text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
# Classes and subclasses
In this notebook, I will show you the basics of classes and subclasses in Python. As you've seen in the lectures from this week, `Trax` uses layer classes as building blocks for deep learning models, so it is important to understand how classes and subclasses behave in order to be able to build custom layers when needed.
By completing this notebook, you will:
- Be able to define classes and subclasses in Python
- Understand how inheritance works in subclasses
- Be able to work with instances
# Part 1: Parameters, methods and instances
First, let's define a class `My_Class`.
```
class My_Class: #Definition of My_class
x = None
```
`My_Class` has one parameter `x` without any value. You can think of parameters as the variables that every object assigned to a class will have. So, at this point, any object of class `My_Class` would have a variable `x` equal to `None`. To check this, I'll create two instances of that class and get the value of `x` for both of them.
```
instance_a= My_Class() #To create an instance from class "My_Class" you have to call "My_Class"
instance_b= My_Class()
print('Parameter x of instance_a: ' + str(instance_a.x)) #To get a parameter 'x' from an instance 'a', write 'a.x'
print('Parameter x of instance_b: ' + str(instance_b.x))
```
For an existing instance you can assign new values for any of its parameters. In the next cell, assign a value of `5` to the parameter `x` of `instance_a`.
```
### START CODE HERE (1 line) ###
instance_a.x = 5
### END CODE HERE ###
print('Parameter x of instance_a: ' + str(instance_a.x))
```
## 1.1 The `__init__` method
When you want to assign values to the parameters of your class when an instance is created, it is necessary to define a special method: `__init__`. The `__init__` method is called when you create an instance of a class. It can have multiple arguments to initialize the paramenters of your instance. In the next cell I will define `My_Class` with an `__init__` method that takes the instance (`self`) and an argument `y` as inputs.
```
class My_Class:
def __init__(self, y): # The __init__ method takes as input the instance to be initialized and a variable y
self.x = y # Sets parameter x to be equal to y
```
In this case, the parameter `x` of an instance from `My_Class` would take the value of an argument `y`.
The argument `self` is used to pass information from the instance being created to the method `__init__`. In the next cell, create an instance `instance_c`, with `x` equal to `10`.
```
### START CODE HERE (1 line) ###
instance_c = My_Class(10)
### END CODE HERE ###
print('Parameter x of instance_c: ' + str(instance_c.x))
```
Note that in this case, you had to pass the argument `y` from the `__init__` method to create an instance of `My_Class`.
## 1.2 The `__call__` method
Another important method is the `__call__` method. It is performed whenever you call an initialized instance of a class. It can have multiple arguments and you can define it to do whatever you want like
- Change a parameter,
- Print a message,
- Create new variables, etc.
In the next cell, I'll define `My_Class` with the same `__init__` method as before and with a `__call__` method that adds `z` to parameter `x` and prints the result.
```
class My_Class:
def __init__(self, y): # The __init__ method takes as input the instance to be initialized and a variable y
self.x = y # Sets parameter x to be equal to y
def __call__(self, z): # __call__ method with self and z as arguments
self.x += z # Adds z to parameter x when called
print(self.x)
```
Let’s create `instance_d` with `x` equal to 5.
```
instance_d = My_Class(5)
```
And now, see what happens when `instance_d` is called with argument `10`.
```
instance_d(10)
```
Now, you are ready to complete the following cell so any instance from `My_Class`:
- Is initialized taking two arguments `y` and `z` and assigns them to `x_1` and `x_2`, respectively. And,
- When called, takes the values of the parameters `x_1` and `x_2`, sums them, prints and returns the result.
```
class My_Class:
def __init__(self, y, z): #Initialization of x_1 and x_2 with arguments y and z
### START CODE HERE (2 lines) ###
self.x_1 = y
self.x_2 = z
### END CODE HERE ###
def __call__(self): #When called, adds the values of parameters x_1 and x_2, prints and returns the result
### START CODE HERE (1 line) ###
result = self.x_1 + self.x_2
### END CODE HERE ###
print("Addition of {} and {} is {}".format(self.x_1,self.x_2,result))
return result
```
Run the next cell to check your implementation. If everything is correct, you shouldn't get any errors.
```
instance_e = My_Class(10,15)
def test_class_definition():
assert instance_e.x_1 == 10, "Check the value assigned to x_1"
assert instance_e.x_2 == 15, "Check the value assigned to x_2"
assert instance_e() == 25, "Check the __call__ method"
print("\033[92mAll tests passed!")
test_class_definition()
```
## 1.3 Custom methods
In addition to the `__init__` and `__call__` methods, your classes can have custom-built methods to do whatever you want when called. To define a custom method, you have to indicate its input arguments, the instructions that you want it to perform and the values to return (if any). In the next cell, `My_Class` is defined with `my_method` that multiplies the values of `x_1` and `x_2`, sums that product with an input `w`, and returns the result.
```
class My_Class:
def __init__(self, y, z): #Initialization of x_1 and x_2 with arguments y and z
self.x_1 = y
self.x_2 = z
def __call__(self): #Performs an operation with x_1 and x_2, and returns the result
a = self.x_1 - 2*self.x_2
return a
def my_method(self, w): #Multiplies x_1 and x_2, adds argument w and returns the result
result = self.x_1*self.x_2 + w
return result
```
Create an instance `instance_f` of `My_Class` with any integer values that you want for `x_1` and `x_2`. For that instance, see the result of calling `My_method`, with an argument `w` equal to `16`.
```
### START CODE HERE (1 line) ###
instance_f = My_Class(1,10)
### END CODE HERE ###
print("Output of my_method:",instance_f.my_method(16))
```
As you can corroborate in the previous cell, to call a custom method `m`, with arguments `args`, for an instance `i` you must write `i.m(args)`. With that in mind, methods can call others within a class. In the following cell, try to define `new_method` which calls `my_method` with `v` as input argument. Try to do this on your own in the cell given below.
```
class My_Class:
def __init__(self, y, z): #Initialization of x_1 and x_2 with arguments y and z
self.x_1 = None
self.x_2 = None
def __call__(self): #Performs an operation with x_1 and x_2, and returns the result
a = None
return a
def my_method(self, w): #Multiplies x_1 and x_2, adds argument w and returns the result
b = None
return b
def new_method(self, v): #Calls My_method with argument v
### START CODE HERE (1 line) ###
result = self.my_method(v)
### END CODE HERE ###
return result
```
<b>SPOILER ALERT</b> Solution:
```
# hidden-cell
class My_Class:
def __init__(self, y, z): #Initialization of x_1 and x_2 with arguments y and z
self.x_1 = y
self.x_2 = z
def __call__(self): #Performs an operation with x_1 and x_2, and returns the result
a = self.x_1 - 2*self.x_2
return a
def my_method(self, w): #Multiplies x_1 and x_2, adds argument w and returns the result
b = self.x_1*self.x_2 + w
return b
def new_method(self, v): #Calls My_method with argument v
result = self.my_method(v)
return result
instance_g = My_Class(1,10)
print("Output of my_method:",instance_g.my_method(16))
print("Output of new_method:",instance_g.new_method(16))
```
# Part 2: Subclasses and Inheritance
`Trax` uses classes and subclasses to define layers. The base class in `Trax` is `layer`, which means that every layer from a deep learning model is defined as a subclass of the `layer` class. In this part of the notebook, you are going to see how subclasses work. To define a subclass `sub` from class `super`, you have to write `class sub(super):` and define any method and parameter that you want for your subclass. In the next cell, I define `sub_c` as a subclass of `My_Class` with only one method (`additional_method`).
```
class sub_c(My_Class): #Subclass sub_c from My_class
def additional_method(self): #Prints the value of parameter x_1
print(self.x_1)
```
## 2.1 Inheritance
When you define a subclass `sub`, every method and parameter is inherited from `super` class, including the `__init__` and `__call__` methods. This means that any instance from `sub` can use the methods defined in `super`. Run the following cell and see for yourself.
```
instance_sub_a = sub_c(1,10)
print('Parameter x_1 of instance_sub_a: ' + str(instance_sub_a.x_1))
print('Parameter x_2 of instance_sub_a: ' + str(instance_sub_a.x_2))
print("Output of my_method of instance_sub_a:",instance_sub_a.my_method(16))
```
As you can see, `sub_c` does not have an initialization method `__init__`, it is inherited from `My_class`. However, you can overwrite any method you want by defining it again in the subclass. For instance, in the next cell define a class `sub_c` with a redefined `my_Method` that multiplies `x_1` and `x_2` but does not add any additional argument.
```
class sub_c(My_Class): #Subclass sub_c from My_class
def my_method(self): #Multiplies x_1 and x_2 and returns the result
### START CODE HERE (1 line) ###
b = self.x_1*self.x_2
### END CODE HERE ###
return b
```
To check your implementation run the following cell.
```
test = sub_c(3,10)
assert test.my_method() == 30, "The method my_method should return the product between x_1 and x_2"
print("Output of overridden my_method of test:",test.my_method()) #notice we didn't pass any parameter to call my_method
#print("Output of overridden my_method of test:",test.my_method(16)) #try to see what happens if you call it with 1 argument
```
In the next cell, two instances are created, one of `My_Class` and another one of `sub_c`. The instances are initialized with equal `x_1` and `x_2` parameters.
```
y,z= 1,10
instance_sub_a = sub_c(y,z)
instance_a = My_Class(y,z)
print('My_method for an instance of sub_c returns: ' + str(instance_sub_a.my_method()))
print('My_method for an instance of My_Class returns: ' + str(instance_a.my_method(10)))
```
As you can see, even though `sub_c` is a subclass from `My_Class` and both instances are initialized with the same values, `My_method` returns different results for each instance because you overwrote `My_method` for `sub_c`.
<b>Congratulations!</b> You just reviewed the basics behind classes and subclasses. Now you can define your own classes and subclasses, work with instances and overwrite inherited methods. The concepts within this notebook are more than enough to understand how layers in `Trax` work.
| github_jupyter |
```
!pip install --upgrade git+https://github.com/EmGarr/kerod.git
%tensorflow_version 2.x
import tensorflow as tf
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
```
# Download COCO/2017
Download and preprocess COCO/2017 to the following format (required by od networks):
```python
dataset = {
'images' : A tensor of float32 and shape [1, height, widht, 3],
'images_info': A tensor of float32 and shape [1, 2] ,
'bbox': A tensor of float32 and shape [1, num_boxes, 4],
'labels': A tensor of int32 and shape [1, num_boxes],
'num_boxes': A tensor of int32 and shape [1, 1],
'weights': A tensor of float32 and shape [1, num_boxes]
}
```
If you need to download the dataset in a specific directory you can use the argument `data_dir` of `tfds.load`.
```
import functools
import tensorflow as tf
import tensorflow_datasets as tfds
from kerod.dataset.preprocessing import preprocess, expand_dims_for_single_batch
from kerod.core.standard_fields import BoxField
ds_train, ds_info = tfds.load(name="coco/2017", split="train", shuffle_files=True, with_info=True)
ds_train = ds_train.map(functools.partial(preprocess, bgr=True), num_parallel_calls=tf.data.experimental.AUTOTUNE)
# Filter example with no boxes after preprocessing
ds_train = ds_train.filter(lambda x, y: tf.shape(y[BoxField.BOXES])[0] > 1)
ds_train = ds_train.map(expand_dims_for_single_batch, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_train = ds_train.prefetch(tf.data.experimental.AUTOTUNE)
ds_test = tfds.load(name="coco/2017", split="validation", shuffle_files=False)
ds_test = ds_test.map(
functools.partial(preprocess, horizontal_flip=False, bgr=True),
num_parallel_calls=tf.data.experimental.AUTOTUNE)
# Filter example with no boxes after preprocessing
ds_test = ds_test.filter(lambda x, y: tf.shape(y[BoxField.BOXES])[0] > 1)
ds_test = ds_test.map(expand_dims_for_single_batch, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_test = ds_test.prefetch(tf.data.experimental.AUTOTUNE)
ds_info
```
# Load and train the network
```
from kerod.core.standard_fields import BoxField
from kerod.core.learning_rate_schedule import LearningRateScheduler
from kerod.model import factory
from tensorflow.keras.callbacks import TensorBoard, ModelCheckpoint
# Number of classes of COCO
classes = ds_info.features['objects']['label'].names
num_classes = len(classes)
model_faster_rcnn = factory.build_model(num_classes)
base_lr = 0.02
optimizer = tf.keras.optimizers.SGD(learning_rate=base_lr, momentum=0.9)
model_faster_rcnn.compile(optimizer=optimizer, loss=None)
#The numbering of epochs (LearningRateScheduler) starts at 0.
# Which means the decrease will happens on the epoch 9:
#(8 + 1: numbering of fit logging starts at 1)
callbacks = [
LearningRateScheduler(base_lr, 1, epochs=[8, 10]),
TensorBoard(),
ModelCheckpoint('checkpoints/')
]
model_faster_rcnn.fit(ds_train, validation_data=ds_test, epochs=12, callbacks=callbacks)
# Save the weights for the serving
model_faster_rcnn.save_weights('final_weights.h5')
# Export a saved model for serving purposes
model_faster_rcnn.export_for_serving('serving')
```
## Visualisation of few images
```
from od.utils.drawing import BoxDrawer
drawer = BoxDrawer(classes)
for i, example in enumerate(ds_val):
inputs, ground_truths = example
out = model_faster_rcnn.predict_on_batch(inputs)
boxes, scores, labels, valid_detections = out
# Will draw the results
drawer(
inputs['images'],
boxes,
scores=scores,
labels=labels,
num_valid_detections=valid_detections
)
if i == 5:
break
```
## Tensorboard
```
# Load TENSORBOARD
%load_ext tensorboard
# Start TENSORBOARD
%tensorboard --logdir logs
```
## Coco evaluation
### Load the dataset
```
import tensorflow_datasets as tfds
ds_val, ds_info = tfds.load(name="coco/2017", split="validation", shuffle_files=False, with_info=True)
# category_ids basicaly map the index 0 the id
# e.g: 0 -> 1, 2 -> 3, 79 -> 90
category_ids = [1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 13, 14, 15, 16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 27, 28, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43, 44, 46, 47, 48, 49, 50, 51, 52, 53, 54, 55, 56, 57, 58, 59, 60, 61, 62, 63, 64, 65, 67, 70, 72, 73, 74, 75, 76, 77, 78, 79, 80, 81, 82, 84, 85, 86, 87, 88, 89, 90]
```
### Super dirty but the evaluation works
1. perform the analysis
```
import json
import numpy as np
import tensorflow as tf
from kerod.core.standard_fields import DatasetField, BoxField
from kerod.core.box_ops import convert_to_center_coordinates
from kerod.dataset.preprocessing import resize_to_min_dim
results = []
for i, example in enumerate(ds_val):
print(i)
# preprocess image
image = example['image'][:, :, ::-1]
image = resize_to_min_dim(image, 800.0, 1333.0)
image_information = tf.cast(tf.shape(image)[:2], dtype=tf.float32)
inputs = {
'images': tf.expand_dims(image, axis=0),
'images_information':tf.expand_dims(image_information, axis=0)
}
# predict
boxes, scores, labels, valid_detections = model_faster_rcnn.predict_on_batch(inputs)
# Post processing and append to coco results
bbox = boxes[0] * tf.tile(
tf.expand_dims(tf.cast(example['image'].shape[:2], tf.float32), axis=0),
[1, 2])
scores = scores[0]
labels = labels[0]
for i in range(valid_detections[0]):
# Convert from [y_min, x_min, y_max, x_max] to coco format [x_min, y_min, w, h]
sbox = bbox[i].numpy()
sbox = [sbox[1], sbox[0], sbox[3] - sbox[1], sbox[2] - sbox[0]]
res = {
'image_id': int(example['image/id']),
'category_id': category_ids[int(labels[i])],
'bbox': [round(float(c), 4) for c in sbox],
'score': round(float(scores[i]), 4),
}
results.append(res)
with open('coco_results.json', 'w') as f:
json.dump(results, f)
```
2. install the coco library to compute the performances
```
!wget http://images.cocodataset.org/annotations/annotations_trainval2017.zip
!unzip annotations_trainval2017.zip
!pip install 'git+https://github.com/cocodataset/cocoapi.git#subdirectory=PythonAPI'
```
3. compute the performances
```
from pycocotools.coco import COCO
from pycocotools.cocoeval import COCOeval
with open('coco_results_corrected.json', 'r') as f:
results = json.load(f)
coco = COCO('./annotations/instances_val2017.json')
ret = {}
cocoDt = coco.loadRes(results)
cocoEval = COCOeval(coco, cocoDt, 'bbox')
cocoEval.evaluate()
cocoEval.accumulate()
cocoEval.summarize()
```
| github_jupyter |
# Vanishing
This jupyter notebook computes some vanishing results for the coherent cohomology of the Siegel variety in positive characteristic. Let $ p \geq 2$ be a prime number and $N \geq 3$ an integer such that $p \nmid N$. Let $X$ be a smooth and projective toroidal compactification of the Siegel variety of dimension $\frac{g(g+1)}{2}$ over $\mathbb{F}_p$. We denote by $D$ the border of the toroidal compactification. If $\lambda$ is a character of $\text{GL}_g$, then we denote by $\nabla(\lambda)$ the costandard automorphic vector bundle of highest weight $\lambda$. We implement an algorithm which computes characters $\lambda$ of $\text{GL}_g$ such that the coherent cohomology $H^i(X,\nabla(\lambda)(-D))$ vanishes.
```
from tqdm import tqdm
import matplotlib.pyplot as plt
class SiegelVariety:
def __init__(self,g,p):
self.g = g
self.p = p
self.d = g*(g+1)/2
if g < 2:
raise ValueError('The genus g must be greater than 1')
elif p not in Primes():
raise ValueError('The number ' + str(p) + ' is not prime')
elif p < self.g**2:
raise ValueError('The prime ' + str(p) + ' is lower than the dimension of the flag bundle ' + str(self.g**2))
self.L = RootSystem("A"+str(self.g-1)).ambient_space() #Ag-1
self.LG = RootSystem("C"+str(self.g)).ambient_space() #Cg
self.W = WeylGroup(self.L)
self.WG = WeylGroup(self.LG)
A = WeylCharacterRing(['A',self.g-1])
C = WeylCharacterRing(['C',self.g])
self.phi_L = A.positive_roots()
self.phi_G = C.positive_roots()
self.Delta_L = A.simple_roots()
self.Delta_G = C.simple_roots()
#Cotangent bundle of the Siegel variety
OmegaWeight = []
for i in range(self.g):
if i == 0:
OmegaWeight.append(2)
else:
OmegaWeight.append(0)
Omega1 = A(OmegaWeight)
#Exterior product of differentials over the Siegel variety
self.nu = []
for i in range(1,self.d+1):
temp = []
Omega = Omega1.exterior_power(i)
for x in Omega.weight_multiplicities():
temp.append(x)
self.nu.append(temp)
self.mu = []
for nu_j in self.nu:
temp = []
for char in nu_j:
temp.append(self.L(self.changeConvention(char)))
self.mu.append(temp)
#Known vanishing results - in degrees i>k for k = 0 to d-1
self.Cvan = []
for i in range(self.d):
self.Cvan.append([])
#Change root convention
def changeConvention(self,char):
res = []
for i in range(self.g-1,-1,-1):
res.append(-char[i])
return res
#Test equality of characters
def equality(self,el1,el2):
for i in range(self.g):
if el1[i] != el2[i]:
return False
return True
#Create a list of subsets of given cardinal
def powerset_length(self,s,n):
res = []
x = len(s)
for i in range(1 << x):
temp = []
temp = [s[j] for j in range(x) if (i & (1 << j))]
if len(temp) == n:
res.append(temp)
return res
#Test if a list contains an element
def contains(self,lst,el):
for x in lst:
if self.equality(x,el):
return True
return False
#Test our vanishing result
def vanishes(self,k, char):
return self.contains(self.Cvan[k],char)
#Construct relevant set of roots
def buildIprime(self,T):
Iprime = []
for i in range(1,len(T)+1):
if T[i-1] != 0:
Iprime.append(self.Delta_L[i])
return Iprime
def buildRoots(self,Iprime):
phi_Iprime = []
phiLIprime = []
for i in range(1 << len(Iprime)):
temp = sum(Iprime[j] for j in range(len(Iprime)) if (i & (1 << j)))
if temp != 0:
tempList = []
for k in range(self.g):
tempList.append(temp[k])
if self.LG(tempList) in self.phi_G:
phi_Iprime.append(temp)
for x in self.phi_L:
if x not in phi_Iprime:
phiLIprime.append(x)
return phi_Iprime,phiLIprime
#Check L_Iprime-dominance of characters
def LIprimedominant(self,Iprime,phiLIprime,char):
for alpha in phiLIprime:
value = 0
for j in range(self.g):
value += char[j]*alpha[j]
if value < 0:
return False
for alpha in Iprime:
value = 0
for j in range(self.g):
value += char[j]*alpha[j]
if value != 0:
return False
return True
#Check L-dominance
def Ldominant(self,char):
for alpha in self.Delta_L:
value = 0
for j in range(self.g):
value += char[j]*alpha[j]
if value < 0:
return False
return True
#Construction of a set of dominant weights between kmin and kmax
def buildWeights(self,kmin,kmax):
kvalues = []
kvalue = []
for i in range(self.g):
kvalue.append(kmin)
stop = False
stopGlobal = False
icurrent = 0
while not stopGlobal:
stop = False
kvaluetemp = kvalue.copy()
kvalues.append(kvaluetemp)
while not stop:
if kvalue[icurrent] < kmax:
kvalue[icurrent] += 1
icurrent = 0
stop = True
elif icurrent < self.g-1:
found = False
nextValue = 0
i = icurrent+1
while not found and i < self.g:
if kvalue[i] < kmax:
nextValue = kvalue[i]+1
found = True
i+=1
if found:
kvalue[icurrent] = nextValue
icurrent += 1
else:
stop = True
stopGlobal = True
else:
stop = True
stopGlobal = True
return kvalues
#If it returns True, it means that the automorphic line bundle of weight char
#is D-ample on the flag bundle of type Iprime
def ample(self,Iprime,char):
for w in self.WG:
for alpha in self.phi_G:
alpha = alpha.associated_coroot()
v = w.action(alpha)
num = 0
for j in range(self.g):
num += char[j]*v[j]
den = 0
for j in range(self.g):
den += char[j]*alpha[j]
if den != 0:
if max(num/den,-num/den) > self.p-1:
return False
for alpha in self.phi_G:
if not self.contains(self.phi_L,alpha):
alpha = alpha.associated_coroot()
value = 0
for j in range(self.g):
value += char[j]*alpha[j]
if value >= 0:
return False
for alpha in self.Delta_L:
if alpha not in Iprime:
value = 0
for j in range(self.g):
value += char[j]*alpha[j]
if value <= 0:
return False
return True
#Check a partial degeneration of a spectral sequence
def degeneration(self,Iprime,phiLIprime,tworhoIprime,e,char):
lamb = self.L(char)
lambp = lamb-self.mu[self.d-e-1][0]
r = len(phiLIprime)
if self.LIprimedominant(Iprime,phiLIprime,lambp) and self.ample(Iprime,lambp+tworhoIprime):
for k in range(e+1):
set_of_M = self.powerset_length(phiLIprime,r-k)
for M in set_of_M:
if len(M) != 0:
s_M = sum(M)
for j in range(0,len(self.mu[self.d-e+k-1])):
if k!=0 or j != 0:
if e+1 < self.d:
value = lambp - s_M +tworhoIprime+self.mu[self.d-e+k-1][j]
if self.Ldominant(value) and value not in self.Cvan[e+1]:
return False
return True
else:
return False
#Compute a set of vanishing results from the flag bundle of type Iprime
#for degrees > e and between kmin and kmax
def compute(self,T,e,kmin,kmax):
res = False
Iprime = self.buildIprime(T)
phi_Iprime, phiLIprime = self.buildRoots(Iprime)
tworhoIprime = sum(x for x in phiLIprime)
kvalues = self.buildWeights(kmin,kmax)
timer = tqdm(kvalues)
for char in timer:
if not self.contains(self.Cvan[e],self.L(char)):
if self.degeneration(Iprime,phiLIprime,tworhoIprime,e,char):
res = True
for k in range(e,self.d):
self.Cvan[k].append(self.L(char))
return res
#Compute a set of vanishing results between kmin and kmax
def computeAll(self,kmin,kmax):
out = False
Iprimes = []
for i in range(1 << self.g-1):
T = [0 for i in range(self.g-1)]
for j in range(self.g-1):
if (i & (1 << j)):
T[j] = 1
Iprimes.append(T)
step = 1
max_step = len(Iprimes)*(self.d)
for T in Iprimes:
for e in range(self.d):
print('Step '+str(step)+'/'+str(max_step)+' : e = '+str(e)+', Iprime = '+str(T))
res = self.compute(T,e,kmin,kmax)
out = out or res
step = step +1
return out
#Technical conversion of the set of vanishing results
def convert(self):
res = []
added = []
for c in self.Cvan:
a = []
for x in c:
if x not in added:
b = []
for i in range(self.g):
b.append(x[i])
a.append(b)
added.append(x)
res.append(a)
return res
#Save vanishing results in a file
def save(self):
res = self.convert()
for k in range(self.d):
filepath = 'g'+str(self.g)+'p'+str(self.p)+'_'+str(k)+'.txt'
with open('save/'+filepath,'w') as f:
for item in res[k]:
for x in item:
f.write("%s " % x)
f.write("\n")
print('Results saved in '+filepath)
#Load vanishing results from a file
def load(self):
for k in range(self.d):
filepath = 'g'+str(self.g)+'p'+str(self.p)+'_'+str(k)+'.txt'
with open('save/'+filepath,'r') as f:
data = f.read()
list_data = data.split('\n')
chars = []
for x in list_data:
y = x.split(' ')
y.pop()
char = [int(z) for z in y]
if len(char) == self.g:
chars.append(char)
for i in range(k,self.d):
for char in chars:
if not self.vanishes(i,char):
self.Cvan[i].append(char)
print('Results loaded from '+filepath)
#Get statistics about the results
def statistics(self):
res = self.convert()
for i in range(self.d):
print('H^* is concentrated in degrees [0:' + str(i) + '] for ' + str(len(res[i])) + ' characters')
#Get psmall weights for Sp_2g twisted by -w0
def psmall(self,kmax):
kmin = 0
kvalues = self.buildWeights(kmin,kmax)
rho = 1/2*sum(self.phi_G)
res = []
for char in kvalues:
is_psmall = True
for alpha in self.phi_G:
prod = 0
for j in range(self.g):
prod += (char[j]+rho[j])*alpha[j]
if max(prod,-prod) > self.p:
is_psmall = False
break
if is_psmall:
res += [tuple(self.changeConvention(char))]
filepath = 'g'+str(self.g)+'p'+str(self.p)+'_psmall'+'.txt'
with open('save/'+filepath,'w') as f:
for char in res:
for x in char:
f.write("%s " % x)
f.write("\n")
print('p-small weights saved in '+filepath)
print(res)
```
# Basic use examples
We create the Siegel threefold $X$ over $\mathbb{F}_7$.
```
X = SiegelVariety(g = 2, p = 7)
```
If the next line returns True, it means that the automorphic line bundle $\mathcal{L}_{(-2,-8)}$ is $D$-ample on the complete flag variety $Y$ over $X$.
```
X.ample([],[-2,-8])
```
The next line compute vanishing results for characters $\lambda = (k_1,k_2)$ with $ -50 \leq k_2 \leq k_1 \leq 0$ using the function $g_{I_0,e}$ in the case where $I_0 = \emptyset$ and $e = 0$. The results are registered in the list $C_{\text{van}}$. It returns True if the algorithm has found new vanishing results.
```
X.compute([], e = 0, kmin = -50, kmax = 0)
```
The next line runs the compute method for each $I_0 \subset I$ and $0 \leq e \leq d$. We only need to specify the range of characters $\lambda = (k_1,k_2)$ we want to consider. It returns True if the algorithm has found new vanishing results. You may want to run this command several times until it returns False.
```
X.computeAll(-50,0)
```
The next line returns True if we know that $H^i(X,\nabla(-4,-6)(-D)) = 0$ for all $i>1$.
```
X.vanishes(1,(-4,-6))
```
If the next line returns False, it means we don't know if $H^i(X,\nabla(-4,-6)(-D)) = 0$ for all $i>0$.
```
X.vanishes(0,(-4,-6))
X.statistics()
```
# Save/Load results
```
X.save()
X.load()
```
# Other computations
We compute and save vanishing results for different values of $p$ when $g = 2$
```
p_values = [5,7,11,31]
for p in p_values:
X_p = SiegelVariety(g = 2, p = p)
found = True
while found:
found = X_p.computeAll(-50,0)
X_p.save()
```
We compute and save vanishing results for different values of $p$ when $g = 3$
```
p_values = [11,31,691]
for p in p_values:
X_p = SiegelVariety(g = 3, p = p)
i = 2
while i>0:
X_p.computeAll(-25,0)
i -= 1
X_p.save()
```
We compute and save some $\text{Sp}_{2g}$ p-small weights for different values of $p$ when $g = 2$
```
p_values = [5,7,11,31]
for p in p_values:
X_p = SiegelVariety(g = 2, p = p)
X_p.psmall(6*p)
```
# Plotting tools
```
#Plot our results when g = 2 or g = 3
def plot(variety):
colors = ['k','b','r','g','c']
if variety.g == 2:
res = variety.convert()
plt.clf()
fig = plt.figure(figsize=(12,6))
for k in range(variety.d):
if len(res[k]) != 0:
x, y = zip(*res[k])
plt.scatter(x,y, label='Concentrated in degrees [0 : ' + str(k) + ']' , color = colors[k%len(colors)])
plt.title('Vanishing results for Siegel threefold with p = ' + str(variety.p))
plt.legend()
plt.show()
elif variety.g == 3:
res = variety.convert()
plt.clf()
fig = plt.figure(figsize=(25,15))
ax = plt.axes(projection='3d')
for k in range(variety.d):
if len(res[k]) != 0:
x, y, z = zip(*res[k])
ax.scatter3D(x,y,z, label='Concentrated in degrees [0 : ' + str(k) + ']' , color = colors[k%len(colors)])
plt.title('Vanishing results for Siegel variety of genus 3 with p = ' + str(variety.p))
plt.legend()
plt.show()
else:
print('The genus must be 2 or 3')
X = SiegelVariety(g = 2, p = 11)
X.load()
plot(X)
Y = SiegelVariety(g = 3, p = 11)
Y.load()
plot(Y)
```
| github_jupyter |
# Working with Text data
```
import numpy as np
import matplotlib.pyplot as plt
% matplotlib inline
```
#### Applying bag-of-words to a toy dataset
```
bards_words = ["The fool doth think he is wise,",
"but the wise man knows himself to be a fool"]
from sklearn.feature_extraction.text import CountVectorizer
vect = CountVectorizer()
vect.fit(bards_words)
print("Vocabulary size: {}".format(len(vect.vocabulary_)))
print("Vocabulary content:\n {}".format(vect.vocabulary_))
bag_of_words = vect.transform(bards_words)
print("bag_of_words: {}".format(repr(bag_of_words)))
print("Dense representation of bag_of_words:\n{}".format(
bag_of_words.toarray()))
vect.get_feature_names()
vect.inverse_transform(bag_of_words)
```
### Task 1
Compute bigrams and trigrams of words as well. How does that change the vocabulary size? How would you imagine this changes the vocabulary size in a real application?
# Download data from http://ai.stanford.edu/~amaas/data/sentiment/
# Delete the ``train/unsup`` folder.
# Sentiment analysis of movie reviews
```
from sklearn.datasets import load_files
reviews_train = load_files("aclImdb/train/")
# load_files returns a bunch, containing training texts and training labels
text_train, y_train = reviews_train.data, reviews_train.target
print("type of text_train: {}".format(type(text_train)))
print("length of text_train: {}".format(len(text_train)))
print("text_train[1]:\n{}".format(text_train[1]))
text_train = [doc.replace(b"<br />", b" ") for doc in text_train]
print("Samples per class (training): {}".format(np.bincount(y_train)))
reviews_test = load_files("aclImdb/test/")
text_test, y_test = reviews_test.data, reviews_test.target
print("Number of documents in test data: {}".format(len(text_test)))
print("Samples per class (test): {}".format(np.bincount(y_test)))
text_test = [doc.replace(b"<br />", b" ") for doc in text_test]
```
### Representing text data as Bag of Words

### Task 2
Use the ``CountVectorizer`` to build a vocabulary and create a bag of word representation of the training data.
How big is the vocabulary?
Display some of the words in the vocabulary using ``get_feature_names()``.
What is 5 most common words?
What is the 5 most common words with ``stop_words='english'``?
```
vect = CountVectorizer()
# ... solution here ...
```
### Task 3
Build a ``LogisticRegression`` model on the dataset.
Extract the features with the largest coefficients (10 most positive and 10 most negative) and visualize them in a bar plot.
Do these make sense?
Then evaluate the model on the test set.
```
from sklearn.linear_model import LogisticRegression
# ...
```
### Task 4
Use stop words and a minimum document frequency to limit the number of features. How does that impact the result?
Then add bigrams.
### Task 5
Use GridSearchCV to adjust the ``C`` parameter in LogisticRegression.
## Pipelines
Scikit-learn has a ``Pipeline`` class that allows chaining of multiple transformations with a model. We can chain the ``CountVectorizer`` with the ``LogisticRegression`` and search over the ``C`` and the size of the n-grams at the same time
### Task 6
Complete the code below to search over whether to use unigrams or unigrams and bigrams.
```
from sklearn.pipeline import make_pipeline
pipeline = make_pipeline(CountVectorizer(), LogisticRegression())
# there are two steps in the pipeline, here are their names:
print(pipeline.named_steps.keys())
print(pipeline.named_steps)
param_grid = {'countvectorizer__ngram_range': #fill in ngram settings,
'logisticregression__C': # fill in settings of C}
grid = GridSearchCV(pipeline, param_grid, cv=5, verbose=10)
grid.fit(text_train, y_train)
# look at grid.cv_results_
```
| github_jupyter |
# Implementing scikitlearn estimators
In this notebook we implement scikit learn estimators to estimate the sparsity pattern in deepmod. Code has been modified already, we just need to build a new training loop.
```
import numpy as np
import pandas as pd
import torch
from DeePyMoD_SBL.data import Burgers
from DeePyMoD_SBL.deepymod_torch.library_functions import library_1D_in
from DeePyMoD_SBL.deepymod_torch.DeepMod import DeepModDynamic
from DeePyMoD_SBL.deepymod_torch.output import Tensorboard, progress
from DeePyMoD_SBL.deepymod_torch.losses import reg_loss, mse_loss
from DeePyMoD_SBL.deepymod_torch.training import train
from DeePyMoD_SBL.deepymod_torch.sparsity import scaling
if torch.cuda.is_available():
torch.set_default_tensor_type('torch.cuda.FloatTensor')
import matplotlib.pyplot as plt
import time
%load_ext autoreload
%autoreload 2
```
# Making data
```
x = np.linspace(-2, 5, 75)
t = np.linspace(0.5, 5.0, 25)
x_grid, t_grid = np.meshgrid(x, t, indexing='ij')
dataset = Burgers(0.1, 1.0)
u = dataset.solution(x_grid, t_grid)
X = np.concatenate((t_grid.reshape(-1, 1), x_grid.reshape(-1, 1)), axis=1)
y = u.reshape(-1, 1)
X_train = torch.tensor(X, dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y, dtype=torch.float32, requires_grad=True)
noise_level = 0.05
X_train = torch.tensor(X, dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y + noise_level * np.std(y) * np.random.randn(y.size, 1), dtype=torch.float32, requires_grad=True)
```
# Finished run
Let's start by trying to one to run on a finished run:
```
config = {'n_in': 2, 'hidden_dims': [20, 20, 20, 20], 'n_out': 1, 'library_function':library_1D_in, 'library_args':{'poly_order':2, 'diff_order': 2}, 'fit_method':'lstsq'}
model = DeepModDynamic(**config)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train(model, X_train, y_train, optimizer, 5000, loss_func_args={'l1':0.0})
model.fit.coeff_vector
```
So that's already decent. Now let's build a function for the SKLearn estimator
```
from sklearn.linear_model import Lasso
regression_func = Lasso(alpha = 1e-4, fit_intercept=False, warm_start=True)
_, time_deriv_list, _, _, theta = model(X_train)
# Normalizing output
dt = (time_deriv_list[0] / torch.norm(time_deriv_list[0])).detach().cpu().numpy()
library = (theta / torch.norm(theta, dim=0, keepdim=True)).detach().cpu().numpy()
regression_func.fit(library, dt)
library
regression_func.coef_
torch.tensor(regression_func.coef_ != 0.0, dtype=torch.bool)
```
So that works. Let's set it up in a function
```
def determine_sparsity_mask(estimator, X, y):
# Normalizing inputs
y_normed = (y / torch.norm(y, keepdim=True)).detach().cpu().numpy()
X_normed = (X / torch.norm(X, dim=0, keepdim=True)).detach().cpu().numpy()
estimator.fit(X_normed, y_normed)
sparsity_mask = torch.tensor(estimator.coef_ != 0.0, dtype=torch.bool)
print(sparsity_mask)
print(estimator.coef_)
return sparsity_mask
determine_sparsity_mask(regression_func, theta, time_deriv_list[0])
model.fit.sparsity_mask
def train_dynamic(model, data, target, optimizer, max_iterations, sparsity_estimator):
'''Trains the deepmod model with MSE, regression and l1 cost function. Updates model in-place.'''
start_time = time.time()
number_of_terms = [coeff_vec.shape[0] for coeff_vec in model(data)[3]]
board = Tensorboard(number_of_terms)
# Training
print('| Iteration | Progress | Time remaining | Cost | MSE | Reg | L1 |')
for iteration in torch.arange(0, max_iterations + 1):
# Calculating prediction and library and scaling
prediction, time_deriv_list, sparse_theta_list, coeff_vector_list, theta = model(data)
coeff_vector_scaled_list = scaling(coeff_vector_list, sparse_theta_list, time_deriv_list)
# Calculating loss
loss_reg = reg_loss(time_deriv_list, sparse_theta_list, coeff_vector_list)
loss_mse = mse_loss(prediction, target)
loss = torch.sum(loss_reg) + torch.sum(loss_mse)
# Writing
if iteration % 100 == 0:
progress(iteration, start_time, max_iterations, loss.item(), torch.sum(loss_mse).item(), torch.sum(loss_reg).item(), torch.sum(loss_reg).item())
board.write(iteration, loss, loss_mse, loss_reg, loss_reg, coeff_vector_list, coeff_vector_scaled_list)
# Optimizer step
optimizer.zero_grad()
loss.backward()
optimizer.step()
# Updating sparsity pattern
if iteration % 200 == 0:
with torch.no_grad():
model.fit.sparsity_mask = [determine_sparsity_mask(sparsity_estimator, theta, time_deriv_list[0])]
board.close()
lasso_estimator = Lasso(alpha = 1e-4, fit_intercept=False, warm_start=True)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train_dynamic(model, X_train, y_train, optimizer, 5000, lasso_estimator)
model.fit.sparsity_mask
model.fit.coeff_vector
```
Fuck it works!!! Now let's try it out with other estimators
# Trying with lassocv
```
config = {'n_in': 2, 'hidden_dims': [20, 20, 20, 20], 'n_out': 1, 'library_function':library_1D_in, 'library_args':{'poly_order':2, 'diff_order': 2}, 'fit_method':'lstsq'}
model = DeepModDynamic(**config)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train(model, X_train, y_train, optimizer, 5000, loss_func_args={'l1':0.0})
from sklearn.linear_model import LassoCV
lassocv_estimator = LassoCV(fit_intercept=False)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train_dynamic(model, X_train, y_train, optimizer, 5000, lassocv_estimator)
model.fit.sparsity_mask
model.fit.coeff_vector
lassocv_estimator.coef_
lasso_estimator.alpha
```
# LassoIC
```
config = {'n_in': 2, 'hidden_dims': [20, 20, 20, 20], 'n_out': 1, 'library_function':library_1D_in, 'library_args':{'poly_order':2, 'diff_order': 2}, 'fit_method':'lstsq'}
model = DeepModDynamic(**config)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train(model, X_train, y_train, optimizer, 5000, loss_func_args={'l1':0.0})
from sklearn.linear_model import LassoLarsIC
lassocv_estimator = LassoLarsIC(fit_intercept=False)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train_dynamic(model, X_train, y_train, optimizer, 5000, lassocv_estimator)
lassocv_estimator.coef_
model.fit.coeff_vector
```
# Testing code
```
import numpy as np
import pandas as pd
import torch
from DeePyMoD_SBL.data import Burgers
from DeePyMoD_SBL.deepymod_torch.library_functions import library_1D_in
from DeePyMoD_SBL.deepymod_torch.DeepMod import DeepModDynamic
from DeePyMoD_SBL.deepymod_torch.training import train_dynamic
from sklearn.linear_model import LassoLarsIC
if torch.cuda.is_available():
torch.set_default_tensor_type('torch.cuda.FloatTensor')
import matplotlib.pyplot as plt
import time
%load_ext autoreload
%autoreload 2
```
# Making data
```
x = np.linspace(-2, 5, 75)
t = np.linspace(0.5, 5.0, 25)
x_grid, t_grid = np.meshgrid(x, t, indexing='ij')
dataset = Burgers(0.1, 1.0)
u = dataset.solution(x_grid, t_grid)
X = np.concatenate((t_grid.reshape(-1, 1), x_grid.reshape(-1, 1)), axis=1)
y = u.reshape(-1, 1)
X_train = torch.tensor(X, dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y, dtype=torch.float32, requires_grad=True)
y_train.shape
noise_level = 0.0
X_train = torch.tensor(X, dtype=torch.float32, requires_grad=True)
y_train = torch.tensor(y + noise_level * np.std(y) * np.random.randn(y.size, 1), dtype=torch.float32, requires_grad=True)
```
# Testing normal
```
estimator = LassoLarsIC(fit_intercept=False)
config = {'n_in': 2, 'hidden_dims': [30, 30, 30, 30, 30], 'n_out': 1, 'library_function':library_1D_in, 'library_args':{'poly_order':2, 'diff_order': 2}, 'sparsity_estimator': estimator}
model = DeepModDynamic(**config)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train_dynamic(model, X_train, y_train, optimizer, 15000, loss_func_args={'sparsity_update_period': 200, 'start_sparsity_update': 5000})
config = {'n_in': 2, 'hidden_dims': [30, 30, 30, 30, 30], 'n_out': 1, 'library_function':library_1D_in, 'library_args':{'poly_order':2, 'diff_order': 2}, 'sparsity_estimator': estimator}
model = DeepModDynamic(**config)
optimizer = torch.optim.Adam(model.network_parameters(), betas=(0.99, 0.999), amsgrad=True)
train_dynamic(model, X_train, y_train, optimizer, 15000, loss_func_args={'sparsity_update_period': 200, 'start_sparsity_update': 5000})
model.sparsity_estimator.coef_
model.constraints.coeff_vector
```
let's do another 5000, see if its better:
```
train_dynamic(model, X_train, y_train, optimizer, 5000, loss_func_args={'sparsity_update_period': 200, 'start_sparsity_update': 0})
model.sparsity_estimator.coef_
model.constraints.coeff_vector
```
# Analysing
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
from tensorboard.backend.event_processing.event_accumulator import EventAccumulator
def load_tensorboard(path):
# Assumes one file per folder
event_file = next(filter(lambda filename: filename[:6] =='events', os.listdir(path)))
summary_iterator = EventAccumulator(str(path + event_file)).Reload()
tags = summary_iterator.Tags()['scalars']
steps = np.array([event.step for event in summary_iterator.Scalars(tags[0])])
data = np.array([[event.value for event in summary_iterator.Scalars(tag)] for tag in tags]).T
df = pd.DataFrame(data=data, index=steps, columns=tags)
return df
df = load_tensorboard('runs/Apr20_09-11-22_4b6076e78386/')
df.keys()
plt.semilogy(df['MSE_0'])
plt.semilogy(df['Regression_0'])
coeff_keys = [key for key in df.keys() if key[:5]=='coeff']
for key in coeff_keys:
plt.plot(df[key], label=f'{key[-1]}')
plt.legend()
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Automatic differentiation and gradient tape
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/beta/tutorials/eager/automatic_differentiation"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/site/en/r2/tutorials/eager/automatic_differentiation.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
</table>
In the previous tutorial we introduced `Tensor`s and operations on them. In this tutorial we will cover [automatic differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation), a key technique for optimizing machine learning models.
## Setup
```
from __future__ import absolute_import, division, print_function, unicode_literals
!pip install tensorflow==2.0.0-beta0
import tensorflow as tf
```
## Gradient tapes
TensorFlow provides the [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) API for automatic differentiation - computing the gradient of a computation with respect to its input variables. Tensorflow "records" all operations executed inside the context of a `tf.GradientTape` onto a "tape". Tensorflow then uses that tape and the gradients associated with each recorded operation to compute the gradients of a "recorded" computation using [reverse mode differentiation](https://en.wikipedia.org/wiki/Automatic_differentiation).
For example:
```
x = tf.ones((2, 2))
with tf.GradientTape() as t:
t.watch(x)
y = tf.reduce_sum(x)
z = tf.multiply(y, y)
# Derivative of z with respect to the original input tensor x
dz_dx = t.gradient(z, x)
for i in [0, 1]:
for j in [0, 1]:
assert dz_dx[i][j].numpy() == 8.0
```
You can also request gradients of the output with respect to intermediate values computed during a "recorded" `tf.GradientTape` context.
```
x = tf.ones((2, 2))
with tf.GradientTape() as t:
t.watch(x)
y = tf.reduce_sum(x)
z = tf.multiply(y, y)
# Use the tape to compute the derivative of z with respect to the
# intermediate value y.
dz_dy = t.gradient(z, y)
assert dz_dy.numpy() == 8.0
```
By default, the resources held by a GradientTape are released as soon as GradientTape.gradient() method is called. To compute multiple gradients over the same computation, create a `persistent` gradient tape. This allows multiple calls to the `gradient()` method. as resources are released when the tape object is garbage collected. For example:
```
x = tf.constant(3.0)
with tf.GradientTape(persistent=True) as t:
t.watch(x)
y = x * x
z = y * y
dz_dx = t.gradient(z, x) # 108.0 (4*x^3 at x = 3)
dy_dx = t.gradient(y, x) # 6.0
del t # Drop the reference to the tape
```
### Recording control flow
Because tapes record operations as they are executed, Python control flow (using `if`s and `while`s for example) is naturally handled:
```
def f(x, y):
output = 1.0
for i in range(y):
if i > 1 and i < 5:
output = tf.multiply(output, x)
return output
def grad(x, y):
with tf.GradientTape() as t:
t.watch(x)
out = f(x, y)
return t.gradient(out, x)
x = tf.convert_to_tensor(2.0)
assert grad(x, 6).numpy() == 12.0
assert grad(x, 5).numpy() == 12.0
assert grad(x, 4).numpy() == 4.0
```
### Higher-order gradients
Operations inside of the `GradientTape` context manager are recorded for automatic differentiation. If gradients are computed in that context, then the gradient computation is recorded as well. As a result, the exact same API works for higher-order gradients as well. For example:
```
x = tf.Variable(1.0) # Create a Tensorflow variable initialized to 1.0
with tf.GradientTape() as t:
with tf.GradientTape() as t2:
y = x * x * x
# Compute the gradient inside the 't' context manager
# which means the gradient computation is differentiable as well.
dy_dx = t2.gradient(y, x)
d2y_dx2 = t.gradient(dy_dx, x)
assert dy_dx.numpy() == 3.0
assert d2y_dx2.numpy() == 6.0
```
## Next Steps
In this tutorial we covered gradient computation in TensorFlow. With that we have enough of the primitives required to build and train neural networks.
```
```
| github_jupyter |
### Created by Tirthajyoti Sarkar, Ph.D., Jan 2018
# Mean-shift Clustering Technique
Mean-shift clustering aims to discover blobs in a smooth density of samples. It is a centroid based algorithm, which works by updating candidates for centroids to be the mean of the points within a given region. These candidates are then filtered in a post-processing stage to eliminate near-duplicates to form the final set of centroids.
Given a candidate centroid $x_i$ for iteration $t$, the candidate is updated according to the following equation:
$$x_i^{t+1} = x_i^t + m(x_i^t)$$
Where $N(x_i)$ is the neighborhood of samples within a given distance around $x_i$ and $m$ is the mean shift vector that is computed for each centroid that points towards a region of the maximum increase in the density of points. This is computed using the following equation, effectively updating a centroid to be the mean of the samples within its neighborhood:
$$m(x_i) = \frac{\sum_{x_j \in N(x_i)}K(x_j - x_i)x_j}{\sum_{x_j \in N(x_i)}K(x_j - x_i)}$$
**The algorithm automatically sets the number of clusters, instead of relying on a parameter bandwidth, which dictates the size of the region to search through**. This parameter can be set manually, but can be estimated using the provided estimate_bandwidth function, which is called if the bandwidth is not set.
**The algorithm is not highly scalable, as it requires multiple nearest neighbor searches** during the execution of the algorithm. The algorithm is guaranteed to converge, however the algorithm will stop iterating when the change in centroids is small.
## Make the synthetic data
```
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from sklearn.cluster import MeanShift
from sklearn import metrics
from sklearn.datasets.samples_generator import make_blobs
# Generate sample data
centers = [[1, 1], [-1, -1], [1, -1]]
X, labels_true = make_blobs(n_samples=300, centers=centers, cluster_std=0.4,random_state=101)
X.shape
plt.figure(figsize=(8,5))
plt.scatter(X[:,0],X[:,1],edgecolors='k',c='orange',s=75)
plt.grid(True)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.show()
```
## Clustering
```
ms_model = MeanShift().fit(X)
cluster_centers = ms_model.cluster_centers_
labels = ms_model.labels_
n_clusters = len(cluster_centers)
labels = ms_model.labels_
```
#### Number of detected clusters and their centers
```
print("Number of clusters detected by the algorithm:", n_clusters)
print("Cluster centers detected at:\n\n", cluster_centers)
plt.figure(figsize=(8,5))
plt.scatter(X[:,0],X[:,1],edgecolors='k',c=ms_model.labels_,s=75)
plt.grid(True)
plt.xticks(fontsize=15)
plt.yticks(fontsize=15)
plt.show()
```
#### Homogeneity
Homogeneity metric of a cluster labeling given a ground truth.
A clustering result satisfies homogeneity if all of its clusters contain only data points which are members of a single class. This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
```
print ("Homogeneity score:", metrics.homogeneity_score(labels_true,labels))
```
#### Completeness
Completeness metric of a cluster labeling given a ground truth.
A clustering result satisfies completeness if all the data points that are members of a given class are elements of the same cluster. This metric is independent of the absolute values of the labels: a permutation of the class or cluster label values won’t change the score value in any way.
```
print("Completeness score:",metrics.completeness_score(labels_true,labels))
```
## Time complexity and model quality as the data size grows
```
import time
from tqdm import tqdm
n_samples = [10,20,50,100,200,500,1000,2000,3000,5000,7500,10000]
centers = [[1, 1], [-1, -1], [1, -1]]
t_ms = []
homo_ms=[]
complete_ms=[]
for i in tqdm(n_samples):
X,labels_true = make_blobs(n_samples=i, centers=centers, cluster_std=0.4,random_state=101)
t1 = time.time()
ms_model = MeanShift().fit(X)
t2=time.time()
t_ms.append(t2-t1)
homo_ms.append(metrics.homogeneity_score(labels_true,ms_model.labels_))
complete_ms.append(metrics.completeness_score(labels_true,ms_model.labels_))
plt.figure(figsize=(8,5))
plt.title("Time complexity of Mean Shift\n",fontsize=20)
plt.scatter(n_samples,t_ms,edgecolors='k',c='green',s=100)
plt.plot(n_samples,t_ms,'k--',lw=3)
plt.grid(True)
plt.xticks(fontsize=15)
plt.xlabel("Number of samples",fontsize=15)
plt.yticks(fontsize=15)
plt.ylabel("Time taken for model (sec)",fontsize=15)
plt.show()
plt.figure(figsize=(8,5))
plt.title("Homogeneity score with data set size\n",fontsize=20)
plt.scatter(n_samples,homo_ms,edgecolors='k',c='green',s=100)
plt.plot(n_samples,homo_ms,'k--',lw=3)
plt.grid(True)
plt.xticks(fontsize=15)
plt.xlabel("Number of samples",fontsize=15)
plt.yticks(fontsize=15)
plt.ylabel("Homogeneity score",fontsize=15)
plt.show()
plt.figure(figsize=(8,5))
plt.title("Completeness score with data set size\n",fontsize=20)
plt.scatter(n_samples,complete_ms,edgecolors='k',c='green',s=100)
plt.plot(n_samples,complete_ms,'k--',lw=3)
plt.grid(True)
plt.xticks(fontsize=15)
plt.xlabel("Number of samples",fontsize=15)
plt.yticks(fontsize=15)
plt.ylabel("Completeness score",fontsize=15)
plt.show()
```
## How well the cluster detection works in the presence of noise?
Create data sets with varying degree of noise std. dev and run the model to detect clusters.
```
noise = [0.01,0.05,0.1,0.2,0.3,0.4,0.5,0.6,0.7,0.8,0.9,1.0,1.25,1.5,1.75,2.0]
n_clusters = []
for i in noise:
centers = [[1, 1], [-1, -1], [1, -1]]
X, labels_true = make_blobs(n_samples=200, centers=centers, cluster_std=i,random_state=101)
ms_model=MeanShift().fit(X)
n_clusters.append(len(ms_model.cluster_centers_))
print("Detected number of clusters:",n_clusters)
plt.figure(figsize=(8,5))
plt.title("Cluster detection with noisy data\n",fontsize=20)
plt.scatter(noise,n_clusters,edgecolors='k',c='green',s=100)
plt.grid(True)
plt.xticks(fontsize=15)
plt.xlabel("Noise std.dev",fontsize=15)
plt.yticks(fontsize=15)
plt.ylabel("Number of clusters detected",fontsize=15)
plt.show()
```
** We see that the cluster detection works well up to a certain level of noise std. dev, after which the mean of the blobs shifts to the overall centroid and the number of detected clusters tends to 1**
| github_jupyter |
Follow the Sherlock data repo for complete installation instruction: https://github.com/mitmedialab/sherlock-project/tree/8d6411d793dfcfacae0bd300b806e023d0644e95
```
import pandas as pd
import numpy as np
import sys
import tensorflow as tf
import matplotlib.pyplot as plt
from ast import literal_eval
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import f1_score
import re
sys.path.append("..")
from src.features.build_features import build_features
from src.features.build_features import _get_data
from src.deploy.train_sherlock import train_sherlock
from src.deploy.predict_sherlock import predict_sherlock
_get_data()
testdf = pd.read_csv('../../Benchmark-Labeled-Data/data_test.csv')
test_metadata = pd.read_csv('../../RawCSV/Metadata/meta_data.csv')
test_merged = pd.merge(testdf,test_metadata,on='Record_id')
y_true = test_merged.y_act.values.tolist()
test_merged['list_vals'] = ""
test_merged
for row in test_merged.itertuples():
if row.Index%100 == 0: print(row.Index)
col = getattr(row,'Attribute_name')
csv_name = '../../RawCSV/RawCSVFiles/' + getattr(row,'name')
df = pd.read_csv(csv_name,encoding='latin1')
try:
df_col = df[col].tolist()
test_merged.at[row.Index,'list_vals'] = df_col
except KeyError: test_merged.at[row.Index,'list_vals'] = []
sherlock_df = test_merged[["list_vals", "y_act"]]
for index, row in sherlock_df.iterrows():
if row["list_vals"] == []:
sherlock_df.at[index, "list_vals"] = [""]
else:
templst = []
flag = 1
for x in row["list_vals"]:
if x != x: continue
try: abc = int(x)
except:
flag = 0
break
try: abc = float(x)
except:
flag = 0
break
for x in row["list_vals"]:
if x != x:
templst.append("0")
continue
if flag: temp = x
else: temp = x
templst.append(temp)
sherlock_df.at[index, "list_vals"] = templst
sherlock_df
sherlock_df1 = sherlock_df['list_vals']
sherlock_df1
X_test = build_features(sherlock_df1)
predicted_labels = predict_sherlock(X_test, nn_id='sherlock')
predicted_labels
label_df = pd.DataFrame(predicted_labels,columns=['label'])
df = test_merged[['y_act', 'Attribute_name','sample_1','sample_2','sample_3','sample_4','sample_5','total_vals', 'num_nans', '%_nans', 'num_of_dist_val' ,'%_dist_val']]
df = df.fillna(0)
df['label'] = label_df['label']
df['ColumnA'] = df[df.columns[2:7]].apply(lambda x: '$#$'.join(x.dropna().astype(str)), axis=1 )
curdf = pd.read_csv('Semantic2FeatureType_Mapping.csv') # Load Semantic Types to Feature Type Mappings
curdic = {}
for i,row in curdf.iterrows():
if row['type'] not in curdic:
curdic[row['type']] = []
if row['l0'] == row['l0']: curdic[row['type']].append(row['l0'])
if row['l1'] == row['l1']: curdic[row['type']].append(row['l1'])
if row['l2'] == row['l2']: curdic[row['type']].append(row['l2'])
if row['l3'] == row['l3']: curdic[row['type']].append(row['l3'])
delimeters = r"(,|;|\|)"
delimeters = re.compile(delimeters)
del_pattern = r"\b[0-9]+[a-zA-Z \% \$]+"
del_reg = re.compile(del_pattern)
def func(lst):
lst = list(lst.split('$#$'))
try:
lst = [float(i) for i in lst]
except ValueError: f=1
if all(isinstance(x, int) for x in lst) or all(isinstance(x, float) for x in lst): return 1
else: return 0
df['isNumeric'] = df['ColumnA'].apply(lambda x: func(x))
ysherlock = []
yact = []
for i,row in df.iterrows():
if len(curdic[row['label']]) == 1:
ysherlock.append(curdic[row['label']][0])
yact.append(row['y_act'])
elif row['label'] in ['age', 'result', 'plays', 'ranking']:
if row['isNumeric'] == 1: ysherlock.append('Numeric')
elif del_reg.match(str(row['sample_1'])) or del_reg.match(str(row['sample_2'])) or del_reg.match(str(row['sample_3'])) or del_reg.match(str(row['sample_4'])) or del_reg.match(str(row['sample_5'])):
ysherlock.append('Embedded Number')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['sales', 'rank', 'elevation', 'weight'] :
if row['isNumeric'] == 1: ysherlock.append('Numeric')
else: ysherlock.append('Embedded Number')
yact.append(row['y_act'])
elif row['label'] in ['area','position', 'depth']:
if row['isNumeric'] == 1: ysherlock.append('Numeric')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['command']:
templst = [len(str(row['sample_1']).split(' ')), len(str(row['sample_2']).split(' ')), len(str(row['sample_3']).split(' ')), len(str(row['sample_4']).split(' ')), len(str(row['sample_5']).split(' '))]
# print(templst)
if np.mean(templst) > 3: ysherlock.append('Sentence')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['code']:
if row['%_dist_val'] > 99.99 or row['num_of_dist_val'] == 1 or row['total_vals'] == row['num_nans']: ysherlock.append('Not-Generalizable')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['day','duration', 'year']:
try:
pd.Timestamp(row['sample_1'])
ysherlock.append('Datetime')
except ValueError: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['order']:
if row['isNumeric'] == 1: ysherlock.append('Context-Specific')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['range']:
if del_reg.match(str(row['sample_1'])) or del_reg.match(str(row['sample_2'])) or del_reg.match(str(row['sample_3'])) or del_reg.match(str(row['sample_4'])) or del_reg.match(str(row['sample_5'])):
ysherlock.append('Embedded Number')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
elif row['label'] in ['genre','collection']:
if len(delimeters.findall(str(str(row['sample_1'])))) > 1: ysherlock.append('List')
else: ysherlock.append('Categorical')
yact.append(row['y_act'])
# else:
# print(row)
# print(row['label'])
len(ysherlock)
dict_label = {
'Numeric': 0,
'Categorical': 1,
'Datetime':2,
'Sentence':3,
'URL': 4,
'Embedded Number': 5,
'List': 6,
'Not-Generalizable': 7,
'Custom Object': 8,
'Context-Specific': 8
}
ysherlock1 = [dict_label[x] for x in ysherlock]
from sklearn.metrics import accuracy_score, confusion_matrix
print(accuracy_score(yact, ysherlock1))
print(confusion_matrix(yact, ysherlock1))
```
| github_jupyter |
```
from plot_lib import plot_data, plot_model, set_default
set_default()
import torch
import torchvision
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader
from torchvision import datasets, transforms
from torchvision.datasets import FashionMNIST
import matplotlib.pyplot as plt
import numpy
from torchsummary import summary
from torchvision.utils import save_image
import os
# Creating the folder "pictures" where I will save the reconstructed images
path = './pictures'
if not os.path.exists(path):
os.makedirs(path)
# function to count number of parameters
def get_n_params(model):
np=0
for p in list(model.parameters()):
np += p.nelement()
return np
# Convert vector to image
def to_img(x):
x = 0.5 * (x + 1)
x = x.view(x.size(0), 28, 28)
return x
# Displaying routine
def display_images(in_, out, n=1):
for N in range(n):
if in_ is not None:
in_pic = to_img(in_.cpu().data)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(in_pic[i+4*N])
plt.axis('off')
out_pic = to_img(out.cpu().data)
plt.figure(figsize=(18, 6))
for i in range(4):
plt.subplot(1,4,i+1)
plt.imshow(out_pic[i+4*N])
plt.axis('off')
# Define data loading step
batch_size = 128 #256
img_transform = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))
])
dataset = FashionMNIST('./data', transform=img_transform, download=True)
dataloader = DataLoader(dataset, batch_size=batch_size, shuffle=True)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# Define model architecture and reconstruction loss
class Autoencoder(nn.Module):
def __init__(self):
super().__init__()
self.encoder = nn.Sequential( nn.Conv2d(in_channels=1, out_channels=16, kernel_size=3, stride=3, padding=1),
nn.LeakyReLU(), nn.AvgPool2d( kernel_size=2,stride=2, padding=0 ),
nn.Conv2d(16, 8, kernel_size=3, stride=2, padding=1) , nn.LeakyReLU(),
nn.AvgPool2d( kernel_size=2, stride=1, padding=0 )
)
self.decoder = nn.Sequential( nn.ConvTranspose2d(8, 16, kernel_size=3, stride=2, padding=0),
nn.LeakyReLU(), nn.ConvTranspose2d(16, 8, kernel_size=5, stride=3, padding=1),
nn.LeakyReLU(), nn.ConvTranspose2d( 8, 1, kernel_size=2, stride=2, padding=1), nn.Tanh()
)
#self.fc = nn.Linear(8*2*2, 10)
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
return x
model = Autoencoder().to(device)
criterion = nn.MSELoss()
#Print the parameters size of the model
summary(model, (1,28,28))
print(model.parameters)
# Configure the optimiser
learning_rate = 1e-3
L2_regularization = 1e-5
optimizer = torch.optim.Adam(
model.parameters(),
lr=learning_rate, weight_decay = L2_regularization
)
# Train standard or denoising autoencoder (AE)
num_epochs = 20
# do = nn.Dropout() # comment out for standard AE
for epoch in range(num_epochs):
for data in dataloader:
img, label = data
img.requires_grad_()
#img = img.view(img.size(0), -1)
img = img.view(-1,1,28,28)
# img_bad = do(img).to(device) # comment out for standard AE
# ===================forward=====================
output = model(img) # feed <img> (for std AE) or <img_bad> (for denoising AE)
#output = model(label)
#output = output.view(-1,1,28,28)
loss = criterion(output, img.data)
# ===================backward====================
optimizer.zero_grad()
loss.backward()
optimizer.step()
# ===================log========================
print('epoch[{}/{}], loss : {}'.format(epoch + 1, num_epochs, loss.item()))
#Saving the images at each epoch
save_image(output, "./pictures/{}.png".format(epoch+1))
#displaying the images
display_images(None, output) # pass (None, output) for std AE, (img_bad, output) for denoising AE
```
| github_jupyter |
```
import torch
import torch.utils.data
from torch.autograd import Variable
import torch.nn as nn
import torch.optim as optim
import numpy as np
import h5py
from data_utils import get_data
import matplotlib.pyplot as plt
from solver_pytorch import Solver
# Load data from all .mat files, combine them, eliminate EOG signals, shuffle and
# seperate training data, validation data and testing data.
# Also do mean subtraction on x.
data = get_data('../project_datasets',num_validation=100, num_test=100)
for k in data.keys():
print('{}: {} '.format(k, data[k].shape))
# class flatten to connect to FC layer
class Flatten(nn.Module):
def forward(self, x):
N, C, H = x.size() # read in N, C, H
return x.view(N, -1)
# turn x and y into torch type tensor
dtype = torch.FloatTensor
X_train = Variable(torch.Tensor(data.get('X_train'))).type(dtype)
y_train = Variable(torch.Tensor(data.get('y_train'))).type(torch.IntTensor)
X_val = Variable(torch.Tensor(data.get('X_val'))).type(dtype)
y_val = Variable(torch.Tensor(data.get('y_val'))).type(torch.IntTensor)
X_test = Variable(torch.Tensor(data.get('X_test'))).type(dtype)
y_test = Variable(torch.Tensor(data.get('y_test'))).type(torch.IntTensor)
# train a 1D convolutional neural network
# optimize hyper parameters
best_model = None
parameters =[] # a list of dictionaries
parameter = {} # a dictionary
best_params = {} # a dictionary
best_val_acc = 0.0
# hyper parameters in model
filter_nums = [20]
filter_sizes = [20]
pool_sizes = [4]
# hyper parameters in solver
batch_sizes = [100]
lrs = [5e-4]
for filter_num in filter_nums:
for filter_size in filter_sizes:
for pool_size in pool_sizes:
linear_size = int((X_test.shape[2]-filter_size)/4)+1
linear_size = int((linear_size-pool_size)/pool_size)+1
linear_size *= filter_num
for batch_size in batch_sizes:
for lr in lrs:
model = nn.Sequential(
nn.Conv1d(22, filter_num, kernel_size=filter_size, stride=4),
nn.ReLU(inplace=True),
nn.Dropout(p=0.5),
nn.BatchNorm1d(num_features=filter_num),
nn.MaxPool1d(kernel_size=pool_size, stride=pool_size),
Flatten(),
nn.Linear(linear_size, 20),
nn.ReLU(inplace=True),
nn.Linear(20, 4)
)
model.type(dtype)
solver = Solver(model, data,
lr = lr, batch_size=batch_size,
verbose=True, print_every=50)
solver.train()
# save training results and parameters of neural networks
parameter['filter_num'] = filter_num
parameter['filter_size'] = filter_size
parameter['pool_size'] = pool_size
parameter['batch_size'] = batch_size
parameter['lr'] = lr
parameters.append(parameter)
print('Accuracy on the validation set: ', solver.best_val_acc)
print('parameters of the best model:')
print(parameter)
if solver.best_val_acc > best_val_acc:
best_val_acc = solver.best_val_acc
best_model = model
best_solver = solver
best_params = parameter
# Plot the loss function and train / validation accuracies of the best model
plt.subplot(2,1,1)
plt.plot(best_solver.loss_history)
plt.title('Training loss history')
plt.xlabel('Iteration')
plt.ylabel('Training loss')
plt.subplot(2,1,2)
plt.plot(best_solver.train_acc_history, '-o', label='train accuracy')
plt.plot(best_solver.val_acc_history, '-o', label='validation accuracy')
plt.xlabel('Iteration')
plt.ylabel('Accuracies')
plt.legend(loc='upper center', ncol=4)
plt.gcf().set_size_inches(10, 10)
plt.show()
print('Accuracy on the validation set: ', best_val_acc)
print('parameters of the best model:')
print(best_params)
# test set
y_test_pred = model(X_test)
_, y_pred = torch.max(y_test_pred,1)
test_accu = np.mean(y_pred.data.numpy() == y_test.data.numpy())
print('Test accuracy', test_accu, '\n')
```
| github_jupyter |
# Textos (cadenas de caracteres)
```
'Hola Mundo'
"Hola Mundo"
'Este texto incluye unas " " '
"Esta 'palabra' se encuentra escrita entre comillas simples"
"Esta \"palabra\" se encuentra escrita entre comillas dobles"
'Esta \'palabra\' se encuentra escrita entre comillas dobles'
```
## La función print()
Es una instrucción que nos permite mostrar correctamente el valor de una cadena (u otros valores/variables) por pantalla.
```
"Una cadena"
'otra cadena'
'otra cadena más'
print("Una cadena")
print('otra cadena')
print('otra cadena más')
```
#### Acepta carácteres especiales como las tabulaciones /t o los saltos de línea /n
```
print("Un texto\tuna tabulación")
print("Un texto\nuna nueva línea")
```
#### Para evitar los carácteres especiales, debemos indicar que una cadena es cruda (raw)
```
print("C:\nombre\directorio")
print(r"C:\nombre\directorio") # r => raw (cruda)
```
#### Podemos utilizar """ *(triple comillas)* para cadenas multilínea
```
print("""Una línea
otra línea
otra línea\tuna tabulación""")
```
#### También es posible asignar cadenas a variables
La forma correcta de mostrarlas es con la instrucción print().
```
c = "Esto es una cadena\ncon dos líneas"
c
print(c)
```
## Operaciones
Una de las operaciones de las cadenas es la concatenación (o suma de cadenas)
```
c + c
print(c+c)
s = "Una cadena" " compuesta de dos cadenas"
print(s)
c1 = "Una cadena"
c2 = "otra cadena"
print("Una cadena " + c2)
```
#### También es posible utilizar la multiplicación de cadenas
```
diez_espacios = " " * 10
print(diez_espacios + "un texto a diez espacios")
```
## Índices en las cadenas
Los índices nos permiten posicionarnos en un carácter específico de una cadena.
Representan un número [índice], que empezando por el 0 indica el carácter de la primera posición, y así sucesivamente.
```
palabra = "Python"
palabra[0] # carácter en la posición 0
palabra[3]
```
#### El índice negativo -1, hace referencia al carácter de la última posición, el -2 al penúltimo y así sucesivamente
```
palabra[-1]
palabra[-0]
palabra[-2]
palabra[-6]
palabra[5]
```
## Slicing en las cadenas
El slicing es una capacidad de las cadenas que devuelve un subconjunto o subcadena utilizando dos índices [inicio:fin]:
- El primer índice indica donde empieza la subcadena (se incluye el carácter).
- El segundo índice indica donde acaba la subcadena (se excluye el carácter).
```
palabra = "Python"
palabra[0:2]
palabra[2:]
palabra[:2]
```
#### Si en el slicing no se indica un índice se toma por defecto el principio y el final (incluídos)
```
palabra[:]
palabra[:2] + palabra[2:]
palabra[-2:]
```
#### Si un índice se encuentra fuera del rango de la cadena, dará error
```
palabra[99]
```
#### Pero con slicing ésto no pasa y simplemente se ignora el espacio hueco
```
palabra[:99]
palabra[99:]
```
## Inmutabilidad
Una propiedad de las cadenas es que no se pueden modificar. Si intentamos reasignar un carácter, no nos dejará:
```
palabra[0] = "N"
```
#### Sin embargo, utilizando slicing y concatenación podemos generar nuevas cadenas fácilmente:
```
palabra = "N" + palabra[1:]
palabra
```
### Funciones
Un ejemplo de función útil que soportan las cadenas es len(), que nos permite saber su longitud (el número de caracteres que contienen).
```
len(palabra)
```
#### Hay más funciones, pero las iremos descubriendo a lo largo del curso.
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Keras 전처리 레이어를 사용한 구조적 데이터 분류
<table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/tutorials/structured_data/preprocessing_layers"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/tutorials/structured_data/preprocessing_layers.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/tutorials/structured_data/preprocessing_layers.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
<td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/tutorials/structured_data/preprocessing_layers.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드</a></td>
</table>
이 튜토리얼에서는 구조적 데이터(예: CSV의 표 형식 데이터)를 분류하는 방법을 보여줍니다. [Keras](https://www.tensorflow.org/guide/keras)를 사용하여 모델을 정의하고, [전처리 레이어](https://keras.io/guides/preprocessing_layers/)를 CSV의 열에서 모델 훈련에 사용되는 특성으로 매핑하는 브리지로 사용합니다. 이 튜토리얼에는 다음을 위한 전체 코드가 포함되어 있습니다.
- [Pandas](https://pandas.pydata.org/)를 사용하여 CSV 파일을 로드합니다.
- [tf.data](https://www.tensorflow.org/guide/datasets)를 사용하여 행을 일괄 처리하고 셔플하는 입력 파이프라인을 빌드합니다.
- Keras 전처리 레이어를 사용하여 CSV의 열에서 모델을 훈련하는 데 사용되는 특성으로 매핑합니다.
- Keras를 사용하여 모델을 빌드, 훈련 및 평가합니다.
참고: 이 튜토리얼은 [특성 열의 구조적 데이터 분류하기](https://www.tensorflow.org/tutorials/structured_data/feature_columns)와 유사합니다. 이 버전은 `tf.feature_column` 대신 새로운 실험용 Keras [전처리 레이어](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing)를 사용합니다. Keras Preprocessing Layer는 더 직관적이며 배포를 단순화하기 위해 모델 내에 쉽게 포함될 수 있습니다.
## 데이터세트
PetFinder [데이터세트](https://www.kaggle.com/c/petfinder-adoption-prediction)의 단순화된 버전을 사용합니다. CSV에는 수천 개의 행이 있습니다. 각 행은 애완 동물을 설명하고 각 열은 속성을 설명합니다. 이 정보를 사용하여 애완 동물의 입양 여부를 예측합니다.
다음은 이 데이터세트에 대한 설명입니다. 숫자 열과 범주 열이 모두 있습니다. 이 튜토리얼에서 사용하지 않는 자유 텍스트 열이 있습니다.
열 | 설명 | 특성 유형 | 데이터 형식
--- | --- | --- | ---
유형 | 동물의 종류(개, 고양이) | 범주형 | 문자열
나이 | 애완 동물의 나이 | 수치 | 정수
품종 1 | 애완 동물의 기본 품종 | 범주형 | 문자열
색상 1 | 애완 동물의 색상 1 | 범주형 | 문자열
색상 2 | 애완 동물의 색상 2 | 범주형 | 문자열
MaturitySize | 성장한 크기 | 범주형 | 문자열
FurLength | 모피 길이 | 범주형 | 문자열
예방 접종 | 애완 동물이 예방 접종을 받았습니다 | 범주형 | 문자열
불임 시술 | 애완 동물이 불임 시술을 받았습니다 | 범주형 | 문자열
건강 | 건강 상태 | 범주형 | 문자열
회비 | 입양비 | 수치 | 정수
설명 | 이 애완 동물에 대한 프로필 작성 | 텍스트 | 문자열
PhotoAmt | 이 애완 동물의 업로드된 총 사진 | 수치 | 정수
AdoptionSpeed | 입양 속도 | 분류 | 정수
## TensorFlow 및 기타 라이브러리 가져오기
```
!pip install -q sklearn
import numpy as np
import pandas as pd
import tensorflow as tf
from sklearn.model_selection import train_test_split
from tensorflow.keras import layers
from tensorflow.keras.layers.experimental import preprocessing
```
## Pandas를 사용하여 데이터 프레임 만들기
[Pandas](https://pandas.pydata.org/)는 구조적 데이터를 로드하고 처리하는 데 유용한 여러 유틸리티가 포함된 Python 라이브러리입니다. Pandas를 사용하여 URL에서 데이터세트를 다운로드하고 데이터 프레임에 로드합니다.
```
import pathlib
dataset_url = 'http://storage.googleapis.com/download.tensorflow.org/data/petfinder-mini.zip'
csv_file = 'datasets/petfinder-mini/petfinder-mini.csv'
tf.keras.utils.get_file('petfinder_mini.zip', dataset_url,
extract=True, cache_dir='.')
dataframe = pd.read_csv(csv_file)
dataframe.head()
```
## 목표 변수 만들기
Kaggle 대회에서의 작업은 애완 동물이 입양되는 속도를 예측하는 것입니다(예: 첫 주, 첫 달, 첫 3개월 등). 튜토리얼을 위해 단순화해 봅시다. 여기에서는 입양 속도를 이진 분류 문제로 변환하고 단순히 애완 동물이 입양되었는지 여부를 예측합니다.
레이블 열을 수정한 후, 0은 애완 동물이 입양되지 않았음을 나타내고 1은 입양되었음을 나타냅니다.
```
# In the original dataset "4" indicates the pet was not adopted.
dataframe['target'] = np.where(dataframe['AdoptionSpeed']==4, 0, 1)
# Drop un-used columns.
dataframe = dataframe.drop(columns=['AdoptionSpeed', 'Description'])
```
## 데이터 프레임을 훈련, 검증 및 테스트로 분할하기
다운로드한 데이터세트는 단일 CSV 파일입니다. 이를 훈련, 검증 및 테스트 세트로 분할합니다.
```
train, test = train_test_split(dataframe, test_size=0.2)
train, val = train_test_split(train, test_size=0.2)
print(len(train), 'train examples')
print(len(val), 'validation examples')
print(len(test), 'test examples')
```
## tf.data를 사용하여 입력 파이프라인 만들기
다음으로 데이터를 셔플하고 일괄 처리하기 위해 [tf.data](https://www.tensorflow.org/guide/datasets)로 데이터 프레임을 래핑합니다. 매우 큰 CSV 파일(메모리에 적합하지 않을 정도로 큰 파일)을 사용하는 경우, tf.data를 사용하여 디스크에서 직접 읽을 수 있습니다. 이 튜토리얼에서는 다루지 않습니다.
```
# A utility method to create a tf.data dataset from a Pandas Dataframe
def df_to_dataset(dataframe, shuffle=True, batch_size=32):
dataframe = dataframe.copy()
labels = dataframe.pop('target')
ds = tf.data.Dataset.from_tensor_slices((dict(dataframe), labels))
if shuffle:
ds = ds.shuffle(buffer_size=len(dataframe))
ds = ds.batch(batch_size)
ds = ds.prefetch(batch_size)
return ds
```
이제 입력 파이프라인을 생성했으므로 반환되는 데이터의 형식을 확인하기 위해 호출해 보겠습니다. 출력을 읽기 쉽게 유지하기 위해 작은 배치 크기를 사용했습니다.
```
batch_size = 5
train_ds = df_to_dataset(train, batch_size=batch_size)
[(train_features, label_batch)] = train_ds.take(1)
print('Every feature:', list(train_features.keys()))
print('A batch of ages:', train_features['Age'])
print('A batch of targets:', label_batch )
```
데이터세트가 데이터 프레임의 행에서 열 값에 매핑되는 열 이름의 사전(데이터 프레임에서)을 반환하는 것을 볼 수 있습니다.
## 전처리 레이어의 사용을 시연합니다.
Keras 전처리 레이어 API를 사용하면 Keras 네이티브 입력 처리 파이프라인을 빌드할 수 있습니다. 3개의 전처리 레이어를 사용하여 특성 전처리 코드를 보여줍니다.
- [`Normalization`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Normalization) - 데이터의 특성별 정규화입니다.
- [`CategoryEncoding`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/CategoryEncoding) 카테고리 인코딩 레이어입니다.
- [`StringLookup`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/StringLookup) - 어휘의 문자열을 정수 인덱스로 매핑합니다.
- [`IntegerLookup`](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/IntegerLookup) - 어휘의 정수를 정수 인덱스로 매핑합니다.
[여기](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing)에서 사용 가능한 전처리 레이어의 목록을 찾을 수 있습니다.
### 숫자 열
각 숫자 특성에 대해 Normalization() 레이어를 사용하여 각 특성의 평균이 0이고 표준 편차가 1인지 확인합니다.
`get_normalization_layer` 함수는 특성별 정규화를 숫자 특성에 적용하는 레이어를 반환합니다.
```
def get_normalization_layer(name, dataset):
# Create a Normalization layer for our feature.
normalizer = preprocessing.Normalization()
# Prepare a Dataset that only yields our feature.
feature_ds = dataset.map(lambda x, y: x[name])
# Learn the statistics of the data.
normalizer.adapt(feature_ds)
return normalizer
photo_count_col = train_features['PhotoAmt']
layer = get_normalization_layer('PhotoAmt', train_ds)
layer(photo_count_col)
```
참고: 숫자 특성(수백 개 이상)이 많은 경우, 먼저 숫자 특성을 연결하고 단일 [normalization](https://www.tensorflow.org/api_docs/python/tf/keras/layers/experimental/preprocessing/Normalization) 레이어를 사용하는 것이 더 효율적입니다.
### 범주 열
이 데이터세트에서 Type은 문자열(예: 'Dog'또는 'Cat')으로 표시됩니다. 모델에 직접 문자열을 공급할 수 없습니다. 전처리 레이어는 문자열을 원-핫 벡터로 처리합니다.
`get_category_encoding_layer` 함수는 어휘의 값을 정수 인덱스로 매핑하고 특성을 원-핫 인코딩하는 레이어를 반환합니다.
```
def get_category_encoding_layer(name, dataset, dtype, max_tokens=None):
# Create a StringLookup layer which will turn strings into integer indices
if dtype == 'string':
index = preprocessing.StringLookup(max_tokens=max_tokens)
else:
index = preprocessing.IntegerLookup(max_values=max_tokens)
# Prepare a Dataset that only yields our feature
feature_ds = dataset.map(lambda x, y: x[name])
# Learn the set of possible values and assign them a fixed integer index.
index.adapt(feature_ds)
# Create a Discretization for our integer indices.
encoder = preprocessing.CategoryEncoding(max_tokens=index.vocab_size())
# Prepare a Dataset that only yields our feature.
feature_ds = feature_ds.map(index)
# Learn the space of possible indices.
encoder.adapt(feature_ds)
# Apply one-hot encoding to our indices. The lambda function captures the
# layer so we can use them, or include them in the functional model later.
return lambda feature: encoder(index(feature))
type_col = train_features['Type']
layer = get_category_encoding_layer('Type', train_ds, 'string')
layer(type_col)
```
종종 모델에 숫자를 직접 입력하지 않고 대신 해당 입력의 원-핫 인코딩을 사용합니다. 애완 동물의 나이를 나타내는 원시 데이터를 고려합니다.
```
type_col = train_features['Age']
category_encoding_layer = get_category_encoding_layer('Age', train_ds,
'int64', 5)
category_encoding_layer(type_col)
```
## 사용할 열 선택하기
여러 유형의 전처리 레이어를 사용하는 방법을 살펴보았습니다. 이제 레이어를 모델을 훈련하는 데 사용합니다. [Keras 함수형 API](https://www.tensorflow.org/guide/keras/functional)를 사용하여 모델을 빌드합니다. Keras 함수형 API는 [tf.keras.Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) API보다 더 유연한 모델을 생성하는 방법입니다.
이 튜토리얼의 목표는 전처리 레이어를 처리하는 데 필요한 전체 코드(예: 메커니즘)를 보여주는 것입니다. 모델을 훈련하기 위해 몇 개의 열이 임의로 선택되었습니다.
요점: 목표가 정확한 모델을 빌드하는 것이라면 자신의 더 큰 데이터세트를 시도하고 포함할 가장 의미 있는 특성과 표현 방법에 대해 신중하게 고려하세요.
이전에는 입력 파이프라인을 보여주기 위해 작은 배치 크기를 사용했습니다. 이제 더 큰 배치 크기로 새 입력 파이프라인을 생성해 보겠습니다.
```
batch_size = 256
train_ds = df_to_dataset(train, batch_size=batch_size)
val_ds = df_to_dataset(val, shuffle=False, batch_size=batch_size)
test_ds = df_to_dataset(test, shuffle=False, batch_size=batch_size)
all_inputs = []
encoded_features = []
# Numeric features.
for header in ['PhotoAmt', 'Fee']:
numeric_col = tf.keras.Input(shape=(1,), name=header)
normalization_layer = get_normalization_layer(header, train_ds)
encoded_numeric_col = normalization_layer(numeric_col)
all_inputs.append(numeric_col)
encoded_features.append(encoded_numeric_col)
# Categorical features encoded as integers.
age_col = tf.keras.Input(shape=(1,), name='Age', dtype='int64')
encoding_layer = get_category_encoding_layer('Age', train_ds, dtype='int64',
max_tokens=5)
encoded_age_col = encoding_layer(age_col)
all_inputs.append(age_col)
encoded_features.append(encoded_age_col)
# Categorical features encoded as string.
categorical_cols = ['Type', 'Color1', 'Color2', 'Gender', 'MaturitySize',
'FurLength', 'Vaccinated', 'Sterilized', 'Health', 'Breed1']
for header in categorical_cols:
categorical_col = tf.keras.Input(shape=(1,), name=header, dtype='string')
encoding_layer = get_category_encoding_layer(header, train_ds, dtype='string',
max_tokens=5)
encoded_categorical_col = encoding_layer(categorical_col)
all_inputs.append(categorical_col)
encoded_features.append(encoded_categorical_col)
```
## 모델 생성, 컴파일 및 훈련하기
이제 엔드 투 엔드 모델을 만들 수 있습니다.
```
all_features = tf.keras.layers.concatenate(encoded_features)
x = tf.keras.layers.Dense(32, activation="relu")(all_features)
x = tf.keras.layers.Dropout(0.5)(x)
output = tf.keras.layers.Dense(1)(x)
model = tf.keras.Model(all_inputs, output)
model.compile(optimizer='adam',
loss=tf.keras.losses.BinaryCrossentropy(from_logits=True),
metrics=["accuracy"])
```
연결 그래프를 시각화해 보겠습니다.
```
# rankdir='LR' is used to make the graph horizontal.
tf.keras.utils.plot_model(model, show_shapes=True, rankdir="LR")
```
### 모델 훈련하기
```
model.fit(train_ds, epochs=10, validation_data=val_ds)
loss, accuracy = model.evaluate(test_ds)
print("Accuracy", accuracy)
```
## 새로운 데이터로 추론하기
요점: 전처리 코드가 모델 자체에 포함되어 있기 때문에 여러분이 개발한 모델은 이제 CSV 파일에서 행을 직접 분류할 수 있습니다.
이제 Keras 모델을 저장하고 다시 로드할 수 있습니다. TensorFlow 모델에 대한 자세한 내용은 [여기](https://www.tensorflow.org/tutorials/keras/save_and_load)에서 튜토리어를 따르세요.
```
model.save('my_pet_classifier')
reloaded_model = tf.keras.models.load_model('my_pet_classifier')
```
새 샘플에 대한 예측값을 얻으려면, `model.predict()`를 호출하면 됩니다. 다음 두 가지만 수행해야 합니다.
1. 배치 차원을 갖도록 스칼라를 목록으로 래핑합니다(모델은 단일 샘플이 아닌 데이터 배치만 처리함).
2. 각 특성에 대해 `convert_to_tensor`를 호출합니다.
```
sample = {
'Type': 'Cat',
'Age': 3,
'Breed1': 'Tabby',
'Gender': 'Male',
'Color1': 'Black',
'Color2': 'White',
'MaturitySize': 'Small',
'FurLength': 'Short',
'Vaccinated': 'No',
'Sterilized': 'No',
'Health': 'Healthy',
'Fee': 100,
'PhotoAmt': 2,
}
input_dict = {name: tf.convert_to_tensor([value]) for name, value in sample.items()}
predictions = reloaded_model.predict(input_dict)
prob = tf.nn.sigmoid(predictions[0])
print(
"This particular pet had a %.1f percent probability "
"of getting adopted." % (100 * prob)
)
```
요점: 일반적으로 더 크고 복잡한 데이터세트를 사용한 딥 러닝을 통해 최상의 결과를 얻을 수 있습니다. 작은 데이터세트로 작업할 때는 의사 결정 트리 또는 랜덤 포레스트를 강력한 기준으로 사용하는 것이 좋습니다. 이 튜토리얼의 목표는 구조적 데이터를 처리하는 메커니즘을 보여주기 위한 것이므로 향후 자체 데이터세트를 처리할 때 시작점으로 사용할 수 있는 코드를 살펴보았습니다.
## 다음 단계
구조적 데이터의 분류에 대해 자세히 알아보는 가장 좋은 방법은 직접 시도해 보는 것입니다. 처리할 데이터세트를 찾고 위와 유사한 코드를 사용하여 분류하도록 모델을 훈련할 수 있습니다. 정확성을 높이려면 모델에 포함할 특성과 표현 방법을 신중하게 고려하세요.
| github_jupyter |
## NumPy
### The Scientific Python Trilogy
Why is Python so popular for research work?
MATLAB has typically been the most popular "language of technical computing", with strong built-in support for efficient numerical analysis with matrices (the *mat* in MATLAB is for Matrix, not Maths), and plotting.
Other dynamic languages have cleaner, more logical syntax (Ruby, Haskell)
But Python users developed three critical libraries, matching the power of MATLAB for scientific work:
* Matplotlib, the plotting library created by [John D. Hunter](https://en.wikipedia.org/wiki/John_D._Hunter)
* NumPy, a fast matrix maths library created by [Travis Oliphant](https://www.anaconda.com/people/travis-oliphant)
* IPython, the precursor of the notebook, created by [Fernando Perez](http://fperez.org)
By combining a plotting library, a matrix maths library, and an easy-to-use interface allowing live plotting commands
in a persistent environment, the powerful capabilities of MATLAB were matched by a free and open toolchain.
We've learned about Matplotlib and IPython in this course already. NumPy is the last part of the trilogy.
### Limitations of Python Lists
The normal Python list is just one dimensional. To make a matrix, we have to nest Python lists:
```
x = [list(range(5)) for N in range(5)]
x
x[2][2]
```
Applying an operation to every element is a pain:
```
x + 5
[[elem + 5 for elem in row] for row in x]
```
Common useful operations like transposing a matrix or reshaping a 10 by 10 matrix into a 20 by 5 matrix are not easy to code in raw Python lists.
### The NumPy array
NumPy's array type represents a multidimensional matrix $M_{i,j,k...n}$
The NumPy array seems at first to be just like a list. For example, we can index it and iterate over it:
```
import numpy as np
my_array = np.array(range(5))
my_array
my_array[2]
for element in my_array:
print("Hello" * element)
```
We can also see our first weakness of NumPy arrays versus Python lists:
```
my_array.append(4)
```
For NumPy arrays, you typically don't change the data size once you've defined your array,
whereas for Python lists, you can do this efficiently. However, you get back lots of goodies in return...
### Elementwise Operations
Most operations can be applied element-wise automatically!
```
my_array + 2
```
These "vectorized" operations are very fast: (the `%%timeit` magic reports how long it takes to run a cell; there is [more information](https://ipython.readthedocs.io/en/stable/interactive/magics.html#magic-timeit) available if interested)
```
import numpy as np
big_list = range(10000)
big_array = np.arange(10000)
%%timeit
[x**2 for x in big_list]
%%timeit
big_array**2
```
### arange and linspace
NumPy has two methods for quickly defining evenly-spaced arrays of (floating-point) numbers. These can be useful, for example, in plotting.
The first method is `arange`:
```
x = np.arange(0, 10, 0.1) # Start, stop, step size
```
This is similar to Python's `range`, although note that we can't use non-integer steps with the latter!
```
y = list(range(0, 10, 0.1))
```
The second method is `linspace`:
```
import math
values = np.linspace(0, math.pi, 100) # Start, stop, number of steps
values
```
Regardless of the method used, the array of values that we get can be used in the same way.
In fact, NumPy comes with "vectorised" versions of common functions which work element-by-element when applied to arrays:
```
%matplotlib inline
from matplotlib import pyplot as plt
plt.plot(values, np.sin(values))
```
So we don't have to use awkward list comprehensions when using these.
### Multi-Dimensional Arrays
NumPy's true power comes from multi-dimensional arrays:
```
np.zeros([3, 4, 2]) # 3 arrays with 4 rows and 2 columns each
```
Unlike a list-of-lists in Python, we can reshape arrays:
```
x = np.array(range(40))
x
y = x.reshape([4, 5, 2])
y
```
And index multiple columns at once:
```
y[3, 2, 1]
```
Including selecting on inner axes while taking all from the outermost:
```
y[:, 2, 1]
```
And subselecting ranges:
```
y[2:, :1, :] # Last 2 axes, 1st row, all columns
```
And [transpose](https://en.wikipedia.org/wiki/Transpose) arrays:
```
y.transpose()
```
You can get the dimensions of an array with `shape`:
```
y.shape
y.transpose().shape
```
Some numpy functions apply by default to the whole array, but can be chosen to act only on certain axes:
```
x = np.arange(12).reshape(4,3)
x
x.mean(1) # Mean along the second axis, leaving the first.
x.mean(0) # Mean along the first axis, leaving the second.
x.mean() # mean of all axes
```
### Array Datatypes
A Python `list` can contain data of mixed type:
```
x = ['hello', 2, 3.4]
type(x[2])
type(x[1])
```
A NumPy array always contains just one datatype:
```
np.array(x)
```
NumPy will choose the least-generic-possible datatype that can contain the data:
```
y = np.array([2, 3.4])
y
```
You can access the array's `dtype`, or check the type of individual elements:
```
y.dtype
type(y[0])
z = np.array([3, 4, 5])
z
type(z[0])
```
The results are, when you get to know them, fairly obvious string codes for datatypes:
NumPy supports all kinds of datatypes beyond the python basics.
NumPy will convert python type names to dtypes:
```
x = [2, 3.4, 7.2, 0]
int_array = np.array(x, dtype=int)
float_array = np.array(x, dtype=float)
int_array
float_array
int_array.dtype
float_array.dtype
```
### Broadcasting
This is another really powerful feature of NumPy.
By default, array operations are element-by-element:
```
np.arange(5) * np.arange(5)
```
If we multiply arrays with non-matching shapes we get an error:
```
np.arange(5) * np.arange(6)
np.zeros([2,3]) * np.zeros([2,4])
m1 = np.arange(100).reshape([10, 10])
m2 = np.arange(100).reshape([10, 5, 2])
m1 + m2
```
Arrays must match in all dimensions in order to be compatible:
```
np.ones([3, 3]) * np.ones([3, 3]) # Note elementwise multiply, *not* matrix multiply.
```
**Except**, that if one array has any Dimension 1, then the data is **REPEATED** to match the other.
```
col = np.arange(10).reshape([10, 1])
col
row = col.transpose()
row
col.shape # "Column Vector"
row.shape # "Row Vector"
row + col
10 * row + col
```
This works for arrays with more than one unit dimension.
### Newaxis
Broadcasting is very powerful, and numpy allows indexing with `np.newaxis` to temporarily create new one-long dimensions on the fly.
```
import numpy as np
x = np.arange(10).reshape(2, 5)
y = np.arange(8).reshape(2, 2, 2)
x
y
x[:, :, np.newaxis, np.newaxis].shape
y[:, np.newaxis, :, :].shape
res = x[:, :, np.newaxis, np.newaxis] * y[:, np.newaxis, :, :]
res.shape
np.sum(res)
```
Note that `newaxis` works because a $3 \times 1 \times 3$ array and a $3 \times 3$ array contain the same data,
differently shaped:
```
threebythree = np.arange(9).reshape(3, 3)
threebythree
threebythree[:, np.newaxis, :]
```
### Dot Products
NumPy multiply is element-by-element, not a dot-product:
```
a = np.arange(9).reshape(3, 3)
a
b = np.arange(3, 12).reshape(3, 3)
b
a * b
```
To get a dot-product, (matrix inner product) we can use a built in function:
```
np.dot(a, b)
```
Though it is possible to represent this in the algebra of broadcasting and newaxis:
```
a[:, :, np.newaxis].shape
b[np.newaxis, :, :].shape
a[:, :, np.newaxis] * b[np.newaxis, :, :]
(a[:, :, np.newaxis] * b[np.newaxis, :, :]).sum(1)
```
Or if you prefer:
```
(a.reshape(3, 3, 1) * b.reshape(1, 3, 3)).sum(1)
```
We use broadcasting to generate $A_{ij}B_{jk}$ as a 3-d matrix:
```
a.reshape(3, 3, 1) * b.reshape(1, 3, 3)
```
Then we sum over the middle, $j$ axis, [which is the 1-axis of three axes numbered (0,1,2)] of this 3-d matrix. Thus we generate $\Sigma_j A_{ij}B_{jk}$.
We can see that the broadcasting concept gives us a powerful and efficient way to express many linear algebra operations computationally.
### Record Arrays
These are a special array structure designed to match the CSV "Record and Field" model. It's a very different structure
from the normal NumPy array, and different fields *can* contain different datatypes. We saw this when we looked at CSV files:
```
x = np.arange(50).reshape([10, 5])
record_x = x.view(dtype={'names': ["col1", "col2", "another", "more", "last"],
'formats': [int]*5 })
record_x
```
Record arrays can be addressed with field names like they were a dictionary:
```
record_x['col1']
```
We've seen these already when we used NumPy's CSV parser.
### Logical arrays, masking, and selection
Numpy defines operators like == and < to apply to arrays *element by element*:
```
x = np.zeros([3, 4])
x
y = np.arange(-1, 2)[:, np.newaxis] * np.arange(-2, 2)[np.newaxis, :]
y
iszero = x == y
iszero
```
A logical array can be used to select elements from an array:
```
y[np.logical_not(iszero)]
```
Although when printed, this comes out as a flat list, if assigned to, the *selected elements of the array are changed!*
```
y[iszero] = 5
y
```
### Numpy memory
Numpy memory management can be tricksy:
```
x = np.arange(5)
y = x[:]
y[2] = 0
x
```
It does **not** behave like lists!
```
x = list(range(5))
y = x[:]
y[2] = 0
x
```
We must use `np.copy` to force separate memory. Otherwise NumPy tries its hardest to make slices be *views* on data.
Now, this has all been very theoretical, but let's go through a practical example, and see how powerful NumPy can be.
| github_jupyter |
# Anna KaRNNa
In this notebook, we'll build a character-wise RNN trained on Anna Karenina, one of my all-time favorite books. It'll be able to generate new text based on the text from the book.
This network is based off of Andrej Karpathy's [post on RNNs](http://karpathy.github.io/2015/05/21/rnn-effectiveness/) and [implementation in Torch](https://github.com/karpathy/char-rnn). Also, some information [here at r2rt](http://r2rt.com/recurrent-neural-networks-in-tensorflow-ii.html) and from [Sherjil Ozair](https://github.com/sherjilozair/char-rnn-tensorflow) on GitHub. Below is the general architecture of the character-wise RNN.
<img src="assets/charseq.jpeg" width="500">
```
import time
from collections import namedtuple
import numpy as np
import tensorflow as tf
```
First we'll load the text file and convert it into integers for our network to use. Here I'm creating a couple dictionaries to convert the characters to and from integers. Encoding the characters as integers makes it easier to use as input in the network.
```
with open('anna.txt', 'r') as f:
text=f.read()
vocab = sorted(set(text))
vocab_to_int = {c: i for i, c in enumerate(vocab)}
int_to_vocab = dict(enumerate(vocab))
encoded = np.array([vocab_to_int[c] for c in text], dtype=np.int32)
```
Let's check out the first 100 characters, make sure everything is peachy. According to the [American Book Review](http://americanbookreview.org/100bestlines.asp), this is the 6th best first line of a book ever.
```
text[:100]
```
And we can see the characters encoded as integers.
```
encoded[:100]
```
Since the network is working with individual characters, it's similar to a classification problem in which we are trying to predict the next character from the previous text. Here's how many 'classes' our network has to pick from.
```
len(vocab)
```
## Making training mini-batches
Here is where we'll make our mini-batches for training. Remember that we want our batches to be multiple sequences of some desired number of sequence steps. Considering a simple example, our batches would look like this:
<img src="assets/sequence_batching@1x.png" width=500px>
<br>
We start with our text encoded as integers in one long array in `encoded`. Let's create a function that will give us an iterator for our batches. I like using [generator functions](https://jeffknupp.com/blog/2013/04/07/improve-your-python-yield-and-generators-explained/) to do this. Then we can pass `encoded` into this function and get our batch generator.
The first thing we need to do is discard some of the text so we only have completely full batches. Each batch contains $N \times M$ characters, where $N$ is the batch size (the number of sequences) and $M$ is the number of steps. Then, to get the total number of batches, $K$, we can make from the array `arr`, you divide the length of `arr` by the number of characters per batch. Once you know the number of batches, you can get the total number of characters to keep from `arr`, $N * M * K$.
After that, we need to split `arr` into $N$ sequences. You can do this using `arr.reshape(size)` where `size` is a tuple containing the dimensions sizes of the reshaped array. We know we want $N$ sequences (`batch_size` below), let's make that the size of the first dimension. For the second dimension, you can use `-1` as a placeholder in the size, it'll fill up the array with the appropriate data for you. After this, you should have an array that is $N \times (M * K)$.
Now that we have this array, we can iterate through it to get our batches. The idea is each batch is a $N \times M$ window on the $N \times (M * K)$ array. For each subsequent batch, the window moves over by `n_steps`. We also want to create both the input and target arrays. Remember that the targets are the inputs shifted over one character.
The way I like to do this window is use `range` to take steps of size `n_steps` from $0$ to `arr.shape[1]`, the total number of steps in each sequence. That way, the integers you get from `range` always point to the start of a batch, and each window is `n_steps` wide.
> **Exercise:** Write the code for creating batches in the function below. The exercises in this notebook _will not be easy_. I've provided a notebook with solutions alongside this notebook. If you get stuck, checkout the solutions. The most important thing is that you don't copy and paste the code into here, **type out the solution code yourself.**
```
def get_batches(arr, batch_size, n_steps):
'''Create a generator that returns batches of size
batch_size x n_steps from arr.
Arguments
---------
arr: Array you want to make batches from
batch_size: Batch size, the number of sequences per batch
n_steps: Number of sequence steps per batch
'''
# Get the number of characters per batch and number of batches we can make
characters_per_batch = batch_size * n_steps
n_batches = len(arr) // characters_per_batch
# Keep only enough characters to make full batches
arr = arr[:n_batches * characters_per_batch]
# Reshape into batch_size rows
arr = arr.reshape((batch_size, -1))
for n in range(0, arr.shape[1], n_steps):
# The features
x = arr[:,n:n+n_steps]
# The targets, shifted by one
y = np.zeros(x.shape, dtype=x.dtype)
y_tmp = arr[:,n+1:n+n_steps+1]
y[:,:y_tmp.shape[1]] = y_tmp
yield x, y
```
Now I'll make my data sets and we can check out what's going on here. Here I'm going to use a batch size of 10 and 50 sequence steps.
```
batches = get_batches(encoded, 10, 50)
x, y = next(batches)
print('x\n', x[:10, :10])
print('\ny\n', y[:10, :10])
```
If you implemented `get_batches` correctly, the above output should look something like
```
x
[[55 63 69 22 6 76 45 5 16 35]
[ 5 69 1 5 12 52 6 5 56 52]
[48 29 12 61 35 35 8 64 76 78]
[12 5 24 39 45 29 12 56 5 63]
[ 5 29 6 5 29 78 28 5 78 29]
[ 5 13 6 5 36 69 78 35 52 12]
[63 76 12 5 18 52 1 76 5 58]
[34 5 73 39 6 5 12 52 36 5]
[ 6 5 29 78 12 79 6 61 5 59]
[ 5 78 69 29 24 5 6 52 5 63]]
y
[[63 69 22 6 76 45 5 16 35 35]
[69 1 5 12 52 6 5 56 52 29]
[29 12 61 35 35 8 64 76 78 28]
[ 5 24 39 45 29 12 56 5 63 29]
[29 6 5 29 78 28 5 78 29 45]
[13 6 5 36 69 78 35 52 12 43]
[76 12 5 18 52 1 76 5 58 52]
[ 5 73 39 6 5 12 52 36 5 78]
[ 5 29 78 12 79 6 61 5 59 63]
[78 69 29 24 5 6 52 5 63 76]]
```
although the exact numbers will be different. Check to make sure the data is shifted over one step for `y`.
## Building the model
Below is where you'll build the network. We'll break it up into parts so it's easier to reason about each bit. Then we can connect them up into the whole network.
<img src="assets/charRNN.png" width=500px>
### Inputs
First off we'll create our input placeholders. As usual we need placeholders for the training data and the targets. We'll also create a placeholder for dropout layers called `keep_prob`. This will be a scalar, that is a 0-D tensor. To make a scalar, you create a placeholder without giving it a size.
> **Exercise:** Create the input placeholders in the function below.
```
def build_inputs(batch_size, num_steps):
''' Define placeholders for inputs, targets, and dropout
Arguments
---------
batch_size: Batch size, number of sequences per batch
num_steps: Number of sequence steps in a batch
'''
# Declare placeholders we'll feed into the graph
inputs = tf.placeholder(dtype=tf.int32,shape=[batch_size, num_steps], name = 'inputs')
targets = tf.placeholder(dtype=tf.int32, shape =[batch_size, num_steps], name = 'targets')
# Keep probability placeholder for drop out layers
keep_prob = tf.placeholder(tf.float32)
return inputs, targets, keep_prob
```
### LSTM Cell
Here we will create the LSTM cell we'll use in the hidden layer. We'll use this cell as a building block for the RNN. So we aren't actually defining the RNN here, just the type of cell we'll use in the hidden layer.
We first create a basic LSTM cell with
```python
lstm = tf.contrib.rnn.BasicLSTMCell(num_units)
```
where `num_units` is the number of units in the hidden layers in the cell. Then we can add dropout by wrapping it with
```python
tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
```
You pass in a cell and it will automatically add dropout to the inputs or outputs. Finally, we can stack up the LSTM cells into layers with [`tf.contrib.rnn.MultiRNNCell`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/contrib/rnn/MultiRNNCell). With this, you pass in a list of cells and it will send the output of one cell into the next cell. Previously with TensorFlow 1.0, you could do this
```python
tf.contrib.rnn.MultiRNNCell([cell]*num_layers)
```
This might look a little weird if you know Python well because this will create a list of the same `cell` object. However, TensorFlow 1.0 will create different weight matrices for all `cell` objects. But, starting with TensorFlow 1.1 you actually need to create new cell objects in the list. To get it to work in TensorFlow 1.1, it should look like
```python
def build_cell(num_units, keep_prob):
lstm = tf.contrib.rnn.BasicLSTMCell(num_units)
drop = tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
return drop
tf.contrib.rnn.MultiRNNCell([build_cell(num_units, keep_prob) for _ in range(num_layers)])
```
Even though this is actually multiple LSTM cells stacked on each other, you can treat the multiple layers as one cell.
We also need to create an initial cell state of all zeros. This can be done like so
```python
initial_state = cell.zero_state(batch_size, tf.float32)
```
Below, we implement the `build_lstm` function to create these LSTM cells and the initial state.
```
def build_lstm(lstm_size, num_layers, batch_size, keep_prob):
''' Build LSTM cell.
Arguments
---------
keep_prob: Scalar tensor (tf.placeholder) for the dropout keep probability
lstm_size: Size of the hidden layers in the LSTM cells
num_layers: Number of LSTM layers
batch_size: Batch size
'''
### Build the LSTM Cell
# Use a basic LSTM cell
def build_cell (lstm_size, keep_prob):
# Use a basic LSTM cell
lstm = tf.contrib.rnn.BasicLSTMCell(lstm_size)
# Add dropout to the cell
drop = tf.contrib.rnn.DropoutWrapper(lstm, output_keep_prob=keep_prob)
return drop
# Stack up multiple LSTM layers, for deep learning
cell = tf.contrib.rnn.MultiRNNCell([build_cell(lstm_size, keep_prob) for _ in range(num_layers)])
initial_state = cell.zero_state(batch_size, tf.float32)
return cell, initial_state
```
### RNN Output
Here we'll create the output layer. We need to connect the output of the RNN cells to a full connected layer with a softmax output. The softmax output gives us a probability distribution we can use to predict the next character, so we want this layer to have size $C$, the number of classes/characters we have in our text.
If our input has batch size $N$, number of steps $M$, and the hidden layer has $L$ hidden units, then the output is a 3D tensor with size $N \times M \times L$. The output of each LSTM cell has size $L$, we have $M$ of them, one for each sequence step, and we have $N$ sequences. So the total size is $N \times M \times L$.
We are using the same fully connected layer, the same weights, for each of the outputs. Then, to make things easier, we should reshape the outputs into a 2D tensor with shape $(M * N) \times L$. That is, one row for each sequence and step, where the values of each row are the output from the LSTM cells. We get the LSTM output as a list, `lstm_output`. First we need to concatenate this whole list into one array with [`tf.concat`](https://www.tensorflow.org/api_docs/python/tf/concat). Then, reshape it (with `tf.reshape`) to size $(M * N) \times L$.
One we have the outputs reshaped, we can do the matrix multiplication with the weights. We need to wrap the weight and bias variables in a variable scope with `tf.variable_scope(scope_name)` because there are weights being created in the LSTM cells. TensorFlow will throw an error if the weights created here have the same names as the weights created in the LSTM cells, which they will be default. To avoid this, we wrap the variables in a variable scope so we can give them unique names.
> **Exercise:** Implement the output layer in the function below.
```
def build_output(lstm_output, lstm_size, out_size):
''' Build a softmax layer, return the softmax output and logits.
Arguments
---------
lstm_output: List of output tensors from the LSTM layer
lstm_size (in_size): Size of the input tensor, for example, size of the LSTM cells
out_size: Size of this softmax layer
'''
# Reshape output so it's a bunch of rows, one row for each step for each sequence.
# NOTE: LSTM output returns a list of outputs for each output so we need to concatenate them to form an array of M x N X L
# Concatenate lstm_output over axis 1 (the columns)
print('LSTM OUTPUT SIZE IS' , lstm_output.get_shape())
seq_output = tf.concat(lstm_output, axis=1)
# Reshape seq_output to a 2D tensor with lstm_size columns
x = tf.reshape(seq_output,[-1,lstm_size])
# Connect the RNN outputs to a softmax layer
with tf.variable_scope('softmax'):
# Create the weight and bias variables here
softmax_w = tf.Variable(tf.truncated_normal((lstm_size, out_size), stddev=0.1))
softmax_b = tf.Variable(tf.zeros(out_size))
# Since output is a bunch of rows of RNN cell outputs, logits will be a bunch
# of rows of logit outputs, one for each step and sequence
logits = tf.matmul(x, softmax_w) + softmax_b
# Use softmax to get the probabilities for predicted characters
out = tf.nn.softmax(logits, name='predictions')
return out, logits
```
### Training loss
Next up is the training loss. We get the logits and targets and calculate the softmax cross-entropy loss. First we need to one-hot encode the targets, we're getting them as encoded characters. Then, reshape the one-hot targets so it's a 2D tensor with size $(M*N) \times C$ where $C$ is the number of classes/characters we have. Remember that we reshaped the LSTM outputs and ran them through a fully connected layer with $C$ units. So our logits will also have size $(M*N) \times C$.
Then we run the logits and targets through `tf.nn.softmax_cross_entropy_with_logits` and find the mean to get the loss.
>**Exercise:** Implement the loss calculation in the function below.
```
def build_loss(logits, targets, lstm_size, num_classes):
''' Calculate the loss from the logits and the targets.
Arguments
---------
logits: Logits from final fully connected layer
targets: Targets for supervised learning
lstm_size: Number of LSTM hidden units
num_classes: Number of classes in targets
'''
# One-hot encode targets and reshape to match logits, one row per sequence per step
y_one_hot = tf.one_hot(targets, depth=num_classes) # shape: shape_of_targets X num_classes =
# [batch_size, num_steps, num_classes ]
print("shape of y_one_hot before reshape: " , y_one_hot.get_shape())
y_reshaped =tf.reshape(y_one_hot, logits.get_shape()) # shape: M*N x num_classes
# Softmax cross entropy loss
loss = tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=y_reshaped)
loss = tf.reduce_mean(loss)
return loss
```
### Optimizer
Here we build the optimizer. Normal RNNs have have issues gradients exploding and disappearing. LSTMs fix the disappearance problem, but the gradients can still grow without bound. To fix this, we can clip the gradients above some threshold. That is, if a gradient is larger than that threshold, we set it to the threshold. This will ensure the gradients never grow overly large. Then we use an AdamOptimizer for the learning step.
```
def build_optimizer(loss, learning_rate, grad_clip):
''' Build optmizer for training, using gradient clipping.
Arguments:
loss: Network loss
learning_rate: Learning rate for optimizer
'''
# Optimizer for training, using gradient clipping to control exploding gradients
tvars = tf.trainable_variables()
#
grads, _ = tf.clip_by_global_norm(tf.gradients(loss, tvars), grad_clip)
train_op = tf.train.AdamOptimizer(learning_rate)
optimizer = train_op.apply_gradients(zip(grads, tvars))
# NOTE: Calling minimize() takes care of both computing the gradients and applying them to the variables.
#If you want to process the gradients before applying them you can instead use the optimizer in three steps:
# Compute the gradients with compute_gradients() OR tf.gradients(loss, train_vars).
# Process the gradients as you wish.
# Apply the processed gradients with apply_gradients().
return optimizer
```
### Build the network
Now we can put all the pieces together and build a class for the network. To actually run data through the LSTM cells, we will use [`tf.nn.dynamic_rnn`](https://www.tensorflow.org/versions/r1.0/api_docs/python/tf/nn/dynamic_rnn). This function will pass the hidden and cell states across LSTM cells appropriately for us. It returns the outputs for each LSTM cell at each step for each sequence in the mini-batch. It also gives us the final LSTM state. We want to save this state as `final_state` so we can pass it to the first LSTM cell in the the next mini-batch run. For `tf.nn.dynamic_rnn`, we pass in the cell and initial state we get from `build_lstm`, as well as our input sequences. Also, we need to one-hot encode the inputs before going into the RNN.
> **Exercise:** Use the functions you've implemented previously and `tf.nn.dynamic_rnn` to build the network.
```
class CharRNN:
def __init__(self, num_classes, batch_size=64, num_steps=50,
lstm_size=128, num_layers=2, learning_rate=0.001,
grad_clip=5, sampling=False):
# When we're using this network for sampling later, we'll be passing in
# one character at a time, so providing an option for that
if sampling == True:
batch_size, num_steps = 1, 1
else:
batch_size, num_steps = batch_size, num_steps
tf.reset_default_graph()
# Build the input placeholder tensors
self.inputs, self.targets, self.keep_prob = build_inputs(batch_size, num_steps)
# Build the LSTM cell
cell, self.initial_state = build_lstm(lstm_size, num_layers, batch_size, keep_prob)
### Run the data through the RNN layers
# First, one-hot encode the input tokens
# our features are one hot encoded characters
x_one_hot = tf.one_hot(self.inputs, num_classes)
print("shape of one hot encoded inputs: ", x_one_hot.get_shape())
# NOTE: we do not reshape x_one_hot, as we input them into dynamic rnn cell
# Run each sequence step through the RNN with tf.nn.dynamic_rnn
# 'outputs' is a tensor of shape [batch_size, max_time = num_steps, cell_state_size = lstm_size]
# 'state' is a tensor of shape [batch_size, cell_state_size] for a single cell
# 'state' is a N-tuple where N is the number of LSTMCells containing a
# tf.contrib.rnn.LSTMStateTuple for each cell
outputs, state = tf.nn.dynamic_rnn(cell, x_one_hot, initial_state=self.initial_state)
print("shape of the outputs is: ", outputs.get_shape())
print("type of the state is: ", type(state))
self.final_state = state
# Get softmax predictions and logits
self.prediction, self.logits = build_output(outputs, lstm_size, num_classes)
# Loss and optimizer (with gradient clipping)
self.loss = build_loss(self.logits, self.targets, lstm_size, num_classes)
self.optimizer = build_optimizer(self.loss, learning_rate, grad_clip)
```
## Hyperparameters
Here are the hyperparameters for the network.
* `batch_size` - Number of sequences running through the network in one pass.
* `num_steps` - Number of characters in the sequence the network is trained on. Larger is better typically, the network will learn more long range dependencies. But it takes longer to train. 100 is typically a good number here.
* `lstm_size` - The number of units in the hidden layers.
* `num_layers` - Number of hidden LSTM layers to use
* `learning_rate` - Learning rate for training
* `keep_prob` - The dropout keep probability when training. If you're network is overfitting, try decreasing this.
Here's some good advice from Andrej Karpathy on training the network. I'm going to copy it in here for your benefit, but also link to [where it originally came from](https://github.com/karpathy/char-rnn#tips-and-tricks).
> ## Tips and Tricks
>### Monitoring Validation Loss vs. Training Loss
>If you're somewhat new to Machine Learning or Neural Networks it can take a bit of expertise to get good models. The most important quantity to keep track of is the difference between your training loss (printed during training) and the validation loss (printed once in a while when the RNN is run on the validation data (by default every 1000 iterations)). In particular:
> - If your training loss is much lower than validation loss then this means the network might be **overfitting**. Solutions to this are to decrease your network size, or to increase dropout. For example you could try dropout of 0.5 and so on.
> - If your training/validation loss are about equal then your model is **underfitting**. Increase the size of your model (either number of layers or the raw number of neurons per layer)
> ### Approximate number of parameters
> The two most important parameters that control the model are `lstm_size` and `num_layers`. I would advise that you always use `num_layers` of either 2/3. The `lstm_size` can be adjusted based on how much data you have. The two important quantities to keep track of here are:
> - The number of parameters in your model. This is printed when you start training.
> - The size of your dataset. 1MB file is approximately 1 million characters.
>These two should be about the same order of magnitude. It's a little tricky to tell. Here are some examples:
> - I have a 100MB dataset and I'm using the default parameter settings (which currently print 150K parameters). My data size is significantly larger (100 mil >> 0.15 mil), so I expect to heavily underfit. I am thinking I can comfortably afford to make `lstm_size` larger.
> - I have a 10MB dataset and running a 10 million parameter model. I'm slightly nervous and I'm carefully monitoring my validation loss. If it's larger than my training loss then I may want to try to increase dropout a bit and see if that helps the validation loss.
> ### Best models strategy
>The winning strategy to obtaining very good models (if you have the compute time) is to always err on making the network larger (as large as you're willing to wait for it to compute) and then try different dropout values (between 0,1). Whatever model has the best validation performance (the loss, written in the checkpoint filename, low is good) is the one you should use in the end.
>It is very common in deep learning to run many different models with many different hyperparameter settings, and in the end take whatever checkpoint gave the best validation performance.
>By the way, the size of your training and validation splits are also parameters. Make sure you have a decent amount of data in your validation set or otherwise the validation performance will be noisy and not very informative.
```
batch_size = 100 # Sequences per batch
num_steps = 100 # Number of sequence steps per batch
lstm_size = 512 # Size of hidden layers in LSTMs
num_layers = 2 # Number of LSTM layers
learning_rate = 0.001 # Learning rate
keep_prob = 0.5 # Dropout keep probability
print("number of steps per epoch ~ {}".format( 1985223/batch_size/num_steps))
```
## Time for training
This is typical training code, passing inputs and targets into the network, then running the optimizer. Here we also get back the final LSTM state for the mini-batch. Then, we pass that state back into the network so the next batch can continue the state from the previous batch. And every so often (set by `save_every_n`) I save a checkpoint.
Here I'm saving checkpoints with the format
`i{iteration number}_l{# hidden layer units}.ckpt`
> **Exercise:** Set the hyperparameters above to train the network. Watch the training loss, it should be consistently dropping. Also, I highly advise running this on a GPU.
```
epochs = 20
# Print losses every N interations
print_every_n = 50
# Save every N iterations
save_every_n = 200
model = CharRNN(len(vocab), batch_size=batch_size, num_steps=num_steps,
lstm_size=lstm_size, num_layers=num_layers,
learning_rate=learning_rate)
saver = tf.train.Saver()
saver = tf.train.Saver(max_to_keep=100)
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
# Use the line below to load a checkpoint and resume training
#saver.restore(sess, 'checkpoints/______.ckpt')
counter = 0
for e in range(epochs):
# Train network
new_state = sess.run(model.initial_state)
loss = 0
for x, y in get_batches(encoded, batch_size, num_steps):
counter += 1
start = time.time()
feed = {model.inputs: x,
model.targets: y,
model.keep_prob: keep_prob,
model.initial_state: new_state}
batch_loss, new_state, _ = sess.run([model.loss,
model.final_state,
model.optimizer],
feed_dict=feed)
if (counter % print_every_n == 0):
end = time.time()
print('Epoch: {}/{}... '.format(e+1, epochs),
'Training Step: {}... '.format(counter),
'Training loss: {:.4f}... '.format(batch_loss),
'{:.4f} sec/batch'.format((end-start)))
if (counter % save_every_n == 0):
saver.save(sess, "checkpoints/i{}_l{}.ckpt".format(counter, lstm_size))
saver.save(sess, "checkpoints/i{}_l{}.ckpt".format(counter, lstm_size))
```
#### Saved checkpoints
Read up on saving and loading checkpoints here: https://www.tensorflow.org/programmers_guide/variables
```
tf.train.get_checkpoint_state('checkpoints')
```
## Sampling
Now that the network is trained, we'll can use it to generate new text. The idea is that we pass in a character, then the network will predict the next character. We can use the new one, to predict the next one. And we keep doing this to generate all new text. I also included some functionality to prime the network with some text by passing in a string and building up a state from that.
The network gives us predictions for each character. To reduce noise and make things a little less random, I'm going to only choose a new character from the top N most likely characters.
```
# fcn to sample from n most likely chars
def pick_top_n(preds, vocab_size, top_n=5):
p = np.squeeze(preds) # probabilities of each char
p[np.argsort(p)[:-top_n]] = 0 # set the probs for everything other than top_n to zero
p = p / np.sum(p) # normalize
c = np.random.choice(vocab_size, 1, p=p)[0] # choose a number out of vocab_size with prob=p
return c
def sample(checkpoint, n_samples, lstm_size, vocab_size, prime="The "):
'''
prime: to prime the network with some text by passing in a string and
building up a state from that (not starting with 0 initial state).
'''
samples = [c for c in prime]
model = CharRNN(len(vocab), lstm_size=lstm_size, sampling=True) # note sampling=True,
#sets batch_size=1, num_steps=1
saver = tf.train.Saver()
with tf.Session() as sess:
saver.restore(sess, checkpoint) # note: restore must be called inside a session
new_state = sess.run(model.initial_state)
# run through rnn with the prime and generate the first character
for c in prime:
x = np.zeros((1, 1)) # batch_size=1, num_steps=1
x[0,0] = vocab_to_int[c]
feed = {model.inputs: x,
model.keep_prob: 1.,
model.initial_state: new_state}
preds, new_state = sess.run([model.prediction, model.final_state],
feed_dict=feed)
c = pick_top_n(preds, len(vocab))
samples.append(int_to_vocab[c])
for i in range(n_samples):
x[0,0] = c
feed = {model.inputs: x,
model.keep_prob: 1.,
model.initial_state: new_state}
preds, new_state = sess.run([model.prediction, model.final_state],
feed_dict=feed)
c = pick_top_n(preds, len(vocab))
samples.append(int_to_vocab[c])
return ''.join(samples)
```
Here, pass in the path to a checkpoint and sample from the network.
