, fignum)
clf.fit(X_train, y_train)
p3 = go.Scatter(x=data_to_plotly(X_test),
y=clf.predict(X_test),
mode='lines', showlegend=False,
line=dict(color='blue', width=2)
)
p4 = go.Scatter(x=data_to_plotly(X_train),
y=y_train,
mode='markers', showlegend=False,
marker=dict(color='red')
)
fig.append_trace(p3, 1, fignum)
fig.append_trace(p4, 1, fignum)
fignum += 1
for i in map(str, range(1, 3)):
x = 'xaxis' + i
y = 'yaxis' + i
fig['layout'][x].update(title='x', zeroline=False)
fig['layout'][y].update(title='y', zeroline=False)
py.iplot(fig)
```
### License
Code source:
Gaël Varoquaux
Modified for documentation by Jaques Grobler
License:
BSD 3 clause
```
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'Ordinary Least Squares and Ridge Regression Variance.ipynb', 'scikit-learn/plot-ols-ridge-variance/', ' Ordinary Least Squares and Ridge Regression Variance | plotly',
' ',
title = 'Ordinary Least Squares and Ridge Regression Variance | plotly',
name = 'Ordinary Least Squares and Ridge Regression Variance',
has_thumbnail='true', thumbnail='thumbnail/ols.jpg',
language='scikit-learn', page_type='example_index',
display_as='linear_models', order=7,
ipynb= '~Diksha_Gabha/3186')
```
| github_jupyter |
```
from __future__ import (print_function, absolute_import, division,
unicode_literals, with_statement)
from sklearn.model_selection import train_test_split
from sklearn import datasets
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import ParameterGrid
import numpy as np
from cleanlab.classification import LearningWithNoisyLabels
from cleanlab.noise_generation import generate_noisy_labels
from cleanlab.util import value_counts
from cleanlab.latent_algebra import compute_inv_noise_matrix
```
`cleanlab` can be used with any classifier and dataset for multiclass
learning with noisy labels. Its comprised of components from the theory and
algorithms of **confident learning**. It's a Python class that wraps around
any classifier as long as .fit(X, y, sample_weight),
.predict(X), .predict_proba(X) are defined.
See https://l7.curtisnorthcutt.com/cleanlab-python-package for docs.
## Here we show the performance with LogisiticRegression classifier
## versus LogisticRegression \*without\* cleanlab on the Iris dataset.
```
# Seed for reproducibility
seed = 2
clf = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000)
rp = LearningWithNoisyLabels(clf=clf, seed=seed)
np.random.seed(seed=seed)
# Get iris dataset
iris = datasets.load_iris()
X = iris.data # we only take the first two features.
y = iris.target
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2)
try:
get_ipython().run_line_magic('matplotlib', 'inline')
from matplotlib import pyplot as plt
_ = plt.figure(figsize=(12, 8))
color_list = plt.cm.tab10(np.linspace(0, 1, 6))
_ = plt.scatter(X_train[:, 1], X_train[:, 3],
color=[color_list[z] for z in y_train], s=50)
ax = plt.gca()
# plt.axis('off')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
_ = ax.get_xaxis().set_ticks([])
_ = ax.get_yaxis().set_ticks([])
_ = plt.title("Iris dataset (feature 3 vs feature 1)", fontsize=30)
except Exception as e:
print(e)
print("Plotting is only supported in an iPython interface.")
# Generate lots of noise.
noise_matrix = np.array([
[0.5, 0.0, 0.0],
[0.5, 1.0, 0.5],
[0.0, 0.0, 0.5],
])
py = value_counts(y_train)
# Create noisy labels
s = generate_noisy_labels(y_train, noise_matrix)
try:
get_ipython().run_line_magic('matplotlib', 'inline')
from matplotlib import pyplot as plt
_ = plt.figure(figsize=(15, 8))
color_list = plt.cm.tab10(np.linspace(0, 1, 6))
for k in range(len(np.unique(y_train))):
X_k = X_train[y_train == k] # data for class k
_ = plt.scatter(
X_k[:, 1],
X_k[:, 3],
color=[color_list[noisy_label] for noisy_label in s[y_train == k]],
s=200,
marker=r"${a}$".format(a=str(k)),
linewidth=1,
)
_ = plt.scatter(
x=X_train[s != y_train][:, 1],
y=X_train[s != y_train][:, 3],
color=[color_list[z] for z in s],
s=400,
facecolors='none',
edgecolors='black',
linewidth=2,
alpha=0.5,
)
ax = plt.gca()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
_ = ax.get_xaxis().set_ticks([])
_ = ax.get_yaxis().set_ticks([])
_ = plt.title("Iris dataset (features 3 and 1). Label errors circled.",
fontsize=30)
except Exception as e:
print(e)
print("Plotting is only supported in an iPython interface.")
print('WITHOUT confident learning,', end=" ")
clf = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000)
_ = clf.fit(X_train, s)
pred = clf.predict(X_test)
print("Iris dataset test accuracy:", round(accuracy_score(pred, y_test), 2))
print("\nNow we show improvement using cleanlab to characterize the noise")
print("and learn on the data that is (with high confidence) labeled correctly.")
print()
print('WITH confident learning (noise matrix given),', end=" ")
_ = rp.fit(X_train, s, noise_matrix=noise_matrix)
pred = rp.predict(X_test)
print("Iris dataset test accuracy:", round(accuracy_score(pred, y_test), 2))
print('WITH confident learning (noise / inverse noise matrix given),', end=" ")
inv = compute_inv_noise_matrix(py, noise_matrix)
_ = rp.fit(X_train, s, noise_matrix=noise_matrix, inverse_noise_matrix=inv)
pred = rp.predict(X_test)
print("Iris dataset test accuracy:", round(accuracy_score(pred, y_test), 2))
print('WITH confident learning noise not given,', end=" ")
clf = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000)
rp = LearningWithNoisyLabels(clf=clf, seed=seed)
_ = rp.fit(X_train, s)
pred = rp.predict(X_test)
print("Iris dataset test accuracy:", round(accuracy_score(pred, y_test), 2))
```
## Performance of confident learning across varying settings.
To learn more, inspect ```cleanlab/pruning.py``` and ```cleanlab/classification.py```.
```
param_grid = {
"prune_method": ["prune_by_noise_rate", "prune_by_class", "both"],
"converge_latent_estimates": [True, False],
}
# Fit LearningWithNoisyLabels across all parameter settings.
params = ParameterGrid(param_grid)
scores = []
for param in params:
clf = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000)
rp = LearningWithNoisyLabels(clf=clf, n_jobs=1, **param)
_ = rp.fit(X_train, s) # s is the noisy y_train labels
scores.append(accuracy_score(rp.predict(X_test), y_test))
# Print results sorted from best to least
for i in np.argsort(scores)[::-1]:
print("Param settings:", params[i])
print(
"Iris dataset test accuracy (using confident learning):\t",
round(scores[i], 2),
"\n",
)
param_grid = {
"prune_method": ["prune_by_noise_rate", "prune_by_class", "both"],
"converge_latent_estimates": [True, False],
}
# Fit LearningWithNoisyLabels across all parameter settings.
params = ParameterGrid(param_grid)
scores = []
for param in params:
clf = LogisticRegression(solver='lbfgs', multi_class='auto', max_iter=1000)
rp = LearningWithNoisyLabels(clf=clf, n_jobs=1, **param)
_ = rp.fit(X_train, s) # s is the noisy y_train labels
scores.append(accuracy_score(rp.predict(X_test), y_test))
# Print results sorted from best to least
for i in np.argsort(scores)[::-1]:
print("Param settings:", params[i])
print(
"Iris dataset test accuracy (using confident learning):\t",
round(scores[i], 2),
"\n",
)
```
| github_jupyter |
```
#!/usr/bin/env python3
# This script takes a sdf for a set molecules with a SMILES column as input and filters out repeating entries,
# molecules without prices and molecules with lower availability than 100 mg.
import pandas as pd
import numpy as np
from openeye import oechem, oedepict, oemolprop
import oenotebook as oenb
import matplotlib.pyplot as plt
%matplotlib inline
# Convert SDF file of full similarity list to csv for easier manipulation
ifs = oechem.oemolistream()
ofs = oechem.oemolostream()
ifs.SetFormat(oechem.OEFormat_SDF)
ofs.SetFormat(oechem.OEFormat_CSV)
#for mol in ifs.GetOEGraphMols():
if ifs.open("eMol_similarity_set_2017_07.sdf"):
if ofs.open("eMol_similarity_set_2017_07.csv"):
for mol in ifs.GetOEGraphMols():
oechem.OEWriteMolecule(ofs, mol)
else:
OEThrow.Fatal("Unable to create 'output.mol2'")
else:
OEThrow.Fatal("Unable to open 'input.sdf'")
print("SDF file converted to CSV: eMol_similarity_set_2017_07.csv")
df_eMol_sim = pd.read_csv("eMol_similarity_set_2017_07.csv") # This file contains full starting set of compounds
print("Number of molecules: ", df_eMol_sim.shape[0])
df_eMol_sim
# Eliminate repeating entries
df_eMol_sim.drop_duplicates(inplace=True)
df_eMol_sim = df_eMol_sim.reset_index(drop=True)
print("Number of unique enteries :", df_eMol_sim.shape[0])
# Eliminate entries without price
df_eMol_sim_price = df_eMol_sim[np.isfinite(df_eMol_sim["Price_USD"])].reset_index(drop=True)
print("Number of unique enteries with price :", df_eMol_sim_price.shape[0])
# Eliminate entries not in Tier1
df_eMol_sim_price_tier1 = df_eMol_sim_price[df_eMol_sim_price["Supplier_tier"] == 1.0].reset_index(drop=True)
print("Number of unique enteries with price from Tier1 :",df_eMol_sim_price_tier1.shape[0])
# Eliminate entries with availability less than 100 mg
df_eMol_sim_price_tier1_100mg = df_eMol_sim_price_tier1[df_eMol_sim_price_tier1["Amount_mg"] >= 100.0].reset_index(drop=True)
print("Number of unique enteries with price from Tier1, 100 mg availability:",df_eMol_sim_price_tier1_100mg.shape[0])
df_eMol_sim_price_tier1_100mg.to_csv("df_eMol_sim_price_tier1_100mg.csv")
df_eMol_sim_price_tier1_100mg
# Eliminate repeating molecules based on canonical isomeric SMILES
df_eMol_sim_unique_molecules = df_eMol_sim_price_tier1_100mg.drop_duplicates(subset = "SMILES")
print("Number of unique molecules: ", df_eMol_sim_unique_molecules.shape[0])
# Eliminate compound with eMolecules SKU 112319653, because it takes too long for conformer generation.
df_eMol_sim_unique_molecules = df_eMol_sim_unique_molecules[df_eMol_sim_unique_molecules["TITLE"] != 112319653]
print("Number of unique molecules: ", df_eMol_sim_unique_molecules.shape[0])
df_eMol_sim_unique_molecules.to_csv("df_eMol_sim_unique_molecules.csv")
df_eMol_sim_unique_molecules_smiles = df_eMol_sim_unique_molecules.loc[:, ("SMILES", "TITLE")]
df_eMol_sim_unique_molecules_smiles.to_csv("df_eMol_sim_unique_molecules_smiles.smi")
df_eMol_sim_unique_molecules_smiles
```
| github_jupyter |
# Using MVR to predict porosity from well log data
Put together by Thomas Martin, thomasmartin@mines.edu, all errors are mine
```
!pip install scikit-learn --upgrade
# If you have installation questions, please reach out
import pandas as pd # data storage
import numpy as np # math and stuff
import sklearn
import datetime
from sklearn import linear_model
import statsmodels.api as sm
from sklearn.metrics import mean_squared_error, median_absolute_error
from sklearn.utils.class_weight import compute_sample_weight
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, max_error, mean_squared_error
from sklearn.model_selection import GridSearchCV
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt # plotting utility
from google.colab import drive
drive.mount('/content/drive')
df = pd.read_csv('drive/My Drive/1_lewis_research/core_to_wl_merge/Merged_dataset_inner_imputed_12_21_2020.csv')
df = df.drop(['Unnamed: 0', 'Unnamed: 0.1', 'LiveTime2','ScanTime2', 'LiveTime1','ScanTime1',
'ref_num', 'API', 'well_name', 'sample_num', 'sample' ], axis=1)
df.describe()
```
## Loading in Dataset
```
df.columns.values
df = df[df.He_por >= 0]
dataset = df[[
'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID', 'GR_smooth', 'PE_smooth',
'He_por'
]]
```
In the next code block, we will remove the rows without data, and change string NaN's to np.nans
```
dataset.replace('NaN',np.nan, regex=True, inplace=True)#
np.shape(dataset)
dataset.head(3)
X = dataset[[ 'CAL', 'GR', 'DT', 'SP', 'DENS', 'PE',
'RESD', 'PHIN', 'PHID', 'GR_smooth', 'PE_smooth']]
Y = dataset[['He_por']]
Y_array = np.array(Y.values)
```
## Starting to set up the ML model params
```
seed = 7 # random seed is only used if you want to compare exact answers with friends
test_size = 0.25 # how much data you want to withold, .15 - 0.3 is a good starting point
X_train, X_test, y_train, y_test = train_test_split(X.values, Y_array, test_size=test_size)
regr = linear_model.LinearRegression()
regr.fit(X_train, y_train)
print('Intercept: \n', regr.intercept_)
print('Coefficients: \n', regr.coef_)
preds = regr.predict(X_test)
rmse5 = mean_squared_error(y_test, preds, squared=False)
print("Mean Squared Error: %f" % (rmse5))
max5 = max_error(y_test, preds)
print("Max Error: %f" % (max5))
MAE2 = median_absolute_error(y_test, preds)
print("Median Abs Error: %f" % (MAE2))
x = datetime.datetime.now()
d = {'target': [Y.columns.values],
'MSE': [rmse5 ],
'MAE': [MAE2],
'MaxError': [max5],
'day': [x.day],
'month':[x.month],
'year':[x.year],
'model':['MVR'],
'version':[sklearn.__version__]}
results = pd.DataFrame(data=d)
results.to_csv('drive/My Drive/1_lewis_research/analysis/experiments/mvr/mvr_results/por_mvr.csv')
results
from sklearn.metrics import explained_variance_score
explained_variance_score(y_test, preds, multioutput='uniform_average')
```
| github_jupyter |
```
import pandas as pd
import seaborn as sns
%matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.colors as clr
import numpy as np
from matplotlib.colors import ListedColormap
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import cm
from matplotlib.colors import ListedColormap, LinearSegmentedColormap
import pandas as pd
import matplotlib.pylab as plt
import numpy as np
import scipy
import seaborn as sns
import glob
df = pd.read_csv("9112_combined_A_scores.csv")
df['chr'] = df['name'].apply(lambda x:x.split(":")[0])
df['start'] = df['name'].apply(lambda x:x.split(":")[1].split("-")[0])
df['end'] = df['name'].apply(lambda x:x.split(":")[1].split("-")[1])
print (df.head())
rename_cols = []
for c in df.columns:
if "HbFBase" in c:
rename_cols.append("HbFBase")
else:
rename_cols.append(c)
df.columns = rename_cols
names = ['HbFBase','CADD',"DeepSEA"]
for n in names:
df['%s_rank'%(n)] = -df[n].rank(ascending=False)
df.head()
df['start'] = df['start'].astype(int)
df = df.sort_values(['chr','start'])
df.head()
def to_matrix(df,n,top=500):
c=n
size=100
tmp = df.copy()
tmp = tmp.sort_values(n,ascending=False).head(n=top)
my_index_list = tmp.index.tolist()
out = []
for i in my_index_list:
line = []
current_chr = df.at[i,"chr"]
for j in range(-size,size+1):
try:
chr = df.at[i+j,"chr"]
except:
# print (i+j)
chr = "None"
if chr == current_chr:
value = df.at[i+j,"%s_rank"%(c)]
else:
value = np.nan
line.append(value)
out.append(line)
out_df = pd.DataFrame(out)
sel_cols = out_df.columns.tolist()
# print (out_df.head())
out_df.index = my_index_list
out_df['chr'] = tmp['chr']
out_df['start'] = tmp['start']
out_df['end'] = tmp['end']
out_df['name'] = tmp['name']
out_df['value'] = "."
