code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import haversine as hv
import numpy as np
import ipywidgets
# Read in
location = pd.read_csv("Data/location.txt", delimiter="\t", names=["longitude","latitude","a","b"])
del location["a"], location["b"]
# From examples_week10.ipynb (IS590DV Class example)
# Tohoku : 142.369, 38.322
Tohoku=(142.369, 38.322)
# Calculate the distances
distance=[]
for row in location.iterrows():
station = (row[1][0],row[1][1])
distance.append(hv.haversine(Tohoku,station))
# Add distance column to location DataFrame
location["distance"]=distance
location["order"]=location.index
# Sort by distacnes (acsending)
location = location.sort_values(by=['distance'])
order=list(location["order"])
# Read in detector data
detector=pd.read_csv("Data/data_tohoku_norm_transpose.csv",names=np.arange(0,438), index_col=False)
ordered_detector = detector[order[0]]
for each in order[1:]:
ordered_detector = pd.concat([ordered_detector,detector[each]],axis=1)
ordered_detector
# +
# Add time range column, from week 10 example
time = pd.date_range("2:46PM", "6:46PM", freq="1s")
time -= time[0]
print(len(time))
print(len(ordered_detector))
ordered_detector["time"] = time
ordered_detector.set_index("time", inplace=True)
# -
@ipywidgets.interact(Time=(0,14400,1), Num=list(np.arange(0,438)))
def make_plot(Time, Num=0):
fig = plt.figure(figsize=(15,10))
# Plot the detectore on US map
ax11 = plt.subplot2grid((12,8), (1,0), colspan=4, rowspan=6)
ax11.scatter(location["longitude"],location["latitude"], c=detector.iloc[Time], cmap="Reds")
# Reference; https://stackoverflow.com/questions/8202605/matplotlib-scatterplot-colour-as-a-function-of-a-third-variable
ax11.set_title("Visualizing Tohoku Earthquake Detectors")
ax11.axes.get_xaxis().set_visible(False)
ax11.axes.get_yaxis().set_visible(False)
# Reference: https://stackoverflow.com/questions/2176424/hiding-axis-text-in-matplotlib-plots
ax21 = plt.subplot2grid((12,8), (0,5), colspan=3, rowspan=3)
ax22 = plt.subplot2grid((12,8), (4,5), colspan=3, rowspan=3)
ax22.plot(detector[Num])
ax22.set_title(label = "Waveform of Detector No."+str(Num))
plt.show()
# # Test area
# ?plt.subplot2grid
|
part1/Component+1_1127.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Regularized Linear Regression
import numpy as np
import matplotlib.pyplot as plt
import scipy.io as scio
train_data = scio.loadmat("./ex5data1.mat")
# print(train_data)
X = train_data['X']
Y = train_data['y']
Xtest = train_data['Xtest']
Ytest = train_data['ytest']
Xval = train_data['Xval']
Yval = train_data['yval']
# +
# Plot the training set
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax1.set_title('Data distribution')
plt.xlabel('Change in water level (x)')
plt.ylabel('Water following out of the dam(y)')
plt.scatter(X, Y, color='red', marker='x')
plt.show()
# +
def hypothesis(X, theta):
return np.matmul(X, theta)
def compute_loss_reg(X, Y, theta, lamb=1):
H = hypothesis(X, theta)
return (np.sum((H - Y)**2) + lamb * np.sum(theta[1:] ** 2)) / (2 * len(X))
def train(Xtrain, Ytrain, epoch, lamb, learning_rate):
theta = np.ones([Xtrain.shape[1], 1])
for i in range(epoch):
H = hypothesis(Xtrain, theta)
delta = np.matmul((H - Ytrain).T, Xtrain) / len(Xtrain)
# print(delta.shape, theta.shape)
delta[:,1:] += lamb * theta[1:].T / len(Xtrain)
# delta[0] -= lamb * (theta.T)[0] / len(Xtrain)
theta -= learning_rate * delta.T
return theta
# Training
Xtrain = np.c_[np.ones(len(X)), X]
theta = train(Xtrain, Y, epoch=4000, lamb=1, learning_rate=0.002)
loss = compute_loss_reg(Xtrain, Y, theta, lamb=1)
print(loss)
# +
# Plot the model
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax1.set_title('Data distribution')
plt.xlabel('Change in water level (x)')
plt.ylabel('Water following out of the dam(y)')
plt.scatter(X, Y, color='red', marker='x')
plt.plot(X, hypothesis(Xtrain, theta))
plt.legend(('Line regression','Training Data'), loc='best')
plt.show()
# +
def plot_learning_curve(X, error_train, error_val):
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax1.set_title('Learning curve for linear regression')
plt.xlabel('Number of training samples')
plt.ylabel('Error ')
plt.plot(range(2, len(X) + 1), error_train)
plt.plot(range(2, len(X) + 1), error_val)
plt.legend(('Train','Cross validation'), loc='best')
plt.show()
# Plot learning curve for training and validation set
error_train = []
error_val = []
for i in range(2, len(X) + 1):
Xtrain = np.c_[np.ones(len(X[:i])), X[:i]]
Ytrain = Y[:i]
theta = train(Xtrain, Ytrain, epoch=1500, lamb=1, learning_rate=0.001)
error_train.append(compute_loss_reg(Xtrain, Ytrain, theta, lamb=0))
error_val.append(compute_loss_reg(np.c_[np.ones(len(Xval)), Xval], Yval, theta, lamb=0))
# print(error_train)
# print(error_val)
plot_learning_curve(X, error_train, error_val)
# -
# ### Polynomial regression
# +
def feature_poly(X, degree):
for i in range(2, degree + 1):
X = np.c_[X, X[:,0]**i]
return X
def feature_norm(X):
mean = np.mean(X, axis=0)
std = np.std(X, axis=0)
return (X - mean) / std
def create_feature(X):
Xtrain = feature_poly(X, 8)
Xtrain = feature_norm(Xtrain)
return np.c_[np.ones(len(Xtrain)), Xtrain]
def plot_poly(X, Y, theta):
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax1.set_title('Data distribution')
plt.xlabel('Change in water level (x)')
plt.ylabel('Water following out of the dam(y)')
plt.scatter(X, Y, color='red', marker='x')
x = np.arange(-80, 80, 0.5).reshape(-1, 1)
plt.plot(x, hypothesis(create_feature(x), theta))
plt.legend(('Line regression','Training Data'), loc='best')
plt.show()
def cal_poly_by_lambda(X, Y, lamb, epoch=40000):
Xtrain = create_feature(X)
theta = train(Xtrain, Y, epoch, lamb=lamb, learning_rate=0.004)
# print(theta, lamb)
# loss = compute_loss_reg(Xtrain, Y, theta, lamb)
# print('loss:', loss)
return theta
def plot_poly_by_lambda(X, Y, lamb):
theta = cal_poly_by_lambda(X, Y, lamb)
plot_poly(X, Y, theta)
def cal_learning_curve_by_lamba(X, Y, lamb):
error_train = []
error_val = []
for i in range(2, len(X) + 1):
theta = cal_poly_by_lambda(X[:i], Y[:i], lamb, epoch=1000)
error_train.append(compute_loss_reg(create_feature(X[:i]), Y[:i], theta, lamb))
error_val.append(compute_loss_reg(create_feature(Xval), Yval, theta, lamb))
return error_train, error_val
def plot_learning_curve_by_lamba(X, Y, lamb):
error_train, error_val = cal_learning_curve_by_lamba(X, Y, lamb)
plot_learning_curve(X, error_train, error_val)
# -
# #### Lambda = 0
# Overfit
plot_poly_by_lambda(X, Y, lamb=0)
# Training loss is always low, but validation loss is higher
plot_learning_curve_by_lamba(X, Y, lamb=0)
# #### Lambda = 1
# It's good
plot_poly_by_lambda(X, Y, lamb=1)
# Loss in both training set and validation set are almost the same
plot_learning_curve_by_lamba(X, Y, lamb=1)
# #### Lambda = 100
# High bias, cannot fit
plot_poly_by_lambda(X, Y, lamb=100)
# Loss are high
plot_learning_curve_by_lamba(X, Y, lamb=100)
# #### Different lambda
# +
# Plot the loss for different lambda
lambs = [0, 0.001, 0.003, 0.01, 0.03, 0.1, 0.3, 1, 3, 10]
error_train = []
error_val = []
for lamb in lambs:
theta = cal_poly_by_lambda(X, Y, lamb)
error_train.append(compute_loss_reg(create_feature(X), Y, theta, lamb))
error_val.append(compute_loss_reg(create_feature(Xval), Yval, theta, lamb))
fig = plt.figure()
ax1 = fig.add_subplot(1, 1, 1)
ax1.set_title('Learning curve for linear regression')
plt.xlabel('Lambda')
plt.ylabel('Error ')
plt.plot(lambs, error_train)
plt.plot(lambs, error_val)
plt.legend(('Train','Cross validation'), loc='best')
plt.show()
# -
# #### Computing test set error
# Lambda = 0.3 should be the best one
lamb = 0.3
theta = cal_poly_by_lambda(X, Y, lamb)
loss = compute_loss_reg(create_feature(Xtest), Ytest, theta, lamb)
print(loss)
# #### Plotting learning curves with randomly selected examples
# +
def cal_learning_curve_by_lamba_randomly(X, Y, lamb):
error_train = []
error_val = []
for i in range(2, len(X) + 1):
losses_train = []
losses_val = []
for _ in range(50):
indexes = np.random.permutation(len(X))[:i]
theta = cal_poly_by_lambda(X[indexes], Y[indexes], lamb, epoch=1000)
losses_train.append(compute_loss_reg(create_feature(X[indexes]), Y[indexes], theta, lamb))
losses_val.append(compute_loss_reg(create_feature(Xval[indexes]), Yval[indexes], theta, lamb))
error_train.append(np.mean(np.array(losses_train)))
error_val.append(np.mean(np.array(losses_val)))
return error_train, error_val
error_train, error_val = cal_learning_curve_by_lamba_randomly(X, Y, lamb=0.01)
plot_learning_curve(X, error_train, error_val)
|
exercise5/estimation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS" tags=[]
# # Path translation and file synchronization
# + [markdown] kernel="SoS" tags=[]
# * **Difficulty level**: intermediate
# * **Time need to lean**: 20 minutes or less
# * A remote host might have different paths from the local host, making the execution of tasks difficult
# * SoS automatically translates paths specified in `_input`, `_depends` and `_output` according to host configurations
# * Options `to_host` and `from_host` specify files and directories send before task execution and retrieve after task execution, respectively.
# * Use of named path could make your workflow more portable and easier to read.
# + [markdown] kernel="SoS"
# ## Translation of input and output paths
# + [markdown] kernel="SoS"
# When local and remote hosts do not share file systems (or share only some file systems), things can get a bit complicated because SoS will need to decide what paths to use on the remote host. There are a few things to understand here:
#
# **The current project directory, and all input, output and dependent files that are involved need to be under paths defined for local and remote host.** This is usually not a problem if you are working under your home directory and you have `home` defined under `paths` of both local and remote hosts, but can become more complicated if your tasks involves system directories such as `resource`, `temp`, and `scratch` that are outside of `home`. In these cases, all involved directories need to be defined for both local and remote hosts.
#
# **Unless specified otherwise, the tasks will be executed under the remote version of the current working directory.**. That is to say, the execution of tasks will leave files on remote hosts that will not be automatically removed, and in a worse scenario **might overwrite remote files without warning**. This is why we recommend that you set remote `home` to a directory other than the true `home` (e.g. `/home/user_name/scratch`, or `/home/user_name/sos_temp`). In this way SoS will write to sos-specified directories on remote hosts and will not containminate your real `home` directory.
#
# **Unless specified otherwise, input and dependent files will be copied to remote host before execution, and output files will be copied to local host after the completion of the task.** It is therefore important for you to plan ahead and avoid synchronization of large files that should stay on remote hosts.
# + [markdown] kernel="SoS" tags=[]
# ## Working directory of tasks (Option `workdir`)
#
# The `workdir` of task is default to the current working directory, or, in the case of remote execution, the remote counterpart of the current working directory.
#
# Option `workdir` controls the working directory of the task. For example, the following step downloads a file to the `resource` directory using [action `download`](download.html).
# + kernel="SoS" tags=[]
task: queue='localhost', workdir='resource'
download:
ftp://speedtest.tele2.net/512KB.zip
# + kernel="SoS" tags=[]
!ls resource
# + [markdown] kernel="SoS" tags=[]
# ## Sending additional files before task execution (Option `to_host`)
# + [markdown] kernel="SoS" tags=[]
# Option `to_host` specifies additional files or directories that would be synchronized to the remote host before tasks are executed. It can be specified as
#
# * A single file or directory (with respect to local file system), or
# * A list of files or directories, or
#
# The files or directories will be translated using the host-specific path maps. Note that if a symbolic link is specified in `to_host`, both the symbolic link and the path it refers to would be synchronized to the remote host.
#
# Just to demontrate how to use this option, let us copy all notebooks in this directory to a remote host and count the number of them.
# + kernel="SoS"
# %preview -n wc.txt
output: 'wc.txt'
task: to_host='task*.ipynb', queue='bcb'
sh: expand=True
wc -l *.ipynb > {_output}
# + [markdown] kernel="SoS" tags=[]
# ## Retrieving additional files after task completion (Option `from_host`)
# + [markdown] kernel="SoS" tags=[]
# Option `from_host` specifies additional files or directories that would be synchronized from the remote host after tasks are executed. It can be specified as
#
# * A single file or directory (with respect to local file system), or
# * A list of files or directories, or
#
# The files or directories will be translated using the host-specific path maps to determine what remote files to retrieve.
# + [markdown] kernel="SoS"
# ## Absolute paths and named paths
# + [markdown] kernel="SoS"
# The use of relative paths are highly recommended because relative paths are not system dependent. Although `data/sample1.csv` can be under different paths on local and remote hosts, SoS handles the mapping of current project directory and `data/sample1.csv` would represent the same file under local and remote hosts.
#
# Things get a lot more complicated when absolute paths are involved. In the following example, `_output` is specified with absolute path, the task still magically works on a cluster system with home directory `/home/bpeng1` because SoS automatically translates input and output files, and knows the output should be `/home/bpeng1/scratch/sos/sos-docs/src/user_guide/random_output.txt` on the remote host. The output files are correctly synchronized to local host.
# + kernel="SoS"
# %preview -n random_output.txt
output: '/Users/bpeng1/sos/sos-docs/src/user_guide/random_output.txt'
task: queue='htc', mem='4G'
import random
with open(_output, 'w') as out:
out.write(f'Random number is {random.randint(0, 1000)}')
# + [markdown] kernel="SoS"
# However, if you execute the workflow directly on the remote host using option `-r`, it would fail because '/Users' is not a writable directory on the remote host.
# + kernel="SoS"
# %env --expect-error
# %run -r htc-headnode
output: '/Users/bpeng1/sos/sos-docs/src/user_guide/random_output.txt'
import random
with open(_output, 'w') as out:
out.write(f'Random number is {random.randint(0, 1000)}')
# + [markdown] kernel="SoS"
# This problem could be solved by the use of host-specific paths. For example, if you are running the work on `htc-headnode`, you can change the output to use the correct path for this host.
# + kernel="SoS"
# %run -r htc-headnode
output: '/home/bpeng1/sos/sos-docs/src/user_guide/random_output.txt'
import random
with open(_output, 'w') as out:
out.write(f'Random number is {random.randint(0, 1000)}')
# + [markdown] kernel="SoS"
# A better choice that would make your workflow more "portable" would be using [named paths](targets.ipynb). For example, if you use `#home` on `htc-headnode` which has the correct named paths defined, the workflow would execute successfully.
# + kernel="SoS"
# %run -r htc-headnode
output: '#home/sos/sos-docs/src/user_guide/random_output.txt'
import random
with open(_output, 'w') as out:
out.write(f'Random number is {random.randint(0, 1000)}')
# + [markdown] kernel="SoS" tags=[]
# ## Further reading
#
# * [The `task_statement`](task_statement.html)
|
src/user_guide/task_files.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="sYAdiHnE8g8w"
# # Pandas
#
# <!--<badge>--><a href="https://colab.research.google.com/github/TheAIDojo/Machine_Learning_Bootcamp/blob/main/Week 02 - Data Science Libraries/2- Pandas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a><!--</badge>-->
#
#
# One of the downsides of NumPy is that it is difficult to work with data that is heterogenous (that is, of mixed data types). Such data is very common, however. Pandas (https://pandas.pydata.org/) is a Python library makes working with such data significantly easier. Similar to NumPy, Pandas is often used to work with tabular data. The two primary data structures are the [Series](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.Series.html) (one-dimensional arrays) and the [DataFrame](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.html#pandas.DataFrame) (two-dimensional arrays, which can be thought of as dictionaries of Series).
#
# More details on Pandas can be found in the following tutorials.
#
# * [Quick Start](https://pandas.pydata.org/pandas-docs/stable/getting_started/10min.html)
# * [Cookbook](https://pandas.pydata.org/pandas-docs/stable/user_guide/cookbook.html#cookbook)
# * [Cheatsheet](https://pandas.pydata.org/Pandas_Cheat_Sheet.pdf)
#
# + [markdown] id="ChSpL5z_8g8w"
# ### Series
#
# Series are one-dimensional arrays in elements have indexes (essentially labels). They can be created in a number of ways, including from a Python dictionary. If no indexes are provided when defined, then integer indexes 0, 1, 2, ..., are used. Series can also be given names.
# + colab={"base_uri": "https://localhost:8080/"} id="I_Rk7Gr-8g8w" outputId="78f560ec-27bc-4102-bb44-a89257ddd9b2" pycharm={"is_executing": false}
import pandas as pd
import numpy as np
series1 = pd.Series([10, 20, 30, 10]) # no indexes or name specified
series2 = pd.Series(
[1, 2, 3, 4.0, 5], index=["first", "second", "third", "fourth", "fifth"], name="s2"
)
series3 = pd.Series({3: "a", "d": 0, "7": 8}, name="s3") # from a dictionary.
series4 = pd.Series(42, range(10), name="s4") # from a scalar, with indexes.
print(series1)
print("-" * 40)
print(series2)
print("-" * 40)
print(series1[2])
print("-" * 40)
print(series3)
print("-" * 40)
print(series4)
# + [markdown] id="3XNXBGBM8g8x"
# You can examine a series using several builtin properties and methods.
# + colab={"base_uri": "https://localhost:8080/"} id="bIm_JCiU8g8x" outputId="422fdb6e-59f4-4c12-cc66-26df777c1e86" pycharm={"is_executing": false}
print(series2.size)
print(series2.shape)
print(series2.dtype)
print(series2.index)
# + [markdown] id="hT85xUZr8g8x"
# ## Accessing Elements and Slicing
#
# Series work much like NumPy ndarrays and can be sliced.
# + colab={"base_uri": "https://localhost:8080/"} id="xVuuvjBI8g8y" outputId="8c17344d-e93c-4efd-8014-ec8ea4bcbff5" pycharm={"is_executing": false}
series = pd.Series({"a": 1, "b": 2, "c": 3})
series["a"] = 10
print(series)
# + colab={"base_uri": "https://localhost:8080/"} id="IzKEvjNo8g8y" outputId="6876109d-b9d4-4aa9-b53f-bfad7f5d96e9" pycharm={"is_executing": false}
series = pd.Series(range(100, 1000, 25), index=list(range(100, 136)), name="s")
series[100] # note we're referencing by label
# + colab={"base_uri": "https://localhost:8080/"} id="_TC1IcoK8g8z" outputId="57ca003d-ca6d-4de7-c131-6def5cdb0428" pycharm={"is_executing": false}
series[[102, 101, 100, 109]] # note we're referencing by label
# + colab={"base_uri": "https://localhost:8080/"} id="6xbwOc5M8g8z" outputId="42c97163-69a8-4ca3-ef87-58a72a49f164" pycharm={"is_executing": false}
series[10:15] # note we're referencing by position
# + colab={"base_uri": "https://localhost:8080/"} id="LEuOxUms8g8z" outputId="a3e98b4d-a59d-4076-b52b-2e6e3fbe3253" pycharm={"is_executing": false}
series[series > 800] # slicing using a boolean condition
# + colab={"base_uri": "https://localhost:8080/"} id="rHFaHTWk8g8z" outputId="30e5175d-4aa0-4f7b-cded-e3ef0816d08b" pycharm={"is_executing": false}
series > 800 # The condition itself returns a new series.
# + colab={"base_uri": "https://localhost:8080/"} id="aWcIH3wo8g80" outputId="967a0b80-a22b-4468-f1a8-22ecce93aee1" pycharm={"is_executing": false}
series.mean()
# + colab={"base_uri": "https://localhost:8080/"} id="CNOHSFY_8g80" outputId="2b99ffa9-a6ee-485b-f7fa-9c608addfcdb" pycharm={"is_executing": false}
series[series < series.mean()]
# + colab={"base_uri": "https://localhost:8080/"} id="65ITMzW08g80" outputId="c770b2d0-f44d-416f-a332-4c7adf83ab6c" pycharm={"is_executing": false}
series == 150
# + [markdown] id="fUPabblz8g80"
# ## DataFrames
#
# DataFrame objects can be viewed as two-dimensional arrays--rows and columns in which the columns may be of different data types. You can create a DataFrame in one of several ways. One way is simply to pass a tabular structure of data (a nested list, or a NumPy ndarray).
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="EnfyAywT8g80" outputId="cd7d7ddd-5800-4f20-fda7-f58611f27237" pycharm={"is_executing": false}
df = pd.DataFrame([[1, 2, 3], [4, 5, 6]])
df
# + [markdown] id="ttpQvopp8g80"
# You can specify labels for the indexes and columns.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="6_vRatYP8g80" outputId="b298768b-8c1b-4d16-f520-2e39a697e31a" pycharm={"is_executing": false}
ar = np.array([[1, 2, 3], [4, 5, 6], [20, 5, 1]])
df = pd.DataFrame(ar, columns=["a", "b", "c"], index=["row1", "row2", "row3"])
df.query("b==5")
# + [markdown] id="TbUrxk4H8g80"
# You can also create a DataFrame from a dictionary of Series.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="Wq5fEZrv8g80" outputId="a73bd12d-934d-484a-8bf0-9c1d43fe7458" pycharm={"is_executing": false}
s1 = pd.Series([1, 4], index=["row1", "row2"])
s2 = pd.Series([2, 5], index=["row1", "row2"])
s3 = pd.Series([3, 6], index=["row1", "row2"])
df = pd.DataFrame({"a": s1, "b": s2, "c": s3})
df
# + [markdown] id="OQbdFMm28g81"
# DataFrames can also be created from a list of dictionaries.
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="WRqQQ_cX8g81" outputId="031283f1-79dc-4096-fc17-c03d73206c0a" pycharm={"is_executing": false}
df = pd.DataFrame([{"a": 1, "b": 2, "c": 3}, {"a": 4, "b": 5, "c": 6}])
df.index = ["row1", "row2"]
df
# + [markdown] id="4mhXDC7q8g81"
# ## Accessing, inserting, deleting, and manipulating columns
#
# Once created, you can refer to columns of data using the column name.
# + colab={"base_uri": "https://localhost:8080/"} id="W0YYk1rx8g81" outputId="ea3b0ce3-b5be-4db0-8b8e-d1d212548105" pycharm={"is_executing": false}
df["a"]
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="E2HbVgjm8g81" outputId="9cecc1c3-d8cd-4e82-b0cf-195e48cf638f" pycharm={"is_executing": false}
df[["a", "c"]]
# + colab={"base_uri": "https://localhost:8080/"} id="Jg29lL8O8g81" outputId="c96ac013-0694-4c14-eb8a-2a73f5c34b6b" pycharm={"is_executing": false}
print(df.columns)
print(df.index)
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="T6w8SAVs8g81" outputId="a8a0ff7c-8434-4426-df8b-c44f91bd77c1" pycharm={"is_executing": false}
df["d"] = df["c"] + 1
df
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="d_hHnfT78g81" outputId="39758e7f-a5b5-4f4c-e53a-fbf1aa809d4e" pycharm={"is_executing": false}
df = df * 2
df
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="Su9dTUnC8g81" outputId="a40352cf-bab8-4e81-a08e-1092d973b18e" pycharm={"is_executing": false}
df["c"] = pd.Series({"row1": 6.5})
df
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="vD3Nl46s8g82" outputId="33526928-42df-4ed4-a365-4ccccb9cfc30" pycharm={"is_executing": false}
df = df.drop(["c"], axis=1)
df
# + [markdown] id="V5t3nBkQ8g82"
# Note the datatype of the following:
# + colab={"base_uri": "https://localhost:8080/"} id="DT54VBb58g82" outputId="8bac505b-d48d-4dae-bccd-e1e9a42d480c" pycharm={"is_executing": false}
type(df["a"])
# + colab={"base_uri": "https://localhost:8080/"} id="THdb0OjA8g82" outputId="727faf63-e126-496b-c575-94d17fbb8be3" pycharm={"is_executing": false}
type(df[["a", "b"]])
# + [markdown] id="4uSVTD4l8g82"
# ## Accessing rows/indexes
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="JW937a-h8g82" outputId="c7bd6d62-4207-48e3-949d-0999573ce710" pycharm={"is_executing": false}
df = pd.DataFrame(
[{"a": 1, "b": 2, "c": 3}, {"a": 4, "b": 5, "c": 6}, {"a": 9, "b": 8, "c": 9}]
)
df.index = ["row1", "row2", "row3"]
df
# + colab={"base_uri": "https://localhost:8080/"} id="eNZIwBIt8g82" outputId="adf55077-fe72-4be0-d4d3-c21228b4604a" pycharm={"is_executing": false}
df.loc["row2"] # by index label
# + colab={"base_uri": "https://localhost:8080/"} id="8m9g7boM8g82" outputId="a1f8b13c-fec9-4f1b-e3ba-a7fe6e75307e" pycharm={"is_executing": false}
df.iloc[1] # by index position
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="3vATseVG8g82" outputId="86f53142-b07d-40d2-cae7-cc1f1bc285cc" pycharm={"is_executing": false}
df[1:]
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="5_YGq98c8g82" outputId="69c93e18-ac18-4dc9-8ba8-c2c93815a56a" pycharm={"is_executing": false}
df.iloc[1:]
# + [markdown] id="FYy-t9t88g83"
# ## Basic Operations on DataFrames
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="zlvdAVYz8g83" outputId="d511e0da-5d66-4c01-e920-d2d5d79e7897" pycharm={"is_executing": false}
df1 = pd.DataFrame(
[[1, 2, 3, 4, 5], [6, 7, 8, 9, 10]], index=["x", "y"], columns=list("abcde")
)
df1
# + colab={"base_uri": "https://localhost:8080/", "height": 162} id="QGqJHswx8g83" outputId="6b04ccb0-0a3e-4158-9b1b-20fafc8239c0" pycharm={"is_executing": false}
df2 = pd.DataFrame(
[[1, 2, 3, 4, 5], [6, 7, 8, 9, 10]], index=["x", "y"], columns=list("abcde")
)
df2 = 1 - df2 % 2
print(df2)
df2 = df2 == 1
df2
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="bxFEBQxk8g83" outputId="0d77c66a-d9c3-4ca0-d770-391265ed1653" pycharm={"is_executing": false}
df3 = df1 > 4
df3
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="GTrFqL1Y8g83" outputId="bc3ab581-4195-400e-92f4-acd84fb80e2d" pycharm={"is_executing": false}
df4 = df2 & df3
df4
# + [markdown] id="DFGp5PqH8g83"
# ## Basic Statistics and other information
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="OM786wLa8g83" outputId="09df088b-a7a3-4a7b-d91a-9a5d5710a3c4" pycharm={"is_executing": false}
df1 = pd.DataFrame(
[[1, 2, 3, 4, 5], [6, 7, 8, 9, 10]],
index=["x", "y"],
columns=["a", "b", "c", "d", "e"],
)
df1
# + colab={"base_uri": "https://localhost:8080/"} id="kf3LjvZ-8g83" outputId="1a2aa614-7944-4a50-b8ef-14b0e038ff1f" pycharm={"is_executing": false}
df1.mean(axis=0) # takes mean for each column
# + colab={"base_uri": "https://localhost:8080/"} id="Re4aIwsw8g83" outputId="abe34e08-e8a7-4d97-e7a7-bb1c44593ac9" pycharm={"is_executing": false}
df1.mean(axis=1) # takes mean for each row
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="Z2TAD2Wp8g83" outputId="066f7c45-db45-438f-8f73-28011cf50aac" pycharm={"is_executing": false}
df1.describe() # Generate descriptive statistics.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="dcJtO6Rh8g83" outputId="9115f87b-862f-40d6-a30e-317bdf1f57c5" pycharm={"is_executing": false}
# we can create DataFrames from Numpy Arrays
numpy_array = np.arange(1000).reshape(
(500, 2)
) # create numpy array with shape (500 ,2 ) start from 0 to 1000
df2 = pd.DataFrame(numpy_array, columns=["a", "b"])
df2.head() # get the first 5 rows from the datframe
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="O30tYBNm8g84" outputId="efad92c0-91bd-42b1-b895-2bd338c6b620" pycharm={"is_executing": false}
df2.tail() # get the last 5 rows from the dataframe
# + [markdown] id="wbpFxSub8g84"
# ## *Reading and Writing CSV files*
#
# Reading in csv files is generally fairly easy in Pandas, but there are many options. See [read_csv](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.read_csv.html) and [to_csv](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_csv.html)
# + id="50o4ysgm8g84" pycharm={"is_executing": false}
df2.to_csv("test.csv", header=True, index=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 392} id="KSJxy7T38g84" outputId="2c6a89ca-2133-44b1-f392-f98e6d3dace3" pycharm={"is_executing": false}
df3 = pd.read_csv("test.csv", header=0)
df3
# + id="D1pwZueG8g84" outputId="2f8a93d9-18da-40cc-f5c2-18389062010b" pycharm={"is_executing": false}
df3 = pd.read_csv("test.csv", header=0, index_col=0)
df3
# + [markdown] id="L1EbgpHs8g84"
# # Plotting in Pandas
#
# It is possible to create figures and charts by invoking methods directly on Pandas objects.
# + id="9rG256Q98g84" outputId="17eef403-e649-4a30-ab8d-67d8f346ebc5" pycharm={"is_executing": false}
import matplotlib.pyplot as plt
x = np.linspace(0, 10, 100)
y = x**2
df = pd.DataFrame(np.array([x, y]).T, columns=["a", "b"])
df.plot()
# + id="TxztBdUt8g84" outputId="68206e15-977a-486c-9f91-b54f6e1b4671" pycharm={"is_executing": false}
x = np.random.rand(10)
y = np.random.rand(10)
df = pd.DataFrame(np.array([x, y]).T, columns=["a", "b"])
df.plot.bar()
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="0jAARlW-8g84" outputId="0b7f8838-eb86-4256-8d94-095e69a479cd" pycharm={"is_executing": false}
x = np.random.randn(200)
df = pd.DataFrame(x, columns=["a"])
df.plot.hist()
# + [markdown] id="AEI4ZmTDaudr"
# # Missing data with pandas
#
# + [markdown] id="DRpcGx1hazsB"
# Pandas treat None and NaN as essentially interchangeable for indicating missing or null values. To facilitate this convention, there are several useful functions for detecting, removing, and replacing null values in Pandas DataFrame :
#
# isnull()
#
# notnull()
#
# dropna()
#
# fillna()
#
# replace()
#
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="WPZh08Kd8g84" outputId="d349b0cd-8367-410b-c320-f7c3a4a6f6d6" pycharm={"is_executing": false}
# Dictionary of lists
data = {
"First Score": [100, 90, np.nan, 95],
"Second Score": [30, 45, 56, np.nan],
"Third Score": [np.nan, 40, 80, 98],
}
# Creating a dataframe from list
df = pd.DataFrame(data)
# Using isnull() function
df.isnull()
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="ztTof67t9ppD" outputId="4d62efce-c06d-4fa3-f57d-ac2d24323ad1"
data = {
"First Score": [100, 90, np.nan, 95],
"Second Score": [30, 45, 56, np.nan],
"Third Score": [np.nan, 40, 80, 98],
}
# creating a dataframe using dictionary
df = pd.DataFrame(data)
# using notnull() function
df.notnull()
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="5Aliu4i_mgIU" outputId="a0e9d0df-24af-46f1-985c-3a9ce56522ac"
data = {
"First Score": [100, 90, np.nan, 95],
"Second Score": [30, 45, 56, np.nan],
"Third Score": [np.nan, 40, 80, 98],
}
# creating a dataframe from dictionary
df = pd.DataFrame(data)
# filling missing value using fillna()
df.fillna(0, inplace=True)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="n3ma5IyGmmmS" outputId="4852028e-361e-40fe-f4ae-c8c0551082a2"
data = {
"First_Score": [100, 90, 80, 95],
"Second_Score": [30, 45, 56, 35],
"Third_Score": [np.nan, 40, 80, 98],
}
# creating a dataframe from dictionary
df = pd.DataFrame(data)
# filling missing value using fillna()
df.Third_Score.fillna(int(df.Third_Score.mean()), inplace=True)
df
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="5k8JUEkWnBKy" outputId="3f5263e2-225f-462d-87f3-33365639850f"
data = {
"First_Score": [100, 90, 80, 95],
"Second_Score": [30, 45, 56, 35],
"Third_Score": [np.nan, 40, 80, 98],
}
# creating a dataframe from dictionary
df = pd.DataFrame(data)
# filling missing value using replace()
df.replace(to_replace=np.nan, value=-99, inplace=True)
df
# + id="XfpTxiznw_9q"
|
Week 02 - Data Science Libraries/2- Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pretrained Models
#
# ## Base learners using pretrained models
#
# ### Base learners using Resnet152, AlexNet, Densenet161
#
# Built using the function that Tyler created
# The tb_preprocess.py script should be in the same folder as the notebook.
#
# ##### Need to add other models
# +
modelnames = {'alexnet':'alexnet','resnet152':'resnet152','densenet161':'densenet161'}
#Set the Model name
model_param = 'resnet152'
filepath = '.'
# +
import os
import random
import numpy as np
import pandas as pd
import torch
import torchvision
import torch.nn as nn
import torch.nn.functional as F
import torchvision.datasets as datasets
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
import time; _START_RUNTIME = time.time()
from torchvision import models
#Use the preprocess function from tb_preprocess.py to load the datasets
from tb_preprocess import load_data
train_loader, val_loader, train_loader_red, val_loader_red = load_data()
idx2class = {'covid-19': 0, 'normal': 1, 'pnuemonia': 2}
classes = ('Covid-19', 'Normal', 'Pneumonia')
# -
# ### Use the model that is pretrained.
#
# ##### Model selection is hardcoded for now
#
# +
red_model = models.__dict__[model_param](pretrained=True)
#Instantiating CUDA device
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
#Verifying CUDA
print(device)
#Move the input and model to GPU for speed if available
red_model.to(device)
# -
# #### Change the last linear layer to match the number of classes. In this case 3.
#
# ##### Eg:- The standard Alexnet model has 1000 out features for the last Linear layer. Since the data set only has 3 classes, change the last layer to have 3 outs
# +
if model_param == 'alexnet':
red_model.classifier[6] = torch.nn.Linear(red_model.classifier[6].in_features, 3, bias=True)
elif model_param == 'resnet152':
red_model.fc = torch.nn.Linear(2048, 3, bias=True)
elif model_param == 'densenet161':
red_model.classifier = torch.nn.Linear(2208, 3, bias=True)
red_model.eval()
if model_param == 'alexnet':
red_model.features = torch.nn.DataParallel(red_model.features)
red_model.cuda()
else:
red_model = torch.nn.DataParallel(red_model).cuda()
# -
# #### Criterion and Optimizer
# +
import torch.optim as optim
#Loss
criterion = nn.CrossEntropyLoss()
#Optimizer(SGD)
optimizer = optim.SGD(red_model.parameters(), lr=0.001, momentum=0.9)
# -
# ### Training Alexnet
# +
#Epochs is set to 1. Performace is slow
#Need to try CUDA to speed up
n_epochs = 25
def train_model(model, train_dataloader, n_epoch=n_epochs, optimizer=optimizer, criterion=criterion):
import torch.optim as optim
# prep model for training
model.train()
for epoch in range(n_epoch):
EPOCH_START = time.time()
curr_epoch_loss = []
for data, target in train_dataloader:
data, target = data.cuda(), target.cuda()
optimizer.zero_grad()
output = model(data)
#print(output)
loss = criterion(output, target)
loss.backward()
optimizer.step()
curr_epoch_loss.append(loss.cpu().data.numpy())
print(f"Epoch {epoch}: curr_epoch_loss={np.mean(curr_epoch_loss)}")
print("Training time = {:.2f} seconds".format(time.time() - EPOCH_START))
return model
# -
# ### Save the model
# +
trainedmodel = model_param + '.pth'
if os.path.exists(trainedmodel):
print('Model is trained - ' + model_param)
red_model = torch.load(trainedmodel)
else:
red_model = train_model(red_model, train_loader)
torch.save(red_model, trainedmodel)
print("Total running time = {:.2f} seconds".format(time.time() - _START_RUNTIME))
# -
# #### Testing
# +
def eval_model(model, dataloader):
model.eval()
Y_pred = []
Y_test = []
for data, target in dataloader:
data, target = data.cuda(), target.cuda()
output = model(data)
#print(output.data)
#print(target)
_, y_predicted = torch.max(output.data, 1)
Y_pred = np.append(Y_pred,y_predicted.detach().cpu().numpy())
Y_test = np.append(Y_test,target.detach().cpu().numpy())
return Y_pred, Y_test
from sklearn.metrics import accuracy_score,classification_report, confusion_matrix, ConfusionMatrixDisplay
y_pred, y_true = eval_model(red_model, val_loader)
acc = accuracy_score(y_true, y_pred)
print(("Validation Accuracy: " + str(acc)))
# Classification Report
print(classification_report(y_true, y_pred))
# Confusion Matrix
print('Confusion Matrix')
print(confusion_matrix(y_true, y_pred))
confusion_matrix_df = pd.DataFrame(confusion_matrix(y_true, y_pred)).rename(columns=idx2class, index=idx2class)
cm = confusion_matrix(y_true, y_pred)
#print(cm)
print('Confusion Matrix Display')
cm_display = ConfusionMatrixDisplay(cm,classes).plot()
# -
# ## Testing Accuracy
#
# Classification accuracy of the test model.
# +
correct = 0
total = 0
# no_grad impacts the autograd engine and deactivate it. It will reduce memory usage and speed up computations
with torch.no_grad():
for data in val_loader:
images, labels = data[0].to(device), data[1].to(device)
outputs = red_model(images)
#print(outputs)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on test X-rays : %d %%' % ( 100 * correct / total))
# -
# ## Individual Classes
#
# +
#Testing classification accuracy for individual classes.
class_correct = list(0. for i in range(3))
class_total = list(0. for i in range(3))
with torch.no_grad():
for data in val_loader:
images, labels = data[0].to(device), data[1].to(device)
outputs = red_model(images)
_, predicted = torch.max(outputs, 1)
c = (predicted == labels).squeeze()
for i in range(4):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
for i in range(3):
print('Accuracy of %5s : %2d %%' % (
classes[i], 100 * class_correct[i] / class_total[i]))
|
Code/red_base_learners_cuda.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# utils
import re
import os
from glob import glob
import concurrent.futures
import time
import pandas as pd
from datetime import datetime
# tesseract
import pytesseract
# opencv
import cv2
os.environ['OMP_THREAD_LIMIT'] = '6'
# -
df = pd.read_csv('./is_ocr.csv')
# +
# drop unused columns
unused_columns = [
'process_class',
'process_processing_date',
'page_is_ocr',
'process_process_time',
'process_is_complete',
'document_processing_date',
'document_processing_time',
'page_image',
'page_piece'
]
new_df = df
for column in unused_columns:
new_df = new_df.drop(column, axis=1)
# add new column to dataframe
new_df['crappy_ocr_text'] = None
# add new column to join process_id and document_id
new_df['id'] = new_df['process_id'].map(lambda x: str(x) + '_') + new_df['document_id'].map(lambda x: str(x) + '_') + new_df['page_number'].map(str)
new_df = new_df.set_index('id')
new_df = new_df.sort_index()
new_df.head()
# +
def extract_and_add_to_df(path):
"""
Receive a image path, extract it's content using tesseract and add it to dataframe.
"""
# get image id
current_id = re.sub('./crappy_images\/crappy_([\d]+_[\d]+_[\d]+)\.jpg', '\\1', path)
# read image
image = cv2.imread(path)
# extract text
text = pytesseract.image_to_string(image, lang='por+eng')
# # add to dataframe
# new_df.loc[current_id, 'crappy_ocr_text'] = text
return [current_id, text]
# +
crappy_folder = './crappy_images/'
image_list = glob(crappy_folder + '*.jpg')
size = len(image_list)
values = []
with concurrent.futures.ProcessPoolExecutor(max_workers=6) as executor:
count = 0
current_line = 0
times = 0
print('[{}] Info: Pipeline has started. Ammount of data: {}'.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S'), size))
for img_path, result in zip(image_list, executor.map(extract_and_add_to_df, image_list)):
values.append(result)
current_line += 1
count += 1
if count == 1000:
times += 1
print('[{}] Info: Pipeline has processed and generated {} crappy images.'.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S'), times * count))
count = 0
# -
for value in values:
current_id, text = value
new_df.loc[current_id, 'crappy_ocr_text'] = text
new_df.to_csv('./ocr_results.csv')
|
ocr_pipeline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('ignore')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
from sklearn import cluster, mixture
from sklearn.metrics import calinski_harabasz_score, davies_bouldin_score, silhouette_score
data = pd.read_csv("./data_transformed.csv", index_col=0)
data
def create_clustering_algorithms(n_clusters):
minibatch = cluster.MiniBatchKMeans(n_clusters=n_clusters)
agglomerative_ward = cluster.AgglomerativeClustering(n_clusters=n_clusters, linkage='ward')
agglomerative_average = cluster.AgglomerativeClustering(n_clusters=n_clusters, linkage="average")
birch = cluster.Birch(n_clusters=n_clusters)
gmm = mixture.GaussianMixture(n_components=n_clusters)
clustering_algorithms = (('MiniBatchKMeans', minibatch),
('Agglomerative_Ward', agglomerative_ward),
('Agglomerative_Average', agglomerative_average),
('Birch', birch),
('GaussianMixture', gmm))
return clustering_algorithms
algorithms_names = ['MiniBatchKMeans', 'Agglomerative_Ward', 'Agglomerative_Average', 'Birch', 'GaussianMixture']
# + code_folding=[8]
def calculate_cluster_metrics(n_cluster_min, n_cluster_max, dataset):
silhouette_scores = pd.DataFrame(index=range(2, 15), columns=algorithms_names)
davies_bouldin_scores = pd.DataFrame(index=range(2, 15), columns=algorithms_names)
calinski_harabasz_scores = pd.DataFrame(index=range(2, 15), columns=algorithms_names)
for n_clusters in range(n_cluster_min, n_cluster_max):
clustering_algorithms = create_clustering_algorithms(n_clusters)
for name, algorithm in clustering_algorithms:
algorithm.fit(dataset)
if hasattr(algorithm, 'labels_'):
y_pred = algorithm.labels_.astype(np.int)
else:
y_pred = algorithm.predict(dataset)
db_score = davies_bouldin_score(dataset, y_pred)
ch_score = calinski_harabasz_score(dataset, y_pred)
s_score = silhouette_score(dataset, y_pred)
davies_bouldin_scores.loc[n_clusters, name] = db_score
calinski_harabasz_scores.loc[n_clusters, name] = ch_score
silhouette_scores.loc[n_clusters, name] = s_score
return silhouette_scores, davies_bouldin_scores, calinski_harabasz_scores
# -
silhouette_scores, davies_bouldin_scores, calinski_harabasz_scores = calculate_cluster_metrics(2, 15, data)
def plot_cluster_metrics(silhouette_scores, davies_bouldin_scores, calinski_harabasz_scores):
fig, axes = plt.subplots(nrows=1, ncols=3, figsize=(15, 5))
davies_bouldin_scores.plot(ax=axes[0], legend=None, title='davies_bouldain')
calinski_harabasz_scores.plot(ax=axes[1], legend=None, title='calinski_harabasz')
silhouette_scores.plot(ax=axes[2], title='silhouette');
plot_cluster_metrics(silhouette_scores, davies_bouldin_scores, calinski_harabasz_scores)
# For davies_bouldain and silhouette score Agglomerative Clustering with Average Linkage is significantly better than all other algorithms,
# while for calinski_harabasz score it is surprisingly the worst algorithm. Moreover, most scores decrease with number of clusters, suggesting that data is not easily separated and there is no one natural number of clusters that maximize (or minimize, regarding davies bouldain score) aformentioned scores.
full_data = pd.read_csv("./AllBooks_baseline_DTM_Labelled.csv", index_col=0)
# Only 100 most popular words
data_top_100 = full_data.loc[:, full_data.sum(axis=0).sort_values(ascending=False)[:200].index]
silhouette_scores_top_100, davies_bouldin_scores_top_100, calinski_harabasz_scores_top_100 = calculate_cluster_metrics(2, 15, data_top_100)
plot_cluster_metrics(silhouette_scores_top_100, davies_bouldin_scores_top_100, calinski_harabasz_scores_top_100)
# For data consisting of 100 most popular words results are similar, with agglomerative clustering with average linkage still being the outlier.
# +
from yellowbrick.cluster import KElbowVisualizer
model =cluster.KMeans()
visualizer = KElbowVisualizer(model, k=(2,15),timings=False)
visualizer.fit(data)
visualizer.show();
# -
# No significant point in elbow plot suggests once again that data is not easily separable.
from yellowbrick.cluster import intercluster_distance
intercluster_distance(cluster.KMeans(5, random_state=42), data);
# Projecting data into 2 dimensions shows that somehow clustering algorithm found way to separate at least one distinct cluster
from yellowbrick.cluster import silhouette_visualizer
silhouette_visualizer(cluster.KMeans(5, random_state=42), data, colors='yellowbrick');
# Analyzing silhouette diagrams results in a bit of a headache:
# green cluster and very small yellow cluster are very well separated.
# Blue cluster is descent, while both red and purple have many observations that might have been asigned to the wrong clusterster.
#
# TODO:
# - different clustering algorithms: DBSCAN, HDBSCAN, genie, meanshift, affinity propagation
# - use other metrics based on truth labels: Adjusted Rand index, Fowlkes-Mallows index
|
Projekty/Projekt2/Grupa2/NguyenPiatyszekPingielski/Milestone_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="g_nWetWWd_ns"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="2pHVBk_seED1"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" colab={} colab_type="code" id="N_fMsQ-N8I7j"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] colab_type="text" id="pZJ3uY9O17VN"
# # Сохраняй и загружай модели
# + [markdown] colab_type="text" id="M4Ata7_wMul1"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/save_and_restore_models"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Читай на TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/ru/tutorials/keras/save_and_restore_models.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запусти в Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/ru/tutorials/keras/save_and_restore_models.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучай код на GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="mBdde4YJeJKF"
# Прогресс обучения моделей можно сохранять во время и после обучения: тренировку можно возобновить с того места, где ты остановился. Это обычно помогает избежать долгих бесперервыных сессий обучения. Сохраняя модель, ты также можешь поделиться ею с другими, чтобы они могли воспроизвести результаты ее работы. Большинство практиков машинного обучения помимо самой модели и использованных техник также публикуют:
#
# * Код, при помощи которого обучалась модель
# * Тренировочные веса, или параметры модели
#
# Публикация этих данных помогает другим понять как работает модель, а также они смогут проверить как она ведет себя с новыми данными.
#
# Внимание! Будь осторожен с кодом, которому ты не доверяешь. Обязательно прочти [Как использовать TensorFlow безопасно?](https://github.com/tensorflow/tensorflow/blob/master/SECURITY.md)
#
# ### Варианты
#
# Существуют разные способы сохранять модели TensorFlow - все зависит от API, которые ты использовал в своей модели. В этом уроке используется [tf.keras](https://www.tensorflow.org/guide/keras), высокоуровневый API для построения и обучения моделей в TensorFlow. Для всех остальных подходов читай руководство по TensorFlow [Сохраняй и загружай модели](https://www.tensorflow.org/guide/saved_model) или [Сохранение в Eager](https://www.tensorflow.org/guide/eager#object_based_saving).
# + [markdown] colab_type="text" id="xCUREq7WXgvg"
# ## Настройка
#
# ### Настроим и импортируем зависимости
# + [markdown] colab_type="text" id="7l0MiTOrXtNv"
# Установим и импортируем TensorFlow и все зависимые библиотеки:
# + colab={} colab_type="code" id="RzIOVSdnMYyO"
# !pip install h5py pyyaml
# + [markdown] colab_type="text" id="SbGsznErXWt6"
# ### Загрузим датасет
#
# Мы воспользуемся [датасетом MNIST](http://yann.lecun.com/exdb/mnist/) для обучения нашей модели, чтобы показать как сохранять веса. Ускорим процесс, используя только первые 1000 образцов данных:
# + colab={} colab_type="code" id="7Nm7Tyb-gRt-"
from __future__ import absolute_import, division, print_function
import os
import tensorflow as tf
from tensorflow import keras
tf.__version__
# + colab={} colab_type="code" id="9rGfFwE9XVwz"
(train_images, train_labels), (test_images, test_labels) = tf.keras.datasets.mnist.load_data()
train_labels = train_labels[:1000]
test_labels = test_labels[:1000]
train_images = train_images[:1000].reshape(-1, 28 * 28) / 255.0
test_images = test_images[:1000].reshape(-1, 28 * 28) / 255.0
# + [markdown] colab_type="text" id="anG3iVoXyZGI"
# ### Построим модель
# + [markdown] colab_type="text" id="wynsOBfby0Pa"
# Давай построим простую модель, на которой мы продемонстрируем как сохранять и загружать веса моделей:
# + colab={} colab_type="code" id="0HZbJIjxyX1S"
# Возвращает короткую последовательную модель
def create_model():
model = tf.keras.models.Sequential([
keras.layers.Dense(512, activation=tf.nn.relu, input_shape=(784,)),
keras.layers.Dropout(0.2),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
model.compile(optimizer=tf.train.AdamOptimizer(),
loss=tf.keras.losses.sparse_categorical_crossentropy,
metrics=['accuracy'])
return model
# Создадим модель
model = create_model()
model.summary()
# + [markdown] colab_type="text" id="soDE0W_KH8rG"
# ## Сохраняем контрольные точки
# + [markdown] colab_type="text" id="mRyd5qQQIXZm"
# Основная задача заключается в том, чтобы автоматически сохранять модель как *во время*, так и *по окончании* обучения. Таким образом ты сможешь снова использовать модель без необходимости обучать ее заново, или просто продолжить с места, на котором обучение было приостановлено.
#
# Эту задачу выполняет функция обратного вызова `tf.keras.callbacks.ModelCheckpoint`. Эта функция также может быть настроена при помощи нескольких аргументов.
#
# ### Использование функции
#
# Обучим нашу модель и передадим ей функцию `ModelCheckpoint`:
# + colab={} colab_type="code" id="IFPuhwntH8VH"
checkpoint_path = "training_1/cp.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
# Создадим контрольную точку при помощи callback функции
cp_callback = tf.keras.callbacks.ModelCheckpoint(checkpoint_path,
save_weights_only=True,
verbose=1)
model = create_model()
model.fit(train_images, train_labels, epochs = 10,
validation_data = (test_images,test_labels),
callbacks = [cp_callback]) # передаем callback обучению
# + [markdown] colab_type="text" id="rlM-sgyJO084"
# Это создаст одну совокупность файлов контрольных точек TensorFlow, которые обновлялись в конце каждой эпохи:
# + colab={} colab_type="code" id="gXG5FVKFOVQ3"
# !ls {checkpoint_dir}
# + [markdown] colab_type="text" id="wlRN_f56Pqa9"
# Теперь создадим новую необученную модель. Когда мы восстанавливаем модель только из весов, новая модель должна быть точно такой же структуры, как и старая. Посколько архитектура модель точно такая же, то мы можем опубликовать веса из другой *инстанции* модели.
#
# Также мы оценим точность новой модели на проверочных данных. Необученная модель будет лишь изредка угадывать правильную категорию обзоров фильмов (точность будет около 10%):
# + colab={} colab_type="code" id="Fp5gbuiaPqCT"
model = create_model()
loss, acc = model.evaluate(test_images, test_labels)
print("Необученная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="1DTKpZssRSo3"
# А теперь загрузим веса из контрольной точки и проверим еще раз:
# + colab={} colab_type="code" id="2IZxbwiRRSD2"
model.load_weights(checkpoint_path)
loss,acc = model.evaluate(test_images, test_labels)
print("Восстановленная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="bpAbKkAyVPV8"
# ### Параметры вызова контрольной точки
#
# У callback функции есть несколько параметров, которые дают контрольным точкам уникальные имена, а также корректируют частоту сохранения.
#
# Обучим новую модель и укажем параметр чтобы сохранять контрольные точки через каждые 5 эпох:
#
# + colab={} colab_type="code" id="mQF_dlgIVOvq"
# Укажем эпоху в имени файла (переведем ее в строки при помощи `str.format`)
checkpoint_path = "training_2/cp-{epoch:04d}.ckpt"
checkpoint_dir = os.path.dirname(checkpoint_path)
cp_callback = tf.keras.callbacks.ModelCheckpoint(
checkpoint_path, verbose=1, save_weights_only=True,
# Сохраняем веса через каждые 5 эпох
period=5)
model = create_model()
model.fit(train_images, train_labels,
epochs = 50, callbacks = [cp_callback],
validation_data = (test_images,test_labels),
verbose=0)
# + [markdown] colab_type="text" id="1zFrKTjjavWI"
# Теперь посмотрим на получившиеся контрольные точки и выберем последнюю:
# + colab={} colab_type="code" id="p64q3-V4sXt0"
# ! ls {checkpoint_dir}
# + colab={} colab_type="code" id="1AN_fnuyR41H"
latest = tf.train.latest_checkpoint(checkpoint_dir)
latest
# + [markdown] colab_type="text" id="Zk2ciGbKg561"
# Помни: по умолчанию TensorFlow сохраняет только 5 последних контрольных точек.
#
# Для проверки восстановим модель по умолчанию и загрузим последнюю контрольную точку:
# + colab={} colab_type="code" id="3M04jyK-H3QK"
model = create_model()
model.load_weights(latest)
loss, acc = model.evaluate(test_images, test_labels)
print("Восстановленная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="c2OxsJOTHxia"
# ## Как выглядят эти файлы?
# + [markdown] colab_type="text" id="JtdYhvWnH2ib"
# Код выше сохраняет веса модели как совокупность [контрольных точек](https://www.tensorflow.org/guide/saved_model#save_and_restore_variables) - форматированных файлов, которые содержат только обученные веса в двоичном формате. Они включают в себя:
# * Один или несколько шардов (shard, пер. "Часть данных"), в которых хранятся веса твоей модели
# * Индекс, который указывает какие веса хранятся в каждом шарде
#
# Если ты обучаешь модель на одном компьютере, то тогда у тебя будет всего один шард, оканчивающийся на `.data-00000-of-00001`
# + [markdown] colab_type="text" id="S_FA-ZvxuXQV"
# ## Сохраняем веса вручную
#
# Выше мы посмотрели как загружать веса в модель.
#
# Сохранять веса вручную так же просто, просто воспользуйся методом `Model.save_weights`:
# + colab={} colab_type="code" id="R7W5plyZ-u9X"
# Сохраняем веса
model.save_weights('./checkpoints/my_checkpoint')
# Восстанавливаем веса
model = create_model()
model.load_weights('./checkpoints/my_checkpoint')
loss,acc = model.evaluate(test_images, test_labels)
print("Восстановленная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="kOGlxPRBEvV1"
# ## Сохраняем модель целиком
#
# Ты также можешь сохранить модель целиком в единый файл, который будет содержать все веса, конфигурацию модели и даже оптимизатор конфигурации (однако это зависит от выбранных параметров). Это позволит тебе восстановить модель и продолжить обучение позже, ровно с того момента, где ты остановился, и без правки изначального кода.
#
# Сохранять рабочую модель полностью весьма полезно. Например, ты можешь потом восстановить ее в TensorFlow.js ([HDF5](https://js.tensorflow.org/tutorials/import-keras.html), [Сохраненные модели](https://js.tensorflow.org/tutorials/import-saved-model.html)) и затем обучать и запускать ее в веб-браузерах, или конвертировать ее в формат для мобильных устройств, используя TensorFlow Lite ([HDF5](https://www.tensorflow.org/lite/convert/python_api#exporting_a_tfkeras_file_), [Сохраненные модели](https://www.tensorflow.org/lite/convert/python_api#exporting_a_savedmodel_))
# + [markdown] colab_type="text" id="SkGwf-50zLNn"
# ### Сохраняем в формате HDF5
#
# В Keras есть встроенный формат для сохранения модель при помощи стандарта [HDF5](https://en.wikipedia.org/wiki/Hierarchical_Data_Format). Для наших целей сохраненная модель будет использована как единый двоичный объект *blob*.
# + colab={} colab_type="code" id="m2dkmJVCGUia"
model = create_model()
# Используй keras.optimizer чтобы восстановить оптимизатор из файла HDF5
model.compile(optimizer=keras.optimizers.Adam(),
loss=tf.keras.losses.sparse_categorical_crossentropy,
metrics=['accuracy'])
model.fit(train_images, train_labels, epochs=5)
# Сохраним модель полностью в единый HDF5 файл
model.save('my_model.h5')
# + [markdown] colab_type="text" id="GWmttMOqS68S"
# Теперь воссоздадим модель из этого файла:
# + colab={} colab_type="code" id="5NDMO_7kS6Do"
# Воссоздадим точно такую же модель, включая веса и оптимизатор:
new_model = keras.models.load_model('my_model.h5')
new_model.summary()
# + [markdown] colab_type="text" id="JXQpbTicTBwt"
# Проверим ее точность:
# + colab={} colab_type="code" id="jwEaj9DnTCVA"
loss, acc = new_model.evaluate(test_images, test_labels)
print("Восстановленная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="dGXqd4wWJl8O"
# Данная техника сохраняет все:
#
# * Веса модели
# * Конфигурацию (ее структуру)
# * Параметры оптимизатора
#
# Keras сохраняет модели путем исследования ее архтикетуры. В настоящее время он не может сохранять оптимизаторы TensorFlow из `tf.train`. Когда ты используешь их тебе будет нужно скомпилировать модель еще раз после загрузки. Таким образом ты получишь параметры оптимизатора.
#
# + [markdown] colab_type="text" id="kPyhgcoVzqUB"
# ### Сохраняем как `saved_model`
# + [markdown] colab_type="text" id="LtcN4VIb7JkK"
# Обрати внимание: этот метод сохранения моделей `tf.keras` является экспериментальным и может измениться в будущих версиях.
# + [markdown] colab_type="text" id="DSWiSB0Q8c46"
# Построим новую модель:
# + colab={} colab_type="code" id="sI1YvCDFzpl3"
model = create_model()
model.fit(train_images, train_labels, epochs=5)
# + [markdown] colab_type="text" id="iUvT_3qE8hV5"
# Создадим `saved_model`:
# + colab={} colab_type="code" id="sq8fPglI1RWA"
saved_model_path = tf.contrib.saved_model.save_keras_model(model, "./saved_models")
# + [markdown] colab_type="text" id="MjpmyPfh8-1n"
# Сохраненные модели будут помещены в папку и отмечены текущей датой и временем в названии:
# + colab={} colab_type="code" id="ZtOvxA7V0iTv"
# !ls saved_models/
# + [markdown] colab_type="text" id="B7qfpvpY9HCe"
# Загрузим новую модель Keras из уже сохраненной:
# + colab={} colab_type="code" id="0YofwHdN0pxa"
new_model = tf.contrib.saved_model.load_keras_model(saved_model_path)
new_model
# + [markdown] colab_type="text" id="uWwgNaz19TH2"
# Запустим загруженную модель:
# + colab={} colab_type="code" id="Pc9e6G6w1AWG"
# Оптимизатор не был восстановлен, поэтому мы укажим новый
new_model.compile(optimizer=tf.train.AdamOptimizer(),
loss=tf.keras.losses.sparse_categorical_crossentropy,
metrics=['accuracy'])
loss, acc = new_model.evaluate(test_images, test_labels)
print("Загруженная модель, точность: {:5.2f}%".format(100*acc))
# + [markdown] colab_type="text" id="eUYTzSz5VxL2"
# ## Что дальше?
#
# Это был короткий урок по сохранению и загрузке своих моделей при помощи `tf.kers`.
#
# * В [руководстве по tf.keras](https://www.tensorflow.org/guide/keras) рассказывается подробнее о том, как можно сохранять и загружать модели при помощи `tf.keras`
#
# * Статья [Сохраняй в Eager](https://www.tensorflow.org/guide/eager#object_based_saving) рассказывает как сохранять модель во время Eager Execution
#
# * В руководстве [Сохраняй и загружай модели](https://www.tensorflow.org/guide/saved_model) содержится подробный урок обо всех низкоуровневых деталях сохранения моделей TensorFlow
|
site/ru/tutorials/keras/save_and_restore_models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Assignment 1
# ### Question 1
# Write a program which will find all such numbers which are divisible by 7 but are not a multiple of 5, between 2000 and 3200 (both included). The numbers obtained should be printed in a comma-separated sequence on a single line.
lst = range(2000,3201)
for i in lst:
if i%7==0 and i%5!=0:
print(i,end=',')
# ### Question 2
# Write a Python program to accept the user's first and last name and then getting them printed in the the reverse order with a space between first name and last name.
fstname = input("Enter your First Name : ")
lstname = input("Enter your Last Name : ")
f = fstname[::-1]
l = lstname[::-1]
print(l,'',f)
# ### Question 3
# Write a Python program to find the volume of a sphere with diameter 12 cm. Formula: V=4/3 * π * r^3
d = 12
r = d/2
V = (4/3)*(22/7)*r*r*r
print(V)
|
Assignment 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Natural Language Processing with Embeddig Layer
from IPython.display import Image
# %load_ext nb_black
import numpy as np
import pandas as pd
import tensorflow
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import (
Activation,
BatchNormalization,
Dense,
Flatten,
LSTM,
)
from tensorflow.keras.preprocessing.text import one_hot, Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
# ## Esempio classificazione testi
# define documents
docs = [
"Well done!",
"Good work",
"Great effort",
"nice work",
"Excellent!",
"Weak",
"Poor effort!",
"not good",
"poor work",
"Could have done better.",
]
# define class labels
labels = np.array([1, 1, 1, 1, 1, 0, 0, 0, 0, 0])
for x, y in zip(docs, labels):
print(f"Frase: {x} -> Classe: {y}")
# ### Preprocessing dei dati
# Le parole verranno trasformare in uno scalare, non più in un vettore di "uno" e "zeri" il cui indice rappresenta la parola.
vocab_size = 14
encoded_docs = [one_hot(d, vocab_size) for d in docs]
print(encoded_docs)
# I dati, come per un normale dataset, devono avere lo stesso numero di features. Per far ciò si procede con il padding, ovvero aggiungendo "zeri" fino a raggiungere la lunghezza necessaria.
max_length = 4
padded_docs = pad_sequences(encoded_docs, maxlen=max_length, padding="post")
print(padded_docs)
tokenizer = Tokenizer(
num_words=vocab_size, # numero massimo di parole da considerare compreso OOV
)
tokenizer.fit_on_texts(docs)
sequences = tokenizer.texts_to_sequences(docs)
sequences = pad_sequences(sequences, maxlen=max_length, padding="post", dtype="int8")
sequences
Image('ohe_sentence.png')
docs[0]
docs[9]
tokenizer.index_word
tokenizer.sequences_to_texts(sequences)
tokenizer.word_index
# ## Embedding Layer
from tensorflow.keras.layers import Embedding
vocab_size
# A questo punto è possibile costruire il modello.
model = Sequential()
model.add(Embedding(vocab_size + 1, 3, input_length=max_length))
####################
model.add(Flatten())
####################
model.add(Dense(1, activation="sigmoid"))
model.compile(optimizer="adam", loss="binary_crossentropy", metrics=["accuracy"])
print(model.summary())
model.fit(padded_docs, labels, epochs=50, verbose=0)
loss, accuracy = model.evaluate(padded_docs, labels, verbose=0)
print("Accuracy: %0.2f" % (accuracy * 100))
padded_docs.shape
labels.shape
model = Sequential()
model.add(Embedding(vocab_size + 1, 3, input_length=4))
###################
model.add(LSTM(32))
###################
model.add(Dense(1, activation="sigmoid"))
model.compile(optimizer="RMSprop", loss="binary_crossentropy", metrics=["accuracy"])
model.fit(padded_docs, labels, epochs=50, verbose=1)
loss, accuracy = model.evaluate(padded_docs, labels, verbose=0)
print("Accuracy: %0.2f" % (accuracy * 100))
Image('Steps.png')
emb_layer = model.get_layer(index=0)
emb_layer.embeddings
# ## Introduzione a GloVe
# E' un algoritmo non supervisionato che serve per ottenere una rappresentazione vettoriale delle parole.
# La rappresentazione vettoriale delle parole permette di inserire nello stesso claster (quindi vicine in termini spaziali) parole simili.
# +
emb_index = {}
f = open("vectors.txt")
for line in f:
values = line.split()
word = values[0]
coefs = np.array(values[1:], dtype="float32")
emb_index[word] = coefs
f.close()
# -
emb_index["canzoni"]
EMBEDDING_DIM = emb_index["canzoni"].shape[0]
EMBEDDING_DIM
vocab_size
embedding_matrix = np.zeros((vocab_size + 1, EMBEDDING_DIM))
for word, i in tokenizer.word_index.items():
# print(i, word)
if word in emb_index.keys():
emb_vector = emb_index[word]
embedding_matrix[i] = emb_vector
embedding_matrix.shape
embedding_matrix[0]
embedding_matrix[1]
model = Sequential()
model.add(
Embedding(
vocab_size + 1, 100, input_length=4, weights=[embedding_matrix], trainable=False
)
)
###################
model.add(LSTM(32))
###################
model.add(Dense(1, activation="sigmoid"))
model.compile(optimizer="RMSprop", loss="binary_crossentropy", metrics=["accuracy"])
model.fit(padded_docs, labels, epochs=50, verbose=1)
loss, accuracy = model.evaluate(padded_docs, labels, verbose=0)
print("Accuracy: %0.2f" % (accuracy * 100))
|
IntroDeepLearning/Serata 3 - Recurrent Neural Net/NLP_with_RNN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="7KzCr5LXMjBO"
# ## Tutorial notebook
# + [markdown] pycharm={"name": "#%% md\n"}
# This Jupyter notebook shows how to perform a basic analysis of γ-ray photon-count maps using the convolutional neural network-based method presented in [arXiv:2107.09070](http://arxiv.org/abs/2107.09070).
#
# In this example, the photon-count maps consist of **three** different emission components:
# 1. *Fermi* bubbles (Poissonian)
# 2. Galactic Center Excess (point source-like, single population)
# 3. Isotropic point sources (point source-like, two populations in each map).
#
# As discussed in the paper, for the point source-like templates the Poissonian case is included as the limit of ultra-faint point source emission (<< 1 photon expected per source) where the neural network can no longer distinguish point sources from Poissonian emission.
#
# To consider different scenarios (e.g. other templates, more training data, different network architectures, etc.), simply modify the sample parameter file ```GCE_NN/parameter_files/parameters.py``` accordingly.
# The available templates can be viewed in the function ```get_templates()``` in ```GCE/data_utils.py```.
#
# Also, if you don't have access to a GPU and just want to try out the code, it is recommended to reduce the number of training steps in the ```parameters.py```
# file in the folder ```parameter_files``` under "Training settings" from ```2500``` to e.g. ```p_train['num_steps'] = 500``` to reduce the
# training time. In this case, you will see a warning
# ```
# "WARNING:tensorflow:There are non-GPU devices in `tf.distribute.Strategy`...
# ```
# + id="VLj9BMSPMF9V" colab={"base_uri": "https://localhost:8080/"} outputId="da87b8d4-a5c5-4c2b-c51f-e493dc7dbe2f"
from matplotlib import pyplot as plt
# %matplotlib inline
import numpy as np
import healpy as hp
import os
# + colab={"base_uri": "https://localhost:8080/"} id="KIx_p5alYlU5" outputId="18e4bbe0-da02-410c-89f8-6efb72116610"
# %pip list | grep gce-nn # check if gce-nn module is there
import GCE.gce
# + [markdown] id="bE7XSJ7rOjgo"
# First, we need to **initialize** an analysis object.
# + colab={"base_uri": "https://localhost:8080/"} id="21lsX63-OjNU" outputId="8abe98d5-54b6-4426-f5f5-b121668b95c2"
gce = GCE.gce.Analysis()
# + [markdown] id="gfeV5afmO7pp"
# Now, let's **load the parameters** from the parameter file in the parameter_files folder.
# + colab={"base_uri": "https://localhost:8080/"} id="4hC6e-jZPA-m" outputId="267d80b0-684a-49fb-c272-787d7dff7d02"
gce.load_params("../parameter_files/parameters.py")
# + [markdown] id="VOpAIsRrdfZB"
# We can take a look at the loaded parameters:
# + colab={"base_uri": "https://localhost:8080/"} id="zfC6OAjPdknM" outputId="dea66d4d-f70e-431b-e3cc-b7ec974f19d6"
gce.print_params()
# + [markdown] id="Z_LGGzbjeKZH"
# The parameters are stored in gce.p and can also be accessed group-wise. For example, the Poissonian (P) and point-source (PS) templates used in this analysis can be viewed with
# + colab={"base_uri": "https://localhost:8080/"} id="LZeVTwPveJjS" outputId="3f467553-faa5-4cb5-b074-a68206dce210"
gce.p.mod
# + [markdown] id="iDqev5FQemIK"
# and the data-related settings (such as the exposure map, the mask for the region of interest, as well as whether the *Fermi* point-spread function at 2 GeV shall be applied) are stored in
# + colab={"base_uri": "https://localhost:8080/"} id="9deORIGwe3G8" outputId="1c8a99fd-1796-43cc-dd12-2cff027c928b"
gce.p.data
# + [markdown] id="rRIVxF75dQJh"
# Now, let's generate some simulated Monte Carlo photon-count maps for each of the templates. The relevant parameters are stored in the field "tt" (training and testing data) - most importantly the priors, as well as the number of maps given by "n_chunk" (each chunk will be saved in a single file) times the number of simulations per chunk.
# + colab={"base_uri": "https://localhost:8080/"} id="j1wpn081dMHD" outputId="bfbdd9c5-1090-4e80-b80b-80d011fbcd87"
gce.p.tt
# + [markdown] id="6esqIeBWhaDe"
# To **generate** the template maps, we can simply run
# + id="kgvxgXvmh-3H" colab={"base_uri": "https://localhost:8080/"} outputId="4fe4ee61-73c5-4f53-ae3d-b1682c1f7e32"
# Ray settings (for parallelized data generation)
# ray_settings = {"num_cpus": 4, "object_store_memory": 2000000000}
ray_settings = {"num_cpus": 4} # select the number of CPUs here
gce.generate_template_maps(ray_settings, n_example_plots=5, job_id=0)
# + [markdown] id="JgSUGOmXJ4Dl"
# Some example maps (whose number is determined by ```n_example_plots``` above) for each template can be viewed in the folder ```GCE_NN/data/Template_maps/Example_128```.
# + [markdown] id="IZy9C4Cez6Pu"
# The next step is to **combine** (i.e. sum up) the individual template maps to obtain the final training, validation, and testing maps. Internally, this is done in two steps: 1) the filenames of the template maps for each of these three subsets are stored in a file, and 2) the template maps are combined and saved.
# + id="jv3KUnRmzynd" colab={"base_uri": "https://localhost:8080/"} outputId="98d2f596-716e-4eb6-ea53-9cc425d3ee53"
gce.combine_template_maps(save_filenames=True, do_combine=True)
# + [markdown] id="UfprvUAl2uKc"
# NOTE: if data has already been generated, the corresponding parameters can be directly loaded from the template maps / combined maps folders, e.g.
#
# ```
# gce.load_params("../data/Template_maps/Test_128")
# gce.load_params("../data/Combined_maps/Test_comb_128")
# ```
#
# + [markdown] id="BFUd4HLV1uUV"
# Next, we need to build the **data processing pipeline** that will feed the combined photon-count maps to the neural network.
# + colab={"base_uri": "https://localhost:8080/"} id="oY_3_oH71mmS" outputId="18a863f3-c50a-4a2d-fa22-500b56d9fbd4"
gce.build_pipeline()
# + [markdown] id="0t-F3xFZMb3V"
# We can use the method ```get_samples()``` to get photon-count maps and their associated labels from the datasets **train** (used for training), **val** (used as an independent validation dataset during training), and **test** (used for testing once the training is finished)
# + colab={"base_uri": "https://localhost:8080/"} id="NCq1omWZ2_5y" outputId="6cded615-f41b-4fb9-ec81-624b587e051f"
samples = gce.datasets["test"].get_samples(1)
data, labels = samples["data"], samples["label"] # samples contains data and labels (flux fractions & SCD histograms)
print("Shapes:")
print(" Data", data.shape) # n_samples x n_pix_in_ROI
print(" Flux fractions", labels[0].shape) # n_samples x n_templates
print(" SCD histograms", labels[1].shape) # n_samples x n_bins x n_PS_templates
# + [markdown] id="JtAvpwmg2FCs"
# Let's take a look at a combined map. The maps are compressed and only contain the pixels that lie within the ROI - the method ```decompress()``` returns the full-sky map that can be fed to the healpy functions.
# + [markdown] id="suiTYvwM32ll"
# We will plot
# 1. the **photon-count map**,
# 2. the rescaled version in **'flux' space** as shown to the neural network (divided by exposure correction), and
# 3. the *Fermi* **exposure correction**.
# + pycharm={"name": "#%%\n"}
# NOTE: the maps are stored in NEST format
map_to_plot = 0
r = gce.p.data["outer_rad"] + 1
hp.cartview(gce.decompress(data[map_to_plot] * gce.template_dict["rescale_compressed"]), nest=True,
title="Simulated data: Count space", lonra=[-r, r], latra=[-r, r])
hp.cartview(gce.decompress(data[map_to_plot]), nest=True,
title="Simulated data: Flux space", lonra=[-r, r], latra=[-r, r])
hp.cartview(gce.decompress(gce.template_dict["rescale_compressed"], fill_value=np.nan), nest=True,
title="Fermi exposure correction", lonra=[-r, r], latra=[-r, r])
plt.show()
# + [markdown] id="sZdi-sqRRkOS"
# Let's also plot the real *Fermi* map in our region of interest. Of course, it looks quite different from our simulated maps because we only included the *Fermi* bubbles, the GCE, and isotropic point sources in this example (so we are completely ignoring the diffuse Galactic foregrounds, which are responsible for the majority of photon counts).
# + colab={"base_uri": "https://localhost:8080/", "height": 375} id="xQb4l2YTQVy1" outputId="49da64e5-eb55-4b7a-d7ef-a6e10778a3f6"
fermi_counts = gce.datasets["test"].get_fermi_counts()
hp.cartview(gce.decompress(fermi_counts * gce.generators["test"].settings_dict["rescale_compressed"]), nest=True,
title="Fermi data: Count space", max=100, lonra=[-r, r], latra=[-r, r])
# hp.cartview(gce.decompress(fermi_counts), nest=True, title="Fermi data: Flux space", max=100)
plt.show()
# + [markdown] id="XSlNFiUVT7m7"
# Now, it's time to **build** our neural network:
#
# + pycharm={"name": "#%%\n"}
gce.build_nn()
# + [markdown] id="cgTUkgH9UG-j"
# *NOTE*: Once the neural network has been trained, **loading** is as easy as ```gce.load_nn()```.
# + id="eg7I_lFQhFek"
# gce.load_nn()
# + [markdown] id="0fpRCAyIUWxm"
# Let's **train** our neural network to predict
# 1. the **flux fractions** of the different templates (using a negative maximum log-likelihood loss function), and
# 2. the **SCD histograms** of the GCE and isotropic point source populations (using the *Earth Mover's pinball loss*, see [arXiv:2106.02051](https://arxiv.org/abs/2106.02051)).
# + colab={"base_uri": "https://localhost:8080/"} id="7GFC2rqsUVa_" outputId="4da7e2e5-d228-4d81-d24f-2f58659e9278"
gce.train_nn("flux_fractions")
# + colab={"base_uri": "https://localhost:8080/"} id="eiDLtxMkUuxB" outputId="172388a8-828b-4a17-99b7-502723681801"
gce.train_nn("histograms")
# + [markdown] id="UBibNbv-U6GA"
# Finally, let's **evaluate** the performance of our neural network on simulated test data.
# + id="r7wRxSZ_VFU_"
n_samples = 20
test_samples = gce.datasets["test"].get_samples(n_samples)
test_data, test_ffs, test_hists = test_samples["data"], test_samples["label"][0], test_samples["label"][1]
tau = np.arange(5, 100, 5) * 0.01 # quantile levels for SCD histograms, from 5% to 95% in steps of 5%
pred = gce.predict(test_data, tau=tau, multiple_taus=True) # get the NN predictions
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="bbWYHid2VUJs" outputId="94fbe736-af3b-402e-c5e0-8643d27be654"
# Make some plots (will be saved in the models folder)
gce.plot_nn_architecture()
gce.plot_flux_fractions(test_ffs, pred)
gce.plot_histograms(test_hists, pred, plot_inds=np.arange(9))
gce.plot_maps(test_data, decompress=True, plot_inds=np.arange(9))
plt.show()
# + [markdown] id="4BnWAJt1Tf0j" pycharm={"name": "#%% md\n"}
# Clearly, the training dataset is too small and the training was too short to obtain accurate and precise predictions. Still, the neural networks have already learned *something*, and the predictions are roughly in the right ballpark.
|
examples/gce_nn_example_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sqlite3
df_bal = pd.read_csv('./data/balance.csv', delimiter=';', dtype='str')
df_2015 = pd.read_csv('./data/2015.csv', dtype='str')
df_2015 = df_2015[df_2015['TRANSACTION ID'].notna()]
df_2016 = pd.read_csv('./data/2016.csv', dtype='str')
df_2016 = df_2016[df_2016['TRANSACTION ID'].notna()]
df_2017 = pd.read_csv('./data/2017.csv', dtype='str')
df_2017 = df_2017[df_2017['TRANSACTION ID'].notna()]
df_trans = pd.concat([df_2015, df_2016, df_2017])
df_trans.columns = [i.strip().lower() for i in df_trans.columns]
df_bal.columns = [i.strip().lower() for i in df_bal.columns]
df_comb = pd.merge(df_trans, df_bal, how='left', on='date')
df_comb.to_csv('./data/combined_transactions.csv')
df_comb['words'] = df_comb['description'].apply(lambda x: x.split())
# +
def sort_categories(desc_words):
if 'Call' in desc_words or 'Put' in desc_words:
return 'option'
elif 'Bought' in desc_words or 'Sold' in desc_words:
return 'stock'
elif 'EXPIRATION' in desc_words or 'EXERCISE' in desc_words:
return 'expiration'
elif 'ASSIGNMENT' in desc_words:
return 'assignment'
elif 'DIVIDEND' in desc_words:
return 'dividend'
elif 'FUNDING' in desc_words:
return 'funds'
elif 'MARGIN' in desc_words:
return 'margin interest'
# elif 'OFF-CYCLE' in desc_words:
# return 'off-cycle interest'
elif 'FUTURES' in desc_words:
return 'futures'
# elif 'MARK' in desc_words:
# return 'mark to market'
# elif 'REBATE' in desc_words:
# return 'rebate'
# elif 'INTERNAL' in desc_words and 'CASH' in desc_words:
# return 'internal cash transfer'
elif 'INTEREST' in desc_words and 'ADJUSTMENT' in desc_words:
return 'interest adjustment'
# elif 'INTERNAL' in desc_words and 'ACCOUNT' in desc_words:
# return 'internal account transfer'
# elif 'MISCELLANEOUS' in desc_words:
# return 'miscellaneous'
else:
return 'miscellaneous'
def get_underlying(symbol):
try:
underlying = symbol.split()[0]
return underlying
except BaseException:
return
# -
df_comb['category'] = df_comb['words'].apply(lambda x: sort_categories(x))
df_comb['underlying'] = df_comb['symbol'].apply(lambda x: get_underlying(x))
df_comb
df_comb.columns
df_comb.to_excel('./data/processed.xlsx', index=False)
df_comb[df_comb['category']=='stocks']
df_comb[((df_comb['category']=='stock') | (df_comb['category']=='option')) & (df_comb['underlying']=='NFLX')]
df_comb[['price', 'commission']].fillna(0)
|
data_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #Numpy Tutorial
# Numpy is a computational library for Python that is optimized for operations on multi-dimensional arrays. In this notebook we will use numpy to work with 1-d arrays (often called vectors) and 2-d arrays (often called matrices).