```
tf.train.latest_checkpoint('checkpoints')
# Sample 2000 chars from the latest checkpoint
checkpoint = tf.train.latest_checkpoint('checkpoints')
samp = sample(checkpoint, 2000, lstm_size, len(vocab), prime="Far")
print(samp)
checkpoint = 'checkpoints/i200_l512.ckpt'
samp = sample(checkpoint, 1000, lstm_size, len(vocab), prime="Far")
print(samp)
# Sample 1000 chars from the checkpoint i600
checkpoint = 'checkpoints/i600_l512.ckpt'
samp = sample(checkpoint, 1000, lstm_size, len(vocab), prime="Far")
print(samp)
# Sample 1000 chars from the checkpoint i1200
checkpoint = 'checkpoints/i1200_l512.ckpt'
samp = sample(checkpoint, 1000, lstm_size, len(vocab), prime="Far")
print(samp)
```
| github_jupyter |
```
!pip3 install opacus
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader, random_split
from sklearn import preprocessing
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import accuracy_score
batch_size = 50
class ChurnDataset(Dataset):
def __init__(self, csv_file):
df = pd.read_csv(csv_file)
df = df.drop(["Surname", "CustomerId", "RowNumber"], axis=1)
# Grouping variable names
self.categorical = ["Geography", "Gender"]
self.target = "Exited"
# One-hot encoding of categorical variables
self.churn_frame = pd.get_dummies(df, prefix=self.categorical)
# Save target and predictors
self.X = self.churn_frame.drop(self.target, axis=1)
self.y = self.churn_frame["Exited"]
scaler = StandardScaler()
X_array = scaler.fit_transform(self.X)
self.X = pd.DataFrame(X_array)
def __len__(self):
return len(self.churn_frame)
def __getitem__(self, idx):
# Convert idx from tensor to list due to pandas bug (that arises when using pytorch's random_split)
if isinstance(idx, torch.Tensor):
idx = idx.tolist()
return [self.X.iloc[idx].values, self.y[idx]]
def get_CHURN_model():
model = nn.Sequential(nn.Linear(13, 64),
nn.ReLU(),
nn.Linear(64, 64),
nn.ReLU(),
nn.Linear(64, 1))
return model
def get_dataloader(csv_file, batch_size):
# Load dataset
dataset = ChurnDataset(csv_file)
# Split into training and test
train_size = int(0.8 * len(dataset))
test_size = len(dataset) - train_size
trainset, testset = random_split(dataset, [train_size, test_size])
trainloader = DataLoader(trainset, batch_size=batch_size, shuffle=True)
testloader = DataLoader(testset, batch_size=batch_size, shuffle=True)
return trainloader, testloader, trainset, testset
def train(trainloader, net, optimizer, n_epochs=100):
device = "cpu"
# Define the model
#net = get_CHURN_model()
net = net.to(device)
#criterion = nn.CrossEntropyLoss()
criterion = nn.BCEWithLogitsLoss()
# Train the net
loss_per_iter = []
loss_per_batch = []
for epoch in range(n_epochs):
running_loss = 0.0
for inputs, labels in trainloader:
inputs = inputs.to(device)
labels = labels.to(device)
# Zero the parameter gradients
optimizer.zero_grad()
# Forward + backward + optimize
outputs = net(inputs.float())
loss = criterion(outputs, labels.float().unsqueeze(1))
loss.backward()
optimizer.step()
# Save loss to plot
running_loss += loss.item()
loss_per_iter.append(loss.item())
print("Epoch {} - Training loss: {}".format(epoch, running_loss/len(trainloader)))
running_loss = 0.0
return net
csv_file = "data/churn.csv"
trainloader, testloader, train_ds, test_ds = get_dataloader(csv_file, batch_size)
net = get_CHURN_model()
optimizer = optim.Adam(net.parameters(), weight_decay=0.0001, lr=0.003)
model = train(trainloader, net, optimizer, 50)
max_per_sample_grad_norm = 1.5
sample_rate = batch_size/len(train_ds)
noise_multiplier = 0.8
from opacus import PrivacyEngine
net = get_CHURN_model()
optimizer = optim.Adam(net.parameters(), weight_decay=0.0001, lr=0.003)
privacy_engine = PrivacyEngine(
net,
max_grad_norm=max_per_sample_grad_norm,
noise_multiplier = noise_multiplier,
sample_rate = sample_rate,
)
privacy_engine.attach(optimizer)
model = train(trainloader, net, optimizer, batch_size)
epsilon, best_alpha = privacy_engine.get_privacy_spent()
print (f" ε = {epsilon:.2f}, δ = {privacy_engine.target_delta}")
```
| github_jupyter |
```
import numpy as np
from tensorflow.keras import layers, models, optimizers, metrics
from matplotlib import pyplot as plt
import pandas as pd
import tensorflow as tf
normal = pd.read_csv("../ecg/ptbdb_normal.csv")
abnormal = pd.read_csv("../ecg/ptbdb_abnormal.csv")
normal = normal.values[:,:-1]
abnormal = abnormal.values[:,:-1]
normal = normal - 0.5
abnormal = abnormal - 0.5
def show_ecg(signal):
plt.figure(figsize=(12, 1))
plt.plot(np.arange(signal.size), signal, c="black")
plt.show()
show_ecg(normal[10])
show_ecg(abnormal[10])
normal.shape, abnormal.shape
X = np.concatenate((normal, abnormal))
y = np.concatenate((np.zeros(normal.shape[0]), np.ones(abnormal.shape[0])))
X = np.pad(X, ((0, 0), (0, 5)), mode='edge')
X = np.expand_dims(X, axis=2)
X.shape, y.shape
p = np.random.permutation(y.size - 1)
X = X[p]
y = y[p]
from __future__ import print_function, division
from tensorflow.keras.datasets import mnist
from tensorflow.keras.layers import Input, Dense, Reshape, Flatten, Dropout, multiply
from tensorflow.keras.layers import BatchNormalization, Activation, Embedding, ZeroPadding1D
from tensorflow.keras.layers import LeakyReLU
from tensorflow.keras.layers import UpSampling1D, Conv1D
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.optimizers import Adam
import matplotlib.pyplot as plt
import numpy as np
class ACGAN():
def __init__(self):
# Input shape
self.img_rows = 192
self.channels = 1
self.img_shape = (self.img_rows, self.channels)
self.num_classes = 2
self.latent_dim = 100
optimizer = Adam(0.0002, 0.5)
losses = ['binary_crossentropy', 'sparse_categorical_crossentropy']
# Build and compile the discriminator
self.discriminator = self.build_discriminator()
self.discriminator.compile(loss=losses,
optimizer=optimizer,
metrics=['accuracy'])
# Build the generator
self.generator = self.build_generator()
# The generator takes noise and the target label as input
# and generates the corresponding digit of that label
noise = Input(shape=(self.latent_dim,))
label = Input(shape=(1,))
img = self.generator([noise, label])
# For the combined model we will only train the generator
self.discriminator.trainable = False
# The discriminator takes generated image as input and determines validity
# and the label of that image
valid, target_label = self.discriminator(img)
# The combined model (stacked generator and discriminator)
# Trains the generator to fool the discriminator
self.combined = Model([noise, label], [valid, target_label])
self.combined.compile(loss=losses,
optimizer=optimizer)
def build_generator(self):
model = Sequential()
model.add(Dense(8 * 24, activation="relu", input_dim=self.latent_dim))
model.add(Reshape((16, 12)))
model.add(BatchNormalization(momentum=0.8))
model.add(UpSampling1D(size=3))
model.add(Conv1D(256, kernel_size=5, padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(momentum=0.8))
model.add(UpSampling1D(size=2))
model.add(Conv1D(128, kernel_size=5, padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(momentum=0.8))
model.add(UpSampling1D(size=2))
model.add(Conv1D(64, kernel_size=5, padding="same"))
model.add(Activation("relu"))
model.add(BatchNormalization(momentum=0.8))
model.add(Conv1D(self.channels, kernel_size=3, padding='same'))
model.add(Activation("tanh"))
model.summary()
noise = Input(shape=(self.latent_dim,))
label = Input(shape=(1,), dtype='int32')
label_embedding = Flatten()(Embedding(self.num_classes, self.latent_dim)(label))
model_input = multiply([noise, label_embedding])
img = model(model_input)
return Model([noise, label], img)
def build_discriminator(self):
model = Sequential()
model.add(Conv1D(16, kernel_size=7, strides=2, input_shape=self.img_shape, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Conv1D(32, kernel_size=7, strides=2, padding="same"))
model.add(ZeroPadding1D(padding=(0, 1)))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(BatchNormalization(momentum=0.8))
model.add(Conv1D(64, kernel_size=7, strides=2, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(BatchNormalization(momentum=0.8))
model.add(Conv1D(128, kernel_size=7, strides=1, padding="same"))
model.add(LeakyReLU(alpha=0.2))
model.add(Dropout(0.25))
model.add(Flatten())
model.summary()
img = Input(shape=self.img_shape)
# Extract feature representation
features = model(img)
# Determine validity and label of the image
validity = Dense(1, activation="sigmoid")(features)
label = Dense(self.num_classes, activation="softmax")(features)
return Model(img, [validity, label])
def train(self, epochs, batch_size=128, sample_interval=50):
# Load the dataset
(X_train, y_train) = (X, y)
# Configure inputs
y_train = y_train.reshape(-1, 1)
# Adversarial ground truths
valid = np.ones((batch_size, 1))
fake = np.zeros((batch_size, 1))
for epoch in range(epochs):
# ---------------------
# Train Discriminator
# ---------------------
# Select a random batch of images
idx = np.random.randint(0, X_train.shape[0], batch_size)
imgs = X_train[idx]
# Sample noise as generator input
noise = np.random.normal(0, 1, (batch_size, self.latent_dim))
# The labels of the digits that the generator tries to create an
# image representation of
sampled_labels = np.random.randint(0, 2, (batch_size, 1))
# Generate a half batch of new images
gen_imgs = self.generator.predict([noise, sampled_labels])
# Image labels. 0-9
img_labels = y_train[idx]
# Train the discriminator
d_loss_real = self.discriminator.train_on_batch(imgs, [valid, img_labels])
d_loss_fake = self.discriminator.train_on_batch(gen_imgs, [fake, sampled_labels])
d_loss = 0.5 * np.add(d_loss_real, d_loss_fake)
# ---------------------
# Train Generator
# ---------------------
# Train the generator
g_loss = self.combined.train_on_batch([noise, sampled_labels], [valid, sampled_labels])
# Plot the progress
if epoch % (sample_interval // 5) == 0:
print ("%d [D loss: %f, acc.: %.2f%%, op_acc: %.2f%%] [G loss: %f]" % (epoch, d_loss[0], 100*d_loss[3], 100*d_loss[4], g_loss[0]))
# If at save interval => save generated image samples
if epoch % sample_interval == 0:
self.save_model()
self.sample_images(epoch)
def sample_images(self, epoch):
noise = np.random.normal(0, 1, (10, self.latent_dim))
sampled_labels = np.array(([0] * 5) + ([1] * 5))
gen_imgs = self.generator.predict([noise, sampled_labels])
fig, axs = plt.subplots(10, 1)
for i in range(sampled_labels.size):
axs[i].plot(gen_imgs[i,:,0], color='black')
plt.show()
def save_model(self):
pass
acgan = ACGAN()
acgan.train(epochs=14000, batch_size=32, sample_interval=50)
```
| github_jupyter |
# Automated Machine Learning
_**Classification with Deployment using a Bank Marketing Dataset**_
## Contents
1. [Introduction](#Introduction)
1. [Setup](#Setup)
1. [Train](#Train)
1. [Results](#Results)
1. [Deploy](#Deploy)
1. [Test](#Test)
1. [Acknowledgements](#Acknowledgements)
## Introduction
In this example we use the UCI Bank Marketing dataset to showcase how you can use AutoML for a classification problem and deploy it to an Azure Container Instance (ACI). The classification goal is to predict if the client will subscribe to a term deposit with the bank.
If you are using an Azure Machine Learning Compute Instance, you are all set. Otherwise, go through the [configuration](../../../configuration.ipynb) notebook first if you haven't already to establish your connection to the AzureML Workspace.
Please find the ONNX related documentations [here](https://github.com/onnx/onnx).
In this notebook you will learn how to:
1. Create an experiment using an existing workspace.
2. Configure AutoML using `AutoMLConfig`.
3. Train the model using local compute with ONNX compatible config on.
4. Explore the results, featurization transparency options and save the ONNX model
5. Inference with the ONNX model.
6. Register the model.
7. Create a container image.
8. Create an Azure Container Instance (ACI) service.
9. Test the ACI service.
In addition this notebook showcases the following features
- **Blocking** certain pipelines
- Specifying **target metrics** to indicate stopping criteria
- Handling **missing data** in the input
## Setup
As part of the setup you have already created an Azure ML `Workspace` object. For AutoML you will need to create an `Experiment` object, which is a named object in a `Workspace` used to run experiments.
```
import json
import logging
from matplotlib import pyplot as plt
import pandas as pd
import os
import azureml.core
from azureml.core.experiment import Experiment
from azureml.core.workspace import Workspace
from azureml.core.dataset import Dataset
from azureml.train.automl import AutoMLConfig
from azureml.interpret import ExplanationClient
```
This sample notebook may use features that are not available in previous versions of the Azure ML SDK.
Accessing the Azure ML workspace requires authentication with Azure.
The default authentication is interactive authentication using the default tenant. Executing the `ws = Workspace.from_config()` line in the cell below will prompt for authentication the first time that it is run.
If you have multiple Azure tenants, you can specify the tenant by replacing the `ws = Workspace.from_config()` line in the cell below with the following:
```
from azureml.core.authentication import InteractiveLoginAuthentication
auth = InteractiveLoginAuthentication(tenant_id = 'mytenantid')
ws = Workspace.from_config(auth = auth)
```
If you need to run in an environment where interactive login is not possible, you can use Service Principal authentication by replacing the `ws = Workspace.from_config()` line in the cell below with the following:
```
from azureml.core.authentication import ServicePrincipalAuthentication
auth = auth = ServicePrincipalAuthentication('mytenantid', 'myappid', 'mypassword')
ws = Workspace.from_config(auth = auth)
```
For more details, see [aka.ms/aml-notebook-auth](http://aka.ms/aml-notebook-auth)
```
ws = Workspace.from_config()
# choose a name for experiment
experiment_name = "automl-classification-bmarketing-all"
experiment = Experiment(ws, experiment_name)
output = {}
output["Subscription ID"] = ws.subscription_id
output["Workspace"] = ws.name
output["Resource Group"] = ws.resource_group
output["Location"] = ws.location
output["Experiment Name"] = experiment.name
pd.set_option("display.max_colwidth", -1)
outputDf = pd.DataFrame(data=output, index=[""])
outputDf.T
```
## Create or Attach existing AmlCompute
You will need to create a compute target for your AutoML run. In this tutorial, you create AmlCompute as your training compute resource.
> Note that if you have an AzureML Data Scientist role, you will not have permission to create compute resources. Talk to your workspace or IT admin to create the compute targets described in this section, if they do not already exist.
#### Creation of AmlCompute takes approximately 5 minutes.
If the AmlCompute with that name is already in your workspace this code will skip the creation process.
As with other Azure services, there are limits on certain resources (e.g. AmlCompute) associated with the Azure Machine Learning service. Please read [this article](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-manage-quotas) on the default limits and how to request more quota.
```
from azureml.core.compute import ComputeTarget, AmlCompute
from azureml.core.compute_target import ComputeTargetException
# Choose a name for your CPU cluster
cpu_cluster_name = "cpu-cluster-4"
# Verify that cluster does not exist already
try:
compute_target = ComputeTarget(workspace=ws, name=cpu_cluster_name)
print("Found existing cluster, use it.")
except ComputeTargetException:
compute_config = AmlCompute.provisioning_configuration(
vm_size="STANDARD_DS12_V2", max_nodes=6
)
compute_target = ComputeTarget.create(ws, cpu_cluster_name, compute_config)
compute_target.wait_for_completion(show_output=True)
```
# Data
### Load Data
Leverage azure compute to load the bank marketing dataset as a Tabular Dataset into the dataset variable.
### Training Data
```
data = pd.read_csv(
"https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_train.csv"
)
data.head()
# Add missing values in 75% of the lines.
import numpy as np
missing_rate = 0.75
n_missing_samples = int(np.floor(data.shape[0] * missing_rate))
missing_samples = np.hstack(
(
np.zeros(data.shape[0] - n_missing_samples, dtype=np.bool),
np.ones(n_missing_samples, dtype=np.bool),
)
)
rng = np.random.RandomState(0)
rng.shuffle(missing_samples)
missing_features = rng.randint(0, data.shape[1], n_missing_samples)
data.values[np.where(missing_samples)[0], missing_features] = np.nan
if not os.path.isdir("data"):
os.mkdir("data")
# Save the train data to a csv to be uploaded to the datastore
pd.DataFrame(data).to_csv("data/train_data.csv", index=False)
ds = ws.get_default_datastore()
ds.upload(
src_dir="./data", target_path="bankmarketing", overwrite=True, show_progress=True
)
# Upload the training data as a tabular dataset for access during training on remote compute
train_data = Dataset.Tabular.from_delimited_files(
path=ds.path("bankmarketing/train_data.csv")
)
label = "y"
```
### Validation Data
```
validation_data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_validate.csv"
validation_dataset = Dataset.Tabular.from_delimited_files(validation_data)
```
### Test Data
```
test_data = "https://automlsamplenotebookdata.blob.core.windows.net/automl-sample-notebook-data/bankmarketing_test.csv"
test_dataset = Dataset.Tabular.from_delimited_files(test_data)
```
## Train
Instantiate a AutoMLConfig object. This defines the settings and data used to run the experiment.
|Property|Description|
|-|-|
|**task**|classification or regression or forecasting|
|**primary_metric**|This is the metric that you want to optimize. Classification supports the following primary metrics: <br><i>accuracy</i><br><i>AUC_weighted</i><br><i>average_precision_score_weighted</i><br><i>norm_macro_recall</i><br><i>precision_score_weighted</i>|
|**iteration_timeout_minutes**|Time limit in minutes for each iteration.|
|**blocked_models** | *List* of *strings* indicating machine learning algorithms for AutoML to avoid in this run. <br><br> Allowed values for **Classification**<br><i>LogisticRegression</i><br><i>SGD</i><br><i>MultinomialNaiveBayes</i><br><i>BernoulliNaiveBayes</i><br><i>SVM</i><br><i>LinearSVM</i><br><i>KNN</i><br><i>DecisionTree</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>GradientBoosting</i><br><i>TensorFlowDNN</i><br><i>TensorFlowLinearClassifier</i><br><br>Allowed values for **Regression**<br><i>ElasticNet</i><br><i>GradientBoosting</i><br><i>DecisionTree</i><br><i>KNN</i><br><i>LassoLars</i><br><i>SGD</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>TensorFlowLinearRegressor</i><br><i>TensorFlowDNN</i><br><br>Allowed values for **Forecasting**<br><i>ElasticNet</i><br><i>GradientBoosting</i><br><i>DecisionTree</i><br><i>KNN</i><br><i>LassoLars</i><br><i>SGD</i><br><i>RandomForest</i><br><i>ExtremeRandomTrees</i><br><i>LightGBM</i><br><i>TensorFlowLinearRegressor</i><br><i>TensorFlowDNN</i><br><i>Arima</i><br><i>Prophet</i>|
|**allowed_models** | *List* of *strings* indicating machine learning algorithms for AutoML to use in this run. Same values listed above for **blocked_models** allowed for **allowed_models**.|
|**experiment_exit_score**| Value indicating the target for *primary_metric*. <br>Once the target is surpassed the run terminates.|
|**experiment_timeout_hours**| Maximum amount of time in hours that all iterations combined can take before the experiment terminates.|
|**enable_early_stopping**| Flag to enble early termination if the score is not improving in the short term.|
|**featurization**| 'auto' / 'off' Indicator for whether featurization step should be done automatically or not. Note: If the input data is sparse, featurization cannot be turned on.|
|**n_cross_validations**|Number of cross validation splits.|
|**training_data**|Input dataset, containing both features and label column.|
|**label_column_name**|The name of the label column.|
**_You can find more information about primary metrics_** [here](https://docs.microsoft.com/en-us/azure/machine-learning/service/how-to-configure-auto-train#primary-metric)
```
automl_settings = {
"experiment_timeout_hours": 0.3,
"enable_early_stopping": True,
"iteration_timeout_minutes": 5,
"max_concurrent_iterations": 4,
"max_cores_per_iteration": -1,
# "n_cross_validations": 2,
"primary_metric": "AUC_weighted",
"featurization": "auto",
"verbosity": logging.INFO,
}
automl_config = AutoMLConfig(
task="classification",
debug_log="automl_errors.log",
compute_target=compute_target,
experiment_exit_score=0.9984,
blocked_models=["KNN", "LinearSVM"],
enable_onnx_compatible_models=True,
training_data=train_data,
label_column_name=label,
validation_data=validation_dataset,
**automl_settings,
)
```
Call the `submit` method on the experiment object and pass the run configuration. Execution of local runs is synchronous. Depending on the data and the number of iterations this can run for a while. Validation errors and current status will be shown when setting `show_output=True` and the execution will be synchronous.
```
remote_run = experiment.submit(automl_config, show_output=False)
```
Run the following cell to access previous runs. Uncomment the cell below and update the run_id.
```
# from azureml.train.automl.run import AutoMLRun
# remote_run = AutoMLRun(experiment=experiment, run_id='<run_ID_goes_here')
# remote_run
# Wait for the remote run to complete
remote_run.wait_for_completion()
# Retrieve the best Run object
best_run = remote_run.get_best_child()
```
## Transparency
View featurization summary for the best model - to study how different features were transformed. This is stored as a JSON file in the outputs directory for the run.
```
# Download the featurization summary JSON file locally
best_run.download_file(
"outputs/featurization_summary.json", "featurization_summary.json"
)
# Render the JSON as a pandas DataFrame
with open("featurization_summary.json", "r") as f:
records = json.load(f)
pd.DataFrame.from_records(records)
```
## Results
```
from azureml.widgets import RunDetails
RunDetails(remote_run).show()
```
### Retrieve the Best Model's explanation
Retrieve the explanation from the best_run which includes explanations for engineered features and raw features. Make sure that the run for generating explanations for the best model is completed.
```
# Wait for the best model explanation run to complete
from azureml.core.run import Run
model_explainability_run_id = remote_run.id + "_" + "ModelExplain"
print(model_explainability_run_id)
model_explainability_run = Run(
experiment=experiment, run_id=model_explainability_run_id
)
model_explainability_run.wait_for_completion()
# Get the best run object
best_run = remote_run.get_best_child()
```
#### Download engineered feature importance from artifact store
You can use ExplanationClient to download the engineered feature explanations from the artifact store of the best_run.
```
client = ExplanationClient.from_run(best_run)
engineered_explanations = client.download_model_explanation(raw=False)
exp_data = engineered_explanations.get_feature_importance_dict()
exp_data
```
#### Download raw feature importance from artifact store
You can use ExplanationClient to download the raw feature explanations from the artifact store of the best_run.
```
client = ExplanationClient.from_run(best_run)
engineered_explanations = client.download_model_explanation(raw=True)
exp_data = engineered_explanations.get_feature_importance_dict()
exp_data
```
### Retrieve the Best ONNX Model
Below we select the best pipeline from our iterations. The `get_output` method returns the best run and the fitted model. The Model includes the pipeline and any pre-processing. Overloads on `get_output` allow you to retrieve the best run and fitted model for *any* logged metric or for a particular *iteration*.
Set the parameter return_onnx_model=True to retrieve the best ONNX model, instead of the Python model.
```
best_run, onnx_mdl = remote_run.get_output(return_onnx_model=True)
```
### Save the best ONNX model
```
from azureml.automl.runtime.onnx_convert import OnnxConverter
onnx_fl_path = "./best_model.onnx"
OnnxConverter.save_onnx_model(onnx_mdl, onnx_fl_path)
```
### Predict with the ONNX model, using onnxruntime package
```
import sys
import json
from azureml.automl.core.onnx_convert import OnnxConvertConstants
from azureml.train.automl import constants
from azureml.automl.runtime.onnx_convert import OnnxInferenceHelper
def get_onnx_res(run):
res_path = "onnx_resource.json"
run.download_file(
name=constants.MODEL_RESOURCE_PATH_ONNX, output_file_path=res_path
)
with open(res_path) as f:
result = json.load(f)
return result
if sys.version_info < OnnxConvertConstants.OnnxIncompatiblePythonVersion:
test_df = test_dataset.to_pandas_dataframe()
mdl_bytes = onnx_mdl.SerializeToString()
onnx_result = get_onnx_res(best_run)
onnxrt_helper = OnnxInferenceHelper(mdl_bytes, onnx_result)
pred_onnx, pred_prob_onnx = onnxrt_helper.predict(test_df)
print(pred_onnx)
print(pred_prob_onnx)
else:
print("Please use Python version 3.6 or 3.7 to run the inference helper.")
```
## Deploy
### Retrieve the Best Model
Below we select the best pipeline from our iterations. The `get_best_child` method returns the Run object for the best model based on the default primary metric. There are additional flags that can be passed to the method if we want to retrieve the best Run based on any of the other supported metrics, or if we are just interested in the best run among the ONNX compatible runs. As always, you can execute `??remote_run.get_best_child` in a new cell to view the source or docs for the function.
```
??remote_run.get_best_child
```
#### Widget for Monitoring Runs
The widget will first report a "loading" status while running the first iteration. After completing the first iteration, an auto-updating graph and table will be shown. The widget will refresh once per minute, so you should see the graph update as child runs complete.
**Note:** The widget displays a link at the bottom. Use this link to open a web interface to explore the individual run details
```
best_run = remote_run.get_best_child()
model_name = best_run.properties["model_name"]
script_file_name = "inference/score.py"
best_run.download_file("outputs/scoring_file_v_1_0_0.py", "inference/score.py")
```
### Register the Fitted Model for Deployment
If neither `metric` nor `iteration` are specified in the `register_model` call, the iteration with the best primary metric is registered.
```
description = "AutoML Model trained on bank marketing data to predict if a client will subscribe to a term deposit"
tags = None
model = remote_run.register_model(
model_name=model_name, description=description, tags=tags
)
print(
remote_run.model_id
) # This will be written to the script file later in the notebook.
```
### Deploy the model as a Web Service on Azure Container Instance
```
from azureml.core.model import InferenceConfig
from azureml.core.webservice import AciWebservice
from azureml.core.webservice import Webservice
from azureml.core.model import Model
from azureml.core.environment import Environment
inference_config = InferenceConfig(entry_script=script_file_name)
aciconfig = AciWebservice.deploy_configuration(
cpu_cores=2,
memory_gb=2,
tags={"area": "bmData", "type": "automl_classification"},
description="sample service for Automl Classification",
)
aci_service_name = model_name.lower()
print(aci_service_name)
aci_service = Model.deploy(ws, aci_service_name, [model], inference_config, aciconfig)
aci_service.wait_for_deployment(True)
print(aci_service.state)
```
### Get Logs from a Deployed Web Service
Gets logs from a deployed web service.
```
# aci_service.get_logs()
```
## Test
Now that the model is trained, run the test data through the trained model to get the predicted values. This calls the ACI web service to do the prediction.
Note that the JSON passed to the ACI web service is an array of rows of data. Each row should either be an array of values in the same order that was used for training or a dictionary where the keys are the same as the column names used for training. The example below uses dictionary rows.
```
# Load the bank marketing datasets.
from numpy import array
X_test = test_dataset.drop_columns(columns=["y"])
y_test = test_dataset.keep_columns(columns=["y"], validate=True)
test_dataset.take(5).to_pandas_dataframe()
X_test = X_test.to_pandas_dataframe()
y_test = y_test.to_pandas_dataframe()
import requests
X_test_json = X_test.to_json(orient="records")
data = '{"data": ' + X_test_json + "}"
headers = {"Content-Type": "application/json"}
resp = requests.post(aci_service.scoring_uri, data, headers=headers)
y_pred = json.loads(json.loads(resp.text))["result"]
actual = array(y_test)
actual = actual[:, 0]
print(len(y_pred), " ", len(actual))
```
### Calculate metrics for the prediction
Now visualize the data as a confusion matrix that compared the predicted values against the actual values.
```
%matplotlib notebook
from sklearn.metrics import confusion_matrix
import itertools
cf = confusion_matrix(actual, y_pred)
plt.imshow(cf, cmap=plt.cm.Blues, interpolation="nearest")
plt.colorbar()
plt.title("Confusion Matrix")
plt.xlabel("Predicted")
plt.ylabel("Actual")
class_labels = ["no", "yes"]
tick_marks = np.arange(len(class_labels))
plt.xticks(tick_marks, class_labels)
plt.yticks([-0.5, 0, 1, 1.5], ["", "no", "yes", ""])
# plotting text value inside cells
thresh = cf.max() / 2.0
for i, j in itertools.product(range(cf.shape[0]), range(cf.shape[1])):
plt.text(
j,
i,
format(cf[i, j], "d"),
horizontalalignment="center",
color="white" if cf[i, j] > thresh else "black",
)
plt.show()
```
### Delete a Web Service
Deletes the specified web service.
```
aci_service.delete()
```
## Acknowledgements
This Bank Marketing dataset is made available under the Creative Commons (CCO: Public Domain) License: https://creativecommons.org/publicdomain/zero/1.0/. Any rights in individual contents of the database are licensed under the Database Contents License: https://creativecommons.org/publicdomain/zero/1.0/ and is available at: https://www.kaggle.com/janiobachmann/bank-marketing-dataset .
_**Acknowledgements**_
This data set is originally available within the UCI Machine Learning Database: https://archive.ics.uci.edu/ml/datasets/bank+marketing
[Moro et al., 2014] S. Moro, P. Cortez and P. Rita. A Data-Driven Approach to Predict the Success of Bank Telemarketing. Decision Support Systems, Elsevier, 62:22-31, June 2014
| github_jupyter |
```
import xarray as xr
import glob
from xgcm import Grid
import matplotlib.pyplot as plt
import cmocean
import cartopy.crs as ccrs
%pylab inline
import warnings
warnings.simplefilter('ignore') # filter some warning messages
in_file = 'roho160_clim_GLORYS_20170115_to_20191130_zarr.nc'
lat1, lon1 = 59.7,4.8
lat2, lon2 = 59.2,5
# choose a variiable and season to plot
var = 'u'
seasn = 2
seasons_nm = ['DJF','MAM','JJA','SON']
print(in_file)
print('Point 1: ', lat1,'N, ',lon1,'W')
print('Point 2: ', lat2,'N, ',lon2,'W')
print('Variable: ',var)
print('Season: ', seasons_nm[seasn])
def compute_depth_layers(ds, grid, hmin=0.1):
## from https://raphaeldussin.medium.com/modern-python-tools-for-the-roms-ocean-model-bfca8642db01
""" compute depths of ROMS vertical levels (Vtransform = 2) """
# compute vertical transformation functional
S_rho = (ds.hc * ds.s_rho + ds.Cs_r * ds.h) / (ds.hc + ds.h)
S_w = (ds.hc * ds.s_w + ds.Cs_w * ds.h) / (ds.hc + ds.h)
# compute depth of rho (layers) and w (interfaces) points
z_rho = ds.zeta + (ds.zeta + ds.h) * S_rho
z_w = ds.zeta + (ds.zeta + ds.h) * S_w
# transpose arrays and fill NaNs with a minimal depth
ds['z_rho'] = z_rho.transpose(*('season', 's_rho','yh','xh'),
transpose_coords=False).fillna(hmin)
ds['z_w'] = z_w.transpose(*('season', 's_w','yh','xh'),
transpose_coords=False).fillna(hmin)
# interpolate depth of levels at U and V points
ds['z_u'] = grid.interp(ds['z_rho'], 'X', boundary='fill')
ds['z_v'] = grid.interp(ds['z_rho'], 'Y', boundary='fill')
# compute layer thickness as difference between interfaces
ds['dz'] = grid.diff(ds['z_w'], 'Z')
# add z_rho and z_w to xarray coordinates
ds = ds.set_coords(['z_rho', 'z_w', 'z_v', 'z_u'])
return ds
def rot_coords(ds):
# modified from Trond's functions
# https://github.com/trondkr/romstools/blob/master/VolumeFlux/tools.py
# converto to curvilinear coordinates
# calculate u,v in xh,yh grid points
u_rho = (ds.u[:,:,:,:-1].values+ds.u[:,:,:,1::].values)/2.0
v_rho = (ds.v[:,:,:-1,:].values+ds.v[:,:,1::,:].values)/2.0
print(np.shape(u_rho),np.shape(v_rho))
# for each vertical level
angle = np.full((np.shape(v_rho)),np.nan)
for iy,i in enumerate(ds.s_rho.values):
angle[:,iy,:,:] = ds.angle.values
print("angle : {}".format(np.shape(angle)))
u=u_rho
v=v_rho
print("u : {} v: {}".format(np.shape(u), np.shape(v)))
u_rotated = u*np.cos(angle)-v*np.sin(angle)
v_rotated = v*np.cos(angle)+u*np.sin(angle)
print("u : {} and v: {}".format(np.shape(u_rotated),
np.shape(v_rotated)))
cv_dz = xr.DataArray(
name="u",
data=u_rotated,
coords={'season': (['season'],['DJF','JJA','MAM','SON']),
's_rho': (['s_rho'], ds.s_rho.values),
'lat': (['yh','xh'], ds["lat_rho"].values),
'lon': (['yh','xh'], ds["lon_rho"].values),
'z_rho': (['season','s_rho','yh', 'xh'],ds.z_rho.values)},
dims=['season','s_rho',"yh","xh"]
).to_dataset()
speed=np.square(u_rotated**2 + v_rotated**2)
cv_dz["v"]=(['season','s_rho','yh', 'xh'], v_rotated)
cv_dz["speed"]=(['season','s_rho','yh', 'xh'], speed)
cv_dz["temp"]=(['season','s_rho','yh', 'xh'], ds.temp.values)
#print(cv_dz)
return cv_dz
def line_points(temp):
# modified from Trond's functions
# https://github.com/trondkr/romstools/blob/master/VolumeFlux/tools.py
# find indices for both points
lo = temp.lon.values - lon1
la = temp.lat.values - lat1
diff = (lo*lo) + (la*la)
jj1, ii1 = np.where(diff==diff.min())
jj1 = jj1[0]
ii1 = ii1[0]
print(jj1,ii1, 'yh,xh')
lo = temp.lon.values - lon2
la = temp.lat.values - lat2
diff = (lo*lo) + (la*la)
jj2, ii2 = np.where(diff==diff.min())
jj2 = jj2[0]
ii2 = ii2[0]
#print(jj2,ii2, 'yh,xh')
#print(temp.lon[jj1,ii1].values, temp.lat[jj1,ii1].values)
#print(temp.lon[jj2,ii2].values, temp.lat[jj2,ii2].values)
# find the line - in indices
if ii1 != ii2:
aj = (jj2 - jj1)/(ii2-ii1)
bj = jj1 - aj * ii1
if jj1 != jj2:
ai = (ii2 - ii1)/(jj2-jj1)
bi = ii1 - ai * jj1
if (abs(aj)) <= 1:
if (ii2<ii1): #swap
i = ii1
j = jj1
ii1 = ii2
jj1 = jj2
ii2 = i
jj2 = j
# nearest
n = 0
near = np.zeros((ii2-ii1+1,5))
for i in range(ii1,ii2+1):
j = aj*i + bj
near[n,0]=i
near[n,1]=j
near[n,2]=np.floor(j)
near[n,3]=np.ceil(j)
near[n,4]=np.round(j)
n += 1
else:
if (jj2<jj1): #swap
i = ii1
j = jj1
ii1 = ii2
jj1 = jj2
ii2 = i
jj2 = j
# nearest
n = 0
near = np.zeros((jj2-jj1+1,5))
for j in range(jj1,jj2+1):
i = ai*j + bi
near[n,0]=j
near[n,1]=i
near[n,2]=np.floor(i)
near[n,3]=np.ceil(i)
near[n,4]=np.round(i)
n += 1
return near, aj
def v_transect(temp, near, aj):
# modified from Trond's functions
# https://github.com/trondkr/romstools/blob/master/VolumeFlux/tools.py
# calculate vertical transect
transect = np.zeros((temp.z_rho.shape[0],near.shape[0]))
for n in range(near.shape[0]):
if (abs(aj)<=1):
if (near[n,2]==near[n,3]):
transect[:,n] = temp[int(near[n,2]), int(near[n,0]),:]
else:
transect[:,n] = (int(near[n,1]) - int(near[n,2])) * temp[int(near[n,3]), int(near[n,0]),:] + \
(int(near[n,3]) - int(near[n,1])) * temp[int(near[n,2]), int(near[n,0]),:]
else:
if (near[n,2] == near[n,3]):
transect[:,n] = temp[int(near[n,0]), int(near[n,2]),:]
else:
transect[:,n] = (int(near[n,1]) - int(near[n,2])) * temp[int(near[n,0]), int(near[n,3]), :] + \
(int(near[n,3]) - int(near[n,1])) * temp[int(near[n,0]), int(near[n,2]), :]
# lats and lons of line
lons = np.zeros(near.shape[0])
lats = np.zeros(near.shape[0])
l = near.shape[0]
if lon2-lon1<0: # flip
for i in range(near.shape[0]):
lons[i]=temp.lon[near[l-i-1,0].astype(int),near[l-i-1,4].astype(int)].values
lats[i]=temp.lat[near[l-i-1,0].astype(int),near[l-i-1,4].astype(int)].values
else:
for i in range(near.shape[0]):
lons[i]=temp.lon[near[i,0].astype(int),near[i,4].astype(int)].values
lats[i]=temp.lat[near[i,0].astype(int),near[i,4].astype(int)].values
return transect, lats, lons
# open zarr file
dz = xr.open_zarr(in_file, consolidated=True)
# calculate climatology
dz_clim = dz.groupby('time.season').mean('time',keep_attrs=True)
#dz_clim
# Create xgcm grid object
grid_dz = Grid(dz_clim, coords={'X': {'center': 'xh', 'outer': 'xq'},
'Y': {'center': 'yh', 'outer': 'yq'},
'Z': {'center': 's_rho', 'outer': 's_w'}},
periodic=False)
# Add depths of layers and interfaces to dataset
dz_clz = compute_depth_layers(dz_clim, grid_dz)
#dz_clz
cv_dz = rot_coords(dz_clz) # transform to curvilinear coordinates
# transform vertical coordinates on one variable
target_depth_levels = np.arange(0,-300,-10)
snapshot = cv_dz.isel(season=seasn)
sndz = grid_dz.transform(snapshot[var], 'Z', target_depth_levels, target_data=snapshot['z_rho'], method='linear')
# plot at -50 depth to show line
plt.figure(figsize=[10,8])
ax = plt.axes(projection=ccrs.PlateCarree())
p = sndz.sel(z_rho=-50).plot(ax=ax, x='lon', y='lat',
cmap=cmocean.cm.thermal,
add_labels=True,
transform=ccrs.PlateCarree())
plt.plot([lon1,lon2],[lat1,lat2],'kd-')
# background
url = 'http://map1c.vis.earthdata.nasa.gov/wmts-geo/wmts.cgi'
p.axes.add_wmts(url, 'BlueMarble_NextGeneration')
p.axes.set_extent([3,7.5,58.5,61.5])
pnts, aj = line_points(sndz)
transect, lats, lons = v_transect(sndz, pnts, aj)
fig = plt.figure(figsize=[10,6])
ax1 = fig.add_subplot(111)
plt.contourf(lons,sndz.z_rho,transect, 20,cmap=cmocean.cm.balance)
cbar = plt.colorbar()
cbar.set_label(dz_clim[var].long_name+' ('+dz_clim[var].units+')')
plt.ylabel('Z (m)')
plt.xlabel('Longitude')
ax2 = ax1.twiny()
ax2.plot(lats, np.full(len(lats), np.nan)) # Create a dummy plot
ax2.cla()
xx = np.round(ax2.get_xticks() * (lats[-1]-lats[0])+lats[0],1)
ax2.set_xticklabels(xx)
plt.xlabel('Latitude')
plt.title('Season: '+seasons_nm[seasn],loc='left')
plt.show()
```
| github_jupyter |
# HyperLearning AI - Introduction to Python
An introductory course to the Python 3 programming language, with a curriculum aligned to the Certified Associate in Python Programming (PCAP) examination syllabus (PCAP-31-02).<br/>
https://knowledgebase.hyperlearning.ai/courses/introduction-to-python
## 07. Functions and Modules Part 2
https://knowledgebase.hyperlearning.ai/en/courses/introduction-to-python/modules/7/functions-and-modules-part-2
In this module we will formally introduce Python modules and packages, including how to write and use Python modules, how to construct and distribute Python packages, how to hide Python module entities, how to document Python modules, and Python hashbangs. Specifically we will cover:
* **Python Modules** - importing modules, qualifying entities, initialising modules, writing and using modules, the name variable, Python hashbangs, and module documentation
* **Python Packages** - creating packages, packages vs directories, the init file, and hiding module entities
### 1. Modules
#### 1.1. Importing Modules
```
# Import our numbertools module
import numbertools
# Call functions from the module
print(numbertools.is_int(0.5))
print(numbertools.is_even(1_000_002))
print(numbertools.is_prime(277))
print(numbertools.is_fibonacci(12))
print(numbertools.is_perfect_square(1444))
# Access variables from the module
print(numbertools.mobius_phi)
# Create an alias for a module
import numbertools as nt
# Call module entities qualified with the module alias
print(nt.is_perfect_square(9801))
print(nt.mobius_phi)
# List all the function and variable names in a given module
print(dir(nt))
print(dir(math))
# Import specific entities
from math import cos, pi, radians, sin, tan
# Use the specifically imported entities
print(round(cos(math.radians(90)), 0))
print(round(sin(math.radians(90)), 0))
print(round(tan(math.radians(45)), 0))
print(round(pi, 10))
```
#### 1.2. Module Search Path
```
# Examine and modify sys.path
import sys
# Examine sys.path
print(sys.path)
# Append a location to sys.path
sys.path.append('/foo/bar/code')
print(sys.path)
```
#### 1.5. Module Documentation
```
# Access and display a module's docstring
print(numbertools.__doc__)
# Access and display a built-in module's docstring
print(math.__doc__)
# Access and display a function's docstring
print(numbertools.is_fibonacci.__doc__)
# Access and display a built-in function's docstring
print(sin.__doc__)
# Display help on the numbertools module
help(numbertools)
# Display help on a specific function
help(numbertools.is_perfect_square)
# Display help on a specifc built-in function
help(len)
# Display help on the built-in math module
help(math)
```
#### 1.7. PYC Files
```
# Generate a byte code file of our numbertools module
import py_compile
py_compile.compile('numbertools.py')
# Generate a byte code file of our numbertools module in a custom location
py_compile.compile('numbertools.py', cfile='/tmp/numbertools.pyc')
```
### 2. Packages
#### 2.1. Importing Packages
```
# Import a module from a nested package in the Pandas library
import pandas.api.types as types
print(types.is_integer_dtype(str))
# Alternatively use from to import a specific module
from pandas.api import types
print(types.is_integer_dtype(str))
# Alternatively use from to import a specific function from a specific module
from pandas.api.types import is_integer_dtype
print(is_integer_dtype(str))
```
#### 2.2.8. Install from Local Wheel
```
# Import our generated distribution package myutils
import myutils
# Alternatively import a specific module from a specific package
import myutils.collections.dictutils as dictutils
# Call one of our myutils bespoke functions
my_dict = {
1: ['python', 3.8],
2: ['java', 11],
3: ['scala', 2.13]
}
# Convert a dictionary to a Pandas DataFrame using our user-defined dictutils.convert_to_dataframe() function
df = dictutils.convert_to_dataframe(my_dict, ['Language', 'Version'])
df.head()
```
| github_jupyter |
**1**. (25 points)
- Write a **recursive** function that returns the length of the hailstone sequence staring with a positive integer $n$. (15 points)
The hailstone sequence is defined by the following rules:
```
- If n is 1, stop
- If n is even, divide by 2 and repeat
- If n is odd, multiply by 3 and add 1 and repeat
```
For example, the hailstone sequence starting with $n = 3$ has length 8:
```
- 3, 10, 5, 16, 8, 4, 2, 1
```
Use the `functools` package to avoid duplicate function calls.
- Find the number that gives the longest sequence for starting numbers less than 100,000. Report the number and the length of the generated sequence. (10 points)
```
from functools import lru_cache
@lru_cache(None)
def hailstone(n, k=1):
"""Reprots length of hailstone (Collatz) sequence startign with n."""
if n == 1:
return k
else:
if n % 2 == 0:
return hailstone(n // 2, k+1)
else:
return hailstone(n*3 + 1, k+1)
best = [0, 0]
for i in range(1, 100000):
s = hailstone(i)
if s > best[1]:
best = (i, s)
best
```
An alternative solution.
```
@lru_cache(None)
def hailstone_alt(n):
"""Reprots length of hailstone (Collatz) sequence startign with n."""
if n == 1:
return 1
else:
if n % 2 == 0:
return 1 + hailstone_alt(n // 2)
else:
return 1 + hailstone_alt(n*3 + 1)
hailstone_alt(3)
best = [0, 0]
for i in range(1, 100000):
s = hailstone_alt(i)
if s > best[1]:
best = (i, s)
best
```
**2**. (25 points)
- Create a `pnadas` DataFrame called `df` from the data set at https://bit.ly/2ksKr8f, taking care to only read in the `time` and `value` columns. (5 points)
- Fill all rows with missing values with the value from the last non-missing value (i.e. forward fill) (5 points)
- Convert to a `pandas` Series `s` using `time` as the index (5 points)
- Create a new series `s1` with the rolling average using a shifting window of size 7 and a minimum period of 1 (5 points)
- Report the `time` and value for the largest rolling average (5 points)
```
import pandas as pd
df = pd.read_csv('https://bit.ly/2ksKr8f', usecols=['time', 'value'])
df = df.fillna(method='ffill')
df.head()
```
Note: The pd.Series constructor has quite unintuitive behavior when the `index` argument is provided. See `DataFrame_to_Series.ipynb` for this.
```
s = pd.Series(data=df['value'])
s.index = df['time']
s.head()
s1 = s.rolling(7, 1).mean()
s1.head()
s1.sort_values(ascending=False).head(1)
```
**3**. (25 points)
- Get information in JSON format about startship 23 from the Star Wars API https://swapi.co/api using the `requests` package (5 points)
- Report the time interval between `created` and `edited` in minutes using the `pendulum` package (5 points)
- Replace the URL values stored at the `films` key with the titles of the actual films (5 points)
- Save the new JSON (with film titles and not URLs) to a file `ship.json` (5 points)
- Read in the JSON file you have just saved as a Python dictionary (5 points)
```
import requests
url = 'https://swapi.co/api/starships/23'
ship = requests.get(url).json()
ship
import pendulum
created = pendulum.parse(ship['created'])
edited = pendulum.parse(ship['edited'])
(edited - created).in_minutes()
films = [requests.get(film).json()['title'] for film in ship['films']]
films
ship['films'] = films
import json
with open('ship.json', 'w') as f:
json.dump(ship, f)
with open('ship.json') as f:
ship = json.load(f)
ship
```
**4**. (25 points)
Use SQL to answer the following questions using the SQLite3 database `anemia.db`:
- Count the number of male and female patients (5 points)
- Find the average age of male and female patients (as of right now) (5 points)
- Show the sex, hb and name of patients with severe anemia ordered by severity. Severe anemia is defined as
- Hb < 7 if female
- Hb < 8 if male
(15 points)
You many assume `pid` is the PRIMARY KEY in the patients table and the FOREIGN KEY in the labs table.
Note: Hb is short for hemoglobin levels.
Hint: In SQLite3, you can use `DATE('now')` to get today's date.
```
%load_ext sql
%sql sqlite:///anemia.db
%%sql
SELECT * FROM sqlite_master WHERe type='table'
%%sql
SELECT * FROM patients LIMIT 3
%%sql
SELECT * FROM labs LIMIT 3
%%sql
SELECT sex, COUNT(sex)
FROM patients
GROUP BY sex
%%sql
SELECT date('now')
%%sql
SELECT sex, round(AVG(date('now') - birthday), 1)
FROM patients
GROUP BY sex
%%sql
SELECT sex, hb, name
FROM patients, labs
WHERE patients.pid = labs.pid AND
((sex = 'M' AND hb < 8) OR (sex = 'F' AND hb < 7))
ORDER BY hb
```
| github_jupyter |
# A Brief Introduction to Jupyter Notebooks
Jupyter Notebooks combine an execution environment for R/Python/Julia/Haskell with written instructions/documentation/descriptions as markdown. They are organized in _cells_, and each cell has a type. For use, two types of cells are relevant, Markdown and Code.
## Markdown Cells
The Markdown cells contain textual information that is formatted using Markdown. An explanation for how Markdown works can be found as part of the [Juypter Documentation](https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html). For this exercise, we use the Markdown cells to provide explanations, define tasks, and ask questions.
## Code Cells
The code cells contain exectuable code. Each cell is executed on its own and there is no fixed order for the execution. To execute the code in a cell, you simply have to click on the _Run_ button at the top of the page or hit Ctrl+Enter. Code has to be written in R, Python, Julia, or Haskell. All cells within the same notebook must use the same language, which is defined by the kernel of the notebook. We provide the notebooks to you with the Python kernel enabled. You may switch to a different kernel using the menu bar by clicking on Kernel-->Change Kernel and then selecting the language of your choice.
## The internal state
While the cells are executed on their own, they all share the same state. For example, if you load a library in one cell, it will be available in all cells. Similarly, if you store something in a variable, this variable will be globally accessible from all other cells. The state can be reset by restarting the kernel. This usually happens automatically, when you re-open the notebook. You can also trigger this manually using the menu bar by clicking on Kernel-->Restart.
## The Output
The output of a code cell appears below the cell. By default, the result of the last executed line is printed. Other textual outputs can be generated by using the print-commands of the programming languange. The generation of plots and similar elements will be covered in this exercise.
## Hello World
Below this cell, you find a code cell that contains the code for printing "Hello World" in Python. Execute the cell and see what happens.
```
print("Hello World")
```
You actually do not require the print to achieve the same result, because the return value of the final line is printed automatically. The cell below accomplishes (almost) the same thing.
```
"Hello World"
```
# Programming in this Exercise
You may solve the problems in whichever programming language you desire. You may or may not use Jupyter Notebooks. Within this exercise, we are interested in the interpretation of results, not the programming to achieve this. We give general guidance on how to solve problems with Python as part of the descriptions of the exercises. However, other languages, especially R, are also suited for solving the exerices. However, in case you have programming problems, we will only help you (within reasonable limits) if you use Python.
Please be reminded that these exercises are primarily designed for Computer Science M.Sc. students. Thus, we assume that all students poses basic programming skills and are able to learn new programming languages on their own. We will also usually not comment on your code quality. In case you have no experience in programming (at all) you may find this exercise difficult. If you only have experience with other languages, but not Python, you should be able to solve all tasks using help from the internet without problems. Google and StackOverflow are your friends.
| github_jupyter |
... ***CURRENTLY UNDER DEVELOPMENT*** ...
## Obtain synthetic waves and water level timeseries under a climate change scenario (future AWTs occurence probability & future TCs occurrence probability)
inputs required:
* Historical DWTs (for plotting)
* Historical wave families (for plotting)
* Synthetic DWTs ENSO climate change
* Probability of TCs under climate change
* Historical intradaily hydrograph parameters
* TCs waves
* Fitted multivariate extreme model for the waves associated to each DWT
in this notebook:
* Generate synthetic time series of wave conditions
In the case of **waves associated to a TC event**, the associated simulated waves from WWIII simulation that have been reconstructed in notebook 07 are use whenever the simulated TC enters the 4 degrees radio. The probability of entering the 4 degree radio (from the 14 degree radio) is modified according to future changes under a climate change scenario
```
#!/usr/bin/env python
# -*- coding: utf-8 -*-
# common
import os
import os.path as op
# pip
import numpy as np
import xarray as xr
import pandas as pd
from datetime import datetime
import matplotlib.pyplot as plt
# DEV: override installed teslakit
import sys
sys.path.insert(0, op.join(os.path.abspath(''), '..', '..','..', '..'))
# teslakit
from teslakit.database import Database
from teslakit.climate_emulator import Climate_Emulator
from teslakit.waves import AWL, Aggregate_WavesFamilies
from teslakit.plotting.outputs import Plot_FitSim_Histograms
from teslakit.plotting.extremes import Plot_FitSim_AnnualMax, Plot_FitSim_GevFit, Plot_Fit_QQ
from teslakit.plotting.waves import Plot_Waves_Histogram_FitSim
from teslakit.plotting.climate_change import Plot_RCP_ocurrence
```
## Database and Site parameters
```
# --------------------------------------
# Teslakit database
p_data = r'/Users/albacid/Projects/TeslaKit_projects'
# offshore
db = Database(p_data)
db.SetSite('ROI')
# climate change - S6
db_S6 = Database(p_data)
db_S6.SetSite('ROI_CC_S6')
# climate change - S5
db_S5 = Database(p_data)
db_S5.SetSite('ROI_CC_S5')
# climate emulator simulation modified path
p_S6_CE_sims = op.join(db_S6.paths.site.EXTREMES.climate_emulator, 'Simulations')
# --------------------------------------
# Load data for climate emulator simulation climate change: ESTELA DWT and TCs (MU, TAU)
DWTs_sim = db_S5.Load_ESTELA_DWT_sim() # DWTs climate change
TCs_params = db.Load_TCs_r2_sim_params() # TCs parameters (copula generated)
TCs_RBFs = db.Load_TCs_sim_r2_rbf_output() # TCs numerical_IH-RBFs_interpolation output
probs_TCs = db.Load_TCs_probs_synth() # TCs synthetic probabilities
pchange_TCs = probs_TCs['category_change_cumsum'].values[:]
l_mutau_wt = db.Load_MU_TAU_hydrograms() # MU - TAU intradaily hidrographs for each DWT
MU_WT = np.array([x.MU.values[:] for x in l_mutau_wt]) # MU and TAU numpy arrays
TAU_WT = np.array([x.TAU.values[:] for x in l_mutau_wt])
# --------------------------------------
# Load climate change data
lon_rcp, lat_rcp, RCP85ratioHIST_occurrence = db.Load_RCP85()
# solve first 10 DWTs simulations
DWTs_sim = DWTs_sim.isel(n_sim=slice(0, 10))
print(DWTs_sim)
```
## Obtain future TCs occurence probability at the site
```
# ROI point
lon_p = 167.5
lat_p = 9.75
# find closest point at RCP data
lon_g, lat_g = np.meshgrid(lon_rcp, lat_rcp)
dif = np.sqrt((lon_g - lon_p)**2 + (lat_g - lat_p)**2)
min_ind = np.where(dif == np.min(dif))
ix_lon, ix_lat = min_ind[0], min_ind[1]
# TCs ocurrence probability at site
TCs_occurrence_prob = RCP85ratioHIST_occurrence[ix_lon, ix_lat]
# Plot global map and location of Site
Plot_RCP_ocurrence(lon_g, lat_g, RCP85ratioHIST_occurrence, ix_lon, ix_lat);
#--------------------------------------
# Modify future probability of a TC affecting the site (modify probability of TC in r1, entering r2)
pchange_TCs += pchange_TCs * TCs_occurrence_prob/100.0
```
## Climate Emulator - Simulation
```
# --------------------------------------
# Climate Emulator extremes model fitting
# Load Climate Emulator
CE = Climate_Emulator(db.paths.site.EXTREMES.climate_emulator)
CE.Load()
# set a new path for S6 simulations
# waves associated to future ENSO probability (no TCs) already simulated but copied manually from:
# ROI_CC_S5/EXTREMES/climate_emulator/Simulations/WAVES_noTCs
CE.Set_Simulation_Folder(p_S6_CE_sims, copy_WAVES_noTCs = False)
# optional: list variables to override distribution to empirical
#CE.sim_icdf_empirical_override = ['sea_Hs_31',
# 'swell_1_Hs_1','swell_1_Tp_1',
# 'swell_1_Hs_2','swell_1_Tp_2',]
# set simulated waves min-max filter
CE.sim_waves_filter.update({
'hs': (0, 8),
'tp': (2, 25),
'ws': (0, 0.06),
})
# --------------------------------------
# Climate Emulator simulation
# each DWT series will generate a different set of waves
for n in DWTs_sim.n_sim:
print('- Sim: {0} -'.format(int(n)+1))
# Select DWTs simulation
DWTs = DWTs_sim.sel(n_sim=n)
# Load previously simulated waves (no TCs)
WVS_sim, _, _ = CE.LoadSim(n_sim=int(n))
# Simulate TCs and update simulated waves
TCs_sim, WVS_upd = CE.Simulate_TCs(DWTs, WVS_sim, TCs_params, TCs_RBFs, pchange_TCs, MU_WT, TAU_WT)
# store simulation data
CE.SaveSim(WVS_sim, TCs_sim, WVS_upd, int(n))
```
| github_jupyter |
```
%%html
<h1> Initial Setup</h1>
#The following packages must be installed after anaconda is installed. They are commented off here.
#!pip install jira
#!pip install numpy
#!pip install pandas
#!pip install xslwriter
#!pip install json
#!pip intsall datetime
#!pip install functools
from jira import JIRA
import numpy as np
import pandas as pd
import xlsxwriter
import json
from datetime import datetime
from datetime import timedelta
pd.set_option('display.max_columns', None)
pd.set_option('display.max_rows', None)
pd.set_option('display.show_dimensions', True)
json_data_file = domain = domain = username = password = None
cpath = "./jira.json"
while not json_data_file:
try:
json_data_file = open(cpath)
except FileNotFoundError:
cpath = input('Directory Path of jira.json: ')
cpath = cpath + '/jira.json'
data = json.load(json_data_file)
username = data['auth']['username']
password = data['auth']['password']
bugqueryadd = data['bugqueryadd']
epicqueryadd = data['epicqueryadd']
storyqueryadd = data['storyqueryadd']
domain = data['domain']
columns = data['columns']
fields = data['fields']
outfile = data['outfile']
#if not domain:
# domain = input("Jira Domain (e.g https://XXX:PPP/jira): ")
#Only username and password will be accepted outside of the file
if not username:
username = input("Username: ")
if not password:
password = getpass.getpass("Password: ")
def get_jira_client(domain, username, password):
options = {'server': domain}
return JIRA(options, basic_auth=(username, password))
writer = pd.ExcelWriter(outfile)
jira = get_jira_client(domain, username, password)
#Important dates/labels that set the baseline for this run
qtrStart = '2018-07-04'
qtrEnd = '2018-09-25'
qtrStartDate = pd.to_datetime(qtrStart, format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
qtrEndDate = pd.to_datetime(qtrEnd, format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
relp = 'R18'
reln = 'R19'
#for relp:
#last deadline for inserting stories
releaseStoryDeadline = datetime(2018, 7, 24)
#monitoring deadline for removing stories
releaseStoryRemovalMonitoringStart = datetime(2018, 6, 12)
releaseStoryRemovalMonitoringEnd = datetime(2018, 9, 11)
bins = [datetime(2018, 6, 19), datetime(2018, 7, 3), datetime(2018, 7, 17),
datetime(2018, 7, 31), datetime(2018, 8, 14), datetime(2018, 8, 28), datetime(2018, 9, 11),
datetime(2018, 9, 25), datetime(2018, 10, 9)]
binLabels = ['reg-sprint-24', 'r18-sprint-25', 'r18-sprint-26',
'r18-sprint-27', 'reg-sprint-28',
'reg-sprint-29', 'r19-sprint-30', 'r19-sprint-31']
epics = jira.search_issues('type=epic and ' + epicqueryadd, json_result=True, maxResults=20000, fields = fields)
stories = jira.search_issues('type=story and ' + storyqueryadd, json_result=True, maxResults=20000, fields = fields, expand='changelog')
#prep the stories and epics dataframes
#fix the column names
#extract comment data
#extract all the history from stories and build all the workflow fields
for issue in stories['issues']:
#merge the textual fields of comments, summary
alltext = [comment['body'] for comment in issue['fields']['comment']['comments']]
if (issue['fields']['summary'] != None):
alltext.append(issue['fields']['summary'])
if (issue['fields']['description'] != None):
alltext.append(issue['fields']['description'])
try:
issue['fields']['textinfo'] = ' '.join(alltext)
except TypeError:
print(alltext)
#for stories only, record the important parts of change log as separate columns
issue['fields']['Open Set By'] = []
issue['fields']['Approval Set By'] = []
issue['fields']['Closed Set By'] = []
issue['fields']['Code Review Set By'] = []
issue['fields']['In Analysis Set By'] = []
issue['fields']['In Progress Set By'] = []
issue['fields']['In UI/UX Set By'] = []
issue['fields']['Ready for Estimation Set By'] = []
issue['fields']['Testing Set By'] = []
issue['fields']['Resolved Set By'] = []
issue['fields']['Reopened Set By'] = []
changelog = issue['changelog']
for history in changelog['histories']:
for item in history['items']:
#print (item['field'])
if (item['field'] == 'Fix Version') and (item['fromString'] == relp):
#a story was moved out of the current fix version?
issue['fields']['FixVersion Change Date'] = pd.to_datetime(history['created'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
#print(issue['key'], ' fix version changed from ', item['fromString'], ' to ', item['toString'])
if item['field'] == 'status':
#need to ensure if there are multiple times a certain status is updated, we capture it
#the first or last time based on the specific status.
timestamp = pd.to_datetime(history['created'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
event = item['toString'] + ' ' + 'Set By'
author = history['author']['name']
issue['fields'][event].append((author, timestamp))
#issue['fields'][item['toString'] + ' ' + 'Set To Date'] = history['created']
#issue['fields'][item['toString'] + ' ' + 'Set By'] = history['author']['name']
issue['fields']['Open Set By'] = min(issue['fields']['Open Set By'], key = lambda t: t[1]) if issue['fields']['Open Set By'] else None
issue['fields']['Approval Set By'] = max(issue['fields']['Approval Set By'], key = lambda t: t[1]) if issue['fields']['Approval Set By'] else None
issue['fields']['Closed Set By'] = max(issue['fields']['Closed Set By'], key = lambda t: t[1]) if issue['fields']['Closed Set By'] else None
issue['fields']['Code Review Set By'] = min(issue['fields']['Code Review Set By'], key = lambda t: t[1]) if issue['fields']['Code Review Set By'] else None
issue['fields']['In Analysis Set By'] = min(issue['fields']['In Analysis Set By'], key = lambda t: t[1]) if issue['fields']['In Analysis Set By'] else None
issue['fields']['In Progress Set By'] = min(issue['fields']['In Progress Set By'], key = lambda t: t[1]) if issue['fields']['In Progress Set By'] else None
issue['fields']['In UI/UX Set By'] = min(issue['fields']['In UI/UX Set By'], key = lambda t: t[1]) if issue['fields']['In UI/UX Set By'] else None
issue['fields']['Ready for Estimation Set By'] = min(issue['fields']['Ready for Estimation Set By'], key = lambda t: t[1]) if issue['fields']['Ready for Estimation Set By'] else None
issue['fields']['Testing Set By'] = min(issue['fields']['Testing Set By'], key = lambda t: t[1]) if issue['fields']['Testing Set By'] else None
issue['fields']['Resolved Set By'] = min(issue['fields']['Resolved Set By'], key = lambda t: t[1]) if issue['fields']['Resolved Set By'] else None
issue['fields']['Reopened Set By'] = min(issue['fields']['Reopened Set By'], key = lambda t: t[1]) if issue['fields']['Reopened Set By'] else None
issue['fields']['Open Set To Date'] = issue['fields']['Open Set By'][1] if issue['fields']['Open Set By'] else None
issue['fields']['Open Set By'] = issue['fields']['Open Set By'][0] if issue['fields']['Open Set By'] else None
issue['fields']['Approval Set To Date'] = issue['fields']['Approval Set By'][1] if issue['fields']['Approval Set By'] else None
issue['fields']['Approval Set By'] = issue['fields']['Approval Set By'][0] if issue['fields']['Approval Set By'] else None
issue['fields']['Closed Set To Date'] = issue['fields']['Closed Set By'][1] if issue['fields']['Closed Set By'] else None
issue['fields']['Closed Set By'] = issue['fields']['Closed Set By'][0] if issue['fields']['Closed Set By'] else None
issue['fields']['Code Review Set To Date'] = issue['fields']['Code Review Set By'][1] if issue['fields']['Code Review Set By'] else None
issue['fields']['Code Review Set By'] = issue['fields']['Code Review Set By'][0] if issue['fields']['Code Review Set By'] else None
issue['fields']['In Analysis Set To Date'] = issue['fields']['In Analysis Set By'][1] if issue['fields']['In Analysis Set By'] else None
issue['fields']['In Analysis Set By'] = issue['fields']['In Analysis Set By'][0] if issue['fields']['In Analysis Set By'] else None
issue['fields']['In Progress Set To Date'] = issue['fields']['In Progress Set By'][1] if issue['fields']['In Progress Set By'] else None
issue['fields']['In Progress Set By'] = issue['fields']['In Progress Set By'][0] if issue['fields']['In Progress Set By'] else None
issue['fields']['In UI/UX Set To Date'] = issue['fields']['In UI/UX Set By'][1] if issue['fields']['In UI/UX Set By'] else None
issue['fields']['In UI/UX Set By'] = issue['fields']['In UI/UX Set By'][0] if issue['fields']['In UI/UX Set By'] else None
issue['fields']['Ready for Estimation Set To Date'] = issue['fields']['Ready for Estimation Set By'][1] if issue['fields']['Ready for Estimation Set By'] else None
issue['fields']['Ready for Estimation Set By'] = issue['fields']['Ready for Estimation Set By'][0] if issue['fields']['Ready for Estimation Set By'] else None
issue['fields']['Testing Set To Date'] = issue['fields']['Testing Set By'][1] if issue['fields']['Testing Set By'] else None
issue['fields']['Testing Set By'] = issue['fields']['Testing Set By'][0] if issue['fields']['Testing Set By'] else None
issue['fields']['Resolved Set To Date'] = issue['fields']['Resolved Set By'][1] if issue['fields']['Resolved Set By'] else None
issue['fields']['Resolved Set By'] = issue['fields']['Resolved Set By'][0] if issue['fields']['Resolved Set By'] else None
issue['fields']['Reopened Set To Date'] = issue['fields']['Reopened Set By'][1] if issue['fields']['Reopened Set By'] else None
issue['fields']['Reopened Set By'] = issue['fields']['Reopened Set By'][0] if issue['fields']['Reopened Set By'] else None
for issue in epics['issues']:
alltext = [comment['body'] for comment in issue['fields']['comment']['comments']]
alltext.append(issue['fields']['summary'])
#alltext.append(issue['fields']['description'])
issue['fields']['textinfo'] = ' '.join(alltext)
epic_list = []
for epic in epics['issues']:
epic['fields']['key'] = epic['key']
epic_list.append(epic['fields'])
epics_df = pd.DataFrame(epic_list)
story_list = []
for story in stories['issues']:
story['fields']['key'] = story['key']
story_list.append(story['fields'])
stories_df = pd.DataFrame(story_list)
#replacement of custom field's by their names is only done inside the dataframe
# Fetch all fields
allfields=jira.fields()
# Make a map from field name -> field id
nameMap = {field['name']:field['id'] for field in allfields}
idMap = {field['id']:field['name'] for field in allfields}
for column in epics_df.columns:
if ('custom' in column):
epics_df.rename(columns={column: idMap[column]}, inplace=True)
for column in stories_df.columns:
if ('custom' in column):
stories_df.rename(columns={column: idMap[column]}, inplace=True)
stories_df['Team'] = stories_df['Team'].apply(lambda x: x[0].get('value') if (type(x) == list) else None)
stories_df['status'] = stories_df['status'].apply(lambda x: x.get('name'))
stories_df['reporter'] = stories_df['reporter'].apply(lambda x: x.get('name'))
stories_df['fixVersions'] = stories_df['fixVersions'].apply(lambda x: x[0]['name'] if ((type(x) == list) and x and (type(x[0]) == dict)) else None)
stories_df['Platform'] = stories_df['Platform'].apply(lambda x: x[0].get('value'))
stories_df['created'] = pd.to_datetime(stories_df['created'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
stories_df['resolution'] = stories_df['resolution'].apply(lambda x: x['name'] if type(x) == dict else None)
#insert a column for jira link
stories_df['story_link'] = '=HYPERLINK("' + domain + '/browse/' + stories_df['key'] + '","' + stories_df['key'] + '")'
#eliminate stories that are marked not needed
stories_df = stories_df[stories_df['resolution'] != 'Not Needed']
#extract the sprint information from the sprints field and create a separate sprints-issue dataframe
#this is only possible once we have the stories dataframe
from functools import reduce
#Takes a list of sprints of the form:
#['com.atlassian.greenhopper.service.sprint.Sprint@1b7eb58a[id=519,rapidViewId=219,state=CLOSED,name=Knight Riders Sprint 2018 - 22,startDate=2018-05-23T21:16:06.149+05:30,endDate=2018-06-05T19:44:00.000+05:30,completeDate=2018-06-06T20:45:27.547+05:30,sequence=519]',
# 'com.atlassian.greenhopper.service.sprint.Sprint@2a28663d[id=542,rapidViewId=219,state=ACTIVE,name=Knight Riders Sprint 2018-23,startDate=2018-06-06T22:14:10.412+05:30,endDate=2018-06-19T20:42:00.000+05:30,completeDate=<null>,sequence=542]']
# and returns one list with a dictionary object for each sprint located. The object also contains the issue key
# the other is
# we return a dictionary
def getSprintInfo(issueKey, sprint):
#locate the part in square braces
start = sprint.find('[') + 1
end = sprint.find(']', start)
dict_sprint = dict(x.split('=') for x in sprint[start:end].split(','))
dict_sprint['issue_key'] = issueKey
return dict_sprint
#we return a list of dictionaries, where each dictionary is a sprint paired with the issue.
def getSprints (issueKey, sprints):
if type(sprints) == list:
return [getSprintInfo(issueKey, sprint) for sprint in sprints]
else:
return []
x1 = []
for index, row in stories_df.iterrows():
if row['Sprint']:
x1 = x1 + (getSprints(row['key'], row['Sprint']))
sprints_df = pd.DataFrame(x1)
sprints_df['endDate'] = pd.to_datetime(sprints_df['endDate'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
sprints_df['startDate'] = pd.to_datetime(sprints_df['startDate'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
sprints_df['completeDate'] = pd.to_datetime(sprints_df['completeDate'], format='%Y-%m-%dT%H:%M:%S.%f', errors='coerce')
#Limit all stories/sprints to this quarter.
sprints_df = sprints_df[(sprints_df['endDate'] >= qtrStartDate) & (sprints_df['startDate'] < qtrEndDate)]
#calcuate the age of the stories in the last state it is in
now = datetime.now() + pd.Timedelta('010:30:00')
stories_df['Age In Days'] = stories_df.apply(lambda x: (now - x[x['status'] + ' Set To Date']).days, axis = 1)
%%html
<h1> Base Query Statistics</h1>
<h3>Impact: None</h3>
<h4>Action: None</h4>
#Basic statistics before we start separating
print('No. Epics: ', epics_df['key'].unique().size)
print('No. Stories: ', stories_df['key'].unique().size)
print('No. Sprints: ', sprints_df['name'].unique().size)
print('No of stories without linked epics: ', sum(pd.isnull(stories_df['Epic Link'])))
print ('Stories not Closed: ', stories_df[stories_df['status'] != 'Closed']['key'].unique().size)
print ('Stories without a fixVersion: ', stories_df[stories_df['fixVersions'] == None]['key'].unique().size)
storiesFixVersionsStatus_df = stories_df[['fixVersions', 'status', 'key']].copy()
storiesFixVersionsStatus_df.groupby(['fixVersions', 'status']).agg(['count'])
%%html
<h1>Fix Version Changed Changed after deadline for current release </h1>
<h3>Impact: These are release scope changes that disrupt planning</h3>
<h4>Action: Prevent Future Scope changes</h4>
stories_df[pd.notnull(stories_df['FixVersion Change Date']) & (stories_df['FixVersion Change Date'] > releaseStoryRemovalMonitoringStart) & (stories_df['fixVersions'] != relp)][['key', 'Team', 'reporter', 'summary', 'FixVersion Change Date', 'fixVersions']].sort_values('Team')
%%html
<h1>Stories given wrong fixVersions</h1>
<h3>Impact: The codebase has been changed, yet the fix version is NOT the current release!</h3>
<h4>Action: The fixVersions for these stories must be set to current release</h4>
#Stories that in Code Review/Testing or Approval in reln need to be flagged
df = stories_df[((stories_df['fixVersions'].isin([reln, 'Backlog']) | pd.isnull(stories_df['fixVersions'])) &
(stories_df['status'].isin(['Code Review', 'In Progress', 'Approval', 'Closed'])) )]
df[['key', 'status', 'fixVersions', 'summary']]
#first merge - create the epics and stories merge
scope_df = pd.merge(epics_df, stories_df, how='right', on=None, left_on='key', right_on='Epic Link',
left_index=False, right_index=False, sort=True,
suffixes=('_epic', '_story'), copy=True, indicator=False,
validate=None)
#Combine the sprints with the epics + stories dataframe and we can then drop the duplicate issue_key field.
sprintsWithStoriesAndEpics_df = pd.merge(scope_df, sprints_df, how='left', on=None, left_on='key_story', right_on='issue_key',
left_index=False, right_index=False,
suffixes=('_story', '_sprint'),
copy=True, indicator=True,
validate=None).drop(columns = ['issue_key'])
#We can drop stories that are in future sprints
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[sprintsWithStoriesAndEpics_df['state'] != 'FUTURE']
%%html
<h1>Stories violating Sprint or Release Commitments</h1>
<h3>Impact: These stories were inserted after sprints started or after release deadline and hence they disrupt planning</h3>
<h4>Action: Prevent Future Scope changes</h4>
<h5>Note:Upto 24 hour grace period has been allowed for marking stories open
#find the stories which were opened more than a day later than the sprint started
#or those were inserted after the development sprints were over
sprintsWithStoriesAndEpics_dfCopy = sprintsWithStoriesAndEpics_df[pd.notnull(sprintsWithStoriesAndEpics_df['startDate'])]
sprintsWithStoriesAndEpics_dfCopy = sprintsWithStoriesAndEpics_dfCopy[['Team_story', 'startDate', 'state', 'Open Set To Date', 'reporter_story', 'Story Points', 'key_story', 'name', 'fixVersions_story']].copy()
sprintsWithStoriesAndEpics_dfCopy['sprintLeadTime'] = (sprintsWithStoriesAndEpics_dfCopy['Open Set To Date'] - sprintsWithStoriesAndEpics_dfCopy['startDate']).dt.days
sprintsWithStoriesAndEpics_dfCopy['sprintCommitment'] = sprintsWithStoriesAndEpics_dfCopy['sprintLeadTime'] <= 1
#sprintsWithStoriesAndEpics_dfCopy['key_story'].unique().size
sprintsWithStoriesAndEpics_dfCopy['beyondReleaseDeadline'] = sprintsWithStoriesAndEpics_dfCopy['Open Set To Date'] >= releaseStoryDeadline
df = sprintsWithStoriesAndEpics_dfCopy[(sprintsWithStoriesAndEpics_dfCopy['sprintCommitment'] != True)|(sprintsWithStoriesAndEpics_dfCopy['beyondReleaseDeadline'] == True)].sort_values(by='key_story')
#df = sprintsWithStoriesAndEpics_dfCopy
#df = df[df['state'] == 'ACTIVE']
#write out the source data onto disk
#however we want to write only the records which are duplicates. Better idea to remove the non duplicates.
df.to_excel(writer, index=False, sheet_name='Late Commitments', freeze_panes=(1,0), columns=['Team_story', 'startDate', 'Open Set To Date', 'reporter_story', 'Story Points', 'key_story', 'name', 'sprintLeadTime', 'sprintCommitment'])
df.sort_values(['startDate', 'Team_story'], ascending = False)
%%html
<h1>Stories that took too long in Analysis or Development or QA or Approvals</h1>
<h3>Impact: These stories may need analysis on why they took longer than threshold in either the Dev/QA/Prod buckets</h3>
<h4>Action: Find preventive strategies for future</h4>
#For stories that are closed, lets find the time it took for us to go through each state completely,
#the points of the story, the number of sprints it took, the team the story is in.
#We are ignoring the Reopen workflow.
sprintsWithStoriesAndEpics_df['Analysis Duration'] = (sprintsWithStoriesAndEpics_df['Ready for Estimation Set To Date'] - sprintsWithStoriesAndEpics_df['created_story']).dt.days
sprintsWithStoriesAndEpics_df['Dev Duration'] = (sprintsWithStoriesAndEpics_df['Testing Set To Date'] - sprintsWithStoriesAndEpics_df['Open Set To Date']).dt.days
sprintsWithStoriesAndEpics_df['QA Duration'] = (sprintsWithStoriesAndEpics_df['Approval Set To Date'] - sprintsWithStoriesAndEpics_df['Testing Set To Date']).dt.days
sprintsWithStoriesAndEpics_df['Approval Duration'] = (sprintsWithStoriesAndEpics_df['Closed Set To Date'] - sprintsWithStoriesAndEpics_df['Approval Set To Date']).dt.days
df = sprintsWithStoriesAndEpics_df
df = df[(df['Analysis Duration'] > 60) | (df['Dev Duration'] > 7) | (df['QA Duration'] > 2) | (df['Approval Duration'] > 1)]
df[['key_story', 'fixVersions_story', 'status_story', 'state', 'Team_story', 'Analysis Duration', 'Dev Duration', 'QA Duration', 'Approval Duration']].sort_values(['state', 'Approval Duration', 'QA Duration', 'Dev Duration'], ascending = False)
%%html
<h1>Current Releases Analysis</h1>
#Lets remove the stories which we do not care about - not in relp or reln
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[((sprintsWithStoriesAndEpics_df['fixVersions_story'] == relp) | (sprintsWithStoriesAndEpics_df['fixVersions_story'] == reln))]
#Lets remove the stories which we do not care about - closed
stories_df = stories_df[stories_df['status'] != "Closed"]
scope_df = scope_df[scope_df['status_story'] != "Closed"]
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[sprintsWithStoriesAndEpics_df['status_story'] != "Closed"]
#select the latest sprint that the stories are in and then we can filter the ones that sprints that are closed.
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df.loc[sprintsWithStoriesAndEpics_df.groupby("key_story")["startDate"].idxmax()]
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[pd.notnull(sprintsWithStoriesAndEpics_df.index)]
#only after the above is done, we can filter the stories that have their latest sprints closed
%%html
<h1> Stories in current releases that are not yet assigned to sprints or in inactive sprints</h1>
<h3>Impact: These stories are in current/next release and not yet assigned to a sprint or are in inactive sprints</h3>
<h4>Action: These may to be fixed</h4>
sprintsWithStoriesAndEpics_df[(sprintsWithStoriesAndEpics_df['_merge'] == 'left_only') |
(sprintsWithStoriesAndEpics_df['state'] == 'CLOSED')][
['key_story', 'fixVersions_story', 'reporter_story', 'summary_story', 'status_story', 'name']].sort_values(
['fixVersions_story'])
#eliminiate the stories that are not assigned to sprints.
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[sprintsWithStoriesAndEpics_df['_merge'] != 'left_only']
#eliminate the stories with recent inactive sprints
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[sprintsWithStoriesAndEpics_df['state'] != 'CLOSED']
%%html
<h1> Stories that violate the Age criteria</h1>
<h3>Impact: These stories have been sitting in their current state for too long</h3>
<h4>Action: Needs attention from scrum masters or product or devops</h4>
#Calculate sprint Age.
#sprintsWithStoriesAndEpics_df['key_story'][165]
#There are two scenarios we have not considered - if the sprint is not active anymore, the age should be zero
#We can remove stories in inactive sprints and report them as having no sprints!
#The second case if the last status change happened earlier than sprint start date.
sprintsWithStoriesAndEpics_df['Sprint Age In Days'] = sprintsWithStoriesAndEpics_df.apply(lambda x: (now - max(x[x['status_story'] + ' Set To Date'], x['startDate'])).days, axis = 1)
#List the stories with their status, age and sprint age.
sprintsWithStoriesAndEpics_df = sprintsWithStoriesAndEpics_df[(sprintsWithStoriesAndEpics_df['Sprint Age In Days'] > 3) | (sprintsWithStoriesAndEpics_df['Age In Days'] > 3)]
sprintsWithStoriesAndEpics_df[['key_story', 'Team_story', 'fixVersions_story', 'summary_story', 'status_story', 'Age In Days', 'Sprint Age In Days', 'Open Set To Date']].sort_values(by=['Age In Days', 'Sprint Age In Days'], ascending = False)
%%html
<h1> Calculate the Stories not having any mention of AC or Acceptance.</h1>
#this is a list of strings
#scope_df['textinfo'] = scope_df['textinfo_story'] + scope_df['textinfo_epic']
scope_df['textinfo'] = scope_df['textinfo_story']
scope_df['Invalid AC'] = scope_df['textinfo'].str.contains('Acceptance|AC', case = False, regex = True) == False
#write out the source data onto disk
#however we want to write only the records which are duplicates. Better idea to remove the non duplicates.
scope_df[scope_df['Invalid AC']].to_excel(writer, index=False, sheet_name='Invalid AC', freeze_panes=(1,0), columns=['Team_story', 'key_story', 'reporter_story'])
invalid_ac_df = scope_df[['reporter_story', 'Invalid AC']].copy()
#produce statistics for valid/invalid AC
invalid_ac_df.groupby(['reporter_story']).sum().sort_values(by=['Invalid AC'], ascending=False).head()
writer.save()
```
| github_jupyter |
# Extract electricity prices from VENRON data set
# Introduction
In this notebook we use `Fonduer` to extract relations from the `VENRON` dataset.
This code is a modified version of their original hardware [tutorial](https://github.com/HazyResearch/fonduer-tutorials/tree/master/hardware).
The `Fonduer` pipeline (as outlined in the [paper](https://arxiv.org/abs/1703.05028)), and the iterative KBC process:
1. KBC Initialization
2. Candidate Generation and Multimodal Featurization
3. Probabilistic Relation Classification
4. Error Analysis and Iterative KBC
## Setup
First we import the relevant libraries and connect to the local database.
Follow the README instructions to setup the connection to the postgres DB correctly.
If the database has existing candidates with generated features, the will not be overriden.
To re-run the entire pipeline including initialization drop the database first.
```
! dropdb -h postgres -h postgres --if-exists elec_price_vol
! createdb -h postgres -h postgres elec_price_vol
# source .venv/bin/activate
%load_ext autoreload
%autoreload 2
%matplotlib inline
import os
import sys
import logging
PARALLEL = 8 # 4 # assuming a quad-core machine
ATTRIBUTE = "elec_price_vol"
DB_USERNAME = 'user'
DB_PASSWORD = 'venron'
conn_string = f'postgresql://{DB_USERNAME}:{DB_PASSWORD}@postgres:5432/{ATTRIBUTE}'
dataset = 'gold' # 'full'
docs_path = f'data/{dataset}/html/'
pdf_path = 'data/pdf/'
gold_file = 'data/electricity_gold.csv'
max_docs = 10 # 114
```
## 1.1 Parsing and Transforming the Input Documents into Unified Data Models
We first initialize a `Meta` object, which manages the connection to the database automatically, and enables us to save intermediate results.
```
from fonduer import Meta, init_logging
# Configure logging for Fonduer
init_logging(log_dir="logs", level=logging.INFO) # DEBUG LOGGING
session = Meta.init(conn_string).Session()
from fonduer.parser.preprocessors import HTMLDocPreprocessor
from fonduer.parser.models import Document, Sentence
from fonduer.parser import Parser
has_documents = session.query(Document).count() > 0
corpus_parser = Parser(session, structural=True, lingual=True, visual=True, pdf_path=pdf_path)
if (not has_documents):
doc_preprocessor = HTMLDocPreprocessor(docs_path, max_docs=max_docs)
%time corpus_parser.apply(doc_preprocessor, parallelism=PARALLEL)
print(f"Documents: {session.query(Document).count()}")
print(f"Sentences: {session.query(Sentence).count()}")
# Initialize NLP library for vector similarities
import sys
!{sys.executable} -m spacy download en_core_web_lg
```
## 1.2 Dividing the Corpus into Test and Train
We'll split the documents 80/10/10 into train/dev/test splits. Note that here we do this in a non-random order to preserve the consistency and we reference the splits by 0/1/2 respectively.
```
docs = session.query(Document).order_by(Document.name).all()
ld = len(docs)
train_docs = set()
dev_docs = set()
test_docs = set()
splits = (0.8, 0.9)
data = [(doc.name, doc) for doc in docs]
data.sort(key=lambda x: x[0])
for i, (doc_name, doc) in enumerate(data):
if i < splits[0] * ld:
train_docs.add(doc)
elif i < splits[1] * ld:
dev_docs.add(doc)
else:
test_docs.add(doc)
from pprint import pprint
pprint([x.name for x in train_docs][0:5])
print(f"Number of documents split: {len(docs)}")
```
# Phase 2: Mention Extraction, Candidate Extraction Multimodal Featurization
Given the unified data model from Phase 1, `Fonduer` extracts relation
candidates based on user-provided **matchers** and **throttlers**. Then,
`Fonduer` leverages the multimodality information captured in the unified data
model to provide multimodal features for each candidate.
## 2.1 Mention Extraction & Candidate Generation
1. Define mention classes
2. Use matcher functions to define the format of potential mentions
3. Define Mentionspaces (Ngrams)
4. Run Mention extraction (all possible ngrams in the document, API [ReadTheDocs](https://fonduer.readthedocs.io/en/stable/user/candidates.html#fonduer.candidates.MentionExtractor))
```
from fonduer.candidates import MentionExtractor
from fonduer.candidates.models import Mention
from my_subclasses import mention_classes, mention_spaces, matchers
hasMentions = session.query(Mention).count() > 0
if (not hasMentions):
# 4.) Mention extraction
mention_extractor = MentionExtractor(
session, mention_classes, mention_spaces, matchers
)
docs = session.query(Document).order_by(Document.name).all()
mention_extractor.apply(docs, parallelism=PARALLEL)
mentions = session.query(Mention).all()
print(f"Total Mentions: {len(mentions)}")
from fonduer_utils import prune_duplicate_mentions
from fonduer.utils.data_model_utils import get_ancestor_tag_names
Station = mention_classes[0]
station_throttler = lambda s: ('head' in get_ancestor_tag_names(s) and 'title' in get_ancestor_tag_names(s))
# Performance increase (reduce quadratic candidates combination by deleting duplicate mentions)
mentions = prune_duplicate_mentions(session, mentions, Station, station_throttler)
# DEBUG: Test if at least one station mention is for meadmktplace types
list([x for x in mentions if x.document.name.upper() == "11_NP 15 PAGES" and isinstance(x, Station)])
```
## 2.2 Candidate Extraction
1. Define Candidate Class
2. Define trottlers to reduce the number of possible candidates
3. Extract candidates (View the API for the CandidateExtractor on [ReadTheDocs](https://fonduer.readthedocs.io/en/stable/user/candidates.html#fonduer.candidates.MentionExtractor).)
In the last part we specified that these `Candidates` belong to the training set by specifying `split=0`; recall that we're referring to train/dev/test as splits 0/1/2.
```
import re
from my_subclasses import candidate_classes, throttlers
from fonduer.candidates import CandidateExtractor
from fonduer.utils.visualizer import Visualizer
# 1.) Define Candidate class
StationPrice = candidate_classes[0]
has_candidates = session.query(StationPrice).filter(StationPrice.split == 0).count() > 0
# 2.) Candidate extraction
# NOTE: Without nested_relations flag DocumentMentions and FigureMentions are filtered out.
# Otherwise they would require a rewrite of the featurizers, due to preprocessing we duplicate the img-url and doc-name
candidate_extractor = CandidateExtractor(session, [StationPrice], throttlers=throttlers) # , nested_relations=True)
for i, docs in enumerate([train_docs, dev_docs, test_docs]):
if (not has_candidates):
candidate_extractor.apply(docs, split=i, parallelism=PARALLEL)
print(f"Number of Candidates in split={i}: {session.query(StationPrice).filter(StationPrice.split == i).count()}")
train_cands = candidate_extractor.get_candidates(split = 0)
dev_cands = candidate_extractor.get_candidates(split = 1)
test_cands = candidate_extractor.get_candidates(split = 2)
cands = [train_cands, dev_cands, test_cands]
# 3.) Visualize some candidate for error analysis
# pprint(train_cands[0][0])
# vis = Visualizer(pdf_path)
# Display a candidate
# vis.display_candidates([train_cands[0][0]])
```
## 2.2 Multimodal Featurization
Unlike dealing with plain unstructured text, `Fonduer` deals with richly formatted data, and consequently featurizes each candidate with a baseline library of multimodal features.
### Featurize with `Fonduer`'s optimized Postgres Featurizer
We now annotate the candidates in our training, dev, and test sets with features. The `Featurizer` provided by `Fonduer` allows this to be done in parallel to improve performance.
View the API provided by the `Featurizer` on [ReadTheDocs](https://fonduer.readthedocs.io/en/stable/user/features.html#fonduer.features.Featurizer).
At the end of this phase, `Fonduer` has generated the set of candidates and the feature matrix. Note that Phase 1 and 2 are relatively static and typically are only executed once during the KBC process.
```
from fonduer.features import Featurizer
from fonduer.features.models import Feature
from fonduer.features.feature_extractors import FeatureExtractor
featurizer = Featurizer(
session,
[StationPrice],
feature_extractors=FeatureExtractor(["textual", "structural", "tabular", "visual"])
)
has_features = session.query(Feature).count() > 0
if (not has_features):
# Training set
%time featurizer.apply(split=0, train=True, parallelism=PARALLEL)
%time F_train = featurizer.get_feature_matrices(train_cands)
print(F_train[0].shape)
# Dev set
%time featurizer.apply(split=1, parallelism=PARALLEL)
%time F_dev = featurizer.get_feature_matrices(dev_cands)
print(F_dev[0].shape)
# Test set
%time featurizer.apply(split=2, parallelism=PARALLEL)
%time F_test = featurizer.get_feature_matrices(test_cands)
print(F_test[0].shape)
else:
%time F_train = featurizer.get_feature_matrices(train_cands)
%time F_dev = featurizer.get_feature_matrices(dev_cands)
%time F_test = featurizer.get_feature_matrices(test_cands)
F = [F_train, F_dev, F_test]
```
# Phase 3: Probabilistic Relation Classification
In this phase, `Fonduer` applies user-defined **labeling functions**, which express various heuristics, patterns, and [weak supervision](http://hazyresearch.github.io/snorkel/blog/weak_supervision.html) strategies to label our data, to each of the candidates to create a label matrix that is used by our data programming engine.
1. Load Gold Data
---
Iterate the following steps
2. Create labeling functions
3. Apply labeling functions and measure accuracy of each LF (based on gold data).
4. Build a generative model by combining the labeling functions
5. Iterate on labeling function based on the models score
---
6. Finally build a descriminative model and test on the test set
### 3.1) Loading Gold LF
```
from fonduer.supervision.models import GoldLabel
from electricity_utils import get_gold_func
from fonduer.supervision import Labeler
from my_subclasses import stations_mapping_dict
# 1.) Load the gold data
gold = get_gold_func(gold_file, attribute=ATTRIBUTE, stations_mapping_dict=stations_mapping_dict)
docs = corpus_parser.get_documents()
labeler = Labeler(session, [StationPrice])
%time labeler.apply(docs=docs, lfs=[[gold]], table=GoldLabel, train=True, parallelism=PARALLEL)
```
### 3.2) Creating Labeling Functions
We have 3 states that we can return from a LF: `ABSTAIN`, `FALSE` or `TRUE`.
A library of data model utilities
which can be used to write labeling functions are outline in [Read the
Docs](http://fonduer.readthedocs.io/en/stable/user/data_model_utils.html).
### 3.3) Applying the Labeling Functions
Next, we need to actually run the LFs over all of our training candidates, producing a set of `Labels` and `LabelKeys` (just the names of the LFs) in the database. Note that this will delete any existing `Labels` and `LabelKeys` for this candidate set.
View the API provided by the `Labeler` on [ReadTheDocs](https://fonduer.readthedocs.io/en/stable/user/supervision.html#fonduer.supervision.Labeler).
We can also view statistics about the resulting label matrix.
* **Coverage** is the fraction of candidates that the labeling function emits a non-zero label for.
* **Overlap** is the fraction candidates that the labeling function emits a non-zero label for and that another labeling function emits a non-zero label for.
* **Conflict** is the fraction candidates that the labeling function emits a non-zero label for and that another labeling function emits a conflicting non-zero label for.
In addition, because we have already loaded the gold labels, we can view the emperical accuracy of these labeling functions when compared to our gold labels using the `analysis` module of [Snorkel](https://github.com/snorkel-team/snorkel)
### 3.4) Build Generative Model
Now, we'll train a model of the LFs to estimate their accuracies. Once the model is trained, we can combine the outputs of the LFs into a single, noise-aware training label set for our extractor. Intuitively, we'll model the LFs by observing how they overlap and conflict with each other. To do so, we use [Snorkel](https://github.com/snorkel-team/snorkel)'s single-task label model.
We then print out the marginal probabilities for each training candidate.
```
from fonduer.utils.data_model_utils import *
from electricity_utils import eval_LFs
from snorkel.labeling import labeling_function
from snorkel.labeling import LFAnalysis
from snorkel.labeling.model import LabelModel
from fonduer_utils import get_applied_lfs, get_neighbor_cell_ngrams_own, _min_range_diff, min_row_diff, min_col_diff
import matplotlib.pyplot as plt
import re
def run_labeling_functions():
ABSTAIN = -1
FALSE = 0
TRUE = 1
@labeling_function()
def LF_other_station_table(c):
station_span = c.station.context.get_span().lower()
neighbour_cells = get_neighbor_cell_ngrams_own(c.price, dist=100, directions=True, n_max = 4, absolute = True)
up_cells = [x for x in neighbour_cells if len(x) > 1 and x[1] == 'DOWN' and x[0] in stations_list]
# No station name in upper cells
if (len(up_cells) == 0):
return ABSTAIN
# Check if the next upper aligned station-span corresponds to the candidate span (or equivalents)
closest_header = up_cells[len(up_cells)-1]
return TRUE if closest_header[0] in stations_mapping_dict[station_span] else FALSE
@labeling_function()
def LF_station_non_meta_tag(c):
html_tags = get_ancestor_tag_names(c.station)
return FALSE if ('head' in html_tags and 'title' in html_tags) else ABSTAIN
# Basic constraint for the price LFs to be true -> no wrong station (increase accuracy)
def base(c):
return (
LF_station_non_meta_tag(c) != 0 and
LF_other_station_table(c) != 0 and
LF_off_peak_head(c) != 0 and
LF_purchases(c)
)
# 2.) Create labeling functions
@labeling_function()
def LF_on_peak_head(c):
return TRUE if 'on peak' in get_aligned_ngrams(c.price, n_min=2, n_max=2) and base(c) else ABSTAIN
@labeling_function()
def LF_off_peak_head(c):
return FALSE if 'off peak' in get_aligned_ngrams(c.price, n_min=2, n_max=2) else ABSTAIN
@labeling_function()
def LF_price_range(c):
price = float(c.price.context.get_span())
return TRUE if price > 0 and price < 1000 and base(c) else FALSE
@labeling_function()
def LF_price_head(c):
return TRUE if 'price' in get_aligned_ngrams(c.price) and base(c) else ABSTAIN
@labeling_function()
def LF_firm_head(c):
return TRUE if 'firm' in get_aligned_ngrams(c.price)and base(c) else ABSTAIN
@labeling_function()
def LF_dollar_to_left(c):
return TRUE if '$' in get_left_ngrams(c.price, window=2) and base(c) else ABSTAIN
@labeling_function()
def LF_purchases(c):
return FALSE if 'purchases' in get_aligned_ngrams(c.price, n_min=1) else ABSTAIN
station_price_lfs = [
LF_other_station_table,
LF_station_non_meta_tag,
# indicator
LF_price_range,
# negative indicators
LF_off_peak_head,
LF_purchases,
# positive indicators
LF_on_peak_head,
LF_price_head,
LF_firm_head,
LF_dollar_to_left,
]
# 3.) Apply the LFs on the training set
labeler = Labeler(session, [StationPrice])
labeler.apply(split=0, lfs=[station_price_lfs], train=True, clear=True, parallelism=PARALLEL)
L_train = labeler.get_label_matrices(train_cands)
# Check that LFs are all applied (avoid crash)
applied_lfs = L_train[0].shape[1]
has_non_applied = applied_lfs != len(station_price_lfs)
print(f"Labeling functions on train_cands not ABSTAIN: {applied_lfs} (/{len(station_price_lfs)})")
if (has_non_applied):
applied_lfs = get_applied_lfs(session)
non_applied_lfs = [l.name for l in station_price_lfs if l.name not in applied_lfs]
print(f"Labling functions {non_applied_lfs} are not applied.")
station_price_lfs = [l for l in station_price_lfs if l.name in applied_lfs]
# 4.) Evaluate their accuracy
L_gold_train = labeler.get_gold_labels(train_cands, annotator='gold')
# Sort LFs for LFAnalysis because LFAnalysis does not sort LFs,
# while columns of L_train are sorted alphabetically already.
sorted_lfs = sorted(station_price_lfs, key=lambda lf: lf.name)
LFAnalysis(L=L_train[0], lfs=sorted_lfs).lf_summary(Y=L_gold_train[0].reshape(-1))
# 5.) Build generative model
gen_model = LabelModel(cardinality=2)
gen_model.fit(L_train[0], n_epochs=500, log_freq=100)
train_marginals_lfs = gen_model.predict_proba(L_train[0])
plt.hist(train_marginals_lfs[:, TRUE], bins=20)
plt.show()
# Apply on dev-set
labeler.apply(split=1, lfs=[station_price_lfs], clear=True, parallelism=PARALLEL)
L_dev = labeler.get_label_matrices(dev_cands)
L_gold_dev = labeler.get_gold_labels(dev_cands, annotator='gold')
LFAnalysis(L=L_dev[0], lfs=sorted_lfs).lf_summary(Y=L_gold_dev[0].reshape(-1))
return (gen_model, train_marginals_lfs)
if (dataset == 'full'):
(gen_model, train_marginals_lfs) = run_labeling_functions()
eval_LFs(train_marginals_lfs, train_cands, gold)
# # Query for analysis
# labels = session.query(Label).all()
# gold_labels = session.query(GoldLabel).all()
# gold_labels_map = { gold_label.candidate_id: gold_label for gold_label in gold_labels }
# from fonduer.candidates.models import Candidate
# DB_FALSE = FALSE +1
# DB_ABSTAIN = ABSTAIN +1
# DB_TRUE = TRUE +1
# def get_incorrect_instances(lf):
# def is_wrong_label(label):
# if (lf.name not in label.keys):
# return False # Abstain
# assigned_label = label.values[label.keys.index(lf.name)]
# gold_label = gold_labels_map[label.candidate_id] # [x for x in gold_labels if x.candidate_id == label.candidate_id][0]
# return gold_label.values[0] != assigned_label
# return [x.candidate for x in labels if is_wrong_label(x)]
# lf = station_price_lfs[5]
# wrong_cands = get_incorrect_instances(lf)
# pprint(f"Labeling Function: {lf.name} has wrongly labelled the candidate(1/{len(wrong_cands)}):")
# if (len(wrong_cands) > 0):
# wrong_cand = wrong_cands[100]
# pprint(wrong_cand)
# pprint('LF is True' if lf(wrong_cand) == 1 else 'LF is False')
# vis = Visualizer(pdf_path)
# # Display a candidate
# vis.display_candidates([wrong_cand])
# else:
# print("There are no wrong candidates for this labeling function")
```
## Training the Discriminative Model
Fonduer uses the machine learning framework [Emmental](https://github.com/SenWu/emmental) to support all model training.
```
import emmental
import numpy as np
from emmental.modules.embedding_module import EmbeddingModule
from emmental.data import EmmentalDataLoader
from emmental.model import EmmentalModel
from emmental.learner import EmmentalLearner
from fonduer.learning.utils import collect_word_counter
from fonduer.learning.dataset import FonduerDataset
from fonduer.learning.task import create_task
ABSTAIN = -1
FALSE = 0
TRUE = 1
def train_model(cands, F, train_marginals, model_type="LogisticRegression"):
# Extract candidates and features
train_cands = cands[0]
F_train = F[0]
# 1.) Setup training config
config = {
"meta_config": {"verbose": True},
"model_config": {"model_path": None, "device": 0, "dataparallel": False},
"learner_config": {
"n_epochs": 50,
"optimizer_config": {"lr": 0.001, "l2": 0.0},
"task_scheduler": "round_robin",
},
"logging_config": {
"evaluation_freq": 1,
"counter_unit": "epoch",
"checkpointing": False,
"checkpointer_config": {
"checkpoint_metric": {f"{ATTRIBUTE}/{ATTRIBUTE}/train/loss": "min"},
"checkpoint_freq": 1,
"checkpoint_runway": 2,
"clear_intermediate_checkpoints": True,
"clear_all_checkpoints": True,
},
},
}
emmental.init(Meta.log_path)
emmental.Meta.update_config(config=config)
# 2.) Collect word counter from training data
word_counter = collect_word_counter(train_cands)
# 3.) Generate word embedding module for LSTM model
# (in Logistic Regression, we generate it since Fonduer dataset requires word2id dict)
# Geneate special tokens
arity = 2
specials = []
for i in range(arity):
specials += [f"~~[[{i}", f"{i}]]~~"]
emb_layer = EmbeddingModule(
word_counter=word_counter, word_dim=300, specials=specials
)
# 4.) Generate dataloader for training set
# Filter out noise samples
diffs = train_marginals.max(axis=1) - train_marginals.min(axis=1)
train_idxs = np.where(diffs > 1e-6)[0]
train_dataloader = EmmentalDataLoader(
task_to_label_dict={ATTRIBUTE: "labels"},
dataset=FonduerDataset(
ATTRIBUTE,
train_cands[0],
F_train[0],
emb_layer.word2id,
train_marginals,
train_idxs,
),
split="train",
batch_size=100,
shuffle=True,
)
# 5.) Training
tasks = create_task(
ATTRIBUTE, 2, F_train[0].shape[1], 2, emb_layer, model=model_type # "LSTM"
)
model = EmmentalModel(name=f"{ATTRIBUTE}_task")
for task in tasks:
model.add_task(task)
emmental_learner = EmmentalLearner()
emmental_learner.learn(model, [train_dataloader])
return (model, emb_layer)
from electricity_utils import entity_level_f1
from fonduer_utils import schema_match_filter
price_col_keywords = ["price", "weighted avg."]
DEBUG = False
def eval_model(model, emb_layer, cands, F, schema_filter=False):
# Extract candidates and features
train_cands = cands[0]
dev_cands = cands[1]
test_cands = cands[2]
F_train = F[0]
F_dev = F[1]
F_test = F[2]
# apply schema filter
def apply(cands):
return schema_match_filter(
cands,
"station",
"price",
price_col_keywords,
stations_mapping_dict,
0.05,
DEBUG,
)
# Generate dataloader for test data
test_dataloader = EmmentalDataLoader(
task_to_label_dict={ATTRIBUTE: "labels"},
dataset=FonduerDataset(
ATTRIBUTE, test_cands[0], F_test[0], emb_layer.word2id, 2
),
split="test",
batch_size=100,
shuffle=False,
)
test_preds = model.predict(test_dataloader, return_preds=True)
positive = np.where(np.array(test_preds["probs"][ATTRIBUTE])[:, TRUE] > 0.6)
true_pred = [test_cands[0][_] for _ in positive[0]]
true_pred = apply(true_pred) if schema_filter else true_pred
test_results = entity_level_f1(true_pred, gold_file, ATTRIBUTE, test_docs, stations_mapping_dict=stations_mapping_dict)
# Run on dev and train set for validation
# We run the predictions also on our training and dev set, to validate that everything seems to work smoothly
# Generate dataloader for dev data
dev_dataloader = EmmentalDataLoader(
task_to_label_dict={ATTRIBUTE: "labels"},
dataset=FonduerDataset(
ATTRIBUTE, dev_cands[0], F_dev[0], emb_layer.word2id, 2
),
split="test",
batch_size=100,
shuffle=False,
)
dev_preds = model.predict(dev_dataloader, return_preds=True)
positive_dev = np.where(np.array(dev_preds["probs"][ATTRIBUTE])[:, TRUE] > 0.6)
true_dev_pred = [dev_cands[0][_] for _ in positive_dev[0]]
true_dev_pred = apply(true_dev_pred) if schema_filter else true_dev_pred
dev_results = entity_level_f1(true_dev_pred, gold_file, ATTRIBUTE, dev_docs, stations_mapping_dict=stations_mapping_dict)
# Generate dataloader for train data
train_dataloader = EmmentalDataLoader(
task_to_label_dict={ATTRIBUTE: "labels"},
dataset=FonduerDataset(
ATTRIBUTE, train_cands[0], F_train[0], emb_layer.word2id, 2
),
split="test",
batch_size=100,
shuffle=False,
)
train_preds = model.predict(train_dataloader, return_preds=True)
positive_train = np.where(np.array(train_preds["probs"][ATTRIBUTE])[:, TRUE] > 0.6)
true_train_pred = [train_cands[0][_] for _ in positive_train[0]]
true_train_pred = apply(true_train_pred) if schema_filter else true_train_pred
train_results = entity_level_f1(true_train_pred, gold_file, ATTRIBUTE, train_docs, stations_mapping_dict=stations_mapping_dict)
return [train_results, dev_results, test_results]
```
## Evaluating on the Test Set
```
# Based on gold labels or labeling functions (gold/full data set)
train_marginals_gold = np.array([[0,1] if gold(x) else [1,0] for x in train_cands[0]])
train_marginals = train_marginals_gold if dataset == 'gold' else train_marginals_lfs
from electricity_utils import summarize_results
# Build model and evaluate for Logistic Regression
(lr_model, lr_emb_layer) = train_model(cands, F, train_marginals, "LogisticRegression" )
print("Evaluate Logistic Regression method")
lr_results = eval_model(lr_model, lr_emb_layer, cands, F)
(prec_total, rec_total, f1_total) = summarize_results(lr_results)
print(f"TOTAL DOCS PAIRWISE (LogisticRegression): Precision={prec_total}, Recall={rec_total}, F1={f1_total}")
print("Evaluate Logistic Regression method with schema matching")
lr_results = eval_model(lr_model, lr_emb_layer, cands, F, True)
(prec_total, rec_total, f1_total) = summarize_results(lr_results)
print(f"TOTAL DOCS PAIRWISE (LogisticRegression): Precision={prec_total}, Recall={rec_total}, F1={f1_total}")
# Build model and evaluate for LSTM
(lstm_model, lstm_emb_layer) = train_model(cands, F, train_marginals, "LSTM" )
print("Evaluate LSTM method")
lstm_results = eval_model(lstm_model, lstm_emb_layer, cands, F)
(prec_total, rec_total, f1_total) = summarize_results(lstm_results)
print(f"TOTAL DOCS PAIRWISE (LSTM): Precision={prec_total}, Recall={rec_total}, F1={f1_total}")
print("Evaluate LSTM method with schema matching")
lstm_results = eval_model(lstm_model, lstm_emb_layer, cands, F, True)
(prec_total, rec_total, f1_total) = summarize_results(lstm_results)
print(f"TOTAL DOCS PAIRWISE (LSTM): Precision={prec_total}, Recall={rec_total}, F1={f1_total}")
```
# Phase 4: Error Analysis & Iterative KBC
- Analyise the false positive (FP) and false negative (FN) candidates
- Use the visualization tool to better understand which labeling functions might be responsible
- Test the labeling functions on this candidates to verify they work as expected
```
# from electricity_utils import entity_to_candidates
# def display_cand(cand_nr):
# # Get a list of candidates that match the FN[10] entity
# fp_cands = entity_to_candidates(FP[cand_nr], test_cands[0])
# # Display a candidate
# fp_cand = fp_cands[0]
# print(fp_cand)
# print(f"Number of FP: {cand_nr}/{len(FP)}")
# vis.display_candidates([fp_cand])
# return fp_cand
# maximum = len(FP)-1
# from ipywidgets import widgets
# from functools import partial
# from IPython.display import display, clear_output
# class Counter:
# def __init__(self, initial=0, maximum=0, minimum=0):
# self.value = initial
# self.maximum = maximum
# self.minimum = 0
# self.cand = display_cand(initial)
# def increment(self, amount=1):
# if (self.value+amount > self.maximum):
# return self.value
# self.value += amount
# return self.value
# def decrement(self, amount=1):
# if (self.value-amount < 0):
# return self.value
# self.value -= amount
# return self.value
# def __iter__(self, sentinal=False):
# return iter(self.increment, sentinal)
# def display_all(cand_nr):
# # Clear previous
# clear_output(wait=True)
# # Redraw
# display(minus)
# display(plus)
# return display_cand(cand_nr)
# def btn_inc(counter, w):
# counter.increment()
# counter.cand = display_all(counter.value)
# def btn_dec(counter, w):
# counter.decrement()
# counter.cand = display_all(counter.value)
# counter = Counter(40, maximum)
# minus = widgets.Button(description='<')
# minus.on_click(partial(btn_dec, counter))
# plus = widgets.Button(description='>')
# plus.on_click(partial(btn_inc, counter))
# display(minus)
# display(plus)
# # Get a list of candidates that match the FN[10] entity
# tp_cands = entity_to_candidates(TP[40], test_cands[0])
# # Display a candidate
# print(f"Number of TP: {len(TP)}")
# print(tp_cands[0])
# vis.display_candidates([tp_cands[0]])
# # Get a list of candidates that match the FN[10] entity
# fn_cands = entity_to_candidates(FN[2], test_cands[0])
# # Display a candidate
# print(f"Number of FN: {len(FN)}")
# print(fn_cands)
# vis.display_candidates([fn_cands[0]])
# result = re.compile(station_rgx, flags=re.I).search(mentions[len(mentions)-7].document.name)
# result
# import spacy
# from itertools import chain, tee, groupby, product
# from fonduer.utils.data_model_utils.tabular import _get_aligned_sentences
# from itertools import groupby
# import operator
# def get_col(m):
# s = m.context.sentence
# if (not s.is_tabular()):
# return -1
# if (s.cell.col_start != s.cell.col_end):
# return -1
# return s.cell.col_start
# def get_headers(mentions_col):
# m_sentences = [m.context.sentence for m in mentions_col]
# min_row = min([x.cell.row_start for x in m_sentences])
# s = m_sentences[0]
# aligned = [x.text for x in _get_aligned_sentences(s, axis=1) if x not in m_sentences and x.cell.row_end < min_row]
# # TODO: HEADER cell-annotation condition
# return aligned
# def get_sim(mentions_col_it, fid, pos_keyw, id_dict):
# headers = " , ".join(get_headers(list(mentions_col_it)))
# pos_keyw_vec = nlp(" , ".join(pos_keyw + id_dict[fid.context.get_span().lower()]))
# headers_vec = nlp(headers)
# # vectorize with word2vec and measure the similarity to positive/negative schema column keywords
# return pos_keyw_vec.similarity(headers_vec)
# def schema_match_filter(cands, id_field, filter_field, pos_keyw = [], id_dict = {}, variance=0.05, DEBUG=False):
# filtered_cands = []
# # group them by document, itertools requires sorting
# cands.sort(key=lambda c: c.document.name)
# for doc, doc_it in groupby(cands, lambda c: c.document.name):
# # group them by the candidate id field (e.g. all prices for one station-id)
# doc_cands = list(doc_it)
# doc_cands.sort(key=lambda c: getattr(c, id_field))
# for fid, doc_cand_it in groupby(doc_cands, lambda c: getattr(c, id_field)):
# it1, it2, it3 = tee(doc_cand_it, 3)
# # group by col
# doc_ms = [getattr(c, filter_field) for c in iter(it1)]
# doc_ms.sort(key=lambda m: get_col(m))
# ms_by_cols = { col:list(it) for col, it in groupby(doc_ms, lambda m: get_col(m)) }
# # ignore non tabular or multi-col/row
# if (-1 in ms_by_cols.keys()):
# filtered_cands += [c for c in iter(it2) if getattr(c, filter_field) in ms_by_cols[-1]]
# # Compare headers of each column based on semantic similarity (word vectors)
# similarities = { col:get_sim(it, fid, pos_keyw, id_dict) for col, it in ms_by_cols.items() if col != -1 }
# sim_sorted = [(col, sim) for col, sim in sorted(similarities.items(), key=lambda i: i[1], reverse=True)]
# maximum = sim_sorted[0]
# # If there is a conflict (multiple assigned columns)
# # only take the maximum similarity as true for this candidate match
# if (len(sim_sorted) > 1 and DEBUG):
# print("#####################################")
# print(f"Similarity for {fid.context.get_span()} in doc {doc}")
# print(similarities)
# print(f"The maximum similarity is for entries in column {maximum}")
# print()
# for col, it in ms_by_cols.items():
# print(f"Col {col} with {len(list(it))} entries and headers:")
# pprint(get_headers(list(it)))
# print()
# # Filter only the k maximal similar column candidates based on variance
# for i in sim_sorted:
# if (i[1] >= maximum[1]-variance):
# if (len(sim_sorted) > 1 and DEBUG):
# print("KEEP", i)
# filtered_cands += [c for c in iter(it3) if getattr(c, filter_field) in ms_by_cols[i[0]]]
# # only max column
# # counts = { col:len(list(it)) for col, it in ms_by_cols.items() if col != -1 }
# # maximum = max(counts.items(), key=operator.itemgetter(1))[0]
# # if (len(counts) > 1):
# # print("max and all", doc, maximum, counts, get_header(ms_by_cols[maximum][0]))
# # pprint(ms_by_cols)
# # print()
# return filtered_cands
# nlp = spacy.load("en_core_web_lg")
# price_pos_keywords = ["price", "firm", "on peak", "weighted avg."]
# result = schema_match_filter(train_cands[0], "station", "price", price_pos_keywords, stations_mapping_dict)
# print(len(result), "vs", len(train_cands[0]))
```
| github_jupyter |
<a href="https://colab.research.google.com/github/ceos-seo/odc-colab/blob/master/notebooks/02.03.Colab_Median_Mosaic_L8.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
Downloads the odc-colab Python module and runs it to setup ODC.
```
!wget -nc https://raw.githubusercontent.com/ceos-seo/odc-colab/master/odc_colab.py
from odc_colab import odc_colab_init
odc_colab_init(install_odc_gee=True)
```
Downloads an existing index and populates the new ODC environment with it.
```
from odc_colab import populate_db
populate_db()
```
<a id="top"></a>
# Landsat-8 Cloud-Filtered Median Mosaic
This notebook uses global Landsat-8 data from Google Earth Engine which has been indexed to work with the Open Data Cube. The selected data is used to create a custom Landsat-8 cloud-filtered median mosaic for any time period and location. The mosaic can be output as a GeoTIFF product for analysis in external GIS tools. The median mosaic is quite common and reflects the "median" value of cloud-free pixels for all spectral bands in the time series.
Users should review the "Cloud Statistics" notebook for more information about the temporal and spatial samples that are used for any median mosaic. An understanding of the underlying data is important for creating a valid mosaic for further analyses. In many cases, cloud contamination will create poor or time-biased mosaics unless there are sufficient cloud-free pixels in the time series. With a careful review of the cloud coverage over a given region and time period, it is possible to improve the mosaics and avoid false outputs.
Most mosaics are created for annual or seasonal time periods. In cloudy regions there may not be any clear (non-cloudy) pixels for the defined time window. If this is the case, those pixels will be masked from the final median mosaic product and be shown as "white" regions in the output image.
This baseline notebook runs in about 5 minutes. The default region (0.15 degrees square) and time window (one year) uses about 20% of the allocated RAM memory. Selecting larger regions and time windows should be done carefully to avoid exceeding the system limits or having long run times. For example, regions near 0.5-degree square will "crash" the code and not run to completion.
```
# Suppress Warning Messages
import warnings
warnings.filterwarnings('ignore')
# Load Data Cube Configuration
from odc_gee import earthengine
dc = earthengine.Datacube(app='Median Mosaic')
# Import Data Cube API
import utils.data_cube_utilities.data_access_api as dc_api
api = dc_api.DataAccessApi()
# Import Utilities
import xarray as xr
import numpy as np
import matplotlib.pyplot as plt
# Define the Product and Platform
# This data is indexed from Google Earth Engine data sources
product = "ls8_google"
platform = "LANDSAT_8"
```
## <span id="define_extents">Define the Extents of the Analysis [▴](#top)</span>
```
# MODIFY HERE
# Select a Latitude-Longitude point for the center of the analysis region
# Select the size of the box (in degrees) surrounding the center point
# Mombasa, Kenya
lat_long = (-4.03, 39.62)
box_size_deg = 0.15
# Calculate the latitude and longitude bounds of the analysis box
latitude = (lat_long[0]-box_size_deg/2, lat_long[0]+box_size_deg/2)
longitude = (lat_long[1]-box_size_deg/2, lat_long[1]+box_size_deg/2)
# Select a time range
# The inputs require a format (Min,Max) using this date format (YYYY-MM-DD)
# The Landsat-8 allowable time range is: 2013-04-07 to current
time_extents = ('2020-01-01', '2020-12-31')
# The code below renders a map that can be used to view the region.
# It is possible to find new regions using the map below.
# Use your mouse to zoom in/out to explore new regions
# Click on the map to view Lat-Lon coordinates of any location that could define the region boundary
from utils.data_cube_utilities.dc_display_map import display_map
display_map(latitude,longitude)
```
## <span id="load_data">Load the Data and Mask the Clouds [▴](#top)</span>
```
landsat_dataset = dc.load(latitude=latitude,longitude=longitude,platform=platform,time=time_extents,
product=product,measurements=['red', 'green', 'blue', 'nir', 'swir1', 'swir2', 'pixel_qa'])
from utils.data_cube_utilities.clean_mask import landsat_qa_clean_mask
cloud_mask = landsat_qa_clean_mask(landsat_dataset, platform=platform)
cleaned_dataset = landsat_dataset.where(cloud_mask)
```
## <span id="mosaics">Create Median Mosaic and View the Results [▴](#top)</span>
```
# Load the median mosaic function and create the median mosaic
from utils.data_cube_utilities.dc_mosaic import create_median_mosaic
median_composite = create_median_mosaic(cleaned_dataset, cloud_mask)
# Load the plotting utility
from utils.data_cube_utilities.dc_rgb import rgb
# Show Median Mosaic
# Select the output image bands for each color (Red, Green, Blue)
# Users can create other combinations of bands, as desired
# True-Color = red, green, blue (this is the common true-color RGB image)
# False-Color = swir2, nir, green (this is commonly used for Landsat data viewing)
median_rgb = median_composite[['swir2', 'nir', 'green']].to_array()
# Define the plot settings and show the plots
# Users may want to alter the figure sizes or plot titles
# The "vmax" value controls the brightness of the images and can be adjusted
median_rgb.plot.imshow(vmin=0, vmax=5000, figsize=(10,10))
plt.title("Median Mosaic")
plt.axis('off')
plt.show()
```
## <span id="export">Create GeoTIFF Output Product [▴](#top)</span>
```
from utils.data_cube_utilities.import_export import export_slice_to_geotiff
# MODIFY HERE
# Change the name of the output file, or it will be overwritten for each run
# The output file can be found using the "file" icon on the Colab menu on the left
export_slice_to_geotiff(median_composite, './output/DEMO_median_composite.tif')
```
| github_jupyter |
# pandas Data
> Marcos Duarte
> Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
> Federal University of ABC, Brazil
```
import numpy as np
import pandas as pd
%matplotlib notebook
# tk qt notebook inline ipympl
import matplotlib as mpl
import matplotlib.pyplot as plt
import sys, os
sys.path.insert(1, r'./../functions')
path2 = r'./../../../X/Clau/'
name = 'WBDS01walkT06mkr.txt'
fname = os.path.join(path2, name)
```
## pandas with one index
```
df = pd.read_csv(fname, sep='\t', header=0, index_col=0, dtype=np.float64, engine='c')
df.columns = df.columns.str.replace('\.', '')
df.head()
ax = df.plot(y='RASISX', figsize=(8, 3), title='A plot of kinematics')
ax.set_ylabel('Position [mm]')
plt.tight_layout(pad=0, h_pad=0, rect=[0, 0, 1, .95])
def plot_widget(df):
"""general plot widget of a pandas dataframe
"""
from ipywidgets import widgets
col_w = widgets.SelectMultiple(options=df.columns,
value=[df.columns[0]],
description='Column')
clear_w = widgets.Checkbox(value=True,
description='Clear axis')
container = widgets.HBox(children=[col_w, clear_w])
display(container)
fig, ax = plt.subplots(1, 1, figsize=(9, 4))
if col_w.value:
df.plot(y=col_w.value[0], ax=ax)
plt.tight_layout()
plt.show()
def plot(change):
if clear_w.value:
ax.clear()
for c in col_w.value:
df.plot(y=c, ax=ax)
col_w.observe(plot, names='value')
plot_widget(df)
```
## pandas multiindex
Data with hierarchical column index ([multiindex](http://pandas.pydata.org/pandas-docs/stable/advanced.html#creating-a-multiindex-hierarchical-index-object)) where columns have multiple levels.
```
df = pd.read_csv(fname, sep='\t', header=0, index_col=0, dtype=np.float64, engine='c')
# format columns as multindexes and relabel them
cols = [s[:-1] for s in df.columns.str.replace('\.', '')]
df.columns = [cols, list('XYZ')*int(df.shape[1]/3)]
df.columns.set_names(names=['Marker', 'Coordinate'], level=[0, 1], inplace=True)
#df = df.swaplevel(0, 1, axis=1) # for 'Coordinate' to go first
df.head()
df['RASIS'].head()
# df.RASIS.head()
df.RASIS.X.head()
# df['RASIS']['X'].head()
df.xs('X', level='Coordinate', axis=1).head()
df.loc[:, (slice(None), 'X')].head()
df.swaplevel(0, 1, axis=1).head() # for 'Coordinate' to go first
ax = df.plot(y=('RASIS', 'X'), subplots=True, figsize=(8, 2), rot=0)
ax = df.plot(y='RASIS', subplots=True, sharex=True, figsize=(8, 4), rot=0,
title='A plot of kinematics')
plt.tight_layout(pad=0, h_pad=0, rect=[0, 0, 1, .95])
values = df.reset_index(drop=False).values
values[0, :5]
df.head()
x = df.swaplevel(0, 1, axis=1)
x2 = x.unstack(level=-1)
x2.head()
x.head()
```
| github_jupyter |
```
#import IBM API key and service URL from ibm-credentials.env
import os
from dotenv import load_dotenv
load_dotenv('ibm-nlu-credentials.env')
API_KEY = os.getenv('NATURAL_LANGUAGE_UNDERSTANDING_APIKEY')
URL=os.getenv('NATURAL_LANGUAGE_UNDERSTANDING_URL')
#import lib
import json
from ibm_watson import NaturalLanguageUnderstandingV1
from ibm_cloud_sdk_core.authenticators import IAMAuthenticator
from ibm_watson.natural_language_understanding_v1 import Features, EntitiesOptions, KeywordsOptions
#load API key
authenticator = IAMAuthenticator(API_KEY)
natural_language_understanding = NaturalLanguageUnderstandingV1(
version='2022-03-22',
authenticator=authenticator
)
natural_language_understanding.set_service_url(URL)
#csv to 2-D list
import csv
#import numpy as np
data_arr=[]
title_arr=[]
id_arr=[]
country_arr=[]
sector_arr=[]
date_arr=[]
with open("locanto_demo.csv",newline='',encoding='utf-8') as f:
reader=csv.reader(f)
boolVal = 0
for row in reader:
data_arr.append(row)
temp = str(row).split(',')
while boolVal == 1:
title_arr.append(temp[0])
for i in temp:
try:
if isinstance(int(i.replace("'","")), int):
if(i.replace("'", "") != '000'):
id_arr.append(i)
except Exception as e:
i = i #placeholder so the code doesn't stop when it finds a non number value
# checking if the second last value in the row is any sector that has a comma in it's title as it
# separates the sector into two values
finalValue = temp[len(temp)-2]
if (finalValue == "Accounting" or finalValue == "Hospitality" or finalValue == "Marketing" or finalValue == "Retail"):
sector_arr.append(finalValue)
sector_arr.append(temp[len(temp)-1])
date_arr.append(temp[len(temp)-3])
country_arr.append(temp[len(temp)-4])
else:
sector_arr.append(temp[len(temp)-1])
date_arr.append(temp[len(temp)-2])
country_arr.append(temp[len(temp)-3])
boolVal = 0
boolVal = 1
#print(id_arr)
print(len(title_arr))
print(len(id_arr))
print(len(country_arr))
#NLU
line=1
print("{\"ads\": [")
while line < len(data_arr):
#while line < 4:
response = natural_language_understanding.analyze(
text=data_arr[line][2],
features=Features(
keywords=KeywordsOptions(emotion=False, sentiment=False,limit=1))).get_result()
# take the specific string values out of the array and remove unwanted characters
unwantedChars = "\"][\'"
temp_title = title_arr[line-1]
temp_id = id_arr[line-1].replace("'","")
temp_country = country_arr[line-1].replace("'","")
temp_sector = sector_arr[line-1].replace("'","")
temp_date = date_arr[line-1].replace("'", "")
# as these are the first and last values in each row, they have extra unwanted characters
for unwantedChar in unwantedChars:
temp_title = temp_title.replace(unwantedChar, "")
temp_sector = temp_sector.replace(unwantedChar,"")
print(f"{'{'}\n")
print(f" \"{'Job_title'}\": {'{'}\n \"{'title'}\": \"{temp_title}\"\n {'}'},\n")
print(f" \"{'Ad_id'}\": {'{'}\n \"{'id'}\": \"{temp_id}\"\n {'}'},\n")
print(f" \"{'Country'}\": {'{'}\n \"{'country'}\": \"{temp_country}\"\n {'}'},\n")
print(f" \"{'Sector'}\": {'{'}\n \"{'sector'}\": \"{temp_sector}\"\n {'}'},\n")
print(f" \"{'Date'}\": {'{'}\n \"{'date'}\": \"{temp_date}\"\n {'}'},\n")
print(f" {(json.dumps(response, indent=2))[1:-1]}\n")
str = (('}', '},') [line < (len(data_arr)-1)])
print(str)
line = line + 1
print("]}")
```
| github_jupyter |
<a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_08_1_kaggle_intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# T81-558: Applications of Deep Neural Networks
**Module 8: Kaggle Data Sets**
* Instructor: [Jeff Heaton](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
* For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# Module 8 Material
* **Part 8.1: Introduction to Kaggle** [[Video]](https://www.youtube.com/watch?v=v4lJBhdCuCU&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_1_kaggle_intro.ipynb)
* Part 8.2: Building Ensembles with Scikit-Learn and Keras [[Video]](https://www.youtube.com/watch?v=LQ-9ZRBLasw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_2_keras_ensembles.ipynb)
* Part 8.3: How Should you Architect Your Keras Neural Network: Hyperparameters [[Video]](https://www.youtube.com/watch?v=1q9klwSoUQw&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_3_keras_hyperparameters.ipynb)
* Part 8.4: Bayesian Hyperparameter Optimization for Keras [[Video]](https://www.youtube.com/watch?v=sXdxyUCCm8s&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_4_bayesian_hyperparameter_opt.ipynb)
* Part 8.5: Current Semester's Kaggle [[Video]](https://www.youtube.com/watch?v=PHQt0aUasRg&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_08_5_kaggle_project.ipynb)
# Part 8.1: Introduction to Kaggle
[Kaggle](http://www.kaggle.com) runs competitions where data scientists compete to provide the best model to fit the data. A simple project to get started with Kaggle is the [Titanic data set](https://www.kaggle.com/c/titanic-gettingStarted). Most Kaggle competitions end on a specific date. Website organizers have currently scheduled the Titanic competition to end on December 31, 20xx (with the year usually rolling forward). However, they have already extended the deadline several times, and an extension beyond 2014 is also possible. Second, the Titanic data set is considered a tutorial data set. There is no prize, and your score in the competition does not count towards becoming a Kaggle Master.
### Kaggle Ranks
You achieve Kaggle ranks by earning gold, silver, and bronze medals.
* [Kaggle Top Users](https://www.kaggle.com/rankings)
* [Current Top Kaggle User's Profile Page](https://www.kaggle.com/stasg7)
* [Jeff Heaton's (your instructor) Kaggle Profile](https://www.kaggle.com/jeffheaton)
* [Current Kaggle Ranking System](https://www.kaggle.com/progression)
### Typical Kaggle Competition
A typical Kaggle competition will have several components. Consider the Titanic tutorial:
* [Competition Summary Page](https://www.kaggle.com/c/titanic)
* [Data Page](https://www.kaggle.com/c/titanic/data)
* [Evaluation Description Page](https://www.kaggle.com/c/titanic/details/evaluation)
* [Leaderboard](https://www.kaggle.com/c/titanic/leaderboard)
### How Kaggle Competition Scoring
Kaggle is provided with a data set by the competition sponsor, as seen in Figure 8.SCORE. Kaggle divides this data set as follows:
* **Complete Data Set** - This is the complete data set.
* **Training Data Set** - This dataset provides both the inputs and the outcomes for the training portion of the data set.
* **Test Data Set** - This dataset provides the complete test data; however, it does not give the outcomes. Your submission file should contain the predicted results for this data set.
* **Public Leaderboard** - Kaggle does not tell you what part of the test data set contributes to the public leaderboard. Your public score is calculated based on this part of the data set.
* **Private Leaderboard** - Likewise, Kaggle does not tell you what part of the test data set contributes to the public leaderboard. Your final score/rank is calculated based on this part. You do not see your private leaderboard score until the end.
**Figure 8.SCORE: How Kaggle Competition Scoring**

### Preparing a Kaggle Submission
You do not submit the code to your solution to Kaggle. For competitions, you are scored entirely on the accuracy of your submission file. A Kaggle submission file is always a CSV file that contains the **Id** of the row you are predicting and the answer. For the titanic competition, a submission file looks something like this:
```
PassengerId,Survived
892,0
893,1
894,1
895,0
896,0
897,1
...
```
The above file states the prediction for each of the various passengers. You should only predict on ID's that are in the test file. Likewise, you should render a prediction for every row in the test file. Some competitions will have different formats for their answers. For example, a multi-classification will usually have a column for each class and your predictions for each class.
