text stringlengths 2.5k 6.39M | kind stringclasses 3
values |
|---|---|
<script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
<script>
window.dataLayer = window.dataLayer || [];
function gtag(){dataLayer.push(arguments);}
gtag('js', new Date());
gtag('config', 'UA-59152712-8');
</script>
# Sommerfeld Boundary Conditions
<font color='green'><b> Validated for all coordinate systems available in NRPy+ </b></font>
## Authors: Terrence Pierre Jacques & Zach Etienne
## Abstract
The aim of this notebook is to describe the mathematical motivation behind the Sommerfeld boundary condition, document how it is implemented within the NRPy+ infrastructure in Cartesian coordinates, and record ongoing validation tests against the [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk).
**Notebook Status:** <font color='green'><b> Validated against the Einstein Toolkit </b></font>
**Validation Notes:** The Sommerfeld boundary condition as implemented in NRPy+ has been validated against [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk)(ETK) for the case of a scalar wave propagating across a 3D Cartesian grid. We have agreement to roundoff error with the Einstein Toolkit. Specifically, we have achieved:
1. Roundoff level agreement for the wave propagating toward each of the individual faces
1. Roundoff level agreement using any combination of input parameters available in the [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk) (variable value at infinity, radial power, and wave speed)
[comment]: <> (Introduction: TODO)
<a id='toc'></a>
# Table of Contents
$$\label{toc}$$
This notebook is organized as follows
1. [Step 1](#initializenrpy): Set core NRPy+ parameters for numerical grids
1. [Step 2](#sbc): Definition and mathematical motivation
1. [Step 2.a](#intro): Introduction & background mathematics
1. [Step 2.b](#sbc_prelims): Preliminaries - The scalar wave equation in curvilinear coordinates
1. [Step 2.c](#sbc_ansatz): Sommerfeld boundary condition ansatz
1. [Step 2.d](#sbc_ansatz_dtf): Applying the ansatz to $\partial_t f$
1. [Step 2.e](#curvicoords): Implementation in generic curvilinear coordinates
1. [Step 2.e.i](#cartcoords_byhand): Sommerfeld boundary conditions implementation in Cartesian coordinates, derived by hand
1. [Step 2.e.ii](#cartcoords_bynrpysympy): Sommerfeld boundary conditions implementation in Cartesian coordinates, derived by NRPy+/SymPy
1. [Step 3](#numalg): Numerical algorithm overview
1. [Step 3.a](#class): Sommerfeld python class and parameters
1. [Step 3.b](#partial_rf): Calculate $\partial_r f$
1. [Step 3.c](#cfunc): `apply_bcs_sommerfeld()` C function
1. [Step 3.d](#k): Solving for the subdominant radial falloff proportionality constant $k$
1. [Step 3.e](#innerbcs): Inner Boundary Conditions
1. [Step 4](#py_validate): Python file validation
1. [Step 5](#interface): NRPy+ Interface for Applying Sommerfeld Boundary Conditions
1. [Step 6](#etk_validation): Validation against the [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk)
1. [Latex Output](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
<a id='initializenrpy'></a>
# Step 1: Set core NRPy+ parameters for numerical grids \[Back to [top](#toc)\]
$$\label{initializenrpy}$$
Import needed NRPy+ core modules and set working directories.
```
# Step P1: Import needed NRPy+ core modules:
import NRPy_param_funcs as par # NRPy+: Parameter interface
import reference_metric as rfm # NRPy+: Reference metric support
import grid as gri # NRPy+: Functions having to do with numerical grids
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
import shutil, os, sys # Standard Python modules for multiplatform OS-level functions
# Create C code output directory:
Ccodesdir = os.path.join("SommerfeldBoundaryCondition_Validate")
# First remove C code output directory if it exists
# Courtesy https://stackoverflow.com/questions/303200/how-do-i-remove-delete-a-folder-that-is-not-empty
shutil.rmtree(Ccodesdir, ignore_errors=True)
# Then create a fresh directory
cmd.mkdir(Ccodesdir)
par.set_parval_from_str("grid::DIM",3)
DIM = par.parval_from_str("grid::DIM")
```
<a id='sbc'></a>
# Step 2: Sommerfeld Boundary Conditions \[Back to [top](#toc)\]
$$\label{sbc}$$
<a id='intro'></a>
## Step 2.a: Introduction & background mathematics \[Back to [top](#toc)\]
$$\label{intro}$$
When we numerically solve an initial value problem, appropriate initial data must be provided, coupled to a technique for evolving the initial data forward in time (to construct the solution at $t>0$), and we must impose boundary conditions.
The subject of this notebook is implementation of Sommerfeld boundary conditions, which are often used when solving hyperbolic systems of partial differential equations (PDEs). Sommerfeld boundary conditions are also referred to as a radiation or transparent boundary conditions.
The essential idea of a transparent boundary is creating a boundary in which wave fronts can pass through with minimal reflections. In other words, the boundary condition acts to map our numerical solution to outside our numerical domain in a smooth fashion. Because this mapping is assumed to be linear, this treatment occurs at the same time-level as calculations within our numerical domain. This point will be revisited later.
Suppose we have a dynamical variable $f$; i.e., a variable in our hyperbolic system of PDEs that satisfies the equation
$$
\partial_t f = \text{something},
$$
where generally we refer to "something" as the "right-hand side" or "RHS" of the hyperbolic PDE, where we formulate the PDE such that the RHS contains no explicit time derivatives, but generally does contain spatial derivatives (typically computed in NRPy+ using finite differencing, though may in general be computed using other, e.g., pseudospectral techniques).
To construct the solution at times after the initial data, we adopt the [Method of Lines (MoL)](Tutorial-ScalarWave.ipynb) approach, which integrates the equations forward *in time* using standard explicit techniques typically used when solving *ordinary* differential equations. In doing so, MoL evaluates the RHS of the PDE at all points in our numerical domain (typically using finite difference derivatives), except the ghost zones (i.e., the gridpoints neighboring the precise point needed to evaluate a derivative using e.g., a finite difference derivative).
After each RHS evaluation we must fill in the data on the boundaries (i.e., the ghost zones), so that data exist at all gridpoints (including the boundaries) at the next MoL substep. In doing so, we have two options:
1. Perform an MoL substep to push the *interior* solution $f$ forward in time one substep, and then update the boundary values of $f$.
1. During the MoL substep, immediately after evaluating the RHS of the $\partial t f$ equation, update the boundary values of the RHS of the $\partial t f$ equation. Then push the solution $f$ forward in time *at all gridpoints, including ghost zones* by one substep.
Our implementation of the Sommerfeld boundary condition implements the second option, filling in the data for $\partial_t f$ (cf., our [extrapolation boundary condition implementation](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb)) on the boundaries (i.e., ghost zones).
<a id='sbc_prelims'></a>
## Step 2.b: Preliminaries - The scalar wave equation in curvilinear coordinates \[Back to [top](#toc)\]
$$\label{sbc_prelims}$$
Our Sommerfeld boundary condition implementation assumes $f$ behaves as a spherically symmetric, outgoing wave at the boundaries. Under these assumptions, the waves satisfy the wave equation in spherical coordinates with angular parts set to zero. I.e., under these assumptions the wavefunction $u(r,t)$ will satisfy:
\begin{align}
0 = \Box\, u &= \hat{D}^{\nu} \hat{D}_{\nu} u \\
&= \hat{g}^{\mu\nu} \hat{D}_{\mu} \hat{D}_{\nu} u \\
&= \hat{g}^{\mu\nu} \hat{D}_{\mu} \partial_{\nu} u \\
&= \hat{g}^{\mu\nu} \left[\partial_{\mu} (\partial_{\nu} u) - \hat{\Gamma}^\alpha_{\mu\nu} (\partial_{\alpha} u) \right],
\end{align}
here the hatted metric is defined as $\hat{g}^{tt} = -1/v^2$ (where $v$ is the wavespeed), $\hat{g}^{rr}=1$, $\hat{g}^{\theta\theta}=1/r^2$, and $\hat{g}^{\phi\phi}=1/(r^2\sin^2\theta)$ are the only nonzero metric terms for the spherical (contravariant) metric. However, the fact that $u=u(r,t)$ does not depend on the angular pieces greatly simplifies the expression:
\begin{align}
\Box\, u
&= \hat{g}^{\mu\nu} \left[\partial_{\mu} (\partial_{\nu} u) - \hat{\Gamma}^\alpha_{\mu\nu} (\partial_{\alpha} u) \right] \\
&= \left(-\frac{1}{v^2}\partial_t^2 + \partial_r^2\right)u - \hat{\Gamma}^\alpha_{\mu\nu} (\partial_{\alpha} u) \\
&= \left(-\frac{1}{v^2}\partial_t^2 + \partial_r^2\right)u - \hat{g}^{\mu\nu} \left[\hat{\Gamma}^t_{\mu\nu} \partial_{t} + \hat{\Gamma}^r_{\mu\nu} \partial_{r}\right]u.
\end{align}
We will now refer the reader to the [scalar wave in curvilinear coordinates notebook](Tutorial-ScalarWaveCurvilinear.ipynb) for the remainder of the derivation. The bottom line is, after removing terms implying angular variation in $u$, one obtains the wave equation for spherically symmetric waves:
$$
\frac{1}{v^2} \partial_t^2 u = \partial_r^2 u + \frac{2}{r} \partial_r u,
$$
which has the general solution
$$
u(r,t) = A \frac{u(r + vt)}{r} + B \frac{u(r - vt)}{r},
$$
where the (left) right term represents an (ingoing) outgoing wave.
<a id='sbc_ansatz'></a>
## Step 2.c: Sommerfeld boundary condition ansatz \[Back to [top](#toc)\]
$$\label{sbc_ansatz}$$
Inspired by the solution to the scalar wave equation, our Sommerfeld boundary condition will assume the solution $f(r,t)$ acts as an *outgoing* spherical wave ($A=0$), with an asymptotic value $f_0$ at $r\to\infty$ and a correction term for incoming waves or transient non-wavelike behavior at the boundaries, with $r^n$ falloff (ignoring higher-order radial falloffs):
$$
f = f_0 + \frac{u(r-vt)}{r} + \frac{c}{r^n},
$$
where $c$ is a constant.
<a id='sbc_ansatz_dtf'></a>
## Step 2.d: Applying the ansatz to $\partial_t f$ \[Back to [top](#toc)\]
$$\label{sbc_ansatz_dtf}$$
As described in the above section, we will not apply Sommerfeld boundary conditions to $f$ but $\partial_t f$ instead:
$$
\partial_t f = -v \frac{u'(r-vt)}{r}.
$$
To get a better understanding of the $u'(r-vt)$ term, let's compute the radial partial derivative as well:
\begin{align}
\partial_r f &= \frac{u'(r-vt)}{r} - \frac{u(r-vt)}{r^2} - n \frac{c}{r^{n+1}} \\
\implies \frac{u'(r-vt)}{r} &= \partial_r f + \frac{u(r-vt)}{r^2} + n \frac{c}{r^{n+1}}
\end{align}
Thus we get
\begin{align}
\partial_t f &= -v \frac{u'(r-vt)}{r} \\
&= -v \left[\partial_r f + \frac{u(r-vt)}{r^2} + n \frac{c}{r^{n+1}} \right]
\end{align}
To take care of the (as-yet) unknown $\frac{u(r-vt)}{r^2}$ term, notice our ansatz
$$
f = f_0 + \frac{u(r-vt)}{r} + \frac{c}{r^n}
$$
implies that
\begin{align}
\frac{f - f_0}{r} &= \frac{u(r-vt)}{r^2} + \frac{c}{r^{n+1}} \\
\implies \frac{u(r-vt)}{r^2} &= \frac{f - f_0}{r} - \frac{c}{r^{n+1}}
\end{align}
so we have
\begin{align}
\partial_t f &= -v \left[\partial_r f + \frac{u(r-vt)}{r^2} + n \frac{c}{r^{n+1}} \right]\\
&= -v \left[\partial_r f + \frac{f - f_0}{r} - \frac{c}{r^{n+1}} + n \frac{c}{r^{n+1}} \right] \\
&= -v \left[\partial_r f + \frac{f - f_0}{r}\right] + \frac{k}{r^{n+1}},
\end{align}
where $k=-v c(n-1)$ is just another constant.
Thus we have derived our boundary condition:
$$
\boxed{
\partial_t f = -\frac{v}{r} \left[r \partial_r f + (f - f_0)\right] + \frac{k}{r^{n+1}}.
}
$$
<a id='curvicoords'></a>
## Step 2.e: Implementation in generic curvilinear coordinates \[Back to [top](#toc)\]
$$\label{curvicoords}$$
The core equation
$$
\boxed{
\partial_t f = -\frac{v}{r} \left[r \partial_r f + (f - f_0)\right] + \frac{k}{r^{n+1}}.
}
$$
is implemented in NRPy+ using its `reference_metric.py` module ([Tutorial notebook](Tutorial-Reference_Metric.ipynb)), as this module requires *all* coordinate systems to define the spherical coordinate $r$ in terms of input quantities `(xx0,xx1,xx2)`. Thus we need only rewrite the above equation in terms of `(xx0,xx1,xx2)`. Defining $x^i$=`(xx0,xx1,xx2)`, we have, using the chain rule:
\begin{align}
\partial_t f &= -\frac{v}{r} \left[r \partial_r f + (f - f_0)\right] + \frac{k}{r^{n+1}} \\
&= -\frac{v}{r(x^i)} \left[r \frac{\partial x^i}{\partial r} \partial_i f + (f - f_0)\right] + \frac{k}{r^{n+1}}.
\end{align}
$\frac{\partial x^i}{\partial r}$ can be impossible to compute directly, as we are given $r(x^i)$ but not necessarily $x^i(r)$. The key here is to note that we are actually given $x^j_{\rm Sph} = (r(x^i),\theta(x^i),\phi(x^i))$ for all coordinate systems, so we can define the Jacobian
$$\frac{\partial x^j_{\rm Sph}(x^i)}{\partial x^i},$$
and NRPy+ can invert this matrix to give us
$$\frac{\partial x^i}{\partial x^j_{\rm Sph}},$$
In summary, the implementation of Sommerfeld boundary conditions in arbitrary curvilinear coordinates $x^i=$`(xx0,xx1,xx2)` is given by
$$
\boxed{
\partial_t f = -\frac{v}{r(x^i)} \left[r(x^i) \frac{\partial x^i}{\partial r} \partial_i f + (f - f_0)\right] + \frac{k}{r^{n+1}}.
}
$$
In the next subsections, we'll work through implementation of this general equation in the special case of Cartesian coordinates, first by hand, and then *automatically*, for Cartesian *or other curvilinear coordinate systems supported by NRPy+* using `reference_metric.py`.
<a id='cartcoords_byhand'></a>
### Step 2.e.i: Sommerfeld boundary conditions implementation in Cartesian coordinates, derived by hand \[Back to [top](#toc)\]
$$\label{cartcoords_byhand}$$
Let's now work this out for Cartesian coordinates in NRPy+, first by hand, and then using `reference_metric.py`:
In Cartesian coordinates $\frac{\partial f}{\partial r}$ may be expanded as
$$
\frac{\partial f}{\partial r} = \frac{\partial x}{\partial r} \partial_x f + \frac{\partial y}{\partial r}\partial_y f + \frac{\partial z}{\partial r}\partial_z f.
$$
Defining $x^i$ to be the $i$th component of the Cartesian reference metric, we have
\begin{align}
x^0 = x &= r\sin\theta \cos\phi \implies \frac{\partial x}{\partial r}=\frac{x}{r}, \\
x^1 = y &= r\sin\theta \sin\phi \implies \frac{\partial y}{\partial r}=\frac{y}{r}, \\
x^2 = z &= r\cos\theta \implies \frac{\partial z}{\partial r}=\frac{z}{r}.
\end{align}
Based on this, we can rewrite the above as
\begin{align}
\frac{\partial f}{\partial r}
&= \frac{x}{r} \partial_x f +\frac{y}{r} \partial_y f +\frac{z}{r} \partial_z f \\
&= \frac{x^0}{r} \partial_0 f +\frac{x^1}{r} \partial_1 f +\frac{x^2}{r} \partial_2 f \\
&= \frac{x^i}{r} \partial_i f,
\end{align}
yielding the Sommerfeld boundary condition in *Cartesian coordinates*
$$
\partial_t f = -\frac{v}{r} \left[x^i \partial_i f + \left( f - f_0 \right) \right] + \frac{k}{r^{n+1}}.
$$
<a id='cartcoords_bynrpysympy'></a>
### Step 2.e.ii: Sommerfeld boundary conditions implementation in Cartesian coordinates, derived automatically by NRPy+/SymPy \[Back to [top](#toc)\]
$$\label{cartcoords_bynrpysympy}$$
Now let's use NRPy+'s `reference_metric.py` to obtain the same expression for $\frac{\partial f}{\partial r}$ in Cartesian coordinates; i.e.,
$$
\frac{\partial f}{\partial r} = \frac{x^i}{r} \partial_i f,
$$
but using the generic coordinate system interface (`CoordSystem=Cartesian`; feel free to change it to another coordinate system of your choice). Note in the above that me we must also know the functional form of $r(x^i)$, so we use NRPy's `outputC` module to output the C code to calculate $r(x^i)$ and $\frac{\partial x^i}{\partial r} \partial_i f$, as shown below.
<a id='numalg'></a>
# Step 3: Numerical algorithm overview \[Back to [top](#toc)\]
$$\label{numalg}$$
To implement the above Sommerfeld boundary condition, we must specify for each dynamical variable:
* its wave speed at the boundary,
* its asymptotic value at infinity, and
* its $r^{n+1}$ power that must be applied for the non-wavelike behavior at the boundary.
Note in our ansatz
$$
f = f_0 + \frac{u(r-vt)}{r} + \frac{c}{r^n},
$$
in this expansion it would be natural to have $n = 2$, but in the boundary condition we have
$$
\partial_t f = -\frac{v}{r(x^i)} \left[r(x^i) \frac{\partial x^i}{\partial r} \partial_i f + (f - f_0)\right] + \frac{k}{r^{n+1}}.
$$
Thus, the exponent in the $k$ term should be 3. And indeed, in our own implementations we have found that $n=2$ exhibits the best results for minimal reflections. The [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk) documentation page also states similar results, writing "empirically, we have found that taking (exponent $ = 3$) almost completely eliminates the bad transient caused by the radiative boundary condition on its own". The following set of code cells implement the above equation, excluding the $\frac{k}{r^{n+1}}$ term, for a Cartesian grid.
Our procedure in implementing this boundary condition is as follows:
0) Define data for $f$ at **all** points in our numerical domain
1) Evaluate $\frac{df}{dt}$ at points in the **interior** only, using the prescribed equations to evaluate the RHS and ghost zones for the finite differences at the outer boundaries
2) For points in the ghost zones, apply the Sommerfeld condition to obtain $\frac{df}{dt}$ in the ghost zones, assuming advection and radial fall off behavior. When evaluating the spatial derivatives, use forward or backward finite differences for directions perpendicular to the outer boundary face being considered, and centered derivatives for the directions which lie in the plane of the outer boundary face. Furthermore, to minimize error we loop through inner boundary points and work outwards, so that the forward or backward stencils never include untreated points.
3) Account for non-wavelike evolution of our numerical solution at the boundary
4) Perform RK-update for all points (interior + ghost zones). Since the mapping to outside our numerical domain is linear, treatment of points in the interior and points on the boundary must be at the same time level.
<a id='class'></a>
## Step 3.a: Sommerfeld python class and parameters \[Back to [top](#toc)\]
$$\label{class}$$
First we define the python class __sommerfeld_boundary_condition_class__, where we store all variables and functions related to the our implementation of the Sommerfeld boundary condition. When calling the class, users can set default values for each dynamical variable's value at infinity, radial fall-off power, and wave speed at the boundary.
Next we define the function _sommerfeld_params_ which writes these parameters to lists in C, which will be used by the main C code.
```
%%writefile $Ccodesdir/Sommerfeld.py
class sommerfeld_boundary_condition_class():
"""
Class for generating C code to apply Sommerfeld boundary conditions
"""
# class variables should be the resulting dicts
# Set class variable default values
# radial falloff power n = 3 has been found to yield the best results
# - see Tutorial-SommerfeldBoundaryCondition.ipynb Step 2 for details
def __init__(self, fd_order=2, vars_at_inf_default = 0., vars_radial_falloff_power_default = 3., vars_speed_default = 1.):
evolved_variables_list, _, _ = gri.gridfunction_lists()
# set class finite differencing order
self.fd_order = fd_order
NRPy_FD_order = par.parval_from_str("finite_difference::FD_CENTDERIVS_ORDER")
if NRPy_FD_order < fd_order:
print("ERROR: The global central finite differencing order within NRPy+ must be greater than or equal to the Sommerfeld boundary condition's finite differencing order")
sys.exit(1)
# Define class dictionaries to store sommerfeld parameters for each EVOL gridfunction
# EVOL gridfunction asymptotic value at infinity
self.vars_at_infinity = {}
# EVOL gridfunction wave speed at outer boundaries
self.vars_speed = {}
# EVOL gridfunction radial falloff power
self.vars_radial_falloff_power = {}
# Set default values for each specific EVOL gridfunction
for gf in evolved_variables_list:
self.vars_at_infinity[gf.upper() + 'GF'] = vars_at_inf_default
self.vars_radial_falloff_power[gf.upper() + 'GF'] = vars_radial_falloff_power_default
self.vars_speed[gf.upper() + 'GF'] = vars_speed_default
def sommerfeld_params(self):
# Write parameters to C file
# Creating array for EVOL gridfunction values at infinity
var_at_inf_string = "{"
for _gf,val in self.vars_at_infinity.items():
var_at_inf_string += str(val) + ", "
var_at_inf_string = var_at_inf_string[:-2] + "};"
# Creating array for EVOL gridfunction values of radial falloff power
vars_radial_falloff_power_string = "{"
for _gf,val in self.vars_radial_falloff_power.items():
vars_radial_falloff_power_string += str(val) + ", "
vars_radial_falloff_power_string = vars_radial_falloff_power_string[:-2] + "};"
# Creating array for EVOL gridfunction values of wave speed at outer boundaries
var_speed_string = "{"
for _gf,val in self.vars_speed.items():
var_speed_string += str(val) + ", "
var_speed_string = var_speed_string[:-2] + "};"
# Writing to values to sommerfeld_params.h file
out_str = """
// Sommerfeld EVOL grid function parameters
const REAL evolgf_at_inf[NUM_EVOL_GFS] = """+var_at_inf_string+"""
const REAL evolgf_radial_falloff_power[NUM_EVOL_GFS] = """+vars_radial_falloff_power_string+"""
const REAL evolgf_speed[NUM_EVOL_GFS] = """+var_speed_string+"""
"""
return out_str
```
<a id='partial_rf'></a>
## Step 3.b: Calculate $\partial_r f$ \[Back to [top](#toc)\]
$$\label{partial_rf}$$
Next we generate the C code for calculating $\partial_r f$ for each dynamical variable $f$ in our coordinate system of choice.
```
%%writefile -a $Ccodesdir/Sommerfeld.py
@staticmethod
def dfdr_function(fd_order):
# function to write c code to calculate dfdr term in Sommerfeld boundary condition
# Read what # of dimensions being used
DIM = par.parval_from_str("grid::DIM")
# Set up the chosen reference metric from chosen coordinate system, set within NRPy+
CoordSystem = par.parval_from_str("reference_metric::CoordSystem")
rfm.reference_metric()
# Simplifying the results make them easier to interpret.
do_simplify = True
if "Sinh" in CoordSystem:
# Simplification takes too long on Sinh* coordinate systems
do_simplify = False
# Construct Jacobian matrix, output Jac_dUSph_dDrfmUD[i][j] = \partial x_{Sph}^i / \partial x^j:
Jac_dUSph_dDrfmUD = ixp.zerorank2()
for i in range(3):
for j in range(3):
Jac_dUSph_dDrfmUD[i][j] = sp.diff(rfm.xxSph[i],rfm.xx[j])
# Invert Jacobian matrix, output to Jac_dUrfm_dDSphUD.
Jac_dUrfm_dDSphUD, dummyDET = ixp.generic_matrix_inverter3x3(Jac_dUSph_dDrfmUD)
# Jac_dUrfm_dDSphUD[i][0] stores \partial x^i / \partial r
if do_simplify:
for i in range(3):
Jac_dUrfm_dDSphUD[i][0] = sp.simplify(Jac_dUrfm_dDSphUD[i][0])
# Declare \partial_i f, which is actually computed later on
fdD = ixp.declarerank1("fdD") # = [fdD0, fdD1, fdD2]
contraction = sp.sympify(0)
for i in range(3):
contraction += fdD[i]*Jac_dUrfm_dDSphUD[i][0]
if do_simplify:
contraction = sp.simplify(contraction)
r_str_and_contraction_str = outputC([rfm.xxSph[0],contraction],
["*_r","*_partial_i_f"],filename="returnstring",params="includebraces=False")
```
Here we generate the C code used to calculate all relevant spatial derivatives $\partial_i f$, using second order accurate finite differences. Specifically, if our ghost zone point lies on one of the faces, on an edge or corner, we use forward or backward differences depending on the specific direction, and centered differences otherwise. Note that all derivatives along the normal of the boundary faces are forward or backward, to minimize using non-updated points in the derivative calculations.
For example, consider some point with Cartesian coordinates $(i,j,k)$ on our grid, the derivative of $f$ along the $x$ direction will be the forward (backward with change of signs on coefficients)
$$
\frac{\partial f_{ijk}}{\partial x} \approx \frac{1}{2\Delta x} \left( -3f_{i,j,k} + 4f_{i+1,j,k} - f_{i+2,j,k} \right),
$$
or the centered difference approximation
$$
\frac{\partial f_{ijk}}{\partial x} \approx \frac{1}{2\Delta x} \left( f_{i+1,j,k} - f_{i-1,j,k} \right).
$$
We determine the signs of the coefficients (corresponding to using either a forward or backward difference) by determining what face the point lies within. The above is applied for all three Cartesian directions. Note the use if the `SHIFTSTENCIL` variable, which helps determine when to use forward/backward difference to take derivatives along normals to boundary faces, or when to use central differences to either take derivatives parallel to the faces or at points on edges and corners.
```
%%writefile -a $Ccodesdir/Sommerfeld.py
def gen_central_2oFD_stencil_str(intdirn):
if intdirn == 0:
return "(gfs[IDX4S(which_gf,i0+1,i1,i2)]-gfs[IDX4S(which_gf,i0-1,i1,i2)])*0.5" # Does not include the 1/dx multiplication
if intdirn == 1:
return "(gfs[IDX4S(which_gf,i0,i1+1,i2)]-gfs[IDX4S(which_gf,i0,i1-1,i2)])*0.5" # Does not include the 1/dy multiplication
return "(gfs[IDX4S(which_gf,i0,i1,i2+1)]-gfs[IDX4S(which_gf,i0,i1,i2-1)])*0.5" # Does not include the 1/dz multiplication
def gen_central_4oFD_stencil_str(intdirn):
if intdirn == 0:
return """(-c2*gfs[IDX4S(which_gf,i0+2,i1,i2)]
+c1*gfs[IDX4S(which_gf,i0+1,i1,i2)]
-c1*gfs[IDX4S(which_gf,i0-1,i1,i2)]
+c2*gfs[IDX4S(which_gf,i0-2,i1,i2)])""" # Does not include the 1/dx multiplication
if intdirn == 1:
return """(-c2*gfs[IDX4S(which_gf,i0,i1+2,i2)]
+c1*gfs[IDX4S(which_gf,i0,i1+1,i2)]
-c1*gfs[IDX4S(which_gf,i0,i1-1,i2)]
+c2*gfs[IDX4S(which_gf,i0,i1-2,i2)])""" # Does not include the 1/dy multiplication
return """(-c2*gfs[IDX4S(which_gf,i0,i1,i2+2)]
+c1*gfs[IDX4S(which_gf,i0,i1,i2+1)]
-c1*gfs[IDX4S(which_gf,i0,i1,i2-1)]
+c2*gfs[IDX4S(which_gf,i0,i1,i2-2)])""" # Does not include the 1/dz multiplication
def gen_central_6oFD_stencil_str(intdirn):
if intdirn == 0:
return """( c3*gfs[IDX4S(which_gf,i0+3,i1,i2)]
-c2*gfs[IDX4S(which_gf,i0+2,i1,i2)]
+c1*gfs[IDX4S(which_gf,i0+1,i1,i2)]
-c1*gfs[IDX4S(which_gf,i0-1,i1,i2)]
+c2*gfs[IDX4S(which_gf,i0-2,i1,i2)]
-c3*gfs[IDX4S(which_gf,i0-3,i1,i2)])""" # Does not include the 1/dx multiplication
if intdirn == 1:
return """( c3*gfs[IDX4S(which_gf,i0,i1+3,i2)]
-c2*gfs[IDX4S(which_gf,i0,i1+2,i2)]
+c1*gfs[IDX4S(which_gf,i0,i1+1,i2)]
-c1*gfs[IDX4S(which_gf,i0,i1-1,i2)]
+c2*gfs[IDX4S(which_gf,i0,i1-2,i2)]
-c3*gfs[IDX4S(which_gf,i0,i1-3,i2)])""" # Does not include the 1/dy multiplication
return """( c3*gfs[IDX4S(which_gf,i0,i1,i2+3)]
-c2*gfs[IDX4S(which_gf,i0,i1,i2+2)]
+c1*gfs[IDX4S(which_gf,i0,i1,i2+1)]
-c1*gfs[IDX4S(which_gf,i0,i1,i2-1)]
+c2*gfs[IDX4S(which_gf,i0,i1,i2-2)]
-c3*gfs[IDX4S(which_gf,i0,i1,i2-3)])""" # Does not include the 1/dz multiplication
def gen_central_fd_stencil_str(intdirn, fd_order):
if fd_order==2:
return gen_central_2oFD_stencil_str(intdirn)
if fd_order==4:
return gen_central_4oFD_stencil_str(intdirn)
return gen_central_6oFD_stencil_str(intdirn)
def output_dfdx(intdirn, fd_order):
dirn = str(intdirn)
dirnp1 = str((intdirn+1)%3) # if dirn='0', then we want this to be '1'; '1' then '2'; and '2' then '0'
dirnp2 = str((intdirn+2)%3) # if dirn='0', then we want this to be '2'; '1' then '0'; and '2' then '1'
preface = """
// On a +x"""+dirn+""" or -x"""+dirn+""" face, do up/down winding as appropriate:
if(abs(FACEXi["""+dirn+"""])==1 || i"""+dirn+"""+NGHOSTS >= Nxx_plus_2NGHOSTS"""+dirn+""" || i"""+dirn+"""-NGHOSTS <= 0) {
int8_t SHIFTSTENCIL"""+dirn+""" = FACEXi["""+dirn+"""];
if(i"""+dirn+"""+NGHOSTS >= Nxx_plus_2NGHOSTS"""+dirn+""") SHIFTSTENCIL"""+dirn+""" = -1;
if(i"""+dirn+"""-NGHOSTS <= 0) SHIFTSTENCIL"""+dirn+""" = +1;
SHIFTSTENCIL"""+dirnp1+""" = 0;
SHIFTSTENCIL"""+dirnp2+""" = 0;
"""
if fd_order == 2:
return preface + """
fdD"""+dirn+"""
= SHIFTSTENCIL"""+dirn+"""*(-1.5*gfs[IDX4S(which_gf,i0+0*SHIFTSTENCIL0,i1+0*SHIFTSTENCIL1,i2+0*SHIFTSTENCIL2)]
+2.*gfs[IDX4S(which_gf,i0+1*SHIFTSTENCIL0,i1+1*SHIFTSTENCIL1,i2+1*SHIFTSTENCIL2)]
-0.5*gfs[IDX4S(which_gf,i0+2*SHIFTSTENCIL0,i1+2*SHIFTSTENCIL1,i2+2*SHIFTSTENCIL2)]
)*invdx"""+dirn+""";
// Not on a +x"""+dirn+""" or -x"""+dirn+""" face, using centered difference:
} else {
fdD"""+dirn+""" = """+gen_central_fd_stencil_str(intdirn, 2)+"""*invdx"""+dirn+""";
}
"""
if fd_order == 4:
return preface + """
fdD"""+dirn+"""
= SHIFTSTENCIL"""+dirn+"""*(u0*gfs[IDX4S(which_gf,i0+0*SHIFTSTENCIL0,i1+0*SHIFTSTENCIL1,i2+0*SHIFTSTENCIL2)]
+u1*gfs[IDX4S(which_gf,i0+1*SHIFTSTENCIL0,i1+1*SHIFTSTENCIL1,i2+1*SHIFTSTENCIL2)]
+u2*gfs[IDX4S(which_gf,i0+2*SHIFTSTENCIL0,i1+2*SHIFTSTENCIL1,i2+2*SHIFTSTENCIL2)]
+u3*gfs[IDX4S(which_gf,i0+3*SHIFTSTENCIL0,i1+3*SHIFTSTENCIL1,i2+3*SHIFTSTENCIL2)]
+u4*gfs[IDX4S(which_gf,i0+4*SHIFTSTENCIL0,i1+4*SHIFTSTENCIL1,i2+4*SHIFTSTENCIL2)]
)*invdx"""+dirn+""";
// Not on a +x"""+dirn+""" or -x"""+dirn+""" face, using centered difference:
} else {
fdD"""+dirn+""" = """+gen_central_fd_stencil_str(intdirn, 4)+"""*invdx"""+dirn+""";
}
"""
if fd_order == 6:
return preface + """
fdD"""+dirn+"""
= SHIFTSTENCIL"""+dirn+"""*(u0*gfs[IDX4S(which_gf,i0+0*SHIFTSTENCIL0,i1+0*SHIFTSTENCIL1,i2+0*SHIFTSTENCIL2)]
+u1*gfs[IDX4S(which_gf,i0+1*SHIFTSTENCIL0,i1+1*SHIFTSTENCIL1,i2+1*SHIFTSTENCIL2)]
+u2*gfs[IDX4S(which_gf,i0+2*SHIFTSTENCIL0,i1+2*SHIFTSTENCIL1,i2+2*SHIFTSTENCIL2)]
+u3*gfs[IDX4S(which_gf,i0+3*SHIFTSTENCIL0,i1+3*SHIFTSTENCIL1,i2+3*SHIFTSTENCIL2)]
+u4*gfs[IDX4S(which_gf,i0+4*SHIFTSTENCIL0,i1+4*SHIFTSTENCIL1,i2+4*SHIFTSTENCIL2)]
+u5*gfs[IDX4S(which_gf,i0+5*SHIFTSTENCIL0,i1+5*SHIFTSTENCIL1,i2+5*SHIFTSTENCIL2)]
+u6*gfs[IDX4S(which_gf,i0+6*SHIFTSTENCIL0,i1+6*SHIFTSTENCIL1,i2+6*SHIFTSTENCIL2)]
)*invdx"""+dirn+""";
// Not on a +x"""+dirn+""" or -x"""+dirn+""" face, using centered difference:
} else {
fdD"""+dirn+""" = """+gen_central_fd_stencil_str(intdirn, 6)+"""*invdx"""+dirn+""";
}
"""
print("Error: fd_order = "+str(fd_order)+" currently unsupported.")
sys.exit(1)
contraction_term_func = """
// Function to calculate the radial derivative of a grid function
void contraction_term(const paramstruct *restrict params, const int which_gf, const REAL *restrict gfs, REAL *restrict xx[3],
const int8_t FACEXi[3], const int i0, const int i1, const int i2, REAL *restrict _r, REAL *restrict _partial_i_f) {
#include "RELATIVE_PATH__set_Cparameters.h" /* Header file containing correct #include for set_Cparameters.h;
* accounting for the relative path */
// Initialize derivatives to crazy values, to ensure that
// we will notice in case they aren't set properly.
REAL fdD0=1e100;
REAL fdD1=1e100;
REAL fdD2=1e100;
REAL xx0 = xx[0][i0];
REAL xx1 = xx[1][i1];
REAL xx2 = xx[2][i2];
int8_t SHIFTSTENCIL0;
int8_t SHIFTSTENCIL1;
int8_t SHIFTSTENCIL2;
"""
if fd_order == 4:
contraction_term_func +="""
// forward/backward finite difference coefficients
const REAL u0 =-25./12.;
const REAL u1 = 4.;
const REAL u2 = -3.;
const REAL u3 = 4./3.;
const REAL u4 = -1./4.;
// central finite difference coefficients
const REAL c1 = 2./3.;
const REAL c2 = 1./12.;
"""
if fd_order == 6:
contraction_term_func +="""
// forward/backward finite difference coefficients
const REAL u0 = -49./20.;
const REAL u1 = 6.;
const REAL u2 = -15./2.;
const REAL u3 = 20./3.;
const REAL u4 = -15./4.;
const REAL u5 = 6./5.;
const REAL u6 = -1./6.;
// central finite difference coefficients
const REAL c1 = 3./4.;
const REAL c2 = 3./20.;
const REAL c3 = 1./60;
"""
for i in range(DIM):
if "fdD"+str(i) in r_str_and_contraction_str:
contraction_term_func += output_dfdx(i, fd_order)
contraction_term_func += "\n" + r_str_and_contraction_str
contraction_term_func +="""
} // END contraction_term function
"""
return contraction_term_func
```
<a id='cfunc'></a>
## Step 3.c: `apply_bcs_sommerfeld()` C function \[Back to [top](#toc)\]
$$\label{cfunc}$$
Here, we build up the main C code and define the function `apply_bcs_sommerfeld()` to be used by NRPy's MoL time stepping algorithm.
```
%%writefile -a $Ccodesdir/Sommerfeld.py
def write_sommerfeld_main_Ccode(self, Ccodesdir):
main_Ccode = """
// Boundary condition driver routine: Apply BCs to all
// boundary faces of the 3D numerical domain, filling in the
// outer boundary ghost zone layers, starting with the innermost
// layer and working outward.
"""
main_Ccode += self.sommerfeld_params()
main_Ccode += self.dfdr_function(self.fd_order)
main_Ccode += """
void apply_bcs_sommerfeld(const paramstruct *restrict params, REAL *restrict xx[3],
const bc_struct *restrict bcstruct, const int NUM_GFS,
const int8_t *restrict gfs_parity, REAL *restrict gfs,
REAL *restrict rhs_gfs) {
#pragma omp parallel for
for(int which_gf=0;which_gf<NUM_GFS;which_gf++) {
const REAL char_speed = evolgf_speed[which_gf];
const REAL var_at_infinity = evolgf_at_inf[which_gf];
const REAL radial_falloff_power = evolgf_radial_falloff_power[which_gf];
#include "RELATIVE_PATH__set_Cparameters.h" /* Header file containing correct #include for set_Cparameters.h;
* accounting for the relative path */
for(int which_gz = 0; which_gz < NGHOSTS; which_gz++) {
for(int pt=0;pt<bcstruct->num_ob_gz_pts[which_gz];pt++) {
const int i0 = bcstruct->outer[which_gz][pt].outer_bc_dest_pt.i0;
const int i1 = bcstruct->outer[which_gz][pt].outer_bc_dest_pt.i1;
const int i2 = bcstruct->outer[which_gz][pt].outer_bc_dest_pt.i2;
const int8_t FACEX0 = bcstruct->outer[which_gz][pt].FACEi0;
const int8_t FACEX1 = bcstruct->outer[which_gz][pt].FACEi1;
const int8_t FACEX2 = bcstruct->outer[which_gz][pt].FACEi2;
const int8_t FACEXi[3] = {FACEX0, FACEX1, FACEX2};
// Initialize derivatives to crazy values, to ensure that
// we will notice in case they aren't set properly.
REAL r = 1e100;
REAL partial_i_f = 1e100;
```
Finally, we calculate $\frac{df}{dt}$ without the $\frac{k}{r^{n+1}}$ term;
$$
\frac{\partial f}{\partial t} = -\frac{v}{r(x^i)} \left[r(x^i) \frac{\partial x^i}{\partial r} \partial_i f + (f - f_0)\right].
$$
```
%%writefile -a $Ccodesdir/Sommerfeld.py
contraction_term(params, which_gf, gfs, xx, FACEXi, i0, i1, i2, &r, &partial_i_f);
const REAL invr = 1./r;
const REAL source_rhs = -char_speed*(partial_i_f + invr*(gfs[IDX4S(which_gf,i0,i1,i2)] - var_at_infinity));
rhs_gfs[IDX4S(which_gf,i0,i1,i2)] = source_rhs;
```
<a id='k'></a>
## Step 3.d: Solving for $k$ \[Back to [top](#toc)\]
$$\label{k}$$
Here we formulate a way to approximate $k$. If our solution satisfies the advection equation both at the ghost zones and at interior points close to the boundaries, then we may find $k$ by determining the portion of $f$ that is not accounted for by the equation
$$
\frac{\partial f}{\partial t} = \left[\frac{\partial f}{\partial t} \right]_{adv} = -\frac{v}{r(x^i)} \left[r(x^i) \frac{\partial x^i}{\partial r} \partial_i f + (f - f_0)\right].
$$
The above is the advection equation we arrive at assuming $f$ behaves purely as an outgoing spherical wave. For an interior point directly adjacent to a ghost zone point, $f$ must satisfy **both** the time evolution equation for prescribed points within the interior, $\left[\frac{\partial f}{\partial t} \right]_{evol}$, and the advection equation $\left[\frac{\partial f}{\partial t} \right]_{adv}$. We then find the difference as
$$
\delta = \left[\frac{\partial f}{\partial t} \right]_{evol} - \left[\frac{\partial f}{\partial t} \right]_{adv} = \frac{k}{r^{n+1}},
$$
i.e. $\delta$ represents the numerical departure from the expected purely wave-like behavior at that point. We solve for $\delta$ at this interior point and express $k$ as
$$
k = \delta r^{n+1}_{int}.
$$
Thus, the $\frac{k}{r^{n+1}}$ term for the associated ghost zone point may be expressed as
$$
\frac{k}{r^{n+1}_{gz}} = \delta \left( \frac{r_{int}}{r_{gz}} \right) ^{n+1}.
$$
We approximate $k$ in this fashion using the code below. Note that we activate this term only when *radial_falloff_power* > 0, which set to 3 by default.
```
%%writefile -a $Ccodesdir/Sommerfeld.py
/************* For radial falloff and the extrapolated k term *************/
if (radial_falloff_power > 0) {
// Move one point away from gz point to compare pure advection to df/dt|interior
const int i0_offset = i0+FACEX0;
const int i1_offset = i1+FACEX1;
const int i2_offset = i2+FACEX2;
// Initialize derivatives to crazy values, to ensure that
// we will notice in case they aren't set properly.
REAL r_offset = 1e100;
REAL partial_i_f_offset = 1e100;
contraction_term(params, which_gf, gfs, xx, FACEXi, i0_offset, i1_offset, i2_offset, &r_offset, &partial_i_f_offset);
const REAL invr_offset = 1./r_offset;
// Pure advection: [FIXME: Add equation (appearing in Jupyter notebook documentation)]
const REAL extrap_rhs = char_speed*(partial_i_f_offset + invr_offset*(gfs[IDX4S(which_gf,i0_offset,i1_offset,i2_offset)] - var_at_infinity));
// Take difference between pure advection and df/dt|interior
const REAL diff_between_advection_and_f_rhs =
rhs_gfs[IDX4S(which_gf,i0_offset,i1_offset,i2_offset)] + extrap_rhs;
// Solve for k/(r_gz)^n+1 term
rhs_gfs[IDX4S(which_gf,i0,i1,i2)] += diff_between_advection_and_f_rhs*pow(r_offset*invr,radial_falloff_power);
}
} // END for(int pt=0;pt<num_ob_gz_pts[which_gz];pt++)
```
<a id='innerbcs'></a>
## Step 3.e: Inner Boundary Conditions \[Back to [top](#toc)\]
$$\label{innerbcs}$$
Finally, we apply parity conditions for inner boundary conditions. Since the Sommerfeld boundary condition treats the right hand sides, these data are thus copied over to the right hand sides of points in the inner boundaries, according to appropriate parity conditions. For a detailed discussion on inner boundaries and parity conditions, see [Tutorial-Start_to_Finish-Curvilinear_BCs](Tutorial-Start_to_Finish-Curvilinear_BCs.ipynb).
```
%%writefile -a $Ccodesdir/Sommerfeld.py
// Apply INNER (parity) boundary conditions:
for(int pt=0;pt<bcstruct->num_ib_gz_pts[which_gz];pt++) {
const int i0dest = bcstruct->inner[which_gz][pt].inner_bc_dest_pt.i0;
const int i1dest = bcstruct->inner[which_gz][pt].inner_bc_dest_pt.i1;
const int i2dest = bcstruct->inner[which_gz][pt].inner_bc_dest_pt.i2;
const int i0src = bcstruct->inner[which_gz][pt].inner_bc_src_pt.i0;
const int i1src = bcstruct->inner[which_gz][pt].inner_bc_src_pt.i1;
const int i2src = bcstruct->inner[which_gz][pt].inner_bc_src_pt.i2;
rhs_gfs[IDX4S(which_gf,i0dest,i1dest,i2dest)] =
bcstruct->inner[which_gz][pt].parity[gfs_parity[which_gf]] * rhs_gfs[IDX4S(which_gf, i0src,i1src,i2src)];
} // END for(int pt=0;pt<num_ib_gz_pts[which_gz];pt++)
} // END for(int which_gz = 0; which_gz < NGHOSTS; which_gz++)
} // END for(int which_gf=0;which_gf<NUM_GFS;which_gf++)
} // END function
"""
```
Here we write the entire function to a C file.
```
%%writefile -a $Ccodesdir/Sommerfeld.py
with open(os.path.join(Ccodesdir,"boundary_conditions/apply_bcs_sommerfeld.h"),"w") as file:
file.write(main_Ccode)
def write_sommerfeld_file(self, Ccodesdir):
self.write_sommerfeld_main_Ccode(Ccodesdir)
print("""\nSuccessfully generated Sommerfeld boundary condition C code""")
```
<a id='py_validate'></a>
# Step 4: Python file validation \[Back to [top](#toc)\]
$$\label{py_validate}$$
Here we validate the python code generated by this notebook, [SommerfeldBoundaryCondition_Validate/Sommerfeld.py](../edit/SommerfeldBoundaryCondition_Validate/Sommerfeld.py), against the trusted code in [CurviBoundaryConditions/CurviBoundaryConditions.py](../edit/CurviBoundaryConditions/CurviBoundaryConditions.py) line by line. Passing corresponds to complete agreement between the files.
Note that there is more content [CurviBoundaryConditions/CurviBoundaryConditions.py](../edit/CurviBoundaryConditions/CurviBoundaryConditions.py) relating to more than just the Sommerfeld boundary condition, so we start the comparison where the Sommerfeld code begins.
```
# Then compare all files generated by this notebook
# (output moved in previous code cell to validate/)
# and the separate Python module (output to Baikal
# and BaikalVacuum).
import difflib
def compare_two_files(filepath1,filepath2, file1_idx1=None):
with open(filepath1) as file1, open(filepath2) as file2:
# Read the lines of each file
file1_lines = file1.readlines()
file2_lines = file2.readlines()
if file1_idx1!=None:
file1_lines = file1_lines[file1_idx1:]
# print(file1_lines)
num_diffs = 0
file1_lines_noleadingwhitespace = []
for line in file1_lines:
if line.strip() == "": # If the line contains only whitespace, remove all leading whitespace
file1_lines_noleadingwhitespace.append(line.lstrip())
else:
file1_lines_noleadingwhitespace.append(line)
file2_lines_noleadingwhitespace = []
for line in file2_lines:
if line.strip() == "": # If the line contains only whitespace, remove all leading whitespace
file2_lines_noleadingwhitespace.append(line.lstrip())
else:
file2_lines_noleadingwhitespace.append(line)
for line in difflib.unified_diff(file1_lines_noleadingwhitespace, file2_lines_noleadingwhitespace,
fromfile=filepath1,
tofile =filepath2):
sys.stdout.writelines(line)
num_diffs = num_diffs + 1
if num_diffs == 0:
print("PASSED: "+filepath2+" matches trusted version")
else:
print("FAILED (see diff above): "+filepath2+" does NOT match trusted version")
import os
notebook_cfile = 'SommerfeldBoundaryCondition_Validate/Sommerfeld.py'
nrpy_cfile = 'CurviBoundaryConditions/CurviBoundaryConditions.py'
idx1=258
compare_two_files(nrpy_cfile, notebook_cfile, idx1)
```
<a id='interface'></a>
# Step 5: NRPy+ Interface for Applying Sommerfeld Boundary Conditions \[Back to [top](#toc)\]
$$\label{interface}$$
To apply the Sommerfeld boundary condition to any given grid function, its wave speed at the boundaries, asymptotic value at infinity, and radial exponent of the k term (*radial_falloff_power*) must be specified. In general, a (*radial_falloff_power*) of 3 has been found to yield the best results, i.e. minimal initial transients and reflections.
Here we showcase the features of the NRPy+ interface for implementing this boundary condition and defining these values. The interface is a python class structure that allows the user to specify default values for each grid function, which may then be changed. This may be useful when the user wants to define the default values for several grid functions but wishes to specifically alter few others.
To begin, we define our global NRPy finite differencing order and our coordinate system of choice. Our Sommerfeld boundary condition driver check that the finite differencing chosen for the driver is less than or equal to the global finite differencing order.
```
import finite_difference as fin # NRPy+: Finite difference C code generation module
FD_order = 6
par.set_parval_from_str("finite_difference::FD_CENTDERIVS_ORDER",FD_order)
# Set the coordinate system for the numerical grid
# Choices are: Spherical, SinhSpherical, SinhSphericalv2, Cylindrical, SinhCylindrical,
# SymTP, SinhSymTP
CoordSystem = "Spherical"
par.set_parval_from_str("reference_metric::CoordSystem",CoordSystem)
```
Next, let's define a few grid functions. Note that the boundary condition should be used only for variables defined in three spatial dimensions.
```
# Step P3: Defining a couple of grid functions
uu, vv, ww, xx, yy, zz, = gri.register_gridfunctions("EVOL",["uu","vv","ww","xx","yy","zz"])
```
NRPy+ can now access these grid function names, and will store default values for the boundary condition. First we import `CurviBoundaryConditions.CurviBoundaryConditions`, and define a class instance _bcs_ using `sommerfeld_boundary_condition_class`.
```
import CurviBoundaryConditions.CurviBoundaryConditions as cbcs
cbcs.Set_up_CurviBoundaryConditions(os.path.join(Ccodesdir,"boundary_conditions/"),
Cparamspath=os.path.join("../"), BoundaryCondition='Sommerfeld')
bcs = cbcs.sommerfeld_boundary_condition_class(fd_order=4,
vars_radial_falloff_power_default=3,
vars_speed_default=1.,
vars_at_inf_default=1.)
# bcs.vars_radpower.items()
bcs.vars_at_infinity['VVGF'] = 0.0
bcs.write_sommerfeld_file(Ccodesdir)
```
Using the instance _bcs_ we may change these default values using the grid function names, since NRPy+ stores these values in python dictionaries. We then print out the contents of the dictionaries.
```
# Changing values for uu
bcs.vars_at_infinity['UUGF'] = 5.
bcs.vars_speed['UUGF'] = 0.5
# Changing values for zz
bcs.vars_at_infinity['ZZGF'] = 4.
bcs.vars_speed['ZZGF'] = 2.**0.5
print('GF values at infinity =', bcs.vars_at_infinity.items())
print('GF speeds = ', bcs.vars_speed.items())
print('GF radial powers = ' ,bcs.vars_radial_falloff_power.items())
```
Finally, we write these values into the `sommerfeld_params.h` file, and generate the `radial_derivative.h` file, which defines the function used to calculate the $\partial_r f$ term, which our C code may read from later, using the function write_sommerfeld_files(), which takes the C codes directory path and finite differencing order as inputs. __Only second and fourth order finite differences are supported at this time.__
<a id='etk_validation'></a>
# Step 6: Validation against the [Einstein Toolkit's NewRad boundary condition driver](https://www.einsteintoolkit.org/EinsteinEvolve/thornguide/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk) \[Back to [top](#toc)\]
$$\label{etk_validation}$$
Here we showcase some validation results of our Sommerfeld boundary condition as implemented in NRPy+ against [ETK's NewRad boundary condition driver](https://www.einsteintoolkit.org/thornguide/EinsteinEvolve/NewRad/documentation.html#XEinsteinEvolve_NewRad_Alcubierre:2002kk), storing the ETK data and subsequent plots in the [SommerfeldBoundaryCondition folder](SommerfeldBoundaryCondition). Specifically, we do so by
1. Generating plane wave initial data for ETK using the [ETK_thorn-IDScalarWaveNRPy notebook](Tutorial-ETK_thorn-IDScalarWaveNRPy.ipynb), with the NRPy+ code generation documented [here](Tutorial-ScalarWave.ipynb)
2. Generating the ETK evolution C codes using the [ETK_thorn-WaveToyNRPy notebook](Tutorial-ETK_thorn-WaveToyNRPy.ipynb), with the NRPy+ code generation generation also documented [here](Tutorial-ScalarWave.ipynb)
3. Compare results to [Tutorial-Start_to_Finish-ScalarWaveCurvilinear](Tutorial-Start_to_Finish-ScalarWaveCurvilinear.ipynb), adding 1e-16 to the log$_{10}$(relative error) plots to avoid taking the log$_{10}$ of zero where we have perfect agreement. $t$=0.3 represents 6 time steps into the systems
For both the evolution thorn and NRPy+ code we define the gridfunctions __UUGF__ and __VVGF__, use RK4 time stepping, and fourth order finite differencing. For the boundary condition parameters, we set uu_at_infinity = 2.0, vv_at_infinity = 0.0, and char_speed = 1.0, and var_radial_falloff_power = 3.0 for both evolution variables.
First we show validation results for the case of a scalar wave propagating in the +x direction (initial data, documented [here](Tutorial-ScalarWave.ipynb), with kk0 = 1, kk1 = kk2 = 0), overlaying $u\left(x, y=0,z=0,t=0.3\right)$ from the ETK thorn and from NRPy+, and plotting the relative difference between the two.
```
from IPython.display import Image
from IPython.display import display
x_axis_plot = Image("SommerfeldBoundaryCondition/NRPy_vs_ETK_x-axis.png", width=400, height=400)
E_relx_axis_plot = Image("SommerfeldBoundaryCondition/E_rel_x-axis.png", width=400, height=400)
display(x_axis_plot,E_relx_axis_plot)
```
Lastly we show validation results for the case of a scalar wave propagating along the -x, -y, +z diagonal, and taking a slice along the y-axis at x=z=0 (initial data, documented [here](Tutorial-ScalarWave.ipynb), with kk0 = -1, kk1 = -1, kk2 = 1), overlaying $u\left(x=0, y,z=0,t=0.3\right)$ from the ETK thorn and from NRPy+, and plotting the relative difference between the two.
```
diagonal = Image("SommerfeldBoundaryCondition/NRPy_vs_ETK_diagonal.png", width=400, height=400)
E_rel_diagonal = Image("SommerfeldBoundaryCondition/E_rel_diagonal.png", width=400, height=400)
display(diagonal, E_rel_diagonal)
```
<a id='latex_pdf_output'></a>
# Step 7: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
$$\label{latex_pdf_output}$$
The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
[Tutorial-SommerfeldBoundaryCondition.pdf](Tutorial-SommerfeldBoundaryCondition.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
```
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-SommerfeldBoundaryCondition")
```
| github_jupyter |
# Anaconda Installation Instructions
## Installation
A recommended way to install python and its various modules, packages, and libraries is to install it as part of the Anaconda distribution.
Most users will find this distribution convenient. Once you've installed Anaconda, the
python package will be available to you. In other words, you do not have to install python from the python.org website.
Go to the Anaconda distribution website: https://www.anaconda.com/distribution/
Click on "download" for Python 3.7 version. Follow the instructions for the Anaconda3 Installer package.
The graphical installer will prompt you to click "continue" after you see ReadMe and Software License Agreement.
You will need to enter your own password for your machine to install software. The directory anaconda3 will be located in your opt directory when installed.
## Checking Your Installation
Open up a bash shell. Change directory to the location of where you want to work.
Type "jupyter notebook" in your shell and hit enter. Wait for a new browser tab to open up at http://localhost:8889/tree . At the home page of jupyter notebook in the web browser,
look to the upper right for "New" button with a down arrow. Click on it and select "Python 3".
This will bring up a new notebook. Click on "File" and select "rename". Type in a new notebook name to save it.
In a new cell, type: "import pandas as pd" and hit shift + enter to execute cell. At the second line, type "import numpy as np" and hit shift + enter. On the third line, print("Hello World!") and hit shift + enter. If there are no errors, then your installation was successful.
Open up a bash shell. To see the version of python that you have, type "python -V".
To see the version of Anaconda that you have, type "conda -V". To see the packages that come with Anaconda, type "conda list".
To upgrade Anaconda to a newer version, use "conda update conda".
For a specific version, use "conda update anaconda=VersionNumber". An example would be, "conda update anaconda=2019.10".
To downgrade to a particular version of python, use "conda search python" to see all your version options. You can then use "conda install python=3.6.4", for instance.
To uninstall Anaconda, type "rm -rf ~/anaconda3". This will remove the software from your machine.
## Resources
Here are resources that you can refer to for this class:
Anaconda documentation: https://docs.anaconda.com/
Python documentation https://docs.python.org/3/
Pandas documentation: https://pandas.pydata.org/pandas-docs/stable/
Statsmodels documentation: https://www.statsmodels.org/stable/index.html#
PyMC3 documentation: https://docs.pymc.io/
```
# end
```
| github_jupyter |
## Advanced Algorithmic Trading DT - V8
## Updates from Last Version
Find Testing of model
Do for:
- Random Forest
- Bagging
- Boosting
#### Import Packages
```
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import seaborn as sns
import math
import datetime
import gc
from sklearn.ensemble import (BaggingRegressor, RandomForestRegressor, AdaBoostRegressor)
from sklearn.model_selection import ParameterGrid
from sklearn.tree import DecisionTreeRegressor
from sklearn import linear_model
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import BayesianRidge
from sklearn.metrics import mean_squared_error
from technical_indicators import * # import all function
from sklearn.model_selection import TimeSeriesSplit
from sklearn.model_selection import train_test_split
#import parfit as pf
from sklearn.metrics import r2_score
from sklearn.preprocessing import scale
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
from sklearn.cross_decomposition import PLSRegression
```
#### Set Parameters
```
# Set the random seed, number of estimators and the "step factor" used to plot the graph of MSE for each method
random_state = 42 # Seed
n_jobs = -1 # -1 --> all Processors # Parallelisation factor for bagging, random forests (controls the number of processor cores used)
n_estimators = 200 # total number of estimators ot use in the MSE graph
step_factor = 10 # controls cranularity of calculation by stepping through the number of estimators
axis_step = int(n_estimators / step_factor) # 1000/10 = 100 separate calculations will be performed for each of the 3 ensebmle methods
```
#### Read in Data via GitHub URL
```
url = "https://raw.githubusercontent.com/meenmo/Stat479_Project/master/Data/IBM.csv"
df_ORIGINAL = pd.read_csv(url)
```
***
## Clean Data & Create Technical Indicator Variables
- Create Deep copy of dataframe
- Use Adjusted Close Data
- Drop Close
- Rename "Adj. Close" as "Close"
- Create Lagged Features
- Drop NaN
- Create Technical Indicator Variables
- Drop NaN
- Re-set index as Date
```
df_features = df_ORIGINAL.copy(deep=True) # Create Deep
df_features.drop(['Close'], axis = 1, inplace = True) # drop close column
df_features.columns = ['Date', 'High', 'Low', 'Open', 'Volume', 'Close'] # Close is actually Adj. Close
df_features['Date'] = pd.to_datetime(df_features['Date'])
#df_features.head() # sanity check
"""
Creates Lagged Returns
- given OHLCV dataframe
- numer of lagged days
"""
def create_lag_features(df, lag_days):
df_ret = df.copy()
# iterate through the lag days to generate lag values up to lag_days + 1
for i in range(1,lag_days + 2):
df_lag = df_ret[['Date', 'Close']].copy()
# generate dataframe to shift index by i day.
df_lag['Date'] = df_lag['Date'].shift(-i)
df_lag.columns = ['Date', 'value_lag' + str(i)]
# combine the valuelag
df_ret = pd.merge(df_ret, df_lag, how = 'left', left_on = ['Date'], right_on = ['Date'])
#frees memory
del df_lag
# calculate today's percentage lag
df_ret['Today'] = (df_ret['Close'] - df_ret['value_lag1'])/(df_ret['value_lag1']) * 100.0
# calculate percentage lag
for i in range(1, lag_days + 1):
df_ret['lag' + str(i)] = (df_ret['value_lag'+ str(i)] - df_ret['value_lag'+ str(i+1)])/(df_ret['value_lag'+str(i+1)]) * 100.0
# drop unneeded columns which are value_lags
for i in range(1, lag_days + 2):
df_ret.drop(['value_lag' + str(i)], axis = 1, inplace = True)
return df_ret
### Run Function
df_features = create_lag_features(df_features, 5) # 5 lag features
#df_features.head(7)
# drop earlier data with missing lag features
df_features.dropna(inplace=True)
# reset index
df_features.reset_index(drop = True, inplace = True)
#### GENERATE TECHNICAL INDICATORS FEATURES
df_features = standard_deviation(df_features, 14)
df_features = relative_strength_index(df_features, 14) # periods
df_features = average_directional_movement_index(df_features, 14, 13) # n, n_ADX
df_features = moving_average(df_features, 21) # periods
df_features = exponential_moving_average(df_features, 21) # periods
df_features = momentum(df_features, 14) #
df_features = average_true_range(df_features, 14)
df_features = bollinger_bands(df_features, 21)
df_features = ppsr(df_features)
df_features = stochastic_oscillator_k(df_features)
df_features = stochastic_oscillator_d(df_features, 14)
df_features = trix(df_features, 14)
df_features = macd(df_features, 26, 12)
df_features = mass_index(df_features)
df_features = vortex_indicator(df_features, 14)
df_features = kst_oscillator(df_features, 10, 10, 10, 15, 10, 15, 20, 30)
df_features = true_strength_index(df_features, 25, 13)
#df_features = accumulation_distribution(df_features, 14) # Causes Problems, apparently
df_features = chaikin_oscillator(df_features)
df_features = money_flow_index(df_features, 14)
df_features = on_balance_volume(df_features, 14)
df_features = force_index(df_features, 14)
df_features = ease_of_movement(df_features, 14)
df_features = commodity_channel_index(df_features, 14)
df_features = keltner_channel(df_features, 14)
df_features = ultimate_oscillator(df_features)
df_features = donchian_channel(df_features, 14)
#drop earlier data with missing lag features
df_features.dropna(inplace=True)
df_features = df_features.reset_index(drop = True)
###########################################################################################
# Store Variables now for plots later
daily_index = df_features.index
daily_returns = df_features["Today"]
daily_price = df_features["Close"]
# Re-set "Date" as the index
df_features = df_features.set_index('Date')
### Sanity Check
df_features.head(10)
```
## Standardize Data & Create X & y
- Drop all data used to create technical indicators (this is done in the book)
- Then Standardize, necessary for PLS
- Run PLS
- Select Appropriate number of components
- Create X & y
NOTE: some technical indicators use Present data, but for simplicity, just ignore this
```
### Standardize Data
##########################################################################################
# Drop Columns
list_of_columns_to_exclude = ["High", "Low", "Open", "Volume","Close", "Today"]
X_temp_standardized = df_features.copy(deep=True)
X_temp_standardized.drop(list_of_columns_to_exclude, axis = 1, inplace = True) # drop columns
# Standardize
X_temp_standardized
dates = X_temp_standardized.index # get dates to set as index after data is standardized
names = X_temp_standardized.columns # Get column names first
X_temp_standardized = StandardScaler().fit_transform(X_temp_standardized)
# Convert to DataFrame
X_temp_standardized = pd.DataFrame(X_temp_standardized, columns=names, index=dates)
X = X_temp_standardized
### Get y
##########################################################################################
y_temp = pd.DataFrame(df_features["Today"], index=X.index) # can only standardize a dataframe
sc = StandardScaler()
y = sc.fit_transform(y_temp) # Standardize, cause we did it for our original variables
y = pd.DataFrame(y, index=X.index, columns=["Today"]) # convert back to dataframe
y = y["Today"] # now re-get y as a Pandas Series
### Sanity Check
print("Shape of X: ", X.shape)
print("Shape of y: ", y.shape)
# Check Types
print(type(X)) # Needs to be <class 'pandas.core.frame.DataFrame'>
print(type(y)) # Needs ro be <class 'pandas.core.series.Series'>
```
#### Split: Train & Validatte / Test
- Train & Validate: < '2018-01-01'
- Test: >= '2018-01-01'
```
X_train_all = X.loc[(X.index < '2018-01-01')]
y_train_all = y[X_train_all.index]
# # creates all test data which is all after January 2018
X_test = X.loc[(X.index >= '2018-01-01'),:]
y_test = y[X_test.index]
### Sanity Check
print("Shape of X_train_all: ", X_train_all.shape)
print("Shape of y_train_all: ", y_train_all.shape)
print("Shape of X_test: ", X_test.shape)
print("Shape of y_test: ", y_test.shape)
```
## Time Series Train Test Split ----
### Random Forest
```
"""
Execute Random Forest for differnt number of Time Series Splits
"""
def Call_Random_Forest(numSplits):
### Prepare Random Forest
##############################################################################
# Initialize Random Forest Instance
rf = RandomForestRegressor(n_estimators=150, n_jobs=-1, random_state=123, max_features="sqrt", min_samples_split=4, max_depth=30, min_samples_leaf=1)
rf_mse = [] # MSE
rf_r2 = [] # R2
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
# # Print Statements
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
# print("Split: ", splitCount)
# print('Observations: ', (X_train.shape[0] + X_val.shape[0]))
# #print('Cutoff date, or first date in validation data: ', X_val.iloc[0,0])
# print('Training Observations: ', (X_train.shape[0]))
# print('Testing Observations: ', (X_val.shape[0]))
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
### Run Random Forest
rf.fit(X_train, y_train)
prediction = rf.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
rf_mse.append(mse)
rf_r2.append(r2)
# print("rf_mse: ", rf_mse)
# print("rf_r2: ", rf_r2)
### Time Series Split
##############################################################################
# Plot the chart of MSE versus number of estimators
plt.figure(figsize=(12, 7))
plt.title('Random Forest - MSE & R-Squared')
### MSE
plt.plot(list(range(1,splitCount+1)), rf_mse, 'b-', color="blue", label='MSE')
plt.plot(list(range(1,splitCount+1)), rf_r2, 'b-', color="green", label='R-Squared')
plt.plot(list(range(1,splitCount+1)), np.array([0] * splitCount), 'b-', color="red", label='Zero')
plt.legend(loc='upper right')
plt.xlabel('Train/Test Split Number')
plt.ylabel('Mean Squared Error & R-Squared')
plt.show()
print("rf_r2: ", rf_r2)
#print(rf.feature_importances_)
print("Mean r2: ", np.mean(rf_r2))
```
### Bagging
```
"""
Execute Bagging for differnt number of Time Series Splits
"""
def Call_Bagging(numSplits):
### Prepare Bagging
##############################################################################
# Initialize Bagging Instance
bagging = BaggingRegressor(n_estimators=150, n_jobs=-1, random_state=123)
bagging_mse = [] # MSE
bagging_r2 = [] # R2
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
# # Print Statements
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
# print("Split: ", splitCount)
# print('Observations: ', (X_train.shape[0] + X_test.shape[0]))
# #print('Cutoff date, or first date in validation data: ', X_val.iloc[0,0])
# print('Training Observations: ', (X_train.shape[0]))
# print('Testing Observations: ', (X_test.shape[0]))
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
### Run Random Forest
bagging.fit(X_train, y_train)
prediction = bagging.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
bagging_mse.append(mse)
bagging_r2.append(r2)
### Time Series Split
##############################################################################
# Plot the chart of MSE versus number of estimators
plt.figure(figsize=(12, 7))
plt.title('Bagging - MSE & R-Squared')
### MSE
plt.plot(list(range(1,splitCount+1)), bagging_mse, 'b-', color="blue", label='MSE')
plt.plot(list(range(1,splitCount+1)), bagging_r2, 'b-', color="green", label='R-Squared')
plt.plot(list(range(1,splitCount+1)), np.array([0] * splitCount), 'b-', color="red", label='Zero')
plt.legend(loc='upper right')
plt.xlabel('Train/Test Split Number')
plt.ylabel('Mean Squared Error & R-Squared')
plt.show()
print("bagging_r2: ", bagging_r2)
print(np.mean(bagging_r2))
```
### Boosting
```
"""
Execute Random Forest for differnt number of Time Series Splits
"""
def Call_Boosting(numSplits):
### Prepare Boosting
##############################################################################
# Initialize Boosting Instance
boosting = AdaBoostRegressor(DecisionTreeRegressor(),
n_estimators=150, random_state=123,learning_rate=0.01)
boosting_mse = [] # MSE
boosting_r2 = [] # R2
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
# # Print Statements
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
# print("Split: ", splitCount)
# print('Observations: ', (X_train.shape[0] + X_test.shape[0]))
# #print('Cutoff date, or first date in validation data: ', X_val.iloc[0,0])
# print('Training Observations: ', (X_train.shape[0]))
# print('Testing Observations: ', (X_test.shape[0]))
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
### Run Random Forest
boosting.fit(X_train, y_train)
prediction = boosting.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
boosting_mse.append(mse)
boosting_r2.append(r2)
### Time Series Split
##############################################################################
# Plot the chart of MSE versus number of estimators
plt.figure(figsize=(12, 7))
plt.title('Boosting - MSE & R-Squared')
### MSE
plt.plot(list(range(1,splitCount+1)), boosting_mse, 'b-', color="blue", label='MSE')
plt.plot(list(range(1,splitCount+1)), boosting_r2, 'b-', color="green", label='R-Squared')
plt.plot(list(range(1,splitCount+1)), np.array([0] * splitCount), 'b-', color="red", label='Zero')
plt.legend(loc='upper right')
plt.xlabel('Train/Test Split Number')
plt.ylabel('Mean Squared Error & R-Squared')
plt.show()
print("boosting_r2: ", boosting_r2)
```
### Linear Regression
```
"""
Execute Linear Regression for different number of Time Series Splits
"""
def Call_Linear(numSplits):
### Prepare Random Forest
##############################################################################
# Initialize Random Forest Instance
linear = LinearRegression(n_jobs=-1, normalize=True, fit_intercept=False) # if we don't fit the intercept we get a better prediction
linear_mse = [] # MSE
linear_r2 = [] # R2
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
# # Print Statements
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
# print("Split: ", splitCount)
# print('Observations: ', (X_train.shape[0] + X_test.shape[0]))
# #print('Cutoff date, or first date in validation data: ', X_val.iloc[0,0])
# print('Training Observations: ', (X_train.shape[0]))
# print('Testing Observations: ', (X_test.shape[0]))
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
### Run Random Forest
linear.fit(X_train, y_train)
prediction = linear.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
r2 = np.corrcoef(y_val, prediction)[0, 1]
r2 = r2*r2 # square of correlation coefficient --> R-squared
linear_mse.append(mse)
linear_r2.append(r2)
### Time Series Split
##############################################################################
# Plot the chart of MSE versus number of estimators
plt.figure(figsize=(12, 7))
plt.title('Linear Regression - MSE & R-Squared')
### MSE
plt.plot(list(range(1,splitCount+1)), linear_mse, 'b-', color="blue", label='MSE')
plt.plot(list(range(1,splitCount+1)), linear_r2, 'b-', color="green", label='R-Squared')
plt.plot(list(range(1,splitCount+1)), np.array([0] * splitCount), 'b-', color="red", label='Zero')
plt.legend(loc='upper right')
plt.xlabel('Train/Test Split Number')
plt.ylabel('Mean Squared Error & R-Squared')
plt.show()
print("linear_r2: ", linear_r2)
```
### Misc. Graphs ---- Price, Returns & Cumulative Returns
```
# figure dimenstions
length = 15
height = 5
### Prices
plt.figure(figsize=(length, height))
plt.title('IBM Adj Close Price Graph')
plt.plot(daily_index, daily_price, 'b-', color="blue", label='Prices')
plt.legend(loc='upper right')
plt.xlabel('Days')
plt.ylabel('Prices')
plt.show()
### Returns
plt.figure(figsize=(length, height))
plt.title('IBM Daily Returns')
plt.plot(daily_index, daily_returns, 'b-', color="blue", label='Returns')
plt.legend(loc='upper right')
plt.xlabel('Days')
plt.ylabel('Returns')
plt.show()
### Cumulative Returns
plt.figure(figsize=(length, height))
plt.title('IBM Cumulative Returns')
cumulative_returns = daily_returns.cumsum()
plt.plot(daily_index, cumulative_returns, 'b-', color="green", label='Cumulative Returns')
plt.legend(loc='upper right')
plt.xlabel('Days')
plt.ylabel('Cumulative Return')
plt.show()
```
### First - A Note on R-Squared
##### What Does A Negative R Squared Value Mean?
- What does R-squared tell us?
- It tells us whether a horizonal line through the vertical mean of the data is a better predictor
- For a Linear Regression
- R-squared is just the coreelation coefficient squared
- R-squared can't be negative, becasue at 0, it becomes the horizontal line
- For All other Model
- For practical purposes, the lowest R2 you can get is zero, but only because the assumption is that if your regression line is not better than using the mean, then you will just use the mean value.
- However if your regression line is worse than using the mean value, the r squared value that you calculate will be negative.
- Note that the reason R2 can't be negative in the linear regression case is just due to chance and how linear regression is contructed
***
***
***
## Grid Search for Best Model
### Random Forest
```
"""
Execute Random Forest for differnt number of Time Series Splits
Parameters
- numSplits: Number of Train/Validation splits to run in order to compute our final model (bigger --> more accurate & more complex)
- minSamplesSplit:
- maxDepth: Max Depth of the DT
- minSamplesLeaf:
Returns
- Average R-Squared of the DT model across all Train/Validation splits
"""
def Call_Random_Forest_Grid_Search(numSplits, minSamplesSplit, maxDepth, minSamplesLeaf):
### Prepare Random Forest
##############################################################################
# Initialize Random Forest Instance
rf_mse = [] # MSE
rf_r2 = [] # R2
rf = RandomForestRegressor(n_estimators=150, n_jobs=-1, random_state=123, max_features="sqrt",
min_samples_split=minSamplesSplit,
max_depth=maxDepth,
min_samples_leaf=minSamplesLeaf)
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
### Run Random Forest
rf.fit(X_train, y_train)
prediction = rf.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
rf_mse.append(mse)
rf_r2.append(r2)
return np.mean(rf_r2)
# Call_Random_Forest_Grid_Search(numSplits, minSamplesSplit, maxDepth, minSamplesLeaf)
minSamplesSplit_list = [2,5,10,15,20]
maxDepth_list = [15,20,25]
minSamplesLeaf_list = [2,5,10]
best_model_parameters = [0,0,0]
max_r2 = -100
count = 0
# Loop over all possible parameters
for minSamplesSplit in minSamplesSplit_list:
for maxDepth in maxDepth_list:
for minSamplesLeaf in minSamplesLeaf_list:
count += 1
temp_mean_r2 = Call_Random_Forest_Grid_Search(20, minSamplesSplit, maxDepth, minSamplesLeaf) # Call Random Forest Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2:
max_r2 = temp_mean_r2 # store new max
best_model_parameters[0] = minSamplesSplit
best_model_parameters[1] = maxDepth
best_model_parameters[2] = minSamplesLeaf
print("Best R2: ", max_r2)
best_model_parameters
minSamplesSplit_list = [4,5,6]
maxDepth_list = [25,30,35]
minSamplesLeaf_list = [1,2,3]
best_model_parameters = [0,0,0]
max_r2 = -100
count = 0
# Loop over all possible parameters
for minSamplesSplit in minSamplesSplit_list:
for maxDepth in maxDepth_list:
for minSamplesLeaf in minSamplesLeaf_list:
count += 1
temp_mean_r2 = Call_Random_Forest_Grid_Search(20, minSamplesSplit, maxDepth, minSamplesLeaf) # Call Random Forest Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2:
max_r2 = temp_mean_r2 # store new max
best_model_parameters[0] = minSamplesSplit
best_model_parameters[1] = maxDepth
best_model_parameters[2] = minSamplesLeaf
print("Best R2: ", max_r2)
best_model_parameters
minSamplesSplit_list = [2,3,4]
maxDepth_list = [29,30,31]
minSamplesLeaf_list = [1,2]
best_model_parameters = [0,0,0]
max_r2 = -100
count = 0
# Loop over all possible parameters
for minSamplesSplit in minSamplesSplit_list:
for maxDepth in maxDepth_list:
for minSamplesLeaf in minSamplesLeaf_list:
count += 1
temp_mean_r2 = Call_Random_Forest_Grid_Search(20, minSamplesSplit, maxDepth, minSamplesLeaf) # Call Random Forest Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2:
max_r2 = temp_mean_r2 # store new max
best_model_parameters[0] = minSamplesSplit
best_model_parameters[1] = maxDepth
best_model_parameters[2] = minSamplesLeaf
print("Best R2: ", max_r2)
print(best_model_parameters)
```
# Best Model: Random Forest
##### According to Grid Search, our best model is:
- [4, 30, 1]
- minSamplesSplit = 4
- maxDepth = 30
- minSamplesLeaf = 1
## Revised RandomForest
```
"""
Execute Random Forest for differnt number of Time Series Splits
Parameters
- numSplits: Number of Train/Validation splits to run in order to compute our final model (bigger --> more accurate & more complex)
- maxDepth: Max Depth of the DT
- minSamplesLeaf: min samples to split (basically pruning)
Returns
- Average R-Squared of the DT model across all Train/Validation splits
"""
def Call_Random_Forest_Grid_Search(numSplits, maxDepth, minSamplesLeaf):
### Prepare Random Forest
##############################################################################
# Initialize Random Forest Instance
rf_mse = [] # MSE
rf_r2 = [] # R2
rf = RandomForestRegressor(n_estimators=150, n_jobs=-1, random_state=123, max_features="sqrt",
max_depth=maxDepth,
min_samples_leaf=minSamplesLeaf)
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
### Run Random Forest
rf.fit(X_train, y_train)
prediction = rf.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
rf_mse.append(mse)
rf_r2.append(r2)
return np.mean(rf_r2)
# Call_Random_Forest_Grid_Search(numSplits, minSamplesSplit, maxDepth, minSamplesLeaf)
minSamplesLeaf_list = [2,5,10,15]
maxDepth_list = [15,20,25,30]
best_model_parameters = [0,0]
max_r2 = -100
count = 0
rf_df = pd.DataFrame()
# Loop over all possible parameters
for minSamplesLeaf in minSamplesLeaf_list:
for maxDepth in maxDepth_list:
count += 1
temp_mean_r2 = Call_Random_Forest_Grid_Search(20, maxDepth, minSamplesLeaf) # Call Random Forest Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2:
max_r2 = temp_mean_r2 # store new max
best_model_parameters[0] = minSamplesLeaf
best_model_parameters[1] = maxDepth
rf_df.loc[count,'min_sample_leaf'] = minSamplesLeaf
rf_df.loc[count,'max_depth'] = maxDepth
rf_df.loc[count,'mean_r2'] = temp_mean_r2
print("Best R2: ", max_r2)
print(best_model_parameters)
import seaborn as sns
# plot nicely
x_index = np.unique(rf_df.loc[:,'min_sample_leaf'])
y_index = np.unique((rf_df.loc[:,'max_depth']))
a = pd.DataFrame(index = x_index, columns = y_index)
for i in x_index:
for j in y_index:
a.loc[i,j] = list(rf_df.loc[(rf_df['min_sample_leaf'] == i) & (rf_df['max_depth'] == j),'mean_r2'])[0]#.astype(float)
a = a.apply(pd.to_numeric)
sns.heatmap(a)
plt.xlabel('max_depth')
plt.ylabel('min_samples_leaf')
plt.title('r2')
plt.show()
```
### Bagging
```
"""
Execute Bagging for differnt number of Time Series Splits
Parameters
- numSplits: Number of Train/Validation splits to run in order to compute our final model (bigger --> more accurate & more complex)
- min_samples_leaf: minimum samples to split
- max_depth: depth of tree
Returns
- Average R-Squared of the DT model across all Train/Validation splits"""
def Call_Bagging_Grid_Search(numSplits, min_samples_leaf, max_depth):
### Prepare Bagging
##############################################################################
# Initialize Bagging Instance
bagging_mse = [] # MSE
bagging_r2 = [] # R2
bagging = BaggingRegressor(DecisionTreeRegressor(min_samples_leaf = min_samples_leaf, max_depth = max_depth),
n_estimators=150, random_state=123,
#n_jobs = -1,
max_samples=100,
max_features=20)
# bagging = BaggingRegressor(n_estimators=150, n_jobs=-1, random_state=123)
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
# # Print Statements
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
# print("Split: ", splitCount)
# print('Observations: ', (X_train.shape[0] + X_test.shape[0]))
# #print('Cutoff date, or first date in validation data: ', X_val.iloc[0,0])
# print('Training Observations: ', (X_train.shape[0]))
# print('Testing Observations: ', (X_test.shape[0]))
# print("~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~~")
### Run Bagging
bagging.fit(X_train, y_train)
prediction = bagging.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
bagging_mse.append(mse)
bagging_r2.append(r2)
return np.mean(bagging_r2)
min_sample_leaf = [5,10,15]
max_depth = [5,10,15]
best_model_parameters_bag = [0,0]
max_r2_bag = -100
count = 0
bag_df = pd.DataFrame()
# Loop over all possible parameters
for minSampleLeaf in min_sample_leaf:
for maxDepth in max_depth:
count += 1
temp_mean_r2 = Call_Bagging_Grid_Search(20, minSampleLeaf, maxDepth) # Call Boosting Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2_bag:
max_r2_bag = temp_mean_r2 # store new max
best_model_parameters_bag[0] = minSampleLeaf
best_model_parameters_bag[1] = maxDepth
bag_df.loc[count,'min_sample_leaf'] = minSampleLeaf
bag_df.loc[count,'max_depth'] = maxDepth
bag_df.loc[count,'mean_r2'] = temp_mean_r2
print("Best R2: ", max_r2_bag)
print(best_model_parameters_bag)
import seaborn as sns
# plot nicely
x_index = np.unique(bag_df.loc[:,'min_sample_leaf'])
y_index = np.unique((bag_df.loc[:,'max_depth']))
a = pd.DataFrame(index = x_index, columns = y_index)
for i in x_index:
for j in y_index:
a.loc[i,j] = list(bag_df.loc[(bag_df['min_sample_leaf'] == i) & (bag_df['max_depth'] == j),'mean_r2'])[0]#.astype(float)
a = a.apply(pd.to_numeric)
sns.heatmap(a)
plt.xlabel('max_depth')
plt.ylabel('min_samples_leaf')
plt.title('r2')
plt.show()
```
# Best Model: Bagging
##### According to Grid Search, our best model is:
- minsampleleaf = 5
- maxdepth = 15
- I had to put max-samples as 100, because some of the split has max samples of only slightly over 100
# AdaBoost
```
"""
Execute Boosting for differnt number of Time Series Splits
Parameters
- numSplits: Number of Train/Validation splits to run in order to compute our final model (bigger --> more accurate & more complex)
- min_samples_leaf: Mininum number of samples to split. if a leaf node has greater than this parameter, we can still split
- max_depth: depth of tree
- learning_rate: gradient descent alpha
Returns
- Average R-Squared of the DT model across all Train/Validation splits"""
def Call_Boosting_Grid_Search(numSplits, min_samples_leaf, max_depth, learning_rate):
### Prepare Boosting
##############################################################################
# Initialize Bagging Instance
boost_mse = [] # MSE
boost_r2 = [] # R2
boost = AdaBoostRegressor(DecisionTreeRegressor(max_features = 'sqrt', random_state = 123, min_samples_leaf = min_samples_leaf,
max_depth = max_depth),
n_estimators=150, random_state=123,learning_rate=learning_rate)
### Time Series Split
##############################################################################
splits = TimeSeriesSplit(n_splits=numSplits) # 3 splits
splitCount = 0 # dummy count var to track current split num in print statements
for train_index, test_index in splits.split(X_train_all):
splitCount += 1
# Train Split
X_train = X_train_all.iloc[train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_train_all.iloc[test_index,:]
y_val = y[X_val.index]
### Run Boosting
boost.fit(X_train, y_train)
prediction = boost.predict(X_val)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
boost_mse.append(mse)
boost_r2.append(r2)
return np.mean(boost_r2)
### WARNING TAKES VERY LONG TO RUN
min_sample_leaf = [5,10,15]
max_depth = [5,10,15]
learning_rate = [0.01,0.1,1]
best_model_parameters_boost = [0,0,0]
max_r2_boosting = -100
count = 0
boosting_df = pd.DataFrame()
# Loop over all possible parameters
for minSampleLeaf in min_sample_leaf:
for maxDepth in max_depth:
for learn in learning_rate:
count += 1
temp_mean_r2 = Call_Boosting_Grid_Search(20, minSampleLeaf, maxDepth, learn) # Call Boosting Train/Validation
print("temp_mean ", count, ": ", temp_mean_r2)
if temp_mean_r2 > max_r2_boosting:
max_r2_boosting = temp_mean_r2 # store new max
best_model_parameters_boost[0] = minSampleLeaf
best_model_parameters_boost[1] = maxDepth
best_model_parameters_boost[2] = learn
boosting_df.loc[count,'min_sample_leaf'] = minSampleLeaf
boosting_df.loc[count,'max_depth'] = maxDepth
boosting_df.loc[count, 'Learning_rate'] = learn
boosting_df.loc[count,'mean_r2'] = temp_mean_r2
print("Best R2: ", max_r2_boosting)
print(best_model_parameters_boost)
display(boosting_df)
import seaborn as sns
# plot nicely
for learn in ([0.01,0.1,1]):
x_index = np.unique(boosting_df.loc[:,'min_sample_leaf'])
y_index = np.unique((boosting_df.loc[:,'max_depth']))
a = pd.DataFrame(index = x_index, columns = y_index)
for i in x_index:
for j in y_index:
a.loc[i,j] = list(boosting_df.loc[(boosting_df['min_sample_leaf'] == i) & (boosting_df['max_depth'] == j) & (boosting_df['Learning_rate'] == learn),'mean_r2'])[0]#.astype(float)
a = a.apply(pd.to_numeric)
sns.heatmap(a)
plt.title('learning_rate' + str(learn))
plt.xlabel('max_depth')
plt.ylabel('min_samples_leaf')
plt.show()
```
## Best Model:
#### According to Grid Search, our best model is:
- min_samples_leaf = 5
- max_depth = 15
- learning_rate = 1
***
# FINAL PREDICTION - Naive (Fixed via Retraining the Model, below)
```
train_mse=[]
train_r2=[]
test_mse=[]
test_r2=[]
# RandomForestRegressor
rf = RandomForestRegressor(n_estimators=150, n_jobs=-1, random_state=123, max_features="sqrt",
max_depth=30,
min_samples_leaf=10)
rf.fit(X_train_all, y_train_all)
mse_train = mean_squared_error(y_train_all, rf.predict(X_train_all))
r2_train = r2_score(y_train_all, rf.predict(X_train_all))
mse_test = mean_squared_error(y_test, rf.predict(X_test))
r2_test = r2_score(y_test, rf.predict(X_test))
train_mse.append(mse_train)
train_r2.append(r2_train)
test_mse.append(mse_test)
test_r2.append(r2_test)
print('MSE on Train:', mse_train)
print('r2 on Train:', r2_train)
print('MSE on Test:', mse_test)
print('r2 on Test:', r2_test)
# bagging
bag = BaggingRegressor(DecisionTreeRegressor(max_depth = 15, min_samples_leaf = 5),
n_estimators=150, n_jobs=1, random_state=123,
max_samples=1500,
max_features=20)
bag.fit(X_train_all, y_train_all)
mse_train = mean_squared_error(y_train_all, bag.predict(X_train_all))
r2_train = r2_score(y_train_all, bag.predict(X_train_all))
mse_test = mean_squared_error(y_test, bag.predict(X_test))
r2_test = r2_score(y_test, bag.predict(X_test))
train_mse.append(mse_train)
train_r2.append(r2_train)
test_mse.append(mse_test)
test_r2.append(r2_test)
print('MSE on Train:', mse_train)
print('r2 on Train:', r2_train)
print('MSE on Test:', mse_test)
print('r2 on Test:', r2_test)
# BOOSTING
boost = AdaBoostRegressor(DecisionTreeRegressor(max_features = 'sqrt', random_state = 123, min_samples_leaf = 5,
max_depth = 15),
n_estimators=150, random_state=123,learning_rate=1)
boost.fit(X_train_all, y_train_all)
mse_train = mean_squared_error(y_train_all, boost.predict(X_train_all))
r2_train = r2_score(y_train_all, boost.predict(X_train_all))
mse_test = mean_squared_error(y_test, boost.predict(X_test))
r2_test = r2_score(y_test, boost.predict(X_test))
train_mse.append(mse_train)
train_r2.append(r2_train)
test_mse.append(mse_test)
test_r2.append(r2_test)
print('MSE on Train:', mse_train)
print('r2 on Train:', r2_train)
print('MSE on Test:', mse_test)
print('r2 on Test:', r2_test)
plt.plot(['RF','Bagging','Boosting'],train_mse,label='MSE')
plt.plot(train_r2,label='R^2')
plt.legend()
plt.title('MSE & R^2 of Training Set')
plt.show()
plt.plot(['RF','Bagging','Boosting'],test_mse,label='MSE')
plt.plot(test_r2,label='R^2')
plt.legend()
plt.title('MSE & R^2 of Test Set')
plt.show()
```
#### Random Forest
```
# plot best model which is Random Forest
plt.figure(figsize=(15, 7))
plt.plot(sc.inverse_transform(np.array(y_test)))
plt.plot(sc.inverse_transform(rf.predict(X_test)))
plt.plot(list(range(1,210+1)), np.array([0] * 210), 'b-', color="red", label='Zero')
plt.legend(["Realized Returns","Forecasted Returns","Zero"])
plt.ylabel('% Returns')
plt.xlabel('days')
plt.title('Random Forest: Predicted Returns vs Actual Returns')
plt.show()
```
#### Bagging
```
# plot best model which is Bagging
plt.figure(figsize=(15, 7))
plt.plot(sc.inverse_transform(np.array(y_test)))
plt.plot(sc.inverse_transform(bag.predict(X_test)))
plt.plot(list(range(1,210+1)), np.array([0] * 210), 'b-', color="red", label='Zero')
plt.legend(["Realized Returns","Forecasted Returns","Zero"])
plt.ylabel('% Returns')
plt.xlabel('days')
plt.title('Bagging: Predicted Returns vs Actual Returns')
plt.show()
```
#### Boosting
```
# plot best model which is boosting
plt.figure(figsize=(15, 7))
plt.plot(sc.inverse_transform(np.array(y_test)))
plt.plot(sc.inverse_transform(boost.predict(X_test)))
plt.plot(list(range(1,210+1)), np.array([0] * 210), 'b-', color="red", label='Zero')
plt.legend(["Realized Returns","Forecasted Returns","Zero"])
plt.ylabel('% Returns')
plt.xlabel('days')
plt.title('Boosting: Predicted Returns vs Actual Returns')
plt.show()
print("Cumulative Realized Returns: ", sc.inverse_transform(np.array(y_test)).cumsum()[-1])
print("Cumulative Random Forest Returns ", sc.inverse_transform(rf.predict(X_test)).cumsum()[-1])
print("Cumulative Boosting Returns ", sc.inverse_transform(bag.predict(X_test)).cumsum()[-1])
print("Cumulative Boosting Returns ", sc.inverse_transform(boost.predict(X_test)).cumsum()[-1])
sorted_returns = np.array(sc.inverse_transform(np.array(y_test)))
#print(sorted(sorted_returns, reverse=False)[0:4])
returns_minus_worst_4_days = np.array(sorted(sorted_returns, reverse=False)[4:]).cumsum()[-1]
returns_minus_worst_4_days
```
#### Cumulative Returns
```
# plot best model which is boosting
plt.figure(figsize=(15, 7))
plt.plot(sc.inverse_transform(np.array(y_test)).cumsum())
plt.plot(sc.inverse_transform(boost.predict(X_test)).cumsum())
plt.plot(list(range(1,210+1)), np.array([0] * 210), 'b-', color="red", label='Zero')
plt.legend(["Cumulative Realized Returns","Cumulative Forecasted Returns","Zero"])
plt.ylabel('% Returns')
plt.xlabel('days')
plt.title('Boosting: Predicted Returns vs Actual Returns')
plt.show()
```
***
# FINAL PREDICTION - Fixed from V7
```
rf_metrics = []
bag_metrics = []
boost_metrics = []
rf_prediction = []
bag_prediction = []
boost_prediction = []
```
#### Set Final Model Parameters
```
### Number of Testing Splits/Times we Re-train the model on latest data
N = 10
### Random Forest
max_depth_rf = 30
min_samples_leaf_rf = 10
# ### Base Decision Tree (Bagging & Boosting)
# max_depth_ = 15
# min_samples_leaf_ = 5
### Bagging
max_depth_bagging = 15
min_samples_leaf_bagging = 5
max_samples_bagging = 1500
max_features = 20
### Boosting
max_depth_boosting = 15
min_samples_leaf_boosting = 5
learning_rate = 1
```
#### Run Final Models (Random Forest, Bagging, Boosting)
```
## WARNING Takes long to run
splits = TimeSeriesSplit(n_splits=N)
rf = RandomForestRegressor(n_estimators=150, n_jobs=-1, random_state=123, max_features="sqrt",
max_depth=max_depth_rf,
min_samples_leaf=min_samples_leaf_rf)
bag = BaggingRegressor(DecisionTreeRegressor(max_features = 'sqrt', random_state = 123,
max_depth = max_depth_bagging,
min_samples_leaf = min_samples_leaf_bagging),
n_estimators=150,
n_jobs=-1,
random_state=123,
max_samples=max_samples_bagging,
max_features=max_features)
boost = AdaBoostRegressor(DecisionTreeRegressor(max_features = 'sqrt', random_state = 123,
max_depth = max_depth_boosting),
min_samples_leaf = min_samples_leaf_boosting,
n_estimators=150,
random_state=123,
learning_rate=learning_rate)
# predict first fold of test set, need to change 21 if using different than 10 cv
### Run Random Forest
rf.fit(X_train_all, y_train_all)
prediction = rf.predict(X_test.iloc[0:21,:])
rf_prediction.append(prediction)
y_val = y[X_test.iloc[0:21,:].index]
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
rf_metrics.append([mse,r2])
# ### Run bagging
bag.fit(X_train_all, y_train_all)
prediction = bag.predict(X_test.iloc[0:21,:])
bag_prediction.append(prediction)
y_val = y[X_test.iloc[0:21,:].index]
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
bag_metrics.append([mse,r2])
## Run boosting
boost.fit(X_train_all, y_train_all)
prediction = boost.predict(X_test.iloc[0:21,:])
boost_prediction.append(prediction)
y_val = y[X_test.iloc[0:21,:].index]
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
boost_metrics.append([mse,r2])
splitCount = 0
for train_index, test_index in splits.split(X_test):
splitCount += 1
# Calculate NEW Train/Test indices
train_index = X_train_all.shape[0] + len(train_index)
# Train Split
X_train = X.iloc[0:train_index,:]
y_train = y[X_train.index]
# Validate Split
X_val = X_test.iloc[test_index[0]:(test_index[-1]+1),:]
y_val = y[X_val.index]
# print('Train_start', X_train.index[0])
# print('Train:', X_train.index[-1])
# print('Val_start:', X_val.index[0])
# print('Val_end:', X_val.index[-1])
### Run Random Forest
rf.fit(X_train, y_train)
prediction = rf.predict(X_val)
rf_prediction.append(prediction)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
rf_metrics.append([mse,r2])
### Run bagging
bag.fit(X_train, y_train)
prediction = bag.predict(X_val)
bag_prediction.append(prediction)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
bag_metrics.append([mse,r2])
## Run boosting
boost.fit(X_train, y_train)
prediction = boost.predict(X_val)
boost_prediction.append(prediction)
mse = mean_squared_error(y_val, prediction)
r2 = r2_score(y_val, prediction)
boost_metrics.append([mse,r2])
print(splitCount)
#make into one array instead of 10x10 matrix
rf_prediction = np.array(rf_prediction)
rf_prediction = np.concatenate(rf_prediction, axis=0)
bag_prediction = np.array(bag_prediction)
bag_prediction = np.concatenate(bag_prediction, axis=0)
boost_prediction = np.array(boost_prediction)
boost_prediction = np.concatenate(boost_prediction, axis=0)
rf_mse = mean_squared_error(y_test, rf_prediction)
rf_r2 = r2_score(y_test, rf_prediction)
bag_mse = mean_squared_error(y_test, bag_prediction)
bag_r2 = r2_score(y_test, bag_prediction)
boost_mse = mean_squared_error(y_test, boost_prediction)
boost_r2 = r2_score(y_test, boost_prediction)
plt.plot(['RF','Bagging','Boosting'],[rf_mse, bag_mse, boost_mse],label='MSE')
plt.plot([rf_r2, bag_r2, boost_r2],label='R^2')
plt.legend()
plt.title('MSE & R^2 of final test prediction')
plt.show()
```
| github_jupyter |
```
!pip install torch
!pip install torchsummary
import torch
import torch.nn as nn
from torchsummary import summary
"""
Implementation based on original paper NeurIPS 2016 https://papers.nips.cc/paper/6096-learning-a-probabilistic-latent-space-of-object-shapes-via-3d-generative-adversarial-modeling.pdf
"""
```
## Discriminator
```
class Discriminator(torch.nn.Module):
def __init__(self, in_channels=3, dim=64, out_conv_channels=512):
super(Discriminator, self).__init__()
conv1_channels = int(out_conv_channels / 8)
conv2_channels = int(out_conv_channels / 4)
conv3_channels = int(out_conv_channels / 2)
self.out_conv_channels = out_conv_channels
self.out_dim = int(dim / 16)
self.conv1 = nn.Sequential(
nn.Conv3d(
in_channels=in_channels, out_channels=conv1_channels, kernel_size=4,
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv1_channels),
nn.LeakyReLU(0.2, inplace=True)
)
self.conv2 = nn.Sequential(
nn.Conv3d(
in_channels=conv1_channels, out_channels=conv2_channels, kernel_size=4,
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv2_channels),
nn.LeakyReLU(0.2, inplace=True)
)
self.conv3 = nn.Sequential(
nn.Conv3d(
in_channels=conv2_channels, out_channels=conv3_channels, kernel_size=4,
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv3_channels),
nn.LeakyReLU(0.2, inplace=True)
)
self.conv4 = nn.Sequential(
nn.Conv3d(
in_channels=conv3_channels, out_channels=out_conv_channels, kernel_size=4,
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(out_conv_channels),
nn.LeakyReLU(0.2, inplace=True)
)
self.out = nn.Sequential(
nn.Linear(out_conv_channels * self.out_dim * self.out_dim * self.out_dim, 1),
nn.Sigmoid(),
)
def forward(self, x):
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.conv4(x)
# Flatten and apply linear + sigmoid
x = x.view(-1, self.out_conv_channels * self.out_dim * self.out_dim * self.out_dim)
x = self.out(x)
return x
```
## Generator
```
class Generator(torch.nn.Module):
def __init__(self, in_channels=512, out_dim=64, out_channels=1, noise_dim=200, activation="sigmoid"):
super(Generator, self).__init__()
self.in_channels = in_channels
self.out_dim = out_dim
self.in_dim = int(out_dim / 16)
conv1_out_channels = int(self.in_channels / 2.0)
conv2_out_channels = int(conv1_out_channels / 2)
conv3_out_channels = int(conv2_out_channels / 2)
self.linear = torch.nn.Linear(noise_dim, in_channels * self.in_dim * self.in_dim * self.in_dim)
self.conv1 = nn.Sequential(
nn.ConvTranspose3d(
in_channels=in_channels, out_channels=conv1_out_channels, kernel_size=(4, 4, 4),
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv1_out_channels),
nn.ReLU(inplace=True)
)
self.conv2 = nn.Sequential(
nn.ConvTranspose3d(
in_channels=conv1_out_channels, out_channels=conv2_out_channels, kernel_size=(4, 4, 4),
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv2_out_channels),
nn.ReLU(inplace=True)
)
self.conv3 = nn.Sequential(
nn.ConvTranspose3d(
in_channels=conv2_out_channels, out_channels=conv3_out_channels, kernel_size=(4, 4, 4),
stride=2, padding=1, bias=False
),
nn.BatchNorm3d(conv3_out_channels),
nn.ReLU(inplace=True)
)
self.conv4 = nn.Sequential(
nn.ConvTranspose3d(
in_channels=conv3_out_channels, out_channels=out_channels, kernel_size=(4, 4, 4),
stride=2, padding=1, bias=False
)
)
if activation == "sigmoid":
self.out = torch.nn.Sigmoid()
else:
self.out = torch.nn.Tanh()
def project(self, x):
"""
projects and reshapes latent vector to starting volume
:param x: latent vector
:return: starting volume
"""
return x.view(-1, self.in_channels, self.in_dim, self.in_dim, self.in_dim)
def forward(self, x):
x = self.linear(x)
x = self.project(x)
x = self.conv1(x)
x = self.conv2(x)
x = self.conv3(x)
x = self.conv4(x)
return self.out(x)
```
## Test
```
def test_gan3d(print_summary=True):
noise_dim = 200 # latent space vector dim
in_channels = 512 # convolutional channels
dim = 64 # cube volume
model_generator = Generator(in_channels=512, out_dim=dim, out_channels=1, noise_dim=noise_dim)
noise = torch.rand(1, noise_dim)
generated_volume = model_generator(noise)
print("Generator output shape", generated_volume.shape)
model_discriminator = Discriminator(in_channels=1, dim=dim, out_conv_channels=in_channels)
out = model_discriminator(generated_volume)
print("Discriminator output", out.item())
if print_summary:
print("\n\nGenerator summary\n\n")
summary(model_generator, (1, noise_dim))
print("\n\nDiscriminator summary\n\n")
summary(model_discriminator, (1,dim,dim,dim))
test_gan3d()
```
| github_jupyter |
# Project 2 - Mc907/Mo651 - Mobile Robotics
### Student:
Luiz Eduardo Cartolano - RA: 183012
### Instructor:
Esther Luna Colombini
### Github Link:
[Project Repository](https://github.com/luizcartolano2/mc907-mobile-robotics)
### Youtube Link:
[Link to Video](https://youtu.be/uqNeEhWo0dA)
### Subject of this Work:
The general objective of this work is to implement and evaluate at least 1 robot control behavior per group member.
### Goals:
1. Implement and evaluate at least 1 robot control behavior per group member (AvoidObstacle, WallFollow, Go- ToGoal) using models based on PID, Fuzzy, Neural Networks, etc;
2. Propose a behavior coordination strategy (state machine, planner, AR, subsumption, etc.)
# Code Starts Here
Import of used libraries
```
from lib import vrep
import sys, time
from src import robot as rb
from src.utils import vrep2array
import math
from time import time
import numpy as np
import cv2
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
import skfuzzy
import skfuzzy as fuzz
import skfuzzy.control as ctrl
# from reinforcement_learning.train import *
```
# Defining the kinematic model of the Pionner P3DX
For this project, we are going to use the configuration of the mobile robot being characterized by the position (x,y) and the orientation in a Cartesian coordinate.
Using the follow parameters:
1. $V_R$: linear velocity of the right wheel.
2. $V_L$: linear velocity of the left wheel.
3. $W$: angular velocity of the mobile robot.
4. $X$: abscissa of the robot.
5. $Y$: intercept of the robot.
6. $X,Y$ : the actual position coordinates.
7. $\theta$: orientation of the robot.
8. $L$: the distance between the driving wheels.
The kinematic model is given by these equations [1](https://www.hindawi.com/journals/cin/2016/9548482/abs/):
<br>
\begin{align}
\frac{dX}{dt} & = \frac{V_L + V_R}{2} \cdot cos(\theta) \\
\frac{dY}{dt} & = \frac{V_L + V_R}{2} \cdot sen(\theta) \\
\frac{d \theta}{dt} & = \frac{V_L - V_R}{2} \\
\end{align}
<br>
Where ($X$,$Y$ and $\theta$) are the robot actual position and orientation angle in world reference frame. In simulation, we use the discrete form to build a model of the robot. The discrete form of the kinematic model is given by the following equations:<br>
<br>
\begin{align}
X_{k+1} & = X_k + T \cdot \frac{V_{lk} + V_{rk}}{2} \cdot cos(\theta_k + \frac{d \theta}{dt} ) \\
Y_{k+1} & = Y_k + T \cdot \frac{V_{lk} + V_{rk}}{2} \cdot sen(\theta_k + \frac{d \theta}{dt}) \\
\theta_{k+1} & = \theta_k + T \cdot \frac{V_{lk} + V_{rk}}{L} \\
\end{align}
<br>
where $X_{k+1}$ and $Y_{k+1}$ represent the position of the center axis of the mobile robot and $T$ is the sampling time.
```
class Pose:
"""
A class used to store the robot pose.
...
Attributes
----------
x : double
The x position of the robot on the map
y : double
The y position of the robot on the map
orientation : double
The angle theta of the robot on the map
Methods
-------
The class doesn't have any methods
"""
def __init__(self, x=None, y=None, orientation=None):
self.x = x
self.y = y
self.orientation = orientation
class Odometry():
"""
A class used to implement methods that allow a robot to calculate his own odometry.
...
Attributes
----------
robot : obj
The robot object
lastPose : obj Pose
Store the robot's pose during his movement
lastTimestamp : time
Store the last timestamp
left_vel : double
Store the velocity of the left robot wheel
right_vel : double
Store the velocity of the right robot wheel
delta_time : double
Store how much time has passed
delta_theta : double
Store how the orientation change
delta_space : double
Store how the (x,y) change
Methods
-------
ground_truth_updater()
Function to update the ground truth, the real pose of the robot at the simulator
odometry_pose_updater()
Function to estimate the pose of the robot based on the kinematic model
"""
def __init__(self, robot):
self.robot = robot
self.lastPose = None
self.lastTimestamp = time()
self.left_vel = 0
self.right_vel = 0
self.delta_time = 0
self.delta_theta = 0
self.delta_space = 0
def ground_truth_updater(self):
"""
Function to update the ground truth, the real pose of the robot at the simulator
"""
# get the (x,y,z) position of the robot at the simulator
pose = self.robot.get_current_position()
# get the orientation of the robot (euler angles)
orientation = self.robot.get_current_orientation()
# return an pose object (x,y,theta)
return Pose(x=pose[0], y=pose[1], orientation=orientation[2])
def odometry_pose_updater(self):
"""
Function to estimate the pose of the robot based on the knematic model
"""
if self.lastPose is None:
self.lastPose = self.ground_truth_updater()
return self.lastPose
# get the actual timestamp
time_now = time()
# get the robot linear velocity for the left and right wheel
left_vel, right_vel = self.robot.get_linear_velocity()
# calculate the difference between the acutal and last timestamp
delta_time = time_now - self.lastTimestamp
# calculate the angle deslocation - based on the kinematic model
delta_theta = (right_vel - left_vel) * (delta_time / self.robot.ROBOT_WIDTH)
# calculate the distance deslocation - based on the kinematic model
delta_space = (right_vel + left_vel) * (delta_time / 2)
# auxiliary function to sum angles
add_deltha = lambda start, delta: (((start+delta)%(2*math.pi))-(2*math.pi)) if (((start+delta)%(2*math.pi))>math.pi) else ((start+delta)%(2*math.pi))
# calculate the new X pose
x = self.lastPose.x + (delta_space * math.cos(add_deltha(self.lastPose.orientation, delta_theta/2)))
# calculate the new Y pose
y = self.lastPose.y + (delta_space * math.sin(add_deltha(self.lastPose.orientation, delta_theta/2)))
# calculate the new Orientation pose
theta = add_deltha(self.lastPose.orientation, delta_theta)
# uptade the state of the class
self.lastPose = Pose(x, y, theta)
self.lastTimestamp = time_now
self.left_vel = left_vel
self.right_vel = right_vel
self.delta_time = delta_time
self.delta_theta = delta_theta
self.delta_space = delta_space
return self.lastPose
```
# Defining the class that controls the robot walker
For this project we are going to use two different controllers in order to make the robot avoid obstacles in the map. The first one is a classical fuzzy based system, and the second one, is a more modern approach, based on artificial intelligence, called reinforcement learning.
### Controllers:
**1. Fuzzy**
Fuzzy logic is a very common technique in the Artificial Intelligence branch. It is a method that introduces the concept of partially correct and / or wrong, different from Boolean logic, in which only binary values are allowed. This fact allows generalist logic in which it is not necessary to deal with all possible cases, ideal for applications with limited memory and / or time.
A fuzzy control system is a control system based on fuzzy logic—a mathematical system that analyzes analog input values in terms of logical variables that take on continuous values between 0 and 1, in contrast to classical or digital logic, which operates on discrete values of either 1 or 0 (true or false, respectively).
The input variables in a fuzzy control system are in general mapped by sets of membership functions similar to this, known as "fuzzy sets". The process of converting a crisp input value to a fuzzy value is called "fuzzification". A control system may also have various types of switch, or "ON-OFF", inputs along with its analog inputs, and such switch inputs of course will always have a truth value equal to either 1 or 0, but the scheme can deal with them as simplified fuzzy functions that happen to be either one value or another. Given "mappings" of input variables into membership functions and truth values, the microcontroller then makes decisions for what action to take, based on a set of "rules", each of the form:
~~~
IF brake temperature IS warm AND speed IS not very fast
THEN brake pressure IS slightly decreased.
~~~
For this project the implemented fuzzy was very simples, and aims just to make the robot abble to avoid obstacles on his way. He uses the ultrassonic sensors for his three inputs (front, left and right distance) and outputs the linear velocity for both weels.
Some fundamental concepts to understand logic are:
1. **Degree of Relevance:** value in the range $[0,1]$ that determines the degree to which a given element belongs to a set, allowing a gradual transition from falsehood to truth.
2. **Fuzzy Set:** A set A in X is expressed as a set of ordered pairs: $A = {{(x, \mu_A (X)) | x \in X }}$.
3. **Fuzzy Rules:** are created to evaluate the antecedent (input) and apply the result to the consequent (output). They are partially activated depending on the antecedent.
4. **Fuzzy Steps:**
1. *Fuzification:* stage in which subjective linguistic variables and pertinence functions are defined.
2. *Inference:* stage at which rules are defined and evaluated.
3. *Defuzification:* step in which the resulting regions are converted to values for the system output variable. The best known methods of defuzification are: centroid, bisector, lowest of maximum (SOM), middle of maximum (MOM), highest of maximum (LOM).
Now, we are given a more detailed explanation about the implemented system, showing how we model the inputs, outputs, system rules and the defuzzify methods.
1. **Inputs and Outputs:** For this fuzzy we use three antecedents (all of them has the same shape of the one show bellow) and two consequents. The antecedents works mapping the left, right and front sensors of the ultrassonic sensors of the robot. As we can see, inputs are divide in three sets, low, medium and high distances that aims to tell the system how far he is from some object in the map. The consequents, by the other side, aims to mapping the velocity of both wheels of the robot, they are split in four velocities.
Fuzzy Antecedent | Fuzzy Consequent
:-------------------------:|:-------------------------:
 | 
2. **Rules:** The system is implemented using eleven rules, that we enough to make the robot able to escape from obstacles from the map with a stable control. The rules can be describe as follow:
~~~
left['low'] AND right['low'] AND (front['medium'] OR front['far'])),
output_left['low'], output_right['low']
left['low'] AND right['low'] AND front['low'],
output_left['reverse'], output_right['low']
left['medium'] OR left['far'] AND right['low'] AND front['low'],
output_left['low'], output_right['high']
left['medium'] OR left['far'] AND right['low'] AND front['medium'] OR front['far'],
output_left['low'], output_right['high']
left['far'] AND right['medium'] AND front['low'],
output_left['low'], output_right['high']
left['far'] AND right['far'] AND front['low'],
output_left['high'], output_right['low']
left['medium'] AND right['medium'] AND front['low'],
output_left['high'], output_right['low']
left['medium'] AND right['far'] AND front['low'],
output_left['high'], output_right['low']
left['low'] AND right['medium'] OR right['far'] AND front['low'],
output_left['high'], output_right['low']
left['low'] AND right['medium'] OR right['far'] AND front['medium'] OR front['far'],
output_left['high'], output_right['low']
left['medium'] OR left['far'] AND right['medium'] OR right['far'] AND front['medium'] OR front['far'],
output_left['medium'], output_right['medium']
~~~
3. **Defuzzification:** In order to understand how the choosen defuzzification afects the controller we test two different models, the minimum of the maximuns (SOM) and maximum of the maximuns(LOM), the consequences of this approach will be commented at the Results section.
```
class FuzzyControler():
"""
A class used to implement methods that allow a robot to walk, based on a fuzzy logic controller.
...
Attributes
----------
forward: skfuzzy object
Skfuzzy input object
left: skfuzzy object
Skfuzzy input object
right: skfuzzy object
Skfuzzy input object
output_left: skfuzzy object
Skfuzzy output object
output_right: skfuzzy object
Skfuzzy output object
rules: skfuzzy object
List of rules to the fuzzy
control: skfuzzy object
Skfuzzy controller object
simulator: skfuzzy object
Skfuzzy simulator object
Methods:
-------
create_inputs()
Function to create skfuzzy input functions
create_outputs()
Function to create skfuzzy output functions
create_rules()
Function to create skfuzzy rules
create_control()
Function to create skfuzzy controller
show_fuzzy()
Function to show the fuzzy rules as a graph
create_simulator()
Function that controls the fuzzy pipeline
simulate()
Function that give outputs velocity based on input distance
"""
def __init__(self, behavior):
self.front = None
self.left = None
self.right = None
self.output_left = None
self.output_right = None
self.rules = []
self.control = None
self.simulator = None
self.behavior = behavior
def create_inputs(self):
# set the variable universe as near, medium and far
self.front = ctrl.Antecedent(np.arange(0, 5.01, 0.01), 'front')
self.front['low'] = fuzz.trapmf(self.front.universe, [0, 0, 0.6, 1])
self.front['medium'] = fuzz.trimf(self.front.universe, [0.6, 1, 1.4])
self.front['far'] = fuzz.trapmf(self.front.universe, [1, 1.5, 5, 5])
self.left = ctrl.Antecedent(np.arange(0, 5.01, 0.01), 'left')
self.left['low'] = fuzz.trapmf(self.left.universe, [0, 0, 0.6, 1])
self.left['medium'] = fuzz.trimf(self.left.universe, [0.6, 1, 1.4])
self.left['far'] = fuzz.trapmf(self.left.universe, [1, 1.5, 5, 5])
self.right = ctrl.Antecedent(np.arange(0, 5.01, 0.01), 'right')
self.right['low'] = fuzz.trapmf(self.right.universe, [0, 0, 0.6, 1])
self.right['medium'] = fuzz.trimf(self.right.universe, [0.6, 1, 1.4])
self.right['far'] = fuzz.trapmf(self.right.universe, [1, 1.5, 5, 5])
return
def create_outputs(self):
self.output_left = ctrl.Consequent(np.arange(-1, 2.01, 0.1), 'output_left')
self.output_left['reverse'] = fuzz.trapmf(self.output_left.universe, [-1,-1, 0, 0.2])
self.output_left['low'] = fuzz.trimf(self.output_left.universe, [0,1, 1.3])
self.output_left['medium'] = fuzz.trimf(self.output_left.universe, [1,1.5, 1.75])
self.output_left['high'] = fuzz.trimf(self.output_left.universe, [1.2,1.8, 2])
self.output_left.defuzzify_method = 'lom'
self.output_right = ctrl.Consequent(np.arange(-1, 2.01, 0.1), 'output_right')
self.output_right['reverse'] = fuzz.trapmf(self.output_left.universe, [-1,-1, 0, 0.2])
self.output_right['low'] = fuzz.trimf(self.output_left.universe, [0,1, 1.3])
self.output_right['medium'] = fuzz.trimf(self.output_left.universe, [1,1.5, 1.75])
self.output_right['high'] = fuzz.trimf(self.output_left.universe, [1.2,1.8, 2])
self.output_right.defuzzify_method = 'lom'
return
def create_rules(self, front, left, right, output_left, output_right):
rule1 = ctrl.Rule(antecedent=(left['low'] & right['low'] & (front['medium'] | front['far'])),
consequent=(output_left['low'], output_right['low']))
rule2 = ctrl.Rule(antecedent=(left['low'] & right['low'] & front['low']),
consequent=(output_left['reverse'], output_right['low']))
rule3 = ctrl.Rule(antecedent=((left['medium'] | left['far']) & right['low'] & front['low']),
consequent=(output_left['low'], output_right['high']))
rule4 = ctrl.Rule(antecedent=((left['medium'] | left['far']) & right['low'] & (front['medium'] | front['far'])),
consequent=(output_left['low'], output_right['high']))
rule5 = ctrl.Rule(antecedent=(left['far'] & right['medium'] & front['low']),
consequent=(output_left['low'], output_right['high']))
rule6 = ctrl.Rule(antecedent=(left['far'] & right['far'] & front['low']),
consequent=(output_left['high'], output_right['low']))
rule7 = ctrl.Rule(antecedent=(left['medium'] & right['medium'] & front['low']),
consequent=(output_left['high'], output_right['low']))
rule8 = ctrl.Rule(antecedent=(left['medium'] & right['far'] & front['low']),
consequent=(output_left['high'], output_right['low']))
rule9 = ctrl.Rule(antecedent=(left['low'] & (right['medium'] | right['far']) & front['low']),
consequent=(output_left['high'], output_right['low']))
rule10 = ctrl.Rule(antecedent=(left['low'] & (right['medium'] | right['far']) & (front['medium'] | front['far'])),
consequent=(output_left['high'], output_right['low']))
rule11 = ctrl.Rule(antecedent=((left['medium'] | left['far']) & (right['medium'] | right['far']) & (front['medium'] | front['far'])),
consequent=(output_left['medium'], output_right['medium']))
for i in range(1, 12):
self.rules.append(eval("rule" + str(i)))
return
def create_control(self):
# call function to create robot input
self.create_inputs()
# call function to create robot output
self.create_outputs()
if self.behavior == "avoid_obstacle":
# call function to create rules
self.create_rules(self.front, self.left, self.right, self.output_left, self.output_right)
# create controller
self.control = skfuzzy.control.ControlSystem(self.rules)
return
def show_fuzzy(self):
if self.control is None:
raise Exception("Control not created yet!")
else:
self.control.view()
return
def create_simulator(self):
if self.control is None:
# crete controller if it doensn't exist
self.create_control()
# create simulator object
self.simulator = ctrl.ControlSystemSimulation(self.control)
return
def simulate(self, input_foward=None, input_left=None, input_right=None):
if self.simulator is None:
# crete simulator if it doensn't exist
self.create_simulator()
# if there is no input raise exception
if input_foward is None or input_left is None or input_right is None:
raise Exception("Inputs can't be none")
# simulte the robot linear velocity based on given inputs
self.simulator.input['front'] = input_foward
self.simulator.input['left'] = input_left
self.simulator.input['right'] = input_right
self.simulator.compute()
return self.simulator.output['output_left'], self.simulator.output['output_right']
```
**2. Reinforcement Learning**
Reinforcement learning is an area of Machine Learning. Reinforcement. It is about taking suitable action to maximize reward in a particular situation. Reinforcement learning differs from the supervised learning in a way that in supervised learning the training data has the answer key with it so the model is trained with the correct answer itself whereas in reinforcement learning, there is no answer but the reinforcement agent decides what to do to perform the given task. In the absence of training dataset, it is bound to learn from its experience.
**Main points in Reinforcement learning:**
1. Input: The input should be an initial state from which the model will start
2. Output: There are many possible output as there are variety of solution to a particular problem
3. Training: The training is based upon the input, The model will return a state and the user will decide to reward or punish the model based on its output.
4. The model keeps continues to learn.
5. The best solution is decided based on the maximum reward.
**Types of Reinforcement:**
1. Positive - is defined as when an event, occurs due to a particular behavior, increases the strength and the frequency of the behavior.
2. Negative - is defined as strengthening of a behavior because a negative condition is stopped or avoided.
One of the most common RL algorithms is the **Q-Learning**, a basic form which uses Q-values (also called action values) to iteratively improve the behavior of the learning agent. A brief introduction can be done with the following informations:
1. **Q-Values:** Q-values are defined for states and actions. $Q(S, A)$ is an estimation of how good is it to take the action A at the state S.
2. **Rewards and Episodes:** an agent over the course of its lifetime starts from a start state, makes a number of transitions from its current state to a next state based on its choice of action and also the environment the agent is interacting in. At every step of transition, the agent from a state takes an action, observes a reward from the environment, and then transits to another state.
3. **TD-Update:** the Temporal Difference or TD-Update rule can be represented as follows: $Q(S,A) = Q(S,A) + \alpha \cdot (R + \gamma \cdot (Q',S') - Q(S,A))$. Where the variables can be described as follow:
1. S: current state
2. A: current action
3. S': next state
4. A': next action
5. R: curren reward
6. $\gamma$: discounting factor for future rewards
7. $\alpha$: learning rate
4. **Choosing the Action:** the policy for choosing an action is very simple. It goes as follows :
1. with probability $(1-\epsilon)$ choose the action which has the highest Q-value
2. with probability $(\epsilon)$ choose any action at random
In order to implement the behavior for this project we need to make two main components, the enviroment (who communicate with the V-Rep interface) and the training function that implements the policy and the Q-Learning algorithm, the last one was done based on [this](https://www.geeksforgeeks.org/q-learning-in-python/) implementation.
Now, we are going to explain, with more details the enviroment and training implementations, both codes can be found [here](https://github.com/luizcartolano2/mc907-mobile-robotics/blob/project2/reinforcement_learning/environment.py) and [here](https://github.com/luizcartolano2/mc907-mobile-robotics/blob/project2/reinforcement_learning/train.py), and a function call can be seen bellow.
**Enviroment Implementation:**
The enviroment, on a Q-Learning situation, must have the hability to start and restart the simulation, to inform states and, more important, to take actions and identify the consequences (reward) of that actions.
1. **State:**
Since we aim to create a reinforcement behavior that "teaches" the robot how to avoid obstacles in the scene, we simply choose as a state the ultrassonic sensors observations, that is, the read distance in all of the robot sensors, what was implemented as follow:
~~~
observations = {}
observations['proxy_sensor'] = [np.array(self.read_ultrassonic_sensors())]
~~~
2. **Actions:**
The robot actions were limited to three options:
1. Walk straight: $[1.5,1.5]$.
2. Turn Left: $[0.5,1.5]$.
3. Turn Right: $[1.5,0.5]$.
3. **Reset Function:**
In order to training the model the code have to be able to restart the simulation at the start of every episode, in order to make it along the V-Rep simulator, it was necessary to implement a restart function, that stop the simultion and start it again, between both actions a delay is required in order to make sure the older simulation were completely killed. Also, the reset function has to return to the training function the intial state of the simulation.
This was done with the following lines of code:
~~~
stop = vrep.simxStopSimulation(self.clientID,vrep.simx_opmode_blocking)
time.sleep(5)
start = vrep.simxStartSimulation(self.clientID, vrep.simx_opmode_blocking)
observations = {}
observations['proxy_sensor'] = [np.array(self.read_ultrassonic_sensors())]
~~~
4. **Rewards model:**
The robot rewards were given based on the following situations:
1. Punishment to be close to objects:
``` (np.array(observations['proxy_sensor']) < 0.7).sum() * -2 ```
2. Punishment to be very close to objects:
``` (np.array(observations['proxy_sensor']) < 0.2).sum() * -10 ```
3. Rewarded for movement:
``` np.clip(np.sum(np.absolute(action)) * 2, 0, 2) ```
4. Reward for walking:
``` if dist > 0.1: reward['proxy_sensor'] += 50 ```
5. Punishment for dying:
``` if np.any(np.array(observations['proxy_sensor']) < 0.1): reward['proxy_sensor'] -= 100000 ```
5. **Step Update Function:**
The step/update function receive an action as input, perform and evalute it. At the end, it checks if the episode is done (the robot colide with something). Due to V-Rep problemns at implementation, we consider that, if any of the sensors were ten centimeters or less from an object, the robot is considered as "dead".
**Q-Learning Implementation:**
We can split the Q-Learning part in three main functionalities, as we follow explain:
1. **Action Policy:**
The action policy is the function to created to updade the action probabilities, it works as follow:
~~~
action_probabilities = np.ones(num_actions,dtype=float) * epsilon / num_actions
best_action = np.argmax(Q[state])
action_probabilities[best_action] += (1.0 - epsilon)
~~~
2. **Q-Learning Loop:**
The Q-Leanrning function iterate over the episodes, choosing actions based on their probabilities and updating the probabilities based on the TD-Rule and the state reward.
3. **Save/Load model:**
In order to save the model over many iterations and be able to keep improving the quality of the controller, at the end of every simulation, we create save the state/probability dictionary at a text file and, at the start of every simulation, we load that values into the dictionary.
#### Observation
Once that the reinforcement behavior were a test we develop it in a separate branch and using .py files, not a Jupyter Notebook, so here we are just going to show how to call the Q-Learning function, and to present the obtained results in a following section. Links to the implemented files are available to.
```
# create the simulation enviroment
env = Robot()
# calls the Q-Learning function
Q, stats = qLearning(env, 500)
# save the learned model
with open('model.txt', 'w') as f:
json.dump({str(k): str(tuple(v)) for k, v in Q.items()}, f)
# plot few results charts
plotting.plot_episode_stats(stats)
```
# Controls robot actions
A state machine, is a mathematical model of computation. It is an abstract machine that can be in exactly one of a finite number of states at any given time. The state machine can change from one state to another in response to some external inputs and/or a condition is satisfied, the change from one state to another is called a transition.
For this project, we implemented two behaviors with same goal, to avoid obstacles, but, the one who uses reinforcement learning were tested in a separate situation, so he isn't implemented at the state machine. The one that is implemented, the fuzzy controller, has two simple states, the one before he is initialized, and the one his working. For the firts stage we create all the used objects and transit to next stage. At the next stage, we read the sensors inputs, call the fuzzy simulator in order to get the outputs and change the robot velocities.
The fuzzy controller class has his own state machine implemented in order to create all the need behaviors, that is, it checks if all conditions are satisfied and, if not, create the antecedents, consequents, rules and put all together in a controller object, and then, it make the simulations.
```
def state_machine(behavior="avoid_obstacle"):
# stage
stage = 0
if behavior == "follow_wall":
raise Exception("Not implemented!")
elif behavior == "avoid_obstacle":
while True:
if stage == 0:
# first we create the robot and the walker object
robot = rb.Robot()
fuzzy = FuzzyControler(behavior=behavior)
# instantiate the odometry calculator
odometry_calculator = Odometry(robot=robot)
stage = 1
if stage == 1:
sensors = robot.read_ultrassonic_sensors()
front_sensors = min(sensors[3], sensors[4])
left_sensors = min(sensors[0], sensors[1], sensors[2])
right_sensors = min(sensors[5], sensors[6], sensors[7])
left_vel, right_vel = fuzzy.simulate(input_foward=front_sensors, input_left=left_sensors, input_right=right_sensors)
robot.set_left_velocity(left_vel)
robot.set_right_velocity(right_vel)
else:
raise Exception("Not implemented!")
```
# Main function - Execute the code here!
Here is a simple signal handler implement in order to make the simulator execution last for a given time period.
```
import signal
from contextlib import contextmanager
class TimeoutException(Exception): pass
@contextmanager
def time_limit(seconds):
def signal_handler(signum, frame):
raise TimeoutException("Timed out!")
signal.signal(signal.SIGALRM, signal_handler)
signal.alarm(seconds)
try:
yield
finally:
signal.alarm(0)
try:
ground_truth = []
odometry = []
lines = []
corners = []
points_kmeans = []
with time_limit(90):
state_machine()
except TimeoutException as e:
print("Timed out!")
```
# Results
In order to show the obtained results we are going to demonstrate a video with the running simulation for both behaviors. For the fuzzy system we are going to evaluate it beyond some different start pose situations and changing the input/output universe. The reinforcement learning, by the other side, has changes only at the enviroment, also, we are going to show how it performed as the episodes were growing.
An important observation we want to do respect about the topic that ask to evaluate the model using the ground truth position and the odometry one, since both controllers aims to avoid obstacles in the scene, and that, both of them have a react behavior, none of them uses the pose as input for the behavior, so it doesn't make any difference for the obtained results.
The robot can be seen in action at [youtube](https://youtu.be/uqNeEhWo0dA). And the video behavior will be following commented. Before go into the obtained results we are going to first explain how the tests were done. For both the fuzzy controller and the reinforcement learning.
### Fuzzy Controller
In order to test and understand the fuzzy behavior in a range of different situations, we create five different scenarios for it and test each of them in three different start poses at the scene. The start poses are show in the follow images:
First Pose | Second Pose | Third Pose
:-------------------------:|:-------------------------:|:-------------------------:
 |  | 
Beyond the different starting we change, for each experiment, or the shape/range of the fuzzy antecedents or the deffuzitication method. Each experiment can be described as follow:
1. **Experiment 1:**
For the first experiment the fuzzy set were the default one, and the deffuzitication method was the smallest of the maximuns (SOM).
2. **Experiment 2:**
For the second one, we expand the distance considered as "close" and still use the the smallest of the maximuns (SOM).
3. **Experiment 3:**
For the third one, we expand the distance considered as "medium" and still use the the smallest of the maximuns (SOM).
4. **Experiment 4:**
For the fourth one, we expand the distance considered as "medium", reduce the considered as "far" and still use the the smallest of the maximuns (SOM).
5. **Experiment 5:**
For the last one, we keep the default antecedent and change the deffuzification method for the largest of the maximuns (LOM).
The antecedent sets are show here:
Experiment 1 | Experiment 2 | Experiment 3 | Experiment 4 | Experiment 5
:-------------------------:|:-------------------------:|:-------------------------:|:-------------------------:|:-------------------------:
 |  |  |  | 
As we can observe in the [video](https://youtu.be/uqNeEhWo0dA), there isn't significant difference between the experiments 1, 3 e 4. The bigger ones happens when we change the "close" range of the input and, mainly, when we change the defuzzification method.
We can say that the experiment 1 show a robust and stable controller, where the robot is able to escape from obstacles with consistency and without taking major risks, besides that, he also walks with a good speed and do smooth movements.
For the second experiments we experience the worst fuzzy results. The robot achieves his goal of don't collide with other obstacles, but, he has sudden movements and once he considers that a lot of obstacles are close to him, a lot of bad decisions are taking, including moments when it basically turns on its own axis.
For the third experiment we have a closer situation for the first one, except for one aspect, he walks to close to the scene objects in a lot of situations.
The fourt and fifth experiments exhibit similar behaviors, they are consistent on their actions, don't take big risks, but they move to slow.
### Reinforcement Learning
The reinforcement learning was test on only one scenario, the same of Experiment 1. We achieve pretty good results, considering the lack of training time. After about 2500 episodes the robot were able, as we can observe in the video, to avoid the first obstacle. However, better results can be longed for, once that he doesn't survive a lot of time at the enviroment.
The scenario, and few charts describing the results are displayed below:
Enviroment | Episode Length | Episode Reward
:-------------------------:|:-------------------------:|:-------------------------:
 |  | 
As we expect, lenght and reward grow as the time passed, showing that we are on the right path.
# Discussion
Both controllers achieved satisfactory results, as expected at the start of the development.
### Fuzzy Controller
The fuzzy one, turned out to be a very good option when we talk about a reactive behavior, the implementation was quite easy and the results were satisfactory. Analysing the experiments we also could extract interesting insights. For example, we realize that increase the ```close``` or the ```medium``` intervals of the antecedents sets makes the robots moves less soft and he starts to take more risk and lowest consistent actions.
By the other side, consider less objects in a ```far``` distance and take the maximum of the maximuns as the defuzzification methods both made the robot movements more conservative, walking at a much lower speed. That behavior can be easily explained for both situations. For the first one, the robot starts to "imagine" that objects are close to him than he expects, so he acts more carefully. For the second one, once we start to defuzzify our set taking the maximum value, any sensor who make a ```low``` or ```medium``` read will overlap the other ones, and the robot will, again, act carefully.
Lastly, we decide that the first configuration (Experiment 1) of antecedents and defuzzification were the best one, and set it as the 'defaul' one. The choice was based on the robot's consistency and robustness. That results makes a lot of sense when we analyse how the controller were structered in the antecedent/defuzzication mix. That is, the antecedents were created in a way that the values tend to overlap so decisions aren't abrupt. Besides that, the defuzzification is a bit more confident, so, tends to higher speeds.
### Reinforcement Learning
The reinforcement approach showed to be an amazing approach for hard to note problems using artificial intelligence. As we saw on the charts of lenght and reward over the episodes, the idea of maximizing the reward obtained makes the robot learn with his mistakes, that is, the bad action take at some state. We didn't have time to train the robot over different scenarios, so he still not able, yet, to stay alive for a very long time in the enviroment. But, he can avoid some obstacles.
The rewards were modeled in a way that be close to an object has a negative return, a punishment. Also, make a move at the scene, walk, give a positive reward, a prize. By the end, the worst punish is die. Nevertheless, we believe that the way the state was defined wasn't the best possible and, possibly, is one of the reasons the results were not spectacular. An additional improvement that can be made in order to achieve better results is change the state for it to map only the front, left and right ultrassonic sensors.
Another problem faced was to determine when the robot had hit an object on the scene. The decision was check all the ultrassonic sensors and if any of then were ten centimeters or less from an object, we consider a hit. The decision fulfilled what was expected, but had some situations where wasn't clear if it really had hit something.
# Conclusions And Final Observations
In general, the work presented satisfactory results, especially considering that the objective of this work was to introduce the students to the concepts of fuzzy logic applies to robotics, being able to understand its operation and having the ability to evaluate solutions and know how to modify them for optimal results. It is also noteworthy that the results obtained by applying the solution to the proposed problem were significantly positive. Also, that was the first time ever working with reinforcement learning, that also presents very good results and it was really fun to work with.
Weaknesses of the work, which need to be improved in future iterations, are mostly related to how the fuzification criteria were defined. An improvement thought for the project is the use of genetic algorithms to choose the best ways to create the membership functions and to choose the fuzification method. For the reinforcement learning approach, we need to make a better model of the ```state-reward``` relation.
| github_jupyter |
Today we're going to walk through an example of predicting tumor and normal status directly from gene expression values. We'll be using the python package scikit learn to construct our SVM classifier. For machine learning, we highly recommend this package.
Lots of documentation is available:
http://scikit-learn.org/stable/documentation.html
We're going to be working on a support vector machine classifier. As we dig into the details, make sure you're referring to the documentation for more information:
http://scikit-learn.org/stable/modules/generated/sklearn.svm.SVC.html
```
import numpy as np
from sklearn import svm
from sklearn import preprocessing
# Define a useful helper function to read in our PCL files and store the gene names,
# matrix of values, and sample names
# We'll use this function later, but we don't need to dig into how it works here.
def read_dataset(filename):
data_fh = open(filename)
samples = data_fh.readline().strip().split('\t') # sample ids tab delimited
gids = [] # gene ids will be stored here
genes_samples = [] # genes x samples -- gene major
for line in data_fh:
toks = line.strip().split('\t')
gids.append(toks[0]) # add gene id
vals = [float(x) for x in toks[1:]]
zarray = preprocessing.scale(vals) # make each gene's expression values comparable
genes_samples.append(zarray)
data_fh.close()
#because we want samples x genes instead of genes x samples, we need to transpose
samples_genes = np.transpose(np.array(genes_samples))
return {'genes': gids, 'matrix': samples_genes, 'samples': samples}
# Use the function that we defined to read in our dataset
bric = read_dataset('../29_Data_ML-II/METABRIC_dataset.pcl')
# Now we need to figure out which samples in metabric are tumors and which are normal.
# We will store this in status_list (matching each example in the dataset), so that we
# can provide this to scikit learn's SVM implementation.
status = {} # hold tumor/normal status encoded as 1 (tumor)/2 (normal)
label_fh = open('tumor_normal_label.txt')
for line in label_fh:
toks = line.strip().split()
if toks[1] == 'Tumor':
status[toks[0]] = 1
elif toks[1] == 'Normal':
status[toks[0]] = 2
status_list = []
for sample in bric['samples']:
status_list.append(status[sample])
# Now we're going to construct a classifier. First we need to set up our parameters
svm_classifier = svm.SVC(C=0.000001, kernel='linear')
# Once our parameters are set, we can fit the classifier to our data
svm_classifier.fit(bric['matrix'], status_list)
# Once we have our classifier, we can apply it back to the examples and get our score
# Since this is binary classification. We get an accuracy.
score = svm_classifier.score(bric['matrix'], status_list)
print("Training Accuracy: " + str(score))
```
Congratulations! You've built your first SVM, and on training data it separates tumor data from normal data with over 90% accuracy! Now that we've done this with some biomedical data, let's take a step back and talk about things we should consider as we build a model.
_Q1: What are our labels?_
_Q2: What are our features?_
_Q3: What are our examples?_
### Overfitting in machine learning ###
When you train a computer to build a model that describes data that you've seen, a challenge known as "overfitting" can arise. When fitting the model, we want to find a model that fits the data as well as possible. However, real data is noisy. The model that fits data we have with the least error may capture the main features of the data, but may also capture noise in the data that we don't intend to model. When a model fits noise in training data, we call this problem overfitting.
For example, imagine that a professor wants to test a group of students' knowledge of calculus. She gives the students previous exam questions and answers to study. However, in the final exam, she uses the same questions to test the students. Some of the students could do very well because they memorized answers to the questions even though they don't understand calculus. The professor realizes this problem and then gives the students a new set of questions to test them. The students who memorized all the answers to previous exam questions may fail the new exam because they have no idea how to solve the new problems. We would say that those students have "overfit" to training data.
How can overfitting be a problem with machine learning? Don't we want the model to fit the data as well as possible? The reason is we want a model that captures the features that will also exist in some new data. If the model fits the noise in the data, the model will perform poorly on new data sets!
Let's use simulations to illustrate the overfitting problem. We are going to simulate two variables x and y and we let y = x + e, where e is some noise. That is, y is a linear function of x.
```
## Load necessary Python packages
import numpy as np # numpy makes it convenient to load/modify matrices of data
import sklearn.linear_model as lm # this scikit learn module has code to fit a line
import matplotlib.pyplot as plt # this lets us plot our results
from sklearn.metrics import mean_squared_error # we use this to see how well our model fits data
%matplotlib inline
# This code will make our data by adding random noise to a linear relationship
# Simulate two variables x and y
# y=x+e, e is some noise
x = np.linspace(0., 2, 10)
y = x + 0.5*np.random.randn(len(x))
```
Let's plot the data. The code in the box below will do this. As we can see, the relation between x and y is linear but with some random noise.
```
# This uses matplotlib to show points. You've seen a little bit of this before in the kmeans code
# We're using it for examples but you don't have to understand how this works.
# If you one day want to plot your results using python, you might want to keep this code
# as a reference.
plt.figure(figsize=(8,6))
plt.scatter(x[:100], y[:100])
plt.xlabel("x")
plt.ylabel("y")
#plt.plot(x, y)
```
Next, we want to train linear regression models on x and use the models to predict y. The models we are going to use are:
1. A simple linear regression model: Y~X
2. A complex multiple regression model: Y ~ X + X^2 + X^3 + X^4 ... + X^10
We want to choose the model that will most accurately predict y.