#
# For a the full user guide and reference for numpy see: http://docs.scipy.org/doc/numpy/
import numpy as np # importing this way allows us to refer to numpy as np
# # Creating Numpy Arrays
# New arrays can be made in several ways. We can take an existing list and convert it to a numpy array:
mylist = [1., 2., 3., 4.]
mynparray = np.array(mylist)
mynparray
# You can initialize an array (of any dimension) of all ones or all zeroes with the ones() and zeros() functions:
one_vector = np.ones(4)
print one_vector # using print removes the array() portion
one2Darray = np.ones((2, 4)) # an 2D array with 2 "rows" and 4 "columns"
print one2Darray
zero_vector = np.zeros(4)
print zero_vector
# You can also initialize an empty array which will be filled with values. This is the fastest way to initialize a fixed-size numpy array however you must ensure that you replace all of the values.
empty_vector = np.empty(5)
print empty_vector
# #Accessing array elements
# Accessing an array is straight forward. For vectors you access the index by referring to it inside square brackets. Recall that indices in Python start with 0.
mynparray[2]
# 2D arrays are accessed similarly by referring to the row and column index separated by a comma:
my_matrix = np.array([[1, 2, 3], [4, 5, 6]])
print my_matrix
print my_matrix[1, 2]
# Sequences of indices can be accessed using ':' for example
print my_matrix[0:2, 2] # recall 0:2 = [0, 1]
print my_matrix[0, 0:3]
# You can also pass a list of indices.
fib_indices = np.array([1, 1, 2, 3])
random_vector = np.random.random(10) # 10 random numbers between 0 and 1
print random_vector
print random_vector[fib_indices]
# You can also use true/false values to select values
my_vector = np.array([1, 2, 3, 4])
select_index = np.array([True, False, True, False])
print my_vector[select_index]
# For 2D arrays you can select specific columns and specific rows. Passing ':' selects all rows/columns
select_cols = np.array([True, False, True]) # 1st and 3rd column
select_rows = np.array([False, True]) # 2nd row
print my_matrix[select_rows, :] # just 2nd row but all columns
print my_matrix[:, select_cols] # all rows and just the 1st and 3rd column
# #Operations on Arrays
# You can use the operations '\*', '\*\*', '\\', '+' and '-' on numpy arrays and they operate elementwise.
my_array = np.array([1., 2., 3., 4.])
print my_array*my_array
print my_array**2
print my_array - np.ones(4)
print my_array + np.ones(4)
print my_array / 3
print my_array / np.array([2., 3., 4., 5.]) # = [1.0/2.0, 2.0/3.0, 3.0/4.0, 4.0/5.0]
# You can compute the sum with np.sum() and the average with np.average()
print np.sum(my_array)
print np.average(my_array)
print np.sum(my_array)/len(my_array)
# #The dot product
# An important mathematical operation in linear algebra is the dot product.
#
# When we compute the dot product between two vectors we are simply multiplying them elementwise and adding them up. In numpy you can do this with np.dot()
array1 = np.array([1., 2., 3., 4.])
array2 = np.array([2., 3., 4., 5.])
print np.dot(array1, array2)
print np.sum(array1*array2)
# Recall that the Euclidean length (or magnitude) of a vector is the squareroot of the sum of the squares of the components. This is just the squareroot of the dot product of the vector with itself:
array1_mag = np.sqrt(np.dot(array1, array1))
print array1_mag
print np.sqrt(np.sum(array1*array1))
# We can also use the dot product when we have a 2D array (or matrix). When you have an vector with the same number of elements as the matrix (2D array) has columns you can right-multiply the matrix by the vector to get another vector with the same number of elements as the matrix has rows. For example this is how you compute the predicted values given a matrix of features and an array of weights.
my_features = np.array([[1., 2.], [3., 4.], [5., 6.], [7., 8.]])
print my_features
my_weights = np.array([0.4, 0.5])
print my_weights
my_predictions = np.dot(my_features, my_weights) # note that the weights are on the right
print my_predictions # which has 4 elements since my_features has 4 rows
# Similarly if you have a vector with the same number of elements as the matrix has *rows* you can left multiply them.
my_matrix = my_features
my_array = np.array([0.3, 0.4, 0.5, 0.6])
print np.dot(my_array, my_matrix) # which has 2 elements because my_matrix has 2 columns
# #Multiplying Matrices
# If we have two 2D arrays (matrices) matrix_1 and matrix_2 where the number of columns of matrix_1 is the same as the number of rows of matrix_2 then we can use np.dot() to perform matrix multiplication.
matrix_1 = np.array([[1., 2., 3.],[4., 5., 6.]])
print matrix_1
matrix_2 = np.array([[1., 2.], [3., 4.], [5., 6.]])
print matrix_2
print np.dot(matrix_1, matrix_2)
|
Studying Materials/Course 2 Regression/Multiple Regression/numpy-tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolutional Neural Network (CNN)
#
# `
# Convolutional Neural Networks (ConvNets or CNNs) are a category of Neural Networks proven effective in image recognition and classification. ConvNets have been successful in identifying faces, objects and traffic signs apart from powering vision in robots and self-driving cars.
# `
#
# More details in the link: https://ujjwalkarn.me/2016/08/11/intuitive-explanation-convnets/
#
# ## Introduction
#
# In this lab, we will build a Convolutional Neural Network to automatically detecting P and S phases in the seismic waveforms. This lab is modified from study entitled ["Generalized Seismic Phase Detection with Deep Learning" by <NAME> et al., 2019](https://arxiv.org/abs/1805.01075)
#
# The training dataset are provided in the Waveform.npy and Label.npy. The waveforms (X) are composed of three components (N,E,Z) with the window length of 4 seconds. The sampling rate is 100 Hz. Therefore, for each training seismgram, there are 400*3 data points. The Labels (Y) distinguish 3 classes (P,S, and Noise windows) with 3 numbers (0,1,2). In order to perform multiple classification by CNN, we need to do one-hot encoding for the labels. The link of why we need one-hot encoding is attached: https://machinelearningmastery.com/why-one-hot-encode-data-in-machine-learning/. By using one-hot encoding we change the labels 0,1,and 2 into [1,0,0],[0,1,0],and[0,0,1]
#
# We then split the training dataset into two parts: one for training, one for testing. We use the testing dataset to select best model. To measure the performance of best trained model, we plot the [confusion matrix](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/#:~:text=A%20confusion%20matrix%20is%20a,related%20terminology%20can%20be%20confusing.), [precision-recall curve](https://scikit-learn.org/stable/auto_examples/model_selection/plot_precision_recall.html) and [ROC curve](https://towardsdatascience.com/understanding-auc-roc-curve-68b2303cc9c5).
#
# #### Notice
#
# If you meet a bug from Keras packages (version problem), please try to change the import source.
# For example, you can switch `from keras.layers import Conv1D` to `from tensorflow.keras.layers import Conv1D`
#
# +
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import scipy.stats as stats
from obspy.signal.trigger import trigger_onset
# sklearn packages
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import confusion_matrix, precision_recall_curve, roc_curve
# keras packages
from keras import backend as K
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.models import Sequential, Model
from keras.layers import Input, Conv1D, MaxPooling1D, UpSampling1D,Flatten,Dense,Dropout,BatchNormalization
from keras.utils import np_utils
from keras.optimizers import Adam
# -
# ## Read Data
#
# Load waveform (X) and label (Y) dataset from Southern California Earthquake Data Center http://scedc.caltech.edu/research-tools/deeplearning.html. The dataset used in this labe includes 10000 samples (1% of total dataset). The following section plot 3 examples of P/S waves and Noise windows. The window length are all 4 seconds with sampling rate of 100 Hz. The P and S wave arrivals occurs at the center of the windows.
#
# In order to perform multiple classification with CNN, we need to do one-hot encoding for the labels [[link]](https://machinelearningmastery.com/why-one-hot-encode-data-in-machine-learning/). By using one-hot encoding we change the labels 0,1,and 2 into [1,0,0],[0,1,0],and[0,0,1] respectively. We use [1,0,0],[0,1,0],and[0,0,1] to represent P phase, noise, and S pahse respectively.
#
# +
X=np.load('Waveform.npy')
Y=np.load('Label.npy')
labels=['P','S','Noise']
# Plot examples of 3 classes
matplotlib.rc('font', **{'size' : 15})
order=[0,2,1]
plt.figure(figsize=(8,8))
for k in range(3):
plt.subplot(3,1,k+1)
for i in range(3):
plt.plot(np.arange(400)*0.01,X[order[k],:,i]+i)
plt.title(labels[k])
plt.yticks([])
if k<2:
plt.xticks([])
plt.show()
# convert integers to dummy variables (one hot encoding)
encoder = LabelEncoder()
encoded_Y = encoder.fit_transform(Y)
en_Y = np_utils.to_categorical(encoded_Y)
# split dataset into training set and validation set
X_train, X_val, y_train, y_val = train_test_split(X, en_Y, test_size=0.33, random_state=42)
# -
# ## Building the Model
#
# Training a convolutional nerual network is similar to training a (fully-connected) nerual network. You can find the definition of loss function, optimizer, activation functions, epoch and batch size in the lab of nerual network.
#
# The largest difference between CNN and NN is that CNN use layers called Conv1D or Conv2D. In our lab, waveforms are time series not a 2D images. So we use the [Conv1D](https://keras.io/api/layers/convolution_layers/convolution1d/). The first argument for Conv1D is the number of filters. It means the dimensionality of the output space (i.e. the number of output filters in the convolution). It must be a integer. The second argument is kernel size. It specifies the length of the 1D convolution window. Another important argument is strides, specifying the stride length of the convolution. It means the downsampling rate, if you set stride equals 2, the output time series would downsample by 2. It has similar effect as [pooling layers](https://keras.io/api/layers/pooling_layers/max_pooling1d/). The first layer is very special, you need to define the input shape (input_shape). In our case the shape of input is 400*3. The window length of a recording of waveform is 4 seconds and the sampling rate is 100 Hz. So we had 400 points for a waveform recording. The number 3 means the number of channels (N,E,Z).
#
# We usually use relu function for the activation functions in the Conv1D and Dense layers, however, for the last layer, we should use softmax. The softmax function takes the output vector, and scales all values such that they sum up to 1. In this way, we get a vector of probabilities. The first entry in the output corresponds to the probability that the input image is a 0, the second entry that the input is 1, etc.:
#
# $$
# P = \left[\begin{matrix} p(0) \\ p(1) \\ p(2) \\ ... \\ p(9) \end{matrix} \right] \quad , \quad \sum_{i=0}^9 P_i = 1
# $$
#
# We now have to choose a loss function. For multi-class classification tasks, _categorical cross-entropy_ is usually a good choice. This loss function is defined as follows:
#
# $$
# \mathcal{L} = - \sum_{c=0}^N y_c \log \left( p_c \right)
# $$
#
# where $y_c$ is the label of class $c$, and $p$ is the predicted probability. Note that $y_c$ is either 0 or 1, and that $0 < p_c < 1$. With our chosen loss function, we are ready for the final assembly of the model.
#
# In addition, we add Dropout. You can learn more about it if you are interested. [Dropout](https://towardsdatascience.com/machine-learning-part-20-dropout-keras-layers-explained-8c9f6dc4c9ab) is a technique used to prevent a model from overfitting. Dropout works by randomly setting the outgoing edges of hidden units (neurons that make up hidden layers) to 0 at each update of the training phase.
#
# We build the model with the following code:
# ```
# model = Sequential()
# model.add(Conv1D(16, 3, activation='relu',strides=2,input_shape=(n_in,3)))
# model.add(Conv1D(32, 3, strides=2,activation='relu'))
# model.add(Conv1D(64, 3, strides=2,activation='relu'))
# model.add(Conv1D(128, 3, strides=2,activation='relu'))
# model.add(Flatten())
# model.add(Dense(128, activation='relu'))
# model.add(Dropout(0.5))
# model.add(Dense(3, activation='softmax'))
# ```
# The model structure is shown below:
#
# 
#
#
# +
# 3 classes
n_in=400
model = Sequential()
# add convolutional layers
model.add(Conv1D(16, 3, activation='relu',strides=2,input_shape=(n_in,3)))
model.add(Conv1D(32, 3, strides=2,activation='relu'))
model.add(Conv1D(64, 3, strides=2,activation='relu'))
model.add(Conv1D(128, 3, strides=2,activation='relu'))
# Flatten before fully connected layers
model.add(Flatten())
model.add(Dense(128, activation='relu'))
# Dropout to prevent a model from overfitting. 0.5 means 50% neurals are deactivated.
model.add(Dropout(0.5))
# Softmax is suitable for multiple classification problem
model.add(Dense(3, activation='softmax'))
model.summary()
adam=Adam(learning_rate=0.0005, beta_1=0.9, beta_2=0.999, amsgrad=False)
model.compile(loss='categorical_crossentropy', optimizer=adam, metrics=['accuracy'])
# Early stop
es = EarlyStopping(monitor='val_accuracy', mode='max', verbose=1, patience=5)
mc = ModelCheckpoint('CNNclassifier.h5', monitor='val_accuracy', mode='max', verbose=0, save_best_only=True)
history=model.fit(X_train, y_train, epochs=100, batch_size=128, validation_data=(X_val, y_val),
callbacks=[es,mc], verbose=0)
# -
# ## Training History
#
# We have recorded the history of training in a variable named 'history'. We wll then visualize the history of the training/testing loss. In addition to loss, we can plot the metrics change with the training epoch. In the following plots, you can see the training loss would be smaller than testing loss after certain epoch. It means the model starts to overfit after that epoch and we should stop training then.
#
# plot metrics
plt.figure(figsize=(7,7))
plt.subplot(211)
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.legend(['train_loss','val_loss'])
plt.subplot(212)
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.legend(['train_accuracy','val_accuracy'])
plt.xlabel('epoch')
scores = model.evaluate(X_val, y_val, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# ## [Plotting Confusion Matrix](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/#:~:text=A%20confusion%20matrix%20is%20a,related%20terminology%20can%20be%20confusing.)
#
# In this section, we would plot the confusion matrix. You could learn more about it through the link
y_pred = model.predict(X_val)
y_val_nonhot=np.round(y_val.argmax(axis=1))
y_pred_nonhot=np.round(y_pred.argmax(axis=1))
cm = confusion_matrix(y_val_nonhot, y_pred_nonhot)
print(cm)
plt.figure(figsize=(6,6))
plt.imshow(cm, interpolation='nearest', cmap='jet')
plt.colorbar()
tick_marks = np.arange(3)
plt.xticks(tick_marks, labels, rotation=45)
plt.yticks(tick_marks, labels)
plt.ylim([2.5,-0.5])
plt.xlim([-0.5,2.5])
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
# ## [Plotting Precision-Recall Curve](https://scikit-learn.org/stable/auto_examples/model_selection/plot_precision_recall.html)
# +
# precision recall curve
plt.figure(figsize=(7,7))
precision = dict()
recall = dict()
for i in range(3):
precision[i], recall[i], _ = precision_recall_curve(y_val[:, i],y_pred[:, i])
plt.plot(recall[i], precision[i], lw=2, label='{}'.format(labels[i]))
plt.xlabel("recall")
plt.ylabel("precision")
plt.legend(loc="best")
plt.title("precision vs. recall curve")
plt.show()
# -
# ## [Plotting ROC Curve](https://towardsdatascience.com/understanding-auc-roc-curve-68b2303cc9c5)
#
# +
# roc curve
plt.figure(figsize=(7,7))
fpr = dict()
tpr = dict()
for i in range(3):
fpr[i], tpr[i], _ = roc_curve(y_val[:, i], y_pred[:, i])
plt.plot(fpr[i], tpr[i], lw=2, label='{}'.format(labels[i]))
plt.xlabel("false positive rate")
plt.ylabel("true positive rate")
plt.legend(loc="best")
plt.title("ROC curve")
plt.show()
# -
# ## Exercise 1
#
# Please finish training the provided model. Then plot loss/accuracy history curve, confusion matrix, precision-recall curve, and ROC curve. Please use plt.savefig to save these figures and include them in the final reports.
#
# ## Exercise 2
#
# Please try 3 additional models and plot corresponding loss/accuracy history curve, confusion matrix, precision-recall curve, and ROC curve.
# Please use plt.savefig to save these figures and include them in the final reports.
# #### Model 1 (Deeper Structure)
# Filter size in all Conv1D layers are 3.
# Strides in all Conv1D layers are 2.
#
# 
#
#
# #### Model 2 (Change Filter Size)
# Filter size in the first Conv1D layer is 7, second layer is 5, and third layer is 3.
# Strides in all Conv1D layers are 2.
#
# 
#
#
# #### Model 3 (Larger Stride)
# Filter size in all Conv1D layers are 3.
# Strides in all Conv1D layers are 3.
#
# 
#
# ## Exercise 3
#
# By testing the performance of previous models, you have experiences on how to build a CNN model. Please experiment with different model configurations (number of layers, number of filters, activation functions, number of epochs, etc.) and then use model.summary() function to print the best configuration you used. Please include the summary information in the final report.
|
PhasePicking.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Science Mathematics
# # Simple Linear Regression
# # In-Class Activity
# Refer to your class handout for background information.
import numpy as np
from scipy import stats
# Let's instantiate the data set.
submarine_sightings = np.array([1,2,3,4,5,6,7,8,9,10])
cyber_activity_metric = np.array([0.021025,0.022103,0.023237,0.024428,0.025681,0.026997,0.028381,0.029836,0.031366,0.032974])
# Now, let's calculate our regression values.
slope, intercept, r_value, p_value, std_err = stats.linregress(submarine_sightings,cyber_activity_metric)
slope, intercept, r_value, p_value, std_err
# Next, print the R^2 value. How good is your fit?
# + tags=[]
print('r-squared:', r_value**2)
# -
# ### Question 1:
# +
'''THis is my answer
I think the data science is cool
'''
# 2:
x = mx+ b
# -
# ***Now save your output. Go to File -> Print Preview and save your final output as a PDF. Turn in to your Instructor, along with any additional sheets.
# ### Question C
#
# *Does a linear relationship exist between these data sets? How do you know?*
#
# Yes, a linear relationship exists; the slope is positive and the standard error is pretty small.
#
# ### Question D
#
# *Based on your calculations, is there a correlative relationship between cyber activity and submarines within the region?*
#
# The correlation is very strong. This can be easily double-checked visually with a quick graph:
import matplotlib.pyplot as plt
plt.plot(submarine_sightings, cyber_activity_metric, 'bo')
# # Question 2
#
# You are developing a convolutional neural network for identifying Chinese
# military images on social media. Your loss function and gradient
# functions are:
#
# f(m,b)=m^2+b^2
#
# ∇f(m,b)=[2m,2b]
#
# Answer the following:
#
# a. Using a learning rate of 0.1, calculate the first 5 gradient
# descent iterations, beginning at (1,5). Did your model converge?
#
#
# + tags=[]
step = 0.1
pos = [1,5,"f(m,b)"] #f(m,b) = m**2 + b**2)
for i in range(5):
pos[0] = round((pos[0] - (step * 2 * pos[0])),2)
pos[1] = round((pos[1] - (step * 2 * pos[1])),2)
pos[2] = round((pos[0]**2 + pos[1]**2),2)
print("Step {}".format(i))
print (pos)
print ("\n")
# -
# It converges, albeit slowly and I think it didn't quite get there
# b. Increase the learning rate to 0.5, and repeat step a. Does your model converge?
# + tags=[]
step = 0.5
pos = [1,5,"f(m,b)"] #f(m,b) = m**2 + b**2)
for i in range(5):
pos[0] = round((pos[0] - (step * 2 * pos[0])),2)
pos[1] = round((pos[1] - (step * 2 * pos[1])),2)
pos[2] = round((pos[0]**2 + pos[1]**2),2)
print("Step {}".format(i))
print (pos)
print ("\n")
# -
# It seems to have overconverged - it went to all zeroes in the first step
# c. Explain the significance of the learning rate. Did it have an impact on the convergence of your model?
# It appears that the learning rate needs to be set at a good rate to properly converge without overshooting. I wonder if you could learn at a medium rate, then if you overshoot, step back and feed that output into another algorithm with a smaller learning rate to dial it in a little better. Or, perhaps, do short runs with various learning rates to estimate which one will converge at a reasonable speed, then feed that into a convergence algorithm.
# For fun, we could graph the convergence process...
# + tags=[]
from mpl_toolkits import mplot3d
import array as arr
step = 0.07
pos = [1,5,"f(m,b)"] #f(m,b) = m**2 + b**2)
xg,yg,zg=[],[],[] # will use these to shove the points into a 3d matplotlib plot
for i in range(25):
pos[0] = round((pos[0] - (step * 2 * pos[0])),2)
pos[1] = round((pos[1] - (step * 2 * pos[1])),2)
pos[2] = round((pos[0]**2 + pos[1]**2),2)
xg.append(pos[0]) # there's probably a more elegant way
yg.append(pos[1]) # but this ensures correct type
zg.append(pos[2])
#print("Step {}: \t{}".format(i,pos))
fig = plt.figure()
ax = plt.axes(projection='3d')
ax.scatter3D(xg,yg,zg, 'ro')
# -
|
Data_Science_Mathematics_Session_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Chapter 10 of [A guided tour of mathematical methods for the physical sciences](http://www.cambridge.org/nz/academic/subjects/physics/mathematical-methods/guided-tour-mathematical-methods-physical-sciences-3rd-edition#KUoGXYx5FTwytcUg.97) explores the Laplacian. In Chapter 5 and 6, you learned about the gradient and the divergence. Now, we are combining this to form the Laplacian operator.
# ### The second derivative of a 1D function
# In the book, we discuss the second derivative to explore the nature of the extrema of functions. Is the second derivative of a function $f(x)$ greater than zero, then $f(x)$ has a minimum. With $\frac{d^2 f}{d x^2}<0$, $f(x)$ has a maximum. It is easy to prove that the shortest distance between two points is a straight line. You probably knew that, but an analysis of the second derivative of the shape of paths between two points, results in $$\frac{d^2 h}{dx^2} = 0.$$
# ### The Laplacian
# In higher dimensions, Chapter 10 shows that the shape $h(x,y)$ of a soap film obeys $$\nabla^2 h(x,y) =0.$$ We already learned about the gradient operator $\nabla$ in Chapter 5, but $\nabla^2 = \nabla\cdot\nabla$: the divergence of the gradient. This differential equation is called the Laplace equation. In some books, you will find the Laplacian operator $\nabla^2$ written as $\Delta$.
#
# If we consider a square frame to blow bubbles, it makes sense to treat the problem in Cartesian coordinates, where
# $$\nabla^2 h(x,y) = \frac{\partial^2 h}{\partial x^2}+\frac{\partial^2 h}{\partial y^2} = 0.$$
#
# This homogeneous second order partial differential equation can be solved by separation of variables and applying the boundary conditions. Here, we'll address a few cases of the shape of a soap film. First, note that solutions where
# $$ \frac{\partial^2 h}{\partial x^2}=\frac{\partial^2 h}{\partial y^2} = 0$$ satisfy the Laplace equation. The soap film can have a shape where both the x- and y-dependence is described by constant and/or linear terms. First, we load some of the tools that we'll use:
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
import numpy as np
# %matplotlib notebook
# We define a grid:
x = np.linspace(-1, 1, num=10)
y = np.linspace(-1, 1, num=10)
X, Y = np.meshgrid(x, y)
# ### The shape of a soap film in a frame with straight sides
# A shape that's a solution to the Laplace equation is $h(x,y) = x + y$. Try inserting it into the Laplace equation!
def h(x,y):
return x+y
# Create a 3D figure of the film in the frame:
# +
fig = plt.figure()
ax = fig.gca(projection='3d')
# Compute and plot the shape of the soap film:
Z = h(X,Y)
ax.plot_surface(X, Y, Z, cmap=cm.coolwarm)
# Plot the wire frame:
maxx = np.ones_like(x)
ax.plot(x, maxx, h(x,maxx),'k',linewidth=2)
minx = -1*np.ones_like(x)
ax.plot(x, minx, h(x,minx),'k',linewidth=2)
maxy = np.ones_like(y)
ax.plot(maxy, y, h(maxy,y),'k',linewidth=2)
miny = -np.ones_like(y)
ax.plot(miny, y, h(miny,y),'k',linewidth=2)
# And for fun, plot a horizontal stick to hold the soap bubble frame:
ax.plot([-1,-2], [-1,-2],[h(-1,-1),h(-1,-1)],'k',linewidth=2)
plt.show()
# -
# You can rotate and zoom the figure above to get a good idea of the soap film shape.
#
# ### A soap film with a saddle shape
# In the previous example, the math (the Laplace equation) tells us the soap film forms a flat plane in that particular flat frame (but we have not included gravity as a force on the soap film, or you blowing in the frame)
#
# Maybe not a very exciting or unexpected result? What if we used a different shaped frame? A function to compute the soap film height $h(x,y) = x*y$. This too, you can confirm is a solution to the Laplace equation. Compute the shape of this soap film.
def h(x,y):
return x*y
Z = h(X,Y)
# ### And plot this frame and film:
# +
fig = plt.figure()
ax = fig.gca(projection='3d')
ax.plot_surface(X, Y, Z, cmap=cm.coolwarm)
#Plot the wire frame:
maxx = np.ones_like(x)
ax.plot(x, maxx, h(x,maxx),'k',linewidth=2)
minx = -1*np.ones_like(x)
ax.plot(x, minx, h(x,minx),'k',linewidth=2)
maxy = np.ones_like(y)
ax.plot(maxy, y, h(maxy,y),'k',linewidth=2)
miny = -np.ones_like(y)
ax.plot(miny, y, h(miny,y),'k',linewidth=2)
#Plot a "stick" to hold the soap bubble frame:
ax.plot([-1,-2], [-1,-2],[h(-1,-1),h(-1,-1)],'k',linewidth=2)
plt.show()
# -
# This frame still has straight sides, but if you rotate (drag and try it!) the graph, you will see the film has a saddle shape. This makes sense, because for the Laplace equation to hold, when the x-dependence has a maximum, the y-dependence is at a minimum, so the second partial derivatives add to zero: $$\nabla^2 h(x,y) = \frac{\partial^2 h}{\partial x^2}+\frac{\partial^2 h}{\partial y^2} = 0.$$
# ### Homework: Try your own frame shape!
# The Laplace equation and its inhomogeneous cousin, the Poisson equation, are used all throughout physics. The rest of Chapter 10 shows you how to derive the Laplace equation in cylindrical and spherical coordinates. The latter will prove handy -- for example --- when we tackle the gravitational field of the Earth, or the electric field of a charged sphere.
#
# With the codes here, and what you know about solutions to the Laplace equation so far, you can now plot the soap film shape for different solutions. The solution $h(x,y) = x^2 - y^2$, for example. After you tried that, let us return to the [overview of jupyter notebooks](https://pal.blogs.auckland.ac.nz/2017/12/02/jupyter-notebooks-for-mathematical-methods-in-the-physical-sciences/)!
|
10_Laplacian.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Contents
# ========
# - [Introduction](#Introduction)
# - [Block Using the Sorted Neighborhood Blocker](#Block-Using-the-Sorted-Neighborhood-Blocker)
# - [Block Tables to Produce a Candidate Set of Tuple Pairs](#Block-Tables-to-Produce-a-Candidate-Set-of-Tuple-Pairs)
# - [Handling Missing Values](#Handling-Missing-Values)
# - [Window Size](#Window-Size)
# - [Stable Sort Order](#Stable-Sort-Order)
# - [Sorted Neighborhood Blocker Limitations](#Sorted-Neighborhood-Blocker-limitations)
# # Introduction
# <font color='red'>WARNING: The sorted neighborhood blocker is still experimental and has not been fully tested yet. Use this blocker at your own risk.</font>
#
# Blocking is typically done to reduce the number of tuple pairs considered for matching. There are several blocking methods proposed. The *py_entitymatching* package supports a subset of such blocking methods (#ref to what is supported). One such supported blocker is the sorted neighborhood blocker. This IPython notebook illustrates how to perform blocking using the sorted neighborhood blocker.
#
# Note, often the sorted neighborhood blocking technique is used on a single table. In this case we have implemented sorted neighborhood blocking between two tables. We first enrich the tables with whether the table is the left table, or right table. Then we merge the tables. At this point we perform sorted neighborhood blocking, which is to pass a sliding window of `window_size` (default 2) across the merged dataset. Within the sliding window all tuple pairs that have one tuple from the left table and one tuple from the right table are returned.
# First, we need to import *py_entitymatching* package and other libraries as follows:
# + nbpresent={"id": "9a89351b-e44f-47ad-afff-b148744173af"}
# Import py_entitymatching package
import py_entitymatching as em
import os
import pandas as pd
# + [markdown] nbpresent={"id": "0f59e4ac-032a-4d59-9172-8ee653831acb"}
# Then, read the input tablse from the datasets directory
# + nbpresent={"id": "2401accd-3160-4b07-aed4-de2f9a4dea35"}
# Get the datasets directory
datasets_dir = em.get_install_path() + os.sep + 'datasets'
# Get the paths of the input tables
path_A = datasets_dir + os.sep + 'person_table_A.csv'
path_B = datasets_dir + os.sep + 'person_table_B.csv'
# + nbpresent={"id": "e51b3877-75c8-431d-bdbd-77362bbf2191"}
# Read the CSV files and set 'ID' as the key attribute
A = em.read_csv_metadata(path_A, key='ID')
B = em.read_csv_metadata(path_B, key='ID')
# + nbpresent={"id": "ac2eb60b-bf26-4a6a-a453-120cc7f660c4"}
A.head()
# + nbpresent={"id": "afadc046-692d-42ad-9c72-523493597682"}
B.head()
# + [markdown] nbpresent={"id": "ca7f0c34-9c21-4b6e-8010-eda1030df041"}
# # Block Using the Sorted Neighborhood Blocker
#
# Once the tables are read, we can do blocking using sorted neighborhood blocker.
# + [markdown] nbpresent={"id": "45585660-2ba9-4211-adee-ace14fe8745f"}
# With the sorted neighborhood blocker, you can only block between two tables to produce a candidate set of tuple pairs.
# + [markdown] nbpresent={"id": "726bd6c9-23a5-4543-a201-f84864433f20"}
# ## Block Tables to Produce a Candidate Set of Tuple Pairs
# + nbpresent={"id": "d2001a06-fe74-4ebd-896c-992803828753"}
# Instantiate attribute equivalence blocker object
sn = em.SortedNeighborhoodBlocker()
# + [markdown] nbpresent={"id": "06bcba06-ff85-43b7-bb04-ce1566a29dd9"}
# For the given two tables, we will assume that two persons with different `zipcode` values do not refer to the same real world person. So, we apply attribute equivalence blocking on `zipcode`. That is, we block all the tuple pairs that have different zipcodes.
# + nbpresent={"id": "2cdc68f4-5874-43d1-a378-b0bd31552ef8"}
# Use block_tables to apply blocking over two input tables.
C1 = sn.block_tables(A, B,
l_block_attr='birth_year', r_block_attr='birth_year',
l_output_attrs=['name', 'birth_year', 'zipcode'],
r_output_attrs=['name', 'birth_year', 'zipcode'],
l_output_prefix='l_', r_output_prefix='r_', window_size=3)
# + nbpresent={"id": "7b4967f5-2f99-4394-bfe9-29ff15334d39"}
# Display the candidate set of tuple pairs
C1.head()
# + [markdown] nbpresent={"id": "386dbb7d-085e-4946-b376-93e686eb62f5"}
# Note that the tuple pairs in the candidate set have the same zipcode.
#
# The attributes included in the candidate set are based on l_output_attrs and r_output_attrs mentioned in block_tables command (the key columns are included by default). Specifically, the list of attributes mentioned in l_output_attrs are picked from table A and the list of attributes mentioned in r_output_attrs are picked from table B. The attributes in the candidate set are prefixed based on l_output_prefix and r_ouptut_prefix parameter values mentioned in block_tables command.
# + nbpresent={"id": "fa6af6e5-471b-4296-98f4-1f4d4ee2869a"}
# Show the metadata of C1
em.show_properties(C1)
# + nbpresent={"id": "6ec70bd1-adea-40af-9f30-304f6236c5ca"}
id(A), id(B)
# + [markdown] nbpresent={"id": "51679b6c-2667-4ed9-88aa-caff4c81edbf"}
# Note that the metadata of C1 includes key, foreign key to the left and right tables (i.e A and B) and pointers to left and right tables.
# -
# ### Handling Missing Values
# If the input tuples have missing values in the blocking attribute, then they are ignored by default. This is because, including all possible tuple pairs with missing values can significantly increase the size of the candidate set. But if you want to include them, then you can set `allow_missing` paramater to be True.
# Introduce some missing values
A1 = em.read_csv_metadata(path_A, key='ID')
A1.ix[0, 'zipcode'] = pd.np.NaN
A1.ix[0, 'birth_year'] = pd.np.NaN
A1
# Use block_tables to apply blocking over two input tables.
C2 = sn.block_tables(A1, B,
l_block_attr='zipcode', r_block_attr='zipcode',
l_output_attrs=['name', 'birth_year', 'zipcode'],
r_output_attrs=['name', 'birth_year', 'zipcode'],
l_output_prefix='l_', r_output_prefix='r_',
allow_missing=True) # setting allow_missing parameter to True
len(C1), len(C2)
C2
# The candidate set C2 includes all possible tuple pairs with missing values.
# ## Window Size
#
# A tunable parameter to the Sorted Neighborhood Blocker is the Window size. To perform the same result as above with a larger window size is via the `window_size` argument. Note that it has more results than C1.
C3 = sn.block_tables(A, B,
l_block_attr='birth_year', r_block_attr='birth_year',
l_output_attrs=['name', 'birth_year', 'zipcode'],
r_output_attrs=['name', 'birth_year', 'zipcode'],
l_output_prefix='l_', r_output_prefix='r_', window_size=5)
len(C1)
len(C3)
# ## Stable Sort Order
#
# One final challenge for the Sorted Neighborhood Blocker is making the sort order stable. If the column being sorted on has multiple identical keys, and those keys are longer than the window size, then different results may occur between runs. To always guarantee the same results for every run, make sure to make the sorting column unique. One method to do so is to append the id of the tuple onto the end of the sorting column. Here is an example.
A["birth_year_plus_id"]=A["birth_year"].map(str)+'-'+A["ID"].map(str)
B["birth_year_plus_id"]=B["birth_year"].map(str)+'-'+A["ID"].map(str)
C3 = sn.block_tables(A, B,
l_block_attr='birth_year_plus_id', r_block_attr='birth_year_plus_id',
l_output_attrs=['name', 'birth_year_plus_id', 'birth_year', 'zipcode'],
r_output_attrs=['name', 'birth_year_plus_id', 'birth_year', 'zipcode'],
l_output_prefix='l_', r_output_prefix='r_', window_size=5)
C3.head()
# # Sorted Neighborhood Blocker limitations
#
# Since the sorted neighborhood blocker requires position in sorted order, unlike other blockers, blocking on a candidate set or checking two tuples is not applicable. Attempts to call `block_candset` or `block_tuples` will raise an assertion.
|
notebooks/guides/step_wise_em_guides/Performing Blocking Using Built-In Blockers (Sorted Neighborhood Blocker).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="snYKsZJ1Atdn"
import numpy as np
import matplotlib.pyplot as plt
import scipy.integrate as integrate
import scipy.special as special
from google.colab import files
# + id="95Yj-oufAxra"
class Simple_Dist:
# param = [(x,density)]
# constructed left-to-right on [init[0][0],end]
# dist pairs stored as inclusive left end
def __init__(self, init, end):
self.param = init
self.start = init[0][0]
self.end = end
def display(self):
points = []
a_param = self.param + [(self.end,self.param[-1][1])]
for i, v in enumerate(a_param[:-1]):
points.append(v)
points.append((a_param[i+1][0],v[1]))
x,y = zip(*points)
plt.plot(x,y)
plt.title("Median Density")
plt.ylabel("Density")
plt.xlabel("Value")
def cdf(self,threshold):
basis = []
val = []
a_param = self.param + [(self.end,self.param[-1][1])]
for i, v in enumerate(a_param):
if v[0] < threshold:
basis.append(v[0])
val.append(v[1])
else:
basis.append(threshold)
cdf = 0
for i, d in enumerate(val):
cdf += d*(basis[i+1]-basis[i])
return cdf
def invcdf(self, prob):
a_param = self.param + [(self.end,self.param[-1][1])]
cdf = 0
for i, v in enumerate(a_param[:-1]):
if cdf+v[1]*(a_param[i+1][0]-a_param[i][0])>prob:
exceed = cdf+v[1]*(a_param[i+1][0]-a_param[i][0])-prob
return a_param[i+1][0]-exceed/v[1]
else:
cdf += v[1]*(a_param[i+1][0]-a_param[i][0])
return self.end
def median(self):
return self.invcdf(0.5)
# Vote indicates
def update_dist_interval(self, left, right, vote, alpha = 0.1):
temp = self.param+[(self.end,self.param[-1][1])]
# Insert new data(s)
# index1 = next(i for i,v in enumerate(self.param+[(self.end,self.param[-1][1])]) if right <= v[0])
index1 = 0
for i in range(len(temp)):
if temp[i][0] > right:
break
index1 += 1
if index1 < len(self.param) and self.param[index1][0] != right:
self.param.insert(index1,(right,self.param[index1-1][1]))
elif index1 >= len(self.param) and right != self.end:
self.param.insert(index1,(right,self.param[index1-1][1]))
index2 = 0
for i in range(len(temp)):
if temp[i][0] >= left:
break
index2 += 1
if index2 < len(self.param) and self.param[index2][0] != left:
self.param.insert(index2,(left,self.param[index2-1][1]))
elif index2 >= len(self.param):
self.param.insert(index2,(left,self.param[index2-1][1]))
# print('start')
# for p in self.param:
# print(p)
# print('end')
# Modify distribution
if vote == -1:
# r_mass = 1 - self.cdf(right)
for i, v in enumerate(self.param):
# print(v)
if left <= v[0] < right:
# print('increase')
self.param[i] = (v[0],v[1] * alpha)
else:
self.param[i] = (v[0],v[1] * (1 - alpha))
# print('decrease')
norm = self.cdf(self.end)
for i, v in enumerate(self.param):
self.param[i] = (v[0], v[1]/norm)
else:
# l_mass = self.cdf(right)
for i, v in enumerate(self.param):
# print(v)
if left <= v[0] < right:
# print('increase')
self.param[i] = (v[0],v[1] * (1 - alpha))
else:
self.param[i] = (v[0],v[1] * alpha)
# print('decrease')
norm = self.cdf(self.end)
for i, v in enumerate(self.param):
self.param[i] = (v[0], v[1]/norm)
# print('end')
def update_dist(self, threshold, vote, alpha = 0.1):
# Insert new data
index = next(i for i,v in enumerate(self.param+[(self.end,self.param[-1][1])]) if threshold < v[0])
if index < len(self.param) and self.param[index][0] != threshold:
self.param.insert(index,(threshold,self.param[index-1][1]))
elif index >= len(self.param):
self.param.insert(index,(threshold,self.param[index-1][1]))
# Modify distribution
if vote == -1:
r_mass = 1 - self.cdf(threshold)
for i, v in enumerate(self.param):
if v[0] < threshold:
self.param[i] = (v[0],v[1] * (1 - alpha))
else:
self.param[i] = (v[0],v[1] * alpha)
norm = self.cdf(self.end)
for i, v in enumerate(self.param):
self.param[i] = (v[0], v[1]/norm)
else:
l_mass = self.cdf(threshold)
for i, v in enumerate(self.param):
if v[0] >= threshold:
self.param[i] = (v[0],v[1] * (1 - alpha))
else:
self.param[i] = (v[0],v[1] * alpha)
norm = self.cdf(self.end)
for i, v in enumerate(self.param):
self.param[i] = (v[0], v[1]/norm)
# + colab={"base_uri": "https://localhost:8080/"} id="mFsSV9gNA508" outputId="6a98e463-7794-49bd-f73c-2cd7f6212d1b"
# true metric
m_star = (-0.94,-0.34) #200 degree
m_star = (0.98,0.17) #11.47 degree
m_star = m_star/np.linalg.norm(m_star) # normalize
# obtain true theta
t_true = np.arccos(m_star[0])
if t_true > np.pi/2: # make sure theta either in [0, pi/2] or [pi, 3/2*pi]
t_true = np.pi*2-t_true
print(t_true)
def eta(x):
return 1/(1+np.exp(5*x))
zeta = 0.5 # f_X /sim U(-1,1)
# implementation of proposition 1
# return a classifier with t
def h_bar(t):
m11, m00 = np.cos(t), np.sin(t)
def hb(x):
if m11+m00 >= 0:
return int(eta(x)>=m00/(m11+m00))
else:
return int(eta(x)<=m00/(m11+m00))
return hb
# confusion matrix, analytical solution
def C11(t): # P(Y=1, h=1)
m11,m00 = np.cos(t), np.sin(t)
x_prime = 0.
h = h_bar(t)
if m00 == 0:
x_prime = 1
elif m11/m00 <= 0:
x_prime = -1
else:
x_prime = np.log(m11/m00)/5
if x_prime > 1:
x_prime = 1
elif x_prime < -1:
x_prime = -1
if m00+m11 >= 0:
return (x_prime-0.2*np.log(1+np.exp(5*x_prime))+1+0.2*np.log(1+np.exp(-5))) # x-0.2ln(1+e^5x)
else:
return (1-0.2*np.log(1+np.exp(5))-x_prime+0.2*np.log(1+np.exp(5*x_prime)))
def C10(t): # P(Y=0, h=1)
return 1-C11(t)
def C01(t): # P(Y=1, h=0)
return 1-C00(t)
def C00(t): # P(Y=0, h=0)
m11,m00 = np.cos(t), np.sin(t)
x_prime = 0
h = h_bar(t)
if m00 == 0:
x_prime = 1
elif (m00+m11)/m00-1 <= 0:
x_prime = -1
else:
x_prime = np.log(m11/m00)/5
if x_prime > 1:
x_prime = 1
elif x_prime < -1:
x_prime = -1
if m00+m11 >= 0:
return (0.2*np.log(1+np.exp(5))-0.2*np.log(1+np.exp(5*x_prime))) # ln(1+e^5x)
else:
return (0.2*np.log(1+np.exp(5*x_prime))-0.2*np.log(1+np.exp(-5)))
# metric evaluation
def phi(t):
m11, m00 = np.cos(t_true), np.sin(t_true)
return m11*C11(t)+m00*C00(t)
# return t*(1-t)+0.6*t
# query function (always maximize phi function)
# alpha: error rate —— with probability alpha, the oracle will return wrong answer
def query(t_1, t_2, alpha):
if phi(t_1) < phi(t_2):
if np.random.rand() > alpha:
return 1 # prefer t2
else:
return 0
else:
if np.random.rand() > alpha:
return 0 # prefer t1
else:
return 1
# + id="SATUNFD4A9Kg"
# implements algorithm 1
# analytical version
# alpha: error rate of oracle
def max_quasiconcave_metric(eps, alpha, iteration):
t_a = 0
t_b = np.pi/2
dist = Simple_Dist(init=[(t_a,1/(t_b-t_a))],end=t_b)
m_bar = np.zeros(2)
C_bar = 0
for iter in range(iteration):
x1 = dist.invcdf(1/3)
x2 = dist.invcdf(2/3)
if query(x1,x2,alpha):
dist.update_dist_interval(left=x1, right=t_b, vote=1, alpha=alpha)
# dist.update_dist(x1,1, alpha = alpha)
else:
dist.update_dist_interval(left=t_a, right=x2, vote=1, alpha=alpha)
# dist.update_dist(x2,-1, alpha = alpha)
# dist.display()
ct = dist.median()
C_ct = np.array([[C00(ct), C01(ct)],[C10(ct), C11(ct)]])
m_bar[0], m_bar[1] = np.cos(ct), np.sin(ct)
C_bar = C_ct
del dist
return m_bar,C_bar
# + colab={"base_uri": "https://localhost:8080/"} id="l0JKlKVQBA0F" outputId="d665f155-0669-414f-c689-e6b9e16acbfe"
m,C = max_quasiconcave_metric(1e-4, 0.1, 100)
print("true metric: "+str(m_star))
print("elicited metric: "+str(m))
print("confusion matrix: \n"+str(C))
# + [markdown] id="LA_r0kD7BJA4"
# # Visualization of $\phi$
# + colab={"base_uri": "https://localhost:8080/", "height": 365} id="f6CEUjQKBDcB" outputId="e1f06420-1120-4038-e1b3-f6960727e1d0"
# Plot phi function versus different thetas
ph = []
for i in np.arange(0, np.pi*1.5, np.pi/36):
ph.append(phi(i))
plt.figure(figsize=(16,5))
plt.plot(np.arange(0, np.pi*1.5, np.pi/36), ph, "^-")
for p in np.arange(0.5, 1.5, 0.5):
plt.axvline(x=np.pi*p, c='r', ls='--', alpha=0.7)
plt.axvline(x=t_true, c='g')
t_elicited = np.arccos(m[0])
if t_elicited > np.pi/2: # make sure theta either in [0, pi/2] or [pi, 3/2*pi]
t_elicited = np.pi*2-t_elicited
plt.axvline(x=t_elicited, c='b')
# plt.axvline(x=np.arccos(np.pi/4), c='black')
plt.xticks(np.arange(0, np.pi*1.5, np.pi/36), rotation=60, size="small")
plt.title("phi change with theta")
plt.xlabel("theta/radian")
plt.ylabel("phi")
plt.show()
# + id="QombDkMrBIE1"
num_itr = 1000
step = 0.01
re_arr = np.zeros((50,num_itr))
for itr in range(num_itr):
t_true = np.random.rand()*np.pi/2
m_star = (np.cos(t_true),np.sin(t_true))
# print(itr)
for lp, al in enumerate(np.arange(0., .5, step)):
m,C = max_quasiconcave_metric(1e-4, al, 100)
# re.append(abs(phi2(m[0], m[1])-phi2(m_star[0], m_star[1])))
t = np.arccos(m[0])
if t > np.pi/2:
t = np.pi*2-t
# re_arr[lp][itr] = np.linalg.norm(t-t_true)
re_arr[lp][itr] = np.linalg.norm(m-m_star)
re_max = np.amax(re_arr, 1)
re_min = np.amin(re_arr, 1)
re_mean = np.mean(re_arr, 1)
re_median = np.median(re_arr, 1)
re_std = np.std(re_arr, 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="e8lcwoy2C6wM" outputId="a20d6a19-8edd-44f9-d2e1-07d98da46702"
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(np.arange(0., 1., step), re_max, "-", color='b', alpha=0.2) # plot maximum deviation
# ax.plot(np.arange(0., 1., step), re_min, "-", color='b', alpha=0.2) # plot minimum deviation
ax.plot(np.arange(0., .5, step), re_mean, "^-", color='b', alpha=1, label='mean') # plot mean relative error
ax.errorbar(np.arange(0., .5, step), re_mean, yerr=re_std, capsize=4)
# cm = plt.get_cmap('viridis')
# i=0
# up_prev = re_median
# low_prev = re_median
# for i,pct in enumerate(range(40, 0, -10)):
# re_up = np.percentile(re_arr, 100-pct, 1)
# re_low = np.percentile(re_arr, pct, 1)
# ax.fill_between(np.arange(0., .5, step), re_up, up_prev, color=cm.colors[i*60], alpha=pct/100+0.3, label=str(100-2*pct)+'% interval')
# ax.fill_between(np.arange(0., .5, step), re_low, low_prev, color=cm.colors[i*60], alpha=pct/100+0.3)
# up_prev = re_up
# low_prev = re_low
# ax.fill_between(np.arange(0., .5, step), re_max, up_prev, color=cm.colors[(i+1)*60], alpha=0.3, label='100% interval')
# ax.fill_between(np.arange(0., .5, step), re_min, low_prev, color=cm.colors[(i+1)*60], alpha=0.3)
# plt.yticks(np.arange(0, 1, step=0.1))
plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.title("distance from metric vs alpha")
plt.xlabel("alpha")
plt.ylabel("2-norm distance from true metric")
plt.legend(loc='upper left')
plt.grid()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 415} id="fODZrCob0Zd9" outputId="454fd251-a3ea-456c-a574-0faac15cbed6"
re_hoeffding = np.sqrt(-np.log(0.975)/(2*1000))
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(np.arange(0., 1., step), re_max, "-", color='b', alpha=0.2) # plot maximum deviation
# ax.plot(np.arange(0., 1., step), re_min, "-", color='b', alpha=0.2) # plot minimum deviation
ax.plot(np.arange(0., .5, step), re_mean, "-", color='b', alpha=1, label='mean') # plot mean relative error
ax.errorbar(np.arange(0., .5, step), re_mean, yerr=re_hoeffding, capsize=4)
# cm = plt.get_cmap('viridis')
# i=0
# up_prev = re_median
# low_prev = re_median
# for i,pct in enumerate(range(40, 0, -10)):
# re_up = np.percentile(re_arr, 100-pct, 1)
# re_low = np.percentile(re_arr, pct, 1)
# ax.fill_between(np.arange(0., .5, step), re_up, up_prev, color=cm.colors[i*60], alpha=pct/100+0.3, label=str(100-2*pct)+'% interval')
# ax.fill_between(np.arange(0., .5, step), re_low, low_prev, color=cm.colors[i*60], alpha=pct/100+0.3)
# up_prev = re_up
# low_prev = re_low
# ax.fill_between(np.arange(0., .5, step), re_max, up_prev, color=cm.colors[(i+1)*60], alpha=0.3, label='100% interval')
# ax.fill_between(np.arange(0., .5, step), re_min, low_prev, color=cm.colors[(i+1)*60], alpha=0.3)
# plt.yticks(np.arange(0, 1, step=0.1))
plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.title("Distance From Metric vs alpha")
plt.xlabel("alpha")
plt.ylabel("2-norm distance from true metric")
plt.legend(loc='upper left')
plt.grid()
plt.show()
# + [markdown] id="ZRQwwsYqMhgU"
# ## Non-PTA comparison
# + id="0YZdVo5RIbnf"
# implements algorithm 1
# analytical version
# alpha: error rate of oracle
def max_quasiconcave_metric_nPTA(eps, alpha, iteration):
t_a = 0
t_b = np.pi/2
m_bar = np.zeros(2)
C_bar = 0
iter = 0
# while np.linalg.norm(t_a-t_b) > eps:
for iter in range(iteration):
# divide the searching range into equally seperated intervals
t_c = (3*t_a+t_b)/4
t_d = (t_a+t_b)/2
t_e = (t_a+3*t_b)/4
# compute Confusion Matrices
C_a = np.array([[C00(t_a), C01(t_a)],[C10(t_a), C11(t_a)]])
C_b = np.array([[C00(t_b), C01(t_b)],[C10(t_b), C11(t_b)]])
C_c = np.array([[C00(t_c), C01(t_c)],[C10(t_c), C11(t_c)]])
C_d = np.array([[C00(t_d), C01(t_d)],[C10(t_d), C11(t_d)]])
C_e = np.array([[C00(t_e), C01(t_e)],[C10(t_e), C11(t_e)]])
# pairwise comparisons
ca = query(t_c, t_a, alpha)
dc = query(t_d, t_c, alpha)
ed = query(t_e, t_d, alpha)
be = query(t_b, t_e, alpha)
# sanity check for out-of-order responses
if ca and not dc:
ca = not ca
if dc and not ed:
dc = not dc
if ed and not be:
ed = not ed
# determine the next iter search range based on oracle resposne to query
if ca:
t_b = t_d
elif not ca and dc:
t_b = t_d
elif not dc and ed:
t_a = t_c
t_b = t_e
elif not ed and be:
t_a = t_d
else:
t_a = t_d
m_bar[0], m_bar[1] = np.cos(t_d), np.sin(t_d)
C_bar = C_d
iter += 1
# print("iteration run:"+str(iter))
return m_bar,C_bar
# + id="Wuk8CaoRMiVw"
num_itr = 1000
step = 0.01
re_arr_nPTA = np.zeros((50,num_itr))
for itr in range(num_itr):
t_true = np.random.rand()*np.pi/2
m_star = (np.cos(t_true),np.sin(t_true))
for lp, al in enumerate(np.arange(0., .5, step)):
m,C = max_quasiconcave_metric_nPTA(1e-4, al, 25)
# re.append(abs(phi2(m[0], m[1])-phi2(m_star[0], m_star[1])))
t = np.arccos(m[0])
if t > np.pi/2:
t = np.pi*2-t
# re_arr_nPTA[lp][itr] = np.linalg.norm(t-t_true)
re_arr_nPTA[lp][itr] = np.linalg.norm(m-m_star)
re_max_nPTA = np.amax(re_arr_nPTA, 1)
re_min_nPTA = np.amin(re_arr_nPTA, 1)
re_mean_nPTA = np.mean(re_arr_nPTA, 1)
re_median_nPTA = np.median(re_arr_nPTA, 1)
re_std_nPTA = np.std(re_arr_nPTA, 1)
# + colab={"base_uri": "https://localhost:8080/", "height": 432} id="P9fcmeG2MvGv" outputId="1bf786ba-47a5-4444-d5d0-9d874b290f7f"
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(np.arange(0., .5, step), re_max, "-", color='b', alpha=0.2) # plot maximum deviation
# ax.plot(np.arange(0., .5, step), re_min, "-", color='b', alpha=0.2) # plot minimum deviation
# ax.plot(np.arange(0., .5, step), re_mean_nPTA, "-", alpha=1, label='non-PTA mean',color='y') # plot mean relative error
# ax.plot(np.arange(0., .5, step), re_mean, "-", alpha=1, label='PTA mean',color='g') # plot mean relative error
ax.errorbar(np.arange(0., .5, step), re_mean_nPTA, yerr=re_std_nPTA, capsize=4,label='non-PTA mean')
ax.errorbar(np.arange(0., .5, step), re_mean, yerr=re_std, capsize=4,label='PTA mean')
# plt.yticks(np.arange(0, 1, step=0.1))
plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.title("Distance From Metric vs alpha")
plt.grid()
plt.xlabel("alpha")
plt.ylabel("2-norm distance from true metric")
plt.legend(loc='upper left')
plt.show()
plt.savefig('Loss vs alpha (std)')
# + colab={"base_uri": "https://localhost:8080/", "height": 432} id="suIDyT4h4KWo" outputId="892fd238-2271-4857-f925-ea92c2593765"
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(np.arange(0., .5, step), re_max, "-", color='b', alpha=0.2) # plot maximum deviation
# ax.plot(np.arange(0., .5, step), re_min, "-", color='b', alpha=0.2) # plot minimum deviation
# ax.plot(np.arange(0., .5, step), re_mean_nPTA, "-", alpha=1, label='non-PTA mean') # plot mean relative error
# ax.plot(np.arange(0., .5, step), re_mean, "-", alpha=1, label='PTA mean') # plot mean relative error
ax.errorbar(np.arange(0., .5, step), re_mean_nPTA, yerr=re_hoeffding, capsize=4,label='non-PTA mean')
ax.errorbar(np.arange(0., .5, step), re_mean, yerr=re_hoeffding, capsize=4,label='PTA mean')
# plt.yticks(np.arange(0, 1, step=0.1))
plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.grid()
plt.title("Distance From etric vs alpha")
plt.xlabel("alpha")
plt.ylabel("2-norm distance from true metric")
plt.legend(loc='upper left')
plt.show()
plt.savefig('Loss vs alpha (hoeffding)')
# + [markdown] id="YTVGHzd1J3ov"
# ## Loss vs num_iter
# + id="AxKneGLRPnvX"
step = 0.01
num_itr = 1000
re_arr_iters = np.zeros((4, 19, num_itr))
for i, al in enumerate([0, 0.12, 0.25, 0.5]):
for j, iteration in enumerate(np.append(np.arange(1,10, 1), np.arange(10, 110, 10))):
for itr in range(num_itr):
t_true = np.random.rand()*np.pi/2
m_star = (np.cos(t_true),np.sin(t_true))
# print(itr)
m,C = max_quasiconcave_metric(1e-4, al, iteration)
# re.append(abs(phi2(m[0], m[1])-phi2(m_star[0], m_star[1])))
t = np.arccos(m[0])
if t > np.pi/2:
t = np.pi*2-t
re_arr_iters[i][j][itr] = np.linalg.norm(m-m_star)
re_max = np.amax(re_arr_iters, 2)
re_min = np.amin(re_arr_iters, 2)
re_mean = np.mean(re_arr_iters, 2)
re_median = np.median(re_arr_iters, 2)
re_std = np.std(re_arr_iters, 2)
# + id="jCxAbQzYkr3W"
num_itr = 1000
step = 0.01
re_arr_nPTA_iters = np.zeros((4, 19, num_itr))
for i, al in enumerate([0, 0.12, 0.25, 0.5]):
for j, iteration in enumerate(np.append(np.arange(1,10, 1), np.arange(10, 110, 10))):
for itr in range(num_itr):
t_true = np.random.rand()*np.pi/2
m_star = (np.cos(t_true),np.sin(t_true))
# print(itr)
m,C = max_quasiconcave_metric_nPTA(1e-4, al, iteration)
# re.append(abs(phi2(m[0], m[1])-phi2(m_star[0], m_star[1])))
t = np.arccos(m[0])
if t > np.pi/2:
t = np.pi*2-t
re_arr_nPTA_iters[i][j][itr] = np.linalg.norm(m-m_star)
re_max_nPTA = np.amax(re_arr_nPTA_iters, 2)
re_min_nPTA = np.amin(re_arr_nPTA_iters, 2)
re_mean_nPTA = np.mean(re_arr_nPTA_iters, 2)
re_median_nPTA = np.median(re_arr_nPTA_iters, 2)
re_std_nPTA = np.std(re_arr_nPTA_iters, 2)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="jN8oUng5TSWu" outputId="c69b2a2d-c1fe-4380-9ec4-fcd35bbc6052"
alphas = [0, 0.12, 0.25, 0.5]
num_iters = np.append(np.arange(1,10, 1), np.arange(10, 110, 10))
hoeffding_iters = np.sqrt(-np.log(0.975)/(np.multiply(2,num_iters)))
# print(hoeffding_iters)
for i in range(4):
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(num_iters, re_mean_nPTA[i], "-", alpha=1, label='non-PTA mean') # plot mean relative error
# ax.plot(num_iters, re_mean[i], "-", alpha=1, label='PTA mean') # plot mean relative error
ax.errorbar(num_iters, re_mean_nPTA[i], yerr=hoeffding_iters, capsize=4, label='non-PTA mean')
ax.errorbar(num_iters, re_mean[i], yerr=hoeffding_iters, capsize=4, label='PTA mean')
# plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.title("Loss vs num_iter alpha={}".format(alphas[i]))
plt.grid()
plt.xlabel("num_iter")
plt.ylabel("2-norm loss")
plt.legend(loc='upper left')
plt.savefig("Loss vs num_iter alpha={}.png".format(alphas[i]))
files.download("Loss vs num_iter alpha={}.png".format(alphas[i]))
plt.show()
# + [markdown] id="8WS1GHn560Dl"
# # log plots
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Ypek-iQfjDw-" outputId="bba30a82-537a-458f-ae7c-c23cf84099be"
alphas = [0, 0.12, 0.25, 0.5]
num_iters = np.append(np.arange(1,10, 1), np.arange(10, 110, 10))
hoeffding_iters = np.sqrt(-np.log(0.975)/(np.multiply(2,num_iters)))
# print(hoeffding_iters)
for i in range(4):
fig, ax = plt.subplots()
fig.set_size_inches(16, 6)
# ax.plot(num_iters, re_mean_nPTA[i], "-", alpha=1, label='non-PTA mean') # plot mean relative error
# ax.plot(num_iters, re_mean[i], "-", alpha=1, label='PTA mean') # plot mean relative error
ax.errorbar(num_iters, re_mean_nPTA[i], yerr=hoeffding_iters, capsize=4, label='non-PTA mean')
ax.errorbar(num_iters, re_mean[i], yerr=hoeffding_iters, capsize=4, label='PTA mean')
# plt.xticks(np.arange(0, .5, step=step), rotation=60, size="small")
plt.title("Loss vs num_iter alpha={}".format(alphas[i]))
plt.grid()
plt.xlabel("num_iter (log-scale)")
plt.ylabel("2-norm loss")
plt.legend(loc='upper left')
# plt.yscale('log')
plt.xscale('log')
plt.savefig("Loss vs num_iter alpha={} (xscale=log).png".format(alphas[i]))
files.download("Loss vs num_iter alpha={} (xscale=log).png".format(alphas[i]))
plt.show()
# + id="24Q2Hb7m4zYL"
|
PTA_LPM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8
# language: python
# name: python3
# ---
# <center>
# <img src="https://gitlab.com/ibm/skills-network/courses/placeholder101/-/raw/master/labs/module%201/images/IDSNlogo.png" width="300" alt="cognitiveclass.ai logo" />
# </center>
#
# # **SpaceX Falcon 9 first stage Landing Prediction**
#
# # Lab 1: Collecting the data
#
# Estimated time needed: **45** minutes
#
# In this capstone, we will predict if the Falcon 9 first stage will land successfully. SpaceX advertises Falcon 9 rocket launches on its website with a cost of 62 million dollars; other providers cost upward of 165 million dollars each, much of the savings is because SpaceX can reuse the first stage. Therefore if we can determine if the first stage will land, we can determine the cost of a launch. This information can be used if an alternate company wants to bid against SpaceX for a rocket launch. In this lab, you will collect and make sure the data is in the correct format from an API. The following is an example of a successful and launch.
#
# 
#
# Several examples of an unsuccessful landing are shown here:
#
# 
#
# Most unsuccessful landings are planned. Space X performs a controlled landing in the oceans.
#
# ## Objectives
#
# In this lab, you will make a get request to the SpaceX API. You will also do some basic data wrangling and formating.
#
# * Request to the SpaceX API
# * Clean the requested data
#
# ***
#
# ## Import Libraries and Define Auxiliary Functions
#
# We will import the following libraries into the lab
#
# +
# Requests allows us to make HTTP requests which we will use to get data from an API
import requests
# Pandas is a software library written for the Python programming language for data manipulation and analysis.
import pandas as pd
# NumPy is a library for the Python programming language, adding support for large, multi-dimensional arrays and matrices, along with a large collection of high-level mathematical functions to operate on these arrays
import numpy as np
# Datetime is a library that allows us to represent dates
import datetime
# Setting this option will print all collumns of a dataframe
pd.set_option('display.max_columns', None)
# Setting this option will print all of the data in a feature
pd.set_option('display.max_colwidth', None)
# -
# Below we will define a series of helper functions that will help us use the API to extract information using identification numbers in the launch data.
#
# From the <code>rocket</code> column we would like to learn the booster name.
#
# Takes the dataset and uses the rocket column to call the API and append the data to the list
def getBoosterVersion(data):
for x in data['rocket']:
response = requests.get("https://api.spacexdata.com/v4/rockets/"+str(x)).json()
BoosterVersion.append(response['name'])
# From the <code>launchpad</code> we would like to know the name of the launch site being used, the logitude, and the latitude.
#
# Takes the dataset and uses the launchpad column to call the API and append the data to the list
def getLaunchSite(data):
for x in data['launchpad']:
response = requests.get("https://api.spacexdata.com/v4/launchpads/"+str(x)).json()
Longitude.append(response['longitude'])
Latitude.append(response['latitude'])
LaunchSite.append(response['name'])
# From the <code>payload</code> we would like to learn the mass of the payload and the orbit that it is going to.
#
# Takes the dataset and uses the payloads column to call the API and append the data to the lists
def getPayloadData(data):
for load in data['payloads']:
response = requests.get("https://api.spacexdata.com/v4/payloads/"+load).json()
PayloadMass.append(response['mass_kg'])
Orbit.append(response['orbit'])
# From <code>cores</code> we would like to learn the outcome of the landing, the type of the landing, number of flights with that core, whether gridfins were used, wheter the core is reused, wheter legs were used, the landing pad used, the block of the core which is a number used to seperate version of cores, the number of times this specific core has been reused, and the serial of the core.
#
# Takes the dataset and uses the cores column to call the API and append the data to the lists
def getCoreData(data):
for core in data['cores']:
if core['core'] != None:
response = requests.get("https://api.spacexdata.com/v4/cores/"+core['core']).json()
Block.append(response['block'])
ReusedCount.append(response['reuse_count'])
Serial.append(response['serial'])
else:
Block.append(None)
ReusedCount.append(None)
Serial.append(None)
Outcome.append(str(core['landing_success'])+' '+str(core['landing_type']))
Flights.append(core['flight'])
GridFins.append(core['gridfins'])
Reused.append(core['reused'])
Legs.append(core['legs'])
LandingPad.append(core['landpad'])
# Now let's start requesting rocket launch data from SpaceX API with the following URL:
#
spacex_url="https://api.spacexdata.com/v4/launches/past"
response = requests.get(spacex_url)
# Check the content of the response
#
print(response.content)
# You should see the response contains massive information about SpaceX launches. Next, let's try to discover some more relevant information for this project.
#
# ### Task 1: Request and parse the SpaceX launch data using the GET request
#
# To make the requested JSON results more consistent, we will use the following static response object for this project:
#
static_json_url='https://cf-courses-data.s3.us.cloud-object-storage.appdomain.cloud/IBM-DS0321EN-SkillsNetwork/datasets/API_call_spacex_api.json'
# We should see that the request was successfull with the 200 status response code
#
response.status_code
# Now we decode the response content as a Json using <code>.json()</code> and turn it into a Pandas dataframe using <code>.json_normalize()</code>
#
# Use json_normalize meethod to convert the json result into a dataframe
json_content = response.json()
data = pd.json_normalize(json_content)
# Using the dataframe <code>data</code> print the first 5 rows
#
# Get the head of the dataframe
data.head()
# You will notice that a lot of the data are IDs. For example the rocket column has no information about the rocket just an identification number.
#
# We will now use the API again to get information about the launches using the IDs given for each launch. Specifically we will be using columns <code>rocket</code>, <code>payloads</code>, <code>launchpad</code>, and <code>cores</code>.
#
# +
# Lets take a subset of our dataframe keeping only the features we want and the flight number, and date_utc.
data = data[['rocket', 'payloads', 'launchpad', 'cores', 'flight_number', 'date_utc']]
# We will remove rows with multiple cores because those are falcon rockets with 2 extra rocket boosters and rows that have multiple payloads in a single rocket.
data = data[data['cores'].map(len)==1]
data = data[data['payloads'].map(len)==1]
# Since payloads and cores are lists of size 1 we will also extract the single value in the list and replace the feature.
data['cores'] = data['cores'].map(lambda x : x[0])
data['payloads'] = data['payloads'].map(lambda x : x[0])
# We also want to convert the date_utc to a datetime datatype and then extracting the date leaving the time
data['date'] = pd.to_datetime(data['date_utc']).dt.date
# Using the date we will restrict the dates of the launches
data = data[data['date'] <= datetime.date(2020, 11, 13)]
# -
# * From the <code>rocket</code> we would like to learn the booster name
#
# * From the <code>payload</code> we would like to learn the mass of the payload and the orbit that it is going to
#
# * From the <code>launchpad</code> we would like to know the name of the launch site being used, the longitude, and the latitude.
#
# * From <code>cores</code> we would like to learn the outcome of the landing, the type of the landing, number of flights with that core, whether gridfins were used, whether the core is reused, whether legs were used, the landing pad used, the block of the core which is a number used to seperate version of cores, the number of times this specific core has been reused, and the serial of the core.
#
# The data from these requests will be stored in lists and will be used to create a new dataframe.
#
#Global variables
BoosterVersion = []
PayloadMass = []
Orbit = []
LaunchSite = []
Outcome = []
Flights = []
GridFins = []
Reused = []
Legs = []
LandingPad = []
Block = []
ReusedCount = []
Serial = []
Longitude = []
Latitude = []
# These functions will apply the outputs globally to the above variables. Let's take a looks at <code>BoosterVersion</code> variable. Before we apply <code>getBoosterVersion</code> the list is empty:
#
BoosterVersion
# Now, let's apply <code> getBoosterVersion</code> function method to get the booster version
#
# Call getBoosterVersion
getBoosterVersion(data)
# the list has now been update
#
BoosterVersion[0:5]
# we can apply the rest of the functions here:
#
# Call getLaunchSite
getLaunchSite(data)
# Call getPayloadData
getPayloadData(data)
# Call getCoreData
getCoreData(data)
# Finally lets construct our dataset using the data we have obtained. We we combine the columns into a dictionary.
#
launch_dict = {'FlightNumber': list(data['flight_number']),
'Date': list(data['date']),
'BoosterVersion':BoosterVersion,
'PayloadMass':PayloadMass,
'Orbit':Orbit,
'LaunchSite':LaunchSite,
'Outcome':Outcome,
'Flights':Flights,
'GridFins':GridFins,
'Reused':Reused,
'Legs':Legs,
'LandingPad':LandingPad,
'Block':Block,
'ReusedCount':ReusedCount,
'Serial':Serial,
'Longitude': Longitude,
'Latitude': Latitude}
# Then, we need to create a Pandas data frame from the dictionary launch_dict.
#
# Create a data from launch_dict
launch_df = pd.DataFrame.from_dict(launch_dict)
# Show the summary of the dataframe
#
# Show the head of the dataframe
launch_df.head()
# ### Task 2: Filter the dataframe to only include `Falcon 9` launches
#
# Finally we will remove the Falcon 1 launches keeping only the Falcon 9 launches. Filter the data dataframe using the <code>BoosterVersion</code> column to only keep the Falcon 9 launches. Save the filtered data to a new dataframe called <code>data_falcon9</code>.
#
# Hint data['BoosterVersion']!='Falcon 1'
data_falcon9 = launch_df[launch_df['BoosterVersion']!='Falcon 1']
# Now that we have removed some values we should reset the FlgihtNumber column
#
data_falcon9.loc[:,'FlightNumber'] = list(range(1, data_falcon9.shape[0]+1))
data_falcon9
# ## Data Wrangling
#
# We can see below that some of the rows are missing values in our dataset.
#
data_falcon9.isnull().sum()
# Before we can continue we must deal with these missing values. The <code>LandingPad</code> column will retain None values to represent when landing pads were not used.
#
# ### Task 3: Dealing with Missing Values
#
# Calculate below the mean for the <code>PayloadMass</code> using the <code>.mean()</code>. Then use the mean and the <code>.replace()</code> function to replace `np.nan` values in the data with the mean you calculated.
#
# +
# Calculate the mean value of PayloadMass column
mean_payload = data_falcon9.PayloadMass.mean()
# Replace the np.nan values with its mean value
data_falcon9.PayloadMass = data_falcon9.PayloadMass.replace(np.nan, mean_payload)
# -
# You should see the number of missing values of the <code>PayLoadMass</code> change to zero.
#
# Now we should have no missing values in our dataset except for in <code>LandingPad</code>.
#
# We can now export it to a <b>CSV</b> for the next section,but to make the answers consistent, in the next lab we will provide data in a pre-selected date range.
#
# <code>data_falcon9.to_csv('dataset_part\_1.csv', index=False)</code>
#
# ## Authors
#
# <a href="https://www.linkedin.com/in/joseph-s-50398b136/?utm_medium=Exinfluencer&utm_source=Exinfluencer&utm_content=000026UJ&utm_term=10006555&utm_id=NA-SkillsNetwork-Channel-SkillsNetworkCoursesIBMDS0321ENSkillsNetwork26802033-2021-01-01"><NAME></a> has a PhD in Electrical Engineering, his research focused on using machine learning, signal processing, and computer vision to determine how videos impact human cognition. Joseph has been working for IBM since he completed his PhD.
#
# ## Change Log
#
# | Date (YYYY-MM-DD) | Version | Changed By | Change Description |
# | ----------------- | ------- | ---------- | ----------------------------------- |
# | 2020-09-20 | 1.1 | Joseph | get result each time you run |
# | 2020-09-20 | 1.1 | Azim | Created Part 1 Lab using SpaceX API |
# | 2020-09-20 | 1.0 | Joseph | Modified Multiple Areas |
#
# Copyright © 2021 IBM Corporation. All rights reserved.
#
|
Data Collection API Lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Homework 2
#
# Name: <NAME>
# <br>
# ID: u1265976
# ## Question1
# #### Problem Satement
# The aim is to distinguish between the presence and absence of cardiac arrhythmia and to classify it in one of the 16 groups. <br>Class 01 refers to 'normal' ECG.<br>
# Classes 02 to 15 refers to different classes of arrhythmia .<br>
# Class 16 refers to the rest of unclassified ones. <br>
#
#
# The motto of the project is to built differnet machine learning models and neural netwoek to perfomr the classification.
# Dataset : https://archive.ics.uci.edu/ml/datasets/Arrhythmia
# #### Dataset Specification
# This database contains 279 attributes, 206 of which are linear valued and the rest are nominal.
#
# 1 Age: Age in years , linear <br>
# 2 Sex: Sex (0 = male; 1 = female) , nominal <br>
# 3 Height: Height in centimeters , linear <br>
# 4 Weight: Weight in kilograms , linear <br>
# 5 QRS duration: Average of QRS duration in msec., linear<br>
# 6 P-R interval: Average duration between onset of P and Q waves in msec., linear <br>
# 7 Q-T interval: Average duration between onset of Q and offset of T waves in msec., linear <br>
# 8 T interval: Average duration of T wave in msec., linear <br>
# 9 P interval: Average duration of P wave in msec., linear <br>
# Vector angles in degrees on front plane of:, linear <br>
# 10 QRS <br>
# 11 T <br>
# 12 P <br>
# 13 QRST <br>
# 14 J<br>
#
# 15 Heart rate: Number of heart beats per minute ,linear
# <br>
# Of channel DI:
# Average width, in msec., of: linear
# 16 Q wave <br>
# 17 R wave <br>
# 18 S wave <br>
# 19 R' wave, small peak just after R <br>
# 20 S' wave <br>
# <br>
# 21 Number of intrinsic deflections, linear
# <br>
# 22 Existence of ragged R wave, nominal
# <br>
# 23 Existence of diphasic derivation of R wave, nominal
# <br>
# 24 Existence of ragged P wave, nominal
# <br>
# 25 Existence of diphasic derivation of P wave, nominal
# <br>
# 26 Existence of ragged T wave, nominal
# <br>
# 27 Existence of diphasic derivation of T wave, nominal
# <br>
# Of channel DII:
# <br>
# 28 .. 39 (similar to 16 .. 27 of channel DI)
# <br>
# Of channels DIII:
# <br>
# 40 .. 51
# <br>
# Of channel AVR:
# 52 .. 63
# <br>
# Of channel AVL:
# 64 .. 75
# <br>
# Of channel AVF:
# 76 .. 87
# <br>
# Of channel V1:
# 88 .. 99
# <br>
# Of channel V2:
# 100 .. 111
# <br>
# Of channel V3:
# 112 .. 123
# <br>
# Of channel V4:
# 124 .. 135
# <br>
# Of channel V5:
# 136 .. 147
# <br>
# Of channel V6:
# 148 .. 159
#
# <br>
# Of channel DI:
# Amplitude , * 0.1 milivolt, of
#
# 160 JJ wave, linear <br>
# 161 Q wave, linear <br>
# 162 R wave, linear <br>
# 163 S wave, linear <br>
# 164 R' wave, linear <br>
# 165 S' wave, linear <br>
# 166 P wave, linear <br>
# 167 T wave, linear <br>
# <br>
# 168 QRSA , Sum of areas of all segments divided by 10, ( Area= width * height / 2 ), linear
# 169 QRSTA = QRSA + 0.5 * width of T wave * 0.1 * height of T wave. (If T is diphasic then the bigger segment is considered), linear
#
# <br>
# Of channel DII:
# 170 .. 179
# <br>
# Of channel DIII:
# 180 .. 189
# <br>
# Of channel AVR:
# 190 .. 199
# <br>
# Of channel AVL:
# 200 .. 209
# <br>
# Of channel AVF:
# 210 .. 219
# <br>
# Of channel V1:
# 220 .. 229
# <br>
# Of channel V2:
# 230 .. 239
# <br>
# Of channel V3:
# 240 .. 249
# <br>
# Of channel V4:
# 250 .. 259
# <br>
# Of channel V5:
# 260 .. 269
# <br>
# Of channel V6:
# 270 .. 279
# #### Project Milestone
# Week 1: Exploratory Analysis <br>
# week 2: Perform Lietreature Review <br>
# Week 3 and 4: Inspect different Validation Techniques <br>
# week 5 and 6. Build Machine Learning Models <br>
# Week 7 and 8: Develop Neural network with different architecture <br>
# week 9: Compare the results of Machine Learning and Deep Learning Models.(Documentation and presentation of results)<br>
# ## Question2
# #### For the flow chart please open the pdf in the present directory.
# +
#Import Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.metrics import f1_score
from sklearn.preprocessing import MinMaxScaler,StandardScaler
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.metrics import roc_curve, auc, roc_auc_score
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from imblearn.over_sampling import RandomOverSampler
from imblearn.over_sampling import SMOTE
import warnings
warnings.filterwarnings('ignore')
# -
#Load the Data
data = pd.read_csv('hepatitis.data',header=None)
data.replace('?',np.nan,inplace=True)
X = data[data.columns[1:20]]
y = data[data.columns[0]]
# +
#converting to numeric or continus to float
cat_var = [2,3,4,5,6,7,8,9,10,11,12,13,19]
num_var = [1,14,15,16,17,18]
# -
for i in num_var:
X.loc[:,i] = X.loc[:,i].astype(float)
for i in cat_var:
X.loc[:,i] = X.loc[:,i].astype(str)
X_train1, X_test, y_train1, y_test = train_test_split(X, y, test_size=0.20, random_state=42)
X_train, X_val, y_train, y_val = train_test_split(X_train1, y_train1, test_size=0.25, random_state=42)
train_mean={}
train_mode={}
for i in num_var:
train_mean[i] = X_train[i].mean()
for i in cat_var:
train_mode[i] = X_train[i].mode()[0]
# +
#Replacing with mean amd mode in train
for i in num_var:
X_train.loc[:,i].replace(np.nan,train_mean[i],inplace= True)
for i in cat_var:
X_train.loc[:,i].replace("nan",train_mode[i],inplace= True)
# +
#Replacing with train mean and mode in test and validation
for i in num_var:
X_val.loc[:,i].replace(np.nan,train_mean[i],inplace= True)
X_test.loc[:,i].replace(np.nan,train_mean[i],inplace= True)
for i in cat_var:
X_val.loc[:,i].replace("nan",train_mode[i],inplace= True)
X_test.loc[:,i].replace("nan",train_mode[i],inplace= True)
# -
X_train=pd.get_dummies(X_train)
X_val=pd.get_dummies(X_val)
X_test=pd.get_dummies(X_test)
clf = StandardScaler()
clf.fit(X_train)
X_train = pd.DataFrame(clf.transform(X_train))
X_val = pd.DataFrame(clf.transform(X_val))
X_test = pd.DataFrame(clf.transform(X_test))
# # RandomOverSampler
ros = RandomOverSampler(random_state=42)
X_res, y_res = ros.fit_resample(X_train, y_train)
# ### KNN classifier -RandomOverSampler
#KNN classifier
model = KNeighborsClassifier(n_neighbors=3) # Load our classifier
model.fit(X_res, y_res)
prediction_knn = model.predict(X_val) # Make predictions with our trained model on the test data
print(f1_score(y_val, prediction_knn) * 100)
# ### Logistic Regression - RandomOverSampler
logreg = LogisticRegression(C=1e5,solver='liblinear')
logreg.fit(X_res, y_res)
prediction_lr = logreg.predict(X_val)
print(f1_score(y_val, prediction_lr) * 100)
# ### RandomForest - RandomOverSampler
clf = RandomForestClassifier(max_depth=2, random_state=0)
clf.fit(X_res, y_res)
prediction_rf = clf.predict(X_val)
print(f1_score(y_val, prediction_rf) * 100)
# # SMOTE
sm = SMOTE(random_state=42)
X_res_s, y_res_s = sm.fit_resample(X_train, y_train)
# ### KNN Classifier -SMOTE
#KNN classifier
model_2 = KNeighborsClassifier(n_neighbors=3) # Load our classifier
model_2.fit(X_res_s, y_res_s)
prediction_knn_s = model_2.predict(X_val) # Make predictions with our trained model on the test data
print(f1_score(y_val, prediction_knn_s) * 100)
# ### Logistic Regression-SMOTE
logreg_2 = LogisticRegression(C=1e5,solver='liblinear')
logreg_2.fit(X_res_s, y_res_s)
prediction_lr_s = logreg_2.predict(X_val)
print(f1_score(y_val, prediction_lr_s) * 100)
# ### Random Forest - SMOTE
clf_2 = RandomForestClassifier(max_depth=2, random_state=0)
clf_2.fit(X_res_s, y_res_s)
prediction_rf_s = clf_2.predict(X_val)
print(f1_score(y_val, prediction_rf_s) * 100)
# ## ROC of RandomOverSample
prediction_random = model_2.predict(X_test)
print(prediction_random)
print(y_test)
fpr, tpr, thresholds = roc_curve(y_test,prediction_random,pos_label=2)
areaUnderROC = auc(fpr, tpr)
plt.figure()
plt.plot(fpr, tpr, color='r', lw=2, label='ROC curve')
plt.plot([0, 1], [0, 1], color='k', lw=2, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic: Area under the curve = {0:0.2f}'.format(areaUnderROC))
plt.legend(loc="lower right")
plt.show()
# ## ROC of SMOTE
prediction_smote= model_2.predict(X_test)
fpr, tpr, thresholds = roc_curve(y_test,prediction_smote,pos_label=2)
areaUnderROC = auc(fpr, tpr)
plt.figure()
plt.plot(fpr, tpr, color='r', lw=2, label='ROC curve')
plt.plot([0, 1], [0, 1], color='k', lw=2, linestyle='--')
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.05])
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.title('Receiver operating characteristic: Area under the curve = {0:0.2f}'.format(areaUnderROC))
plt.legend(loc="lower right")
plt.show()
|
Assignments/HW2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import glob
reviews_dir = './data_ml_2020/movies_reviews'
# +
negative_reviews = []
positive_reviews = []
for file in glob.glob(reviews_dir + "/neg/*.txt"):
with open(file, "r") as f:
negative_reviews.append(f.read().replace("\n", ""))
for file in glob.glob(reviews_dir + "/pos/*.txt"):
with open(file, "r") as f:
positive_reviews.append(f.read().replace("\n", ""))
# -
# Use Glob to get the file paths and add them to an array.
#
# Then match them with 0 or 1. 0 if they are negative, 1 if they are positive.
# +
import numpy as np
X = np.concatenate((negative_reviews, positive_reviews))
Y = np.concatenate((np.zeros((len(negative_reviews))), np.ones((len(positive_reviews)))))
# +
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y,test_size=0.4,random_state=20)
# -
# CountVectorizer transforms the dataset into a vector matrix (counts each word). Then, the TfidfTransformer reduces the impact of common words such as "The". This is the transformation step where we get data ready for modeling.