# Select Kaggle Competitions
There have been many exciting competitions on Kaggle; these are some of my favorites. Some select predictive modeling competitions, which use tabular data include:
* [Otto Group Product Classification Challenge](https://www.kaggle.com/c/otto-group-product-classification-challenge)
* [Galaxy Zoo - The Galaxy Challenge](https://www.kaggle.com/c/galaxy-zoo-the-galaxy-challenge)
* [Practice Fusion Diabetes Classification](https://www.kaggle.com/c/pf2012-diabetes)
* [Predicting a Biological Response](https://www.kaggle.com/c/bioresponse)
Many Kaggle competitions include computer vision datasets, such as:
* [Diabetic Retinopathy Detection](https://www.kaggle.com/c/diabetic-retinopathy-detection)
* [Cats vs Dogs](https://www.kaggle.com/c/dogs-vs-cats)
* [State Farm Distracted Driver Detection](https://www.kaggle.com/c/state-farm-distracted-driver-detection)
# Module 8 Assignment
You can find the first assignment here: [assignment 8](https://github.com/jeffheaton/t81_558_deep_learning/blob/master/assignments/assignment_yourname_class8.ipynb)
| github_jupyter |
```
import numpy as np
import pandas as pd
from sklearn.linear_model import LogisticRegression
from sklearn.utils import shuffle
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import log_loss
from sklearn.ensemble import RandomForestClassifier
from datetime import datetime
# Load the data sets
data_dir = './data/'
df_Household = pd.read_csv(data_dir + 'Household.csv')
df_Trip = pd.read_csv(data_dir + 'Trip.csv')
df_Person = pd.read_csv(data_dir + 'Person.csv')
df_Blockgroup_UrbanVillage = pd.read_csv(data_dir + 'Blockgroup_UrbanVillage.csv')
#print (df_Household.head())
print (df_Blockgroup_UrbanVillage.head())
```
## Process Persons (Race demographics)
```
# load race information
df_persons = df_Person[['hhid','race_afam','race_aiak','race_asian','race_hapi','race_hisp','race_white','race_other']]
df_persons.dropna(axis = 0, inplace=True)
#race_afam race_aiak race_asian race_hapi race_hisp race_white race_other
df_race = df_persons.groupby(['hhid'], as_index=False).agg({'race_afam':sum,'race_aiak':sum,
'race_asian':sum,'race_hapi':sum,
'race_white':sum,'race_other':sum,
'race_hisp':sum})
df_race['race_total'] = df_race['race_afam'] + df_race['race_asian'] + df_race['race_hisp'] + df_race['race_white'] + df_race['race_other']
df_race['pct_white'] = df_race['race_white'] / df_race['race_total']
print (df_race)
```
## Process Households
```
# load household data
df_households = df_Household[['bg_household','hhid','final_lat','final_lng','hhsize','vehicle_count','numchildren',
'hhincome_broad','car_share','rent_own','res_dur','offpark','hh_wt_final']]
df_households['bg_household'] = df_households['bg_household'].astype(float).astype(int).astype(str)
# merge with seattle block group data
df_seattle = df_Blockgroup_UrbanVillage[['BLOCKGROUP','URBAN_VILLAGE_NAME','URBAN_VILLAGE_TYPE']]
df_seattle['bg_household'] = df_seattle['BLOCKGROUP'].astype(str)
df_households = pd.merge(left=df_households, right=df_seattle, how='left', on='bg_household')
df_households.drop(['BLOCKGROUP'], axis = 1, inplace=True)
df_households['URBAN_VILLAGE_NAME'] = df_households['URBAN_VILLAGE_NAME'].fillna("Outside Seattle")
df_households['URBAN_VILLAGE_TYPE'] = df_households['URBAN_VILLAGE_TYPE'].fillna("Outside Seattle")
df_households = df_households.rename(columns={'final_lat':'hh_lat', 'final_lng':'hh_lng'})
df_households = df_households.rename(columns={'URBAN_VILLAGE_NAME':'hh_uv', 'URBAN_VILLAGE_TYPE':'hh_uv_type'})
# Assign income variables
df_households['income'] = np.where(df_households['hhincome_broad']==1, "Under $25,000", "")
df_households['income'] = np.where(df_households['hhincome_broad']==2, "$25,000-$49,999", df_households['income'])
df_households['income'] = np.where(df_households['hhincome_broad']==3, "$50,000-$74,999", df_households['income'])
df_households['income'] = np.where(df_households['hhincome_broad']==4, "$75,000-$99,999", df_households['income'])
df_households['income'] = np.where(df_households['hhincome_broad']==5, "$100,000 or more", df_households['income'])
df_households['income'] = np.where(df_households['hhincome_broad']==6, "Prefer not to answer", df_households['income'])
# Assign home ownership
df_households['homeownership'] = np.where(df_households['rent_own']==1, "Own", "Other")
df_households['homeownership'] = np.where(df_households['rent_own']==2, "Rent", df_households['homeownership'])
# Assign residency tenure
df_households['tenure'] = np.where(df_households['res_dur']==1, "Less than a year", "")
df_households['tenure'] = np.where(df_households['res_dur']==2, "Between 1 and 2 years", df_households['tenure'])
df_households['tenure'] = np.where(df_households['res_dur']==3, "Between 2 and 3 years", df_households['tenure'])
df_households['tenure'] = np.where(df_households['res_dur']==4, "Between 3 and 5 years", df_households['tenure'])
df_households['tenure'] = np.where(df_households['res_dur']==5, "Between 5 and 10 years", df_households['tenure'])
df_households['tenure'] = np.where(df_households['res_dur']==6, "Between 10 and 20 years", df_households['tenure'])
df_households['tenure'] = np.where(df_households['res_dur']==7, "More than 20 years", df_households['tenure'])
df_households = pd.merge(left=df_households, right=df_race, how='left', on='hhid')
print (df_households.head())
print (df_households)
```
## Process Trips
```
df_trips = df_Trip[['tripid','hhid','origin_lat','origin_lng','dest_lat','dest_lng',
'bg_origin','bg_dest','personid','google_duration','trip_path_distance',
'depart_time_timestamp','arrival_time_timestamp','daynum','o_purp','d_purp',
'mode_1','travelers_total','traveldate','trip_wt_final']]
df_trips['bg_dest'] = df_trips['bg_dest'].astype(str)
df_trips['bg_origin'] = df_trips['bg_origin'].astype(str)
# merge with seattle block group data
df_seattle = df_Blockgroup_UrbanVillage[['BLOCKGROUP','URBAN_VILLAGE_NAME','URBAN_VILLAGE_TYPE']]
df_seattle['BLOCKGROUP'] = df_seattle['BLOCKGROUP'].astype(str)
df_trips = pd.merge(left=df_trips, right=df_seattle, how='left', left_on='bg_origin', right_on='BLOCKGROUP')
df_trips = df_trips.rename(columns={'URBAN_VILLAGE_NAME':'uv_origin', 'URBAN_VILLAGE_TYPE':'uvType_origin'})
df_trips.drop(['BLOCKGROUP'], axis = 1, inplace=True)
df_trips['uv_origin'] = df_trips['uv_origin'].fillna("Outside Seattle")
df_trips['uvType_origin'] = df_trips['uvType_origin'].fillna("Outside Seattle")
df_trips = pd.merge(left=df_trips, right=df_seattle, how='left', left_on='bg_dest', right_on='BLOCKGROUP')
df_trips = df_trips.rename(columns={'URBAN_VILLAGE_NAME':'uv_dest', 'URBAN_VILLAGE_TYPE':'uvType_dest'})
df_trips.drop(['BLOCKGROUP'], axis = 1, inplace=True)
df_trips['uv_dest'] = df_trips['uv_dest'].fillna("Outside Seattle")
df_trips['uvType_dest'] = df_trips['uvType_dest'].fillna("Outside Seattle")
# Drop missing variables, clean up column
df_trips['mode_1'] = df_trips['mode_1'].fillna(0)
df_trips['mode_1'] = df_trips['mode_1'].astype(str).replace(' ', '0')
df_trips['mode_1'] = df_trips['mode_1'].astype(str).astype(int)
df_trips['travelers_total'] = df_trips['travelers_total'].astype(str).replace(' ', '0')
df_trips['travelers_total'] = df_trips['travelers_total'].astype(str).astype(int)
# drop rows where the duration or distance is null or an empty space
df_trips = df_trips[df_trips['google_duration'].notnull()]
df_trips = df_trips[df_trips['trip_path_distance'].notnull()]
df_trips = df_trips[df_trips['google_duration'] != " "]
df_trips = df_trips[df_trips['trip_path_distance'] != " "]
df_trips['google_duration'] = df_trips['google_duration'].astype(float)
df_trips['trip_path_distance'] = df_trips['trip_path_distance'].astype(float)
df_trips = df_trips[df_trips['trip_path_distance'].notnull()]
# Create OD Pairs for urban villages and block groups
df_trips['uv_od_pair'] = df_trips['uv_origin'] + " to " + df_trips['uv_dest']
df_trips['bg_od_pair'] = df_trips['bg_origin'].astype(str) + " to " + df_trips['bg_dest'].astype(str)
# Assign mode variables
df_trips['mode'] = np.where(df_trips['mode_1']==1, "Walk", "Other")
df_trips['mode'] = np.where(df_trips['mode_1']==2, "Bike", df_trips['mode'])
df_trips['mode'] = np.where((df_trips['mode_1']>=3) & (df_trips['mode_1']<=17) & (df_trips['travelers_total']==1), "Drive Alone", df_trips['mode'])
df_trips['mode'] = np.where(((df_trips['mode_1']==21) | (df_trips['mode_1']==22) |
(df_trips['mode_1']==33) | (df_trips['mode_1']==34) | (df_trips['mode_1']==18))
& (df_trips['travelers_total']==1), "Drive Alone", df_trips['mode'])
df_trips['mode'] = np.where((df_trips['mode_1']>=3) & (df_trips['mode_1']<=17) & (df_trips['travelers_total']!=1), "Drive w Others", df_trips['mode'])
df_trips['mode'] = np.where(((df_trips['mode_1']==21) | (df_trips['mode_1']==22) |
(df_trips['mode_1']==33) | (df_trips['mode_1']==34) | (df_trips['mode_1']==18))
& (df_trips['travelers_total']>1), "Drive w Others", df_trips['mode'])
df_trips['mode'] = np.where((df_trips['mode_1']==23) | (df_trips['mode_1']==41) | (df_trips['travelers_total']==42), "Transit", df_trips['mode'])
df_trips['mode'] = np.where((df_trips['mode_1']==32) | (df_trips['travelers_total']==52), "Transit", df_trips['mode'])
df_trips['drive_alone'] = np.where(df_trips['mode']=="Drive Alone", 1, 0)
# Assign purpose variables
df_trips['purpose'] = np.where(df_trips['d_purp']==1, "Go Home", "Other")
df_trips['purpose'] = np.where(df_trips['d_purp']==6, "School", df_trips['purpose'])
df_trips['purpose'] = np.where((df_trips['d_purp']==10) | (df_trips['d_purp']==11), "Work", df_trips['purpose'])
# Assign time period variables
df_trips['depart_time_timestamp'] = pd.to_datetime(df_trips['depart_time_timestamp'], errors='coerce')
df_trips['depart_day'] = df_trips['depart_time_timestamp'].dt.dayofweek
df_trips['depart_time'] = df_trips['depart_time_timestamp'].dt.hour
df_trips['depart_period'] = np.where((df_trips['depart_day']>=0) & (df_trips['depart_day']<=4) &
(df_trips['depart_time']>=7) & (df_trips['depart_time']<=9), "Weekday AM", "")
df_trips['depart_period'] = np.where((df_trips['depart_day']>=0) & (df_trips['depart_day']<=4) &
(df_trips['depart_time']>=10) & (df_trips['depart_time']<=15), "Weekday Mid", df_trips['depart_period'])
df_trips['depart_period'] = np.where((df_trips['depart_day']>=0) & (df_trips['depart_day']<=4) &
(df_trips['depart_time']>=16) & (df_trips['depart_time']<=19), "Weekday PM", df_trips['depart_period'])
df_trips['depart_period'] = np.where((df_trips['depart_day']>=0) & (df_trips['depart_day']<=4) &
(df_trips['depart_time']<7), "Other", df_trips['depart_period'])
df_trips['depart_period'] = np.where((df_trips['depart_day']>=0) & (df_trips['depart_day']<=4) &
(df_trips['depart_time']>19), "Late Night", df_trips['depart_period'])
df_trips['depart_period'] = np.where((df_trips['depart_day']>=5) & (df_trips['depart_day']<=6), "Late Night", df_trips['depart_period'])
#dts = dfBad[cols].apply(lambda x: pd.to_datetime(x, errors='coerce', format='%m/%d/%Y'))
#pd.to_datetime(df_trips['depart_time_timestamp'])
#df_trips['depart_time_timestamp'] = datetime.strptime(df_trips['depart_time_timestamp'], '%b %d %Y %I:%M%p')
#to_datetime
#print (df_trips.dtypes)
print (df_trips.head())
```
## Origin and Destination Summary
```
#filter for only trips starting and ending in Seattle
df_trips_Seattle = df_trips[(df_trips['uv_origin'] != 'Outside Seattle') | (df_trips['uv_dest'] != 'Outside Seattle')]
#print (df_trips_Seattle)
#aggregate by urban village origin
df_UV_Origins = df_trips_Seattle.groupby(['uv_origin'], as_index=False).agg({'google_duration':['mean','std','skew'],'trip_path_distance':['mean'],
'origin_lat':['mean','std','skew'],'origin_lng':['mean']})
print (df_UV_Origins.head())
#aggregate by urban village destination
df_UV_Destinations = df_trips_Seattle.groupby(['uv_dest'], as_index=False).agg({'google_duration':['mean'],'trip_path_distance':['mean'],
'dest_lat':['mean'],'dest_lng':['mean']})
#print (df_UV_Destinations.head())
#aggregate by urban village OD Pair
df_UV_ODPair = df_trips_Seattle.groupby(['uv_od_pair'], as_index=False).agg({'google_duration':['mean'],'trip_path_distance':['mean'],
'origin_lat':['mean'],'origin_lat':['mean'],
'dest_lat':['mean'],'dest_lng':['mean']})
#print (df_UV_ODPair.head())
##aggregate by blockgroup origin
df_BG_Origins = df_trips_Seattle.groupby(['bg_origin'], as_index=False).agg({'google_duration':['mean'],'trip_path_distance':['mean'],
'origin_lat':['mean'],'origin_lng':['mean']})
#print (df_BG_Origins)
#aggregate by blockgroup destination
df_BG_Destinations = df_trips_Seattle.groupby(['bg_dest'], as_index=False).agg({'google_duration':['mean'],'trip_path_distance':['mean'],
'dest_lat':['mean'],'dest_lng':['mean']})
#print (df_BG_Destinations.head())
#aggregate by urban village OD Pair
df_BG_ODPair = df_trips_Seattle.groupby(['bg_od_pair'], as_index=False).agg({'google_duration':['mean'],'trip_path_distance':['mean'],
'origin_lat':['mean'],'origin_lng':['mean'],
'dest_lat':['mean'],'dest_lng':['mean']})
df_BG_ODPair.columns = df_BG_ODPair.columns.droplevel(level=1)
df_BG_ODPair.to_csv(data_dir + 'df_BG_ODPair.csv', mode='w', header=True, index=False)
df_BG_Origins.to_csv(data_dir + 'BG_Origins.csv', mode='w', header=True, index=False)
```
## Merged, Normalized Trip Dataset
```
df_trip_household= pd.merge(left=df_trips, right=df_households, how='left', left_on='hhid', right_on='hhid')
print (df_trip_household.head())
df_trip_household.to_csv(data_dir + 'Trip_Household_Merged.csv', mode='w', header=True, index=False)
df_UV_Origins.to_csv(data_dir + 'UV_Origins.csv', mode='w', header=True, index=False)
```
| github_jupyter |
Before you turn this problem in, make sure everything runs as expected. First, **restart the kernel** (in the menubar, select Kernel$\rightarrow$Restart) and then **run all cells** (in the menubar, select Cell$\rightarrow$Run All).
Make sure you fill in any place that says `YOUR CODE HERE`, as well as your email used on the [moodle](https://moodle.ens.psl.eu/user/index.php?id=1020) below:
```
NAME = ""
```
---
# Homewrok 1: MLP from scratch
In this homework, you will code a [Multilayer perceptron](https://en.wikipedia.org/wiki/Multilayer_perceptron) with one hidden layer to classify cloud of points in 2D.
## 1. Some utilities and your dataset
You should not modify the code in this section
```
# all of these libraries are used for plotting
import numpy as np
import matplotlib.pyplot as plt
# Plot the dataset
def plot_data(ax, X, Y):
plt.axis('off')
ax.scatter(X[:, 0], X[:, 1], s=1, c=Y, cmap='bone')
from sklearn.datasets import make_moons
X, Y = make_moons(n_samples=2000, noise=0.1)
%matplotlib inline
x_min, x_max = -1.5, 2.5
y_min, y_max = -1, 1.5
fig, ax = plt.subplots(1, 1, facecolor='#4B6EA9')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
plot_data(ax, X, Y)
plt.show()
```
This is your dataset: two moons each one corresponding to one class (black or white in the picture above).
In order to make it more fun and illustrative, the code below allows you to see the decision boundary of your classifier. Unfortunately, animation is not working on colab...
```
# Define the grid on which we will evaluate our classifier
xx, yy = np.meshgrid(np.arange(x_min, x_max, .1),
np.arange(y_min, y_max, .1))
to_forward = np.array(list(zip(xx.ravel(), yy.ravel())))
# plot the decision boundary of our classifier
def plot_decision_boundary(ax, X, Y, classifier):
# forward pass on the grid, then convert to numpy for plotting
Z = classifier.forward(to_forward)
Z = Z.reshape(xx.shape)
# plot contour lines of the values of our classifier on the grid
ax.contourf(xx, yy, Z>0.5, cmap='Blues')
# then plot the dataset
plot_data(ax, X,Y)
```
## 2. MLP in numpy
Here you need to code your implementation of the [ReLU](https://en.wikipedia.org/wiki/Rectifier_(neural_networks) activation and the [Sigmoid](https://en.wikipedia.org/wiki/Sigmoid_function).
```
class MyReLU(object):
def forward(self, x):
# the relu is y_i = max(0, x_i)
# YOUR CODE HERE
raise NotImplementedError()
def backward(self, grad_output):
# the gradient is 1 for the inputs that were above 0, 0 elsewhere
# YOUR CODE HERE
raise NotImplementedError()
def step(self, learning_rate):
# no need to do anything here, since ReLU has no parameters
# YOUR CODE HERE
raise NotImplementedError()
class MySigmoid(object):
def forward(self, x):
# the sigmoid is y_i = 1./(1+exp(-x_i))
# YOUR CODE HERE
raise NotImplementedError()
def backward(self, grad_output):
# the partial derivative is e^-x / (e^-x + 1)^2
# YOUR CODE HERE
raise NotImplementedError()
def step(self, learning_rate):
# no need to do anything here since Sigmoid has no parameters
# YOUR CODE HERE
raise NotImplementedError()
```
Probably a good time to test your functions...
```
test_relu = MyReLU()
test_relu.forward(X[10])
test_relu.backward(1.)
test_sig = MySigmoid()
test_sig.forward(np.ones(1))
test_sig.backward(np.ones(1))
```
A bit more complicated, you need now to implement your linear layer i.e. multiplication by a matrix W and summing with a bias b.
```
class MyLinear(object):
def __init__(self, n_input, n_output):
# initialize two random matrices for W and b (use np.random.randn)
# YOUR CODE HERE
raise NotImplementedError()
def forward(self, x):
# save a copy of x, you'll need it for the backward
# return Wx + b
# YOUR CODE HERE
raise NotImplementedError()
def backward(self, grad_output):
# y_i = \sum_j W_{i,j} x_j + b_i
# d y_i / d W_{i, j} = x_j
# d loss / d y_i = grad_output[i]
# so d loss / d W_{i,j} = x_j * grad_output[i] (by the chain rule)
# YOUR CODE HERE
raise NotImplementedError()
# d y_i / d b_i = 1
# d loss / d y_i = grad_output[i]
# YOUR CODE HERE
raise NotImplementedError()
# now we need to compute the gradient with respect to x to continue the back propagation
# d y_i / d x_j = W_{i, j}
# to compute the gradient of the loss, we have to sum over all possible y_i in the chain rule
# d loss / d x_j = \sum_i (d loss / d y_i) (d y_i / d x_j)
# YOUR CODE HERE
raise NotImplementedError()
def step(self, learning_rate):
# update self.W and self.b in the opposite direction of the stored gradients, for learning_rate
# YOUR CODE HERE
raise NotImplementedError()
```
As we did in practicals, you need now to code your network (what we called my_composition in the [practicals](https://github.com/dataflowr/notebooks/blob/master/Module2/02_backprop.ipynb)). Recall with a Sigmoid layer, you should use the BCE loss.
```
class Sequential(object):
def __init__(self, layers):
# YOUR CODE HERE
raise NotImplementedError()
def forward(self, x):
# YOUR CODE HERE
raise NotImplementedError()
def compute_loss(self, out, label):
# use the BCE loss
# -(label * log(output) + (1-label) * log(1-output))
# save the gradient, and return the loss
# beware of dividing by zero in the gradient.
# split the computation in two cases, one where the label is 0 and another one where the label is 1
# add a small value (1e-10) to the denominator
# YOUR CODE HERE
raise NotImplementedError()
def backward(self):
# apply backprop sequentially, starting from the gradient of the loss
# YOUR CODE HERE
raise NotImplementedError()
def step(self, learning_rate):
# take a gradient step for each layers
# YOUR CODE HERE
raise NotImplementedError()
h=50
# define your network with your Sequential
# it should be a linear layer with 2 inputs and h outputs, followed by a ReLU
# then a linear layer with h inputs and 1 outputs, followed by a sigmoid
# feel free to try other architectures
# YOUR CODE HERE
raise NotImplementedError()
# unfortunately animation is not working on colab
# you should comment the following line if on colab
%matplotlib notebook
fig, ax = plt.subplots(1, 1, facecolor='#4B6EA9')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
losses = []
learning_rate = 1e-2
for it in range(10000):
# pick a random example id
j = np.random.randint(1, len(X))
# select the corresponding example and label
example = X[j:j+1]
label = Y[j]
# do a forward pass on the example
# YOUR CODE HERE
raise NotImplementedError()
# compute the loss according to your output and the label
# YOUR CODE HERE
raise NotImplementedError()
# backward pass
# YOUR CODE HERE
raise NotImplementedError()
# gradient step
# YOUR CODE HERE
raise NotImplementedError()
# draw the current decision boundary every 250 examples seen
if it % 250 == 0 :
plot_decision_boundary(ax, X,Y, net)
fig.canvas.draw()
plot_decision_boundary(ax, X,Y, net)
fig.canvas.draw()
%matplotlib inline
plt.plot(losses)
```
## 3. Using a Pytorch module
In this last part, use `toch.nn.Module` to recode `MyLinear` and `MyReLU` so that these modules will be pytorch compatible.
```
import torch
import torch.nn as nn
# y = Wx + b
class MyLinear_mod(nn.Module):
def __init__(self, n_input, n_output):
super(MyLinear_mod, self).__init__()
# define self.A and self.b the weights and biases
# initialize them with a normal distribution
# use nn.Parameters
# YOUR CODE HERE
raise NotImplementedError()
def forward(self, x):
# YOUR CODE HERE
raise NotImplementedError()
class MyReLU_mod(nn.Module):
def __init__(self):
super(MyReLU_mod, self).__init__()
def forward(self, x):
# YOUR CODE HERE
raise NotImplementedError()
# the grid for plotting the decision boundary should be now made of tensors.
to_forward = torch.from_numpy(np.array(list(zip(xx.ravel(), yy.ravel())))).float()
```
Define your network using `MyLinear_mod`, `MyReLU_mod` and [`nn.Sigmoid`](https://pytorch.org/docs/stable/nn.html#sigmoid)
```
h=50
# define your network with nn.Sequential
# use MyLinear_mod, MyReLU_mod and nn.Sigmoid (from pytorch)
# YOUR CODE HERE
raise NotImplementedError()
from torch import optim
optimizer = optim.SGD(net.parameters(), lr=1e-2)
X_torch = torch.from_numpy(X).float()
Y_torch = torch.from_numpy(Y).float()
# you should comment the following line if on colab
%matplotlib notebook
fig, ax = plt.subplots(1, 1, facecolor='#4B6EA9')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
losses = []
criterion = nn.BCELoss()
for it in range(10000):
# pick a random example id
j = np.random.randint(1, len(X))
# select the corresponding example and label
example = X_torch[j:j+1]
label = Y_torch[j:j+1].unsqueeze(1)
# do a forward pass on the example
# YOUR CODE HERE
raise NotImplementedError()
# compute the loss according to your output and the label
# YOUR CODE HERE
raise NotImplementedError()
# zero the gradients
# YOUR CODE HERE
raise NotImplementedError()
# backward pass
# YOUR CODE HERE
raise NotImplementedError()
# gradient step
# YOUR CODE HERE
raise NotImplementedError()
# draw the current decision boundary every 250 examples seen
if it % 250 == 0 :
plot_decision_boundary(ax, X,Y, net)
fig.canvas.draw()
plot_decision_boundary(ax, X,Y, net)
fig.canvas.draw()
%matplotlib inline
plt.plot(losses)
```
| github_jupyter |
```
import numpy as np
import tensorflow as tf
import sys
sys.path.insert(1, '../libs')
from functions import *
from utils import *
STYLE_WEIGHTS = [3.0]
CONTENT_WEIGHTS = [1.0]
MODEL_SAVE_PATHS = ['../../models/style_weight_2e0.ckpt']
# for inferring (stylize)
INFERRING_CONTENT_DIR = '../../_inference/content'
INFERRING_STYLE_DIR = '../../_inference/style'
OUTPUTS_DIR = '../../_inference/output'
ENCODER_WEIGHTS_PATH = '../../vgg19_normalised.npz'
model_save_path = MODEL_SAVE_PATHS
content_imgs_path = list_images(INFERRING_CONTENT_DIR)
style_imgs_path = list_images(INFERRING_STYLE_DIR)
contents_path = content_imgs_path
styles_path = style_imgs_path
output_dir = OUTPUTS_DIR
encoder_path = ENCODER_WEIGHTS_PATH
model_path = model_save_path
resize_height=None
resize_width=None
content = tf.placeholder(tf.float32, shape=(1, None, None, 3), name='content')
style = tf.placeholder(tf.float32, shape=(1, None, None, 3), name='style')
tf.reshape(content, shape=tf.shape(style))
if isinstance(contents_path, str):
contents_path = [contents_path]
if isinstance(styles_path, str):
styles_path = [styles_path]
with tf.Graph().as_default(), tf.Session() as sess:
# build the dataflow graph
content = tf.placeholder(tf.float32, shape=(1, None, None, 3), name='content')
style = tf.placeholder(tf.float32, shape=(1, None, None, 3), name='style')
# content = tf.placeholder(tf.float32, shape=INPUT_SHAPE, name='content')
# style = tf.placeholder(tf.float32, shape=INPUT_SHAPE, name='style')
stn = STNet(encoder_path)
output_image = stn.transform(content, style)
sess.run(tf.global_variables_initializer())
# restore the trained model and run the style transferring
saver = tf.train.Saver()
saver.restore(sess, model_path)
outputs = []
for content_path in contents_path:
content_img = get_images(content_path,
height=resize_height, width=resize_width)
for style_path in styles_path:
style_img = get_images(style_path)
result = sess.run(output_image,
feed_dict={content: content_img, style: style_img})
outputs.append(result[0])
save_images(outputs, contents_path, styles_path, output_dir, suffix=suffix)
class SANet:
'''
Style-Attentional Network learns the mapping
between the content features and the style features
by slightly modifying the self-attention mechanism
'''
def __init__(self, num_filter):
self.num_filter = num_filter
# def map(self, content, style, scope='attention'):
# with tf.variable_scope(scope, reuse = tf.AUTO_REUSE):
# f = conv(content, self.num_filter // 8, kernel=1, stride=1, scope='f_conv') # [bs, h, w, c']
# g = conv(style, self.num_filter // 8, kernel=1, stride=1, scope='g_conv') # [bs, h, w, c']
# h = conv(style, self.num_filter , kernel=1, stride=1, scope='h_conv') # [bs, h, w, c]
# # N = h * w
# s = tf.matmul(hw_flatten(g), hw_flatten(f), transpose_b=True) # # [bs, N, N]
# attention = tf.nn.softmax(s) # attention map
# o = tf.matmul(attention, hw_flatten(h)) # [bs, N, C]
# # gamma = tf.get_variable("gamma", [1], initializer=tf.constant_initializer(0.0))
# o = tf.reshape(o, shape=content.shape) # [bs, h, w, C]
# o = conv(o, self.num_filter, kernel=1, stride=1, scope='attn_conv')
# # o = gamma * o + content
# o = o + content
# return o
def map(self, content, style, scope='attention'):
with tf.variable_scope(scope, reuse = tf.AUTO_REUSE):
f = conv(content, self.num_filter // 8, kernel=1, stride=1, scope='f_conv') # [bs, h, w, c']
g = conv(style, self.num_filter // 8, kernel=1, stride=1, scope='g_conv') # [bs, h, w, c']
h = conv(style, self.num_filter , kernel=1, stride=1, scope='h_conv') # [bs, h, w, c]
s = tf.matmul(g, f, transpose_b=True)
attention = tf.nn.softmax(s)
o = tf.matmul(attention, h)
# o = tf.reshape(o, shape=content.shape)
o = conv(o, self.num_filter, kernel=1, stride=1, scope='attn_conv')
o = o + content
return o
```
| github_jupyter |
# The Implicit Backward Time Centered Space (BTCS) Difference Equation for the Heat Equation
#### John S Butler john.s.butler@tudublin.ie [Course Notes](https://johnsbutler.netlify.com/files/Teaching/Numerical_Analysis_for_Differential_Equations.pdf) [Github](https://github.com/john-s-butler-dit/Numerical-Analysis-Python)
## Overview
This notebook will implement the implicit Backward Time Centered Space (FTCS) Difference method for the Heat Equation.
## The Heat Equation
The Heat Equation is the first order in time ($t$) and second order in space ($x$) Partial Differential Equation:
$$ \frac{\partial u}{\partial t} = \frac{\partial^2 u}{\partial x^2},$$
the equation describes heat transfer on a domain
$$ \Omega = \{ t \geq, 0\leq x \leq 1\}. $$
with an initial condition at time $t=0$ for all $x$ and boundary condition on the left ($x=0$) and right side ($x=1$).
## Backward Time Centered Space (BTCS) Difference method
This notebook will illustrate the Backward Time Centered Space (BTCS) Difference method for the Heat Equation with the __initial conditions__
$$ u(x,0)=2x, \ \ 0 \leq x \leq \frac{1}{2}, $$
$$ u(x,0)=2(1-x), \ \ \frac{1}{2} \leq x \leq 1, $$
and __boundary condition__
$$ u(0,t)=0, u(1,t)=0. $$
```
# LIBRARY
# vector manipulation
import numpy as np
# math functions
import math
# THIS IS FOR PLOTTING
%matplotlib inline
import matplotlib.pyplot as plt # side-stepping mpl backend
import warnings
warnings.filterwarnings("ignore")
```
## Discete Grid
The region $\Omega$ is discretised into a uniform mesh $\Omega_h$. In the space $x$ direction into $N$ steps giving a stepsize of
$$ h=\frac{1-0}{N},$$
resulting in
$$x[i]=0+ih, \ \ \ i=0,1,...,N,$$
and into $N_t$ steps in the time $t$ direction giving a stepsize of
$$ k=\frac{1-0}{N_t}$$
resulting in
$$t[i]=0+ik, \ \ \ k=0,...,K.$$
The Figure below shows the discrete grid points for $N=10$ and $Nt=100$, the known boundary conditions (green), initial conditions (blue) and the unknown values (red) of the Heat Equation.
```
N=10
Nt=100
h=1/N
k=1/Nt
r=k/(h*h)
time_steps=15
time=np.arange(0,(time_steps+.5)*k,k)
x=np.arange(0,1.0001,h)
X, Y = np.meshgrid(x, time)
fig = plt.figure()
plt.plot(X,Y,'ro');
plt.plot(x,0*x,'bo',label='Initial Condition');
plt.plot(np.ones(time_steps+1),time,'go',label='Boundary Condition');
plt.plot(x,0*x,'bo');
plt.plot(0*time,time,'go');
plt.xlim((-0.02,1.02))
plt.xlabel('x')
plt.ylabel('time (ms)')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.title(r'Discrete Grid $\Omega_h,$ h= %s, k=%s'%(h,k),fontsize=24,y=1.08)
plt.show();
```
## Discrete Initial and Boundary Conditions
The discrete initial conditions are
$$ w[i,0]=2x[i], \ \ 0 \leq x[i] \leq \frac{1}{2} $$
$$ w[i,0]=2(1-x[i]), \ \ \frac{1}{2} \leq x[i] \leq 1 $$
and the discete boundary conditions are
$$ w[0,j]=0, w[10,j]=0, $$
where $w[i,j]$ is the numerical approximation of $U(x[i],t[j])$.
The Figure below plots values of $w[i,0]$ for the inital (blue) and boundary (red) conditions for $t[0]=0.$
```
w=np.zeros((N+1,time_steps+1))
b=np.zeros(N-1)
# Initial Condition
for i in range (1,N):
w[i,0]=2*x[i]
if x[i]>0.5:
w[i,0]=2*(1-x[i])
# Boundary Condition
for k in range (0,time_steps):
w[0,k]=0
w[N,k]=0
fig = plt.figure(figsize=(8,4))
plt.plot(x,w[:,0],'o:',label='Initial Condition')
plt.plot(x[[0,N]],w[[0,N],0],'go',label='Boundary Condition t[0]=0')
#plt.plot(x[N],w[N,0],'go')
plt.xlim([-0.1,1.1])
plt.ylim([-0.1,1.1])
plt.title('Intitial and Boundary Condition',fontsize=24)
plt.xlabel('x')
plt.ylabel('w')
plt.legend(loc='best')
plt.show()
```
## The Implicit Backward Time Centered Space (BTCS) Difference Equation
The implicit Backward Time Centered Space (BTCS) difference equation of the Heat Equation is derived by discretising
$$ \frac{\partial u_{ij+1}}{\partial t} = \frac{\partial^2 u_{ij+1}}{\partial x^2},$$
around $(x_i,t_{j+1})$ giving the difference equation
$$
\frac{w_{ij+1}-w_{ij}}{k}=\frac{w_{i+1j+1}-2w_{ij+1}+w_{i-1j+1}}{h^2}
$$
Rearranging the equation we get
$$
-rw_{i-1j+1}+(1+2r)w_{ij+1}-rw_{i+1j+1}=w_{ij}
$$
for $i=1,...9$ where $r=\frac{k}{h^2}$.
This gives the formula for the unknown term $w_{ij+1}$ at the $(ij+1)$ mesh points
in terms of $x[i]$ along the jth time row.
Hence we can calculate the unknown pivotal values of $w$ along the first row of $j=1$ in terms of the known boundary conditions.
This can be written in matrix form
$$ A\mathbf{w}_{j+1}=\mathbf{w}_{j} +\mathbf{b}_{j+1} $$
for which $A$ is a $9\times9$ matrix:
$$
\left(\begin{array}{cccc cccc}
1+2r&-r& 0&0&0 &0&0&0\\
-r&1+2r&-r&0&0&0 &0&0&0\\
0&-r&1+2r &-r&0&0& 0&0&0\\
0&0&-r&1+2r &-r&0&0& 0&0\\
0&0&0&-r&1+2r &-r&0&0& 0\\
0&0&0&0&-r&1+2r &-r&0&0\\
0&0&0&0&0&-r&1+2r &-r&0\\
0&0&0&0&0&0&-r&1+2r&-r\\
0&0&0&0&0&0&0&-r&1+2r\\
\end{array}\right)
\left(\begin{array}{c}
w_{1j+1}\\
w_{2j+1}\\
w_{3j+1}\\
w_{4j+1}\\
w_{5j+1}\\
w_{6j+1}\\
w_{7j+1}\\
w_{8j+1}\\
w_{9j+1}\\
\end{array}\right)=
\left(\begin{array}{c}
w_{1j}\\
w_{2j}\\
w_{3j}\\
w_{4j}\\
w_{5j}\\
w_{6j}\\
w_{7j}\\
w_{8j}\\
w_{9j}\\
\end{array}\right)+
\left(\begin{array}{c}
rw_{0j+1}\\
0\\
0\\
0\\
0\\
0\\
0\\
0\\
rw_{10j+1}\\
\end{array}\right).
$$
It is assumed that the boundary values $w_{0j+1}$ and $w_{10j+1}$ are known for $j=1,2,...$, and $w_{i0}$ for $i=0,...,10$ is the initial condition.
The Figure below shows the values of the $9\times 9$ matrix in colour plot form for $r=\frac{k}{h^2}$.
```
A=np.zeros((N-1,N-1))
for i in range (0,N-1):
A[i,i]=1+2*r
for i in range (0,N-2):
A[i+1,i]=-r
A[i,i+1]=-r
Ainv=np.linalg.inv(A)
fig = plt.figure(figsize=(12,4));
plt.subplot(121)
plt.imshow(A,interpolation='none');
plt.xticks(np.arange(N-1), np.arange(1,N-0.9,1));
plt.yticks(np.arange(N-1), np.arange(1,N-0.9,1));
clb=plt.colorbar();
clb.set_label('Matrix elements values');
clb.set_clim((-1,1));
plt.title('Matrix A r=%s'%(np.round(r,3)),fontsize=24)
plt.subplot(122)
plt.imshow(Ainv,interpolation='none');
plt.xticks(np.arange(N-1), np.arange(1,N-0.9,1));
plt.yticks(np.arange(N-1), np.arange(1,N-0.9,1));
clb=plt.colorbar();
clb.set_label('Matrix elements values');
clb.set_clim((-1,1));
plt.title(r'Matrix $A^{-1}$ r=%s'%(np.round(r,3)),fontsize=24)
fig.tight_layout()
plt.show();
```
## Results
To numerically approximate the solution at $t[1]$ the matrix equation becomes
$$ \mathbf{w}_{1}=A^{-1}(\mathbf{w}_{0} +\mathbf{b}_{0}) $$
where all the right hand side is known.
To approximate solution at time $t[2]$ we use the matrix equation
$$ \mathbf{w}_{2}=A^{-1}(\mathbf{w}_{1} +\mathbf{b}_{1}). $$
Each set of numerical solutions $w[i,j]$ for all $i$ at the previous time step is used to approximate the solution $w[i,j+1]$.
The Figure below shows the numerical approximation $w[i,j]$ of the Heat Equation using the FTCS method at $x[i]$ for $i=0,...,10$ and time steps $t[j]$ for $j=1,...,15$. The left plot shows the numerical approximation $w[i,j]$ as a function of $x[i]$ with each color representing the different time steps $t[j]$. The right plot shows the numerical approximation $w[i,j]$ as colour plot as a function of $x[i]$, on the $x[i]$ axis and time $t[j]$ on the $y$ axis.
The solution is stable for $r>\frac{1}{2}$ unlike in the explicit method.
```
fig = plt.figure(figsize=(12,6))
plt.subplot(121)
for j in range (1,time_steps+1):
b[0]=r*w[0,j]
b[N-2]=r*w[N,j]
w[1:(N),j]=np.dot(Ainv,w[1:(N),j-1]+b)
plt.plot(x,w[:,j],'o:',label='t[%s]=%s'%(j,time[j]))
plt.xlabel('x')
plt.ylabel('w')
#plt.legend(loc='bottom', bbox_to_anchor=(0.5, -0.1))
plt.legend(bbox_to_anchor=(-.4, 1), loc=2, borderaxespad=0.)
plt.subplot(122)
plt.imshow(w.transpose())
plt.xticks(np.arange(len(x)), x)
plt.yticks(np.arange(len(time)), time)
plt.xlabel('x')
plt.ylabel('time')
clb=plt.colorbar()
clb.set_label('Temperature (w)')
plt.suptitle('Numerical Solution of the Heat Equation r=%s'%(np.round(r,3)),fontsize=24,y=1.08)
fig.tight_layout()
plt.show()
```
## Local Trunction Error
The local truncation error of the classical explicit difference approach to
\begin{equation}
\frac{\partial U}{\partial t} - \frac{\partial^2 U}{\partial x^2}=0,
\end{equation}
with
\begin{equation}
F_{ij+1}(w)=\frac{w_{ij+1}-w_{ij}}{k}-\frac{w_{i+1j+1}-2w_{ij+1}+w_{i-1j+1}}{h^2}=0,
\end{equation}
is
\begin{equation}
T_{ij+1}=F_{ij+1}(U)=\frac{U_{ij+1}-U_{ij}}{k}-\frac{U_{i+1j+1}-2U_{ij+1}+U_{i-1j+1}}{h^2},
\end{equation}
By Taylors expansions we have
\begin{eqnarray*}
U_{i+1j}&=&U((i+1)h,(j+1)k)=U(x_i+h,t_{j+1})\\
&=&U_{ij+1}+h\left(\frac{\partial U}{\partial x} \right)_{ij+1}+\frac{h^2}{2}\left(\frac{\partial^2 U}{\partial x^2} \right)_{ij+1}+\frac{h^3}{6}\left(\frac{\partial^3 U}{\partial x^3} \right)_{ij+1} +...\\
U_{i-1j}&=&U((i-1)h,(j+1)k)=U(x_i-h,t_{j+1})\\
&=&U_{ij+1}-h\left(\frac{\partial U}{\partial x} \right)_{ij+1}+\frac{h^2}{2}\left(\frac{\partial^2 U}{\partial x^2} \right)_{ij+1}-\frac{h^3}{6}\left(\frac{\partial^3 U}{\partial x^3} \right)_{ij+1} +...\\
U_{ij}&=&U(ih,(j)k)=U(x_i,t_j)\\
&=&U_{ij+1}-k\left(\frac{\partial U}{\partial t} \right)_{ij+1}+\frac{k^2}{2}\left(\frac{\partial^2 U}{\partial t^2} \right)_{ij+1}-\frac{k^3}{6}\left(\frac{\partial^3 U}{\partial t^3} \right)_{ij+1} +...
\end{eqnarray*}
substitution into the expression for $T_{ij+1}$ then gives
\begin{eqnarray*}
T_{ij+1}&=&\left(\frac{\partial U}{\partial t} - \frac{\partial^2 U}{\partial x^2} \right)_{ij+1}+\frac{k}{2}\left(\frac{\partial^2 U}{\partial t^2} \right)_{ij+1}
-\frac{h^2}{12}\left(\frac{\partial^4 U}{\partial x^4} \right)_{ij+1}\\
& & +\frac{k^2}{6}\left(\frac{\partial^3 U}{\partial t^3} \right)_{ij+1}
-\frac{h^4}{360}\left(\frac{\partial^6 U}{\partial x^6} \right)_{ij+1}+ ...
\end{eqnarray*}
But $U$ is the solution to the differential equation so
\begin{equation} \left(\frac{\partial U}{\partial t} - \frac{\partial^2 U}{\partial x^2} \right)_{ij+1}=0,\end{equation}
the principal part of the local truncation error is
\begin{equation}
\frac{k}{2}\left(\frac{\partial^2 U}{\partial t^2} \right)_{ij+1}-\frac{h^2}{12}\left(\frac{\partial^4 U}{\partial x^4} \right)_{ij+1}.
\end{equation}
Hence the truncation error is
\begin{equation}
T_{ij}=O(k)+O(h^2).
\end{equation}
## Stability Analyis
To investigating the stability of the fully implicit BTCS difference method of the Heat Equation, we will use the von Neumann method.
The FTCS difference equation is:
$$\frac{1}{k}(w_{pq+1}-w_{pq})=\frac{1}{h_x^2}(w_{p-1q+1}-2w_{pq+1}+w_{p+1q+1}),$$
approximating
$$\frac{\partial U}{\partial t}=\frac{\partial^2 U}{\partial x^2}$$
at $(ph,k(q+1))$. Substituting $w_{pq}=e^{i\beta x}\xi^{q}$ into the difference equation gives:
$$e^{i\beta ph}\xi^{q+1}-e^{i\beta ph}\xi^{q}=r\{e^{i\beta (p-1)h}\xi^{q+1}-2e^{i\beta ph}\xi^{q+1}+e^{i\beta (p+1)h}\xi^{q+1} \}
$$
where $r=\frac{k}{h_x^2}$. Divide across by $e^{i\beta (p)h}\xi^{q}$ leads to
$$ \xi-1=r \xi (e^{i\beta (-1)h} -2+e^{i\beta h}), $$
$$\xi-\xi r (2\cos(\beta h)-2)= 1,$$
$$\xi(1+4r\sin^2(\beta\frac{h}{2})) =1$$
Hence
$$\xi=\frac{1}{(1+4r\sin^2(\beta\frac{h}{2}))} \leq 1$$
therefore the equation is unconditionally stable as $0 < \xi \leq 1$ for all $r$ and all $\beta$ .
## References
[1] G D Smith Numerical Solution of Partial Differential Equations: Finite Difference Method Oxford 1992
[2] Butler, J. (2019). John S Butler Numerical Methods for Differential Equations. [online] Maths.dit.ie. Available at: http://www.maths.dit.ie/~johnbutler/Teaching_NumericalMethods.html [Accessed 14 Mar. 2019].
[3] Wikipedia contributors. (2019, February 22). Heat equation. In Wikipedia, The Free Encyclopedia. Available at: https://en.wikipedia.org/w/index.php?title=Heat_equation&oldid=884580138 [Accessed 14 Mar. 2019].
| github_jupyter |
```
from quchem_ibm.Qiskit_Chemistry import *
transformation='BK'
from quchem.Hamiltonian_Generator_Functions import *
from quchem.Graph import *
## HAMILTONIAN start
Molecule = 'LiH'
geometry = [('Li', (0., 0., 0.)), ('H', (0., 0., 1.45))]
basis = 'sto-6g'
### Get Hamiltonian
Hamilt = Hamiltonian_PySCF(Molecule,
run_scf=1, run_mp2=1, run_cisd=1, run_ccsd=1, run_fci=1,
basis=basis,
multiplicity=1,
geometry=geometry) # normally None!
QubitHamiltonian = Hamilt.Get_Qubit_Hamiltonian(threshold=None, transformation=transformation)
### HAMILTONIAN end
#####################################
print(QubitHamiltonian)
fci_energy = Hamilt.molecule.fci_energy
print(fci_energy)
```
From PHYS. REV. X, **8**, 031022 (2018):
$$LiH_{BK}^{HartreeFock} = | 101000000000\rangle$$
```
from quchem.Ansatz_Generator_Functions import *
n_electrons=Hamilt.molecule.n_electrons
n_qubits=Hamilt.molecule.n_qubits
ansatz_obj = Ansatz(n_electrons,n_qubits)
print('JW ground state = ', ansatz_obj.Get_JW_HF_state_in_OCC_basis())
print('BK ground state = ', ansatz_obj.Get_BK_HF_state_in_OCC_basis())
# qubits_to_remove = Find_I_Z_indices_in_Hamiltonian(QubitHamiltonian, Hamilt.molecule.n_qubits)
# print('qubits only acted on by I or Z:', qubits_to_remove)
# input_state = ansatz_obj.Get_BK_HF_state_in_OCC_basis() if transformation=='BK' else ansatz_obj.Get_JW_HF_state_in_OCC_basis()
# # Remove_Z_terms_from_Hamiltonian
# NewQubitHamiltonian = Remove_Z_terms_from_Hamiltonian(
# QubitHamiltonian,
# input_state,
# qubits_to_remove,
# check_reduction=True)
# NewQubitHamiltonian
# qubitNo_re_label_dict, NewQubitHamiltonian_relabelled = Re_label_Hamiltonian(NewQubitHamiltonian)
# NewQubitHamiltonian_relabelled
NewQubitHamiltonian_relabelled=QubitHamiltonian
```
# Find what new FCI energy is
- here should be the same as true answer!
```
from openfermion import qubit_operator_sparse
from scipy.sparse import csc_matrix
new_Molecular_H_MATRIX = csc_matrix(qubit_operator_sparse(NewQubitHamiltonian_relabelled))
# new_Molecular_H_MATRIX = np.flip(new_Molecular_H_MATRIX)
from scipy.sparse.linalg import eigs
try:
eig_values, eig_vectors = eigs(new_Molecular_H_MATRIX)
except:
from scipy.linalg import eig
eig_values, eig_vectors = eig(new_Molecular_H_MATRIX.todense())
new_FCI_Energy = min(eig_values)
index = np.where(eig_values==new_FCI_Energy)[0][0]
ground_state_vector = eig_vectors[:, index]
print('new_FCI = ', new_FCI_Energy, 'VS old FCI:', fci_energy)
print(np.isclose(new_FCI_Energy, fci_energy))
np.dot(ground_state_vector.conj().T, new_Molecular_H_MATRIX.dot(ground_state_vector))
```
# Ansatz Circuit
EMTPY!!!!!!!!!!
```
q_reg = QuantumRegister(12)
Ansatz_circuit = QuantumCircuit(q_reg)
# Ansatz_circuit.draw()
```
# Standard VQE
```
from tqdm.notebook import tqdm
standard_VQE_circuits, standard_I_term = Build_Standard_VQE_circuits_MEMORY_EFF(
NewQubitHamiltonian_relabelled,
Ansatz_circuit,
q_reg)
j=200
print(standard_VQE_circuits[j]['qubitOp'])
circ = QuantumCircuit.from_qasm_str(standard_VQE_circuits[j]['circuit'])
circ.draw()
```
# Graph
```
from tqdm.notebook import tqdm
Hamiltonian_graph_obj = Openfermion_Hamiltonian_Graph(NewQubitHamiltonian_relabelled)
commutativity_flag = 'AC' ## <- defines relationship between sets!!!
plot_graph = False
Graph_colouring_strategy='largest_first'
anti_commuting_sets = Hamiltonian_graph_obj.Get_Clique_Cover_as_QubitOp(commutativity_flag, Graph_colouring_strategy=Graph_colouring_strategy, plot_graph=plot_graph)
anti_commuting_sets
```
# Seq Rot circuits
```
n_qubits= len(new_input_state)
rotation_reduction_check=False
Seq_Rot_VQE_circuits, Seq_Rot_I_term = Get_Seq_Rot_Unitary_Part_circuits_MEMORY_EFF(
anti_commuting_sets,
Ansatz_circuit,
q_reg,
n_qubits,
S_index_dict=None,
rotation_reduction_check=rotation_reduction_check)
j=3
circ = QuantumCircuit.from_qasm_str(Seq_Rot_VQE_circuits[j]['circuit'])
circ.draw()
# n_qubits= len(new_input_state)
# rotation_reduction_check=False
# Seq_Rot_VQE_circuits, Seq_Rot_I_term = Get_Seq_Rot_Unitary_Part_circuits(
# anti_commuting_sets,
# Ansatz_circuit,
# q_reg,
# n_qubits,
# S_index_dict=None,
# rotation_reduction_check=rotation_reduction_check)
```
# LCU circuits
```
# n_qubits= len(new_input_state)
# check_ansatz_state = False
# LCU_VQE_circuits, LCU_I_term= Get_LCU_Unitary_Part_circuits(anti_commuting_sets, ground_state_vector, n_qubits,
# N_index_dict=None,check_ansatz_state=check_ansatz_state)
n_qubits= len(new_input_state)
LCU_VQE_circuits, LCU_I_term=Get_LCU_Unitary_Part_circuits_MEMORY_EFF(
anti_commuting_sets,
Ansatz_circuit,
q_reg,
n_qubits,
N_index_dict=None)
```
# Save experiment input
```
list(NewQubitHamiltonian_relabelled)
```
| github_jupyter |
##### Copyright 2018 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Transfer Learning with TensorFlow Hub for TFLite
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_lite/tflite_c02_transfer_learning.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_lite/tflite_c02_transfer_learning.ipynb">
<img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
View source on GitHub</a>
</td>
</table>
## Setup
```
import os
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
print("Version: ", tf.__version__)
print("Eager mode: ", tf.executing_eagerly())
print("Hub version: ", hub.__version__)
print("GPU is", "available" if tf.config.list_physical_devices('GPU') else "NOT AVAILABLE")
```
## Select the Hub/TF2 module to use
Hub modules for TF 1.x won't work here, please use one of the selections provided.
```
module_selection = ("mobilenet_v2", 224, 1280) #@param ["(\"mobilenet_v2\", 224, 1280)", "(\"inception_v3\", 299, 2048)"] {type:"raw", allow-input: true}
handle_base, pixels, FV_SIZE = module_selection
MODULE_HANDLE ="https://tfhub.dev/google/tf2-preview/{}/feature_vector/4".format(handle_base)
IMAGE_SIZE = (pixels, pixels)
print("Using {} with input size {} and output dimension {}".format(
MODULE_HANDLE, IMAGE_SIZE, FV_SIZE))
```
## Data preprocessing
Use [TensorFlow Datasets](http://tensorflow.org/datasets) to load the cats and dogs dataset.
This `tfds` package is the easiest way to load pre-defined data. If you have your own data, and are interested in importing using it with TensorFlow see [loading image data](../load_data/images.ipynb)
```
import tensorflow_datasets as tfds
tfds.disable_progress_bar()
```
The `tfds.load` method downloads and caches the data, and returns a `tf.data.Dataset` object. These objects provide powerful, efficient methods for manipulating data and piping it into your model.
Since `"cats_vs_dogs"` doesn't define standard splits, use the subsplit feature to divide it into (train, validation, test) with 80%, 10%, 10% of the data respectively.
```
(train_examples, validation_examples, test_examples), info = tfds.load(
'cats_vs_dogs',
split=['train[80%:]', 'train[80%:90%]', 'train[90%:]'],
with_info=True,
as_supervised=True,
)
num_examples = info.splits['train'].num_examples
num_classes = info.features['label'].num_classes
```
### Format the Data
Use the `tf.image` module to format the images for the task.
Resize the images to a fixes input size, and rescale the input channels
```
def format_image(image, label):
image = tf.image.resize(image, IMAGE_SIZE) / 255.0
return image, label
```
Now shuffle and batch the data
```
BATCH_SIZE = 32 #@param {type:"integer"}
train_batches = train_examples.shuffle(num_examples // 4).map(format_image).batch(BATCH_SIZE).prefetch(1)
validation_batches = validation_examples.map(format_image).batch(BATCH_SIZE).prefetch(1)
test_batches = test_examples.map(format_image).batch(1)
```
Inspect a batch
```
for image_batch, label_batch in train_batches.take(1):
pass
image_batch.shape
```
## Defining the model
All it takes is to put a linear classifier on top of the `feature_extractor_layer` with the Hub module.
For speed, we start out with a non-trainable `feature_extractor_layer`, but you can also enable fine-tuning for greater accuracy.
```
do_fine_tuning = False #@param {type:"boolean"}
```
Load TFHub Module
```
feature_extractor = hub.KerasLayer(MODULE_HANDLE,
input_shape=IMAGE_SIZE + (3,),
output_shape=[FV_SIZE],
trainable=do_fine_tuning)
print("Building model with", MODULE_HANDLE)
model = tf.keras.Sequential([
feature_extractor,
tf.keras.layers.Dense(num_classes)
])
model.summary()
#@title (Optional) Unfreeze some layers
NUM_LAYERS = 7 #@param {type:"slider", min:1, max:50, step:1}
if do_fine_tuning:
feature_extractor.trainable = True
for layer in model.layers[-NUM_LAYERS:]:
layer.trainable = True
else:
feature_extractor.trainable = False
```
## Training the model
```
if do_fine_tuning:
model.compile(
optimizer=tf.keras.optimizers.SGD(lr=0.002, momentum=0.9),
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
else:
model.compile(
optimizer='adam',
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
EPOCHS = 5
hist = model.fit(train_batches,
epochs=EPOCHS,
validation_data=validation_batches)
```
## Export the model
```
CATS_VS_DOGS_SAVED_MODEL = "exp_saved_model"
```
Export the SavedModel
```
tf.saved_model.save(model, CATS_VS_DOGS_SAVED_MODEL)
%%bash -s $CATS_VS_DOGS_SAVED_MODEL
saved_model_cli show --dir $1 --tag_set serve --signature_def serving_default
loaded = tf.saved_model.load(CATS_VS_DOGS_SAVED_MODEL)
print(list(loaded.signatures.keys()))
infer = loaded.signatures["serving_default"]
print(infer.structured_input_signature)
print(infer.structured_outputs)
```
## Convert using TFLite's Converter
Load the TFLiteConverter with the SavedModel
```
converter = tf.lite.TFLiteConverter.from_saved_model(CATS_VS_DOGS_SAVED_MODEL)
```
### Post-training quantization
The simplest form of post-training quantization quantizes weights from floating point to 8-bits of precision. This technique is enabled as an option in the TensorFlow Lite converter. At inference, weights are converted from 8-bits of precision to floating point and computed using floating-point kernels. This conversion is done once and cached to reduce latency.
To further improve latency, hybrid operators dynamically quantize activations to 8-bits and perform computations with 8-bit weights and activations. This optimization provides latencies close to fully fixed-point inference. However, the outputs are still stored using floating point, so that the speedup with hybrid ops is less than a full fixed-point computation.
```
converter.optimizations = [tf.lite.Optimize.DEFAULT]
```
### Post-training integer quantization
We can get further latency improvements, reductions in peak memory usage, and access to integer only hardware accelerators by making sure all model math is quantized. To do this, we need to measure the dynamic range of activations and inputs with a representative data set. You can simply create an input data generator and provide it to our converter.
```
def representative_data_gen():
for input_value, _ in test_batches.take(100):
yield [input_value]
converter.representative_dataset = representative_data_gen
```
The resulting model will be fully quantized but still take float input and output for convenience.
Ops that do not have quantized implementations will automatically be left in floating point. This allows conversion to occur smoothly but may restrict deployment to accelerators that support float.
### Full integer quantization
To require the converter to only output integer operations, one can specify:
```
converter.target_spec.supported_ops = [tf.lite.OpsSet.TFLITE_BUILTINS_INT8]
```
### Finally convert the model
```
tflite_model = converter.convert()
tflite_model_file = 'converted_model.tflite'
with open(tflite_model_file, "wb") as f:
f.write(tflite_model)
```
##Test the TFLite model using the Python Interpreter
```
# Load TFLite model and allocate tensors.
interpreter = tf.lite.Interpreter(model_path=tflite_model_file)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]["index"]
output_index = interpreter.get_output_details()[0]["index"]
from tqdm import tqdm
# Gather results for the randomly sampled test images
predictions = []
test_labels, test_imgs = [], []
for img, label in tqdm(test_batches.take(10)):
interpreter.set_tensor(input_index, img)
interpreter.invoke()
predictions.append(interpreter.get_tensor(output_index))
test_labels.append(label.numpy()[0])
test_imgs.append(img)
#@title Utility functions for plotting
# Utilities for plotting
class_names = ['cat', 'dog']
def plot_image(i, predictions_array, true_label, img):
predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
plt.grid(False)
plt.xticks([])
plt.yticks([])
img = np.squeeze(img)
plt.imshow(img, cmap=plt.cm.binary)
predicted_label = np.argmax(predictions_array)
if predicted_label == true_label:
color = 'green'
else:
color = 'red'
plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
100*np.max(predictions_array),
class_names[true_label]),
color=color)
```
NOTE: Colab runs on server CPUs. At the time of writing this, TensorFlow Lite doesn't have super optimized server CPU kernels. For this reason post-training full-integer quantized models may be slower here than the other kinds of optimized models. But for mobile CPUs, considerable speedup can be observed.
```
#@title Visualize the outputs { run: "auto" }
index = 0 #@param {type:"slider", min:0, max:9, step:1}
plt.figure(figsize=(6,3))
plt.subplot(1,2,1)
plot_image(index, predictions, test_labels, test_imgs)
plt.show()
```
Download the model.
**NOTE: You might have to run to the cell below twice**
```
labels = ['cat', 'dog']
with open('labels.txt', 'w') as f:
f.write('\n'.join(labels))
try:
from google.colab import files
files.download('converted_model.tflite')
files.download('labels.txt')
except:
pass
```
# Prepare the test images for download (Optional)
This part involves downloading additional test images for the Mobile Apps only in case you need to try out more samples
```
!mkdir -p test_images
from PIL import Image
for index, (image, label) in enumerate(test_batches.take(50)):
image = tf.cast(image * 255.0, tf.uint8)
image = tf.squeeze(image).numpy()
pil_image = Image.fromarray(image)
pil_image.save('test_images/{}_{}.jpg'.format(class_names[label[0]], index))
!ls test_images
!zip -qq cats_vs_dogs_test_images.zip -r test_images/
try:
files.download('cats_vs_dogs_test_images.zip')
except:
pass
```
| github_jupyter |
# The Jones Vector
**Scott Prahl**
*April 2020*
```
import numpy as np
import matplotlib.pyplot as plt
import pypolar.jones as jones
import pypolar.visualization as vis
np.set_printoptions(suppress=True) # to print 1e-16 as zero
```
## Introduction
This notebook generates Jones vectors for different polarization states and derives parameters from them. It shows that `pypolar.jones` produces results that match those from standard references.
Complete details of the assumptions can be found in [Jupyter notebook on Conventions](./02-Jones-Conventions.html)
Resources used or mentioned in this notebook.
* Jones, "A New Calculus for the Treatment of Optical Systems", *JOSA*, **38**, 681 (1948).
* Azzam, *Ellipsometry and Polarized Light*, 1977.
* Collett, *Field Guide to Polarization*, 2005.
* Fowles, *Introduction to Modern Optics*, 1975.
* Goldstein, *Polarized Light*, 2003.
* Kliger, *Polarized Light in Optics and Spectroscopy*, 1990
* Shurcliff, *Polarized Light Production and Use*, 1962.
* Tompkins, *Handbook of Ellipsometry*, 2005.
### Comparison with Kliger Appendix A page 275
Here the Jones vectors (and polarization angles) are
```
def printit(J):
alpha=np.degrees(jones.ellipse_azimuth(J))
tanomega = jones.ellipticity(J)
beta = np.degrees(jones.amplitude_ratio_angle(J))
delta = np.degrees(jones.phase(J))
print("%6.2f %6.3f %6.2f %7.2f [%13s, %13s]" %
(alpha, tanomega,beta,delta,
J[0].__format__('.3f'),J[1].__format__('.3f')))
print("alpha tan(omega) beta ϕy-ϕx Standard Normalized")
light = jones.field_horizontal()
#print("Jones vector for horizontally-polarized light")
printit(light)
light = jones.field_vertical()
#print("Jones vector for vertically-polarized light")
printit(light)
light = jones.field_linear(np.radians(45))
#print("Jones vector for 45° linearly polarized light")
printit(light)
light = jones.field_linear(np.radians(-45))
#print("Jones vector for -45° linearly polarized light")
printit(light)
light = jones.field_linear(np.radians(30))
#print("Jones vector for 30° linearly polarized light")
printit(light)
light = jones.field_linear(np.radians(-60))
#print("Jones vector for -60° linearly polarized light")
printit(light)
light = jones.field_right_circular()
#print("Jones vector for right circularly polarized light")
printit(light)
light = jones.field_left_circular()
#print("Jones vector for left circularly polarized light")
printit(light)
J = 1/np.sqrt(5)*np.array([2,1j])
printit(J)
J = 1/np.sqrt(5)*np.array([2,-1j])
printit(J)
J = 1/np.sqrt(5)*np.array([1,2j])
printit(J)
J = 1/np.sqrt(5)*np.array([1,-2j])
printit(J)
J = 1/2*np.array([np.sqrt(2),1+1j])
printit(J)
J = 1/2*np.array([np.sqrt(2),1-1j])
printit(J)
J = 1/2/np.sqrt(2)*np.array([np.sqrt(6),1+1j])
printit(J)
J = np.sqrt(6)/4*np.array([2/np.sqrt(6),-1-1j])
printit(J)
```
### Comparison with Shurcliff Table 2.1 page 23
```
def printit(J):
alpha=np.degrees(jones.ellipse_azimuth(J))
tanomega = jones.ellipticity(J)
ratio = jones.amplitude_ratio(J)
delta = np.degrees(jones.phase(J))
print("%6.2f %6.3f %7.2f %7.2f [%13s, %13s]" %
(alpha, tanomega,ratio,delta,
J[0].__format__('.3f'),J[1].__format__('.3f')))
print(" tilt b/a Eyo/Exo ϕy-ϕx Standard Normalized")
J = np.array([1,0])
printit(J)
J = np.array([0,1])
printit(J)
J = 1/np.sqrt(2)*np.array([1,1])
printit(J)
J = 1/np.sqrt(2)*np.array([1,-1])
printit(J)
J = 1/np.sqrt(2)*np.array([-1j,1])
printit(J)
J = 1/np.sqrt(2)*np.array([1j,1])
printit(J)
J = 1/np.sqrt(5)*np.array([-2j,1])
printit(J)
J = 1/np.sqrt(5)*np.array([-1j,2])
printit(J)
J = 0.325*np.array([2.73,1+1j])
printit(J)
```
### Comparison with Wikipedia or Fowles (page 34)
These treatments use $e^{kz-\omega t}$ we need to use `jones.use_alternate_convention(True)` to account for this different convention.
```
def printit(J):
alpha=np.degrees(jones.ellipse_azimuth(J))
tanomega = jones.ellipticity(J)
ratio = jones.amplitude_ratio(J)
delta = np.degrees(jones.phase(J))
print("%6.2f %6.3f %7.2f %7.2f [%13s, %13s]" %
(alpha, tanomega,ratio,delta,
J[0].__format__('.3f'),J[1].__format__('.3f')))
# to account for different sign convention
jones.use_alternate_convention(True)
print(" tilt b/a Eyo/Exo ϕy-ϕx Standard Normalized")
light = jones.field_horizontal()
print("Jones vector for horizontally-polarized light")
printit(light)
light = jones.field_vertical()
print("Jones vector for vertically-polarized light")
printit(light)
light = jones.field_linear(np.radians(45))
print("Jones vector for 45° linearly polarized light")
printit(light)
light = jones.field_linear(np.radians(-45))
print("Jones vector for -45° linearly polarized light")
printit(light)
light = jones.field_right_circular()
print("Jones vector for right circularly polarized light")
printit(light)
light = jones.field_left_circular()
print("Jones vector for left circularly polarized light")
printit(light)
jones.use_alternate_convention(False)
```
## Round tripping through code
This tests construction and deconstruction of Jones vectors.
```
def testit(azimuth, ellipticity_angle, phi_x, E0):
J = jones.field_elliptical(azimuth, ellipticity_angle, phi_x, E0)
alpha = jones.ellipse_azimuth(J)
phix = np.angle(J[0])
epsilon = jones.ellipticity_angle(J)
delta = jones.phase(J)
Ex0, Ey0 = np.abs(J)
e0 = np.sqrt(Ex0**2+Ey0**2)
print("%6.2f %6.2f " % (np.degrees(azimuth),np.degrees(alpha)), end='')
print("%6.2f %6.2f " % (np.degrees(ellipticity_angle),np.degrees(epsilon)), end='')
print("%6.2f %6.2f " % (np.degrees(phi_x),np.degrees(phix)), end='')
print("%6.2f %6.2f " % (E0,e0))
print("azimuth calc epsilon calc phi_x calc E0 calc")
ellipticity_angle = np.radians(12)
phi_x = np.radians(-17)
E0 = 3
for az in [-89, -30, -40, 0, 40]:
azimuth = np.radians(az)
testit(azimuth, ellipticity_angle, phi_x, E0)
print()
azimuth = np.radians(55)
E0 = 5
for p in [-89, -30, -40, 0, 40]:
phi_x = np.radians(p)
testit(azimuth, ellipticity_angle, phi_x, E0)
print()
azimuth = np.radians(-15)
phi_x = np.radians(17)
E0 = 0.5
for p in [-44, -30, 0, 15, 40]:
ellipticity_angle = np.radians(p)
testit(azimuth, ellipticity_angle, phi_x, E0)
```
## Intensities
```
light = jones.field_horizontal()
inten = jones.intensity(light)
print("Intensity for horizontally-polarized light %.3f" % inten)
light = jones.field_vertical()
inten = jones.intensity(light)
print("Intensity for vertically-polarized light %.3f" % inten)
light = jones.field_linear(np.radians(45))
inten = jones.intensity(light)
print("Intensity for 45° linearly polarized light %.3f" % inten)
light = jones.field_right_circular()
inten = jones.intensity(light)
print("Intensity for right circularly polarized light %.3f" % inten)
light = jones.field_left_circular()
inten = jones.intensity(light)
print("Intensity for left circularly polarized light %.3f" % inten)
```
## The polarization variable $\chi$
The polarization variable is defined as
$$
\chi= \frac{E_y}{E_x}
$$
which happens to be equal to
$$
\chi = \frac{\tan\alpha+ j\tan\varepsilon}{1-j\tan\alpha\tan\varepsilon}
$$
where $\alpha$ is the azimuth of the ellipse and $\varepsilon=\tan{b/a}$ is the ellipticity angle ($b$ and $a$ are the minor and major axes of the ellipse.
```
azimuth = np.radians(-15)
phi_x = np.radians(17)
E0 = 0.5
for p in [-44, -30, 0, 15, 40]:
ellipticity_angle = np.radians(p)
tane = np.tan(ellipticity_angle)
tana = np.tan(azimuth)
J = jones.field_elliptical(azimuth, ellipticity_angle, phi_x, E0)
chi1 = (tana+1j*tane)/(1-1j*tana*tane)
print('jones = ', jones.polarization_variable(J))
print('expected = ', chi1)
print()
```
| github_jupyter |
# Rocket Problem with AeroSandbox Dynamics Engine
## Overview of the Dynamics Engine
### Overview & Motivation
In aerospace problems, we often end up with system dynamics that look (relatively) similar: our objects of interest are (roughly) rigid bodies moving around in 3D space (which we sometimes approximate as 2D or 1D).
While there's nothing wrong with implementing the dynamics manually for a problem [as we did in the previous tutorial](./01%20-%20Rocket%20Problem%20with%20Manual%20Dynamics.ipynb), it's a bit tedious and error-prone to manually implement the same dynamics every time we want to simulate an aerospace system.
So, AeroSandbox provides some (optional) shorthands that can be used to save hundreds of lines of code.
### Definitions, Coordinate Systems, and Assumptions
AeroSandbox's 3D dynamics engine essentially implements the full (nonlinear) equations of motion from Chapter 9 of [*"Flight Vehicle Aerodynamics"* by Mark Drela](https://mitpress.mit.edu/books/flight-vehicle-aerodynamics), along with the same (standard) coordinate system assumptions.
First, we define relevant axes systems:
* Earth Axes: Using the North, East, Down (NED) convention. Note that this implies some things:
* $z_e$ **points down!** If you want altitude, you want $-z_e$.
* Assumes negligible local curvature of the earth. Don't use this for your hypersonic vehicles without corrections.
* Body Axes: An axis system that is "painted on" (i.e., rotates with) the vehicle. Uses the convention where:
* $x_b$ points forward on the vehicle.
* $y_b$ points rightward on the vehicle.
* $z_b$ points down on the vehicle.
In other words, body axes are equivalent to geometry axes rotated 180° about the $y_g$ axis.
Specifically, we parameterize the state of a rigid 3D body in space with the following 12 state variables:
* $x_e$: $x$-position, in Earth axes. [meters]
* $y_e$: $y$-position, in Earth axes. [meters]
* $z_e$: $z$-position, in Earth axes. [meters]
* $u$: $x$-velocity, in body axes. [m/s]
* $v$: $y$-velocity, in body axes. [m/s]
* $w$: $z$-velocity, in body axes. [m/s]
* $\phi$: roll angle. Uses yaw-pitch-roll Euler angle convention. [rad]
* $\theta$: pitch angle. Uses yaw-pitch-roll Euler angle convention. [rad]
* $\psi$: yaw angle. Uses yaw-pitch-roll Euler angle convention. [rad]
* $p$: $x$-angular-velocity, in body axes. [rad/sec]
* $q$: $y$-angular-velocity, in body axes. [rad/sec]
* $r$: $z$-angular-velocity, in body axes. [rad/sec]
Force inputs to the system can be declared using the following inputs (in body axes):
* $X$: $x_b$-direction force. [N]
* $Y$: $y_b$-direction force. [N]
* $Z$: $z_b$-direction force. [N]
* $L$: Moment about the $x_b$-axis. Assumed these moments are applied about the center of mass. [Nm]
* $M$: Moment about the $y_b$-axis. Assumed these moments are applied about the center of mass. [Nm]
* $N$: Moment about the $z_b$-axis. Assumed these moments are applied about the center of mass. [Nm]
Mass properties are also defined:
* $m$: Mass of the body. [kg]
* $I_{xx}$: Respective component of the (symmetric) moment of inertia tensor.
* $I_{yy}$: Respective component of the (symmetric) moment of inertia tensor.
* $I_{zz}$: Respective component of the (symmetric) moment of inertia tensor.
* $I_{xy}$: Respective component of the (symmetric) moment of inertia tensor.
* $I_{xz}$: Respective component of the (symmetric) moment of inertia tensor.
* $I_{yz}$: Respective component of the (symmetric) moment of inertia tensor.
As are a few other quantities:
* $g$: Magnitude of gravitational acceleration. Assumed to act in the $z_e$ ("downward") direction, so a nominal value would be `9.81`. [m/s^2]
* $h_x$: $x_b$-component of onboard angular momentum (e.g. propellers), in body axes. [kg*m^2/sec]
* $h_y$: $y_b$-component of onboard angular momentum (e.g. propellers), in body axes. [kg*m^2/sec]
* $h_z$: $z_b$-component of onboard angular momentum (e.g. propellers), in body axes. [kg*m^2/sec]
## Example on Rocket Problem
Here, we pose the same rocket optimal control problem using the AeroSandbox dynamics engine.
```
import aerosandbox as asb
import aerosandbox.numpy as np
### Environment
opti = asb.Opti()
### Time discretization
N = 500 # Number of discretization points
time_final = 100 # seconds
time = np.linspace(0, time_final, N)
### Constants
mass_initial = 500e3 # Initial mass, 500 metric tons
z_e_final = -100e3 # Final altitude, 100 km
g = 9.81 # Gravity, m/s^2
alpha = 1 / (300 * g) # kg/(N*s), Inverse of specific impulse, basically - don't worry about this
dyn = asb.DynamicsPointMass1DVertical(
mass_props=asb.MassProperties(mass=opti.variable(init_guess=mass_initial, n_vars=N)),
z_e=opti.variable(init_guess=np.linspace(0, z_e_final, N)), # Altitude (negative due to Earth-axes convention)
w_e=opti.variable(init_guess=-z_e_final / time_final, n_vars=N), # Velocity
)
dyn.add_gravity_force(g=g)
thrust = opti.variable(init_guess=g * mass_initial, n_vars=N)
dyn.add_force(Fz=-thrust)
dyn.constrain_derivatives(
opti=opti,
time=time,
)
### Fuel burn
opti.constrain_derivative(
derivative=-alpha * thrust,
variable=dyn.mass_props.mass,
with_respect_to=time,
method="midpoint",
)
### Boundary conditions
opti.subject_to([
dyn.z_e[0] == 0,
dyn.w_e[0] == 0,
dyn.mass_props.mass[0] == mass_initial,
dyn.z_e[-1] == z_e_final,
])
### Path constraints
opti.subject_to([
dyn.mass_props.mass >= 0,
thrust >= 0
])
### Objective
opti.minimize(-dyn.mass_props.mass[-1]) # Maximize the final mass == minimize fuel expenditure
### Solve
sol = opti.solve(verbose=False)
print(f"Solved in {sol.stats()['iter_count']} iterations.")
dyn.substitute_solution(sol)
print(dyn)
```
Cool, so the problem solves!
This is an instance where the abstraction provided by the AeroSandbox dynamics engine isn't really needed. The dynamics here are so simple (they're 1-dimensional!) that we can just as easily implement our own integrators.
But where the dynamics engine really shines is when problems get more complicated - 2D and 3D problems with gyroscopic effects, interacting flight dynamics modes, etc. Let's take a look at an example in the following tutorial.
| github_jupyter |
```
!pip install av
! wget https://raw.githubusercontent.com/pytorch/vision/6de158c473b83cf43344a0651d7c01128c7850e6/references/video_classification/transforms.py
# Download HMDB51 data and splits from serre lab website
! wget http://serre-lab.clps.brown.edu/wp-content/uploads/2013/10/hmdb51_org.rar
! wget http://serre-lab.clps.brown.edu/wp-content/uploads/2013/10/test_train_splits.rar
pip install git+https://github.com/Atze00/MoViNet-pytorch.git
# Extract and organize video data..
! mkdir -p video_data test_train_splits
! unrar e test_train_splits.rar test_train_splits
! rm test_train_splits.rar
! unrar e hmdb51_org.rar
! rm hmdb51_org.rar
! mv *.rar video_data
import os
for files in os.listdir('video_data'):
foldername = files.split('.')[0]
os.system("mkdir -p video_data/" + foldername)
os.system("unrar e video_data/"+ files + " video_data/"+foldername)
! rm video_data/*.rar
import time
import torchvision
import torch.nn.functional as F
import torchvision.transforms as transforms
import torch.optim as optim
from torch.utils.data import random_split, DataLoader
import torch
import transforms as T
from movinets import MoViNet
from movinets.config import _C
torch.manual_seed(97)
num_frames = 16 # 16
clip_steps = 2
Bs_Train = 16
Bs_Test = 16
transform = transforms.Compose([
T.ToFloatTensorInZeroOne(),
T.Resize((200, 200)),
T.RandomHorizontalFlip(),
#T.Normalize(mean=[0.43216, 0.394666, 0.37645], std=[0.22803, 0.22145, 0.216989]),
T.RandomCrop((172, 172))])
transform_test = transforms.Compose([
T.ToFloatTensorInZeroOne(),
T.Resize((200, 200)),
#T.Normalize(mean=[0.43216, 0.394666, 0.37645], std=[0.22803, 0.22145, 0.216989]),
T.CenterCrop((172, 172))])
hmdb51_train = torchvision.datasets.HMDB51('video_data/', 'test_train_splits/', num_frames,frame_rate=5,
step_between_clips = clip_steps, fold=1, train=True,
transform=transform, num_workers=2)
hmdb51_test = torchvision.datasets.HMDB51('video_data/', 'test_train_splits/', num_frames,frame_rate=5,
step_between_clips = clip_steps, fold=1, train=False,
transform=transform_test, num_workers=2)
train_loader = DataLoader(hmdb51_train, batch_size=Bs_Train, shuffle=True)
test_loader = DataLoader(hmdb51_test, batch_size=Bs_Test, shuffle=False)
def train_iter(model, optimz, data_load, loss_val):
samples = len(data_load.dataset)
model.train()
model.cuda()
model.clean_activation_buffers()
optimz.zero_grad()
for i, (data,_ , target) in enumerate(data_load):
out = F.log_softmax(model(data.cuda()), dim=1)
loss = F.nll_loss(out, target.cuda())
loss.backward()
optimz.step()
optimz.zero_grad()
model.clean_activation_buffers()
if i % 50 == 0:
print('[' + '{:5}'.format(i * len(data)) + '/' + '{:5}'.format(samples) +
' (' + '{:3.0f}'.format(100 * i / len(data_load)) + '%)] Loss: ' +
'{:6.4f}'.format(loss.item()))
loss_val.append(loss.item())
def evaluate(model, data_load, loss_val):
model.eval()
samples = len(data_load.dataset)
csamp = 0
tloss = 0
model.clean_activation_buffers()
with torch.no_grad():
for data, _, target in data_load:
output = F.log_softmax(model(data.cuda()), dim=1)
loss = F.nll_loss(output, target.cuda(), reduction='sum')
_, pred = torch.max(output, dim=1)
tloss += loss.item()
csamp += pred.eq(target.cuda()).sum()
model.clean_activation_buffers()
aloss = tloss / samples
loss_val.append(aloss)
print('\nAverage test loss: ' + '{:.4f}'.format(aloss) +
' Accuracy:' + '{:5}'.format(csamp) + '/' +
'{:5}'.format(samples) + ' (' +
'{:4.2f}'.format(100.0 * csamp / samples) + '%)\n')
def train_iter_stream(model, optimz, data_load, loss_val, n_clips = 2, n_clip_frames=8):
"""
In causal mode with stream buffer a single video is fed to the network
using subclips of lenght n_clip_frames.
n_clips*n_clip_frames should be equal to the total number of frames presents
in the video.
n_clips : number of clips that are used
n_clip_frames : number of frame contained in each clip
"""
#clean the buffer of activations
samples = len(data_load.dataset)
model.cuda()
model.train()
model.clean_activation_buffers()
optimz.zero_grad()
for i, (data,_, target) in enumerate(data_load):
data = data.cuda()
target = target.cuda()
l_batch = 0
#backward pass for each clip
for j in range(n_clips):
output = F.log_softmax(model(data[:,:,(n_clip_frames)*(j):(n_clip_frames)*(j+1)]), dim=1)
loss = F.nll_loss(output, target)
_, pred = torch.max(output, dim=1)
loss = F.nll_loss(output, target)/n_clips
loss.backward()
l_batch += loss.item()*n_clips
optimz.step()
optimz.zero_grad()
#clean the buffer of activations
model.clean_activation_buffers()
if i % 50 == 0:
print('[' + '{:5}'.format(i * len(data)) + '/' + '{:5}'.format(samples) +
' (' + '{:3.0f}'.format(100 * i / len(data_load)) + '%)] Loss: ' +
'{:6.4f}'.format(l_batch))
loss_val.append(l_batch)
def evaluate_stream(model, data_load, loss_val, n_clips = 2, n_clip_frames=8):
model.eval()
model.cuda()
samples = len(data_load.dataset)
csamp = 0
tloss = 0
with torch.no_grad():
for data, _, target in data_load:
data = data.cuda()
target = target.cuda()
model.clean_activation_buffers()
for j in range(n_clips):
output = F.log_softmax(model(data[:,:,(n_clip_frames)*(j):(n_clip_frames)*(j+1)]), dim=1)
loss = F.nll_loss(output, target)
_, pred = torch.max(output, dim=1)
tloss += loss.item()
csamp += pred.eq(target).sum()
aloss = tloss / len(data_load)
loss_val.append(aloss)
print('\nAverage test loss: ' + '{:.4f}'.format(aloss) +
' Accuracy:' + '{:5}'.format(csamp) + '/' +
'{:5}'.format(samples) + ' (' +
'{:4.2f}'.format(100.0 * csamp / samples) + '%)\n')
N_EPOCHS = 1
model = MoViNet(_C.MODEL.MoViNetA0, causal = True, pretrained = True )
start_time = time.time()
trloss_val, tsloss_val = [], []
model.classifier[3] = torch.nn.Conv3d(2048, 51, (1,1,1))
optimz = optim.Adam(model.parameters(), lr=0.00005)
for epoch in range(1, N_EPOCHS + 1):
print('Epoch:', epoch)
train_iter_stream(model, optimz, train_loader, trloss_val)
evaluate_stream(model, test_loader, tsloss_val)
print('Execution time:', '{:5.2f}'.format(time.time() - start_time), 'seconds')
N_EPOCHS = 1
model = MoViNet(_C.MODEL.MoViNetA0, causal = False, pretrained = True )
start_time = time.time()
trloss_val, tsloss_val = [], []
model.classifier[3] = torch.nn.Conv3d(2048, 51, (1,1,1))
optimz = optim.Adam(model.parameters(), lr=0.00005)
for epoch in range(1, N_EPOCHS + 1):
print('Epoch:', epoch)
train_iter(model, optimz, train_loader, trloss_val)
evaluate(model, test_loader, tsloss_val)
print('Execution time:', '{:5.2f}'.format(time.time() - start_time), 'seconds')
```
| github_jupyter |
# Convolutional Neural Network Example
Build a convolutional neural network with TensorFlow v2.
This example is using a low-level approach to better understand all mechanics behind building convolutional neural networks and the training process.
- Author: Aymeric Damien
- Project: https://github.com/aymericdamien/TensorFlow-Examples/
## CNN Overview
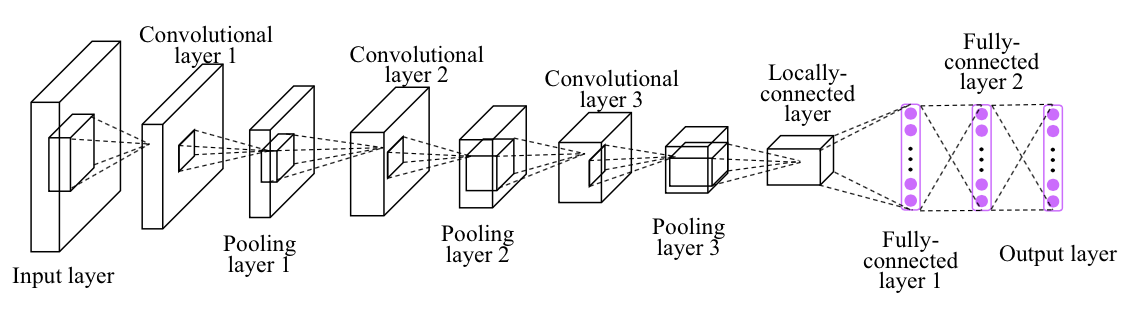
## MNIST Dataset Overview
This example is using MNIST handwritten digits. The dataset contains 60,000 examples for training and 10,000 examples for testing. The digits have been size-normalized and centered in a fixed-size image (28x28 pixels) with values from 0 to 255.
In this example, each image will be converted to float32 and normalized to [0, 1].

More info: http://yann.lecun.com/exdb/mnist/
```
from __future__ import absolute_import, division, print_function
import tensorflow as tf
from tensorflow.keras import Model, layers
import numpy as np
# MNIST dataset parameters.
num_classes = 10 # total classes (0-9 digits).
# Training parameters.
learning_rate = 0.001
training_steps = 200
batch_size = 128
display_step = 10
# Network parameters.
conv1_filters = 32 # number of filters for 1st conv layer.
conv2_filters = 64 # number of filters for 2nd conv layer.
fc1_units = 1024 # number of neurons for 1st fully-connected layer.
# Prepare MNIST data.
from tensorflow.keras.datasets import mnist
(x_train, y_train), (x_test, y_test) = mnist.load_data()
# Convert to float32.
x_train, x_test = np.array(x_train, np.float32), np.array(x_test, np.float32)
# Normalize images value from [0, 255] to [0, 1].
x_train, x_test = x_train / 255., x_test / 255.
# Use tf.data API to shuffle and batch data.
train_data = tf.data.Dataset.from_tensor_slices((x_train, y_train))
train_data = train_data.repeat().shuffle(5000).batch(batch_size).prefetch(1)
# Create TF Model.
class ConvNet(Model):
# Set layers.
def __init__(self):
super(ConvNet, self).__init__()
# Convolution Layer with 32 filters and a kernel size of 5.
self.conv1 = layers.Conv2D(32, kernel_size=5, activation=tf.nn.relu)
# Max Pooling (down-sampling) with kernel size of 2 and strides of 2.
self.maxpool1 = layers.MaxPool2D(2, strides=2)
# Convolution Layer with 64 filters and a kernel size of 3.
self.conv2 = layers.Conv2D(64, kernel_size=3, activation=tf.nn.relu)
# Max Pooling (down-sampling) with kernel size of 2 and strides of 2.
self.maxpool2 = layers.MaxPool2D(2, strides=2)
# Flatten the data to a 1-D vector for the fully connected layer.
self.flatten = layers.Flatten()
# Fully connected layer.
self.fc1 = layers.Dense(1024)
# Apply Dropout (if is_training is False, dropout is not applied).
self.dropout = layers.Dropout(rate=0.5)
# Output layer, class prediction.
self.out = layers.Dense(num_classes)
# Set forward pass.
def call(self, x, is_training=False):
x = tf.reshape(x, [-1, 28, 28, 1])
x = self.conv1(x)
x = self.maxpool1(x)
x = self.conv2(x)
x = self.maxpool2(x)
x = self.flatten(x)
x = self.fc1(x)
x = self.dropout(x, training=is_training)
x = self.out(x)
if not is_training:
# tf cross entropy expect logits without softmax, so only
# apply softmax when not training.
x = tf.nn.softmax(x)
return x
# Build neural network model.
conv_net = ConvNet()
# Cross-Entropy Loss.
# Note that this will apply 'softmax' to the logits.
def cross_entropy_loss(x, y):
# Convert labels to int 64 for tf cross-entropy function.
y = tf.cast(y, tf.int64)
# Apply softmax to logits and compute cross-entropy.
loss = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=y, logits=x)
# Average loss across the batch.
return tf.reduce_mean(loss)
# Accuracy metric.
def accuracy(y_pred, y_true):
# Predicted class is the index of highest score in prediction vector (i.e. argmax).
correct_prediction = tf.equal(tf.argmax(y_pred, 1), tf.cast(y_true, tf.int64))
return tf.reduce_mean(tf.cast(correct_prediction, tf.float32), axis=-1)
# Stochastic gradient descent optimizer.
optimizer = tf.optimizers.Adam(learning_rate)
# Optimization process.
def run_optimization(x, y):
# Wrap computation inside a GradientTape for automatic differentiation.
with tf.GradientTape() as g:
# Forward pass.
pred = conv_net(x, is_training=True)
# Compute loss.
loss = cross_entropy_loss(pred, y)
# Variables to update, i.e. trainable variables.
trainable_variables = conv_net.trainable_variables
# Compute gradients.
gradients = g.gradient(loss, trainable_variables)
# Update W and b following gradients.
optimizer.apply_gradients(zip(gradients, trainable_variables))
# Run training for the given number of steps.
for step, (batch_x, batch_y) in enumerate(train_data.take(training_steps), 1):
# Run the optimization to update W and b values.
run_optimization(batch_x, batch_y)
if step % display_step == 0:
pred = conv_net(batch_x)
loss = cross_entropy_loss(pred, batch_y)
acc = accuracy(pred, batch_y)
print("step: %i, loss: %f, accuracy: %f" % (step, loss, acc))
# Test model on validation set.
pred = conv_net(x_test)
print("Test Accuracy: %f" % accuracy(pred, y_test))
# Visualize predictions.
import matplotlib.pyplot as plt
# Predict 5 images from validation set.
n_images = 5
test_images = x_test[:n_images]
predictions = conv_net(test_images)
# Display image and model prediction.
for i in range(n_images):
plt.imshow(np.reshape(test_images[i], [28, 28]), cmap='gray')
plt.show()
print("Model prediction: %i" % np.argmax(predictions.numpy()[i]))
```
| github_jupyter |
```
pwd
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import geopandas as gpd
import geoplot
# read data
### flow
flow_df = pd.read_csv("../data/flows_sa2_months_2018-02-01.csv")
print("flow shape:", flow_df.shape)
print(flow_df.head(10))
print()
### transaction 1
trans_age_df = pd.read_csv("../data/transaction_age_bins.csv")
print("trans_age_df: ", trans_age_df.shape)
print(trans_age_df.head(10))
print()
### transaction 2
trans_mcc_df = pd.read_csv("../data/transaction_mcc.csv")
print("trans_mcc_df: ", trans_mcc_df.shape)
print(trans_mcc_df.head(10))
print()
# Change sa2 values to string
flow_df['sa2'] = flow_df.sa2.astype(str)
flow_df['agent_home_sa2'] = [x[:-2] for x in flow_df['agent_home_sa2'].astype(str)]
trans_age_df['source_sa2'] = trans_age_df['source_sa2'].astype(str)
trans_age_df['target_sa2'] = trans_age_df['target_sa2'].astype(str)
trans_mcc_df['source_sa2'] = trans_mcc_df['source_sa2'].astype(str)
trans_mcc_df['target_sa2'] = trans_mcc_df['target_sa2'].astype(str)
# Shapes
print("flow shape:", flow_df.shape)
print("trans_age_df: ", trans_age_df.shape)
print("trans_mcc_df: ", trans_mcc_df.shape)
# describe
print(flow_df.describe())
print(flow_df.dtypes)
print(trans_age_df.describe())
print(trans_age_df.dtypes)
print(trans_mcc_df.describe())
print(trans_mcc_df.dtypes)
# check a few columns' info
# Note: total number of SA2 in South Australia is 172.
# Many residents' locations from the credit card data are outside South Australia region.
print("--- number of sa2 in flow_df ---")
print("Flow SA2", len(np.unique(flow_df.sa2, return_counts = True)[0])) #
print(np.unique(flow_df.sa2, return_counts = True)) #
print()
print("Agent home SA2", len(np.unique(flow_df.agent_home_sa2, return_counts = True)[0])) #
print(np.unique(flow_df.agent_home_sa2, return_counts = True)) #
print()
print("--- sa2 in trans_age_df ---")
print("Source sa2", len(np.unique(trans_age_df.source_sa2, return_counts = True)[0])) #
print(np.unique(trans_age_df.source_sa2, return_counts = True)) #
print()
print("Target sa2", len(np.unique(trans_age_df.target_sa2, return_counts = True)[0])) #
print(np.unique(trans_age_df.target_sa2, return_counts = True)) #
print()
print("--- sa2 in trans_mcc_df ---")
print("Source sa2", len(np.unique(trans_mcc_df.source_sa2, return_counts = True)[0])) #
print(np.unique(trans_mcc_df.source_sa2, return_counts = True)) #
print()
print("Target sa2", len(np.unique(trans_mcc_df.target_sa2, return_counts = True)[0])) #
print(np.unique(trans_mcc_df.target_sa2, return_counts = True)) #
print()
print("--- activity types in trans_mcc_df ---")
print(np.unique(trans_mcc_df.mcc, return_counts = True)) # it is activity types.
print()
# Q: what are the 'Cell Size Limit', 'OUTST', and 'nan'? Can I just drop the values?
```
# Process Three Dataframes
```
# flow_df
# Remove the agents whose home are outside South Australia
south_australia_sa4_set = ['401','402','403','404','405','406','407']
flow_df = flow_df.loc[np.array([x[:3] in south_australia_sa4_set for x in flow_df.agent_home_sa2])]
flow_df
# edit trans_age_df
# remove the invalide values in source_sa2 and target_sa2
invalid_value_list = ['Cell Size Limit', 'nan', 'OUTST']
trans_age_df=trans_age_df.loc[~trans_age_df.source_sa2.isin(invalid_value_list)]
trans_age_df=trans_age_df.loc[~trans_age_df.target_sa2.isin(invalid_value_list)]
trans_age_df
# edit trans_mcc_df
# remove the invalide values in source_sa2 and target_sa2
invalid_value_list = ['Cell Size Limit', 'nan', 'OUTST']
trans_mcc_df=trans_mcc_df.loc[~trans_mcc_df.source_sa2.isin(invalid_value_list)]
trans_mcc_df=trans_mcc_df.loc[~trans_mcc_df.target_sa2.isin(invalid_value_list)]
trans_mcc_df
```
#### save cleaned files
```
trans_mcc_df.to_pickle("../data_process/trans_mcc_df.pkl")
trans_age_df.to_pickle("../data_process/trans_age_df.pkl")
flow_df.to_pickle("../data_process/flow_df.pkl")
```
# Spatial visualization
```
# Read SA2
sa2_shape = gpd.read_file("../data/sa2/SA2_2016_AUST.shp")
sa2_shape.dtypes
# Basic info of sa2 for the whole Australia
# sa2 shapefile is quite self-explainary.
# It includes the names and ids of sa2, sa3, and sa4 areas.
print(type(sa2_shape))
print(sa2_shape.head(10))
print(sa2_shape.dtypes)
# Keep only South Australia area.
# Info: file:///Users/shenhaowang/Downloads/StatePublicHealthPlan_Final.pdf
# Note: Based on the info above, South Australia has 7 SA4 regions.
south_australia_sa4_set = ['401','402','403','404','405','406','407']
sa2_south_au = sa2_shape.loc[sa2_shape.SA4_CODE16.isin(south_australia_sa4_set)]
print(sa2_south_au.shape) # 172 SA2 regions are left.
print(sa2_south_au.head(10))
# SA area
geoplot.polyplot(sa2_south_au,
edgecolor = 'black',
linewidth = 0.5,
figsize = (15,15))
# group vars
#
flow_source_sa2_group = flow_df.groupby(by=["agent_home_sa2"]).sum()
flow_target_sa2_group = flow_df.groupby(by=["sa2"]).sum()
flow_source_sa2_group['o_sa2'] = flow_source_sa2_group.index
flow_target_sa2_group['d_sa2'] = flow_target_sa2_group.index
flow_source_sa2_group.columns = ['state', 'unique_agents_home_based',
'sum_stay_duration_home_based',
'total_stays_home_based', 'o_sa2']
flow_target_sa2_group.columns = ['state', 'unique_agents_d_based',
'sum_stay_duration_d_based',
'total_stays_d_based', 'd_sa2']
print(flow_source_sa2_group.head(5))
print(flow_target_sa2_group.head(5))
#
trans_age_source_sa2_group = trans_age_df.groupby(by = ['source_sa2']).sum()
trans_age_target_sa2_group = trans_age_df.groupby(by = ['target_sa2']).sum()
trans_age_source_sa2_group['o_sa2'] = trans_age_source_sa2_group.index
trans_age_target_sa2_group['d_sa2'] = trans_age_target_sa2_group.index
trans_age_source_sa2_group.columns = ['count_credit_o_from_age_file', 'amount_credit_o_from_age_file', 'o_sa2']
trans_age_target_sa2_group.columns = ['count_credit_d_from_age_file', 'amount_credit_d_from_age_file', 'd_sa2']
print(trans_age_source_sa2_group.head(5))
print(trans_age_target_sa2_group.head(5))
#
trans_mcc_source_sa2_group = trans_mcc_df.groupby(by = ['source_sa2']).sum()
trans_mcc_target_sa2_group = trans_mcc_df.groupby(by = ['target_sa2']).sum()
trans_mcc_source_sa2_group['o_sa2'] = trans_mcc_source_sa2_group.index
trans_mcc_target_sa2_group['d_sa2'] = trans_mcc_target_sa2_group.index
trans_mcc_source_sa2_group.columns = ['count_credit_o_from_mcc_file', 'amount_credit_o_from_mcc_file', 'o_sa2']
trans_mcc_target_sa2_group.columns = ['count_credit_o_from_mcc_file', 'amount_credit_o_from_mcc_file', 'd_sa2']
print(trans_mcc_source_sa2_group.head(5))
print(trans_mcc_target_sa2_group.head(5))
# merge
#
sa2_south_au_info = sa2_south_au.merge(flow_source_sa2_group, left_on = 'SA2_MAIN16', right_on = 'o_sa2', how = 'outer')
sa2_south_au_info = sa2_south_au_info.merge(flow_target_sa2_group, left_on = 'SA2_MAIN16', right_on = 'd_sa2', how = 'outer')
#
sa2_south_au_info=sa2_south_au_info.merge(trans_age_source_sa2_group, left_on = 'SA2_MAIN16', right_on = 'o_sa2', how = 'outer')
sa2_south_au_info=sa2_south_au_info.merge(trans_age_target_sa2_group, left_on = 'SA2_MAIN16', right_on = 'd_sa2', how = 'outer')
sa2_south_au_info=sa2_south_au_info.merge(trans_mcc_source_sa2_group, left_on = 'SA2_MAIN16', right_on = 'o_sa2', how = 'outer')
sa2_south_au_info=sa2_south_au_info.merge(trans_mcc_target_sa2_group, left_on = 'SA2_MAIN16', right_on = 'd_sa2', how = 'outer')
sa2_south_au_info
print(sa2_south_au_info.columns)
# visualize counts of credit card usage for destination and origin locations from the age file.
print("Visualize counts of credit card usage for destination and origin locations")
ax = geoplot.polyplot(sa2_south_au,
edgecolor = 'black',
linewidth = 0.02,
figsize = (8,8))
ax.set_title("Counts of credit card usage in destination locations", fontsize=16)
geoplot.choropleth(sa2_south_au_info,
hue = sa2_south_au_info['count_credit_d_from_age_file'],
edgecolor = 'black',
linewidth = 0.02,
ax = ax,
cmap='Greens')
#
ax = geoplot.polyplot(sa2_south_au,
edgecolor = 'black',
linewidth = 0.02,
figsize = (8,8))
ax.set_title("Counts of credit card usage in origin locations", fontsize=16)
geoplot.choropleth(sa2_south_au_info,
hue = sa2_south_au_info['count_credit_o_from_age_file'],
edgecolor = 'black',
linewidth = 0.02,
ax = ax,
cmap='Greens')
import folium
import networkx
```
## Build network
## Baseline gravity model
Key question: what are the inputs and outputs?
| github_jupyter |
# 基于 GCN 的有监督学习
图神经网络(GNN)结合了图结构和机器学习的优势. GraphScope提供了处理学习任务的功能。本次教程,我们将会展示GraphScope如何使用GCN算法训练一个模型。
本次教程的学习任务是在文献引用网络上的点分类任务。在点分类任务中,算法会确定[Cora](https://linqs.soe.ucsc.edu/data)数据集上每个顶点的标签。在```Cora```数据集中,由学术出版物作为顶点,出版物之间的引用作为边,如果出版物A引用了出版物B,则图中会存在一条从A到B的边。Cora数据集中的节点被分为了七个主题类,我们的模型将会训练来预测出版物顶点的主题。
在这一任务中,我们使用图聚合网络(GCN)算法来训练模型。有关这一算法的更多信息可以参考["Knowing Your Neighbours: Machine Learning on Graphs"](https://medium.com/stellargraph/knowing-your-neighbours-machine-learning-on-graphs-9b7c3d0d5896)
这一教程将会分为以下几个步骤:
- 启动GraphScope的学习引擎,并将图关联到引擎上
- 使用内置的GCN模型定义训练过程,并定义相关的超参
- 开始训练
```
# Install graphscope package if you are NOT in the Playground
!pip3 install graphscope
!pip3 uninstall -y importlib_metadata # Address an module conflict issue on colab.google. Remove this line if you are not on colab.
# Import the graphscope module.
import graphscope
graphscope.set_option(show_log=False) # enable logging
# Load cora dataset
from graphscope.dataset import load_cora
graph = load_cora()
```
然后,我们需要定义一个特征列表用于图的训练。训练特征集合必须从点的属性集合中选取。在这个例子中,我们选择了属性集合中所有以"feat_"为前缀的属性作为训练特征集,这一特征集也是Cora数据中点的特征集。
借助定义的特征列表,接下来,我们使用 [graphlearn](https://graphscope.io/docs/reference/session.html#graphscope.Session.graphlearn) 方法来开启一个学习引擎。
在这个例子中,我们在 "graphlearn" 方法中,指定在数据中 "paper" 类型的顶点和 "cites" 类型边上进行模型训练。
利用 "gen_labels" 参数,我们将 "paper" 点数据集进行划分,其中75%作为训练集,10%作为验证集,15%作为测试集。
```
# define the features for learning
paper_features = []
for i in range(1433):
paper_features.append("feat_" + str(i))
# launch a learning engine.
lg = graphscope.graphlearn(
graph,
nodes=[("paper", paper_features)],
edges=[("paper", "cites", "paper")],
gen_labels=[
("train", "paper", 100, (0, 75)),
("val", "paper", 100, (75, 85)),
("test", "paper", 100, (85, 100)),
],
)
```
这里我们使用内置的GCN模型定义训练过程。你可以在[Graph Learning Model](https://graphscope.io/docs/learning_engine.html#data-model)获取更多内置学习模型的信息。
在本次示例中,我们使用tensorflow作为NN后端训练器。
```
import graphscope.learning
from graphscope.learning.examples import GCN
from graphscope.learning.graphlearn.python.model.tf.optimizer import get_tf_optimizer
from graphscope.learning.graphlearn.python.model.tf.trainer import LocalTFTrainer
# supervised GCN.
def train(config, graph):
def model_fn():
return GCN(
graph,
config["class_num"],
config["features_num"],
config["batch_size"],
val_batch_size=config["val_batch_size"],
test_batch_size=config["test_batch_size"],
categorical_attrs_desc=config["categorical_attrs_desc"],
hidden_dim=config["hidden_dim"],
in_drop_rate=config["in_drop_rate"],
neighs_num=config["neighs_num"],
hops_num=config["hops_num"],
node_type=config["node_type"],
edge_type=config["edge_type"],
full_graph_mode=config["full_graph_mode"],
)
graphscope.learning.reset_default_tf_graph()
trainer = LocalTFTrainer(
model_fn,
epoch=config["epoch"],
optimizer=get_tf_optimizer(
config["learning_algo"], config["learning_rate"], config["weight_decay"]
),
)
trainer.train_and_evaluate()
# define hyperparameters
config = {
"class_num": 7, # output dimension
"features_num": 1433,
"batch_size": 140,
"val_batch_size": 300,
"test_batch_size": 1000,
"categorical_attrs_desc": "",
"hidden_dim": 128,
"in_drop_rate": 0.5,
"hops_num": 2,
"neighs_num": [5, 5],
"full_graph_mode": False,
"agg_type": "gcn", # mean, sum
"learning_algo": "adam",
"learning_rate": 0.01,
"weight_decay": 0.0005,
"epoch": 5,
"node_type": "paper",
"edge_type": "cites",
}
```
在定义完训练过程和超参后,现在我们可以使用学习引擎和定义的超参开始训练过程。
```
train(config, lg)
```
| github_jupyter |
<table align="left">
<td>
<a href="https://colab.research.google.com/github/nyandwi/machine_learning_complete/blob/main/5_intro_to_machine_learning/5_intro_to_machine_learning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Read In Colab"/></a>
</td>
</table>
*This notebook was created by [Jean de Dieu Nyandwi](https://twitter.com/jeande_d) for the love of machine learning community. For any feedback, errors or suggestion, he can be reached on email (johnjw7084 at gmail dot com), [Twitter](https://twitter.com/jeande_d), or [LinkedIn](https://linkedin.com/in/nyandwi).*
<a name='0'></a>
## Intro to Machine Learning
*Covering:*
* [1. Intro to Machine Learning Paradigm](#1)
* [2. Machine Learning Workflow](#2)
* [3. Evaluation Metrics](#3)
* [4. Underfitting (Low Bias) and Overfitting (High Variance)](#4)
<a name='1'></a>
## 1. Intro to Machine Learning Paradigm
Machine Learning is a new programming paradigm in which instead of explicitly programming computers to perform tasks, we let them learn from data in order to find the underlying patterns in the data.
Here is an excerpt of ML definition from Wikipedia:
***Machine learning (ML) is the study of computer algorithms that improve automatically through experience and by the use of data. It is seen as a part of artificial intelligence. Machine learning algorithms build a model based on sample data, known as "training data", in order to make predictions or decisions without being explicitly programmed to do so. - Wikipedia.***
What does that mean?
Simple, we do not program machines. We show them the data and they figure out the rest. Take an example: if you wanted to build a program that can recognize human and horse with traditional programming, you would have to write every single instruction which differentiate horse from human, or set of codes that represent human/horse. But with Machine Learning, you can feed the image of the horse/humans in different scenarios, and the machine can figure out rules which can help us identify horse and human.
This brings us to the next topic, Machine Learning Vs Traditional programming.
### Machine Learning Vs Traditional programming
The key inputs to any typical machine learning model are data and the results (or labels). The job of the model is to determine the set of rules that map the data and labels. Let's think about it using our previous example: if you provide a bunch of pictures of humans and horses to the ML system along with their labels(their names), you can get the rules which can help you perform the task of classifying horse and human.
On the flip side, in standard programming, this is what we do and if you are a coder you know it well. In order to get results, you have to provide data and rules.

### Types of Machine Learning
In broad, there are 3 types of Machine learning:
* **Supervised learning** where you have the data and the labels. Think of labels as what represents the input data.
* **Unsupervised learning**: Where you do not have the labels
* **Reinforcement learning** in which the goal is to optimize the rewards. Reinforcement learning has got its application in areas such as robotics.
Most ML problems fall in the category of supervised learning. Let's take an example. In our example earlier, if horse images are labelled as horses and same for humans, it's easy for a machine to relate each image with its associated label.
An example of unsupervised learning is a customer segmentation. Let's say that you want to provide promotions to a group of your clients based on their purchasing history, but you don't know these groups and their interests well. You just only have data. Using unsupervised techniques such as clustering, you can group customers who share the same interests and will likely all appreciate the promotion that you're offering. That's just one example, there are more applications of unsupervised learning.
### Categories of ML Problems
In terms of problems that we can solve with machine learning, there are 3 categories which are closely connected to the types we saw in the last section.
* **Classification**: This falls in the supervised learning type. As we saw, the example can be to classify a horse or human. Whether you have two categories or more, they are all classification problems. With two categories, it is usually termed as binary classification, and multi-classification for more than two categories.
* **Regression**: This is also a supervised type. The goal here is to predict a continuous value. Example is predicting the price of a house given its size, region, number of rooms, etc...
* **Clustering** where the goal is to group some entities based on given characteristics. An example can be to group the customers based on some similar characteristics.
### ML Applications
Nowadays, most tech products/services possess some sort of machine learning algorithm running in the background, from browsers to our mobile phone. It's fun, my mobile is able to recognize me in any photo that I am part of. Or things like character recognition where your phone camera can read characters.
It's fair to say that Machine Learning has transformed many industries, from banking, healthcare, production, streaming, to autonomous vehicles. Here are other detailed scenarios highlighting the applications of machine learning in the real world settings:
* A bank or any credit card provider can detect fraud in real-time. Banks can also predict if a given customer requesting a loan will pay it back or not based on their financial history.
* A Medical Analyst can diagnose a disease in a handful of minutes, or predict the likelihood or course of diseases or survival rate(Prognosis).
* An engineer in a given industry can detect failure or defect on the equipment.
* A telecommunication company can learn that a given customer is not satisfied with the service and is likely to opt-out from the service (churn).
* Our email inboxes are smart enough to differentiate spams and important emails.
* A given ads agency can place the relevant ads on a website that are likely to attract visitors.
* A driverless car can confidently know that an object in front is a pedestrian.
* A streaming service can suggest the best media to their clients based on their interests.
<a name='2'></a>
## 2. Machine Learning Workflow
Different from standard programming, ML is made of code and data. Some steps in ML involve data while a small part involve code(ML algorithm).
Here is an end to end ML workflow:
* Defining the problem
* Collecting the data
* Establishing the baseline
* Exploratory Data Analysis
* Feature Engineering
* Choosing/Creating/Training a ML model
* Performing Error analysis
* Deploying a model
Let's elaborate it more, step by step.
### 1. Problem definition
Problem definition is the important and the initial step in any ML project. This is where you make sure you understand the problem really well. Understanding the problem will give you proper intuitions about the next steps to follow such as right learning algorithms, etc.
### 2. Collecting the data
After you have defined your problem, the next step is to find the relevant data.
Nowadays, there are many open source datasets, be it images, texts, or structured data (data in tabular fashion) on platforms like Kaggle, Google Datasets, UCL, and some government websites. Your job as ML Engineer is to find the relevant data that you can use to solve a presented problem.
But there are times that you will have to collect your dataset, especially if you are solving a problem that no one solved before. In this case, consider the time that you will have to spend collecting data and the cost. You also do not need to wait until you have your desired data points before you can start. Embrace ML development early on so that you can learn if you (really) need more data or improvements you can make for further collections. This idea is inspired by Andrew Ng.
Also, when collecting the data, quality is better than quantity. There are times where small data but good data can outwork big poor data.
The amount of data you need is going to depend on the problem you're solving and its scope, but whatever the problem is, aiming to collect good data is the way to go.
### 3. Establishing a baseline
Without a benchmark, you won't know how to evaluate your results properly.
A baseline is the simplest model that can solve your project. It does not have to be a model. It can be an open source application, a statistical analysis or intuitions you get from data at a quick glance.
Take an example: Let's say that you want to build a cat and dog classifier and you have 2000 images, where 1400 are cats and 600 are dogs. Before you build a model there is already a 70% chance that any random image you pick up will be a cat. In this case, 70% is your simple baseline and it means your goal is to predict cats with 70% accuracy or more.
If you can't beat the baseline, sometimes it means the project is not worth pursuing.
### 4. Exploratory Data Analysis(EDA)
Before manipulating the data, it is quite important to go through it with the goal of learning the dataset. This can be overlooked, but doing it well will help you to know the effective strategies to be applied while cleaning the data.
Go through some values, plot some features, and try to understand the correlation between them. If the work is vision-related, visualize some images, spot what’s missing in the images. Are they diverse enough? What types of image scenarios can you add? Or what are images that can mislead the model?
Here are other things that you would want to inspect in the data:
* Duplicate values
* Corrupted or data with unsupported formats (ex: having an image of .txt and with 0 Kilobytes)
* Class imbalances
* Biases that can be present in the data
Before performing EDA, split the data into the training set, validation and test sets to avoid data leakage.
### 5. Data Preprocessing
Data processing is perhaps the biggest part of an ML project. There is a notion that Data Scientists and Machine Learning Engineers spend more than 80% preparing the data and this makes sense, the real world datasets are messy.
In this step, it is where you convert the raw data to go in a format that can be accepted by the ML algorithm. It can mean manipulating features to be in their proper formats or create new features from the existing ones.
Feature engineering being a part of data processing is also where things get creative. In terms of structured data, the way you engineer a numerical feature is going to be different to how you engineer the categorical features. Also in unstructured data, the way you manipulate images is going to be different to how you manipulate texts or sounds.
As the next parts will cover the practical implementations of this step in various data types (tabular, images, texts), let's be general about things you're likely going to deal with while manipulating the features:
* **Imputing missing values**: Missing values can either be filled, removed or left as they are. There are various strategies for missing values such as mean, median, frequent imputations, backward and forward fill, and iterative imputations. The right imputation technique depends on the problem and the dataset. While imputing missing values is appreciated for most machine learning algorithms, tree models can not suffer from them. So if you are using models like Random Forest or Decision tree, you can leave missing values as they are.
* **Encoding categorical features**: Categorical features are all types of features that have categorical values. For example, A gender feature having the values Male and Female is a categorical feature. You will want to encode such types of features. The techniques for encoding them are label encoding where you can assign 1 to Male, and 2 to Female, or one hot encoding where you can get the binary representations (0s and 1s) in one hot matrix. You will see this in practice.
* **Normalizing/Standardizing the numeric features**: Most ML models will work well when the input values are scaled to small values and that will result in training fast as well as converging fast. Normalization will set the values to between 0 and 1 whereas Standardization rescale the features to have mean of 0 and unit standard deviation. If you know that your data has normal or gausian distribution, normalization can be a good choice. Otherwise, Standardization will work well by default.
* **Dealing with date features**: The date in the format we know it may not be recognized by the model. If we have Day-MM--YY, we can create a day, month and column features.
Data preparation is a huge work and it depends on the dataset you are working with. The type of feature processing that you can apply is unique and will depend on the problem at hand and the available dataset.
For more about handling categorical features, check this [article](https://jeande.medium.com/your-onestop-guide-on-handling-categorical-features-5988caaef78a), feature scaling [here](https://jeande.medium.com/the-ultimate-and-practical-guide-on-feature-scaling-d03fbe2cb25e), and handling missing values [here](https://jeande.medium.com/a-comprehensive-guide-for-handling-missing-values-990c999c49ed).
### 6. Choosing/Creating a ML model
Choosing and Creating a model is the tiniest part in a typical machine learning workflow.
There are different types of models but most of them will fall in these categories: linear models such as linear/logistic regression, tree based models like decision trees, ensembles such as Gradient Boost or XGBoost, and neural networks.
All of these are what you can choose from when creating a model for your problem. Getting a model that can ultimately solve your problem is no free lunch scenario. You will have to experiment with different models and tune hyper-parameters, things like that.
To reduce your modelling curve, here are a few things that you can consider while choosing a machine learning model.
* **The scope of the problem**: There are a set of problems which directly give a clue of the right learning algorithm. Take an example: If you are going to build an image classifier from big data, neural networks (Convolutional neural networks specifically) might be a go to algorithm.
* **The size of the dataset**: Linear models tend to work well in small data problems whereas ensembles and neural networks can work well in complex problems.
* **The level of interpretability**: If you want the results of your model to be explainable, neural networks will not be a good choice. Tree based models such as decision trees can be explainable compared to other models.
* **Training time**: Complex models (neural nets and ensembles) will take too long to train and thus exhausting the computation resources. On the other hand, linear models can train faster.
So as you can see, there is a trade off. You want explainability, choose models which can provide that for you. You have a small dataset or you care about the training time, same thing, choose a reasonable algorithm.
It is also worth mentioning again that choosing and training a model is an iterative process. With the grouth of ML frameworks such as Scikit-Learn, TensorFlow or PyTorch, building is easy. But getting the model to converge is another thing. It is often data improvement aided by error analysis that will ultimately improve the model.
For more about model seletion, here is the [article](https://jeande.medium.com/solving-a-puzzle-machine-learning-algorithm-selection-2c4a1b5ccf35) I wrote a while back.
### 7. Performing Error Analysis
Performing the error analysis will guide you throughout the data and the model improvement. In order to spot the errors, we have to iteratively ask the right questions.
Because a good model comes from good data, we will want to keep the mode fixed to see the types of errors that are happening. Here are questions that we can ask ourselves:
* Is the model doing poorly on all classes or is it one specific class?
* Is it because there are not enough data points for that particular class compared to other classes?
* There are trade-offs and limits on how much you can do to reduce the error. Is there room for improvement?
Often, the improvement will not come from tuning the model, but spending time to increase the data quality.
When improving the data, you can create artificial data (a.ka data augmentation). This will work well most of the time. As this step is iterative too, keep doing error analysis and continuously aim to improve the data.
### 8. Deploying, Monitoring and Maintaining the Model
Model deployment is the last part in this workflow. When everything has gone right, your model converged or you came to find the reliable data, the next step will be to deploy the model so that the users can start to make requests and get predictions or enhanced services (ML in action).
Model deployment is not in the scope of this introduction. If you want to learn more about it, I recommend [Machine Learning Engineering for Production (MLOps) Specialization - Deeplearning.AI](https://www.coursera.org/specializations/machine-learning-engineering-for-production-mlops).
<a name='3'></a>
## 3. Evaluation Metrics
Earlier in this introduction to Machine Learning, we saw that most problems are either regression or classification. In this section, we will learn the evaluation metrics that are used in evaluating the performance of the machine learning models.
Let's kick this off with the regression metrics!
### Regression Metrics
In regression, the goal is to predict the continuous value. The difference between the actual value and the predicted value is called the error.
`Error = Actual value - Predicted value`
The square of the error over all samples is called mean squarred error.
`MSE = SQUARE(Actual value - Predicted value)/Number of Samples`
*MSE Actual Formula*:
$$\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2$$
Taking the square root of the mean squared error will give the Root Mean Squared Error(RMSE). RMSE is the most used regression metric.
*RMSE Actual Formula*:
$$\sqrt{\frac 1n\sum_{i=1}^n(y_i-\hat{y}_i)^2}$$
There are times that you will work with the datasets containing outliers. In this case, the commonly used metric is called Mean Absolute Error (MAE). As simple as calculating MSE, MAE is also the absolute of the error.
`MAE = ABSOLUTE (Actual value - Predicted Value)`
*MAE Actual Formula*
$$\frac 1n\sum_{i=1}^n|y_i-\hat{y}_i|$$
Like said, MAE is very sensitive to outliers. This will make it a suitable metric for all kinds of problems which are likely to have abnormal scenarios such as time series.
### Classification Metrics
In classification problems, the goal is to predict the categories/class. Accuracy is the most used metric. The accuracy shows the ability of the model in making the correct predictions. Take an example, in a horse/human classifier. If you have 250 training images for horses and the same number for humans, and the model can confidently predict 400 images, then the accuracy is 400/500 = 0.8, so your model is 80% accurate.
The accuracy is simply an indicator of how your model is in making correct predictions and it will only be useful if you have a balanced dataset (like we had 250 images for horses and 250 images for humans).
When we have a skewed dataset or when there are imbalances, we need a different perspective on how we evaluate the model. Take an example, if we have 450 images for horses and 50 images for humans, there is a chance of 90% (450/500) that the horse will be correctly predicted, because the dataset is dominated by the horses. But how about humans? Well, it's obvious that the model will struggle predicting them correctly.
This is where we introduce other metrics that can be far more useful than accuracy, such as precision, recall, and F1 score.
Precision shows the percentage of the positive predictions that are actually positive. To quote [Google ML Crash Course](https://developers.google.com/machine-learning/crash-course/classification/precision-and-recall), precision answer the following question: `What proportion of positive identifications was actually correct?`
The recall on the other hand shows the percentage of the actual positive samples that were classified correctly. It answers this question: `What proportion of actual positives was identified correctly?`
There is a tradeoff between precision and recall. Often, increasing precision will decrease recall and vice versa. To simplify things, we combine both of these two metrics into a single metric called the F1 score.
F1 score is the harmonic mean of precision and recall, and it shows how good the model is at classifying all classes without having to balance between precision and recall. If either precision or recall is very low, the F1 score is going to be low too.
Both accuracy, precision, and recall can be calculated easily by using a [confusion matrix](https://jeande.tech/how-to-read-a-confusion-matrix). A confusion matrix shows the number of correct and incorrect predictions made by a classifier in all available classes.
More intuitively, a confusion matrix is made of 4 main elements: True negatives, false negatives, true positives, and false positives.
* **True Positives(TP)**: Number of samples that are correctly classified as positive, and their actual label is positive.
* **False Positives (FP)**: Number of samples that are incorrectly classified as positive, when in fact their actual label is negative.
* **True Negatives (TN)**: Number of samples that are correctly classified as negative, and their actual label is negative.
* **False Negatives (FN)**: Number of samples that are incorrectly classified as negative, when in fact their actual label is positive.
The accuracy that we talked about is the number of correct examples over total examples. So, that is
`Accuracy = (TP + TN) / (TP + TN + FP + FN)`
Precision is the model accuracy on predicting positive examples.
`Precision = TP / (TP + FP)`
On the other hand, Recall is the model ability to predict the positive examples correctly.
`Recall = TP / (TP+FN)`
The higher the recall and precision, the better the model is at making accurate predictions but there is a tradeoff between them. Increasing precision will reduce the recall and vice versa.
A classifier that doesn't have false positives has a precision of 1, and a classifier that doesn't have false negatives has a recall of 1. Ideally, a perfect classifier will have the precision and recall of 1.
We can combine both precision and recall to get another metric called F1 Score. F1 Score is the harmonic mean of precision and recall.
`F1 Score = 2 *(precision * recall) / (precision + recall)`
Take an example of the following confusion matrix.
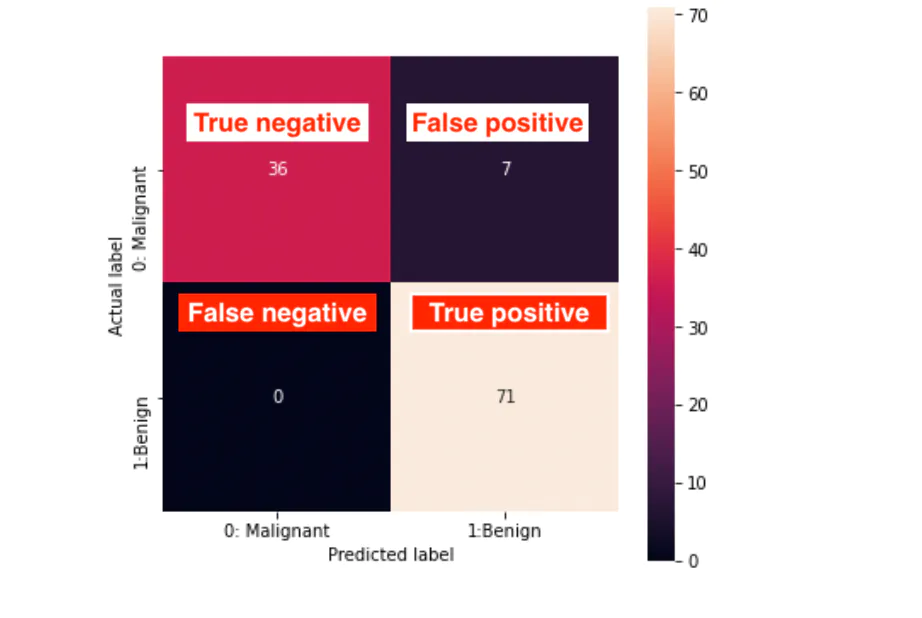
From the above confusion matrix:
* Accuracy = `(TP + TN) / (TP + TN + FP + FN) = (71 +36)/(71+36+7+0) = 0.93 or 93%`
* Precision = `TP / (TP + FP) = 71/(71+7) =0.91 or 91%`
* Recall = `TP / (TP + FN) = 71/(71+0) = 1, or 100%`
* F1 score = `2PR / (P + R) = 2x0.91x1/(0.91+1) = 0.95, or 95%`
Both accuracy, confusion matrix, precision, recall, and F1 score are implemented easily in Scikit-Learn, a Machine Learning framework used to build classical ML algorithms.
<a name='4'></a>
## 4. Underfitting and Overfitting
Building a machine learning model that can fit the training data well is not a trivial task. Often, at the initial training, the model will either underfit or overfit the data. Some machine learning models proves that really well. Take an example: When training a decision trees, it is very likely that they will overfit the data at first.
There is a trade off between underfitting/overfitting, and so it's important to understand the difference between them and how to handle each and each. Understanding and handling underfitting/overfitting is a critical task in diagonizing machine learning models.
### Underfitting (Low Bias)
Underfitting happens when the model does poor on the training data. It can be caused by the fact that the model is simple for the training data or the data does not contains the things that you are trying to predict. Good data has high `predictive power`, and poor data has low predictive power.
Here are some of the techniques that can be used to deal with a model which has low bias(underfit):
* Use complex models. If you are using linear models, try other complex models like Random forests or Support Vector Machines. Not to mention neural networks if you are dealing with unstructured data (images, texts, sounds)
* Add more training data and use good features. Good features have high predictive power.
* Reduce the regularization.
* If you're using neural networks, increase the number of epochs/training iterations. If the epochs are very low, the model may not be able to learn the underlying rules in data and so it will not perform well.
### Overfitting (High Variance)
Overfitting is the reverse of underfitting. An overfitted model will do well on the training data but will be so poor when supplied with a new data (the data that the model never saw).
Overfitting is caused by using model which is too complex for the dataset and few training examples.
Here are techniques to handle overfitting:
* Try simple models or simplify the current model. Some machine learning algorithms can be simplified. Take an example: in neural networks, you can reduce the number of layers or neurons. Also in classical algorithms like Support Vector Machines, you can try different kernels, a linear kernel is simple than a polynomial kernel.
* Find more training data.
* Stop the training early (a.k.a Early Stopping)
* Use other different regularization techniques like dropout(in neural networks).
To summarize this, it is very important to be able to understand why the model is not doing well. If the model is being poor on the data it was trained on, you know it is underfitting and you know what to do about it.
Also with the exception of improving/expanding the training data, often you have to tune hyperparameters to get a model that can generalize well. While there are techniques that simplified hyperparameter search(like [Grid search](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.GridSearchCV.html#sklearn.model_selection.GridSearchCV), [Random search](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html), [Keras Tuner]), it is important to understand the hyperparameters of the model you are using so that you can know their proper search space.
### [BACK TO TOP](#0)
| github_jupyter |
# Welcome to ExKaldi
In this section, we will train a N-Grams language model and query it.
Althrough __SriLM__ is avaliable in ExKaldi, we recommend __KenLM__ toolkit.
```
import exkaldi
import os
dataDir = "librispeech_dummy"
```
Firstly, prepare the lexicons. We have generated and saved a __LexiconBank__ object in file already (3_prepare_lexicons). So restorage it directly.
```
lexFile = os.path.join(dataDir, "exp", "lexicons.lex")
lexicons = exkaldi.load_lex(lexFile)
lexicons
```
We will use training text corpus to train LM model. Even though we have prepared a transcription file in the data directory, we do not need the utterance-ID information at the head of each line, so we must take a bit of work to produce a new text.
We can lend a hand of the exkaldi __Transcription__ class.
```
textFile = os.path.join(dataDir, "train", "text")
trans = exkaldi.load_transcription(textFile)
trans
newTextFile = os.path.join(dataDir, "exp", "train_lm_text")
trans.save(fileName=newTextFile, discardUttID=True)
```
But actually, you don't need do this. If you use a __Transcription__ object to train the language model, the information of utterance ID will be discarded automatically.
Now we train a 2-grams model with __KenLM__ backend.
```
arpaFile = os.path.join(dataDir, "exp", "2-gram.arpa")
exkaldi.lm.train_ngrams_kenlm(lexicons, order=2, text=trans, outFile=arpaFile, config={"-S":"20%"})
```
ARPA model can be transform to binary format in order to accelerate loading and reduce memory cost.
Although __KenLM__ Python API supports reading ARPA format, but in exkaldi, we only expected KenLM Binary format.
```
binaryLmFile = os.path.join(dataDir, "exp", "2-gram.binary")
exkaldi.lm.arpa_to_binary(arpaFile, binaryLmFile)
```
Use the binary LM file to initialize a Python KenLM n-grams object.
```
model = exkaldi.lm.KenNGrams(binaryLmFile)
model
```
__KenNGrams__ is simple wrapper of KenLM python Model. Check model information:
```
model.info
```
You can query this model with a sentence.
```
model.score_sentence("HELLO WORLD", bos=True, eos=True)
```
There is a example to compute the perplexity of test corpus in order to evaluate the language model.
```
evalTrans = exkaldi.load_transcription( os.path.join(dataDir, "test", "text") )
score = model.score(evalTrans)
score
type(score)
```
___score___ is an exkaldi __Metric__ (a subclass of Python dict) object.
We design a group of classes to hold Kaldi text format table and exkaldi own text format data:
__ListTable__: spk2utt, utt2spk, words, phones and so on.
__Transcription__: transcription corpus, n-best decoding result and so on.
__Metric__: AM score, LM score, LM perplexity, Sentence lengthes and so on.
__IndexTable__: The index of binary data.
__WavSegment__: The wave information.
All these classes are subclasses of Python dict. They have some common and respective methods and attributes.
In this case, for example, we can compute the average value of __Metric__.
```
score.mean()
```
More precisely, the weighted average by the length os sentences.
```
score.mean( weight= evalTrans.sentence_length() )
```
Actually, we use perplexity more to evaluate it.
```
model.perplexity(evalTrans)
```
Back to Language Model. If you want to use query ARPA model directly. You can use this function.
Actually, we use the perplexity score to
```
model = exkaldi.load_ngrams(arpaFile)
model.info
```
As the termination of this section, we generate the Grammar fst for futher steps.
```
Gfile = os.path.join(dataDir, "exp", "G.fst")
exkaldi.decode.graph.make_G(lexicons, arpaFile, outFile=Gfile, order=2)
```
| github_jupyter |
# Basic Gates Kata Workbook
**What is this workbook?**
A workbook is a collection of problems, accompanied by solutions to them. The explanations focus on the logical steps required to solve a problem; they illustrate the concepts that need to be applied to come up with a solution to the problem, explaining the mathematical steps required.