Let's use ski-learn to train these two models:
```
# You don't need to know how this code works. We're not going to focus on regression
# during this course. You may want to have it to refer to in the future.
### simple regression
lr = lm.LinearRegression()
lr.fit(x[:,np.newaxis], y);
y_lr = lr.predict(x[:, np.newaxis])
### multiple regression
lrp = lm.LinearRegression()
lrp.fit(np.vander(x, N=10, increasing=True), y)
y_lrp = lrp.predict(np.vander(x, N=10, increasing=True))
x_plot = np.linspace(0., 2, 1000)
y_plot = lrp.predict(np.vander(x_plot, N=10, increasing=True))
```
Let's plot the fitting results.
```
plt.figure(figsize=(8,6))
plt.scatter(x, y)
plt.plot(x, y_lr, 'g',label='Simple regression')
plt.title("Linear regression")
plt.plot(x_plot, y_plot,label='Multiple regression')
plt.legend(loc=2)
```
Let's calculate the MSE for simple regression model:
```
mean_squared_error(y, y_lr)
```
Let's calculate the MSE for multiple regression model:
```
mean_squared_error(y, y_lrp)
```
The multiple regression model fits the data perferlly (MSE is almost 0). The predicted values are the exact the same as the observed values since the prediction curve goes through every point. However, the simple regression model captures the linear relation between x and y but it didn't predict perfectlly well with the observed values. Then, shoud we choose multiple regression model rather than simple regression model since the former fitts the data much better than the latter?
_Q4: Which model do you think is the better model? Why?_
Remember that we want to find a model that fits the data well and, most importantly, can predict well on some new data. Let's simulate some new data and see the prediction performance of each model on the new data.
```
x_new = np.linspace(0., 2, 10)
y_new = x + 0.5*np.random.randn(len(x_new))
y_lr_new = lr.predict(x_new[:, np.newaxis])
y_lrp_new = lrp.predict(np.vander(x_new, N=10, increasing=True))
```
Let's plot the old models applied to the new data.
```
plt.figure(figsize=(8,6))
plt.scatter(x_new, y_new)
plt.plot(x, y_lr, 'g',label='Simple regression')
plt.title("Linear regression")
plt.plot(x_plot, y_plot,label='Multiple regression')
plt.legend(loc=2)
```
MSE for simple regression on new data:
```
mean_squared_error(y_new, y_lr_new)
```
MSE for multiple regression on new data:
```
mean_squared_error(y_new, y_lrp_new)
```
The multiple regression model will almost certainly perform worse than simple regression model on the new data (we don't know for sure in your case, because new data are simulated each time - check with your neighbors to see what they get as well, or feel free to clear and re-run the code to see another example). This is because the multiple regression model overfits the training data. It captures not only the true linear relation between x and y but also the random noise. However, simple regression only captures linear relation.
This also demonstrates that it is not a good idea to train and evaluate a model on the same data set. If so, we tend to choose the model that overfits the data. However, in real data analysis, you will occasionally see papers reporting nearly perfect model fitting results. If you look closely, you will find that the authors fit and evaluate the model on the same data set. You now know that this is a typical overfitting problem. In your future research, be careful with the overfitting problem when you try some machine learning models on your data!
To avoid overfitting, there are several methods. One is to use regularization in the model to reduce the model complexity. The other is to train the model on one dataset and evaluate the model on a separate dataset. For now, we'll cover evaluating on a separate dataset.
## Homework: BRCA Tumor/Normal - Revisited!
We are lucky enough to have an independent validation dataset of breast cancers from The Cancer Genome Atlas (TCGA). Let's see how our classifier does here!
```
# Let's read in the dataset and mark examples as tumor or normal depending on
# how they are annotated the sample description file (BRCA.547.PAM50.SigClust.Subtypes.txt)
tcga = read_dataset('../29_Data_ML-II/TCGA_dataset.pcl')
tcga_status = {} # hol tumor/normal status encoded as 1 (tumor)/2 (normal)
label_fh = open('BRCA.547.PAM50.SigClust.Subtypes.txt')
for line in label_fh:
toks = line.strip().split()
if toks[1] == 'tumor-adjacent normal':
tcga_status[toks[0]] = 2
else:
tcga_status[toks[0]] = 1
tcga_status_list = []
for sample in tcga['samples']:
tcga_status_list.append(tcga_status[sample])
# The first lines here are just the code from above copied down for convenience.
# Now we're going to construct a classifier. First we need to set up our parameters
svm_classifier = svm.SVC(C=0.000000001, kernel='linear')
# Once our parameters are set, we can fit the classifier to our data
svm_classifier.fit(bric['matrix'], status_list)
# Once we have our classifier, we can apply it back to the examples and get our score
# Since this is binary classification. We get an accuracy.
score = svm_classifier.score(bric['matrix'], status_list)
print("Training Accuracy: " + str(score))
# Ok - now let's apply our classifier from before to these data:
tcga_score = svm_classifier.score(tcga['matrix'], tcga_status_list)
print("Testing Accuracy: " + str(tcga_score))
```
_Q0: Run the code in the cell above this and report the training and testing accuracy observed with C = 0.000000001 (1 pt)_
_Q1: Do you think that your breast cancer classifier is under or overfitting your data? Why or why not? (3 pts)_
_Q2: Based on your answer to Q1, should you raise, lower, or keep C the same here? (1 pt)_
_Q3: Justify your answer to Q2 (3 pts)
_Q4: Try a different C. Report your training and testing accuracy (2 pts)._
| github_jupyter |
# TensorFlow: Convolutional NN
**Components Of The Model**
* Model Function (`cnn_model_fn`)
- accept features, class labels, mode and model params as args
- define the layers
- define a dictionary for output of predictions
- create `EstimatorSpec` object for the appropriate mode
+ train, predict, eval
+ create one-hot from class labels for train and eval
+ eval needs dict of metrics to use
* Main Function (`main`)
- accept mode and model params as args
- call a function to get data
+ load MNIST from TF in this case
- create the estimator with `cnn_model_fn` and model params
- create `*_input_fn` where `*` is the mode (e.g. train)
+ uses `numpy_input_fn` from the TF API for numpy data
- run the classifier/estimator using the appropriate mode
- perform any work necessary to display or return results
```
import numpy as np
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.INFO)
## Model Function
def cnn_model_fn(features, labels, mode, params):
"""Model function for CNN."""
# Input Layer
# Reshape X to 4-D tensor: [batch_size, width, height, channels]
# MNIST images are 28x28 pixels, and have one color channel
input_layer = tf.reshape(features["x"], [-1, 28, 28, 1])
# Computes 32 features using a 5x5 filter with ReLU activation
# Input Tensor Shape: [batch_size, 28, 28, 1]
# Output Tensor Shape: [batch_size, 28, 28, 32]
conv1 = tf.layers.conv2d(
inputs=input_layer,
filters=32,
kernel_size=[5, 5],
padding="same", # use "valid" to not preserve WxH
activation=tf.nn.relu)
# First max pooling layer with a 2x2 filter and stride of 2
# Input Tensor Shape: [batch_size, 28, 28, 32]
# Output Tensor Shape: [batch_size, 14, 14, 32]
pool1 = tf.layers.max_pooling2d(inputs=conv1, pool_size=[2, 2], strides=2)
# Computes 64 feature maps using a 5x5 filter.
# Input Tensor Shape: [batch_size, 14, 14, 32]
# Output Tensor Shape: [batch_size, 14, 14, 64]
conv2 = tf.layers.conv2d(
inputs=pool1,
filters=64,
kernel_size=[5, 5],
padding="same",
activation=tf.nn.relu)
# Input Tensor Shape: [batch_size, 14, 14, 64]
# Output Tensor Shape: [batch_size, 7, 7, 64]
pool2 = tf.layers.max_pooling2d(inputs=conv2, pool_size=[2, 2], strides=2)
# Flatten tensor into a batch of vectors for input to dense layer
pool2_flat = tf.reshape(pool2, [-1, 7 * 7 * 64])
# Input Tensor Shape: [batch_size, 7 * 7 * 64]
# Output Tensor Shape: [batch_size, 1024]
dense = tf.layers.dense(inputs=pool2_flat, units=1024, activation=tf.nn.relu)
# Add dropout operation; 0.6 probability that element will be kept
dropout = tf.layers.dropout(
inputs=dense, rate=0.4, training=mode == tf.estimator.ModeKeys.TRAIN)
# Logits layer
# Input Tensor Shape: [batch_size, 1024]
# Output Tensor Shape: [batch_size, 10]
logits = tf.layers.dense(inputs=dropout, units=10)
# this dict will be returned for predictions
predictions = {
# actual class predictions
"classes": tf.argmax(input=logits, axis=1),
# class probabilities from softmax on logits
"probabilities": tf.nn.softmax(logits, name="softmax_tensor")
}
if mode == tf.estimator.ModeKeys.PREDICT:
return tf.estimator.EstimatorSpec(mode=mode, predictions=predictions)
# Calculate loss (for both TRAIN and EVAL modes)
onehot_labels = tf.one_hot(indices=tf.cast(labels, tf.int32), depth=10)
loss = tf.losses.softmax_cross_entropy(
onehot_labels=onehot_labels, logits=logits)
# Configure the training Op (for TRAIN mode)
if mode == tf.estimator.ModeKeys.TRAIN:
optimizer = tf.train.GradientDescentOptimizer(learning_rate=0.001)
train_op = optimizer.minimize(
loss=loss,
global_step=tf.train.get_global_step())
return tf.estimator.EstimatorSpec(mode=mode, loss=loss, train_op=train_op)
# evaluation metrics (for EVAL mode)
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(
labels=labels, predictions=predictions["classes"])}
return tf.estimator.EstimatorSpec(
mode=mode, loss=loss, eval_metric_ops=eval_metric_ops)
## Main Function
def main(mode='train', model_params={'learning_rate': 0.001}):
# Load training and eval data
mnist = tf.contrib.learn.datasets.load_dataset("mnist")
train_data = mnist.train.images # Returns np.array
train_labels = np.asarray(mnist.train.labels, dtype=np.int32)
eval_data = mnist.test.images # Returns np.array
eval_labels = np.asarray(mnist.test.labels, dtype=np.int32)
# Create the Estimator
mnist_classifier = tf.estimator.Estimator(
model_fn=cnn_model_fn,
params=model_params,
model_dir="/tmp/mnist_convnet_model")
# Train the model
if mode == 'train':
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": train_data},
y=train_labels,
batch_size=100,
num_epochs=10,
shuffle=True)
mnist_classifier.train(
input_fn=train_input_fn)
elif mode == 'predict':
predict_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": eval_data},
num_epochs=1,
shuffle=False)
preds = mnist_classifier.predict(
input_fn=predict_input_fn)
return np.array([p for p in preds])
elif mode == 'eval':
# Evaluate the model and print results
eval_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": eval_data},
y=eval_labels,
num_epochs=1,
shuffle=False)
eval_results = mnist_classifier.evaluate(input_fn=eval_input_fn)
print(eval_results)
main()
main(mode='eval')
preds = main(mode='predict')
preds[0]
```
| github_jupyter |
<a href="https://colab.research.google.com/github/aly202012/Teaching/blob/master/Clustering.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
```
# clustaring
```
هو شكل من أشكال التعلم الآلي غير الخاضع للإشراف حيث يتم تجميع الملاحظات في مجموعات بناءً على أوجه التشابه في قيم البيانات أو الميزات. يعتبر هذا النوع من التعلم الآلي غير خاضع للإشراف لأنه لا يستخدم قيم التسمية المعروفة سابقًا لتدريب نموذج ؛ في نموذج التجميع ، يكون الملصق هو الكتلة التي يتم تعيين الملاحظة إليها ، بناءً على ميزاتها فقط.
```
import pandas as pd
# load the training dataset
data = pd.read_csv('Seed_Data.csv')
# Display a random sample of 10 observations (just the features)
# هذا الكود يظهر بالضبط الابعاد الحقيقيه للبيانات وهي 9 اعمده
#features = data[data.columns[0:]]
# هذا الكود يستخدم للفصل بين الهدف والسمات الخاصه به
features = data[data.columns[0:6]]
#features.sample(10)
features.head(10)
#data.shape
# (210, 8)
```
الآن ، بالطبع يصعب تصور الفضاء السداسي الأبعاد في عالم ثلاثي الأبعاد ، أو في مخطط ثنائي الأبعاد ؛ لذلك سنستفيد من تقنية رياضية تسمى تحليل المكونات الرئيسية (PCA) لتحليل العلاقات بين الميزات وتلخيص كل ملاحظة على أنها إحداثيات لمكونين رئيسيين - وبعبارة أخرى ، سنقوم بترجمة قيم الميزة سداسية الأبعاد إلى إحداثيات ثنائية الأبعاد.
```
from sklearn.preprocessing import MinMaxScaler
from sklearn.decomposition import PCA
# Normalize the numeric features so they're on the same scale
scaled_features = MinMaxScaler().fit_transform(features[data.columns[0:6]])
# Get two principal components
pca = PCA(n_components=2).fit(scaled_features)
features_2d = pca.transform(scaled_features)
features_2d[0:10]
# نلاحظ تغير في الابعاد للبيانات
#features_2d.shape
# (210, 2)
# تم التحويل الي ثلاثي الابعاد حتي يسهل رسمهم
import matplotlib.pyplot as plt
%matplotlib inline
plt.scatter(features_2d[:,0],features_2d[:,1])
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Data')
plt.show()
#importing the libraries
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
%matplotlib inline
# Create 10 models with 1 to 10 clusters
wcss = []
for i in range(1, 11):
kmeans = KMeans(n_clusters = i)
# Fit the data points
kmeans.fit(features.values)
# Get the WCSS (inertia) value
wcss.append(kmeans.inertia_)
#Plot the WCSS values onto a line graph
plt.plot(range(1, 11), wcss)
plt.title('WCSS by Clusters')
plt.xlabel('Number of clusters')
plt.ylabel('WCSS')
plt.show()
# هذا الرسم البياني يظهر امكانيه تقسيمنا للبيانات الي 3 مجموعات فقط بالطبع يمكننا التقسيم الي عدد مجموعات اكثر ولكن هذا غير مناسب
# وذلك حسب نظريه الكوع
# التالي هو استخدام تقنيه للتجميع تسمي بال كا مين
from sklearn.cluster import KMeans
# Create a model based on 3 centroids
model = KMeans(n_clusters=3, init='k-means++', n_init=100, max_iter=1000)
# Fit to the data and predict the cluster assignments for each data point
km_clusters = model.fit_predict(features.values)
# View the cluster assignments
km_clusters
# حدث التقسيم ثلاثي الابعاد الان لنري الرسم البياني
def plot_clusters(samples, clusters):
col_dic = {0:'blue',1:'green',2:'orange'}
mrk_dic = {0:'*',1:'x',2:'+'}
colors = [col_dic[x] for x in clusters]
markers = [mrk_dic[x] for x in clusters]
for sample in range(len(clusters)):
plt.scatter(samples[sample][0], samples[sample][1], color = colors[sample], marker=markers[sample], s=100)
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Assignments')
plt.show()
plot_clusters(features_2d, km_clusters)
```
في بعض الأحيان ، يتم استخدام التجميع كخطوة أولية نحو إنشاء نموذج تصنيف. تبدأ بتحديد مجموعات مميزة من نقاط البيانات ، ثم تقوم بتعيين تسميات فئة لتلك المجموعات. يمكنك بعد ذلك استخدام هذه البيانات المصنفة لتدريب نموذج التصنيف.
```
seed_species = data[data.columns[7]]
plot_clusters(features_2d, seed_species.values)
```
المجموعات الهرمية¶
تقدم طرق التجميع الهرمي افتراضات توزيعية أقل عند مقارنتها بطرق K-mean. ومع ذلك ، فإن طرق K-mean بشكل عام أكثر قابلية للتوسع ، وأحيانًا تكون كثيرة جدًا.
ينشئ التجميع الهرمي مجموعات إما بطريقة تقسيمية أو بطريقة تكتلية. طريقة الانقسام هي نهج "من أعلى لأسفل" يبدأ بمجموعة البيانات بأكملها ثم إيجاد الأقسام بطريقة متدرجة. التجميع التجميعي هو نهج "من الأسفل إلى الأعلى **. في هذا المعمل ستعمل مع المجموعات التراكمية التي تعمل تقريبًا على النحو التالي:
يتم حساب مسافات الربط بين كل نقطة من نقاط البيانات.
يتم تجميع النقاط بشكل ثنائي مع أقرب جار لها.
يتم حساب مسافات الربط بين المجموعات.
يتم دمج المجموعات زوجيًا في مجموعات أكبر.
يتم تكرار الخطوتين 3 و 4 حتى تصبح جميع نقاط البيانات في مجموعة واحدة.
يمكن حساب وظيفة الربط بعدة طرق:
يقيس ربط الأقسام الزيادة في التباين للمجموعات المرتبطة ،
متوسط الارتباط يستخدم متوسط المسافة الزوجية بين أعضاء المجموعتين ،
يستخدم الارتباط الكامل أو الأقصى الحد الأقصى للمسافة بين أعضاء المجموعتين.
تُستخدم عدة مقاييس مختلفة للمسافة لحساب وظائف الربط:
المسافة الإقليدية أو l2 هي الأكثر استخدامًا. هذا المقياس هو الخيار الوحيد لطريقة ربط وارد.
تعتبر مسافة مانهاتن أو إل 1 قوية بالنسبة للقيم المتطرفة ولها خصائص أخرى مثيرة للاهتمام.
تشابه جيب التمام هو حاصل الضرب النقطي بين متجهات الموقع مقسومًا على مقادير المتجهات. لاحظ أن هذا المقياس هو مقياس للتشابه ، في حين أن المقياسين الآخرين هما مقياس الاختلاف. يمكن أن يكون التشابه مفيدًا جدًا عند العمل مع بيانات مثل الصور أو المستندات النصية.
```
# Agglomerative Clustering
# ويسمي ايضا بالتجميع التكتلي او العنقودي
from sklearn.cluster import AgglomerativeClustering
agg_model = AgglomerativeClustering(n_clusters=3)
agg_clusters = agg_model.fit_predict(features.values)
agg_clusters
import matplotlib.pyplot as plt
%matplotlib inline
def plot_clusters(samples, clusters):
col_dic = {0:'blue',1:'green',2:'orange'}
mrk_dic = {0:'*',1:'x',2:'+'}
colors = [col_dic[x] for x in clusters]
markers = [mrk_dic[x] for x in clusters]
for sample in range(len(clusters)):
plt.scatter(samples[sample][0], samples[sample][1], color = colors[sample], marker=markers[sample], s=100)
plt.xlabel('Dimension 1')
plt.ylabel('Dimension 2')
plt.title('Assignments')
plt.show()
plot_clusters(features_2d, agg_clusters)
```
| github_jupyter |
2D surface fields of annual mean temperature, salinity, DIN, Chl-a(Region
0-10 S,35 -50 E) for 2005. For Chl-a, two types of figures: using linear and log scale of chlorophyll. For each variable, it was calculated the area-averaged value taking into account that the grid is irregular.
```
#import modules
#allows plots to appear beneath cell
%matplotlib notebook
import numpy as np
import pandas as pd
import numpy.ma as ma
import netCDF4 as nc4
from netCDF4 import Dataset
import matplotlib.pyplot as plt
from matplotlib import gridspec
import matplotlib.colors as colors
from matplotlib import colors, ticker, cm
import matplotlib.patches as mpatches
import cartopy as cart
from mpl_toolkits.basemap import Basemap
import xarray as xrr
from numpy.ma import masked_where
#load in monthly SST and SSS data from 2005:
m_list = ['01', '02', '03', '04', '05', '06', '07', '08', '09', '10', '11', '12']
sst=np.zeros((120,180,12))
sss=np.zeros((120,180,12))
va=0
for m in m_list:
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/means/2005/ORCA0083-N06_2005m'+str(m)+'T.nc','r',format='NETCDF4')
sst_=fyd.variables['sst'][0,1373:1493,3865:4045]
sst[:,:,va]=sst_
sss_=fyd.variables['sss'][0,1373:1493,3865:4045]
sss[:,:,va]=sss_
va=va+1
sst[np.abs(sst) > 3000.] = np.nan
sst[sst == 0.] = np.nan
sss[np.abs(sss) > 3000.] = np.nan
sss[sss == 0.] = np.nan
#load of Dissolved Inorganic Nitrogen (DIN) and Chl-a concentration (CHD and CHN) data:
m_list = ['01', '02', '03', '04', '05', '06', '07', '08', '09', '10', '11', '12']
din=np.zeros((120,180,12))
chlND=np.zeros((120,180,12))
chlD=np.zeros((120,180,12))
va=0
for m in m_list:
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/medusa/2005/ORCA0083-N06_2005m'+str(m)+'P.nc','r',format='NETCDF4')
din_=fyd.variables['DIN'][0,0,1373:1493,3865:4045]
din[:,:,va]=din_
chl_nd=fyd.variables['CHN'][0,0,1373:1493,3865:4045]
chlND[:,:,va]=chl_nd
chl_d=fyd.variables['CHD'][0,0,1373:1493,3865:4045]
chlD[:,:,va]=chl_d
va=va+1
din[np.abs(din) > 3000.] = np.nan
din[din == 0.] = np.nan
chlND[np.abs(chlND) > 3000.] = np.nan
chlND[chlND == 0.] = np.nan
chlD[np.abs(chlD) > 3000.] = np.nan
chlD[chlD == 0.] = np.nan
chl_a = chlND + chlD
#din=np.zeros((1,1,120,179,12))
#chl=np.zeros((1,1,120,179,12))
latS=fyd.variables['nav_lat'][1373:1493,3865:4045]
lonS=fyd.variables['nav_lon'][1373:1493,3865:4045]
#load in bathymetry
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/domain/bathymetry_ORCA12_V3.3.nc','r',format='NETCDF4')
bathy=fyd.variables['Bathymetry'][1373:1493,3865:4045]
# load in mask and set land to = 1 and ocean = masked for land contour.
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/domain/mask.nc','r',format='NETCDF4')
mask=fyd.variables['tmask'][0,0,1373:1493,3865:4045]
maskc=mask
mask = masked_where(np.abs(mask) > 0, mask)
mask[mask == 0] = 1
#SST annual mean
sst_m=np.nanmean(sst,axis=2)
#SSS annual mean
sss_m=np.nanmean(sss,axis=2)
#DIN annual mean
din_m=np.nanmean(din,axis=2)
#CHL annual mean
chl_m=np.nanmean(chl_a,axis=2)
#2D annual mean SST surface field
ticks=[26.75, 27, 27.25, 27.5, 27.75, 28, 28.25, 28.5]
low=26.75
high=28.5
ran=0.025
plt.figure(figsize=(7, 5))
plt.plot(111)
P1 = plt.contourf(lonS,latS,sst_m,np.arange(low, high, ran),extend='both',cmap=plt.cm.spectral)
plt.contourf(lonS,latS,mask,vmin=0, vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1)
#plt.contour(lonS,latS,bathy,colors='k',levels=[200])
plt.contour(lonS,latS,maskc,colors='k',levels=[0], linewidths=0.5)
cbar.set_ticks(ticks)
cbar.set_label('$^\circ$ C',rotation=0)
plt.title('Annual Mean SST - 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
#2D annual mean SSS surface field
ticks=[33.25, 33.5, 33.75, 34, 34.25, 34.5, 34.75, 35, 35.25, 35.5, 35.75, 36]
low=33.25
high=36
ran=0.025
plt.figure(figsize=(7, 5))
plt.plot(111)
P1 = plt.contourf(lonS,latS,sss_m,np.arange(low, high, ran),extend='both',cmap=plt.cm.spectral)
plt.contourf(lonS,latS,mask,vmin=0, vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1)
#plt.contour(lonS,latS,bathy,colors='k',levels=[200])
plt.contour(lonS,latS,maskc,colors='k',levels=[0], linewidths=0.5)
cbar.set_ticks(ticks)
cbar.set_label('g/kg',rotation=0,labelpad=12)
plt.title('Annual Mean SSS - 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
#2D annual mean DIN surface field
ticks=[0.04, 0.06, 0.10, 0.14, 0.18,
0.22, 0.26, 0.3, 0.34, 0.38, 0.42, 0.46, 0.5, 0.54,0.58]
low=0.052
high=0.61
ran=0.01
plt.figure(figsize=(7, 5))
plt.plot(111)
P1 = plt.contourf(lonS,latS,din_m,np.arange(low, high, ran),extend='both',cmap=plt.cm.spectral)
plt.contourf(lonS,latS,mask,vmin=0, vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1)
#plt.contour(lonS,latS,bathy,colors='k',levels=[200])
plt.contour(lonS,latS,maskc,colors='k',levels=[0], linewidths=0.5)
cbar.set_ticks(ticks)
cbar.set_label('mmol N/m$^3$',rotation=-90,labelpad=14)
plt.title('Annual Mean DIN - 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
#2D annual mean Chl-a surface field
ticks=[0.05, 0.1, 0.15, 0.2, 0.25, 0.3, 0.35, 0.4, 0.45, 0.5, 0.55, 0.6]
low=0.05
high=0.63
ran=0.01
plt.figure(figsize=(7, 5))
plt.plot(111)
P1 = plt.contourf(lonS,latS,chl_m,np.arange(low, high, ran),extend='both',cmap=plt.cm.spectral)
plt.contourf(lonS,latS,mask,vmin=0, vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1)
#plt.contour(lonS,latS,bathy,colors='k',levels=[200])
plt.contour(lonS,latS,maskc,colors='k',levels=[0], linewidths=0.5)
cbar.set_ticks(ticks)
cbar.set_label('mg/m$^3$',rotation=-90,labelpad=14)
plt.title('Annual Mean Chl-a 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
#For Chl-a, also a logaritmic plot
from matplotlib.colors import LogNorm
from matplotlib import ticker, cm
low=0.065
high=0.65
ran=0.01
plt.figure(figsize=(5, 4))
plt.plot(111)
#levs = np.logspace(-1.2,0.3,6)
levs = np.logspace(-1.11,-0.21,7)
P1 = plt.contourf(lonS,latS,chl_m,levs,norm=LogNorm())
plt.contourf(lonS,latS,mask,vmin=0,vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1,orientation='vertical',
norm=LogNorm(),ticks=[8e-2,1e-1,2.5e-1,6e-1])
#cbar.ax.set_yticklabels(['{:.0e}'.format(x) for x in levs])
cbar.set_label('mg/m$^3$',rotation=0,labelpad=14)
plt.title('Annual Mean Chl-a (log), 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
from matplotlib.colors import LogNorm
import matplotlib.ticker as ticker
low=0.065
high=0.65
ran=0.01
plt.figure(figsize=(5, 4))
plt.plot(111)
plt.rcParams['text.usetex'] = True
levs = np.logspace(-1.11,-0.21,8)
P1 = plt.contourf(lonS,latS,log)#levs norm=LogNorm())
plt.contourf(lonS,latS,mask,vmin=0, vmax=2,cmap=plt.cm.Greys)
cbar = plt.colorbar(P1,orientation='vertical',spacing='proportional',
norm=LogNorm(),format='$10^{%.2f}$',
ticks=[-1.11,-1,-0.75,-0.5,-0.25, -0.2])
cbar.set_label('mg/m$^3$',rotation=0,labelpad=13)
plt.title('Annual Mean Chl-a (log), 2005')
plt.ylim((-10,0))
plt.xlim((35,50))
plt.xlabel('E Longitude')
plt.ylabel('S Latitude')
#For each variable, the area-averaged value
#taking into account that the grid is irregular.
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/domain/mesh_hgr.nc','r',format='NETCDF4')
e1t=fyd.variables['e1t'][0,1373:1493,3865:4045]
e2t=fyd.variables['e2t'][0,1373:1493,3865:4045]
areas = e1t*e2t
fyd=nc4.Dataset('/group_workspaces/jasmin2/nemo/vol1/ORCA0083-N006/domain/mask.nc','r',format='NETCDF4')
mask = fyd.variables['tmask'][0,0,1373:1493,3865:4045]
m_area=areas*mask
area_tot=np.sum(m_area,dtype=np.float32)
#sst
average_sst = np.nansum(m_area*sst_m)/area_tot
print 'The sst area averaged value for this region in 2005 is:\n',average_sst,'°C'
#sss
average_sss = np.nansum(m_area*sss_m)/area_tot
print 'The sss area averaged value for this region in 2005 is:\n',average_sss,'g/kg'
#din
average_din = np.nansum(m_area*din_m)/area_tot
print 'The DIN area averaged value for this region in 2005 is:\n',average_din,'mmol N/m³'
#chl-a
average_chl = np.nansum(m_area*chl_m)/area_tot
print 'The Chl-a area averaged value for this region in 2005 is:\n',average_chl,'mg/m³'
```
| github_jupyter |
## Diagnosis visualization
- We will use the ACT ontology, and we will use a mapping file that will allow us to go from ICD9CM to ICD10CM
- The file used ACT_ICD10_ICD9_3_colmns_actual_nodeName.csv is available in the shared folder.
```
######################
## ICD descriptions ##
######################
ncats <- read.delim( file = "./ACT_ICD10_ICD9_3_colmns_actual_nodeName.csv",
sep = ",",
colClasses = "character",
header = TRUE)
#clean the file removing those that are empty or those that belongs to groups of diagnostics
ncats <- ncats[ ncats$C_BASECODE != "", ]
ncats <- ncats[- grep("-", ncats$C_BASECODE), ]
#split the diagnosis code column to have on one side the code and in the other the version
ncats$icd_code <- sapply(strsplit( as.character(ncats$C_BASECODE), "[:]"), '[', 2)
ncats$icd_version <- sapply(strsplit( as.character(ncats$C_BASECODE), "[:]"), '[', 1)
ncats$icd_version <- gsub( "CM", "", ncats$icd_version)
ncats$icd_version <- gsub( "ICD", "", ncats$icd_version)
#create an additional column where we remove the dots from all the codes, since some sites are
#providing the codes without dots
ncats$codeNoDots <- gsub("[.]", "", ncats$icd_code)
#create an additional column with the highest level of the hierarchy, to be able to group
#the different diagnosis in around 20 different categories
ncats$Category <- sapply(strsplit( as.character(ncats$C_FULLNAME), "[\\]"), '[', 3)
ncats$Category <- sapply(strsplit( as.character(ncats$Category), "[(]"), '[', 1)
ncats$Category <- trimws(ncats$Category)
#map from ICD9 to 10 based on the hierarchy
icd10 <- ncats[ ncats$icd_version == 10, ]
icd9 <- ncats[ ncats$icd_version == 9, ]
icd10$description <- sapply(strsplit( as.character(icd10$C_FULLNAME), "[\\]"), tail, 1)
icd9$description <- sapply(strsplit( as.character(icd9$C_FULLNAME), "[\\]"), '[',
length(unlist(strsplit(icd9$C_FULLNAME, "[\\]")))-1)
for( i in 1:nrow(icd9) ){
lng <- length(unlist(strsplit(icd9$C_FULLNAME[i], "[\\]")))
icd9$description[i] <- sapply(strsplit( as.character(icd9$C_FULLNAME)[i], "[\\]"), '[', lng-1)
}
totalDiagnosis <- rbind( icd9, icd10)
#####################
## Diagnosis files ##
#####################
diagnosisCombined <- read.csv(file = "./Diagnoses-Combined200405.csv",
sep = ",",
header = TRUE,
colClasses = "character")
diagnosisCombined <- diagnosisCombined[, c("siteid", "icd_code", "icd_version", "num_patients")]
diagnosisCombined$perc_patients <- round(100*(as.numeric( diagnosisCombined$num_patients ) / 15637 ),2)
diagnosisPerCountry <- read.delim(file = "./Diagnoses-CombinedByCountry200405.csv",
sep = ",",
header = TRUE,
colClasses = "character")
diagnosisPerCountry <- diagnosisPerCountry[, c("siteid", "icd_code", "icd_version", "num_patients") ]
demographicsPerCountry <- read.delim(file = "./Demographics-CombinedByCountry200405.csv",
sep = ",",
header = TRUE,
colClasses = "character")
demographicsPerCountry <- demographicsPerCountry[demographicsPerCountry$sex == "All",
c("siteid", "total_patients") ]
diagnosisPerCountry$siteid <- as.character( diagnosisPerCountry$siteid )
diagnosisPerCountry <- merge( diagnosisPerCountry, demographicsPerCountry, by = "siteid")
diagnosisPerCountry$perc_patients <- round(100*(as.numeric( diagnosisPerCountry$num_patients ) / as.numeric( diagnosisPerCountry$total_patients )),2)
diagnosisPerCountry <- diagnosisPerCountry[, c("siteid", "icd_code", "icd_version", "num_patients", "perc_patients")]
diagnosis <- rbind( diagnosisCombined, diagnosisPerCountry)
###########################################################################
### Filter the diagnosis by number and percentage of affected patients ###
##########################################################################
numberOfPatientsFilter = 10
selection <- diagnosis[ as.numeric( diagnosis$num_patients) >= numberOfPatientsFilter, ]
percentageFilter = 0.1
selection <- diagnosis[ as.numeric( diagnosis$perc_patients) > percentageFilter, ]
##################################################
## Heatmap with ICD description and categories ##
#################################################
icdMapping <- totalDiagnosis[, c("codeNoDots", "Category", "description")]
icdMapping <- icdMapping[ ! duplicated( icdMapping ), ]
selection$codeNoDots <- gsub("[.]", "", selection$icd_code)
selectionDesc <- merge( selection, icdMapping, all.x = TRUE,by = "codeNoDots")
selectionDesc$lbl <- ifelse( is.na(selectionDesc$description), as.character(selectionDesc$icd_code),
as.character(selectionDesc$description))
selectionDesc$Category <- factor(selectionDesc$Category, levels = c("Certain conditions originating in the perinatal period",
"Congenital malformations, deformations and chromosomal abnormalities",
"Symptoms, signs and abnormal clinical and laboratory findings, not elsewhere classified",
"Certain infectious and parasitic diseases",
"Factors influencing health status and contact with health services",
"Pregnancy, childbirth and the puerperium",
"Diseases of the nervous system",
"Diseases of the blood and blood-forming organs and certain disorders involving the immune mechanism",
"Diseases of the circulatory system",
"Neoplasms",
"Diseases of the musculoskeletal system and connective tissue" ,
"Injury, poisoning and certain other consequences of external causes",
"Diseases of the digestive system",
"Diseases of the respiratory system",
"Diseases of the genitourinary system",
"Endocrine, nutritional and metabolic diseases",
"Diseases of the ear and mastoid process",
"Diseases of the eye and adnexa",
"Mental and behavioral disorders",
"Diseases of the skin and subcutaneous tissue",
"External causes of morbidity"),
labels = c("Perinatal",
"Congenital&Chromosomal",
"Symptoms",
"Infectious",
"Factors influencing health status",
"Pregnance, childbirth",
"Nervous",
"Blood",
"Circulatory",
"Neoplams",
"Musculoskeletal",
"Injury,poisoning",
"Digestive",
"Respiratory",
"Genitourinary",
"Endocrine/metabolic",
"Ear and mastoid",
"Eye and adnexa",
"Mental",
"Skin",
"External causes"))
###########################
## Heatmap representation #
###########################
ggplot(data = selectionDesc, aes(x = lbl, y = siteid, alpha=perc_patients)) +
scale_alpha(range = c(0.5, 1))+
geom_tile(aes(fill = siteid), colour = "white") +
facet_grid(. ~ Category, scales = "free", switch = "x")+
scale_fill_manual(values = c("Italy" = "#009E73", "France" = "#0072B2", "Germany" = "#E69F00", "USA" = "#D55E00", "Combined" = "#444444")) +
theme_bw() +
theme(panel.grid=element_blank(), axis.text.y = element_text(size=5),plot.title = element_text(size=7),
axis.title.x=element_blank(),
axis.text.x=element_blank(),
axis.ticks.x=element_blank(),
strip.text.x = element_text(angle = 90, size = 5)) +
coord_cartesian(expand=FALSE)+
labs(title = "Diagnoses (Date 2020-04-05) | 12 sites | 15,637 patients ")
##################################################
## Heatmap showing the specific diagnosis codes ##
##################################################
ggplot(selectionDesc, aes(y=description, x=siteid, fill=siteid, alpha=perc_patients)) +
geom_tile() +
scale_fill_manual(values = c("Italy" = "#009E73", "France" = "#0072B2", "Germany" = "#E69F00", "USA" = "#D55E00", "Combined" = "#444444")) +
theme_bw() +
theme(panel.grid=element_blank(), axis.text.y = element_text(size=5),plot.title = element_text(size=7)) +
coord_cartesian(expand=FALSE)+
facet_grid(Category ~ ., scales = "free")+theme(
strip.text.y = element_text(
size = 5))+
labs(title = "Diagnoses (Date 2020-04-05) | 12 sites | 15,637 patients ")
######################
## Barplot with ICD ##
######################
ggplot(data=selectionByCountry, aes(x=reorder(lbl,perc_patients), y=perc_patients)) +
geom_bar(aes(fill= siteid), stat="identity", position=position_dodge()) +
theme_bw()+
theme(axis.text.x = element_text(angle =45, hjust = 1), axis.text.y = element_text(size=5))+
labs(title = paste0("Number of patients by diagnostic code (>=",min(selection$num_patients),"patients)" ),
x = "diagnostic code", y = "percentage of patients")+ coord_flip()+
scale_fill_manual("legend", values = c("Italy" = "#009E73", "France" = "#0072B2", "Germany" = "#E69F00", "USA" = "#D55E00", "Combined" = "#444444"))
```
| github_jupyter |
**Decision Tree**
O Algorítimo cria uma árvore de decisão baseado nos dados de treinamento para classificar novos registros
Vantagens :
- Fácil interpretação / entendimento
- Os dados não precisam ser normalizados ou padronizados para treinar o modelo
- Bastante rápido para classificar novos registros
Desvantagens :
- Potencial para a criação de árvores muito complexas (lembrando que a construção da mesma problema np-completo)
- Tem problemas de variância (pouca sensibilidade a mudanças na base de treinamento o que acaba levando a overfiting). As vezes é preciso "podar" a base de treinamento
```
import pandas as pd
import numpy as np
from sklearn.preprocessing import LabelEncoder
from sklearn.naive_bayes import GaussianNB
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler,MinMaxScaler
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.metrics import confusion_matrix, accuracy_score, classification_report
from sklearn.compose import make_column_transformer
from sklearn.tree import DecisionTreeClassifier
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# carregando a base de dados de censo
base = pd.read_csv('../../res/census.csv')
# separando os dados de previsao e classificacao
previsores = base.iloc[:, 0:14].values
classificadores = base.iloc[:, 14].values
#gerando uma copia dos dados originais para fazer mais testes abaixo
previsores_escalonados=previsores.copy()
#efetuando correcoes nos dados do censo
#transformando dados categorios da base em dados discretos
labelencoder_prev = LabelEncoder()
previsores[:, 1] = labelencoder_prev.fit_transform(previsores[:, 1])
previsores[:, 3] = labelencoder_prev.fit_transform(previsores[:, 3])
previsores[:, 5] = labelencoder_prev.fit_transform(previsores[:, 5])
previsores[:, 6] = labelencoder_prev.fit_transform(previsores[:, 6])
previsores[:, 7] = labelencoder_prev.fit_transform(previsores[:, 7])
previsores[:, 8] = labelencoder_prev.fit_transform(previsores[:, 8])
previsores[:, 9] = labelencoder_prev.fit_transform(previsores[:, 9])
previsores[:, 13] = labelencoder_prev.fit_transform(previsores[:, 13])
#preprocess = make_column_transformer(( OneHotEncoder(categories='auto'), [1,3,5,6,7,8,9,13] ),remainder="passthrough")
#previsores = preprocess.fit_transform(previsores).toarray()
# Criando uma tabela de correlacao de pearson para entender a correlacao entre as variaveis
#gerando uma copia dos dados com a label encodada
alldata= previsores.copy()
alldata=np.append(alldata,classificadores.reshape(-1,1),axis=1)
alldata[:,14] = labelencoder_prev.fit_transform(alldata[:,14])
#print(alldata)
alldata=alldata.astype(float)
alldata =pd.DataFrame(alldata,columns=base.columns)
#gerando a matriz de correlacao a partir destes dados
corr = alldata.corr()
#print(alldata)
#print(corr)
ds=len(alldata.columns)
cmap = sns.diverging_palette(10, 255, as_cmap=True)
plt.figure(figsize=(20, 11))
plt.subplot(1, 2, 1)
plt.title("Pearson Correlation")
ax = sns.heatmap(corr, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=1, annot=True, cbar_kws={"shrink": .5})
ax.set_ylim(ds, 0)
plt.tight_layout()
plt.show()
#fazendo o one hot encoder para a copia da base (para os valores discretos)
#preprocess = make_column_transformer(( OneHotEncoder(categories='auto'), [1,3,5,6,7,8,9,13] ),remainder="passthrough")
#previsores_escalonados = preprocess.fit_transform(previsores_escalonados).toarray()
#removendo colunas com correlacao alta para verificar se a parformance melhora
previsores_less_cols=previsores.copy()
previsores_less_cols= np.delete(previsores,2,axis=1) #final-weight
previsores_less_cols= np.delete(previsores_less_cols,12,axis=1) #native-country
#previsores_less_cols= np.delete(previsores_less_cols,1,axis=1) #workclass
#separando os valores de teste e treinamento para os previsores escalonados e nao escalonados
previsores_treinamento, previsores_teste, classificadores_treinamento1, classificadores_teste1 = train_test_split(previsores, classificadores, test_size=0.15, random_state=0)
previsores_treinamentolc, previsores_testelc, classificadores_treinamentolc, classificadores_testelc = train_test_split(previsores_less_cols, classificadores, test_size=0.15, random_state=0)
# instanciando o naive bayes com o scikit
classificador = DecisionTreeClassifier(criterion='entropy',random_state=0)
classificador.fit(previsores_treinamento, classificadores_treinamento1)
previsoes = classificador.predict(previsores_teste)
#verificando a importancia de cada feature para o algoritimo
print(np.round(classificador.feature_importances_*100, 2))
print(np.sum(classificador.feature_importances_))
classificador.fit(previsores_treinamentolc, classificadores_treinamentolc)
# rodando previsoes com o dados de teste (copia)
#previsoes_dados_escalonados = classificador.predict(previsores_escalonados_teste)
# fazendo o fit com os dados normais
#classificador.fit(previsores_treinamento, classificadores_treinamento1)
previsoeslc = classificador.predict(previsores_testelc)
print(np.round(classificador.feature_importances_*100, 2))
print(np.sum(classificador.feature_importances_))
#testes dessa instancia algoritimo
# o dado de precisao per se nao quer dizer muita coisa e preciso verificar outras metricas
#precisao_escalonados = accuracy_score(classificadores_teste, previsoes_dados_escalonados)
precisao = accuracy_score(classificadores_teste1, previsoes)
precisaolc = accuracy_score(classificadores_testelc, previsoeslc)
# uma dessas metricas eh a matriz de confusao ... ela e capaz de mostrar o desempenho do algoritimo para cada classe
matrizlc = confusion_matrix(classificadores_testelc, previsoeslc)
matriz = confusion_matrix(classificadores_teste1, previsoes)
#o scikit tambem possui uma classe utilitaria que prove um report mais detalhado...
reportlc = classification_report(classificadores_testelc, previsoeslc)
report = classification_report(classificadores_teste1, previsoes)
print("Precisão :\n")
print(precisao)
print("\n")
print(precisaolc)
print("\nMatriz de confusão :\n")
print(matriz)
print(matrizlc)
print("\nReport :\n")
print (report)
print (reportlc)
```
<br>**TODO :Verificar a base de treinamento para melhorar a distribuicao das classes e verificar se ha alguma melhora**
| github_jupyter |
# Stickmodel Demo
stickmodel can be used to calculate internal forces and moments for statically determinate bar problems
```
import numpy as np
from matplotlib import pyplot as plt
from wingstructure.structure.stickmodel import solve_equilibrium
```
## define bar
```
# straight bar with two segments
nodes = np.array([[0,0,0], [0,1,0], [0,2,0]], dtype=np.float)
def display_bar(nodes, anotate=False):
plt.plot(nodes[:,1], nodes[:, 2], 'o-')
plt.axis('equal')
plt.axis('off')
if not anotate:
return
for i, node in enumerate(nodes):
plt.annotate(f'node-{i}', node[1:]-np.array([0.1, 0.1]))
display_bar(nodes, True)
plt.xlim(-0.5, 2.5);
```
## define loads
```
forces = np.array([
[0, 0.5, 0, 0, 1, 1, 0],
[0, 1.5, 0, 0, 0, 2, 1]
])
def display_force(ax, start, vec, color='k', arrow_size=1.0):
if (vec==0.0).all():
return
else:
ax.arrow(*start, *vec, fc=color, ec=color, head_width=arrow_size*0.07,
head_length=arrow_size*0.1, linewidth=2.5*arrow_size)
def display_forces(forces, fac=0.3, arrow_size=1.0):
ax = plt.gca()
for force in forces:
display_force(ax, force[1:3], (fac*force[4:6]), arrow_size=arrow_size)
display_bar(nodes)
display_forces(forces, fac=0.1)
plt.xlim(-0.5, 2.5);
```
## calculate equilibrium
*solve_equilibrium* calculates state of equilibrium
```
sol = solve_equilibrium(nodes, forces, prescribed={2:np.zeros(6)})
def display_sol(nodes, sol, fac=0.3):
ax = plt.gca()
for i in range(nodes.shape[0]-1):
display_force(ax, nodes[i, 1:], fac*np.array([sol[i, 1], 0.0]), color=f'C0{i+1}')
display_force(ax, nodes[i, 1:], fac*np.array([0.0, sol[i, 2]]), color=f'C0{i+1}')
display_force(ax, nodes[i+1, 1:], -fac*np.array([sol[i+1, 1], 0.0]), color=f'C0{i+1}')
display_force(ax, nodes[i+1, 1:], -fac*np.array([0.0, sol[i+1, 2]]), color=f'C0{i+1}')
display_bar(nodes)
display_forces(forces, fac=0.1)
display_sol(nodes, sol, fac=0.1)
plt.xlim(-0.5, 2.5);
```
# more complex model
```
n = 10
l = 3.0
nodes = np.vstack((np.zeros(n), np.linspace(0, l, n), np.zeros(n))).T
forces = np.zeros((n-1, 7))
# point of attack for forces
el_mid_pts = nodes[:-1, :] + np.diff(nodes, axis=0)/2
forces[:, 0:3] = el_mid_pts
# magnitudes of forces, only z value
forces[:, 5] = np.cos(el_mid_pts[:,1]*np.pi/2/l)
# element force is acting on
forces[:,-1] = range(n-1)
display_bar(nodes)
display_forces(forces, fac=0.3)
sol1 = np.cumsum(forces[::-1, 5])
sol2 = solve_equilibrium(nodes, forces, prescribed={n-1:np.zeros(6)})
plt.plot(nodes[:-1,1][::-1], sol1)
plt.plot(nodes[:, 1], sol2[:, 2], '--')
```
| github_jupyter |
**Chapter 4 – Training Linear Models**
_This notebook contains all the sample code and solutions to the exercises in chapter 4._
<table align="left">
<td>
<a href="https://colab.research.google.com/github/ageron/handson-ml2/blob/master/04_training_linear_models.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
</td>
<td>
<a target="_blank" href="https://kaggle.com/kernels/welcome?src=https://github.com/ageron/handson-ml2/blob/add-kaggle-badge/04_training_linear_models.ipynb"><img src="https://kaggle.com/static/images/open-in-kaggle.svg" /></a>
</td>
</table>
# Setup
First, let's import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures. We also check that Python 3.5 or later is installed (although Python 2.x may work, it is deprecated so we strongly recommend you use Python 3 instead), as well as Scikit-Learn ≥0.20.
```
# Python ≥3.5 is required
import sys
assert sys.version_info >= (3, 5)
# Scikit-Learn ≥0.20 is required
import sklearn
assert sklearn.__version__ >= "0.20"
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "training_linear_models"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
# Linear regression using the Normal Equation
```
import numpy as np
X = 2 * np.random.rand(100, 1)
y = 4 + 3 * X + np.random.randn(100, 1)
plt.plot(X, y, "b.")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.axis([0, 2, 0, 15])
save_fig("generated_data_plot")
plt.show()
X_b = np.c_[np.ones((100, 1)), X] # add x0 = 1 to each instance
theta_best = np.linalg.inv(X_b.T.dot(X_b)).dot(X_b.T).dot(y)
theta_best
X_new = np.array([[0], [2]])
X_new_b = np.c_[np.ones((2, 1)), X_new] # add x0 = 1 to each instance
y_predict = X_new_b.dot(theta_best)
y_predict
plt.plot(X_new, y_predict, "r-")
plt.plot(X, y, "b.")
plt.axis([0, 2, 0, 15])
plt.show()
```
The figure in the book actually corresponds to the following code, with a legend and axis labels:
```
plt.plot(X_new, y_predict, "r-", linewidth=2, label="Predictions")
plt.plot(X, y, "b.")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.legend(loc="upper left", fontsize=14)
plt.axis([0, 2, 0, 15])
save_fig("linear_model_predictions_plot")
plt.show()
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X, y)
lin_reg.intercept_, lin_reg.coef_
lin_reg.predict(X_new)
```
The `LinearRegression` class is based on the `scipy.linalg.lstsq()` function (the name stands for "least squares"), which you could call directly:
```
theta_best_svd, residuals, rank, s = np.linalg.lstsq(X_b, y, rcond=1e-6)
theta_best_svd
```
This function computes $\mathbf{X}^+\mathbf{y}$, where $\mathbf{X}^{+}$ is the _pseudoinverse_ of $\mathbf{X}$ (specifically the Moore-Penrose inverse). You can use `np.linalg.pinv()` to compute the pseudoinverse directly:
```
np.linalg.pinv(X_b).dot(y)
```
# Linear regression using batch gradient descent
```
eta = 0.1 # learning rate
n_iterations = 1000
m = 100
theta = np.random.randn(2,1) # random initialization
for iteration in range(n_iterations):
gradients = 2/m * X_b.T.dot(X_b.dot(theta) - y)
theta = theta - eta * gradients
theta
X_new_b.dot(theta)
theta_path_bgd = []
def plot_gradient_descent(theta, eta, theta_path=None):
m = len(X_b)
plt.plot(X, y, "b.")
n_iterations = 1000
for iteration in range(n_iterations):
if iteration < 10:
y_predict = X_new_b.dot(theta)
style = "b-" if iteration > 0 else "r--"
plt.plot(X_new, y_predict, style)
gradients = 2/m * X_b.T.dot(X_b.dot(theta) - y)
theta = theta - eta * gradients
if theta_path is not None:
theta_path.append(theta)
plt.xlabel("$x_1$", fontsize=18)
plt.axis([0, 2, 0, 15])
plt.title(r"$\eta = {}$".format(eta), fontsize=16)
np.random.seed(42)
theta = np.random.randn(2,1) # random initialization
plt.figure(figsize=(10,4))
plt.subplot(131); plot_gradient_descent(theta, eta=0.02)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.subplot(132); plot_gradient_descent(theta, eta=0.1, theta_path=theta_path_bgd)
plt.subplot(133); plot_gradient_descent(theta, eta=0.5)
save_fig("gradient_descent_plot")
plt.show()
```
# Stochastic Gradient Descent
```
theta_path_sgd = []
m = len(X_b)
np.random.seed(42)
n_epochs = 50
t0, t1 = 5, 50 # learning schedule hyperparameters
def learning_schedule(t):
return t0 / (t + t1)
theta = np.random.randn(2,1) # random initialization
for epoch in range(n_epochs):
for i in range(m):
if epoch == 0 and i < 20: # not shown in the book
y_predict = X_new_b.dot(theta) # not shown
style = "b-" if i > 0 else "r--" # not shown
plt.plot(X_new, y_predict, style) # not shown
random_index = np.random.randint(m)
xi = X_b[random_index:random_index+1]
yi = y[random_index:random_index+1]
gradients = 2 * xi.T.dot(xi.dot(theta) - yi)
eta = learning_schedule(epoch * m + i)
theta = theta - eta * gradients
theta_path_sgd.append(theta) # not shown
plt.plot(X, y, "b.") # not shown
plt.xlabel("$x_1$", fontsize=18) # not shown
plt.ylabel("$y$", rotation=0, fontsize=18) # not shown
plt.axis([0, 2, 0, 15]) # not shown
save_fig("sgd_plot") # not shown
plt.show() # not shown
theta
from sklearn.linear_model import SGDRegressor
sgd_reg = SGDRegressor(max_iter=1000, tol=1e-3, penalty=None, eta0=0.1, random_state=42)
sgd_reg.fit(X, y.ravel())
sgd_reg.intercept_, sgd_reg.coef_
```
# Mini-batch gradient descent
```
theta_path_mgd = []
n_iterations = 50
minibatch_size = 20
np.random.seed(42)
theta = np.random.randn(2,1) # random initialization
t0, t1 = 200, 1000
def learning_schedule(t):
return t0 / (t + t1)
t = 0
for epoch in range(n_iterations):
shuffled_indices = np.random.permutation(m)
X_b_shuffled = X_b[shuffled_indices]
y_shuffled = y[shuffled_indices]
for i in range(0, m, minibatch_size):
t += 1
xi = X_b_shuffled[i:i+minibatch_size]
yi = y_shuffled[i:i+minibatch_size]
gradients = 2/minibatch_size * xi.T.dot(xi.dot(theta) - yi)
eta = learning_schedule(t)
theta = theta - eta * gradients
theta_path_mgd.append(theta)
theta
theta_path_bgd = np.array(theta_path_bgd)
theta_path_sgd = np.array(theta_path_sgd)
theta_path_mgd = np.array(theta_path_mgd)
plt.figure(figsize=(7,4))
plt.plot(theta_path_sgd[:, 0], theta_path_sgd[:, 1], "r-s", linewidth=1, label="Stochastic")
plt.plot(theta_path_mgd[:, 0], theta_path_mgd[:, 1], "g-+", linewidth=2, label="Mini-batch")
plt.plot(theta_path_bgd[:, 0], theta_path_bgd[:, 1], "b-o", linewidth=3, label="Batch")
plt.legend(loc="upper left", fontsize=16)
plt.xlabel(r"$\theta_0$", fontsize=20)
plt.ylabel(r"$\theta_1$ ", fontsize=20, rotation=0)
plt.axis([2.5, 4.5, 2.3, 3.9])
save_fig("gradient_descent_paths_plot")
plt.show()
```
# Polynomial regression
```
import numpy as np
import numpy.random as rnd
np.random.seed(42)
m = 100
X = 6 * np.random.rand(m, 1) - 3
y = 0.5 * X**2 + X + 2 + np.random.randn(m, 1)
plt.plot(X, y, "b.")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.axis([-3, 3, 0, 10])
save_fig("quadratic_data_plot")
plt.show()
from sklearn.preprocessing import PolynomialFeatures
poly_features = PolynomialFeatures(degree=2, include_bias=False)
X_poly = poly_features.fit_transform(X)
X[0]
X_poly[0]
lin_reg = LinearRegression()
lin_reg.fit(X_poly, y)
lin_reg.intercept_, lin_reg.coef_
X_new=np.linspace(-3, 3, 100).reshape(100, 1)
X_new_poly = poly_features.transform(X_new)
y_new = lin_reg.predict(X_new_poly)
plt.plot(X, y, "b.")
plt.plot(X_new, y_new, "r-", linewidth=2, label="Predictions")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.legend(loc="upper left", fontsize=14)
plt.axis([-3, 3, 0, 10])
save_fig("quadratic_predictions_plot")
plt.show()
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import Pipeline
for style, width, degree in (("g-", 1, 300), ("b--", 2, 2), ("r-+", 2, 1)):
polybig_features = PolynomialFeatures(degree=degree, include_bias=False)
std_scaler = StandardScaler()
lin_reg = LinearRegression()
polynomial_regression = Pipeline([
("poly_features", polybig_features),
("std_scaler", std_scaler),
("lin_reg", lin_reg),
])
polynomial_regression.fit(X, y)
y_newbig = polynomial_regression.predict(X_new)
plt.plot(X_new, y_newbig, style, label=str(degree), linewidth=width)
plt.plot(X, y, "b.", linewidth=3)
plt.legend(loc="upper left")
plt.xlabel("$x_1$", fontsize=18)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.axis([-3, 3, 0, 10])
save_fig("high_degree_polynomials_plot")
plt.show()
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
def plot_learning_curves(model, X, y):
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=10)
train_errors, val_errors = [], []
for m in range(1, len(X_train)):
model.fit(X_train[:m], y_train[:m])
y_train_predict = model.predict(X_train[:m])
y_val_predict = model.predict(X_val)
train_errors.append(mean_squared_error(y_train[:m], y_train_predict))
val_errors.append(mean_squared_error(y_val, y_val_predict))
plt.plot(np.sqrt(train_errors), "r-+", linewidth=2, label="train")
plt.plot(np.sqrt(val_errors), "b-", linewidth=3, label="val")
plt.legend(loc="upper right", fontsize=14) # not shown in the book
plt.xlabel("Training set size", fontsize=14) # not shown
plt.ylabel("RMSE", fontsize=14) # not shown
lin_reg = LinearRegression()
plot_learning_curves(lin_reg, X, y)
plt.axis([0, 80, 0, 3]) # not shown in the book
save_fig("underfitting_learning_curves_plot") # not shown
plt.show() # not shown
from sklearn.pipeline import Pipeline
polynomial_regression = Pipeline([
("poly_features", PolynomialFeatures(degree=10, include_bias=False)),
("lin_reg", LinearRegression()),
])
plot_learning_curves(polynomial_regression, X, y)
plt.axis([0, 80, 0, 3]) # not shown
save_fig("learning_curves_plot") # not shown
plt.show() # not shown
```
# Regularized models
```
np.random.seed(42)
m = 20
X = 3 * np.random.rand(m, 1)
y = 1 + 0.5 * X + np.random.randn(m, 1) / 1.5
X_new = np.linspace(0, 3, 100).reshape(100, 1)
from sklearn.linear_model import Ridge
ridge_reg = Ridge(alpha=1, solver="cholesky", random_state=42)
ridge_reg.fit(X, y)
ridge_reg.predict([[1.5]])
ridge_reg = Ridge(alpha=1, solver="sag", random_state=42)
ridge_reg.fit(X, y)
ridge_reg.predict([[1.5]])
from sklearn.linear_model import Ridge
def plot_model(model_class, polynomial, alphas, **model_kargs):
for alpha, style in zip(alphas, ("b-", "g--", "r:")):
model = model_class(alpha, **model_kargs) if alpha > 0 else LinearRegression()
if polynomial:
model = Pipeline([
("poly_features", PolynomialFeatures(degree=10, include_bias=False)),
("std_scaler", StandardScaler()),
("regul_reg", model),
])
model.fit(X, y)
y_new_regul = model.predict(X_new)
lw = 2 if alpha > 0 else 1
plt.plot(X_new, y_new_regul, style, linewidth=lw, label=r"$\alpha = {}$".format(alpha))
plt.plot(X, y, "b.", linewidth=3)
plt.legend(loc="upper left", fontsize=15)
plt.xlabel("$x_1$", fontsize=18)
plt.axis([0, 3, 0, 4])
plt.figure(figsize=(8,4))
plt.subplot(121)
plot_model(Ridge, polynomial=False, alphas=(0, 10, 100), random_state=42)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.subplot(122)
plot_model(Ridge, polynomial=True, alphas=(0, 10**-5, 1), random_state=42)
save_fig("ridge_regression_plot")
plt.show()
```
**Note**: to be future-proof, we set `max_iter=1000` and `tol=1e-3` because these will be the default values in Scikit-Learn 0.21.
```
sgd_reg = SGDRegressor(penalty="l2", max_iter=1000, tol=1e-3, random_state=42)
sgd_reg.fit(X, y.ravel())
sgd_reg.predict([[1.5]])
from sklearn.linear_model import Lasso
plt.figure(figsize=(8,4))
plt.subplot(121)
plot_model(Lasso, polynomial=False, alphas=(0, 0.1, 1), random_state=42)
plt.ylabel("$y$", rotation=0, fontsize=18)
plt.subplot(122)
plot_model(Lasso, polynomial=True, alphas=(0, 10**-7, 1), random_state=42)
save_fig("lasso_regression_plot")
plt.show()
from sklearn.linear_model import Lasso
lasso_reg = Lasso(alpha=0.1)
lasso_reg.fit(X, y)
lasso_reg.predict([[1.5]])
from sklearn.linear_model import ElasticNet
elastic_net = ElasticNet(alpha=0.1, l1_ratio=0.5, random_state=42)
elastic_net.fit(X, y)
elastic_net.predict([[1.5]])
np.random.seed(42)
m = 100
X = 6 * np.random.rand(m, 1) - 3
y = 2 + X + 0.5 * X**2 + np.random.randn(m, 1)
X_train, X_val, y_train, y_val = train_test_split(X[:50], y[:50].ravel(), test_size=0.5, random_state=10)
```
Early stopping example:
```
from copy import deepcopy
poly_scaler = Pipeline([
("poly_features", PolynomialFeatures(degree=90, include_bias=False)),
("std_scaler", StandardScaler())
])
X_train_poly_scaled = poly_scaler.fit_transform(X_train)
X_val_poly_scaled = poly_scaler.transform(X_val)
sgd_reg = SGDRegressor(max_iter=1, tol=-np.infty, warm_start=True,
penalty=None, learning_rate="constant", eta0=0.0005, random_state=42)
minimum_val_error = float("inf")
best_epoch = None
best_model = None
for epoch in range(1000):
sgd_reg.fit(X_train_poly_scaled, y_train) # continues where it left off
y_val_predict = sgd_reg.predict(X_val_poly_scaled)
val_error = mean_squared_error(y_val, y_val_predict)
if val_error < minimum_val_error:
minimum_val_error = val_error
best_epoch = epoch
best_model = deepcopy(sgd_reg)
```
Create the graph:
```
sgd_reg = SGDRegressor(max_iter=1, tol=-np.infty, warm_start=True,
penalty=None, learning_rate="constant", eta0=0.0005, random_state=42)
n_epochs = 500
train_errors, val_errors = [], []
for epoch in range(n_epochs):
sgd_reg.fit(X_train_poly_scaled, y_train)
y_train_predict = sgd_reg.predict(X_train_poly_scaled)
y_val_predict = sgd_reg.predict(X_val_poly_scaled)
train_errors.append(mean_squared_error(y_train, y_train_predict))
val_errors.append(mean_squared_error(y_val, y_val_predict))
best_epoch = np.argmin(val_errors)
best_val_rmse = np.sqrt(val_errors[best_epoch])
plt.annotate('Best model',
xy=(best_epoch, best_val_rmse),
xytext=(best_epoch, best_val_rmse + 1),
ha="center",
arrowprops=dict(facecolor='black', shrink=0.05),
fontsize=16,
)
best_val_rmse -= 0.03 # just to make the graph look better
plt.plot([0, n_epochs], [best_val_rmse, best_val_rmse], "k:", linewidth=2)
plt.plot(np.sqrt(val_errors), "b-", linewidth=3, label="Validation set")
plt.plot(np.sqrt(train_errors), "r--", linewidth=2, label="Training set")
plt.legend(loc="upper right", fontsize=14)
plt.xlabel("Epoch", fontsize=14)
plt.ylabel("RMSE", fontsize=14)
save_fig("early_stopping_plot")
plt.show()
best_epoch, best_model
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
t1a, t1b, t2a, t2b = -1, 3, -1.5, 1.5
t1s = np.linspace(t1a, t1b, 500)
t2s = np.linspace(t2a, t2b, 500)
t1, t2 = np.meshgrid(t1s, t2s)
T = np.c_[t1.ravel(), t2.ravel()]
Xr = np.array([[1, 1], [1, -1], [1, 0.5]])
yr = 2 * Xr[:, :1] + 0.5 * Xr[:, 1:]
J = (1/len(Xr) * np.sum((T.dot(Xr.T) - yr.T)**2, axis=1)).reshape(t1.shape)
N1 = np.linalg.norm(T, ord=1, axis=1).reshape(t1.shape)
N2 = np.linalg.norm(T, ord=2, axis=1).reshape(t1.shape)
t_min_idx = np.unravel_index(np.argmin(J), J.shape)
t1_min, t2_min = t1[t_min_idx], t2[t_min_idx]
t_init = np.array([[0.25], [-1]])
def bgd_path(theta, X, y, l1, l2, core = 1, eta = 0.05, n_iterations = 200):
path = [theta]
for iteration in range(n_iterations):
gradients = core * 2/len(X) * X.T.dot(X.dot(theta) - y) + l1 * np.sign(theta) + l2 * theta
theta = theta - eta * gradients
path.append(theta)
return np.array(path)
fig, axes = plt.subplots(2, 2, sharex=True, sharey=True, figsize=(10.1, 8))
for i, N, l1, l2, title in ((0, N1, 2., 0, "Lasso"), (1, N2, 0, 2., "Ridge")):
JR = J + l1 * N1 + l2 * 0.5 * N2**2
tr_min_idx = np.unravel_index(np.argmin(JR), JR.shape)
t1r_min, t2r_min = t1[tr_min_idx], t2[tr_min_idx]
levelsJ=(np.exp(np.linspace(0, 1, 20)) - 1) * (np.max(J) - np.min(J)) + np.min(J)
levelsJR=(np.exp(np.linspace(0, 1, 20)) - 1) * (np.max(JR) - np.min(JR)) + np.min(JR)
levelsN=np.linspace(0, np.max(N), 10)
path_J = bgd_path(t_init, Xr, yr, l1=0, l2=0)
path_JR = bgd_path(t_init, Xr, yr, l1, l2)
path_N = bgd_path(np.array([[2.0], [0.5]]), Xr, yr, np.sign(l1)/3, np.sign(l2), core=0)
ax = axes[i, 0]
ax.grid(True)
ax.axhline(y=0, color='k')
ax.axvline(x=0, color='k')
ax.contourf(t1, t2, N / 2., levels=levelsN)
ax.plot(path_N[:, 0], path_N[:, 1], "y--")
ax.plot(0, 0, "ys")
ax.plot(t1_min, t2_min, "ys")
ax.set_title(r"$\ell_{}$ penalty".format(i + 1), fontsize=16)
ax.axis([t1a, t1b, t2a, t2b])
if i == 1:
ax.set_xlabel(r"$\theta_1$", fontsize=16)
ax.set_ylabel(r"$\theta_2$", fontsize=16, rotation=0)
ax = axes[i, 1]
ax.grid(True)
ax.axhline(y=0, color='k')
ax.axvline(x=0, color='k')
ax.contourf(t1, t2, JR, levels=levelsJR, alpha=0.9)
ax.plot(path_JR[:, 0], path_JR[:, 1], "w-o")
ax.plot(path_N[:, 0], path_N[:, 1], "y--")
ax.plot(0, 0, "ys")
ax.plot(t1_min, t2_min, "ys")
ax.plot(t1r_min, t2r_min, "rs")
ax.set_title(title, fontsize=16)
ax.axis([t1a, t1b, t2a, t2b])
if i == 1:
ax.set_xlabel(r"$\theta_1$", fontsize=16)
save_fig("lasso_vs_ridge_plot")
plt.show()
```
# Logistic regression
```
t = np.linspace(-10, 10, 100)
sig = 1 / (1 + np.exp(-t))
plt.figure(figsize=(9, 3))
plt.plot([-10, 10], [0, 0], "k-")
plt.plot([-10, 10], [0.5, 0.5], "k:")
plt.plot([-10, 10], [1, 1], "k:")
plt.plot([0, 0], [-1.1, 1.1], "k-")
plt.plot(t, sig, "b-", linewidth=2, label=r"$\sigma(t) = \frac{1}{1 + e^{-t}}$")
plt.xlabel("t")
plt.legend(loc="upper left", fontsize=20)
plt.axis([-10, 10, -0.1, 1.1])
save_fig("logistic_function_plot")
plt.show()
from sklearn import datasets
iris = datasets.load_iris()
list(iris.keys())
print(iris.DESCR)
X = iris["data"][:, 3:] # petal width
y = (iris["target"] == 2).astype(np.int) # 1 if Iris virginica, else 0
```
**Note**: To be future-proof we set `solver="lbfgs"` since this will be the default value in Scikit-Learn 0.22.
```
from sklearn.linear_model import LogisticRegression
log_reg = LogisticRegression(solver="lbfgs", random_state=42)
log_reg.fit(X, y)
X_new = np.linspace(0, 3, 1000).reshape(-1, 1)
y_proba = log_reg.predict_proba(X_new)
plt.plot(X_new, y_proba[:, 1], "g-", linewidth=2, label="Iris virginica")
plt.plot(X_new, y_proba[:, 0], "b--", linewidth=2, label="Not Iris virginica")
```
The figure in the book actually is actually a bit fancier:
```
X_new = np.linspace(0, 3, 1000).reshape(-1, 1)
y_proba = log_reg.predict_proba(X_new)
decision_boundary = X_new[y_proba[:, 1] >= 0.5][0]
plt.figure(figsize=(8, 3))
plt.plot(X[y==0], y[y==0], "bs")
plt.plot(X[y==1], y[y==1], "g^")
plt.plot([decision_boundary, decision_boundary], [-1, 2], "k:", linewidth=2)
plt.plot(X_new, y_proba[:, 1], "g-", linewidth=2, label="Iris virginica")
plt.plot(X_new, y_proba[:, 0], "b--", linewidth=2, label="Not Iris virginica")
plt.text(decision_boundary+0.02, 0.15, "Decision boundary", fontsize=14, color="k", ha="center")
plt.arrow(decision_boundary, 0.08, -0.3, 0, head_width=0.05, head_length=0.1, fc='b', ec='b')
plt.arrow(decision_boundary, 0.92, 0.3, 0, head_width=0.05, head_length=0.1, fc='g', ec='g')
plt.xlabel("Petal width (cm)", fontsize=14)
plt.ylabel("Probability", fontsize=14)
plt.legend(loc="center left", fontsize=14)
plt.axis([0, 3, -0.02, 1.02])
save_fig("logistic_regression_plot")
plt.show()
decision_boundary
log_reg.predict([[1.7], [1.5]])
from sklearn.linear_model import LogisticRegression
X = iris["data"][:, (2, 3)] # petal length, petal width
y = (iris["target"] == 2).astype(np.int)
log_reg = LogisticRegression(solver="lbfgs", C=10**10, random_state=42)
log_reg.fit(X, y)
x0, x1 = np.meshgrid(
np.linspace(2.9, 7, 500).reshape(-1, 1),
np.linspace(0.8, 2.7, 200).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_proba = log_reg.predict_proba(X_new)
plt.figure(figsize=(10, 4))
plt.plot(X[y==0, 0], X[y==0, 1], "bs")
plt.plot(X[y==1, 0], X[y==1, 1], "g^")
zz = y_proba[:, 1].reshape(x0.shape)
contour = plt.contour(x0, x1, zz, cmap=plt.cm.brg)
left_right = np.array([2.9, 7])
boundary = -(log_reg.coef_[0][0] * left_right + log_reg.intercept_[0]) / log_reg.coef_[0][1]
plt.clabel(contour, inline=1, fontsize=12)
plt.plot(left_right, boundary, "k--", linewidth=3)
plt.text(3.5, 1.5, "Not Iris virginica", fontsize=14, color="b", ha="center")
plt.text(6.5, 2.3, "Iris virginica", fontsize=14, color="g", ha="center")
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.axis([2.9, 7, 0.8, 2.7])
save_fig("logistic_regression_contour_plot")
plt.show()
X = iris["data"][:, (2, 3)] # petal length, petal width
y = iris["target"]
softmax_reg = LogisticRegression(multi_class="multinomial",solver="lbfgs", C=10, random_state=42)
softmax_reg.fit(X, y)
x0, x1 = np.meshgrid(
np.linspace(0, 8, 500).reshape(-1, 1),
np.linspace(0, 3.5, 200).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_proba = softmax_reg.predict_proba(X_new)
y_predict = softmax_reg.predict(X_new)
zz1 = y_proba[:, 1].reshape(x0.shape)
zz = y_predict.reshape(x0.shape)
plt.figure(figsize=(10, 4))
plt.plot(X[y==2, 0], X[y==2, 1], "g^", label="Iris virginica")
plt.plot(X[y==1, 0], X[y==1, 1], "bs", label="Iris versicolor")
plt.plot(X[y==0, 0], X[y==0, 1], "yo", label="Iris setosa")
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x0, x1, zz, cmap=custom_cmap)
contour = plt.contour(x0, x1, zz1, cmap=plt.cm.brg)
plt.clabel(contour, inline=1, fontsize=12)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="center left", fontsize=14)
plt.axis([0, 7, 0, 3.5])
save_fig("softmax_regression_contour_plot")
plt.show()
softmax_reg.predict([[5, 2]])
softmax_reg.predict_proba([[5, 2]])
```
# Exercise solutions
## 1. to 11.
See appendix A.
## 12. Batch Gradient Descent with early stopping for Softmax Regression
(without using Scikit-Learn)
Let's start by loading the data. We will just reuse the Iris dataset we loaded earlier.
```
X = iris["data"][:, (2, 3)] # petal length, petal width
y = iris["target"]
```
We need to add the bias term for every instance ($x_0 = 1$):
```
X_with_bias = np.c_[np.ones([len(X), 1]), X]
```
And let's set the random seed so the output of this exercise solution is reproducible:
```
np.random.seed(2042)
```
The easiest option to split the dataset into a training set, a validation set and a test set would be to use Scikit-Learn's `train_test_split()` function, but the point of this exercise is to try understand the algorithms by implementing them manually. So here is one possible implementation:
```
test_ratio = 0.2
validation_ratio = 0.2
total_size = len(X_with_bias)
test_size = int(total_size * test_ratio)
validation_size = int(total_size * validation_ratio)
train_size = total_size - test_size - validation_size
rnd_indices = np.random.permutation(total_size)
X_train = X_with_bias[rnd_indices[:train_size]]
y_train = y[rnd_indices[:train_size]]
X_valid = X_with_bias[rnd_indices[train_size:-test_size]]
y_valid = y[rnd_indices[train_size:-test_size]]
X_test = X_with_bias[rnd_indices[-test_size:]]
y_test = y[rnd_indices[-test_size:]]
```
The targets are currently class indices (0, 1 or 2), but we need target class probabilities to train the Softmax Regression model. Each instance will have target class probabilities equal to 0.0 for all classes except for the target class which will have a probability of 1.0 (in other words, the vector of class probabilities for ay given instance is a one-hot vector). Let's write a small function to convert the vector of class indices into a matrix containing a one-hot vector for each instance:
```
def to_one_hot(y):
n_classes = y.max() + 1
m = len(y)
Y_one_hot = np.zeros((m, n_classes))
Y_one_hot[np.arange(m), y] = 1
return Y_one_hot
```
Let's test this function on the first 10 instances:
```
y_train[:10]
to_one_hot(y_train[:10])
```
Looks good, so let's create the target class probabilities matrix for the training set and the test set:
```
Y_train_one_hot = to_one_hot(y_train)
Y_valid_one_hot = to_one_hot(y_valid)
Y_test_one_hot = to_one_hot(y_test)
```
Now let's implement the Softmax function. Recall that it is defined by the following equation:
$\sigma\left(\mathbf{s}(\mathbf{x})\right)_k = \dfrac{\exp\left(s_k(\mathbf{x})\right)}{\sum\limits_{j=1}^{K}{\exp\left(s_j(\mathbf{x})\right)}}$
```
def softmax(logits):
exps = np.exp(logits)
exp_sums = np.sum(exps, axis=1, keepdims=True)
return exps / exp_sums
```
We are almost ready to start training. Let's define the number of inputs and outputs:
```
n_inputs = X_train.shape[1] # == 3 (2 features plus the bias term)
n_outputs = len(np.unique(y_train)) # == 3 (3 iris classes)
```
Now here comes the hardest part: training! Theoretically, it's simple: it's just a matter of translating the math equations into Python code. But in practice, it can be quite tricky: in particular, it's easy to mix up the order of the terms, or the indices. You can even end up with code that looks like it's working but is actually not computing exactly the right thing. When unsure, you should write down the shape of each term in the equation and make sure the corresponding terms in your code match closely. It can also help to evaluate each term independently and print them out. The good news it that you won't have to do this everyday, since all this is well implemented by Scikit-Learn, but it will help you understand what's going on under the hood.
So the equations we will need are the cost function:
$J(\mathbf{\Theta}) =
- \dfrac{1}{m}\sum\limits_{i=1}^{m}\sum\limits_{k=1}^{K}{y_k^{(i)}\log\left(\hat{p}_k^{(i)}\right)}$
And the equation for the gradients:
$\nabla_{\mathbf{\theta}^{(k)}} \, J(\mathbf{\Theta}) = \dfrac{1}{m} \sum\limits_{i=1}^{m}{ \left ( \hat{p}^{(i)}_k - y_k^{(i)} \right ) \mathbf{x}^{(i)}}$
Note that $\log\left(\hat{p}_k^{(i)}\right)$ may not be computable if $\hat{p}_k^{(i)} = 0$. So we will add a tiny value $\epsilon$ to $\log\left(\hat{p}_k^{(i)}\right)$ to avoid getting `nan` values.
```
eta = 0.01
n_iterations = 5001
m = len(X_train)
epsilon = 1e-7
Theta = np.random.randn(n_inputs, n_outputs)
for iteration in range(n_iterations):
logits = X_train.dot(Theta)
Y_proba = softmax(logits)
if iteration % 500 == 0:
loss = -np.mean(np.sum(Y_train_one_hot * np.log(Y_proba + epsilon), axis=1))
print(iteration, loss)
error = Y_proba - Y_train_one_hot
gradients = 1/m * X_train.T.dot(error)
Theta = Theta - eta * gradients
```
And that's it! The Softmax model is trained. Let's look at the model parameters:
```
Theta
```
Let's make predictions for the validation set and check the accuracy score:
```
logits = X_valid.dot(Theta)
Y_proba = softmax(logits)
y_predict = np.argmax(Y_proba, axis=1)
accuracy_score = np.mean(y_predict == y_valid)
accuracy_score
```
Well, this model looks pretty good. For the sake of the exercise, let's add a bit of $\ell_2$ regularization. The following training code is similar to the one above, but the loss now has an additional $\ell_2$ penalty, and the gradients have the proper additional term (note that we don't regularize the first element of `Theta` since this corresponds to the bias term). Also, let's try increasing the learning rate `eta`.
```
eta = 0.1
n_iterations = 5001
m = len(X_train)
epsilon = 1e-7
alpha = 0.1 # regularization hyperparameter
Theta = np.random.randn(n_inputs, n_outputs)
for iteration in range(n_iterations):
logits = X_train.dot(Theta)
Y_proba = softmax(logits)
if iteration % 500 == 0:
xentropy_loss = -np.mean(np.sum(Y_train_one_hot * np.log(Y_proba + epsilon), axis=1))
l2_loss = 1/2 * np.sum(np.square(Theta[1:]))
loss = xentropy_loss + alpha * l2_loss
print(iteration, loss)
error = Y_proba - Y_train_one_hot
gradients = 1/m * X_train.T.dot(error) + np.r_[np.zeros([1, n_outputs]), alpha * Theta[1:]]
Theta = Theta - eta * gradients
```
Because of the additional $\ell_2$ penalty, the loss seems greater than earlier, but perhaps this model will perform better? Let's find out:
```
logits = X_valid.dot(Theta)
Y_proba = softmax(logits)
y_predict = np.argmax(Y_proba, axis=1)
accuracy_score = np.mean(y_predict == y_valid)
accuracy_score
```
Cool, perfect accuracy! We probably just got lucky with this validation set, but still, it's pleasant.
Now let's add early stopping. For this we just need to measure the loss on the validation set at every iteration and stop when the error starts growing.
```
eta = 0.1
n_iterations = 5001
m = len(X_train)
epsilon = 1e-7
alpha = 0.1 # regularization hyperparameter
best_loss = np.infty
Theta = np.random.randn(n_inputs, n_outputs)
for iteration in range(n_iterations):
logits = X_train.dot(Theta)
Y_proba = softmax(logits)
error = Y_proba - Y_train_one_hot
gradients = 1/m * X_train.T.dot(error) + np.r_[np.zeros([1, n_outputs]), alpha * Theta[1:]]
Theta = Theta - eta * gradients
logits = X_valid.dot(Theta)
Y_proba = softmax(logits)
xentropy_loss = -np.mean(np.sum(Y_valid_one_hot * np.log(Y_proba + epsilon), axis=1))
l2_loss = 1/2 * np.sum(np.square(Theta[1:]))
loss = xentropy_loss + alpha * l2_loss
if iteration % 500 == 0:
print(iteration, loss)
if loss < best_loss:
best_loss = loss
else:
print(iteration - 1, best_loss)
print(iteration, loss, "early stopping!")
break
logits = X_valid.dot(Theta)
Y_proba = softmax(logits)
y_predict = np.argmax(Y_proba, axis=1)
accuracy_score = np.mean(y_predict == y_valid)
accuracy_score
```
Still perfect, but faster.
Now let's plot the model's predictions on the whole dataset:
```
x0, x1 = np.meshgrid(
np.linspace(0, 8, 500).reshape(-1, 1),
np.linspace(0, 3.5, 200).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
X_new_with_bias = np.c_[np.ones([len(X_new), 1]), X_new]
logits = X_new_with_bias.dot(Theta)
Y_proba = softmax(logits)
y_predict = np.argmax(Y_proba, axis=1)
zz1 = Y_proba[:, 1].reshape(x0.shape)
zz = y_predict.reshape(x0.shape)
plt.figure(figsize=(10, 4))
plt.plot(X[y==2, 0], X[y==2, 1], "g^", label="Iris virginica")
plt.plot(X[y==1, 0], X[y==1, 1], "bs", label="Iris versicolor")
plt.plot(X[y==0, 0], X[y==0, 1], "yo", label="Iris setosa")
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x0, x1, zz, cmap=custom_cmap)
contour = plt.contour(x0, x1, zz1, cmap=plt.cm.brg)
plt.clabel(contour, inline=1, fontsize=12)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.legend(loc="upper left", fontsize=14)
plt.axis([0, 7, 0, 3.5])
plt.show()
```
And now let's measure the final model's accuracy on the test set:
```
logits = X_test.dot(Theta)
Y_proba = softmax(logits)
y_predict = np.argmax(Y_proba, axis=1)
accuracy_score = np.mean(y_predict == y_test)
accuracy_score
```
Our perfect model turns out to have slight imperfections. This variability is likely due to the very small size of the dataset: depending on how you sample the training set, validation set and the test set, you can get quite different results. Try changing the random seed and running the code again a few times, you will see that the results will vary.
| github_jupyter |
## Reinforcement Learning for seq2seq
This time we'll solve a problem of transribing hebrew words in english, also known as g2p (grapheme2phoneme)
* word (sequence of letters in source language) -> translation (sequence of letters in target language)
Unlike what most deep learning researchers do, we won't only train it to maximize likelihood of correct translation, but also employ reinforcement learning to actually teach it to translate with as few errors as possible.
### About the task
One notable property of Hebrew is that it's consonant language. That is, there are no wovels in the written language. One could represent wovels with diacritics above consonants, but you don't expect people to do that in everyay life.
Therefore, some hebrew characters will correspond to several english letters and others - to none, so we should use encoder-decoder architecture to figure that out.
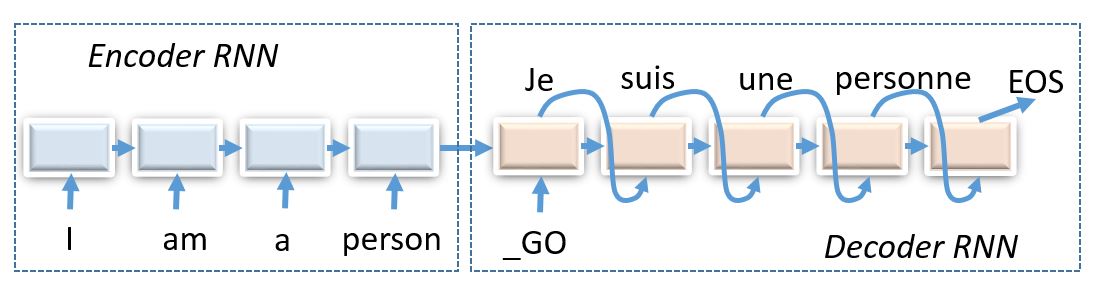
_(img: esciencegroup.files.wordpress.com)_
Encoder-decoder architectures are about converting anything to anything, including
* Machine translation and spoken dialogue systems
* [Image captioning](http://mscoco.org/dataset/#captions-challenge2015) and [image2latex](https://openai.com/requests-for-research/#im2latex) (convolutional encoder, recurrent decoder)
* Generating [images by captions](https://arxiv.org/abs/1511.02793) (recurrent encoder, convolutional decoder)
* Grapheme2phoneme - convert words to transcripts
We chose simplified __Hebrew->English__ machine translation for words and short phrases (character-level), as it is relatively quick to train even without a gpu cluster.
```
# If True, only translates phrases shorter than 20 characters (way easier).
EASY_MODE = True
# Please keep it until you're done debugging your code
# If false, works with all phrases (please switch to this mode for homework assignment)
# way we translate. Either "he-to-en" or "en-to-he"
MODE = "he-to-en"
# maximal length of _generated_ output, does not affect training
MAX_OUTPUT_LENGTH = 50 if not EASY_MODE else 20
REPORT_FREQ = 100 # how often to evaluate validation score
```
### Step 1: preprocessing
We shall store dataset as a dictionary
`{ word1:[translation1,translation2,...], word2:[...],...}`.
This is mostly due to the fact that many words have several correct translations.
We have implemented this thing for you so that you can focus on more interesting parts.
__Attention python2 users!__ You may want to cast everything to unicode later during homework phase, just make sure you do it _everywhere_.
```
import numpy as np
from collections import defaultdict
word_to_translation = defaultdict(list) # our dictionary
bos = '_'
eos = ';'
with open("main_dataset.txt", encoding='utf8') as fin:
for line in fin:
en, he = line[:-1].lower().replace(bos, ' ').replace(eos,
' ').split('\t')
word, trans = (he, en) if MODE == 'he-to-en' else (en, he)
if len(word) < 3:
continue
if EASY_MODE:
if max(len(word), len(trans)) > 20:
continue
word_to_translation[word].append(trans)
print("size = ", len(word_to_translation))
# get all unique lines in source language
all_words = np.array(list(word_to_translation.keys()))
# get all unique lines in translation language
all_translations = np.array(
[ts for all_ts in word_to_translation.values() for ts in all_ts])
```
### split the dataset
We hold out 10% of all words to be used for validation.
```
from sklearn.model_selection import train_test_split
train_words, test_words = train_test_split(
all_words, test_size=0.1, random_state=42)
```
### Building vocabularies
We now need to build vocabularies that map strings to token ids and vice versa. We're gonna need these fellas when we feed training data into model or convert output matrices into english words.
```
from voc import Vocab
inp_voc = Vocab.from_lines(''.join(all_words), bos=bos, eos=eos, sep='')
out_voc = Vocab.from_lines(''.join(all_translations), bos=bos, eos=eos, sep='')
# Here's how you cast lines into ids and backwards.
batch_lines = all_words[:5]
batch_ids = inp_voc.to_matrix(batch_lines)
batch_lines_restored = inp_voc.to_lines(batch_ids)
print("lines")
print(batch_lines)
print("\nwords to ids (0 = bos, 1 = eos):")
print(batch_ids)
print("\nback to words")
print(batch_lines_restored)
```
Draw word/translation length distributions to estimate the scope of the task.
```
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=[8, 4])
plt.subplot(1, 2, 1)
plt.title("words")
plt.hist(list(map(len, all_words)), bins=20)
plt.subplot(1, 2, 2)
plt.title('translations')
plt.hist(list(map(len, all_translations)), bins=20)
```
### Step 3: deploy encoder-decoder (1 point)
__assignment starts here__
Our architecture consists of two main blocks:
* Encoder reads words character by character and outputs code vector (usually a function of last RNN state)
* Decoder takes that code vector and produces translations character by character
Than it gets fed into a model that follows this simple interface:
* __`model.symbolic_translate(inp, **flags) -> out, logp`__ - takes symbolic int32 matrix of hebrew words, produces output tokens sampled from the model and output log-probabilities for all possible tokens at each tick.
* if given flag __`greedy=True`__, takes most likely next token at each iteration. Otherwise samples with next token probabilities predicted by model.
* __`model.symbolic_score(inp, out, **flags) -> logp`__ - takes symbolic int32 matrices of hebrew words and their english translations. Computes the log-probabilities of all possible english characters given english prefices and hebrew word.
That's all! It's as hard as it gets. With those two methods alone you can implement all kinds of prediction and training.
```
# set flags here if necessary
import theano
theano.config.floatX = 'float32'
import theano.tensor as T
import lasagne
from basic_model_theano import BasicTranslationModel
model = BasicTranslationModel(inp_voc, out_voc,
emb_size=64, hid_size=128)
# Play around with symbolic_translate and symbolic_score
inp = T.constant(np.random.randint(0, 10, [3, 5], dtype='int32'))
out = T.constant(np.random.randint(0, 10, [3, 5], dtype='int32'))
# translate inp (with untrained model)
sampled_out, logp = model.symbolic_translate(inp, greedy=False)
dummy_translate = theano.function([], sampled_out, updates=model.auto_updates)
print("\nSymbolic_translate output:\n", sampled_out, logp)
print("\nSample translations:\n", dummy_translate())
# score logp(out | inp) with untrained input
logp = model.symbolic_score(inp, out)
dummy_score = theano.function([], logp)
print("\nSymbolic_score output:\n", logp)
print("\nLog-probabilities (clipped):\n", dummy_score()[:, :2, :5])
# Prepare any operations you want here
inp = T.imatrix("input tokens [batch,time]")
trans, _ = <build symbolic translations with greedy = True >
translate_fun = theano.function([inp], trans, updates=model.auto_updates)
def translate(lines):
"""
You are given a list of input lines.
Make your neural network translate them.
:return: a list of output lines
"""
# Convert lines to a matrix of indices
lines_ix = <YOUR CODE >
# Compute translations in form of indices (call your function)
trans_ix = <YOUR CODE >
# Convert translations back into strings
return out_voc.to_lines(trans_ix)
print("Sample inputs:", all_words[:3])
print("Dummy translations:", translate(all_words[:3]))
assert trans.ndim == 2 and trans.dtype.startswith(
'int'), "trans must be a tensor of integers (token ids)"
assert translate(all_words[:3]) == translate(
all_words[:3]), "make sure translation is deterministic (use greedy=True and disable any noise layers)"
assert type(translate(all_words[:3])) is list and (type(translate(all_words[:1])[0]) is str or type(
translate(all_words[:1])[0]) is unicode), "translate(lines) must return a sequence of strings!"
print("Tests passed!")
```
### Scoring function
LogLikelihood is a poor estimator of model performance.
* If we predict zero probability once, it shouldn't ruin entire model.
* It is enough to learn just one translation if there are several correct ones.
* What matters is how many mistakes model's gonna make when it translates!
Therefore, we will use minimal Levenshtein distance. It measures how many characters do we need to add/remove/replace from model translation to make it perfect. Alternatively, one could use character-level BLEU/RougeL or other similar metrics.
The catch here is that Levenshtein distance is not differentiable: it isn't even continuous. We can't train our neural network to maximize it by gradient descent.
```
import editdistance # !pip install editdistance
def get_distance(word, trans):
"""
A function that takes word and predicted translation
and evaluates (Levenshtein's) edit distance to closest correct translation
"""
references = word_to_translation[word]
assert len(references) != 0, "wrong/unknown word"
return min(editdistance.eval(trans, ref) for ref in references)
def score(words, bsize=100):
"""a function that computes levenshtein distance for bsize random samples"""
assert isinstance(words, np.ndarray)
batch_words = np.random.choice(words, size=bsize, replace=False)
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
return np.array(distances, dtype='float32')
# should be around 5-50 and decrease rapidly after training :)
[score(test_words, 10).mean() for _ in range(5)]
```
## Step 2: Supervised pre-training
Here we define a function that trains our model through maximizing log-likelihood a.k.a. minimizing crossentropy.
```
from agentnet.learning.generic import get_values_for_actions, get_mask_by_eos
class llh_trainer:
# variable for correct answers
input_sequence = T.imatrix("input sequence [batch,time]")
reference_answers = T.imatrix("reference translations [batch, time]")
# Compute log-probabilities of all possible tokens at each step. Use model interface.
logprobs_seq = <YOUR CODE >
# compute mean crossentropy
crossentropy = - get_values_for_actions(logprobs_seq, reference_answers)
mask = get_mask_by_eos(T.eq(reference_answers, out_voc.eos_ix))
loss = T.sum(crossentropy * mask)/T.sum(mask)
# Build weight updates. Use model.weights to get all trainable params.
updates = <YOUR CODE >
train_step = theano.function(
[input_sequence, reference_answers], loss, updates=updates)
```
Actually run training on minibatches
```
import random
def sample_batch(words, word_to_translation, batch_size):
"""
sample random batch of words and random correct translation for each word
example usage:
batch_x,batch_y = sample_batch(train_words, word_to_translations,10)
"""
# choose words
batch_words = np.random.choice(words, size=batch_size)
# choose translations
batch_trans_candidates = list(map(word_to_translation.get, batch_words))
batch_trans = list(map(random.choice, batch_trans_candidates))
return inp_voc.to_matrix(batch_words), out_voc.to_matrix(batch_trans)
bx, by = sample_batch(train_words, word_to_translation, batch_size=3)
print("Source:")
print(bx)
print("Target:")
print(by)
from IPython.display import clear_output
from tqdm import tqdm, trange # or use tqdm_notebook,tnrange
loss_history = []
editdist_history = []
for i in trange(25000):
loss = llh_trainer.train_step(
*sample_batch(train_words, word_to_translation, 32))
loss_history.append(loss)
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
plt.figure(figsize=(12, 4))
plt.subplot(131)
plt.title('train loss / traning time')
plt.plot(loss_history)
plt.grid()
plt.subplot(132)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(133)
plt.title('val score / traning time')
plt.plot(editdist_history)
plt.grid()
plt.show()
print("llh=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
```
## Preparing for reinforcement learning (2 points)
First we need to define loss function as a custom theano operation.
The simple way to do so is
```
@theano.compile.as_op(input_types,output_type(s),infer_shape)
def my_super_function(inputs):
return outputs
```
__Your task__ is to implement `_compute_levenshtein` function that takes matrices of words and translations, along with input masks, then converts those to actual words and phonemes and computes min-levenshtein via __get_distance__ function above.
```
@theano.compile.as_op([T.imatrix]*2, [T.fvector], lambda _, shapes: [shapes[0][:1]])
def _compute_levenshtein(words_ix, trans_ix):
"""
A custom theano operation that computes levenshtein loss for predicted trans.
Params:
- words_ix - a matrix of letter indices, shape=[batch_size,word_length]
- words_mask - a matrix of zeros/ones,
1 means "word is still not finished"
0 means "word has already finished and this is padding"
- trans_mask - a matrix of output letter indices, shape=[batch_size,translation_length]
- trans_mask - a matrix of zeros/ones, similar to words_mask but for trans_ix
Please implement the function and make sure it passes tests from the next cell.
"""
# convert words to strings
words = <restore words(a list of strings) from words_ix >
assert type(words) is list and type(
words[0]) is str and len(words) == len(words_ix)
# convert translations to lists
translations = <restore trans(a list of lists of phonemes) from trans_ix
assert type(translations) is list and type(
translations[0]) is str and len(translations) == len(trans_ix)
# computes levenstein distances. can be arbitrary python code.
distances = <apply get_distance to each pair of[words, translations] >
assert type(distances) in (list, tuple, np.ndarray) and len(
distances) == len(words_ix)
distances = np.array(list(distances), dtype='float32')
return distances
# forbid gradient
from theano.gradient import disconnected_grad
def compute_levenshtein(*args):
return disconnected_grad(_compute_levenshtein(*[arg.astype('int32') for arg in args]))
```
Simple test suite to make sure your implementation is correct. Hint: if you run into any bugs, feel free to use print from inside _compute_levenshtein.
```
# test suite
# sample random batch of (words, correct trans, wrong trans)
batch_words = np.random.choice(train_words, size=100)
batch_trans = list(map(random.choice, map(
word_to_translation.get, batch_words)))
batch_trans_wrong = np.random.choice(all_translations, size=100)
batch_words_ix = T.constant(inp_voc.to_matrix(batch_words))
batch_trans_ix = T.constant(out_voc.to_matrix(batch_trans))
batch_trans_wrong_ix = T.constant(out_voc.to_matrix(batch_trans_wrong))
# assert compute_levenshtein is zero for ideal translations
correct_answers_score = compute_levenshtein(
batch_words_ix, batch_trans_ix).eval()
assert np.all(correct_answers_score ==
0), "a perfect translation got nonzero levenshtein score!"
print("Everything seems alright!")
# assert compute_levenshtein matches actual scoring function
wrong_answers_score = compute_levenshtein(
batch_words_ix, batch_trans_wrong_ix).eval()
true_wrong_answers_score = np.array(
list(map(get_distance, batch_words, batch_trans_wrong)))
assert np.all(wrong_answers_score ==
true_wrong_answers_score), "for some word symbolic levenshtein is different from actual levenshtein distance"
print("Everything seems alright!")
```
Once you got it working...
* You may now want to __remove/comment asserts__ from function code for a slight speed-up.
* There's a more detailed tutorial on custom theano ops here: [docs](http://deeplearning.net/software/theano/extending/extending_theano.html), [example](https://gist.github.com/justheuristic/9f4ffef6162a8089c3260fc3bbacbf46).
## Self-critical policy gradient (2 points)
In this section you'll implement algorithm called self-critical sequence training (here's an [article](https://arxiv.org/abs/1612.00563)).
The algorithm is a vanilla policy gradient with a special baseline.
$$ \nabla J = E_{x \sim p(s)} E_{y \sim \pi(y|x)} \nabla log \pi(y|x) \cdot (R(x,y) - b(x)) $$
Here reward R(x,y) is a __negative levenshtein distance__ (since we minimize it). The baseline __b(x)__ represents how well model fares on word __x__.
In practice, this means that we compute baseline as a score of greedy translation, $b(x) = R(x,y_{greedy}(x)) $.

Luckily, we already obtained the required outputs: `model.greedy_translations, model.greedy_mask` and we only need to compute levenshtein using `compute_levenshtein` function.
```
class trainer:
input_sequence = T.imatrix("input tokens [batch,time]")
# use model to __sample__ symbolic translations given input_sequence
sample_translations, sample_logp = <your code here >
auto_updates = model.auto_updates
# use model to __greedy__ symbolic translations given input_sequence
greedy_translations, greedy_logp = <your code here >
greedy_auto_updates = model.auto_updates
# Note: you can use model.symbolic_translate(...,unroll_scan=True,max_len=MAX_OUTPUT_LENGTH)
# to run much faster at a cost of longer compilation
rewards = - compute_levenshtein(input_sequence, sample_translations)
baseline = <compute __negative__ levenshtein for greedy mode >
# compute advantage using rewards and baseline
advantage = <your code - compute advantage >
# compute log_pi(a_t|s_t), shape = [batch, seq_length]
logprobs_phoneme = get_values_for_actions(sample_logp, sample_translations)
# policy gradient
J = logprobs_phoneme*advantage[:, None]
mask = get_mask_by_eos(T.eq(sample_translations, out_voc.eos_ix))
loss = - T.sum(J*mask) / T.sum(mask)
# regularize with negative entropy. Don't forget the sign!
# note: for entropy you need probabilities for all tokens (sample_logp), not just phoneme_logprobs
entropy = <compute entropy matrix of shape[batch, seq_length], H = -sum(p*log_p), don't forget the sign!>
assert entropy.ndim == 2, "please make sure elementwise entropy is of shape [batch,time]"
loss -= 0.01*T.sum(entropy*mask) / T.sum(mask)
# compute weight updates, clip by norm
grads = T.grad(loss, model.weights)
grads = lasagne.updates.total_norm_constraint(grads, 50)
updates = lasagne.updates.adam(grads, model.weights, learning_rate=1e-5)
train_step = theano.function([input_sequence], loss,
updates=auto_updates+greedy_auto_updates+updates)
```
# Policy gradient training
```
for i in trange(100000):
loss_history.append(
trainer.train_step(sample_batch(
train_words, word_to_translation, 32)[0])
)
if (i+1) % REPORT_FREQ == 0:
clear_output(True)
current_scores = score(test_words)
editdist_history.append(current_scores.mean())
plt.figure(figsize=(8, 4))
plt.subplot(121)
plt.title('val score distribution')
plt.hist(current_scores, bins=20)
plt.subplot(122)
plt.title('val score / traning time')
plt.plot(editdist_history)
plt.grid()
plt.show()
print("J=%.3f, mean score=%.3f" %
(np.mean(loss_history[-10:]), np.mean(editdist_history[-10:])))
model.translate("EXAMPLE;")
```
### Results
```
for word in train_words[:10]:
print("%s -> %s" % (word, translate([word])[0]))
test_scores = []
for start_i in trange(0, len(test_words), 32):
batch_words = test_words[start_i:start_i+32]
batch_trans = translate(batch_words)
distances = list(map(get_distance, batch_words, batch_trans))
test_scores.extend(distances)
print("Supervised test score:", np.mean(test_scores))
# ^^ If you get Out Of Memory, please replace this with batched computation
```
## Step 6: Make it actually work (5++ pts)
<img src=https://github.com/yandexdataschool/Practical_RL/raw/master/yet_another_week/_resource/do_something_scst.png width=400>
In this section we want you to finally __restart with EASY_MODE=False__ and experiment to find a good model/curriculum for that task.
We recommend you to start with the following architecture
```
encoder---decoder
P(y|h)
^
LSTM -> LSTM
^ ^
biLSTM -> LSTM
^ ^
input y_prev
```
__Note:__ you can fit all 4 state tensors of both LSTMs into a in a single state - just assume that it contains, for example, [h0, c0, h1, c1] - pack it in encode and update in decode.
Here are some cool ideas on what you can do then.
__General tips & tricks:__
* In some tensorflow versions and for some layers, it is required that each rnn/gru/lstm cell gets it's own `tf.variable_scope(unique_name, reuse=False)`.
* Otherwise it will complain about wrong tensor sizes because it tries to reuse weights from one rnn to the other.
* You will likely need to adjust pre-training time for such a network.
* Supervised pre-training may benefit from clipping gradients somehow.
* SCST may indulge a higher learning rate in some cases and changing entropy regularizer over time.
* It's often useful to save pre-trained model parameters to not re-train it every time you want new policy gradient parameters.
* When leaving training for nighttime, try setting REPORT_FREQ to a larger value (e.g. 500) not to waste time on it.
__Formal criteria:__
To get 5 points we want you to build an architecture that:
* _doesn't consist of single GRU_
* _works better_ than single GRU baseline.
* We also want you to provide either learning curve or trained model, preferably both
* ... and write a brief report or experiment log describing what you did and how it fared.
### Attention
There's more than one way to connect decoder to encoder
* __Vanilla:__ layer_i of encoder last state goes to layer_i of decoder initial state
* __Every tick:__ feed encoder last state _on every iteration_ of decoder.
* __Attention:__ allow decoder to "peek" at one (or several) positions of encoded sequence on every tick.
The most effective (and cool) of those is, of course, attention.
You can read more about attention [in this nice blog post](https://distill.pub/2016/augmented-rnns/). The easiest way to begin is to use "soft" attention with "additive" or "dot-product" intermediate layers.
__Tips__
* Model usually generalizes better if you no longer allow decoder to see final encoder state
* Once your model made it through several epochs, it is a good idea to visualize attention maps to understand what your model has actually learned
* There's more stuff [here](https://github.com/yandexdataschool/Practical_RL/blob/master/week8_scst/bonus.ipynb)
* If you opted for hard attention, we recommend [gumbel-softmax](https://blog.evjang.com/2016/11/tutorial-categorical-variational.html) instead of sampling. Also please make sure soft attention works fine before you switch to hard.
### UREX
* This is a way to improve exploration in policy-based settings. The main idea is that you find and upweight under-appreciated actions.
* Here's [video](https://www.youtube.com/watch?v=fZNyHoXgV7M&feature=youtu.be&t=3444)
and an [article](https://arxiv.org/abs/1611.09321).
* You may want to reduce batch size 'cuz UREX requires you to sample multiple times per source sentence.
* Once you got it working, try using experience replay with importance sampling instead of (in addition to) basic UREX.
### Some additional ideas:
* (advanced deep learning) It may be a good idea to first train on small phrases and then adapt to larger ones (a.k.a. training curriculum).
* (advanced nlp) You may want to switch from raw utf8 to something like unicode or even syllables to make task easier.
* (advanced nlp) Since hebrew words are written __with vowels omitted__, you may want to use a small Hebrew vowel markup dataset at `he-pron-wiktionary.txt`.
```
assert not EASY_MODE, "make sure you set EASY_MODE = False at the top of the notebook."
```
`[your report/log here or anywhere you please]`
__Contributions:__ This notebook is brought to you by
* Yandex [MT team](https://tech.yandex.com/translate/)
* Denis Mazur ([DeniskaMazur](https://github.com/DeniskaMazur)), Oleg Vasilev ([Omrigan](https://github.com/Omrigan/)), Dmitry Emelyanenko ([TixFeniks](https://github.com/tixfeniks)) and Fedor Ratnikov ([justheuristic](https://github.com/justheuristic/))
* Dataset is parsed from [Wiktionary](https://en.wiktionary.org), which is under CC-BY-SA and GFDL licenses.
| github_jupyter |
# Advanced TTS demos
[](https://colab.research.google.com/github/r9y9/ttslearn/blob/master/notebooks/ch11_Advanced-demos.ipynb)
このページ(ノートブック形式)では、第11章で少し触れた「非自己回帰型ニューラルボコーダ」を用いた、発展的な音声合成のデモを示します。
書籍ではJSUTコーパスのみを扱いましたが、ここではJVSコーパスを用いた多話者音声合成など、他のコーパスを利用した音声合成のデモも紹介します。
このページのデモは、書籍では解説していないことに注意してください。
非自己回帰型ニューラルボコーダの実装には、[kan-bayashi/ParallelWaveGAN](https://github.com/kan-bayashi/ParallelWaveGAN) を利用します。
多話者音声合成の実装は、書籍では実装の解説はしていませんが、第9章、第10章の内容に、軽微な修正を加えることで実現可能です。
興味のある読者は、extra_recipes のソースコードを参照してください。
## 準備
### ttslearn のインストール
```
%%capture
try:
import ttslearn
except ImportError:
!pip install ttslearn
import ttslearn
ttslearn.__version__
```
### パッケージのインポート
```
%pylab inline
import IPython
from IPython.display import Audio
import librosa
import librosa.display
from tqdm.notebook import tqdm
import torch
import random
```
## JSUT
### Tacotron + Parallel WaveGAN (16kHz)
```
from ttslearn.contrib import Tacotron2PWGTTS
if torch.cuda.is_available():
device = torch.device("cuda")
else:
device = torch.device("cpu")
print("Device:", device)
pwg_engine = Tacotron2PWGTTS(device=device)
%time wav, sr = pwg_engine.tts("あらゆる現実を、すべて自分のほうへねじ曲げたのだ。")
IPython.display.display(Audio(wav, rate=sr))
```
### Tacotron + Parallel WaveGAN (24kHz)
```
from ttslearn.pretrained import create_tts_engine
pwg_engine = create_tts_engine("tacotron2_pwg_jsut24k", device=device)
%time wav, sr = pwg_engine.tts("あらゆる現実を、すべて自分のほうへねじ曲げたのだ。")
IPython.display.display(Audio(wav, rate=sr))
```
### Tacotron + HiFi-GAN (24kHz)
```
from ttslearn.pretrained import create_tts_engine
pwg_engine = create_tts_engine("tacotron2_hifipwg_jsut24k", device=device)
%time wav, sr = pwg_engine.tts("あらゆる現実を、すべて自分のほうへねじ曲げたのだ。")
IPython.display.display(Audio(wav, rate=sr))
```
## JVS
### Multi-speaker Tacotron + Parallel WaveGAN (16kHz)
```
pwg_engine = create_tts_engine("multspk_tacotron2_pwg_jvs16k", device=device)
for spk in ["jvs001", "jvs010", "jvs030", "jvs050", "jvs100"]:
text = "タコスと寿司、あなたはどっちが好きですか?わたしは" + ("寿司" if random.random() > 0.2 else "タコス") + "が好きです。"
wav, sr = pwg_engine.tts(text, spk_id=pwg_engine.spk2id[spk])
print(f"Speaker: {spk}")
print(text)
IPython.display.display(Audio(wav, rate=sr))
```
### Multi-speaker Tacotron + Parallel WaveGAN (24kHz)
```
pwg_engine = create_tts_engine("multspk_tacotron2_pwg_jvs24k", device=device)
for spk in ["jvs001", "jvs010", "jvs030", "jvs050", "jvs100"]:
text = "タコスと寿司、あなたはどっちが好きですか?わたしは" + ("寿司" if random.random() > 0.2 else "タコス") + "が好きです。"
wav, sr = pwg_engine.tts(text, spk_id=pwg_engine.spk2id[spk])
print(f"Speaker: {spk}")
print(text)
IPython.display.display(Audio(wav, rate=sr))
```
### Multi-speaker Tacotron + HiFi-GAN (24kHz)
```
pwg_engine = create_tts_engine("multspk_tacotron2_hifipwg_jvs24k", device=device)
for spk in ["jvs001", "jvs010", "jvs030", "jvs050", "jvs100"]:
text = "タコスと寿司、あなたはどっちが好きですか?わたしは" + ("寿司" if random.random() > 0.2 else "タコス") + "が好きです。"
wav, sr = pwg_engine.tts(text, spk_id=pwg_engine.spk2id[spk])
print(f"Speaker: {spk}")
print(text)
IPython.display.display(Audio(wav, rate=sr))
```
## Common voice (ja)
### Multi-speaker Tacotron + Parallel WaveGAN (16kHz)
```
pwg_engine = create_tts_engine("multspk_tacotron2_pwg_cv16k", device=device)
# NOTE: some speaker's voice have significant amount of noise (e.g., speaker 0)
for spk_id in [5, 6, 12, 15, 19]:
text = ("今日" if random.random() > 0.5 else "明日") + "の天気は、" + ("晴れ時々曇り" if random.random() > 0.5 else "晴れ") + "です。"
wav, sr = pwg_engine.tts(text, spk_id=spk_id)
print(f"Speaker ID: {spk_id}")
print(text)
IPython.display.display(Audio(wav, rate=sr))
```
### Multi-speaker Tacotron + Parallel WaveGAN (24kHz)
```
pwg_engine = create_tts_engine("multspk_tacotron2_pwg_cv24k", device=device)
# NOTE: some speaker's voice have significant amount of noise (e.g., speaker 0)
for spk_id in [5, 6, 12, 15, 19]:
text = ("今日" if random.random() > 0.5 else "明日") + "の天気は、" + ("晴れ時々曇り" if random.random() > 0.5 else "晴れ") + "です。"
wav, sr = pwg_engine.tts(text, spk_id=spk_id)
print(f"Speaker ID: {spk_id}")
print(text)
IPython.display.display(Audio(wav, rate=sr))
```
## 参考
- Parallel WaveGAN: https://arxiv.org/abs/1910.11480
- HiFi-GAN: https://arxiv.org/abs/2010.05646
- Parallel WaveGANを含むGANベースの非自己回帰型ニューラルボコーダの実装: https://github.com/kan-bayashi/ParallelWaveGAN
| github_jupyter |
# Glass Classification and Analysis
## What is glass classification?
Basically there are different types of glasses in the dataset which we have to use and predict and analyse the score.
For predcition we have to use model that are there in the machine learning and from that we have to find which model is more efficient.
## Exploring and Analyzing data
### Importing necessary libraries
```
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings('ignore')
import seaborn as sns
plt.rcParams['figure.figsize'] = [10,6]
# Reading data from the file and storing it in dataframe i.e. 'data' using
# pandas library.
data = pd.read_csv('glass.csv')
data.head() # head is used to print the first 5 rows from the dataset.
# Finding total number of rows and columns in the dataset using shape method.
data.shape
```
As shown there are total number of rows=214 and total number of columns=10 which shows 9 different types of glasses and 1 column with the name as 'type'.
```
data.info()
```
There are no null values in any of the column in the dataframe as shown above using the info() method.
```
type = data['Type'].groupby(data['Type']).count()
type
```
As mentioned in the above cell output that majority data is of type 1 and 2.
Describe method is used to get mean, standard deviation, min, max, etc of all the columns in the dataframe as shown below.
```
data.describe()
# We will find the correlation using the corr() method whith 'Type' column.
data.corr()['Type'].sort_values()
plt.figure(figsize=(10,6))
ax = sns.heatmap(data.corr(), cmap="YlGnBu", annot=True)
ax
fig = plt.figure(figsize = (15,10))
ax = fig.gca()
data.hist(ax=ax)
plt.show()
sns.countplot(data['Type'])
```
we have to take care here about one thing that is, training set is small. so what we will do ,that we will train and classify roughly first and then we can improve over it considering over or under fitting.
#### Preprocessing Data
```
#Import MinMaxScaler module from scikit-learn library.
from sklearn.preprocessing import MinMaxScaler
# Now we will drop the Type column from the dataframe so that we can easily
# normalize the data set.
X = data.drop('Type',axis=1)
y = data['Type']
x = X.values # Returns a numpy array.
min_max_scaler = MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df = pd.DataFrame(x_scaled)
df
```
#### Splitting Data
```
# Now we will split the dataset into the training and testing data using train_test_split module
# from scikit-learn library.
from sklearn.model_selection import train_test_split
# Dividing the dataset into train and test data with a 7:3 ratio as shown below.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.30, random_state=42)
print(X_train.shape)
```
### Different Classification techniques with the accuracy of the model
#### DecisionTreeClassifier model
##### 1. ID3 Decision Tree
ID3 stands for Iterative Dichotomiser 3 and is named such because the algorithm iteratively (repeatedly) dichotomizes(divides) features into two or more groups at each step. ID3 uses a top-down greedy approach to build a decision tree.
```
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
clf = DecisionTreeClassifier(criterion='entropy')
clf = clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
accuracy_score(y_test, y_pred)
```
#### Advantages of ID3 Decision Tree Algorithm
- Builds the fastest tree.
- Understandable prediction rules are created from the training data.
- Only need to test enough attributes until all data is classified.
- Whole dataset is searched to create tree.
#### Disadvantages of ID3 Decision Tree Algorithm
- Data may be over-fitted or over-classified, if a small sample is tested.
- Only one attribute at a time is tested for making a decision.
- Classifying continuous data may be computationally expensive, as many trees must be generated to see where to break the continum.
#### 2. CART Decision Tree
Classification And Regression Trees (CART) algorithm is a classification algorithm for building a decision tree based on Gini's impurity index as splitting criterion. CART is a binary tree build by splitting node into two child nodes repeatedly.
This algorithm works repeatedly in three steps as mentioned below:
1. Find each feature’s best split. For each feature with K different values there exist K-1 possible splits. Find the split, which maximizes the splitting criterion. The resulting set of splits contains best splits (one for each feature).
2. Find the node’s best split. Among the best splits from Step i find the one, which maximizes the splitting criterion.
3. Split the node using best node split from Step ii and repeat from Step i until stopping criterion is satisfied.
```
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import mean_squared_error
import math
clf = DecisionTreeClassifier()
clf.fit(X_train,y_train)
y_pred = clf.predict(X_test)
asc = accuracy_score(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
rmse = math.sqrt(mse)
print(asc, mse, rmse)
```
#### Advantages of CART Decision Tree Algorithm
- Transparent and easy to understand.
- Decision trees can inherently perform multiclass classification.
- They can handle both numerical and categorical data.
- Nonlinear relationships among features do not affect the performance of the decision trees.
#### Disadvantages of CART Decision Tree Algorithm
- A small change in the dataset can make the tree structure unstable which can cause variance.
- It does not work well if there are smooth limits.
- It has high variance and it is unstable.
- Decision tree learners create underfit trees if some classes are imbalanced.
#### 3. RandomForestClassifier model
###### What is Random Forest Classifier?
A random forest is a machine learning technique that's used to solve regression and classification problems. It utilizes ensemble learning, which is a technique that combines many classifiers to provide solutions to complex problems. A random forest algorithm consists of many decision trees.
Random forest is a supervised learning algorithm which is used for both classification as well as regression. Similarly, random forest algorithm creates decision trees on data samples and then gets the prediction from each of them and finally selects the best solution by means of voting.
```
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import mean_squared_error
from math import sqrt
clf = RandomForestClassifier(n_estimators=100, max_depth=2, random_state=0)
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
asc = accuracy_score(y_test, y_pred)
mse = mean_squared_error(y_test, y_pred)
rmse = math.sqrt(mse)
print(asc, mse, rmse)
```
#### Advantages of Random Forest
- Random Forest is capable of performing both Classification and Regression tasks.
- It is capable of handling large datasets with high dimensionality.
- It enhances the accuracy of the model and prevents the overfitting issue.
#### Disadvantages of Random Forest
- Although random forest can be used for both classification and regression tasks, it is not more suitable for Regression tasks.
## Conclusion
As there are different classification techniques to predict the accuracy of the glass dataset. We have tried 3 models as shown above such that we can select the model with the high accuracy result. We used 'ID3 Decision Tree', 'CART Decision Tree', 'Random Forest Classifier' and from that we find that the accuracy of the 'CART decision tree' is highest among all and for our dataset the 'CART decision tree' is the best model for predicting the results.
| github_jupyter |
This notebook is accompanied with a [series of blog posts](https://medium.com/@margaretmz/selfie2anime-with-tflite-part-1-overview-f97500800ffe). To follow along with this Colab Notebook we recommend that you also read [this blog post](https://medium.com/@margaretmz/selfie2anime-with-tflite-part-2-tflite-model-84002cf521dc) simultaneously.
**Authors**: [Margaret Maynard-Reid](https://twitter.com/margaretmz) and [Sayak Paul](https://twitter.com/RisingSayak)
**Reviewers**: [Khanh LeViet](https://twitter.com/khanhlvg) and [Hoi Lam](https://twitter.com/hoitab)
Shoutout to Khanh LeViet and Lu Wang from the TensorFlow Lite team for their guidance. Main codebase of UGATIT is here: https://github.com/taki0112/UGATIT.
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/margaretmz/selfie2anime-e2e-tutorial/blob/master/ml/Selfie2Anime_Model_Conversion.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/margaretmz/selfie2anime-e2e-tutorial/tree/master/ml/Selfie2Anime_Model_Conversion.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## Initial setup
```
!pip install tensorflow==1.14
import tensorflow as tf
print(tf.__version__)
```
You can safely ignore the warnings.
```
import os
import tempfile
```
## Loading the checkpoints
To use the Kaggle API, sign up for a Kaggle account at https://www.kaggle.com. Then go to the 'Account' tab of your user profile (https://www.kaggle.com/account) and select 'Create API Token'. This will trigger the download of `kaggle.json`, a file containing your API credentials.
```
os.environ['KAGGLE_USERNAME'] = "" # TODO: enter your Kaggle user name here
os.environ['KAGGLE_KEY'] = "" # TODO: enter your Kaggle key here
!kaggle datasets download -d t04glovern/ugatit-selfie2anime-pretrained
!unzip -qq /content/ugatit-selfie2anime-pretrained.zip
```
**Note**: There are other versions of the UGATIT model that you can check [here](https://github.com/taki0112/UGATIT/#pretrained-model). Here, we are using an optimized one.
## Some utils
```
!git clone https://github.com/taki0112/UGATIT
%cd UGATIT
# Reference: https://dev.to/0xbf/use-dot-syntax-to-access-dictionary-key-python-tips-10ec
class DictX(dict):
def __getattr__(self, key):
try:
return self[key]
except KeyError as k:
raise AttributeError(k)
def __setattr__(self, key, value):
self[key] = value
def __delattr__(self, key):
try:
del self[key]
except KeyError as k:
raise AttributeError(k)
def __repr__(self):
return '<DictX ' + dict.__repr__(self) + '>'
# This is needed just to initialize `UGATIT` class
args = dict(phase='test',
light=True,
dataset='selfie2anime',
epoch=100,
iteration=10000,
batch_size=1,
print_freq=1000,
save_freq=1000,
decay_flag=True,
decay_epoch=50,
lr=0.0001,
GP_ld=10,
adv_weight=1,
cycle_weight=10,
identity_weight=10,
cam_weight=1000,
gan_type='lsgan',
smoothing=True,
ch=64,
n_res=4,
n_dis=6,
n_critic=1,
sn=True,
img_size=256,
img_ch=3,
augment_flag=False,
checkpoint_dir='/content',
result_dir='/content',
log_dir='/content',
sample_dir='/content')
# Wrap the arguments in a dictionary because this particular format is required
# in order to instantiate the `UGATIT` class
data = DictX(args)
```
## UGATIT class for convenience
Run this block of code to get access to some helper functions. Otherwise, the rest of this Colab may not run correctly.
```
#@title
from ops import *
from utils import *
from glob import glob
import time
from tensorflow.contrib.data import prefetch_to_device, shuffle_and_repeat, map_and_batch
import numpy as np
class UGATIT(object) :
def __init__(self, sess, args):
self.light = args.light
if self.light :
self.model_name = 'UGATIT_light'
else :
self.model_name = 'UGATIT'
self.sess = sess
self.phase = args.phase
self.checkpoint_dir = args.checkpoint_dir
self.result_dir = args.result_dir
self.log_dir = args.log_dir
self.dataset_name = args.dataset
self.augment_flag = args.augment_flag
self.epoch = args.epoch
self.iteration = args.iteration
self.decay_flag = args.decay_flag
self.decay_epoch = args.decay_epoch
self.gan_type = args.gan_type
self.batch_size = args.batch_size
self.print_freq = args.print_freq
self.save_freq = args.save_freq
self.init_lr = args.lr
self.ch = args.ch
""" Weight """
self.adv_weight = args.adv_weight
self.cycle_weight = args.cycle_weight
self.identity_weight = args.identity_weight
self.cam_weight = args.cam_weight
self.ld = args.GP_ld
self.smoothing = args.smoothing
""" Generator """
self.n_res = args.n_res
""" Discriminator """
self.n_dis = args.n_dis
self.n_critic = args.n_critic
self.sn = args.sn
self.img_size = args.img_size
self.img_ch = args.img_ch
self.sample_dir = os.path.join(args.sample_dir, self.model_dir)
check_folder(self.sample_dir)
# self.trainA, self.trainB = prepare_data(dataset_name=self.dataset_name, size=self.img_size
self.trainA_dataset = glob('./dataset/{}/*.*'.format(self.dataset_name + '/trainA'))
self.trainB_dataset = glob('./dataset/{}/*.*'.format(self.dataset_name + '/trainB'))
self.dataset_num = max(len(self.trainA_dataset), len(self.trainB_dataset))
print()
print("##### Information #####")
print("# light : ", self.light)
print("# gan type : ", self.gan_type)
print("# dataset : ", self.dataset_name)
print("# max dataset number : ", self.dataset_num)
print("# batch_size : ", self.batch_size)
print("# epoch : ", self.epoch)
print("# iteration per epoch : ", self.iteration)
print("# smoothing : ", self.smoothing)
print()
print("##### Generator #####")
print("# residual blocks : ", self.n_res)
print()
print("##### Discriminator #####")
print("# discriminator layer : ", self.n_dis)
print("# the number of critic : ", self.n_critic)
print("# spectral normalization : ", self.sn)
print()
print("##### Weight #####")
print("# adv_weight : ", self.adv_weight)
print("# cycle_weight : ", self.cycle_weight)
print("# identity_weight : ", self.identity_weight)
print("# cam_weight : ", self.cam_weight)
##################################################################################
# Generator
##################################################################################
def generator(self, x_init, reuse=False, scope="generator"):
channel = self.ch
with tf.variable_scope(scope, reuse=reuse) :
x = conv(x_init, channel, kernel=7, stride=1, pad=3, pad_type='reflect', scope='conv')
x = instance_norm(x, scope='ins_norm')
x = relu(x)
# Down-Sampling
for i in range(2) :
x = conv(x, channel*2, kernel=3, stride=2, pad=1, pad_type='reflect', scope='conv_'+str(i))
x = instance_norm(x, scope='ins_norm_'+str(i))
x = relu(x)
channel = channel * 2
# Down-Sampling Bottleneck
for i in range(self.n_res):
x = resblock(x, channel, scope='resblock_' + str(i))
# Class Activation Map
cam_x = global_avg_pooling(x)
cam_gap_logit, cam_x_weight = fully_connected_with_w(cam_x, scope='CAM_logit')
x_gap = tf.multiply(x, cam_x_weight)
cam_x = global_max_pooling(x)
cam_gmp_logit, cam_x_weight = fully_connected_with_w(cam_x, reuse=True, scope='CAM_logit')
x_gmp = tf.multiply(x, cam_x_weight)
cam_logit = tf.concat([cam_gap_logit, cam_gmp_logit], axis=-1)
x = tf.concat([x_gap, x_gmp], axis=-1)
x = conv(x, channel, kernel=1, stride=1, scope='conv_1x1')
x = relu(x)
heatmap = tf.squeeze(tf.reduce_sum(x, axis=-1))
# Gamma, Beta block
gamma, beta = self.MLP(x, reuse=reuse)
# Up-Sampling Bottleneck
for i in range(self.n_res):
x = adaptive_ins_layer_resblock(x, channel, gamma, beta, smoothing=self.smoothing, scope='adaptive_resblock' + str(i))
# Up-Sampling
for i in range(2) :
x = up_sample(x, scale_factor=2)
x = conv(x, channel//2, kernel=3, stride=1, pad=1, pad_type='reflect', scope='up_conv_'+str(i))
x = layer_instance_norm(x, scope='layer_ins_norm_'+str(i))
x = relu(x)
channel = channel // 2
x = conv(x, channels=3, kernel=7, stride=1, pad=3, pad_type='reflect', scope='G_logit')
x = tanh(x)
return x, cam_logit, heatmap
def MLP(self, x, use_bias=True, reuse=False, scope='MLP'):
channel = self.ch * self.n_res
if self.light :
x = global_avg_pooling(x)
with tf.variable_scope(scope, reuse=reuse):
for i in range(2) :
x = fully_connected(x, channel, use_bias, scope='linear_' + str(i))
x = relu(x)
gamma = fully_connected(x, channel, use_bias, scope='gamma')
beta = fully_connected(x, channel, use_bias, scope='beta')
gamma = tf.reshape(gamma, shape=[self.batch_size, 1, 1, channel])
beta = tf.reshape(beta, shape=[self.batch_size, 1, 1, channel])
return gamma, beta
##################################################################################
# Discriminator
##################################################################################
def discriminator(self, x_init, reuse=False, scope="discriminator"):
D_logit = []
D_CAM_logit = []
with tf.variable_scope(scope, reuse=reuse) :
local_x, local_cam, local_heatmap = self.discriminator_local(x_init, reuse=reuse, scope='local')
global_x, global_cam, global_heatmap = self.discriminator_global(x_init, reuse=reuse, scope='global')
D_logit.extend([local_x, global_x])
D_CAM_logit.extend([local_cam, global_cam])
return D_logit, D_CAM_logit, local_heatmap, global_heatmap
def discriminator_global(self, x_init, reuse=False, scope='discriminator_global'):
with tf.variable_scope(scope, reuse=reuse):
channel = self.ch
x = conv(x_init, channel, kernel=4, stride=2, pad=1, pad_type='reflect', sn=self.sn, scope='conv_0')
x = lrelu(x, 0.2)
for i in range(1, self.n_dis - 1):
x = conv(x, channel * 2, kernel=4, stride=2, pad=1, pad_type='reflect', sn=self.sn, scope='conv_' + str(i))
x = lrelu(x, 0.2)
channel = channel * 2
x = conv(x, channel * 2, kernel=4, stride=1, pad=1, pad_type='reflect', sn=self.sn, scope='conv_last')
x = lrelu(x, 0.2)
channel = channel * 2
cam_x = global_avg_pooling(x)
cam_gap_logit, cam_x_weight = fully_connected_with_w(cam_x, sn=self.sn, scope='CAM_logit')
x_gap = tf.multiply(x, cam_x_weight)
cam_x = global_max_pooling(x)
cam_gmp_logit, cam_x_weight = fully_connected_with_w(cam_x, sn=self.sn, reuse=True, scope='CAM_logit')
x_gmp = tf.multiply(x, cam_x_weight)
cam_logit = tf.concat([cam_gap_logit, cam_gmp_logit], axis=-1)
x = tf.concat([x_gap, x_gmp], axis=-1)
x = conv(x, channel, kernel=1, stride=1, scope='conv_1x1')
x = lrelu(x, 0.2)
heatmap = tf.squeeze(tf.reduce_sum(x, axis=-1))
x = conv(x, channels=1, kernel=4, stride=1, pad=1, pad_type='reflect', sn=self.sn, scope='D_logit')
return x, cam_logit, heatmap
def discriminator_local(self, x_init, reuse=False, scope='discriminator_local'):
with tf.variable_scope(scope, reuse=reuse) :
channel = self.ch
x = conv(x_init, channel, kernel=4, stride=2, pad=1, pad_type='reflect', sn=self.sn, scope='conv_0')
x = lrelu(x, 0.2)
for i in range(1, self.n_dis - 2 - 1):
x = conv(x, channel * 2, kernel=4, stride=2, pad=1, pad_type='reflect', sn=self.sn, scope='conv_' + str(i))
x = lrelu(x, 0.2)
channel = channel * 2
x = conv(x, channel * 2, kernel=4, stride=1, pad=1, pad_type='reflect', sn=self.sn, scope='conv_last')
x = lrelu(x, 0.2)
channel = channel * 2
cam_x = global_avg_pooling(x)
cam_gap_logit, cam_x_weight = fully_connected_with_w(cam_x, sn=self.sn, scope='CAM_logit')
x_gap = tf.multiply(x, cam_x_weight)
cam_x = global_max_pooling(x)
cam_gmp_logit, cam_x_weight = fully_connected_with_w(cam_x, sn=self.sn, reuse=True, scope='CAM_logit')
x_gmp = tf.multiply(x, cam_x_weight)
cam_logit = tf.concat([cam_gap_logit, cam_gmp_logit], axis=-1)
x = tf.concat([x_gap, x_gmp], axis=-1)
x = conv(x, channel, kernel=1, stride=1, scope='conv_1x1')
x = lrelu(x, 0.2)
heatmap = tf.squeeze(tf.reduce_sum(x, axis=-1))
x = conv(x, channels=1, kernel=4, stride=1, pad=1, pad_type='reflect', sn=self.sn, scope='D_logit')
return x, cam_logit, heatmap
##################################################################################
# Model
##################################################################################
def generate_a2b(self, x_A, reuse=False):
out, cam, _ = self.generator(x_A, reuse=reuse, scope="generator_B")
return out, cam
def generate_b2a(self, x_B, reuse=False):
out, cam, _ = self.generator(x_B, reuse=reuse, scope="generator_A")
return out, cam
def discriminate_real(self, x_A, x_B):
real_A_logit, real_A_cam_logit, _, _ = self.discriminator(x_A, scope="discriminator_A")
real_B_logit, real_B_cam_logit, _, _ = self.discriminator(x_B, scope="discriminator_B")
return real_A_logit, real_A_cam_logit, real_B_logit, real_B_cam_logit
def discriminate_fake(self, x_ba, x_ab):
fake_A_logit, fake_A_cam_logit, _, _ = self.discriminator(x_ba, reuse=True, scope="discriminator_A")
fake_B_logit, fake_B_cam_logit, _, _ = self.discriminator(x_ab, reuse=True, scope="discriminator_B")
return fake_A_logit, fake_A_cam_logit, fake_B_logit, fake_B_cam_logit
def gradient_panalty(self, real, fake, scope="discriminator_A"):
if self.gan_type.__contains__('dragan'):
eps = tf.random_uniform(shape=tf.shape(real), minval=0., maxval=1.)
_, x_var = tf.nn.moments(real, axes=[0, 1, 2, 3])
x_std = tf.sqrt(x_var) # magnitude of noise decides the size of local region
fake = real + 0.5 * x_std * eps
alpha = tf.random_uniform(shape=[self.batch_size, 1, 1, 1], minval=0., maxval=1.)
interpolated = real + alpha * (fake - real)
logit, cam_logit, _, _ = self.discriminator(interpolated, reuse=True, scope=scope)
GP = []
cam_GP = []
for i in range(2) :
grad = tf.gradients(logit[i], interpolated)[0] # gradient of D(interpolated)
grad_norm = tf.norm(flatten(grad), axis=1) # l2 norm
# WGAN - LP
if self.gan_type == 'wgan-lp' :
GP.append(self.ld * tf.reduce_mean(tf.square(tf.maximum(0.0, grad_norm - 1.))))
elif self.gan_type == 'wgan-gp' or self.gan_type == 'dragan':
GP.append(self.ld * tf.reduce_mean(tf.square(grad_norm - 1.)))
for i in range(2) :
grad = tf.gradients(cam_logit[i], interpolated)[0] # gradient of D(interpolated)
grad_norm = tf.norm(flatten(grad), axis=1) # l2 norm
# WGAN - LP
if self.gan_type == 'wgan-lp' :
cam_GP.append(self.ld * tf.reduce_mean(tf.square(tf.maximum(0.0, grad_norm - 1.))))
elif self.gan_type == 'wgan-gp' or self.gan_type == 'dragan':
cam_GP.append(self.ld * tf.reduce_mean(tf.square(grad_norm - 1.)))
return sum(GP), sum(cam_GP)
def build_model(self):
if self.phase == 'train' :
self.lr = tf.placeholder(tf.float32, name='learning_rate')
""" Input Image"""
Image_Data_Class = ImageData(self.img_size, self.img_ch, self.augment_flag)
trainA = tf.data.Dataset.from_tensor_slices(self.trainA_dataset)
trainB = tf.data.Dataset.from_tensor_slices(self.trainB_dataset)
gpu_device = '/gpu:0'
trainA = trainA.apply(shuffle_and_repeat(self.dataset_num)).apply(map_and_batch(Image_Data_Class.image_processing, self.batch_size, num_parallel_batches=16, drop_remainder=True)).apply(prefetch_to_device(gpu_device, None))
trainB = trainB.apply(shuffle_and_repeat(self.dataset_num)).apply(map_and_batch(Image_Data_Class.image_processing, self.batch_size, num_parallel_batches=16, drop_remainder=True)).apply(prefetch_to_device(gpu_device, None))
trainA_iterator = trainA.make_one_shot_iterator()
trainB_iterator = trainB.make_one_shot_iterator()
self.domain_A = trainA_iterator.get_next()
self.domain_B = trainB_iterator.get_next()
""" Define Generator, Discriminator """
x_ab, cam_ab = self.generate_a2b(self.domain_A) # real a
x_ba, cam_ba = self.generate_b2a(self.domain_B) # real b
x_aba, _ = self.generate_b2a(x_ab, reuse=True) # real b
x_bab, _ = self.generate_a2b(x_ba, reuse=True) # real a
x_aa, cam_aa = self.generate_b2a(self.domain_A, reuse=True) # fake b
x_bb, cam_bb = self.generate_a2b(self.domain_B, reuse=True) # fake a
real_A_logit, real_A_cam_logit, real_B_logit, real_B_cam_logit = self.discriminate_real(self.domain_A, self.domain_B)
fake_A_logit, fake_A_cam_logit, fake_B_logit, fake_B_cam_logit = self.discriminate_fake(x_ba, x_ab)
""" Define Loss """
if self.gan_type.__contains__('wgan') or self.gan_type == 'dragan' :
GP_A, GP_CAM_A = self.gradient_panalty(real=self.domain_A, fake=x_ba, scope="discriminator_A")
GP_B, GP_CAM_B = self.gradient_panalty(real=self.domain_B, fake=x_ab, scope="discriminator_B")
else :
GP_A, GP_CAM_A = 0, 0
GP_B, GP_CAM_B = 0, 0
G_ad_loss_A = (generator_loss(self.gan_type, fake_A_logit) + generator_loss(self.gan_type, fake_A_cam_logit))
G_ad_loss_B = (generator_loss(self.gan_type, fake_B_logit) + generator_loss(self.gan_type, fake_B_cam_logit))
D_ad_loss_A = (discriminator_loss(self.gan_type, real_A_logit, fake_A_logit) + discriminator_loss(self.gan_type, real_A_cam_logit, fake_A_cam_logit) + GP_A + GP_CAM_A)
D_ad_loss_B = (discriminator_loss(self.gan_type, real_B_logit, fake_B_logit) + discriminator_loss(self.gan_type, real_B_cam_logit, fake_B_cam_logit) + GP_B + GP_CAM_B)
reconstruction_A = L1_loss(x_aba, self.domain_A) # reconstruction
reconstruction_B = L1_loss(x_bab, self.domain_B) # reconstruction
identity_A = L1_loss(x_aa, self.domain_A)
identity_B = L1_loss(x_bb, self.domain_B)
cam_A = cam_loss(source=cam_ba, non_source=cam_aa)
cam_B = cam_loss(source=cam_ab, non_source=cam_bb)
Generator_A_gan = self.adv_weight * G_ad_loss_A
Generator_A_cycle = self.cycle_weight * reconstruction_B
Generator_A_identity = self.identity_weight * identity_A
Generator_A_cam = self.cam_weight * cam_A
Generator_B_gan = self.adv_weight * G_ad_loss_B
Generator_B_cycle = self.cycle_weight * reconstruction_A
Generator_B_identity = self.identity_weight * identity_B
Generator_B_cam = self.cam_weight * cam_B
Generator_A_loss = Generator_A_gan + Generator_A_cycle + Generator_A_identity + Generator_A_cam
Generator_B_loss = Generator_B_gan + Generator_B_cycle + Generator_B_identity + Generator_B_cam
Discriminator_A_loss = self.adv_weight * D_ad_loss_A
Discriminator_B_loss = self.adv_weight * D_ad_loss_B
self.Generator_loss = Generator_A_loss + Generator_B_loss + regularization_loss('generator')
self.Discriminator_loss = Discriminator_A_loss + Discriminator_B_loss + regularization_loss('discriminator')
""" Result Image """
self.fake_A = x_ba
self.fake_B = x_ab
self.real_A = self.domain_A
self.real_B = self.domain_B
""" Training """
t_vars = tf.trainable_variables()
G_vars = [var for var in t_vars if 'generator' in var.name]
D_vars = [var for var in t_vars if 'discriminator' in var.name]
self.G_optim = tf.train.AdamOptimizer(self.lr, beta1=0.5, beta2=0.999).minimize(self.Generator_loss, var_list=G_vars)
self.D_optim = tf.train.AdamOptimizer(self.lr, beta1=0.5, beta2=0.999).minimize(self.Discriminator_loss, var_list=D_vars)
"""" Summary """
self.all_G_loss = tf.summary.scalar("Generator_loss", self.Generator_loss)
self.all_D_loss = tf.summary.scalar("Discriminator_loss", self.Discriminator_loss)
self.G_A_loss = tf.summary.scalar("G_A_loss", Generator_A_loss)
self.G_A_gan = tf.summary.scalar("G_A_gan", Generator_A_gan)
self.G_A_cycle = tf.summary.scalar("G_A_cycle", Generator_A_cycle)
self.G_A_identity = tf.summary.scalar("G_A_identity", Generator_A_identity)
self.G_A_cam = tf.summary.scalar("G_A_cam", Generator_A_cam)
self.G_B_loss = tf.summary.scalar("G_B_loss", Generator_B_loss)
self.G_B_gan = tf.summary.scalar("G_B_gan", Generator_B_gan)
self.G_B_cycle = tf.summary.scalar("G_B_cycle", Generator_B_cycle)
self.G_B_identity = tf.summary.scalar("G_B_identity", Generator_B_identity)
self.G_B_cam = tf.summary.scalar("G_B_cam", Generator_B_cam)
self.D_A_loss = tf.summary.scalar("D_A_loss", Discriminator_A_loss)
self.D_B_loss = tf.summary.scalar("D_B_loss", Discriminator_B_loss)
self.rho_var = []
for var in tf.trainable_variables():
if 'rho' in var.name:
self.rho_var.append(tf.summary.histogram(var.name, var))
self.rho_var.append(tf.summary.scalar(var.name + "_min", tf.reduce_min(var)))
self.rho_var.append(tf.summary.scalar(var.name + "_max", tf.reduce_max(var)))
self.rho_var.append(tf.summary.scalar(var.name + "_mean", tf.reduce_mean(var)))
g_summary_list = [self.G_A_loss, self.G_A_gan, self.G_A_cycle, self.G_A_identity, self.G_A_cam,
self.G_B_loss, self.G_B_gan, self.G_B_cycle, self.G_B_identity, self.G_B_cam,
self.all_G_loss]
g_summary_list.extend(self.rho_var)
d_summary_list = [self.D_A_loss, self.D_B_loss, self.all_D_loss]
self.G_loss = tf.summary.merge(g_summary_list)
self.D_loss = tf.summary.merge(d_summary_list)
else :
""" Test """
self.test_domain_A = tf.placeholder(tf.float32, [1, self.img_size, self.img_size, self.img_ch], name='test_domain_A')
self.test_domain_B = tf.placeholder(tf.float32, [1, self.img_size, self.img_size, self.img_ch], name='test_domain_B')
self.test_fake_B, _ = self.generate_a2b(self.test_domain_A)
self.test_fake_A, _ = self.generate_b2a(self.test_domain_B)
def train(self):
# initialize all variables
tf.global_variables_initializer().run()
# saver to save model
self.saver = tf.train.Saver()
# summary writer
self.writer = tf.summary.FileWriter(self.log_dir + '/' + self.model_dir, self.sess.graph)
# restore check-point if it exits
could_load, checkpoint_counter = self.load(self.checkpoint_dir)
if could_load:
start_epoch = (int)(checkpoint_counter / self.iteration)
start_batch_id = checkpoint_counter - start_epoch * self.iteration
counter = checkpoint_counter
print(" [*] Load SUCCESS")
else:
start_epoch = 0
start_batch_id = 0
counter = 1
print(" [!] Load failed...")