# +
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
cv = CountVectorizer()
X_train_cv = cv.fit_transform(X_train)
tf_trans = TfidfTransformer()
X_train_tf = tf_trans.fit_transform(X_train_cv)
# -
# The docs suggest using either MultinomialNB or ComplementNB. I decided to use both to test which one is better.
# As you can see from the results below, CNB has a slight edge over the MNB, so if I was to only use one for a dataset that required more training I'd definitely use CNB.
# +
from sklearn.naive_bayes import MultinomialNB, ComplementNB
from sklearn.metrics import accuracy_score
mnb_model = MultinomialNB().fit(X_train_tf, Y_train)
cnb_model = ComplementNB().fit(X_train_tf, Y_train)
# +
print(f"Score for MNB: {mnb_model.score(X_train_tf, Y_train)}")
print(f"Score for CNB: {cnb_model.score(X_train_tf, Y_train)}")
print(f"Accuracy Score for MNB: {accuracy_score(Y_train, mnb_model.predict(X_train_tf))}")
print(f"Accuracy Score for CNB: {accuracy_score(Y_train, cnb_model.predict(X_train_tf))}")
# +
X_test_cv = cv.transform(X_test)
X_test_tf = tf_trans.transform(X_test_cv)
print(f"Score for MNB: {mnb_model.score(X_test_tf, Y_test)}")
print(f"Score for CNB: {cnb_model.score(X_test_tf, Y_test)}")
print(f"Accuracy for MNB: {accuracy_score(Y_test, mnb_model.predict(X_test_tf))}")
print(f"Accuracy for CNB: {accuracy_score(Y_test, cnb_model.predict(X_test_tf))}")
# +
import os
predict_reviews = []
predict_data = []
for file in glob.glob(reviews_dir + "/review/*.txt"):
with open(file, "r", encoding="utf8") as f:
predict_reviews.append(f.read().replace("\n", ""))
filename = os.path.splitext(os.path.basename(file))[0]
title = " ".join([word.capitalize() for word in filename.split(" ")[0].replace("_", " ").split(" ")])
rating = filename.split(" ")[1].replace("p", ".").replace("o", "/")
predict_data.append({"title": title, "rating": rating})
# +
X_predict_cv = cv.transform(predict_reviews)
X_predict_tf = tf_trans.transform(X_predict_cv)
prediction = mnb_model.predict(X_predict_tf)
def predict(i: int) -> str:
if i == 0:
return "Negative"
if i == 1:
return "Positive"
i = 0
for p in prediction:
print(f"{predict_data[i]['title']} | {predict_data[i]['rating']} | {predict(p)}")
i += 1
# +
prediction = cnb_model.predict(X_predict_tf)
i = 0
for p in prediction:
print(f"{predict_data[i]['title']} | {predict_data[i]['rating']} | {predict(p)}")
i += 1
# -
# All the reviews were above average except for Godmothered, but when I looked at the review, although it was very cruel about the movie a majority of the review was dedicated to praise for the star <NAME>. So even though the review was overall negative, I can see why the model decided to assign it a 'positive' rating - because it was (in a way)...
#
# Maybe with more training it would properly predict that dataset.
|
Text Classification with Naive Bayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# select a GPU
import os
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = "1"
# +
#imports
import numpy as np
import matplotlib.pyplot as plt
import h5py
import scipy.io
from sklearn.metrics import confusion_matrix
import pandas as pd
from DCASE_plots import plot_confusion_matrix
import librosa
import soundfile as sound
import keras
import tensorflow
print("Librosa version = ",librosa.__version__)
print("Pysoundfile version = ",sound.__version__)
print("keras version = ",keras.__version__)
print("tensorflow version = ",tensorflow.__version__)
# -
#Task 1a dev validation set
ThisPath = '../Task1b/'
File = ThisPath + 'evaluation_setup/fold1_evaluate.csv'
sr = 44100
SampleDuration = 10
NumFreqBins = 128
NumFFTPoints = 2048
HopLength = int(NumFFTPoints/2)
NumTimeBins = int(np.ceil(SampleDuration*sr/HopLength))
# +
#load filenames and labels
dev_test_df = pd.read_csv(File,sep='\t', encoding='ASCII')
Inds_device_a=np.where(dev_test_df['filename'].str.contains("-a.wav")==True)[0]
Inds_device_b=np.where(dev_test_df['filename'].str.contains("-b.wav")==True)[0]
Inds_device_c=np.where(dev_test_df['filename'].str.contains("-c.wav")==True)[0]
Inds_device_bc=np.concatenate((Inds_device_b,Inds_device_c),axis=-1)
wavpaths = dev_test_df['filename'].tolist()
ClassNames = np.unique(dev_test_df['scene_label'])
y_val_labels = dev_test_df['scene_label'].astype('category').cat.codes.values
#swap codes for 2 and 1 to match the DCASE ordering of classes
a1=np.where(y_val_labels==2)
a2=np.where(y_val_labels==3)
y_val_labels.setflags(write=1)
y_val_labels[a1] = 3
y_val_labels[a2] = 2
# +
#load wav files and get log-mel spectrograms, deltas, and delta-deltas
def deltas(X_in):
X_out = (X_in[:,:,2:,:]-X_in[:,:,:-2,:])/10.0
X_out = X_out[:,:,1:-1,:]+(X_in[:,:,4:,:]-X_in[:,:,:-4,:])/5.0
return X_out
LM_val = np.zeros((len(wavpaths),NumFreqBins,NumTimeBins,1),'float32')
for i in range(len(wavpaths)):
audio_data,fs = sound.read(ThisPath + wavpaths[i],stop=SampleDuration*sr)
LM_val[i,:,:,0]= librosa.feature.melspectrogram(audio_data,
sr=sr,
n_fft=NumFFTPoints,
hop_length=HopLength,
n_mels=NumFreqBins,
fmin=0.0,
fmax=sr/2,
htk=True,
norm=None)
LM_val=np.log(LM_val)
LM_deltas_val = deltas(LM_val)
LM_deltas_deltas_val = deltas(LM_deltas_val)
LM_val = np.concatenate((LM_val[:,:,4:-4,:],LM_deltas_val[:,:,2:-2,:],LM_deltas_deltas_val),axis=-1)
# -
#load and run the model
best_model = keras.models.load_model('DCASE_Task1b_development.h5')
y_pred_val = np.argmax(best_model.predict(LM_val),axis=1)
# +
#get metrics for all devices combined
Overall_accuracy = np.sum(y_pred_val==y_val_labels)/LM_val.shape[0]
print("overall accuracy: ", Overall_accuracy)
plot_confusion_matrix(y_val_labels, y_pred_val, ClassNames,normalize=True,title="Task 1b, all devices")
conf_matrix = confusion_matrix(y_val_labels,y_pred_val)
conf_mat_norm_recall = conf_matrix.astype('float32')/conf_matrix.sum(axis=1)[:,np.newaxis]
conf_mat_norm_precision = conf_matrix.astype('float32')/conf_matrix.sum(axis=0)[:,np.newaxis]
recall_by_class = np.diagonal(conf_mat_norm_recall)
precision_by_class = np.diagonal(conf_mat_norm_precision)
mean_recall = np.mean(recall_by_class)
mean_precision = np.mean(precision_by_class)
print("per-class accuracy (recall): ",recall_by_class)
print("per-class precision: ",precision_by_class)
print("mean per-class recall: ",mean_recall)
print("mean per-class precision: ",mean_precision)
# +
#get metrics for device A only
Overall_accuracy = np.sum(y_pred_val[Inds_device_a]==y_val_labels[Inds_device_a])/len(Inds_device_a)
print("overall accuracy: ", Overall_accuracy)
plot_confusion_matrix(y_val_labels[Inds_device_a], y_pred_val[Inds_device_a], ClassNames,normalize=True,title="Task 1b, Device A")
conf_matrix = confusion_matrix(y_val_labels[Inds_device_a],y_pred_val[Inds_device_a])
conf_mat_norm_recall = conf_matrix.astype('float32')/conf_matrix.sum(axis=1)[:,np.newaxis]
conf_mat_norm_precision = conf_matrix.astype('float32')/conf_matrix.sum(axis=0)[:,np.newaxis]
recall_by_class = np.diagonal(conf_mat_norm_recall)
precision_by_class = np.diagonal(conf_mat_norm_precision)
mean_recall = np.mean(recall_by_class)
mean_precision = np.mean(precision_by_class)
print("per-class accuracy (recall): ",recall_by_class)
print("per-class precision: ",precision_by_class)
print("mean per-class recall: ",mean_recall)
print("mean per-class precision: ",mean_precision)
# +
#get metrics for device B only
Overall_accuracy = np.sum(y_pred_val[Inds_device_b]==y_val_labels[Inds_device_b])/len(Inds_device_b)
print("overall accuracy: ", Overall_accuracy)
plot_confusion_matrix(y_val_labels[Inds_device_b], y_pred_val[Inds_device_b], ClassNames,normalize=True,title="Task 1b, Device B")
conf_matrix = confusion_matrix(y_val_labels[Inds_device_b],y_pred_val[Inds_device_b])
conf_mat_norm_recall = conf_matrix.astype('float32')/conf_matrix.sum(axis=1)[:,np.newaxis]
conf_mat_norm_precision = conf_matrix.astype('float32')/conf_matrix.sum(axis=0)[:,np.newaxis]
recall_by_class = np.diagonal(conf_mat_norm_recall)
precision_by_class = np.diagonal(conf_mat_norm_precision)
mean_recall = np.mean(recall_by_class)
mean_precision = np.mean(precision_by_class)
print("per-class accuracy (recall): ",recall_by_class)
print("per-class precision: ",precision_by_class)
print("mean per-class recall: ",mean_recall)
print("mean per-class precision: ",mean_precision)
# +
#get metrics for device C only
Overall_accuracy = np.sum(y_pred_val[Inds_device_c]==y_val_labels[Inds_device_c])/len(Inds_device_c)
print("overall accuracy: ", Overall_accuracy)
plot_confusion_matrix(y_val_labels[Inds_device_c], y_pred_val[Inds_device_c], ClassNames,normalize=True,title="Task 1b, Device C")
conf_matrix = confusion_matrix(y_val_labels[Inds_device_c],y_pred_val[Inds_device_c])
conf_mat_norm_recall = conf_matrix.astype('float32')/conf_matrix.sum(axis=1)[:,np.newaxis]
conf_mat_norm_precision = conf_matrix.astype('float32')/conf_matrix.sum(axis=0)[:,np.newaxis]
recall_by_class = np.diagonal(conf_mat_norm_recall)
precision_by_class = np.diagonal(conf_mat_norm_precision)
mean_recall = np.mean(recall_by_class)
mean_precision = np.mean(precision_by_class)
print("per-class accuracy (recall): ",recall_by_class)
print("per-class precision: ",precision_by_class)
print("mean per-class recall: ",mean_recall)
print("mean per-class precision: ",mean_precision)
# +
#get metrics for device B and C
Overall_accuracy = np.sum(y_pred_val[Inds_device_bc]==y_val_labels[Inds_device_bc])/len(Inds_device_bc)
print("overall accuracy: ", Overall_accuracy)
plot_confusion_matrix(y_val_labels[Inds_device_bc], y_pred_val[Inds_device_bc], ClassNames,normalize=True,title="Task 1b, Device B and C")
conf_matrix = confusion_matrix(y_val_labels[Inds_device_bc],y_pred_val[Inds_device_bc])
conf_mat_norm_recall = conf_matrix.astype('float32')/conf_matrix.sum(axis=1)[:,np.newaxis]
conf_mat_norm_precision = conf_matrix.astype('float32')/conf_matrix.sum(axis=0)[:,np.newaxis]
recall_by_class = np.diagonal(conf_mat_norm_recall)
precision_by_class = np.diagonal(conf_mat_norm_precision)
mean_recall = np.mean(recall_by_class)
mean_precision = np.mean(precision_by_class)
print("per-class accuracy (recall): ",recall_by_class)
print("per-class precision: ",precision_by_class)
print("mean per-class recall: ",mean_recall)
print("mean per-class precision: ",mean_precision)
# -
|
DCASE_Task1b_inference.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Map View Selection
#
# With this application, users can select and downsample data interactively in 2D plan view.
#
# <img align="right" width="50%" src="./images/plot_selection.png">
#
#
# New user? Visit the [Getting Started](../installation.rst) page.
# ## Application
# The following sections provide details on the different parameters controling the application. Interactive widgets shown below are for demonstration purposes only.
# +
from geoapps.plotting import PlotSelection2D
app = PlotSelection2D(h5file=r"../../../assets/FlinFlon.geoh5")
app.main
# -
# ## Object/Data Selection
#
# List of objects available to pull data from.
app.data_panel
# See the [Object Selection](object_data_selection.ipynb) page for more details.
# ## Resolution
#
# Determine the minimum distance between data points.
app.resolution
# For `Grid2D` objects, the resolution of the grid is downsampled uniformely to the nearest interval. For example, requesting a 15 m minimum resolution on a 10 m base grid will result in a 20 m sub-sampling.
#
# For `Points`, `Curve` and `Surface` objects, the downsampling is done radially using the [Scipy.spatial.cKDTree](https://docs.scipy.org/doc/scipy/reference/generated/scipy.spatial.cKDTree.html) routine.
# ## Window Selection
#
# Manual selection of an area of interest and data resolution.
# ### Northing
#
# Vertical center position (m)
app.center_y
# ### Easting
#
# Horizontal center position (m)
app.center_x
# ### Width
#
# Window size along East (m)
app.width
# ### Height
#
# Window size along North (m)
app.height
# ### Azimuth
#
# Orientation of the selection window, in degree from North.
app.azimuth
# ## Zoom on selection
#
# Set the plotting area to the extent of the window selection, or keep fix to the target object extent.
app.zoom_extent
# Need help? Contact us at <EMAIL>
# + nbsphinx="hidden"
import matplotlib.pyplot as plt
app.figure.savefig("images/plot_selection.png", dpi=300, bbox_inches="tight")
|
docs/content/applications/view_selection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Sylwiaes/numpy_pandas_cwiczenia/blob/main/01_numpy_cwiczenia/021_030_exercises.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="WLEgCW5F1hho"
# ## Numpy
#
# ### Spis treści:
# * [Import biblioteki](#0)
# * [Ćwiczenie 21](#1)
# * [Ćwiczenie 22](#2)
# * [Ćwiczenie 23](#3)
# * [Ćwiczenie 24](#4)
# * [Ćwiczenie 25](#5)
# * [Ćwiczenie 26](#6)
# * [Ćwiczenie 27](#7)
# * [Ćwiczenie 28](#8)
# * [Ćwiczenie 29](#9)
# * [Ćwiczenie 30](#10)
# + [markdown] id="zeLOLmTj1kBM"
# ### <a name='0'></a>Import biblioteki
# + id="HXMB7SMe1Wnz" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="09a49488-fc1f-46e6-e96b-f1298dfa1560"
import numpy as np
np.__version__
# + [markdown] id="cyc70pMe1oCg"
# ### <a name='1'></a> Ćwiczenie 21
# Wygeneruj poniższą tablicę numpy. Zapisz tablicę do pliku bianarnego o nazwie _'array.npy'_ i następnie wczytaj z powrotem ten plik do innej zmiennej.
#
#
# ```
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11]])
# ```
#
#
# __Wskazówka:__ Użyj funkcji _np.save()_ oraz _np.load()_.
# + id="MhvZKgtPKAOF" colab={"base_uri": "https://localhost:8080/"} outputId="e364e9da-25ea-4be6-a365-b66da13a7452"
# tutaj wpisz rozwiązanie
A = np.arange(12).reshape(3, -1)
A
# + id="6CxG1mH3eGx9"
np.save(fname='array.npy', A)
# + colab={"base_uri": "https://localhost:8080/"} id="vVMoKCOhe3xx" outputId="bba603d9-5e09-48b8-84b1-689546ace736"
B = np.load('array.npy')
B
# + [markdown] id="J9pY1rtr1sC0"
# ### <a name='2'></a> Ćwiczenie 22
# Wygeneruj ponizszą tablicę numpy:
#
#
# ```
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11]])
# ```
#
#
# Zapisz tablicę do pliku tekstowego o nazwie _'array.txt'_ z dokładnością do drugiego miejsca po przecinku i następnie wczytaj z powrotem ten plik do innej zmiennej.
#
# __Wskazówka:__ Użyj funkcji _np.savetxt()_ oraz _np.loadtxt()_.
#
# + id="Ac7M7bDhLAlf"
# tutaj wpisz rozwiązanie
A = np.arange(12).reshape(3, -1)
np.savetxt(fname='array.txt', X=A, fmt='%0.2f')
# + colab={"base_uri": "https://localhost:8080/"} id="wMa72GmRfihR" outputId="c47da357-d998-43ee-8752-3bcdaccea5f5"
B = np.loadtxt('array.txt')
B
# + [markdown] id="XQLYT2m11wVG"
# ### <a name='3'></a> Ćwiczenie 23
# Wygeneruj i przekształć poniższą tablicę numpy na listę.
#
#
# ```
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11]])
# ```
#
#
#
# __Wskazówka:__ Użyj metody _np.array.tolist()_.
# + id="Dtu-EJViT1Os" colab={"base_uri": "https://localhost:8080/"} outputId="c3504cdf-2b30-42e2-fe61-1c100fc28e36"
# tutaj wpisz rozwiązanie
A = np.arange(12).reshape(3, -1)
A.tolist()
# + [markdown] id="4xTKSUzg10MK"
# ### <a name='4'></a> Ćwiczenie 24
# Podana jest poniższa tablica numpy:
#
# ```
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11]])
# ```
# Używając operatora wycinania przekształć tablicę w poniższe:
#
# * zamiana wierszy (pierwszy z ostatnim)
#
# ```
# array([[ 8, 9, 10, 11],
# [ 4, 5, 6, 7],
# [ 0, 1, 2, 3]])
# ```
#
#
#
#
#
#
# + id="1iqUDmG3V7sM" colab={"base_uri": "https://localhost:8080/"} outputId="5023f7a2-ac47-46d6-9381-1ded05f9eb3d"
# tutaj wpisz rozwiązanie
A = np.arange(12).reshape(3, -1)
A[::-1]
# + [markdown] id="QwFXH2dIEhMp"
# * zamiana kolumn (odwrócona kolejność)
#
#
# ```
# array([[ 3, 2, 1, 0],
# [ 7, 6, 5, 4],
# [11, 10, 9, 8]])
# ```
#
# + id="dux6cn8YV-q_" colab={"base_uri": "https://localhost:8080/"} outputId="4dd35c26-642f-4395-b1a2-607465246350"
# tutaj wpisz rozwiązanie
A[:, ::-1]
# + [markdown] id="i5mm0dexEoDe"
# * zamiana wierszy i kolumn (odwrócone kolejności)
#
#
# ```
# array([[11, 10, 9, 8],
# [ 7, 6, 5, 4],
# [ 3, 2, 1, 0]])
# ```
# + id="ZqkvfYoKWHLp" colab={"base_uri": "https://localhost:8080/"} outputId="9ab1e47a-6613-41ea-f616-ea78c36015f2"
# tutaj wpisz rozwiązanie
A[::-1, ::-1]
# + [markdown] id="o3HKIKlP15Ro"
# ### <a name='5'></a> Ćwiczenie 25
# Podaną poniżej tablicę numpy:
#
# ```
# array([[1., 1., 1., 1.],
# [1., 1., 1., 1.],
# [1., 1., 1., 1.],
# [1., 1., 1., 1.]])
# ```
# Przekształć na tablicę:
#
#
# ```
# array([[0., 0., 0., 0., 0., 0.],
# [0., 1., 1., 1., 1., 0.],
# [0., 1., 1., 1., 1., 0.],
# [0., 1., 1., 1., 1., 0.],
# [0., 1., 1., 1., 1., 0.],
# [0., 0., 0., 0., 0., 0.]])
# ```
# __Wskazówka:__ Użyj funkcji _np.pad()_.
#
#
#
#
# + id="enLY_Qz4qGdJ" colab={"base_uri": "https://localhost:8080/"} outputId="ae0d522f-b8fc-4e90-d1a8-ab3ef5ec7635"
# tutaj wpisz rozwiązanie
A = np.ones(shape=(4, 4))
np.pad(A, pad_width=1, constant_values=0)#pad_with-dodanie kolumn po brzegach i wierszy
# + [markdown] id="PunXd_Ey16so"
# ### <a name='6'></a> Ćwiczenie 26
# Podaną poniżej tablicę numpy:
#
# ```
# array([[0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 0, 0],
# [0, 0, 0, 0, 0, 0]])
# ```
# Przekształć na tablicę:
#
#
# ```
# array([[10, 0, 10, 0, 10, 0],
# [ 5, 0, 5, 0, 5, 0],
# [10, 0, 10, 0, 10, 0],
# [ 5, 0, 5, 0, 5, 0],
# [10, 0, 10, 0, 10, 0],
# [ 5, 0, 5, 0, 5, 0]])
# ```
# __Wskazówka:__ Użyj operatora wycinania oraz przypisz odpowiednie wartości.
#
#
#
#
# + id="xOVKpjvMr7bX" colab={"base_uri": "https://localhost:8080/"} outputId="b295912a-42eb-4971-ef87-5ff29eafd807"
# tutaj wpisz rozwiązanie
A = np.zeros(shape=(6, 6), dtype='int')
A[::2, ::2] = 10 #wycięcie co 2 elementu z wiersza i z kolumny
A[1::2, ::2] #wycinanie od indeksu 1 co 2 element oraz co 2 kolumna
A
# + [markdown] id="hOhqDRz618Tx"
# ### <a name='7'></a> Ćwiczenie 27
# Połącz podane poniżej tablice $A$ oraz $B$ w jedną.
#
#
# ```
# A = np.arange(12).reshape(-1, 4)
# B = np.array([[4, 3, 7, 2],
# [0, 5, 2, 6]])
# ```
#
#
#
#
# __Oczekiwany rezultat:__
#
#
# ```
# array([[ 0, 1, 2, 3],
# [ 4, 5, 6, 7],
# [ 8, 9, 10, 11],
# [ 4, 3, 7, 2],
# [ 0, 5, 2, 6]])
# ```
# __Wskazówka:__ Użyj funkcji _np.append()_.
#
# + id="CCHfKU4Y3Z-X" colab={"base_uri": "https://localhost:8080/"} outputId="9e16dad5-9694-4011-8e80-cdf6e5d1193a"
# tutaj wpisz rozwiązanie
A = np.arange(12).reshape(-1, 4)
B = np.array([[4, 3, 7, 2],
[0, 5, 2, 6]])
np.append(A, B, axis=0)
# + [markdown] id="TXUd_4631-gw"
# ### <a name='8'></a> Ćwiczenie 28
# Z podanych poniżej tablic $A$ i $B$ wydobądź elementy wspólne (przecięcie) tablic.
#
# ```
# A = np.arange(8).reshape(-1, 4)
# B = np.array([[9, 10, 11, 3],
# [2, 8, 0, 9]])
# ```
#
#
#
# __Wskazówka:__ Użyj funkcji _np.intersect1d()_.
# + id="pgvyad4UCI6Q" colab={"base_uri": "https://localhost:8080/"} outputId="5a907272-860f-4d30-b5ab-42da039ca855"
# tutaj wpisz rozwiązanie
A = np.arange(8).reshape(-1, 4)
B = np.array([[9, 10, 11, 3],
[2, 8, 0, 9]])
np.intersect1d(A, B)
# + [markdown] id="tgI6xEn-2AQo"
# ### <a name='9'></a> Ćwiczenie 29
# Podana jest poniższa tablica numpy $A$.
# Znajdź unikalne wartości tej tablicy.
#
#
# ```
# A = np.array([[5, 1, 2, 1, 2],
# [9, 1, 9, 7, 5],
# [4, 1, 5, 7, 9]])
# ```
#
#
# __Wskazówka:__ Użyj funkcji _np.unique()_.
# + id="h3Q6RADIELkH" colab={"base_uri": "https://localhost:8080/"} outputId="c1d4b2bc-906e-433a-d453-9bab2d962897"
# tutaj wpisz rozwiązanie
A = np.array([[5, 1, 2, 1, 2],
[9, 1, 9, 7, 5],
[4, 1, 5, 7, 9]])
np.unique(A)
# + [markdown] id="wxhpf-uq2B1I"
# ### <a name='10'></a> Ćwiczenie 30
# Podana jest poniższa tablica:
#
#
# ```
# A = np.array([[0.4, 0.3, 0.3],
# [0.1, 0.1, 0.8],
# [0.2, 0.5, 0.3]])
# ```
#
#
#
# Zwróć indeksy na których stoją maksymalne wartości dla danego wiersza.
#
# __Wskazówka:__ Użyj funkcji _np.argmax()_.
# + id="YcQFRYUyFWbY" colab={"base_uri": "https://localhost:8080/"} outputId="b35a618c-abcb-48e1-bc7c-a54633aab3cb"
# tutaj wpisz rozwiązanie
A = np.array([[0.4, 0.3, 0.3],
[0.1, 0.1, 0.8],
[0.2, 0.5, 0.3]])
np.argmax(A, axis=1)
|
01_numpy_cwiczenia/021_030_exercises.ipynb
|
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # ML.Net - StopWords
// ## <NAME> -> Cientista de Dados 👋
// (<EMAIL>)
//
// [](https://www.linkedin.com/in/davi-ramos/)
// [](https://twitter.com/Daviinfo/)
// <a href="https://github.com/DaviRamos"><img src="https://img.shields.io/github/followers/DaviRamos.svg?label=GitHub&style=social" alt="GitHub"></a>
// ML.NET Nuget packages installation
//#r "nuget:Microsoft.ML,1.3.1"
#r "nuget:Microsoft.ML"
// ## Using C# Class
using Microsoft.ML;
using Microsoft.ML.Data;
using System;
using System.Collections.Generic;
using System.Text;
// ## Declare data-classes for input data and predictions
// +
public class TextData
{
public string Text { get; set; }
}
public class TextTokens
{
public string[] Tokens { get; set; }
}
// -
// ## Função Auxiliar para Imprimir os Tokens
private static void PrintTokens(TextTokens tokens)
{
Console.WriteLine(Environment.NewLine);
var sb = new StringBuilder();
foreach (var token in tokens.Tokens)
{
sb.AppendLine(token);
}
Console.WriteLine(sb.ToString());
}
// +
var context = new MLContext();
var emptyData = new List<TextData>();
var data = context.Data.LoadFromEnumerable(emptyData);
var tokenization = context.Transforms.Text.TokenizeIntoWords("Tokens", "Text", separators: new[] { ' ', '.', ',' })
.Append(context.Transforms.Text.RemoveDefaultStopWords("Tokens", "Tokens",
Microsoft.ML.Transforms.Text.StopWordsRemovingEstimator.Language.English));
var stopWordsModel = tokenization.Fit(data);
var engine = context.Model.CreatePredictionEngine<TextData, TextTokens>(stopWordsModel);
var newText = engine.Predict(new TextData { Text = "This is a test sentence, and it is a long one." });
PrintTokens(newText);
var customTokenization = context.Transforms.Text.TokenizeIntoWords("Tokens", "Text", separators: new[] { ' ', '.', ',' })
.Append(context.Transforms.Text.RemoveStopWords("Tokens", "Tokens", new[] { "and", "a" }));
var customStopWordsModel = customTokenization.Fit(data);
var customEngine = context.Model.CreatePredictionEngine<TextData, TextTokens>(customStopWordsModel);
var newCustomText = customEngine.Predict(new TextData { Text = "This is a test sentence, and it is a long one." });
PrintTokens(newCustomText);
Console.ReadLine();
// -
|
Notebooks/ML.Net - StopWords.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="pMHdlTffbT2X"
# # Week 4 Lecture 1
#
# ## Determinants
#
# We did examples of $2\times 2$ determinants in the theory lecture. Let's take a look at an example of a larger determinant.
#
# Consider the matrix $A = \begin{bmatrix}1 & 4 & -1\\ 3 & 2 & 0\\ -1 & 4 & 3 \end{bmatrix}$. We can take the determinant by doing
# + colab={"base_uri": "https://localhost:8080/"} id="cqLDcVrjbWiX" outputId="a590e953-2c27-4c0a-ad8e-ba51a9b8d04c"
import numpy as np
import scipy.linalg as la
A = np.array([[1, 4, -1], [3, 2, 0], [-1, 4, 3]])
print(la.det(A))
# + [markdown] id="p-x3CfbPhqc-"
# We can do this with another matrix $B = \begin{bmatrix} 2 & 4 & 6\\ 0 & 3 & 1\\ 0 & 0 & -5 \end{bmatrix}$
# + colab={"base_uri": "https://localhost:8080/"} id="389iL4VMhdTF" outputId="7baedae2-3486-4b6c-a843-5b43d9607ed8"
B = np.array([[2, 4, 6], [0, 3, 1], [0, 0, -5]])
print(la.det(B))
# + [markdown] id="0RElBrd7h40c"
# Notice that this is just the product of the diagonal. In any triangular matrix the determinant will be the product of its diagonal.
# + [markdown] id="deqf-hPYFIhb"
# ## Eigenvalues and Eigenvectors
#
# In the last lecture, we discussed a class of iterative methods for solving the system $A\mathbf{x} = \mathbf{b}$. In particular, we found an equation for the error of our estimated solutions:
#
# $\mathbf{e}_k = M^k\mathbf{e}_0$.
#
# We saw before that this equation is straightforward if $M$ is just a number. We would like to use a similar method with the matrix version, but to do so we need to know when and how we are allowed to treat matrix multiplication like scalar multiplication. In particular, we would like to know when we are allowed to write
#
# $M\mathbf{x} = \lambda\mathbf{x},\hspace{1in}$ (1)
#
# where $M$ is a matrix, $\mathbf{x}$ is a vector and $\lambda$ is a number.
#
# The notation here is a bit confusing. We are ultimately interested in using the matrix $M$ from last lecture in equation (1), but the equation is perfectly valid for *any* square matrix $M$, not just that particular one. In addition, the vector $\mathbf{x}$ in (1) has nothing to do with the solution to our original system $A\mathbf{x} = \mathbf{b}$. You should think of equation (1) as follows: You are given some fixed matrix $M$ (it can be any square matrix - can you see why it has to be square?) and you are looking for a vector $\mathbf{x}$ and a number $\lambda$ such that multiplying $\mathbf{x}$ by $M$ is the same as multiplying $\mathbf{x}$ by the number $\lambda$. Since $\mathbf{x} = \mathbf{0}$ (and any value of $\lambda$) is an obvious solution to this problem for any matrix $M$, we will only care about cases where $\mathbf{x} \neq \mathbf{0}$. (It is still possible for $\lambda = 0$ even if $\mathbf{x} \neq \mathbf{0}$, and we are still interested in these solutions.)
#
# The number $\lambda$ in this equation is called an *eigenvalue* of $M$ and the vector $\mathbf{x}$ is called an *eigenvector* of $M$ corresponding to $\lambda$.
#
# Let's look at a few simple examples. Let
#
# $M = \begin{pmatrix} 1 & 2 \\ 0 & 3 \end{pmatrix}$ and $\mathbf{x} = \begin{pmatrix} 1 \\ 1 \end{pmatrix}$.
#
# We have
#
# $M\mathbf{x} = \begin{pmatrix} 1 & 2 \\ 0 & 3 \end{pmatrix}\begin{pmatrix} 1 \\ 1 \end{pmatrix} = \begin{pmatrix} 3 \\ 3 \end{pmatrix} = 3\mathbf{x}$.
#
# This means that 3 is an eigenvalue of $M$ with corresponding eigenvector $[1, 1]^{T}$. (Remember that the $T$ stands for transpose and just means "make this a column vector.")
#
# Similarly, if we keep the same matrix $M$ but let
#
# $\mathbf{x} = \begin{pmatrix} 10 \\ 10 \end{pmatrix}$,
#
# then we have
#
# $M\mathbf{x} = \begin{pmatrix} 1 & 2 \\ 0 & 3 \end{pmatrix}\begin{pmatrix} 10 \\ 10 \end{pmatrix} = \begin{pmatrix} 30 \\ 30 \end{pmatrix} = 3\mathbf{x}$,
#
# so $[10, 10]^{T}$ is also an eigenvector corresponding to the eigenvalue 3. In fact, it is easy to check that any multiple of $[1, 1]^{T}$ is also an eigenvector with corresponding eigenvalue 3.
#
# However, if we instead let
#
# $\mathbf{x} = \begin{pmatrix} 0 \\ 1 \end{pmatrix}$,
#
# then we have
#
# $M\mathbf{x} = \begin{pmatrix} 1 & 2 \\ 0 & 3 \end{pmatrix}\begin{pmatrix} 0 \\ 1 \end{pmatrix} = \begin{pmatrix} 2 \\ 3 \end{pmatrix}$.
#
# This cannot possibly be written as $\lambda \mathbf{x}$ because any multiple of $\mathbf{x}$ would have a zero in the first entry. That means that $[0, 1]^{T}$ is *not* an eigenvector of $M$.
#
# It turns out (and you should experiment until you are convinced that this is true) that most vectors are not eigenvectors of $M$. The only possible eigenvectors are multiples of $[1, 1]^T$ and multiples of $[1, 0]^T$. Moreover, there are only two eigenvalues of $M$. We already saw that 3 is an eigenvalue of $M$ with corresponding eigenvectors that are multiples of $[1, 1]^T$. It is also easy to check that 1 is an eigenvalue of $M$ with corresponding eigenvectors that are multiples of $[1, 0]^T$. Notice that $M$ was a $2\times 2$ matrix and we found 2 eigenvalues, each with a family of corresponding eigenvectors. (By "family", I mean that all the eigenvectors were multiples of each other. The word I should really use is "subspace", but we won't go into the technical details of subspaces in this class.) This pattern is not a coincidence. It turns out (although we will not prove it) that *every* $n\times n$ matrix has $n$ eigenvalues, and almost every $n\times n$ matrix has $n$ different families of eigenvectors.
#
# There are a couple caveats about this theorem that are worth remembering. First, the rule that there are $n$ eigenvalues of an $n\times n$ matrix works just like the rule that there are $n$ solutions to an $n$th degree polynomial, which you should already be familiar with from a basic algebra class. In particular, it is possible for eigenvalues to be complex, and it is possible for the same eigenvalue to be repeated multiple times. If one of the eigenvalues is complex, then its corresponding eigenvectors can also be written with complex entries. If an eigenvalue is repeated $k$ times, then it will (almost always) have $k$ different families of eigenvectors. It is actually possible for repeated eigenvalues to share the same family of eigenvectors (which is where the "almost" in our theorem comes from). We won't worry about that issue in our class, because it is quite rare and makes the analysis much more complicated.
#
# The extra videos (available under "Additional Resources" in the canvas modules) show how to find eigenvalues and eigenvectors of a $2\times 2$ matrix by hand. It is also possible to use that method for $3\times 3$ matrices (but it is much more difficult). For anything larger than a $3\times 3$ matrix it is not usually possible to find eigenvalues/eigenvectors by hand. Since we are primarily interested in large matrices, we will not worry about finding these values on paper. Instead, we will let python do the hard work for us.
# + [markdown] id="vQWqPE-6FIht"
# In python, we can find the eigenvalues and eigenvectors of a matrix $M$ with the `eig` function, which is located in the `numpy.linalg` package. The `eig` function takes a square matrix $M$ as an argument and returns two things: A 1D array containing all of the eigenvalues of $M$ and a matrix containing all of the eigenvectors of $M$. For example,
# + id="k7fHW64bFIhu" outputId="e999b618-65df-4c48-e812-81cfdefad9b4"
import numpy as np
M = np.array([[1, 2], [0, 3]])
w, V = np.linalg.eig(M)
print(w)
# + id="A-xaBj1nFIhv" outputId="1a21d0f2-b299-4475-d690-6edc14d8a781"
print(V)
# + [markdown] id="0dBF4AVTFIhw"
# The array `w` is easy to interpret. Each entry is an eigenvalue of $M$. The matrix `V` is slightly more complicated. Each column of V is an eigenvector of $M$, and the order of V matches the order of w. That is, the first column of V is an eigenvector corresponding to the first entry of w and the second column of V is an eigenvector corresponding to the second entry of w. In general, the column `V[;, i]` is an eigenvector corresponding to the eigenvalue `w[i]`. (**Warning:** The syntax `V[:, i]` pulls out the entries from the `i`th column of V, but it makes a 1D array instead of a column vector. If you want a column vector, then you will have to reshape this array.)
#
# Python does not just use this format for V because it's a convenient way to store all of the eigenvectors; the matrix $V$ has some important mathematical significance. Before we can see it, though, we need to rearrange the eigenvalues slightly. Instead of storing the eigenvalues in a 1D array, it is often more mathematically useful to make a diagonal matrix with the eigenvalues along the main diagonal (and the rest of the entries zero). This can be done in python with the `diag` function, which is part of the `numpy` package.
# + id="A_Xo_9bYFIhw" outputId="014f6bd4-543f-4b64-aa02-c48a3a68cf90"
D = np.diag(w)
print(D)
# + [markdown] id="8tiwFb-MFIhx"
# There is an important mathematical relationship between $M$, $V$ and $D$. In particular, it turns out that we can rewrite $M$ as follows:
#
# $M = VDV^{-1}$.
#
# (We will verify this in python but not prove it.) This formula is called an *eigendecomposition* of $M$. You can check the formula in python with the code
# + id="NNJwXvDhFIhx" outputId="4a76d7d8-7bfe-4a92-aa4d-c22621f17b44"
print(M)
# + id="p99nEYt4FIhy" outputId="c69c89b1-c647-4893-c860-0d7dee8dfca3"
print(V @ D @ scipy.linalg.inv(V))
# + [markdown] id="bYASmEeiLwXE"
# Now lets look at the exmaple from the theory lecture.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="Q-KrN4lhMwRq" outputId="0a6e0ced-2d87-4d72-bd46-bb2b55cd0d74"
import numpy as np
import matplotlib.pyplot as plt
A = np.array([[4, 1], [3, 2]])
w, V = np.linalg.eig (A)
v1 = V[:,0]
v2 = V[:,1]
v1_x = np.arange(0, v1[0] + v1[0]/100, v1[0]/100)
v1_y = np.arange(0, v1[1] + v1[1]/100, v1[1]/100)
v2_x = np.arange(0, v2[0] + v2[0]/100, v2[0]/100)
v2_y = np.arange(0, v2[1] + v2[1]/100, v2[1]/100)
plt.subplot(2,2,1)
plt.plot(v1_x, v1_y, linewidth = 4)
plt.plot(v2_x, v2_y, linewidth = 4)
v1 = A@v1
v2 = A@v2
v1_x = np.arange(0, v1[0] + v1[0]/100, v1[0]/100)
v1_y = np.arange(0, v1[1] + v1[1]/100, v1[1]/100)
v2_x = np.arange(0, v2[0] + v2[0]/100, v2[0]/100)
v2_y = np.arange(0, v2[1] + v2[1]/100, v2[1]/100)
plt.subplot(2,2,2)
plt.plot(v1_x, v1_y, linestyle = "--", linewidth = 4)
plt.plot(v2_x, v2_y, linestyle = "--", linewidth = 4)
v = np.random.randn(2,1)
v_x = np.arange(0, v[0] + v[0]/100, v[0]/100)
v_y = np.arange(0, v[1] + v[1]/100, v[1]/100)
plt.subplot(2,2,3)
plt.plot(v_x, v_y, linewidth = 4)
v = A@v
v_x = np.arange(0, v[0] + v[0]/100, v[0]/100)
v_y = np.arange(0, v[1] + v[1]/100, v[1]/100)
plt.subplot(2,2,4)
plt.plot(v_x, v_y, linestyle = "--", linewidth = 4)
# + [markdown] id="Hj74pW1hFIhy"
# ## More about eigendecomposition
#
# It is worth noting that the eigenvectors and eigenvalues of a matrix might be complex, so there might be lots of imaginary numbers on the right side of the above equation. We won't worry about these complex numbers for two reasons: First, they all cancel out when you multiply $VDV^{-1}$, so there won't be any complex numbers in our final answers. Second, we will ultimately only care about the magnitude of eigenvalues, so we will only really work with real numbers. (Remember, the magnitude of a complex number $a + bi$ is $\sqrt{a^2 + b^2}$. You can use the python command `np.abs()` to find the magnitude of a complex number.)
#
# The eigendecomposition is particularly useful for computing powers of $M$. To see this, notice that
#
# $M^2 = MM = \left(VDV^{-1}\right)\left(VDV^{-1}\right) = VDV^{-1}VDV^{-1} = VDDV^{-1} = VD^2V^{-1}$,
#
# because the innermost $V$ and $V^{-1}$ cancel out. (It is important to remember that you cannot reorder matrix multiplication, so we are only allowed to cancel a $V^{-1}$ and $V$ if they appear beside each other.)
#
# Similarly,
#
# $M^3 = M^2M = \left(VD^2V^{-1}\right)\left(VDV^{-1}\right) = VD^2V^{-1}VDV^{-1} = VD^2DV^{-1} = VD^3V^{-1}$.
#
# The pattern should quickly become apparent. In general, we have
#
# $M^k = VD^kV^{-1}$.
#
# The reason this is so convenient is that $D$ is a diagonal matrix and it is easy to compute powers of a diagonal matrix: We just raise each entry of the matrix to the same power. For example,
#
# $\left(\begin{array}{c} 2 & 0 \\ 0 & 1/3 \end{array}\right)^k = \left(\begin{array}{c} 2^k & 0 \\ 0 & (1/3)^k \end{array}\right)$.
|
determinants, eigenvalues and eigenvectors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/linked0/deep-learning/blob/master/AAMY/Cats_vs_Dogs.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="1RZAgiODbaD8" colab_type="code" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# + id="J37KEK7rdBIZ" colab_type="code" colab={}
try:
# Use the %tensorflow_version magic if in colab.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + id="keWET94NeDSf" colab_type="code" colab={}
import os
import matplotlib.pyplot as plt
import numpy as np
# + id="yjf_o7wjetq8" colab_type="code" colab={}
import logging
logger = tf.get_logger()
logger.setLevel(logging.ERROR)
# + id="37tSNtyie1MI" colab_type="code" colab={}
_URL = 'https://storage.googleapis.com/mledu-datasets/cats_and_dogs_filtered.zip'
zip_dir = tf.keras.utils.get_file('cats_and_dogs_filterted.zip', origin=_URL, extract=True)
# + id="pDMZOWJQfNrZ" colab_type="code" outputId="01d93682-5a20-4f30-b357-7c5f4b1ce4ca" colab={"base_uri": "https://localhost:8080/", "height": 169}
zip_dir_base = os.path.dirname(zip_dir)
# !find $zip_dir_base -type d -print
# + id="U1iTSKDFftGW" colab_type="code" colab={}
base_dir = os.path.join(os.path.dirname(zip_dir), 'cats_and_dogs_filtered')
train_dir = os.path.join(base_dir, 'train')
validation_dir = os.path.join(base_dir, 'validation')
train_cats_dir = os.path.join(train_dir, 'cats')
train_dogs_dir = os.path.join(train_dir, 'dogs')
validation_cats_dir = os.path.join(validation_dir, 'cats')
validation_dogs_dir = os.path.join(validation_dir, 'dogs')
# + id="M4l21wxMgdo4" colab_type="code" colab={}
num_cats_tr = len(os.listdir(train_cats_dir))
num_dogs_tr = len(os.listdir(train_dogs_dir))
num_cats_val = len(os.listdir(validation_cats_dir))
num_dogs_val = len(os.listdir(validation_dogs_dir))
total_train = num_cats_tr + num_dogs_tr
total_val = num_cats_val + num_dogs_val
# + id="4F78wCsait4e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 150} outputId="32ca28a8-f107-49ae-ebed-b696dc8b9e8b"
print('total training cat images:', num_cats_tr)
print('total training dog images:', num_dogs_tr)
print('total validation cat images:', num_cats_val)
print('total validation dog images:', num_dogs_val)
print("--")
print("Total training images:", total_train)
print("Total validation images:", total_val)
# + id="AcSLSLRyiVu4" colab_type="code" colab={}
BATCH_SIZE = 100
IMG_SHAPE = 150
# + id="eQQERtj5jCru" colab_type="code" colab={}
from tensorflow.keras.preprocessing.image import ImageDataGenerator
# + id="U8eUoeO4i0NG" colab_type="code" colab={}
train_image_generator = ImageDataGenerator(rescale=1./255)
validation_image_generator = ImageDataGenerator(rescale=1./255)
# + id="rPlzBRSSjJ3J" colab_type="code" colab={}
# + id="RNbomGFWjMSN" colab_type="code" colab={}
|
AAMY/Cats_vs_Dogs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Lab 04 : Test set evaluation -- demo
# For Google Colaboratory
import sys, os
if 'google.colab' in sys.modules:
# mount google drive
from google.colab import drive
drive.mount('/content/gdrive')
path_to_file = '/content/gdrive/My Drive/CS4243_codes/codes/labs_lecture05/lab04_test_set'
print(path_to_file)
# move to Google Drive directory
os.chdir(path_to_file)
# !pwd
import torch
import torch.nn as nn
import torch.optim as optim
from random import randint
import utils
# ### Download the data and print the sizes
# +
from utils import check_mnist_dataset_exists
data_path=check_mnist_dataset_exists()
train_data=torch.load(data_path+'mnist/train_data.pt')
train_label=torch.load(data_path+'mnist/train_label.pt')
test_data=torch.load(data_path+'mnist/test_data.pt')
test_label=torch.load(data_path+'mnist/test_label.pt')
# -
# ### Make a ONE layer net class.
class one_layer_net(nn.Module):
def __init__(self, input_size, output_size):
super(one_layer_net , self).__init__()
# 一层全连接网络MLP
self.linear_layer = nn.Linear( input_size, output_size , bias=False)
def forward(self, x):
# 直接输出
scores = self.linear_layer(x)
return scores
# ### Build the net
net=one_layer_net(784,10)
print(net)
# ### Choose the criterion, optimizer, batchsize, learning rate
# +
criterion = nn.CrossEntropyLoss()
optimizer=torch.optim.SGD( net.parameters() , lr=0.01 )
bs=200
# -
# ### Do 30 passes through the training set
# +
for epoch in range(30):
running_loss=0
running_error=0
num_batches=0
shuffled_indices=torch.randperm(60000)
for count in range(0,60000,bs):
optimizer.zero_grad()
indices=shuffled_indices[count:count+bs]
minibatch_data = train_data[indices]
minibatch_label= train_label[indices]
inputs = minibatch_data.view(bs,784)
inputs.requires_grad_()
scores=net( inputs )
loss = criterion( scores , minibatch_label)
loss.backward()
optimizer.step()
# compute and accumulate stats
running_loss += loss.detach().item()
error = utils.get_error( scores.detach() , minibatch_label)
running_error += error.item()
num_batches+=1
# compute stats for the full training set
total_loss = running_loss/num_batches
total_error = running_error/num_batches
print('epoch=',epoch, '\t loss=', total_loss , '\t error=', total_error*100 ,'percent')
# -
# ### Now that the network is trained and do 10% of error on the training set, we are going to see how well it is doing on the test set...
# +
running_error=0
num_batches=0
for i in range(0,10000,bs):
# extract the minibatch
minibatch_data = test_data[i:i+bs]
minibatch_label= test_label[i:i+bs]
# reshape the minibatch
inputs = minibatch_data.view(bs,784)
# feed it to the network
scores=net( inputs )
# compute the error made on this batch
error = utils.get_error( scores , minibatch_label)
# add it to the running error
running_error += error.item()
num_batches+=1
# compute error rate on the full test set
total_error = running_error/num_batches
print( 'error rate on test set =', total_error*100 ,'percent')
# -
# ### Choose image at random from the test set and see how good/bad are the predictions
# +
# choose a picture at random
idx=randint(0, 10000-1)
im=test_data[idx]
# diplay the picture
utils.show(im)
# feed it to the net and display the confidence scores
scores = net( im.view(1,784))
probs= torch.softmax(scores, dim=1)
utils.show_prob_mnist(probs)
# -
|
codes/labs_lecture05/lab04_test_set/test_set_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Encoding the Board
def cross(a, b):
return [s+t for s in a for t in b]
def extract_units(unitlist, boxes):
"""Initialize a mapping from box names to the units that the boxes belong to
Parameters
----------
unitlist(list)
a list containing "units" (rows, columns, diagonals, etc.) of boxes
boxes(list)
a list of strings identifying each box on a sudoku board (e.g., "A1", "C7", etc.)
Returns
-------
dict
a dictionary with a key for each box (string) whose value is a list
containing the units that the box belongs to (i.e., the "member units")
"""
for current_box in boxes:
for unit in unitlist:
if current_box in unit:
# defaultdict avoids this raising a KeyError when new keys are added
units[current_box].append(unit)
return units
def extract_peers(units, boxes):
"""Initialize a mapping from box names to a list of peer boxes (i.e., a flat list
of boxes that are in a unit together with the key box)
Parameters
----------
units(dict)
a dictionary with a key for each box (string) whose value is a list
containing the units that the box belongs to (i.e., the "member units")
boxes(list)
a list of strings identifying each box on a sudoku board (e.g., "A1", "C7", etc.)
Returns
-------
dict
a dictionary with a key for each box (string) whose value is a set
containing all boxes that are peers of the key box (boxes that are in a unit
together with the key box)
"""
# the value for keys that aren't in the dictionary are initialized as an empty list
for key_box in boxes:
for unit in units[key_box]:
for peer_box in unit:
if peer_box != key_box:
# defaultdict avoids this raising a KeyError when new keys are added
peers[key_box].add(peer_box)
return peers
# +
rows = 'ABCDEFGHI'
cols = '123456789'
boxes = cross(rows, cols)
row_units = [cross(r, cols) for r in rows]
column_units = [cross(rows, c) for c in cols]
square_units = [cross(rs, cs) for rs in ('ABC', 'DEF', 'GHI') for cs in ('123', '456', '789')]
unitlist = row_units + column_units + square_units
# TODO: Update the unit list to add the new diagonal units
unitlist = unitlist + [[a+b for a,b in zip(rows, cols[::-1])]] + [[a+b for a,b in zip(rows, cols)]]
units = extract_units(unitlist, boxes)
peers = extract_peers(units, boxes)
print(unitlist)
# -
def grid_values(grid):
"""Convert grid string into {<box>: <value>} dict with '.' value for empties.
Args:
grid: Sudoku grid in string form, 81 characters long
Returns:
Sudoku grid in dictionary form:
- keys: Box labels, e.g. 'A1'
- values: Value in corresponding box, e.g. '8', or '123456789' if it is empty.
"""
values = []
all_digits = '123456789'
for c in grid:
if c == '.':
values.append(all_digits)
elif c in all_digits:
values.append(c)
assert len(grid) == 81
return dict(zip(boxes, values))
def display(values):
"""
Display the values as a 2-D grid.
Input: The sudoku in dictionary form
Output: None
"""
width = 1+max(len(values[s]) for s in boxes)
line = '+'.join(['-'*(width*3)]*3)
for r in rows:
print(''.join(values[r+c].center(width)+('|' if c in '36' else '')
for c in cols))
if r in 'CF': print(line)
return
grid = '..3.2.6..9..3.5..1..18.64....81.29..7.......8..67.82....26.95..8..2.3..9..5.1.3..'
values = grid_values(grid)
display(values)
# # Eliminate the impossible solution in each grid
def eliminate(values):
"""Eliminate values from peers of each box with a single value.
Go through all the boxes, and whenever there is a box with a single value,
eliminate this value from the set of values of all its peers.
Args:
values: Sudoku in dictionary form.
Returns:
Resulting Sudoku in dictionary form after eliminating values.
"""
solved_values = [box for box in values.keys() if len(values[box]) == 1]
for box in solved_values:
digit = values[box]
for peer in peers[box]:
values[peer] = values[peer].replace(digit, '')
return values
display(eliminate(values))
# # Only Choice:
# If there is only one box in a unit which would allow a certain digit, then that box must be assigned that digit.
def only_choice(values):
"""Finalize all values that are the only choice for a unit.
Go through all the units, and whenever there is a unit with a value
that only fits in one box, assign the value to this box.
Input: Sudoku in dictionary form.
Output: Resulting Sudoku in dictionary form after filling in only choices.
"""
for unit in unitlist:
for digit in '123456789':
dplaces = [box for box in unit if digit in values[box]]
if len(dplaces) == 1:
values[dplaces[0]] = digit
return values
display(only_choice(values))
# # Constraint Propagation
def reduce_puzzle(values):
"""
Iterate eliminate() and only_choice(). If at some point, there is a box with no available values, return False.
If the sudoku is solved, return the sudoku.
If after an iteration of both functions, the sudoku remains the same, return the sudoku.
Input: A sudoku in dictionary form.
Output: The resulting sudoku in dictionary form.
"""
stalled = False
while not stalled:
# Check how many boxes have a determined value
solved_values_before = len([box for box in values.keys() if len(values[box]) == 1])
# Your code here: Use the Eliminate Strategy
eliminate(values)
# Your code here: Use the Only Choice Strategy
only_choice(values)
# Check how many boxes have a determined value, to compare
solved_values_after = len([box for box in values.keys() if len(values[box]) == 1])
# If no new values were added, stop the loop.
stalled = solved_values_before == solved_values_after
# Sanity check, return False if there is a box with zero available values:
if len([box for box in values.keys() if len(values[box]) == 0]):
return False
return values
display(reduce_puzzle(values))
# # Harder Sudoku
grid2 = '4.....8.5.3..........7......2.....6.....8.4......1.......6.3.7.5..2.....1.4......'
values2 = grid_values(grid2)
display(reduce_puzzle(values2))
# # Search
def search(values):
# First, reduce the puzzle using the previous function
values = reduce_puzzle(values)
if values is False:
return False
if all(len(values[s]) == 1 for s in boxes):
return values
# Choose one of the unfilled squares with the fewest possibilities
n, s = min((len(values[s]), s) for s in boxes if len(values[s]) > 1)
# Now use recursion to solve each one of the resulting sudokus, and if one returns a value (not False), return that answer!
for value in values[s]:
new_sudoku = values.copy()
new_sudoku[s] = value
attempt = search(new_sudoku)
if attempt:
return attempt
display(search(values2))
|
Sudoku/Sudoku.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
tensor = tf.constant([1, 2, 3, 4, 5, 6, 7])
print(tensor)
tensor = tf.constant(-1.0, shape=[2, 3])
print(tensor)
tensor = tf.constant([1, 2, 3, 4, 5, 6, 7])
print(tensor)
node1 = tf.constant(3.0, tf.float32)
node2 = tf.constant(4.0)
print(node1, node2)
sess = tf.Session()
print(sess)
print(sess.run([node1, node2]))
node3 = tf.add(node1, node2)
print('node3:', node3)
print(sess.run(node3))
print(sess.run(node1 + node2))
# +
a = tf.placeholder(tf.float32)
b = tf.placeholder(tf.float32)
adder_node = a + b
# -
print(sess.run(adder_node, {a: 3, b: 4.5}))
print(sess.run(adder_node, {a: [1, 3], b: [2, 4]}))
add_and_triple = adder_node * 3
print(sess.run(add_and_triple, {a: 3, b: 4.5}))
sess.close()
# +
import tensorflow as tf
a = tf.constant(5, name='input_a')
b = tf.constant(3, name='input_b')
c = tf.multiply(a, b, name='multiply_c')
d = tf.add(a, b, name='add_d')
e = tf.add(c, d, name='add_e')
sess = tf.Session()
output = sess.run(e)
writer = tf.summary.FileWriter('./my_graph', sess.graph)
writer.close()
sess.close()
# -
W = tf.Variable([.3], tf.float32)
b = tf.Variable([-.3], tf.float32)
x = tf.placeholder(tf.float32)
linear_model = W * x + b
# +
sess = tf.Session()
init = tf.global_variables_initializer()
sess.run(init)
# -
print(sess.run(linear_model, {x:[1, 2, 3, 4]}))
y = tf.placeholder(tf.float32)
squared_deltas = tf.square(linear_model - y)
loss = tf.reduce_sum(squared_deltas)
print(sess.run(loss, {x:[1, 2, 3, 4], y:[0, -1, -2, -3]}))
fixW = tf.assign(W, [-1.])
fixb = tf.assign(b, [1.])
sess.run([fixW, fixb])
# [array([-1.], dtype = float32), array([1.], dtype = float32)]
print(sess.run(loss, {x:[1, 2, 3, 4], y:[0, -1, -2, -3]}))
optimizer = tf.train.GradientDescentOptimizer(0.01)
train = optimizer.minimize(loss)
# +
sess.run(init)
# reset values to incorrect defaults
for i in range(1000):
sess.run(train, {x: [1, 2, 3, 4], y: [0, -1, -2, -3]})
print(sess.run([W, b]))
# [array([-0.9999969], dtype = float32), array([0.99999082], dtype = float32)]
|
TF_CodingTech.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import os
data_dir = "/projectnb/llamagrp/feyzanb/data/turkish_data"
file_names = ["Posts from 2016-01-01 to 2016-07-07.xls",
"Posts from 2017-03-07 to 2017-10-08.xls",
"Posts from 2018-02-26 to 2018-06-29.xls",
"Posts from 2018-10-28 to 2018-12-31.xls",
"Posts from 2016-07-08 to 2017-03-06.xls",
"Posts from 2017-10-09 to 2018-02-25.xls",
"Posts from 2018-06-30 to 2018-10-27.xls"]
df = pd.read_excel(os.path.join(data_dir, "Posts from 2016-01-01 to 2016-07-07.xls"))\
.iloc[:,[0,17]]
df.columns = ["guid", "headline"]
for f in file_names[1:]:
df_temp = pd.read_excel(os.path.join(data_dir, f)).iloc[:,[0,17]]
df_temp.columns = ["guid", "headline"]
print(df.columns)
df = pd.concat([df, df_temp])
print(df.columns)
df.columns
len(df)
# +
search_for_1 = ["silahlı saldırı", "toplu saldırı", "saldırgan", "rastgele ateş",
"toplu silahlı saldırı", "silahlı saldırgan dehşet", "bireysel silahlanma",
"silah kontrolü", "toplu katliam", "silah yasası"]
search_for_2 = ["ABD", "Amerika", "Birleşik Devletler", "Trump"]
result_1 = df['headline'].str.contains('|'.join(search_for_1))
result_2 = df['headline'].str.contains('|'.join(search_for_2))
result = result_1 & result_2
print("Number of headlines including words related to gun violence: ", result_1.sum())
print("Number of headlines including words related to the US: ", result_2.sum())
print("Number of headlines that include at least one from each category: ", result.sum())
# +
# export filtered & shuffled data to csv file
data_filtered = df.loc[result, ['guid', 'headline']].sample(frac=1, random_state=13)
data_filtered.head()
# data_filtered_relevant = data.loc[pd.Series(data['Not Relevant'] == 0) & result,
# ['URL', 'German Headline', 'Not Relevant']].sample(frac=1, random_state=13)
# data_filtered.to_csv(r'german_filtered.csv')
# data_filtered_relevant.to_csv(r'german_filtered_relevant.csv')
# -
data_filtered.to_csv(r'turkish.csv')
|
dataset/arabic/arabic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 列表List
# - 一个列表可以储存任意大小的数据集合,你可以理解为他是一个容器
# ## 先来一个例子爽一爽
# 
# ## 创建一个列表
# - a = [1,2,3,4,5]
a3 = list('abcdef')
a3
a = [1,2,3]
[1] + a + [100]
a * 3
# ## 列表的一般操作
# 
# ## 列表索引操作
# - Mylist[index]
# - 正序索引,逆序索引
# - 列表一定注意越界
# - 
# ## 列表切片操作
# - Mylist[start:end]
# - 正序切片,逆序切片
i = 0
while i < len(a):
print(a[i])
i += 1
# if i >= 3:
# break
# ## 列表 +、*、in 、not in
# ## 使用for循环遍历元素
# - for 循环可以遍历一切可迭代元素
# ## EP:
# - 使用while 循环遍历列表
# ## 列表的比较
# - \>,<,>=,<=,==,!=
# ## 列表生成式
# [x for x in range(10)]
[x for x in range(10) ]
[x for x in range(10) if x % 2 == 0 ]
[x for x in range(10) if x % 2 == 0 ]
[ x if x%2 == 0 else 100 for x in range(10) ]
a = [x for x in range(5)]
a.append(5) # append修改原列表
a
# ## 列表的方法
# 
a1 = [1,2,3,4]
a1.insert(1,100)#插入
a1
a1.remove(2)#删除
a1
a1.reverse()#翻转
a1
aq = [0,-1,1,5,-6,-10]
aq.sort()#升序排列
aq
aq.sort(reverse = True)#降序排列
aq # *表示强制命名
a1 = [0,-1,2,3,3,4]
sorted(a1,reverse=True)# sorted不更改源数据
list_ = [['困的黁',18],['你看',22],['男1',2],['男2',0]]
sorted(list_,key = lambda x : x[1])
list_ = [['高风艳',80,10000],['皮卡丘',22,100],['男1',2,20000],['男2',0,-99]]
sorted(list_,key=lambda x:x[2])
# +
# def B2():
# print('哈哈哈')
# def B1():
# func()
# B1(B2)
# -
import matplotlib.pyplot as plt
plt.plot([1,2,3],[1,2,3])
import matplotlib.pyplot as plt
x = []
y = []
for i in range(-10,11):
x.append(i)
y.append(i ** 2)
plt.plot(x,y)
import matplotlib.pyplot as plt
x = []
y = []
for i in range(-10,11):
x.append(i)
y.append(i ** 2)
plt.plot(x,y,'r--')
import matplotlib.pyplot as plt
x = []
y = []
for i in range(-10,11):
x.append(i)
y.append(i ** 2)
plt.plot(x,y,'r--')
plt.plot(x,y)
# +
# coding:utf-8
__author__ = 'taohao'
import matplotlib.pyplot as plt
from matplotlib import animation
import numpy as np
import math
def drawHeart():
t = np.linspace(0, math.pi, 1000)
x = np.sin(t)
y = np.cos(t) + np.power(x, 2.0/3)
plt.plot(x, y, color='red', linewidth=2, label='h')
plt.plot(-x, y, color='red', linewidth=2, label='-h')
plt.xlabel('t')
plt.ylabel('h')
plt.ylim(-2, 2)
plt.xlim(-2, 2)
plt.legend()
plt.show()
drawHeart()
# -
from turtle import *
pensize(1)
pencolor('red')
fillcolor('pink')
speed(5)
up()
goto(-30, 100)
down()
begin_fill()
left(90)
circle(120,180)
circle(360,70)
left(38)
circle(360,70)
circle(120,180)
end_fill()
up()
goto(-100,-100)
down()
# ## 将字符串分割成列表
# - split 按照自定义的内容拆分
sum([1,2,3,4,5])
# ## EP:
# 
# 
# coding:utf-8
import turtle as t
t.pensize(4) # 设置画笔的大小
t.colormode(255) # 设置GBK颜色范围为0-255
t.color((255,155,192),"pink") # 设置画笔颜色和填充颜色(pink)
t.setup(840,500) # 设置主窗口的大小为840*500
t.speed(10) # 设置画笔速度为10
#鼻子
t.pu() # 提笔
t.goto(-100,100) # 画笔前往坐标(-100,100)
t.pd() # 下笔
t.seth(-30) # 笔的角度为-30°
t.begin_fill() # 外形填充的开始标志
a=0.4
for i in range(120):
if 0<=i<30 or 60<=i<90:
a=a+0.08
t.lt(3) #向左转3度
t.fd(a) #向前走a的步长
else:
a=a-0.08
t.lt(3)
t.fd(a)
t.end_fill() # 依据轮廓填充
t.pu() # 提笔
t.seth(90) # 笔的角度为90度
t.fd(25) # 向前移动25
t.seth(0) # 转换画笔的角度为0
t.fd(10)
t.pd()
t.pencolor(255,155,192) # 设置画笔颜色
t.seth(10)
t.begin_fill()
t.circle(5) # 画一个半径为5的圆
t.color(160,82,45) # 设置画笔和填充颜色
t.end_fill()
t.pu()
t.seth(0)
t.fd(20)
t.pd()
t.pencolor(255,155,192)
t.seth(10)
t.begin_fill()
t.circle(5)
t.color(160,82,45)
t.end_fill()
#头
t.color((255,155,192),"pink")
t.pu()
t.seth(90)
t.fd(41)
t.seth(0)
t.fd(0)
t.pd()
t.begin_fill()
t.seth(180)
t.circle(300,-30) # 顺时针画一个半径为300,圆心角为30°的园
t.circle(100,-60)
t.circle(80,-100)
t.circle(150,-20)
t.circle(60,-95)
t.seth(161)
t.circle(-300,15)
t.pu()
t.goto(-100,100)
t.pd()
t.seth(-30)
a=0.4
for i in range(60):
if 0<=i<30 or 60<=i<90:
a=a+0.08
t.lt(3) #向左转3度
t.fd(a) #向前走a的步长
else:
a=a-0.08
t.lt(3)
t.fd(a)
t.end_fill()
#耳朵
t.color((255,155,192),"pink")
t.pu()
t.seth(90)
t.fd(-7)
t.seth(0)
t.fd(70)
t.pd()
t.begin_fill()
t.seth(100)
t.circle(-50,50)
t.circle(-10,120)
t.circle(-50,54)
t.end_fill()
t.pu()
t.seth(90)
t.fd(-12)
t.seth(0)
t.fd(30)
t.pd()
t.begin_fill()
t.seth(100)
t.circle(-50,50)
t.circle(-10,120)
t.circle(-50,56)
t.end_fill()
#眼睛
t.color((255,155,192),"white")
t.pu()
t.seth(90)
t.fd(-20)
t.seth(0)
t.fd(-95)
t.pd()
t.begin_fill()
t.circle(15)
t.end_fill()
t.color("black")
t.pu()
t.seth(90)
t.fd(12)
t.seth(0)
t.fd(-3)
t.pd()
t.begin_fill()
t.circle(3)
t.end_fill()
t.color((255,155,192),"white")
t.pu()
t.seth(90)
t.fd(-25)
t.seth(0)
t.fd(40)
t.pd()
t.begin_fill()
t.circle(15)
t.end_fill()
t.color("black")
t.pu()
t.seth(90)
t.fd(12)
t.seth(0)
t.fd(-3)
t.pd()
t.begin_fill()
t.circle(3)
t.end_fill()
#腮
t.color((255,155,192))
t.pu()
t.seth(90)
t.fd(-95)
t.seth(0)
t.fd(65)
t.pd()
t.begin_fill()
t.circle(30)
t.end_fill()
#嘴
t.color(239,69,19)
t.pu()
t.seth(90)
t.fd(15)
t.seth(0)
t.fd(-100)
t.pd()
t.seth(-80)
t.circle(30,40)
t.circle(40,80)
#身体
t.color("red",(255,99,71))
t.pu()
t.seth(90)
t.fd(-20)
t.seth(0)
t.fd(-78)
t.pd()
t.begin_fill()
t.seth(-130)
t.circle(100,10)
t.circle(300,30)
t.seth(0)
t.fd(230)
t.seth(90)
t.circle(300,30)
t.circle(100,3)
t.color((255,155,192),(255,100,100))
t.seth(-135)
t.circle(-80,63)
t.circle(-150,24)
t.end_fill()
#手
t.color((255,155,192))
t.pu()
t.seth(90)
t.fd(-40)
t.seth(0)
t.fd(-27)
t.pd()
t.seth(-160)
t.circle(300,15)
t.pu()
t.seth(90)
t.fd(15)
t.seth(0)
t.fd(0)
t.pd()
t.seth(-10)
t.circle(-20,90)
t.pu()
t.seth(90)
t.fd(30)
t.seth(0)
t.fd(237)
t.pd()
t.seth(-20)
t.circle(-300,15)
t.pu()
t.seth(90)
t.fd(20)
t.seth(0)
t.fd(0)
t.pd()
t.seth(-170)
t.circle(20,90)
#脚
t.pensize(10)
t.color((240,128,128))
t.pu()
t.seth(90)
t.fd(-75)
t.seth(0)
t.fd(-180)
t.pd()
t.seth(-90)
t.fd(40)
t.seth(-180)
t.color("black")
t.pensize(15)
t.fd(20)
t.pensize(10)
t.color((240,128,128))
t.pu()
t.seth(90)
t.fd(40)
t.seth(0)
t.fd(90)
t.pd()
t.seth(-90)
t.fd(40)
t.seth(-180)
t.color("black")
t.pensize(15)
t.fd(20)
#尾巴
t.pensize(4)
t.color((255,155,192))
t.pu()
t.seth(90)
t.fd(70)
t.seth(0)
t.fd(95)
t.pd()
t.seth(0)
t.circle(70,20)
t.circle(10,330)
t.circle(70,30)
# ## 列表的复制
# - copy 浅复制
# - deepcopy import copy 深复制
# - http://www.pythontutor.com/visualize.html#mode=edit
# ## 列表排序
# - sort
# - sorted
# - 列表的多级排序
# - 匿名函数
# ## EP:
# - 手动排序该列表[5,3,8,0,17],以升序或者降序
# - 1
# 
scores = []
for i in range(4):
s = eval(input('>>'))
scores.append(s)
mnum = max(scores)
for i in scores:
if i >= mnum-10:
print('A')
elif i>=mnum-20:
print('B')
elif i >= mnum-30:
print('C')
elif i>=mnum-40:
print('D')
else:
print('F')
# - 2
# 
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
numbers.reverse()
print(numbers)
# - 3
# 
i = 1
nums = {}
while i<101:
nums[i] = 0
i+=1
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
for i in numbers:
nums[i] += 1
for i in range(1,101):
if nums[i]!=0:
print(i,nums[i])
# - 4
# 
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
mean = sum(numbers) / len(numbers)
bn = 0
ln = 0
for i in numbers:
if i>=mean:
bn+=1
else:
ln+=1
print('大于等于平均',bn)
print('小于平均',ln)
# - 5
# 
import random
randn = []
for i in range(1000):
n = random.randint(0,9)
randn.append(n)
print('0:',randn.count(0),'1:',randn.count(1),'2:',randn.count(2),'3:',randn.count(3),'4:',randn.count(4),'5:',randn.count
(5),'6:',randn.count(6),'7:',randn.count(7),'8:',randn.count(8),'9:',randn.count(9))
# - 6
# 
def indexOfSmallestElement(lst):
small = min(lst)
cs = lst.count(small)
if cs>1:
return lst.index(small)
a = []
a = input('>>').split(' ')
numbers = list(map(int,a))
indexOfSmallestElement(numbers)
# - 7
# 
# 
import random
def shuffle(lst):
b = []
while len(lst)!=0:
n = random.randint(0,len(lst)-1)
c = numbers.pop(n)
b.append(c)
print(b)
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
shuffle(numbers)
# - 8
# 
def eliminateDuplicates(lst):
return list(set(lst))
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
eliminateDuplicates(numbers)
# - 9
# 
def isSorted(lst):
n = 0
for i in range(len(lst)-1):
if lst[i+1]>=lst[i]:
n+=1
if n== len(lst)-1:
print('yes')
else:
print('no')
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
isSorted(numbers)
# - 10
# 
def bubbleSort(nums):
for i in range(len(nums)-1):
for j in range(len(nums)-i-1):
if nums[j] > nums[j+1]:
nums[j], nums[j+1] = nums[j+1], nums[j]
return nums
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
bubbleSort(numbers)
# - 11
# 
# - 12
# 
def isConsecutiveFour(values):
if len(values)>=4:
for i in range(len(values)-3):
if values[i]==values[i+1]==values[i+2]==values[i+3]:
print('Yes')
break
else:
print('No')
else:
print('No')
a = []
a = input('>>').split(' ')
numbers = list(map(int, a))
isConsecutiveFour(numbers)
|
9.20.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib notebook
# %reload_ext autoreload
# %autoreload 2
# %load_ext cython
import csv
import sys, os, ctypes
import math
import numpy as np
import matplotlib.pyplot as plt
from collections import defaultdict
from numpy import genfromtxt
import itertools
from ipywidgets import IntProgress
from IPython.display import display, clear_output
# +
# Turn a sequence of numbers into a debug friendly format
def stringify(b,spacer='', max=None):
rtn = spacer.join(map(lambda x: '_' if x is None else str(int(x)), b))
if max is not None and len(rtn) > max:
return rtn[:max] + "..."
return rtn
# Differential encodings cares about transitions not symbols; so
# we can take a modulated signal and just return how many symbols
# are in a row.
def group_like(l):
return [len(list(c)) for s, c in itertools.groupby(l)]
# -
group_like([1, 1, 1, 1, 0, 0])
# + language="cython"
# import numpy as np
# cimport numpy as np
# import cython
#
# # We will use levenshtein edit distance as our error metric; but it's
# # pretty slow in native python so we'll do it in cython and it's roughly
# # as fast as a pure C implementation
# @cython.boundscheck(False) # Deactivate bounds checking
# @cython.wraparound(False) # Deactivate negative indexing.
# def levenshtein(np.ndarray s, np.ndarray t):
# ''' From Wikipedia article; Iterative with two matrix rows. '''
# cdef int is_eq = True
# cdef int len_s = len(s)
# cdef int len_t = len(t)
# cdef int i = 0
# cdef int j = 0
# cdef int cost = 0
# cdef long[:] _s = s;
# cdef long[:] _t = t;
# if len_s == len_t:
# for i in range(len_s):
# if _s[i] != _t[i]:
# is_eq = False
# break
# if is_eq:
# return 0
# elif len_s == 0: return len_t
# elif len_t == 0: return len_s
# _v0 = np.zeros((len_t + 1), dtype=np.int32)
# cdef int len_v0 = len(_v0)
# cdef int[:] v0 = _v0
# _v1 = np.zeros((len_t + 1), dtype=np.int32)
# cdef int[:] v1 = _v1
# for i in range(len(v0)):
# v0[i] = i
# for i in range(len_s):
# v1[0] = i + 1
# for j in range(len_t):
# cost = 0 if _s[i] == _t[j] else 1
# v1[j + 1] = min(v1[j] + 1, v0[j + 1] + 1, v0[j] + cost)
# for j in range(len_v0):
# v0[j] = v1[j]
#
# return v1[len_t]
# levenshtein(np.array([1,0]), np.array([0,1]))
# +
def conditional_reseed(seed):
if seed is not None:
np.random.seed(seed)
def make_data_stream(n, seed = None):
conditional_reseed(seed)
return np.random.randint(0, 2, size=n)
# -
make_data_stream(50)
# Encode a list of bits as per https://en.wikipedia.org/wiki/Differential_Manchester_encoding
def encode_diff_manchester(d, start = None, inv = 0):
# start state doesn't matter; set it randomly to avoid relying on it below
state = np.random.randint(0, 2) if start is None else start
rtn = []
for bit in d:
if bit ^ inv:
rtn += [state, state]
state = state ^ 1
else:
rtn += [state, state ^ 1]
return rtn
# +
ds = make_data_stream(25)
eds = encode_diff_manchester(ds)
print('This should stack transitions under every |, between the bars is the bit that generated the two bits below')
print(' ' + stringify(ds, ' | '))
print(stringify(eds, ' '))
# -
# Take in a data stream and mimic sampling it at some supersampling factor `f`,
# with random phase and frequency error.
def make_sampled_stream(d, f, seed = None, f_error = .05, drift = .05):
conditional_reseed(seed)
inc = 1. / (f * np.random.uniform(1 - f_error, 1 + f_error))
phase = np.random.rand() / 2.
rtn = []
# Faster to generate all these at once rather than in the loop
phase_drifts = np.random.uniform(-drift/f, drift/f, size=math.ceil((1+len(d))*f))
idx = 0
while phase < len(d):
this_floor = math.floor(phase)
rtn.append(d[this_floor])
phase += inc + phase_drifts[idx % len(phase_drifts)]
idx+=1
return rtn
f = 3
sampled_eds = make_sampled_stream(eds, f,seed=11)
print("With f=3, the sampled stream should ideally have three symbols perfectly for each modulated bit but random error has been added")
print('Data | ' + stringify(ds, ' | '))
print('Modulated: | ' + stringify(eds, ' | '))
print('Sampled: |' + stringify(sampled_eds, ' '))
stringify(group_like(sampled_eds),' ')
def decode_sampled_diff_manchester(sampled, f):
groups = group_like(sampled)
read_since_output = 0
grps_since_output = 0
rtn = []
for g in groups:
read_since_output += g
grps_since_output += 1
if read_since_output >= math.ceil(1.5 * f):
if grps_since_output == 2:
rtn.append(0)
elif grps_since_output == 1:
rtn.append(1)
# This shouldn't happen except for error;
# most likely guess is a '0'.
elif grps_since_output == 3:
rtn.append(0)
# We also don't handle the case where 'read_since_output'
# is too high -- if it's much more than 2 * f, that is
# likely an encoding error. This is sort of self correcting
# if we are out of phase; since we handle both errors by
# effectively dropping data it'll resync.
# For actual usage; probably throw in some indication of
# where the decodeing is valid / invalid. Later we'll use
# edit distance which will sort this out for us.
read_since_output = grps_since_output = 0
return np.array(rtn)
# +
# Generate `cnt` data streams of `size` length, encode, sample with noise,
# demodulate and calculate the average error introduced into the decoded
# stream.
def evaluate_f_error(f, size=500, seed=None, cnt=1000):
err = 0
for i in range(cnt):
ds = make_data_stream(size)
eds = encode_diff_manchester(ds)
sampled_eds = make_sampled_stream(eds, f, f_error = .05, drift = .05)
dsdf = np.array(decode_sampled_diff_manchester(sampled_eds, f))
err += levenshtein(ds, dsdf)
return err / cnt
def evaluate_fs(fs):
progressBar = IntProgress(min=0, max=len(fs))
display(progressBar)
rtn = []
for f in fs:
progressBar.value += 1
rtn.append(evaluate_f_error(f))
print("Done!")
return np.array(rtn)
# -
# This takes a minute; go get a coffee or something
fs = np.arange(1., 5.25, .05)
# %time fs_errors = evaluate_fs(fs)
plt.figure()
plt.plot(fs, fs_errors, label="Avg edit distance")
perfect_fs = np.copy(fs)
perfect_fs[fs_errors > 0] = None
plt.plot(perfect_fs, fs_errors, '-')
plt.xlabel('sample rate / data rate')
plt.ylabel('Error')
plt.legend()
|
jupyter/encoding-test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="jjneMzGxDhGd" colab={}
# Install MTCNN library for face detection
# !pip install mtcnn
# + colab_type="code" id="Po6pXsPbTTyu" colab={}
# Import required packages
import os
import numpy as np
import cv2
from PIL import Image
from numpy import asarray
from mtcnn.mtcnn import MTCNN
from keras.models import load_model,model_from_json
# + colab_type="code" id="2EFsZHawnnZ5" colab={}
# Load pretrained Inception-ResNet-v1 model
# Update model and weights path according to your working environment
model_path = "Models/Inception_ResNet_v1.json"
weights_path = "Models/facenet_keras_weights.h5"
json_file = open(model_path, 'r')
loaded_model_json = json_file.read()
json_file.close()
enc_model = model_from_json(loaded_model_json)
enc_model.load_weights(weights_path)
# + colab_type="code" id="LINQ1ZzCbEYw" colab={}
# Initialize a MTCNN face detector
mtcnn_detector = MTCNN()
# + colab_type="code" id="060X7dGbUaJJ" colab={}
# Function to detect and extract face from a image
def detect_face(filename, required_size=(160, 160),normalize = True):
img = Image.open(filename)
# convert to RGB
img = img.convert('RGB')
# convert to array
pixels = np.asarray(img)
# detect faces in the image
results = mtcnn_detector.detect_faces(pixels)
# extract the bounding box from the first face
x1, y1, width, height = results[0]['box']
x1, y1 = abs(x1), abs(y1)
x2, y2 = x1 + width, y1 + height
# extract the face
face = pixels[y1:y2, x1:x2]
# resize pixels to the model size
image = Image.fromarray(face)
image = image.resize(required_size)
face_array = asarray(image)
if normalize == True:
mean = np.mean(face_array, axis=(0,1,2), keepdims=True)
std = np.std(face_array, axis=(0,1,2), keepdims=True)
std_adj = np.maximum(std, 1.0)
return (face_array - mean) / std
else :
return face_array
# + colab_type="code" id="bMzpMGNe4HgU" colab={}
# Compute Face encodings and load IDs of known persons
# Update face database path according to your working environment
known_faces_encodings = []
known_faces_ids = []
known_faces_path = "Face_database/"
for filename in os.listdir(known_faces_path):
# Detect faces
face = detect_face(known_faces_path+filename,normalize = True)
# Compute face encodings
feature_vector = enc_model.predict(face.reshape(1,160,160,3))
feature_vector/= np.sqrt(np.sum(feature_vector**2))
known_faces_encodings.append(feature_vector)
# Save Person IDs
label = filename.split('.')[0]
known_faces_ids.append(label)
known_faces_encodings = np.array(known_faces_encodings).reshape(len(known_faces_encodings),128)
known_faces_ids = np.array(known_faces_ids)
# + colab_type="code" id="4S60cggIBBR2" colab={}
# No. of known IDs loaded from database
print(known_faces_ids.shape[0])
# + colab_type="code" id="iyV6Eyd3XkA8" colab={}
# Function to recognize a face (if it is in known_faces)
def recognize(img,known_faces_encodings,known_faces_ids,threshold = 0.75):
scores = np.zeros((len(known_faces_ids),1),dtype=float)
enc = enc_model.predict(img.reshape(1,160,160,3))
enc/= np.sqrt(np.sum(enc**2))
scores = np.sqrt(np.sum((enc-known_faces_encodings)**2,axis=1))
match = np.argmin(scores)
if scores[match] > threshold :
return ("UNKNOWN",0)
else :
return (known_faces_ids[match],scores[match])
# + colab_type="code" id="1okFKhLStsHD" colab={}
# Function to perform real-time face recognition through a webcam
def face_recognition(mode,file_path,known_faces_encodings,known_faces_ids,
detector = 'haar', threshold = 0.75):
if detector == 'haar':
# Load the cascade
face_cascade = cv2.CascadeClassifier('Models/haarcascade_frontalface_default.xml')
if mode == 'webcam':
# To capture webcam feed. Change argument for differnt webcams
cap = cv2.VideoCapture(0)
elif mode == 'video':
# To capture video feed
cap = cv2.VideoCapture(file_path)
while True:
# Read the frame
_, img = cap.read()
# Stop if end of video file
if _ == False:
break;
if detector == 'haar':
#Convert to grayscale
gray = cv2.cvtColor(img, cv2.COLOR_BGR2GRAY)
# Detect the faces
faces = face_cascade.detectMultiScale(gray, 1.1, 4)
elif detector == 'mtcnn' :
results = mtcnn_detector.detect_faces(img)
if(len(results)==0):
continue
faces = []
for i in range(len(results)):
x,y,w,h = results[i]['box']
x, y = abs(x), abs(y)
faces.append([x,y,w,h])
# Draw the rectangle around each face
for (x, y, w, h) in faces:
image = Image.fromarray(img[y:y+h, x:x+w])
image = image.resize((160,160))
face_array = asarray(image)
# Normalize
mean = np.mean(face_array, axis=(0,1,2), keepdims=True)
std = np.std(face_array, axis=(0,1,2), keepdims=True)
std_adj = np.maximum(std, 1.0)
face_array_normalized = (face_array - mean) / std
# Recognize
label = recognize(face_array_normalized,known_faces_encodings,known_faces_ids,threshold = 0.75)
cv2.rectangle(img, (x, y), (x+w, y+h), (0, 0, 0), 2)
cv2.putText(img, label[0], (x, y-10), cv2.FONT_HERSHEY_SIMPLEX, 0.9, (255,255,255), 2)
# Display
cv2.imshow('Face_Recognition', img)
# Stop if escape key is pressed
key = cv2.waitKey(25) & 0xff
if key==27:
break
# Release the VideoCapture object
cap.release()
# + colab_type="code" id="zoVzEG5Qz4zF" colab={}
# Execute Face recognition on a webcam feed.
# Note : Threshold has to be adjusted according to your requirements !
face_recognition('webcam',None,known_faces_encodings,known_faces_ids,
detector = 'haar',threshold = 0.75)
# + colab_type="code" id="yeRZlEJXVJZh" colab={}
# Execute Face recognition on a Video file.
# Note : Threshold has to be adjusted according to your requirements !
face_recognition('video',"test.mp4",known_faces_encodings,known_faces_ids,
detector = 'mtcnn',threshold = 0.75)
|
Face_Recognition.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# # When To Stop Fuzzing
#
# In the past chapters, we have discussed several fuzzing techniques. Knowing _what_ to do is important, but it is also important to know when to _stop_ doing things. In this chapter, we will learn when to _stop fuzzing_ – and use a prominent example for this purpose: The *Enigma* machine that was used in the second world war by the navy of Nazi Germany to encrypt communications, and how Alan Turing and I.<NAME> used _fuzzing techniques_ to crack ciphers for the Naval Enigma machine.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# Turing did not only develop the foundations of computer science, the Turing machine. Together with his assistant <NAME>, he also invented estimators of the probability of an event occuring that has never previously occured. We show how the Good-Turing estimator can be used to quantify the *residual risk* of a fuzzing campaign that finds no vulnerabilities. Meaning, we show how it estimates the probability of discovering a vulnerability when no vulnerability has been observed before throughout the fuzzing campaign.