Note that a workbook should not be the primary source of knowledge on the subject matter; it assumes that you've already read a tutorial or a textbook and that you are now seeking to improve your problem-solving skills. You should attempt solving the tasks of the respective kata first, and turn to the workbook only if stuck or for reinforcement. While a textbook emphasizes knowledge acquisition, a workbook emphasizes skill acquisition.
This workbook describes the solutions to the problems offered in the [Basic Gates Kata](./BasicGates.ipynb).
Since the tasks are offered as programming problems, the explanations also cover some elements of Q# that might be non-obvious for a novitiate.
**What you should know for this workbook**
You should be familiar with the following concepts and associated techniques **prior to** beginning work on the Basic Gates Quantum Kata.
1. [Complex numbers](../tutorials/ComplexArithmetic/ComplexArithmetic.ipynb).
2. Basic linear algebra (multiplying column vectors by matrices), per the first part of [this tutorial](../tutorials/LinearAlgebra/LinearAlgebra.ipynb).
3. [The concept of qubit and its properties](../tutorials/Qubit/Qubit.ipynb).
4. [Single-qubit gates](../tutorials/SingleQubitGates/SingleQubitGates.ipynb).
You can also consult the [complete Quantum Katas learning path](https://github.com/microsoft/QuantumKatas#learning-path).
# Part 1. Single-Qubit Gates
## Task 1.1. State flip: $|0\rangle$ to $|1\rangle$ and vice versa
**Input:** A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
**Goal:** Change the state of the qubit to $\alpha |1\rangle + \beta |0\rangle$.
**Example:**
If the qubit is in state $|0\rangle$, change its state to $|1\rangle$.
If the qubit is in state $|1\rangle$, change its state to $|0\rangle$.
### Solution
We can recognize that the Pauli X gate will change the state $|0\rangle$ to $|1\rangle$ and vice versa, and $\alpha |0\rangle + \beta |1\rangle$ to $\alpha |1\rangle + \beta |0\rangle$.
As a reminder, the Pauli X gate is defined by the following matrix:
$$
X =
\begin{bmatrix}
0 & 1\\
1 & 0
\end{bmatrix}
$$
We can see how it affects, for example, the basis state $|0\rangle$:
$$X|0\rangle=
\begin{bmatrix}
0 & 1\\
1 & 0
\end{bmatrix}
\begin{bmatrix}
1\\
0
\end{bmatrix}
=
\begin{bmatrix}
0 \cdot 1 + 1 \cdot 0\\
1 \cdot 1 + 0 \cdot 0
\end{bmatrix}
=
\begin{bmatrix}
0\\
1
\end{bmatrix}
=|1\rangle
$$
Similarly, we can consider the effect of the X gate on the superposition state $|\psi\rangle = 0.6|0\rangle + 0.8|1\rangle$:
$$X|\psi\rangle=
\begin{bmatrix}
0 & 1\\
1 & 0
\end{bmatrix}
\begin{bmatrix}
0.6\\
0.8
\end{bmatrix}
=
\begin{bmatrix}
0 \cdot 0.6 + 1 \cdot 0.8\\
1 \cdot 0.6 + 0 \cdot 0.8
\end{bmatrix}
=
\begin{bmatrix}
0.8\\
0.6
\end{bmatrix}
= 0.8|0\rangle + 0.6|1\rangle
$$
```
%kata T101_StateFlip
operation StateFlip (q : Qubit) : Unit is Adj+Ctl {
X(q);
}
```
[Return to Task 1.1 of the Basic Gates kata.](./BasicGates.ipynb#Task-1.1.-State-flip:-$|0\rangle$-to-$|1\rangle$-and-vice-versa)
## Task 1.2. Basis change: $|0\rangle$ to $|+\rangle$ and $|1\rangle$ to $|-\rangle$ (and vice versa)
**Input**: A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
**Goal**: Change the state of the qubit as follows:
* If the qubit is in state $|0\rangle$, change its state to $|+\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle + |1\rangle\big)$.
* If the qubit is in state $|1\rangle$, change its state to $|-\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle - |1\rangle\big)$.
* If the qubit is in superposition, change its state according to the effect on basis vectors.
### Solution
We can recognize that the Hadamard gate changes states $|0\rangle$ and $|1\rangle$ to $|+\rangle$ and $|-\rangle$, respectively, and vice versa.
As a reminder, the Hadamard gate is defined by the following matrix:
$$
\frac{1}{\sqrt{2}}\begin{bmatrix}
1 & 1 \\
1 & -1
\end{bmatrix}
$$
For example, we can work out $H|1\rangle$ as follows:
$$
H|1\rangle=
\frac{1}{\sqrt{2}}\begin{bmatrix}
1 & 1 \\
1 & -1
\end{bmatrix}
\begin{bmatrix}
0\\
1\\
\end{bmatrix}
=
\frac{1}{\sqrt{2}}\begin{bmatrix}
1 \cdot 0 + 1 \cdot 1 \\
1 \cdot 0 + (-1) \cdot 1
\end{bmatrix}
=
\frac{1}{\sqrt{2}}\begin{bmatrix}
1\\
-1
\end{bmatrix}
= \frac{1}{\sqrt{2}} \big(|0\rangle - |1\rangle\big) = |-\rangle
$$
Similarly, we can consider the effect of the Hadamard gate on the superposition state $|\psi\rangle = 0.6|0\rangle + 0.8|1\rangle$ (rounding the numbers to 4 decimal places):
$$
H|\psi⟩ =
\frac{1}{\sqrt{2}}\begin{bmatrix}
1 & 1 \\
1 & -1
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\beta\\
\end{bmatrix}
=
\frac{1}{\sqrt{2}}\begin{bmatrix}
\alpha + \beta\\
\alpha - \beta\\
\end{bmatrix}
= 0.7071\begin{bmatrix}
1.4\\
-0.2\\
\end{bmatrix}
= \begin{bmatrix}
0.98994\\
-0.14142\\
\end{bmatrix}
= 0.9899|0\rangle - 0.1414|1\rangle
$$
```
%kata T102_BasisChange
operation BasisChange (q : Qubit) : Unit is Adj+Ctl {
H(q);
}
```
[Return to Task 1.2 of the Basic Gates kata](./BasicGates.ipynb#Task-1.2.-Basis-change:-$|0\rangle$-to-$|+\rangle$-and-$|1\rangle$-to-$|-\rangle$-(and-vice-versa)).
## Task 1.3. Sign flip: $|+\rangle$ to $|-\rangle$ and vice versa.
**Input**: A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
**Goal** : Change the qubit state to $\alpha |0\rangle - \beta |1\rangle$ (i.e. flip the sign of the $|1\rangle$ component of the superposition).
### Solution
The action of the Pauli Z gate is exactly what is required by this question.
This gate leaves the sign of the $|0\rangle$ component of the superposition unchanged but flips the sign of the $|1\rangle$ component of the superposition.
As a reminder, the Pauli Z gate is defined by the following matrix:
$$
Z =
\begin{bmatrix}
1 & 0\\
0 & -1
\end{bmatrix}
$$
Let's see its effect on the only computational basis state that it changes, $|1\rangle$:
$$
Z|1\rangle =
\begin{bmatrix}
1 & 0\\
0 & -1
\end{bmatrix}
\begin{bmatrix}
0\\
1\\
\end{bmatrix}
=
\begin{bmatrix}
1 \cdot 0 + 0 \cdot1\\
0 \cdot 1 + -1 \cdot 1\\
\end{bmatrix}
=
\begin{bmatrix}
0\\
-1\\
\end{bmatrix}
=
-\begin{bmatrix}
0\\
1\\
\end{bmatrix}
= -|1\rangle
$$
In general applying the Z gate to a single qubit superposition state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$ gives
$$
Z|\psi\rangle =
\begin{bmatrix}
1 & 0 \\
0 & -1
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\beta\\
\end{bmatrix}
=
\begin{bmatrix}
1\cdot\alpha + 0\cdot\beta\\
0\cdot\alpha + -1\cdot\beta\\
\end{bmatrix}
=
\begin{bmatrix}
\alpha\\
-\beta\\
\end{bmatrix}
= \alpha |0\rangle -\beta |1\rangle
$$
```
%kata T103_SignFlip
operation SignFlip (q : Qubit) : Unit is Adj+Ctl {
Z(q);
}
```
[Return to Task 1.3 of the Basic Gates kata](./BasicGates.ipynb#Task-1.3.-Sign-flip:-$|+\rangle$--to-$|-\rangle$--and-vice-versa.).
## Task 1.4. Amplitude change: $|0\rangle$ to $\cos{α} |0\rangle + \sin{α} |1\rangle$.
**Inputs:**
1. Angle α, in radians, represented as Double.
2. A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
**Goal:** Change the state of the qubit as follows:
- If the qubit is in state $|0\rangle$, change its state to $\cos{α} |0\rangle + \sin{α} |1\rangle$.
- If the qubit is in state $|1\rangle$, change its state to $-\sin{α} |0\rangle + \cos{α} |1\rangle$.
- If the qubit is in superposition, change its state according to the effect on basis vectors.
### Solution
We can recognize that we need to use one of the rotation gates Rx, Ry, and Rz (named because they "rotate" the qubit state in the three dimensional space visualized as the Bloch sphere about the x, y, and z axes, respectively), since they involve angle parameters. Of these three gates, only Ry rotates the basis states $|0\rangle$ and $|1\rangle$ to have real amplitudes (the other two gates introduce complex coefficients).
As a reminder,
$$
R_{y}(\theta) =
\begin{bmatrix}
\cos \frac{\theta}{2} & -\sin \frac{\theta}{2}\\
\sin \frac{\theta}{2} & \cos \frac{\theta}{2}
\end{bmatrix}
$$
Let's see its effect on the $|0\rangle$ state:
$$
R_y(\theta)|0\rangle =
\begin{bmatrix}
\cos \frac{\theta}{2} & -\sin \frac{\theta}{2}\\
\sin \frac{\theta}{2} & \cos \frac{\theta}{2}
\end{bmatrix}
\begin{bmatrix}
1\\
0\\
\end{bmatrix}
=
\begin{bmatrix}
\cos \frac{\theta}{2}\cdot1 - \sin \frac{\theta}{2}\cdot0\\
\sin \frac{\theta}{2}\cdot1 + \cos \frac{\theta}{2}\cdot0
\end{bmatrix}
=
\begin{bmatrix}
\cos \frac{\theta}{2}\\
\sin \frac{\theta}{2}
\end{bmatrix}
= \cos\frac{\theta}{2} |0\rangle + \sin\frac{\theta}{2} |1\rangle
$$
Recall that when applying a gate, you can tell what its matrix does to the basis states by looking at its columns: the first column of the matrix is the state into which it will transform the $|0\rangle$ state, and the second column is the state into which it will transform the $|1\rangle$ state.
In the example used by the testing harness we are given $\beta = 0.6, \gamma = 0.8$ and $\alpha = 1.0471975511965976 = \frac{\pi}{3}$.
Since $\cos \frac{\pi}{3} = 0.5$ and $\sin \frac{\pi}{3} = 0.8660$, working to 4 decimal places, we can compute:
$$
R_{y}(\theta) |\psi\rangle
=
\begin{bmatrix}
\cos \frac{\theta}{2} & -\sin \frac{\theta}{2}\\
\sin \frac{\theta}{2} & \cos \frac{\theta}{2}
\end{bmatrix}
\begin{bmatrix}
\beta\\
\gamma
\end{bmatrix}
=
\begin{bmatrix}
cos \frac{\theta}{2}\cdot\beta - sin \frac{\theta}{2}\cdot\gamma\\
sin \frac{\theta}{2}\cdot\beta +cos \frac{\theta}{2}\cdot\gamma
\end{bmatrix}
=
\begin{bmatrix}
0.6\cdot\cos \frac{\pi}{3} -0.8\cdot\sin \frac{\pi}{3}\\
0.6\cdot\sin \frac{\pi}{3} +0.8\cdot\cos \frac{\pi}{3}
\end{bmatrix}
=
\begin{bmatrix}
0.3 - 0.6928\\
0.5196 + 0.4
\end{bmatrix}
=
\begin{bmatrix}
-0.3928\\
0.9196
\end{bmatrix}
$$
Notice that we used $\frac{\theta}{2} = \alpha$; this means that in the Q# code we need to pass the angle $\theta = 2\alpha$.
```
%kata T104_AmplitudeChange
operation AmplitudeChange (alpha : Double, q : Qubit) : Unit is Adj+Ctl {
Ry(2.0 * alpha, q);
}
```
[Return to Task 1.4 of the Basic Gates kata](./BasicGates.ipynb#Task-1.4.-Amplitude-change:-$|0\rangle$-to-$\cos{α}-|0\rangle-+-\sin{α}-|1\rangle$.).
## Task 1.5. Phase flip
**Input:** A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
**Goal:** Change the qubit state to $\alpha |0\rangle + \color{red}i\beta |1\rangle$ (add a relative phase $i$ to $|1\rangle$ component of the superposition).
### Solution
We can recognize that the S gate performs this particular relative phase addition to the $|1\rangle$ basis state. As a reminder,
$$
S =
\begin{bmatrix}
1 & 0\\
0 & i
\end{bmatrix}
$$
Let's see the effect of this gate on the general superposition $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
$$
\begin{bmatrix}
1 & 0 \\
0 & i
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\beta\\
\end{bmatrix}
=
\begin{bmatrix}
1\cdot\alpha + 0\cdot\beta\\
0\cdot\alpha + i\cdot\beta
\end{bmatrix}
=
\begin{bmatrix}
\alpha\\
i\beta\\
\end{bmatrix}
$$
It is therefore easy to see that when $|\psi\rangle = 0.6|0\rangle + 0.8|1\rangle, S|\psi\rangle = 0.6|0\rangle + 0.8i|1\rangle$.
```
%kata T105_PhaseFlip
operation PhaseFlip (q : Qubit) : Unit is Adj+Ctl {
S(q);
}
```
### Solution 2
See the next task, Phase Change, for an explanation of using R1 gate to implement the same transformation:
```
%kata T105_PhaseFlip
open Microsoft.Quantum.Math;
operation PhaseFlip (q : Qubit) : Unit is Adj+Ctl {
R1(0.5 * PI(), q);
}
```
[Return to Task 1.5 of the Basic Gates kata](./BasicGates.ipynb#Task-1.5.-Phase-flip).
## Task 1.6. Phase change
**Inputs:**
1. Angle α, in radians, represented as Double.
2. A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
**Goal:** Change the state of the qubit as follows:
- If the qubit is in state $|0\rangle$, don't change its state.
- If the qubit is in state $|1\rangle$, change its state to $e^{i\alpha} |1\rangle$.
- If the qubit is in superposition, change its state according to the effect on basis vectors: $\beta |0\rangle + \color{red}{e^{i\alpha}} \gamma |1\rangle$.
### Solution
We know that:
$$
R1(\alpha)
=
\begin{bmatrix}
1 & 0\\
0 & \color{red}{e^{i\alpha}}
\end{bmatrix}
$$
So we have:
$$
R1(\beta |0\rangle + \gamma |1\rangle) =
\begin{bmatrix}
1 & 0 \\
0 & \color{red}{e^{i\alpha}}
\end{bmatrix}
\begin{bmatrix}
\beta\\
\gamma\\
\end{bmatrix}
=
\begin{bmatrix}
1.\beta + 0.\gamma\\
0.\beta + \color{red}{e^{i\alpha}}\gamma
\end{bmatrix}
=
\begin{bmatrix}
\beta\\
\color{red}{e^{i\alpha}}\gamma
\end{bmatrix}
= \beta |0\rangle + \color{red}{e^{i\alpha}} \gamma |1\rangle
$$
> Note that the results produced by the test harness can be unexpected.
If you run the kata several times and examine the output, you'll notice that success is signaled even though the corresponding amplitudes of the desired and actual states look very different.
>
> So what's going on? The full state simulator used in these tests performs the computations "up to a global phase", that is, sometimes the resulting state acquires a global phase that doesn't affect the computations or the measurement outcomes, but shows up in DumpMachine output. (You can read more about the global phase in the [Qubit tutorial](../tutorials/Qubit/Qubit.ipynb#Relative-and-Global-Phase).)
>
> For example, in one run you can get the desired state $(0.6000 + 0000i)|0\rangle + (-0.1389 +0.7878i)|1\rangle$ and the actual state $(-0.1042 + 0.5909i)|0\rangle + (-0.7518 -0.2736i)|1\rangle$.
You can verify that the ratios of amplitudes of the respective basis states are equal: $\frac{-0.1042 + 0.5909i}{0.6} = -0.173667 +0.984833 i = \frac{-0.7518 -0.2736i}{-0.1389 +0.7878i}$, so the global phase acquired by the state is (-0.173667 +0.984833 i).
You can also check that the absolute value of this multiplier is approximately 1, so it doesn't impact the measurement probabilities.
>
> The testing harness for this and the rest of the tasks checks that your solution implements the required transformation exactly, without introducing any global phase, so it shows up only in the helper output and does not affect the verification of your solution.
```
%kata T106_PhaseChange
operation PhaseChange (alpha : Double, q : Qubit) : Unit is Adj+Ctl {
R1(alpha, q);
}
```
Suppose now that $\alpha = \frac{\pi}{2}$.
Then $e^{i\alpha}= \cos\frac{\pi}{2} + i\sin\frac{\pi}{2}$.
And, since $\cos\frac{\pi}{2}= 0$ and $\sin\frac{\pi}{2} = 1$, then we have that $\cos\frac{\pi}{2} + i \sin\frac{\pi}{2} = i$, and
$R1(\frac{\pi}{2}) = S$, which we used in the second solution to task 1.5, above.
[Return to Task 1.6 of the Basic Gates kata](./BasicGates.ipynb#Task-1.6.-Phase-Change).
## Task 1.7. Global phase change
**Input:** A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
**Goal**: Change the state of the qubit to $- \beta |0\rangle - \gamma |1\rangle$.
> Note: this change on its own is not observable - there is no experiment you can do on a standalone qubit to figure out whether it acquired the global phase or not.
> However, you can use a controlled version of this operation to observe the global phase it introduces.
> This is used in later katas as part of more complicated tasks.
### Solution
We recognize that a global phase change can be accomplished by using the R rotation gate with the PauliI (identity) gate.
As a reminder, the R gate is defined as $R_{\mu}(\theta) = \exp(\frac{\theta}{2}i\cdot\sigma_{\mu})$, where $\sigma_{\mu}$ is one of the Pauli gates I, X, Y or Z.
> Note that a global phase is not detectable and has no physical meaning - it disappears when you take a measurement of the state.
> You can read more about this in the [Single-qubit measurements tutorial](../tutorials/SingleQubitSystemMeasurements/SingleQubitSystemMeasurements.ipynb#Measurements-in-arbitrary-orthogonal-bases).
For the problem at hand, we'll use the rotation gate $R_{\mu}(\theta) = \exp(\frac{\theta}{2}i\cdot\sigma_{\mu})$ with $\sigma_{\mu} = I$.
$R(PauliI, 2\pi) = \exp(\frac{2\pi}{2} iI) = \exp(i\pi) I = (\cos\pi + i\sin\pi) I$ and, since $\cos\pi = -1$ and $\sin\pi = 0$, we have that $R(PauliI, 2\pi) = -I$:
$$
R(\beta |0\rangle + \gamma |1\rangle)
=
-1\begin{bmatrix}
1 & 0 \\
0 & 1
\end{bmatrix}
\begin{bmatrix}
\beta\\
\gamma\\
\end{bmatrix}
=
\begin{bmatrix}
-1 & 0 \\
0 & -1
\end{bmatrix}
\begin{bmatrix}
\beta\\
\gamma\\
\end{bmatrix}
=
\begin{bmatrix}
-1\cdot\beta + 0\cdot\gamma\\
0\cdot\beta + -1\cdot\gamma \\
\end{bmatrix}
=
\begin{bmatrix}
-\beta\\
-\gamma\\
\end{bmatrix}
=
- \beta |0\rangle - \gamma |1\rangle
$$
The test harness for this test shows the result of applying the *controlled* variant of your solution to be able to detect the phase change.
```
%kata T107_GlobalPhaseChange
open Microsoft.Quantum.Math;
operation GlobalPhaseChange (q : Qubit) : Unit is Adj+Ctl {
R(PauliI, 2.0 * PI(), q);
}
```
[Return to Task 1.7 of the Basic Gates kata](./BasicGates.ipynb#Task-1.7.-Global-phase-change).
## Task 1.8. Bell state change - 1
**Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
**Goal:** Change the two-qubit state to $|\Phi^{-}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle - |11\rangle\big)$.
### Solution
We recognize that the goal is another Bell state. In fact, it is one of the four Bell states.
We remember from Task 1.3 that the Pauli Z gate will change the state of the $|0\rangle$ basis state of a single qubit, so this gate seems like a good candidate for what we want to achieve. This gate leaves the sign of the $|0\rangle$ basis state of a superposition unchanged, but flips the sign of the $|1\rangle$ basis state of the superposition.
Don't forget that the Z gate acts on only a single qubit, and we have two here.
Lets also remember how the Bell state is made up from its individual qubits.
If the two qubits are A and B, where A is `qs[0]` and B is `qs[1]`, we can write that
$|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|0_{A}0_{B}\rangle + |1_{A}1_{B}\rangle\big)$, and also remember that this is a superposition.
If we apply the Z gate to the qubit A, it will flip the phase of the basis state $|1_A\rangle$. As this phase is in a sense spread across the entangled state, with $|1_A\rangle$ basis state being part of the second half of the superposition, this application has the effect of flipping the sign of the whole basis state $|1_A1_B\rangle$, as you can see by running the solution below.
> If you run the solution below several times, you will notice that you get the state $\frac{1}{\sqrt{2}} \big(-|00\rangle + |11\rangle\big)$ at a guess as frequently as the goal state. Here is why.
>
> We can write this second state as $-\frac{1}{\sqrt{2}} \big(|00\rangle -|11\rangle\big)$ = $(-1)\frac{1}{\sqrt{2}} \big(|00\rangle - |11\rangle\big)$.
This state is the same as our goal state $|\Phi^{-}\rangle$ with an unmeasurable global phase of $-1$.
The exact same calculations can be done if we apply Z to the qubit B, so that's another possible solution.
```
%kata T108_BellStateChange1
operation BellStateChange1 (qs : Qubit[]) : Unit is Adj+Ctl {
Z(qs[0]); // or Z(qs[1]);
}
```
[Return to Task 1.8 of the Basic Gates kata](./BasicGates.ipynb#Task-1.8.-Bell-state-change---1).
## Task 1.9. Bell state change - 2
**Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
**Goal:** Change the two-qubit state to $|\Psi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|01\rangle + |10\rangle\big)$.
### Solution ##
We have seen in Task 1.1 that the Pauli X gate flips $|0\rangle$ to $|1\rangle$ and vice versa, and as we seem to need some flipping of states, perhaps this gate may be of use. (Bearing in mind, of course, that the X gate operates on a single qubit).
Let's compare the starting state $\frac{1}{\sqrt{2}} \big(|0_A0_B\rangle + |1_A1_B\rangle\big)$ with the goal state $\frac{1}{\sqrt{2}} \big(1_A0_B\rangle + |0_A1_B\rangle\big)$ term by term and see how we need to transform it to reach the goal.
Using our nomenclature from Task 1.8, we can now see by comparing terms that $|0_{A}\rangle$ has flipped to $|1_A\rangle$ to get the first term, and $|1_{A}\rangle$ has flipped to $|0\rangle$ to get the second term. This allows us to say that the correct gate to use is Pauli X, applied to `qs[0]`.
```
%kata T109_BellStateChange2
operation BellStateChange2 (qs : Qubit[]) : Unit is Adj+Ctl {
X(qs[0]); // or X(qs[1]);
}
```
[Return to Task 1.9 of the Basic Gates kata](./BasicGates.ipynb#Task-1.9.-Bell-state-change---2).
## Task 1.10. Bell state change - 3
**Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
**Goal:** Change the two-qubit state, without adding a global phase, to $|\Psi^{-}\rangle = \frac{1}{\sqrt{2}} \big(|01\rangle - |10\rangle\big)$.
### Solution ##
We remember from Task 1.3 that the Pauli-Z gate leaves the sign of the $|0\rangle$ component of the single qubit superposition unchanged but flips the sign of the $|1\rangle$ component of the superposition. We have also just seen in Task 1.9 how to change our input state to the state $\frac{1}{\sqrt{2}} \big(|01\rangle + |10\rangle\big)$, which is almost our goal state (disregarding the phase change for the moment). So it would seem that a combination of these two gates will be what we need here. The remaining question is in what order to apply them, and to which qubit.
First of all, which qubit? Looking back at Task 1.9, it seems clear that we need to use qubit `qs[0]`, like we did there.
Second, in what order should we apply the gates? Remember that the Pauli-Z gate flips the phase of the $|1\rangle$ component of the superposition and leaves the $|0\rangle$ component alone.
Let's experiment with applying X to `qs[0]` first. Looking at our "halfway answer" state $\frac{1}{\sqrt{2}} \big(|01\rangle + |10\rangle\big)$, we can see that if we apply the Z gate to `qs[0]`, it will leave the $|0_{A}\rangle$ alone but flip the phase of $|1_{A}\rangle$ to $-|1_{A}\rangle$, thus flipping the phase of the $|11\rangle$ component of our Bell state.
```
%kata T110_BellStateChange3
operation BellStateChange3 (qs : Qubit[]) : Unit is Adj+Ctl {
X(qs[0]);
Z(qs[0]);
}
```
[Return to Task 1.10 of the Basic Gates kata](./BasicGates.ipynb#Task-1.10.-Bell-state-change---3).
# Part II. Multi-Qubit Gates
## Task 2.1. Two-qubit gate - 1
**Input:** Two unentangled qubits (stored in an array of length 2).
The first qubit will be in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$, the second - in state $|0\rangle$
(this can be written as two-qubit state $\big(\alpha |0\rangle + \beta |1\rangle \big) \otimes |0\rangle = \alpha |00\rangle + \beta |10\rangle$.
**Goal:** Change the two-qubit state to $\alpha |00\rangle + \beta |11\rangle$.
### Solution
Let's denote the first qubit in state $\alpha |0\rangle + \beta |1\rangle$ as A and the second qubit in state $|0\rangle$ as B.
Compare our input state $\alpha |0_A0_B\rangle + \beta |1_A0_B\rangle$ with the goal state $\alpha |0_A0_B\rangle + \beta |1_A1_B\rangle$.
We want to pass our input qubit through a gate or gates (to be decided) that do the following. If qubit A is in the $|0\rangle$ state, then we want to leave qubit B alone (the first term of the superposition).
However, if A is in the $|1\rangle$ state, we want to flip qubit B from $|0\rangle$ into $|1\rangle$ state. In other words, the state of B is to be made contingent upon the state of A.
This gate exists and is called a CNOT (controlled-not) gate. Depending upon the state of the **control** qubit (A in our case), the value of the controlled or **target** qubit (B in our case) is inverted or unchanged. Thus we get the goal state $\alpha |00\rangle + \beta |11\rangle$.
```
%kata T201_TwoQubitGate1
operation TwoQubitGate1 (qs : Qubit[]) : Unit is Adj+Ctl {
CNOT(qs[0], qs[1]);
}
```
[Return to Task 2.1 of the Basic Gates kata](./BasicGates.ipynb#Task-2.1.-Two-qubit-gate---1).
## Task 2.2. Two-qubit gate - 2
**Input:** Two unentangled qubits (stored in an array of length 2) in state $|+\rangle \otimes |+\rangle = \frac{1}{2} \big( |00\rangle + |01\rangle + |10\rangle \color{blue}+ |11\rangle \big)$.
**Goal:** Change the two-qubit state to $\frac{1}{2} \big( |00\rangle + |01\rangle + |10\rangle \color{red}- |11\rangle \big)$.
### Solution
Firstly we notice that we are dealing with an unentangled pair of qubits. In vector form this is:
In vector form the transformation we need is
$$\frac{1}{2}
\begin{bmatrix}
1\\
1\\
1\\
1\\
\end{bmatrix} \rightarrow
\frac{1}{2}
\begin{bmatrix}
1\\
1\\
1\\
-1\\
\end{bmatrix}
$$
All that needs to happen to change the input into the goal is that the $|11\rangle$ basis state needs to have its sign flipped.
We remember that the Pauli Z gate flips signs in the single qubit case, so we need to investigate if there is a 2-qubit version of this gate that we can use here. We can also recall task 1.6 which dealt with phase shifts and, remembering that $e^{i\cdot\pi} = -1$, we can think of the transformation we're looking for as a phase shift.
It can be useful to investigate a general case and then use it to perform a specific state change, so let's look for a 2-qubit variant of the phase shift.
Similarly to task 2.1, the phase shift only occurs on one of the basis states, so this suggests it might be a conditional shift. If we could have our phase shift applied to `qs[1]` conditional on `qs[0]` being in the state $|1\rangle$, then we would have a description of our gate. If we now look though a list of gates in the [Single-qubit gates tutorial](../tutorials/SingleQubitGates/SingleQubitGates.ipynb), we'll find the R1 phase shift gate with angle parameter $\theta$ (radians), defined as
$$
R1(\alpha)
=
\begin{bmatrix}
1 & 0\\
0 & \color{red}{e^{i\alpha}}
\end{bmatrix}
$$
The controlled variant of this gate will look like this:
$$
CR1(\alpha) =
\begin{bmatrix}
1 & 0 & 0 & 0\\
0 & 1 & 0 & 0\\
0 & 0 & 1 & 0\\
0 & 0 & 0 &\color{red}{e^{i\alpha}}
\end{bmatrix}
$$
This gate is almost Pauli I, the identity gate, with the different in just the last column, showing what will happen to the $|11\rangle$ basis state. Applying it to our input state for $\alpha = \pi$, we'll get:
$$
\frac{1}{2}
\begin{bmatrix}
1 & 0 & 0 & 0\\
0 & 1 & 0 & 0\\
0 & 0 & 1 & 0\\
0 & 0 & 0 &\color{red}{e^{i\alpha}}
\end{bmatrix}
\begin{bmatrix}
1\\
1\\
1\\
1\\
\end{bmatrix}
=
\frac{1}{2}
\begin{bmatrix}
1\\
1\\
1\\
1\cdot\color{red}{e^{i\alpha}}\\
\end{bmatrix}
=
\frac{1}{2} \big( |00\rangle + |01\rangle + |10\rangle \color{red}- |11\rangle \big)
$$
The last thing we notice if we look through the [list of operations in the Microsoft.Quantum.Canon namespace](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.canon) is the CZ (Controlled Z) gate, a special case of CR1 that implements exactly this gate.
```
%kata T202_TwoQubitGate2
operation TwoQubitGate2 (qs : Qubit[]) : Unit is Adj+Ctl {
CZ(qs[0], qs[1]);
}
```
Alternatively, we can express this gate using the intrinsic gate Z and its controlled variant using the Controlled functor:
```
%kata T202_TwoQubitGate2
operation TwoQubitGate2 (qs : Qubit[]) : Unit is Adj+Ctl {
Controlled Z([qs[0]], qs[1]);
}
```
[Return to Task 2.2 of the Basic Gates kata](./BasicGates.ipynb#Task-2.2.-Two-qubit-gate---2).
## Task 2.3. Two-qubit gate - 3
**Input:** Two unentangled qubits (stored in an array of length 2) in an arbitrary two-qubit state $\alpha |00\rangle + \color{blue}\beta |01\rangle + \color{blue}\gamma |10\rangle + \delta |11\rangle$.
**Goal:** Change the two-qubit state to $\alpha |00\rangle + \color{red}\gamma |01\rangle + \color{red}\beta |10\rangle + \delta |11\rangle$.
> This task can be solved using one intrinsic gate; as an exercise, try to express the solution using several (possibly controlled) Pauli gates.
## Solution
A visual comparison of the two states easily reveals that the amplitudes of the $|01\rangle$ and the $|01\rangle$ components of the state have been swapped. This suggests that we might look for a swap gate that operates on 2 qubits, by changing the components of the 2 qubits to which the amplitudes are 'attached'.
Let's investigate the first possibility. There is a swap gate that might fit the bill; its matrix representation is:
$$
SWAP =
\begin{bmatrix}
1 & 0 & 0 & 0\\
0 & 0 & 1 & 0\\
0 & 1 & 0 & 0\\
0 & 0 & 0 & 1\\
\end{bmatrix}
$$
and our input state vector is:
$$
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\end{bmatrix}$$
So operating on our input state vector with the SWAP gate gives us
$$
\begin{bmatrix}
1 & 0 & 0 & 0\\
0 & 0 & 1 & 0\\
0 & 1 & 0 & 0\\
0 & 0 & 0 & 1\\
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\color{blue}\beta\\
\color{blue}\gamma\\
\delta\\
\end{bmatrix}
=
\begin{bmatrix}
\alpha\\
\color{red}\gamma\\
\color{red}\beta\\
\delta\\
\end{bmatrix}
=
|00\rangle + \color{red}\gamma |01\rangle + \color{red}\beta |10\rangle + \delta |11\rangle
$$
and we can confirm this with the task solution:
```
%kata T203_TwoQubitGate3
operation TwoQubitGate3 (qs : Qubit[]) : Unit is Adj+Ctl {
SWAP(qs[0], qs[1]);
}
```
> If you run this solution a few times you might see an apparent anomaly. The test harness uses an input state that has positive values of $\alpha$ and $\delta$ and negative values of $\beta$ and $\gamma$, while
the "actual state" reported (the state prepared by your solution) can come out with negative values of $\alpha$ and $\delta$ and positive values of $\beta$ and $\gamma$.
We have seen this before in the previous tasks: we can write the apparently anomalous state as $(-1)(\alpha|00\rangle + \beta |01\rangle + \gamma |10\rangle + \delta |11\rangle)$ and see that it differs from the goal state by a global phase of $\pi$ (remember that $e^{i\pi}=-1$). This doesn't mean that your implementation introduced this phase; sometimes the full state simulator used in the test harness produces a global phase in its calculations.
Let's now follow the hint in the question and try to express the solution using several (possibly controlled) Pauli gates.
If we look at the available cotrolled gates, CR and its special case CZ produce rotations, and that's not really what we want. So perhaps we are being pointed towards CNOT? If we carefully compare the input with the goal state, we see that the bits in the two basis states of the two qubits are being flipped, which results in a swap. What we need to do is to turn $|01\rangle$ into $|10\rangle$ and $|10\rangle$ into $|01\rangle$ while leaving the other two basis states unchanged.
With some experimentation with sequences of CNOT gates we can arrive at the following sequence of transformations:
<table>
<col width="150"/>
<col width="150"/>
<col width="150"/>
<col width="150"/>
<tr>
<th style="text-align:center">Starting state</th>
<th style="text-align:center">After CNOT$_{01}$</th>
<th style="text-align:center">After CNOT$_{10}$</th>
<th style="text-align:center">After CNOT$_{01}$</th>
</tr>
<tr>
<td style="text-align:center">$|00\rangle$</td>
<td style="text-align:center">$|00\rangle$</td>
<td style="text-align:center">$|00\rangle$</td>
<td style="text-align:center">$|00\rangle$</td>
</tr>
<tr>
<td style="text-align:center">$|01\rangle$</td>
<td style="text-align:center">$|01\rangle$</td>
<td style="text-align:center">$|11\rangle$</td>
<td style="text-align:center">$|10\rangle$</td>
</tr>
<tr>
<td style="text-align:center">$|10\rangle$</td>
<td style="text-align:center">$|11\rangle$</td>
<td style="text-align:center">$|01\rangle$</td>
<td style="text-align:center">$|01\rangle$</td>
</tr>
<tr>
<td style="text-align:center">$|11\rangle$</td>
<td style="text-align:center">$|10\rangle$</td>
<td style="text-align:center">$|10\rangle$</td>
<td style="text-align:center">$|11\rangle$</td>
</tr>
</table>
```
%kata T203_TwoQubitGate3
operation TwoQubitGate3 (qs : Qubit[]) : Unit is Adj+Ctl {
CNOT(qs[0], qs[1]);
CNOT(qs[1], qs[0]);
CNOT(qs[0], qs[1]);
}
```
[Return to Task 2.3 of the Basic Gates kata](./BasicGates.ipynb#Task-2.3.-Two-qubit-gate---3).
## Task 2.4. Toffoli gate
**Input:** Three qubits (stored in an array of length 3) in an arbitrary three-qubit state
$\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \zeta|101\rangle + \color{blue}\eta|110\rangle + \color{blue}\theta|111\rangle$.
**Goal:** Flip the state of the third qubit if the state of the first two is $|11\rangle$, i.e., change the three-qubit state to $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \zeta|101\rangle + \color{red}\theta|110\rangle + \color{red}\eta|111\rangle$.
### Solution
This is essentially bookwork, because there is only one gate that performs this state change (and the task title already gave it away!) The Toffoli gate is:
$$
\begin{bmatrix}
1 & 0 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 1 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 1 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 1 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 1 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 1 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 0 & 1\\
0 & 0 & 0 & 0 & 0 & 0 & 1 & 0\\
\end{bmatrix}
$$
and our initial state is:
$$
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\zeta\\
\eta\\
\theta\\
\end{bmatrix}
$$
So we have:
$$
\begin{bmatrix}
1 & 0 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 1 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 1 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 1 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 1 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 1 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 0 & 1\\
0 & 0 & 0 & 0 & 0 & 0 & 1 & 0\\
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\zeta\\
\color{blue}\eta\\
\color{blue}\theta\\
\end{bmatrix}
=
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\zeta\\
\color{red}\theta\\
\color{red}\eta\\
\end{bmatrix}
=
\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \zeta|101\rangle + \color{red}\theta|110\rangle + \color{red}\eta|111\rangle
$$
```
%kata T204_ToffoliGate
operation ToffoliGate (qs : Qubit[]) : Unit is Adj+Ctl {
CCNOT(qs[0], qs[1], qs[2]);
}
```
[Return to Task 2.4 of the Basic Gates kata](./BasicGates.ipynb#Task-2.4.-Toffoli-gate).
## Task 2.5. Fredkin gate
**Input:** Three qubits (stored in an array of length 3) in an arbitrary three-qubit state
$\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \color{blue}\zeta|101\rangle + \color{blue}\eta|110\rangle + \theta|111\rangle$.
**Goal:** Swap the states of second and third qubit if and only if the state of the first qubit is $|1\rangle$, i.e., change the three-qubit state to $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \color{red}\eta|101\rangle + \color{red}\zeta|110\rangle + \theta|111\rangle$.
### Solution
Again this is essentially bookwork, because there is only one gate that performs this state change (and the task title already gave it away!)
The Fredkin gate is also known as the controlled swap gate (Controlled SWAP):
$$
\begin{bmatrix}
1 & 0 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 1 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 1 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 1 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 1 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 1 & 0\\
0 & 0 & 0 & 0 & 0 & 1 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 0 & 1\\
\end{bmatrix}
$$
and our initial state is:
$$
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\zeta\\
\eta\\
\theta\\
\end{bmatrix}
$$
So we have:
$$
\begin{bmatrix}
1 & 0 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 1 & 0 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 1 & 0 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 1 & 0 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 1 & 0 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 1 & 0\\
0 & 0 & 0 & 0 & 0 & 1 & 0 & 0\\
0 & 0 & 0 & 0 & 0 & 0 & 0 & 1\\
\end{bmatrix}
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\color{blue} \zeta\\
\color{blue} \eta\\
\theta\\
\end{bmatrix}
=
\begin{bmatrix}
\alpha\\
\beta\\
\gamma\\
\delta\\
\epsilon\\
\color{red} \eta\\
\color{red} \zeta\\
\theta\\
\end{bmatrix}
=
\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \color{red}\eta|101\rangle + \color{red}\zeta|110\rangle + \theta|111\rangle
$$
Notice carefully how the qubits are passed to the gate: `[qs[0]], (qs[1], [qs[2])`. The `Controlled` functor produces an operation that takes two parameters: the first one is an array of control qubits (in this case a single-element array consisting of the first qubit), and the second parameter is a tuple of all parameters you'd pass to the original gate (in this gate two single-qubit parameters that would be arguments to a SWAP gate).
```
%kata T205_FredkinGate
operation FredkinGate (qs : Qubit[]) : Unit is Adj+Ctl {
Controlled SWAP([qs[0]], (qs[1], qs[2]));
}
```
[Return to Task 2.5 of the Basic Gates kata](./BasicGates.ipynb#Task-2.5.-Fredkin-gate).
| github_jupyter |
# Fidelity
## Implementation
```
import os.path
import pandas as pd
import numpy as np
from os import listdir
from os.path import isfile, join
def bin_data(dt1, dt2, c = 10):
dt1 = dt1.copy()
dt2 = dt2.copy()
# quantile binning of numerics
num_cols = dt1.dtypes[dt1.dtypes!='object'].index
for col in num_cols:
# determine breaks based on `dt1`
breaks = dt1[col].quantile(np.linspace(0, 1, c+1)).unique()
dt1[col] = pd.cut(dt1[col], bins=breaks, include_lowest=True).astype(str)
dt2_vals = pd.to_numeric(dt2[col], 'coerce')
dt2_bins = pd.cut(dt2_vals, bins=breaks, include_lowest=True).astype(str)
dt2_bins[dt2_vals < min(breaks)] = '_other_'
dt2_bins[dt2_vals > max(breaks)] = '_other_'
dt2[col] = dt2_bins
# top-C binning of categoricals
cat_cols = dt1.dtypes[dt1.dtypes=='object'].index
for col in cat_cols:
# determine top values based on `dt1`
top_vals = dt1[col].value_counts().head(c).index.tolist()
dt1[col].replace(np.setdiff1d(dt1[col].unique().tolist(), top_vals), '_other_', inplace=True)
dt2[col].replace(np.setdiff1d(dt2[col].unique().tolist(), top_vals), '_other_', inplace=True)
return [dt1, dt2]
def hellinger(p1, p2):
return np.sqrt(1 - np.sum(np.sqrt(p1*p2)))
def kullback_leibler(p1, p2):
idx = p1>0
return np.sum(p1[idx] * np.log(p1[idx]/p2[idx]))
def jensen_shannon(p1, p2):
m = 0.5 * (p1 + p2)
return 0.5 * kullback_leibler(p1, m) + 0.5 * kullback_leibler(p2, m)
def fidelity(dt1, dt2, c = 100, k = 1):
[dt1_bin, dt2_bin] = bin_data(dt1, dt2, c = c)
# build grid of all cross-combinations
cols = trn.columns
interactions = pd.DataFrame(np.array(np.meshgrid(cols, cols, cols)).reshape(3, len(cols)**3).T)
interactions.columns = ['dim1', 'dim2', 'dim3']
if k == 1:
interactions = interactions.loc[(interactions['dim1']==interactions['dim2']) & (interactions['dim2']==interactions['dim3'])]
elif k == 2:
interactions = interactions.loc[(interactions['dim1']<interactions['dim2']) & (interactions['dim2']==interactions['dim3'])]
elif k == 3:
interactions = interactions.loc[(interactions['dim1']<interactions['dim2']) & (interactions['dim2']<interactions['dim3'])]
else:
raise('k>3 not supported')
results = []
for idx in range(interactions.shape[0]):
row = interactions.iloc[idx]
val1 = dt1_bin[row.dim1] + dt1_bin[row.dim2] + dt1_bin[row.dim3]
val2 = dt2_bin[row.dim1] + dt2_bin[row.dim2] + dt2_bin[row.dim3]
freq1 = val1.value_counts(normalize=True).to_frame(name='p1')
freq2 = val2.value_counts(normalize=True).to_frame(name='p2')
freq = freq1.join(freq2, how='outer').fillna(0.0)
p1 = freq['p1']
p2 = freq['p2']
out = pd.DataFrame({
'k': k,
'dim1': [row.dim1], 'dim2': [row.dim2], 'dim3': [row.dim3],
'tvd': [np.sum(np.abs(p1 - p2)) / 2],
'mae': [np.mean(np.abs(p1 - p2))],
'max': [np.max(np.abs(p1 - p2))],
'l1d': [np.sum(np.abs(p1 - p2))],
'l2d': [np.sqrt(np.sum((p1 - p2)**2))],
'hellinger': [hellinger(p1, p2)],
'jensen_shannon': [jensen_shannon(p1, p2)]})
results.append(out)
return pd.concat(results)
```
## Test Drive
```
trn = pd.read_csv('data/credit-default_trn.csv.gz')
syn = pd.read_csv('data/credit-default_mostly.csv.gz')
#syn = pd.read_csv('data/credit-default_synthpop.csv.gz')
fidelity(trn, syn, k=1, c=100).agg('mean')
```
## Benchmark
```
# benchmark all
datasets = ['adult', 'credit-default', 'bank-marketing', 'online-shoppers']
fns = ['mostly', 'copulagan', 'ctgan', 'tvae', 'gaussian_copula', 'gretel', 'synthpop',
'mostly_e1', 'mostly_e2', 'mostly_e4', 'mostly_e8', 'mostly_e16',
'flip10', 'flip20', 'flip30', 'flip40', 'flip50',
'flip60', 'flip70', 'flip80', 'flip90',
'val']
results = []
for dataset in datasets:
trn = pd.read_csv('data/' + dataset + '_trn.csv.gz')
for fn in fns:
syn_fn = 'data/' + dataset + '_' + fn + '.csv.gz'
print(syn_fn)
if (os.path.exists(syn_fn)):
syn = pd.read_csv(syn_fn)
fid1 = fidelity(trn, syn, k=1, c=100)
fid2 = fidelity(trn, syn, k=2, c=10)
fid3 = fidelity(trn, syn, k=3, c=5)
out = pd.concat([fid1, fid2, fid3])
out['dataset'] = dataset
out['synthesizer'] = fn
results.append(out)
x = pd.concat(results)
x.to_csv('fidelity.csv', index=False)
x
x.groupby(['dataset', 'synthesizer', 'k']).agg('mean').head(20)
```
| github_jupyter |
```
# Import libraries
import numpy as np
import pandas as pd
import scipy as sp
import matplotlib
import matplotlib.pyplot as plt
from matplotlib.font_manager import FontProperties # for unicode fonts
import psycopg2
import sys
import datetime as dt
import mp_utils as mp
import sklearn
from sklearn.pipeline import Pipeline
# used for train/test splits and cross validation
from sklearn.cross_validation import train_test_split
from sklearn.grid_search import GridSearchCV, RandomizedSearchCV
from sklearn.ensemble import BaggingClassifier
# used to impute mean for data and standardize for computational stability
from sklearn.preprocessing import Imputer
from sklearn.preprocessing import StandardScaler
# logistic regression is our favourite model ever
from sklearn.linear_model import LogisticRegression
from sklearn.linear_model import LogisticRegressionCV # l2 regularized regression
from sklearn.linear_model import LassoCV
# used to calculate AUROC/accuracy
from sklearn import metrics
# used to create confusion matrix
from sklearn.metrics import confusion_matrix
from sklearn.cross_validation import cross_val_score
# gradient boosting - must download package https://github.com/dmlc/xgboost
import xgboost as xgb
# default colours for prettier plots
col = [[0.9047, 0.1918, 0.1988],
[0.2941, 0.5447, 0.7494],
[0.3718, 0.7176, 0.3612],
[1.0000, 0.5482, 0.1000],
[0.4550, 0.4946, 0.4722],
[0.6859, 0.4035, 0.2412],
[0.9718, 0.5553, 0.7741],
[0.5313, 0.3359, 0.6523]];
marker = ['v','o','d','^','s','o','+']
ls = ['-','-','-','-','-','s','--','--']
%matplotlib inline
from __future__ import print_function
```
# Notebook outline
This notebook will evaluate the mortality prediction model in the following contexts:
* using a random time segment for each patient in the data
* training using a random time segment, but evaluating using:
* 4 hours before death (or random time, if lived)
* same as the above, but at 8, 16, and 24 hours before death
```
# below config used on pc70
sqluser = 'alistairewj'
dbname = 'mimic'
schema_name = 'mimiciii'
# Connect to local postgres version of mimic
con = psycopg2.connect(dbname=dbname, user=sqluser)
cur = con.cursor()
cur.execute('SET search_path to ' + schema_name)
# exclusion criteria:
# - less than 16 years old
# - stayed in the ICU less than 4 hours
# - never have any chartevents data (i.e. likely administrative error)
query = \
"""
with t1 as
(
select ie.icustay_id
, adm.HOSPITAL_EXPIRE_FLAG
, ROW_NUMBER() over (partition by ie.subject_id order by intime) as rn
from icustays ie
inner join admissions adm
on ie.hadm_id = adm.hadm_id
inner join patients pat
on ie.subject_id = pat.subject_id
and ie.intime > (pat.dob + interval '16' year)
where adm.HAS_CHARTEVENTS_DATA = 1
and
not (
(lower(diagnosis) like '%organ donor%' and deathtime is not null)
or (lower(diagnosis) like '%donor account%' and deathtime is not null)
)
and (ie.outtime - ie.intime) >= interval '4' hour
)
select
icustay_id
, HOSPITAL_EXPIRE_FLAG
from t1
"""
co = pd.read_sql_query(query,con)
co.set_index('icustay_id',inplace=True)
# extract static vars into a separate dataframe
df_static = pd.read_sql_query('select * from mpap_static_vars',con)
for dtvar in ['intime','outtime','deathtime']:
df_static[dtvar] = pd.to_datetime(df_static[dtvar])
df_static.set_index('icustay_id',inplace=True)
cur.close()
con.close()
vars_static = [u'male', u'emergency', u'age',
u'cmed', u'csurg', u'surg', u'nsurg',
u'surg_other', u'traum', u'nmed',
u'omed', u'ortho', u'gu', u'gyn', u'ent']
```
# connect to the database and extract severity of illness scores
```
# Connect to local postgres version of mimic
con = psycopg2.connect(dbname=dbname, user=sqluser)
cur = con.cursor()
cur.execute('SET search_path to ' + schema_name)
query = \
"""
select
icustay_id
, oasis
from oasis
"""
oa = pd.read_sql_query(query,con)
oa.set_index('icustay_id',inplace=True)
cur.execute('SET search_path to ' + schema_name)
query = \
"""
select s.icustay_id, s.sofa
from sofa s
order by s.icustay_id
"""
sofa = pd.read_sql_query(query,con)
sofa.set_index('icustay_id',inplace=True)
cur.execute('SET search_path to ' + schema_name)
query = \
"""
select s.icustay_id, s.saps
from saps s
order by s.icustay_id
"""
saps = pd.read_sql_query(query,con)
saps.set_index('icustay_id',inplace=True)
cur.execute('SET search_path to ' + schema_name)
query = \
"""
select s.icustay_id, s.sapsii
from sapsii s
order by s.icustay_id
"""
sapsii = pd.read_sql_query(query,con)
sapsii.set_index('icustay_id',inplace=True)
cur.execute('SET search_path to ' + schema_name)
query = \
"""
select icustay_id
, APSIII
from apsiii
order by icustay_id
"""
apsiii = pd.read_sql_query(query,con)
apsiii.set_index('icustay_id',inplace=True)
cur.close()
con.close()
```
## Model 1: Using first 24 hours of data
Now we loop through all the design matrices and get an idea of the CV performance of each.
Here are some additional models worth considering:
```python
models = {'l2logreg': LogisticRegressionCV(penalty='l2',cv=5,fit_intercept=True),
'lasso': LassoCV(cv=5,fit_intercept=True),
'xgb': xgb.XGBClassifier(max_depth=3, n_estimators=300, learning_rate=0.05),
'logreg': LogisticRegression(fit_intercept=True)}
```
```
#analyses = ['base', 'base_nodeathfix', '00', '04', '08','16',
# '24','fixed', 'wt8', 'wt16', 'wt24',
# 'wt8_00', 'wt8_08', 'wt8_16', 'wt8_24']
seeds = {'base': 473010,
'base_nodeathfix': 217632,
'00': 724311,
'04': 952227,
'08': 721297,
'16': 968879,
'24': 608972,
'fixed': 585794,
'wt8': 176381,
'wt16': 658229,
'wt24': 635170,
'wt8_00': 34741,
'wt8_08': 95467,
'wt8_16': 85349,
'wt8_24': 89642,
'wt24_fixed': 761456}
data_ext = 'base'
# SVM parameters tuned by cross-validation
#svm_parameters = {'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
# 'C': [1, 10]}
# use a full grid over all parameters
# specify parameters and distributions to sample from
N_FEAT = X.shape[1]
param_dist = {"max_depth": [3, 7, None],
"max_features": sp.stats.randint(1, N_FEAT),
"min_samples_split": sp.stats.randint(1, N_FEAT),
"min_samples_leaf": sp.stats.randint(1, N_FEAT),
"n_estimators": sp.stats.randint(50, 500),
"criterion": ["gini", "entropy"]}
# set up randomized search for RF
n_iter_search = 20
rf_random_search = RandomizedSearchCV(sklearn.ensemble.RandomForestClassifier(),
param_distributions=param_dist,
n_iter=n_iter_search)
models = {'xgb': xgb.XGBClassifier(max_depth=3, n_estimators=300, learning_rate=0.05),
'lasso': LassoCV(cv=5,fit_intercept=True),
'logreg': LogisticRegression(fit_intercept=True),
'rf': sklearn.ensemble.RandomForestClassifier(),
#'svm': GridSearchCV(sklearn.svm.SVC(kernel='rbf',class_weight='balanced',probability=False),
# svm_parameters, cv=5, scoring='roc_auc')
}
results = dict()
np.random.seed(seed=seeds[data_ext])
# load the data into a numpy array
X, y, X_header = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext='_' + data_ext)
print('{} - ========= {} ========='.format(dt.datetime.now(), data_ext))
scores = list()
for i, mdl in enumerate(models):
if mdl == 'xgb':
# no pre-processing of data necessary for xgb
estimator = Pipeline([(mdl, models[mdl])])
else:
estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
strategy="mean",
axis=0)),
("scaler", StandardScaler()),
(mdl, models[mdl])])
curr_score = cross_val_score(estimator, X, y, scoring='roc_auc',cv=5)
print('{} - {:10s} {:0.4f} [{:0.4f}, {:0.4f}]'.format(dt.datetime.now(), mdl,
np.mean(curr_score),
np.min(curr_score), np.max(curr_score)))
# save the score to a dictionary
results[mdl] = curr_score
#analyses = ['base', 'base_nodeathfix', '00', '04', '08','16',
# '24','fixed', 'wt8', 'wt16', 'wt24',
# 'wt8_00', 'wt8_08', 'wt8_16', 'wt8_24']
seeds = {'base': 473010,
'base_nodeathfix': 217632,
'00': 724311,
'04': 952227,
'08': 721297,
'16': 968879,
'24': 608972,
'fixed': 585794,
'wt8': 176381,
'wt16': 658229,
'wt24': 635170,
'wt8_00': 34741,
'wt8_08': 95467,
'wt8_16': 85349,
'wt8_24': 89642,
'wt24_fixed': 761456}
data_ext = 'wt24_fixed'
# SVM parameters tuned by cross-validation
#svm_parameters = {'kernel': ['rbf'], 'gamma': [1e-3, 1e-4],
# 'C': [1, 10]}
# use a full grid over all parameters
# specify parameters and distributions to sample from
N_FEAT = X.shape[1]
param_dist = {"max_depth": [3, 7, None],
"max_features": sp.stats.randint(1, N_FEAT),
"min_samples_split": sp.stats.randint(1, N_FEAT),
"min_samples_leaf": sp.stats.randint(1, N_FEAT),
"n_estimators": sp.stats.randint(50, 500),
"criterion": ["gini", "entropy"]}
# set up randomized search for RF
n_iter_search = 20
rf_random_search = RandomizedSearchCV(sklearn.ensemble.RandomForestClassifier(),
param_distributions=param_dist,
n_iter=n_iter_search)
models = {'xgb': xgb.XGBClassifier(max_depth=3, n_estimators=300, learning_rate=0.05),
'lasso': LassoCV(cv=5,fit_intercept=True),
'logreg': LogisticRegression(fit_intercept=True),
'rf': sklearn.ensemble.RandomForestClassifier(),
#'svm': GridSearchCV(sklearn.svm.SVC(kernel='rbf',class_weight='balanced',probability=False),
# svm_parameters, cv=5, scoring='roc_auc')
}
results = dict()
np.random.seed(seed=seeds[data_ext])
# load the data into a numpy array
X, y, X_header = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext='_' + data_ext)
print('{} - ========= {} ========='.format(dt.datetime.now(), data_ext))
scores = list()
for i, mdl in enumerate(models):
if mdl == 'xgb':
# no pre-processing of data necessary for xgb
estimator = Pipeline([(mdl, models[mdl])])
else:
estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
strategy="mean",
axis=0)),
("scaler", StandardScaler()),
(mdl, models[mdl])])
curr_score = cross_val_score(estimator, X, y, scoring='roc_auc',cv=5)
print('{} - {:10s} {:0.4f} [{:0.4f}, {:0.4f}]'.format(dt.datetime.now(), mdl,
np.mean(curr_score),
np.min(curr_score), np.max(curr_score)))
# save the score to a dictionary
results[mdl] = curr_score
# compare to severity of illness scores
df = co
# merge in the various severity scores
df = df.merge(oa, how='left', left_index=True,right_index=True,suffixes=('','_oasis'))
df = df.merge(sofa, how='left', left_index=True,right_index=True,suffixes=('','_sofa'))
df = df.merge(saps, how='left', left_index=True,right_index=True,suffixes=('','_saps'))
df = df.merge(sapsii, how='left', left_index=True,right_index=True,suffixes=('','_sapsii'))
df = df.merge(apsiii, how='left', left_index=True,right_index=True,suffixes=('','_apsiii'))
for v in df.columns:
if v != 'hospital_expire_flag':
print('{:8s} - {:0.4f}'.format(v,metrics.roc_auc_score(df['hospital_expire_flag'],df[v])))
# print the results
mdl = 'xgb'
print('=================== {} ==================='.format(mdl))
for data_ext in np.sort(results_val.keys()):
curr_score = results[mdl][data_ext]
print('{:15s} - {:0.4f} [{:0.4f} - {:0.4f}]'.format(data_ext, np.mean(curr_score), np.min(curr_score), np.max(curr_score)))
```
The above reported cross-validation performance in a variety of settings. We're also interested in *evaluating* the same model in the various settings. That is, training a model using random offsets, and then evaluating how it performs 4 hours before death, 8 hours, etc.
```
# extract the data used to train the model
data_ext = 'base'
np.random.seed(seed=seeds[data_ext])
# load the data into a numpy array
X, y, X_header = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext=data_ext)
# load into a dictionary the other various datasets/models
X_val = dict()
y_val = dict()
X_header_val = dict()
results_val = dict() # stores AUROCs across datasets
mdl_val = dict() # stores the model trained across k-folds
for i, data_ext in enumerate(analyses):
# load the data into a numpy array
X_val[data_ext], y_val[data_ext], X_header_val[data_ext] = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext=data_ext)
results_val[data_ext] = dict()
print('{} - Finished loading data'.format(dt.datetime.now()))
np.random.seed(seed=seeds[data_ext])
# create k-fold indices
K = 5 # number of folds
idxK = np.random.permutation(X.shape[0])
idxK = np.mod(idxK,K)
mdl = 'xgb'
mdl_val[mdl] = list()
for data_ext in X_val:
results_val[data_ext][mdl] = list() # initialize list for scores
# no pre-processing of data necessary for xgb
estimator = Pipeline([(mdl, models[mdl])])
for k in range(K):
# train the model using all but the kth fold
curr_mdl = estimator.fit(X[idxK != k, :],y[idxK != k])
for data_ext in X_val:
# get prediction on this dataset
curr_prob = curr_mdl.predict_proba(X_val[data_ext][idxK == k, :])
curr_prob = curr_prob[:,1]
# calculate score (AUROC)
curr_score = metrics.roc_auc_score(y_val[data_ext][idxK == k], curr_prob)
# add score to list of scores
results_val[data_ext][mdl].append(curr_score)
# save the current model
mdl_val[mdl].append(curr_mdl)
print('{} - Finished fold {} of {}.'.format(dt.datetime.now(), k+1, K))
# print the results
mdl = 'xgb'
print('=================== {} ==================='.format(mdl))
for data_ext in np.sort(results_val.keys()):
curr_score = results_val[data_ext][mdl]
print('{:15s} - {:0.4f} [{:0.4f} - {:0.4f}]'.format(data_ext, np.mean(curr_score), np.min(curr_score), np.max(curr_score)))
```
Repeat the same experiment as above, but this time, let's train a model with the outcome "did the patient die in the next 24 hours?"
```
# extract the data
np.random.seed(seed=seeds[data_ext])
data_ext = 'base'
# load the data into a numpy array
X, y, X_header = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext=data_ext,
diedWithin=24)
# load into a dictionary the other various datasets/models
X_val = dict()
y_val = dict()
X_header_val = dict()
results_val = dict() # stores AUROCs across datasets
mdl_val = dict() # stores the model trained across k-folds
for i, data_ext in enumerate(analyses):
# load the data into a numpy array
X_val[data_ext], y_val[data_ext], X_header_val[data_ext] = mp.load_design_matrix(co,
df_additional_data=df_static[vars_static],
data_ext='_' + data_ext)
results_val[data_ext] = dict()
print('{} - Finished loading data'.format(dt.datetime.now()))
np.random.seed(seed=seeds[data_ext])
# create k-fold indices
K = 5 # number of folds
idxK = np.random.permutation(X.shape[0])
idxK = np.mod(idxK,K)
mdl = 'xgb'
mdl_val[mdl] = list()
for data_ext in X_val:
results_val[data_ext][mdl] = list() # initialize list for scores
# no pre-processing of data necessary for xgb
estimator = Pipeline([(mdl, models[mdl])])
for k in range(K):
# train the model using all but the kth fold
curr_mdl = estimator.fit(X[idxK != k, :],y[idxK != k])
for data_ext in X_val:
# get prediction on this dataset
curr_prob = curr_mdl.predict_proba(X_val[data_ext][idxK == k, :])
curr_prob = curr_prob[:,1]
# calculate score (AUROC)
curr_score = metrics.roc_auc_score(y_val[data_ext][idxK == k], curr_prob)
# add score to list of scores
results_val[data_ext][mdl].append(curr_score)
# save the current model
mdl_val[mdl].append(curr_mdl)
print('{} - Finished fold {} of {}.'.format(dt.datetime.now(), k+1, K))
# print the results
mdl = 'xgb'
print('=================== {} ==================='.format(mdl))
for data_ext in np.sort(results_val.keys()):
curr_score = results_val[data_ext][mdl]
print('{:15s} - {:0.4f} [{:0.4f} - {:0.4f}]'.format(data_ext, np.mean(curr_score), np.min(curr_score), np.max(curr_score)))
```
Now we have an estimate of how well these models do in cross-validation. The next step will be to take the best model and optimize it appropriately using only a training subset of data.
```
# create training / test sets
np.random.seed(seed=324875)
icustay_id = co.index.values
idxTest = np.random.rand(X.shape[0]) > 0.20
X_train = X[~idxTest,:]
y_train = y[~idxTest]
iid_train = icustay_id[~idxTest]
X_test = X[idxTest,:]
y_test = y[idxTest]
iid_test = icustay_id[~idxTest]
# optimize hyperparameters of a model using only the training set
# takes ~20 minutes
# first train it w/o grid search
xgb_nopreproc = xgb.XGBClassifier(max_depth=3, n_estimators=300, learning_rate=0.05)
xgb_nopreproc = xgb_nopreproc.fit(X_train, y_train)
# parameters with multiple values will be used in the grid search
grid_params = {
'max_depth': [4,7], # max depth of the tree
'learning_rate': [0.05, 0.3], # step size shrinkage, makes earlier trees less important over time
'n_estimators': [300, 1000], # number of trees built
'subsample': [0.3, 0.8] # subsample the data when fitting each tree (prevent overfitting)
}
default_params = {'colsample_bytree': 1,
'colsample_bylevel':1,
'silent':1,
'reg_lambda':1, # L2 regularization on weights
'reg_alpha':0, # L1 regularization on weights
'objective':'binary:logistic'}
init_model = xgb.XGBClassifier(**default_params)
# the pipeline here is redundant - but could be useful if you want to add any custom preprocessing
# for example, creating binary features from categories, etc...
# the custom function only has to implement 'fit' and 'transform'
estimator = Pipeline([("xgb", GridSearchCV(init_model, grid_params, verbose=1))])
xgb_model_cv = estimator.fit(X_train,y_train)
# generate class probabilities
y_prob = xgb_model_cv.predict_proba(X_test)
y_prob = y_prob[:, 1]
# predict class labels for the test set
y_pred = (y_prob > 0.5).astype(int)
# get the original xgb predictions without cross-validation
# gives us a rough idea of the improvement of selecting some of the parameters
y_prob_nocv = xgb_nopreproc.predict_proba(X_test)[:,1]
print('\n --- Performance on 20% held out test set --- \n')
# generate evaluation metrics
print('Accuracy = {:0.3f}'.format(metrics.accuracy_score(y_test, y_pred)))
print('AUROC = {:0.3f} (unoptimized model was {:0.3f})'.format(metrics.roc_auc_score(y_test, y_prob),
metrics.roc_auc_score(y_test, y_prob_nocv)))
mp.print_cm(y_test, y_pred)
```
Given the above optimized hyperparameters, train the final model.
```
#best_params = xgb_model_cv.get_params()['xgb'].best_params_
xgb_model = xgb.XGBClassifier(**default_params)
#xgb_model = xgb_model.set_params(**best_params)
xgb_model = xgb_model.fit(X_train, y_train)
# feature importance!
plt.figure(figsize=[14,40])
ax = plt.gca()
mp.plot_xgb_importance_fmap(xgb_model, X_header=X_header, ax=ax)
plt.show()
```
SVM is just too slow :(
```python
# speed up SVM
estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
strategy="mean",
axis=0)),
("scaler", sklearn.preprocessing.MinMaxScaler()),
("svm", sklearn.svm.SVC(cache_size=6000))])
for n in [100,1000,10000]:
print(n)
%timeit estimator.fit(X[0:n,:],y[0:n])
```
~10,000 samples take ~5s and that's using default parameters.
```python
# speed up SVM with bagging
n_estimators = 10
estimator = Pipeline([("imputer", Imputer(missing_values='NaN',
strategy="mean",
axis=0)),
("scaler", sklearn.preprocessing.MinMaxScaler()),
("svm_bagged", BaggingClassifier(sklearn.svm.SVC(kernel='linear',
probability=False,
class_weight='balanced',
cache_size=6000),
max_samples = 1.0 / n_estimators,
n_estimators=n_estimators,
bootstrap=False))])
for n in [100,1000,10000]:
print(n)
%timeit estimator.fit(X[0:n,:],y[0:n])
```
| github_jupyter |
# Deploy Document Classification Custom Skill
This tutorial shows how to deploy a document classification custom skill for Cognitive Search. We will use the document classifier that was created by *01_Train_AML_Model.ipynb*. If you have not already, please run that script.
For more information on using custom skills with Cognitive Search, please see this [page](https://docs.microsoft.com/en-us/azure/search/cognitive-search-custom-skill-interface).
### 0.0 Important Variables you need to set for this tutorial
Enter your workspace, resource and subscription credentials below
```
# Machine Learning Service Workspace configuration
my_workspace_name = ''
my_azure_subscription_id = ''
my_resource_group = ''
# Azure Kubernetes Service configuration
my_aks_location = 'eastus'
my_aks_compute_target_name = 'aks-comptarget'
my_aks_service_name = 'aks-service'
my_leaf_domain_label = 'ssl1' # web service url prefix
```
### 1.0 Import Packages
```
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License.
import numpy as np
import azureml
from azureml.core import Workspace, Run
# display the core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
```
### 2.0 Connect to Workspace
Create a workspace object. If you already have a workspace and a config.json file you can use `ws = Workspace.from_config()` instead.
```
from azureml.core import Workspace
from azureml.core.model import Model
ws = Workspace.get(name = my_workspace_name, resource_group = my_resource_group, subscription_id = my_azure_subscription_id)
print(ws.name, ws.location, ws.resource_group, sep = '\t')
```
### 3.0 Register Model
The last step in the training script wrote the file outputs/sklearn_mnist_model.pkl in a directory named outputs.
Register the model in the workspace so that you (or other collaborators) can query, examine, and deploy this model.
```
model_name="newsgroup_classifier"
model = Model.register(model_path="outputs/newsgroup_classifier.pkl",
model_name=model_name,
tags={"data": "newsgroup", "document": "classification"},
description="document classifier for newsgroup20",
workspace=ws)
print(model.id)
```
### 4.0 Create Scoring Script
Create the scoring script, called score.py, used by the web service call to show how to use the model.
You must include two required functions into the scoring script:
- The init() function, which typically loads the model into a global object. This function is run only once when the Docker container is started.
- The run(input_data) function uses the model to predict a value based on the input data. Inputs and outputs to the run typically use JSON for serialization and de-serialization, but other formats are supported.
*The **run function** has been specifically tailored to deploy the model as a custom skill. This means that inputs & outputs are formatted correctly and any errors will be returned in a format usable by Cognitive Search*.
```
%%writefile score.py
import json
import numpy as np
import pandas as pd
import os
import pickle
import joblib
from azureml.core.model import Model
def init():
global model
# retreive the path to the model file using the model name
model_path = Model.get_model_path(model_name='newsgroup_classifier')
model = joblib.load(model_path)
def convert_to_df(my_dict):
df = pd.DataFrame(my_dict["values"])
data = df['data'].tolist()
index = df['recordId'].tolist()
return pd.DataFrame(data, index = index)
def run(raw_data):
data = json.loads(raw_data)
# Converting the input dictionary to a dataframe
try:
df = convert_to_df(data)
# Returning error message for each item in batch if data not in correct format
except:
df = pd.DataFrame(data)
index = df['recordId'].tolist()
message = "Request for batch is not in correct format"
output_list = [{'recordId': i, 'data': {}, "errors": [{'message': message}]} for i in index]
return {'values': output_list}
output_list = []
for index, row in df.iterrows():
output = {'recordId': index, 'data': {}}
try:
output['data']['type'] = str(model.predict([row['content']])[0])
# Returning exception if an error occurs
except Exception as ex:
output['errors'] = [{'message': str(ex)}]
output_list.append(output)
return {'values': output_list}
```
### 5.0 Create Environment and Inference Configuration
```
from azureml.core.conda_dependencies import CondaDependencies
from azureml.core import Environment
pip = ["azureml-defaults", "scikit-learn", "pandas", "joblib"]
conda_deps = CondaDependencies.create(conda_packages=None, pip_packages=pip)
myenv = Environment(name='myenv')
myenv.python.conda_dependencies = conda_deps
from azureml.core.model import InferenceConfig
inf_config = InferenceConfig(entry_script='score.py', environment=myenv)
```
### 6.0 Create Azure Kubernetes Service Configuration File
Estimated time to complete: about 10 minutes
Create an Azure Kubernetes Service deployment configuration file. Notice that we enable SSL since Azure Search only allows secure endpoints as custom skills.
```
# create AKS compute target
from azureml.core.compute import ComputeTarget, AksCompute
config = AksCompute.provisioning_configuration(location= my_aks_location)
config.enable_ssl(leaf_domain_label= my_leaf_domain_label, overwrite_existing_domain=True)
aks = ComputeTarget.create(ws, my_aks_compute_target_name, config)
aks.wait_for_completion(show_output=True)
# if you already created a configuration file, you can just attach:
#config = AksCompute.attach_configuration(resource_group= my_resource_group, cluster_name='enter cluser name here')
#config.enable_ssl(leaf_domain_label= my_leaf_domain_label, overwrite_existing_domain=True)
#aks = ComputeTarget.attach(ws, my_aks_compute_target_name, config)
#aks.wait_for_completion(show_output=True)
print(aks.ssl_configuration.cname, aks.ssl_configuration.status)
```
### 7.0 Define the Deployment Configuration
```
from azureml.core.webservice import AksWebservice, Webservice
# If deploying to a cluster configured for dev/test, ensure that it was created with enough
# cores and memory to handle this deployment configuration. Note that memory is also used by
# dependencies and AML components.
aks_config = AksWebservice.deploy_configuration(autoscale_enabled=True,
autoscale_min_replicas=1,
autoscale_max_replicas=3,
autoscale_refresh_seconds=10,
autoscale_target_utilization=70,
auth_enabled=True,
cpu_cores=1, memory_gb=2,
scoring_timeout_ms=5000,
replica_max_concurrent_requests=2,
max_request_wait_time=5000)
```
### 8.0 Deploy a web service
Deploy a web service using the AKS image. Then get the web service HTTPS endpoint and the key to use to call the service
```
from azureml.core.model import Model
document_classifier = Model(ws, model_name)
# deploy an AKS web service using the image
#aks_config = AksWebservice.deploy_configuration()
service = Model.deploy(workspace=ws,
name=my_aks_service_name,
models=[document_classifier],
inference_config=inf_config,
deployment_config=aks_config,
deployment_target=aks,
overwrite=True)
service.wait_for_deployment(show_output = True)
primary, secondary = service.get_keys()
print('Scoring Uri: ' + service.scoring_uri)
print('Primary key: ' + primary)
```
### 9.0 Test Deployed Service
#### 9.1 Import 20newsgroups Test Dataset
```
from sklearn.datasets import fetch_20newsgroups
categories = ['comp.graphics', 'sci.space']
newsgroups_test = fetch_20newsgroups(subset='test', categories=categories)
X_test = newsgroups_test.data
y_test = [categories[x] for x in newsgroups_test.target]
```
#### 9.2 Format Data in Correct Structure for Cognitive Search
For more information on custom skills see this [link](https://docs.microsoft.com/en-us/azure/search/cognitive-search-custom-skill-interface).
```
# send a random row from the test set to score
random_index = np.random.randint(0, len(X_test)-1)
input_data = {"values":[{"recordId": "0", "data": {"content": newsgroups_test.data[random_index]}}]}
print(input_data)
```
#### 9.3 Send HTTP Request and View Results
```
import requests
import json
input_json = json.dumps(input_data)
headers = { 'Content-Type':'application/json'}
headers['Authorization']= f'Bearer {primary}'
# for AKS deployment you'd need the service key in the header as well
# api_key = service.get_key()
# headers = {'Content-Type':'application/json', 'Authorization':('Bearer '+ api_key)}
resp = requests.post(service.scoring_uri, input_json, headers=headers)
print("POST to url", service.scoring_uri)
print("label:", y_test[random_index])
print("prediction:", resp.text)
```
## 10.0 Integrate the custom skill
```
print('Scoring Uri: ' + service.scoring_uri)
print('Primary key: ' + primary)
```
Nice work! You're now ready to add the custom skill to your skillset.
Add the following skill to your skillset:
```json
{
"@odata.type": "#Microsoft.Skills.Custom.WebApiSkill",
"description": "A document classification custom skill",
"uri": "<your-scoring-uri>",
"httpHeaders": {
"Authorization": "Bearer <your-primary-key>"
},
"batchSize": 1,
"context": "/document",
"inputs": [
{
"name": "content",
"source": "/document/content"
}
],
"outputs": [
{
"name": "type",
"targetName": "type"
}
]
}
```
Don't forget to also add an [output field mapping](https://docs.microsoft.com/azure/search/cognitive-search-output-field-mapping) to your indexer so that the data gets mapped into the search index correctly:
```json
{
"sourceFieldName": "/document/type",
"targetFieldName": "type"
}
```
| github_jupyter |
# Tema 14: Bases de datos con SQLite (Enunciados)
*Estos ejercicios son optativos para hacer al final de la unidad y están pensados para apoyar tu aprendizaje*.
## *NOTA IMPORTANTE*
Todos los ejercicios deberás realizarlos en **scripts** creados en el mismo directorio donde trabajarás con las bases de datos.
**1.1) A lo largo de estos ejercicios vamos a crear un pequeño sistema para gestionar los platos del menú de un restaurante. Por ahora debes empezar creando un script llamado restaurante.py y dentro una función crear_bd() que creará una pequeña base de datos restaurante.db con las siguientes dos tablas:**
** *Si ya existen deberá tratar la excepción y mostrar que las tablas ya existen. En caso contrario mostrará que se han creado correctamente.* **
```sql
CREATE TABLE categoria(
id INTEGER PRIMARY KEY AUTOINCREMENT,
nombre VARCHAR(100) UNIQUE NOT NULL)
```
```sql
CREATE TABLE plato(
id INTEGER PRIMARY KEY AUTOINCREMENT,
nombre VARCHAR(100) UNIQUE NOT NULL,
categoria_id INTEGER NOT NULL,
FOREIGN KEY(categoria_id) REFERENCES categoria(id))
```
**Nota:** *La línea **FOREIGN KEY(categoria_id) REFERENCES categoria(id)** indica un tipo de clave especial (foránea), por la cual se crea una relación entre la categoría de un plato con el registro de categorías.*
** Llama a la función y comprueba que la base de datos se crea correctamente.**
**1.2) Crea una función llamada agregar_categoria() que pida al usuario un nombre de categoría y se encargue de crear la categoría en la base de datos (ten en cuenta que si ya existe dará un error, por que el nombre es UNIQUE).**
** Ahora, crea un pequeño menú de opciones dentro del script, que te de la bienvenida al sistema y te permita Crear una categoría o Salir. Añade las siguientes tres categorías utilizando este menú de opciones:**
- Primeros
- Segundos
- Postres
**1.3) Crea una función llamada agregar_plato() que muestre al usuario las categorías disponibles y le permita escoger una (escribiendo un número).**
**Luego le pedirá introducir el nombre del plato y lo añadirá a la base de datos, teniendo en cuenta que la categoria del plato concuerde con el id de la categoría y que el nombre del plato no puede repetirse (no es necesario comprobar si la categoría realmente existe, en ese caso simplemente no se insertará el plato).**
**Agrega la nueva opción al menú de opciones y añade los siguientes platos:**
- **Primeros**: Ensalada del tiempo / Zumo de tomate
- **Segundos**: Estofado de pescado / Pollo con patatas
- **Postres**: Flan con nata / Fruta del tiempo
**1.4) Crea una función llamada mostrar_menu() que muestre el menú con todos los platos de forma ordenada: los primeros, los segundos y los postres. Optativamente puedes adornar la forma en que muestras el menú por pantalla.**
**2) En este ejercicios debes crear una interfaz gráfica con tkinter (menu.py) que muestre de forma elegante el menú del restaurante.**
- Tú eliges el nombre del restaurante y el precio del menú, así como las tipografías, colores, adornos y tamaño de la ventana.
- El único requisito es que el programa se conectará a la base de datos para buscar la lista categorías y platos.
- Algunas ideas: https://www.google.es/search?tbm=isch&q=dise%C3%B1o+menu+restaurantes
### Ejemplo

| github_jupyter |
# ArcGIS and IBM Watson
## Machine Learning Integration for Pedestrian Activity Classification

# 1. Retrieve Training Data from Survey123
# 2. Retrieve Models from IBM Watson
```
import os
from watson_developer_cloud import VisualRecognitionV3
import json
from IPython.display import Image
from IPython.core.display import HTML
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
visual_recognition = VisualRecognitionV3(
'2018-03-19',
iam_api_key=os.environ['BLUEMIX_API_KEY'])
visual_recognition.list_classifiers(verbose=True)
classifier_name = 'Esri_Classification_CobbCounty_Cam128_v2'
classifier_id = None
for classifier in visual_recognition.list_classifiers()['classifiers']:
if classifier['name'] == classifier_name:
classifier_id = classifier['classifier_id']
print(classifier_id)
test_images_dir = "{0}\\TestData\\128".format(os.getcwd())
high_dir = "{0}\\{1}".format(test_images_dir, "High")
med_dir = "{0}\\{1}".format(test_images_dir, "Medium")
low_dir = "{0}\\{1}".format(test_images_dir, "Low")
high_img_path = "{0}\\{1}".format(high_dir, os.listdir(high_dir)[0])
med_img_path = "{0}\\{1}".format(med_dir, os.listdir(med_dir)[0])
low_img_path = "{0}\\{1}".format(low_dir, os.listdir(low_dir)[0])
```
# 3. Test Classification on Sample Images
## Low Pedestrian Density Sample
```
Image(filename= low_img_path)
with open(low_img_path, 'rb') as images_file:
classes = visual_recognition.classify(
images_file,
threshold='0.0',
classifier_ids=classifier_id)
# print(json.dumps(classes, indent=2))
output_classes = classes['images'][0]['classifiers'][0]['classes']
df = pd.DataFrame(data=output_classes)
df = df.set_index('class')
df.plot(kind='barh')
```
## Medium Pedestrian Density Sample
```
Image(filename= med_img_path)
with open(med_img_path, 'rb') as images_file:
classes = visual_recognition.classify(
images_file,
threshold='0.0',
classifier_ids=classifier_id)
# print(json.dumps(classes, indent=2))
output_classes = classes['images'][0]['classifiers'][0]['classes']
df = pd.DataFrame(data=output_classes)
df = df.set_index('class')
df.plot(kind='barh')
```
## High Pedestrian Density Sample
```
Image(filename= high_img_path)
with open(high_img_path, 'rb') as images_file:
classes = visual_recognition.classify(
images_file,
threshold='0.0',
classifier_ids=classifier_id)
# print(json.dumps(classes, indent=2))
output_classes = classes['images'][0]['classifiers'][0]['classes']
df = pd.DataFrame(data=output_classes)
df = df.set_index('class')
df.plot(kind='barh')
```
| github_jupyter |
# Custom Modeling
Script to test the custom modeling .py scripts
## Imports
```
# imports
import os
import time
import datetime
import json
import gc
from numba import jit
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from tqdm import tqdm_notebook
# Preprocessing
from sklearn.preprocessing import MinMaxScaler,StandardScaler,Imputer,LabelEncoder,PolynomialFeatures
import lightgbm as lgb
import xgboost as xgb
#from catboost import CatBoostRegressor, CatBoostClassifier
from sklearn import metrics
# Model selection
from sklearn.model_selection import train_test_split,cross_validate
from sklearn.model_selection import KFold
from sklearn.model_selection import GridSearchCV
```
### Metric Definition
```
def group_mean_log_mae(y_true, y_pred, types, floor=1e-9):
"""
Fast metric computation for this competition: https://www.kaggle.com/c/champs-scalar-coupling
Code is from this kernel: https://www.kaggle.com/uberkinder/efficient-metric
"""
maes = (y_true-y_pred).abs().groupby(types).mean()
return np.log(maes.map(lambda x: max(x, floor))).mean()
```
### Modeling Custom Function
```
def train_model_regression(X, X_test, y, params, folds, model_type='lgb', eval_metric='mae', columns=None, plot_feature_importance=False, model=None,
verbose=10000, early_stopping_rounds=200, n_estimators=50000):
"""
A function to train a variety of regression models.
Returns dictionary with oof predictions, test predictions, scores and, if necessary, feature importances.
:params: X - training data, can be pd.DataFrame or np.ndarray (after normalizing)
:params: X_test - test data, can be pd.DataFrame or np.ndarray (after normalizing)
:params: y - target
:params: folds - folds to split data
:params: model_type - type of model to use
:params: eval_metric - metric to use
:params: columns - columns to use. If None - use all columns
:params: plot_feature_importance - whether to plot feature importance of LGB
:params: model - sklearn model, works only for "sklearn" model type
"""
columns = X.columns if columns is None else columns
X_test = X_test[columns]
# to set up scoring parameters
metrics_dict = {'mae': {'lgb_metric_name': 'mae',
'catboost_metric_name': 'MAE',
'sklearn_scoring_function': metrics.mean_absolute_error},
'group_mae': {'lgb_metric_name': 'mae',
'catboost_metric_name': 'MAE',
'scoring_function': group_mean_log_mae},
'mse': {'lgb_metric_name': 'mse',
'catboost_metric_name': 'MSE',
'sklearn_scoring_function': metrics.mean_squared_error}
}
result_dict = {}
# out-of-fold predictions on train data
oof = np.zeros(len(X))
# averaged predictions on train data
prediction = np.zeros(len(X_test))
# list of scores on folds
scores = []
feature_importance = pd.DataFrame()
# split and train on folds
for fold_n, (train_index, valid_index) in enumerate(folds.split(X)):
print(f'Fold {fold_n + 1} started at {time.ctime()}')
if type(X) == np.ndarray:
X_train, X_valid = X[columns][train_index], X[columns][valid_index]
y_train, y_valid = y[train_index], y[valid_index]
else:
X_train, X_valid = X[columns].iloc[train_index], X[columns].iloc[valid_index]
y_train, y_valid = y.iloc[train_index], y.iloc[valid_index]
if model_type == 'lgb':
model = lgb.LGBMRegressor(**params, n_estimators = n_estimators, n_jobs = -1)
model.fit(X_train, y_train,
eval_set=[(X_train, y_train), (X_valid, y_valid)], eval_metric=metrics_dict[eval_metric]['lgb_metric_name'],
verbose=verbose, early_stopping_rounds=early_stopping_rounds)
y_pred_valid = model.predict(X_valid)
y_pred = model.predict(X_test, num_iteration=model.best_iteration_)
if model_type == 'xgb':
train_data = xgb.DMatrix(data=X_train, label=y_train, feature_names=X.columns)
valid_data = xgb.DMatrix(data=X_valid, label=y_valid, feature_names=X.columns)
watchlist = [(train_data, 'train'), (valid_data, 'valid_data')]
model = xgb.train(dtrain=train_data, num_boost_round=400, evals=watchlist, early_stopping_rounds=200, verbose_eval=verbose, params=params)
y_pred_valid = model.predict(xgb.DMatrix(X_valid, feature_names=X.columns), ntree_limit=model.best_ntree_limit)
y_pred = model.predict(xgb.DMatrix(X_test, feature_names=X.columns), ntree_limit=model.best_ntree_limit)
if model_type == 'sklearn':
model = model
model.fit(X_train, y_train)
y_pred_valid = model.predict(X_valid).reshape(-1,)
score = metrics_dict[eval_metric]['sklearn_scoring_function'](y_valid, y_pred_valid)
print(f'Fold {fold_n}. {eval_metric}: {score:.4f}.')
print('')
y_pred = model.predict(X_test).reshape(-1,)
if model_type == 'cat':
model = CatBoostRegressor(iterations=20000, eval_metric=metrics_dict[eval_metric]['catboost_metric_name'], **params,
loss_function=metrics_dict[eval_metric]['catboost_metric_name'])
model.fit(X_train, y_train, eval_set=(X_valid, y_valid), cat_features=[], use_best_model=True, verbose=False)
y_pred_valid = model.predict(X_valid)
y_pred = model.predict(X_test)
oof[valid_index] = y_pred_valid.reshape(-1,)
if eval_metric != 'group_mae':
scores.append(metrics_dict[eval_metric]['sklearn_scoring_function'](y_valid, y_pred_valid))
else:
scores.append(metrics_dict[eval_metric]['scoring_function'](y_valid, y_pred_valid, X_valid['type']))
prediction += y_pred
if model_type == 'lgb' and plot_feature_importance:
# feature importance
fold_importance = pd.DataFrame()
fold_importance["feature"] = columns
fold_importance["importance"] = model.feature_importances_
fold_importance["fold"] = fold_n + 1
feature_importance = pd.concat([feature_importance, fold_importance], axis=0)
prediction /= folds.n_splits
print('CV mean score: {0:.4f}, std: {1:.4f}.'.format(np.mean(scores), np.std(scores)))
result_dict['oof'] = oof
result_dict['prediction'] = prediction
result_dict['scores'] = scores
if model_type == 'lgb':
if plot_feature_importance:
feature_importance["importance"] /= folds.n_splits
cols = feature_importance[["feature", "importance"]].groupby("feature").mean().sort_values(
by="importance", ascending=False)[:50].index
best_features = feature_importance.loc[feature_importance.feature.isin(cols)]
plt.figure(figsize=(16, 12));
sns.barplot(x="importance", y="feature", data=best_features.sort_values(by="importance", ascending=False));
plt.title('LGB Features (avg over folds)');
result_dict['feature_importance'] = feature_importance
return result_dict
def reduce_mem_usage(df, verbose=True):
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage().sum() / 1024**2
for col in df.columns:
col_type = df[col].dtypes
if col_type in numerics:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024**2
if verbose: print('Mem. usage decreased to {:5.2f} Mb ({:.1f}% reduction)'.format(end_mem, 100 * (start_mem - end_mem) / start_mem))
return df
```
## Data Load
```
# Train Data
train = pd.read_csv('input/train.csv')
# Test Data
test = pd.read_csv('input/test.csv')
# Submission Data
sub = pd.read_csv('input/sample_submission.csv')
#General Data
structures = pd.read_csv('input/structures.csv')
len(test['type'])
len(sub['id'])
```
## EDA
```
# We take a first look at the dataset
train.info()
print ('#################################################')
print ('#################################################')
test.info()
```
## Feature Engineering
##### Chemical Bond Calculation
```
from tqdm import tqdm_notebook as tqdm
atomic_radius = {'H':0.38, 'C':0.77, 'N':0.75, 'O':0.73, 'F':0.71} # Without fudge factor
fudge_factor = 0.05
atomic_radius = {k:v + fudge_factor for k,v in atomic_radius.items()}
print(atomic_radius)
electronegativity = {'H':2.2, 'C':2.55, 'N':3.04, 'O':3.44, 'F':3.98}
#structures = pd.read_csv(structures, dtype={'atom_index':np.int8})
atoms = structures['atom'].values
atoms_en = [electronegativity[x] for x in tqdm(atoms)]
atoms_rad = [atomic_radius[x] for x in tqdm(atoms)]
structures['EN'] = atoms_en
structures['rad'] = atoms_rad
display(structures.head())
i_atom = structures['atom_index'].values
p = structures[['x', 'y', 'z']].values
p_compare = p
m = structures['molecule_name'].values
m_compare = m
r = structures['rad'].values
r_compare = r
source_row = np.arange(len(structures))
max_atoms = 28
bonds = np.zeros((len(structures)+1, max_atoms+1), dtype=np.int8)
bond_dists = np.zeros((len(structures)+1, max_atoms+1), dtype=np.float32)
print('Calculating bonds')
for i in tqdm(range(max_atoms-1)):
p_compare = np.roll(p_compare, -1, axis=0)
m_compare = np.roll(m_compare, -1, axis=0)
r_compare = np.roll(r_compare, -1, axis=0)
mask = np.where(m == m_compare, 1, 0) #Are we still comparing atoms in the same molecule?
dists = np.linalg.norm(p - p_compare, axis=1) * mask
r_bond = r + r_compare
bond = np.where(np.logical_and(dists > 0.0001, dists < r_bond), 1, 0)
source_row = source_row
target_row = source_row + i + 1 #Note: Will be out of bounds of bonds array for some values of i
target_row = np.where(np.logical_or(target_row > len(structures), mask==0), len(structures), target_row) #If invalid target, write to dummy row
source_atom = i_atom
target_atom = i_atom + i + 1 #Note: Will be out of bounds of bonds array for some values of i
target_atom = np.where(np.logical_or(target_atom > max_atoms, mask==0), max_atoms, target_atom) #If invalid target, write to dummy col
bonds[(source_row, target_atom)] = bond
bonds[(target_row, source_atom)] = bond
bond_dists[(source_row, target_atom)] = dists
bond_dists[(target_row, source_atom)] = dists
bonds = np.delete(bonds, axis=0, obj=-1) #Delete dummy row
bonds = np.delete(bonds, axis=1, obj=-1) #Delete dummy col
bond_dists = np.delete(bond_dists, axis=0, obj=-1) #Delete dummy row
bond_dists = np.delete(bond_dists, axis=1, obj=-1) #Delete dummy col
print('Counting and condensing bonds')
bonds_numeric = [[i for i,x in enumerate(row) if x] for row in tqdm(bonds)]
bond_lengths = [[dist for i,dist in enumerate(row) if i in bonds_numeric[j]] for j,row in enumerate(tqdm(bond_dists))]
bond_lengths_mean = [ np.mean(x) for x in bond_lengths]
bond_lengths_std = [ np.std(x) for x in bond_lengths]
n_bonds = [len(x) for x in bonds_numeric]
#bond_data = {'bond_' + str(i):col for i, col in enumerate(np.transpose(bonds))}
#bond_data.update({'bonds_numeric':bonds_numeric, 'n_bonds':n_bonds})
bond_data = {'n_bonds':n_bonds, 'bond_lengths_mean': bond_lengths_mean,'bond_lengths_std':bond_lengths_std }
bond_df = pd.DataFrame(bond_data)
structures = structures.join(bond_df)
display(structures.head(20))
def map_atom_info(df, atom_idx):
df = pd.merge(df, structures, how = 'left',
left_on = ['molecule_name', f'atom_index_{atom_idx}'],
right_on = ['molecule_name', 'atom_index'])
df = df.drop('atom_index', axis=1)
df = df.rename(columns={'atom': f'atom_{atom_idx}',
'x': f'x_{atom_idx}',
'y': f'y_{atom_idx}',
'z': f'z_{atom_idx}'})
df = reduce_mem_usage(df)
return df
train = map_atom_info(train, 0)
train = map_atom_info(train, 1)
test = map_atom_info(test, 0)
test = map_atom_info(test, 1)
train_p_0 = train[['x_0', 'y_0', 'z_0']].values
train_p_1 = train[['x_1', 'y_1', 'z_1']].values
test_p_0 = test[['x_0', 'y_0', 'z_0']].values
test_p_1 = test[['x_1', 'y_1', 'z_1']].values
train['dist'] = np.linalg.norm(train_p_0 - train_p_1, axis=1)
test['dist'] = np.linalg.norm(test_p_0 - test_p_1, axis=1)
train['dist_x'] = (train['x_0'] - train['x_1']) ** 2
test['dist_x'] = (test['x_0'] - test['x_1']) ** 2
train['dist_y'] = (train['y_0'] - train['y_1']) ** 2
test['dist_y'] = (test['y_0'] - test['y_1']) ** 2
train['dist_z'] = (train['z_0'] - train['z_1']) ** 2
test['dist_z'] = (test['z_0'] - test['z_1']) ** 2
'''
This will create 2 features:
1) will show the first letter of `type`
2) Will show the rest of characters
'''
train['type_0'] = train['type'].apply(lambda x: x[0])
test['type_0'] = test['type'].apply(lambda x: x[0])
train['type_1'] = train['type'].apply(lambda x: x[1:])
test['type_1'] = test['type'].apply(lambda x: x[1:])
train['dist_to_type_mean'] = train['dist'] / train.groupby('type')['dist'].transform('mean')
test['dist_to_type_mean'] = test['dist'] / test.groupby('type')['dist'].transform('mean')
train['dist_to_type_0_mean'] = train['dist'] / train.groupby('type_0')['dist'].transform('mean')
test['dist_to_type_0_mean'] = test['dist'] / test.groupby('type_0')['dist'].transform('mean')
train['dist_to_type_1_mean'] = train['dist'] / train.groupby('type_1')['dist'].transform('mean')
test['dist_to_type_1_mean'] = test['dist'] / test.groupby('type_1')['dist'].transform('mean')
train[f'molecule_type_dist_mean'] = train.groupby(['molecule_name', 'type'])['dist'].transform('mean')
test[f'molecule_type_dist_mean'] = test.groupby(['molecule_name', 'type'])['dist'].transform('mean')
```
#### More Features
```
def create_features(df):
df['molecule_couples'] = df.groupby('molecule_name')['id'].transform('count')
df['molecule_dist_mean'] = df.groupby('molecule_name')['dist'].transform('mean')
df['molecule_dist_min'] = df.groupby('molecule_name')['dist'].transform('min')
df['molecule_dist_max'] = df.groupby('molecule_name')['dist'].transform('max')
df['atom_0_couples_count'] = df.groupby(['molecule_name', 'atom_index_0'])['id'].transform('count')
df['atom_1_couples_count'] = df.groupby(['molecule_name', 'atom_index_1'])['id'].transform('count')
df[f'molecule_atom_index_0_x_1_std'] = df.groupby(['molecule_name', 'atom_index_0'])['x_1'].transform('std')
df[f'molecule_atom_index_0_y_1_mean'] = df.groupby(['molecule_name', 'atom_index_0'])['y_1'].transform('mean')
df[f'molecule_atom_index_0_y_1_mean_diff'] = df[f'molecule_atom_index_0_y_1_mean'] - df['y_1']
df[f'molecule_atom_index_0_y_1_mean_div'] = df[f'molecule_atom_index_0_y_1_mean'] / df['y_1']
df[f'molecule_atom_index_0_y_1_max'] = df.groupby(['molecule_name', 'atom_index_0'])['y_1'].transform('max')
df[f'molecule_atom_index_0_y_1_max_diff'] = df[f'molecule_atom_index_0_y_1_max'] - df['y_1']
df[f'molecule_atom_index_0_y_1_std'] = df.groupby(['molecule_name', 'atom_index_0'])['y_1'].transform('std')
df[f'molecule_atom_index_0_z_1_std'] = df.groupby(['molecule_name', 'atom_index_0'])['z_1'].transform('std')
df[f'molecule_atom_index_0_dist_mean'] = df.groupby(['molecule_name', 'atom_index_0'])['dist'].transform('mean')
df[f'molecule_atom_index_0_dist_mean_diff'] = df[f'molecule_atom_index_0_dist_mean'] - df['dist']
df[f'molecule_atom_index_0_dist_mean_div'] = df[f'molecule_atom_index_0_dist_mean'] / df['dist']
df[f'molecule_atom_index_0_dist_max'] = df.groupby(['molecule_name', 'atom_index_0'])['dist'].transform('max')
df[f'molecule_atom_index_0_dist_max_diff'] = df[f'molecule_atom_index_0_dist_max'] - df['dist']
df[f'molecule_atom_index_0_dist_max_div'] = df[f'molecule_atom_index_0_dist_max'] / df['dist']
df[f'molecule_atom_index_0_dist_min'] = df.groupby(['molecule_name', 'atom_index_0'])['dist'].transform('min')
df[f'molecule_atom_index_0_dist_min_diff'] = df[f'molecule_atom_index_0_dist_min'] - df['dist']
df[f'molecule_atom_index_0_dist_min_div'] = df[f'molecule_atom_index_0_dist_min'] / df['dist']
df[f'molecule_atom_index_0_dist_std'] = df.groupby(['molecule_name', 'atom_index_0'])['dist'].transform('std')
df[f'molecule_atom_index_0_dist_std_diff'] = df[f'molecule_atom_index_0_dist_std'] - df['dist']
df[f'molecule_atom_index_0_dist_std_div'] = df[f'molecule_atom_index_0_dist_std'] / df['dist']
df[f'molecule_atom_index_1_dist_mean'] = df.groupby(['molecule_name', 'atom_index_1'])['dist'].transform('mean')
df[f'molecule_atom_index_1_dist_mean_diff'] = df[f'molecule_atom_index_1_dist_mean'] - df['dist']
df[f'molecule_atom_index_1_dist_mean_div'] = df[f'molecule_atom_index_1_dist_mean'] / df['dist']
df[f'molecule_atom_index_1_dist_max'] = df.groupby(['molecule_name', 'atom_index_1'])['dist'].transform('max')
df[f'molecule_atom_index_1_dist_max_diff'] = df[f'molecule_atom_index_1_dist_max'] - df['dist']
df[f'molecule_atom_index_1_dist_max_div'] = df[f'molecule_atom_index_1_dist_max'] / df['dist']
df[f'molecule_atom_index_1_dist_min'] = df.groupby(['molecule_name', 'atom_index_1'])['dist'].transform('min')
df[f'molecule_atom_index_1_dist_min_diff'] = df[f'molecule_atom_index_1_dist_min'] - df['dist']
df[f'molecule_atom_index_1_dist_min_div'] = df[f'molecule_atom_index_1_dist_min'] / df['dist']
df[f'molecule_atom_index_1_dist_std'] = df.groupby(['molecule_name', 'atom_index_1'])['dist'].transform('std')
df[f'molecule_atom_index_1_dist_std_diff'] = df[f'molecule_atom_index_1_dist_std'] - df['dist']
df[f'molecule_atom_index_1_dist_std_div'] = df[f'molecule_atom_index_1_dist_std'] / df['dist']
df[f'molecule_atom_1_dist_mean'] = df.groupby(['molecule_name', 'atom_1'])['dist'].transform('mean')
df[f'molecule_atom_1_dist_min'] = df.groupby(['molecule_name', 'atom_1'])['dist'].transform('min')
df[f'molecule_atom_1_dist_min_diff'] = df[f'molecule_atom_1_dist_min'] - df['dist']
df[f'molecule_atom_1_dist_min_div'] = df[f'molecule_atom_1_dist_min'] / df['dist']
df[f'molecule_atom_1_dist_std'] = df.groupby(['molecule_name', 'atom_1'])['dist'].transform('std')
df[f'molecule_atom_1_dist_std_diff'] = df[f'molecule_atom_1_dist_std'] - df['dist']
df[f'molecule_type_0_dist_std'] = df.groupby(['molecule_name', 'type_0'])['dist'].transform('std')
df[f'molecule_type_0_dist_std_diff'] = df[f'molecule_type_0_dist_std'] - df['dist']
df[f'molecule_type_dist_mean'] = df.groupby(['molecule_name', 'type'])['dist'].transform('mean')
df[f'molecule_type_dist_mean_diff'] = df[f'molecule_type_dist_mean'] - df['dist']
df[f'molecule_type_dist_mean_div'] = df[f'molecule_type_dist_mean'] / df['dist']
df[f'molecule_type_dist_max'] = df.groupby(['molecule_name', 'type'])['dist'].transform('max')
df[f'molecule_type_dist_min'] = df.groupby(['molecule_name', 'type'])['dist'].transform('min')
df[f'molecule_type_dist_std'] = df.groupby(['molecule_name', 'type'])['dist'].transform('std')
df[f'molecule_type_dist_std_diff'] = df[f'molecule_type_dist_std'] - df['dist']
df = reduce_mem_usage(df)
return df
def map_atom_info_2df(df_1,df_2, atom_idx):
df = pd.merge(df_1, df_2, how = 'left',
left_on = ['molecule_name', f'atom_index_{atom_idx}'],
right_on = ['molecule_name', 'atom_index'])
df = df.drop('atom_index', axis=1)
return df
def create_closest(df):
df_temp = df.loc[:,["molecule_name","atom_index_0","atom_index_1","dist","x_0","y_0","z_0","x_1","y_1","z_1"]].copy()
df_temp_ = df_temp.copy()
df_temp_ = df_temp_.rename(columns={'atom_index_0': 'atom_index_1',
'atom_index_1': 'atom_index_0',
'x_0': 'x_1',
'y_0': 'y_1',
'z_0': 'z_1',
'x_1': 'x_0',
'y_1': 'y_0',
'z_1': 'z_0'})
df_temp = pd.concat(objs=[df_temp,df_temp_],axis=0)
df_temp["min_distance"] = df_temp.groupby(['molecule_name', 'atom_index_0'])['dist'].transform('min')
df_temp=df_temp.drop_duplicates(subset=['molecule_name', 'atom_index_0'])
df_temp = df_temp[df_temp["min_distance"]==df_temp["dist"]]
df_temp = df_temp.drop(['x_0','y_0','z_0','min_distance'], axis=1)
df_temp = df_temp.rename(columns={'atom_index_0': 'atom_index',
'atom_index_1': 'atom_index_closest',
'distance': 'distance_closest',
'x_1': 'x_closest',
'y_1': 'y_closest',
'z_1': 'z_closest'})
for atom_idx in [0,1]:
df = map_atom_info_2df(df,df_temp, atom_idx)
df = df.rename(columns={'atom_index_closest': f'atom_index_closest_{atom_idx}',
'distance_closest': f'distance_closest_{atom_idx}',
'x_closest': f'x_closest_{atom_idx}',
'y_closest': f'y_closest_{atom_idx}',
'z_closest': f'z_closest_{atom_idx}'})
return df
```
#### Cosine Angles
```
def cosine_features(df):
df["distance_0"]=((df['x_0']-df['x_closest_0'])**2+(df['y_0']-df['y_closest_0'])**2+(df['z_0']-df['z_closest_0'])**2)**(1/2)
df["distance_1"]=((df['x_1']-df['x_closest_1'])**2+(df['y_1']-df['y_closest_1'])**2+(df['z_1']-df['z_closest_1'])**2)**(1/2)
df["vec_0_x"]=(df['x_0']-df['x_closest_0'])/df["distance_0"]
df["vec_0_y"]=(df['y_0']-df['y_closest_0'])/df["distance_0"]
df["vec_0_z"]=(df['z_0']-df['z_closest_0'])/df["distance_0"]
df["vec_1_x"]=(df['x_1']-df['x_closest_1'])/df["distance_1"]
df["vec_1_y"]=(df['y_1']-df['y_closest_1'])/df["distance_1"]
df["vec_1_z"]=(df['z_1']-df['z_closest_1'])/df["distance_1"]
df["vec_x"]=(df['x_1']-df['x_0'])/df["dist"]
df["vec_y"]=(df['y_1']-df['y_0'])/df["dist"]
df["vec_z"]=(df['z_1']-df['z_0'])/df["dist"]
df["cos_0_1"]=df["vec_0_x"]*df["vec_1_x"]+df["vec_0_y"]*df["vec_1_y"]+df["vec_0_z"]*df["vec_1_z"]
df["cos_0"]=df["vec_0_x"]*df["vec_x"]+df["vec_0_y"]*df["vec_y"]+df["vec_0_z"]*df["vec_z"]
df["cos_1"]=df["vec_1_x"]*df["vec_x"]+df["vec_1_y"]*df["vec_y"]+df["vec_1_z"]*df["vec_z"]
df=df.drop(['vec_0_x','vec_0_y','vec_0_z','vec_1_x','vec_1_y','vec_1_z','vec_x','vec_y','vec_z'], axis=1)
return df
```
#### Bonds Calculation
```
# NOT USED
def bonds_compute(df):
atom_rad = [self.atomic_radius[x] for x in df['atom'].values]
df['rad'] = atom_rad
position = df[['x','y','z']].values
p_temp = position
molec_name = df['molecule_name'].values
m_temp = molec_name
radius = df['rad'].values
r_temp = radius
bond = 0
dist_keep = 0
dist_bond = 0
no_bond = 0
dist_no_bond = 0
dist_matrix = np.zeros((df.shape[0],2*29))
dist_matrix_bond = np.zeros((df.shape[0],2*29))
dist_matrix_no_bond = np.zeros((df.shape[0],2*29))
for i in range(29):
p_temp = np.roll(p_temp,-1,axis=0)
m_temp = np.roll(m_temp,-1,axis=0)
r_temp = np.roll(r_temp,-1,axis=0)
mask = (m_temp==molec_name)
dist = np.linalg.norm(position-p_temp,axis=1) * mask
dist_temp = np.roll(np.linalg.norm(position-p_temp,axis=1)*mask,i+1,axis=0)
diff_radius_dist = (dist-(radius+r_temp)) * (dist<(radius+r_temp)) * mask
diff_radius_dist_temp = np.roll(diff_radius_dist,i+1,axis=0)
bond += (dist<(radius+r_temp)) * mask
bond_temp = np.roll((dist<(radius+r_temp)) * mask,i+1,axis=0)
no_bond += (dist>=(radius+r_temp)) * mask
no_bond_temp = np.roll((dist>=(radius+r_temp)) * mask,i+1,axis=0)
bond += bond_temp
no_bond += no_bond_temp
dist_keep += dist * mask
dist_matrix[:,2*i] = dist
dist_matrix[:,2*i+1] = dist_temp
dist_matrix_bond[:,2*i] = dist * (dist<(radius+r_temp)) * mask
dist_matrix_bond[:,2*i+1] = dist_temp * bond_temp
dist_matrix_no_bond[:,2*i] = dist * (dist>(radius+r_temp)) * mask
dist_matrix_no_bond[:,2*i+1] = dist_temp * no_bond_temp
df['n_bonds'] = bond
df['n_no_bonds'] = no_bond
df['dist_mean'] = np.nanmean(np.where(dist_matrix==0,np.nan,dist_matrix), axis=1)
df['dist_median'] = np.nanmedian(np.where(dist_matrix==0,np.nan,dist_matrix), axis=1)
df['dist_std_bond'] = np.nanstd(np.where(dist_matrix_bond==0,np.nan,dist_matrix), axis=1)
df['dist_mean_bond'] = np.nanmean(np.where(dist_matrix_bond==0,np.nan,dist_matrix), axis=1)
df['dist_median_bond'] = np.nanmedian(np.where(dist_matrix_bond==0,np.nan,dist_matrix), axis=1)
df['dist_mean_no_bond'] = np.nanmean(np.where(dist_matrix_no_bond==0,np.nan,dist_matrix), axis=1)
df['dist_std_no_bond'] = np.nanstd(np.where(dist_matrix_no_bond==0,np.nan,dist_matrix), axis=1)
df['dist_median_no_bond'] = np.nanmedian(np.where(dist_matrix_no_bond==0,np.nan,dist_matrix), axis=1)
df['dist_std'] = np.nanstd(np.where(dist_matrix==0,np.nan,dist_matrix), axis=1)
df['dist_min'] = np.nanmin(np.where(dist_matrix==0,np.nan,dist_matrix), axis=1)
df['dist_max'] = np.nanmax(np.where(dist_matrix==0,np.nan,dist_matrix), axis=1)
df['range_dist'] = np.absolute(X['dist_max']-df['dist_min'])
df['dist_bond_min'] = np.nanmin(np.where(dist_matrix_bond==0,np.nan,dist_matrix), axis=1)
df['dist_bond_max'] = np.nanmax(np.where(dist_matrix_bond==0,np.nan,dist_matrix), axis=1)
df['range_dist_bond'] = np.absolute(X['dist_bond_max']-df['dist_bond_min'])
df['dist_no_bond_min'] = np.nanmin(np.where(dist_matrix_no_bond==0,np.nan,dist_matrix), axis=1)
df['dist_no_bond_max'] = np.nanmax(np.where(dist_matrix_no_bond==0,np.nan,dist_matrix), axis=1)
df['range_dist_no_bond'] = np.absolute(X['dist_no_bond_max']-df['dist_no_bond_min'])
df['n_diff'] = pd.DataFrame(np.around(dist_matrix_bond,5)).nunique(axis=1).values #5
df = reduce_mem_usage(X)
return df
train = create_features(train)
test = create_features(test)
train = create_closest(train)
test = create_closest(test)
train = cosine_features(train)
test = cosine_features(test)
good_columns = ['type',
'bond_lengths_mean_y',
'bond_lengths_std_y',
'bond_lengths_mean_x',
'molecule_atom_index_0_dist_min_div',
'molecule_atom_index_0_dist_std_div',
'molecule_atom_index_0_dist_mean',
'molecule_atom_index_0_dist_max',
'dist_y',
'molecule_atom_index_1_dist_std_diff',
'z_0',
'molecule_type_dist_min',
'molecule_atom_index_0_y_1_mean_div',
'dist_x',
'x_0',
'y_0',
'molecule_type_dist_std',
'molecule_atom_index_0_y_1_std',
'molecule_dist_mean',
'molecule_atom_index_0_dist_std_diff',
'dist_z',
'molecule_atom_index_0_dist_std',
'molecule_atom_index_0_x_1_std',
'molecule_type_dist_std_diff',
'molecule_type_0_dist_std',
'dist',
'molecule_atom_index_0_dist_mean_diff',
'molecule_atom_index_1_dist_min_div',
'molecule_atom_index_1_dist_mean_diff',
'y_1',
'molecule_type_dist_mean_div',
'molecule_dist_max',
'molecule_atom_index_0_dist_mean_div',
'z_1',
'molecule_atom_index_0_z_1_std',
'molecule_atom_index_1_dist_mean_div',
'molecule_atom_index_1_dist_min_diff',
'molecule_atom_index_1_dist_mean',
'molecule_atom_index_1_dist_min',
'molecule_atom_index_1_dist_max',
'molecule_type_0_dist_std_diff',
'molecule_atom_index_0_dist_min_diff',
'molecule_type_dist_mean_diff',
'x_1',
'molecule_atom_index_0_y_1_max',
'molecule_atom_index_0_y_1_mean_diff',
'molecule_atom_1_dist_std_diff',
'molecule_atom_index_0_y_1_mean',
'molecule_atom_1_dist_std',
'molecule_type_dist_max']
```
### Label Encoding
```
categoricals = train.select_dtypes(include='object').columns
categoricals = test.select_dtypes(include='object').columns
# Train Categoricals
for c in categoricals:
lbl = LabelEncoder()
lbl.fit(list(train[c].values))
train[c] = lbl.transform(list(train[c].values))
# Test Categoricals
for c in categoricals:
lbl = LabelEncoder()
lbl.fit(list(test[c].values))
test[c] = lbl.transform(list(test[c].values))
print('train size',train.shape)
print('test size',test.shape)
```
## Modeling
#### Label Define
```
# We define the label
y = train['scalar_coupling_constant']
X = train[good_columns].copy()
X_test = test[good_columns].copy()
duplicate_columns = X.columns[X.columns.duplicated()]
duplicate_columns
X = X.drop(['dist_y'], axis=1);
X_test = X_test.drop(['dist_y'], axis=1);
X = X.drop(['dist_x'], axis=1);
X_test = X_test.drop(['dist_x'], axis=1);
duplicate_columns = X.columns[X.columns.duplicated()]
duplicate_columns
# XGB Matrix Creation
dtrain = xgb.DMatrix(X, label=y)
```
#### K-Folds
```
# Setting a 5-fold stratified cross-validation (note: shuffle=True)
skf = KFold(n_splits=5, shuffle=True, random_state=8)
```
#### Parameter Tuning
```
params = {'booster' : 'gbtree',
# Parameters that we are going to tune.
'max_depth':8,
'min_child_weight': 1,
'eta':0.3,
'subsample': 1,
'colsample_bytree': 1,
# Other parameters
'objective':'reg:linear',
'eval_metric' : 'mae',
}
```
#### Model Training
```
result_dict_xgb = train_model_regression(X=X, X_test=X_test, y=y, params=params, folds=skf, model_type='xgb', eval_metric='group_mae', plot_feature_importance=True,
verbose=1000, early_stopping_rounds=16, n_estimators=10000)
```
## Submission
```
result_dict_xgb.keys()
sub['scalar_coupling_constant'] = result_dict_xgb['prediction']
sub.head()
sub.to_csv('TRR_c_XGB_Molecular_Properties_3.csv', index=False)
```
Score of XXXX
## Plot oof VS target
```
plot_data = pd.DataFrame(y)
plot_data.index.name = 'id'
plot_data['yhat'] = result_dict_xgb['oof']
plot_data['type'] = lbl.inverse_transform(X['type'])
def plot_oof_preds(ctype, llim, ulim):
plt.figure(figsize=(6,6))
sns.scatterplot(x='scalar_coupling_constant',y='yhat',
data=plot_data.loc[plot_data['type']==ctype,
['scalar_coupling_constant', 'yhat']]);
plt.xlim((llim, ulim))
plt.ylim((llim, ulim))
plt.plot([llim, ulim], [llim, ulim])
plt.xlabel('scalar_coupling_constant')
plt.ylabel('predicted')
plt.title(f'{ctype}', fontsize=18)
plt.show()
plot_oof_preds('1JHC', 0, 250)
plot_oof_preds('1JHN', 0, 100)
plot_oof_preds('2JHC', -50, 50)
plot_oof_preds('2JHH', -50, 50)
plot_oof_preds('2JHN', -25, 25)
plot_oof_preds('3JHC', -25, 100)
plot_oof_preds('3JHH', -20, 20)
plot_oof_preds('3JHN', -15, 15)
```
| github_jupyter |
##### Copyright 2020 Google
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Precomputed analysis
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://quantumai.google/cirq/experiments/qaoa/precomputed_analysis"><img src="https://quantumai.google/site-assets/images/buttons/quantumai_logo_1x.png" />View on QuantumAI</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/quantumlib/ReCirq/blob/master/docs/qaoa/precomputed_analysis.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/colab_logo_1x.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/quantumlib/ReCirq/blob/master/docs/qaoa/precomputed_analysis.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/github_logo_1x.png" />View source on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/ReCirq/docs/qaoa/precomputed_analysis.ipynb"><img src="https://quantumai.google/site-assets/images/buttons/download_icon_1x.png" />Download notebook</a>
</td>
</table>
Use precomputed optimal angles to measure the expected value of $\langle C \rangle$ across a variety of problem types, sizes, $p$-depth, and random instances.
## Setup
Install the ReCirq package:
```
try:
import recirq
except ImportError:
!pip install git+https://github.com/quantumlib/ReCirq
```
Now import Cirq, ReCirq and the module dependencies:
```
import recirq
import cirq
import numpy as np
import pandas as pd
```
## Load the raw data
Go through each record, load in supporting objects, flatten everything into records, and put into a massive dataframe.
```
from recirq.qaoa.experiments.precomputed_execution_tasks import \
DEFAULT_BASE_DIR, DEFAULT_PROBLEM_GENERATION_BASE_DIR, DEFAULT_PRECOMPUTATION_BASE_DIR
records = []
for record in recirq.iterload_records(dataset_id="2020-03-tutorial", base_dir=DEFAULT_BASE_DIR):
dc_task = record['task']
apre_task = dc_task.precomputation_task
pgen_task = apre_task.generation_task
problem = recirq.load(pgen_task, base_dir=DEFAULT_PROBLEM_GENERATION_BASE_DIR)['problem']
record['problem'] = problem.graph
record['problem_type'] = problem.__class__.__name__
record['optimum'] = recirq.load(apre_task, base_dir=DEFAULT_PRECOMPUTATION_BASE_DIR)['optimum']
record['bitstrings'] = record['bitstrings'].bits
recirq.flatten_dataclass_into_record(record, 'task')
recirq.flatten_dataclass_into_record(record, 'precomputation_task')
recirq.flatten_dataclass_into_record(record, 'generation_task')
recirq.flatten_dataclass_into_record(record, 'optimum')
records.append(record)
df_raw = pd.DataFrame(records)
df_raw['timestamp'] = pd.to_datetime(df_raw['timestamp'])
df_raw.head()
```
## Narrow down to relevant data
Drop unnecessary metadata and use bitstrings to compute the expected value of the energy. In general, it's better to save the raw data and lots of metadata so we can use it if it becomes necessary in the future.
```
from recirq.qaoa.simulation import hamiltonian_objectives, hamiltonian_objective_avg_and_err
import cirq_google as cg
def compute_energy_w_err(row):
permutation = []
for i, q in enumerate(row['qubits']):
fi = row['final_qubits'].index(q)
permutation.append(fi)
energy, err = hamiltonian_objective_avg_and_err(row['bitstrings'], row['problem'], permutation)
return pd.Series([energy, err], index=['energy', 'err'])
# Start cleaning up the raw data
df = df_raw.copy()
# Don't need these columns for present analysis
df = df.drop(['gammas', 'betas', 'circuit', 'violation_indices',
'precomputation_task.dataset_id',
'generation_task.dataset_id',
'generation_task.device_name'], axis=1)
# p is specified twice (from a parameter and from optimum)
assert (df['optimum.p'] == df['p']).all()
df = df.drop('optimum.p', axis=1)
# Compute energies
df = df.join(df.apply(compute_energy_w_err, axis=1))
df = df.drop(['bitstrings', 'qubits', 'final_qubits', 'problem'], axis=1)
# Normalize
df['energy_ratio'] = df['energy'] / df['min_c']
df['err_ratio'] = df['err'] * np.abs(1/df['min_c'])
df['f_val_ratio'] = df['f_val'] / df['min_c']
df
```
## Plots
```
%matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
sns.set_style('ticks')
plt.rc('axes', labelsize=16, titlesize=16)
plt.rc('xtick', labelsize=14)
plt.rc('ytick', labelsize=14)
plt.rc('legend', fontsize=14, title_fontsize=16)
# theme colors
QBLUE = '#1967d2'
QRED = '#ea4335ff'
QGOLD = '#fbbc05ff'
QGREEN = '#34a853ff'
QGOLD2 = '#ffca28'
QBLUE2 = '#1e88e5'
C = r'\langle C \rangle'
CMIN = r'C_\mathrm{min}'
COVERCMIN = f'${C}/{CMIN}$'
def percentile(n):
def percentile_(x):
return np.nanpercentile(x, n)
percentile_.__name__ = 'percentile_%s' % n
return percentile_
```
### Raw swarm plots of all data
```
import numpy as np
from matplotlib import pyplot as plt
pretty_problem = {
'HardwareGridProblem': 'Hardware Grid',
'SKProblem': 'SK Model',
'ThreeRegularProblem': '3-Regular MaxCut'
}
for problem_type in ['HardwareGridProblem', 'SKProblem', 'ThreeRegularProblem']:
df1 = df
df1 = df1[df1['problem_type'] == problem_type]
for p in sorted(df1['p'].unique()):
dfb = df1
dfb = dfb[dfb['p'] == p]
dfb = dfb.sort_values(by='n_qubits')
plt.subplots(figsize=(7,5))
n_instances = dfb.groupby('n_qubits').count()['energy_ratio'].unique()
if len(n_instances) == 1:
n_instances = n_instances[0]
label = f'{n_instances}'
else:
label = f'{min(n_instances)} - {max(n_instances)}'
#sns.boxplot(dfb['n_qubits'], dfb['energy_ratio'], color=QBLUE, saturation=1)
#sns.boxplot(dfb['n_qubits'], dfb['f_val_ratio'], color=QGREEN, saturation=1)
sns.swarmplot(x=dfb['n_qubits'], y=dfb['energy_ratio'], color=QBLUE)
sns.swarmplot(x=dfb['n_qubits'], y=dfb['f_val_ratio'], color=QGREEN)
plt.axhline(1, color='grey', ls='-')
plt.axhline(0, color='grey', ls='-')
plt.title(f'{pretty_problem[problem_type]}, {label} instances, p={p}')
plt.xlabel('# Qubits')
plt.ylabel(COVERCMIN)
plt.tight_layout()
plt.show()
```
### Compare SK and hardware grid vs. n
```
pretty_problem = {
'HardwareGridProblem': 'Hardware Grid',
'SKProblem': 'SK Model',
'ThreeRegularProblem': '3-Regular MaxCut'
}
df1 = df
df1 = df1[
((df1['problem_type'] == 'SKProblem') & (df1['p'] == 3))
| ((df1['problem_type'] == 'HardwareGridProblem') & (df1['p'] == 3))
]
df1 = df1.sort_values(by='n_qubits')
MINQ = 3
df1 = df1[df1['n_qubits'] >= MINQ]
plt.subplots(figsize=(8, 6))
plt.xlim((8, 23))
# SK
dfb = df1
dfb = dfb[dfb['problem_type'] == 'SKProblem']
sns.swarmplot(x=dfb['n_qubits'], y=dfb['energy_ratio'], s=5, linewidth=0.5, edgecolor='k', color=QRED)
sns.swarmplot(x=dfb['n_qubits'], y=dfb['f_val_ratio'], s=5, linewidth=0.5, edgecolor='k', color=QRED,
marker='s')
dfg = dfb.groupby('n_qubits').mean().reset_index()
# --------
# Hardware
dfb = df1
dfb = dfb[dfb['problem_type'] == 'HardwareGridProblem']
sns.swarmplot(x=dfb['n_qubits'], y=dfb['energy_ratio'], s=5, linewidth=0.5, edgecolor='k', color=QBLUE)
sns.swarmplot(x=dfb['n_qubits'], y=dfb['f_val_ratio'], s=5, linewidth=0.5, edgecolor='k', color=QBLUE,
marker='s')
dfg = dfb.groupby('n_qubits').mean().reset_index()
# -------
plt.axhline(1, color='grey', ls='-')
plt.axhline(0, color='grey', ls='-')
plt.xlabel('# Qubits')
plt.ylabel(COVERCMIN)
from matplotlib.patches import Patch
from matplotlib.lines import Line2D
from matplotlib.legend_handler import HandlerTuple
lelements = [
Line2D([0], [0], color=QBLUE, marker='o', ms=7, ls='', ),
Line2D([0], [0], color=QRED, marker='o', ms=7, ls='', ),
Line2D([0], [0], color='k', marker='s', ms=7, ls='', markerfacecolor='none'),
Line2D([0], [0], color='k', marker='o', ms=7, ls='', markerfacecolor='none'),
]
plt.legend(lelements, ['Hardware Grid', 'SK Model', 'Noiseless', 'Experiment', ], loc='best',
title=f'p = 3',
handler_map={tuple: HandlerTuple(ndivide=None)}, framealpha=1.0)
plt.tight_layout()
plt.show()
```
### Hardware grid vs. p
```
dfb = df
dfb = dfb[dfb['problem_type'] == 'HardwareGridProblem']
dfb = dfb[['p', 'instance_i', 'n_qubits', 'energy_ratio', 'f_val_ratio']]
P_LIMIT = max(dfb['p'])
def max_over_p(group):
i = group['energy_ratio'].idxmax()
return group.loc[i][['energy_ratio', 'p']]
def count_p(group):
new = {}
for i, c in enumerate(np.bincount(group['p'], minlength=P_LIMIT+1)):
if i == 0:
continue
new[f'p{i}'] = c
return pd.Series(new)
dfgy = dfb.groupby(['n_qubits', 'instance_i']).apply(max_over_p).reset_index()
dfgz = dfgy.groupby(['n_qubits']).apply(count_p).reset_index()
# In the paper, we restrict to n > 10
# dfgz = dfgz[dfgz['n_qubits'] > 10]
dfgz = dfgz.set_index('n_qubits').sum(axis=0)
dfgz /= (dfgz.sum())
dfgz
dfb = df
dfb = dfb[dfb['problem_type'] == 'HardwareGridProblem']
dfb = dfb[['p', 'instance_i', 'n_qubits', 'energy_ratio', 'f_val_ratio']]
# In the paper, we restrict to n > 10
# dfb = dfb[dfb['n_qubits'] > 10]
dfg = dfb.groupby('p').agg(['median', percentile(25), percentile(75), 'mean', 'std']).reset_index()
plt.subplots(figsize=(5.5,4))
plt.errorbar(x=dfg['p'], y=dfg['f_val_ratio', 'mean'],
yerr=(dfg['f_val_ratio', 'std'],
dfg['f_val_ratio', 'std']),
fmt='o-',
capsize=7,
color=QGREEN,
label='Noiseless'
)
plt.errorbar(x=dfg['p'], y=dfg['energy_ratio', 'mean'],
yerr=(dfg['energy_ratio', 'std'],
dfg['energy_ratio', 'std']),
fmt='o-',
capsize=7,
color=QBLUE,
label='Experiment'
)
plt.xlabel('p')
plt.ylabel('Mean ' + COVERCMIN)
plt.ylim((0, 1))
plt.text(0.05, 0.9, r'Hardware Grid', fontsize=16, transform=plt.gca().transAxes, ha='left', va='bottom')
plt.legend(loc='center right')
ax2 = plt.gca().twinx() # instantiate a second axes that shares the same x-axis
dfgz_p = [int(s[1:]) for s in dfgz.index]
dfgz_y = dfgz.values
ax2.bar(dfgz_p, dfgz_y, color=QBLUE, width=0.9, lw=1, ec='k')
ax2.tick_params(axis='y')
ax2.set_ylim((0, 2))
ax2.set_yticks([0, 0.25, 0.50])
ax2.set_yticklabels(['0%', None, '50%'])
ax2.set_ylabel('Fraction best' + ' ' * 41, fontsize=14)
plt.tight_layout()
```
| github_jupyter |
# Managing your content
As an organization matures and expands its GIS, users add items of various types and properties with varying relationships to one another. Any administrator must regularly manage items contained in various groups and owned by various users. In this section we demonstrate how to work with individual items in a GIS. This guide shows how to retrieve item properties, delete an existing item, and how to examine relationships between items.
Topics covered in this page:
* [Properties of an item](#properties-of-an-item)
* [Updating item properties](#updating-item-properties)
* [Downloading your items](#downloading-your-items)
* [Deleting content](#deleting-content)
* [Delete protection](#delete-protection)
* [Finding relationships between items](#finding-relationships-between-items)
<a id="properties-of-an-item"></a>
## Properties of an item
An `Item` in your GIS is rich with multiple properties. You can access them as properties on the [`Item`]((https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item)) object;
```
#connect to GIS
from arcgis.gis import GIS
gis = GIS("https://pythonapi.playground.esri.com/portal", "arcgis_python", "amazing_arcgis_123")
#access an Item
volcanoes_item = gis.content.get('452afa4ce761441995cb6d8c69d854d2')
volcanoes_item
# item id
volcanoes_item.id
# title
volcanoes_item.title
# tags
volcanoes_item.tags
```
<a id="updating-item-properties"></a>
### Updating item properties
You can update any of the `Item`s properties using the [`update()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.update) method. It accepts parameters similar to [`add()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.ContentManager.add) method.