out_df['strand'] = "."
print (df["%s_rank"%(c)].mean())
out_df = out_df.fillna(df["%s_rank"%(c)].mean())
out_df[['chr','start','end','name','value','strand']+sel_cols].to_csv("%s.computeMatrix.bed"%(c),header=False,index=False,sep="\t")
return out_df[sel_cols]
color_dict ={}
color_dict['HbFBase']='red'
color_dict['CADD']='green'
color_dict['DeepSEA']='blue'
fig, ax = plt.subplots()
for n in names:
print (n)
result_df = to_matrix(df,n)
mean_line = pd.DataFrame(result_df.mean())
# sns.lineplot(data = mean_line)
test = pd.melt(result_df)
sns.lineplot(x="variable", y="value", data=test,c=color_dict[n],ax=ax,label=n)
ax.set_xticklabels(['']+list(range(-100,101,25)))
ax.set_yticklabels(['']+list(range(5000,999,-1000))+[1])
plt.ylabel("Average rank")
plt.xlabel("Downstream / Upstream neighbors")
plt.savefig("Score_ranks_comparison_top500.pdf", bbox_inches='tight')
```
| github_jupyter |
```
%pylab inline
%load_ext autoreload
%autoreload 2
import sys
sys.path.insert(0,'..')
from tensorflow_probability.python.internal.backend import jax as tf
import tensorflow_probability as tfp; tfp = tfp.experimental.substrates.jax
from jax_nf.real_nvp import RealNVP
from flax import nn
import jax
import jax.numpy as np
import numpy as onp
import flax
tfd = tfp.distributions
tfb = tfp.bijectors
@nn.module
def AffineCoupling(x, nunits, apply_scaling=True):
# Let's have one more non stupid lyerre
net = nn.leaky_relu(nn.Dense(x, 128))
net = nn.leaky_relu(nn.Dense(net, 128))
shift = nn.Dense(net, nunits)
if apply_scaling:
scaler = tfb.Scale(np.clip(np.exp(nn.Dense(net, nunits)), 1e-3, 1e3))
else:
scaler = tfb.Identity()
return tfb.Chain([ tfb.Shift(shift), scaler])
# Instantiate the splines
d = 2
dummy_input = np.zeros((1, d//2))
_, params1 = AffineCoupling.init(jax.random.PRNGKey(0), dummy_input, d//2)
from functools import partial
affine1 = partial(AffineCoupling.call, params1)
nvp = tfd.TransformedDistribution(
tfd.Normal(0,1),
bijector=RealNVP(1, bijector_fn=affine1),
event_shape=(2,))
samps = nvp.sample((1000,), seed=jax.random.PRNGKey(1))
hist2d(samps[:,0], samps[:,1],64);
# Sweet :-D Hurray for TFP
# Let's try to learn a density
d=2
@nn.module
def AffineFlow(x):
affine1 = AffineCoupling.shared(name='affine1')
affine2 = AffineCoupling.shared(name='affine2')
affine3 = AffineCoupling.shared(name='affine3',apply_scaling=False)
affine4 = AffineCoupling.shared(name='affine4',apply_scaling=False)
affine5 = AffineCoupling.shared(name='affine5',apply_scaling=False)
affine6 = AffineCoupling.shared(name='affine6',apply_scaling=False)
# Computes the likelihood of these x
chain = tfb.Chain([
RealNVP(d//2, bijector_fn=affine1),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine2),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine3),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine4),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine5),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine6)
])
nvp = tfd.TransformedDistribution(
tfd.Normal(0,1),
bijector=chain,
event_shape=(d,))
return nvp.log_prob(x)
dummy_input = np.zeros((1,d))
res, params = AffineFlow.init(jax.random.PRNGKey(0), dummy_input)
model = nn.Model(AffineFlow, params)
# Ok, sweet
@jax.jit
def train_step(optimizer, batch):
def loss_fn(model):
log_prob = model(batch['x'])
return -np.mean(log_prob)
loss, grad = jax.value_and_grad(loss_fn)(optimizer.target)
optimizer = optimizer.apply_gradient(grad)
return loss, optimizer
# Now let's draw ou famous two moons
from sklearn import datasets
batch_size=1024
def get_batch():
x, y = datasets.make_moons(n_samples=batch_size, noise=.05)
return {'x': x}
# okokokok, let's try it out
optimizer = flax.optim.Adam(learning_rate=0.001).create(model)
losses = []
for i in range(2000):
batch = get_batch()
l, optimizer = train_step(optimizer, batch)
losses.append(l)
if i %100 ==0:
print(l)
plot(losses[100:])
# ok, fine, let's see if we can rebuild our flow
@nn.module
def AffineFlowSampler(key, n_samples):
affine1 = AffineCoupling.shared(name='affine1')
affine2 = AffineCoupling.shared(name='affine2')
affine3 = AffineCoupling.shared(name='affine3',apply_scaling=False)
affine4 = AffineCoupling.shared(name='affine4',apply_scaling=False)
affine5 = AffineCoupling.shared(name='affine5',apply_scaling=False)
affine6 = AffineCoupling.shared(name='affine6',apply_scaling=False)
# Computes the likelihood of these x
chain = tfb.Chain([
RealNVP(d//2, bijector_fn=affine1),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine2),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine3),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine4),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine5),
tfb.Permute([1,0]),
RealNVP(d//2, bijector_fn=affine6)
])
nvp = tfd.TransformedDistribution(
tfd.Normal(0,1),
bijector=chain,
event_shape=(d,))
return nvp.sample(n_samples, seed=key)
sampler = nn.Model(AffineFlowSampler, optimizer.target.params)
samps = sampler(jax.random.PRNGKey(1),1000)
hist2d(samps[:,0], samps[:,1],64, [[-1.5,2.5],[-1.,1.5]]);
```
| github_jupyter |
<a href="https://colab.research.google.com/github/satyajitghana/TSAI-DeepNLP-END2.0/blob/main/14_BERT_BART/BART_For_Paraphrasing_w_Simple_Transformers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
! pip3 install simpletransformers==0.61.13 --quiet
%%bash
mkdir data
wget https://storage.googleapis.com/paws/english/paws_wiki_labeled_final.tar.gz -P data
tar -xvf data/paws_wiki_labeled_final.tar.gz -C data
mv data/final/* data
rm -r data/final
wget http://qim.fs.quoracdn.net/quora_duplicate_questions.tsv -P data
%%file utils.py
import warnings
import pandas as pd
def load_data(
file_path, input_text_column, target_text_column, label_column, keep_label=1
):
df = pd.read_csv(file_path, sep="\t", error_bad_lines=False)
df = df.loc[df[label_column] == keep_label]
df = df.rename(
columns={input_text_column: "input_text", target_text_column: "target_text"}
)
df = df[["input_text", "target_text"]]
df["prefix"] = "paraphrase"
return df
def clean_unnecessary_spaces(out_string):
if not isinstance(out_string, str):
warnings.warn(f">>> {out_string} <<< is not a string.")
out_string = str(out_string)
out_string = (
out_string.replace(" .", ".")
.replace(" ?", "?")
.replace(" !", "!")
.replace(" ,", ",")
.replace(" ' ", "'")
.replace(" n't", "n't")
.replace(" 'm", "'m")
.replace(" 's", "'s")
.replace(" 've", "'ve")
.replace(" 're", "'re")
)
return out_string
import os
from datetime import datetime
import logging
import pandas as pd
from sklearn.model_selection import train_test_split
from simpletransformers.seq2seq import Seq2SeqModel, Seq2SeqArgs
from utils import load_data, clean_unnecessary_spaces
logging.basicConfig(level=logging.INFO)
transformers_logger = logging.getLogger("transformers")
transformers_logger.setLevel(logging.ERROR)
# Google Data
train_df = pd.read_csv("data/train.tsv", sep="\t").astype(str)
eval_df = pd.read_csv("data/dev.tsv", sep="\t").astype(str)
train_df = train_df.loc[train_df["label"] == "1"]
eval_df = eval_df.loc[eval_df["label"] == "1"]
train_df = train_df.rename(
columns={"sentence1": "input_text", "sentence2": "target_text"}
)
eval_df = eval_df.rename(
columns={"sentence1": "input_text", "sentence2": "target_text"}
)
train_df = train_df[["input_text", "target_text"]]
eval_df = eval_df[["input_text", "target_text"]]
train_df["prefix"] = "paraphrase"
eval_df["prefix"] = "paraphrase"
# Quora Data
# The Quora Dataset is not separated into train/test, so we do it manually the first time.
df = load_data(
"data/quora_duplicate_questions.tsv", "question1", "question2", "is_duplicate"
)
q_train, q_test = train_test_split(df)
q_train.to_csv("data/quora_train.tsv", sep="\t")
q_test.to_csv("data/quora_test.tsv", sep="\t")
# The code block above only needs to be run once.
# After that, the two lines below are sufficient to load the Quora dataset.
# q_train = pd.read_csv("data/quora_train.tsv", sep="\t")
# q_test = pd.read_csv("data/quora_test.tsv", sep="\t")
train_df = pd.concat([train_df, q_train])
eval_df = pd.concat([eval_df, q_test])
train_df = train_df[["prefix", "input_text", "target_text"]]
eval_df = eval_df[["prefix", "input_text", "target_text"]]
train_df = train_df.dropna()
eval_df = eval_df.dropna()
train_df["input_text"] = train_df["input_text"].apply(clean_unnecessary_spaces)
train_df["target_text"] = train_df["target_text"].apply(clean_unnecessary_spaces)
eval_df["input_text"] = eval_df["input_text"].apply(clean_unnecessary_spaces)
eval_df["target_text"] = eval_df["target_text"].apply(clean_unnecessary_spaces)
len(train_df), len(eval_df)
train_df
train_df_sample = train_df.sample(frac=0.2, random_state=42)
eval_df_sample = eval_df.sample(frac=0.05, random_state=42)
len(train_df_sample), len(eval_df_sample)
model_args = Seq2SeqArgs()
model_args.do_sample = True
model_args.eval_batch_size = 16
model_args.evaluate_during_training = True
model_args.evaluate_during_training_steps = 2500
model_args.evaluate_during_training_verbose = True
model_args.fp16 = False
model_args.learning_rate = 5e-5
model_args.max_length = 128
model_args.max_seq_length = 128
model_args.num_beams = None
model_args.num_return_sequences = 3
model_args.num_train_epochs = 1
model_args.overwrite_output_dir = True
model_args.reprocess_input_data = True
model_args.save_eval_checkpoints = False
model_args.save_steps = -1
model_args.top_k = 50
model_args.top_p = 0.95
model_args.train_batch_size = 16
model_args.use_multiprocessing = False
model_args.wandb_project = "Paraphrasing with BART"
model = Seq2SeqModel(
encoder_decoder_type="bart",
encoder_decoder_name="facebook/bart-base",
args=model_args,
)
model.train_model(train_df_sample, eval_data=eval_df_sample)
model.eval_batch_size = 16
%env TOKENIZERS_PARALLELISM=0
model.eval_model(eval_df_sample)
```
W&B Training Logs: https://wandb.ai/satyajit_meow/Paraphrasing%20with%20BART?workspace=user-satyajit_meow
```
model = Seq2SeqModel(
encoder_decoder_type="bart",
encoder_decoder_name="outputs/checkpoint-1673-epoch-1",
args=model_args,
)
to_predict = [
prefix + ": " + str(input_text)
for prefix, input_text in zip(eval_df_sample["prefix"].tolist(), eval_df_sample["input_text"].tolist())
]
truth = eval_df_sample["target_text"].tolist()
preds = model.predict(to_predict)
to_predict[:5]
preds[:5]
# Saving the predictions if needed
os.makedirs("predictions", exist_ok=True)
with open(f"predictions/predictions_{datetime.now()}.txt", "w") as f:
for i, text in enumerate(to_predict):
f.write(str(text).split(':', 1)[1].strip() + "\n\n")
f.write("Truth:\n")
f.write(truth[i] + "\n\n")
f.write("Prediction:\n")
for pred in preds[i]:
f.write(str(pred) + "\n")
f.write(
"________________________________________________________________________________\n"
)
import random
for i, text in enumerate(to_predict[:128]):
print(f'Text > {str(text).split(":", 1)[1].strip()}')
print(f'Pred < {preds[i][0]}')
print(f'Truth = {truth[i]}')
print()
```
| github_jupyter |
### Time_Series_Load_and_Explore_Data
Jay Urbain, PhD
Credits:
- Introduction to Time Series Forecasting with Python, Jason Brownlee.
- Python Data Science Handbook, Jake VanderPlas.
- Chris Albion, https://chrisalbon.com/python/data_wrangling/pandas_time_series_basics/
We will use the Daily Female Births Dataset as an example. This dataset describes the number of daily female births in California in 1959. The units are a count and there are 365 observations. The source of the dataset is credited to Newton (1988).
```
datadir = '/Users/jayurbain/Dropbox/Python-time-series-forecasting/time_series_forecasting_with_python/code/chapter_04'
```
#### Load Daily Female Births Dataset
Pandas represented time series datasets as a Series. A Series1 is a one-dimensional array with a time label for each row.
http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.html
The series has a name, which is the column name of the data column. You can see that each row has an associated date. This is in fact not a column, but instead a time index for value. As an index, there can be multiple values for one time, and values may be spaced evenly or unevenly across times. The main function for loading CSV data in Pandas is the read csv() function. We can use this to load the time series as a Series object, instead of a DataFrame, as follows:
```
# load dataset using read_csv()
from pandas import read_csv
series = read_csv(datadir + '/' + 'daily-total-female-births.csv', header=0, parse_dates=[0], index_col=0, squeeze=True)
print(type(series))
print(series.head())
```
Note the arguments to the read csv() function. We provide it a number of hints to ensure the data is loaded as a Series.