#
# We discuss means to speed up [coverage-based fuzzers](Coverage.ipynb) and introduce a range of estimation and extrapolation methodologies to assess and extrapolate fuzzing progress and residual risk.
#
# **Prerequisites**
#
# * _The chapter on [Coverage](Coverage.ipynb) discusses how to use coverage information for an executed test input to guide a coverage-based mutational greybox fuzzer_.
# * Some knowledge of statistics is helpful.
# + slideshow={"slide_type": "skip"}
import fuzzingbook_utils
# + slideshow={"slide_type": "skip"}
import Fuzzer
import Coverage
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## The Enigma Machine
#
# It is autumn in the year of 1938. Turing has just finished his PhD at Princeton University demonstrating the limits of computation and laying the foundation for the theory of computer science. Nazi Germany is rearming. It has reoccupied the Rhineland and annexed Austria against the treaty of Versailles. It has just annexed the Sudetenland in Czechoslovakia and begins preparations to take over the rest of Czechoslovakia despite an agreement just signed in Munich.
#
# Meanwhile, the British intelligence is building up their capability to break encrypted messages used by the Germans to communicate military and naval information. The Germans are using [Enigma machines](https://en.wikipedia.org/wiki/Enigma_machine) for encryption. Enigma machines use a series of electro-mechanical rotor cipher machines to protect military communication. Here is a picture of an Enigma machine:
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# 
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# By the time Turing joined the British Bletchley park, the Polish intelligence reverse engineered the logical structure of the Enigma machine and built a decryption machine called *Bomba* (perhaps because of the ticking noise they made). A bomba simulates six Enigma machines simultaneously and tries different decryption keys until the code is broken. The Polish bomba might have been the very _first fuzzer_.
#
# Turing took it upon himself to crack ciphers of the Naval Enigma machine, which were notoriously hard to crack. The Naval Enigma used, as part of its encryption key, a three letter sequence called *trigram*. These trigrams were selected from a book, called *Kenngruppenbuch*, which contained all trigrams in a random order.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ### The Kenngruppenbuch
#
# Let's start with the Kenngruppenbuch (K-Book).
#
# We are going to use the following Python functions.
# * `shuffle(elements)` - shuffle *elements* and put items in random order.
# * `choice(elements, p=weights)` - choose an item from *elements* at random. An element with twice the *weight* is twice as likely to be chosen.
# * `log(a)` - returns the natural logarithm of a.
# * `a ** b` - is the a to the power of b (a.k.a. [power operator](https://docs.python.org/3/reference/expressions.html#the-power-operator))
# + slideshow={"slide_type": "skip"}
import string
# + slideshow={"slide_type": "skip"}
import numpy
from numpy.random import choice
from numpy.random import shuffle
from numpy import log
# + [markdown] slideshow={"slide_type": "subslide"}
# We start with creating the set of trigrams:
# + slideshow={"slide_type": "fragment"}
letters = list(string.ascii_letters[26:]) # upper-case characters
trigrams = [str(a + b + c) for a in letters for b in letters for c in letters]
shuffle(trigrams)
# + slideshow={"slide_type": "fragment"}
trigrams[:10]
# + [markdown] slideshow={"slide_type": "fragment"}
# These now go into the Kenngruppenbuch. However, it was observed that some trigrams were more likely chosen than others. For instance, trigrams at the top-left corner of any page, or trigrams on the first or last few pages were more likely than one somewhere in the middle of the book or page. We reflect this difference in distribution by assigning a _probability_ to each trigram, using Benford's law as introduced in [Probabilistic Fuzzing](ProbabilisticGrammarFuzzer.ipynb).
# + [markdown] slideshow={"slide_type": "subslide"}
# Recall, that Benford's law assigns the $i$-th digit the probability $\log_{10}\left(1 + \frac{1}{i}\right)$ where the base 10 is chosen because there are 10 digits $i\in [0,9]$. However, Benford's law works for an arbitrary number of "digits". Hence, we assign the $i$-th trigram the probability $\log_b\left(1 + \frac{1}{i}\right)$ where the base $b$ is the number of all possible trigrams $b=26^3$.
# + slideshow={"slide_type": "fragment"}
k_book = {} # Kenngruppenbuch
for i in range(1, len(trigrams) + 1):
trigram = trigrams[i - 1]
# choose weights according to Benford's law
k_book[trigram] = log(1 + 1 / i) / log(26**3 + 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's a random trigram from the Kenngruppenbuch:
# + slideshow={"slide_type": "subslide"}
random_trigram = choice(list(k_book.keys()), p=list(k_book.values()))
random_trigram
# + [markdown] slideshow={"slide_type": "fragment"}
# And this is its probability:
# + slideshow={"slide_type": "fragment"}
k_book[random_trigram]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fuzzing the Enigma
#
# In the following, we introduce an extremely simplified implementation of the Naval Enigma based on the trigrams from the K-book. Of course, the encryption mechanism of the actual Enigma machine is much more sophisticated and worthy of a much more detailed investigation. We encourage the interested reader to follow up with further reading listed in the Background section.
#
# The personell at Bletchley Park can only check whether an encoded message is encoded with a (guessed) trigram.
# Our implementation `naval_enigma()` takes a `message` and a `key` (i.e., the guessed trigram). If the given key matches the (previously computed) key for the message, `naval_enigma()` returns `True`.
# + slideshow={"slide_type": "skip"}
from Fuzzer import RandomFuzzer
from Fuzzer import Runner
# + slideshow={"slide_type": "subslide"}
class EnigmaMachine(Runner):
def __init__(self, k_book):
self.k_book = k_book
self.reset()
def reset(self):
"""Resets the key register"""
self.msg2key = {}
def internal_msg2key(self, message):
"""Internal helper method.
Returns the trigram for an encoded message."""
if not message in self.msg2key:
# Simulating how an officer chooses a key from the Kenngruppenbuch to encode the message.
self.msg2key[message] = choice(list(self.k_book.keys()), p=list(self.k_book.values()))
trigram = self.msg2key[message]
return trigram
def naval_enigma(self, message, key):
"""Returns true if 'message' is encoded with 'key'"""
if key == self.internal_msg2key(message):
return True
else:
return False
# + [markdown] slideshow={"slide_type": "subslide"}
# To "fuzz" the `naval_enigma()`, our job will be to come up with a key that matches a given (encrypted) message. Since the keys only have three characters, we have a good chance to achieve this in much less than a seconds. (Of course, longer keys will be much harder to find via random fuzzing.)
# + slideshow={"slide_type": "fragment"}
class EnigmaMachine(EnigmaMachine):
def run(self, tri):
"""PASS if cur_msg is encoded with trigram tri"""
if self.naval_enigma(self.cur_msg, tri):
outcome = self.PASS
else:
outcome = self.FAIL
return (tri, outcome)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now we can use the `EnigmaMachine` to check whether a certain message is encoded with a certain trigram.
# + slideshow={"slide_type": "fragment"}
enigma = EnigmaMachine(k_book)
enigma.cur_msg = "BrEaK mE. L0Lzz"
enigma.run("AAA")
# + [markdown] slideshow={"slide_type": "fragment"}
# The simplest way to crack an encoded message is by brute forcing. Suppose, at Bletchley park they would try random trigrams until a message is broken.
# + slideshow={"slide_type": "subslide"}
class BletchleyPark(object):
def __init__(self, enigma):
self.enigma = enigma
self.enigma.reset()
self.enigma_fuzzer = RandomFuzzer(
min_length=3,
max_length=3,
char_start=65,
char_range=26)
def break_message(self, message):
"""Returning the trigram for an encoded message"""
self.enigma.cur_msg = message
while True:
(trigram, outcome) = self.enigma_fuzzer.run(self.enigma)
if outcome == self.enigma.PASS:
break
return trigram
# + [markdown] slideshow={"slide_type": "subslide"}
# How long does it take Bletchley park to find the key using this brute forcing approach?
# + slideshow={"slide_type": "skip"}
from Timer import Timer
# + slideshow={"slide_type": "fragment"}
enigma = EnigmaMachine(k_book)
bletchley = BletchleyPark(enigma)
with Timer() as t:
trigram = bletchley.break_message("BrEaK mE. L0Lzz")
# + [markdown] slideshow={"slide_type": "fragment"}
# Here's the key for the current message:
# + slideshow={"slide_type": "fragment"}
trigram
# + [markdown] slideshow={"slide_type": "fragment"}
# And no, this did not take long:
# + slideshow={"slide_type": "fragment"}
'%f seconds' % t.elapsed_time()
# + slideshow={"slide_type": "fragment"}
'Bletchley cracks about %d messages per second' % (1/t.elapsed_time())
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Turing's Observations
# Okay, lets crack a few messages and count the number of times each trigram is observed.
# + slideshow={"slide_type": "skip"}
from collections import defaultdict
# + slideshow={"slide_type": "fragment"}
n = 100 # messages to crack
# + slideshow={"slide_type": "fragment"}
observed = defaultdict(int)
for msg in range(0, n):
trigram = bletchley.break_message(msg)
observed[trigram] += 1
# list of trigrams that have been observed
counts = [k for k, v in observed.items() if int(v) > 0]
t_trigrams = len(k_book)
o_trigrams = len(counts)
# + slideshow={"slide_type": "subslide"}
"After cracking %d messages, we observed %d out of %d trigrams." % (
n, o_trigrams, t_trigrams)
# + slideshow={"slide_type": "fragment"}
singletons = len([k for k, v in observed.items() if int(v) == 1])
# + slideshow={"slide_type": "fragment"}
"From the %d observed trigrams, %d were observed only once." % (
o_trigrams, singletons)
# + [markdown] slideshow={"slide_type": "fragment"}
# Given a sample of previously used entries, Turing wanted to _estimate the likelihood_ that the current unknown entry was one that had been previously used, and further, to estimate the probability distribution over the previously used entries. This lead to the development of the estimators of the missing mass and estimates of the true probability mass of the set of items occuring in the sample. Good worked with Turing during the war and, with Turing’s permission, published the analysis of the bias of these estimators in 1953.
# + [markdown] slideshow={"slide_type": "subslide"}
# Suppose, after finding the keys for n=100 messages, we have observed the trigram "ABC" exactly $X_\text{ABC}=10$ times. What is the probability $p_\text{ABC}$ that "ABC" is the key for the next message? Empirically, we would estimate $\hat p_\text{ABC}=\frac{X_\text{ABC}}{n}=0.1$. We can derive the empirical estimates for all other trigrams that we have observed. However, it becomes quickly evident that the complete probability mass is distributed over the *observed* trigrams. This leaves no mass for *unobserved* trigrams, i.e., the probability of discovering a new trigram. This is called the missing probability mass or the discovery probability.
# + [markdown] slideshow={"slide_type": "subslide"}
# Turing and Good derived an estimate of the *discovery probability* $p_0$, i.e., the probability to discover an unobserved trigram, as the number $f_1$ of trigrams observed exactly once divided by the total number $n$ of messages cracked:
# $$
# p_0 = \frac{f_1}{n}
# $$
# where $f_1$ is the number of singletons and $n$ is the number of cracked messages.
# + [markdown] slideshow={"slide_type": "fragment"}
# Lets explore this idea for a bit. We'll extend `BletchleyPark` to crack `n` messages and record the number of trigrams observed as the number of cracked messages increases.
# + slideshow={"slide_type": "subslide"}
class BletchleyPark(BletchleyPark):
def break_message(self, message):
"""Returning the trigram for an encoded message"""
# For the following experiment, we want to make it practical
# to break a large number of messages. So, we remove the
# loop and just return the trigram for a message.
#
# enigma.cur_msg = message
# while True:
# (trigram, outcome) = self.enigma_fuzzer.run(self.enigma)
# if outcome == self.enigma.PASS:
# break
trigram = enigma.internal_msg2key(message)
return trigram
def break_n_messages(self, n):
"""Returns how often each trigram has been observed,
and #trigrams discovered for each message."""
observed = defaultdict(int)
timeseries = [0] * n
# Crack n messages and record #trigrams observed as #messages increases
cur_observed = 0
for cur_msg in range(0, n):
trigram = self.break_message(cur_msg)
observed[trigram] += 1
if (observed[trigram] == 1):
cur_observed += 1
timeseries[cur_msg] = cur_observed
return (observed, timeseries)
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's crack 2000 messages and compute the GT-estimate.
# + slideshow={"slide_type": "fragment"}
n = 2000 # messages to crack
# + slideshow={"slide_type": "fragment"}
bletchley = BletchleyPark(enigma)
(observed, timeseries) = bletchley.break_n_messages(n)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let us determine the Good-Turing estimate of the probability that the next trigram has not been observed before:
# + slideshow={"slide_type": "fragment"}
singletons = len([k for k, v in observed.items() if int(v) == 1])
gt = singletons / n
gt
# + [markdown] slideshow={"slide_type": "subslide"}
# We can verify the Good-Turing estimate empirically and compute the empirically determined probability that the next trigram has not been observed before. To do this, we repeat the following experiment repeats=1000 times, reporting the average: If the next message is a new trigram, return 1, otherwise return 0. Note that here, we do not record the newly discovered trigrams as observed.
# + slideshow={"slide_type": "fragment"}
repeats = 1000 # experiment repetitions
# + slideshow={"slide_type": "fragment"}
newly_discovered = 0
for cur_msg in range(n, n + repeats):
trigram = bletchley.break_message(cur_msg)
if(observed[trigram] == 0):
newly_discovered += 1
newly_discovered / repeats
# + [markdown] slideshow={"slide_type": "subslide"}
# Looks pretty accurate, huh? The difference between estimates is reasonably small, probably below 0.03. However, the Good-Turing estimate did not nearly require as much computational resources as the empirical estimate. Unlike the empirical estimate, the Good-Turing estimate can be computed during the campaign. Unlike the empirical estimate, the Good-Turing estimate requires no additional, redundant repetitions.
# + [markdown] slideshow={"slide_type": "subslide"}
# In fact, the Good-Turing (GT) estimator often performs close to the best estimator for arbitrary distributions ([Try it here!](#Kenngruppenbuch)). Of course, the concept of *discovery* is not limited to trigrams. The GT estimator is also used in the study of natural languages to estimate the likelihood that we haven't ever heard or read the word we next encounter. The GT estimator is used in ecology to estimate the likelihood of discovering a new, unseen species in our quest to catalog all _species_ on earth. Later, we will see how it can be used to estimate the probability to discover a vulnerability when none has been observed, yet (i.e., residual risk).
# + [markdown] slideshow={"slide_type": "subslide"}
# Alan Turing was interested in the _complement_ $(1-GT)$ which gives the proportion of _all_ messages for which the Brits have already observed the trigram needed for decryption. For this reason, the complement is also called sample coverage. The *sample coverage* quantifies how much we know about decryption of all messages given the few messages we have already decrypted.
# + [markdown] slideshow={"slide_type": "fragment"}
# The probability that the next message can be decrypted with a previously discovered trigram is:
# + slideshow={"slide_type": "fragment"}
1 - gt
# + [markdown] slideshow={"slide_type": "subslide"}
# The *inverse* of the GT-estimate (1/GT) is a _maximum likelihood estimate_ of the expected number of messages that we can decrypt with previously observed trigrams before having to find a new trigram to decrypt the message. In our setting, the number of messages for which we can expect to reuse previous trigrams before having to discover a new trigram is:
# + slideshow={"slide_type": "fragment"}
1 / gt
# + [markdown] slideshow={"slide_type": "subslide"}
# But why is GT so accurate? Intuitively, despite a large sampling effort (i.e., cracking $n$ messages), there are still $f_1$ trigrams that have been observed only once. We could say that such "singletons" are very rare trigrams. Hence, the probability that the next messages is encoded with such a rare but observed trigram gives a good upper bound on the probability that the next message is encoded with an evidently much rarer, unobserved trigram. Since Turing's observation 80 years ago, an entire statistical theory has been developed around the hypothesis that rare, observed "species" are good predictors of unobserved species.
#
# Let's have a look at the distribution of rare trigrams.
# + slideshow={"slide_type": "fragment"}
# %matplotlib inline
# + slideshow={"slide_type": "skip"}
import matplotlib.pyplot as plt
# + slideshow={"slide_type": "subslide"}
frequencies = [v for k, v in observed.items() if int(v) > 0]
frequencies.sort(reverse=True)
# Uncomment to see how often each discovered trigram has been observed
# print(frequencies)
# frequency of rare trigrams
plt.figure(num=None, figsize=(12, 4), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(1, 2, 1)
plt.hist(frequencies, range=[1, 21], bins=numpy.arange(1, 21) - 0.5)
plt.xticks(range(1, 21))
plt.xlabel('# of occurances (e.g., 1 represents singleton trigrams)')
plt.ylabel('Frequency of occurances')
plt.title('Figure 1. Frequency of Rare Trigrams')
# trigram discovery over time
plt.subplot(1, 2, 2)
plt.plot(timeseries)
plt.xlabel('# of messages cracked')
plt.ylabel('# of trigrams discovered')
plt.title('Figure 2. Trigram Discovery Over Time');
# + slideshow={"slide_type": "subslide"}
# Statistics for most and least often observed trigrams
singletons = len([v for k, v in observed.items() if int(v) == 1])
total = len(frequencies)
print("%3d of %3d trigrams (%.3f%%) have been observed 1 time (i.e., are singleton trigrams)."
% (singletons, total, singletons * 100 / total))
print("%3d of %3d trigrams ( %.3f%%) have been observed %d times."
% (1, total, 1 / total, frequencies[0]))
# + [markdown] slideshow={"slide_type": "subslide"}
# The *majority of trigrams* have been observed only once, as we can see in Figure 1 (left). In other words, a the majority of observed trigrams are "rare" singletons. In Figure 2 (right), we can see that discovery is in full swing. The trajectory seems almost linear. However, since there is a finite number of trigrams (26^3 = 17,576) trigram discovery will slow down and eventually approach an asymptote (the total number of trigrams).
#
# ### Boosting the Performance of BletchleyPark
# Some trigrams have been observed very often. We call these "abundant" trigrams.
# + slideshow={"slide_type": "subslide"}
print("Trigram : Frequency")
for trigram in sorted(observed, key=observed.get, reverse=True):
if observed[trigram] > 10:
print(" %s : %d" % (trigram, observed[trigram]))
# + [markdown] slideshow={"slide_type": "subslide"}
# We'll speed up the code breaking by _trying the abundant trigrams first_.
#
# First, we'll find out how many messages can be cracked by the existing brute forcing strategy at Bledgley park, given a maximum number of attempts. We'll also track the number of messages cracked over time (`timeseries`).
# + slideshow={"slide_type": "subslide"}
class BletchleyPark(BletchleyPark):
def __init__(self, enigma):
super().__init__(enigma)
self.cur_attempts = 0
self.cur_observed = 0
self.observed = defaultdict(int)
self.timeseries = [None] * max_attempts * 2
def break_message(self, message):
"""Returns the trigram for an encoded message, and
track #trigrams observed as #attempts increases."""
self.enigma.cur_msg = message
while True:
self.cur_attempts += 1 # NEW
(trigram, outcome) = self.enigma_fuzzer.run(self.enigma)
self.timeseries[self.cur_attempts] = self.cur_observed # NEW
if outcome == self.enigma.PASS:
break
return trigram
def break_max_attempts(self, max_attempts):
"""Returns #messages successfully cracked after a given #attempts."""
cur_msg = 0
n_messages = 0
while True:
trigram = self.break_message(cur_msg)
# stop when reaching max_attempts
if self.cur_attempts >= max_attempts:
break
# update observed trigrams
n_messages += 1
self.observed[trigram] += 1
if (self.observed[trigram] == 1):
self.cur_observed += 1
self.timeseries[self.cur_attempts] = self.cur_observed
cur_msg += 1
return n_messages
# + [markdown] slideshow={"slide_type": "subslide"}
# `original` is the number of messages cracked by the bruteforcing strategy, given 100k attempts. Can we beat this?
# + slideshow={"slide_type": "fragment"}
max_attempts = 100000
# + slideshow={"slide_type": "fragment"}
bletchley = BletchleyPark(enigma)
original = bletchley.break_max_attempts(max_attempts)
original
# + [markdown] slideshow={"slide_type": "fragment"}
# Now, we'll create a boosting strategy by trying trigrams first that we have previously observed most often.
# + slideshow={"slide_type": "subslide"}
class BoostedBletchleyPark(BletchleyPark):
def break_message(self, message):
"""Returns the trigram for an encoded message, and
track #trigrams observed as #attempts increases."""
self.enigma.cur_msg = message
# boost cracking by trying observed trigrams first
for trigram in sorted(self.prior, key=self.prior.get, reverse=True):
self.cur_attempts += 1
(_, outcome) = self.enigma.run(trigram)
self.timeseries[self.cur_attempts] = self.cur_observed
if outcome == self.enigma.PASS:
return trigram
# else fall back to normal cracking
return super().break_message(message)
# + [markdown] slideshow={"slide_type": "subslide"}
# `boosted` is the number of messages cracked by the boosted strategy.
# + slideshow={"slide_type": "fragment"}
boostedBletchley = BoostedBletchleyPark(enigma)
boostedBletchley.prior = observed
boosted = boostedBletchley.break_max_attempts(max_attempts)
boosted
# + [markdown] slideshow={"slide_type": "subslide"}
# We see that the boosted technique cracks substantially more messages. It is worthwhile to record how often each trigram is being used as key and try them in the order of their occurence.
#
# ***Try it***. *For practical reasons, we use a large number of previous observations as prior (`boostedBletchley.prior = observed`). You can try to change the code such that the strategy uses the trigram frequencies (`self.observed`) observed **during** the campaign itself to boost the campaign. You will need to increase `max_attempts` and wait for a long while.*
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's compare the number of trigrams discovered over time.
# + slideshow={"slide_type": "subslide"}
# print plots
line_old, = plt.plot(bletchley.timeseries, label="Bruteforce Strategy")
line_new, = plt.plot(boostedBletchley.timeseries, label="Boosted Strategy")
plt.legend(handles=[line_old, line_new])
plt.xlabel('# of cracking attempts')
plt.ylabel('# of trigrams discovered')
plt.title('Trigram Discovery Over Time');
# + [markdown] slideshow={"slide_type": "fragment"}
# We see that the boosted fuzzer is constantly superior over the random fuzzer.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# ## Estimating the Probability of Path Discovery
#
# <!-- ## Residual Risk: Probability of Failure after an Unsuccessful Fuzzing Campaign -->
# <!-- Residual risk is not formally defined in this section, so I made the title a bit more generic -- AZ -->
#
# So, what does Turing's observation for the Naval Enigma have to do with fuzzing _arbitrary_ programs? Turing's assistant I.J. Good extended and published Turing's work on the estimation procedures in Biometrica, a journal for theoretical biostatistics that still exists today. Good did not talk about trigrams. Instead, he calls them "species". Hence, the GT estimator is presented to estimate how likely it is to discover a new species, given an existing sample of individuals (each of which belongs to exactly one species).
#
# Now, we can associate program inputs to species, as well. For instance, we could define the path that is exercised by an input as that input's species. This would allow us to _estimate the probability that fuzzing discovers a new path._ Later, we will see how this discovery probability estimate also estimates the likelihood of discovering a vulnerability when we have not seen one, yet (residual risk).
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"}
# Let's do this. We identify the species for an input by computing a hash-id over the set of statements exercised by that input. In the [Coverage](Coverage.ipynb) chapter, we have learned about the [Coverage class](Coverage.ipynb#A-Coverage-Class) which collects coverage information for an executed Python function. As an example, the function [`cgi_decode()`](Coverage.ipynb#A-CGI-Decoder) was introduced. The function `cgi_decode()` takes a string encoded for a website URL and decodes it back to its original form.
#
# Here's what `cgi_decode()` does and how coverage is computed.
# + slideshow={"slide_type": "skip"}
from Coverage import Coverage, cgi_decode
# + slideshow={"slide_type": "fragment"}
encoded = "Hello%2c+world%21"
with Coverage() as cov:
decoded = cgi_decode(encoded)
# + slideshow={"slide_type": "subslide"}
decoded
# + slideshow={"slide_type": "fragment"}
print(cov.coverage());
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Trace Coverage
# First, we will introduce the concept of execution traces, which are a coarse abstraction of the execution path taken by an input. Compared to the definition of path, a trace ignores the sequence in which statements are exercised or how often each statement is exercised.
#
# * `pickle.dumps()` - serializes an object by producing a byte array from all the information in the object
# * `hashlib.md5()` - produces a 128-bit hash value from a byte array
# + slideshow={"slide_type": "skip"}
import pickle
import hashlib
# + slideshow={"slide_type": "fragment"}
def getTraceHash(cov):
pickledCov = pickle.dumps(cov.coverage())
hashedCov = hashlib.md5(pickledCov).hexdigest()
return hashedCov
# + [markdown] slideshow={"slide_type": "subslide"}
# Remember our model for the Naval Enigma machine? Each message must be decrypted using exactly one trigram while multiple messages may be decrypted by the same trigram. Similarly, we need each input to yield exactly one trace hash while multiple inputs can yield the same trace hash.
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's see whether this is true for our `getTraceHash()` function.
# + slideshow={"slide_type": "subslide"}
inp1 = "a+b"
inp2 = "a+b+c"
inp3 = "abc"
with Coverage() as cov1:
cgi_decode(inp1)
with Coverage() as cov2:
cgi_decode(inp2)
with Coverage() as cov3:
cgi_decode(inp3)
# + [markdown] slideshow={"slide_type": "fragment"}
# The inputs `inp1` and `inp2` execute the same statements:
# + slideshow={"slide_type": "fragment"}
inp1, inp2
# + slideshow={"slide_type": "fragment"}
cov1.coverage() - cov2.coverage()
# + [markdown] slideshow={"slide_type": "subslide"}
# The difference between both coverage sets is empty. Hence, the trace hashes should be the same:
# + slideshow={"slide_type": "fragment"}
getTraceHash(cov1)
# + slideshow={"slide_type": "fragment"}
getTraceHash(cov2)
# + slideshow={"slide_type": "fragment"}
assert getTraceHash(cov1) == getTraceHash(cov2)
# + [markdown] slideshow={"slide_type": "fragment"}
# In contrast, the inputs `inp1` and `inp3` execute _different_ statements:
# + slideshow={"slide_type": "fragment"}
inp1, inp3
# + slideshow={"slide_type": "fragment"}
cov1.coverage() - cov3.coverage()
# + [markdown] slideshow={"slide_type": "fragment"}
# Hence, the trace hashes should be different, too:
# + slideshow={"slide_type": "subslide"}
getTraceHash(cov1)
# + slideshow={"slide_type": "fragment"}
getTraceHash(cov3)
# + slideshow={"slide_type": "fragment"}
assert getTraceHash(cov1) != getTraceHash(cov3)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Measuring Trace Coverage over Time
# In order to measure trace coverage for a `function` executing a `population` of fuzz inputs, we slightly adapt the `population_coverage()` function from the [Chapter on Coverage](Coverage.ipynb#Coverage-of-Basic-Fuzzing).
# + slideshow={"slide_type": "subslide"}
def population_trace_coverage(population, function):
cumulative_coverage = []
all_coverage = set()
cumulative_singletons = []
cumulative_doubletons = []
singletons = set()
doubletons = set()
for s in population:
with Coverage() as cov:
try:
function(s)
except BaseException:
pass
cur_coverage = set([getTraceHash(cov)])
# singletons and doubletons -- we will need them later
doubletons -= cur_coverage
doubletons |= singletons & cur_coverage
singletons -= cur_coverage
singletons |= cur_coverage - (cur_coverage & all_coverage)
cumulative_singletons.append(len(singletons))
cumulative_doubletons.append(len(doubletons))
# all and cumulative coverage
all_coverage |= cur_coverage
cumulative_coverage.append(len(all_coverage))
return all_coverage, cumulative_coverage, cumulative_singletons, cumulative_doubletons
# + [markdown] slideshow={"slide_type": "subslide"}
# Let's see whether our new function really contains coverage information only for *two* traces given our three inputs for `cgi_decode`.
# + slideshow={"slide_type": "fragment"}
all_coverage = population_trace_coverage([inp1, inp2, inp3], cgi_decode)[0]
assert len(all_coverage) == 2
# + [markdown] slideshow={"slide_type": "fragment"}
# Unfortunately, the `cgi_decode()` function is too simple. Instead, we will use the original Python [HTMLParser](https://docs.python.org/3/library/html.parser.html) as our test subject.
# + slideshow={"slide_type": "skip"}
from Fuzzer import RandomFuzzer
from Coverage import population_coverage
from html.parser import HTMLParser
# + slideshow={"slide_type": "fragment"}
trials = 50000 # number of random inputs generated
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's run a random fuzzer for $n=50000$ times and plot trace coverage over time.
# + slideshow={"slide_type": "fragment"}
# create wrapper function
def my_parser(inp):
parser = HTMLParser() # resets the HTMLParser object for every fuzz input
parser.feed(inp)
# + slideshow={"slide_type": "subslide"}
# create random fuzzer
fuzzer = RandomFuzzer(min_length=1, max_length=100,
char_start=32, char_range=94)
# create population of fuzz inputs
population = []
for i in range(trials):
population.append(fuzzer.fuzz())
# execute and measure trace coverage
trace_timeseries = population_trace_coverage(population, my_parser)[1]
# execute and measure code coverage
code_timeseries = population_coverage(population, my_parser)[1]
# plot trace coverage over time
plt.figure(num=None, figsize=(12, 4), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(1, 2, 1)
plt.plot(trace_timeseries)
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of traces exercised')
plt.title('Trace Coverage Over Time')
# plot code coverage over time
plt.subplot(1, 2, 2)
plt.plot(code_timeseries)
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of statements covered')
plt.title('Code Coverage Over Time');
# + [markdown] slideshow={"slide_type": "subslide"}
# Above, we can see trace coverage (left) and code coverage (right) over time. Here are our observations.
# 1. **Trace coverage is more robust**. There are less sudden jumps in the graph compared to code coverage.
# 2. **Trace coverage is more fine grained.** There more traces than statements covered at the end (y-axis)
# 3. **Trace coverage grows more steadily**. Code coverage exercise more than half the statements with the first input that it exercises after 50k inputs. Instead, the number of traces covered grows slowly and steadily since each input can yield only one execution trace.
#
# It is for this reason that one of the most prominent and successful fuzzers today, american fuzzy lop (AFL), uses a similar *measure of progress* (a hash computed over the branches exercised by the input).
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Evaluating the Discovery Probability Estimate
#
# Let's find out how the Good-Turing estimator performs as estimate of discovery probability when we are fuzzing to discover execution traces rather than trigrams.
#
# To measure the empirical probability, we execute the same population of inputs (n=50000) and measure in regular intervals (measurement=100 intervals). During each measurement, we repeat the following experiment repeats=500 times, reporting the average: If the next input yields a new trace, return 1, otherwise return 0. Note that during these repetitions, we do not record the newly discovered traces as observed.
# + slideshow={"slide_type": "fragment"}
repeats = 500 # experiment repetitions
measurements = 100 # experiment measurements
# + slideshow={"slide_type": "subslide"}
emp_timeseries = []
all_coverage = set()
step = int(trials / measurements)
for i in range(0, trials, step):
if i - step >= 0:
for j in range(step):
inp = population[i - j]
with Coverage() as cov:
try:
my_parser(inp)
except BaseException:
pass
all_coverage |= set([getTraceHash(cov)])
discoveries = 0
for _ in range(repeats):
inp = fuzzer.fuzz()
with Coverage() as cov:
try:
my_parser(inp)
except BaseException:
pass
if getTraceHash(cov) not in all_coverage:
discoveries += 1
emp_timeseries.append(discoveries / repeats)
# + [markdown] slideshow={"slide_type": "subslide"}
# Now, we compute the Good-Turing estimate over time.
# + slideshow={"slide_type": "fragment"}
gt_timeseries = []
singleton_timeseries = population_trace_coverage(population, my_parser)[2]
for i in range(1, trials + 1, step):
gt_timeseries.append(singleton_timeseries[i - 1] / i)
# + [markdown] slideshow={"slide_type": "fragment"}
# Let's go ahead and plot both time series.
# + slideshow={"slide_type": "fragment"}
line_emp, = plt.semilogy(emp_timeseries, label="Empirical")
line_gt, = plt.semilogy(gt_timeseries, label="Good-Turing")
plt.legend(handles=[line_emp, line_gt])
plt.xticks(range(0, measurements + 1, int(measurements / 5)),
range(0, trials + 1, int(trials / 5)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('discovery probability')
plt.title('Discovery Probability Over Time');
# + [markdown] slideshow={"slide_type": "subslide"}
# Again, the Good-Turing estimate appears to be *highly accurate*. In fact, the empirical estimator has a much lower precision as indicated by the large swings. You can try and increase the number of repetitions (repeats) to get more precision for the empirical estimates, however, at the cost of waiting much longer.
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Discovery Probability Quantifies Residual Risk
#
# Alright. You have gotten a hold of a couple of powerful machines and used them to fuzz a software system for several months without finding any vulnerabilities. Is the system vulnerable?
#
# Well, who knows? We cannot say for sure; there is always some residual risk. Testing is not verification. Maybe the next test input that is generated reveals a vulnerability.
#
# Let's say *residual risk* is the probability that the next test input reveals a vulnerability that has not been found, yet. Böhme \cite{stads} has shown that the Good-Turing estimate of the discovery probability is also an estimate of the maxmimum residual risk.
#
# **Proof sketch (Residual Risk)**. Here is a proof sketch that shows that an estimator of discovery probability for an arbitrary definition of species gives an upper bound on the probability to discover a vulnerability when none has been found: Suppose, for each "old" species A (here, execution trace), we derive two "new" species: Some inputs belonging to A expose a vulnerability while others belonging to A do not. We know that _only_ species that do not expose a vulnerability have been discovered. Hence, _all_ species exposing a vulnerability and _some_ species that do not expose a vulnerability remain undiscovered. Hence, the probability to discover a new species gives an upper bound on the probability to discover (a species that exposes) a vulnerability. **QED**.
#
# An estimate of the discovery probability is useful in many other ways.
#
# 1. **Discovery probability**. We can estimate, at any point during the fuzzing campaign, the probability that the next input belongs to a previously unseen species (here, that it yields a new execution trace, i.e., exercises a new set of statements).
# 2. **Complement of discovery probability**. We can estimate the proportion of *all* inputs the fuzzer can generate for which we have already seen the species (here, execution traces). In some sense, this allows us to quantify the *progress of the fuzzing campaign towards completion*: If the probability to discovery a new species is too low, we might as well abort the campaign.
# 3. **Inverse of discovery probability**. We can predict the number of test inputs needed, so that we can expect the discovery of a new species (here, execution trace).
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## How Do We Know When to Stop Fuzzing?
#
# In fuzzing, we have measures of progress such as [code coverage](Coverage.ipynb) or [grammar coverage](GrammarCoverageFuzzer.ipynb). Suppose, we are interested in covering all statements in the program. The _percentage_ of statements that have already been covered quantifies how "far" we are from completing the fuzzing campaign. However, sometimes we know only the _number_ of species $S(n)$ (here, statements) that have been discovered after generating $n$ fuzz inputs. The percentage $S(n)/S$ can only be computed if we know the _total number_ of species $S$. Even then, not all species may be feasible.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ### A Success Estimator
#
# If we do not _know_ the total number of species, then let's at least _estimate_ it: As we have seen before, species discovery slows down over time. In the beginning, many new species are discovered. Later, many inputs need to be generated before discovering the next species. In fact, given enough time, the fuzzing campaign approaches an _asymptote_. It is this asymptote that we can estimate.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# In 1984, <NAME>, a well-known theoretical bio-statistician, has developed an estimator $\hat S$ which estimates the asymptotic total number of species $S$:
# \begin{align}
# \hat S_\text{Chao1} = \begin{cases}
# S(n) + \frac{f_1^2}{2f_2} & \text{if $f_2>0$}\\
# S(n) + \frac{f_1(f_1-1)}{2} & \text{otherwise}
# \end{cases}
# \end{align}
# * where $f_1$ and $f_2$ is the number of singleton and doubleton species, respectively (that have been observed exactly once or twice, resp.), and
# * where $S(n)$ is the number of species that have been discovered after generating $n$ fuzz inputs.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# So, how does Chao's estimate perform? To investigate this, we generate trials=400000 fuzz inputs using a fuzzer setting that allows us to see an asymptote in a few seconds. We measure trace coverage coverage. After half-way into our fuzzing campaign (trials/2=100000), we generate Chao's estimate $\hat S$ of the asymptotic total number of species. Then, we run the remainer of the campaign to see the "empirical" asymptote.
# + slideshow={"slide_type": "fragment"}
trials = 400000
fuzzer = RandomFuzzer(min_length=2, max_length=4,
char_start=32, char_range=32)
population = []
for i in range(trials):
population.append(fuzzer.fuzz())
_, trace_ts, f1_ts, f2_ts = population_trace_coverage(population, my_parser)
# + slideshow={"slide_type": "subslide"}
time = int(trials / 2)
time
# + slideshow={"slide_type": "fragment"}
f1 = f1_ts[time]
f2 = f2_ts[time]
Sn = trace_ts[time]
if f2 > 0:
hat_S = Sn + f1 * f1 / (2 * f2)
else:
hat_S = Sn + f1 * (f1 - 1) / 2
# + [markdown] slideshow={"slide_type": "fragment"}
# After executing `time` fuzz inputs (half of all), we have covered these many traces:
# + slideshow={"slide_type": "fragment"}
time
# + slideshow={"slide_type": "subslide"}
Sn
# + [markdown] slideshow={"slide_type": "fragment"}
# We can estimate there are this many traces in total:
# + slideshow={"slide_type": "fragment"}
hat_S
# + [markdown] slideshow={"slide_type": "fragment"}
# Hence, we have achieved this percentage of the estimate:
# + slideshow={"slide_type": "fragment"}
100 * Sn / hat_S
# + [markdown] slideshow={"slide_type": "fragment"}
# After executing `trials` fuzz inputs, we have covered these many traces:
# + slideshow={"slide_type": "fragment"}
trials
# + slideshow={"slide_type": "fragment"}
trace_ts[trials - 1]
# + [markdown] slideshow={"slide_type": "subslide"}
# The accuracy of Chao's estimator is quite reasonable. It isn't always accurate -- particularly at the beginning of a fuzzing campaign when the [discovery probability](WhenIsEnough.ipynb#Measuring-Trace-Coverage-over-Time) is still very high. Nevertheless, it demonstrates the main benefit of reporting a percentage to assess the progress of a fuzzing campaign towards completion.
#
# ***Try it***. *Try setting and `trials` to 1 million and `time` to `int(trials / 4)`.*
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ### Extrapolating Fuzzing Success
# <!-- ## Cost-Benefit Analysis: Extrapolating the Number of Species Discovered -->
#
# Suppose you have run the fuzzer for a week, which generated $n$ fuzz inputs and discovered $S(n)$ species (here, covered $S(n)$ execution traces). Instead, of running the fuzzer for another week, you would like to *predict* how many more species you would discover. In 2003, <NAME> and her team developed an extrapolation methodology to do just that. We are interested in the number $S(n+m^*)$ of species discovered if $m^*$ more fuzz inputs were generated:
#
# \begin{align}
# \hat S(n + m^*) = S(n) + \hat f_0 \left[1-\left(1-\frac{f_1}{n\hat f_0 + f_1}\right)^{m^*}\right]
# \end{align}
# * where $\hat f_0=\hat S - S(n)$ is an estimate of the number $f_0$ of undiscovered species, and
# * where $f_1$ the number of singleton species, i.e., those we have observed exactly once.
#
# The number $f_1$ of singletons, we can just keep track of during the fuzzing campaign itself. The estimate of the number $\hat f_0$ of undiscovered species, we can simply derive using Chao's estimate $\hat S$ and the number of observed species $S(n)$.
#
# Let's see how Chao's extrapolator performs by comparing the predicted number of species to the empirical number of species.
# + slideshow={"slide_type": "subslide"}
prediction_ts = [None] * time
f0 = hat_S - Sn
for m in range(trials - time):
prediction_ts.append(Sn + f0 * (1 - (1 - f1 / (time * f0 + f1)) ** m))
# + slideshow={"slide_type": "subslide"}
plt.figure(num=None, figsize=(12, 3), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(1, 3, 1)
plt.plot(trace_ts, color='white')
plt.plot(trace_ts[:time])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of traces exercised')
plt.subplot(1, 3, 2)
line_cur, = plt.plot(trace_ts[:time], label="Ongoing fuzzing campaign")
line_pred, = plt.plot(prediction_ts, linestyle='--',
color='black', label="Predicted progress")
plt.legend(handles=[line_cur, line_pred])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of traces exercised')
plt.subplot(1, 3, 3)
line_emp, = plt.plot(trace_ts, color='grey', label="Actual progress")
line_cur, = plt.plot(trace_ts[:time], label="Ongoing fuzzing campaign")
line_pred, = plt.plot(prediction_ts, linestyle='--',
color='black', label="Predicted progress")
plt.legend(handles=[line_emp, line_cur, line_pred])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of traces exercised');
# + [markdown] slideshow={"slide_type": "subslide"}
# The prediction from Chao's extrapolator looks quite accurate. We make a prediction at $time=trials/4$. Despite an extrapolation by 3 times (i.e., at trials), we can see that the predicted value (black, dashed line) closely matches the empirical value (grey, solid line).
#
# ***Try it***. Again, try setting and `trials` to 1 million and `time` to `int(trials / 4)`.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Lessons Learned
#
# * One can measure the _progress_ of a fuzzing campaign (as species over time, i.e., $S(n)$).
# * One can measure the _effectiveness_ of a fuzzing campaign (as asymptotic total number of species $S$).
# * One can estimate the _effectiveness_ of a fuzzing campaign using the Chao1-estimator $\hat S$.
# * One can extrapolate the _progress_ of a fuzzing campaign, $\hat S(n+m^*)$.
# * One can estimate the _residual risk_ (i.e., the probability that a bug exists that has not been found) using the Good-Turing estimator $GT$ of the species discovery probability.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Next Steps
#
# This chapter is the last in the book! If you want to continue reading, have a look at the [Appendices](99_Appendices.ipynb). Otherwise, _make use of what you have learned and go and create great fuzzers and test generators!_
# + [markdown] slideshow={"slide_type": "slide"}
# ## Background
#
# * A **statistical framework for fuzzing**, inspired from ecology. <NAME>. [STADS: Software Testing as Species Discovery](https://mboehme.github.io/paper/TOSEM18.pdf). ACM TOSEM 27(2):1--52
# * Estimating the **discovery probability**: I.<NAME>. 1953. [The population frequencies of species and the
# estimation of population parameters](https://www.jstor.org/stable/2333344). Biometrika 40:237–264.
# * Estimating the **asymptotic total number of species** when each input can belong to exactly one species: <NAME>. 1984. [Nonparametric estimation of the number of classes in a population](https://www.jstor.org/stable/4615964). Scandinavian Journal of Statistics 11:265–270
# * Estimating the **asymptotic total number of species** when each input can belong to one or more species: <NAME>. 1987. [Estimating the population size for capture-recapture data with unequal catchability](https://www.jstor.org/stable/2531532). Biometrics 43:783–791
# * **Extrapolating** the number of discovered species: <NAME>, <NAME>, and <NAME>. 2003. [Predicting the Number of New Species in Further Taxonomic Sampling](http://chao.stat.nthu.edu.tw/wordpress/paper/2003_Ecology_84_P798.pdf). Ecology 84, 3 (2003), 798–804.
# + [markdown] button=false new_sheet=true run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## Exercises
# <NAME> <NAME> developed an estimator for the case where each input belongs to exactly one species. For instance, each input yields exactly one execution trace (see function [`getTraceHash`](#Trace-Coverage)). However, this is not true in general. For instance, each input exercises multiple statements and branches in the source code. Generally, each input can belong to one *or more* species.
#
# In this extended model, the underlying statistics are quite different. Yet, all estimators that we have discussed in this chapter turn out to be almost identical to those for the simple, single-species model. For instance, the Good-Turing estimator $C$ is defined as
# $$C=\frac{Q_1}{n}$$
# where $Q_1$ is the number of singleton species and $n$ is the number of generated test cases.
# Throughout the fuzzing campaign, we record for each species the *incidence frequency*, i.e., the number of inputs that belong to that species. Again, we define a species $i$ as *singleton species* if we have seen exactly one input that belongs to species $i$.
# + [markdown] slideshow={"slide_type": "fragment"} solution2="shown" solution2_first=true
# ### Exercise 1: Estimate and Evaluate the Discovery Probability for Statement Coverage
#
# In this exercise, we create a Good-Turing estimator for the simple fuzzer.
# + [markdown] slideshow={"slide_type": "fragment"} solution2="shown" solution2_first=true
# #### Part 1: Population Coverage
#
# Implement a function `population_stmt_coverage()` as in [the section on estimating discovery probability](#Estimating-the-Discovery-Probability) that monitors the number of singletons and doubletons over time, i.e., as the number $i$ of test inputs increases.
# + slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
from Coverage import population_coverage, Coverage
...
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Here we go:
# + slideshow={"slide_type": "skip"} solution2="hidden"
def population_stmt_coverage(population, function):
cumulative_coverage = []
all_coverage = set()
cumulative_singletons = []
cumulative_doubletons = []
singletons = set()
doubletons = set()
for s in population:
with Coverage() as cov:
try:
function(s)
except BaseException:
pass
cur_coverage = cov.coverage()
# singletons and doubletons
doubletons -= cur_coverage
doubletons |= singletons & cur_coverage
singletons -= cur_coverage
singletons |= cur_coverage - (cur_coverage & all_coverage)
cumulative_singletons.append(len(singletons))
cumulative_doubletons.append(len(doubletons))
# all and cumulative coverage
all_coverage |= cur_coverage
cumulative_coverage.append(len(all_coverage))
return all_coverage, cumulative_coverage, cumulative_singletons, cumulative_doubletons
# + [markdown] slideshow={"slide_type": "fragment"} solution2="shown" solution2_first=true
# #### Part 2: Population
#
# Use the random `fuzzer(min_length=1, max_length=1000, char_start=0, char_range=255)` from [the chapter on Fuzzers](Fuzzer.ipynb) to generate a population of $n=10000$ fuzz inputs.
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
from Fuzzer import RandomFuzzer
from html.parser import HTMLParser
...
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** This is fairly straightforward:
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
trials = 2000 # increase to 10000 for better convergences. Will take a while..
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We create a wrapper function...
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
def my_parser(inp):
parser = HTMLParser() # resets the HTMLParser object for every fuzz input
parser.feed(inp)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# ... and a random fuzzer:
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
fuzzer = RandomFuzzer(min_length=1, max_length=1000,
char_start=0, char_range=255)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We fill the population:
# + cell_style="split" slideshow={"slide_type": "skip"} solution2="hidden"
population = []
for i in range(trials):
population.append(fuzzer.fuzz())
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
# #### Part 3: Estimating Probabilities
#
# Execute the generated inputs on the Python HTML parser (`from html.parser import HTMLParser`) and estimate the probability that the next input covers a previously uncovered statement (i.e., the discovery probability) using the Good-Turing estimator.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Here we go:
# + slideshow={"slide_type": "skip"} solution2="hidden"
measurements = 100 # experiment measurements
step = int(trials / measurements)
gt_timeseries = []
singleton_timeseries = population_stmt_coverage(population, my_parser)[2]
for i in range(1, trials + 1, step):
gt_timeseries.append(singleton_timeseries[i - 1] / i)
# + [markdown] slideshow={"slide_type": "fragment"} solution2="hidden" solution2_first=true
# #### Part 4: Empirical Evaluation
#
# Empirically evaluate the accuracy of the Good-Turing estimator (using $10000$ repetitions) of the probability to cover new statements using the experimental procedure at the end of [the section on estimating discovery probability](#Estimating-the-Discovery-Probability).
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** This is as above:
# + slideshow={"slide_type": "skip"} solution2="hidden"
# increase to 10000 for better precision (less variance). Will take a while..
repeats = 100
# + slideshow={"slide_type": "skip"} solution2="hidden"
emp_timeseries = []
all_coverage = set()
for i in range(0, trials, step):
if i - step >= 0:
for j in range(step):
inp = population[i - j]
with Coverage() as cov:
try:
my_parser(inp)
except BaseException:
pass
all_coverage |= cov.coverage()
discoveries = 0
for _ in range(repeats):
inp = fuzzer.fuzz()
with Coverage() as cov:
try:
my_parser(inp)
except BaseException:
pass
# If intersection not empty, a new stmt was (dis)covered
if cov.coverage() - all_coverage:
discoveries += 1
emp_timeseries.append(discoveries / repeats)
# + slideshow={"slide_type": "skip"} solution2="hidden"
# %matplotlib inline
import matplotlib.pyplot as plt
line_emp, = plt.semilogy(emp_timeseries, label="Empirical")
line_gt, = plt.semilogy(gt_timeseries, label="Good-Turing")
plt.legend(handles=[line_emp, line_gt])
plt.xticks(range(0, measurements + 1, int(measurements / 5)),
range(0, trials + 1, int(trials / 5)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('discovery probability')
plt.title('Discovery Probability Over Time');
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="shown" solution2_first=true solution_first=true
# ### Exercise 2: Extrapolate and Evaluate Statement Coverage
#
# In this exercise, we use Chao's extrapolation method to estimate the success of fuzzing.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# #### Part 1: Create Population
#
# Use the random `fuzzer(min_length=1, max_length=1000, char_start=0, char_range=255)` to generate a population of $n=400000$ fuzz inputs.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Here we go:
# + slideshow={"slide_type": "skip"} solution2="hidden"
trials = 400 # Use 400000 for actual solution. This takes a while!
# + slideshow={"slide_type": "skip"} solution2="hidden"
population = []
for i in range(trials):
population.append(fuzzer.fuzz())
_, stmt_ts, Q1_ts, Q2_ts = population_stmt_coverage(population, my_parser)
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# #### Part 2: Compute Estimate
#
# Compute an estimate of the total number of statements $\hat S$ after $n/4=100000$ fuzz inputs were generated. In the extended model, $\hat S$ is computed as
# \begin{align}
# \hat S_\text{Chao1} = \begin{cases}
# S(n) + \frac{Q_1^2}{2Q_2} & \text{if $Q_2>0$}\\
# S(n) + \frac{Q_1(Q_1-1)}{2} & \text{otherwise}
# \end{cases}
# \end{align}
# * where $Q_1$ and $Q_2$ is the number of singleton and doubleton statements, respectively (i.e., statements that have been exercised by exactly one or two fuzz inputs, resp.), and
# * where $S(n)$ is the number of statements that have been (dis)covered after generating $n$ fuzz inputs.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Here we go:
# + slideshow={"slide_type": "skip"} solution2="hidden"
time = int(trials / 4)
Q1 = Q1_ts[time]
Q2 = Q2_ts[time]
Sn = stmt_ts[time]
if Q2 > 0:
hat_S = Sn + Q1 * Q1 / (2 * Q2)
else:
hat_S = Sn + Q1 * (Q1 - 1) / 2
print("After executing %d fuzz inputs, we have covered %d **(%.1f %%)** statements.\n" % (time, Sn, 100 * Sn / hat_S) +
"After executing %d fuzz inputs, we estimate there are %d statements in total.\n" % (time, hat_S) +
"After executing %d fuzz inputs, we have covered %d statements." % (trials, stmt_ts[trials - 1]))
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "subslide"} solution="hidden" solution2="hidden" solution2_first=true solution_first=true toc-hr-collapsed=false
# #### Part 3: Compute and Evaluate Extrapolator
#
# Compute and evaluate Chao's extrapolator by comparing the predicted number of statements to the empirical number of statements.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** Here's our solution:
# + slideshow={"slide_type": "skip"} solution2="hidden"
prediction_ts = [None] * time
Q0 = hat_S - Sn
for m in range(trials - time):
prediction_ts.append(Sn + Q0 * (1 - (1 - Q1 / (time * Q0 + Q1)) ** m))
# + slideshow={"slide_type": "skip"} solution2="hidden"
plt.figure(num=None, figsize=(12, 3), dpi=80, facecolor='w', edgecolor='k')
plt.subplot(1, 3, 1)
plt.plot(stmt_ts, color='white')
plt.plot(stmt_ts[:time])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of statements exercised')
plt.subplot(1, 3, 2)
line_cur, = plt.plot(stmt_ts[:time], label="Ongoing fuzzing campaign")
line_pred, = plt.plot(prediction_ts, linestyle='--',
color='black', label="Predicted progress")
plt.legend(handles=[line_cur, line_pred])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of statements exercised')
plt.subplot(1, 3, 3)
line_emp, = plt.plot(stmt_ts, color='grey', label="Actual progress")
line_cur, = plt.plot(stmt_ts[:time], label="Ongoing fuzzing campaign")
line_pred, = plt.plot(prediction_ts, linestyle='--',
color='black', label="Predicted progress")
plt.legend(handles=[line_emp, line_cur, line_pred])
plt.xticks(range(0, trials + 1, int(time)))
plt.xlabel('# of fuzz inputs')
plt.ylabel('# of statements exercised');
|
docs/notebooks/WhenToStopFuzzing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="39203a85-c522-4e5a-a9a2-b8551e5dfe12" _uuid="e584282e2739b57f39ecdbac9381f66b4a15c102"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
from subprocess import check_output
print(check_output(["ls", "../input"]).decode("utf8"))
# Any results you write to the current directory are saved as output.
# + [markdown] _cell_guid="1116e340-646b-467b-990b-cec660407918" _uuid="71775b009fd5a354bf17db8c0943b2f9c79e9b20"
# ### **Loading dataset into pandas dataframe**
# + _cell_guid="e9cb07cb-4252-4072-ba98-4d1bae11ba4a" _uuid="4e7d26e7dddbf23c16f7f15791c963a8e0fe68d7"
df_card = pd.read_csv('../input/creditcard.csv')
# + [markdown] _cell_guid="41fbfbed-b689-4132-9bff-fba9d03b5849" _kg_hide-input=true _uuid="d6d83e03d73b01ab729b391463fccf0b83545a5c"
# ### Head of dataset
# + _cell_guid="495230c9-cbe8-4d2a-9c4c-aabb7bda4718" _uuid="de853733bce38ba735e4659ebc737de4a2fbb72a"
df_card.head()
# + [markdown] _cell_guid="21f28f1f-226d-46a2-b6ec-040aa7d779f1" _uuid="9015d3248d301add9e5849103166020e3182a468"
# ### Describing the dataset
# + _cell_guid="71c359e9-2aa8-400e-ba9c-0ec3abd6bbc6" _uuid="a9ecba985bc503afeaec8bfb046570bbe3ba3217"
df_card[['Time', 'Amount']].describe()
# + [markdown] _cell_guid="13b10600-88f3-4885-ba36-9854de5939e0" _uuid="b1a846265125efb8186971195f2820b856d89174"
# #### Checking the data types of dataframe columns
# + _cell_guid="ef2445fd-fcc8-4945-84b1-296cfa38efbc" _uuid="95afaacdd304fce997ac7172510be2bf18160519"
df_card.info()
# + [markdown] _cell_guid="eb848b44-55f6-48e6-96a0-0c1652c27ee2" _uuid="327c166a02c2ef86ff208b6812caeec33b653c51"
# ### Very imbalanced classes. There are a lot more regular transactions than fraudulent
#
# 1. 0 284315
# 2. 1 492
# + _cell_guid="39afb00c-9926-4c96-a1d7-b1b7f5d0fa75" _uuid="7a0fbfd8f4583157f59a8b9e4047e80d443902be"
df_card['Class'].value_counts()
# + _cell_guid="32b778b6-58a2-4c2f-82ca-acc466011066" _uuid="b1bec69641eb8a6c2bc605e7e1d8f6ac1d83b598"
print("Fraudulent transactions account for {:.2f}% of the dataset"
.format(df_card['Class'].value_counts()[1]/len(df_card)*100))
# + [markdown] _cell_guid="549ba82b-5ecc-4611-85ad-b0a5092e91b3" _uuid="4ff21d6e2930a68f186e3b7b9549baac9fdba9aa"
# ### Fraudulent transactions have a slightly higher mean value
# + _cell_guid="e07d569b-caca-4e50-8691-ef2da8ef8a72" _uuid="338778f6ea8a379533bd01250cf7cb1efa7d3181"
df_card[['Amount', 'Class']].groupby('Class').mean()
# + [markdown] _cell_guid="5bb70a47-8252-497f-93d2-59a65ab25893" _uuid="d8b614133afe6eef837487f7bd31c78d77885bc5"
# ### Even though regular transactions have a higher transaction amount
# + _cell_guid="8d9c6b77-5129-46df-9082-8ba6322367c8" _uuid="d4a054e15ab6127f5e2eb1b6596a5ed02aeab084"
df_card[['Amount', 'Class']].groupby('Class').max()
# + [markdown] _cell_guid="f0cb3fc1-6210-4872-b5d1-bc4d0c38100d" _uuid="9448e8e01501f3334bbfb8660a2f4821807274f2"
# ### The transaction amount that repeated the most is a very small value: **$1.00**. Would it be fraudulent or not?
# + _cell_guid="523c676c-413c-49cb-96f8-e3a160189612" _uuid="5422a633abe975a96ae039db64cb3c01d019a837"
df_card['Amount'].value_counts()
# + [markdown] _cell_guid="01b32f92-625c-41c6-8444-7dac76bc0c6f" _uuid="72a50d2014991cb8126c6c07d211fbbb6c849b20"
# ### Sorting the fraudulent transactions by Amount
# + _cell_guid="4c4c5b7b-0349-4326-a7a0-763730dbdba9" _uuid="9196524e461e25f7c235224b6b76296883c69550"
df_card[df_card['Class'] == 1][['Amount', 'Class']].sort_values(by='Amount', ascending=False).head(10)
# + [markdown] _cell_guid="996cccd4-fc5b-4a7c-87c3-49ebd5b5c650" _uuid="cba021d97f89cadc41d0b879af9ee61cfbf49628"
# ### Now we can check that the most repeated amount for fraudulent transactions is $1.00! This could indicate that this is just a "checking" amount, a value used to test if the transaction is approved.
#
# + _cell_guid="8fe81dc6-07bd-4aaa-970e-973cbb389ace" _uuid="cb42e9564bc37554f3d3ce31e4f670ed8b144eda"
df_card[df_card['Class'] == 1][['Amount', 'Class']]['Amount'].value_counts()
# + [markdown] _cell_guid="71435950-3f8d-42e7-af9c-d1bffdae4fc2" _uuid="d2e7cb2bd90ea3c9bd2e69b9fae60c9deb676369"
# ### **Exploratory Data Analysis (EDA)**
# + _cell_guid="d32ed84b-917e-4f76-9d95-e18f729e411e" _uuid="af2ebb8ef0d382959b7e1e9de65072c6f030aa98"
def get_transactions_average():
## Fraudulent transactions mean
fraudulent_transactions_mean = df_card[df_card['Class'] == 1]['Amount'].mean()
## Regular transactions mean
normal_transactions_mean = df_card[df_card['Class'] == 0]['Amount'].mean()
## Creating an array with the mean values
return [fraudulent_transactions_mean, normal_transactions_mean]
# + _cell_guid="7a38d38d-0583-4707-bc69-5a895e914ac0" _uuid="eb0927c5d0b892e664cdc62177374fff16098de6"
# Get the mean values for each transaction type
mean_arr = get_transactions_average()
# Calculate the overall mean
overall_mean = df_card['Amount'].mean()
# + [markdown] _cell_guid="a69cc952-e8be-4d55-aace-f5f4caae4700" _uuid="0184c73d8b6df4e1289ff8bca9178048643f2e25"
# #### Plotting the mean values in a bar plot. We can check the regular transactions are aroud the overall mean value, but the fraudulent ones are slightly above the mean
# + _cell_guid="8d01d696-580b-4983-9de4-a139789188ce" _uuid="d9a273efeae67718b6c39cd306fabaa41dab53a5"
fig = plt.figure(figsize=(10, 8))
## Labels to replace the elements' indexes in the x-axis
xticks_labels = ['Fraudulent transactions', 'Regular transactions']
## X-axis elements
xticks_elements = [item for item in range(0,len(mean_arr))]
ax = plt.gca()
## Plot the bar char custom bar colors
plt.bar(xticks_elements, mean_arr, color='#2F4F4F')
## Map the xticks to their string descriptions, then rotate them to make them more readable
plt.xticks(xticks_elements, xticks_labels, rotation=70)
## Draw a horizontal line to show the overall mean to compare with each category's mean
plt.axhline(overall_mean, color='#e50000', animated=True, linestyle='--')
## Annotate the line to explain its purpose
ax.annotate('Overall Mean', xy=(0.5, overall_mean), xytext=(0.5, 110),
arrowprops=dict(facecolor='#e50000', shrink=0.05))
## Set the x-axis label
plt.xlabel('Transactions')
## Set the y-axis label
plt.ylabel('Average amount in $ Dollar')
## Show the plot
plt.show()
# + _cell_guid="aff62e04-6797-475f-964b-50d7e8212775" _uuid="0ac2aee82e3f5d00429538d794106af534bc5b24"
# Describing the amount values for the fraulent transactions
describe_arr = df_card[df_card['Class'] == 1]['Amount'].describe()
describe_arr
# + _cell_guid="ed1f919d-c706-4122-b210-b0146120666e" _uuid="2135f5c3bd3904ee609b4e63340b0d7d564fc9a6"
## Creates a new figure
plt.figure(figsize=(10, 8))
## Filter out the fraudulent transactions from dataframe
df_fraudulent = df_card[df_card['Class']==1]
## Creates a boxplot from fraudulent transactions data
sns.boxplot(x="Class", y="Amount",
data=df_fraudulent, palette='muted')
## Most values are clustered around small values, but the max transactions amount is smaller
## than those from regular transactions
# + _cell_guid="4eea03a4-e824-425d-bd4d-4146d5c908ab" _uuid="5cec461b6fc4bb930dcf6c03f894aacc9337721b"
## Creates a new figure
plt.figure(figsize=(10, 8))
## Filter out the normal transactions from dataframe
df_regular = df_card[df_card['Class']==0]
## Creates a boxplot from the regular transactions data
sns.boxplot(x="Class", y="Amount",
data=df_regular, palette='muted')
## Most transactions are grouped around small amounts
# + _cell_guid="d38daa15-07e6-4a6a-92a7-33cbf6dd2c58" _uuid="e6872309137c360bd7da178289bfc81ebd729993"
## Creates a new figure
plt.figure(figsize=(10, 8))
## Draw a distribution plot (histogram) from amount values
sns.distplot(df_card['Amount'], kde=True, hist=True, norm_hist=True)
## Check that most of the transactions are clustered around small values
# + _cell_guid="47e7c822-88eb-40f3-ad08-720880cf4641" _uuid="889ea271edeb76d2769f7acc38f61fa9ecc31a78"
## Creates a new figure
plt.figure(figsize=(10, 8))
## Draw a distribution plot (histogram) from fraudulent transactions data
sns.distplot(df_fraudulent['Amount'], kde=True, hist=True, norm_hist=True)
## Check that most transactions are clustered around $0 and $500.
# + _cell_guid="c9e18a47-0549-41e2-b492-0f35f925a1b3" _uuid="a5233bc2a79dd56d4d3df2ec45ecce599cfac59b"
df_card.head()
# + [markdown] _cell_guid="a86ba050-5bc0-4d25-9413-52c1c0aca07c" _uuid="e00b7d845e05dce259350224b65ed63e83206616"
# ### Preparing data for model
# + _cell_guid="0af15315-d27b-4e39-959d-7fa623af0f85" _uuid="945a02041db57f14e5657c798d1416014cb559d7"
## Dataset split import
from sklearn.model_selection import train_test_split
# + _cell_guid="e1cb42fa-2625-4503-a79c-d45a3beaa906" _uuid="e51fb0debd3b2edcbf651cddcecb69d37e011364"
## Scale the amount feature before fitting the models
sc= StandardScaler()
df_card["scaled_amount"]= sc.fit_transform(df_card.iloc[:,29].values.reshape(-1,1))
## Drops the old amount, once the scaled one has been added to the dataframe
df_card.drop('Amount', axis=1, inplace=True)
# + _cell_guid="e5252f03-1f11-4c65-8919-9dab412f254e" _kg_hide-input=false _kg_hide-output=false _uuid="c86890cbf5171c319b464597b8c7a27b39ae0f03"
## Set the features to the X variable
X = df_card.drop(['Time', 'Class'], axis=1)
## Set the target column to the y_target variable
y_target = df_card['Class']
# + [markdown] _cell_guid="e16cc161-ccba-46f9-87b9-5af7f5a32511" _uuid="b1f902aaf1089ae7975bbdd6e48d12eadd360a8c"
# ### Models
# + _cell_guid="4fb6ef5e-68ed-4436-922a-5803dba58e30" _uuid="4b14ba480692d3de425b1382084b902f3528e029"
## Models and evaluation metrics imports
from sklearn.ensemble import RandomForestClassifier
from sklearn import svm
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import recall_score
from sklearn.metrics import roc_curve, auc, roc_auc_score, average_precision_score
from sklearn.metrics import classification_report
from sklearn.metrics import accuracy_score
from sklearn.model_selection import GridSearchCV
# + [markdown] _cell_guid="135c9186-fb2e-4791-9d51-f553c5c5caa6" _uuid="ce6fbdcdf3d3bb2f2015b957c135cd6ff4c88d97"
# ## Model utility functions
# + _cell_guid="844d0a2e-d6e3-465b-a848-9586ee14651b" _uuid="22af00e0c6645b10970c17154b0b32cd512b907b"
## Split the data into train and test set
X_train, X_test, y_train, y_test = train_test_split(X, y_target, random_state=42)
# + _cell_guid="1708b51b-a8af-4d90-9642-4fc59733882d" _uuid="cbd09425fe20c24de4af3d8df231a110e89c1531"
## This is a generic function to calculate the auc score which is used several times in this notebook
def evaluate_model_auc(model, X_test_parameter, y_test_parameter):
## The predictions
y_pred = model.predict(X_test_parameter)
## False positive rate, true positive rate and treshold
fp_rate, tp_rate, treshold = roc_curve(y_test_parameter, y_pred)
## Calculate the auc score
auc_score = auc(fp_rate, tp_rate)
## Returns the score to the model
return (auc_score)
# + _cell_guid="d91205ce-0b59-46f4-944f-a72752141a80" _uuid="9b5f22b5f5f097c15c1c72d52af309330769a19f"
## This is a generic function to plot the area under the curve (AUC) for a model
def plot_auc(model, X_test, y_test):
## Predictions
y_pred = model.predict(X_test)
## Calculates auc score
fp_rate, tp_rate, treshold = roc_curve(y_test, y_pred)
auc_score = auc(fp_rate, tp_rate)
## Creates a new figure and adds its parameters
plt.figure()
plt.title('ROC Curve')
## Plot the data - false positive rate and true positive rate
plt.plot(fp_rate, tp_rate, 'b', label = 'AUC = %0.2f' % auc_score)
plt.legend(loc = 'lower right')
plt.plot([0, 1], [0, 1],'r--')
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel('True Positive Rate')
plt.xlabel('False Positive Rate')
# + _cell_guid="b19a9ea8-cd6a-42c9-8fc9-bd9dff867d20" _uuid="8f5ac14473e98e9d6fda04907f0e42a1958d832b"
## This is a generic utility function to calculate a model's score
def evaluate_model_score(model, X_test, y_test):
## Return the score value to the model
return model.score(X_test, y_test)
# + _cell_guid="31663779-e5db-49d5-92f0-4127f0061f47" _uuid="459874839b65154aa9dcaca8f957cc5607e2459e"
## This is a generic function to create a classification report and return it to the model. The target
## variables have been mapped to the transaction types
def evaluate_classification_report(model, y_test):
return classification_report(y_test, model.predict(X_test), target_names=['Regular transaction',
'Fraudulent transaction'])
# + _cell_guid="e87fc4ae-d4e5-4252-958c-217267e98528" _uuid="7d1fe4fd834e7e3d975b13043a8e5f6fc857fa3c"
## This utility function evaluates a model using some common metrics such as accurary and auc. Also, it
## prints out the classification report for the specific model
def evaluate_model(model_param, X_test_param, y_test_param):
print("Model evaluation")
print("Accuracy: {:.5f}".format(evaluate_model_score(model_param, X_test_param, y_test_param)))
print("AUC: {:.5f}".format(evaluate_model_auc(model_param, X_test_param, y_test_param)))
print("\n#### Classification Report ####\n")
print(evaluate_classification_report(model_param, y_test_param))
plot_auc(model_param, X_test_param, y_test_param)
# + _cell_guid="c297810e-385e-4128-ae5f-d7dc281fcba8" _uuid="6c46c99041fb306ec936d160e699b876cc745d2e"
## This is a shared function used to print out the results of a gridsearch process
def gridsearch_results(gridsearch_model):
print('Best score: {} '.format(gridsearch_model.best_score_))
print('\n#### Best params ####\n')
print(gridsearch_model.best_params_)
# + _cell_guid="e111983a-bf50-4813-be3b-34caf7842374" _uuid="cf07d2a537d93402c462fd1711756f020f4d5ff7"
# Returns the Random Forest model which the n_estimators returns the highest score in order to improve
# the results of the default classifier
# min_estimator - min number of estimators to run
# max_estimator - max number of estimators to run
# X_train, y_train, X_test, y_test - splitted dataset
# scoring function: accuracy or auc
def model_selection(min_estimator, max_estimator, X_train_param, y_train_param,
X_test_param, y_test_param, scoring='accuracy'):
scores = []
## Returns the classifier with highest accuracy score
if (scoring == 'accuracy'):
for n in range(min_estimator, max_estimator):
rfc_selection = RandomForestClassifier(n_estimators=n, random_state=42).fit(X_train_param, y_train_param)
score = evaluate_model_score(rfc_selection, X_test_param, y_test_param)
print('Number of estimators: {} - Score: {:.5f}'.format(n, score))
scores.append((rfc_selection, score))
## Returns the classifier with highest auc score
elif (scoring == 'auc'):
for n in range(min_estimator, max_estimator):
rfc_selection = RandomForestClassifier(n_estimators=n, random_state=42).fit(X_train_param, y_train_param)
score = evaluate_model_auc(rfc_selection, X_test_param, y_test_param)
print('Number of estimators: {} - AUC: {:.5f}'.format(n, score))
scores.append((rfc_selection, score))
return sorted(scores, key=lambda x: x[1], reverse=True)[0][0]
# + [markdown] _cell_guid="6d912c8a-bc8d-4c08-9a57-4a31b6b8391b" _uuid="c4661f5620cb85a4b4b53505d947626bf940a53f"
#
# ### **Dealing with imbalanced classes**
#
# + _cell_guid="cfc97131-8bab-4a96-a0e6-a1949de1f74c" _uuid="283f7be1ea52096749724c8560c74e9b4f0c5e2f"
## Importing SMOTE
from imblearn.over_sampling import SMOTE
## Importing resample
from sklearn.utils import resample
# + [markdown] _cell_guid="12428303-46a0-4036-906b-c6cf0035aa9b" _uuid="ea1a1cb0a3ea58b7b30dfd00af8514a2f589ff28"
# ### SMOTE
# Models like RFC and SVC have a parameter that penalizes imbalanced datasets in order to get more accurate results. However, we are going to balance the data using a technique called SMOTE to create synthetic data points from the monirity class using KNearest Neighbors.