```
# update the tags
volcanoes_item.update(item_properties={'tags':'python, vei, empirical, in-situ'})
volcanoes_item.tags
# updating thumbnail
volcanoes_item.update(thumbnail=r'pathway\to\your\directory\IMAGE_NAME.jpg')
volcanoes_item
```
<a id="downloading-your-items"></a>
## Downloading your items
You can download various components of your items, such as the thumbnail, data, or metadata. Downloading assists you in the process of archiving content for a GIS your organization is retiring, or for publishing and migrating content from one GIS to another.
### Download item data
The [`get_data()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.get_data) function reads the binary or text data associated with an item and returns it as a byte array.
* If the text data is JSON, it converts the data to a Python dictionary. If it is another text format, it returns the file as a string.
* If data is not text, binary files are returned along with the path to where the data is downloaded.
```
ports_csv_item = gis.content.get('a0b157b9ce2440a39551967c7c789835')
ports_csv_item
```
The data for a csv item is the csv file itself. It's downloaded to your default temporary directory
```
ports_csv_item.get_data()
```
### Download metadata
You can download metadata into an XML file using the `download_metadata()` method. Once you have the XML file you can edit it and then modify existing portal items by entering the file name as the metadata parameter of the [`Item.update()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.update) method. You can also add it with the metadata parameter on the [`ContentManager.add()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.ContentManager.add) method when creating new content. See the specific documentation for enabling metadata with [ArcGIS Online](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.update) or [ArcGIS Enterprise](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.update).
```
ports_csv_item.download_metadata(save_folder=r'\pathway\to\download\the\metadata\')
```
### Download thumbnail
Finally, you can download the thumbnail of an item using the `download_thumbnail()` method.
```
ports_csv_item.download_thumbnail(save_folder= r'pathway\to\your\directory\')
```
## Deleting content
You can delete any item using the `delete()` method of the `Item` object. When deleting a hosted layer, this operation also deletes the hosted services powering the item.
```
item_for_deletion = gis.content.get('a1752743422b45f791b7eb4dbc5a8010')
item_for_deletion
item_for_deletion.delete()
```
#### Delete protection
You can protect items from getting deleted inadvertently. Use the `protect()` of the `Item` object for this.
```
# let us protect the ports item we accessed earlier
ports_csv_item.protect(enable = True)
# attempting to delete will return an error
ports_csv_item.delete()
```
As expected an exception is raised. You can disable delete protection by setting the `enable` parameter to `False`.
<a id ="finding-relationships-between-items"></a>
## Finding relationships between items
You can add many [types of items](http://enterprise.arcgis.com/en/portal/latest/use/supported-items.htm) to your web GIS. Depending upon the type of item, you can then perform different operations on that item. Once an item is added to the GIS, it seldom exists in isolation but often relates to other items. You might [publish](http://enterprise.arcgis.com/en/portal/latest/use/publish-features.htm) new items from other items to create a new service, or create a new service as a result of a geoprocessing operation on a source item. You may also add one or more layer items to compose a web map or web scene item. Whenever you perform such operations, you establish a relationship between the items. A GIS supports different [relationship types](https://developers.arcgis.com/rest/users-groups-and-items/relationship-types.htm) depending upon the items involved. These relationships represent how items in your GIS are connected to one another. They also convey the impact removing an item from your GIS may have on other items. See the [Relationships between web services and portal items](https://enterprise.arcgis.com/en/server/latest/administer/linux/relationships-between-web-services-and-portal-items.htm) article for more information.
The `Item` class has 3 methods that allow you to determine the relationships to and from an item:
* [`dependent_upon()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.dependent_upon)
* [`dependent_to()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.dependent_to)
* [`related_items()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.related_items)
Let us observe how the ports csv item and the feature layer published from this item share a relationship
> **NOTE:** The Relationship Type functionality is currently evolving within Enterprise and ArcGIS Online implementations. Some items do not currently return all relationships they have between other items.
```
ports_feature_layer = gis.content.get('238d3e97434f40fda38bc846a97b5cfe')
ports_feature_layer
```
Since we know this feature layer item is published from the csv, we can specify the `relationship_type` parameter as `Service2Data`. The direction of relationship would be `forward` as the current item is a service and we are looking for the original data used to publish it.
```
ports_feature_layer.related_items('Service2Data', 'forward')
```
On the ports csv item, the `relationship_type` remains the same whereas the direction is reversed
```
ports_csv_item.related_items('Service2Data', 'reverse')
```
### Relationships of web map items
As seen above, source data and the services published from the data share a relationship. Web Map items share a `Map2Service` relationship with the items used as layers in the map. Let's examine what results we get with a web map using the methods described above.
```
webmap_item = gis.content.get('e92ec599e2a64c69945fe5cd833c8cb8')
webmap_item
```
The `dependent_upon()` method lists all forward relationships for an item. It also lists the items it depends on, and the type of dependency between the two items. In the case of this web map, it has three operational layers dependent upon hosted feature layers (with corresponsding item id values) and a base map layer that depends upon a map service (with corresponding url).
```
webmap_item.dependent_upon()
```
Calling `dependent_to()` lists the items which are dependent upon this web map item, which in this case is none.
```
webmap_item.dependent_to()
```
The table in [this documentation](http://resources.arcgis.com/en/help/arcgis-rest-api/index.html#/Relationship_types/02r3000000mm000000/) gives you the list of all supported relationship types that can exist between two items in your GIS.
### Adding relationships
It is beneficial to add a relationship to establish how information flows from one item to another across a GIS. In the web map example above, the map is dependent upon 4 other items. However, for the hosted feature layers (powered by feature services) the dependency type is listed as `id` instead of `Map2Service`. We can change that by adding a new relationship.
You can add a relationship by using the [`add_relationship()`](https://esri.github.io/arcgis-python-api/apidoc/html/arcgis.gis.toc.html#arcgis.gis.Item.add_relationship) method to pass the item to which the current item is related and the type of relationship you want to create.
```
#from the example above, use the item id of first relationship to get the related item
webmap_related_item = gis.content.get('77561ef541054730af5597ff4d1a3d98')
webmap_related_item
# add a relationship
webmap_item.add_relationship(rel_item= webmap_related_item, rel_type= 'Map2Service')
```
Now that a relationship is added, the property is reflected on the web map item. After the update, querying for related items on the web map item promptly returns the feaure layer collection item.
```
webmap_item.related_items('Map2Service', 'forward')
```
### Deleting relationships
You can remove defunct relationships by calling the `delete_relationship()` method and passing the relationship type and related item.
| github_jupyter |
<center><h1>Python Pandas Tutorial</h1><center>
## Pandas is Python Data Analysis Library
pandas is an open source, BSD-licensed(can use for commercial means) library providing high-performance, easy-to-use data structures and data analysis tools for the Python programming language
* Widely used
* Open Source
* Active Development
* Great Documentation
Home Page: http://pandas.pydata.org/
Using Documentation from: http://pandas.pydata.org/pandas-docs/stable/
Fantastic Cheat Sheet: http://pandas.pydata.org/Pandas_Cheat_Sheet.pdf
Best book by Panda's creator Wes Kinney (2nd Edition 2017): http://shop.oreilly.com/product/0636920050896.do
```
import pandas as pd
#pd.reset_option('display.max_rows')
pd.options.display.max_rows = 40
# Pandas is a big package took a while...
import numpy as np # another big library with various numeric functions
import matplotlib.pyplot as plt
```
# Panda's two fundamental data structures: Series and DataFrame.
### Series
A Series is a one-dimensional array-like object containing a sequence of values (
similar types to NumPy types) and an associated array of data labels - index.
Simplest Series is from an array of data.
```
# Let's create some Series!
s = pd.Series([1,4,3.5,3,np.nan,0,-5])
s
s+4
# NaN = Not a Number (used for missing numerical values)
# https://pandas.pydata.org/pandas-docs/stable/user_guide/missing_data.html
s2 = s * 4
s2
s2**2
### Often you want Series with an index identifying each data point with a label
labeledSeries = pd.Series([24, 77, -35, 31], index=['d', 'e', 'a', 'g'])
labeledSeries
## A bit similar to dictionary isn't it?
labeledSeries['g']
labeledSeries.index
labeledSeries.values
labeledSeries[['a','d']] # NOTE double list brackets!!
# select values via a boolean array
labeledSeries[labeledSeries > 30]
# So Series behaves like a fixed-length, ordered dictionary with extra helper methods
'd' in labeledSeries
```
### Can create series from dictionary by simply passing to constructor pd.Series(mydict)
```
citydict = {'Riga': 650000, 'Tukums':20000, 'Ogre': 25000, 'Carnikava': 3000}
citydict
cseries = pd.Series(citydict)
cseries
## Overwriting default index
clist = ['Jurmala', 'Riga', 'Tukums', 'Ogre', 'Daugavpils']
cseries2 = pd.Series(citydict, index = clist)
cseries2
# notice Carnikava was lost, since our index did not have it!
# and order was preserved from the given index list!
# For missing data
cseries2.isnull()
cseries2.dropna()
cseries2
cseries3 = cseries + cseries2
cseries3
# So NaN + number = NaN
cseries.name = "Latvian Cities"
cseries.index.name = "City"
cseries
cseries.index
cseries.index = ['CarnikavaIsNotaCity','OgreEatsHumans', 'RigaIsOld', 'TukumsSmukums']
cseries
# Series values are mutable
cseries['RigaIsOld']=625000
cseries
# How to rename individual index elements?
cseries.index[2]='RigaIsOldButFantastic'
cseries
# We use Renaming method to rename individual elements
cseries.rename(index={'RigaIsOld':'RigaRocks'})
```
### Integer (Position-based) vs Label-based Indexes
Working with pandas objects indexed by integers is something that often trips up
new users due to some differences with indexing semantics on built-in Python data
structures like lists and tuples. For example, you might not expect the following code
to generate an error:
```
ser = pd.Series(np.arange(3.))
ser
ser[-1]
```
In this case, pandas could “fall back” on integer indexing, but it’s difficult to do this in
general without introducing subtle bugs.
Here we have an index containing 0, 1, 2,
but inferring what the user wants (label-based indexing or position-based) is difficult:
```
ser
## With a non-integer index there is no potential for ambiguity:
ser2 = pd.Series(np.arange(3.), index=['a', 'b', 'c'])
ser2[-1]
ser2[::-1]
## To keep things consistent, if you have an axis index containing integers, data selection
##will always be label-oriented.
# For more precise handling, use loc (for labels) or iloc (for integers):
ser2.loc['b']
# Note: label indexing includes the endpoint, integer indexing does not
ser.loc[:1]
ser.iloc[:1]
```
* loc gets rows (or columns) with particular labels from the index.
* iloc gets rows (or columns) at particular positions in the index (so it only takes integers).
# Date Range creation
Date ranges are used as indexes for time series data:
* https://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html#time-series
```
dates = pd.date_range('20180521', periods=15)
dates
pd.date_range('20200416', periods=15, freq="W")
pd.date_range('20200416', periods=7, freq="W-THU")
# more on data_range frequency here
# https://stackoverflow.com/questions/35339139/where-is-the-documentation-on-pandas-freq-tags
# Datetime is in the standard library (so all Python installations will have it)
from datetime import date
date.today()
# We can get a data range starting from today
months = pd.date_range(date.today().strftime("%Y-%m-%d"), periods = 10, freq='BMS')
months
```
## Reading data files
```
city_data = pd.read_csv("data/iedz_skaits_2018.csv", index_col=0)
city_data.head()
type(city_data)
city_series = city_data.squeeze()
type(city_series)
city_series.head()
city_series["Salaspils"]
city_series.sum()
city_series.describe()
city_series[city_series < 1000]
bitmap = city_series < 1000
bitmap.sample(20) # kādēļ sample() nevis head()
city_series[bitmap].sort_index()
```
## DataFrame
A DataFrame represents a rectangular table of data and contains an ordered collection of columns.
Each column can be a different value type (numeric, string,
boolean, etc.).
The DataFrame has both a row and column index;
Think of it
as an ordered dict of Series all sharing the same row index.
Underneath data is stored as one or more two-dimensional blocks (similar to ndarray)
rather than a list, dict, or some other collection of
one-dimensional arrays.
```
# Many ways of Data Frame creation
# One Common way is common is
# from a dict of equal-length lists or NumPy arrays
data = {'city': ['Riga', 'Riga', 'Riga', 'Jurmala', 'Jurmala', 'Jurmala'],
'year': [1990, 2000, 2018, 2001, 2002, 2003],
'popul': [0.9, 0.75, 0.62, 0.09, 0.08, 0.06]}
df = pd.DataFrame(data)
df
df2 = pd.DataFrame(data, columns=['year','city', 'popul','budget'])
df2
# missing column simply given Nans
df2['budget']=300000000
df2
df2['budget']=[300000, 250000, 400000, 200000, 250000, 200000] # need to pass all values
df2
# Many ways of changing individual values
## Recommended way of changing in place (same dataframe)
df2.iat[3,2]=0.063
df2
df2["budget"]
# delete column by its name
del df2["budget"]
df2
df = pd.DataFrame(np.random.randn(15,5), index=dates, columns=list('ABCDE'))
# We passed 15 rows of 5 random elements and set index to dates and columns to our basic list elements
df
df2 = pd.DataFrame({ 'A' : 1.,
'B' : pd.Timestamp('20130102'),
'C' : pd.Series(1,index=list(range(4)),dtype='float32'),
'D' : np.array([3] * 4,dtype='int32'),
'E' : pd.Categorical(["test","train","test","train"]),
'F' : 'foo' })
df2
#most columns need matching length!
df3 = pd.DataFrame({ 'A' : 1.,
'B' : pd.Timestamp('20180523'),
'C' : s,
'D' : [x**2 for x in range(7)],
'E' : pd.Categorical(['test','train']*3+["train"]),
'F' : 'aha'
})
df3
## different datatypes for columns!
df3.dtypes
df3.head()
df3.tail(3)
df.index
df3.index
df3.values
df3.describe()
df.info()
# Transpose
df3.T
df.sort_index(axis=1,ascending=True)
## Sort by Axis in reverse
df.sort_index(axis=1,ascending=False)
df3.sort_values(by='C', ascending=False)
# Notice NaN gets last
```
### Selection
Note: while standard Python / Numpy expressions for selecting and setting are intuitive and come in handy for interactive work, for production code, we recommend the optimized pandas data access methods, .at, .iat, .loc and .iloc.
```
df3['D']
df3[:5]
df3[2:5]
df3[2:5:2]
df3[::-1]
```
## Selection by Label
For getting a cross section using a label:
```
df
df.loc[dates[0]]
df.loc[dates[2:5]]
## Selecting on a multi-axis by label:
df.loc[:, ['A','B','C']]
df.loc[dates[2:5], ['A','B','C']]
df.loc['20180525':'20180601',['B','C']]
# Reduction in the dimensions of the returned object:
df.loc['20180526', ["B", "D"]]
## Getting scalars (single values)
df.loc['20180526', ["D"]]
# same as above
df.at[dates[5],'D']
## Selection by Position
df.iloc[3]
# By integer slices, acting similar to numpy/python:
df.iloc[2:5,:2]
# By lists of integer position locations, similar to the numpy/python style:
df.iloc[[3,5,1],[1,4,2]]
df.iloc[2,2]
# For getting fast access to a scalar (equivalent to the prior method):
df.iat[2,2]
```
## Boolean Indexing
```
## Using a single column’s values to select data.
df[df.A > 0.2]
df[df > 0]
df[df > 1]
# TODO - fix this!
s1 = pd.Series([x**3 for x in range(15)], index=pd.date_range('20180521', periods=15))
s1
df['F'] = s1
df
## This is apparently a bug! https://github.com/pandas-dev/pandas/issues/10440
df['F']=42
df
df['G']=[x**3 for x in range(15)] # passing a fresh list to particular column
df
df.at[dates[1], 'A'] = 33
df
df.iat[4,4]= 42
df
df3 = df.copy()
df3
df3[df3 > 0.2 ] = -df3
df3
# Missing Data
# pandas primarily uses the value np.nan to represent missing data. It is by default not included in computations
df['H'] = s1
df
df.fillna(value=3.14)
# there is also df.dropna() to drop any ROWS! with missing data
```
## Operations
DataFrame methods and properties:
* https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html
Series methods and properties:
* https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.html
Data Science Handbook:
* [Data manipulation with Pandas](https://jakevdp.github.io/PythonDataScienceHandbook/index.html#3.-Data-Manipulation-with-Pandas)
```
df.mean()
df.max()
# Other axis
df.mean(axis=1)
## String operations (df.str.*)
str1 = pd.Series(['APPle', 'baNAna', np.NaN, 42, 'mangO'])
str1
str1.str.lower()
str1.str.len()
## Apply
df.apply(lambda x: x*3) # ie same as df*3
df = pd.DataFrame({'key': ['A', 'B', 'C', 'A', 'B', 'C'], 'data': range(6)}, columns=['key', 'data'])
df
df.groupby('key')
df.groupby('key').sum()
```
### Time series
```
ts = pd.Series(np.random.randn(3650), index=pd.date_range('11/18/2008', periods=3650))
ts = ts.cumsum() # cumulative sum
ts.plot()
ts["2014-01-01":"2016-01-01"] = np.NaN
ts.plot()
rolling_avg = ts.rolling(window=90).mean()
rolling_avg
rolling_avg.plot()
```
## File operations
```
# CSV
# Writing to a csv file.
df.to_csv("testing.csv")
# Reading from csv
new_df = pd.read_csv("testing.csv", index_col=0)
new_df.head()
# Excel
df.to_excel('myx.xlsx', sheet_name='Sheet1')
df6=pd.read_excel('myx.xlsx', 'Sheet1', index_col=None, na_values=['NA'])
df6.head()
# basic DataFrame information:
df.info()
df.info(memory_usage="deep") # more reliable info
```
| github_jupyter |
# Demo ARIMA Time Series Forecasting on Ray local
<b>Suggestion: Make a copy of this notebook. This way you will retain the original, executed notebook outputs. Make edits in the copied notebook. </b>
### Description:
This notebook goes along with the tutorial <a href="https://towardsdatascience.com/scaling-time-series-forecasting-with-ray-arima-and-prophet-e6c856e605ee">How to Train Faster Time Series Forecasting Using Ray, part 1 of 2<a>.
This notebook demonstrates Time Series Forecasting ARIMA algorithm on Ray. Example data is NYC yellow taxi from: https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page <br>
Forecast goal: Given 6 months historical taxi trips data for NYC, your task is to predict #pickups at each location in NYC at monthly level for the next 2 months.
### Demo notes:
Output shows timings using MEDIUM dataset <br>
Both demo datasets are available in this github repo under data/ <br>
SMALL dataset contains original, actual 260 items "clean_taxi_monthly.parquet" <br>
MEDIUM dataset contains 2860 items with extra fakes "clean_taxi_monthly_fake_medium.parquet" <br>
```
# install open-source Ray if you haven't already
# !pip install "ray[default] installs the latest version; otherwise use a specific version
# !pip install "ray[default]==1.9.0"
# install ARIMA library
# !pip install pmdarima
# install Anyscale to run Ray easily on a Cloud
# !pip install anyscale
###########
# Import libraries
###########
# Open-source libraries
import os # Python os functions
import time # Python time functions
import warnings # Python warnings
warnings.filterwarnings('ignore')
import ray # Run distributed code
import numpy as np # Numerical processing
import pandas as pd # Dataframe (tabular data) processing
import matplotlib as mpl # Graph plotting
import matplotlib.pyplot as plt
%matplotlib inline
import pickle
# Open-source ARIMA forecasting libraries
arima_model_type = "pmdarima.arima.arima.ARIMA"
import pmdarima as pm
from pmdarima.model_selection import train_test_split
!python --version
print(f"ray: {ray.__version__}")
print(f"numpy: {np.__version__}")
print(f"pandas: {pd.__version__}")
print(f"matplotlib: {mpl.__version__}")
print(f"pmdarima: {pm.__version__}")
AVAILABLE_LOCAL_CPU = os.cpu_count()
```
# Change how you want to run Ray below.
<b>Depending on whether you want to run Ray Local or Ray in a Cloud:</b>
<ul>
<li><b>To run Ray Local, change below variables, then continue running cells in the notebook</b>: <br>
RUN_RAY_LOCAL = True; RUN_RAY_ON_A_CLOUD = False</li>
<li><b>To run Ray in a Cloud, change below variables, then continue running cells in the notebook</b>: <br>
RUN_RAY_LOCAL = False; RUN_RAY_ON_A_CLOUD = True </li>
</ul>
```
###########
# CHANGE VARIABLES BELOW.
# To run Ray Local: RUN_RAY_LOCAL = True; RUN_RAY_ON_A_CLOUD = False
# To run Ray in a Cloud: RUN_RAY_LOCAL = False; RUN_RAY_ON_A_CLOUD = True
###########
RUN_RAY_LOCAL = True
RUN_RAY_ON_A_CLOUD = False
###########
# Run Ray Local on your laptop for testing purposes
# Dashboard doc: https://docs.ray.io/en/master/ray-dashboard.html#ray-dashboard
###########
if RUN_RAY_LOCAL:
# num_cpus, num_gpus are optional parameters
# by default Ray will detect and use all available
NUM_CPU = AVAILABLE_LOCAL_CPU
print(f"You are running Ray Local with {NUM_CPU} CPUs")
# start up ray locally
if ray.is_initialized():
ray.shutdown()
ray.init()
else:
print("You are not running Ray Local")
###########
# Run Ray in the Cloud using Anyscale
# View your cluster on console.anyscale.com
###########
if RUN_RAY_ON_A_CLOUD:
print("You are running Ray on a Cloud")
# !pip install anyscale # install anyscale if you haven't already
import anyscale
# You can specify more pip installs, clone github, or copy code/data here in the runtime env.
# Everything in the runtime environment will override the cluster environment.
# https://docs.anyscale.com/user-guide/configure/dependency-management/anyscale-environments
my_env={ "working_dir": ".",
"pip": ["pmdarima"],
}
# start up ray in any cloud
if ray.is_initialized():
ray.shutdown()
ray.init(
"anyscale://christy-forecast3",
# runtime_env=my_env,
# optionally put pip installs in the cluster config instead of runtime_env
cluster_env="christy-forecast:4"
)
else:
print("You are not running Ray on a Cloud")
```
# Read 8 months clean NYC taxi data
New York City Yellow Taxi ride volumes per location (8 months of historical data). <ul>
<li>Original source: https://www1.nyc.gov/site/tlc/about/tlc-trip-record-data.page</li>
<li>Clean monthly source: https://github.com/christy/AnyscaleDemos/blob/main/forecasting_demos/data/clean_taxi_monthly_fake_medium.parquet?raw=true </li>
</ul>
Normally there is a data cleaning/prep step to convert raw data -> cleaned data. We'll dig into details of ETL later. <br>
For now, let's just start with cleaned, aggregated monthly data for ARIMA and Prophet, since those algorithms are typically for strategic-level forecasting, not typically for detailed-level forecasting.
```
############
# Read pandas dataframe
# If you cloned this notebook from github the data should be in your data/ folder
###########
# read 8 months of clean, aggregated monthly taxi data
# filename = "https://github.com/christy/AnyscaleDemos/blob/main/forecasting_demos/data/clean_taxi_monthly_fake_medium.parquet?raw=true"
# filename = "data/clean_taxi_monthly.parquet"
filename = "data/clean_taxi_monthly_fake_medium.parquet"
g_month = pd.read_parquet(filename)
# rename "time" column, since prophet expects that, arima doesn't care
g_month.reset_index(inplace=True)
g_month.rename(columns={"pickup_monthly": "time"}, inplace=True)
display(g_month.head())
# Train a model per item_id
item_list = list(g_month["pulocationid"].unique())
print(f"Number unique items = {len(item_list)}")
###########
# Assume these are already-existing functions.
###########
###########
# Define a train_model function, default train on 6 months, inference 2
###########
def train_model_ARIMA(
theDF: pd.DataFrame,
item_col: str,
item_value: str,
target_col: str,
train_size: int = 6,
) -> list:
"""This function trains a model using ARIMA algorithm.
Args:
theDF (pd.DataFrame): Input data. It must be indexed by "time".
item_col (str): Name of the column containing item_id or SKU.
item_value (str): Value of the item_id or SKU being forecasted.
target_col (str): Name of the column containing the actual value.
train_size (int, optional): Count of number of timestamps to use
for training. Defaults to 6.
Returns:
list: [
train (pd.DataFrame): Training data.
test (pd.DataFrame): Test data for evaluation.
model (pmdarima.arima.arima.ARIMA): ARIMA model for inference.
]
"""
import warnings
warnings.filterwarnings("ignore")
# split data into train/test
train, test = train_test_split(
theDF.loc[(theDF[item_col] == item_value), :], train_size=train_size
)
# train and fit auto.arima model
model = pm.auto_arima(
y=train[target_col],
X=train.loc[:, (train.columns != target_col) & (train.columns != item_col)],
)
return [train, test, model]
###########
# Define inference_model function
###########
def inference_model_ARIMA(
model: "pmdarima.arima.arima.ARIMA",
test: pd.DataFrame,
item_col: str,
target_col: str,
) -> pd.DataFrame:
"""This function inferences a model using ARIMA algorithm. It uses
the actual values, if known, in the test evaluation dataframe
and concats them into the forecast output dataframe,
for easier evaluation later.
Args:
model (pmdarima.arima.arima.ARIMA): ARIMA model for inference.
test (pd.DataFrame): Test data for evaluation.
item_col (str): Name of the column containing item_id or SKU.
target_col (str): Name of the column containing the actual value.
Returns:
pd.DataFrame: Forecast as pandas dataframe containing the forecast along
with actual values.
"""
# inference on test data
forecast = pd.DataFrame(
model.predict(
n_periods=test.shape[0],
X=test.loc[:, (test.columns != target_col) & (test.columns != item_col)],
index=test.index,
)
)
# put both actual_value and predicted_value in forecast, for easier eval later
forecast.columns = ["predicted_value"]
forecast.predicted_value = forecast.predicted_value.astype(np.int32)
forecast = pd.concat([forecast, test.loc[:, target_col].reset_index()], axis=1)
forecast.set_index("time", inplace=True)
return forecast
```
# Regular Python
```
# keyboard interrupt - this takes too long!
#%%timeit
###########
# Main Regular Python program flow to train and inference ARIMA models
###########
# initialize objects
train = []
test = []
model = []
forecast = []
start = time.time()
# Train every model
train, test, model = map(
list,
zip(
*(
[
train_model_ARIMA(
g_month.set_index("time"),
item_col="pulocationid",
item_value=v,
target_col="trip_quantity",
train_size=6,
)
for p, v in enumerate(item_list)
]
)
),
)
# Inference every model
forecast = [
inference_model_ARIMA(
model[p], test[p], item_col="pulocationid", target_col="trip_quantity"
)
for p in range(len(item_list))
]
time_regular_python = time.time() - start
print(f"Done! ARIMA on Regular Python finished in {time_regular_python} seconds")
# Run once more without timeit to set the variables and inspect
###########
# Main Regular Python program flow to train and inference ARIMA models
###########
# initialize objects
train = []
test = []
model = []
forecast = []
start = time.time()
# Train every model
train, test, model = map(
list,
zip(
*(
[
train_model_ARIMA(
g_month.set_index("time"),
item_col="pulocationid",
item_value=v,
target_col="trip_quantity",
train_size=6,
)
for p, v in enumerate(item_list)
]
)
),
)
# Inference every model
forecast = [
inference_model_ARIMA(
model[p], test[p], item_col="pulocationid", target_col="trip_quantity"
)
for p in range(len(item_list))
]
time_regular_python = time.time() - start
print(f"Done! ARIMA on Regular Python finished in {time_regular_python} seconds")
###########
# inspect a few forecasts
###########
assert len(model) == len(item_list)
assert len(forecast) == len(item_list)
print(f"len(forecast): {len(forecast)}")
# plot first two forecasts
plt.figure(figsize=(8, 5))
for p, v in enumerate(item_list[0:2]):
display(forecast[p])
plt.plot(train[p]["trip_quantity"], label="Train")
plt.plot(test[p]["trip_quantity"], label="Test")
plt.plot(forecast[p]["predicted_value"], label="Forecast")
plt.legend(loc="best")
```
# Ray distributed Python
```
###########
# Convert existing functions to Ray parallelized functions
###########
###########
# Define a train_model function, default train on 6 months, inference 2
###########
@ray.remote(num_returns=3)
def train_model_ARIMA_remote(
theDF: pd.DataFrame,
item_col: str,
item_value: str,
target_col: str,
train_size: int = 6,
) -> list:
"""This function trains a model using ARIMA algorithm for use by Ray
parallelization engine for distributed training on a cluster.
Args:
theDF (pd.DataFrame): [description]
item_col (str): Name of the column containing item_id or SKU.
item_value (str): Value of the item_id or SKU being forecasted.
target_col (str): Name of the column containing the actual value.
train_size (int, optional): Count of number of timestamps to use
for training. Defaults to 6.
Returns:
list: [
train (pd.DataFrame): Training data.
test (pd.DataFrame): Test data for evaluation.
model (pmdarima.arima.arima.ARIMA): ARIMA model for inference.
]
"""
import warnings
warnings.filterwarnings("ignore")
# split data into train/test
train, test = train_test_split(
theDF.loc[(theDF[item_col] == item_value), :], train_size=train_size
)
# train and fit auto.arima model
model = pm.auto_arima(
y=train[target_col],
X=train.loc[:, (train.columns != target_col) & (train.columns != item_col)],
)
# here is the extra pickle step only required for statsmodels objects
# Explanation why https://alkaline-ml.com/pmdarima/1.0.0/serialization.html
# return [train, test, model]
return [train, test, pickle.dumps(model)]
###########
# Define inference_model function
###########
@ray.remote
def inference_model_ARIMA_remote(
model_pickle: bytes, test: pd.DataFrame, item_col: str, target_col: str
) -> pd.DataFrame:
"""This function can run on a cluster distributed by the Ray parallelization
engine to inference a model using ARIMA algorithm. It uses the
actual values, if known, in the test evaluation dataframe and
concats them into the forecast output dataframe, for easier
evaluation later.
Args:
model_pickle (bytes): Serialized ARIMA model.
test (pd.DataFrame): Ray remote reference to the pandas test dataframe.
item_col (str): Name of the column containing item_id or SKU.
target_col (str): Name of the column containing the actual value.
Returns:
pd.DataFrame: forecast is a Ray remote reference to a pandas dataframe
containing the forecast along with actual values.
"""
# Here is extra unpickle step
model = pickle.loads(model_pickle)
# inference on test data
forecast = pd.DataFrame(
model.predict(
n_periods=test.shape[0],
X=test.loc[:, (test.columns != target_col) & (test.columns != item_col)],
index=test.index,
)
)
# put both actual_value and predicted_value in forecast, for easier eval later
forecast.columns = ["predicted_value"]
forecast.predicted_value = forecast.predicted_value.astype(np.int32)
forecast = pd.concat([forecast, test.loc[:, target_col].reset_index()], axis=1)
forecast.set_index("time", inplace=True)
return forecast
#%%timeit
###########
# Main Ray distributed program flow to train and inference ARIMA models
###########
model = []
train = []
test = []
forecast_obj_refs = []
# initialize data in ray object store on each cluster
input_data_ref = ray.put(g_month.set_index("time"))
start = time.time()
# Train every model
train, test, model = map(
list,
zip(
*(
[
train_model_ARIMA_remote.remote(
input_data_ref,
item_col="pulocationid",
item_value=v,
target_col="trip_quantity",
train_size=6,
)
for p, v in enumerate(item_list)
]
)
),
)
# Inference every model
forecast_obj_refs = [
inference_model_ARIMA_remote.remote(
model[p], test[p], item_col="pulocationid", target_col="trip_quantity"
)
for p in range(len(item_list))
]
# ray.get() means block until all objectIDs requested are available
forecast_ray = ray.get(forecast_obj_refs)
time_ray_local = time.time() - start
print(f"Done! ARIMA on Ray Local finished in {time_ray_local} seconds")
```
# Verify forecasts
```
# Run the Ray local code again to get the forecasts
###########
# Main Ray distributed program flow to train and inference ARIMA models
###########
model = []
train = []
test = []
forecast_obj_refs = []
# initialize data in ray object store on each cluster
input_data_ref = ray.put(g_month.set_index("time"))
start = time.time()
# Train every model
train, test, model = map(
list,
zip(
*(
[
train_model_ARIMA_remote.remote(
input_data_ref,
item_col="pulocationid",
item_value=v,
target_col="trip_quantity",
train_size=6,
)
for p, v in enumerate(item_list)
]
)
),
)
# Inference every model
forecast_obj_refs = [
inference_model_ARIMA_remote.remote(
model[p], test[p], item_col="pulocationid", target_col="trip_quantity"
)
for p in range(len(item_list))
]
# ray blocking step, to get the forecasts
# ray.get() means block until all objectIDs requested are available
forecast_ray = ray.get(forecast_obj_refs)
time_ray_local = time.time() - start
print(f"Done! ARIMA on Ray Local finished in {time_ray_local} seconds")
# Calculate speedup:
speedup = time_regular_python / time_ray_local
print(f"Speedup from running Ray parallel code on your laptop: {np.round(speedup, 1)}x"
f", or {(np.round(speedup, 0)-1) * 100}%")
# Verify ray forecast is same as regular Python forecast
assert len(forecast_ray) == len(forecast)
assert len(forecast_ray[0]) == len(forecast[0])
assert forecast_ray[0].equals(forecast[0])
###########
# inspect a few forecasts
###########
assert len(model) == len(item_list)
assert len(forecast) == len(item_list)
print(f"len(forecast): {len(forecast_ray)}")
# plot first two forecasts
train = ray.get(train)
test = ray.get(test)
plt.figure(figsize=(8, 5))
for p in range(len(item_list[0:2])):
display(forecast_ray[p])
plt.plot(train[p]["trip_quantity"], label="Train")
plt.plot(test[p]["trip_quantity"], label="Test")
plt.plot(forecast_ray[p]["predicted_value"], label="Forecast")
plt.legend(loc="best")
# fancier plots
# plot first two forecasts
fig, axs = plt.subplots(2, 1, figsize=(8, 5), sharex=True)
for p, v in enumerate(item_list[0:2]):
print(f"Forecast for item {v}:")
display(forecast_ray[p])
ax = axs[p]
train[p].trip_quantity.plot(ax=ax, label="Train")
test[p].trip_quantity.plot(ax=ax, label="Test")
forecast_ray[p].predicted_value.plot(ax=ax, label="Forecast")
ax.legend(loc="best")
ax.set_title(f"item {v}")
###########
# verify intermediate model artifacts, if desired
###########
# ray.get the model artifacts, in case you want to check them
model = ray.get(model)
assert len(train) == len(test)
assert len(train) == len(model)
# verify types of objects you got back from ray...
for p in range(len(item_list[0:1])): # just look at first in the list
print(f"type(train[{p}]): {type(train[p])}")
print(f"type(test[{p}]): {type(test[p])}")
display(train[p])
display(test[p])
print()
print(f"type(model[{p}]) before unpickling: {type(model[p])}")
model[p] = pickle.loads(model[p])
print(f"type(model[{p}]) after unpickling: {type(model[p])}")
display(model[p])
```
# Now run the same code as Ray Local, but this time run using Anyscale in any Cloud.
<b>
<ol>
<li>Go back to top of notebook </li>
<li>Change variables RUN_RAY_LOCAL = False; RUN_RAY_ON_A_CLOUD = True <br>
... And run the next 2 cells to propertly shutdown/start Ray </li>
<li>Come back here to bottom of notebook <br>
Run cell below.</li>
</ul>
</b>
```
%%timeit
###########
# Main Ray distributed program flow to train and inference ARIMA models
###########
model = []
train = []
test = []
# This step required for Ray 1.8 and below
# initialize data in ray object store on each cluster
input_data_ref = ray.put(g_month.set_index("time"))
start = time.time()
# Train every model
train, test, model = map(
list,
zip(
*(
[
train_model_ARIMA_remote.remote(
input_data_ref,
item_col="pulocationid",
item_value=v,
target_col="trip_quantity",
train_size=6,
)
for p, v in enumerate(item_list)
]
)
),
)
# Inference every model
forecast_obj_refs = [
inference_model_ARIMA_remote.remote(
model[p], test[p], item_col="pulocationid", target_col="trip_quantity"
)
for p in range(len(item_list))
]
# ray blocking step, to get the forecasts
# ray.get() means block until all objectIDs requested are available
forecast_ray = ray.get(forecast_obj_refs)
time_ray_cloud = time.time() - start
print(f"Done! ARIMA on Ray in Cloud finished in {time_ray_cloud} seconds")
time_ray_cloud = 23
# Calculate speedup running parallel Python Ray in a Cloud:
speedup = time_regular_python / time_ray_cloud
print(f"Speedup from running Ray parallel code in a Cloud: {np.round(speedup, 1)}x"
f", or {(np.round(speedup, 0)-1) * 100}%")
ray.shutdown()
```
| github_jupyter |
```
# default_exp trainer.vits
```
# Debugging
```
import math
import os
import random
import time
import torch
from torch import nn
import torch.nn.functional as F
import torch.utils.data
import numpy as np
import librosa
import librosa.util as librosa_util
from librosa.util import normalize, pad_center, tiny
from scipy.signal import get_window
from scipy.io.wavfile import read
from librosa.filters import mel as librosa_mel_fn
MAX_WAV_VALUE = 32768.0
torch.backends.cudnn.benchmark = True
def dynamic_range_compression_torch(x, C=1, clip_val=1e-5):
"""
PARAMS
------
C: compression factor
"""
return torch.log(torch.clamp(x, min=clip_val) * C)
def dynamic_range_decompression_torch(x, C=1):
"""
PARAMS
------
C: compression factor used to compress
"""
return torch.exp(x) / C
def spectral_normalize_torch(magnitudes):
output = dynamic_range_compression_torch(magnitudes)
return output
def spectral_de_normalize_torch(magnitudes):
output = dynamic_range_decompression_torch(magnitudes)
return output
mel_basis = {}
hann_window = {}
def spectrogram_torch(y, n_fft, sampling_rate, hop_size, win_size, center=False):
if torch.min(y) < -1.0:
print("min value is ", torch.min(y))
if torch.max(y) > 1.0:
print("max value is ", torch.max(y))
global hann_window
dtype_device = str(y.dtype) + "_" + str(y.device)
wnsize_dtype_device = str(win_size) + "_" + dtype_device
if wnsize_dtype_device not in hann_window:
hann_window[wnsize_dtype_device] = torch.hann_window(win_size).to(
dtype=y.dtype, device=y.device
)
y = torch.nn.functional.pad(
y.unsqueeze(1),
(int((n_fft - hop_size) / 2), int((n_fft - hop_size) / 2)),
mode="reflect",
)
y = y.squeeze(1)
spec = torch.stft(
y,
n_fft,
hop_length=hop_size,
win_length=win_size,
window=hann_window[wnsize_dtype_device],
center=center,
pad_mode="reflect",
normalized=False,
onesided=True,
)
spec = torch.sqrt(spec.pow(2).sum(-1) + 1e-6)
return spec
def spec_to_mel_torch(spec, n_fft, num_mels, sampling_rate, fmin, fmax):
global mel_basis
dtype_device = str(spec.dtype) + "_" + str(spec.device)
fmax_dtype_device = str(fmax) + "_" + dtype_device
if fmax_dtype_device not in mel_basis:
mel = librosa_mel_fn(sampling_rate, n_fft, num_mels, fmin, fmax)
mel_basis[fmax_dtype_device] = torch.from_numpy(mel).to(
dtype=spec.dtype, device=spec.device
)
spec = torch.matmul(mel_basis[fmax_dtype_device], spec)
spec = spectral_normalize_torch(spec)
return spec
def mel_spectrogram_torch(
y, n_fft, num_mels, sampling_rate, hop_size, win_size, fmin, fmax, center=False
):
if torch.min(y) < -1.0:
print("min value is ", torch.min(y))
if torch.max(y) > 1.0:
print("max value is ", torch.max(y))
global mel_basis, hann_window
dtype_device = str(y.dtype) + "_" + str(y.device)
fmax_dtype_device = str(fmax) + "_" + dtype_device
wnsize_dtype_device = str(win_size) + "_" + dtype_device
if fmax_dtype_device not in mel_basis:
mel = librosa_mel_fn(sampling_rate, n_fft, num_mels, fmin, fmax)
mel_basis[fmax_dtype_device] = torch.from_numpy(mel).to(
dtype=y.dtype, device=y.device
)
if wnsize_dtype_device not in hann_window:
hann_window[wnsize_dtype_device] = torch.hann_window(win_size).to(
dtype=y.dtype, device=y.device
)
y = torch.nn.functional.pad(
y.unsqueeze(1),
(int((n_fft - hop_size) / 2), int((n_fft - hop_size) / 2)),
mode="reflect",
)
y = y.squeeze(1)
spec = torch.stft(
y,
n_fft,
hop_length=hop_size,
win_length=win_size,
window=hann_window[wnsize_dtype_device],
center=center,
pad_mode="reflect",
normalized=False,
onesided=True,
)
spec = torch.sqrt(spec.pow(2).sum(-1) + 1e-6)
spec = torch.matmul(mel_basis[fmax_dtype_device], spec)
spec = spectral_normalize_torch(spec)
return spec
```
# Trainer
```
# export
import json
import os
from pathlib import Path
from pprint import pprint
import torch
from torch.cuda.amp import autocast, GradScaler
import torch.distributed as dist
from torch.nn import functional as F
from torch.nn.parallel import DistributedDataParallel as DDP
from torch.optim.lr_scheduler import ExponentialLR
from torch.utils.data import DataLoader
from tensorboardX import SummaryWriter
import time
from uberduck_ml_dev.models.common import MelSTFT
from uberduck_ml_dev.utils.plot import (
plot_attention,
plot_gate_outputs,
plot_spectrogram,
)
from uberduck_ml_dev.text.util import text_to_sequence, random_utterance
from uberduck_ml_dev.text.symbols import symbols_with_ipa
from uberduck_ml_dev.trainer.base import TTSTrainer
from uberduck_ml_dev.models.vits import (
DEFAULTS,
MultiPeriodDiscriminator,
SynthesizerTrn,
)
from uberduck_ml_dev.data_loader import (
TextAudioSpeakerLoader,
TextAudioSpeakerCollate,
DistributedBucketSampler,
)
from uberduck_ml_dev.vendor.tfcompat.hparam import HParams
from uberduck_ml_dev.utils.plot import save_figure_to_numpy, plot_spectrogram
from uberduck_ml_dev.utils.utils import slice_segments, clip_grad_value_
```
# Losses
```
# export
def feature_loss(fmap_r, fmap_g):
loss = 0
for dr, dg in zip(fmap_r, fmap_g):
for rl, gl in zip(dr, dg):
rl = rl.float().detach()
gl = gl.float()
loss += torch.mean(torch.abs(rl - gl))
return loss * 2
def discriminator_loss(disc_real_outputs, disc_generated_outputs):
loss = 0
r_losses = []
g_losses = []
for dr, dg in zip(disc_real_outputs, disc_generated_outputs):
dr = dr.float()
dg = dg.float()
r_loss = torch.mean((1 - dr) ** 2)
g_loss = torch.mean(dg**2)
loss += r_loss + g_loss
r_losses.append(r_loss.item())
g_losses.append(g_loss.item())
return loss, r_losses, g_losses
def generator_loss(disc_outputs):
loss = 0
gen_losses = []
for dg in disc_outputs:
dg = dg.float()
l = torch.mean((1 - dg) ** 2)
gen_losses.append(l)
loss += l
return loss, gen_losses
def kl_loss(z_p, logs_q, m_p, logs_p, z_mask):
"""
z_p, logs_q: [b, h, t_t]
m_p, logs_p: [b, h, t_t]
"""
z_p = z_p.float()
logs_q = logs_q.float()
m_p = m_p.float()
logs_p = logs_p.float()
z_mask = z_mask.float()
kl = logs_p - logs_q - 0.5
kl += 0.5 * ((z_p - m_p) ** 2) * torch.exp(-2.0 * logs_p)
kl = torch.sum(kl * z_mask)
l = kl / torch.sum(z_mask)
return l
```
# VITS Trainer
```
# export
class VITSTrainer(TTSTrainer):
REQUIRED_HPARAMS = [
"betas",
"c_kl",
"c_mel",
"eps",
"lr_decay",
"segment_size",
"training_audiopaths_and_text",
"val_audiopaths_and_text",
"warm_start_name_g",
"warm_start_name_d",
]
def __init__(self, *args, **kwargs):
super().__init__(*args, **kwargs)
self.log_interval = 10
for param in self.REQUIRED_HPARAMS:
if not hasattr(self, param):
raise Exception(f"VITSTrainer missing a required param: {param}")
self.mel_stft = MelSTFT(
device=self.device,
rank=self.rank,
padding=(self.filter_length - self.hop_length) // 2,
)
def init_distributed(self):
if not self.distributed_run:
return
if self.rank is None or self.world_size is None:
raise Exception(
"Rank and wrld size must be provided when distributed training"
)
dist.init_process_group(
"nccl",
init_method="tcp://localhost:54321",
rank=self.rank,
world_size=self.world_size,
)
torch.cuda.set_device(self.rank)
def _log_training(self, scalars, images):
print("log training placeholder...")
if self.rank != 0 or self.global_step % self.log_interval != 0:
return
for k, v in scalars.items():
pieces = k.split("_")
key = "/".join(pieces)
self.log(key, self.global_step, scalar=v)
for k, v in images.items():
pieces = k.split("_")
key = "/".join(pieces)
self.log(key, self.global_step, image=v)
def _log_validation(self):
print("log validation...")
pass
def save_checkpoint(self, checkpoint_name, model, optimizer, learning_rate, epoch):
if self.rank != 0:
return
if hasattr(model, "module"):
state_dict = model.module.state_dict()
else:
state_dict = model.state_dict()
os.makedirs(self.checkpoint_path, exist_ok=True)
torch.save(
{
"model": state_dict,
"global_step": self.global_step,
"optimizer": optimizer.state_dict(),
"learning_rate": learning_rate,
"epoch": epoch,
},
os.path.join(self.checkpoint_path, f"{checkpoint_name}.pt"),
)
def warm_start(self, net_g, net_d, optim_g, optim_d):
if not (self.warm_start_name_g and self.warm_start_name_d):
return net_g, net_d, optim_g, optim_d, 0
if self.warm_start_name_g:
checkpoint = torch.load(self.warm_start_name_g)
net_g.load_state_dict(checkpoint["model"])
optim_g.load_state_dict(checkpoint["optimizer"])
if self.warm_start_name_d:
checkpoint = torch.load(self.warm_start_name_d)
net_d.load_state_dict(checkpoint["model"])
optim_d.load_state_dict(checkpoint["optimizer"])
self.global_step = checkpoint["global_step"]
self.learning_rate = checkpoint["learning_rate"]
start_epoch = checkpoint["epoch"]
return net_g, net_d, optim_g, optim_d, start_epoch
def _batch_to_device(self, *args):
ret = []
if self.device == "cuda":
for arg in args:
arg = arg.cuda(self.rank, non_blocking=True)
ret.append(arg)
return ret
else:
return args
def _evaluate(self, generator, val_loader):
print("Validation ...")
generator.eval()
with torch.no_grad():
for batch_idx, batch in enumerate(val_loader):
(
x,
x_lengths,
spec,
spec_lengths,
y,
y_lengths,
speakers,
) = self._batch_to_device(*batch)
x = x[:1]
x_lengths = x_lengths[:1]
spec = spec[:1]
spec_lengths[:1]
y = y[:1]
y_lengths = y_lengths[:1]
speakers = speakers[:1]
break
if self.distributed_run:
y_hat, attn, mask, *_ = generator.module.infer(
x, x_lengths, speakers, max_len=1000
)
else:
y_hat, attn, mask, *_ = generator.infer(
x, x_lengths, speakers, max_len=1000
)
y_hat_lengths = mask.sum([1, 2]).long() * self.hparams.hop_length
mel = self.mel_stft.spec_to_mel(spec)
y_hat_mel = self.mel_stft.mel_spectrogram(y_hat.squeeze(1).float())
self.log(
"Val/mel_gen",
self.global_step,
image=save_figure_to_numpy(plot_spectrogram(y_hat_mel[0].data.cpu())),
)
self.log(
"Val/mel_gt",
self.global_step,
image=save_figure_to_numpy(plot_spectrogram(mel[0].data.cpu())),
)
self.log(
"Val/audio_gen", self.global_step, audio=y_hat[0, :, : y_hat_lengths[0]]
)
self.log("Val/audio_gt", self.global_step, audio=y[0, :, : y_lengths[0]])
generator.train()
def _train_and_evaluate(
self, epoch, nets, optims, schedulers, scaler: GradScaler, loaders
):
net_g, net_d = nets
optim_g, optim_d = optims
scheduler_g, scheduler_d = schedulers
train_loader, val_loader = loaders
train_loader.batch_sampler.set_epoch(epoch)
net_g.train()
net_d.train()
# TODO (zach): remove when you want to.
# self._evaluate(net_g, val_loader)
for batch_idx, batch in enumerate(train_loader):
print(f"global step: {self.global_step}")
print(f"batch idx: {batch_idx}")
(
x,
x_lengths,
spec,
spec_lengths,
y,
y_lengths,
speakers,
) = self._batch_to_device(*batch)
with autocast(enabled=self.fp16_run):
(
y_hat,
l_length,
attn,
ids_slice,
x_mask,
z_mask,
(z, z_p, m_p, logs_p, m_q, logs_q),
) = net_g(x, x_lengths, spec, spec_lengths, speakers)
mel = self.mel_stft.spec_to_mel(spec)
# NOTE(zach): slight difference from the original VITS
# implementation due to padding differences in the spectrograms
y_mel = slice_segments(
mel, ids_slice, self.segment_size // self.hop_length
)
y_hat_mel = self.mel_stft.mel_spectrogram(y_hat.squeeze(1))
y = slice_segments(y, ids_slice * self.hop_length, self.segment_size)
# Discriminator
y_d_hat_r, y_d_hat_g, _, _ = net_d(y, y_hat.detach())
with autocast(enabled=False):
loss_disc, losses_disc_r, losses_disc_g = discriminator_loss(
y_d_hat_r, y_d_hat_g
)
loss_disc_all = loss_disc
optim_d.zero_grad()
scaler.scale(loss_disc_all).backward()
scaler.unscale_(optim_d)
scaler.step(optim_d)
with autocast(enabled=self.fp16_run):
# Generator
y_d_hat_r, y_d_hat_g, fmap_r, fmap_g = net_d(y, y_hat)
with autocast(enabled=False):
loss_dur = torch.sum(l_length.float())
loss_mel = F.l1_loss(y_mel, y_hat_mel) * self.c_mel
loss_kl = kl_loss(z_p, logs_q, m_p, logs_p, z_mask) * self.c_kl
loss_fm = feature_loss(fmap_r, fmap_g)
loss_gen, losses_gen = generator_loss(y_d_hat_g)
loss_gen_all = loss_gen + loss_fm + loss_mel + loss_dur + loss_kl
optim_g.zero_grad()
scaler.scale(loss_gen_all).backward()
scaler.unscale_(optim_g)
scaler.step(optim_g)
scaler.update()
if self.rank == 0 and self.global_step % self.log_interval == 0:
grad_norm_g = clip_grad_value_(net_g.parameters(), None)
grad_norm_d = clip_grad_value_(net_d.parameters(), None)
self._log_training(
scalars=dict(
loss_g_total=loss_gen_all,
loss_d_total=loss_disc_all,
gradnorm_d=grad_norm_d,
gradnorm_g=grad_norm_g,
loss_g_fm=loss_fm,
loss_g_dur=loss_dur,
loss_g_mel=loss_mel,
loss_g_kl=loss_kl,
),
images=dict(
slice_mel_org=save_figure_to_numpy(
plot_spectrogram(y_mel[0].data.cpu())
),
slice_mel_gen=save_figure_to_numpy(
plot_spectrogram(y_hat_mel[0].data.cpu())
),
all_mel=save_figure_to_numpy(
plot_spectrogram(mel[0].data.cpu())
),
all_attn=save_figure_to_numpy(
plot_attention(attn[0, 0].data.cpu())
),
),
)
self.global_step += 1
if self.rank == 0:
self._evaluate(net_g, val_loader)
def train(self):
if self.distributed_run:
self.init_distributed()
train_dataset = TextAudioSpeakerLoader(
self.training_audiopaths_and_text,
self.hparams,
debug=self.debug,
debug_dataset_size=self.debug_dataset_size,
)
train_sampler = DistributedBucketSampler(
train_dataset,
self.batch_size,
[32, 300, 400, 500, 600, 700, 800, 900, 1000],
num_replicas=self.world_size,
rank=self.rank,
shuffle=True,
)
collate_fn = TextAudioSpeakerCollate()
train_loader = DataLoader(
train_dataset,
num_workers=0,
shuffle=False,
pin_memory=True,
collate_fn=collate_fn,
batch_sampler=train_sampler,
)
val_dataset, val_loader = None, None
if self.rank == 0:
val_dataset = TextAudioSpeakerLoader(
self.val_audiopaths_and_text,
self.hparams,
debug=self.debug,
debug_dataset_size=self.debug_dataset_size,
)
val_loader = DataLoader(
val_dataset,
num_workers=0,
shuffle=False,
batch_size=self.batch_size,
pin_memory=True,
drop_last=False,
collate_fn=collate_fn,
)
model_kwargs = {k: v for k, v in DEFAULTS.values().items() if hasattr(self, k)}
net_g = SynthesizerTrn(
len(symbols_with_ipa),
self.filter_length // 2 + 1,
self.segment_size // self.hop_length,
n_speakers=self.n_speakers,
**model_kwargs,
)
net_d = MultiPeriodDiscriminator(self.use_spectral_norm)
if self.device == "cuda":
net_g = net_g.cuda(self.rank)
net_d = net_d.cuda(self.rank)
optim_g = torch.optim.AdamW(
net_g.parameters(),
self.learning_rate,
betas=self.betas,
eps=self.eps,
)
optim_d = torch.optim.AdamW(
net_d.parameters(), self.learning_rate, betas=self.betas, eps=self.eps
)
start_epoch = 0
net_g, net_d, optim_g, optim_d, start_epoch = self.warm_start(
net_g,
net_d,
optim_g,
optim_d,
)
if self.distributed_run:
net_g = DDP(net_g, device_ids=[self.rank])
net_d = DDP(net_d, device_ids=[self.rank])
scheduler_g = ExponentialLR(
optim_g, gamma=self.lr_decay, last_epoch=start_epoch - 1
)
scheduler_d = ExponentialLR(
optim_d, gamma=self.lr_decay, last_epoch=start_epoch - 1
)
scaler = GradScaler(enabled=self.fp16_run)
for epoch in range(start_epoch, self.epochs):
self._train_and_evaluate(
epoch,
[net_g, net_d],
[optim_g, optim_d],
[scheduler_g, scheduler_d],
scaler,
[train_loader, val_loader],
)
if epoch % self.epochs_per_checkpoint == 0:
self.save_checkpoint(
f"{self.checkpoint_name}_G_{self.global_step}",
net_g,
optim_g,
self.learning_rate,
epoch,
)
self.save_checkpoint(
f"{self.checkpoint_name}_D_{self.global_step}",
net_d,
optim_d,
self.learning_rate,
epoch,
)
```
| github_jupyter |
```
!pip install --upgrade tables
!pip install eli5
!pip install xgboost
import pandas as pd
import numpy as np
from sklearn.dummy import DummyRegressor
from sklearn.tree import DecisionTreeRegressor
from sklearn.ensemble import RandomForestRegressor
import xgboost as xgb
from sklearn.metrics import mean_absolute_error as mae
from sklearn.model_selection import cross_val_score, KFold
import eli5
from eli5.sklearn import PermutationImportance
cd "/content/drive/My Drive/Colab Notebooks/dw_matrix/matrix_two/dw_matrix_cargb-"
```
Wczytujemy nasz zbiór
```
df = pd.read_hdf('data/car.h5')
df.shape
```
Feature Engineering
```
SUFFIX_CAT = '__cat'
for feat in df.columns:
if isinstance(df[feat][0], list): continue
factorized_values = df[feat].factorize()[0]
if SUFFIX_CAT in feat:
df[feat] = factorized_values
else:
df[feat + SUFFIX_CAT] = factorized_values
cat_feats = [x for x in df.columns if SUFFIX_CAT in x]
cat_feats = [x for x in cat_feats if 'price' not in x]
len(cat_feats)
X = df[cat_feats].values
y = df['price_value'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
np.mean(scores), np.std(scores)
```
Usprawniamy
```
def run_model(model, feats):
X = df[feats].values
y = df['price_value'].values
scores = cross_val_score(model, X, y, cv=3, scoring='neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
```
Uruchamiamy Model
```
run_model( DecisionTreeRegressor(max_depth=5), cat_feats)
```
Uruchamiamy model Random Forest
```
model = RandomForestRegressor(max_depth=5, n_estimators=50, random_state=0)
run_model(model, cat_feats)
```
XGBost
```
xgb_params = {
'max_depth': 5,
'n_estimators': 50,
'leraning_rate': 0.1,
'seed': 0
}
run_model(xgb.XGBRegressor(**xgb_params), cat_feats)
```
Zobaczmy które cechy według xgb są istotne
```
m = xgb.XGBRegressor(max_depth=5, n_estimators=50, learning_rate=0.1, seed=0)
m.fit(X, y)
imp = PermutationImportance(m, random_state=0).fit(X, y)
eli5.show_weights(imp, feature_names=cat_feats)
len(cat_feats)
feats = [ 'param_napęd__cat','param_rok-produkcji__cat','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat', 'param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
```
Sprawdzamy model dla 20 cech
```
run_model(xgb.XGBRegressor(**xgb_params), feats)
```
Analizujemy kolumny
```
df['param_napęd'].unique()
df['param_rok-produkcji'].unique()
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x))
feats = [ 'param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc__cat','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat', 'param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
df['param_moc'] = df['param_moc'].map(lambda x: -1 if str(x) == 'None' else int(x.split(' ')[0]))
df['param_rok-produkcji'] = df['param_rok-produkcji'].map(lambda x: -1 if str(x) == 'None' else int(x))
feats = [ 'param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa__cat','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat', 'param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
run_model(xgb.XGBRegressor(**xgb_params), feats)
```
Sprawdzamy 'parametr_pojemność_skokowa'
```
#df['param_pojemność-skokowa'].unique()
```
Sa warości - liczby
```
feats = [ 'param_napęd__cat','param_rok-produkcji','param_stan__cat','param_skrzynia-biegów__cat','param_faktura-vat__cat','param_moc','param_marka-pojazdu__cat','feature_kamera-cofania__cat','param_typ__cat','param_pojemność-skokowa','seller_name__cat','feature_wspomaganie-kierownicy__cat','param_model-pojazdu__cat', 'param_wersja__cat','param_kod-silnika__cat','feature_system-start-stop__cat','feature_asystent-pasa-ruchu__cat','feature_czujniki-parkowania-przednie__cat','feature_łopatki-zmiany-biegów__cat','feature_regulowane-zawieszenie__cat']
df['param_pojemność-skokowa'] = df['param_pojemność-skokowa'].map(lambda x: -1 if str(x) == 'None' else int( str(x).split('cm')[0].replace(' ', '')))
run_model(xgb.XGBRegressor(**xgb_params), feats)
```
| github_jupyter |
# Robust Wireless Sensing using Probabilistic and Statistical Assessments :Artifact - Interactive Demo
<img src="./overview2.jpg" width="100%" style="float:left" />
## Step 1. Training the underlying model and use it to predict test samples.
## Step 2. Training an anomaly detector for each class according to the training samples.
## Step 3. Evaluate the ability of the underlying model to predict the test sample.
## Step 4. Performance of the RISE.
## Preliminaries
This interactive Jupyter notebook provides an example to show the performance of RISE.
## Instructions for Experimental Workflow:
Select each cell in turn and use “Cell” > “Run Cell” from the menu to run specific cells. Note that some cells depend on previous cells being executed. If any errors occur, ensure all previous cells have been executed.
## Important Notes
#### Some cells can take a few minutes to complete; please wait for the results until step to the next cell.
## Step 1. Performance of the underlying model without RISE
Here we provide the results of each experimental environment for each case study.
You can change the scene names at two places in each piece of code to get results for different scenarios.
```
#Import different data sets
##-------------------------------Case study 1: Gesture recognition----------------------------------------##
#### For WiG, you can change S1 S2 S3 S4 S5
filepath_AR = '/root/RISE-Version2/Jupyter/WiG_test/S1/'
from WiG_test.S1.test_start import start
x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# ### For WiAG, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/WiAG_test/S4/'
# from WiAG_test.S4.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For WiAG_C, you can change L1 L2 L3 L4 L5
# filepath_AR = '/root/RISE-Version2/Jupyter/WiAG_C_test/L5/'
# from WiAG_C_test.L5.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For WiAG_O, you can change L1 L2 L3 L4 L5
# filepath_AR = '/root/RISE-Version2/Jupyter/WiAG_O_test/L4/'
# from WiAG_O_test.L4.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For TACT, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/TACT_test/S1/'
# from TACT_test.S1.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For AllSee, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/AllSee_test/S5/'
# from AllSee_test.S5.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For EI, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/EI_test/S2/'
# from EI_test.S2.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start(filepath_AR)
##-------------------------------Case study 2: Gait recognition----------------------------------------##
# #### For WiWho, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/WiWho_test/S5/'
# from WiWho_test.S5.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For WifiU, you can change S1 S2 S3 S4 S5
# filepath_AR = '/root/RISE-Version2/Jupyter/WifiU_test/S5/'
# from WifiU_test.S4.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
##-------------------------------Case study 3: Activity recognition----------------------------------------##
# #### For VibWrite_R, you can change N _1 _2 _3 _4
# filepath_AR = '/root/RISE-Version2/Jupyter/VibWrite_R_test/_4/'
# from VibWrite_R_test._4.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For VibWrite_A, you can change N _1 _2 _3 _4
# filepath_AR = '/root/RISE-Version2/Jupyter/VibWrite_A_test/_4/'
# from VibWrite_A_test._4.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For Taprint_R, you can change S D M W
# filepath_AR = '/root/RISE-Version2/Jupyter/Taprint_R_test/W/'
# from Taprint_R_test.W.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For Taprint_A, you can change S D M W
# filepath_AR = '/root/RISE-Version2/Jupyter/Taprint_A_test/W/'
# from Taprint_A_test.W.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
# #### For UDO_Free, you can change RP LP DC DB
# filepath_AR = '/root/RISE-Version2/Jupyter/UDO_Free/DB/'
# from UDO_Free.DB.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_ dif,class_num,class_index = start()
# #### For M_Touch, you can change S DP T M
# filepath_AR = '/root/RISE-Version2/Jupyter/M_Touch/M/'
# from M_Touch.M.test_start import start
# x_train1, y_train1, x_test1, y_test1, myclassifier, y_true_dif, y_pred_dif,class_num,class_index = start()
```
## Step 2. Training an Anomaly Detector for each class according to the training data.
For convenience, we have saved trained anomaly detectors for each experimental environment. However, you can also run the following cell to train the anomaly detector.
```
for tt in range(class_num):
print('clf_p_prob_'+ str(tt)+'\n')
# ############################ Trainning Anomaly Detector ###############################
# from sklearn.model_selection import StratifiedKFold
# import numpy as np
# from sklearn import preprocessing
# from nonconformist.nc import MarginErrFunc
# import warnings
# warnings.filterwarnings("ignore", message="Numerical issues were encountered ")
# import sys
# sys.path.insert(0,'/root/RISE-Version2/')
# from Statistical_vector.statistical_vector import train_statistical_vector, test_statistical_vector_param, non_condition_p
# import random
# from sklearn import svm
# import joblib
# import sklearn.ensemble
# from sklearn import neighbors
# from sklearn.linear_model import LogisticRegression
# from sklearn.ensemble import AdaBoostClassifier
# from sklearn.ensemble import VotingClassifier
# ## Probability vector and statistical vector are calculated according to the training data
# skf = StratifiedKFold(n_splits=3, random_state=0,shuffle=True)
# cal_proba = np.empty(shape=[0, (class_num)*4])
# cal_score = np.empty(shape=[0, 1*4])
# train_proba = np.empty(shape=[0, (class_num)*4])
# train_nonconformity = np.empty(shape=[0, (class_num)*4])
# train_p = np.empty(shape=[0, (class_num)*4])
# cal_label = np.empty(shape=[0,1])
# train_label = np.empty(shape=[0,1])
# for train_index, cal_index in skf.split(x_train1, y_train1):
# data_train = x_train1[train_index, :]
# label_train = y_train1[train_index]
# data_cal = x_train1[cal_index, :]
# label_cal = y_train1[cal_index]
# train_p1_svm_, train_proba1_svm_ = train_statistical_vector(data_train, label_train, data_cal, label_cal,
# classification_model = myclassifier[0], non_Func = MarginErrFunc(), significance=None)
# train_p1_rf_, train_proba1_rf_ = train_statistical_vector(data_train, label_train, data_cal, label_cal,
# classification_model = myclassifier[1], non_Func = MarginErrFunc(), significance=None)
# train_p1_lr_, train_proba1_lr_ = train_statistical_vector(data_train, label_train, data_cal, label_cal,
# classification_model = myclassifier[4], non_Func = MarginErrFunc(), significance=None)
# train_p1_gbc_, train_proba1_gbc_ = train_statistical_vector(data_train, label_train, data_cal, label_cal,
# classification_model = myclassifier[5], non_Func = MarginErrFunc(), significance=None)
# train_proba1 = np.hstack((train_proba1_svm_, train_proba1_rf_, train_proba1_lr_, train_proba1_gbc_))
# train_p1 = np.hstack((train_p1_svm_, train_p1_rf_, train_p1_lr_, train_p1_gbc_))
# train_proba = np.append(train_proba, train_proba1, axis=0)
# train_p = np.append(train_p, train_p1, axis=0)
# train_label = np.append(train_label,y_train1[train_index])
# p_thr = 0.1
# rows_ = []
# for row_ in range(train_p.shape[0]):
# if (max(train_p[row_,0:0+class_num]) > p_thr) & (max(train_p[row_,class_num:class_num*1+class_num]) > p_thr) \
# & (max(train_p[row_,class_num*2:class_num*2+class_num]) > p_thr) & (max(train_p[row_,class_num*3:class_num*3+class_num]) > p_thr) :
# rows_.append(row_)
# pro_n_p = np.hstack((train_proba, train_p))
# group_it = pro_n_p[rows_,:]
# train_label1 = train_label[rows_]
# group_it = preprocessing.scale(group_it)
# index = [t for t in range(group_it.shape[0])]
# random.shuffle(index)
# group_it1 = group_it[index]
# train_p_lable1 = train_label1[index]
# ####Training an anomaly detector for each class
# ##Generalize probability vectors and statistical vectors for each class
# names = locals()
# fe_p_prob = np.hstack((train_p_lable1.reshape(-1,1),group_it1))
# for tt in range(class_num):
# names['fe_p_prob_%s'%tt] = fe_p_prob[fe_p_prob[:,0]==tt]
# names['fe_p_prob_%s'%tt] = np.delete(names['fe_p_prob_%s'%tt], 0, 1)
# print('\n--------------- Training one-class SVM model for each class -----------------\n')
# ##training one-class SVM model for each class
# for tt in range(class_num):
# names['./save_model/clf_p_prob_%s'%tt] = svm.OneClassSVM(nu=0.5,kernel="linear" )
# names['./save_model/clf_p_prob_%s'%tt].fit(names['fe_p_prob_%s'%tt])
# print('clf_p_prob_'+ str(tt)+'\n')
# ##save one-class SVM model for each class
# joblib.dump(names['./save_model/clf_p_prob_%s'%tt],'./save_model/clf_p_prob_%s'%tt+'.model')
# ##Ensemble Learning
# clf_dif1 = sklearn.ensemble.RandomForestClassifier(n_estimators=100,random_state=0)
# clf_dif2 = svm.SVC(probability = True, random_state=0)
# clf_dif3 = neighbors.KNeighborsClassifier(n_neighbors=10)
# clf_dif4 = LogisticRegression(random_state=0)
# clf_dif7 = AdaBoostClassifier(random_state=0)
# clf_pit = VotingClassifier(estimators = [('rf',clf_dif1),('svm',clf_dif2),
# ('knn',clf_dif3),('lr',clf_dif4),('AdaBoost',clf_dif7)],voting='hard')
# clf_pit.fit(group_it1,train_p_lable1)
# joblib.dump(clf_pit,'./save_model/clf_pit.model')
```
## Step 3. Evaluate the ability of the underlying model to predict the test data.
For convenience, we have saved the approval or rejection of the predicted results of the test sample by the anomaly detector. Approval is denoted by 1, rejection is denoted by -1. However, you can also run the following cell to get the result.
```
import numpy as np
accept_or_reject = np.load(filepath_AR + 'accept_or_reject.npy')
print(accept_or_reject)
# #####Calculate the probability vector and statistical vector of the test set
# from sklearn.model_selection import StratifiedShuffleSplit
# calibration_portion = 0.5
# split = StratifiedShuffleSplit(n_splits=1,
# test_size=calibration_portion)
# for train, cal in split.split(x_train1,y_train1):
# cal_scores1_svm = np.empty(cal.reshape(-1,1).shape,dtype=float)
# cal_scores1_rf = np.empty(cal.reshape(-1,1).shape,dtype=float)
# cal_scores1_lr = np.empty(cal.reshape(-1,1).shape,dtype=float)
# cal_scores1_gbc = np.empty(cal.reshape(-1,1).shape,dtype=float)
# test_svmnc1_score = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_rfnc1_score = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_lrnc1_score = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_gbcnc1_score = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_svm1_proba = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_rf1_proba = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_lr1_proba = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# test_gbc1_proba = np.empty(np.array([x_test1.shape[0],class_num]),dtype=float)
# for repeat in range(10):
# train_sample = np.random.choice(train.size, train.size, replace=True)
# data_train = x_train1[train_sample, :]
# label_train = y_train1[train_sample]
# data_cal = x_train1[cal, :]
# label_cal = y_train1[cal]
# data_test = x_test1
# cal_scores_svm, test_svmnc_score, test_svm_proba = test_statistical_vector_param(data_train, label_train, data_cal, label_cal, data_test,
# classification_model=myclassifier[0], non_Func = MarginErrFunc(), significance=None)
# cal_scores1_svm = np.hstack((cal_scores1_svm,cal_scores_svm.reshape(-1,1)))
# test_svmnc1_score = np.dstack((test_svmnc1_score,test_svmnc_score))
# test_svm1_proba = np.dstack((test_svm1_proba,test_svm_proba))
# cal_scores_rf, test_rfnc_score, test_rf_proba = test_statistical_vector_param(data_train, label_train, data_cal, label_cal, data_test,
# classification_model=myclassifier[1], non_Func = MarginErrFunc(), significance=None)
# cal_scores1_rf = np.hstack((cal_scores1_rf,cal_scores_rf.reshape(-1,1)))
# test_rfnc1_score = np.dstack((test_rfnc1_score,test_rfnc_score))
# test_rf1_proba = np.dstack((test_rf1_proba,test_rf_proba))
# cal_scores_lr, test_lrnc_score, test_lr_proba = test_statistical_vector_param(data_train, label_train, data_cal, label_cal, data_test,
# classification_model=myclassifier[4], non_Func = MarginErrFunc(), significance=None)
# cal_scores1_lr = np.hstack((cal_scores1_lr,cal_scores_lr.reshape(-1,1)))
# test_lrnc1_score = np.dstack((test_lrnc1_score,test_lrnc_score))
# test_lr1_proba = np.dstack((test_lr1_proba,test_lr_proba))
# cal_scores_gbc, test_gbcnc_score, test_gbc_proba = test_statistical_vector_param(data_train, label_train, data_cal, label_cal, data_test,
# classification_model=myclassifier[5], non_Func = MarginErrFunc(), significance=None)
# cal_scores1_gbc = np.hstack((cal_scores1_gbc,cal_scores_gbc.reshape(-1,1)))
# test_gbcnc1_score = np.dstack((test_gbcnc1_score,test_gbcnc_score))
# test_gbc1_proba = np.dstack((test_gbc1_proba,test_gbc_proba))
# ##Nonconformity score of the validation set of the test set
# cal_scores2_svm = np.mean(np.delete(cal_scores1_svm,0,1), axis=1)
# cal_scores2_rf = np.mean(np.delete(cal_scores1_rf,0,1), axis=1)
# cal_scores2_lr = np.mean(np.delete(cal_scores1_lr,0,1), axis=1)
# cal_scores2_gbc = np.mean(np.delete(cal_scores1_gbc,0,1), axis=1)
# ##Nonconformity score of the test set
# test_svmnc2_score = np.mean(np.delete(test_svmnc1_score,0,2), axis=2)
# test_rfnc2_score = np.mean(np.delete(test_rfnc1_score,0,2), axis=2)
# test_lrnc2_score = np.mean(np.delete(test_lrnc1_score,0,2), axis=2)
# test_gbcnc2_score = np.mean(np.delete(test_gbcnc1_score,0,2), axis=2)
# ##The probability vector of the test set
# test_proba_svm_ = np.mean(np.delete(test_svm1_proba,0,2), axis=2)
# test_proba_rf_ = np.mean(np.delete(test_rf1_proba,0,2), axis=2)
# test_proba_lr_ = np.mean(np.delete(test_lr1_proba,0,2), axis=2)
# test_proba_gbc_ = np.mean(np.delete(test_gbc1_proba,0,2), axis=2)
# ##The statistical vector of the test set
# test_p_svm_ = non_condition_p(cal_scores2_svm, test_svmnc2_score)
# test_p_rf_ = non_condition_p(cal_scores2_rf, test_rfnc2_score)
# test_p_lr_ = non_condition_p(cal_scores2_lr, test_lrnc2_score)
# test_p_gbc_ = non_condition_p(cal_scores2_gbc, test_gbcnc2_score)
# test_group_it = np.hstack((test_proba_svm_, test_proba_rf_, test_proba_lr_, test_proba_gbc_,
# test_p_svm_, test_p_rf_, test_p_lr_, test_p_gbc_))
# test_group_it = preprocessing.scale(test_group_it)
# ##Determine whether the prediction of the original model is correct
# discarded_sample = []
# accept_sample = []
# discarded_right_sample = []
# accept_right_sample = []
# accept_or_reject = []
# dis_test = np.empty((test_group_it.shape[0],class_num),float)
# for aa in range(class_num):
# aa_prob = joblib.load('./save_model/clf_p_prob_%s'%aa+'.model')
# dis_test[:,aa] = aa_prob.decision_function(test_group_it).flatten()
# for t in range(len(y_pred_dif)):
# result_it_d = np.argmax(dis_test[t,:])
# svmit = joblib.load('./save_model/clf_pit.model')
# result_it = svmit.predict(test_group_it[t].reshape(1,-1))
# if (y_pred_dif[t] == result_it == result_it_d):
# accept_sample.append(t)
# accept_or_reject.append(1)
# if(y_pred_dif[t] == y_true_dif[t]):
# accept_right_sample.append(t)
# else:
# discarded_sample.append(t)
# accept_or_reject.append(-1)
# if(y_pred_dif[t] == y_true_dif[t]):
# discarded_right_sample.append(t)
# print('\n--------------- Evaluate the ability of the underlying model to predict the test data -----------------\n')
# print(accept_or_reject)
```
## Step 4. The performance of the RISE.
Accuracy: The ratio of the number of correctly predicted samples to the total number of testing samples.
Precision:Of all detected drifting samples, how many are correct?
Recall:Of all drifting samples, how many are actually detected by Rise?
F1-score:A high F1-score means Rise can detect most drifting samples while rarely mis-classifying normal samples.
```
discarded_sample = np.load(filepath_AR + 'discarded_sample.npy')
discarded_right_sample = np.load(filepath_AR + 'discarded_right_sample.npy')
accept_sample = np.load(filepath_AR + 'accept_sample.npy')
accept_right_sample = np.load(filepath_AR + 'accept_right_sample.npy')
reject_num = len(discarded_sample)
reject_num_right = len(discarded_right_sample)
accept_num = len(accept_sample)
accept_num_right = len(accept_right_sample)
TP = reject_num - reject_num_right
FP = reject_num_right
FN = accept_num - accept_num_right
TN = accept_num_right
Accuracy = (TP+TN)/(TP+FP+FN+TN)
Precision = TP/(TP+FP)
Recall = TP/(TP+FN)
F1_Score = 2 * Precision * Recall / (Precision + Recall)
print('\n ------------------ The performance of the RISE --------------------\n')
print('True Positive:',TP)
print('False Positive:',FP)
print('False Negative:',FN)
print('True Negative:', TN)
print('Accuracy:',Accuracy)
print('Precision:',Precision)
print('Recall:',Recall)
print('F1_Score:', F1_Score)
```
| github_jupyter |
# Load hdf file and post-process
For a given model (as guitar_model above), set internal data using hdf5 input. Useful for post-processing only.
Prerequisite : a complete simulation saved in hdf file (e.g. results of run.py)
### Usage:
* call load_model function : create a model (dynamical system, interactions and so on) and load results from a previous simu from hdf file
* call post-processing tools
```
# Reload automatically all python modules before each cell exec
%load_ext autoreload
%autoreload 2
# standard python packages
import sys
import time
import os
import pickle
import numpy as np
from model_tools import load_model
# visu
import matplotlib.pyplot as plt
%matplotlib inline
import scipy.io
import h5py
import simulation_campaigns
import collections
import pickle
import subprocess
#pklfilename = 'campaign_1812.pkl'
pklfilename = 'campaign_2018.pkl'
#pkl_file = open('campaign_new_0612.pkl', 'rb')
pkl_file = open(pklfilename, 'rb')
remote_data = pickle.load(pkl_file)
for name in remote_data:
print("Campaign name : " + name)
if name.find('results') < 0:
print("Available freqs:")
print(list(remote_data[name].keys()))
print("\n")
```
## Select input file(s) and load model(s)
```
import scipy.io as sio
res = sio.loadmat('one_contact/guitare_obst0_frequs.mat')
sig = sio.loadmat('one_contact/guitare_obst0_amortissements.mat')
hf = sio.loadmat('one_contact/guitare_obst0_h.mat')
def load(filename):
if filename.find('bass') >= 0:
matlab_input = './bass_guitar/pb2'
elif filename.find('fretless') >= 0:
matlab_input = './fretless_bass_guitar/bsf'
elif filename.find('guitar_single') >=0 :
matlab_input = './one_contact/guitare_obst0'
elif filename.find('single') >=0 :
matlab_input = './one_contact/pb1'
m, s, f, e = load_model(filename)
frets_file = matlab_input + '_h.mat'
all_frets_positions = scipy.io.loadmat(frets_file)['h'][:, 0]
print(m.modal_values)
#if filename.find('converted') >= 0:
# m._convert[...]=False
# h5source = h5py.File(filename, 'r')
# restit = h5source.attrs['restit']
# h5source.close()
#else:
# m.convert_modal_output(s)
# restit = None
return m, s, f, all_frets_positions, e#, restit
camp = 'bass_one'
names = [name for name in remote_data.keys() if name.find(camp) >=0]
print(names)
index = 0
campaign = remote_data[names[index]]
results_path = remote_data["results_paths"][1]
fileslist = {}
for freq in campaign:
fileslist[np.float(freq)] = os.path.join(results_path, campaign[freq][5])
fileslist = collections.OrderedDict(sorted(fileslist.items()))
print(names[index])
print('select one of the available freqs:')
for freq in fileslist:
print(freq)
selected_freq = 4014080.0
print(fileslist[selected_freq])
reference_file = fileslist[selected_freq]
extra_freqs = [31360.0 ,62720.0, 125440.0, 250880.0,501760.0, 1003520.0, 2007040.0]
#5000.0, 100000.0, 10000000.0]
#extra_freqs = [fr for fr in fileslist if fr != selected_freq]#[10000.0, 10000000.0]
extra_files = {}
models, strings, frets = {}, {}, {}
guitar_model, guitar_string, guitar_frets, frets_pos, restit = load(reference_file)
for freq in extra_freqs:
models[freq], strings[freq], frets[freq], fpos, restit = load(fileslist[freq])
fig_path = 'figs_bass_onec'
if not os.path.exists(fig_path):
os.mkdir(fig_path)
```
## Plot trajectories
Use plotTraj(dynamical_system, list of dof to be plotted)
If list of dof is not set, all dof will be plotted.
Result = dof as a function of time + zoom specific ranges (as in the paper)
To save the resulting figure(s), add argument filename when calling plotTraj
Arg 'iplot' is used to change figure number. Set same iplot if you want to plot all curves on the same figure.
Default iplot=0.
```
inter = guitar_frets[0]
inter.contact_index
guitar_model.time_step
fig_path = './fig_campaign_2018/one_contact/'
print(fig_path)
# Select dof to plot traj
dof = 990
#ground = frets_pos[dof]#- guitar_string.diameter*0.5
#print(ground)
freq = guitar_model.fs
filename = os.path.join(fig_path, 'traj_' + str(freq) + '_x=' + str(dof) + '.pdf')
guitar_model.plot_traj(guitar_string, dof=dof, iplot=0, filename=filename)
#m2.plot_traj(s2, dof=dof, iplot=0)#, filename=filename)
#current_model.plot_traj(current_string, dof=dof, iplot=0)#, filename=filename)
#m3.plot_traj(s3, dof=dof, iplot=0)#, filename=filename)
#dof = 9
#ground = frets_pos[dof]#- guitar_string.diameter*0.5
#filename = os.path.join(fig_path, 'traj_' + str(freq) + '_x=' + str(dof) + '.pdf')
#guitar_model.plot_traj(guitar_string, dof=dof, iplot=1, filename=filename)#, ground=ground)
#guitar_model2.plot_traj(guitar_string2, dof=dof, iplot=0, )#, ground=ground
ticks = ['x--', '^:', 'o--', 's', '*--', '--', ':'] * 10
ticks = ['--', ':', '--', '--', '--', '--', ':'] * 10
leg =[]
traj_ref = guitar_model.data_ds[guitar_string][990,:]
plt.plot(guitar_model.time, traj_ref, ':k')
leg.append('ref')
i = 1
for freq in models:
traj = models[freq].data_ds[strings[freq]][990,:]
time = models[freq].time
#plt.plot(time, (traj-traj_ref), ticks[i])
leg.append(freq)
plt.plot(time, traj, ticks[i])
i += 1
plt.legend(leg)
plt.xlim(1.45,1.55)
plt.ylim(0.017,0.019)
filename = os.path.join(fig_path, 'traj_convergence_zoom.pdf')
plt.savefig(filename)
#plt.xlim(2.97, 3.05)
#plt.ylim(0.014, 0.02)
inter = guitar_model.data_interactions
ic = 0
time = guitar_model.time
y = inter[guitar_frets[ic]][0][:]
#ydot = inter[frets[ic]][:,1]
plt.figure()
for ic in range(len(guitar_frets)):
lamb = inter[guitar_frets[ic]][1][:]
#print(np.where(lamb >=1e-22))
plt.plot(time, lamb, 'o')
#plt.plot(time, ydot,'--x')
plt.xlim(0,0.05)
#plt.ylim(-10,10)
ind = np.where(y<0)
ind2 = np.where(y>=0)
plt.figure(44)
#plt.clf()
plt.plot(time[ind2], y[ind2], 'k:o', time[ind], y[ind], 'rx:', )
plt.xlim(0.003, 0.009)
#plt.figure()
#plt.plot(time, lamb, 'k:o')
```
# Plot modes
Use guitar_model.plot_modes(guitar_string,plot_shape)
plot_shape = (x,y) where x,y is the resolution of the grid of plots.
Example:
plot_modes(guitar_string, (4,2)) : splits time range in 8 and plots dof=f(x) for those 8 time instants on 8 figures
Set filename arg to save resulting figure in a file.
```
# Plot 10 figures on two columns, for 10 time instants
guitar_model.plot_modes(guitar_string,plot_shape=[5,2], iplot=0)
plt.savefig(os.path.join(fig_path, 'modes.pdf'))
# Same thing but save output in modes.pdf
#guitar_model.plot_modes(guitar_string,plot_shape=[5,3], filename='modes.pdf', iplot=1)
# plot modes for time[12] and time[5000] on one column
#guitar_model.plot_modes(guitar_string, times =[12, 1200], plot_shape=(2,1), filename='modes2.pdf', iplot=2)
```
## Plot contactogram
!! Works only if enable_interactions_output=True in model (guitar_model) constructor
```
guitar_model.contactogram(guitar_string, 33)
plt.savefig(os.path.join(fig_path, 'contacto.pdf'))
guitar_model.nb_time_steps_output
```
## Create a movie mode=f(time)
```
moviename = os.path.basename(reference_file)
moviename = os.path.join(fig_path, os.path.splitext(moviename)[0] + '.mp4')
print(moviename)
#moviename = 'tmp.mp4'
guitar_model.make_movie(guitar_string, moviename)
if os.path.exists(('./last.mp4')):
os.remove('./last.mp4')
os.symlink(moviename, 'last.mp4')
```
<video controls src="last.mp4" />
#%%html
#<video width="320" height="240" controls>
# <source src="./last.mp4" type="video/mp4">
#</video>
| github_jupyter |
# [deplacy](https://koichiyasuoka.github.io/deplacy/) voor syntactische analyse
## met [spaCy-Alpino](https://github.com/KoichiYasuoka/spaCy-Alpino)
```
!pip install deplacy spacy_alpino
import spacy_alpino
nlp=spacy_alpino.load()
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [Camphr-Udify](https://camphr.readthedocs.io/en/latest/notes/udify.html)
```
!pip install deplacy camphr 'unofficial-udify>=0.3.0' en-udify@https://github.com/PKSHATechnology-Research/camphr_models/releases/download/0.7.0/en_udify-0.7.tar.gz
import pkg_resources,imp
imp.reload(pkg_resources)
import spacy
nlp=spacy.load("en_udify")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [Stanza](https://stanfordnlp.github.io/stanza)
```
!pip install deplacy stanza
import stanza
stanza.download("nl")
nlp=stanza.Pipeline("nl")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [UDPipe 2](http://ufal.mff.cuni.cz/udpipe/2)
```
!pip install deplacy
def nlp(t):
import urllib.request,urllib.parse,json
with urllib.request.urlopen("https://lindat.mff.cuni.cz/services/udpipe/api/process?model=nl&tokenizer&tagger&parser&data="+urllib.parse.quote(t)) as r:
return json.loads(r.read())["result"]
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [Trankit](https://github.com/nlp-uoregon/trankit)
```
!pip install deplacy trankit transformers
import trankit
nlp=trankit.Pipeline("dutch")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [spacy-udpipe](https://github.com/TakeLab/spacy-udpipe)
```
!pip install deplacy spacy-udpipe
import spacy_udpipe
spacy_udpipe.download("nl")
nlp=spacy_udpipe.load("nl")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [Turku-neural-parser-pipeline](https://turkunlp.org/Turku-neural-parser-pipeline/)
```
!pip install deplacy ufal.udpipe configargparse 'tensorflow<2' torch==0.4.1 torchtext==0.3.1 torchvision==0.2.1
!test -d Turku-neural-parser-pipeline || git clone --depth=1 https://github.com/TurkuNLP/Turku-neural-parser-pipeline
!cd Turku-neural-parser-pipeline && git submodule update --init --recursive && test -d models_nl_alpino || python fetch_models.py nl_alpino
import sys,subprocess
nlp=lambda t:subprocess.run([sys.executable,"full_pipeline_stream.py","--gpu","-1","--conf","models_nl_alpino/pipelines.yaml"],cwd="Turku-neural-parser-pipeline",input=t,encoding="utf-8",stdout=subprocess.PIPE).stdout
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [spaCy](https://spacy.io/)
```
!pip install deplacy
!sudo pip install -U spacy
!sudo python -m spacy download nl_core_news_md
import pkg_resources,imp
imp.reload(pkg_resources)
import spacy
nlp=spacy.load("nl_core_news_md")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [spaCy-COMBO](https://github.com/KoichiYasuoka/spaCy-COMBO)
```
!pip install deplacy spacy_combo
import spacy_combo
nlp=spacy_combo.load("nl_alpino")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [NLP-Cube](https://github.com/Adobe/NLP-Cube)
```
!pip install deplacy nlpcube
from cube.api import Cube
nlp=Cube()
nlp.load("nl")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [COMBO-pytorch](https://gitlab.clarin-pl.eu/syntactic-tools/combo)
```
!pip install --index-url https://pypi.clarin-pl.eu/simple deplacy combo
import combo.predict
nlp=combo.predict.COMBO.from_pretrained("dutch-ud27")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [spaCy-jPTDP](https://github.com/KoichiYasuoka/spaCy-jPTDP)
```
!pip install deplacy spacy_jptdp
import spacy_jptdp
nlp=spacy_jptdp.load("nl_alpino")
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
## met [Frog](https://languagemachines.github.io/frog)
```
!sudo apt install frog frogdata
!pip install deplacy
def nlp(t):
import subprocess
u=""
for s in subprocess.check_output(["frog"],input=t.encode("utf-8")).decode("utf-8").split("\n"):
t=s.split("\t")
u+=s+"\n" if len(t)!=10 else "\t".join([t[0],t[1],t[2],t[4].split("(")[0],t[4],"_",t[8],t[9],"_","_"])+"\n"
return u
doc=nlp("Toch houd ik ze vast, ondanks alles, omdat ik nog steeds aan de innerlijke goedheid van den mens geloof.")
import deplacy
deplacy.render(doc)
deplacy.serve(doc,port=None)
# import graphviz
# graphviz.Source(deplacy.dot(doc))
```
| github_jupyter |
```
import datetime
lipidname = "PXSM"
tail = "TCC CCCCCC"
link = "A A"
head = "C P"
description = "; A general model sphingomyelin (SM) lipid \n; C16:0 palmitic acid , C24:0 lignoceroyl \n"
modeledOn="; This topology follows the standard Martini 2.0 lipid definitions and building block rules.\n; Reference(s): \n; S.J. Marrink, A.H. de Vries, A.E. Mark. Coarse grained model for semi-quantitative lipid simulations. JPC-B, 108:750-760, \n; 2004. doi:10.1021/jp036508g \n; S.J. Marrink, H.J. Risselada, S. Yefimov, D.P. Tieleman, A.H. de Vries. The MARTINI force field: coarse grained model for \n; biomolecular simulations. JPC-B, 111:7812-7824, 2007. doi:10.1021/jp071097f \n; T.A. Wassenaar, H.I. Ingolfsson, R.A. Bockmann, D.P. Tieleman, S.J. Marrink. Computational lipidomics with insane: a versatile \n; tool for generating custom membranes for molecular simulations. JCTC, 150410125128004, 2015. doi:10.1021/acs.jctc.5b00209\n; Created: "
now = datetime.datetime.now()
membrane="testmembrane"
insane="../insane+SF.py"
mdparams="../test.mdp"
martinipath="../martini.ff/"
ITPCatalogue="./epithelial.cat"
ITPMasterFile="martini_v2_epithelial.itp"
modeledOn+= now.strftime("%Y.%m.%d")+"\n"
# Cleaning up intermediate files from previous runs
!rm -f *#*
!rm -f *step*
!rm -f {membrane}*
import fileinput
import os.path
print("Create itp")
!python {martinipath}/lipid-martini-itp-v06.py -o {lipidname}.itp -alname {lipidname} -name {lipidname} -alhead '{head}' -allink '{link}' -altail '{tail}'
#update description and parameters
with fileinput.FileInput(lipidname+".itp", inplace=True) as file:
for line in file:
if line == "; This is a ...\n":
print(description, end='')
elif line == "; Was modeled on ...\n":
print(modeledOn, end='')
else:
print(line, end='')
#Add this ITP file to the catalogue file
if not os.path.exists(ITPCatalogue):
ITPCatalogueData = []
else:
with open(ITPCatalogue, 'r') as file :
ITPCatalogueData = file.read().splitlines()
ITPCatalogueData = [x for x in ITPCatalogueData if not x==lipidname+".itp"]
ITPCatalogueData.append(lipidname+".itp")
with open(ITPCatalogue, 'w') as file :
file.writelines("%s\n" % item for item in ITPCatalogueData)
#build ITPFile
with open(martinipath+ITPMasterFile, 'w') as masterfile:
for ITPfilename in ITPCatalogueData:
with open(ITPfilename, 'r') as ITPfile :
for line in ITPfile:
masterfile.write(line)
print("Done")
# build a simple membrane to visualize this species
!python2 {insane} -o {membrane}.gro -p {membrane}.top -d 0 -x 3 -y 3 -z 3 -sol PW -center -charge 0 -orient -u {lipidname}:1 -l {lipidname}:1 -itpPath {martinipath}
import os #Operating system specific commands
import re #Regular expression library
print("Test")
print("Grompp")
grompp = !gmx grompp -f {mdparams} -c {membrane}.gro -p {membrane}.top -o {membrane}.tpr
success=True
for line in grompp:
if re.search("ERROR", line):
success=False
if re.search("Fatal error", line):
success=False
#if not success:
print(line)
if success:
print("Run")
!export GMX_MAXCONSTRWARN=-1
!export GMX_SUPPRESS_DUMP=1
run = !gmx mdrun -v -deffnm {membrane}
summary=""
logfile = membrane+".log"
if not os.path.exists(logfile):
print("no log file")
print("== === ====")
for line in run:
print(line)
else:
try:
file = open(logfile, "r")
fe = False
for line in file:
if fe:
success=False
summary=line
elif re.search("^Steepest Descents.*converge", line):
success=True
summary=line
break
elif re.search("Fatal error", line):
fe = True
except IOError as exc:
sucess=False;
summary=exc;
if success:
print("Success")
else:
print(summary)
```
| github_jupyter |
# Seldon Core Real Time Stream Processing with KNative Eventing
In this example we will show how you can enable real time stream processing in Seldon Core by leveraging the KNative Eventing integration.
In this example we will deploy a simple model containerised with Seldon Core and we will leverage the basic Seldon Core integration with KNative Eventing which will allow us to connect it so it can receive cloud events as requests and return a cloudevent-enabled response which can be collected by other components.
## Pre-requisites
You will require the following in order to go ahead:
* Istio 1.42+ Installed ([Documentation Instructions](https://istio.io/latest/docs/setup/install/))
* KNative Eventing 0.13 installed ([Documentation Instructions](https://knative.dev/docs/admin/install/))
* Seldon Core v1.1+ installed with Istio Ingress Enabled ([Documentation Instructions](https://docs.seldon.io/projects/seldon-core/en/latest/workflow/install.html#ingress-support))
## Deploy your Seldon Model
We will first deploy our model using Seldon Core. In this case we'll use one of the [pre-packaged model servers](https://docs.seldon.io/projects/seldon-core/en/latest/servers/overview.html).
We first createa configuration file:
```
%%writefile ./assets/simple-iris-deployment.yaml
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: iris-deployment
spec:
predictors:
- graph:
implementation: SKLEARN_SERVER
modelUri: gs://seldon-models/v1.10.0-dev/sklearn/iris
name: simple-iris-model
children: []
name: default
replicas: 1
```
### Run the model in our cluster
Now we run the Seldon Deployment configuration file we just created.
```
!kubectl apply -f assets/simple-iris-deployment.yaml
```
### Check that the model has been deployed
```
!kubectl get pods | grep iris
```
## Create a Trigger to reach our model
We want to create a trigger that is able to reach directly to the service.
We will be using the following seldon deployment:
```
!kubectl get sdep | grep iris
```
### Create trigger configuration
```
%%writefile ./assets/seldon-knative-trigger.yaml
apiVersion: eventing.knative.dev/v1beta1
kind: Trigger
metadata:
name: seldon-eventing-sklearn-trigger
spec:
broker: default
filter:
attributes:
type: seldon.iris-deployment.default.request
subscriber:
ref:
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
name: iris-deployment
```
Create this trigger file which will send all cloudevents of type `"seldon.<deploymentName>.request"`.
```
!kubectl apply -f assets/seldon-knative-trigger.yaml
```
CHeck that the trigger is working correctly (you should see "Ready: True"), together with the URL that will be reached.
```
!kubectl get trigger
```
### Send a request to the KNative Eventing default broker
To send requests we can do so by sending a curl command from a pod inside of the cluster.
```
!kubectl run --quiet=true -it --rm curl --image=radial/busyboxplus:curl --restart=Never -- \
curl -v "default-broker.default.svc.cluster.local" \
-H "Ce-Id: 536808d3-88be-4077-9d7a-a3f162705f79" \
-H "Ce-specversion: 0.3" \
-H "Ce-Type: seldon.iris-deployment.default.request" \
-H "Ce-Source: seldon.examples.streaming.curl" \
-H "Content-Type: application/json" \
-d '{"data": { "ndarray": [[1,2,3,4]]}}'
```
### Check our model has received it
We can do this by checking the logs (we can query the logs through the service name) and see that the request has been processed
```
!kubectl logs svc/iris-deployment-default simple-iris-model | tail -6
```
## Connect a source to listen to the results of the seldon model
Our Seldon Model is producing results which are sent back to KNative.
This means that we can connect other subsequent services through a trigger that filters for those response cloudevents.
### First create the service that willl print the results
This is just a simple pod that prints all the request data into the console.
```
%%writefile ./assets/event-display-deployment.yaml
# event-display app deploment
apiVersion: apps/v1
kind: Deployment
metadata:
name: event-display
spec:
replicas: 1
selector:
matchLabels: &labels
app: event-display
template:
metadata:
labels: *labels
spec:
containers:
- name: helloworld-python
image: gcr.io/knative-releases/github.com/knative/eventing-sources/cmd/event_display
---
# Service that exposes event-display app.
# This will be the subscriber for the Trigger
kind: Service
apiVersion: v1
metadata:
name: event-display
spec:
selector:
app: event-display
ports:
- protocol: TCP
port: 80
targetPort: 8080
```
### Now run the event display resources
```
!kubectl apply -f assets/event-display-deployment.yaml
```
### Check that the event display has been deployed
```
!kubectl get pods | grep event
```
### Create trigger for event display
We now can create a trigger that sends all the requests of the type and source created by the seldon deployment to our event display pod
```
%%writefile ./assets/event-display-trigger.yaml
# Trigger to send events to service above
apiVersion: eventing.knative.dev/v1alpha1
kind: Trigger
metadata:
name: event-display
spec:
broker: default
filter:
attributes:
type: seldon.iris-deployment.default.response
source: seldon.iris-deployment
subscriber:
ref:
apiVersion: v1
kind: Service
name: event-display
```
### Apply that trigger
```
!kubectl apply -f assets/event-display-trigger.yaml
```
### Check our triggers are correctly set up
We now should see the event trigger available.
```
!kubectl get trigger
```
## Send a couple of requests more
We can use the same process we outlined above to send a couple more events.
```
!kubectl run --quiet=true -it --rm curl --image=radial/busyboxplus:curl --restart=Never -- \
curl -v "default-broker.default.svc.cluster.local" \
-H "Ce-Id: 536808d3-88be-4077-9d7a-a3f162705f79" \
-H "Ce-Specversion: 0.3" \
-H "Ce-Type: seldon.iris-deployment.default.request" \
-H "Ce-Source: dev.knative.samples/helloworldsource" \
-H "Content-Type: application/json" \
-d '{"data": { "ndarray": [[1,2,3,4]]}}'
```
### Visualise the requests that come from the service
```
!kubectl logs svc/event-display | tail -40
```
| github_jupyter |
<a href="https://colab.research.google.com/github/agayev169/pytorch_examples/blob/master/dcgan.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
## GPU
```
!nvidia-smi
```
## Imports
```
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
from torchvision import datasets, transforms
import matplotlib.pyplot as plt
import numpy as np
import time
```
## Constants
```
device = "cuda" if torch.cuda.is_available() else "cpu"
batch_size = 128
epochs_n = 20
lr = 0.0002
```
## Model
### Discriminator
```
class Discriminator(nn.Module):
def __init__(self):
super(Discriminator, self).__init__()
self.conv1 = nn.Conv2d(1, 64, 4, 2, 1)
self.conv2 = nn.Conv2d(64, 128, 4, 2, 1)
self.bn2 = nn.BatchNorm2d(128)
self.conv3 = nn.Conv2d(128, 256, 4, 2, 1)
self.bn3 = nn.BatchNorm2d(256)
self.conv4 = nn.Conv2d(256, 1, 4, 2, 1)
def forward(self, x):
x = self.conv1(x) # x, 64, 14, 14
x = F.relu(x)
x = self.conv2(x) # x, 128, 7, 7
x = self.bn2(x)
x = F.relu(x)
x = self.conv3(x) # x, 256, 3, 3
x = self.bn3(x)
x = F.relu(x)
x = self.conv4(x) # x, 1, 1, 1
x = torch.sigmoid(x)
return x.view(-1, 1).squeeze(1)
```
### Generator
```
class Generator(nn.Module):
def __init__(self):
super(Generator, self).__init__()
self.conv1_t = nn.ConvTranspose2d(100, 512, 5, 1, 0)
self.bn1 = nn.BatchNorm2d(512)
self.conv2_t = nn.ConvTranspose2d(512, 256, 4, 2, 0)
self.bn2 = nn.BatchNorm2d(256)
self.conv3_t = nn.ConvTranspose2d(256, 128, 4, 2, 0)
self.bn3 = nn.BatchNorm2d(128)
self.conv4_t = nn.ConvTranspose2d(128, 1, 3, 1, 0)
def forward(self, x): # x: x, 100, 1, 1
x = self.conv1_t(x) # x, 256, 5, 5
x = self.bn1(x)
x = F.relu(x)
x = self.conv2_t(x) # x, 128, 12, 12
x = self.bn2(x)
x = F.relu(x)
x = self.conv3_t(x) # x, 64, 26, 26
x = self.bn3(x)
x = F.relu(x)
x = self.conv4_t(x) # x, 1, 28, 28
x = torch.tanh(x)
return x
```
### Weights initialization
```
def weights_init(m):
classname = m.__class__.__name__
if classname.find('Conv') != -1:
nn.init.normal_(m.weight.data, 0.0, 0.02)
elif classname.find('BatchNorm') != -1:
nn.init.normal_(m.weight.data, 1.0, 0.02)
nn.init.constant_(m.bias.data, 0)
```
### Test
### Train
```
def train(generator, discriminator, device, train_loader, opt_gen, opt_disc, epochs_n, criterion=None, criterion_sum=None):
if not criterion:
criterion = nn.BCELoss()
criterion_sum = nn.BCELoss(reduction="sum")
fixed_noise = torch.normal(mean=0, std=1, size=(5, 100, 1, 1)).to(device)
results = np.empty(shape=(epochs_n + 1, 5, 28, 28))
results[0] = generator(fixed_noise).to("cpu").detach().numpy().reshape(5, 28, 28)
for epoch in range(epochs_n):
epoch_begin = time.time()
total_loss_gen = 0
total_loss_disc = 0
D_x = 0
D_z = 0
for x, _ in train_loader:
fake_labels = torch.zeros(x.size()[0]).to(device)
real_labels = torch.ones (x.size()[0]).to(device)
# generator
opt_gen.zero_grad()
z = torch.normal(mean=0, std=1, size=(x.size()[0], 100, 1, 1)).to(device)
gen_out = generator(z)
disc_out = discriminator(gen_out)
D_z += np.sum(disc_out.to("cpu").detach().numpy())
loss_gen = criterion(disc_out, real_labels)
total_loss_gen += criterion_sum(disc_out, real_labels).item()
loss_gen.backward()
opt_gen.step()
# discriminator
opt_disc.zero_grad()
x = x.to(device)
disc_out_real = discriminator(x)
D_x += np.sum(disc_out_real.to("cpu").detach().numpy())
loss_disc_real = criterion(disc_out_real, real_labels)
total_loss_disc += criterion_sum(disc_out_real, real_labels).item()
z = torch.normal(mean=0, std=1, size=(x.size()[0], 100, 1, 1)).to(device)
gen_out = generator(z)
disc_out_fake = discriminator(gen_out.detach())
loss_disc_fake = criterion(disc_out_fake, fake_labels)
total_loss_disc += criterion_sum(disc_out_fake, fake_labels).item()
disc_total_loss = loss_disc_fake + loss_disc_real
disc_total_loss.backward()
opt_disc.step()
D_z /= len(train_loader.dataset)
D_x /= len(train_loader.dataset)
total_loss_gen = total_loss_gen / len(train_loader.dataset)
total_loss_disc = total_loss_disc / len(train_loader.dataset)
results[epoch + 1] = generator(fixed_noise).to("cpu").detach().numpy().reshape(5, 28, 28)
epoch_time = time.time() - epoch_begin
print(f"TRAIN --- epoch: {(epoch + 1):3d} --- generator loss: {total_loss_gen:.6f} --- discriminator loss: {total_loss_disc:.6f} --- D_x: {D_x:.3f} --- D_z: {D_z:.3f} --- time: {epoch_time:.4f}")
epoch += 1
return results
```
## Data
```
train_loader = torch.utils.data.DataLoader(
datasets.MNIST("data/", train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5,), (0.5,))
])),
batch_size=batch_size, shuffle=True
)
```
## Training
```
generator = Generator().to(device)
discriminator = Discriminator().to(device)
generator.train()
discriminator.train()
generator.apply(weights_init)
discriminator.apply(weights_init)
opt_gen = optim.Adam( generator.parameters(), lr=lr, betas=(0.5, 0.999))
opt_disc = optim.Adam(discriminator.parameters(), lr=lr, betas=(0.5, 0.999))
imgs = train(generator, discriminator, device, train_loader, opt_gen, opt_disc, epochs_n)
```
## Generating images
```
fig, ax = plt.subplots(len(imgs), 5, figsize=(15, len(imgs) * 3))
for i in range(len(imgs) * 5):
ax[i // 5, i % 5].imshow(imgs[i // 5][i % 5], cmap="gray")
ax[i // 5, i % 5].axis("off")
z = torch.normal(0, 1, size=(25, 100, 1, 1)).to(device)
out = generator(z).to("cpu").detach().numpy().reshape(-1, 28, 28)
print(discriminator(generator(z)))
fig, ax = plt.subplots(5, 5, figsize=(15, 15))
for i in range(25):
ax[i // 5, i % 5].imshow(out[i], cmap="gray")
ax[i // 5, i % 5].axis("off")
```
| github_jupyter |
# Python Module for Amazon Web Services Price Information
This notebook illustrates the capabilities of a lightweight Python module for accessing the Amazon Web Services (AWS) price lists.
The classes available in the module follow the hierarchy of the AWS price information:
* `AWSOffersIndex` class represents the AWS offer index information. This is the entry point as it lists all the supported AWS services for which price information is available.
* `AWSOffer` class represents price information for one, specific, AWS service. This is called an _offer_ in Amazon's pricing parlance.
* One AWS offer can contain many products. They are represented with the `AWSProduct` class.
* Pricing information for each offer's product is represented with the `AWSProductPricing` class.
* Product pricing data is given in pricing tiers and they are represented with the `AWSProductPriceTier` class.
Import all the classes that represent varous types of AWS price information.
```
from aws_price_list import *
```
## AWS Offer Index
Create an object that represents the AWS Offer Index. This command will attempt to HTTP `GET` the JSON file with information about all the available AWS offers (these would be _services_ like EC2 or S3).
```
oi = AWSOffersIndex()
```
Available information about the AWS offer index:
```
oi.format
oi.published
oi.disclaimer
oi.accessed
oi.endpoint
```
Having all time-related attributes as `datetime.datetime` objects makes it possible to print date-time information in various string formats or perform date-time computations easily.
```
oi.accessed.strftime('%c %Z')
```
The list of AWS offers (services) for which price information is available:
```
oi.offers
```
The offer index information can be reloaded:
```
oi.reload()
oi.accessed.strftime('%c %z')
```
## AWS Offer
The `offer()` method accesses the price information about the specific offer (service). This command will HTTP `GET` the JSON file with the prices and terms of the specified AWS offer:
```
offer = oi.offer('AmazonS3')
```
Various information about the offer's prices and terms:
```
offer.format
offer.disclaimer
offer.published
offer.accessed
offer.endpoint
offer.code
offer.version
```
Listing of all available products for the AWS offer:
```
offer.products
```
That is quite a lot! Exact number:
```
len(offer.products)
```
Price information for offer's products can come under different _terms_. To find out which terms are available:
```
offer.terms
```
The offer information can be reloaded:
```
offer.reload()
offer.accessed
```
## AWS Product
To get price information for a product, use the `product()` method with the product's SKU and one of the offer's terms. The default term, when not specified, is `OnDemand`.
If there is no product information for the requested term a `ValueError` exception is raised.
```
prod = offer.product('WP9ANXZGBYYSGJEA')
```
Various product information:
```
prod.sku
prod.family
prod.attributes
```
All attributes are available in a dictionary so to access any of them is simple:
```
prod.attributes['location']
prod.term_type
```
## AWS Product Pricing
```
pricing = prod.pricing
pricing
```
Various product pricing information:
```
pricing[0].code
pricing[0].product_sku
pricing[0].attributes
pricing[0].effective_from
```
To calculate the price for a given amount (unit conversion not supported):
```
pricing[0].get_price(458.64)
```
The unit of the calculated price above:
```
pricing[0].price_unit
```
## AWS Product Pricing Tier
The most atomic pricing information about an AWS offer's product is accessed via:
```
tiers = pricing[0].tiers
tiers
```
Various price tier information:
```
for t in tiers:
print(t.rate_code)
for t in tiers:
print(t.description)
for t in tiers:
print(t.applies_to)
```
The pricing tiers are always sorted on the tier's begin range value:
```
for t in tiers:
print('from: {}, to: {}, price: ${} {}'
.format(t.begin_range, t.end_range, t.price, t.unit))
```
| github_jupyter |
**This notebook is an exercise in the [Data Cleaning](https://www.kaggle.com/learn/data-cleaning) course. You can reference the tutorial at [this link](https://www.kaggle.com/alexisbcook/parsing-dates).**
---
In this exercise, you'll apply what you learned in the **Parsing dates** tutorial.
# Setup
The questions below will give you feedback on your work. Run the following cell to set up the feedback system.
```
from learntools.core import binder
binder.bind(globals())
from learntools.data_cleaning.ex3 import *
print("Setup Complete")
```
# Get our environment set up
The first thing we'll need to do is load in the libraries and dataset we'll be using. We'll be working with a dataset containing information on earthquakes that occured between 1965 and 2016.
```
# modules we'll use
import pandas as pd
import numpy as np
import seaborn as sns
import datetime
# read in our data
earthquakes = pd.read_csv("../input/earthquake-database/database.csv")
# set seed for reproducibility
np.random.seed(0)
```
# 1) Check the data type of our date column
You'll be working with the "Date" column from the `earthquakes` dataframe. Investigate this column now: does it look like it contains dates? What is the dtype of the column?
```
# TODO: Your code here!
earthquakes.dtypes
```
Once you have answered the question above, run the code cell below to get credit for your work.
```
# Check your answer (Run this code cell to receive credit!)
q1.check()
# Line below will give you a hint
#q1.hint()
```
# 2) Convert our date columns to datetime
Most of the entries in the "Date" column follow the same format: "month/day/four-digit year". However, the entry at index 3378 follows a completely different pattern. Run the code cell below to see this.
```
earthquakes[3378:3383]
```
This does appear to be an issue with data entry: ideally, all entries in the column have the same format. We can get an idea of how widespread this issue is by checking the length of each entry in the "Date" column.
```
date_lengths = earthquakes.Date.str.len()
date_lengths.value_counts()
```
Looks like there are two more rows that has a date in a different format. Run the code cell below to obtain the indices corresponding to those rows and print the data.
```
indices = np.where([date_lengths == 24])[1]
print('Indices with corrupted data:', indices)
earthquakes.loc[indices]
```
Given all of this information, it's your turn to create a new column "date_parsed" in the `earthquakes` dataset that has correctly parsed dates in it.
**Note**: When completing this problem, you are allowed to (but are not required to) amend the entries in the "Date" and "Time" columns. Do not remove any rows from the dataset.
```
# TODO: Your code here
earthquakes.loc[3378, 'Date'] = '02/23/1975'
earthquakes.loc[7512, 'Date'] = '04/28/1985'
earthquakes.loc[20650, 'Date'] = '03/13/2011'
earthquakes['date_parsed'] = pd.to_datetime(earthquakes.Date, format='%m/%d/%Y')
# Check your answer
q2.check()
# Lines below will give you a hint or solution code
#q2.hint()
#q2.solution()
```
# 3) Select the day of the month
Create a Pandas Series `day_of_month_earthquakes` containing the day of the month from the "date_parsed" column.
```
# try to get the day of the month from the date column
day_of_month_earthquakes = earthquakes.date_parsed.dt.day
# Check your answer
q3.check()
# Lines below will give you a hint or solution code
#q3.hint()
#q3.solution()
```
# 4) Plot the day of the month to check the date parsing
Plot the days of the month from your earthquake dataset.
```
# TODO: Your code here!
sns.histplot(day_of_month_earthquakes, kde=False)
```
Does the graph make sense to you?
```
# Check your answer (Run this code cell to receive credit!)
q4.check()
# Line below will give you a hint
#q4.hint()
```
# (Optional) Bonus Challenge
For an extra challenge, you'll work with a [Smithsonian dataset](https://www.kaggle.com/smithsonian/volcanic-eruptions) that documents Earth's volcanoes and their eruptive history over the past 10,000 years
Run the next code cell to load the data.
```
volcanos = pd.read_csv("../input/volcanic-eruptions/database.csv")
```
Try parsing the column "Last Known Eruption" from the `volcanos` dataframe. This column contains a mixture of text ("Unknown") and years both before the common era (BCE, also known as BC) and in the common era (CE, also known as AD).
```
volcanos['Last Known Eruption'].sample(5)
```
# (Optional) More practice
If you're interested in graphing time series, [check out this tutorial](https://www.kaggle.com/residentmario/time-series-plotting-optional).
You can also look into passing columns that you know have dates in them the `parse_dates` argument in `read_csv`. (The documention [is here](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.read_csv.html).) Do note that this method can be very slow, but depending on your needs it may sometimes be handy to use.
# Keep going
In the next lesson, learn how to [**work with character encodings**](https://www.kaggle.com/alexisbcook/character-encodings).
| github_jupyter |
# Algorithm for force plate calibration
Marcos Duarte
This notebook demonstrates the algorithm for force plate calibration proposed by Cedraro et al. (2008, 2009).
A force plate (FP) is an electromechanical device that measures the components of the vectors force $(\mathbf{F} = [F_X,\, F_Y,\, F_Z])$ and moment of force or torque $(\mathbf{M} = [M_X,\, M_Y,\, M_Z])$ applied to the it. The FP is composed by a transducer that transforms a mechanical deformation to an electrical signal usually using strain gauges or piezoelectric sensors. The transformation from electrical signals (input) to force and moment of force (output) as a function of time in a six-component FP usually is given by the following linear relationship:
$$
\mathbf{L}(t) = \mathbf{C}\mathbf{V}(t)
$$
Where $\mathbf{L}(t)$ is the force plate output vector $([\mathbf{F}(t), \mathbf{M}(t)]^T)$, in N and Nm, $\mathbf{V}(t)$ is the vector of electrical signals (six voltage signals, in V) and $\mathbf{C}$ is known as the six-by-six (constant) calibration matrix (in N/V or Nm/V). Note that we used the term vector here to refer to an uni-dimensional matrix (usual in scientific computing), which is different from vector/scalar concept in Mechanics.
The expansion of the former equiation at a given instant is:
$$
\begin{bmatrix}
F_x \\ F_y \\ F_z \\ M_x \\ M_y \\ M_z
\end{bmatrix}\, = \,
\begin{bmatrix}
C_{11} && C_{12} && C_{13} && C_{14} && C_{15} && C_{16} \\
C_{21} && C_{22} && C_{23} && C_{24} && C_{25} && C_{26} \\
C_{31} && C_{32} && C_{33} && C_{34} && C_{35} && C_{36} \\
C_{41} && C_{42} && C_{43} && C_{44} && C_{45} && C_{46} \\
C_{51} && C_{52} && C_{53} && C_{54} && C_{55} && C_{56} \\
C_{61} && C_{62} && C_{63} && C_{64} && C_{65} && C_{66}
\end{bmatrix}\,
\begin{bmatrix}
V_1 \\ V_2 \\ V_3 \\ V_4 \\ V_5 \\ V_6
\end{bmatrix}
$$
The terms off-diagonal are known as the crosstalk terms and represent the effect of a load applied in one direction on the other direction. For a FP with none or small crosstalk, the off-diagonal terms are zero or very small compared to the main-diagonal terms. Note that the equation above is in fact a system of six linear independent equations with six unknowns each (where $V_1 ... V_6$ are the measured inputs):
\begin{cases}
F_x &=& C_{11}V_1 + C_{12}V_2 + C_{13}V_3 + C_{14}V_4 + C_{15}V_5 + C_{16}V_6 \\
F_y &=& C_{21}V_1 + C_{22}V_2 + C_{23}V_3 + C_{24}V_4 + C_{25}V_5 + C_{26}V_6 \\
F_z &=& C_{31}V_1 + C_{32}V_2 + C_{33}V_3 + C_{34}V_4 + C_{35}V_5 + C_{36}V_6 \\
M_x &=& C_{41}V_1 + C_{42}V_2 + C_{43}V_3 + C_{44}V_4 + C_{45}V_5 + C_{46}V_6 \\
M_y &=& C_{51}V_1 + C_{52}V_2 + C_{53}V_3 + C_{54}V_4 + C_{55}V_5 + C_{56}V_6 \\
M_z &=& C_{61}V_1 + C_{62}V_2 + C_{63}V_3 + C_{64}V_4 + C_{65}V_5 + C_{66}V_6
\end{cases}
Of course, an important aspect of the FP functionning is that it should be calibrated, i.e., the calibration matrix must be known and accurate (it comes with the force plate when you buy one). Cedraro et al. (2008) proposed a method for in situ re-calibration of FP and their algorithm is presented next.
## Algorithm
Consider that in a re-calibration procedure we apply on the FP known forces, $\mathbf{F}_I = [F_{X_I},\, F_{Y_I},\, F_{Z_I}]^T$, at known places, $\mathbf{COP} = [X_{COP},\, Y_{COP},\, Z_{COP}]$ (the center of pressure coordinates in the FP reference frame).
The moments of forces, $\mathbf{M}_I = [M_{X_I},\, M_{Y_I},\, M_{Z_I}]^T$, due to these forces can be found using the equation $\mathbf{M}_I = \mathbf{COP} \times \mathbf{F}_I$, which can be expressed in matrix form as:
$$
\mathbf{M}_I =
\begin{bmatrix}
0 && -Z_{COP} && Y_{COP} \\
Z_{COP} && 0 && -X_{COP} \\
-Y_{COP} && X_{COP} && 0
\end{bmatrix}\, \mathbf{F}_I \, = \, \mathbf{A}_{COP}\mathbf{F}_I
$$
$\mathbf{A}_{COP}$ (a [skew-symmetric matrix](https://en.wikipedia.org/wiki/Skew-symmetric_matrix)) is simply the COP position in matrix form in order to calculate the [cross product with matrix multiplication](https://en.wikipedia.org/wiki/Cross_product).
These known loads on the FP can also be represented as:
$$
\mathbf{L}_I =
\begin{bmatrix}
\mathbf{F}_I \\
\mathbf{M}_I
\end{bmatrix}
$$
### Linear re-calibration
For a linear re-calibration, the relationship between the measured FP output, $\mathbf{L}$, and the known loads, $\mathbf{L}_I$, is approximated by a linear equation:
$$ \mathbf{L}_I = \mathbf{C}\mathbf{L} + \mathbf{E} $$
Where $\mathbf{C}$ now is the six-by-six re-calibration matrix (with dimensionless units) and $\mathbf{E}$ is a gaussian, uncorrelated, zero mean
noise six-by-one matrix.
The re-calibration matrix can be found by solving the equation above and then $\mathbf{C}$ can be later used to re-calibrate the FP output:
$$ \mathbf{L}_C = \mathbf{C}\mathbf{L} $$
Where $\mathbf{L}_C$ is the re-calibrated FP output. For a perfectly calibrated FP, $\mathbf{L}_C = \mathbf{L}$ and $\mathbf{C} = \mathbf{I}$, the six-by-six identity matrix.
Cedraro et al. (2008, 2009) proposed to use a calibrated three-component load cell (LC) to measure the loads $\mathbf{F}_I(t)$ applied on the FP at $k$ known measurements sites. The LC measures the loads in its own coordinate system $(xyz)$: $\mathbf{F}_{LC}(t) = [F_x(t),\, F_y(t),\, F_z(t)]^T$, which is probaly rotated (by an unknown value, represented by rotation matrix $\mathbf{R}^k$) in relation to the FP coordinate system (the coordinate systems are also translated to each other but the translation is known and given by the COP position).
For each measurement site, the equation for the determination of the re-calibration matrix will be given by:
$$ \mathbf{P}^k\mathbf{R}^k\mathbf{F}^k_{LC}(t)= \mathbf{P}^k\mathbf{F}_I^k(t) = \mathbf{C}\mathbf{L}^k(t) + \mathbf{E}^k(t) \quad k = 1, ..., n $$
Where:
$$
\mathbf{P}^k =
\begin{bmatrix}
\mathbf{I}_3 \\
\mathbf{A}_{COP}
\end{bmatrix}
$$
and $I_3$ is the three-by-three identity matrix.
Using a typical load cell, with a flat bottom, on top the FP, a realistic assumption is to consider that $z$ of LC is aligned to $Z$ of FP (the vertical direction); in this case the rotation matrix is:
$$
\mathbf{R}^k =
\begin{bmatrix}
\cos\alpha^k && -\sin\alpha^k && 0 \\
\sin\alpha^k && \cos\alpha^k && 0 \\
0 && 0 && 1
\end{bmatrix}
$$
Cedraro et al. (2008) propose the following algorithm to estimate $\mathbf{C}$:
1. The misalignments, $\alpha^k$, are initialized: $\mathbf{\alpha} = [\alpha^1, \cdots, \alpha^n]$;
2. $\mathbf{C}$ is calculated by a least-squares approach;
3. The residual errors are estimated as: $\mathbf{E}^k(t) = \mathbf{P}^k\mathbf{R}^k\mathbf{F}^k_{LC}(t) - \mathbf{C}\mathbf{L}^k(t)$;
4. The increment $\mathbf{\Delta\alpha}$ is calculated by minimizing the cost function $\sum_{k,t}\mathbf{E}^k(t)^T\mathbf{E}^k(t)$, assuming dimensional unitary weights;
5. The parameters are updated: $\mathbf{\alpha} = \mathbf{\alpha} + \mathbf{\Delta\alpha}$.
The iteration of steps 2–5 stops when each $\Delta\alpha^k < \varepsilon_0$, where $\varepsilon_0=10^{-10}$ is the chosen threshold.
### Simulation
Let's simulate some data to test this calibration procedure. Cedraro et al. (2008) employed sinusoids, cosenoids, and ramps as sintetic signals to simulate the calibration process:
```
import numpy as np
from numpy.linalg import inv
import matplotlib.pyplot as plt
%matplotlib inline
import seaborn as sns
sns.set_context("notebook", font_scale=1.4,
rc={"lines.linewidth": 3, "lines.markersize": 8, "axes.titlesize": 'x-large'})
# simulated forces measured by the load cell in its local coordinate system
samples = np.linspace(1, 6000, 6000)
ns = samples.shape[0]
Flc = np.array([100*np.sin(5*2*np.pi*samples/ns) + 2*np.random.randn(6000),
100*np.cos(5*2*np.pi*samples/ns) + 2*np.random.randn(6000),
samples/15 + 200 + 5*np.random.randn(6000)])
# plots
fig, axs = plt.subplots(3, 1, figsize=(8, 5), sharex='all')
axs[0].plot(samples, Flc[0])
axs[0].set_ylabel('Fx (N)')
axs[0].locator_params(axis='y', nbins=3)
axs[0].yaxis.set_label_coords(-.08, 0.5)
axs[1].plot(samples, Flc[1])
axs[1].set_ylabel('Fy (N)')
axs[1].locator_params(axis='y', nbins=3)
axs[1].yaxis.set_label_coords(-.08, 0.5)
axs[2].plot(samples, Flc[2])
axs[2].set_ylabel('Fz (N)')
axs[2].set_xlabel('Samples')
axs[2].locator_params(axis='y', nbins=3)
axs[2].yaxis.set_label_coords(-.08, 0.5)
plt.tight_layout(pad=.5, h_pad=.025)
plt.show()
```
And Cedraro et al. (2008) also proposed five measurement sites and a re-calibration matrix for the simulated re-calibration:
```
# simulated true re-calibration matrix
C = np.array([[ 1.0354, -0.0053, -0.0021, -0.0289, -0.0402, 0.0081],
[ 0.0064, 1.0309, -0.0031, 0.0211, 0.0135, -0.0001],
[ 0.0000, -0.0004, 1.0022, -0.0005, -0.0182, 0.0300],
[-0.0012, -0.0385, 0.0002, 0.9328, 0.0007, 0.0017],
[ 0.0347, 0.0003, 0.0008, -0.0002, 0.9325, -0.0024],
[-0.0004, -0.0013, -0.0003, -0.0023, 0.0035, 1.0592]])
# five k measurements sites (in m)
COP = np.array([[ 0, 112, 112, -112, -112],
[ 0, 192, -192, 192, -192],
[-124, -124, -124, -124, -124]])/1000
# number of sites
nk = COP.shape[1]
# function for the COP skew-symmetric matrix
Acop = lambda x,y,z : np.array([[.0, -z, y], [z, .0, -x], [-y, x, .0]])
# same simulated forces measured by the load cell in all sites
Flc = np.tile(Flc, nk)
```
Let's generate the loads measured by the FP given the re-calibration matrix and the simulated forces measured by the load cell (we will consider no rotation for now). For that we will have to solve the equation:
$$ \mathbf{L}_I = \mathbf{C}\mathbf{L} $$
Which is:
$$ \mathbf{L} = \mathbf{C}^{-1}\mathbf{L}_I $$
$\mathbf{C}$ is a square (6-by-6) matrix and the computation of its inverse is straightforward.
```
# simulated loads measured by LC
Li = np.empty((6, ns*nk))
P = np.empty((6, 3, nk))
for k, cop in enumerate(COP.T):
P[:, :, k] = np.vstack((np.eye(3), Acop(*cop)))
Li[:, k*ns:(k+1)*ns] = P[:, :, k] @ Flc[:, k*ns:(k+1)*ns]
# simulated loads applied on FP
L = inv(C) @ Li
```
In the calculations above we took advantage of the [new operator for matrix multiplcation in Python 3](https://www.python.org/dev/peps/pep-0465/): `@` (mnemonic: `@` is `*` for mATrices).
We can now simulate the re-calibration procedure by determining the re-calibration matrix using these loads. Of course, the re-calibration matrix to be determined should be equal to the simulated re-calibration matrix we started with, but this is the fun of the simulation - we know where we want to go.