- header=0: We must specify the header information at row 0.
- parse dates=[0]: We give the function a hint that data in the first column contains dates that need to be parsed. This argument takes a list, so we provide it a list of one element, which is the index of the first column.
- index col=0: We hint that the first column contains the index information for the time series.
- squeeze=True: We hint that we only have one data column and that we are interested in a Series and not a DataFrame.
#### Exploring Time Series Data
```
# summarize first few lines of a file
series.head(10)
print(series.tail(10))
# summarize the dimensions of a time series
from pandas import Series
print(series.size)
```
#### Querying By Time
```
print(series.loc['1959-01'])
```
#### Descriptive Statistics
Calculating descriptive statistics on your time series can help get an idea of the distribution and spread of values. This may help with ideas of data scaling and even data cleaning that you can perform later as part of preparing your dataset for modeling. The describe() function creates a 7 number summary of the loaded time series including mean, standard deviation, median, minimum, and maximum of the observations.
```
print(series.describe())
```
#### Using pandas
```
from datetime import datetime
import pandas as pd
%matplotlib inline
import matplotlib.pyplot as pyplot
```
Create a dataframe
```
data = {'date': ['2014-05-01 18:47:05.069722', '2014-05-01 18:47:05.119994', '2014-05-02 18:47:05.178768', '2014-05-02 18:47:05.230071', '2014-05-02 18:47:05.230071', '2014-05-02 18:47:05.280592', '2014-05-03 18:47:05.332662', '2014-05-03 18:47:05.385109', '2014-05-04 18:47:05.436523', '2014-05-04 18:47:05.486877'],
'battle_deaths': [34, 25, 26, 15, 15, 14, 26, 25, 62, 41]}
df = pd.DataFrame(data, columns = ['date', 'battle_deaths'])
print(df)
```
Convert df['date'] from string to datetime
```
df['date'] = pd.to_datetime(df['date'])
```
Set df['date'] as the index and delete the column
```
df.index = df['date']
del df['date']
df
```
View all observations that occured in 2014
```
df['2014']
```
View all observations that occured in May 2014
```
df['2014-05']
```
Observations after May 3rd, 2014
```
df[datetime(2014, 5, 3):]
```
Count the number of observations per timestamp
```
df.groupby(level=0).count()
```
Mean value of battle_deaths per day
```
df.resample('D').mean()
```
Total value of battle_deaths per day
```
df.resample('D').sum()
```
Plot of the total battle deaths per day
```
df.resample('D').sum().plot()
```
| github_jupyter |
```
library(tidyverse)
library(cowplot)
library(gridExtra)
`%+replace%` <- ggplot2::`%+replace%`
theme_zietzm <- function(base_size = 11.5, base_family = "") {
# Starts with theme_bw and then modify some parts
# Theme options are documentated at http://docs.ggplot2.org/current/theme.html
ggplot2::theme_classic(base_size = base_size, base_family = base_family) %+replace%
ggplot2::theme(
strip.background = ggplot2::element_rect(fill = NA, colour = 'grey90', size = 0),
strip.text = element_text(vjust = 1, size = 10),
plot.margin = ggplot2::margin(t=2, r=2, b=2, l=2, unit='pt'),
legend.spacing = grid::unit(0.1, 'cm'),
legend.key = ggplot2::element_blank(),
panel.border=element_rect(fill = NA, color = 'black', size = 0.5),
axis.line=element_line(size=0),
)
}
```
# 1. Subfigure A
Differences in overall degree distributions for networks of the same kind of data
```
ppi_df <- read_tsv('../../data/task3/3.all_nodes/ppi.tsv.xz') %>%
select(-starts_with('name')) %>%
gather('id_side', 'id', id_a:id_b) %>%
group_by(.dots=c("id")) %>%
summarize_at(vars(train, test_new), funs(sum)) %>%
rename(biased=train, unbiased=test_new) %>%
gather('network_type', 'degree', biased:unbiased) %>%
mutate(name = 'ppi')
tftg_df <- read_tsv('../../data/task3/3.all_nodes/tftg.tsv.xz') %>%
select(-starts_with('name')) %>%
gather('id_side', 'id', id_a:id_b) %>%
group_by(.dots=c("id_side", "id")) %>%
summarize_at(vars(train, test_new), funs(sum)) %>%
rename(biased=train, unbiased=test_new) %>%
gather('network_type', 'degree', biased:unbiased) %>%
mutate(name = id_side %>% recode(id_a = 'tftg_source', id_b = 'tftg_target')) %>%
ungroup() %>%
select(-id_side)
biorxiv_df <- read_tsv('../../data/task3/3.all_nodes/biorxiv.tsv.xz') %>%
select(-starts_with('name')) %>%
gather('id_side', 'id', id_a:id_b) %>%
group_by(.dots=c("id")) %>%
summarize_at(vars(train, test_new), funs(sum)) %>%
rename(biased=train, unbiased=test_new) %>%
gather('network_type', 'degree', biased:unbiased) %>%
mutate(
name = 'co_author',
network_type = network_type %>% recode(biased = "<2018", unbiased = ">=2018")
)
histogram_vis_df <- bind_rows(ppi_df, tftg_df, biorxiv_df) %>%
mutate(
name = name %>% recode_factor(
ppi = 'PPI',
tftg_source = 'TF-TG: TF',
tftg_target = 'TF-TG: TG',
co_author = 'Co-authorship',
),
network_type = network_type %>% recode_factor(
biased = 'Literature-derived',
unbiased = 'Systematic',
)
)
head(histogram_vis_df, 2)
histogram_labels <- data.frame(
name = c('PPI', 'PPI', 'TF-TG: TF', 'TF-TG: TF', 'TF-TG: TG', 'TF-TG: TG',
'Co-authorship', 'Co-authorship'),
x = c(250, 50, 50, 300, 8, 25, 15, 25),
y = c(550, 2000, 40, 12, 500, 200, 1500, 500),
label = factor(c('Literature-derived', 'Systematic',
'Literature-derived', 'Systematic',
'Literature-derived', 'Systematic',
'<2018', '>=2018'), levels = c('Systematic', 'Literature-derived', '<2018', '>=2018')),
network_type = factor(c('Literature-derived', 'Systematic',
'Literature-derived', 'Systematic',
'Literature-derived', 'Systematic',
'<2018', '>=2018'), levels = c('Literature-derived', 'Systematic', '<2018', '>=2018'))
)
histogram_dists <- (
ggplot(histogram_vis_df, aes(x = degree, fill = network_type))
+ geom_histogram(position = position_identity(), alpha = 0.5, bins = 50)
+ facet_wrap("name", scales = 'free', nrow = 2)
+ scale_fill_brewer(palette = "Set1")
+ scale_color_brewer(palette = "Set1")
+ ylab('Nodes')
+ xlab('Node degree')
+ theme_zietzm()
+ labs(fill = 'Network type')
+ theme(legend.position = "none")
+ geom_text(data = histogram_labels, aes(x = x, y = y, label = label, color = network_type),
alpha = 0.8, size = 4, nudge_x = -5, hjust = 'left')
)
```
# 2. Subfigure B
Differences in node degree by sampling method
```
format_degrees <- function(df) {
# Helper function to compute degree
df %>%
group_by(id_a) %>%
summarise(
degree_a_train = sum(train),
degree_a_test_recon = sum(test_recon),
degree_a_test_new = sum(test_new)
) %>%
full_join(
df %>%
group_by(id_b) %>%
summarise(
degree_b_train = sum(train),
degree_b_test_recon = sum(test_recon),
degree_b_test_new = sum(test_new)
),
by = c("id_a" = "id_b")
) %>%
replace(is.na(.), 0) %>%
mutate(
degree_train = degree_a_train + degree_b_train,
degree_test_recon = degree_a_test_recon + degree_b_test_recon,
degree_test_new = degree_a_test_new + degree_b_test_new
) %>%
select(degree_train, degree_test_recon, degree_test_new)
}
ppi_df <- read_tsv('../../data/task3/2.edges//ppi.tsv.xz')
tftg_df <- read_tsv('../../data/task3/2.edges//tftg.tsv.xz')
head(ppi_df, 2)
df <- bind_rows(
ppi_df %>%
format_degrees %>%
mutate(network = 'PPI'),
tftg_df %>%
format_degrees %>%
mutate(network = 'TF-TG')
)
vis_df <- bind_rows(
df %>%
select(x = degree_test_recon, y = degree_train, network) %>%
mutate(task = 'Reconstruction'),
df %>%
select(x = degree_test_recon, y = degree_test_new, network) %>%
mutate(task = 'Systematic')
) %>%
mutate(task = task %>% recode_factor(Reconstruction = 'Subsampled holdout',
Systematic = 'Systematic holdout'))
palette <- 'Spectral'
direction <- -1
degree_bias <- (
ggplot(vis_df, aes(x = x + 1, y = y + 1))
+ stat_binhex(aes(color = ..count..), bins = 15)
+ geom_abline(slope = 1, intercept = 0, color = 'black', linetype = 'dashed')
+ xlab('Literature degree')
+ ylab('Holdout degree')
+ facet_grid(rows = vars(network), cols = vars(task))
+ scale_x_log10()
+ scale_y_log10()
+ theme_zietzm()
+ scale_fill_distiller(palette = palette, direction = direction, name = 'Nodes')
+ scale_color_distiller(palette = palette, direction = direction, name = 'Nodes')
+ coord_fixed(xlim = c(1, 1000), ylim = c(1, 1000), ratio = 1)
+ theme(legend.position = c(0.875, 0.08),
legend.background = element_rect(fill = NA, colour = NA),
legend.direction = 'horizontal',
legend.text = element_text(size = 7),
legend.key.size = unit(0.35, "cm"),
legend.title = element_text(size = 9),
legend.title.align = 0.5,
)
+ guides(fill = guide_colorbar(title.position = 'top'),
color = guide_colorbar(title.position = 'top'))
)
options(repr.plot.width=5, repr.plot.height=4)
degree_bias
```
# 3. Save figure
```
options(repr.plot.width=9, repr.plot.height=4)
combined <- plot_grid(
histogram_dists,
degree_bias + theme(plot.margin = unit(c(0.1, -1, 0.05, -0.5), 'cm')),
labels = c("A", "B")
)
ggsave('../../img/degree_bias.png', combined, width = 9, height = 4, dpi = 400)
combined
```
| github_jupyter |
# Compare decay calculation results of radioactivedecay high precision mode and PyNE
This notebook compares decay calculation results between the Python package [radioactivedecay](https://pypi.org/project/radioactivedecay/) `v0.4.2`, using its high precision mode, and [PyNE](https://pyne.io/) `v0.7.5`. The PyNE decay data is read in by radioactivedecay, so both codes are using the same underlying decay data for the calculations.
Note the following radionuclides were removed from the decay dataset fed to radioactivedecay, as these radionuclides are part of chains where two radionuclides have degenerate half-lives: Po-191, Pt-183, Bi-191, Pb-187, Tl-187, Rn-195, At-195, Hg-183, Au-183, Pb-180, Hg-176, Tl-177, Pt-172, Ir-172, Lu-153 and Ta-157. radioactivedecay cannot calculate decays for chains containing radionuclides with identical half-lives, and the PyNE treatment for these chains currently suffers from a [bug](https://github.com/pyne/pyne/issues/1342).
First import the necessary modules for the comparison.
```
import radioactivedecay as rd
import pyne
from pyne import nucname, data
from pyne.material import Material, from_activity
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
print("Package versions used: radioactivedecay", rd.__version__, "PyNE", pyne.__version__)
```
### Load PyNE decay dataset into radioactivedecay, check a simple case
Load the decay dataset created using the PyNE data into radioactivedecay.
```
pynedata = rd.DecayData(dataset_name="pyne_truncated", dir_path=".", load_sympy=True)
```
Define some functions to perform radioactive decay calculations for a single radionuclide with radioactivedecay and PyNE. To compare the results we have to remove the stable radionuclides from the inventory returned by PyNE. We also have to convert the canonical radionuclide ids into string format (e.g. `10030000` to `H-3`), and sort the inventory alphabetically.
```
def rd_decay(nuclide, time):
"""Perform a decay calculation for one radionuclide with radioactivedecay."""
return rd.InventoryHP({nuclide: 1.0}, decay_data=pynedata).decay(time).activities()
def add_hyphen(name):
"""Add a hypen to radionuclide name string e.g. H3 to H-3."""
for i in range(1, len(name)):
if not name[i].isdigit(): continue
name = name[:i] + "-" + name[i:]
break
return name
def strip_stable(inv):
"""Remove stable nuclides from a PyNE inventory."""
new_inv = dict()
for id in inv:
if data.decay_const(id) <= 0.0: continue
new_inv[add_hyphen(nucname.name(id))] = inv[id]
return new_inv
def pyne_decay(nuclide, time):
"""Perform a decay calculation for one radionuclide with PyNE."""
inv = strip_stable(from_activity({nucname.id(nuclide): 1.0}).decay(time).activity())
return dict(sorted(inv.items(), key=lambda x: x[0]))
```
First let's compare the decay results for a single case - the decay of 1 unit (Bq or Ci) of Pb-212 for 0.0 seconds:
```
inv_rd = rd_decay("Pb-212", 0.0)
inv_pyne = pyne_decay("Pb-212", 0.0)
print(inv_rd)
print(inv_pyne)
```
radioactivedecay returns activities for all the radionuclides in the decay chain below Pb-212, even though they have zero activity. PyNE does not return activities for two of the radionuclides (Bi-212 and Po-212), presumably because it evaluated their activities to be exactly zero. A small non-zero activity is returned for Ti-208, and the activity of Pb-212 deviates slightly from unity. The likely explanation for these results is round-off errors.
For the below comparisons, it makes sense to compare the activities of the radionuclides returned by both radioactivedecay and PyNE, whilst checking the activities of the radionuclides missing from the inventories returned by either radioactivedecay or PyNE are negligible. This function cuts down the inventories returned by radioactivedecay and PyNE to just the radionuclides present in both inventories. It also reports how many radionuclides were removed from each inventory to do this.
```
def match_inventories(inv1, inv2):
"""Cuts down the two inventories so they only contain the radionuclides present in both.