# + _cell_guid="3880baba-b58d-4a4b-973b-f9f421538f52" _uuid="53cb66e20a820e6367b3b6a1b51240f5d12e3626"
## Making a copy of the dataset (could've been done using df.copy())
dataset = df_card[df_card.columns[1:]]
## Defines the features to the dataset_features variable
dataset_features = dataset.drop(['Class'], axis=1)
## Defines the target feature to the dataset_target variable
dataset_target = dataset['Class']
# + _cell_guid="863296c5-bb95-4966-97bc-9a0c3e22b879" _uuid="5103d179a8f671fb71b343db3d175100884ba2de"
## Split the data once again
X_train_2, X_test_2, y_train_2, y_test_2 = train_test_split(dataset_features,
dataset_target,
random_state=42)
# + _cell_guid="0924705e-e6e7-4295-af69-9b2d8510ee58" _uuid="91dbc0ccd4aeffab9b081a26e3b213fe8b2e0645"
## This function generates a balanced X_train and y_train from the original dataset to fit the model
def get_balanced_train_data(df):
sm = SMOTE(random_state=42, ratio = 1.0)
X_train_res, y_train_res = sm.fit_sample(X_train_2, y_train_2)
## Returns balanced X_train & y_train
return (X_train_res, y_train_res)
# + _cell_guid="66f6d1c3-b267-4025-8a3c-097655f7e745" _uuid="7288001e9fced6dd89d681c31e480b05c758e2b3"
## Calling the function to get scalled training data
(X_train_resampled, y_train_resampled) = get_balanced_train_data(df_card)
# + [markdown] _cell_guid="107dc535-8a32-4619-9d80-ac3fd4a47207" _uuid="5533c967659f0513deecdc0bf22ab2353faa971a"
# ## **SVM**
# With default parameters
# + _cell_guid="3353b172-d368-4dda-9a6e-73b75378e0ed" _uuid="aef335d6a6fe3dc436aad43a666b76547e780e79"
## Creating a SVC model with default parameters
svc = svm.SVC()
svc.fit(X_train_2, y_train_2)
# + _cell_guid="832659bb-c4c7-489b-9d31-9cff4587fc63" _uuid="d380eff6e5c1f1eef92dd7fc8602eab5ed481a6a"
## Evaluating the model
evaluate_model(svc, X_test_2, y_test_2)
# + [markdown] _cell_guid="0c36c91f-f73c-42a5-ab6f-a5b807c6f13a" _uuid="a726ed25d5272b136e556a941534f54fa2ca9abf"
# ### Cross validation with parameter tuning
# Setting parameters
# + _cell_guid="aadf3b7b-2e3c-4a06-a55c-64df95b68cb8" _uuid="b0ed0553481e043851ecb99eaec8ca59837c3cc6"
## Parameters grid to be tested on the model
parameters = {
'C': [1, 5, 10, 15],
'degree':[1, 2, 3, 5],
'kernel': ['linear'],
'class_weight': ['balanced', {0:1, 1:10}, {0:1, 1:15}, {0:1, 1:20}],
'gamma': [0.01, 0.001, 0.0001, 0.00001]
}
# + _cell_guid="42bcadaf-d1a4-4292-a2ea-4fe35c00d8dc" _uuid="33540e7cc5726a6e351c23a34011217df28d54e5"
## Creates a gridsearch to find the best parameters for this dataset.
clf = GridSearchCV(estimator=svm.SVC(random_state=42),
## Passes the parameter grid as argument (these parameters will be tested
## when this model is created)
param_grid=parameters,
## Run the processes in all CPU cores
n_jobs=-1,
## Set the scoring method to 'roc_auc'
scoring='roc_auc')
# + _cell_guid="ea5089dd-ebff-4dc5-986a-5ba67f6337fc" _uuid="7fa8c7cea0771d9c8d9851c37813e1865a01712b"
## Fit the gridsearch model to the data
# clf.fit(X_train_2[:5000], y_train_2[:5000])
# + _cell_guid="733794cc-d521-4c41-bfd7-4d685f8d327d" _uuid="1d728b144ba1143d3f76f5296d21ec72bb152c17"
## Find the model with the best score achieved and the best parameters to use
# gridsearch_results(clf)
# + [markdown] _cell_guid="d0970f03-03d1-4c46-ad08-e2cdd9d0e4cb" _uuid="81210862b556fda446f758542c5abdfd2fcf2749"
# ### Using the optimal parameters
# + _cell_guid="c099a148-54cf-41af-9665-50f66b1c0a01" _uuid="a6feb4eaacc2a3403a612a80d3bddc2377588a46"
## Creates a SVC model with the optimal parameters found in the previous step
svc_grid_search = svm.SVC(C=1,
kernel='linear',
degree=1,
class_weight={0:1, 1:10},
gamma=0.01,
random_state=42)
svc_grid_search.fit(X_train_2[:5000], y_train_2[:5000])
# + _cell_guid="0b8c0357-7d52-42d4-afc5-d5fb2e9365fc" _uuid="d8044d78e881f8100d506137fb0e5b2c6af6c4b1"
## Evaluate the model
evaluate_model(svc_grid_search, X_test_2, y_test_2)
# + [markdown] _cell_guid="88aafdda-1f1a-4a6d-8106-c03252957de2" _uuid="98728f4c9f627cacb1a2256e76c3bfbb185f522d"
# ## **Random Forest Classifier**
# With default parameters
# + _cell_guid="bff1d2fa-cc91-4a25-b812-9041b13ef960" _uuid="80f05eb1cd4b0fc02b2dc58db5c41daf87c38e27"
## Creates a Random Forest Classifier with default parameters
model_rfc = RandomForestClassifier().fit(X_train_2, y_train_2)
# + _cell_guid="5c6f6dfc-c7e5-4721-9aa6-e33132b19c78" _uuid="47ca9ceac0d954d02946f91124f01866f5e0db7c"
## Evaluate the model
evaluate_model(model_rfc, X_test, y_test)
# + [markdown] _cell_guid="f1e41691-7e7e-43c8-a6e7-4acce304d605" _uuid="7a4ae00d4ce2b8e698cf92f65766fba7bfabfb81"
# ### Random Forest Classifier presentes a good performance right out of the box, but can we improve it? Let's test using the balanced dataset using parameter tuning
# + [markdown] _cell_guid="c13d13d4-73d1-4010-9c23-8e184f8aff87" _uuid="0a155f520506b7ace65560c67ab3072bf265a2bb"
# ### Selecting a model with best # of estimators
# + _cell_guid="ce5ad4d9-77a9-42eb-9483-a0a834e102f6" _uuid="b309b09cf143a524cfee4dd760df6d9843a2d64c"
## Creating a model selecting the best number of estimators
rfc_model = model_selection(5, 15, X_train, y_train, X_test, y_test, scoring='auc')
# + _cell_guid="e837de0b-6435-4209-91df-22bc0295d821" _uuid="024b13151d09f5ac3af6e6ce4c5a200c9b61b5c0"
## Evaluate the model
evaluate_model(rfc_model, X_test, y_test)
# + [markdown] _cell_guid="f99c1cb0-ba2b-46ab-ac2c-30a58de11604" _uuid="11b9147643777350cf482f22f02c08db02f064db"
# ### Training with balanced dataset
# + _cell_guid="5d181b50-189c-44e3-b38d-2f91dae41c71" _uuid="0f7ea01bd5b5cf486ce4866925f7809ed16d4c61"
## Select the model with the best number of estimators using the balanced dataset
rfc_smote = model_selection(5, 15, X_train_resampled, y_train_resampled,
X_test_2, y_test_2, scoring='auc')
# + _cell_guid="3fc0636d-b7b7-4b48-85fd-cf81134b8354" _uuid="74bc785edcf30c854afdad718c592190d9f10451"
## Evaluate the model with AUC metric
evaluate_model(rfc_smote, X_test_2, y_test_2)
# + _cell_guid="df7d00c5-d83e-4d27-bccf-44be5560e51c" _uuid="c52ab5c366d6265947905a18ef2060b0e79aa3d6"
## Show the most important features from the dataset
sorted(rfc_smote.feature_importances_, reverse=True)[:5]
# + _cell_guid="84171613-c24d-4e2d-b63f-2e1a3230b437" _uuid="0ed95ed137e25ce4970e3381940c77b0c271be11"
## Itemgetter import
from operator import itemgetter
# + _cell_guid="565fbdc8-4be9-4b1d-a6e0-c614dc5b5617" _uuid="3b338c01d03e73c047db0f8d6bcb6f738aad6b52"
## Loading features and importance
features = [i for i in X.columns.values]
importance = [float(i) for i in rfc_smote.feature_importances_]
feature_importance = []
## Creating a list of tuples concatenating feature names and its importance
for item in range(0, len(features)):
feature_importance.append((features[item], importance[item]))
## Sorting the list
feature_importance.sort(key=itemgetter(1), reverse=True)
## Printing the top 5 most important features
feature_importance[:5]
# + [markdown] _cell_guid="22a33e3c-b9a4-40bc-8cae-0897c0f3ac9b" _uuid="6c799a712bf0baa6365db46953a1230f7b13ed26"
# ### **Grid search random forest**
# + _cell_guid="ef6ea9a5-abf0-40f5-8a2b-bf72129853b4" _uuid="171ea1a6e1eced74d97b538fd859dbb7c0965d79"
## Parameters to use with the RFC model
parameters_rfc = {
'n_estimators': [5, 6, 7, 8, 9, 10, 13, 15],
# 'class_weight': ['balanced'],
'max_depth': [None, 5, 10, 15, 20, 25, 30, 35, 40],
'min_samples_leaf': [1, 2, 3, 4, 5]
}
# + _cell_guid="d4e588d2-5bd5-45c0-b054-ba948bb3b1f5" _uuid="fc5628af09e612642bc11a0715effa691f325772"
## Gridsearch to get the best parameters for RFC
rfc_grid_search = GridSearchCV(estimator=RandomForestClassifier(random_state=42,
n_jobs=-1),
param_grid=parameters_rfc,
cv=10,
scoring='roc_auc',
return_train_score=True)
# + _cell_guid="9e5893d3-1900-4bed-93c2-c09212f6c956" _uuid="d274e7c1a3b6418e98970975071ad4c98dd32d5e"
## Train the gridsearch model
## Using only part of the dataset because the entire data takes too long to train; the same
## applies to the other models
## Takes too long
rfc_grid_search.fit(X_train_2[:10000], y_train_2[:10000])
# + _cell_guid="c0f8049d-abb8-4b9f-93f9-80272370421c" _uuid="bc37f5c966bfd0f7f09b59dff7139e955262f2bc"
## Check the results of cross validation
cv_results = pd.DataFrame(rfc_grid_search.cv_results_)
## Sort the values to get the best result
cv_results.sort_values(by='rank_test_score').head()
# + _cell_guid="0632c791-b233-47c7-91d7-3972bbf5402d" _uuid="fe9d7ae0e591897260d0d15ab18c2f35db49757b"
## Model with the best score and the best parameters
#
gridsearch_results(rfc_grid_search)
# + [markdown] _cell_guid="fb4ce92c-27b4-45ea-b58e-e87fbed60950" _uuid="633ab09b5ed3f889c7f1dffefef140981534d99d"
# ### Training model with optimal parameters
# + _cell_guid="1f9e0827-79e9-4b90-b718-dba1817e65c0" _uuid="cf5e994e8e4f26a430fb1311c4dfd1a0303a0e4f"
## RFC model using the parameters found by gridsearch
rfc = RandomForestClassifier(random_state=42,
n_estimators=7, min_samples_leaf=1, max_depth=5)
## Fit the data
rfc.fit(X_train_2, y_train_2)
# + _cell_guid="1bad5481-bd9d-431c-b469-3232badde7c7" _uuid="ac38cc17b3662518a43dc3643c44b617bd8455aa"
## Evaluate the model
evaluate_model(rfc, X_test_2, y_test_2)
# + [markdown] _cell_guid="b97501e2-81bc-4808-b088-767e757d769c" _uuid="a57d9e947f94ab4f4e000ca4cd07c1b9b150ab1b"
# ### With balanced dataset
# + _cell_guid="3399adcf-6e3d-447a-9040-440c1d6ea814" _uuid="2987d2aff6beba783da670c747c90b211bc70d13"
## Running gridsearch again to find the best results for the scalled dataset
rfc_grid_search_balanced = GridSearchCV(estimator=RandomForestClassifier(random_state=42,
n_jobs=-1),
param_grid=parameters_rfc,
cv=10,
scoring='roc_auc',
return_train_score=True)
# + _cell_guid="0c749e10-39b1-4ba1-8cf3-2849cbef3140" _uuid="804a2a8684a0feb4bb46c717372cfbfea0fe7c49"
## Fitting the data
## Takes too long
# rfc_grid_search_balanced.fit(X_train_resampled[:5000], y_train_resampled[:5000])
# + _cell_guid="413740e1-2633-4b66-8c87-6e0b786e4e43" _uuid="a635c1d4b6d34e6d959b48b4ae1c387d2924ee18"
## Best score and best parameters
#gridsearch_results(rfc_grid_search_balanced)
#13 4 None
# + [markdown] _cell_guid="3d294e08-579f-432d-854f-cd38f408f4da" _uuid="e757478383a0f683c11eaf27063742ce45507446"
# It takes too long to run the gridsearch process on this model for this dataset. For this reason, I decided to run on my local machine and wait until the process finishes. After completed, I obtained the following parameters:
# 1. **n_estimators**: 13
# 2. **min_samples_leaf**: 4
# 3. **max_depth**: default (None)
#
# Note that a lower **max_depth** parameter will lower the precision for the 'balanced' dataset.
# + [markdown] _cell_guid="c175a953-d78c-48f3-93bf-ecaeeaf4c93e" _uuid="0a68bfb6081c6f966f9c619792d8cc49139b7b0a"
# ### Training model with optimal parameters
# + _cell_guid="c05a266a-1a01-4323-9287-5eb431a9baf2" _uuid="cbe91d14d3b14ba52a81d09dfb5e1dae9eac108b"
## Creating a new model with the selected parameters
rfc_balanced = RandomForestClassifier(random_state=42,
n_estimators=13, min_samples_leaf=4, max_depth=None)
rfc_balanced.fit(X_train_resampled, y_train_resampled)
# + _cell_guid="38a68c66-7dfb-4b15-8919-f5913fb95ec2" _uuid="c30f0a57bedce12f049eddee8bf8d26e3e95b09d"
## Evaluate the model
evaluate_model(rfc_balanced, X_test_2, y_test_2)
# + [markdown] _cell_guid="63b1c289-2b8b-4f99-9968-f1f6899f2dfc" _uuid="5124b0f4fa0cad3a9e4ee7714d2fdec2ed3f5f21"
# ## **Logistic regression**
# + _cell_guid="65f17c16-78eb-4a70-bd99-37cfdaed5687" _uuid="dfbb91b302f864a5cba8950e1eb83856b49382b3"
## Parameters grid for Logistic Regression model
param_grid_lreg = {
'C': [0.001, 0.01, 0.1, 1, 10, 15],
'class_weight': ['balanced', {0:1, 0:10}, {0:1, 1:15}, {0:1, 1:20}],
'penalty': ['l1', 'l2']
}
# + _cell_guid="2dc474f6-b59b-48ec-9a7b-7a7e8c516676" _uuid="e24e74962fbf9110eb1a04184ec0830e5b60a860"
## Running gridsearch to find best parameters for Logistic Regression model
lreg_grid_search = GridSearchCV(estimator=LogisticRegression(random_state=42),
param_grid=param_grid_lreg, cv=10, scoring='roc_auc')
# + _cell_guid="7311480f-90a1-442d-b095-f31ca12634f6" _uuid="a005f89ff41b14e2ef165d3030d8797b4bca028e"
## Fitting the data (it takes a long time)
# lreg_grid_search.fit(X_train_2[:2000], y_train_2[:2000])
# + _cell_guid="c9e1a672-4fd3-4784-8ee2-6855876c3bb1" _uuid="d143f542c9ddbabf58d193d13e67ba71a92742c5"
## Printing the best results for this model
# gridsearch_results(lreg_grid_search)
# + [markdown] _cell_guid="0af4aa70-2f2e-4f5d-90f0-2289c5597117" _uuid="145b23064ce103dbdf2d64dd1b1330781de40197"
# ### Parameter tuning
# + _cell_guid="077ee7bf-6db5-4f58-aedb-6abe56dafbdb" _uuid="75b93b4d82b4fd98f28b3165adf0956bc650c9d8"
## Creating a model with gridsearch parameters
lreg = LogisticRegression(C=1, penalty='l1', random_state=42,
class_weight={0:1, 1:10})
## Fitting the model
lreg.fit(X_train_2, y_train_2)
# + _cell_guid="446246ea-e9ae-47f5-9588-940be7d51a35" _uuid="aeab858318ea562a0a91dcacd69964fec00d5ab8"
## Evaluate the model
evaluate_model(lreg, X_test_2, y_test_2)
# + [markdown] _cell_guid="29e0e16f-872f-4a8b-bdeb-985fe3f33887" _uuid="999d2376473f881ab54a70ab785254656ad13d84"
# ## Final words
# As you can see, we can get good results with some models out of the box such as Random Forest Classifier. However, the imbalanced nature of this dataset might impact the overall result. For this reason, I've tried several classifier algorithms along with different parameter tuning and a varied set of evaluation metrics in order to achieve stable results.
#
# This is just an example of how to use some basic machine learning techniques such as: data manipulation, EDA, data scaling, balancing (SMOTE), gridsearch, cross validation, and model evaluation.
# + _cell_guid="b0bd2dbe-1997-476f-a106-a753a66a20a6" _uuid="aaffe7e3638cdb50edbf2db0d307b4e011032373"
|
Classification/0.94 AUC with imbalanced dataset/Credit card fraud detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.10 64-bit (''PythonDataConda'': conda)'
# name: python3710jvsc74a57bd0ac97be953db092153337d196dff28cd269b2245e0bae87498e32021eb0bea4fd
# ---
# ## Observations and Insights
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as st
import numpy as np
# Study data files
mouse_metadata_path = "./data/Mouse_metadata.csv"
study_results_path = "./data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
combined_data = pd.merge(mouse_metadata, study_results, on="Mouse ID", how="outer")
combinded_data = pd.DataFrame(combined_data)
# Display the data table for preview
combined_data.head()
# -
# Checking the number of mice.
combined_data.count()
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
duplicates = combined_data[combined_data.duplicated(subset=["Mouse ID", "Timepoint"], keep=False)]
duplicates
# Checking the number of mice in the clean DataFrame.
cleaned_data = combined_data.drop_duplicates(subset=["Mouse ID", "Timepoint"], keep="first")
#cleaned_data.count()
len(cleaned_data["Mouse ID"].unique())
# ## Summary Statistics
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
trimmed_data = cleaned_data.loc[:, ["Drug Regimen", "Tumor Volume (mm3)"]]
grouped_data = trimmed_data.groupby("Drug Regimen")
# mean, median, variance, standard deviation, and SEM of the tumor volume.
average = round(grouped_data.mean(), 2)
middle = round(grouped_data.median(), 2)
standard_dev = round(grouped_data.std(), 2)
variance = round(standard_dev**2)
standard_error = round(grouped_data.sem(), 2)
# Assemble the resulting series into a single summary dataframe.
summary_Tumor_Volume = pd.DataFrame({"Average": average["Tumor Volume (mm3)"], "Median": middle["Tumor Volume (mm3)"],"Variance": variance["Tumor Volume (mm3)"], "Standard Deviation": standard_dev["Tumor Volume (mm3)"], "SEM": standard_error["Tumor Volume (mm3)"]})
summary_Tumor_Volume.head()
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Using the aggregation method, produce the same summary statistics in a single line
grouped_data.aggregate({"Tumor Volume (mm3)": ['mean', 'median', 'std', 'var', 'sem']})
# ## Bar and Pie Charts
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
sample_size = cleaned_data["Drug Regimen"].value_counts()
sample_size = pd.DataFrame(sample_size)
sample_size = sample_size.reset_index(0)
sample_size = sample_size.rename(columns={"index":"Drug Regimen", "Drug Regimen":"Sample Size"})
sample_size
#x_values = np.arange(0, sample_size["Drug Regimen"].count(), 1)
sample_size.plot.bar(x="Drug Regimen", y="Sample Size", rot=45)
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
x_values = np.arange(0, sample_size["Sample Size"].count(), 1)
Drug_Regimen= sample_size["Drug Regimen"]
plt.xticks(x_values, Drug_Regimen, rotation=45)
plt.bar(x_values, sample_size["Sample Size"], color="blue", alpha=.25)
plt.ylim(0, max(sample_size["Sample Size"])+25)
plt.ylabel("Sample Size")
plt.xlabel("Drug Regimen")
plt.title("Drug Trial Sizes")
plt.show()
# Generate a pie plot showing the distribution of female versus male mice using pandas
gender_proportion = cleaned_data["Sex"].value_counts()
gender_proportion.plot.pie(title="Gender Proportion", explode=(0.1, 0), autopct="%1.1f%%", shadow=True)
# Generate a pie plot showing the distribution of female versus male mice using pyplot
plt.pie(gender_proportion, labels=["Male", "Female"], explode=(0.1,0), autopct="%1.1f%%", shadow=True)
plt.title("Gender Proportion")
plt.show()
# ## Quartiles, Outliers and Boxplots
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
four_regimens = cleaned_data.loc[(cleaned_data["Drug Regimen"]=='Capomulin') | (cleaned_data["Drug Regimen"]=='Ramicane') | (cleaned_data["Drug Regimen"]=='Infubinol') | (cleaned_data["Drug Regimen"]=='Ceftamin'), ["Mouse ID", "Drug Regimen", "Timepoint", "Tumor Volume (mm3)"]]
four_regimens = four_regimens.rename(columns={"Tumor Volume (mm3)":"Vol (mm3)"})
# Get data corresponding to last (greatest) Timepoint for each mouse
four_regimens_last = four_regimens.sort_values("Timepoint").groupby("Mouse ID").tail(1)
# Group by Drug Regimen
four_regimens_grouped = four_regimens_last.groupby("Drug Regimen")
# +
# Generate a box plot of the final tumor volume of each mouse across four regimens of interest
four_regimens_grouped.boxplot(column='Vol (mm3)', layout=(1,4))
plt.show()
# +
# Calculate the IQR and quantitatively determine if there are any potential outliers.
quartiles = four_regimens_grouped['Vol (mm3)'].quantile([0.25, 0.5, 0.75]).unstack()
lowerq = quartiles[0.25]
upperq = quartiles[0.75]
IQR = upperq - lowerq
# Determine outliers using upper and lower bounds
lower_bound = lowerq - (1.5*IQR)
upper_bound = upperq + (1.5*IQR)
# sort values by timepoint, unique by mouse id, keep last
#print(four_regimens)
sorted_data = four_regimens.sort_values('Timepoint')
#print(sorted_data)
last_data = sorted_data.drop_duplicates(subset=['Mouse ID'], keep='last')
#print(last_data)
treatments=['Capomulin', 'Ramicane', 'Infubinol', 'Ceftamin']
volumes=[]
for drug in treatments:
volumes = last_data.loc[last_data['Drug Regimen']==drug, ['Vol (mm3)']]
print(f"{drug} lower outliers are {lower_outliers}")
# -
# ## Line and Scatter Plots
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
# ## Correlation and Regression
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
|
Pymaceuticals/pymaceuticals_starter.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# --------------------------
# HTML Display
# Examples of building and displaying HTML content directly in a Jupyter Notebook
# <NAME>
# <EMAIL>
# http://www.josephtrue.com/
# https://github.com/joseph-true
# --------------------------
# -
# These libraries support what we want to do
from IPython.core.display import display, HTML
import random
#
# Simple example of displaying some HTML
display(HTML('<h1>Hello, world!</h1>'))
# +
#
# Retrieve HTML from a file
f = open("html-test-file.htm", "r")
strHtml = f.read()
f.close
# Display the raw HTML from the file
strHtml
# -
# Display the HTML content from the file
display(HTML(strHtml))
#
# Build a div tag
# Color resource: https://htmlcolorcodes.com/
display(HTML('<br>'))
display(HTML('<div align="right" style="background-color:#7fb3d5;width:300px">TEST</div>'))
display(HTML('<br>'))
# +
#
# Build it dynamically blending code, values and HTML
strDynHTML = ''
for i in range(10):
x = random.randint(25,400)
# open the div tag, set div width with random number, add margin, close the div
strDynHTML = strDynHTML + '<div align="right" style="background-color:#7fb3d5;'
strDynHTML = strDynHTML + 'width:' + str(x) + 'px;>'
strDynHTML = strDynHTML + 'margin-top:2px; margin-bottom:2px;">'
strDynHTML = strDynHTML + '<b>' + str(x) + '</b>'
strDynHTML = strDynHTML + '</div>'
display(HTML(strDynHTML))
# -
|
html-display/html-display-test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + nbsphinx="hidden"
# THIS CELL SETS STUFF UP FOR DEMO / COLLAB. THIS CELL CAN BE IGNORED.
#-------------------------------------GET RID OF TF DEPRECATION WARNINGS--------------------------------------#
import warnings
warnings.filterwarnings('ignore', category=FutureWarning)
import tensorflow as tf
tf.compat.v1.logging.set_verbosity(tf.compat.v1.logging.ERROR)
#----------------------------------INSTALL PSYCHRNN IF IN A COLAB NOTEBOOK-------------------------------------#
# Installs the correct branch / release version based on the URL. If no branch is provided, loads from master.
try:
import google.colab
IN_COLAB = True
except:
IN_COLAB = False
if IN_COLAB:
import json
import re
import ipykernel
import requests
from requests.compat import urljoin
from notebook.notebookapp import list_running_servers
kernel_id = re.search('kernel-(.*).json',
ipykernel.connect.get_connection_file()).group(1)
servers = list_running_servers()
for ss in servers:
response = requests.get(urljoin(ss['url'], 'api/sessions'),
params={'token': ss.get('token', '')})
for nn in json.loads(response.text):
if nn['kernel']['id'] == kernel_id:
relative_path = nn['notebook']['path'].split('%2F')
if 'blob' in relative_path:
blob = relative_path[relative_path.index('blob') + 1]
# !pip install git+https://github.com/murraylab/PsychRNN@$blob
else:
# !pip install git+https://github.com/murraylab/PsychRNN
# -
# # Simple Example
# This example walks through the steps and options involved in setting up and training a recurrent neural network on a cognitive task.
#
# Most users will want to [define their own tasks](NewTask.ipynb), but for the purposes of getting familiar with the package features, we will use one of the [built-in tasks](../apidoc/tasks.rst#implemented-example-tasks), the 2-alternative forced choice [Perceptual Discrimination](../apidoc/tasks.rst#module-psychrnn.tasks.perceptual_discrimination) task.
#
# This example will use the [Basic](../apidoc/backend.rst#psychrnn.backend.models.basic.Basic) implementation of [RNN](../apidoc/backend.rst#psychrnn.backend.rnn.RNN). If you are new to RNNs, we recommend you stick with the Basic implementation. PsychRNN also includes [BasicScan](../apidoc/backend.rst#psychrnn.backend.models.basic.BasicScan) and [LSTM](../apidoc/backend.rst#psychrnn.backend.models.lstm.LSTM) implementations of RNN. If you want to use a different architecture, you can [define a new model](NewModel.ipynb), but that should not be necessary for most use cases.
# +
from psychrnn.tasks.perceptual_discrimination import PerceptualDiscrimination
from psychrnn.backend.models.basic import Basic
import tensorflow as tf
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# ## Initialize Task
# First we define some global parameters that we will use when setting up the task and the model:
dt = 10 # The simulation timestep.
tau = 100 # The intrinsic time constant of neural state decay.
T = 2000 # The trial length.
N_batch = 50 # The number of trials per training update.
N_rec = 50 # The number of recurrent units in the network.
name = 'basicModel' # Unique name used to determine variable scope for internal use.
pd = PerceptualDiscrimination(dt = dt, tau = tau, T = T, N_batch = N_batch) # Initialize the task object
# ## Initialize Model
#
# When we initialize the model, we pass in a dictionary of parameters that will determine how the network is set up.
# ### Set Up Network Parameters
# [PerceptualDiscrimination.get_task_params()](../apidoc/tasks.rst#psychrnn.tasks.task.Task.get_task_params) puts the passed in parameters and other generated parameters into a dictionary we can then use to initialize our Basic RNN model.
network_params = pd.get_task_params()
print(network_params)
# We add in a few params that any [RNN](../apidoc/backend.rst#psychrnn.backend.rnn.RNN) needs but that the [Task](../apidoc/tasks.rst#psychrnn.tasks.task.Task) doesn't generate for us.
network_params['name'] = name # Unique name used to determine variable scope.
network_params['N_rec'] = N_rec # The number of recurrent units in the network.
# There are some other optional parameters we can add in. Additional parameter options like those for [biological constraints](BiologicalConstraints.ipynb), [loading weights](SavingLoadingWeights.ipynb), and [other features](OtherFeatures.ipynb) are also available:
# +
network_params['rec_noise'] = 0.0 # Noise into each recurrent unit. Default: 0.0
network_params['W_in_train'] = True # Indicates whether W_in is trainable. Default: True
network_params['W_rec_train'] = True # Indicates whether W_rec is trainable. Default: True
network_params['W_out_train'] = True # Indicates whether W_out is trainable. Default: True
network_params['b_rec_train'] = True # Indicates whether b_rec is trainable. Default: True
network_params['b_out_train'] = True # Indicates whether b_out is trainable. Default: True
network_params['init_state_train'] = True # Indicates whether init_state is trainable. Default: True
network_params['transfer_function'] = tf.nn.relu # Transfer function to use for the network. Default: tf.nn.relu.
network_params['loss_function'] = "mean_squared_error"# String indicating what loss function to use. If not `mean_squared_error` or `binary_cross_entropy`, params["loss_function"] defines the custom loss function. Default: "mean_squared_error".
network_params['load_weights_path'] = None # When given a path, loads weights from file in that path. Default: None
# network_params['initializer'] = # Initializer to use for the network. Default: WeightInitializer (network_params) if network_params includes W_rec or load_weights_path as a key, GaussianSpectralRadius (network_params) otherwise.
# -
# #### Initialization Parameters
#
# When ``network_params['initializer']`` is not set, the following optional parameters will be passed to the initializer. See [WeightInitializer](../apidoc/backend.rst#psychrnn.backend.initializations.WeightInitializer) for more details. If ``network_params['W_rec']`` and ``network_params['load_weights_path']`` are not set, these parameters will be passed to the [GaussianSpectralRadius Initializer](../apidoc/backend.rst#psychrnn.backend.initializations.GaussianSpectralRadius). Not all optional parameters are shown here. See [Biological Constraints](BiologicalConstraints.ipynb) and [Loading Model with Weights](SavingLoadingWeights.ipynb#Loading-Model-with-Weights) for more options.
# +
network_params['which_rand_init'] = 'glorot_gauss' # Which random initialization to use for W_in and W_out. Will also be used for W_rec if which_rand_W_rec_init is not passed in. Options: 'const_unif', 'const_gauss', 'glorot_unif', 'glorot_gauss'. Default: 'glorot_gauss'.
network_params['which_rand_W_rec_init'] = network_params['which_rand_init'] # 'Which random initialization to use for W_rec. Options: 'const_unif', 'const_gauss', 'glorot_unif', 'glorot_gauss'. Default: which_rand_init.
network_params['init_minval'] = -.1 # Used by const_unif_init() as minval if 'const_unif' is passed in for which_rand_init or which_rand_W_rec_init. Default: -.1.
network_params['init_maxval'] = .1 # Used by const_unif_init() as maxval if 'const_unif' is passed in for which_rand_init or which_rand_W_rec_init. Default: .1.
# -
# #### Regularization Parameters
# Parameters for regularizing the loss are passed in through network_params as well. By default, there is no regularization. Below are options for regularizations to include. See [Regularizer](../apidoc/backend.rst#psychrnn.backend.regularizations.Regularizer) for details.
# +
network_params['L1_in'] = 0 # Parameter for weighting the L1 input weights regularization. Default: 0.
network_params['L1_rec'] = 0 # Parameter for weighting the L1 recurrent weights regularization. Default: 0.
network_params['L1_out'] = 0 # Parameter for weighting the L1 output weights regularization. Default: 0.
network_params['L2_in'] = 0 # Parameter for weighting the L2 input weights regularization. Default: 0.
network_params['L2_rec'] = 0 # Parameter for weighting the L2 recurrent weights regularization. Default: 0.
network_params['L2_out'] = 0 # Parameter for weighting the L2 output weights regularization. Default: 0.
network_params['L2_firing_rate'] = 0 # Parameter for weighting the L2 regularization of the relu thresholded states. Default: 0.
network_params['custom_regularization'] = None # Custom regularization function. Default: None.
# -
# ### Instantiate Model
basicModel = Basic(network_params)
# ## Train Model
# ### Set Up Training Parameters
# Set the training parameters for our model. All of the parameters below are optional.
train_params = {}
train_params['save_weights_path'] = None # Where to save the model after training. Default: None
train_params['training_iters'] = 100000 # number of iterations to train for Default: 50000
train_params['learning_rate'] = .001 # Sets learning rate if use default optimizer Default: .001
train_params['loss_epoch'] = 10 # Compute and record loss every 'loss_epoch' epochs. Default: 10
train_params['verbosity'] = False # If true, prints information as training progresses. Default: True
train_params['save_training_weights_epoch'] = 100 # save training weights every 'save_training_weights_epoch' epochs. Default: 100
train_params['training_weights_path'] = None # where to save training weights as training progresses. Default: None
train_params['optimizer'] = tf.compat.v1.train.AdamOptimizer(learning_rate=train_params['learning_rate']) # What optimizer to use to compute gradients. Default: tf.train.AdamOptimizer(learning_rate=train_params['learning_rate'])
train_params['clip_grads'] = True # If true, clip gradients by norm 1. Default: True
# Example usage of the optional fixed_weights parameter is available in the [Biological Constraints](BiologicalConstraints.ipynb#Fixed-Weights) tutorial
train_params['fixed_weights'] = None # Dictionary of weights to fix (not allow to train). Default: None
# Example usage of the optional performance_cutoff and performance_measure parameters is available in [Curriculum Learning](curriculum_learning.ipynb#Train-Models) tutorial.
train_params['performance_cutoff'] = None # If performance_measure is not None, training stops as soon as performance_measure surpases the performance_cutoff. Default: None.
train_params['performance_measure'] = None # Function to calculate the performance of the network using custom criteria. Default: None.]
# ### Train Model on Task using Training Parameters
losses, initialTime, trainTime = basicModel.train(pd, train_params)
plt.plot(losses)
plt.ylabel("Loss")
plt.xlabel("Training Iteration")
plt.title("Loss During Training")
# ## Test Model
# Get a batch of trials from the task to test the network on.
x,y,m, _ = pd.get_trial_batch()
# Plot the x value of the trial -- for the PerceptualDiscrimination, this includes two input neurons with different coherence.
plt.plot(range(0, len(x[0,:,:])*dt,dt), x[0,:,:])
plt.ylabel("Input Magnitude")
plt.xlabel("Time (ms)")
plt.title("Input Data")
plt.legend(["Input Channel 1", "Input Channel 2"])
# Run the trained model on this trial (not included in the training set).
output, state_var = basicModel.test(x)
plt.plot(range(0, len(output[0,:,:])*dt,dt),output[0,:,:])
plt.ylabel("Activity of Output Unit")
plt.xlabel("Time (ms)")
plt.title("Output on New Sample")
plt.legend(["Output Channel 1", "Output Channel 2"])
plt.plot(range(0, len(state_var[0,:,:])*dt,dt),state_var[0,:,:])
plt.ylabel("State Variable Value")
plt.xlabel("Time (ms)")
plt.title("Evolution of State Variables over Time")
# ## Get & Save Model Weights
# We can get the weights used by the model in dictionary form using [get_weights](../apidoc/backend.rst#psychrnn.backend.rnn.RNN.get_weights), or we can save the weights directly to a file using [save](../apidoc/backend.rst#psychrnn.backend.rnn.RNN.save).
# +
weights = basicModel.get_weights()
print(weights.keys())
# -
basicModel.save("./weights/saved_weights")
# ## Cleanup
# Clean up the model to clear out the tensorflow namespace
basicModel.destruct()
|
docs/notebooks/PerceptualDiscrimination.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Spark
// language: ''
// name: sparkkernel
// ---
// # This notebook shows how to manage the events table with Spark
// We first need to the kudu jars to the spark session
// %%configure -f
{ "conf": {"spark.jars.packages": "org.apache.kudu:kudu-spark2_2.11:1.4.0" }}
sc
// +
import org.apache.kudu.spark.kudu._
import org.apache.spark.sql.functions._
//we need to define a udf function for converting two strings into a map
val asMap = udf((keys:String, values: String) => keys.split("#").zip(values.split("#")).toMap)
//then we define a first dataframe df1 associated to the kudu table
val df1 = spark.sqlContext.read.options(Map("kudu.master" -> "master:7051","kudu.table" -> "Events3")).kudu
//the next dataframe contains an additional column of type Map containing a map for the two arrays: keys and values
//val df2 = df1.withColumn("tags", asMap($"attributesKeys",$"attributesValues"))
// -
val source = df1.select("source")
val res = source.distinct.collect
df1.printSchema()
source.groupBy("source").count().show()
df1.filter("source == 'http://services.infoblu.it/BITW/data?id=teamd26lkc'").groupBy("ts").count().show()
df1.filter("source == 'http://services.infoblu.it/BITW/data?id=teamd26lkc'").filter("ts == '1519994000000'")
val df_sample = df1.filter("source == 'http://services.infoblu.it/BITW/data?id=teamd26lkc' and ts == '1519994000000'")
val df_sample_with_attributes = df_sample.withColumn("tags", asMap($"attributesKeys",$"attributesValues"))
df_sample_with_attributes.take(10)
val a = SystemE
// %%cleanup -f
|
iot_ingestion_manager/notes/events.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CniKr6Fo-3iy" colab_type="text"
# ## In this notebook, we will load a CNN model called the [VGG16](https://arxiv.org/abs/1409.1556) model, with pre-trained weights on the [ImageNet](http://www.image-net.org) dataset. We will use *activation maximization* to visualize the features that this model has learnt.
# + [markdown] id="ro0Y6oUL4oC7" colab_type="text"
# ### ImageNet is an image database organized according to the [WordNet](https://wordnet.princeton.edu/) hierarchy. Each meaningful concept in WordNet is called a "synonym set" or "synset". ImageNet has more than 100,000 synsets with an average of 1000 images to illustrate each synset.
# + [markdown] id="hYp7pwZaB-bX" colab_type="text"
# ### Step 1 Install and import all dependencies. (You can ignore the error messages in the outputs.)
# + id="78qgQJpFL1kB" colab_type="code" colab={}
import warnings
warnings.filterwarnings('ignore')
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.FATAL)
# !pip install Keras-Applications
# !pip install --quiet --force-reinstall git+https://github.com/raghakot/keras-vis.git -U
# !pip install --quiet --force-reinstall scipy==1.2
# + id="Zg3twKQyMVTe" colab_type="code" colab={}
from keras.applications.vgg16 import VGG16
from vis.visualization import visualize_activation
from vis.utils import utils
from keras.models import Sequential, Model
from keras.layers import Dense, Dropout, Activation, Flatten
from keras import activations
import matplotlib.pyplot as plt
# + [markdown] id="XR7wj5UoHnzQ" colab_type="text"
# ### Step 2 First, let's understand what's inside the ImageNet dataset. Below, we are downloading synonym sets for each of the 1000 classes of images in ImageNet. The descriptions can also be viewed [here.](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a)
# + id="X8ITEW_ADrxm" colab_type="code" colab={}
import pickle
from urllib.request import urlopen
classidx_to_description_dict = pickle.load(urlopen('https://gist.githubusercontent.com/yrevar/6135f1bd8dcf2e0cc683/raw/d133d61a09d7e5a3b36b8c111a8dd5c4b5d560ee/imagenet1000_clsid_to_human.pkl'))
# + [markdown] id="YWC5I8XiCCnm" colab_type="text"
# ### Step 3 Download and load the pre-trained model. For this exercise, we are using the [VGG16](https://arxiv.org/abs/1409.1556) architecture with weights pre-trained on [ImageNet](http://www.image-net.org) dataset. Note that in the last layer (predictions dense layer), the model output is classified into 1000 classes.
# + id="Mt_py1fjuLpK" colab_type="code" colab={}
model = VGG16()
model.summary()
# + [markdown] id="QcPATVq_Lh-E" colab_type="text"
# ### Step 4 Use activation maximization to visualize the images that maximize output of each filter in the prediction layer. Prediction layer is the layer in the model with 1000 filters, each representing a separate class of dataset.
# ### Question: What features has the model learnt for class *Volcano* and *Baseball*, separately? How about other classes? Share with your table the interesting visualizations you get! Modify variable *filter_idx* to explore the features learned by different filters in the prediction layer.</br></br> You can find the *filter name* to *filter index* mapping [here.](https://gist.github.com/yrevar/942d3a0ac09ec9e5eb3a)
# + id="ULSSO23rzWIq" colab_type="code" colab={}
prediction_layer_name = "predictions" # The prediction layer. Refer model.summary() to get names of all layers.
# + id="3uvG1sHuMPAx" colab_type="code" colab={}
def get_layer_index_from_layer_name(layer_name):
for idx, layer in enumerate(model.layers):
if layer.name == layer_name:
return idx
prediction_layer_index = get_layer_index_from_layer_name(prediction_layer_name)
# + id="TSPAWCti7Bic" colab_type="code" colab={}
filter_idx = 980 # Index of filter you want to visualize. In this case 980 is Volcano. For the predictions layer, this corresponds to class indexes seen in classidx_to_description_dict
# + id="sAwUGJJ_QZAJ" colab_type="code" colab={}
original_activation = model.layers[prediction_layer_index].activation
# Modify activation of last layer to linear
model.layers[prediction_layer_index].activation = activations.linear
model = utils.apply_modifications(model)
# + id="k-CP_VI2P2Hi" colab_type="code" colab={}
fig=plt.figure(figsize=(20, 20))
print("Visualizing image for class index " + str(filter_idx) + ": " + classidx_to_description_dict[filter_idx])
img = visualize_activation(model, prediction_layer_index, filter_indices=[filter_idx], max_iter=100, tv_weight=1., lp_norm_weight=0.)
ax2 = fig.add_subplot(2, 2, 1)
plt.imshow(img)
# Restoring model to its original state
model.layers[prediction_layer_index].activation = original_activation
model = utils.apply_modifications(model)
# + [markdown] id="YZl-eBGxmoct" colab_type="text"
# ### Step 5 Try modifying variable *layer_name* to explore the features learned by different layers.
# ### Question: What features has model learnt for layer block1_conv1, block2_conv1, block3_conv1, block4_conv1 and block5_conv1, separately? Experiment with other layers yourself: From shallow to deeper layers, how are the features learnt evolving?
# + id="9YzrkSsfa2eq" colab_type="code" colab={}
layer_name = "block1_conv1" # Name of the layer whose filters you want to visualize. Refer model.summary() to get names of all layers.
layer_index = get_layer_index_from_layer_name(layer_name)
original_activation = model.layers[layer_index].activation
model.layers[layer_index].activation = activations.linear
model = utils.apply_modifications(model)
columns = 5
rows = 2
fig = plt.figure(figsize=(20, 20))
for i in range(0,10):
img = visualize_activation(model, layer_index, filter_indices=[i], max_iter=100, tv_weight=1., lp_norm_weight=0.)
fig.add_subplot(rows, columns, i+1)
plt.imshow(img)
plt.show()
# Restoring model to its original state
model.layers[layer_index].activation = original_activation
model = utils.apply_modifications(model)
# + id="cMhYjtR9u1ET" colab_type="code" colab={}
|
vgg16_interpretation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Created this notebook to play around with synthetic data generation.
# I am using this source as my inspiration: https://www.youtube.com/watch?time_continue=187&v=VJBY2eVtf7o&feature=emb_logo&ab_channel=KeithGalli
#
# Once I play with it I am using this for my tasks in FAD and also PAD project:
#
# Following steps in video I first import the modules that I need:
#
# +
#Importing modules:
import numpy as np
import pandas as pd
import datetime
import random
#now I create the columns using my example from task 4 in FAD tasks:
# creating the Numpy array
np.random.seed(42)
# creating the dataframe
womenapplied=pd.DataFrame({'Number of applicants':np.random.randint(0,10,size=30)})
womenaccepted=pd.DataFrame({'Accepted':np.random.randint(0,5,size=30)})
df=pd.DataFrame(womenapplied + womenaccepted)
#creating categorical variable for women and men applicants and accepted candidates
#source: https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html?highlight=index%20values
print(df)
# -
for i in range(50):
Department_A= random.choice(list(Department_A.keys()))
df.loc[i]=[i,Department_A]
df.to_csv('test_data.csv')
# +
#Trying the same using a np array ( not random module):
women_applied=np.random.randint(1,20, size=30)
women_accepted=np.random.randint(1,15, size=30)
men_applied=np.random.randint(1,10, size=30)
men_accepted=np.random.randint(1,20, size=30)
Data_women=women_applied, women_accepted
Data_men=men_applied,men_accepted
#Company employment divided by gender dataframe :
df = pd.DataFrame((da women_applied, women_accepted, men_applied, men_accepted),
columns=['Women applied', 'Women accepted', 'Men applied', 'Men accepted'])
print(df)
# +
# creating the dataframe
df=pd.DataFrame({'Number of applicants':np.random.randint(0,10,size=25)})
# displaying the dataframe
print(df)
# -
# *****
#
# ### Playing with synthetic data for task 4 , simpsons paradox:
#
# ***
# Following approach from this source now:
#
# https://easystats.github.io/correlation/reference/simulate_simpson.html
#
#
# %matplotlib inline
# +
#I firt import the modules needed and I create my variables:
import numpy as np
import pandas as pd
import random
import matplotlib.pyplot as plt
import seaborn as sns
#Creating Synthetic data
np.random.seed(0)
a=np.random.randint(1,25,30)#number of observations for applied women group
b=np.random.randint(1,10,30)#number of observations for women accepted in the job
print(a)
print(b)
# +
#Plotting this data
x=np.linspace( 1,30,num=30,endpoint=False)
y=a
plt.style.use('ggplot')
plt.plot(x, y)
plt.show()
# +
#Using shuffle to modify 'a'and 'b'
a_s=np.random.shuffle(a)
print(a_s)
# +
#Trying again.
#Now I am creating and plotting data for adults that tried a certain drug and got cured vs adults that were not cured.
import numpy as np
import pandas as pd
import random
import matplotlib.pyplot as plt
import seaborn as sns
np.random.seed(0)
a=np.random.randint(3,20,30)#number of observations of cured adults with drug A
b=np.random.randint(1,10,30)#number of observations of non cured adults with drug B
print(a)
print(b)
x = np.linspace(1, 30, 30)
y1=a
y2=b
sns.scatterplot(x,y1)
#plt.barchar(x, y1, y2)
#yfit = [a + b * xi for xi in x]
#plt.plot(x,y1,y2)
plt.show()
# +
#Creating and plotting data from men that applied and were accepted in role:
np.random.seed(0)
a=np.random.randint(3,15,30)#number of observations for applied women group
b=np.random.randint(5,26,30)#number of observations for women accepted in the job
x = np.linspace(1, 30, 30)
y1=a
y2=b
sns.regplot(x,y1 )
plt.show()
# +
#Using multiple subplots to expose which number of adults got cured with drug A:
#setting variables:
a=np.random.randint(5,25,100)#number of adults that were cured with Drug A
b=np.random.randint(5,68,100)#number of adults not cured with Drug A
#Plotting:
x1=np.linspace(5, 30, 100)
x2=np.linspace(5, 30, 100)
y1=np.random.randint(5,25,100)
y2=np.random.randint(5,68,100)
fig, (ax1, ax2) = plt.subplots(2, 1)
fig.suptitle('Effects on adults that tried drug A')
ax1.plot(x1, y1, 'o-')
ax1.set_ylabel('Cured')
ax2.plot(x2, y2, '.-')
ax2.set_xlabel('Count')
ax2.set_ylabel('Non cured')
plt.show()
# +
#Effects of the drug in children:
#setting variables:
a=np.random.randint(2,30,70)#number of children that were cured with Drug A
b=np.random.randint(7,15,40)#number of children not cured with Drug A
#Plotting:
x1=np.linspace(5, 30, 100)
x2=np.linspace(5, 30, 100)
y1=np.random.randint(2,30,100)
y2=np.random.randint(7,15,100)
fig, (ax1, ax2) = plt.subplots(2, 1)
fig.suptitle('Effects on children that tried drug A')
ax1.plot(x1, y1, 'o-')
ax1.set_ylabel('Cured')
ax2.plot(x2, y2, '.-')
ax2.set_xlabel('Count')
ax2.set_ylabel('Non cured')
plt.show()
# +
#combining both:
a=np.random.randint(5,25,100)#Adults cured
b=np.random.randint(2,30,100)#number of children that were cured with Drug A
#Plotting for adults cured compared with children cured:
x=np.linspace(1, 30, 100)
y=a*x+b
sns.lmplot(x, y)
plt.show()
# -
# ### Attempting this again:
#
# Now I am using an example from this source: https://www.youtube.com/watch?v=wgLUDw8eLB4&feature=emb_logo&ab_channel=singingbanana
#
# We have two drugs A and B and 100 people trying them during a testing time of 2 days.
# The first and second day drug B seem to have better results, however looking at the data from the two doys combined drugs A cures more people overall.
# Lets reproduce this and plot it using Python:
# +
#Creating the synthetic data based on this example:
import numpy as np
#Data for day 1:
#Drug A:
Drug_A=np.array([2,4,7,8,9,3,2,4,3,6,7,8,1,9,4,3,2,1,6,7])
print(Drug_A)
#Finding sum of Drug A array:
Drug_Asum=np.sum(Drug_A)
print(Drug_Asum)
# +
#Data for day 1:
#Drug B:
Drug_B=np.array([3,5,6,8,1,3,1,3,1,0])
print(Drug_B)
#Finding sum of Drug A array:
Drug_Bsum=np.sum(Drug_B)
print(Drug_Bsum)
# +
#Plotting both drugs results for day 1:
import matplotlib.pyplot as plt
#Organizing data:
x=np.linspace(1,2,100)
y1=Drug_A*x
y2=Drug_B*x
plt.bar(x,y1,y2)
plt.show()
# -
|
Drafts/Playing with synthetic data creation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IMDB | IST 652 FINAL PROJECT | <NAME> & <NAME>
# # =======================================================
# # PART 2: A - SCRAPING & SCRIPTING
# # =======================================================
# ## THE LIBRARIES
from bs4 import BeautifulSoup
import json
import csv
import pandas as pd
from urllib.parse import quote
import requests
import time
import re
# ## THE DATA
# ### Kaggle
# _Data from kaggle.com_
kaggle = pd.read_csv("movies.csv", encoding = "ISO-8859-1")
len(kaggle)
kaggle_cleaned = pd.read_csv("working_movies_usa.csv", encoding = "ISO-8859-1" )
len(kaggle_cleaned)
# ### IMDB
# _Data from scraping imdb.com (see below for details)_
imdb_707 = pd.read_csv("movies_IST707.csv", encoding = "ISO-8859-1" )
len(imdb_707)
imdb_ids = pd.read_csv("ids_from_imdb.csv", encoding = "ISO-8859-1" )
len(imdb_ids)
imdb_scores = pd.read_csv("V2_IMDB_score_data.csv", encoding = "ISO-8859-1")
len(imdb_scores)
# ### TMDB
# _Data from hitting the TMDB api (see below for details)_
# First attempt
tmdb_movies = pd.read_csv("tmdb_movies_csv.csv", encoding = "ISO-8859-1")
len(tmdb_movies)
# Taking every actor in our kaggle dataset and getting their movie credits using TMDB's api
tmdb_actors = pd.read_csv("tmdb_20k.csv", encoding = "ISO-8859-1")
len(tmdb_actors)
# Removing things without budget, revenue, production studio or genre
tmdb_actors_cleaned = pd.read_csv("tmdb_20k_cleaned.csv", encoding = "ISO-8859-1")
len(tmdb_actors_cleaned)
tmdb_v2 = pd.read_csv("tmdb_from_imdb_v2.csv", encoding = "ISO-8859-1")
len(tmdb_v2)
# ### The-Numbers
# _Data from scraping the-numbers.com (see below for details)_
tn_1 = pd.read_csv("V2_TN_budget_data_and_url.csv", encoding = "ISO-8859-1")
len(tn_1)
tn_2 = pd.read_csv("V2_TN_reports_dates.csv", encoding = "ISO-8859-1")
len(tn_2)
# ## THE SCRAPING & API CALLING
# ### IMDB
# +
# ==============================================================
# GETTING THE IMDB ID & SCORE
# ==============================================================
def get_info_from_movies(movies):
ids_for_movies_in_year = []
for i, movie in enumerate(movies):
link_with_id = movie.find('a', {'href': re.compile('/title/tt')})
imdb_id = link_with_id.attrs['href'].split('/')[2]
clean = "".join(line.strip() for line in movie.text.split("\n"))
# valiant regex attempt
# rating = re.compile('\)(.*)')
# name = re.compile('\..*\ ')
# date = re.compile('(\d{3}).')
title_rating_string = clean.split('0Rate')[0]
rating = title_rating_string.split(')')[1]
name = title_rating_string.split('.')[1].split('(')[0]
date = title_rating_string.split('(')[1].split(')')[0]
movie_dict = {
'imdb_id': imdb_id,
'name': name,
'imdb_rating': rating,
'date': date
}
ids_for_movies_in_year.append(movie_dict)
return(ids_for_movies_in_year)
def get_imdb_html(year, urlending):
url = ('https://www.imdb.com/search/title/?title_type=feature&boxoffice_gross_us=1,&release_date='+str(year)+'-01-01,'+str(year)+'-12-31&countries=us&view=simple&count=250'+urlending)
headers = {'Accept-Language': 'en-US'}
movies_html = requests.get(url.format(), headers=headers).content
soup = BeautifulSoup(movies_html, 'html.parser')
soup_main = soup.find("div", {"id": "main"})
movies_list = soup_main.find('div', class_="lister list detail sub-list")
movies = soup_main.find_all('div', class_="lister-item mode-simple")
return movies
def get_imdb_scores(year, urlending):
movies = get_imdb_html(year, urlending)
return get_info_from_movies(movies)
# +
def run_imdb_scores_script():
all_the_ids = []
for year in range(1970, 2020):
all_the_ids += get_imdb_scores(year, '')
all_the_ids += get_imdb_scores(year, '&start=251')
all_the_ids_df = pd.DataFrame(all_the_ids)
all_the_ids_df.to_csv('imdb_ids.csv')
# save a small (2018) subset
# all_the_2018_ids_df = pd.DataFrame(all_the_ids[48])
# all_the_2018_ids_df.to_csv
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# # UNCOMMENT TO RUN <3
# run_imdb_scores_script()
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# -
# ### TMDB
# +
# ==============================================================
# GETTING THE SUPPLIMENTAL INFO FROM TMBD API
# ==============================================================
# NOTE: This is a fun time capsule of how we wrote these files
# at the start of this project!! We'd like to think we've
# cleaned things up a bit!!
def write_csv(data):
df = pd.DataFrame(data)
df.to_csv('2018_movies.csv', index=False)
headers = {'Accept-Language': 'en-US'}
payload = "{}"
api_key = open("tmdb_api_key.txt")
api_key = api_key.read()
def get_tmdb_info(imdb_id_file):
all_movie_data = []
with open(imdb_id_file, encoding='utf-8') as csvfile:
movies = csv.reader(csvfile)
for movie in movies:
try:
url = "https://api.themoviedb.org/3/movie/"
thing_looking_for = movie[1]
my_api_key = "?api_key=" + api_key
full_url = url + thing_looking_for + my_api_key
res = requests.get(full_url, payload, headers=headers)
data = res.content.decode('UTF-8')
jdata = json.loads(data)
try:
title = jdata['title']
budget = jdata['budget']
genres = jdata['genres']
production_companies = jdata['production_companies']
release_date = jdata['release_date']
revenue = jdata['revenue']
profit = revenue - budget
popularity = jdata['popularity']
vote_average = jdata['vote_average']
vote_count = jdata['vote_count']
except KeyError:
title = 'NA'
budget = 'NA'
genres = 'NA'
production_companies = 'NA'
release_date = 'NA'
revenue = 'NA'
profit = 'NA'
popularity = 'NA'
vote_average = 'NA'
vote_count = 'NA'
movie_data = {
'release_date': release_date,
'title': title,
'budget': budget,
'genres': genres,
'production_companies': production_companies,
'revenue': revenue,
'profit': profit,
'popularity': popularity,
'vote_average': vote_average,
'vote_count': vote_count
}
all_movie_data.append(movie_data)
except UnicodeDecodeError:
director_data = {}
all_movie_data_df = pd.DataFrame(all_movie_data)
print(all_movie_data_df)
write_csv(all_movie_data)
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# # UNCOMMENT TO RUN <3
# get_tmdb_info('imdb_ids_2018.csv')
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# -
# ### The-Numbers
# ##### BUDGET CSV
# +
# 1. get the page (get_tn_data)
# 2. get the soup (get_soup)
# 3. get the data from the soup (get_data_from_soup)
def get_data_from_soup(soup_data):
all_movies = []
for data in soup_data[1:]:
data_array = data.text.split('\n')
movie_data = {
'num': data_array[0],
'release_date': data_array[1],
'name': data_array[2],
'production_budget': data_array[3],
'domestic_gross': data_array[4],
'worldwide_gross': data_array[5]
}
all_movies.append(movie_data)
return all_movies
def get_soup(num):
url = ('https://www.the-numbers.com/movie/budgets/all'+num)
headers = {'Accept-Language': 'en-US'}
movies_html = requests.get(url.format(), headers=headers).content
soup = BeautifulSoup(movies_html, 'html.parser')
soup_data = soup.find_all("tr")
return soup_data
def get_tn_data(num):
tn_soup = get_soup(num)
return get_data_from_soup(tn_soup)
# +
# 1. create array to house all data
# 2. quick workaround for our first page
# 3. iterate through the the-numbers url
# 4. save to df, save to csv
def run_TN_script():
all_pages = []
all_pages += get_tn_data('')
for i in range(1,59):
results = get_tn_data('/'+ str(i) + '01')
all_pages += results
all_pages_df = pd.DataFrame(all_pages)
all_pages_df.to_csv('TN_budget_data.csv')
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# # UNCOMMENT TO RUN <3
# run_TN_script()
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# -
# ##### REPORTS CSV
# +
def format_data(data_array):
movie_data = {
'Released': data_array[2],
'ReleasedWorldwide': data_array[3],
'Released_2': data_array[4],
'ReleasedWorldwide_2': data_array[5],
'Title': data_array[6],
'TheatricalDistributor': data_array[7],
'Genre': data_array[8],
'Source': data_array[9],
'ProductionMethod': data_array[10],
'CreativeType': data_array[11],
'ProductionBudget': data_array[12],
'OpeningWeekendTheaters': data_array[13],
'MaximumTheaters': data_array[14],
'TheatricalEngagements': data_array[15],
'OpeningWeekendRevenue': data_array[16],
'DomesticBoxOffice': data_array[17],
'Infl.Adj.Dom.BoxOffice': data_array[18],
'InternationalBoxOffice': data_array[19],
'WorldwideBoxOffice': data_array[20]
}
return movie_data
def get_report(year):
url = ('https://www.the-numbers.com/movies/report/All/All/All/All/All/All/All/All/All/.1/None/'+str(year)+'/'+ str(year + 1)+'/None/None/None/None/None/None?show-release-date=On&view-order-by=domestic-box-office&show-release-year=On&view-order-direction=desc&show-production-budget=On&show-opening-weekend-theaters=On&show-domestic-box-office=On&show-maximum-theaters=On&show-inflation-adjusted-domestic-box-office=On&show-theatrical-engagements=On&show-international-box-office=On&show-opening-weekend-revenue=On&show-worldwide-box-office=On&show-worldwide-release-date=On&show-worldwide-release-year=On&show-theatrical-distributor=On&show-genre=On&show-source=On&show-production-method=On&show-creative-type=On')
headers = {'Accept-Language': 'en-US'}
movies_html = requests.get(url.format(), headers=headers).content
soup = BeautifulSoup(movies_html, 'html.parser')
soup_data = soup.find_all("tr")
all_movies = []
for data in soup_data[1:]:
data_array = data.text.split('\n')
try:
url = data.find_all('a')[0]
cast_data = get_cast(url)
# Saving summary data for V2
# summary_data = get_summary(url)
data_object = format_data(data_array)
data_object.update(cast_data)
all_movies.append(data_object)
except:
print('no report')
return all_movies
def get_summary(url):
url = 'https://www.the-numbers.com' + url.attrs['href']
headers = {'Accept-Language': 'en-US'}
movies_html = requests.get(url.format(), headers=headers).content
soup = BeautifulSoup(movies_html, 'html.parser')
soup_main = soup.find("div", {"id": "summary"})
return "coming soon"
def get_cast(url):
url = 'https://www.the-numbers.com' + url.attrs['href'].split("#")[0]+"#tab=cast-and-crew"
headers = {'Accept-Language': 'en-US'}
movies_html = requests.get(url.format(), headers=headers).content
soup = BeautifulSoup(movies_html, 'html.parser')
soup_main = soup.find("div", {"id": "cast-and-crew"})
soup_data = soup_main.find_all("div", class_="cast_new")
cast_data = {}
leads = []
supporting = []
production = []
for data in soup_data:
if 'Lead' in data.h1.text:
cast = data.find_all("tr")
for castmember in cast:
leads.append(castmember.text.strip().split('\n')[0])
if 'Supporting' in data.h1.text:
cast = data.find_all("tr")
for castmember in cast:
supporting.append(castmember.text.strip().split('\n')[0])
if 'Production' in data.h1.text:
cast = data.find_all("tr")
for castmember in cast:
production.append({castmember.text.strip().split('\n')[2]: castmember.text.strip().split('\n')[0]})
cast_data.update({'star': leads[0]})
cast_data.update({'leads': leads})
cast_data.update({'supporting': supporting})
cast_data.update({'production': production})
return cast_data
# +
def run_TN_reports_script():
all_pages = []
for year in range(2000,2020):
results = get_report(year)
all_pages += results
pd.DataFrame(all_pages).to_csv('TN_reports_data.csv')
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# # UNCOMMENT TO RUN <3
# run_TN_reports_script()
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# -
# ## THE CLEANING & PREP
# ##### Baby's First Lambda
# +
def clean_merged_file(merged_file):
big_movies = pd.read_csv(merged_file, encoding = "ISO-8859-1")
big_movies_clean = pd.DataFrame({
"id": big_movies['id'],
"imdb_id": big_movies['imdb_id'],
"name": big_movies['name'],
"budget": big_movies['budget'],
"revenue": big_movies['revenue'],
"runtime": big_movies['runtime'],
"score": big_movies['score'],
"vote_count": big_movies['vote_count'],
"released": big_movies['released'],
"tagline": big_movies['tagline'],
"production_companies": big_movies['production_companies'],
"genres": big_movies['genres']
})
return big_movies_clean
def get_all_from_list(list_of_things, num, key_to_get):
if list_of_things == '[]':
return 'na'
else:
try:
return eval(list_of_things)[num][key_to_get]
except:
return eval(list_of_things)[0][key_to_get]
# NOTE: This section only worked for some iterations of our data
# A clear sign we must refactor!
def widen_df(big_movies_clean):
# the slash at the end of the line is so we can split it into two lines
# PRODUCTION COMPANIES
big_movies_clean['production_company_1'] = big_movies.apply \
(lambda x: get_all_from_list(x['production_companies'], 0, 'name'),axis=1)
big_movies_clean['production_company_2'] = big_movies.apply \
(lambda x: get_all_from_list(x['production_companies'], 1, 'name'),axis=1)
big_movies_clean['production_company_3'] = big_movies.apply \
(lambda x: get_all_from_list(x['production_companies'], 2, 'name'),axis=1)
# # GENRES
big_movies_clean['genre_1'] = big_movies.apply \
(lambda x: get_all_from_list(x['genres'], 0, 'name'),axis=1)
big_movies_clean['genre_2'] = big_movies.apply \
(lambda x: get_all_from_list(x['genres'], 1, 'name'),axis=1)
big_movies_clean['genre_3'] = big_movies.apply \
(lambda x: get_all_from_list(x['genres'], 2, 'name'),axis=1)
# +
def run_clean_file_script():
big_movies_clean = clean_merged_file("testing_first_merge.csv")
big_movies_clean_v2 = widen_df(big_movies_clean)
big_movies_clean_v2.to_csv('big_movies_clean_v2.csv')
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# # UNCOMMENT TO RUN <3
# run_clean_file_script()
# # ++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++++
# -
# # =======================================================
# # PART 2: B - ANALYSIS
# # =======================================================
# ## THE LIBRARIES
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.patches as mpatchesphew
# ## THE INITIAL EDA
# +
# ======================================================================
# DOING EXPLORATORY DATA ANALYSIS ON A SMALL SUBSET (2018) OF THE DATA
# ======================================================================
# NOTE: The comments in this section are a nice melding of Ali practicality
# and Kendra color. Please do not fault Ali for the bad puns and word play
# -------------------------------------
# BUT FIRST, WE PREP!!
# -------------------------------------
# STEP 1: readin' and cleanin'
movies = pd.read_csv('2018_movies.csv')
movies.head()
movies.shape
# just say nah to na
# droping the first row of NaNs
movies = movies.drop([0,])
movies.shape
# dropping na and NaN in place
movies.dropna(inplace = True)
# STEP 2: droppin' and removin'
# say bye bye to those without budget
# (removing movies without the information we need)
index_names = movies[movies["budget"] == 0].index
# We can see that 325 movies in our df have a budget of 0 dollars...
# We have to drop these movies
index_names
# Repeating above... but with revenue
movies.drop(index_names, inplace = True)
index_names = movies[movies["revenue"] == 0].index
movies.drop(index_names, inplace = True)
# STEP 3: formattin' and finessin'
# Type-casting isn't just for Hollywood
# Checking to see the data type for the release_date column
movies.release_date.dtype
#It shows that it is saved as an object, we want to convert this to date format
#Changing the data type to date by using the pd_to_datetime function, this will allow us to extract each element of the date
movies["release_date"] = pd.to_datetime(movies["release_date"])
#now we want to extract the month, day, and year and create new columns named month, day and year
movies["month"], movies["day"], movies["year"] = movies["release_date"].dt.month, movies["release_date"].dt.day, movies["release_date"].dt.year
# STEP 4: surmisin' and summarisin'
# Getting summary statistics for our df
movies.describe()
# -
# It's interesting to note that the lowest budget for a movie in our df is only 258,157 and the maximum budget is 300,000,000. The lowest revenue is 4,537. The maximum revenue is 2,046,240,000. This is a massive revenue. We aggregated a profit column and the minimum profit is - 60,477,350 and maximum profit is 1,746,240,000. This is a massive profit. Their is a large range in popularity scores. The maximum popularity score is 71.54 and the minimum is 3.54. The mean popularity is 18.4 and 75% of the movies have a popularity score less than 21.2. This makes us question if the maximum popularity score might be an error, or it might correspond to the movie with the highest profit. We should investigate what movie this score references. The vote_average column has a range of 4 - 8.4, with an average of 6.5. The vote_count has a maximum of 14,913. This might reference the same movie that had the largest popularity. The max value in this column, also appears to be an outlier, as 75% of the movies have less than 2151 votes and the average vote count is 1701
# STEP 5: column creatin'
#We have decided that we want to aggregate a percent profit column as well, in an attempt to normalize the data
#To do this we are diving the profit column by the budget column and multiply the result by 100 and saving it in a
#new column named percent_profit
movies["percent_profit"] = movies["profit"]/movies["budget"]*100
# Saving a column as-is for future use
movies_original_df = movies
movies.budget.describe()
# First, we are going to discretize the budget column.
# We are discretizing the budget column into four groups: extremely_low, low, high and extremely_high. To do this we first
# need to create a list of the categories
#
categories = ["extremely_low", "low", "high", "extremely_high"]
# Now we need to show where we want to insert the breaks. We have decided that extremely low budgets are budgets less
# than 13,000,000, low have budgets between 13,000,000 and 30,000,000, high have budgets between 30,000,000 and
# 62,192,550, and extremely_high have budgets between 62,192,550 and 300,000,000. We chose the values based on the
# quartiles.
#Saving the movies df as movies_discretized_df
movies_discretized_df = movies
#Discretizing the budget columns using the cut function from pandas
movies_discretized_df["budget"] = pd.cut(movies_discretized_df["budget"], [0, 13000000, 30000000, 62192550, 300000001], labels = categories)
#Now we are going to repeat the steps to discretize the revenue column
movies_discretized_df.revenue.describe()
#We are using the same categories as above
# `extremely_low` revenue are revenues less than 21,458,200, `low` are revenues between 21,458,200 and
# 62,954,020, `high` revenues are revenues between 62,954,020 and 187,976,900, and `extremely_high` revenues between
# 187,976,900 and 2,046,240,000.
#
movies_discretized_df["revenue"] = pd.cut(movies_discretized_df["revenue"], [0, 21458200, 62954020, 187976900, 2046240001], labels = categories)
#Now we are going to repeat the steps to discretized the profit column
movies_discretized_df.profit.describe()
#Now we are going to repeat the steps to discretized the profit column
movies_discretized_df.profit.describe()
'''negative profit are profits less than $0, low profits are profits between $0 and
$29,314,900, high profits are profits between $29,314,900 and $140,784,100, and extremely_high profits between
$140,784,100 and $1,746,240,001.
'''
categories = ["negative", "low", "high", "extremely_high"]
movies_discretized_df["profit"] = pd.cut(movies_discretized_df["profit"], [-60477351, 0, 29314900, 140784100, 1746240001], labels = categories)
movies_discretized_df.profit.dtype
#Now we are going to repeat the steps to discretize the popularity column
movies_discretized_df.popularity.describe()
categories = ["extremely_low", "low", "high", "extremely_high"]
'''extremely_low popularity are popularities less than 12.442, low popularities are popularities between 12.442 and
15.7405, high popularity are popularities between 15.7405 and 21.23025 and extremely_high popularity between 21.23025
and 71.538'''
movies_discretized_df["popularity"] = pd.cut(movies_discretized_df["popularity"], [0, 12.442, 15.7405, 21.23025, 71.538], labels = categories)
movies_discretized_df["popularity"].head()
#Now we are going to repeat the steps to discretize the vote avg
movies_discretized_df.vote_average.describe()
#We are using the same categories as above
'''extremely_low vote_average are vote averages less than 6, low are between 6 to 6.5, high between 6.5 and 7 and
extremely_high 7 and 8.5'''
movies_discretized_df["vote_average"] = pd.cut(movies_discretized_df["vote_average"], [0, 6, 6.5, 7, 8.5], labels = categories)
movies_discretized_df["vote_average"].head()
#We are using the same categories as above
'''extremely_low vote counts are vote counts less than 440, low are between 440 and 1151, high are between 1151 and 2522
and extremely_high are between 2522 and 14913'''
movies_discretized_df["vote_count"] = pd.cut(movies_discretized_df["vote_count"], [0, 440, 1151, 2522, 14914], labels = categories)
movies_discretized_df["vote_count"].head()
movies_discretized_df.percent_profit.describe()
'''extremely_low are percent profits between -100 and 0, low between 6.5 and 108, high between 108 and 436 and
extremely_high between 436 and 6527'''
categories = ["negative", "low", "high", "extremely_high"]
movies_discretized_df["percent_profit"] = pd.cut(movies_discretized_df["percent_profit"], [-100, 0, 108, 436, 6528], labels = categories )
movies_discretized_df["percent_profit"]
movies_discretized_df.day.describe()
# +
#We are setting new categories for the day column by creating a new column for week
'''week_1 is the first 7 days of the month, week_2 is days 8 - 14, week_3 is days 15 - 21, and week_4 are the
rest of the days'''
categories = ["week_1", "week_2", "week_3", "week_4"]
movies_discretized_df["week"] = pd.cut(movies_discretized_df["day"], [0, 8, 15, 22, 32], labels = categories)
movies_discretized_df.head()
#We have successfully discretized the df, now we can remove the day and release_date column
movies_discretized_df.drop(columns=['day', 'release_date'], inplace = True)
#Checking to make sure that it worked
movies_discretized_df.head()
# +
#Question 1:
#How are the amounts of percent_profits distributed across budget levels?
'''We want to compare the budget category percentage make up for each percent_profit level. To do this we need to
get the count for each budget level, the count for each percent_profit level by budget level and then divide
the count of the percent_profit/count of budget level and multiply by 100. We have to do this for each
budget level and level of percent_profits. We think that we could potentially answer this question by group bys.'''
movies_discretized_count = movies_discretized_df.groupby(["budget", "percent_profit"])["budget"].count()
'''Taking the output from the line above and converting it to a data frame. We are using pandas, which we important as pd.
First, we call the package we are using then the function from that package and then what we want to run the function on.
pd.function(item to use). We are using the DataFrame function from the pandas package on the series created by our group by'''
movies_discretized_count_df = pd.DataFrame(movies_discretized_count)
#Checking to see what our df looks like.
movies_discretized_count_df
#Changing the column name from budget to counts
movies_discretized_count_df.columns = ["counts"]
#Checking to see what our df looks like.
movies_discretized_count_df
# -
#We want to get a total count for the number of percent_profit counts for each budget level. We will experiment to see how this is possible
'''This shows that we have 2 indexes budget and percent_profit... We want to create columns from each index
We are creating a new column named budget by extracting the values from the first index (0) which is the budget
index'''
movies_discretized_count_df["budget_category"]=movies_discretized_count_df.index.get_level_values(0)
#We are creating a new column named total_donations by extracting the values from the second index (1) which is total_donations
movies_discretized_count_df["percent_profit_category"] = movies_discretized_count_df.index.get_level_values(1)
#Checking to make sure it worked...
movies_discretized_count_df
#It did!
#Now we want to remove the indexes so, we can create a new group by to get the sum of the counts for each group
#To do this we are using the reset_index(drop = True) This will drop our group by indexes and allow us to create a new one.
movies_discretized_count_df = movies_discretized_count_df.reset_index(drop = True)
movies_discretized_count_df
#Now we are getting the sum of each budget category.
budget_discretized_count_df = movies_discretized_count_df.groupby(["budget_category"])["counts"].sum()
#Checking the results
budget_discretized_count_df
'''We ultimately want a column that contains the total counts for each budget group. We are probably doing this in
a roundabout way, but as I am extremely new to python this is the best way I can think of doing it. We are going to create
a new column that replicates the income_level called income_level_count and then we will use the replace function to
replace the 1s with their total count, the 2s with their total count... '''
#First, replicating the income level column in a column named budget_category_count
movies_discretized_count_df["budget_category_count"] = movies_discretized_count_df["budget_category"]
#Now replacing the income level with the total count for each income level
movies_discretized_count_df["budget_category_count"] = movies_discretized_count_df["budget_category_count"].replace(["extremely_low"], 35)
movies_discretized_count_df["budget_category_count"] = movies_discretized_count_df["budget_category_count"].replace(["low"], 35)
movies_discretized_count_df["budget_category_count"] = movies_discretized_count_df["budget_category_count"].replace(["high"], 32)
movies_discretized_count_df["budget_category_count"] = movies_discretized_count_df["budget_category_count"].replace(["extremely_high"], 34)
#Checking to see if that worked:
movies_discretized_count_df
#Okay, we are one step closer... Now, we need to create a column that takes the counts/budget_category_counts * 100
movies_discretized_count_df["percent"] = movies_discretized_count_df["counts"]/movies_discretized_count_df["budget_category_count"] *100
#Looking at our data frame... It worked!!!
movies_discretized_count_df
#We no longer need the count columns
movies_discretized_count_df.drop(["counts", "budget_category_count"], axis = 1, inplace = True )
'''Attempting to graph this data using a grouped bar chart:
formula: df.pivot(columns, group, values).plot(kind = "type of graph", color = ["color to use, can be a list of colors"],
title = "you can set the title of your graph here")'''
graph = movies_discretized_count_df.pivot("budget_category", "percent_profit_category",
"percent").plot(kind="bar", color = ["crimson", "salmon", "palegreen", "darkgreen"],
title = "Percent of Percent Profit to Budget Category")
#Changing the y label of our graph to Percent
plt.ylabel("Percent")
#Changing the x axis label of our graph to Budget Category
plt.xlabel("Budget Category")
#Making it so the tick labels are not angled
plt.xticks(rotation = 0)
#How to change the tick labels (we ended up not needing this, but want to keep for future reference)
#plt.Axes.set_xticklabels(graph, labels = ['extremely low', 'low', 'high', 'extremely high'])
#moving the legend position to underneath the graph, also setting it to have 4 columns so the legend is in a
#straight single line and adding a legend title
plt.legend( loc = "lower center", bbox_to_anchor = (.5, -.4), ncol = 4, title = "Percent Profit Category")
# This graph proved very interesting. Movies with an extremely low budget have the highest percentage make-up of making an extremely
# high percent profit. Movies with an extremely high budget are the most likely to be profitable overall, being that they
# are the least likely to have a negative profit, with only 5.9% of the movies classified as having an extremely high
# budget in our dataset made a negative profit. Movies with an low or high budget only make an extremely high
# percent profit less than 17.1% and 15.6% of the time respectively. They also have the highest chance of making a low or
# negative profit out of all of the budget categories. Based, on this analysis, percent profits are not uniformally
# distributed across budget levels. Movies with an extremely high budget are the least likely to have a negative percent
# profit. Movies with an extremely low budget are the most likely to have an extremely high percent profit. Our
# recommendation to studios, would be to either have a extremely low or extremely high budget and to veer away from
# productions with an extremely low or high budget. Further analysis for tighter recommendatios is needed.
#
# +
#Question: Do big name production companies impact the percent profit?
#To answer this question we are first going to create a for loop that will loop through the production_companies column
#in order movies_discretized_df and store the production company in a list called production_company. The only issue
#with this method, is that if a movie has more than one production company this will not be shown and only the last
#company included in the for loop will be given credit.
production_company = []
for movie in movies_discretized_df['production_companies']:
if "Universal" in movie:
production_company.append("Universal")
elif "Sony" in movie:
production_company.append("Sony")
elif "Fox" in movie:
production_company.append("Fox")
elif "DreamWorks" in movie:
production_company.append("DW")
elif "MGM" in movie:
production_company.append("MGM")
elif "Paramount" in movie:
production_company.append("Paramount")
elif "Disney" in movie:
production_company.append("Disney")
elif "<NAME>" in movie:
production_company.append("WB")
else:
production_company.append("None")
# -
movies_discretized_df["main_production_co"] = production_company
movies_discretized_df.head()
#Now we want to group by production company and percent profit
'''We want to compare the production company percentage make up for each percent_profit level. To do this we need to
get the count for each production company, the count for each percent_profit level by production company and then divide
the count of the percent_profit/count of production company and multiply by 100. We have to do this for each
production company and level of percent_profits. We think that we could potentially answer this question by group bys.'''
movies_discretized_count_q2 = movies_discretized_df.groupby(["main_production_co", "percent_profit"])["main_production_co"].count()
'''Taking the output from the line above and converting it to a data frame. We are using pandas, which we important as pd.
First, we call the package we are using then the function from that package and then what we want to run the function on.
pd.function(item to use). We are using the DataFrame function from the pandas package on the series created by our group by'''
movies_discretized_count_df_q2 = pd.DataFrame(movies_discretized_count_q2)
#Checking to see what our df looks like.
movies_discretized_count_df_q2
#Changing the column name from budget to counts
movies_discretized_count_df_q2.columns = ["counts"]
#Checking to see what our df looks like.
movies_discretized_count_df_q2.head()
#We want to get a total count for the number of percent_profit counts for each production company.
'''This shows that we have 2 indexes budget and percent_profit... We want to create columns from each index
We are creating a new column named budget by extracting the values from the first index (0) which is the budget
index'''
movies_discretized_count_df_q2["production_company"]=movies_discretized_count_df_q2.index.get_level_values(0)
#We are creating a new column named total_donations by extracting the values from the second index (1) which is total_donations
movies_discretized_count_df_q2["percent_profit_category"] = movies_discretized_count_df_q2.index.get_level_values(1)
#Checking to make sure it worked...
movies_discretized_count_df_q2
#It did!
#Now we want to remove the indexes so, we can create a new group by to get the sum of the counts for each group
#To do this we are using the reset_index(drop = True) This will drop our group by indexes and allow us to create a new one.
movies_discretized_count_df_q2 = movies_discretized_count_df_q2.reset_index(drop = True)
#Now we are getting the sum of each production company category.
production_company_discretized_count_df_q2 = movies_discretized_count_df_q2.groupby(["production_company"])["counts"].sum()
#Checking the results
production_company_discretized_count_df_q2
'''We ultimately want a column that contains the total counts for each production company. We are going to create
a new column that replicates the production company called production_company_count and then we will use the replace function to
replace the 1s with their total count, the 2s with their total count... '''
#First, replicating the income level column in a column named budget_category_count
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company"]
#Now replacing the income level with the total count for each income level
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["DW"], 1)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["Disney"], 8)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["Fox"], 11)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["None"], 66)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["Paramount"], 9)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["Sony"], 8)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["Universal"], 20)
movies_discretized_count_df_q2["production_company_count"] = movies_discretized_count_df_q2["production_company_count"].replace(["WB"], 13)
movies_discretized_count_df_q2
#Okay, we are one step closer... Now, we need to create a column that takes the counts/budget_category_counts * 100
movies_discretized_count_df_q2["percent"] = movies_discretized_count_df_q2["counts"]/movies_discretized_count_df_q2["production_company_count"] *100
#Looking at our data frame... It worked!!!
movies_discretized_count_df_q2.head()
#We no longer need the count columns
movies_discretized_count_df_q2.drop(["counts", "production_company_count"], axis = 1, inplace = True )
'''Attempting to graph this data using a grouped bar chart:
formula: df.pivot(columns, group, values).plot(kind = "type of graph", color = ["color to use, can be a list of colors"],
title = "you can set the title of your graph here")'''
graph = movies_discretized_count_df_q2.pivot("production_company", "percent_profit_category",
"percent").plot(kind="bar", color = ["crimson", "salmon", "palegreen", "darkgreen"],
title = "Percent of Percent Profit to Production Company")
#Changing the y label of our graph to Percent
plt.ylabel("Percent")
#Changing the x axis label of our graph to Budget Category
plt.xlabel("Production Company")
#Making it so the tick labels are not angled
plt.xticks(rotation = 0)
#How to change the tick labels (we ended up not needing this, but want to keep for future reference)
#plt.Axes.set_xticklabels(graph, labels = ['extremely low', 'low', 'high', 'extremely high'])
#moving the legend position to underneath the graph, also setting it to have 4 columns so the legend is in a
#straight single line and adding a legend title
plt.legend( loc = "lower center", bbox_to_anchor = (.5, -.4), ncol = 4, title = "Percent Profit Category")
# This graph provides some insights, however, most of our movies have more than one main production company and only
# one production company is being shown. For example, DreamWorks and Universal had a movie named <NAME> and it was
# profitable. However, based on the way that we assigned a main production company, only Universal was given credit for
# that movie.
# Question:
#Does time of the month the movie is released affect percent profit?
'''We want to compare the percent_profit level percentage make up for each time of month. To do this we need to
get the count for each time of month, the count for each percent_profit level by time of month company and then divide
the count of the percent_profit/count of time of month and multiply by 100. We have to do this for each
time of month and level of percent_profits.'''
movies_discretized_count_q3 = movies_discretized_df.groupby(["week", "percent_profit"])["week"].count()
'''Taking the output from the line above and converting it to a data frame. We are using pandas, which we important as pd.
First, we call the package we are using then the function from that package and then what we want to run the function on.
pd.function(item to use). We are using the DataFrame function from the pandas package on the series created by our group by'''
movies_discretized_count_df_q3 = pd.DataFrame(movies_discretized_count_q3)
#Checking to see what our df looks like.
movies_discretized_count_df_q3
#Changing the column name from week to counts
movies_discretized_count_df_q3.columns = ["counts"]
#Checking to see what our df looks like.
movies_discretized_count_df_q3.head()
#We want to get a total count for the number of percent_profit counts for each week.
'''This shows that we have 2 indexes week and percent_profit... We want to create columns from each index
We are creating a new column named week by extracting the values from the first index (0) which is the week
index'''
movies_discretized_count_df_q3["week"]=movies_discretized_count_df_q3.index.get_level_values(0)
#We are creating a new column named total_donations by extracting the values from the second index (1) which is percent_profit
movies_discretized_count_df_q3["percent_profit_category"] = movies_discretized_count_df_q3.index.get_level_values(1)
#Checking to make sure it worked...
movies_discretized_count_df_q3
#It did!