The re-calibration matrix can be found by solving the following equation (considering the angles equal zero for now):
$$ \mathbf{L}_I = \mathbf{C}\mathbf{L} $$
$$ \mathbf{L}_I \mathbf{L}^{-1} = \mathbf{C}\mathbf{L} \mathbf{L}^{-1} = \mathbf{C}\mathbf{I}$$
$$ \mathbf{C} = \mathbf{L}_I\mathbf{L}^{-1} $$
The problem is that $\mathbf{L}$ in general is a non-square matrix and its inverse is not defined (unless you perform exactly six measurements and then $\mathbf{L}$ would be a six-by-six square matrix, but this is too restrictive). However, we still can solve the equation with some extra manipulation:
$$ \mathbf{L}_I = \mathbf{C}\mathbf{L} $$
$$ \mathbf{L}_I \mathbf{L}^T = \mathbf{C}\mathbf{L} \mathbf{L}^T $$
$$ \mathbf{L}_I \mathbf{L}^T(\mathbf{L}\mathbf{L}^T)^{-1} = \mathbf{C}\mathbf{L} \mathbf{L}^T (\mathbf{L}\mathbf{L}^T)^{-1} = \mathbf{C}\mathbf{I} $$
$$ \mathbf{C} = \mathbf{L}_I\mathbf{L}^T(\mathbf{L}\mathbf{L}^T)^{-1} $$
Note that $\mathbf{L} \mathbf{L}^T$ is a square matrix and is invertible (also [nonsingular](https://en.wikipedia.org/wiki/Invertible_matrix)) if $\mathbf{L}$ is L.I. ([linearly independent rows/columns](https://en.wikipedia.org/wiki/Linear_independence)). The matrix $\mathbf{L}^T(\mathbf{L}\mathbf{L}^T)^{-1}$ is known as the [generalized inverse or Moore–Penrose pseudoinverse](https://en.wikipedia.org/wiki/Moore%E2%80%93Penrose_pseudoinverse), a generalization of the inverse matrix. If we denote this pseudoinverse matrix by $\mathbf{L}^+$, we can state the solution of the equation simply as:
$$ \mathbf{L}_I = \mathbf{C}\mathbf{L} $$
$$ \mathbf{C} = \mathbf{L}_I \mathbf{L}^+ $$
To compute the Moore–Penrose pseudoinverse, we could calculate it by the naive approach in Python:
```python
from numpy.linalg import inv
Linv = L.T @ inv(L @ L.T)
```
But both Numpy and Scipy have functions to calculate the pseudoinverse, which might give greater numerical stability (but read [Inverses and pseudoinverses. Numerical issues, speed, symmetry](http://vene.ro/blog/inverses-pseudoinverses-numerical-issues-speed-symmetry.html)).
Of note, [numpy.linalg.pinv](http://docs.scipy.org/doc/numpy/reference/generated/numpy.linalg.pinv.html) calculates the pseudoinverse of a matrix using its singular-value decomposition (SVD) and including all large singular values (using the [LAPACK (Linear Algebra Package)](https://en.wikipedia.org/wiki/LAPACK) routine gesdd), whereas [scipy.linalg.pinv](http://docs.scipy.org/doc/scipy/reference/generated/scipy.linalg.pinv.html#scipy.linalg.pinv) calculates a pseudoinverse of a matrix using a least-squares solver (using the LAPACK method gelsd) and [scipy.linalg.pinv2](http://docs.scipy.org/doc/scipy/reference/generated/scipy.linalg.pinv2.html) also uses SVD to find the pseudoinverse (also using the LAPACK routine gesdd).
Let's use [scipy.linalg.pinv2](http://docs.scipy.org/doc/scipy/reference/generated/scipy.linalg.pinv2.html):
```
from scipy.linalg import pinv2
Lpinv = pinv2(L)
```
Then, the re-calibration matrix is:
```
C2 = Li @ Lpinv
```
Which is indeed the same as the initial calibration matrix:
```
np.allclose(C, C2)
```
The residual error between the old loads and new loads after re-calibration is:
```
E = Li - C2 @ L
e = np.sum(E * E)
print('Average residual error between old and new loads:', e)
```
### Optimization
Let's now implement the full algorithm considering the likely rotation of the load cell during a re-calibration.
The idea is to guess initial values for the angles, estmate the re-calibration matrix, estimate new values for the angles that minimize the equation for the residuals and then estimate again the re-calibration matrix in an iterative approach until the estimated angles converge to the actual angles of the load cell in the different sites. This is a typical problem of [optimization](https://en.wikipedia.org/wiki/Mathematical_optimization) where the angles are the design variables and the equation for the residuals is the cost function (see this [notebook about optimization](http://nbviewer.jupyter.org/github/demotu/BMC/blob/master/notebooks/Optimization.ipynb)).
Let's code the optimization in a complete function for the force plate re-calibration, named `fpcalibra.py`, with the following signature:
```python
C, ang = fpcalibra(Lfp, Flc, COP, threshold=1e-10)
```
Let's import this function and run its example:
```
import sys
sys.path.insert(1, r'./../functions') # add to pythonpath
from fpcalibra import fpcalibra
>>> import numpy as np
>>> from numpy.linalg import inv
>>>
>>> # simulated true re-calibration matrix
>>> C = np.array([[ 1.0354, -0.0053, -0.0021, -0.0289, -0.0402, 0.0081],
>>> [ 0.0064, 1.0309, -0.0031, 0.0211, 0.0135, -0.0001],
>>> [ 0.0000, -0.0004, 1.0022, -0.0005, -0.0182, 0.0300],
>>> [-0.0012, -0.0385, 0.0002, 0.9328, 0.0007, 0.0017],
>>> [ 0.0347, 0.0003, 0.0008, -0.0002, 0.9325, -0.0024],
>>> [-0.0004, -0.0013, -0.0003, -0.0023, 0.0035, 1.0592]])
>>> # simulated 5 measurements sites (in m)
>>> COP = np.array([[ 0, 112, 112, -112, -112],
>>> [ 0, 192, -192, 192, -192],
>>> [-124, -124, -124, -124, -124]])/1000
>>> nk = COP.shape[1]
>>> # simulated forces measured by the load cell (in N) before rotation
>>> samples = np.linspace(1, 6000, 6000)
>>> ns = samples.shape[0]
>>> Flc = np.empty((3, nk*ns))
>>> for k in range(nk):
>>> Flc[:, k*ns:(k+1)*ns] = np.array([100*np.sin(5*2*np.pi*samples/ns) + 2*np.random.randn(ns),
>>> 100*np.cos(5*2*np.pi*samples/ns) + 2*np.random.randn(ns),
>>> samples/15 + 200 + 5*np.random.randn(ns)])
>>> # function for the COP skew-symmetric matrix
>>> Acop = lambda x,y,z : np.array([[.0, -z, y], [z, .0, -x], [-y, x, .0]])
>>> # simulated loads measured by the force plate
>>> Li = np.empty((6, ns*nk))
>>> P = np.empty((6, 3, nk))
>>> for k, cop in enumerate(COP.T):
>>> P[:, :, k] = np.vstack((np.eye(3), Acop(*cop)))
>>> Li[:, k*ns:(k+1)*ns] = P[:, :, k] @ Flc[:, k*ns:(k+1)*ns]
>>> Lfp = inv(C) @ Li
>>> # simulated angles of rotaton of the measurement sites
>>> ang = np.array([20, -10, 0, 15, -5])/180*np.pi
>>> # function for the rotation matrix
>>> R = lambda a : np.array([[np.cos(a), -np.sin(a), 0], [np.sin(a), np.cos(a), 0], [ 0, 0, 1]])
>>> # simulated forces measured by the load cell after rotation
>>> for k in range(nk):
>>> Flc[:, k*ns:(k+1)*ns] = R(ang[k]).T @ Flc[:, k*ns:(k+1)*ns]
>>>
>>> C2, ang2 = fpcalibra(Lfp, Flc, COP)
>>>
>>> e = np.sqrt(np.sum(C2-C)**2)
>>> print('Residual between simulated and optimal re-calibration matrices:', e)
>>> e = np.sqrt(np.sum(ang2-ang)**2)
>>> print('Residual between simulated and optimal rotation angles:', e)
```
The simulation works as expected and the function was able to estimate accurately the known initial re-calibration matrix and angles of rotation.
## Non-linear algorithm for force plate calibration
Cappello et al. (2011) extended the algorithm described earlier and proposed an algorithm for non-linear re-calibration of FPs.
The idea is that a load applied on the FP produces bending which depends on the point of force application and in turn will result in systematic errors in the COP determination. Consequently, this non-linearity could be modeled and compensated with a re-calibration which takes into account the COP coordinates measured by the FP and added to the linear re-calibration we deduced above (Cappello et al., 2011).
The re-calibration equation will be (Cappello et al., 2011):
$$ \begin{array}{l l}
\mathbf{L}_C = \mathbf{C}_0\mathbf{L} + \,
\begin{bmatrix}
C_{x_{11}} & C_{x_{12}} & 0 & C_{x_{14}} & C_{x_{15}} & C_{x_{16}} \\
C_{x_{21}} & C_{x_{22}} & 0 & C_{x_{24}} & C_{x_{25}} & C_{x_{26}} \\
C_{x_{31}} & C_{x_{32}} & 0 & C_{x_{34}} & C_{x_{35}} & C_{x_{36}} \\
C_{x_{41}} & C_{x_{42}} & 0 & C_{x_{44}} & C_{x_{45}} & C_{x_{46}} \\
C_{x_{51}} & C_{x_{52}} & 0 & C_{x_{54}} & C_{x_{55}} & C_{x_{56}} \\
C_{x_{61}} & C_{x_{62}} & 0 & C_{x_{64}} & C_{x_{65}} & C_{x_{66}}
\end{bmatrix}\,
\begin{bmatrix}
F_x \\ F_y \\ F_z \\ M_x \\ M_y \\ M_z
\end{bmatrix} COP_x +
\begin{bmatrix}
C_{y_{11}} & C_{y_{12}} & 0 & 0 & C_{y_{15}} & C_{y_{16}} \\
C_{y_{21}} & C_{y_{22}} & 0 & 0 & C_{y_{25}} & C_{y_{26}} \\
C_{y_{31}} & C_{y_{32}} & 0 & 0 & C_{y_{35}} & C_{y_{36}} \\
C_{y_{41}} & C_{y_{42}} & 0 & 0 & C_{y_{45}} & C_{y_{46}} \\
C_{y_{51}} & C_{y_{52}} & 0 & 0 & C_{y_{55}} & C_{y_{56}} \\
C_{y_{61}} & C_{y_{62}} & 0 & 0 & C_{y_{65}} & C_{y_{66}}
\end{bmatrix}\,
\begin{bmatrix}
F_x \\ F_y \\ F_z \\ M_x \\ M_y \\ M_z
\end{bmatrix} COP_y
\\[6pt]
\mathbf{L}_C = (\mathbf{C}_0 + \mathbf{C}_x COP_x + \mathbf{C}_y COP_y)\mathbf{L} = \mathbf{C}_{NL}\mathbf{L}
\end{array} $$
Where $\mathbf{C}_0$ is the linear re-calibration matrix, $\mathbf{L}$ is the measured FP output, $\mathbf{C}_x$ and $\mathbf{C}_y$ are the non-linear re-calibration matrices.
To estimate $\mathbf{C}_{NL}$, Cappello et al. (2011) suggest to employ the algorithm proposed by Cedraro et al. (2008) to estimate the linear re-calibration described earlier.
```
# number of sites
nk = COP.shape[1]
# number of samples
ns = int(Lfp.shape[1]/nk)
# function for the COP skew-symmetric matrix
Acop = lambda x,y,z : np.array([[.0, -z, y], [z, .0, -x], [-y, x, .0]])
P = np.empty((6, 3, nk))
for k, cop in enumerate(COP.T):
P[:, :, k] = np.vstack((np.eye(3), Acop(*cop)))
# function for the 2D rotation matrix
R = lambda a : np.array([[np.cos(a), -np.sin(a), 0], [np.sin(a), np.cos(a), 0], [ 0, 0, 1]])
# Pseudoiverse of the loads measured by the force plate
if method.lower() == 'svd':
Lpinv = pinv2(Lfp)
else:
Lpinv = pinv(Lfp)
# cost function for the optimization
def costfun(ang, P, R, Flc, CLfp, nk, ns, E):
for k in range(nk):
E[:,k*ns:(k+1)*ns] = (P[:,:,k] @ R(ang[k])) @ Flc[:,k*ns:(k+1)*ns] - CLfp[:,k*ns:(k+1)*ns]
return np.sum(E * E)
# inequality constraints
bnds = [(-np.pi/2, np.pi/2) for k in range(nk)]
# some initialization
ang0 = np.zeros(nk)
E = np.empty((6, ns*nk))
da = []
delta_ang = 10*threshold
Li = np.empty((6, ns*nk))
start = time.time()
# the optimization
while np.all(delta_ang > threshold):
for k in range(nk):
Li[:,k*ns:(k+1)*ns] = (P[:,:,k] @ R(ang0[k])) @ Flc[:,k*ns:(k+1)*ns]
C = Li @ Lpinv
CLfp = C @ Lfp
res = minimize(fun=costfun, x0=ang0, args=(P, R, Flc, CLfp, nk, ns, E),
bounds=bnds, method='TNC', options={'disp': False})
delta_ang = np.abs(res.x - ang0)
ang0 = res.x
da.append(delta_ang.sum())
tdelta = time.time() - start
print('\nOptimization finished in %.1f s after %d steps.\n' %(tdelta, len(da)))
print('Optimal calibration matrix:\n', C)
print('\nOptimal angles:\n', res.x*180/np.pi)
print('\n')
return C, res.x
```
## References
- [Cedraro A, Cappello A, Chiari L (2008) A portable system for in-situ re-calibration of force platforms: theoretical validation. Gait & Posture, 28, 488–494](http://www.ncbi.nlm.nih.gov/pubmed/18450453).
- [Cedraro A, Cappello A, Chiari L (2009) A portable system for in-situ re-calibration of force platforms: experimental validation. Gait & Posture, 29, 449–453](http://www.ncbi.nlm.nih.gov/pubmed/19111467).
- [Cappello A, Bagala F, Cedraro A, Chiari L (2011) Non-linear re-calibration of force platforms. Gait & Posture, 33, 724–726](http://www.ncbi.nlm.nih.gov/pubmed/21392999).
```
%load_ext version_information
%version_information numpy, scipy, matplotlib, ipython, jupyter, pandas
```
## Function fpcalibra.py
```
# %load ./../functions/fpcalibra.py
"""Force plate calibration algorithm.
"""
__author__ = 'Marcos Duarte, https://github.com/demotu/BMC'
__version__ = 'fpcalibra.py v.1.0.1 2016/08/19'
__license__ = "MIT"
import numpy as np
from scipy.linalg import pinv, pinv2
from scipy.optimize import minimize
import time
def fpcalibra(Lfp, Flc, COP, threshold=1e-10, method='SVD'):
"""Force plate calibration algorithm.
For a force plate (FP) re-calibration, the relationship between the
measured FP output (L) and the known loads (Li) is approximated by:
Li = C@L + E (@ is the operator for matrix multiplication).
Where C is the 6-by-6 re-calibration matrix and E is a gaussian,
uncorrelated, zero mean noise six-by-one matrix.
The re-calibration matrix can be found by solving the equation above and
then C can be later used to re-calibrate the FP output: Lc = C@L.
Where Lc is the re-calibrated FP output.
Cedraro et al. (2008) [1]_ proposed to use a calibrated three-component
load cell to measure the forces applied on the FP at known measurement
sites and an algorithm for the re-calibration.
This code implements the re-calibration algorithm, see [2]_
Parameters
----------
Lfp : numpy 2-D array (6, nsamples*nksites)
loads [Fx, Fy, Fz, Mx, My, Mz] (in N and Nm) measured by the force
plate due to the corresponding forces applied at the measurement sites
Flc : numpy 2-D array (3, nsamples*nksites)
forces [Fx, Fy, Fz] (in N) measured by the load cell at the
measurement sites
COP : numpy 2-D array (3, nksites)
positions [COPx, COPy, COPz] (in m) of the load cell at the
measurement sites
threshold : float, optional
threshold to stop the optimization (default 1e-10)
method : string, optional
method for the pseudiinverse calculation, 'SVD' (default) or 'lstsq'
SVD is the Singular Value Decomposition and lstsq is least-squares
Returns
-------
C : numpy 2-D (6-by-6) array
optimal force plate re-calibration matrix (in dimensionless units)
ang : numpy 1-D array [ang0, ..., angk]
optimal angles of rotation (in rad) of the load cells at the
measurement sites
References
----------
.. [1] Cedraro A, Cappello A, Chiari L (2008) Gait & Posture, 28, 488–494.
.. [2] http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/ForcePlateCalibration.ipynb
Example
-------
>>> from fpcalibra import fpcalibra
>>> import numpy as np
>>> from numpy.linalg import inv
>>>
>>> # simulated true re-calibration matrix
>>> C = np.array([[ 1.0354, -0.0053, -0.0021, -0.0289, -0.0402, 0.0081],
>>> [ 0.0064, 1.0309, -0.0031, 0.0211, 0.0135, -0.0001],
>>> [ 0.0000, -0.0004, 1.0022, -0.0005, -0.0182, 0.0300],
>>> [-0.0012, -0.0385, 0.0002, 0.9328, 0.0007, 0.0017],
>>> [ 0.0347, 0.0003, 0.0008, -0.0002, 0.9325, -0.0024],
>>> [-0.0004, -0.0013, -0.0003, -0.0023, 0.0035, 1.0592]])
>>> # simulated 5 measurements sites (in m)
>>> COP = np.array([[ 0, 112, 112, -112, -112],
>>> [ 0, 192, -192, 192, -192],
>>> [-124, -124, -124, -124, -124]])/1000
>>> nk = COP.shape[1]
>>> # simulated forces measured by the load cell (in N) before rotation
>>> samples = np.linspace(1, 6000, 6000)
>>> ns = samples.shape[0]
>>> Flc = np.empty((3, nk*ns))
>>> for k in range(nk):
>>> Flc[:, k*ns:(k+1)*ns] = np.array([100*np.sin(5*2*np.pi*samples/ns) + 2*np.random.randn(ns),
>>> 100*np.cos(5*2*np.pi*samples/ns) + 2*np.random.randn(ns),
>>> samples/15 + 200 + 5*np.random.randn(ns)])
>>> # function for the COP skew-symmetric matrix
>>> Acop = lambda x,y,z : np.array([[.0, -z, y], [z, .0, -x], [-y, x, .0]])
>>> # simulated loads measured by the force plate
>>> Li = np.empty((6, ns*nk))
>>> P = np.empty((6, 3, nk))
>>> for k, cop in enumerate(COP.T):
>>> P[:, :, k] = np.vstack((np.eye(3), Acop(*cop)))
>>> Li[:, k*ns:(k+1)*ns] = P[:, :, k] @ Flc[:, k*ns:(k+1)*ns]
>>> Lfp = inv(C) @ Li
>>> # simulated angles of rotaton of the measurement sites
>>> ang = np.array([20, -10, 0, 15, -5])/180*np.pi
>>> # function for the rotation matrix
>>> R = lambda a : np.array([[np.cos(a), -np.sin(a), 0], [np.sin(a), np.cos(a), 0], [ 0, 0, 1]])
>>> # simulated forces measured by the load cell after rotation
>>> for k in range(nk):
>>> Flc[:, k*ns:(k+1)*ns] = R(ang[k]).T @ Flc[:, k*ns:(k+1)*ns]
>>>
>>> C2, ang2 = fpcalibra(Lfp, Flc, COP)
>>>
>>> e = np.sqrt(np.sum(C2-C)**2)
>>> print('Residual between simulated and optimal re-calibration matrices:', e)
>>> e = np.sqrt(np.sum(ang2-ang)**2)
>>> print('Residual between simulated and optimal rotation angles:', e)
"""
# number of sites
nk = COP.shape[1]
# number of samples
ns = int(Lfp.shape[1]/nk)
# function for the COP skew-symmetric matrix
Acop = lambda x,y,z : np.array([[.0, -z, y], [z, .0, -x], [-y, x, .0]])
P = np.empty((6, 3, nk))
for k, cop in enumerate(COP.T):
P[:, :, k] = np.vstack((np.eye(3), Acop(*cop)))
# function for the 2D rotation matrix
R = lambda a : np.array([[np.cos(a), -np.sin(a), 0], [np.sin(a), np.cos(a), 0], [ 0, 0, 1]])
# Pseudoiverse of the loads measured by the force plate
if method.lower() == 'svd':
Lpinv = pinv2(Lfp)
else:
Lpinv = pinv(Lfp)
# cost function for the optimization
def costfun(ang, P, R, Flc, CLfp, nk, ns, E):
for k in range(nk):
E[:,k*ns:(k+1)*ns] = (P[:,:,k] @ R(ang[k])) @ Flc[:,k*ns:(k+1)*ns] - CLfp[:,k*ns:(k+1)*ns]
return np.sum(E * E)
# inequality constraints
bnds = [(-np.pi/2, np.pi/2) for k in range(nk)]
# some initialization
ang0 = np.zeros(nk)
E = np.empty((6, ns*nk))
da = []
delta_ang = 10*threshold
Li = np.empty((6, ns*nk))
start = time.time()
# the optimization
while np.all(delta_ang > threshold):
for k in range(nk):
Li[:,k*ns:(k+1)*ns] = (P[:,:,k] @ R(ang0[k])) @ Flc[:,k*ns:(k+1)*ns]
C = Li @ Lpinv
CLfp = C @ Lfp
res = minimize(fun=costfun, x0=ang0, args=(P, R, Flc, CLfp, nk, ns, E),
bounds=bnds, method='TNC', options={'disp': False})
delta_ang = np.abs(res.x - ang0)
ang0 = res.x
da.append(delta_ang.sum())
tdelta = time.time() - start
print('\nOptimization finished in %.1f s after %d steps.\n' %(tdelta, len(da)))
print('Optimal calibration matrix:\n', C)
print('\nOptimal angles:\n', res.x*180/np.pi)
print('\n')
return C, res.x
```
| github_jupyter |
```
import csv
from bpemb import BPEmb
from cleantext import clean
from fastai.callbacks import *
from fastai.imports import torch
from fastai.text import *
import pandas as pd
torch.cuda.set_device(2)
bpemb_de = BPEmb(lang="de", vs=25000, dim=300)
# construct the vocabulary by added a padding token with the ID 25000 (because of the bpemb_de vocab size)
itos = dict(enumerate(bpemb_de.words + ['xxpad']))
voc = Vocab(itos)
def load_data(filename):
texts = []
labels = []
with open(filename) as csvfile:
# follow the 10kGNAD creator's setup
reader = csv.reader(csvfile, delimiter=';', quotechar='\'')
for row in reader:
labels.append(row[0])
texts.append(row[1])
df = pd.DataFrame({'label': labels, 'text': texts})
df['text'] = df['text'].apply(lambda x: bpemb_de.encode_ids_with_bos_eos(clean(x, lang='de')))
return df
df_train_valid = load_data("10kGNAD/train.csv")
# the last 1000 training samples are used for validation
df_train = df_train_valid.iloc[:-1000]
df_valid = df_train_valid.iloc[-1000:]
df_test = load_data("10kGNAD/test.csv")
data_lm = TextLMDataBunch.from_ids('uf_de_exp', bs=128, vocab=voc, train_ids=df_train['text'], valid_ids=df_valid['text'])
learn_lm = language_model_learner(data_lm, AWD_LSTM, drop_mult=0.5)
learn_lm.load('/mnt/data/group07/johannes/germanlm/exp_10/models/2019_ 4_14_20_48_17_552279')
learn_lm.fit_one_cycle(1, 1e-2, moms=(0.8,0.7))
learn_lm.unfreeze()
learn_lm.fit_one_cycle(8, 1e-3, moms=(0.8,0.7))
learn_lm.save_encoder('enc')
classes = df_train['label'].unique().tolist()
for dfx in [df_train, df_valid, df_test]:
dfx['label'] = dfx['label'].apply(lambda x: classes.index(x))
# NB: set the corrext padding idx
data_train = TextClasDataBunch.from_ids('uf_de_exp', pad_idx=25000, classes=classes, bs=32, vocab=voc, train_lbls=df_train['label'], train_ids=df_train['text'], valid_ids=df_valid['text'], valid_lbls=df_valid['label'])
# store the test dataset within another TextClasDataBunch
data_test = TextClasDataBunch.from_ids('uf_de_exp', pad_idx=25000, classes=classes, bs=32, vocab=voc, train_lbls=df_train['label'], train_ids=df_train['text'], valid_ids=df_test['text'], valid_lbls=df_test['label'])
learn = text_classifier_learner(data_train, AWD_LSTM, drop_mult=0.5)
learn.load_encoder('enc', device='cuda:2')
factor = 2.6
lr = 4e-3
lrs = [lr / (factor ** (4 - x)) for x in range(4)] + [lr]
learn.fit(1, lrs)
learn.freeze_to(-2)
learn.fit(1, lrs)
learn.callbacks += [
SaveModelCallback(learn, name='best', monitor='accuracy'),
EarlyStoppingCallback(learn, monitor='accuracy', patience=10),
]
learn.unfreeze()
learn.fit(100, lrs)
learn.validate(data_test.valid_dl)
```
| github_jupyter |
```
# look at tools/set_up_magics.ipynb
yandex_metrica_allowed = True ; get_ipython().run_cell('# one_liner_str\n\nget_ipython().run_cell_magic(\'javascript\', \'\', \'// setup cpp code highlighting\\nIPython.CodeCell.options_default.highlight_modes["text/x-c++src"] = {\\\'reg\\\':[/^%%cpp/]} ;\')\n\n# creating magics\nfrom IPython.core.magic import register_cell_magic, register_line_magic\nfrom IPython.display import display, Markdown, HTML\nimport argparse\nfrom subprocess import Popen, PIPE\nimport random\nimport sys\nimport os\nimport re\nimport signal\nimport shutil\nimport shlex\nimport glob\n\n@register_cell_magic\ndef save_file(args_str, cell, line_comment_start="#"):\n parser = argparse.ArgumentParser()\n parser.add_argument("fname")\n parser.add_argument("--ejudge-style", action="store_true")\n args = parser.parse_args(args_str.split())\n \n cell = cell if cell[-1] == \'\\n\' or args.no_eof_newline else cell + "\\n"\n cmds = []\n with open(args.fname, "w") as f:\n f.write(line_comment_start + " %%cpp " + args_str + "\\n")\n for line in cell.split("\\n"):\n line_to_write = (line if not args.ejudge_style else line.rstrip()) + "\\n"\n if line.startswith("%"):\n run_prefix = "%run "\n if line.startswith(run_prefix):\n cmds.append(line[len(run_prefix):].strip())\n f.write(line_comment_start + " " + line_to_write)\n continue\n run_prefix = "%# "\n if line.startswith(run_prefix):\n f.write(line_comment_start + " " + line_to_write)\n continue\n raise Exception("Unknown %%save_file subcommand: \'%s\'" % line)\n else:\n f.write(line_to_write)\n f.write("" if not args.ejudge_style else line_comment_start + r" line without \\n")\n for cmd in cmds:\n display(Markdown("Run: `%s`" % cmd))\n get_ipython().system(cmd)\n\n@register_cell_magic\ndef cpp(fname, cell):\n save_file(fname, cell, "//")\n\n@register_cell_magic\ndef asm(fname, cell):\n save_file(fname, cell, "//")\n \n@register_cell_magic\ndef makefile(fname, cell):\n assert not fname\n save_file("makefile", cell.replace(" " * 4, "\\t"))\n \n@register_line_magic\ndef p(line):\n try:\n expr, comment = line.split(" #")\n display(Markdown("`{} = {}` # {}".format(expr.strip(), eval(expr), comment.strip())))\n except:\n display(Markdown("{} = {}".format(line, eval(line))))\n \ndef show_file(file, clear_at_begin=True, return_html_string=False):\n if clear_at_begin:\n get_ipython().system("truncate --size 0 " + file)\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n elem.innerText = xmlhttp.responseText;\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (errors___OBJ__ < 10 && !entrance___OBJ__) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n \n <font color="white"> <tt>\n <p id="__OBJ__" style="font-size: 16px; border:3px #333333 solid; background: #333333; border-radius: 10px; padding: 10px; "></p>\n </tt> </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__ -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n \nBASH_POPEN_TMP_DIR = "./bash_popen_tmp"\n \ndef bash_popen_terminate_all():\n for p in globals().get("bash_popen_list", []):\n print("Terminate pid=" + str(p.pid), file=sys.stderr)\n p.terminate()\n globals()["bash_popen_list"] = []\n if os.path.exists(BASH_POPEN_TMP_DIR):\n shutil.rmtree(BASH_POPEN_TMP_DIR)\n\nbash_popen_terminate_all() \n\ndef bash_popen(cmd):\n if not os.path.exists(BASH_POPEN_TMP_DIR):\n os.mkdir(BASH_POPEN_TMP_DIR)\n h = os.path.join(BASH_POPEN_TMP_DIR, str(random.randint(0, 1e18)))\n stdout_file = h + ".out.html"\n stderr_file = h + ".err.html"\n run_log_file = h + ".fin.html"\n \n stdout = open(stdout_file, "wb")\n stdout = open(stderr_file, "wb")\n \n html = """\n <table width="100%">\n <colgroup>\n <col span="1" style="width: 70px;">\n <col span="1">\n </colgroup> \n <tbody>\n <tr> <td><b>STDOUT</b></td> <td> {stdout} </td> </tr>\n <tr> <td><b>STDERR</b></td> <td> {stderr} </td> </tr>\n <tr> <td><b>RUN LOG</b></td> <td> {run_log} </td> </tr>\n </tbody>\n </table>\n """.format(\n stdout=show_file(stdout_file, return_html_string=True),\n stderr=show_file(stderr_file, return_html_string=True),\n run_log=show_file(run_log_file, return_html_string=True),\n )\n \n cmd = """\n bash -c {cmd} &\n pid=$!\n echo "Process started! pid=${{pid}}" > {run_log_file}\n wait ${{pid}}\n echo "Process finished! exit_code=$?" >> {run_log_file}\n """.format(cmd=shlex.quote(cmd), run_log_file=run_log_file)\n # print(cmd)\n display(HTML(html))\n \n p = Popen(["bash", "-c", cmd], stdin=PIPE, stdout=stdout, stderr=stdout)\n \n bash_popen_list.append(p)\n return p\n\n\n@register_line_magic\ndef bash_async(line):\n bash_popen(line)\n \n \ndef show_log_file(file, return_html_string=False):\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function halt__OBJ__(elem, color)\n {\n elem.setAttribute("style", "font-size: 14px; background: " + color + "; padding: 10px; border: 3px; border-radius: 5px; color: white; "); \n }\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n if (entrance___OBJ__ < 0) {\n entrance___OBJ__ = 0;\n }\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n if (elem.innerHTML != xmlhttp.responseText) {\n elem.innerHTML = xmlhttp.responseText;\n }\n if (elem.innerHTML.includes("Process finished.")) {\n halt__OBJ__(elem, "#333333");\n } else {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (!entrance___OBJ__) {\n if (errors___OBJ__ < 6) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n } else {\n halt__OBJ__(elem, "#994444");\n }\n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n\n <p id="__OBJ__" style="font-size: 14px; background: #000000; padding: 10px; border: 3px; border-radius: 5px; color: white; ">\n </p>\n \n </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__.md -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n\n \nclass TInteractiveLauncher:\n tmp_path = "./interactive_launcher_tmp"\n def __init__(self, cmd):\n try:\n os.mkdir(TInteractiveLauncher.tmp_path)\n except:\n pass\n name = str(random.randint(0, 1e18))\n self.inq_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".inq")\n self.log_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".log")\n \n os.mkfifo(self.inq_path)\n open(self.log_path, \'w\').close()\n open(self.log_path + ".md", \'w\').close()\n\n self.pid = os.fork()\n if self.pid == -1:\n print("Error")\n if self.pid == 0:\n exe_cands = glob.glob("../tools/launcher.py") + glob.glob("../../tools/launcher.py")\n assert(len(exe_cands) == 1)\n assert(os.execvp("python3", ["python3", exe_cands[0], "-l", self.log_path, "-i", self.inq_path, "-c", cmd]) == 0)\n self.inq_f = open(self.inq_path, "w")\n interactive_launcher_opened_set.add(self.pid)\n show_log_file(self.log_path)\n\n def write(self, s):\n s = s.encode()\n assert len(s) == os.write(self.inq_f.fileno(), s)\n \n def get_pid(self):\n n = 100\n for i in range(n):\n try:\n return int(re.findall(r"PID = (\\d+)", open(self.log_path).readline())[0])\n except:\n if i + 1 == n:\n raise\n time.sleep(0.1)\n \n def input_queue_path(self):\n return self.inq_path\n \n def close(self):\n self.inq_f.close()\n os.waitpid(self.pid, 0)\n os.remove(self.inq_path)\n # os.remove(self.log_path)\n self.inq_path = None\n self.log_path = None \n interactive_launcher_opened_set.remove(self.pid)\n self.pid = None\n \n @staticmethod\n def terminate_all():\n if "interactive_launcher_opened_set" not in globals():\n globals()["interactive_launcher_opened_set"] = set()\n global interactive_launcher_opened_set\n for pid in interactive_launcher_opened_set:\n print("Terminate pid=" + str(pid), file=sys.stderr)\n os.kill(pid, signal.SIGKILL)\n os.waitpid(pid, 0)\n interactive_launcher_opened_set = set()\n if os.path.exists(TInteractiveLauncher.tmp_path):\n shutil.rmtree(TInteractiveLauncher.tmp_path)\n \nTInteractiveLauncher.terminate_all()\n \nyandex_metrica_allowed = bool(globals().get("yandex_metrica_allowed", False))\nif yandex_metrica_allowed:\n display(HTML(\'\'\'<!-- YANDEX_METRICA_BEGIN -->\n <script type="text/javascript" >\n (function(m,e,t,r,i,k,a){m[i]=m[i]||function(){(m[i].a=m[i].a||[]).push(arguments)};\n m[i].l=1*new Date();k=e.createElement(t),a=e.getElementsByTagName(t)[0],k.async=1,k.src=r,a.parentNode.insertBefore(k,a)})\n (window, document, "script", "https://mc.yandex.ru/metrika/tag.js", "ym");\n\n ym(59260609, "init", {\n clickmap:true,\n trackLinks:true,\n accurateTrackBounce:true\n });\n </script>\n <noscript><div><img src="https://mc.yandex.ru/watch/59260609" style="position:absolute; left:-9999px;" alt="" /></div></noscript>\n <!-- YANDEX_METRICA_END -->\'\'\'))\n\ndef make_oneliner():\n html_text = \'("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "")\'\n html_text += \' + "<""!-- MAGICS_SETUP_PRINTING_END -->"\'\n return \'\'.join([\n \'# look at tools/set_up_magics.ipynb\\n\',\n \'yandex_metrica_allowed = True ; get_ipython().run_cell(%s);\' % repr(one_liner_str),\n \'display(HTML(%s))\' % html_text,\n \' #\'\'MAGICS_SETUP_END\'\n ])\n \n\n');display(HTML(("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "") + "<""!-- MAGICS_SETUP_PRINTING_END -->")) #MAGICS_SETUP_END
```
# HTTP, libcurl, more cmake
<p><a href="https://www.youtube.com/watch?v=oXEKbLwc6y8&list=PLjzMm8llUm4AmU6i_hPU0NobgA4VsBowc&index=26" target="_blank">
<h3>Видеозапись семинара</h3>
</a></p>
[Ридинг Яковлева про HTTP](https://github.com/victor-yacovlev/mipt-diht-caos/tree/master/practice/http-curl)
[Ридинг Яковлева про cmake](https://github.com/victor-yacovlev/mipt-diht-caos/blob/master/practice/linux_basics/cmake.md)
## HTTP
[HTTP (HyperText Transfer Protocol)](https://ru.wikipedia.org/wiki/HTTP) — протокол прикладного/транспортного уровня передачи данных.
Изначально был создан как протокол прикладного уровня для передачи документов в html формате (теги и все вот это). Но позже был распробован и сейчас может используется для передачи произвольных данных, что характерно для транспортного уровня.
Отправка HTTP запроса:
* <a href="#get_term" style="color:#856024"> Из терминала </a>
* <a href="#netcat" style="color:#856024"> С помощью netcat, telnet </a> на уровне TCP, самостоятельно формируя HTTP запрос.
* <a href="#curl" style="color:#856024"> С помощью curl </a> на уровне HTTP
* <a href="#get_python" style="color:#856024"> Из python </a> на уровне HTTP
* <a href="#get_c" style="color:#856024"> Из программы на C </a> на уровне HTTP
* <a href="#touch_http" style="color:#856024"> Более разнообразное использование HTTP </a>
#### HTTP 1.1 и HTTP/2
На семинаре будем рассматривать HTTP 1.1, но стоит знать, что текущая версия протокола существенно более эффективна.
[Как HTTP/2 сделает веб быстрее / Хабр](https://habr.com/ru/company/nix/blog/304518/)
| HTTP 1.1 | HTTP/2 |
|----------|--------|
| одно соединение - один запрос, <br> как следствие вынужденная конкатенация, встраивание и спрайтинг (spriting) данных | несколько запросов на соединение |
| все нужные заголовки каждый раз отправляются полностью | сжатие заголовков, позволяет не отправлять каждый раз одни и те же заголовки |
| | возможность отправки данных по инициативе сервера |
| текстовый протокол | двоичный протокол |
| | приоритезация потоков - клиент может сообщать, что ему более важно|
## libcurl
Библиотека умеющая все то же, что и утилита curl.
## cmake
Решает задачу кроссплатформенной сборки
* Фронтенд для систем непосредственно занимающихся сборкой
* cmake хорошо интегрирован с многими IDE
* CMakeLists.txt в корне дерева исходников - главный конфигурационный файл и главный индикатор того, что проект собирается с помощью cmake
Примеры:
* <a href="#сmake_simple" style="color:#856024"> Простой пример </a>
* <a href="#сmake_curl" style="color:#856024"> Пример с libcurl </a>
[Введение в CMake / Хабр](https://habr.com/ru/post/155467/)
[Документация для libCURL](https://curl.haxx.se/libcurl/c/)
<a href="#hw" style="color:#856024">Комментарии к ДЗ</a>
## <a name="get_term"></a> HTTP из терминала
#### <a name="netcat"></a> На уровне TCP
```
%%bash
# make request string
VAR=$(cat <<HEREDOC_END
GET / HTTP/1.1
Host: ejudge.atp-fivt.org
HEREDOC_END
)
# Если работаем в терминале, то просто пишем "nc ejudge.atp-fivt.org 80" и вводим запрос
# ↓↓↓↓↓↓↓↓↓↓↓↓↓↓↓↓ - имитация ввода в stdin. "-q1" - чтобы netcat не закрылся сразу после закрытия stdin
echo -e "$VAR\n" | nc -q1 ejudge.atp-fivt.org 80 | head -n 14
# ↑↑↑↑↑↑↑↑↑↑↑↑ - обрезаем только начало вывода, чтобы не затопило выводом
# Можно еще исползовать telnet: "telnet ejudge.atp-fivt.org 80"
import time
a = TInteractiveLauncher("telnet ejudge.atp-fivt.org 80 | head -n 10")
a.write("""\
GET / HTTP/1.1
Host: ejudge.atp-fivt.org
""")
time.sleep(1)
a.close()
%%bash
VAR=$(cat <<HEREDOC_END
USER pechatnov@yandex.ru
HEREDOC_END
)
# попытка загрузить почту по POP3 протоколу (не получится, там надо с шифрованием заморочиться)
echo -e "$VAR\n" | nc -q1 pop.yandex.ru 110
```
#### <a name="curl"></a> Сразу на уровне HTTP
curl - возволяет делать произвольные HTTP запросы
wget - в первую очередь предназначен для скачивания файлов. Например, умеет выкачивать страницу рекурсивно
```
%%bash
curl ejudge.atp-fivt.org | head -n 10
%%bash
wget ejudge.atp-fivt.org -O - | head -n 10
```
## <a name="get_python"></a> HTTP из python
```
import requests
data = requests.get("http://ejudge.atp-fivt.org").content.decode()
print(data[:200])
```
## <a name="get_c"></a> HTTP из C
Пример от Яковлева. (Ниже более подробно рассмотрим библиотеку)
```
%%cpp curl_easy.c
%run gcc -Wall curl_easy.c -lcurl -o curl_easy.exe
%run ./curl_easy.exe | head -n 5
#include <curl/curl.h>
#include <assert.h>
int main() {
CURL *curl = curl_easy_init();
assert(curl);
CURLcode res;
curl_easy_setopt(curl, CURLOPT_URL, "http://ejudge.atp-fivt.org");
res = curl_easy_perform(curl);
curl_easy_cleanup(curl);
assert(res == 0);
return 0;
}
```
#### <a name="touch_http"></a> Потрогаем HTTP более разнообразно
Установка:
<br>https://install.advancedrestclient.com/ - программка для удобной отправки разнообразных http запросов
<br>`pip3 install --user wsgidav cheroot` - webdav сервер
```
!mkdir webdav_dir 2>&1 | grep -v "File exists" || true
!rm -r webdav_dir/*
!echo "Hello!" > webdav_dir/file.txt
a = TInteractiveLauncher("wsgidav --port=9024 --root=./webdav_dir --auth=anonymous --host=0.0.0.0")
!curl localhost:9024 | head -n 4
!curl -X "PUT" localhost:9024/curl_added_file.txt --data-binary @curl_easy.c
!curl -X "COPY" localhost:9024/curl_added_file.txt -H "Destination: /curl_added_file_2.txt"
!ls webdav_dir
!cat webdav_dir/curl_added_file.txt | grep main -C 2
!curl -X "DELETE" localhost:9024/curl_added_file.txt
!curl -X "DELETE" localhost:9024/curl_added_file_2.txt
!ls webdav_dir
os.kill(a.get_pid(), signal.SIGINT)
a.close()
```
## libcurl
Установка: `sudo apt-get install libcurl4-openssl-dev` (Но это не точно! Воспоминания годичной давности. Напишите мне пожалуйста получится или не получится)
Документация: https://curl.haxx.se/libcurl/c/CURLOPT_WRITEFUNCTION.html
Интересный факт: размер chunk'a всегда равен 1.
Модифицирпованный пример от Яковлева
```
%%cpp curl_medium.c
%run gcc -Wall curl_medium.c -lcurl -o curl_medium.exe
%run ./curl_medium.exe "http://ejudge.atp-fivt.org" | head -n 5
#include <unistd.h>
#include <string.h>
#include <stdlib.h>
#include <assert.h>
#include <curl/curl.h>
typedef struct {
char *data;
size_t length;
size_t capacity;
} buffer_t;
static size_t callback_function(
char *ptr, // буфер с прочитанными данными
size_t chunk_size, // размер фрагмента данных; всегда равен 1
size_t nmemb, // количество фрагментов данных
void *user_data // произвольные данные пользователя
) {
buffer_t *buffer = user_data;
size_t total_size = chunk_size * nmemb;
size_t required_capacity = buffer->length + total_size;
if (required_capacity > buffer->capacity) {
required_capacity *= 2;
buffer->data = realloc(buffer->data, required_capacity);
assert(buffer->data);
buffer->capacity = required_capacity;
}
memcpy(buffer->data + buffer->length, ptr, total_size);
buffer->length += total_size;
return total_size;
}
int main(int argc, char *argv[]) {
assert(argc == 2);
const char* url = argv[1];
CURL *curl = curl_easy_init();
assert(curl);
CURLcode res;
// регистрация callback-функции записи
curl_easy_setopt(curl, CURLOPT_WRITEFUNCTION, callback_function);
// указатель &buffer будет передан в callback-функцию
// параметром void *user_data
buffer_t buffer = {.data = NULL, .length = 0, .capacity = 0};
curl_easy_setopt(curl, CURLOPT_WRITEDATA, &buffer);
curl_easy_setopt(curl, CURLOPT_URL, url);
res = curl_easy_perform(curl);
assert(res == 0);
write(STDOUT_FILENO, buffer.data, buffer.length);
free(buffer.data);
curl_easy_cleanup(curl);
}
```
## cmake
Установка: `apt-get install cmake cmake-extras`
#### <a name="cmake_simple"></a> Простой пример
Источник: [Введение в CMake / Хабр](https://habr.com/ru/post/155467/). Там же можно найти множество более интересных примеров.
```
!mkdir simple_cmake_example 2>&1 | grep -v "File exists" || true
%%cmake simple_cmake_example/CMakeLists.txt
cmake_minimum_required(VERSION 2.8) # Проверка версии CMake.
# Если версия установленой программы
# старее указаной, произайдёт аварийный выход.
add_executable(main main.cpp) # Создает исполняемый файл с именем main
# из исходника main.cpp
%%cpp simple_cmake_example/main.cpp
%run mkdir simple_cmake_example/build #// cоздаем директорию для файлов сборки
%# // переходим в нее, вызываем cmake, чтобы он создал правильный Makefile
%# // а затем make, который по Makefile правильно все соберет
%run cd simple_cmake_example/build && cmake .. && make
%run simple_cmake_example/build/main #// запускаем собранный бинарь
%run ls -la simple_cmake_example #// смотрим, а что же теперь есть в основной директории
%run ls -la simple_cmake_example/build #// ... и в директории сборки
%run rm -r simple_cmake_example/build #// удаляем директорию с файлами сборки
#include <iostream>
int main(int argc, char** argv)
{
std::cout << "Hello, World!" << std::endl;
return 0;
}
```
#### <a name="cmake_curl"></a> Пример с libcurl
```
!mkdir curl_cmake_example || true
!cp curl_medium.c curl_cmake_example/main.c
%%cmake curl_cmake_example/CMakeLists.txt
%run mkdir curl_cmake_example/build
%run cd curl_cmake_example/build && cmake .. && make
%run curl_cmake_example/build/main "http://ejudge.atp-fivt.org" | head -n 5 #// запускаем собранный бинарь
%run rm -r curl_cmake_example/build
cmake_minimum_required(VERSION 2.8)
set(CMAKE_C_FLAGS "-std=gnu11") # дополнительные опции компилятора Си
# найти библиотеку CURL; опция REQUIRED означает,
# что библиотека является обязательной для сборки проекта,
# и если необходимые файлы не будут найдены, cmake
# завершит работу с ошибкой
find_package(CURL REQUIRED)
# это библиотека в проекте не нужна, просто пример, как написать обработку случаев, когда библиотека не найдена
find_package(SDL)
if(NOT SDL_FOUND)
message(">>>>> Failed to find SDL (not a problem)")
else()
message(">>>>> Managed to find SDL, can add include directories, add target libraries")
endif()
# это библиотека в проекте не нужна, просто пример, как подключить модуль интеграции с pkg-config
find_package(PkgConfig REQUIRED)
# и ненужный в этом проекте FUSE через pkg-config
pkg_check_modules(
FUSE # имя префикса для названий выходных переменных
# REQUIRED # опционально можно писать, чтобы было required
fuse3 # имя библиотеки, должен существовать файл fuse3.pc
)
if(NOT FUSE_FOUND)
message(">>>>> Failed to find FUSE (not a problem)")
else()
message(">>>>> Managed to find FUSE, can add include directories, add target libraries")
endif()
# добавляем цель собрать исполняемый файл из перечисленных исходнико
add_executable(main main.c)
# добавляет в список каталогов для цели main,
# которые превратятся в опции -I компилятора для всех
# каталогов, которые перечислены в переменной CURL_INCLUDE_DIRECTORIES
target_include_directories(main PUBLIC ${CURL_INCLUDE_DIRECTORIES})
# include_directories(${CURL_INCLUDE_DIRECTORIES}) # можно вот так
# для цели my_cool_program указываем библиотеки, с которыми
# программа будет слинкована (в результате станет опциями -l и -L)
target_link_libraries(main ${CURL_LIBRARIES})
```
# <a name="hw"></a> Комментарии к ДЗ
* `Connection: close` - чтобы с вами keep-alive не поддерживали
* Комментарий от [Михаила Циона](https://github.com/MVCionOld):
<br> От себя хочу добавить про использование `сURL`'a. Одним из хэдеров в `http`-запросе есть `User-Agent`, которые сигнализирует сайту про, что "вы" это то браузер, поисковый бот/скраппер, мобильный телефоны или холодильник. Некоторые сайты нормально открываются в браузере, но при попытке получить исходный `HTML` код с помощью `cURL` эти запросы могут отклоняться. Могут возвращаться коды ответов, например, 403, то есть доступ запрещён.
<br> Зачастую боты не несут никакой пользы, но в то же время создают нагрузку на сервис и/или ведут другую вредоносную активность. Насколько мне известно, есть два способа бороться с такими негодяями: проверять `User-Agent` и использование `JavaScript`. Во втором случае это инъекции на куки, асинхронная генерация страницы и тд. Что касается агента - банально денаить конкретные паттерны. У `сURL`'a есть своя строка для агента, в основном меняется только версия, например `curl/7.37.0`.
<br> Возможно, кто-то сталкивался с тем, что при написании скраппера основанного на `сURL`'e вы получали `BadRequest` (например, при тестировании задачи **inf21-2**), хотя сайт прекрасно открывался. Это как раз первый случай.
<br> Однако, можно менять агента, например, из терминала:
<br> `curl -H "User-Agent: Mozilla/5.0" url`
<br> при использовании `libcurl`:
<br> `curl_easy_setopt(curl, CURLOPT_USERAGENT, "Mozilla/5.0");`
| github_jupyter |
```
#
# COMMENTS TO DO
#
#Condensed code based on the code from: https://jmetzen.github.io/2015-11-27/vae.html
%matplotlib inline
import tensorflow as tf
import tensorflow.contrib.layers as layers
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
import numpy as np
import os
import time
import glob
from tensorflow.examples.tutorials.mnist import input_data
def plot(samples, w, h, fw, fh, iw=28, ih=28):
fig = plt.figure(figsize=(fw, fh))
gs = gridspec.GridSpec(w, h)
gs.update(wspace=0.05, hspace=0.05)
for i, sample in enumerate(samples):
ax = plt.subplot(gs[i])
plt.axis('off')
ax.set_xticklabels([])
ax.set_yticklabels([])
ax.set_aspect('equal')
plt.imshow(sample.reshape(iw, ih), cmap='Greys_r')
return fig
def encoder(images, num_outputs_h0=8, num_outputs_h1=16, kernel_size=5, stride=2, num_hidden_fc=1024, z_dim=100):
print("Encoder")
h0 = layers.convolution2d(
inputs=images,
num_outputs=num_outputs_h0,
kernel_size=kernel_size,
stride=stride,
activation_fn=tf.nn.relu,
scope='e_cnn_%d' % (0,)
)
print("Convolution 1 -> {}".format(h0))
h1 = layers.convolution2d(
inputs=h0,
num_outputs=num_outputs_h1,
kernel_size=kernel_size,
stride=stride,
activation_fn=tf.nn.relu,
scope='e_cnn_%d' % (1,)
)
print("Convolution 2 -> {}".format(h1))
h1_dim = h1.get_shape().as_list()[1]
h2_flat = tf.reshape(h1, [-1, h1_dim * h1_dim * num_outputs_h1])
print("Reshape -> {}".format(h2_flat))
h2_flat =layers.fully_connected(
inputs=h2_flat,
num_outputs=num_hidden_fc,
activation_fn=tf.nn.relu,
scope='e_d_%d' % (0,)
)
print("FC 1 -> {}".format(h2_flat))
z_mean =layers.fully_connected(
inputs=h2_flat,
num_outputs=z_dim,
activation_fn=None,
scope='e_d_%d' % (1,)
)
print("Z mean -> {}".format(z_mean))
z_log_sigma_sq =layers.fully_connected(
inputs=h2_flat,
num_outputs=z_dim,
activation_fn=None,
scope='e_d_%d' % (2,)
)
return z_mean, z_log_sigma_sq
def decoder(z, num_hidden_fc=1024, h1_reshape_dim=7, kernel_size=5, h1_channels=16, h2_channels = 8, output_channels=1, strides=2, output_dims=784):
print("Decoder")
batch_size = tf.shape(z)[0]
h0 =layers.fully_connected(
inputs=z,
num_outputs=num_hidden_fc,
activation_fn=tf.nn.relu,
scope='d_d_%d' % (0,)
)
print("FC 1 -> {}".format(h0))
h1 =layers.fully_connected(
inputs=h0,
num_outputs=h1_reshape_dim*h1_reshape_dim*h1_channels,
activation_fn=tf.nn.relu,
scope='d_d_%d' % (1,)
)
print("FC 2 -> {}".format(h1))
h1_reshape = tf.reshape(h1, [-1, h1_reshape_dim, h1_reshape_dim, h1_channels])
print("Reshape -> {}".format(h1_reshape))
wdd2 = tf.get_variable('wd2', shape=(kernel_size, kernel_size, h2_channels, h1_channels), initializer=tf.contrib.layers.xavier_initializer())
bdd2 = tf.get_variable('bd2', shape=(h2_channels,), initializer=tf.constant_initializer(0))
h2 = tf.nn.conv2d_transpose(h1_reshape, wdd2, output_shape=(batch_size, h1_reshape_dim*2, h1_reshape_dim*2, h2_channels), strides=(1, strides, strides, 1), padding='SAME')
h2_out = tf.nn.relu(h2 + bdd2)
h2_out = tf.reshape(h2_out, (batch_size, h1_reshape_dim*2, h1_reshape_dim*2, h2_channels))
print("DeConv 1 -> {}".format(h2_out))
h2_dim = h2_out.get_shape().as_list()[1]
wdd3 = tf.get_variable('wd3', shape=(kernel_size, kernel_size, output_channels, h2_channels), initializer=tf.contrib.layers.xavier_initializer())
bdd3 = tf.get_variable('bd3', shape=(output_channels,), initializer=tf.constant_initializer(0))
h3 = tf.nn.conv2d_transpose(h2_out, wdd3, output_shape=(batch_size, h2_dim*2, h2_dim*2, output_channels), strides=(1, strides, strides, 1), padding='SAME')
h3_out = tf.nn.sigmoid(h3 + bdd3)
#Workaround to use dinamyc batch size...
h3_out = tf.reshape(h3_out, (batch_size, h2_dim*2, h2_dim*2, output_channels))
print("DeConv 2 -> {}".format(h3_out))
h3_reshape = tf.reshape(h3_out, [-1, output_dims])
print("Reshape -> {}".format(h3_reshape))
return h3_reshape
mnist = input_data.read_data_sets('DATASETS/MNIST_TF', one_hot=True)
#For reconstructing the same or a different image (denoising)
images = tf.placeholder(tf.float32, shape=(None, 784))
images_28x28x1 = tf.reshape(images, [-1, 28, 28, 1])
images_target = tf.placeholder(tf.float32, shape=(None, 784))
is_training_placeholder = tf.placeholder(tf.bool)
learning_rate_placeholder = tf.placeholder(tf.float32)
z_dim = 100
with tf.variable_scope("encoder") as scope:
z_mean, z_log_sigma_sq = encoder(images_28x28x1)
with tf.variable_scope("reparameterization") as scope:
eps = tf.random_normal(shape=tf.shape(z_mean), mean=0.0, stddev=1.0, dtype=tf.float32)
# z = mu + sigma*epsilon
z = tf.add(z_mean, tf.multiply(tf.sqrt(tf.exp(z_log_sigma_sq)), eps))
with tf.variable_scope("decoder") as scope:
x_reconstr_mean = decoder(z)
scope.reuse_variables()
##### SAMPLING #######
z_input = tf.placeholder(tf.float32, shape=[None, z_dim])
x_sample = decoder(z_input)
#reconstr_loss = tf.reduce_sum(tf.nn.sigmoid_cross_entropy_with_logits(logits=x_reconstr_mean, labels=images_target), reduction_indices=1)
offset=1e-7
obs_ = tf.clip_by_value(x_reconstr_mean, offset, 1 - offset)
reconstr_loss = -tf.reduce_sum(images_target * tf.log(obs_) + (1-images_target) * tf.log(1 - obs_), 1)
latent_loss = -.5 * tf.reduce_sum(1. + z_log_sigma_sq - tf.pow(z_mean, 2) - tf.exp(z_log_sigma_sq), reduction_indices=1)
cost = tf.reduce_mean(reconstr_loss + latent_loss)
optimizer=tf.train.AdamOptimizer(learning_rate=learning_rate_placeholder).minimize(cost)
init = tf.global_variables_initializer()
save_path = "MODELS_CVAE_MNIST/CONV_VAE_MNIST.ckpt"
CVAE_SAVER = tf.train.Saver()
with tf.Session() as sess:
sess.run(init)
CVAE_SAVER.restore(sess, save_path)
print("Model restored in file: {}".format(save_path))
random_gen = sess.run(x_sample,feed_dict={z_input: np.random.randn(100, z_dim)})
fig=plot(random_gen, 10, 10, 10, 10)
plt.show()
```
# Creating MNIST subset
```
labels = 10
subset_size_per_label = 10
for label in range(labels):
indexes = np.where(mnist.train.labels[:,label] == 1)[0]
np.random.shuffle(indexes)
if label == 0:
X_mini=mnist.train.images[indexes[:subset_size_per_label]].copy()
Y_mini=mnist.train.labels[indexes[:subset_size_per_label]].copy()
else:
X_mini=np.vstack((X_mini,mnist.train.images[indexes[:subset_size_per_label]].copy()))
Y_mini=np.vstack((Y_mini,mnist.train.labels[indexes[:subset_size_per_label]].copy()))
fig=plot(X_mini, 10, 10, 10, 10)
plt.show()
print(np.argmax(Y_mini, axis=1))
```
# Interpolating
```
from numpy.linalg import norm
import progressbar
def slerp(p0, p1, t):
omega = np.arccos(np.dot(p0/norm(p0), p1/norm(p1)))
so = np.sin(omega)
return np.sin((1.0-t)*omega) / so * p0 + np.sin(t*omega)/so * p1
def linear(p0, p1, t):
return p0 * (1-t) + p1 * t
def interpolate(sample1, sample2, alphaValues, sess, method="linear"):
x_together = np.vstack((sample1, sample2))
z_samples = sess.run(z, feed_dict={images: x_together})
#fig=plot(z_samples, 1, 2, 10, 10, 10, 10)
#plt.show()
interpolation_steps = alphaValues.shape[0]
z_interpolations = np.zeros((interpolation_steps, z_dim))
for i, alpha in enumerate(alphaValues):
if method == "slerp":
z_interpolations[i] = slerp(z_samples[0], z_samples[1], alpha)
else:
z_interpolations[i] = linear(z_samples[0], z_samples[1], alpha)
x_interpolated = sess.run(x_sample, feed_dict={z_input: z_interpolations})
#fig=plot(x_interpolated, 1, INTERPOLATION_STEPS, 10, 10)
#plt.show()
return x_interpolated
labels = 10
INTERPOLATION_STEPS = 10
alphaValues = np.linspace(0, 1, INTERPOLATION_STEPS)
n_gen = labels * subset_size_per_label * (subset_size_per_label - 1) * INTERPOLATION_STEPS
print("Total gen: {}".format(n_gen))
x_pool = np.zeros((n_gen, X_mini.shape[1]))
y_pool = np.zeros((n_gen, Y_mini.shape[1]))
with tf.Session() as sess:
sess.run(init)
CVAE_SAVER.restore(sess, save_path)
print("Model restored in file: {}".format(save_path))
bar = progressbar.ProgressBar(max_value=n_gen)
bar.start()
counter = 0
for label in range(labels):
offset = label * subset_size_per_label
for i in range(subset_size_per_label):
samples_ind = list(range(subset_size_per_label))
samples_ind.remove(i)
x_sample_1 = X_mini[offset + i].copy()
for j in samples_ind:
x_sample_2 = X_mini[offset + j].copy()
x_output=interpolate(x_sample_1, x_sample_2, alphaValues, sess, method="linear")
x_pool[counter:counter+INTERPOLATION_STEPS] = x_output.copy()
y_pool[counter:counter+INTERPOLATION_STEPS, label] = 1
counter+=INTERPOLATION_STEPS
bar.update(counter)
bar.finish()
fig=plot(x_pool[:100], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[:100], axis=1))
fig=plot(x_pool[-100:], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[-100:], axis=1))
perm = np.random.permutation(x_pool.shape[0])
x_pool = x_pool[perm]
y_pool = y_pool[perm]
perm = np.random.permutation(x_pool.shape[0])
x_pool = x_pool[perm]
y_pool = y_pool[perm]
fig=plot(x_pool[:100], 10, 10, 10, 10)
plt.show()
print(np.argmax(y_pool[:100], axis=1))
```
# Storing MINI-MNIST and GEN-MNIST
```
fx = open("DATASETS/MNIST_ALT/X_MINI_100", "wb")
np.save(fx, X_mini)
fx.close()
fy = open("DATASETS/MNIST_ALT/Y_MINI_100", "wb")
np.save(fy, Y_mini)
fy.close()
fx = open("DATASETS/MNIST_ALT/X_GEN_9K_CVAE", "wb")
np.save(fx, x_pool)
fx.close()
fy = open("DATASETS/MNIST_ALT/Y_GEN_9K_CVAE", "wb")
np.save(fy, y_pool)
fy.close()
```
| github_jupyter |
```
import pandas as pd
import os
import matplotlib.pyplot as plt
%matplotlib inline
import time
import numpy as np
parentDirectory = os.path.abspath(os.path.join(os.path.join(os.getcwd(), os.pardir), os.pardir))
DATA_DIR = parentDirectory +'/data/'
FIGURES_DIR = parentDirectory +'/figures/'
df = pd.read_parquet(DATA_DIR+'food_timeseries.parquet')
selected_codes = ['FR','DE','US','IT','CA','GB',
'ES','AU','MX','BR','IN','DK'
'NG','KE','EG','ID','SE','JP']
categories = list(df['category'].unique())
df = df.loc[df['name']!='Bánh mì']
weeks_2019 = list(df.iloc[:5]['ts'].apply(lambda x: pd.Series(x['max_ratio'])).iloc[0].index[:52])
weeks_2020 = list(df.iloc[:5]['ts'].apply(lambda x: pd.Series(x['max_ratio'])).iloc[0].index[52:])
items = df['mid'].unique()
list_items = []
for c,gr1 in df.groupby('category'):
print(c)
for item,gr2 in gr1.groupby('name'):
gr2.loc[:,'total_2019'] = gr2['ts'].apply(lambda x: np.sum([x['max_ratio'][i] for i in weeks_2019])).copy()
gr2.loc[:,'total_2020'] = gr2['ts'].apply(lambda x: np.sum([x['max_ratio'][i] for i in weeks_2020])).copy()
gr2.loc[:,'total_2019_2020'] = gr2['ts'].apply(lambda x: np.sum([x['max_ratio'][i] for i in weeks_2019+weeks_2020])).copy()
for cnt,row in gr2[['country_code','total_2019','total_2020','total_2019_2020']].iterrows():
entry = {}
entry['country'] = row['country_code']
entry['category'] = c
entry['total_2019'] = row['total_2019']
entry['total_2020'] = row['total_2020']
entry['total_2019_2020'] = row['total_2019_2020']
entry['item'] = item
list_items.append(entry)
df_analysis = pd.DataFrame(list_items)
for c,gr in df_analysis.groupby('category'):
print(c)
for country, gr2 in gr.groupby('country'):
y1 = gr2['total_2019'].rank(ascending = True).values
y2 = gr2['total_2020'].rank(ascending = True).values
y3 = gr2['total_2019_2020'].rank(ascending = True).values
if country == 'AU':
S1 = (pd.Series( y1, index = gr2['item'].values))
S2 = (pd.Series( y2, index = gr2['item'].values))
S3 = (pd.Series( y3, index = gr2['item'].values))
else:
S1 = S1.add(pd.Series( y1, index = gr2['item'].values), fill_value = 0)
S2 = S2.add(pd.Series( y2, index = gr2['item'].values), fill_value = 0)
S3 = S3.add(pd.Series( y3, index = gr2['item'].values), fill_value = 0)
print('average top 10 2019-2020:')
my_list = [i.split('(')[0].strip() for i in (list((S3/12).dropna().sort_values().index[-10:]))[::-1]]
my_string = ', '.join(map(str, my_list))
print(my_string)
print('\n')
print('all sorted by volume 2019-2020:')
my_list = [i.split('(')[0].strip() for i in (list((S3/12).dropna().sort_values().index))[::-1]]
my_string = ', '.join(map(str, my_list))
print(my_string)
print('---------------------------------------------')
```
| github_jupyter |
```
%matplotlib inline
import pandas as pd
import numpy as np
import pymc3 as pm
import theano.tensor as tt
import scipy.stats as st
from scipy import optimize
import matplotlib.pylab as plt
import theano.tensor as tt
import theano
plt.style.use('seaborn-darkgrid')
```
# Laplace approximation in PyMC3
Here first try to port a [similar Stan example](http://discourse.mc-stan.org/t/algebraic-sovler-problems-laplace-approximation-in-stan/2172)
- TL;dr summary
- http://discourse.mc-stan.org/t/algebraic-sovler-problems-laplace-approximation-in-stan/2172/37
There are also some previous efforts
- Laplace approximation in PyMC3
- https://healthyalgorithms.com/2015/06/22/laplace-approximation-in-python-another-cool-trick-with-pymc3/
- https://healthyalgorithms.com/2015/07/10/laplace-approximation-in-pymc3-revisited/
## Setup data and fit with NUTS
Make sure there is no funny business here
```
N, M, sigma = 100, 10, 2.
index = np.random.randint(0, M, size=N)
X = np.random.normal(0, sigma, size=M)
Y = np.random.poisson(np.exp(X[index]))
X
with pm.Model() as m:
sd = pm.HalfNormal('sd', 1.)
group_mu = pm.Normal('g_mu', 0., sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index]), observed=Y)
trace = pm.sample(1000, tune=1000)
pm.traceplot(trace, lines={'sd': sigma, 'g_mu': X});
pm.summary(trace).round(2)
```
## Direct port of Stan model
> Consider a model like this:
$$
\begin{align*} y &\sim p(y | x,\theta) \\ x \mid \theta &\sim N(0,Q(\theta)^{-1})\\ \theta &\sim \pi(\theta) \end{align*}
$$
> The most important thing to note is that, because $p(x,y,\theta) = p(x \mid y,\theta)p(\theta \mid y ) p(y)$ it follows that
$$
p(\theta \mid y ) = \frac{p(x,y,\theta)}{p(x \mid y, \theta) p(y)} \propto \frac{p(y\mid x,\theta)p(x \mid \theta) p(\theta)}{p(x \mid y, \theta)}.
$$
> This identity hold for every $x$. The only problem is that except in special cases (like a model with a Gaussianl likelihood), we don’t know $p(x \mid y, \theta)$.
> The trick that we use is to approximate the conditional $p(x \mid \theta,y)$ by a Gaussian that matches the location and curvature at the mode. To do this we need to find
$$
x^*(\theta) = \arg \max_x p(x \mid y,\theta)
$$
> and compute the Hessian of $p(x \mid y,\theta)$ at $x^*(\theta)$. A quick calculation shows that this Hessian is
$$
Q(\theta) + H(\theta),
$$
> where $H_{ij} = \frac{\partial^2}{\partial x_i \partial x_j} \log( p(x \mid y,\theta))$. The Gaussian approximation is then
$$
p(x \mid y,\theta) \approx N(x^*(\theta), (Q(\theta) + H(\theta))^{-1}).
$$
> We then use the above expression for $p(\theta \mid y)$ evaluating the RHS at $x=x^*(\theta)$ and get
$$
p(\theta \mid y) \propto \left(\frac{|Q(\theta)|}{|Q(\theta) + H(\theta)|}\right)^{1/2} \exp\left(-\frac{1}{2}x^*(\theta)^TQ(\theta)x^*(\theta) +\log p(y \mid x^*(\theta),\theta) \right)\pi(\theta).
$$
Data and transformed data:
```stan
data {
int N;
int M;
int y[N];
int<lower=1, upper=M> index[N];
}
```
```stan
transformed data {
vector[M] xzero = rep_vector(0.0, M);
real number_of_samples[M];
int sums[M];
for (j in 1:M) {
sums[j] = 0;
number_of_samples[j]=0.0;
}
for (i in 1:N) {
sums[index[i]] += y[i];
number_of_samples[index[i]] +=1.0;
}
// xzero = log((to_vector(sums) + 0.1) ./ to_vector(number_of_samples));
{ // Beware of empty categories!!!!!!
int tmp = M;
real summm=0.0;
for (i in 1:M) {
if(number_of_samples[i]==0){
tmp = tmp-1;
} else {
summm = summm + sums[i]/number_of_samples[i];
}
}
xzero = rep_vector(summm/tmp,M);
}
}
```
```
# transformed data block
sums = np.zeros(M)
number_of_samples = np.zeros(M)
for i in range(N):
sums[index[i]] += Y[i]
number_of_samples[index[i]] += 1
# xzero = np.log((sums + 0.1) / number_of_samples)
tmp = M
summm=0.0
for i in range(M):
if number_of_samples[i]==0:
tmp = tmp-1
else:
summm = summm + sums[i]/number_of_samples[i]
xzero = np.repeat(summm/tmp, M)
```
```stan
functions {
vector conditional_grad(vector x, vector sigma, real[] number_of_samples, int[] sums) {
vector[dims(x)[1]] result;
result = (to_vector(sums)-to_vector(number_of_samples).*exp(x)) - x/sigma[1]^2;
return result;
}
vector conditional_neg_hessian(vector x, real sigma, real[] number_of_samples) {
vector[dims(x)[1]] result;
result = to_vector(number_of_samples).*exp(x) + 1/sigma^2;
return result;
}
}
```
```
# functions block
def conditional_grad(x, sigma, number_of_samples, sums):
return (sums - number_of_samples * tt.exp(x)) - x / sigma**2
def conditional_neg_hessian(x, sigma, number_of_samples):
return number_of_samples * tt.exp(x) + 1 / sigma**2
```
### Implement algebra_solver
```stan
transformed parameters {
vector[1] sigma_tmp;
vector[M] conditional_mode;
sigma_tmp[1] = sigma;
conditional_mode = algebra_solver(conditional_grad, xzero, sigma_tmp, number_of_samples, sums );
}
```
```
# rewrite functions
# conditional_grad
def func(x, sigma):
return sums - number_of_samples * np.exp(x) - x / sigma**2
# conditional_hessian
def jac(x, sigma):
return np.diag(-number_of_samples * np.exp(x) - 1 / sigma**2)
def x_star_theta(theta):
sol = optimize.root(func, np.zeros(M), jac=jac, method='hybr', args=theta)
return sol.x
```
### Validate solver
```
point = m.test_point
point
%%timeit
x_star = x_star_theta(np.exp(point['sd_log__']))
%%timeit
point1 = pm.find_MAP(start=point, model=m, vars=[group_mu], progressbar=None)
x_star = x_star_theta(np.exp(point['sd_log__']))
point1 = pm.find_MAP(start=point, model=m, vars=[group_mu], progressbar=None)
x_star2 = point1['g_mu']
_, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.plot(x_star, x_star2, 'o');
```
### Wrap it in theano
ref https://docs.pymc.io/advanced_theano.html
```
import theano.tests.unittest_tools
class XstarTheta(tt.Op):
itypes = [tt.dscalar]
otypes = [tt.dvector]
def perform(self, node, inputs, outputs):
theta, = inputs
x = x_star_theta(theta)
outputs[0][0] = np.array(x)
def grad(self, inputs, g):
theta, = inputs
x = self(theta)
grad_tmp = 2 * x / (number_of_samples * theta**3 * tt.exp(x) + theta)
return [tt.sum(g[0] * grad_tmp)]
theano.config.compute_test_value = 'off'
theano.tests.unittest_tools.verify_grad(XstarTheta(), [np.array(0.2)])
theano.tests.unittest_tools.verify_grad(XstarTheta(), [np.array(1e-5)])
theano.tests.unittest_tools.verify_grad(XstarTheta(), [np.array(1e5)])
```
### Put it all togther
```stan
parameters {
//vector[M] group_mean;
real<lower=0> sigma;
}
model {
vector[M] laplace_precisions;
sigma ~ normal(0,2);
laplace_precisions = conditional_neg_hessian(conditional_mode, sigma,number_of_samples);
// p(y | x^*) p(x^* |sigma )/p(x^* | sigma, y)
for (i in 1:N) {
target += poisson_log_lpmf(y[i] | conditional_mode[index[i]]);
}
target += -0.5*dot_self(conditional_mode)/sigma^2 -M*log(sigma) - 0.5*sum(log(laplace_precisions));
}
generated quantities {
vector[M] x;
{
vector[M] laplace_precisions = conditional_neg_hessian(conditional_mode, sigma,number_of_samples);
for (i in 1:M) {
x[i] = normal_rng(conditional_mode[i],inv_sqrt(laplace_precisions[i]));
}
}
}
```
```
tt_XstarTheta = XstarTheta()
with pm.Model() as m_la:
sd = pm.HalfNormal('sd', 1.)
conditional_mode = tt_XstarTheta(sd)
laplace_precisions = conditional_neg_hessian(conditional_mode, sd,
number_of_samples)
obs = pm.Poisson('obs', tt.exp(conditional_mode[index]), observed=Y)
pm.Potential('x∣y,θ',
-0.5 * (conditional_mode.dot(conditional_mode)) / sd**2 -
M * tt.log(sd) - 0.5 * tt.sum(tt.log(laplace_precisions)))
group_mu = pm.Deterministic(
'g_mu',
pm.tt_rng().normal(1, conditional_mode,
1 / tt.sqrt(laplace_precisions)))
trace_la = pm.sample(1000, tune=1000)
pm.traceplot(trace_la, lines={'sd': sigma, 'g_mu': X});
df_la = pm.summary(trace_la)
df_la.round(2)
```
Compare with MCMC
```
df_nuts = pm.summary(trace)
ind = df_la.index
x = df_nuts['mean'][ind]
xlower = df_nuts['hpd_2.5'][ind]
xupper = df_nuts['hpd_97.5'][ind]
y = df_la['mean'][ind]
ylower = df_la['hpd_2.5'][ind]
yupper = df_la['hpd_97.5'][ind]
_, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.errorbar(
x,
y,
yerr=[y - ylower, yupper - y],
xerr=[x - xlower, xupper - x],
fmt='o',
alpha=.5)
ax.set_ylabel('Laplace approximation')
ax.set_xlabel('MCMC with NUTS')
ax.plot(
[min(xlower), max(xupper)], [min(xlower), max(xupper)], ls="--", c=".3");
```
### Check with a larger data set
```
N, M, sigma = 1000, 100, 2.
index2 = np.random.randint(0, M, size=N)
X2 = np.random.normal(0, sigma, size=M)
Y2 = np.random.poisson(np.exp(X2[index2]))
with pm.Model() as m2:
sd = pm.HalfNormal('sd', 1.)
group_mu = pm.Normal('g_mu', 0., sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index2]), observed=Y2)
trace2 = pm.sample(1000, tune=1000)
# transformed data block
sums = np.zeros(M)
number_of_samples = np.zeros(M)
for i in range(N):
sums[index2[i]] += Y2[i]
number_of_samples[index2[i]] += 1
# xzero = np.log((sums + 0.1) / number_of_samples)
tmp = M
summm = 0.0
for i in range(M):
if number_of_samples[i] == 0:
tmp = tmp - 1
else:
summm = summm + sums[i] / number_of_samples[i]
xzero = np.repeat(summm / tmp, M)
with pm.Model() as m_la2:
sd = pm.HalfNormal('sd', 1.)
conditional_mode = tt_XstarTheta(sd)
laplace_precisions = conditional_neg_hessian(conditional_mode, sd,
number_of_samples)
obs = pm.Poisson('obs', tt.exp(conditional_mode[index2]), observed=Y2)
pm.Potential('x∣y,θ',
-0.5 * (conditional_mode.dot(conditional_mode)) / sd**2 -
M * tt.log(sd) - 0.5 * tt.sum(tt.log(laplace_precisions)))
group_mu = pm.Deterministic(
'g_mu',
pm.tt_rng().normal(1, conditional_mode,
1 / tt.sqrt(laplace_precisions)))
trace_la2 = pm.sample(1000, tune=1000)
df_nuts = pm.summary(trace2)
df_la = pm.summary(trace_la2)
ind = df_la.index
x = df_nuts['mean'][ind]
xlower = df_nuts['hpd_2.5'][ind]
xupper = df_nuts['hpd_97.5'][ind]
y = df_la['mean'][ind]
ylower = df_la['hpd_2.5'][ind]
yupper = df_la['hpd_97.5'][ind]
_, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.errorbar(
x,
y,
yerr=[y - ylower, yupper - y],
xerr=[x - xlower, xupper - x],
fmt='o',
alpha=.5)
ax.set_ylabel('Laplace approximation')
ax.set_xlabel('MCMC with NUTS')
ax.plot(
[min(xlower), max(xupper)], [min(xlower), max(xupper)], ls="--", c=".3");
```
## Make it more automatic
Automatically infer the Mode of $p(x \mid y,\theta)$ and the Hessian around the mode
```
# set up model again following cell above
N, M, sigma = 100, 10, 2.
with pm.Model() as model:
sd = pm.HalfNormal('sd', 1.)
group_mu = pm.Normal('g_mu', 0., sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index]), observed=Y)
```
Q: If I am taking the model likelihood $p (x , \theta \mid y)$ while fix $\theta$ is that the same as taking the mode of $p (x \mid y, \theta)$?
```
from pymc3.blocking import DictToArrayBijection, ArrayOrdering
start = model.test_point
vars = pm.theanof.inputvars(group_mu)
bij = DictToArrayBijection(ArrayOrdering(vars), start)
logp_func = bij.mapf(model.fastlogp)
x0 = bij.map(start)
dlogp_func = bij.mapf(model.fastdlogp(vars))
dlogp_func(x0)
# transformed data block
sums = np.zeros(M)
number_of_samples = np.zeros(M)
for i in range(N):
sums[index[i]] += Y[i]
number_of_samples[index[i]] += 1
# xzero = np.log((sums + 0.1) / number_of_samples)
tmp = M
summm = 0.0
for i in range(M):
if number_of_samples[i] == 0:
tmp = tmp - 1
else:
summm = summm + sums[i] / number_of_samples[i]
xzero = np.repeat(summm / tmp, M)
def func(x, sigma):
return sums - number_of_samples * np.exp(x) - x / sigma**2
# conditional_hessian
def jac(x, sigma):
return np.diag(-number_of_samples * np.exp(x) - 1 / sigma**2)
def x_star_theta(theta):
sol = optimize.root(func, np.zeros(M), jac=jac, method='hybr', args=theta)
return sol.x
func(x0, np.exp(start['sd_log__']))
dlogp_func_jac = bij.mapf(model.fastd2logp(vars))
dlogp_func_jac(x0)
jac(x0, np.exp(start['sd_log__']))
```
Recompiling the dlogp and d2logp is quite slow, ideally we compile it once only
```
# modify ValueGradFunction from PyMC3
# ValueGradFunction(self.logpt, grad_vars, extra_vars, **kwargs)
class GradHessianFunction(object):
"""Create a theano function that computes a gradient and its Hessian.
Parameters
----------
cost : theano variable
The value that we compute with its gradient.
grad_vars : list of named theano variables
The arguments with respect to which the gradient is computed.
extra_vars : list of named theano variables
Other arguments of the function that are assumed constant. They
are stored in shared variables and can be set using
`set_extra_values`.
dtype : str, default=theano.config.floatX
The dtype of the arrays.
casting : {'no', 'equiv', 'save', 'same_kind', 'unsafe'}, default='no'
Casting rule for casting `grad_args` to the array dtype.
See `numpy.can_cast` for a description of the options.
Keep in mind that we cast the variables to the array *and*
back from the array dtype to the variable dtype.
kwargs
Extra arguments are passed on to `theano.function`.
Attributes
----------
size : int
The number of elements in the parameter array.
profile : theano profiling object or None
The profiling object of the theano function that computes value and
gradient. This is None unless `profile=True` was set in the
kwargs.
"""
def __init__(self,
cost,
grad_vars,
extra_vars=None,
dtype=None,
casting='no',
**kwargs):
if extra_vars is None:
extra_vars = []
names = [arg.name for arg in grad_vars + extra_vars]
if any(name is None for name in names):
raise ValueError('Arguments must be named.')
if len(set(names)) != len(names):
raise ValueError('Names of the arguments are not unique.')
if cost.ndim > 0:
raise ValueError('Cost must be a scalar.')
self._grad_vars = grad_vars
self._extra_vars = extra_vars
self._extra_var_names = set(var.name for var in extra_vars)
self._cost = cost
self._ordering = ArrayOrdering(grad_vars)
self.size = self._ordering.size
self._extra_are_set = False
if dtype is None:
dtype = theano.config.floatX
self.dtype = dtype
for var in self._grad_vars:
if not np.can_cast(var.dtype, self.dtype, casting):
raise TypeError('Invalid dtype for variable %s. Can not '
'cast to %s with casting rule %s.' %
(var.name, self.dtype, casting))
if not np.issubdtype(var.dtype, np.floating):
raise TypeError('Invalid dtype for variable %s. Must be '
'floating point but is %s.' % (var.name,
var.dtype))
givens = []
self._extra_vars_shared = {}
for var in extra_vars:
shared = theano.shared(var.tag.test_value, var.name + '_shared__')
self._extra_vars_shared[var.name] = shared
givens.append((var, shared))
self._vars_joined, self._cost_joined = self._build_joined(
self._cost, grad_vars, self._ordering.vmap)
grad = tt.grad(self._cost_joined, self._vars_joined)
grad.name = '__grad'
f = tt.flatten(grad)
idx = tt.arange(f.shape[0], dtype='int32')
def grad_i(i):
return tt.grad(f[i], self._vars_joined)
# negative full hessian
hessian = -theano.map(grad_i, idx)[0]
hessian.name = '__hessian'
inputs = [self._vars_joined]
self._theano_function = theano.function(
inputs, [grad, hessian], givens=givens, **kwargs)
self._dlogp_func = theano.function(
inputs, [grad], givens=givens, **kwargs)
self._d2logp_func = theano.function(
inputs, [hessian], givens=givens, **kwargs)
def set_extra_values(self, extra_vars):
self._extra_are_set = True
for var in self._extra_vars:
self._extra_vars_shared[var.name].set_value(extra_vars[var.name])
def get_extra_values(self):
if not self._extra_are_set:
raise ValueError('Extra values are not set.')
return {
var.name: self._extra_vars_shared[var.name].get_value()
for var in self._extra_vars
}
def __call__(self, array, extra_vars=None):
if extra_vars is not None:
self.set_extra_values(extra_vars)
if not self._extra_are_set:
raise ValueError('Extra values are not set.')
if array.shape != (self.size, ):
raise ValueError('Invalid shape for array. Must be %s but is %s.' %
((self.size, ), array.shape))
dlogp, d2logp = self._theano_function(array)
return dlogp, d2logp
def dlogp(self, array, extra_vars=None):
if extra_vars is not None:
self.set_extra_values(extra_vars)
if not self._extra_are_set:
raise ValueError('Extra values are not set.')
if array.shape != (self.size, ):
raise ValueError('Invalid shape for array. Must be %s but is %s.' %
((self.size, ), array.shape))
return self._dlogp_func(array)[0]
def d2logp(self, array, extra_vars=None):
if extra_vars is not None:
self.set_extra_values(extra_vars)
if not self._extra_are_set:
raise ValueError('Extra values are not set.')
if array.shape != (self.size, ):
raise ValueError('Invalid shape for array. Must be %s but is %s.' %
((self.size, ), array.shape))
return self._d2logp_func(array)[0]
@property
def profile(self):
"""Profiling information of the underlying theano function."""
return self._theano_function.profile
def dict_to_array(self, point):
"""Convert a dictionary with values for grad_vars to an array."""
array = np.empty(self.size, dtype=self.dtype)
for varmap in self._ordering.vmap:
array[varmap.slc] = point[varmap.var].ravel().astype(self.dtype)
return array
def array_to_dict(self, array):
"""Convert an array to a dictionary containing the grad_vars."""
if array.shape != (self.size, ):
raise ValueError('Array should have shape (%s,) but has %s' %
(self.size, array.shape))
if array.dtype != self.dtype:
raise ValueError(
'Array has invalid dtype. Should be %s but is %s' %
(self._dtype, self.dtype))
point = {}
for varmap in self._ordering.vmap:
data = array[varmap.slc].reshape(varmap.shp)
point[varmap.var] = data.astype(varmap.dtyp)
return point
def array_to_full_dict(self, array):
"""Convert an array to a dictionary with grad_vars and extra_vars."""
point = self.array_to_dict(array)
for name, var in self._extra_vars_shared.items():
point[name] = var.get_value()
return point
def _build_joined(self, cost, args, vmap):
args_joined = tt.vector('__args_joined')
args_joined.tag.test_value = np.zeros(self.size, dtype=self.dtype)
joined_slices = {}
for vmap in vmap:
sliced = args_joined[vmap.slc].reshape(vmap.shp)
sliced.name = vmap.var
joined_slices[vmap.var] = sliced
replace = {var: joined_slices[var.name] for var in args}
return args_joined, theano.clone(cost, replace=replace)
varnames = [var.name for var in vars]
extra_vars = [var for var in model.free_RVs if var.name not in varnames]
dlogp_d2logp_func = GradHessianFunction(model.logpt, vars, extra_vars)
array0 = dlogp_d2logp_func.dict_to_array(point)
dlogp_d2logp_func.d2logp(array0, point)
jac(array0, np.exp(point['sd_log__']))
def x_star_theta(point):
sol = optimize.root(
func,
np.zeros(M),
jac=jac,
method='hybr',
args=np.exp(point['sd_log__']))
return sol.x
def dlogp_pm(array, point):
return dlogp_d2logp_func.dlogp(array, point)
def d2logp_pm(array, point):
return -dlogp_d2logp_func.d2logp(array, point)
def x_star_theta_pm(point):
sol = optimize.root(
dlogp_pm, np.zeros(M), jac=d2logp_pm, method='hybr', args=point)
return sol.x
%%timeit
thetas = np.linspace(.01, 10, 5)
for theta in thetas:
point['sd_log__'] = np.log(theta)
x_star = x_star_theta(point)
%%timeit
thetas = np.linspace(.01, 10, 5)
for theta in thetas:
point['sd_log__'] = np.log(theta)
x_star = x_star_theta_pm(point)
x_star_theta(point)
x_star_theta_pm(point)
Htheta = pm.theanof.hessian(model.logpt, vars)
```
## An approximation step method that could be combine with `CompoundStep`
```
from pymc3.step_methods.arraystep import BlockedStep
from pymc3.model import modelcontext
class LaplaceApprox(BlockedStep):
"""
Step method that approximate a node using Laplace approximation
Parameters
----------
vars : list
List of variables to do approximation over.
model: pymc3 model
Qtheta: Precision matrix of x: x | theta ~ N(0, Q(theta)^-1)
"""
def __init__(self, vars, Qtheta, model=None):
model = modelcontext(model)
self.vars = vars
self.m = model
varnames = [var.name for var in vars]
extra_vars = [
var for var in model.free_RVs if var.name not in varnames
]
self._dlogp_d2logp_func = GradHessianFunction(model.logpt, vars,
extra_vars)
x_approx = self._dlogp_d2logp_func._vars_joined
with self.m:
Htheta = pm.theanof.hessian(self.m.logpt, vars)
pm.Deterministic(
'x_approx_sd', 1 / tt.sqrt(tt.diag(Htheta)))
# I am not sure if Qtheta is really necessary ????
# seems the conditinal Hessian return above is already Q(θ) + H(θ) ????
pm.Potential('x|y,theta',
pm.MvNormal.dist(x_approx,
tau=Htheta).logp(x_approx))
# pm.Potential('x|y,theta',
# pm.MvNormal.dist(x_approx,
# tau=Qtheta + Htheta).logp(x_approx))
def dlogp_pm(self, array, point):
return self._dlogp_d2logp_func.dlogp(array, point)
def d2logp_pm(self, array, point):
return self._dlogp_d2logp_func.d2logp(array, point)
def x_star_theta_pm(self, point):
array0 = self._dlogp_d2logp_func.dict_to_array(self.m.test_point)
sol = optimize.root(
self.dlogp_pm,
np.zeros_like(array0), # array0 or a random initial point for the solver
jac=self.d2logp_pm,
method='hybr',
args=point)
return sol.x
def step(self, point):
xmode = self.x_star_theta_pm(point)
point = self._dlogp_d2logp_func.array_to_full_dict(xmode)
return point
with pm.Model() as m_la2:
sd = pm.HalfNormal('sd', 1.)
group_mu = pm.Normal('g_mu', 0., sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index]), observed=Y)
step1 = pm.NUTS(vars=[sd])
step2 = LaplaceApprox(vars=[group_mu], Qtheta=tt.eye(M)*sd**-2)
trace_la2 = pm.sample(1000, tune=1000, step=[step2, step1])
x_mode = trace_la2['g_mu']
x_mode_sd = trace_la2['x_approx_sd']
g_mu_approx = st.norm.rvs(x_mode, x_mode_sd)
trace_la2.add_values({'g_mu_approx': g_mu_approx})
pm.traceplot(
trace_la2,
varnames=['sd', 'g_mu_approx'],
lines={
'sd': sigma,
'g_mu_approx': X
});
df_1 = pm.summary(trace, varnames=['sd', 'g_mu'])
df_2 = pm.summary(trace_la2, varnames=['sd', 'g_mu_approx'])
x = df_1['mean']
xlower = df_1['hpd_2.5']
xupper = df_1['hpd_97.5']
y = df_2['mean']
ylower = df_2['hpd_2.5']
yupper = df_2['hpd_97.5']
_, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.errorbar(
x,
y,
yerr=[y - ylower, yupper - y],
xerr=[x - xlower, xupper - x],
fmt='o',
alpha=.5)
ax.set_ylabel('Laplace approximation')
ax.set_xlabel('MCMC with NUTS')
ax.plot(
[min(xlower), max(xupper)], [min(xlower), max(xupper)], ls="--", c=".3");
```
## Another example
```
N, M, sigma = 500, 20, 3.5
index = np.random.randint(0, M, size=N)
X = np.random.normal(0, sigma, size=M)
Y = np.random.poisson(np.exp(X[index]))
with pm.Model() as m_la2:
sd = pm.HalfNormal('sd', 2.)
group_mu = pm.Normal('g_mu', 0, sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index]), observed=Y)
step1 = pm.NUTS(vars=[sd])
step2 = LaplaceApprox(vars=[group_mu], Qtheta=tt.eye(M)*sd**-2)
trace_la2 = pm.sample(1000, tune=1000, step=[step2, step1])
pm.traceplot(trace_la2);
x_mode = trace_la2['g_mu']
x_mode_sd = trace_la2['x_approx_sd']
g_mu_approx = st.norm.rvs(x_mode, x_mode_sd)
trace_la2.add_values({'g_mu_approx': g_mu_approx})
with pm.Model() as m:
sd = pm.HalfNormal('sd', 2.)
group_mu = pm.Normal('g_mu', 0, sd, shape=M)
obs = pm.Poisson('obs', tt.exp(group_mu[index]), observed=Y)
trace = pm.sample(1000, tune=1000)
df_1 = pm.summary(trace, varnames=['sd', 'g_mu'])
df_2 = pm.summary(trace_la2, varnames=['sd', 'g_mu_approx'])
x = df_1['mean']
xlower = df_1['hpd_2.5']
xupper = df_1['hpd_97.5']
y = df_2['mean']
ylower = df_2['hpd_2.5']
yupper = df_2['hpd_97.5']
_, ax = plt.subplots(1, 1, figsize=(5, 5))
ax.errorbar(
x,
y,
yerr=[y - ylower, yupper - y],
xerr=[x - xlower, xupper - x],
fmt='o',
alpha=.5)
ax.set_ylabel('Laplace approximation')
ax.set_xlabel('MCMC with NUTS')
ax.plot(
[min(xlower), max(xupper)], [min(xlower), max(xupper)], ls="--", c=".3");
```
## Next step: INLA
- Blog posts and tutorials
- http://www.flutterbys.com.au/stats/tut/tut12.9.html
- https://www.precision-analytics.ca/blog-1/inla
- http://www.martinmodrak.cz/2018/02/02/a-gentle-stan-vs.-inla-comparison/
- References
- http://www.statslab.cam.ac.uk/~rjs57/RSS/0708/Rue08.pdf
- https://arxiv.org/pdf/1604.00860.pdf
- https://arxiv.org/pdf/1403.4630.pdf
- https://arxiv.org/pdf/1503.00256.pdf
| github_jupyter |
# Iron March Anonymous Wikipedia Edits
This notebook briefly examines how to examine whether IP addresses associated with the fascist Iron March forum had edited Wikipedia anonymously. These IP addresses were part of a dataset that was [leaked](https://www.bellingcat.com/resources/how-tos/2019/11/06/massive-white-supremacist-message-board-leak-how-to-access-and-interpret-the-data/). The idea to do this was suggested by [@z3dster](https://twitter.com/z3dster/status/1193930176863883264) on Twitter.
## IP Addresses
First lets load in all the IP address from the torrent dataset. You'll need to obtain the dataset yourself, it's nt included as part of this notebook repository. Once you have it update the path appropriately. We're going to keep track of the date that the account was created because IP addresses from ISPs can change all the time.
As the Bellingcat article points out the IP addresses need to be taken with a huge grain of salt, because users could be behind proxies or VPNs, and the ISP for your IP address changes all the time. Presenting the edit with a WHOIS lookup for the IP address helps in evaluating the data. But it must be stressed that this is far from certain.
When we look up edits in Wikipedia we could assume that the IP address was recorded when the account was created. But this is a big assumption because the IP address could be updated with every login, depending on how the Iron March platform operated. Looking at the leaked database it appears that the forum might have been hosted using [Invision Community](https://en.wikipedia.org/wiki/Invision_Community). But this is proprietary (closed source) so it's difficult to say how the IP address was recorded.
```
import os
import pandas
data_dir = '/Users/edsu/Downloads/iron_march_201911/'
users = pandas.read_csv(os.path.join(data_dir, 'csv/core_members.csv'), parse_dates=['joined'])
users.head()
```
It's interesting to quickly see the growth of users over time. To do that lets convert the epoch time in the joined column to a proper datetime.
```
users['joined'] = pandas.to_datetime(users['joined'], unit='s')
joined = users.resample('M', on='joined').count().member_id
joined = joined.reset_index()
joined.columns = ['joined', 'users']
joined.head()
import altair
altair.renderers.enable('notebook')
chart = altair.Chart(joined, width=800, title="Iron March Account Creation")
chart = chart.configure_axisX(labelAngle=45)
chart = chart.mark_bar().encode(
altair.X('yearmonth(joined):T', title="Time"),
altair.Y('users', title="Accounts per Month"),
)
chart = chart.configure_mark(color='#095')
chart
```
So it was clearly in a growth phase when it was shut down. And perhaps it's just a coincidence, but growth seems to have accelerated during the 2016 presidential election...
## Wikipedia Edits
Now lets create a function that uses Wikipedia's [XTools](https://xtools.wmflabs.org/) service, specifically its [API](https://xtools.readthedocs.io/en/stable/api/), to return the page edits for a given IP address for a particular wikipedia site (e.g. *en.wikipedia.org* for a particular date range (which defaults to the last year).
```
import datetime
import requests
def get_edits(ip, wikipedia="en.wikipedia.org", start=None, end=None):
if start is None and end is None:
end = datetime.date.today()
start = end - datetime.timedelta(days=365)
url = 'https://xtools.wmflabs.org/api/user/nonautomated_edits/{}/{}/all/{}/{}'.format(
wikipedia,
ip,
start.strftime('%Y-%m-%d'),
end.strftime('%Y-%m-%d')
)
results = requests.get(url).json()
return results['nonautomated_edits']
```
Now we can test the function out on an IP address for the US House of Representatives [143.231.249.135](https://en.wikipedia.org/wiki/Special:Contributions/143.231.249.135).
```
get_edits('143.231.249.135')
```
## Whois
It might be useful to get a sense of where IP addresses are coming from, since they can change a lot depending on the ISP, or whether the IP is behind an organizational proxy.
```
import ipwhois
whois_cache = {}
def whois(ip):
if ip in whois_cache:
return whois_cache[ip]
ip_whois = ipwhois.IPWhois(ip)
try:
result = ip_whois.lookup_rdap()
whois_cache[ip] = result
return result
except:
return {'asn_description': 'Unknown'}
whois('143.231.249.135')
```
We can try to use the *asn_description* in our output to visually flag ISPs.
```
whois('143.231.249.135')['asn_description']
```
## Anonymous Edits
Now lets take a look at the Iron March IP addresses to see if any of them edited English Wikipedia anonymously. It will take some time to do the lookups so the function prints out a **.** for every IP address checked, and a **x** for every IP address that had some edits.
```
import sys
def anon_edits(users, wikipedia="en.wikipedia.org"):
edits = []
for user in users.itertuples():
sys.stdout.write('.')
sys.stdout.flush()
start = user.joined
end = datetime.date(2017, 10, 1)
found = False
for edit in get_edits(user.ip_address, wikipedia, start=start, end=end):
found = True
edit['wikipedia'] = wikipedia
result = {
"user": user.ip_address,
"whois": whois(user.ip_address),
"edit": edit,
"url": 'https://{e[wikipedia]}/w/index.php?diff={e[rev_id]}'.format(e=edit)
}
edits.append(result)
if found:
sys.stdout.write('x')
sys.stdout.flush()
return edits
en = anon_edits(users)
len(en)
def print_edits(edits):
for e in edits:
if not e['whois']:
e['whois'] = {'asn_description': 'unknown'}
print('{e[edit][page_title]} by {e[user]} ({e[whois][asn_description]})\n{e[url]}\n'.format(e=e))
print_edits(en)
```
As you can see from the WHOIS information there are lots of ISPs in this data. So interpreting the edits is very problematic since a user may have signed up for Iron March from one IP address and then the ISP assigned it to another user. But if you look closely you can spot some edits that do look politically motivated.
Here is what some anonymous edits to the Wikimedia Commons look like.
```
commons = anon_edits(users, 'commons.wikimedia.org')
print_edits(commons)
import csv
def save_edits(edits, filename):
fieldnames = ['ip', 'asn', 'timestamp', 'page', 'url']
output = csv.DictWriter(open(filename, 'w'), fieldnames=fieldnames)
for e in edits:
output.writerow({
"ip": e['user'],
'timestamp': e['edit']['timestamp'],
"asn": e['whois']['asn_description'],
"page": e['edit']['page_title'],
"url": e['url']
})
save_edits(en, 'data/ironmarch-en.csv')
```
| github_jupyter |
# Lecture 4
## - Multivariable linear regression
### Hypothesis
- if there is one x
- \\(H(x) = Wx + b\\)
- if there are more x
- \\(H(x_1,x_2,x_3) = W_1x_1 + W_2x_2 + W_3x_3 + b\\)
### Cost function
- \\( cost(W,b) = \frac{1}{m}\sum_{i=1}^{m} (H(x_1^i, x_2^i, x_3^i) - y^i)^2 \\)
<hr/>
## - Matrix
### Hypothesis using matrix
- \\((x_1,x_2,x_3) * \left( \begin{array}{c}W_{1}\\W_{2}\\W_{3}\\\end{array}\right) = (x_1W_1 + x_2W_2 + x_3W_3)\\)
- if there are a lot instance
- \\(\left( \begin{array}{c}x_{11} x_{12} x_{13}\\x_{21} x_{22} x_{23}\\x_{31} x_{32} x_{33}\\x_{41} x_{42} x_{43}\\x_{51} x_{52} x_{53}\\\end{array}\right) * \left( \begin{array}{c}W_{1}\\W_{2}\\W_{3}\\\end{array}\right) = \left( \begin{array}{c}
x_{11}W_1 + x_{12}W_2 + x_{13}W_3\\x_{21}W_1+ x_{22}W_2+ x_{23}W_3\\x_{31}W_1+ x_{32}W_2+ x_{33}W_3\\x_{41}W_1+ x_{42}W_2+ x_{43}W_3\\x_{51}W_1+ x_{52}W_2+ x_{53}W_3\\\end{array}\right)\\)
- [5,3] * [3,1] = [5,1]
- \\(H(X) = XW\\)
### example
| x_1 | x_2 | x_3 | Y |
|:--------: |---------- |:--------: |---------- |
| 73 | 80 | 75 | 152 |
| 93 | 88 | 93 | 185 |
| 89 | 91 |90 | 180 |
| 96 | 98 |100 | 196 |
| 73 | 66 |70 | 142 |
```
import tensorflow as tf
# without Matrix
x1_data = [73.,93.,89.,96.,73.]
x2_data = [80.,88.,91.,98.,66.]
x3_data = [75.,93.,90.,100.,70.]
y_data = [152.,185.,180.,196.,142.]
x1 = tf.placeholder("float32")
x2 = tf.placeholder("float32")
x3 = tf.placeholder("float32")
Y = tf.placeholder("float32")
w1 = tf.Variable(tf.random_normal([1]), name = 'weight1')
w2 = tf.Variable(tf.random_normal([1]), name = 'weight1')
w3 = tf.Variable(tf.random_normal([1]), name = 'weight1')
b = tf.Variable(tf.random_normal([1]), name = 'bias')
hypothesis = x1 * w1 + x2 * w2 + x3 * w3 + b
cost = tf.reduce_mean(tf.square(hypothesis - Y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate = 1e-5)
train = optimizer.minimize(cost)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, hy_val, _ = sess.run([cost, hypothesis, train],
feed_dict = {x1:x1_data, x2:x2_data, x3:x3_data, Y:y_data})
if step % 10 == 0:
print(step, "Cost: ",cost_val,"\nprediction:\n",hy_val,"\n")
#with Matrix
x_data = [[73.,80.,75.,],
[93.,88.,93.,],
[89.,91.,90.,],
[96.,98.,100.,],
[73.,66.,70.,]]
y_data = [[152.],
[185.],
[180.],
[196.],
[142.]]
#필요할때 마다 추가가 가능하도록 None 설정
X = tf.placeholder("float32",shape=[None,3])
Y = tf.placeholder("float32",shape=[None,1])
W = tf.Variable(tf.random_normal([3,1]), name='weight')
b = tf.Variable(tf.random_normal([1]), name='bias')
hypothesis = tf.matmul(X,W) + b
cost = tf.reduce_mean(tf.square(hypothesis - Y))
optimizer = tf.train.GradientDescentOptimizer(learning_rate = 1e-5)
train = optimizer.minimize(cost)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, hy_val, _ = sess.run([cost, hypothesis, train],
feed_dict = {X:x_data, Y:y_data})
if step % 10 == 0:
print(step, "Cost: ",cost_val,"\nprediction:\n",hy_val,"\n")
```
| github_jupyter |
## High and Low Pass Filters
Now, you might be wondering, what makes filters high and low-pass; why is a Sobel filter high-pass and a Gaussian filter low-pass?
Well, you can actually visualize the frequencies that these filters block out by taking a look at their fourier transforms. The frequency components of any image can be displayed after doing a Fourier Transform (FT). An FT looks at the components of an image (edges that are high-frequency, and areas of smooth color as low-frequency), and plots the frequencies that occur as points in spectrum. So, let's treat our filters as small images, and display them in the frequency domain!
```
import numpy as np
import matplotlib.pyplot as plt
import cv2
%matplotlib inline
# Define gaussian, sobel, and laplacian (edge) filters
gaussian = (1/9)*np.array([[1, 1, 1],
[1, 1, 1],
[1, 1, 1]])
sobel_x= np.array([[-1, 0, 1],
[-2, 0, 2],
[-1, 0, 1]])
sobel_y= np.array([[-1,-2,-1],
[0, 0, 0],
[1, 2, 1]])
# laplacian, edge filter
laplacian=np.array([[0, 1, 0],
[1,-4, 1],
[0, 1, 0]])
filters = [gaussian, sobel_x, sobel_y, laplacian]
filter_name = ['gaussian','sobel_x', \
'sobel_y', 'laplacian']
# perform a fast fourier transform on each filter
# and create a scaled, frequency transform image
f_filters = [np.fft.fft2(x) for x in filters]
fshift = [np.fft.fftshift(y) for y in f_filters]
frequency_tx = [np.log(np.abs(z)+1) for z in fshift]
# display 4 filters
for i in range(len(filters)):
plt.subplot(2,2,i+1),plt.imshow(frequency_tx[i],cmap = 'gray')
plt.title(filter_name[i]), plt.xticks([]), plt.yticks([])
plt.show()
```
Areas of white or light gray, allow that part of the frequency spectrum through! Areas of black mean that part of the spectrum is blocked out of the image.
Recall that the low frequencies in the frequency spectrum are at the center of the frequency transform image, and high frequencies are at the edges. You should see that the Gaussian filter allows only low-pass frequencies through, which is the center of the frequency transformed image. The sobel filters block out frequencies of a certain orientation and a laplace (detects edges regardless of orientation) filter, should block out low-frequencies!
You are encouraged to load in an image, apply a filter to it using `filter2d` then visualize what the fourier transform of that image looks like before and after a filter is applied.
```
## TODO: load in an image, and filter it using a kernel of your choice
## apply a fourier transform to the original *and* filtered images and compare them
```
| github_jupyter |
[](https://colab.research.google.com/github/mravanba/comp551-notebooks/blob/master/Perceptron_and_LinearSVM.ipynb)
# Perceptron
The Perceptron algorithm finds the linear decision boundary by considering each example $x^{(n)}, y^{(n)} \in \mathcal{D}$, where $y^{(n)} \in \{-1,+1\}$.
If $\hat{y}^{(n)} = w^\top x^{(n)}$ has a different sign from $y^{(n)}$ the weights are updated to *increase* $\hat{y}^{(n)} {y}^{(n)}$. The gradient of
$\hat{y}^{(n)} {y}^{(n)}$ wrt. $w$ is $\frac{\partial}{\partial w} y^{(n)}(w^\top x^{(n)}) = y^{(n)} x^{(n)}$. Therefore, if the example is misclassified
the Perceptron learning algorithm simply updates $w$ using
$$
w^{\{t+1\}} \leftarrow w^{\{t\}} + y^{(n)} x^{(n)}
$$
If the data is linearly separable, the algorithm is guaranteed to converge.
However, if the data is not linearly separable, this procedure does not converge, and oscillates. Below, we use a `max_iters` to in case the data is not linearly seperable. We also record the update *history* so that we can visualize the learning. To be consistent with previous classification methods, we assume the input labels are in $\{0,1\}$.
```
import numpy as np
#%matplotlib notebook
%matplotlib inline
import matplotlib.pyplot as plt
from IPython.core.debugger import set_trace
import warnings
warnings.filterwarnings('ignore')
class Perceptron:
def __init__(self, add_bias=True, max_iters=10000, record_updates=False):
self.max_iters = max_iters
self.add_bias = add_bias
self.record_updates = record_updates
if record_updates:
self.w_hist = [] # records the weight
self.n_hist = [] # records the data-point selected
def fit(self, x, y):
if x.ndim == 1:
x = x[:, None]
if self.add_bias:
N = x.shape[0]
x = np.column_stack([x,np.ones(N)])
N,D = x.shape
w = np.zeros(D) #initialize the weights
if self.record_updates:
w_hist = [w]
#y = np.sign(y -.1) #to get +1 for class 1 and -1 for class 0
y = 2*y - 1 # converting 0,1 to -1,+1
t = 0
change = True #if the weight does not change the algorithm has converged
while change and t < self.max_iters:
change = False
for n in np.random.permutation(N):
yh = np.sign(np.dot(x[n,:], w)) #predict the output of the training sample
if yh == y[n]:
continue #skip the samples which are correctly classified
#w = w + (y[n]-yh)*x[n,:] #update the weights
w = w + y[n]*x[n,:]
if self.record_updates:
self.w_hist.append(w)
self.n_hist.append(n)
change = True
t += 1
if t >= self.max_iters:
break
if change:
print(f'did not converge after {t} updates')
else:
print(f'converged after {t} iterations!')
self.w = w
return self
def predict(self, x):
if x.ndim == 1:
x = x[:, None]
Nt = x.shape[0]
if self.add_bias:
x = np.column_stack([x,np.ones(Nt)])
yh = np.sign(np.dot(self.w, x))
return (yh + 1)//2 # converting -/+1 to classes 0,1
```
Let's apply this to do binary classification with Iris flowers dataset.
Here, we choose the labels and two features to make the task linearly separable.
```
from sklearn import datasets
dataset = datasets.load_iris()
x, y = dataset['data'][:,2:], dataset['target'] #slice last two features of Iris dataset
x, y = x[y < 2, :], y[y< 2] #slice class 0 and 1
model = Perceptron(record_updates=True)
yh = model.fit(x,y)
```
Let's plot the decision bounday $w^\top x = 0$ using the update history.
```
plt.plot(x[y==0,0], x[y==0,1], 'k.' )
plt.plot(x[y==1,0], x[y==1,1], 'b.' )
x_line = np.linspace(np.min(x[:,0]), np.max(x[:,0]), 100)
for t,w in enumerate(model.w_hist):
coef = -w[0]/w[1] #slope of the decision boundary
plt.plot(x_line, coef*x_line - w[2]/w[1], 'r-',
alpha=t/len(model.w_hist), label=f't={t}')
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.title('convergence of Perceptron algorithm.')
plt.show()
dataset = datasets.load_iris()
x, y = dataset['data'][:,:], dataset['target']
print(x.shape)
```
Next let's try an example where the data is not linearly separable.
```
dataset = datasets.load_iris()
x, y = dataset['data'][:,[1,2]], dataset['target'] #slice feature 1 and 2 of Iris dataset
y = y > 1
model = Perceptron(record_updates=True)
yh = model.fit(x,y)
plt.plot(x[y==0,0], x[y==0,1], 'k.' )
plt.plot(x[y==1,0], x[y==1,1], 'b.' )
x_line = np.linspace(np.min(x[:,0]), np.max(x[:,0]), 100)
for t,w in enumerate(model.w_hist):
coef = -w[0]/w[1]
plt.plot(x_line, coef*x_line - w[2]/w[1], 'r-', alpha=t/len(model.w_hist), label=f't={t}')
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.ylim(-1,10)
plt.title('Perceptron when the data is not linearly separable')
plt.show()
```
# Linear Support Vector Machine
As we saw in lectures, soft margin SVM uses **hinge loss** $L(y, z) = \max(0, 1-yz)$. This is in contrast to the Perceptron's loss function $L(y,z) = \max(0, -yz)$. In addition, while Perceptron uses SGD with a learning rate of $\alpha=1$, we can choose other procedures.
```
class GradientDescent:
def __init__(self, learning_rate=.001, max_iters=1e4, epsilon=1e-8, record_history=False):
self.learning_rate = learning_rate
self.max_iters = max_iters
self.record_history = record_history
self.epsilon = epsilon
if record_history:
self.w_history = []
def run(self, gradient_fn, x, y, w):
grad = np.inf
t = 1
while np.linalg.norm(grad) > self.epsilon and t < self.max_iters:
grad = gradient_fn(x, y, w)
w = w - self.learning_rate * grad
if self.record_history:
self.w_history.append(w)
t += 1
return w
```
Below is a simple implementation of Linear SVM, where the only difference with our previous implementation of logistic regression is the choice of loss function and the fact that the input labels are in $\{-1,+1\}$ rather than $\{0,1\}$ (note that in the implementation below, to keep things simple, we are applying the L2 regularization to the intercept as well.)
```
def cost_fn(x, y, w, lambdaa):
N, D = x.shape # not really used!
z = np.dot(x, w) # N
J = np.mean(np.maximum(0, 1- y*z)) + (lambdaa/2.) * np.linalg.norm(w)**2 #loss of the SVM
return J
class LinearSVM:
def __init__(self, add_bias=True, lambdaa = .01):
self.add_bias = add_bias
self.lambdaa = .01
pass
def fit(self, x, y, optimizer):
if x.ndim == 1:
x = x[:, None]
if self.add_bias:
N = x.shape[0]
x = np.column_stack([x,np.ones(N)])
N,D = x.shape
y = 2*y - 1 # converting 0,1 to -1,+1
def subgradient(x, y, w):
N,D = x.shape
yh = np.dot(x, w)
violations = np.nonzero(yh*y < 1)[0] # get those indexes for which yh*y<1
grad = -np.dot(x[violations,:].T, y[violations])/N #compute x^Ty for those indexes and scale it down by N
grad += self.lambdaa * w #add the gradients from the weight regularization term
return grad
w0 = np.zeros(D)
self.w = optimizer.run(subgradient, x, y, w0)
return self
def predict(self, x):
if self.add_bias:
x = np.column_stack([x,np.ones(N)])
yh = (np.sign(x@self.w) + 1)//2 #converting -1,+1 to 0,1
return yh
```
Let's try again to fit the Iris dataset of previous example this time using linear SVM; this is the setting where the data is not linearly separable.
```
dataset = datasets.load_iris()
x, y = dataset['data'][:,[1,2]], dataset['target']
y = y > 1
optimizer = GradientDescent(learning_rate=.01, max_iters=300, record_history=True)
model = LinearSVM(lambdaa=.00001)
model.fit(x,y, optimizer)
plt.plot(x[y==0,0], x[y==0,1], 'k.' )
plt.plot(x[y==1,0], x[y==1,1], 'b.' )
x_line = np.linspace(np.min(x[:,0]), np.max(x[:,0]), 100)
for t,w in enumerate(optimizer.w_history):
coef = -w[0]/w[1]
plt.plot(x_line, coef*x_line - w[2]/w[1], 'r-', alpha=t/len(optimizer.w_history), label=f't={t}')#, alpha=(t+1)/(len(model.w_hist)+.1))
plt.xlabel(r'$x_1$')
plt.ylabel(r'$x_2$')
plt.ylim(-1,10)
plt.title('Perceptron when the data is not linearly separable')
plt.show()
```
| github_jupyter |
# Covariance and Correlation
Covariance measures how two variables vary in tandem from their means.
For example, let's say we work for an e-commerce company, and they are interested in finding a correlation between page speed (how fast each web page renders for a customer) and how much a customer spends.
numpy offers covariance methods, but we'll do it the "hard way" to show what happens under the hood. Basically we treat each variable as a vector of deviations from the mean, and compute the "dot product" of both vectors. Geometrically this can be thought of as the angle between the two vectors in a high-dimensional space, but you can just think of it as a measure of similarity between the two variables.
First, let's just make page speed and purchase amount totally random and independent of each other; a very small covariance will result as there is no real correlation:
```
%matplotlib inline
import numpy as np
from pylab import *
def de_mean(x):
xmean = mean(x)
return [xi - xmean for xi in x]
def covariance(x, y):
n = len(x)
return dot(de_mean(x), de_mean(y)) / (n-1)
pageSpeeds = np.random.normal(3.0, 1.0, 1000)
purchaseAmount = np.random.normal(50.0, 10.0, 1000)
scatter(pageSpeeds, purchaseAmount)
covariance (pageSpeeds, purchaseAmount)
```
Now we'll make our fabricated purchase amounts an actual function of page speed, making a very real correlation. The negative value indicates an inverse relationship; pages that render in less time result in more money spent:
```
purchaseAmount = np.random.normal(50.0, 10.0, 1000) / pageSpeeds
scatter(pageSpeeds, purchaseAmount)
covariance (pageSpeeds, purchaseAmount)
```
But, what does this value mean? Covariance is sensitive to the units used in the variables, which makes it difficult to interpret. Correlation normalizes everything by their standard deviations, giving you an easier to understand value that ranges from -1 (for a perfect inverse correlation) to 1 (for a perfect positive correlation):
```
def correlation(x, y):
stddevx = x.std()
stddevy = y.std()
return covariance(x,y) / stddevx / stddevy #In real life you'd check for divide by zero here
correlation(pageSpeeds, purchaseAmount)
```
numpy can do all this for you with numpy.corrcoef. It returns a matrix of the correlation coefficients between every combination of the arrays passed in:
```
np.corrcoef(pageSpeeds, purchaseAmount)
```
(It doesn't match exactly just due to the math precision available on a computer.)
We can force a perfect correlation by fabricating a totally linear relationship (again, it's not exactly -1 just due to precision errors, but it's close enough to tell us there's a really good correlation here):
```
purchaseAmount = 100 - pageSpeeds * 3
scatter(pageSpeeds, purchaseAmount)
correlation (pageSpeeds, purchaseAmount)
```
Remember, correlation does not imply causality!
## Activity
numpy also has a numpy.cov function that can compute Covariance for you. Try using it for the pageSpeeds and purchaseAmounts data above. Interpret its results, and compare it to the results from our own covariance function above.
| github_jupyter |
# How many cases of COVID-19 does each U.S. state really have?
> Reported U.S. case counts are based on the number of administered tests. Since not everyone is tested, this number is biased. We use Bayesian techniques to estimate the true number of cases.
- author: Joseph Richards
- image: images/covid-state-case-estimation.png
- hide: false
- comments: true
- categories: [MCMC, US, states, cases]
- permalink: /covid-19-us-case-estimation/
- toc: false
> Warning: This analysis contains the results of a predictive model. There are a number of assumptions made which include some speculation. Furthermore, this analysis was not prepared or reviewed by an Epidimiologist. Therefore, the assumptions and methods presented should be scrutinized carefully before arriving at any conclusions.
```
#hide
# Setup and imports
%matplotlib inline
import warnings
warnings.simplefilter('ignore')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pymc3 as pm
import requests
from IPython.display import display, Markdown
#hide
# Data utilities:
def get_statewise_testing_data():
'''
Pull all statewise data required for model fitting and
prediction
Returns:
* df_out: DataFrame for model fitting where inclusion
requires testing data from 7 days ago
* df_pred: DataFrame for count prediction where inclusion
only requires testing data from today
'''
# Pull testing counts by state:
out = requests.get('https://covidtracking.com/api/states')
df_out = pd.DataFrame(out.json())
df_out.set_index('state', drop=True, inplace=True)
# Pull time-series of testing counts:
ts = requests.get('https://covidtracking.com/api/states/daily')
df_ts = pd.DataFrame(ts.json())
# Get data from last week
date_last_week = df_ts['date'].unique()[7]
df_ts_last_week = _get_test_counts(df_ts, df_out.index, date_last_week)
df_out['num_tests_7_days_ago'] = \
(df_ts_last_week['positive'] + df_ts_last_week['negative'])
df_out['num_pos_7_days_ago'] = df_ts_last_week['positive']
# Get data from today:
df_out['num_tests_today'] = (df_out['positive'] + df_out['negative'])
# State population:
df_pop = pd.read_excel(('https://github.com/jwrichar/COVID19-mortality/blob/'
'master/data/us_population_by_state_2019.xlsx?raw=true'),
skiprows=2, skipfooter=5)
r = requests.get(('https://raw.githubusercontent.com/jwrichar/COVID19-mortality/'
'master/data/us-state-name-abbr.json'))
state_name_abbr_lookup = r.json()
df_pop.index = df_pop['Geographic Area'].apply(
lambda x: str(x).replace('.', '')).map(state_name_abbr_lookup)
df_pop = df_pop.loc[df_pop.index.dropna()]
df_out['total_population'] = df_pop['Total Resident\nPopulation']
# Tests per million people, based on today's test coverage
df_out['tests_per_million'] = 1e6 * \
(df_out['num_tests_today']) / df_out['total_population']
df_out['tests_per_million_7_days_ago'] = 1e6 * \
(df_out['num_tests_7_days_ago']) / df_out['total_population']
# People per test:
df_out['people_per_test'] = 1e6 / df_out['tests_per_million']
df_out['people_per_test_7_days_ago'] = \
1e6 / df_out['tests_per_million_7_days_ago']
# Drop states with messed up / missing data:
# Drop states with missing total pop:
to_drop_idx = df_out.index[df_out['total_population'].isnull()]
print('Dropping %i/%i states due to lack of population data: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
df_pred = df_out.copy(deep=True) # Prediction DataFrame
# Criteria for model fitting:
# Drop states with missing test count 7 days ago:
to_drop_idx = df_out.index[df_out['num_tests_7_days_ago'].isnull()]
print('Dropping %i/%i states due to lack of tests: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
# Drop states with no cases 7 days ago:
to_drop_idx = df_out.index[df_out['num_pos_7_days_ago'] == 0]
print('Dropping %i/%i states due to lack of positive tests: %s' %
(len(to_drop_idx), len(df_out), ', '.join(to_drop_idx)))
df_out.drop(to_drop_idx, axis=0, inplace=True)
# Criteria for model prediction:
# Drop states with missing test count today:
to_drop_idx = df_pred.index[df_pred['num_tests_today'].isnull()]
print('Dropping %i/%i states in prediction data due to lack of tests: %s' %
(len(to_drop_idx), len(df_pred), ', '.join(to_drop_idx)))
df_pred.drop(to_drop_idx, axis=0, inplace=True)
# Cast counts to int
df_pred['negative'] = df_pred['negative'].astype(int)
df_pred['positive'] = df_pred['positive'].astype(int)
return df_out, df_pred
def _get_test_counts(df_ts, state_list, date):
ts_list = []
for state in state_list:
state_ts = df_ts.loc[df_ts['state'] == state]
# Back-fill any gaps to avoid crap data gaps
state_ts.fillna(method='bfill', inplace=True)
record = state_ts.loc[df_ts['date'] == date]
ts_list.append(record)
df_ts = pd.concat(ts_list, ignore_index=True)
return df_ts.set_index('state', drop=True)
#hide
# Model utilities
def case_count_model_us_states(df):
# Normalize inputs in a way that is sensible:
# People per test: normalize to South Korea
# assuming S.K. testing is "saturated"
ppt_sk = np.log10(51500000. / 250000)
df['people_per_test_normalized'] = (
np.log10(df['people_per_test_7_days_ago']) - ppt_sk)
n = len(df)
# For each country, let:
# c_obs = number of observed cases
c_obs = df['num_pos_7_days_ago'].values
# c_star = number of true cases
# d_obs = number of observed deaths
d_obs = df[['death', 'num_pos_7_days_ago']].min(axis=1).values
# people per test
people_per_test = df['people_per_test_normalized'].values
covid_case_count_model = pm.Model()
with covid_case_count_model:
# Priors:
mu_0 = pm.Beta('mu_0', alpha=1, beta=100, testval=0.01)
# sig_0 = pm.Uniform('sig_0', lower=0.0, upper=mu_0 * (1 - mu_0))
alpha = pm.Bound(pm.Normal, lower=0.0)(
'alpha', mu=8, sigma=3, shape=1)
beta = pm.Bound(pm.Normal, upper=0.0)(
'beta', mu=-1, sigma=1, shape=1)
# beta = pm.Normal('beta', mu=0, sigma=1, shape=3)
sigma = pm.HalfNormal('sigma', sigma=0.5, testval=0.1)
# sigma_1 = pm.HalfNormal('sigma_1', sigma=2, testval=0.1)
# Model probability of case under-reporting as logistic regression:
mu_model_logit = alpha + beta * people_per_test
tau_logit = pm.Normal('tau_logit',
mu=mu_model_logit,
sigma=sigma,
shape=n)
tau = np.exp(tau_logit) / (np.exp(tau_logit) + 1)
c_star = c_obs / tau
# Binomial likelihood:
d = pm.Binomial('d',
n=c_star,
p=mu_0,
observed=d_obs)
return covid_case_count_model
#hide
df, df_pred = get_statewise_testing_data()
# Initialize the model:
mod = case_count_model_us_states(df)
# Run MCMC sampler
with mod:
trace = pm.sample(500, tune=500, chains=1)
#hide_input
n = len(trace['beta'])
# South Korea:
ppt_sk = np.log10(51500000. / 250000)
# Compute predicted case counts per state right now
logit_now = pd.DataFrame([
pd.Series(np.random.normal((trace['alpha'][i] + trace['beta'][i] * (np.log10(df_pred['people_per_test']) - ppt_sk)),
trace['sigma'][i]), index=df_pred.index)
for i in range(len(trace['beta']))])
prob_missing_now = np.exp(logit_now) / (np.exp(logit_now) + 1)
predicted_counts_now = np.round(df_pred['positive'] / prob_missing_now.mean(axis=0)).astype(int)
predicted_counts_now_lower = np.round(df_pred['positive'] / prob_missing_now.quantile(0.975, axis=0)).astype(int)
predicted_counts_now_upper = np.round(df_pred['positive'] / prob_missing_now.quantile(0.025, axis=0)).astype(int)
case_increase_percent = list(map(lambda x, y: (((x - y) / float(y))),
predicted_counts_now, df_pred['positive']))
df_summary = pd.DataFrame(
data = {
'Cases Reported': df_pred['positive'],
'Cases Estimated': predicted_counts_now,
'Percent Increase': case_increase_percent,
'Tests per Million People': df_pred['tests_per_million'].round(1),
'Cases Estimated (range)': list(map(lambda x, y: '(%i, %i)' % (round(x), round(y)),
predicted_counts_now_lower, predicted_counts_now_upper)),
'Cases per Million': ((df_pred['positive'] / df_pred['total_population']) * 1e6),
'Positive Test Rate': (df_pred['positive'] / (df_pred['positive'] + df_pred['negative']))
},
index=df_pred.index)
from datetime import datetime
display(Markdown("## Summary for the United States on %s:" % str(datetime.today())[:10]))
display(Markdown(f"**Reported Case Count:** {df_summary['Cases Reported'].sum():,}"))
display(Markdown(f"**Predicted Case Count:** {df_summary['Cases Estimated'].sum():,}"))
case_increase_percent = 100. * (df_summary['Cases Estimated'].sum() - df_summary['Cases Reported'].sum()) / df_summary['Cases Estimated'].sum()
display(Markdown("**Percentage Underreporting in Case Count:** %.1f%%" % case_increase_percent))
#hide
df_summary.loc[:, 'Ratio'] = df_summary['Cases Estimated'] / df_summary['Cases Reported']
df_summary.columns = ['Reported Cases', 'Est Cases', '% Increase',
'Tests per Million', 'Est Range',
'Cases per Million', 'Positive Test Rate',
'Ratio']
df_display = df_summary[['Reported Cases', 'Est Cases', 'Est Range', 'Ratio',
'Tests per Million', 'Cases per Million',
'Positive Test Rate']].copy()
```
## COVID-19 Case Estimates, by State
### Definition Of Fields:
- **Reported Cases**: The number of cases reported by each state, which is a function of how many tests are positive.
- **Est Cases**: The predicted number of cases, accounting for the fact that not everyone is tested.
- **Est Range**: The 95% confidence interval of the predicted number of cases.
- **Ratio**: `Estimated Cases` divided by `Reported Cases`.
- **Tests per Million**: The number of tests administered per one million people. The less tests administered per capita, the larger the difference between reported and estimated number of cases, generally.
- **Cases per Million**: The number of **reported** cases per on million people.
- **Positive Test Rate**: The **reported** percentage of positive tests.
```
#hide_input
df_display.sort_values(
by='Est Cases', ascending=False).style.background_gradient(
cmap='Oranges').format(
{'Ratio': "{:.1f}"}).format(
{'Tests per Million': "{:.1f}"}).format(
{'Cases per Million': "{:.1f}"}).format(
{'Positive Test Rate': "{:.0%}"})
#hide_input
df_plot = df_summary.copy(deep=True)
# Compute predicted cases per million
df_plot['predicted_counts_now_pm'] = 1e6 * (
df_pred['positive'] / prob_missing_now.mean(axis=0)) / df_pred['total_population']
df_plot['predicted_counts_now_lower_pm'] = 1e6 * (
df_pred['positive'] / prob_missing_now.quantile(0.975, axis=0))/ df_pred['total_population']
df_plot['predicted_counts_now_upper_pm'] = 1e6 * (
df_pred['positive'] / prob_missing_now.quantile(0.025, axis=0))/ df_pred['total_population']
df_plot.sort_values('predicted_counts_now_pm', ascending=False, inplace=True)
xerr = [
df_plot['predicted_counts_now_pm'] - df_plot['predicted_counts_now_lower_pm'],
df_plot['predicted_counts_now_upper_pm'] - df_plot['predicted_counts_now_pm']]
fig, axs = plt.subplots(1, 1, figsize=(15, 15))
ax = plt.errorbar(df_plot['predicted_counts_now_pm'], range(len(df_plot)-1, -1, -1),
xerr=xerr, fmt='o', elinewidth=1, label='Estimate')
ax = plt.yticks(range(len(df_plot)), df_plot.index[::-1])
ax = plt.errorbar(df_plot['Cases per Million'], range(len(df_plot)-1, -1, -1),
xerr=None, fmt='.', color='k', label='Reported')
ax = plt.xlabel('COVID-19 Case Counts Per Million People', size=20)
ax = plt.legend(fontsize='xx-large', loc=4)
ax = plt.grid(linestyle='--', color='grey', axis='x')
```
## Appendix: Model Diagnostics
### Derived relationship between Test Capacity and Case Under-reporting
Plotted is the estimated relationship between test capacity (in terms of people per test -- larger = less testing) and the likelihood a COVID-19 case is reported (lower = more under-reporting of cases).
The lines represent the posterior samples from our MCMC run (note the x-axis is plotted on a log scale). The rug plot shows the current test capacity for each state (black '|') and the capacity one week ago (cyan '+'). For comparison, South Korea's testing capacity is currently at the very left of the graph (200 people per test).