# loop for epoch
start_time = time.time()
past_g_loss = -1.
lr = self.init_lr
for epoch in range(start_epoch, self.epoch):
# lr = self.init_lr if epoch < self.decay_epoch else self.init_lr * (self.epoch - epoch) / (self.epoch - self.decay_epoch)
if self.decay_flag :
#lr = self.init_lr * pow(0.5, epoch // self.decay_epoch)
lr = self.init_lr if epoch < self.decay_epoch else self.init_lr * (self.epoch - epoch) / (self.epoch - self.decay_epoch)
for idx in range(start_batch_id, self.iteration):
train_feed_dict = {
self.lr : lr
}
# Update D
_, d_loss, summary_str = self.sess.run([self.D_optim,
self.Discriminator_loss, self.D_loss], feed_dict = train_feed_dict)
self.writer.add_summary(summary_str, counter)
# Update G
g_loss = None
if (counter - 1) % self.n_critic == 0 :
batch_A_images, batch_B_images, fake_A, fake_B, _, g_loss, summary_str = self.sess.run([self.real_A, self.real_B,
self.fake_A, self.fake_B,
self.G_optim,
self.Generator_loss, self.G_loss], feed_dict = train_feed_dict)
self.writer.add_summary(summary_str, counter)
past_g_loss = g_loss
# display training status
counter += 1
if g_loss == None :
g_loss = past_g_loss
print("Epoch: [%2d] [%5d/%5d] time: %4.4f d_loss: %.8f, g_loss: %.8f" % (epoch, idx, self.iteration, time.time() - start_time, d_loss, g_loss))
if np.mod(idx+1, self.print_freq) == 0 :
save_images(batch_A_images, [self.batch_size, 1],
'./{}/real_A_{:03d}_{:05d}.png'.format(self.sample_dir, epoch, idx+1))
# save_images(batch_B_images, [self.batch_size, 1],
# './{}/real_B_{:03d}_{:05d}.png'.format(self.sample_dir, epoch, idx+1))
# save_images(fake_A, [self.batch_size, 1],
# './{}/fake_A_{:03d}_{:05d}.png'.format(self.sample_dir, epoch, idx+1))
save_images(fake_B, [self.batch_size, 1],
'./{}/fake_B_{:03d}_{:05d}.png'.format(self.sample_dir, epoch, idx+1))
if np.mod(idx + 1, self.save_freq) == 0:
self.save(self.checkpoint_dir, counter)
# After an epoch, start_batch_id is set to zero
# non-zero value is only for the first epoch after loading pre-trained model
start_batch_id = 0
# save model for final step
self.save(self.checkpoint_dir, counter)
@property
def model_dir(self):
n_res = str(self.n_res) + 'resblock'
n_dis = str(self.n_dis) + 'dis'
if self.smoothing :
smoothing = '_smoothing'
else :
smoothing = ''
if self.sn :
sn = '_sn'
else :
sn = ''
return "{}_{}_{}_{}_{}_{}_{}_{}_{}_{}{}{}".format(self.model_name, self.dataset_name,
self.gan_type, n_res, n_dis,
self.n_critic,
self.adv_weight, self.cycle_weight, self.identity_weight, self.cam_weight, sn, smoothing)
def save(self, checkpoint_dir, step):
checkpoint_dir = os.path.join(checkpoint_dir, self.model_dir)
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
self.saver.save(self.sess, os.path.join(checkpoint_dir, self.model_name + '.model'), global_step=step)
def load(self, checkpoint_dir):
print(" [*] Reading checkpoints...")
checkpoint_dir = os.path.join(checkpoint_dir, self.model_dir)
ckpt = tf.train.get_checkpoint_state(checkpoint_dir)
if ckpt and ckpt.model_checkpoint_path:
ckpt_name = os.path.basename(ckpt.model_checkpoint_path)
self.saver.restore(self.sess, os.path.join(checkpoint_dir, ckpt_name))
counter = int(ckpt_name.split('-')[-1])
print(" [*] Success to read {}".format(ckpt_name))
return True, counter
else:
print(" [*] Failed to find a checkpoint")
return False, 0
def test(self):
tf.global_variables_initializer().run()
test_A_files = glob('./dataset/{}/*.*'.format(self.dataset_name + '/testA'))
test_B_files = glob('./dataset/{}/*.*'.format(self.dataset_name + '/testB'))
self.saver = tf.train.Saver()
could_load, checkpoint_counter = self.load(self.checkpoint_dir)
self.result_dir = os.path.join(self.result_dir, self.model_dir)
check_folder(self.result_dir)
if could_load :
print(" [*] Load SUCCESS")
else :
print(" [!] Load failed...")
# write html for visual comparison
index_path = os.path.join(self.result_dir, 'index.html')
index = open(index_path, 'w')
index.write("<html><body><table><tr>")
index.write("<th>name</th><th>input</th><th>output</th></tr>")
for sample_file in test_A_files : # A -> B
print('Processing A image: ' + sample_file)
sample_image = np.asarray(load_test_data(sample_file, size=self.img_size))
image_path = os.path.join(self.result_dir,'{0}'.format(os.path.basename(sample_file)))
fake_img = self.sess.run(self.test_fake_B, feed_dict = {self.test_domain_A : sample_image})
save_images(fake_img, [1, 1], image_path)
index.write("<td>%s</td>" % os.path.basename(image_path))
index.write("<td><img src='%s' width='%d' height='%d'></td>" % (sample_file if os.path.isabs(sample_file) else (
'../..' + os.path.sep + sample_file), self.img_size, self.img_size))
index.write("<td><img src='%s' width='%d' height='%d'></td>" % (image_path if os.path.isabs(image_path) else (
'../..' + os.path.sep + image_path), self.img_size, self.img_size))
index.write("</tr>")
for sample_file in test_B_files : # B -> A
print('Processing B image: ' + sample_file)
sample_image = np.asarray(load_test_data(sample_file, size=self.img_size))
image_path = os.path.join(self.result_dir,'{0}'.format(os.path.basename(sample_file)))
fake_img = self.sess.run(self.test_fake_A, feed_dict = {self.test_domain_B : sample_image})
save_images(fake_img, [1, 1], image_path)
index.write("<td>%s</td>" % os.path.basename(image_path))
index.write("<td><img src='%s' width='%d' height='%d'></td>" % (sample_file if os.path.isabs(sample_file) else (
'../..' + os.path.sep + sample_file), self.img_size, self.img_size))
index.write("<td><img src='%s' width='%d' height='%d'></td>" % (image_path if os.path.isabs(image_path) else (
'../..' + os.path.sep + image_path), self.img_size, self.img_size))
index.write("</tr>")
index.close()
```
## Build and initialize the model
[Reference](https://github.com/tensorflow/magenta/blob/85ef5267513f62f4a40b01b2a1ee488f90f64a13/magenta/models/arbitrary_image_stylization/arbitrary_image_stylization_convert_tflite.py#L46) of the following utility.
```
def load_checkpoint(sess, checkpoint):
"""Loads a checkpoint file into the session.
Args:
sess: tf.Session, the TF session to load variables from the checkpoint to.
checkpoint: str, path to the checkpoint file.
"""
model_saver = tf.train.Saver(tf.global_variables())
checkpoint = os.path.expanduser(checkpoint)
if tf.gfile.IsDirectory(checkpoint):
checkpoint = tf.train.latest_checkpoint(checkpoint)
tf.logging.info('loading latest checkpoint file: {}'.format(checkpoint))
model_saver.restore(sess, checkpoint)
```
## Exporting to `SavedModel`
Note that we will only be using the `Selfie2Anime` variant.
```
saved_model_dir = tempfile.mkdtemp()
with tf.Graph().as_default(), tf.Session() as sess:
gan = UGATIT(sess, data)
gan.build_model()
load_checkpoint(sess, '/content/checkpoint/UGATIT_light_selfie2anime_lsgan_4resblock_6dis_1_1_10_10_1000_sn_smoothing')
# Write SavedModel for serving or conversion to TF Lite
tf.saved_model.simple_save(
sess,
saved_model_dir,
inputs={
gan.test_domain_A.name: gan.test_domain_A,
},
outputs={gan.test_fake_B.name: gan.test_fake_B})
tf.logging.debug('Export transform SavedModel to',
saved_model_dir)
```
Note the path of the `SavedModel` from the above logs. We will be needing this for the subsequent steps. The warnings can be ignored.
```
# Inspecting model size
print(os.path.getsize(os.path.join(saved_model_dir, 'saved_model.pb')))
```
## TF Lite conversion
```
!pip install -q tensorflow==2.2.0
```
**Important**: Restart the runtime by selecting menu item, Runtime > Restart runtime.
```
import tensorflow as tf
print(tf.__version__)
def convert_to_tflite(saved_model_path, tflite_model_path):
model = tf.saved_model.load(saved_model_path)
concrete_func = model.signatures[
tf.saved_model.DEFAULT_SERVING_SIGNATURE_DEF_KEY]
concrete_func.inputs[0].set_shape([1, 256, 256, 3])
concrete_func.outputs[0].set_shape([1, 256, 256, 3])
converter = tf.lite.TFLiteConverter.from_concrete_functions([concrete_func])
converter.optimizations = [tf.lite.Optimize.OPTIMIZE_FOR_SIZE]
tflite_model = converter.convert()
with tf.io.gfile.GFile(tflite_model_path, 'wb') as f:
f.write(tflite_model)
print('Fixed-point Quantized model:', tflite_model_path,
'Size:', len(tflite_model) / 1024, "kb")
convert_to_tflite('/tmp/tmp22_x9l4i/', 'selfie2anime.tflite') # Note that the path might change since we are using `tempfile`
```
Warnings can be ignored here.
## Running inference with the TF Lite model
### Gather an example image
```
!wget https://pbs.twimg.com/profile_images/1235595938921459713/h26CpAPb_400x400.jpg
import numpy as np
import matplotlib.pyplot as plt
```
### View the image
```
def load_image(path):
image_raw = tf.io.read_file(path)
image = tf.image.decode_image(image_raw, channels=3)
return image
test_image_original = load_image("h26CpAPb_400x400.jpg")
print(test_image_original.shape)
plt.imshow(test_image_original)
plt.show()
```
### Preprocess the image
```
def resize(image):
resized_image = tf.image.resize(image, [256, 256], method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
resized_image = tf.cast(resized_image, tf.float32)
resized_image = tf.expand_dims(resized_image, 0)
return resized_image
test_image_resized = resize(test_image_original)
test_image_resized.shape
```
### Run inference on the preprocessed image
```
with tf.io.gfile.GFile('selfie2anime.tflite', 'rb') as f:
model_content = f.read()
# Initialze TensorFlow Lite inpterpreter.
interpreter = tf.lite.Interpreter(model_content=model_content)
interpreter.allocate_tensors()
input_index = interpreter.get_input_details()[0]['index']
output = interpreter.tensor(interpreter.get_output_details()[0]["index"])
# Set model input
interpreter.set_tensor(input_index, test_image_resized)
# Run inference
interpreter.invoke()
# Visualize results
plt.subplot(121)
plt.title('Selfie')
plt.imshow(test_image_original)
plt.subplot(122)
plt.title('Anime')
plt.imshow(output()[0])
```
| github_jupyter |
```
import keras
from __future__ import absolute_import
from __future__ import print_function
import numpy as np
import random
from keras.datasets import mnist
from keras.models import Model
from keras.layers import Input, Flatten, Dense, Dropout,Flatten, Lambda
from keras.layers import Conv2D, Activation,AveragePooling2D,MaxPooling2D
from keras.optimizers import RMSprop
from keras import backend as K
num_classes=10
epochs=20
def euclid_dis(vects):
x,y = vects
sum_square = K.sum(K.square(x-y), axis=1, keepdims=True)
return K.sqrt(K.maximum(sum_square, K.epsilon()))
def eucl_dist_output_shape(shapes):
shape1, shape2 = shapes
return (shape1[0], 1)
def contrastive_loss(y_true, y_pred):
margin = 1
square_pred = K.square(y_pred)
margin_square = K.square(K.maximum(margin - y_pred, 0))
return K.mean(y_true * square_pred + (1 - y_true) * margin_square)
def create_pairs(x, digit_indices):
pairs = []
labels = []
n=min([len(digit_indices[d]) for d in range(num_classes)]) -1
for d in range(num_classes):
for i in range(n):
z1, z2 = digit_indices[d][i], digit_indices[d][i+1]
pairs += [[x[z1], x[z2]]]
inc = random.randrange(1, num_classes)
dn = (d + inc) % num_classes
z1, z2 = digit_indices[d][i], digit_indices[dn][i]
pairs += [[x[z1], x[z2]]]
labels += [1,0]
return np.array(pairs), np.array(labels)
def create_base_net(input_shape):
input = Input(shape = input_shape)
x = Conv2D(32, (3,3), activation = 'relu')(input)
x = AveragePooling2D(pool_size = (2,2))(x)
x = Conv2D(64, (3,3), activation = 'tanh')(x)
x = MaxPooling2D(pool_size = (2,2))(x)
x = Dropout(0.25)(x)
x = Flatten()(x)
x = Dense(128, activation = 'tanh')(x)
x = Dropout(0.5)(x)
x = Dense(64,activation = 'tanh')(x)
x = Dropout(0.5)(x)
x = Dense(10,activation = 'tanh')(x)
model = Model(input, x)
model.summary()
return model
def compute_accuracy(y_true, y_pred):
'''Compute classification accuracy with a fixed threshold on distances.
'''
pred = y_pred.ravel() < 0.5
return np.mean(pred == y_true)
def accuracy(y_true, y_pred):
'''Compute classification accuracy with a fixed threshold on distances.
'''
return K.mean(K.equal(y_true, K.cast(y_pred < 0.5, y_true.dtype)))
# the data, split between train and test sets
(x_train, y_train), (x_test, y_test) = mnist.load_data()
x_train = x_train.reshape(x_train.shape[0], 28, 28,1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
# input_shape = (1, 28, 28)
print(x_train.shape)
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
x_train /= 255
x_test /= 255
input_shape = x_train.shape[1:]
input_shape = (28, 28, 1)
# create training+test positive and negative pairs
digit_indices = [np.where(y_train == i)[0] for i in range(num_classes)]
tr_pairs, tr_y = create_pairs(x_train, digit_indices)
digit_indices = [np.where(y_test == i)[0] for i in range(num_classes)]
te_pairs, te_y = create_pairs(x_test, digit_indices)
# network definition
base_network = create_base_net(input_shape)
input_a = Input(shape=input_shape)
input_b = Input(shape=input_shape)
processed_a = base_network(input_a)
processed_b = base_network(input_b)
distance = Lambda(euclid_dis,
output_shape=eucl_dist_output_shape)([processed_a, processed_b])
model = Model([input_a, input_b], distance)
rms = RMSprop()
model.compile(loss=contrastive_loss, optimizer=rms, metrics=[accuracy])
model.fit([tr_pairs[:, 0], tr_pairs[:, 1]], tr_y,
batch_size=128,
epochs=epochs,
validation_data=([te_pairs[:, 0], te_pairs[:, 1]], te_y))
# compute final accuracy on training and test sets
y_pred = model.predict([tr_pairs[:, 0], tr_pairs[:, 1]])
tr_acc = compute_accuracy(tr_y, y_pred)
y_pred = model.predict([te_pairs[:, 0], te_pairs[:, 1]])
te_acc = compute_accuracy(te_y, y_pred)
print('* Accuracy on training set: %0.2f%%' % (100 * tr_acc))
print('* Accuracy on test set: %0.2f%%' % (100 * te_acc))
import matplotlib.pyplot as plt
from PIL import Image
number_of_items = 20
import tensorflow as tf
im = tf.keras.preprocessing.image.array_to_img(
tr_pairs[1,0],
data_format=None,
scale=True,
dtype=None
)
plt.figure(figsize=(10, 5))
for item in range(number_of_items):
display = plt.subplot(1, number_of_items,item+1)
im = tf.keras.preprocessing.image.array_to_img( tr_pairs[item,0], data_format=None, scale=True,dtype=None)
plt.imshow(im, cmap="gray")
display.get_xaxis().set_visible(False)
display.get_yaxis().set_visible(False)
plt.show()
plt.figure(figsize=(10, 5))
for item in range(number_of_items):
display = plt.subplot(1, number_of_items,item+1)
im = tf.keras.preprocessing.image.array_to_img( tr_pairs[item,1], data_format=None, scale=True,dtype=None)
plt.imshow(im, cmap="gray")
display.get_xaxis().set_visible(False)
display.get_yaxis().set_visible(False)
plt.show()
for i in range(number_of_items):
print(y_pred[i])
plt.figure(figsize=(20, 2))
plt.imshow(im, cmap="gray")
plt.show()
```
| github_jupyter |
# Cox model
```
import arviz as az
import numpy as np
import pymc3 as pm
import scipy as sp
import theano.tensor as tt
from pymc3 import (NUTS, Gamma, Metropolis, Model, Normal, Poisson, find_MAP,
forestplot, sample, starting, traceplot)
from theano import function as fn
from theano import printing
print('Running on PyMC3 v{}'.format(pm.__version__))
%config InlineBackend.figure_format = 'retina'
az.style.use('arviz-darkgrid')
```
Here is the original model, implemented in BUGS:
```R
model
{
# Set up data
for(i in 1:Nsubj) {
for(j in 1:T) {
# risk set = 1 if obs.t >= t
Y[i,j] <- step(obs.t[i] - t[j] + eps)
# counting process jump = 1 if obs.t in [ t[j], t[j+1] )
# i.e. if t[j] <= obs.t < t[j+1]
dN[i, j] <- Y[i, j] * step(t[j + 1] - obs.t[i] - eps) * FAIL[i]
}
}
# Model
for(j in 1:T) {
for(i in 1:Nsubj) {
dN[i, j] ~ dpois(Idt[i, j]) # Likelihood
Idt[i, j] <- Y[i, j] * exp(beta[1]*pscenter[i] + beta[2]*
hhcenter[i] + beta[3]*ncomact[i] + beta[4]*rleader[i] + beta[5]*dleader[i] + beta[6]*inter1[i] + beta[7]*inter2[i]) * dL0[j] # Intensity
}
dL0[j] ~ dgamma(mu[j], c)
mu[j] <- dL0.star[j] * c # prior mean hazard
}
c ~ dgamma(0.0001, 0.00001)
r ~ dgamma(0.001, 0.0001)
for (j in 1 : T) { dL0.star[j] <- r * (t[j + 1] - t[j]) }
# next line indicates number of covariates and is for the corresponding betas
for(i in 1:7) {beta[i] ~ dnorm(0.0,0.00001)}
}
```
```
dta = dict(T=73, Nsubj=430, eps=0.0, t=[1, 21, 85, 128, 129, 148, 178, 204,
206, 210, 211, 212, 225, 238, 241,
248, 259, 273, 275, 281, 286, 289,
301, 302, 303, 304, 313, 317, 323,
344, 345, 349, 350, 351, 355, 356,
359, 364, 385, 386, 389, 390, 391,
392, 394, 395, 396, 397, 398, 399,
400, 406, 415, 416, 426, 427, 434,
435, 437, 441, 447, 448, 449, 450,
451, 453, 455, 456, 458, 459, 460,
461, 462, 463],
obs_t = [460, 313, 435, 350, 435, 350, 350, 460, 460, 448, 225, 225, 396, 435, 396, 396, 453, 396, 456, 397, 397, 396, 395, 275, 449, 395, 395, 462, 302, 302, 458, 461, 396, 241, 389, 458, 304, 304, 395, 395, 364, 460, 415, 463, 396, 459, 441, 435, 396, 458, 437, 396, 356, 356, 396, 455, 396, 462, 399, 400, 350, 350, 395, 395, 441, 355, 85, 458, 128, 396, 386, 386, 386, 462, 458, 390, 390, 396, 396, 396, 427, 458, 395, 275, 275, 395, 359, 395, 395, 441, 395, 463, 178, 275, 463, 396, 396, 259, 396, 396, 458, 441, 396, 463, 396, 463, 435, 396, 437, 396, 398, 463, 460, 462, 460, 460, 210, 396, 435, 458, 385, 323, 323, 359, 396, 396, 460, 238, 441, 450, 392, 458, 396, 458, 396, 396, 462, 435, 396, 394, 396, 435, 458, 1, 395, 395, 451, 462, 458, 462, 396, 286, 396, 349, 449, 462, 455, 21, 463, 461, 461, 456, 435, 396, 460, 462, 462, 435, 435, 460, 386, 396, 458, 386, 461, 441, 435, 435, 463, 456, 396, 275, 460, 406, 460, 406, 317, 406, 461, 396, 359, 458, 463, 435, 462, 458, 396, 396, 273, 396, 435, 281, 275, 396, 447, 225, 447, 396, 435, 416, 396, 248, 396, 435, 435, 396, 461, 385, 396, 458, 458, 396, 461, 396, 448, 396, 396, 460, 455, 456, 463, 462, 458, 463, 396, 462, 395, 456, 396, 463, 396, 435, 459, 396, 396, 396, 395, 435, 455, 395, 461, 344, 396, 395, 396, 317, 396, 395, 426, 461, 396, 289, 441, 395, 396, 458, 396, 396, 435, 396, 395, 396, 441, 345, 396, 359, 435, 435, 396, 396, 395, 458, 461, 458, 212, 301, 458, 456, 395, 396, 395, 435, 396, 396, 303, 458, 460, 400, 396, 462, 359, 458, 396, 206, 441, 396, 458, 396, 462, 396, 396, 275, 396, 395, 435, 435, 462, 225, 458, 462, 396, 396, 289, 396, 303, 455, 400, 400, 359, 461, 396, 462, 460, 463, 463, 463, 204, 435, 435, 396, 396, 396, 463, 458, 396, 455, 435, 396, 396, 463, 396, 461, 463, 460, 441, 460, 435, 435, 460, 455, 460, 395, 460, 460, 460, 435, 449, 463, 462, 129, 391, 396, 391, 391, 434, 356, 462, 396, 349, 225, 396, 435, 461, 391, 391, 351, 211, 461, 212, 434, 148, 356, 458, 456, 455, 435, 463, 463, 462, 435, 463, 437, 460, 396, 406, 451, 460, 435, 396, 460, 455, 396, 398, 456, 458, 396, 456, 449, 396, 128, 396, 462, 463, 396, 396, 396, 435, 460, 396, 458],
FAIL= [1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1],
pscenter= [
-.01434325, -.01460965, .01322687, .00971885, -.03223412, -.01113493, -.01359567, -.03357866, -.0387039, -.0553269, -.03238896, -.07464545, -.07325128, -.07062459, -.07464545, -.07032613, -.0703005, .00965232, -.01408955, .00577483, -.00219072, -.00084567, .01643198, .06509522, .06824313, .07300876, .07300876, .01394272, .06824313, .02063087, .00383186, -.02573045, -.02410864, -.02272752, .05120398, -.00997729, -.00550709, -.02062663, -.03077685, -.01688493, .01035959, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .01149963, .0034338, .0376236, .00733331, .01520069, .03832785, .03832785, -.02622275, -.02622275, -.02622275, -.01492678, -.02897806, -.02897806, -.02897806, -.02847666, -.031893, -.03919478, -.04224754, -.04743705, -.0510477, -.031893, -.01129093, .01706207, .00193999, -.01503116, .003101, -.00083466, .02395027, -.07952866, -.08559135, -.07251801, -.06586029, -.08432532, -.0613939, -.081205, -.07540084, -.08488011, -.08488011, -.08488011, -.07492433, -.08907269, -.09451609, -.05301854, -.08980743, -.0771635, -.0771635, -.08650947, -.07856082, -.0771635, -.08204606, -.08178245, -.05263504, -.05355574, -.05109092, -.04696729, -.04696729, -.04696729, -.05257489, -.05303248, -.05348096, -.04983674, -.04699414, .00584956, -.00792241, -.01719816, -.02138029, -.01576016, -.04274812, -.04014061, .0471441, .0471441, .0471441, .0471441, .0471441, .0471441, .0471441, .04233112, .0471441, .04233112, .050568, .07388823, .0493324, .04512087, .03205975, .02913185, .06010427, .05324252, .06973204, .05579907, .01212243, .07962459, .05054695, .06672142, .14026688, .01734403, .06078221, .06543709, .06438115, .20126908, -.03138622, -.02180659, .01637333, -.02415774, .01828684, .03106104, .04268495, .01897239, .01591935, -.02367065, -.0619156, -.06403028, -.06851645, -.04821694, -.03889525, -.05023452, -.05013452, -.01557191, -.01171948, -.01362136, -.01174715, -.02707938, -.02634164, -.02634164, -.02634164, -.00692153, -.02381614, -.00890537, -.00611669, -.00894752, -.03551984, -.0252678, -.01513384, -.01016569, -.03551984, -.03773227, -.01978032, .06803483, .06706496, .10551275, .15091534, .03092981, .06556855, .10781559, .12671031, .0936299, .09362991, .09362991, .08294538, .09362991, .09362991, .09362991, .01177025, .02610553, .03546937, .03546937, .03546937, .034415, -.00305626, .04973665, .05103208, .07546701, .05306436, .00824125, .01961115, .01202359, -.02919447, -.01016712, .01756074, -.04035511, -.04753104, -.04463152, -.04845615, -.05010044, .00031411, -.07911871, -.08799869, -.07980882, -.09393142, -.08000018, -.07666632, -.07817401, -.07444922, -.07226554, -.08216553, -.0777643, -.07752042, -.05767992, -.04727952, -.03774814, -.06870384, -.05999847, -.05947695, .02989959, .04627543, .02772475, .02883079, .03642944, .02871235, .04148949, .04240279, .07747082, .07626323, .04268012, .03225577, .06468724, -.05140995, -.05399637, -.05351515, .07302427, .02432223, .0490674, .0490674, .0490674, .0490674, .09013112, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .10476315, .07008056, .08666077, .01546215, .01667466, .03417671, .05253941, .04293926, .01496588, .02692172, -.03827151, .04809769, .08742411, .04533176, .01455173, .01831875, .02710811, .09834951, .09952456, .06993483, .02945534, .038731, .1181948, .04435538, .04435538, -.02357505, .05824019, .05820741, -.02357505, .09324722, .15534712, .07207468, .04692869, -.03490683, -.04404809, -.05054474, -.05325826, -.0474724, -.04905931, .01068221, .02879751, .00852646, .02693032, .01835589, .02989959, .02989959, .02989959, .04976377, .04439012, .03397319, .02989959, .02989959, .05468828, .04463226, .05886378, .06311052, .02989959, .04595331, .04203459, .01231324, -.01399783, .04595331, .00145386, .04601278, .06459354, -.0007196, .00012216, -.07614055, -.08435525, -.07957162, -.10299519, -.08156988, -.08225659, -.07449063, -.00210284, -.00797183, -.025355, -.01258251, -.04372031, -.03985972, -.03545086, -.03384566, -.04025533, -.07523724, -.05947702, -.061286, -.07666647, -.07663169, -.05902354, -.07652324, -.07645561, -.06258684, -.09604834, -.08813326, -.03292062, -.07848112, -.08239502, -.08316891, -.07244316, -.075417, -.07652324, -.07922532, -.08755959, -.08583414, -.07450142, -.08066016, -.06057205, -.07652324, -.06249051, -.08781742, -.086076, -.07652324, -.07696518, -.0618688, -.06073988, -.06524737, -.04419825, -.04489509, -.04390368, -.04358438, -.04489509, -.04520512, -.04187583, -.03653955, -.03973426, -.03753508, -.03569439, -.06789339, .06689456, .05526327, .05139003, .02641841, .04891529, .07078697, .06862645, .06832582, .04104258, -.00120631, .01947345, .04891779, .04891779, .03561932, .02576244, .03158225, .03608047, .08685057, .04632537, .06841581, -.02899643],
hhcenter= [ -.78348798, -.63418788, -.91218799, -.98388809, -.23518796, .11481193, -1.415588, -1.2535881, -.55738801, -.88128799, -1.109488, .05721192, -1.045788, -.30888793, .29651192, -.36688802, -.50058788, .02271203, -.59088796, -.04198809, .50561196, -.07418796, .98481184, .78921205, .09431199, -.06488796, 2.1662121, .08891205, 1.4004121, 1.316112, 1.9362121, 2.0107121, 1.150712, .31951192, -.23918791, -.1562881, -.9575879, -.07728811, .29641202, 1.2273121, 1.7717118, 1.5764117, .14181189, .72131211, 1.279212, .68241197, -.72808808, -.00488802, -.23938794, -1.000788, .55081207, -.52348799, 1.780612, -.35888812, .36481193, 1.5480118, -.03078791, 1.389112, .30211189, .70901209, -.16668792, 1.435812, .47001198, 2.0838118, 1.1673121, .18461208, -.30608794, 1.4470119, .23301201, -.58458799, .44011191, -.61948794, -.41388795, .263212, .66171199, .92451197, .78081208, .90991193, 1.6920118, 1.334012, 1.2101121, .41591194, -.48498794, -.73278803, -1.093588, .09911207, -.93418807, -.46908805, .0205119, .0535119, -.14228792, -.55708808, -.45498797, -.54008788, -.30998799, -.10958811, -.0960879, -.01338812, -.88168806, -.51788801, .36801198, .46621206, .13271193, -.11208793, -.76768798, -.54508799, -1.2773881, .16641192, .95871216, -.48238799, 1.6281118, -.18848796, -.49718806, -.41348812, -.31628796, -.59528798, -.11718794, -.57058805, -.59488791, -.21248789, -.65658802, -.56298798, -.52698797, -.65758795, -.04988809, .55341202, -.76328796, .254612, 1.3500118, -.54958791, 1.665812, .14671211, 1.963912, .29161194, -.56838793, 1.9371119, .90991193, -.39558789, .39521196, -.55208796, -.05268808, -.77368802, -.45428798, .05841212, -.45308802, -.12458798, .01431207, -.28228804, .79281193, -.26358792, -.54738802, -.38158795, -.54118794, -.72828788, -.58128804, .355912, -.24078794, -1.0384881, -.75038809, -.41018793, -.43538806, -1.566388, -.53388804, -.28388807, -1.2348881, -.69028801, -1.620088, -.78128809, -.54648799, -.92738789, .11871199, .26851204, .61571199, .82891208, 1.1985121, 1.012012, 1.0602121, -.02988811, .79301196, .67731196, .43991187, .9404121, .5254119, 1.0365119, 1.6220121, .61671191, -.50318807, 2.6073117, .02361206, -.60438794, -.79278797, -.18108793, -.48178813, -.44038793, -.22628804, -.07398792, .519512, .40211204, .582012, 1.830512, .80441195, .58801204, -.56368798, -1.5451881, .45991209, -.23448797, -.36918804, 1.3247118, .19541197, -.20818801, 1.163012, -.78228801, -.6048879, -.575288, 1.3241119, .0147119, -.76518792, -.37478802, -.35508797, -.90038794, -1.250888, -.46608803, -.98488802, -1.5185881, -.90908808, -1.048188, -.90138787, -.77278799, -1.248988, -.34448811, -.61628789, .38531187, -.51728791, -.00878807, -.60078806, -.45358798, .46301201, -.22048803, -.71518797, -.76478809, -.75028795, -.4952881, .01731209, -.83718795, .57951194, .54291207, .45341209, .16941194, 1.054112, .61721212, 2.2717118, 1.1593118, 2.0280118, .92281204, 1.0100121, -.1866879, 2.6503119, 2.3914118, -.19948788, -.36418793, -.9259879, -.71058792, -.1104879, .16971211, 1.474812, 1.9360118, 2.5344119, 2.0171118, 1.9387121, .55071193, -.03918811, .20681195, .40421203, -.75518793, -.45678803, -1.0271881, .77211195, 1.146812, -1.147788, -1.565588, -.34888789, 1.303812, 1.952312, 1.639112, .07731203, .25901201, -.45608804, -.5028879, .03641204, -.03808804, .38571194, .31831196, -.17648788, -.44528791, -.55918807, -.53108805, .39721206, -.06328794, -.34038803, -.05988808, -.89548796, -.03518792, .045512, -.1859879, -.039288, -.82568806, .01431207, .40091208, -.2531881, .030412, -.31918809, -.54958791, -.79078788, .36691192, -.324388, -1.0082881, -1.232188, -.53248805, -.23678799, -.89188808, .25111201, -.6766879, -.3565881, -.61228794, -.21078797, -1.0343881, -.58358806, -.15588804, -.39238808, -.67818803, -.19498797, 1.099412, 1.2767119, -.64068788, -.50678796, -.64058799, -.86918801, 1.4048119, -.59648794, .23331194, .68371207, .11251191, -.17128797, .17081194, -.44218799, -.48708794, .09591202, .20131211, -.20108791, -.02158805, -.48188803, -.3012881, -.55008787, -1.146188, -.82128805, -.87638801, -.54488796, -.60288805, -1.003088, -.25078794, -.14818807, -.14738794, -.80938786, -.85988802, -.90188807, -.94998807, -.75718802, -.37418792, -.66708797, 1.0981121, 1.1441121, .47381189, -.12958808, -.34358808, -.84328789, -.33498809, -.98088807, -.6903879, -1.284988, -.80838794, -.91838807, -.81848806, -.34488794, -.83438796, .12971191, .99381214, -.91608804, -.31808802, -.01018806, .98171192, -.91638798, -1.043988, -1.0103881, 1.451612, -.01528808, .02441196, -.41458794, .25691202, .18601207, -.815988, -.02908798, -.59088796, -.35608789, .79691201, 1.8123121, -.98588794, 1.548912, 2.3653121, -.09238812, .96741205, .05891208, -.15618797, -.5660879, -.28338811, -.10088798, 1.1663117, .21981196, .07151202, -.009088, -.49578807, .15441208, -.44488809, -.2677879, -.54388803, -.25468799, .68631202, -.88128799, -.84628791, -1.2549881, -.36198804],
ncomact= [ 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 1, 1, 1, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 0, 1, 1, 1, 0, 0, 0, 0, 0, 1, 0, 1, 1, 0, 1, 0, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 1, 1, 1, 1, 0, 1, 1, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 0, 0, 0, 1, 0, 0, 1, 1, 0, 1, 0, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 1, 0, 0, 0, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 1, 0, 0, 1, 1, 0, 1, 1, 0, 1, 0, 1, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 1, 0, 1, 0, 1, 0, 0, 0, 1, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 1, 0, 1, 1, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 1, 0, 1, 0, 0, 0, 1, 1, 1, 1, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 1, 0, 1, 0, 0, 0, 1, 0, 1, 0, 1, 0, 1, 1, 0, 0, 1, 0, 0, 0, 0, 1],
rleader= [ 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
dleader= [ 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 1, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 1, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0, 0],
inter1= [ -.01434325, -.01460965, 0, 0, 0, -.01113493, 0, 0, 0, -.0553269, -.03238896, 0, 0, -.07062459, -.07464545, -.07032613, 0, 0, -.01408955, 0, -.00219072, 0, 0, 0, 0, 0, .07300876, .01394272, 0, 0, 0, 0, 0, 0, .05120398, 0, -.00550709, -.02062663, -.03077685, -.01688493, 0, .01149963, 0, .01149963, .01149963, 0, 0, 0, 0, 0, 0, 0, 0, 0, .01149963, .0034338, .0376236, .00733331, 0, .03832785, .03832785, -.02622275, -.02622275, -.02622275, -.01492678, 0, 0, -.02897806, -.02847666, 0, 0, -.04224754, -.04743705, -.0510477, -.031893, 0, 0, 0, -.01503116, .003101, -.00083466, .02395027, -.07952866, 0, 0, -.06586029, 0, -.0613939, -.081205, -.07540084, -.08488011, -.08488011, 0, -.07492433, -.08907269, -.09451609, 0, -.08980743, 0, -.0771635, 0, 0, -.0771635, -.08204606, 0, -.05263504, 0, -.05109092, -.04696729, 0, -.04696729, 0, -.05303248, -.05348096, 0, 0, .00584956, -.00792241, -.01719816, 0, -.01576016, 0, -.04014061, 0, 0, 0, 0, 0, .0471441, 0, .04233112, 0, .04233112, 0, 0, .0493324, .04512087, .03205975, .02913185, 0, .05324252, 0, 0, 0, 0, .05054695, 0, .14026688, .01734403, .06078221, 0, 0, 0, -.03138622, 0, .01637333, 0, 0, 0, 0, .01897239, .01591935, 0, -.0619156, 0, -.06851645, 0, -.03889525, -.05023452, -.05013452, 0, 0, -.01362136, 0, 0, -.02634164, 0, 0, 0, 0, -.00890537, -.00611669, 0, 0, 0, -.01513384, 0, -.03551984, 0, -.01978032, 0, .06706496, .10551275, 0, .03092981, .06556855, 0, 0, 0, .09362991, 0, 0, 0, 0, 0, 0, .02610553, .03546937, 0, 0, .034415, 0, 0, 0, .07546701, 0, 0, 0, 0, -.02919447, -.01016712, 0, 0, 0, 0, -.04845615, -.05010044, 0, 0, 0, 0, 0, 0, -.07666632, 0, 0, -.07226554, -.08216553, -.0777643, 0, 0, -.04727952, 0, -.06870384, -.05999847, 0, 0, 0, .02772475, .02883079, .03642944, 0, .04148949, 0, 0, 0, .04268012, .03225577, 0, -.05140995, -.05399637, 0, 0, .02432223, 0, .0490674, .0490674, .0490674, 0, 0, 0, 0, 0, 0, 0, 0, .10476315, 0, 0, 0, 0, 0, .07008056, 0, 0, .01667466, 0, .05253941, .04293926, 0, .02692172, 0, 0, .08742411, .04533176, 0, .01831875, 0, .09834951, .09952456, 0, .02945534, .038731, 0, .04435538, 0, -.02357505, 0, 0, -.02357505, .09324722, 0, 0, 0, -.03490683, 0, -.05054474, 0, -.0474724, -.04905931, 0, .02879751, 0, 0, 0, 0, 0, 0, 0, .04439012, 0, .02989959, .02989959, .05468828, .04463226, 0, 0, 0, 0, 0, .01231324, -.01399783, .04595331, .00145386, 0, .06459354, -.0007196, 0, -.07614055, -.08435525, 0, -.10299519, 0, 0, 0, -.00210284, -.00797183, 0, 0, 0, 0, -.03545086, 0, 0, 0, 0, -.061286, -.07666647, 0, -.05902354, -.07652324, -.07645561, 0, 0, 0, -.03292062, 0, 0, 0, 0, -.075417, 0, -.07922532, 0, -.08583414, -.07450142, -.08066016, 0, 0, -.06249051, 0, 0, 0, 0, -.0618688, 0, -.06524737, -.04419825, -.04489509, 0, 0, 0, -.04520512, -.04187583, 0, 0, -.03753508, 0, 0, 0, 0, 0, 0, 0, 0, .06862645, 0, 0, -.00120631, .01947345, 0, 0, .03561932, 0, .03158225, .03608047, 0, 0, 0, -.02899643],
inter2= [-.78348798, -.63418788, 0, 0, 0, .11481193, 0, 0, 0, -.88128799, -1.109488, 0, 0, -.30888793, .29651192, -.36688802, 0, 0, -.59088796, 0, .50561196, 0, 0, 0, 0, 0, 2.1662121, .08891205, 0, 0, 0, 0, 0, 0, -.23918791, 0, -.9575879, -.07728811, .29641202, 1.2273121, 0, 1.5764117, 0, .72131211, 1.279212, 0, 0, 0, 0, 0, 0, 0, 0, 0, .36481193, 1.5480118, -.03078791, 1.389112, 0, .70901209, -.16668792, 1.435812, .47001198, 2.0838118, 1.1673121, 0, 0, 1.4470119, .23301201, 0, 0, -.61948794, -.41388795, .263212, .66171199, 0, 0, 0, 1.6920118, 1.334012, 1.2101121, .41591194, -.48498794, 0, 0, .09911207, 0, -.46908805, .0205119, .0535119, -.14228792, -.55708808, 0, -.54008788, -.30998799, -.10958811, 0, -.01338812, 0, -.51788801, 0, 0, .13271193, -.11208793, 0, -.54508799, 0, .16641192, .95871216, 0, 1.6281118, 0, -.49718806, -.41348812, 0, 0, -.11718794, -.57058805, -.59488791, 0, -.65658802, 0, -.52698797, 0, 0, 0, 0, 0, 1.3500118, 0, 1.665812, 0, 1.963912, 0, 0, 1.9371119, .90991193, -.39558789, .39521196, 0, -.05268808, 0, 0, 0, 0, -.12458798, 0, -.28228804, .79281193, -.26358792, 0, 0, 0, -.72828788, 0, .355912, 0, 0, 0, 0, -.43538806, -1.566388, 0, -.28388807, 0, -.69028801, 0, -.78128809, -.54648799, -.92738789, 0, 0, .61571199, 0, 0, 1.012012, 0, 0, 0, 0, .43991187, .9404121, 0, 0, 0, .61671191, 0, 2.6073117, 0, -.60438794, 0, -.18108793, -.48178813, 0, -.22628804, -.07398792, 0, 0, 0, 1.830512, 0, 0, 0, 0, 0, 0, -.36918804, 1.3247118, 0, 0, 1.163012, 0, 0, 0, 1.3241119, 0, 0, 0, 0, -.90038794, -1.250888, 0, 0, 0, 0, -1.048188, -.90138787, 0, 0, 0, 0, 0, 0, -.00878807, 0, 0, .46301201, -.22048803, -.71518797, 0, 0, -.4952881, 0, -.83718795, .57951194, 0, 0, 0, 1.054112, .61721212, 2.2717118, 0, 2.0280118, 0, 0, 0, 2.6503119, 2.3914118, 0, -.36418793, -.9259879, 0, 0, .16971211, 0, 1.9360118, 2.5344119, 2.0171118, 0, 0, 0, 0, 0, 0, 0, 0, .77211195, 0, 0, 0, 0, 0, 1.952312, 0, 0, .25901201, 0, -.5028879, .03641204, 0, .38571194, 0, 0, -.44528791, -.55918807, 0, .39721206, 0, -.34038803, -.05988808, 0, -.03518792, .045512, 0, -.039288, 0, .01431207, 0, 0, .030412, -.31918809, 0, 0, 0, -.324388, 0, -1.232188, 0, -.23678799, -.89188808, 0, -.6766879, 0, 0, 0, 0, 0, 0, 0, -.67818803, 0, 1.099412, 1.2767119, -.64068788, -.50678796, 0, 0, 0, 0, 0, .68371207, .11251191, -.17128797, .17081194, 0, -.48708794, .09591202, 0, -.20108791, -.02158805, 0, -.3012881, 0, 0, 0, -.87638801, -.54488796, 0, 0, 0, 0, -.14738794, 0, 0, 0, 0, -.75718802, -.37418792, 0, 1.0981121, 1.1441121, .47381189, 0, 0, 0, -.33498809, 0, 0, 0, 0, -.91838807, 0, -.34488794, 0, .12971191, .99381214, -.91608804, 0, 0, .98171192, 0, 0, 0, 0, -.01528808, 0, -.41458794, .25691202, .18601207, 0, 0, 0, -.35608789, .79691201, 0, 0, 1.548912, 0, 0, 0, 0, 0, 0, 0, 0, 1.1663117, 0, 0, -.009088, -.49578807, 0, 0, -.2677879, 0, -.25468799, .68631202, 0, 0, 0, -.36198804])
def load_data_cox(dta):
array = lambda x : np.array(dta[x], dtype=float)
t = array('t')
obs_t = array('obs_t')
pscenter = array('pscenter')
hhcenter = array('hhcenter')
ncomact = array('ncomact')
rleader = array('rleader')
dleader = array('dleader')
inter1 = array('inter1')
inter2 = array('inter2')
fail = array('FAIL')
return (t, obs_t, pscenter, hhcenter, ncomact,
rleader, dleader, inter1, inter2, fail)
(t, obs_t, pscenter, hhcenter, ncomact, rleader,
dleader, inter1, inter2, fail) = load_data_cox(dta)
X = np.array([pscenter, hhcenter, ncomact, rleader, dleader, inter1, inter2])
X.shape
with Model() as model:
T = len(t) - 1
nsubj = len(obs_t)
# risk set equals one if obs_t >= t
Y = np.array([[int(obs >= time) for time in t] for obs in obs_t])
# counting process. jump = 1 if obs_t \in [t[j], t[j+1])
dN = np.array([[Y[i,j]*int(t[j+1] >= obs_t[i])*fail[i] for j in range(T)] for i in
range(nsubj)])
c = Gamma('c', .0001, .00001)
r = Gamma('r', .001, .0001)
dL0_star = r*np.diff(t)
# prior mean hazard
mu = dL0_star * c
dL0 = Gamma('dL0', mu, c, shape=T)
beta = Normal('beta', np.zeros(7),
np.ones(7)*100, shape=7)
linear_model = tt.exp(tt.dot(X.T, beta))
idt = Y[:, :-1] * tt.outer(linear_model, dL0)
dn_like = Poisson('dn_like', idt, observed=dN)
with model:
trace = sample(2000, n_init=10000, init='advi_map')
traceplot(trace, var_names=['c', 'r']);
forestplot(trace, var_names=['beta']);
%load_ext watermark
%watermark -n -u -v -iv -w
```
| github_jupyter |
# Descriptor
#### This notebook showcases the functions used in descriptor analysis.
#### That is, determining the keypoints descriptor or unqiue identifier.
#### This descriptor is composed of the orientation histograms in local neighborhoods near the keypoint.
## Imports
```
# Handles relative import
import os, sys
dir2 = os.path.abspath('')
dir1 = os.path.dirname(dir2)
if not dir1 in sys.path: sys.path.append(dir1)
import cv2
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import const
import octaves as octaves_lib
import keypoints as keypoints_lib
import reference_orientation as reference_lib
import descriptor as descriptor_lib
```
## Find a Keypoint
```
img = cv2.imread('../images/box_in_scene.png', flags=cv2.IMREAD_GRAYSCALE)
img = cv2.normalize(img, None, alpha=0, beta=1, norm_type=cv2.NORM_MINMAX, dtype=cv2.CV_32F)
img = img[0:300, 100:400]
octave_idx = 4
gauss_octaves = octaves_lib.build_gaussian_octaves(img)
gauss_octave = gauss_octaves[octave_idx]
dog_octave = octaves_lib.build_dog_octave(gauss_octave)
extrema = octaves_lib.find_dog_extrema(dog_octave)
keypoint_coords = keypoints_lib.find_keypoints(extrema, dog_octave)
keypoints = reference_lib.assign_reference_orientations(keypoint_coords, gauss_octave, octave_idx)
keypoint = keypoints[0]
magnitudes, orientations = reference_lib.gradients(gauss_octave)
coord = keypoint.coordinate
sigma = keypoint.sigma
shape = gauss_octave.shape
s, y, x = coord.round().astype(int)
pixel_dist = octaves_lib.pixel_dist_in_octave(octave_idx)
max_width = (np.sqrt(2) * const.descriptor_locality * sigma) / pixel_dist
max_width = max_width.round().astype(int)
in_frame = descriptor_lib.patch_in_frame(coord, max_width, shape)
print(f'This keypoint is in frame: {in_frame}')
```
## Relative Coordinates
At this point, a keypoint has an orientation (see notebook 3).
This orientation becomes the local neighborhoods x axis.
In other words, there is a change of reference frame.
This is visualized here by showing each points relative x and y coordinate.
```
orientation_patch = orientations[s,
y - max_width: y + max_width,
x - max_width: x + max_width]
magnitude_patch = magnitudes[s,
y - max_width: y + max_width,
x - max_width: x + max_width]
patch_shape = magnitude_patch.shape
center_offset = [coord[1] - y, coord[2] - x]
rel_patch_coords = descriptor_lib.relative_patch_coordinates(center_offset, patch_shape, pixel_dist, sigma, keypoint.orientation)
plt.imshow(rel_patch_coords[1])
plt.title(f'rel X coords')
plt.colorbar()
plt.show()
plt.imshow(rel_patch_coords[0])
plt.title(f'rel Y coords')
plt.colorbar()
plt.show()
```
## Gaussian Weighting of Neighborhood
```
magnitude_patch = descriptor_lib.mask_outliers(magnitude_patch, rel_patch_coords, const.descriptor_locality)
orientation_patch = (orientation_patch - keypoint.orientation) % (2 * np.pi)
weights = descriptor_lib.weighting_matrix(center_offset, patch_shape, octave_idx, sigma, const.descriptor_locality)
plt.imshow(weights)
```
# Descriptor Patch
```
magnitude_patch = magnitude_patch * weights
plt.imshow(magnitude_patch)
```
# Descriptor Patch of Each Histogram
```
coords_rel_to_hists = rel_patch_coords[None] - descriptor_lib.histogram_centers[..., None, None]
hists_magnitude_patch = descriptor_lib.mask_outliers(magnitude_patch[None], coords_rel_to_hists, const.inter_hist_dist, 1)
nr_cols = 4
fig, axs = plt.subplots(nr_cols, nr_cols, figsize=(7, 7))
for idx, masked_magntiude in enumerate(hists_magnitude_patch):
row = idx // nr_cols
col = idx % nr_cols
axs[row, col].imshow(masked_magntiude)
axs[row, col].axis('off')
plt.tight_layout()
```
## Histograms to SIFT Feature
```
hists_magnitude_patch = descriptor_lib.interpolate_2d_grid_contribution(hists_magnitude_patch, coords_rel_to_hists)
hists = descriptor_lib.interpolate_1d_hist_contribution(hists_magnitude_patch, orientation_patch)
sift_feature = descriptor_lib.normalize_sift_feature(hists.ravel())
```
## Visualize Descriptor on Input Image
```
abs_coord = keypoint.absolute_coordinate[1:][::-1]
coord = keypoint.coordinate
sigma = keypoint.sigma
shape = gauss_octave.shape
s, y, x = coord.round().astype(int)
center_offset = [coord[1] - y, coord[2] - x]
pixel_dist = octaves_lib.pixel_dist_in_octave(octave_idx)
width = const.descriptor_locality * sigma
theta = keypoint.orientation
c, s = np.cos(theta), np.sin(theta)
rot_mat = np.array(((c, -s), (s, c)))
arrow = np.matmul(rot_mat, np.array([1, 0])) * 50
hist_centers = descriptor_lib.histogram_centers.T
hist_centers = hist_centers * sigma
hist_centers = np.matmul(rot_mat, hist_centers)
hist_centers = (hist_centers + abs_coord[:,None]).round().astype(int)
color = (1, 0, 0)
darkened = cv2.addWeighted(img, 0.5, np.zeros(img.shape, img.dtype),0,0)
col_img = cv2.cvtColor(darkened, cv2.COLOR_GRAY2RGB)
# Horizontal lines
for i in range(5):
offset = np.array([0, width/2]) * i
l = np.array([-width, -width]) + offset
r = np.array([width, -width]) + offset
l = (np.matmul(rot_mat, l) + abs_coord).round().astype(int)
r = (np.matmul(rot_mat, r) + abs_coord).round().astype(int)
col_img = cv2.line(col_img, l, r, color=color, thickness=1)
# Vertical lines
for i in range(5):
offset = np.array([width/2, 0]) * i
t = np.array([-width, -width]) + offset
b = np.array([-width, width]) + offset
t = (np.matmul(rot_mat, t) + abs_coord).round().astype(int)
b = (np.matmul(rot_mat, b) + abs_coord).round().astype(int)
col_img = cv2.line(col_img, t, b, color=color, thickness=1)
plt.figure(figsize=(8, 8))
plt.imshow(col_img)
plt.axis('off')
plt.title('red arrow is x axis relative to keypoint')
xs, ys = hist_centers
plt.scatter(xs, ys, c=[x for x in range(len(xs))], cmap='autumn_r')
plt.arrow(abs_coord[0], abs_coord[1], arrow[0], arrow[1], color='red', width=1, head_width=10)
plt.show()
print(f'The red arrow represtns a rotation of {np.rad2deg(keypoint.orientation)} degrees.')
```
# Histogram Content
```
cmap = matplotlib.cm.get_cmap('autumn_r')
fig, axs = plt.subplots(4, 4, figsize=(8, 8))
for idx, hist in enumerate(hists):
row = idx // 4
col = idx % 4
color = cmap((idx + 1) / len(hists))
axs[row, col].bar(list(range(const.nr_descriptor_bins)), hist, color=color)
plt.tight_layout()
plt.show()
```
## The SIFT feature a.k.a Concatenated Histograms
```
colors = [cmap((idx+1) / len(hists)) for idx in range(16)]
colors = np.repeat(colors, const.nr_descriptor_bins, axis=0)
plt.figure(figsize=(20, 4))
plt.bar(range(len(sift_feature)), sift_feature, color=colors)
```
| github_jupyter |
# Demo: How to scrape multiple things from multiple pages
The goal is to scrape info about the five top-grossing movies for each year, for 10 years. I want the title and rank of the movie, and also, how much money did it gross at the box office. In the end I will put the scraped data into a CSV file.
```
from bs4 import BeautifulSoup
import requests
url = 'https://www.boxofficemojo.com/yearly/chart/?yr=2018'
html = requests.get(url)
soup = BeautifulSoup(html.text, 'html.parser')
# I discover the data I want is in an HTML table with no class or ID
tables = soup.find_all( 'table' )
print(len(tables))
# I had to test a few numbers before I got the correct tables[] and rows[] numbers
# I just kept changing the number and printing until I found it
rows = tables[6].find_all('tr')
# print(len(rows))
# print(rows[2])
cells = rows[2].find_all('td')
title = cells[1].text
print(title)
# get top 5 movies on this page - I know the first row is [2]
for i in range(2, 7):
cells = rows[i].find_all('td')
title = cells[1].text
print(title)
# I would like to get the total gross number also
for i in range(2, 7):
cells = rows[i].find_all('td')
gross = cells[3].text
print(gross)
# next I want to get rank (1-5), title and gross all on one line
for i in range(2, 7):
cells = rows[i].find_all('td')
print(cells[0].text, cells[1].text, cells[3].text)
# I want to do this for 10 years, ending with 2018
# first create a list of the years I want
years = []
start = 2018
for i in range(0, 10):
years.append(start - i)
print(years)
# create a base url so I can open each year's page
base_url = 'https://www.boxofficemojo.com/yearly/chart/?yr='
# test it
# print(base_url + years[0]) -- ERROR
print( base_url + str(years[0]) )
# collect all necessary pieces from above to make a loop that gets top 5 movies
# for each of the 10 years
for year in years:
url = base_url + str(year)
html = requests.get(url)
soup = BeautifulSoup(html.text, 'html.parser')
tables = soup.find_all( 'table' )
rows = tables[6].find_all('tr')
for i in range(2, 7):
cells = rows[i].find_all('td')
print(cells[0].text, cells[1].text, cells[3].text)
# I realize now that each line needs to have the year also
# and maybe I should clean the gross so it's a pure integer
# so test that - using .strip() and .replace() chained together -
num = '$293,004,164'
print(num.strip('$').replace(',', ''))
miniyears = [2017, 2014]
for year in miniyears:
url = base_url + str(year)
html = requests.get(url)
soup = BeautifulSoup(html.text, 'html.parser')
tables = soup.find_all( 'table' )
rows = tables[6].find_all('tr')
for i in range(2, 7):
cells = rows[i].find_all('td')
gross = cells[3].text.strip('$').replace(',', '')
print(year, cells[0].text, cells[1].text, gross)
# I should really save my data into a csv
import csv
# open new file for writing -
csvfile = open("movies.csv", 'w', newline='', encoding='utf-8')
# make a new variable, c, for Python's CSV writer object -
c = csv.writer(csvfile)
#write header row to csv
c.writerow( ['year', 'rank', 'title', 'gross'] )
# modified code from above
for year in years:
url = base_url + str(year)
html = requests.get(url)
soup = BeautifulSoup(html.text, 'html.parser')
tables = soup.find_all( 'table' )
rows = tables[6].find_all('tr')
for i in range(2, 7):
cells = rows[i].find_all('td')
gross = cells[3].text.strip('$').replace(',', '')
# print(year, cells[0].text, cells[1].text, gross)
# instead of printing, I need to make a list and write that list to the CSV as one row
c.writerow( [year, cells[0].text, cells[1].text, gross] )
# close the file
csvfile.close()
```
The result is a CSV file, named movies.csv, that has 51 rows: the header row plus 5 movies for each year from 2009 through 2018. It has four columns: year, rank, title, and gross.
Note that **only the final cell above** is needed to create this CSV, by scraping 10 separate web pages. Everything *above* the final cell above is just instruction, demonstration. It is intended to show the problem-solving you need to go through to get to a desired scraping result.
| github_jupyter |
# DoWhy: Interpreters for Causal Estimators
This is a quick introduction to the use of interpreters in the DoWhy causal inference library.
We will load in a sample dataset, use different methods for estimating the causal effect of a (pre-specified)treatment variable on a (pre-specified) outcome variable and demonstrate how to interpret the obtained results.
First, let us add the required path for Python to find the DoWhy code and load all required packages
```
%load_ext autoreload
%autoreload 2
import numpy as np
import pandas as pd
import logging
import dowhy
from dowhy import CausalModel
import dowhy.datasets
```
Now, let us load a dataset. For simplicity, we simulate a dataset with linear relationships between common causes and treatment, and common causes and outcome.
Beta is the true causal effect.
```
data = dowhy.datasets.linear_dataset(beta=1,
num_common_causes=5,
num_instruments = 2,
num_treatments=1,
num_discrete_common_causes=1,
num_samples=10000,
treatment_is_binary=True,
outcome_is_binary=False)
df = data["df"]
print(df[df.v0==True].shape[0])
df
```
Note that we are using a pandas dataframe to load the data.
## Identifying the causal estimand
We now input a causal graph in the GML graph format.
```
# With graph
model=CausalModel(
data = df,
treatment=data["treatment_name"],
outcome=data["outcome_name"],
graph=data["gml_graph"],
instruments=data["instrument_names"]
)
model.view_model()
from IPython.display import Image, display
display(Image(filename="causal_model.png"))
```
We get a causal graph. Now identification and estimation is done.
```
identified_estimand = model.identify_effect(proceed_when_unidentifiable=True)
print(identified_estimand)
```
## Method 1: Propensity Score Stratification
We will be using propensity scores to stratify units in the data.
```
causal_estimate_strat = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_stratification",
target_units="att")
print(causal_estimate_strat)
print("Causal Estimate is " + str(causal_estimate_strat.value))
```
### Textual Interpreter
The textual Interpreter describes (in words) the effect of unit change in the treatment variable on the outcome variable.
```
# Textual Interpreter
interpretation = causal_estimate_strat.interpret(method_name="textual_effect_interpreter")
```
### Visual Interpreter
The visual interpreter plots the change in the standardized mean difference (SMD) before and after Propensity Score based adjustment of the dataset. The formula for SMD is given below.
$SMD = \frac{\bar X_{1} - \bar X_{2}}{\sqrt{(S_{1}^{2} + S_{2}^{2})/2}}$
Here, $\bar X_{1}$ and $\bar X_{2}$ are the sample mean for the treated and control groups.
```
# Visual Interpreter
interpretation = causal_estimate_strat.interpret(method_name="propensity_balance_interpreter")
```
This plot shows how the SMD decreases from the unadjusted to the stratified units.
## Method 2: Propensity Score Matching
We will be using propensity scores to match units in the data.
```
causal_estimate_match = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_matching",
target_units="atc")
print(causal_estimate_match)
print("Causal Estimate is " + str(causal_estimate_match.value))
# Textual Interpreter
interpretation = causal_estimate_match.interpret(method_name="textual_effect_interpreter")
```
Cannot use propensity balance interpretor here since the interpreter method only supports propensity score stratification estimator.
## Method 3: Weighting
We will be using (inverse) propensity scores to assign weights to units in the data. DoWhy supports a few different weighting schemes:
1. Vanilla Inverse Propensity Score weighting (IPS) (weighting_scheme="ips_weight")
2. Self-normalized IPS weighting (also known as the Hajek estimator) (weighting_scheme="ips_normalized_weight")
3. Stabilized IPS weighting (weighting_scheme = "ips_stabilized_weight")
```
causal_estimate_ipw = model.estimate_effect(identified_estimand,
method_name="backdoor.propensity_score_weighting",
target_units = "ate",
method_params={"weighting_scheme":"ips_weight"})
print(causal_estimate_ipw)
print("Causal Estimate is " + str(causal_estimate_ipw.value))
# Textual Interpreter
interpretation = causal_estimate_ipw.interpret(method_name="textual_effect_interpreter")
interpretation = causal_estimate_ipw.interpret(method_name="confounder_distribution_interpreter", fig_size=(8,8), font_size=12, var_name='W4', var_type='discrete')
```
| github_jupyter |
<center>
<img src="https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork/labs/Module%202/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
</center>
# Multiple Linear Regression
Estimated time needed: **15** minutes
## Objectives
After completing this lab you will be able to:
* Use scikit-learn to implement Multiple Linear Regression
* Create a model, train it, test it and use the model
<h1>Table of contents</h1>
<div class="alert alert-block alert-info" style="margin-top: 20px">
<ol>
<li><a href="https://#understanding-data">Understanding the Data</a></li>
<li><a href="https://#reading_data">Reading the Data in</a></li>
<li><a href="https://#multiple_regression_model">Multiple Regression Model</a></li>
<li><a href="https://#prediction">Prediction</a></li>
<li><a href="https://#practice">Practice</a></li>
</ol>
</div>
<br>
<hr>
### Importing Needed packages
```
import matplotlib.pyplot as plt
import pandas as pd
import pylab as pl
import numpy as np
%matplotlib inline
```
### Downloading Data
To download the data, we will use !wget to download it from IBM Object Storage.
```
!wget -O FuelConsumption.csv https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBMDeveloperSkillsNetwork-ML0101EN-SkillsNetwork/labs/Module%202/data/FuelConsumptionCo2.csv
```
**Did you know?** When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
<h2 id="understanding_data">Understanding the Data</h2>
### `FuelConsumptionCo2.csv`:
We have downloaded a fuel consumption dataset, **`FuelConsumptionCo2.csv`**, which contains model-specific fuel consumption ratings and estimated carbon dioxide emissions for new light-duty vehicles for retail sale in Canada. [Dataset source](http://open.canada.ca/data/en/dataset/98f1a129-f628-4ce4-b24d-6f16bf24dd64?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01)
* **MODELYEAR** e.g. 2014
* **MAKE** e.g. Acura
* **MODEL** e.g. ILX
* **VEHICLE CLASS** e.g. SUV
* **ENGINE SIZE** e.g. 4.7
* **CYLINDERS** e.g 6
* **TRANSMISSION** e.g. A6
* **FUELTYPE** e.g. z
* **FUEL CONSUMPTION in CITY(L/100 km)** e.g. 9.9
* **FUEL CONSUMPTION in HWY (L/100 km)** e.g. 8.9
* **FUEL CONSUMPTION COMB (L/100 km)** e.g. 9.2
* **CO2 EMISSIONS (g/km)** e.g. 182 --> low --> 0
<h2 id="reading_data">Reading the data in</h2>
```
df = pd.read_csv("FuelConsumptionCo2.csv")
# take a look at the dataset
df.head()
```
Let's select some features that we want to use for regression.
```
cdf = df[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY','FUELCONSUMPTION_COMB','CO2EMISSIONS']]
cdf.head(9)
```
Let's plot Emission values with respect to Engine size:
```
plt.scatter(cdf.ENGINESIZE, cdf.CO2EMISSIONS, color='blue')
plt.xlabel("Engine size")
plt.ylabel("Emission")
plt.show()
```
#### Creating train and test dataset
Train/Test Split involves splitting the dataset into training and testing sets respectively, which are mutually exclusive. After which, you train with the training set and test with the testing set.
This will provide a more accurate evaluation on out-of-sample accuracy because the testing dataset is not part of the dataset that have been used to train the model. Therefore, it gives us a better understanding of how well our model generalizes on new data.
We know the outcome of each data point in the testing dataset, making it great to test with! Since this data has not been used to train the model, the model has no knowledge of the outcome of these data points. So, in essence, it is truly an out-of-sample testing.
Let's split our dataset into train and test sets. Around 80% of the entire dataset will be used for training and 20% for testing. We create a mask to select random rows using the **np.random.rand()** function:
```
msk = np.random.rand(len(df)) < 0.8
train = cdf[msk]
test = cdf[~msk]
```
#### Train data distribution
```
plt.scatter(train.ENGINESIZE, train.CO2EMISSIONS, color='blue')
plt.xlabel("Engine size")
plt.ylabel("Emission")
plt.show()
```
<h2 id="multiple_regression_model">Multiple Regression Model</h2>
In reality, there are multiple variables that impact the co2emission. When more than one independent variable is present, the process is called multiple linear regression. An example of multiple linear regression is predicting co2emission using the features FUELCONSUMPTION_COMB, EngineSize and Cylinders of cars. The good thing here is that multiple linear regression model is the extension of the simple linear regression model.
```
from sklearn import linear_model
regr = linear_model.LinearRegression()
x = np.asanyarray(train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
y = np.asanyarray(train[['CO2EMISSIONS']])
regr.fit (x, y)
# The coefficients
print ('Coefficients: ', regr.coef_)
print ('Intercept: ',regr.intercept_)
```
As mentioned before, **Coefficient** and **Intercept** are the parameters of the fitted line.
Given that it is a multiple linear regression model with 3 parameters and that the parameters are the intercept and coefficients of the hyperplane, sklearn can estimate them from our data. Scikit-learn uses plain Ordinary Least Squares method to solve this problem.
#### Ordinary Least Squares (OLS)
OLS is a method for estimating the unknown parameters in a linear regression model. OLS chooses the parameters of a linear function of a set of explanatory variables by minimizing the sum of the squares of the differences between the target dependent variable and those predicted by the linear function. In other words, it tries to minimizes the sum of squared errors (SSE) or mean squared error (MSE) between the target variable (y) and our predicted output ($\hat{y}$) over all samples in the dataset.
OLS can find the best parameters using of the following methods:
* Solving the model parameters analytically using closed-form equations
* Using an optimization algorithm (Gradient Descent, Stochastic Gradient Descent, Newton’s Method, etc.)
<h2 id="prediction">Prediction</h2>
```
y_hat= regr.predict(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
x = np.asanyarray(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_COMB']])
y = np.asanyarray(test[['CO2EMISSIONS']])
print("Residual sum of squares: %.2f"
% np.mean((y_hat - y) ** 2))
# Explained variance score: 1 is perfect prediction
print('Variance score: %.2f' % regr.score(x, y))
```
**Explained variance regression score:**\
Let $\hat{y}$ be the estimated target output, y the corresponding (correct) target output, and Var be the Variance (the square of the standard deviation). Then the explained variance is estimated as follows:
$\texttt{explainedVariance}(y, \hat{y}) = 1 - \frac{Var{ y - \hat{y}}}{Var{y}}$\
The best possible score is 1.0, the lower values are worse.
<h2 id="practice">Practice</h2>
Try to use a multiple linear regression with the same dataset, but this time use FUELCONSUMPTION_CITY and FUELCONSUMPTION_HWY instead of FUELCONSUMPTION_COMB. Does it result in better accuracy?
```
regr = linear_model.LinearRegression()
x = np.asanyarray(train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(train[['CO2EMISSIONS']])
regr.fit (x, y)
print ('Coefficients: ', regr.coef_)
print ('Intercept: ',regr.intercept_)
y_= regr.predict(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
x = np.asanyarray(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(test[['CO2EMISSIONS']])
print("Residual sum of squares: %.2f"% np.mean((y_ - y) ** 2))
print('Variance score: %.2f' % regr.score(x, y))
```
<details><summary>Click here for the solution</summary>
```python
regr = linear_model.LinearRegression()
x = np.asanyarray(train[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(train[['CO2EMISSIONS']])
regr.fit (x, y)
print ('Coefficients: ', regr.coef_)
y_= regr.predict(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
x = np.asanyarray(test[['ENGINESIZE','CYLINDERS','FUELCONSUMPTION_CITY','FUELCONSUMPTION_HWY']])
y = np.asanyarray(test[['CO2EMISSIONS']])
print("Residual sum of squares: %.2f"% np.mean((y_ - y) ** 2))
print('Variance score: %.2f' % regr.score(x, y))
```
</details>
<h2>Want to learn more?</h2>
IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: <a href="https://www.ibm.com/analytics/spss-statistics-software?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01">SPSS Modeler</a>
Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at <a href="https://www.ibm.com/cloud/watson-studio?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01">Watson Studio</a>
### Thank you for completing this lab!
## Author
Saeed Aghabozorgi
### Other Contributors
<a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDeveloperSkillsNetworkML0101ENSkillsNetwork20718538-2021-01-01" target="_blank">Joseph Santarcangelo</a>
## Change Log
| Date (YYYY-MM-DD) | Version | Changed By | Change Description |
| ----------------- | ------- | ---------- | ---------------------------------- |
| 2020-11-03 | 2.1 | Lakshmi | Made changes in URL |
| 2020-08-27 | 2.0 | Lavanya | Moved lab to course repo in GitLab |
| | | | |
| | | | |
## <h3 align="center"> © IBM Corporation 2020. All rights reserved. <h3/>
| github_jupyter |
In this notebook, we'll learn how to use GANs to do semi-supervised learning.
In supervised learning, we have a training set of inputs $x$ and class labels $y$. We train a model that takes $x$ as input and gives $y$ as output.
In semi-supervised learning, our goal is still to train a model that takes $x$ as input and generates $y$ as output. However, not all of our training examples have a label $y$. We need to develop an algorithm that is able to get better at classification by studying both labeled $(x, y)$ pairs and unlabeled $x$ examples.
To do this for the SVHN dataset, we'll turn the GAN discriminator into an 11 class discriminator. It will recognize the 10 different classes of real SVHN digits, as well as an 11th class of fake images that come from the generator. The discriminator will get to train on real labeled images, real unlabeled images, and fake images. By drawing on three sources of data instead of just one, it will generalize to the test set much better than a traditional classifier trained on only one source of data.
```
%matplotlib inline
import pickle as pkl
import time
import matplotlib.pyplot as plt
import numpy as np
from scipy.io import loadmat
import tensorflow as tf
!mkdir data
from urllib.request import urlretrieve
from os.path import isfile, isdir
from tqdm import tqdm
data_dir = 'data/'
if not isdir(data_dir):
raise Exception("Data directory doesn't exist!")
class DLProgress(tqdm):
last_block = 0
def hook(self, block_num=1, block_size=1, total_size=None):
self.total = total_size
self.update((block_num - self.last_block) * block_size)
self.last_block = block_num
if not isfile(data_dir + "train_32x32.mat"):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='SVHN Training Set') as pbar:
urlretrieve(
'http://ufldl.stanford.edu/housenumbers/train_32x32.mat',
data_dir + 'train_32x32.mat',
pbar.hook)
if not isfile(data_dir + "test_32x32.mat"):
with DLProgress(unit='B', unit_scale=True, miniters=1, desc='SVHN Training Set') as pbar:
urlretrieve(
'http://ufldl.stanford.edu/housenumbers/test_32x32.mat',
data_dir + 'test_32x32.mat',
pbar.hook)
trainset = loadmat(data_dir + 'train_32x32.mat')
testset = loadmat(data_dir + 'test_32x32.mat')
idx = np.random.randint(0, trainset['X'].shape[3], size=36)
fig, axes = plt.subplots(6, 6, sharex=True, sharey=True, figsize=(5,5),)
for ii, ax in zip(idx, axes.flatten()):
ax.imshow(trainset['X'][:,:,:,ii], aspect='equal')
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
plt.subplots_adjust(wspace=0, hspace=0)
def scale(x, feature_range=(-1, 1)):
# scale to (0, 1)
x = ((x - x.min())/(255 - x.min()))
# scale to feature_range
min, max = feature_range
x = x * (max - min) + min
return x
class Dataset:
def __init__(self, train, test, val_frac=0.5, shuffle=True, scale_func=None):
# Even though a validation fraction is defined above, they use bad form here and don't use the validation data set; really, should use validation to help guide hyperparameter choices. Here, we are just going to look at accuracy to cut to the chase
split_idx = int(len(test['y'])*(1 - val_frac))
self.test_x, self.valid_x = test['X'][:,:,:,:split_idx], test['X'][:,:,:,split_idx:]
self.test_y, self.valid_y = test['y'][:split_idx], test['y'][split_idx:]
self.train_x, self.train_y = train['X'], train['y']
# The SVHN dataset comes with lots of labels, but for the purpose of this exercise,
# we will pretend that there are only 1000.
# We use this mask to say which labels we will allow ourselves to use.
#Thus, the label_mask variable will be used to keep track of which labels have access to and which pretend don't have access to
self.label_mask = np.zeros_like(self.train_y)
self.label_mask[0:1000] = 1
self.train_x = np.rollaxis(self.train_x, 3)
self.valid_x = np.rollaxis(self.valid_x, 3)
self.test_x = np.rollaxis(self.test_x, 3)
if scale_func is None:
self.scaler = scale
else:
self.scaler = scale_func
self.train_x = self.scaler(self.train_x)
self.valid_x = self.scaler(self.valid_x)
self.test_x = self.scaler(self.test_x)
self.shuffle = shuffle
# The main function to access the data set class is this batches method described below; just need to specify which batch size you want and use which_set to determine if want to use train or test set
def batches(self, batch_size, which_set="train"):
x_name = which_set + "_x"
y_name = which_set + "_y"
num_examples = len(getattr(dataset, y_name))
if self.shuffle:
idx = np.arange(num_examples)
np.random.shuffle(idx)
setattr(dataset, x_name, getattr(dataset, x_name)[idx])
setattr(dataset, y_name, getattr(dataset, y_name)[idx])
if which_set == "train":
dataset.label_mask = dataset.label_mask[idx]
dataset_x = getattr(dataset, x_name)
dataset_y = getattr(dataset, y_name)
for ii in range(0, num_examples, batch_size):
x = dataset_x[ii:ii+batch_size]
y = dataset_y[ii:ii+batch_size]
if which_set == "train":
# When we use the data for training, we need to include
# the label mask, so we can pretend we don't have access
# to some of the labels, as an exercise of our semi-supervised
# learning ability
yield x, y, self.label_mask[ii:ii+batch_size]
else:
yield x, y
def model_inputs(real_dim, z_dim):
inputs_real = tf.placeholder(tf.float32, (None, *real_dim), name='input_real')# feeding in input images
inputs_z = tf.placeholder(tf.float32, (None, z_dim), name='input_z')# feeding in random noise to guide generator
y = tf.placeholder(tf.int32, (None), name='y')# loading in label classes
label_mask = tf.placeholder(tf.int32, (None), name='label_mask')#placeholder to tell us whether or not can use the label
return inputs_real, inputs_z, y, label_mask
def generator(z, output_dim, reuse=False, alpha=0.2, training=True, size_mult=128):
with tf.variable_scope('generator', reuse=reuse):
# First fully connected layer
x1 = tf.layers.dense(z, 4 * 4 * size_mult * 4)
# Reshape it to start the convolutional stack
x1 = tf.reshape(x1, (-1, 4, 4, size_mult * 4))
x1 = tf.layers.batch_normalization(x1, training=training)
x1 = tf.maximum(alpha * x1, x1)
x2 = tf.layers.conv2d_transpose(x1, size_mult * 2, 5, strides=2, padding='same')
x2 = tf.layers.batch_normalization(x2, training=training)
x2 = tf.maximum(alpha * x2, x2)
x3 = tf.layers.conv2d_transpose(x2, size_mult, 5, strides=2, padding='same')
x3 = tf.layers.batch_normalization(x3, training=training)
x3 = tf.maximum(alpha * x3, x3)
# Output layer--32X32 with 3 output channels
logits = tf.layers.conv2d_transpose(x3, output_dim, 5, strides=2, padding='same')
out = tf.tanh(logits)
return out
def discriminator(x, reuse=False, alpha=0.2, drop_rate=0., num_classes=10, size_mult=64):
#This will be a multi-class classifier. Don't use any max pooling or average pooling to reduce size; instead use a convolution of stride 2 (see The All Convolutional Net for reason why). Also no batch norm just in the first layer (or else hard to make pixels have correct mean and standard dev) but then do use it later. Also use drop out a lot, even in the first layer, because it's a regularization technique to make test error is not too much higher than the training error, and because only using 1000 images, want to make sure don't overfit. Also use leaky relu (the max commands below, which make it so get non negative value, so have a slope of alpha as function of the input)
with tf.variable_scope('discriminator', reuse=reuse):
x = tf.layers.dropout(x, rate=drop_rate/2.5)
# Input layer is 32x32x3
x1 = tf.layers.conv2d(x, size_mult, 3, strides=2, padding='same')
relu1 = tf.maximum(alpha * x1, x1)
relu1 = tf.layers.dropout(relu1, rate=drop_rate)
x2 = tf.layers.conv2d(relu1, size_mult, 3, strides=2, padding='same')
bn2 = tf.layers.batch_normalization(x2, training=True)
relu2 = tf.maximum(alpha * x2, x2)
x3 = tf.layers.conv2d(relu2, size_mult, 3, strides=2, padding='same')
bn3 = tf.layers.batch_normalization(x3, training=True)
relu3 = tf.maximum(alpha * bn3, bn3)
relu3 = tf.layers.dropout(relu3, rate=drop_rate)
x4 = tf.layers.conv2d(relu3, 2 * size_mult, 3, strides=1, padding='same')
bn4 = tf.layers.batch_normalization(x4, training=True)
relu4 = tf.maximum(alpha * bn4, bn4)
x5 = tf.layers.conv2d(relu4, 2 * size_mult, 3, strides=1, padding='same')
bn5 = tf.layers.batch_normalization(x5, training=True)
relu5 = tf.maximum(alpha * bn5, bn5)
x6 = tf.layers.conv2d(relu5, 2 * size_mult, 3, strides=2, padding='same')
bn6 = tf.layers.batch_normalization(x6, training=True)
relu6 = tf.maximum(alpha * bn6, bn6)
relu6 = tf.layers.dropout(relu6, rate=drop_rate)
x7 = tf.layers.conv2d(relu5, 2 * size_mult, 3, strides=1, padding='valid')
# Don't use bn on this layer, because bn would set the mean of each feature
# to the bn mu parameter.
# This layer is used for the feature matching loss, which only works if
# the means can be different when the discriminator is run on the data than
# when the discriminator is run on the generator samples.
relu7 = tf.maximum(alpha * x7, x7)
# In layer 7 above is where use feature matching to make sure average feature value in training data is roughly same as average feature value for generator data. Because taking average, want to make sure average of one type of data will chnage when move to another kind of data, so don't use batch norm because sets mean to be exactly same as the bias parameter, and then feature matching loss wouldn't be able to find any difference at all. Can use weight normalization, but it's a bit more complicated
# Flatten it by global average pooling
# Global average pooling means that for every feature map, take average over all spatial domian of the feature map and return just one value. So, here there is a width by heigh by channel by batch size tensor, and after global max pooling will have tensor of batch size by channel
features = tf.reduce_mean(relu7, axis = (1,2))# the axes are as follows: 0=batch, 1=height, 2 = width, 3 = channels of the feature map, so here take average over 1 and 2 to get rid of those spatial dimensions
# Set class_logits to be the inputs to a softmax distribution over the different classes
class_logits = tf.layers.dense(features, num_classes)# I have implemented the 11th class to zero below
# Set gan_logits such that P(input is real | input) = sigmoid(gan_logits).
# Keep in mind that class_logits gives you the probability distribution over all the real
# classes and the fake class. You need to work out how to transform this multiclass softmax
# distribution into a binary real-vs-fake decision that can be described with a sigmoid.
# Numerical stability is very important.
# You'll probably need to use this numerical stability trick:
# log sum_i exp a_i = m + log sum_i exp(a_i - m).
# This is numerically stable when m = max_i a_i.
# (It helps to think about what goes wrong when...
# 1. One value of a_i is very large
# 2. All the values of a_i are very negative
# This trick and this value of m fix both those cases, but the naive implementation and
# other values of m encounter various problems)
real_class_logits = class_logits
fake_class_logits = 0.0# use this because only used a 10 class matmul
max_val = tf.reduce_max(real_class_logits,axis = 1, keep_dims = True)
stable_real_class_logits = real_class_logits - max_val
gan_logits = max_val + tf.log(tf.reduce_sum(tf.exp(stable_real_class_logits),1)) - fake_class_logits# add the max back in here so its equal but numerically stable
out = tf.nn.softmax(class_logits)
return out, class_logits, gan_logits, features
def model_loss(input_real, input_z, output_dim, y, num_classes, label_mask, alpha=0.2, drop_rate=0.):
"""
Get the loss for the discriminator and generator
:param input_real: Images from the real dataset
:param input_z: Z input
:param output_dim: The number of channels in the output image
:param y: Integer class labels
:param num_classes: The number of classes
:param alpha: The slope of the left half of leaky ReLU activation
:param drop_rate: The probability of dropping a hidden unit
:return: A tuple of (discriminator loss, generator loss)
"""
# These numbers multiply the size of each layer of the generator and the discriminator,
# respectively. You can reduce them to run your code faster for debugging purposes.
g_size_mult = 2#32 # smaller numbers ot make them run faster are something like 2 or 4
d_size_mult = 4#64
# Here we run the generator and the discriminator
g_model = generator(input_z, output_dim, alpha=alpha, size_mult=g_size_mult)
d_on_data = discriminator(input_real, alpha=alpha, drop_rate=drop_rate, size_mult=d_size_mult)
# The above line calculates batch norm statistics (e.g., mean of all features on real data and
# standard dev on features for real data) based on real input data
d_model_real, class_logits_on_data, gan_logits_on_data, data_features = d_on_data
d_on_samples = discriminator(g_model, reuse=True, alpha=alpha, drop_rate=drop_rate, size_mult=d_size_mult)
# in the above call to the discriminator, it runs on samples from the generator
# and calculates totally different batch norm statistics based on the generator (fake) data
# compared to the stats from the discriminator run on real data
d_model_fake, class_logits_on_samples, gan_logits_on_samples, sample_features = d_on_samples
# Can look at paper "Improved techniques for training GANs" for different techniques to try rather than running discriminator on real and separately on fake data
# Here we compute `d_loss`, the loss for the discriminator.
# This should combine two different losses:
# 1. The loss for the GAN problem, where we minimize the cross-entropy for the binary
# real-vs-fake classification problem.
# 2. The loss for the SVHN digit classification problem, where we minimize the cross-entropy
# for the multi-class softmax. For this one we use the labels. Don't forget to ignore
# use `label_mask` to ignore the examples that we are pretending are unlabeled for the
# semi-supervised learning problem.
# Use logit for sigmoid data and set up cross_entropy loss
d_loss_real = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits = gan_logits_on_data,
labels = tf.ones_like(gan_logits_on_data)))# dealing with binary classification, which is why useing sigmoid cross entropy
d_loss_fake = tf.reduce_mean(
tf.nn.sigmoid_cross_entropy_with_logits(logits = gan_logits_on_samples,
labels = tf.zeros_like(gan_logits_on_samples)))# dealing with binary classification, which is why useing sigmoid cross entropy
# Below is the loss for the supervised portion of the semi-supervised learning
# Need to remember to pay attention only to the label mask values, so zero out cross entropy where don't have the label
y = tf.squeeze(y)
class_cross_entropy = tf.nn.softmax_cross_entropy_with_logits(logits = class_logits_on_data,
labels = tf.one_hot(y,num_classes,
dtype=tf.float32))
class_cross_entropy = tf.squeeze(class_cross_entropy)
label_mask = tf.squeeze(tf.to_float(label_mask))
d_loss_class = tf.reduce_sum(label_mask * class_cross_entropy)/tf.maximum(1.0, tf.reduce_sum(label_mask))# Can't just take the tf.reduce_mean because there may be some or all that are zero, which is why have to mutliply the label mask by cross entropy in numerator, and then to calculate demoniaror, have to sum up all the ones in the label masks. If there is a minibatch that has all zero label masks, then would have a denominator (and numberator) or zero, which is why have the max part in the denominator
d_loss = d_loss_fake + d_loss_real + d_loss_class
# Here we set `g_loss` to the "feature matching" loss invented by Tim Salimans at OpenAI.
# This loss consists of minimizing the absolute difference between the expected features (i.e., mean features)
# on the data and the expected features on the generated samples.
# This loss works better for semi-supervised learning than the tradition GAN losses.
data_moments = tf.reduce_mean(data_features, axis = 0)
sample_moments = tf.reduce_mean(sample_features, axis = 0)
g_loss = tf.reduce_mean(tf.abs(sample_moments - data_moments))#helps make sure values in discriminator average is the same no matter if run on data or generated samples, which will help force generator to calculate average of generated samples to be same as the data
# Moment matching is when take stats from one data set and ask them to be similar to statistics from another dataset, which is why use the word moments up above; each stat we extract is a moment
pred_class = tf.cast(tf.argmax(class_logits_on_data, 1), tf.int32)
eq = tf.equal(tf.squeeze(y), pred_class)
correct = tf.reduce_sum(tf.to_float(eq))
masked_correct = tf.reduce_sum(label_mask * tf.to_float(eq))
return d_loss, g_loss, correct, masked_correct, g_model
def model_opt(d_loss, g_loss, learning_rate, beta1):
"""
Get optimization operations
:param d_loss: Discriminator loss Tensor
:param g_loss: Generator loss Tensor
:param learning_rate: Learning Rate Placeholder
:param beta1: The exponential decay rate for the 1st moment in the optimizer
:return: A tuple of (discriminator training operation, generator training operation)
"""
# Get weights and biases to update. Get them separately for the discriminator and the generator
t_vars = tf.trainable_variables()
d_vars = [vars for vars in t_vars if vars.name.startswith( 'discriminator')]
g_vars = [vars for vars in t_vars if vars.name.startswith('generator')]
#this is just to make sure all the variables we are dealing with are associated
#with the generator or discriminator
for t in t_vars:
assert t in d_vars or t in g_vars
# Minimize both players' costs simultaneously
d_train_opt = tf.train.AdamOptimizer(learning_rate = learning_rate, beta1 = beta1).minimize(d_loss, var_list = d_vars)
g_train_opt = tf.train.AdamOptimizer(learning_rate = learning_rate, beta1 = beta1).minimize(g_loss, var_list = g_vars)
shrink_lr = tf.assign(learning_rate, learning_rate * 0.9)
return d_train_opt, g_train_opt, shrink_lr
class GAN:
"""
A GAN model.
:param real_size: The shape of the real data.
:param z_size: The number of entries in the z code vector.
:param learnin_rate: The learning rate to use for Adam.
:param num_classes: The number of classes to recognize.
:param alpha: The slope of the left half of the leaky ReLU activation
:param beta1: The beta1 parameter for Adam.
"""
def __init__(self, real_size, z_size, learning_rate, num_classes=10, alpha=0.2, beta1=0.5):
tf.reset_default_graph()
self.learning_rate = tf.Variable(learning_rate, trainable=False)
inputs = model_inputs(real_size, z_size)
self.input_real, self.input_z, self.y, self.label_mask = inputs
self.drop_rate = tf.placeholder_with_default(.5, (), "drop_rate")
loss_results = model_loss(self.input_real, self.input_z,
real_size[2], self.y, num_classes,
label_mask=self.label_mask,
alpha=0.2,
drop_rate=self.drop_rate)
self.d_loss, self.g_loss, self.correct, self.masked_correct, self.samples = loss_results
self.d_opt, self.g_opt, self.shrink_lr = model_opt(self.d_loss, self.g_loss, self.learning_rate, beta1)
def view_samples(epoch, samples, nrows, ncols, figsize=(5,5)):
fig, axes = plt.subplots(figsize=figsize, nrows=nrows, ncols=ncols,
sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
ax.axis('off')
img = ((img - img.min())*255 / (img.max() - img.min())).astype(np.uint8)
ax.set_adjustable('box-forced')
im = ax.imshow(img)
plt.subplots_adjust(wspace=0, hspace=0)
return fig, axes
def train(net, dataset, epochs, batch_size, figsize=(5,5)):
saver = tf.train.Saver()
sample_z = np.random.normal(0, 1, size=(50, z_size))
samples, train_accuracies, test_accuracies = [], [], []
steps = 0
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for e in range(epochs):
print("Epoch",e)
t1e = time.time()
num_examples = 0
num_correct = 0
for x, y, label_mask in dataset.batches(batch_size):#this is simplified; for a real applicaiton
#would use a tensorflow queue to run things asynchronously
assert 'int' in str(y.dtype)
steps += 1
num_examples += label_mask.sum()
# Sample random noise for G
batch_z = np.random.normal(0, 1, size=(batch_size, z_size))# also simplified here compared to a
# real application in which would use a tensorflow random number generator that would
# happen on a GPU rather than a CPU
# Run optimizers
t1 = time.time()
_, _, correct = sess.run([net.d_opt, net.g_opt, net.masked_correct],
feed_dict={net.input_real: x, net.input_z: batch_z,
net.y : y, net.label_mask : label_mask})
t2 = time.time()
num_correct += correct
sess.run([net.shrink_lr])
train_accuracy = num_correct / float(num_examples)# only calculating accuracy for labeled data and it's updated on the fly with each new minibatch at the moment actually training
print("\t\tClassifier train accuracy: ", train_accuracy)# this is average throughout entire time of epoch, so it will be a little higher than what it actually is because the avarege is a little back in time, so test accuracy may be a little higher than train accuracy because train accuracy is over whole epoch rather than instataneously
num_examples = 0
num_correct = 0
for x, y in dataset.batches(batch_size, which_set="test"):
assert 'int' in str(y.dtype)
num_examples += x.shape[0]
correct, = sess.run([net.correct], feed_dict={net.input_real: x,
net.y : y,
net.drop_rate: 0.})
num_correct += correct
test_accuracy = num_correct / float(num_examples)
print("\t\tClassifier test accuracy", test_accuracy)
print("\t\tStep time: ", t2 - t1)
t2e = time.time()
print("\t\tEpoch time: ", t2e - t1e)
gen_samples = sess.run(
net.samples,
feed_dict={net.input_z: sample_z})
samples.append(gen_samples)
_ = view_samples(-1, samples, 5, 10, figsize=figsize)
plt.show()
# Save history of accuracies to view after training
train_accuracies.append(train_accuracy)
test_accuracies.append(test_accuracy)
saver.save(sess, './checkpoints/generator.ckpt')
with open('samples.pkl', 'wb') as f:
pkl.dump(samples, f)
return train_accuracies, test_accuracies, samples
!mkdir checkpoints
real_size = (32,32,3)
z_size = 100
learning_rate = 0.0003
net = GAN(real_size, z_size, learning_rate)
# This is where most of the action happens and where can get a feel for hyper parameter tuning
# Each time run notebook, will get slightly different results
# Training and test accuracy should increase, and the training accuracy will be much higher (e.g., 0.2 higher) than test accuracy because using so few data
# In real life, should use a validation test set to determine whne stop training
# Test accuracy of 0.69 or above is good
# This notebook was set up to do fairly good semisupervised learning, not create new images (so the output of digits in this notebook will not be great), which is fine here because we just want to give the discriminator a bit more data to train with
dataset = Dataset(trainset, testset)
batch_size = 128
epochs = 25
train_accuracies, test_accuracies, samples = train(net,
dataset,
epochs,
batch_size,
figsize=(10,5))
fig, ax = plt.subplots()
plt.plot(train_accuracies, label='Train', alpha=0.5)
plt.plot(test_accuracies, label='Test', alpha=0.5)
plt.title("Accuracy")
plt.legend()
```
When you run the fully implemented semi-supervised GAN, you should usually find that the test accuracy peaks at 69-71%. It should definitely stay above 68% fairly consistently throughout the last several epochs of training.
This is a little bit better than a [NIPS 2014 paper](https://arxiv.org/pdf/1406.5298.pdf) that got 64% accuracy on 1000-label SVHN with variational methods. However, we still have lost something by not using all the labels. If you re-run with all the labels included, you should obtain over 80% accuracy using this architecture (and other architectures that take longer to run can do much better).
```
_ = view_samples(-1, samples, 5, 10, figsize=(10,5))
!mkdir images
for ii in range(len(samples)):
fig, ax = view_samples(ii, samples, 5, 10, figsize=(10,5))
fig.savefig('images/samples_{:03d}.png'.format(ii))
plt.close()
```
Congratulations! You now know how to train a semi-supervised GAN. This exercise is stripped down to make it run faster and to make it simpler to implement. In the original work by Tim Salimans at OpenAI, a GAN using [more tricks and more runtime](https://arxiv.org/pdf/1606.03498.pdf) reaches over 94% accuracy using only 1,000 labeled examples.
| github_jupyter |
# Deep learning for computer vision
This notebook will teach you to build and train convolutional networks for image recognition. Brace yourselves.
# CIFAR dataset
This week, we shall focus on the image recognition problem on cifar10 dataset
* 60k images of shape 3x32x32
* 10 different classes: planes, dogs, cats, trucks, etc.
<img src="cifar10.jpg" style="width:80%">
```
# when running in colab, un-comment this
# !wget https://raw.githubusercontent.com/yandexdataschool/Practical_DL/fall19/week03_convnets/cifar.py
import numpy as np
from cifar import load_cifar10
X_train, y_train, X_val, y_val, X_test, y_test = load_cifar10("cifar_data")
class_names = np.array(['airplane', 'automobile', 'bird', 'cat', 'deer',
'dog', 'frog', 'horse', 'ship', 'truck'])
print(X_train.shape,y_train.shape)
import matplotlib.pyplot as plt
%matplotlib inline
plt.figure(figsize=[12,10])
for i in range(12):
plt.subplot(3,4,i+1)
plt.xlabel(class_names[y_train[i]])
plt.imshow(np.transpose(X_train[i],[1,2,0]))
```
# Building a network
Simple neural networks with layers applied on top of one another can be implemented as `torch.nn.Sequential` - just add a list of pre-built modules and let it train.
```
import torch, torch.nn as nn
import torch.nn.functional as F
# a special module that converts [batch, channel, w, h] to [batch, units]
class Flatten(nn.Module):
def forward(self, input):
return input.view(input.size(0), -1)
```
Let's start with a dense network for our baseline:
```
model = nn.Sequential()
# reshape from "images" to flat vectors
model.add_module('flatten', Flatten())
# dense "head"
model.add_module('dense1', nn.Linear(3 * 32 * 32, 64))
model.add_module('dense1_relu', nn.ReLU())
model.add_module('dense2_logits', nn.Linear(64, 10)) # logits for 10 classes
```
As in our basic tutorial, we train our model with negative log-likelihood aka crossentropy.
```
def compute_loss(X_batch, y_batch):
X_batch = torch.as_tensor(X_batch, dtype=torch.float32)
y_batch = torch.as_tensor(y_batch, dtype=torch.int64)
logits = model(X_batch)
return F.cross_entropy(logits, y_batch).mean()
# example
compute_loss(X_train[:5], y_train[:5])
```
### Training on minibatches
* We got 40k images, that's way too many for a full-batch SGD. Let's train on minibatches instead
* Below is a function that splits the training sample into minibatches
```
# An auxilary function that returns mini-batches for neural network training
def iterate_minibatches(X, y, batchsize):
indices = np.random.permutation(np.arange(len(X)))
for start in range(0, len(indices), batchsize):
ix = indices[start: start + batchsize]
yield X[ix], y[ix]
opt = torch.optim.SGD(model.parameters(), lr=0.01)
train_loss = []
val_accuracy = []
import time
num_epochs = 100 # total amount of full passes over training data
batch_size = 50 # number of samples processed in one SGD iteration
for epoch in range(num_epochs):
# In each epoch, we do a full pass over the training data:
start_time = time.time()
model.train(True) # enable dropout / batch_norm training behavior
for X_batch, y_batch in iterate_minibatches(X_train, y_train, batch_size):
# train on batch
loss = compute_loss(X_batch, y_batch)
loss.backward()
opt.step()
opt.zero_grad()
train_loss.append(loss.data.numpy())
# And a full pass over the validation data:
model.train(False) # disable dropout / use averages for batch_norm
for X_batch, y_batch in iterate_minibatches(X_val, y_val, batch_size):
logits = model(torch.as_tensor(X_batch, dtype=torch.float32))
y_pred = logits.max(1)[1].data.numpy()
val_accuracy.append(np.mean(y_batch == y_pred))
# Then we print the results for this epoch:
print("Epoch {} of {} took {:.3f}s".format(
epoch + 1, num_epochs, time.time() - start_time))
print(" training loss (in-iteration): \t{:.6f}".format(
np.mean(train_loss[-len(X_train) // batch_size :])))
print(" validation accuracy: \t\t\t{:.2f} %".format(
np.mean(val_accuracy[-len(X_val) // batch_size :]) * 100))
```
Don't wait for full 100 epochs. You can interrupt training after 5-20 epochs once validation accuracy stops going up.
```
```
```
```
```
```
```
```
```
```
### Final test
```
model.train(False) # disable dropout / use averages for batch_norm
test_batch_acc = []
for X_batch, y_batch in iterate_minibatches(X_test, y_test, 500):
logits = model(torch.as_tensor(X_batch, dtype=torch.float32))
y_pred = logits.max(1)[1].data.numpy()
test_batch_acc.append(np.mean(y_batch == y_pred))
test_accuracy = np.mean(test_batch_acc)
print("Final results:")
print(" test accuracy:\t\t{:.2f} %".format(
test_accuracy * 100))
if test_accuracy * 100 > 95:
print("Double-check, than consider applying for NIPS'17. SRSly.")
elif test_accuracy * 100 > 90:
print("U'r freakin' amazin'!")
elif test_accuracy * 100 > 80:
print("Achievement unlocked: 110lvl Warlock!")
elif test_accuracy * 100 > 70:
print("Achievement unlocked: 80lvl Warlock!")
elif test_accuracy * 100 > 60:
print("Achievement unlocked: 70lvl Warlock!")
elif test_accuracy * 100 > 50:
print("Achievement unlocked: 60lvl Warlock!")
else:
print("We need more magic! Follow instructons below")
```
## Task I: small convolution net
### First step
Let's create a mini-convolutional network with roughly such architecture:
* Input layer
* 3x3 convolution with 10 filters and _ReLU_ activation
* 2x2 pooling (or set previous convolution stride to 3)
* Flatten
* Dense layer with 100 neurons and _ReLU_ activation
* 10% dropout
* Output dense layer.
__Convolutional layers__ in torch are just like all other layers, but with a specific set of parameters:
__`...`__
__`model.add_module('conv1', nn.Conv2d(in_channels=3, out_channels=10, kernel_size=3)) # convolution`__
__`model.add_module('pool1', nn.MaxPool2d(2)) # max pooling 2x2`__
__`...`__
Once you're done (and compute_loss no longer raises errors), train it with __Adam__ optimizer with default params (feel free to modify the code above).
If everything is right, you should get at least __50%__ validation accuracy.
```
```
```
```
```
```
```
```
```
```
__Hint:__ If you don't want to compute shapes by hand, just plug in any shape (e.g. 1 unit) and run compute_loss. You will see something like this:
__`RuntimeError: size mismatch, m1: [5 x 1960], m2: [1 x 64] at /some/long/path/to/torch/operation`__
See the __1960__ there? That's your actual input shape.
## Task 2: adding normalization
* Add batch norm (with default params) between convolution and ReLU
* nn.BatchNorm*d (1d for dense, 2d for conv)
* usually better to put them after linear/conv but before nonlinearity
* Re-train the network with the same optimizer, it should get at least 60% validation accuracy at peak.
```
```
```
```
```
```
```
```
```
```
```
```
```
```
## Task 3: Data Augmentation
There's a powerful torch tool for image preprocessing useful to do data preprocessing and augmentation.
Here's how it works: we define a pipeline that
* makes random crops of data (augmentation)
* randomly flips image horizontally (augmentation)
* then normalizes it (preprocessing)
```
from torchvision import transforms
means = np.array((0.4914, 0.4822, 0.4465))
stds = np.array((0.2023, 0.1994, 0.2010))
transform_augment = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomRotation([-30, 30]),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(means, stds),
])
from torchvision.datasets import CIFAR10
train_loader = CIFAR10("./cifar_data/", train=True, transform=transform_augment)
train_batch_gen = torch.utils.data.DataLoader(train_loader,
batch_size=32,
shuffle=True,
num_workers=1)
for (x_batch, y_batch) in train_batch_gen:
print('X:', type(x_batch), x_batch.shape)
print('y:', type(y_batch), y_batch.shape)
for i, img in enumerate(x_batch.numpy()[:8]):
plt.subplot(2, 4, i+1)
plt.imshow(img.transpose([1,2,0]) * stds + means )
raise NotImplementedError("Plese use this code in your training loop")
# TODO use this in your training loop
```
When testing, we don't need random crops, just normalize with same statistics.
```
transform_test = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize(means, stds),
])
test_loader = <YOUR CODE>
```
# Homework 2.2: The Quest For A Better Network
In this assignment you will build a monster network to solve CIFAR10 image classification.
This notebook is intended as a sequel to seminar 3, please give it a try if you haven't done so yet.
(please read it at least diagonally)
* The ultimate quest is to create a network that has as high __accuracy__ as you can push it.
* There is a __mini-report__ at the end that you will have to fill in. We recommend reading it first and filling it while you iterate.
## Grading
* starting at zero points
* +20% for describing your iteration path in a report below.
* +20% for building a network that gets above 20% accuracy
* +10% for beating each of these milestones on __TEST__ dataset:
* 50% (50% points)
* 60% (60% points)
* 65% (70% points)
* 70% (80% points)
* 75% (90% points)
* 80% (full points)
## Restrictions
* Please do NOT use pre-trained networks for this assignment until you reach 80%.
* In other words, base milestones must be beaten without pre-trained nets (and such net must be present in the e-mail). After that, you can use whatever you want.
* you __can__ use validation data for training, but you __can't'__ do anything with test data apart from running the evaluation procedure.
## Tips on what can be done:
* __Network size__
* MOAR neurons,
* MOAR layers, ([torch.nn docs](http://pytorch.org/docs/master/nn.html))
* Nonlinearities in the hidden layers
* tanh, relu, leaky relu, etc
* Larger networks may take more epochs to train, so don't discard your net just because it could didn't beat the baseline in 5 epochs.
* Ph'nglui mglw'nafh Cthulhu R'lyeh wgah'nagl fhtagn!
### The main rule of prototyping: one change at a time
* By now you probably have several ideas on what to change. By all means, try them out! But there's a catch: __never test several new things at once__.
### Optimization
* Training for 100 epochs regardless of anything is probably a bad idea.
* Some networks converge over 5 epochs, others - over 500.
* Way to go: stop when validation score is 10 iterations past maximum
* You should certainly use adaptive optimizers
* rmsprop, nesterov_momentum, adam, adagrad and so on.
* Converge faster and sometimes reach better optima
* It might make sense to tweak learning rate/momentum, other learning parameters, batch size and number of epochs
* __BatchNormalization__ (nn.BatchNorm2d) for the win!
* Sometimes more batch normalization is better.
* __Regularize__ to prevent overfitting
* Add some L2 weight norm to the loss function, PyTorch will do the rest
* Can be done manually or with weight_decay parameter of a optimizer ([for example SGD's doc](https://pytorch.org/docs/stable/optim.html#torch.optim.SGD)).
* Dropout (`nn.Dropout`) - to prevent overfitting
* Don't overdo it. Check if it actually makes your network better
### Convolution architectures
* This task __can__ be solved by a sequence of convolutions and poolings with batch_norm and ReLU seasoning, but you shouldn't necessarily stop there.
* [Inception family](https://hacktilldawn.com/2016/09/25/inception-modules-explained-and-implemented/), [ResNet family](https://towardsdatascience.com/an-overview-of-resnet-and-its-variants-5281e2f56035?gi=9018057983ca), [Densely-connected convolutions (exotic)](https://arxiv.org/abs/1608.06993), [Capsule networks (exotic)](https://arxiv.org/abs/1710.09829)
* Please do try a few simple architectures before you go for resnet-152.
* Warning! Training convolutional networks can take long without GPU. That's okay.
* If you are CPU-only, we still recomment that you try a simple convolutional architecture
* a perfect option is if you can set it up to run at nighttime and check it up at the morning.
* Make reasonable layer size estimates. A 128-neuron first convolution is likely an overkill.
* __To reduce computation__ time by a factor in exchange for some accuracy drop, try using __stride__ parameter. A stride=2 convolution should take roughly 1/4 of the default (stride=1) one.
### Data augmemntation
* getting 5x as large dataset for free is a great
* Zoom-in+slice = move
* Rotate+zoom(to remove black stripes)
* Add Noize (gaussian or bernoulli)
* Simple way to do that (if you have PIL/Image):
* ```from scipy.misc import imrotate,imresize```
* and a few slicing
* Other cool libraries: cv2, skimake, PIL/Pillow
* A more advanced way is to use torchvision transforms:
```
transform_train = transforms.Compose([
transforms.RandomCrop(32, padding=4),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize((0.4914, 0.4822, 0.4465), (0.2023, 0.1994, 0.2010)),
])
trainset = torchvision.datasets.CIFAR10(root=path_to_cifar_like_in_seminar, train=True, download=True, transform=transform_train)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=128, shuffle=True, num_workers=2)
```
* Or use this tool from Keras (requires theano/tensorflow): [tutorial](https://blog.keras.io/building-powerful-image-classification-models-using-very-little-data.html), [docs](https://keras.io/preprocessing/image/)
* Stay realistic. There's usually no point in flipping dogs upside down as that is not the way you usually see them.
```
```
```
```
```
```
```
```
```
# you might as well write your solution here :)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
```
# Distributed training in TensorFlow
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/r1/tutorials/distribute/keras.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/r1/tutorials/distribute/keras.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
</td>
</table>
## Overview
The `tf.distribute.Strategy` API provides an abstraction for distributing your training
across multiple processing units. The goal is to allow users to enable distributed training using existing models and training code, with minimal changes.
This tutorial uses the `tf.distribute.MirroredStrategy`, which
does in-graph replication with synchronous training on many GPUs on one machine.
Essentially, it copies all of the model's variables to each processor.
Then, it uses [all-reduce](http://mpitutorial.com/tutorials/mpi-reduce-and-allreduce/) to combine the gradients from all processors and applies the combined value to all copies of the model.
`MirroredStategy` is one of several distribution strategy available in TensorFlow core. You can read about more strategies at [distribution strategy guide](../../guide/distribute_strategy.ipynb).
### Keras API
This example uses the `tf.keras` API to build the model and training loop. For custom training loops, see [this tutorial](training_loops.ipynb).
## Import Dependencies
```
from __future__ import absolute_import, division, print_function, unicode_literals
# Import TensorFlow
try:
# %tensorflow_version only exists in Colab.
%tensorflow_version 2.x
except Exception:
pass
import tensorflow.compat.v1 as tf
import tensorflow_datasets as tfds
import os
```
## Download the dataset
Download the MNIST dataset and load it from [TensorFlow Datasets](https://www.tensorflow.org/datasets). This returns a dataset in `tf.data` format.
Setting `with_info` to `True` includes the metadata for the entire dataset, which is being saved here to `ds_info`.
Among other things, this metadata object includes the number of train and test examples.
```
datasets, ds_info = tfds.load(name='mnist', with_info=True, as_supervised=True)
mnist_train, mnist_test = datasets['train'], datasets['test']
```
## Define Distribution Strategy
Create a `MirroredStrategy` object. This will handle distribution, and provides a context manager (`tf.distribute.MirroredStrategy.scope`) to build your model inside.
```
strategy = tf.distribute.MirroredStrategy()
print ('Number of devices: {}'.format(strategy.num_replicas_in_sync))
```
## Setup Input pipeline
If a model is trained on multiple GPUs, the batch size should be increased accordingly so as to make effective use of the extra computing power. Moreover, the learning rate should be tuned accordingly.
```
# You can also do ds_info.splits.total_num_examples to get the total
# number of examples in the dataset.
num_train_examples = ds_info.splits['train'].num_examples
num_test_examples = ds_info.splits['test'].num_examples
BUFFER_SIZE = 10000
BATCH_SIZE_PER_REPLICA = 64
BATCH_SIZE = BATCH_SIZE_PER_REPLICA * strategy.num_replicas_in_sync
```
Pixel values, which are 0-255, [have to be normalized to the 0-1 range](https://en.wikipedia.org/wiki/Feature_scaling). Define this scale in a function.
```
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
```
Apply this function to the training and test data, shuffle the training data, and [batch it for training](https://www.tensorflow.org/api_docs/python/tf/data/Dataset#batch).
```
train_dataset = mnist_train.map(scale).shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
eval_dataset = mnist_test.map(scale).batch(BATCH_SIZE)
```
## Create the model
Create and compile the Keras model in the context of `strategy.scope`.
```
with strategy.scope():
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10, activation='softmax')
])
model.compile(loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
```
## Define the callbacks.
The callbacks used here are:
* *Tensorboard*: This callback writes a log for Tensorboard which allows you to visualize the graphs.
* *Model Checkpoint*: This callback saves the model after every epoch.
* *Learning Rate Scheduler*: Using this callback, you can schedule the learning rate to change after every epoch/batch.
For illustrative purposes, add a print callback to display the *learning rate* in the notebook.
```
# Define the checkpoint directory to store the checkpoints
checkpoint_dir = './training_checkpoints'
# Name of the checkpoint files
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt_{epoch}")
# Function for decaying the learning rate.
# You can define any decay function you need.
def decay(epoch):
if epoch < 3:
return 1e-3
elif epoch >= 3 and epoch < 7:
return 1e-4
else:
return 1e-5
# Callback for printing the LR at the end of each epoch.
class PrintLR(tf.keras.callbacks.Callback):
def on_epoch_end(self, epoch, logs=None):
print ('\nLearning rate for epoch {} is {}'.format(
epoch + 1, tf.keras.backend.get_value(model.optimizer.lr)))
callbacks = [
tf.keras.callbacks.TensorBoard(log_dir='./logs'),
tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_prefix,
save_weights_only=True),
tf.keras.callbacks.LearningRateScheduler(decay),
PrintLR()
]
```
## Train and evaluate
Now, train the model in the usual way, calling `fit` on the model and passing in the dataset created at the beginning of the tutorial. This step is the same whether you are distributing the training or not.
```
model.fit(train_dataset, epochs=10, callbacks=callbacks)
```
As you can see below, the checkpoints are getting saved.
```
# check the checkpoint directory
!ls {checkpoint_dir}
```
To see how the model perform, load the latest checkpoint and call `evaluate` on the test data.
Call `evaluate` as before using appropriate datasets.
```
model.load_weights(tf.train.latest_checkpoint(checkpoint_dir))
eval_loss, eval_acc = model.evaluate(eval_dataset)
print ('Eval loss: {}, Eval Accuracy: {}'.format(eval_loss, eval_acc))
```
To see the output, you can download and view the TensorBoard logs at the terminal.
```
$ tensorboard --logdir=path/to/log-directory
```
```
!ls -sh ./logs
```
## Export to SavedModel
If you want to export the graph and the variables, SavedModel is the best way of doing this. The model can be loaded back with or without the scope. Moreover, SavedModel is platform agnostic.
```
path = 'saved_model/'
tf.keras.experimental.export_saved_model(model, path)
```
Load the model without `strategy.scope`.
```
unreplicated_model = tf.keras.experimental.load_from_saved_model(path)
unreplicated_model.compile(
loss='sparse_categorical_crossentropy',
optimizer=tf.keras.optimizers.Adam(),
metrics=['accuracy'])
eval_loss, eval_acc = unreplicated_model.evaluate(eval_dataset)
print ('Eval loss: {}, Eval Accuracy: {}'.format(eval_loss, eval_acc))
```
## What's next?
Read the [distribution strategy guide](../../guide/distribute_strategy_tf1.ipynb).
Note: `tf.distribute.Strategy` is actively under development and we will be adding more examples and tutorials in the near future. Please give it a try. We welcome your feedback via [issues on GitHub](https://github.com/tensorflow/tensorflow/issues/new).
| github_jupyter |
# ARDRegressor with PowerTransformer
This Code template is for regression analysis using the ARDRegressor with Feature Transformation technique PowerTransformer.
### Required Packages
```
import warnings
import numpy as np
import pandas as pd
import seaborn as se
import matplotlib.pyplot as plt
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import PowerTransformer
from sklearn.model_selection import train_test_split
from sklearn.metrics import r2_score, mean_absolute_error, mean_squared_error
from sklearn.linear_model import ARDRegression
warnings.filterwarnings('ignore')
```
### Initialization
Filepath of CSV file
```
#filepath
file_path= ""
```
List of features which are required for model training.
```
#x_values
features=[]
```
Target feature for prediction.
```
#y_value
target=''
```
### Data Fetching
Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
```
df=pd.read_csv(file_path)
df.head()
```
### Feature Selections
It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
We will assign all the required input features to X and target/outcome to Y.
```
X=df[features]
Y=df[target]
```
### Data Preprocessing
Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
```
def NullClearner(df):
if(isinstance(df, pd.Series) and (df.dtype in ["float64","int64"])):
df.fillna(df.mean(),inplace=True)
return df
elif(isinstance(df, pd.Series)):
df.fillna(df.mode()[0],inplace=True)
return df
else:return df
def EncodeX(df):
return pd.get_dummies(df)
```
Calling preprocessing functions on the feature and target set.
```
x=X.columns.to_list()
for i in x:
X[i]=NullClearner(X[i])
X=EncodeX(X)
Y=NullClearner(Y)
X.head()
```
#### Correlation Map
In order to check the correlation between the features, we will plot a correlation matrix. It is effective in summarizing a large amount of data where the goal is to see patterns.
```
f,ax = plt.subplots(figsize=(18, 18))
matrix = np.triu(X.corr())
se.heatmap(X.corr(), annot=True, linewidths=.5, fmt= '.1f',ax=ax, mask=matrix)
plt.show()
```
### Data Splitting
The train-test split is a procedure for evaluating the performance of an algorithm. The procedure involves taking a dataset and dividing it into two subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
```
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
```
### Feature Transformation
Power transforms are a family of parametric, monotonic transformations that are applied to make data more Gaussian-like. This is useful for modeling issues related to heteroscedasticity (non-constant variance), or other situations where normality is desired.
[More on PowerTransformer module and parameters](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.PowerTransformer.html)
### Model
Bayesian ARD regression.
Fit the weights of a regression model, using an ARD prior. The weights of the regression model are assumed to be in Gaussian distributions. Also estimate the parameters lambda (precisions of the distributions of the weights) and alpha (precision of the distribution of the noise). The estimation is done by an iterative procedures (Evidence Maximization)
#### Parameters:
> - **n_iter: int, default=300** -> Maximum number of iterations.
> - **tol: float, default=1e-3** -> Stop the algorithm if w has converged.
> - **alpha_1: float, default=1e-6** -> Hyper-parameter : shape parameter for the Gamma distribution prior over the alpha parameter.
> - **alpha_2: float, default=1e-6** -> Hyper-parameter : inverse scale parameter (rate parameter) for the Gamma distribution prior over the alpha parameter.
> - **lambda_1: float, default=1e-6** -> Hyper-parameter : shape parameter for the Gamma distribution prior over the lambda parameter.
> - **lambda_2: float, default=1e-6** -> Hyper-parameter : inverse scale parameter (rate parameter) for the Gamma distribution prior over the lambda parameter.
> - **compute_score: bool, default=False** -> If True, compute the objective function at each step of the model.
> - **threshold_lambda: float, default=10 000** -> threshold for removing (pruning) weights with high precision from the computation.
> - **fit_intercept: bool, default=True** -> whether to calculate the intercept for this model. If set to false, no intercept will be used in calculations (i.e. data is expected to be centered).
> - **normalize: bool, default=False** -> This parameter is ignored when fit_intercept is set to False. If True, the regressors X will be normalized before regression by subtracting the mean and dividing by the l2-norm. If you wish to standardize, please use StandardScaler before calling fit on an estimator with normalize=False.
> - **copy_X: bool, default=True** -> If True, X will be copied; else, it may be overwritten.
> - **verbose: bool, default=False** -> Verbose mode when fitting the model.
```
model=make_pipeline(PowerTransformer(), ARDRegression())
model.fit(x_train,y_train)
```
#### Model Accuracy
We will use the trained model to make a prediction on the test set.Then use the predicted value for measuring the accuracy of our model.
score: The score function returns the coefficient of determination R2 of the prediction.
```
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
```
> **r2_score**: The **r2_score** function computes the percentage variablility explained by our model, either the fraction or the count of correct predictions.
> **mae**: The **mean abosolute error** function calculates the amount of total error(absolute average distance between the real data and the predicted data) by our model.
> **mse**: The **mean squared error** function squares the error(penalizes the model for large errors) by our model.
```
y_pred=model.predict(x_test)
print("R2 Score: {:.2f} %".format(r2_score(y_test,y_pred)*100))
print("Mean Absolute Error {:.2f}".format(mean_absolute_error(y_test,y_pred)))
print("Mean Squared Error {:.2f}".format(mean_squared_error(y_test,y_pred)))
```
#### Prediction Plot
First, we make use of a plot to plot the actual observations, with x_train on the x-axis and y_train on the y-axis.
For the regression line, we will use x_train on the x-axis and then the predictions of the x_train observations on the y-axis.
```
n=len(x_test) if len(x_test)<20 else 20
plt.figure(figsize=(14,10))
plt.plot(range(n),y_test[0:n], color = "green")
plt.plot(range(n),model.predict(x_test[0:n]), color = "red")
plt.legend(["Actual","prediction"])
plt.title("Predicted vs True Value")
plt.xlabel("Record number")
plt.ylabel(target)
plt.show()
```
#### Creator: Nikhil Shrotri , Github: [Profile](https://github.com/nikhilshrotri)
| github_jupyter |
# Define and run a distributed training pipeline
In this notebook we will use **MLRun** to run all the functions we've written in the [mlrun-mpijob-classify](mlrun_mpijob_classify.ipynb) and [nuclio-serving-tf-images](nuclio-serving-tf-images.ipynb) in a **Kubeflow Pipeline**.
**Kubeflow Pipelines** will supply the orchastration to run the pipeline, while **MLRun** will supply an easy interface to define the pipeline and lunch the serving function at the end.
We will show how to:
* Run remote functions from notebooks using `code_to_function`
* Run saved functions from our DB using `import_function`
* How to define and lunch a Kubeflow Pipeline
* How to access the DB from the code and list the pipeline's entries
```
# nuclio: ignore
import nuclio
from mlrun import new_function, code_to_function, get_run_db, mount_v3io, mlconf, new_model_server, v3io_cred, import_function
import os
mlconf.dbpath = 'http://mlrun-api:8080'
base_dir = '/User/mlrun/examples'
images_path = os.path.join(base_dir, 'images')
model_name = 'cat_vs_dog_v1'
```
## Import and define ML functions for our pipeline (utils, training, serving)
Using `code_to_function` we parse the given python file and build a function from it
```
# Build a function from the python source code
# The code contains functions for data import and labeling
utilsfn = code_to_function(name='file_utils',
filename='../src/utils.py',
image='mlrun/mlrun:0.4.6',
kind='job')
# Add mount access to the function
utilsfn.apply(mount_v3io())
utilsfn.export('../yaml/utils.yaml')
```
Using `import_function` we import the horovod training function from our DB.
As we can see, all the function deployment parameters were saved, like Replicas, GPU Configuration, Mounts, Runtime and the code source.
> Please verify that the `HOROVOD_FILE` path (specified in `spec.command`) is available from the cluster (Local path and Mounted path may vary)
```
# read the training function object from MLRun DB
trainer_fn = import_function('db://horovod-trainer')
print(trainer_fn.to_yaml())
```
Using `filename=<jupyter notebook file>` in the `new_model_server` we parse the given Jupyter Notebook and build our model server from it.
> All the annotations given in the notebook will be parsed and saved to the function normally
The model server will deploy the model given under `models={<model_name>:<model_file_path>}` as `model_class=<model_class_name>` .
Just like any other MLRun function we can set our environment variables, workers and add mounts.
The model server will provide us with a `/<model_name>/predict` endpoint where we can query the model.
```
# inference function
inference_function = new_model_server('tf-images-server',
filename='./nuclio-serving-tf-images.ipynb',
model_class='TFModel')
inference_function.with_http(workers=2)
inference_function.apply(mount_v3io())
```
## Create and run the pipeline
In this part we define the Kubeflow Pipeline to run our process.
MLRun helps us doing that by requiring us to only add `<fn>.as_step()` in order to turn our functions to a pipeline step for kubeflow. All the parameters and inputs can be then set regularly and will be deployed as defined in the pipeline.
The pipeline order is defined by the following:
* We can specify `<fn>.after(<previous fn>)`
* We can specify that a function has a parameter or input, taken from a previous function.
Ex: `models={'cat_vs_dog_v1': train.outputs['model']}` in the inference function definition, taking the model file from the training function.
Notice that you need to `log_artifact` in your function and write it's name in the function's `outputs` parameter to expose it to the pipeline for later use.
```
import kfp
from kfp import dsl
artifacts_path = 'v3io:///users/admin/mlrun/kfp/{{workflow.uid}}/'
@dsl.pipeline(
name='Image classification training pipeline',
description='Shows how to use mlrun with horovod.'
)
def hvd_pipeline(
image_archive= 'http://iguazio-sample-data.s3.amazonaws.com/catsndogs.zip',
images_path = '/User/mlrun/examples/images',
source_dir= '/User/mlrun/examples/images/cats_n_dogs',
checkpoints_dir= '/User/mlrun/examples/checkpoints',
model_path= '/User/mlrun/examples/models/cats_n_dogs.h5',
model_name= 'cat_vs_dog_v1'
):
open_archive = utilsfn.as_step(name='download',
handler='open_archive',
out_path=images_path,
params={'target_dir': images_path},
inputs={'archive_url': image_archive},
outputs=['content'])
label = utilsfn.as_step(name='label',
handler='categories_map_builder',
out_path=images_path,
params={'source_dir': source_dir},
outputs=['categories_map',
'file_categories']).after(open_archive)
train = trainer_fn.as_step(name='train',
params={'epochs': 1,
'checkpoints_dir': checkpoints_dir,
'model_path': model_path,
'data_path': source_dir},
inputs={
'categories_map': label.outputs['categories_map'],
'file_categories': label.outputs['file_categories']},
outputs=['model']).apply(v3io_cred())
# deploy the model using nuclio functions
deploy = inference_function.deploy_step(project='nuclio-serving',
models={model_name: train.outputs['model']})
# for debug generate the pipeline dsl
kfp.compiler.Compiler().compile(hvd_pipeline, 'hvd_pipeline.yaml')
client = kfp.Client(namespace='default-tenant')
arguments = {}
run_result = client.create_run_from_pipeline_func(hvd_pipeline, arguments, experiment_name='horovod1')
# connect to the run db
db = get_run_db().connect()
# query the DB with filter on workflow ID (only show this workflow)
db.list_runs('', labels=f'workflow={run_result.run_id}').show()
```
| github_jupyter |
# Métodos Numéricos Aplicados à Transferência de Calor
## Introdução
O normal do nosso fluxo de trabalho é iniciar importando as bibliotecas que vamos utilizar no decorrer do material.
Um maior detalhamento sobre elas já foi feito na aula anterior, de modo que agora podemos utilizar diretamente:
```
import handcalcs.render
import matplotlib.pyplot as plt
import numpy as np
import plotly.express as px
import plotly.figure_factory as ff
import plotly.graph_objects as go
import plotly.io as pio
from tqdm.notebook import tqdm
```
O bloco a seguir é opcional, ele vai alterar o estilo padrão de nossas figuras, e aumentar um pouco o seu tamanho, melhorando a apresentação em nossa aula:
```
# Definindo um novo estilo para as figuras [opcional]
# Esse bloco modifica alguns dos valores padrões para
plt.rcdefaults()
# https://matplotlib.org/3.1.0/gallery/style_sheets/style_sheets_reference.html
plt.style.use("ggplot")
# https://matplotlib.org/3.1.1/tutorials/introductory/customizing.html
plt.rcParams.update({"figure.dpi": 100, "figure.figsize": (6, 6)})
px.defaults.template = "ggplot2"
px.defaults.height = 600
pio.templates.default = "ggplot2"
```
## Exercícios Resolvidos
### Convecção e difusão transiente unidimensional
Resolver a EDP:
\begin{equation}
\dfrac{\partial T}{\partial t} = \alpha \dfrac{\partial^2 T}{\partial x^2} - u\dfrac{\partial T}{\partial x}, \quad 0\leq x \leq 1 ; 0\leq t \leq 8
\end{equation}
Condições de contorno:
\begin{equation}
T(0,t)=T(1,t)=0
\end{equation}
Condição inicial:
\begin{equation}
T(x,0) = 1 - ( 10 x - 1 )^2 \quad \text{ se $0 \leq x \leq 0,2$}, \quad \text{ senão } T(x,0) = 0
\end{equation}
Discretizando com as derivadas espaciais numa representação por diferença central e a derivada temporal com diferença ascendente:
\begin{equation}
\dfrac{T_{i,n+1}-T_{i,n}}{\Delta t}=\alpha \dfrac{T_{i-1,n}-2T_{i,n}+T_{i+1,n}}{(\Delta x)^2} -u\dfrac{T_{i+1,n}-T_{i-1,n}}{2\Delta x}, \quad 1 \le i \le I - 2, \quad n > 0,
\end{equation}
\begin{equation}
T_{i=0,n} = 0,
\end{equation}
\begin{equation}
T_{i=I-1,n} = 0.
\end{equation}
Agora devemos isolar a incógnita do nosso problema: o termo $T_{i,n+1}$. Perceba que todos os termos à direita são conhecidos, e usamos essa informação para avançar progressivamente no tempo:
\begin{equation}
T_{i,n+1} = T_{i,n} + \Delta t \left( \alpha \dfrac{T_{i-1,n}-2T_{i,n}+T_{i+1,n}}{(\Delta x)^2} -u\dfrac{T_{i+1,n}-T_{i-1,n}}{2\Delta x} \right), \quad 1 \le i \le I - 2, \quad n > 0,
\end{equation}
Veja como podemos programar a solução para o problema, e note, principalmente, como ela se assemelha muito com a notação discretizada da equação acima. Aqui está o código:
```
def equação_exemplo_1(coord_x, coord_t, alpha, u):
# Condição inicial
T = np.zeros(shape = (coord_x.size, coord_t.size))
for i, x in enumerate(coord_x):
if x <= 0.2:
T[i] = 1. - (10. * x - 1) ** 2.
# Condições de contorno
T[0, :] = 0.0
T[-1, :] = 0.0
# Passo de tempo e resolução da malha
dt = coord_t[1] - coord_t[0]
dx = coord_x[1] - coord_x[0]
# Aqui resolve-se a equação
for n in tqdm(range(0, coord_t.size - 1)):
for i in range(1, coord_x.size - 1):
T[i, n + 1] = T[i, n] + dt * (
alpha * (T[i - 1, n] - 2.0 * T[i, n] + T[i + 1, n]) / dx**2.0
- u * (T[i + 1, n] - T[i - 1, n]) / (2.0 * dx)
)
return T
```
Hora de executar o cálculo utilizando a função que definimos no bloco anterior:
```
coord_x = np.linspace(0., 1., num=101)
coord_t = np.linspace(0., 8., num=2001)
T = equação_exemplo_1(
coord_x,
coord_t,
alpha = 0.001,
u = 0.08
)
```
Por fim, exemplificamos a exibição dos resultados com `plotly`:
```
fig = go.Figure()
for n in range(0, coord_t.size, 500):
fig.add_trace(
go.Scatter(
x=coord_x, y=T[:,n],
mode='lines',
name=f't={coord_t[n]}'
)
)
fig.update_layout(
title='Convecção e difusão transiente',
xaxis_title='x',
yaxis_title='Temperatura'
)
fig.show()
fig = px.imshow(
T.T,
x = coord_x,
y = coord_t,
color_continuous_scale = 'RdBu_r',
title = "Temperatura",
labels = dict(x = "x", y = "tempo"),
aspect = "auto",
origin = "lower"
)
fig.show()
```
### Escoamento em Cavidade com Transferência de Calor
Aqui está o sistema de equações diferenciais: a equação da continuidade, duas equações para os componentes de velocidade $u,v$ e uma equação para a temperatura $\Theta$:
$$ \frac{\partial u}{\partial x} + \frac{\partial v}{\partial y} = 0 $$
$$\frac{\partial u}{\partial t}+u\frac{\partial u}{\partial x}+v\frac{\partial u}{\partial y} = - \frac{\partial p}{\partial x}+ \frac{1}{Re} \left(\frac{\partial^2 u}{\partial x^2}+\frac{\partial^2 u}{\partial y^2} \right) $$
$$\frac{\partial v}{\partial t}+u\frac{\partial v}{\partial x}+v\frac{\partial v}{\partial y} = - \frac{\partial p}{\partial y}+ \frac{1}{Re} \left(\frac{\partial^2 v}{\partial x^2}+\frac{\partial^2 v}{\partial y^2}\right) + Ri ~ \Theta $$
$$ \frac{\partial \Theta}{\partial t} + u\frac{\partial \Theta}{\partial x} + v\frac{\partial \Theta}{\partial y} = \frac{1}{Re ~ Pr} \left(\frac{\partial^2 \Theta}{\partial x^2}+\frac{\partial^2 \Theta}{\partial y^2} \right) $$
* Equações discretas:
Primeiro, vamos discretizar a equação do momento para $u$, da seguinte maneira:
$$
\begin{split}
& \frac{u_{i,j}^{n+1}-u_{i,j}^{n}}{\Delta t}+u_{i,j}^{n}\frac{u_{i+1,j}^{n}-u_{i-1,j}^{n}}{2 \Delta x}+v_{i,j}^{n}\frac{u_{i,j+1}^{n}-u_{i,j-1}^{n}}{2\Delta y} = \\
& \qquad -\frac{p_{i+1,j}^{n}-p_{i-1,j}^{n}}{2\Delta x}+\frac{1}{Re}\left(\frac{u_{i+1,j}^{n}-2u_{i,j}^{n}+u_{i-1,j}^{n}}{\Delta x^2}+\frac{u_{i,j+1}^{n}-2u_{i,j}^{n}+u_{i,j-1}^{n}}{\Delta y^2}\right)
\end{split}
$$
Da mesma forma para a equação do momento para $v$:
$$
\begin{split}
&\frac{v_{i,j}^{n+1}-v_{i,j}^{n}}{\Delta t}+u_{i,j}^{n}\frac{v_{i+1,j}^{n}-v_{i-1,j}^{n}}{2\Delta x}+v_{i,j}^{n}\frac{v_{i,j+1}^{n}-v_{i,j-1}^{n}}{2\Delta y} = \\
& \qquad - \frac{p_{i,j+1}^{n}-p_{i,j-1}^{n}}{2\Delta y}
+\frac{1}{Re}\left(\frac{v_{i+1,j}^{n}-2v_{i,j}^{n}+v_{i-1,j}^{n}}{\Delta x^2}+\frac{v_{i,j+1}^{n}-2v_{i,j}^{n}+v_{i,j-1}^{n}}{\Delta y^2}\right) + Ri ~ \Theta_{i,j}
\end{split}
$$
A obtenção da equação de pressão-Poisson discretizada está fora do escopo dessa aula, por ser parte da disciplina de mecânica dos fluidos computacional, mais detalhes podem ser obtidos nos passos 10 e 11 do curso [CFD com Python](https://github.com/fschuch/CFDPython-BR). De qualquer maneira, a equação pode ser escrita assim:
$$
\begin{split}
& \frac{p_{i+1,j}^{n}-2p_{i,j}^{n}+p_{i-1,j}^{n}}{\Delta x^2}+\frac{p_{i,j+1}^{n}-2p_{i,j}^{n}+p_{i,j-1}^{n}}{\Delta y^2} = \\
& \qquad \left[ \frac{1}{\Delta t}\left(\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}+\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right) -\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x} - 2\frac{u_{i,j+1}-u_{i,j-1}}{2\Delta y}\frac{v_{i+1,j}-v_{i-1,j}}{2\Delta x} - \frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right]
\end{split}
$$
Por fim, a equação para temperatura:
$$
\begin{split}
&\frac{\Theta_{i,j}^{n+1}-\Theta_{i,j}^{n}}{\Delta t}+u_{i,j}^{n}\frac{\Theta_{i+1,j}^{n}-\Theta_{i-1,j}^{n}}{2\Delta x}+v_{i,j}^{n}\frac{\Theta_{i,j+1}^{n}-\Theta_{i,j-1}^{n}}{2\Delta y} = \\
& \qquad + \frac{1}{Re ~ Pr}\left(\frac{\Theta_{i+1,j}^{n}-2\Theta_{i,j}^{n}+\Theta_{i-1,j}^{n}}{\Delta x^2}+\frac{\Theta_{i,j+1}^{n}-2\Theta_{i,j}^{n}+\Theta_{i,j-1}^{n}}{\Delta y^2}\right)
\end{split}
$$
**Exercitando:** Você pode escrever essas equações em suas próprias anotações, manualmente, seguindo cada termo mentalmente à medida que as escreve.
Como antes, vamos reorganizar as equações da maneira que as iterações devem proceder no código. Primeiro, as equações de momento para a velocidade no passo de tempo subsequente.