#Now we want to remove the indexes so, we can create a new group by to get the sum of the counts for each group
#To do this we are using the reset_index(drop = True) This will drop our group by indexes and allow us to create a new one.
movies_discretized_count_df_q3 = movies_discretized_count_df_q3.reset_index(drop = True)
# Now we are getting the sum of each production company category.
week_discretized_count_df_q3 = movies_discretized_count_df_q3.groupby(["week"])["counts"].sum()
#Checking the results
week_discretized_count_df_q3
'''We ultimately want a column that contains the total counts for each week. We are going to create
a new column that replicates the week called week_count and then we will use the replace function to
replace the 1s with their total count, the 2s with their total count... '''
#First, replicating the income level column in a column named budget_category_count
movies_discretized_count_df_q3["week_count"] = movies_discretized_count_df_q3["week"]
#Now replacing the income level with the total count for each income level
movies_discretized_count_df_q3["week_count"] = movies_discretized_count_df_q3["week_count"].replace(["week_1"], 37)
movies_discretized_count_df_q3["week_count"] = movies_discretized_count_df_q3["week_count"].replace(["week_2"], 34)
movies_discretized_count_df_q3["week_count"] = movies_discretized_count_df_q3["week_count"].replace(["week_3"], 33)
movies_discretized_count_df_q3["week_count"] = movies_discretized_count_df_q3["week_count"].replace(["week_4"], 32)
movies_discretized_count_df_q3.head()
#Okay, we are one step closer... Now, we need to create a column that takes the counts/week_count * 100
movies_discretized_count_df_q3["percent"] = movies_discretized_count_df_q3["counts"]/movies_discretized_count_df_q3["week_count"] *100
#Looking at our data frame... It worked!!!
movies_discretized_count_df_q3.head()
#We no longer need the count columns
movies_discretized_count_df_q3.drop(["counts", "week_count"], axis = 1, inplace = True )
'''Attempting to graph this data using a grouped bar chart:
formula: df.pivot(columns, group, values).plot(kind = "type of graph", color = ["color to use, can be a list of colors"],
title = "you can set the title of your graph here")'''
graph = movies_discretized_count_df_q3.pivot("week", "percent_profit_category",
"percent").plot(kind="bar", color = ["crimson", "salmon", "palegreen", "darkgreen"],
title = "Percent of Percent Profit to Week")
#Changing the y label of our graph to Percent
plt.ylabel("Percent")
#Changing the x axis label of our graph to Budget Category
plt.xlabel("Week")
#Making it so the tick labels are not angled
plt.xticks(rotation = 0)
#How to change the tick labels (we ended up not needing this, but want to keep for future reference)
#plt.Axes.set_xticklabels(graph, labels = ['extremely low', 'low', 'high', 'extremely high'])
#moving the legend position to underneath the graph, also setting it to have 4 columns so the legend is in a
#straight single line and adding a legend title
plt.legend( loc = "lower center", bbox_to_anchor = (.5, -.4), ncol = 4, title = "Percent Profit Category")
# This is interesting in that it shows that movies released within the first two weeks of the month tend to be more
# profitable. We would like to look at a breakdown of month to percent profit for further analysis
# Taking a brief detour back to our non-discretized df
movies_original_df.head()
# QUESTION: Do "Good" Movies Make Money? -- We're defining "Good" as vote average
plt.plot(movies_original_df.profit, movies_original_df.vote_average, 'o')
plt.title('Do "Good" Movies Make Money?')
plt.xlabel('Profit')
plt.ylabel('Vote Average')
plt.show()
# QUESTION: Does Popularity = Profit?
plt.plot(movies_original_df.profit, movies_original_df.popularity, 'o')
plt.title('Does Popularity = Profits?')
plt.xlabel('Profit')
plt.ylabel('Popularity')
plt.show()
# QUESTION: How does budget impact vote average?
plt.plot(movies_original_df.budget, movies_original_df.vote_average, 'o')
plt.title('How does Budget Impact Vote Average?')
plt.xlabel('Budget')
plt.ylabel('Vote Average')
plt.show()
# QUESTION: How does budget impact popularity?
plt.plot(movies_original_df.budget, movies_original_df.popularity, 'o')
plt.title('How does Budget Impact Popularity?')
plt.xlabel('Budget')
plt.ylabel('Popularity')
plt.show()
# +
# QUESTION: Is there a relationship between "Above Average Movies" and Budget/Price?
below_avg = movies_original_df[movies_original_df.vote_average < 6.5]
above_avg = movies_original_df[movies_original_df.vote_average >= 6.5]
plt.plot(below_avg.budget, below_avg.profit, 'o', label="below average")
plt.plot(above_avg.budget, above_avg.profit, 'o', label="above average")
plt.title('BUDGET vs PROFIT by AVERAGE VOTE!')
plt.xlabel('BUDGET')
plt.ylabel('PROFIT')
plt.legend()
plt.show()
# +
# QUESTION: Is there a relationship between "Above Average Movies" and Budget/Price?
below_avg = movies_original_df[movies_original_df.vote_average < 6.5]
above_avg = movies_original_df[movies_original_df.vote_average >= 6.5]
plt.plot(below_avg.budget, below_avg.percent_profit, 'o', label="below average")
plt.plot(above_avg.budget, above_avg.percent_profit, 'o', label="above average")
plt.title('BUDGET vs PERCENT PROFIT by AVERAGE VOTE!')
plt.xlabel('BUDGET')
plt.ylabel('PROFIT')
plt.legend()
plt.show()
# +
# BIG QUESTION: What role do production companies play in the entertainment industry?
# Is there a relationship between production studio and average vote?
# Production studio and budget?
# Production studio and percent profit?
# +
# Adding the BIG EIGHT Production Studios to the DF
# WARNER BROS
wb = []
for movie in movies_original_df['production_companies']:
if "Warner Bros" in movie:
wb.append(True)
else:
wb.append(False)
movies_original_df['wb'] = wb
# MGM
mgm = []
for movie in movies_original_df['production_companies']:
if "MGM" in movie:
mgm.append(True)
else:
mgm.append(False)
movies_original_df['mgm'] = mgm
# DREAMWORKS
dw = []
for movie in movies_original_df['production_companies']:
if "DreamWorks" in movie:
dw.append(True)
else:
dw.append(False)
movies_original_df['dw'] = dw
# SONY
sony = []
for movie in movies_original_df['production_companies']:
if "Sony" in movie:
sony.append(True)
else:
sony.append(False)
movies_original_df['sony'] = sony
# DISNEY
disney = []
for movie in movies_original_df['production_companies']:
if "Disney" in movie:
disney.append(True)
else:
disney.append(False)
movies_original_df['disney'] = disney
#FOX
fox = []
for movie in movies_original_df['production_companies']:
if "Century Fox" in movie:
fox.append(True)
else:
fox.append(False)
movies_original_df['fox'] = fox
# PARAMOUNT
paramount = []
for movie in movies_original_df['production_companies']:
if "Paramount" in movie:
paramount.append(True)
else:
paramount.append(False)
movies_original_df['paramount'] = paramount
#UNIVERSAL
universal = []
for movie in movies_original_df['production_companies']:
if "Universal" in movie:
universal.append(True)
else:
universal.append(False)
movies_original_df['universal'] = universal
# +
sony = movies_original_df[movies_original_df.sony == True]
wb = movies_original_df[movies_original_df.wb == True]
disney = movies_original_df[movies_original_df.disney == True]
fox = movies_original_df[movies_original_df.fox == True]
universal = movies_original_df[movies_original_df.universal == True]
paramount = movies_original_df[movies_original_df.paramount == True]
dw = movies_original_df[movies_original_df.dw == True]
mgm = movies_original_df[movies_original_df.mgm == True]
plt.plot(sony.budget, sony.revenue, 'o', label="Sony")
plt.plot(wb.budget, wb.revenue, 'o', label="Warner Bros.")
plt.plot(disney.budget, disney.revenue, 'o', label="Disney")
plt.plot(fox.budget, fox.revenue, 'o', label="Fox")
plt.plot(universal.budget, universal.revenue, 'o', label="Universal")
plt.plot(paramount.budget, paramount.revenue, 'o', label="Paramount")
plt.plot(dw.budget, dw.revenue, 'o', label="DreamWorks")
plt.plot(mgm.budget, mgm.revenue, 'o', label="MGM")
plt.title('BUDGET vs REVENUE by PRODUCTION COMPANY')
plt.xlabel('BUDGET')
plt.ylabel('REVENUE')
plt.legend( loc = "lower center", bbox_to_anchor = (.5, -.4), ncol = 4, title = "Production Company")
plt.show()
# +
sony = movies_original_df[movies_original_df.sony == True]
wb = movies_original_df[movies_original_df.wb == True]
disney = movies_original_df[movies_original_df.disney == True]
fox = movies_original_df[movies_original_df.fox == True]
universal = movies_original_df[movies_original_df.universal == True]
paramount = movies_original_df[movies_original_df.paramount == True]
dw = movies_original_df[movies_original_df.dw == True]
mgm = movies_original_df[movies_original_df.mgm == True]
plt.plot(sony.budget, sony.percent_profit, 'o', label="Sony")
plt.plot(wb.budget, wb.percent_profit, 'o', label="<NAME>.")
plt.plot(disney.budget, disney.percent_profit, 'o', label="Disney")
plt.plot(fox.budget, fox.percent_profit, 'o', label="Fox")
plt.plot(universal.budget, universal.percent_profit, 'o', label="Universal")
plt.plot(paramount.budget, paramount.percent_profit, 'o', label="Paramount")
plt.plot(dw.budget, dw.percent_profit, 'o', label="DreamWorks")
plt.plot(mgm.budget, mgm.percent_profit, 'o', label="MGM")
plt.title('BUDGET vs PERCENT PROFIT by PRODUCTION COMPANY')
plt.xlabel('BUDGET')
plt.ylabel('PERCENT PROFIT')
plt.legend( loc = "lower center", bbox_to_anchor = (.5, -.4), ncol = 4, title = "Production Company")
plt.show()
# -
|
assets/all_html/2019_09_04_Final_Project_Ali_Ho_Kendra_Osburn_P2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def searchInsert(nums, target):
if target in nums:
return nums.index(target)
if target < nums[0]:
return 0
for i in range(len(nums) - 1):
if nums[i] < target and nums[i + 1] > target:
return i + 1
return len(nums)
print(searchInsert([1,1,3,5,6], 1))
# -
|
Anjani/Leetcode/Array/Search Insert Position.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import io
from typing import Dict
from pathlib import Path
from pprint import pprint
from IPython.display import clear_output, display, display_markdown
from ipywidgets import FileUpload, Layout, Output
from fastai.vision import load_learner, Image, open_image
# -
layout = Layout(width="100%", min_height="180px", align="center")
v = FileUpload(multiple=False, accept="image/*", layout=layout)
o = Output()
# +
def my_transform(x):
x.data = 1 - x.data
return x
learner_path = Path("~/.fastai/data/mnist_png").expanduser()
l = load_learner(learner_path)
# -
def predict(i: Image) -> Dict:
cat, _, probs = l.predict(i)
return {
'category': cat.obj,
'confidence': probs.numpy().round(4)[_]
}
def _handle_upload(change):
[upload] = change["new"].values()
stream = io.BytesIO(upload["content"])
image = open_image(stream).apply_tfms(None, size=128)
p = predict(image)
with o:
clear_output()
display_markdown(f"# {str(p)}", raw=True)
display(image)
v.observe(_handle_upload, "value")
display(v)
display(o)
|
fastai-learner-on-mobile/widgit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h2>Machine Learning Model for Predicting Cars' Fuel Consumption</h2>
# <h3><b>Abstract:</b></h3><br>
# This Jupiter Notebook project represents a Machine Learning Multiple Linear Regression model that was developed by using the Python programming language. The developed model enables to predict the cars' fuel consumption in function of factors such as age of cars, cars' weight, number of cylinders in vehicles' engines, horsepower and others. A well-known fuel consumption <a href='https://archive.ics.uci.edu/ml/datasets/Auto+MPG'>dataset</a> was used for the purposes of the project. The dataset was subsequently split in two datasets - train and test datasets. The train dataset was used to train the regression model and the test dataset was used to predict the fuel consumption with the fitted model. The value of the coefficient of determination R-squared = 0.92 shows that about 92% of the variation in the values of the dependent variable 'fuel consumption' can be explained with the influence of the independent factors in the model and the rest 8% depend on other factors which the model does not take into account.
# <p><b>Author: </b><NAME>, PhD</p>
# <b>Last Updated: </b>19 August 2020</p>
# <h2>Table of Contents</h2>
#
# <a>1. Libraries and Modules</a><br>
# <a>2. Data Import</a><br>
# <a>3. Data Wrangling and Cleansing</a><br>
# <a>4. Exploratory Data Analysis and Data Visualization</a><br>
# <a>5. Model Development, Training and Prediction</a><br>
# <a style="padding-left: 15px;">5.1. Model Training</a><br>
# <a style="padding-left: 15px;">5.2. Model Predictions and Evaluation Metrics</a><br>
# <a>6. References</a><br>
# <a>About the Author</a>
#
# <h2 id='1. Libraries and Modules'>1. Libraries and Modules</h2>
# Import libraries and modules
import pandas as pd
import datetime
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
# <h2 id='2. Data Import'>2. Data Import</h2>
# +
# Declare column names
col_names = ['mpg', 'cylinders', 'displacement', 'horsepower', 'weight', 'acceleration', 'model_year', 'origin', 'car_name']
# Assign the content of the '.data' file to a new dataframe called df
df = pd.read_fwf('auto-mpg.data', names = col_names)
# Show the first 5 data rows
df.head()
# -
df.tail()
# Show the number of datarows and columns
df.shape
# Display the data types
df.dtypes
# <h2 id='3. Data Wrangling and Cleansing'>3. Data Wrangling and Cleansing</h2>
# Calculate and show a descriptive statistics
df.describe()
# <b>Note:</b> Statistics for the 'horsepower' column is not showing as it is an object (string) value column!
# Check for missing values
df.isnull().sum()
# Replace the missing values in the 'horsepower' column
df['horsepower'] = df['horsepower'].replace('?', '0')
# +
# Cast the 'horsepower' column from an object (string) to a numeric (float64)
df['horsepower'] = pd.to_numeric(df['horsepower'])
# Display the data types
df.dtypes
# +
# Convert the fuel consumption measurement unit from US miles per galon(mpg) to litres per 100km(L/100km)
fuel_conversion_rate = 235.2146
df['mpg'] = round(fuel_conversion_rate/df['mpg'],1)
# Rename the 'mpg' column to 'fuel consumption'
df.rename(columns = {'mpg':'fuel_consumption'}, inplace = True)
# Show the first 5 data rows
df.head()
# -
# Count and display in a table the distribution of the cars names by a unique name
df_car_names = df['car_name'].value_counts().to_frame()
df_car_names
# Assign to the df_cylinders dataframe and show the distribution of the cars grouped by an engine type (number of cylinders)
df_cylinders = df['cylinders'].value_counts().to_frame()
df_cylinders
# Create and display in a table a new dataframe(df_horsepower) from existing one(df) having 2 columns(cylinders and horsepower)
df_horsepower = df[['cylinders', 'horsepower']].copy()
df_horsepower
# Calculate and show the average horsepower of cars by the number of cylinders (3, 4, 5, 6 and 8).
df_av_hp = df_horsepower.groupby(['cylinders']).mean()
df_av_hp
# +
# Assign to a separate variable the calculated average HP value by engine type (number of cylinders)
hp_av_3 = round(df_av_hp.iloc[0,0])
hp_av_4 = round(df_av_hp.iloc[1,0])
hp_av_5 = round(df_av_hp.iloc[2,0])
hp_av_6 = round(df_av_hp.iloc[3,0])
hp_av_8 = round(df_av_hp.iloc[4,0])
# Print the averaged HP values
print('-----------------------------------------------------------------')
print('average horse power for 3-cylinder engines =', hp_av_3)
print('average horse power for 4-cylinder engines =', hp_av_4)
print('average horse power for 5-cylinder engines =', hp_av_5)
print('average horse power for 6-cylinder engines =', hp_av_6)
print('average horse power for 8-cylinder engines =', hp_av_8)
print('-----------------------------------------------------------------')
# Assign to a variable and print the number of the datarows in the df dataframe
counter = df.shape[0]
#print('counter = ', counter)
# Check for 0 values by engine type in the horsepower column and assign the calculated average values
for i in range(counter):
if df.loc[i, 'cylinders'] == 3 and df.loc[i, 'horsepower'] == 0:
df.loc[i, 'horsepower'] = hp_av_3
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
elif df.loc[i, 'cylinders'] == 4 and df.loc[i, 'horsepower'] == 0:
df.loc[i, 'horsepower'] = hp_av_4
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
elif df.loc[i, 'cylinders'] == 5 and df.loc[i, 'horsepower'] == 0:
df.loc[i, 'horsepower'] = hp_av_5
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
elif df.loc[i, 'cylinders'] == 6 and df.loc[i, 'horsepower'] == 0:
df.loc[i, 'horsepower'] = hp_av_6
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
elif df.loc[i, 'cylinders'] == 8 and df.loc[i, 'horsepower'] == 0:
df.loc[i, 'horsepower'] = hp_av_8
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
else:
pass
#print('Row number =', i, 'cylinders =', df.loc[i, 'cylinders'], df.loc[i, 'horsepower'])
# +
# Declare variables for the full current date as well as for the current year only
current_full_date = datetime.datetime.now()
current_year = current_full_date.year
# Print the current year
#print(current_year)
# Represent the year from a 2-digit number to a 4-digit number (e.g. from 70 to 1970) to calculate the age of the cars.
# The below formula is only valid for cars produced in 20th century (see the original dataset)!
df['model_year'] = df['model_year'] + 1900
# Rename the 'model_year' column to a 'car_age' column and calculate the cars' age.
df.rename(columns = {'model_year':'car_age'}, inplace = True)
df['car_age'] = current_year - df['car_age']
# Display the first 5 datarows including the calculated car age
df.head()
# -
# Convert the cars' weight from pounds[lb] to kilograms[kg]
weight_conversion_rate = 0.4536
df['weight'] = round(df['weight']*weight_conversion_rate)
# +
# Drop the 'origin' and 'car_name' columns from the df dataframe
df.drop(columns = ['origin', 'car_name'], axis = 1, inplace = True)
# Display the frist 5 data rows
df.head()
# -
# Save to a '.csv' file the working df dataframe
df.to_csv('df_hp_missing_values_fixed.csv', sep = ',')
# <h2 id='4. Exploratory Data Analysis and Data Visualization'>4. Exploratory Data Analysis and Data Visualization</h2>
# Calculate and print descriptive statistics to see the 'horsepower', 'weight' and 'car_age' values.
df.describe()
# +
# Create a list variable containing the columns name excluding the removed "origin" and "car_name" columns
col_names_corr = ['fuel_consumption', 'cylinders', 'displacement', 'horsepower', 'weight', 'acceleration', 'car_age']
# Dsiplay scatter plot diagram for the feature variable 'cylinders' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'cylinders', y = 'fuel_consumption', data = df)
# -
# Display scatter plot diagram for the feature variable 'displacement' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'displacement', y = 'fuel_consumption', data = df)
# Display scatter plot diagram for the feature variable 'horsepower' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'horsepower', y = 'fuel_consumption', data = df)
# Display scatter plot diagram for the feature variable 'weight' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'weight', y = 'fuel_consumption', data = df)
# Display scatter plot diagram for the feature variable 'acceleration' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'acceleration', y = 'fuel_consumption', data = df)
# Display scatter plot diagram for the feature variable 'car_age' and the label variable 'fuel_consumption'
sns.scatterplot(x = 'car_age', y = 'fuel_consumption', data = df)
# Display the correlations table/matrix
df.corr()
# Assign to a variable, plot on a heatmap and display the correlation matrix
sns.set(style = 'white')
corr_matrix = df.corr()
sns.heatmap(corr_matrix, vmin=None, vmax=None, cmap='YlGnBu', center=None, robust=False, annot=True, fmt='.2g', annot_kws=None, linewidths=2, linecolor='white', cbar=True, cbar_kws=None, cbar_ax=None, square=False, xticklabels='auto', yticklabels='auto', mask=None, ax=None)
# Display the correlation between the label 'fuel_consumption' variable and the featured variables sorted in a descending order.
df.corr()['fuel_consumption'].sort_values(ascending = False)
# <h2 id='5. Model Development, Training and Prediction'>5. Model Development, Training and Prediction</h2>
# <h3 id='5.1. Model Training'>5.1. Model Training</h3>
# +
# Create a list variable containing the columns name excluding the label variable 'fuel_consumption' as well as the "origin" and "car_name" columns.
col_names_regr = ['cylinders', 'displacement', 'horsepower', 'weight', 'acceleration', 'car_age']
X = df[col_names_regr]
Y = df['fuel_consumption']
# Split the data into train and test datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state = 1)
# Print the number of the datarows used to train and test the multiple linear regression model
print("The number of rows in the train dataset are", X_train.shape[0])
print("The number of rows in the test dataset are", X_test.shape[0])
# Train the regression model by using the fit() function
MLR = LinearRegression()
MLR.fit(X_train, Y_train)
# -
# <h3 id='5.2. Model Predictions and Evaluation Metrics'>5.2. Model Predictions and Evaluation Metrics</h3>
# +
# Make predictions with the fitted model
Y_hat = MLR.predict(X_test)
# Display the value of the coefficient of determiantion(R-squared) calculated with the test dataset
R_squared_test = MLR.score(X_test, Y_test)
# Round the R-squared value
R_squared_test = round(R_squared_test, 3)
# Print the values of the regression coefficients, some predicted values and the R-squared value
print('\nThe intercept value of the multiple linear regression model is b0 =', round(MLR.intercept_,8))
print('\nThe coefficients b1, b2, ..., b11 of the multiple linear regression model of type')
print('\nY_hat = b0 + b1.X1 + b2.X2 + b3.X3 + b4.X4 + b5.X5 + b6.X6 have values: ')
print(MLR.coef_)
print('\nThe Multiple Linear Regression Model is:')
print('fuel_consumption = ')
print(' -15.51909647')
print(' +0.30048785.cylinders')
print(' -0.00453743.displacement')
print(' +0.02804279.horsepower')
print(' +0.00586874.weight')
print(' +0.0878004.acceleration')
print(' +0.31352031.car_age')
print('\nThe predicted values for cars fuel consumption per 100km are:')
print(Y_hat, sep = '\n')
print('\nThe coefficient of determination R-squared calculated for the test dataset is: ')
print('R-squared_test =', round(R_squared_test,2))
# -
# <h2 id='6. References'>6. References</h2>
# [1] <a href='https://www.python-course.eu/pandas_replacing_values.php'>Accessing and Changing values of DataFrames</a><br>
# [2] <a href='https://pythonhow.com/accessing-dataframe-columns-rows-and-cells/'>Accessing pandas dataframe columns, rows, and cells</a><br>
# [3] <a href='https://archive.ics.uci.edu/ml/machine-learning-databases/auto-mpg/'> Auto MPG Data Folder</a><br>
# [4] <a href='https://archive.ics.uci.edu/ml/datasets/Auto+MPG'>Auto MPG Data Set</a><br>
# [5] <a href='https://www.geeksforgeeks.org/check-multiple-conditions-in-if-statement-python/'>Check multiple conditions in if statement – Python</a><br>
# [6] <a href='https://stackoverflow.com/questions/34682828/extracting-specific-selected-columns-to-new-dataframe-as-a-copy'>Extracting specific selected columns to new DataFrame as a copy</a><br>
# [7] <a href='https://www.datacamp.com/community/tutorials/for-loops-in-python'>For Loops in Python</a><br>
# [8] <a href='https://www.thecalculatorsite.com/conversions/fuelconsumption.php'>Fuel Consumption Converter</a><br>
# [9] <a href='https://stackoverflow.com/questions/19632728/how-do-i-get-a-python-program-to-do-nothing/19632742'>How do I get a python program to do nothing?</a><br>
# [10] <a href='https://stackoverflow.com/questions/15943769/how-do-i-get-the-row-count-of-a-pandas-dataframe'>How do I get the row count of a pandas DataFrame?</a><br>
# [11] <a href='https://towardsdatascience.com/how-to-change-datatypes-in-pandas-in-4-minutes-677addf9a409'>How To Change DataTypes In Pandas in 4 Minutes</a><br>
# [12] <a href='https://stackoverflow.com/questions/30071886/how-to-get-current-time-in-python-and-break-up-into-year-month-day-hour-minu'>How to get current time in python and break up into year, month, day, hour, minute?</a><br>
# [13] <a href='https://stackoverflow.com/questions/41906199/how-to-make-a-new-line-in-a-jupyter-markdown-cell'>How to make a new line in a jupyter markdown cell</a><br>
# [14] <a href='https://stackoverflow.com/questions/28080066/how-to-reference-a-ipython-notebook-cell-in-markdown'>How to reference a IPython notebook cell in markdown?</a><br>
# [15] <a href='https://stackoverflow.com/questions/31554574/html-padding-style'>HTML Padding Style</a><br>
# [16] <a href='https://pandas.pydata.org/pandas-docs/stable/user_guide/io.html#io-fwf-reader'>IO tools (text, CSV, HDF5, …)</a><br>
# [17] <a href='https://jupyter.brynmawr.edu/services/public/dblank/Jupyter%20Notebook%20Users%20Manual.ipynb'>Jupyter Notebook Users Manual</a><br>
# [18] <a href='https://realpython.com/linear-regression-in-python/'>Linear Regression in Python by <NAME></a><br>
# [19] <a href='https://en.wikipedia.org/wiki/Machine_learning'>Machine learning</a><br>
# [20] <a href='https://www.ritchieng.com/machine-learning-linear-regression/'>Machine Learning Linear Regression</a><br>
# [21] <a href='https://www.datacamp.com/community/tutorials/markdown-in-jupyter-notebook'>Markdown in Jupyter Notebook</a><br>
# [22] <a href='https://www.mathsisfun.com/metric-imperial-conversion-charts.html'>Metric Conversion Chart</a><br>
# [23] <a href='https://www.asknumbers.com/mpg-to-L100km.aspx'>MPG to L/100 KM Conversion</a><br>
# [24] <a href='https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.core.groupby.GroupBy.mean.html'>pandas.core.groupby.GroupBy.mean</a><br>
# [25] <a href='https://pandas.pydata.org/pandas-docs/version/0.25.3/reference/api/pandas.core.groupby.GroupBy.mean.html'>pandas.core.groupby.GroupBy.mean (additional source)</a><br>
# [26] <a href='https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.replace.html'>pandas.DataFrame.replace</a><br>
# [27] <a href='https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.to_csv.html'>pandas.DataFrame.to_csv</a><br>
# [28] <a href='https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_numeric.html#pandas.to_numeric'>pandas.to_numeric</a><br>
# [29] <a href='https://seaborn.pydata.org/examples/many_pairwise_correlations.html'>Plotting a diagonal correlation matrix</a><br>
# [30] <a href='https://en.wikipedia.org/wiki/Python_(programming_language)'>Python (programming language)</a><br>
# [31] <a href='https://www.guru99.com/round-function-python.html'>Python round() function with Examples</a><br>
# [32] <a href='https://en.wikipedia.org/wiki/Regression_analysis'>Regression analysis</a><br>
# [33] <a href='https://datatofish.com/replace-nan-values-with-zeros/'>Replace NaN Values with Zeros in Pandas DataFrame</a><br>
# [34] <a href='https://seaborn.pydata.org/generated/seaborn.heatmap.html'>seaborn.heatmap</a><br>
# [35] <a href='https://seaborn.pydata.org/generated/seaborn.pairplot.html'>seaborn.pairplot</a><br>
# [36] <a href='https://pythonbasics.org/seaborn-pairplot/'>Seaborn pairplot example</a><br>
# [37] <a href='https://indianaiproduction.com/seaborn-pairplot/'>Seaborn Pairplot In Detail | Python Seaborn Tutorial</a><br>
# [38] <a href='https://seaborn.pydata.org/generated/seaborn.scatterplot.html'>seaborn.scatterplot</a><br>
# [39] <a href='https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html'>sklearn.linear_model.LinearRegression</a><br>
# [40] <a href='https://jupyter-notebook.readthedocs.io/en/stable/notebook.html'>The Jupyter Notebook User Documentation</a><br>
# [41] <a href='https://docs.python.org/2/library/datetime.html#datetime.datetime.now'>The Python Standard Library: 8.1. datetime — Basic date and time types</a><br>
# [42] <a href='https://www.yourmechanic.com/article/what-is-engine-displacement'>What Is Engine Displacement?</a><br>
# [43] <a href='https://stackoverflow.com/questions/16923281/writing-a-pandas-dataframe-to-csv-file'>Writing a pandas DataFrame to CSV file</a><br>
# <h2 id='About the Author'>About the Author</h2>
# <p><a href = 'https://www.linkedin.com/in/stavridimitrov/'><NAME></a> is a Data Science enthusiast who has an extensive experience in data analysis, software design, development, testing, implementation and applications support. He has worked as a Data Scientist, Data Analyst, Software Developer and Application Support Analyst / Programmer. While working as a University Lecturer/Assistant Professor, Stavri has been involved in research work and teaching activities. Over the years he has participated in research projects and studies in the area of software development, transportation engineering and supply chains. Stavri holds a PhD degree in Engineering from the University of Auckland, New Zealand.</p>
|
Machine Learning Model for Predicting Cars' Fuel Consumption.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# %pylab inline
import pandas
d = pandas.read_csv("data/movie_reviews.tsv", delimiter="\t")
# Holdout split
split = 0.7
d_train = d[:int(split*len(d))]
d_test = d[int((1-split)*len(d)):]
# +
## tf-idf
# -
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer(strip_accents='unicode', stop_words='english', min_df=1, max_features=None, norm="l2")
features_w2v = vectorizer.fit_transform(d_train.review)
test_features_w2v = vectorizer.transform(d_test.review)
# +
from sklearn.metrics import accuracy_score, roc_auc_score, classification_report, roc_curve, precision_score, recall_score
def performance(y_true, pred, color="g", ann=True):
acc = accuracy_score(y_true, pred[:,1] > 0.5)
precision = precision_score(y_true, pred[:,1] > 0.5, average='binary')
recall = recall_score(y_true, pred[:,1] > 0.5, average='binary')
auc = roc_auc_score(y_true, pred[:,1])
fpr, tpr, thr = roc_curve(y_true, pred[:,1])
plot(fpr, tpr, color, linewidth="3")
xlabel("False positive rate")
ylabel("True positive rate")
if ann:
annotate("Acc: %0.2f" % acc, (0.1,0.8), size=14)
annotate("Precision: %0.2f" % precision, (0.1,0.7), size=14)
annotate("Recall: %0.2f" % recall, (0.1,0.6), size=14)
annotate("AUC: %0.2f" % auc, (0.1,0.5), size=14)
# +
# Random Forest
from sklearn.ensemble import RandomForestClassifier
model_rf = RandomForestClassifier(n_estimators=100, n_jobs=-1)
model_rf.fit(features_w2v, d_train.sentiment)
pred_rf = model_rf.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred_rf)
# +
# Navie Bayes
from sklearn.naive_bayes import MultinomialNB
model_NB = MultinomialNB()
model_NB.fit(features_w2v, d_train.sentiment)
pred_NB = model_NB.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred_NB, color="b")
# +
# K-Nearest Neighbours
from sklearn.neighbors import KNeighborsClassifier
model_KNN = KNeighborsClassifier(n_neighbors=5)
model_KNN.fit(features_w2v, d_train.sentiment)
pred_KNN = model_KNN.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred_KNN, color="c")
# +
# Neural network models
from sklearn.neural_network import MLPClassifier
model_NNM = MLPClassifier(solver='lbfgs', alpha=1e-5,hidden_layer_sizes=(5, 2), random_state=1)
model_NNM.fit(features_w2v, d_train.sentiment)
pred_NNM = model_NNM.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred_NNM, color="r")
print len(d_test.sentiment)
# +
# Support Vector Classification
from sklearn.svm import SVC
model_SVC = SVC(probability=True)
model_SVC.fit(features_w2v, d_train.sentiment)
pred_SVC = model_SVC.predict_proba(test_features_w2v)
performance(d_test.sentiment, pred_SVC, color="pink")
# -
performance(d_test.sentiment, pred_rf,ann = False)
performance(d_test.sentiment, pred_NB, color="b",ann = False)
performance(d_test.sentiment, pred_KNN, color="c",ann = False)
performance(d_test.sentiment, pred_NNM, color="r",ann = False)
performance(d_test.sentiment, pred_SVC, color="pink",ann = False)
|
sentiment analysis for movie review 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from urllib import urlopen
data_address = "https://www.group.pictet/corporate/fr/home/intermediaries/lpp_indices/lpp2000.html"
test_address = "http://pythonscraping.com/pages/page1.html"
warandpeace_address = "http://www.pythonscraping.com/pages/warandpeace.html"
html = urlopen(test_address)
html_data = urlopen(data_address)
html_litt = urlopen(warandpeace_address)
from bs4 import BeautifulSoup
bs_obj = BeautifulSoup(html.read())
data_bs_obj = BeautifulSoup(html_data.read())
bs_obj.contents
litt_bs = BeautifulSoup(html_litt)
nameList = litt_bs.findAll("span", {"class":"green"})
for name in nameList:
print(name.get_text())
litt_bs
|
web_scraping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Reducing Failure-Inducing Inputs
#
# A standard problem in debugging is this: Your program fails after processing some large input. Only a _part_ of this input, however, is responsible for the failure. _Reducing_ the input to a failure-inducing minimum not only eases debugging – it also helps in understanding why and when the program fails. In this chapter, we present techniques that _automatically reduce and simplify failure-inducing inputs to a minimum_, notably the popular _Delta Debugging_ technique.
# -
from bookutils import YouTubeVideo
YouTubeVideo("6fmJ5l257bM")
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Prerequisites**
#
# * Using the "delta debugging" technique for reduction has no specific prerequisites.
# * To understand the `DeltaDebugger` implementation, reading [the chapter on tracing](Tracer.ipynb) is recommended.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# This chapter is adapted from [a similar chapter in "The Fuzzing Book"](https://www.fuzzingbook.org/html/Reducer.html). The material has been adapted to be independent from the `fuzzingbook` infrastructure, to build on general delta debugging (`dd`), and to provide a simpler invocation interface.
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from debuggingbook.DeltaDebugger import <identifier>
# ```
#
# and then make use of the following features.
#
#
# A _reducer_ takes a failure-inducing input and reduces it to the minimum that still reproduces the failure. This chapter provides a `DeltaDebugger` class that implements such a reducer.
#
# Here is a simple example: An arithmetic expression causes an error in the Python interpreter:
#
# ```python
# >>> def myeval(inp: str) -> Any:
# >>> return eval(inp)
# >>> with ExpectError(ZeroDivisionError):
# >>> myeval('1 + 2 * 3 / 0')
# ```
# Can we reduce this input to a minimum? _Delta Debugging_ is a simple and robust reduction algorithm. We provide a `DeltaDebugger` class that is used in conjunction with a (failing) function call:
#
# ```python
# with DeltaDebugger() as dd:
# fun(args...)
# dd
# ```
#
# The class automatically determines minimal arguments that cause the function to fail with the same exception as the original. Printing out the class object reveals the minimized call.
#
# ```python
# >>> with DeltaDebugger() as dd:
# >>> myeval('1 + 2 * 3 / 0')
# >>> dd
# ```
# The input is reduced to the maximum: We get the essence of the division by zero.
#
# There also is an interface to access the reduced input(s) programmatically. The method `min_args()` returns a dictionary in which all function arguments are minimized:
#
# ```python
# >>> dd.min_args()
# ```
# In contrast, `max_args()` returns a dictionary in which all function arguments are maximized, but still pass:
#
# ```python
# >>> dd.max_args()
# ```
# The method `min_arg_diff()` returns a triple of
# * passing input,
# * failing input, and
# * their minimal failure-inducing difference:
#
# ```python
# >>> dd.min_arg_diff()
# ```
# And you can also access the function itself, as well as its original arguments.
#
# ```python
# >>> dd.function().__name__, dd.args()
# ```
# `DeltaDebugger` processes (i.e., minimizes or maximizes) all arguments that support a `len()` operation and that can be indexed – notably _strings_ and _lists_. If a function has multiple arguments, all arguments that can be processed will be processed.
#
# This chapter also provides a number of superclasses to `DeltaDebugger`, notably `CallCollector`, which obtains the first function call for `DeltaDebugger`. `CallReducer` classes allow for implementing alternate call reduction strategies.
#
#
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Why Reducing?
#
# A common problem in debugging is that given an input, only a _small part of that input may be responsible for the failure_. A central part of debugging is to _identify_ these parts – and to simplify (or _reduce_) the input to a minimal form that reproduces the failure – but does and contains as little else as possible.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# Here's an example of such a situation. We have a `mystery()` method that – given its code – can occasionally fail. But under which circumstances does this actually happen? We have deliberately obscured the exact condition in order to make this non-obvious.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# -
import Tracer
from bookutils import quiz
def mystery(inp: str) -> None:
x = inp.find(chr(0o17 + 0o31))
y = inp.find(chr(0o27 + 0o22))
if x >= 0 and y >= 0 and x < y:
raise ValueError("Invalid input")
else:
pass
# To find an input that causes the function to fail, let us _fuzz_ it – that is, feed it with random inputs – until we find a failing input. There's [entire books about fuzzing](https://fuzzingbook.org); but here, a very simple `fuzz()` function for this purpose will already suffice.
# To build a fuzzer, we need random inputs – and thus a source for randomness. The function `random.randrange(a, b)` returns a random number in the range (a, b).
import random
random.randrange(32, 128)
# We can use `random.randrange()` to compose random (printable) characters:
def fuzz() -> str:
length = random.randrange(10, 70)
fuzz = ""
for i in range(length):
fuzz += chr(random.randrange(32, 127))
return fuzz
# Here are some random strings produced by our `fuzz()` function:
for i in range(6):
print(repr(fuzz()))
# Let us now use `fuzz()` to find an input where `mistery()` fails:
while True:
fuzz_input = fuzz()
try:
mystery(fuzz_input)
except ValueError:
break
# This is an input that causes `mystery()` to fail:
failing_input = fuzz_input
failing_input
len(failing_input)
from ExpectError import ExpectError
with ExpectError(ValueError):
mystery(failing_input)
# Something in this input causes `mystery()` to fail. But what is it?
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Manual Input Reduction
#
# One important step in the debugging process is _reduction_ – that is, to identify those circumstances of a failure that are relevant for the failure to occur, and to _omit_ (if possible) those parts that are not. As Kernighan and Pike \cite{Kernighan1999} put it:
#
# > For every circumstance of the problem, check whether it is relevant for the problem to occur. If it is not, remove it from the problem report or the test case in question.
# -
# Specifically for inputs, they suggest a _divide and conquer_ process:
#
# > Proceed by binary search. Throw away half the input and see if the output is still wrong; if not, go back to the previous state and discard the other half of the input.
#
# This is something we can easily try out, using our last generated input:
failing_input
# For instance, we can see whether the error still occurs if we only feed in the first half:
half_length = len(failing_input) // 2 # // is integer division
first_half = failing_input[:half_length]
first_half
with ExpectError(ValueError):
mystery(first_half)
# Nope – the first half alone does not suffice. Maybe the second half?
second_half = failing_input[half_length:]
assert first_half + second_half == failing_input
second_half
with ExpectError(ValueError):
mystery(second_half)
# This did not go so well either. We may still proceed by cutting away _smaller chunks_ – say, one character after another. If our test is deterministic and easily repeated, it is clear that this process eventually will yield a reduced input. But still, it is a rather inefficient process, especially for long inputs. What we need is a _strategy_ that effectively minimizes a failure-inducing input – a strategy that can be automated.
# ## Delta Debugging
# One strategy to effectively reduce failure-inducing inputs is _delta debugging_ \cite{Zeller2002}. Delta Debugging implements the "binary search" strategy, as listed above, but with a twist: If neither half fails (also as above), it keeps on cutting away smaller and smaller chunks from the input, until it eliminates individual characters. Thus, after cutting away the first half, we cut away
# the first quarter, the second quarter, and so on.
# Let us illustrate this on our example, and see what happens if we cut away the first quarter.
quarter_length = len(failing_input) // 4
input_without_first_quarter = failing_input[quarter_length:]
input_without_first_quarter
with ExpectError(ValueError):
mystery(input_without_first_quarter)
# Ah! This has failed, and reduced our failing input by 25%. Let's remove another quarter.
input_without_first_and_second_quarter = failing_input[quarter_length * 2:]
input_without_first_and_second_quarter
with ExpectError(ValueError):
mystery(input_without_first_and_second_quarter)
# This is not too surprising, as we had that one before:
second_half
input_without_first_and_second_quarter
# How about removing the third quarter, then?
input_without_first_and_third_quarter = failing_input[quarter_length:
quarter_length * 2] + failing_input[quarter_length * 3:]
input_without_first_and_third_quarter
with ExpectError(ValueError):
mystery(input_without_first_and_third_quarter)
# Yes! This has succeeded. Our input is now 50% smaller.
# We have now tried to remove pieces that make up $\frac{1}{2}$ and $\frac{1}{4}$ of the original failing string. In the next iteration, we would go and remove even smaller pieces – $\frac{1}{8}$, $\frac{1}{16}$ and so on. We continue until we are down to $\frac{1}{26}$ – that is, individual characters.
# However, this is something we happily let a computer do for us – and this is what the _Delta Debugging_ algorithm does. Delta Debugging implements the strategy sketched above: It first removes larger chunks of size $\frac{1}{2}$; if this does not fail, then we proceed to chunks of size $\frac{1}{4}$, then $\frac{1}{8}$ and so on.
# Our `ddmin()` implementation uses the exact same Python code as Zeller in \cite{Zeller2002}; the only difference is that it has been adapted to work on Python 3. The variable `n` (initially 2) indicates the granularity – in each step, chunks of size $\frac{1}{n}$ are cut away. If none of the test fails (`some_complement_is_failing` is False), then `n` is doubled – until it reaches the length of the input.
PASS = 'PASS'
FAIL = 'FAIL'
UNRESOLVED = 'UNRESOLVED'
# ignore
from typing import Sequence, Any, Callable, Optional, Type, Tuple
from typing import Dict, Union, Set, List, FrozenSet, cast
def ddmin(test: Callable, inp: Sequence, *test_args: Any) -> Sequence:
"""Reduce the input inp, using the outcome of test(fun, inp)."""
assert test(inp, *test_args) != PASS
n = 2 # Initial granularity
while len(inp) >= 2:
start = 0
subset_length = int(len(inp) / n)
some_complement_is_failing = False
while start < len(inp):
complement = (inp[:int(start)] + inp[int(start + subset_length):]) # type: ignore
if test(complement, *test_args) == FAIL:
inp = complement
n = max(n - 1, 2)
some_complement_is_failing = True
break
start += subset_length
if not some_complement_is_failing:
if n == len(inp):
break
n = min(n * 2, len(inp))
return inp
# To see how `ddmin()` works, let us run it on our failing input. We need to define a `test` function that returns PASS or FAIL, depending on the test outcome. This `generic_test()` assumes that the function fails if it raises an exception (such as an `AssertException`), and passes otherwise. The optional argument `expected_exc` specifies the name of exception to be checked for; this ensures we reduce only for the kind of error raised in the original failure.
def generic_test(inp: Sequence, fun: Callable,
expected_exc: Optional[Type] = None) -> str:
result = None
detail = ""
try:
result = fun(inp)
outcome = PASS
except Exception as exc:
detail = f" ({type(exc).__name__}: {str(exc)})"
if expected_exc is None:
outcome = FAIL
elif type(exc) == type(expected_exc) and str(exc) == str(expected_exc):
outcome = FAIL
else:
outcome = UNRESOLVED
print(f"{fun.__name__}({repr(inp)}): {outcome}{detail}")
return outcome
# We can now invoke `ddmin()` in our setting. With each step, we see how the remaining input gets smaller and smaller, until only two characters remain:
ddmin(generic_test, failing_input, mystery, ValueError('Invalid input'))
# Now we know why `mystery()` fails – it suffices that the input contains two matching parentheses. Delta Debugging determines this in 25 steps. Its result is _1-minimal_, meaning that every character contained is required to produce the error; removing any (as seen in the last two tests, above) no longer causes the test to fail. This property is guaranteed by the delta debugging algorithm, which in its last stage always tries to delete characters one by one.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# A reduced test case such as the one above has many advantages:
#
# * A reduced test case __reduces the _cognitive load_ of the programmer__. The test case is shorter and focused, and thus does not burden the programmer with irrelevant details. A reduced input typically leads to shorter executions and smaller program states, both of which reduce the search space as it comes to understanding the bug. In our case, we have eliminated lots of irrelevant input – only the two characters the reduced input contains are relevant.
#
# * A reduced test case __is easier to communicate__. All one needs here is the summary: `mystery() fails on "()"`, which is much better than `mystery() fails on a 4100-character input (attached)`.
#
# * A reduced test case helps in __identifying duplicates__. If similar bugs have been reported already, and all of them have been reduced to the same cause (namely that the input contains matching parentheses), then it becomes obvious that all these bugs are different symptoms of the same underlying cause – and would all be resolved at once with one code fix.
# -
# How effective is delta debugging? In the best case (when the left half or the right half fails), the number of tests is logarithmic proportional to the length $n$ of an input (i.e., $O(\log_2 n)$); this is the same complexity as binary search. In the worst case, though, delta debugging can require a number of tests proportional to $n^2$ (i.e., $O(n^2)$) – this happens in the case when we are down to character granularity, and we have to repeatedly tried to delete all characters, only to find that deleting the last character results in a failure \cite{Zeller2002}. (This is a pretty pathological situation, though.)
# In general, delta debugging is a robust algorithm that is easy to implement, easy to deploy, and easy to use – provided that the underlying test case is deterministic and runs quickly enough to warrant a number of experiments. In general, any debugging task should start with simplifying the test case as much as possible – and this is where delta debugging can help.
# ## A Simple DeltaDebugger Interface
#
# As defined above, using `ddmin()` still requires the developer to set up a special testing function – and writing or using even a generic tester (like `generic_test()`) takes some effort. We want to simplify the setup such that only two lines of Python is required.
#
# Our aim is to have a `DeltaDebugger` class that we can use in conjunction with a failing (i.e., exception raising) function call:
#
# ```python
# with DeltaDebugger() as dd:
# mystery(failing_input)
# dd
# ```
# Here, at the end of the `with` statement, printing out `dd` shows us the minimal input that causes the failure.
# ### Excursion: Implementing DeltaDebugger
# Our interface consist of six building blocks:
#
# 1. We collect the name and args of the first call in the `with` body, as well as the exception it raises.
# 2. We set up an infrastructure such that we can repeat calls with different arguments.
# 3. We make sure that multiple tests with the same arguments can return outcomes from a cache.
# 4. We create a `DeltaDebugger` class that implements the general Delta Debugging algorithm – an algorithm than can minimize failing inputs as well as maximize passing inputs.
# 5. We provide an infrastructure that applies Delta Debugging on multiple arguments.
# 6. Finally, custom methods like `min_args()` allow to invoke delta debugging on arguments.
# #### Collecting a Call
#
# We start by creating an infrastructure that collects a call. The `CallCollector` class saves the first call observed in `_function`, `_args`, and `_exception` attributes, respectively; it then turns tracing off.
import sys
from types import FunctionType, FrameType, TracebackType
from StackInspector import StackInspector
class NoCallError(ValueError):
pass
class CallCollector(StackInspector):
"""
Collect an exception-raising function call f().
Use as `with CallCollector(): f()`
"""
def __init__(self) -> None:
"""Initialize collector"""
self.init()
def init(self) -> None:
"""Reset for new collection."""
self._function: Optional[Callable] = None
self._args: Dict[str, Any] = {}
self._exception: Optional[BaseException] = None
self.original_trace_function: Optional[Callable] = None
def traceit(self, frame: FrameType, event: str, arg: Any) -> None:
"""Tracing function. Collect first call, then turn tracing off."""
if event == 'call':
name = frame.f_code.co_name
if name.startswith('__'):
# Internal function
return
if self._function is not None:
# Already set
return
func = self.search_func(name, frame)
if func:
self._function = func
else:
# Create new function from given code
self._function = self.create_function(frame)
self._args = {} # Create a local copy of args
for var in frame.f_locals:
if var in frame.f_code.co_freevars:
continue # Local var, not an argument
self._args[var] = frame.f_locals[var]
# Turn tracing off
sys.settrace(self.original_trace_function)
def after_collection(self) -> None:
"""Called after collection. To be defined in subclasses."""
pass
def args(self) -> Dict[str, Any]:
"""Return the dictionary of collected arguments."""
return self._args
def function(self) -> Callable:
"""Return the function called."""
if self._function is None:
raise NoCallError("No function call collected")
return self._function
def exception(self) -> Optional[BaseException]:
"""Return the exception produced, or `None` if none."""
return self._exception
def format_call(self, args: Optional[Dict[str, Any]] = None) -> str:
...
def format_exception(self, exc: Optional[BaseException] = None) -> str:
...
def call(self, new_args: Optional[Dict[str, Any]] = None) -> Any:
...
# A `CallCollector` is used like a `Tracer` from the [chapter on tracing](Tracer.ipynb), using a `with` block to collect a single function call.
class CallCollector(CallCollector):
def __enter__(self) -> Any:
"""Called at begin of `with` block. Turn tracing on."""
self.init()
self.original_trace_function = sys.gettrace()
sys.settrace(self.traceit)
return self
def __exit__(self, exc_tp: Type, exc_value: BaseException,
exc_traceback: TracebackType) -> Optional[bool]:
"""Called at end of `with` block. Turn tracing off."""
sys.settrace(self.original_trace_function)
if not self._function:
if exc_tp:
return False # re-raise exception
else:
raise NoCallError("No call collected")
if self.is_internal_error(exc_tp, exc_value, exc_traceback):
return False # Re-raise exception
self._exception = exc_value
self.after_collection()
return True # Ignore exception
# Here are the attributes as collected by `CallCollector` for our `mystery()` function. Note that the `mystery()` exception is "swallowed" by the `CallCollector`.
with CallCollector() as call_collector:
mystery(failing_input)
call_collector.function()
call_collector.args()
call_collector.exception()
# If an error occurs _before_ the first function call takes place, the exception is simply re-raised.
with ExpectError(NameError):
with CallCollector() as c:
some_error() # type: ignore
# #### Repeating a Call
#
# Our second step is an infrastructure such that we can call the function collected earlier with alternate arguments. We can call the function directly via the collected `_function` attribute:
call_collector.function()("foo")
with ExpectError(ValueError):
call_collector.function()(failing_input)
# We can also provide the arguments collected during the call:
with ExpectError(ValueError):
call_collector.function()(**call_collector.args())
# Our `call()` method calls the collected function using this construct. It also allows to _change_ individual arguments by providing a `new_args` dictionary of variable names to new values.
class CallCollector(CallCollector):
def call(self, new_args: Optional[Dict[str, Any]] = None) -> Any:
"""
Call collected function. If `new_args` is given,
override arguments from its {var: value} entries.
"""
if new_args is None:
new_args = {}
args = {} # Create local copy
for var in self.args():
args[var] = self.args()[var]
for var in new_args:
args[var] = new_args[var]
return self.function()(**args)
# Using simply `call()` without arguments reproduces the failure:
with CallCollector() as call_collector:
mystery(failing_input)
with ExpectError(ValueError):
call_collector.call()
# We can also supply alternate arguments (and get alternate outcomes):
call_collector.call({'inp': 'foo'})
# We close with two helper functions that come handy for logging and error messages:
class CallCollector(CallCollector):
def format_call(self, args: Optional[Dict[str, Any]] = None) -> str:
"""Return a string representing a call of the function with given args."""
if args is None:
args = self.args()
return self.function().__name__ + "(" + \
", ".join(f"{arg}={repr(args[arg])}" for arg in args) + ")"
def format_exception(self, exc: Optional[BaseException] = None) -> str:
"""Return a string representing the given exception."""
if exc is None:
exc = self.exception()
s = type(exc).__name__
if str(exc):
s += ": " + str(exc)
return s
with CallCollector() as call_collector:
mystery(failing_input)
call_collector.format_call()
call_collector.format_exception()
# #### Testing, Logging, and Caching
#
# Our next to last step is an infrastructure that implements delta debugging for the collected call.
# We first introduce a `CallReducer` class as an abstract superclass for all kinds of reducers.
# Its `run()` method tests the function and returns PASS, FAIL, or UNRESOLVED. As with `generic_test()`, above, we check for exception type and exact error message.
class CallReducer(CallCollector):
def __init__(self, *, log: Union[bool, int] = False) -> None:
"""Initialize. If `log` is True, enable logging."""
super().__init__()
self.log = log
self.reset()
def reset(self) -> None:
"""Reset the number of tests."""
self.tests = 0
def run(self, args: Dict[str, Any]) -> str:
"""
Run collected function with `args`. Return
* PASS if no exception occurred
* FAIL if the collected exception occurred
* UNRESOLVED if some other exception occurred.
Not to be used directly; can be overloaded in subclasses.
"""
try:
result = self.call(args)
except Exception as exc:
self.last_exception = exc
if (type(exc) == type(self.exception()) and
str(exc) == str(self.exception())):
return FAIL
else:
return UNRESOLVED # Some other failure
self.last_result = result
return PASS
# The `test()` method runs a single test (with logging, if wanted); the `reduce_arg()` method will eventually reduce an input to the minimum.
class CallReducer(CallReducer):
def test(self, args: Dict[str, Any]) -> str:
"""Like run(), but also log detail and keep statistics."""
outcome = self.run(args)
if outcome == PASS:
detail = ""
else:
detail = f" ({self.format_exception(self.last_exception)})"
self.tests += 1
if self.log:
print(f"Test #{self.tests} {self.format_call(args)}: {outcome}{detail}")
return outcome
def reduce_arg(self, var_to_be_reduced: str, args: Dict[str, Any]) -> Sequence:
"""
Determine and return a minimal value for var_to_be_reduced.
To be overloaded in subclasses.
"""
return args[var_to_be_reduced]
# Here's some logging output from the `test()` function:
# +
with CallReducer(log=True) as reducer:
mystery(failing_input)
reducer.test({'inp': failing_input})
reducer.test({'inp': '123'})
reducer.test({'inp': '123'})
# -
# The `CachingCallReducer` variant saves test results, such that we don't have to run the same tests again and again:
class CachingCallReducer(CallReducer):
"""Like CallReducer, but cache test outcomes."""
def init(self) -> None:
super().init()
self._cache: Dict[FrozenSet, str] = {}
def test(self, args: Dict[str, Any]) -> str:
# Create a hashable index
try:
index = frozenset((k, v) for k, v in args.items())
except TypeError:
index = frozenset()
if not index:
# Non-hashable value – do not use cache
return super().test(args)
if index in self._cache:
return self._cache[index]
outcome = super().test(args)
self._cache[index] = outcome
return outcome
# If we now repeat a test with the same argument, its outcome can be found in the cache:
# +
with CachingCallReducer(log=True) as reducer:
mystery(failing_input)
reducer.test({'inp': failing_input})
reducer.test({'inp': '123'})
reducer.test({'inp': '123'})
# -
# #### General Delta Debugging
#
# The `DeltaDebugger` class finally implements Delta Debugging on arguments. Our implementation uses the _general_ `dd` delta debugging algorithm from \cite{Zeller2002}. In contrast to `ddmin`, it returns a _pair_ of a maximized passing input and a minimized failing input. The algorithm can be customized, however, to leave the passing input fixed and _only_ to minimize the failing input (as with `ddmin`), or vice versa.
# Internally, `dd` does not directly work on a list of elements; instead, it works on sets of _indices_ into such a list. The function `to_set()` converts a collection into such a set.
def to_set(inp: Sequence) -> Set:
"""Convert inp into a set of indices"""
return set(range(len(inp)))
to_set("abcd")
# The function `from_set()` converts a set of indices back into the original collection. For this, it uses a function `empty()` that returns an empty collection that has the same type as the given input `inp`.
def empty(inp: Any) -> Any:
"""Return an "empty" element of the same type as inp"""
return type(inp)()
empty("abc"), empty([1, 2, 3]), empty({0, -1, -2})
# The function `add_to()` tries out various ways to add an element to a given collection.
def add_to(collection: Any, elem: Any) -> Any:
"""Add element to collection; return new collection."""
if isinstance(collection, str):
return collection + elem # Strings
try: # Lists and other collections
return collection + type(collection)([elem])
except TypeError:
pass
try: # Sets
return collection | type(collection)([elem])
except TypeError:
pass
raise ValueError("Cannot add element to collection")
add_to("abc", "d"), add_to([1, 2, 3], 4), add_to(set([1, 2, 3]), 4)
# Using `empty()` and `add_to()`, we can now implement `from_set()`:
def from_set(the_set: Any, inp: Sequence) -> Any:
"""Convert a set of indices into `inp` back into a collection."""
ret = empty(inp)
for i, c in enumerate(inp):
if i in the_set:
ret = add_to(ret, c)
return ret
from_set({1, 2}, "abcd")
# To split a set of elements into `n` subsets of equal size, we use this helper function, based on [this discussion in StackOverflow](https://stackoverflow.com/questions/2130016/splitting-a-list-into-n-parts-of-approximately-equal-length/37414115#37414115).
def split(elems: Any, n: int) -> List:
assert 1 <= n <= len(elems)
k, m = divmod(len(elems), n)
try:
subsets = list(elems[i * k + min(i, m):(i + 1) * k + min(i + 1, m)]
for i in range(n))
except TypeError:
# Convert to list and back
subsets = list(type(elems)(
list(elems)[i * k + min(i, m):(i + 1) * k + min(i + 1, m)])
for i in range(n))
assert len(subsets) == n
assert sum(len(subset) for subset in subsets) == len(elems)
assert all(len(subset) > 0 for subset in subsets)
return subsets
for n in range(1, 8):
print(split([1, 2, 3, 4, 5, 6, 7], n))
split("abcd", 3)
split({1, 2, 3, 4, 5, 6, 7}, 3)
# With these, we can now implement general delta debugging. Our implementation follows \cite{Zeller2002} with the following optimizations:
#
# 1. We can control whether only to minimize or to maximize ("mode")
# 2. The operations "Reduce to subset" and "Increase to subset" are only taken while the number of subsets is still 2.
# 3. If "Reduce to subset" and "Increase to subset" are successful, the offset is set to `i` (not `0`) to distribute reduction operations more evenly across the input. (Thanks to <NAME> and <NAME> to point out this issue!)
class NotFailingError(ValueError):
pass
class NotPassingError(ValueError):
pass
class DeltaDebugger(CachingCallReducer):
def dd(self, var_to_be_reduced: str, fail_args: Dict[str, Any],
*, mode: str = '-') -> Tuple[Sequence, Sequence, Sequence]:
"""General Delta Debugging.
`var_to_be_reduced` - the name of the variable to reduce.
`fail_args` - a dict of (failure-inducing) function arguments,
with `fail_args[var_to_be_reduced]` - the input to apply dd on.
`mode`- how the algorithm should operate:
'-' (default): minimize input (`ddmin`),
'+': maximizing input (`ddmax`),
'+-': minimizing pass/fail difference (`dd`)
Returns a triple (`pass`, `fail`, `diff`) with
* maximized passing input (`pass`),
* minimized failing input (`fail`), and
* their difference `diff`
(elems that are in `fail`, but not in `pass`).
"""
def test(c: Set) -> str:
# Set up args
test_args = {}
for var in fail_args:
test_args[var] = fail_args[var]
test_args[var_to_be_reduced] = from_set(c, fail_inp)
return self.test(test_args)
def ret(c_pass: Set, c_fail: Set) -> \
Tuple[Sequence, Sequence, Sequence]:
return (from_set(c_pass, fail_inp),
from_set(c_fail, fail_inp),
from_set(c_fail - c_pass, fail_inp))
n = 2 # Initial granularity
fail_inp = fail_args[var_to_be_reduced]
c_pass = to_set([])
c_fail = to_set(fail_inp)
offset = 0
minimize_fail = '-' in mode
maximize_pass = '+' in mode
# Validate inputs
if test(c_pass) == FAIL:
if maximize_pass:
s_pass = repr(from_set(c_pass, fail_inp))
raise NotPassingError(
f"Input {s_pass} expected to pass, but fails")
else:
return ret(c_pass, c_pass)
if test(c_fail) == PASS:
if minimize_fail:
s_fail = repr(from_set(c_fail, fail_inp))
raise NotFailingError(
f"Input {s_fail} expected to fail, but passes")
else:
return ret(c_fail, c_fail)
# Main loop
while True:
if self.log > 1:
print("Passing input:", repr(from_set(c_pass, fail_inp)))
print("Failing input:", repr(from_set(c_fail, fail_inp)))
print("Granularity: ", n)
delta = c_fail - c_pass
if len(delta) < n:
return ret(c_pass, c_fail)
deltas = split(delta, n)
reduction_found = False
j = 0
while j < n:
i = (j + offset) % n
next_c_pass = c_pass | deltas[i]
next_c_fail = c_fail - deltas[i]
if minimize_fail and n == 2 and test(next_c_pass) == FAIL:
if self.log > 1:
print("Reduce to subset")
c_fail = next_c_pass
offset = i # was offset = 0 in original dd()
reduction_found = True
break
elif maximize_pass and n == 2 and test(next_c_fail) == PASS:
if self.log > 1:
print("Increase to subset")
c_pass = next_c_fail
offset = i # was offset = 0 in original dd()
reduction_found = True
break
elif minimize_fail and test(next_c_fail) == FAIL:
if self.log > 1:
print("Reduce to complement")
c_fail = next_c_fail
n = max(n - 1, 2)
offset = i
reduction_found = True
break
elif maximize_pass and test(next_c_pass) == PASS:
if self.log > 1:
print("Increase to complement")
c_pass = next_c_pass
n = max(n - 1, 2)
offset = i
reduction_found = True
break
else:
j += 1 # choose next subset
if not reduction_found:
if self.log > 1:
print("No reduction found")
if n >= len(delta):
return ret(c_pass, c_fail)
if self.log > 1:
print("Increase granularity")
n = min(n * 2, len(delta))
# By default, `dd()` minimizes inputs – just like `ddmin()`.
with DeltaDebugger() as dd:
mystery(failing_input)
# Its output is a triple of maximized passing input (if wanted), minimized failing input, and difference. Here is this triple for `mystery()`, just as with `ddmin()`:
mystery_pass, mystery_fail, mystery_diff = dd.dd('inp', {'inp': failing_input})
# The first element (`mystery_pass`) is the maximal passing input:
mystery_pass
# The second element (`mystery_fail`) is the minimal failing input:
mystery_fail
# And the third element (`mystery_diff`) is the difference between the two:
mystery_diff
# (Note that we will introduce more comfortable APIs later.)
# We can follow the operation of `dd()` by increasing the logging level. We see how with every test, the difference between the passing and the failing input gets smaller and smaller.
with DeltaDebugger(log=2) as dd:
mystery(failing_input)
dd.dd('inp', {'inp': failing_input})
# #### Processing Multiple Arguments
#
# What happens if a function has multiple arguments? First, we check if they are _reducible_ – that is, they provide a `len()` length function and a way to access indexed elements. This holds for all strings and all lists, as well as other ordered collections.
def is_reducible(value: Any) -> bool:
# Return True if `value` supports len() and indexing.
try:
_ = len(value)
except TypeError:
return False
try:
_ = value[0]
except TypeError:
return False
except IndexError:
return False
return True
# Our method `process_args()` processes recorded call arguments, one after the one, until all are minimized or maximized. Processing them individually (rather than, say, all at once) allows to maintain a stable _context_ during reduction.
# This method also does all the housekeeping, checking arguments and results, and raising errors if need be.
class FailureNotReproducedError(ValueError):
pass
class DeltaDebugger(DeltaDebugger):
def check_reproducibility(self) -> None:
# Check whether running the function again fails
assert self.function(), \
"No call collected. Use `with dd: func()` first."
assert self.args(), \
"No arguments collected. Use `with dd: func(args)` first."
self.reset()
outcome = self.test(self.args())
if outcome == UNRESOLVED:
raise FailureNotReproducedError(
"When called again, " +
self.format_call(self.args()) +
" raised " +
self.format_exception(self.last_exception) +
" instead of " +
self.format_exception(self.exception()))
if outcome == PASS:
raise NotFailingError("When called again, " +
self.format_call(self.args()) +
" did not fail")
assert outcome == FAIL
class DeltaDebugger(DeltaDebugger):
def process_args(self, strategy: Callable, **strategy_args: Any) -> \
Tuple[Dict[str, Any], Dict[str, Any], Dict[str, Any]]:
"""
Reduce all reducible arguments, using `strategy`(var, `strategy_args`).
Can be overloaded in subclasses.
"""
pass_args = {} # Local copy
fail_args = {} # Local copy
diff_args = {}
for var in self.args():
fail_args[var] = self.args()[var]
diff_args[var] = self.args()[var]
pass_args[var] = self.args()[var]
if is_reducible(pass_args[var]):
pass_args[var] = empty(pass_args[var])
vars_to_be_processed = set(fail_args.keys())
pass_processed = 0
fail_processed = 0
self.check_reproducibility()
# We take turns in processing variables until all are processed
while len(vars_to_be_processed) > 0:
for var in vars_to_be_processed:
if not is_reducible(fail_args[var]):
vars_to_be_processed.remove(var)
break
if self.log:
print(f"Processing {var}...")
maximized_pass_value, minimized_fail_value, diff = \
strategy(var, fail_args, **strategy_args)
if (maximized_pass_value is not None and
len(maximized_pass_value) > len(pass_args[var])):
pass_args[var] = maximized_pass_value
# FIXME: diff_args may not be correct for multiple args
diff_args[var] = diff
if self.log:
print(f"Maximized {var} to",
repr(maximized_pass_value))
vars_to_be_processed = set(fail_args.keys())
pass_processed += 1
if (minimized_fail_value is not None and
len(minimized_fail_value) < len(fail_args[var])):
fail_args[var] = minimized_fail_value
diff_args[var] = diff
if self.log:
print(f"Minimized {var} to",
repr(minimized_fail_value))
vars_to_be_processed = set(fail_args.keys())
fail_processed += 1
vars_to_be_processed.remove(var)
break
assert pass_processed == 0 or self.test(pass_args) == PASS, \
f"{self.format_call(pass_args)} does not pass"
assert fail_processed == 0 or self.test(fail_args) == FAIL, \
f"{self.format_call(fail_args)} does not fail"
if self.log and pass_processed > 0:
print("Maximized passing call to",
self.format_call(pass_args))
if self.log and fail_processed > 0:
print("Minimized failing call to",
self.format_call(fail_args))
return pass_args, fail_args, diff_args
# For more housekeeping, we define the `after_collection()` method that will be invoked at the end of the `with` block. It checks for a number of additional preconditions.
class DeltaDebugger(DeltaDebugger):
def after_collection(self) -> None:
# Some post-collection checks
if self._function is None:
raise NoCallError("No function call observed")
if self.exception() is None:
raise NotFailingError(
f"{self.format_call()} did not raise an exception")
if self.log:
print(f"Observed {self.format_call()}" +
f" raising {self.format_exception(self.exception())}")
# #### Public API
#
# We finish the implementation with public methods that allow users to run delta debugging and obtain the diagnostics.
class DeltaDebugger(DeltaDebugger):
def min_args(self) -> Dict[str, Any]:
"""Return 1-minimal arguments."""
pass_args, fail_args, diff = self.process_args(self.dd, mode='-')
return fail_args
class DeltaDebugger(DeltaDebugger):
def max_args(self) -> Dict[str, Any]:
"""Return 1-maximal arguments."""
pass_args, fail_args, diff = self.process_args(self.dd, mode='+')
return pass_args
class DeltaDebugger(DeltaDebugger):
def min_arg_diff(self) -> Tuple[Dict[str, Any], Dict[str, Any], Dict[str, Any]]:
"""Return 1-minimal difference between arguments."""
return self.process_args(self.dd, mode='+-')
# The `__repr__()` method returns a string representation of the minimized call.
class DeltaDebugger(DeltaDebugger):
def __repr__(self) -> str:
"""Return a string representation of the minimized call."""
return self.format_call(self.min_args())
# ### End of Excursion
# To see how the `DeltaDebugger` works, let us run it on our failing input. The expected usage is as introduced earlier – we wrap the failing function in a `with` block, and then print out the debugger to see the reduced arguments. We see that `DeltaDebugger` easily reduces the arguments to the minimal failure-inducing input:
with DeltaDebugger() as dd:
mystery(failing_input)
dd
# We can turn on logging for `DeltaDebugger` to see how it proceeds. With each step, we see how the remaining input gets smaller and smaller, until only two characters remain:
with DeltaDebugger(log=True) as dd:
mystery(failing_input)
dd
# It is also possible to access the debugger programmatically:
with DeltaDebugger() as dd:
mystery(failing_input)
dd.args()
dd.min_args()
quiz("What happens if the function under test does not raise an exception?",
[
"Delta debugging searches for the minimal input"
" that produces the same result",
"Delta debugging starts a fuzzer to find an exception",
"Delta debugging raises an exception"
"Delta debugging runs forever in a loop",
], '0 ** 0 + 1 ** 0 + 0 ** 1 + 1 ** 1')
# Indeed, `DeltaDebugger` checks if an exception occurs. If not, you obtain a `NotFailingError`.
with ExpectError(NotFailingError):
with DeltaDebugger() as dd:
mystery("An input that does not fail")
# Delta Debugging also assumes that the function under test is _deterministic_. If it occasionally fails and occasionally passes, you will get random results.
# ## Usage Examples
#
# Let us apply `DeltaDebugger` on a number of examples.
# ### Reducing remove_html_markup()
#
# For our ongoing `remove_html_markup()` example, we can reduce the failure-inducing input to a minimum, too:
from Assertions import remove_html_markup # minor dependency
with DeltaDebugger(log=True) as dd:
remove_html_markup('"x > y"')
dd.min_args()
# ### Reducing Multiple Arguments
#
# If a function has multiple reducible variables, they get reduced in turns. This `string_error()` function fails whenever `s1` is a substring of `s2`:
def string_error(s1: str, s2: str) -> None:
assert s1 not in s2, "no substrings"
# Running `DeltaDebugger` on `string_error` shows how first `s1` is reduced, then `s2`, then `s1` again.
# +
with DeltaDebugger(log=True) as dd:
string_error("foo", "foobar")
string_error_args = dd.min_args()
string_error_args
# -
# We see that the failure also occurs if both strings are empty:
with ExpectError(AssertionError):
string_error(string_error_args['s1'], string_error_args['s2'])
# ### Invoking an Interactive Debugger
#
# The results from delta debugging can be immediately used to invoke an nvoke an [interactive debugger](Debugger) on the minimized input. To this end, we need to turn the dictionary returned by `min_args()` into a arguments of the (failing) function call.
# Python provides a simple way to turn dictionaries into function calls. The construct
#
# ```python
# fun(**args)
# ```
#
# invokes the function `fun`, with all parameters assigned from the respective values in the dictionary.
# With this, we can immediately invoke a `Debugger` on the failing run with minimized arguments:
from Debugger import Debugger # minor dependency
# ignore
from bookutils import next_inputs
# ignore
next_inputs(['print', 'quit'])
with ExpectError(AssertionError):
with Debugger():
string_error(**string_error_args)
# ### Reducing other Collections
#
# Our `DeltaDebugger` is not limited to strings. It can reduce any argument `x` for which a `len(x)` operation and an indexing operation `x[i]` is defined – notably lists. Here is how to apply `DeltaDebugger` on a list:
def list_error(l1: List, l2: List, maxlen: int) -> None:
assert len(l1) < len(l2) < maxlen, "invalid string length"
with DeltaDebugger() as dd:
list_error(l1=[1, 2, 3, 4, 5, 6, 7, 8, 9, 10], l2=[1, 2, 3], maxlen=5)
dd
# ## Debugging Inputs
#
# Sometimes, it may be useful to not _minimize_ the input, but rather _maximize_ it – that is, to find the _maximum_ input that does _not_ fail. For instance, you may have an input of which you want to _preserve_ as much as possible – to repair it, or to establish a _context_ that is as close as possible to the real input.
# This is possible by using the `max_arg()` method. It implements the `ddmax` variant of the general Delta Debugging algorithm \cite{Kirschner2020}. With each step, it tries to add more and more characters to the passing input until it is _1-maximal_ – that is, any additional character that would be added from the failing input also would cause the function to fail.
with DeltaDebugger(log=True) as dd:
mystery(failing_input)
max_passing_input = dd.max_args()['inp']
max_passing_input
# Note that this is precisely the failure-inducing input _except_ for the first parenthesis. Adding this single character would cause the input to cause a failure.
# ## Failure-Inducing Differences
# If one wants to look for _differences_ that distinguish passing from failing runs, Delta Debugging also has a direct method for this – by both maximizing the passing input and minimizing the failing input until they meet somewhere in the middle. The remaining difference is what makes the difference between passing and failing.
# To compute the failure-inducing differences for `mystery()`, use the `min_arg_diff()` method:
with DeltaDebugger(log=True) as dd:
mystery(failing_input)
max_passing_args, min_failing_args, diff = dd.min_arg_diff()
max_passing_args['inp'], min_failing_args['inp'], diff['inp']
# Minimizing failure-inducing differences is especially efficient on large inputs, since the number of differences between a passing and a failing input is much smaller than the inputs themselves. Here is the failure-inducing difference as determined by Delta Debugging:
diff['inp']
# ## Reducing Program Code
#
# One particularly fun application of reducers is on _program code_. Technically speaking, program code is just another input to a computation; and we can actually automatically determine which minimum of program code is required to produce a failure, using Delta Debugging. Such minimization of code is typically used as it comes to debugging programs that accept code as their input, such as _compilers_ and _interpreters_. However, it can also pinpoint failure causes in the (input) code itself.
# As an example, let us apply Delta Debugging on the code from [the chapter on assertions](Assertions.html). You do not need to have read the chapter; the important part is that this chapter provides an implementation of `remove_html_markup()` that we want to use.
# ignore
try:
del remove_html_markup
except NameError:
pass
import Assertions # minor dependency
# Here is the source code of all the chapter; this is several hundred lines long.
import inspect
assertions_source_lines, _ = inspect.getsourcelines(Assertions)
# print_content("".join(assertions_source_lines), ".py")
assertions_source_lines[:10]
len(assertions_source_lines)
# We can take this code and execute it. Nothing particular should happen here, as our imports only import definitions of functions, classes, and global variables.
def compile_and_run(lines: List[str]) -> None:
# To execute 'Assertions' in place, we need to define __name__ and __package__
exec("".join(lines), {'__name__': '<string>',
'__package__': 'debuggingbook',
'Any': Any,
'Type': Type,
'TracebackType': TracebackType,
'Optional': Optional},
{})
compile_and_run(assertions_source_lines)
from Assertions import remove_html_markup # minor dependency
# Let us add some code to it – a "My Test" assertion that tests that `remove_html_markup()`, applied on a string with double quotes, should keep these in place:
def compile_and_test_html_markup_simple(lines: List[str]) -> None:
compile_and_run(lines +
[
'''''',
'''assert remove_html_markup('"foo"') == '"foo"', "My Test"\n'''
])
# This assertion fails. (As always, `remove_html_markup()` is buggy.)
with ExpectError(AssertionError):
compile_and_test_html_markup_simple(assertions_source_lines)
# The question we want to address in this section is: Given this assertion, can we automatically determine which part of the `Assertions` code lines in `assertions_source_lines` is relevant for producing the failure?
# ### Reducing Code Lines
#
# Since our `Assertions` source code comes as a list of lines, we can apply our `DeltaDebugger` on it. The result will be the list of source lines that is necessary to make the assertion fail.
quiz("What will the reduced set of lines contain?",
[
"All of the source code in the assertions chapter.",
"Only the source code of `remove_html_markup()`",
"Only a subset of `remove_html_markup()`",
"No lines at all."
], '[x for x in range((1 + 1) ** (1 + 1)) if x % (1 + 1) == 1][1]')
# Let us see what the `DeltaDebugger` produces.
with DeltaDebugger(log=False) as dd:
compile_and_test_html_markup_simple(assertions_source_lines)
# We get exactly _two_ lines of code:
reduced_lines = dd.min_args()['lines']
len(reduced_lines)
# And these are:
from bookutils import print_content
print_content("".join(reduced_lines), ".py")
# On these lines, our test actually still fails:
with ExpectError(AssertionError):
compile_and_test_html_markup_simple(reduced_lines)
# This failure may come as a surprise – `remove_html_markup()` is reduced to a function which does not even return a value. However, this is how it causes our "My Test" assertion to fail: In Python, a function without an explicit `return` statement returns `None`. This value is definitely not the string the "My Test" assertion expects, so it fails.
# At the same time, we also have a function `test_square_root()` which is equally devoid of any meaning – its code line does not even stem from its original implementation. Note, however, how the set of four lines is actually 1-minimal – removing any further line would result in a syntax error.
# To ensure we do not remove code that actually would be necessary for normal behavior, let us add another check – one that checks for the _normal_ functionality of `remove_html_markup()`. If this one fails (say, after the code has been tampered with too much), it raises an exception – but a _different_ one from the original failure:
def compile_and_test_html_markup(lines: List[str]) -> None:
compile_and_run(lines +
[
'',
'''if remove_html_markup('<foo>bar</foo>') != 'bar':\n''',
''' raise RuntimeError("Missing functionality")\n''',
'''assert remove_html_markup('"foo"') == '"foo"', "My Test"\n'''
])
# On our "reduced" code, we now obtain a different exception.
with ExpectError():
compile_and_test_html_markup(reduced_lines)
# Such an outcome that is different from the original failure causes our `DeltaDebugger` not treating this as a failure, but rather as a `UNRESOLVED` outcome, indicating that the test cannot determine whether it passed or failed. The `ddmin` algorithm treats such unresolved outcomes as if they were passing; hence, the algorithm treats its minimization attempt as unsuccessful.
# How does this change things? When we reduce the `Assertions` source code with the extended assertions, we now get a different result:
with DeltaDebugger(log=False) as dd:
compile_and_test_html_markup(assertions_source_lines)
reduced_assertions_source_lines = dd.min_args()['lines']
# Our result actually is the source code of `remove_html_markup()` – and _only_ the source code. This is a success, as Delta Debugging has eliminated all the other parts of the `Assertions` source code; these neither contribute to the correct functioning of `remove_html_markup()`, nor to the failure at hand.
print_content(''.join(reduced_assertions_source_lines), '.py')
# All in all, we have reduced the number of relevant lines in `Assertions` to about 3% of the original source code.
len(reduced_assertions_source_lines) / len(assertions_source_lines)
# The astute reader may notice that `remove_html_markup()`, as shown above, is slightly different from the original version in the [chapter on assertions](Assertions.ipynb). Here's the original version for comparison:
remove_html_markup_source_lines, _ = inspect.getsourcelines(Assertions.remove_html_markup)
print_content(''.join(remove_html_markup_source_lines), '.py')
quiz("In the reduced version, what has changed?",
[
"Comments are deleted",
"Blank lines are deleted",
"Initializations are deleted",
"The assertion is deleted",
], '[(1 ** 0 - -1 ** 0) ** n for n in range(0, 3)]')
# Indeed, Delta Debugging has determined all these as being irrelevant for reproducing the failure – and consequently, has deleted them.
# ### Reducing Code Characters
# We can reduce the code further by removing individual _characters_ rather than lines. To this end, we convert our (already reduced) `remove_html_markup()` code into a list of characters.
reduced_assertions_source_characters = list("".join(reduced_assertions_source_lines))
print(reduced_assertions_source_characters[:30])
# Our `compile_and_test_html_markup()` works (and fails) as before: It still joins the given strings into one and executes them. (Remember that in Python, "characters" are simply strings of length one.)
with ExpectError(AssertionError):
compile_and_test_html_markup(reduced_assertions_source_characters)
# Let's see what Delta Debugging makes of that – and also, how long it takes. The `Timer` class gives us a simple means to measure time.
from Timer import Timer
with DeltaDebugger(log=False) as dd:
compile_and_test_html_markup(reduced_assertions_source_characters)
# Here's the reduced result:
with Timer() as t:
further_reduced_assertions_source_characters = dd.min_args()['lines']
print_content("".join(further_reduced_assertions_source_characters), ".py")
# There's a number of observations we can make about this code.
#
# * All superfluous blanks and even newlines have been removed.
# * As a curiosity, the initialization of `quote` and `out` to `""` is now merged into a single (semantics-preserving) statement.
# * The semantics and effect of `<` and `>` characters is preserved, as mandated by our `RuntimeError` check.
# * Double quotes still have the effect of not being included in the returned value: the remaining `quote` has no effect.
#
# Semantics-wise, this reduced variant still yields the "original" failure; the biggest semantic differences, though, are in the condition and code associated with double quotes – which actually also is the location of the defect to be fixed. This is how reducing code can also point to not only _necessary_ locations, but also _defective_ locations.
# Mind you that reducing code is not cheap, and especially not if you remove by characters. It has taken `DeltaDebugger` several thousand tests to obtain the result above:
dd.tests
# And to do so, it even required _several seconds_. This may be little for a human, but from a CPU standpoint, this is an enormous effort.
t.elapsed_time()
# ### Reducing Syntax Trees
# When reducing code (or generally speaking, recursive structures), using a _syntactic_ approach can be a much better alternative to the _line-by-line_ or _character-by-character_ approaches discussed above. The idea is that one represents the input as a _tree_ (rather than a sequence of strings), in which a reducer would work on entire subtrees, deleting or reducing parts of the tree.
# We illustrate this concept on _syntax trees_ representing Python code. Python provides us with simple means to interactively convert code into syntax trees (and back again). So, in order to reduce code, we can
#
# 1. _parse_ the program code into a syntax tree (called *abstract syntax tree* or *AST*);
# 2. reduce the syntax tree to a minimum, executing it to test reductions; and
# 3. _unparse_ the tree to obtain textual code again.
#
# Since transformations on the AST are much less likely to produce syntax errors, reducing ASTs is much more efficient than reducing program code as text.
# In the [chapter on slicing](Slicer.ipynb), we already have seen several examples on how to work with ASTs. In our context, an AST also offers additional possibilities for reducing. Notably, instead of just _deleting_ code fragments, we can also _replace_ them with simpler fragments. For instance, we can replace arithmetic expressions with constants, or conditional statements `if cond: body` with the associated body `body`.