```
#hide_input
# Plot pop/test vs. Prob of case detection for all posterior samples:
x = np.linspace(0.0, 4.0, 101)
logit_pcase = pd.DataFrame([
trace['alpha'][i] + trace['beta'][i] * x
for i in range(n)])
pcase = np.exp(logit_pcase) / (np.exp(logit_pcase) + 1)
fig, ax = plt.subplots(1, 1, figsize=(14, 9))
for i in range(n):
ax = plt.plot(10**(ppt_sk + x), pcase.iloc[i], color='grey', lw=.1, alpha=.5)
plt.xscale('log')
plt.xlabel('State-wise population per test', size=14)
plt.ylabel('Probability a true case is detected', size=14)
# rug plots:
ax=plt.plot(df_pred['people_per_test'], np.zeros(len(df_pred)),
marker='|', color='k', ls='', ms=20,
label='U.S. State-wise Test Capacity Now')
ax=plt.plot(df['people_per_test_7_days_ago'], np.zeros(len(df)),
marker='+', color='c', ls='', ms=10,
label='U.S. State-wise Test Capacity 7 Days Ago')
ax = plt.legend(fontsize='x-large')
```
## About this Analysis
This analysis was done by [Joseph Richards](https://twitter.com/joeyrichar).
This project[^1] uses the testing rates per state from [https://covidtracking.com/](https://covidtracking.com/), which reports case counts and mortality by state. This is used to **estimate the number of unreported (untested) COVID-19 cases in each U.S. state.**
The analysis makes a few assumptions:
1. The probability that a case is reported by a state is a function of the number of tests run per person in that state. Hence the degree of under-reported cases is a function of tests run per capita.
2. The underlying mortality rate is the same across every state.
3. Patients take time to succumb to COVID-19, so the mortality counts *today* reflect the case counts *7 days ago*. E.g., mortality rate = (cumulative deaths today) / (cumulative cases 7 days ago).
The model attempts to find the most likely relationship between state-wise test volume (per capita) and under-reporting, such that the true underlying mortality rates between the individual states are as similar as possible. The model simultaneously finds the most likely posterior distribution of mortality rates, the most likely *true* case count per state, and the test volume vs. case underreporting relationship.
[^1]: Full details about the model are available at: https://github.com/jwrichar/COVID19-mortality
| github_jupyter |
# CapsNetS2I - Capsule Neural Network Architecture for Joint Intent Detection and Slot Filling
<img src="high-level-arch.png" width="50%" height="50%">
### Import necessary modules and libraries
```
import model
import data_loader
import flags
import tensorflow as tf
import os
import warnings
warnings.filterwarnings('ignore')
tf.logging.set_verbosity(tf.logging.ERROR)
```
### Load data
First, we define the paths for the pre-trained word embeddings, train and test datasets
```
word2vec_path = '../data-capsnets/word-vec/cc.ro.300.vec'
training_data_path = '../data-capsnets/scenario0/train.txt'
test_data_path = '../data-capsnets/scenario0/test.txt'
```
#### Load pre-trained word embeddings
There are a total of ... word vectors, so loading these takes a while (~ 10 minutes)
```
print('------------------load word2vec begin-------------------')
w2v = data_loader.load_w2v(word2vec_path)
print('------------------load word2vec end---------------------')
```
#### Load train and test datasets
```
data = data_loader.read_input_data(w2v, training_data=training_data_path, test_data=test_data_path, test=True)
```
#### Visualize the contents of the data dictionary
`intents_dict` - maps ids to intent labels
<br>
`slots_dict` - maps ids to slot labels
<br>
For slots we use the **IOB** (Inside, Outside, Beginning) notation - useful for slots that span multiple words (i.e. _"douazeci de grade"_ --> B-grade I-grade I-grade)
```
import pprint
pp = pprint.PrettyPrinter(indent=4)
print('Intent class dictionary')
pp.pprint(data['intents_dict'])
print('Slots class dictionary')
pp.pprint(data['slots_dict'])
print('Max sentence length: %d words\n' % data['max_len'])
test_sample_idx = 110
print('Test sample')
print(data['x_text_te'][test_sample_idx])
print('Intent: %s' % data['y_intents_te'][test_sample_idx])
print('Slots: %s' % data['y_slots_te'][test_sample_idx])
print('Sample containing word embedding indices:')
print(data['x_te'][test_sample_idx])
```
## Set TensorFlow application flags
These flags contain application-wide information such as the scenario, the checkpoint directory containing the saved model, as well as the hyperparameters of the model: learning rate, batch size, number of epochs, the dimensionality of the prediction and
output vectors of the capsule neural network model, etc.
```
vocab_size, word_emb_size = data['embedding'].shape
_, max_sentence_length = data['x_tr'].shape
intents_number = len(data['intents_dict'])
slots_number = len(data['slots_dict'])
hidden_size = 64
FLAGS = tf.app.flags.FLAGS
tf.app.flags.DEFINE_string('ckpt_dir', './saved_models/', 'check point dir')
tf.app.flags.DEFINE_string('scenario_num', '0', 'Scenario number')
tf.app.flags.DEFINE_string('errors_dir', './errors/', 'Errors dir')
tf.app.flags.DEFINE_float('keep_prob', 0.8, 'embedding dropout keep rate for training')
tf.app.flags.DEFINE_integer('hidden_size', hidden_size, 'embedding vector size')
tf.app.flags.DEFINE_integer('batch_size', 32, 'batch size')
tf.app.flags.DEFINE_integer('num_epochs', 20, 'num of epochs')
tf.app.flags.DEFINE_integer('vocab_size', vocab_size, 'vocab size of word vectors')
tf.app.flags.DEFINE_integer('max_sentence_length', max_sentence_length, 'max number of words in one sentence')
tf.app.flags.DEFINE_integer('intents_nr', intents_number, 'intents_number') #
tf.app.flags.DEFINE_integer('slots_nr', slots_number, 'slots_number') #
tf.app.flags.DEFINE_integer('word_emb_size', word_emb_size, 'embedding size of word vectors')
tf.app.flags.DEFINE_boolean('use_embedding', True, 'whether to use embedding or not.')
tf.app.flags.DEFINE_float('learning_rate', 0.01, 'learning rate')
tf.app.flags.DEFINE_integer('slot_routing_num', 2, 'slot routing num')
tf.app.flags.DEFINE_integer('intent_routing_num', 3, 'intent routing num')
tf.app.flags.DEFINE_integer('intent_output_dim', 16, 'intent output dimension')
tf.app.flags.DEFINE_integer('slot_output_dim', 2 * hidden_size, 'slot output dimension')
tf.app.flags.DEFINE_integer('d_a', 20, 'self attention weight hidden units number')
tf.app.flags.DEFINE_integer('r', 5, 'number of self attention heads')
tf.app.flags.DEFINE_float('alpha', 0.0001, 'coefficient for self attention loss')
tf.app.flags.DEFINE_integer('n_splits', 3, 'Number of cross-validation splits')
tf.app.flags.DEFINE_string('f', '', 'kernel')
```
#### Reset flags
We should run this whenever we want to make a change in one flag, as redefinition of an existing flag is not possible.
```
for name in list(FLAGS):
delattr(FLAGS, name)
```
## Load trained model from checkpoint directory
```
tf.reset_default_graph()
config = tf.ConfigProto()
sess = tf.Session(config=config)
# Instantiate Model
capsnet = model.CapsNet(FLAGS)
ckpt_dir = FLAGS.ckpt_dir + 'scenario' + FLAGS.scenario_num + '/'
if os.path.exists(ckpt_dir):
print('Restoring Variables from Checkpoint for testing')
saver = tf.train.Saver()
saver.restore(sess, tf.train.latest_checkpoint(ckpt_dir))
else:
print('No trained model exists in checkpoint dir!')
import numpy as np
text = 'Buna Pepper as vrea sa aprinzi lumina in hol'
text_len = np.asarray(len(text.split(' ')))
max_len = data['max_len']
# Map words to their corresponding word embedding indices
text_vec = []
for w in text.split(' '):
if w in w2v.vocab:
text_vec.append(w2v.vocab[w].index)
else:
print('Word %s not in vocabulary!' % w)
break
text_vec = np.asarray(text_vec)
print('Sentence with w2v indices:')
print(text_vec)
# Pad sentence with 0s so that it fills the maximum sentence length
text_vec_pad = np.append(text_vec, np.zeros((max_len - text_len,), dtype=np.int64))
print('Padded sentence:')
print(text_vec_pad)
```
Expand dimensions of the input data to fit the dimensions that the model expects:
<br>
`input_x` - shape (batch_size, max_len)
<br>
`sentence_len` - shape (batch_size,)
```
print('text_vec_pad shape: ' + str(text_vec_pad.shape))
text_vec_pad_dim = np.expand_dims(text_vec_pad, axis=0)
print('text_vec_pad_dim shape: ' + str(text_vec_pad_dim.shape))
print('text_len shape: ' + str(text_len.shape))
text_len_dim = np.expand_dims(text_len, axis=0)
print('text_len_dim shape: ' + str(text_len_dim.shape))
```
## Obtain prediction
```
import util
# Feed the sample to the model to obtain slot and intent predictions
[intent_outputs, slots_outputs, slot_weights_c] = sess.run([
capsnet.intent_output_vectors, capsnet.slot_output_vectors, capsnet.slot_weights_c],
feed_dict={capsnet.input_x: text_vec_pad_dim, capsnet.sentences_length: text_len_dim,
capsnet.keep_prob: 1.0})
intent_outputs_reduced_dim = tf.squeeze(intent_outputs, axis=[1, 4])
intent_outputs_norm = util.safe_norm(intent_outputs_reduced_dim)
slot_weights_c_reduced_dim = tf.squeeze(slot_weights_c, axis=[3, 4])
[intent_predictions, slot_predictions] = sess.run([intent_outputs_norm, slot_weights_c_reduced_dim])
```
Each of the **13 IntentCaps** output a **16-dimensional vector**.
<br>
For slot filling, we are interested in the **routing weights** between **WordCaps** and **SlotCaps** -- in this way we predict the slot type of each individual word. There is a routing weight associated to each word capsule - slot capsule pair (15 x 8 total)
```
print('IntentCaps output shape: ' + str(intent_outputs.shape))
print('WordCaps - SlotCaps routing weights: ' + str(slot_weights_c.shape))
intent_pred = np.argmax(intent_predictions, axis=1)
slots_pred = np.argmax(slot_predictions, axis=2)
intent_label = data['intents_dict'][intent_pred[0]]
slot_labels = [data['slots_dict'][x] for x in slots_pred[0]]
print(text)
print('Intent prediction: ' + intent_label)
print('Slots prediction: ' + str(slot_labels[:text_len]))
```
## More predictions
```
text = 'Da drumul la lumina in hol'
text_len = np.asarray(len(text.split(' ')))
max_len = data['max_len']
# Map words to their corresponding word embedding indices
text_vec = []
for w in text.split(' '):
if w in w2v.vocab:
text_vec.append(w2v.vocab[w].index)
else:
print('Word %s not in vocabulary!' % w)
break
text_vec = np.asarray(text_vec)
# Pad sentence with 0s so that it fills the maximum sentence length
text_vec_pad = np.append(text_vec, np.zeros((max_len - text_len,), dtype=np.int64))
text_vec_pad_dim = np.expand_dims(text_vec_pad, axis=0)
text_len_dim = np.expand_dims(text_len, axis=0)
# Feed the sample to the model to obtain slot and intent predictions
[intent_outputs, slots_outputs, slot_weights_c] = sess.run([
capsnet.intent_output_vectors, capsnet.slot_output_vectors, capsnet.slot_weights_c],
feed_dict={capsnet.input_x: text_vec_pad_dim, capsnet.sentences_length: text_len_dim,
capsnet.keep_prob: 1.0})
intent_outputs_reduced_dim = tf.squeeze(intent_outputs, axis=[1, 4])
intent_outputs_norm = util.safe_norm(intent_outputs_reduced_dim)
slot_weights_c_reduced_dim = tf.squeeze(slot_weights_c, axis=[3, 4])
[intent_predictions, slot_predictions] = sess.run([intent_outputs_norm, slot_weights_c_reduced_dim])
intent_pred = np.argmax(intent_predictions, axis=1)
slots_pred = np.argmax(slot_predictions, axis=2)
intent_label = data['intents_dict'][intent_pred[0]]
slot_labels = [data['slots_dict'][x] for x in slots_pred[0]]
print(text)
print('Intent prediction: ' + intent_label)
print('Slots prediction: ' + str(slot_labels[:text_len]))
```
| github_jupyter |
## Imports
```
pip install torchviz
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
# from torch.optim.lr_scheduler import StepLR, ReduceLROnPlateau, CosineAnnealingLR
from torchvision import datasets, transforms
from torchviz import make_dot
import os
import time
import random
import networkx as nx
import yaml
import matplotlib.pyplot as plt
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
```
## helper functions
```
def plot_results(list_of_epochs, list_of_train_losses, list_of_train_accuracies, list_of_val_accuracies):
plt.figure(figsize=(20, 9))
plt.subplot(1, 2, 1)
plt.plot(list_of_epochs, list_of_train_losses, label='training loss')
plt.legend()
plt.subplot(1, 2, 2)
plt.plot(list_of_epochs, list_of_train_accuracies, label='training accuracy')
plt.plot(list_of_epochs, list_of_val_accuracies, label='validation accuracy')
plt.legend()
if not os.path.isdir('./result_plots'):
os.makedirs('./result_plots')
plt.savefig('./result_plots/accuracy_plot_per_epoch.jpg')
plt.close()
```
## training routine
```
def set_lr(optim, epoch_num, lrate):
"""adjusts lr to starting lr thereafter reduced by 10% at every 20 epochs"""
lrate = lrate * (0.1 ** (epoch_num // 20))
for params in optim.param_groups:
params['lr'] = lrate
def train(model, train_dataloader, optim, loss_func, epoch_num, lrate):
model.train()
loop_iter = 0
training_loss = 0
training_accuracy = 0
for training_data, training_label in train_dataloader:
set_lr(optim, epoch_num, lrate)
training_data, training_label =\
training_data.to(device), training_label.to(device)
optim.zero_grad()
pred_raw = model(training_data)
curr_loss = loss_func(pred_raw, training_label)
curr_loss.backward()
optim.step()
training_loss += curr_loss.data
pred = pred_raw.data.max(1)[1]
curr_accuracy =\
float(pred.eq(training_label.data).sum()) * 100. / len(training_data)
training_accuracy += curr_accuracy
loop_iter += 1
if loop_iter % 100 == 0:
print(
f"epoch {epoch_num}, loss: {curr_loss.data}, accuracy: {curr_accuracy}")
data_size = len(train_dataloader.dataset) // batch_size
return training_loss / data_size, training_accuracy / data_size
```
## accuracy metric
```
def accuracy(model, test_data_loader):
model.eval()
success = 0
with torch.no_grad():
for test_data, test_label in test_data_loader:
test_data, test_label = test_data.to(device), test_label.to(device)
pred_raw = model(test_data)
pred = pred_raw.data.max(1)[1]
success += pred.eq(test_label.data).sum()
return float(success) * 100. / len(test_data_loader.dataset)
```
## data loader and load data
```
batch_size = 64
def load_dataset(batch_size):
transform_train_dataset = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.4983, 0.4795, 0.4382), (0.2712, 0.2602, 0.2801)),
])
transform_test_dataset = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.4983, 0.4795, 0.4382), (0.2712, 0.2602, 0.2801)),
])
train_dataloader = torch.utils.data.DataLoader(
datasets.CIFAR10('dataset', transform=transform_train_dataset,
train=True, download=True),
batch_size=batch_size,
shuffle=True
)
test_dataloader = torch.utils.data.DataLoader(
datasets.CIFAR10('dataset', transform=transform_test_dataset,
train=False),
batch_size=batch_size,
shuffle=False
)
return train_dataloader, test_dataloader
train_dataloader, test_dataloader = load_dataset(batch_size)
```
## graph class def
```
class RndGraph(object):
def __init__(self, num_nodes, graph_probability, nearest_neighbour_k=4,
num_edges_attach=5):
self.num_nodes = num_nodes
self.graph_probability = graph_probability
self.nearest_neighbour_k = nearest_neighbour_k
self.num_edges_attach = num_edges_attach
def make_graph_obj(self):
graph_obj = nx.random_graphs.connected_watts_strogatz_graph(
self.num_nodes, self.nearest_neighbour_k, self.graph_probability)
return graph_obj
def get_graph_config(self, graph_obj):
incoming_edges = {}
incoming_edges[0] = []
node_list = [0]
last = []
for n in graph_obj.nodes():
neighbor_list = list(graph_obj.neighbors(n))
neighbor_list.sort()
edge_list = []
passed_list = []
for nbr in neighbor_list:
if n > nbr:
edge_list.append(nbr + 1)
passed_list.append(nbr)
if not edge_list:
edge_list.append(0)
incoming_edges[n + 1] = edge_list
if passed_list == neighbor_list:
last.append(n + 1)
node_list.append(n + 1)
incoming_edges[self.num_nodes + 1] = last
node_list.append(self.num_nodes + 1)
return node_list, incoming_edges
def save_graph(self, graph_obj, path_to_write):
if not os.path.isdir("cached_graph_obj"):
os.mkdir("cached_graph_obj")
#nx.write_yaml(graph_obj, "./cached_graph_obj/" + path_to_write)
with open("./cached_graph_obj/" + path_to_write, 'w') as fh:
yaml.dump(graph_obj, fh)
def load_graph(self, path_to_read):
#return nx.read_yaml("./cached_graph_obj/" + path_to_read)
with open("./cached_graph_obj/" + path_to_read, 'r') as fh:
return yaml.load(fh, Loader=yaml.Loader)
```
## randwire def
```
def initialize_weights(layer):
if isinstance(layer, nn.Conv2d):
torch.nn.init.xavier_uniform_(layer.weight)
if layer.bias is not None:
torch.nn.init.zeros_(layer.bias)
class SepConv2d(nn.Module):
def __init__(self, input_ch, output_ch, kernel_length=3, dilation_size=1,
padding_size=1, stride_length=1, bias_flag=True):
super(SepConv2d, self).__init__()
self.conv_layer = nn.Conv2d(input_ch, input_ch, kernel_length,
stride_length, padding_size, dilation_size,
bias=bias_flag, groups=input_ch)
self.pointwise_layer = nn.Conv2d(input_ch, output_ch, kernel_size=1,
stride=1, padding=0, dilation=1,
groups=1, bias=bias_flag)
def forward(self, x):
return self.pointwise_layer(self.conv_layer(x))
class UnitLayer(nn.Module):
def __init__(self, input_ch, output_ch, stride_length=1):
super(UnitLayer, self).__init__()
self.dropout = 0.3
self.unit_layer = nn.Sequential(
nn.ReLU(),
SepConv2d(input_ch, output_ch, stride_length=stride_length),
nn.BatchNorm2d(output_ch),
nn.Dropout(self.dropout)
)
def forward(self, x):
return self.unit_layer(x)
class GraphNode(nn.Module):
def __init__(self, input_degree, input_ch, output_ch, stride_length=1):
super(GraphNode, self).__init__()
self.input_degree = input_degree
if len(self.input_degree) > 1:
self.params = nn.Parameter(torch.ones(
len(self.input_degree), requires_grad=True))
self.unit_layer = UnitLayer(
input_ch, output_ch, stride_length=stride_length)
def forward(self, *ip):
if len(self.input_degree) > 1:
op = (ip[0] * torch.sigmoid(self.params[0]))
for idx in range(1, len(ip)):
op += (ip[idx] * torch.sigmoid(self.params[idx]))
return self.unit_layer(op)
else:
return self.unit_layer(ip[0])
class RandWireGraph(nn.Module):
def __init__(self, num_nodes, graph_prob, input_ch, output_ch, train_mode,
graph_name):
super(RandWireGraph, self).__init__()
self.num_nodes = num_nodes
self.graph_prob = graph_prob
self.input_ch = input_ch
self.output_ch = output_ch
self.train_mode = train_mode
self.graph_name = graph_name
# get graph nodes and in edges
rnd_graph_node = RndGraph(self.num_nodes, self.graph_prob)
if self.train_mode is True:
print("train_mode: ON")
rnd_graph = rnd_graph_node.make_graph_obj()
self.node_list, self.incoming_edge_list =\
rnd_graph_node.get_graph_config(rnd_graph)
rnd_graph_node.save_graph(rnd_graph, graph_name)
else:
rnd_graph = rnd_graph_node.load_graph(graph_name)
self.node_list, self.incoming_edge_list =\
rnd_graph_node.get_graph_config(rnd_graph)
# define input Node
self.list_of_modules = nn.ModuleList(
[GraphNode(self.incoming_edge_list[0], self.input_ch,
self.output_ch, stride_length=2)])
# define the rest Node
self.list_of_modules.extend(
[GraphNode(self.incoming_edge_list[n], self.output_ch,
self.output_ch) for n in self.node_list if n > 0])
def forward(self, x):
mem_dict = {}
# start vertex
op = self.list_of_modules[0].forward(x)
mem_dict[0] = op
# the rest vertex
for n in range(1, len(self.node_list) - 1):
# print(node, self.in_edges[node][0], self.in_edges[node])
if len(self.incoming_edge_list[n]) > 1:
op = self.list_of_modules[n].forward(
*[mem_dict[incoming_vtx] for incoming_vtx
in self.incoming_edge_list[n]])
else:
op = self.list_of_modules[n].forward(
mem_dict[self.incoming_edge_list[n][0]])
mem_dict[n] = op
op = mem_dict[self.incoming_edge_list[self.num_nodes + 1][0]]
for incoming_vtx in range(
1, len(self.incoming_edge_list[self.num_nodes + 1])):
op += mem_dict[
self.incoming_edge_list[self.num_nodes + 1][incoming_vtx]]
return op / len(self.incoming_edge_list[self.num_nodes + 1])
```
## randwire NN model def
```
class RandWireNNModel(nn.Module):
def __init__(self, num_nodes, graph_prob, input_ch, output_ch, train_mode):
super(RandWireNNModel, self).__init__()
self.num_nodes = num_nodes
self.graph_prob = graph_prob
self.input_ch = input_ch
self.output_ch = output_ch
self.train_mode = train_mode
self.dropout = 0.3
self.class_num = 10
self.conv_layer_1 = nn.Sequential(
nn.Conv2d(in_channels=3, out_channels=self.output_ch,
kernel_size=3, padding=1),
nn.BatchNorm2d(self.output_ch),
)
self.conv_layer_2 = nn.Sequential(
RandWireGraph(self.num_nodes, self.graph_prob, self.input_ch,
self.output_ch*2, self.train_mode,
graph_name="conv_layer_2")
)
self.conv_layer_3 = nn.Sequential(
RandWireGraph(self.num_nodes, self.graph_prob, self.input_ch*2,
self.output_ch*4, self.train_mode,
graph_name="conv_layer_3")
)
self.conv_layer_4 = nn.Sequential(
RandWireGraph(self.num_nodes, self.graph_prob, self.input_ch*4,
self.output_ch*8, self.train_mode,
graph_name="conv_layer_4")
)
self.classifier_layer = nn.Sequential(
nn.Conv2d(in_channels=self.input_ch*8, out_channels=1280,
kernel_size=1),
nn.BatchNorm2d(1280)
)
self.output_layer = nn.Sequential(
nn.Dropout(self.dropout),
nn.Linear(1280, self.class_num)
)
def forward(self, x):
x = self.conv_layer_1(x)
x = self.conv_layer_2(x)
x = self.conv_layer_3(x)
x = self.conv_layer_4(x)
x = self.classifier_layer(x)
# global average pooling
_, _, h, w = x.size()
x = F.avg_pool2d(x, kernel_size=[h, w])
x = torch.squeeze(x)
x = self.output_layer(x)
return x
```
## hyperparams initialization
```
num_epochs = 5
graph_probability = 0.7
node_channel_count = 64
num_nodes = 16
lrate = 0.1
#batch_size = 64
train_mode = True
```
## training loop
```
rand_wire_model = RandWireNNModel(num_nodes, graph_probability,
node_channel_count, node_channel_count, train_mode).to(device)
optim_module = optim.SGD(rand_wire_model.parameters(), lr=lrate,
weight_decay=1e-4, momentum=0.8)
loss_func = nn.CrossEntropyLoss().to(device)
epochs = []
test_accuracies = []
training_accuracies = []
training_losses = []
best_test_accuracy = 0
start_time = time.time()
for ep in range(1, num_epochs + 1):
epochs.append(ep)
training_loss, training_accuracy = train(rand_wire_model, train_dataloader,
optim_module, loss_func, ep, lrate)
test_accuracy = accuracy(rand_wire_model, test_dataloader)
test_accuracies.append(test_accuracy)
training_losses.append(training_loss.cpu())
training_accuracies.append(training_accuracy)
print('test acc: {0:.2f}%, best test acc: {1:.2f}%'.format(
test_accuracy, best_test_accuracy))
if best_test_accuracy < test_accuracy:
model_state = {
'model': rand_wire_model.state_dict(),
'accuracy': test_accuracy,
'ep': ep,
}
if not os.path.isdir('model_checkpoint'):
os.mkdir('model_checkpoint')
model_filename = "ch_count_" + str(node_channel_count) + "_prob_" +\
str(graph_probability)
torch.save(model_state,
'./model_checkpoint/' + model_filename + 'ckpt.t7')
best_test_accuracy = test_accuracy
plot_results(epochs, training_losses, training_accuracies,
test_accuracies)
print("model train time: ", time.time() - start_time)
```
## test model
```
def num_model_params(model_obj):
num_params = 0
for l in list(model_obj.parameters()):
l_p = 1
for p in list(l.size()):
l_p *= p
num_params += l_p
return num_params
print("total model params: ", num_model_params(rand_wire_model))
if os.path.exists("./model_checkpoint"):
rand_wire_nn_model = RandWireNNModel(num_nodes, graph_probability, node_channel_count, node_channel_count,
train_mode=False).to(device)
model_filename = "ch_count_" + str(node_channel_count) + "_prob_" + str(graph_probability)
model_checkpoint = torch.load('./model_checkpoint/' + model_filename + 'ckpt.t7')
rand_wire_nn_model.load_state_dict(model_checkpoint['model'])
last_ep = model_checkpoint['ep']
best_model_accuracy = model_checkpoint['accuracy']
print(f"best model accuracy: {best_model_accuracy}%, last epoch: {last_ep}")
rand_wire_nn_model.eval()
success = 0
for test_data, test_label in test_dataloader:
test_data, test_label = test_data.to(device), test_label.to(device)
pred_raw = rand_wire_nn_model(test_data)
pred = pred_raw.data.max(1)[1]
success += pred.eq(test_label.data).sum()
print(f"test accuracy: {float(success) * 100. / len(test_dataloader.dataset)} %")
else:
assert False, "File not found. Please check again."
```
## visualize model graph
```
x = torch.randn(2, 3, 32, 32, device=device)
y = rand_wire_nn_model(x)
g = make_dot(y.mean(), params=dict(rand_wire_nn_model.named_parameters()))
g.format='svg'
g.filename = 'image2'
g.render(view=False)
g
```
| github_jupyter |
```
#!pip install rouge
import pandas as pd
def scoring(bertdf,userdf):
#1.1 clean up bert df, extract the first row, and reset index
bertsum = bertdf.iloc[0:1]
bertsum =bertsum.to_string(header=False,index=False,index_names=False)
bertsum= [int(s) for s in bertsum.split(',')]
bertdf=bertdf.drop(0).reset_index(drop=True)
#1.2 clean up user df, extract the first row, and reset index
usersum = userdf.iloc[0:1]
usersum =usersum.to_string(header=False,index=False,index_names=False)
usersum= [int(s) for s in usersum.split(',')]
userdf=userdf.drop(0).reset_index(drop=True)
#2.1 convert to list
berttitle= bertdf.columns[0]
cleanedbert =bertdf.iloc[bertsum , : ]
bertdf =cleanedbert[berttitle].tolist()
bert = ' '.join([str(elem) for elem in bertdf])
#2.2 convert to list
usertitle= userdf.columns[0]
cleaneduser =userdf.iloc[usersum , : ]
userdf =cleaneduser[usertitle].tolist()
user = ' '.join([str(elem) for elem in userdf])
#scoring, we use rouge-l (ROUGE-L: Longest Common Subsequence based statistics, takes sentences into account)
from rouge import Rouge
rouge = Rouge()
scores = rouge.get_scores(bert, user)
f_score =scores[0]["rouge-l"]["f"]
precision =scores[0]["rouge-l"]["p"]
recall =scores[0]["rouge-l"]["r"]
#print(f1,precision,recall)
print("BERT: "+str(bertsum) +"\nUSER: "+str(usersum))
print("\n\nROUGE scoring:\n\n"+
"Precision is :"+"{:.2%}".format(precision)+
"\nRecall is :"+"{:.2%}".format(recall)+
"\nF Score is :"+"{:.2%}".format(f_score))
print("\nPrecision: how much BERT summary exceeds human summary, (if less than 100% means user removed sentences)\n"
"Recall: how much BERT summary explains the human summary, (if less than 100% means user added sentences)\n"
"F Score: aggregation of BERT performance,(if 100% means perfect match)")
return
#input 2 csv file and convert it to dataframe
bert = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testbert.csv")
user = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testuser_same.csv")
print("SUMMARY PERFECT: If summariser is same as human: \n")
#scoring (machine, human) - this order is important
scoring(bert,user)
#input 2 csv file and convert it to dataframe
bert = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testbert.csv")
user = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testuser_add.csv")
print("SUMMARISER NOT PERFECT: BERT has less sentences: \n")
#scoring (machine, human) - this order is important
scoring(bert,user)
#input 2 csv file and convert it to dataframe
bert = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testbert.csv")
user = pd.read_csv(r"C:\Users\User\Desktop\TIPP\11 NVidia project\data\testuser_minus.csv")
print("SUMMARISER NOT PERFECT: BERT has more sentences: \n")
#scoring (machine, human) - this order is important
scoring(bert,user)
```
| github_jupyter |
#**Part 1 - Data gathering and preprocessing**
**Libraries**
```
import numpy as np #Linear_Algebra
import matplotlib.pyplot as plt
import pandas as pd #Data_Processing
import pandas_datareader as pdr
from scipy import stats
%matplotlib inline
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
pip install -q yfinance --upgrade
#Import Yahoo Finance
import yfinance as yf
yf.pdr_override()
#CISCO data
SELECTED_STOCK = 'CSCO'
start = '2010-12-17'
end = '2019-12-17'
#Download NVIDIA stock price data for the past 10 yrs to date
stock_data = pdr.get_data_yahoo(SELECTED_STOCK, start, end)
stock_data.head(10)
```
**Feature Engineering**
```
#Getting the Open price
stock_data_open = stock_data.Open.values
reshaped_stock_data_open = np.reshape(stock_data_open, (-1, 1))
reshaped_stock_data_open
#validity check
np.mean(reshaped_stock_data_open)==np.mean(stock_data_open)
```
**Feature Scaling**
```
from sklearn.preprocessing import MinMaxScaler
sc = MinMaxScaler(feature_range = (0,1))
scaled_data = sc.fit_transform(reshaped_stock_data_open)
def timestamp(n_period, scaled_data):
x_train = []
y_train = [] #1 output to predict
for i in range(n_period,len(scaled_data)):
x_train.append(scaled_data[i-n_period:i,0])
y_train.append(scaled_data[i,0])
x_train, y_train = np.array(x_train), np.array(y_train)
#reshaping
x_train = np.reshape(x_train, (x_train.shape[0], x_train.shape[1], 1))
return x_train, y_train
x_train, y_train = timestamp(60, scaled_data)
```
#**Part 2 - Building the RNN**
```
import warnings
warnings.simplefilter("ignore")
#Importing the keras libraries and packages
from tensorflow.python.keras.layers import Dense, LSTM, Dropout
from tensorflow.python.keras import Sequential
regressor = Sequential()
#Adding the first LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True, input_shape = (x_train.shape[1], 1)))
regressor.add(Dropout(rate = 0.2))
x_train.shape[1]
#Adding the second LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True))
regressor.add(Dropout(rate = 0.2))
#Adding the third LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50, return_sequences=True))
regressor.add(Dropout(rate = 0.2))
#Adding the fourth LSTM Layer and some Dropout regularisation
regressor.add(LSTM(units=50))
regressor.add(Dropout(rate = 0.2))
#Adding the output layer
regressor.add(Dense(units=1))
#compiling the RNN
regressor.compile(optimizer='adam', loss='mean_squared_error')
#fitting the RNN to the training set
regressor.fit(x_train, y_train, epochs=50, batch_size = 32)
```
**Save the model**
```
regressor = regressor.save("regressor.h5")
```
**Load the model**
```
from tensorflow.python.keras.models import load_model
regressor = load_model("regressor.h5")
```
#**Part 3 - Making the predictions and visualising the results**
| github_jupyter |
# Example use case of the NGM
This notebook provides a simple use case of the Neural Graphical Modelling (NGM) in continuous-time for time series data sampled from an underlying dynamical system, based on our paper *"Graphical modelling in continuous-time: consistency guarantees and algorithms using Neural ODEs"*.
## What is the neural graphical modelling method?
It is a continuous-time alternative to the more common discrete-time graphical models, such as Granger causality. While appealing due to their simplicity, discrete-time methods are at odds with the continuous-time nature of the unfolding processes over time, and cumbersome to interpret with irregularly sampled time series or with processes observed at a scale that does not reflect that of causal associations.
Here, we propose graphical modelling by explicitly considering the latent mechanism continuously in time using the mathematics of differential equations. The objective then is to consistently recover interactions between stochastic processes in infinitesimal intervals of time. The key advantages of the resulting graphical model are that it is directly applicable to the general setting of irregularly-sampled multivariate time series, it does not impose constraints on the non-linearities or dimensionality of the underlying system, and under appropriate conditions, identifies associations exactly.
Let us import some necessary packages before illustrating the use of NGM.
```
import numpy as np
import torch
import sys
import matplotlib.pyplot as plt
sys.path.append("../")
from utils import simulate_lorenz_96, compare_graphs
import NMC as models
import importlib
```
We will use the Lorenz model as the underlying data generating mechanism, with 10 variables and 1000 regularly-sampled observations.
```
# Simulate data.
p = 10
T = 1000
num_points = T
data, GC = simulate_lorenz_96(p, T=T, sigma=1, delta_t=0.05, sd=0.0, F=5)
# Format for NeuralODE.
times = np.linspace(0, T, num_points)
times_np = np.hstack([times[:, None]])
times = torch.from_numpy(times_np[:, :, None].astype(np.float32))
data = torch.from_numpy(data[:, None, :].astype(np.float32))
```
Let us plot the data. The right-most panel is the binary matrix indicating the presence / absence of edges in the underlying graph of dependencies. This is our target.
```
fig, axs = plt.subplots(1, 2, figsize=(7, 2.3))
fig.tight_layout(pad=0.2, w_pad=2, h_pad=3)
axs[0].plot(data[:100].squeeze())
cax = axs[1].matshow(GC)
fig.colorbar(cax)
plt.show()
```
We will now define the model
```
# Specify device.
device = "cpu" # Select device: "cpu", "cuda", "cuda:1", etc.
device = torch.device(device)
importlib.reload(models)
# Function specification.
func = models.MLPODEF(dims=[p, 12, 1], GL_reg=0.1)
func = func.to(device)
```
And train using adjoint backpropagation to approximate the observed trajectories. The intermediate plots during training show in the leftmost panel: the observed data, in the middle panel: the approximated trajectories, in the rightmost panel: the approximated causal structure.
```
importlib.reload(models)
# Group Lasso training.
models.train(func, data, n_steps=1000, plot_freq=20, device=device)
# Adaptive Group lasso training.
weights = func.group_weights()
func.GL_reg *= 1 / weights
func.reset_parameters()
models.train(func, data, n_steps=1000, plot_freq=20, device=device)
```
Let us inspect the recovered graph and compute true positive rates (TPR) and false discovery rates (FDR)
```
# Inspect estimated causal graph and SHD computation.
W_est = func.causal_graph(w_threshold=0.0)
print("Estimated strength of interactions between stochastic processes")
print(W_est)
print("")
print("TPR and FDR of our estimate with respect to truth:")
compare_graphs(GC, W_est)
```
A plot is perhaps more informative
```
graph = func.causal_graph(w_threshold=0.0)
plt.matshow(graph, cmap="Reds")
plt.colorbar()
plt.show()
```
| github_jupyter |
<!--BOOK_INFORMATION-->
<img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
*This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by Jake VanderPlas; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
*The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
*No changes were made to the contents of this notebook from the original.*
<!--NAVIGATION-->
< [Aggregations: Min, Max, and Everything In Between](02.04-Computation-on-arrays-aggregates.ipynb) | [Contents](Index.ipynb) | [Comparisons, Masks, and Boolean Logic](02.06-Boolean-Arrays-and-Masks.ipynb) >
# Computation on Arrays: Broadcasting
We saw in the previous section how NumPy's universal functions can be used to *vectorize* operations and thereby remove slow Python loops.
Another means of vectorizing operations is to use NumPy's *broadcasting* functionality.
Broadcasting is simply a set of rules for applying binary ufuncs (e.g., addition, subtraction, multiplication, etc.) on arrays of different sizes.
## Introducing Broadcasting
Recall that for arrays of the same size, binary operations are performed on an element-by-element basis:
```
import numpy as np
a = np.array([0, 1, 2])
b = np.array([5, 5, 5])
a + b
```
Broadcasting allows these types of binary operations to be performed on arrays of different sizes–for example, we can just as easily add a scalar (think of it as a zero-dimensional array) to an array:
```
a + 5
```
We can think of this as an operation that stretches or duplicates the value ``5`` into the array ``[5, 5, 5]``, and adds the results.
The advantage of NumPy's broadcasting is that this duplication of values does not actually take place, but it is a useful mental model as we think about broadcasting.
We can similarly extend this to arrays of higher dimension. Observe the result when we add a one-dimensional array to a two-dimensional array:
```
M = np.ones((3, 3))
M
M + a
```
Here the one-dimensional array ``a`` is stretched, or broadcast across the second dimension in order to match the shape of ``M``.
While these examples are relatively easy to understand, more complicated cases can involve broadcasting of both arrays. Consider the following example:
```
a = np.arange(3)
b = np.arange(3)[:, np.newaxis]
print(a)
print(b)
a + b
```
Just as before we stretched or broadcasted one value to match the shape of the other, here we've stretched *both* ``a`` and ``b`` to match a common shape, and the result is a two-dimensional array!
The geometry of these examples is visualized in the following figure (Code to produce this plot can be found in the [appendix](06.00-Figure-Code.ipynb#Broadcasting), and is adapted from source published in the [astroML](http://astroml.org) documentation. Used by permission).

The light boxes represent the broadcasted values: again, this extra memory is not actually allocated in the course of the operation, but it can be useful conceptually to imagine that it is.
## Rules of Broadcasting
Broadcasting in NumPy follows a strict set of rules to determine the interaction between the two arrays:
- Rule 1: If the two arrays differ in their number of dimensions, the shape of the one with fewer dimensions is *padded* with ones on its leading (left) side.
- Rule 2: If the shape of the two arrays does not match in any dimension, the array with shape equal to 1 in that dimension is stretched to match the other shape.
- Rule 3: If in any dimension the sizes disagree and neither is equal to 1, an error is raised.
To make these rules clear, let's consider a few examples in detail.
### Broadcasting example 1
Let's look at adding a two-dimensional array to a one-dimensional array:
```
M = np.ones((2, 3))
a = np.arange(3)
```
Let's consider an operation on these two arrays. The shape of the arrays are
- ``M.shape = (2, 3)``
- ``a.shape = (3,)``
We see by rule 1 that the array ``a`` has fewer dimensions, so we pad it on the left with ones:
- ``M.shape -> (2, 3)``
- ``a.shape -> (1, 3)``
By rule 2, we now see that the first dimension disagrees, so we stretch this dimension to match:
- ``M.shape -> (2, 3)``
- ``a.shape -> (2, 3)``
The shapes match, and we see that the final shape will be ``(2, 3)``:
```
M + a
```
### Broadcasting example 2
Let's take a look at an example where both arrays need to be broadcast:
```
a = np.arange(3).reshape((3, 1))
b = np.arange(3)
```
Again, we'll start by writing out the shape of the arrays:
- ``a.shape = (3, 1)``
- ``b.shape = (3,)``
Rule 1 says we must pad the shape of ``b`` with ones:
- ``a.shape -> (3, 1)``
- ``b.shape -> (1, 3)``
And rule 2 tells us that we upgrade each of these ones to match the corresponding size of the other array:
- ``a.shape -> (3, 3)``
- ``b.shape -> (3, 3)``
Because the result matches, these shapes are compatible. We can see this here:
```
a + b
```
### Broadcasting example 3
Now let's take a look at an example in which the two arrays are not compatible:
```
M = np.ones((3, 2))
a = np.arange(3)
```
This is just a slightly different situation than in the first example: the matrix ``M`` is transposed.
How does this affect the calculation? The shape of the arrays are
- ``M.shape = (3, 2)``
- ``a.shape = (3,)``
Again, rule 1 tells us that we must pad the shape of ``a`` with ones:
- ``M.shape -> (3, 2)``
- ``a.shape -> (1, 3)``
By rule 2, the first dimension of ``a`` is stretched to match that of ``M``:
- ``M.shape -> (3, 2)``
- ``a.shape -> (3, 3)``
Now we hit rule 3–the final shapes do not match, so these two arrays are incompatible, as we can observe by attempting this operation:
```
M + a
```
Note the potential confusion here: you could imagine making ``a`` and ``M`` compatible by, say, padding ``a``'s shape with ones on the right rather than the left.
But this is not how the broadcasting rules work!
That sort of flexibility might be useful in some cases, but it would lead to potential areas of ambiguity.
If right-side padding is what you'd like, you can do this explicitly by reshaping the array (we'll use the ``np.newaxis`` keyword introduced in [The Basics of NumPy Arrays](02.02-The-Basics-Of-NumPy-Arrays.ipynb)):
```
a[:, np.newaxis].shape
M + a[:, np.newaxis]
```
Also note that while we've been focusing on the ``+`` operator here, these broadcasting rules apply to *any* binary ``ufunc``.
For example, here is the ``logaddexp(a, b)`` function, which computes ``log(exp(a) + exp(b))`` with more precision than the naive approach:
```
np.logaddexp(M, a[:, np.newaxis])
```
For more information on the many available universal functions, refer to [Computation on NumPy Arrays: Universal Functions](02.03-Computation-on-arrays-ufuncs.ipynb).
## Broadcasting in Practice
Broadcasting operations form the core of many examples we'll see throughout this book.
We'll now take a look at a couple simple examples of where they can be useful.
### Centering an array
In the previous section, we saw that ufuncs allow a NumPy user to remove the need to explicitly write slow Python loops. Broadcasting extends this ability.
One commonly seen example is when centering an array of data.
Imagine you have an array of 10 observations, each of which consists of 3 values.
Using the standard convention (see [Data Representation in Scikit-Learn](05.02-Introducing-Scikit-Learn.ipynb#Data-Representation-in-Scikit-Learn)), we'll store this in a $10 \times 3$ array:
```
X = np.random.random((10, 3))
```
We can compute the mean of each feature using the ``mean`` aggregate across the first dimension:
```
Xmean = X.mean(0)
Xmean
```
And now we can center the ``X`` array by subtracting the mean (this is a broadcasting operation):
```
X_centered = X - Xmean
```
To double-check that we've done this correctly, we can check that the centered array has near zero mean:
```
X_centered.mean(0)
```
To within machine precision, the mean is now zero.
### Plotting a two-dimensional function
One place that broadcasting is very useful is in displaying images based on two-dimensional functions.
If we want to define a function $z = f(x, y)$, broadcasting can be used to compute the function across the grid:
```
# x and y have 50 steps from 0 to 5
x = np.linspace(0, 5, 50)
y = np.linspace(0, 5, 50)[:, np.newaxis]
z = np.sin(x) ** 10 + np.cos(10 + y * x) * np.cos(x)
```
We'll use Matplotlib to plot this two-dimensional array (these tools will be discussed in full in [Density and Contour Plots](04.04-Density-and-Contour-Plots.ipynb)):
```
%matplotlib inline
import matplotlib.pyplot as plt
plt.imshow(z, origin='lower', extent=[0, 5, 0, 5],
cmap='viridis')
plt.colorbar();
```
The result is a compelling visualization of the two-dimensional function.
<!--NAVIGATION-->
< [Aggregations: Min, Max, and Everything In Between](02.04-Computation-on-arrays-aggregates.ipynb) | [Contents](Index.ipynb) | [Comparisons, Masks, and Boolean Logic](02.06-Boolean-Arrays-and-Masks.ipynb) >
| github_jupyter |
```
import argparse
import logging
import os
import torch
import logging
from torch.utils.data import DataLoader, TensorDataset
from torch.utils.data.distributed import DistributedSampler
from torch.utils.data.sampler import RandomSampler, SequentialSampler
import sys
sys.path.append(os.path.join(os.path.dirname("__file__"), '..'))
import time
import random
import torch.nn.functional as F
from decode_abstract_models import *
from seq2seq.ReaSCAN_dataset import *
from seq2seq.helpers import *
from torch.optim.lr_scheduler import LambdaLR
def isnotebook():
try:
shell = get_ipython().__class__.__name__
if shell == 'ZMQInteractiveShell':
return True # Jupyter notebook or qtconsole
elif shell == 'TerminalInteractiveShell':
return False # Terminal running IPython
else:
return False # Other type (?)
except NameError:
return False # Probably standard Python interpreter
path_to_data = "../../../data-files/gSCAN-Simple/data-compositional-splits.txt"
data_json = json.load(open(path_to_data, "r"))
NUM = 200
agent_positions_batch = []
target_positions_batch = []
target_commands = []
for ex in data_json["examples"]["situational_1"]:
target_commands += [ex["target_commands"]]
situation_repr = ex['situation']
agent = torch.tensor(
(int(situation_repr["agent_position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["agent_position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
target = torch.tensor(
(int(situation_repr["target_object"]["position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["target_object"]["position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
agent_positions_batch.append(agent)
target_positions_batch.append(target)
if len(agent_positions_batch) == NUM:
break
agent_positions_batch = torch.stack(agent_positions_batch, dim=0)
target_positions_batch = torch.stack(target_positions_batch, dim=0)
hi_model = HighLevelModel()
hidden_states = hi_model(agent_positions_batch, target_positions_batch, tag="situation_encode")
actions = torch.zeros(hidden_states.size(0), 1).long()
actions_sequence = []
actions_length = torch.zeros(hidden_states.size(0), 1).long()
for i in range(1):
hidden_states, actions = hi_model(
hmm_states=hidden_states,
hmm_actions=actions,
tag="_hmm_step_fxn"
)
actions_length += (actions!=0).long()
actions_sequence += [actions]
grid_size = 6
x_target = torch.zeros(hidden_states.shape[0], (grid_size*2-1)).long()
y_target = torch.zeros(hidden_states.shape[0], (grid_size*2-1)).long()
indices = hidden_states + 5
x_target[range(x_target.shape[0]), indices[:,0]] = 1
y_target[range(y_target.shape[0]), indices[:,1]] = 1
actions_sequence = torch.cat(actions_sequence, dim=-1)
for i in range(actions_sequence.size(0)):
pred = (hi_model.actions_list_to_sequence(actions_sequence[i,:actions_length[i]].tolist()))
actual = target_commands[i]
assert pred == actual
```
#### try some interventions
```
data_json = json.load(open(path_to_data, "r"))
training_set = ReaSCANDataset(
data_json,
"../../../data-files/gSCAN-Simple/", split="train",
input_vocabulary_file="input_vocabulary.txt",
target_vocabulary_file="target_vocabulary.txt",
generate_vocabulary=False, k=0
)
training_set.read_dataset(
max_examples=100,
simple_situation_representation=False
)
train_data, _ = training_set.get_dual_dataset()
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=50)
hi_model = HighLevelModel(
# None
)
# Shuffle the dataset and loop over it.
for step, batch in enumerate(train_dataloader):
# main batch
input_batch, target_batch, situation_batch, \
agent_positions_batch, target_positions_batch, \
input_lengths_batch, target_lengths_batch, \
dual_input_batch, dual_target_batch, dual_situation_batch, \
dual_agent_positions_batch, dual_target_positions_batch, \
dual_input_lengths_batch, dual_target_lengths_batch = batch
high_hidden_states = hi_model(
agent_positions_batch=agent_positions_batch.unsqueeze(dim=-1),
target_positions_batch=target_positions_batch.unsqueeze(dim=-1),
tag="situation_encode"
)
high_actions = torch.zeros(
high_hidden_states.size(0), 1
).long()
dual_high_hidden_states = hi_model(
agent_positions_batch=dual_agent_positions_batch.unsqueeze(dim=-1),
target_positions_batch=dual_target_positions_batch.unsqueeze(dim=-1),
tag="situation_encode"
)
dual_high_actions = torch.zeros(
dual_high_hidden_states.size(0), 1
).long()
break # just steal one batch
intervene_time = 1
intervene_attribute = 0
# get the intercepted dual hidden states.
for j in range(intervene_time):
dual_high_hidden_states, dual_high_actions = hi_model(
hmm_states=dual_high_hidden_states,
hmm_actions=dual_high_actions,
tag="_hmm_step_fxn"
)
train_max_decoding_steps = 20
# main intervene for loop.
cf_high_hidden_states = high_hidden_states
cf_high_actions = high_actions
intervened_target_batch = [torch.ones(high_hidden_states.size(0), 1).long()] # SOS tokens
intervened_target_lengths_batch = torch.zeros(high_hidden_states.size(0), 1).long()
# we need to take of the SOS and EOS tokens.
for j in range(train_max_decoding_steps-2):
# intercept like antra!
if j == intervene_time:
# only swap out this part.
cf_high_hidden_states[:,intervene_attribute] = dual_high_hidden_states[:,intervene_attribute]
# comment out two lines below if it is not for testing.
# cf_high_hidden_states = dual_high_hidden_states
# cf_high_actions = dual_high_actions
cf_high_hidden_states, cf_high_actions = hi_model(
hmm_states=cf_high_hidden_states,
hmm_actions=cf_high_actions,
tag="_hmm_step_fxn"
)
# record the output for loss calculation.
intervened_target_batch += [cf_high_actions]
intervened_target_lengths_batch += (cf_high_actions!=0).long()
intervened_target_batch += [torch.zeros(high_hidden_states.size(0), 1).long()] # pad for extra eos
intervened_target_lengths_batch += 2
intervened_target_batch = torch.cat(intervened_target_batch, dim=-1)
for i in range(high_hidden_states.size(0)):
intervened_target_batch[i,intervened_target_lengths_batch[i,0]-1] = 2
(intervened_target_batch[:,:target_batch.size(1)] != target_batch).sum(1)
intervened_target_batch[:,:target_batch.size(1)]
```
#### Hidden states of high level model of the compositional generalization split.
```
# train hidden states
NUM = 200
agent_positions_batch = []
target_positions_batch = []
target_commands = []
for ex in data_json["examples"]["train"]:
target_commands += [ex["target_commands"]]
situation_repr = ex['situation']
agent = torch.tensor(
(int(situation_repr["agent_position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["agent_position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
target = torch.tensor(
(int(situation_repr["target_object"]["position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["target_object"]["position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
agent_positions_batch.append(agent)
target_positions_batch.append(target)
if len(agent_positions_batch) == NUM:
break
agent_positions_batch = torch.stack(agent_positions_batch, dim=0)
target_positions_batch = torch.stack(target_positions_batch, dim=0)
hi_model = HighLevelModel()
hidden_states = hi_model(agent_positions_batch, target_positions_batch, tag="situation_encode")
hidden_states
NUM = 200
cg_agent_positions_batch = []
cg_target_positions_batch = []
cg_target_commands = []
for ex in data_json["examples"]["situational_1"]:
cg_target_commands += [ex["target_commands"]]
situation_repr = ex['situation']
agent = torch.tensor(
(int(situation_repr["agent_position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["agent_position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
target = torch.tensor(
(int(situation_repr["target_object"]["position"]["row"]) * int(situation_repr["grid_size"])) +
int(situation_repr["target_object"]["position"]["column"]), dtype=torch.long).unsqueeze(dim=0)
cg_agent_positions_batch.append(agent)
cg_target_positions_batch.append(target)
if len(cg_agent_positions_batch) == NUM:
break
cg_agent_positions_batch = torch.stack(cg_agent_positions_batch, dim=0)
cg_target_positions_batch = torch.stack(cg_target_positions_batch, dim=0)
hi_model = HighLevelModel()
hidden_states = hi_model(cg_agent_positions_batch, cg_target_positions_batch, tag="situation_encode")
hidden_states[0]
data_json["examples"]["situational_1"][10]
# the first should be positive and the second should be negative.
```
Let us see, if your intervention produce any similar examples as above.
```
data_json = json.load(open(path_to_data, "r"))
training_set = ReaSCANDataset(
data_json,
"../../../data-files/gSCAN-Simple/", split="train",
input_vocabulary_file="input_vocabulary.txt",
target_vocabulary_file="target_vocabulary.txt",
generate_vocabulary=False, k=0
)
training_set.read_dataset(
max_examples=1000,
simple_situation_representation=False
)
train_data, _ = training_set.get_dual_dataset()
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=50)
# Shuffle the dataset and loop over it.
for step, batch in enumerate(train_dataloader):
# main batch
input_batch, target_batch, situation_batch, \
agent_positions_batch, target_positions_batch, \
input_lengths_batch, target_lengths_batch, \
dual_input_batch, dual_target_batch, dual_situation_batch, \
dual_agent_positions_batch, dual_target_positions_batch, \
dual_input_lengths_batch, dual_target_lengths_batch = batch
high_hidden_states = hi_model(
agent_positions_batch=agent_positions_batch.unsqueeze(dim=-1),
target_positions_batch=target_positions_batch.unsqueeze(dim=-1),
tag="situation_encode"
)
high_actions = torch.zeros(
high_hidden_states.size(0), 1
).long()
print(high_hidden_states)
break
# Shuffle the dataset and loop over it.
for step, batch in enumerate(train_dataloader):
# main batch
input_batch, target_batch, situation_batch, \
agent_positions_batch, target_positions_batch, \
input_lengths_batch, target_lengths_batch, \
dual_input_batch, dual_target_batch, dual_situation_batch, \
dual_agent_positions_batch, dual_target_positions_batch, \
dual_input_lengths_batch, dual_target_lengths_batch = batch
high_hidden_states = hi_model(
agent_positions_batch=agent_positions_batch.unsqueeze(dim=-1),
target_positions_batch=target_positions_batch.unsqueeze(dim=-1),
tag="situation_encode"
)
high_actions = torch.zeros(
high_hidden_states.size(0), 1
).long()
dual_high_hidden_states = hi_model(
agent_positions_batch=dual_agent_positions_batch.unsqueeze(dim=-1),
target_positions_batch=dual_target_positions_batch.unsqueeze(dim=-1),
tag="situation_encode"
)
dual_high_actions = torch.zeros(
dual_high_hidden_states.size(0), 1
).long()
intervene_attribute = 1
intervene_time = random.choice([1,2,3])
# get the intercepted dual hidden states.
for j in range(intervene_time):
dual_high_hidden_states, dual_high_actions = hi_model(
hmm_states=dual_high_hidden_states,
hmm_actions=dual_high_actions,
tag="_hmm_step_fxn"
)
train_max_decoding_steps = 20
# main intervene for loop.
cf_high_hidden_states = high_hidden_states
cf_high_actions = high_actions
# we need to take of the SOS and EOS tokens.
for j in range(train_max_decoding_steps-1):
# intercept like antra!
if j == intervene_time:
# only swap out this part.
cf_high_hidden_states[:,intervene_attribute] = dual_high_hidden_states[:,intervene_attribute]
print(cf_high_hidden_states)
break
cf_high_hidden_states, cf_high_actions = hi_model(
hmm_states=cf_high_hidden_states,
hmm_actions=cf_high_actions,
tag="_hmm_step_fxn"
)
cg_count = 0
for i in range(input_batch.size(0)):
if cf_high_hidden_states[i][0] > 0 and cf_high_hidden_states[i][1] < 0 and cf_high_hidden_states[i][2] == 0:
cg_count += 1
print(f"cg_count: {cg_count}/{input_batch.size(0)}")
```
Following sections are for counterfactual training for new attribute splits
```
path_to_data = "../../../data-files/ReaSCAN-novel-attribute/data-compositional-splits.txt"
data_json = json.load(open(path_to_data, "r"))
training_set = ReaSCANDataset(
data_json,
"../../../data-files/ReaSCAN-novel-attribute/", split="train",
input_vocabulary_file="input_vocabulary.txt",
target_vocabulary_file="target_vocabulary.txt",
generate_vocabulary=False, k=0
)
training_set.read_dataset(
max_examples=100,
simple_situation_representation=False
)
train_data, _ = training_set.get_dual_dataset(novel_attribute=True)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=50)
for step, batch in enumerate(train_dataloader):
input_sequence, target_sequence, situation, \
agent_positions, target_positions, \
input_lengths, target_lengths, \
dual_input_sequence, dual_target_sequence, dual_situation, \
dual_agent_positions, dual_target_positions, \
dual_input_lengths, dual_target_lengths, = batch
```
| github_jupyter |
# Segmentierung mit Stardist
Dieses Notebook ist dem 2D-Beispiel https://github.com/mpicbg-csbd/stardist/tree/master/examples/2D der GitHub Implementierung entnommen.
Nun wenden wir uns der Segmentierung mit DeepLearning zu. Hier verwenden wir wieder die *E. Coli* Daten.
```
from __future__ import print_function, unicode_literals, absolute_import, division
%load_ext autoreload
%autoreload 2
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
%config InlineBackend.figure_format = 'retina'
from glob import glob
from tqdm import tqdm
from tifffile import imread
from csbdeep.utils import Path, download_and_extract_zip_file
from stardist import fill_label_holes, relabel_image_stardist, random_label_cmap
from stardist.matching import matching_dataset
#np.random.seed(42)
lbl_cmap = random_label_cmap()
```
# Data
This notebook demonstrates how the training data for *StarDist* should look like and whether the annotated objects can be appropriately described by star-convex polygons.
<div class="alert alert-block alert-info">
The training data that needs to be provided for StarDist consists of corresponding pairs of raw images and pixelwise annotated ground truth images (masks), where every pixel has a unique integer value indicating the object id (or 0 for background).
</div>
```
X_glob = sorted(glob('/extdata/readonly/f-prak-v15/e-coli-swarming/train/input/*.tif'))
Y_glob = sorted(glob('/extdata/readonly/f-prak-v15/e-coli-swarming/train/labels/*.tif'))
def labelname(name):
return name[:-5]+name[-4:]
assert all(Path(x).name == labelname(Path(y).name) for x,y in zip(X_glob, Y_glob))
```
Load only a small subset
```
X_glob, Y_glob = X_glob[:10], Y_glob[:10]
X = list(map(imread, X_glob))
Y = list(map(imread, Y_glob))
```
# Example image
```
i = min(4, len(X)-1)
img, lbl = X[i], fill_label_holes(Y[i])
assert img.ndim in (2,3)
img = img if img.ndim==2 else img[...,:3]
# assumed axes ordering of img and lbl is: YX(C)
plt.figure(figsize=(16,10))
plt.subplot(121); plt.imshow(img,cmap='gray'); plt.axis('off'); plt.title('Raw image')
plt.subplot(122); plt.imshow(lbl,cmap=lbl_cmap); plt.axis('off'); plt.title('GT labels')
None;
```
# Fitting ground-truth labels with star-convex polygons
In unseren Ground Truth labels sind Objekte einfach zusammenhängende Pixel. Stardist betrachtet Objekte jedoch als Polygone. Macht euch diesen Unterschied nochmals klar! Im Folgenden ist `n_rays` die Anzahl der Strahlen, also auch die Anzahl der Polygonpunkte, mit der ein Objekt dargstellt wird. Es wird überprüft, wie viele Polygonpunkte benötigt werden, um die Objekte auch mit der Polygon Darstellung gut zu beschreiben. Dafür wird die Interscetion over Union (IoU) von label-Objekt mit dem Polygon-Objekt berechnet für verschiedene Anzahlen an Strahlen.
```
n_rays = [2**i for i in range(2,8)]
scores = []
for r in tqdm(n_rays):
Y_reconstructed = [relabel_image_stardist(lbl, n_rays=r) for lbl in Y]
mean_iou = matching_dataset(Y, Y_reconstructed, thresh=0, show_progress=False).mean_true_score
scores.append(mean_iou)
plt.figure(figsize=(8,5))
plt.plot(n_rays, scores, 'o-')
plt.xlabel('Number of rays for star-convex polygon')
plt.ylabel('Reconstruction score (mean intersection over union)')
plt.title("Accuracy of ground truth reconstruction (should be > 0.8 for a reasonable number of rays)")
None;
```
## Example image reconstructed with various number of rays
```
fig, ax = plt.subplots(2,3, figsize=(16,11))
for a,r in zip(ax.flat,n_rays):
a.imshow(relabel_image_stardist(lbl, n_rays=r), cmap=lbl_cmap)
a.set_title('Reconstructed (%d rays)' % r)
a.axis('off')
plt.tight_layout();
```
| github_jupyter |
```
from rcwc import *
import numpy as np
import cvxpy as cp
from sklearn.neighbors import NearestNeighbors as kNN
from matplotlib import pyplot as plt
import os, time
from matplotlib import cm
import matplotlib
from matplotlib.ticker import ScalarFormatter
import seaborn as sns
np.random.seed(94)
n = 50
pts = np.random.rand(n,2)
# Building spatially correlated dataset
x = pts[:,0] + pts[:,1] - 1
colormap = matplotlib.colors.ListedColormap(sns.color_palette("coolwarm",8))
plt.scatter(pts[:,0], pts[:,1], c=x, cmap=colormap)
plt.clim(-1,1)
plt.colorbar()
plt.xlabel('x')
plt.ylabel('y', rotation='0', labelpad=20)
plt.yticks([0,0.5,1])
plt.tight_layout()
# Creating edges
knearest = kNN(n_neighbors=6).fit(pts)
dists, neighbors = knearest.kneighbors(pts)
neighbors = neighbors[:,1:] #get rid of self edges
# Generating samples
m = 1000 # num samples
k = 25 # size of sample
nrecruits = 2 # number of new recruits
A = np.zeros((m,n))
b = np.ones((m,n)) * 1/n
start_node = np.random.randint(low=0,high=n)
A[:,start_node] = 1
for i in range(m):
sample = set([start_node])
recruiters = set([start_node])
while len(recruiters) > 0 and len(sample) < k:
recruits = set()
for r in recruiters:
possible_recruits = [neigh for neigh in tuple(neighbors[r])]
idx = np.random.choice(len(possible_recruits), size=min(k-len(sample), min(nrecruits, len(possible_recruits))))
recruits.update([possible_recruits[id] for id in tuple(idx)])
sample.update(recruits)
if len(sample) >= k:
break
recruiters = recruits
for elem in sample:
A[i,elem] = 1
# Plot a random sample (gold is root, red is in sample, all points are in target)
i = np.random.randint(0,m)
plt.scatter(pts[:,0], pts[:,1])
plt.scatter(pts[A[i,:] == 1,0], pts[A[i,:] == 1,1], c='red')
plt.scatter(pts[start_node,0], pts[start_node,1],c='gold')
# Plot individual probability of being sampled
sample_prob = np.sum(A, axis=0) / m
mask = sample_prob > 0
plt.scatter(pts[:,0], pts[:,1], c='grey')
plt.scatter(pts[mask,0], pts[mask,1], c=sample_prob[mask])
plt.colorbar()
# Basic estimator (sample mean)
a_mean = A / np.sum(A, axis=1, keepdims=True)
print('Worst-case error: {}'.format(evaluate_weights_grothendieck(a_mean, b)))
print('Spatial values error: {}'.format(evaluate_weights(a_mean, b, x)))
%%time
# RCWC estimator
a_rcwc = rcwc(A, b)
# Evaluating RCWC results
print('Worst-case error: {}'.format(evaluate_weights_grothendieck(a_rcwc, b, is_verbose=False)))
print('Spatial values error: {}'.format(evaluate_weights(a_rcwc, b, x)))
```
| github_jupyter |
# This tutorial shows how to make and manipulate a power spectrum of two light curves using Stingray.
```
import numpy as np
from stingray import Lightcurve, Powerspectrum, AveragedPowerspectrum
import matplotlib.pyplot as plt
import matplotlib.font_manager as font_manager
%matplotlib inline
font_prop = font_manager.FontProperties(size=16)
```
# Power spectrum example
## 1. Create a light curve
There are two ways to make `Lightcurve` objects. We'll show one way here. Check out "Lightcurve/Lightcurve\ tutorial.ipynb" for more examples.
Generate an array of relative timestamps that's 8 seconds long, with dt = 0.03125 s, and make two signals in units of counts. The signal is a sine wave with amplitude = 300 cts/s, frequency = 2 Hz, phase offset = 0 radians, and mean = 1000 cts/s. We then add Poisson noise to the light curve.
```
dt = 0.03125 # seconds
exposure = 8. # seconds
times = np.arange(0, exposure, dt) # seconds
signal = 300 * np.sin(2.*np.pi*times/0.5) + 1000 # counts/s
noisy = np.random.poisson(signal*dt) # counts
```
Now let's turn `noisy` into a `Lightcurve` object.
```
lc = Lightcurve(times, noisy)
```
Here we plot it to see what it looks like.
```
fig, ax = plt.subplots(1,1,figsize=(10,6))
ax.plot(lc.time, lc.counts, lw=2, color='blue')
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Counts (cts)", fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16)
ax.tick_params(axis='y', labelsize=16)
ax.tick_params(which='major', width=1.5, length=7)
ax.tick_params(which='minor', width=1.5, length=4)
plt.show()
```
## 2. Pass the light curve to the `Powerspectrum` class to create a `Powerspectrum` object.
You can also specify the optional attribute `norm` if you wish to normalize power to squared fractional rms, Leahy, or squared absolute normalization. The default normalization is 'none'.
```
ps = Powerspectrum(lc)
print(ps)
```
Since the negative Fourier frequencies (and their associated powers) are discarded, the number of time bins per segment `n` is twice the length of `freq` and `power`.
```
print("\nSize of positive Fourier frequencies:", len(ps.freq))
print("Number of data points per segment:", ps.n)
```
# Properties
A `Powerspectrum` object has the following properties :
1. `freq` : Numpy array of mid-bin frequencies that the Fourier transform samples.
2. `power` : Numpy array of the power spectrum.
3. `df` : The frequency resolution.
4. `m` : The number of power spectra averaged together. For a `Powerspectrum` of a single segment, `m=1`.
5. `n` : The number of data points (time bins) in one segment of the light curve.
6. `nphots1` : The total number of photons in the light curve.
```
print(ps.freq)
print(ps.power)
print(ps.df)
print(ps.m)
print(ps.n)
print(ps.nphots1)
```
We can plot the power as a function of Fourier frequency. Notice how there's a spike at our signal frequency of 2 Hz!
```
fig, ax1 = plt.subplots(1,1,figsize=(9,6), sharex=True)
ax1.plot(ps.freq, ps.power, lw=2, color='blue')
ax1.set_ylabel("Frequency (Hz)", fontproperties=font_prop)
ax1.set_ylabel("Power (raw)", fontproperties=font_prop)
ax1.set_yscale('log')
ax1.tick_params(axis='x', labelsize=16)
ax1.tick_params(axis='y', labelsize=16)
ax1.tick_params(which='major', width=1.5, length=7)
ax1.tick_params(which='minor', width=1.5, length=4)
for axis in ['top', 'bottom', 'left', 'right']:
ax1.spines[axis].set_linewidth(1.5)
plt.show()
```
You'll notice that the power spectrum is a bit noisy. This is because we're only using one segment of data. Let's try averaging together power spectra from multiple segments of data.
# Averaged power spectrum example
You could use a long `Lightcurve` and have `AveragedPowerspectrum` chop it into specified segments, or give a list of `Lightcurve`s where each segment of `Lightcurve` is the same length. We'll show the first way here.
## 1. Create a long light curve.
Generate an array of relative timestamps that's 1600 seconds long, and a signal in count units, with the same properties as the previous example. We then add Poisson noise and turn it into a `Lightcurve` object.
```
long_dt = 0.03125 # seconds
long_exposure = 1600. # seconds
long_times = np.arange(0, long_exposure, long_dt) # seconds
# In count rate units here
long_signal = 300 * np.sin(2.*np.pi*long_times/0.5) + 1000
# Multiply by dt to get count units, then add Poisson noise
long_noisy = np.random.poisson(long_signal*dt)
long_lc = Lightcurve(long_times, long_noisy)
fig, ax = plt.subplots(1,1,figsize=(10,6))
ax.plot(long_lc.time, long_lc.counts, lw=2, color='blue')
ax.set_xlim(0,20)
ax.set_xlabel("Time (s)", fontproperties=font_prop)
ax.set_ylabel("Counts (cts)", fontproperties=font_prop)
ax.tick_params(axis='x', labelsize=16)
ax.tick_params(axis='y', labelsize=16)
ax.tick_params(which='major', width=1.5, length=7)
ax.tick_params(which='minor', width=1.5, length=4)
plt.show()
```
## 2. Pass the light curve to the `AveragedPowerspectrum` class with a specified `segment_size`.
If the exposure (length) of the light curve cannot be divided by `segment_size` with a remainder of zero, the last incomplete segment is thrown out, to avoid signal artefacts. Here we're using 8 second segments.
```
avg_ps = AveragedPowerspectrum(long_lc, 8.)
```
We can check how many segments were averaged together by printing the `m` attribute.
```
print("Number of segments: %d" % avg_ps.m)
```
`AveragedPowerspectrum` has the same properties as `Powerspectrum`, but with `m` $>$1.
Let's plot the averaged power spectrum!
```
fig, ax1 = plt.subplots(1,1,figsize=(9,6))
ax1.plot(avg_ps.freq, avg_ps.power, lw=2, color='blue')
ax1.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax1.set_ylabel("Power (raw)", fontproperties=font_prop)
ax1.set_yscale('log')
ax1.tick_params(axis='x', labelsize=16)
ax1.tick_params(axis='y', labelsize=16)
ax1.tick_params(which='major', width=1.5, length=7)
ax1.tick_params(which='minor', width=1.5, length=4)
for axis in ['top', 'bottom', 'left', 'right']:
ax1.spines[axis].set_linewidth(1.5)
plt.show()
```
Now we'll show examples of all the things you can do with a `Powerspectrum` or `AveragedPowerspectrum` object using built-in stingray methods.
# Normalizating the power spectrum
The three kinds of normalization are:
* `leahy`: Leahy normalization. Makes the Poisson noise level $= 2$. See *Leahy et al. 1983, ApJ, 266, 160L*.
* `frac`: Fractional rms-squared normalization, also known as rms normalization. Makes the Poisson noise level $= 2 / meanrate$. See *Belloni & Hasinger 1990, A&A, 227, L33*, and *Miyamoto et al. 1992, ApJ, 391, L21.*
* `abs`: Absolute rms-squared normalization, also known as absolute normalization. Makes the Poisson noise level $= 2 \times meanrate$. See *insert citation*.
* `none`: No normalization applied. This is the default.
```
avg_ps_leahy = AveragedPowerspectrum(long_lc, 8, norm='leahy')
avg_ps_frac = AveragedPowerspectrum(long_lc, 8., norm='frac')
avg_ps_abs = AveragedPowerspectrum(long_lc, 8., norm='abs')
fig, [ax1, ax2, ax3] = plt.subplots(3,1,figsize=(6,12))
ax1.plot(avg_ps_leahy.freq, avg_ps_leahy.power, lw=2, color='black')
ax1.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax1.set_ylabel("Power (Leahy)", fontproperties=font_prop)
ax1.set_yscale('log')
ax1.tick_params(axis='x', labelsize=14)
ax1.tick_params(axis='y', labelsize=14)
ax1.tick_params(which='major', width=1.5, length=7)
ax1.tick_params(which='minor', width=1.5, length=4)
ax1.set_title("Leahy norm.", fontproperties=font_prop)
ax2.plot(avg_ps_frac.freq, avg_ps_frac.power, lw=2, color='black')
ax2.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax2.set_ylabel("Power (rms)", fontproperties=font_prop)
ax2.tick_params(axis='x', labelsize=14)
ax2.tick_params(axis='y', labelsize=14)
ax2.set_yscale('log')
ax2.tick_params(which='major', width=1.5, length=7)
ax2.tick_params(which='minor', width=1.5, length=4)
ax2.set_title("Fractional rms-squared norm.", fontproperties=font_prop)
ax3.plot(avg_ps_abs.freq, avg_ps_abs.power, lw=2, color='black')
ax3.set_xlabel("Frequency (Hz)", fontproperties=font_prop)
ax3.set_ylabel("Power (abs)", fontproperties=font_prop)
ax3.tick_params(axis='x', labelsize=14)
ax3.tick_params(axis='y', labelsize=14)
ax3.set_yscale('log')
ax3.tick_params(which='major', width=1.5, length=7)
ax3.tick_params(which='minor', width=1.5, length=4)
ax3.set_title("Absolute rms-squared norm.", fontproperties=font_prop)
for axis in ['top', 'bottom', 'left', 'right']:
ax1.spines[axis].set_linewidth(1.5)
ax2.spines[axis].set_linewidth(1.5)
ax3.spines[axis].set_linewidth(1.5)
plt.tight_layout()
plt.show()
```
# Re-binning a power spectrum in frequency
Typically, rebinning is done on an averaged, normalized power spectrum.
## 1. We can linearly re-bin a power spectrum
(although this is not done much in practice)
```
print("DF before:", avg_ps.df)
# Both of the following ways are allowed syntax:
# lin_rb_ps = Powerspectrum.rebin(avg_ps, 0.25, method='mean')
lin_rb_ps = avg_ps.rebin(0.25, method='mean')
print("DF after:", lin_rb_ps.df)
```
## 2. And we can logarithmically/geometrically re-bin a power spectrum
In this re-binning, each bin size is 1+f times larger than the previous bin size, where `f` is user-specified and normally in the range 0.01-0.1. The default value is `f=0.01`.
```
# Both of the following ways are allowed syntax:
# log_rb_ps, log_rb_freq, binning = Powerspectrum.rebin_log(avg_ps, f=0.02)
log_rb_ps, log_rb_freq, binning = avg_ps.rebin_log(f=0.02)
```
Note that `rebin` returns a `Powerspectrum` or `AveragedPowerspectrum` object (depending on the input object), whereas `rebin_log` returns three `np.ndarray`s.
```
# print(type(lin_rb_ps))
print(type(log_rb_ps))
print(type(log_rb_freq))
print(type(binning))
```
| github_jupyter |
##### Copyright 2020 The TensorFlow Hub Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://www.tensorflow.org/text/tutorials/classify_text_with_bert"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/text/blob/master/docs/tutorials/classify_text_with_bert.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/text/blob/master/docs/tutorials/classify_text_with_bert.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/text/docs/tutorials/classify_text_with_bert.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
</td>
<td>
<a href="https://tfhub.dev/google/collections/bert/1"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
</td>
</table>
# Classify text with BERT
This tutorial contains complete code to fine-tune BERT to perform sentiment analysis on a dataset of plain-text IMDB movie reviews.
In addition to training a model, you will learn how to preprocess text into an appropriate format.
In this notebook, you will:
- Load the IMDB dataset
- Load a BERT model from TensorFlow Hub
- Build your own model by combining BERT with a classifier
- Train your own model, fine-tuning BERT as part of that
- Save your model and use it to classify sentences
If you're new to working with the IMDB dataset, please see [Basic text classification](https://www.tensorflow.org/tutorials/keras/text_classification) for more details.
## About BERT
[BERT](https://arxiv.org/abs/1810.04805) and other Transformer encoder architectures have been wildly successful on a variety of tasks in NLP (natural language processing). They compute vector-space representations of natural language that are suitable for use in deep learning models. The BERT family of models uses the Transformer encoder architecture to process each token of input text in the full context of all tokens before and after, hence the name: Bidirectional Encoder Representations from Transformers.
BERT models are usually pre-trained on a large corpus of text, then fine-tuned for specific tasks.
## Setup
```
# A dependency of the preprocessing for BERT inputs
!pip install -q tensorflow-text
```
You will use the AdamW optimizer from [tensorflow/models](https://github.com/tensorflow/models).
```
!pip install -q tf-models-official
import os
import shutil
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_text as text
from official.nlp import optimization # to create AdamW optmizer
import matplotlib.pyplot as plt
tf.get_logger().setLevel('ERROR')
```
## Sentiment Analysis
This notebook trains a sentiment analysis model to classify movie reviews as *positive* or *negative*, based on the text of the review.
You'll use the [Large Movie Review Dataset](https://ai.stanford.edu/~amaas/data/sentiment/) that contains the text of 50,000 movie reviews from the [Internet Movie Database](https://www.imdb.com/).
### Download the IMDB dataset
Let's download and extract the dataset, then explore the directory structure.
```
url = 'https://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz'
dataset = tf.keras.utils.get_file('aclImdb_v1.tar.gz', url,
untar=True, cache_dir='.',
cache_subdir='')
dataset_dir = os.path.join(os.path.dirname(dataset), 'aclImdb')
train_dir = os.path.join(dataset_dir, 'train')
# remove unused folders to make it easier to load the data
remove_dir = os.path.join(train_dir, 'unsup')
shutil.rmtree(remove_dir)
```
Next, you will use the `text_dataset_from_directory` utility to create a labeled `tf.data.Dataset`.
The IMDB dataset has already been divided into train and test, but it lacks a validation set. Let's create a validation set using an 80:20 split of the training data by using the `validation_split` argument below.
Note: When using the `validation_split` and `subset` arguments, make sure to either specify a random seed, or to pass `shuffle=False`, so that the validation and training splits have no overlap.
```
AUTOTUNE = tf.data.AUTOTUNE
batch_size = 32
seed = 42
raw_train_ds = tf.keras.preprocessing.text_dataset_from_directory(
'aclImdb/train',
batch_size=batch_size,
validation_split=0.2,
subset='training',
seed=seed)
class_names = raw_train_ds.class_names
train_ds = raw_train_ds.cache().prefetch(buffer_size=AUTOTUNE)
val_ds = tf.keras.preprocessing.text_dataset_from_directory(
'aclImdb/train',
batch_size=batch_size,
validation_split=0.2,
subset='validation',
seed=seed)
val_ds = val_ds.cache().prefetch(buffer_size=AUTOTUNE)
test_ds = tf.keras.preprocessing.text_dataset_from_directory(
'aclImdb/test',
batch_size=batch_size)
test_ds = test_ds.cache().prefetch(buffer_size=AUTOTUNE)
```
Let's take a look at a few reviews.
```
for text_batch, label_batch in train_ds.take(1):
for i in range(3):
print(f'Review: {text_batch.numpy()[i]}')
label = label_batch.numpy()[i]
print(f'Label : {label} ({class_names[label]})')
```
## Loading models from TensorFlow Hub
Here you can choose which BERT model you will load from TensorFlow Hub and fine-tune. There are multiple BERT models available.
- [BERT-Base](https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3), [Uncased](https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3) and [seven more models](https://tfhub.dev/google/collections/bert/1) with trained weights released by the original BERT authors.
- [Small BERTs](https://tfhub.dev/google/collections/bert/1) have the same general architecture but fewer and/or smaller Transformer blocks, which lets you explore tradeoffs between speed, size and quality.
- [ALBERT](https://tfhub.dev/google/collections/albert/1): four different sizes of "A Lite BERT" that reduces model size (but not computation time) by sharing parameters between layers.
- [BERT Experts](https://tfhub.dev/google/collections/experts/bert/1): eight models that all have the BERT-base architecture but offer a choice between different pre-training domains, to align more closely with the target task.
- [Electra](https://tfhub.dev/google/collections/electra/1) has the same architecture as BERT (in three different sizes), but gets pre-trained as a discriminator in a set-up that resembles a Generative Adversarial Network (GAN).
- BERT with Talking-Heads Attention and Gated GELU [[base](https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_base/1), [large](https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_large/1)] has two improvements to the core of the Transformer architecture.
The model documentation on TensorFlow Hub has more details and references to the
research literature. Follow the links above, or click on the [`tfhub.dev`](http://tfhub.dev) URL
printed after the next cell execution.
The suggestion is to start with a Small BERT (with fewer parameters) since they are faster to fine-tune. If you like a small model but with higher accuracy, ALBERT might be your next option. If you want even better accuracy, choose
one of the classic BERT sizes or their recent refinements like Electra, Talking Heads, or a BERT Expert.
Aside from the models available below, there are [multiple versions](https://tfhub.dev/google/collections/transformer_encoders_text/1) of the models that are larger and can yield even better accuracy, but they are too big to be fine-tuned on a single GPU. You will be able to do that on the [Solve GLUE tasks using BERT on a TPU tutorial](https://www.tensorflow.org/text/tutorials/bert_glue).
You'll see in the code below that switching the tfhub.dev URL is enough to try any of these models, because all the differences between them are encapsulated in the SavedModels from TF Hub.
```
#@title Choose a BERT model to fine-tune
bert_model_name = 'small_bert/bert_en_uncased_L-4_H-512_A-8' #@param ["bert_en_uncased_L-12_H-768_A-12", "bert_en_cased_L-12_H-768_A-12", "bert_multi_cased_L-12_H-768_A-12", "small_bert/bert_en_uncased_L-2_H-128_A-2", "small_bert/bert_en_uncased_L-2_H-256_A-4", "small_bert/bert_en_uncased_L-2_H-512_A-8", "small_bert/bert_en_uncased_L-2_H-768_A-12", "small_bert/bert_en_uncased_L-4_H-128_A-2", "small_bert/bert_en_uncased_L-4_H-256_A-4", "small_bert/bert_en_uncased_L-4_H-512_A-8", "small_bert/bert_en_uncased_L-4_H-768_A-12", "small_bert/bert_en_uncased_L-6_H-128_A-2", "small_bert/bert_en_uncased_L-6_H-256_A-4", "small_bert/bert_en_uncased_L-6_H-512_A-8", "small_bert/bert_en_uncased_L-6_H-768_A-12", "small_bert/bert_en_uncased_L-8_H-128_A-2", "small_bert/bert_en_uncased_L-8_H-256_A-4", "small_bert/bert_en_uncased_L-8_H-512_A-8", "small_bert/bert_en_uncased_L-8_H-768_A-12", "small_bert/bert_en_uncased_L-10_H-128_A-2", "small_bert/bert_en_uncased_L-10_H-256_A-4", "small_bert/bert_en_uncased_L-10_H-512_A-8", "small_bert/bert_en_uncased_L-10_H-768_A-12", "small_bert/bert_en_uncased_L-12_H-128_A-2", "small_bert/bert_en_uncased_L-12_H-256_A-4", "small_bert/bert_en_uncased_L-12_H-512_A-8", "small_bert/bert_en_uncased_L-12_H-768_A-12", "albert_en_base", "electra_small", "electra_base", "experts_pubmed", "experts_wiki_books", "talking-heads_base"]
map_name_to_handle = {
'bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3',
'bert_en_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_cased_L-12_H-768_A-12/3',
'bert_multi_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_multi_cased_L-12_H-768_A-12/3',
'small_bert/bert_en_uncased_L-2_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-128_A-2/1',
'small_bert/bert_en_uncased_L-2_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-256_A-4/1',
'small_bert/bert_en_uncased_L-2_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-512_A-8/1',
'small_bert/bert_en_uncased_L-2_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-768_A-12/1',
'small_bert/bert_en_uncased_L-4_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-128_A-2/1',
'small_bert/bert_en_uncased_L-4_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-256_A-4/1',
'small_bert/bert_en_uncased_L-4_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-512_A-8/1',
'small_bert/bert_en_uncased_L-4_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-768_A-12/1',
'small_bert/bert_en_uncased_L-6_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-128_A-2/1',
'small_bert/bert_en_uncased_L-6_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-256_A-4/1',
'small_bert/bert_en_uncased_L-6_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-512_A-8/1',
'small_bert/bert_en_uncased_L-6_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-768_A-12/1',
'small_bert/bert_en_uncased_L-8_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-128_A-2/1',
'small_bert/bert_en_uncased_L-8_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-256_A-4/1',
'small_bert/bert_en_uncased_L-8_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-512_A-8/1',
'small_bert/bert_en_uncased_L-8_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-768_A-12/1',
'small_bert/bert_en_uncased_L-10_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-128_A-2/1',
'small_bert/bert_en_uncased_L-10_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-256_A-4/1',
'small_bert/bert_en_uncased_L-10_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-512_A-8/1',
'small_bert/bert_en_uncased_L-10_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-768_A-12/1',
'small_bert/bert_en_uncased_L-12_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-128_A-2/1',
'small_bert/bert_en_uncased_L-12_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-256_A-4/1',
'small_bert/bert_en_uncased_L-12_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-512_A-8/1',
'small_bert/bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-768_A-12/1',
'albert_en_base':
'https://tfhub.dev/tensorflow/albert_en_base/2',
'electra_small':
'https://tfhub.dev/google/electra_small/2',
'electra_base':
'https://tfhub.dev/google/electra_base/2',
'experts_pubmed':
'https://tfhub.dev/google/experts/bert/pubmed/2',
'experts_wiki_books':
'https://tfhub.dev/google/experts/bert/wiki_books/2',
'talking-heads_base':
'https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_base/1',
}
map_model_to_preprocess = {
'bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'bert_en_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_cased_preprocess/3',
'small_bert/bert_en_uncased_L-2_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-2_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-2_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-2_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-4_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-4_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-4_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-4_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-6_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-6_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-6_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-6_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-8_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-8_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-8_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-8_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-10_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-10_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-10_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-10_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-12_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-12_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-12_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'small_bert/bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'bert_multi_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_multi_cased_preprocess/3',
'albert_en_base':
'https://tfhub.dev/tensorflow/albert_en_preprocess/3',
'electra_small':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'electra_base':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'experts_pubmed':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'experts_wiki_books':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
'talking-heads_base':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/3',
}
tfhub_handle_encoder = map_name_to_handle[bert_model_name]
tfhub_handle_preprocess = map_model_to_preprocess[bert_model_name]
print(f'BERT model selected : {tfhub_handle_encoder}')
print(f'Preprocess model auto-selected: {tfhub_handle_preprocess}')
```
## The preprocessing model
Text inputs need to be transformed to numeric token ids and arranged in several Tensors before being input to BERT. TensorFlow Hub provides a matching preprocessing model for each of the BERT models discussed above, which implements this transformation using TF ops from the TF.text library. It is not necessary to run pure Python code outside your TensorFlow model to preprocess text.
The preprocessing model must be the one referenced by the documentation of the BERT model, which you can read at the URL printed above. For BERT models from the drop-down above, the preprocessing model is selected automatically.
Note: You will load the preprocessing model into a [hub.KerasLayer](https://www.tensorflow.org/hub/api_docs/python/hub/KerasLayer) to compose your fine-tuned model. This is the preferred API to load a TF2-style SavedModel from TF Hub into a Keras model.
```
bert_preprocess_model = hub.KerasLayer(tfhub_handle_preprocess)
```
Let's try the preprocessing model on some text and see the output:
```
text_test = ['this is such an amazing movie!']
text_preprocessed = bert_preprocess_model(text_test)
print(f'Keys : {list(text_preprocessed.keys())}')
print(f'Shape : {text_preprocessed["input_word_ids"].shape}')
print(f'Word Ids : {text_preprocessed["input_word_ids"][0, :12]}')
print(f'Input Mask : {text_preprocessed["input_mask"][0, :12]}')
print(f'Type Ids : {text_preprocessed["input_type_ids"][0, :12]}')
```
As you can see, now you have the 3 outputs from the preprocessing that a BERT model would use (`input_words_id`, `input_mask` and `input_type_ids`).
Some other important points:
- The input is truncated to 128 tokens. The number of tokens can be customized, and you can see more details on the [Solve GLUE tasks using BERT on a TPU tutorial](https://www.tensorflow.org/text/tutorials/bert_glue).
- The `input_type_ids` only have one value (0) because this is a single sentence input. For a multiple sentence input, it would have one number for each input.
Since this text preprocessor is a TensorFlow model, It can be included in your model directly.
## Using the BERT model
Before putting BERT into your own model, let's take a look at its outputs. You will load it from TF Hub and see the returned values.
```
bert_model = hub.KerasLayer(tfhub_handle_encoder)
bert_results = bert_model(text_preprocessed)
print(f'Loaded BERT: {tfhub_handle_encoder}')
print(f'Pooled Outputs Shape:{bert_results["pooled_output"].shape}')
print(f'Pooled Outputs Values:{bert_results["pooled_output"][0, :12]}')
print(f'Sequence Outputs Shape:{bert_results["sequence_output"].shape}')
print(f'Sequence Outputs Values:{bert_results["sequence_output"][0, :12]}')
```
The BERT models return a map with 3 important keys: `pooled_output`, `sequence_output`, `encoder_outputs`:
- `pooled_output` to represent each input sequence as a whole. The shape is `[batch_size, H]`. You can think of this as an embedding for the entire movie review.
- `sequence_output` represents each input token in the context. The shape is `[batch_size, seq_length, H]`. You can think of this as a contextual embedding for every token in the movie review.
- `encoder_outputs` are the intermediate activations of the `L` Transformer blocks. `outputs["encoder_outputs"][i]` is a Tensor of shape `[batch_size, seq_length, 1024]` with the outputs of the i-th Transformer block, for `0 <= i < L`. The last value of the list is equal to `sequence_output`.
For the fine-tuning you are going to use the `pooled_output` array.
## Define your model
You will create a very simple fine-tuned model, with the preprocessing model, the selected BERT model, one Dense and a Dropout layer.
Note: for more information about the base model's input and output you can use just follow the model's url for documentation. Here specifically you don't need to worry about it because the preprocessing model will take care of that for you.
```
def build_classifier_model():
text_input = tf.keras.layers.Input(shape=(), dtype=tf.string, name='text')
preprocessing_layer = hub.KerasLayer(tfhub_handle_preprocess, name='preprocessing')
encoder_inputs = preprocessing_layer(text_input)
encoder = hub.KerasLayer(tfhub_handle_encoder, trainable=True, name='BERT_encoder')
outputs = encoder(encoder_inputs)
net = outputs['pooled_output']
net = tf.keras.layers.Dropout(0.1)(net)
net = tf.keras.layers.Dense(1, activation=None, name='classifier')(net)
return tf.keras.Model(text_input, net)
```
Let's check that the model runs with the output of the preprocessing model.
```
classifier_model = build_classifier_model()
bert_raw_result = classifier_model(tf.constant(text_test))
print(tf.sigmoid(bert_raw_result))
```
The output is meaningless, of course, because the model has not been trained yet.
Let's take a look at the model's structure.
```
tf.keras.utils.plot_model(classifier_model)
```
## Model training
You now have all the pieces to train a model, including the preprocessing module, BERT encoder, data, and classifier.
### Loss function
Since this is a binary classification problem and the model outputs a probability (a single-unit layer), you'll use `losses.BinaryCrossentropy` loss function.
```
loss = tf.keras.losses.BinaryCrossentropy(from_logits=True)
metrics = tf.metrics.BinaryAccuracy()
```
### Optimizer
For fine-tuning, let's use the same optimizer that BERT was originally trained with: the "Adaptive Moments" (Adam). This optimizer minimizes the prediction loss and does regularization by weight decay (not using moments), which is also known as [AdamW](https://arxiv.org/abs/1711.05101).
For the learning rate (`init_lr`), we use the same schedule as BERT pre-training: linear decay of a notional initial learning rate, prefixed with a linear warm-up phase over the first 10% of training steps (`num_warmup_steps`). In line with the BERT paper, the initial learning rate is smaller for fine-tuning (best of 5e-5, 3e-5, 2e-5).
```
epochs = 5
steps_per_epoch = tf.data.experimental.cardinality(train_ds).numpy()
num_train_steps = steps_per_epoch * epochs
num_warmup_steps = int(0.1*num_train_steps)
init_lr = 3e-5
optimizer = optimization.create_optimizer(init_lr=init_lr,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
optimizer_type='adamw')
```
### Loading the BERT model and training
Using the `classifier_model` you created earlier, you can compile the model with the loss, metric and optimizer.
```
classifier_model.compile(optimizer=optimizer,
loss=loss,
metrics=metrics)
```
Note: training time will vary depending on the complexity of the BERT model you have selected.
```
print(f'Training model with {tfhub_handle_encoder}')
history = classifier_model.fit(x=train_ds,
validation_data=val_ds,
epochs=epochs)
```
### Evaluate the model
Let's see how the model performs. Two values will be returned. Loss (a number which represents the error, lower values are better), and accuracy.
```
loss, accuracy = classifier_model.evaluate(test_ds)
print(f'Loss: {loss}')
print(f'Accuracy: {accuracy}')
```
### Plot the accuracy and loss over time
Based on the `History` object returned by `model.fit()`. You can plot the training and validation loss for comparison, as well as the training and validation accuracy:
```
history_dict = history.history
print(history_dict.keys())
acc = history_dict['binary_accuracy']
val_acc = history_dict['val_binary_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
fig = plt.figure(figsize=(10, 6))
fig.tight_layout()
plt.subplot(2, 1, 1)
# "bo" is for "blue dot"
plt.plot(epochs, loss, 'r', label='Training loss')
# b is for "solid blue line"
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
# plt.xlabel('Epochs')
plt.ylabel('Loss')
plt.legend()
plt.subplot(2, 1, 2)
plt.plot(epochs, acc, 'r', label='Training acc')
plt.plot(epochs, val_acc, 'b', label='Validation acc')
plt.title('Training and validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc='lower right')
```
In this plot, the red lines represents the training loss and accuracy, and the blue lines are the validation loss and accuracy.
## Export for inference
Now you just save your fine-tuned model for later use.
```
dataset_name = 'imdb'
saved_model_path = './{}_bert'.format(dataset_name.replace('/', '_'))
classifier_model.save(saved_model_path, include_optimizer=False)
```
Let's reload the model so you can try it side by side with the model that is still in memory.
```
reloaded_model = tf.saved_model.load(saved_model_path)
```
Here you can test your model on any sentence you want, just add to the examples variable below.
```
def print_my_examples(inputs, results):
result_for_printing = \
[f'input: {inputs[i]:<30} : score: {results[i][0]:.6f}'
for i in range(len(inputs))]
print(*result_for_printing, sep='\n')
print()
examples = [
'this is such an amazing movie!', # this is the same sentence tried earlier
'The movie was great!',
'The movie was meh.',
'The movie was okish.',
'The movie was terrible...'
]
reloaded_results = tf.sigmoid(reloaded_model(tf.constant(examples)))
original_results = tf.sigmoid(classifier_model(tf.constant(examples)))
print('Results from the saved model:')
print_my_examples(examples, reloaded_results)
print('Results from the model in memory:')
print_my_examples(examples, original_results)
```
If you want to use your model on [TF Serving](https://www.tensorflow.org/tfx/guide/serving), remember that it will call your SavedModel through one of its named signatures. In Python, you can test them as follows:
```
serving_results = reloaded_model \
.signatures['serving_default'](tf.constant(examples))
serving_results = tf.sigmoid(serving_results['classifier'])
print_my_examples(examples, serving_results)
```
## Next steps
As a next step, you can try [Solve GLUE tasks using BERT on a TPU tutorial](https://www.tensorflow.org/text/tutorials/bert_glue) which runs on a TPU and shows you how to work with multiple inputs.
| github_jupyter |
```
%load_ext autoreload
%autoreload 2
%aimport utils_1_1
import pandas as pd
import numpy as np
import altair as alt
from altair_saver import save
from constants_1_1 import SITE_FILE_TYPES
from utils_1_1 import (
get_site_file_paths,
get_site_file_info,
get_site_ids,
read_full_cli_df,
get_visualization_subtitle,
get_country_color_map,
get_siteid_color_maps,
apply_theme,
)
from web import for_website
alt.data_transformers.disable_max_rows() # Allow using rows more than 5000
df = read_full_cli_df()
df
pdf = df
# Negative values to zeros
pdf.loc[pdf['num_patients_all_still_in_hospital'] < 0, 'num_patients_all_still_in_hospital'] = np.nan
pdf.loc[pdf['num_patients_ever_severe_still_in_hospital'] < 0, 'num_patients_ever_severe_still_in_hospital'] = np.nan
# add never severe values
pdf['num_patients_never_severe_in_hospital'] = pdf['num_patients_all_still_in_hospital'] - pdf['num_patients_ever_severe_still_in_hospital']
pdf = pdf.rename(columns={
'num_patients_all_still_in_hospital': 'all',
'num_patients_ever_severe_still_in_hospital': 'ever severe',
'num_patients_never_severe_in_hospital': 'never severe'
})
# Wide to long
pdf = pd.melt(pdf, id_vars=[
'siteid',
'days_since_admission',
'color',
'country'
])
pdf = pdf.rename(columns={"variable": 'category', "value": 'num_patients'})
pdf
COUNTRY_NAMES = list(get_country_color_map().keys())
COUNTRY_COLORS = list(get_country_color_map().values())
COUNTRY_COLOR_MAP = {COUNTRY_NAMES[i]: COUNTRY_COLORS[i] for i in range(len(COUNTRY_NAMES))}
COUNTRY_COLOR_MAP
SITE_NAMES = list(get_siteid_color_maps().keys())
SITE_COLORS = list(get_siteid_color_maps().values())
SITE_COLOR_MAP = {SITE_NAMES[i]: SITE_COLORS[i] for i in range(len(SITE_NAMES))}
SITE_COLOR_MAP
def line(
df=None,
c_field=None,
c_domain=None,
c_range=None,
x_field=None,
y_field=None,
title=''
):
# Selections
legend_selection = alt.selection_multi(fields=[c_field], bind="legend")
nearest = alt.selection(type="single", nearest=True, on="mouseover", encodings=["x", "y"], empty='none', clear="mouseout")
# Rule
rule = (
alt.Chart(df)
.mark_rule(color="red")
.encode(x=f"{x_field}:Q", size=alt.value(0.5))
.transform_filter(nearest)
)
line = (
alt.Chart(df)
.mark_line(size=3)
.encode(
x=alt.X(
f"{x_field}:Q",
scale=alt.Scale(nice=False, clamp=False, padding=10),
title="Days since positive"
),
y=alt.Y(
f'sum({y_field}):Q',
axis=alt.Axis(format="r"),
scale=alt.Scale(nice=False, clamp=False, padding=10),
title="Number of patients"
),
color=alt.Color(c_field, scale=alt.Scale(domain=c_domain, range=c_range), title=None),
opacity=alt.value(0.7),
tooltip=[
alt.Tooltip(c_field, title="Country"),
alt.Tooltip(f'sum({y_field}):Q', title="Number of patients", format="r"),
alt.Tooltip(x_field, title="Days since positive")
]
)
)
circle_size = 10
circle = (
line
.mark_circle(size=circle_size, opacity=0.7)
.encode(size=alt.condition(~nearest, alt.value(circle_size), alt.value(circle_size * 2)))
.add_selection(nearest)
)
plot = (
(line + circle.add_selection(nearest) + rule)
.transform_filter(alt.FieldOneOfPredicate(field=c_field, oneOf=c_domain))
.transform_filter(legend_selection)
.add_selection(legend_selection)
.properties(
title={
"text": title,
"subtitle": get_visualization_subtitle(data_release='06-16-2020'),
"subtitleColor": "gray",
},
width=500, height=400
)
.interactive()
)
return plot
ever = line(
df=pdf[pdf['category'] == 'ever severe'],
c_field='country',
c_domain=list(df['country'].unique()),
c_range=list(df['color'].unique()),
x_field='days_since_admission',
y_field='num_patients',
title='Ever Severe Patients Still In Hospital'
)
never = line(
df=pdf[pdf['category'] == 'never severe'],
c_field='country',
c_domain=list(df['country'].unique()),
c_range=list(df['color'].unique()),
x_field='days_since_admission',
y_field='num_patients',
title='Never Severe Patients Still In Hospital'
)
final = apply_theme((ever | never).resolve_scale(y='shared'), legend_orient="right")
for_website(final, "clinicalcourse1.1", "clinicalcourse1.1")
fpdf = pdf[pdf['category'] != 'all']
plot = line(
df=fpdf,
c_field='category',
c_domain=['ever severe', 'never severe'],
c_range=['#DC3A11', '#3366CC'],
x_field='days_since_admission',
y_field='num_patients',
title='Ever and Never Severe Patients Still In Hospital'
)
apply_theme(plot.resolve_scale(y='shared'), legend_orient="right")
```
| github_jupyter |
```
CSV_PATH = "/home/zhimin90/CPT/CSVs/"
from datetime import timedelta, date
from dateutil.relativedelta import relativedelta
import pandas as pd
import numpy as np
from sodapy import Socrata
import dill
from sklearn.preprocessing import MinMaxScaler,StandardScaler
from KDEpy import FFTKDE, NaiveKDE
client = Socrata("data.cityofchicago.org", None)
results = client.get("wqdh-9gek",order="request_date DESC", limit=100000)
# Convert to pandas DataFrame
results_df = pd.DataFrame.from_records(results)
test_df = results_df
test_df.columns = pd.Series(test_df.columns).apply(lambda x: x.upper()).values
xbound = (-87.9361,-87.5245)
ybound = (41.6447,42.023)
test_df = test_df[test_df.LATITUDE.notna()].sort_values(['REQUEST_DATE','COMPLETION_DATE'], ascending=[0,0])
test_df['REQUEST_DATE'] = pd.to_datetime(test_df['REQUEST_DATE'])
test_df['COMPLETION_DATE'] = pd.to_datetime(test_df['COMPLETION_DATE'])
test_df['LATITUDE'] = pd.to_numeric(test_df['LATITUDE'])
test_df['LONGITUDE'] = pd.to_numeric(test_df['LONGITUDE'])
df = test_df
map_arr = []
interval_int = 30 #use 30 days data to predict next 7 days
series_range = 7 #days
time_interval = timedelta(days=interval_int)
date_start = min(df['REQUEST_DATE'])
date_end = max(df['REQUEST_DATE'])
geo_price_map = df[['REQUEST_DATE', 'COMPLETION_DATE','LATITUDE', 'LONGITUDE']]
filter1a = pd.to_numeric(geo_price_map["LONGITUDE"]) > xbound[0]
filter1b = pd.to_numeric(geo_price_map["LONGITUDE"]) < xbound[1]
filter1c = pd.to_numeric(geo_price_map["LATITUDE"]) > ybound[0]
filter1d = pd.to_numeric(geo_price_map["LATITUDE"]) < ybound[1]
print("sum of remaining is: " + str(sum(filter1a&filter1b&filter1c&filter1d)))
geo_price_map = geo_price_map[filter1a&filter1b&filter1c&filter1d]
for int_cur_date in range(0, (date_end - date_start).days - interval_int, int(series_range)):
geo_price_map_filtered = geo_price_map[geo_price_map['LONGITUDE'].notnull()]
filter2 = geo_price_map_filtered['REQUEST_DATE'] > (date_end - timedelta(days=int_cur_date+interval_int))
filter3 = geo_price_map_filtered['REQUEST_DATE'] <= (date_end - timedelta(days=int_cur_date))
print(date_end - timedelta(days=int_cur_date+interval_int))
print(date_end - timedelta(days=int_cur_date))
geo_price_map_filtered = geo_price_map_filtered.where(filter2 & filter3)
print("pothole count: " + str(len(geo_price_map_filtered.notnull().index)))
print("_"*20)
map_arr.append(geo_price_map_filtered)
map_arr.reverse()
pothole_count = []
for df in map_arr:
pothole_count.append(df.count())
def get_kde( x, y, xmin, xmax, ymin, ymax, xx, yy, positions):
values = np.array([x, y]).T
#values = values.reshape(values.shape[1], values.shape[0])
#print("values is: " + str(values))
#grid, points = get_kernel(values)
points = get_kernel(values, positions)
#kernel.set_bandwidth(bw_method=kernel.factor / 30.)
f = np.reshape(points, xx.shape)
#print(points.shape)
#print(grid)
#return grid, f
return f
def get_kernel(data, positions):
#print(data.shape)
#print(data)
estimator = FFTKDE(kernel='gaussian', norm=2, bw=0.001)
#grid, points = estimator.fit(data, weights=None).evaluate(grid_size)
points = estimator.fit(data, weights=None).evaluate(positions)
#grid, points = estimator.fit(data, weights=None).evaluate(grid_size)
#kernel = gaussian_kde(dataset=values, bw_method="silverman" )
#return grid, points
return points
grid_size = 1000
density_matrix_t_series = []
# Define the borders
x = [-87.9361,-87.5245]
y = [41.6447,42.023]
deltaX = (max(x) - min(x))/10
deltaY = (max(y) - min(y))/10
xmin = min(x) - deltaX
xmax = max(x) + deltaX
ymin = min(y) - deltaY
ymax = max(y) + deltaY
xx, yy = np.mgrid[xmin:xmax:(grid_size*1j), ymin:ymax:(grid_size*1j)]
positions = np.dstack([xx.ravel(), yy.ravel()])
positions = positions.reshape(positions.shape[1], positions.shape[2])
grid_matrix = positions
for i, df in enumerate(map_arr):
if df["LONGITUDE"].count() > 400:
#grid, points = get_kde(df["LONGITUDE"].dropna().to_numpy(), df["LATITUDE"].dropna().to_numpy() , xmin, xmax, ymin, ymax, xx, yy, positions)
points = get_kde(df["LONGITUDE"].dropna().to_numpy(), df["LATITUDE"].dropna().to_numpy() , xmin, xmax, ymin, ymax, xx, yy, positions)
density_matrix_t_series.append(points)
print("@" + str(i))
s = round(len(density_matrix_t_series)*0)
f_in = open(CSV_PATH +'Scalers_2020.pkl', "rb")
scaler,scaler2 = dill.load(f_in)
f_in.close()
dm_series_np = np.array(density_matrix_t_series[s:])
flattened_matrix_np = np.reshape(dm_series_np, (dm_series_np.shape[0]*dm_series_np.shape[1], dm_series_np.shape[2]))
normalized_matrices_test = scaler2.transform(scaler.transform(flattened_matrix_np))
x_test = normalized_matrices_test[0:-normalized_matrices_test.shape[1]].copy()
y_test = normalized_matrices_test[normalized_matrices_test.shape[1]-1:-1].copy()
x_test2 = np.reshape(x_test, (x_test.shape[0], 1, x_test.shape[1]))
y_test2 = y_test
import tensorflow.keras as keras
model = keras.models.load_model(CSV_PATH + 'TensorFlowModel_2020_train_save')
from datetime import timedelta, date
from dateutil.relativedelta import relativedelta
def predictor(model, data_in, grid, start_frame_date, end_frame_date, time_shift):
xx, yy = grid
offset = yy.shape[0]
print("offset = yy.shape[0]" + str(offset))
xx = xx.ravel()
yy = yy.ravel()
xdelta = abs(xx[1] - xx[1+offset])
ydelta = abs(yy[0] - yy[1+offset])
print("xdelta"+str(xdelta))
print("ydelta"+str(ydelta))
columns = [ 'start_date', 'end_date', 'poly_coordinate', 'density']
pred = model.predict(data_in)
data = scaler.inverse_transform(scaler2.inverse_transform(pred))
data_reshaped = data.reshape((int(data.shape[0]/data.shape[1]), data.shape[1], data.shape[1]))
print(data_reshaped.shape)
#each cell is a density estimate from KDE that that has been aggregated by number of potholes over time
#This time interval of density cell is input frame time + timeshift the target frame in the model that has shifted forward by
row_dict = {'start_date' : None, 'end_date' : None, 'poly_coordinate': None, 'density': 0}
#append = pd.DataFrame(columns=columns)
dict_list = []
for t, matrix in enumerate(data_reshaped):
xy_matrix = np.flip(np.rot90(matrix),0)
print(xy_matrix.shape)
row_dict['start_date'] = pd.to_datetime(start_frame_date) + timedelta(days=(time_shift*(t+1)))
row_dict['end_date'] = pd.to_datetime(end_frame_date) + timedelta(days=(time_shift*(t+1)))
for i, row in enumerate(xy_matrix):
for j, cell in enumerate(row):
pos_index = i + j*xy_matrix.shape[1]
#generate density cell (square) polycoordinate [[cxmin,cymin],[cxmax, cymin],[cxmin, cymax],[cxmax, cymax]]
row_dict['poly_coordinate'] = [[xx[pos_index],yy[pos_index]],[xx[pos_index]+xdelta,yy[pos_index]],[xx[pos_index]+xdelta,yy[pos_index]+ydelta], [xx[pos_index],yy[pos_index]+ydelta]]
row_dict['density'] = cell
dict_list.append(row_dict.copy())
return pd.DataFrame(dict_list)
Last_time_frame = y_test2[-(y_test2.shape[1]+1):-1]
start_frame_date = min(map_arr[-1]['REQUEST_DATE'][map_arr[-1]['REQUEST_DATE'].notna()])
end_frame_date = max(map_arr[-1]['REQUEST_DATE'][map_arr[-1]['REQUEST_DATE'].notna()])
time_shift = 7 #days
dataframe = predictor(model,np.reshape(Last_time_frame,(Last_time_frame.shape[0],1,Last_time_frame.shape[1])), (xx, yy), start_frame_date, end_frame_date, time_shift)
import geopandas as gpd
from shapely.geometry import Polygon
df = dataframe
df["int_density"] = df.density.astype(int)
list = []
for index, row in df.iterrows():
list.append( [row['start_date'], row['end_date'],Polygon( row['poly_coordinate']), row['density'], row['int_density']] )
gdf = gpd.GeoDataFrame(list, columns =['start_date','end_date', 'geometry', 'density', 'int_density'])
xmin, ymin, xmax, ymax = gdf.total_bounds
grid_size = 10
xgrid = np.arange(xmin, xmax, (xmax-xmin)/grid_size)
ygrid = np.arange(ymin, ymax, (ymax-ymin)/grid_size)
print(xgrid,ygrid)
c = 0
gdf["zone"] = None
for row in xgrid:
for col in ygrid:
boundbox = Polygon([[row,col],[row+(xmax-xmin)/grid_size,col],[row+(xmax-xmin)/grid_size,col+(ymax-ymin)/grid_size],[row,col+(ymax-ymin)/grid_size],[row,col]])
bb_df = gpd.GeoSeries(boundbox)
bool_within_bb = gdf.geometry.intersects(boundbox)
index_within_bb = gdf[bool_within_bb].index
gdf.iloc[index_within_bb,5] = c
c+=1
print(c)
print("count rows within count: " + str(len(index_within_bb)))
print("-"*25)
gdf[gdf["zone"].isnull()]
gdf_dissolved = gdf.dissolve(by=['int_density','zone'])
gdf_dissolved
max(gdf_dissolved["density"])
gdf_dissolved[["geometry","density"]].plot(column='density',figsize=(10,10))
gdf_dissolved.iloc[:2,]
def applyInsert(geometry,start_d,end_d,density):
print(geometry,start_d,end_d,density)
print("-"*100)
gdf_dissolved.iloc[:5,:].apply(lambda row: applyInsert(row.geometry, row.start_date, row.end_date, row.density), axis=1)
gdf_dissolved.iloc[:5,:]
```
| github_jupyter |
```
import pickle
import numpy as np
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Sequential
from keras.layers import *
from keras import Model
from sklearn.model_selection import train_test_split
from keras.callbacks import EarlyStopping
import keras.metrics
import tensorflow as tf
from sklearn.metrics import precision_recall_curve
from sklearn.metrics import classification_report
from torchtext import data
import pandas as pd
from sklearn.metrics import accuracy_score
from torch.utils.data import Dataset, DataLoader
from collections import Counter
from sklearn.utils import class_weight
from sklearn.metrics import accuracy_score
from sklearn.metrics import precision_recall_fscore_support
def read_data(filename):
dfile = open(filename, 'rb')
data = pickle.load(dfile)
dfile.close()
return data
X_train, Y_train, labels_train = read_data('TCR/data_train_tcr')
X_test, Y_test, labels_test = read_data('TCR/data_test_tcr')
unique_tokens = read_data('TCR/unique_tokens_tcr')
MAX_NB_WORDS = 5000
MAX_SEQUENCE_LENGTH = 175
EMBEDDING_DIM = 300
VAL_SIZE = 0.15
unique_pos, unique_deps, unique_words = unique_tokens[0], unique_tokens[1], unique_tokens[2]
tokenizer1 = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer1.fit_on_texts(unique_pos)
word_index1 = tokenizer1.word_index
tokenizer2 = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer2.fit_on_texts(unique_words)
word_index2 = tokenizer2.word_index
tokenizer3 = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer3.fit_on_texts(unique_deps)
word_index3 = tokenizer3.word_index
#train
seq1 = tokenizer1.texts_to_sequences(X_train[0])
seq11 = pad_sequences(seq1, maxlen=MAX_SEQUENCE_LENGTH)
seq2 = tokenizer2.texts_to_sequences(X_train[2])
seq12 = pad_sequences(seq2, maxlen=MAX_SEQUENCE_LENGTH)
seq3 = tokenizer3.texts_to_sequences(X_train[1])
seq13 = pad_sequences(seq3, maxlen=MAX_SEQUENCE_LENGTH)
#test
seq1 = tokenizer1.texts_to_sequences(X_test[0])
seq11_test = pad_sequences(seq1, maxlen=MAX_SEQUENCE_LENGTH)
seq2 = tokenizer2.texts_to_sequences(X_test[2])
seq12_test = pad_sequences(seq2, maxlen=MAX_SEQUENCE_LENGTH)
seq3 = tokenizer3.texts_to_sequences(X_test[1])
seq13_test = pad_sequences(seq3, maxlen=MAX_SEQUENCE_LENGTH)
# FOLDS 5-fold cross validation
# fold 1
nb_validation_samples = int(VAL_SIZE*seq11.shape[0])
fold1_x_train1 = seq11[:-nb_validation_samples]
fold1_x_train2 = seq12[:-nb_validation_samples]
fold1_x_train3 = seq13[:-nb_validation_samples]
fold1_y_train = Y_train[:-nb_validation_samples]
fold1_lab_train = labels_train[:-nb_validation_samples]
fold1_x_val1 = seq11[-nb_validation_samples:]
fold1_x_val2 = seq12[-nb_validation_samples:]
fold1_x_val3 = seq13[-nb_validation_samples:]
fold1_y_val = Y_train[-nb_validation_samples:]
fold1_lab_val = labels_train[-nb_validation_samples:]
# FOLD 2
# print(nb_validation_samples, seq11.shape, seq12.shape, seq13.shape)
fold2_x_train1 = np.concatenate((seq11[:-2*nb_validation_samples],seq11[-nb_validation_samples:]))
fold2_x_train2 = np.concatenate((seq12[:-2*nb_validation_samples],seq12[-nb_validation_samples:]))
fold2_x_train3 = np.concatenate((seq13[:-2*nb_validation_samples],seq13[-nb_validation_samples:]))
fold2_y_train = np.concatenate((Y_train[:-2*nb_validation_samples], Y_train[-nb_validation_samples:]))
fold2_lab_train = np.concatenate((labels_train[:-2*nb_validation_samples],labels_train[-nb_validation_samples:]))
fold2_x_val1 = seq11[-2*nb_validation_samples:-nb_validation_samples]
fold2_x_val2 = seq12[-2*nb_validation_samples:-nb_validation_samples]
fold2_x_val3 = seq13[-2*nb_validation_samples:-nb_validation_samples]
fold2_y_val = Y_train[-2*nb_validation_samples:-nb_validation_samples]
fold2_lab_val = labels_train[-2*nb_validation_samples:-nb_validation_samples]
#fold 3
fold3_x_train1 = np.concatenate((seq11[:-3*nb_validation_samples],seq11[-2*nb_validation_samples:]))
fold3_x_train2 = np.concatenate((seq12[:-3*nb_validation_samples],seq12[-2*nb_validation_samples:]))
fold3_x_train3 = np.concatenate((seq13[:-3*nb_validation_samples],seq13[-2*nb_validation_samples:]))
fold3_y_train = np.concatenate((Y_train[:-3*nb_validation_samples], Y_train[-2*nb_validation_samples:]))
fold3_lab_train = np.concatenate((labels_train[:-3*nb_validation_samples],labels_train[-2*nb_validation_samples:]))
fold3_x_val1 = seq11[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_x_val2 = seq12[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_x_val3 = seq13[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_y_val = Y_train[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_lab_val = labels_train[-3*nb_validation_samples:-2*nb_validation_samples]
#fold 4
fold4_x_train1 = np.concatenate((seq11[:-4*nb_validation_samples],seq11[-3*nb_validation_samples:]))
fold4_x_train2 = np.concatenate((seq12[:-4*nb_validation_samples],seq12[-3*nb_validation_samples:]))
fold4_x_train3 = np.concatenate((seq13[:-4*nb_validation_samples],seq13[-3*nb_validation_samples:]))
fold4_y_train = np.concatenate((Y_train[:-4*nb_validation_samples], Y_train[-3*nb_validation_samples:]))
fold4_lab_train = np.concatenate((labels_train[:-4*nb_validation_samples],labels_train[-3*nb_validation_samples:]))
fold4_x_val1 = seq11[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_x_val2 = seq12[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_x_val3 = seq13[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_y_val = Y_train[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_lab_val = labels_train[-4*nb_validation_samples:-3*nb_validation_samples]
# fold 5
fold5_x_train1 = seq11[nb_validation_samples:]
fold5_x_train2 = seq12[nb_validation_samples:]
fold5_x_train3 = seq13[nb_validation_samples:]
fold5_y_train = Y_train[nb_validation_samples:]
fold5_lab_train = labels_train[nb_validation_samples:]
fold5_x_val1 = seq11[0:nb_validation_samples]
fold5_x_val2 = seq12[0:nb_validation_samples]
fold5_x_val3 = seq13[0:nb_validation_samples]
fold5_y_val = Y_train[0:nb_validation_samples]
fold5_lab_val = labels_train[0:nb_validation_samples]
pos_vec = read_data('pos.vector')
dep_vec = read_data('deps.vector')
word_vec = {}
word_vec['PADDING'] = 300
f = open('glove.42B.300d.txt')
for line in f:
values = line.split()
word = values[0]
coefs = np.asarray(values[1:], dtype='float32')
word_vec[word.lower()] = line
f.close()
# pos tags
embedding_matrix1 = np.zeros((len(word_index1) + 1, 28))
for word, i in word_index1.items():
embedding_vector = pos_vec.get(word)
if embedding_vector is not None:
embedding_matrix1[i] = np.asarray(embedding_vector.split()[1:], dtype='float32')
#word vec
embedding_matrix2 = np.zeros((len(word_index2) + 1, EMBEDDING_DIM))
for word, i in word_index2.items():
embedding_vector = word_vec.get(word)
if embedding_vector is not None:
embedding_matrix2[i] = np.asarray(embedding_vector.split()[1:], dtype='float32')
# deps vec
embedding_matrix3 = np.zeros((len(word_index3) + 1, len(dep_vec['PADDING'])))
for word, i in word_index3.items():
embedding_vector = dep_vec.get(word)
if embedding_vector is not None:
embedding_matrix3[i] = np.asarray(embedding_vector, dtype='float32')
def get_class_weights(training_labels):
class_weights = class_weight.compute_class_weight('balanced',np.unique(training_labels),training_labels)
uni = list(np.unique(training_labels))
labelset = ['CLINK', 'CLINK-R', 'O']
weights = []
for i in labelset:
try:
idx = uni.index(i)
weights.append(class_weights[idx])
except:
weights.append(0)
return weights
```
# To Extract Causal Features
```
def defineModel(l1,l2,l3,l4,d1,out,d):
embedding_layer1 = Embedding(len(word_index2) + 1,EMBEDDING_DIM,weights=[embedding_matrix2],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer2 = Embedding(len(word_index1) + 1,28,weights=[embedding_matrix1],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer3 = Embedding(len(word_index3) + 1,77,weights=[embedding_matrix3],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
wi = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
wi2 = embedding_layer1(wi)
pi_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
pi2_sen = embedding_layer2(pi_sen)
di_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
di2_sen = embedding_layer3(di_sen)
lstm1_sen = Bidirectional(LSTM(l1, activation='tanh', dropout=d, return_sequences=True), name = 'bid1causal_sen')(pi2_sen) # pos encoded features
lstm2_sen = Bidirectional(LSTM(l2, activation='tanh', dropout=d, return_sequences=True), name= 'bid2causal_sen')(di2_sen) # dep features
lstm3 = Bidirectional(LSTM(l4, activation='tanh', dropout=d+0.1, return_sequences=True), name = 'bid3causal')(wi2) # woed features
hid_sen = concatenate([lstm1_sen, lstm2_sen, lstm3])
lstm5 = Bidirectional(LSTM(l4, activation='tanh', dropout=d), name = 'bid3causallstm2_sen')(hid_sen)
yii = Dense(d1, activation='relu', name='dense1')(lstm5)
yi = Dense(out, activation="softmax", name='dense2')(yii)
model = Model(inputs=[pi_sen,di_sen,wi],outputs=yi)
return model
def getfolddata(num):
if num==1:
return [fold1_x_train1,fold1_x_train3,fold1_x_train2], fold1_y_train, fold1_lab_train, [fold1_x_val1,fold1_x_val3,fold1_x_val2] , fold1_y_val, fold1_lab_val
elif num==2:
return [fold2_x_train1,fold2_x_train3,fold2_x_train2], fold2_y_train, fold2_lab_train, [fold2_x_val1,fold2_x_val3,fold2_x_val2] , fold2_y_val, fold2_lab_val
elif num==3:
return [fold3_x_train1,fold3_x_train3,fold3_x_train2], fold3_y_train, fold3_lab_train, [fold3_x_val1,fold3_x_val3,fold3_x_val2] , fold3_y_val, fold3_lab_val
elif num==4:
return [fold4_x_train1,fold4_x_train3,fold4_x_train2], fold4_y_train, fold4_lab_train, [fold4_x_val1,fold4_x_val3,fold4_x_val2] , fold4_y_val, fold4_lab_val
elif num==5:
return [fold5_x_train1,fold5_x_train3,fold5_x_train2], fold5_y_train, fold5_lab_train, [fold5_x_val1,fold5_x_val3,fold5_x_val2] , fold5_y_val, fold5_lab_val
def trainModel():
num_classes = 3
epochs = 50
batchsize = 64
lrs = [0.001,0.1,0.1,0.001,0.01]
drop = [0.1,0.2,0.3,0.3,0.2]
file1 = 'TCR/chkpt/'
out = num_classes
for fold in [1,2,3,4,5]:
checkpoint_filepath = file1 + f'model_causal_tcr_fold{fold}'
training_data, y_train, training_labels, val_data, y_val, val_labels = getfolddata(fold)
weights = get_class_weights(training_labels)
set_nodes = [32, 32, 64, 64, 32]
l1 = set_nodes[0]
l2 = set_nodes[1]
l3 = set_nodes[2]
l4 = set_nodes[3]
d1 = set_nodes[4]
d = drop[fold-1]
lr = lrs[fold-1]
optimizer = tf.keras.optimizers.RMSprop(learning_rate=lr)
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_filepath,save_weights_only=True,monitor='val_accuracy',mode='max',save_best_only=True)
callback = tf.keras.callbacks.EarlyStopping(monitor='val_accuracy', patience=20)
model = defineModel(l1,l2,l3,l4,d1,out,d)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'], loss_weights=weights)
model.fit(x = training_data, y = y_train, epochs = epochs, batch_size = batchsize,validation_data=(val_data,y_val), callbacks=[callback, model_checkpoint_callback], verbose=0)
model.load_weights(checkpoint_filepath)
model.save(f"tcr_causal_fold{fold}.h5")
del model
trainModel()
```
# To Extract Temporal Features
```
X_train_temp, Y_train_temp, labels_train_temp = read_data('TCR/data_train_temporal_tcr')
unique_tokens_temp = read_data('TCR/unique_tokens_temporal_tcr')
unique_pos_temp, unique_deps_temp, unique_words_temp = unique_tokens[0], unique_tokens[1], unique_tokens[2]
tokenizer1_temp = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer1_temp.fit_on_texts(unique_pos_temp)
word_index1_temp = tokenizer1_temp.word_index
tokenizer2_temp = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer2_temp.fit_on_texts(unique_words_temp)
word_index2_temp = tokenizer2_temp.word_index
tokenizer3_temp = Tokenizer(num_words=MAX_NB_WORDS)
tokenizer3_temp.fit_on_texts(unique_deps_temp)
word_index3_temp = tokenizer3_temp.word_index
#train
seq1_temp = tokenizer1_temp.texts_to_sequences(X_train_temp[0])
seq11_temp = pad_sequences(seq1_temp, maxlen=MAX_SEQUENCE_LENGTH)
seq2_temp = tokenizer2_temp.texts_to_sequences(X_train_temp[2])
seq12_temp = pad_sequences(seq2_temp, maxlen=MAX_SEQUENCE_LENGTH)
seq3_temp = tokenizer3_temp.texts_to_sequences(X_train_temp[1])
seq13_temp = pad_sequences(seq3_temp, maxlen=MAX_SEQUENCE_LENGTH)
# FOLDS 5-fold cross validation
# fold 1
nb_validation_samples = int(VAL_SIZE*seq11.shape[0])
fold1_x_train1_temp = seq11_temp[:-nb_validation_samples]
fold1_x_train2_temp = seq12_temp[:-nb_validation_samples]
fold1_x_train3_temp = seq13_temp[:-nb_validation_samples]
fold1_y_train_temp = Y_train_temp[:-nb_validation_samples]
fold1_lab_train_temp = labels_train_temp[:-nb_validation_samples]
fold1_x_val1_temp = seq11_temp[-nb_validation_samples:]
fold1_x_val2_temp = seq12_temp[-nb_validation_samples:]
fold1_x_val3_temp = seq13_temp[-nb_validation_samples:]
fold1_y_val_temp = Y_train_temp[-nb_validation_samples:]
fold1_lab_val_temp = labels_train_temp[-nb_validation_samples:]
# FOLD 2
# print(nb_validation_samples, seq11.shape, seq12.shape, seq13.shape)
fold2_x_train1_temp = np.concatenate((seq11_temp[:-2*nb_validation_samples],seq11_temp[-nb_validation_samples:]))
fold2_x_train2_temp = np.concatenate((seq12_temp[:-2*nb_validation_samples],seq12_temp[-nb_validation_samples:]))
fold2_x_train3_temp = np.concatenate((seq13_temp[:-2*nb_validation_samples],seq13_temp[-nb_validation_samples:]))
fold2_y_train_temp = np.concatenate((Y_train_temp[:-2*nb_validation_samples], Y_train_temp[-nb_validation_samples:]))
fold2_lab_train_temp = np.concatenate((labels_train_temp[:-2*nb_validation_samples],labels_train_temp[-nb_validation_samples:]))
fold2_x_val1_temp = seq11_temp[-2*nb_validation_samples:-nb_validation_samples]
fold2_x_val2_temp = seq12_temp[-2*nb_validation_samples:-nb_validation_samples]
fold2_x_val3_temp = seq13_temp[-2*nb_validation_samples:-nb_validation_samples]
fold2_y_val_temp = Y_train_temp[-2*nb_validation_samples:-nb_validation_samples]
fold2_lab_val_temp = labels_train_temp[-2*nb_validation_samples:-nb_validation_samples]
#fold 3
fold3_x_train1_temp = np.concatenate((seq11_temp[:-3*nb_validation_samples],seq11_temp[-2*nb_validation_samples:]))
fold3_x_train2_temp = np.concatenate((seq12_temp[:-3*nb_validation_samples],seq12_temp[-2*nb_validation_samples:]))
fold3_x_train3_temp = np.concatenate((seq13_temp[:-3*nb_validation_samples],seq13_temp[-2*nb_validation_samples:]))
fold3_y_train_temp = np.concatenate((Y_train_temp[:-3*nb_validation_samples], Y_train_temp[-2*nb_validation_samples:]))
fold3_lab_train_temp = np.concatenate((labels_train_temp[:-3*nb_validation_samples],labels_train_temp[-2*nb_validation_samples:]))
fold3_x_val1_temp = seq11_temp[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_x_val2_temp = seq12_temp[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_x_val3_temp = seq13_temp[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_y_val_temp = Y_train_temp[-3*nb_validation_samples:-2*nb_validation_samples]
fold3_lab_val_temp = labels_train_temp[-3*nb_validation_samples:-2*nb_validation_samples]
#fold 4
fold4_x_train1_temp = np.concatenate((seq11_temp[:-4*nb_validation_samples],seq11_temp[-3*nb_validation_samples:]))
fold4_x_train2_temp = np.concatenate((seq12_temp[:-4*nb_validation_samples],seq12_temp[-3*nb_validation_samples:]))
fold4_x_train3_temp = np.concatenate((seq13_temp[:-4*nb_validation_samples],seq13_temp[-3*nb_validation_samples:]))
fold4_y_train_temp = np.concatenate((Y_train_temp[:-4*nb_validation_samples], Y_train_temp[-3*nb_validation_samples:]))
fold4_lab_train_temp = np.concatenate((labels_train_temp[:-4*nb_validation_samples],labels_train_temp[-3*nb_validation_samples:]))
fold4_x_val1_temp = seq11_temp[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_x_val2_temp = seq12_temp[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_x_val3_temp = seq13_temp[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_y_val_temp = Y_train_temp[-4*nb_validation_samples:-3*nb_validation_samples]
fold4_lab_val_temp = labels_train_temp[-4*nb_validation_samples:-3*nb_validation_samples]
# fold 5
fold5_x_train1_temp = seq11_temp[nb_validation_samples:]
fold5_x_train2_temp = seq12_temp[nb_validation_samples:]
fold5_x_train3_temp = seq13_temp[nb_validation_samples:]
fold5_y_train_temp = Y_train_temp[nb_validation_samples:]
fold5_lab_train_temp = labels_train_temp[nb_validation_samples:]
fold5_x_val1_temp = seq11_temp[0:nb_validation_samples]
fold5_x_val2_temp = seq12_temp[0:nb_validation_samples]
fold5_x_val3_temp = seq13_temp[0:nb_validation_samples]
fold5_y_val_temp = Y_train_temp[0:nb_validation_samples]
fold5_lab_val_temp = labels_train_temp[0:nb_validation_samples]
def getfolddata_temp(num):
if num==1:
return [fold1_x_train1_temp,fold1_x_train3_temp,fold1_x_train2_temp], fold1_y_train_temp, fold1_lab_train_temp, [fold1_x_val1_temp,fold1_x_val3_temp,fold1_x_val2_temp] , fold1_y_val_temp, fold1_lab_val_temp
elif num==2:
return [fold2_x_train1_temp,fold2_x_train3_temp,fold2_x_train2_temp], fold2_y_train_temp, fold2_lab_train_temp, [fold2_x_val1_temp,fold2_x_val3_temp,fold2_x_val2_temp] , fold2_y_val_temp, fold2_lab_val_temp
elif num==3:
return [fold3_x_train1_temp,fold3_x_train3_temp,fold3_x_train2_temp], fold3_y_train_temp, fold3_lab_train_temp, [fold3_x_val1_temp,fold3_x_val3_temp,fold3_x_val2_temp] , fold3_y_val_temp, fold3_lab_val_temp
elif num==4:
return [fold4_x_train1_temp,fold4_x_train3_temp,fold4_x_train2_temp], fold4_y_train_temp, fold4_lab_train_temp, [fold4_x_val1_temp,fold4_x_val3_temp,fold4_x_val2_temp] , fold4_y_val_temp, fold4_lab_val_temp
elif num==5:
return [fold5_x_train1_temp,fold5_x_train3_temp,fold5_x_train2_temp], fold5_y_train_temp, fold5_lab_train_temp, [fold5_x_val1_temp,fold5_x_val3_temp,fold5_x_val2_temp] , fold5_y_val_temp, fold5_lab_val_temp
# pos tags
embedding_matrix1 = np.zeros((len(word_index1) + 1, 28))
for word, i in word_index1.items():
embedding_vector = pos_vec.get(word)
if embedding_vector is not None:
embedding_matrix1[i] = np.asarray(embedding_vector.split()[1:], dtype='float32')
#word vec
embedding_matrix2 = np.zeros((len(word_index2) + 1, EMBEDDING_DIM))
for word, i in word_index2.items():
embedding_vector = word_vec.get(word)
if embedding_vector is not None:
embedding_matrix2[i] = np.asarray(embedding_vector.split()[1:], dtype='float32')
# deps vec
embedding_matrix3 = np.zeros((len(word_index3) + 1, len(dep_vec['PADDING'])))
for word, i in word_index3.items():
embedding_vector = dep_vec.get(word)
if embedding_vector is not None:
embedding_matrix3[i] = np.asarray(embedding_vector, dtype='float32')
def get_class_weights_temp(training_labels):
class_weights = class_weight.compute_class_weight('balanced',np.unique(training_labels),training_labels)
uni = list(np.unique(training_labels))
labelset = ['AFTER', 'BEFORE', 'SIMULTANEOUS']
weights = []
for i in labelset:
try:
idx = uni.index(i)
weights.append(class_weights[idx]*2)
except:
weights.append(0)
return weights
def defineModel(l1,l2,l3,l4,d1,out,d):
embedding_layer1 = Embedding(len(word_index2) + 1,EMBEDDING_DIM,weights=[embedding_matrix2],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer2 = Embedding(len(word_index1) + 1,28,weights=[embedding_matrix1],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer3 = Embedding(len(word_index3) + 1,77,weights=[embedding_matrix3],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
wi = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
wi2 = embedding_layer1(wi)
pi_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
pi2_sen = embedding_layer2(pi_sen)
di_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
di2_sen = embedding_layer3(di_sen)
lstm1_sen = Bidirectional(LSTM(l1, activation='tanh', dropout=d, return_sequences=True), name = 'bid1temp_sen')(pi2_sen) # pos features
lstm2_sen = Bidirectional(LSTM(l2, activation='tanh', dropout=d, return_sequences=True), name= 'bid2temp_sen')(di2_sen) # dep features
lstm3 = Bidirectional(LSTM(l4, activation='tanh', dropout=d+0.1, return_sequences=True), name = 'bid3temp')(wi2) # woed features
hid_sen = concatenate([lstm1_sen, lstm2_sen, lstm3])
lstm5 = Bidirectional(LSTM(l4, activation='tanh', dropout=d), name = 'bid3templstm2_sen')(hid_sen)
yii = Dense(d1, activation='relu', name='dense1temp')(lstm5)
yi = Dense(out, activation="softmax", name='dense2temp')(yii)
model = Model(inputs=[pi_sen,di_sen,wi],outputs=yi)
return model
def trainModel_temporal():
num_classes = 3
epochs = 50
batchsize = 64
lr = 0.005
d = 0.3
file1 = 'TCR/chkpt/'
out = num_classes
for fold in [1,2,3,4,5]:
checkpoint_filepath = file1 + f'model_temp_tcr_fold{fold}'
training_data, y_train, training_labels, val_data, y_val, val_labels = getfolddata_temp(fold)
weights = get_class_weights_temp(training_labels)
set_nodes = [32, 32, 64, 64, 32]
l1 = set_nodes[0]
l2 = set_nodes[1]
l3 = set_nodes[2]
l4 = set_nodes[3]
d1 = set_nodes[4]
optimizer = tf.keras.optimizers.Adam(learning_rate=lr)
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_filepath,save_weights_only=True,monitor='val_accuracy',mode='max',save_best_only=True)
callback = tf.keras.callbacks.EarlyStopping(monitor='val_accuracy', patience=20)
model = defineModel(l1,l2,l3,l4,d1,out,d)
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'], loss_weights=weights)
model.fit(x = training_data, y = y_train, epochs = epochs, batch_size = batchsize,validation_data=(val_data,y_val), callbacks=[callback, model_checkpoint_callback])
model.load_weights(checkpoint_filepath)
model.save(f"tcr_temp_fold{fold}.h5")
del model
trainModel_temporal()
```
# Joint Model for Causal Relation Classification
```
def get_class_weights(training_labels):
class_weights = class_weight.compute_class_weight('balanced',np.unique(training_labels),training_labels)
uni = list(np.unique(training_labels))
labelset = ['CLINK', 'CLINK-R', 'O']
weights = []
for i in labelset:
try:
idx = uni.index(i)
weights.append(class_weights[idx])
except:
weights.append(0)
return weights
def defineModel(l1,l2,l3,l4,d1,out,d):
embedding_layer1 = Embedding(len(word_index2) + 1,EMBEDDING_DIM,weights=[embedding_matrix2],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer2 = Embedding(len(word_index1) + 1,28,weights=[embedding_matrix1],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
embedding_layer3 = Embedding(len(word_index3) + 1,77,weights=[embedding_matrix3],input_length=MAX_SEQUENCE_LENGTH,trainable=False)
wi = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
wi2 = embedding_layer1(wi)
pi_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
pi2_sen = embedding_layer2(pi_sen)
di_sen = Input(shape=(MAX_SEQUENCE_LENGTH,), dtype='int32')
di2_sen = embedding_layer3(di_sen)
lstm1temp = Bidirectional(LSTM(l1, activation='tanh', dropout=d, return_sequences=True), name = 'bid1temp_sen')(pi2_sen)
lstm1temp.trainable = False
lstm2temp = Bidirectional(LSTM(l2, activation='tanh', dropout=d, return_sequences=True), name= 'bid2temp_sen')(di2_sen)
lstm2temp.trainable = False
lstm3temp = Bidirectional(LSTM(l4, activation='tanh', dropout=d+0.1, return_sequences=True), name = 'bid3temp')(wi2)
lstm3temp.trainable = False
hid_temp = concatenate([lstm1temp, lstm2temp, lstm3temp])
lstm4temp = Bidirectional(LSTM(l4, activation='tanh', dropout=d), name = 'bid3templstm2_sen')(hid_temp)
lstm4temp.trainable = False
lstm1causal = Bidirectional(LSTM(l1, activation='tanh', dropout=d, return_sequences=True), name = 'bid1causal_sen')(pi2_sen)
lstm1causal.trainable = False
lstm2causal = Bidirectional(LSTM(l2, activation='tanh', dropout=d, return_sequences=True), name= 'bid2causal_sen')(di2_sen)
lstm2causal.trainable = False
lstm3causal = Bidirectional(LSTM(l3, activation='tanh', dropout=0.45, return_sequences=True), name = 'bid3causal')(wi2)
lstm3causal.trainable = False
hid_causal = concatenate([lstm1causal, lstm2causal, lstm3causal])
lstm4causal = Bidirectional(LSTM(l4, activation='tanh', dropout=d), name = 'bid3causallstm2_sen')(hid_causal)
lstm4causal.trainable = False
merged_features = concatenate([lstm4temp, lstm4causal])
yii = Dense(d1, activation='relu', name='denselayer1')(merged_features)
yi = Dense(out, activation="softmax", name='denselayer2')(yii)
model = Model(inputs=[pi_sen,di_sen,wi],outputs=yi)
return model
```
# Train
```
def trainModelJoint():
num_classes = 3
epochs = 50
batchsizes = [ 64, 64, 128, 128, 128]
lrs = [0.001, 0.001, 0.001, 0.005, 0.01]
drop = [0.1, 0.2, 0.1, 0.1, 0.1 ]
file1 = 'TCR/chkpt/'
out = num_classes
for fold in [1,2,3,4,5]:
checkpoint_filepath = file1 + f'model_joint_tcr_fold{fold}'
training_data, y_train, training_labels, val_data, y_val, val_labels = getfolddata(fold)
weights = get_class_weights_temp(training_labels)
set_nodes = [32, 32, 64, 64, 32]
l1 = set_nodes[0]
l2 = set_nodes[1]
l3 = set_nodes[2]
l4 = set_nodes[3]
d1 = set_nodes[4]
lr = lrs[fold-1]
d = drop[fold-1]
batchsize = batchsizes[fold-1]
optimizer = tf.keras.optimizers.Adam(learning_rate=lr)
model_checkpoint_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_filepath,save_weights_only=True,monitor='val_accuracy',mode='max',save_best_only=True)
callback = tf.keras.callbacks.EarlyStopping(monitor='val_accuracy', patience=20)
model = defineModel(l1,l2,l3,l4,d1,out,d)
model.load_weights(f'TCR/tcr_causal_fold{fold}.h5', by_name =True) # to extract causal features
model.load_weights(f'TCR/tcr_temp_fold{fold}.h5', by_name =True) # to extract temporal features
model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'], loss_weights=weights)
model.fit(x = training_data, y = y_train, epochs = epochs, batch_size = batchsize,validation_data=(val_data,y_val), callbacks=[callback, model_checkpoint_callback])
model.load_weights(checkpoint_filepath)
model.save(f"TCR/tcr_joint_fold{fold}.h5")
del model
model = trainModelJoint()
#save all best models
```
# Best Model Results
```
def format_report(report, scores, accuracy, fold):
# [ 'causes' ,'caused by', 'OTHER' ]
print("")
print(f"Test set result for fold {fold}")
print(f" {'{0:>10}'.format('precision')} {'{0:>10}'.format('recall')} {'{0:>10}'.format('f1-score')}")
print(f" causes {'{0:>10}'.format(round(report['0']['precision']*100.0, 1))} {'{0:>10}'.format(round(report['0']['recall']*100.0, 1))} {'{0:>10}'.format(round(report['0']['f1-score']*100.0, 1))}")
print(f" caused by {'{0:>10}'.format(round(report['1']['precision']*100.0, 1))} {'{0:>10}'.format(round(report['1']['recall']*100.0, 1))} {'{0:>10}'.format(round(report['1']['f1-score']*100.0, 1))}")
print("")
print(f" accuracy {'{0:>10}'.format('')} {'{0:>10}'.format('')} {'{0:>10}'.format(round(accuracy*100, 1))}")
print(f" micro avg {'{0:>10}'.format(round(scores[0]*100.0, 1))} {'{0:>10}'.format(round(scores[1]*100.0, 1))} {'{0:>10}'.format(round(scores[2]*100.0, 1))}")
drop = [0.1, 0.2, 0.1, 0.1, 0.1 ]
for fold in [1,2,3,4,5]:
model = defineModel(32,32,64,64,32,3, drop[fold-1])
model.load_weights(f'TCR/tcr_joint_fold{fold}.h5', by_name=True)
data_test = [seq11_test,seq13_test,seq12_test]
classes = np.argmax(model.predict(x = data_test), axis=-1)
y_test_classes = Y_test.argmax(1)
y_pred_classes = classes
accuracy = accuracy_score(y_test_classes, y_pred_classes)
report = classification_report(y_true=y_test_classes, y_pred=y_pred_classes, zero_division=0, output_dict=True, digits= 3, labels=[0,1,2,3,4,5,6,7,8,9,10,11,12,13])
scores = precision_recall_fscore_support(y_true=y_test_classes, y_pred=y_pred_classes, average='micro', labels=[0,1,2,3,4,5,6,7,8,9,10,11,12,13])
format_report(report, scores, accuracy, fold)
```
| github_jupyter |
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# 1D Alfven Wave `GiRaFFEfood` Initial Data for `GiRaFFE`
## This module provides another initial data option for `GiRaFFE`, drawn from [this paper](https://arxiv.org/abs/1310.3274) .