Also returns inventories of the radionuclides unique to each inventory."""
s1 = set(inv1.keys())
s2 = set(inv2.keys())
s = s1.intersection(s2)
inv1_intersection = {nuclide: inv1[nuclide] for nuclide in s}
inv2_intersection = {nuclide: inv2[nuclide] for nuclide in s}
inv1_difference = {nuclide: inv1[nuclide] for nuclide in s1.difference(s2)}
inv2_difference = {nuclide: inv2[nuclide] for nuclide in s2.difference(s1)}
return inv1_intersection, inv1_difference, inv2_intersection, inv2_difference
```
### Generate decay calculation comparisons between radioactivedecay and PyNE
We now systematically compare the results of decay calculations performed using radioactivedecay and PyNE. The strategy is the set initial inventories containing 1 Bq of each radionuclide in the decay dataset, and then decay for various time periods that are factor multiples of that radionuclides half-life. The factor multiples used are zero and each order of magnitude between 10<sup>-6</sup> and 10<sup>6</sup>, inclusive.
We calculate the absolute activity error for each radionuclide returned by both radioactivedecay and PyNE in the decayed inventories, as well as the relative activity error to the PyNE activity. We store the results in a Pandas DataFrame. We also store the results for the radionuclides that are not returned in either the radioactivedecay or PyNE inventories for examination.
Note: this next code block may take over an hour to run.
```
hl_factors = [0.0, 1e-6, 1e-5, 1e-4, 1e-3, 1e-2, 1e-1, 1e0, 1e1, 1e2, 1e3, 1e4, 1e5, 1e6]
rows = []
rows_pyne_missing = []
rows_rd_missing = []
for hl_factor in hl_factors:
for nuclide in pynedata.nuclides:
decay_time = hl_factor*rd.Nuclide(nuclide, decay_data=pynedata).half_life()
rd_inv = rd_decay(nuclide, decay_time)
pyne_inv = pyne_decay(nuclide, decay_time)
rd_inv, rd_unique, pyne_inv, pyne_unique = match_inventories(rd_inv, pyne_inv)
if len(rd_unique) != 0:
for nuc, act in rd_unique.items():
rows_pyne_missing.append({
"parent": nuclide,
"hl_factor": hl_factor,
"decay_time": decay_time,
"pyne_missing_nuclide": nuc,
"rd_activity": act
})
if len(pyne_unique) != 0:
for nuc, act in pyne_unique.items():
rows_rd_missing.append({
"parent": nuclide,
"hl_factor": hl_factor,
"decay_time": decay_time,
"rd_missing_nuclide": nuc,
"pyne_activity": act
})
for nuc, pyne_act in pyne_inv.items():
rd_act = rd_inv[nuc]
if pyne_act == 0.0:
if rd_act == 0.0:
abs_err = 0.0
rel_err = 0.0
else:
abs_err = abs(rd_act)
rel_err = np.inf
else:
abs_err = abs(pyne_act-rd_act)
rel_err = abs_err/abs(pyne_act)
rows.append({
"parent": nuclide,
"hl_factor": hl_factor,
"decay_time": decay_time,
"nuclide": nuc,
"pyne_activity": pyne_act,
"rd_activity": rd_act,
"abs_err": abs_err,
"rel_err": rel_err,
})
print(hl_factor, "complete")
df = pd.DataFrame(rows)
df_pyne_missing = pd.DataFrame(rows_pyne_missing)
df_rd_missing = pd.DataFrame(rows_rd_missing)
```
### Examine cases where radionuclides not returned by radioactivedecay and PyNE
First we check the cases where radionuclides returned by PyNE are not present in decayed inventory from radioactivedecay.
```
print("Radionuclides not returned by radioactivedecay:", df_rd_missing.rd_missing_nuclide.unique())
print("These cases arise from the decay chains of:", df_rd_missing.parent.unique())
print("Total number of missing cases:", len(df_rd_missing))
```
radioactivedecay does not return activities for these radionuclides as the chains all pass through radionuclides which PyNE reports as having an undefined half-life:
```
print(data.half_life("He-9"), data.half_life("W-192"), data.half_life("Dy-170"), data.half_life("H-5"))
```
PyNE clearly has some assumption to infer the half-lives and simulate these decay chains.
Now we check the radionuclides not returned by PyNE:
```
print("Total number of missing cases:", len(df_pyne_missing))
df_pyne_missing.sort_values(by=["rd_activity"], ascending=False).head(n=10)
```
The maximum activity of any radionuclide returned by radioactivedecay but not returned by PyNE is 2.4E-14 Bq. The activities in all other cases are lower than this. This could be related to PyNE filtering out any results it considers negligible.
### Comparing decayed activities between radioactivedecay and PyNE
We now compare the decayed activities of radionuclides returned both by radioactivedecay and by PyNE. For the 237095 comparisons, the mean and maximum absolute errors are 1.9E-17 Bq and 5.8E-14 Bq, respectively:
```
df.describe()
df.sort_values(by=["abs_err", "rel_err", "pyne_activity"], ascending=False).head(n=10)
```
For around 8% of the activities compared the activity calculated by radioactivedecay and PyNE are identical:
```
len(df[df.abs_err == 0.0])
```
Now plot the errors. For cases where the PyNE activity is 0.0, we set these activity values to 10<sup>-70</sup> and the relative errors to 10<sup>17</sup> to force the points to show on the panel (a) log-log graph.
```
df.loc[(df["pyne_activity"] == 0.0) & (df["rd_activity"] != 0.0), "pyne_activity"] = 1e-70
df.loc[df["rel_err"] == np.inf, "rel_err"] = 1e17
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(12.8,4.8))
cmap_colors = plt.get_cmap("plasma").colors
colors = ["black"]
n = len(hl_factors)-1
colors.extend([""] * n)
for i in range(0, n):
colors[i+1] = cmap_colors[int(round((i*(len(cmap_colors)-1)/(n-1))))]
for i in range(0, len(hl_factors)):
ax[0].plot(df[df.hl_factor == hl_factors[i]].pyne_activity, df[df.hl_factor == hl_factors[i]].rel_err,
marker=".", linestyle="", label=hl_factors[i], color=colors[i])
ax[0].set(xlabel="PyNE activity (Bq)", ylabel="relative error", xscale="log", yscale="log")
ax[0].legend(loc="upper left", title="half-life factor")
ax[0].text(-0.15, 1.0, "(a)", transform=ax[0].transAxes)
ax[0].set_xlim(1e-104, 1e4)
ax[0].set_ylim(1e-18, 1e18)
for i in range(0, len(hl_factors)):
ax[1].plot(df[df.hl_factor == hl_factors[i]].abs_err, df[df.hl_factor == hl_factors[i]].rel_err,
marker=".", linestyle="", label=hl_factors[i], color=colors[i])
ax[1].set(xlabel="absolute error", ylabel="relative error", xscale="log", yscale="log")
ax[1].legend(loc="upper right", title="half-life factor")
ax[1].set_xlim(1e-25, 1e-7)
_ = ax[1].text(-0.15, 1.0, "(b)", transform=ax[1].transAxes)
```
In all cases the differences in activities reported by radioactivedecay and by PyNE are small (< 1E-13 Bq). Relative errors tend to increase as the radioactivity reported by PyNE decreases from 1 Bq. Relative errors greater than 1E-4 only occur when the PyNE activity is smaller than 2.5E-11.
```
df[df.rel_err > 1E-4].pyne_activity.max()
```
Export the DataFrames to CSV files:
```
df.to_csv('radioactivedecay_high_precision_pyne.csv')
df_pyne_missing.to_csv('radioactivedecay_high_precision_pyne_pyne_missing.csv')
df_rd_missing.to_csv('radioactivedecay_high_precision_pyne_rd_missing.csv')
```
### Summary
The activity results reported by radioactivedecay and PyNE differ by less than 1E-13 Bq, given an initial inventory of 1 Bq of the parent radionuclide. PyNE used double precision floating point arithmetic for its decay calculations, while radioactivedecay used SymPy computations with a precision of 320 significant figures. PyNE used some slightly different treatments than radioactivedecay for calculating decay chains passing through almost stable radionuclides, and for filtering out radionuclides with negligible activities.
In summary we can the results calculated by radioactivedecay and PyNE using the PyNE decay dataset are identical to within reasonable expectations for numerical precision.
| github_jupyter |
```
from pyspark.sql import SparkSession
spark = SparkSession \
.builder \
.appName("Python Spark Feedforward neural network example") \
.config("spark.some.config.option", "some-value") \
.getOrCreate()
from pyspark.ml.classification import MultilayerPerceptronClassifier
from pyspark.ml.evaluation import MulticlassClassificationEvaluator
# Load training data
df = spark.read.format('com.databricks.spark.csv').\
options(header='true', \
inferschema='true').load("./data/WineData.csv",header=True);
df.show(5)
df.printSchema()
# Convert to float format
def string_to_float(x):
return float(x)
#
def condition(r):
if (0<= r <= 4):
label = "low"
elif(4< r <= 6):
label = "medium"
else:
label = "high"
return label
from pyspark.sql.functions import udf
from pyspark.sql.types import StringType, DoubleType
string_to_float_udf = udf(string_to_float, DoubleType())
quality_udf = udf(lambda x: condition(x), StringType())
#df= df.withColumn("quality", string_to_float_udf("quality")).withColumn("Cquality", quality_udf("quality"))
df= df.withColumn("quality", quality_udf("quality"))
df.printSchema()
df.show()
# convert the data to dense vector
def transData(data):
return data.rdd.map(lambda r: [r[-1], Vectors.dense(r[:-1])]).toDF(['label','features'])
from pyspark.sql import Row
from pyspark.ml.linalg import Vectors
data= transData(df)
data.show()
from pyspark.ml.feature import IndexToString, StringIndexer, VectorIndexer
# Index labels, adding metadata to the label column.
# Fit on whole dataset to include all labels in index.
labelIndexer = StringIndexer(inputCol="label", outputCol="indexedLabel").fit(data)
labelIndexer.transform(data).show(6)
# Automatically identify categorical features, and index them.
# Set maxCategories so features with > 4 distinct values are treated as continuous.
featureIndexer =VectorIndexer(inputCol="features", \
outputCol="indexedFeatures", \
maxCategories=4).fit(data)
featureIndexer.transform(data).show(6)
data.printSchema()
# Split the data into train and test
(trainingData, testData) = data.randomSplit([0.6, 0.4])
data.show()
# specify layers for the neural network:
# input layer of size 11 (features), two intermediate of size 5 and 4
# and output of size 7 (classes)
layers = [11, 5, 4, 4, 3 , 7]
# create the trainer and set its parameters
FNN = MultilayerPerceptronClassifier(labelCol="indexedLabel", featuresCol="indexedFeatures",\
maxIter=100, layers=layers, blockSize=128, seed=1234)
# Convert indexed labels back to original labels.
labelConverter = IndexToString(inputCol="prediction", outputCol="predictedLabel",
labels=labelIndexer.labels)
# Chain indexers and forest in a Pipeline
from pyspark.ml import Pipeline
pipeline = Pipeline(stages=[labelIndexer, featureIndexer, FNN, labelConverter])
# train the model
# Train model. This also runs the indexers.
model = pipeline.fit(trainingData)
# Make predictions.
predictions = model.transform(testData)
# Select example rows to display.
predictions.select("features","label","predictedLabel").show(5)
# Select (prediction, true label) and compute test error
evaluator = MulticlassClassificationEvaluator(
labelCol="indexedLabel", predictionCol="prediction", metricName="accuracy")
accuracy = evaluator.evaluate(predictions)
print("Predictions accuracy = %g, Test Error = %g" % (accuracy,(1.0 - accuracy)))
```
| github_jupyter |
```
import scipy.io, os
import numpy as np
import matplotlib.pyplot as plt
from netCDF4 import Dataset
from fastjmd95 import rho
from matplotlib.colors import ListedColormap
import seaborn as sns; sns.set()
sns.set()
from mpl_toolkits.axes_grid1.inset_locator import inset_axes
import matplotlib as mpl
colours=sns.color_palette('colorblind', 10)
my_cmap = ListedColormap(colours)
color_list=colours
```
## Code to plot the meridional overturning and density structure from the North Atlantic
Data used are from the ECCOv4 State Estimate available: https://ecco-group.org/products-ECCO-V4r4.html
The plot below features the North Atlantic basin, but the data includes the Southern Ocean and the arctic, with only the Indian and Pacific basins removed.
The plot can be adjusted latitudinally, and data to plot the Indian and Pacific ocean are available.
Initially we load the required data:
```
gridInfo=np.load('latLonDepthLevelECCOv4.npz')
zLev=gridInfo['depthLevel'][:]
depthPlot=zLev.cumsum()
lat=gridInfo['lat'][:]
lon=gridInfo['lon'][:]
dens=np.load('density20yr.npy')
masks=np.load('regimeMasks.npz')
maskMD=masks['maskMD']
maskSSV=masks['maskSSV']
maskNSV=masks['maskNSV']
maskTR=masks['maskTR']
maskSO=masks['maskSO']
maskNL=masks['maskNL']
PSI_A=np.load('PSI_atlantic.npz')
PSI_MD_A=PSI_A['PSI_MD_A']
PSI_TR_A=PSI_A['PSI_TR_A']
PSI_SSV_A=PSI_A['PSI_SSV_A']
PSI_NSV_A=PSI_A['PSI_NSV_A']
PSI_SO_A=PSI_A['PSI_SO_A']
PSI_NL_A=PSI_A['PSI_NL_A']
PSI_all_A=PSI_A['PSI_all_A']
```
### Define the functions needed to plot the data
```
levs=[32,33,34, 34.5, 35, 35.5,36,36.5,37,37.25,37.5,37.75,38]
cols=plt.cm.viridis([300,250, 200,150, 125, 100, 50,30, 10,15,10,9,1])
Land=np.ones(np.nansum(PSI_all_A, axis=0).shape)*np.nan
Land[np.nansum(PSI_all_A, axis=0)==0.0]=0
land3D=np.ones(dens.shape)
land3D[dens==0]=np.nan
def zPlotSurf(ax, data,zMin, zMax,label,mm,latMin,latMax,RGB,Ticks,saveName='test'):
land=np.ones(np.nanmean(data, axis=0).shape)*np.nan
land[np.nansum(data, axis=0)==0.0]=0
n=50
levels = np.linspace(-20, 20, n+1)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],-np.nanmean(data, axis=0)[zMin:zMax,latMin:latMax], levels=np.linspace(-20, 20, n+1),cmap=plt.cm.seismic, extend='both')
n2=30
densityPlot=np.nanmean((dens*land3D*mm), axis=2)
assert(len(levs)==len(cols))
CS=ax.contour(lat[0,latMin:latMax],-depthPlot[zMin:zMax],densityPlot[zMin:zMax,latMin:latMax],
levels=levs,
linewidths=3,colors=cols, extend='both')
ax.tick_params(axis='y', labelsize=20)
if Ticks == 0:
ax.set_xticklabels( () )
elif Ticks == 1:
ax.set_xticklabels( () )
ax.set_yticklabels( () )
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],land[zMin:zMax,latMin:latMax], 1,cmap=plt.cm.Set2)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],Land[zMin:zMax,latMin:latMax], 50,cmap=plt.cm.bone)
yL=ax.get_ylim()
xL=ax.get_xlim()
plt.text(xL[0]+0.02*np.ptp(xL), yL[0]+0.4*np.ptp(yL), label, fontsize=20, size=30,
weight='bold', bbox={'facecolor':'white', 'alpha':0.7}, va='bottom')
def zPlotDepth(ax, data,zMin, zMax,label,mm,latMin,latMax,RGB,Ticks,saveName='test'):
land=np.ones(np.nanmean(data, axis=0).shape)*np.nan
land[np.nansum(data, axis=0)==0.0]=0
n=50
levels = np.linspace(-20, 20, n+1)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],-np.nanmean(data, axis=0)[zMin:zMax,latMin:latMax], levels=np.linspace(-20, 20, n+1),cmap=plt.cm.seismic, extend='both')
n2=30
densityPlot=np.nanmean((dens*land3D*mm), axis=2)
ax.contour(lat[0,latMin:latMax],-depthPlot[zMin:zMax],densityPlot[zMin:zMax,latMin:latMax], colors=cols,
levels=levs,
linewidths=3, extend='both')
if Ticks == 0:
ax.tick_params(axis='y', labelsize=20)
#ax.set_xticklabels( () )
elif Ticks== 1:
#ax.set_xticklabels( () )
ax.set_yticklabels( () )
plt.tick_params(axis='both', labelsize=20)
#plt.clim(cmin, cmax)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],land[zMin:zMax,latMin:latMax], 1,cmap=plt.cm.Set2)
ax.contourf(lat[0,latMin:latMax],-depthPlot[zMin:zMax],Land[zMin:zMax,latMin:latMax], 50,cmap=plt.cm.bone)
yL=ax.get_ylim()
xL=ax.get_xlim()
plt.text(xL[0]+0.03*np.ptp(xL), yL[0]+0.03*np.ptp(yL), label, fontsize=20, size=30,
weight='bold', bbox={'facecolor':RGB, 'alpha':1}, va='bottom')
```
The figure is a composite of different subplots, calling the different functions to plot the surface and the deep.