A equação do momento na direção de $u$:
$$
\begin{split}
u_{i,j}^{n+1} = u_{i,j}^{n} & - u_{i,j}^{n} \frac{\Delta t}{2\Delta x} \left(u_{i+1,j}^{n}-u_{i-1,j}^{n}\right) - v_{i,j}^{n} \frac{\Delta t}{2\Delta y} \left(u_{i,j+1}^{n}-u_{i,j-1}^{n}\right) \\
& - \frac{\Delta t}{2\Delta x} \left(p_{i+1,j}^{n}-p_{i-1,j}^{n}\right) \\
& + \frac{1}{Re} \left(\frac{\Delta t}{\Delta x^2} \left(u_{i+1,j}^{n}-2u_{i,j}^{n}+u_{i-1,j}^{n}\right) + \frac{\Delta t}{\Delta y^2} \left(u_{i,j+1}^{n}-2u_{i,j}^{n}+u_{i,j-1}^{n}\right)\right)
\end{split}
$$
```
def cavidade_u(u, v, dt, dx, dy, p, re):
return (
u[1:-1, 1:-1]
- u[1:-1, 1:-1] * dt / (2*dx) * (u[2:, 1:-1] - u[0:-2, 1:-1])
- v[1:-1, 1:-1] * dt / (2*dy) * (u[1:-1, 2:] - u[1:-1, 0:-2])
- dt / (2 * dx) * (p[2:, 1:-1] - p[0:-2, 1:-1])
+ (1.0 / re)
* (
dt / dx ** 2 * (u[2:, 1:-1] - 2 * u[1:-1, 1:-1] + u[0:-2, 1:-1])
+ dt / dy ** 2 * (u[1:-1, 2:] - 2 * u[1:-1, 1:-1] + u[1:-1, 0:-2])
)
)
```
A equação do momento na direção de $v$:
$$
\begin{split}
v_{i,j}^{n+1} = v_{i,j}^{n} & - u_{i,j}^{n} \frac{\Delta t}{2\Delta x} \left(v_{i+1,j}^{n}-v_{i-1,j}^{n}\right) - v_{i,j}^{n} \frac{\Delta t}{2\Delta y} \left(v_{i,j+1}^{n}-v_{i,j-1}^{n})\right) \\
& - \frac{\Delta t}{2\Delta y} \left(p_{i,j+1}^{n}-p_{i,j-1}^{n}\right) \\
& + \frac{1}{Re} \left(\frac{\Delta t}{\Delta x^2} \left(v_{i+1,j}^{n}-2v_{i,j}^{n}+v_{i-1,j}^{n}\right) + \frac{\Delta t}{\Delta y^2} \left(v_{i,j+1}^{n}-2v_{i,j}^{n}+v_{i,j-1}^{n}\right)\right) + \Delta t ~ Ri ~ \Theta_{i,j}
\end{split}
$$
```
def cavidade_v(u, v, theta, dt, dx, dy, p, re, ri):
return (
v[1:-1, 1:-1]
- u[1:-1, 1:-1] * dt / (2*dx) * (v[2:, 1:-1] - v[0:-2, 1:-1])
- v[1:-1, 1:-1] * dt / (2*dy) * (v[1:-1, 2:] - v[1:-1, 0:-2])
- dt / (2 * dy) * (p[1:-1, 2:] - p[1:-1, 0:-2])
+ (1.0 / re)
* (
dt / dx ** 2 * (v[2:, 1:-1] - 2 * v[1:-1, 1:-1] + v[0:-2, 1:-1])
+ dt / dy ** 2 * (v[1:-1, 2:] - 2 * v[1:-1, 1:-1] + v[1:-1, 0:-2])
)
) + ri * dt * theta[1:-1, 1:-1]
```
Agora, reorganizamos a equação de pressão-Poisson:
$$
\begin{split}
p_{i,j}^{n} = & \frac{\left(p_{i+1,j}^{n}+p_{i-1,j}^{n}\right) \Delta y^2 + \left(p_{i,j+1}^{n}+p_{i,j-1}^{n}\right) \Delta x^2}{2\left(\Delta x^2+\Delta y^2\right)} \\
& -\frac{\Delta x^2\Delta y^2}{2\left(\Delta x^2+\Delta y^2\right)} \\
& \times \left[\frac{1}{\Delta t}\left(\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}+\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right)-\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x}\frac{u_{i+1,j}-u_{i-1,j}}{2\Delta x} -2\frac{u_{i,j+1}-u_{i,j-1}}{2\Delta y}\frac{v_{i+1,j}-v_{i-1,j}}{2\Delta x}-\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\frac{v_{i,j+1}-v_{i,j-1}}{2\Delta y}\right]
\end{split}
$$
```
def build_up_b(u, v, dt, dx, dy):
"""Constrói o termo entre chaves na equação acima"""
b = np.zeros_like(u)
b[1:-1, 1:-1] = (
1
/ dt
* (
(u[2:, 1:-1] - u[0:-2, 1:-1]) / (2 * dx)
+ (v[1:-1, 2:] - v[1:-1, 0:-2]) / (2 * dy)
)
- ((u[2:, 1:-1] - u[0:-2, 1:-1]) / (2 * dx)) ** 2
- 2
* (
(u[1:-1, 2:] - u[1:-1, 0:-2])
/ (2 * dy)
* (v[2:, 1:-1] - v[0:-2, 1:-1])
/ (2 * dx)
)
- ((v[1:-1, 2:] - v[1:-1, 0:-2]) / (2 * dy)) ** 2
)
return b
def pressure_poisson(u, v, dt, dx, dy, p, nit=100):
b = build_up_b(u, v, dt, dx, dy)
for q in range(nit):
pn = p.copy()
p[1:-1, 1:-1] = (
(
(pn[2:, 1:-1] + pn[0:-2, 1:-1]) * dy ** 2
+ (pn[1:-1, 2:] + pn[1:-1, 0:-2]) * dx ** 2
)
/ (2 * (dx ** 2 + dy ** 2))
- dx ** 2 * dy ** 2 / (2 * (dx ** 2 + dy ** 2)) * b[1:-1, 1:-1]
)
p[-1, :] = p[-2, :] # dp/dx = 0 at x = 2
p[:, 0] = p[:, 1] # dp/dy = 0 at y = 0
p[0, :] = p[1, :] # dp/dx = 0 at x = 0
p[:, -1] = 0 # p = 0 at y = 2
return p
```
Quase lá! Só falta a equação para temperatura:
$$
\begin{split}
\Theta_{i,j}^{n+1} = \Theta_{i,j}^{n} & - u_{i,j}^{n} \frac{\Delta t}{2\Delta x} \left(\Theta_{i+1,j}^{n}-\Theta_{i-1,j}^{n}\right) - v_{i,j}^{n} \frac{\Delta t}{2\Delta y} \left(\Theta_{i,j+1}^{n}-\Theta_{i,j-1}^{n})\right) \\
& + \frac{1}{Re ~ Pr} \left(\frac{\Delta t}{\Delta x^2} \left(\Theta_{i+1,j}^{n}-2\Theta_{i,j}^{n}+\Theta_{i-1,j}^{n}\right) + \frac{\Delta t}{\Delta y^2} \left(\Theta_{i,j+1}^{n}-2\Theta_{i,j}^{n}+\Theta_{i,j-1}^{n}\right)\right)
\end{split}
$$
```
def cavidade_theta(u, v, theta, dt, dx, dy, p, re, pr):
return (
theta[1:-1, 1:-1]
- u[1:-1, 1:-1] * dt / (2*dx) * (theta[2:, 1:-1] - theta[0:-2, 1:-1])
- v[1:-1, 1:-1] * dt / (2*dy) * (theta[1:-1, 2:] - theta[1:-1, 0:-2])
+ (1.0 / re / pr)
* (
dt / dx ** 2 * (theta[2:, 1:-1] - 2 * theta[1:-1, 1:-1] + theta[0:-2, 1:-1])
+ dt
/ dy ** 2
* (theta[1:-1, 2:] - 2 * theta[1:-1, 1:-1] + theta[1:-1, 0:-2])
)
)
```
A condição inicial é $u, v, p, \Theta = 0 $ em todos os lugares, e as condições de contorno são:
$u=1$ e $\Theta=0$ em $y=2$ (a "tampa");
$u, v = 0$ e $\Theta=1$ nas fronteiras restantes;
$\frac{\partial p}{\partial y}=0$ em $y=0$;
$p=0$ em $y=2$
$\frac{\partial p}{\partial x}=0$ em $x=0$ e $x=2$
Agora todos juntos em uma nova função para efetivamente resolver o escoamento em uma cavidade:
```
def cavidade(x, y, t, re, ri, pr):
# Condição inicial
u = np.zeros((x.size, y.size))
v = np.zeros((x.size, y.size))
p = np.zeros((x.size, y.size))
theta = np.zeros((x.size, y.size))
# Passo de tempo e resolução da malha
dt = t[1] - t[0]
dx = x[1] - x[0]
dy = y[1] - y[0]
# Laço temporal
for n in tqdm(range(t.size)):
un = u.copy()
vn = v.copy()
thetan = theta.copy()
p = pressure_poisson(un, vn, dt, dx, dy, p)
u[1:-1, 1:-1] = cavidade_u(un, vn, dt, dx, dy, p, re)
v[1:-1, 1:-1] = cavidade_v(un, vn, thetan, dt, dx, dy, p, re, ri)
theta[1:-1, 1:-1] = cavidade_theta(un, vn, thetan, dt, dx, dy, p, re, pr)
# Condições de contorno
u[:, 0] = 0
u[0, :] = 0
u[-1, :] = 0
u[:, -1] = 1 # Definir velocidade na tampa da cavidade igual a 1
v[:, 0] = 0
v[:, -1] = 0
v[0, :] = 0
v[-1, :] = 0
theta[-1, :] = 1
theta[:, 0] = 1
theta[0, :] = 1
theta[:, -1] = 0 # Definir theta na tampa da cavidade igual a 1
return u, v, theta, p
```
Hora da ação:
```
# Coordenadas
x = np.linspace(start = 0.0, stop = 2.0, num=21)
y = np.linspace(start = 0.0, stop = 2.0, num=21)
t = np.arange(start = 0.0, stop = 5.0, step = 0.001)
u, v, theta, p = cavidade(x, y, t, re = 40.0, ri = 0.0, pr = 1.0)
```
Verificando os resultados:
```
fig = px.imshow(
theta.T,
x=x,
y=y,
color_continuous_scale="RdBu_r",
title="Temperatura",
labels=dict(x="x", y="y"),
origin="lower",
)
fig.show()
fig = ff.create_streamline(
x[1:-1], y[1:-1], u.T[1:-1, 1:-1], v.T[1:-1, 1:-1], marker_color="black",
)
fig.add_trace(
go.Contour(
z=theta.T,
x=x,
y=y,
colorscale="RdBu_r",
colorbar=dict(title="Temperatura", titleside="right"),
)
)
fig.update_layout(
title="Escoamento em Cavidade com Transferência de Calor",
xaxis_title="x",
yaxis_title="y",
autosize=False,
width=800,
height=800,
)
fig.show()
```
> Leitura recomendada:
> * Gostaria de se apronfundar mais no assunto? Veja a aula [Métodos Numéricos com Python](https://github.com/fschuch/metodos-numericos-com-python), que inclui a solução do problema da cavidade com transferência de calor utilizando conceitos Python mais avançados, como a criação e manipulação de classes em programação orientada ao objeto (OOP).
-----
> **Felipe N. Schuch**,<br>
> Pesquisador em Fluidodinâmica Computacional na PUCRS, com interesse em: Escoamentos turbulentos, transferência de calor e massa, e interação fluido-estrutura; Processamento e visualização de dados em Python; Jupyter Notebook como uma ferramenta de colaboração, pesquisa e ensino.<br>
> [felipeschuch@outlook.com](mailto:felipeschuch@outlook.com "Email") [@fschuch](https://twitter.com/fschuch "Twitter") [Aprenda.py](https://fschuch.github.io/aprenda.py "Blog") [@aprenda.py](https://www.instagram.com/aprenda.py/ "Instagram")<br>
-----
| github_jupyter |
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
df=pd.read_excel('Chapter5_HR_DataSet.xlsx')
df.describe()
df.columns
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df.drop('Quitting',axis=1))
x_standardized_features = scaler.transform(df.drop('Quitting',axis=1))
x_standardized_features
x_standardized_features = pd.DataFrame(x_standardized_features,columns=df.columns[:-1])
x_standardized_features.head()
y=df['Quitting']
y
from sklearn.model_selection import train_test_split
```
# Cross validation
```
seed= 1000
np.random.seed(seed)
X_train, X_test, y_train, y_test = train_test_split(x_standardized_features,y, test_size=0.30)
len(X_train)
```
# Applying KNN model
```
from sklearn.neighbors import KNeighborsClassifier
np.random.seed(seed)
KNN = KNeighborsClassifier(n_neighbors=1, metric='euclidean')
KNN.fit(X_train,y_train)
pred = KNN.predict(X_test)
```
# Prediction and evaluation
```
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,pred))
print(classification_report(y_test,pred))
```
# Choosing K
```
error_rate = []
for i in range(1,50):
KNN = KNeighborsClassifier(n_neighbors=i,metric='euclidean')
KNN.fit(X_train,y_train)
pred_i = KNN.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
plt.figure(figsize=(10,6))
plt.plot(range(1,50),error_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
KNN = KNeighborsClassifier(n_neighbors=1,metric='euclidean')
KNN.fit(X_train,y_train)
pred = KNN.predict(X_test)
print('WITH K=1')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
from sklearn.metrics import classification_report,confusion_matrix
np.random.seed(seed)
KNN = KNeighborsClassifier(n_neighbors=5,metric='euclidean')
KNN.fit(X_train,y_train)
y_pred = KNN.predict(X_test)
print('\n')
print(confusion_matrix(y_test,y_pred))
print('\n')
print(classification_report(y_test,y_pred))
from sklearn import metrics
Scores = []
for k in range(1, 51):
KNN = KNeighborsClassifier(n_neighbors=k,metric='euclidean')
KNN.fit(X_train, y_train)
y_pred = KNN.predict(X_test)
Scores.append(metrics.accuracy_score(y_test, y_pred))
Scores
plt.figure(figsize=(10,8))
plt.plot(range(1, 51), Scores)
plt.xlabel('K Values')
plt.ylabel('Testing Accuracy')
plt.title('K Determination Using KNN', fontsize=20)
```
| github_jupyter |
# Introduction
Welcome to the Eureka! tutorial - today we will learn how to run Eureka!'s S3 data reduction module, which takes 2D JWST data and reduces it to 1D spectra.
Eureka! is an open-source python package available for download at https://github.com/kevin218/Eureka (lead developers are Sebastian Zieba, Giannina Guzman Caloca, Kevin Stevenson, and Laura Kreidberg). Check out the docs at https://eurekadocs.readthedocs.io/en/latest/. Additional contribution are welcome! check out the issues page on github.
**One word of caution is that the package is under heavy development, so this tutorial is based on a stable release branch (release/v0.1). If you are using the ers-transit environment, it installs the stable release branch for you.** If you work with more recent versions of the code, it may not be backward compatible with this tutorial.
## Goals
- walk through all the major steps in data reduction
- get comfortable with the Eureka! structure and syntax
- most importantly, make sure none of the steps are a black box.
## Import standard python packages and Eureka!
```
import sys, os, time
import numpy as np
import matplotlib.pyplot as plt
from importlib import reload
import eureka.S3_data_reduction.s3_reduce as s3
from eureka.lib import readECF as rd
from eureka.lib import logedit
from eureka.lib import readECF as rd
from eureka.lib import manageevent as me
from eureka.S3_data_reduction import optspex
from eureka.lib import astropytable
from eureka.lib import util
from eureka.S3_data_reduction import plots_s3
```
### Step 0: Initialization
```
# Starts timer to monitor how long data reduction takes
t0 = time.time()
# Names the event (has to match the event name used for the *.ecf files)
eventlabel = 'wasp43b'
# Initialize metadata object to store all extra information
# related to the event and the data reduction
meta = s3.Metadata()
meta.eventlabel = eventlabel
# Initialize data object to store data from the observation
dat = s3.Data()
```
Try printing how much time has passed since the timer was initialized. Run the cell again. Do you see the time change?
```
print(time.time() - t0) #time elapsed since the timer start
# Load Eureka! control file and store values in Metadata object
ecffile = 'S3_' + eventlabel + '.ecf'
ecf = rd.read_ecf(ecffile)
rd.store_ecf(meta, ecf)
```
Information from the ECF ("Eureka control file") is now stored in a Metadata object. This includes all the high level information about the data reduction (which JWST instrument was used? do we want to display plots? where is the data stored? what size is the extraction window? etc.)
To see the current contents of the Metadata object, type ``meta.__dict__.keys``.
What is the value of ``meta.bg_deg``? Can you change it?
### Step 1: Make directories to store reduced data, create log file, read in data
```
# Create directories for Stage 3 processing
datetime= time.strftime('%Y-%m-%d_%H-%M-%S')
meta.workdir = 'S3_' + datetime + '_' + meta.eventlabel
if not os.path.exists(meta.workdir):
os.makedirs(meta.workdir)
if not os.path.exists(meta.workdir+"/figs"):
os.makedirs(meta.workdir+"/figs")
# Load instrument module
exec('from eureka.S3_data_reduction import ' + meta.inst + ' as inst', globals())
reload(inst)
# Open new log file
meta.logname = './'+meta.workdir + '/S3_' + meta.eventlabel + ".log"
log = logedit.Logedit(meta.logname)
log.writelog("\nStarting Stage 3 Reduction")
# Create list of file segments
meta = util.readfiles(meta)
num_data_files = len(meta.segment_list)
log.writelog(f'\nFound {num_data_files} data file(s) ending in {meta.suffix}.fits')
stdspec = np.array([])
```
*Important check!* Were the correct files read in? They are stored in ``meta.segment_list``.
### Step 2: read the data (and look at it!)
```
# pick a single file to read and reduce as a test
m = 17
# Read in data frame and header
log.writelog(f'Reading file {m+1} of {num_data_files}')
dat = inst.read(meta.segment_list[m], dat, returnHdr=True)
```
#### What data are we using?
The full description of the data is available [here](https://stsci.app.box.com/s/8r6kqh9m53jkwkff0scmed6zx42g307e/file/804595804746)). To quickly summarize, we are using simulated NIRCam grism time series data from the ERS Simulated Spectra Team. The simulation assumes a WASP-43 b-like planet with physically realistic spectral features added. The simulated data are based on the following observational design:
- GRISMR+F322W2 pupil and filter
- RAPID readout mode
- 19 Groups per integrations
- 1287 integrations
- 1 Exposure
- 4 Output amplifiers
The data themselves are divided into “segments,” with each individual segment (seg001, seg002, etc.) containing a subset of the overall dataset. This is how flight data will be delivered. The segments are numbered in their order of observation.
We will use the Stage 2 Output from the [JWST data reduction pipeline](https://jwst-pipeline.readthedocs.io/en/latest/jwst/data_products/stages.html). For NIRCam, Stage 2 consists of the flat field correction, WCS/wavelength solution, and photometric calibration (counts/sec -> MJy). Note that this is specifically for NIRCam: the steps in Stage 2 change a bit depending on the instrument. The Stage 2 outputs are rougly equivalent to a “flt” file from HST.
The files have the suffix ``/*calints.fits`` which contain fully calibrated images (MJy) for each individual integration. This is the one you want if you’re starting with Stage 2 and want to do your own spectral extraction.
### Let's take a look at the data!
What is stored in the data object?
```
print(dat.__dict__.keys())
```
The calibrated 2D data, error array, data quality are stored in `data`, `err`, and `dq`. `wave` is the wavelength.
The header information is stored in mhdr (main header) and shdr (science header). Use the headers to check whether the data is really from NIRCam.
```
dat.mhdr['INSTRUME']
```
What units are the data stored in?
```
dat.shdr['BUNIT']
```
#### What does the data look like??
```
plt.imshow(dat.data[0], origin = 'lower', aspect='auto', vmin=0, vmax=2e6)
ax = plt.gca()
plt.colorbar(label='Brightness (MJy/sr)') #we will convert to photoelectrons later
ax.set_xlabel('wavelength direction')
ax.set_ylabel('spatial direction')
```
What happens if we change the contrast with the vmax parameter? what is the approximate background level?
```
plt.imshow(dat.data[0], origin = 'lower', aspect='auto', vmin=0, vmax=10000)
ax = plt.gca()
plt.colorbar(label='Brightness (MJy/sr)')
ax.set_xlabel('wavelength direction')
ax.set_ylabel('spatial direction')
```
How big should the extraction window be? Should it be symmetric? (Hint: we want to capture all the flux from the target star, but minimize the background)
Let's plot the spatial profile to see how wide the PSF is.
```
plt.plot(dat.data[0][:,1000]) #plots column 1000
plt.xlabel("Spatial pixel")
plt.ylabel("Flux (MJy/sr)")
```
Flux is mostly concentrated over a few pixels. But the wings are pretty wide! This is easier to see in log space:
```
plt.plot(np.log10(dat.data[0][:,1000])) #plots log10 of column 1000
ind_max = np.argmax(dat.data[0][:,1000]) #finds row where counts peak
plt.axvline(ind_max, color = 'red') #marks peak counts with a red vertical line
plt.xlabel("Spatial pixel")
plt.ylabel("Log10 Flux (MJy/sr)")
```
### Decide which regions we want to use for the background and for the spectrum¶
```
# Get number of integrations and frame dimensions
meta.n_int, meta.ny, meta.nx = dat.data.shape
# Saves source postion (accounting for window size)
meta.src_xpos = dat.shdr['SRCXPOS']-meta.xwindow[0]
meta.src_ypos = dat.shdr['SRCYPOS']-meta.ywindow[0]
```
**Check which extraction window is saved in Metadata**
```
print(meta.xwindow)
print(meta.ywindow)
# Trim data to subarray region of interest
dat, meta = util.trim(dat, meta)
```
### Handle some unit conversion
```
#Convert units (eg. for NIRCam: MJy/sr -> DN -> Electrons)
dat, meta = inst.unit_convert(dat, meta, log)
# Record integration mid-times in BJD_TDB
dat.bjdtdb = dat.int_times['int_mid_BJD_TDB']
```
### Mask bad pixels
```
# Create bad pixel mask
dat.submask = np.ones(dat.subdata.shape)
# Check if arrays have NaNs
dat.submask = util.check_nans(dat.subdata, dat.submask, log)
dat.submask = util.check_nans(dat.suberr, dat.submask, log)
dat.submask = util.check_nans(dat.subv0, dat.submask, log)
# Manually mask regions [colstart, colend, rowstart, rowend]
if hasattr(meta, 'manmask'):
log.writelog(" Masking manually identified bad pixels")
for i in range(len(meta.manmask)):
ind, colstart, colend, rowstart, rowend = meta.manmask[i]
dat.submask[rowstart:rowend,colstart:colend] = 0
# Perform outlier rejection of sky background along time axis
log.writelog('Performing background outlier rejection')
meta.bg_y1 = int(meta.src_ypos - meta.bg_hw)
meta.bg_y2 = int(meta.src_ypos + meta.bg_hw)
dat.submask = inst.flag_bg(dat, meta)
```
### How many bad pixels were masked?
```
ny, nx = dat.submask.shape[1], dat.submask.shape[2]
print(ny, nx)
print(1. - np.sum(dat.submask[5])/(nx*ny)) # fraction of bad pixels in the integration
```
### Subtract the background
```
dat = util.BGsubtraction(dat, meta, log, meta.isplots_S3)
# Plots background-subtracted image (be patient, this can take a second!)
if meta.isplots_S3 >= 3:
for n in range(meta.n_int):
#make image+background plots
plots_s3.image_and_background(dat, meta, n)
# Try doing a linear column-by-column fit to the background instead
meta.bg_deg = 1 #fits a polynomial of degree 1 to each column
dat = util.BGsubtraction(dat, meta, log, meta.isplots_S3)
# Plots background-subtracted image (be patient, this can take a second!)
if meta.isplots_S3 >= 3:
for n in range(meta.n_int):
#make image+background plots
plots_s3.image_and_background(dat, meta, n)
```
### Time to extract the spectrum!!
```
# Select only aperture region
ap_y1 = int(meta.src_ypos - meta.spec_hw)
ap_y2 = int(meta.src_ypos + meta.spec_hw)
dat.apdata = dat.subdata[:,ap_y1:ap_y2]
dat.aperr = dat.suberr [:,ap_y1:ap_y2]
dat.apmask = dat.submask[:,ap_y1:ap_y2]
dat.apbg = dat.subbg [:,ap_y1:ap_y2]
dat.apv0 = dat.subv0 [:,ap_y1:ap_y2]
# Compute median frame
meta.medsubdata = np.median(dat.subdata, axis=0)
meta.medapdata = np.median(dat.apdata, axis=0)
# Extract standard spectrum and its variance
dat.stdspec = np.sum(dat.apdata, axis=1)
dat.stdvar = np.sum(dat.aperr**2, axis=1)
```
What does the standard spectrum look like?
```
plt.plot(dat.subwave[meta.src_ypos], dat.stdspec[0])
```
Now let's do optimal extraction (this de-weights the wings of the spectrum that are background-dominated)
```
# Extract optimal spectrum with uncertainties
log.writelog(" Performing optimal spectral extraction")
dat.optspec = np.zeros((dat.stdspec.shape))
dat.opterr = np.zeros((dat.stdspec.shape))
gain = 1 #FINDME: need to determine correct gain
for n in range(meta.n_int):
dat.optspec[n], dat.opterr[n], mask = optspex.optimize(dat.apdata[n], dat.apmask[n], dat.apbg[n], dat.stdspec[n], gain, dat.apv0[n], p5thresh=meta.p5thresh, p7thresh=meta.p7thresh, fittype=meta.fittype, window_len=meta.window_len, deg=meta.prof_deg, n=dat.intstart+n, isplots=meta.isplots_S3, eventdir=meta.workdir, meddata=meta.medapdata)
# Plotting results
if meta.isplots_S3 >= 3:
for n in range(meta.n_int):
#make optimal spectrum plot
plots_s3.optimal_spectrum(dat, meta, n)
```
### Save results and generate final figures
```
# Append results
if len(stdspec) == 0:
wave_2d = dat.subwave
wave_1d = dat.subwave[meta.src_ypos]
stdspec = dat.stdspec
stdvar = dat.stdvar
optspec = dat.optspec
opterr = dat.opterr
bjdtdb = dat.bjdtdb
else:
stdspec = np.append(stdspec, dat.stdspec, axis=0)
stdvar = np.append(stdvar, dat.stdvar, axis=0)
optspec = np.append(optspec, dat.optspec, axis=0)
opterr = np.append(opterr, dat.opterr, axis=0)
bjdtdb = np.append(bjdtdb, dat.bjdtdb, axis=0)
# Calculate total time
total = (time.time() - t0)/60.
log.writelog('\nTotal time (min): ' + str(np.round(total,2)))
# Save results
log.writelog('Saving results')
me.saveevent(meta, meta.workdir + '/S3_' + meta.eventlabel + "_Meta_Save", save=[])
# Save results
log.writelog('Saving results')
me.saveevent(dat, meta.workdir + '/S3_' + meta.eventlabel + "_Data_Save", save=[])
log.writelog('Saving results as astropy table...')
astropytable.savetable(meta, bjdtdb, wave_1d, stdspec, stdvar, optspec, opterr)
log.writelog('Generating figures')
if meta.isplots_S3 >= 1:
# 2D light curve without drift correction
plots_s3.lc_nodriftcorr(meta, wave_1d, optspec)
log.closelog()
```
# Now let's make lightcurves!
This is Stage 4 of the Eureka! pipeline. It takes the time series of spectra from Stage 3 and bins them into wavelength channels.
```
#importing Eureka! Stage 4 modules
import eureka.S4_generate_lightcurves.s4_genLC as s4
import eureka.S4_generate_lightcurves.plots_s4 as plots_s4
# Load Eureka! control file and store values in Event object
ecffile = 'S4_' + eventlabel + '.ecf'
ecf = rd.read_ecf(ecffile)
rd.store_ecf(meta, ecf)
# Create directories for Stage 3 processing
datetime= time.strftime('%Y-%m-%d_%H-%M-%S')
meta.lcdir = meta.workdir + '/S4_' + datetime + '_' + str(meta.nspecchan) + 'chan'
if not os.path.exists(meta.lcdir):
os.makedirs(meta.lcdir)
if not os.path.exists(meta.lcdir+"/figs"):
os.makedirs(meta.lcdir+"/figs")
# Copy existing S3 log file
meta.s4_logname = './'+meta.lcdir + '/S4_' + meta.eventlabel + ".log"
#shutil.copyfile(ev.logname, ev.s4_logname, follow_symlinks=True)
log = logedit.Logedit(meta.s4_logname, read=meta.logname)
log.writelog("\nStarting Stage 4: Generate Light Curves\n")
```
### Store the 1D optimally extracted spectra, wavelengths, and times
```
table = astropytable.readtable(meta)
optspec, wave_1d, bjdtdb = np.reshape(table['optspec'].data, (-1, meta.subnx)), \
table['wave_1d'].data[0:meta.subnx], table['bjdtdb'].data[::meta.subnx]
#Replace NaNs with zero
optspec[np.where(np.isnan(optspec))] = 0
```
### Specify wavelength bins
```
# Determine wavelength bins
binsize = (meta.wave_max - meta.wave_min)/meta.nspecchan
meta.wave_low = np.round([i for i in np.linspace(meta.wave_min, meta.wave_max-binsize, meta.nspecchan)],3)
meta.wave_hi = np.round([i for i in np.linspace(meta.wave_min+binsize, meta.wave_max, meta.nspecchan)],3)
```
## Make the lightcurves!!!
### Important sanity check: do the error bars on the data points look correct to you ? If not, why not?
```
log.writelog("Generating light curves")
n_int, nx = optspec.shape
meta.lcdata = np.zeros((meta.nspecchan, n_int))
meta.lcerr = np.zeros((meta.nspecchan, n_int))
for i in range(meta.nspecchan):
log.writelog(f"Bandpass {i} = %.3f - %.3f" % (meta.wave_low[i],meta.wave_hi[i]))
# Compute valid indices within wavelength range
index = np.where((wave_1d >= meta.wave_low[i])*(wave_1d <= meta.wave_hi[i]))[0]
# Sum flux for each spectroscopic channel
meta.lcdata[i] = np.sum(optspec[:,index],axis=1)
# Add uncertainties in quadrature
meta.lcerr[i] = np.sqrt(np.sum(optspec[:,index]**2,axis=1))
# Plot each spectroscopic light curve
if meta.isplots_S4 >= 3:
plots_s4.binned_lightcurve(meta, bjdtdb, i)
# Save results
log.writelog('Saving results')
me.saveevent(meta, meta.lcdir + '/S4_' + meta.eventlabel + "_Meta_Save", save=[])
log.closelog()
```
## Final note: the error bars are in fact not correct, due to a bug in the NIRCam error arrays. This is a reminder to always check whether the results from your pipeline make sense!!
| github_jupyter |
<a href="https://colab.research.google.com/github/NeuromatchAcademy/course-content/blob/master/tutorials/W2D3_BiologicalNeuronModels/student/W2D3_Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Intro
**Our 2021 Sponsors, including Presenting Sponsor Facebook Reality Labs**
<p align='center'><img src='https://github.com/NeuromatchAcademy/widgets/blob/master/sponsors.png?raw=True'/></p>
## Overview
Today you will learn about a few interesting properties of biological neurons and synapses. In his intro lecture Upi Bhalla will start with an overview of the complexity of the neurons and synapses in the brain. He will also introduce a mathematical description of action potential generation and propagation by which neurons communicate with each other. Then, in a series of short tutorials Richard Naud will introduce simple neuron and synapse models. These tutorials will give you insights about how neurons may generate irregular spike patterns and synchronize their activity. In the first tutorial you will learn about the input-output transfer function of the leaky integrate and fire neuron model. In the second tutorial you will use this model to understand how statistics of inputs affects transfer of synchrony. In the third tutorial you will explore the short-term dynamics of synapses which means that synaptic weight is dependent on the recent history of spiking activity of the pre-synaptic neurons. In the bonus tutorial, you can learn about spike timing dependent plasticity and explore how synchrony in the input may shape the synaptic weight distribution. Finally, in the outro lecture Yiota Poirazi will explain how the simplified description of neurons can be expanded to include more biological complexity. She will provide evidence of how dendritic morphology may expand the computational repertoire of individual neurons.
The models we use in today’s lecture fall in the category of how models (W1D1). You will use several concepts from linear systems (W2D2). The insights developed in these tutorials will be useful to understand the dynamics of neural networks (W3D4). Moreover, you will learn about the origin of statistics of neuronal activity which will be useful for several tutorials. For example, the understanding of synchrony will be very useful in appreciating the problem of causality (W3D5).
Neuron and synapse models are essential building blocks of mechanistic models of brain function and dysfunction. One of the common questions in neuroscience is to identify the causes of changes in the statistics of spiking activity patterns. Whether these changes are caused by changes in neuron/synapse properties or by a change in the input or by a combination of both? With the contents of this tutorial, you should have a framework to think about which changes in spike patterns are due to neuron/synapse or input changes.
## Video
```
# @markdown
from ipywidgets import widgets
out2 = widgets.Output()
with out2:
from IPython.display import IFrame
class BiliVideo(IFrame):
def __init__(self, id, page=1, width=400, height=300, **kwargs):
self.id=id
src = "https://player.bilibili.com/player.html?bvid={0}&page={1}".format(id, page)
super(BiliVideo, self).__init__(src, width, height, **kwargs)
video = BiliVideo(id=f"BV18A411v7Yy", width=854, height=480, fs=1)
print("Video available at https://www.bilibili.com/video/{0}".format(video.id))
display(video)
out1 = widgets.Output()
with out1:
from IPython.display import YouTubeVideo
video = YouTubeVideo(id=f"MAOOPv3whZ0", width=854, height=480, fs=1, rel=0)
print("Video available at https://youtube.com/watch?v=" + video.id)
display(video)
out = widgets.Tab([out1, out2])
out.set_title(0, 'Youtube')
out.set_title(1, 'Bilibili')
display(out)
```
## Slides
```
# @markdown
from IPython.display import IFrame
IFrame(src=f"https://mfr.ca-1.osf.io/render?url=https://osf.io/gyfr2/?direct%26mode=render%26action=download%26mode=render", width=854, height=480)
```
| github_jupyter |
# Diagrama de Gantt para seguimiento de un proyecto
### Campos del dataset:
- Etapa del proyecto.
- Tarea a realizar.
- Fecha de inicio de la tarea.
- Fecha de fin de la tarea.
- Porcentaje de la tarea que ya se completó.
Se podría agregar un campo con una descripción completa de la tarea a realizar
```
import pandas as pd
df = pd.DataFrame()
df['Etapa'] = ['Concepción e inicio del proyecto', 'Definición y planificación', 'MVP1', 'MVP1', 'MVP2','MVP2', 'Presentación de resultados', 'Cierre del proyecto']
df['Tarea'] = ['task1', 'task2', 'task3', 'task4', 'task5', 'task6', 'task7', 'task8']
df['FechaInicio'] = ['1/01/2021','8/01/2021','15/01/2021','18/01/2021','20/01/2021','27/01/2021','2/02/2021','10/02/2021']
df['FechaFin'] = ['7/01/2021','15/01/2021','21/01/2021','25/01/2021','28/01/2021','2/02/2021','10/02/2021','12/02/2021']
df['Pct_completo']=[1,1, 0.9, 0.75, 0.6, 0.45, 0.3, 0.2]
df
```
- Le doy formato datetime a las fechas de inicio y fin de cada tarea.
- Calculo la cantidad de días que me lleva realizar cada tarea.
- Asigno un color para cada una de las etapas del proyecto.
Los colores los busco en: https://colorbrewer2.org/#type=sequential&scheme=BuGn&n=3
```
import matplotlib.pyplot as plt
import numpy as np
from datetime import datetime as dt
df['Start']=[dt.strptime(a, '%d/%m/%Y') for a in df['FechaInicio']]
df['End']=[dt.strptime(a, '%d/%m/%Y') for a in df['FechaFin']]
# project start date
proj_start = df.Start.min()
# number of days from project start to task start
df['start_num'] = (df.Start-proj_start).dt.days
# number of days from project start to end of tasks
df['end_num'] = (df.End-proj_start).dt.days
# days between start and end of each task
df['days_start_to_end'] = df.end_num - df.start_num
# create a column with the color for each project
def color(row):
c_dict = {'Concepción e inicio del proyecto':'#E64646', 'Definición y planificación':'#E69646',
'MVP1':'#34D05C', 'MVP2':'#34D0C3', 'Presentación de resultados':'#3475D0', 'Cierre del proyecto':'#c51b7d'}
return c_dict[row['Etapa']]
df['color'] = df.apply(color, axis=1)
# days between start and current progression of each task
df['current_num'] = (df.days_start_to_end * df.Pct_completo)
df
```
### Gráfico para el seguimiento del proyecto
```
from matplotlib.patches import Patch
fig, ax = plt.subplots(1, figsize=(16,4))
# bars
ax.barh(df.Tarea, df.current_num, left=df.start_num, color=df.color)
ax.barh(df.Tarea, df.days_start_to_end, left=df.start_num, color=df.color, alpha=0.5)
# texts
for idx, row in df.iterrows():
ax.text(row.end_num+0.2, idx,
f"{int(row.Pct_completo*100)}%",
va='center', alpha=0.8, fontsize = 12, color='white')
ax.text(row.start_num-2, idx,
f"{row.Tarea}",
va='center', alpha=0.8, fontsize = 10, color='white' )
##### LEGENDS #####
c_dict = {'Concepción e inicio del proyecto':'#E64646', 'Definición y planificación':'#E69646',
'MVP1':'#34D05C', 'MVP2':'#34D0C3', 'Presentación de resultados':'#3475D0', 'Cierre del proyecto':'#c51b7d'}
legend_elements = [Patch(facecolor=c_dict[i], label=i) for i in c_dict]
legend = plt.legend(handles=legend_elements, bbox_to_anchor =(0.75, -0.1), ncol = 3)
plt.setp(legend.get_texts(), color='white')
frame = legend.get_frame()
frame.set_color('#023858')
##### TITLE #####
plt.suptitle('\nSeguimiento de proyectos \n',fontsize = 20, fontweight = "bold",verticalalignment= 'baseline',
horizontalalignment='center', color='white')
plt.title(label='"Nombre del proyecto"\n',fontdict = {'fontsize': 16, 'fontweight':"bold", 'verticalalignment': 'baseline',
'horizontalalignment': 'center', 'color':'white'})
##### TICKS #####
# Asigno marcas en el eje x cada 7 días
xticks = np.arange(0, df.end_num.max()+1, 7)
ax.set_xticks(xticks)
# Asigno el formato de fecha que quiero ver para esas marcas
xticks_labels = pd.date_range(proj_start, end=df.End.max()).strftime("%d/%m")
ax.set_xticklabels(xticks_labels[::7])
# Quiero que el gráfico inicie 3 días antes de la fecha de la primer tarea y finalice 3 días después de la fecha de la última tarea.
xticks_minor = np.arange(df.start_num.min()-3, df.end_num.max()+3, 1)
ax.set_xticks(xticks_minor, minor=True)
#especifico el color de fondo del gráfico
fig.set_facecolor('#023858')
#fig.set_alpha(0.6)
ax.set_facecolor('#023858')
# especifico los colores de los ejes
ax.spines['bottom'].set_color('white')
ax.spines['top'].set_color('white')
ax.spines['left'].set_color('#023858')
ax.spines['right'].set_color('#023858')
ax.xaxis.label.set_color('white')
ax.tick_params(axis='x', colors='white')
ax.get_yaxis().set_visible(False) #Elimino los nombres del eje y
#especifico que quiero grillas verticales en un determinado color
ax.grid(axis='x', #solo lineas verticales
color='#045a8d',
linestyle='dotted', linewidth=1) #linea de puntos y tamaño 1)
plt.show()
```
| github_jupyter |
# Advanced Features
When analyzing the real world datasets, we may have the following targets:
1. certain variables must be selected when some prior information is given;
2. selecting the weak signal variables when the prediction performance is mainly interested;
3. identifying predictors when group structure are provided;
4. pre-excluding a part of predictors when datasets have ultra high-dimensional predictors;
5. specify the division of sample in cross validation;
6. specify the initial active set before splicing.
In the following content, we will illustrate the statistic methods to reach these targets in a one-by-one manner, and give quick examples to show how to perform the statistic methods in `abessLm` and the same steps can be implemented in all methods. Actually, in our methods, the targets can be properly handled by simply change some default arguments in the functions.
## Nuisance Regression
Nuisance regression refers to best subset selection with some prior information that some variables are required to stay in the active set. For example, if we are interested in a certain gene and want to find out what other genes are associated with the response when this particular gene shows effect.
In the `abessLm()` (or other methods), the argument `always_select` is designed to realize this goal. User can pass a vector containing the indexes of the target variables to `always_select`. Here is an example.
```
import numpy as np
from abess.datasets import make_glm_data
from abess.linear import abessLm
np.random.seed(0)
# gene data
n = 100
p = 20
k = 5
dt = make_glm_data(n = n, p = p, k = k, family = 'gaussian')
print('real coefficients:\n', dt.coef_, '\n')
print('real coefficients\' indexes:\n', np.nonzero(dt.coef_)[0])
model = abessLm(support_size = range(0, 6))
model.fit(dt.x, dt.y)
print('fitted coefficients:\n', model.coef_, '\n')
print('fitted coefficients\' indexes:\n', np.nonzero(model.coef_)[0])
```
The coefficients are located in \[2, 5, 10, 11, 18\].
But if we suppose that the 7th and 8th variables are worthy to be included in the model, we can call:
```
model = abessLm(support_size = range(0, 6), always_select = [7, 8])
model.fit(dt.x, dt.y)
print('fitted coefficients:\n', model.coef_, '\n')
print('fitted coefficients\' indexes:\n', np.nonzero(model.coef_)[0])
```
Now the variables we chosen are always in the model.
## Regularized Adaptive Best Subset Selection
In some cases, especially under low signal-to-noise ratio (SNR) setting or predictors are highly correlated, the vallina type of $L_0$ constrained model may not be satisfying and a more sophisticated trade-off between bias and variance is needed. Under this concern, the `abess` pakcage provides option of best subset selection with $L_2$ norm regularization called the regularized bess. The model has this following form:
$$
\begin{align}
\arg\min_\beta L(\beta) + \alpha \|\beta\|_2^2.
\end{align}
$$
To implement the regularized bess, user need to specify a value to an additive argument `alpha` in the `abessLm()` function (or other methods). This value corresponds to the penalization parameter in the model above.
Let’s test the regularized best subset selection against the no-regularized one over 100 replicas in terms of prediction performance. With argument `snr` in `make_glm_data()`, we can add white noise into generated data.
```
loss = np.zeros((2, 100))
coef = np.repeat([1, 0], [5, 25])
for i in range(100):
np.random.seed(i)
train = make_glm_data(n = 100, p = 30, k = 5, family = 'gaussian', coef_ = coef, snr = 0.05)
np.random.seed(i + 100)
test = make_glm_data(n = 100, p = 30, k = 5, family = 'gaussian', coef_ = coef, snr = 0.05)
# normal
model = abessLm()
model.fit(train.x, train.y)
loss[0, i] = np.linalg.norm(model.predict(test.x) - test.y)
# regularized
model = abessLm(alpha = 0.7)
model.fit(train.x, train.y)
loss[1, i] = np.linalg.norm(model.predict(test.x) - test.y)
print("normal model's loss:", np.mean(loss[0,:]))
print("regularized model's loss:", np.mean(loss[1,:]))
```
The regularized model has a lower test loss. And we can also make a boxplot:
```
import matplotlib.pyplot as plt
plt.boxplot([loss[0,:], loss[1,:]], labels = ['ABESS', 'RABESS'])
plt.show()
```
We see that the regularized best subset select ("RABESS" in figure) indeed reduces the prediction error.
## Best group subset selection
Best group subset selection (BGSS) aims to choose a small part of non-overlapping groups to achieve the best interpretability on the response variable. BGSS is practically useful for the analysis of ubiquitously existing variables with certain group structures. For instance, a categorical variable with several levels is often represented by a group of dummy variables. Besides, in a nonparametric additive model, a continuous component can be represented by a set of basis functions (e.g., a linear combination of spline basis functions). Finally, specific prior knowledge can impose group structures on variables. A typical example is that the genes belonging to the same biological pathway can be considered as a group in the genomic data analysis.
The BGSS can be achieved by solving:
$$
\min_{\beta\in \mathbb{R}^p} \frac{1}{2n} ||y-X\beta||_2^2,\quad s.t.\ ||\beta||_{0,2}\leq s .
$$
where $||\beta||_{0,2} = \sum_{j=1}^J I(||\beta_{G_j}||_2\neq 0)$ in which $||\cdot||_2$ is the $L_2$ norm and model size $s$ is a positive integer to be determined from data. Regardless of the NP-hard of this problem, Zhang et al develop a certifiably polynomial algorithm to solve it. This algorithm is integrated in the `abess` package, and user can handily select best group subset by assigning a proper value to the `group` arguments:
We still use the dataset `dt` generated before, which has 100 samples, 5 useful variables and 15 irrelevant varibales.
```
print('real coefficients:\n', dt.coef_, '\n')
```
Support we have some prior information that every 5 variables as a group:
```
group = np.linspace(0, 3, 4).repeat(5)
print('group index:\n', group)
```
Then we can set the `group` argument in function. Besides, the `support_size` here indicates the number of groups, instead of the number of variables.
```
model = abessLm(support_size = range(0, 3))
model.fit(dt.x, dt.y, group = group)
print('coefficients:\n', model.coef_)
```
The fitted result suggest that only two groups are selected (since `support_size` is from 0 to 2) and the selected variables are shown before.
## Integrate SIS
Ultra-high dimensional predictors increase computational cost but reduce estimation accuracy for any statistical procedure. To reduce dimensionality from high to a relatively acceptable level, a fairly general asymptotic framework, named feature screening (sure independence screening) is proposed to tackle even exponentially growing dimension. The feature screening can theoretically maintain all effective predictors with a high probability, which is called "the sure screening property".
In our program, to carrying out the Integrate SIS, user need to pass an integer smaller than the number of the predictors to the `screening_size`. Then the program will first calculate the marginal likelihood of each predictor and reserve those predictors with the `screening_size` largest marginal likelihood. Then, the ABESS algorithm is conducted only on this screened subset.
Here is an example.
```
n = 100
p = 1000
k = 3
np.random.seed(2)
# gene data
dt = make_glm_data(n = n, p = p, k = k, family = 'gaussian')
print('real coefficients\' indexes:', np.nonzero(dt.coef_)[0])
# fit
model = abessLm(support_size = range(0, 5), screening_size = 100)
model.fit(dt.x, dt.y)
print('fitted coefficients\' indexes:', np.nonzero(model.coef_)[0])
```
## User-specified cross validation division
Sometimes, especially when running a test, we would like to fix the train and valid data used in cross validation, instead of choosing them randomly.
One simple method is to fix a random seed, such as `numpy.random.seed()`. But in some cases, we would also like to specify which samples would be in the same "fold", which has great flexibility.
In our program, an additional argument `cv_fold_id` is for this user-specified cross validation division. An integer array with the same size of input samples can be given, and those with same integer would be assigned to the same "fold" in K-fold CV.
```
n = 100
p = 1000
k = 3
np.random.seed(2)
dt = make_glm_data(n = n, p = p, k = k, family = 'gaussian')
# cv_fold_id has a size of `n`
# cv_fold_id has `cv` different integers
cv_fold_id = [1 for i in range(30)] + [2 for i in range(30)] + [3 for i in range(40)]
model = abessLm(support_size = range(0, 5), cv = 3)
model.fit(dt.x, dt.y, cv_fold_id = cv_fold_id)
print('fitted coefficients\' indexes:', np.nonzero(model.coef_)[0])
```
## User-specified initial active set
We believe that it worth allowing given an initial active set so that the splicing process starts from this set for each sparsity.
It might come from prior analysis, whose result is not quite precise but better than random selection, so the algorithm can run more efficiently. Or you just want to give different initial sets to test the stability of the algorithm.
*Note that this is NOT equal to `always_select`, since they can be exchanged to inactive set when splicing.*
To specify initial active set, an additive argument `A_init` should be given in `fit()`.
```
n = 100
p = 10
k = 3
np.random.seed(2)
dt = make_glm_data(n = n, p = p, k = k, family = 'gaussian')
model = abessLm(support_size = range(0, 5))
model.fit(dt.x, dt.y, A_init = [0, 1, 2])
```
Some strategies for initial active set are:
- If $sparsity = len(A\_init)$, the splicing process would start from $A\_init$.
- If $sparsity > len(A\_init)$, the initial set includes $A\_init$ and other variables `inital screening` chooses.
- If $sparsity < len(A\_init)$, the initial set includes part of $A\_init$.
- If both `A_init` and `always_select` are given, `always_select` first.
- For warm-start, `A_init` will only affect splicing under the first sparsity in `support_size`.
- For CV, `A_init` will affect each fold but not the re-fitting on full data.
## R tutorial
For R tutorial, please view [https://abess-team.github.io/abess/articles/v07-advancedFeatures.html](https://abess-team.github.io/abess/articles/v07-advancedFeatures.html).
| github_jupyter |
# Lesson 4: Data reclassification
- https://kodu.ut.ee/~kmoch/geopython2021/L4/reclassify.html
```
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import os
fp = "corine_tartu.shp"
data = gpd.read_file(fp)
%matplotlib inline
data.head(5)
fp_clc = "corine_legend/clc_legend.csv"
data_legend = pd.read_csv(fp_clc, sep=';', encoding='latin1')
data_legend.head(5)
display(data.dtypes)
display(data_legend.dtypes)
# please don't actually do it right now, it might cause extra troubles later
# data = data.merge(data_legend, how='inner', left_on='code_12', right_on='CLC_CODE')
def change_type(row):
code_as_int = int(row['code_12'])
return code_as_int
data['clc_code_int'] = data.apply(change_type, axis=1)
data.head(2)
data = data.merge(data_legend, how='inner', left_on='clc_code_int', right_on='CLC_CODE', suffixes=('', '_legend'))
selected_cols = ['ID','Remark','Shape_Area','CLC_CODE','LABEL3','RGB','geometry']
# Select data
data = data[selected_cols]
# What are the columns now?
data.columns
# Check coordinate system information
data.crs
data_proj = data.to_crs(epsg=3301)
# Calculate the area of bogs
data_proj['area'] = data_proj.area
# What do we have?
data_proj['area'].head(2)
data_proj.plot(column='CLC_CODE', linewidth=0.05)
print(list(data_proj['CLC_CODE'].unique()))
print(list(data_proj['LABEL3'].unique()))
bogs = data_proj.loc[data['LABEL3'] == 'Peat bogs'].copy()
bogs.head(2)
bogs['area_km2'] = bogs['area'] / 1000000
# What is the mean size of our bogs?
l_mean_size = bogs['area_km2'].mean()
l_mean_size
fig, ax = plt.subplots()
bogs['area_km2'].plot.hist(bins=10)
# Add title
plt.title("Bogs area_km2 histogram")
def binaryClassifier(row, source_col, output_col, threshold):
# If area of input geometry is lower that the threshold value
if row[source_col] < threshold:
# Update the output column with value 0
row[output_col] = 0
# If area of input geometry is higher than the threshold value update with value 1
else:
row[output_col] = 1
# Return the updated row
return row
bogs['small_big'] = None
bogs = bogs.apply(binaryClassifier, source_col='area_km2', output_col='small_big', threshold=l_mean_size, axis=1)
bogs.plot(column='small_big', linewidth=0.05, cmap="seismic")
```
## Classification based on common classification schemes
```
fp = "population_admin_units.shp"
acc = gpd.read_file(fp)
import pysal.viz.mapclassify as mc
# Define the number of classes
n_classes = 5
# Create a Natural Breaks classifier
classifier = mc.NaturalBreaks.make(k=n_classes)
acc.dtypes
import numpy as np
def change_type_defensively(row):
try:
return int(row['population'])
except Exception:
return np.nan
acc['population_int'] = acc.apply(change_type_defensively, axis=1)
acc.head(5)
# Classify the data
acc['population_classes'] = acc[['population_int']].apply(classifier)
# Let's see what we have
acc.head()
acc.plot(column="population_classes", linewidth=0, legend=True)
# Plot
fig, ax = plt.subplots()
acc["population_int"].plot.hist(bins=100);
# Add title
plt.title("Amount of inhabitants column histogram")
grouped = acc.groupby('population_classes')
# legend_dict = { 'class from to' : 'white'}
legend_dict = {}
for cl, valds in grouped:
minv = valds['population_int'].min()
maxv = valds['population_int'].max()
print("Class {}: {} - {}".format(cl, minv, maxv))
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
import collections
# legend_dict, a special ordered dictionary (which reliably remembers order of adding things) that holds our class description and gives it a colour on the legend (we leave it "background" white for now)
legend_dict = collections.OrderedDict([])
for cl, valds in grouped:
minv = valds['population_int'].min()
maxv = valds['population_int'].max()
legend_dict.update({"Class {}: {} - {}".format(cl, minv, maxv): "white"})
# Plot preps for several plot into one figure
fig, ax = plt.subplots()
# plot the dataframe, with the natural breaks colour scheme
acc.plot(ax=ax, column="population_classes", linewidth=0, legend=True)
# the custom "patches" per legend entry of our additional labels
patchList = []
for key in legend_dict:
data_key = mpatches.Patch(color=legend_dict[key], label=key)
patchList.append(data_key)
# plot the custom legend
plt.legend(handles=patchList, loc='lower center', bbox_to_anchor=(0.5, -0.5), ncol=1)
# Add title
plt.title("Amount of inhabitants natural breaks classifier")
plt.tight_layout()
```
| github_jupyter |
```
import dataset
import json
import csv
import tweepy
from sqlalchemy.exc import ProgrammingError
import pandas as pd
import dataset
db = dataset.connect("DB_URL")
#insert your credentials here
import os
from dotenv import load_dotenv
load_dotenv()
CONSUMER_KEY = os.getenv('CONSUMER_KEY')
CONSUMER_SECRET = os.getenv('CONSUMER_SECRET')
ACCESS_KEY = os.getenv('ACCESS_KEY')
ACCESS_SECRET = os.getenv('ACCESS_SECRET')
class StreamListener(tweepy.StreamListener):
def on_status(self, status):
filter_words = ["police", "officer", "cop"]
conditions = (not 'RT @' in status.text) and any(word in status.text for word in filter_words)
if conditions:
description = status.user.description
loc = status.user.location
text = status.text
coords = status.coordinates
geo = status.geo
name = status.user.screen_name
user_created = status.user.created_at
id_str = status.id_str
created = status.created_at
source = status.user.url
language = status.lang
if geo is not None:
geo = json.dumps(geo)
if coords is not None:
coords = json.dumps(coords)
table = db["tweets"]
try:
table.insert(dict(
user_description=description,
user_location=loc,
coordinates=coords,
text=text,
geo=geo,
user_name=name,
user_created=user_created,
id_str=id_str,
created=created,
source = source,
language = language,
))
except ProgrammingError as err:
print(err)
def on_error(self, status_code):
if status_code == 420:
#return False in on_data disconnects the stream
return False
auth = tweepy.OAuthHandler(CONSUMER_KEY, CONSUMER_SECRET)
auth.set_access_token(ACCESS_KEY, ACCESS_SECRET)
api = tweepy.API(auth)
stream_listener = StreamListener()
stream = tweepy.Stream(auth=api.auth, listener=stream_listener)
stream.filter(track=["police", "cop", "officer"])
df_raw = pd.DataFrame(db['tweets'])
df_raw = df_raw[df_raw['language'].isin(["en", "und"]) ]
df = df_raw[["id_str", "text"]]
df['reddit'] = 0
df.rename(columns={'id_str': 'ids'}, inplace=True)
df
df_reddit = pd.read_csv('reddit_tweets.csv')
df_combined = pd.concat([df_reddit, df])
df_combined
#created CSV
df_combined.to_csv("combined_tweets.csv", index=False)
```
| github_jupyter |
## All of the module notebooks combined into this single notebook.
```
storage_account = 'steduanalytics__update_this'
use_test_env = True
if use_test_env:
stage1 = 'abfss://test-env@' + storage_account + '.dfs.core.windows.net/stage1'
stage2 = 'abfss://test-env@' + storage_account + '.dfs.core.windows.net/stage2'
stage3 = 'abfss://test-env@' + storage_account + '.dfs.core.windows.net/stage3'
else:
stage1 = 'abfss://stage1@' + storage_account + '.dfs.core.windows.net'
stage2 = 'abfss://stage2@' + storage_account + '.dfs.core.windows.net'
stage3 = 'abfss://stage3@' + storage_account + '.dfs.core.windows.net'
# Extracted from Clever_setup_and_update
# Process resource usage
df = spark.read.csv(stage1 + '/clever', header='true', inferSchema='true')
df = df.withColumn('sis_id',df.sis_id.cast('string'))
df.write.format('parquet').mode('overwrite').save(stage2 + '/clever/resource_usage_students')
# Anonymize data and load into stage3
from pyspark.sql.functions import sha2, lit
df = spark.read.format('parquet').load(stage2 + '/clever/resource_usage_students')
df = df.withColumn('sis_id', sha2(df.sis_id, 256)).withColumn('clever_user_id',lit('*')).withColumn('clever_school_id',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/clever/resource_usage_students')
# Create sql on-demand db for Clever data
def create_spark_db(db_name, source_path):
spark.sql('CREATE DATABASE IF NOT EXISTS ' + db_name)
spark.sql("create table if not exists " + db_name + ".resource_usage_students using PARQUET location '" + source_path + "/resource_usage_students'")
db_prefix = 'test_' if use_test_env else ''
create_spark_db(db_prefix + 's2_clever', stage2 + '/clever')
create_spark_db(db_prefix + 's3_clever', stage3 + '/clever')
# Extracted from contoso_sis_setup_and_update
# Process studentsectionmark and studentattendance
df = spark.read.csv(stage1 + '/contoso_sis/studentsectionmark.csv', header='true', inferSchema='true')
df = df.withColumn('id',df.id.cast('string')).withColumn('student_id',df.student_id.cast('string'))
df.write.format('parquet').mode('overwrite').save(stage2 + '/contoso_sis/studentsectionmark')
df = spark.read.csv(stage1 + '/contoso_sis/studentattendance.csv', header='true', inferSchema='true')
df = df.withColumn('id',df.id.cast('string')).withColumn('student_id',df.student_id.cast('string'))
df.write.format('parquet').mode('overwrite').save(stage2 + '/contoso_sis/studentattendance')
# Anonymize data and load into stage3
df = spark.read.format('parquet').load(stage2 + '/contoso_sis/studentsectionmark')
df = df.withColumn('id', sha2(df.id, 256)).withColumn('student_id',sha2(df.student_id, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/contoso_sis/studentsectionmark')
df = spark.read.format('parquet').load(stage2 + '/contoso_sis/studentattendance')
df = df.withColumn('id', sha2(df.id, 256)).withColumn('student_id',sha2(df.student_id, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/contoso_sis/studentattendance')
# Create spark db to allow for access to the data in the delta-lake via SQL on-demand.
def create_spark_db(db_name, source_path):
spark.sql('CREATE DATABASE IF NOT EXISTS ' + db_name)
spark.sql("create table if not exists " + db_name + ".studentsectionmark using PARQUET location '" + source_path + "/studentsectionmark'")
spark.sql("create table if not exists " + db_name + ".studentattendance using PARQUET location '" + source_path + "/studentattendance'")
db_prefix = 'test_' if use_test_env else ''
create_spark_db(db_prefix + 's2_contoso_sis', stage2 + '/contoso_sis')
create_spark_db(db_prefix + 's3_contoso_sis', stage3 + '/contoso_sis')
# Extracted from iReady_setup_and_update
# Process personalized_instruction_by_lesson_math.csv
def remove_spaces(str): return str.replace(' ', '').replace('(','_').replace(')','_').replace('=', '__')
def process(filename):
df = spark.read.csv(stage1 + '/iready/' + filename + '.csv', header='true', inferSchema='true')
newColumns = map(remove_spaces, df.columns)
df = df.toDF(*newColumns)
df = df.withColumn('StudentID',df.StudentID.cast('string')) # StudentID needs to be a string to allow for hashing when moving into stage3
df.write.format('parquet').mode('overwrite').save(stage2 + '/iready/' + filename)
process('comprehensive_student_lesson_activity_with_standards_ela')
process('comprehensive_student_lesson_activity_with_standards_math')
process('diagnostic_and_instruction_ela_ytd_window')
process('diagnostic_and_instruction_math_ytd_window')
process('diagnostic_results_ela')
process('diagnostic_results_math')
process('personalized_instruction_by_lesson_ela')
process('personalized_instruction_by_lesson_math')
# Anonymize data and load into stage3
from pyspark.sql.functions import sha2, lit
df = spark.read.format('parquet').load(stage2 + '/iready/comprehensive_student_lesson_activity_with_standards_ela')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/comprehensive_student_lesson_activity_with_standards_ela')
df = spark.read.format('parquet').load(stage2 + '/iready/comprehensive_student_lesson_activity_with_standards_math')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/comprehensive_student_lesson_activity_with_standards_math')
df = spark.read.format('parquet').load(stage2 + '/iready/diagnostic_and_instruction_ela_ytd_window')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*')).withColumn('UserName', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/diagnostic_and_instruction_ela_ytd_window')
df = spark.read.format('parquet').load(stage2 + '/iready/diagnostic_and_instruction_math_ytd_window')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*')).withColumn('UserName', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/diagnostic_and_instruction_math_ytd_window')
df = spark.read.format('parquet').load(stage2 + '/iready/diagnostic_results_ela')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/diagnostic_results_ela')
df = spark.read.format('parquet').load(stage2 + '/iready/diagnostic_results_math')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/diagnostic_results_math')
df = spark.read.format('parquet').load(stage2 + '/iready/personalized_instruction_by_lesson_ela')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/personalized_instruction_by_lesson_ela')
df = spark.read.format('parquet').load(stage2 + '/iready/personalized_instruction_by_lesson_math')
df = df.withColumn('StudentID', sha2(df.StudentID, 256)).withColumn('LastName',lit('*')).withColumn('FirstName',lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/iready/personalized_instruction_by_lesson_math')
# Extracted from M365_setup_and_update
stage1_m365 = stage1 + '/m365/DIPData'
stage1_m365_activity = stage1 + '/m365/DIPData/Activity/ApplicationUsage'
# Process Roster data from stage 1 to stage 2
#
# Sets up the edu_dl (stage 2 data lake) with whatever data is found in the DIP inbound folder.
# This includes:
# - adding column names
# - casting values into a schema
# Calendar
df = spark.read.csv(stage1_m365 + '/Roster/Calendar.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Calendar')
df = spark.sql("select _c0 Id, _c1 Name, _c2 Description, cast(_c3 as int) SchoolYear, cast(_c4 as boolean) IsCurrent, _c5 ExternalId, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c7, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c8 as boolean) IsActive, _c9 OrgId from Calendar")
df.write.format("parquet").mode("overwrite").save(stage2 + '/m365/Calendar')
# Course
df = spark.read.csv(stage1_m365 + '/Roster/Course.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Course')
df = spark.sql("select _c0 Id, _c1 Name, _c2 Code, _c3 Description, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 CalendarId from Course")
df.write.format("parquet").mode("overwrite").save(stage2 + '/m365/Course')
# Org
df = spark.read.csv(stage1_m365 + '/Roster/Org.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Org')
df = spark.sql("select _c0 Id, _c1 Name, _c2 Identifier, _c3 ExternalId, to_timestamp(_c4, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c6 as boolean) IsActive, _c7 ParentOrgId, _c8 RefOrgTypeId, _c9 SourceSystemId from Org")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/Org')
# Person
df = spark.read.csv(stage1_m365 + '/Roster/Person.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Person')
df_Person = spark.sql("select _c0 Id, _c1 FirstName, _c2 MiddleName, _c3 LastName, _c4 GenerationCode, _c5 Prefix, _c6 EnabledUser, _c7 ExternalId, to_timestamp(_c8, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c9, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c10 as boolean) IsActive, _c11 SourceSystemId from Person")
df_Person.write.format('parquet').mode('overwrite').save(stage2 + '/m365/Person')
# PersonIdentifier
df = spark.read.csv(stage1_m365 + '/Roster/PersonIdentifier.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'PersonIdentifier')
df = spark.sql("select _c0 Id, _c1 Identifier, _c2 Description, _c3 RefIdentifierTypeId, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 PersonId, _c9 SourceSystemId from PersonIdentifier")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/PersonIdentifier')
# RefDefinition
df = spark.read.csv(stage1_m365 + '/Roster/RefDefinition.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'RefDefinition')
df = spark.sql("select _c0 Id, _c1 RefType, _c2 Namespace, _c3 Code, cast(_c4 as int) SortOrder, _c5 Description, cast(_c6 as boolean) IsActive from RefDefinition")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/RefDefinition')
# Section
df = spark.read.csv(stage1_m365 + '/Roster/Section.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Section')
df = spark.sql("select _c0 Id, _c1 Name, _c2 Code, _c3 Location, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 CourseId, _c9 RefSectionTypeId, _c10 SessionId, _c11 OrgId from Section")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/Section')
# Session
df = spark.read.csv(stage1_m365 + '/Roster/Session.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'Session')
df = spark.sql("select _c0 Id, _c1 Name, to_timestamp(_c2, 'MM/dd/yyyy hh:mm:ss a') BeginDate, to_timestamp(_c3, 'MM/dd/yyyy hh:mm:ss a') EndDate, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 CalendarId, _c9 ParentSessionId, _c10 RefSessionTypeId from Session")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/Session')
# StaffOrgAffiliation
df = spark.read.csv(stage1_m365 + '/Roster/StaffOrgAffiliation.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'StaffOrgAffiliation')
df = spark.sql("select _c0 Id, cast(_c1 as boolean) IsPrimary, to_timestamp(_c2, 'MM/dd/yyyy hh:mm:ss a') EntryDate, to_timestamp(_c3, 'MM/dd/yyyy hh:mm:ss a') ExitDate, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 OrgId, _c9 PersonId, _c10 RefStaffOrgRoleId from StaffOrgAffiliation")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/StaffOrgAffiliation')
# StaffSectionMembership
df = spark.read.csv(stage1_m365 + '/Roster/StaffSectionMembership.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'StaffSectionMembership')
df = spark.sql("select _c0 Id, cast(_c1 as boolean) IsPrimaryStaffForSection, to_timestamp(_c2, 'MM/dd/yyyy hh:mm:ss a') EntryDate, to_timestamp(_c3, 'MM/dd/yyyy hh:mm:ss a') ExitDate, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 PersonId, _c9 RefStaffSectionRoleId, _c10 SectionId from StaffSectionMembership")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/StaffSectionMembership')
# StudentOrgAffiliation
df = spark.read.csv(stage1_m365 + '/Roster/StudentOrgAffiliation.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'StudentOrgAffiliation')
df = spark.sql("select _c0 Id, cast(_c1 as boolean) IsPrimary, to_timestamp(_c2, 'MM/dd/yyyy hh:mm:ss a') EntryDate, to_timestamp(_c3, 'MM/dd/yyyy hh:mm:ss a') ExitDate, _c4 ExternalId, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c6, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c7 as boolean) IsActive, _c8 OrgId, _c9 PersonId, _c10 RefGradeLevelId, _c11 RefStudentOrgRoleId, _c12 RefEnrollmentStatusId from StudentOrgAffiliation")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/StudentOrgAffiliation')
# StudentSectionMembership
df = spark.read.csv(stage1_m365 + '/Roster/StudentSectionMembership.csv', header='false')
if (df.count() > 0):
sqlContext.registerDataFrameAsTable(df, 'StudentSectionMembership')
df = spark.sql("select _c0 Id, to_timestamp(_c1, 'MM/dd/yyyy hh:mm:ss a') EntryDate, to_timestamp(_c2, 'MM/dd/yyyy hh:mm:ss a') ExitDate, _c3 ExternalId, to_timestamp(_c4, 'MM/dd/yyyy hh:mm:ss a') CreateDate, to_timestamp(_c5, 'MM/dd/yyyy hh:mm:ss a') LastModifiedDate, cast(_c6 as boolean) IsActive, _c7 PersonId, _c8 RefGradeLevelWhenCourseTakenId, _c9 RefStudentSectionRoleId, _c10 SectionId from StudentSectionMembership")
df.write.format('parquet').mode('overwrite').save(stage2 + '/m365/StudentSectionMembership')
# Process Activity data from stage1 into stage2.
#
# If this is the first load, it loads all activity data.
# If this is a subsequent load, it determines the max date currently stored and only loads data from after that date.
def append_to_activity_table(max_date=False):
df = spark.read.csv(stage1_m365_activity, header='false')
sqlContext.registerDataFrameAsTable(df, 'Activity')
df_Activity = spark.sql("select _c0 SignalType, to_timestamp(_c1) StartTime, _c2 UserAgent, _c3 SignalId, _c4 SISClassId, _c5 OfficeClassId, _c6 ChannelId, _c7 AppName, _c8 ActorId, _c9 ActorRole, _c10 SchemaVersion, _c11 AssignmentId, _c12 SubmissionId, _c13 Action, _c14 AssginmentDueDate, _c15 ClassCreationDate, _c16 Grade, _c17 SourceFileExtension, _c18 MeetingDuration, '' PersonId from Activity")
if (max_date):
df_Activity = df_Activity.filter(df_Activity.StartTime > max_date)
if (df_Activity.count() == 0):
print('No new activity data to load')
else:
print('Adding activity data later than: ' + str(max_date))
# The assumption here is that there will always be data in these inbound files
sqlContext.registerDataFrameAsTable(df_Activity, 'Activity')
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/PersonIdentifier'), 'PersonIdentifier')
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/RefDefinition'), 'RefDefinition')
df1 = spark.sql( \
"select act.SignalType, act.StartTime, act.UserAgent, act.SignalId, act.SISClassId, act.OfficeClassId, act.ChannelId, \
act.AppName, act.ActorId, act.ActorRole, act.SchemaVersion, act.AssignmentId, act.SubmissionId, act.Action, act.AssginmentDueDate, \
act.ClassCreationDate, act.Grade, act.SourceFileExtension, act.MeetingDuration, pi.PersonId \
from PersonIdentifier pi, RefDefinition rd, Activity act \
where \
pi.RefIdentifierTypeId = rd.Id \
and rd.RefType = 'RefIdentifierType' \
and rd.Code = 'ActiveDirectoryId' \
and pi.Identifier = act.ActorId \
")
df1.write.format("parquet").mode("append").save(stage2 + '/m365/Activity0p2')
try:
df = spark.read.format('parquet').load(stage2 + '/m365/Activity0p2')
sqlContext.registerDataFrameAsTable(df, 'Activity')
# Bad data with a date in the future can prevent the uploading of new activity data,
# so we ensure that the watermark is calculated on good data by filtering with CURRENT_TIMESTAMP
df1 = spark.sql("select StartTime from Activity where StartTime < CURRENT_TIMESTAMP")
max_date = df1.agg({'StartTime': 'max'}).first()[0]
print(max_date)
append_to_activity_table(max_date)
except:
print("No Activity data has been loaded into stage2 data lake yet.")
append_to_activity_table()
# Anonymize the M365 data from stage2 and load into stage3
from pyspark.sql.functions import sha2, lit
# Activity
df = spark.read.format('parquet').load(stage2 + '/m365/Activity0p2')
df = df.withColumn('PersonId', sha2(df.PersonId, 256)).withColumn('ActorId', sha2(df.ActorId, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/Activity0p2')
# Calendar, Course, Org
spark.read.format('parquet').load(stage2 + '/m365/Calendar').write.format('parquet').mode('overwrite').save(stage3 + '/m365/Calendar')
spark.read.format('parquet').load(stage2 + '/m365/Course').write.format('parquet').mode('overwrite').save(stage3 + '/m365/Course')
spark.read.format('parquet').load(stage2 + '/m365/Org').write.format('parquet').mode('overwrite').save(stage3 + '/m365/Org')
# Person
df = spark.read.format('parquet').load(stage2 + '/m365/Person')
df = df.withColumn('Id', sha2(df.Id, 256)).withColumn('FirstName', lit('*')).withColumn("MiddleName", lit('*')).withColumn('LastName', lit('*')).withColumn('ExternalId', sha2(df.ExternalId, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/Person')
# PersonIdentifier
df = spark.read.format('parquet').load(stage2 + '/m365/PersonIdentifier')
df = df.withColumn('PersonId', sha2(df.Id, 256)).withColumn('Identifier', lit('*')).withColumn("ExternalId", lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/PersonIdentifier')
# RefDefinition, Section, Session
spark.read.format('parquet').load(stage2 + '/m365/RefDefinition').write.format('parquet').mode('overwrite').save(stage3 + '/m365/RefDefinition')
spark.read.format('parquet').load(stage2 + '/m365/Section').write.format('parquet').mode('overwrite').save(stage3 + '/m365/Section')
spark.read.format('parquet').load(stage2 + '/m365/Session').write.format('parquet').mode('overwrite').save(stage3 + '/m365/Session')
# StaffOrgAffiliation
df = spark.read.format('parquet').load(stage2 + '/m365/StaffOrgAffiliation')
df = df.withColumn('PersonId', sha2(df.PersonId, 256)).withColumn('ExternalId', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/StaffOrgAffiliation')
# StaffSectionMembership
df = spark.read.format('parquet').load(stage2 + '/m365/StaffSectionMembership')
df = df.withColumn('PersonId', sha2(df.PersonId, 256)).withColumn('ExternalId', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/StaffSectionMembership')
# StudentOrgAffiliation
df = spark.read.format('parquet').load(stage2 + '/m365/StudentOrgAffiliation')
df = df.withColumn('PersonId', sha2(df.PersonId, 256)).withColumn('ExternalId', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/StudentOrgAffiliation')
# StudentSectionMembership
df = spark.read.format('parquet').load(stage2 + '/m365/StudentSectionMembership')
df = df.withColumn('PersonId', sha2(df.PersonId, 256)).withColumn('ExternalId', lit('*'))
df.write.format('parquet').mode('overwrite').save(stage3 + '/m365/StudentSectionMembership')
# Create spark db to allow for access to the data in the delta-lake via SQL on-demand.
# This is only creating metadata for SQL on-demand, pointing to the data in the delta-lake.
# This also makes it possible to connect in Power BI via the azure sql data source connector.
def create_spark_db(db_name, source_path):
spark.sql('CREATE DATABASE IF NOT EXISTS ' + db_name)
spark.sql("create table if not exists " + db_name + ".Activity using PARQUET location '" + source_path + "/Activity0p2'")
spark.sql("create table if not exists " + db_name + ".Calendar using PARQUET location '" + source_path + "/Calendar'")
spark.sql("create table if not exists " + db_name + ".Course using PARQUET location '" + source_path + "/Course'")
spark.sql("create table if not exists " + db_name + ".Org using PARQUET location '" + source_path + "/Org'")
spark.sql("create table if not exists " + db_name + ".Person using PARQUET location '" + source_path + "/Person'")
spark.sql("create table if not exists " + db_name + ".PersonIdentifier using PARQUET location '" + source_path + "/PersonIdentifier'")
spark.sql("create table if not exists " + db_name + ".RefDefinition using PARQUET location '" + source_path + "/RefDefinition'")
spark.sql("create table if not exists " + db_name + ".Section using PARQUET location '" + source_path + "/Section'")
spark.sql("create table if not exists " + db_name + ".Session using PARQUET location '" + source_path + "/Session'")
spark.sql("create table if not exists " + db_name + ".StaffOrgAffiliation using PARQUET location '" + source_path + "/StaffOrgAffiliation'")
spark.sql("create table if not exists " + db_name + ".StaffSectionMembership using PARQUET location '" + source_path + "/StaffSectionMembership'")
spark.sql("create table if not exists " + db_name + ".StudentOrgAffiliation using PARQUET location '" + source_path + "/StudentOrgAffiliation'")
spark.sql("create table if not exists " + db_name + ".StudentSectionMembership using PARQUET location '" + source_path + "/StudentSectionMembership'")
db_prefix = 'test_' if use_test_env else ''
create_spark_db(db_prefix + 's2_m365', stage2 + '/m365')
create_spark_db(db_prefix + 's3_m365', stage3 + '/m365')
# Extracted from Contoso_ISD_setup
from pyspark.sql.functions import sha2, lit
# Process studentsectionmark and studentattendance
df = spark.read.csv(stage1 + '/contoso_sis/studentsectionmark.csv', header='true', inferSchema='true')
df = df.withColumn('id',df.id.cast('string')).withColumn('student_id',df.student_id.cast('string'))
df.write.format('parquet').mode('overwrite').save(stage2 + '/contoso_sis/studentsectionmark')
df = spark.read.csv(stage1 + '/contoso_sis/studentattendance.csv', header='true', inferSchema='true')
df = df.withColumn('id',df.id.cast('string')).withColumn('student_id',df.student_id.cast('string'))
df.write.format('parquet').mode('overwrite').save(stage2 + '/contoso_sis/studentattendance')
# Anonymize data and load into stage3
df = spark.read.format('parquet').load(stage2 + '/contoso_sis/studentsectionmark')
df = df.withColumn('id', sha2(df.id, 256)).withColumn('student_id',sha2(df.student_id, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/contoso_sis/studentsectionmark')
df = spark.read.format('parquet').load(stage2 + '/contoso_sis/studentattendance')
df = df.withColumn('id', sha2(df.id, 256)).withColumn('student_id',sha2(df.student_id, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/contoso_sis/studentattendance')
# Create spark db to allow for access to the data in the delta-lake via SQL on-demand.
def create_spark_db(db_name, source_path):
spark.sql('CREATE DATABASE IF NOT EXISTS ' + db_name)
spark.sql("create table if not exists " + db_name + ".studentsectionmark using PARQUET location '" + source_path + "/studentsectionmark'")
spark.sql("create table if not exists " + db_name + ".studentattendance using PARQUET location '" + source_path + "/studentattendance'")
db_prefix = 'test_' if use_test_env else ''
create_spark_db(db_prefix + 's2_contoso_sis', stage2 + '/contoso_sis')
create_spark_db(db_prefix + 's3_contoso_sis', stage3 + '/contoso_sis')
# Extracted from Contoso_ISD_setup_and_update
# Process sectionmark data
# Convert id values to use the Person.Id and Section.Id values set in the Education Data Platform.
from pyspark.sql.functions import sha2, lit
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/contoso_sis/studentsectionmark'), 'SectionMark')
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/Person'), 'Person')
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/Section'), 'Section')
df = spark.sql("select sm.id Id, p.Id PersonId, s.Id SectionId, cast(sm.numeric_grade_earned as int) NumericGrade, \
sm.alpha_grade_earned AlphaGrade, sm.is_final_grade IsFinalGrade, cast(sm.credits_attempted as int) CreditsAttempted, cast(sm.credits_earned as int) CreditsEarned, \
sm.grad_credit_type GraduationCreditType, sm.id ExternalId, CURRENT_TIMESTAMP CreateDate, CURRENT_TIMESTAMP LastModifiedDate, true IsActive \
from SectionMark sm, Person p, Section s \
where sm.student_id = p.ExternalId \
and sm.section_id = s.ExternalId")
df.write.format('parquet').mode('overwrite').save(stage2 + '/ContosoISD/SectionMark')
df.write.format('parquet').mode('overwrite').save(stage2 + '/ContosoISD/SectionMark2')
# Add SectionMark data to stage3 (anonymized parquet lake)
df = df.withColumn('PersonId', sha2(df.PersonId, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/ContosoISD/SectionMark')
df.write.format('parquet').mode('overwrite').save(stage3 + '/ContosoISD/SectionMark2')
# Repeat the above process, this time for student attendance
# Convert id values to use the Person.Id, Org.Id and Section.Id values
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/contoso_sis/studentattendance'), 'Attendance')
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/Org'), 'Org')
df = spark.sql("select att.id Id, p.Id PersonId, att.school_year SchoolYear, o.Id OrgId, to_date(att.attendance_date,'MM/dd/yyyy') AttendanceDate, \
att.all_day AllDay, att.Period Period, s.Id SectionId, att.AttendanceCode AttendanceCode, att.PresenceFlag PresenceFlag, \
att.attendance_status AttendanceStatus, att.attendance_type AttendanceType, att.attendance_sequence AttendanceSequence \
from Attendance att, Org o, Person p, Section s \
where att.student_id = p.ExternalId \
and att.school_id = o.ExternalId \
and att.section_id = s.ExternalId")
df.write.format('parquet').mode('overwrite').save(stage2 +'/ContosoISD/Attendance')
# Add Attendance data to stage3 (anonymized parquet lake)
df = df.withColumn('PersonId', sha2(df.PersonId, 256))
df.write.format('parquet').mode('overwrite').save(stage3 + '/ContosoISD/Attendance')
# Add 'Department' column to Course (hardcoded to "Math" for this Contoso example)
sqlContext.registerDataFrameAsTable(spark.read.format('parquet').load(stage2 + '/m365/Course'), 'Course')
df = spark.sql("select Id, Name, Code, Description, ExternalId, CreateDate, LastModifiedDate, IsActive, CalendarId, 'Math' Department from Course")
df.write.format('parquet').mode('overwrite').save(stage2 + '/ContosoISD/Course')
df.write.format('parquet').mode('overwrite').save(stage3 + '/ContosoISD/Course')
# Create spark db to allow for access to the data in the delta-lake via SQL on-demand.
# This is only creating metadata for SQL on-demand, pointing to the data in the delta-lake.
# This also makes it possible to connect in Power BI via the azure sql data source connector.
def create_spark_db(db_name, source_path):
spark.sql('CREATE DATABASE IF NOT EXISTS ' + db_name)
spark.sql(f"create table if not exists " + db_name + ".Activity using PARQUET location '" + source_path + "/m365/Activity0p2'")
spark.sql(f"create table if not exists " + db_name + ".Calendar using PARQUET location '" + source_path + "/m365/Calendar'")
spark.sql(f"create table if not exists " + db_name + ".Org using PARQUET location '" + source_path + "/m365/Org'")
spark.sql(f"create table if not exists " + db_name + ".Person using PARQUET location '" + source_path + "/m365/Person'")
spark.sql(f"create table if not exists " + db_name + ".PersonIdentifier using PARQUET location '" + source_path + "/m365/PersonIdentifier'")
spark.sql(f"create table if not exists " + db_name + ".RefDefinition using PARQUET location '" + source_path + "/m365/RefDefinition'")
spark.sql(f"create table if not exists " + db_name + ".Section using PARQUET location '" + source_path + "/m365/Section'")
spark.sql(f"create table if not exists " + db_name + ".Session using PARQUET location '" + source_path + "/m365/Session'")
spark.sql(f"create table if not exists " + db_name + ".StaffOrgAffiliation using PARQUET location '" + source_path + "/m365/StaffOrgAffiliation'")
spark.sql(f"create table if not exists " + db_name + ".StaffSectionMembership using PARQUET location '" + source_path + "/m365/StaffSectionMembership'")
spark.sql(f"create table if not exists " + db_name + ".StudentOrgAffiliation using PARQUET location '" + source_path + "/m365/StudentOrgAffiliation'")
spark.sql(f"create table if not exists " + db_name + ".StudentSectionMembership using PARQUET location '" + source_path + "/m365/StudentSectionMembership'")
spark.sql(f"create table if not exists " + db_name + ".Course using PARQUET location '" + source_path + "/ContosoISD/Course'")
spark.sql(f"create table if not exists " + db_name + ".Attendance using PARQUET location '" + source_path + "/ContosoISD/Attendance'")
spark.sql(f"create table if not exists " + db_name + ".SectionMark using PARQUET location '" + source_path + "/ContosoISD/SectionMark'")
spark.sql(f"create table if not exists " + db_name + ".SectionMark2 using PARQUET location '" + source_path + "/ContosoISD/SectionMark2'")
db_prefix = 'test_' if use_test_env else ''
create_spark_db(db_prefix + 's2_ContosoISD', stage2)
create_spark_db(db_prefix + 's3_ContosoISD', stage3)
```
| github_jupyter |
```
import os
import numpy as np
import pandas as pd
from numpy import linalg
from surprise import Reader, Dataset
from surprise import SVD
from surprise import SVDpp
os.chdir('/Recommendation system of films/data/')
df = pd.read_csv('ratings.csv', delimiter=',')
df = df.drop(['timestamp'], axis=1)
df.info()
df.describe()
import plotly.express as px
import seaborn as sns
import matplotlib.pyplot as plt
plt.figure(figsize=(15,7))
sns.set(style="darkgrid")
sns.countplot(df['rating'])
real=df.groupby(['rating'])['userId'].count()
real=pd.DataFrame(real)
fig = px.line(real)
fig.show()
real=df.groupby(['rating'])['movieId'].count()
real=pd.DataFrame(real)
fig = px.line(real)
fig.show()
real=df.groupby(['userId'])['rating'].count()
real=pd.DataFrame(real)
fig = px.line(real)
fig.show()
plt.figure(figsize=(15,10))
labels=['0.5','1','1.5','2','2.5','3','3.5','4','4.5','5']
colors = ["SkyBlue","PeachPuff",'lightcoral','gold','indigo','teal','magenta','deeppink','green','gray']
plt.pie(df['rating'].value_counts(),labels=labels,colors=colors,
autopct='%1.2f%%', shadow=True, startangle=140)
plt.show()
df['userId'] = df['userId'].astype(str)
df['movieId'] = df['movieId'].astype(str)
df['userId'] = 'person_'+df['userId'].astype(str)
df['movieId'] = 'movie_'+df['movieId'].astype(str)
df_new=df.copy()
df_new=df_new.rename(columns={"movieId": "userId/movieId"})
df_new = df_new.pivot_table(index=['userId'], columns='userId/movieId', values='rating', aggfunc=np.sum).reset_index()
df_new.index=df_new['userId'].values
df_new=df_new.drop(['userId'], axis=1)
df_new
minimum_rating = min(df['rating'].values)
maximum_rating = max(df['rating'].values)
print(minimum_rating,maximum_rating)
reader = Reader(rating_scale=(minimum_rating,maximum_rating))
data = Dataset.load_from_df(df[['userId', 'movieId', 'rating']], reader)
svdplpl = SVDpp(lr_all=0.005, reg_all=0.02)
svdplpl.fit(data.build_full_trainset())
df_svdplpl=df_new.copy()
for user in df_new.index:
for movie in df_new.columns:
if str(df_svdplpl.loc[user, movie])=='nan':
df_svdplpl.at[user, movie] = round(svdplpl.predict(user, movie).est,4)
df_svdplpl
```
manual svd++
```
i_bound=10
import random
from sklearn.metrics import mean_squared_error
from IPython.display import clear_output
import matplotlib.pyplot as plt
df_train=df_new.copy()
df_train=df_train.fillna(0)
df_train=df_train.loc[df_train.index[0:20],:].copy()
df_train_sum=pd.DataFrame(df_train.sum()).copy()
cols=df_train_sum[df_train_sum[0]!=0].index[0:20].tolist()
df_train=df_train.loc[:,cols].copy()
df_train_sum2=pd.DataFrame(df_train.T.sum()).copy()
inds=df_train_sum2[df_train_sum2[0]!=0].index.tolist()
df_train=df_train.loc[inds,:].copy()
df_train
df_bi = pd.DataFrame([random.random() for i in range(df_train.shape[0])])
df_bi['person_id']=df_train.index
df_bi.columns=['bi_value', 'person_id']
df_bi=df_bi.loc[:,['person_id','bi_value']]
for i in range(i_bound):
df_bi['pi_value_'+str(i)]=[random.random() for i in range(df_train.shape[0])]
df_bi
df_ba = pd.DataFrame([random.random() for i in range(df_train.shape[1])])
df_ba['movie_id']=df_train.columns
df_ba.columns=['ba_value', 'movie_id']
df_ba=df_ba.loc[:,['movie_id','ba_value']]
for i in range(i_bound):
df_ba['qa_value_'+str(i)]=[random.random() for i in range(df_train.shape[1])]
df_ba
df_train_fit=df_train.copy().reset_index()
df_train_fit=df_train_fit.rename(columns={"index": "person_id"})
lc = len(df_train_fit.columns)
df_train_fit = pd.melt(df_train_fit
, id_vars=['person_id'], value_vars = df_train_fit.columns[1:lc],
var_name='movie_id', value_name='rating')
df_train_fit = df_train_fit[df_train_fit['rating']!=0.0].copy()
df_train_fit.reset_index(drop=True, inplace=True)
df_train_fit
def analyse_errors(big_df, df_bi, df_ba, mu, i_bound):
big_df = big_df.merge(df_bi, how='left', on=['person_id'])
big_df = big_df.merge(df_ba, how='left', on=['movie_id'])
big_df['vect_dot']=0
for i in range(i_bound):
big_df['vect_dot'] = big_df['vect_dot'] + big_df['pi_value_'+str(i)]*big_df['qa_value_'+str(i)]
big_df['rating_pred']=mu + big_df['bi_value'] + big_df['ba_value'] + big_df['vect_dot']
return big_df, np.sqrt(mean_squared_error(big_df['rating'].values, big_df['rating_pred'].values))
def svd_plus_plus_fitting(df_train_fit, df_bi, df_ba, n_steps, gamma, lambdaa, i_bound):
first_rmse = 0
last_rmse = 0
steps=[]
rmses=[]
target_line=[]
mu = df_train_fit.mean().mean()
for step in range(n_steps):
steps.append(step)
part = random.sample(range(0, df_train_fit.shape[0]), 1)
item = df_train_fit.loc[part,:]
item.reset_index(drop=True, inplace=True)
item, _ = analyse_errors(big_df=item, df_bi=df_bi, df_ba=df_ba, mu=mu, i_bound=i_bound)
_, rmse = analyse_errors(big_df=df_train_fit, df_bi=df_bi, df_ba=df_ba, mu=mu, i_bound=i_bound)
if (step==0):
first_rmse=rmse
else:
last_rmse=rmse
rmses.append(rmse)
target_line.append(0)
item['rating_error']=item['rating'] - item['rating_pred']
movem = item['rating_error'].values.tolist()[0]
item=item.drop(['rating','vect_dot','rating_pred'], axis=1)
item['bi_value'] = item['bi_value'] + gamma*(item['rating_error'] - lambdaa*item['bi_value'])
item['ba_value'] = item['ba_value'] + gamma*(item['rating_error'] - lambdaa*item['ba_value'])
copy_pi_vector_pi = item[item.columns[3:(i_bound+3)]].copy()
copy_pi_vector_qa = item[item.columns[(i_bound+4):(len(item.columns)-1)]].copy()
copy_pi_vector_qa.columns = copy_pi_vector_pi.columns
copy_qa_vector_pi = item[item.columns[3:(i_bound+3)]].copy()
copy_qa_vector_qa = item[item.columns[(i_bound+4):(len(item.columns)-1)]].copy()
copy_qa_vector_pi.columns = copy_qa_vector_qa.columns
item[item.columns[3:(i_bound+3)]] = copy_pi_vector_pi + gamma*(copy_pi_vector_qa*movem -
lambdaa*copy_pi_vector_pi)
item[item.columns[(i_bound+4):(len(item.columns)-1)]] = copy_qa_vector_qa + gamma*(copy_qa_vector_pi*movem -
lambdaa*copy_qa_vector_qa)
item = item.drop(['rating_error'], axis=1)
item_bi = item[item.columns[0:1].tolist() + item.columns[2:(i_bound+3)].tolist()].copy()
item_ba = item[item.columns[1:2].tolist() + item.columns[(i_bound+3):].tolist()].copy()
pids = item_bi['person_id'].values.tolist()
mids = item_ba['movie_id'].values.tolist()
item_bi.index = df_bi[df_bi['person_id'].isin(pids)].index
df_bi[df_bi['person_id'].isin(pids)]=item_bi
item_ba.index=df_ba[df_ba['movie_id'].isin(mids)].index
df_ba[df_ba['movie_id'].isin(mids)]=item_ba
print('ITERATION NUMBER:',(step+1),' of ',n_steps,'RMSE=',rmse)
return df_bi, df_ba
n_steps=5000
gamma = 0.00095
lambdaa = 0.00005
df_bi_new, df_ba_new = svd_plus_plus_fitting(df_train_fit=df_train_fit, df_bi=df_bi, df_ba=df_ba,
n_steps=n_steps, gamma=gamma, lambdaa=lambdaa, i_bound=i_bound)
mu = df_train_fit.mean().mean()
l_bound=0.5
u_bound=5
df_svd_pp=df_train.copy()
for usr in df_train.index:
for flm in df_train.columns:
if df_svd_pp.loc[usr, flm]==0.0:
v1=df_ba_new[df_ba_new['movie_id']==flm][df_ba_new.columns[2:]].values[0]
v2=df_bi_new[df_bi_new['person_id']==usr][df_bi_new.columns[2:]].values[0]
ba_value = df_ba_new[df_ba_new['movie_id']==flm]['ba_value'].values.tolist()[0]
bi_value = df_bi_new[df_bi_new['person_id']==usr]['bi_value'].values.tolist()[0]
rate = mu + bi_value + ba_value + round(np.dot(v1,v2),4)
if rate<l_bound:
rate=l_bound
elif rate>u_bound:
rate=u_bound
df_svd_pp.at[usr, flm] = rate
df_svd_pp
df_svd_pp.max().max()
df_svd_pp.min().min()
df_svd_pp.apply(lambda x: sum(x.isnull()), axis=0).unique()
```
| github_jupyter |
# MNIST Dataset
Here we try our first neural network using Keras, which is a high level library which wraps most of the functionality in tensorflow.
Pay most attention to the section named `Keras Model`. The numpy, plotting functions should only be secondary considerations.
```
from keras.models import Sequential
from keras.layers import Dense
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets('./', one_hot=True)
x, y = mnist.train.next_batch(20)
x.shape
```
The above extracted 20 images from disk. The 784 comes from the fact that its a flattened 28x28 image.
```
28*28
```
`y` is the one-hot encoded labels. Meaning instead of simply stating that label is 0-9, it puts a 1 where the label is supposed to be and 0 else where. See below to have a better understanding:
```
y.shape
y
```
Visualise the first image (x[0]). Note that we need to reshape the flattened image before visualising:
```
plt.imshow(x[0].reshape(28,28))
plt.show()
```
## Keras Model
```
model = Sequential()
model.add(Dense(100, activation='relu', input_dim=784))
model.add(Dense(10, activation='softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
model.summary()
def get_batch(dataset, batch_size = 256):
while (1):
yield dataset.next_batch(batch_size)
batch_size = 256
test_gen = get_batch(mnist.test, batch_size)
steps_per_epoch = mnist.test.num_examples// batch_size
model.evaluate_generator(test_gen, steps_per_epoch)
batch_size = 256
data_gen = get_batch(mnist.train, batch_size)
steps_per_epoch = mnist.train.num_examples//batch_size
model.fit_generator(data_gen, steps_per_epoch, epochs=1)
model.evaluate_generator(test_gen, steps_per_epoch)
```
It is really important to be able to reload the model after you've been training it for hours on end (usually). So save the model.
```
from keras.models import load_model
model.save('my_model.h5')
model2 = load_model('my_model.h5')
model2.evaluate_generator(test_gen, steps_per_epoch)
x, y = next(test_gen)
x.shape
model.predict(x[:2])
plt.imshow(x[0].reshape(28,28))
plt.imshow(x[1].reshape(28,28))
model.predict_classes(x[:2])
```
## Conclusion
There are only 3 things you need to remember from this lesson.
1. Model Architecture.
2. model.fit(), don't worry too much about the generator part just yet. When you do lesson 5 we will ignore the generator functions. This is only useful if the data is too big to fit in memory. Essentially `fit()` is used to train your model.
3. model.predict() and model.predict_classes()
| github_jupyter |
### Please help by upvoting this kernel if you feel useful.
This note book will guide you quickly for the neccesary funtions to use in data science.
# Contents
1. [Numpy(Numerical Python)](#Numpy)
1. [Basic operations](#BasicOperation)
2. [Array creation](#creation)
3. [Indexing & Slicing](#slicing)
2. [Pandas](#Pandas)
1. [Data Structures](#DataStructures)
1. [Series](#Series)
2. [Data Frame](#DataFrames)
2. [Handle Missing Data](#MissingDataHandle)
1. [Drop missing data](#dropna)
2. [Fill missing data](#fillna)
3. [Data Operations](#DataOperations)
1. [Custom functions(applymap)](#CustomFunction)
2. [Statistical functions(max,mean,std)](#StatFunction)
3. [Grouping(groupby)](#groupby)
4. [Sorting(sort_values)](#Sorting)
3. [matplotlib](#matplotlib)
1. [Basic Plot](#basicplot)
2. [Multiple plots](#MultiplePlots)
3. [Subplots](#Subplots)
4. [Properties](#Properties)
1. [Alpha](#Alpha)
2. [Annotation](#Annotation)
5. [Types of plots](#Types)
1. [histogram](#histogram)
2. [scatter](#scatter)
3. [heatmap](#heatmap)
3. [pie](#pie)
4. [errorbar](#errorbar)
## 1) Numpy(Numerical Python)[^](#Numpy)<a id="Numpy" ></a><br>
is a library consisting of multidimensional array objects and a collection of routines for processing those arrays. Using NumPy, mathematical and logical operations on arrays can be performed.
```
#First we need to import the numpy library
import numpy as np # linear algebra
```
### 1) Basic operations[^](#BasicOperation)<a id="BasicOperation" ></a><br>
```
#1) Create arrays
data = np.array([[1, 2], [3, 4]]) # create 2D array
print(data)
data_complex = np.array([1, 2, 3], dtype = complex)
print(data_complex)
#2) see datatype
print(np.dtype(np.int64))
print(np.dtype('i1')) #int8=i1, int16=i2, int64=i4
#3) shape
print(data.shape)
#4) arange
data_arange = np.arange(15) #one dimentional array
print(data_arange)
#5) reshape, change shape
reshaped = data_arange.reshape((3,5))
print(reshaped.shape)
#6) itemsize : length of each element of array in bytes
print(reshaped.itemsize)
```
### 2) Array creations[^](#creation)<a id="creation" ></a><br>
```
#1) empty : array of random values (not initialized)
empty = np.empty([2,3]) #Default dtype is float
print(empty)
#2) zeros :
zeros = np.zeros([2,3], dtype = int)
print(zeros)
#3) ones :
ones = np.ones([2,3], dtype = int)
print(ones)
#4) create based on exising list
list_data = [1,2,3]
np_data = np.asarray(list_data)
print(np_data)
#5) create based on exising tuple
tuple_data = (1,2,3,5)
np_data_tuple = np.asarray(tuple_data)
print(np_data_tuple)
#6) from buffer
str_data = 'String date'.encode()
np_str = np.frombuffer(str_data, dtype = 'S1')
print(np_str)
#7) range func
range_data = np.asarray(range(5))
print(range_data)
#8) linspace : eg -> linspace(start, stop, num, endpoint, retstep, dtype)
linspace_data = np.linspace(10,20,5)
print(linspace_data)
linspace_data_1 = np.linspace(1,2,5, retstep = True)
print(linspace_data_1)
#9) logspace : numbers that are evenly spaced on a log scale (numpy.logspace(start, stop, num, endpoint, base, dtype))
log_data = np.logspace(1.0, 2.0, num = 10)
print(log_data)
```
### 3) Indexing & Slicing[^](#slicing)<a id="slicing" ></a><br>
```
#1) slice : slice(start:stop:step)
data = np.arange(10)
sliced = slice(2,7,2)
print(data)
print(data[sliced])
#2) same above with array with colon
data = np.arange(10)
sliced_index = data[2:7:2]
print(sliced_index)
#3) few indexed operations
data = np.array([[1,2,3],[3,4,5],[4,5,6]])
print('Original array is:')
print(data)
# this returns array of items in the second column
print('The items in the second column are:')
print(data[...,1])
# Now we will slice all items from the second row
print('The items in the second row are:')
print(data[1,...])
# Now we will slice all items from column 1 onwards
print('The items column 1 onwards are:')
print(data[...,1:])
```
## 2) Pandas[^](#Pandas)<a id="Pandas" ></a><br>
### 1) Data Structures[^](#DataStructures)<a id="DataStructures" ></a><br>
1. Series - labled 1D array
2. Data Frames -
3. Panel - 3 dimensional
4. Panel 4D - 4 dimensional
```
#First we need to import the pandas library
import pandas as pd
import numpy as np
```
#### 1) Series(One dimentinal array)[^](#Series)<a id="Series" ></a><br>
```
#### 1) Series(One dimentinal array)[^](#Series)<a id="Series" ></a><br>###Create Series
#1) list
list_series = pd.Series(list('abcdef'))
print(list_series)
#2) ndarray
arr_series = pd.Series(np.array(["one","two"]))
print(arr_series)
#3) dict
dict_series = pd.Series([120,230],index=["one","two"])
print(dict_series)
#4) scalar
scalar_series = pd.Series(3.,index=["a","b","c"])
print(scalar_series)
#### Access data of a series
print(dict_series[1]) # index
print(scalar_series[0:1]) # index range
print(dict_series.loc['one']) # index name
print(list_series.iloc[2]) # index position
```
#### 2) Data Frame(Two dimentinal array)[^](#DataFrame)<a id="DataFrame" ></a><br>
like a spread sheet
```
###Create Data Frames
#1) list
data_list = {'city':["London","Sydney"],'year':[2001,2005]}
list_df = pd.DataFrame(data_list)
print(list_df)
#2) dict
dict_data = {'London':{2001:100},'Sydney':{2005:200}}
dict_df = pd.DataFrame(dict_data)
print(dict_df)
#3) Series
series_data = pd.Series([120,230],index=["one","two"])
series_df = pd.DataFrame({'value':series_data})
print(series_df)
#4) narray
array_data = np.array([2001,2005,2006])
arr_df = pd.DataFrame({'year':array_data})
print(arr_df)
#4) dataframe
df_data = pd.DataFrame({'year':array_data})
df_df = pd.DataFrame(array_data)
print(df_df)
#Using above data frames
#View Data
print(list_df.city) # specific column
list_df.describe # whole dataset
print(arr_df.head(1)) #top records
print(arr_df.index) #list indexs
print(dict_df.columns) #list columns
print(list_df['year']) #specific column by name give column
print(dict_df.loc[2001]) #view by key gives row
print(dict_df.iloc[0:1]) #view by index gives rows
print(dict_df.iat[1,1]) #view by index gives value
print(list_df[list_df['year']>2003]) #view by condition, column greater than a value
```
* ### 2) Handle Missing Data[^](#MissingDataHandle)<a id="MissingDataHandle" ></a><br>
#### 1) Drop missing data[^](#dropna)<a id="dropna" ></a><br>
```
import pandas as pd
df = pd.DataFrame({'col1':{2001:100,2002:300},'col2':{2002:200}})
print("df : \n",df)
df_droped = df.dropna()
print("droped df : \n",df_droped)
```
#### 2) Fill missing data[^](#fillna)<a id="fillna" ></a><br>
```
import pandas as pd
df = pd.DataFrame({'col1':{2001:100,2002:300},'col2':{2002:200}})
print("df : \n",df)
df_filled = df.fillna(0)
print("filled df : \n",df_filled)
```
### 2) Data Operations[^](#DataOperations)<a id="DataOperations" ></a><br>
#### 1) Custom Function(applymap)[^](#CustomFunction)<a id="CustomFunction" ></a><br>
```
import pandas as pd
df_movie_rating = pd.DataFrame({'movie 1':[5,4,3,3,2,1],'movie 2':[4,2,1,2,3,5]},
index=['Tom','Jeff','Pterm','Ann','Ted','Paul'])
df_movie_rating
def movie_grade(rating):
if rating==5:
return 'A'
if rating==4:
return 'B'
if rating==3:
return 'C'
else:
return 'F'
print(movie_grade(4))
df_movie_rating.applymap(movie_grade)
```
#### 2) Statistical functions(max,mean,std)[^](#StatFunction)<a id="StatFunction" ></a><br>
```
import pandas as pd
df_test_scores = pd.DataFrame({'test 1':[98,89,34,23,45],'test 2':[23,34,50,76,80]}
,index=['Sam','Ann','Tom','Fed','Jef'])
df_test_scores
print("max : ",df_test_scores.max())
print("min : ",df_test_scores.min())
print("mean : ",df_test_scores.mean())
print("std : ",df_test_scores.std())
```
#### 3) Grouping (groupby)[^](#groupby)<a id="groupby" ></a><br>
```
df_names = pd.DataFrame({'first':['George','Bill','Ronald','Jimmy','George'],
'last':['Bush','Clienton','Regon','Carter','Washington']})
df_names
df_names_grouped = df_names.groupby('first')
df_names_grouped.get_group('George')
```
#### 4) Sorting (sort_values)[^](#Sorting)<a id="Sorting" ></a><br>
```
df_names.sort_values('first') # indexes will remain same unless you are re indexing
```
## 3) matplotlib[^](#matplotlib)<a id="matplotlib" ></a><br>
### 1) Basic plot[^](#basicplot)<a id="basicplot" ></a><br>
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
randomNumbers = np.random.rand(10)
print(randomNumbers)
style.use('ggplot')
plt.plot(randomNumbers,'g',label='line one',linewidth=2)
plt.xlabel('Range')
plt.ylabel('Numbers')
plt.title('Random number plot')
plt.legend()
plt.show()
```
### 2) Multiple plots[^](#MultiplePlots)<a id="MultiplePlots" ></a><br>
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
web_customers_monday = [12,34,5,232,232,232,53,5,64,34]
web_customers_tuesday = [3,23,12,21,500,54,34,65,87,92]
web_customers_wednesday = [32,82,23,22,332,242,153,73,12,23]
time_hrs = [2,4,6,7,8,10,12,15,18,20]
style.use('ggplot')
plt.plot(time_hrs,web_customers_monday,'r',label='monday',linewidth=1)
plt.plot(time_hrs,web_customers_tuesday,'g',label='tuesday',linewidth=1.2)
plt.plot(time_hrs,web_customers_wednesday,'b',label='wednesday',linewidth=1.5)
plt.title('Web site traffic')
plt.xlabel('Hrs')
plt.ylabel('Number of users')
plt.legend()
plt.show()
```
### 3) Sub plots[^](#Subplots)<a id="Subplots" ></a><br>
```
#subplot(row,cloum,position)
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
web_customers_monday = [12,34,5,232,232,232,53,5,64,34]
web_customers_tuesday = [3,23,12,21,500,54,34,65,87,92]
web_customers_wednesday = [32,82,23,22,332,242,153,73,12,23]
time_hrs = [2,4,6,7,8,10,12,15,18,20]
style.use('ggplot')
plt.figure(figsize=(8,4))
plt.subplots_adjust(hspace=1,wspace=1)
plt.subplot(2,2,1)
plt.title('Monday')
plt.plot(time_hrs,web_customers_monday,'r',label='monday',linewidth=1,linestyle='-')
plt.subplot(2,2,2)
plt.title('Tuesday')
plt.plot(time_hrs,web_customers_tuesday,'g',label='tuesday',linewidth=1.2)
plt.subplot(2,2,3)
plt.title('Wednesday')
plt.plot(time_hrs,web_customers_wednesday,'b',label='wednesday',linewidth=1.5)
plt.xlabel('Hrs')
plt.ylabel('Number of users')
plt.show()
```
### 4) Properties [^](#Properties)<a id="Properties" ></a><br>
Line properties
1. alpha
2. animated
Plot graphics
1. line style
2. line width
3. marker style
#### 1) Alpha [^](#Alpha)<a id="Alpha" ></a><br>
```
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
web_customers = [12,34,5,232,232,232,53,5,64,34]
time_hrs = [2,4,6,7,8,10,12,15,18,20]
style.use('ggplot')
plt.plot(time_hrs,web_customers,alpha=0.4)
plt.title('Web site traffic')
plt.xlabel('Hrs')
plt.ylabel('Number of users')
plt.show()
```
#### 2) Annotation [^](#Annotation)<a id="Annotation" ></a><br>
```
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
#Alpha for line transparency
web_customers = [12,34,10,232,200,180,53,5,64,34]
time_hrs = [2,4,6,7,8,10,12,15,18,20]
style.use('ggplot')
plt.plot(time_hrs,web_customers,alpha=0.7)
plt.title('Web site traffic')
plt.xlabel('Hrs')
plt.ylabel('Number of users')
#plt.annotate('annotation text','ha=horizontal align',va='vertical align',xytext=text position,
#xy=arrow position,arrowprops=properties of arrow)
plt.annotate('Max',ha='center',va='bottom',xytext=(5,232),xy=(7,232),arrowprops={'facecolor':'green'})
plt.annotate('Min',ha='center',va='bottom',xytext=(13,5),xy=(15,5),arrowprops={'facecolor':'green'})
plt.show()
```
### 1) Types of plots (types)[^](#Types)<a id="Types" ></a><br>
1. Histogram
2. Heat Map
3. Scatter Plot
4. Pie Chart
5. Error Bar
#### 1) Histogram (histogram)[^](#histogram)<a id="histogram" ></a><br>
```
#Histogram
from sklearn.datasets import load_boston
import matplotlib.pyplot as plt
from matplotlib import style
%matplotlib inline
bostan_real_state_data = load_boston()
#print(bostan_real_state_data.DESCR)
x_axis = bostan_real_state_data.data
y_axis = bostan_real_state_data.target
style.use('ggplot')
plt.figure(figsize=(8,8))
plt.hist(y_axis,bins=50)
plt.xlabel('price')
plt.ylabel('number of houses')
plt.show()
```
#### 2) Scatter (scatter)[^](#scatter)<a id="scatter" ></a><br>
```
#Scatter plot
style.use('ggplot')
plt.figure(figsize=(6,6))
plt.scatter(x_axis[:,5],y_axis)
plt.show()
```
#### 3) Heat Map (heatmap)[^](#heatmap)<a id="heatmap" ></a><br>
```
#Heat Map
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
flight_data = sns.load_dataset('flights')
#flight_data.head()
flight_data = flight_data.pivot('month','year','passengers')
sns.heatmap(flight_data)
```
#### 4) Pie chart (pie)[^](#pie)<a id="pie" ></a><br>
```
#Pie Charts
import matplotlib.pyplot as plt
%matplotlib inline
job_data = ['40','20','12','23','15']
labels = ['IT','Finace','marketing','Admin','HR']
explode = (0.05,0.04,0,0,0) #spilit the chart
#autopct= percent value embedded
plt.pie(job_data,labels=labels,explode=explode,autopct='%1.1f%%',startangle=70)
plt.axis('equal') # equal size chart
plt.show()
```
#### 5) Error bar (errorbar)[^](#errorbar)<a id="errorbar" ></a><br>
Error bars use mainly to identify errors
```
import numpy as np
import matplotlib.pyplot as plt
# example data
x = np.arange(0.1, 4, 0.5)
y = np.exp(-x)
# example variable error bar values
yerr = 0.1 + 0.2*np.sqrt(x)
xerr = 0.1 + yerr
# First illustrate basic pyplot interface, using defaults where possible.
plt.figure()
plt.errorbar(x, y, xerr=0.2, yerr=0.4)
plt.title("Simplest errorbars, 0.2 in x, 0.4 in y")
plt.show()
```
| github_jupyter |
```
"""
data preparation for model-based task:
1. extract the data with selected features;
2. set the rare categorical values to 'other';
3. fit a label encoder and a one-hot encoder for new data set
"""
##==================== Package ====================##
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import LabelEncoder
from dummyPy import OneHotEncoder
import random
import pickle # to store temporary variable
##==================== File-Path (fp) ====================##
## raw data (for read)
fp_train = "../../Datasets/ctr/train.csv"
fp_test = "../../Datasets/ctr/test.csv"
## subsample training set
fp_sub_train_f = "../../Datasets/ctr/sub_train_f.csv"
fp_col_counts = "../../Datasets/ctr/col_counts"
## data after selecting features (LR_fun needed)
## and setting rare categories' value to 'other' (feature filtering)
fp_train_f = "../../Datasets/ctr/train_f.csv"
fp_test_f = "../../Datasets/ctr/test_f.csv"
## storing encoder for labeling / one-hot encoding task
fp_lb_enc = "../../Datasets/ctr/lb_enc"
fp_oh_enc = "../../Datasets/ctr/oh_enc"
##==================== pre-Processing ====================##
## some simple original features is selected for dataset
'''features are used
C1: int, 1001, 1002, ...
banner_pos: int, 0,1,2,3,...
site_domain: object, large set of object variables
site_id: object, large set of object variables
site_category:object, large set of object variables
app_id: object, large set of object variables
app_category: object, small set of object variables
device_type: int, 0,1,2,3,4
device_conn_type:int, 0,1,2,3
C14: int, small set of int variables
C15: int, ...
C16: int, ...
'''
## feature names
cols = ['C1',
'banner_pos',
'site_domain',
'site_id',
'site_category',
'app_id',
'app_category',
'device_type',
'device_conn_type',
'C14',
'C15',
'C16']
cols_train = ['id', 'click']
cols_test = ['id']
cols_train.extend(cols)
cols_test.extend(cols)
## data reading
df_train_ini = pd.read_csv(fp_train, nrows = 10)
df_train_org = pd.read_csv(fp_train, chunksize = 1000000, iterator = True)
df_test_org = pd.read_csv(fp_test, chunksize = 1000000, iterator = True)
#----- counting features' categories numbers -----#
## 1.init_dict
cols_counts = {} # the categories count for each feature
for col in cols:
cols_counts[col] = df_train_ini[col].value_counts()
cols_counts
## 2.counting through train-set
for chunk in df_train_org:
for col in cols:
cols_counts[col] = cols_counts[col].append(chunk[col].value_counts())
## 3.counting through test-set
for chunk in df_test_org:
for col in cols:
cols_counts[col] = cols_counts[col].append(chunk[col].value_counts())
## 4.merge the deduplicates index in counting vectors
for col in cols:
cols_counts[col] = cols_counts[col].groupby(cols_counts[col].index).sum()
# sort the counts
cols_counts[col] = cols_counts[col].sort_values(ascending=False)
## 5.store the value_counting
pickle.dump(cols_counts, open(fp_col_counts, 'wb'))
## 6.show the distribution of value_counts
fig = plt.figure(1)
for i, col in enumerate(cols):
ax = fig.add_subplot(4, 3, i+1)
ax.fill_between(np.arange(len(cols_counts[col])), cols_counts[col].get_values())
# ax.set_title(col)
plt.show()
#----- set rare to 'other' -----#
# cols_counts = pickle.load(open(fp_col_counts, 'rb'))
## save at most k indices of the categorical variables
## and set the rest to 'other'
k = 99
col_index = {}
for col in cols:
col_index[col] = cols_counts[col][0: k].index
df_train_org = pd.read_csv(fp_train, dtype = {'id': str}, chunksize = 1000000, iterator = True)
df_test_org = pd.read_csv(fp_test, dtype = {'id': str}, chunksize = 1000000, iterator = True)
## train set
hd_flag = True # add column names at 1-st row
for chunk in df_train_org:
df = chunk.copy()
for col in cols:
df[col] = df[col].astype('object')
# assign all the rare variables as 'other'
df.loc[~df[col].isin(col_index[col]), col] = 'other'
with open(fp_train_f, 'a') as f:
df.to_csv(f, columns = cols_train, header = hd_flag, index = False)
hd_flag = False
## test set
hd_flag = True # add column names at 1-st row
for chunk in df_test_org:
df = chunk.copy()
for col in cols:
df[col] = df[col].astype('object')
# assign all the rare variables as 'other'
df.loc[~df[col].isin(col_index[col]), col] = 'other'
with open(fp_test_f, 'a') as f:
df.to_csv(f, columns = cols_test, header = hd_flag, index = False)
hd_flag = False
#----- generate encoder for label encoding -----#
#----- generate encoder for one-hot encoding -----#
'''
notes: here we do not apply label/one-hot transform
as we do it later in the iteration of model training on chunks
'''
## 1.label encoding
lb_enc = {}
for col in cols:
col_index[col] = np.append(col_index[col], 'other')
for col in cols:
lb_enc[col] = LabelEncoder()
lb_enc[col].fit(col_index[col])
## store the label encoder
pickle.dump(lb_enc, open(fp_lb_enc, 'wb'))
## 2.one-hot encoding
oh_enc = OneHotEncoder(cols)
df_train_f = pd.read_csv(fp_train_f, index_col=None, chunksize=500000, iterator=True)
df_test_f = pd.read_csv(fp_test_f, index_col=None, chunksize=500000, iterator=True)
for chunk in df_train_f:
oh_enc.fit(chunk)
for chunk in df_test_f:
oh_enc.fit(chunk)
## store the one-hot encoder
pickle.dump(oh_enc, open(fp_oh_enc, 'wb'))
#----- construct of original train set (sub-sampling randomly) -----#
n = sum(1 for line in open(fp_train_f)) - 1 # total size of train data (about 46M)
s = 2000000 # desired train set size (2M)
## the 0-indexed header will not be included in the skip list
skip = sorted(random.sample(range(1, n+1), n-s))
df_train = pd.read_csv(fp_train_f, skiprows = skip)
df_train.columns = cols_train
## store the sub-sampling train set as .csv
df_train.to_csv(fp_sub_train_f, index=False)
print(' - end - ')
for col in cols:
print col,lb_enc[col].classes_
```
| github_jupyter |
```
%matplotlib inline
import numpy as np
from numpy.random import rand
import matplotlib.pyplot as plt
#----------------------------------------------------------------------
## BLOCK OF FUNCTIONS USED IN THE MAIN CODE
#----------------------------------------------------------------------
def initialstate(N):
''' generates a random spin configuration for initial condition'''
# NxN matrix with random inters -1,1
state = 2*np.random.randint(2, size=(1,N))-1
return state
def mcmove(config, beta):
'''Monte Carlo move using Metropolis algorithm '''
#Config is the NxN spin matrix configuration
#In each iteration NxN updates
for i in range(N):
b = np.random.randint(0, N)
s = config[0,b]
nb = config[0,(b+1)%N] + config[0,(b-1)%N]
#print nb
cost = 2*s*nb
if cost < 0:
s *= -1
elif rand() < np.exp(-cost*beta):
s *= -1
config[0,b] = s
#print config
return config
def calcEnergy(config):
'''Energy of a given configuration'''
energy = 0
for i in range(N):
S = config[0,i]
nb = config[0,(i+1)%N] + config[0,(i-1)%N]
#print nb
energy += -nb*S
return energy/2. #Divide by 4 ensures , avoid multiple counting
def calcMag(config):
'''Magnetization of a given configuration'''
#magnetiation is just sum of all spins \sum_{i} S_i
mag = np.sum(np.sum(config))
return mag
## change the parameter below if you want to simulate a smaller system
nt = 2**4 # number of temperature points
N = 2**4 # size of the lattice, N
eqSteps = 2**8 # number of MC sweeps for equilibration
mcSteps = 2**8 # number of MC sweeps for calculation
n1, n2 = 1.0/(mcSteps*N*1), 1.0/(mcSteps*mcSteps*N*1)
#tm is the transition temperature
T=np.linspace(.01, 4, nt)
Energy = np.zeros(nt); Magnetization = np.zeros(nt)
SpecificHeat = np.zeros(nt); Susceptibility = np.zeros(nt)
#----------------------------------------------------------------------
# MAIN PART OF THE CODE
#----------------------------------------------------------------------
#m is the temperature index
for m in range(len(T)):
config = initialstate(N)
iT=1.0/T[m]; iT2=iT*iT;
for l in range(eqSteps): # equilibrate
mcmove(config, iT) # Monte Carlo moves
#Calculate quantities post equilibriation
E1 = M1 = E2 = M2 = 0
for i in range(mcSteps):
mcmove(config, iT)
Ene = calcEnergy(config) # calculate the energy
Mag = calcMag(config) # calculate the magnetisation
E1 = E1 + Ene #Cumulative energy
M1 = M1 + Mag
M2 = M2 + Mag*Mag
E2 = E2 + Ene*Ene
Energy[m] = n1*E1 #Weighing for MC averages.
Magnetization[m] = n1*M1
SpecificHeat[m] = (n1*E2 - n2*E1*E1)*iT2
Susceptibility[m] = (n1*M2 - n2*M1*M1)*iT2
f = plt.figure(figsize=(15, 8)); # plot the calculated values
sp = f.add_subplot(1, 2, 1 );
plt.plot(T, Energy, 'o', color="#348ABD", label='simulation');
plt.plot(T, -np.tanh(1.0/T), color="#A60628", lw=2,label='analytical')
plt.xlabel("Temperature (T)", fontsize=20);
plt.ylabel("Energy ", fontsize=20); plt.legend(fontsize=16)
sp = f.add_subplot(1, 2, 2 );
plt.plot(T, SpecificHeat, 'o', color="#348ABD", label='simulation');
plt.plot(T, (1.0/T**2)*(np.cosh(1.0/T))**(-2), color="#A60628", lw=2,label='analytical')
plt.xlabel("Temperature (T)", fontsize=20); plt.ylim([-0.05, .56])
plt.ylabel("Specific Heat ", fontsize=20); plt.legend(fontsize=16)
```
| github_jupyter |
# Module 6: Convolutions by examples
We'll build our first Convolutional Neural Network (CNN) from scratch.
## 1. Preparations
```
%matplotlib inline
import math,sys,os,numpy as np
from numpy.linalg import norm
from matplotlib import pyplot as plt
import torch
import torchvision
from torchvision import models,transforms,datasets
```
Download MNIST data on disk and convert it to pytorch compatible formating.
```torchvision.datasets``` features support (download, formatting) for a collection of popular datasets. The list of available datasets in ```torchvision``` can be found [here](http://pytorch.org/docs/master/torchvision/datasets.html).
Note that the download is performed only once. The function will always check first if the data is already on disk.
```
root_dir = './data/MNIST/'
torchvision.datasets.MNIST(root=root_dir,download=True)
```
MNIST datasets consists of small images of hand-written digits. The images are grayscale and have size 28 x 28. There are 60,000 training images and 10,000 testing images.
```
train_set = torchvision.datasets.MNIST(root=root_dir, train=True, download=True)
```
Define and initialize a data loader for the MNIST data already downloaded on disk.
```
MNIST_dataset = torch.utils.data.DataLoader(train_set, batch_size=1, shuffle=True, num_workers=1)
```
For the current notebook, we can format data as _numpy ndarrays_ which are easier to plot in matplotlib. The same operations can be easily performed on _pytorch Tensors_.
```
images = train_set.data.numpy().astype(np.float32)/255
labels = train_set.targets.numpy()
print(images.shape,labels.shape)
```
## 2. Data visualization
For convenience we define a few functions for formatting and plotting our image data
```
# plot multiple images
def plots(ims, interp=False, titles=None):
ims=np.array(ims)
mn,mx=ims.min(),ims.max()
f = plt.figure(figsize=(12,24))
for i in range(len(ims)):
sp=f.add_subplot(1, len(ims), i+1)
if not titles is None: sp.set_title(titles[i], fontsize=18)
plt.imshow(ims[i], interpolation=None if interp else 'none', vmin=mn,vmax=mx)
# plot a single image
def plot(im, interp=False):
f = plt.figure(figsize=(3,6), frameon=True)
plt.imshow(im, interpolation=None if interp else 'none')
plt.gray()
plt.close()
plot(images[5000])
labels[5000]
plots(images[5000:5005], titles=labels[5000:5005])
```
## 3. A simple classifier
In this section we will construct a basic binary classifier.
Our classifier will tell us whether a given image depicts a _one_ or an _eight_.
We fetch all images from the _eight_ class and from the _one_ class.
```
n=len(images)
eights=[images[i] for i in range(n) if labels[i]==8]
ones=[images[i] for i in range(n) if labels[i]==1]
len(eights), len(ones)
plots(eights[:5])
plots(ones[:5])
```
We keep the first 1000 digits for the test set and we average all the remaining digits.
```
raws8 = np.mean(eights[1000:],axis=0)
plot(raws8)
```
We now do the same thing with the ones:
```
raws1 = np.mean(ones[1000:],axis=0)
plot(raws1)
```
We built a 'typical representative' of the eights and a 'typical representative' of the ones. Now for a new sample from the test set, we compute the distance between this sample and our two representatives and classify the sample with the label of the closest representative.
For the distance between images, we just take the pixelwise squared distance.
```
# sum of squared distance
def sse(a,b): return ((a-b)**2).sum()
# return 1 if closest to 8 and 0 otherwise
def is8_raw_n2(im): return 1 if sse(im,raws1) > sse(im,raws8) else 0
nb_8_predicted_8, nb_1_predicted_8 = [np.array([is8_raw_n2(im) for im in ims]).sum() for ims in [eights[:1000],ones[:1000]]]
nb_8_predicted_1, nb_1_predicted_1 = [np.array([(1-is8_raw_n2(im)) for im in ims]).sum() for ims in [eights[:1000],ones[:1000]]]
# just to check
print(nb_8_predicted_1+nb_8_predicted_8, nb_1_predicted_1+nb_1_predicted_8)
```
<img src="https://upload.wikimedia.org/wikipedia/commons/thumb/2/26/Precisionrecall.svg/1024px-Precisionrecall.svg.png" alt="Drawing" style="width: 500px;"/>
source [wikipedia](https://en.wikipedia.org/wiki/Precision_and_recall)
```
def compute_scores(nb_8_predicted_8,nb_8_predicted_1,nb_1_predicted_1,nb_1_predicted_8):
Precision_8 = nb_8_predicted_8/(nb_8_predicted_8+nb_1_predicted_8)
Recall_8 = nb_8_predicted_8/(nb_8_predicted_1+nb_8_predicted_8)
Precision_1 = nb_1_predicted_1/(nb_1_predicted_1+nb_8_predicted_1)
Recall_1 = nb_1_predicted_1/(nb_1_predicted_1+nb_1_predicted_8)
return Precision_8, Recall_8, Precision_1, Recall_1
Precision_8, Recall_8, Precision_1, Recall_1 = compute_scores(nb_8_predicted_8,nb_8_predicted_1,nb_1_predicted_1,nb_1_predicted_8)
print('precision 8:', Precision_8, 'recall 8:', Recall_8)
print('precision 1:', Precision_1, 'recall 1:', Recall_1)
print('accuracy :', (Recall_1+Recall_8)/2)
```
This is our baseline for our binary classification task. Now your task will be to do better with convolutions!
## 4. Filters and convolutions
Let start with this visual explanation of [Interactive image kernels](http://setosa.io/ev/image-kernels/)
In some fields, convolution or filtering can be better understood as _correlations_.
In practice we slide the filter matrix over the image (a bigger matrix) always selecting patches from the image with the same size as the filter. We compute the dot product between the filter and the image patch and store the scalar response which reflects the degree of similarity/correlation between the filter and image patch.
Here is a simple 3x3 filter, ie a 3x3 matrix (see [Sobel operator](https://en.wikipedia.org/wiki/Sobel_operator) for more examples)
```
top=[[-1,-1,-1],
[ 1, 1, 1],
[ 0, 0, 0]]
plot(top)
```
We now create a toy image, to understand how convolutions operate.
```
cross = np.zeros((28,28))
cross += np.eye(28)
for i in range(4):
cross[12+i,:] = np.ones(28)
cross[:,12+i] = np.ones(28)
plot(cross)
```
Our `top` filter should highlight top horizontal border in the image.
```
from scipy.ndimage.filters import convolve, correlate
corr_cross = correlate(cross,top)
plot(corr_cross)
?correlate
```
What is done on the border of the image?
## Padding
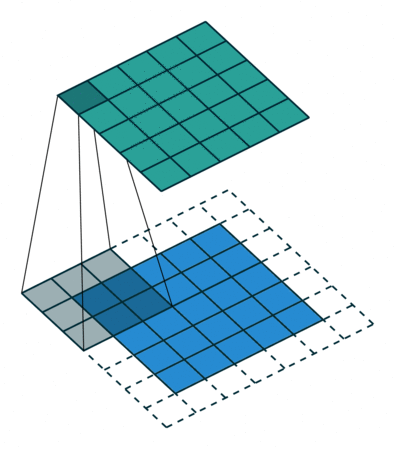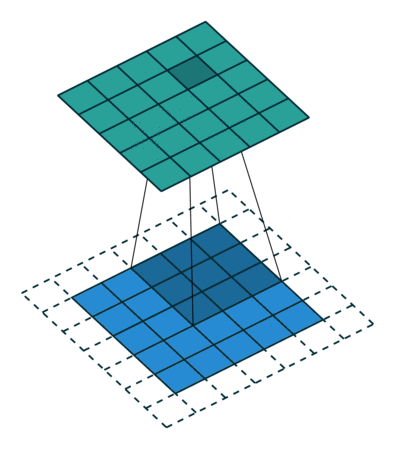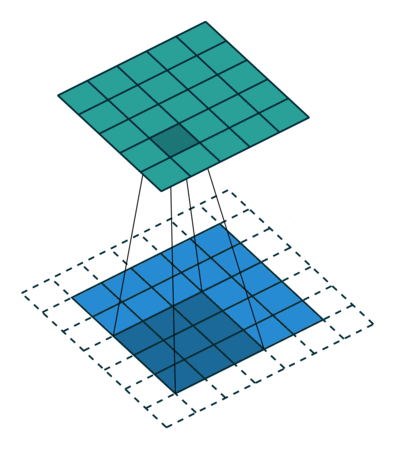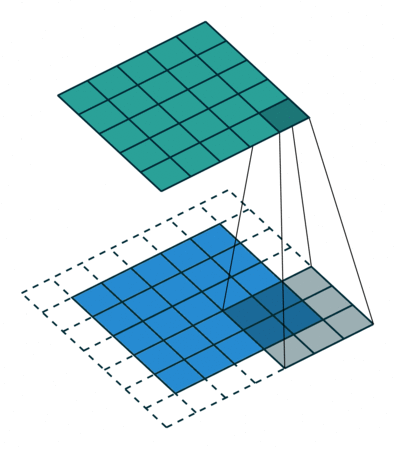
source: [Convolution animations](https://github.com/vdumoulin/conv_arithmetic/blob/master/README.md)
```
# to see the role of padding
corr_cross = correlate(cross,top, mode='constant')
plot(corr_cross)
corrtop = correlate(images[5000], top)
plot(corrtop)
```
By rotating the filter with 90 degrees and calling the ```convolve``` function we get the same response as with the previously called ```correlate``` function.
```
np.rot90(top, 1)
convtop = convolve(images[5000], np.rot90(top,2))
plot(convtop)
np.allclose(convtop, corrtop)
```
Let's generate a few more variants of our simple 3x3 filter
```
straights=[np.rot90(top,i) for i in range(4)]
plots(straights)
```
We proceed similarly to generate a set of filters with a different behavior
```
br=[[ 0, 0, 1],
[ 0, 1,-1.5],
[ 1,-1.5, 0]]
diags = [np.rot90(br,i) for i in range(4)]
plots(diags)
```
We can compose filters to obtain more complex patterns
```
rots = straights + diags
corrs_cross = [correlate(cross, rot) for rot in rots]
plots(corrs_cross)
rots = straights + diags
corrs = [correlate(images[5000], rot) for rot in rots]
plots(corrs)
```
Next we illustrate the effect of downsampling.
We select the most basic downsampling technique: __max pooling__. We keep only the maximum value for sliding windows of size ```7x7```.
__Max pooling__ is a handy technique with a few useful perks:
- since it selects the maximum values it ensures invariance to translations
- reducing the size is helpful since data becomes more compact and easier to compare
- we will see later in this course that since max pooling reduces the size of our images, the operations performed later on in the network have bigger receptive field / concern a bigger patch in the input image and allow the discovery of higher level patterns.
```
import skimage
from skimage.measure import block_reduce
def pool(im): return block_reduce(im, (7,7), np.max)
plots([pool(im) for im in corrs])
```
We now build a classifier with convolutions.
To this end we select a set of training images depicting _eights_ and _ones_, we convolve them with our set of filters, pool them and average them for each class and filter. We will thus obtain a set of _representative_ signatures for _eights_ and for _ones_.
Given a new test image we compute its features by convolution and pooling with the same filters and then compare them with the _representative_ features. The class with the most _similar_ features is chosen as prediction.
We keep 1000 images of _eight_ for the test set and use the remaining ones for the training: we convolve them with our bank of filters, perform max pooling on the responses and store them in ```pool8```.
```
pool8 = [np.array([pool(correlate(im, rot)) for im in eights[1000:]]) for rot in rots]
len(pool8), pool8[0].shape
```
We plot the result of the first filter+pooling on the first 5 _eights_ in our set.
```
plots(pool8[0][0:5])
```
For the 4 first _eights_ in our set, we plot the result of the 8 filters+pooling
```
plots([pool8[i][0] for i in range(8)])
plots([pool8[i][1] for i in range(8)])
plots([pool8[i][2] for i in range(8)])
plots([pool8[i][3] for i in range(8)])
```
We normalize the data in order to smoothen activations and bring them to similar ranges of values
```
def normalize(arr): return (arr-arr.mean())/arr.std()
```
Next we compute the average _eight_ by averaging all responses for each filter from _rots_.
```
filts8 = np.array([ims.mean(axis=0) for ims in pool8])
filts8 = normalize(filts8)
```
We should obtain a set of canonical _eights_ responses for each filter.
```
plots(filts8)
```
We proceed similarly with training samples from the _one_ class and plot the canonical _ones_.
```
pool1 = [np.array([pool(correlate(im, rot)) for im in ones[1000:]]) for rot in rots]
filts1 = np.array([ims.mean(axis=0) for ims in pool1])
filts1 = normalize(filts1)
plots(filts1)
```
Do you notice any differences between ```filts8``` and ```filts1```? Which ones?
We define a function that correlates a given image with all filters from ```rots``` and max pools the responses.
```
def pool_corr(im): return np.array([pool(correlate(im, rot)) for rot in rots])
plots(pool_corr(eights[1000]))
#check
plots([pool8[i][0] for i in range(8)])
np.allclose(pool_corr(eights[1000]),[pool8[i][0] for i in range(8)])
# function used for a voting based classifier that will indicate which one of the
# two classes is most likely given the sse distances
# n2 comes from norm2
# is8_n2 returns 1 if it thinks it's an eight and 0 otherwise
def is8_n2(im): return 1 if sse(pool_corr(im),filts1) > sse(pool_corr(im),filts8) else 0
```
We perform a check to see if our function actually works. We correlate the an image of _eight_ with ```filts8``` and ```filts1```. It should give smaller distance for the _eights_.
```
sse(pool_corr(eights[0]), filts8), sse(pool_corr(eights[0]), filts1)
plot(eights[0])
```
We now test our classifier on the 1000 images of _eights_ and 1000 images of _ones_
```
nb_8_predicted_8, nb_1_predicted_8 = [np.array([is8_n2(im) for im in ims]).sum() for ims in [eights[:1000],ones[:1000]]]
nb_8_predicted_1, nb_1_predicted_1 = [np.array([(1-is8_n2(im)) for im in ims]).sum() for ims in [eights[:1000],ones[:1000]]]
Precisionf_8, Recallf_8, Precisionf_1, Recallf_1 = compute_scores(nb_8_predicted_8,nb_8_predicted_1,nb_1_predicted_1,nb_1_predicted_8)
print('precision 8:', Precisionf_8, 'recall 8:', Recallf_8)
print('precision 1:', Precisionf_1, 'recall 1:', Recallf_1)
print('accuracy :', (Recallf_1+Recallf_8)/2)
print('accuracy baseline:', (Recall_1+Recall_8)/2)
```
We improved the accuracy while reducing the embedding size from a $28\times 28 = 784$ vector to a $4\times 4\times 8 = 128$ vector.
We have successfully built a classifier for _eights_ and _ones_ using features extracted with a bank of pre-defined features and a set of training samples.
## 5. Practicals: improving classification with Convolutional Neural Net
You will now build a neural net that will learn the weights of the filters.
The first layer of your network will be a convolutional layer with $8$ filters of size $3\times 3$. Then you will apply a Max Pooling layer to reduce the size of the image to $4\times 4$ as we did above. This will produce a (once flatten) a vector of size $128 = 4\times 4\times 8$. From this vector, you need to predict if the corresponding input is a $1$ or a $8$. So you are back to a classification problem as seen in previous lesson.
You need to fill the code written below to construct your CNN. You will need to look for documentation about [torch.nn](https://pytorch.org/docs/stable/nn.html) in the Pytorch doc.
```
import torch.nn as nn
import torch.nn.functional as F
class classifier(nn.Module):
def __init__(self):
super(classifier, self).__init__()
# fill the missing entries below
self.conv1 = nn.Conv2d(in_channels=1, out_channels=8, kernel_size=3, padding=?)
self.fc = nn.Linear(in_features=128, out_features=2)
def forward(self,x):
# implement your network here, use F.max_pool2d, F.log_softmax and do not forget to flatten your vector
x = self.conv1(x)
#
# Your code here
#
#
return x
conv_class = classifier()
```
Your code should work fine on a batch of 3 images.
```
batch_3images = train_set.data[0:2].type(torch.FloatTensor).resize_(3, 1, 28, 28)
#conv_class(batch_3images)
```
The following lines of code implement a data loader for the train set and the test set. No modification is needed.
```
bs = 64
l8 = np.array(0)
eights_dataset = [[torch.from_numpy(e.astype(np.float32)).unsqueeze(0), torch.from_numpy(l8.astype(np.int64))] for e in eights]
l1 = np.array(1)
ones_dataset = [[torch.from_numpy(e.astype(np.float32)).unsqueeze(0), torch.from_numpy(l1.astype(np.int64))] for e in ones]
train_dataset = eights_dataset[1000:] + ones_dataset[1000:]
test_dataset = eights_dataset[:1000] + ones_dataset[:1000]
train_loader = torch.utils.data.DataLoader(train_dataset,
batch_size=bs, shuffle=True)
test_loader = torch.utils.data.DataLoader(test_dataset,
batch_size=bs, shuffle=True)
```
You need now to code the training loop. Store the loss and accuracy for each epoch.
```
def train(model,data_loader,loss_fn,optimizer,n_epochs=1):
model.train(True)
loss_train = np.zeros(n_epochs)
acc_train = np.zeros(n_epochs)
for epoch_num in range(n_epochs):
running_corrects = 0.0
running_loss = 0.0
size = 0
for data in data_loader:
inputs, labels = data
bs = labels.size(0)
#
#
# Your code here
#
#
size += bs
epoch_loss = running_loss.item() / size
epoch_acc = running_corrects.item() / size
loss_train[epoch_num] = epoch_loss
acc_train[epoch_num] = epoch_acc
print('Train - Loss: {:.4f} Acc: {:.4f}'.format(epoch_loss, epoch_acc))
return loss_train, acc_train
conv_class = classifier()
# choose the appropriate loss
loss_fn =
# your SGD optimizer
learning_rate = 1e-3
optimizer_cl =
# and train for 10 epochs
l_t, a_t = train(conv_class,train_loader,loss_fn,optimizer_cl,n_epochs = 10)
```
Let's learn for 10 more epochs
```
l_t1, a_t1 = train(conv_class,train_loader,loss_fn,optimizer_cl,n_epochs = 10)
```
Our network seems to learn but we now need to check its accuracy on the test set.
```
def test(model,data_loader):
model.train(False)
running_corrects = 0.0
running_loss = 0.0
size = 0
for data in data_loader:
inputs, labels = data
bs = labels.size(0)
#
# Your code here
#
size += bs
print('Test - Loss: {:.4f} Acc: {:.4f}'.format(running_loss / size, running_corrects.item() / size))
test(conv_class,test_loader)
```
Change the optimizer to Adam.
How many parameters did your network learn?
You can see them as follows:
```
for m in conv_class.children():
print('weights :', m.weight.data)
print('bias :', m.bias.data)
for m in conv_class.children():
T_w = m.weight.data.numpy()
T_b = m.bias.data.numpy()
break
plots([T_w[i][0] for i in range(8)])
T_b
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import geopandas as gpd
import psycopg2
from geoalchemy2 import Geometry, WKTElement
from sqlalchemy import *
from shapely.geometry import MultiPolygon
from zipfile import ZipFile
import requests
import sys
import yaml
with open('../../config/postgres.yaml') as f:
engine_configs = yaml.load(f, Loader=yaml.FullLoader)
try:
engine = create_engine('postgresql://{username}:{password}@{host}:{port}/{dbname}'.format(**engine_configs))
except Exception as e:
print("Uh oh, can't connect. Invalid dbname, user or password?")
print(e)
def process_geometry_SQL_insert(gdf):
gdf['geom'] = gdf['geometry'].apply(lambda x: WKTElement((MultiPolygon([x]) if x.geom_type == 'Polygon' else x).wkt, srid=4326))
gdf = gdf.drop('geometry', 1)
return gdf
# Often when reading in a ShapeFile from Basemap, you'll get: "ValueError: readshapefile can only handle 2D shape types"
# A trick can be to convert your geometry in your GeoPandas Dataframe and restoring the new flattened 2D geometry
# series back into a shapefile and try again.
# edit from http://stackoverflow.com/questions/33417764/basemap-readshapefile-valueerror
from shapely.geometry import Polygon, MultiPolygon, shape, Point
def convert_3D_2D(geometry):
'''
Takes a GeoSeries of 3D Multi/Polygons (has_z) and returns a list of 2D Multi/Polygons
'''
new_geo = []
for p in geometry:
if p.has_z:
if p.geom_type == 'Polygon':
lines = [xy[:2] for xy in list(p.exterior.coords)]
new_p = Polygon(lines)
new_geo.append(new_p)
elif p.geom_type == 'MultiPolygon':
new_multi_p = []
for ap in p:
lines = [xy[:2] for xy in list(ap.exterior.coords)]
new_p = Polygon(lines)
new_multi_p.append(new_p)
new_geo.append(MultiPolygon(new_multi_p))
return new_geo
CITY='chicago'
```
### Neighborhoods
```
sql = """INSERT INTO spatial_groups (city, core_geom, core_id, lower_ids, spatial_name, approx_geom)
SELECT a.city, a.core_geom, a.core_id, array_agg(a.core_id), 'core', ST_multi(a.core_geom)
FROM spatial_groups a
where a.city='{city}' and a.spatial_name = 'ego'
GROUP BY a.core_id, a.core_geom, a.city;
""".format(city=CITY, tempname=CITY.lower())
result = engine.execute(text(sql))
```
## Land use
```
land_gdf = gpd.read_file('zip://../../data/chicago/land_use/land_use.zip', dtype={'LANDUSE': str})
land_gdf = land_gdf.to_crs({'init': 'epsg:4326'})
land_gdf.head()
land_gdf['landuse'] = 'none'
land_gdf.loc[(land_gdf['LANDUSE'].str[:2].isin({'11'})) | (land_gdf['LANDUSE'].isin({'1216'})), 'landuse'] = 'residential'
land_gdf.loc[(land_gdf['LANDUSE'].str[:2].isin({'12', '13', '14', '15', '20'})) & (~land_gdf['LANDUSE'].isin({'1510', '1511', '1512', '1520', '1550', '1561', '1565'})), 'landuse'] = 'commercial'
land_gdf.loc[land_gdf['LANDUSE'].str[:1].isin({'3'}), 'landuse'] = 'recreational'
land_gdf.loc[land_gdf['LANDUSE'].str[:1].isin({'4'}), 'landuse'] = 'vacant'
land_gdf.head()
```
## Net area
```
land_gdf_unique = land_gdf.copy()
land_gdf_unique.loc[:, 'x'] = land_gdf_unique.geometry.centroid.x
land_gdf_unique.loc[:, 'y'] = land_gdf_unique.geometry.centroid.y
land_gdf_unique = land_gdf_unique.drop_duplicates(subset=['x', 'y'])[['geometry', 'landuse']]
ins_gdf = process_geometry_SQL_insert(land_gdf_unique)
ins_gdf.to_sql('temptable_unique_{}'.format(CITY.lower()), engine, if_exists='replace', index=False, dtype={'geom': Geometry('MultiPolygon', srid=4326)})
sql = """
UPDATE temptable_unique_{tempname} p SET geom=ST_Multi(ST_buffer(p.geom, 0.0))
WHERE NOT ST_Isvalid(p.geom);
""".format(city=CITY, tempname=CITY.lower())
result = engine.execute(text(sql))
## This deletes the blocks that are related to streets
sql = """
DELETE FROM block b
WHERE city='{city}' and NOT EXISTS (select * from temptable_unique_{tempname} t where st_intersects(t.geom, b.geom) and t.landuse <> 'none');
""".format(city=CITY, tempname=CITY.lower())
result = engine.execute(text(sql))
sql = """
DELETE
FROM temptable_unique_{tempname} t
USING unused_areas u
WHERE u.city = '{city}' AND ST_Intersects(u.geom, t.geom) AND (NOT ST_Touches(u.geom, t.geom))
AND (ST_Contains(u.geom, t.geom) OR ST_AREA(ST_Intersection(t.geom, u.geom))/ST_Area(t.geom) > 0.5);
""".format(city=CITY, tempname=CITY.lower())
result = engine.execute(text(sql))
sql = """
INSERT INTO spatial_groups_net_area (sp_id, city, spatial_name, used_area)
SELECT sp_id, city, spatial_name, SUM(ST_Area(ST_Intersection(s.approx_geom, t.geom)::geography))/1000000.