# Let us illustrate how this works, again choosing `remove_html_markup()` as our ongoing example. One more time, we create a function with associated test.
fun_source = inspect.getsource(remove_html_markup)
print_content(fun_source, '.py')
# #### From Code to Syntax Trees
#
# Let us parse this piece of code into an AST. This is done by the `ast.parse()` function.
import ast
fun_tree: ast.Module = ast.parse(fun_source)
# The parsed tree contains the function definition:
from bookutils import show_ast
show_ast(fun_tree)
# Let us add some tests to this, using the same scheme:
test_source = (
'''if remove_html_markup('<foo>bar</foo>') != 'bar':\n''' +
''' raise RuntimeError("Missing functionality")\n''' +
'''assert remove_html_markup('"foo"') == '"foo"', "My Test"'''
)
test_tree: ast.Module = ast.parse(test_source)
print_content(ast.unparse(test_tree), '.py')
# We can merge the function definition tree and the test tree into a single one:
import copy
fun_test_tree = copy.deepcopy(fun_tree)
fun_test_tree.body += test_tree.body
# Such a tree can be compiled into a code object, using Python's `compile()` function:
fun_test_code = compile(fun_test_tree, '<string>', 'exec')
# and the resulting code object can be executed directly, using the Python `exec()` function. We see that our test fails as expected.
with ExpectError(AssertionError):
exec(fun_test_code, {}, {})
# #### Traversing Syntax Trees
#
# Our goal is now to reduce this tree (or at least the subtree with the function definition) to a minimum.
# To this end, we manipulate the AST through the `ast` Python module. The [official Python `ast` reference](http://docs.python.org/3/library/ast) is complete, but a bit brief; the documentation ["Green Tree Snakes - the missing Python AST docs"](https://greentreesnakes.readthedocs.io/en/latest/) provides an excellent introduction.
# The two means for exploring and changing ASTs are the classes `NodeVisitor` and `NodeTransformer`, respectively. We start with creating a list of all nodes in the tree, using a `NodeVisitor` subclass.
#
# Its `visit()` method is called for every node in the tree, which we achieve by having it return `self.generic_visit()` for the current node. It saves all visited nodes in the `_all_nodes` attribute.
from ast import NodeTransformer, NodeVisitor, AST
class NodeCollector(NodeVisitor):
"""Collect all nodes in an AST."""
def __init__(self) -> None:
super().__init__()
self._all_nodes: List[AST] = []
def generic_visit(self, node: AST) -> None:
self._all_nodes.append(node)
return super().generic_visit(node)
def collect(self, tree: AST) -> List[AST]:
"""Return a list of all nodes in tree."""
self._all_nodes = []
self.visit(tree)
return self._all_nodes
# This is how our `NodeCollector()` class produces a list of all nodes:
fun_nodes = NodeCollector().collect(fun_tree)
len(fun_nodes)
fun_nodes[:30]
# Such a list of nodes is what we can feed into Delta Debugging in order to reduce it. The idea is that with every test, we take the tree and for each node in the tree, we check whether it is still in the list – if not, we remove it. Thus, by reducing the list of nodes, we simultaneously reduce the tree as well.
# #### Deleting Nodes
#
# In our next step, we write some code that, given such a list of nodes, _prunes_ the tree such that _only_ elements in the list are still contained. To this end, we proceed in four steps:
#
# 1. We traverse the original AST, _marking_ all nodes as "to be deleted".
# 2. We traverse the given list of nodes, clearing their markers.
# 3. We copy the original tree (including the markers) into a new tree – the one to be reduced.
# 4. We traverse the new tree, now deleting all marked nodes.
# Why do we go through such an extra effort? The reason is that our list of nodes contains references into the _original_ tree – a tree that needs to stay unchanged such that we can reuse it for later. The new tree (the copy) has the same nodes, but at different addresses, so our original references cannot be used anymore. Markers, however, just like any other attributes, are safely copied from the original into the new tree.
# The `NodeMarker()` visitor marks all nodes in a tree:
class NodeMarker(NodeVisitor):
def visit(self, node: AST) -> AST:
node.marked = True # type: ignore
return super().generic_visit(node)
# The `NodeReducer()` transformer reduces all marked nodes. If a method `visit_<node class>()` is defined, it will be invoked; otherwise, `visit_Node()` is invoked, which _deletes_ the node (and its subtree) by returning `None`.
class NodeReducer(NodeTransformer):
def visit(self, node: AST) -> Any:
method = 'visit_' + node.__class__.__name__
visitor = getattr(self, method, self.visit_Node)
return visitor(node)
def visit_Module(self, node: AST) -> Any:
# Can't remove modules
return super().generic_visit(node)
def visit_Node(self, node: AST) -> Any:
"""Default visitor for all nodes"""
if node.marked: # type: ignore
return None # delete it
return super().generic_visit(node)
# Our function `copy_and_reduce()` puts these pieces together:
def copy_and_reduce(tree: AST, keep_list: List[AST]) -> AST:
"""Copy tree, reducing all nodes that are not in keep_list."""
# Mark all nodes except those in keep_list
NodeMarker().visit(tree)
for node in keep_list:
# print("Clearing", node)
node.marked = False # type: ignore
# Copy tree and delete marked nodes
new_tree = copy.deepcopy(tree)
NodeReducer().visit(new_tree)
return new_tree
# Let us apply this in practice. We take the first assignment in our tree...
fun_nodes[4]
# ... whose subtree happens to be the assignment to `tag`:
ast.unparse(fun_nodes[4])
# We keep all nodes _except_ for this one.
keep_list = fun_nodes.copy()
del keep_list[4]
# Let us now create a copy of the tree in which the assignment is missing:
new_fun_tree = cast(ast.Module, copy_and_reduce(fun_tree, keep_list))
show_ast(new_fun_tree)
# The new tree no longer contains the initial assignment to `tag`:
print_content(ast.unparse(new_fun_tree), '.py')
# If we add our tests and then execute this code, we get an error, as `tag` is now no longer initialized:
new_fun_tree.body += test_tree.body
fun_code = compile(new_fun_tree, "<string>", 'exec')
with ExpectError(UnboundLocalError):
exec(fun_code, {}, {})
# If we have _no_ node in the keep list, the whole tree gets deleted:
empty_tree = copy_and_reduce(fun_tree, [])
ast.unparse(empty_tree)
# #### Reducing Trees
#
# We can put all these steps together in a single function. `compile_and_test_ast()` takes a tree and a list of nodes, reduces the tree to those nodes in the list, and then compiles and runs the reduced AST.
def compile_and_test_ast(tree: ast.Module, keep_list: List[AST],
test_tree: Optional[ast.Module] = None) -> None:
new_tree = cast(ast.Module, copy_and_reduce(tree, keep_list))
# print(ast.unparse(new_tree))
if test_tree is not None:
new_tree.body += test_tree.body
try:
code_object = compile(new_tree, '<string>', 'exec')
except Exception:
raise SyntaxError("Cannot compile")
exec(code_object, {}, {})
with ExpectError(AssertionError):
compile_and_test_ast(fun_tree, fun_nodes, test_tree)
# When we run our delta debugger on the AST, this is the list of remaining nodes we obtain:
with DeltaDebugger() as dd:
compile_and_test_ast(fun_tree, fun_nodes, test_tree)
reduced_nodes = dd.min_args()['keep_list']
len(reduced_nodes)
# This is the associated tree:
reduced_fun_tree = copy_and_reduce(fun_tree, reduced_nodes)
show_ast(reduced_fun_tree)
# And this is its textual representation:
print_content(ast.unparse(reduced_fun_tree), '.py')
dd.tests
# We see that some code was deleted – notably the assertion at the end – but otherwise, our deletion strategy was not particularly effective. This is because in Python, one cannot simply delete the single statement in a controlled body – this raises a syntax error. One would have to replace it with `pass` (or some other statement with no effect) to stay syntactically valid. Still, the syntax-based reduction would still single out `remove_html_markup()` from the `Assertions` source code – and do so even faster, as it would apply on one definition (rather than one line) after another.
# #### Transforming Nodes
#
# To further boost our syntactic reduction strategy, we implement a set of additional reduction operators. First, as already discussed, we do not simply delete an assignment, but we replace it with a `pass` statement. To obtain the tree for `pass`, we simply parse it and access the subtree.
class NodeReducer(NodeReducer):
PASS_TREE = ast.parse("pass").body[0]
def visit_Assign(self, node: ast.Assign) -> AST:
if node.marked: # type: ignore
# Replace by pass
return self.PASS_TREE
return super().generic_visit(node)
# In a similar vein, we can replace comparison operators with `False`:
class NodeReducer(NodeReducer):
FALSE_TREE = ast.parse("False").body[0].value # type: ignore
def visit_Compare(self, node: ast.Compare) -> AST:
if node.marked: # type: ignore
# Replace by False
return self.FALSE_TREE
return super().generic_visit(node)
# If we have a Boolean operator, we attempt to replace it with its left operand:
class NodeReducer(NodeReducer):
def visit_BoolOp(self, node: ast.BoolOp) -> AST:
if node.marked: # type: ignore
# Replace by left operator
return node.values[0]
return super().generic_visit(node)
# And if we find an `If` clause, we attempt to replace it by its body:
class NodeReducer(NodeReducer):
def visit_If(self, node: ast.If) -> Union[AST, List[ast.stmt]]:
if node.marked: # type: ignore
# Replace by body
return node.body
return super().generic_visit(node)
# Let us try to reduce our code with these additional reducers enabled:
with DeltaDebugger() as dd:
compile_and_test_ast(fun_tree, fun_nodes, test_tree)
# This is the reduced code we get. We see that all references to `quote` have gone, as has the handling of single quotes – none of this is relevant for the failure:
reduced_nodes = dd.min_args()['keep_list']
reduced_fun_tree = copy_and_reduce(fun_tree, reduced_nodes)
print_content(ast.unparse(reduced_fun_tree), '.py')
# Again, the best insights come from comparing this reduced version to the original implementation – and we learn that the problem is not related to the `quote` variable, or to the handling of single quotes; the problem is simply that when the input contains double quotes, these are not added to the final string.
# With our reduction code, however, we only touch the surface of what could actually be possible. So far, we implement exactly one reduction per node – but of course, there are many alternatives an expression or statement could be reduced to. We will explore some of these in the [exercises](#Exercises), below; also be sure to check out the [background](#Background) on code reduction.
# ## Synopsis
#
# A _reducer_ takes a failure-inducing input and reduces it to the minimum that still reproduces the failure. This chapter provides a `DeltaDebugger` class that implements such a reducer.
# Here is a simple example: An arithmetic expression causes an error in the Python interpreter:
def myeval(inp: str) -> Any:
return eval(inp)
with ExpectError(ZeroDivisionError):
myeval('1 + 2 * 3 / 0')
# Can we reduce this input to a minimum? _Delta Debugging_ is a simple and robust reduction algorithm. We provide a `DeltaDebugger` class that is used in conjunction with a (failing) function call:
#
# ```python
# with DeltaDebugger() as dd:
# fun(args...)
# dd
# ```
#
# The class automatically determines minimal arguments that cause the function to fail with the same exception as the original. Printing out the class object reveals the minimized call.
with DeltaDebugger() as dd:
myeval('1 + 2 * 3 / 0')
dd
# The input is reduced to the maximum: We get the essence of the division by zero.
# There also is an interface to access the reduced input(s) programmatically. The method `min_args()` returns a dictionary in which all function arguments are minimized:
dd.min_args()
# In contrast, `max_args()` returns a dictionary in which all function arguments are maximized, but still pass:
dd.max_args()
# The method `min_arg_diff()` returns a triple of
# * passing input,
# * failing input, and
# * their minimal failure-inducing difference:
dd.min_arg_diff()
# And you can also access the function itself, as well as its original arguments.
dd.function().__name__, dd.args()
# `DeltaDebugger` processes (i.e., minimizes or maximizes) all arguments that support a `len()` operation and that can be indexed – notably _strings_ and _lists_. If a function has multiple arguments, all arguments that can be processed will be processed.
# This chapter also provides a number of superclasses to `DeltaDebugger`, notably `CallCollector`, which obtains the first function call for `DeltaDebugger`. `CallReducer` classes allow for implementing alternate call reduction strategies.
# ignore
from ClassDiagram import display_class_hierarchy
# ignore
display_class_hierarchy([DeltaDebugger],
public_methods=[
StackInspector.caller_frame,
StackInspector.caller_function,
StackInspector.caller_globals,
StackInspector.caller_locals,
StackInspector.caller_location,
StackInspector.search_frame,
StackInspector.search_func,
StackInspector.is_internal_error,
StackInspector.our_frame,
CallCollector.__init__,
CallCollector.__enter__,
CallCollector.__exit__,
CallCollector.function,
CallCollector.args,
CallCollector.exception,
CallCollector.call,
CallReducer.__init__,
CallReducer.reduce_arg,
DeltaDebugger.dd,
DeltaDebugger.min_args,
DeltaDebugger.max_args,
DeltaDebugger.min_arg_diff,
DeltaDebugger.__repr__
],
project='debuggingbook')
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# * Reducing failure-inducing inputs to a minimum is helpful for testing and debugging.
# * _Delta debugging_ is a simple and robust algorithm to easily reduce inputs of test cases, as well as their code.
# * Precisely specifying failure conditions helps avoiding false diagnoses.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Next Steps
#
# Our next chapter focuses on [finding _failure-inducing code changes_](ChangeDebugger.ipynb), using delta debugging and version control systems.
# -
# ## Background
#
# The "lexical" delta debugging algorithm discussed here – both in its simplifying `ddmin` as well as in its general `dd` form – stem from \cite{Zeller2002}; actually, `ddmin` is the exact Python implementation as used by Zeller in 2002. The `ddmax` variant was first evaluated in \cite{Kirschner2020}. This chapter is the first to show how both `ddmin` and `ddmax` can be implemented as small variations of `dd`.
# The idea of systematically reducing inputs has been discovered a number of times, although not as automatic and generic as delta debugging. \cite{Slutz1998}, for instance, discusses systematic reduction of SQL statements for SQL databases; the general process as manual work is well described by \cite{Kernighan1999}.
# The deficits of delta debugging as it comes to syntactically complex inputs were first discussed in *compiler testing*, and _reducing tree inputs_ rather than string inputs was quickly discovered as an alternative. *Hierarchical Delta Debugging* (*HDD*) \cite{Misherghi2006} applies delta debugging on subtrees of a parse tree, systematically reducing a parse tree to a minimum. _Generalized Tree Reduction_ \cite{Herfert2017} generalizes this idea to apply arbitrary _patterns_ such as replacing a term by a compatible term in a subtree. Using _grammars_ to reduce inputs was first implemented in the _Perses_ tool \cite{Sun2018}. [A Python implementation of grammar-based input reduction](https://www.fuzzingbook.org/html/Reducer.html#Grammar-Based-Input-Reduction) is part of "The Fuzzing Book".
# While applying delta debugging to code lines does a decent job, _syntactic_ and especially _language-specific_ approaches can do a much better job for the programming language at hand:
#
# * *C-Reduce* \cite{Regehr2012} is a reducer specifically targeting the reduction of programming languages. Besides reductions in the style of delta debugging or tree transformations, C-Reduce comes with more than 30 source-to-source transformations that replace aggregates by scalars, remove function parameters at a definition and all call sites, change functions to return `void` and deleting all `return` statements, and many more. While specifically instantiated for the C language (and used for testing C compilers), these principles extend to arbitrary programming languages following an ALGOL-like syntax.
#
# * Kalhauge and Palsberg \cite{Kalhauge2019} introduce *binary reduction of dependency graphs*, a general solution for reducing arbitrary inputs with dependencies. Their *J-Reduce* tool specifically targets Java programs, and again is much faster than delta debugging and achieves a higher reduction rate.
# This [blog post](https://www.drmaciver.com/2019/01/notes-on-test-case-reduction/) by David McIver contains lots of insights on how to apply reduction in practice, in particular multiple runs with different abstraction levels.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Exercises
#
# How to best reduce inputs is still an underdeveloped field of research, with lots of opportunities.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 1: Advanced Syntactic Code Reduction
#
# Extend the code in ["Transforming Nodes"](#Transforming-Nodes) such that _multiple_ reduction possibilities for a node are being considered. For instance:
#
# * Replace a `BoolOp` node by `True`.
# * Replace a `BoolOp` node by `False`.
# * Replace a `BoolOp` node by its left operand.
# * Replace a `BoolOp` node by its right operand.
#
# or:
#
# * Replace an `If` node by its "then" body.
# * Replace an `If` node by its "else" body.
#
# or:
#
# * Replace all instances of a variable by a constant.
#
# or:
#
# * Replace expressions by a constant.
#
# Have a look at the [official Python `ast` reference](http://docs.python.org/3/library/ast) for a list of nodes (and some ideas on what to replace them by). The documentation ["Green Tree Snakes - the missing Python AST docs"](https://greentreesnakes.readthedocs.io/en/latest/) provides an excellent introduction on visitors and transformers. Make copious use of AST visualization and tests to ensure your syntax trees are still correct.
#
# Strategy-wise, you should first create a list of _possible_ reductions; and then pass to Delta Debugging a "keep list" of reductions that should _not_ be applied. When Delta Debugging minimizes this list, it will apply as many reductions as possible.
|
notebooks/DeltaDebugger.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("..")
import network as net
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# initialize hyper parameters
# our network will have 2 input neurons for input features or two inputs of a logic gate
# output layer will contain only one neuron to emulate logic gate's output
layer_dims = [2, 3, 1]
# length of activaiton_funcs should be [len(layer_dims) - 1] because the first layer is input layer and does not require computations
activation_funcs = ["tanh", "sigmoid"]
num_iterations = 20000
learning_rate = 0.075
# +
# initialize training data for or gate
X_train_orig = np.array([[0, 0],
[0, 1],
[1, 0],
[1, 1]])
# currently the shape of our training data is [4 x 2] i.e [m x n] so, we need to transpose it to make [n x m]
X_train = X_train_orig.T
Y_train_or_orig = np.array([[0],
[1],
[1],
[1]])
# transpose Y_train_or_orig to get [1 x m] shape
Y_train_or = Y_train_or_orig.T
# +
# instantiate the neural network object
dnn = net.DeepNeuralNetwork(layer_dims, activation_funcs, num_iterations, learning_rate)
# train the network
final_cost, costs = dnn.train_network(X_train, Y_train_or, cost_interval=100)
print("Final cost: " + str(final_cost))
print("Cost at iteration 0: " + str(costs[0]))
#print("costs: " + str(costs))
plt.plot(costs)
# -
# prediction time
X = np.array([[0],
[1]])
result = np.squeeze(dnn.predict(X))
print("result: " + str(result >= 0.5) + "\nactual output: " + str(result))
|
tests/or_gate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/01.png" width="1250" height="700" align="middle" alt="abstraction"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/02.png" width="1250" height="700" align="middle" alt="wli"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/03.jpg" width="700" height="500" align="middle" alt="nav"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/04.png" width="1000" height="700" align="middle" alt="backend_vs_frontend"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/05.png" width="700" height="500" align="middle" alt="lenguajes"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/06.jpg" width="800" height="700" align="middle" alt="jupyter"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/07.png" width="800" height="500" align="middle" alt="wil_joke"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/08.png" width="800" height="500" align="middle" alt="wil_joke"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/intro/09.png" width="800" height="500" align="middle" alt="wil_joke"/>
# + [markdown] slideshow={"slide_type": "slide"}
# ## <center> **Sistemas Operativos**
# <center><img src="img/SO/01_02.png" width="800" height="500" align="middle" alt="wil_joke"/>
# + [markdown] slideshow={"slide_type": "slide"}
# <center><img src="img/SO/04.jpg" width="400" height="500" align="middle" alt="wil_joke"/>
# -
# la terminal
# comandos del terminal
# como funciona el internet
# servidor cliente
# foto de mariposa
# como funciona el internet
# url
# servidores
# estandares de la web movil friendy
#
# no romper la web
#
# + [markdown] slideshow={"slide_type": "slide"}
# html code
# + [markdown] slideshow={"slide_type": "slide"}
# css
# + [markdown] slideshow={"slide_type": "slide"}
# js
# + [markdown] slideshow={"slide_type": "slide"}
# editores
# -
# estandates
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# arquitectura statica
# -
# arquitectura dinamica
#
#
#
#
#
#
|
.ipynb_checkpoints/slides-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gitFloyd/AAI-Project-3/blob/main/Pistachio_Detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="rWuCR4VnjYwv" outputId="a377947a-d86d-4bd5-a07b-05d824b12768"
# !pip install split-folders
# + id="ds1TP2uvjdUQ"
import pandas as pd
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import pathlib
import random
import os
import splitfolders
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.preprocessing import image_dataset_from_directory
from tensorflow.keras.layers.experimental import preprocessing
from tensorflow.keras.applications.vgg16 import VGG16
from tensorflow.keras.optimizers import SGD, RMSprop, Adam, Adagrad, Adadelta
from keras.layers import Dense, Dropout, Activation, Flatten, BatchNormalization, Conv2D, MaxPool2D, MaxPooling2D
from tensorflow.keras.callbacks import ReduceLROnPlateau, ModelCheckpoint, EarlyStopping
from tensorflow.keras import Model
from tensorflow.keras.layers import Dense, GlobalAveragePooling2D
# + id="IbfpSPQljfGA"
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '2'
# + colab={"base_uri": "https://localhost:8080/"} id="HgkTqVwvNaLq" outputId="b412c327-7993-4892-d199-223e275cc0b6"
from google.colab import drive
drive.mount('/content/drive')
# + id="sMmqal8WNryD"
data_dir ='/content/drive/MyDrive/Colab-Notebooks/Pistachio_Image_Dataset'
# + colab={"base_uri": "https://localhost:8080/"} id="QY4FliS5j0ro" outputId="bff2c40a-ce2d-4250-d16e-18e06cf6c754"
splitfolders.ratio(data_dir, output="output", seed=101, ratio=(.8, .1, .1))
# + id="7lD9D7kXkBag"
train_path = './output/train'
test_path = './output/test'
val_path = './output/val'
# + id="rucuHVsTkEmS"
img = mpimg.imread('./output/train/Kirmizi_Pistachio/kirmizi 62.jpg')
img1 = mpimg.imread('./output/train/Siirt_Pistachio/siirt 603.jpg')
# + colab={"base_uri": "https://localhost:8080/"} id="qhQNKaNJkJsQ" outputId="55ebb7f9-68b2-4be8-fc39-a870442fca5f"
img.shape, img1.shape
# + id="Bi41MX1EkOBS"
img_size = 512
batch = 32
# + id="IGjbNlpYkUzo"
labels = []
for i in os.listdir(train_path):
labels+=[i]
# + colab={"base_uri": "https://localhost:8080/"} id="NEg2v5N8kZy7" outputId="9d03848c-22d1-4c46-dda0-fef7860476d9"
labels
# + id="oMYqShbFlDQP"
def load_random_imgs_from_folder(folder,label):
plt.figure(figsize=(15,15))
for i in range(3):
file = random.choice(os.listdir(folder))
image_path = os.path.join(folder, file)
img=mpimg.imread(image_path)
ax=plt.subplot(1,3,i+1)
ax.title.set_text(label)
plt.xlabel(f'Name: {file}')
plt.imshow(img)
# + colab={"base_uri": "https://localhost:8080/", "height": 631} id="75XbntNInD8N" outputId="65f4a5a4-40a0-47ea-eb54-1cac1235c633"
for label in labels:
load_random_imgs_from_folder(f"{data_dir}/{label}",label)
# + id="hm6nBvrAnWRj"
train_datagen = ImageDataGenerator(rescale = 1.0/255.0,
rotation_range = 0.5,
width_shift_range = 0.2,
height_shift_range = 0.2,
shear_range = 0.2,
zoom_range = 0.1,
horizontal_flip = True,
fill_mode = 'nearest'
)
test_val_datagen = ImageDataGenerator(rescale = 1.0/255.0)
# + colab={"base_uri": "https://localhost:8080/"} id="V5RkkcBVSO1F" outputId="bad3d78a-1912-4cc8-abdd-cef45b105132"
train_generator = train_datagen.flow_from_directory(directory = train_path,
batch_size = batch,
class_mode = "categorical",
target_size = (img_size,img_size)
)
val_generator = test_val_datagen.flow_from_directory(directory = val_path,
batch_size = batch,
class_mode = "categorical",
target_size = (img_size,img_size)
)
test_generator = test_val_datagen.flow_from_directory(directory = test_path,
batch_size = batch,
class_mode = "categorical",
target_size = (img_size,img_size)
)
# + colab={"base_uri": "https://localhost:8080/"} id="qqxHYDguSbS1" outputId="ac834cfd-ad5e-46b0-d7c3-beb6c158ae1f"
data_train = image_dataset_from_directory(
data_dir,
label_mode='categorical',
validation_split=0.2,
subset="training",
seed=0,
color_mode="rgb",
image_size=(img_size,img_size),
batch_size=32,
)
data_test = image_dataset_from_directory(
data_dir,
validation_split=0.2,
subset="validation",
label_mode='categorical',
seed=0,
color_mode="rgb",
image_size=(img_size,img_size),
batch_size=32,
)
# + id="BP3guShgSfyI"
train_data_small = data_train.take(5)
# + colab={"base_uri": "https://localhost:8080/"} id="QNA1YY3ySl-0" outputId="803cc533-3430-40cd-f4d0-a62a351a5b38"
base_model = VGG16(weights='imagenet', include_top=False,
input_shape=(img_size, img_size,3))
# freeze extraction layers
base_model.trainable = False
# add custom top layers
x = base_model.output
x = GlobalAveragePooling2D()(x)
x = Dropout(0.2)(x)
x = Dense(4096,activation="relu")(x)
x = Dense(4096,activation="relu")(x)
x = Dropout(0.2)(x)
x = Dense(2096,activation="relu")(x)
predictions = Dense(2, activation='sigmoid')(x)
model = Model(inputs=base_model.input, outputs=predictions)
# confirm unfrozen layers
for layer in model.layers:
if layer.trainable==True:
print(layer)
# + colab={"base_uri": "https://localhost:8080/"} id="B1TYLWj_Sw9y" outputId="ce029e1d-dbad-4315-9202-31f030ace1ed"
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="jQPrOqC1S1KX" outputId="d17caa02-c168-44b1-8c8a-5ad3016fd8c8"
from tensorflow.keras.utils import plot_model
from IPython.display import Image
plot_model(model, to_file='convnet.png', show_shapes=True,show_layer_names=True)
Image(filename='convnet.png')
# + id="H9ygan8SS69-"
callbacks = [EarlyStopping(monitor='val_loss', patience=5, verbose=1),
ModelCheckpoint('model.hdf5',
save_best_only=True)]
# + id="P2dnmsomTAfd"
opt = Adam(learning_rate=0.005)
model.compile(
loss='categorical_crossentropy',
optimizer=opt,
metrics=['accuracy']
)
# + colab={"background_save": true, "base_uri": "https://localhost:8080/"} id="b3E4NRotTDuU" outputId="f97b3768-97f9-42c9-98c1-016f34a3a347"
#history=model.fit(data_train,
#epochs=50,
#validation_data=data_test,
#validation_steps=int(0.1 * len(data_test)),
#verbose=1,
#callbacks=callbacks)
|
Pistachio_Detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %%writefile load_test.py
import molotov
@molotov.scenario(100)
async def scenario_one(session):
async with session.get("http://localhost:5000") as resp:
assert resp.status == 200
# +
# %%writefile small.py
from flask import Flask, redirect, request
app = Flask('basic app')
@app.route('/', methods=['GET', 'POST'])
def index():
if request.method == 'POST':
redirect('https://www.google.com/search?q=%s' % request.args['q'])
else:
return '<h1>GET request from Flask!</h1>'
# +
# FLASK_APP=small.py flask run
# molotov -v -r 100 load_test.py
# -
|
python_utils/load_testing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch1_0
# language: python
# name: pytorch1_0
# ---
# %matplotlib inline
# +
import numpy as np
import pylab
from scipy.optimize import curve_fit
from numba import njit
from scipy.misc import derivative
np.random.seed(17)
# -
def plot_f(x, funcs):
if not isinstance(funcs, list):
funcs = [funcs]
for func in funcs:
pylab.plot(x, func(x), label=func.__name__)
pylab.legend(loc='upper left')
pylab.grid(True)
pylab.show()
def get_act(func, popt):
def f(x):
return func(x, *popt)
return f
# +
def softplus(x):
return np.log(1+np.exp(x))
def relu(x):
return np.clip(x, a_min=0, a_max=None)
def relu6(x):
return np.clip(x, a_min=0, a_max=6)
def leakyrelu(x):
res = np.array(x)
neg_x_idx = x < 0
res[neg_x_idx] = 0.01*x[neg_x_idx]
return res
def get_leaky_relu(alpha):
def LR(x):
res = np.array(x)
neg_x_idx = x < 0
res[neg_x_idx] = alpha*x[neg_x_idx]
return res
LR.alpha = alpha
return LR
def elu(x, alpha=1.0):
res = np.array(x)
neg_x_idx = x <= 0
x = x[neg_x_idx]
res[neg_x_idx] = alpha*(np.exp(x)-1)
return res
def celu(x, alpha=1.0):
res = np.array(x)
neg_x_idx = x < 0
x = x[neg_x_idx]
res[neg_x_idx] = alpha*(np.exp(x/alpha)-1)
return res
def selu(x, alpha=1.6732632423543772848170429916717, scale=1.0507009873554804934193349852946):
res = np.array(x)
neg_x_idx = x < 0
x = x[neg_x_idx]
res[neg_x_idx] = alpha*(np.exp(x)-1)
return scale*res
def tanh(x):
return np.tanh(x)
def sigmoid(x):
return 1.0 / (1.0 + np.exp(-x))
def swish(x):
return x * (1.0 / (1.0 + np.exp(-x)))
# +
@njit
def ratio_func54(x, w0,w1,w2,w3,w4, w5, d1, d2, d3, d4):
c1 = 0
xp = (x-c1)
xp1 = xp
xp2 = xp1*xp
xp3 = xp2*xp
xp4 = xp3*xp
xp5 = xp4*xp
P = w0 + w1*xp1 + w2*xp2 + w3*xp3 + w4*xp4 + w5*xp5
Q = 1.0 + d1*xp1 + d2*xp2 + d3*xp3 + d4*xp4
return P/Q
@njit
def ratio_func_abs54(x, w0,w1,w2,w3,w4, w5, d1, d2, d3, d4):
c1 = 0
xp = (x-c1)
xp1 = xp
xp2 = xp1*xp
xp3 = xp2*xp
xp4 = xp3*xp
xp5 = xp4*xp
P = w0 + w1*xp1 + w2*xp2 + w3*xp3 + w4*xp4 + w5*xp5
Q = 1.0 + np.abs(d1)* np.abs(xp1) + np.abs(d2)* np.abs(xp2) + np.abs(d3)* np.abs(xp3) + np.abs(d4)* np.abs(xp4)
return P/Q
# -
def fit_func(func, ref_func, x, p0=None, maxfev=10000000, bounds=None):
y = ref_func(x)
popt, _ = curve_fit(func, x, y, p0=p0, maxfev=maxfev, bounds=bounds)
#print(popt)
return popt, get_act(func, popt)
lr000 = get_leaky_relu(0.0)
lr001 = get_leaky_relu(0.01)
lr025 = get_leaky_relu(0.25)
lr030 = get_leaky_relu(0.30)
lr020 = get_leaky_relu(0.20)
lrm050 = get_leaky_relu(-0.50)
# +
x = np.arange(-3,3,0.000001)
result = []
for lrf in [lr000]:
popt, act_f = fit_func(ratio_func_abs54, lrf, x, bounds=(-np.inf, np.inf))
print(lrf.alpha, popt.tolist())
result.append([popt, act_f])
plot_f(np.arange(-5,5,0.00001), [act_f, lrf])
# +
x = np.arange(-10,10,0.000001)
popt, act_f = fit_func(ratio_func_abs54, relu6, x, bounds=(-np.inf, np.inf))
print('relu6', popt.tolist())
plot_f(np.arange(-7,7,0.00001), [act_f, relu6])
# -
popt_sigmoid = [1/2, 1/4, 1/18, 1/144, 1/2016, 1/60480, 0, 1/9, 0, 1/1000]
popt_tanh = [1/2, 1/4, 1/18, 1/144, 1/2016, 1/60480, 0, 1/9, 0, 1/1000]
popt_swish = [1/2, 1/4, 1/18, 1/144, 1/2016, 1/60480, 0, 1/9, 0, 1/1000]
popt_lrelu0_01 = [0.02979246288832245, 0.6183773789612337, 2.3233520651936534, 3.0520265972657823, 1.4854800152744463, 0.251037168372827, -1.1420122633346115, 4.393228341365807, 0.8715444974667658, 0.34720651643419215]
popt_lrelu0_20 = [0.025577756009581332, 0.6618281545012629, 1.5818297539580468, 2.944787587381909, 0.9528779431354413, 0.23319680694163697, -0.5096260509947604, 4.183768902183391, 0.3783209020348012, 0.3240731442906416]
popt_lrelu0_25 = [0.02423485464722387, 0.6770971779085044, 1.4385836314706064, 2.9549799006291724, 0.8567972159918334, 0.2322961171003388, -0.41014745814143555, 4.1469196374300115, 0.3029254642283438, 0.32002849530519256]
popt_lrelu0_30 = [0.022823661027641513, 0.6935843817924783, 1.308474321805162, 2.976815988084191, 0.7716529650279255, 0.23252265245280854, -0.3284954321510746, 4.115579017543179, 0.2415560267417864, 0.31659365394646605]
popt_lrelu0_50_neg =[0.026504409606513814, 0.8077291240826262, 13.566116392373088, 7.002178997009714, 11.614777812309141, 0.6872037476855452, -13.706489934094302, 6.077817327962073, 12.325352286416361, -0.540068802253311]
popt = [0.022823661027641513, 0.6935843817924783, 1.308474321805162, 2.976815988084191, 0.7716529650279255, 0.23252265245280854, -0.3284954321510746, 4.115579017543179, 0.2415560267417864, 0.31659365394646605]
act_f = get_act(ratio_func_abs54, popt_sigmoid)
plot_f(np.arange(-5,5,0.00001), [act_f])
|
activations/pau/find_coefficients.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# Content provided under a Creative Commons Attribution license, CC-BY 4.0.
# (c) 2014 L.Barba, P.Bardet, A.Wickenheiser (The George Washington University).
# Thanks: <NAME>, <NAME>, <NAME>, NSF for support via CAREER award #1149784 to LAB and GW Office and Teaching and Learning for seed grant to LAB and AMW.
# -
# ##### Version 0.1 -- February 2014
# # JITcode 1, lesson 1
# This is lesson 1 of the first *Just-in-Time (JIT) module* for teaching computing to engineers, in context. The first module lays the foundations for building computational skills. It is not meant to support a particular engineering course, so it can be used by freshman students. The context problems should be interesting to any science-minded student.
#
# Lesson 1 builds competency on these basic skills:
#
# * reading data from a file in comma-separated format (CSV)
# * plotting data
# * analyzing data with statistics
# * writing an image of a plot to a file
#
# ## Context — Earth temperature over time
# Is global temperature rising? How much? This is a question of burning importance in today's world!
#
# Data about global temperatures are available from several sources: NASA, the National Climatic Data Center (NCDC) and the University of East Anglia in the UK. Check out the [University Corporation for Atmospheric Research](https://www2.ucar.edu/climate/faq/how-much-has-global-temperature-risen-last-100-years) (UCAR) for an in-depth discussion.
#
# The [NASA Goddard Space Flight Center](http://svs.gsfc.nasa.gov/goto?3901) is one of our sources of global climate data. They produced this video showing a color map of the changing global surface **temperature anomalies** from 1880 to 2011.
#
# The term [_global temperature anomaly_](https://www.ncdc.noaa.gov/monitoring-references/faq/anomalies.php) means the difference in temperature with respect to a reference value or a long-term average. It is a very useful way of looking at the problem and in many ways better than absolute temperature. For example, a winter month may be colder than average in Washington DC, and also in Miami, but the absolute temperatures will be different in both places.
from IPython.display import YouTubeVideo
YouTubeVideo('lyb4gau3LyI')
# How would we go about understanding the _trends_ from the data on global temperature?
#
# The first step in analyzing unknown data is to generate some simple plots. We are going to look at the temperature-anomaly history, contained in a file, and make our first plot to explore this data.
#
# We are going to smooth the data and then we'll fit a line to it to find a trend, plotting along the way to see how it all looks.
#
# Let's get started!
#
# The first thing to do is to load our favorite library: the **NumPy** library for array operations.
import numpy
# Make sure you have studied the introduction to [_JITcode_ in Python](http://nbviewer.ipython.org/github/barbagroup/JITcode-MechE/blob/master/lessons/00_Lesson00_QuickPythonIntro.ipynb) to know a bit about this library and why we need it.
# ## Step 1: Read a data file
# The data is contained in the file:
#
# `GlobalTemperatureAnomaly-1958-2008.csv`
#
# with the year on the first column and 12 monthly averages of temperature anomaly listed sequentially on the second column. We will read the file, then make an initial plot to see what it looks like.
#
# To load the file, we use a function from the NumPy library called `loadtxt()`. To tell Python where to look for this function, we precede the function name with the library name, and use a dot between the two names. This is how it works:
numpy.loadtxt(fname='./resources/GlobalTemperatureAnomaly-1958-2008.csv', delimiter=',')
# Note that we called the function with two parameters: the file name and path, and the delimiter that separates each value on a line (a comma). Both parameters are strings (made up of characters) and we put them in single quotes.
#
# As the output of the function, we get an array. Because it's rather big, Python shows only a few rows and columns of the array.
#
# So far, so good. Now, what if we want to manipulate this data? Or plot it? We need to refer to it with a name. We've only just read the file, but we did not assign the array any name! Let's try again.
T=numpy.loadtxt(fname='./resources/GlobalTemperatureAnomaly-1958-2008.csv', delimiter=',')
# That's interesting. Now, we don't see any output from the function call. Why? It's simply that the output was stored into the variable `T`, so to see it, we can do:
print(T)
# Ah, there it is! Let's find out how big the array is. For that, we use a cool NumPy function called `shape()`:
numpy.shape(T)
# Again, we've told Python where to find the function shape() by attaching it to the library name with a dot. However, NumPy arrays also happen to have a property shape that will return the same value, so we can get the same result another way:
T.shape
# It's just shorter. The array `T` holding our temperature-anomaly data has two columns and 612 rows. Since we said we had monthly data, how many years is that?
612/12
# That's right: from 1958 through 2008.
# ## Step 2: Plot the data
# We will display the data in two ways: as a time series of the monthly temperature anomalies versus time, and as a histogram. To be fancy, we'll put both plots in one figure.
#
# Let's first load our plotting library, called `matplotlib`. To get the plots inside the notebook (rather than as popups), we use a special command, `%matplotlib inline`:
from matplotlib import pyplot
# %matplotlib inline
# What's this `from` business about? `matplotlib` is a pretty big (and awesome!) library. All that we need is a subset of the library for creating 2D plots, so we ask for the `pyplot` module of the `matplotlib` library.
#
# Plotting the time series of temperature is as easy as calling the function [`plot()`](http://matplotlib.org/1.5.1/api/pyplot_api.html#matplotlib.pyplot.plot) from the module `pyplot`.
#
# But remember the shape of `T`? It has two columns and the temperature-anomaly values are in the second column. We extract the values of the second column by specifying 1 as the second index (the first column has index 0) and using the colon notation `:` to mean *all rows*. Check it out:
pyplot.plot(T[:,1])
# You can add a semicolon at the end of the plotting command to avoid that stuff that appeared on top of the figure, that `Out[x]: [< ...>]` ugliness. Try it.
#
# *Do you see a trend in the data?*
#
# The plot above is certainly useful, but wouldn't it be nicer if we could look at the data relative to the year, instead of the location of the data in the array?
#
# The plot function can take another input; let's get the year displayed as well.
pyplot.plot(T[:,0],T[:,1]);
# The temperature anomaly certainly seems to show an increasing trend. But we're not going to stop there, of course. It's not that easy to convince people that the planet is warming, as you know.
#
# Plotting a histogram is as easy as calling the function `hist()`. Why should it be any harder?
pyplot.hist(T[:,1]);
# *What does this plot tell you about the data?* It's more interesting than just an increasing trend, that's for sure. You might want to look at more statistics now: mean, median, standard deviation ... NumPy makes that easy for you:
meanT = numpy.mean(T[:,1])
medianT = numpy.median(T[:,1])
print( meanT, medianT)
# You can control several parameters of the [`hist()`](http://matplotlib.org/1.3.1/api/pyplot_api.html?highlight=hist#matplotlib.pyplot.hist) plot. Learn more by reading the manual page (yes, you have to read the manual sometimes!). The first option is the number of bins—the default is 10—but you can also change the appearance (color, transparency). Try some things out.
pyplot.hist(T[:,1], 20, normed=1, facecolor='g', alpha=0.55);
# This is fun. Finally, we'll put both plots on the same figure using the [`subplot()`](http://matplotlib.org/api/pyplot_api.html?highlight=subplot#matplotlib.pyplot.subplot) function, which creates a grid of plots. The argument tells this function how many rows and columns of sub-plots we want, and where in the grid each plot will go.
#
# To help you see what each plotting command is doing, we added comments, which in Python follow the `#` symbol.
pyplot.figure(figsize=(12,4)) # the size of the figure area
pyplot.subplot(121) # creates a grid of 1 row, 2 columns and selects the first plot
pyplot.plot(T[:,0],T[:,1],'g') # our time series, but now green
pyplot.xlim(1958,2008) # set the x-axis limits
pyplot.subplot(122) # prepares for the second plot
pyplot.hist(T[:,1], 20, normed=1, facecolor='g', alpha=0.55);
# ## Step 3: Smooth the data and do regression
# You see a lot of fluctuations on the time series, so you might be asking yourself "How can I smooth it out?" No? Let's do it anyway.
#
# One possible approach to smooth the data (there are others) is using a *moving average*, also known as a sliding-window average. This is defined as:
#
# $$\hat{x}_{i,n} = \frac{1}{n} \sum_{j=1}^{n} x_{i-j}$$
#
# The only parameter to the moving average is the value $n$. As you can see, the moving average smooths the set of data points by creating a new data set consisting of local averages (of the $n$ previous data points) at each point in the new set.
#
# A moving average is technically a _convolution_, and luckily NumPy has a built-in function for that, `convolve()`. We use it like this:
#
N = 12
window = numpy.ones(N)/N
smooth = numpy.convolve(T[:,1], window, 'same')
pyplot.figure(figsize=(10, 4))
pyplot.plot(T[:,0], smooth, 'r')
pyplot.xlim(1958,2008);
# Did you notice the function [`ones()`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.ones.html)? It creates an array filled with ... you guessed it: ones!
#
# We use a _window_ of 12 data points, meaning that the plot shows the average temperature over the last 12 months. Looking at the plot, we can still see a trend, but the range of values is smaller. Let's plot the original time series together with the smoothed version:
pyplot.figure(figsize=(10, 4))
pyplot.plot(T[:,0], T[:,1], 'g', linewidth=1) # we specify the line width here ...
pyplot.plot(T[:,0], smooth, 'r', linewidth=2) # making the smoothed data a thicker line
pyplot.xlim(1958, 2008);
# That is interesting! The smoothed data follows the trend nicely but has much less noise. Well, that is what filtering data is all about.
#
# Let's now fit a straight line through the temperature-anomaly data, to see the trends. We need to perform a least-squares linear regression to find the slope and intercept of a line
#
# $$y = mx+b$$
#
# that fits our data. Thankfully, Python and NumPy are here to help with the `polyfit()` function. The function takes three arguments: the two array variables $x$ and $y$, and the order of the polynomial for the fit (in this case, 1 for linear regression).
#
year = T[:,0] # it's time to use a more friendly name for column 1 of our data
m,b = numpy.polyfit(year, T[:,1], 1)
pyplot.figure(figsize=(10, 4))
pyplot.plot(year, T[:,1], 'g', linewidth=1)
pyplot.plot(year, m * year + b, 'k--', linewidth=2)
pyplot.xlim(1958, 2008);
# There is more than one way to do this. Another of the favorite Python libraries is **SciPy**, and it has a `linregress(x,y)` function that will work as well. But let's not get carried away.
# ## Step 4: Checking for auto-correlation in the data
# We won't go into details, but you will learn more about all this if you take a course on experimental methods—for example, at GW, the Mechanical and Aerospace Engineering department offers _"Methods of Engineering Experimentation"_ (MAE-3120).
#
# The fact is that in **time series** (like global temperature anomaly, stock values, etc.), the fluctuations in the data are not random: adjacent data points are not independent. We say that there is _auto-correlation_ in the data.
#
# The problem with auto-correlation is that various techniques in statistical analysis rely on the assumption that scatter (or error) is random. If you apply these techniques willy-nilly, you can get false trends, overestimate uncertainties or exaggerate the goodness of a fit. All bad things!
#
# For the global temperature anomaly, this discussion is crucial: _many critics claim that since there is auto-correlation in the data, no reliable trends can be obtained_
#
# As a well-educated engineering student who cares about the planet, you will appreciate this: we _can_ estimate the trend for the global temperature anomalies taking into account that the data points are not independent. We just need to use more advanced techniques of data analysis.
#
# To finish off this lesson, your first in data analysis with Python, we'll put all our nice plots in one figure frame, and add the _residual_. Because the residual is not random "white" noise, you can conclude that there is auto-correlation in this time series.
#
# Finally, we'll save the plot to an image file using the `savefig()` command of Pyplot—this will be useful to you when you have to prepare reports for your engineering courses!
pyplot.figure(figsize=(10, 8)) # the size of the figure area
pyplot.subplot(311) # creates a grid of 3 columns, 1 row and place the first plot
pyplot.plot(year, T[:,1], 'g', linewidth=1) # we specify the line width here ...
pyplot.plot(year, smooth, 'r', linewidth=2) # making the smoothed data a thicker line
pyplot.xlim(1958, 2008)
pyplot.subplot(312)
pyplot.plot(year, T[:,1], 'g', linewidth=1)
pyplot.plot(year, m * year + b, 'k--', linewidth=2)
pyplot.xlim(1958, 2008)
pyplot.subplot(313)
pyplot.plot(year, T[:,1] - m * year + b, 'o', linewidth=2)
pyplot.xlim(1958, 2008)
pyplot.savefig("TemperatureAnomaly.png")
# ## Step 5: Generating useful output
# Here, we'll use our linear fit to project the temperature into the future. We'll also save some image files that we could later add to a document or report based on our findings. First, let's create an expectation of the temperature difference up to the year 2100.
#
spacing = (2008 + 11 / 12 - 1958) / 612
length = (2100 - 1958) / spacing
length = int(length) #we'll need an integer for the length of our array
years = numpy.linspace(1958, 2100, num = length)
temp = m * years + b#use our linear regression to estimate future temperature change
pyplot.figure(figsize=(10, 4))
pyplot.plot(years, temp)
pyplot.xlim(1958, 2100)
out=(years, temp) #create a tuple out of years and temperature we can output
out = numpy.array(out).T #form an array and transpose it
# Ok, that estimation looks reasonable. Let's save the data that describes it back to a .csv file, like the one we originally imported.
#
numpy.savetxt('./resources/GlobalTemperatureEstimate-1958-2100.csv', out, delimiter=",")
# Now, lets make a nicer picture that we can show to back up some of our information. We can plot the linear regression as well as the original data and then save the figure.
#
pyplot.figure(figsize = (10, 4))
pyplot.plot(year, T[:,1], 'g')
pyplot.plot(years, temp, 'k--')
pyplot.xlim(1958, 2100)
pyplot.savefig('./resources/GlobalTempPlot.png')
# Nice! Now we've got some stuff that we could use in a report, or show to someone unfamiliar with coding. Remember to play with our settings; I'm sure you could get an even nicer-looking plot if you try!
# ##### Dig Deeper & Think
# 1. How is the global temperature anomaly calculated?
# 2. What does it mean and why is it employed instead of the global mean temperature to quantify global warming?
# 3. Why is it important to check that the residuals are independent and random when performing linear regression?
# 4. In this particular case, is it possible to still estimate a trend with confidence?
# 5. What is your best estimate of the global temperature by the end of the 22nd century?
# ##### What did we learn?
# You should have played around with the embedded code in this notebook, and also written your own version of all the code in a separate Python script to learn:
#
# * how to read data from a comma-separated file
# * how to plot the data
# * how to do some basic analysis on the data
# * how to write to a file
# ---
# + active=""
# Please ignore the cell bellow. It simply loads a style to make this notebook look pretty.
# -
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
|
module00_Introduction_to_Python/01_Lesson01_Playing_with_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pyefiuna/Laboratorio/blob/master/notebooks/Distribuciones%20de%20Probabilidad/Distribucion_Binomial.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DlqMfhq3BwGV"
# 
# # PROBABILIDAD Y ESTADÍSTICA
# ---
#
# + [markdown] id="bX1rIDKUCign"
# ## Distribuciones de probabilidad
# ### Distribución Binomial
# + [markdown] id="l5nAlRx6GPjU"
# ### Materiales y recursos
# ##### Teoría:
# * [Distribución Binomial](https://drive.google.com/file/d/1nsSupN3fb80MjWT8yokAAgJBLtAmdk40/view?ts=5f19b9dc)
#
# ##### Actividades de laboratorio
# * [Vídeos de ejercicios de distribución binomial resueltos (en excel)](https://www.youtube.com/playlist?list=PLwx_FbswjRXy0PUH06vFIl3YJpy5daNqC)
# * [Enunciados de ejercicios de distribución binomial](https://eaula.ing.una.py/pluginfile.php/76573/mod_resource/content/1/5.%20Distribuci%C3%B3n%20binomial%20Laboratorio%20-%20Enunciados.pdf)
#
# + [markdown] id="WqsUQvtQDpYM"
# ### Importar módulos necesarios
# * En esta sección, estaremos usando el módulo [scipy.stats](https://docs.scipy.org/doc/scipy/reference/tutorial/stats.html), el cual nos provee la clases `binom` , para la resolución de problemas de distribución de probabilidades binomiales.
# * Complementariamente usaremos el módulo matplotlib.pyplot para realizar todos los gráficos que necesitemos.
# + id="aYw6xQEfDwNv"
from scipy.stats import binom # De esta manera importamos solamente la clase deseada y no todo el módulo scipy.stats
import matplotlib.pyplot as plt # De esta manera importamos el módulo matplotlib.pyplot y le asignamos el alias de "plt" por conveniencia a la hora de trabajar con el mismo
# + [markdown] id="4MrlLbWRKJ90"
# # Ejercicios de distribución de probabilidad binomial
# 1. La probabilidad de que el comprador de un osciloscopio haga uso del servicio técnico dentro del plazo de garantía es 0,2. Para los 5 osciloscopios que cierta empresa ha vendido independientemente a 5 compradores este mes:
# > (a). ¿Cuál es la probabilidad de que exactamente 3 de los compradores hagan uso de la garantía?
# (b). ¿Cuál es la probabilidad de que 3 o más compradores hagan uso de la garantía?
# + id="ee5xtCXcFlWd"
# Declaramos las variables que necesitaremos para el ejercicio:
cant_ensayos = 5
prob_exito = 0.2
# Creamos un objeto del tipo "binom" distribución binomial, con los parametros anteriores:
dist_binomial = binom(cant_ensayos, prob_exito)
# A partir de este momento, solo necesitamos llamar al objeto dist_binomial cada
# vez que necesitemos hacer un cálculo referente a la distribución con binomial
# con los parámetros dados en el ejercicio.
# + [markdown] id="RvhMrey1Maju"
# (a). Para hallar la probabilidad de que **exactamente** 3 de los compradores hagan uso de la garantía, debemos aplicar el método `pmf()` _probability mass function_ al cual debemos pasar como parémetro x el número de ocurrencias del cual necesitamos calcular la probabilidad, para la función binomial dada.
# + id="ad-uLNL5MNRU" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="76b9416f-6b1f-4699-a3ad-653ceca1caa4"
# Para ello aplicamos la función pmf() a nuestra distribución binomial, a la cual habíamos llamado dist_binomial, de la siguiente manera:
prob_de_3 = dist_binomial.pmf(3) # Guardamos el valor calculado dentro de la variable "prob_de_3"
# A continuación imprimimos lo calculado haciendo uso de la funcion print()
# También redondeamos el resultado a 5 digitos decimales
print('La probabilidad de que exactamente 3 compradores hagan uso de su garantía es:', round(prob_de_3, 5))
# + [markdown] id="IG_Y_zeWS7qF"
# Antes de resolver el ítem (b), recordemos que:
# > $p(0) + p(1) + p(2) + p(3) + p(4) + p(5) = 1$
#
# es decir, que si queremos hallar la probabilidad de que al menos dos clientes hagan uso de la garantía podemos hallar:
# > $ p(x>2) = 1 - [p(0) + p(1) + p(2)]$
#
# (b). Para hallar la probabilidad de que 3 o más compradores hagan uso de la garantía tenemos dos opciones:
# * Aplicar la fórmula indicada más arriba. Usando el método `cdf(x)` _cumulative distribution function_ podemos calcular la probabilidad acumulada, para x = 2, de que 0, 1 y 2 compradores hagan uso de la garantía, o
# * Hacer uso del método `sf()` _survival function_ la cual calcula el valor $1-cdf(x)$ dónde cdf es la probabilidad acumulada de x.
# + id="T8ictH-AOP9q" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7ec88a41-d1be-41a0-ad5a-47b20e441d3b"
# Hallemos la respuesta usando la primera opción, la probabilidad acumulada para x=2 está dada por:
prob_acum_2 = dist_binomial.cdf(2) # Recordemos que: prob_acum_2 = p(0) + p(1) + p(2)
prob_almenos_3 = 1 - prob_acum_2
# Imprimimos la respuesta redondeada a 5 dígitos decimales
print('La probabilidad de que al menos 3 personas hagan uso de la garantía es:', round(prob_almenos_3, 5))
# + id="hh_qI3MYSnTw" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b3cb6753-e827-41d7-fb53-19f829a0ee50"
# Hallemos ahora la respuesta usando la segunda opción, para evitar confución vamos a usar otra variable: "p_al_menos_3":
p_al_menos_3 = dist_binomial.sf(2)
# Imprimimos la respuesta redondeada a 5 dígitos decimales:
print('La probabilidad de que al menos 3 personas hagan uso de la garantía es:', round(p_al_menos_3, 5))
# Podemos comparar ambos resultados y comprobar que ambos métodos son factibles
# + [markdown] id="fFsS0jZZc0Qu"
# ### Opcional
# De manera a conocer el potencial que nos ofrece el módulo `scipy.stats` haremos uso de algunas de sus funciones para hallar otros parámetros de la función binomial dada (n=5, p= 0,2)
# + id="VDGRDLpMZbrZ" colab={"base_uri": "https://localhost:8080/", "height": 90} outputId="aecfca50-a3dd-4935-a20a-ae3e3c511fd7"
# Si quisieramos calcular la media, varianza, sesgo y kurtosis para los parámetros dados, solo tendríamos que hacer uso de:
media, var, sesgo, kurt = dist_binomial.stats(moments='mvsk')
print('Media:', media)
print('Varianza:', var)
print('Sesgo:', sesgo)
print('Kurtosis:', kurt)
# + [markdown] id="SuVCKoZ3eHGF"
# También podríamos valernos de una lista de valores para x, en este caso de 0 a 5 clientes que hacen uso de la garantía y obtener una lista de probabilidades para cada valor correspondiente de x. Esto es particularmente útil para graficar la distribución de probabilidades del problema.
# + id="6jBh-oKYa--E" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="15abbe0c-bd19-4dd4-c384-bb1975544b36"
# Creamos una variable x con un rango de valores de 0 a 5
x = range(6) # La función range nos devuelve un objeto iterable que inicia en 0 (por defecto) y termina en el valor dado sin incluirlo es decir 0, 1, 2, 3, 4, 5.
# Calculemos las probabilidades para cada valor de x, las cuales guardaremos en la variable p
p = dist_binomial.pmf(x)
# Si imprimimos p, vemos que es una lista (vector) con el valor de probabilidad para cada valor de x dado
print(p) # Recordemos que x va de 0 a 5, por lo tanto p(3) es el cuarto valor de la lista p
# + id="TuDWB5c_ckqK" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="68129ce5-b097-4ab8-cc42-04f58bc88a5b"
# Finalmente podemos graficar la función haciendo uso del módulo matplotlib.plotlib, al que llamamos con el alias de plt
fig, ax = plt.subplots(1, 1) # Creamos un lienzo para nuestro gráfico
# El gráfico está dado por:
ax.plot(x, p, 'bo') # Agregamos nuestro gráfico en forma de puntos azules con el comando 'bo' -> b: blue (azul)- o: puntos
ax.vlines(x= x, ymin= 0, ymax= p, colors='b') # Agregamos las líneas verticales desde ymin= 0 hasta ymax = p(x), de color azul
# Los demás complementos, opcionales:
ax.grid() # Agrega las grillas al gráfico
ax.set_title('Distribución de probabilidades')
ax.set_xlabel('Cantidad de clientes')
ax.set_ylabel('Probabilidad de que usen la garantía')
plt.show()
# + [markdown] id="CNVhmB_Ho4C5"
# 2. Un examen de estadística de elección múltiple contenía 20 preguntas y cada una de ellas 5 respuestas. Solo hay una respuesta correcta por pregunta. Si un estudiante desconocía todas las respuestas y contestó al azar.
# > (a).¿Cuál es la probabilidad de que conteste correctamente a 5 preguntas? (b).¿Cuál es la probabilidad de que conteste correctamente a lo más 5 preguntas?
# + id="trWNeboyg1Rp"
# Declaramos las variables que necesitaremos para el ejercicio:
resp_correctas = 1
resp_posibles = 5
cant_ensayos = 20
prob_exito = resp_correctas / resp_posibles
# Creamos un objeto del tipo "binom" distribución binomial, con los parametros anteriores:
dist_binomial = binom(cant_ensayos, prob_exito)
# A partir de este momento, solo necesitamos llamar al objeto dist_binomial cada
# vez que necesitemos hacer un cálculo referente a la distribución con binomial
# con los parámetros dados en el ejercicio.
# + [markdown] id="2Ie_G4t1rj9U"
# (a).¿Cuál es la probabilidad de que conteste correctamente a 5 preguntas? Para responder a esta pregunta usamos el método `pmf()` de la clase `binom`. Para ello, recordemos que en el paso anterior ya creamos un objeto del tipo `binom` al que llamamos `dist_binomial` el cual ya contiene los datos del problema.
# + id="nCqX6TyOqy4P" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="41ae8180-48b4-49da-f323-6833e099130b"
# Para ello aplicamoc la función pmf() a nuestra distribución binomial:
prob_de_exact_5 = dist_binomial.pmf(5) # Guardamos el valor calculado dentro de la variable "prob_de_exact_5"
# A continuación imprimimos lo calculado haciendo uso de la funcion print()
# También redondeamos el resultado a 5 digitos decimales
print('La probabilidad de que responda correctamente a exactamente 5 preguntas es:', round(prob_de_exact_5, 5))
# + [markdown] id="ZIVcO4mDsydT"
# (b).¿Cuál es la probabilidad de que conteste correctamente a lo más 5 preguntas?. O dicho de otra manera cuál es la probabilidad de que conteste 0, 1, 2, 3, 4 o 5 respuestas correctamente. Para ello, hacemos uso del método `cdf(x)` lo aplicamos a nuestro objeto `dist_binom` con $x = 5$ y tendremos la respuesta buscada.
# + id="KkkpvZyCsqjY" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d5784430-0c01-4b9e-a164-7ed6f7806cfc"
prob_hasta_5 = dist_binomial.cdf(5)
# A continuación imprimimos lo calculado haciendo uso de la funcion print()
# También redondeamos el resultado a 5 digitos decimales
print('La probabilidad de que responda correctamente a lo más 5 preguntas es:', round(prob_hasta_5, 5))
# + [markdown] id="Up7VV0gTuMle"
# ### Opcional
# Graficar la distribución de probabilidades para responder correctamente 0, 1, 2... 20 preguntas del examen.
# + id="W-jUxtewu1qm"
# Primero creamos el eje x
x = range(21) # recordemos que esto crea un objeto iterable "range" con valores desde 0 (por defecto) hasta 20 (21 valores en total)
# Luego calculamos los valores de probabilidad para cada x:
p = dist_binomial.pmf(x)
# + id="kWrZ0sQ9t-ke" colab={"base_uri": "https://localhost:8080/", "height": 517} outputId="03baf46b-804b-4c67-9a70-deefce36b3e9"
fig, ax = plt.subplots(1, 1, figsize= (16,8)) # Creamos un lienzo para nuestro gráfico
# El gráfico está dado por:
ax.plot(x, p, 'bo') # Agregamos nuestro gráfico en forma de puntos azules con el comando 'bo' -> b: blue (azul)- o: puntos
ax.vlines(x= x, ymin= 0, ymax= p, colors='b') # Agregamos las líneas verticales desde ymin= 0 hasta ymax = p(x), de color azul
# Los demás complementos, opcionales:
ax.grid() # Agrega las grillas al gráfico
ax.set_title('Distribución de probabilidades', fontsize='x-large', fontweight= 'bold') # Agrega un título, con tamaño de fuente extra grande y en negritas
ax.set_xlabel('Cantidad de preguntas', fontsize='large') # Agrega texto al eje x
ax.set_ylabel('Probabilidad de responder correctamente', fontsize='large') # Agrega texto al eje y
plt.xticks(x) # Modifica los puntos representados en el eje x, y la grilla para que coincidan exactamente a intervalos enteros
plt.show() # Para mostrar el gráfico
# + [markdown] id="OGotdtCSDVLU"
# 3. Un experimento consiste en lanzar un dado varias veces. Determine la probabilidad de que aparezca:
# > (a). La cara 6 entre 29 y 32 veces inclusive, si el dado es lanzado 170 veces
# (b). La cara 6 menos de 22 veces, si el dado es lanzado 170 veces
# (c). Algún número par, si el dado es lanzado 5 veces
# + id="nW0zWnMVvcD6"
# Empezamos por definir los parámetros para las primeras dos preguntas.
# Declaramos las variables que necesitaremos para el ejercicio:
casos_favorables = 1 # Que salga la cara 6
casos_posibles = 6 # Por las 6 caras del dado
cant_ensayos = 170
prob_exito = casos_favorables / casos_posibles
# Creamos un objeto del tipo "binom" distribución binomial, con los parametros anteriores:
dist_binomial = binom(cant_ensayos, prob_exito)
# A partir de este momento, solo necesitamos llamar al objeto dist_binomial cada
# vez que necesitemos hacer un cálculo referente a la distribución con binomial
# con los parámetros dados en el ejercicio.
# + [markdown] id="kN2xdarjGC9G"
# (a). La cara 6 entre 29 y 32 veces inclusive, si el dado es lanzado 170 veces. Para resolver esto podemos hallar la probabilidad acumulada de 32 y a esta restar la probabilidad acumulada de 28, haciendo uso del método `cdf(x)`
# + id="jczf7hnEFBat" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="d4af223c-fbf6-4c80-b427-8d5e7ebe70a1"
# Hallamos la probabilidad acumulada de 32:
prob_acum_32 = dist_binomial.cdf(32)
# Hallamos la probabilidad acumulada de 28:
prob_acum_28 = dist_binomial.cdf(28)
# Hallamos la diferencia de ambas
prob_entre_29_y_32 = prob_acum_32 - prob_acum_28
# Imprimimos el resultado redondeado a 5 dígitos decimales
print('La probabilidad de que la cara 6 salga entre 29 y 32 veces es de:', round(prob_entre_29_y_32, 5))
# + [markdown] id="ieYGhMyQIrNR"
# (b). La cara 6 menos de 22 veces, si el dado es lanzado 170 veces. Es decir la probabilidad acumulada de 21.
# + id="yFa1RRzAHunp" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0559f7ec-c3e4-4a81-9927-cd3bed45db6d"
prob_acum_21 = dist_binomial.cdf(21)
# Imprimimos el resultado redondeado a 5 dígitos decimales
print('La probabilidad de que la cara 6 salga menos de 22 veces es:', round(prob_acum_21, 5))
# + [markdown] id="H_2Y30tiLMq0"
# (c). Algún número par, si el dado es lanzado 5 veces. Para responder a esta pregunta debemos volver a crear el objeto dist_binomial, o crear uno distinto,para los datos dados. Hecho esto, debemos considerar que la respuesta estará satisfecha si obtenemos 1, 2, 3, 4 o 5 numeros pares.
#
# + id="AaO5V0l8JaMM"
# Declaramos las variables con los nuevos datos:
casos_favorables = 3 # Cantidad de números pares del dado: 2, 4 y 6
casos_posibles = 6 # El total de caras del dado
prob_exito = casos_favorables / casos_posibles
cant_ensayos = 5
# Para este ejemplo, simplmente vamos a reutilizar nuestra variable
dist_binomial = binom(cant_ensayos, prob_exito)
# + id="OT7aSH8zM7ds" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="80857b26-7f12-4ea0-f287-f229a81fe12c"
# Una forma de hacer esto es calcular la probabilidad para x= 1, 2, 3, 4, 5 y sumar todos los resultados:
# Calcular las probabilidades para x=[1,2,3,4,5] puede hacerse en un solo paso, como vimos al hacer los gráficos
p = dist_binomial.pmf(range(1,6)) # Esto nos devuelve un vector con todas las probabilidades, para x=[1,2,3,4,5]
prob_algun_par = sum(p) # Esto nos permite hallar la suma de todos los valores obtenidos en el array p.
# Imprimimos el resultado
print('La probabilidad de que salga algú número par, si el dado es lanzado 5 veces, es:', round(prob_algun_par, 5))
# + [markdown] id="S2M2kPyBZU0h"
# 4. Una fábrica produce en cada turno 100 piezas de forma que la probabilidad de que una sea defectuosa es 0.05. En elcontrol de calidad se revisan todas las piezas y se depositan las defectuosas en un recipiente que se vacía al final de cada turno.
# > ¿Cuántas piezas ha de contener el recipiente para que la probabilidad de que su capacidad no se vea rebasada al final de cada turno sea mayor o igual a 0,95?
# + [markdown] id="HFik6tjyaBK6"
# En otras palabras, tenemos que hallar la cantidad de botellas defectuosas (x) para la cual la probabilidad acumulada, sea mayor o igual a 0,95.
# Para esto hacemos usa del método `ppf()` _percent point function_ el cual es la inversa del método `cdf()`
# + id="iuHi8iUUXv46" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="1fa47614-591e-411c-c212-2ea071064071"
# Definimos las variables conocidas:
cant_ensayos = 100
prob_exito = 0.05
# Creamos un objeto del tipo "binom" distribución binomial, con los parametros anteriores:
dist_binomial = binom(cant_ensayos, prob_exito)
# Hallamos x, para que satisfaga p(x) >= 0.95
x = dist_binomial.ppf(0.95)
# Imprimimos el resultado
print('El recipiente debe contener al menos:', int(x), 'botellas para que la probabilidad de que su capacidad no se vea rebasada al final de cada turno sea mayor o igual a 0,95')
# + [markdown] id="xX2XHkqodWXf"
# 5. Se estima que la probabilidad de que una muestra de agua del lago Ypacarai de 1 $cm^{3}$ contenga alguna bacteria es de 0,86.
# > (a). Si se toman en forma independiente 8 muestras de agua de 1 $cm^{3}$ cada una, ¿cuál es la probabilidad de que por lo menos una muestra contenga alguna colonia de bacterias? (b). ¿Cuál es el número mínimo de muestras de 1 $cm^{3}$ que deberían tomarse para tener al menos 99% de probabilidad de observar al menos una colonia?
# + id="Stv8cq84b6vg"
# Definimos las variables del problema:
cant_ensayos = 8
prob_exito = 0.86
# Creamos un objeto del tipo "binom" distribución binomial, con los parametros anteriores:
dist_binomial = binom(cant_ensayos, prob_exito)
# + [markdown] id="Ri1EmhCbn_jr"
# > (a). Si se toman en forma independiente 8 muestras de agua de 1 $cm^{3}$ cada una, ¿cuál es la probabilidad de que por lo menos una muestra contenga alguna colonia de bacterias?
#
# Para esto usamos la función `sf(x)` con $x = 0$
# + id="53GuYkS1n9L_" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="fe4eefe9-f13f-4361-831d-7fd653c59bfe"
prob_al_menos_1 = dist_binomial.sf(0)
# Imprimimos el resultado redondeado a 5 digitos decimales
print('La probabilidad de que al menos una de las muestras contenga alguna colonia de bacterias es:', prob_al_menos_1)
# + [markdown] id="nqPux4eWqMAS"
# > (b). ¿Cuál es el número mínimo de muestras de 1 $cm^{3}$ que deberían tomarse para tener al menos 99% de probabilidad de observar al menos una colonia?
#
# En este caso, lo que tendremos que ir variando es el la cantidad de ensayos, y con esto encontrar el número mínimo de ensayos a realizar para que almenos una muestra contenga alguna colonia de bacterias con una probabilidad de 99%. Como en la parte (a) vemos que para 8 ensayos la probabilidad de obtener al menos una colonia de bacterias era más de 99,99%, tomaremos 8 como nuestro límite superior para la cantidad de ensayos a realizar.
# + id="ddZOq6x-p0P-" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="0f3306b0-1b45-4181-966a-54c721b80267"
# Definimos la cantidad de ensayos como el intervalo de 1 a 8
cant_ensayos = range(1, 9) # Rango desde 1 (inclusive) hasta 9 (excluyendo al 9)
# Esta vez no definimos un único objeto "binom" ya que la forma de la distribución binomial cambia con la cantidad de ensayos
# Por lo tanto para calcular la probabilidad para cada caso solicitado escribimos la siguiente expresión:
p = binom.sf(0, cant_ensayos, prob_exito)
print(p)
# + [markdown] id="ib6omzvDyOWA"
# Al imprimir p obtenemos las probabilidades de encontrar al menos una colonia de bacterias para los distintos valores de muestra.
# > $p = [0.86, 0.9804, 0.997256, 0.99961584, 0.99994622, 0.99999247, 0.99999895, 0.99999985]$
#
# Vemos que el 3er elemento satisface la condición de que su probabilidad sea de 99% por lo tanto la respuesta buscada es:
# $n = 3$
# + id="mPIV3pRbxVtr"
|
notebooks/Distribuciones de Probabilidad/Distribucion_Binomial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cap_env
# language: python
# name: cap_env
# ---
# # On this notebook the test and training sets will be defined.
# +
# Basic imports
import os
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import datetime as dt
import scipy.optimize as spo
import sys
# %matplotlib inline
# %pylab inline
pylab.rcParams['figure.figsize'] = (20.0, 10.0)
# %load_ext autoreload
# %autoreload 2
sys.path.append('../')
# -
# ## Let's test the scikit learn example for TimeSeriesSplit (with some modifications)
# +
from sklearn.model_selection import TimeSeriesSplit
num_samples = 30
dims = 2
X = np.random.random((num_samples,dims))
y = np.array(range(num_samples))
tscv = TimeSeriesSplit(n_splits=3)
print(tscv)
TimeSeriesSplit(n_splits=3)
for train_index, test_index in tscv.split(X):
print("TRAIN_indexes:", train_index, "TEST_indexes:", test_index)
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
# -
# ### It may be useful for validation purposes. The test set will be separated before, anyway. The criterion to follow is to always keep causality.
# ## Let's get the data and preserve one part as the test set.
# Note: The way the test set will be used, is still not defined. Also, the definition of X and y may depend on the length of the base time interval used for training. But, in any case, it is a good practise to separate a fraction of the data for test, that will be untouched regardless of all those decisions.
data_df = pd.read_pickle('../../data/data_df.pkl')
print(data_df.shape)
data_df.head(10)
# ### I will save about two years worth of data for the test set (it wouldn't be correct to save a fixed fraction of the total set because the size of the "optimal" training set is still to be defined; I may end up using much less than the total dataset).