```
# Set general figure options
# figure layout
xs = 15.5 # figure width in inches
nx = 2 # number of axes in x dimension
ny = 3 # number of sub-figures in y dimension (each sub-figure has two axes)
nya = 2 # number of axes per sub-figure
idy = [2.0, 1.0] # size of the figures in the y dimension
xm = [0.07, 0.07,0.9, 0.07] # x margins of the figure (left to right)
ym = [1.5] + ny*[0.07, 0.1] + [0.3] # y margins of the figure (bottom to top)
# pre-calculate some things
xcm = np.cumsum(xm) # cumulative margins
ycm = np.cumsum(ym) # cumulative margins
idx = (xs - np.sum(xm))/nx
idy_off = [0] + idy
ys = np.sum(idy)*ny + np.sum(ym) # size of figure in y dimension
# make the figure!
fig = plt.figure(figsize=(xs, ys))
# loop through sub-figures
ix,iy=0,0
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_TR_A,1,50,'TR', maskTR,200, 310, color_list[1],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
else:
xticks = ax.get_xticks()
ax.set_xticklabels(['{:0.0f}$^\circ$N'.format(xtick) for xtick in xticks])
elif iys == 1:
zPlotSurf(ax, PSI_TR_A,0,10,'', maskTR,200, 310, color_list[1],'')
# remove x ticks
ax.set_xticks([])
ix,iy=0,1
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_NL_A,1,50,'NL', maskNL,200, 310, color_list[-1],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_NL_A,0,10,'', maskNL,200, 310, color_list[4],'')
# remove x ticks
ax.set_xticks([])
############### n-SV
ix,iy=0,2
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_NSV_A,1,50,'N-SV', maskNSV,200, 310, color_list[4],'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_NSV_A,0,10,'', maskNSV,200, 310, color_list[-1],'')
# remove x ticks
ax.set_xticks([])
#
#_______________________________________________________________________
# S-SV
ix,iy=1,2
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
# ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_SSV_A,1,50,'S-SV', maskSSV,200, 310, color_list[2],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_SSV_A,0,10,'', maskSSV,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
#%%%%%%%%%%%%%%%%%%%%%%%%% SO
ix,iy=1,1
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_SO_A,1,50,'SO', maskSO,200, 310, color_list[-3],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
elif iys == 1:
zPlotSurf(ax, PSI_SO_A,0,10,'', maskSO,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
#%%%%%%%MD
ix,iy=1,0
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
#ax = plt.axes(loc)
for iys in range(nya):
# (bottom left corner x, bottom left corner y, width, height)
loc = ((xcm[ix] + (ix*idx))/xs,
(ycm[nya*iy + iys] + np.sum(idy)*iy+ idy_off[iys])/ys,
idx/xs,
idy[iys]/ys)
#print(loc[0], loc[1], loc[0] + loc[2], loc[1] + loc[3])
# create the axis
ax = plt.axes(loc)
# split between your two figure types
if iys == 0:
zPlotDepth(ax, PSI_MD_A,1,50,'MD', maskMD,200, 310, color_list[0],1,'')
# if not the bottom figure remove x ticks
if iy > 0:
ax.set_xticks([])
else:
xticks = ax.get_xticks()
ax.set_xticklabels(['{:0.0f}$^\circ$N'.format(xtick) for xtick in xticks])
elif iys == 1:
zPlotSurf(ax, PSI_MD_A,0,10,'', maskMD,200, 310, color_list[-3],1,'')
# remove x ticks
ax.set_xticks([])
cmap = plt.get_cmap('viridis')
cmap = mpl.colors.ListedColormap(cols)
ncol = len(levs)
axes = plt.axes([(xcm[0])/(xs), (ym[0]-0.6)/ys, (2*idx + xm[1])/(xs*2), (0.2)/ys])
cb = fig.colorbar(plt.cm.ScalarMappable(norm=mpl.colors.Normalize(-0.5, ncol - 0.5), cmap=cmap),
cax=axes, orientation='horizontal')
cb.ax.set_xticks(np.arange(ncol))
cb.ax.set_xticklabels(['{:0.2f}'.format(lev) for lev in levs])
cb.ax.tick_params(labelsize=20)
cb.set_label(label=r'Density, $\sigma_2$',weight='bold', fontsize=20)
cmap = plt.get_cmap('seismic')
ncol = len(cols)
axes = plt.axes([(xcm[2]+2*idx)/(xs*2), (ym[0]-0.6)/ys, (2*idx+xm[3])/(xs*2), (0.2)/ys])
cb = fig.colorbar(plt.cm.ScalarMappable(norm=mpl.colors.Normalize(-20,20), cmap=cmap),
cax=axes, label='title', orientation='horizontal', extend='both',format='%.0f',
boundaries=np.linspace(-20, 20, 41))
cb.ax.tick_params(labelsize=20)
cb.set_label(label=r'SV ($10^{6}m^{2}s^{-2}$)',weight='bold', fontsize=20)
# save as a png
#fig.savefig('psiRho_NAtl_sigma2.png', dpi=200, bbox_inches='tight')
```
| github_jupyter |
# Everything is better with friends: Using SAS in Python applications with SASPy and open-source tools
## Half-Day Tutorial • SAS Global Forum 2020
## Section 1. Python Code Conventions and Data Structures
### Example 1.1. Meet the Python Environment
<b><u>Instructions</u></b>: Click anywhere in the code cell immediately below, and run the cell using Shift-Enter. Then attempt the Exercises that follow, only looking at the explanatory notes for hints when needed.
```
import warnings
warnings.filterwarnings('ignore')
import platform
print(platform.sys.version)
help('modules')
```
**Line-by-Line Code Explanation**:
* Lines 1-2: Load the `warnings` module, and use the `filterwarnings` method to suppress warnings globally. (This is needed because of warnings generated when Line 7 is executed in SAS University Edition.)
* Lines 4-5: Load the `platform` module, and print Python and OS version information.
* Line 7: Print all modules currently available to be loaded into the Python kernel.
**Exercise 1.1.1**. True or False: Changing Line 1 to `IMPORT WARNINGS` would result in an execution error.
**Exercise 1.1.2**. True or False: The example code should result in an execution error because there are no terminating semicolons.
**Notes about Example 1.1**:
1. To increase performance, only a small number of modules in Python's standard library are available to use directly by default, so the `warnings` and `platform` modules need to be explicitly loaded before use. Python has a large standard library because of its "batteries included" philosophy.
2. Numerous third-party modules are also actively developed and made freely available through sites like https://github.com/ and https://pypi.org/. Two of the third-party modules needed for this tutorial are pandas, which we'll use for DataFrame objects below, and saspy, which allows Python scripts to connect to a SAS kernel for executing SAS code. Both of these modules come pre-installed with SAS University Edition.
3. This example illustrates four ways Python syntax differs from SAS:
* Unlike SAS, capitalization matters in Python. Changing Line 4 to `IMPORT PLATFORM` would produce an error.
* Unlike SAS, semicolons are optional in Python, and they are typically only used to separate multiple statements placed on the same line. E.g., Lines 4-5 could be combined into `import platform; print(platform.sys.version)`
* Unlike SAS, dot-notation has a consistent meaning in Python and can be used to reference objects nested inside each other at any depth. E.g., the `platform` module object invokes the sub-module object `sys` nested inside of it, and `sys` invokes the object `version` nested inside of it. (Think Russian nesting dolls or turduckens.)
* Unlike SAS, single and double quotes always have identical behavior in Python. E.g., `help('modules')` would produce exactly the same results as `help("modules")`.
4. If an error is displayed, an incompatible kernel has been chosen. This Notebook was developed using the Python 3.5 kernel provided with SAS University Edition as of January 2020.
### Example 1.2. Hello, Data!
<b><u>Instructions</u></b>: Click anywhere in the code cell immediately below, and run the cell using Shift-Enter. Then attempt the Exercises that follow, only looking at the explanatory notes for hints when needed.
```
hello_world_str = 'Hello, Jupyter!'
print(hello_world_str)
print()
if hello_world_str == 'Hello, Jupyter!':
print(type(hello_world_str))
else:
print("Error: The string doesn't have the expected value!")
```
**Line-by-Line Code Explanation**:
* Lines 1-3: Create a string object (`str` for short) named `hello_world_str`, and print it's value, followed by a blank line.
* Lines 4-7: Check to see if `hello_world_str` has the expected value. If so, print it's type. Otherwise, print an error message.
**Exercise 1.2.1**. Which of the following changes to the above example would result in an error? (pick all that apply):
* [ ] Removing an equal sign (`=`) in the expression `if hello_world_str == 'Hello, Jupyter!'`
* [ ] Removing the statement: `print()`
* [ ] Unindenting `print(type(hello_world_str))`
**Exercise 1.2.2**. Write several lines of Python code to produce the following output:
```
42
<class 'int'>
```
```
```
**Notes about Example 1.2**:
1. This example illustrates three more ways Python differs from SAS:
* Unlike SAS, variables are dynamically typed in Python. After Line 1 has been used to create `hello_world_str`, it can be assigned a new value later with a completely different type. E.g., we could later use `hello_world_str = 42` to change `type(hello_world_str)` to `<class 'int'>`.
* Unlike SAS, single-equals (`=`) only ever means assignment, and double-equals (`==`) only ever tests for equality, in Python. E.g., changing Line 4 to `if hello_world_str = 'Hello, Jupyter!'` would produce an error.
* Unlike SAS, indentation is significant and used to determine scope in Python. E.g., unindenting Line 5 would produce an error since the `if` statement would no longer have a body.
### Example 1.3. Python Lists and Indexes
<b><u>Instructions</u></b>: Click anywhere in the code cell immediately below, and run the cell using Shift-Enter. Then attempt the Exercises that follow, only looking at the explanatory notes for hints when needed.
```
hello_world_list = ['Hello', 'list']
print(hello_world_list)
print()
print(type(hello_world_list))
```
**Line-by-Line Code Explanation**:
* Line 1: Create a list object named `hello_world_list`, which contains two strings.
* Lines 2-4: Print the contents of `hello_world_list`, followed by a blank line and its type.
**Exercise 1.3.1**. Would the Python statement `print(hello_world_list[1])` display the value `'Hello'` or `'World'`?
**Exercise 1.3.2**. True or False: A Python list may only contain values of the same type.
**Notes about Example 1.3**.
1. Values in lists are always kept in insertion order, meaning the order they appear in the list's definition, and they can be individually accessed using numerical indexes within bracket notation:
* `hello_world_list[0]` returns `'Hello'`
* `hello_world_list[1]` returns `'list'`.
2. The left-most element of a list is always at index `0`. Unlike SAS, customized indexing is only available for more sophisticated data structures in Python (e.g., a dictionary, as in Example 1.4 below).
3. Lists are the most fundamental Python data structure and are related to SAS data-step arrays. However, unlike a SAS data-step array, a Python list object may contain values of different types (such as `str` or `int`). However, processing the values of a list without checking their types may cause errors if it contains unexpected values.
### Example 1.4. Python Dictionaries
<b><u>Instructions</u></b>: Click anywhere in the code cell immediately below, and run the cell using Shift-Enter. Then attempt the Exercises that follow, only looking at the explanatory notes for hints when needed.
```
hello_world_dict = {
'salutation' : ['Hello' , 'dict'],
'valediction' : ['Goodbye' , 'list'],
'part of speech' : ['interjection', 'noun'],
}
print(hello_world_dict)
print()
print(type(hello_world_dict))
```
**Line-by-Line Code Explanation**:
* Line 1-5 : Create a dictionary object (`dict` for short) named `hello_world_dict`, which contains three key-value pairs, where each key is a string and each value is a list of two strings.
* Lines 6-8: Print the contents of `hello_world_dict`, followed by a blank line and its type.
**Exercise 1.4.1**. What would be displayed by executing the statement `print(hello_world_dict['salutation'])`?
**Exercise 1.4.2**. Write a single line of Python code to print the initial element of the list associated with the key `valediction`.
```
```
**Notes about Example 1.4**:
1. Dictionaries are another fundamental Python data structure, which map keys (appearing before the colons in Lines 2-4) to values (appearing after the colons in Lines 2-4). The value associated with each key can be accessed using bracket notation:
* hello_world_dict['salutation'] returns ['Hello', 'dict']
* hello_world_dict['valediction'] returns ['Goodbye', 'list']
* hello_world_dict['part of speech'] returns ['interjection', 'noun']
2. Whenever indexable data structures are nested in Python, indexing methods can be combined. E.g., `hello_world_dict['salutation'][0] == ['Hello', 'dict'][0] == 'Hello'`.
3. Dictionaries are more generally called _associative arrays_ or _maps_ and are related to SAS formats and data-step hash tables.
4. In Python 3.5, the print order of key-value pairs may not match insertion order, meaning the order key-value pairs are listed when the dictionary is created. As of Python 3.7 (released in June 2018), insertion order is preserved.