**Notebook Status:** <font color='orange'><b> Self-Validated </b></font>
**Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). **Additional validation tests may have been performed, but are as yet, undocumented. (TODO)**
### NRPy+ Source Code for this module: [GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py](../edit/GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py)
## Introduction:
### Alfvén Wave:
This is a flat-spacetime test with initial data
\begin{align}
A_x &= 0 \\
A_y &= \left \{ \begin{array}{lll}\gamma_\mu x - 0.015 & \mbox{if} & x \leq -0.1/\gamma_\mu \\
1.15 \gamma_\mu x - 0.03g(x) & \mbox{if} & -0.1/\gamma_\mu \leq x \leq 0.1/\gamma_\mu \\
1.3 \gamma_\mu x - 0.015 & \mbox{if} & x \geq 0.1/\gamma_\mu \end{array} \right. , \\
A_z = &\ y - \gamma_\mu (1-\mu)x ,
\end{align}
which generates the magnetic field in the wave frame,
\begin{align}
B'^{x'}(x') = &\ 1.0,\ B'^y(x') = 1.0, \\
B'^z(x') = &\ \left \{ \begin{array}{lll} 1.0 & \mbox{if} & x' \leq -0.1 \\
1.0+0.15 f(x') & \mbox{if} & -0.1 \leq x' \leq 0.1 \\
1.3 & \mbox{if} & x' \geq 0.1 \end{array} \right. .
\end{align}
The electric field in the wave frame is then given by
$$E'^{x'}(x') = -B'^z(0,x') \ \ , \ \ E'^y(x') = 0.0 \ \ , \ \ E'^z(x') = 1.0 .$$
These are converted to the grid frame by
\begin{align}
B^x(0,x) = &\ B'^{x'}(\gamma_\mu x) , \\
B^y(0,x) = &\ \gamma_\mu [ B'^y(\gamma_\mu x) - \mu E'^z(\gamma_\mu x) ] , \\
B^z(0,x) = &\ \gamma_\mu [ B'^z(\gamma_\mu x) + \mu E'^y(\gamma_\mu x) ] ,
\end{align}
and
\begin{align}
E^x(0,x) = &\ E'^{x'}(\gamma_\mu x) , \\
E^y(0,x) = &\ \gamma_\mu [ E'^y(\gamma_\mu x) + \mu B'^z(\gamma_\mu x) ] ,\\
E^z(0,x) = &\ \gamma_\mu [ E'^z(\gamma_\mu x) - \mu B'^y(\gamma_\mu x) ],
\end{align}
and the velocity is given by $$\mathbf{v} = \frac{\mathbf{E} \times \mathbf{B}}{B^2}$$ in flat spacetime. Additionally, $f(x)=1+\sin (5\pi x)$, $-1<\mu<1$ is the wave speed relative to the grid frame and $\gamma_\mu = (1-\mu^2)^{-1/2}$, and $g(x) = \cos (5\pi \gamma_\mu x)/\pi$.
For the eventual purpose of testing convergence, any quantity $Q$ evolves as $Q(t,x) = Q(0,x-\mu t)$
See the [Tutorial-GiRaFFEfood_NRPy](Tutorial-GiRaFFEfood_NRPy.ipynb) tutorial notebook for more general detail on how this is used.
<a id='toc'></a>
# Table of Contents:
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Import core NRPy+ modules and set NRPy+ parameters
1. [Step 2](#vector_ak): Set the vector $A_k$
1. [Step 3](#vectors_for_velocity): Set the vectors $B^i$ and $E^i$ for the velocity
1. [Step 4](#vi): Calculate $v^i$
1. [Step 5](#code_validation): Code Validation against `GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests` NRPy+ module
1. [Step 6](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Import core NRPy+ modules and set NRPy+ parameters \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
Here, we will import the NRPy+ core modules and set the reference metric to Cartesian, set commonly used NRPy+ parameters, and set C parameters that will be set from outside the code eventually generated from these expressions. We will also set up a parameter to determine what initial data is set up, although it won't do much yet.
```
# Step 0: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import os,sys
nrpy_dir_path = os.path.join("..")
if nrpy_dir_path not in sys.path:
sys.path.append(nrpy_dir_path)
# Step 0.a: Import the NRPy+ core modules and set the reference metric to Cartesian
from outputC import * # NRPy+: Core C code output module
import finite_difference as fin # NRPy+: Finite difference C code generation module
import NRPy_param_funcs as par # NRPy+: Parameter interface
import grid as gri # NRPy+: Functions having to do with numerical grids
import indexedexp as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
import reference_metric as rfm # NRPy+: Reference metric support
par.set_parval_from_str("reference_metric::CoordSystem","Cartesian")
rfm.reference_metric()
# Step 1a: Set commonly used parameters.
thismodule = "GiRaFFEfood_NRPy_1D"
# Set the spatial dimension parameter to 3.
par.set_parval_from_str("grid::DIM", 3)
DIM = par.parval_from_str("grid::DIM")
```
##### <a id='vector_ak'></a>
# Step 2: Set the vector $A_k$ \[Back to [top](#toc)\]
$$\label{vector_ak}$$
The vector potential is given as
\begin{align}
A_x &= 0 \\
A_y &= \left \{ \begin{array}{lll}\gamma_\mu x - 0.015 & \mbox{if} & x \leq -0.1/\gamma_\mu \\
1.15 \gamma_\mu x - 0.03g(x) & \mbox{if} & -0.1/\gamma_\mu \leq x \leq 0.1/\gamma_\mu \\
1.3 \gamma_\mu x - 0.015 & \mbox{if} & x \geq 0.1/\gamma_\mu \end{array} \right. , \\
A_z &= y - \gamma_\mu (1-\mu)x .
\end{align}
However, to take full advantage of NRPy+'s automated function generation capabilities, we want to write this without the `if` statements, replacing them with calls to `fabs()`. To do so, we will use rewrite these functions in terms of maxima and minima. Critically, we will define these functions in the following way:
\begin{align}
\min(a,b) &= \tfrac{1}{2} \left( a+b - \lvert a-b \rvert \right) \\
\max(a,b) &= \tfrac{1}{2} \left( a+b + \lvert a-b \rvert \right). \\
\end{align}
For real numbers, these operate exactly as expected. In the case $a>b$,
\begin{align}
\min(a,b) &= \tfrac{1}{2} \left( a+b - (a-b) \right) = b \\
\max(a,b) &= \tfrac{1}{2} \left( a+b + (a-b) \right) = a, \\
\end{align}
and in the case $a<b$, the reverse holds:
\begin{align}
\min(a,b) &= \tfrac{1}{2} \left( a+b - (b-a) \right) = a \\
\max(a,b) &= \tfrac{1}{2} \left( a+b + (b-a) \right) = b, \\
\end{align}
In code, we will represent this as:
```
min_noif(a,b) = sp.Rational(1,2)*(a+b-nrpyAbs(a-b))
max_noif(a,b) = sp.Rational(1,2)*(a+b+nrpyAbs(a-b))
```
Note the `_noif` suffix to avoid conflicts with other functions, and the use of `nrpyAbs()` function, which will always be interpreted as the C function `fabs()` (Sympy `Abs()` may get interpreted as $\sqrt{zz^*}$, for instance).
For convenience, we define two new positions, $x^+ = x+0.1/\gamma_\mu$ and $x^- = x-0.1/\gamma_\mu$. With these tools, we rewrite the $y$-component of the vector potential:
\begin{align}
A_y =& 1.15 \gamma_\mu x - 0.15 \gamma_\mu \left( \min(x^-,0)-\max(x^+,0) \right) \\
&+ g(x) \frac{\max(x^-,0) \min(x^+,0)}{\max(x^-,\epsilon) \min(x^+,\epsilon)} \\\
&- 0.015 \frac{\min(x^-,0) \max(x^+,0)}{\min(x^-,\epsilon) \max(x^+,\epsilon)}
\end{align}
First, however, we must set $$\gamma_\mu = (1-\mu^2)^{-1/2}$$ and $$g(x) = \cos (5\pi \gamma_\mu x)/\pi$$.
```
mu_AW = par.Cparameters("REAL",thismodule,["mu_AW"], -0.5) # The wave speed
M_PI = par.Cparameters("#define",thismodule,["M_PI"], "")
gammamu = 1/sp.sqrt(1-mu_AW**2)
# We'll use reference_metric.py to define x and y
x = rfm.xxCart[0]
y = rfm.xxCart[1]
g_AW = sp.cos(5*M_PI*gammamu*x)/M_PI
```
Now, we can define the vector potential. We will create three copies of this variable, because the potential is uniquely defined in three zones. Data for $x \leq -0.1/\gamma_\mu$ shall be referred to as "left", data for $-0.1/\gamma_\mu \leq x \leq 0.1/\gamma_\mu$ as "center", and data for $x \geq 0.1/\gamma_\mu$ as "right".
Starting on the left,
\begin{align}
A_x &= 0 \\
A_y &= \gamma_\mu x - 0.015 \\
A_z &= y - \gamma_\mu (1-\mu)x .
\end{align}
```
AD = ixp.register_gridfunctions_for_single_rank1("EVOL","AD")
AleftD = ixp.zerorank1()
AleftD[0] = sp.sympify(0)
AleftD[1] = gammamu*x-0.015
AleftD[2] = y-gammamu*(1-mu_AW)*x
```
In the center,
\begin{align}
A_x &= 0 \\
A_y &= 1.15 \gamma_\mu x - 0.03g(x) \\
A_z &= y - \gamma_\mu (1-\mu)x .
\end{align}
```
AcenterD = ixp.zerorank1()
AcenterD[0] = sp.sympify(0)
AcenterD[1] = 1.15*gammamu*x-0.03*g_AW
AcenterD[2] = y-gammamu*(1-mu_AW)*x
```
And on the right,
\begin{align}
A_x &= 0 \\
A_y &= 1.3 \gamma_\mu x - 0.015 \\
A_z &= y - \gamma_\mu (1-\mu)x .
\end{align}
```
ArightD = ixp.zerorank1()
ArightD[0] = sp.sympify(0)
ArightD[1] = 1.3*gammamu*x-0.015
ArightD[2] = y-gammamu*(1-mu_AW)*x
```
<a id='vectors_for_velocity'></a>
# Step 3: Set the vectors $B^i$ and $E^i$ for the velocity \[Back to [top](#toc)\]
$$\label{vectors_for_velocity}$$
Now, we will set the magnetic and electric fields that we will need to define the initial velocities. First, we need to define $$f(x)=1+\sin (5\pi x);$$ note that in the definition of $B^i$, we need $f(x')$ where $x'=\gamma_\mu x$.
```
xprime = gammamu*x
f_AW = 1.0 + sp.sin(5.0*M_PI*xprime)
print(f_AW)
```
We will now set the magnetic field in the wave frame:
\begin{align}
B'^{x'}(x') = &\ 1.0,\ B'^y(x') = 1.0, \\
B'^z(x') = &\ \left \{ \begin{array}{lll} 1.0 & \mbox{if} & x' \leq -0.1 \\
1.0+0.15 f(x') & \mbox{if} & -0.1 \leq x' \leq 0.1 \\
1.3 & \mbox{if} & x' \geq 0.1 \end{array} \right. .
\end{align}
```
BleftpU = ixp.zerorank1()
BleftpU[0] = sp.sympify(1.0)
BleftpU[1] = sp.sympify(1.0)
BleftpU[2] = sp.sympify(1.0)
BcenterpU = ixp.zerorank1()
BcenterpU[0] = sp.sympify(1.0)
BcenterpU[1] = sp.sympify(1.0)
BcenterpU[2] = 1.0 + 0.15*f_AW
BrightpU = ixp.zerorank1()
BrightpU[0] = sp.sympify(1.0)
BrightpU[1] = sp.sympify(1.0)
BrightpU[2] = sp.sympify(1.3)
```
Now, we will set the electric field in the wave frame:
\begin{align}
E'^{x'}(x') &= -B'^z(0,x'), \\
E'^y(x') &= 0.0, \\
E'^z(x') &= 1.0 .
\end{align}
```
EleftpU = ixp.zerorank1()
EleftpU[0] = -BleftpU[2]
EleftpU[1] = sp.sympify(0.0)
EleftpU[2] = sp.sympify(1.0)
EcenterpU = ixp.zerorank1()
EcenterpU[0] = -BcenterpU[2]
EcenterpU[1] = sp.sympify(0.0)
EcenterpU[2] = sp.sympify(1.0)
ErightpU = ixp.zerorank1()
ErightpU[0] = -BrightpU[2]
ErightpU[1] = sp.sympify(0.0)
ErightpU[2] = sp.sympify(1.0)
```
Next, we must transform the fields into the grid frame. We'll do the magnetic fields first.
\begin{align}
B^x(0,x) = &\ B'^{x'}(\gamma_\mu x) , \\
B^y(0,x) = &\ \gamma_\mu [ B'^y(\gamma_\mu x) - \mu E'^z(\gamma_\mu x) ] , \\
B^z(0,x) = &\ \gamma_\mu [ B'^z(\gamma_\mu x) + \mu E'^y(\gamma_\mu x) ] ,
\end{align}
```
BleftU = ixp.zerorank1()
BleftU[0] = BleftpU[0]
BleftU[1] = gammamu*(BleftpU[1]-mu_AW*EleftpU[2])
BleftU[2] = gammamu*(BleftpU[2]+mu_AW*EleftpU[1])
BcenterU = ixp.zerorank1()
BcenterU[0] = BcenterpU[0]
BcenterU[1] = gammamu*(BcenterpU[1]-mu_AW*EcenterpU[2])
BcenterU[2] = gammamu*(BcenterpU[2]+mu_AW*EcenterpU[1])
BrightU = ixp.zerorank1()
BrightU[0] = BrightpU[0]
BrightU[1] = gammamu*(BrightpU[1]-mu_AW*ErightpU[2])
BrightU[2] = gammamu*(BrightpU[2]+mu_AW*ErightpU[1])
```
And now the electric fields:
\begin{align}
E^x(0,x) = &\ E'^{x'}(\gamma_\mu x) , \\
E^y(0,x) = &\ \gamma_\mu [ E'^y(\gamma_\mu x) + \mu B'^z(\gamma_\mu x) ] ,\\
E^z(0,x) = &\ \gamma_\mu [ E'^z(\gamma_\mu x) - \mu B'^y(\gamma_\mu x) ],
\end{align}
```
EleftU = ixp.zerorank1()
EleftU[0] = EleftpU[0]
EleftU[1] = gammamu*(EleftpU[1]+mu_AW*BleftpU[2])
EleftU[2] = gammamu*(EleftpU[2]-mu_AW*BleftpU[1])
EcenterU = ixp.zerorank1()
EcenterU[0] = EcenterpU[0]
EcenterU[1] = gammamu*(EcenterpU[1]+mu_AW*BcenterpU[2])
EcenterU[2] = gammamu*(EcenterpU[2]-mu_AW*BcenterpU[1])
ErightU = ixp.zerorank1()
ErightU[0] = ErightpU[0]
ErightU[1] = gammamu*(ErightpU[1]+mu_AW*BrightpU[2])
ErightU[2] = gammamu*(ErightpU[2]-mu_AW*BrightpU[1])
```
<a id='vi'></a>
# Step 4: Calculate $v^i$ \[Back to [top](#toc)\]
$$\label{vi}$$
Now, we calculate $$\mathbf{v} = \frac{\mathbf{E} \times \mathbf{B}}{B^2},$$ which is equivalent to $$v^i = [ijk] \frac{E^j B^k}{B^2},$$ where $[ijk]$ is the Levi-Civita symbol and $B^2 = \gamma_{ij} B^i B^j$ is a trivial dot product in flat space.
```
import WeylScal4NRPy.WeylScalars_Cartesian as weyl
LeviCivitaSymbolDDD = weyl.define_LeviCivitaSymbol_rank3()
Bleft2 = BleftU[0]*BleftU[0] + BleftU[1]*BleftU[1] + BleftU[2]*BleftU[2]
Bcenter2 = BcenterU[0]*BcenterU[0] + BcenterU[1]*BcenterU[1] + BcenterU[2]*BcenterU[2]
Bright2 = BrightU[0]*BrightU[0] + BrightU[1]*BrightU[1] + BrightU[2]*BrightU[2]
ValenciavU = ixp.register_gridfunctions_for_single_rank1("AUX","ValenciavU")
ValenciavleftU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
ValenciavleftU[i] += LeviCivitaSymbolDDD[i][j][k] * EleftU[j] * BleftU[k] / Bleft2
ValenciavcenterU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
ValenciavcenterU[i] += LeviCivitaSymbolDDD[i][j][k] * EcenterU[j] * BcenterU[k] / Bcenter2
ValenciavrightU = ixp.zerorank1()
for i in range(DIM):
for j in range(DIM):
for k in range(DIM):
ValenciavrightU[i] += LeviCivitaSymbolDDD[i][j][k] * ErightU[j] * BrightU[k] / Bright2
```
<a id='code_validation'></a>
# Step 5: Code Validation against `GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests` NRPy+ module \[Back to [top](#toc)\]
$$\label{code_validation}$$
Here, as a code validation check, we verify agreement in the SymPy expressions for the `GiRaFFE` Aligned Rotator initial data equations we intend to use between
1. this tutorial and
2. the NRPy+ [`GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py`](../edit/GiRaFFEfood_NRPy/GiRaFFEfood_NRPy_1D_tests.py) module.
```
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
import GiRaFFEfood_NRPy.GiRaFFEfood_NRPy_1D_tests as gfho
gfho.GiRaFFEfood_NRPy_1D_tests()
print("Consistency check between GiRaFFEfood_NRPy tutorial and NRPy+ module: ALL SHOULD BE ZERO.")
for i in range(DIM):
print("ValenciavleftU["+str(i)+"] - gfho.ValenciavleftU["+str(i)+"] = " + str(ValenciavleftU[i] - gfho.ValenciavleftU[i]))
print("AleftD["+str(i)+"] - gfho.AleftD["+str(i)+"] = " + str(AleftD[i] - gfho.AleftD[i]))
print("ValenciavcenterU["+str(i)+"] - gfho.ValenciavcenterU["+str(i)+"] = " + str(ValenciavcenterU[i] - gfho.ValenciavcenterU[i]))
print("AcenterD["+str(i)+"] - gfho.AcenterD["+str(i)+"] = " + str(AcenterD[i] - gfho.AcenterD[i]))
print("ValenciavrightU["+str(i)+"] - gfho.ValenciavrightU["+str(i)+"] = " + str(ValenciavrightU[i] - gfho.ValenciavrightU[i]))
print("ArightD["+str(i)+"] - gfho.ArightD["+str(i)+"] = " + str(ArightD[i] - gfho.ArightD[i]))
```
<a id='latex_pdf_output'></a>
# Step 6: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-GiRaFFEfood_NRPy_1D_tests.pdf](Tutorial-GiRaFFEfood_NRPy_1D_tests.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
!jupyter nbconvert --to latex --template latex_nrpy_style.tplx --log-level='WARN' Tutorial-GiRaFFEfood_NRPy_1D_tests.ipynb
!pdflatex -interaction=batchmode Tutorial-GiRaFFEfood_NRPy_1D_tests.tex
!pdflatex -interaction=batchmode Tutorial-GiRaFFEfood_NRPy_1D_tests.tex
!pdflatex -interaction=batchmode Tutorial-GiRaFFEfood_NRPy_1D_tests.tex
!rm -f Tut*.out Tut*.aux Tut*.log
```
| github_jupyter |
```
import numpy as np
import pandas as pd
%matplotlib inline
import matplotlib.pyplot as plt
from crossing_tree import fbm, crossings, crossing_tree
from sklearn.preprocessing import StandardScaler
random_state = np.random.RandomState(0x0BADC0D)
gen_ = fbm(2**15+1, H=0.60, time=True)
# gen_ = fbm(2**22+1, H=0.95, time=True)
# gen_ = fbm(2**22+1, H=0.90, time=True)
gen_.initialize(random_state)
T, X = gen_()
print(np.std(np.diff(X)))
print(np.mean(np.abs(np.diff(X))))
print(np.median(np.abs(np.diff(X))))
print(np.percentile(np.abs(np.diff(X)), 95))
print(1.0 / np.sqrt(len(T) - 1))
# scale_ = np.diff(X).std()
# scale_ = np.median(np.abs(np.diff(X)))
scale_ = np.mean(np.abs(np.diff(X))) * 1024 # * 16384
# %%timeit -n 20
xi, ti = crossings(X, T, scale_, 0)
xi2, ti2 = crossings(X, T, scale_, 0.5)
print(xi.shape)
print(ti.shape)
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111)
ax.plot(T[:(1<<20)], X[:(1<<20)], "y", alpha=0.5)
ax.plot(ti[:11], xi[:11], "-k")
ax.plot(ti2[:11], xi2[:11], "-r")
ax.set_title("Scale %g"%(scale_,))
# scale_ *= 2
plt.show()
from scipy.stats import ks_2samp
# scale_ = np.median(np.abs(np.diff(X)))
scale_ = np.diff(X).std()
scale_ = 1.96 / np.sqrt( len(T)-1)
# scale_ = np.median(np.abs(np.diff(X)))
# scale_ = np.mean(np.abs(np.diff(X)))
for k in range(10):
xi, ti = crossings(X, T, scale_, 0)
xi2, ti2 = crossings(X, T, scale_, 0.25)
scale_ *= 2
dti_s = np.diff(ti)
dti_s /= np.sqrt(np.mean(dti_s**2))
dti2_s = np.diff(ti2)
dti2_s /= np.sqrt(np.mean(dti2_s**2))
fig = plt.figure(figsize=(12, 5))
ax = fig.add_subplot(121)
ax.hist(dti_s, bins=100, color="k", lw=0);
ax = fig.add_subplot(122)
ax.hist(dti2_s, bins=100, color="r", lw=0);
plt.show()
print(ks_2samp(dti_s, dti2_s))
```
Tree
```
# scale_ = np.median(np.abs(np.diff(X)))
scale = np.diff(X).std()/2
# scale = 1.96 / np.sqrt( len(T)-1)
# scale = np.median(np.abs(np.diff(X)))
# scale = np.mean(np.abs(np.diff(X)))
origin = 0.
xi, ti, offspring, excursions, subcrossings, durations = crossing_tree(X, T, scale, origin)
print(len(offspring))
def structure(offspring):
iter_ = iter(offspring)
try:
value_ = next(iter_)
except StopIteration:
raise TypeError('reduce() of empty sequence')
yield value_
for index in iter_:
value_ = value_[index]
yield value_
list(structure(offspring))
```
<hr/>
```
%load_ext cython
%%cython
#-a
import numpy as _np
cimport numpy as np
cimport cython
from libc.math cimport isnan, fabs, floor, ceil, NAN
np.import_array()
ctypedef fused real:
cython.floating
@cython.boundscheck(False)
@cython.wraparound(False)
@cython.cdivision(True)
@cython.initializedcheck(False)
@cython.nonecheck(False)
def integer_xing(real[:] x, real[:] t, real scale, real origin):
cdef np.intp_t n_samples = x.shape[0]
cdef np.intp_t[::1] size = _np.empty(n_samples - 1, dtype=_np.int)
cdef real[::1] first = _np.empty(n_samples - 1, dtype=_np.float)
cdef np.intp_t i
cdef real first_, last_, direction, prev_last = NAN
cdef np.intp_t total = 0, size_
with nogil:
# Detect integer-level crossings, ignoring re-crossings of the same level
for i in range(n_samples - 1):
direction, size_ = 0.0, 0
if x[i] < x[i+1]:
first_, last_ = ceil((x[i] - origin) / scale), floor((x[i+1] - origin) / scale)
direction = +1.0
elif x[i] > x[i+1]:
first_, last_ = floor((x[i] - origin) / scale), ceil((x[i+1] - origin) / scale)
direction = -1.0
if direction != 0.0:
size_ = <int>fabs(last_ + direction - first_)
if size_ > 0 and prev_last == first_:
first_ += direction
size_ -= 1
if size_ > 0:
prev_last = last_
first[i], size[i] = first_, size_
total += size_
cdef real[::1] xi = _np.empty(total, dtype=_np.float)
cdef real[::1] ti = _np.empty(total, dtype=_np.float)
cdef np.int_t k, j = 0
cdef long double x_slope_, t_slope_, first_xi_, first_ti_
with nogil:
# Interpolate the crossing times and crossing levels
for i in range(n_samples-1):
size_ = size[i]
if size_ > 0:
x_slope_ = +scale if x[i+1] > x[i] else -scale
t_slope_ = (t[i+1] - t[i]) / (x[i+1] - x[i])
first_ = first[i] * scale + origin
for k in range(size_):
xi[j] = first_ + x_slope_ * k
ti[j] = t[i] + t_slope_ * (xi[j] - x[i])
j += 1
## Marginally slower
# size_ = size[i]
# if size_ > 0:
# t_slope_ = (t[i+1] - t[i]) / (x[i+1] - x[i])
# xi[j] = first[i] * scale
# ti[j] = t[i] + t_slope_ * (xi[j] - x[i])
# j += 1
# if size_ > 1:
# x_slope_ = +scale if x[i+1] > x[i] else -scale
# for k in range(size_ - 1):
# xi[j] = xi[j-1] + x_slope_
# ti[j] = ti[j-1] + t_slope_ * x_slope_
# j += 1
return xi, ti
```
| github_jupyter |
```
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from sklearn.preprocessing import MinMaxScaler
from datetime import datetime
from datetime import timedelta
sns.set()
class Model:
def __init__(
self,
learning_rate,
num_layers,
size,
size_layer,
output_size,
forget_bias = 0.1,
):
def lstm_cell(size_layer):
return tf.nn.rnn_cell.GRUCell(size_layer)
rnn_cells = tf.nn.rnn_cell.MultiRNNCell(
[lstm_cell(size_layer) for _ in range(num_layers)],
state_is_tuple = False,
)
self.X = tf.placeholder(tf.float32, (None, None, size))
self.Y = tf.placeholder(tf.float32, (None, output_size))
drop = tf.contrib.rnn.DropoutWrapper(
rnn_cells, output_keep_prob = forget_bias
)
self.hidden_layer = tf.placeholder(
tf.float32, (None, num_layers * size_layer)
)
self.outputs, self.last_state = tf.nn.dynamic_rnn(
drop, self.X, initial_state = self.hidden_layer, dtype = tf.float32
)
rnn_W = tf.Variable(tf.random_normal((size_layer, output_size)))
rnn_B = tf.Variable(tf.random_normal([output_size]))
self.logits = tf.matmul(self.outputs[-1], rnn_W) + rnn_B
self.cost = tf.reduce_mean(tf.square(self.Y - self.logits))
self.optimizer = tf.train.AdamOptimizer(learning_rate).minimize(
self.cost
)
df = pd.read_csv('../dataset/GOOG-year.csv')
date_ori = pd.to_datetime(df.iloc[:, 0]).tolist()
df.head()
minmax = MinMaxScaler().fit(df.iloc[:, 1:].astype('float32'))
df_log = minmax.transform(df.iloc[:, 1:].astype('float32'))
df_log = pd.DataFrame(df_log)
df_log.head()
num_layers = 1
size_layer = 128
timestamp = 5
epoch = 500
dropout_rate = 0.7
future_day = 50
tf.reset_default_graph()
modelnn = Model(
0.01, num_layers, df_log.shape[1], size_layer, df_log.shape[1], dropout_rate
)
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
for i in range(epoch):
init_value = np.zeros((1, num_layers * size_layer))
total_loss = 0
for k in range(0, df_log.shape[0] - 1, timestamp):
index = min(k + timestamp, df_log.shape[0] - 1)
batch_x = np.expand_dims(df_log.iloc[k:index, :].values, axis = 0)
batch_y = df_log.iloc[k + 1 : index + 1, :].values
last_state, _, loss = sess.run(
[modelnn.last_state, modelnn.optimizer, modelnn.cost],
feed_dict = {
modelnn.X: batch_x,
modelnn.Y: batch_y,
modelnn.hidden_layer: init_value,
},
)
loss = np.mean(loss)
init_value = last_state
total_loss += loss
total_loss /= df_log.shape[0] // timestamp
if (i + 1) % 100 == 0:
print('epoch:', i + 1, 'avg loss:', total_loss)
output_predict = np.zeros((df_log.shape[0] + future_day, df_log.shape[1]))
output_predict[0, :] = df_log.iloc[0, :]
upper_b = (df_log.shape[0] // timestamp) * timestamp
init_value = np.zeros((1, num_layers * size_layer))
for k in range(0, (df_log.shape[0] // timestamp) * timestamp, timestamp):
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(
df_log.iloc[k : k + timestamp, :], axis = 0
),
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
output_predict[k + 1 : k + timestamp + 1, :] = out_logits
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(df_log.iloc[upper_b:, :], axis = 0),
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
output_predict[upper_b + 1 : df_log.shape[0] + 1, :] = out_logits
df_log.loc[df_log.shape[0]] = out_logits[-1, :]
date_ori.append(date_ori[-1] + timedelta(days = 1))
for i in range(future_day - 1):
out_logits, last_state = sess.run(
[modelnn.logits, modelnn.last_state],
feed_dict = {
modelnn.X: np.expand_dims(df_log.iloc[-timestamp:, :], axis = 0),
modelnn.hidden_layer: init_value,
},
)
init_value = last_state
output_predict[df_log.shape[0], :] = out_logits[-1, :]
df_log.loc[df_log.shape[0]] = out_logits[-1, :]
date_ori.append(date_ori[-1] + timedelta(days = 1))
df_log = minmax.inverse_transform(output_predict)
date_ori = pd.Series(date_ori).dt.strftime(date_format = '%Y-%m-%d').tolist()
def anchor(signal, weight):
buffer = []
last = signal[0]
for i in signal:
smoothed_val = last * weight + (1 - weight) * i
buffer.append(smoothed_val)
last = smoothed_val
return buffer
current_palette = sns.color_palette('Paired', 12)
fig = plt.figure(figsize = (15, 10))
ax = plt.subplot(111)
x_range_original = np.arange(df.shape[0])
x_range_future = np.arange(df_log.shape[0])
ax.plot(
x_range_original,
df.iloc[:, 1],
label = 'true Open',
color = current_palette[0],
)
ax.plot(
x_range_future,
anchor(df_log[:, 0], 0.5),
label = 'predict Open',
color = current_palette[1],
)
ax.plot(
x_range_original,
df.iloc[:, 2],
label = 'true High',
color = current_palette[2],
)
ax.plot(
x_range_future,
anchor(df_log[:, 1], 0.5),
label = 'predict High',
color = current_palette[3],
)
ax.plot(
x_range_original,
df.iloc[:, 3],
label = 'true Low',
color = current_palette[4],
)
ax.plot(
x_range_future,
anchor(df_log[:, 2], 0.5),
label = 'predict Low',
color = current_palette[5],
)
ax.plot(
x_range_original,
df.iloc[:, 4],
label = 'true Close',
color = current_palette[6],
)
ax.plot(
x_range_future,
anchor(df_log[:, 3], 0.5),
label = 'predict Close',
color = current_palette[7],
)
ax.plot(
x_range_original,
df.iloc[:, 5],
label = 'true Adj Close',
color = current_palette[8],
)
ax.plot(
x_range_future,
anchor(df_log[:, 4], 0.5),
label = 'predict Adj Close',
color = current_palette[9],
)
box = ax.get_position()
ax.set_position(
[box.x0, box.y0 + box.height * 0.1, box.width, box.height * 0.9]
)
ax.legend(
loc = 'upper center',
bbox_to_anchor = (0.5, -0.05),
fancybox = True,
shadow = True,
ncol = 5,
)
plt.title('overlap stock market')
plt.xticks(x_range_future[::30], date_ori[::30])
plt.show()
fig = plt.figure(figsize = (20, 8))
plt.subplot(1, 2, 1)
plt.plot(
x_range_original,
df.iloc[:, 1],
label = 'true Open',
color = current_palette[0],
)
plt.plot(
x_range_original,
df.iloc[:, 2],
label = 'true High',
color = current_palette[2],
)
plt.plot(
x_range_original,
df.iloc[:, 3],
label = 'true Low',
color = current_palette[4],
)
plt.plot(
x_range_original,
df.iloc[:, 4],
label = 'true Close',
color = current_palette[6],
)
plt.plot(
x_range_original,
df.iloc[:, 5],
label = 'true Adj Close',
color = current_palette[8],
)
plt.xticks(x_range_original[::60], df.iloc[:, 0].tolist()[::60])
plt.legend()
plt.title('true market')
plt.subplot(1, 2, 2)
plt.plot(
x_range_future,
anchor(df_log[:, 0], 0.5),
label = 'predict Open',
color = current_palette[1],
)
plt.plot(
x_range_future,
anchor(df_log[:, 1], 0.5),
label = 'predict High',
color = current_palette[3],
)
plt.plot(
x_range_future,
anchor(df_log[:, 2], 0.5),
label = 'predict Low',
color = current_palette[5],
)
plt.plot(
x_range_future,
anchor(df_log[:, 3], 0.5),
label = 'predict Close',
color = current_palette[7],
)
plt.plot(
x_range_future,
anchor(df_log[:, 4], 0.5),
label = 'predict Adj Close',
color = current_palette[9],
)
plt.xticks(x_range_future[::60], date_ori[::60])
plt.legend()
plt.title('predict market')
plt.show()
fig = plt.figure(figsize = (15, 10))
ax = plt.subplot(111)
ax.plot(x_range_original, df.iloc[:, -1], label = 'true Volume')
ax.plot(x_range_future, anchor(df_log[:, -1], 0.5), label = 'predict Volume')
box = ax.get_position()
ax.set_position(
[box.x0, box.y0 + box.height * 0.1, box.width, box.height * 0.9]
)
ax.legend(
loc = 'upper center',
bbox_to_anchor = (0.5, -0.05),
fancybox = True,
shadow = True,
ncol = 5,
)
plt.xticks(x_range_future[::30], date_ori[::30])
plt.title('overlap market volume')
plt.show()
fig = plt.figure(figsize = (20, 8))
plt.subplot(1, 2, 1)
plt.plot(x_range_original, df.iloc[:, -1], label = 'true Volume')
plt.xticks(x_range_original[::60], df.iloc[:, 0].tolist()[::60])
plt.legend()
plt.title('true market volume')
plt.subplot(1, 2, 2)
plt.plot(x_range_future, anchor(df_log[:, -1], 0.5), label = 'predict Volume')
plt.xticks(x_range_future[::60], date_ori[::60])
plt.legend()
plt.title('predict market volume')
plt.show()
```
| github_jupyter |
<div class="devsite-table-wrapper"><table class="tfo-notebook-buttons" align="left">
<td><a target="_blank" href="https://www.tensorflow.org/tfx/tutorials/transform/simple">
<img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a></td>
<td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/tfx/blob/master/docs/tutorials/transform/simple.ipynb">
<img src="https://www.tensorflow.org/images/colab_logo_32px.png">Run in Google Colab</a></td>
<td><a target="_blank" href="https://github.com/tensorflow/tfx/blob/master/docs/tutorials/transform/simple.ipynb">
<img width=32px src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">View source on GitHub</a></td>
</table></div>
##### Copyright © 2019 Google Inc.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Preprocess data with TensorFlow Transform
***The Feature Engineering Component of TensorFlow Extended (TFX)***
This example colab notebook provides a very simple example of how <a target='_blank' href='https://www.tensorflow.org/tfx/transform/'>TensorFlow Transform (<code>tf.Transform</code>)</a> can be used to preprocess data using exactly the same code for both training a model and serving inferences in production.
TensorFlow Transform is a library for preprocessing input data for TensorFlow, including creating features that require a full pass over the training dataset. For example, using TensorFlow Transform you could:
* Normalize an input value by using the mean and standard deviation
* Convert strings to integers by generating a vocabulary over all of the input values
* Convert floats to integers by assigning them to buckets, based on the observed data distribution
TensorFlow has built-in support for manipulations on a single example or a batch of examples. `tf.Transform` extends these capabilities to support full passes over the entire training dataset.
The output of `tf.Transform` is exported as a TensorFlow graph which you can use for both training and serving. Using the same graph for both training and serving can prevent skew, since the same transformations are applied in both stages.
## Python check and imports
First, we'll make sure that we're using Python 2. Then, we'll go ahead and install and import the stuff we need.
```
from __future__ import print_function
import sys, os
# Confirm that we're using Python 2
assert sys.version_info.major is 2, 'Oops, not running Python 2'
try:
import tensorflow_transform as tft
except ImportError:
# this will take a minute, ignore the warnings
!pip install -q tensorflow-transform
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import pprint
import tempfile
import tensorflow as tf
import tensorflow_transform as tft
import tensorflow_transform.beam.impl as tft_beam
from tensorflow_transform.tf_metadata import dataset_metadata
from tensorflow_transform.tf_metadata import dataset_schema
tf.logging.set_verbosity(tf.logging.ERROR)
```
## Data: Create some dummy data
We'll create some simple dummy data for our simple example:
* `raw_data` is the initial raw data that we're going to preprocess
* `raw_data_metadata` contains the schema that tells us the types of each of the columns in `raw_data`. In this case, it's very simple.
```
raw_data = [
{'x': 1, 'y': 1, 's': 'hello'},
{'x': 2, 'y': 2, 's': 'world'},
{'x': 3, 'y': 3, 's': 'hello'}
]
raw_data_metadata = dataset_metadata.DatasetMetadata(
dataset_schema.from_feature_spec({
'y': tf.FixedLenFeature([], tf.float32),
'x': tf.FixedLenFeature([], tf.float32),
's': tf.FixedLenFeature([], tf.string),
}))
```
## Transform: Create a preprocessing function
The _preprocessing function_ is the most important concept of tf.Transform. A preprocessing function is where the transformation of the dataset really happens. It accepts and returns a dictionary of tensors, where a tensor means a <a target='_blank' href='https://www.tensorflow.org/api_docs/python/tf/Tensor'><code>Tensor</code></a> or <a target='_blank' href='https://www.tensorflow.org/api_docs/python/tf/SparseTensor'><code>SparseTensor</code></a>. There are two main groups of API calls that typically form the heart of a preprocessing function:
1. **TensorFlow Ops:** Any function that accepts and returns tensors, which usually means TensorFlow ops. These add TensorFlow operations to the graph that transforms raw data into transformed data one feature vector at a time. These will run for every example, during both training and serving.
2. **TensorFlow Transform Analyzers:** Any of the analyzers provided by tf.Transform. Analyzers also accept and return tensors, but unlike TensorFlow ops they only run once, during training, and typically make a full pass over the entire training dataset. They create <a target='_blank' href='https://www.tensorflow.org/api_docs/python/tf/constant'>tensor constants</a>, which are added to your graph. For example, `tft.min` computes the minimum of a tensor over the training dataset. tf.Transform provides a fixed set of analyzers, but this will be extended in future versions.
Caution: When you apply your preprocessing function to serving inferences, the constants that were created by analyzers during training do not change. If your data has trend or seasonality components, plan accordingly.
```
def preprocessing_fn(inputs):
"""Preprocess input columns into transformed columns."""
x = inputs['x']
y = inputs['y']
s = inputs['s']
x_centered = x - tft.mean(x)
y_normalized = tft.scale_to_0_1(y)
s_integerized = tft.compute_and_apply_vocabulary(s)
x_centered_times_y_normalized = (x_centered * y_normalized)
return {
'x_centered': x_centered,
'y_normalized': y_normalized,
's_integerized': s_integerized,
'x_centered_times_y_normalized': x_centered_times_y_normalized,
}
```
## Putting it all together
Now we're ready to transform our data. We'll use Apache Beam with a direct runner, and supply three inputs:
1. `raw_data` - The raw input data that we created above
2. `raw_data_metadata` - The schema for the raw data
3. `preprocessing_fn` - The function that we created to do our transformation
<aside class="key-term"><b>Key Term:</b> <a target='_blank' href='https://beam.apache.org/'>Apache Beam</a> uses a <a target='_blank' href='https://beam.apache.org/documentation/programming-guide/#applying-transforms'>special syntax to define and invoke transforms</a>. For example, in this line:
<blockquote><code>result = pass_this | 'name this step' >> to_this_call</code></blockquote>
The method <code>to_this_call</code> is being invoked and passed the object called <code>pass_this</code>, and <a target='_blank' href='https://stackoverflow.com/questions/50519662/what-does-the-redirection-mean-in-apache-beam-python'>this operation will be referred to as <code>name this step</code> in a stack trace</a>. The result of the call to <code>to_this_call</code> is returned in <code>result</code>. You will often see stages of a pipeline chained together like this:
<blockquote><code>result = apache_beam.Pipeline() | 'first step' >> do_this_first() | 'second step' >> do_this_last()</code></blockquote>
and since that started with a new pipeline, you can continue like this:
<blockquote><code>next_result = result | 'doing more stuff' >> another_function()</code></blockquote></aside>
```
def main():
# Ignore the warnings
with tft_beam.Context(temp_dir=tempfile.mkdtemp()):
transformed_dataset, transform_fn = ( # pylint: disable=unused-variable
(raw_data, raw_data_metadata) | tft_beam.AnalyzeAndTransformDataset(
preprocessing_fn))
transformed_data, transformed_metadata = transformed_dataset # pylint: disable=unused-variable
print('\nRaw data:\n{}\n'.format(pprint.pformat(raw_data)))
print('Transformed data:\n{}'.format(pprint.pformat(transformed_data)))
if __name__ == '__main__':
main()
```
## Is this the right answer?
Previously, we used `tf.Transform` to do this:
```
x_centered = x - tft.mean(x)
y_normalized = tft.scale_to_0_1(y)
s_integerized = tft.compute_and_apply_vocabulary(s)
x_centered_times_y_normalized = (x_centered * y_normalized)
```
#### x_centered
With input of `[1, 2, 3]` the mean of x is 2, and we subtract it from x to center our x values at 0. So our result of `[-1.0, 0.0, 1.0]` is correct.
#### y_normalized
We wanted to scale our y values between 0 and 1. Our input was `[1, 2, 3]` so our result of `[0.0, 0.5, 1.0]` is correct.
#### s_integerized
We wanted to map our strings to indexes in a vocabulary, and there were only 2 words in our vocabulary ("hello" and "world"). So with input of `["hello", "world", "hello"]` our result of `[0, 1, 0]` is correct.
#### x_centered_times_y_normalized
We wanted to create a new feature by crossing `x_centered` and `y_normalized` using multiplication. Note that this multiplies the results, not the original values, and our new result of `[-0.0, 0.0, 1.0]` is correct.
| github_jupyter |
The model trained in this notebook is the best performing model I was able to get (on the task of MovieLens rating prediction) after a bit of experimentation with hyperparameters. It uses:
- matrix factorization architecture
- embedding size = 32
- embedding L2 penalty
- dropout (applied to embedding vectors)
It's used for lessons 3 and 4 of the embeddings course, which are on exploring learned embeddings with Gensim and t-SNE, respectively. (On the assumption that the model with the best error will probably have the 'best' embeddings, in terms of identifying interesting/useful latent properties.
TODO: I imagine there could still be some further gains with a little more experimentation.
- Training and val errors are so close, I wonder about cutting dropout prob in half or something
- And/or doubling embedding size
- Adding biases back in?
```
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
import tensorflow as tf
from tensorflow import keras
import os
import random
RUNNING_ON_KERNELS = 'KAGGLE_WORKING_DIR' in os.environ
input_dir = '../input' if RUNNING_ON_KERNELS else '../input/movielens_preprocessed'
ratings_path = os.path.join(input_dir, 'rating.csv')
df = pd.read_csv(ratings_path, usecols=['userId', 'movieId', 'rating', 'y'])
tf.set_random_seed(1); np.random.seed(1); random.seed(1)
movie_embedding_size = user_embedding_size = 32
user_id_input = keras.Input(shape=(1,), name='user_id')
movie_id_input = keras.Input(shape=(1,), name='movie_id')
movie_r12n = keras.regularizers.l1_l2(l1=0, l2=1e-6)
user_r12n = keras.regularizers.l1_l2(l1=0, l2=1e-7)
dropout = .2
# Had good results with 'glorot_uniform' embeddings initializer, but this seems to cause some issues
# with model deserialization
user_embedded = keras.layers.Embedding(df.userId.max()+1, user_embedding_size,
embeddings_regularizer=user_r12n,
input_length=1, name='user_embedding')(user_id_input)
user_embedded = keras.layers.Dropout(dropout)(user_embedded)
movie_embedded = keras.layers.Embedding(df.movieId.max()+1, movie_embedding_size,
embeddings_regularizer=movie_r12n,
input_length=1, name='movie_embedding')(movie_id_input)
movie_embedded = keras.layers.Dropout(dropout)(movie_embedded)
dotted = keras.layers.Dot(2)([user_embedded, movie_embedded])
out = keras.layers.Flatten()(dotted)
biases = 0
if biases:
bias_r12n = None
bias_r12n = keras.regularizers.l1_l2(l1=1e-4, l2=1e-7) # XXX 1e-6 -> 1e-4
bias_init = 'zeros'
movie_b = keras.layers.Embedding(df.movieId.max()+1, 1,
name='movie_bias',
embeddings_initializer=bias_init,
embeddings_regularizer=bias_r12n,
)(movie_id_input)
movie_b = keras.layers.Flatten()(movie_b)
user_b = keras.layers.Embedding(df.userId.max()+1, 1,
name='user_bias',
embeddings_initializer=bias_init,
embeddings_regularizer=bias_r12n,
)(user_id_input)
user_b = keras.layers.Flatten()(user_b)
out = keras.layers.Add()([user_b, movie_b, out])
model = keras.Model(
inputs = [user_id_input, movie_id_input],
outputs = out,
)
model.compile(
tf.train.AdamOptimizer(0.001),
loss='MSE',
metrics=['MAE'],
)
tf.set_random_seed(1); np.random.seed(1); random.seed(1)
history = model.fit(
[df.userId, df.movieId],
df.y,
batch_size=10**4,
epochs=30,
verbose=2,
validation_split=.05,
);
model.save('movie_svd_model_32.h5')
```
| github_jupyter |
```
#Load in utility classes
%run ~/projects/testlipids/testlipids.ipynb
import os #Operating system specific commands
import re #Regular expression library
membrane="epithelial"
insane="./insane+SF.py"
mdparams="./test.mdp"
martinipath="./martini.ff"
# TODO: 3000 lipids
# TODO: neuronal membrane
# TODO: plasma membrane
# Cleaning up intermediate files from previous runs
!rm -f *#*
!rm -f *step*
!rm -f {membrane}.gro
!rm -f {membrane}.edr
!rm -f {membrane}.log
!rm -f {membrane}.pdb
!rm -f {membrane}.tpr
!rm -f {membrane}.trr
cmdline = "-u PBSM:0.007 -u PXSM:0.004 -u DPSM:0.017 -u PNSM:0.008 -u DPMC:0.007 -u POMC:0.043 -u DOMC:0.009 -u PIMC:0.009 -u OIMC:0.004 -u OEMC:0.005 -u PAMC:0.012 -u PUMC:0.029 -u POME:0.001 -u DOME:0.006 -u OIME:0.001 -u OQME:0.002 -u OAME:0.014 -u OUME:0.006 -u IAME:0.007 -u IQME:0.001 -u LPPC:0.008 -u DPPC:0.003 -u POPC:0.147 -u DOPC:0.011 -u PIPC:0.088 -u OIPC:0.05 -u PAPC:0.052 -u PUPC:0.05 -u POPE:0.007 -u OPPE:0.002 -u DOPE:0.007 -u PAPE:0.006 -u PUPE:0.002 -u OIPE:0.002 -u PQPE:0.002 -u OQPE:0.001 -u OAPE:0.002 -u OUPE:0.001 -u POPG:0.004 -u DOPG:0.003 -u DODG:0.004 -u PODG:0.007 -l OGPS:0.004 -l DPPS:0.001 -l POPS:0.097 -l DOPS:0.006 -l PAPI:0.029 -l PAP1:0.014 -l PAP2:0.014 -l PQPI:0.02 -l POPI:0.013 -l PUPI:0.008 -l DOPI:0.008 -l PIPI:0.006 -l PEPI:0.004 -l PBSM:0.003 -l PXSM:0.002 -l DPSM:0.007 -l PNSM:0.003 -l DPMC:0.003 -l POMC:0.017 -l DOMC:0.004 -l PIMC:0.003 -l OIMC:0.002 -l OEMC:0.002 -l PAMC:0.005 -l PUMC:0.012 -l POME:0.005 -l DOME:0.021 -l OIME:0.004 -l OQME:0.006 -l OAME:0.054 -l OUME:0.024 -l IAME:0.025 -l IQME:0.003 -l LPPC:0.003 -l DPPC:0.001 -l POPC:0.058 -l DOPC:0.004 -l PIPC:0.035 -l OIPC:0.02 -l PAPC:0.021 -l PUPC:0.02 -l POPA:0.01 -l DOPA:0.005 -l POPE:0.025 -l OPPE:0.009 -l DOPE:0.027 -l PAPE:0.021 -l PUPE:0.008 -l OIPE:0.008 -l PQPE:0.007 -l OQPE:0.004 -l OAPE:0.009 -l OUPE:0.005 -l POPG:0.004 -l DOPG:0.002 -l DODG:0.002 -l PODG:0.004 "
print("Build")
build = !python2 {insane} -o {membrane}.gro -p {membrane}.top -d 0 -x 35 -y 35 -z 15 -sol PW -salt 0.15 -center -charge 0 -orient {cmdline}
for line in build:
print(line)
# remove extra descriptive lines in the gro file that would otherwise overrun the buffer during GROMPP
import datetime
generationDate = datetime.datetime.now().strftime("%Y.%m.%d")
!sed -i '1s/.*/Complex Epithelial membrane generated ({generationDate}) using INSANE/' {membrane}.gro
# remove extra descriptive lines in the top file that would otherwise overrun the buffer during GROMPP
import datetime
generationDate = datetime.datetime.now().strftime("%Y.%m.%d")
!sed -i '13s/.*/Complex Epithelial membrane generated ({generationDate}) using INSANE/' {membrane}.top
print("Grompp")
grompp = !gmx grompp -f {mdparams} -c {membrane}.gro -p {membrane}.top -o {membrane}.tpr
success=True
for line in grompp:
if re.search("Fatal error", line):
success=False
#if not success:
print(line)
if success:
print("Run")
!export GMX_MAXCONSTRWARN=-1
!export GMX_SUPPRESS_DUMP=1
run = !gmx mdrun -v -deffnm {membrane}
summary=""
logfile = membrane+".log"
if not os.path.exists(logfile):
print("no log file")
print("== === ====")
for line in run:
print(line)
else:
try:
file = open(logfile, "r")
fe = False
for line in file:
if fe:
success=False
summary=line
elif re.search("^Steepest Descents.*converge", line):
success=True
summary=line
break
elif re.search("Fatal error", line):
fe = True
except IOError as exc:
sucess=False;
summary=exc;
if success:
print("Success")
else:
print(summary)
# test lipids
#Load in utility classes
%run ./testlipids.ipynb
#load lipids from GRO file
lipids=lipidsfromsystem("./epithelial")
print(lipids)
testlipids(lipids,membrane,insane,mdparams,martinipath).execute(True).report()
# test lipids
#Load in utility classes
%run ./testlipids.ipynb
# test an individual lipis species
import os #Operating system specific commands
#re-compile
lipid = 'DPMC'
results=[]
if build(os.getcwd(),membrane,lipid,insane).execute().report(results).success:
comp = compile(os.getcwd(),membrane,lipid,mdparams,martinipath).execute().report(results)
print(lipid)
if not comp.success:
for line in comp.output:
print(line)
else:
for line in results:
print(line)
print("done")
```
| github_jupyter |
## Classification example for Landsat 8 imagery
**Author:** René Kopeinig<br>
**Description:** Classification example for Landsat 8 imagery based on the scientfic work "MAD-MEX: Automatic Wall-to-Wall Land Cover Monitoring for the Mexican REDD-MRV Program Using All Landsat Data" by S.Gebhardt et. al 2014. Please find the link to the paper here: https://www.mdpi.com/2072-4292/6/5/3923
```
from IPython.display import Image
import ee, folium
ee.Initialize()
%matplotlib inline
```
### Get and visualize the Landsat 8 input data
```
area_of_interest = ee.Geometry.Rectangle([-98.75, 19.15, -98.15,18.75])
mexico_landcover_2010_landsat = ee.Image("users/renekope/MEX_LC_2010_Landsat_v43").clip(area_of_interest)
landsat8_collection = ee.ImageCollection('LANDSAT/LC8_L1T_TOA').filterDate('2016-01-01', '2018-04-19').min()
landsat8_collection = landsat8_collection.slice(0,9)
vis = {
'bands': ['B6', 'B5', 'B2'],
'min': 0,
'max': 0.5,
'gamma': [0.95, 1.1, 1],
'region':area_of_interest}
image = landsat8_collection.clip(area_of_interest)
mapid = image.getMapId(vis)
map = folium.Map(location=[19.15,-98.75],zoom_start=9, height=500,width=700)
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='Landsat 8 ',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
### Functions to derive vegetation indices and other raster operations
```
def NDVI(image):
return image.normalizedDifference(['B5', 'B4'])
def SAM(image):
band1 = image.select("B1")
bandn = image.select("B2","B3","B4","B5","B6","B7","B8","B9");
maxObjSize = 256;
b = band1.divide(bandn);
spectralAngleMap = b.atan();
spectralAngleMap_sin = spectralAngleMap.sin();
spectralAngleMap_cos = spectralAngleMap.cos();
sum_cos = spectralAngleMap_cos.reduce(ee.call("Reducer.sum"));
sum_sin = spectralAngleMap_sin.reduce(ee.call("Reducer.sum"));
return ee.Image.cat(sum_sin, sum_cos, spectralAngleMap_sin, spectralAngleMap_cos);
#Enhanced Vegetation Index
def EVI(image):
# L(Canopy background)
# C1,C2(Coefficients of aerosol resistance term)
# GainFactor(Gain or scaling factor)
gain_factor = ee.Image(2.5);
coefficient_1 = ee.Image(6);
coefficient_2 = ee.Image(7.5);
l = ee.Image(1);
nir = image.select("B5");
red = image.select("B4");
blue = image.select("B2");
evi = image.expression(
"Gain_Factor*((NIR-RED)/(NIR+C1*RED-C2*BLUE+L))",
{
"Gain_Factor":gain_factor,
"NIR":nir,
"RED":red,
"C1":coefficient_1,
"C2":coefficient_2,
"BLUE":blue,
"L":l
}
)
return evi
#Atmospherically Resistant Vegetation Index
def ARVI(image):
red = image.select("B4")
blue = image.select("B2")
nir = image.select("B5")
red_square = red.multiply(red)
arvi = image.expression(
"NIR - (REDsq - BLUE)/(NIR+(REDsq-BLUE))",{
"NIR": nir,
"REDsq": red_square,
"BLUE": blue
}
)
return arvi
#Leaf Area Index
def LAI(image):
nir = image.select("B5")
red = image.select("B4")
coeff1 = ee.Image(0.0305);
coeff2 = ee.Image(1.2640);
lai = image.expression(
"(((NIR/RED)*COEFF1)+COEFF2)",
{
"NIR":nir,
"RED":red,
"COEFF1":coeff1,
"COEFF2":coeff2
}
)
return lai
def tasseled_cap_transformation(image):
#Tasseled Cap Transformation for Landsat 8 based on the
#scientfic work "Derivation of a tasselled cap transformation based on Landsat 8 at-satellite reflectance"
#by M.Baigab, L.Zhang, T.Shuai & Q.Tong (2014). The bands of the output image are the brightness index,
#greenness index and wetness index.
b = image.select("B2", "B3", "B4", "B5", "B6", "B7");
#Coefficients are only for Landsat 8 TOA
brightness_coefficents= ee.Image([0.3029, 0.2786, 0.4733, 0.5599, 0.508, 0.1872])
greenness_coefficents= ee.Image([-0.2941, -0.243, -0.5424, 0.7276, 0.0713, -0.1608]);
wetness_coefficents= ee.Image([0.1511, 0.1973, 0.3283, 0.3407, -0.7117, -0.4559]);
fourth_coefficents= ee.Image([-0.8239, 0.0849, 0.4396, -0.058, 0.2013, -0.2773]);
fifth_coefficents= ee.Image([-0.3294, 0.0557, 0.1056, 0.1855, -0.4349, 0.8085]);
sixth_coefficents= ee.Image([0.1079, -0.9023, 0.4119, 0.0575, -0.0259, 0.0252]);
#Calculate tasseled cap transformation
brightness = image.expression(
'(B * BRIGHTNESS)',
{
'B':b,
'BRIGHTNESS': brightness_coefficents
})
greenness = image.expression(
'(B * GREENNESS)',
{
'B':b,
'GREENNESS': greenness_coefficents
})
wetness = image.expression(
'(B * WETNESS)',
{
'B':b,
'WETNESS': wetness_coefficents
})
fourth = image.expression(
'(B * FOURTH)',
{
'B':b,
'FOURTH': fourth_coefficents
})
fifth = image.expression(
'(B * FIFTH)',
{
'B':b,
'FIFTH': fifth_coefficents
})
sixth = image.expression(
'(B * SIXTH)',
{
'B':b,
'SIXTH': sixth_coefficents
})
bright = brightness.reduce(ee.call("Reducer.sum"));
green = greenness.reduce(ee.call("Reducer.sum"));
wet = wetness.reduce(ee.call("Reducer.sum"));
four = fourth.reduce(ee.call("Reducer.sum"));
five = fifth.reduce(ee.call("Reducer.sum"));
six = sixth.reduce(ee.call("Reducer.sum"));
tasseled_cap = ee.Image(bright).addBands(green).addBands(wet).addBands(four).addBands(five).addBands(six)
return tasseled_cap.rename('brightness','greenness','wetness','fourth','fifth','sixth')
```
### Derive and visualize Tasseled Cap Transformation
```
tct = tasseled_cap_transformation(landsat8_collection)
image = tct.clip(area_of_interest)
vis_tct = {'min':-1,'max':2,'size':'800',
'bands':['brightness','greenness','wetness'],
'region':area_of_interest}
mapid = image.getMapId(vis_tct)
map = folium.Map(location=[19.15,-98.75],zoom_start=9, height=500,width=700)
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='Tasseled Cap Transformation',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
### Derive indices, spectral angles. Build and visualize image stack
```
ndvi = NDVI(landsat8_collection)
sam = SAM(landsat8_collection)
evi = EVI(landsat8_collection)
arvi = ARVI(landsat8_collection)
lai = LAI(landsat8_collection)
spectral_indices_stack = ee.Image(ndvi).addBands(lai).addBands(sam).addBands(arvi).addBands(evi).addBands(tct).addBands(landsat8_collection)
image = ndvi.clip(area_of_interest)
vis_ndvi = {'min':-1,'max':1,'size':'800',
'region':area_of_interest}
mapid = image.getMapId(vis_ndvi)
map = folium.Map(location=[19.15,-98.75],zoom_start=9, height=500,width=700)
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='NDVI',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
### Define classification function
```
def classification(raster_input, training_dataset,number_of_training_points, region, classification_algorithm):
bands = raster_input.bandNames()
points = ee.FeatureCollection.randomPoints(region, number_of_training_points, number_of_training_points, 1)
training = training_dataset.addBands(raster_input).reduceToVectors(
reducer='mean',
geometry=points,
geometryType='centroid',
scale=30,
crs='EPSG:4326'
)
classifier = ee.Classifier.randomForest().train(
features=training,
classProperty='label',
inputProperties=raster_input.bandNames(),
)
out = raster_input.classify(classifier)
return out
```
### Derive classification function
```
output = classification(spectral_indices_stack, mexico_landcover_2010_landsat, 10000, area_of_interest, 'Cart')
palette = ['5d9cd4','007e00','003c00','aaaa00','aa8000','8baa00','ffb265','00d900','aa007f','ff55ff','ff557f','ff007f','ff55ff','aaffff','00ffff','55aaff','e29700','bd7e00','966400','a2ecb1','c46200','aa5500','6d3600','00aa7f','008a65','005941','e9e9af','faff98',
'00007f','c7c8bc','4d1009','000000','fef7ff','6daa50','3a7500','0b5923','ffaaff','ffd1fa']
palette = ','.join(palette)
# make a visualizing variable
vis_classification = {'min': 0, 'max': len(palette), 'palette': palette, 'region':area_of_interest}
```
### Display training data of classification
```
image = mexico_landcover_2010_landsat.clip(area_of_interest)
mapid = image.getMapId(vis_classification)
map = folium.Map(location=[19.15,-98.75],zoom_start=9, height=500,width=700)
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='Training Data',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
### Display classification output
Please be patient. It may take a few moments. You might have to run this cell several times.
```
image = output.clip(area_of_interest)
mapid = image.getMapId(vis_classification)
map = folium.Map(location=[19.15,-98.75],zoom_start=9, height=500,width=700)
folium.TileLayer(
tiles=mapid['tile_fetcher'].url_format,
attr='Map Data © <a href="https://earthengine.google.com/">Google Earth Engine</a>',
overlay=True,
name='Classification Output',
).add_to(map)
map.add_child(folium.LayerControl())
map
```
| github_jupyter |
```
import math
import numpy as np
import pandas as pd
import scikitplot
import seaborn as sns
from matplotlib import pyplot
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import classification_report
import tensorflow as tf
from tensorflow.keras import optimizers
from tensorflow.keras.datasets import mnist
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Flatten, Dense, Conv2D, MaxPooling2D
from tensorflow.keras.layers import Dropout, BatchNormalization, LeakyReLU, Activation
from tensorflow.keras.callbacks import Callback, EarlyStopping, ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from keras.utils import np_utils
# Adicionando a path das pastas
train_dir = "./images/train" #passing the path with training images
test_dir = "./images/validation" #passing the path with testing images
#original size of the image
img_size = 48
# processando o pre training set
train_datagen = ImageDataGenerator(rescale = 1./255,
shear_range = 0.2,
zoom_range = 0.2,
horizontal_flip = True)
# processando o pre testing set
validation_datagen = ImageDataGenerator(rescale = 1./255,
validation_split = 0.2)
# depois tem que alterar pra binary pra ver qual performs better
train_set = train_datagen.flow_from_directory(directory = train_dir,
target_size = (img_size,img_size),
batch_size = 32,
color_mode = "grayscale",
class_mode = "categorical",
subset = "training"
)
validation_set = validation_datagen.flow_from_directory( directory = test_dir,
target_size = (img_size,img_size),
batch_size = 32,
color_mode = "grayscale",
class_mode = "categorical",
subset = "validation"
)
# num_classes usado na hora do dense
num_classes = 7
# Inicializing the cnn
cnn = Sequential()
# First Layer
cnn.add(
Conv2D(
filters = 64,
kernel_size=(5,5),
input_shape=(img_size, img_size, 1),
activation = 'elu',
name = 'Conv2D_1'
)
)
cnn.add(BatchNormalization(name='batchnorm_1'))
# Second Layer
cnn.add(
Conv2D(
filters=64,
kernel_size=(5,5),
activation='elu',
name='conv2d_2'
)
)
cnn.add(BatchNormalization(name='batchnorm_2'))
cnn.add(MaxPooling2D(pool_size=(2,2), name='maxpool2d_1'))
cnn.add(Dropout(0.4, name='dropout_1'))
# Third Layer
cnn.add(
Conv2D(
filters=128,
kernel_size=(3,3),
activation='elu',
name='conv2d_3'
)
)
cnn.add(BatchNormalization(name='batchnorm_3'))
# Fouth Layer
cnn.add(
Conv2D(
filters=128,
kernel_size=(3,3),
activation='elu',
name='conv2d_4'
)
)
cnn.add(BatchNormalization(name='batchnorm_4'))
cnn.add(MaxPooling2D(pool_size=(2,2), name='maxpool2d_2'))
cnn.add(Dropout(0.4, name='dropout_2'))
# flattening
cnn.add(Flatten(name='flatten'))
# densing
cnn.add(
Dense(
128,
activation='elu',
kernel_initializer='he_normal',
name='dense_1'
)
)
cnn.add(BatchNormalization(name='batchnorm_7'))
cnn.add(Dropout(0.6, name='dropout_4'))
cnn.add(
Dense(
num_classes,
activation='softmax',
name='out_layer'
)
)
cnn.compile(
loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy']
)
# usando GPU pode aumentar a qntd de epochs
epochs = 25
# dizem que batch size = 32 funciona melhor, mas vi mtts com 64
batch_size = 32
cnn.summary()
# training the cnn
history = cnn.fit(x = train_set,epochs = epochs,validation_data = validation_set)
cnn.save("CNN_PHOSTOS_3.h5")
```
| github_jupyter |
```
import fitsio
import numpy as np
from matplotlib import pyplot as plt
from astropy.table import Table,unique,join
import healpy as hp
dirz = '/global/cfs/cdirs/desi/survey/catalogs/SV1/redshift_comps/daily/v1/ELG'
fa = Table.read(dirz+'/alltiles_ELGzinfo.fits')
pixfn = '/global/cfs/cdirs/desi/target/catalogs/dr9/0.50.0/pixweight/main/resolve/dark/pixweight-1-dark.fits'
hdr = fitsio.read_header(pixfn,ext=1)
nside,nest = hdr['HPXNSIDE'],hdr['HPXNEST']
print(nside,nest)
def radec2thphi(ra,dec):
return (-dec+90.)*np.pi/180.,ra*np.pi/180.
def thphi2radec(theta,phi):
return 180./np.pi*phi,-(180./np.pi*theta-90)
th,phi =radec2thphi(fa['TARGET_RA'],fa['TARGET_DEC'])
hpx = hp.ang2pix(nside,th,phi,nest=nest)
fp = fitsio.read(pixfn)
dg = np.zeros(len(fa))
for i in range(0,len(dg)):
dg[i] = fp['GALDEPTH_G'][hpx[i]]
fa['GALDEPTH_G'] = dg
#select data with good redshifts and enough observing time
rdmin = 1000
wz = fa['FIBERSTATUS'] == 0
wz &= fa['ZWARN'] == 0
wz &= fa['R_DEPTH_EBVAIR'] > rdmin
fz = unique(fa[wz],keys=['TARGETID'])
print(len(fz))
wn = (fz['PHOTSYS'] == 'N')
fzn = fz[wn]
ws = (fz['PHOTSYS'] == 'S')
fzs = fz[ws]
print(len(fzn),len(fzs))
plt.hist(fzs['GALDEPTH_G'])
plt.xlabel('GALDEPTH_G')
plt.ylabel('# of good unique SV1 ELGs, R_DEPTH_EBVAIR >'+str(rdmin))
plt.title('DECaLS')
plt.show()
plt.hist(fzn['GALDEPTH_G'])
plt.xlabel('GALDEPTH_G')
plt.ylabel('# of good unique SV1 ELGs, R_DEPTH_EBVAIR >'+str(rdmin))
plt.title('BASS/MzLS')
plt.show()
ws = fzs['GALDEPTH_G'] > 1500
print(sum(fzs[ws]['elgqso_weight']),len(fzs[~ws]['elgqso_weight']))
nbin = 15
rng = (0.1,1.6)
bs = (rng[1]-rng[0])/nbin
a = plt.hist(fzs[ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzs[ws]['elgqso_weight'],label='GALDEPTH_G>1500')
b = plt.hist(fzs[~ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzs[~ws]['elgqso_weight'],label='GALDEPTH_G<1500')
plt.clf()
zl = a[1][:-1]+0.05
an = a[0]/np.sum(a[0])/bs
bn = b[0]/np.sum(b[0])/bs
plt.errorbar(zl,an,np.sqrt(a[0])/np.sum(a[0])/bs,label='GALDEPTH_G>1500')
plt.errorbar(zl+0.01,bn,np.sqrt(b[0])/np.sum(b[0])/bs,label='GALDEPTH_G<1500')
plt.legend()
plt.xlabel('redshift')
plt.ylabel('dN/dz ELG')
plt.title('DECaLS')
plt.show()
plt.plot(zl,bn/an)
ol = np.ones(len(bn))
plt.plot(zl,ol,':k')
plt.ylim(0.7,1.3)
plt.xlabel('redshift')
plt.ylabel('dN/dz low/high depth ratio')
plt.title('DECaLS ELGs, splitting on GALDEPTH_G at 1500')
ws = fzn['GALDEPTH_G'] > 550
print(sum(fzn[ws]['elgqso_weight']),len(fzn[~ws]['elgqso_weight']))
nbin = 15
rng = (0.1,1.6)
bs = (rng[1]-rng[0])/nbin
a = plt.hist(fzn[ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzn[ws]['elgqso_weight'],label='GALDEPTH_G>550')
b = plt.hist(fzn[~ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzn[~ws]['elgqso_weight'],label='GALDEPTH_G<550')
plt.clf()
zl = a[1][:-1]+0.05
an = a[0]/np.sum(a[0])/bs
bn = b[0]/np.sum(b[0])/bs
plt.errorbar(zl,an,np.sqrt(a[0])/np.sum(a[0])/bs,label='GALDEPTH_G>550')
plt.errorbar(zl+0.01,bn,np.sqrt(b[0])/np.sum(b[0])/bs,label='GALDEPTH_G<550')
plt.legend()
plt.xlabel('redshift')
plt.ylabel('dN/dz ELG')
plt.title('BASS/MzLS')
plt.show()
plt.plot(zl,bn/an)
ol = np.ones(len(bn))
plt.plot(zl,ol,':k')
plt.ylim(0.7,1.3)
plt.xlabel('redshift')
plt.ylabel('dN/dz low/high depth ratio')
plt.title('BASS/MzLS ELGs, splitting on GALDEPTH_G at 550')
from desitarget.sv1 import sv1_targetmask
tarbit2 = sv1_targetmask.desi_mask['ELG_FDR_GFIB']
ws = (fzs['SV1_DESI_TARGET'] & tarbit2) > 0
fzsf = fzs[ws]
print(len(fzsf))
ws = (fzn['SV1_DESI_TARGET'] & tarbit2) > 0
fznf = fzn[ws]
print(len(fznf))
ws = fzsf['GALDEPTH_G'] > 1500
print(sum(fzsf[ws]['elgqso_weight']),len(fzsf[~ws]['elgqso_weight']))
nbin = 15
rng = (0.1,1.6)
bs = (rng[1]-rng[0])/nbin
a = plt.hist(fzsf[ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzsf[ws]['elgqso_weight'],label='GALDEPTH_G>1500')
b = plt.hist(fzsf[~ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fzsf[~ws]['elgqso_weight'],label='GALDEPTH_G<1500')
plt.clf()
zl = a[1][:-1]+0.05
an = a[0]/np.sum(a[0])/bs
bn = b[0]/np.sum(b[0])/bs
plt.errorbar(zl,an,np.sqrt(a[0])/np.sum(a[0])/bs,label='GALDEPTH_G>1500')
plt.errorbar(zl,bn,np.sqrt(b[0])/np.sum(b[0])/bs,label='GALDEPTH_G<1500')
plt.legend()
plt.xlabel('redshift')
plt.ylabel('dN/dz ELG')
plt.title('DECaLS FDR GFIB')
plt.show()
plt.plot(zl,bn/an)
ol = np.ones(len(bn))
plt.plot(zl,ol,':k')
plt.ylim(0.7,1.3)
plt.xlabel('redshift')
plt.ylabel('dN/dz low/high depth ratio')
plt.title('DECaLS FDR GIFB ELGs, splitting on GALDEPTH_G at 1500')
ws = fznf['GALDEPTH_G'] > 550
print(sum(fznf[ws]['elgqso_weight']),len(fznf[~ws]['elgqso_weight']))
nbin = 15
rng = (0.1,1.6)
bs = (rng[1]-rng[0])/nbin
a = plt.hist(fznf[ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fznf[ws]['elgqso_weight'],label='GALDEPTH_G>550')
b = plt.hist(fznf[~ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fznf[~ws]['elgqso_weight'],label='GALDEPTH_G<550')
plt.clf()
zl = a[1][:-1]+0.05
an = a[0]/np.sum(a[0])/bs
bn = b[0]/np.sum(b[0])/bs
plt.errorbar(zl,an,np.sqrt(a[0])/np.sum(a[0])/bs,label='GALDEPTH_G>550')
plt.errorbar(zl+0.01,bn,np.sqrt(b[0])/np.sum(b[0])/bs,label='GALDEPTH_G<550')
plt.legend()
plt.xlabel('redshift')
plt.ylabel('dN/dz ELG')
plt.title('BASS/MzLS FDR GFIB')
plt.show()
plt.plot(zl,bn/an)
ol = np.ones(len(bn))
plt.plot(zl,ol,':k')
plt.ylim(0.7,1.3)
plt.xlabel('redshift')
plt.ylabel('dN/dz low/high depth ratio')
plt.title('BASS/MzLS FDR GFIB ELGs, splitting on GALDEPTH_G at 550')
plt.hist(fzn['EBV'])
ws = fznf['EBV'] > 0.06
print(sum(fznf[ws]['elgqso_weight']),len(fznf[~ws]['elgqso_weight']))
nbin = 15
rng = (0.1,1.6)
bs = (rng[1]-rng[0])/nbin
a = plt.hist(fznf[ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fznf[ws]['elgqso_weight'],label='EBV>0.06')
b = plt.hist(fznf[~ws]['Z'],histtype='step',bins=nbin,range=rng,weights=fznf[~ws]['elgqso_weight'],label='EBV_G<0.06')
plt.clf()
zl = a[1][:-1]+0.05
an = a[0]/np.sum(a[0])/bs
bn = b[0]/np.sum(b[0])/bs
plt.errorbar(zl,an,np.sqrt(a[0])/np.sum(a[0])/bs,label='EBV>0.06')
plt.errorbar(zl+0.01,bn,np.sqrt(b[0])/np.sum(b[0])/bs,label='EBV<0.06')
plt.legend()
plt.xlabel('redshift')
plt.ylabel('dN/dz ELG')
plt.title('BASS/MzLS FDR GFIB')
plt.show()
plt.plot(zl,bn/an)
ol = np.ones(len(bn))
plt.plot(zl,ol,':k')
plt.ylim(0.7,1.3)
plt.xlabel('redshift')
plt.ylabel('dN/dz low/high EBV ratio')
plt.title('BASS/MzLS FDR GFIB ELGs, splitting on EBV at 0.06')
plt.hist(fzs['EBV'])
```
| github_jupyter |
# Quantum Spins
We are going consider a magnetic insulator,
a Mott insulator. In this problem we have a valence electron per
atom, with very localized wave-functions. The Coulomb repulsion between
electrons on the same orbital is so strong, that electrons are bound to
their host atom, and cannot move. For this reason, charge disappears
from the equation, and the only remaining degree of freedom is the spin
of the electrons. The corresponding local state can therefore be either
$|\uparrow\rangle$ or $|\downarrow\rangle$. The only interaction taking
place is a process that “flips” two anti-aligned neighboring spins
$|\uparrow\rangle|\downarrow\rangle \rightarrow |\downarrow\rangle|\uparrow\rangle$.
Let us now consider a collection of spins residing on site of a
–one-dimensional for simplicity– lattice. An arbitrary state of
$N$-spins can be described by using the $S^z$ projection
($\uparrow,\downarrow$) of each spin as: $|s_1,s_2,..., s_N\rangle$. As
we can easily see, there are $2^N$ of such configurations.
Let us now consider a collection of spins residing on site of a
–one-dimensional for simplicity– lattice. An arbitrary state of
$N$-spins can be described by using the $S^z$ projection
($\uparrow,\downarrow$) of each spin as: $|s_1,s_2,..., s_N\rangle$. As
we can easily see, there are $2^N$ of such configurations.
We shall describe the interactions between neighboring spins using the
so-called Heisenberg Hamiltonian:
$$\hat{H}=\sum_{i=1}^{N-1} \hat{\mathbf{S}}_i \cdot \hat{\mathbf{S}}_{i+1}
$$
where $\hat{\mathbf{S}}_i = (\hat{S}^x,\hat{S}^y,\hat{S}^z)$ is the spin
operator acting on the spin on site $i$.
Since we are concerned about spins one-half, $S=1/2$, all these
operators have a $2\times2$ matrix representation, related to the
well-known Pauli matrices:
$$S^z = \left(
\begin{array}{cc}
1/2 & 0 \\
0 & -1/2
\end{array}
\right),
S^x = \left(
\begin{array}{cc}
0 & 1/2 \\
1/2 & 0
\end{array}
\right),
S^y = \left(
\begin{array}{cc}
0 & -i/2 \\
i/2 & 0
\end{array}
\right),
$$
These matrices act on two-dimensional vectors defined by the basis
states $|\uparrow\rangle$ and $|\downarrow\rangle$. It is useful to
introduce the identities:
$$\hat{S}^\pm = \left(\hat{S}^x \pm i \hat{S}^y\right),$$ where $S^+$
and $S^-$ are the spin raising and lowering operators. It is intuitively
easy to see why by looking at how they act on the basis states:
$\hat{S}^+|\downarrow\rangle = |\uparrow\rangle$ and
$\hat{S}^-|\uparrow\rangle = |\downarrow\rangle$. Their corresponding
$2\times2$ matrix representations are: $$S^+ = \left(
\begin{array}{cc}
0 & 1 \\
0 & 0
\end{array}
\right),
S^- = \left(
\begin{array}{cc}
0 & 0 \\
1 & 0
\end{array}
\right),
$$
We can now re-write the Hamiltonian (\[heisenberg\]) as:
$$\hat{H}=\sum_{i=1}^{N-1}
\hat{S}^z_i \hat{S}^z_{i+1} +
\frac{1}{2}\left[
\hat{S}^+_i \hat{S}^-_{i+1} +
\hat{S}^-_i \hat{S}^+_{i+1}
\right]
$$ The first term in this expression is diagonal and
does not flip spins. This is the so-called Ising term. The second term
is off-diagonal, and involves lowering and raising operators on
neighboring spins, and is responsible for flipping anti-aligned spins.
This is the “$XY$” part of the Hamiltonian.
The Heisenberg spin chain is a paradigmatic model in condensed matter.
Not only it is attractive due to its relative simplicity, but can also
describe real materials that can be studied experimentally. The
Heisenberg chain is also a prototypical integrable system, that can be
solved exactly by the Bethe Ansatz, and can be studied using
bosonization techniques and conformal field theory.
The Heisenberg spin chain is a paradigmatic model in condensed matter.
Not only it is attractive due to its relative simplicity, but can also
describe real materials that can be studied experimentally. The
Heisenberg chain is also a prototypical integrable system, that can be
solved exactly by the Bethe Ansatz, and can be studied using
bosonization techniques and conformal field theory.
In these lectures, we will be interested in obtaining its ground state
properties of this model by numerically solving the time-independent
Schrödinger equation: $$\hat{H}|\Psi\rangle = E|\Psi\rangle,$$ where $H$
is the Hamiltonian of the problem, $|\Psi\rangle$ its eigenstates, with
the corresponding eigenvalues, or energies $E$.
Exact diagonalization
=====================

#### Pictorial representation of the Hamiltonian building recursion explained in the text. At each step, the block size is increased by adding a spin at a time.
In this section we introduce a technique that will allow us to calculate
the ground state, and even excited states of small Heisenberg chains.
Exact Diagonalization (ED) is a conceptually simple technique which
basically consists of diagonalizing the Hamiltonian matrix by brute
force. Same as for the spin operators, the Hamiltonian also has a
corresponding matrix representation. In principle, if we are able to
compute all the matrix elements, we can use a linear algebra package to
diagonalize it and obtain all the eigenvalues and eigenvectors @lapack.
In these lectures we are going to follow a quite unconventional
procedure to describe how this technique works. It is important to point out that this
is a quite inefficient and impractical way to diagonalize the
Hamiltonian, and more sophisticated techniques are generally used in
practice.
Two-spin problem
----------------
The Hilbert space for the two-spin problem consists of four possible
configurations of two spins
$$\left\{ |\uparrow\uparrow\rangle,|\uparrow\downarrow\rangle,|\downarrow\uparrow\rangle,|\downarrow\downarrow\rangle \right\}$$
The problem is described by the Hamiltonian: $$\hat{H}=
\hat{S}^z_1 \hat{S}^z_2 +
\frac{1}{2}\left[
\hat{S}^+_1 \hat{S}^-_2 +
\hat{S}^-_1 \hat{S}^+_2
\right]$$
The corresponding matrix will have dimensions $4 \times 4$. In order to
compute this matrix we shall use some simple matrix algebra to first
obtain the single-site operators in the expanded Hilbert space. This is
done by following the following simple scheme: And operator $O_1$ acting
on the left spin, will have the following $4 \times 4$ matrix form:
$$\tilde{O}_1 = O_1 \otimes {1}_2
$$ Similarly, for an operator $O_2$ acting on the right
spin: $$\tilde{O}_2 = {1}_2 \otimes O_2
$$ where we introduced the $n \times n$ identity matrix
${1}_n$. The product of two operators acting on different sites
can be obtained as: $$\tilde{O}_{12} = O_1 \otimes O_2$$ It is easy to
see that the Hamiltonian matrix will be given by: $$H_{12}=
S^z \otimes S^z +
\frac{1}{2}\left[
S^+ \otimes S^- +
S^- \otimes S^+
\right]$$ where we used the single spin ($2 \times 2$) matrices $S^z$
and $S^\pm$. We leave as an exercise for the reader to show that the
final form of the matrix is:
$$H_{12} = \left(
\begin{array}{cccc}
1/4 & 0 & 0 & 0 \\
0 & -1/4 & 1/2 & 0 \\
0 & 1/2 & -1/4 & 0 \\
0 & 0 & 0 & 1/4 \\
\end{array}
\right),
$$
Obtaining the eigenvalues and eigenvectors is also a straightforward
exercise: two of them are already given, and the entire problem now
reduces to diagonalizing a two by two matrix. We therefore obtain the
well known result: The ground state
$|s\rangle = 1/\sqrt{2}\left[ |\uparrow\downarrow\rangle - |\downarrow\uparrow\rangle \right]$,
has energy $E_s=-3/4$, and the other three eigenstates
$\left\{|\uparrow\uparrow\rangle,|\downarrow\downarrow\rangle,1/\sqrt{2}\left[ |\uparrow\downarrow\rangle + |\downarrow\uparrow\rangle \right] \right\}$
form a multiplet with energy $E_t=1/4$.
Many spins
----------
Imagine now that we add a third spin to the right of our two spins. We
can use the previous result to obtain the new $8 \times 8$ Hamiltonian
matrix as: $$H_{3}=
H_{2} \otimes {1}_2 +
\tilde{S}^z_2 \otimes S^z +
\frac{1}{2}\left[
\tilde{S}^+_2 \otimes S^- +
\tilde{S}^-_2 \otimes S^+
\right]$$ Here we used the single spin $S^z_1$, $S^\pm_1$, and the
\`tilde\` matrices defined in Eqs.(\[tildeL\]) and (\[tildeR\]):
$$\tilde{S}^z_2 = {1}_2 \otimes S^z,$$ and
$$\tilde{S}^\pm_2 = {1}_2 \otimes S^\pm,$$
It is easy to see that this leads to a recursion scheme to construct the
$2^i \times 2^i$ Hamiltonian matrix the $i^\mathrm{th}$ step as:
$$H_{i}=
H_{i-1} \otimes {1}_2 +
\tilde{S}^z_{i-1} \otimes S^z +
\frac{1}{2}\left[
\tilde{S}^+_{i-1} \otimes S^- +
\tilde{S}^-_{i-1} \otimes S^+
\right]$$
with $$\tilde{S}^z_{i-1} = {1}_{2^{i-2}} \otimes S^z,$$ and
$$\tilde{S}^\pm_{i-1} = {1}_{2^{i-2}} \otimes S^\pm,$$
This recursion algorithm can be visualized as a left ‘block’, to which
we add new ‘sites’ or spins to the right, one at a time, as shown in
Fig.\[fig:block\].The block has a ‘block Hamiltonian’, $H_L$, that is
iteratively built by connecting to the new spins through the
corresponding interaction terms.
```
%matplotlib inline
import numpy as np
from matplotlib import pyplot
# PARAMETERS
nsites = 4
#Single site operators
sz0 = np.zeros(shape=(2,2)) # single site Sz
splus0 = np.zeros(shape=(2,2)) # single site S+
sz0[0,0] = -0.5
sz0[1,1] = 0.5
splus0[1,0] = 1.0
term_szsz = np.zeros(shape=(4,4)) #auxiliary matrix to store Sz.Sz
term_szsz = np.kron(sz0,sz0)
term_spsm = np.zeros(shape=(4,4)) #auxiliary matrix to store 1/2 S+.S-
term_spsm = np.kron(splus0,np.transpose(splus0))*0.5
term_spsm += np.transpose(term_spsm)
h12 = term_szsz+term_spsm
H = np.zeros(shape=(2,2))
for i in range(1,nsites):
diml = 2**(i-1) # 2^i
dim = diml*2
print ("ADDING SITE ",i," DIML= ",diml)
Ileft = np.eye(diml)
Iright = np.eye(2)
# We add the term for the interaction S_i.S_{i+1}
aux = np.zeros(shape=(dim,dim))
aux = np.kron(H,Iright)
H = aux
H = H + np.kron(Ileft,h12)
w, v = np.linalg.eigh(H) #Diagonalize the matrix
print(w)
from matplotlib import pyplot
pyplot.rcParams['axes.linewidth'] = 2 #set the value globally
%matplotlib inline
Beta = np.arange(0.1,10,0.1)
et = np.copy(Beta)
n = 0
for x in Beta:
p = np.exp(-w*x)
z = np.sum(p)
et[n] = np.dot(w,p)/z
print (x,et[n])
n+=1
pyplot.plot(1/Beta,et,lw=2);
```
#### Challenge 12.1:
Compute the energy and specific heat of the spin chain with $L=12$ as a function of temperature, for $T < 4$.
# A practical exact diagonalization algorithm
1. Initialization: Topology of the lattice, neighbors, and signs.
2. Contruction of a basis suitable for the problem.
3. Constuction of the matrix element of the Hamiltonian.
4. Diagonalization of the matrix.
5. Calculation of observables or expectation values.
As we shall see below, we are going to need the concept of “binary
word”. A binary word is the binary representation of an integer (in
powers of ‘two’, i.e. $n=\sum_{i}b_{i}.2^{i}$), and consist of a
sequence of ‘bits’. A bit $b_{i}$ in the binary basis correspond to our
digits in the decimal system, and can assumme the values ‘zero’ or
‘one’. At this stage we consider appropriate to introduce the logical
operators `AND, OR, XOR`. The binary operators act between bits and
their multiplication tables are listed below
**AND** | 0 | 1
:---:|:---:|:---:
**0** | 0 | 0
**1** | 0 | 1
**OR** | 0 | 1
:---:|:---:|:---:
**0** | 0 | 1
**1** | 1 | 1
**XOR** | 0 | 1
:---:|:---:|:---:
**0** | 0 | 1
**1** | 1 | 0
The bit manipulation is a very useful tool, and most of the programming
languages provide the needed commands.
## Initilalization and definitions
In the program, it is convenient to store in arrays all those quantities
that will be used more frequently. In particular, we must determine the
topology of the cluster, labeling the sites, and storing the components
of the lattice vectors. We must also generate arrays with the nearest
neighbors, and the next-nearest neighbors, according to our needs.
## Construction of the basis
Memory limitations impose severe restrictions on the size of the
clusters that can be studied with this method. To understand this point,
note that although the lowest energy state can be written in the
$\{|\phi
_{n}\rangle \}$ basis as
$|\psi _{0}\rangle =\sum_{m}c_{m}|\phi _{m}\rangle $, this expression is
of no practical use unless $|\phi _{m}\rangle $ itself is expressed in a
convenient basis to which the Hamiltonian can be easily applied. A
natural orthonormal basis for fermion systems is the occupation number
representation, describing all the possible distributions of $N_{e}$
electrons over $N$ sites, while for spin systems it is covenient to work
in a basis where the $S_{z}$ is defined at every site, schematically
represented as $|n\rangle =|\uparrow \downarrow \uparrow ...\rangle $.
The size of this type of basis set
grows exponentially with the system size. In practice this problem can be
considerably alleviated by the use of symmetries of the Hamiltonian that
reduces the matrix to a block-form. The most obvious symmetry is the
number of particles in the problem which is usually conserved at least
for fermionic problems. The total projection of the spin
$S_{total}^{z}$, is also a good quantum number. For translationally
invariant problems, the total momentum $\mathbf{k%
}$ of the system is also conserved introducing a reduction of $1/N$ in
the number of states (this does not hold for models with open boundary
conditions or explicit disorder). In addition, several Hamiltonians have
additional symmetries. On a square lattice, rotations in $\pi /2$ about
a given site, spin inversion, and reflections with respect to the
lattice axis are good quantum numbers (although care must be taken in
their implementation since some of these operations are combinations of
others and thus not independent).
In the following we shall consider a spin-1/2 Heisenberg chain as a
practical example. In this model it is useful to represent the spins
pointing in the ‘up’ direction by a digit ‘1’, and the down-spins by a
‘0’. Following this rule, a state in the $S^{z}$ basis can be
represented by a sequence of ones and zeroes, i.e., a “binary word”.
Thus, two Néel configurations in the 4-site chain can be seen as
$$\begin{aligned}
\mid \uparrow \downarrow \uparrow \downarrow \rangle \equiv |1010\rangle , \\
\mid \downarrow \uparrow \downarrow \uparrow \rangle \equiv |0101\rangle .\end{aligned}$$
Once the up-spins have been placed the whole configuration has been
uniquely determined since the remaining sites can only be occupied by
down-spins. The resulting binary number can be easily converted into
integers $i\equiv
\sum_{l}b_{l}.2^{l}$, where the summation is over all the sites of the
lattice, and $b_{l}$ can be $1$ or $0$. For example:
$$\begin{array}{lllll}
2^{3} & 2^{2} & 2^{1} & 2^{0} & \\
1 & 0 & 1 & 0 & \rightarrow 2^{1}+2^{3}=10, \\
0 & 1 & 0 & 1 & \rightarrow 2^{0}+2^{2}=5.
\end{array}$$
Using the above convention, we can construct the whole basis for the
given problem. However, we must consider the memory limitations of our
computer, introducing some symetries to make the problem more tractable.
The symetries are operations that commute with the Hamiltonian, allowing
us to divide the basis in subspaces with well defined quantum numbers.
By means of similarity transformations we can generate a sequence of
smaller matrices along the diagonal (i.e. the Hamiltonian matrix is
“block diagonal”), and each of them can be diagonalized independently.
For fermionic systems, the simplest symmetries are associated to the
conservation of the number of particles and the projection
$S_{total}^{z}$ of the total spin in the $z$ direction. In the spin-1/2
Heisenberg model, a matrix with $2^{N}\times 2^{N}$ elements can be
reduced to $2S+1$ smaller matrices, corresponding to the projections $%
m=(N_{\uparrow }-N_{\downarrow })/2=-S,-S+1,...,S-1,S$, with $S=N/2$.
The dimension of each of these subspaces is obtained from the
combinatory number $\left(
\begin{array}{c}
N \\
N_{\uparrow }
\end{array}
\right) = \frac{N!}{N_{\uparrow }!N_{\downarrow }!}$.
#### Example: 4-site Heisenberg chain
The total basis has
$2^{4}=16$ states that can be grouped in
$4+1=5$ subspaces with well defined values of the quantum
number $m=S^z$. The dimensions of these subspaces are:
m | dimension
:--:|:--:
-2 | 1
-1 | 4
0 | 6
2 | 4
1 | 1
Since we know that the ground state of the Hilbert
chain is a singlet, we are only interested in the subspace with
</span>$S_{total}^{z}=m=0$<span>. For our example, this subspace is
given by (in increasing order): </span>
$$\begin{eqnarray}
\mid \downarrow \downarrow \uparrow \uparrow \rangle & \equiv &|0011\rangle
\equiv |3\rangle , \\
\mid \downarrow \uparrow \downarrow \uparrow \rangle &\equiv &|0101\rangle
\equiv |5\rangle , \\
\mid \downarrow \uparrow \uparrow \downarrow \rangle &\equiv &|0110\rangle
\equiv |6\rangle , \\
\mid \uparrow \downarrow \downarrow \uparrow \rangle &\equiv &|1001\rangle
\equiv |9\rangle , \\
\mid \uparrow \downarrow \uparrow \downarrow \rangle &\equiv &|1010\rangle
\equiv |10\rangle , \\
\mid \uparrow \uparrow \downarrow \downarrow \rangle &\equiv &|1010\rangle
\equiv |12\rangle .
\end{eqnarray}
$$
Generating the possible configurations of $N$ up-spins (or
spinless fermions) in $L$ sites is equivalent to the problem of
generating the corresponding combinations of zeroes and ones in
lexicographic order.
## Construction of the Hamiltonian matrix elements
We now have to address the problem of generating all the Hamiltonian matrix elements
. These are obtained by the application of the
Hamiltonian on each state of the basis $|\phi \rangle $, generating all
the values
$H_{\phi ,\phi ^{\prime }}=\langle \phi ^{\prime }|{{\hat{H}}}%
|\phi \rangle $. We illustrate this procedure in the following
pseudocode:
    **for** each term ${\hat{H}_i}$ of the Hamiltonian ${\hat{H}}$
        **for** all the states $|\phi\rangle $ in the basis
            $\hat{H}_i|\phi\rangle = \langle \phi^{\prime }|{\hat{H}_i}|\phi\rangle |\phi^{\prime }\rangle $
            
$H_{\phi,\phi^{\prime }}=H_{\phi,\phi^{\prime }} + \langle \phi^{\prime }|{\hat{H}_i}|\phi\rangle $
        **end for**
    **end for**
As the states are representated by binary words, the spin operators (as
well as the creation and destructions operators) can be be easily
implemented using the logical operators `AND, OR, XOR`.
As a practical example let us consider the spin-1/2 Heisenberg
Hamiltonian on the 4-sites chain:
$${{\hat{H}}}=J\sum_{i=0}^{3}\mathbf{S}_{i}.\mathbf{S}_{i+1}=J%
\sum_{i=0}^{3}S_{i}^{z}.S_{i+1}^{z}+\frac{J}{2}%
\sum_{i=0}^{3}(S_{i}^{+}S_{i+1}^{-}+S_{i}^{-}S_{i+1}^{+}), $$
with $\mathbf{S}_{4}=\mathbf{S}_{0}$, due to the periodic boundary
conditions. To evaluate the matrix elements in the basis (\[basis0\]) we
must apply each term of ${{\hat{H}}}$ on such states. The first term is
called Ising term, and is diagonal in this basis (also called Ising
basis). The last terms, or fluctuation terms, give strictly off-diagonal
contributions to the matrix in this representation (when symmetries are
considered, they can also have diagonal contributions). These
fluctuations cause an exchange in the spin orientation between neighbors
with opposite spins, e.g. $\uparrow \downarrow \rightarrow $
$\downarrow \uparrow $. The way to implement spin-flip operations of
this kind on the computer is defining new ‘masks’. A mask for this
operation is a binary number with ‘zeroes’ everywhere, except in the
positions of the spins to be flipped, e.g 00..0110...00. Then, the
logical operator `XOR` is used between the initial state and the mask to
invert the bits (spins) at the positions indicated by the mask. For
example, (0101)$\mathtt{.}$`XOR.`(1100)=(1001), i.e., 5`.XOR.`12=9. It
is useful to store all the masks for a given geometry in memory,
generating them immediately after the tables for the sites and neighbors.
<span>In the Heisenberg model the spin flips can be implemented using
masks, with ‘zeroes’ everywhere, except in the positions of the spins to
be flipped. To illustrate this let us show the effect of the
off-diagonal terms on one of the Néel configurations:</span>
$$\begin{eqnarray}
S_{0}^{+}S_{1}^{-}\,+S_{0}^{-}S_{1}^{+}\,|0101\rangle &\equiv &(0011)\mathtt{%
.XOR.}(0101)=3\mathtt{.XOR.}5=(0110)=6\equiv \,|0110\rangle , \\
S_{1}^{+}S_{2}^{-}+S_{1}^{-}S_{2}^{+}\,\,|0101\rangle &\equiv &(0110)\mathtt{%
.XOR.}(0101)=6\mathtt{.XOR.}5=(0011)=3\equiv \,|0011\rangle , \\
S_{2}^{+}S_{3}^{-}+S_{2}^{-}S_{3}^{+}\,\,|0101\rangle &\equiv &(1100)\mathtt{%
.XOR.}(0101)=12\mathtt{.XOR.}5=(1001)=9\equiv \,|1001\rangle , \\
S_{3}^{+}S_{0}^{-}\,+S_{3}^{-}S_{0}^{+}|0101\rangle &\equiv &(1001)\mathtt{%
.XOR.}(0101)=9\mathtt{.XOR.}5=(1100)=12\equiv \,|1100\rangle .%
\end{eqnarray}$$
<span>After applying the Hamiltonian (\[h4\]) on our basis states, the
reader can verify as an exercise that we obtain </span>
$$\begin{eqnarray}
{{\hat{H}}}\,|0101\rangle &=&-J\,|0101\rangle +\frac{J}{2}\left[
\,|1100\rangle +\,|1001\rangle +\,|0011\rangle +\,|0110\rangle \right] , \\
{{\hat{H}}}\,|1010\rangle &=&-J\,|1010\rangle +\frac{J}{2}\left[
\,|1100\rangle +\,|1001\rangle +\,|0011\rangle +\,|0110\rangle \right] , \\
{{\hat{H}}}\,|0011\rangle &=&\frac{J}{2}\left[ \,\,|0101\rangle
+\,|1010\rangle \right] , \\
{{\hat{H}}}\,|0110\rangle &=&\frac{J}{2}\left[ \,\,|0101\rangle
+\,|1010\rangle \right] , \\
{{\hat{H}}}\,|1001\rangle &=&\frac{J}{2}\left[ \,|0101\rangle +\,|1010\rangle
\right] , \\
{{\hat{H}}}\,|1100\rangle &=&\frac{J}{2}\left[ \,|0101\rangle +\,|1010\rangle
\right] .%
\end{eqnarray}$$
<span>The resulting matrix for the operator </span>$\hat{H}$<span>in the
</span>$S^{z}$<span> representation is: </span> $$H=J\left(
\begin{array}{llllll}
0 & 1/2 & 0 & 0 & 1/2 & 0 \\
1/2 & -1 & 1/2 & 1/2 & 0 & 1/2 \\
0 & 1/2 & 0 & 0 & 1/2 & 0 \\
0 & 1/2 & 0 & 0 & 1/2 & 0 \\
1/2 & 0 & 1/2 & 1/2 & -1 & 1/2 \\
0 & 1/2 & 0 & 0 & 1/2 & 0
\end{array}
\right) . $$
```
class BoundaryCondition:
RBC, PBC = range(2)
class Direction:
RIGHT, TOP, LEFT, BOTTOM = range(4)
L = 4
Nup = 2
maxdim = 2**L
bc = BoundaryCondition.PBC
# Tables for energy
hdiag = np.zeros(4) # Diagonal energies
hflip = np.zeros(4) # off-diagonal terms
hdiag[0] = +0.25
hdiag[1] = -0.25
hdiag[2] = -0.25
hdiag[3] = +0.25
hflip[0] = 0.
hflip[1] = 0.5
hflip[2] = 0.5
hflip[3] = 0.
#hflip *= 2
#hdiag *= 0.
# Lattice geometry (1D chain)
nn = np.zeros(shape=(L,4), dtype=np.int16)
for i in range(L):
nn[i, Direction.RIGHT] = i-1
nn[i, Direction.LEFT] = i+1
if(bc == BoundaryCondition.RBC): # Open Boundary Conditions
nn[0, Direction.RIGHT] = -1 # This means error
nn[L-1, Direction.LEFT] = -1
else: # Periodic Boundary Conditions
nn[0, Direction.RIGHT] = L-1 # We close the ring
nn[L-1, Direction.LEFT] = 0
# We build basis
basis = []
dim = 0
for state in range(maxdim):
basis.append(state)
dim += 1
print ("Basis:")
print (basis)
# We build Hamiltonian matrix
H = np.zeros(shape=(dim,dim))
def IBITS(n,i):
return ((n >> i) & 1)
for i in range(dim):
state = basis[i]
# Diagonal term
for site_i in range(L):
site_j = nn[site_i, Direction.RIGHT]
if(site_j != -1): # This would happen for open boundary conditions
two_sites = IBITS(state,site_i) | (IBITS(state,site_j) << 1)
value = hdiag[two_sites]
H[i,i] += value
for i in range(dim):
state = basis[i]
# Off-diagonal term
for site_i in range(L):
site_j = nn[site_i, Direction.RIGHT]
if(site_j != -1):
mask = (1 << site_i) | (1 << site_j)
two_sites = IBITS(state,site_i) | (IBITS(state,site_j) << 1)
value = hflip[two_sites]
if(value != 0.):
new_state = (state ^ mask)
j = new_state
H[i,j] += value
print(H)
d, v = np.linalg.eigh(H) #Diagonalize the matrix
print("===================================================================================================================")
print(d)
print(v[:,0])
# Using quantum number conservation : Sz
L=8
Nup=4
maxdim = 2**L
# Lattice geometry (1D chain)
nn = np.zeros(shape=(L,4), dtype=np.int16)
for i in range(L):
nn[i, Direction.RIGHT] = i-1
nn[i, Direction.LEFT] = i+1
if(bc == BoundaryCondition.RBC): # Open Boundary Conditions
nn[0, Direction.RIGHT] = -1 # This means error
nn[L-1, Direction.LEFT] = -1
else: # Periodic Boundary Conditions
nn[0, Direction.RIGHT] = L-1 # We close the ring
nn[L-1, Direction.LEFT] = 0
basis = []
dim = 0
for state in range(maxdim):
n_ones = 0
for bit in range(L):
n_ones += IBITS(state,bit)
if(n_ones == Nup):
basis.append(state)
dim += 1
print ("Dim=",dim)
print ("Basis:")
print (basis)
#hflip[:] *= 2
# We build Hamiltonian matrix
H = np.zeros(shape=(dim,dim))
def IBITS(n,i):
return ((n >> i) & 1)
for i in range(dim):
state = basis[i]
# Diagonal term
for site_i in range(L):
site_j = nn[site_i, Direction.RIGHT]
if(site_j != -1): # This would happen for open boundary conditions
two_sites = IBITS(state,site_i) | (IBITS(state,site_j) << 1)
value = hdiag[two_sites]
H[i,i] += value
def bisect(state, basis):
# Binary search, only works in a sorted list of integers
# state : integer we seek
# basis : list of sorted integers in increasing order
# ret_val : return value, position on the list, -1 if not found
ret_val = -1
dim = len(basis)
# for i in range(dim):
# if(state == basis[i]):
# return i
origin = 0
end = dim-1
middle = (origin+end)/2
while(1):
index_old = middle
middle = (origin+end)//2
if(state < basis[middle]):
end = middle
else:
origin = middle
if(basis[middle] == state):
break
if(middle == index_old):
if(middle == end):
end = end - 1
else:
origin = origin + 1
ret_val = middle
return ret_val
for i in range(dim):
state = basis[i]
# Off-diagonal term
for site_i in range(L):
site_j = nn[site_i, Direction.RIGHT]
if(site_j != -1):
mask = (1 << site_i) | (1 << site_j)
two_sites = IBITS(state,site_i) | (IBITS(state,site_j) << 1)
value = hflip[two_sites]
if(value != 0.):
new_state = (state ^ mask)
j = bisect(new_state, basis)
H[i,j] += value
print(H)
d, v = np.linalg.eigh(H) #Diagonalize the matrix
print(d,np.min(d))
print(v[:,0])
```
Obtaining the ground-state: Lanczos diagonalization
---------------------------------------------------
Once we have a superblock matrix, we can apply a library routine to
obtain the ground state of the superblock $|\Psi\rangle$. The two
algorithms widely used for this purpose are the Lanczos and Davidson
diagonalization. Both are explained to great extent in Ref.@noack, so we
refer the reader to this material for further information. In these
notes we will briefly explain the Lanczos procedure.
The basic idea of the Lanczos method is that a special basis can be constructed
where the Hamiltonian has a tridiagonal representation. This is carried
out iteratively as shown below. First, it is necessary to select an
arbitrary seed vector $|\phi _{0}\rangle $ in the Hilbert space of the
model being studied. If we are seeking the ground-state of the model,
then it is necessary that the overlap between the actual ground-state
$|\psi
_{0}\rangle $, and the initial state $|\phi _{0}\rangle $ be nonzero. If
no “a priori” information about the ground state is known, this
requirement is usually easily satisfied by selecting an initial state
with *randomly* chosen coefficients in the working basis that is being
used. If some other information of the ground state is known, like its
total momentum and spin, then it is convenient to initiate the
iterations with a state already belonging to the subspace having those
quantum numbers (and still with random coefficients within this
subspace).
After $|\phi _{0}\rangle $ is selected, define a new vector by applying
the Hamiltonian ${{\hat{H}}}$, over the initial state. Subtracting the
projection over $|\phi _{0}\rangle $, we obtain
$$|\phi _{1}\rangle ={{\hat{H}}}|\phi _{0}\rangle -\frac{{\langle }\phi _{0}|{{%
\hat{H}}}|{\phi }_{0}{\rangle }}{\langle \phi _{0}|\phi _{0}\rangle }|\phi
_{0}\rangle ,$$ that satisfies $\langle \phi _{0}|\phi _{1}\rangle =0$.
Now, we can construct a new state that is orthogonal to the previous two
as,
$$|\phi _{2}\rangle ={{\hat{H}}}|\phi _{1}\rangle -{\frac{{\langle \phi _{1}|{%
\hat{H}}|\phi _{1}\rangle }}{{\langle \phi _{1}|\phi _{1}\rangle }}}|\phi
_{1}\rangle -{\frac{{\langle \phi _{1}|\phi _{1}\rangle }}{{\langle \phi
_{0}|\phi _{0}\rangle }}}|\phi _{0}\rangle .$$ It can be easily checked
that $\langle \phi _{0}|\phi _{2}\rangle
=\langle \phi _{1}|\phi _{2}\rangle =0$. The procedure can be
generalized by defining an orthogonal basis recursively as,
$$|\phi _{n+1}\rangle ={{\hat{H}}}|\phi _{n}\rangle -a_{n}|\phi _{n}\rangle
-b_{n}^{2}|\phi _{n-1}\rangle ,$$ where $n=0,1,2,...$, and the
coefficients are given by
$$a_{n}={\frac{{\langle \phi _{n}|{\hat{H}}|\phi _{n}\rangle }}{{\langle \phi
_{n}|\phi _{n}\rangle }}},\qquad b_{n}^{2}={\frac{{\langle \phi _{n}|\phi
_{n}\rangle }}{{\langle \phi _{n-1}|\phi _{n-1}\rangle }}},$$
supplemented by $b_{0}=0$, $|\phi _{-1}\rangle =0$. In this basis, it
can be shown that the Hamiltonian matrix becomes, $$H=\left(
\begin{array}{lllll}
a_{0} & b_{1} & 0 & 0 & ... \\
b_{1} & a_{1} & b_{2} & 0 & ... \\
0 & b_{2} & a_{2} & b_{3} & ... \\
0 & 0 & b_{3} & a_{3} & ... \\
\vdots
{} &
\vdots
&
\vdots
&
\vdots
&
\end{array}
\right)$$ i.e. it is tridiagonal as expected. Once in this form the
matrix can be diagonalized easily using standard library subroutines.
However, note that to diagonalize completely a Hamiltonian on a finite
cluster, a number of iterations equal to the size of the Hilbert space
(or the subspace under consideration) are needed. In practice this would
demand a considerable amount of CPU time. However, one of the advantages
of this technique is that accurate enough information about the ground
state of the problem can be obtained after a small number of iterations
(typically of the order of $\sim 100$ or less).
Another way to formulate the problem is by obtaining the tridiagonal
form of the Hamiltonian starting from a Krylov basis, which is spanned
by the vectors
$$\left\{|\phi_0\rangle,\hat{H}|\phi_0\rangle,\hat{H}^2|\phi_0\rangle,...,\hat{H}^n|\phi_0\rangle\right\}$$
and asking that each vector be orthogonal to the previous two. Notice
that each new iteration of the process requires one application of the
Hamiltonian. Most of the time this simple procedure works for practical
purposes, but care must be payed to the possibility of losing
orthogonality between the basis vectors. This may happen due to the
finite machine precision. In that case, a re-orthogonalization procedure
may be required.
Notice that the new super-Hamiltonian matrix has dimensions
$D_L D_R d^2 \times D_L D_R d^2$. This could be a large matrix. In
state-of-the-art simulations with a large number of states, one does not
build this matrix in memory explicitly, but applies the operators to the
state directly in the diagonalization routine.
```
def lanczos(m, seed, maxiter, tol, use_seed, calc_gs, force_maxiter = False):
x1 = seed
x2 = seed
gs = seed
a = np.zeros(100)
b = np.zeros(100)
z = np.zeros((100,100))
lvectors = []
control_max = maxiter;
if(maxiter == -1):
force_maxiter = False
if(control_max == 0):
gs = 1
maxiter = 1
return(e0,gs)
x1[:] = 0
x2[:] = 0
gs[:] = 0
maxiter = 0
a[:] = 0.0
b[:] = 0.0
if(use_seed):
x1 = seed
else :
x1 = np.random.random(x1.shape[0])*2-1.
b[0] = np.sqrt(np.dot(x1,x1))
x1 = x1 / b[0]
x2[:] = 0
b[0] = 1.
e0 = 9999
nmax = min(99, gs.shape[0])
for iter in range(1,nmax+1):
eini = e0
if(b[iter - 1] != 0.):
aux = x1
x1 = -b[iter-1] * x2
x2 = aux / b[iter-1]
aux = np.dot(m,x2)
x1 = x1 + aux
a[iter] = np.dot(x1, x2)
x1 = x1 - x2*a[iter]
b[iter] = np.sqrt(np.dot(x1, x1))
lvectors.append(x2)
z.resize((iter+1,iter+1))
z[:,:] = 0
for i in range(0,iter-1):
z[i,i+1] = b[i+1]
z[i+1,i] = b[i+1]
z[i,i] = a[i+1]
z[iter-1,iter-1]=a[iter]
d, v = np.linalg.eig(z)
col = 0
n = 0
e0 = 9999
for e in d:
if(e < e0):
e0 = e
col = n
n+=1
e0 = d[col]
print ("Iter = ",iter," Ener = ",e0)
if((force_maxiter and iter >= control_max) or (iter >= gs.shape[0] or iter == 99 or abs(b[iter]) < tol) or \
((not force_maxiter) and abs(eini-e0) <= tol)):
# converged
gs = 0.
for i in range(0,iter):
gs += v[i,col]*lvectors[i]
print ("E0 = ", e0, np.sqrt(np.dot(gs,gs)))
maxiter = iter
return(e0,gs) # We return with ground states energy
seed = np.zeros(H.shape[0])
e0, gs = lanczos(H,seed,6,1.e-5,False,False)
print(gs)
print(np.dot(gs,gs))
```
| github_jupyter |
```
import pandas as pd
sightings = pd.read_csv('koala-survey-sightings-data.csv', encoding='utf-8', parse_dates=[['Date', 'Time']])
# They used a shorthand for some of the genus
sightings['TreeSpecies'] = sightings['TreeSpecies'].replace({"E.": "Eucalyptus", "L.":"Lophostemon", "C.": "Corymbia", "A.": "Angophora"}, regex=True)
sightings['TreeSpecies'] = sightings['TreeSpecies'].dropna()
sightings.drop_duplicates(subset=['TreeSpecies'], inplace=True)
sightings[['TreeSpecies']].to_csv('koala-sightings-species.csv')
# ^ Upload that file to the ALA and download the results and field guide with a mapping to the correct species.
#sightings_with_species_info = pd.merge(sightings, unique_df, right_on="Scientific Name - original", left_on="TreeSpecies", how="left")
import requests
from urllib.parse import urlsplit
import os
processed_df = pd.read_csv('SEQ_Koala_Survey_Data_2010_-_2015.csv')
for _, row in processed_df.iterrows():
doc_id = urlsplit(row["guid"]).path.split('/')[-1]
r = requests.get(row["guid"] + ".json")
with open(os.path.join('apni', doc_id + '.json'), 'wb') as f:
f.write( r.content )
import requests
from bs4 import BeautifulSoup
import pandas as pd
from urllib.parse import urlsplit
import os
processed_df = pd.read_csv('SEQ_Koala_Survey_Data_2010_-_2015.csv')
for _, row in processed_df.iterrows():
try:
doc_id = urlsplit(row["guid"]).path.split('/')[-1]
path = os.path.join('apni', doc_id + '.txt')
if not os.path.exists(path):
response = requests.get(
url="https://en.wikipedia.org/wiki/{0}".format(row["scientificName"].replace(" ", "_")),
)
soup = BeautifulSoup(response.content, 'html.parser')
d = soup.find(id="Description").parent.next_sibling.next_sibling.get_text()
with open(os.path.join('apni', doc_id + '.txt'), 'w') as f:
f.write(d)
else:
print(path, " already exists")
except Exception as e:
print(e, row[0])
# generate plants.json
processed_df = pd.read_csv('SEQ_Koala_Survey_Data_2010_-_2015.csv')
index = {}
for _, row in processed_df.iterrows():
doc_id = urlsplit(row["guid"]).path.split('/')[-1]
path = os.path.join('apni', doc_id + '.txt')
index[row['Supplied Name']] = {'id': doc_id, 'scientificName': row['scientificName'], 'family': row['family'] }
print('<option value="{0}">{1}</option>'.format(row['Supplied Name'], row['Supplied Name']))
import json
with open('plants.json', 'w') as f:
f.write(json.dumps(index))
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.