FROM temptable_unique_{tempname} t
INNER JOIN spatial_groups s ON ST_Intersects(s.approx_geom, t.geom) AND NOT ST_Touches(s.approx_geom, t.geom)
WHERE s.city = '{city}' AND s.spatial_name='core'
GROUP BY sp_id, city, spatial_name;
""".format(city=CITY, tempname=CITY.lower())
result = engine.execute(text(sql))
```
## Refresh materialized views
```
sql = """
REFRESH MATERIALIZED VIEW spatial_groups_unused_areas;
"""
result = engine.execute(text(sql))
2
```
| github_jupyter |
```
import numpy as np
import matplotlib.pyplot as plt
```
### Least Square Method using Normal Equation Solution
### 正規方程式を用いて最小二乗法を行う
適当にランダムで線形データにノイズが乗ったようなものを作成する
```
X = 2 * np.random.rand(100, 1) # Randomize with a Uniform Distribution
y = 4 + 3 *X + np.random.randn(100, 1) # Randomize with a Normalized Distribution, Y = 4 + 3X + Noise
plt.scatter(X, y)
plt.xlabel('X')
plt.ylabel('y')
plt.show()
X_b = np.c_[np.ones((100, 1)), X] # Add 1 to heads of each row of X. This is constant term which equivalent to b of y= ax + b
theta_best = np.linalg.inv(X_b.T.dot(X_b)).dot(X_b.T).dot(y)
print(theta_best)
```
真値は4, 3なのでまあまあ近い。<br>
次に、予測結果をplotしてみる。
```
X_new = np.array([[0], [2]])
X_new_b = np.c_[np.ones((2,1)), X_new]
y_predict = X_new_b.dot(theta_best)
plt.plot(X_new, y_predict, 'r-')
plt.plot(X, y, 'b.')
plt.axis([0,2, 0, 15])
plt.show()
```
いい感じに回帰できてる<br>
いま自分でコードを書いたけど、scikit-learnなら次のようになる
```
from sklearn.linear_model import LinearRegression
lin_reg = LinearRegression()
lin_reg.fit(X, y) # fit method returns Bias Param to ingercept_ member variable, and other Weights to coef_ member variable
print(lin_reg.intercept_, lin_reg.coef_)
```
手で書いたやつと同じ結果になった。<br>
予測をするならこちら、先ほどのグラフの赤線の左端と右端になるはず。
```
print(lin_reg.predict(X_new))
```
正規方程式の特徴:<br>
① 特徴量の数nに対して$O(n^2)$<br>
② サンプルデータ数mに対して$O(m)$<br>
→ 特徴量やサンプル数が非常に多い場合は正規方程式は向かない、その場合は勾配法を使う
### Gradient Descent
### 勾配法
Deep Learningでも使ってる方法<br>
線形回帰に最小二乗誤差(Mean Square Error, MSE)を適用する時凸関数であることが保証されている→ 学習率が十分小さく長い時間計算して良いなら必ず大域的最適解に収束する
Batch Gradient Descent<br>
バッチ勾配降下法 データ全部使う
```
eta = 0.1 # Learning Rate
n_iterations = 1000
m = 100 # Number of Samples
theta = np.random.randn(2,1) # Initialize Parameters with a Normal Distribution
for iteration in range(n_iterations):
gradients = 2/m * X_b.T.dot(X_b.dot(theta) - y) # This gradient is from MSE definition
theta = theta - eta * gradients
print(theta)
```
正規方程式で見つけた最適解と一致した→ 勾配法が正しく機能した
Stocastic Gradient Descent (SGD)<br>
確率的勾配降下法<br>
データがめちゃくちゃ多くなると全部のデータ見てパラメータ更新するのは時間がかかりすぎる。そこでサンプル一つにつき更新を行う。<br>
サンプル一つずつ重みを更新するため、全体のコスト関数は上下を繰り返す。BGDなら必ず下がったが。。。<br>
そうすると最適値付近でも値が暴れるので、学習率を少しずつ下げたりしたい。(学習スケジュール)
```
n_epochs = 50
t0, t1 = 5, 50
def learning_schedule(t):
return t0 / (t + t1)
theta = np.random.randn(2, 1)
for epoch in range(n_epochs):
for i in range(m):
random_index = np.random.randint(m) # random int number within 0 ~ m
xi = X_b[random_index: random_index+1]
yi = y[random_index: random_index + 1]
gradients = 2 * xi.T.dot(xi.dot(theta) - yi)
eta = learning_schedule(epoch * m + i)
theta = theta - eta*gradients
print(theta)
```
結構ぶれてるけど近い<br>
これも同様にscikit-learnで書いてみると
```
from sklearn.linear_model import SGDRegressor
sgd_reg = SGDRegressor(n_iter=50, penalty=None, eta0=0.1) # penalty is a parameter for regulatization, and the learning schedule is default one
sgd_reg.fit(X, y.ravel()) # ravel is a method like flatten
print(sgd_reg.intercept_, sgd_reg.coef_)
```
ミニバッチ勾配降下法もscikit-learnにはある。BGDとSGDの中間。
### Polynomial Regression
### 多項式回帰
```
m = 100
X = 6 * np.random.rand(m, 1) - 3
y = 0.5 * X**2 + X + 2*np.random.randn(m,1)
plt.plot(X, y, 'b.')
plt.xlabel('X')
plt.ylabel('y')
plt.show()
from sklearn.preprocessing import PolynomialFeatures
poly_features = PolynomialFeatures(degree = 2, include_bias = False) # Choose degree of polynomial
X_poly = poly_features.fit_transform(X)
print(X[0]) # Data
print(X_poly[0]) # Calculated Polynomial Feature
```
生データXに対して高次の特徴量X_polyが計算されている<br>
回帰の計算方法は特徴量が増えても変わらないので、後のコードは線形回帰の時と同じ
```
lin_reg = LinearRegression()
lin_reg.fit(X_poly, y)
print(lin_reg.intercept_, lin_reg.coef_)
```
真値はそれぞれ[0, 1, 0.5]なのでまあこんなもんかという感じ<br>
過学習させるのが簡単なのでやってみる
```
poly_features_over = PolynomialFeatures(degree = 30, include_bias = False) # Choose degree of polynomial
X_poly_over = poly_features_over.fit_transform(X)
lin_reg_over = LinearRegression()
lin_reg_over.fit(X_poly_over, y)
X_new = np.reshape(np.arange(-3, 3, 0.1), [60, 1])
y_predict = lin_reg.predict(poly_features.fit_transform(X_new))
y_predict_over = lin_reg_over.predict(poly_features_over.fit_transform(X_new))
plt.plot(X, y, 'b.')
plt.plot(X_new, y_predict, 'g')
plt.plot(X_new, y_predict_over, 'r')
plt.ylim([-5, 10])
plt.xlim([-3, 3])
plt.legend(('Data', '2', '30'))
plt.show()
```
さすがに30次の多項式では過学習してる
一般に過学習をどうやって判断するか?<br>
→ 与えるデータが増えた時にほとんど成績に変化がなければ、過小適合<br>
→ Training set と Validation setのさが大きければ過学習
ここでデータを増やした時にどれだけLossが下がるか見る関数を定義
```
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
def plot_learning_curves(model, X, y):
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2)
train_errors, val_errors = [], []
for m in range(1, len(X_train)):
model.fit(X_train[:m], y_train[:m])
y_train_predict = model.predict(X_train[:m])
y_val_predict = model.predict(X_val)
train_errors.append(mean_squared_error(y_train_predict, y_train[:m]))
val_errors.append(mean_squared_error(y_val_predict, y_val))
plt.plot(np.sqrt(train_errors), 'r', label='train')
plt.plot(np.sqrt(val_errors), 'b', label='val')
plt.legend(('train', 'val'))
plt.xlabel('Training set size')
plt.ylabel('RMSE')
plt.ylim([0, 4])
```
これを使ってとりあえず線形回帰で確認してみる
```
lin_reg = LinearRegression()
plot_learning_curves(lin_reg, X, y)
```
いつもみている学習曲線とは違うので注意。これは横軸がデータのサイズである。Deep Learning系は計算が重いので、この方法は使わないのだと思われる。<br>
しかし、パラメータが少ない対象への機械学習ならこの方法が有効と考える。<br>
この例ではデータ量が増加してもLossは下がらず、またTrainとValidationの差が無い。従って、過小適合していると考えられる。
```
from sklearn.pipeline import Pipeline
polynomial_regression = Pipeline([
("poly_features", PolynomialFeatures(degree=10, include_bias = False)),
("lin_reg", LinearRegression())
])
plot_learning_curves(polynomial_regression, X, y)
```
データサイズが大きくなるとtrainとvalの差がなくなっている→ 汎化性能が向上している→ データを増やせば性能が上がる
### Ridge Regression
### リッジ回帰
回帰する関数の形が特殊でわけはなく、Lossに正則化項を加えていることが特徴。線形回帰+正則化。コスト関数を$J(\theta)$として<br>
$J(\theta) = MSE(\theta) + \alpha \frac{1}{2} \Sigma_{i=1}^n{\theta_i^2}$<br>
正則化項は特徴量のスケールの影響を強く受けるので使う時は必ずスケーリングをしておくこと・
```
from sklearn.linear_model import Ridge
poly_features_over = PolynomialFeatures(degree = 10, include_bias = False) # Choose degree of polynomial
X_over = poly_features_over.fit_transform(X)
alpha_array = [1e-1, 10]
y_predict_over = np.zeros((len(alpha_array), len(X_new), 1))
for i, alph in enumerate(alpha_array):
ridge_reg = Ridge(alpha=alph, solver = 'cholesky') # Solve Ridge regression with closed form
ridge_reg.fit(X_over, y)
y_predict_over[i] = ridge_reg.predict(poly_features_over.fit_transform(X_new))
plt.plot(X, y, 'b.')
plt.plot(X_new, y_predict_over[0], 'g')
plt.plot(X_new, y_predict_over[1], 'r')
plt.ylim([-5, 10])
plt.xlim([-3, 3])
plt.legend(('Data', 'alhpa='+str(alpha_array[0]), 'alpha='+str(alpha_array[1])))
plt.show()
```
10次の回帰も滑らかになっていい感じ
### Lasso Regression
正則化項が2乗でなく $l1$ノルム(この場合絶対値)<br>
$J(\theta) = MSE(\theta) + \alpha\Sigma_{i=1}^n{|\theta_i|}$<br>
Ridge回帰と比較するとあまり寄与しない特徴量の係数がほぼ0になるように訓練される(係数が疎になる)特徴がある
```
from sklearn.linear_model import Lasso
poly_features_over = PolynomialFeatures(degree = 10, include_bias = False) # Choose degree of polynomial
X_over = poly_features_over.fit_transform(X)
alpha_array = [1e-7, 10]
y_predict_over = np.zeros((len(alpha_array), len(X_new)))
for i, alph in enumerate(alpha_array):
ridge_reg = Lasso(alpha=alph) # Solve Ridge regression with closed form
ridge_reg.fit(X_over, y)
y_predict_over[i] = ridge_reg.predict(poly_features_over.fit_transform(X_new))
plt.plot(X, y, 'b.')
plt.plot(X_new, y_predict_over[0], 'g')
plt.plot(X_new, y_predict_over[1], 'r')
plt.ylim([-5, 10])
plt.xlim([-3, 3])
plt.legend(('Data', 'alhpa='+str(alpha_array[0]), 'alpha='+str(alpha_array[1])))
plt.show()
```
たしかにRidgeの$\alpha=10$とLassoの$\alpha=1e-7$を比べても、Lassoの方が高次の項消えてる感じがする
### Elastic Net
Ridge回帰とLasso回帰の混ぜ合わせ、パラメータを調節できる。<br>
$J(\theta) = MSE(\theta) + r\alpha\Sigma_{i=1}^n{|\theta_i|} + \alpha \frac{1-r}{2} \Sigma_{i=1}^n{\theta_i^2}$
```
from sklearn.linear_model import ElasticNet
from sklearn.linear_model import Ridge
poly_features_over = PolynomialFeatures(degree = 10, include_bias = False) # Choose degree of polynomial
X_over = poly_features_over.fit_transform(X)
alph = 1
l1_ratio_array = [0, 0.5, 1]
y_predict_over = np.zeros((len(l1_ratio_array), len(X_new)))
for i, l1_ratio in enumerate(l1_ratio_array):
elastic_net = ElasticNet(alpha=alph, l1_ratio = l1_ratio) # Solve Ridge regression with closed form
elastic_net.fit(X_over, y)
y_predict_over[i] = elastic_net.predict(poly_features_over.fit_transform(X_new))
plt.plot(X, y, 'b.')
plt.plot(X_new, y_predict_over[0], 'g')
plt.plot(X_new, y_predict_over[1], 'r')
plt.plot(X_new, y_predict_over[2], 'b')
plt.ylim([-5, 10])
plt.xlim([-3, 3])
# plt.legend(('Data', 'r='+str(l1_ratio_array[0]), 'r='+str(l1_ratio_array[1]), 'r='+str(l1_ratio_array[2])))
plt.show()
```
↑ なんだかうまく動いてない感じがするが....<br>
正則化について、デフォはRidge。データの前提条件として、意味のある特徴量は少ないなど(スパースなモデルが欲しい時とか)あればLasso。<br>
しかし特徴量が多い時Lassoは変な挙動をすることがあるので一般的にはLassoを使うならElasticNetが良い。
### Logistic Regression
### ロジスティック回帰
$\sigma(t) = \frac{1}{1 + exp(-t)}$<br>
$J(\theta) = -\frac{1}{m}\Sigma_{i=1}^{m}[y^{(i)}log(p^{(i)}) + (1 - y^{(i)}) log(1-p^{(i)})]$
正規方程式みたいに閉形式で最適解はでないけど凸関数であることはわかっているので勾配法で解く<br>
分類器を作りたいのでIrisデータセットで試す
<img src="https://image.slidesharecdn.com/s06t1python4-151110035703-lva1-app6891/95/s06-t1-python4-28-638.jpg?cb=1447127885">
ロジスティック回帰は1クラスの分類器なので適当にirisのうちでVirginica種を見つけることを目標にする
```
from sklearn import datasets
from sklearn.linear_model import LogisticRegression
iris = datasets.load_iris() # load iris dataset
print(list(iris.keys()))
X_all = iris["data"]
y = (iris["target"] == 2).astype(np.int) # Virginica
log_reg = LogisticRegression()
log_reg.fit(X_all, y)
X = iris["data"][:,2:]
x0, x1, x2, x3 = np.meshgrid(
np.linspace(2.9, 7, 500).reshape(-1, 1),
np.linspace(0.8, 2.7, 200).reshape(-1, 1),
3.4,
1.5,
)
X_new = np.c_[x0.ravel(), x1.ravel(), x2.ravel(), x3.ravel()]
y_proba = log_reg.predict_proba(X_new)
plt.figure(figsize=(10, 4))
plt.plot(X[y==0, 0], X[y==0, 1], "b.")
plt.plot(X[y==1, 0], X[y==1, 1], "g.")
zz = y_proba[:, 1].reshape(x0.shape)
print(x1.shape)
contour = plt.contour(x0[:,:,0,0], x1[:,:,0,0], zz[:,:,0,0], cmap=plt.cm.brg)
left_right = np.array([2.9, 7])
boundary = -(log_reg.coef_[0][0] * left_right + log_reg.intercept_[0]) / log_reg.coef_[0][1]
plt.clabel(contour, inline=1, fontsize=12)
plt.plot(left_right, boundary, "k--", linewidth=3)
plt.xlabel("Petal length", fontsize=14)
plt.ylabel("Petal width", fontsize=14)
plt.axis([2.9, 7, 0.8, 2.7])
plt.show()
```
4つの特徴量のうちの2次元断面<br>
### まとめ<br>
深層学習以外の機械学習案件も多いので基本を見直してみました<br>
Lassoとかあんまり考えたことなかったけどちゃんと使い分けとか特徴があるのでそこを理解した上で適切な手法を選択したい。<br>
sklearnの関数はオプションのところで思わぬデフォ値になってたりするので注意しよう
| github_jupyter |

<a href="https://hub.callysto.ca/jupyter/hub/user-redirect/git-pull?repo=https%3A%2F%2Fgithub.com%2Fcallysto%2Fhackathon&branch=master&subPath=ColonizingMars/ChallengeTemplates/challenge-option-2-how-could-we-colonize-mars.ipynb&depth=1" target="_parent"><img src="https://raw.githubusercontent.com/callysto/curriculum-notebooks/master/open-in-callysto-button.svg?sanitize=true" width="123" height="24" alt="Open in Callysto"/></a>
# Data Scientist Challenge: How could we colonize Mars?
Use this notebook if you are interested in proposing ways to colonize Mars.
## How
Use data to answer questions such as:
1. How do we decide who will go?
population proportions, demographics, health, qualifications, genetic diversity
2. What do we need to bring?
3. What are some essential services?
4. What kinds of jobs should people do?
5. How do we feed people there?
Consider: supply, manage, distribute, connect
6. Where should we land?
7. What structures should we design and build?
8. Should we terraform Mars? How?
9. How should Mars be governed?
Pick as many questions from the above section (or come up with your own). Complete the sections within this notebook.
### Section I: About You
Double click this cell and tell us:
1. Your name
2. Your email address
3. Why you picked this challenge
4. The questions you picked
For example
1. Your name: Not-my Name
2. Your email address: not_my_real_address@domain.com
3. Why you picked this challenge: I don't think we should attempt to colonize Mars
4. The questions you picked: Why does humanity tend to colonize? Why not focus on making Earth a better place?
### Section II: The data you used
Please provide the following information:
1. Name of dataset
2. Link to dataset
3. Why you picked the dataset
If you picked multiple datasets, separate them using commas ","
```
# Use this cell to import libraries
import pandas as pd
import plotly_express as px
import numpy as np
# Use this cell to read the data - use the tutorials if you are not sure how to do this
```
### Section III: Data Analysis and Visualization
Use as many code cells as you need - remember to add a title, as well as appropriate x and y labels to your visualizations.
Ensure to briefly comment on what you see.
A sample is provided.
```
# Sample code
x_values = np.array([i for i in range(-200,200)])
y_values = x_values**3
px.line(x=x_values,
y=y_values,
title="Line plot of x and y values",
labels = {'x':'Independent Variable x','y':'Dependent Variable y'})
```
#### Observations
We see that the dependent variable y increases exponentially. Indeed, when the independent variable x = 10, then the dependent variable y = 1000, when the independent variable x = 100, then y = 1'000,000.
It appears the relationship between the two variables is as follows:
$$y = x^3$$
### Section IV: Conclusion
It is crucial that you connect what you learned via the dataset to the main question(s) you are asking.
Use this space to propose a solution to the question you picked. Ensure it is clear in your answer what area of development you chose to focus on and your proposed solution based on the dataset(s) you worked on.
[](https://github.com/callysto/curriculum-notebooks/blob/master/LICENSE.md)
| github_jupyter |
```
# Adds link to the scripts folder
import sys
import os
sys.path.append("../scripts/")
import matplotlib.pyplot as plt
import numpy as np
from trajectory import Trajectory, create_trajectory_list, filter, create_all_patient_trajectories
from hivevo.patients import Patient
import filenames
import copy
```
# Mean frequency over time
```
def get_mean_in_time(trajectories, nb_bins=15, freq_range=[0.4, 0.6]):
"""
Computes the mean frequency in time of a set of trajectories from the point they are seen in the freq_range window.
Returns the middle of the time bins and the computed frequency mean.
"""
# Create bins and select trajectories going through the freq_range
time_bins = np.linspace(-1000, 2000, nb_bins)
trajectories = [traj for traj in trajectories if np.sum(np.logical_and(
traj.frequencies >= freq_range[0], traj.frequencies < freq_range[1]), dtype=bool)]
# Offset trajectories to set t=0 at the point they are seen in the freq_range and adds all the frequencies / times
# to arrays for later computation of mean
t_traj = np.array([])
f_traj = np.array([])
for traj in trajectories:
idx = np.where(np.logical_and(traj.frequencies >=
freq_range[0], traj.frequencies < freq_range[1]))[0][0]
traj.t = traj.t - traj.t[idx]
t_traj = np.concatenate((t_traj, traj.t))
f_traj = np.concatenate((f_traj, traj.frequencies))
# Binning of all the data in the time bins
filtered_fixed = [traj for traj in trajectories if traj.fixation == "fixed"]
filtered_lost = [traj for traj in trajectories if traj.fixation == "lost"]
freqs, fixed, lost = [], [], []
for ii in range(len(time_bins) - 1):
freqs = freqs + [f_traj[np.logical_and(t_traj >= time_bins[ii], t_traj < time_bins[ii + 1])]]
fixed = fixed + [len([traj for traj in filtered_fixed if traj.t[-1] < time_bins[ii]])]
lost = lost + [len([traj for traj in filtered_lost if traj.t[-1] < time_bins[ii]])]
# Computation of the mean in each bin, active trajectories contribute their current frequency,
# fixed contribute 1 and lost contribute 0
mean = []
for ii in range(len(freqs)):
mean = mean + [np.sum(freqs[ii]) + fixed[ii]]
mean[-1] /= (len(freqs[ii]) + fixed[ii] + lost[ii])
return 0.5 * (time_bins[1:] + time_bins[:-1]), mean
```
## Region env
```
patient_names = ["p1", "p2", "p3", "p4", "p5", "p6", "p8", "p9", "p11"]
region = "env"
nb_bins = 15
fontsize = 16
trajectories = create_all_patient_trajectories(region, patient_names)
trajectories = [traj for traj in trajectories if traj.t[-1] != 0]
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
freq_range = [0.2, 0.4]
time_bins, mean_syn1 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn1 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.4, 0.6]
time_bins, mean_syn2 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn2 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.6, 0.8]
time_bins, mean_syn3 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn3 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
plt.figure(figsize=(14,10))
plt.plot(time_bins, mean_syn1, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn1, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn2, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn2, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn3, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn3, '.-', label="Non-synonymous")
plt.xlabel("Time [days]", fontsize=fontsize)
plt.ylabel("Frequency", fontsize=fontsize)
plt.legend(fontsize=fontsize)
plt.title(f"Region {region}", fontsize=fontsize)
plt.ylim([0,1])
plt.grid()
plt.show()
```
## Region pol
```
patient_names = ["p1", "p2", "p3", "p4", "p5", "p6", "p8", "p9", "p11"]
region = "pol"
nb_bins = 15
fontsize = 16
trajectories = create_all_patient_trajectories(region, patient_names)
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
freq_range = [0.2, 0.4]
time_bins, mean_syn1 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn1 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.4, 0.6]
time_bins, mean_syn2 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn2 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.6, 0.8]
time_bins, mean_syn3 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn3 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
plt.figure(figsize=(14,10))
plt.plot(time_bins, mean_syn1, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn1, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn2, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn2, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn3, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn3, '.-', label="Non-synonymous")
plt.xlabel("Time [days]", fontsize=fontsize)
plt.ylabel("Frequency", fontsize=fontsize)
plt.legend(fontsize=fontsize)
plt.title(f"Region {region}", fontsize=fontsize)
plt.ylim([0,1])
plt.grid()
plt.show()
```
## Region gag
```
patient_names = ["p1", "p2", "p3", "p4", "p5", "p6", "p8", "p9", "p11"]
region = "gag"
nb_bins = 15
fontsize = 16
trajectories = create_all_patient_trajectories(region, patient_names)
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
freq_range = [0.2, 0.4]
time_bins, mean_syn1 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn1 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.4, 0.6]
time_bins, mean_syn2 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn2 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
freq_range = [0.6, 0.8]
time_bins, mean_syn3 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn3 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
plt.figure(figsize=(14,10))
plt.plot(time_bins, mean_syn1, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn1, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn2, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn2, '.-', label="Non-synonymous")
plt.plot(time_bins, mean_syn3, '.-', label="Synonymous")
plt.plot(time_bins, mean_non_syn3, '.-', label="Non-synonymous")
plt.xlabel("Time [days]", fontsize=fontsize)
plt.ylabel("Frequency", fontsize=fontsize)
plt.legend(fontsize=fontsize)
plt.title(f"Region {region}", fontsize=fontsize)
plt.ylim([0,1])
plt.grid()
plt.show()
```
## Comparison between regions
```
patient_names = ["p1", "p2", "p3", "p4", "p5", "p6", "p8", "p9", "p11"]
freq_range = [0.4, 0.5]
nb_bins = 15
fontsize = 16
region = "env"
trajectories = create_all_patient_trajectories(region, patient_names)
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
time_bins, mean_syn1 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn1 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
region = "gag"
trajectories = create_all_patient_trajectories(region, patient_names)
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
time_bins, mean_syn2 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn2 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
region = "pol"
trajectories = create_all_patient_trajectories(region, patient_names)
syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == True])
non_syn_traj = copy.deepcopy([traj for traj in trajectories if traj.synonymous == False])
time_bins, mean_syn3 = get_mean_in_time(syn_traj, nb_bins, freq_range)
time_bins, mean_non_syn3 = get_mean_in_time(non_syn_traj, nb_bins, freq_range)
plt.figure(figsize=(14,10))
plt.plot(time_bins, mean_syn1, '.-', label="Syn env")
plt.plot(time_bins, mean_syn2, '.-', label="Syn gag")
plt.plot(time_bins, mean_syn3, '.-', label="Syn pol")
plt.xlabel("Time [days]", fontsize=fontsize)
plt.ylabel("Frequency", fontsize=fontsize)
plt.legend(fontsize=fontsize)
plt.title(f"Synonymous [0.4, 0.6]", fontsize=fontsize)
plt.ylim([0,1])
plt.grid()
plt.show()
plt.figure(figsize=(14,10))
plt.plot(time_bins, mean_non_syn1, '.-', label="Non-syn env")
plt.plot(time_bins, mean_non_syn2, '.-', label="Non-syn gag")
plt.plot(time_bins, mean_non_syn3, '.-', label="Non-syn pol")
plt.xlabel("Time [days]", fontsize=fontsize)
plt.ylabel("Frequency", fontsize=fontsize)
plt.legend(fontsize=fontsize)
plt.title(f"Non-synonymous [0.4, 0.6]", fontsize=fontsize)
plt.ylim([0,1])
plt.grid()
plt.show()
```
| github_jupyter |
```
%matplotlib inline
```
Autograd: Automatic Differentiation
===================================
Central to all neural networks in PyTorch is the ``autograd`` package.
Let’s first briefly visit this, and we will then go to training our
first neural network.
The ``autograd`` package provides automatic differentiation for all operations
on Tensors. It is a define-by-run framework, which means that your backprop is
defined by how your code is run, and that every single iteration can be
different.
Let us see this in more simple terms with some examples.
Tensor
--------
``torch.Tensor`` is the central class of the package. If you set its attribute
``.requires_grad`` as ``True``, it starts to track all operations on it. When
you finish your computation you can call ``.backward()`` and have all the
gradients computed automatically. The gradient for this tensor will be
accumulated into ``.grad`` attribute.
To stop a tensor from tracking history, you can call ``.detach()`` to detach
it from the computation history, and to prevent future computation from being
tracked.
To prevent tracking history (and using memory), you can also wrap the code block
in ``with torch.no_grad():``. This can be particularly helpful when evaluating a
model because the model may have trainable parameters with `requires_grad=True`,
but for which we don't need the gradients.
There’s one more class which is very important for autograd
implementation - a ``Function``.
``Tensor`` and ``Function`` are interconnected and build up an acyclic
graph, that encodes a complete history of computation. Each tensor has
a ``.grad_fn`` attribute that references a ``Function`` that has created
the ``Tensor`` (except for Tensors created by the user - their
``grad_fn is None``).
If you want to compute the derivatives, you can call ``.backward()`` on
a ``Tensor``. If ``Tensor`` is a scalar (i.e. it holds a one element
data), you don’t need to specify any arguments to ``backward()``,
however if it has more elements, you need to specify a ``gradient``
argument that is a tensor of matching shape.
```
import torch
```
Create a tensor and set requires_grad=True to track computation with it
```
x = torch.ones(2, 2, requires_grad=True)
print(x)
```
Do an operation of tensor:
```
y = x + 2
print(y)
```
``y`` was created as a result of an operation, so it has a ``grad_fn``.
```
print(y.grad_fn)
```
Do more operations on y
```
z = y * y * 3
out = z.mean()
print(z, out)
```
``.requires_grad_( ... )`` changes an existing Tensor's ``requires_grad``
flag in-place. The input flag defaults to ``False`` if not given.
```
a = torch.randn(2, 2)
a = ((a * 3) / (a - 1))
print(a.requires_grad)
a.requires_grad_(True)
print(a.requires_grad)
b = (a * a).sum()
print(b.grad_fn)
```
Gradients
---------
Let's backprop now
Because ``out`` contains a single scalar, ``out.backward()`` is
equivalent to ``out.backward(torch.tensor(1))``.
```
out.backward()
```
print gradients d(out)/dx
```
print(x.grad)
```
You should have got a matrix of ``4.5``. Let’s call the ``out``
*Tensor* “$o$”.
We have that $o = \frac{1}{4}\sum_i z_i$,
$z_i = 3(x_i+2)^2$ and $z_i\bigr\rvert_{x_i=1} = 27$.
Therefore,
$\frac{\partial o}{\partial x_i} = \frac{3}{2}(x_i+2)$, hence
$\frac{\partial o}{\partial x_i}\bigr\rvert_{x_i=1} = \frac{9}{2} = 4.5$.
You can do many crazy things with autograd!
```
x = torch.randn(3, requires_grad=True)
y = x * 2
while y.data.norm() < 1000:
y = y * 2
print(y)
gradients = torch.tensor([0.1, 1.0, 0.0001], dtype=torch.float)
y.backward(gradients)
print(x.grad)
```
You can also stop autograd from tracking history on Tensors
with ``.requires_grad=True`` by wrapping the code block in
``with torch.no_grad()``:
```
print(x.requires_grad)
print((x ** 2).requires_grad)
with torch.no_grad():
print((x ** 2).requires_grad)
```
**Read Later:**
Documentation of ``autograd`` and ``Function`` is at
https://pytorch.org/docs/autograd
| github_jupyter |
# CHAPTER 20 - Learning Probabilistic Models
### George Tzanetakis, University of Victoria
## WORKPLAN
The section number is based on the 4th edition of the AIMA textbook and is the suggested
reading for this week. Each list entry provides just the additional sections. For example the Expected reading include the sections listed under Basic as well as the sections listed under Expected. Some additional readings are suggested for Advanced.
1. Basic: Sections **20.1**, **20.2.1**, **20.2.2**, and **Summary**
2. Expected: Same as Basic plus **20.3**, **20.3.1**, **20.3.3**
3. Advanced: All the chapter including bibligraphical and historical notes
We have covered a variety of probabilistic models that model uncertainty and allow us to do inference in different ways. In this notebook we describe some of the ways we can estimate probabistic modesl from data.
These techniques provide the connection between statistics, probability, and machine learning.
The ideas are based on Chapter 20 of the Artificial Intelligence: a Modern Approach textbook and specifically section 20.2 Learning from Complete Data.
**Density estimation** refers to the task of learning the probabiity density function (for continuous models) or the probability distribution function (for discrete models) given some data that we assume was generated from that modal. **Complete data** means that we have data for all the **variables** in our model.
The most common type of learning is **parameter learning** where we assume a particular structure for our model and characterize it by estimating a set of parameters. For example we might assume a normal or Gaussian multi-variate distribution and estimate the mean vector and the covariance matrix that characterize it. As another example, we might be given the structure of a Bayesian network (in terms of parent/child coonditional relationships) and learn the conditional probabilty tables. We will also briefly discuss the problem of learning structure as well as non-parametric density estimation in which we don't need to make any assumptions about the model.
# Learning with Complete Data
## A random variable class
Define a helper random variable class based on the scipy discrete random variable functionality providing both numeric and symbolic RVs. You don't need to look at the implementation - the usage will be obvious through the examples below.
```
%matplotlib inline
import matplotlib.pyplot as plt
from scipy import stats
import numpy as np
class Random_Variable:
def __init__(self, name, values, probability_distribution):
self.name = name
self.values = values
self.probability_distribution = probability_distribution
if all(type(item) is np.int_ for item in self.values):
self.type = 'numeric'
self.rv = stats.rv_discrete(name = name,
values = (values, probability_distribution))
elif all(type(item) is str for item in values):
self.type = 'symbolic'
self.rv = stats.rv_discrete(name = name,
values = (np.arange(len(values)), probability_distribution))
self.symbolic_values = values
else:
self.type = 'undefined'
def sample(self,size):
if (self.type =='numeric'):
return self.rv.rvs(size=size)
elif (self.type == 'symbolic'):
numeric_samples = self.rv.rvs(size=size)
mapped_samples = [self.values[x] for x in numeric_samples]
return mapped_samples
def prob_of_value(self, value):
indices = np.where(self.values == value)
return self.probability_distribution[indices[0][0]]
```
# Likelihood of model given some data
First let's review the concept of likelihood of a model given some data
Lets start by creating a random variable corresponding to a 6-faced dice where there are two faces with the numbers 1,2 and 3 therefore each number appears with equal probability. We can generate random samples from this model.
```
values = np.int64([1, 2, 3])
probabilities = [2/6., 2/6., 2/6.]
dice1 = Random_Variable('dice1', values, probabilities)
samples = dice1.sample(30)
print(samples)
```
Lets also create a random variable where three of the faces have the number 2, 2 faces have the number 1 and 1 face has the number 3. We can also generate random samples from this model.
```
values = np.int_([1, 2, 3])
probabilities = [2./6, 3./6, 1./6]
dice2 = Random_Variable('dice2', values, probabilities)
samples = dice2.sample(30)
print(samples)
```
The likelihood of a sequence of samples given a model can be obtained by taking the product of the corresponding
probabilities. We can see that for this particular sequence of data the likelihood of the model for dice2 is higher. So if we have some data and some specific models we can select the model with the highest likelihood.
```
data = [1,2,2,1,1,3,1,2,3,2]
print(dice1.prob_of_value(1))
print(dice1.prob_of_value(3))
print(dice2.prob_of_value(3))
def likelihood(data, model):
likelihood = 1.0
for d in data:
likelihood *= model.prob_of_value(d)
return likelihood
print("Likelihood for dice1: %f" % likelihood(data,dice1))
print("Likelihood for dice2: %f" % likelihood(data,dice2))
```
Notice that even with only 10 values the likelihood gets relatively small and we can expect it will get smaller as the sequences of data get smaller. We can also use log-likelihood to avoid this problem.
```
data = [1,2,2,1,1,3,1,2,3,2]
print(dice1.prob_of_value(1))
print(dice1.prob_of_value(3))
print(dice2.prob_of_value(3))
def log_likelihood(data, model):
likelihood = 0.0
for d in data:
likelihood += np.log(model.prob_of_value(d))
return likelihood
print("Likelihood for dice1: %f" % log_likelihood(data,dice1))
print("Likelihood for dice2: %f" % log_likelihood(data,dice2))
```
In the case above we examined two possible models. One could ask the question of all possible models for a particular problem can we find the one with the highest likelihood ? If we have a dice with six faces that can only have the number 1, 2, and 3 then there is a finite amount of models and we can calculate their likelihoods as we did above. However if we relax the requirement to have a dice and simply have the values 1,2 and 3 but with arbitrary associated probabilities then we have an infinte number of possible models. Without going into the math it turns out that at least for this particular case the model that will have the maximum likelihood can be simply obtained by counting the relative frequencies of the values in the data. This is called maximum likelihood estimation of model parameters.
```
import collections
data = [1,2,2,1,1,3,1,2,3,2,2,2,2]
counts = collections.Counter(data)
print(counts)
est_probability_distribution = [counts[1]/float(len(data)), counts[2]/float(len(data)), counts[3]/float(len(data))]
print(est_probability_distribution)
values = np.int_([1, 2, 3])
probabilities = est_probability_distribution
model = Random_Variable('model', values, probabilities)
samples = model.sample(30)
print(samples)
```
# Maximum-likelihood parameter learning for Discrete Models
Lets start by creating a random variable corresponding to a bag of candy with two types lime and cherry similar to what we did in the previous notebook. We can easily generate random samples from this model. For example in the code below we geenerate 100 samples.
```
values = ['c', 'l']
probabilities = [0.2, 0.8]
dice1 = Random_Variable('bag1', values, probabilities)
samples = dice1.sample(100)
print(samples)
```
Now imagine that you are just given these samples and you are told that they were from a bag of candy but you don't know the percentage of each candy type in the bag and you need to estimate it. Let's call the probability a candy from the bag is cherry $\theta$. Then our task of parameter learning is to estimate $\theta$ from the provided samples. In the previous notebook without much explanation I stated that the "best" possible model in a maximum likelihood sense can be easily obtained by simply counting the percentage of each candy type in our bag.
As you can see the estimated paremeter $\theta$ is close but not the same as the original value which was $0.2$. If we had more samples this estimate becomes more accurate.
We can see that with this simple example we have the ability to "learn" a model. Once we have a "learned" model from the data we can use it to make predictions or inference in general as well as generate samples if needed.
```
import collections
counts = collections.Counter(samples)
print(counts)
est_probability_distribution = [counts['c']/float(len(samples)), counts['l']/float(len(samples))]
print(est_probability_distribution)
```
Using the counts seems intuitive and I told you that for the case of discrete random variables this provides the maximum likelihood estimate but can we prove this assertion?
Here is how we can do it. Each time we have a candy of a particular type we multiply the associated probability top get the likelihood of the sequence (assuming i.i.d. samples). If there are $c$ cherry candies and $l=N-c$ limes then we can write the likelihood as follows:
$$ P({\bf d} | h_{\theta}) = \prod_{j=1}^{N} P(d_j | h_{\theta}) = \theta^{c} * (1-\theta)^{l}$$
Note: check how the mathematical expression above is notated. It is using LaTeX notation which can be embedded in markdown cells. It is a useful thing to learn to produce nice looking equations in both notebook and papers.
The maximum-likelihood hypothesis is given by the value of $\theta$ that maximizes the exression above. The same value can be obtained by maximizing the **log likelihood**. Note that we have used log-likelihood before to avoid small numerical likelihood values when computing over long sequences. Here we use it because it allows us to simplify our expression to prove our approach to maximum likelihood parameter estimation. By taking the log we convert the product to a sum which is easer to maximize.
$$ L({\bf d}| h_{\theta}) = \log{P({\bf d} | h_{\theta})} = \sum_{j=1}^{N}\log{P(d_j| h_{\theta})} = c \log{ \theta} + l \log{(1-\theta)} $$
To find the maximum-likelihood value of $\theta$, we differentiate the $L$ with respect to $\theta$ and se the resulting expression to zero:
$$
\frac{L({\bf d} / h_{\theta})}{d \theta} = \frac{c}{\theta} - \frac{l}{1-\theta} = 0
$$
Solving for $\theta$ we get:
$$
\theta = \frac{c}{c+l} = \frac{c}{N}
$$
This might seem like a lot of work to prove something obvious but now we actually know that of all the infinite possible models of bags we could have - the one we estimate by counting the proportion of candy is the "best" in a maximum likelihood sense.
The approach we followed can be used for a variety of probabilistic models. The steps are as follows:
1. Write down an expression for the likelihood of the data as a function of the parameters and use log to simplify it for step 2
2. Write down the derivative of the log likelihood with respect to each parameter
3. Find the parameter values such that the derivatives are zero.
Note: If we are lucky we are able to perform steps 2 and 3 analytically and derive an exact ML parameter estimate. There are many cases especially when dealing with continuous models (which we cover below) in which
maximimizing the likelihood function analytically is not possible and one needs to resort to numerical methods
which do not provide an exact solution.
As another example of analytical ML parameter estimation, the book has one more example in which there is an extra random variable wrapper and the model has three parameters $\theta_1, \theta_2, \theta_3$.
For example by basicallly filtering the data and counting we can do ML parameter estimation for Naive Bayes models as well as Baysian Networks with discrete random variables. You have already seen to some extent how this can be done during lectures as well as in the assignments. Notice that the structure of the Bayesian network allows us to simplify the problem of ML parameter estimation by factoring different groups of variables based on their conditional structural relationships.
# Maximum-Likelihood parameter learning for continuous models
Continuous probability models are heavily used in real-world applications. Like I mentioned in the previous video - in many cases we need to resort to numerical optimization methods to perform parameter estimatino. However in some cases we can get the exact answer analytically. Let's consider the simple example of learning the parameters of a Gaussian density function on a single variable. Similarly to what we did in the previous video for learning the parameters of a disrete random variable we will first generate some data and then estimate the parameters from the data.
The data will be generated using a Gaussian density function on a single variable. The corresponding equation is:
$$ P(x) = \frac{1}{\sqrt{2 \pi \sigma}} e^{-\frac{x-\mu}{2\sigma^2}}$$
The parameters of this model are the mean $\mu$ and the standard deviation $\sigma$.
```
mu = 0.0
sigma = 0.2
s = np.random.normal(mu, sigma, 10)
print(s)
samples = np.random.normal(mu, sigma, 10000)
```
Let the observed values by $x_1, \dots, x_N$. Then the log-likelihood is:
$$
L = \sum_{j=1}^{N} \log{\frac{1}{\sqrt{2 \pi \sigma}}} e^{-\frac{x-\mu}{2\sigma^2}} = N(-\log \sqrt{2\pi} - \log \sigma) - \sum_{j=1}^{N} \frac{x-\mu}{2\sigma^2}
$$
Setting the derivatives to zero we obtain:
$$
\frac{\partial L}{\partial \mu} = - \frac{1}{\sigma^2}\sum_{j=1}^{N} (x_j-\mu) = 0
$$
which implies:
$$
\mu = \frac{\sum_j x_k}{N}
$$
So the maximum likelihood value of the mean is the sample average. Similarly you can find that the maximum likelihood value of the standard deviation is the square root of the sample variance. You can check the textbook for the details of the standard deviation $\sigma$.
Let's check how we can calculate these ML parameter estimates for the data that we have.
```
estimated_mean1 = np.sum(samples) / len(samples)
print(estimated_mean1)
estimated_mean2 = np.mean(samples)
print(estimated_mean2)
estimated_std = np.std(samples)
print(estimated_std)
```
So armed with simple filtering, counting and calculating sample mean and sample standard deviation we have everything we need to estimate the probabilities of a Naive Bayes model that contains a mixture of continuous and discrete variables. For the discrete variables we count and estimate directly the probabilities. For the continuous variables, we first estimate the ML parameters (sample mean and standard deviation) and then for a particular value of the feature we use the single varaible Gaussian density equation to derive a probability value for that value.
Some notes for further reading - not needed for the "final" assignment for those interested in digging deeper.
**Note1**: In a Baysian network with continuous variables you have the problem of having continuous parent and a continuous child variable. These can be addressed with linear Gaussian models. More details in the textbook
**Note2**: Similarly to Baysian learning in discrete models one can follow a similar approach and use a hypothesis prior to guide the learning. The textbook shows an example that use **beta distributions** you can check out.
**Note3**: If you remember when we covered Baysian network we looked at approximate inference using direct sampling and rejection sampling. You will notice that the approach we followed was similar to statistical learning in the sense that we generated samples and then used counting to estimate probabilities. So at a basic level inference and learning can be considered the same process. We start with a few things that we know and then using data we update what we know.
**Note4**: It is also possible to learn the structure of a Bayesian network from data. The basic idea is to search over the space of possible model. To do so we will need some method to determine when a good structure has been found. More details can be found in the book.
# Bayesian Learning
The two important terms we will cover are data and hypotheses or models. The hypotheses are different probabilistic theories. Let's consider the example described by the book.
We have a candy manufacturer that produces bags of candy wrapped in the same opaque wrapper. The flavors are cherry and lime. There are 5 kinds of bags:
h1: 100% cherry
h2: 75% cherry and 25% lime
h3: 50% cherry and 50% lime
h4: 25% cherry and 75% lime
h5: 100% lime
Given a new bag of candy the random variable *H* takes one of these 5 values: h1, h2, h3, h4, h5. We don't know which type it is and we gradually unwrap candy D1, D2, D3,..., DN where each of those is a random varilable with value $cherry$ and $lime$ (the bags are really big so replacement does not make a difference).
The task is given a sequence of observations D1, ... DN to predict the flavor of the next piece of candy.
As an extreme example consider we observe a sequence of 100 lime candies then we have a high confidence the bag is of type h1 and therefore the next candy will be lime.
One approach to solving this type of problem is the maximum likelihood described above. This consists of selecting the "best" hypothesis using the maximum likelihood and then doing the prediction using that hypothesis. If each candy bag is equally likely then this works.
```
data_book = ['l'] * 10
data_other = ['l','c','l','l','l','l','l','c','l','c']
counts_book = collections.Counter(data_book)
print(counts_book['c'], counts_book['l'])
counts_other = collections.Counter(data_other)
print(counts_other['c'], counts_other['l'])
h1 = [1.0, 0.0]
h2 = [0.75, 0.25]
h3 = [0.5, 0.5]
h4 = [0.25, 0.75]
h5 = [0.0, 1.0]
prior = [0.1, 0.2, 0.4, 0.2, 0.1]
prior = [0.2, 0.2, 0.2, 0.2, 0.2]
def likelihood(d, h):
counts = collections.Counter(d)
return np.power(h[0], counts['c']) * np.power(h[1], counts['l'])
print('Likelihood of h1 for data_book:', likelihood(data_book, h1))
print('Likelihood of h5 for data_book:', likelihood(data_book, h5))
print('Likelihood of h3 for data_book:', likelihood(data_book, h3))
print('Likelihood of h3 for data_other:', likelihood(data_other, h4))
```
Now consider that we have a prior probability distribution for the hypotheses. For the candy bag scenario
let's say we know from the manufacturer that 10% of candy bags are h1, 20% are h2, 40% h3, 20% h4 and 10% h5.
An alternative more general approach, called $Baysian Learning$ is to calculate the probability of each hypothesis given the data and then use all the hypotheses, weighted by their probabilitities rather than
just selecting the "best" by maximum likelihood to perform the prediction. Notice that with this approach we can take into account the prior probability over the hypotheses.
Mathematically we can calculate the probability of each hypothesis by weighting the likilehood by the prior:
$${\bf P}(h_{i} | {\bf d} ) = \alpha {\bf P}({\bf d} | h_{i}){\bf P}(h_i) $$
Suppose we want to make a prediction about an unknown quantity X such as predicting what the next candy will be. Then we have:
$$ {\bf P}(X | {\bf d}) = \sum_{i} {\bf P}(X| {\bf d}, h_i){\bf P}(h_i|{\bf d}) = \sum_{i} {\bf P}(X|h_i) {\bf P}(h_i | {\bf d})$$
where we assume that predictions are weighted averages over the predictions of the individual hypotheses. The key terms are the **hypothesis** prior $P(h_i)$ and the **likelihood** of the data under each hypothesis $P({\bf d} | h_i)$.
Assuming that the observatois are i.i.d we can calculate easily the likelihood like we did before and the priors for the hypotheses are given:
$$ P({\bf d} | h_i) = \prod_{j} P(d_k, h_i) $$
```
data_book = ['l','l','l','l','c','c','c','c','c','c']
import numpy as np
posterior = np.zeros(5)
posteriors = np.zeros((11, 5))
posteriors[0] = prior
for n in range(1,11):
for (i,h) in enumerate([h1,h2,h3,h4,h5]):
posterior[i] = (prior[i] * likelihood(data_book[:n], h))
posterior /= np.sum(posterior)
posteriors[n] = posterior
x = np.arange(0,11)
y1 = posteriors[:,0]
y2 = posteriors[:,1]
y3 = posteriors[:,2]
y4 = posteriors[:,3]
y5 = posteriors[:,4]
print(x)
print(y3)
```
Let's try to create a figure simular to 20.1 from the textbook that plots the posterior probabilities as a function of the number of observatons for each hypothesis. Each colored line corresponds to one of the five hypotheses (bag types of candy).
```
from bokeh.plotting import figure, show
from bokeh.io import output_notebook
output_notebook()
p = figure()
p.legend.title = 'Bayesian Learning'
for (y,label,color) in zip([y1,y2,y3,y4,y5],
['P(h1|d)', 'P(h2|d)','P(h3|d)', 'P(h4|d)', 'P(h5/d)'],
['red','green','blue','yellow', 'orange']):
p.line(x, y, legend=label, line_width=2,color=color)
p.circle(x,y, legend=label, color = color)
show(p)
```
The plot above shows how the posterior probabilities of each hypothesis get updated as we receive more data. To make a prediction
we consider all hypotheses and weight them by their posterior probabilities.
Let's first consider how to do a prediction without observing any data. Let's say we want the probability that the next candy is lime.
If we only had one hypothesis we can directly use the corresponding probability. For example if we have an h3 bag then the probability the next candy is lime is 0.5 - if we have an h5 bag the probability the next candy is like is 1.0.
Now if we have multiple hypothesis and their prior we simply weigh them by the prior. So for $h1, h2, h3, h4, h5$ and the prior
$P(h1) = 0.1, P(h2 = 0.2) P(h3 = 0.4) P(h4 = 0.2) P(h1 = 0.1)$
we would have:
$ 0.1 * 0 + 0.25 * 0.2 + 0.5 * 0.4 + 0.75 * 0.2 + 0.1 * 1 = 0.5$
If we know that d1 is lime then we use the posterior probabilities
after d1 that we calculated above to weigh the probabilities.
```
lime_probs = [h1[1], h2[1], h3[1], h4[1], h5[1]]
pd0 = posteriors[0,:]
print('Posteriors:',pd0)
print('Lime probabilities for each hypotheses:', lime_probs)
print('P(next=lime):', np.dot(pd0, lime_probs))
pd1 = posteriors[1,:]
print('Posteriors:',pd1)
print('Lime probabilities for each hypotheses:', lime_probs)
print('P(next=lime):', np.dot(pd1, lime_probs))
prob_next_lime_bayes = np.zeros(11)
for n in range(0,11):
for (i,h) in enumerate([h1,h2,h3,h4,h5]):
if (n == 0):
posterior[i] = prior[i]
else: # n > 0
posterior[i] = (prior[i] * likelihood(data_book[:n], h))
lime_probs = [h1[1], h2[1], h3[1], h4[1], h5[1]]
posterior /= np.sum(posterior)
prob_next_lime_bayes[n] = np.dot(posterior, lime_probs)
print(prob_next_lime_bayes)
```
We can plot the probability that the next candy is lime in a way similar to Figure 20.1 of the textbook.
```
p = figure()
p.line(x, prob_next_lime_bayes, line_width=2,color=color)
p.circle(x,prob_next_lime_bayes, color = color)
show(p)
```
Bayesian learning is very powerful but can be computationally expensive as we have to consider all possible hypotheses and this can become prohibitive in large problems. A common approximation is to make predictions based on the most probable hypothesis. This is called the maximum a posteriori or MAP hypothesis. Typically as we get more data the probability of competing hypothesis vanishes and therefore with enough data the MAP estimate tends to be the same as the Bayesian one.
So in practical terms rather than taking a sum we take a maximum. Let's modify the code for plotting the prediction to follow this approach.
```
prob_next_lime_map = np.zeros(11)
lime_probs = [h1[1], h2[1], h3[1], h4[1], h5[1]]
for n in range(0,11):
for (i,h) in enumerate([h1,h2,h3,h4,h5]):
if (n == 0):
posterior[i] = prior[i]
else: # n > 0
posterior[i] = (prior[i] * likelihood(data_book[:n], h))
posterior /= np.sum(posterior)
# this is the previous weighted sum
#prob_next_lime[n] = np.dot(posterior, lime_probs)
# instead we find the maximum posterior hypothesis and predict based on it
max_i = np.argmax(posterior)
prob_next_lime_map[n] = lime_probs[max_i]
print(prob_next_lime_map)
p = figure()
p.line(x, prob_next_lime_bayes, line_width=2,color='blue', legend='Bayes')
p.circle(x,prob_next_lime_bayes, color = 'blue', legend='Bayes')
p.line(x, prob_next_lime_map, line_width=2,color='red', legend='MAP')
p.circle(x,prob_next_lime_map, color = 'red', legend='MAP')
show(p)
```
Notice that in the case of uniform hypothesis prior then the MAP learning reduces to choosing the hypothesis that maximizes the likelihood. So to summarize we have three large families/types of statistical learning:
1. **Bayesian Learning** is the most powerful and flexible case in which all hypotheses are considered weighted by their probabilities
2. **Maximum a Posteriori (MAP)** learning is a common approximation that only considers the most probable hypothesis. It tends to be easier to compute that BL as it does not require a big summation or integration over the possible hypotheses.
3. **Maximum Likelihood (ML)** learning assumes that there is a uniform prior among the hypotheses. This is the simplest approach.
With enough data they all converge.
**Note1:** In machine learning each particular model/classifier can be considered a hypothesis. Overfitting can occur when the hypothesis space is very expressive and can capture a lot of variation in the data due to noise.
Bayesian and MAP learning methods use the prior to penalize complexity. That way one can control the tradeoff between the complexity of a hypothesis and its degree of fit to the data.
**Note2:** As I have mentioned before a lot of probability calculation are basically combinations of sums and products and the Bayesian learning computations above are no exception. In MAP learning instead of doing a weighted summation (dot-product) of the probabilities for different hypothesis we select the max. A similar differentiation can be observed in Hidden Markov Models when considering the difference between maximum likelihood estimation and filtering. In a filtering operation we are interested to know the current hidden state
given a sequence of observations. In maximum likelihood state estimation we are interested to know the entire sequence of hidden state that is most likely given a sequence of data. One question to ask is whether the result of filtering for a particular state would be always the same as the corresponding state in the maximum likelihood sequence of states. The answer is no.
To understand this consider what is called a trellis diagram that shows the transitions between states in an HMM.
To make it concrete let's say that the states are sunny and cloudy and we are interested in whether the state at step 3 is S1 (sunny). There are multiple paths through the trellis that would result in that outcome. For example
1. sunny, sunny, sunny
2. sunny, cloudy, sunny
3. could, sunny, sunny
4. cloudy, cloudy, sunny
For filtering we would calculate the probabilities of each of these paths by taking into account both the transition and observation model and then we would sum them all up. So in this case all paths are considered.
If you view each path as a hypotheses then this can be viewed as a type of Bayesian Learning.
In contrast in maximum likelihood state estimation we compute the probability of each path as before but now at
the end we select the path that is most probable. That gives us single path and associated probability so the summation we used in filtering becomes a max operator. This corresponds to MAP learning.
<img src="images/trellis.png" width="50%"/>
# Learning from incomplete data
There are many scenarios in learning probabilistic models in which there are hidden variables. This means that we have data for the evidence variables and we are interested in doing inference for some query variables but we do **NOT** have data for the hidden variables.
There are many scenarios where this is the case. For example the book describes:
1. Unsupervised clustering with mixtures of Gaussians
2. Learning Bayesian networks with hidden variables
3. Learning HMMs from observations without associated states
In all these cases, a common approach is to use the Expectation-Maximization (EM) algorithm.
This is an iterative algorithm in which at each step we improve our estimation of the parameters of a probabilistic model. The basic idea is simple to describe. Our probabilistic model is characterized by several parameters that we will denote as $\theta$. For example for a discrete Naive Bayes classifier this would be the probabilities of each feature given the class, for a continuous Naive Bayes classifier, this would be the mean and variance of each feature given the class, for a Bayesian network these would be the conditional probability tables, for a HMM these would be the transition model and sensor model probabilities.
There are two steps:
1. In the **expectation step (E-step)** we use the current iteration model characterized by $\theta_i$ to compute the expected values of the hidden variables for each example. In other words we use our current probability model to infer reasonable choices for the hidden variables for each example.
2. In the **maximization step (M-step)** we have now "complete" data in the sense that have the data values for the non hidden variables that we had from the beginning as well as expected data we obtained by using our current guess of the parameters of the model. In this step we can perform maximum-likelihood parameter estimation using complete data and obtained an updated model characterized by a new $\theta_{i+}$.
These two steps are repeated until convergence (i.e the parameters between iterations don't change - or more accurately change by a very small amount).
Another way to think of it is that if we have a probabilistic model we can sample it to generate data as well as use it to fill in missing values in the data that we have. After filling in these missing values we can learn a new model and repeat the process. In several notebooks we have seen this approach where we use a model to generate some data and then use that data to estimate a model (typicaly the estimated model is close to the original model). The EM-algorithm essentially alternates between these two steps.
Once you understand this basic principle, all sorts of variations and improvements can be used. For example the E-step can be done with approximate rather than exact inference or the M-step can be done with non-parametric estimation. The notations for specific problems can become relatively intimidating at first glance but if you really understand the basic principle you should be able to understand them for specific cases.
## Simple binary classification example
In this notebook I will show a very simple example of this idea. Hopefully this will give you some general intuition about this approach. Then you can review the specific book examples that are more complicated (learning Gaussian mixtures, Bayesian networks with hidden variables, and learning hmm parameters). We will end by showing the general mathematical notation.
Let's consider a simple binary classification problem with one continous attribute. For example this could be classifying whether some one is a professional basketball player or not based on their height. We can generate some synthetic data for this problem by simply sampling two Gaussian distribution. Let's say that professional basketball players have an average height of 190cm and the average height of other people is 175cm. For simiplicity we will consider they both have a standard deviation of 10cm.
```
import numpy as np
# generate twenty samples of each class
bball_samples = np.random.normal(190, 10, 20)
other_samples = np.random.normal(175, 10, 20)
print(bball_samples)
print(other_samples)
# generate 1000 samples of each class and plot histogram
bball_mean_height = 190
other_mean_height = 175
bball_samples = np.random.normal(bball_mean_height, 10, 1000)
other_samples = np.random.normal(other_mean_height, 10, 1000)
from matplotlib import pyplot
bins = np.linspace(150, 220, 100)
pyplot.hist(bball_samples, bins, alpha=0.5, label='x')
pyplot.hist(other_samples, bins, alpha=0.5, label='y')
pyplot.legend(loc='upper right')
pyplot.show()
```
You can clearly see in the histogram the height-distribution and overlap. You can also see that there is an equal number of instances for each class and that the standard deviation is the same.
Now suppose that you are just given the nba_samples and other_samples and told that these are labeled samples for training a Naive Bayes classifier. You also know that they both have a standard deviation of 10cm so we will keep that. In this case the only parameter we are trying to estimate is the mean of each class. So 𝜃=(𝜇𝑛𝑏𝑎,𝜇𝑜𝑡ℎ𝑒𝑟).
Given this data the maximum-likelihood estimate for the means is easily obtained by taking the statistical mean of the samples.
```
mu_bball = np.mean(bball_samples)
mu_other = np.mean(other_samples)
print(mu_bball, mu_other)
```
Now that we have "learned" a model we can use it to predict. Suppose you are given a test height - lets say 183cm. You can calcuate the $P(183/nba)$ and $P(183/other)$ by using the corresponding probability density functions characterized by $\mu_{nba}$ and $\sigma = 10$ and $\mu_{other}$ and $\sigma = 10$.
```
from scipy.stats import norm
test_height = 183
p_bball = norm(mu_bball, 10).pdf(test_height)
p_other = norm(mu_other, 10).pdf(test_height)
print(p_bball, p_other)
if (p_bball > p_other):
print(str(test_height) + " is more likely a professional basketball player")
else:
print(str(test_height) + " is more likely NOT a professional basketball player")
```
## Unsupervised learning
Now let's make a very simple change to the problem above. Suppose that we know that we have two classes and we have heights. We also know that the standard deviation for each class is 10cm. However we are not given the labels of the heights but just a dataset of heights.
```
heights = np.hstack([bball_samples,other_samples])
np.random.shuffle(heights)
print(heights)
```
If we know the means of the two classes, we can predict the class of each instance (height)
```
estimated_bball_samples = []
estimated_other_samples = []
for h in heights[:20]:
p_bball = norm(mu_bball, 10).pdf(h)
p_other = norm(mu_other, 10).pdf(h)
if (p_bball > p_other):
print('bball', h)
else:
print('other', h)
```
So the idea of the EM-algorithm would be let's start with a reasonable guess of the two means, then predict the heights, the re-estimate the means and keep repeating until convergence.
```
mu_bball = 170
mu_other = 165
for i in range(0,12):
estimated_bball_samples = []
estimated_other_samples = []
print((i,mu_bball, mu_other))
for h in heights:
# E-step - use current model to estimate values for the hidden variable (class membership)
p_bball = norm(mu_bball, 10).pdf(h)
p_other = norm(mu_other, 10).pdf(h)
if (p_bball > p_other):
estimated_bball_samples.append(h)
else:
estimated_other_samples.append(h)
# M-step - using the estimated class values re-caculate the parameters of the model i.e the means
mu_bball = np.mean(estimated_bball_samples)
mu_other = np.mean(estimated_other_samples)
```
This is a simple example of the EM-algorithm with only one parameter to estimate per class to show the basic principle. In more real-world scenarios the probabilistic model can be much more complex with many parameters.
In addition the prediction step typically is done using likelihoods and weighted samples rather than simple
prediction as in this example. Therefore the E-step and M-step tend to be more complicated and have complex update equations but the basic principle is the same.
**Advanced**: A great tutorial article on EM is:
Bilmes JA. A gentle tutorial of the EM algorithm and its application to parameter estimation for Gaussian mixture and hidden Markov models. International Computer Science Institute. 1998 Apr 21;4(510):126.
http://www.leap.ee.iisc.ac.in/sriram/teaching/MLSP_18/refs/GMM_Bilmes.pdf
| github_jupyter |
```
# Copyright 2021 NVIDIA Corporation. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
```
# Getting Started Outbrain: Download and Convert
## Overview
Outbrain dataset was published in [Kaggle Outbrain click prediction](https://www.kaggle.com/c/outbrain-click-prediction) competition, where the ‘Kagglers’ were challenged to predict on which ads and other forms of sponsored content its global users would click. One of the top finishers' preprocessing and feature engineering pipeline is taken into consideration here, and this pipeline was restructured using NVTabular and cuDF.
```
import os
# Get dataframe library - cudf or pandas
from nvtabular.dispatch import get_lib, random_uniform, reinitialize
df_lib = get_lib()
```
## Download the dataset
First, you need to [download](https://www.kaggle.com/c/outbrain-click-prediction/data) the Kaggle Outbrain click prediction challenge and set DATA_BUCKET_FOLDER with the dataset path.
```
DATA_BUCKET_FOLDER = os.environ.get("INPUT_DATA_DIR", "~/nvt-examples/outbrain/data/")
```
The OUTPUT_BUCKET_FOLDER is the folder where the preprocessed dataset will be saved.
```
OUTPUT_BUCKET_FOLDER = os.environ.get("OUTPUT_DATA_DIR", "./outbrain-preprocessed/")
os.makedirs(OUTPUT_BUCKET_FOLDER, exist_ok=True)
```
## Preparing Our Dataset
Here, we merge the component tables of our dataset into a single data frame, using [cuDF](https://github.com/rapidsai/cudf), which is a GPU DataFrame library for loading, joining, aggregating, filtering, and otherwise manipulating data. We do this because NVTabular applies a workflow to a single table. We also re-initialize managed memory. `rmm.reinitialize()` provides an easy way to initialize RMM (RAPIDS Memory Manager) with specific memory resource options across multiple devices. The reason we re-initialize managed memory here is to allow us to perform memory intensive merge operation. Note that dask-cudf can also be used here.
```
# use managed memory for device memory allocation
reinitialize(managed_memory=True)
# Alias for read_csv
read_csv = df_lib.read_csv
# Merge all the CSV files together
documents_meta = read_csv(DATA_BUCKET_FOLDER + "documents_meta.csv", na_values=["\\N", ""])
merged = (
read_csv(DATA_BUCKET_FOLDER + "clicks_train.csv", na_values=["\\N", ""])
.merge(
read_csv(DATA_BUCKET_FOLDER + "events.csv", na_values=["\\N", ""]),
on="display_id",
how="left",
suffixes=("", "_event"),
)
.merge(
read_csv(DATA_BUCKET_FOLDER + "promoted_content.csv", na_values=["\\N", ""]),
on="ad_id",
how="left",
suffixes=("", "_promo"),
)
.merge(documents_meta, on="document_id", how="left")
.merge(
documents_meta,
left_on="document_id_promo",
right_on="document_id",
how="left",
suffixes=("", "_promo"),
)
)
```
## Splitting into train and validation datasets
We use a time-stratified sample to create a validation set that is more recent, and save both our train and validation sets to parquet files to be read by NVTabular. Note that you should run the cell below only once, then save your `train` and `valid` data frames as parquet files. If you want to rerun this notebook you might end up with a different train-validation split each time because samples are drawn from a uniform distribution.
```
# Do a stratified split of the merged dataset into a training/validation dataset
merged["day_event"] = (merged["timestamp"] / 1000 / 60 / 60 / 24).astype(int)
random_state = df_lib.Series(random_uniform(size=len(merged)))
valid_set, train_set = merged.scatter_by_map(
((merged.day_event <= 10) & (random_state > 0.2)).astype(int)
)
train_set.head()
```
We save the dataset to disk.
```
train_filename = os.path.join(OUTPUT_BUCKET_FOLDER, "train_gdf.parquet")
valid_filename = os.path.join(OUTPUT_BUCKET_FOLDER, "valid_gdf.parquet")
train_set.to_parquet(train_filename, compression=None)
valid_set.to_parquet(valid_filename, compression=None)
merged = train_set = valid_set = None
reinitialize(managed_memory=False)
```
| github_jupyter |
# Deep learning the collisional cross sections of the peptide universe from a million experimental values
Florian Meier, Niklas D. Köhler, Andreas-David Brunner, Jean-Marc H. Wanka, Eugenia Voytik, Maximilian T. Strauss, Fabian J. Theis, Matthias Mann
Pre-print: https://doi.org/10.1101/2020.05.19.102285
Publication: pending
revised 09/2020
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import glob
from scipy import optimize
```
#### Import raw data from MaxQuant output
```
# Load evidence.txt files from folder
filenames = glob.glob("data/evidence*.txt")
evidences = [pd.read_csv(filename, sep='\t', engine='python', header=0) for filename in filenames]
# Combine all evidences in one dataframe
evidence_all = pd.concat(evidences, sort=False, ignore_index = True)
# Clean up
del evidences
evidence_all.head()
# Drop reverse hits
# Drop features with no intensity value
evidence_all = evidence_all.loc[(evidence_all['Reverse'] != '+') & (evidence_all['Intensity'] > 0)]
'{0} CCS values in the entire data set.'.format(len(evidence_all))
# Number of CCS values per Experiment
evidence_all['Experiment'].value_counts(), evidence_all['Experiment'].value_counts().sum()
```
### Construction of a very large scale peptide CCS data set
```
# Analysis of whole-proteome digests
group = ['CElegans_Tryp',
'Drosophila_LysC',
'Drosophila_LysN',
'Drosophila_Trp',
'Ecoli_LysC',
'Ecoli_LysN',
'Ecoli_trypsin',
'HeLa_LysC',
'HeLa_LysN',
'HeLa_Trp_2',
'HeLa_Trypsin_1',
'Yeast_LysC',
'Yeast_LysN',
'Yeast_Trypsin']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
len(evidence_tmp)
```
<b>Figure 1b.<b/> Overview of the CCS data set in this study by organism.
```
print('Number of LC-MS/MS runs: {0}'.format(len(set(evidence_tmp['Raw file']))))
print('Number of peptide spectrum matches: {0}'.format(len(evidence_tmp['Raw file'])))
print('Number of unique CCS values (mod. sequence/charge): {0}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str))),
len(set(evidence_tmp['Sequence']))))
print('Number of unique peptide sequences: {0}'.format(len(set(evidence_tmp['Sequence']))))
group = ['CElegans_Tryp']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of PSMs/CCS values (C.elegans): {}'.format(len(evidence_tmp['Sequence'])))
print('Number of unique CCS values (C.elegans): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
group = ['Drosophila_LysC', 'Drosophila_LysN', 'Drosophila_Trp']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of PSMs/CCS values (Drosophila): {}'.format(len(evidence_tmp['Sequence'])))
print('Number of unique CCS values (Drosophila): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
group = ['Ecoli_LysC', 'Ecoli_LysN', 'Ecoli_trypsin']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of PSMs/CCS values (E.coli): {}'.format(len(evidence_tmp['Sequence'])))
print('Number of unique CCS values (E.coli): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
group = ['HeLa_LysC', 'HeLa_LysN', 'HeLa_Trp_2', 'HeLa_Trypsin_1']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of PSMs/CCS values (HeLa): {}'.format(len(evidence_tmp['Sequence'])))
print('Number of unique CCS values (HeLa): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
group = ['Yeast_LysC', 'Yeast_LysN', 'Yeast_Trypsin']
evidence_tmp= evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of PSMs/CCS values (Yeast): {}'.format(len(evidence_tmp['Sequence'])))
print('Number of unique CCS values (Yeast): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
group = ['CElegans_Tryp',
'Drosophila_LysC',
'Drosophila_Trp',
'Ecoli_LysC',
'Ecoli_trypsin',
'HeLa_LysC',
'HeLa_Trp_2',
'HeLa_Trypsin_1',
'Yeast_LysC',
'Yeast_Trypsin']
evidence_tmp= evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of unique CCS values (Trypsin and LysC): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
print('Number of unique sequences (Trypsin and LysC): {}'.format(len(set(evidence_tmp['Sequence']))))
# Unique peptide sequences contributed by trypsin and LysC data set
np.round(338192/426845*100,0)
group = ['Drosophila_LysN',
'Ecoli_LysN',
'HeLa_LysN',
'Yeast_LysN']
evidence_tmp= evidence_all.loc[evidence_all['Experiment'].isin(group)]
print('Number of unique CCS values (LysN): {}'.format(
len(set(evidence_tmp['Modified sequence'].astype(str) + evidence_tmp['Charge'].astype(str)))))
print('Number of unique sequences (LysN): {}'.format(len(set(evidence_tmp['Sequence']))))
# Unique peptide sequences contributed by LysN data set
np.round(89093/426845*100,0)
# Occurence of multiple features per modified sequence/charge state in single LC-MS runs.
group = ['CElegans_Tryp',
'Drosophila_LysC',
'Drosophila_LysN',
'Drosophila_Trp',
'Ecoli_LysC',
'Ecoli_LysN',
'Ecoli_trypsin',
'HeLa_LysC',
'HeLa_LysN',
'HeLa_Trp_2',
'HeLa_Trypsin_1',
'Yeast_LysC',
'Yeast_LysN',
'Yeast_Trypsin']
evidence_tmp = evidence_all.loc[evidence_all['Experiment'].isin(group)]
hist_data = evidence_tmp.groupby(['Modified sequence', 'Charge', 'Raw file'], ).count()
len(hist_data), len(evidence_tmp)
fig = plt.figure(figsize=(6,6))
ax = plt.axes()
plt.hist(x = hist_data['Sequence'], bins = 5, range = (1,6), align = 'left',
weights= (np.zeros_like(hist_data['Sequence']) + 100) / hist_data['Sequence'].size,
rwidth = 0.5)
plt.xlabel('No. of features')
plt.ylabel('% of total')
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.ylim((0,100))
plt.xticks(range(1,6))
plt.yticks(range(0,101,10))
plt.savefig("figures/Suppl_Fig_1.jpg");
plt.show()
```
<b>Supplementary Figure 1.</b> Number of detected features per modified peptide sequence and charge
state in single LC-TIMS-MS experiments.
```
# Reduce to unique modified sequence/charge state
evidence_unique = evidence_tmp.loc[evidence_tmp.groupby(['Modified sequence', 'Charge'])['Intensity'].idxmax()]
len(evidence_unique)
# Reduce to unique sequence
evidence_unique_seq = evidence_tmp.loc[evidence_tmp.groupby(['Sequence'])['Intensity'].idxmax()]
len(evidence_unique_seq)
# Peptide length distribution
print("Median peptide length: {:10.1f} amino acids".format(evidence_unique_seq['Length'].median()))
print("Minimum peptide length: {:10.1f} amino acids".format(evidence_unique_seq['Length'].min()))
print("Maximum peptide length: {:10.1f} amino acids".format(evidence_unique_seq['Length'].max()))
evidence_unique['Length'].hist()
plt.xlabel('Amino acid count')
plt.ylabel('Count');
cmap = plt.get_cmap("RdYlBu")
colors = cmap(np.linspace(0, 1, num=20))
# C-terminal amino acids
fig, ax = plt.subplots()
size = 0.4
vals = np.array(evidence_unique_seq['Sequence'].str[-1:].value_counts())
labels = evidence_unique_seq['Sequence'].str[-1:].value_counts().index.tolist()
ax.pie(vals, radius=1, labels = labels, colors=colors,
wedgeprops=dict(width=size, edgecolor='w'))
ax.set(aspect="equal")
plt.savefig("figures/Figure_1_c.pdf")
plt.show()
```
<b>Figure 1c.</b> Frequency of peptide C-terminal amino acids.
```
np.sum(vals), vals, labels
# N-terminal amino acids
fig, ax = plt.subplots()
size = 0.4
vals = np.array(evidence_unique_seq['Sequence'].str[:1].value_counts())
labels = evidence_unique_seq['Sequence'].str[:1].value_counts().index.tolist()
ax.pie(vals, radius=1, colors=colors, labels = labels,
wedgeprops=dict(width=size, edgecolor='w'))
ax.set(aspect="equal")
plt.savefig("figures/Figure_1_d.pdf")
plt.show()
```
<b>Figure 1d.</b> Frequency of peptide N-terminal amino acids.
```
np.sum(vals), vals, labels
# Evaluate peptide charge distribution
evidence_unique['Charge'].value_counts() / np.sum(evidence_unique['Charge'].value_counts()) * 100
# Drop charge 1 features
evidence_unique = evidence_unique.loc[(evidence_unique['Charge'] != 1)]
len(evidence_unique)
# Charge distribution
charge_color = [colors[0], colors[6], colors[18]]
fig, ax = plt.subplots()
size = 0.6
vals = np.array(evidence_unique['Charge'].value_counts())
labels = evidence_unique['Charge'].value_counts().index.tolist()
ax.pie(vals, radius=1, labels = labels, colors=charge_color,
wedgeprops=dict(width=size, edgecolor='w'))
ax.set(aspect="equal")
plt.savefig("figures/Figure_1_e_Charge.pdf")
plt.show()
# m/z vs. CCS distribution
charge_color = {2: colors[0], 3: colors[6], 4: colors[18]}
grid = sns.JointGrid(x='m/z', y='CCS', data=evidence_unique)
g = grid.plot_joint(plt.scatter, c=evidence_unique['Charge'].apply(lambda x: charge_color[x]), alpha=0.1, s=0.1)
g.fig.set_figwidth(8)
g.fig.set_figheight(6)
sns.kdeplot(data=evidence_unique, x=evidence_unique.loc[evidence_unique['Charge']== 2, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color=colors[0])
sns.kdeplot(data=evidence_unique, x=evidence_unique.loc[evidence_unique['Charge']== 3, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color=colors[6])
sns.kdeplot(data=evidence_unique, x=evidence_unique.loc[evidence_unique['Charge']== 4, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color=colors[18])
sns.kdeplot(data=evidence_unique, y=evidence_unique.loc[evidence_unique['Charge']== 2, 'CCS'], ax=g.ax_marg_y,
legend=False, fill=True, color=colors[0])
sns.kdeplot(data=evidence_unique, y=evidence_unique.loc[evidence_unique['Charge']== 3, 'CCS'], ax=g.ax_marg_y,
legend=False, fill=True, color=colors[6])
sns.kdeplot(data=evidence_unique, y=evidence_unique.loc[evidence_unique['Charge']== 4, 'CCS'], ax=g.ax_marg_y,
legend=False, fill=True, color=colors[18])
g.savefig("figures/Figure_1_e.jpg")
len(evidence_unique.loc[evidence_unique['Charge']== 2]), \
len(evidence_unique.loc[evidence_unique['Charge']== 3]), \
len(evidence_unique.loc[evidence_unique['Charge']== 4])
len(evidence_unique.loc[evidence_unique['Charge']== 2]) + \
len(evidence_unique.loc[evidence_unique['Charge']== 3]) + \
len(evidence_unique.loc[evidence_unique['Charge']== 4])
```
### Analysis of ion mobility trend lines
```
def trendline_func(x, a, b):
return a * np.power(x, b)
# Subset tryptic peptides (C-terminal R or K)
evidence_tryptic = evidence_unique.loc[
(evidence_unique['Sequence'].str[-1:] == 'R') | (evidence_unique['Sequence'].str[-1:] == 'K')]
len(evidence_tryptic)
# Split data set by charge state
CCS_fit_charge2 = evidence_tryptic[evidence_tryptic['Charge'] == 2]
CCS_fit_charge3 = evidence_tryptic[evidence_tryptic['Charge'] == 3]
CCS_fit_charge4 = evidence_tryptic[evidence_tryptic['Charge'] == 4]
# Fit to power-law trend line
params_charge2, params_covariance_charge2 = optimize.curve_fit(trendline_func, CCS_fit_charge2['m/z'], CCS_fit_charge2['CCS'])
params_charge3, params_covariance_charge3 = optimize.curve_fit(trendline_func, CCS_fit_charge3['m/z'], CCS_fit_charge3['CCS'])
params_charge4, params_covariance_charge4 = optimize.curve_fit(trendline_func, CCS_fit_charge4['m/z'], CCS_fit_charge4['CCS'])
print('2+')
print(params_charge2, params_covariance_charge2)
print('---')
print('3+')
print(params_charge3, params_covariance_charge3)
print('---')
print('4+')
print(params_charge4, params_covariance_charge4)
# m/z vs. CCS distribution
charge_color = {2: colors[0], 3: colors[6], 4: colors[18]}
grid = sns.JointGrid(x='m/z', y='CCS', data = evidence_tryptic)
g = grid.plot_joint(plt.scatter, c = evidence_tryptic['Charge'].apply(lambda x: charge_color[x]), alpha=0.1, s=0.1)
plt.plot(np.arange(300,1800,1), trendline_func(np.arange(300,1800,1), params_charge2[0], params_charge2[1]),
color = "black", ls = 'dashed', lw = 1)
plt.plot(np.arange(300,1800,1), trendline_func(np.arange(300,1800,1), params_charge3[0], params_charge3[1]),
color = "black", ls = 'dashed', lw = 1)
plt.plot(np.arange(300,1800,1), trendline_func(np.arange(300,1800,1), params_charge4[0], params_charge4[1]),
color = "black", ls = 'dashed', lw = 1)
g.fig.set_figwidth(8)
g.fig.set_figheight(6)
g.savefig("figures/Suppl_Fig_2a.jpg")
```
<b>Supplementary Figure 2a.</b> Distribution of tryptic peptides in the m/z vs. CCS space color-coded
by charge state as in Figure 1. Fitted power-law (A*x^b)) trend lines (dashed lines) visualize the
correlation of ion mass and mobility in each charge state.
```
CCS_fit_charge2['Delta fit'] = (CCS_fit_charge2['CCS'] - trendline_func(CCS_fit_charge2['m/z'], params_charge2[0],
params_charge2[1])) / CCS_fit_charge2['CCS'] * 100
plt.figure(figsize=(4,2))
ax = CCS_fit_charge2['Delta fit'].hist(bins = 100, range=(-20,20), color = colors[0])
plt.grid(b = False)
plt.xticks(np.arange(-20, 21, step=5), (-20, '', -10, '', 0, '', 10, '', 20))
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.xlabel('Residual (%)')
plt.ylabel('Count');
plt.savefig("figures/Suppl_Fig_2b.jpg")
```
<b>Supplementary Figure 2b.</b> Residuals for charge state 2.
```
CCS_fit_charge3['Delta fit'] = (CCS_fit_charge3['CCS'] - trendline_func(CCS_fit_charge3['m/z'], params_charge3[0],
params_charge3[1])) / CCS_fit_charge3['CCS'] * 100
plt.figure(figsize=(4,2))
ax = CCS_fit_charge3['Delta fit'].hist(bins = 100, range=(-20,20), color = colors[6])
plt.grid(b = False)
plt.xticks(np.arange(-20, 21, step=5), (-20, '', -10, '', 0, '', 10, '', 20))
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.xlabel('Residual (%)')
plt.ylabel('Count');
plt.savefig("figures/Suppl_Fig_2c.jpg")
```
<b>Supplementary Figure 2c.</b> Residuals for charge state 3.
```
CCS_fit_charge4['Delta fit'] = (CCS_fit_charge4['CCS'] - trendline_func(CCS_fit_charge4['m/z'], params_charge4[0],
params_charge4[1])) / CCS_fit_charge4['CCS'] * 100
plt.figure(figsize=(4,2))
ax = CCS_fit_charge4['Delta fit'].hist(bins = 100, range=(-20,20), color = colors[18])
plt.grid(b = False)
plt.xticks(np.arange(-20, 21, step=5), (-20, '', -10, '', 0, '', 10, '', 20))
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
plt.xlabel('Residual (%)')
plt.ylabel('Count');
plt.savefig("figures/Suppl_Fig_2d.jpg")
```
<b>Supplementary Figure 2d.</b> Residuals for charge state 4.
```
# Percentile distribution around trend line
CCS_fit = CCS_fit_charge2['Delta fit'].append(CCS_fit_charge3['Delta fit']).append(CCS_fit_charge4['Delta fit'])
np.percentile(CCS_fit, 2.5), np.percentile(CCS_fit, 97.5)
```
### Analysis of peak capacity
```
def calc_K0_from_CCS(CCS, charge, mass):
mass = mass + charge * 1.00727647
k0 = CCS * np.sqrt(305 * mass * 28 / (28 + mass)) * 1/18500 * 1/charge
return k0
# Subset tryptic peptides (C-terminal R or K)
evidence_tryptic = evidence_unique.loc[
(evidence_unique['Sequence'].str[-1:] == 'R') | (evidence_unique['Sequence'].str[-1:] == 'K')]
len(evidence_tryptic)
evidence_tryptic['1/K0'] = calc_K0_from_CCS(evidence_tryptic['CCS'], evidence_tryptic['Charge'], evidence_tryptic['Mass'])
evidence_tryptic['1/K0'].hist();
charge_color = {2: colors[0], 3: colors[6], 4: colors[18]}
grid = sns.JointGrid(x='m/z', y='1/K0', data = evidence_tryptic)
g = grid.plot_joint(plt.scatter, c = evidence_tryptic['Charge'].apply(lambda x: charge_color[x]), alpha = 0.5, s = 0.5)
g.fig.set_figwidth(6)
g.fig.set_figheight(4)
sns.kdeplot(data=evidence_tryptic, x=evidence_tryptic.loc[evidence_tryptic['Charge']== 2, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color = colors[0])
sns.kdeplot(data=evidence_tryptic, x=evidence_tryptic.loc[evidence_tryptic['Charge']== 3, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color = colors[6])
sns.kdeplot(data=evidence_tryptic, x=evidence_tryptic.loc[evidence_tryptic['Charge']== 4, 'm/z'], ax=g.ax_marg_x,
legend=False, fill=True, color = colors[18])
sns.kdeplot(data=evidence_tryptic, y=evidence_tryptic.loc[evidence_tryptic['Charge']== 2, '1/K0'], ax=g.ax_marg_y,
legend=False, fill=True, color = colors[0])
sns.kdeplot(data=evidence_tryptic, y=evidence_tryptic.loc[evidence_tryptic['Charge']== 3, '1/K0'], ax=g.ax_marg_y,
legend=False, fill=True, color = colors[6])
sns.kdeplot(data=evidence_tryptic, y=evidence_tryptic.loc[evidence_tryptic['Charge']== 4, '1/K0'], ax=g.ax_marg_y,
legend=False, fill=True, color = colors[18]);
plt.xlim(300,1650)
plt.ylim(0.7,1.5)
plt.savefig("figures/Suppl_Fig_3a.jpg");
```
<b>Supplementary Figure 3a.</b> Distribution of tryptic peptides in the m/z vs. ion mobility (1/K0)
space color-coded by charge state as in Figure 1.
```
xbins = 1350
ybins = 44
hist, xedges, yedges = np.histogram2d(evidence_tryptic['m/z'],
evidence_tryptic['1/K0'],
bins=[xbins,ybins],
range=[[300, 1650], [0.7, 1.5]])
over_threshold = hist > 1
print('Fraction of peptides:')
print(np.round((np.nansum((10**np.log10(hist-1))) / hist.sum().sum() ), 2))
print('Occupancy of the 2D space:')
print(np.round(sum((sum(over_threshold))/(xbins*ybins)),2))
fig = plt.figure(figsize=(6,4))
plt.xlabel('m/z')
plt.ylabel('1/K0');
XB = xedges
YB = yedges
X,Y = np.meshgrid(xedges,yedges)
Z = over_threshold.T
plt.pcolormesh(X, Y, Z);
plt.savefig("figures/Suppl_Fig_3b.jpg")
```
<b>Supplementary Figure 3b.</b> Estimating the peak capacity (Φ) of two-
dimensional peptide separation with TIMS-MS. In an ideally orthogonal 2D separation, the total
peak capacity would be ΦMS \* ΦTIMS. Assuming an ion mobility resolution of 60 ((1/K0) / Δ(1/K0)),
the average peak full width at half maximum is 0.018 Vs cm-2 in the peptide 1/K0 range (0.7-1.5
Vs cm-2). This would result in a theoretical peak capacity of ΦMS \* 44. However, the correlation of
mass and mobility reduces the effective peak capacity and the 2D histogram analysis (1350 m/z x
44 ion mobility bins) shows that 96% of the peptides occupy about 27% of the total area (yellow
vs. purple area). Using this as a correction factor, we estimate the peak capacity of TIMS-MS to
about ΦMS \* 12.
| github_jupyter |
# Random Signals
*This jupyter notebook is part of a [collection of notebooks](../index.ipynb) on various topics of Digital Signal Processing.
## Cumulative Distribution Functions
A random process can be characterized by the statistical properties of its amplitude values. [Cumulative distribution functions](https://en.wikipedia.org/wiki/Cumulative_distribution_function) (CDFs) are one possibility to do so.
### Univariate Cumulative Distribution Function
The univariate CDF $P_x(\theta, k)$ of a continuous-amplitude real-valued random signal $x[k]$ is defined as
\begin{equation}
P_x(\theta, k) := \Pr \{ x[k] \leq \theta\}
\end{equation}
where $\Pr \{ \cdot \}$ denotes the probability that the given condition holds. The univariate CDF quantifies the probability that for the entire ensemble and for a fixed time index $k$ the amplitude $x[k]$ is smaller or equal to $\theta$. The term '*univariate*' reflects the fact that only one random process is considered.
The CDF shows the following properties which can be concluded directly from its definition
\begin{equation}
\lim_{\theta \to -\infty} P_x(\theta, k) = 0
\end{equation}
and
\begin{equation}
\lim_{\theta \to \infty} P_x(\theta, k) = 1
\end{equation}
The former property results from the fact that all amplitude values $x[k]$ are larger than $- \infty$, the latter from the fact that all amplitude values lie within $- \infty$ and $\infty$. The univariate CDF $P_x(\theta, k)$ is furthermore a non-decreasing function
\begin{equation}
P_x(\theta_1, k) \leq P_x(\theta_2, k) \quad \text{for } \theta_1 \leq \theta_2
\end{equation}
The probability that $\theta_1 < x[k] \leq \theta_2$ is given as
\begin{equation}
\Pr \{\theta_1 < x[k] \leq \theta_2\} = P_x(\theta_2, k) - P_x(\theta_1, k)
\end{equation}
Hence, the probability that a continuous-amplitude random signal takes a specific value $x[k]=\theta$ is zero when calculated by means of the CDF. This motivates the definition of probability density functions introduced later.
### Bivariate Cumulative Distribution Function
The statistical dependencies between two signals are frequently of interest in statistical signal processing. The bivariate or joint CDF $P_{xy}(\theta_x, \theta_y, k_x, k_y)$ of two continuous-amplitude real-valued random signals $x[k]$ and $y[k]$ is defined as
\begin{equation}
P_{xy}(\theta_x, \theta_y, k_x, k_y) := \Pr \{ x[k_x] \leq \theta_x \wedge y[k_y] \leq \theta_y \}
\end{equation}
The joint CDF quantifies the probability for the entire ensemble of sample functions that for a fixed $k_x$ the amplitude value $x[k_x]$ is smaller or equal to $\theta_x$ and that for a fixed $k_y$ the amplitude value $y[k_y]$ is smaller or equal to $\theta_y$. The term '*bivariate*' reflects the fact that two random processes are considered. The following properties can be concluded from its definition
\begin{align}
\lim_{\theta_x \to -\infty} P_{xy}(\theta_x, \theta_y, k_x, k_y) &= 0 \\
\lim_{\theta_y \to -\infty} P_{xy}(\theta_x, \theta_y, k_x, k_y) &= 0
\end{align}
and
\begin{equation}
\lim_{\substack{\theta_x \to \infty \\ \theta_y \to \infty}} P_{xy}(\theta_x, \theta_y, k_x, k_y) = 1
\end{equation}
The bivariate CDF can also be used to characterize the statistical properties of one random signal $x[k]$ at two different time-instants $k_x$ and $k_y$ by setting $y[k] = x[k]$
\begin{equation}
P_{xx}(\theta_1, \theta_2, k_1, k_2) := \Pr \{ x[k_1] \leq \theta_1 \wedge y[k_2] \leq \theta_2 \}
\end{equation}
The definition of the bivariate CDF can be extended straightforward to the case of more than two random variables. The resulting CDF is termed as multivariate CDF.
## Probability Density Functions
[Probability density functions](https://en.wikipedia.org/wiki/Probability_density_function) (PDFs) describe the probability for one or multiple random signals to take on a specific value. Again the univariate case is discussed first.
### Univariate Probability Density Function
The univariate PDF $p_x(\theta, k)$ of a continuous-amplitude real-valued random signal $x[k]$ is defined as the derivative of the univariate CDF
\begin{equation}
p_x(\theta, k) = \frac{\partial}{\partial \theta} P_x(\theta, k)
\end{equation}
This can be seen as the differential equivalent of the limit case $\lim_{\theta_2 \to \theta_1} \Pr \{\theta_1 < x[k] \leq \theta_2\}$. As a consequence of above definition, the CDF can be computed from the PDF by integration
\begin{equation}
P_x(\theta, k) = \int\limits_{-\infty}^{\theta} p_x(\alpha, k) \, \mathrm{d}\alpha
\end{equation}
Due to the properties of the CDF and its definition, the PDF shows the following properties
\begin{equation}
p_x(\theta, k) \geq 0
\end{equation}
and
\begin{equation}
\int\limits_{-\infty}^{\infty} p_x(\theta, k) \, \mathrm{d}\theta = \lim_{\theta \to \infty} P_x(\theta, k) = 1
\end{equation}
The univariate PDF has only positive values and the area below the PDF is equal to one. The latter property may be used for normalization.
#### Example - Estimate of an univariate PDF by the histogram
In the process of calculating a [histogram](https://en.wikipedia.org/wiki/Histogram), the entire range of amplitude values of a random signal is split into a series of intervals (bins). For a given random signal the number of samples is counted which fall into one of these intervals. This is repeated for all intervals. The counts are finally normalized with respect to the total number of samples. This process constitutes a numerical estimation of the PDF of a random process.
In the following example the histogram of an ensemble of random signals is computed for each time index $k$. The CDF is computed by taking the cumulative sum over the histogram bins. This constitutes a numerical approximation of above integral
\begin{equation}
\int\limits_{-\infty}^{\theta} p_x(\alpha, k) \, \mathrm{d}\alpha \approx \sum_{i=0}^{N} p_x(\theta_i, k) \, \Delta\theta_i
\end{equation}
where $p_x(\theta_i, k)$ denotes the $i$-th bin of the PDF and $\Delta\theta_i$ its width.
```
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
K = 32 # number of temporal samples
N = 10000 # number of sample functions
bins = 100 # number of bins for the histogram
# draw sample functions from a random process
np.random.seed(2)
x = np.random.normal(size=(N, K))
x += np.tile(np.cos(2*np.pi/K*np.arange(K)), [N, 1])
# compute the histogram
px = np.zeros((bins, K))
for k in range(K):
px[:, k], edges = np.histogram(x[:, k], bins=bins, range=(-4,4), density=True)
# compute the CDF
Px = np.cumsum(px, axis=0) * 8/bins
# plot the PDF
plt.figure(figsize=(10,6))
plt.pcolor(np.arange(K), edges, px)
plt.title(r'Estimated PDF $\hat{p}_x(\theta, k)$')
plt.xlabel(r'$k$')
plt.ylabel(r'$\theta$')
plt.colorbar()
plt.autoscale(tight=True)
# plot the CDF
plt.figure(figsize=(10,6))
plt.pcolor(np.arange(K), edges, Px, vmin=0, vmax=1)
plt.title(r'Estimated CDF $\hat{P}_x(\theta, k)$')
plt.xlabel(r'$k$')
plt.ylabel(r'$\theta$')
plt.colorbar()
plt.autoscale(tight=True)
```
**Exercise**
* Change the number of sample functions `N` or/and the number of `bins` and rerun the examples. What changes? Why?
Solution: In numerical simulations of random processes only a finite number of sample functions and temporal samples can be considered. This holds also for the number of intervals (bins) used for the histogram. As a result, numerical approximations of the CDF/PDF will be subject to statistical uncertainties that typically will become smaller if the number of sample functions `N` is increased.
### Bivariate Probability Density Function
The bivariate or joint PDF $p_{xy}(\theta_x, \theta_y, k_x, k_y)$ of two continuous-amplitude real-valued random signals $x[k]$ and $y[k]$ is defined as
\begin{equation}
p_{xy}(\theta_x, \theta_y, k_x, k_y) := \frac{\partial^2}{\partial \theta_x \partial \theta_y} P_{xy}(\theta_x, \theta_y, k_x, k_y)
\end{equation}
This constitutes essentially the generalization of the univariate PDF. The bivariate PDF quantifies the joint probability that $x[k]$ takes the value $\theta_x$ and that $y[k]$ takes the value $\theta_y$ for the entire ensemble of sample functions. Analogous to the univariate case, the bivariate CDF is given by integration
\begin{equation}
P_{xy}(\theta_x, \theta_y, k_x, k_y) = \int\limits_{-\infty}^{\theta_x} \int\limits_{-\infty}^{\theta_y} p_{xy}(\alpha, \beta, k_x, k_y) \, \mathrm{d}\alpha \mathrm{d}\beta
\end{equation}
Due to the properties of the bivariate CDF and its definition the bivariate PDF shows the following properties
\begin{equation}
p_{xy}(\theta_x, \theta_y, k_x, k_y) \geq 0
\end{equation}
and
\begin{equation}
\int_{-\infty}^{\infty} \int_{-\infty}^{\infty} p_{xy}(\theta_x, \theta_y, k_x, k_y) \, \mathrm{d}\theta_x \mathrm{d}\theta_y = 1
\end{equation}
For the special case of one signal only, the bivariate PDF
\begin{equation}
p_{xx}(\theta_1, \theta_2, k_1, k_2) := \frac{\partial^2}{\partial \theta_1 \partial \theta_2} P_{xx}(\theta_1, \theta_2, k_1, k_2)
\end{equation}
describes the probability that the random signal $x[k]$ takes the values $\theta_1$ at time instance $k_1$ and $\theta_2$ at time instance $k_2$. Hence, $p_{xx}(\theta_1, \theta_2, k_1, k_2)$ provides insights into the temporal dependencies of the amplitudes of the random signal $x[k]$.
| github_jupyter |
# Convolutional Neural Networks: Application
Welcome to Course 4's second assignment! In this notebook, you will:
- Implement helper functions that you will use when implementing a TensorFlow model
- Implement a fully functioning ConvNet using TensorFlow
**After this assignment you will be able to:**
- Build and train a ConvNet in TensorFlow for a classification problem
We assume here that you are already familiar with TensorFlow. If you are not, please refer the *TensorFlow Tutorial* of the third week of Course 2 ("*Improving deep neural networks*").
### <font color='darkblue'> Updates to Assignment <font>
#### If you were working on a previous version
* The current notebook filename is version "1a".
* You can find your work in the file directory as version "1".
* To view the file directory, go to the menu "File->Open", and this will open a new tab that shows the file directory.
#### List of Updates
* `initialize_parameters`: added details about tf.get_variable, `eval`. Clarified test case.
* Added explanations for the kernel (filter) stride values, max pooling, and flatten functions.
* Added details about softmax cross entropy with logits.
* Added instructions for creating the Adam Optimizer.
* Added explanation of how to evaluate tensors (optimizer and cost).
* `forward_propagation`: clarified instructions, use "F" to store "flatten" layer.
* Updated print statements and 'expected output' for easier visual comparisons.
* Many thanks to Kevin P. Brown (mentor for the deep learning specialization) for his suggestions on the assignments in this course!
## 1.0 - TensorFlow model
In the previous assignment, you built helper functions using numpy to understand the mechanics behind convolutional neural networks. Most practical applications of deep learning today are built using programming frameworks, which have many built-in functions you can simply call.
As usual, we will start by loading in the packages.
```
import math
import numpy as np
import h5py
import matplotlib.pyplot as plt
import scipy
from PIL import Image
from scipy import ndimage
import tensorflow as tf
from tensorflow.python.framework import ops
from cnn_utils import *
%matplotlib inline
np.random.seed(1)
```
Run the next cell to load the "SIGNS" dataset you are going to use.
```
# Loading the data (signs)
X_train_orig, Y_train_orig, X_test_orig, Y_test_orig, classes = load_dataset()
```
As a reminder, the SIGNS dataset is a collection of 6 signs representing numbers from 0 to 5.
<img src="images/SIGNS.png" style="width:800px;height:300px;">
The next cell will show you an example of a labelled image in the dataset. Feel free to change the value of `index` below and re-run to see different examples.
```
# Example of a picture
index = 6
plt.imshow(X_train_orig[index])
print ("y = " + str(np.squeeze(Y_train_orig[:, index])))
```
In Course 2, you had built a fully-connected network for this dataset. But since this is an image dataset, it is more natural to apply a ConvNet to it.
To get started, let's examine the shapes of your data.
```
X_train = X_train_orig/255.
X_test = X_test_orig/255.
Y_train = convert_to_one_hot(Y_train_orig, 6).T
Y_test = convert_to_one_hot(Y_test_orig, 6).T
print ("number of training examples = " + str(X_train.shape[0]))
print ("number of test examples = " + str(X_test.shape[0]))
print ("X_train shape: " + str(X_train.shape))
print ("Y_train shape: " + str(Y_train.shape))
print ("X_test shape: " + str(X_test.shape))
print ("Y_test shape: " + str(Y_test.shape))
conv_layers = {}
```
### 1.1 - Create placeholders
TensorFlow requires that you create placeholders for the input data that will be fed into the model when running the session.
**Exercise**: Implement the function below to create placeholders for the input image X and the output Y. You should not define the number of training examples for the moment. To do so, you could use "None" as the batch size, it will give you the flexibility to choose it later. Hence X should be of dimension **[None, n_H0, n_W0, n_C0]** and Y should be of dimension **[None, n_y]**. [Hint: search for the tf.placeholder documentation"](https://www.tensorflow.org/api_docs/python/tf/placeholder).
```
# GRADED FUNCTION: create_placeholders
def create_placeholders(n_H0, n_W0, n_C0, n_y):
"""
Creates the placeholders for the tensorflow session.
Arguments:
n_H0 -- scalar, height of an input image
n_W0 -- scalar, width of an input image
n_C0 -- scalar, number of channels of the input
n_y -- scalar, number of classes
Returns:
X -- placeholder for the data input, of shape [None, n_H0, n_W0, n_C0] and dtype "float"
Y -- placeholder for the input labels, of shape [None, n_y] and dtype "float"
"""
### START CODE HERE ### (≈2 lines)
X = tf.placeholder(tf.float32, shape=[None, n_H0, n_W0, n_C0])
Y = tf.placeholder(tf.float32, shape=[None, n_y])
### END CODE HERE ###
return X, Y
X, Y = create_placeholders(64, 64, 3, 6)
print ("X = " + str(X))
print ("Y = " + str(Y))
```
**Expected Output**
<table>
<tr>
<td>
X = Tensor("Placeholder:0", shape=(?, 64, 64, 3), dtype=float32)
</td>
</tr>
<tr>
<td>
Y = Tensor("Placeholder_1:0", shape=(?, 6), dtype=float32)
</td>
</tr>
</table>
### 1.2 - Initialize parameters
You will initialize weights/filters $W1$ and $W2$ using `tf.contrib.layers.xavier_initializer(seed = 0)`. You don't need to worry about bias variables as you will soon see that TensorFlow functions take care of the bias. Note also that you will only initialize the weights/filters for the conv2d functions. TensorFlow initializes the layers for the fully connected part automatically. We will talk more about that later in this assignment.
**Exercise:** Implement initialize_parameters(). The dimensions for each group of filters are provided below. Reminder - to initialize a parameter $W$ of shape [1,2,3,4] in Tensorflow, use:
```python
W = tf.get_variable("W", [1,2,3,4], initializer = ...)
```
#### tf.get_variable()
[Search for the tf.get_variable documentation](https://www.tensorflow.org/api_docs/python/tf/get_variable). Notice that the documentation says:
```
Gets an existing variable with these parameters or create a new one.
```
So we can use this function to create a tensorflow variable with the specified name, but if the variables already exist, it will get the existing variable with that same name.
```
# GRADED FUNCTION: initialize_parameters
def initialize_parameters():
"""
Initializes weight parameters to build a neural network with tensorflow. The shapes are:
W1 : [4, 4, 3, 8]
W2 : [2, 2, 8, 16]
Note that we will hard code the shape values in the function to make the grading simpler.
Normally, functions should take values as inputs rather than hard coding.
Returns:
parameters -- a dictionary of tensors containing W1, W2
"""
tf.set_random_seed(1) # so that your "random" numbers match ours
### START CODE HERE ### (approx. 2 lines of code)
W1 = tf.get_variable("W1", [4, 4, 3, 8], initializer = tf.contrib.layers.xavier_initializer(seed = 0))
W2 = tf.get_variable("W2", [2, 2, 8, 16], initializer = tf.contrib.layers.xavier_initializer(seed = 0))
### END CODE HERE ###
parameters = {"W1": W1,
"W2": W2}
return parameters
tf.reset_default_graph()
with tf.Session() as sess_test:
parameters = initialize_parameters()
init = tf.global_variables_initializer()
sess_test.run(init)
print("W1[1,1,1] = \n" + str(parameters["W1"].eval()[1,1,1]))
print("W1.shape: " + str(parameters["W1"].shape))
print("\n")
print("W2[1,1,1] = \n" + str(parameters["W2"].eval()[1,1,1]))
print("W2.shape: " + str(parameters["W2"].shape))
```
** Expected Output:**
```
W1[1,1,1] =
[ 0.00131723 0.14176141 -0.04434952 0.09197326 0.14984085 -0.03514394
-0.06847463 0.05245192]
W1.shape: (4, 4, 3, 8)
W2[1,1,1] =
[-0.08566415 0.17750949 0.11974221 0.16773748 -0.0830943 -0.08058
-0.00577033 -0.14643836 0.24162132 -0.05857408 -0.19055021 0.1345228
-0.22779644 -0.1601823 -0.16117483 -0.10286498]
W2.shape: (2, 2, 8, 16)
```
### 1.3 - Forward propagation
In TensorFlow, there are built-in functions that implement the convolution steps for you.
- **tf.nn.conv2d(X,W, strides = [1,s,s,1], padding = 'SAME'):** given an input $X$ and a group of filters $W$, this function convolves $W$'s filters on X. The third parameter ([1,s,s,1]) represents the strides for each dimension of the input (m, n_H_prev, n_W_prev, n_C_prev). Normally, you'll choose a stride of 1 for the number of examples (the first value) and for the channels (the fourth value), which is why we wrote the value as `[1,s,s,1]`. You can read the full documentation on [conv2d](https://www.tensorflow.org/api_docs/python/tf/nn/conv2d).
- **tf.nn.max_pool(A, ksize = [1,f,f,1], strides = [1,s,s,1], padding = 'SAME'):** given an input A, this function uses a window of size (f, f) and strides of size (s, s) to carry out max pooling over each window. For max pooling, we usually operate on a single example at a time and a single channel at a time. So the first and fourth value in `[1,f,f,1]` are both 1. You can read the full documentation on [max_pool](https://www.tensorflow.org/api_docs/python/tf/nn/max_pool).
- **tf.nn.relu(Z):** computes the elementwise ReLU of Z (which can be any shape). You can read the full documentation on [relu](https://www.tensorflow.org/api_docs/python/tf/nn/relu).
- **tf.contrib.layers.flatten(P)**: given a tensor "P", this function takes each training (or test) example in the batch and flattens it into a 1D vector.
* If a tensor P has the shape (m,h,w,c), where m is the number of examples (the batch size), it returns a flattened tensor with shape (batch_size, k), where $k=h \times w \times c$. "k" equals the product of all the dimension sizes other than the first dimension.
* For example, given a tensor with dimensions [100,2,3,4], it flattens the tensor to be of shape [100, 24], where 24 = 2 * 3 * 4. You can read the full documentation on [flatten](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/flatten).
- **tf.contrib.layers.fully_connected(F, num_outputs):** given the flattened input F, it returns the output computed using a fully connected layer. You can read the full documentation on [full_connected](https://www.tensorflow.org/api_docs/python/tf/contrib/layers/fully_connected).
In the last function above (`tf.contrib.layers.fully_connected`), the fully connected layer automatically initializes weights in the graph and keeps on training them as you train the model. Hence, you did not need to initialize those weights when initializing the parameters.
#### Window, kernel, filter
The words "window", "kernel", and "filter" are used to refer to the same thing. This is why the parameter `ksize` refers to "kernel size", and we use `(f,f)` to refer to the filter size. Both "kernel" and "filter" refer to the "window."
**Exercise**
Implement the `forward_propagation` function below to build the following model: `CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED`. You should use the functions above.
In detail, we will use the following parameters for all the steps:
- Conv2D: stride 1, padding is "SAME"
- ReLU
- Max pool: Use an 8 by 8 filter size and an 8 by 8 stride, padding is "SAME"
- Conv2D: stride 1, padding is "SAME"
- ReLU
- Max pool: Use a 4 by 4 filter size and a 4 by 4 stride, padding is "SAME"
- Flatten the previous output.
- FULLYCONNECTED (FC) layer: Apply a fully connected layer without an non-linear activation function. Do not call the softmax here. This will result in 6 neurons in the output layer, which then get passed later to a softmax. In TensorFlow, the softmax and cost function are lumped together into a single function, which you'll call in a different function when computing the cost.
```
# GRADED FUNCTION: forward_propagation
def forward_propagation(X, parameters):
"""
Implements the forward propagation for the model:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Note that for simplicity and grading purposes, we'll hard-code some values
such as the stride and kernel (filter) sizes.
Normally, functions should take these values as function parameters.
Arguments:
X -- input dataset placeholder, of shape (input size, number of examples)
parameters -- python dictionary containing your parameters "W1", "W2"
the shapes are given in initialize_parameters
Returns:
Z3 -- the output of the last LINEAR unit
"""
# Retrieve the parameters from the dictionary "parameters"
W1 = parameters['W1']
W2 = parameters['W2']
### START CODE HERE ###
# CONV2D: stride of 1, padding 'SAME'
Z1 = tf.nn.conv2d(X,W1, strides = [1,1,1,1], padding = 'SAME')
# RELU
A1 = tf.nn.relu(Z1)
# MAXPOOL: window 8x8, stride 8, padding 'SAME'
P1 = tf.nn.max_pool(A1, ksize = [1,8,8,1], strides = [1,8,8,1], padding = 'SAME')
# CONV2D: filters W2, stride 1, padding 'SAME'
Z2 = tf.nn.conv2d(P1,W2, strides = [1,1,1,1], padding = 'SAME')
# RELU
A2 = tf.nn.relu(Z2)
# MAXPOOL: window 4x4, stride 4, padding 'SAME'
P2 = tf.nn.max_pool(A2, ksize = [1,4,4,1], strides = [1,4,4,1], padding = 'SAME')
# FLATTEN
F = tf.contrib.layers.flatten(P2)
# FULLY-CONNECTED without non-linear activation function (not not call softmax).
# 6 neurons in output layer. Hint: one of the arguments should be "activation_fn=None"
Z3 = tf.contrib.layers.fully_connected(F, 6, activation_fn=None)
### END CODE HERE ###
return Z3
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(Z3, {X: np.random.randn(2,64,64,3), Y: np.random.randn(2,6)})
print("Z3 = \n" + str(a))
```
**Expected Output**:
```
Z3 =
[[-0.44670227 -1.57208765 -1.53049231 -2.31013036 -1.29104376 0.46852064]
[-0.17601591 -1.57972014 -1.4737016 -2.61672091 -1.00810647 0.5747785 ]]
```
### 1.4 - Compute cost
Implement the compute cost function below. Remember that the cost function helps the neural network see how much the model's predictions differ from the correct labels. By adjusting the weights of the network to reduce the cost, the neural network can improve its predictions.
You might find these two functions helpful:
- **tf.nn.softmax_cross_entropy_with_logits(logits = Z, labels = Y):** computes the softmax entropy loss. This function both computes the softmax activation function as well as the resulting loss. You can check the full documentation [softmax_cross_entropy_with_logits](https://www.tensorflow.org/api_docs/python/tf/nn/softmax_cross_entropy_with_logits).
- **tf.reduce_mean:** computes the mean of elements across dimensions of a tensor. Use this to calculate the sum of the losses over all the examples to get the overall cost. You can check the full documentation [reduce_mean](https://www.tensorflow.org/api_docs/python/tf/reduce_mean).
#### Details on softmax_cross_entropy_with_logits (optional reading)
* Softmax is used to format outputs so that they can be used for classification. It assigns a value between 0 and 1 for each category, where the sum of all prediction values (across all possible categories) equals 1.
* Cross Entropy is compares the model's predicted classifications with the actual labels and results in a numerical value representing the "loss" of the model's predictions.
* "Logits" are the result of multiplying the weights and adding the biases. Logits are passed through an activation function (such as a relu), and the result is called the "activation."
* The function is named `softmax_cross_entropy_with_logits` takes logits as input (and not activations); then uses the model to predict using softmax, and then compares the predictions with the true labels using cross entropy. These are done with a single function to optimize the calculations.
** Exercise**: Compute the cost below using the function above.
```
# GRADED FUNCTION: compute_cost
def compute_cost(Z3, Y):
"""
Computes the cost
Arguments:
Z3 -- output of forward propagation (output of the last LINEAR unit), of shape (number of examples, 6)
Y -- "true" labels vector placeholder, same shape as Z3
Returns:
cost - Tensor of the cost function
"""
### START CODE HERE ### (1 line of code)
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(logits = Z3, labels = Y))
### END CODE HERE ###
return cost
tf.reset_default_graph()
with tf.Session() as sess:
np.random.seed(1)
X, Y = create_placeholders(64, 64, 3, 6)
parameters = initialize_parameters()
Z3 = forward_propagation(X, parameters)
cost = compute_cost(Z3, Y)
init = tf.global_variables_initializer()
sess.run(init)
a = sess.run(cost, {X: np.random.randn(4,64,64,3), Y: np.random.randn(4,6)})
print("cost = " + str(a))
```
**Expected Output**:
```
cost = 2.91034
```
## 1.5 Model
Finally you will merge the helper functions you implemented above to build a model. You will train it on the SIGNS dataset.
**Exercise**: Complete the function below.
The model below should:
- create placeholders
- initialize parameters
- forward propagate
- compute the cost
- create an optimizer
Finally you will create a session and run a for loop for num_epochs, get the mini-batches, and then for each mini-batch you will optimize the function. [Hint for initializing the variables](https://www.tensorflow.org/api_docs/python/tf/global_variables_initializer)
#### Adam Optimizer
You can use `tf.train.AdamOptimizer(learning_rate = ...)` to create the optimizer. The optimizer has a `minimize(loss=...)` function that you'll call to set the cost function that the optimizer will minimize.
For details, check out the documentation for [Adam Optimizer](https://www.tensorflow.org/api_docs/python/tf/train/AdamOptimizer)
#### Random mini batches
If you took course 2 of the deep learning specialization, you implemented `random_mini_batches()` in the "Optimization" programming assignment. This function returns a list of mini-batches. It is already implemented in the `cnn_utils.py` file and imported here, so you can call it like this:
```Python
minibatches = random_mini_batches(X, Y, mini_batch_size = 64, seed = 0)
```
(You will want to choose the correct variable names when you use it in your code).
#### Evaluating the optimizer and cost
Within a loop, for each mini-batch, you'll use the `tf.Session` object (named `sess`) to feed a mini-batch of inputs and labels into the neural network and evaluate the tensors for the optimizer as well as the cost. Remember that we built a graph data structure and need to feed it inputs and labels and use `sess.run()` in order to get values for the optimizer and cost.
You'll use this kind of syntax:
```
output_for_var1, output_for_var2 = sess.run(
fetches=[var1, var2],
feed_dict={var_inputs: the_batch_of_inputs,
var_labels: the_batch_of_labels}
)
```
* Notice that `sess.run` takes its first argument `fetches` as a list of objects that you want it to evaluate (in this case, we want to evaluate the optimizer and the cost).
* It also takes a dictionary for the `feed_dict` parameter.
* The keys are the `tf.placeholder` variables that we created in the `create_placeholders` function above.
* The values are the variables holding the actual numpy arrays for each mini-batch.
* The sess.run outputs a tuple of the evaluated tensors, in the same order as the list given to `fetches`.
For more information on how to use sess.run, see the documentation [tf.Sesssion#run](https://www.tensorflow.org/api_docs/python/tf/Session#run) documentation.
```
# GRADED FUNCTION: model
def model(X_train, Y_train, X_test, Y_test, learning_rate = 0.009,
num_epochs = 100, minibatch_size = 64, print_cost = True):
"""
Implements a three-layer ConvNet in Tensorflow:
CONV2D -> RELU -> MAXPOOL -> CONV2D -> RELU -> MAXPOOL -> FLATTEN -> FULLYCONNECTED
Arguments:
X_train -- training set, of shape (None, 64, 64, 3)
Y_train -- test set, of shape (None, n_y = 6)
X_test -- training set, of shape (None, 64, 64, 3)
Y_test -- test set, of shape (None, n_y = 6)
learning_rate -- learning rate of the optimization
num_epochs -- number of epochs of the optimization loop
minibatch_size -- size of a minibatch
print_cost -- True to print the cost every 100 epochs
Returns:
train_accuracy -- real number, accuracy on the train set (X_train)
test_accuracy -- real number, testing accuracy on the test set (X_test)
parameters -- parameters learnt by the model. They can then be used to predict.
"""
ops.reset_default_graph() # to be able to rerun the model without overwriting tf variables
tf.set_random_seed(1) # to keep results consistent (tensorflow seed)
seed = 3 # to keep results consistent (numpy seed)
(m, n_H0, n_W0, n_C0) = X_train.shape
n_y = Y_train.shape[1]
costs = [] # To keep track of the cost
# Create Placeholders of the correct shape
### START CODE HERE ### (1 line)
X, Y = create_placeholders(n_H0, n_W0, n_C0, n_y)
### END CODE HERE ###
# Initialize parameters
### START CODE HERE ### (1 line)
parameters = initialize_parameters()
### END CODE HERE ###
# Forward propagation: Build the forward propagation in the tensorflow graph
### START CODE HERE ### (1 line)
Z3 = forward_propagation(X, parameters)
### END CODE HERE ###
# Cost function: Add cost function to tensorflow graph
### START CODE HERE ### (1 line)
cost = compute_cost(Z3, Y)
### END CODE HERE ###
# Backpropagation: Define the tensorflow optimizer. Use an AdamOptimizer that minimizes the cost.
### START CODE HERE ### (1 line)
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost)
### END CODE HERE ###
# Initialize all the variables globally
init = tf.global_variables_initializer()
# Start the session to compute the tensorflow graph
with tf.Session() as sess:
# Run the initialization
sess.run(init)
# Do the training loop
for epoch in range(num_epochs):
minibatch_cost = 0.
num_minibatches = int(m / minibatch_size) # number of minibatches of size minibatch_size in the train set
seed = seed + 1
minibatches = random_mini_batches(X_train, Y_train, minibatch_size, seed)
for minibatch in minibatches:
# Select a minibatch
(minibatch_X, minibatch_Y) = minibatch
"""
# IMPORTANT: The line that runs the graph on a minibatch.
# Run the session to execute the optimizer and the cost.
# The feedict should contain a minibatch for (X,Y).
"""
### START CODE HERE ### (1 line)
_ , temp_cost = sess.run([optimizer, cost], feed_dict={X: minibatch_X, Y: minibatch_Y})
### END CODE HERE ###
minibatch_cost += temp_cost / num_minibatches
# Print the cost every epoch
if print_cost == True and epoch % 5 == 0:
print ("Cost after epoch %i: %f" % (epoch, minibatch_cost))
if print_cost == True and epoch % 1 == 0:
costs.append(minibatch_cost)
# plot the cost
plt.plot(np.squeeze(costs))
plt.ylabel('cost')
plt.xlabel('iterations (per tens)')
plt.title("Learning rate =" + str(learning_rate))
plt.show()
# Calculate the correct predictions
predict_op = tf.argmax(Z3, 1)
correct_prediction = tf.equal(predict_op, tf.argmax(Y, 1))
# Calculate accuracy on the test set
accuracy = tf.reduce_mean(tf.cast(correct_prediction, "float"))
print(accuracy)
train_accuracy = accuracy.eval({X: X_train, Y: Y_train})
test_accuracy = accuracy.eval({X: X_test, Y: Y_test})
print("Train Accuracy:", train_accuracy)
print("Test Accuracy:", test_accuracy)
return train_accuracy, test_accuracy, parameters
```
Run the following cell to train your model for 100 epochs. Check if your cost after epoch 0 and 5 matches our output. If not, stop the cell and go back to your code!
```
_, _, parameters = model(X_train, Y_train, X_test, Y_test)
```
**Expected output**: although it may not match perfectly, your expected output should be close to ours and your cost value should decrease.
<table>
<tr>
<td>
**Cost after epoch 0 =**
</td>
<td>
1.917929
</td>
</tr>
<tr>
<td>
**Cost after epoch 5 =**
</td>
<td>
1.506757
</td>
</tr>
<tr>
<td>
**Train Accuracy =**
</td>
<td>
0.940741
</td>
</tr>
<tr>
<td>
**Test Accuracy =**
</td>
<td>
0.783333
</td>
</tr>
</table>
Congratulations! You have finished the assignment and built a model that recognizes SIGN language with almost 80% accuracy on the test set. If you wish, feel free to play around with this dataset further. You can actually improve its accuracy by spending more time tuning the hyperparameters, or using regularization (as this model clearly has a high variance).
Once again, here's a thumbs up for your work!
```
fname = "images/thumbs_up.jpg"
image = np.array(ndimage.imread(fname, flatten=False))
my_image = scipy.misc.imresize(image, size=(64,64))
plt.imshow(my_image)
```
| github_jupyter |
# Chat Intents
## Applying labels
**Summary**
This notebook provides a way to automatically extract and apply labels to document clusters. See the `chatintents_tutorial.ipynb` notebook for a tutorial of the chatintents package, which simplifies and makes it easier to use the methods outlined below.
```
import collections
from pathlib import Path
import numpy as np
import pandas as pd
import spacy
from spacy import displacy
pd.set_option("display.max_rows", 600)
pd.set_option("display.max_columns", 500)
pd.set_option("max_colwidth", 400)
nlp = spacy.load("en_core_web_sm")
data_clustered = pd.read_csv('../data/processed/sample_clustered.csv')
data_clustered = data_clustered[['text', 'label_st1']]
data_clustered.sample(10)
example_category = data_clustered[data_clustered['label_st1']==31].reset_index(drop=True)
example_category
example_doc = nlp(list(example_category['text'])[12])
print(f'{example_doc}\n')
for token in example_doc:
print(token.text, token.lemma_, token.pos_, token.tag_, token.dep_ , token.is_stop)
displacy.render(example_doc, style="dep")
fig = displacy.render(example_doc, style="dep", jupyter=False)
output_path = Path("../images/dependency_plot.svg") # you can keep there only "dependency_plot.svg" if you want to save it in the same folder where you run the script
output_path.open("w", encoding="utf-8").write(fig)
```
## Helper functions
```
def get_group(df, category_col, category):
"""
Returns documents of a single category
Arguments:
df: pandas dataframe of documents
category_col: str, column name corresponding to categories or clusters
category: int, cluster number to return
Returns:
single_category: pandas dataframe with documents from a single category
"""
single_category = df[df[category_col]==category].reset_index(drop=True)
return single_category
def most_common(lst, n_words):
"""
Get most common words in a list of words
Arguments:
lst: list, each element is a word
n_words: number of top common words to return
Returns:
counter.most_common(n_words): counter object of n most common words
"""
counter=collections.Counter(lst)
return counter.most_common(n_words)
def extract_labels(category_docs, print_word_counts=False):
"""
Extract labels from documents in the same cluster by concatenating
most common verbs, ojects, and nouns
Argument:
category_docs: list of documents, all from the same category or
clustering
print_word_counts: bool, True will print word counts of each type in this category
Returns:
label: str, group label derived from concatentating most common
verb, object, and two most common nouns
"""
verbs = []
dobjs = []
nouns = []
adjs = []
verb = ''
dobj = ''
noun1 = ''
noun2 = ''
# for each document, append verbs, dobs, nouns, and adjectives to
# running lists for whole cluster
for i in range(len(category_docs)):
doc = nlp(category_docs[i])
for token in doc:
if token.is_stop==False:
if token.dep_ == 'ROOT':
verbs.append(token.text.lower())
elif token.dep_=='dobj':
dobjs.append(token.lemma_.lower())
elif token.pos_=='NOUN':
nouns.append(token.lemma_.lower())
elif token.pos_=='ADJ':
adjs.append(token.lemma_.lower())
# for printing out for inspection purposes
if print_word_counts:
for word_lst in [verbs, dobjs, nouns, adjs]:
counter=collections.Counter(word_lst)
print(counter)
# take most common words of each form
if len(verbs) > 0:
verb = most_common(verbs, 1)[0][0]
if len(dobjs) > 0:
dobj = most_common(dobjs, 1)[0][0]
if len(nouns) > 0:
noun1 = most_common(nouns, 1)[0][0]
if len(set(nouns)) > 1:
noun2 = most_common(nouns, 2)[1][0]
# concatenate the most common verb-dobj-noun1-noun2 (if they exist)
label_words = [verb, dobj]
for word in [noun1, noun2]:
if word not in label_words:
label_words.append(word)
if '' in label_words:
label_words.remove('')
label = '_'.join(label_words)
return label
def apply_and_summarize_labels(df, category_col):
"""
Assign groups to original documents and provide group counts
Arguments:
df: pandas dataframe of original documents of interest to
cluster
category_col: str, column name corresponding to categories or clusters
Returns:
summary_df: pandas dataframe with model cluster assignment, number
of documents in each cluster and derived labels
"""
numerical_labels = df[category_col].unique()
# create dictionary of the numerical category to the generated label
label_dict = {}
for label in numerical_labels:
current_category = list(get_group(df, category_col, label)['text'])
label_dict[label] = extract_labels(current_category)
# create summary dataframe of numerical labels and counts
summary_df = (df.groupby(category_col)['text'].count()
.reset_index()
.rename(columns={'text':'count'})
.sort_values('count', ascending=False))
# apply generated labels
summary_df['label'] = summary_df.apply(lambda x: label_dict[x[category_col]], axis = 1)
return summary_df
def combine_ground_truth(df_clusters, df_ground, key):
"""
Combines dataframes of documents with extracted and ground truth labels
Arguments:
df_clusters: pandas dataframe, each row as a document with corresponding extracted label
df_ground: pandas dataframe, each row as a document with corresponding ground truth label
key: str, key to merge tables on
Returns:
df_combined: pandas dataframe, each row as a document with extracted and ground truth labels
"""
df_combined = pd.merge(df_clusters, df_ground, on=key, how = 'left')
return df_combined
def get_top_category(df_label, df_summary):
"""
Returns a dataframe comparing a single model's results to ground truth
label to evalute cluster compositions and derived label relative to labels
and counts of most commmon ground truth category
Arguments:
df_label: pandas dataframe, each row as a document with extracted and ground truth labels
(result of `combine_ground_truth` function)
df_summary: pandas dataframe with model cluster assignment, number
of documents in each cluster and derived labels
(result from `apply_and_summarize_labels` function)
Returns:
df_result: pandas dataframe with each row containing information on
each cluster identified by this model, including count,
extracted label, most represented ground truth label name,
count and percentage of that group
"""
df_label_ground = (df_label.groupby('label')
.agg(top_ground_category=('category', lambda x:x.value_counts().index[0]),
top_cat_count = ('category', lambda x:x.value_counts()[0]))
.reset_index())
df_result = pd.merge(df_summary, df_label_ground, on='label', how='left')
df_result['perc_top_cat'] = df_result.apply(lambda x: int(round(100*x['top_cat_count']/x['count'])), axis=1)
return df_result
```
### Manual inspection
```
example_category = list(get_group(data_clustered, 'label_st1', 46)['text'])
extract_labels(example_category, True)
```
### Without ground truth labels
```
cluster_summary = apply_and_summarize_labels(data_clustered, 'label_st1')
cluster_summary.head(20)
labeled_clusters = pd.merge(data_clustered, cluster_summary[['label_st1', 'label']], on='label_st1', how = 'left')
labeled_clusters.head()
```
If we don't have the ground truth labels (which is the primary use case for this), then the above tables would be the final results. In this case, since we do have the ground truth labels we can investigate how well our model did.
### With ground truth labels
```
data_ground = pd.read_csv('../data/processed/data_sample.csv')[['text', 'category']]
data_ground.head()
labeled_clusters = combine_ground_truth(labeled_clusters, data_ground, 'text')
labeled_clusters.sample(10)
```
The extracted labels (called 'label') match the ground label ('category') quite well for many of the sample documents.
```
labeled_clusters[labeled_clusters['label_st1']==45]
```
#### Count and name of most common category of generated labels and clusters
```
get_top_category(labeled_clusters, cluster_summary)
```
Many of the smaller groups seem to be more pure (the top category is near or at 100%) compared to the larger groups. Thus, it makes sense that many of the extracted labels for the smaller groups tend to be more suitable than some of the extracted labels for the larger clusters, which have more varied representation of different ground truth clusters.
| github_jupyter |
```
import pydicom #read dicom files
import os
import pandas as pd
data_dir = 'C:/Users/casti/Documents/Final_Project/FFE_imagesOversampling/'
patients = os.listdir(data_dir)
labels_df = pd.read_csv('C:/Users/casti/Documents/Final_Project/animallist_Oversampling.csv', index_col=0)
labels_df.head()
for patient in patients:
label = labels_df.at[patient, 'tumor_model']
path = data_dir + patient
slices = [pydicom.read_file(path + '/' + s) for s in os.listdir(path)]
slices.sort(key = lambda x: float(x.ImagePositionPatient[2]))
print(patient, len(slices), slices[0].pixel_array.shape)
# print(slices[0])
len(patients)
import matplotlib.pyplot as plt
import cv2
import numpy as np
import math
%matplotlib inline
IMG_PX_SIZE = 50
HM_SLICES = 20
def chunks(l, n):
#yield succesive n-sized chunks from l. source: Ned Batchelder.
for i in range(0, len(l), n):
yield l[i:i+n]
def mean(l):
return sum(l)/len(l)
def process_data(patient, labels_df, img_px_size=50, hm_slices=20, visualize=False):
label = labels_df.at[patient, 'tumor_model']
path = data_dir + patient
slices = [pydicom.read_file(path + '/' + s) for s in os.listdir(path)]
slices.sort(key = lambda x: float(x.ImagePositionPatient[2]))
new_slices = []
slices = [cv2.resize(np.array(each_slice.pixel_array),(IMG_PX_SIZE, IMG_PX_SIZE)) for each_slice in slices]
chunk_sizes = math.ceil(len(slices) / HM_SLICES)
for slice_chunk in chunks(slices, chunk_sizes):
slice_chunk = list(map(mean, zip(*slice_chunk)))
new_slices.append(slice_chunk)
if len(new_slices) == HM_SLICES-1:
new_slices.append(new_slices[-1])
if len(new_slices) == HM_SLICES-2:
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
if len(new_slices) == HM_SLICES-3:
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
if len(new_slices) == HM_SLICES-4:
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
if len(new_slices) == HM_SLICES-5:
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
new_slices.append(new_slices[-1])
if len(new_slices) == HM_SLICES+2:
new_val=list(map(mean), zip(*[new_slices[HM_SLICES-1], new_slices[HM_SLICES]]))
del new_slices[HM_SLICES]
new_slices[HM_SLICES-1]=new_val
if len(new_slices) == HM_SLICES+2:
new_val=list(map(mean), zip(*[new_slices[HM_SLICES-1], new_slices[HM_SLICES]]))
del new_slices[HM_SLICES]
new_slices[HM_SLICES-1]=new_val
if visualize:
fig = plt.figure(figsize=(20,20))
for num,each_slice in enumerate(new_slices[:1]):
y = fig.add_subplot(4, 5, num + 1)
y.imshow(each_slice)
plt.show()
if label == 1:
label = np.array([0,1])
elif label == 0:
label = np.array([1,0])
return np.array(new_slices), label
much_data = []
for num, patient in enumerate(patients):
if num%20==0:
print(num)
try:
img_data, label = process_data(patient, labels_df, img_px_size=IMG_PX_SIZE, hm_slices=HM_SLICES)
much_data.append([img_data, label])
except KeyError as e:
print('this is unlabeled data')
np.save('muchdata-{}-{}-{}.npy'.format(IMG_PX_SIZE, IMG_PX_SIZE, HM_SLICES), much_data)
from sklearn.model_selection import train_test_split
import tensorflow as tf
from tensorflow.keras.models import Model
import numpy as np
from scipy import ndimage, misc
import matplotlib.pyplot as plt
import tensorflow.keras as keras
from tensorflow.keras import backend as K
from tensorflow.keras.metrics import categorical_crossentropy
from tensorflow.keras.preprocessing import image
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense,GlobalAveragePooling2D
from tensorflow.keras.applications import MobileNet#, imagenet_utils
from tensorflow.keras.applications.mobilenet import preprocess_input
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.layers import Input, Dense, Activation
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.applications.mobilenet import preprocess_input
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import os
import cv2
from tensorflow.keras.applications import VGG16
from tensorflow.keras.layers import AveragePooling3D
from tensorflow.keras.layers import Dropout
from tensorflow.keras.layers import Flatten
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import Input
from tensorflow.keras.models import Model
from tensorflow.keras.datasets import mnist
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Activation, Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.utils import to_categorical
IMG_SIZE_PX=IMG_PX_SIZE
SLICE_COUNT=HM_SLICES
n_classes=2
x = tf.placeholder('float')
y = tf.placeholder('float')
keep_rate = 0.8
keep_prob = tf.placeholder(tf.float32)
def conv3d(x, W):
return tf.nn.conv3d(x, W, strides=[1,1,1,1,1], padding='SAME')
def maxpool3d(x):
# size of window movement of window
return tf.nn.max_pool3d(x, ksize=[1,2,2,2,1], strides=[1,2,2,2,1], padding='SAME')
def convolutional_neural_network(x):
weights = {'W_conv1':tf.Variable(tf.random_normal([3,3,3,1,32])),
'W_conv2':tf.Variable(tf.random_normal([3,3,3,32,64])),
'W_fc':tf.Variable(tf.random_normal([54080,1024])),
'out':tf.Variable(tf.random_normal([1024, n_classes]))}
biases = {'b_conv1':tf.Variable(tf.random_normal([32])),
'b_conv2':tf.Variable(tf.random_normal([64])),
'b_fc':tf.Variable(tf.random_normal([1024])),
'out':tf.Variable(tf.random_normal([n_classes]))}
x = tf.reshape(x, shape=[-1, IMG_SIZE_PX, IMG_SIZE_PX, SLICE_COUNT, 1])
conv1 = tf.nn.relu(conv3d(x, weights['W_conv1']) + biases['b_conv1'])
conv1 = maxpool3d(conv1)
conv2 = tf.nn.relu(conv3d(conv1, weights['W_conv2']) + biases['b_conv2'])
conv2 = maxpool3d(conv2)
fc = tf.reshape(conv2,[-1, 54080])
fc = tf.nn.relu(tf.matmul(fc, weights['W_fc'])+biases['b_fc'])
fc = tf.nn.dropout(fc, keep_rate)
output = tf.matmul(fc, weights['out'])+biases['out']
print(fc)
return output
def train_neural_network(x):
much_data = np.load('muchdata-{}-{}-{}.npy'.format(IMG_PX_SIZE, IMG_PX_SIZE, HM_SLICES), allow_pickle=True)
train_data, validation_data = train_test_split(much_data, random_state=42, test_size=0.25)
prediction = convolutional_neural_network(x)
cost = tf.reduce_mean( tf.nn.sigmoid_cross_entropy_with_logits(labels=y,logits=prediction))
optimizer = tf.train.AdamOptimizer().minimize(cost)
hm_epochs = 100
Accuracy_TRAIN=[]
Accuracy_TEST=[]
LossList=[]
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for epoch in range(hm_epochs):
epoch_loss = 0
success_total=0
attemp_total=0
for data in train_data:
attemp_total +=1
try:
X = data[0]
Y = data[1]
_, c = sess.run([optimizer, cost], feed_dict={x: X, y: Y})
epoch_loss += c
success_total = success_total+1
except Exception as e:
pass
print('Epoch', epoch+1, 'completed out of',hm_epochs,'loss:',epoch_loss, 'success rate:', success_total/attemp_total)
correct = tf.equal(tf.argmax(prediction, 1), tf.argmax(y, 1))
accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
#print('Accuracy_trainig_set:', accuracy)
Accuracytrain = accuracy.eval({x:[i[0] for i in train_data], y:[i[1] for i in train_data]})
print('Accuracy_train:', Accuracytrain)
Accuracytest = accuracy.eval({x:[i[0] for i in validation_data], y:[i[1] for i in validation_data]})
print('Accuracy_test:', Accuracytest)
Accuracy_TRAIN.append(Accuracytrain)
Accuracy_TEST.append(Accuracytest)
LossList.append(epoch_loss)
print('Accuracy training list:', Accuracy_TRAIN)
print('')
print('Accuracy test list:',Accuracy_TEST)
print('')
print('Loss list:',LossList)
# correct = tf.equal(tf.argmax(prediction, 1), tf.argmax(y, 1))
# accuracy = tf.reduce_mean(tf.cast(correct, 'float'))
# print('Accuracy:',accuracy.eval({x:[i[0] for i in validation_data], y:[i[1] for i in validation_data]}))
train_neural_network(x)
from tensorflow.keras.datasets import mnist
from tensorflow.keras import Sequential
from tensorflow.keras.layers import Dense, Activation, Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.utils import to_categorical
baseModel = VGG16(weights="imagenet", include_top=False,
input_tensor=Input(shape=(254, 254, 3)))
headModel = baseModel.output
headModel = AveragePooling2D(pool_size=(2, 2, 2), strides=[1,1,1], padding='SAME')(headModel)
headModel = Flatten(name="flatten")(headModel)
headModel = Dense(64, activation="relu")(headModel)
headModel = Dropout(0.5)(headModel)
headModel = Dense(2, activation="softmax")(headModel)
model = Model(inputs=baseModel.input, outputs=headModel)
for layer in baseModel.layers:
layer.trainable = True
```
| github_jupyter |
# The $\chi^2$ Distribution
## $\chi^2$ Test Statistic
If we make $n$ ranom samples (observations) from Gaussian (Normal) distributions with known means, $\mu_i$, and known variances, $\sigma_i^2$, it is seen that the total squared deviation,
$$
\chi^2 = \sum_{i=1}^{n} \left(\frac{x_i - \mu_i}{\sigma_i}\right)^2\,,
$$
follows a $\chi^2$ distribution with $n$ degrees of freedom.
## Probability Distribution Function
The $\chi^2$ probability distribution function for $k$ degrees of freedom (the number of parameters that are allowed to vary) is given by
$$
f\left(\chi^2\,;k\right) = \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, \chi^{k-2}\,e^{-\chi^2/2}\,,
$$
where if there are no constrained variables the number of degrees of freedom, $k$, is equal to the number of observations, $k=n$. The p.d.f. is often abbreviated in notation from $f\left(\chi^2\,;k\right)$ to $\chi^2_k$.
A reminder that for integer values of $k$, the Gamma function is $\Gamma\left(k\right) = \left(k-1\right)!$, and that $\Gamma\left(x+1\right) = x\Gamma\left(x\right)$, and $\Gamma\left(1/2\right) = \sqrt{\pi}$.
## Mean
Letting $\chi^2=z$, and noting that the form of the Gamma function is
$$
\Gamma\left(z\right) = \int\limits_{0}^{\infty} x^{z-1}\,e^{-x}\,dx,
$$
it is seen that the mean of the $\chi^2$ distribution $f\left(\chi^2 ; k\right)$ is
$$
\begin{align}
\mu &= \textrm{E}\left[z\right] = \displaystyle\int\limits_{0}^{\infty} z\, \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, z^{k/2-1}\,e^{-z\,/2}\,dz \\
&= \displaystyle \frac{\displaystyle 1}{\displaystyle \Gamma\left(k\,/2\right)} \int\limits_{0}^{\infty} \left(\frac{z}{2}\right)^{k/2}\,e^{-z\,/2}\,dz = \displaystyle \frac{\displaystyle 1}{\displaystyle \Gamma\left(k\,/2\right)} \int\limits_{0}^{\infty} x^{k/2}\,e^{-x}\,2 \,dx \\
&= \displaystyle \frac{\displaystyle 2 \,\Gamma\left(k\,/2 + 1\right)}{\displaystyle \Gamma\left(k\,/2\right)} \\
&= \displaystyle 2 \frac{k}{2} \frac{\displaystyle \Gamma\left(k\,/2\right)}{\displaystyle \Gamma\left(k\,/2\right)} \\
&= k.