### Example 1.5. Introduction to Data Frames
<b><u>Instructions</u></b>: Click anywhere in the code cell immediately below, and run the cell using Shift-Enter. Then attempt the Exercises that follow, only looking at the explanatory notes for hints when needed.
```
from pandas import DataFrame
hello_world_df = DataFrame(
{
'salutation' : ['Hello' , 'DataFrame'],
'valediction' : ['Goodbye' , 'dict'],
'part of speech' : ['exclamation', 'noun'],
}
)
print(hello_world_df)
print()
print(hello_world_df.shape)
print()
print(hello_world_df.info())
```
**Line-by-Line Code Explanation**:
* Line 1: Load the definition of a `DataFrame` object from the `pandas` module. (Think of a DataFrame as a rectangular array of values, with all values in a column having the same type.)
* Lines 2-8: Create a DataFrame object (`df` for short) named `hello_world_df` with dimensions 2x3 (2 rows by 3 columns), with each key-value pair in the dictionary in Lines 3-7 becoming a column that is labelled by the key.
* Lines 9-14: Print the contents of `hello_world_df`, following by a blank line, the number of rows and columns in it, another blank line, and some information about it.
**Exercise 1.5.1**. Write a single line of Python code to print the column labelled by `salutation`.
```
```
**Exercise 1.5.2**. Write a single line of Python code to print the final element of the column labeled by `valediction`.
```
```
**Notes About Example 1.5**:
1. The DataFrame object type is not built into Python, which is why we first have to import its definition from the pandas module.
1. DataFrames can be indexed like dictionaries composed of lists. E.g., `hello_world_df['salutation'][0] == ['Hello', 'dict'][0] == 'Hello'`
2. A DataFrame is a tabular data structure with rows and columns, similar to a SAS data set. However, while SAS datasets are typically accessed from disk and processed row-by-row, DataFrames are loaded into memory all at once. This means values in DataFrames can be randomly accessed, but it also means the size of DataFrames can't grow beyond available memory.
3. The dimensions of the DataFrame are determined as follows:
* The keys `'salutation'`, `'valediction'`, and `'part of speech'` of the dictionary passed to the `DataFrame` constructor function become column labels.
* Because each key maps to a list of length two, each column will be two elements tall (with an error occurring if the lists are not of non-uniform length).
4. The DataFrame constructor function can also accept many other object types, including another DataFrame.
| github_jupyter |
# Infected vs Smartphone (Korea)
What percentage of COVID-19 patients could an app cover?
Please direct your questions to [Bin Zhang](https://www.linkedin.com/in/binzhangmd)
```
# Mount for Colab (Optional)
COLAB = False
if COLAB:
from google.colab import drive
drive.mount('/content/drive')
DATASET_BASE = '/content/drive/My Drive/CovidApps/datasets/'
else:
DATASET_BASE = '../datasets/'
DS4C_PATH = DATASET_BASE + 'coronavirusdataset/'
AGE_PATH = DS4C_PATH + 'TimeAge.csv'
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
```
#### Read CSV
```
LAST_UPDATE = '2020-05-14'
age = pd.read_csv(AGE_PATH)
age.tail(10)
print(f'Date range: {len(age.date.unique())} days ({min(age.date)} ~ {max(age.date)})')
age_list = age.age.unique() # 80s := 80+
print('Age groups:', age_list)
```
#### Cases by Age
```
fig, ax = plt.subplots(figsize=(13, 7))
plt.title('Confirmed Cases by Age (as of {0})'.format(LAST_UPDATE), fontsize=15)
sns.barplot(age_list, age.confirmed[-9:])
ax.set_xlabel('age', size=13)
ax.set_ylabel('number of cases', size=13)
plt.show()
```
#### Smartphone Penetration by Age
```
# https://www.statista.com/statistics/897195/south-korea-smartphone-ownership-by-age-group/
data = {'age': [0, 10, 20, 30, 40, 50, 60, 70, 80],
'smartphone': [0.955, 0.955, 0.999, 0.999, 0.997, 0.984, 0.875, 0.363, 0.363]}
smartphone = pd.DataFrame(data)
fig, ax = plt.subplots(figsize=(13, 7))
plt.title('Smartphone Penetration KR, 2018-09', fontsize=15)
sns.barplot(age_list, smartphone['smartphone'])
ax.set_xlabel('age', size=13)
ax.set_ylabel('smartphone', size=13)
plt.show()
```
#### Smartphone Coverage
```
N = len(age_list)
groups = age.confirmed[-9:].to_numpy()
wSp = np.round(groups * smartphone['smartphone'])
woSp = np.round(groups * (1 - smartphone['smartphone']))
width = 0.5 # the width of the bars: can also be len(x) sequence
p1 = plt.bar(age_list, wSp, width)
p2 = plt.bar(age_list, woSp, width,
bottom=wSp)
plt.ylabel('Patients')
plt.title('Patients vs. Smartphone by Age')
plt.legend((p1[0], p2[0]), ('Smartphone', 'Cellphone / Nix'))
plt.show()
cover = np.sum(wSp) / (np.sum(wSp) + np.sum(woSp))
print('Smartphone coverage: ' + '{:.2%}'.format(cover) + ' of infected cases')
```
#### App Coverage (80% Penetation Assumption)
```
APP_COVERAGE = 0.8
yApp = np.round(wSp * APP_COVERAGE)
nApp = np.round(wSp * (1 - APP_COVERAGE))
width = 0.5 # the width of the bars: can also be len(x) sequence
p1 = plt.bar(age_list, yApp, width)
p2 = plt.bar(age_list, nApp, width,
bottom=yApp)
p3 = plt.bar(age_list, woSp, width,
bottom=yApp+nApp)
plt.ylabel('Patients')
plt.title('Patients - Smartphone - App by Age')
plt.legend((p1[0], p2[0], p3[0]), ('Smartphone (App)', 'Smartphone (No App)', 'Cellphone / Nix'))
plt.show()
cover = np.sum(yApp) / (np.sum(wSp) + np.sum(woSp))
print('App coverage: ' + '{:.2%}'.format(cover) + ' of infected cases')
```
| github_jupyter |
# Going deeper with Tensorflow
In this video, we're going to study the tools you'll use to build deep learning models. Namely, [Tensorflow](https://www.tensorflow.org/).
If you're running this notebook outside the course environment, you'll need to install tensorflow:
* `pip install tensorflow` should install cpu-only TF on Linux & Mac OS
* If you want GPU support from offset, see [TF install page](https://www.tensorflow.org/install/)
```
import sys
sys.path.append("..")
import grading
```
# Visualization
Plase note that if you are running on the Coursera platform, you won't be able to access the tensorboard instance due to the network setup there. If you run the notebook locally, you should be able to access TensorBoard on http://127.0.0.1:7007/
```
! killall tensorboard
import os
os.system("tensorboard --logdir=/tmp/tboard --port=7007 &");
import tensorflow as tf
s = tf.InteractiveSession()
```
# Warming up
For starters, let's implement a python function that computes the sum of squares of numbers from 0 to N-1.
```
import numpy as np
def sum_sin(N):
return np.sum(np.arange(N)**2)
%%time
sum_sin(10**8)
```
# Tensoflow teaser
Doing the very same thing
```
# An integer parameter
N = tf.placeholder('int64', name="input_to_your_function")
# A recipe on how to produce the same result
result = tf.reduce_sum(tf.range(N)**2)
result
%%time
result.eval({N: 10**8})
writer = tf.summary.FileWriter("/tmp/tboard", graph=s.graph)
```
# How does it work?
1. Define placeholders where you'll send inputs
2. Make symbolic graph: a recipe for mathematical transformation of those placeholders
3. Compute outputs of your graph with particular values for each placeholder
* `output.eval({placeholder:value})`
* `s.run(output, {placeholder:value})`
So far there are two main entities: "placeholder" and "transformation"
* Both can be numbers, vectors, matrices, tensors, etc.
* Both can be int32/64, floats, booleans (uint8) of various size.
* You can define new transformations as an arbitrary operation on placeholders and other transformations
* `tf.reduce_sum(tf.arange(N)**2)` are 3 sequential transformations of placeholder `N`
* There's a tensorflow symbolic version for every numpy function
* `a+b, a/b, a**b, ...` behave just like in numpy
* `np.mean` -> `tf.reduce_mean`
* `np.arange` -> `tf.range`
* `np.cumsum` -> `tf.cumsum`
* If if you can't find the op you need, see the [docs](https://www.tensorflow.org/api_docs/python).
`tf.contrib` has many high-level features, may be worth a look.
```
with tf.name_scope("Placeholders_examples"):
# Default placeholder that can be arbitrary float32
# scalar, vertor, matrix, etc.
arbitrary_input = tf.placeholder('float32')
# Input vector of arbitrary length
input_vector = tf.placeholder('float32', shape=(None,))
# Input vector that _must_ have 10 elements and integer type
fixed_vector = tf.placeholder('int32', shape=(10,))
# Matrix of arbitrary n_rows and 15 columns
# (e.g. a minibatch your data table)
input_matrix = tf.placeholder('float32', shape=(None, 15))
# You can generally use None whenever you don't need a specific shape
input1 = tf.placeholder('float64', shape=(None, 100, None))
input2 = tf.placeholder('int32', shape=(None, None, 3, 224, 224))
# elementwise multiplication
double_the_vector = input_vector*2
# elementwise cosine
elementwise_cosine = tf.cos(input_vector)
# difference between squared vector and vector itself plus one
vector_squares = input_vector**2 - input_vector + 1
my_vector = tf.placeholder('float32', shape=(None,), name="VECTOR_1")
my_vector2 = tf.placeholder('float32', shape=(None,))
my_transformation = my_vector * my_vector2 / (tf.sin(my_vector) + 1)
print(my_transformation)
dummy = np.arange(5).astype('float32')
print(dummy)
my_transformation.eval({my_vector:dummy, my_vector2:dummy[::-1]})
writer.add_graph(my_transformation.graph)
writer.flush()
```
TensorBoard allows writing scalars, images, audio, histogram. You can read more on tensorboard usage [here](https://www.tensorflow.org/get_started/graph_viz).
# Summary
* Tensorflow is based on computation graphs
* The graphs consist of placehlders and transformations
# Mean squared error
Your assignment is to implement mean squared error in tensorflow.
```
with tf.name_scope("MSE"):
y_true = tf.placeholder("float32", shape=(None,), name="y_true")
y_predicted = tf.placeholder("float32", shape=(None,), name="y_predicted")
# Your code goes here
# You want to use tf.reduce_mean
# mse = tf.<...>
mse = tf.reduce_mean(tf.squared_difference(y_true, y_predicted))
def compute_mse(vector1, vector2):
return mse.eval({y_true: vector1, y_predicted: vector2})
writer.add_graph(mse.graph)
writer.flush()
```
Tests and result submission. Please use the credentials obtained from the Coursera assignment page.
```
import submit
submit.submit_mse(compute_mse, "ssq6554@126.com", "adoEuwVXGdMn5aDs")
```
# Variables
The inputs and transformations have no value outside function call. This isn't too comfortable if you want your model to have parameters (e.g. network weights) that are always present, but can change their value over time.
Tensorflow solves this with `tf.Variable` objects.
* You can assign variable a value at any time in your graph
* Unlike placeholders, there's no need to explicitly pass values to variables when `s.run(...)`-ing
* You can use variables the same way you use transformations
```
# Creating a shared variable
shared_vector_1 = tf.Variable(initial_value=np.ones(5),
name="example_variable")
# Initialize variable(s) with initial values
s.run(tf.global_variables_initializer())
# Evaluating shared variable (outside symbolicd graph)
print("Initial value", s.run(shared_vector_1))
# Within symbolic graph you use them just
# as any other inout or transformation, not "get value" needed
# Setting a new value
s.run(shared_vector_1.assign(np.arange(5)))
# Getting that new value
print("New value", s.run(shared_vector_1))
```
# tf.gradients - why graphs matter
* Tensorflow can compute derivatives and gradients automatically using the computation graph
* True to its name it can manage matrix derivatives
* Gradients are computed as a product of elementary derivatives via the chain rule:
$$ {\partial f(g(x)) \over \partial x} = {\partial f(g(x)) \over \partial g(x)}\cdot {\partial g(x) \over \partial x} $$
It can get you the derivative of any graph as long as it knows how to differentiate elementary operations
```
my_scalar = tf.placeholder('float32')
scalar_squared = my_scalar**2
# A derivative of scalar_squared by my_scalar
derivative = tf.gradients(scalar_squared, [my_scalar, ])
derivative
import matplotlib.pyplot as plt
%matplotlib inline
x = np.linspace(-3, 3)
x_squared, x_squared_der = s.run([scalar_squared, derivative[0]],
{my_scalar:x})
plt.plot(x, x_squared,label="$x^2$")
plt.plot(x, x_squared_der, label=r"$\frac{dx^2}{dx}$")
plt.legend();
```
# Why that rocks
```
my_vector = tf.placeholder('float32', [None])
# Compute the gradient of the next weird function over my_scalar and my_vector
# Warning! Trying to understand the meaning of that function may result in permanent brain damage
weird_psychotic_function = tf.reduce_mean(
(my_vector+my_scalar)**(1+tf.nn.moments(my_vector,[0])[1]) +
1./ tf.atan(my_scalar))/(my_scalar**2 + 1) + 0.01*tf.sin(
2*my_scalar**1.5)*(tf.reduce_sum(my_vector)* my_scalar**2
)*tf.exp((my_scalar-4)**2)/(
1+tf.exp((my_scalar-4)**2))*(1.-(tf.exp(-(my_scalar-4)**2)
)/(1+tf.exp(-(my_scalar-4)**2)))**2
der_by_scalar = tf.gradients(weird_psychotic_function, my_scalar)
der_by_vector = tf.gradients(weird_psychotic_function, my_vector)
# Plotting the derivative
scalar_space = np.linspace(1, 7, 100)
y = [s.run(weird_psychotic_function, {my_scalar:x, my_vector:[1, 2, 3]})
for x in scalar_space]
plt.plot(scalar_space, y, label='function')
y_der_by_scalar = [s.run(der_by_scalar,
{my_scalar:x, my_vector:[1, 2, 3]})
for x in scalar_space]
plt.plot(scalar_space, y_der_by_scalar, label='derivative')
plt.grid()
plt.legend();
```
# Almost done - optimizers
While you can perform gradient descent by hand with automatic grads from above, tensorflow also has some optimization methods implemented for you. Recall momentum & rmsprop?
```
y_guess = tf.Variable(np.zeros(2, dtype='float32'))
y_true = tf.range(1, 3, dtype='float32')
loss = tf.reduce_mean((y_guess - y_true + tf.random_normal([2]))**2)
#loss = tf.reduce_mean((y_guess - y_true)**2)
optimizer = tf.train.MomentumOptimizer(0.01, 0.5).minimize(
loss, var_list=y_guess)
from matplotlib import animation, rc
import matplotlib_utils
from IPython.display import HTML
fig, ax = plt.subplots()
y_true_value = s.run(y_true)
level_x = np.arange(0, 2, 0.02)
level_y = np.arange(0, 3, 0.02)
X, Y = np.meshgrid(level_x, level_y)
Z = (X - y_true_value[0])**2 + (Y - y_true_value[1])**2
ax.set_xlim(-0.02, 2)
ax.set_ylim(-0.02, 3)
s.run(tf.global_variables_initializer())
ax.scatter(*s.run(y_true), c='red')
contour = ax.contour(X, Y, Z, 10)
ax.clabel(contour, inline=1, fontsize=10)
line, = ax.plot([], [], lw=2)
def init():
line.set_data([], [])
return (line,)
guesses = [s.run(y_guess)]
def animate(i):
s.run(optimizer)
guesses.append(s.run(y_guess))
line.set_data(*zip(*guesses))
return (line,)
anim = animation.FuncAnimation(fig, animate, init_func=init,
frames=400, interval=20, blit=True)
try:
HTML(anim.to_html5_video())
# In case the build-in renderers are unaviable, fall back to
# a custom one, that doesn't require external libraries
except RuntimeError:
anim.save(None, writer=matplotlib_utils.SimpleMovieWriter(0.001))
```
# Logistic regression
Your assignment is to implement the logistic regression
Plan:
* Use a shared variable for weights
* Use a matrix placeholder for `X`
We shall train on a two-class MNIST dataset
* please note that target `y` are `{0,1}` and not `{-1,1}` as in some formulae
```
from sklearn.datasets import load_digits
mnist = load_digits(2)
X, y = mnist.data, mnist.target
print("y [shape - %s]:" % (str(y.shape)), y[:10])
print("X [shape - %s]:" % (str(X.shape)))
print('X:\n',X[:3,:10])
print('y:\n',y[:10])
plt.imshow(X[0].reshape([8,8]));
```
It's your turn now!
Just a small reminder of the relevant math:
$$
P(y=1|X) = \sigma(X \cdot W + b)
$$
$$
\text{loss} = -\log\left(P\left(y_\text{predicted} = 1\right)\right)\cdot y_\text{true} - \log\left(1 - P\left(y_\text{predicted} = 1\right)\right)\cdot\left(1 - y_\text{true}\right)
$$
$\sigma(x)$ is available via `tf.nn.sigmoid` and matrix multiplication via `tf.matmul`
```
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=42)
```
__Your code goes here.__ For the training and testing scaffolding to work, please stick to the names in comments.
```
# Model parameters - weights and bias
# weights = tf.Variable(...) shape should be (X.shape[1], 1)
# b = tf.Variable(...)
weights = tf.Variable(initial_value=np.random.randn(X.shape[1], 1)*0.01, name="weights", dtype="float32")
b = tf.Variable(initial_value=0, name="b", dtype="float32")
print(weights)
print(b)
# Placeholders for the input data
# input_X = tf.placeholder(...)
# input_y = tf.placeholder(...)
input_X = tf.placeholder(tf.float32, name="input_X")
input_y = tf.placeholder(tf.float32, name="input_y")
print(input_X)
print(input_y)
# The model code
# Compute a vector of predictions, resulting shape should be [input_X.shape[0],]
# This is 1D, if you have extra dimensions, you can get rid of them with tf.squeeze .
# Don't forget the sigmoid.
# predicted_y = <predicted probabilities for input_X>
predicted_y = tf.squeeze(tf.nn.sigmoid(tf.add(tf.matmul(input_X, weights), b)))
print(predicted_y)
# Loss. Should be a scalar number - average loss over all the objects
# tf.reduce_mean is your friend here
# loss = <logistic loss (scalar, mean over sample)>
loss = -tf.reduce_mean(tf.log(predicted_y)*input_y + tf.log(1-predicted_y)*(1-input_y))
print(loss)
# See above for an example. tf.train.*Optimizer
# optimizer = <optimizer that minimizes loss>
optimizer = tf.train.GradientDescentOptimizer(0.01).minimize(loss)
print(optimizer)
```
A test to help with the debugging
```
validation_weights = 1e-3 * np.fromiter(map(lambda x:
s.run(weird_psychotic_function, {my_scalar:x, my_vector:[1, 0.1, 2]}),
0.15 * np.arange(1, X.shape[1] + 1)),
count=X.shape[1], dtype=np.float32)[:, np.newaxis]
# Compute predictions for given weights and bias
prediction_validation = s.run(
predicted_y, {
input_X: X,
weights: validation_weights,
b: 1e-1})
# Load the reference values for the predictions
validation_true_values = np.loadtxt("validation_predictons.txt")
assert prediction_validation.shape == (X.shape[0],),\
"Predictions must be a 1D array with length equal to the number " \
"of examples in input_X"
assert np.allclose(validation_true_values, prediction_validation)
loss_validation = s.run(
loss, {
input_X: X[:100],
input_y: y[-100:],
weights: validation_weights+1.21e-3,
b: -1e-1})
assert np.allclose(loss_validation, 0.728689)
from sklearn.metrics import roc_auc_score
s.run(tf.global_variables_initializer())
for i in range(5):
s.run(optimizer, {input_X: X_train, input_y: y_train})
loss_i = s.run(loss, {input_X: X_train, input_y: y_train})
print("loss at iter %i:%.4f" % (i, loss_i))
print("train auc:", roc_auc_score(y_train, s.run(predicted_y, {input_X:X_train})))
print("test auc:", roc_auc_score(y_test, s.run(predicted_y, {input_X:X_test})))
```
### Coursera submission
```
grade_submitter = grading.Grader("BJCiiY8sEeeCnhKCj4fcOA")
test_weights = 1e-3 * np.fromiter(map(lambda x:
s.run(weird_psychotic_function, {my_scalar:x, my_vector:[1, 2, 3]}),
0.1 * np.arange(1, X.shape[1] + 1)),
count=X.shape[1], dtype=np.float32)[:, np.newaxis]
```
First, test prediction and loss computation. This part doesn't require a fitted model.
```
prediction_test = s.run(
predicted_y, {
input_X: X,
weights: test_weights,
b: 1e-1})
assert prediction_test.shape == (X.shape[0],),\
"Predictions must be a 1D array with length equal to the number " \
"of examples in X_test"
grade_submitter.set_answer("0ENlN", prediction_test)
loss_test = s.run(
loss, {
input_X: X[:100],
input_y: y[-100:],
weights: test_weights+1.21e-3,
b: -1e-1})
# Yes, the X/y indices mistmach is intentional
grade_submitter.set_answer("mMVpM", loss_test)
grade_submitter.set_answer("D16Rc", roc_auc_score(y_test, s.run(predicted_y, {input_X:X_test})))
```
Please use the credentials obtained from the Coursera assignment page.
```
grade_submitter.submit("ssq6554@126.com", "zfkj43piwD65Symi")
```
| github_jupyter |
# Intepretability on Hateful Twitter Datasets
In this demo, we apply saliency maps (with support of sparse tensors) on the task on the detection of Twitter users who use hateful lexicon using graph machine learning with Stellargraph.
We consider the use-case of identifying hateful users on Twitter motivated by the work in [1] and using the dataset also published in [1]. Classification is based on a graph based on users' retweets and attributes as related to their account activity, and the content of tweets.
We pose identifying hateful users as a binary classification problem. We demonstrate the advantage of connected vs unconnected data in a semi-supervised setting with few training examples.
For connected data, we use Graph Convolutional Networks [2] as implemented in the `stellargraph` library. We pose the problem of identifying hateful tweeter users as node attribute inference in graphs.
We then use the interpretability tool (i.e., saliency maps) implemented in our library to demonstrate how to obtain the importance of the node features and links to gain insights into the model.
**References**
1. "Like Sheep Among Wolves": Characterizing Hateful Users on Twitter. M. H. Ribeiro, P. H. Calais, Y. A. Santos, V. A. F. Almeida, and W. Meira Jr. arXiv preprint arXiv:1801.00317 (2017).
2. Semi-Supervised Classification with Graph Convolutional Networks. T. Kipf, M. Welling. ICLR 2017. arXiv:1609.02907
```
import networkx as nx
import pandas as pd
import numpy as np
import seaborn as sns
import itertools
import os
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.linear_model import LogisticRegressionCV
import stellargraph as sg
from stellargraph.mapper import GraphSAGENodeGenerator, FullBatchNodeGenerator
from stellargraph.layer import GraphSAGE, GCN, GAT
from stellargraph import globalvar
from tensorflow.keras import layers, optimizers, losses, metrics, Model, models
from sklearn import preprocessing, feature_extraction
from sklearn.model_selection import train_test_split
from sklearn import metrics
import matplotlib.pyplot as plt
import seaborn as sns
from scipy.sparse import csr_matrix, lil_matrix
%matplotlib inline
import matplotlib.pyplot as plt
%matplotlib inline
def remove_prefix(text, prefix):
return text[text.startswith(prefix) and len(prefix):]
def plot_history(history):
metrics = sorted(set([remove_prefix(m, "val_") for m in list(history.history.keys())]))
for m in metrics:
# summarize history for metric m
plt.plot(history.history[m])
plt.plot(history.history['val_' + m])
plt.title(m, fontsize=18)
plt.ylabel(m, fontsize=18)
plt.xlabel('epoch', fontsize=18)
plt.legend(['train', 'validation'], loc='best')
plt.show()
```
### Train GCN model on the dataset
```
data_dir = os.path.expanduser("~/data/hateful-twitter-users")
```
### First load and prepare the node features
Each node in the graph is associated with a large number of features (also referred to as attributes).
The list of features is given [here](https://www.kaggle.com/manoelribeiro/hateful-users-on-twitter). We repeated here for convenience.
hate :("hateful"|"normal"|"other")
if user was annotated as hateful, normal, or not annotated.
(is_50|is_50_2) :bool
whether user was deleted up to 12/12/17 or 14/01/18.
(is_63|is_63_2) :bool
whether user was suspended up to 12/12/17 or 14/01/18.
(hate|normal)_neigh :bool
is the user on the neighborhood of a (hateful|normal) user?
[c_] (statuses|follower|followees|favorites)_count :int
number of (tweets|follower|followees|favorites) a user has.
[c_] listed_count:int
number of lists a user is in.
[c_] (betweenness|eigenvector|in_degree|outdegree) :float
centrality measurements for each user in the retweet graph.
[c_] *_empath :float
occurrences of empath categories in the users latest 200 tweets.
[c_] *_glove :float
glove vector calculated for users latest 200 tweets.
[c_] (sentiment|subjectivity) :float
average sentiment and subjectivity of users tweets.
[c_] (time_diff|time_diff_median) :float
average and median time difference between tweets.
[c_] (tweet|retweet|quote) number :float
percentage of direct tweets, retweets and quotes of an user.
[c_] (number urls|number hashtags|baddies|mentions) :float
number of bad words|mentions|urls|hashtags per tweet in average.
[c_] status length :float
average status length.
hashtags :string
all hashtags employed by the user separated by spaces.
**Notice** that c_ are attributes calculated for the 1-neighborhood of a user in the retweet network (averaged out).
First, we are going to load the user features and prepare them for machine learning.
```
users_feat = pd.read_csv(os.path.join(data_dir,
'users_neighborhood_anon.csv'))
```
### Data cleaning and preprocessing
The dataset as given includes a large number of graph related features that are manually extracted.
Since we are going to employ modern graph neural networks methods for classification, we are going to drop these manually engineered features.
The power of Graph Neural Networks stems from their ability to learn useful graph-related features eliminating the need for manual feature engineering.
```
def data_cleaning(feat):
feat = feat.drop(columns=["hate_neigh", "normal_neigh"])
# Convert target values in hate column from strings to integers (0,1,2)
feat['hate'] = np.where(feat['hate']=='hateful', 1, np.where(feat['hate']=='normal', 0, 2))
# missing information
number_of_missing = feat.isnull().sum()
number_of_missing[number_of_missing!=0]
# Replace NA with 0
feat.fillna(0, inplace=True)
# droping info about suspension and deletion as it is should not be use din the predictive model
feat.drop(feat.columns[feat.columns.str.contains("is_|_glove|c_|sentiment")], axis=1, inplace=True)
# drop hashtag feature
feat.drop(['hashtags'], axis=1, inplace=True)
# Drop centrality based measures
feat.drop(columns=['betweenness', 'eigenvector', 'in_degree', 'out_degree'], inplace=True)
feat.drop(columns=['created_at'], inplace=True)
return feat
node_data = data_cleaning(users_feat)
```
The continous features in our dataset have distributions with very long tails. We apply normalization to correct for this.
```
# Ignore the first two columns because those are user_id and hate (the target variable)
df_values = node_data.iloc[:, 2:].values
pt = preprocessing.PowerTransformer(method='yeo-johnson',
standardize=True)
df_values_log = pt.fit_transform(df_values)
node_data.iloc[:, 2:] = df_values_log
# Set the dataframe index to be the same as the user_id and drop the user_id columns
node_data.index = node_data.index.map(str)
node_data.drop(columns=['user_id'], inplace=True)
```
### Next load the graph
Now that we have the node features prepared for machine learning, let us load the retweet graph.
```
g_nx = nx.read_edgelist(path=os.path.expanduser(os.path.join(data_dir,
"users.edges")))
g_nx.number_of_nodes(), g_nx.number_of_edges()
```
The graph has just over 100k nodes and approximately 2.2m edges.
We aim to train a graph neural network model that will predict the "hate"attribute on the nodes.
For computation convenience, we have mapped the target labels **normal**, **hateful**, and **other** to the numeric values **0**, **1**, and **2** respectively.
```
print(set(node_data["hate"]))
list(g_nx.nodes())[:10]
node_data = node_data.loc[list(g_nx.nodes())]
node_data.head()
node_data.index
```
### Splitting the data
For machine learning we want to take a subset of the nodes for training, and use the rest for validation and testing. We'll use scikit-learn again to split our data into training and test sets.
The total number of annotated nodes is very small when compared to the total number of nodes in the graph. We are only going to use 15% of the annotated nodes for training and the remaining 85% of nodes for testing.
First, we are going to select the subset of nodes that are annotated as hateful or normal. These will be the nodes that have 'hate' values that are either 0 or 1.
```
# choose the nodes annotated with normal or hateful classes
annotated_users = node_data[node_data['hate']!=2]
annotated_user_features = annotated_users.drop(columns=['hate'])
annotated_user_targets = annotated_users[['hate']]
```
There are 4971 annoted nodes out of a possible, approximately, 100k nodes.