\end{align}
$$
## Variance
Likewise, the variance is
$$
\begin{align}
\textrm{Var}\left[z\right] &= \textrm{E}\left[\left(z-\textrm{E}\left[z\right]\right)^2\right] = \displaystyle\int\limits_{0}^{\infty} \left(z - k\right)^2\, \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, z^{k/2-1}\,e^{-z\,/2}\,dz \\
&= \displaystyle\int\limits_{0}^{\infty} z^2\, f\left(z \,; k\right)\,dz - 2k\int\limits_{0}^{\infty} z\,\,f\left(z \,; k\right)\,dz + k^2\int\limits_{0}^{\infty} f\left(z \,; k\right)\,dz \\
&= \displaystyle\int\limits_{0}^{\infty} z^2 \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, z^{k/2-1}\,e^{-z\,/2}\,dz - 2k^2 + k^2\\
&= \displaystyle\int\limits_{0}^{\infty} \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, z^{k/2+1}\,e^{-z\,/2}\,dz - k^2\\
&= \frac{\displaystyle 2}{\displaystyle \Gamma\left(k\,/2\right)} \displaystyle\int\limits_{0}^{\infty} \left(\frac{z}{2}\right)^{k/2+1}\,e^{-z\,/2}\,dz - k^2 = \frac{\displaystyle 2}{\displaystyle \Gamma\left(k\,/2\right)} \displaystyle\int\limits_{0}^{\infty} x^{k/2+1}\,e^{-x}\,2\,dx - k^2 \\
&= \displaystyle \frac{\displaystyle 4 \,\Gamma\left(k\,/2 + 2\right)}{\displaystyle \Gamma\left(k\,/2\right)} - k^2 \\
&= \displaystyle 4 \left(\frac{k}{2} + 1\right) \frac{\displaystyle \Gamma\left(k\,/2 + 1\right)}{\displaystyle \Gamma\left(k\,/2\right)} - k^2 \\
&= \displaystyle 4 \left(\frac{k}{2} + 1\right) \frac{k}{2} - k^2 \\
&= k^2 + 2k - k^2 \\
&= 2k,
\end{align}
$$
such that the standard deviation is
$$
\sigma = \sqrt{2k}\,.
$$
Given this information we now plot the $\chi^2$ p.d.f. with various numbers of degrees of freedom to visualize how the distribution's behaviour
```
import numpy as np
import scipy.stats as stats
import matplotlib.pyplot as plt
# Plot the chi^2 distribution
x = np.linspace(0.0, 10.0, num=1000)
[plt.plot(x, stats.chi2.pdf(x, df=ndf), label=fr"$k = ${ndf}") for ndf in range(1, 7)]
plt.ylim(-0.01, 0.5)
plt.xlabel(r"$x=\chi^2$")
plt.ylabel(r"$f\left(x;k\right)$")
plt.title(r"$\chi^2$ distribution for various degrees of freedom")
plt.legend(loc="best")
plt.show();
```
## Cumulative Distribution Function
The cumulative distribution function (CDF) for the $\chi^2$ distribution is (letting $z=\chi^2$)
$$
\begin{split}
F_{\chi^2}\left(x\,; k\right) &= \int\limits_{0}^{x} f_{\chi^2}\left(z\,; k\right) \,dz \\
&= \int\limits_{0}^{x} \frac{\displaystyle 1}{\displaystyle 2^{k/2} \,\Gamma\left(k\,/2\right)}\, z^{k/2-1}\,e^{-z/2} \,dz \\
&= \int\limits_{0}^{x} \frac{\displaystyle 1}{\displaystyle 2 \,\Gamma\left(k\,/2\right)}\, \left(\frac{z}{2}\right)^{k/2-1}\,e^{-z/2} \,dz = \frac{1}{\displaystyle 2 \,\Gamma\left(k\,/2\right)}\int\limits_{0}^{x/2} t^{k/2-1}\,e^{-t} \,2\,dt \\
&= \frac{1}{\displaystyle \Gamma\left(k\,/2\right)}\int\limits_{0}^{x/2} t^{k/2-1}\,e^{-t} \,dt
\end{split}
$$
Noting the form of the [lower incomplete gamma function](https://en.wikipedia.org/wiki/Incomplete_gamma_function) is
$$
\gamma\left(s,x\right) = \int\limits_{0}^{x} t^{s-1}\,e^{-t} \,dt\,,
$$
and the form of the [regularized Gamma function](https://en.wikipedia.org/wiki/Incomplete_gamma_function#Regularized_Gamma_functions_and_Poisson_random_variables) is
$$
P\left(s,x\right) = \frac{\gamma\left(s,x\right)}{\Gamma\left(s\right)}\,,
$$
it is seen that
$$
\begin{split}
F_{\chi^2}\left(x\,; k\right) &= \frac{1}{\displaystyle \Gamma\left(k\,/2\right)}\int\limits_{0}^{x/2} t^{k/2-1}\,e^{-t} \,dt \\
&= \frac{\displaystyle \gamma\left(\frac{k}{2},\frac{x}{2}\right)}{\displaystyle \Gamma\left(\frac{k}{2}\right)} \\
&= P\left(\frac{k}{2},\frac{x}{2}\right)\,.
\end{split}
$$
Thus, it is seen that the compliment to the CDF (the complementary cumulative distribution function (CCDF)),
$$
\bar{F}_{\chi^2}\left(x\,; k\right) = 1-F_{\chi^2}\left(x\,; k\right),
$$
represents a one-sided (one-tailed) $p$-value for observing a $\chi^2$ given a model — that is, the probability to observe a $\chi^2$ value greater than or equal to that which was observed.
```
def chi2_ccdf(x, df):
"""The complementary cumulative distribution function
Args:
x: the value of chi^2
df: the number of degrees of freedom
Returns:
1 - the cumulative distribution function
"""
return 1.0 - stats.chi2.cdf(x=x, df=df)
x = np.linspace(0.0, 10.0, num=1000)
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(14, 4.5))
for ndf in range(1, 7):
axes[0].plot(x, stats.chi2.cdf(x, df=ndf), label=fr"$k = ${ndf}")
axes[1].plot(x, chi2_ccdf(x, df=ndf), label=fr"$k = ${ndf}")
axes[0].set_xlabel(r"$x=\chi^2$")
axes[0].set_ylabel(r"$F\left(x;k\right)$")
axes[0].set_title(r"$\chi^2$ CDF for various degrees of freedom")
axes[0].legend(loc="best")
axes[1].set_xlabel(r"$x=\chi^2$")
axes[1].set_ylabel(r"$\bar{F}\left(x;k\right) = p$-value")
axes[1].set_title(r"$\chi^2$ CCDF ($p$-value) for various degrees of freedom")
axes[1].legend(loc="best")
plt.show();
```
## Binned $\chi^2$ per Degree of Freedom
TODO
## References
- \[1\] G. Cowan, _Statistical Data Analysis_, Oxford University Press, 1998
- \[2\] G. Cowan, "Goodness of fit and Wilk's theorem", Notes, 2013
| github_jupyter |
<a href="https://colab.research.google.com/github/nnuncert/nnuncert/blob/master/notebooks/0_uci_calibration.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Clone repo and install requirements
```
# clone repo
!git clone https://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@github.com/nnuncert/nnuncert
# switch folder and install requirements
%cd nnuncert
!pip install -r requirements.txt
```
# Main imports
```
# %cd nnuncert
import os
import numpy as np
import scipy.stats as spstats
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
import seaborn as sns
from tqdm import tqdm
import hashlib
import nnuncert
from nnuncert.app.uci import RunType, Mode, UCIRun, UCIResults, get_uci_path
from nnuncert.models import make_model, DNNCModel, PNN, type2name
from nnuncert.app import load_uci
from nnuncert.utils.dist import Dist
DATASETS_0 = ["boston", "concrete", "energy", "kin8nm", "powerplant", "wine", "yacht"] #@param
MODELS_0 = ["MC dropout", "PBP", "PNN", "PNN-E", "NLM", "NLM-E", "DNNC-R"] #@param
legend_handler = {
"MC dropout": "MC dropout",
"MC dropout 200": "MC dropout",
"MC dropout 400": "MC dropout",
"PBP": "PBP",
"PNN": "PNN",
"PNN-E": "PNN-E",
"NLM": "NLM",
"NLM-E": "NLM-E",
"DNNC-R": "DNNC-R",
}
name2type_fitkw = {
'MC dropout' : ('MC dropout', {}),
'MC dropout 200' : ('MC dropout', {"conv_factor" : 5}),
'MC dropout 400' : ('MC dropout', {"conv_factor" : 10}),
'PBP' : ('PBP', {}),
'PNN' : ('PNN', {}),
'PNN-E' : ('PNN-E', {}),
'NLM' : ('NLM', {}),
'NLM-E' : ('NLM-E', {}),
'DNNC-R' : ('DNNC-R', {}),
'DNNC-R-STDNN' : ('DNNC-R', {"fit_z_train": False}),
}
```
# Make calibration models
```
# general settings
arch = [[50, "relu", 0]] #@param
epochs = 40 #@param
verbose = 0 #@param
learning_rate = 0.01 #@param
RNG = 42
# set models and some flags
DATASETS = ["boston", "concrete", "energy", "kin8nm", "powerplant", "wine", "yacht"] #@param
MODELS = ["MC dropout", "PBP", "PNN", "PNN-E", "NLM", "NLM-E", "DNNC-R"] #@param
run_type = RunType.CALIBRATION
# keep track of all models
data_track = {}
for dataset in DATASETS:
model_and_pred = {}
# load uci dataset
uci = load_uci(dataset)
uci.get_data("data/uci")
uci.prepare_run()
# "make splits" -> just shuffles train_id
train_id, test_id = run_type.make_splits(uci, rng=RNG)[0]
s = uci.make_train_test_split(train_id=train_id, test_id=test_id)
input_shape = s.x_train.shape[1]
for mod in MODELS:
# init model
mtype, fit_kw = name2type_fitkw[mod]
model = make_model(mtype, input_shape, arch)
# compile model (loss automatically set in model)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# fit NN
if isinstance(model, DNNCModel):
dist = Dist._from_values(s.y_train, method=uci.dist_method,
**uci.dist_kwargs)
fit_kw["dist"] = dist
model.fit(s.x_train, s.y_train, epochs=epochs, verbose=verbose, **fit_kw)
# make predictions
pred_train = model.make_prediction(s.x_train)
# make marginals
y0 = np.linspace(min(uci.data.y), max(uci.data.y), 100)
pred_train.marginals(y0)
# store
model_and_pred[mod] = (model, pred_train)
data_track[dataset] = model_and_pred
```
# Or: load models from repo
This might take a while as the prediction objects for datasets with 10,000 samples require some time.
```
# settings
RNG = 42 # should be 42 if loading from repo
arch = [[50, "relu", 0]] # do not change
learning_rate = 0.01 # do not change
# set models and some flags
DATASETS = ["boston", "concrete", "energy", "kin8nm", "powerplant", "wine", "yacht"] #@param
MODELS = ["MC dropout 400", "PBP", "PNN", "PNN-E", "NLM", "NLM-E", "DNNC-R"] #@param
run_type = RunType.CALIBRATION
data_track = {}
for dataset in DATASETS:
model_and_pred = {}
# load uci dataset
uci = load_uci(dataset)
uci.get_data("data/uci")
uci.prepare_run()
# "make splits" -> just shuffles train_id
train_id, test_id = run_type.make_splits(uci, ratio=0, max_splits=1, rng=RNG)[0]
s = uci.make_train_test_split(train_id=train_id, test_id=test_id)
input_shape = s.x_train.shape[1]
for mod in MODELS:
load_dir = get_uci_path(os.path.join("results", "uci"), dataset, mod, 0, run_type)
# init model
mtype, fit_kw = name2type_fitkw[mod]
arch = [[50, "relu", 0]]
model = make_model(mtype, input_shape, arch)
# compile model (loss automatically set in model)
model.compile(optimizer=tf.keras.optimizers.Adam(learning_rate=learning_rate), metrics=["mae", "mse"])
# load model by giving load_dir as parameter to fit method
if isinstance(model, DNNCModel):
fit_kw["dist"] = None
model.fit(s.x_train, s.y_train, epochs=0, verbose=1, path=load_dir, **fit_kw)
# make prediction and load marginals
pred = model.make_prediction(s.x_train)
pred._marginals = np.load(os.path.join(load_dir, "marginals.npy"))
model_and_pred[mod] = (model, pred)
data_track[dataset] = model_and_pred
```
# Plot
```
# plot helpers for both plots:
from matplotlib.cm import get_cmap
MODEL_COLORS = {
"MC Dropout" : get_cmap("tab20").colors[0], # blue
"MC dropout" : get_cmap("tab20").colors[0], # blue
"MC dropout 200" : get_cmap("tab20").colors[0], # blue
"MC dropout 400" : get_cmap("tab20").colors[0], # blue
'PBP': "#52b2bf", # turquoise
'PNN' : "#d8b709", # yellow
'PNN-E' : "#d87409", # orange
'NLM' : get_cmap("tab20").colors[5], # light green
'NLM-E' : get_cmap("tab20").colors[4], # medium green
'DNNC-R' : "#972d14", # winered
'DNNC-R-STDNN' : get_cmap("tab20").colors[0], # blue
}
rowcol = {0: (0, 0, 2),
1: (0, 2, 4),
2: (0, 4, 6),
3: (0, 6, 8),
4: (1, 1, 3),
5: (1, 3, 5),
6: (1, 5, 7),}
```
## Marginal calibration
```
# some plot helpers
log_axis = {'boston': False,
'concrete': False,
'energy': False,
'kin8nm': False,
'powerplant': False,
'wine': False,
'yacht' : True,
}
bins = {'boston': 50,
'concrete': 50,
'energy': 30,
'kin8nm': 50,
'powerplant': 50,
'wine': 13,
'yacht' : 20,
}
from google.colab import files
from matplotlib import gridspec
from matplotlib.patches import Patch
from matplotlib.lines import Line2D
from nnuncert.app.uci import WineQualityRed
from nnuncert.utils.plotting import _pre_handle, _post_handle
LINEWIDTH_MARGINALS = 1.5
MODELS = ["MC dropout 400", "PBP", "PNN", "PNN-E", "NLM", "NLM-E", "DNNC-R"] #@param
# plot settings
fig = plt.figure(figsize=(14, 6))
height_leg = 0.04
height_row = (1 - height_leg) / 2
gs = gridspec.GridSpec(3, 8, height_ratios=[height_row, height_row, height_leg])
def marginal_plot(d_ssv, uci, preds, models, color_dict, bins=40, noise=None, ax=None, title=None, save_as=None):
fig, ax, return_flag = _pre_handle(ax)
y = uci.data.y
y0 = np.linspace(min(y), max(y), 100)
bar_kw = {"alpha": 1,
"color": "darkgray"}
if isinstance(uci, WineQualityRed):
v = uci.data.y.value_counts(normalize=True)
ax.bar(v.index.array, v.values*2.1, width=0.3, **bar_kw)
else:
# ax.hist(y, bins=bins, density=True, alpha=0.5, color="grey", edgecolor='grey', linewidth=0.1)
ax.hist(y, bins=bins, density=True, **bar_kw)
for m in models:
ax.plot(y0, preds[m][1].marginals(y0, recalc=False), lw=LINEWIDTH_MARGINALS, color=color_dict[m], label=m)
ax.plot(y0, d_ssv.pdf(y0), "--", lw=LINEWIDTH_MARGINALS, color="black")
return _post_handle(fig, ax, return_flag, save_as, title)
axs = []
for i, d in enumerate(DATASETS_0):
# load uci and make empirical dist
uci = load_uci(d)
uci.get_data("data/uci")
uci.prepare_run()
y = uci.data.y
y0 = np.linspace(min(y), max(y), 100)
dist = Dist._from_values(y, method=uci.dist_method, **uci.dist_kwargs)
# load prediction object for all models
preds = data_track[d]
with plt.style.context("ggplot"), plt.rc_context({'xtick.color':'black', 'ytick.color':'black'}):
# with plt.rc_context({'xtick.color':'black', 'ytick.color':'black'}):
# get axis, make plot
(r, c1, c2) = rowcol[i]
ax = plt.subplot(gs[r, c1:c2])
ax.grid(True, linestyle='-.', lw=0.5)
marginal_plot(dist, uci, preds, MODELS, MODEL_COLORS, title=d, ax=ax, bins=bins[d])
# set labels
ax.set_xlabel(r'$y$', color="black")
ax.set_ylabel(r'$p(y| \mathbf{x})$', color="black")
# scale yacht
if log_axis[d] is True:
ax.set_yscale("log")
ax.get_yaxis().set_tick_params(which='minor', size=0)
ax.get_yaxis().set_tick_params(which='minor', width=0)
ax.set_ylabel(r'$log(p(y| \mathbf{x}))$', color="black")
# set limits
ax.set(xlim=(min(y), max(y)))
axs.append(ax)
gs.tight_layout(fig)
fig = plt.gcf()
# make legend elements
legend_elements = [Patch(facecolor="grey", alpha=0.3),
Line2D([0], [0], color='black', linestyle="--", lw=3)]
legend_elements.extend([Line2D([0], [0], color=MODEL_COLORS[m], lw=3) for m in MODELS])
# make legend names
names = ["Histogram of response", "Empirical KDE"]
names += MODELS
# make legend
leg = fig.legend(legend_elements, names, ncol=5, loc='lower center', framealpha=1, fontsize='large')
frame = leg.get_frame()
frame.set_linewidth(0)
```
## Probabilistic calibration
```
from matplotlib import gridspec
from matplotlib.lines import Line2D
from google.colab import files
# from nnuncert.app.uci import MODEL_COLORS
MODELS = ["MC dropout 400", "PBP", "PNN", "PNN-E", "NLM", "NLM-E", "DNNC-R"] #@param
# plot settings
fig = plt.figure(figsize=(14, 6))
height_leg = 0.04
height_row = (1 - height_leg) / 2
gs = gridspec.GridSpec(3, 8, height_ratios=[height_row, height_row, height_leg])
axs = []
# grid for quantiles
min_y, max_y = -0.1, 0.1
ps = np.linspace(0, 1, 20)
for i, d in enumerate(DATASETS):
(r, c1, c2) = rowcol[i]
uci = load_uci(d)
uci.get_data("data/uci")
uci.prepare_run()
s = uci.make_train_test_split(ratio=0, rng=42)
preds = data_track[d]
ax = plt.subplot(gs[r, c1:c2])
ax.set_xlim((0, 1))
ax.grid(True, linestyle='-.', lw=0.5)
for m in MODELS:
pred = preds[m][1]
cdf = pred.cdf(s.y_train)
pj = np.array([np.mean(cdf < p) for p in ps])
ax.plot(ps, pj-ps, lw=2, color=MODEL_COLORS[m])
ax.set_title(d)
ax.set_xlabel(r'$p_j$', color="black")
ax.set_ylabel(r'$\tilde{p}_j - p_j$', color="black")
gs.tight_layout(fig)
ax.hlines(0, 0, 1, linestyles="dashed", lw=1.5, color="black")
# get min, max for all plots and set axis
ymin = np.array([a.get_ylim() for a in fig.axes]).min()
ymax = np.array([a.get_ylim() for a in fig.axes]).max()
for ax in fig.axes:
ax.set_ylim((ymin, ymax))
fig = plt.gcf()
legend_elements = [Line2D([0], [0], color=MODEL_COLORS[m], lw=3) for m in MODELS]
leg = fig.legend(legend_elements, MODELS, ncol=len(MODELS), loc='lower center', framealpha=1, fontsize="large")
frame = leg.get_frame()
frame.set_linewidth(0)
```
| github_jupyter |
# 2-1.2 Intro Python
## Sequence: String
- Accessing String Character with index
- **Accessing sub-strings with index slicing**
- Iterating through Characters of a String
- More String Methods
-----
><font size="5" color="#00A0B2" face="verdana"> <B>Student will be able to</B></font>
- Work with String Characters
- **Slice strings into substrings**
- Iterate through String Characters
- Use String Methods
#
<font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
## Accessing sub-strings
[]( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/251ad8c1-588b-47de-8638-a5bcd0f29800/Unit2_Section1.2a-Index_Slicing-Substrings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/251ad8c1-588b-47de-8638-a5bcd0f29800/Unit2_Section1.2a-Index_Slicing-Substrings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
### Index Slicing [start:stop]
String slicing returns a string section by addressing the start and stop indexes
```python
# assign string to student_name
student_name = "Colette"
# addressing the 3rd, 4th and 5th characters
student_name[2:5]
```
The slice starts at index 2 and ends at index 5 (but does not include index 5)
#
<font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
```
# [ ] review and run example
# assign string to student_name
student_name = "Colette"
# addressing the 3rd, 4th and 5th characters using a slice
print("slice student_name[2:5]:",student_name[2:5])
# [ ] review and run example
# assign string to student_name
student_name = "Colette"
# addressing the 3rd, 4th and 5th characters individually
print("index 2, 3 & 4 of student_name:", student_name[2] + student_name[3] + student_name[4])
# [ ] review and run example
long_word = 'Acknowledgement'
print(long_word[2:11])
print(long_word[2:11], "is the 3rd char through the 11th char")
print(long_word[2:11], "is the index 2, \"" + long_word[2] + "\",", "through index 10, \"" + long_word[10] + "\"")
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 1</B></font>
## slice a string
### start & stop index
```
# [ ] slice long_word to print "act" and to print "tic"
long_word = "characteristics"
# [ ] slice long_word to print "sequence"
long_word = "Consequences"
```
#
<font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
## Accessing beginning of sub-strings
[]( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/368b352f-6061-488c-80a4-d75e455f4416/Unit2_Section1.2b-Index_Slicing_Beginnings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/368b352f-6061-488c-80a4-d75e455f4416/Unit2_Section1.2b-Index_Slicing_Beginnings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
### Index Slicing [:stop]
String slicing returns a string section from index 0 by addressing only the stop index
```python
student_name = "Colette"
# addressing the 1st, 2nd & 3rd characters
student_name[:3]
```
**default start for a slice is index 0**
###
<font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
```
# [ ] review and run example
student_name = "Colette"
# addressing the 1st, 2nd & 3rd characters
print(student_name[:3])
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 2</B></font>
```
# [ ] print the first half of the long_word
long_word = "Consequences"
```
#
<font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
## Accessing ending of sub-strings
[]( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/29beb75a-aee7-43df-9569-e9ad22cffac4/Unit2_Section1.2c-Index_Slicing_Endings.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/29beb75a-aee7-43df-9569-e9ad22cffac4/Unit2_Section1.2c-Index_Slicing_Endings.vtt","srclang":"en","kind":"subtitles","label":"english"}])
### Index Slicing [start:]
String slicing returns a string section including by addressing only the start index
```python
student_name = "Colette"
# addressing the 4th, 5th and 6th characters
student_name[3:]
```
**default end index returns up to and including the last string character**
###
<font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
```
# [ ] review and run example
student_name = "Colette"
# 4th, 5th, 6th and 7th characters
student_name[3:]
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 3</B></font>
```
# [ ] print the second half of the long_word
long_word = "Consequences"
```
#
<font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
## accessing sub-strings by step size
[]( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/62c65917-4979-4d26-9a05-09e1ed02cc51/Unit2_Section1.2d-Index_Slicing-Step_Sizes.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/62c65917-4979-4d26-9a05-09e1ed02cc51/Unit2_Section1.2d-Index_Slicing-Step_Sizes.vtt","srclang":"en","kind":"subtitles","label":"english"}])
### Index Slicing [:], [::2]
- **[:]** returns the entire string
- **[::2]** returns the first char and then steps to every other char in the string
- **[1::3]** returns the second char and then steps to every third char in the string
the number **2**, in the print statement below, represents the **step**
```python
print(long_word[::2])
```
###
<font size="6" color="#00A0B2" face="verdana"> <B>Examples</B></font>
```
# [ ] review and run example
student_name = "Colette"
# return all
print(student_name[:])
# [ ] review and run example
student_name = "Colette"
# return every other
print(student_name[::2])
# [ ] review and run example
student_name = "Colette"
# return every third, starting at 2nd character
print(student_name[1::2])
# [ ] review and run example
long_word = "Consequences"
# starting at 2nd char (index 1) to 9th character, return every other character
print(long_word[1:9:2])
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 4</B></font>
```
# [ ] print the 1st and every 3rd letter of long_word
long_word = "Acknowledgement"
# [ ] print every other character of long_word starting at the 3rd character
long_word = "Acknowledgement"
print(long_word[2::2])
```
<font size="6" color="#00A0B2" face="verdana"> <B>Concepts</B></font>
## Accessing sub-strings continued
[]( http://edxinteractivepage.blob.core.windows.net/edxpages/f7cff1a7-5601-48a1-95a6-fd1fdfabd20e.html?details=[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/2e59f526-fadb-434e-822e-afe3732f75df/Unit2_Section1.2e-Index_Slicing-Reverse.ism/manifest","type":"application/vnd.ms-sstr+xml"}],[{"src":"http://jupyternootbookwams.streaming.mediaservices.windows.net/2e59f526-fadb-434e-822e-afe3732f75df/Unit2_Section1.2e-Index_Slicing-Reverse.vtt","srclang":"en","kind":"subtitles","label":"english"}])
### stepping backwards
```python
print(long_word[::-1])
```
use **[::-1]** to reverse a string
###
<font size="6" color="#00A0B2" face="verdana"> <B>Example</B></font>
```
# [ ] review and run example of stepping backwards using [::-1]
long_word = "characteristics"
# make the step increment -1 to step backwards
print(long_word[::-1])
# [ ] review and run example of stepping backwards using [6::-1]
long_word = "characteristics"
# start at the 7th letter backwards to start
print(long_word[6::-1])
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 5</B></font>
use slicing
```
# [ ] reverse long_word
long_word = "stressed"
# [ ] print the first 5 letters of long_word in reverse
long_word = "characteristics"
```
#
<font size="6" color="#B24C00" face="verdana"> <B>Task 6</B></font>
use slicing
```
# [ ] print the first 4 letters of long_word
# [ ] print the first 4 letters of long_word in reverse
# [ ] print the last 4 letters of long_word in reverse
# [ ] print the letters spanning indexes 3 to 6 of long_word in Reverse
long_word = "timeline"
```
[Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| github_jupyter |
# Space missions feature engineering and predictions
Who does not love Space?
This DataSet was scraped from https://nextspaceflight.com/launches/past/?page=1 and includes all the space missions since the beginning of Space Race (1957)

### Importing Libraries
```
import pandas as pd
```
### Importing the dataset
```
df=pd.read_csv('../input/space-missions-cleaned/Space_Missions_Cleaned.csv')
```
## Feature Engineering
When we try to build a model, we cant just pass null values to it. We need to fill those values somehow and feed it.
```
df.isnull().sum()# To find how many values are missing
```
So, 3360 rocket data is missing..
```
df['Rocket'] = df['Rocket'].fillna(df['Rocket'].mean())
# filling out the missing rocket data by the mean of all missing data
# fillna() allows us to fill the missing data
df.isnull().sum()
```
There are no more null data
```
df.head()
```
Next we need to make sure what columns would be ideal for a model to train. For example Detail,Datum has no need to be included in training data.
```
df=df.drop(['Location','Datum','Detail','DateTime','Launch_Site','Month','Count'],axis=1)
# Dropping unnecessary columns
# axis=1 means we are dropping columns, 0 would be for dropping rows
df.head()
```
Another very important thing is that we cant pass string values to a model for training. We have to convert it to some numerical form for a model to understand.
```
df['Status Mission'].value_counts()# Counts of unique values of Status Mission column
```
The thing we intend to predict here is whether the mission will fail or not. So we have to reduce four unique values into two unique values.
```
df['Status Mission'] =df['Status Mission'].apply(lambda x: x if x == 'Success' else 'Failure')
# converting four unique values namely Success, Failure, Partial Failure and Prelaunch Failure
# into just two values namely Success and Failure
df['Status Mission'].value_counts()
```
Now we have to convert those values into numerical form. The simplest way to do this is make value success 1 and failure 0. LabelEncoder helps us to do just that.
```
from sklearn.preprocessing import LabelEncoder
encoder = LabelEncoder()# creating an object of labelEncoder class
df['Status Mission'] = encoder.fit_transform(df['Status Mission'])# fit_transform() method scales all the data
# and convertes it into 0 and 1.
df[:10]
df['Status Mission'].value_counts()
```
Similiarly, we convert Status Rocket into numerical form
```
encoder = LabelEncoder()
df['Status Rocket']=encoder.fit_transform(df['Status Rocket'])
df.head()
df['Status Rocket'].value_counts()
```
We can predict data for both company and country column, but I decided to drop Country Column.
```
df=df.drop(['Country'],axis=1)
df.head()
```
### One hot encoding the Country Column
```
def onehot_encode(data, column):
dummies = pd.get_dummies(data[column])
data = pd.concat([data, dummies], axis=1)
data.drop(column, axis=1, inplace=True)
return data
df=onehot_encode(df,'Company Name')
```
Segregating the X and y values. What that means is given X data columns, we have to predict y. So, y will only have 1 column and X should not have that column.
```
df.head()
X=df.drop('Status Mission',axis=1)
y=df['Status Mission']
y.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=0.7,random_state=101)
X_train.head()
```
Using StandardScalar to scale the data
```
from sklearn.preprocessing import StandardScaler
scaler=StandardScaler()
scaled_X_train=scaler.fit_transform(X_train)
scaled_X_test=scaler.transform(X_test)
scaled_X_train
```
using Logistic Regression Model for Prediction
```
from sklearn.linear_model import LogisticRegressionCV
log_model=LogisticRegressionCV()
log_model.fit(scaled_X_train,y_train)
y_pred=log_model.predict(scaled_X_test)
y_pred
from sklearn.metrics import accuracy_score,confusion_matrix,classification_report
accuracy_score(y_test,y_pred)
confusion_matrix(y_test,y_pred)
print(classification_report(y_test,y_pred))
```
We see that the acuuracy is 90%, but seeing the classification report we infer that 118 predictions for failure have been given wrong by our model. So this model is not good at all.
```
from sklearn.ensemble import RandomForestClassifier
rfc=RandomForestClassifier(n_estimators=200,max_features='auto',random_state=101)
rfc.fit(scaled_X_train,y_train)
rfc_pred=rfc.predict(scaled_X_test)
accuracy_score(rfc_pred,y_test)
print(confusion_matrix(y_test,rfc_pred))
print(classification_report(y_test,rfc_pred))
```
This model is somewhat better than the logistic regression model but its recall and f1-score is very low which means this is also a poor model.
| github_jupyter |
```
class PointAttribute:
def __init__(s, name, elements, bytes):
s.name = name
s.elements = elements
s.bytes = bytes
class PointAttributes:
POSITION_CARTESIAN = PointAttribute("POSITION_CARTESIAN", 3, 12)
POSITION_PROJECTED_PROFILE = PointAttribute("POSITION_PROJECTED_PROFILE", 2, 8)
COLOR_PACKED = PointAttribute("COLOR_PACKED", 4, 4)
RGB = PointAttribute("RGB", 3, 3)
RGBA = PointAttribute("RGBA", 4, 4)
INTENSITY = PointAttribute("INTENSITY", 1, 2)
CLASSIFICATION = PointAttribute("CLASSIFICATION", 1, 1)
def __init__(s):
s.attributes = []
s.bytes = 0
def add(s, attribute):
s.attributes.append(attribute)
s.bytes = s.bytes + attribute.bytes * attribute.elements
@staticmethod
def fromName(name):
return getattr(PointAttributes, name)
import subprocess
import struct
import json
import sys
import time
exe = "D:/dev/workspaces/CPotree/master/bin/Release_x64/PotreeElevationProfile.exe"
file = "D:/dev/pointclouds/converted/CA13/cloud.js"
coordinates = "{693550.968, 3915914.169},{693890.618, 3916387.819},{694584.820, 3916458.180},{694786.239, 3916307.199}"
width = "14.0"
minLevel = "0"
maxLevel = "4"
attributes = [] # empty list: all available + POSITION_PROJECTED_PROFILE
#attributes = ["--output-attributes", "POSITION_CARTESIAN", POSITION_PROJECTED_PROFILE", "RGB]
start = time.time()
p = subprocess.Popen([exe, file, "--stdout"] + attributes + ["--coordinates", coordinates, "--width", width, "--min-level", minLevel, "--max-level", maxLevel], bufsize=-1, stdout=subprocess.PIPE)
[out, err] = p.communicate()
end = time.time()
print("duration: ", int(1000 * (end - start)), "ms")
headerSize = struct.unpack('i', out[0:4])[0];
header = out[4:4+headerSize].decode("ascii")
buffer = out[4+headerSize:]
print("header:")
print(header)
jHeader = json.loads(header)
numPoints = int(jHeader["points"])
scale = float(jHeader["scale"])
bytesPerPoint = int(jHeader["bytesPerPoint"])
#attributes = jHeader["pointAttributes"]
attributes = []
for attribute in jHeader["pointAttributes"]:
attributes.append(PointAttributes.fromName(attribute))
lx = []
ly = []
lz = []
lpx = []
lpz = []
lc = []
for i in range(numPoints):
byteOffset = bytesPerPoint * i
pbuffer = buffer[byteOffset:byteOffset + bytesPerPoint]
aoffset = 0
for attribute in attributes:
if attribute == PointAttributes.POSITION_CARTESIAN:
ux = struct.unpack('i', pbuffer[aoffset + 0: aoffset + 4])[0]
uy = struct.unpack('i', pbuffer[aoffset + 4: aoffset + 8])[0]
uz = struct.unpack('i', pbuffer[aoffset + 8: aoffset + 12])[0]
x = ux * scale
y = uy * scale
z = uz * scale
lx.append(x)
ly.append(y)
lz.append(z)
elif attribute == PointAttributes.POSITION_PROJECTED_PROFILE:
ux = struct.unpack('i', pbuffer[aoffset + 0: aoffset + 4])[0]
#uy = struct.unpack('i', pbuffer[aoffset + 4: aoffset + 8])[0]
uz = struct.unpack('i', pbuffer[aoffset + 4: aoffset + 8])[0]
x = ux * scale
#y = uy * scale
z = uz * scale
lpx.append(x)
#ly.append(y)
lpz.append(z)
elif attribute == PointAttributes.COLOR_PACKED:
r = pbuffer[aoffset + 0] / 255
g = pbuffer[aoffset + 1] / 255
b = pbuffer[aoffset + 2] / 255
lc.append([r, g, b, 1.0])
elif attribute == PointAttributes.RGB:
r = pbuffer[aoffset + 0] / 255
g = pbuffer[aoffset + 1] / 255
b = pbuffer[aoffset + 2] / 255
lc.append([r, g, b, 1.0])
elif attribute == PointAttributes.INTENSITY:
i = struct.unpack('H', pbuffer[aoffset:aoffset+2])[0] / 255
i = min(i, 1.0)
elif attribute == PointAttributes.CLASSIFICATION:
pass
aoffset = aoffset + attribute.bytes
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
N = len(lx)
x = lx[:N]
y = ly[:N]
colors = lc[:N]
plt.figure(figsize=(15, 6))
plt.scatter(x, y, s=1, c=colors, edgecolors='none')
plt.show()
%matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
N = len(lpx)
x = lpx[:N]
y = lpz[:N]
colors = lc[:N]
plt.figure(figsize=(15, 6))
plt.axes().set_aspect('equal', 'box')
plt.scatter(x, y, s=1, c=colors, edgecolors='none')
plt.xlim([min(x), max(x)])
plt.ylim([min(y), max(y)])
#plt.ylim(-0.5, 4)
plt.show()
```
| github_jupyter |
### Coding the Bose-Hubbard Hamiltonian with QuSpin
The purpose of this tutorial is to teach the interested user to construct bosonic Hamiltonians using QuSpin. To this end, below we focus on the Bose-Hubbard model (BHM) of a 1d chain. The Hamiltonian is
$$ H = -J\sum_{j=0}^{L-1}(b^\dagger_{j+1}b_j + \mathrm{h.c.})-\mu\sum_{j=0}^{L-1} n_j + \frac{U}{2}\sum_{j=0}^{L-1}n_j(n_j-1)$$
where $J$ is the hopping matrix element, $\mu$ -- the chemical potential, and $U$ -- the interaction strength. We label the lattice sites by $j=0,\dots,L-1$, and use periodic boundary conditions.
First, we load the required packages:
```
from quspin.operators import hamiltonian # Hamiltonians and operators
from quspin.basis import boson_basis_1d # Hilbert space boson basis
import numpy as np # generic math functions
```
Next, we define the model parameters:
```
##### define model parameters #####
L=6 # system size
J=1.0 # hopping
U=np.sqrt(2.0) # interaction
mu=2.71 # chemical potential
```
In order to construct the Hamiltonian of the BHM, we need to construct the bosonic basis. This is done with the help of the constructor `boson_basis_1d`. The first required argument is the chain length `L`. As an optional argument one can also specify the number of bosons in the chain `Nb`. We print the basis using the `print()` function.
```
##### construct Bose-Hubbard Hamiltonian #####
# define boson basis with 3 states per site L bosons in the lattice
basis = boson_basis_1d(L,Nb=L) # full boson basis
print(basis)
```
If needed, we can specify the on-site bosonic Hilbert space dimension, i.e. the number of states per site, using the flag `sps=int`. This can help study larger systems of they are dilute.
```
basis = boson_basis_1d(L,Nb=L,sps=3) # particle-conserving basis, 3 states per site
print(basis)
```
Often times, the model under consideration has underlying symmetries. For instance, translation invariance, parity (reflection symmetry), etc. QuSpin allows the user to construct Hamiltonians in symmetry-reduced subspaces. This is done using optional arguments (flags) passed to the basis constructor.
For instance, if we want to construct the basis in the $k=0$ many-body momentum sector, we do this using the flag `kblock=int`. This specifies the many-body momentum of the state via $k=2\pi/L\times\texttt{kblock}$.
Whenever symmetries are present, the `print()` function returns one representative from which one can obtain all 'missing' states by applying the corresponding symmetry operator. It is important to note that, physically, this representative state stands for the linear combination of vectors in the class, not the state that is displayed by `print(basis)`.
```
basis = boson_basis_1d(L,Nb=L,sps=3,kblock=1) # ... and zero momentum sector
print(basis)
```
Additionally, the BHM features reflection symmetry around the middle of the chain. This symmetry block-diagonalises the Hamiltonian into two blocks, corresponding to the negative and positive eigenvalue of the parity operator. The corresponding flag is `pblock=+1,-1`.
```
basis = boson_basis_1d(L,Nb=L,sps=3,kblock=0,pblock=1) # ... + zero momentum and positive parity
print(basis)
```
Now that we have constructed the basis in the symmetry-reduced Hilbert space, we can construct the Hamiltonian. It will be hepful to cast it in the fllowing form:
$$H= -J\sum_{j=0}^{L-1}(b^\dagger_{j+1}b_j + \mathrm{h.c.})-\left(\mu+\frac{U}{2}\right)\sum_{j=0}^{L-1} n_j + \frac{U}{2}\sum_{j=0}^{L-1}n_jn_j $$
We start by defining the site-coupling lists. Suppose we would like to define the operator $\sum_j \mu_j n_j$. To this, end, we can focus on a single summand first, e.g. $2.71 n_{j=3}$. The information encoded in this operator can be summarised as follows:
* the coupling strength is $\mu_{j=3}=2.71$ (site-coupling lists),
* the operator acts on site $j=3$ (site-coupling lists),
* the operator is the density $n$ (operator-string, static/dynamic lists)
In QuSpin, the first two points are grouped together, defininging a list `[mu_j,j]=[2.71,3]`, while the type of operator we specify a bit later (see parantheses). We call this a site-couling list. Summing over multiple sites then results in a nested list of lists:
```
# define site-coupling lists
hop=[[-J,i,(i+1)%L] for i in range(L)] #PBC
interact=[[0.5*U,i,i] for i in range(L)] # U/2 \sum_j n_j n_j
pot=[[-mu-0.5*U,i] for i in range(L)] # -(\mu + U/2) \sum_j j_n
print(hop)
#print(interact)
#print(pot)
```
The site coupling lists specify the sites on which the operators act, yet we need to tell QuSpin which operators are to act on these (pairs of) sites. Thus, we need the following operator strings which enter the static and dynamic lists used to define the Hamiltonian. Since the BHM is time-independent, we use an empty dynamic list
```
# define static and dynamic lists
static=[['+-',hop],['-+',hop],['n',pot],['nn',interact]]
dynamic=[]
print(static)
```
Building the Hamiltonian with QuSpin is now a one-liner using the `hamiltonian` constructor
```
# build Hamiltonian
H=hamiltonian(static,dynamic,basis=basis,dtype=np.float64)
print(H.todense())
```
when the Hamiltonian is constructed, we see three messages saying that it passes three type of symmetries. QuSpin does checks under the hood on the `static` and `dynamic` lists to determine if they satisfy the requested symmetries in the `basis`. They can be disabled by parsing the following flags to the `hamiltonian` constructor: `check_pcon=False`, `check_symm=False` and `check_herm=False`.
We can now diagonalise `H`, and e.g. calculate the entanglement entropy of the ground state.
```
# calculate eigensystem
E,V=H.eigh()
E_GS,V_GS=H.eigsh(k=2,which='SA',maxiter=1E10) # only GS
print("eigenenergies:", E)
#print("GS energy is %0.3f" %(E_GS[0]))
# calculate entanglement entropy per site of GS
subsystem=[i for i in range(L//2)] # sites contained in subsystem
Sent=basis.ent_entropy(V[:,0],sub_sys_A=subsystem,density=True)['Sent_A']
print("GS entanglement per site is %0.3f" %(Sent))
psi_k=V[:,0]
psi_Fock=basis.get_vec(psi_k)
print(psi_k.shape, psi_Fock.shape)
```
| github_jupyter |
```
import numpy as np
int2binary = {}
binary_dim = 8
largest_number = pow(2,binary_dim)
binary = np.unpackbits(np.array([range(largest_number)],dtype=np.uint8).T,axis=1)
for i in range(largest_number):
int2binary[i] = binary[i]
alpha = 0.1
input_dim = 2
hidden_dim = 8
output_dim = 1
synapse_0 = 2 * np.random.random((hidden_dim,input_dim)) - 1
synapse_1 = 2 * np.random.random((output_dim,hidden_dim)) - 1
synapse_h = 2 * np.random.random((hidden_dim,hidden_dim)) - 1
synapse_0_update = np.zeros_like(synapse_0)
synapse_1_update = np.zeros_like(synapse_1)
synapse_h_update = np.zeros_like(synapse_h)
def sigmoid(x):
return 1/(1 + np.exp(-x))
def sigmoid_output_to_derivative(output):
return output*(1-output)
for j in range (5000):
a_int = np.random.randint(largest_number / 2)
a = int2binary[a_int]
b_int = np.random.randint(largest_number / 2)
b = int2binary[b_int]
c_int = a_int + b_int
c = int2binary[c_int]
d = np.zeros_like(c)
overallError = 0
layer_2_deltas = list()
layer_1_values = list()
layer_1_values.append(np.zeros(hidden_dim))
for position in range(binary_dim):
X = np.array([[a[binary_dim - position - 1],b[binary_dim - position - 1]]])
Y = np.array([[c[binary_dim - position - 1]]]).T
# layer_1 = sigmoid(np.dot(X,synapse_0) + np.dot(layer_1_values[-1],synapse_h))
layer_1 = sigmoid(np.dot(X, synapse_0.T) + np.dot(layer_1_values[-1], synapse_h))
layer_2 = sigmoid(np.dot(layer_1,synapse_1))
layer_2_error = y - layer_2
layer_2_deltas.append((layer_2_error) * sigmoid_output_to_derivative(layer_2))
overallError += np.abs(layer_2_error[0])
d[binary_dim - position - 1] = np.round(layer_2[0][0])
layer_1_values.append(copy.deepcopy(layer_1))
future_layer_1_delta = np.zeros(hidden_dim)
for position in range(binary_dim):
X = np.array([[a[position],b[position]]])
layer_1 = layer_1_values[-position - 1]
prev_layer_1 = layer_1_values[-position - 2]
layer_2_delta = layer_2_deltas[-position - 1]
# layer_1_delta = (future_layer_1_delta.dot(synapse_h.T) + \layer_2_delta.dot(synapse_1.T)) * sigmoid_output_to_derivative(layer_1)
layer_1_delta = (future_layer_1_delta.dot(synapse_h.T) + layer_2_delta.dot(synapse_1.T)) * sigmoid_output_to_derivative(layer_1)
synapse_1_update += np.atleast_2d(layer_1).T.dot(layer_2_delta)
synapse_h_update += np.atleast_2d(prev_layer_1).T.dot(layer_1_delta)
synapse_0_update += X.T.dot(layer_1_delta)
future_layer_1_delta = layer_1_delta
synapse_0 += synapse_0_update * alpha
synapse_1 += synapse_1_update * alpha
synapse_h += synapse_h_update * alpha
synapse_0_update *= 0
synapse_1_update *= 0
synapse_h_update *= 0
if (j % 50 == 0):
print("Error:" + str(overallError))
print("Pred:" + str(d))
print("True:" + str(c))
out = 0
for index, x in enumerate(reversed(d)):
out += x * pow(2, index)
print(str(a_int) + " + " + str(b_int) + " = " + str(out))
print("------------")
```
| github_jupyter |
<h1 align="center"> TensorFlow Linear Regression </h1>
Basic TensorFlow: Linear Regression Gradient Descent
**if this tutorial doesn't cover what you are looking for, please leave a comment below the youtube video and I will try to cover what you are interested in.**
<h3 align='Left'> Importing Libraries</h3>
```
import pandas as pd
import pandas_datareader.data as web
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
%matplotlib inline
google = web.DataReader('GOOG', data_source = 'google', start = '3/14/2009', end = '4/14/2016')
google.head()
negative = (len(google)/2.0)*-1
positive = (len(google)/2.0)
google['ticks'] = np.arange(negative,positive,1.0)
google.head()
# Parameters
learning_rate = 0.000001
training_epochs = 25
display_step = 1
# Training Data
train_X = google['ticks'].values
train_Y = google['Open'].values
train_Y = train_Y- train_Y.mean()
n_samples = train_X.shape[0]
# tf Graph Input
rng = np.random
X = tf.placeholder("float")
Y = tf.placeholder("float")
# Set model weights
W = tf.Variable(rng.randn(), name="weight")
b = tf.Variable(0.0, name="bias")
# Construct a linear model
pred = tf.add(tf.mul(X, W), b)
# Initializing the variables
init = tf.initialize_all_variables()
# Mean squared error
cost = tf.reduce_sum(tf.pow(pred-Y, 2))/(2*n_samples)
# Gradient descent
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(cost)
# Initializing the variables
init = tf.initialize_all_variables()
cost_array = []
weight_array = []
intercept_array = []
epic_num = []
# Launch the graph
with tf.Session() as sess:
sess.run(init)
# Fit all training data
for epoch in range(training_epochs):
for (x, y) in zip(train_X, train_Y):
sess.run(optimizer, feed_dict={X: x, Y: y})
#Display logs per epoch step
if ((epoch+1) % display_step == 0):
c = sess.run(cost, feed_dict={X: train_X, Y:train_Y})
epic_num.append(epoch+1)
cost_array.append(c)
weight_array.append(sess.run(W))
intercept_array.append(sess.run(b))
print "Epoch:", '%04d' % (epoch+1), "cost=", "{:.9f}".format(c), \
"W=", sess.run(W), "b=", sess.run(b)
print "Optimization Finished!"
training_cost = sess.run(cost, feed_dict={X: train_X, Y: train_Y})
weight = sess.run(W)
intercept = sess.run(b)
print "Training cost=", training_cost, "W=", weight, "b=",intercept, '\n'
#Graphic display
plt.plot(train_X, train_Y, 'ro', label='Original data')
plt.plot(train_X, sess.run(W) * train_X + sess.run(b), label='Fitted line')
plt.show()
plt.plot(epic_num, cost_array)
plt.ylabel('Cost');
plt.title('Cost over Epics');
plt.xlabel('Epics');
```
| github_jupyter |
```
import pandas as pd
import numpy as np
import pickle
import itertools
import xgboost as xgb
from sklearn.svm import LinearSVC
from sklearn.naive_bayes import MultinomialNB
from sklearn.neural_network import MLPClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.feature_selection import SelectFromModel
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.feature_extraction.text import TfidfTransformer, TfidfVectorizer
from sklearn.metrics import confusion_matrix, f1_score, accuracy_score, precision_score, recall_score
import matplotlib
from sklearn.linear_model import LogisticRegression
from sklearn.neural_network import MLPClassifier
from sklearn.decomposition import PCA
from sklearn.manifold import TSNE
from sklearn.preprocessing import MinMaxScaler, OneHotEncoder
from sklearn.utils.class_weight import compute_class_weight
from IPython.display import display
import matplotlib.pyplot as plt
import seaborn as sns
x_train_text = pd.read_csv('data/t2e/text_train.csv')
x_test_text = pd.read_csv('data/t2e/text_test.csv')
y_train_text = x_train_text['label']
y_test_text = x_test_text['label']
x_train_audio = pd.read_csv('data/s2e/audio_train.csv')
x_test_audio = pd.read_csv('data/s2e/audio_test.csv')
y_train_audio = x_train_audio['label']
y_test_audio = x_test_audio['label']
y_train = y_train_audio # since y_train_audio == y_train_text
y_test = y_test_audio # since y_train_audio == y_train_text
print(x_train_text.shape, y_train_text.shape, x_train_audio.shape, y_train_audio.shape)
emotion_dict = {'ang': 0,
'hap': 1,
'sad': 2,
'fea': 3,
'sur': 4,
'neu': 5}
emo_keys = list(['ang', 'hap', 'sad', 'fea', 'sur', 'neu'])
id_to_emotion = {0: 'ang', 1: 'hap', 2: 'sad', 3: 'fea', 4: 'sur', 5: 'neu'}
def plot_confusion_matrix(cm, classes,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
# plt.figure(figsize=(8,8))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
print("Normalized confusion matrix")
else:
print('Confusion matrix, without normalization')
print(cm)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
def one_hot_encoder(true_labels, num_records, num_classes):
temp = np.array(true_labels[:num_records])
true_labels = np.zeros((num_records, num_classes))
true_labels[np.arange(num_records), temp] = 1
return true_labels
def display_results(y_test, pred_probs, cm=True):
pred = np.argmax(pred_probs, axis=-1)
one_hot_true = one_hot_encoder(y_test, len(pred), len(emotion_dict))
print('Test Set Accuracy = {0:.3f}'.format(accuracy_score(y_test, pred)))
print('Test Set F-score = {0:.3f}'.format(f1_score(y_test, pred, average='macro')))
print('Test Set Precision = {0:.3f}'.format(precision_score(y_test, pred, average='macro')))
print('Test Set Recall = {0:.3f}'.format(recall_score(y_test, pred, average='macro')))
if cm:
plot_confusion_matrix(confusion_matrix(y_test, pred), classes=emo_keys)
cl_weight = dict(pd.Series(x_train_audio['label']).value_counts(normalize=True))
```
## Get Text Features
```
tfidf = TfidfVectorizer(sublinear_tf=True, min_df=5, norm='l2', encoding='latin-1', ngram_range=(1, 2), stop_words='english')
features_text = tfidf.fit_transform(x_train_text.append(x_test_text).transcription).toarray()
x_train_text = features_text[:x_train_text.shape[0]]
x_test_text = features_text[-x_test_text.shape[0]:]
print(features_text.shape, x_train_text.shape, x_test_text.shape)
```
## Combine Text + Audio Features
```
combined_x_train = np.concatenate((np.array(x_train_audio[x_train_audio.columns[2:]]), x_train_text), axis=1)
combined_x_test = np.concatenate((np.array(x_test_audio[x_test_audio.columns[2:]]), x_test_text), axis=1)
print(combined_x_train.shape, combined_x_test.shape)
combined_features_dict = {}
combined_features_dict['x_train'] = combined_x_train
combined_features_dict['x_test'] = combined_x_test
combined_features_dict['y_train'] = np.array(y_train)
combined_features_dict['y_test'] = np.array(y_test)
with open('data/combined/combined_features.pkl', 'wb') as f:
pickle.dump(combined_features_dict, f)
rf_classifier = RandomForestClassifier(n_estimators=600, min_samples_split=25)
rf_classifier.fit(combined_x_train, y_train)
# Predict
pred_probs = rf_classifier.predict_proba(combined_x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/combined_rf_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
with open('trained_models/combined/RF.pkl', 'wb') as f:
pickle.dump(rf_classifier, f)
xgb_classifier = xgb.XGBClassifier(max_depth=7, learning_rate=0.008, objective='multi:softprob',
n_estimators=600, sub_sample=0.8, num_class=len(emotion_dict),
booster='gbtree', n_jobs=4)
xgb_classifier.fit(combined_x_train, y_train)
# Predict
pred_probs = xgb_classifier.predict_proba(combined_x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/combined_xgb_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
with open('trained_models/combined/XGB.pkl', 'wb') as f:
pickle.dump(xgb_classifier, f)
svc_classifier = LinearSVC()
svc_classifier.fit(combined_x_train, y_train)
# Predict
pred = svc_classifier.predict(combined_x_test)
# Results
one_hot_true = one_hot_encoder(y_test, len(pred), len(emotion_dict))
print('Test Set Accuracy = {0:.3f}'.format(accuracy_score(y_test, pred)))
print('Test Set F-score = {0:.3f}'.format(f1_score(y_test, pred, average='macro')))
print('Test Set Precision = {0:.3f}'.format(precision_score(y_test, pred, average='macro')))
print('Test Set Recall = {0:.3f}'.format(recall_score(y_test, pred, average='macro')))
plot_confusion_matrix(confusion_matrix(y_test, pred), classes=emo_keys)
(y_test, pred_probs)
with open('pred_probas/combined_svc_classifier_model.pkl', 'wb') as f:
pickle.dump(svc_classifier, f)
with open('trained_models/combined/SVC.pkl', 'wb') as f:
pickle.dump(svc_classifier, f)
mnb_classifier = MultinomialNB()
mnb_classifier.fit(combined_x_train, y_train)
# Predict
pred_probs = mnb_classifier.predict_proba(combined_x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/combined_mnb_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
with open('trained_models/combined/MNB.pkl', 'wb') as f:
pickle.dump(mnb_classifier, f)
mlp_classifier = MLPClassifier(hidden_layer_sizes=(500, ), activation='relu', solver='adam', alpha=0.0001,
batch_size='auto', learning_rate='adaptive', learning_rate_init=0.01,
power_t=0.5, max_iter=1000, shuffle=True, random_state=None, tol=0.0001,
verbose=False, warm_start=True, momentum=0.8, nesterovs_momentum=True,
early_stopping=False, validation_fraction=0.1, beta_1=0.9, beta_2=0.999,
epsilon=1e-08)
mlp_classifier.fit(combined_x_train, y_train)
# Predict
pred_probs = mlp_classifier.predict_proba(combined_x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/combined_mlp_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
with open('trained_models/combined/MLP.pkl', 'wb') as f:
pickle.dump(mlp_classifier, f)
lr_classifier = LogisticRegression(solver='lbfgs', multi_class='multinomial', max_iter=1000)
lr_classifier.fit(combined_x_train, y_train)
# Predict
pred_probs = lr_classifier.predict_proba(combined_x_test)
# Results
display_results(y_test, pred_probs)
with open('pred_probas/combined_lr_classifier.pkl', 'wb') as f:
pickle.dump(pred_probs, f)
with open('trained_models/combined/LR.pkl', 'wb') as f:
pickle.dump(lr_classifier, f)
ax = xgb.plot_importance(xgb_classifier, max_num_features=10, height=0.5, show_values=False)
fig = ax.figure
fig.set_size_inches(8, 8)
contribution_scores = xgb_classifier.feature_importances_
print(contribution_scores)
# Load predicted probabilities
with open('pred_probas/combined_rf_classifier.pkl', 'rb') as f:
rf_pred_probs = pickle.load(f)
with open('pred_probas/combined_xgb_classifier.pkl', 'rb') as f:
xgb_pred_probs = pickle.load(f)
with open('pred_probas/combined_svc_classifier_model.pkl', 'rb') as f:
svc_preds = pickle.load(f)
with open('pred_probas/combined_mnb_classifier.pkl', 'rb') as f:
mnb_pred_probs = pickle.load(f)
with open('pred_probas/combined_mlp_classifier.pkl', 'rb') as f:
mlp_pred_probs = pickle.load(f)
with open('pred_probas/combined_lr_classifier.pkl', 'rb') as f:
lr_pred_probs = pickle.load(f)
with open('pred_probas/combined_lstm_classifier.pkl', 'rb') as f:
lstm_pred_probs = pickle.load(f)
# Average of the predicted probabilites
ensemble_pred_probs = (xgb_pred_probs +
mlp_pred_probs +
rf_pred_probs +
mnb_pred_probs +
lr_pred_probs)/5.0
# Show metrics
display_results(y_test, ensemble_pred_probs)
```
| github_jupyter |
##### Copyright 2019 The TensorFlow Authors.
```
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#@title MIT License
#
# Copyright (c) 2017 François Chollet
#
# Permission is hereby granted, free of charge, to any person obtaining a
# copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
```
# 保存和恢复模型
<table class="tfo-notebook-buttons" align="left">
<td>
<a target="_blank" href="https://tensorflow.google.cn/beta/tutorials/keras/save_and_restore_models"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 tensorflow.google.cn 上查看</a>
</td>
<td>
<a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/zh-cn/beta/tutorials/keras/save_and_restore_models.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 运行</a>
</td>
<td>
<a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/zh-cn/beta/tutorials/keras/save_and_restore_models.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 Github 上查看源代码</a>
</td>
<td>
<a href="https://storage.googleapis.com/tensorflow_docs/docs/site/zh-cn/beta/tutorials/keras/save_and_restore_models.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
</td>
</table>
Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
[官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
[tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
[docs-zh-cn@tensorflow.org Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
模型可以在训练期间和训练完成后进行保存。这意味着模型可以从任意中断中恢复,并避免耗费比较长的时间在训练上。保存也意味着您可以共享您的模型,而其他人可以通过您的模型来重新创建工作。在发布研究模型和技术时,大多数机器学习从业者分享:
* 用于创建模型的代码
* 模型训练的权重 (weight) 和参数 (parameters) 。
共享数据有助于其他人了解模型的工作原理,并使用新数据自行尝试。
注意:小心不受信任的代码——Tensorflow 模型是代码。有关详细信息,请参阅 [安全使用Tensorflow](https://github.com/tensorflow/tensorflow/blob/master/SECURITY.md)。
### 选项
保存 Tensorflow 的模型有许多方法——具体取决于您使用的 API。本指南使用 [tf.keras](https://tensorflow.google.cn/guide/keras), 一个高级 API 用于在 Tensorflow 中构建和训练模型。有关其他方法的实现,请参阅 TensorFlow [保存和恢复](https://tensorflow.google.cn/guide/saved_model)指南或[保存到 eager](https://tensorflow.google.cn/guide/eager#object-based_saving)。
## 配置
### 安装并导入
安装并导入Tensorflow和依赖项:
```
try:
# Colab only
%tensorflow_version 2.x
except Exception:
pass
!pip install pyyaml h5py # 需要以 HDF5 格式保存模型
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import tensorflow as tf
from tensorflow import keras
print(tf.version.VERSION)
```
### 获取示例数据集
要演示如何保存和加载权重,您将使用 [MNIST 数据集](http://yann.lecun.com/exdb/mnist/). 要加快运行速度,请使用前1000个示例:
```
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
train_labels = train_labels[:1000]
test_labels = test_labels[:1000]
train_images = train_images[:1000].reshape(-1, 28 * 28) / 255.0
test_images = test_images[:1000].reshape(-1, 28 * 28) / 255.0
```
### 定义模型
首先构建一个简单的序列(sequential)模型:
```
# 定义一个简单的序列模型
def create_model():
model = tf.keras.models.Sequential([
keras.layers.Dense(512, activation='relu', input_shape=(784,)),
keras.layers.Dropout(0.2),
keras.layers.Dense(10, activation='softmax')
])
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
return model
# 创建一个基本的模型实例
model = create_model()
# 显示模型的结构
model.summary()
```
## 在训练期间保存模型(以 checkpoints 形式保存)
您可以使用训练好的模型而无需从头开始重新训练,或在您打断的地方开始训练,以防止训练过程没有保存。 `tf.keras.callbacks.ModelCheckpoint` 允许在训练的*过程中*和*结束时*回调保存的模型。
### Checkpoint 回调用法
创建一个只在训练期间保存权重的 `tf.keras.callbacks.ModelCheckpoint` 回调:
```
checkpoint_path = "training_1/cp.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
# 创建一个保存模型权重的回调
cp_callback = tf.keras.callbacks.ModelCheckpoint(filepath=checkpoint_path,
save_weights_only=True,
verbose=1)
# 使用新的回调训练模型
model.fit(train_images,
train_labels,
epochs=10,
validation_data=(test_images,test_labels),
callbacks=[cp_callback]) # 通过回调训练
# 这可能会生成与保存优化程序状态相关的警告。
# 这些警告(以及整个笔记本中的类似警告)是防止过时使用,可以忽略。
```
这将创建一个 TensorFlow checkpoint 文件集合,这些文件在每个 epoch 结束时更新:
```
!ls {checkpoint_dir}
```
创建一个新的未经训练的模型。仅恢复模型的权重时,必须具有与原始模型具有相同网络结构的模型。由于模型具有相同的结构,您可以共享权重,尽管它是模型的不同*实例*。
现在重建一个新的未经训练的模型,并在测试集上进行评估。未经训练的模型将在机会水平(chance levels)上执行(准确度约为10%):
```
# 创建一个基本模型实例
model = create_model()
# 评估模型
loss, acc = model.evaluate(test_images, test_labels)
print("Untrained model, accuracy: {:5.2f}%".format(100*acc))
```
然后从 checkpoint 加载权重并重新评估:
```
# 加载权重
model.load_weights(checkpoint_path)
# 重新评估模型
loss,acc = model.evaluate(test_images, test_labels)
print("Restored model, accuracy: {:5.2f}%".format(100*acc))
```
### checkpoint 回调选项
回调提供了几个选项,为 checkpoint 提供唯一名称并调整 checkpoint 频率。
训练一个新模型,每五个 epochs 保存一次唯一命名的 checkpoint :
```
# 在文件名中包含 epoch (使用 `str.format`)
checkpoint_path = "training_2/cp-{epoch:04d}.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
# 创建一个回调,每 5 个 epochs 保存模型的权重
cp_callback = tf.keras.callbacks.ModelCheckpoint(
filepath=checkpoint_path,
verbose=1,
save_weights_only=True,
period=5)
# 创建一个新的模型实例
model = create_model()
# 使用 `checkpoint_path` 格式保存权重
model.save_weights(checkpoint_path.format(epoch=0))
# 使用新的回调*训练*模型
model.fit(train_images,
train_labels,
epochs=50,
callbacks=[cp_callback],
validation_data=(test_images,test_labels),
verbose=0)
```
现在查看生成的 checkpoint 并选择最新的 checkpoint :
```
! ls {checkpoint_dir}
latest = tf.train.latest_checkpoint(checkpoint_dir)
latest
```
注意: 默认的 tensorflow 格式仅保存最近的5个 checkpoint 。
如果要进行测试,请重置模型并加载最新的 checkpoint :
```
# 创建一个新的模型实例
model = create_model()
# 加载以前保存的权重
model.load_weights(latest)
# 重新评估模型
loss, acc = model.evaluate(test_images, test_labels)
print("Restored model, accuracy: {:5.2f}%".format(100*acc))
```
## 这些文件是什么?
上述代码将权重存储到 [checkpoint](https://tensorflow.google.cn/guide/saved_model#save_and_restore_variables)—— 格式化文件的集合中,这些文件仅包含二进制格式的训练权重。 Checkpoints 包含:
* 一个或多个包含模型权重的分片。
* 索引文件,指示哪些权重存储在哪个分片中。
如果你只在一台机器上训练一个模型,你将有一个带有后缀的碎片: `.data-00000-of-00001`
## 手动保存权重
您将了解如何将权重加载到模型中。使用 `Model.save_weights` 方法手动保存它们同样简单。默认情况下, `tf.keras` 和 `save_weights` 特别使用 TensorFlow [checkpoints](../../guide/keras/checkpoints) 格式 `.ckpt` 扩展名和 ( 保存在 [HDF5](https://js.tensorflow.org/tutorials/import-keras.html) 扩展名为 `.h5` [保存并序列化模型](../../guide/keras/saving_and_serializing#weights-only_saving_in_savedmodel_format) ):
```
# 保存权重
model.save_weights('./checkpoints/my_checkpoint')
# 创建模型实例
model = create_model()
# Restore the weights
model.load_weights('./checkpoints/my_checkpoint')
# Evaluate the model
loss,acc = model.evaluate(test_images, test_labels)
print("Restored model, accuracy: {:5.2f}%".format(100*acc))
```
## 保存整个模型
模型和优化器可以保存到包含其状态(权重和变量)和模型参数的文件中。这可以让您导出模型,以便在不访问原始 python 代码的情况下使用它。而且您可以通过恢复优化器状态的方式,从中断的位置恢复训练。
保存完整模型会非常有用——您可以在 TensorFlow.js ([HDF5](https://js.tensorflow.org/tutorials/import-keras.html), [Saved Model](https://js.tensorflow.org/tutorials/import-saved-model.html)) 加载他们,然后在 web 浏览器中训练和运行它们,或者使用 TensorFlow Lite 将它们转换为在移动设备上运行([HDF5](https://tensorflow.google.cn/lite/convert/python_api#exporting_a_tfkeras_file_), [Saved Model](https://tensorflow.google.cn/lite/convert/python_api#exporting_a_savedmodel_))
### 将模型保存为HDF5文件
Keras 可以使用 [HDF5](https://en.wikipedia.org/wiki/Hierarchical_Data_Format) 标准提供基本保存格式。出于我们的目的,可以将保存的模型视为单个二进制blob:
```
# 创建一个新的模型实例
model = create_model()
# 训练模型
model.fit(train_images, train_labels, epochs=5)
# 将整个模型保存为HDF5文件
model.save('my_model.h5')
```
现在,从该文件重新创建模型:
```
# 重新创建完全相同的模型,包括其权重和优化程序
new_model = keras.models.load_model('my_model.h5')
# 显示网络结构
new_model.summary()
```
检查其准确率(accuracy):
```
loss, acc = new_model.evaluate(test_images, test_labels)
print("Restored model, accuracy: {:5.2f}%".format(100*acc))
```
这项技术可以保存一切:
* 权重
* 模型配置(结构)
* 优化器配置
Keras 通过检查网络结构来保存模型。目前,它无法保存 Tensorflow 优化器(调用自 `tf.train`)。使用这些的时候,您需要在加载后重新编译模型,否则您将失去优化器的状态。
### 通过 `saved_model` 保存
注意:这种保存 `tf.keras` 模型的方法是实验性的,在将来的版本中可能有所改变。
建立一个新模型,然后训练它:
```
model = create_model()
model.fit(train_images, train_labels, epochs=5)
```
创建一个 `saved_model`,并将其放在带有 `tf.keras.experimental.export_saved_model` 的带时间戳的目录中:
```
import time
saved_model_path = "./saved_models/{}".format(int(time.time()))
tf.keras.experimental.export_saved_model(model, saved_model_path)
saved_model_path
```
列出您保存的模型:
```
!ls saved_models/
```
从保存的模型重新加载新的 Keras 模型:
```
new_model = tf.keras.experimental.load_from_saved_model(saved_model_path)
# 显示网络结构
new_model.summary()
```
使用恢复的模型运行预测:
```
model.predict(test_images).shape
# 必须在评估之前编译模型。
# 如果仅部署已保存的模型,则不需要此步骤。
new_model.compile(optimizer=model.optimizer, # 保留已加载的优化程序
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# 评估已恢复的模型
loss, acc = new_model.evaluate(test_images, test_labels)
print("Restored model, accuracy: {:5.2f}%".format(100*acc))
```
| github_jupyter |
```
# Python program for reversal algorithm of array rotation
# Function to reverse arr[] from index start to end
def rverseArray(arr, start, end):
while (start < end):
temp = arr[start]
arr[start] = arr[end]
arr[end] = temp
start += 1
end = end-1
# Function to left rotate arr[] of size n by d
def leftRotate(arr, d):
n = len(arr)
rverseArray(arr, 0, d-1)
rverseArray(arr, d, n-1)
rverseArray(arr, 0, n-1)
# Function to print an array
def printArray(arr):
for i in range(0, len(arr)):
print (arr[i])
# Driver function to test above functions
arr = [1, 2, 3, 4, 5, 6, 7]
leftRotate(arr, 2) # Rotate array by 2
printArray(arr)
# Check if given array is Monotonic
def isMonotonic(A):
return (all(A[i] <= A[i + 1] for i in range(len(A) - 1)) or
all(A[i] >= A[i + 1] for i in range(len(A) - 1)))
# Driver program
A = [6, 5, 4, 4]
# Print required result
print(isMonotonic(A))
# Reversing a list using reversed()
def Reverse(lst):
return [ele for ele in reversed(lst)]
# Driver Code
lst = [10, 11, 12, 13, 14, 15]
print(Reverse(lst))
# Reversing a list using reverse()
def Reverse(lst):
lst.reverse()
return lst
lst = [10, 11, 12, 13, 14, 15]
print(Reverse(lst))
# Reversing a list using slicing technique
def Reverse(lst):
new_lst = lst[::-1]
return new_lst
lst = [10, 11, 12, 13, 14, 15]
print(Reverse(lst))
# Using the in-built function list()
def Cloning(li1):
li_copy = list(li1)
return li_copy
# Driver Code
li1 = [4, 8, 2, 10, 15, 18]
li2 = Cloning(li1)
print("Original List:", li1)
print("After Cloning:", li2)
# Using append()
def Cloning(li1):
li_copy =[]
for item in li1: li_copy.append(item)
return li_copy
# Driver Code
li1 = [4, 8, 2, 10, 15, 18]
li2 = Cloning(li1)
print("Original List:", li1)
print("After Cloning:", li2)
# Using bilt-in method copy()
def Cloning(li1):
li_copy =[]
li_copy = li1.copy()
return li_copy
# Driver Code
li1 = [4, 8, 2, 10, 15, 18]
li2 = Cloning(li1)
print("Original List:", li1)
print("After Cloning:", li2)
# Python program to remove empty tuples from a
# list of tuples function to remove empty tuples
# using list comprehension
def Remove(tuples):
tuples = [t for t in tuples if t]
return tuples
# Driver Code
tuples = [(), ('ram','15','8'), (), ('laxman', 'sita'),
('krishna', 'akbar', '45'), ('',''),()]
print(Remove(tuples))
# Python2 program to remove empty tuples
# from a list of tuples function to remove
# empty tuples using filter
def Remove(tuples):
tuples = filter(None, tuples)
return tuples
# Driver Code
tuples = [(), ('ram','15','8'), (), ('laxman', 'sita'),
('krishna', 'akbar', '45'), ('',''),()]
print (Remove(tuples))
# function which return reverse of a string
def reverse(s):
return s[::-1]
def isPalindrome(s):
# Calling reverse function
rev = reverse(s)
# Checking if both string are equal or not
if (s == rev):
return True
return False
# Driver code
s = "malayalam"
ans = isPalindrome(s)
if ans == 1:
print("Yes")
else:
print("No")
# Python program to check
# if a string is palindrome
# or not
x = "malayalam"
w = ""
for i in x:
w = i + w
if (x==w):
print("YES")
from collections import OrderedDict
seq = ('name', 'age', 'gender')
dict = OrderedDict.fromkeys(seq)
# Output = {'age': None, 'name': None, 'gender': None}
print (str(dict))
dict = OrderedDict.fromkeys(seq, 10)
# Output = {'age': 10, 'name': 10, 'gender': 10}
print (str(dict))
# Python program to check if a string
# contains any special character
# import required package
import re
# Function checks if the string
# contains any special character
def run(string):
# Make own character set and pass
# this as argument in compile method
regex = re.compile('[@_!#$%^&*()<>?/\|}{~:]')
# Pass the string in search
# method of regex object.
if(regex.search(string) == None):
print("String is accepted")
else:
print("String is not accepted.")
# Driver Code
if __name__ == '__main__' :
# Enter the string
string = "Rocky$For$Rockstar"
# calling run function
run(string)
# Python3 program to find list of uncommon words
# Function to return all uncommon words
def UncommonWords(A, B):
# count will contain all the word counts
count = {}
# insert words of string A to hash
for word in A.split():
count[word] = count.get(word, 0) + 1
# insert words of string B to hash
for word in B.split():
count[word] = count.get(word, 0) + 1
# return required list of words
return [word for word in count if count[word] == 1]
# Driver Code
A = "R for Rocky"
B = "Learning from R for Rocky"
# Print required answer
print(UncommonWords(A, B))
from collections import Counter
def remov_duplicates(input):
# split input string separated by space
input = input.split(" ")
# joins two adjacent elements in iterable way
for i in range(0, len(input)):
input[i] = "".join(input[i])
# now create dictionary using counter method
# which will have strings as key and their
# frequencies as value
UniqW = Counter(input)
# joins two adjacent elements in iterable way
s = " ".join(UniqW.keys())
print (s)
# Driver program
if __name__ == "__main__":
input = 'Python is great and Java is also great'
remov_duplicates(input)
```
| github_jupyter |
```
# this script is for the calculation of PDFs in Figure 2 of Rodgers et al. 2021 (https://doi.org/10.5194/esd-2021-50).
# If you have have any questions, please contact the author of this notebook.
# Author: Lei Huang (huanglei[AT]pusan[DOT]ac[DOT]kr)
```
# import
```
%matplotlib inline
import numpy as np
import xarray as xr
import matplotlib.pyplot as plt
import matplotlib as mpl
import glob
import dask.array as da
import pandas as pd
```
# seting parallel
```
## Run the mpirun in command line:
## mpirun --np 6 dask-mpi --scheduler-file scheduler.json --no-nanny --dashboard-address :8785 --memory-limit=60e9
from dask.distributed import Client
client = Client(scheduler_file = 'the_path_for_your_scheduler_json_file')
```
# functions for reading ensembles in parallel
```
# preprocess dataset prior to concatenation
variables = []
exceptcv = ['time', 'nlat', 'nlon', 'z_t',
'lon', 'lat', 'gw', 'landfrac', 'area', *variables]
def def_process_coords(exceptcv = []):
def process_coords(ds, except_coord_vars=exceptcv):
coord_vars = []
for v in np.array(ds.coords):
if not v in except_coord_vars:
coord_vars += [v]
for v in np.array(ds.data_vars):
if not v in except_coord_vars:
coord_vars += [v]
return ds.drop(coord_vars)
return process_coords
# define function to read in files for historical simulations
def read_in(var, exceptcv, domain='lnd/', freq='day_1/', stream='h6', chunks=dict(time=365), ens_s = 0, ens_e = 100):
ens_dir = "mother_directory_for_ensemble_files"
projens_names = [member.split('archive/')[1][:-1] for member in sorted(
glob.glob(ens_dir + "b.e21.BSSP370*.f09_g17*/"))][ens_s:ens_e]
proj_ncfiles = []
for i in np.arange(len(projens_names)):
proj_fnames = sorted(glob.glob(
ens_dir + projens_names[i] + "/" + domain + "proc/tseries/" + freq + "*" + stream + var + "*"))
proj_ncfiles.append(proj_fnames[-2:])
ens_numbers = [members.split('LE2-')[1]
for members in projens_names]
proj_ds = xr.open_mfdataset(proj_ncfiles,
chunks=chunks,
preprocess=def_process_coords(exceptcv),
combine='nested',
concat_dim=[[*ens_numbers], 'time'],
parallel=True)
ens_ds = proj_ds.rename({'concat_dim': 'ensemble'})
return ens_ds
```
# PDF for Nino3.4 SST
```
# read in SST for period of 1980-1989
variables = ['SST']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/ocn/proc/tseries/day_1/*.SST.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/ocn/proc/tseries/day_1/*.SST.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
sst_hist_ds = xr.open_mfdataset(ncfiles,
chunks={'time':365},
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
# read in SST for period of 2090-2099
sst_proj_ds = read_in(var = '.SST.',
exceptcv = exceptcv,
domain = 'ocn/',
freq = 'day_1/',
stream = 'h*',
chunks= dict(time = 365))
# select the regions for Nino3
sst_hist_nino = sst_hist_ds.SST[:,:,...].sel(nlat = slice(168,206), nlon = slice(204,249))
sst_proj_nino = sst_proj_ds.SST.sel(nlat = slice(168,206), nlon = slice(204,249), time = slice('2090-01-02','2100-01-01'))
# tarea is the cell area on the T-grid of POP2
tarea_hist_nino = sst_hist_ds.TAREA.sel(nlat = slice(168,206), nlon = slice(204,249)).broadcast_like(sst_hist_nino).chunk({'time':sst_hist_nino.chunks[1]})
tarea_proj_nino = sst_proj_ds.TAREA.sel(nlat = slice(168,206), nlon = slice(204,249),time = slice('2090-01-02','2100-01-01')).broadcast_like(sst_proj_nino).chunk({'time':sst_proj_nino.chunks[1]})
# calculate the PDF for SST in 1980-1989
# please refer to the document of dask.array.histogram for more information
h_hist_sst_nino_raw, bins_hist_sst_nino_raw = da.histogram(sst_hist_nino,
bins = np.arange(15,40.2,0.2),
weights = tarea_hist_nino,
density = True)
h_hist_sst_nino_raw = h_hist_sst_nino_raw.compute()
# calculate the PDF for SST in 2090-2099
h_proj_sst_nino_raw, bins_proj_sst_nino_raw = da.histogram(sst_proj_nino,
bins = np.arange(15,40.2,0.2),
weights = tarea_proj_nino,
density = True)
h_proj_sst_nino_raw = h_proj_sst_nino_raw.compute()
# save the result to csv file
s1 = np.expand_dims(bins_hist_sst_nino_raw[1:]-0.1, axis = 1)
s2 = np.expand_dims(h_hist_sst_nino_raw, axis = 1)
s3 = np.expand_dims(h_proj_sst_nino_raw, axis = 1)
pd.DataFrame(data = np.concatenate((s1,s2,s3), axis = 1),
columns= ['bins', 'h_hist', 'h_proj']).to_csv('path_csv_file', index = False)
```
# Fire counts in California
```
variables = ['NFIRE']
exceptcv = ['time', 'lat', 'lon', 'landfrac', 'area', *variables]
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/lnd/proc/tseries/day_1/*.NFIRE.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/lnd/proc/tseries/day_1/*.NFIRE.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
nfire_hist_ds = xr.open_mfdataset(ncfiles,
chunks={'time':365},
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
nfire_proj_ds = read_in(var = '.NFIRE.',
exceptcv = exceptcv,
domain = 'lnd/',
freq = 'day_1/',
stream = 'h5',
chunks= dict(time = 365),
ens_s = 10,
ens_e = 100)
nfire_hist_calif = nfire_hist_ds.NFIRE.sel(lat = slice(32,41), lon = slice(235,242))*10000*365*24*3600 # convert the unit to the one shown in Figure 2
nfire_proj_calif = nfire_proj_ds.NFIRE.sel(lat = slice(32,41), lon = slice(235,242), time = slice('2090-01-01','2099-12-31'))*10000*365*24*3600
landfrac_hist_calif = nfire_hist_ds.landfrac.sel(lat = slice(32,41), lon = slice(235,242))
landfrac_proj_calif = nfire_proj_ds.landfrac.sel(lat = slice(32,41), lon = slice(235,242), time = slice('2090-01-01','2099-12-31'))
area_hist_calif = nfire_hist_ds.area.sel(lat = slice(32,41), lon = slice(235,242))
area_proj_calif = nfire_proj_ds.area.sel(lat = slice(32,41), lon = slice(235,242), time = slice('2090-01-01','2099-12-31'))
landfrac_hist_calif = landfrac_hist_calif.broadcast_like(nfire_hist_calif).chunk({'time':nfire_hist_calif.chunks[1]})
landfrac_proj_calif = landfrac_proj_calif.broadcast_like(nfire_proj_calif).chunk({'time':nfire_proj_calif.chunks[1]})
area_hist_calif = area_hist_calif.broadcast_like(nfire_hist_calif).chunk({'time':nfire_hist_calif.chunks[1]})
area_proj_calif = area_proj_calif.broadcast_like(nfire_proj_calif).chunk({'time':nfire_proj_calif.chunks[1]})
nfire_hist_calif = nfire_hist_calif.where(landfrac_hist_calif >= 0.9)
nfire_proj_calif = nfire_proj_calif.where(landfrac_proj_calif >= 0.9)
h_hist_nfire_calif_raw, bins_hist_nfire_calif_raw = np.histogram(nfire_hist_calif,
bins = np.arange(0,2500.1,10),
weights = area_hist_calif,
density = True)
h_proj_nfire_calif_raw, bins_proj_nfire_calif_raw = np.histogram(nfire_proj_calif,
bins = np.arange(0,2500.1,10),
weights = area_proj_calif,
density = True)
s1 = np.expand_dims(bins_hist_nfire_calif_raw[1:] - 5, axis = 1)
s2 = np.expand_dims(h_hist_nfire_calif_raw, axis = 1)
s3 = np.expand_dims(h_proj_nfire_calif_raw, axis = 1)
pd.DataFrame(data = np.concatenate((s1,s2,s3), axis = 1),
columns=['bins', 'h_hist', 'h_proj']).to_csv('path_csv_file', index=False)
```
# PDF for Chlorophyll in NA
```
# In the Biogeochemistry module, chlorophyll concentration equals the sum of diatChl_SURF, diazChl_SURF, and spChl_SURF
## read in chlorophyll for 1980-1989
variables = ['diatChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/ocn/proc/tseries/day_1/*.diatChl_SURF.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/ocn/proc/tseries/day_1/*.diatChl_SURF.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
tchl_hist_ds = xr.open_mfdataset(ncfiles,
chunks=dict(nlat = 192, nlon = 160, time = 365),
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/ocn/proc/tseries/day_1/*.diazChl_SURF.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/ocn/proc/tseries/day_1/*.diazChl_SURF.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
variables = ['diazChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
zchl_hist_ds = xr.open_mfdataset(ncfiles,
chunks=dict(nlat = 192, nlon = 160, time = 365),
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/ocn/proc/tseries/day_1/*.spChl_SURF.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/ocn/proc/tseries/day_1/*.spChl_SURF.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
variables = ['spChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
spchl_hist_ds = xr.open_mfdataset(ncfiles,
chunks=dict(nlat = 192, nlon = 160, time = 365),
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
## read in chlorophyll for 2090-2099
variables = ['diatChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
tchl_proj_ds = read_in(var = '.diatChl_SURF.',
exceptcv = exceptcv,
domain = 'ocn/',
freq = 'day_1/',
stream = 'h*',
chunks= dict(nlat = 192, nlon = 160, time = 365),)
variables = ['diazChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
zchl_proj_ds = read_in(var = '.diazChl_SURF.',
exceptcv = exceptcv,
domain = 'ocn/',
freq = 'day_1/',
stream = 'h*',
chunks= dict(nlat = 192, nlon = 160, time = 365),)
variables = ['spChl_SURF']
exceptcv = ['time', 'nlat', 'nlon', 'z_t', 'TAREA', *variables]
spchl_proj_ds = read_in(var = '.spChl_SURF.',
exceptcv = exceptcv,
domain = 'ocn/',
freq = 'day_1/',
stream = 'h*',
chunks= dict(nlat = 192, nlon = 160, time = 365),)
TLAT = xr.open_dataset(ncfiles[-1]).TLAT
TLONG = xr.open_dataset(ncfiles[-1]).TLONG
chl_hist = tchl_hist_ds.diatChl_SURF[:,:,...] \
+ zchl_hist_ds.diazChl_SURF[:,:,...] \
+ spchl_hist_ds.spChl_SURF[:,:,...]
chl_proj = tchl_proj_ds.diatChl_SURF.sel(time = slice('2090-01-02','2100-01-01')) \
+ zchl_proj_ds.diazChl_SURF.sel(time = slice('2090-01-02','2100-01-01')) \
+ spchl_proj_ds.spChl_SURF.sel(time = slice('2090-01-02','2100-01-01'))
tarea_hist = tchl_hist_ds.TAREA.broadcast_like(chl_hist).chunk({'time':chl_hist.chunks[1]})
tarea_proj = tchl_proj_ds.TAREA.sel(time = slice('2090-01-02','2100-01-01')).broadcast_like(chl_proj).chunk({'time':chl_proj.chunks[1]})
chl_hist_NA = chl_hist.where((TLAT>=40) & (TLAT <= 60) & (TLONG >= 300) & (TLONG <= 345), drop = True)
chl_proj_NA = chl_proj.where((TLAT>=40) & (TLAT <= 60) & (TLONG >= 300) & (TLONG <= 345), drop = True)
tarea_hist_NA = tarea_hist.where((TLAT>=40) & (TLAT <= 60) & (TLONG >= 300) & (TLONG <= 345), drop = True)
tarea_proj_NA = tarea_proj.where((TLAT>=40) & (TLAT <= 60) & (TLONG >= 300) & (TLONG <= 345), drop = True)
h_hist_chl_NA_raw, bins_hist_chl_NA_raw = np.histogram(chl_hist_NA,
bins = np.arange(0,20.2,0.2),
weights = tarea_hist_NA,
density=True)
h_proj_chl_NA_raw, bins_proj_chl_NA_raw = np.histogram(chl_proj_NA,
bins = np.arange(0,20.2,0.2),
weights = tarea_proj_NA,
density=True)
s1 = np.expand_dims(bins_hist_chl_NA_raw[1:]-0.1, axis = 1)
s2 = np.expand_dims(h_hist_chl_NA_raw, axis = 1)
s3 = np.expand_dims(h_proj_chl_NA_raw, axis = 1)
pd.DataFrame(data = np.concatenate((s1,s2,s3), axis = 1),
columns=['bins', 'h_hist', 'h_proj']).to_csv('path_csv_file', index=False)
```
# NEP in Amazon
```
variables = ['NEP']
exceptcv = ['time', 'lat', 'lon', 'gw', 'landfrac', 'area', *variables]
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/lnd/proc/tseries/day_1/*.NEP.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/lnd/proc/tseries/day_1/*.NEP.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
nep_hist_ds = xr.open_mfdataset(ncfiles,
chunks={'time':365},
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
nep_proj_ds = read_in(var = '.NEP.',
exceptcv = exceptcv,
domain = 'lnd/',
freq = 'day_1/',
stream = 'h5',
chunks= dict(time = 365),
ens_s = 10,
ens_e = 100)
nep_hist_amazon = nep_hist_ds.NEP.sel(lat = slice(-10,10), lon = slice(280,310))* 1000000
nep_proj_amazon = nep_proj_ds.NEP.sel(lat = slice(-10,10), lon = slice(280,310), time = slice('2090-01-01','2099-12-31'))* 1000000
landfrac_hist_amazon = nep_hist_ds.landfrac.sel(lat = slice(-10,10), lon = slice(280,310))
landfrac_proj_amazon = nep_proj_ds.landfrac.sel(lat = slice(-10,10), lon = slice(280,310), time = slice('2090-01-01','2099-12-31'))
area_hist_amazon = nep_hist_ds.area.sel(lat = slice(-10,10), lon = slice(280,310))
area_proj_amazon = nep_proj_ds.area.sel(lat = slice(-10,10), lon = slice(280,310), time = slice('2090-01-01','2099-12-31'))
area_hist_amazon = area_hist_amazon.broadcast_like(nep_hist_amazon).chunk({'time':nep_hist_amazon.chunks[1]})
area_proj_amazon = area_proj_amazon.broadcast_like(nep_proj_amazon).chunk({'time':nep_proj_amazon.chunks[1]})
nep_hist_amazon = nep_hist_amazon.where(landfrac_hist_amazon[0,...] >= 0.9)
nep_proj_amazon = nep_proj_amazon.where(landfrac_proj_amazon[0,...] >= 0.9)
h_hist_nep_amazon_raw, bins_hist_nep_amazon_raw = np.histogram(nep_hist_amazon,
bins = np.arange(-60,60.06,1),
weights = area_hist_amazon,
density = True)
h_proj_nep_amazon_raw, bins_proj_nep_amazon_raw = np.histogram(nep_proj_amazon,
bins = np.arange(-60,60.06,1),
weights = area_proj_amazon,
density = True)
s1 = np.expand_dims(bins_hist_nep_amazon_raw[1:] - 0.5, axis = 1)
s2 = np.expand_dims(h_hist_nep_amazon_raw, axis = 1)
s3 = np.expand_dims(h_proj_nep_amazon_raw, axis = 1)
pd.DataFrame(data = np.concatenate((s1,s2,s3), axis = 1),
columns=['bins', 'h_hist', 'h_proj']).to_csv('path_csv_file', index=False)
```
# PDF for precipitation in Nino3.4
```
variables = ['PRECT']
exceptcv = ['time', 'lat', 'lon', 'gw', *variables]
ncfiles = sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTcmip6*/atm/proc/tseries/day_1/*.PRECT.1980*')) \
+ sorted(glob.glob('mother_directory_for_ensemble_files/b.e21.BHISTsmbb*/atm/proc/tseries/day_1/*.PRECT.1980*'))
hist_ens_numbers = [member.split('LE2-')[1][:8] for member in ncfiles]
prect_hist_ds = xr.open_mfdataset(ncfiles,
chunks={'time':365},
combine='nested',
preprocess=def_process_coords(exceptcv),
concat_dim =[[*hist_ens_numbers]],
parallel = True).rename({'concat_dim':'ensemble'})
prect_proj_ds = read_in(var = '.PRECT.',
exceptcv = exceptcv,
domain = 'atm/',
freq = 'day_1/',
stream = 'h1',
chunks= dict(time = 365))
prect_hist_nino = prect_hist_ds.PRECT.sel(lat = slice(-5,5), lon = slice(190,240))* 24* 3600* 1000
prect_proj_nino = prect_proj_ds.PRECT.sel(lat = slice(-5,5), lon = slice(190,240), time = slice('2090-01-01','2099-12-31'))* 24* 3600* 1000
h_hist_prect_nino_raw, bins_hist_prect_nino_raw = np.histogram(prect_hist_nino,
bins = np.arange(0,1000.01,4),
weights = gw_hist_nino,
density = True)
h_proj_prect_nino_raw, bins_proj_prect_nino_raw = np.histogram(prect_proj_nino,
bins = np.arange(0,1000.01,4),
weights = gw_proj_nino,
density = True)
s1 = np.expand_dims(bins_hist_prect_nino_raw[1:] - 2, axis = 1)
s2 = np.expand_dims(h_hist_prect_nino_raw, axis = 1)
s3 = np.expand_dims(h_proj_prect_nino_raw, axis = 1)
pd.DataFrame(data = np.concatenate((s1,s2,s3), axis = 1),
columns=['bins', 'h_hist', 'h_proj']).to_csv('path_csv_file', index=False)
```
| github_jupyter |
# **Runtime Dependencies: Must Run First!**
```
import pandas as pd
from pandas.tseries.offsets import MonthEnd
from datetime import datetime
from matplotlib import pyplot as plt
# Statsmodels API - Standard
import statsmodels.api as sm
# Statsmodels API - Formulaic
import statsmodels.formula.api as smf
# ### Bonus: Multiple Outputs Per Cell
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
```
# **Module 9 - Topic 1: Linear Regression with Statsmodels, Part 2**
## **Module 9.1.5: Multivariate Linear Regression**
In the last notebook, I introduced the CAPM theory and regression with the market to calculate the alpha (y intercept) and beta (slope coefficient) of a stock when plotting excess stock returns vs excess market returns.
In this notebook, I'm going to cover running a Fama French 3 Factor regression with extra x's!
I'm starting below with importing the data sets from the first notebook:
```
loc = "https://github.com/mhall-simon/python/blob/main/data/misc/stocks-factors-capm.xlsx?raw=true"
aapl = pd.read_excel(loc, sheet_name="AAPL", index_col=0, parse_dates=True)
aapl['Return'] = (aapl.Close - aapl.Close.shift(1)) / aapl.Close.shift(1)
aapl.dropna(inplace=True)
aapl.index = aapl.index + MonthEnd(1)
R = pd.DataFrame(aapl.Return)
R = R.rename(columns={"Return":"AAPL"})
amzn = pd.read_excel(loc, sheet_name="AMZN", index_col=0, parse_dates=True)
tsla = pd.read_excel(loc, sheet_name="TSLA", index_col=0, parse_dates=True)
amzn['Return'] = (amzn.Close - amzn.Close.shift(1)) / amzn.Close.shift(1)
tsla['Return'] = (tsla.Close - tsla.Close.shift(1)) / tsla.Close.shift(1)
amzn.dropna(inplace=True)
tsla.dropna(inplace=True)
amzn.index = amzn.index + MonthEnd(1)
tsla.index = tsla.index + MonthEnd(1)
R = pd.merge(R, tsla.Return, left_index=True, right_index=True)
R = pd.merge(R, amzn.Return, left_index=True, right_index=True)
R = R.rename(columns={"Return_x":"TSLA","Return_y":"AMZN"})
dp = lambda x: datetime.strptime(x, "%Y%m")
ff = amzn = pd.read_excel(loc, sheet_name="MktRf", index_col=0, parse_dates=True, date_parser=dp, header=3)
ff.index = ff.index + MonthEnd(1)
R = pd.merge(R, ff, left_index=True, right_index=True)
R.AAPL = R.AAPL - R.RF/12
R.TSLA = R.TSLA - R.RF/12
R.AMZN = R.AMZN - R.RF/12
R = R.rename(columns={'Mkt-RF':'MRP'})
R.head()
```
When we run a multivariate linear regression, we will get a result that looks like this:
$$y =m_1x_1 + m_2x_2 + ... + m_nx_n$$
And here's how we can easily run the regression using statsmodels:
```
Y = R.AAPL
X = R[['MRP','SMB','HML']]
X = sm.add_constant(X)
model = sm.OLS(Y,X)
res = model.fit()
print(res.summary())
```
Very easy!
By including these additional factors, our R squared value increased from 0.365 to 0.495!
Another interesting factor to keep in mind is that the y intercept dropped from -0.0792 to 0.0040, becoming much closer to zero! In the 5 years of observable data, by including the additional factors, there is very little alpha for Apple!
*Tip: If you don't understand the coefficient of determination, Google search it to learn more about OLS and linear regression!*
## **Module 9.1.6: Multivariate Linear Regression, Formulaic Method**
We can also use the formulaic API for statsmodels!
This is exactly the same regression, the only difference being syntax!
```
resf = smf.ols(formula='AMZN ~ MRP + SMB + HML', data=R).fit()
print(resf.summary())
```
## **Statsmodels Linear Regression Summarized**
Using statsmodels, if our data is in a DataFrame, it's really easy to run a lienar regression!
Using the standard API it looks like this:
```python
Y = ___ # The Response Variable
X = ___ # Predictor(s) Variable
X = sm.add_constant(X)
model = sm.OLS(Y,X)
res = model.fit()
print(res.summary())
```
Or we can use the formulaic method:
```python
resf = smf.ols(formula='response ~ predictors', data=df).fit()
print(resf.summary())
```
And we can pull out our coefficients via the following:
```python
res.params['name']
```
```
```
| github_jupyter |
## Tutorial on how to 'delay' the start of particle advection
In many applications, it is needed to 'delay' the start of particle advection. For example because particles need to be released at different times throughout an experiment. Or because particles need to be released at a conatant rate from the same set of locations.
This tutorial will show how this can be done. We start with importing the relevant modules.
```
%matplotlib inline
from parcels import FieldSet, ParticleSet, JITParticle, plotTrajectoriesFile
from parcels import AdvectionRK4
import numpy as np
from datetime import timedelta as delta
```
First import a `FieldSet` (from the Peninsula example, in this case)
```
fieldset = FieldSet.from_parcels('Peninsula_data/peninsula', allow_time_extrapolation = True)
```
Now, there are two ways to delay the start of particles. Either by defining the whole `ParticleSet` at initialisation and giving each particle its own `time`. Or by using the `repeatdt` argument. We will show both options here
### Assigning each particle its own `time` ###
The simplest way to delaye the start of a particle is to use the `time` argument for each particle
```
npart = 10 # number of particles to be released
lon = 3e3 * np.ones(npart)
lat = np.linspace(3e3 , 45e3, npart, dtype=np.float32)
time = np.arange(0, npart) * delta(hours=1).total_seconds() # release every particle one hour later
pset = ParticleSet(fieldset=fieldset, pclass=JITParticle, lon=lon, lat=lat, time=time)
```
Then we can execute the `pset` as usual
```
output_file = pset.ParticleFile(name="DelayParticle_time.nc", outputdt=delta(hours=1))
pset.execute(AdvectionRK4, runtime=delta(hours=24), dt=delta(minutes=5),
output_file=output_file)
output_file.export() # export the trajectory data to a netcdf file
```
And then finally, we can show a movie of the particles. Note that the southern-most particles start to move first.
```
plotTrajectoriesFile('DelayParticle_time.nc', mode='movie2d_notebook')
```
### Using the `repeatdt` argument ###
The second method to delay the start of particle releases is to use the `repeatdt` argument when constructing a `ParticleSet`. This is especially useful if you want to repeatedly release particles from the same set of locations.
```
npart = 10 # number of particles to be released
lon = 3e3 * np.ones(npart)
lat = np.linspace(3e3 , 45e3, npart, dtype=np.float32)
repeatdt = delta(hours=3) # release from the same set of locations every 3 hours
pset = ParticleSet(fieldset=fieldset, pclass=JITParticle, lon=lon, lat=lat, repeatdt=repeatdt)
```
Now we again define an output file and execute the `pset` as usual.
```
output_file = pset.ParticleFile(name="DelayParticle_releasedt", outputdt=delta(hours=1))
pset.execute(AdvectionRK4, runtime=delta(hours=24), dt=delta(minutes=5),
output_file=output_file)
```
And we get an animation where a new particle is released every 3 hours from each start location
```
output_file.export() # export the trajectory data to a netcdf file
plotTrajectoriesFile('DelayParticle_releasedt.nc', mode='movie2d_notebook')
```
Note that, if you want to if you want to at some point stop the repeatdt, the easiest implementation is to use two calls to `pset.execute()`. For example, if in the above example you only want four releases of the pset, you could do the following
```
pset = ParticleSet(fieldset=fieldset, pclass=JITParticle, lon=lon, lat=lat, repeatdt=repeatdt)
output_file = pset.ParticleFile(name="DelayParticle_releasedt_9hrs", outputdt=delta(hours=1))
# first run for 3 * 3 hrs
pset.execute(AdvectionRK4, runtime=delta(hours=9), dt=delta(minutes=5),
output_file=output_file)
# now stop the repeated release
pset.repeatdt = None
# now continue running for the remaining 15 hours
pset.execute(AdvectionRK4, runtime=delta(hours=15), dt=delta(minutes=5),
output_file=output_file)
output_file.export() # export the trajectory data to a netcdf file
plotTrajectoriesFile('DelayParticle_releasedt_9hrs.nc', mode='movie2d_notebook')
```
| github_jupyter |
<p><font size="6"><b>CASE - Sea Surface Temperature data</b></font></p>
> *DS Python for GIS and Geoscience*
> *October, 2021*
>
> *© 2021, Joris Van den Bossche and Stijn Van Hoey. Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
---
```
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import xarray as xr
```
For this use case, we focus on the [Extended Reconstructed Sea Surface Temperature (ERSST)](https://www.ncdc.noaa.gov/data-access/marineocean-data/extended-reconstructed-sea-surface-temperature-ersst-v4), a widely used and trusted gridded compilation of historical Sea Surface Temperature (SST).
> The Extended Reconstructed Sea Surface Temperature (ERSST) dataset is a global monthly sea surface temperature dataset derived from the International Comprehensive Ocean–Atmosphere Dataset (ICOADS). It is produced on a 2° × 2° grid with spatial completeness enhanced using statistical methods. This monthly analysis begins in January 1854 continuing to the present and includes anomalies computed with respect to a 1971–2000 monthly climatology.
First we download the dataset. We will use the [NOAA Extended Reconstructed Sea Surface Temperature (ERSST)](https://psl.noaa.gov/thredds/catalog/Datasets/noaa.ersst/catalog.html?dataset=Datasets/noaa.ersst/sst.mnmean.v4.nc) v4 product. Download the data from this link: https://psl.noaa.gov/thredds/fileServer/Datasets/noaa.ersst/sst.mnmean.v4.nc and store it in a subfolder `data/` from the notebook as `sst.mnmean.v4.nc`.
Reading in the data set, ignoring the `time_bnds` variable:
```
data = './data/sst.mnmean.v4.nc'
ds = xr.open_dataset(data, drop_variables=['time_bnds'], engine="h5netcdf")
```
For this use case, we will focus on the years after 1960, so we slice the data from 1960 and load the data into our computer memory. By only loading the data after the initial slice, we make sure to only load into memory the data we specifically need:
```
ds = ds.sel(time=slice('1960', '2018')).load() # load into memory
ds
```
The data with the extension `nc` is a NetCDF format. NetCDF (Network Common Data Format) is the most widely used format for distributing geoscience data. NetCDF is maintained by the [Unidata](https://www.unidata.ucar.edu/) organization. Check the [netcdf website](https://www.unidata.ucar.edu/software/netcdf/docs/faq.html#whatisit) for more information. Xarray was designed to make reading netCDF files in python as easy, powerful, and flexible as possible.
__Note:__ As the data is in a [OPeNDAP server](https://en.wikipedia.org/wiki/OPeNDAP), we could also load the NETCDF data directly without downloading anything. This would require us to add the `netcdf4` package in our conda environment
### Exploratory data analysis
The data contains a single data variable `sst` and has 3 dimensions: lon, lat and time each described by a coordinate. Let's first get some insight in the structure and content of the data.
<div class="alert alert-success">
**EXERCISE**:
- What is the total amount of elements/values in the xarray data set?
- How many elements are there in the different dimensions
- The metadata of a netcdf file is also interpreted by xarray. Are the attributes on the xarray.Dataset `ds` the same as the attributes of the `sst` data itself?
<details>
<summary>Hints</summary>
- The number of elements or `size` of an array is an attribute of an xarray.DataArray and not of a xarray.Dataset
- Also the `shape` of an array is an attribute of an xarray.DataArray. A xarray.Dataset has the `dims` attribute to query dimension sizes
</details>
</div>
```
# size attribute of array object
ds["sst"].size
# shape attribute of array object
ds["sst"].shape
# dims attribute of dataset object
ds.dims
# attributes of array
ds["sst"].attrs
# attributes of data set
ds.attrs
```
---------
As we work with a single data variable, we will introduce a new variable `sst` which is the `xarray.DataArray` of the SST values. Note that we only keep the attributes on the xarray.DataArray level.
```
sst = ds["sst"]
sst
```
<div class="alert alert-success">
**EXERCISE**:
Make an image plot of the SST in the first month of the data set, January 1960. Adjust the range of the colorbar and switch to the `coolwarm` colormap.
<details>
<summary>Hints</summary>
- xarray can interpret a date string in the [ISO 8601](https://nl.wikipedia.org/wiki/ISO_8601) format as a date, e.g. `2020-01-01`.
- adjust ranges of the colorbar with `vmin` and `vmax`.
</details>
</div>
```
sst.sel(time="1960-01-01").plot.pcolormesh(vmin=-2, vmax=30,
cmap="coolwarm")
```
__Note__
xaray uses xarray.plot.pcolormesh() as the default two-dimensional plot method because it is more flexible than xarray.plot.imshow(). However, for large arrays, imshow can be much faster than pcolormesh. If speed is important to you and you are plotting a regular mesh, consider using imshow.
<div class="alert alert-success">
**EXERCISE**:
How did the SST evolve in time for a specific location on the earth? Make a line plot of the SST at `lon=300`, `lat=50` as a function of time.
Do you recognize the seasonality of the data?
<details>
<summary>Hints</summary>
- Use the `sel` for both the lon/lat selection.
</details>
</div>
```
sst.sel(lon=300, lat=50).plot.line();
```
<div class="alert alert-success">
**EXERCISE**:
What is the evolution of the SST as function of the month of the year?
Calculate the average SST with respect to the _month of the year_ for all positions in the data set and store the result as a variable `ds_mm`.
Use the `ds_mm` variable to make a plot: For longitude `164`, make a comparison in between the monthly average at latitude `-23.4` versus latitude `23.4`. Use a line plot with in the x-axis the month of the year and in the y-axis the average SST.
<details>
<summary>Hints</summary>
- Use the `sel` for both the lon/lat selection.
- If the exact values are not in the coordinate, you can use the `method="nearest"` inside a selection.
</details>
</div>
```
ds_mm = sst.groupby(sst.time.dt.month).mean(dim='time')
ds_mm.sel(lon=164, lat=[-23.4, 23.4], method="nearest").plot.line(hue="lat");
```
<div class="alert alert-success">
**EXERCISE**:
How does the zonal mean climatology for each month of the year changes with the latitude?
Reuse the `ds_mm` from the previous exercise or recalculate the average SST with respect to the _month of the year_ for all positions in the data set and store the result as a variable `ds_mm`.
To check the mean climatology (aggregating over the longitudes) as a function of the latitude for each month in the year, calculate the average SST over the `lon` dimension from `ds_mm`. Plot the result as an image with the month of the year in the x-axis and the latitude in the y-axis.
<details>
<summary>Hints</summary>
- You do not need another `groupby`, but need to calculate a reduction along a dimension.
</details>
</div>
```
ds_mm = sst.groupby(sst.time.dt.month).mean(dim='time')
ds_mm.mean(dim='lon').plot.imshow(x="month", y="lat", vmin=-2, vmax=30, cmap="coolwarm")
# alternative using transpose instead of defining the x and y in the plot function
ds_mm.mean(dim='lon').transpose().plot.imshow(vmin=-2, vmax=30, cmap="coolwarm")
```
<div class="alert alert-success">
**EXERCISE**:
How different is the mean climatology in between January and July?
Reuse the `ds_mm` variable from the previous exercises or recalculate the average SST with respect to the _month of the year_ for all positions in the data set and store the result as a variable `ds_mm`.
Calculate the difference of the mean climatology between January an July and plot the result as an image (map) with the longitude of the year in the x-axis and the latitude in the y-axis.
<details>
<summary>Hints</summary>
- You can subtract xarray just as Numpy arrays. You do not need another `groupby`, but only selections from the `ds_mm` variable.
</details>
</div>
```
(ds_mm.sel(month=1) - ds_mm.sel(month=7)).plot.imshow(vmax=10)
```
### Calculate the residual by removing climatology
To understand how the SST temperature evolved as a function of time during the last decades, we want to remove this climatology from the dataset and examine the residual, called the anomaly, which is the interesting part from a climate perspective.
We will do this by subtracting the monthly average from the values of that specific month. Hence, subtract the average January value over the years from the January data, subtract the average February value over the years from the February data,...
Removing the seasonal climatology is an example of a transformation: it operates over a group, but does not change the size of the dataset as we do the operation on each element (`x - x.mean()`)
This is not the same as the aggregations (e.g. `average`) we applied on each of the groups earlier. When using `groupby`, a calculation to the group can be applied and just like in Pandas, these calculations can either be:
- _aggregation_: reduces the size of the group
- _transformation_: preserves the groups full size
One way to consider is that we `apply` a function to each of the groups. For our anomaly calculation we want to do a _transformation_ and apply the following function:
```
def remove_time_mean(x):
"""Subtract each value by the mean over time"""
return x - x.mean(dim='time')
```
We can `apply` this function to each of the groups:
```
sst = ds["sst"]
ds_anom = sst.groupby('time.month').apply(remove_time_mean)
ds_anom
```
In other words:
> subtract each element by the average over time of all elements of the month the element belongs to
Xarray makes these sorts of transformations easy by supporting groupby arithmetic. This concept is easiest explained by applying it for our application:
```
gb = sst.groupby('time.month') # make groups (in this example each month of the year is a group)
ds_anom = gb - gb.mean(dim='time') # subtract each element of the group/month by the mean of that group/month over time
ds_anom
```
Now we can view the climate signal without the overwhelming influence of the seasonal cycle:
```
ds_anom.sel(lon=300, lat=50).plot.line()
```
Check the difference between Jan. 1 2018 and Jan. 1 1960 to see where the evolution in time is the largest:
```
(ds_anom.sel(time='2018-01-01') - ds_anom.sel(time='1960-01-01')).plot()
```
<div class="alert alert-success">
**EXERCISE**:
Compute the _five-year median_ of the `ds_anom` variable for the location `lon=300`, `lat=50` as well as the 12 month rolling median of the same data set. Store the output as respectively `ds_anom_resample` and `ds_anom_rolling`.
Make a line plot as a function of time for the location `lon=300`, `lat=50` of the original `ds_anom` data, the resampled data and the rolling median data.
<details>
<summary>Hints</summary>
- If you only need a single location, do the slicing (selecting) first instead of calculating them for all positions.
- Use the `resample` and the `rolling` functions.
</details>
</div>
```
# slice the point of interest
ds_anom_loc = ds_anom.sel(lon=300, lat=50)
# compute the resampling and rolling
ds_anom_resample = ds_anom_loc.resample(time='5Y').median(dim='time')
ds_anom_rolling = ds_anom_loc.rolling(time=12, center=True).median()
# make a combined plot
fig, ax = plt.subplots()
ds_anom_loc.plot.line(ax=ax, label="monthly anom")
ds_anom_resample.plot.line(marker='o', label="5 year resample")
ds_anom_rolling.plot.line(label='12 month rolling mean')
fig.legend(loc="upper center", ncol=3)
ax.set_title("");
```
### Make projection aware maps
The previous maps were the default outputs of xarray without specification of the spatial context. For reporting these plots are not appropriate. We can use the [cartopy](https://scitools.org.uk/cartopy/docs/latest/) package to adjust our Matplotlib axis to make them spatially aware.
For more in-depth information on cartopy, see the [visualization-03-cartopy](./visualization-03-cartopy.ipynb) notebook. As a short recap, to make sure the data of xarray can be integrated in a cartopy plot, the crucial element is to define the `transform` argument to to control which coordinate system that the given data is in. You can add the transform keyword with an appropriate `cartopy.crs.CRS` instance from the `import cartopy.crs` module:
```
import cartopy.crs as ccrs
import cartopy
map_proj = ccrs.Robinson() # Define the projection
fig, ax = plt.subplots(figsize = (16,9), subplot_kw={"projection": map_proj})
ax.gridlines()
ax.coastlines()
sst.sel(time="1960-01-01").plot(ax=ax, vmin=-2, vmax=30, center=5,
cmap='coolwarm', transform = ccrs.PlateCarree(), # tranform argument
cbar_kwargs={'shrink':0.75})
```
<div class="alert alert-success">
**EXERCISE**:
Make a plot of the `ds_anom` variable of 2018-01-01 with cartopy.
- Use the `ccrs.Orthographic` with the central lon/lat on -20, 5
- Add coastlines and gridlines to the plot
</div>
```
map_proj = ccrs.Orthographic(-20, 5)
fig, ax = plt.subplots(figsize = (12, 6), subplot_kw={"projection": map_proj})
ax.gridlines()
ax.coastlines()
ds_anom.sel(time='2018-01-01').plot(ax=ax, vmin=-1.5, vmax=1.5,
cmap='coolwarm', transform = ccrs.PlateCarree(),
cbar_kwargs={'shrink':0.5, 'label': 'anomaly'})
```
### Spatial aggregate per basin
Apart from aggregations as a function of time, also spatial aggregations using other (spatial) data sets can be achieved. In the next section, we want to compute the average SST over different ocean basins. The http://iridl.ldeo.columbia.edu/SOURCES/.NOAA/.NODC/.WOA09/.Masks/.basin/ is a data set that contains the main ocean basins in lon/lat:
```
basin = xr.open_dataset("./data/basin.nc")
basin = basin.rename({'X': 'lon', 'Y': 'lat'})
basin["basin"]
```
The name of the basins are included in the attributes of the data set. Using Pandas, we can create a mapping in between the basin names and the index used in the basin data set:
```
basin_names = basin["basin"].attrs['CLIST'].split('\n')
basin_s = pd.Series(basin_names, index=np.arange(1, len(basin_names)+1))
basin_s = basin_s.rename('basin')
basin_s.head()
```
We will use this mapping from identifier to label later in the analysis.
The basin data set provides multiple Z levels. We are interested in the division on surface level (0.0):
```
basin_surface = basin["basin"].sel(Z=0.0).drop_vars("Z")
basin_surface.plot(vmax=10, cmap='tab10')
```
The next step is to align both data sets. For this application, using the 'nearest' available data point will work to map both data sets with each other. Xarray provides the function `interp_like` to interpolate the `basin_surface` to the `sst` variable:
```
basin_surface_interp = basin_surface.interp_like(sst, method='nearest')
basin_surface_interp.plot(vmax=10, cmap='tab10')
```
<div class="alert alert-success">
**EXERCISE**:
Compute the mean SST (over all dimensions) for each of the basins in the `basin_surface` variable starting from the `sst` variable.
Next, we want to plot a horizontal bar chart with the SST for each bar chart. To do so:
- Convert the output to Pandas DataFrame.
- Combine the output with the `basin_s` variable by merging on the index (identifiers of the basin names).
- Create a horizontal barplot of the average temperature for each of the basins using the resulting dataframe.
<details>
<summary>Hints</summary>
- Use a `groupby` with the `basin_surface_interp` as input.
- Joining and merging of tables? See the [Pandas documentation](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.merge.html).
</details>
</div>
```
basin_mean_sst = sst.groupby(basin_surface_interp).mean()
basin_mean_sst = basin_mean_sst.mean(dim="time")
# In such a situation, the ELLIPSIS can be used to aggregate over all dimensions
basin_mean_sst = sst.groupby(basin_surface_interp).mean(dim=...) # ellipsis is shortcut for all dimensions
basin_mean_sst
# Convert to a Pandas DataFrame:
basin_mean_df = basin_mean_sst.to_dataframe()
basin_mean_df
# Merge the data with the `basin_s` data on the index:
basin_mean_df_merged = pd.merge(basin_mean_df, basin_s, left_index=True, right_index=True)
# Create a bar chart of the SST per basin data:
basin_mean_df_merged.sort_values(by="sst").plot.barh(x="basin");
```
-------
Acknowledgements to https://earth-env-data-science.github.io/lectures/xarray/xarray-part2.html
| github_jupyter |
```
%matplotlib inline
```
Training a Classifier
=====================
This is it. You have seen how to define neural networks, compute loss and make
updates to the weights of the network.
Now you might be thinking,
What about data?
----------------
Generally, when you have to deal with image, text, audio or video data,
you can use standard python packages that load data into a numpy array.
Then you can convert this array into a ``torch.*Tensor``.
- For images, packages such as Pillow, OpenCV are useful
- For audio, packages such as scipy and librosa
- For text, either raw Python or Cython based loading, or NLTK and
SpaCy are useful
Specifically for vision, we have created a package called
``torchvision``, that has data loaders for common datasets such as
ImageNet, CIFAR10, MNIST, etc. and data transformers for images, viz.,
``torchvision.datasets`` and ``torch.utils.data.DataLoader``.
This provides a huge convenience and avoids writing boilerplate code.
For this tutorial, we will use the CIFAR10 dataset.
It has the classes: ‘airplane’, ‘automobile’, ‘bird’, ‘cat’, ‘deer’,
‘dog’, ‘frog’, ‘horse’, ‘ship’, ‘truck’. The images in CIFAR-10 are of
size 3x32x32, i.e. 3-channel color images of 32x32 pixels in size.
.. figure:: /_static/img/cifar10.png
:alt: cifar10
cifar10
Training an image classifier
----------------------------
We will do the following steps in order:
1. Load and normalize the CIFAR10 training and test datasets using
``torchvision``
2. Define a Convolutional Neural Network
3. Define a loss function
4. Train the network on the training data
5. Test the network on the test data
1. Load and normalize CIFAR10
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Using ``torchvision``, it’s extremely easy to load CIFAR10.
```
import torch
import torchvision
import torchvision.transforms as transforms
```
The output of torchvision datasets are PILImage images of range **[0, 1]**.
We transform them to Tensors of normalized range **[-1, 1]**.
<div class="alert alert-info"><h4>Note</h4><p>If running on Windows and you get a BrokenPipeError, try setting
the num_worker of torch.utils.data.DataLoader() to 0.</p></div>
```
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
batch_size = 4
trainset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=batch_size,
shuffle=True, num_workers=2)
testset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
testloader = torch.utils.data.DataLoader(testset, batch_size=batch_size,
shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
```
Let us show some of the training images, for fun.
```
import matplotlib.pyplot as plt
import numpy as np
# functions to show an image
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# get some random training images
dataiter = iter(trainloader)
images, labels = dataiter.next()
# show images
imshow(torchvision.utils.make_grid(images))
# print labels
# print(' '.join('%5s' % classes[labels[j]] for j in range(batch_size)))
print(' '.join('{:5s}'.format(classes[labels[j]]) for j in range(batch_size)))
grid_image = torchvision.utils.make_grid(images)
grid_image = grid_image / 2 + 0.5
show_image = np.transpose(grid_image, (1, 2, 0))
plt.imshow(show_image)
```
2. Define a Convolutional Neural Network
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Copy the neural network from the Neural Networks section before and modify it to
take 3-channel images (instead of 1-channel images as it was defined).
```
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, 10)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = torch.flatten(x, 1) # flatten all dimensions except batch
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
net = Net()
```
3. Define a Loss function and optimizer
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
Let's use a Classification Cross-Entropy loss and SGD with momentum.
```
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(net.parameters(), lr=0.001, momentum=0.9)
```
4. Train the network
^^^^^^^^^^^^^^^^^^^^
This is when things start to get interesting.
We simply have to loop over our data iterator, and feed the inputs to the
network and optimize.
```
for epoch in range(2): # loop over the dataset multiple times
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs; data is a list of [inputs, labels]
inputs, labels = data
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 2000 == 1999: # print every 2000 mini-batches
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 2000))
running_loss = 0.0
print('Finished Training')
```
Let's quickly save our trained model:
```
PATH = './cifar_net.pth'
torch.save(net.state_dict(), PATH)
```
See `here <https://pytorch.org/docs/stable/notes/serialization.html>`_
for more details on saving PyTorch models.
5. Test the network on the test data
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
We have trained the network for 2 passes over the training dataset.
But we need to check if the network has learnt anything at all.
We will check this by predicting the class label that the neural network
outputs, and checking it against the ground-truth. If the prediction is
correct, we add the sample to the list of correct predictions.
Okay, first step. Let us display an image from the test set to get familiar.
```
dataiter = iter(testloader)
images, labels = dataiter.next()
# print images
imshow(torchvision.utils.make_grid(images))
print('GroundTruth: ', ' '.join('%5s' % classes[labels[j]] for j in range(4)))
```
Next, let's load back in our saved model (note: saving and re-loading the model
wasn't necessary here, we only did it to illustrate how to do so):
```
net = Net()
net.load_state_dict(torch.load(PATH))
```
Okay, now let us see what the neural network thinks these examples above are:
```
outputs = net(images)
```
The outputs are energies for the 10 classes.
The higher the energy for a class, the more the network
thinks that the image is of the particular class.
So, let's get the index of the highest energy:
```
_, predicted = torch.max(outputs, 1)
print('Predicted: ', ' '.join('%5s' % classes[predicted[j]]
for j in range(4)))
```
The results seem pretty good.
Let us look at how the network performs on the whole dataset.
```
correct = 0
total = 0
# since we're not training, we don't need to calculate the gradients for our outputs
with torch.no_grad():
for data in testloader:
images, labels = data
# calculate outputs by running images through the network
outputs = net(images)
# the class with the highest energy is what we choose as prediction
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: %d %%' % (
100 * correct / total))
```
That looks way better than chance, which is 10% accuracy (randomly picking
a class out of 10 classes).
Seems like the network learnt something.
Hmmm, what are the classes that performed well, and the classes that did
not perform well:
```
# prepare to count predictions for each class
correct_pred = {classname: 0 for classname in classes}
total_pred = {classname: 0 for classname in classes}
# again no gradients needed
with torch.no_grad():
for data in testloader:
images, labels = data
outputs = net(images)
_, predictions = torch.max(outputs, 1)
# collect the correct predictions for each class
for label, prediction in zip(labels, predictions):
if label == prediction:
correct_pred[classes[label]] += 1
total_pred[classes[label]] += 1
# print accuracy for each class
for classname, correct_count in correct_pred.items():
accuracy = 100 * float(correct_count) / total_pred[classname]
print("Accuracy for class {:5s} is: {:.1f} %".format(classname,
accuracy))
```
Okay, so what next?
How do we run these neural networks on the GPU?
Training on GPU
----------------
Just like how you transfer a Tensor onto the GPU, you transfer the neural
net onto the GPU.
Let's first define our device as the first visible cuda device if we have
CUDA available:
```
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# Assuming that we are on a CUDA machine, this should print a CUDA device:
print(device)
```
The rest of this section assumes that ``device`` is a CUDA device.
Then these methods will recursively go over all modules and convert their
parameters and buffers to CUDA tensors:
.. code:: python
```
net.to(device)
```
Remember that you will have to send the inputs and targets at every step
to the GPU too:
.. code:: python
```
inputs, labels = data[0].to(device), data[1].to(device)
```
Why don't I notice MASSIVE speedup compared to CPU? Because your network
is really small.
**Exercise:** Try increasing the width of your network (argument 2 of
the first ``nn.Conv2d``, and argument 1 of the second ``nn.Conv2d`` –
they need to be the same number), see what kind of speedup you get.
**Goals achieved**:
- Understanding PyTorch's Tensor library and neural networks at a high level.
- Train a small neural network to classify images
Training on multiple GPUs
-------------------------
If you want to see even more MASSIVE speedup using all of your GPUs,
please check out :doc:`data_parallel_tutorial`.
Where do I go next?
-------------------
- :doc:`Train neural nets to play video games </intermediate/reinforcement_q_learning>`
- `Train a state-of-the-art ResNet network on imagenet`_
- `Train a face generator using Generative Adversarial Networks`_
- `Train a word-level language model using Recurrent LSTM networks`_
- `More examples`_
- `More tutorials`_
- `Discuss PyTorch on the Forums`_
- `Chat with other users on Slack`_
| github_jupyter |
# Merging
```
# Don't change this cell; just run it.
import numpy as np
import pandas as pd
# Safe settings for Pandas.
pd.set_option('mode.chained_assignment', 'raise')
%matplotlib inline
import matplotlib.pyplot as plt
# Make the plots look more fancy.
plt.style.use('fivethirtyeight')
```
{ucb-page}`Joining_Tables_by_Columns`
Often, data about the same individuals is maintained in more than one table.
For example, one university office might have data about each student's time
to completion of degree, while another has data about the student's tuition
and financial aid.
To understand the students' experience, it may be helpful to put the two
datasets together. If the data are in two tables, each with one row per
student, then we would want to put the columns together, making sure to match
the rows so that each student's information remains on a single row.
Let us do this in the context of a simple example, and then use the method
with a larger dataset.
Suppose we have a data frame for different flavors of ice cream. Each flavor
of ice cream comes with a rating that is in a separate table.
```
cones = pd.DataFrame()
cones['Flavor'] = ['strawberry', 'vanilla', 'chocolate', 'strawberry',
'chocolate']
cones['Price'] = [3.55, 4.75, 6.55, 5.25, 5.75]
cones
ratings = pd.DataFrame()
ratings['Flavor'] = ['strawberry', 'chocolate', 'vanilla']
ratings['Stars'] = [2.5, 3.5, 4]
ratings
```
Each of the tables has a column that contains ice cream flavors. In both
cases, the column has the name `Flavor`. The entries in these columns can be
used to link the two tables.
The method `merge` creates a new table in which each cone in the `cones` table
is augmented with the Stars information in the `ratings` table. For each cone
in `cones`, `merge` finds a row in `ratings` whose `Flavor` matches the cone's
`Flavor`. We have to tell `merge` to use the `Flavor` column for matching,
using the `on` keyword argument.
```
rated = cones.merge(ratings, on='Flavor')
rated
```
Each cone now has not only its price but also the rating of its flavor.
In general, a call to `merge` that augments a table (say `table1`) with information from another table (say `table2`) looks like this:
table1.merge(table2, on=column_for_merging)
In the case above, the matching columns have the name column name: `Flavor`.
This need not be so. For example, let us rename the `Flavor` column in `ratings` to `Kind`:
```
# Rename the 'Flavor' column to 'Kind'
ratings_renamed = ratings.copy()
ratings_renamed.columns = ['Kind', 'Stars']
ratings_renamed
```
Now we have to tell `merge` the name of the column to merge on, for each data frame. The first data frame (`cones` in our case) is called the *left* data frame. The second (`ratings` in our case) is called the *right* data frame. Now the columns have different names in the left and right data frame, we have to use the `left_on` and `right_on` keywords.
```
rated_again = cones.merge(ratings_renamed, left_on='Flavor', right_on='Kind')
rated_again
```
Here is a more general skeleton of a `merge` between `table1` and `table2`, where the corresponding columns may have different names:
table1.merge(table2, left_on=table1_column, right_on=table2_column)
Now that we have done the merge, the new table `rated` (or `rated_again`)
allows us to work out the price per star, which you can think of as an
informal measure of value. Low values are good – they mean that you are paying
less for each rating star.
```
rated['$/Star'] = rated['Price'] / rated['Stars']
rated.sort_values('$/Star').head(3)
```
Though strawberry has the lowest rating among the three flavors, the less
expensive strawberry cone does well on this measure because it doesn't cost a
lot per star.
Suppose there is a table of professional reviews of some ice cream cones, and
we have found the average review for each flavor.
```
reviews = pd.DataFrame()
reviews['Flavor'] = ['vanilla', 'chocolate', 'vanilla', 'chocolate']
reviews['ProfStars'] = [5, 3, 5, 4]
reviews
```
Remember [group by](groupby):
```
average_review = reviews.groupby('Flavor').mean()
average_review
```
Notice that the column that we grouped by — `Flavor` — has become the Index (row labels).
We can merge `cones` and `average_review` by providing the labels of the
columns by which to merge. As you will see, Pandas treats the data frame Index
as being a column, for this purpose.
First we remind ourselves of the contents of `cones`:
```
cones
```
Here is the result of the merge:
```
cones.merge(average_review, left_on='Flavor', right_on='Flavor')
```
Of course in this case the "columns" have the same name, and we can do the same
thing with:
```
cones.merge(average_review, on='Flavor')
```
Notice that `Flavor` is the Index (row labels) for `average_review`, but Pandas
allows this, because it sees the Index name is `Flavor`, and treats it as a
column.
For this reason, merge can also merge with a Series, because the Series can
have a name. At the moment `average_review` is a Dataframe with one column:
`ProfStars`. We can pull out this column as a Series. You may remember that
the Series also gets the Index, and a name, from the column name:
```
avg_rev_as_series = average_review['ProfStars']
avg_rev_as_series
```
Because this Series has a name: `Flavor`, and that is the name of its Index, we can do the same merge with this Series as we did with the Dataframe above:
```
cones.merge(avg_rev_as_series, on='Flavor')
```
## What remains?
Notice that, after our merge, the strawberry cones have disappeared. Merge is
pursuing a particular strategy here, and that is to look for labels that match
in the matching columns. None of the reviews are for strawberry cones, so there
is nothing to which the `strawberry` rows can be merged. This might be what you
want, or it might not be — that depends on the analysis we are trying to
perform with the merged table. If it is not what you want, you may want to ask
merge to use a different strategy.
## Merge strategies
Let us reflect further on the choice that merge made above, when it dropped the
row for strawberry cones. As you saw above, by default, `merge` looks for
labels that are present in *both* of the matching columns. This is the default
merge strategy, called an *inner* merge. We could also call this an
*intersection* merge strategy.
For this default *inner* merge strategy, `merge` first found all the flavor
labels in `cones['Flavor']`:
```
# Different values in cones['Flavor']
cone_flavors = cones['Flavor'].unique()
cone_flavors
```
Then it found all the flavors in `average_review` `'Flavor'` "column" (in this case it found the index):
```
# Different values in average_reviews 'Flavor' - here, the Index
review_flavors = average_review.index.unique()
review_flavors
```
Next `merge` found all the `Flavor` values that are present in *both* data frames. We can call this the *intersection* of the two sets of values. Python has a `set` type to work out intersections and other set operations.
```
flavors_in_both = set(cone_flavors).intersection(review_flavors)
flavors_in_both
```
Merge then throws away any rows in either table that don't have one of these
intersection values in the matching columns. This is how we lost the `strawberry` row from the `cones` table.
This *inner* or *intersection* strategy is often useful — that is why it is the
default. But we may want to do something different. For example, we may want to keep flavors that don't have reviews in our merge result, but get a missing value for the review score. One way of doing that is the *left* merge strategy. Here merge keeps all rows from from the left data frame, but, for each row where there is no corresponding label in the right data frame, it fills the row values from the right data frame with missing values.
```
# A merge using the "left" strategy
cones.merge(average_review, on='Flavor', how='left')
```
## Merging and column names
Sometimes we find ourselves merging two data frames that have column names in common.
For example, imagine we had some user reviews from China:
```
chinese_reviews = pd.DataFrame()
chinese_reviews['Flavor'] = ['vanilla', 'chocolate', 'chocolate']
chinese_reviews['Stars'] = [4.5, 3.5, 4]
chinese_reviews
```
Now imagine we want to merge this data frame into the `rated` data frame. Here's the `rated` data frame:
```
rated
```
Notice that `rated` has a `Stars` column, and `chinese_reviews` has a `Stars` column. Let us see what `merge` does in this situation:
```
china_rated = rated.merge(chinese_reviews, on='Flavor')
china_rated
```
Pandas detected that both data frames had a column called `Stars` and has renamed them accordingly. The column from the left data frame (`rated`) has an `_x` suffix, to give `Stars_x`. The corresponding column from the right data frame has a `_y` suffix: `Stars_y`.
You can change these suffixes with the `suffixes` keyword argument:
```
rated.merge(chinese_reviews, on='Flavor', suffixes=['_left', '_right'])
```
## And more
There is much more information about merging in the [Python Data Science
Handbook merge section](https://jakevdp.github.io/PythonDataScienceHandbook/03.07-merge-and-join.html).
{ucb-page}`Joining_Tables_by_Columns`
| github_jupyter |
# Pandas GroupBy: Your Guide to Grouping Data in Python
___
download link:
[link1](http://archive.ics.uci.edu/ml/datasets/Air+Quality)
[link2](http://archive.ics.uci.edu/ml/datasets/News+Aggregator)
```
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
usecols = ["Date", "Time", "CO(GT)", "T", "RH", "AH"]
fill_value = -200
dates = ["Date", "Time"]
df = pd.read_excel("data/AirQualityUCI.xlsx",
parse_dates=[dates],
usecols=usecols,
na_values=[fill_value])
df.head()
df = df.rename(
columns={
"CO(GT)": "co",
"Date_Time": "tstamp",
"T": "temp_c",
"RH": "rel_hum",
"AH": "abs_hum"})
df.set_index("tstamp", inplace=True)
df.head()
df.index.min()
df.index.max()
day_names = df.index.day_name()
day_names
df.groupby(day_names, sort=True)['co'].mean()
df.groupby(df.index.day_of_week)['co'].mean()
hr = df.index.hour
hr
df.groupby([day_names, hr])["co"].mean().rename_axis(["dow", "hr"])
bins = pd.cut(df["temp_c"], bins=3, labels=("cool", "warm", "hot"))
df[["rel_hum", "abs_hum"]].groupby(bins).agg(["mean", "median"])
df.resample("Q")["co"].agg(["max", "min"])
import datetime as dt
def convertTime(x):
return dt.datetime.fromtimestamp(x/1000, tz=dt.timezone.utc)
df = pd.read_csv('data/newsCorpora.csv',
sep='\t',
header=None,
index_col=0,
names=["title", "url", "outlet", "category", "cluster", "host", "tstamp"],
nrows=900,
parse_dates=["tstamp"],
date_parser= convertTime )
df.head()
df = df.astype(dtype={
"outlet": "category",
"category": "category",
"cluster": "category",
"host": "category",
})
df.head()
df.shape
df.groupby("outlet", sort=False)["title"].apply(lambda x: x.str.contains("Fed").sum()).nlargest(10)
data1 = {'Name':['Jai', 'Princi', 'Gaurav', 'Anuj'],
'Age':[27, 24, 22, 32],
'Address':['Nagpur', 'Kanpur', 'Allahabad', 'Kannuaj'],
'Qualification':['Msc', 'MA', 'MCA', 'Phd']}
# Define a dictionary containing employee data
data2 = {'Name':['Abhi', 'Ayushi', 'Dhiraj', 'Hitesh'],
'Age':[17, 14, 12, 52],
'Address':['Nagpur', 'Kanpur', 'Allahabad', 'Kannuaj'],
'Qualification':['Btech', 'B.A', 'Bcom', 'B.hons']}
# Convert the dictionary into DataFrame
df = pd.DataFrame(data1,index=np.arange(0, 4))
# Convert the dictionary into DataFrame
df1 = pd.DataFrame(data2, index=np.arange(4, 8))
df1
df
pd.concat([df, df1])
pd.merge(df, df1, how='outer')
pd.concat([df, df1], join='inner')
pd.concat([df, df1], axis = 1, join='inner')
df2 = df.append(df1)
df2
pd.concat([df2, df1], axis = 1, join='inner')
data1 = {
'ID':[0, 1, 4, 3],
'Name':['Jai', 'Princi', 'Gaurav', 'Anuj'],
'Age':[27, 24, 22, 32],
'Address':['Nagpur', 'Kanpur', 'Allahabad', 'Kannuaj'],
'Qualification':['Msc', 'MA', 'MCA', 'Phd']}
# Convert the dictionary into DataFrame
df = pd.DataFrame(data1)
df.set_index('ID', inplace=True)
# creating a series
df
d = {'ID':[0,1,2,3], 'Salary':[1000, 2000, 3000, 4000]}
s1 = pd.DataFrame(d)
s1.set_index('ID', inplace=True)
s1
type(s1)
pd.merge(df, s1, left_on='ID', right_on='ID')
pd.concat([df, s1],join='inner', axis=1)
pd.merge(df, s1, on="ID")
pd.merge(df, s1,how='left', on="ID").fillna(0)
pd.merge(df, s1,how='outer', on="ID")
pd.merge(df, s1,how='right', on="ID").fillna('?')
```
| github_jupyter |
# Activity #1: yt
```
# first install yt!
#!pip install yt
# now import!
import yt
```
We'll use a dataset originally from the yt hub: http://yt-project.org/data/
Specifically, we'll use the IsolatedGalaxy dataset: http://yt-project.org/data/IsolatedGalaxy.tar.gz
```
# now, lets grab a dataset & upload it
# here's where mine is stored (in data)
ds = yt.load("/Users/jillnaiman1/data/IsolatedGalaxy/galaxy0030/galaxy0030")
```
This will be a bit of a repeat of a few weeks ago, but here we go!