```
print(annotated_user_targets.hate.value_counts())
# split the data
train_data, test_data, train_targets, test_targets = train_test_split(annotated_user_features,
annotated_user_targets,
test_size=0.85,
random_state=101)
train_targets = train_targets.values
test_targets = test_targets.values
print("Sizes and class distributions for train/test data")
print("Shape train_data {}".format(train_data.shape))
print("Shape test_data {}".format(test_data.shape))
print("Train data number of 0s {} and 1s {}".format(np.sum(train_targets==0),
np.sum(train_targets==1)))
print("Test data number of 0s {} and 1s {}".format(np.sum(test_targets==0),
np.sum(test_targets==1)))
train_targets.shape, test_targets.shape
train_data.shape, test_data.shape
```
We are going to use 745 nodes for training and 4226 nodes for testing.
```
# choosing features to assign to a graph, excluding target variable
node_features = node_data.drop(columns=['hate'])
```
### Dealing with imbalanced data
Because the training data exhibit high imbalance, we introduce class weights.
```
from sklearn.utils.class_weight import compute_class_weight
class_weights = compute_class_weight('balanced',
np.unique(train_targets),
train_targets[:,0])
train_class_weights = dict(zip(np.unique(train_targets),
class_weights))
train_class_weights
```
Our data is now ready for machine learning.
Node features are stored in the Pandas DataFrame `node_features`.
The graph in networkx format is stored in the variable `g_nx`.
### Specify global parameters
Here we specify some parameters that control the type of model we are going to use. For example, we specify the base model type, e.g., GCN, GraphSAGE, etc, as well as model-specific parameters.
```
epochs = 20
```
## Creating the base graph machine learning model in Keras
Now create a `StellarGraph` object from the `NetworkX` graph and the node features and targets. It is `StellarGraph` objects that we use in this library to perform machine learning tasks on.
```
G = sg.StellarGraph(g_nx, node_features=node_features)
print(list(G.nodes())[:10])
```
To feed data from the graph to the Keras model we need a generator. The generators are specialized to the model and the learning task.
For training we map only the training nodes returned from our splitter and the target values.
```
generator = FullBatchNodeGenerator(G, method="gcn", sparse=True)
train_gen = generator.flow(train_data.index,
train_targets, )
base_model = GCN(
layer_sizes=[32, 16],
generator = generator,
bias=True,
dropout=0.5,
activations=["elu", "elu"]
)
x_inp, x_out = base_model.node_model()
prediction = layers.Dense(units=1, activation="sigmoid")(x_out)
```
### Create a Keras model
Now let's create the actual Keras model with the graph inputs `x_inp` provided by the `base_model` and outputs being the predictions from the softmax layer.
```
model = Model(inputs=x_inp, outputs=prediction)
```
We compile our Keras model to use the `Adam` optimiser and the binary cross entropy loss.
```
model.compile(
optimizer=optimizers.Adam(lr=0.005),
loss=losses.binary_crossentropy,
metrics=["acc"],
)
model
```
Train the model, keeping track of its loss and accuracy on the training set, and its performance on the test set during the training. We don't use the test set during training but only for measuring the trained model's generalization performance.
```
test_gen = generator.flow(test_data.index, test_targets)
history = model.fit_generator(
train_gen,
epochs=epochs,
validation_data=test_gen,
verbose=0,
shuffle=False,
class_weight=None,
)
```
### Model Evaluation
Now we have trained the model, let's evaluate it on the test set.
We are going to consider 4 evaluation metrics calculated on the test set: Accuracy, Area Under the ROC curve (AU-ROC), the ROC curve, and the confusion table.
#### Accuracy
```
test_metrics = model.evaluate_generator(test_gen)
print("\nTest Set Metrics:")
for name, val in zip(model.metrics_names, test_metrics):
print("\t{}: {:0.4f}".format(name, val))
all_nodes = node_data.index
all_gen = generator.flow(all_nodes)
all_predictions = model.predict_generator(all_gen).squeeze()[..., np.newaxis]
all_predictions.shape
all_predictions_df = pd.DataFrame(all_predictions,
index=node_data.index)
```
Let's extract the predictions for the test data only.
```
test_preds = all_predictions_df.loc[test_data.index, :]
test_preds.shape
```
The predictions are the probability of the true class that in this case is the probability of a user being hateful.
```
test_predictions = test_preds.values
test_predictions_class = ((test_predictions>0.5)*1).flatten()
test_df = pd.DataFrame({"Predicted_score": test_predictions.flatten(),
"Predicted_class": test_predictions_class,
"True": test_targets[:,0]})
roc_auc = metrics.roc_auc_score(test_df['True'].values,
test_df['Predicted_score'].values)
print("The AUC on test set:\n")
print(roc_auc)
```
## Interpretability by Saliency Maps
To understand which features and edges the model is looking at while making the predictions, we use the interpretability tool in the StellarGraph library (i.e., saliency maps) to demonstrate the importance of node features and edges given a target user.
```
from stellargraph.utils.saliency_maps import IntegratedGradients
int_saliency = IntegratedGradients(model, all_gen)
print(test_data.index)
from stellargraph.utils.saliency_maps import IntegratedGradients, GradientSaliency
#we first select a list of nodes which are confidently classified as hateful.
predicted_hateful_index = set(np.where(all_predictions > 0.9)[0].tolist())
test_indices_set = set([int(k) for k in test_data.index.tolist()])
hateful_in_test = list(predicted_hateful_index.intersection(test_indices_set))
print(hateful_in_test)
#let's pick one node from the predicted hateful users as an example.
idx = 2
target_idx = hateful_in_test[idx]
target_nid = list(G.nodes())[target_idx]
print('target_idx = {}, target_nid = {}'.format(target_idx, target_nid))
print('prediction score for node {} is {}'.format(target_idx, all_predictions[target_idx]))
print('ground truth score for node {} is {}'.format(target_idx, test_targets[test_data.index.tolist().index(str(target_nid))]))
[X,all_targets,A_index, A], y_true_all = all_gen[0]
```
For the prediction of the target node, we then calculate the importance of the features for each node in the graph. Our support for sparse saliency maps makes it efficient to fit the scale like this dataset.
```
#We set the target_idx which is our target node.
node_feature_importance = int_saliency.get_integrated_node_masks(target_idx, 0)
```
As `node_feature_importance` is a matrix where `node_feature_importance[i][j]` indicates the importance of the j-th feature of node i to the prediction of the target node, we sum up the feature importance of each node to measure its node importance.
```
node_importance = np.sum(node_feature_importance, axis=-1)
node_importance_rank = np.argsort(node_importance)[::-1]
print(node_importance[node_importance_rank])
print('node_importance has {} non-zero values'.format(np.where(node_importance != 0)[0].shape[0]))
```
We expect the number of non-zero values of `node_importance` to match the number of nodes in the ego graph.
```
G_ego = nx.ego_graph(g_nx,target_nid, radius=2)
print('The ego graph of the target node has {} neighbors'.format(len(G_ego.nodes())))
```
We then analyze the feature importance of the top-250 important nodes. See the output for the top-5 importance nodes. For each row, the features are sorted according to their importance.
```
feature_names = annotated_users.keys()[1:].values
feature_importance_rank = np.argsort(node_feature_importance[target_idx])[::-1]
df = pd.DataFrame([([k] + list(feature_names[np.argsort(node_feature_importance[k])[::-1]])) for k in node_importance_rank[:250]], columns = range(205))
df.head()
```
As a sanity check, we expect the target node itself to have a relatively high importance.
```
self_feature_importance_rank = np.argsort(node_feature_importance[target_idx])
print(np.sum(node_feature_importance[target_idx]))
print('The node itself is the {}-th important node'.format(1 + node_importance_rank.tolist().index(target_idx)))
df = pd.DataFrame([feature_names[self_feature_importance_rank][::-1]], columns = range(204))
df
```
For different nodes, the same features may have different ranks. To understand the overall importance of the features, we now analyze the average feature importance rank for the above selected nodes. Specifically, we obtain the average rank of each specific feature among the top-250 important nodes.
```
from collections import defaultdict
average_feature_rank = defaultdict(int)
for i in node_importance_rank[:250]:
feature_rank = list(feature_names[np.argsort(node_feature_importance[i])[::-1]])
for j in range(len(feature_rank)):
average_feature_rank[feature_rank[j]] += feature_rank.index(feature_rank[j])
for k in average_feature_rank.keys():
average_feature_rank[k] /= 250.0
sorted_avg_feature_rank = sorted(average_feature_rank.items(), key=lambda a:a[1])
for feat, avg_rank in sorted_avg_feature_rank:
print(feat, avg_rank)
```
It seems for our target node, topics relevant to cleaning, hipster, etc. are important while those such as leaisure, ship, goverment, etc. are not important.
We then calculate the link importance for the edges that are connected to the target node within k hops (k = 2 for our GCN model).
```
link_importance = int_saliency.get_integrated_link_masks(target_idx, 0, steps=2)
(x, y) = link_importance.nonzero()
[X,all_targets,A_index, A], y_true_all = all_gen[0]
print(A_index.shape, A.shape)
G_edge_indices = [(A_index[0, k, 0], A_index[0, k, 1]) for k in range(A_index.shape[1])]
link_dict = {(A_index[0, k, 0], A_index[0, k, 1]):k for k in range(A_index.shape[1])}
```
As a sanity check, we expect the most important edge to connect important nodes.
```
nonzero_importance_val = link_importance[(x,y)].flatten().tolist()[0]
link_importance_rank = np.argsort(nonzero_importance_val)[::-1]
edge_number_in_ego_graph = link_importance_rank.shape[0]
print('There are {} edges within the ego graph of the target node'.format(edge_number_in_ego_graph))
x_rank, y_rank = x[link_importance_rank], y[link_importance_rank]
print('The most important edge connects {}-th important node and {}-th important node'.format(node_importance_rank.tolist().index(x_rank[0]), (node_importance_rank.tolist().index(y_rank[0]))))
```
To ensure that we are getting the correct importance for edges, we then check what happens if we perturb the top-10 most important edges. Specifically, if we remove the top important edges according to the calculated edge importance scores, we should expect to see the prediction of the target node change.
```
from copy import deepcopy
print(A_index.shape)
selected_nodes = np.array([[target_idx]], dtype='int32')
prediction_clean = model.predict([X, selected_nodes, A_index, A]).squeeze()
A_perturb = deepcopy(A)
print('A_perturb.shape = {}'.format(A_perturb.shape))
#we remove top 1% important edges in the graph and see how the prediction changes
topk = int(edge_number_in_ego_graph * 0.01)
for i in range(topk):
edge_x, edge_y = x_rank[i], y_rank[i]
edge_index = link_dict[(edge_x, edge_y)]
A_perturb[0, edge_index] = 0
```
As expected, the prediction score drops after the perturbation. The target node is predicted as non-hateful now.
```
prediction = model.predict([X, selected_nodes, A_index, A_perturb]).squeeze()
print('The prediction score changes from {} to {} after the perturbation'.format(prediction_clean, prediction))
```
NOTES: For UX team, the above notebook shows how we are able to compute the importance of nodes and edges. However, it seems the ego graph of the target node in twitter dataset is often very big so that we may draw only top important nodes/edges on the visualization.
| github_jupyter |
```
import time
import matplotlib
#matplotlib.use('TKAgg')
import SwarmStartleLooming as sw
import numpy as np
import matplotlib.pyplot as plt
np.random.seed(200)
# Initialize Parameters
N = 10
L = 50
total_time = 50.0
dt = 0.001
speed0 = 1.5
alpha = 1
noisep = 0.01
noisev = 0.0
BC = 0 #'along_wall'
IC = 10 # initial condition
repstrength = 1.0
algstrength = 0.5
attstrength = 0.3
reprange = 1.0
algrange = 5.0
attrange = 25.0
repsteep = -20
output = 0.05
int_type = 'voronoi'
startle = True
amplitude_startle = 50.0
duration_startle = 0.070
# initialize system parameters
paraSystem = sw.BaseParams(N=N, L=L, time=total_time, dt=dt, BC=BC, IC=IC, output=output, int_type=int_type, startle=startle)
# initialize prey parameters
paraFish = sw.AgentParams(paraSystem, speed0=speed0, alpha=alpha,
repstrength=repstrength, reprange=reprange, repsteepness=repsteep,
algstrength=algstrength, algrange=algrange,
attstrength=attstrength, attrange=attrange,
noisep=noisep, noisev=noisev,
amplitude_startle=amplitude_startle,
duration_startle=duration_startle,
print_startles=True,
r_m=10*1e6,
tau_m=0.023,
e_l=-0.079,
v_t=-0.061,
vt_std=0.000,
tau_rho=0.001,
rho_null=3.6,
rho_null_std=0.7,
rho_scale=8.16 * 1e6,
exc_scale=30,
noise_std_exc=0.0027,
noise_std_inh=0.000,
vis_input_m=3,
vis_input_b=0,
vis_input_method='max',
vis_input_k=3
)
starttime = time.time()
outData, agentData = sw.SingleRun(paraSystem, paraFish)
endtime = time.time()
print('Total time needed: ' + str(int((endtime - starttime))) + ' seconds or '
+ str(int((endtime - starttime) / 60)) + ' min '
+ 'or ' + str(int((endtime - starttime) / 3600)) + ' hours')
np.random.seed(200)
# Initialize Parameters
N = 40
L = 50
total_time = 50.0
dt = 0.001
speed0 = 1.5
alpha = 1
noisep = 0.01
noisev = 0.0
BC = 0 #'along_wall'
IC = 10 # initial condition
repstrength = 1.0
algstrength = 0.5
attstrength = 0.3
reprange = 1.0
algrange = 5.0
attrange = 25.0
repsteep = -20
output = 0.05
int_type = 'voronoi_matrix'
startle = True
amplitude_startle = 50.0
duration_startle = 0.070
# initialize system parameters
paraSystem = sw.BaseParams(N=N, L=L, time=total_time, dt=dt, BC=BC, IC=IC, output=output, int_type=int_type, startle=startle)
# initialize prey parameters
paraFish = sw.AgentParams(paraSystem, speed0=speed0, alpha=alpha,
repstrength=repstrength, reprange=reprange, repsteepness=repsteep,
algstrength=algstrength, algrange=algrange,
attstrength=attstrength, attrange=attrange,
noisep=noisep, noisev=noisev,
amplitude_startle=amplitude_startle,
duration_startle=duration_startle,
print_startles=True,
r_m=10*1e6,
tau_m=0.023,
e_l=-0.079,
v_t=-0.061,
vt_std=0.000,
tau_rho=0.001,
rho_null=3.6,
rho_null_std=0.7,
rho_scale=8.16 * 1e6,
exc_scale=30,
noise_std_exc=0.0027,
noise_std_inh=0.000,
vis_input_m=3,
vis_input_b=0,
vis_input_method='max',
vis_input_k=3
)
starttime = time.time()
outData, agentData = sw.SingleRun(paraSystem, paraFish)
endtime = time.time()
print('Total time needed: ' + str(int((endtime - starttime))) + ' seconds or '
+ str(int((endtime - starttime) / 60)) + ' min '
+ 'or ' + str(int((endtime - starttime) / 3600)) + ' hours')
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.