```
# print out various stats of this dataset
ds.print_stats()
# this is basically telling us something about the
# number of data points in the dataset
# don't worry if you don't know what levels, grids or cells are at this point
# we'll get to it later
# same thing with field list, its cool if some of these look less familiar then others
ds.field_list
ds.derived_field_list
# this is a 3D simululation of a galaxy, lets check out some stats about the box
ds.domain_right_edge, ds.domain_left_edge
# what this is saying is the box goes from (0,0,0) to (1,1,1) in "code_length" units
# basically, this is just a normalized box
# you can also do fun things like print out max & min densities
ds.r[:].max("density"), ds.r[:].min("density")
# the above is for the whole box
# we can also ask where the maximum density is in this simulation box
ds.r[:].argmax("density")
# so this gives us x/y/z positions for where the maximum
# density is
# ok, lets make a quick plot 1/2 down the z-direction
# if the plot is too big for class try:
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = [3, 3]
p = ds.r[:, :, 0.5].plot("density")
# lets zoom
p.zoom(10)
# so, unless you're an astronomer you might be a little confused about these "kpc" units
# but yt allows us to change them! Behold cool yt units things:
(yt.units.kpc).in_units("cm")
# so we have now changed these weird kpc units
# yt also can do cool things with units like, yt units
# figures out some math stuff like, making things
# into cubed cm
(yt.units.kpc**3).in_units("cm**3")
# so lets set some units of our plot!
# lets change the units of density from g/cm^3 to kg/m^3
p.set_unit("density","kg/m**3")
# we can also include annotations on this plot
p.annotate_velocity()
# this shows how material is moving in this simulation
# this is shown with velocity vectors
# we can combine some of our coding around finding
# max values of density and combine with some
# region plots
# lets project the maximum density along the z axis
# i.e. lets make a plot of the maximum density
# along the z-axis of our plot
p2 = ds.r[:].max("density", axis="z").plot()
# we can zoom this as well
p2.zoom(10)
# if we scroll back up we can see that there is
# indeed a different between this and our slice plot
# here, we are much more "smeared" since we're picking
# only the max density -> everything looks brighter
# we can also do plots based on region selection
# but over specific values of z (and x & y)
# if we recall our box goes from 0->1 in each
# x/y/z direction, we can plot a zoom in
# like so:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].max("density", axis="z").plot()
# so, this shows the maximum density but only in a thin slice
# of the z-axis which is offset from the center
# sicne the galaxy lives at the center, and is the highest
# density gas region, it makes sense that our densities
# are lower and our features look different
# more "fuzzy ball" outside of the galaxy then
# gas flowing onto a galaxy disk
# lets redo the same plot but for the temperature of the gas:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].mean("temperature", axis="z").plot()
# we might want to highlight the temperature of the most dense regions
# why? well maybe we want to, instead of depicting the staight
# temperature, we want to depict the temperature
# of the *majority of the gas*
# we can do this by specifying a "weight" in our projection:
p = ds.r[0.1:0.9, 0.1:0.9, 0.55:0.65].mean("temperature", weight="density", axis="z").plot()
# so why is there this blocky structure? In space, we don't see cubes around galaxies...
# yet anyway...
# this is becuase this is a simulation of a galaxy, not an actual galaxy
# we can show why this might be by plotting the "grids" of this simulation
# over this thing:
p.annotate_grids()
# from this we can see that our grids sort of align where
# the temperature looks funny
# this is a good indicator that we have some numerical
# artifacts in our simulation
# ok! lets try some more analysis-like plots
# some of the helpful yt included plots is
ds.r[:].profile("density", "temperature").plot()
# so this is plotting the temperature of the gas
# in our simulation, in each binned density
# In our actual simulation, we have temperaturates
# at a variety of densities, and this is
# usualy the case, so by default what is plotted
# is the temperature (our 2nd param) plotted
# at each density bin, but weighted by the
# mass of material (gas) in each cell
# we can weight by other things, like in this case
# density:
ds.r[:].profile("density", "temperature", weight_field="density").plot()
# so similar shape (since mass and density are related)
# but a little different
# we can move this to a 2D plot
# to show the cell mass (as a color)
# as a function of both density and temprature
ds.r[:].profile(["density", "temperature"], "cell_mass", weight_field=None).plot()
# note: we can also do a 3D profile object,
# but there is currently no associated plot function with it
```
# Activity #2: Brain data with yt
```
# we can also use yt to play with other sorts of data:
import h5py # might have to pip install
# lets read our datafile into something called "scan_data"
with h5py.File("/Users/jillnaiman1/Downloads/single_dicom.h5", "r") as f:
scan_data = f["/scan"][:]
# if we recall, we had a weird shape of this data:
scan_data.shape
# so to import this data into yt to have
# yt make images for us, we need to do some formatting with numpy
import numpy as np
dsd = yt.load_uniform_grid({'scan': scan_data},
[36, 512, 512],
length_unit = yt.units.cm,
bbox = np.array([[0., 10], [0, 10], [0, 10]]),
)
dsd.r[:].mean("scan", axis="y").plot(); # this takes the mean along the specified axis "y" and plots
# can also do .max or .min
# note here that the number of fields
# availabel is much less:
dsd.field_list
# we can also look at different potions
# of the z-y axis by specifying
# the x-axis
p = dsd.r[0.75,:,:].plot('scan')
```
# Activity #3: Output images and objects (3D) with yt
Note: we'll do more with 3D objects next week/the last week, but this is a good first view of some cool ways we can output objects with yt
```
# lets go back to to our galaxy object
# and make a surface
# first, we'll cut down to a sphere and check
# that out
sphere = ds.sphere("max", (500.0, "kpc"))
sphere.mean("density", axis="y").plot(); # this takes the mean along the specified axis "y" and plots
# lets generate a surface of constant density
# i.e. we'll connect points on a surface
# where the density has a single value
surface = ds.surface(sphere, "density", 1e-27)
surface.export_obj('/Users/jillnaiman1/Downloads/myGalFiles',color_field='temperature')
# the above might take a while
# for checking out our surfaces right here
#http://www2.compute.dtu.dk/projects/GEL/PyGEL/
#!pip install PyGEL3D
from PyGEL3D import gel
from PyGEL3D import js
# for navigating
js.set_export_mode()
m = gel.obj_load("/Users/jillnaiman1/Downloads/myGalFiles.obj")
viewer = gel.GLManifoldViewer()
viewer.display(m)
# press ESC to quit? Yes, but then it takes a while so
# to get rid of the window
del viewer
```
Now, lets try with an inline viewer
```
# Can also display in the notebook
import numpy as np
#js.display(m,wireframe=False)
# comment out after you've run since we'll re-run below
```
Now lets try with an inline viewer & data colors
```
surf_temp = surface['temperature']
surf_temp.shape
# we see that this is infact a long list of values
# temperatures on each surface *face*
# if we look at the shape of the object:
m.positions().shape, surf_temp.shape[0]*3
# we see we have (surf_temp.shape)X3 times
# the number of points in x/y/z
# this is because these are *vertex* values
# so, if we want to color by something, we should use
# 3X the number of faces
js.display(m, data=np.repeat(np.log10(surf_temp),3),wireframe=False)
```
We can also process for 3D printing
```
surface.export_obj('/Users/jillnaiman1/Downloads/myGalFiles_print',dist_fac=0.001)
```
## Outputing images for things like clothing
```
p = ds.r[:, :, 0.5].plot("density")
p.zoom(20)
myImage = p.frb # fixed resoltuion binary
# we can then grab a simple image array
plt.imshow(np.array(myImage['density']))
# or we can turn off labels and grab a lovely image:
p = ds.r[:, :, 0.5].plot("density")
p.zoom(10)
p.hide_colorbar(); p.hide_axes();
p
# save the image
p.save('/Users/jillnaiman1/Downloads/myImage.png')
```
Now you have a lovely image that you can upload and put on things like sweaters or whatnot.
| github_jupyter |
```
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
from six.moves import range
import math
#load data,channge type
load = np.genfromtxt("/Users/yueyuchen/Documents/ML/ML_Immune/Machine_Learning_Immunogenicity/data/bcell.csv",dtype = str, delimiter = ',')
length1 = np.shape(load)[0]
labels1 = np.zeros(length1)
load1 = np.zeros(length1)
for i in range(length1):
labels1[i] = (load[i,1] == 'Positive')
for i in range(length1):
load1[i] = len(load[i,0])
data = []
labels = []
for i in range(length1):
if(load1[i] < 20):
data.append(load[i,0])
labels.append(labels1[i])
length = len(data)
dataset = []
for d in data:
d1 = []
for l in d:
d1.append(np.float32(ord(l)))
for j in range(20-len(d1)):
d1.append(np.float32(0))
dataset.append(d1)
for i in range(len(labels)):
if (labels[i] == 0):
labels[i] = [1,0]
else:
labels[i] = [0,1]
for i in range(length):
dataset[i] = np.asarray(dataset[i])
labels[i] = np.float32(np.asarray(labels[i]))
dataset = np.asarray(dataset)
labels = np.asarray(labels)
shuf = np.append(dataset,labels,axis = 1)
np.random.shuffle(c)
dataset = c[:,:20]
labels = c[:,20:22]
print (np.shape(dataset),np.shape(labels))
trainnum = 200000
testnum = 20000
train_dataset = dataset[:trainnum]
train_labels = labels[:trainnum]
test_dataset = dataset[trainnum:trainnum + testnum]
test_labels = labels[trainnum:trainnum + testnum]
valid_dataset = dataset[trainnum + testnum:trainnum + 2*testnum]
valid_labels = labels[trainnum + testnum:trainnum + 2*testnum]
print (dataset[5],)
graph = tf.Graph()
with graph.as_default():
# Input data.
# Load the training, validation and test data into constants that are
# attached to the graph.
tf_train_dataset = tf.constant(train_dataset)
tf_train_labels = tf.constant(train_labels)
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
# These are the parameters that we are going to be training. The weight
# matrix will be initialized using random values following a (truncated)
# normal distribution. The biases get initialized to zero.
weights = tf.Variable(
tf.truncated_normal([20, 2]))
biases = tf.Variable(tf.zeros([2]))
# Training computation.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(logits, tf_train_labels))
# Optimizer.
# We are going to find the minimum of this loss using gradient descent.
optimizer = tf.train.GradientDescentOptimizer(0.01).minimize(loss)
# Predictions for the training, validation, and test data.
# These are not part of training, but merely here so that we can report
# accuracy figures as we train.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
num_steps = 801
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
with tf.Session(graph=graph) as session:
tf.initialize_all_variables().run()
print('Initialized')
for step in range(num_steps):
_, l, predictions = session.run([optimizer, loss, train_prediction])
if (step % 100 == 0):
print('Loss at step %d: %f' % (step, l))
print('Training accuracy: %.1f%%' % accuracy(
predictions, train_labels))
print('Validation accuracy: %.1f%%' % accuracy(
valid_prediction.eval(), valid_labels))
print('Test accuracy: %.1f%%' % accuracy(test_prediction.eval(), test_labels))
```
| github_jupyter |
When benchmarking you **MUST**
1. close all applications
2. close docker
3. close all but this Web windows
4. all pen editors other than jupyter-lab (this notebook)
```
import os
from cloudmesh.common.Shell import Shell
from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
import pandas as pd
from tqdm.notebook import tqdm
from cloudmesh.common.util import readfile
from cloudmesh.common.util import writefile
from cloudmesh.common.StopWatch import StopWatch
from cloudmesh.common.systeminfo import systeminfo
import ipywidgets as widgets
sns.set_theme(style="whitegrid")
info = systeminfo()
user = info["user"]
node = info["uname.node"]
processors = 4
# Parameters
user = "gregor"
node = "alienware"
processors = 18
p = widgets.IntSlider(
value=processors,
min=1,
max=64,
step=1,
description='Processors:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
u = widgets.Text(value=user, placeholder='The user name', description='User:', disabled=False)
n = widgets.Text(value=node, placeholder='The computer name', description='Computer:', disabled=False)
display(p)
display(u)
display(n)
processors = p.value
user = u.value
node = n.value
print (processors, user, node)
experiments = 10
maximum = 1024 * 100000
intervals = 10
label = f"{user}-{node}-{processors}"
output = f"benchmark/{user}"
delta = int(maximum / intervals)
totals = [64] + list(range(0,maximum, delta))[1:]
points = [int(t/processors) for t in totals]
print (totals)
print(points)
os.makedirs(output, exist_ok=True)
systeminfo = StopWatch.systeminfo({"user": user, "uname.node": node})
writefile(f"{output}/{label}-sysinfo.log", systeminfo)
print (systeminfo)
df = pd.DataFrame(
{"Size": points}
)
df = df.set_index('Size')
experiment_progress = tqdm(range(0, experiments), desc ="Experiment")
experiment = -1
for experiment in experiment_progress:
exoeriment = experiment + 1
log = f"{output}/{label}-{experiment}-log.log"
os.system(f"rm {log}")
name = points[experiment]
progress = tqdm(range(0, len(points)),
desc =f"Benchmark {name}",
bar_format="{desc:<30} {total_fmt} {r_bar}")
i = -1
for state in progress:
i = i + 1
n = points[i]
#if linux, os:
command = f"mpiexec -n {processors} python count-click.py " + \
f"--n {n} --max_number 10 --find 8 --label {label} " + \
f"--user {user} --node={node} " + \
f"| tee -a {log}"
#if windows:
#command = f"mpiexec -n {processors} python count-click.py " + \
# f"--n {n} --max_number 10 --find 8 --label {label} " + \
# f"--user {user} --node={node} " + \
# f">> {log}"
os.system (command)
content = readfile(log).splitlines()
lines = Shell.cm_grep(content, "csv,Result:")
# print(lines)
values = []
times = []
for line in lines:
msg = line.split(",")[7]
t = line.split(",")[4]
total, overall, trials, find, label = msg.split(" ")
values.append(int(overall))
times.append(float(t))
# print (t, overall)
#data = pd.DataFrame(values, times, columns=["Values", "Time"])
#print (data.describe())
#sns.lineplot(data=data, palette="tab10", linewidth=2.5)
# df["Size"] = values
df[f"Time_{experiment}"] = times
# print(df)
df = df.rename_axis(columns="Time")
df
sns.lineplot(data=df, markers=True);
plt.savefig(f'{output}/{label}-line.png');
plt.savefig(f'{output}/{label}-line.pdf');
dfs = df.stack().reset_index()
dfs = dfs.set_index('Size')
dfs = dfs.drop(columns=['Time'])
dfs = dfs.rename(columns={0:'Time'})
dfs
sns.scatterplot(data=dfs, x="Size", y="Time");
plt.savefig(f"{output}/{label}-scatter.pdf")
plt.savefig(f"{output}/{label}-scatter.png")
sns.relplot(x="Size", y="Time", kind="line", data=dfs);
plt.savefig(f"{output}/{label}-relplot.pdf")
plt.savefig(f"{output}/{label}-relplot.png")
df.to_pickle(f"{output}/{label}-df.pkl")
```
| github_jupyter |
## Visualizing VGG-S filters
This is an ipython notebook to generate visualizations of VGG-S filters, for some more info refer to [this blogpost](https://auduno.github.io/2016/06/18/peeking-inside-convnets/).
To run this code, you'll need an installation of Caffe with built pycaffe libraries, as well as the python libraries numpy, scipy and PIL. For instructions on how to install Caffe and pycaffe, refer to the installation guide [here](http://caffe.berkeleyvision.org/installation.html). Before running the ipython notebooks, you'll also need to download the [VGG-S model](https://gist.github.com/ksimonyan/fd8800eeb36e276cd6f9), and modify the variables ```pycaffe_root``` to refer to the path of your pycaffe installation (if it's not already in your python path) and ```model_path``` to refer to the path of the downloaded VGG-S caffe model. Also uncomment the line that enables GPU mode if you have built Caffe with GPU-support and a suitable GPU available.
```
# imports and basic notebook setup
from cStringIO import StringIO
import numpy as np
import os,re,random
import scipy.ndimage as nd
import PIL.Image
import sys
from IPython.display import clear_output, Image, display
from scipy.misc import imresize
pycaffe_root = "/your/path/here/caffe/python" # substitute your path here
sys.path.insert(0, pycaffe_root)
import caffe
model_name="VGGS"
model_path = '/your/path/here/vggs_model/' # substitute your path here
net_fn = './VGG_CNN_S_deploy_mod.prototxt' # added force_backward : true to prototxt
param_fn = model_path + 'VGG_CNN_S.caffemodel'
means = np.float32([104.0, 117.0, 123.0])
#caffe.set_mode_gpu() # uncomment this if gpu processing is available
net = caffe.Classifier(net_fn, param_fn,
mean = means, # ImageNet mean, training set dependent
channel_swap = (2,1,0)) # the model has channels in BGR order instead of RGB
# a couple of utility functions for converting to and from Caffe's input image layout
def preprocess(net, img):
return np.float32(np.rollaxis(img, 2)[::-1]) - net.transformer.mean['data']
def deprocess(net, img):
return np.dstack((img + net.transformer.mean['data'])[::-1])
def blur(img, sigma):
if sigma > 0:
img[0] = nd.filters.gaussian_filter(img[0], sigma, order=0)
img[1] = nd.filters.gaussian_filter(img[1], sigma, order=0)
img[2] = nd.filters.gaussian_filter(img[2], sigma, order=0)
return img
def showarray(a, f, fmt='jpeg'):
a = np.uint8(np.clip(a, 0, 255))
f = StringIO()
PIL.Image.fromarray(a).save(f, fmt)
display(Image(data=f.getvalue()))
def make_step(net, step_size=1.5, end='inception_4c/output', clip=True, focus=None, sigma=None):
'''Basic gradient ascent step.'''
src = net.blobs['data'] # input image is stored in Net's 'data' blob
dst = net.blobs[end]
net.forward(end=end)
one_hot = np.zeros_like(dst.data)
filter_shape = dst.data.shape
if len(filter_shape) > 2:
# backprop only activation in middle of filter
one_hot[0,focus,(filter_shape[2]-1)/2,(filter_shape[3]-1)/2] = 1.
else:
one_hot.flat[focus] = 1.
dst.diff[:] = one_hot
net.backward(start=end)
g = src.diff[0]
src.data[:] += step_size/np.abs(g).mean() * g
if clip:
bias = net.transformer.mean['data']
src.data[:] = np.clip(src.data, -bias, 255-bias)
src.data[0] = blur(src.data[0], sigma)
dst.diff.fill(0.)
def deepdraw(net, base_img, octaves, random_crop=True, visualize=True, focus=None,
clip=True, **step_params):
# prepare base image
image = preprocess(net, base_img) # (3,224,224)
# get input dimensions from net
w = net.blobs['data'].width
h = net.blobs['data'].height
print "starting drawing"
src = net.blobs['data']
src.reshape(1,3,h,w) # resize the network's input image size
for e,o in enumerate(octaves):
if 'scale' in o:
# resize by o['scale'] if it exists
image = nd.zoom(image, (1,o['scale'],o['scale']))
_,imw,imh = image.shape
# select layer
layer = o['layer']
for i in xrange(o['iter_n']):
if imw > w:
if random_crop:
# randomly select a crop
#ox = random.randint(0,imw-224)
#oy = random.randint(0,imh-224)
mid_x = (imw-w)/2.
width_x = imw-w
ox = np.random.normal(mid_x, width_x*0.3, 1)
ox = int(np.clip(ox,0,imw-w))
mid_y = (imh-h)/2.
width_y = imh-h
oy = np.random.normal(mid_y, width_y*0.3, 1)
oy = int(np.clip(oy,0,imh-h))
# insert the crop into src.data[0]
src.data[0] = image[:,ox:ox+w,oy:oy+h]
else:
ox = (imw-w)/2.
oy = (imh-h)/2.
src.data[0] = image[:,ox:ox+w,oy:oy+h]
else:
ox = 0
oy = 0
src.data[0] = image.copy()
sigma = o['start_sigma'] + ((o['end_sigma'] - o['start_sigma']) * i) / o['iter_n']
step_size = o['start_step_size'] + ((o['end_step_size'] - o['start_step_size']) * i) / o['iter_n']
make_step(net, end=layer, clip=clip, focus=focus,
sigma=sigma, step_size=step_size)
if visualize:
vis = deprocess(net, src.data[0])
if not clip: # adjust image contrast if clipping is disabled
vis = vis*(255.0/np.percentile(vis, 99.98))
if i % 1 == 0:
showarray(vis,"./filename"+str(i)+".jpg")
if i % 10 == 0:
print 'finished step %d in octave %d' % (i,e)
# insert modified image back into original image (if necessary)
image[:,ox:ox+w,oy:oy+h] = src.data[0]
print "octave %d image:" % e
showarray(deprocess(net, image),"./octave_"+str(e)+".jpg")
# returning the resulting image
return deprocess(net, image)
octaves = [
{
'layer':'conv5',
'iter_n':200,
'start_sigma':2.5,
'end_sigma':1.1,
'start_step_size':12.*0.25,
'end_step_size':10.*0.25,
},
{
'layer':'conv5',
'iter_n':100,
'start_sigma':1.1,
'end_sigma':0.78*1.1,
'start_step_size':10.*0.25,
'end_step_size':8.*0.25
},
{
'layer':'conv5',
'scale':1.05,
'iter_n':100,
'start_sigma':0.78*1.1,
'end_sigma':0.78,
'start_step_size':8.*0.25,
'end_step_size':6.*0.25
},
{
'layer':'conv5',
'scale':1.05,
'iter_n':50,
'start_sigma':0.78*1.1,
'end_sigma':0.40,
'start_step_size':6.*0.25,
'end_step_size':1.5*0.25
},
{
'layer':'conv5',
'scale':1.05,
'iter_n':25,
'start_sigma':0.4,
'end_sigma':0.1,
'start_step_size':1.5*0.25,
'end_step_size':0.5*0.25
}
]
# get original input size of network
original_w = net.blobs['data'].width
original_h = net.blobs['data'].height
# the background color of the initial image
background_color = np.float32([250.0, 250.0, 250.0])
# generate initial random image
gen_image = np.random.normal(background_color, 8, (original_w, original_h, 3))
# which filter in layer to visualize (conv5 has 512 filters)
imagenet_class = 10
# generate class visualization via octavewise gradient ascent
gen_image = deepdraw(net, gen_image, octaves, focus=imagenet_class,
random_crop=True, visualize=False)
# save image
#img_fn = '_'.join([model_name, "deepdraw", str(imagenet_class)+'.png'])
#PIL.Image.fromarray(np.uint8(gen_image)).save('./' + img_fn)
```
| github_jupyter |
## Model Schema II
```
# Importing libraries
# Math and file management
import numpy as np
import re
from tqdm.auto import tqdm
import sys
import os
import urllib.request
import tarfile
import pickle
import fnmatch
import random
# For Model Building
from keras.layers import Conv2D, MaxPooling2D, Activation,BatchNormalization, UpSampling2D, Dropout, Flatten, Dense, Input, LeakyReLU, Conv2DTranspose,AveragePooling2D, Concatenate
from keras.preprocessing import image
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Model
from keras.models import load_model
from keras.optimizers import Adam
import keras.backend as K
from keras_contrib.layers.normalization.instancenormalization import InstanceNormalization
# For image processing
import skimage.color as imcolor
import PIL as Image
import matplotlib.pyplot as plt
# For processing time measurement
import time
# **THIS STEP SHOULD BE MADE AD HOC DEPENDING ON THE SOURCE OF THE DATASET**
# IN THIS CASE WE HAVE TO CHANGE THE DIRECTORY TO THE PATH WHERE THE DATASET IS LOCATED
def extract(pattern, compression_format, path):
cwd=os.chdir(path)
os.walk(cwd)
for root, dirs, files in os.walk(path):
for name in files:
if fnmatch.fnmatch(name, pattern):
tar = tarfile.open(name, compression_format)
tar.extractall()
tar.close()
return
extract('*.tar.gz',
'r:gz',
r'\Académico\Posgrados\2019 - Maestría en Ciencia de Datos e Innovación Empresarial\Tesis\Datasets')
def image_read(file, size=(256,256)):
'''
This function loads and resizes the image to the passed size and transforms that image into an array
Default image size is set to be 256x256
'''
img = image.load_img(file, target_size=size)
img = image.img_to_array(img)
return img
def image_convert(image_paths,size=256,channels=3):
'''
Redimensions images to Numpy arrays of a certain size and channels. Default values are set to 256x256x3 for coloured
images.
Parameters:
file_paths: a path to the image files
size: an int or a 2x2 tuple to define the size of an image
channels: number of channels to define in the numpy array
'''
# If size is an int
if isinstance(size, int):
# build a zeros matrix of the size of the image
all_images_to_array = np.zeros((len(image_paths), size, size, channels), dtype='int64')
for ind, i in enumerate(image_paths):
# reads image
img = image_read(i)
all_images_to_array[ind] = img.astype('int64')
print('All Images shape: {} size: {:,}'.format(all_images_to_array.shape, all_images_to_array.size))
else:
all_images_to_array = np.zeros((len(image_paths), size[0], size[1], channels), dtype='int64')
for ind, i in enumerate(image_paths):
img = read_img(i)
all_images_to_array[ind] = img.astype('int64')
print('All Images shape: {} size: {:,}'.format(all_images_to_array.shape, all_images_to_array.size))
return all_images_to_array
def find(pattern, path):
result = []
for root, dirs, files in os.walk(path):
for name in files:
if fnmatch.fnmatch(name, pattern):
result.append(os.path.join(root, name))
return result
image_paths=find('*.jpg', r'\Académico\Posgrados\2019 - Maestría en Ciencia de Datos e Innovación Empresarial\Tesis\Datasets\images2')
X_train=image_convert(image_paths)
def rgb_to_lab(img, l=False, ab=False):
"""
Takes in RGB channels in range 0-255 and outputs L or AB channels in range -1 to 1
"""
img = img / 255
lum = imcolor.rgb2lab(img)[:,:,0]
lum = (lum / 50) - 1
lum = lum[...,np.newaxis]
a_b = imcolor.rgb2lab(img)[:,:,1:]
a_b = (a_b + 128) / 255 * 2 - 1
if l:
return lum
else: return a_b
def lab_to_rgb(img):
"""
Takes in LAB channels in range -1 to 1 and out puts RGB chanels in range 0-255
"""
new_img = np.zeros((256,256,3))
for i in range(len(img)):
for j in range(len(img[i])):
pix = img[i,j]
new_img[i,j] = [(pix[0] + 1) * 50,(pix[1] +1) / 2 * 255 - 128,(pix[2] +1) / 2 * 255 - 128]
new_img = imcolor.lab2rgb(new_img) * 255
new_img = new_img.astype('uint8')
return new_img
L = np.array([rgb_to_lab(image, l=True) for image in X_train])
AB = np.array([rgb_to_lab(image, ab=True) for image in X_train])
L_AB_channels = (L,AB)
with open('l_ab_channels.p','wb') as f:
pickle.dump(L_AB_channels,f)
def load_images(filepath):
'''
Loads in pickle files, specifically the L and AB channels
'''
with open(filepath, 'rb') as f:
return pickle.load(f)
X_train_L, X_train_AB = load_images('l_ab_channels.p')
def resnet_block(x ,num_conv=2, num_filters=512,kernel_size=(3,3),padding='same',strides=2):
'''
This function defines a ResNet Block composed of two convolution layers and that returns the sum of the inputs and the
convolution outputs.
Parameters
x: is the tensor which will be used as input to the convolution layer
num_conv: is the number of convolutions inside the block
num_filters: is an int that describes the number of output filters in the convolution
kernel size: is an int or tuple that describes the size of the convolution window
padding: padding with zeros the image so that the kernel fits the input image or not. Options: 'valid' or 'same'
strides: is the number of pixels shifts over the input matrix.
'''
input=x
for i in range(num_conv):
input=Conv2D(num_filters,kernel_size=kernel_size,padding=padding,strides=strides)(input)
input=InstanceNormalization()(input)
input=LeakyReLU(0.2)(input)
return (input + x)
def generator(filters=64,num_enc_layers=4,num_resblock=4,name="Generator"):
'''
The generator per se is an autoencoder built by a series of convolution layers that initially extract features of the
input image.
'''
# defining input
x_0=Input(shape=(256,256,1))
'''
Adding first layer of the encoder model: 64 filters, 5x5 kernel size, 2 so the input size is reduced to half,
input size is the image size: (256,256,1), number of channels 1 for the luminosity channel.
We will use InstanceNormalization through the model and Leaky Relu with and alfa of 0.2
as activation function for the encoder, while relu as activation for the decoder.
between both of them, in the latent space we insert 4 resnet blocks.
'''
for lay in range(num_enc_layers):
x=Conv2D(filters*lay,(3,3),padding='same',strides=2,input_shape=(256,256,1))(x_0)
x=InstanceNormalization()(x)
x=LeakyReLU(0.2)(x)
'''
----------------------------------LATENT SPACE---------------------------------------------
'''
#for r in range(num_resblock):
# x=resnet_block(x)
'''
----------------------------------LATENT SPACE---------------------------------------------
'''
x=Conv2DTranspose(256,(3,3),padding='same',strides=2)(x)
x=InstanceNormalization()(x)
x=Activation('relu')(x)
x=Conv2DTranspose(128,(3,3),padding='same',strides=2)(x)
x=InstanceNormalization()(x)
x=Activation('relu')(x)
x=Conv2DTranspose(64,(3,3),padding='same',strides=2)(x)
x=InstanceNormalization()(x)
x=Activation('relu')(x)
x=Conv2DTranspose(32,(5,5),padding='same',strides=2)(x)
x=InstanceNormalization()(x)
x=Activation('relu')(x)
x=Conv2D(2,(3,3),padding='same')(x)
output=Activation('tanh')(x)
model=Model(x_0,output,name=name)
return model
def discriminator(name="Discriminator"):
# defining input
x_0=Input(shape=(256,256,2))
x=Conv2D(32,(3,3), padding='same',strides=2,input_shape=(256,256,2))(x_0)
x=LeakyReLU(0.2)(x)
x=Dropout(0.25)(x)
x=Conv2D(64,(3,3),padding='same',strides=2)(x)
x=BatchNormalization()(x)
x=LeakyReLU(0.2)(x)
x=Dropout(0.25)(x)
x=Conv2D(128,(3,3), padding='same', strides=2)(x)
x=BatchNormalization()(x)
x=LeakyReLU(0.2)(x)
x=Dropout(0.25)(x)
x=Conv2D(256,(3,3), padding='same',strides=2)(x)
x=BatchNormalization()(x)
x=LeakyReLU(0.2)(x)
x=Dropout(0.25)(x)
x=Flatten()(x)
x=Dense(1)(x)
output=Activation('sigmoid')(x)
model=Model(x_0,output,name=name)
return model
d_image_shape = (256,256,2)
g_image_shape = (256,256,1)
discriminator = discriminator()
discriminator.compile(loss='binary_crossentropy',
optimizer=Adam(lr=0.00008,beta_1=0.5,beta_2=0.999),
metrics=['accuracy'])
#Making the Discriminator untrainable so that the generator can learn from fixed gradient
discriminator.trainable = False
# Build the Generator
generator = generator()
#Defining the combined model of the Generator and the Discriminator
l_channel = Input(shape=g_image_shape)
image = generator(l_channel)
valid = discriminator(image)
combined_network = Model(l_channel, valid)
combined_network.compile(loss='binary_crossentropy',
optimizer=Adam(lr=0.0001,beta_1=0.5,beta_2=0.999))
#creates lists to log the losses and accuracy
gen_losses = []
disc_real_losses = []
disc_fake_losses=[]
disc_acc = []
#train the generator on a full set of 320 and the discriminator on a half set of 160 for each epoch
#discriminator is given real and fake y's while generator is always given real y's
n = 320
y_train_fake = np.zeros([160,1])
y_train_real = np.ones([160,1])
y_gen = np.ones([n,1])
#Optional label smoothing
#y_train_real -= .1
#Pick batch size and number of epochs, number of epochs depends on the number of photos per epoch set above
num_epochs=10
batch_size=32
for epoch in tqdm(range(1,num_epochs+1)):
#shuffle L and AB channels then take a subset corresponding to each networks training size
np.random.shuffle(X_train_L)
l = X_train_L[:n]
np.random.shuffle(X_train_AB)
ab = X_train_AB[:160]
fake_images = generator.predict(l[:160], verbose=1)
#Train on Real AB channels
d_loss_real = discriminator.fit(x=ab, y= y_train_real,batch_size=32,epochs=1,verbose=1)
disc_real_losses.append(d_loss_real.history['loss'][-1])
#Train on fake AB channels
d_loss_fake = discriminator.fit(x=fake_images,y=y_train_fake,batch_size=32,epochs=1,verbose=1)
disc_fake_losses.append(d_loss_fake.history['loss'][-1])
#append the loss and accuracy and print loss
disc_acc.append(d_loss_fake.history['acc'][-1])
#Train the gan by producing AB channels from L
g_loss = combined_network.fit(x=l, y=y_gen,batch_size=32,epochs=1,verbose=1)
#append and print generator loss
gen_losses.append(g_loss.history['loss'][-1])
#every 50 epochs it prints a generated photo and every 100 it saves the model under that epoch
if epoch % 50 == 0:
print('Reached epoch:',epoch)
pred = generator.predict(X_test_L[2].reshape(1,256,256,1))
img = lab_to_rgb(np.dstack((X_test_L[2],pred.reshape(256,256,2))))
plt.imshow(img)
plt.show()
if epoch % 100 == 0:
generator.save('generator_' + str(epoch)+ '_v3.h5')
```
| github_jupyter |
<a href="https://colab.research.google.com/github/malteranalytics/malteranalytics.github.io/blob/master/OpenPose.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# Pose Detection with OpenPose
This notebook uses an open source project [CMU-Perceptual-Computing-Lab/openpose](https://github.com/CMU-Perceptual-Computing-Lab/openpose.git) to detect/track multi person poses on a given youtube video.
## Install OpenPose
```
import os
from os.path import exists, join, basename, splitext
git_repo_url = 'https://github.com/CMU-Perceptual-Computing-Lab/openpose.git'
project_name = splitext(basename(git_repo_url))[0]
if not exists(project_name):
# see: https://github.com/CMU-Perceptual-Computing-Lab/openpose/issues/949
# install new CMake becaue of CUDA10
!wget -q https://cmake.org/files/v3.13/cmake-3.13.0-Linux-x86_64.tar.gz
!tar xfz cmake-3.13.0-Linux-x86_64.tar.gz --strip-components=1 -C /usr/local
# clone openpose
!git clone -q --depth 1 $git_repo_url
!sed -i 's/execute_process(COMMAND git checkout master WORKING_DIRECTORY ${CMAKE_SOURCE_DIR}\/3rdparty\/caffe)/execute_process(COMMAND git checkout f019d0dfe86f49d1140961f8c7dec22130c83154 WORKING_DIRECTORY ${CMAKE_SOURCE_DIR}\/3rdparty\/caffe)/g' openpose/CMakeLists.txt
# install system dependencies
!apt-get -qq install -y libatlas-base-dev libprotobuf-dev libleveldb-dev libsnappy-dev libhdf5-serial-dev protobuf-compiler libgflags-dev libgoogle-glog-dev liblmdb-dev opencl-headers ocl-icd-opencl-dev libviennacl-dev
# install python dependencies
!pip install -q youtube-dl
# build openpose
!cd openpose && rm -rf build || true && mkdir build && cd build && cmake .. && make -j`nproc`
from IPython.display import YouTubeVideo
```
## Detect poses on a test video
We are going to detect poses on the following youtube video:
```
YOUTUBE_ID = 'Gt6nBr1JsDw'
YouTubeVideo(YOUTUBE_ID)
```
Download the above youtube video, cut the first 5 seconds and do the pose detection on that 5 seconds:
```
!rm -rf youtube.mp4
# download the youtube with the given ID
!youtube-dl -f 'bestvideo[ext=mp4]' --output "youtube.%(ext)s" https://www.youtube.com/watch?v=$YOUTUBE_ID
# cut the first 5 seconds
!ffmpeg -y -loglevel info -i youtube.mp4 -t 5 video.mp4
# detect poses on the these 5 seconds
!rm openpose.avi
!cd openpose && ./build/examples/openpose/openpose.bin --video ../video.mp4 --write_json ./output/ --display 0 --write_video ../openpose.avi
# convert the result into MP4
!ffmpeg -y -loglevel info -i openpose.avi output.mp4
```
Finally, visualize the result:
```
def show_local_mp4_video(file_name, width=640, height=480):
import io
import base64
from IPython.display import HTML
video_encoded = base64.b64encode(io.open(file_name, 'rb').read())
return HTML(data='''<video width="{0}" height="{1}" alt="test" controls>
<source src="data:video/mp4;base64,{2}" type="video/mp4" />
</video>'''.format(width, height, video_encoded.decode('ascii')))
show_local_mp4_video('output.mp4', width=960, height=720)
```
| github_jupyter |
# Analysing lipid membrane data
This Jupyter notebook demonstrates the utility of the *refnx* for:
- the co-refinement of three contrast variation datasets of a DMPC (1,2-dimyristoyl-sn-glycero-3-phosphocholine) bilayer measured at the solid-liquid interface with a common model
- the use of the `LipidLeaflet` component to parameterise the model in terms of physically relevant parameters
- the use of Bayesian Markov Chain Monte Carlo (MCMC) to investigate the Posterior distribution of the curvefitting system.
- the intrinsic usefulness of Jupyter notebooks to facilitate reproducible research in scientific data analysis
The first step in most Python scripts is to import modules and functions that are going to be used
```
# use matplotlib for plotting
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import os.path
import refnx, scipy
# the analysis module contains the curvefitting engine
from refnx.analysis import CurveFitter, Objective, Parameter, GlobalObjective, process_chain
# the reflect module contains functionality relevant to reflectometry
from refnx.reflect import SLD, ReflectModel, Structure, LipidLeaflet
# the ReflectDataset object will contain the data
from refnx.dataset import ReflectDataset
```
In order for the analysis to be exactly reproducible the same package versions must be used. The *conda* packaging manager, and *pip*, can be used to ensure this is the case.
```
# version numbers used in this analysis
refnx.version.version, scipy.version.version
```
The `ReflectDataset` class is used to represent a dataset. They can be constructed by supplying a filename
```
pth = os.path.join(os.path.dirname(refnx.__file__), 'analysis', 'test')
data_d2o = ReflectDataset(os.path.join(pth, 'c_PLP0016596.dat'))
data_d2o.name = "d2o"
data_hdmix = ReflectDataset(os.path.join(pth, 'c_PLP0016601.dat'))
data_hdmix.name = "hdmix"
data_h2o = ReflectDataset(os.path.join(pth, 'c_PLP0016607.dat'))
data_h2o.name = "h2o"
```
A `SLD` object is used to represent the Scattering Length Density of a material. It has `real` and `imag` attributes because the SLD is a complex number, with the imaginary part accounting for absorption. The units of SLD are $10^{-6} \mathring{A}^{-2}$
The `real` and `imag` attributes are `Parameter` objects. These `Parameter` objects contain the: parameter value, whether it allowed to vary, any interparameter constraints, and bounds applied to the parameter. The bounds applied to a parameter are probability distributions which encode the log-prior probability of the parameter having a certain value.
```
si = SLD(2.07 + 0j)
sio2 = SLD(3.47 + 0j)
# the following represent the solvent contrasts used in the experiment
d2o = SLD(6.36 + 0j)
h2o = SLD(-0.56 + 0j)
hdmix = SLD(2.07 + 0j)
# We want the `real` attribute parameter to vary in the analysis, and we want to apply
# uniform bounds. The `setp` method of a Parameter is a way of changing many aspects of
# Parameter behaviour at once.
d2o.real.setp(vary=True, bounds=(6.1, 6.36))
d2o.real.name='d2o SLD'
```
The `LipidLeaflet` class is used to describe a single lipid leaflet in our interfacial model. A leaflet consists of a head and tail group region. Since we are studying a bilayer then inner and outer `LipidLeaflet`'s are required.
```
# Parameter for the area per molecule each DMPC molecule occupies at the surface. We
# use the same area per molecule for the inner and outer leaflets.
apm = Parameter(56, 'area per molecule', vary=True, bounds=(52, 65))
# the sum of scattering lengths for the lipid head and tail in Angstrom.
b_heads = Parameter(6.01e-4, 'b_heads')
b_tails = Parameter(-2.92e-4, 'b_tails')
# the volume occupied by the head and tail groups in cubic Angstrom.
v_heads = Parameter(319, 'v_heads')
v_tails = Parameter(782, 'v_tails')
# the head and tail group thicknesses.
inner_head_thickness = Parameter(9, 'inner_head_thickness', vary=True, bounds=(4, 11))
outer_head_thickness = Parameter(9, 'outer_head_thickness', vary=True, bounds=(4, 11))
tail_thickness = Parameter(14, 'tail_thickness', vary=True, bounds=(10, 17))
# finally construct a `LipidLeaflet` object for the inner and outer leaflets.
# Note that here the inner and outer leaflets use the same area per molecule,
# same tail thickness, etc, but this is not necessary if the inner and outer
# leaflets are different.
inner_leaflet = LipidLeaflet(apm,
b_heads, v_heads, inner_head_thickness,
b_tails, v_tails, tail_thickness,
3, 3)
# we reverse the monolayer for the outer leaflet because the tail groups face upwards
outer_leaflet = LipidLeaflet(apm,
b_heads, v_heads, outer_head_thickness,
b_tails, v_tails, tail_thickness,
3, 0, reverse_monolayer=True)
```
The `Slab` Component represents a layer of uniform scattering length density of a given thickness in our interfacial model. Here we make `Slabs` from `SLD` objects, but other approaches are possible.
```
# Slab constructed from SLD object.
sio2_slab = sio2(15, 3)
sio2_slab.thick.setp(vary=True, bounds=(2, 30))
sio2_slab.thick.name = 'sio2 thickness'
sio2_slab.rough.setp(vary=True, bounds=(0, 7))
sio2_slab.rough.name = name='sio2 roughness'
sio2_slab.vfsolv.setp(0.1, vary=True, bounds=(0., 0.5))
sio2_slab.vfsolv.name = 'sio2 solvation'
solv_roughness = Parameter(3, 'bilayer/solvent roughness')
solv_roughness.setp(vary=True, bounds=(0, 5))
```
Once all the `Component`s have been constructed we can chain them together to compose a `Structure` object. The `Structure` object represents the interfacial structure of our system. We create different `Structure`s for each contrast. It is important to note that each of the `Structure`s share many components, such as the `LipidLeaflet` objects. This means that parameters used to construct those components are shared between all the `Structure`s, which enables co-refinement of multiple datasets. An alternate way to carry this out would be to apply constraints to underlying parameters, but this way is clearer. Note that the final component for each structure is a `Slab` created from the solvent `SLD`s, we give those slabs a zero thickness.
```
s_d2o = si | sio2_slab | inner_leaflet | outer_leaflet | d2o(0, solv_roughness)
s_hdmix = si | sio2_slab | inner_leaflet | outer_leaflet | hdmix(0, solv_roughness)
s_h2o = si | sio2_slab | inner_leaflet | outer_leaflet | h2o(0, solv_roughness)
```
The `Structure`s created in the previous step describe the interfacial structure, these structures are used to create `ReflectModel` objects that know how to apply resolution smearing, scaling factors and background.
```
model_d2o = ReflectModel(s_d2o)
model_hdmix = ReflectModel(s_hdmix)
model_h2o = ReflectModel(s_h2o)
model_d2o.scale.setp(vary=True, bounds=(0.9, 1.1))
model_d2o.bkg.setp(vary=True, bounds=(-1e-6, 1e-6))
model_hdmix.bkg.setp(vary=True, bounds=(-1e-6, 1e-6))
model_h2o.bkg.setp(vary=True, bounds=(-1e-6, 1e-6))
```
An `Objective` is constructed from a `ReflectDataset` and `ReflectModel`. Amongst other things `Objective`s can calculate chi-squared, log-likelihood probability, log-prior probability, etc. We then combine all the individual `Objective`s into a `GlobalObjective`.
```
objective_d2o = Objective(model_d2o, data_d2o)
objective_hdmix = Objective(model_hdmix, data_hdmix)
objective_h2o = Objective(model_h2o, data_h2o)
global_objective = GlobalObjective([objective_d2o, objective_hdmix, objective_h2o])
```
A `CurveFitter` object can perform least squares fitting, or MCMC sampling on the `Objective` used to construct it.
```
fitter = CurveFitter(global_objective)
```
We'll just do a normal least squares fit here. MCMC sampling is left as an exercise for the reader.
```
fitter.fit('differential_evolution');
global_objective.plot()
plt.yscale('log')
plt.xlabel('Q / $\AA^{-1}$')
plt.ylabel('Reflectivity')
plt.legend();
```
We can display out what the fit parameters are by printing out an objective:
```
print(global_objective)
```
Let's example the scattering length density profile for each of the systems:
```
fig, ax = plt.subplots()
ax.plot(*s_d2o.sld_profile(), label='d2o')
ax.plot(*s_hdmix.sld_profile(), label='hdmix')
ax.plot(*s_h2o.sld_profile(), label='h2o')
ax.set_ylabel("$\\rho$ / $10^{-6} \AA^{-2}$")
ax.set_xlabel("z / $\AA$")
ax.legend();
```
| github_jupyter |
# Network in Network (NiN)
:label:`sec_nin`
LeNet, AlexNet, and VGG all share a common design pattern:
extract features exploiting *spatial* structure
via a sequence of convolution and pooling layers
and then post-process the representations via fully-connected layers.
The improvements upon LeNet by AlexNet and VGG mainly lie
in how these later networks widen and deepen these two modules.
Alternatively, one could imagine using fully-connected layers
earlier in the process.
However, a careless use of dense layers might give up the
spatial structure of the representation entirely,
*network in network* (*NiN*) blocks offer an alternative.
They were proposed based on a very simple insight:
to use an MLP on the channels for each pixel separately :cite:`Lin.Chen.Yan.2013`.
## (**NiN Blocks**)
Recall that the inputs and outputs of convolutional layers
consist of four-dimensional tensors with axes
corresponding to the example, channel, height, and width.
Also recall that the inputs and outputs of fully-connected layers
are typically two-dimensional tensors corresponding to the example and feature.
The idea behind NiN is to apply a fully-connected layer
at each pixel location (for each height and width).
If we tie the weights across each spatial location,
we could think of this as a $1\times 1$ convolutional layer
(as described in :numref:`sec_channels`)
or as a fully-connected layer acting independently on each pixel location.
Another way to view this is to think of each element in the spatial dimension
(height and width) as equivalent to an example
and a channel as equivalent to a feature.
:numref:`fig_nin` illustrates the main structural differences
between VGG and NiN, and their blocks.
The NiN block consists of one convolutional layer
followed by two $1\times 1$ convolutional layers that act as
per-pixel fully-connected layers with ReLU activations.
The convolution window shape of the first layer is typically set by the user.
The subsequent window shapes are fixed to $1 \times 1$.

:width:`600px`
:label:`fig_nin`
```
from mxnet import np, npx
from mxnet.gluon import nn
from d2l import mxnet as d2l
npx.set_np()
def nin_block(num_channels, kernel_size, strides, padding):
blk = nn.Sequential()
blk.add(nn.Conv2D(num_channels, kernel_size, strides, padding,
activation='relu'),
nn.Conv2D(num_channels, kernel_size=1, activation='relu'),
nn.Conv2D(num_channels, kernel_size=1, activation='relu'))
return blk
```
## [**NiN Model**]
The original NiN network was proposed shortly after AlexNet
and clearly draws some inspiration.
NiN uses convolutional layers with window shapes
of $11\times 11$, $5\times 5$, and $3\times 3$,
and the corresponding numbers of output channels are the same as in AlexNet. Each NiN block is followed by a maximum pooling layer
with a stride of 2 and a window shape of $3\times 3$.
One significant difference between NiN and AlexNet
is that NiN avoids fully-connected layers altogether.
Instead, NiN uses an NiN block with a number of output channels equal to the number of label classes, followed by a *global* average pooling layer,
yielding a vector of logits.
One advantage of NiN's design is that it significantly
reduces the number of required model parameters.
However, in practice, this design sometimes requires
increased model training time.
```
net = nn.Sequential()
net.add(nin_block(96, kernel_size=11, strides=4, padding=0),
nn.MaxPool2D(pool_size=3, strides=2),
nin_block(256, kernel_size=5, strides=1, padding=2),
nn.MaxPool2D(pool_size=3, strides=2),
nin_block(384, kernel_size=3, strides=1, padding=1),
nn.MaxPool2D(pool_size=3, strides=2),
nn.Dropout(0.5),
# There are 10 label classes
nin_block(10, kernel_size=3, strides=1, padding=1),
# The global average pooling layer automatically sets the window shape
# to the height and width of the input
nn.GlobalAvgPool2D(),
# Transform the four-dimensional output into two-dimensional output
# with a shape of (batch size, 10)
nn.Flatten())
```
We create a data example to see [**the output shape of each block**].
```
X = np.random.uniform(size=(1, 1, 224, 224))
net.initialize()
for layer in net:
X = layer(X)
print(layer.name, 'output shape:\t', X.shape)
```
## [**Training**]
As before we use Fashion-MNIST to train the model.
NiN's training is similar to that for AlexNet and VGG.
```
lr, num_epochs, batch_size = 0.1, 10, 128
train_iter, test_iter = d2l.load_data_fashion_mnist(batch_size, resize=224)
d2l.train_ch6(net, train_iter, test_iter, num_epochs, lr, d2l.try_gpu())
```
## Summary
* NiN uses blocks consisting of a convolutional layer and multiple $1\times 1$ convolutional layers. This can be used within the convolutional stack to allow for more per-pixel nonlinearity.
* NiN removes the fully-connected layers and replaces them with global average pooling (i.e., summing over all locations) after reducing the number of channels to the desired number of outputs (e.g., 10 for Fashion-MNIST).
* Removing the fully-connected layers reduces overfitting. NiN has dramatically fewer parameters.
* The NiN design influenced many subsequent CNN designs.
## Exercises
1. Tune the hyperparameters to improve the classification accuracy.
1. Why are there two $1\times 1$ convolutional layers in the NiN block? Remove one of them, and then observe and analyze the experimental phenomena.
1. Calculate the resource usage for NiN.
1. What is the number of parameters?
1. What is the amount of computation?
1. What is the amount of memory needed during training?
1. What is the amount of memory needed during prediction?
1. What are possible problems with reducing the $384 \times 5 \times 5$ representation to a $10 \times 5 \times 5$ representation in one step?
[Discussions](https://discuss.d2l.ai/t/79)
| github_jupyter |
# Table of Contents
1. [Single Run and Snapshot](#key1)<br>
1.1 [Select Halos by Mass](#singlemass)<br>
1.2 [Join Halos to Subhalos](#singlejoin)<br>
1.3 [Connect to Binary Catalog](#binary)
2. [Single Run and Multiple Snapshots](#key2)<br>
2.1 [Select Massive Halos in Each Snapshot](#masssnaps)<br>
2.2 [Calculate Mass Functions in Every Snapshot](#massfns)
3. [Multiple Runs and Single Snapshot](#key3)
4. [More Complex Examples](#key4)<br>
4.1. [Submitting Longer Queries with MyDB](#submit)<br>
4.2. [Select Halos in a Lightcone](#lightcone)
### This notebook gives examples of how to interact with the Indra halo database tables on the SciServer.
Indra database tables contain halo and subhalo catalog information and are accessed by executing SQL queries. In many cases querying the database is much faster than reading the binary catalog files. More example queries that might be useful can be found at the [Virgo - Millennium Database](http://gavo.mpa-garching.mpg.de/Millennium/Help?page=demo/genericqueries) pages.
```
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import indratools as indra
import pandas as pd
import SciServer.CasJobs as cj
import time
```
The halo and subhalo catalogs are stored in tables in the 'Indra' database context. There are two tables for each Indra run, from 2_0_0 to 7_7_7, containing FOF group or Subhalo information for every snapshot.
```
context = "Indra"
X = 2; Y = 3; Z = 4; snapnum = 58
tablename_fof = f'FOF_{X}{Y}{Z}'
tablename_sub = f'SubHalo_{X}{Y}{Z}'
```
Snapshot redshift and scalefactor information is also stored in a table in the Indra database. (`redshift` and `z` are redundant columns, probably for a good reason.)
```
snaps = cj.executeQuery("select * from Snapshots order by redshift",context)
snaps.columns.values
# What redshift did we pick with this snapnum?
snaps['redshift'][snaps['snapnum']==snapnum].values[0]
```
There are two ways of executing SQL with the SciServer.CasJobs API: `executeQuery` and `submitJob`. `executeQuery` returns a dataframe and is equivalent to the 'quick' execution in the SkyServer CasJobs UI: "quick queries are limited to one minute of execution and can only return about one meg of data before they will be cancelled" (see the [CasJobs Guide](https://skyserver.sdss.org/CasJobs/Guide.aspx) for more info). For more complex queries, `submitJob` returns a job ID to be used in e.g. `cj.waitForJob` but doesn't return results directly; see [Section 4](#key4) below for an example.
Selecting a full halo catalog at a low-redshift snapshot requires too much memory for the `executeQuery` command. It is better to select only what you need, or if you want to use the entire catalog (FOF + Subhalo) of one snapshot of one Indra run, you might as well load the binary catalog with `cat = indra.getsubcat((X,Y,Z),snapnum)`.
Here is a quick way to find out the column names of both tables:
```
cj.executeQuery(f'select top 1 * from {tablename_fof}',context)
cj.executeQuery(f'select top 1 * from {tablename_sub}',context)
```
Note that the masses are in units of 1e10 Msun/h (unlike the binary catalog), radii and positions in units of Mpc/h, and velocities in km/s. Note also that in the FOF tables, `np` refers to the number of particles in the main subhalo (e.g. `FirstSubOfHalo` given by `firstSubhaloId`) and not the parent FOF group.
# 1. Single Run and Snapshot
<a id="key1"></a>
## Select by mass
<a id="singlemass"></a>
The primary keys are `fofId` and `subhaloId` for the FOF and Subhalo tables. FOF tables are indexed on `snapnum` and `m_crit200`, and Subhalo tables on `snapnum` and `vDisp`.
Here we select halos above a given mass from a single run and snapshot. We define the minimum mass in the table's units of 1e10 Msun/h but return masses in units of Msun/h.
```
bigNumFOF = np.int(1e10)
bigNumSub = np.int(1e3)*bigNumFOF
minmass = 10000. # minmass = 1e4 corresponds to 1e14 Msun/h.
sql = f"""select np, numSubs, m_crit200*1.0e10 as mass, r_crit200 as radius from {tablename_fof}
where snapnum = {snapnum} and m_crit200 > {minmass}
order by mass desc
"""
cj.executeQuery(sql,context)
```
## Select by mass and select subhalos
<a id="singlejoin"></a>
Here we again select halos above a given mass from a given run and snapshot. We also return subhalos of these halos using SQL `inner join` and the positions of the subhalos. (Note that masses are only defined for FOF groups == main subhalos). We define a new column that keeps track of whether the halo is the main subhalo of the FOF group or not, using SQL `case when` syntax.
```
minmass = 10000. # minmass = 1e4 corresponds to 1e14 Msun/h.
sql = f"""select h.fofId, s.subhaloId, s.x, s.y, s.z,
case when h.firstSubhaloId = s.subhaloId then 1 else 0 end as isMain
from {tablename_fof} h inner join {tablename_sub} s
on h.fofId = s.fofId
and h.snapnum = {snapnum} and h.m_crit200 > {minmass}
"""
df = cj.executeQuery(sql,context)
df
```
Let's plot them! We'll restrict the plot to a sub-volume of the full Gpc/h cube.
```
subdf = df[((df['x'] > 200) & (df['x'] < 600) & (df['y'] < 400) & (df['z'] > 300) & (df['z'] < 700))]
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111,projection='3d')
ax.scatter(subdf['x'][subdf['isMain']==0].values,subdf['y'][subdf['isMain']==0].values,subdf['z'][subdf['isMain']==0].values,marker='.',color='k')
ax.scatter(subdf['x'][subdf['isMain']==1].values,subdf['y'][subdf['isMain']==1].values,subdf['z'][subdf['isMain']==1].values,marker='o',color='m')
ax.set_xlabel('x (Mpc/h)')
ax.set_ylabel('y (Mpc/h)')
ax.set_zlabel('z (Mpc/h)');
```
## Connect DB Halos to Binary Catalog
<a id="binary"></a>
It is possible to connect the halos in the database tables to the corresponding halos as read by the binary catalog. Since all of the catalog information is already in the database tables, the main reason you might want to do this is to pick out the particles in a halo by appropriately indexing the ID arrays returned with `indra.getfofids` and `indra.getsubids`. (See `read_examples.ipynb` for how to do that.)
We'll start with the FOF groups. The database tables only contain those groups with at least one subhalo (which means the group has not been dissolved by the SUBFIND unbinding process).
```
cat = indra.getsubcat((X,Y,Z),snapnum)
hasSubs = np.where(cat['NsubPerHalo'] > 0)[0]
```
To relate the `fofId` values we will index a dataframe that contains all, and only, the FOF halos in this Indra run and at this `snapnum`, ordered by `fofId`.
```
fofdf = cj.executeQuery(f'select fofid, np from {tablename_fof} where snapnum={snapnum} order by fofid',context)
```
As an example, let's match the 4 halos containing the most particles. As noted before, `np` in the database refers to the number of particles in the main subhalo (e.g. `FirstSubOfHalo` given by `firstSubhaloId`) and not the parent FOF group, so the appropriate value in the catalog is the `subLen` of the `FirstSubOfHalo`, where `FirstSubOfHalo` is indexed by the FOF ID.
```
halos = cj.executeQuery(f"select top 4 fofid, np from {tablename_fof} where snapnum={snapnum} order by np desc",context)
binfofids = [hasSubs[fofdf.index[fofdf['fofid'] == halos['fofid'][i]].values[0]] for i in range(len(halos))]
binfofids
for i in range(len(halos)):
print('Np from catalog = {}, Np from DB = {}'.format(cat['subLen'][cat['FirstSubOfHalo'][binfofids[i]]],fofdf['np'][fofdf['fofid']==halos['fofid'][i]].values[0]))
```
The subhalo tables contain all of the subhalos in the binary catalogs, so they are a bit simpler.
```
subdf = cj.executeQuery(f'select subhaloId, np from {tablename_sub} where snapnum={snapnum} order by subhaloId',context)
subhalos = cj.executeQuery(f"select top 4 subhaloId, np from {tablename_sub} where snapnum={snapnum} order by np desc",context)
binsubids = [subdf.index[subdf['subhaloId'] == subhalos['subhaloId'][i]].values[0] for i in range(len(subhalos))]
binsubids
for i in range(len(subhalos)):
print('Np from catalog = {}, Np from DB = {}'.format(cat['subLen'][binsubids[i]],subdf['np'][subdf['subhaloId']==subhalos['subhaloId'][i]].values[0]))
```
# 2. Single Run and Multiple Snapshots
<a id="key2"></a>
## Select massive halos in each snapshot
<a id="masssnaps"></a>
In this example we'll select the most massive halo from each snapshot of a given run, while noting that the most massive halo won't necessarily be "the same" halo across snapshots.
Selecting the most massive halo in SQL uses the aggregate function `MAX` with the `GROUP BY` syntax to aggregate by `snapnum`. To select other columns that correspond to this maximum-mass halo row, use `join` on the results of the aggregating `select` clause. Note that if the two most massive halos have the same mass (unlikely but possible at early snapshots), both are returned.
```
sql = f"""select h.snapnum, h.np, h.numSubs, h.r_crit200 as radius, h.m_crit200*1.0e10 as mass,
h.x, h.y, h.z
from {tablename_fof} h join (select MAX(m_crit200) as mass, snapnum FROM {tablename_fof} GROUP BY snapnum) max
on h.m_crit200 = max.mass and h.snapnum = max.snapnum
order by h.snapnum
"""
df = cj.executeQuery(sql,context)
df.head(8)
```
If you want the top N most massive halos at each snapshot, use the `rank` syntax: `partition by` determines what to group on (in this case `snapnum`), and `order by` determines how to assign ranks within the group.
```
n_massive = 3 # select n_massive most massive halos
sql= f"""
with a as(
select fofid
, rank() over(partition by snapnum order by m_crit200 desc) as rank
from {tablename_fof})
select h.snapnum, h.np, h.numSubs, h.r_crit200 as radius, h.m_crit200*1.0e10 as mass,
h.x, h.y, h.z, a.rank
from a join {tablename_fof} h on h.fofid = a.fofid
where a.rank <= {n_massive}
"""
rankdf = cj.executeQuery(sql,context)
rankdf.head(10)
```
For fun let's plot the most massive halo positions and color them by their `snapnum`.
```
fig = plt.figure(figsize=(8,6))
plt.scatter(df['y'],df['z'],marker='o',s=50,c=df['snapnum'].values)
plt.colorbar()
plt.xlabel('y (Mpc/h)')
plt.ylabel('z (Mpc/h)');
```
## Calculate Mass Functions in Every Snapshot
<a id="massfns"></a>
You can very quickly calculate the number of halos in bins of mass in SQL by using `floor` to round down the masses and then aggregating. (This is the [Millennium DB](http://gavo.mpa-garching.mpg.de/Millennium/Help?page=demo/genericqueries) example H5). Note that if there are no halos in a given mass range in a given snapshot, this will not return a count of 0 but will simply skip it.
```
binsize = .1
sql = f"""select snapnum, power(10.0,{binsize}*floor(.5+log10(m_crit200)/{binsize}))*1e10 as mass, count(*) as num
from {tablename_fof}
where m_crit200 > 0
group by snapnum, floor(.5+log10(m_crit200)/{binsize})
order by 1 desc, 2
"""
df = cj.executeQuery(sql,context)
df
sn = 63
df[df['snapnum']==sn].plot(x='mass',y='num',logx=True,logy=True,title=f"z={snaps['redshift'][snaps['snapnum']==sn].values[0]:.2f}");
sn = 20
df[df['snapnum']==sn].plot(x='mass',y='num',logx=True,logy=True,title=f"z={snaps['redshift'][snaps['snapnum']==sn].values[0]:.2}");
```
# 3. Multiple Runs and Single Snapshot
<a id="key3"></a>
In the current schema, each Indra volume is in a separate table, and it is cumbersome to join many (hundreds of!) tables in one query. Instead, we will define a function that gets what we want from one table, then loop over that function for each desired Indra run.
In this example we will calculate histograms of mass for one snapshot and many Indra runs, then compute the average and variance of the mass functions. This took 2 minutes to compute 64 mass functions, versus 13 minutes computing mass functions by reading the binary catalog files.
```
def select_massfn(snapnum,table,binsize=.1,maxmass=None):
if maxmass is None:
sql = f"""select power(10.0,{binsize}*floor(.5+log10(m_crit200)/{binsize}))*1.0e10 as mass, count(*) as num
from {table}
where m_crit200 > 0 and snapnum = {snapnum}
group by floor(.5+log10(m_crit200)/{binsize})
order by 1
"""
else:
sql = f"""select power(10.0,{binsize}*floor(.5+log10(m_crit200)/{binsize}))*1.0e10 as mass, count(*) as num
from {table}
where m_crit200 > 0 and m_crit200 < {maxmass/1e10} and snapnum = {snapnum}
group by floor(.5+log10(m_crit200)/{binsize})
order by 1
"""
return sql
# let's try a loop of 64 - use 3_Y_Z
# we'll limit max mass to 1e15 in hopes that this means there are no empty bins
runstart = indra.Run((3,0,0)).num
nruns = 64
boxsize=1000.
maxmass = 1.0e15
# how many bins will there be?
df = cj.executeQuery(select_massfn(snapnum,'FOF_300',maxmass=maxmass),context)
nbins = len(df)
t1 = time.time()
nhalo = np.zeros((nbins,nruns),dtype=np.float32)
massfn = np.zeros((nbins,nruns),dtype=np.float32)
for i, runnum in enumerate(range(runstart,runstart+nruns)):
run = indra.Run(runnum)
tablename = f'FOF_{run.X}{run.Y}{run.Z}'
df = cj.executeQuery(select_massfn(snapnum,tablename,maxmass=maxmass),context)
nhalo[:,i] = df['num'].values
massfn[:,i] = np.flipud(np.cumsum(np.flipud(nhalo[:,i])))/boxsize**3
# could be more careful here and save df['mass'] too
t2 = time.time()
print(f'64 mass functions in {t2-t1:.3} sec')
mbins = df['mass'].values # the last run queried
len(df) == nbins
mfmean = np.mean(massfn,1)
mfsig = np.std(massfn,1)
plt.figure(figsize=(8,6))
plt.plot(mbins,mfmean,'k',linewidth=2,label='mean')
plt.fill_between(mbins,mfmean-mfsig,mfmean+mfsig,alpha=.3,color='k')
plt.xscale('log')
plt.yscale('log')
plt.xlabel(r'$M_{200}$ ($M_{\odot}/h$)',size='large')
plt.ylabel(r'$n(>M_{200}) (h^3/Mpc^3)$',size='large');
nhmean = np.mean(nhalo,1)
nhsig = np.std(nhalo,1)
plt.figure(figsize=(8,6))
plt.plot(mbins,nhsig/(nhmean*np.sqrt(nruns)))
plt.xscale('log')
plt.yscale('log')
plt.xlabel(r'$M_{200}$ ($M_{\odot}/h$)',size='large')
plt.ylabel('Standard Error',size='large');
```
# 4. More Complex Examples
<a id="key4"></a>
### Submitting Longer Queries with MyDB
<a id="submit"></a>
For queries that take some time or return a lot of data, use `submitJob` instead of `executeQuery` and save the results to a new table in your personal `MyDB` database context. We first need to drop this table if it already exists, e.g. if you have run this code before and want to run it again. The below pseudocode demonstrates the general steps involved in submitting longer queries into a MyDB table and retrieving the results.
### Select Halos in a Lightcone
<a id="lightcone"></a>
We can use the Spatial3D library to efficiently select halos that are within a lightcone. We will do this by combining a series of cone segments, where each segment gathers everything in one snapshot, for all 64 snapshots or up to the highest desired redshift. The minimum and maximum extent in Mpc/h of these segments have already been loaded into the table `segments`, with the boundaries given by the median redshift between two consecutive snapshots.
```
from astropy.cosmology import FlatLambdaCDM
header = indra.getheader()
Indra = FlatLambdaCDM(H0=header['hubble']*100,Om0=header['omega0'])
Indra.box = header['BoxSize']
Indra.bits = 6 # defines resolution of Peano-Hilbert index
segments = cj.executeQuery('select * from segments order by snapnum desc',context)
segments['redshift'] = cj.executeQuery('select redshift from Snapshots order by snapnum desc',context)
segments.head(5)
```
The below function runs a query that creates and combines cone segments in given redhshift range according to input cone parameters: origin, direction, and opening angle. It selects all the halos within these cone segments, at the appropriate snapshots, with the desired minimum number of particles or halo mass, and saves this as a table in your personal MyDB space. It then queries everything in the table and returns a lightcone object.
```
def lightcone(sim, O, D, angle, lo_z, hi_z,tablename,nplimit=0.,masslimit=0.,DEBUG=False,lc_table=None):
"""
sim: simulation/cosmology object. astropy.cosmology.FlatLambdaCDM with Indra properties
O: origin [x,y,z]
D: direction [dx,dy,dz]
angle: opening half-angle in degrees
lo: lowest redshift
hi: highest redshift
tablename: FOF or SubHalo table name for desired Indra volume
"""
h = sim.H0.value/100
lo_cm = sim.comoving_distance(lo_z).value*h # min comoving dist in Mpc/h
hi_cm = sim.comoving_distance(hi_z).value*h # max comoving dist in Mpc/h
if lc_table is None:
lc_table = "lightcone_"+tablename
sql = f"""
IF OBJECT_ID('{lc_table}', 'U') IS NOT NULL
DROP TABLE {lc_table}
"""
cj.executeQuery(sql,"MyDB")
sql = f"""
declare @box dbo.Box=dbo.Box::New(0,0,0,{sim.box},{sim.box},{sim.box})
declare @lo real={lo_cm}, @hi real={hi_cm}, @angle real=RADIANS(cast({angle} as float))
declare @bits tinyint = {sim.bits}
;
with lightcone as
(
select snapnum
, dbo.Shape::New(dbo.ConeSegment::New({",".join(map(str, O))},{",".join(map(str, D))}
, @angle, dbo.math_max(@lo, lo), dbo.math_min(@hi,hi)).ToString()) as shape
from Indra.segments
where lo <= @hi and hi >= @lo
),
cover as (
select snapnum,shape,c.*
from lightcone cross apply dbo.fCover('H',@bits,@box,1,shape) c
)
select h.fofid,h.np, h.snapnum,h.m_crit200*1.e10 as mass, h.r_crit200
, h.x+sh.shiftx as x
, h.y+sh.shifty as y
, h.z+sh.shiftz as z
into MyDB.{lc_table}
from cover sh
inner loop join Indra.{tablename} h
on h.snapnum=sh.snapnum
and h.phkey between sh.keymin and sh.keymax
and sh.shape.ContainsPoint(h.x+sh.shiftx,h.y+sh.shifty,h.z+sh.shiftz)=1
and h.np > {nplimit}
and h.m_crit200*1.e10 > {masslimit}
"""
if DEBUG:
print(sql)
return
jobid = cj.submitJob(sql,"Indra")
%time cj.waitForJob(jobid,verbose=True)
df = cj.executeQuery(f"select * from {lc_table}","MyDB")
lc = {"O":O, "D":D, "angle":angle, "lo_z":lo_z, "hi_z":hi_z, "table":tablename, "df":df}
return lc
O = [0,0,0] # origin vector
D = [1,.5,1] # direction vector
angle = 1 # opening half-angle in degrees
lc = lightcone(Indra,O,D,angle,lo_z=0,hi_z=6,tablename=tablename_fof)
df = lc['df']
```
Let's plot the results, showing cone segments with alternating colors. Note that many segments fit in one Indra volume - they are defined by the redshifts of the snapshots.
```
fig = plt.figure(figsize=(10,10))
colors = ['blue','yellow']
ax = fig.add_subplot(111,projection='3d')
ax.scatter(df['x'],df['y'],df['z'],s=.1,c=(df['snapnum']%len(colors)).apply(lambda x: colors[x]));
```
We will leave the calculation of observed halo properties, coordinates, distances, etc. as an exercise for the reader.
| github_jupyter |
<a href="https://colab.research.google.com/github/isb-cgc/Community-Notebooks/blob/master/Notebooks/How_to_compare_tumor_features_with_mutation_data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# ISB-CGC Community Notebooks
Check out more notebooks at our [Community Notebooks Repository](https://github.com/isb-cgc/Community-Notebooks)!
```
Title: Co-analyzing tumor radiomics features with somatic mutations
Author: Fabian Seidl, Boris Aguilar
Created: 2021-06-15
URL: https://github.com/isb-cgc/Community-Notebooks/blob/master/Notebooks/How_to_compare_tumor_features_with_mutation_data.ipynb
Purpose: To demonstrate multi-omics data analysis using table joins in Google BigQuery
Notes:
```
# Overview
This notebook showcases an analysis workflow combining tumor radiomics features with mutation data mostly run in Google BigQuery. In this notebook we will:
> 1. Select and format mutation data from the Glioblastoma Multiforme TCGA project
> 2. Join these mutation data to tumor features extracted from radiomics data
> 3. Calculate t-test statistics for tumors with or without mutations in specific genes
The tumor radiomics feature data we used in this notebook are from Bakas et al. Nature Scientific 2017 (https://www.nature.com/articles/sdata2017117) and the mutation data come from the GDC data housed in ISB-CGC BigQuery tables.
First we initialize our python environment and authenticate our session.
```
from google.colab import auth
import pandas_gbq
import pandas as pd
import numpy
import seaborn
from google.cloud import bigquery
bq_project='<your_google_project_here>'
auth.authenticate_user()
client = bigquery.Client(project=bq_project)
print('Authenticated')
```
# Joining the data
The mutation and feature data we will be using in this analysis are already formatted and stored in a dataset; we will be joining them on the unique TCGA barcodes with an SQL query before further downstream analyses.
```
gdc_project = 'GBM'
mutation_table = 'isb-cgc-idc-collaboration.Analysis.{0}_genes_tidy_v1'.format(gdc_project)
feature_table = 'isb-cgc-idc-collaboration.Analysis.unpivoted_tcga_{0}_radiomicFeatures'.format(gdc_project.lower())
# selecting gene mutation data from a previously generated table
join_query1 = """
WITH
table2 AS (
SELECT
'{0}' as Study, gene as symbol,
REPLACE(case_id,'_','-') AS ParticipantBarcode, present
FROM `{1}`
),""".format(gdc_project, mutation_table)
# selecting tumor volumes
join_query2 = """
table1 AS (
SELECT
'{0}' as Study, feature as symbol,
value as volume, ID as ParticipantBarcode
FROM `{1}`
WHERE
feature LIKE 'VOLUME%'
AND ID IN (SELECT DISTINCT ParticipantBarcode FROM table2)
)""".format(gdc_project, feature_table)
# the join operation for the previous two queries
join_query3 = """
SELECT
n1.Study, n1.symbol as symbol1, n1.volume,
n2.symbol as symbol2, n2.present
FROM table1 AS n1
INNER JOIN table2 AS n2
ON
n1.ParticipantBarcode = n2.ParticipantBarcode
AND n1.Study = n2.Study
AND n2.present = 1
GROUP BY
Study, symbol1, present, symbol2, volume
"""
query_job = client.query( join_query1 + join_query2 + join_query3 )
joined_data = query_job.result().to_dataframe()
joined_data.head(5)
```
We can "pipe" this query directly to the next one by simply appending the next step to our previous ones. With the usage of the `GROUP BY` command we can calculate the SUM() and SUM(squared) values for all cases that have a given mutation.
```
# We can amend the third query from the previous cell as below
sum_query = """,
summ_table AS (
SELECT
n1.Study, n1.symbol as symbol1,
n2.symbol as symbol2,
COUNT( n1.ParticipantBarcode) as n_1,
SUM( n1.volume ) as sumx_1,
SUM( n1.volume * n1.volume ) as sumx2_1
FROM table1 AS n1
INNER JOIN table2 AS n2
ON
n1.ParticipantBarcode = n2.ParticipantBarcode
AND n1.Study = n2.Study
AND n2.present = 1
GROUP BY
Study, symbol1, symbol2
)
""".format(gdc_project, mutation_table, feature_table)
select_all = 'SELECT * FROM summ_table'
#print(join_query1 + join_query2 + sum_query)
sum_query_job = client.query( join_query1 + join_query2 + sum_query + select_all)
sum_table = sum_query_job.result().to_dataframe()
sum_table.head(5)
```
The final step of this workflow is to run t-tests on our groups to calculate whether there are significant differences in expression.
```
statistics_query = """,
statistics AS (
SELECT n1.Study, symbol1, symbol2, n_1,
sumx_1 / n_1 as avg1,
( sumx2_1 - sumx_1*sumx_1/n_1 )/(n_1 -1) as var1,
n_t - n_1 as n_0,
(sumx_t - sumx_1)/(n_t - n_1) as avg0,
(sumx2_t - sumx2_1 - (sumx_t-sumx_1)*(sumx_t-sumx_1)/(n_t - n_1) )/(n_t - n_1 -1 ) as var0
FROM summ_table as n1
LEFT JOIN ( SELECT Study, symbol, COUNT( ParticipantBarcode ) as n_t, SUM( volume ) as sumx_t, SUM( volume*volume ) as sumx2_t
FROM table1
GROUP BY Study, symbol ) as n2
ON symbol1 = symbol AND n1.Study = n2.Study
GROUP BY 1,2,3,4,5,6,7,8,9
having var1 > 0 AND var0 > 0 AND n_1 > 5 AND n_0 > 5
)
SELECT Study, symbol1 as radiomic_feature, symbol2 as Ensembl, n_1 as n1, n_0 as n0,
avg1, avg0,
#ABS(avg1 - avg0)/ SQRT( var1 /n_1 + var0/n_0 ) as t,
`cgc-05-0042.functions.jstat_ttest`(ABS(avg1 - avg0)/ SQRT( var1 /n_1 + var0/n_0 ), n_1+n_0-2, 2) as pvalue,
FROM statistics
ORDER BY pvalue ASC"""
#print(join_query1 + join_query2 + sum_query)
stat_query_job = client.query( join_query1 + join_query2 + sum_query + statistics_query )
statistics_table = stat_query_job.result().to_dataframe()
statistics_table.head(5)
```
We can use these data for further analysis or simply generate a summary plot as below:
```
statistics_table['neglog10'] = -numpy.log10(statistics_table['pvalue'])
bxplt = seaborn.boxplot(x=statistics_table['radiomic_feature'],
y=statistics_table['neglog10'],
data=statistics_table
)
labels = [x.get_text().lower() for x in bxplt.get_xticklabels()]
jnk = bxplt.set_xticklabels(labels, rotation=80, size=10)
jnk = bxplt.set(ylabel='-log10(pvalue)')
# The full final query used to generate the data table
print( join_query1 + join_query2 + sum_query + statistics_query )
```
| github_jupyter |
<h2>In-class transcript from Lecture 8, February 4, 2019</h2>
# Imports and defs for lecture
```
# These are the standard imports for CS 111.
# This list may change as the quarter goes on.
import os
import time
import math
import struct
import numpy as np
import numpy.linalg as npla
import scipy
from scipy import sparse
from scipy import linalg
import scipy.sparse.linalg as spla
import matplotlib.pyplot as plt
from matplotlib import cm
from mpl_toolkits.mplot3d import axes3d
%matplotlib tk
bits = {'0':'0000', '1':'0001', '2':'0010', '3':'0011',
'4':'0100', '5':'0101', '6':'0110', '7':'0111',
'8':'1000', '9':'1001', 'a':'1010', 'b':'1011',
'c':'1100', 'd':'1101', 'e':'1110', 'f':'1111'}
drop = {'0':'0', '1':'1', '2':'2', '3':'3', '4':'4', '5':'5', '6':'6', '7':'7',
'8':'0', '9':'1', 'a':'2', 'b':'3', 'c':'4', 'd':'5', 'e':'6', 'f':'7'}
def double_to_hex(f):
s = hex(struct.unpack('<Q', struct.pack('<d', f))[0])
s = s[2:] # remove the 0x prefix
while len(s) < 16: # pad with zeros
s = '0' + s
return s
def fprint(x):
"""Print a 64-bit floating-point number in various formats.
"""
print('input :', x)
# Cast the input to a 64-bit float
x = np.float64(x)
xhex = double_to_hex(x)
print('as float64: {:.16e}'.format(x))
print('as hex : ' + xhex)
if bits[xhex[0]][0] == '0':
sign = '0 means +'
else:
sign = '1 means -'
print('sign :', sign)
expostr = drop[xhex[0]] + xhex[1:3]
expo = int(expostr, 16)
if expo == 0:
print('exponent :', expostr, 'means zero or denormal')
elif expo == 2047:
print('exponent :', expostr, 'means inf or nan')
else:
print('exponent :', expostr, 'means', expo, '- 1023 =', expo - 1023)
mantissa = '1.'
for i in range(3,16):
mantissa = mantissa + bits[xhex[i]]
print('mantissa :', mantissa)
print()
```
# Lecture starts here
```
# Matrix condition number
A = np.array([[1,1000],[2,2001]])
A
b1 = np.array([1.0,2.0])
b1
x1 = npla.solve(A,b1)
x1
b2 = np.array([1.0,2.001])
x2 = npla.solve(A,b2)
x2
npla.norm(x2-x1)
npla.norm(b1 - A@x2)
npla.cond(A)
# Floating point arithmetic
bits
1.0 + .25
x = 1.0
for i in range(60):
print(x)
print(1+x)
print()
if 1.0 == (1.0+x): break
x = x/2.
fprint(1.0)
fprint(2.)
fprint(1./8.)
fprint(0.)
fprint(1./3.)
fprint(np.inf)
fprint(-np.inf)
fprint(np.nan)
a = 1/10
fprint(a)
fprint(a+a+a+a+a+a+a+a+a+a)
print(np.finfo(np.float64))
2**53 / 10**16
x = 1.0
for i in range(60):
print('x:')
fprint(x)
print('1 + x:')
fprint(1+x)
if 1.0 == (1.0+x): break
x = x/2.
```
| github_jupyter |
```
from __future__ import print_function
import lsst.sims.maf
import numpy as np
import matplotlib.pyplot as plt
%matplotlib inline
import lsst.sims.maf.db as db
import lsst.sims.maf.metrics as metrics
import lsst.sims.maf.slicers as slicers
import lsst.sims.maf.metricBundles as metricBundles
lsst.sims.maf.__version__
```
# Background
Metrics are computed against OpSim simulations, the outputs of which are stored in SQLite databases. The latest simulation outputs are avaialable at http://astro-lsst-01.astro.washington.edu:8082. They're described in https://community.lsst.org/t/january-2020-update-fbs-1-4-runs/4006. In order to run metrics against a simulation, you need to have downloaded the associate .db file. In this notebook, we use the simulation database available at http://astro-lsst-01.astro.washington.edu:8082/fbs_db/footprints/footprint_add_mag_cloudsv1.4_10yrs.db.
```
dbdir = '/sims_maf/'
opsdb = db.OpsimDatabase(dbdir+'fbs_1.5/footprints/footprint_add_mag_cloudsv1.5_10yrs.db')
outDir = 'output_directory' #results from the metrics will be stored here
resultsDb = db.ResultsDb(outDir=outDir)
```
What information is in the OpSim output? You can get the names of the columns in the database by running the `columnNames` method on `opsdb`. The database populates these columns with information about every visit in the simulated cadence. In order to access this information, we will need to issue a query against the database, as described below.
```
opsdb.columnNames
```
As described in the sims_maf_contrib Introduction notebook, to run a metric against OpSim output, you need to:
* specify what you want to measure (referred to as the `metric`), e.g. 5-sigma depth
* specify how you want to project the output (referred to as a `slicer`), e.g. a map of the metric on the sky, or a one-dimensional vector ordered by time of observation, or a single number over the whole survey
* specify the subset of the OpSim output that you want to consider (referred to as a `constraint`), e.g. only output for visits in r-band.
Once these are defined, you collect them into a `MetricBundle`, assemble a dictionary (`bundledict`) describing all of yuor `MetricBundles`, associate your `bundledict` with the input simulation database and output results database through a `MetricBundleGroup`, and use the `MetricBundleGroup` to run your metric. Following the Introduction notebook, here's an example that returns the lowest airmass at which r-band observations were made at each point in the sky, at roughly 1-degree resolution, using the footprint_add_mag_cloudsv1.4_10yrs simulation output:
```
# metric = the "maximum" of the "airmass" for each group of visits in the slicer
metric1 = metrics.MinMetric('airmass')
# slicer = a grouping or subdivision of visits for the simulated survey
# based on their position on the sky (using a Healpix grid)
slicer1 = slicers.HealpixSlicer(nside=64)
# sqlconstraint = the sql query (or 'select') that selects all visits in r band
sqlconstraint = 'filter = "r"'
# MetricBundle = combination of the metric, slicer, and sqlconstraint
minairmassSky = metricBundles.MetricBundle(metric1, slicer1, sqlconstraint)
# Our bundleDict will contain just this one metric
bundleDict = {'minairmassSky': minairmassSky}
# Group our bundleDict with the input simulation (opsdb) and location for string results (outDir, resultsDb)
group = metricBundles.MetricBundleGroup(
bundleDict, opsdb, outDir=outDir, resultsDb=resultsDb)
```
Finally, run the metric against the simulation:
```
group.runAll()
```
MAF can plot the map of the minimum airmass, the histogram, and the power spectrum:
```
group.plotAll(closefigs=False)
```
MAF has lots of metric provided in its BaseMetric class. You can get the list of names of them, as shown in the Writing a New Metric notebook, but they won't be very useful unless you can also see what they do. Fortunately, the name of the module is also stored in the BaseMetric registry:
```
# List of provided metrics
for metric, module in zip(metrics.BaseMetric.registry.keys(), metrics.BaseMetric.registry.values()):
print(metric, module)
```
If you want to see the documentation and code for a particular metric, try e.g.:
```
??lsst.sims.maf.metrics.simpleMetrics.MaxMetric
```
# Science Cases
## Magellanic Clouds
### A 3-D Map of the Magellanic System
We will map structure and detect satellites using MSTO and RR Lyrae as tracers.
We desire:
- Footprint: WFD + SCP, avoiding most crowded parts of Galactic Plane
- Filters: ugriz
- N visits per field: 40 per filter in SCP, + WFD
- Depth: u: 25.6 g: 26.8 r: 26.4 i 25.8 z: 24.9 in SCP, otherwise WFD depth
- Cadence: not strongly constrained, but want spacing for proper motions
#### Metrics
From previous notes:
1. The total number of visits per filter per field in the WFD + SCP area with Dec < -60, avoiding the Galactic Plane; could be aggregated into total or average number of visits of all fields over all filters
Existing metrics in MAF: Nvisits
2. The average total depth per filter in the fields above
Existing metrics in MAF:
- CoaddM5 - Coadded depth per healpix, with benchmark value (depending on filter) subtracted.
- fiveSigmaDepth - is this the single-visit depth? (it's ~2 mags shallower than coaddM5)
3. For a candidate stellar population parametrized by age, metallicity, distance, surface density, and spatial profile, the signal to noise of the population, evaluated as:
T/sqrt(T^2 + B^2),
where T is the number of target stars and B the number of foreground contaminant stars plus the number of background unresolved galaxies
Adriano Pieres has a metric similar to this (T/sqrt(B)).
Need to include: contamination filtering (e.g. S/G separation) and proper motion selection
Existing metrics in MAF:
not much! but possibly relevant to create it, there are:
- skyBrightness
- CoaddM5 / fiveSigmaDepth
4. Given an input distribution of dwarf satellites, the number that would be detected above a set S/N threshold, evaluated through @3.
5. Given an input distribution of low surface brightness streams, the number that would be detected above a set S/N threshold, evaluated through @3.
### A Detailed census of variables and transients
We will target periodic and irregular variables, SN light echoes, giant planet transits, microlensing events, and interstellar scintillation.
We desire:
- Footprint: 9 fields on LMC main body, 3 SMC main body fields
- Filters: ugriz
- N visits per field: u: 50 g: 1300 r: 300 i: 300 z: 30 y: 20
- Depth: will be confusion-limited
- Cadence: Two campaigns of 500 15-second consecutive exposure sequences in g for interstellar scintillation; 300 30-second visits in gri spaced roughly logarithmically over 10 years; uzy visits such that reach confusion depth under excellent image quality conditions
#### Metrics
Primary Science: LSST will enable the identification and locations of variable populations in the Clouds using high time and areal coverage, with 2000 visits in each of 12 fields during the 10 years.
Baisc metric:
- N_visits/300 per main body field in gri
I. Obtain light Curves and periods of all variable objects to Mv=6.5 to map populations
A. Find all eclipsing binaries (periods of hrs-yrs), Mv=0-6.5
Existing metrics in MAF:
- periodicDetectMetric (for P=0.5-2 days, amp=0.05-1 mag, 21 and 24th mag stars) https://github.com/rhiannonlynne/notebooks/blob/master/periodicity%20check.ipynb
Metrics needed
B. Find all outbursting cataclysmic variables
B1. New novae and Recurrent novae (outbursts 7 mag in 1-5 yrs), Mv =-7 to -9
B2. Dwarf novae (outbursts 2-8 mag in weeks to years), Mv = 4.5-5.5
Existing metrics in MAF: ObservationTriplets, PeriodogramPurity, FieldstarCount ( https://github.com/LSST-nonproject/sims_maf_contrib., Lund et al.2016, PASP, 128,025002)
Metrics needed
C. Find all periodic pulsating variables
C1. Delta Scuti (periods 0.5-7 hrs), Mv=0.7, amp 0.1-1 mag
C2. RR Lyrae (periods 0.2-1 day), Mv=0.6, amp 0.2-1.2 mag
C3. Long P variables (periods 60-400 days), Mv=-4, 0.3-5 mag
Existing metrics in MAF:
- periodicDetectMatric (for Delta Scute, RR Lyr) https://github.com/rhiannonlynne/notebooks/blob/master/periodicity%20check.ipynb
Metrics needed:
- coverage for LPV
II. Explore light echoes for past SN to probe the physics of explosions (weeks timescale)
Existing metrics
Metrics needed
III. Use interstellar scintillation to probe invisible baryonic matter (min timescale), need two sets of 500 continuous 15s g filter visits separated by a few months)
Existing metrics
Metrics needed
- The number of 500-visit campaigns divided by two.
The challenge in this experiment is to detect small stochastic flux fluctuations at the minute scale. The better the photometric resolution, the better the sensitivity to turbulent structures. The turbulence is characterized by the parameter Rdiff, the average transverse separation corresponding to a given column density fluctuation. Rdiff increases when the turbulence decreases.
-> We can distinguish two effects when dividing the number of visits:
- Basically, the precision of the measured modulation index from every scintillation signal scales with the square root of the number of visits.
- The ultimate sensitivity is also downgraded. Considering a Kolmogorov-Smirnov test to distinguish between a stable light-curve from a scintillating one with an error risk of 10% (a reasonable cut), we can connect the number of measurements with the maximum turbulent parameter Rdiff. Assuming the best 0.5% LSST photometric precision, decreasing the number of measurements from 500 to 250 would degrade the ultimate sensitivity from Rdiff max = 13000km down to 10000km.
IV. Find giant planet transits in LMC to probe planet formation under low metallicity (hrs)
Existing metrics
Metrics needed
V. Use microlensing events in the Clouds and Galactic halo to probe compact objects (timescales weeks-months)
Existing metrics
Metrics needed
- Largest gap between visits for microlensing
- The probability that the visit distribution was drawn from a logarithmic distribution
| github_jupyter |
# Word2Vec
**Learning Objectives**
1. Compile all steps into one function
2. Prepare training data for Word2Vec
3. Model and Training
4. Embedding lookup and analysis
## Introduction
Word2Vec is not a singular algorithm, rather, it is a family of model architectures and optimizations that can be used to learn word embeddings from large datasets. Embeddings learned through Word2Vec have proven to be successful on a variety of downstream natural language processing tasks.
Note: This notebook is based on [Efficient Estimation of Word Representations in Vector Space](https://arxiv.org/pdf/1301.3781.pdf) and
[Distributed
Representations of Words and Phrases and their Compositionality](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf). It is not an exact implementation of the papers. Rather, it is intended to illustrate the key ideas.
These papers proposed two methods for learning representations of words:
* **Continuous Bag-of-Words Model** which predicts the middle word based on surrounding context words. The context consists of a few words before and after the current (middle) word. This architecture is called a bag-of-words model as the order of words in the context is not important.
* **Continuous Skip-gram Model** which predict words within a certain range before and after the current word in the same sentence. A worked example of this is given below.
You'll use the skip-gram approach in this notebook. First, you'll explore skip-grams and other concepts using a single sentence for illustration. Next, you'll train your own Word2Vec model on a small dataset. This notebook also contains code to export the trained embeddings and visualize them in the [TensorFlow Embedding Projector](http://projector.tensorflow.org/).
Each learning objective will correspond to a _#TODO_ in this student lab notebook -- try to complete this notebook first and then review the [solution notebook](https://github.com/GoogleCloudPlatform/training-data-analyst/blob/master/courses/machine_learning/deepdive2/text_classification/solutions/word2vec.ipynb)
## Skip-gram and Negative Sampling
While a bag-of-words model predicts a word given the neighboring context, a skip-gram model predicts the context (or neighbors) of a word, given the word itself. The model is trained on skip-grams, which are n-grams that allow tokens to be skipped (see the diagram below for an example). The context of a word can be represented through a set of skip-gram pairs of `(target_word, context_word)` where `context_word` appears in the neighboring context of `target_word`.
Consider the following sentence of 8 words.
> The wide road shimmered in the hot sun.
The context words for each of the 8 words of this sentence are defined by a window size. The window size determines the span of words on either side of a `target_word` that can be considered `context word`. Take a look at this table of skip-grams for target words based on different window sizes.
Note: For this tutorial, a window size of *n* implies n words on each side with a total window span of 2*n+1 words across a word.

The training objective of the skip-gram model is to maximize the probability of predicting context words given the target word. For a sequence of words *w<sub>1</sub>, w<sub>2</sub>, ... w<sub>T</sub>*, the objective can be written as the average log probability

where `c` is the size of the training context. The basic skip-gram formulation defines this probability using the softmax function.

where *v* and *v<sup>'<sup>* are target and context vector representations of words and *W* is vocabulary size.
Computing the denominator of this formulation involves performing a full softmax over the entire vocabulary words which is often large (10<sup>5</sup>-10<sup>7</sup>) terms.
The [Noise Contrastive Estimation](https://www.tensorflow.org/api_docs/python/tf/nn/nce_loss) loss function is an efficient approximation for a full softmax. With an objective to learn word embeddings instead of modelling the word distribution, NCE loss can be [simplified](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) to use negative sampling.
The simplified negative sampling objective for a target word is to distinguish the context word from *num_ns* negative samples drawn from noise distribution *P<sub>n</sub>(w)* of words. More precisely, an efficient approximation of full softmax over the vocabulary is, for a skip-gram pair, to pose the loss for a target word as a classification problem between the context word and *num_ns* negative samples.
A negative sample is defined as a (target_word, context_word) pair such that the context_word does not appear in the `window_size` neighborhood of the target_word. For the example sentence, these are few potential negative samples (when `window_size` is 2).
```
(hot, shimmered)
(wide, hot)
(wide, sun)
```
In the next section, you'll generate skip-grams and negative samples for a single sentence. You'll also learn about subsampling techniques and train a classification model for positive and negative training examples later in the tutorial.
## Setup
```
# Use the chown command to change the ownership of repository to user.
!sudo chown -R jupyter:jupyter /home/jupyter/training-data-analyst
!pip install -q tqdm
# You can use any Python source file as a module by executing an import statement in some other Python source file.
# The import statement combines two operations; it searches for the named module, then it binds the
# results of that search to a name in the local scope.
import io
import itertools
import numpy as np
import os
import re
import string
import tensorflow as tf
import tqdm
from tensorflow.keras import Model, Sequential
from tensorflow.keras.layers import Activation, Dense, Dot, Embedding, Flatten, GlobalAveragePooling1D, Reshape
from tensorflow.keras.layers.experimental.preprocessing import TextVectorization
```
This notebook uses TF2.x.
Please check your tensorflow version using the cell below.
```
# Show the currently installed version of TensorFlow
print("TensorFlow version: ",tf.version.VERSION)
SEED = 42
AUTOTUNE = tf.data.experimental.AUTOTUNE
```
### Vectorize an example sentence
Consider the following sentence:
`The wide road shimmered in the hot sun.`
Tokenize the sentence:
```
sentence = "The wide road shimmered in the hot sun"
tokens = list(sentence.lower().split())
print(len(tokens))
```
Create a vocabulary to save mappings from tokens to integer indices.
```
vocab, index = {}, 1 # start indexing from 1
vocab['<pad>'] = 0 # add a padding token
for token in tokens:
if token not in vocab:
vocab[token] = index
index += 1
vocab_size = len(vocab)
print(vocab)
```
Create an inverse vocabulary to save mappings from integer indices to tokens.
```
inverse_vocab = {index: token for token, index in vocab.items()}
print(inverse_vocab)
```
Vectorize your sentence.
```
example_sequence = [vocab[word] for word in tokens]
print(example_sequence)
```
### Generate skip-grams from one sentence
The `tf.keras.preprocessing.sequence` module provides useful functions that simplify data preparation for Word2Vec. You can use the `tf.keras.preprocessing.sequence.skipgrams` to generate skip-gram pairs from the `example_sequence` with a given `window_size` from tokens in the range `[0, vocab_size)`.
Note: `negative_samples` is set to `0` here as batching negative samples generated by this function requires a bit of code. You will use another function to perform negative sampling in the next section.
```
window_size = 2
positive_skip_grams, _ = tf.keras.preprocessing.sequence.skipgrams(
example_sequence,
vocabulary_size=vocab_size,
window_size=window_size,
negative_samples=0)
print(len(positive_skip_grams))
```
Take a look at few positive skip-grams.
```
for target, context in positive_skip_grams[:5]:
print(f"({target}, {context}): ({inverse_vocab[target]}, {inverse_vocab[context]})")
```
### Negative sampling for one skip-gram
The `skipgrams` function returns all positive skip-gram pairs by sliding over a given window span. To produce additional skip-gram pairs that would serve as negative samples for training, you need to sample random words from the vocabulary. Use the `tf.random.log_uniform_candidate_sampler` function to sample `num_ns` number of negative samples for a given target word in a window. You can call the funtion on one skip-grams's target word and pass the context word as true class to exclude it from being sampled.
Key point: *num_ns* (number of negative samples per positive context word) between [5, 20] is [shown to work](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) best for smaller datasets, while *num_ns* between [2,5] suffices for larger datasets.
```
# Get target and context words for one positive skip-gram.
target_word, context_word = positive_skip_grams[0]
# Set the number of negative samples per positive context.
num_ns = 4
context_class = tf.reshape(tf.constant(context_word, dtype="int64"), (1, 1))
negative_sampling_candidates, _, _ = tf.random.log_uniform_candidate_sampler(
true_classes=context_class, # class that should be sampled as 'positive'
num_true=1, # each positive skip-gram has 1 positive context class
num_sampled=num_ns, # number of negative context words to sample
unique=True, # all the negative samples should be unique
range_max=vocab_size, # pick index of the samples from [0, vocab_size]
seed=SEED, # seed for reproducibility
name="negative_sampling" # name of this operation
)
print(negative_sampling_candidates)
print([inverse_vocab[index.numpy()] for index in negative_sampling_candidates])
```
### Construct one training example
For a given positive `(target_word, context_word)` skip-gram, you now also have `num_ns` negative sampled context words that do not appear in the window size neighborhood of `target_word`. Batch the `1` positive `context_word` and `num_ns` negative context words into one tensor. This produces a set of positive skip-grams (labelled as `1`) and negative samples (labelled as `0`) for each target word.
```
# Add a dimension so you can use concatenation (on the next step).
negative_sampling_candidates = tf.expand_dims(negative_sampling_candidates, 1)
# Concat positive context word with negative sampled words.
context = tf.concat([context_class, negative_sampling_candidates], 0)
# Label first context word as 1 (positive) followed by num_ns 0s (negative).
label = tf.constant([1] + [0]*num_ns, dtype="int64")
# Reshape target to shape (1,) and context and label to (num_ns+1,).
target = tf.squeeze(target_word)
context = tf.squeeze(context)
label = tf.squeeze(label)
```
Take a look at the context and the corresponding labels for the target word from the skip-gram example above.
```
print(f"target_index : {target}")
print(f"target_word : {inverse_vocab[target_word]}")
print(f"context_indices : {context}")
print(f"context_words : {[inverse_vocab[c.numpy()] for c in context]}")
print(f"label : {label}")
```
A tuple of `(target, context, label)` tensors constitutes one training example for training your skip-gram negative sampling Word2Vec model. Notice that the target is of shape `(1,)` while the context and label are of shape `(1+num_ns,)`
```
print(f"target :", target)
print(f"context :", context )
print(f"label :", label )
```
### Summary
This picture summarizes the procedure of generating training example from a sentence.

## Lab Task 1: Compile all steps into one function
### Skip-gram Sampling table
A large dataset means larger vocabulary with higher number of more frequent words such as stopwords. Training examples obtained from sampling commonly occuring words (such as `the`, `is`, `on`) don't add much useful information for the model to learn from. [Mikolov et al.](https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf) suggest subsampling of frequent words as a helpful practice to improve embedding quality.
The `tf.keras.preprocessing.sequence.skipgrams` function accepts a sampling table argument to encode probabilities of sampling any token. You can use the `tf.keras.preprocessing.sequence.make_sampling_table` to generate a word-frequency rank based probabilistic sampling table and pass it to `skipgrams` function. Take a look at the sampling probabilities for a `vocab_size` of 10.
```
sampling_table = tf.keras.preprocessing.sequence.make_sampling_table(size=10)
print(sampling_table)
```
`sampling_table[i]` denotes the probability of sampling the i-th most common word in a dataset. The function assumes a [Zipf's distribution](https://en.wikipedia.org/wiki/Zipf%27s_law) of the word frequencies for sampling.
Key point: The `tf.random.log_uniform_candidate_sampler` already assumes that the vocabulary frequency follows a log-uniform (Zipf's) distribution. Using these distribution weighted sampling also helps approximate the Noise Contrastive Estimation (NCE) loss with simpler loss functions for training a negative sampling objective.
### Generate training data
Compile all the steps described above into a function that can be called on a list of vectorized sentences obtained from any text dataset. Notice that the sampling table is built before sampling skip-gram word pairs. You will use this function in the later sections.
```
# Generates skip-gram pairs with negative sampling for a list of sequences
# (int-encoded sentences) based on window size, number of negative samples
# and vocabulary size.
def generate_training_data(sequences, window_size, num_ns, vocab_size, seed):
# Elements of each training example are appended to these lists.
targets, contexts, labels = [], [], []
# Build the sampling table for vocab_size tokens.
# TODO 1a -- your code goes here
# Iterate over all sequences (sentences) in dataset.
for sequence in tqdm.tqdm(sequences):
# Generate positive skip-gram pairs for a sequence (sentence).
positive_skip_grams, _ = tf.keras.preprocessing.sequence.skipgrams(
sequence,
vocabulary_size=vocab_size,
sampling_table=sampling_table,
window_size=window_size,
negative_samples=0)
# Iterate over each positive skip-gram pair to produce training examples
# with positive context word and negative samples.
# TODO 1b -- your code goes here
# Build context and label vectors (for one target word)
negative_sampling_candidates = tf.expand_dims(
negative_sampling_candidates, 1)
context = tf.concat([context_class, negative_sampling_candidates], 0)
label = tf.constant([1] + [0]*num_ns, dtype="int64")
# Append each element from the training example to global lists.
targets.append(target_word)
contexts.append(context)
labels.append(label)
return targets, contexts, labels
```
## Lab Task 2: Prepare training data for Word2Vec
With an understanding of how to work with one sentence for a skip-gram negative sampling based Word2Vec model, you can proceed to generate training examples from a larger list of sentences!
### Download text corpus
You will use a text file of Shakespeare's writing for this tutorial. Change the following line to run this code on your own data.
```
path_to_file = tf.keras.utils.get_file('shakespeare.txt', 'https://storage.googleapis.com/download.tensorflow.org/data/shakespeare.txt')
```
Read text from the file and take a look at the first few lines.
```
with open(path_to_file) as f:
lines = f.read().splitlines()
for line in lines[:20]:
print(line)
```
Use the non empty lines to construct a `tf.data.TextLineDataset` object for next steps.
```
# TODO 2a -- your code goes here
```
### Vectorize sentences from the corpus
You can use the `TextVectorization` layer to vectorize sentences from the corpus. Learn more about using this layer in this [Text Classification](https://www.tensorflow.org/tutorials/keras/text_classification) tutorial. Notice from the first few sentences above that the text needs to be in one case and punctuation needs to be removed. To do this, define a `custom_standardization function` that can be used in the TextVectorization layer.
```
# We create a custom standardization function to lowercase the text and
# remove punctuation.
def custom_standardization(input_data):
lowercase = tf.strings.lower(input_data)
return tf.strings.regex_replace(lowercase,
'[%s]' % re.escape(string.punctuation), '')
# Define the vocabulary size and number of words in a sequence.
vocab_size = 4096
sequence_length = 10
# Use the text vectorization layer to normalize, split, and map strings to
# integers. Set output_sequence_length length to pad all samples to same length.
vectorize_layer = TextVectorization(
standardize=custom_standardization,
max_tokens=vocab_size,
output_mode='int',
output_sequence_length=sequence_length)
```
Call `adapt` on the text dataset to create vocabulary.
```
vectorize_layer.adapt(text_ds.batch(1024))
```
Once the state of the layer has been adapted to represent the text corpus, the vocabulary can be accessed with `get_vocabulary()`. This function returns a list of all vocabulary tokens sorted (descending) by their frequency.
```
# Save the created vocabulary for reference.
inverse_vocab = vectorize_layer.get_vocabulary()
print(inverse_vocab[:20])
```
The vectorize_layer can now be used to generate vectors for each element in the `text_ds`.
```
def vectorize_text(text):
text = tf.expand_dims(text, -1)
return tf.squeeze(vectorize_layer(text))
# Vectorize the data in text_ds.
text_vector_ds = text_ds.batch(1024).prefetch(AUTOTUNE).map(vectorize_layer).unbatch()
```
### Obtain sequences from the dataset
You now have a `tf.data.Dataset` of integer encoded sentences. To prepare the dataset for training a Word2Vec model, flatten the dataset into a list of sentence vector sequences. This step is required as you would iterate over each sentence in the dataset to produce positive and negative examples.
Note: Since the `generate_training_data()` defined earlier uses non-TF python/numpy functions, you could also use a `tf.py_function` or `tf.numpy_function` with `tf.data.Dataset.map()`.
```
sequences = list(text_vector_ds.as_numpy_iterator())
print(len(sequences))
```
Take a look at few examples from `sequences`.
```
for seq in sequences[:5]:
print(f"{seq} => {[inverse_vocab[i] for i in seq]}")
```
### Generate training examples from sequences
`sequences` is now a list of int encoded sentences. Just call the `generate_training_data()` function defined earlier to generate training examples for the Word2Vec model. To recap, the function iterates over each word from each sequence to collect positive and negative context words. Length of target, contexts and labels should be same, representing the total number of training examples.
```
targets, contexts, labels = generate_training_data(
sequences=sequences,
window_size=2,
num_ns=4,
vocab_size=vocab_size,
seed=SEED)
print(len(targets), len(contexts), len(labels))
```
### Configure the dataset for performance
To perform efficient batching for the potentially large number of training examples, use the `tf.data.Dataset` API. After this step, you would have a `tf.data.Dataset` object of `(target_word, context_word), (label)` elements to train your Word2Vec model!
```
BATCH_SIZE = 1024
BUFFER_SIZE = 10000
dataset = tf.data.Dataset.from_tensor_slices(((targets, contexts), labels))
dataset = dataset.shuffle(BUFFER_SIZE).batch(BATCH_SIZE, drop_remainder=True)
print(dataset)
```
Add `cache()` and `prefetch()` to improve performance.
```
dataset = dataset.cache().prefetch(buffer_size=AUTOTUNE)
print(dataset)
```
## Lab Task 3: Model and Training
The Word2Vec model can be implemented as a classifier to distinguish between true context words from skip-grams and false context words obtained through negative sampling. You can perform a dot product between the embeddings of target and context words to obtain predictions for labels and compute loss against true labels in the dataset.
### Subclassed Word2Vec Model
Use the [Keras Subclassing API](https://www.tensorflow.org/guide/keras/custom_layers_and_models) to define your Word2Vec model with the following layers:
* `target_embedding`: A `tf.keras.layers.Embedding` layer which looks up the embedding of a word when it appears as a target word. The number of parameters in this layer are `(vocab_size * embedding_dim)`.
* `context_embedding`: Another `tf.keras.layers.Embedding` layer which looks up the embedding of a word when it appears as a context word. The number of parameters in this layer are the same as those in `target_embedding`, i.e. `(vocab_size * embedding_dim)`.
* `dots`: A `tf.keras.layers.Dot` layer that computes the dot product of target and context embeddings from a training pair.
* `flatten`: A `tf.keras.layers.Flatten` layer to flatten the results of `dots` layer into logits.
With the sublassed model, you can define the `call()` function that accepts `(target, context)` pairs which can then be passed into their corresponding embedding layer. Reshape the `context_embedding` to perform a dot product with `target_embedding` and return the flattened result.
Key point: The `target_embedding` and `context_embedding` layers can be shared as well. You could also use a concatenation of both embeddings as the final Word2Vec embedding.
```
class Word2Vec(Model):
def __init__(self, vocab_size, embedding_dim):
super(Word2Vec, self).__init__()
self.target_embedding = Embedding(vocab_size,
embedding_dim,
input_length=1,
name="w2v_embedding", )
self.context_embedding = Embedding(vocab_size,
embedding_dim,
input_length=num_ns+1)
self.dots = Dot(axes=(3,2))
self.flatten = Flatten()
def call(self, pair):
target, context = pair
we = self.target_embedding(target)
ce = self.context_embedding(context)
dots = self.dots([ce, we])
return self.flatten(dots)
```
### Define loss function and compile model
For simplicity, you can use `tf.keras.losses.CategoricalCrossEntropy` as an alternative to the negative sampling loss. If you would like to write your own custom loss function, you can also do so as follows:
``` python
def custom_loss(x_logit, y_true):
return tf.nn.sigmoid_cross_entropy_with_logits(logits=x_logit, labels=y_true)
```
It's time to build your model! Instantiate your Word2Vec class with an embedding dimension of 128 (you could experiment with different values). Compile the model with the `tf.keras.optimizers.Adam` optimizer.
```
# TODO 3a -- your code goes here
```
Also define a callback to log training statistics for tensorboard.
```
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir="logs")
```
Train the model with `dataset` prepared above for some number of epochs.
```
word2vec.fit(dataset, epochs=20, callbacks=[tensorboard_callback])
```
Tensorboard now shows the Word2Vec model's accuracy and loss.
```
!tensorboard --bind_all --port=8081 --logdir logs
```
Run the following command in **Cloud Shell:**
<code>gcloud beta compute ssh --zone <instance-zone> <notebook-instance-name> --project <project-id> -- -L 8081:localhost:8081</code>
Make sure to replace <instance-zone>, <notebook-instance-name> and <project-id>.
In Cloud Shell, click *Web Preview* > *Change Port* and insert port number *8081*. Click *Change and Preview* to open the TensorBoard.

**To quit the TensorBoard, click Kernel > Interrupt kernel**.
## Lab Task 4: Embedding lookup and analysis
Obtain the weights from the model using `get_layer()` and `get_weights()`. The `get_vocabulary()` function provides the vocabulary to build a metadata file with one token per line.
```
# TODO 4a -- your code goes here
```
Create and save the vectors and metadata file.
```
out_v = io.open('vectors.tsv', 'w', encoding='utf-8')
out_m = io.open('metadata.tsv', 'w', encoding='utf-8')
for index, word in enumerate(vocab):
if index == 0: continue # skip 0, it's padding.
vec = weights[index]
out_v.write('\t'.join([str(x) for x in vec]) + "\n")
out_m.write(word + "\n")
out_v.close()
out_m.close()
```
Download the `vectors.tsv` and `metadata.tsv` to analyze the obtained embeddings in the [Embedding Projector](https://projector.tensorflow.org/).
```
try:
from google.colab import files
files.download('vectors.tsv')
files.download('metadata.tsv')
except Exception as e:
pass
```
## Next steps
This tutorial has shown you how to implement a skip-gram Word2Vec model with negative sampling from scratch and visualize the obtained word embeddings.
* To learn more about word vectors and their mathematical representations, refer to these [notes](https://web.stanford.edu/class/cs224n/readings/cs224n-2019-notes01-wordvecs1.pdf).
* To learn more about advanced text processing, read the [Transformer model for language understanding](https://www.tensorflow.org/tutorials/text/transformer) tutorial.
* If you’re interested in pre-trained embedding models, you may also be interested in [Exploring the TF-Hub CORD-19 Swivel Embeddings](https://www.tensorflow.org/hub/tutorials/cord_19_embeddings_keras), or the [Multilingual Universal Sentence Encoder](https://www.tensorflow.org/hub/tutorials/cross_lingual_similarity_with_tf_hub_multilingual_universal_encoder)
* You may also like to train the model on a new dataset (there are many available in [TensorFlow Datasets](https://www.tensorflow.org/datasets)).
| github_jupyter |
```
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python Docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the read-only "../input/" directory
# For example, running this (by clicking run or pressing Shift+Enter) will list all files under the input directory
import os
for dirname, _, filenames in os.walk('.'):
for filename in filenames:
print(os.path.join(dirname, filename))
# You can write up to 20GB to the current directory (/kaggle/working/) that gets preserved as output when you create a version using "Save & Run All"
# You can also write temporary files to /kaggle/temp/, but they won't be saved outside of the current session
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
columns = [
'neighbourhood_group', 'room_type', 'latitude', 'longitude',
'minimum_nights', 'number_of_reviews','reviews_per_month',
'calculated_host_listings_count', 'availability_365',
'price'
]
df = pd.read_csv('AB_NYC_2019.csv', usecols=columns)
df.reviews_per_month = df.reviews_per_month.fillna(0)
# df.isna().sum()
from sklearn.model_selection import train_test_split
df_full_train, df_test = train_test_split(df[columns], test_size=0.2, random_state=1)
df_train, df_val = train_test_split(df_full_train, test_size=0.25, random_state=1)
df_train = df_train.reset_index(drop=True)
df_val = df_val.reset_index(drop=True)
df_test = df_test.reset_index(drop=True)
y_train = np.log1p(df_train.price.values)
y_val = np.log1p(df_val.price.values)
y_test = np.log1p(df_test.price.values)
del df_train["price"]
del df_val["price"]
del df_test["price"]
sns.histplot(y_train, bins=50, color='red')
# Q1
from sklearn.feature_extraction import DictVectorizer
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import roc_auc_score
train_dict = df_train.to_dict(orient='records')
dv = DictVectorizer(sparse=False)
X_train = dv.fit_transform(train_dict)
# dv.get_feature_names()
dt = DecisionTreeRegressor(max_depth=1)
dt.fit(X_train, y_train)
# val_dict = df_val.to_dict(orient='records')
# X_val = dv.transform(val_dict)
# y_pred = dt.predict(X_val)
# roc_auc_score(y_val, y_pred)
#print decision tree
from sklearn.tree import export_text
print(export_text(dt, feature_names=dv.get_feature_names()))
# Q2
# Random Forest of decision trees
from sklearn.feature_extraction import DictVectorizer
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import roc_auc_score
train_dict = df_train.to_dict(orient='records')
dv = DictVectorizer(sparse=False)
X_train = dv.fit_transform(train_dict)
rf = RandomForestRegressor(n_estimators=10, random_state=1, n_jobs=-1)
rf.fit(X_train, y_train)
val_dict = df_val.to_dict(orient='records')
X_val = dv.transform(val_dict)
y_pred = rf.predict(X_val)
from sklearn.metrics import mean_squared_error
rmse = mean_squared_error(y_val, y_pred , squared=False)
rmse
# Q3
from IPython.display import display
import matplotlib.pyplot as plt
scores = []
for n in range(10, 201, 10):
rf = RandomForestRegressor(n_estimators=n, random_state=1, n_jobs=-1)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_val)
rmse = mean_squared_error(y_val, y_pred , squared=False)
scores.append((n,rmse))
df_scores = pd.DataFrame(scores, columns=['n_estimators','rmse'])
display(df_scores)
plt.plot(df_scores.n_estimators, df_scores.rmse)
# Q3
from IPython.display import display
import matplotlib.pyplot as plt
scores = []
for d in [10, 15, 20, 25]:
for n in range(10, 201, 10):
print("Processing d:%s, n:%s"%(d,n))
rf = RandomForestRegressor(n_estimators=n, max_depth=d, random_state=1, n_jobs=-1)
rf.fit(X_train, y_train)
y_pred = rf.predict(X_val)
rmse = mean_squared_error(y_val, y_pred , squared=False)
scores.append((d, n,rmse))
df_scores = pd.DataFrame(scores, columns=["max_depth",'n_estimators','rmse'])
display(df_scores)
for d in [10, 15, 20, 25]:
df_sub = df_scores[df_scores.max_depth==d]
plt.plot(df_sub.n_estimators, df_sub.rmse, label="d=%s"%d)
plt.legend()
# Q5
rf = RandomForestRegressor(n_estimators=10, max_depth=20, random_state=1, n_jobs=-1)
rf.fit(X_train, y_train)
# y_pred = rf.predict(X_val)
# rmse = mean_squared_error(y_val, y_pred , squared=False)
# rmse
# zip(dv.get_feature_names(), rf.feature_importances_.tolist())
d = pd.DataFrame(dv.get_feature_names(), rf.feature_importances_.tolist())
d
# Q6
!pip install xgboost
import xgboost as xgb
features = dv.get_feature_names()
dtrain = xgb.DMatrix(X_train, label=y_train, feature_names=features)
dval = xgb.DMatrix(X_val, label=y_val, feature_names=features)
def train(eta):
xgb_params = {
'eta': eta,
'max_depth': 6,
'min_child_weight': 1,
'objective': 'reg:squarederror',
'nthread': 8,
'seed': 1,
'verbosity': 1,
}
xgb_model = xgb.train(xgb_params, dtrain, num_boost_round=100)
y_pred = xgb_model.predict(dval)
# roc_auc_score(y_val, y_pred)
from sklearn.metrics import mean_squared_error
rmse = mean_squared_error(y_val, y_pred , squared=False)
print("rmse=%s, eta=%s"%(rmse, eta))
for e in [0.3, 0.1, 0.01]:
train(e)
```
| github_jupyter |
```
import math
import scipy
import numpy as np
import pandas as pd
import networkx as nx
import matplotlib.pyplot as plt
import seaborn as sns
%matplotlib inline
sns.set()
from scipy.stats import pearsonr
import osmnx as ox
ox.config(log_console=True, use_cache=True)
ox.__version__
def create_pdf_network():
graph = nx.Graph()
graph.add_edge(1, 4, weight=1)
graph.add_edge(1, 2, weight=1)
graph.add_edge(4, 2, weight=1)
graph.add_edge(2, 3, weight=1)
graph.add_edge(4, 3, weight=1)
graph.add_edge(3, 6, weight=1)
graph.add_edge(3, 7, weight=1)
graph.add_edge(3, 5, weight=1)
graph.add_edge(6, 7, weight=1)
graph = graph.to_undirected()
graph.remove_edges_from(graph.selfloop_edges())
return graph
def average_degree(graph):
degrees = graph.degree()
return np.mean([*dict(degrees).values()])
def visualize2(G, big=False):
if big:
plt.figure(figsize=(12, 8))
labels = G.nodes()
pos=nx.spring_layout(G)
# nodes
nx.draw_networkx_nodes(G,pos,node_size=700)
# edges
nx.draw_networkx_edges(G,pos,
width=6,alpha=0.5,edge_color='b',style='dashed')
# labels
nx.draw_networkx_labels(G,pos,font_size=20,font_family='sans-serif')
plt.axis('off')
plt.show() # display
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q1
## a) Degree distribution,
```
def degree_distribution(G):
vk = dict(G.degree())
vk = list(vk.values()) # we get only the degree values
vk = np.array(vk)
maxk = np.max(vk)
mink = np.min(vk)
kvalues= np.arange(0,maxk+1) # possible values of k
Pk = np.zeros(maxk+1) # P(k)
for k in vk:
Pk[k] = Pk[k] + 1
Pk = Pk/sum(Pk) # the sum of the elements of P(k) must to be equal to one
return kvalues, Pk
graph = create_pdf_network()
def plot_degree_dist(graph, use_log_scale=False, skip_plot=False, ks=None, Pk=None):
# Calculate the data we want if the user doesn't give it to us
if ks is None and Pk is None:
ks, Pk = degree_distribution(graph)
# In case we want only the degree distribution
if not skip_plot:
fig = plt.subplot(1,1,1)
if use_log_scale:
fig.set_xscale('log')
fig.set_yscale('log')
plt.plot(ks,Pk,'bo')
plt.xlabel("k", fontsize=20)
plt.ylabel("P(k)", fontsize=20)
plt.title("Degree distribution", fontsize=20)
#plt.grid(True)
plt.show(True)
return ks, Pk
plot_degree_dist(graph)
```
## b) Local clustering coefficient
```
vcc = []
for i in graph.nodes():
vcc.append(nx.clustering(graph, i))
vcc= np.array(vcc)
print('Clustering of all nodes:', vcc)
```
## c) Transitivity
```
CC = (nx.transitivity(graph))
print("Transitivity = ","%3.4f"%CC)
```
## d) Distance matrix,
```
nx.floyd_warshall_numpy(graph, nodelist=sorted(graph.nodes))
```
## e) Entropy of the degree distribution
```
def shannon_entropy(G):
k,Pk = degree_distribution(G)
H = 0
for p in Pk:
if(p > 0):
H = H - p*math.log(p, 2)
return H
shannon_entropy(graph)
```
## f) Second moment of the degree distribution.
```
def moment_of_degree_distribution(G,m):
M = 0
N = len(G)
for i in G.nodes:
M = M + G.degree(i)**m
M = M/N
return M
k2 = moment_of_degree_distribution(graph, 2)
print("Second moment of the degree distribution = ", k2)
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q2
```
def graph_to_undirected(graph, sum_edges=True, testing=False):
"""Converts a graph to it's undirecte version.
If not sum_edges, we simply use the value of the 'last' edge when iterating.
Otherwise, we sum the value of the incoming/outcoming edge.
"""
if sum_edges:
new_g = nx.Graph()
new_g.add_edges_from(graph.edges, weight=0)
for u, v in graph.edges:
new_g[u][v]['weight'] += graph[u][v]['weight']
new_g
else:
new_g = graph.to_undirected()
# Quick test to show our implementation is correct
if testing:
graph = nx.DiGraph()
graph.add_edge(2, 1, weight=1)
graph.add_edge(1, 2, weight=2)
print("Before:")
visualize(graph)
print("After:")
visualize(graph_to_undirected(graph))
return new_g
def graph_to_unweighted(graph, min_weight=1, testing=False):
"""Converts a graph to it's unweighted version.
Any edge with weight >= min_weight is included.
"""
Gnew = nx.Graph()
for (u,v,w) in graph.edges(data=True):
if w['weight'] >= min_weight :
Gnew.add_edge(u, v)
if testing:
graph = nx.Graph()
graph.add_edge(1, 2, weight=1)
graph.add_edge(2, 3, weight=2)
graph.add_edge(3, 4, weight=3)
print("Before:")
visualize(graph)
print("After:")
visualize(graph_to_unweighted(graph, 3))
return Gnew
def read_edges(filename, digraph=True):
"""Reads an edge list dataset"""
# Read the directed network
graph = nx.read_weighted_edgelist(filename, nodetype=int,
create_using=nx.DiGraph)
return graph
# Run Lesmis network
g_lesmis = read_edges("data/nets/lesmis.txt")
print("Weighted Directed Lesmis: ")
visualize2(g_lesmis, big=True)
print("Unweighted Undirected Lesmis: ")
g_lesmis = graph_to_undirected(g_lesmis, sum_edges=True)
visualize2(graph_to_unweighted(g_lesmis), big=True)
# Run Florida network
g_florida = read_edges("data/nets/eco-foodweb-baywet.edges")
print("Weighted Directed Florida: ")
visualize2(g_florida, big=True)
g_florida = graph_to_undirected(g_florida, sum_edges=True)
print("Unweighted Undirected Florida: ")
visualize2(graph_to_unweighted(g_florida), big=True)
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q3
```
# Read the data
G = read_edges("data/nets/lesmis.txt")
G = graph_to_undirected(G, sum_edges=True)
G = graph_to_unweighted(G)
# let networkx return the adjacency matrix A
A = nx.adj_matrix(G)
A = A.todense()
A = np.array(A, dtype = np.float64)
# define walk length and starting idx
# we start with a random point
walk_length = 100*len(G.nodes)
repetitions=10
visited = np.zeros(len(G.nodes))
for i in range(repetitions):
# Use this to start with the node with the highest amount of neighbours
#current = A.sum(axis=0).argmax()
current = np.random.randint(len(G.nodes))
for _ in range(walk_length):
# Get the list of possible nodes at current step
possible_nodes = np.flatnonzero(A[current])
# Go the next node
next_node = np.random.choice(possible_nodes)
visited[next_node] += 1
#visited.append(next_node)
current = next_node
visited
pearsonr(visited, A.sum(axis=0))
```
Sim, está altamente correlacionado!
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q4
## a)
```
graph = create_pdf_network()
def power_transition_matrix(G, power):
transition_matrix = nx.google_matrix(G, alpha=1)
return transition_matrix**power
power_transition_matrix(graph, 1000)
```
Ele converge!
## b)
$$ ((Aij)^3, Dij) $$ for i, j= 1, . . . , N
```
def create_scatter_plot(power_matrix, dists, plot=True):
x = []
y = []
for i in range(len(graph.nodes)):
for j in range(len(graph.nodes)):
x.append(power_matrix[i, j])
y.append(dists[i, j])
if plot:
sns.scatterplot(x, y)
plt.show()
return x, y
power_matrix = power_transition_matrix(graph, 3)
dists = nx.floyd_warshall_numpy(graph, nodelist=sorted(graph.nodes))
create_scatter_plot(power_matrix, dists);
```
## c)
```
corrs = []
limit = 100
dists = nx.floyd_warshall_numpy(graph, nodelist=sorted(graph.nodes))
for i in range(1, limit):
power_matrix = power_transition_matrix(graph, i)
x, y = create_scatter_plot(power_matrix, dists, plot=False);
corrs.append(pearsonr(x, y)[0])
sns.scatterplot(list(range(1, limit)), corrs)
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q5
$p_k = \sum\limits_{q = k + 1}^\infty {p_q } $
```
ks, Pk = plot_degree_dist(graph, skip_plot=True)
cumulative_Pk = np.cumsum(Pk)
def plot_cumulative_degree_dist(ks, cumulative_Pk, use_log_scale=True,
title="Cumulative Degree distribution"):
fig = plt.figure(figsize=(8, 6))
ax = fig.add_subplot(111)
plt.plot(ks, cumulative_Pk)
if use_log_scale:
ax.set_xscale('log')
ax.set_yscale('log')
plt.xlabel("k", fontsize=20)
plt.ylabel("P(k)", fontsize=20)
plt.title(title, fontsize=20)
plt.show()
plot_cumulative_degree_dist(ks, cumulative_Pk)
```
How is the coefficient of this distribution related to the power law degree distribution?
Se tivermos uma rede que tenha uma power law degree distribution que segue uma exponencial ($ p_k \sim k^{ - \gamma }$), então sua versão cumulativa definida como $p_k = \sum\limits_{q = k + 1}^\infty {p_q } $ irá seguir $p_k \sim k^{ - \gamma + 1} $,
Ou seja, quanto mais se aproximar de uma exponencial com um coefficiente alto, maior o efeito sobre o final da distribuição acumulativa
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q6
```
# Read all the data we will use now and in the future
all_graphs = {}
all_graphs["E-road"] = nx.read_edgelist("data/euroroad.txt")
all_graphs["Authors papers"] = nx.read_gml("data/netscience.gml")
all_graphs["Facebook"] = nx.read_edgelist("data/facebook_combined.txt")
all_graphs["Hamster"] = nx.read_edgelist("data/out.petster-friendships-hamster-uniq",
comments="%")
all_graphs["Astro"] = nx.read_edgelist("data/out.ca-AstroPh",
comments="%")
def get_biggest_component(graph, testing=False):
"""
Fast function taken from:
https://networkx.github.io/documentation/networkx-1.10/reference/generated/networkx.algorithms.components.connected.connected_component_subgraphs.html
If testing is True, use the smallest instead of the biggest connected component
"""
if testing:
return min(nx.connected_component_subgraphs(graph), key=len)
else:
return max(nx.connected_component_subgraphs(graph), key=len)
skip_graphs = ["Astro"]
for graph_name, graph in all_graphs.items():
if graph_name in skip_graphs:
continue
biggest_connect_component = get_biggest_component(graph)
ks, Pk = degree_distribution(biggest_connect_component)
cumulative_Pk = np.cumsum(Pk)
plot_cumulative_degree_dist(ks, cumulative_Pk, "CDF " + graph_name)
```
Comparando os resultados obtidos com os de uma rede que sabemos ser scale-free:
```
scale_free = graph_to_undirected(nx.scale_free_graph(100), sum_edges=False)
biggest_connect_component = get_biggest_component(scale_free)
ks, Pk = degree_distribution(biggest_connect_component)
cumulative_Pk = np.cumsum(Pk)
plot_cumulative_degree_dist(ks, cumulative_Pk, "CDF " + "scale free")
```
Então, percebemos que as redes acima, são de fato scale-free
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q7
```
# Define all the functions we wil use now and in the future
all_funcs = {
"Num nodes": lambda x: len(x.nodes),
"Avg degree": average_degree,
"2nd Momentum": lambda x: moment_of_degree_distribution(x, 2),
"Avg clust coef": nx.average_clustering,
"Diameter": nx.diameter,
"Avg shortest path": nx.average_shortest_path_length,
"Transivity": nx.transitivity,
}
def run_experiment(use_funcs, skip_graphs, testing=False):
"""
In case we are testing, use the smallest connected component instead
of the biggest.
"""
funcs = {x:y for x, y in all_funcs.items() if x in use_funcs}
graphs = {x:y for x, y in all_graphs.items() if x not in skip_graphs}
results = pd.DataFrame(columns=["Name"] + [*funcs.keys()])
for graph_name, graph in graphs.items():
connected_graph = get_biggest_component(graph, testing=testing)
data = [graph_name]
for name, func in funcs.items():
data.append(func(connected_graph))
results.loc[len(results)] = data
return results
run_experiment(["Avg clust coef", "Transivity"], skip_graphs=["Authors papers"])
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q8
```
def distribution_shortest_path(G, title="Distribution of the geodesic distances"):
if nx.is_connected(G) == True:
vl = []
N = len(G.nodes)
diam = nx.diameter(G)
# Build matrix of distances (D)
D = np.zeros(shape=(N, N))
for i in np.arange(0,N):
for j in np.arange(i+1, N):
if(i != j):
aux = nx.shortest_path(G,i,j)
dij = len(aux)-1
D[i][j] = dij
D[j][i] = dij
vl.append(dij)
x = range(0,diam+1)
plt.hist(vl, bins=x, normed=True)
plt.ylabel("P(l)", fontsize=15)
plt.xlabel("Shortest path length (l)", fontsize=15)
plt.title(title, fontsize=20)
plt.grid(True)
plt.show(True)
else:
print("-"*10 + "ERROR!" + "-"*10)
print("The graph has more than one connected component")
graphs = {x:y for x, y in all_graphs.items() if x != "Authors papers"}
for graph_name, graph in graphs.items():
connected_graph = get_biggest_component(graph)
connected_graph = nx.convert_node_labels_to_integers(connected_graph)
distribution_shortest_path(connected_graph, title=graph_name)
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q9
(i) number of nodes, (ii) average degree, (iii) second moment of thedegree distribution, (iv) average clustering coefficient, (v) transitivity, (vi) average shortest pathlength, (vii) diameter
```
use_funcs = {x:y for x, y in all_funcs.items()}
run_experiment(use_funcs, ["Astro"])
```
- - -
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
<br>
# Q10
```
def complexity(G):
k2 = moment_of_degree_distribution(G,2)
k1 = moment_of_degree_distribution(G,1)
return round((k2/k1), 3)
def unmultigraph(graph):
G = nx.Graph()
for u,v,data in graph.edges(data=True):
w = data['length'] if 'length' in data else 1.0
if G.has_edge(u,v):
G[u][v]['weight'] += w
else:
G.add_edge(u, v, weight=w)
G = graph_to_undirected(G, sum_edges=True)
G = graph_to_unweighted(G)
return G
```
### Motuca (PR)
```
motuca = ox.graph_from_place('Motuca', network_type='drive', simplify=False)
print('Complexity: ', end='')
print(complexity(motuca))
print('Average shortest path length: ', end='')
print(nx.average_shortest_path_length(motuca))
print('Average clustering coefficient: ', end='')
motuca = unmultigraph(motuca)
print(nx.average_clustering(motuca))
```
### Mombuca (PR)
```
mombuca = ox.graph_from_place('Mombuca', network_type='drive', simplify=False)
print('Complexity: ', end='')
print(complexity(mombuca))
print('Average shortest path length: ', end='')
print(nx.average_shortest_path_length(mombuca))
print('Average clustering coefficient: ', end='')
mombuca = unmultigraph(mombuca)
print(nx.average_clustering(mombuca))
```
### Guatapara (PR)
```
guatapara = ox.graph_from_place('Guatapara', network_type='drive', simplify=False)
print('Complexity: ', end='')
print(complexity(guatapara))
print('Average shortest path length: ', end='')
print(nx.average_shortest_path_length(guatapara))
print('Average clustering coefficient: ', end='')
guatapara = unmultigraph(guatapara)
print(nx.average_clustering(guatapara))
```
| github_jupyter |
<img src="./result/divvy.png" alt="Drawing" align="left" style="width: 400px;"/>
# Chicago [Divvy](https://www.divvybikes.com/) Bicycle Sharing Data Analysis and Modeling
In this notebook, I conducted a series of exploratory data analysis and modeling on [Chicago Divvy bicycle sharing data](https://www.divvybikes.com/system-data). The goal of this project includes:
* Visualize the bicycle sharing data
* Try to find some interesting pheonona behind the data
* Try to model the bicycle needs behind the data
```
# import necessary packages
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import gc, os
from utils import query_weather, merge
%matplotlib inline
```
# I. Data Preprocessing
### Weather information
Among all the external information, weather has a huge influence on the usage of bicycle in Chicago. In this project, I first write a wrapper to download the weather information from [Weather Underground](https://www.wunderground.com/).
```
# Query data in different years
# You can get free keys from https://www.wunderground.com/
keys = ['***************', '***************', '***************',
'***************', '***************']
years = [2013, 2014, 2015, 2016, 2017]
for key, year in zip(keys, years):
path = './data/weather_' + str(year) + '.csv'
if os.path.isfile(path):
continue
df, _ = query_weather(key=key, year=year, state='IL', area='Chicago')
df.to_csv(path, index=False)
print('File saved:\t', path)
```
### Combine bicycle, station, and weather data
```
if not os.path.isfile('./data/data_raw.csv'):
# year 2013
# load weather information
weather_2013 = pd.read_csv('./data/weather_2013.csv', parse_dates=['date'])
trip_2013 = pd.read_csv('./data/2013/Divvy_Trips_2013.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2013 = pd.read_csv('./data/2013/Divvy_Stations_2013.csv')
# merge information
merged_2013 = merge(trip_2013, station_2013, weather_2013)
# year 2014, Q1 and Q2
# load weather information
weather_2014 = pd.read_csv('./data/weather_2014.csv', parse_dates=['date'])
trip_2014_Q1Q2 = pd.read_csv('./data/2014_Q1Q2/Divvy_Trips_2014_Q1Q2.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2014_Q1Q2 = pd.read_excel('./data/2014_Q1Q2/Divvy_Stations_2014-Q1Q2.xlsx')
# merge information
merged_2014_Q1Q2 = merge(trip_2014_Q1Q2, station_2014_Q1Q2, weather_2014)
# year 2014, Q3 and Q4
trip_2014_Q3_07 = pd.read_csv('./data/2014_Q3Q4/Divvy_Trips_2014-Q3-07.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2014_Q3_0809 = pd.read_csv('./data/2014_Q3Q4/Divvy_Trips_2014-Q3-0809.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2014_Q4 = pd.read_csv('./data/2014_Q3Q4/Divvy_Trips_2014-Q4.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2014_Q3Q4 = pd.read_csv('./data/2014_Q3Q4/Divvy_Stations_2014-Q3Q4.csv')
# merge information
merged_2014_Q3_07 = merge(trip_2014_Q3_07, station_2014_Q3Q4, weather_2014)
merged_2014_Q3_0809 = merge(trip_2014_Q3_0809, station_2014_Q3Q4, weather_2014)
merged_2014_Q4 = merge(trip_2014_Q4, station_2014_Q3Q4, weather_2014)
# year 2015, Q1 and Q2
# load weather information
weather_2015 = pd.read_csv('./data/weather_2015.csv', parse_dates=['date'])
trip_2015_Q1 = pd.read_csv('./data/2015_Q1Q2/Divvy_Trips_2015-Q1.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2015_Q2 = pd.read_csv('./data/2015_Q1Q2/Divvy_Trips_2015-Q2.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2015 = pd.read_csv('./data/2015_Q1Q2/Divvy_Stations_2015.csv')
# merge information
merged_2015_Q1 = merge(trip_2015_Q1, station_2015, weather_2015)
merged_2015_Q2 = merge(trip_2015_Q2, station_2015, weather_2015)
# year 2015, Q3 and Q4
trip_2015_Q3_07 = pd.read_csv('./data/2015_Q3Q4/Divvy_Trips_2015_07.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2015_Q3_08 = pd.read_csv('./data/2015_Q3Q4/Divvy_Trips_2015_08.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2015_Q3_09 = pd.read_csv('./data/2015_Q3Q4/Divvy_Trips_2015_09.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2015_Q4 = pd.read_csv('./data/2015_Q3Q4/Divvy_Trips_2015_Q4.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
# merge information
merged_2015_Q3_07 = merge(trip_2015_Q3_07, station_2015, weather_2015)
merged_2015_Q3_08 = merge(trip_2015_Q3_08, station_2015, weather_2015)
merged_2015_Q3_09 = merge(trip_2015_Q3_09, station_2015, weather_2015)
merged_2015_Q4 = merge(trip_2015_Q4, station_2015, weather_2015)
# year 2016, Q1 and Q2
# load weather information
weather_2016 = pd.read_csv('./data/weather_2016.csv', parse_dates=['date'])
trip_2016_Q1 = pd.read_csv('./data/2016_Q1Q2/Divvy_Trips_2016_Q1.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2016_Q2_04 = pd.read_csv('./data/2016_Q1Q2/Divvy_Trips_2016_04.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2016_Q2_05 = pd.read_csv('./data/2016_Q1Q2/Divvy_Trips_2016_05.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
trip_2016_Q2_06 = pd.read_csv('./data/2016_Q1Q2/Divvy_Trips_2016_06.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2016_Q1Q2 = pd.read_csv('./data/2016_Q1Q2/Divvy_Stations_2016_Q1Q2.csv')
# merge information
merged_2016_Q1 = merge(trip_2016_Q1, station_2016_Q1Q2, weather_2016)
merged_2016_Q2_04 = merge(trip_2016_Q2_04, station_2016_Q1Q2, weather_2016)
merged_2016_Q2_05 = merge(trip_2016_Q2_05, station_2016_Q1Q2, weather_2016)
merged_2016_Q2_06 = merge(trip_2016_Q2_06, station_2016_Q1Q2, weather_2016)
# year 2016, Q3 and Q4
trip_2016_Q3 = pd.read_csv('./data/2016_Q3Q4/Divvy_Trips_2016_Q3.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2016_Q3 = pd.read_csv('./data/2016_Q3Q4/Divvy_Stations_2016_Q3.csv')
trip_2016_Q4 = pd.read_csv('./data/2016_Q3Q4/Divvy_Trips_2016_Q4.csv', low_memory=False,
parse_dates=['starttime', 'stoptime'])
station_2016_Q4 = pd.read_csv('./data/2016_Q3Q4/Divvy_Stations_2016_Q4.csv')
# merge information
merged_2016_Q3 = merge(trip_2016_Q3, station_2016_Q3, weather_2016)
merged_2016_Q4 = merge(trip_2016_Q4, station_2016_Q4, weather_2016)
# year 2017, Q1 and Q2
# load weather information
weather_2017 = pd.read_csv('./data/weather_2017.csv', parse_dates=['date'])
trip_2017_Q1 = pd.read_csv('./data/2017_Q1Q2/Divvy_Trips_2017_Q1.csv', low_memory=False,
parse_dates=['start_time', 'end_time'])
trip_2017_Q1.rename(columns={'start_time': 'starttime', 'end_time': 'stoptime'}, inplace=True)
trip_2017_Q2 = pd.read_csv('./data/2017_Q1Q2/Divvy_Trips_2017_Q2.csv', low_memory=False,
parse_dates=['start_time', 'end_time'])
trip_2017_Q2.rename(columns={'start_time': 'starttime', 'end_time': 'stoptime'}, inplace=True)
station_2017_Q1Q2 = pd.read_csv('./data/2017_Q1Q2/Divvy_Stations_2017_Q1Q2.csv')
# merge information
merged_2017_Q1 = merge(trip_2017_Q1, station_2017_Q1Q2, weather_2017)
merged_2017_Q2 = merge(trip_2017_Q2, station_2017_Q1Q2, weather_2017)
# year 2017, Q3 and Q4
trip_2017_Q3 = pd.read_csv('./data/2017_Q3Q4/Divvy_Trips_2017_Q3.csv', low_memory=False,
parse_dates=['start_time', 'end_time'])
trip_2017_Q3.rename(columns={'start_time': 'starttime', 'end_time': 'stoptime'}, inplace=True)
trip_2017_Q4 = pd.read_csv('./data/2017_Q3Q4/Divvy_Trips_2017_Q4.csv', low_memory=False,
parse_dates=['start_time', 'end_time'])
trip_2017_Q4.rename(columns={'start_time': 'starttime', 'end_time': 'stoptime'}, inplace=True)
station_2017_Q3Q4 = pd.read_csv('./data/2017_Q3Q4/Divvy_Stations_2017_Q3Q4.csv')
# merge information
merged_2017_Q3 = merge(trip_2017_Q3, station_2017_Q3Q4, weather_2017)
merged_2017_Q4 = merge(trip_2017_Q4, station_2017_Q3Q4, weather_2017)
# concatenate and save the merged data
objs = [merged_2013, merged_2014_Q1Q2, merged_2014_Q3_07, merged_2014_Q3_0809, merged_2014_Q4,
merged_2015_Q1, merged_2015_Q2, merged_2015_Q3_07, merged_2015_Q3_08, merged_2015_Q3_09,
merged_2015_Q4, merged_2016_Q1, merged_2016_Q2_04, merged_2016_Q2_05, merged_2016_Q2_06,
merged_2016_Q3, merged_2016_Q4, merged_2017_Q1, merged_2017_Q2, merged_2017_Q3, merged_2017_Q4]
data = pd.concat(objs, axis=0)
data.to_csv('./data/data_raw.csv', index=False)
_ = gc.collect()
```
### Clean the data
* Exclude trip duration that is less than 2 minutes and more than 1 hour
* Exclude trip with missing latitude_start, longitude_start, dpcapacity_start, latitude_end, longitude_end, dpcapacity_end
* Exclude trip without gender information
* Extract year, month, and day information
* Keep useful weather information
```
# read data from ./data/
if not os.path.isfile('./data/data.csv'):
data_raw = pd.read_csv('./data/data_raw.csv', parse_dates=['starttime', 'stoptime'])
data = data_raw[(data_raw['tripduration'] >= 120) & (data_raw['tripduration'] <= 3600)]
data = data[~data['latitude_start'].isnull()]
data = data[~data['latitude_end'].isnull()]
data = data[~data['gender'].isnull()]
data = data[~data['humidity'].isnull()]
# extract detailed time information
date = np.array(list(map(lambda x: (x.year, x.month, x.week, x.dayofweek, x.hour), data['starttime'])))
data['year'] = date[:, 0]
data['month'] = date[:, 1]
data['week'] = date[:, 2]
data['day'] = date[:, 3]
data['hour'] = date[:, 4]
data = data[data['year'] > 2013]
# transform trip duration into minutes
data['tripduration'] = data['tripduration'] / 60.0
# extract the weather events information
def events_map(event):
maps = {'tstorms': 'tstorms', 'rain': 'rain or snow', 'cloudy': 'cloudy', 'unknown': 'unknown',
'mostlycloudy': 'cloudy', 'partlycloudy': 'cloudy', 'clear': 'clear', 'hazy': 'not clear',
'fog': 'not clear', 'snow': 'rain or snow', 'sleet': 'rain or snow'}
return maps[event]
data['events'] = list(map(events_map, data['events']))
# save used informaiton
columns = ['trip_id', 'year', 'month', 'week', 'day', 'hour', 'usertype', 'gender', 'starttime',
'stoptime', 'tripduration', 'temperature', 'events', 'from_station_id', 'from_station_name',
'latitude_start', 'longitude_start', 'dpcapacity_start', 'to_station_id', 'to_station_name',
'latitude_end', 'longitude_end', 'dpcapacity_end']
data.to_csv('./data/data.csv', columns=columns, index=False)
_ = gc.collect()
# read the clean dataset
data = pd.read_csv('./data/data.csv')
data.head()
data.info()
```
# II. Visualization and Analysis
### Trip Distribution
```
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='year', data=data, ax=ax[0])
ax[0].set_title('Trip Count vs. Year', fontsize=16)
ax[0].set_xlabel('Year', fontsize=12)
ax[0].set_ylabel('Count', fontsize=12)
sns.boxplot(x='year', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Year', fontsize=16)
ax[1].set_xlabel('Year', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='month', data=data, ax=ax[0])
ax[0].set_title('Trip Count vs. Month', fontsize=16)
ax[0].set_xlabel('Month', fontsize=12)
ax[0].set_ylabel('Count', fontsize=12)
sns.boxplot(x='month', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Month', fontsize=16)
ax[1].set_xlabel('Month', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='day', data=data, ax=ax[0])
ax[0].set_xticklabels(('Mon', 'Tue', 'Wed', 'Thur', 'Fri', 'Sat', 'Sun'))
ax[0].set_title('Trip Count vs. Day of Week', fontsize=16)
ax[0].set_xlabel('Day of Week', fontsize=12)
ax[0].set_ylabel('Count', fontsize=12)
sns.boxplot(x='day', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Day of Week', fontsize=16)
ax[1].set_xlabel('Day of Week', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
ax[1].set_xticklabels(('Mon', 'Tue', 'Wed', 'Thur', 'Fri', 'Sat', 'Sun'))
plt.show()
# heatmap of trip count vs. (weekday and week no.)
fig, ax = plt.subplots(figsize=(16, 6))
tmp = data[['day', 'week', 'tripduration']].groupby(['day', 'week']).count().reset_index()
pivots = tmp.pivot('day', 'week', 'tripduration')
sns.heatmap(pivots, cbar_kws={'label': 'Trip Counts'})
ax.set_title('Heatmap of Trip Counts vs. (Day of Week and Week No.)', fontsize=16)
ax.set_xlabel('Week No.', fontsize=12)
ax.set_ylabel('Day of Week', fontsize=12)
ax.set_yticklabels(('Mon', 'Tue', 'Wed', 'Thur', 'Fri', 'Sat', 'Sun'), fontsize=10)
plt.tight_layout()
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='hour', data=data, ax=ax[0])
ax[0].set_title('Trip Count vs. Hour', fontsize=16)
ax[0].set_xlabel('Hour', fontsize=12)
ax[0].set_ylabel('Count', fontsize=12)
sns.boxplot(x='hour', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Hour', fontsize=16)
ax[1].set_xlabel('Hour', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# heatmap of trip count vs. (day and hour)
fig, ax = plt.subplots(figsize=(16, 6))
tmp = data[['day', 'hour', 'tripduration']].groupby(['day', 'hour']).count().reset_index()
pivots = tmp.pivot('day', 'hour', 'tripduration')
sns.heatmap(pivots, cbar_kws={'label': 'Trip Counts'})
ax.set_title('Heatmap of Trip Counts vs. (Day of Week and Hour)', fontsize=16)
ax.set_xlabel('Hour', fontsize=12)
ax.set_ylabel('Day of Week', fontsize=12)
ax.set_yticklabels(('Mon', 'Tue', 'Wed', 'Thur', 'Fri', 'Sat', 'Sun'), fontsize=10)
plt.tight_layout()
plt.show()
# Trip distribution
# Subscriber: Annual Membership
# Cumtomer: 24-hour pass
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='usertype', data=data, ax=ax[0])
ax[0].set_yscale('log')
ax[0].set_title('Trip Count vs. User Type', fontsize=16)
ax[0].set_xlabel('User Type', fontsize=12)
ax[0].set_ylabel('Count (log scale)', fontsize=12)
sns.boxplot(x='usertype', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. User Type', fontsize=16)
ax[1].set_xlabel('User Type', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='gender', data=data, ax=ax[0])
ax[0].set_title('Trip Count vs. Gender', fontsize=16)
ax[0].set_xlabel('Gender', fontsize=12)
ax[0].set_ylabel('Count', fontsize=12)
sns.boxplot(x='gender', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Gender', fontsize=16)
ax[1].set_xlabel('Gender', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(16, 6))
sns.countplot(x='events', data=data, ax=ax[0])
ax[0].set_yscale('log')
ax[0].set_title('Trip Count vs. Weather', fontsize=16)
ax[0].set_xlabel('Weather', fontsize=12)
ax[0].set_ylabel('Count (log scale)', fontsize=12)
sns.boxplot(x='events', y='tripduration', data=data, ax=ax[1])
ax[1].set_title('Trip Duration vs. Weather', fontsize=16)
ax[1].set_xlabel('Weather', fontsize=12)
ax[1].set_ylabel('Trip Duration (minutes)', fontsize=12)
plt.show()
# Trip distribution
fig, ax = plt.subplots(nrows=2, ncols=1, figsize=(12, 10))
sns.distplot(data['tripduration'], ax=ax[0],
hist_kws={'histtype': 'bar', 'edgecolor':'black'})
ax[0].set_xlabel('Trip Duration (minutes)', fontsize=12)
ax[0].set_title('Trip Duration Distribution')
sns.distplot(np.log(data['tripduration']), ax=ax[1],
hist_kws={'histtype': 'bar', 'edgecolor':'black'})
ax[1].set_xlabel('Trip Duration (minutes)', fontsize=12)
ax[1].set_title('Trip Duration Distribution (log scale)')
plt.show()
# trip duration vs. temperature
fig, ax = plt.subplots(figsize=(8, 6))
sns.heatmap(data[['tripduration', 'temperature']].corr())
ax.set_xticklabels(labels=['Trip Duration', 'Temperature'], fontsize=12)
ax.set_yticklabels(labels=['Trip Duration', 'Temperature'], fontsize=12)
ax.set_title('Trip Duration and Temperature Correlation', fontsize=14)
plt.axis('image')
plt.show()
```
### Trip Start and Stop Stations
```
# trip start longitude and latitude distribution
g = sns.jointplot('longitude_start', 'latitude_start', data=data, kind='hex')
g.set_axis_labels('Trip Start Longitude', 'Trip Start Latitude')
g.fig.set_figwidth(8)
g.fig.set_figheight(10)
plt.show()
# trip end longitude and latitude distribution
g = sns.jointplot('longitude_end', 'latitude_end', data=data, kind='hex')
g.set_axis_labels('Trip End Longitude', 'Trip End Latitude')
g.fig.set_figwidth(8)
g.fig.set_figheight(10)
plt.show()
# group station according GPS locations
station_start = data[['longitude_start', 'latitude_start', 'tripduration']]
station_start = station_start.groupby(['longitude_start', 'latitude_start'])
station_start = station_start.count().reset_index()
station_end = data[['longitude_end', 'latitude_end', 'tripduration']]
station_end = station_end.groupby(['longitude_end', 'latitude_end'])
station_end = station_end.count().reset_index()
# visualization
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(14, 10))
x = station_start['longitude_start']
y = station_start['latitude_start']
s = station_start['tripduration'] / 200
ax[0].scatter(x, y, s=s, alpha=0.1)
ax[0].set_title('Scatter Plot of Trip Start Stations', fontsize=16)
ax[0].set_xlabel('Longitude', fontsize=12)
ax[0].set_ylabel('Latitude', fontsize=12)
x = station_end['longitude_end']
y = station_end['latitude_end']
s = station_end['tripduration'] / 200
ax[1].scatter(x, y, s=s, alpha=0.1)
ax[1].set_title('Scatter Plot of Trip End Stations', fontsize=16)
ax[1].set_xlabel('Longitude', fontsize=12)
plt.show()
```
# III. Conclusions
Through above visualization figures, I have noticed several interesting phenonmena.
+ From 2014 to 2017, there are increasing demand for sharing bicycles.
- Since the population of Chicago is relative constant, we can assume that people tend to live healthier as time goes on.
+ Peole use sharing bicycles more in summer than winter (more frequently usage and longer trip duration)
+ People use sharing bicycle more frequently in weekdays than weekends, but the average trip duration is longer in weekends than weekdays.
+ During rush hours (~8:00 am and ~5:00 pm), there are increasing demand for sharing bicycles during weekdays, but on weekends, there is no such clear trend.
+ Most users purchased the Annual Membership. But their trip durations are relatively sharter compared with ordinary customers and dependent. Those who purchased 24-hour pass tend to have longest trip.
+ Male users use sharing bicycle more often than female users, but female users have longer trips.
+ Most trips are within 15 minutes.
+ Most usage is in downtown Chicago. There are some stations rarely used.
# IV. What's Next?
### Fixed stations vs. Station-less, which one is better?
| Chicago Divvy Bicycle | China ofo / Mobike Bicycle |
|----------------------------|----------------------------|
| Fixed stations | Station-less |
| Easy to manage | Hard to manage |
| Easy to track | Hard to track single user |
| High cost | Low cost |
| And so on | And so on |
| github_jupyter |
For the original notebook you can check [here](https://digitalsinology.org/classical-chinese-digital-humanities/). I answered the exercises of this notebook, so if you want to write your own answer, you can just ignore my answers.
Classical Chinese DH: Getting started
=====
*By [Donald Sturgeon](http://dsturgeon.net/about)*
\[[View this notebook online](http://digitalsinology.org/classical-chinese-dh-getting-started)\] \[[Download this notebook](http://digitalsinology.org/notebooks/classical-chinese-dh-1.ipynb)\] \[[List of tutorials](http://digitalsinology.org/classical-chinese-digital-humanities/)\]
### Welcome to our first Jupyter Notebook!
A [notebook](http://jupyter-notebook-beginner-guide.readthedocs.org/en/latest/what_is_jupyter.html) is a [hypertext](https://en.wikipedia.org/wiki/Hypertext) document containing a mixture of textual content (like the part you're reading now) and computer programs - lists of instructions written in a programming language (in our case, the [Python](https://en.wikipedia.org/wiki/Python_%28programming_language%29) language) - as well as the output of these programs.
### Using the Jupyter environment
Before getting started with Python itself, it's important to get some basic familiarity with the user interface of the Jupyter environment. Jupyter is fairly intuitive to use, partly because it runs in a web browser and so works a lot like any web page. Basic principles:
* Each "notebook" displays as a single page. Notebooks are opened and saved using the menus and icons shown **within** the Jupyter window (i.e. the menus and icons under the Jupyter logo and icon, **not** the menus / icons belonging to your web browser).
* Notebooks are made up of "cells". Each cell is displayed on the page in a long list, one below another. You can see which parts of the notebook belong to which cell by clicking once on the text - when you do this, this will select the cell containing the text, and show its outline with a grey line.
* Usually a cell contains either text (like this one - in Jupyter this is called a "Markdown" cell), or Python code (like the one below this one).
* You can click on a program cell to edit it, and double-click on a text cell to edit it. Try double-clicking on this cell.
* When you start editing a text cell, the way it is displayed changes so that you can see (and edit) any formatting codes in it. To return the cell back to the "normal" prettified display, you need to "Run" it. You can run a cell by either:
* choosing "Run" from the "Cell" menu above,
* pressing shift-return when the cell is selected, or
* clicking the "Run cell" icon.
* "Run" this cell so that it returns to the original mode of display.
```
for number in range(1,13):
print(str(number) + "*" + str(number) + " = " + str(number*number))
```
The program in a cell doesn't do anything until you ask Jupyter to run (a.k.a. "execute") it - in other words, ask the system to start following the instructions in the program. You can execute a cell by clicking somewhere in it so it's selected, then choosing "Run" from the "Cell" menu (or by pressing shift-return).
When you run a cell containing a Python program, any output that the program generates is displayed directly below that cell. If you modify the program, you'll need to run it again before you will see the modified result.
A lot of the power of Python and Jupyter comes from the ability to easily make use of modules written by other people. Modules are included using lines like "from ... import \*".
A module needs to be installed on your computer before you can use it; many of the most commonly used ones are installed as part of Anaconda.
"Comments" provide a way of explaining to human readers what parts of a program are supposed to do (but are completely ignored by Python itself). Typing the symbol # begins a comment, which continues until the end of the line.
**N.B.** You must install the "ctext" module before running the code below. If you get the error "ImportError: No module named 'ctext'" when you try to run the code, [refer to the instructions](http://digitalsinology.org/classical-chinese-dh-getting-started/) for how to install the ctext module.
```
from ctext import * # This module gives us direct access to data from ctext.org
setapikey("demo") # This allows us access to the data used in these tutorials
paragraphs = gettextasparagrapharray("ctp:analects/xue-er")
print("This chapter is made up of " + str(len(paragraphs)) + " paragraphs. These are:")
# For each paragraph of the chapter data that we downloaded, do the following:
for paragraphnumber in range(0, len(paragraphs)):
print(str(paragraphnumber+1) + ". " + paragraphs[paragraphnumber])
```
'Variables' are named entities that contain some kind of data that can be changed at a later date. We will look at these in much more detail over the next few weeks. For now, you can think of them as named boxes which can contain any kind of data.
Once we have data stored in a variable (like the 'paragraphs' variable above), we can start processing it in whatever way we want. Often we use other variables to track our progress, like the 'longest_paragraph' and 'longest_length' variables in the program below.
```
longest_paragraph = None # We use this variable to record which of the paragraphs we've looked at is longest
longest_length = 0 # We use this one to record how long the longest paragraph we've found so far is
for paragraph_number in range(0, len(paragraphs)):
paragraph_text = paragraphs[paragraph_number];
if len(paragraph_text)>longest_length:
longest_paragraph = paragraph_number
longest_length = len(paragraph_text)
print("The longest paragraph is paragraph number " + str(longest_paragraph+1) + ", which is " + str(longest_length) + " characters long.")
```
Modules allow us to do powerful things like Principle Component Analysis (PCA) and machine learning without having to write any code to perform any of the complex mathematics which lies behind these techniques. They also let us easily plot numerical results within the Jupyter notebook environment.
For example, the following code (which we will go through in much more detail in a future tutorial - don't worry about the contents of it yet) plots the frequencies of the two characters "矣" and "也" in chapters of the Analects versus chapters of the Fengshen Yanyi. (Note: this may take a few seconds to download the data.)
```
import re
import pandas as pd
import matplotlib.pyplot as plt
%matplotlib inline
def makevector(string, termlist, normalize = False):
vector = []
for term in termlist:
termcount = len(re.findall(term, string))
if normalize:
vector.append(termcount/len(string))
else:
vector.append(termcount)
return vector
text1 = gettextaschapterlist("ctp:fengshen-yanyi")
text2 = gettextaschapterlist("ctp:analects")
vectors1 = []
for chapter in text1:
vectors1.append(makevector(chapter, ["矣", "也"], True))
vectors2 = []
for chapter in text2:
vectors2.append(makevector(chapter, ["矣", "也"], True))
df1 = pd.DataFrame(vectors1)
df2 = pd.DataFrame(vectors2)
legend1 = plt.scatter(df1.iloc[:,0], df1.iloc[:,1], color="blue", label="Fengshen Yanyi")
legend2 = plt.scatter(df2.iloc[:,0], df2.iloc[:,1], color="red", label="Analects")
plt.legend(handles = [legend1, legend2])
plt.xlabel("Frequency of 'yi'")
plt.ylabel("Frequency of 'ye'")
```
You can save changes to your notebook using "File" -> "Save and checkpoint". Note that Jupyter often saves your changes for you automatically, so if you *don't* want to save your changes, you might want to make a copy of your notebook first using "File" -> "Make a Copy".
You should try to avoid having the same notebook open in two different browser windows or browser tabs at the same time. (If you do this, both pages may try to save changes to the same file, overwriting each other's work.)
Exercises
----
Before we start writing programs, we need to get familiar with the Jupyter Notebook programming environment. Check that you can complete the following tasks:
* Run each of the program cells in this notebook that are above this cell on your computer, checking that each of the short programs produces the expected output.
* Clear all of the output using "Cell" -> "All output" -> "Clear", then run one or two of them again.
* In Jupyter, each cell in a notebook can be run independently. Sometimes the _order_ in which cells are run is important. Try running the following three cells in order, then see what happens when you run them in a different order. Make sure you understand why in some cases you get different results.
```
number_of_things = 1
print(number_of_things)
number_of_things = number_of_things + 1
print(number_of_things)
```
* Some of the programs in this notebook are very simple. Modify and re-run them to perform the following tasks:
* Print out the squares of the numbers 3 through 20 (instead of 1 through 12)
* Print out the cubes of the numbers 3 through 20 (i.e. 3 x 3 x 3 = 27, 4 x 4 x 4 = 64, etc.)
* Instead of printing passages from the first chapter of the Analects, print passages from the **Daodejing**, and determine the longest passage in it. The **URN** for the Daodejing is: `ctp:dao-de-jing`
```
# Jibancat: print out squares if the number 3 through 20
for i in range(3, 20):
print ('{0}^2 = {1}'.format(i, i ** 2))
# Jibancat: print out cubes if the number 3 through 20
for i in range(3, 20):
print ('{0}^3 = {1}'.format(i, i ** 3))
# this would give the full text of dao-de-jing in a list,
# every elements in the list represent one paragraph
dao_de_jing = gettextasparagrapharray("ctp:dao-de-jing")
# get the index of the longest paragraph
lengthDDJ = list(map(len, dao_de_jing))
lengthDDJ.index(max(lengthDDJ))
dao_de_jing[38]
```
* Often when programming you'll encounter error messages. The following line contains a bug; try running it, and look at the output. Work out which part of the error message is most relevant, and see if you can find an explanation on the web (e.g. on StackOverflow) and fix the mistake.
```
print("The answer to life the universe and everything is: " + 42) # This statement is incorrect and isn't going to work
# we cannot use "+" between a string and a integer
# This would work
print("The answer to life the universe and everything is: ", 42)
```
* Sometimes a program will take a long time to run - or even run forever - and you'll need to stop it. Watch what happens to the circle beside the text "Python 3" at the top-right of the screen when you run the cell below.
* While the cell below is running, try running the cell above. You won't see any output until the cell below has finished running.
* Run the cell below again. While it's running, interrupt its execution by clicking "Kernel" -> "Interrupt".
```
import time
for number in range(1,21):
print(number)
time.sleep(1)
```
* The cell below has been set as a "Markdown" cell, making it a text cell instead of a program ("code") cell. Work out how to make the cell run as a program.
```python
for number in range(1,11):
print("1/" + str(number) + " = " + str(1/number)) # In many programming languages, the symbol "/" means "divided by"
```
You can make a similar code snippet use following markdown language in a text cell.
```python
# you sample code here
```
```
# you can just press "y" on the keyvboard,
# the cell would be forced to be a code cell
for number in range(1,11):
print("1/" + str(number) + " = " + str(1/number)) # In many programming languages, the symbol "/" means "divided by"
```
* Experiment with creating new cells below this one. Make some text cells, type something in them, and run them. Copy and paste some code from above into code cells, and run them too. Try playing around with simple modifications to the code.
* (Optional) You can make your text cells look nicer by including formatting instructions in them. **The way of doing this is called "Markdown" - there are many [good introductions](https://athena.brynmawr.edu/jupyter/hub/dblank/public/Jupyter%20Notebook%20Users%20Manual.ipynb#4.-Using-Markdown-Cells-for-Writing) available online. **
* Lastly, save your modified notebook and close your web browser. Shut down the Python server process, then start it again, and reload your modified notebook. Make sure you can also find the saved notebook file in your computer's file manager (e.g. "Windows Explorer"/"File Explorer" on Windows, or "Finder" on Mac OS X).
**Further reading:**
* [Jupyter Notebook Users Manual](https://athena.brynmawr.edu/jupyter/hub/dblank/public/Jupyter%20Notebook%20Users%20Manual.ipynb), Bryn Mawr College Computer Science - _This provides a thorough introduction to Jupyter features. This guide introduces many more features than we will need to use, but is a great reference._
<div style="float: right;"><a rel="license" href="http://creativecommons.org/licenses/by-sa/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-sa/4.0/88x31.png" /></a></div>
| github_jupyter |
```
import random
import numpy as np
from collections import deque
from keras.models import Sequential, Model
from keras.layers import Dense, Input, Conv2D, Flatten, Activation, MaxPooling2D
from keras.optimizers import Adam
import keras
import logging
import pickle
import os.path
import nnutils
name = "data/CattleG1"
guylaine_input_size = 100
state_width = nnutils.tileWidth
state_height = nnutils.tileHeight
state_channels = 14
ship_input_size = 4
output_size = 6
memory = deque(maxlen=2000)
gamma = 0.95 # discount r
epsilon = 1.0 # exploration
epsilon_min = 0.01
epsilon_decay = 0.995
learning_rate = 0.001
guylaine_input = Input(shape=(guylaine_input_size,), name='ship_guylaine_input')
ship_input = Input(shape=(ship_input_size,), name='ship_input')
x = keras.layers.concatenate([guylaine_input, ship_input])
x = Dense(64, activation='relu')(x)
x = Dense(64, activation='relu')(x)
x = Dense(64, activation='relu')(x)
ship_output = Dense(output_size, activation='sigmoid', name='cattle_output')(x)
model = Model(inputs=[guylaine_input, ship_input], outputs=ship_output)
model.compile(loss='mse',
optimizer=Adam(lr=learning_rate))
model.load_weights(name)
memory = pickle.load(open(name + '_memory', 'rb'))
epsilon = pickle.load(open(name + '_epsilon', 'rb'))
from matplotlib import pyplot as plt
from IPython.display import clear_output
# updatable plot
# a minimal example (sort of)
class PlotLosses(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.i = 0
self.x = []
self.losses = []
self.val_losses = []
self.fig = plt.figure()
self.logs = []
def on_epoch_end(self, epoch, logs={}):
self.logs.append(logs)
self.x.append(self.i)
self.losses.append(logs.get('loss'))
self.val_losses.append(logs.get('val_loss'))
self.i += 1
clear_output(wait=True)
plt.plot(self.x, self.losses, label="loss")
plt.plot(self.x, self.val_losses, label="val_loss")
plt.legend()
plt.show();
plot_losses = PlotLosses()
minBatchSize = batch_size
if (len(memory) < batch_size):
minBatchSize = len(memory)
minibatch = random.sample(memory, minBatchSize)
for guylaine_output, ship_state, action, reward, next_guylaine_output, next_ship_state, done in minibatch:
target = reward
if not done:
target = (reward + gamma * model.predict({'ship_guylaine_input': next_guylaine_output, 'ship_input': next_ship_state}))
target_f = model.predict({'ship_guylaine_input': guylaine_output, 'ship_input': ship_state})
action_index = np.argmax(action)
target_f[0][action_index] = target
model.fit(state, target_f, epochs=1, verbose=0)
if epsilon > epsilon_min:
epsilon *= epsilon_decay
```
| github_jupyter |
# Coding Challange - 06 - Solution
### Question 1:
Write a python class Person that is initialized with age. Create two more classes, Male and Female, that are inherited from the Person class. The two-child classes should contain an instance method marriage(), which returns if the Person can marry or not depending upon the age.
Take marriage age for Male as 21 years and Female as 18 years.
#### Sample Input:
Male - 16
<br>Female - 24
#### Expected Output:
Cannot Marry
<br>Can Marry
```
class Person:
def __init__(self, age):
self.age = age
class Male(Person):
def marriage(self):
if self.age > 21:
print('Can Marry')
else:
print('Cannot Marry')
class Female(Person):
def marriage(self):
if self.age > 18:
print('Can Marry')
else:
print('Cannot Marry')
male_obj = Male(16)
female_obj = Female(24)
male_obj.marriage()
female_obj.marriage()
```
### Question 2:
You are given a task to put a number tag on the books on a bookshelf. Each of the n numbers of books should have a number from 1 to n, and different books should have different numbers. Write a python function to calculate the number of digits required for numbering all those tags.
#### Expected Output:
Enter the number of books:95
<br>181
```
def book_tags(n):
if n == 1:
return 1
s = str(n)
return len(s) + book_tags(n - 1)
no_of_books = int(input('Enter the number of books:'))
print(book_tags(no_of_books))
```
### Question 3:
Write a python program to get the difference between a number and the reverse of that number. Print the difference as a positive integer.
#### Sample Input:
num = 86
#### Expected Output:
18
```
def reverse_diff(num):
return num
def reverse(num):
rev = 0
while(num > 0):
rem = num %10
rev = (rev *10) + rem
num = num //10
return rev
diff = abs(reverse_diff(86) - reverse(86))
print(diff)
```
### Question 4:
Write a python function to print the time given below into military time(24 Hours) format.
10:20:45PM
#### Expected Output:
'22:20:45'
```
def military_time_conversion(s):
time = s.split(":")
if s[-2:] == "PM":
if time[0] != "12":
time[0] = str(int(time[0])+12)
else:
if time[0] == "12":
time[0] = "00"
ntime = ':'.join(time)
return str(ntime[:-2])
military_time_conversion('10:20:45PM')
```
### Question 5:
Using user-defined exception, write a python program to guess the number entered in the code until you get the correct answer.
#### Expected Output:
Enter a number greater than 0: 12
<br>Less than the desired number, please try again!
Enter a number greater than 0: 18
<br>Greater than the desired number, try again!
Enter a number greater than 0: 15
<br>BINGO! That's the number.
```
class NumError(Exception):
pass
class NumSmallError(NumError):
pass
class NumLargeError(NumError):
pass
num = 15
while True:
try:
input_num = int(input("Enter a number greater than 0: "))
if input_num < num:
raise NumSmallError
elif input_num > num:
raise NumLargeError
break
except NumSmallError:
print("Less than the desired number, please try again!\n")
except NumLargeError:
print("Greater than the desired number, try again!\n")
print("BINGO! That's the number.")
```
### Question 6:
Write a python function to find the minimum and maximum values calculated by summing precisely four of the five integers from the list given below.
myList = [1, 3, 5, 7, 9]
#### Expected Output:
Minimum: 16
<br>Maximum: 24
```
def min_max(*numList):
sum=0
for i in range(len(numList)):
sum += numList[i]
print('Minimum: ', sum-max(numList))
print('Maximum: ', sum-min(numList))
myList = [1, 3, 5, 7, 9]
min_max(*myList)
```
### Question 7:
We have a list of numbers given below. Write a python function to ask the user to input a number from the list. Then, push that number towards the end of the list. If the entered number is not present in the list, print the old list.
num_list = [0, 2, 4, 5, 6, 2, 3]
#### Expected Output:
Enter the number to be moved: 2
<br>[0, 4, 5, 6, 3, 2, 2]
```
def move_num(*num_list):
remove_num = [n for i in range(num_list.count(n))]
new_list = [i for i in num_list if i != n]
new_list.extend(remove_num)
return(new_list)
myList = [0, 2, 4, 5, 6, 2, 3]
n = int(input('Enter the number to be moved: '))
print(move_num(*myList))
```
### Question 8:
Write a python program to check if the number is divisible by its digits.
#### Expected Output:
Enter a number: 48
<br>Yes
```
def check_div(num, digit) :
return (digit != 0 and num % digit == 0)
def digits_divide(x) :
temp = x
while (temp > 0) :
digit = temp % 10
if ((check_div(x, digit)) == False) :
return False
temp = temp // 10
return True
n = int(input('Enter a number: '))
if digits_divide(n):
print("Yes")
else:
print("No" )
```
### Question 9:
Write a python function to evaluate the grade ranging from 0 to 100 in an examination. The student fails the exam if the grade is less than 40. But we like to round off grades above 40 based on the following criteria.
If the difference between the grade and the next multiple of 5 is less than 3, round grade up to the next multiple of 5.
#### Expected Output:
Enter the number of subjects:4
<br>Marks of Subjects 1:36
<br>Marks of Subjects 2:48
<br>Marks of Subjects 3:73
<br>Marks of Subjects 4:66
<br>Resultant Grade: [36, 50, 75, 66]
```
def student_grades(grades):
for i in range(len(grades)):
if(grades[i] > 37):
if((grades[i] % 5) != 0):
if(5 - (grades[i] % 5) < 3):
grades[i] += 5 - (grades[i] % 5)
return (grades)
grades_count = int(input('Enter the number of subjects:'))
grades = []
for i in range(grades_count):
grades_item = int(input(f'Marks of Subjects {i + 1}:'))
grades.append(grades_item)
result = student_grades(grades)
print('Resultant Grade: ', result)
```
### Question 10:
Write a python function grow_more to get a sapling height, which increases by one unit in March and by two units in August in a year. The argument of the function takes the number of growth cycles the sapling goes through. Consider the sapling is planted in January.
#### Sample Input:
7
#### Expected Output:
30
#### Explanation:
At 0 --> 1
<br>At 1 --> 2
<br>At 2 --> 3
<br>At 3 --> 6
<br>At 4 --> 7
<br>At 5 --> 14
<br>At 6 --> 15
<br>At 7 --> 30
```
def grow_more(num):
boolean = False
count = 0
for i in range(num + 1):
if(boolean == False):
count += 1
boolean = True
else:
count *= 2
boolean = False
return count
grow_more(7)
```
| github_jupyter |
```
import geopandas as gpd
from tqdm import tqdm
from shapely.geometry import Point, Polygon, MultiPolygon
import shapely
def queen_corners(gdf, sensitivity=2):
"""
Experimental: Fix unprecise corners.
"""
tessellation = gdf.copy()
changes = {}
qid = 0
sindex = tessellation.sindex
for ix, row in tqdm(tessellation.iterrows(), total=tessellation.shape[0]):
corners = []
change = []
cell = row.geometry
coords = cell.exterior.coords
for i in coords:
point = Point(i)
possible_matches_index = list(sindex.intersection(point.bounds))
possible_matches = tessellation.iloc[possible_matches_index]
precise_matches = sum(possible_matches.intersects(point))
if precise_matches > 2:
corners.append(point)
if len(corners) > 2:
for c, it in enumerate(corners):
next_c = c + 1
if c == (len(corners) - 1):
next_c = 0
if corners[c].distance(corners[next_c]) < sensitivity:
change.append([corners[c], corners[next_c]])
elif len(corners) == 2:
if corners[0].distance(corners[1]) > 0:
if corners[0].distance(corners[1]) < sensitivity:
change.append([corners[0], corners[1]])
if change:
for points in change:
x_new = np.mean([points[0].x, points[1].x])
y_new = np.mean([points[0].y, points[1].y])
new = [(x_new, y_new), id]
changes[(points[0].x, points[0].y)] = new
changes[(points[1].x, points[1].y)] = new
qid = qid + 1
for ix, row in tqdm(tessellation.iterrows(), total=tessellation.shape[0]):
cell = row.geometry
coords = list(cell.exterior.coords)
moves = {}
for x in coords:
if x in changes.keys():
moves[coords.index(x)] = changes[x]
keys = list(moves.keys())
delete_points = []
for move, k in enumerate(keys):
if move < len(keys) - 1:
if (
moves[keys[move]][1] == moves[keys[move + 1]][1]
and keys[move + 1] - keys[move] < 5
):
delete_points = delete_points + (
coords[keys[move] : keys[move + 1]]
)
# change the code above to have if based on distance not number
newcoords = [changes[x][0] if x in changes.keys() else x for x in coords]
for coord in newcoords:
if coord in delete_points:
newcoords.remove(coord)
if coords != newcoords:
if not cell.interiors:
# newgeom = Polygon(newcoords).buffer(0)
be = Polygon(newcoords).exterior
mls = be.intersection(be)
if len(list(shapely.ops.polygonize(mls))) > 1:
newgeom = MultiPolygon(shapely.ops.polygonize(mls))
geoms = []
for g, n in enumerate(newgeom):
geoms.append(newgeom[g].area)
newgeom = newgeom[geoms.index(max(geoms))]
else:
newgeom = list(shapely.ops.polygonize(mls))[0]
else:
newgeom = Polygon(newcoords, holes=cell.interiors)
tessellation.loc[ix, "geometry"] = newgeom
return tessellation
path = 'folder/AMS'
tess = gpd.read_file(path + '/elements.gpkg', layer='tessellation')
queen = queen_corners(tess)
queen.to_file(path + 'queen.gpkg', layer='tessellation', driver='GPKG')
```
| github_jupyter |
[](https://pythonista.io)
Python pone a disposición de los desarrolladores mediante la "Autoridad de Empaquetado de Python" (pypa) herramientas que le permiten "empaquetar" sus proyectos para que estos sean distribuidos con facilidad.
El sitio https://packaging.python.org/ ofrece tutoriales, especificaciones y contenidos para facilitar y normar el empaquetado y distribución de paquetes en Python.
Un paquete es una estructura de directorios que incluye una biblioteca de código, documentación, archivos de configuración y datos de un proyecto específico, la cual se encuentra comprimida y puede ser reutilizada por cualquier otro usuario o desarrollador.
## Los módulos *distutils* y *setuptools*.
El módulo *distutils* fue la herramienta de gestión de paquetes original de Python, sin embargo esta ha sido extendida y en la mayoría de los casos, sustituida por el módulo *setuptools*.
```
import setuptools
help(setuptools)
import distutils
help(distutils)
```
## Estructura general de un proyecto.
Un proyecto por lo general tiene una estructura específica. Un ejemplo de dicha estructura puede ser consultado en https://github.com/pypa/sampleproject. Dicha estructura comprende por lo general diversos directorios correspondientes a:
* La bilbioteca de código.
* Archivos de configuración.
* Archivos de datos.
* Archivos para pruebas.
### Archivos de texto que normalmente se incluyen en un proyecto.
* **README.rst**, el cual es un archivo de texto que puede contener una estructura basada en [reStrcuturedText](http://docutils.sourceforge.net/rst.html).
* **LICENSE.txt**, en donde se especifica la licencia bajo la que se libera el código fuente.
* **MANIFEST.in** en le que se indica el contenido del paquete.
* **setup.cfg** en el que se indica la configuración del paquete.
* **setup.py** el script para la creación del paquete.
## El archivo *setup.py*.
Este archivo es el que se utiliza para empaquetar el proyecto y un ejemplo básico es el siguiente.
``` python
from setuptools import setup, find_packages
setup(
name="HelloWorld",
version="0.1",
packages=find_packages(),
)
```
### La función *setup()* de *setuptools*.
Esta es la función primordial para la creación de los paquetes.
### Campos que puede de incluir en la función *setup.()*.
Entre otras cosas, se pueden incluir los siguientes campos:
* *name*
* *version*
* *description*
* *author*
* *author_email*
* *url*
* *download_url*
* *license*
* *packages*
* *py_modules*
**Ejemplo extendido:**
``` python
from setuptools import setup, find_packages
setup(
name="HelloWorld",
version="0.1",
packages=find_packages(),
scripts=['say_hello.py'],
# Project uses reStructuredText, so ensure that the docutils get
# installed or upgraded on the target machine
install_requires=['docutils>=0.3'],
package_data={
# If any package contains *.txt or *.rst files, include them:
'': ['*.txt', '*.rst'],
# And include any *.msg files found in the 'hello' package, too:
'hello': ['*.msg'],
},
# metadata for upload to PyPI
author="Me",
author_email="me@example.com",
description="This is an Example Package",
license="PSF",
keywords="hello world example examples",
url="http://example.com/HelloWorld/", # project home page, if any
project_urls={
"Bug Tracker": "https://bugs.example.com/HelloWorld/",
"Documentation": "https://docs.example.com/HelloWorld/",
"Source Code": "https://code.example.com/HelloWorld/",
}
# could also include long_description, download_url, classifiers, etc.
)
```
### La función *find_packages()* de *setuptools*.
Esta función permite encontrar la estructura de paquetes en un directorio.
En este casó, se identifica a un paquete cuando el subdirectorio contiene un archivo *\_\_init\_\_.py*.
Además de la identificación de paquetes, la función *find_packages()* incluye los parámetros:
* *where*, el cual corresponde al directorio desde el cual se buscarabn los paquetes. Por defecto se utiliza el directorio desde el que se ejecuta la función.
* *exclude*, el cual puede ser un objeto tipo tuple que contiene una lista de aquellos paquetes que no se quieran añadir.
* *include*, el cual corresponde a una tupla que contiene una lista de los paquetes a añadir. Por defecto, añade a todos.
**Ejemplo:**
La función *find_packages()* se ejecutará en el directorio de esta notebook.
```
from setuptools import find_packages
find_packages()
```
## Creación del paquete.
Una vez que el archivo *setup.py* se encuentra disponible, sólo hay que ejecutarlo de la siguiente manera:
``` python
python setup.py sdist --formats=<formato>
```
Los formatos soportados son:
* *zip* para archivos con extensión *.zip*.
* *bztar* para archivos con extensión *.tar.bz*.
* *gztar* para archivos con extensión *.tar.gz*.
* *ztar* para archivos con extensión *.tar.z*.
* *tar* para archivos con extensión *.tar*.
**Ejemplo:**
El archivo [setup.py](setup.py) contiene el siguiente código:
``` python
from setuptools import setup, find_packages
setup(
name="paquete",
version=0.1,
packages=find_packages(),
)
```
```
%run setup.py sdist --formats=zip,gztar
```
El resultado serán un par de archivos en el directorio *[dist](dist)*.
```
!dir dist
```
Cualquiera de los dos paquetes puede ser instalado mediante *pip*.
```
!pip install dist/paquete-0.1.zip
!pip list | grep paquete
help('modules paquete')
```
Debido a que existe el directorio *paquete* en el directorio de trabajo de esta notebook, es necesario camibarla a otro para importar el paquete que está en la biblioteca.
```
%cd ..
import paquete
paquete.saluda()
help(paquete)
```
Para mayor información sobre el uso de *setuptools* y las opciones de empaquetado, puede acudir a https://setuptools.readthedocs.io/en/latest/setuptools.html
<p style="text-align: center"><a rel="license" href="http://creativecommons.org/licenses/by/4.0/"><img alt="Licencia Creative Commons" style="border-width:0" src="https://i.creativecommons.org/l/by/4.0/80x15.png" /></a><br />Esta obra está bajo una <a rel="license" href="http://creativecommons.org/licenses/by/4.0/">Licencia Creative Commons Atribución 4.0 Internacional</a>.</p>
<p style="text-align: center">© José Luis Chiquete Valdivieso. 2019.</p>
| github_jupyter |
**11장 – 심층 신경망 훈련하기**
_이 노트북은 11장에 있는 모든 샘플 코드와 연습문제 해답을 가지고 있습니다._
<table align="left">
<td>
<a target="_blank" href="https://colab.research.google.com/github/rickiepark/handson-ml2/blob/master/11_training_deep_neural_networks.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩에서 실행하기</a>
</td>
</table>
# 설정
먼저 몇 개의 모듈을 임포트합니다. 맷플롯립 그래프를 인라인으로 출력하도록 만들고 그림을 저장하는 함수를 준비합니다. 또한 파이썬 버전이 3.5 이상인지 확인합니다(파이썬 2.x에서도 동작하지만 곧 지원이 중단되므로 파이썬 3을 사용하는 것이 좋습니다). 사이킷런 버전이 0.20 이상인지와 텐서플로 버전이 2.0 이상인지 확인합니다.
```
# 파이썬 ≥3.5 필수
import sys
assert sys.version_info >= (3, 5)
# 사이킷런 ≥0.20 필수
import sklearn
assert sklearn.__version__ >= "0.20"
# 텐서플로 ≥2.0 필수
import tensorflow as tf
from tensorflow import keras
assert tf.__version__ >= "2.0"
%load_ext tensorboard
# 공통 모듈 임포트
import numpy as np
import os
# 노트북 실행 결과를 동일하게 유지하기 위해
np.random.seed(42)
# 깔끔한 그래프 출력을 위해
%matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
mpl.rc('axes', labelsize=14)
mpl.rc('xtick', labelsize=12)
mpl.rc('ytick', labelsize=12)
# 그림을 저장할 위치
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "deep"
IMAGES_PATH = os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID)
os.makedirs(IMAGES_PATH, exist_ok=True)
def save_fig(fig_id, tight_layout=True, fig_extension="png", resolution=300):
path = os.path.join(IMAGES_PATH, fig_id + "." + fig_extension)
print("그림 저장:", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(path, format=fig_extension, dpi=resolution)
```
# 그레이디언트 소실과 폭주 문제
```
def logit(z):
return 1 / (1 + np.exp(-z))
z = np.linspace(-5, 5, 200)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [1, 1], 'k--')
plt.plot([0, 0], [-0.2, 1.2], 'k-')
plt.plot([-5, 5], [-3/4, 7/4], 'g--')
plt.plot(z, logit(z), "b-", linewidth=2)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Saturating', xytext=(3.5, 0.7), xy=(5, 1), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Saturating', xytext=(-3.5, 0.3), xy=(-5, 0), arrowprops=props, fontsize=14, ha="center")
plt.annotate('Linear', xytext=(2, 0.2), xy=(0, 0.5), arrowprops=props, fontsize=14, ha="center")
plt.grid(True)
plt.title("Sigmoid activation function", fontsize=14)
plt.axis([-5, 5, -0.2, 1.2])
save_fig("sigmoid_saturation_plot")
plt.show()
```
## Xavier 초기화와 He 초기화
```
[name for name in dir(keras.initializers) if not name.startswith("_")]
keras.layers.Dense(10, activation="relu", kernel_initializer="he_normal")
init = keras.initializers.VarianceScaling(scale=2., mode='fan_avg',
distribution='uniform')
keras.layers.Dense(10, activation="relu", kernel_initializer=init)
```
## 수렴하지 않는 활성화 함수
### LeakyReLU
```
def leaky_relu(z, alpha=0.01):
return np.maximum(alpha*z, z)
plt.plot(z, leaky_relu(z, 0.05), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([0, 0], [-0.5, 4.2], 'k-')
plt.grid(True)
props = dict(facecolor='black', shrink=0.1)
plt.annotate('Leak', xytext=(-3.5, 0.5), xy=(-5, -0.2), arrowprops=props, fontsize=14, ha="center")
plt.title("Leaky ReLU activation function", fontsize=14)
plt.axis([-5, 5, -0.5, 4.2])
save_fig("leaky_relu_plot")
plt.show()
[m for m in dir(keras.activations) if not m.startswith("_")]
[m for m in dir(keras.layers) if "relu" in m.lower()]
```
LeakyReLU를 사용해 패션 MNIST에서 신경망을 훈련해 보죠:
```
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.fashion_mnist.load_data()
X_train_full = X_train_full / 255.0
X_test = X_test / 255.0
X_valid, X_train = X_train_full[:5000], X_train_full[5000:]
y_valid, y_train = y_train_full[:5000], y_train_full[5000:]
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(100, kernel_initializer="he_normal"),
keras.layers.LeakyReLU(),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
PReLU를 테스트해 보죠:
```
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, kernel_initializer="he_normal"),
keras.layers.PReLU(),
keras.layers.Dense(100, kernel_initializer="he_normal"),
keras.layers.PReLU(),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
### ELU
```
def elu(z, alpha=1):
return np.where(z < 0, alpha * (np.exp(z) - 1), z)
plt.plot(z, elu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1, -1], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title(r"ELU activation function ($\alpha=1$)", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("elu_plot")
plt.show()
```
텐서플로에서 쉽게 ELU를 적용할 수 있습니다. 층을 만들 때 활성화 함수로 지정하면 됩니다:
```
keras.layers.Dense(10, activation="elu")
```
### SELU
Günter Klambauer, Thomas Unterthiner, Andreas Mayr는 2017년 한 [훌륭한 논문](https://arxiv.org/pdf/1706.02515.pdf)에서 SELU 활성화 함수를 소개했습니다. 훈련하는 동안 완전 연결 층만 쌓아서 신경망을 만들고 SELU 활성화 함수와 LeCun 초기화를 사용한다면 자기 정규화됩니다. 각 층의 출력이 평균과
표준편차를 보존하는 경향이 있습니다. 이는 그레이디언트 소실과 폭주 문제를 막아줍니다. 그 결과로 SELU 활성화 함수는 이런 종류의 네트워크(특히 아주 깊은 네트워크)에서 다른 활성화 함수보다 뛰어난 성능을 종종 냅니다. 따라서 꼭 시도해 봐야 합니다. 하지만 SELU 활성화 함수의 자기 정규화 특징은 쉽게 깨집니다. ℓ<sub>1</sub>나 ℓ<sub>2</sub> 정규화, 드롭아웃, 맥스 노름, 스킵 연결이나 시퀀셜하지 않은 다른 토폴로지를 사용할 수 없습니다(즉 순환 신경망은 자기 정규화되지 않습니다). 하지만 실전에서 시퀀셜 CNN과 잘 동작합니다. 자기 정규화가 깨지면 SELU가 다른 활성화 함수보다 더 나은 성능을 내지 않을 것입니다.
```
from scipy.special import erfc
# alpha와 scale은 평균 0과 표준 편차 1로 자기 정규화합니다
# (논문에 있는 식 14 참조):
alpha_0_1 = -np.sqrt(2 / np.pi) / (erfc(1/np.sqrt(2)) * np.exp(1/2) - 1)
scale_0_1 = (1 - erfc(1 / np.sqrt(2)) * np.sqrt(np.e)) * np.sqrt(2 * np.pi) * (2 * erfc(np.sqrt(2))*np.e**2 + np.pi*erfc(1/np.sqrt(2))**2*np.e - 2*(2+np.pi)*erfc(1/np.sqrt(2))*np.sqrt(np.e)+np.pi+2)**(-1/2)
def selu(z, scale=scale_0_1, alpha=alpha_0_1):
return scale * elu(z, alpha)
plt.plot(z, selu(z), "b-", linewidth=2)
plt.plot([-5, 5], [0, 0], 'k-')
plt.plot([-5, 5], [-1.758, -1.758], 'k--')
plt.plot([0, 0], [-2.2, 3.2], 'k-')
plt.grid(True)
plt.title("SELU activation function", fontsize=14)
plt.axis([-5, 5, -2.2, 3.2])
save_fig("selu_plot")
plt.show()
```
기본적으로 SELU 하이퍼파라미터(`scale`과 `alpha`)는 각 뉴런의 평균 출력이 0에 가깝고 표준 편차는 1에 가깝도록 조정됩니다(입력은 평균이 0이고 표준 편차 1로 표준화되었다고 가정합니다). 이 활성화 함수를 사용하면 1,000개의 층이 있는 심층 신경망도 모든 층에 걸쳐 거의 평균이 0이고 표준 편차를 1로 유지합니다. 이를 통해 그레이디언트 폭주와 소실 문제를 피할 수 있습니다:
```
np.random.seed(42)
Z = np.random.normal(size=(500, 100)) # 표준화된 입력
for layer in range(1000):
W = np.random.normal(size=(100, 100), scale=np.sqrt(1 / 100)) # LeCun 초기화
Z = selu(np.dot(Z, W))
means = np.mean(Z, axis=0).mean()
stds = np.std(Z, axis=0).mean()
if layer % 100 == 0:
print("Layer {}: mean {:.2f}, std deviation {:.2f}".format(layer, means, stds))
```
쉽게 SELU를 사용할 수 있습니다:
```
keras.layers.Dense(10, activation="selu",
kernel_initializer="lecun_normal")
```
100개의 은닉층과 SELU 활성화 함수를 사용한 패션 MNIST를 위한 신경망을 만들어 보죠:
```
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="selu",
kernel_initializer="lecun_normal"))
for layer in range(99):
model.add(keras.layers.Dense(100, activation="selu",
kernel_initializer="lecun_normal"))
model.add(keras.layers.Dense(10, activation="softmax"))
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
```
이제 훈련해 보죠. 입력을 평균 0과 표준 편차 1로 바꾸어야 한다는 것을 잊지 마세요:
```
pixel_means = X_train.mean(axis=0, keepdims=True)
pixel_stds = X_train.std(axis=0, keepdims=True)
X_train_scaled = (X_train - pixel_means) / pixel_stds
X_valid_scaled = (X_valid - pixel_means) / pixel_stds
X_test_scaled = (X_test - pixel_means) / pixel_stds
history = model.fit(X_train_scaled, y_train, epochs=5,
validation_data=(X_valid_scaled, y_valid))
```
대신 ReLU 활성화 함수를 사용하면 어떤 일이 일어나는지 확인해 보죠:
```
np.random.seed(42)
tf.random.set_seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[28, 28]))
model.add(keras.layers.Dense(300, activation="relu", kernel_initializer="he_normal"))
for layer in range(99):
model.add(keras.layers.Dense(100, activation="relu", kernel_initializer="he_normal"))
model.add(keras.layers.Dense(10, activation="softmax"))
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model.fit(X_train_scaled, y_train, epochs=5,
validation_data=(X_valid_scaled, y_valid))
```
좋지 않군요. 그레이디언트 폭주나 소실 문제가 발생한 것입니다.
# 배치 정규화
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(100, activation="relu"),
keras.layers.BatchNormalization(),
keras.layers.Dense(10, activation="softmax")
])
model.summary()
bn1 = model.layers[1]
[(var.name, var.trainable) for var in bn1.variables]
bn1.updates
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
이따금 활성화 함수전에 BN을 적용해도 잘 동작합니다(여기에는 논란의 여지가 있습니다). 또한 `BatchNormalization` 층 이전의 층은 편향을 위한 항이 필요 없습니다. `BatchNormalization` 층이 이를 무효화하기 때문입니다. 따라서 필요 없는 파라미터이므로 `use_bias=False`를 지정하여 층을 만들 수 있습니다:
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.BatchNormalization(),
keras.layers.Dense(300, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(100, use_bias=False),
keras.layers.BatchNormalization(),
keras.layers.Activation("relu"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model.fit(X_train, y_train, epochs=10,
validation_data=(X_valid, y_valid))
```
## 그레이디언트 클리핑
모든 케라스 옵티마이저는 `clipnorm`이나 `clipvalue` 매개변수를 지원합니다:
```
optimizer = keras.optimizers.SGD(clipvalue=1.0)
optimizer = keras.optimizers.SGD(clipnorm=1.0)
```
## 사전 훈련된 층 재사용하기
### 케라스 모델 재사용하기
패션 MNIST 훈련 세트를 두 개로 나누어 보죠:
* `X_train_A`: 샌달과 셔츠(클래스 5와 6)을 제외한 모든 이미지
* `X_train_B`: 샌달과 셔츠 이미지 중 처음 200개만 가진 작은 훈련 세트
검증 세트와 테스트 세트도 이렇게 나눕니다. 하지만 이미지 개수는 제한하지 않습니다.
A 세트(8개의 클래스를 가진 분류 문제)에서 모델을 훈련하고 이를 재사용하여 B 세트(이진 분류)를 해결해 보겠습니다. A 작업에서 B 작업으로 약간의 지식이 전달되기를 기대합니다. 왜냐하면 A 세트의 클래스(스니커즈, 앵클 부츠, 코트, 티셔츠 등)가 B 세트에 있는 클래스(샌달과 셔츠)와 조금 비슷하기 때문입니다. 하지만 `Dense` 층을 사용하기 때문에 동일한 위치에 나타난 패턴만 재사용할 수 있습니다(반대로 합성곱 층은 훨씬 많은 정보를 전송합니다. 학습한 패턴을 이미지의 어느 위치에서나 감지할 수 있기 때문입니다. CNN 장에서 자세히 알아 보겠습니다).
```
def split_dataset(X, y):
y_5_or_6 = (y == 5) | (y == 6) # sandals or shirts
y_A = y[~y_5_or_6]
y_A[y_A > 6] -= 2 # class indices 7, 8, 9 should be moved to 5, 6, 7
y_B = (y[y_5_or_6] == 6).astype(np.float32) # binary classification task: is it a shirt (class 6)?
return ((X[~y_5_or_6], y_A),
(X[y_5_or_6], y_B))
(X_train_A, y_train_A), (X_train_B, y_train_B) = split_dataset(X_train, y_train)
(X_valid_A, y_valid_A), (X_valid_B, y_valid_B) = split_dataset(X_valid, y_valid)
(X_test_A, y_test_A), (X_test_B, y_test_B) = split_dataset(X_test, y_test)
X_train_B = X_train_B[:200]
y_train_B = y_train_B[:200]
X_train_A.shape
X_train_B.shape
y_train_A[:30]
y_train_B[:30]
tf.random.set_seed(42)
np.random.seed(42)
model_A = keras.models.Sequential()
model_A.add(keras.layers.Flatten(input_shape=[28, 28]))
for n_hidden in (300, 100, 50, 50, 50):
model_A.add(keras.layers.Dense(n_hidden, activation="selu"))
model_A.add(keras.layers.Dense(8, activation="softmax"))
model_A.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model_A.fit(X_train_A, y_train_A, epochs=20,
validation_data=(X_valid_A, y_valid_A))
model_A.save("my_model_A.h5")
model_B = keras.models.Sequential()
model_B.add(keras.layers.Flatten(input_shape=[28, 28]))
for n_hidden in (300, 100, 50, 50, 50):
model_B.add(keras.layers.Dense(n_hidden, activation="selu"))
model_B.add(keras.layers.Dense(1, activation="sigmoid"))
model_B.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model_B.fit(X_train_B, y_train_B, epochs=20,
validation_data=(X_valid_B, y_valid_B))
model.summary()
model_A = keras.models.load_model("my_model_A.h5")
model_B_on_A = keras.models.Sequential(model_A.layers[:-1])
model_B_on_A.add(keras.layers.Dense(1, activation="sigmoid"))
model_A_clone = keras.models.clone_model(model_A)
model_A_clone.set_weights(model_A.get_weights())
for layer in model_B_on_A.layers[:-1]:
layer.trainable = False
model_B_on_A.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=4,
validation_data=(X_valid_B, y_valid_B))
for layer in model_B_on_A.layers[:-1]:
layer.trainable = True
model_B_on_A.compile(loss="binary_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
history = model_B_on_A.fit(X_train_B, y_train_B, epochs=16,
validation_data=(X_valid_B, y_valid_B))
```
마지막 점수는 어떤가요?
```
model_B.evaluate(X_test_B, y_test_B)
model_B_on_A.evaluate(X_test_B, y_test_B)
```
훌륭하네요! 꽤 많은 정보를 전달했습니다: 오차율이 4배나 줄었네요!
```
(100 - 96.95) / (100 - 99.25)
```
# 고속 옵티마이저
## 모멘텀 옵티마이저
```
optimizer = keras.optimizers.SGD(lr=0.001, momentum=0.9)
```
## 네스테로프 가속 경사
```
optimizer = keras.optimizers.SGD(lr=0.001, momentum=0.9, nesterov=True)
```
## AdaGrad
```
optimizer = keras.optimizers.Adagrad(lr=0.001)
```
## RMSProp
```
optimizer = keras.optimizers.RMSprop(lr=0.001, rho=0.9)
```
## Adam 옵티마이저
```
optimizer = keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999)
```
## Adamax 옵티마이저
```
optimizer = keras.optimizers.Adamax(lr=0.001, beta_1=0.9, beta_2=0.999)
```
## Nadam 옵티마이저
```
optimizer = keras.optimizers.Nadam(lr=0.001, beta_1=0.9, beta_2=0.999)
```
## 학습률 스케줄링
### 거듭제곱 스케줄링
```lr = lr0 / (1 + steps / s)**c```
* 케라스는 `c=1`과 `s = 1 / decay`을 사용합니다
```
optimizer = keras.optimizers.SGD(lr=0.01, decay=1e-4)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
learning_rate = 0.01
decay = 1e-4
batch_size = 32
n_steps_per_epoch = len(X_train) // batch_size
epochs = np.arange(n_epochs)
lrs = learning_rate / (1 + decay * epochs * n_steps_per_epoch)
plt.plot(epochs, lrs, "o-")
plt.axis([0, n_epochs - 1, 0, 0.01])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Power Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
### 지수 기반 스케줄링
```lr = lr0 * 0.1**(epoch / s)```
```
def exponential_decay_fn(epoch):
return 0.01 * 0.1**(epoch / 20)
def exponential_decay(lr0, s):
def exponential_decay_fn(epoch):
return lr0 * 0.1**(epoch / s)
return exponential_decay_fn
exponential_decay_fn = exponential_decay(lr0=0.01, s=20)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 25
lr_scheduler = keras.callbacks.LearningRateScheduler(exponential_decay_fn)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, history.history["lr"], "o-")
plt.axis([0, n_epochs - 1, 0, 0.011])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Exponential Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
이 스케줄 함수는 두 번째 매개변수로 현재 학습률을 받을 수 있습니다:
```
def exponential_decay_fn(epoch, lr):
return lr * 0.1**(1 / 20)
```
에포크가 아니라 반복마다 학습률을 업데이트하려면 사용자 정의 콜백 클래스를 작성해야 합니다:
```
K = keras.backend
class ExponentialDecay(keras.callbacks.Callback):
def __init__(self, s=40000):
super().__init__()
self.s = s
def on_batch_begin(self, batch, logs=None):
# 노트: 에포크마다 `batch` 매개변수가 재설정됩니다
lr = K.get_value(self.model.optimizer.lr)
K.set_value(self.model.optimizer.lr, lr * 0.1**(1 / s))
def on_epoch_end(self, epoch, logs=None):
logs = logs or {}
logs['lr'] = K.get_value(self.model.optimizer.lr)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
lr0 = 0.01
optimizer = keras.optimizers.Nadam(lr=lr0)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
s = 20 * len(X_train) // 32 # 20 에포크 동안 스텝 횟수 (배치 크기 = 32)
exp_decay = ExponentialDecay(s)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[exp_decay])
n_steps = n_epochs * len(X_train) // 32
steps = np.arange(n_steps)
lrs = lr0 * 0.1**(steps / s)
plt.plot(steps, lrs, "-", linewidth=2)
plt.axis([0, n_steps - 1, 0, lr0 * 1.1])
plt.xlabel("Batch")
plt.ylabel("Learning Rate")
plt.title("Exponential Scheduling (per batch)", fontsize=14)
plt.grid(True)
plt.show()
```
### 기간별 고정 스케줄링
```
def piecewise_constant_fn(epoch):
if epoch < 5:
return 0.01
elif epoch < 15:
return 0.005
else:
return 0.001
def piecewise_constant(boundaries, values):
boundaries = np.array([0] + boundaries)
values = np.array(values)
def piecewise_constant_fn(epoch):
return values[np.argmax(boundaries > epoch) - 1]
return piecewise_constant_fn
piecewise_constant_fn = piecewise_constant([5, 15], [0.01, 0.005, 0.001])
lr_scheduler = keras.callbacks.LearningRateScheduler(piecewise_constant_fn)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, [piecewise_constant_fn(epoch) for epoch in history.epoch], "o-")
plt.axis([0, n_epochs - 1, 0, 0.011])
plt.xlabel("Epoch")
plt.ylabel("Learning Rate")
plt.title("Piecewise Constant Scheduling", fontsize=14)
plt.grid(True)
plt.show()
```
### 성능 기반 스케줄링
```
tf.random.set_seed(42)
np.random.seed(42)
lr_scheduler = keras.callbacks.ReduceLROnPlateau(factor=0.5, patience=5)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
optimizer = keras.optimizers.SGD(lr=0.02, momentum=0.9)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid),
callbacks=[lr_scheduler])
plt.plot(history.epoch, history.history["lr"], "bo-")
plt.xlabel("Epoch")
plt.ylabel("Learning Rate", color='b')
plt.tick_params('y', colors='b')
plt.gca().set_xlim(0, n_epochs - 1)
plt.grid(True)
ax2 = plt.gca().twinx()
ax2.plot(history.epoch, history.history["val_loss"], "r^-")
ax2.set_ylabel('Validation Loss', color='r')
ax2.tick_params('y', colors='r')
plt.title("Reduce LR on Plateau", fontsize=14)
plt.show()
```
### tf.keras 스케줄러
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
s = 20 * len(X_train) // 32 # number of steps in 20 epochs (batch size = 32)
learning_rate = keras.optimizers.schedules.ExponentialDecay(0.01, s, 0.1)
optimizer = keras.optimizers.SGD(learning_rate)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 25
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
구간별 고정 스케줄링은 다음을 사용하세요:
```
learning_rate = keras.optimizers.schedules.PiecewiseConstantDecay(
boundaries=[5. * n_steps_per_epoch, 15. * n_steps_per_epoch],
values=[0.01, 0.005, 0.001])
```
### 1사이클 스케줄링
```
K = keras.backend
class ExponentialLearningRate(keras.callbacks.Callback):
def __init__(self, factor):
self.factor = factor
self.rates = []
self.losses = []
def on_batch_end(self, batch, logs):
self.rates.append(K.get_value(self.model.optimizer.lr))
self.losses.append(logs["loss"])
K.set_value(self.model.optimizer.lr, self.model.optimizer.lr * self.factor)
def find_learning_rate(model, X, y, epochs=1, batch_size=32, min_rate=10**-5, max_rate=10):
init_weights = model.get_weights()
iterations = len(X) // batch_size * epochs
factor = np.exp(np.log(max_rate / min_rate) / iterations)
init_lr = K.get_value(model.optimizer.lr)
K.set_value(model.optimizer.lr, min_rate)
exp_lr = ExponentialLearningRate(factor)
history = model.fit(X, y, epochs=epochs, batch_size=batch_size,
callbacks=[exp_lr])
K.set_value(model.optimizer.lr, init_lr)
model.set_weights(init_weights)
return exp_lr.rates, exp_lr.losses
def plot_lr_vs_loss(rates, losses):
plt.plot(rates, losses)
plt.gca().set_xscale('log')
plt.hlines(min(losses), min(rates), max(rates))
plt.axis([min(rates), max(rates), min(losses), (losses[0] + min(losses)) / 2])
plt.xlabel("Learning rate")
plt.ylabel("Loss")
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer=keras.optimizers.SGD(lr=1e-3),
metrics=["accuracy"])
batch_size = 128
rates, losses = find_learning_rate(model, X_train_scaled, y_train, epochs=1, batch_size=batch_size)
plot_lr_vs_loss(rates, losses)
class OneCycleScheduler(keras.callbacks.Callback):
def __init__(self, iterations, max_rate, start_rate=None,
last_iterations=None, last_rate=None):
self.iterations = iterations
self.max_rate = max_rate
self.start_rate = start_rate or max_rate / 10
self.last_iterations = last_iterations or iterations // 10 + 1
self.half_iteration = (iterations - self.last_iterations) // 2
self.last_rate = last_rate or self.start_rate / 1000
self.iteration = 0
def _interpolate(self, iter1, iter2, rate1, rate2):
return ((rate2 - rate1) * (self.iteration - iter1)
/ (iter2 - iter1) + rate1)
def on_batch_begin(self, batch, logs):
if self.iteration < self.half_iteration:
rate = self._interpolate(0, self.half_iteration, self.start_rate, self.max_rate)
elif self.iteration < 2 * self.half_iteration:
rate = self._interpolate(self.half_iteration, 2 * self.half_iteration,
self.max_rate, self.start_rate)
else:
rate = self._interpolate(2 * self.half_iteration, self.iterations,
self.start_rate, self.last_rate)
rate = max(rate, self.last_rate)
self.iteration += 1
K.set_value(self.model.optimizer.lr, rate)
n_epochs = 25
onecycle = OneCycleScheduler(len(X_train) // batch_size * n_epochs, max_rate=0.05)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs, batch_size=batch_size,
validation_data=(X_valid_scaled, y_valid),
callbacks=[onecycle])
```
# 규제를 사용해 과대적합 피하기
## $\ell_1$과 $\ell_2$ 규제
```
layer = keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01))
# or l1(0.1) for ℓ1 regularization with a factor or 0.1
# or l1_l2(0.1, 0.01) for both ℓ1 and ℓ2 regularization, with factors 0.1 and 0.01 respectively
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dense(300, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01)),
keras.layers.Dense(100, activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01)),
keras.layers.Dense(10, activation="softmax",
kernel_regularizer=keras.regularizers.l2(0.01))
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
from functools import partial
RegularizedDense = partial(keras.layers.Dense,
activation="elu",
kernel_initializer="he_normal",
kernel_regularizer=keras.regularizers.l2(0.01))
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
RegularizedDense(300),
RegularizedDense(100),
RegularizedDense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
## 드롭아웃
```
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(300, activation="elu", kernel_initializer="he_normal"),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(100, activation="elu", kernel_initializer="he_normal"),
keras.layers.Dropout(rate=0.2),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
## 알파 드롭아웃
```
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(300, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal"),
keras.layers.AlphaDropout(rate=0.2),
keras.layers.Dense(10, activation="softmax")
])
optimizer = keras.optimizers.SGD(lr=0.01, momentum=0.9, nesterov=True)
model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
n_epochs = 20
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
model.evaluate(X_test_scaled, y_test)
model.evaluate(X_train_scaled, y_train)
history = model.fit(X_train_scaled, y_train)
```
## MC 드롭아웃
```
tf.random.set_seed(42)
np.random.seed(42)
y_probas = np.stack([model(X_test_scaled, training=True)
for sample in range(100)])
y_proba = y_probas.mean(axis=0)
y_std = y_probas.std(axis=0)
np.round(model.predict(X_test_scaled[:1]), 2)
np.round(y_probas[:, :1], 2)
np.round(y_proba[:1], 2)
y_std = y_probas.std(axis=0)
np.round(y_std[:1], 2)
y_pred = np.argmax(y_proba, axis=1)
accuracy = np.sum(y_pred == y_test) / len(y_test)
accuracy
class MCDropout(keras.layers.Dropout):
def call(self, inputs):
return super().call(inputs, training=True)
class MCAlphaDropout(keras.layers.AlphaDropout):
def call(self, inputs):
return super().call(inputs, training=True)
tf.random.set_seed(42)
np.random.seed(42)
mc_model = keras.models.Sequential([
MCAlphaDropout(layer.rate) if isinstance(layer, keras.layers.AlphaDropout) else layer
for layer in model.layers
])
mc_model.summary()
optimizer = keras.optimizers.SGD(lr=0.01, momentum=0.9, nesterov=True)
mc_model.compile(loss="sparse_categorical_crossentropy", optimizer=optimizer, metrics=["accuracy"])
mc_model.set_weights(model.get_weights())
```
이제 MC 드롭아웃을 모델에 사용할 수 있습니다:
```
np.round(np.mean([mc_model.predict(X_test_scaled[:1]) for sample in range(100)], axis=0), 2)
```
## 맥스 노름
```
layer = keras.layers.Dense(100, activation="selu", kernel_initializer="lecun_normal",
kernel_constraint=keras.constraints.max_norm(1.))
MaxNormDense = partial(keras.layers.Dense,
activation="selu", kernel_initializer="lecun_normal",
kernel_constraint=keras.constraints.max_norm(1.))
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=[28, 28]),
MaxNormDense(300),
MaxNormDense(100),
keras.layers.Dense(10, activation="softmax")
])
model.compile(loss="sparse_categorical_crossentropy", optimizer="nadam", metrics=["accuracy"])
n_epochs = 2
history = model.fit(X_train_scaled, y_train, epochs=n_epochs,
validation_data=(X_valid_scaled, y_valid))
```
# 연습문제 해답
## 1. to 7.
부록 A 참조.
## 8. CIFAR10에서 딥러닝
### a.
*문제: 100개의 뉴런을 가진 은닉층 20개로 심층 신경망을 만들어보세요(너무 많은 것 같지만 이 연습문제의 핵심입니다). He 초기화와 ELU 활성화 함수를 사용하세요.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
activation="elu",
kernel_initializer="he_normal"))
```
### b.
*문제: Nadam 옵티마이저와 조기 종료를 사용하여 CIFAR10 데이터셋에 이 네트워크를 훈련하세요. `keras.datasets.cifar10.load_ data()`를 사용하여 데이터를 적재할 수 있습니다. 이 데이터셋은 10개의 클래스와 32×32 크기의 컬러 이미지 60,000개로 구성됩니다(50,000개는 훈련, 10,000개는 테스트). 따라서 10개의 뉴런과 소프트맥스 활성화 함수를 사용하는 출력층이 필요합니다. 모델 구조와 하이퍼파라미터를 바꿀 때마다 적절한 학습률을 찾아야 한다는 것을 기억하세요.*
모델에 출력층을 추가합니다:
```
model.add(keras.layers.Dense(10, activation="softmax"))
```
학습률 5e-5인 Nadam 옵티마이저를 사용해 보죠. 학습률 1e-5, 3e-5, 1e-4, 3e-4, 1e-3, 3e-3, 1e-2를 테스트하고 10번의 에포크 동안 (아래 텐서보드 콜백으로) 학습 곡선을 비교해 보았습니다. 학습률 3e-5와 1e-4가 꽤 좋았기 때문에 5e-5를 시도해 보았고 조금 더 나은 결과를 냈습니다.
```
optimizer = keras.optimizers.Nadam(lr=5e-5)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
```
CIFAR10 데이터셋을 로드하죠. 조기 종료를 사용하기 때문에 검증 세트가 필요합니다. 원본 훈련 세트에서 처음 5,000개를 검증 세트로 사용하겠습니다:
```
(X_train_full, y_train_full), (X_test, y_test) = keras.datasets.cifar10.load_data()
X_train = X_train_full[5000:]
y_train = y_train_full[5000:]
X_valid = X_train_full[:5000]
y_valid = y_train_full[:5000]
```
이제 콜백을 만들고 모델을 훈련합니다:
```
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_model.h5", save_best_only=True)
run_index = 1 # 모델을 훈련할 때마다 증가시킴
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
%tensorboard --logdir=./my_cifar10_logs --port=6006
model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_model.h5")
model.evaluate(X_valid, y_valid)
```
가장 낮은 검증 손실을 내는 모델은 검증 세트에서 약 47% 정확도를 얻었습니다. 이 검증 점수에 도달하는데 39번의 에포크가 걸렸습니다. (GPU가 없는) 제 노트북에서 에포크당 약 10초 정도 걸렸습니다. 배치 정규화를 사용해 성능을 올릴 수 있는지 확인해 보죠.
### c.
*문제: 배치 정규화를 추가하고 학습 곡선을 비교해보세요. 이전보다 빠르게 수렴하나요? 더 좋은 모델이 만들어지나요? 훈련 속도에는 어떤 영향을 미치나요?*
다음 코드는 위의 코드와 배우 비슷합니다. 몇 가지 다른 점은 아래와 같습니다:
* 출력층을 제외하고 모든 `Dense` 층 다음에 (활성화 함수 전에) BN 층을 추가했습니다. 처음 은닉층 전에도 BN 층을 추가했습니다.
* 학습률을 5e-4로 바꾸었습니다. 1e-5, 3e-5, 5e-5, 1e-4, 3e-4, 5e-4, 1e-3, 3e-3를 시도해 보고 20번 에포크 후에 검증 세트 성능이 가장 좋은 것을 선택했습니다.
* run_logdir를 run_bn_* 으로 이름을 바꾸고 모델 파일 이름을 my_cifar10_bn_model.h5로 변경했습니다.
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
model.add(keras.layers.BatchNormalization())
for _ in range(20):
model.add(keras.layers.Dense(100, kernel_initializer="he_normal"))
model.add(keras.layers.BatchNormalization())
model.add(keras.layers.Activation("elu"))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(lr=5e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_bn_model.h5", save_best_only=True)
run_index = 1 # 모델을 훈련할 때마다 증가시킴
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_bn_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
model.fit(X_train, y_train, epochs=100,
validation_data=(X_valid, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_bn_model.h5")
model.evaluate(X_valid, y_valid)
```
* *이전보다 빠르게 수렴하나요?* 훨씬 빠릅니다! 이전 모델은 가장 낮은 검증 손실에 도달하기 위해 39 에포크가 걸렸지만 BN을 사용한 새 모델은 18 에포크가 걸렸습니다. 이전 모델보다 두 배 이상 빠릅니다. BN 층은 훈련을 안정적으로 수행하고 더 큰 학습률을 사용할 수 있기 때문에 수렴이 빨라졌습니다.
* *BN이 더 좋은 모델을 만드나요?* 네! 최종 모델의 성능이 47%가 아니라 55% 정확도로 더 좋습니다. 이는 아주 좋은 모델이 아니지만 적어도 이전보다는 낫습니다(합성곱 신경망이 더 낫겠지만 이는 다른 주제입니다. 14장을 참고하세요).
* *BN이 훈련 속도에 영향을 미치나요?* 모델이 두 배나 빠르게 수렴했지만 각 에포크는 10초가 아니라 16초가 걸렸습니다. BN 층에서 추가된 계산 때문입니다. 따라서 전체적으로 에포크 횟수가 50% 정도 줄었지만 훈련 시간(탁상 시계 시간)은 30% 정도 줄었습니다. 결국 크게 향상되었습니다!
### d.
*문제: 배치 정규화를 SELU로 바꾸어보세요. 네트워크가 자기 정규화하기 위해 필요한 변경 사항을 적용해보세요(즉, 입력 특성 표준화, 르쿤 정규분포 초기화, 완전 연결 층만 순차적으로 쌓은 심층 신경망 등).*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(lr=7e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_selu_model.h5", save_best_only=True)
run_index = 1 # 모델을 훈련할 때마다 증가시킴
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_selu_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
X_means = X_train.mean(axis=0)
X_stds = X_train.std(axis=0)
X_train_scaled = (X_train - X_means) / X_stds
X_valid_scaled = (X_valid - X_means) / X_stds
X_test_scaled = (X_test - X_means) / X_stds
model.fit(X_train_scaled, y_train, epochs=100,
validation_data=(X_valid_scaled, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_selu_model.h5")
model.evaluate(X_valid_scaled, y_valid)
model = keras.models.load_model("my_cifar10_selu_model.h5")
model.evaluate(X_valid_scaled, y_valid)
```
51.4% 정확도를 얻었습니다. 원래 모델보다 더 좋습니다. 하지만 배치 정규화를 사용한 모델만큼 좋지는 않습니다. 최고의 모델에 도달하는데 13 에포크가 걸렸습니다. 이는 원본 모델이나 BN 모델보다 더 빠른 것입니다. 각 에포크는 원본 모델처럼 10초만 걸렸습니다. 따라서 이 모델이 지금까지 가장 빠른 모델입니다(에포크와 탁상 시계 기준으로).
### e.
*문제: 알파 드롭아웃으로 모델에 규제를 적용해보세요. 그다음 모델을 다시 훈련하지 않고 MC 드롭아웃으로 더 높은 정확도를 얻을 수 있는지 확인해보세요.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.Nadam(lr=5e-4)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(patience=20)
model_checkpoint_cb = keras.callbacks.ModelCheckpoint("my_cifar10_alpha_dropout_model.h5", save_best_only=True)
run_index = 1 # 모델을 훈련할 때마다 증가시킴
run_logdir = os.path.join(os.curdir, "my_cifar10_logs", "run_alpha_dropout_{:03d}".format(run_index))
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
callbacks = [early_stopping_cb, model_checkpoint_cb, tensorboard_cb]
X_means = X_train.mean(axis=0)
X_stds = X_train.std(axis=0)
X_train_scaled = (X_train - X_means) / X_stds
X_valid_scaled = (X_valid - X_means) / X_stds
X_test_scaled = (X_test - X_means) / X_stds
model.fit(X_train_scaled, y_train, epochs=100,
validation_data=(X_valid_scaled, y_valid),
callbacks=callbacks)
model = keras.models.load_model("my_cifar10_alpha_dropout_model.h5")
model.evaluate(X_valid_scaled, y_valid)
```
이 모델은 검증 세트에서 50.8% 정확도에 도달합니다. 드롭아웃이 없을 때보다(51.4%) 조금 더 나쁩니다. 하이퍼파라미터 탐색을 좀 많이 수행해 보면 더 나아 질 수 있습니다(드롭아웃 비율 5%, 10%, 20%, 40%과 학습률 1e-4, 3e-4, 5e-4, 1e-3을 시도했습니다). 하지만 이 경우에는 크지 않을 것 같습니다.
이제 MC 드롭아웃을 사용해 보죠. 앞서 사용한 `MCAlphaDropout` 클래스를 복사해 사용하겠습니다:
```
class MCAlphaDropout(keras.layers.AlphaDropout):
def call(self, inputs):
return super().call(inputs, training=True)
```
방금 훈련했던 모델과 (같은 가중치를 가진) 동일한 새로운 모델을 만들어 보죠. 하지만 `AlphaDropout` 층 대신 `MCAlphaDropout` 드롭아웃 층을 사용합니다:
```
mc_model = keras.models.Sequential([
MCAlphaDropout(layer.rate) if isinstance(layer, keras.layers.AlphaDropout) else layer
for layer in model.layers
])
```
그다음 몇 가지 유틸리티 함수를 추가합니다. 첫 번째 함수는 모델을 여러 번 실행합니다(기본적으로 10번). 그다음 평균한 예측 클래스 확률을 반환합니다. 두 번째 함수는 이 평균 확률을 사용해 각 샘플의 클래스를 예측합니다:
```
def mc_dropout_predict_probas(mc_model, X, n_samples=10):
Y_probas = [mc_model.predict(X) for sample in range(n_samples)]
return np.mean(Y_probas, axis=0)
def mc_dropout_predict_classes(mc_model, X, n_samples=10):
Y_probas = mc_dropout_predict_probas(mc_model, X, n_samples)
return np.argmax(Y_probas, axis=1)
```
이제 검증 세트의 모든 샘플에 대해 예측을 만들고 정확도를 계산해 보죠:
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
y_pred = mc_dropout_predict_classes(mc_model, X_valid_scaled)
accuracy = np.mean(y_pred == y_valid[:, 0])
accuracy
```
이 경우에는 실제적인 정확도 향상이 없습니다(50.8%에서 50.9%).
따라서 이 연습문에서 얻은 최상의 모델은 배치 정규화 모델입니다.
### f.
*문제: 1사이클 스케줄링으로 모델을 다시 훈련하고 훈련 속도와 모델 정확도가 향상되는지 확인해보세요.*
```
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.SGD(lr=1e-3)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
batch_size = 128
rates, losses = find_learning_rate(model, X_train_scaled, y_train, epochs=1, batch_size=batch_size)
plot_lr_vs_loss(rates, losses)
plt.axis([min(rates), max(rates), min(losses), (losses[0] + min(losses)) / 1.4])
keras.backend.clear_session()
tf.random.set_seed(42)
np.random.seed(42)
model = keras.models.Sequential()
model.add(keras.layers.Flatten(input_shape=[32, 32, 3]))
for _ in range(20):
model.add(keras.layers.Dense(100,
kernel_initializer="lecun_normal",
activation="selu"))
model.add(keras.layers.AlphaDropout(rate=0.1))
model.add(keras.layers.Dense(10, activation="softmax"))
optimizer = keras.optimizers.SGD(lr=1e-2)
model.compile(loss="sparse_categorical_crossentropy",
optimizer=optimizer,
metrics=["accuracy"])
n_epochs = 15
onecycle = OneCycleScheduler(len(X_train_scaled) // batch_size * n_epochs, max_rate=0.05)
history = model.fit(X_train_scaled, y_train, epochs=n_epochs, batch_size=batch_size,
validation_data=(X_valid_scaled, y_valid),
callbacks=[onecycle])
```
1사이클 방식을 사용해 모델을 15에포크 동안 훈련했습니다. (큰 배치 크기 덕분에) 각 에포크는 3초만 걸렸습니다. 이는 지금까지 훈련한 가장 빠른 모델보다 3배나 더 빠릅니다. 또한 모델 성능도 올라갔습니다(50.8%에서 52.8%). 배치 정규화 모델이 조금 더 성능이 높지만 훈련 속도가 더 느립니다.
| github_jupyter |
```
# look at tools/set_up_magics.ipynb
yandex_metrica_allowed = True ; get_ipython().run_cell('# one_liner_str\n\nget_ipython().run_cell_magic(\'javascript\', \'\', \'// setup cpp code highlighting\\nIPython.CodeCell.options_default.highlight_modes["text/x-c++src"] = {\\\'reg\\\':[/^%%cpp/]} ;\')\n\n# creating magics\nfrom IPython.core.magic import register_cell_magic, register_line_magic\nfrom IPython.display import display, Markdown, HTML\nimport argparse\nfrom subprocess import Popen, PIPE\nimport random\nimport sys\nimport os\nimport re\nimport signal\nimport shutil\nimport shlex\nimport glob\n\n@register_cell_magic\ndef save_file(args_str, cell, line_comment_start="#"):\n parser = argparse.ArgumentParser()\n parser.add_argument("fname")\n parser.add_argument("--ejudge-style", action="store_true")\n args = parser.parse_args(args_str.split())\n \n cell = cell if cell[-1] == \'\\n\' or args.no_eof_newline else cell + "\\n"\n cmds = []\n with open(args.fname, "w") as f:\n f.write(line_comment_start + " %%cpp " + args_str + "\\n")\n for line in cell.split("\\n"):\n line_to_write = (line if not args.ejudge_style else line.rstrip()) + "\\n"\n if line.startswith("%"):\n run_prefix = "%run "\n if line.startswith(run_prefix):\n cmds.append(line[len(run_prefix):].strip())\n f.write(line_comment_start + " " + line_to_write)\n continue\n run_prefix = "%# "\n if line.startswith(run_prefix):\n f.write(line_comment_start + " " + line_to_write)\n continue\n raise Exception("Unknown %%save_file subcommand: \'%s\'" % line)\n else:\n f.write(line_to_write)\n f.write("" if not args.ejudge_style else line_comment_start + r" line without \\n")\n for cmd in cmds:\n display(Markdown("Run: `%s`" % cmd))\n get_ipython().system(cmd)\n\n@register_cell_magic\ndef cpp(fname, cell):\n save_file(fname, cell, "//")\n\n@register_cell_magic\ndef asm(fname, cell):\n save_file(fname, cell, "//")\n \n@register_cell_magic\ndef makefile(fname, cell):\n assert not fname\n save_file("makefile", cell.replace(" " * 4, "\\t"))\n \n@register_line_magic\ndef p(line):\n try:\n expr, comment = line.split(" #")\n display(Markdown("`{} = {}` # {}".format(expr.strip(), eval(expr), comment.strip())))\n except:\n display(Markdown("{} = {}".format(line, eval(line))))\n \ndef show_file(file, clear_at_begin=True, return_html_string=False):\n if clear_at_begin:\n get_ipython().system("truncate --size 0 " + file)\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n elem.innerText = xmlhttp.responseText;\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (errors___OBJ__ < 10 && !entrance___OBJ__) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n \n <font color="white"> <tt>\n <p id="__OBJ__" style="font-size: 16px; border:3px #333333 solid; background: #333333; border-radius: 10px; padding: 10px; "></p>\n </tt> </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__ -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n \nBASH_POPEN_TMP_DIR = "./bash_popen_tmp"\n \ndef bash_popen_terminate_all():\n for p in globals().get("bash_popen_list", []):\n print("Terminate pid=" + str(p.pid), file=sys.stderr)\n p.terminate()\n globals()["bash_popen_list"] = []\n if os.path.exists(BASH_POPEN_TMP_DIR):\n shutil.rmtree(BASH_POPEN_TMP_DIR)\n\nbash_popen_terminate_all() \n\ndef bash_popen(cmd):\n if not os.path.exists(BASH_POPEN_TMP_DIR):\n os.mkdir(BASH_POPEN_TMP_DIR)\n h = os.path.join(BASH_POPEN_TMP_DIR, str(random.randint(0, 1e18)))\n stdout_file = h + ".out.html"\n stderr_file = h + ".err.html"\n run_log_file = h + ".fin.html"\n \n stdout = open(stdout_file, "wb")\n stdout = open(stderr_file, "wb")\n \n html = """\n <table width="100%">\n <colgroup>\n <col span="1" style="width: 70px;">\n <col span="1">\n </colgroup> \n <tbody>\n <tr> <td><b>STDOUT</b></td> <td> {stdout} </td> </tr>\n <tr> <td><b>STDERR</b></td> <td> {stderr} </td> </tr>\n <tr> <td><b>RUN LOG</b></td> <td> {run_log} </td> </tr>\n </tbody>\n </table>\n """.format(\n stdout=show_file(stdout_file, return_html_string=True),\n stderr=show_file(stderr_file, return_html_string=True),\n run_log=show_file(run_log_file, return_html_string=True),\n )\n \n cmd = """\n bash -c {cmd} &\n pid=$!\n echo "Process started! pid=${{pid}}" > {run_log_file}\n wait ${{pid}}\n echo "Process finished! exit_code=$?" >> {run_log_file}\n """.format(cmd=shlex.quote(cmd), run_log_file=run_log_file)\n # print(cmd)\n display(HTML(html))\n \n p = Popen(["bash", "-c", cmd], stdin=PIPE, stdout=stdout, stderr=stdout)\n \n bash_popen_list.append(p)\n return p\n\n\n@register_line_magic\ndef bash_async(line):\n bash_popen(line)\n \n \ndef show_log_file(file, return_html_string=False):\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function halt__OBJ__(elem, color)\n {\n elem.setAttribute("style", "font-size: 14px; background: " + color + "; padding: 10px; border: 3px; border-radius: 5px; color: white; "); \n }\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n if (entrance___OBJ__ < 0) {\n entrance___OBJ__ = 0;\n }\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n if (elem.innerHTML != xmlhttp.responseText) {\n elem.innerHTML = xmlhttp.responseText;\n }\n if (elem.innerHTML.includes("Process finished.")) {\n halt__OBJ__(elem, "#333333");\n } else {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (!entrance___OBJ__) {\n if (errors___OBJ__ < 6) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n } else {\n halt__OBJ__(elem, "#994444");\n }\n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n\n <p id="__OBJ__" style="font-size: 14px; background: #000000; padding: 10px; border: 3px; border-radius: 5px; color: white; ">\n </p>\n \n </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__.md -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n\n \nclass TInteractiveLauncher:\n tmp_path = "./interactive_launcher_tmp"\n def __init__(self, cmd):\n try:\n os.mkdir(TInteractiveLauncher.tmp_path)\n except:\n pass\n name = str(random.randint(0, 1e18))\n self.inq_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".inq")\n self.log_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".log")\n \n os.mkfifo(self.inq_path)\n open(self.log_path, \'w\').close()\n open(self.log_path + ".md", \'w\').close()\n\n self.pid = os.fork()\n if self.pid == -1:\n print("Error")\n if self.pid == 0:\n exe_cands = glob.glob("../tools/launcher.py") + glob.glob("../../tools/launcher.py")\n assert(len(exe_cands) == 1)\n assert(os.execvp("python3", ["python3", exe_cands[0], "-l", self.log_path, "-i", self.inq_path, "-c", cmd]) == 0)\n self.inq_f = open(self.inq_path, "w")\n interactive_launcher_opened_set.add(self.pid)\n show_log_file(self.log_path)\n\n def write(self, s):\n s = s.encode()\n assert len(s) == os.write(self.inq_f.fileno(), s)\n \n def get_pid(self):\n n = 100\n for i in range(n):\n try:\n return int(re.findall(r"PID = (\\d+)", open(self.log_path).readline())[0])\n except:\n if i + 1 == n:\n raise\n time.sleep(0.1)\n \n def input_queue_path(self):\n return self.inq_path\n \n def close(self):\n self.inq_f.close()\n os.waitpid(self.pid, 0)\n os.remove(self.inq_path)\n # os.remove(self.log_path)\n self.inq_path = None\n self.log_path = None \n interactive_launcher_opened_set.remove(self.pid)\n self.pid = None\n \n @staticmethod\n def terminate_all():\n if "interactive_launcher_opened_set" not in globals():\n globals()["interactive_launcher_opened_set"] = set()\n global interactive_launcher_opened_set\n for pid in interactive_launcher_opened_set:\n print("Terminate pid=" + str(pid), file=sys.stderr)\n os.kill(pid, signal.SIGKILL)\n os.waitpid(pid, 0)\n interactive_launcher_opened_set = set()\n if os.path.exists(TInteractiveLauncher.tmp_path):\n shutil.rmtree(TInteractiveLauncher.tmp_path)\n \nTInteractiveLauncher.terminate_all()\n \nyandex_metrica_allowed = bool(globals().get("yandex_metrica_allowed", False))\nif yandex_metrica_allowed:\n display(HTML(\'\'\'<!-- YANDEX_METRICA_BEGIN -->\n <script type="text/javascript" >\n (function(m,e,t,r,i,k,a){m[i]=m[i]||function(){(m[i].a=m[i].a||[]).push(arguments)};\n m[i].l=1*new Date();k=e.createElement(t),a=e.getElementsByTagName(t)[0],k.async=1,k.src=r,a.parentNode.insertBefore(k,a)})\n (window, document, "script", "https://mc.yandex.ru/metrika/tag.js", "ym");\n\n ym(59260609, "init", {\n clickmap:true,\n trackLinks:true,\n accurateTrackBounce:true\n });\n </script>\n <noscript><div><img src="https://mc.yandex.ru/watch/59260609" style="position:absolute; left:-9999px;" alt="" /></div></noscript>\n <!-- YANDEX_METRICA_END -->\'\'\'))\n\ndef make_oneliner():\n html_text = \'("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "")\'\n html_text += \' + "<""!-- MAGICS_SETUP_PRINTING_END -->"\'\n return \'\'.join([\n \'# look at tools/set_up_magics.ipynb\\n\',\n \'yandex_metrica_allowed = True ; get_ipython().run_cell(%s);\' % repr(one_liner_str),\n \'display(HTML(%s))\' % html_text,\n \' #\'\'MAGICS_SETUP_END\'\n ])\n \n\n');display(HTML(("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "") + "<""!-- MAGICS_SETUP_PRINTING_END -->")) #MAGICS_SETUP_END
```
# От С к С++
Будем рассматривать минимально необходимое подмножество С++ на мой взгляд
До этого мы рассматривали подмножество языка С полностью совместимое с С++ (с 20м стандартом).
Рассмотрим на примере стека, как он может быть написан на С и на С++ и как связаны между собой реализации.
```
%%cpp main.c
%run clang -std=c99 -Wall -Werror -fsanitize=address main.c -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
typedef struct stack {
int* a;
int sz;
int max_sz;
} stack_t;
void init_stack(stack_t* stack) {
*stack = (stack_t){0};
}
void destroy_stack(stack_t* stack) {
free(stack->a);
}
void push_stack(stack_t* stack, int elem) {
if (stack->sz == stack->max_sz) {
stack->max_sz += (stack->max_sz == 0);
stack->max_sz *= 2;
(*stack).a = realloc(stack->a, stack->max_sz * sizeof(int));
}
stack->a[stack->sz++] = elem;
}
int top_stack(stack_t* stack) {
return stack->a[stack->sz - 1];
}
void pop_stack(stack_t* stack) {
--stack->sz;
}
int main() {
stack_t* s = (stack_t*)malloc(sizeof(stack_t));
init_stack(s);
push_stack(s, 123);
push_stack(s, 42);
assert(top_stack(s) == 42);
pop_stack(s);
assert(top_stack(s) == 123);
destroy_stack(s);
free(s);
return 0;
}
```
1) Добавим методы
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
#include <new> // Важно для placement new!
// Больше не нужны typedef
struct stack_t {
// Поля такие же
int* a;
int sz;
int max_sz;
// Декларируем методы
stack_t(); // конструктор
~stack_t(); // деструктор
void push(int elem);
int top();
void pop();
};
// void init_stack(stack_t* stack) {
// *stack = (stack_t){0};
// }
stack_t::stack_t() {
this->a = nullptr;
this->sz = 0;
this->max_sz = 0;
}
// void destroy_stack(stack_t* stack) {
// free(stack->a);
// }
stack_t::~stack_t() {
free(this->a);
}
// void push_stack(stack_t* stack, int elem) {
// if (stack->sz == stack->max_sz) {
// stack->max_sz += (stack->max_sz == 0);
// stack->max_sz *= 2;
// (*stack).a = realloc(stack->a, stack->max_sz * sizeof(int));
// }
// stack->a[stack->sz++] = elem;
// }
void stack_t::push(int elem) {
if (this->sz == this->max_sz) {
this->max_sz += (this->max_sz == 0);
this->max_sz *= 2;
this->a = (int*)realloc(this->a, this->max_sz * sizeof(int));
}
this->a[this->sz++] = elem;
}
// int top_stack(stack_t* stack) {
// return stack->a[stack->sz - 1];
// }
int stack_t::top() {
return this->a[this->sz - 1];
}
// void pop_stack(stack_t* stack) {
// --stack->sz;
// }
void stack_t::pop() {
--this->sz;
}
int main() {
stack_t* s = (stack_t*)malloc(sizeof(stack_t));
new (s) stack_t; // init_stack(s);
s->push(123); // push_stack(s, 123);
s->push(42); // push_stack(s, 42);
assert(s->top() == 42); // assert(top_stack(s) == 42);
s->pop(); // pop_stack(s);
assert(s->top() == 123); // assert(top_stack(s) == 123);
s->~stack_t(); // destroy_stack(s);
free(s);
return 0;
}
```
2) Уберем избыточность
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
// Больше не нужны typedef
struct stack_t {
// Поля такие же
int* a;
int sz;
int max_sz;
// Декларируем методы
stack_t(); // конструктор
~stack_t(); // деструктор
void push(int elem);
int top();
void pop();
};
// stack_t::stack_t() {
// this->a = nullptr;
// this->sz = 0;
// this->max_sz = 0;
// }
stack_t::stack_t() {
a = nullptr;
sz = 0;
max_sz = 0;
}
// stack_t::~stack_t() {
// free(this->a);
// }
stack_t::~stack_t() {
free(a);
}
// void stack_t::push(int elem) {
// if (this->sz == this->max_sz) {
// this->max_sz += (this->max_sz == 0);
// this->max_sz *= 2;
// this->a = (int*)realloc(this->a, this->max_sz * sizeof(int));
// }
// this->a[this->sz++] = elem;
// }
void stack_t::push(int elem) {
if (sz == max_sz) {
max_sz += (max_sz == 0);
max_sz *= 2;
a = (int*)realloc(a, max_sz * sizeof(int));
}
a[sz++] = elem;
}
// int stack_t::top() {
// return this->a[this->sz - 1];
// }
int stack_t::top() {
return a[sz - 1];
}
// void stack_t::pop() {
// --this->sz;
// }
void stack_t::pop() {
--sz;
}
int main() {
// variant 1
{
stack_t* s = new stack_t; // stack_t* s = (stack_t*)malloc(sizeof(stack_t));
// new ((void*)s) stack_t;
s->push(123);
s->push(42);
assert(s->top() == 42);
s->pop();
assert(s->top() == 123);
delete s; // s->~stack_t();
// free(s);
}
// variant 2
{
stack_t s; // new ((void*)s) stack_t;
s.push(123);
s.push(42);
assert(s.top() == 42);
s.pop();
assert(s.top() == 123);
// s->~stack_t(); (at the end of scope)
}
// variant 3
{
stack_t* s = new stack_t[2];
s[0].push(123);
s[0].push(42);
assert(s[0].top() == 42);
s[0].pop();
assert(s[0].top() == 123);
s[1].push(123);
s[1].push(42);
assert(s[1].top() == 42);
s[1].pop();
assert(s[1].top() == 123);
delete[] s;
}
return 0;
}
```
3) Ещё подужмем
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
struct stack_t {
int* a;
int sz;
int max_sz;
stack_t() {
a = nullptr;
sz = 0;
max_sz = 0;
}
~stack_t() {
free(a);
}
void push(int elem) {
if (sz == max_sz) {
max_sz += (max_sz == 0);
max_sz *= 2;
a = (int*)realloc(a, max_sz * sizeof(int));
}
a[sz++] = elem;
}
int top() {
return a[sz - 1];
}
void pop() {
--sz;
}
};
int main() {
stack_t s;
s.push(123);
s.push(42);
assert(s.top() == 42);
s.pop();
assert(s.top() == 123);
return 0;
}
```
4) Добавим шаблонов
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
#include <new>
#include <string>
template <typename TElem>
struct stack_t {
TElem* a;
int sz;
int max_sz;
stack_t() {
a = nullptr;
sz = 0;
max_sz = 0;
}
~stack_t() {
while (sz > 0) {
pop();
}
free(a);
}
void push(TElem elem) {
if (sz == max_sz) {
max_sz += (max_sz == 0);
max_sz *= 2;
a = (TElem*)realloc((void*)a, max_sz * sizeof(TElem));
}
new (a + sz) TElem(elem);
++sz;
}
TElem top() {
return a[sz - 1];
}
void pop() {
a[--sz].~TElem();
}
};
// template <typename TElem>
// struct queue_t {
// ....
// };
int main() {
{
stack_t<int> s;
s.push(123);
s.push(42);
assert(s.top() == 42);
s.pop();
assert(s.top() == 123);
}
{
stack_t<char> s;
s.push('A');
s.push('Z');
assert(s.top() == 'Z');
s.pop();
assert(s.top() == 'A');
}
{
stack_t<std::string> s;
s.push("Azaza");
s.push("Brekeke");
assert(s.top() == "Brekeke");
s.pop();
assert(s.top() == "Azaza");
}
return 0;
}
```
Немного больше о связи С++ и С
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
struct stack_t {
int* a;
int sz;
int max_sz;
stack_t();
~stack_t();
void push(int elem);
int top();
void pop();
};
// Если упрощенно, то код в этом блоке компилится как сишный, но при этом он связан с окружающим плюсовым
extern "C" {
void _ZN7stack_tC1Ev(stack_t* s) {
s->a = nullptr;
s->sz = 0;
s->max_sz = 0;
}
void _ZN7stack_tD1Ev(stack_t* s) {
free(s->a);
}
void _ZN7stack_t4pushEi(stack_t* s, int elem) {
if (s->sz == s->max_sz) {
s->max_sz += (s->max_sz == 0);
s->max_sz *= 2;
s->a = (int*)realloc(s->a, s->max_sz * sizeof(int));
}
s->a[s->sz++] = elem;
}
void _ZN7stack_t3popEv(stack_t* s) {
--s->sz;
}
int _ZN7stack_t3topEv(stack_t* s) {
return s->a[s->sz - 1];
}
}
int main() {
stack_t s;
s.push(123);
s.push(42);
assert(s.top() == 42);
s.pop();
assert(s.top() == 123);
return 0;
}
!objdump -t a.exe | grep stack_t
```
# Больше полезностей
https://ravesli.com/urok-192-std-move/
Тут вроде прилично написано про std::move
https://habr.com/ru/post/348198/
Вот тут про lvalue/rvalue и ссылочки всякие
## Перегрузка функций
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
%run objdump -t a.exe | grep sqr
#include <stdio.h>
#include <math.h>
int sqr(int a) {
return a * a;
}
double sqr(double a) {
return a * a;
}
int main() {
printf("%d\n", sqr('a'));
printf("%d\n", sqr(2));
printf("%lf\n", sqr(3.0));
return 0;
}
```
## Перегрузка арифметических операторов
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <math.h>
struct vec_t {
double x;
double y;
double norm() const {
return std::sqrt(x * x + y * y);
}
vec_t operator+(const vec_t& b) const {
return {this->x + b.x, this->y + b.y};
}
vec_t operator-() const {
return {-x, -y};
}
void print() const {
printf("{%lf, %lf}\n", x, y);
}
};
vec_t operator*(const vec_t& a, double k) {
return {a.x * k, a.y * k};
}
int main() {
vec_t{1, 2}.print();
vec_t a = {10, 20};
vec_t b = {100, 200};
(a + b).print();
(-a).print();
(a * -2).print();
return 0;
}
```
## Указатели, ссылки, объекты
```
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
#include <new>
#include <string>
template <typename TElem>
struct stack_t {
TElem* a;
int sz;
int max_sz;
stack_t() {
a = nullptr;
sz = 0;
max_sz = 0;
}
stack_t(const stack_t& other): stack_t() {
*this = other;
}
stack_t(stack_t&& other): stack_t() {
*this = std::move(other);
}
stack_t& operator=(const stack_t& other) {
clear();
for (int i = 0; i < other.sz; ++i) {
push(other.a[i]);
}
return *this;
}
stack_t& operator=(stack_t&& other) {
std::swap(a, other.a);
std::swap(sz, other.sz);
std::swap(max_sz, other.max_sz);
other.clear();
return *this;
}
void clear() {
while (sz > 0) {
pop();
}
}
~stack_t() {
clear();
free(a);
}
void push(TElem elem) {
if (sz == max_sz) {
max_sz += (max_sz == 0);
max_sz *= 2;
// На самом деле так нельзя в общем случае
// не все объекты хорошо перенесут изменение своего адреса в памяти
// a = (TElem*)realloc((void*)a, max_sz * sizeof(TElem));
TElem* new_a = (TElem*)malloc(max_sz * sizeof(TElem));
for (int i = 0; i < sz; ++i) {
new (new_a + i) TElem(std::move(a[i])); // move-конструктором безопасно перемещаем объект в новую память
a[i].~TElem();
}
a = new_a;
}
new (a + sz) TElem(elem);
++sz;
}
TElem top() {
return a[sz - 1];
}
void pop() {
a[--sz].~TElem();
}
};
stack_t<int> create() {
stack_t<int> s;
s.push(1);
return s;
}
int main() {
{
stack_t<int> s;
s.push(123);
s.push(42);
stack_t<int> s2 = s;
assert(s.top() == 42);
s.pop();
assert(s.top() == 123);
assert(s2.top() == 42);
s2.pop();
assert(s2.top() == 123);
}
{
stack_t<int> s;
s.push(123);
s.push(42);
stack_t<int> s2 = std::move(s);
assert(s.sz == 0);
assert(s2.top() == 42);
s2.pop();
assert(s2.top() == 123);
}
{
stack_t<int> s;
s.push(123);
s.push(42);
s = create();
assert(s.sz == 1);
assert(s.top() == 1);
}
{
stack_t<stack_t<int>> s;
s.push(create());
s.push(create());
s.pop();
}
return 0;
}
```
Стандартные контейнеры
vector/queue/priority_queue/set/map/unordered_*/string
Самая нормальная документация, что я знаю тут: https://en.cppreference.com/w/
```
```
# HW
Реализовать на С++ очередь (циклическую/на двух стеках/на указателях)
Опционально шаблонную
Опционально с хорошими конструкторами и операторами присваивания
Как сдавать - выложить на pastebin и прислать в лс
Дедлайн 7 февраля 23:00.
# Задачки про порядок вызова конструкторов и деструкторов
```
%%cpp common.h
#include <stdio.h>
#include <stdlib.h>
#include <string.h>
#include <assert.h>
#include <new>
#include <string>
#define eprintf(...) fprintf(stderr, __VA_ARGS__)
#define logprintf_impl(fmt, line, ...) eprintf(__FILE__ ":" #line " " fmt, __VA_ARGS__)
#define logprintf_impl_2(fmt, line, ...) logprintf_impl(fmt, line, __VA_ARGS__)
#define logprintf(fmt, ...) logprintf_impl_2(fmt, __LINE__, __VA_ARGS__)
// Выделяем на sizeof(size_t) байт больше, чтобы явно сохранить размер выделяемого блока
void* operator new(size_t sz) {
void* ptr = malloc(sz);
printf("allocate %d bytes, addr=%p\n", (int)sz, (void*)ptr);
return ptr;
}
// А здесь удаляем этот расширенный блок и выводим сохраненный размер
void operator delete(void* ptr, size_t sz) noexcept
{
printf("deallocate %d bytes, addr=%p\n", (int)sz, ptr);
free(ptr);
}
// размер структур совпадает с последней цифрой названия
struct obj4 {
char data[4];
obj4(obj4&&) { printf("construct (move) ojb4\n"); }
obj4& operator=(obj4&&) { printf("assign (move) ojb4\n"); return *this; }
obj4() { printf("construct ojb4\n"); }
~obj4() { printf("destruct ojb4\n"); }
};
struct obj5 {
char data[5];
obj5() { printf("construct ojb5\n"); }
~obj5() { printf("destruct ojb5\n"); }
};
struct obj10 {
obj5 o5;
obj4 o4;
char data[1];
obj10() { printf("construct ojb10\n"); }
~obj10() { printf("destruct ojb10\n"); }
};
struct obj20 {
obj4* o4;
obj5 o5;
char data[7];
obj20() { printf("construct ojb20\n"); o4 = new obj4; printf("end of construct ojb20\n"); }
~obj20() { printf("destruct ojb20\n"); delete o4; printf("end of destruct ojb20\n"); }
};
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj4 o4; // constructor
return 0;
// destructor
}
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj4* o4 = new obj4; // allocate, constructor obj4
delete o4; // destructor, deallocate
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj10 o10;
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj20 o20;
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj20* o20 = new obj20;
delete o20;
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 -Wall -Werror -fsanitize=address -fno-exceptions -fno-rtti main.cpp -o a.exe
%run ./a.exe
#include "common.h"
int main() {
obj4 o4_1;
obj4 o4_2;
std::swap(o4_1, o4_2);
return 0;
}
%%cpp lib.h
// void print42();
#include <stdio.h>
inline void print42() {
printf("42\n");
}
template <typename T>
T min(T a, T b);
%%cpp lib.cpp
#include "lib.h"
// #include <stdio.h>
// void print42() {
// printf("42\n");
// }
template <typename T>
T min(T a, T b) {
return (a > b) ? b : a;
}
// int f(int a, int b) {
// return min(a, b);
// }
template
int min<int> (int a, int b);
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include "lib.h"
int main() {
print42();
printf("%d\n", min(10, 30));
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run echo "10.1 20.2" | ./main.exe
#include <stdio.h>
#include <iostream>
#include <iomanip>
struct point_t {
double x;
double y;
point_t operator-(const point_t& b) const {
return point_t{.x = x - b.x, .y = y - b.y};
}
point_t operator*(double k) const {
return {.x = x * k, .y = y * k};
}
static point_t read(FILE* file) {
point_t p;
fscanf(file, "%lf%lf", &p.x, &p.y);
return p;
}
void write(FILE* file) const {
fprintf(file, "{.x = %lf, .y = %lf}", x, y);
}
};
std::istream& operator>>(std::istream& in, point_t& p) {
return in >> p.x >> p.y;
}
std::ostream& operator<<(std::ostream& out, const point_t& p) {
return out << "{" << std::fixed << std::setprecision(3) << p.x << ", " << p.y << "}";
}
int main() {
//(point_t::read(stdin) * 2).write(stdout);
point_t p;
std::cin >> p;
std::cout << (p * 2);
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <cassert>
template <typename T>
struct uniq_ptr {
T* ptr;
uniq_ptr() {
ptr = nullptr;
}
explicit uniq_ptr(T* p) {
ptr = p;
}
uniq_ptr(uniq_ptr<T>&& t) {
ptr = t.ptr;
t.ptr = nullptr;
}
uniq_ptr<T>& operator=(uniq_ptr<T>&& t) {
reset();
ptr = t.ptr;
t.ptr = nullptr;
return *this;
}
T* operator->() {
assert(ptr && "*null");
return ptr;
}
T& operator*() {
assert(ptr && "*null");
return *ptr;
}
void reset() {
if (ptr) {
delete ptr;
}
}
~uniq_ptr() {
reset();
}
};
struct obj_t {
obj_t() { printf("construct obj_t\n"); }
void touch() { printf("is being touched\n"); }
~obj_t() { printf("destruct obj_t\n"); }
};
uniq_ptr<obj_t> create_obj() {
uniq_ptr<obj_t> obj(new obj_t);
printf("0\n");
return obj;
}
int main() {
obj_t* t = NULL;
free(t);
uniq_ptr<obj_t> obj = create_obj();
obj.ptr->touch();
obj->touch();
(*obj).touch();
printf("2\n");
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
struct obj_t {
obj_t() { printf("construct obj_t\n"); }
void touch() { printf("is being touched\n"); }
~obj_t() { printf("destruct obj_t\n"); }
};
int main() {
obj_t* t = NULL;
free(t);
std::shared_ptr<obj_t> obj(new obj_t);
std::shared_ptr<obj_t> obj2 = obj;
obj->touch();
(*obj).touch();
printf("0\n");
obj.reset();
printf("1\n");
obj2.reset();
printf("2\n");
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <vector>
#include <algorithm>
int main() {
std::vector<int> v;
v.push_back(1);
v[0];
v.resize(2);
v.push_back(2);
v.push_back(5);
v.pop_back();
std::sort(v.begin(), v.end(), std::greater<>());
std::sort(v.begin(), v.end(), [](int a, int b) {
return a > b;
});
v.reserve(100);
for (int* i = v.data(); i != v.data() + v.size(); ++i) {
printf("%d ", *i);
}
printf("\n");
for (std::vector<int>::iterator i = v.begin(); i != v.end(); ++i) {
printf("%d ", *i);
}
printf("\n");
for (auto i = v.begin(); i != v.end(); ++i) {
printf("%d ", *i);
}
printf("\n");
int a[232];
for (auto i = std::begin(v); i != std::end(v); ++i) {
printf("%d ", *i);
}
printf("\n");
{
auto&& __x = v;
for (auto __i = std::begin(__x); __i != std::end(__x); ++__i) {
auto& x = *__i;
printf("%d ", x);
}
}
printf("\n");
for (int x : v) {
printf("%d ", x);
}
printf("\n");
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <vector>
namespace my {
void f();
void g();
}
namespace my {
void f() {
printf("X\n");
}
void g() {
printf("Y\n");
f();
}
}
int main() {
using my::g;
g();
return 0;
}
%%cpp main.cpp
%run g++ main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <set>
int main() {
std::set<int> v;
v.insert(4);
v.insert(4);
v.insert(5);
if (v.count(4)) {
printf("4 in set\n");
}
{
auto it = v.find(4);
if (it != v.end()) {
printf("4 in set\n");
}
}
for (int x : v) {
printf("%d ", x);
}
printf("\n");
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <map>
int main() {
std::map<int, int> v;
v[0] = 0;
v[1] = 10;
v[2] = 20;
v[10] = 100;
printf("v[10] = %d\n", v[10]);
if (v.count(4)) {
printf("4 in set\n");
}
{
auto [iter, success] = v.try_emplace(6, 60);
if (success) {
printf("6 added to set\n");
} else {
printf("6 NOT added to set\n");
}
}
for (auto& [key, value] : v) {
printf("(%d %d) ", key, value);
}
printf("\n");
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <unordered_map>
int main() {
std::unordered_map<int, int> v;
v[0] = 0;
v[1] = 10;
v[2] = 20;
v[10] = 100;
printf("v[10] = %d\n", v[10]);
if (v.count(4)) {
printf("4 in set\n");
}
{
auto [iter, success] = v.try_emplace(6, 60);
if (success) {
printf("6 added to set\n");
} else {
printf("6 NOT added to set\n");
}
}
for (auto& [key, value] : v) {
printf("(%d %d) ", key, value);
}
printf("\n");
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <vector>
#include <unordered_map>
int main() {
std::vector<std::string> names = {
"x",
"y",
"x",
"y",
"z",
};
std::unordered_map<std::string, int> name_to_index(names.size());
std::vector<int> indexes;
indexes.reserve(names.size());
for (auto& name : names) {
auto [iter, success] = name_to_index.try_emplace(
name, name_to_index.size());
indexes.push_back(iter->second);
}
for (auto i : indexes) {
printf("%d ", i);
}
printf("\n");
return 0;
}
%%cpp main.cpp
%run g++ -std=c++17 main.cpp lib.cpp -o main.exe
%run ./main.exe
#include <stdio.h>
#include <iostream>
#include <memory>
#include <vector>
#include <unordered_map>
template <typename T>
struct array2d_t {
T* arr;
int n;
int m;
array2d_t(int n_, int m_) {
n = n_;
m = m_;
arr = new T[n * m];
for (int i = 0; i < n * m; ++i) {
arr[i] = {};
}
}
T* operator[](int i) {
return arr + i * m;
}
~array2d_t() {
delete[] arr;
}
};
int main() {
array2d_t<int> arr(5, 5);
arr[3][3] = 142;
for (int i = 0; i < 5; ++i) {
for (int j = 0; j < 5; ++j) {
printf("%d ", arr[i][j]);
}
printf("\n");
}
return 0;
}
%%cpp main.cpp
%run clang++ -std=c++17 -Wall -Werror -fsanitize=address main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string>
#include <variant>
template <typename T>
struct TErrorOr {
std::variant<std::string, T> Variant;
TErrorOr() {
Variant.template emplace<0>("not defined");
}
TErrorOr(T value) {
Variant.template emplace<1>(std::move(value));
}
bool IsOk() const {
return Variant.index() == 1;
}
T& Value() {
return std::get<1>(Variant);
}
std::string& Error() {
return std::get<0>(Variant);
}
};
template <typename T>
TErrorOr<T> CreateError(std::string str) {
TErrorOr<T> err;
err.Variant.template emplace<0>(std::move(str));
return err;
}
TErrorOr<int> f(int a) {
if (a > 40000)
return CreateError<int>("a too big");
return a * a;
}
TErrorOr<int> f2(int a, int b) {
TErrorOr<int> a2 = f(a);
if (!a2.IsOk()) {
return a2;
}
TErrorOr<int> b2 = f(b);
if (!b2.IsOk()) {
return b2;
}
if ((int64_t)a + b > 2000000000) {
return CreateError<int>("a + b too big");
}
return a2.Value() + b2.Value();
}
int main() {
{
int a = 0;
TErrorOr<int> x = f(a);
if (!x.IsOk()) {
printf("Error %s\n", x.Error().c_str());
} else {
printf("Success, res = %d\n", x.Value());
}
}
{
int a = 1000;
int b = 100000;
TErrorOr<int> x = f2(a, b);
if (!x.IsOk()) {
printf("Error %s\n", x.Error().c_str());
} else {
printf("Success, res = %d\n", x.Value());
}
}
return 0;
}
%%cpp main.cpp
%run clang++ -std=c++17 -Wall -Werror -fsanitize=address main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string>
#include <exception>
#include <stdexcept>
int f(int a) {
if (a > 40000)
throw std::runtime_error("a too big");
return a * a;
}
int f2(int a, int b) {
int64_t res = f(a) + f(b);
if (res > 2000000000) {
throw std::runtime_error("a + b too big");
}
return res;
}
int main() {
try {
int a = 1000000;
int x = f(a);
printf("Success, res = %d\n", x);
} catch (const std::exception& e) {
printf("Error: %s\n", e.what());
}
return 0;
}
%%cpp main.cpp
%run clang++ -std=c++17 -Wall -Werror -fsanitize=address main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string>
#include <exception>
#include <stdexcept>
void f() {
std::vector<int> x;
throw std::runtime_error("XXX");
std::vector<int> y;
}
int main() {
std::vector<int> a;
try {
std::vector<int> b;
f();
std::vector<int> c;
} catch (const std::exception& e) {
std::vector<int> d;
}
return 0;
}
swap для кастомных типов
%%cpp main.cpp
%run clang++ -std=c++20 -Wall -Werror -fsanitize=address main.cpp -o a.exe
%run ./a.exe
#include <stdio.h>
#include <stdlib.h>
#include <string>
#include <exception>
#include <stdexcept>
struct A {
// bool operator==(const A&) const = default;
};
int main() {
A{} == A{};
return 0;
}
```
| github_jupyter |
```
import random
import pickle
import matplotlib.pyplot as plt
import numpy as np
from functools import (partial, reduce)
import networkx as nx
import scipy
from sklearn import manifold
from gerrychain.constraints import (
Validator,
single_flip_contiguous,
within_percent_of_ideal_population,
)
from gerrychain import (GeographicPartition, Partition, Graph, MarkovChain,
proposals, updaters, constraints, accept, Election)
from gerrychain.proposals import propose_random_flip
from gerrychain.accept import always_accept
from gerrychain.updaters import Election, Tally, cut_edges
from gerrychain.partition import Partition
from gerrychain.proposals import recom
from gerrychain.metrics import mean_median, efficiency_gap
from sklearn.preprocessing import scale, normalize
grid_width = 4
grid_height = 4
node_size = 50
OPPORTUNITY_PERCENT = 0.5 # suffices for 3x3->3 grid
num_combinations = 117
graph = nx.grid_graph([grid_width , grid_height])
# #####PLOT GRIDS
initial_assignment = {x: x[0] for x in graph.nodes()}
graph = nx.relabel.convert_node_labels_to_integers(graph)
# build initial_assignment
relabeled_assignment = list(initial_assignment.values())
initial_assignment = dict()
for i in range(len(graph.nodes)):
initial_assignment[i] = relabeled_assignment[i]
# red is the minority
# red_nodes = [(1,1),(1,2),(2,1),(2,2)]
red_nodes = red_nodes = [1, 2, 5, 9, 10]
for node in graph.nodes():
graph.node[node]["population"] = 1
if node in red_nodes:
graph.node[node]["red"]=1
graph.node[node]["black"]=0
else:
graph.node[node]["red"]=0
graph.node[node]["black"]=1
# color_dict = {1: "red", 0: "black"}
# plt.figure()
# nx.draw(
# graph,
# pos={x: x for x in graph.nodes()},
# node_color=[color_dict[graph.node[x]["red"]] for x in graph.nodes()],
# node_size=node_size,
# node_shape="s",
# )
# plt.show()
### find a way to compare partitions
### run a chain and get the 10 districts
### pass it to JN's thing
def count_gingles_districts(partition):
num_gingles_districts = 0
for district in partition.parts.keys():
# count minority nodes
minority_nodes = 0
for node in partition.parts[district]:
if graph.nodes[node]["red"] == 1:
minority_nodes += 1
if minority_nodes / len(partition.parts.keys()) > OPPORTUNITY_PERCENT:
num_gingles_districts += 1
return num_gingles_districts
updaters = {
"population": Tally("population"),
"num_gingles_districts": count_gingles_districts,
"cut_edges": cut_edges
}
grid_partition = Partition(graph, assignment=initial_assignment, updaters=updaters)
# ADD CONSTRAINTS
popbound = within_percent_of_ideal_population(grid_partition, 0.1)
# ########Setup Proposal
ideal_population = sum(grid_partition["population"].values()) / len(grid_partition)
tree_proposal = partial(
recom,
pop_col="population",
pop_target=ideal_population,
epsilon=0.05,
node_repeats=1,
)
boundary_chain = MarkovChain(
tree_proposal,
Validator([single_flip_contiguous, popbound]),
accept=always_accept,
initial_state=grid_partition,
total_steps=10000,
)
# store the partitions in a list
partition_list = []
gingles_scores = []
# run the chain
for current_partition in boundary_chain:
seen_before = False
# check if you have seen the partition already
for seen_partition in partition_list:
# TODO: nasty code that needs to be rewritten
seen_list = seen_partition.parts.values()
seen_list = [sorted(list(unit)) for unit in seen_list]
seen_list = sorted(seen_list)
curr_list = current_partition.parts.values()
curr_list = [sorted(list(unit)) for unit in curr_list]
curr_list = sorted(curr_list)
if seen_list == curr_list:
seen_before = True
break
if not seen_before:
gingles_scores.append(current_partition["num_gingles_districts"])
# if haven't seen the partition before, we want to keep it
partition_list.append(current_partition)
if len(partition_list) == num_combinations:
break
print(len(partition_list))
# Don't run this yet
##################################################################################################
####################### SANITY CHECK TO VISUALIZE DISTRICT UNIQUENESS ############################
##################################################################################################
color_dict = {0: "blue", 1: "green", 2: "red"}
for partition in partition_list:
graph = partition.graph
# figure out the node colors
assignment = {}
for x in graph.nodes():
for key in partition.parts.keys():
if x in partition.parts[key]:
assignment[x] = key
plt.figure()
nx.draw(
graph,
pos={x: x for x in graph.nodes()},
node_color=[color_dict[x] for x in list(assignment.values())],
node_size=node_size,
node_shape="s",
)
plt.show()
def sparse_bound_walk_metric(plan1, plan2):
score = dir_sparse_bound_walk_metric(plan1, plan2) + dir_sparse_bound_walk_metric(plan2, plan1)
return score
def dir_sparse_bound_walk_metric(plan1, plan2):
graph = plan1.graph
adj = nx.adjacency_matrix(graph, weight=None)
trans = normalize(adj, norm="l1")
del adj
plan1_bound = reduce(lambda ns, e: set(e) | ns, plan1["cut_edges"], set())
plan2_bound = reduce(lambda ns, e: set(e) | ns, plan2["cut_edges"], set())
if all(list(map(lambda x: x in plan2_bound, plan1_bound))): return 0
to_delete = plan2_bound #| contained_by_plan2
starters = sorted(plan1_bound - to_delete)
to_delete = list(reversed(sorted(list(to_delete))))
plan1_bound = reduce(lambda shifted, x: list(map(lambda y: y-1 if y > x else y, shifted)),
to_delete, starters)
P = trans
del trans
row_mask = np.ones(P.shape[0], dtype=bool)
col_mask = np.ones(P.shape[1], dtype=bool)
row_mask[to_delete] = False
col_mask[to_delete] = False
P = P[row_mask][:,col_mask]
N = scipy.sparse.csr_matrix(scipy.sparse.identity(P.shape[0])) - P
ones = np.ones((P.shape[0], 1))
rowsum = scipy.sparse.linalg.spsolve(N, ones)
score = sum([rowsum[i] for i in plan1_bound])
return score / len(plan1_bound)
##################################################################################################
################################ 2D VISUALIZE ####################################################
##################################################################################################
# Change all the partitions to have graphs that have nodes labeled as integers
def change_node_labels_to_integers(partition_list):
for partition in partition_list:
partition.graph = nx.relabel.convert_node_labels_to_integers(partition.graph)
return partition_list
partition_list = change_node_labels_to_integers(partition_list)
num_plans = len(partition_list)
a = np.zeros([num_plans, num_plans])
for i in range(num_plans):
for j in range(num_plans):
if i > j:
temp = sparse_bound_walk_metric(partition_list[i], partition_list[j])
a[i,j] = temp
a[j,i] = temp
mds = manifold.MDS(n_components=2, max_iter=3000, eps=.00001,
dissimilarity="precomputed", n_jobs=1)
pos = mds.fit(a).embedding_
colors = ["red", "pink", "orange", "yellow", "green", "lime", "cyan",
"blue", "indigo", "mediumpurple", "blueviolet", "purple", "darkmagenta", "fuchsia"]
import time
plt.figure(figsize=(12,8))
for i in range(len(pos)):
plt.plot(pos[i][0],pos[i][1],'.',label=i, markersize=20, color=colors[i % len(colors)])
plt.xlim()
plt.legend()
plt.show()
xs = [tup[0] for tup in pos]
ys = [tup[1] for tup in pos]
# just make the z axis for now
from mpl_toolkits import mplot3d
fig = plt.figure()
ax = plt.axes(projection='3d')
x
# Data for three-dimensional scattered points
ax.scatter3D(xs, ys, gingles_scores, c=range(num_combinations));
# ax.contour3D(xs, ys, gingles_scores, 50, cmap='hsv')
plt.xlim(-3,3)
plt.ylim(-3,3)
ax.set_zlim(0,3)
ax.set_xlabel('1st Principal Component')
ax.set_ylabel('2nd Principal Component')
ax.set_zlabel('# of Gingles districts')
ax.set_title("Diffusion distances, 4x4->4 with 5 minority nodes")
ax.view_init(20, 45)
plt.savefig("pointplot_4x4->4 w 5 minority nodes")
# ok so each point is being squished with another
# what is the similarity between these two partitions that makes them squished??
pos
```
| github_jupyter |
# T1546.011 - Event Triggered Execution: Application Shimming
Adversaries may establish persistence and/or elevate privileges by executing malicious content triggered by application shims. The Microsoft Windows Application Compatibility Infrastructure/Framework (Application Shim) was created to allow for backward compatibility of software as the operating system codebase changes over time. For example, the application shimming feature allows developers to apply fixes to applications (without rewriting code) that were created for Windows XP so that it will work with Windows 10. (Citation: Endgame Process Injection July 2017)
Within the framework, shims are created to act as a buffer between the program (or more specifically, the Import Address Table) and the Windows OS. When a program is executed, the shim cache is referenced to determine if the program requires the use of the shim database (.sdb). If so, the shim database uses hooking to redirect the code as necessary in order to communicate with the OS.
A list of all shims currently installed by the default Windows installer (sdbinst.exe) is kept in:
* <code>%WINDIR%\AppPatch\sysmain.sdb</code> and
* <code>hklm\software\microsoft\windows nt\currentversion\appcompatflags\installedsdb</code>
Custom databases are stored in:
* <code>%WINDIR%\AppPatch\custom & %WINDIR%\AppPatch\AppPatch64\Custom</code> and
* <code>hklm\software\microsoft\windows nt\currentversion\appcompatflags\custom</code>
To keep shims secure, Windows designed them to run in user mode so they cannot modify the kernel and you must have administrator privileges to install a shim. However, certain shims can be used to [Bypass User Access Control](https://attack.mitre.org/techniques/T1548/002) (UAC and RedirectEXE), inject DLLs into processes (InjectDLL), disable Data Execution Prevention (DisableNX) and Structure Exception Handling (DisableSEH), and intercept memory addresses (GetProcAddress).
Utilizing these shims may allow an adversary to perform several malicious acts such as elevate privileges, install backdoors, disable defenses like Windows Defender, etc. (Citation: FireEye Application Shimming) Shims can also be abused to establish persistence by continuously being invoked by affected programs.
## Atomic Tests
```
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
```
### Atomic Test #1 - Application Shim Installation
Install a shim database. This technique is used for privilege escalation and bypassing user access control.
Upon execution, "Installation of AtomicShim complete." will be displayed. To verify the shim behavior, run
the AtomicTest.exe from the <PathToAtomicsFolder>\\T1546.011\\bin directory. You should see a message box appear
with "Atomic Shim DLL Test!" as defined in the AtomicTest.dll. To better understand what is happening, review
the source code files is the <PathToAtomicsFolder>\\T1546.011\\src directory.
**Supported Platforms:** windows
Elevation Required (e.g. root or admin)
#### Dependencies: Run with `powershell`!
##### Description: Shim database file must exist on disk at specified location (#{file_path})
##### Check Prereq Commands:
```powershell
if (Test-Path PathToAtomicsFolder\T1546.011\bin\AtomicShimx86.sdb) {exit 0} else {exit 1}
```
##### Get Prereq Commands:
```powershell
[Net.ServicePointManager]::SecurityProtocol = [Net.SecurityProtocolType]::Tls12
New-Item -Type Directory (split-path PathToAtomicsFolder\T1546.011\bin\AtomicShimx86.sdb) -ErrorAction ignore | Out-Null
Invoke-WebRequest "https://github.com/redcanaryco/atomic-red-team/raw/master/atomics/T1546.011/bin/AtomicShimx86.sdb" -OutFile "PathToAtomicsFolder\T1546.011\bin\AtomicShimx86.sdb"
```
##### Description: AtomicTest.dll must exist at c:\Tools\AtomicTest.dll
##### Check Prereq Commands:
```powershell
if (Test-Path c:\Tools\AtomicTest.dll) {exit 0} else {exit 1}
```
##### Get Prereq Commands:
```powershell
New-Item -Type Directory (split-path c:\Tools\AtomicTest.dll) -ErrorAction ignore | Out-Null
Invoke-WebRequest "https://github.com/redcanaryco/atomic-red-team/raw/master/atomics/T1546.011/bin/AtomicTest.dll" -OutFile c:\Tools\AtomicTest.dll
```
```
Invoke-AtomicTest T1546.011 -TestNumbers 1 -GetPreReqs
```
#### Attack Commands: Run with `command_prompt`
```command_prompt
sdbinst.exe PathToAtomicsFolder\T1546.011\bin\AtomicShimx86.sdb
```
```
Invoke-AtomicTest T1546.011 -TestNumbers 1
```
### Atomic Test #2 - New shim database files created in the default shim database directory
Upon execution, check the "C:\Windows\apppatch\Custom\" folder for the new shim database
https://www.fireeye.com/blog/threat-research/2017/05/fin7-shim-databases-persistence.html
**Supported Platforms:** windows
Elevation Required (e.g. root or admin)
#### Attack Commands: Run with `powershell`
```powershell
Copy-Item $PathToAtomicsFolder\T1546.011\bin\T1546.011CompatDatabase.sdb C:\Windows\apppatch\Custom\T1546.011CompatDatabase.sdb
Copy-Item $PathToAtomicsFolder\T1546.011\bin\T1546.011CompatDatabase.sdb C:\Windows\apppatch\Custom\Custom64\T1546.011CompatDatabase.sdb
```
```
Invoke-AtomicTest T1546.011 -TestNumbers 2
```
### Atomic Test #3 - Registry key creation and/or modification events for SDB
Create registry keys in locations where fin7 typically places SDB patches. Upon execution, output will be displayed describing
the registry keys that were created. These keys can also be viewed using the Registry Editor.
https://www.fireeye.com/blog/threat-research/2017/05/fin7-shim-databases-persistence.html
**Supported Platforms:** windows
Elevation Required (e.g. root or admin)
#### Attack Commands: Run with `powershell`
```powershell
New-ItemProperty -Path HKLM:"\SOFTWARE\Microsoft\Windows NT\CurrentVersion\AppCompatFlags\Custom" -Name "AtomicRedTeamT1546.011" -Value "AtomicRedTeamT1546.011"
New-ItemProperty -Path HKLM:"\SOFTWARE\Microsoft\Windows NT\CurrentVersion\AppCompatFlags\InstalledSDB" -Name "AtomicRedTeamT1546.011" -Value "AtomicRedTeamT1546.011"
```
```
Invoke-AtomicTest T1546.011 -TestNumbers 3
```
## Detection
There are several public tools available that will detect shims that are currently available (Citation: Black Hat 2015 App Shim):
* Shim-Process-Scanner - checks memory of every running process for any shim flags
* Shim-Detector-Lite - detects installation of custom shim databases
* Shim-Guard - monitors registry for any shim installations
* ShimScanner - forensic tool to find active shims in memory
* ShimCacheMem - Volatility plug-in that pulls shim cache from memory (note: shims are only cached after reboot)
Monitor process execution for sdbinst.exe and command-line arguments for potential indications of application shim abuse.
| github_jupyter |
```
%matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from scipy import optimize
import scipy.stats as stat
import pylab
from collections import Counter
```
## Importing the Dataset
```
# Predict whether a customer will buy the new SUV.
# This information will later be given to the advertising department to optimize targetting future customers.
df = pd.read_csv("/content/Social_Network_Ads.csv") # Binary Classification
df.head()
# Length of dataset
len(df)
# Datatype of each column
df.dtypes
```
## Checking for missing values
```
df.isnull().sum()
# No missing values
```
## Check for outliers - Logistic Regression is sensitive to outliers
```
figure = df.boxplot(column="Age")
figure = df.boxplot(column="EstimatedSalary")
```
## Create matrix of features and label
```
X = df.iloc[:,:-1].values # features
y = df.iloc[:,-1].values # label
# X and y are numpy arrays
X.shape, y.shape
```
## Splitting the dataset into Train and Test Set
```
from sklearn.model_selection import train_test_split
# Train - 75% data, Test - 25% data
X_train,X_test,y_train,y_test = train_test_split(X, y, test_size=0.25, random_state=0) # test_size = 0.2 is totally fine
# random_state --> for same split of data
X_train.shape,y_train.shape,X_test.shape,y_test.shape
```
## Feature Scaling (not needed)
Feature scaling is not needed for all ML algorithms. Only for some.
**It is not needed for logistic regression**
```
from sklearn.preprocessing import StandardScaler
sc = StandardScaler()
X_train = sc.fit_transform(X_train)
X_test = sc.transform(X_test)
```
## Training Logistic Regression model on Training set
```
from sklearn.linear_model import LogisticRegression # class
classifier = LogisticRegression() # object, i.e. create the model
# Train the model on training set
classifier.fit(X_train,y_train) # Fit parameters on hypothesis function
# check what the parameters mean by checking out Sklearn documentation
```
## Predicitng a new result
Age = 30, Expected salary = $87,000
Yes or No? To buy SUV
```
# predict() --> direct prediction
# predict_proba() --> probability
pred = classifier.predict([[30, 87000]]) # input must be a 2D matrix
pred
# If feature was scaled (though not needed here), classifier.predict(sc.transform([[30, 87000]]))
pred_prob = classifier.predict_proba([[30, 87000]])
pred_prob
# probabilty of 0 > 1, so final answer is 0
```
## Predicting the Test results - Evaluating the model
```
# Test set --> evaluate model on new observations (future data)
# Input the features to predict the label(estimated), returns a vector(matrix)
y_pred = classifier.predict(X_test)
# Concatenate the 2 vectors (Compare two numerical vectors - estimated and true)
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# reshape vector from horizantal to vertical
# axis = 1 for vertical concatenation
# first column --> predicted label (test set), second column --> true label (real ones - train set)
```
### Four of the main evaluation metrics/methods you'll come across for classification models are:
1. Accuracy
2. Area under ROC curve
3. Confusion matrix
4. Classification report
## Confusion Matrix
The next way to evaluate a classification model is by using a [confusion matrix.](https://en.wikipedia.org/wiki/Confusion_matrix)
A confusion matrix is a quick way to compare estimated labels with actual labels. In essence, giving you an idea of where the model is getting confused.
```
from sklearn.metrics import confusion_matrix
y_pred = classifier.predict(X_test)
confuse_matrix = confusion_matrix(y_test,y_pred) # compare actual labels (test) with predicted labels
confuse_matrix
```
A BETTER visual way is with Seaborn's [heatmap()](https://seaborn.pydata.org/generated/seaborn.heatmap.html) plot.
```
# Function to plot confusion matrix using Seaborn's heatmap()
def plot_confusion_matrix(confuse_matrix):
fig,ax = plt.subplots(figsize=(8,6))
# Set the font scale
sns.set(font_scale=1.5)
ax = sns.heatmap(
confuse_matrix,
annot=True, # Annote the boxes
cbar=False
)
plt.ylabel("Predicted label")
plt.xlabel("True label")
plot_confusion_matrix(confuse_matrix)
# when predict = 0 and actual = 0, we have 65 samples (correct sammples)
# when predict = 1 and actual = 1, we have 24 samples (correct sammples)
# false +ve --> when predict = 1 and actual = 0
# false -ve --> when predict = 0 and actual = 1
# so, model is getting confused in false +ve and false -ve
```
For clarity: (predict target vs true target)
* True positive = model predicts 1 when truth is 1
* False positive = model predicts 1 when truth is 0
* True negative = model predicts 0 when truth is 0
* False negative = model predicts 0 when truth is 1
```
from sklearn.metrics import accuracy_score
accuracy_score(y_test,y_pred)
```
## Classification Report
A classification report is a collection of metrics.
create a classification report using Scikit-Learn's [classification_report()](https://scikit-learn.org/stable/modules/generated/sklearn.metrics.classification_report.html) function.
```
from sklearn.metrics import classification_report
print(classification_report(y_test,y_pred))
```
It returns four columns: precision, recall, f1-score and support.
Each term measures something slightly different:
* **Precision** - Indicates the proportion of positive identifications (model predicted class 1) which were actually correct. A model which produces no false positives has a precision of 1.0.
* **Recall** - Indicates the proportion of actual positives which were correctly classified. A model which produces no false negatives has a recall of 1.0.
* **F1 score** - A combination of precision and recall. A perfect model achieves an F1 score of 1.0.
* **Support** - The number of samples each metric was calculated on.
* **Accuracy** - The accuracy of the model in decimal form. Perfect accuracy is equal to 1.0, in other words, getting the prediction right 100% of the time.
* **Macro avg** - Short for macro average, the average precision, recall and F1 score between classes. Macro avg doesn't take class imbalance into effect. So if you do have class imbalances (more examples of one class than another), you should pay attention to this.
* **Weighted avg** - Short for weighted average, the weighted average precision, recall and F1 score between classes. Weighted means each metric is calculated with respect to how many samples there are in each class. This metric will favour the majority class (e.g. it will give a high value when one class out performs another due to having more samples).
**When should you use each?**
It can be tempting to base your classification models perfomance only on accuracy. And accuracy is a good metric to report, except when you have very imbalanced classes.
To summarize:
* Accuracy is a good measure to start with if all classes are balanced (e.g. same amount of samples which are labelled with 0 or 1)
* Precision and recall become more important when classes are imbalanced.
* If false positive predictions are worse than false negatives, aim for higher precision.
* If false negative predictions are worse than false positives, aim for higher recall.
## Visualize the Training set results
```
# x axis --> 1st feature (Age), y axis --> 2nd feature (Expected salary)
# each points corresponds to a single sample
# we will plot a decision boundary - a linear classifier
# code is pretty much advanced -- only useful for training purposes (no need to understand that much)
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(X_train), y_train # Train set
# create a grid of ranges, step = 0.25 --> dense grid
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
# green - 1, red - 0
# apply predict method on each dense point in the grid
plt.contourf(X1, X2, classifier.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Logistic Regression (Training set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
```
## Visualize the Test Set results
```
from matplotlib.colors import ListedColormap
X_set, y_set = sc.inverse_transform(X_test), y_test # Test set (new, unseen samples)
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 10, stop = X_set[:, 0].max() + 10, step = 0.25),
np.arange(start = X_set[:, 1].min() - 1000, stop = X_set[:, 1].max() + 1000, step = 0.25))
plt.contourf(X1, X2, classifier.predict(sc.transform(np.array([X1.ravel(), X2.ravel()]).T)).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1], c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Logistic Regression (Test set)')
plt.xlabel('Age')
plt.ylabel('Estimated Salary')
plt.legend()
plt.show()
# There are still some incorrect predictions using decision boundary (linear classifier)
# So we will use a stronger classification algorithm (non-linear classifiers), i.e. a prediction curve
```
| github_jupyter |
```
!pip install Category_Encoders
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import SelectKBest
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import classification_report
from category_encoders import OneHotEncoder, OrdinalEncoder
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestRegressor
import category_encoders as ce
import numpy as np
from sklearn.feature_selection import f_regression, SelectKBest
from sklearn.impute import SimpleImputer
from sklearn.linear_model import Ridge, RidgeCV
from sklearn.model_selection import cross_val_score
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.metrics import mean_absolute_error
from xgboost import XGBClassifier
from sklearn.ensemble import GradientBoostingClassifier
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
from sklearn.pipeline import make_pipeline
from google.colab import files
import pandas as pd
uploaded = files.upload()
import pandas as pd
df = pd.read_csv('/content/Path where you want to store the exported CSV file_File Name.csv')
import pandas as pd
# Test on the last 10,000 loans,
# Validate on the 10,000 before that,
# Train on the rest
test = df[-100:]
val = df[-200:-100]
train = df[:-200]
target = 'work_interfere'
X_train = train.drop(['work_interfere'], axis=1)
y_train = train[target]
X_val = val.drop(['work_interfere'], axis=1)
y_val = val[target]
X_test = test.drop(['work_interfere'], axis=1)
y_test = test[target]
model = Pipeline([
('ohe', OneHotEncoder()),
('impute', SimpleImputer()),
('classifier', RandomForestClassifier(n_jobs=-1))
])
model.fit(X_train, y_train)
%%time
model.fit(X_train, y_train)
print('training accuracy:', model.score(X_train, y_train))
print('validation accuracy:', model.score(X_val, y_val))
print('test accuracy:', model.score(X_test, y_test))
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.linear_model import LogisticRegression
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
encoder = ce.OneHotEncoder(use_cat_names=True)
imputer = SimpleImputer()
scaler = StandardScaler()
model = LogisticRegression(max_iter=1000)
X_train_encoded = encoder.fit_transform(X_train)
X_train_imputed = imputer.fit_transform(X_train_encoded)
X_train_scaled = scaler.fit_transform(X_train_imputed)
model.fit(X_train_scaled, y_train)
X_val_encoded = encoder.transform(X_val)
X_val_imputed = imputer.transform(X_val_encoded)
X_val_scaled = scaler.transform(X_val_imputed)
print('Validation Accuracy', model.score(X_val_scaled, y_val))
X_test_encoded = encoder.transform(X_test)
X_test_imputed = imputer.transform(X_test_encoded)
X_test_scaled = scaler.transform(X_test_imputed)
y_pred = model.predict(X_test_scaled)
from sklearn.ensemble import AdaBoostClassifier
ada_classifier = AdaBoostClassifier(n_estimators=50, learning_rate=1.5, random_state=42)
ada_classifier.fit(X_train_encoded,y_train)
print('Validation Accuracy: Adaboost', ada_classifier.score(X_test_encoded, y_test))
# Load xgboost and fit the model
from xgboost import XGBClassifier
xg_classifier = XGBClassifier(n_estimators=50, random_state=42)
xg_classifier.fit(X_train_encoded,y_train)
print('Validation Accuracy: Adaboost', xg_classifier.score(X_test_encoded, y_test))
# Create a new column with 0=long and 1=short class labels
df['treatment'] = le.fit_transform(df['treatment'])
display(df.head())
target = 'work_interfere'
y_train = train[target]
y_val = val[target]
y_test = test[target]
```
| github_jupyter |
Subsets and Splits
No community queries yet
The top public SQL queries from the community will appear here once available.