# +
num_test_samples = 252 * 2
data_train_val_df, data_test_df = data_df.unstack().iloc[:-num_test_samples], data_df.unstack().iloc[-num_test_samples:]
# -
def show_df_basic(df):
print(df.shape)
print('Starting value: %s\nEnding value: %s' % (df.index.get_level_values(0)[0], df.index.get_level_values(0)[-1]))
print(df.head())
show_df_basic(data_train_val_df)
show_df_basic(data_test_df)
# ### I could select the Close values, for example, like below...
data_test_df.loc[slice(None),(slice(None),'Close')].head()
# ### Or like this...
data_test_df.xs('Close', level=1, axis=1).head()
# ### But I think it will be more clear if I swap the levels in the columns
data_train_val_df = data_train_val_df.swaplevel(0, 1, axis=1).stack().unstack()
show_df_basic(data_train_val_df)
data_test_df = data_test_df.swaplevel(0, 1, axis=1).stack().unstack()
show_df_basic(data_test_df)
# ## Now it's very easy to select one of the features:
data_train_val_df['Close']
# ## Let's pickle the data
data_train_val_df.to_pickle('../../data/data_train_val_df.pkl')
data_test_df.to_pickle('../../data/data_test_df.pkl')
# ## No separate validation set will be needed as I will use "time" cross-validation for that.
|
notebooks/dev/n02_separating_the_test_set.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
extracted_data_path = '../data/intermediate/extracted_data.json'
import json
with open(extracted_data_path) as fd:
extracted_data = json.load(fd)
extracted_data
# +
from collections import Counter
nb_review_by_labels = Counter([d[0] for d in extracted_data])
nb_review_by_labels.most_common()
# +
from classifier.pre_process import preprocess_data
preprocessed_data = preprocess_data(extracted_data)
# -
preprocessed_data
# +
from collections import Counter
nb_review_by_labels = Counter([d.split()[0] for d in preprocessed_data])
nb_review_by_labels.most_common()
# -
from classifier.helper import write_json
write_json('../data/intermediate/preprocessed_data.json', preprocessed_data)
|
resources/setup_project/project/notebooks/preprocess_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="1e63aba1"
# # # ?Source https://docs.neptune.ai/how-to-guides/experiment-tracking/organize-ml-experiments
# Down the page there is a link to Colab
# + id="fa4b6849"
# # ! pip install neptune-client
# + id="e4b350e0"
import sys
import os
from pathlib import Path
os.environ["NEPTUNE_API_TOKEN"] = "" # ask for it
os.environ["NEPTUNE_PROJ_NAME"] = "octavf/tree-counting-and-classif"
# source_code_path = os.path.abspath('../src')
source_code_path = Path('/content/drive/MyDrive/vork/ML/trees/tree-counting-and-classification-in-images/src')
sys.path.append(str(source_code_path))
# + [markdown] id="65361e83"
# # Data up & down
# + id="50775966"
from data.neptune_ai.neptune_wrapper import NeptuneWrapper, run
PROJ_NAME = "octavf/tree-counting-and-classif"
API_TOKEN = "" # ask for it
# + [markdown] id="eced1845"
# ## Data up
# + id="0647aea6"
run(PROJ_NAME,
API_TOKEN,
upload_ds=True, download_ds=False,
from_path="/work/training/training_1/",
to_path="dataset/PlaiulRotated30v0.1.2")
# + [markdown] id="426d694e"
# ## Data down
# + id="1077c55c"
run(PROJ_NAME,
API_TOKEN,
upload_ds=False, download_ds=True,
to_path="./dataset/latest")
# + [markdown] id="NJQ2PyV5qGtw"
# # Train model with Neptune logging
# + id="q8ONCrQYqnn-"
# # ! pip install -r /content/drive/MyDrive/vork/ML/trees/tree-counting-and-classification-in-images/requirements.txt
# # ! pip install --upgrade opencv-python setuptools==59.5.0 albumentations==1.0.3
# + id="GUoRotKvqHHc"
from models.train_model import train_model
training_folder_path = Path('/content/drive/MyDrive/vork/ML/trees/training/plaiul')
# + id="VzFa2B7ArU2c"
dataset_path = training_folder_path / 'preds'
# + id="GzpXOhbHqtKi"
train_model(
train_annotations=dataset_path.parent / 'train' / 'labels.csv',
valid_annotations=dataset_path.parent / 'valid' / 'labels.csv',
config_path=source_code_path / 'data' / 'train_config.cfg',
output_path=Path(training_folder_path / 'testNeptuneLogger'),
pretrained_path=Path(training_folder_path / 'deepforest_iter2'),
nbr_gpus=1,
log_in_neptune=True
)
# + id="dl3py-kFuOk5"
|
notebooks/1.1-foc-Neptune_ai_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # To predict diabetes using PIMA diabetes data
# ### Importing libraries
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# -
data = pd.read_csv("./data/pima-data.csv")
data.shape
data.head(5)
# check if any null value is present
data.isnull().values.any()
## Correlation
import seaborn as sns
import matplotlib.pyplot as plt
#get correlations of each features in dataset
corrmat = data.corr()
top_corr_features = corrmat.index
plt.figure(figsize=(20,20))
#plot heat map
g=sns.heatmap(data[top_corr_features].corr(),annot=True,cmap="RdYlGn")
data.corr()
# ## Changing the diabetes column data from boolean to number
diabetes_map = {True: 1, False: 0}
data['diabetes'] = data['diabetes'].map(diabetes_map)
data.head(5)
diabetes_true_count = len(data.loc[data['diabetes'] == True])
diabetes_false_count = len(data.loc[data['diabetes'] == False])
(diabetes_true_count,diabetes_false_count)
# +
## Train Test Split
from sklearn.model_selection import train_test_split
feature_columns = ['num_preg', 'glucose_conc', 'diastolic_bp', 'insulin', 'bmi', 'diab_pred', 'age', 'skin']
predicted_class = ['diabetes']
# +
X = data[feature_columns].values
y = data[predicted_class].values
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.30, random_state=10)
# -
# ## Check how many other missing(zero) values
print("total number of rows : {0}".format(len(data)))
print("number of rows missing glucose_conc: {0}".format(len(data.loc[data['glucose_conc'] == 0])))
print("number of rows missing glucose_conc: {0}".format(len(data.loc[data['glucose_conc'] == 0])))
print("number of rows missing diastolic_bp: {0}".format(len(data.loc[data['diastolic_bp'] == 0])))
print("number of rows missing insulin: {0}".format(len(data.loc[data['insulin'] == 0])))
print("number of rows missing bmi: {0}".format(len(data.loc[data['bmi'] == 0])))
print("number of rows missing diab_pred: {0}".format(len(data.loc[data['diab_pred'] == 0])))
print("number of rows missing age: {0}".format(len(data.loc[data['age'] == 0])))
print("number of rows missing skin: {0}".format(len(data.loc[data['skin'] == 0])))
# +
from sklearn.preprocessing import Imputer
fill_values = Imputer(missing_values=0, strategy="mean", axis=0)
X_train = fill_values.fit_transform(X_train)
X_test = fill_values.fit_transform(X_test)
# +
## Apply Algorithm
from sklearn.ensemble import RandomForestClassifier
random_forest_model = RandomForestClassifier(random_state=10)
random_forest_model.fit(X_train, y_train.ravel())
# +
predict_train_data = random_forest_model.predict(X_test)
from sklearn import metrics
print("Accuracy = {0:.3f}".format(metrics.accuracy_score(y_test, predict_train_data)))
# +
## Hyper Parameter Optimization
params={
"learning_rate" : [0.05, 0.10, 0.15, 0.20, 0.25, 0.30 ] ,
"max_depth" : [ 3, 4, 5, 6, 8, 10, 12, 15],
"min_child_weight" : [ 1, 3, 5, 7 ],
"gamma" : [ 0.0, 0.1, 0.2 , 0.3, 0.4 ],
"colsample_bytree" : [ 0.3, 0.4, 0.5 , 0.7 ]
}
# -
## Hyperparameter optimization using RandomizedSearchCV
from sklearn.model_selection import RandomizedSearchCV
import xgboost
classifier=xgboost.XGBClassifier()
random_search=RandomizedSearchCV(classifier,param_distributions=params,n_iter=5,scoring='roc_auc',n_jobs=-1,cv=5,verbose=3)
def timer(start_time=None):
if not start_time:
start_time = datetime.now()
return start_time
elif start_time:
thour, temp_sec = divmod((datetime.now() - start_time).total_seconds(), 3600)
tmin, tsec = divmod(temp_sec, 60)
print('\n Time taken: %i hours %i minutes and %s seconds.' % (thour, tmin, round(tsec, 2)))
from datetime import datetime
# Here we go
start_time = timer(None) # timing starts from this point for "start_time" variable
random_search.fit(X,y.ravel())
timer(start_time) # timing ends here for "start_time" variable
random_search.best_estimator_
classifier=xgboost.XGBClassifier(base_score=0.5, booster='gbtree', colsample_bylevel=1,
colsample_bytree=0.3, gamma=0.0, learning_rate=0.25,
max_delta_step=0, max_depth=3, min_child_weight=7, missing=None,
n_estimators=100, n_jobs=1, nthread=None,
objective='binary:logistic', random_state=0, reg_alpha=0,
reg_lambda=1, scale_pos_weight=1, seed=None, silent=True,
subsample=1)
from sklearn.model_selection import cross_val_score
score=cross_val_score(classifier,X,y.ravel(),cv=10)
score
score.mean()
|
assets/logos/Diabetes prediction/Diabetes_Prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ItzeelFonseca/daa_2021_1/blob/master/25Noviembre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="qgGIoWqj-0V2"
# # Conteo Frase UNAM
# + colab={"base_uri": "https://localhost:8080/"} id="lzUbJdfS47ex" outputId="d96a28a0-927e-44a2-d727-2495cecb2fe4"
frase ="""El lema que anima a la Universidad Nacional, Por mi raza hablará el espíritu, revela la vocación humanística con la que fue concebida. El autor de esta célebre frase, <NAME>, asumió la rectoría en 1920, en una época en que las esperanzas de la Revolución aún estaban vivas,
había una gran fe en la Patria y el ánimo redentor se extendía en el ambiente."""
frase= frase.strip().replace("\n","").replace(".","").replace(",","").lower().split(" ")
#print(frase)
frecuencias = {}
for index in range (len(frase)):
if frase[index] in frecuencias:
pass
else:
frecuencias[frase[index]] = 1
for pivote in range (index +1,len(frase),1):
#print(frase[index].strip(), "Comparada contra: " , frase[pivote].strip())
if frase[index] == frase[pivote]:
frecuencias [frase[index]] += 1
print(frecuencias)
# + [markdown] id="4rZblae0ATIk"
# # Ejemplo hash
# + colab={"base_uri": "https://localhost:8080/"} id="Oi5jTMlj_XfE" outputId="998ac940-75b7-4dcb-9e6a-92c0b1906bf8"
palabra = 'hola'
print(hash(palabra))
palabra = 'hole'
print(hash(palabra))
# + [markdown] id="4O_Xt7UoGiu0"
# # Ejemplo Frase hash
# + colab={"base_uri": "https://localhost:8080/"} id="p-jBux9sFwz7" outputId="77afc454-d280-4374-ab6b-3b35bde0c55d"
frase ="""El lema que anima a la Universidad Nacional, Por mi raza hablará el espíritu, revela la vocación humanística con la que fue concebida. El autor de esta célebre frase, <NAME>, asumió la rectoría en 1920, en una época en que las esperanzas de la Revolución aún estaban vivas,
había una gran fe en la Patria y el ánimo redentor se extendía en el ambiente."""
frase= frase.strip().replace("\n","").replace(".","").replace(",","").lower().split(" ")
print(frase)
frecuencias = {}
for index in range (len(frase)):
print(hash(frase[index]))
if frase[index] in frecuencias:
frecuencias[frase[index]] += 1
else:
frecuencias[frase[index]] = 1
print(frecuencias)
|
25Noviembre.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelling api response times with
# # 1. requests
# # 2. response time
visitors = 100
# +
from matplotlib import pyplot as plt
# %matplotlib inline
# -
# # Requests are coming within $\lambda$ milliseconds from each other (this is how much we would sleep between requests)
#
# # $\lambda$ = Poisson($\mu$)+1
#
# +
from scipy.stats import poisson
mu = 10
def gen_distances(mu, N=visitors):
return [1. + x for x in poisson.rvs(mu=mu, size=N)]
distances_between_requests = gen_distances(mu)
distances_between_requests[:10]
# -
plt.figure(figsize=(16,2))
plt.scatter([sum(distances_between_requests[:n]) for n in range(visitors)], [0. for _ in range(visitors)], s=1)
;
for _ in range(10):
print(sum(gen_distances(mu)))
# # 1000ms = visitors (100) * mu (10)
# # If we want to have control on RPS, let's ask to configure $\mu$
# # RPS ~~ $\mu$*100
# # $\mu$ = RPS / 100
# # API response time is $\theta$ ms
# (interesting reading: https://blog.newrelic.com/engineering/expected-distributions-website-response-times/)
# # $\theta$ = $\Gamma$($\alpha$) with $\alpha>0$ of gamma distribution
#
# +
from scipy.stats import gamma
alpha = 100
def gen_response_times(alpha, N=visitors):
return gamma.rvs(a=alpha, size=N)
response_times = gen_response_times(alpha)
response_times[:10]
# -
plt.figure(figsize=(16,8))
plt.hist(response_times)
;
# # Group it alltogether
# +
response_times = gen_response_times(alpha)
distances_between_requests = gen_distances(mu)
Xs = [sum(distances_between_requests[:n]) for n in range(visitors)]
plt.figure(figsize=(16,8))
plt.scatter(Xs, response_times)
;
# -
|
example-app/Example-app.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Aligning Data to Darwin Core - Sampling Event with Measurement or Fact using Python
# <NAME>
#
# November 9, 2020
#
# # General information about this notebook
# This notebook was created for the IOOS DMAC Code Sprint Biological Data Session
# The data in this notebook were created specifically as an example and meant solely to be
# illustrative of the process for aligning data to the biological data standard - Darwin Core.
# These data should not be considered actually occurrences of species and any measurements
# are also contrived. This notebook is meant to provide a step by step process for taking
# original data and aligning it to Darwin Core
#
# This notebook is a python implementation of the R notebook [IOOS_DMAC_DataToDWC_Notebook_event.R](https://github.com/ioos/bio_data_guide/blob/master/Standardizing%20Marine%20Biological%20Data/datasets/example_script_with_fake_data/IOOS_DMAC_DataToDwC_Notebook_event.R)
import pandas as pd
import pyworms # pip install git+git://github.com/iobis/pyworms.git
import numpy as np
import uuid
import csv
# Read in the raw data file.
file = 'data/MadeUpDataForBiologicalDataTraining.csv'
df = pd.read_csv(file, header=[0])
# First we need to to decide if we will provide an occurrence only version of the data or
# a sampling event with measurement or facts version of the data. Occurrence only is easier
# to create. It's only one file to produce. However, several pieces of information will be
# left out if we choose that option. If we choose to do sampling event with measurement or
# fact we'll be able to capture all of the data in the file creating a lossless version.
# Here we decide to use the sampling event option to include as much information as we can.
#
# First let's create the eventID and occurrenceID in the original file so that information
# can be reused for all necessary files down the line.
# +
df['eventID'] = df[['region', 'station', 'transect']].apply(lambda x: '_'.join(x.astype(str)), axis=1)
df['occurrenceID'] = uuid.uuid4()
# uuid for each row
#for index, row in df.iterrows():
# df.loc[index, 'occurrenceID'] = uuid.uuid4()
# -
# # Event file
# We will need to create three separate files to comply with the sampling event format.
# We'll start with the event file but we only need to include the columns that are relevant
# to the event file.
event = df[['date', 'lat', 'lon', 'region', 'station', 'transect', 'depth', 'bottom type', 'eventID']].copy()
# Next we need to rename any columns of data that match directly to Darwin Core. We know
# this based on our crosswalk spreadsheet CrosswalkToDarwinCore.csv
event['decimalLatitude'] = event['lat']
event['decimalLongitude'] = event['lon']
event['minimumDepthInMeters'] = event['depth']
event['maximumDepthInMeters'] = event['depth']
event['habitat'] = event['bottom type']
event['island'] = event['region']
# Let's see how it looks:
event.head()
# We need to convert the date to ISO format
event['eventDate'] = pd.to_datetime(
event['date'],
format='%m/%d/%Y',
utc=True)
# We will also have to add any missing required fields
event['basisOfRecord'] = 'HumanObservation'
event['geodeticDatum'] = 'EPSG:4326 WGS84'
# Then we'll remove any columns that we no longer need to clean things up a bit.
event.drop(
columns=['date', 'lat', 'lon', 'region', 'station', 'transect', 'depth', 'bottom type'],
inplace=True)
# We have too many repeating rows of information. We can pare this down using eventID which
# is a unique identifier for each sampling event in the data- which is six, three transects
# per site.
# +
event.drop_duplicates(
subset='eventID',
inplace=True)
event.head(6)
# -
# Finally we write out the event file
event.to_csv(
'MadeUpData_event_frompy.csv',
header=True,
index=False,
date_format='%Y-%m-%d')
# # Occurrence file
# Next we need to create the occurrence file. We start by creating the dataframe.
occurrence = df[['scientific name', 'eventID', 'occurrenceID', 'percent cover']].copy()
# Then we'll rename the columns that align directly with Darwin Core.
occurrence['scientificName'] = occurrence['scientific name']
# Finally we'll add required information that's missing.
occurrence['occurrenceStatus'] = np.where(occurrence['percent cover'] == 0, 'absent', 'present')
# ## Taxonomic Name Matching
# A requirement for OBIS is that all scientific names match to the World Register of
# Marine Species (WoRMS) and a scientificNameID is included. A scientificNameID looks
# like this "urn:lsid:marinespecies.org:taxname:275730" with the last digits after
# the colon being the WoRMS aphia ID. We'll need to go out to WoRMS to grab this
# information.
#
# Create a lookup table of unique scientific names
lut_worms = pd.DataFrame(
columns=['scientificName'],
data=occurrence['scientificName'].unique())
# Add the columns that we can grab information from WoRMS including the required scientificNameID.
# +
headers = ['acceptedname', 'acceptedID', 'scientificNameID', 'kingdom', 'phylum',
'class', 'order', 'family', 'genus', 'scientificNameAuthorship', 'taxonRank']
for head in headers:
lut_worms[head] = ''
# -
# Taxonomic lookup using the library [pyworms](https://github.com/iobis/pyworms)
for index, row in lut_worms.iterrows():
print('Searching for scientific name = %s' % row['scientificName'])
resp = pyworms.aphiaRecordsByMatchNames(row['scientificName'])[0][0]
lut_worms.loc[index, 'acceptedname'] = resp['valid_name']
lut_worms.loc[index, 'acceptedID'] = resp['valid_AphiaID']
lut_worms.loc[index, 'scientificNameID'] = resp['lsid']
lut_worms.loc[index, 'kingdom'] = resp['kingdom']
lut_worms.loc[index, 'phylum'] = resp['phylum']
lut_worms.loc[index, 'class'] = resp['class']
lut_worms.loc[index, 'order'] = resp['order']
lut_worms.loc[index, 'family'] = resp['family']
lut_worms.loc[index, 'genus'] = resp['genus']
lut_worms.loc[index, 'scientificNameAuthorship'] = resp['authority']
lut_worms.loc[index, 'taxonRank'] = resp['rank']
# Merge the lookup table of unique scientific names back with the occurrence data.
occurrence = pd.merge(occurrence, lut_worms, how='left', on='scientificName')
# We're going to remove any unnecessary columns to clean up the file
occurrence.drop(
columns=['scientific name', 'percent cover'],
inplace=True)
# Quick look at what we have before we write out the file
occurrence.head()
# Write out the file.
# +
# sort the columns on scientificName
occurrence.sort_values('scientificName', inplace=True)
# reorganize column order to be consistent with R example:
columns = ["scientificName","eventID","occurrenceID","occurrenceStatus","acceptedname","acceptedID",
"scientificNameID","kingdom","phylum","class","order","family","genus","scientificNameAuthorship",
"taxonRank"]
occurrence.to_csv(
"MadeUpData_Occurrence_frompy.csv",
header=True,
index=False,
quoting=csv.QUOTE_ALL,
columns=columns)
# -
# All done with occurrence!
# # Measurement Or Fact
# The last file we need to create is the measurement or fact file. For this we need to
# combine all of the measurements or facts that we want to include making sure to include
# IDs from the BODC NERC vocabulary where possible.
# Temperature
temperature = df[['eventID', 'temperature', 'date']].copy()
temperature['occurrenceID'] = ''
temperature['measurementType'] = 'temperature'
temperature['measurementTypeID'] = 'http://vocab.nerc.ac.uk/collection/P25/current/WTEMP/'
temperature['measurementValue'] = temperature['temperature']
temperature['measurementUnit'] = 'Celsius'
temperature['measurementUnitID'] = 'http://vocab.nerc.ac.uk/collection/P06/current/UPAA/'
temperature['measurementAccuracy'] = 3
temperature['measurementDeterminedDate'] = pd.to_datetime(temperature['date'],
format='%m/%d/%Y',
utc=True)
temperature['measurementMethod'] = ''
temperature.drop(
columns=['temperature', 'date'],
inplace=True)
# Rugosity
rugosity = df[['eventID', 'rugosity', 'date']].copy()
rugosity['occurrenceID'] = ''
rugosity['measurementType'] = 'rugosity'
rugosity['measurementTypeID'] = ''
rugosity['measurementValue'] = rugosity['rugosity'].map('{:,.6f}'.format)
rugosity['measurementUnit'] = ''
rugosity['measurementUnitID'] = ''
rugosity['measurementAccuracy'] = ''
rugosity['measurementDeterminedDate'] = pd.to_datetime(rugosity['date'],
format='%m/%d/%Y',
utc=True)
rugosity['measurementMethod'] = ''
rugosity.drop(
columns=['rugosity', 'date'],
inplace=True)
# Percent cover
percent_cover = df[['eventID', 'occurrenceID', 'percent cover', 'date']].copy()
percent_cover['measurementType'] = 'Percent Cover'
percent_cover['measurementTypeID'] = 'http://vocab.nerc.ac.uk/collection/P01/current/SDBIOL10/'
percent_cover['measurementValue'] = percent_cover['percent cover']
percent_cover['measurementUnit'] = 'Percent/100m^2'
percent_cover['measurementUnitID'] = ''
percent_cover['measurementAccuracy'] = 5
percent_cover['measurementDeterminedDate'] = pd.to_datetime(percent_cover['date'],
format='%m/%d/%Y',
utc=True)
percent_cover['measurementMethod'] = ''
percent_cover.drop(
columns=['percent cover', 'date'],
inplace=True)
# Concatenate all measurements or facts together.
measurementorfact = pd.concat([temperature, rugosity, percent_cover])
# Let's check to see what it looks like
measurementorfact.head(50)
# Write measurement or fact file
measurementorfact.to_csv('MadeUpDate_mof_frompy.csv',
index=False,
header=True,
date_format='%Y-%m-%d')
|
datasets/example_script_with_fake_data/IOOS DMAC DataToDwC_Notebook_event.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''ml'': conda)'
# language: python
# name: python37464bitmlcondaa8f3f69cf251458aa86fbbc582b758f6
# ---
# +
import pandas as pd
import swifter
data = pd.read_csv('covid19_articles/test_set/combined.csv')
# -
vox_data = pd.read_csv('covid19_articles/test_set/vox.csv')
combined = pd.concat((data, vox_data), ignore_index=True)
combined
# +
import pandas as pd
import swifter
import numpy as np
from evaluate import get_clean, eval_category, eval_emotion, eval_fake, eval_sent
combined = pd.read_csv('covid19_articles/test_set/results_combined.csv', index_col=0)
# -
combined
# +
import numpy as np
from evaluate import get_clean, eval_category, eval_emotion, eval_fake, eval_sent
def get_category(text, title):
try:
cleaned = get_clean(text, title, True)
category = eval_category([cleaned])
return category
except:
return np.nan
# combined[['title','text']].apply(lambda x: f(*x), axis=1)
combined['pred_category'] = combined[['text','title']].swifter.apply(lambda x: get_category(*x), axis=1)
# +
def get_fake(text, title):
try:
cleaned = get_clean(text, title, True)
fake = eval_fake([cleaned])
return fake
except:
return np.nan
combined['pred_fake'] = combined[['text','title']].swifter.apply(lambda x: get_fake(*x), axis=1)
# -
combined.to_csv('covid19_articles/test_set/results_combined.csv')
# +
def get_sent(text, title):
try:
cleaned = get_clean(text, title, False)
sent = eval_sent(cleaned)
return sent
except:
return np.nan
combined['pred_sent'] = combined[['text','title']].swifter.apply(lambda x: get_sent(*x), axis=1)
# -
combined.to_csv('covid19_articles/test_set/results_combined.csv')
# +
def get_emotion(text, title):
try:
cleaned = get_clean(text, title, False)
emotion = eval_emotion(cleaned)
return emotion
except:
return np.nan
combined['pred_emotion'] = combined[['text','title']].swifter.apply(lambda x: get_emotion(*x), axis=1)
# -
combined.isnull().any()
combined.to_csv('covid19_articles/test_set/results_combined.csv')
# +
from collections import Counter
ctr = Counter(combined['pred_emotion'])
# -
combined
|
historical.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="IvAWhBeZvfxR"
# # The Annotated Encoder-Decoder with Attention
#
# Recently, <NAME> wrote a blog post called [The Annotated Transformer](http://nlp.seas.harvard.edu/2018/04/03/attention.html), describing the Transformer model from the paper [Attention is All You Need](https://arxiv.org/abs/1706.03762). This post can be seen as a **prequel** to that: *we will implement an Encoder-Decoder with Attention* using (Gated) Recurrent Neural Networks, very closely following the original attention-based neural machine translation paper ["Neural Machine Translation by Jointly Learning to Align and Translate"](https://arxiv.org/abs/1409.0473) of Bahdanau et al. (2015).
#
# The idea is that going through both blog posts will make you familiar with two very influential sequence-to-sequence architectures. If you have any comments or suggestions, please let me know: [@bastings_nlp](https://twitter.com/bastings_nlp).
# + [markdown] id="A-gyjp6lvfxW"
# # Model Architecture
#
# We will model the probability $p(Y\mid X)$ of a target sequence $Y=(y_1, \dots, y_{N})$ given a source sequence $X=(x_1, \dots, x_M)$ directly with a neural network: an Encoder-Decoder.
#
# <img src="https://github.com/bastings/annotated_encoder_decoder/blob/master/images/bahdanau.png?raw=1" width="636">
#
# #### Encoder
#
# The encoder reads in the source sentence (*at the bottom of the figure*) and produces a sequence of hidden states $\mathbf{h}_1, \dots, \mathbf{h}_M$, one for each source word. These states should capture the meaning of a word in its context of the given sentence.
#
# We will use a bi-directional recurrent neural network (Bi-RNN) as the encoder; a Bi-GRU in particular.
#
# First of all we **embed** the source words.
# We simply look up the **word embedding** for each word in a (randomly initialized) lookup table.
# We will denote the word embedding for word $i$ in a given sentence with $\mathbf{x}_i$.
# By embedding words, our model may exploit the fact that certain words (e.g. *cat* and *dog*) are semantically similar, and can be processed in a similar way.
#
# Now, how do we get hidden states $\mathbf{h}_1, \dots, \mathbf{h}_M$? A forward GRU reads the source sentence left-to-right, while a backward GRU reads it right-to-left.
# Each of them follows a simple recursive formula:
# $$\mathbf{h}_j = \text{GRU}( \mathbf{x}_j , \mathbf{h}_{j - 1} )$$
# i.e. we obtain the next state from the previous state and the current input word embedding.
#
# The hidden state of the forward GRU at time step $j$ will know what words **precede** the word at that time step, but it doesn't know what words will follow. In contrast, the backward GRU will only know what words **follow** the word at time step $j$. By **concatenating** those two hidden states (*shown in blue in the figure*), we get $\mathbf{h}_j$, which captures word $j$ in its full sentence context.
#
#
# #### Decoder
#
# The decoder (*at the top of the figure*) is a GRU with hidden state $\mathbf{s_i}$. It follows a similar formula to the encoder, but takes one extra input $\mathbf{c}_{i}$ (*shown in yellow*).
#
# $$\mathbf{s}_{i} = f( \mathbf{s}_{i - 1}, \mathbf{y}_{i - 1}, \mathbf{c}_i )$$
#
# Here, $\mathbf{y}_{i - 1}$ is the previously generated target word (*not shown*).
#
# At each time step, an **attention mechanism** dynamically selects that part of the source sentence that is most relevant for predicting the current target word. It does so by comparing the last decoder state with each source hidden state. The result is a context vector $\mathbf{c_i}$ (*shown in yellow*).
# Later the attention mechanism is explained in more detail.
#
# After computing the decoder state $\mathbf{s}_i$, a non-linear function $g$ (which applies a [softmax](https://en.wikipedia.org/wiki/Softmax_function)) gives us the probability of the target word $y_i$ for this time step:
#
# $$ p(y_i \mid y_{<i}, x_1^M) = g(\mathbf{s}_i, \mathbf{c}_i, \mathbf{y}_{i - 1})$$
#
# Because $g$ applies a softmax, it provides a vector the size of the output vocabulary that sums to 1.0: it is a distribution over all target words. During test time, we would select the word with the highest probability for our translation.
#
# Now, for optimization, a [cross-entropy loss](https://ml-cheatsheet.readthedocs.io/en/latest/loss_functions.html#cross-entropy) is used to maximize the probability of selecting the correct word at this time step. All parameters (including word embeddings) are then updated to maximize this probability.
#
#
# + [markdown] id="iXvhQ4-uvfxY"
# # Prelims
#
# This tutorial requires **PyTorch >= 0.4.1** and was tested with **Python 3.6**.
#
# Make sure you have those versions, and install the packages below if you don't have them yet.
# + id="9Vnvv3ZIvfxa"
# !pip install --upgrade sacrebleu torch==1.5.1+cu101 torchvision==0.6.1+cu101 -f https://download.pytorch.org/whl/torch_stable.html
# + id="E5QE87_vvfxh" outputId="6eff2988-e575-4eaa-d400-96c72c18786a" colab={"base_uri": "https://localhost:8080/"}
# %matplotlib inline
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import math, copy, time
import matplotlib.pyplot as plt
from torch.nn.utils.rnn import pack_padded_sequence, pad_packed_sequence
from IPython.core.debugger import set_trace
# we will use CUDA if it is available
USE_CUDA = torch.cuda.is_available()
DEVICE=torch.device('cuda:0') # or set to 'cpu'
print("CUDA:", USE_CUDA)
print(DEVICE)
seed = 42
np.random.seed(seed)
torch.manual_seed(seed)
torch.cuda.manual_seed(seed)
# + [markdown] id="60ouSCRbvfxp"
# # Let's start coding!
#
# ## Model class
#
# Our base model class `EncoderDecoder` is very similar to the one in *The Annotated Transformer*.
#
# One difference is that our encoder also returns its final states (`encoder_final` below), which is used to initialize the decoder RNN. We also provide the sequence lengths as the RNNs require those.
# + id="ZNS7Pj1svfxq"
class EncoderDecoder(nn.Module):
"""
A standard Encoder-Decoder architecture. Base for this and many
other models.
"""
def __init__(self, encoder, decoder, src_embed, trg_embed, generator):
super(EncoderDecoder, self).__init__()
self.encoder = encoder
self.decoder = decoder
self.src_embed = src_embed
self.trg_embed = trg_embed
self.generator = generator
def forward(self, src, trg, src_mask, trg_mask, src_lengths, trg_lengths):
"""Take in and process masked src and target sequences."""
encoder_hidden, encoder_final = self.encode(src, src_mask, src_lengths)
return self.decode(encoder_hidden, encoder_final, src_mask, trg, trg_mask)
def encode(self, src, src_mask, src_lengths):
return self.encoder(self.src_embed(src), src_mask, src_lengths)
def decode(self, encoder_hidden, encoder_final, src_mask, trg, trg_mask,
decoder_hidden=None):
return self.decoder(self.trg_embed(trg), encoder_hidden, encoder_final,
src_mask, trg_mask, hidden=decoder_hidden)
# + [markdown] id="J1q_dPjbvfxw"
# To keep things easy we also keep the `Generator` class the same.
# It simply projects the pre-output layer ($x$ in the `forward` function below) to obtain the output layer, so that the final dimension is the target vocabulary size.
# + id="4yBZdeNsvfxx"
class Generator(nn.Module):
"""Define standard linear + softmax generation step."""
def __init__(self, hidden_size, vocab_size):
super(Generator, self).__init__()
self.proj = nn.Linear(hidden_size, vocab_size, bias=False)
def forward(self, x):
return F.log_softmax(self.proj(x), dim=-1)
# + [markdown] id="4WnsqyBuvfx1"
# ## Encoder
#
# Our encoder is a bi-directional GRU.
#
# Because we want to process multiple sentences at the same time for speed reasons (it is more effcient on GPU), we need to support **mini-batches**. Sentences in a mini-batch may have different lengths, which means that the RNN needs to unroll further for certain sentences while it might already have finished for others:
#
# ```
# Example: mini-batch with 3 source sentences of different lengths (7, 5, and 3).
# End-of-sequence is marked with a "3" here, and padding positions with "1".
#
# # +---------------+
# | 4 5 9 8 7 8 3 |
# # +---------------+
# | 5 4 8 7 3 1 1 |
# # +---------------+
# | 5 8 3 1 1 1 1 |
# # +---------------+
# ```
# You can see that, when computing hidden states for this mini-batch, for sentence #2 and #3 we will need to stop updating the hidden state after we have encountered "3". We don't want to incorporate the padding values (1s).
#
# Luckily, PyTorch has convenient helper functions called `pack_padded_sequence` and `pad_packed_sequence`.
# These functions take care of masking and padding, so that the resulting word representations are simply zeros after a sentence stops.
#
# The code below reads in a source sentence (a sequence of word embeddings) and produces the hidden states.
# It also returns a final vector, a summary of the complete sentence, by concatenating the first and the last hidden states (they have both seen the whole sentence, each in a different direction). We will use the final vector to initialize the decoder.
# + id="3w3Wbmk-vfx2"
class Encoder(nn.Module):
"""Encodes a sequence of word embeddings"""
def __init__(self, input_size, hidden_size, num_layers=1, dropout=0.):
super(Encoder, self).__init__()
self.num_layers = num_layers
self.rnn = nn.GRU(input_size, hidden_size, num_layers,
batch_first=True, bidirectional=True, dropout=dropout)
def forward(self, x, mask, lengths):
"""
Applies a bidirectional GRU to sequence of embeddings x.
The input mini-batch x needs to be sorted by length.
x should have dimensions [batch, time, dim].
"""
packed = pack_padded_sequence(x, lengths, batch_first=True)
output, final = self.rnn(packed)
output, _ = pad_packed_sequence(output, batch_first=True)
# we need to manually concatenate the final states for both directions
fwd_final = final[0:final.size(0):2]
bwd_final = final[1:final.size(0):2]
final = torch.cat([fwd_final, bwd_final], dim=2) # [num_layers, batch, 2*dim]
return output, final
# + [markdown] id="dFRYezDZvfx7"
# ### Decoder
#
# The decoder is a conditional GRU. Rather than starting with an empty state like the encoder, its initial hidden state results from a projection of the encoder final vector.
#
# #### Training
# In `forward` you can find a for-loop that computes the decoder hidden states one time step at a time.
# Note that, during training, we know exactly what the target words should be! (They are in `trg_embed`.) This means that we are not even checking here what the prediction is! We simply feed the correct previous target word embedding to the GRU at each time step. This is called teacher forcing.
#
# The `forward` function returns all decoder hidden states and pre-output vectors. Elsewhere these are used to compute the loss, after which the parameters are updated.
#
# #### Prediction
# For prediction time, for forward function is only used for a single time step. After predicting a word from the returned pre-output vector, we can call it again, supplying it the word embedding of the previously predicted word and the last state.
# + id="N_5dHeC5vfx8"
class Decoder(nn.Module):
"""A conditional RNN decoder with attention."""
def __init__(self, emb_size, hidden_size, attention, num_layers=1, dropout=0.5,
bridge=True):
super(Decoder, self).__init__()
self.hidden_size = hidden_size
self.num_layers = num_layers
self.attention = attention
self.dropout = dropout
self.rnn = nn.GRU(emb_size + 2*hidden_size, hidden_size, num_layers,
batch_first=True, dropout=dropout)
# to initialize from the final encoder state
self.bridge = nn.Linear(2*hidden_size, hidden_size, bias=True) if bridge else None
self.dropout_layer = nn.Dropout(p=dropout)
self.pre_output_layer = nn.Linear(hidden_size + 2*hidden_size + emb_size,
hidden_size, bias=False)
def forward_step(self, prev_embed, encoder_hidden, src_mask, proj_key, hidden):
"""Perform a single decoder step (1 word)"""
# compute context vector using attention mechanism
query = hidden[-1].unsqueeze(1) # [#layers, B, D] -> [B, 1, D]
context, attn_probs = self.attention(
query=query, proj_key=proj_key,
value=encoder_hidden, mask=src_mask)
# update rnn hidden state
rnn_input = torch.cat([prev_embed, context], dim=2)
output, hidden = self.rnn(rnn_input, hidden)
pre_output = torch.cat([prev_embed, output, context], dim=2)
pre_output = self.dropout_layer(pre_output)
pre_output = self.pre_output_layer(pre_output)
return output, hidden, pre_output
def forward(self, trg_embed, encoder_hidden, encoder_final,
src_mask, trg_mask, hidden=None, max_len=None):
"""Unroll the decoder one step at a time."""
# the maximum number of steps to unroll the RNN
if max_len is None:
max_len = trg_mask.size(-1)
# initialize decoder hidden state
if hidden is None:
hidden = self.init_hidden(encoder_final)
# pre-compute projected encoder hidden states
# (the "keys" for the attention mechanism)
# this is only done for efficiency
proj_key = self.attention.key_layer(encoder_hidden)
# here we store all intermediate hidden states and pre-output vectors
decoder_states = []
pre_output_vectors = []
# unroll the decoder RNN for max_len steps
for i in range(max_len):
prev_embed = trg_embed[:, i].unsqueeze(1)
output, hidden, pre_output = self.forward_step(
prev_embed, encoder_hidden, src_mask, proj_key, hidden)
decoder_states.append(output)
pre_output_vectors.append(pre_output)
decoder_states = torch.cat(decoder_states, dim=1)
pre_output_vectors = torch.cat(pre_output_vectors, dim=1)
return decoder_states, hidden, pre_output_vectors # [B, N, D]
def init_hidden(self, encoder_final):
"""Returns the initial decoder state,
conditioned on the final encoder state."""
if encoder_final is None:
return None # start with zeros
return torch.tanh(self.bridge(encoder_final))
# + [markdown] id="2MZH2v1uvfyD"
# ### Attention
#
# At every time step, the decoder has access to *all* source word representations $\mathbf{h}_1, \dots, \mathbf{h}_M$.
# An attention mechanism allows the model to focus on the currently most relevant part of the source sentence.
# The state of the decoder is represented by GRU hidden state $\mathbf{s}_i$.
# So if we want to know which source word representation(s) $\mathbf{h}_j$ are most relevant, we will need to define a function that takes those two things as input.
#
# Here we use the MLP-based, additive attention that was used in Bahdanau et al.:
#
# <img src="https://github.com/bastings/annotated_encoder_decoder/blob/master/images/attention.png?raw=1" width="280">
#
#
# We apply an MLP with tanh-activation to both the current decoder state $\bf s_i$ (the *query*) and each encoder state $\bf h_j$ (the *key*), and then project this to a single value (i.e. a scalar) to get the *attention energy* $e_{ij}$.
#
# Once all energies are computed, they are normalized by a softmax so that they sum to one:
#
# $$ \alpha_{ij} = \text{softmax}(\mathbf{e}_i)[j] $$
#
# $$\sum_j \alpha_{ij} = 1.0$$
#
# The context vector for time step $i$ is then a weighted sum of the encoder hidden states (the *values*):
# $$\mathbf{c}_i = \sum_j \alpha_{ij} \mathbf{h}_j$$
# + id="RNd_GG5evfyE"
class BahdanauAttention(nn.Module):
"""Implements Bahdanau (MLP) attention"""
def __init__(self, hidden_size, key_size=None, query_size=None):
super(BahdanauAttention, self).__init__()
# We assume a bi-directional encoder so key_size is 2*hidden_size
key_size = 2 * hidden_size if key_size is None else key_size
query_size = hidden_size if query_size is None else query_size
self.key_layer = nn.Linear(key_size, hidden_size, bias=False)
self.query_layer = nn.Linear(query_size, hidden_size, bias=False)
self.energy_layer = nn.Linear(hidden_size, 1, bias=False)
# to store attention scores
self.alphas = None
def forward(self, query=None, proj_key=None, value=None, mask=None):
assert mask is not None, "mask is required"
# We first project the query (the decoder state).
# The projected keys (the encoder states) were already pre-computated.
query = self.query_layer(query)
# Calculate scores.
scores = self.energy_layer(torch.tanh(query + proj_key))
scores = scores.squeeze(2).unsqueeze(1)
# Mask out invalid positions.
# The mask marks valid positions so we invert it using `mask & 0`.
scores.data.masked_fill_(mask == 0, -float('inf'))
# Turn scores to probabilities.
alphas = F.softmax(scores, dim=-1)
self.alphas = alphas
# The context vector is the weighted sum of the values.
context = torch.bmm(alphas, value)
# context shape: [B, 1, 2D], alphas shape: [B, 1, M]
return context, alphas
# + [markdown] id="JIBemJDPvfyL"
# ## Embeddings and Softmax
# We use learned embeddings to convert the input tokens and output tokens to vectors of dimension `emb_size`.
#
# We will simply use PyTorch's [nn.Embedding](https://pytorch.org/docs/stable/nn.html?highlight=embedding#torch.nn.Embedding) class.
# + [markdown] id="23LwtoQrvfyM"
# ## Full Model
#
# Here we define a function from hyperparameters to a full model.
# + id="Rxi1S1FevfyN"
def make_model(src_vocab, tgt_vocab, emb_size=256, hidden_size=512, num_layers=1, dropout=0.1):
"Helper: Construct a model from hyperparameters."
attention = BahdanauAttention(hidden_size)
model = EncoderDecoder(
Encoder(emb_size, hidden_size, num_layers=num_layers, dropout=dropout),
Decoder(emb_size, hidden_size, attention, num_layers=num_layers, dropout=dropout),
nn.Embedding(src_vocab, emb_size),
nn.Embedding(tgt_vocab, emb_size),
Generator(hidden_size, tgt_vocab))
return model.cuda() if USE_CUDA else model
# + [markdown] id="cB3lZx4EvfyT"
# # Training
#
# This section describes the training regime for our models.
# + [markdown] id="otPkGqOsvfyU"
# We stop for a quick interlude to introduce some of the tools
# needed to train a standard encoder decoder model. First we define a batch object that holds the src and target sentences for training, as well as their lengths and masks.
# + [markdown] id="eVFez8JIvfyW"
# ## Batches and Masking
# + id="Wm9jaJ7evfyX"
class Batch:
"""Object for holding a batch of data with mask during training.
Input is a batch from a torch text iterator.
"""
def __init__(self, src, trg, pad_index=0):
src, src_lengths = src
self.src = src
self.src_lengths = src_lengths
self.src_mask = (src != pad_index).unsqueeze(-2)
self.nseqs = src.size(0)
self.trg = None
self.trg_y = None
self.trg_mask = None
self.trg_lengths = None
self.ntokens = None
if trg is not None:
trg, trg_lengths = trg
self.trg = trg[:, :-1]
self.trg_lengths = trg_lengths
self.trg_y = trg[:, 1:]
self.trg_mask = (self.trg_y != pad_index)
self.ntokens = (self.trg_y != pad_index).data.sum().item()
if USE_CUDA:
self.src = self.src.cuda()
self.src_mask = self.src_mask.cuda()
if trg is not None:
self.trg = self.trg.cuda()
self.trg_y = self.trg_y.cuda()
self.trg_mask = self.trg_mask.cuda()
# + [markdown] id="FVGJEE5evfyd"
# ## Training Loop
# The code below trains the model for 1 epoch (=1 pass through the training data).
# + id="YaYY7--jvfye"
def run_epoch(data_iter, model, loss_compute, print_every=50):
"""Standard Training and Logging Function"""
start = time.time()
total_tokens = 0
total_loss = 0
print_tokens = 0
for i, batch in enumerate(data_iter, 1):
out, _, pre_output = model.forward(batch.src, batch.trg,
batch.src_mask, batch.trg_mask,
batch.src_lengths, batch.trg_lengths)
loss = loss_compute(pre_output, batch.trg_y, batch.nseqs)
total_loss += loss
total_tokens += batch.ntokens
print_tokens += batch.ntokens
if model.training and i % print_every == 0:
elapsed = time.time() - start
print("Epoch Step: %d Loss: %f Tokens per Sec: %f" %
(i, loss / batch.nseqs, print_tokens / elapsed))
start = time.time()
print_tokens = 0
return math.exp(total_loss / float(total_tokens))
# + [markdown] id="I2QGCxvBvfyk"
# ## Training Data and Batching
#
# We will use torch text for batching. This is discussed in more detail below.
# + [markdown] id="zt0Mwd5mvfyl"
# ## Optimizer
#
# We will use the [Adam optimizer](https://arxiv.org/abs/1412.6980) with default settings ($\beta_1=0.9$, $\beta_2=0.999$ and $\epsilon=10^{-8}$).
#
# We will use $0.0003$ as the learning rate here, but for different problems another learning rate may be more appropriate. You will have to tune that.
# + [markdown] id="cCmK9lnjvfym"
# # A First Example
#
# We can begin by trying out a simple copy-task. Given a random set of input symbols from a small vocabulary, the goal is to generate back those same symbols.
# + [markdown] id="IrqJCXrHvfyo"
# ## Synthetic Data
# + id="m7fIAGxFvfyp"
def data_gen(num_words=11, batch_size=16, num_batches=100, length=10, pad_index=0, sos_index=1):
"""Generate random data for a src-tgt copy task."""
for i in range(num_batches):
data = torch.from_numpy(
np.random.randint(1, num_words, size=(batch_size, length)))
data[:, 0] = sos_index
data = data.cuda() if USE_CUDA else data
src = data[:, 1:]
trg = data
src_lengths = [length-1] * batch_size
trg_lengths = [length] * batch_size
yield Batch((src, src_lengths), (trg, trg_lengths), pad_index=pad_index)
# + [markdown] id="FfSSxnGavfyv"
# ## Loss Computation
# + id="QgDt4-CCvfyx"
class SimpleLossCompute:
"""A simple loss compute and train function."""
def __init__(self, generator, criterion, opt=None):
self.generator = generator
self.criterion = criterion
self.opt = opt
def __call__(self, x, y, norm):
x = self.generator(x)
loss = self.criterion(x.contiguous().view(-1, x.size(-1)),
y.contiguous().view(-1))
loss = loss / norm
if self.opt is not None:
loss.backward()
self.opt.step()
self.opt.zero_grad()
return loss.data.item() * norm
# + [markdown] id="OMRWNmIlvfy5"
# ### Printing examples
#
# To monitor progress during training, we will translate a few examples.
#
# We use greedy decoding for simplicity; that is, at each time step, starting at the first token, we choose the one with that maximum probability, and we never revisit that choice.
# + id="B4VOA0xHvfy6"
def greedy_decode(model, src, src_mask, src_lengths, max_len=100, sos_index=1, eos_index=None):
"""Greedily decode a sentence."""
with torch.no_grad():
encoder_hidden, encoder_final = model.encode(src, src_mask, src_lengths)
prev_y = torch.ones(1, 1).fill_(sos_index).type_as(src)
trg_mask = torch.ones_like(prev_y)
output = []
attention_scores = []
hidden = None
for i in range(max_len):
with torch.no_grad():
out, hidden, pre_output = model.decode(
encoder_hidden, encoder_final, src_mask,
prev_y, trg_mask, hidden)
# we predict from the pre-output layer, which is
# a combination of Decoder state, prev emb, and context
prob = model.generator(pre_output[:, -1])
_, next_word = torch.max(prob, dim=1)
next_word = next_word.data.item()
output.append(next_word)
prev_y = torch.ones(1, 1).type_as(src).fill_(next_word)
attention_scores.append(model.decoder.attention.alphas.cpu().numpy())
output = np.array(output)
# cut off everything starting from </s>
# (only when eos_index provided)
if eos_index is not None:
first_eos = np.where(output==eos_index)[0]
if len(first_eos) > 0:
output = output[:first_eos[0]]
return output, np.concatenate(attention_scores, axis=1)
def lookup_words(x, vocab=None):
if vocab is not None:
x = [vocab.itos[i] for i in x]
return [str(t) for t in x]
# + id="dfC2B-ruvfy_"
def print_examples(example_iter, model, n=2, max_len=100,
sos_index=1,
src_eos_index=None,
trg_eos_index=None,
src_vocab=None, trg_vocab=None):
"""Prints N examples. Assumes batch size of 1."""
model.eval()
count = 0
print()
if src_vocab is not None and trg_vocab is not None:
src_eos_index = src_vocab.stoi[EOS_TOKEN]
trg_sos_index = trg_vocab.stoi[SOS_TOKEN]
trg_eos_index = trg_vocab.stoi[EOS_TOKEN]
else:
src_eos_index = None
trg_sos_index = 1
trg_eos_index = None
for i, batch in enumerate(example_iter):
src = batch.src.cpu().numpy()[0, :]
trg = batch.trg_y.cpu().numpy()[0, :]
# remove </s> (if it is there)
src = src[:-1] if src[-1] == src_eos_index else src
trg = trg[:-1] if trg[-1] == trg_eos_index else trg
result, _ = greedy_decode(
model, batch.src, batch.src_mask, batch.src_lengths,
max_len=max_len, sos_index=trg_sos_index, eos_index=trg_eos_index)
print("Example #%d" % (i+1))
print("Src : ", " ".join(lookup_words(src, vocab=src_vocab)))
print("Trg : ", " ".join(lookup_words(trg, vocab=trg_vocab)))
print("Pred: ", " ".join(lookup_words(result, vocab=trg_vocab)))
print()
count += 1
if count == n:
break
# + [markdown] id="yl6GEREMvfzF"
# ## Training the copy task
# + id="CB7ESFaHvfzG"
def train_copy_task():
"""Train the simple copy task."""
num_words = 11
criterion = nn.NLLLoss(reduction="sum", ignore_index=0)
model = make_model(num_words, num_words, emb_size=32, hidden_size=64)
optim = torch.optim.Adam(model.parameters(), lr=0.0003)
eval_data = list(data_gen(num_words=num_words, batch_size=1, num_batches=100))
dev_perplexities = []
if USE_CUDA:
model.cuda()
for epoch in range(10):
print("Epoch %d" % epoch)
# train
model.train()
data = data_gen(num_words=num_words, batch_size=32, num_batches=100)
run_epoch(data, model,
SimpleLossCompute(model.generator, criterion, optim))
# evaluate
model.eval()
with torch.no_grad():
perplexity = run_epoch(eval_data, model,
SimpleLossCompute(model.generator, criterion, None))
print("Evaluation perplexity: %f" % perplexity)
dev_perplexities.append(perplexity)
print_examples(eval_data, model, n=2, max_len=9)
return dev_perplexities
# + id="t7BQjgdQvfzL" outputId="1706c63b-f92b-4ce8-90cc-61cecb4d6d52" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# train the copy task
dev_perplexities = train_copy_task()
def plot_perplexity(perplexities):
"""plot perplexities"""
plt.title("Perplexity per Epoch")
plt.xlabel("Epoch")
plt.ylabel("Perplexity")
plt.plot(perplexities)
plot_perplexity(dev_perplexities)
# + [markdown] id="YiUMt-XFvfzR"
# You can see that the model managed to correctly 'translate' the two examples in the end.
#
# Moreover, the perplexity of the development data nicely went down towards 1.
# + [markdown] id="6qPnbzBlvfzS"
# # A Real World Example
#
# Now we consider a real-world example using the IWSLT German-English Translation task.
# This task is much smaller than usual, but it illustrates the whole system.
#
# The cell below installs torch text and spacy. This might take a while.
# + id="nQHwLCaJvfza"
# !pip install git+git://github.com/pytorch/text spacy
# !python -m spacy download en
# !python -m spacy download de
# + [markdown] id="kFUK2C-lvfzh"
# ## Data Loading
#
# We will load the dataset using torchtext and spacy for tokenization.
#
# This cell might take a while to run the first time, as it will download and tokenize the IWSLT data.
#
# For speed we only include short sentences, and we include a word in the vocabulary only if it occurs at least 5 times. In this case we also lowercase the data.
#
# If you have **issues** with torch text in the cell below (e.g. an `ascii` error), try running `export LC_ALL="en_US.UTF-8"` before you start `jupyter notebook`.
# + id="K80wCzPJvfzi" outputId="93ddbf9c-5797-423b-eb6d-13bf4521d9ad" colab={"base_uri": "https://localhost:8080/"}
# For data loading.
from torchtext import data, datasets
if True:
import spacy
spacy_de = spacy.load('de')
spacy_en = spacy.load('en')
def tokenize_de(text):
return [tok.text for tok in spacy_de.tokenizer(text)]
def tokenize_en(text):
return [tok.text for tok in spacy_en.tokenizer(text)]
UNK_TOKEN = "<unk>"
PAD_TOKEN = "<pad>"
SOS_TOKEN = "<s>"
EOS_TOKEN = "</s>"
LOWER = True
# we include lengths to provide to the RNNs
SRC = data.Field(tokenize=tokenize_de,
batch_first=True, lower=LOWER, include_lengths=True,
unk_token=UNK_TOKEN, pad_token=PAD_TOKEN, init_token=None, eos_token=EOS_TOKEN)
TRG = data.Field(tokenize=tokenize_en,
batch_first=True, lower=LOWER, include_lengths=True,
unk_token=UNK_TOKEN, pad_token=PAD_TOKEN, init_token=SOS_TOKEN, eos_token=EOS_TOKEN)
MAX_LEN = 25 # NOTE: we filter out a lot of sentences for speed
train_data, valid_data, test_data = datasets.IWSLT.splits(
exts=('.de', '.en'), fields=(SRC, TRG),
filter_pred=lambda x: len(vars(x)['src']) <= MAX_LEN and
len(vars(x)['trg']) <= MAX_LEN)
MIN_FREQ = 5 # NOTE: we limit the vocabulary to frequent words for speed
SRC.build_vocab(train_data.src, min_freq=MIN_FREQ)
TRG.build_vocab(train_data.trg, min_freq=MIN_FREQ)
PAD_INDEX = TRG.vocab.stoi[PAD_TOKEN]
# + [markdown] id="6H8AzNjpvfzp"
# ### Let's look at the data
#
# It never hurts to look at your data and some statistics.
# + id="xb-YM_3mvfzq" outputId="d847b573-9496-4ae4-b540-dd1d54d5a3ff" colab={"base_uri": "https://localhost:8080/"}
def print_data_info(train_data, valid_data, test_data, src_field, trg_field):
""" This prints some useful stuff about our data sets. """
print("Data set sizes (number of sentence pairs):")
print('train', len(train_data))
print('valid', len(valid_data))
print('test', len(test_data), "\n")
print("First training example:")
print("src:", " ".join(vars(train_data[0])['src']))
print("trg:", " ".join(vars(train_data[0])['trg']), "\n")
print("Most common words (src):")
print("\n".join(["%10s %10d" % x for x in src_field.vocab.freqs.most_common(10)]), "\n")
print("Most common words (trg):")
print("\n".join(["%10s %10d" % x for x in trg_field.vocab.freqs.most_common(10)]), "\n")
print("First 10 words (src):")
print("\n".join(
'%02d %s' % (i, t) for i, t in enumerate(src_field.vocab.itos[:10])), "\n")
print("First 10 words (trg):")
print("\n".join(
'%02d %s' % (i, t) for i, t in enumerate(trg_field.vocab.itos[:10])), "\n")
print("Number of German words (types):", len(src_field.vocab))
print("Number of English words (types):", len(trg_field.vocab), "\n")
print_data_info(train_data, valid_data, test_data, SRC, TRG)
# + [markdown] id="9BOSgMnWvfz5"
# ## Iterators
# Batching matters a ton for speed. We will use torch text's BucketIterator here to get batches containing sentences of (almost) the same length.
#
# #### Note on sorting batches for RNNs in PyTorch
#
# For effiency reasons, PyTorch RNNs require that batches have been sorted by length, with the longest sentence in the batch first. For training, we simply sort each batch.
# For validation, we would run into trouble if we want to compare our translations with some external file that was not sorted. Therefore we simply set the validation batch size to 1, so that we can keep it in the original order.
# + id="5ZNVkuo9vfz6" outputId="03501af4-91e6-44f8-f994-5485b23a4a37" colab={"base_uri": "https://localhost:8080/"}
train_iter = data.BucketIterator(train_data, batch_size=64, train=True,
sort_within_batch=True,
sort_key=lambda x: (len(x.src), len(x.trg)), repeat=False,
device=DEVICE)
valid_iter = data.Iterator(valid_data, batch_size=1, train=False, sort=False, repeat=False,
device=DEVICE)
def rebatch(pad_idx, batch):
"""Wrap torchtext batch into our own Batch class for pre-processing"""
return Batch(batch.src, batch.trg, pad_idx)
# + [markdown] id="VETFt-2rvf0A"
# ## Training the System
#
# Now we train the model.
#
# On a Titan X GPU, this runs at ~18,000 tokens per second with a batch size of 64.
# + id="xpqZ29IRvf0B"
def train(model, num_epochs=10, lr=0.0003, print_every=100):
"""Train a model on IWSLT"""
if USE_CUDA:
model.cuda()
# optionally add label smoothing; see the Annotated Transformer
criterion = nn.NLLLoss(reduction="sum", ignore_index=PAD_INDEX)
optim = torch.optim.Adam(model.parameters(), lr=lr)
dev_perplexities = []
for epoch in range(num_epochs):
print("Epoch", epoch)
model.train()
train_perplexity = run_epoch((rebatch(PAD_INDEX, b) for b in train_iter),
model,
SimpleLossCompute(model.generator, criterion, optim),
print_every=print_every)
model.eval()
with torch.no_grad():
print_examples((rebatch(PAD_INDEX, x) for x in valid_iter),
model, n=3, src_vocab=SRC.vocab, trg_vocab=TRG.vocab)
dev_perplexity = run_epoch((rebatch(PAD_INDEX, b) for b in valid_iter),
model,
SimpleLossCompute(model.generator, criterion, None))
print("Validation perplexity: %f" % dev_perplexity)
dev_perplexities.append(dev_perplexity)
return dev_perplexities
# + id="NtIVSGCnvf0H" outputId="264452f7-f218-4d61-a6d8-c10199d5bede" colab={"base_uri": "https://localhost:8080/"}
model = make_model(len(SRC.vocab), len(TRG.vocab),
emb_size=256, hidden_size=256,
num_layers=1, dropout=0.2)
dev_perplexities = train(model, print_every=100)
# + id="ZQzd2aZevf0M" outputId="1febe468-5067-490c-a6f2-18089bbfff54" colab={"base_uri": "https://localhost:8080/", "height": 294}
plot_perplexity(dev_perplexities)
# + [markdown] id="0YFtzxNKwKOf"
# ## Save Model
# + id="b6MWxdtPwJhs"
cpu_model = model.to('cpu')
torch.save(cpu_model.state_dict(), 'attention.pt')
# + [markdown] id="0DjsF7FCyiyG"
# Save model metadata
# + id="gSATK2I6ygFP"
import pickle
from collections import defaultdict
stoi = defaultdict(int)
for s, i in SRC.vocab.stoi.items():
if s == UNK_TOKEN or i != 0:
stoi[s] = i
with open('attention_metadata.pkl', 'wb') as f:
metadata = {
'src_stoi': stoi,
'target_itos': TRG.vocab.itos,
'unk_token': UNK_TOKEN,
'pad_token': PAD_TOKEN,
'sos_token': SOS_TOKEN,
'eos_token': EOS_TOKEN,
'src_eos_index': SRC.vocab.stoi[EOS_TOKEN],
'trg_eos_index': TRG.vocab.stoi[EOS_TOKEN],
'trg_sos_index': TRG.vocab.stoi[SOS_TOKEN],
}
pickle.dump(metadata, f)
# + id="_ntjJAXY0z6h"
# !cp attention_metadata.pkl '/content/drive/My Drive/'
# + [markdown] id="o69nzQfhvf0R"
# ## Prediction and Evaluation
#
# Once trained we can use the model to produce a set of translations.
#
# If we translate the whole validation set, we can use [SacreBLEU](https://github.com/mjpost/sacreBLEU) to get a [BLEU score](https://en.wikipedia.org/wiki/BLEU), which is the most common way to evaluate translations.
#
# #### Important sidenote
# Typically you would use SacreBLEU from the **command line** using the output file and original (possibly tokenized) development reference file. This will give you a nice version string that shows how the BLEU score was calculated; for example, if it was lowercased, if it was tokenized (and how), and what smoothing was used. If you want to learn more about how BLEU scores are (and should be) reported, check out [this paper](https://arxiv.org/abs/1804.08771).
#
# However, right now our pre-processed data is only in memory, so we'll calculate the BLEU score right from this notebook for demonstration purposes.
#
# We'll first test the raw BLEU function:
# + id="dtGASOyCvf0S"
import sacrebleu
# + id="BO-yxjnRvf0X" outputId="f4d2aecc-316d-453e-e8d2-8b610128ff41" colab={"base_uri": "https://localhost:8080/"}
# this should result in a perfect BLEU of 100%
hypotheses = ["this is a test"]
references = ["this is a test"]
bleu = sacrebleu.raw_corpus_bleu(hypotheses, [references], .01).score
print(bleu)
# + id="zOBhseHqvf0b" outputId="85170984-abee-44d3-9d60-2161c07a4817" colab={"base_uri": "https://localhost:8080/"}
# here the BLEU score will be lower, because some n-grams won't match
hypotheses = ["this is a test"]
references = ["this is a fest"]
bleu = sacrebleu.raw_corpus_bleu(hypotheses, [references], .01).score
print(bleu)
# + [markdown] id="31jlkaaMvf0f"
# Since we did some filtering for speed, our validation set contains 690 sentences.
# The references are the tokenized versions, but they should not contain out-of-vocabulary UNKs that our network might have seen. So we'll take the references straight out of the `valid_data` object:
# + id="LivuByjivf0g" outputId="6e1e0120-705e-4b8c-f9a0-3ce4e8550f71" colab={"base_uri": "https://localhost:8080/"}
len(valid_data)
# + id="n9m0YsGhvf0k" outputId="367b9e82-3289-4c6d-f3bc-08295971cb29" colab={"base_uri": "https://localhost:8080/"}
references = [" ".join(example.trg) for example in valid_data]
print(len(references))
print(references[0])
# + id="x4uec4wqvf0o" outputId="0090d5f3-a57c-4d50-c108-adb3aef4037e" colab={"base_uri": "https://localhost:8080/", "height": 34}
references[-2]
# + [markdown] id="tR9KmVkfvf0t"
# **Now we translate the validation set!**
#
# This might take a little bit of time.
#
# Note that `greedy_decode` will cut-off the sentence when it encounters the end-of-sequence symbol, if we provide it the index of that symbol.
# + id="hOwxAMsPvf0u" outputId="0ae097dc-0ac7-462c-9d51-4dffedce740d" colab={"base_uri": "https://localhost:8080/"}
hypotheses = []
alphas = [] # save the last attention scores
for batch in valid_iter:
batch = rebatch(PAD_INDEX, batch)
pred, attention = greedy_decode(
model, batch.src, batch.src_mask, batch.src_lengths, max_len=25,
sos_index=TRG.vocab.stoi[SOS_TOKEN],
eos_index=TRG.vocab.stoi[EOS_TOKEN])
hypotheses.append(pred)
alphas.append(attention)
# + id="3GVqhs0Hvf0_" outputId="b0abba86-ec39-4096-9a86-d9a079f6c112" colab={"base_uri": "https://localhost:8080/"}
# we will still need to convert the indices to actual words!
hypotheses[0]
# + id="DYGSSXYHvf1D" outputId="6a6fe217-d992-4f72-ea63-3ed4269f4d7f" colab={"base_uri": "https://localhost:8080/"}
hypotheses = [lookup_words(x, TRG.vocab) for x in hypotheses]
hypotheses[0]
# + id="Sz4WvT58vf1G" outputId="4996be97-c418-41ae-9224-de99ddafd729" colab={"base_uri": "https://localhost:8080/"}
# finally, the SacreBLEU raw scorer requires string input, so we convert the lists to strings
hypotheses = [" ".join(x) for x in hypotheses]
print(len(hypotheses))
print(hypotheses[0])
# + id="HHof7L5Svf1J" outputId="b65dd6e4-e792-4f86-d837-467f9413e7d2" colab={"base_uri": "https://localhost:8080/"}
# now we can compute the BLEU score!
bleu = sacrebleu.raw_corpus_bleu(hypotheses, [references], .01).score
print(bleu)
# + [markdown] id="aNdx-ToZvf1M"
# ## Attention Visualization
#
# We can also visualize the attention scores of the decoder.
# + id="YlHeoiMFvf1O"
def plot_heatmap(src, trg, scores):
fig, ax = plt.subplots()
heatmap = ax.pcolor(scores, cmap='viridis')
ax.set_xticklabels(trg, minor=False, rotation='vertical')
ax.set_yticklabels(src, minor=False)
# put the major ticks at the middle of each cell
# and the x-ticks on top
ax.xaxis.tick_top()
ax.set_xticks(np.arange(scores.shape[1]) + 0.5, minor=False)
ax.set_yticks(np.arange(scores.shape[0]) + 0.5, minor=False)
ax.invert_yaxis()
plt.colorbar(heatmap)
plt.show()
# + id="f8Edr0Gmvf1S" outputId="f07fd427-0b93-4906-f55b-b4b8f4896566" colab={"base_uri": "https://localhost:8080/", "height": 337}
# This plots a chosen sentence, for which we saved the attention scores above.
idx = 5
src = valid_data[idx].src + ["</s>"]
trg = valid_data[idx].trg + ["</s>"]
pred = hypotheses[idx].split() + ["</s>"]
pred_att = alphas[idx][0].T[:, :len(pred)]
print("src", src)
print("ref", trg)
print("pred", pred)
plot_heatmap(src, pred, pred_att)
# + [markdown] id="7hIOugRwvf1X"
# # Congratulations! You've finished this notebook.
#
# What didn't we cover?
#
# - Subwords / Byte Pair Encoding [[paper]](https://arxiv.org/abs/1508.07909) [[github]](https://github.com/rsennrich/subword-nmt) let you deal with unknown words.
# - You can implement a [multiplicative/bilinear attention mechanism](https://arxiv.org/abs/1508.04025) instead of the additive one used here.
# - We used greedy decoding here to get translations, but you can get better results with beam search.
# - The original model only uses a single dropout layer (in the decoder), but you can experiment with adding more dropout layers, for example on the word embeddings and the source word representations.
# - You can experiment with multiple encoder/decoder layers.- Experiment with a benchmarked and improved codebase: [Joey NMT](https://github.com/joeynmt/joeynmt)
# + [markdown] id="lzTgEn1dvf1X"
# If this was useful to your research, please consider citing:
#
# > <NAME>. 2018. The Annotated Encoder-Decoder with Attention. https://bastings.github.io/annotated_encoder_decoder/
#
# Or use the following `Bibtex`:
# ```
# @misc{bastings2018annotated,
# title={The Annotated Encoder-Decoder with Attention},
# author={<NAME>.},
# journal={https://bastings.github.io/annotated\_encoder\_decoder/},
# year={2018}
# }```
|
11 - Attention Mechanism/annotated_encoder_decoder.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mbj5335/hashcat-utils/blob/master/Correlation%20coefficient%20between%20sleep%20and%20long-term%20memory.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="cSlb-2dCXph_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 877} outputId="aec6b127-e7a4-422d-c8e7-a7ca02b253c4"
# !pip install datascience
# + id="lBp85gzaXub9" colab_type="code" colab={}
import numpy as np
from datascience import *
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use("fivethirtyeight")
import warnings
warnings.simplefilter('ignore', FutureWarning)
# + id="FLvxeaKCYUas" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 56} outputId="e8b47bb6-fdd1-41ee-88e5-c28e74669c25"
from google.colab import drive
drive.mount('/content/drive')
# + id="MiH0a9enYqlJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="c8a6e224-5431-4874-f5c6-3db1271751ab"
# !ls /content/drive/My\ Drive/DS200Labs
# + id="jxaU7CUQY8Bj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 385} outputId="5f9f1051-6f0c-4295-ad35-e038875f66d9"
path1 = "/content/drive/My Drive/DS200Labs/sleep_con.csv"
Sleep = Table.read_table(path1)
Sleep.show(10)
# + id="__P5fJxSZG8X" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="3bc9fbc3-7e28-47e8-cbf4-edbc2128747a"
Sleep.scatter('s-time', 'long-term memory', fit_line=True)
# + id="PnRFwCtZcjE_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="2bbc8a8b-ee6d-48bd-e196-7cabc7660889"
Sleep_con = Sleep.to_df()
Sleep_con.shape
# + id="XgN_fcTOc18z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 110} outputId="39e3e664-e056-4073-ac99-50458b794908"
Sleep_con.corr()
|
Correlation coefficient between sleep and long-term memory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.11 ('base2')
# language: python
# name: python3
# ---
# + [markdown] id="_rPjOk7xbe9A"
# The whold training and testing is on Google Colab.
# + colab={"base_uri": "https://localhost:8080/"} id="Z6vkptkhj7Lh" outputId="48480b9b-e1b1-4b31-8cdd-d0a5910a0ca0"
import os
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="UZf_l_SGkIZb" outputId="4895cc2f-8272-4c29-a744-56147d15aef3"
path = "/content/drive/My Drive/segnet/"
os.chdir(path)
os.listdir(path)
# + colab={"base_uri": "https://localhost:8080/"} id="M9XFtbHkazqh" outputId="2475fa75-1754-4afa-cd6a-c921dcd86cc7"
# !/opt/bin/nvidia-smi
# + id="-ewA8Mf4drRM" colab={"base_uri": "https://localhost:8080/"} outputId="4f3d95a6-59c7-4da9-84ad-c170106a4387"
# use this version to avoid bugs
# !pip install torch-summary==1.4.4
# + [markdown] id="pwHM-6uBjX3k"
# # 0. parameters
# ## 0.parameters
# + id="jQSaRIwojX3l"
import torch.utils.data as data
import torch
import numpy as np
import h5py
import matplotlib.pyplot as plt
# %matplotlib inline
import warnings
warnings.filterwarnings('ignore')
from tqdm import tqdm
import torch.nn as nn
from torch.nn import init
import torch.nn.functional as F
import random
import cv2 as cv
from PIL import Image
import time
# + id="108R3g06jX3n"
device = 'cuda' if torch.cuda.is_available() else 'cpu'
MODE = 'TRAIN' # set MODE='TEST' to test without training
n_epochs = 10 # training epochs
class_num = 34
batch_size = 24 # should be no less than 5
learning_rate = 2e-5
weight_decay = 5e-4
log_interval = 10
random_seed = 28
val_percent = 0.1 # training set : validation set = 9:1
torch.manual_seed(random_seed)
bn_momentum = 0.1 # momentum for batch normalization
cate_weight = [1/34]*34 # weight for each class
dir_pre_train_weights = "vgg16_bn-6c64b313.pth" # pre_train weights downloaded from https://download.pytorch.org/models/vgg16_bn-6c64b313.pth
dir_weights = "./weights"
dir_checkpoint = './checkpoints'
# + [markdown] id="PNogkTLqjX3o"
# # 1.Implement a data loader class to handle the downloaded data. (5 points)
# For more information on the dataset please refer to: CityScapes dataset.
# + id="sNDPQBRsSOBH"
color_codes = h5py.File("lab2_test_data.h5", 'r')['color_codes']
# + id="bDrWorCujX3o"
# 'rgb' stores the raw images, while 'seg' stores segmentation maps
class DataFromH5File(data.Dataset):
def __init__(self, filepath):
h5File = h5py.File(filepath, 'r')
# self.color_codes = h5File['color_codes']
self.rgb = h5File['rgb']
self.seg = h5File['seg']
def __getitem__(self, idx):
label = torch.from_numpy(self.seg[idx]).float()
data = torch.from_numpy(self.rgb[idx]).float()
data = data/255.0 # normalization
data = data.permute(2,0,1) # change the image channels into (channel, width, height)
return data, label
def __len__(self):
assert self.rgb.shape[0] == self.seg.shape[0], "Wrong data length" # robustness
return self.rgb.shape[0]
# + id="aUoAgwEPjX3p"
# load training data from lab2_train_data.h5
dataset = DataFromH5File("lab2_train_data.h5")
n_val = int(len(dataset) * val_percent)
n_train = len(dataset) - n_val
# split train & val
train, val = data.random_split(dataset, [n_train, n_val])
train_loader = data.DataLoader(dataset=train, batch_size=batch_size, shuffle=True, pin_memory=True, drop_last = True)
val_loader = data.DataLoader(dataset=val, batch_size=batch_size, shuffle=False, pin_memory=True, drop_last = True) # drop_last=True
# load testing data from lab2_test_data.h5
testset = DataFromH5File("lab2_test_data.h5")
test_loader = data.DataLoader(dataset=testset, batch_size=batch_size, shuffle=False, pin_memory=True)
# + colab={"base_uri": "https://localhost:8080/"} id="Zvzb0C0ojX3p" outputId="8c04fb59-0a89-4c1e-8376-7d64f9da9d94"
# test the data loader
for step, (x, y) in enumerate(train_loader):
print(x.min(),x.max())
print(y.min(),y.max())
print(step)
break
print(len(train_loader), len(val_loader), len(test_loader)) # 669 74 125 when batch_size==4
# + [markdown] id="SVPNv7I1jX3q"
# # 2. Define the model. Provide a schematic of your architecture depicting its overall structure and the relevant parameters. (20 points)
# + [markdown] id="6whmGCmRdQmc"
# ## 2.1 Define the model.
# Badrinarayanan et al. proposed **SegNet**.
# Paper link:https://arxiv.org/pdf/1511.00561.pdf
# Kendall et al. made some improvements and proposed **Bayesian SegNet**.
# Paper link:https://arxiv.org/pdf/1511.02680.pdf
# + id="Td8VznZKaq-1"
# encoder
class Encoder(nn.Module):
def __init__(self, input_channels):
super(Encoder, self).__init__()
self.enco1 = nn.Sequential(
nn.Conv2d(input_channels, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(64, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(64, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(64, momentum=bn_momentum),
nn.ReLU()
)
self.enco2 = nn.Sequential(
nn.Conv2d(64, 128, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(128, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(128, 128, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(128, momentum=bn_momentum),
nn.ReLU()
)
self.enco3 = nn.Sequential(
nn.Conv2d(128, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU()
)
self.enco4 = nn.Sequential(
nn.Conv2d(256, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU()
)
self.enco5 = nn.Sequential(
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU()
)
self.dropout = nn.Dropout(p=0.5)
def forward(self, x):
id = []
x = self.enco1(x)
x, id1 = F.max_pool2d(x, kernel_size=2, stride=2, return_indices=True)
id.append(id1)
x = self.enco2(x)
x, id2 = F.max_pool2d(x, kernel_size=2, stride=2, return_indices=True)
id.append(id2)
x = self.enco3(x)
x, id3 = F.max_pool2d(x, kernel_size=2, stride=2, return_indices=True)
id.append(id3)
x = self.dropout(x)
x = self.enco4(x)
x, id4 = F.max_pool2d(x, kernel_size=2, stride=2, return_indices=True)
id.append(id4)
x = self.dropout(x)
x = self.enco5(x)
x, id5 = F.max_pool2d(x, kernel_size=2, stride=2, return_indices=True)
id.append(id5)
x = self.dropout(x)
return x, id
# encoder + decoder
class BayesianSegNet(nn.Module):
def __init__(self, input_channels, output_channels):
super(BayesianSegNet, self).__init__()
self.weights_new = self.state_dict()
self.encoder = Encoder(input_channels)
self.deco1 = nn.Sequential(
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU()
)
self.deco2 = nn.Sequential(
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 512, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(512, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(512, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU()
)
self.deco3 = nn.Sequential(
nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(256, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(256, 128, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(128, momentum=bn_momentum),
nn.ReLU()
)
self.deco4 = nn.Sequential(
nn.Conv2d(128, 128, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(128, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(128, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(64, momentum=bn_momentum),
nn.ReLU()
)
self.deco5 = nn.Sequential(
nn.Conv2d(64, 64, kernel_size=3, stride=1, padding=1),
nn.BatchNorm2d(64, momentum=bn_momentum),
nn.ReLU(),
nn.Conv2d(64, output_channels, kernel_size=3, stride=1, padding=1),
nn.Softmax() # softmax
)
self.dropout = nn.Dropout(p=0.5)
def forward(self, x):
x, id = self.encoder(x)
x = F.max_unpool2d(x, id[4], kernel_size=2, stride=2)
x = self.deco1(x)
x = self.dropout(x)
x = F.max_unpool2d(x, id[3], kernel_size=2, stride=2)
x = self.deco2(x)
x = self.dropout(x)
x = F.max_unpool2d(x, id[2], kernel_size=2, stride=2)
x = self.deco3(x)
x = self.dropout(x)
x = F.max_unpool2d(x, id[1], kernel_size=2, stride=2)
x = self.deco4(x)
x = F.max_unpool2d(x, id[0], kernel_size=2, stride=2)
x = self.deco5(x)
return x
# delete weights of three fc layers
def load_weights(self, weights_path):
weights = torch.load(weights_path)
del weights["classifier.0.weight"]
del weights["classifier.0.bias"]
del weights["classifier.3.weight"]
del weights["classifier.3.bias"]
del weights["classifier.6.weight"]
del weights["classifier.6.bias"]
names = []
for key, value in self.encoder.state_dict().items():
if "num_batches_tracked" in key:
continue
names.append(key)
for name, dict in zip(names, weights.items()):
self.weights_new[name] = dict[1]
self.encoder.load_state_dict(self.weights_new)
# + [markdown] id="u558tpeDfnAd"
# ## 2.2 Provide a schematic of your architecture depicting its overall structure and the relevant parameters.
#
# + [markdown] id="Q1qAOBBmgF2L"
# 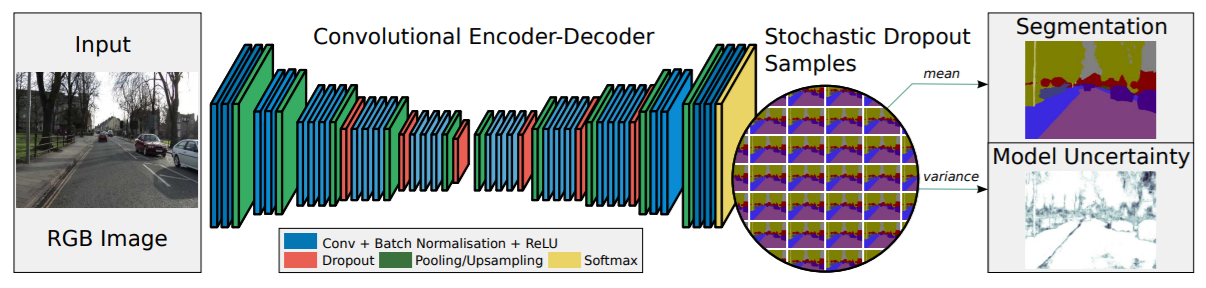
# $$\rm Figure 1.model\; architecture$$
# $$\rm As\; shown\; above,\; the\; encoder\; of\; Bayesian\; SegNet\; is\; the\; same\; as\; VGG16\; without\; fc\; layers.\; Dropout\; is\; added\; to\; the\; last\; three\; convolution\; layers\; of\; the\; encoder,\; and\; the\; first\; three\; convolution\; layers\; of\; the\; decoder.$$
# $$
# \rm Note:Kendall,\; A.,\; Badrinarayanan,\; V.,\; \&\; Cipolla,\; R.\; (2015). Bayesian\; segnet:\; Model\; uncertainty\; in\; deep\; convolutional\; encoder-decoder\; architectures\; for\; scene\; understanding.\; arXiv\; preprint\; arXiv:1511.02680.
# $$
#
# + id="dKVzHiXfaq-2"
from torchsummary import summary
# + colab={"base_uri": "https://localhost:8080/"} id="nfDZiuiWaq-3" outputId="83b6fb80-ce72-415c-bbc0-3c1536ca00b8"
model = BayesianSegNet(input_channels=3, output_channels=class_num) # RGB images so the input_channels=3
model = model.to(device)
x = torch.ones([batch_size, 3, 128, 256]) # input shape
x = x.to(device)
y = model(x)
print(y.shape) # output shape
summary(model, input_size=(3, 128, 256))
# + [markdown] id="5rQgQoaujX3s"
# # 3. Define the loss function and optimizer. (10 points)
# + id="Uo-o330bjX3s"
import torch.optim as optim
# + id="uL9dJHSyjX3t"
optimizer = optim.Adam(model.parameters(), lr=learning_rate, weight_decay=weight_decay)
# cross entropy loss
# To cope with the sample imbalance between different categories, we assign different weights to them.
criterion = nn.CrossEntropyLoss(weight=torch.from_numpy(np.array(cate_weight)).float()).cuda()
# + [markdown] id="pXUulvFDjX3t"
# # 4. Train the network. (5 points)
# + id="OyWKrRJEjX3u"
train_losses = []
train_counter = []
test_losses = []
test_counter = [i*len(train_loader.dataset) for i in range(n_epochs + 1)]
# + id="TSvq6hbierWu"
model.load_weights(dir_pre_train_weights)
# + [markdown] id="9sKZjx4o3ied"
# The segmentation challenge is evaluated using the mean Intersection over Union (mIoU) metric.
# Let $n$ denotes the number of classes, then
# $$
# mIoU = \frac{TP}{TP + FP + FN}
# =\frac{1}{n}\sum_{i = 1}^{n}\frac{p_{ii}}{\sum_{j=1}^{n}p_{ij}+\sum_{j=1}^{n}p_{ji}+p_{ii}}
# $$
# + id="rLNZINSW3L20"
def mIoU(pred, target):
mini = 1
intersection = pred * (pred == target)
# histogram
area_inter, _ = np.histogram(intersection, bins=2, range=(mini, class_num))
area_pred, _ = np.histogram(pred, bins=2, range=(mini, class_num))
area_target, _ = np.histogram(target, bins=2, range=(mini, class_num))
area_union = area_pred + area_target - area_inter
# Intersection area should be smaller than Union area
assert (area_inter <= area_union).all(), "Intersection area should be smaller than Union area"
rate = round(max(area_inter) / max(area_union), 4)
return rate
# + id="QsOg2c9JshYZ"
def validate(epoch):
val_pbar = tqdm(val_loader)
for batch_idx, (data, target) in enumerate(val_pbar):
output = model(data.to(device)).to('cpu') # np.histogram requires cpu type tensor
target = target.squeeze().long()
miou = mIoU(output.argmax(dim=1), target) # data.argmax(dim=1) represents the segmentation results
val_pbar.set_description(f"Validation | Epoch: {epoch} | mIoU: {miou.item():.4f}")
# + id="6ZmMxJ9MjX3u"
def train(epoch):
model.train()
pbar = tqdm(train_loader)
for batch_idx, (data, target) in enumerate(pbar):
data = data.to(device)
target = target.to(device)
optimizer.zero_grad()
output = model(data)
target = target.squeeze().long()
# print('output shape=',output.shape)
# print('target shape=',target.shape)
loss = criterion(output, target)
loss.backward()
optimizer.step()
pbar.set_description(f"Epoch: {epoch} | Loss: {loss.item():.4f}")
if batch_idx % log_interval == 0:
train_losses.append(loss.item())
train_counter.append((batch_idx*batch_size) + ((epoch-1)*len(train_loader.dataset)))
# save the parameters
torch.save(model.state_dict(), '/content/drive/My Drive/segnet/weights/model.pth')
torch.save(optimizer.state_dict(), '/content/drive/My Drive/segnet/weights/optimizer.pth')
# + colab={"base_uri": "https://localhost:8080/"} id="Kkg5b6b-DnW5" outputId="e8ee771d-1d68-462a-bc2d-c56c39d4b55e"
if MODE == 'TEST':
model.load_state_dict(torch.load('/content/drive/My Drive/segnet/weights/23_model.pth'))
else:
for epoch in range(1, n_epochs + 1):
train(epoch)
# validation
if epoch % 4 == 0 or epoch == n_epochs: # or epoch==1:
validate(epoch)
# + [markdown] id="AOHqfHvTubHJ"
# plots of the loss evolution
# + [markdown] id="AxJmbMaWjX3u"
# # 5. Test the resulting network on examples from an independent test set. Implement and present: (40 points)
# a. Predictions for (μ, aleatoric, epistemic) .
# b. Visualizations for (μ, aleatoric, epistemic) on 5 different input examples.
# c. Comment briefly on how the model’s performance could be improved.
# d. Please save your code and results for submission.
# + [markdown] id="y0iw03VJj_au"
# ## 5.1 Predictions and visualizations for $(\mu,aleatoric,epistemic)$
# + id="7dp00qiWhvRw"
from scipy.stats import entropy
# + id="nfWi2BfbJWgQ"
# color the img according to the color_codes with elegant coding
def color(src):
ret = np.zeros((src.shape[0], src.shape[1], 3))
for i in range(class_num):
ret[src==i] = color_codes[i]
return ret
# visualize the segmentation results of 5 random test samples
def visualize():
rand_idx = random.randint(0,len(test_loader)-1) # random test example idx
for batch_idx, (data, target) in enumerate(test_loader):
if batch_idx == rand_idx:
data = data.to(device)
n_samples = 20 # number of bayesian dropout samples
for i in range(5):
outputs = torch.Tensor([model(data[i].unsqueeze(0)).to('cpu').detach().numpy() for _ in range(n_samples)])
output = torch.mean(outputs, 0).squeeze() # mu
aleatoric = entropy(output*255, axis=0)
episdemic = torch.std(outputs, axis=0).squeeze().mean(0)
output = output.argmax(0)
f, ax = plt.subplots(1, 5, figsize=(15,15))
for _ in range(5):
ax[_].axis('off')
ax[0].set_title('Input Image') #set titles for each parts
ax[0].imshow(data[i].permute(1,2,0).cpu())
ax[1].set_title('Ground Truth')
ax[1].imshow(color(target[i].to('cpu').squeeze())/255.0)
ax[2].set_title('Semantic Segmentation')
ax[2].imshow(color(output)/255.0)
ax[3].set_title('Aleatoric Uncertainty')
ax[3].imshow(aleatoric)
ax[4].set_title('Epidemic Uncertainty')
ax[4].imshow(episdemic)
plt.show()
break
# + colab={"base_uri": "https://localhost:8080/", "height": 527} id="lqa9eS4S8Gwa" outputId="2b363474-f570-441b-98c6-92bd8d9fe7d4"
visualize()
# + [markdown] id="QUqww_9ckoA8"
# ## 5.2 Comment briefly on how the model’s performance could be improved.
# + [markdown] id="D-KqtoaYkuKt"
# At the begining, our team used UNet to do the segmentation. However, i tooks nearly 1.5h to train one epoch, which was unaffordable. After searching information about lightweight models, we select SegNet, which enables us to train one epoch **within minutes**.
# **The advantage of SegNet is that it don't need to save the feature maps of the entire encoder part, but only the pooling index, which greatly saves memory. Additionaly, it does not need to deconvolute or learn during the upsampling phase.**
#
# Here's some strategies to improve the model's performance:
# - Since time is limited, we haven't assign different weight to each class. Considering the imbalaced data distribution, **changing the value of cate_weight** will help.
# - Increase **batch size**(using more GPUs). During training, the loss function **fluctuated** severely due to small batch size.
# - Try a couple of activate functions.
# - Fine-tune parameters in session 0.
# + [markdown] id="b7cmDgG4jX3v"
# # References
# [1] https://blog.csdn.net/shwan_ma/article/details/100012808
# [2] https://blog.csdn.net/oYeZhou/article/details/112270908
# [3] https://blog.csdn.net/qq_32939413/article/details/112117734
# [4] <NAME> et al. “SegNet: A Deep Convolutional Encoder-Decoder Architecture for Image Segmentation.” IEEE transactions on pattern analysis and machine intelligence vol. 39,12 (2017): 2481-2495. doi:10.1109/TPAMI.2016.2644615
# [5] <NAME>., <NAME>., & <NAME>. (2015). Bayesian segnet: Model uncertainty in deep convolutional encoder-decoder architectures for scene understanding. arXiv preprint arXiv:1511.02680.
# + id="jmFXBDZVNb8f"
|
BayesianSegNet/main_bayesian_segnet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 7
# ## Training / Test dataset and Normalization
# <hr/>
# +
import numpy as np
import tensorflow as tf
x_data = [[1, 2, 1], [1, 3, 2], [1, 3, 4], [1, 5, 5], [1, 7, 5], [1, 2, 5], [1, 6, 6], [1, 7, 7]]
y_data = [[0, 0, 1], [0, 0, 1], [0, 0, 1], [0, 1, 0], [0, 1, 0], [0, 1, 0], [1, 0, 0], [1, 0, 0]]
# Evaluation our model using this test dataset
x_test = [[2, 1, 1], [3, 1, 2], [3, 3, 4]]
y_test = [[0, 0, 1], [0, 0, 1], [0, 0, 1]]
# +
X = tf.placeholder(tf.float32, shape=[None,3])
Y = tf.placeholder(tf.float32, shape=[None,3])
W = tf.Variable(tf.random_normal([3,3]))
b = tf.Variable(tf.random_normal([3]))
logits = tf.matmul(X, W) + b
hypothesis = tf.nn.softmax(logits)
cost = tf.nn.softmax_cross_entropy_with_logits(logits=logits, labels=Y)
train = tf.train.GradientDescentOptimizer(learning_rate=0.1).minimize(cost)
prediction = tf.arg_max(hypothesis, 1)
is_correct = tf.equal(prediction, tf.arg_max(Y, 1))
accuracy = tf.reduce_mean(tf.to_float(is_correct))
with tf.Session() as sess:
sess.run(tf.global_variables_initializer())
for step in range(2001):
sess.run(train, feed_dict = {X: x_data, Y: y_data})
if step % 400 == 0 :
print('cost:',sess.run(tf.reduce_mean(-tf.reduce_sum(Y * tf.log(hypothesis), axis=1)), feed_dict={X:x_data,Y:y_data}))
ac, pre = sess.run([accuracy, prediction], feed_dict = {X: x_data,Y: y_data})
print('step:',step,'\naccuracy:',ac* 100,'%\n',pre,'\n')
print('----------------------------------------test------------------------------------------')
print('prediction:', sess.run(prediction, feed_dict={X: x_test}))
print('Accuracy:', sess.run(accuracy, feed_dict={X: x_test, Y: y_test}))
# -
# <hr/>
# ## Non - Normalization
# <hr/>
# +
xy = np.array([[828.659973, 833.450012, 908100, 828.349976, 831.659973],
[823.02002, 828.070007, 1828100, 821.655029, 828.070007],
[819.929993, 824.400024, 1438100, 818.97998, 824.159973],
[816, 820.958984, 1008100, 815.48999, 819.23999],
[819.359985, 823, 1188100, 818.469971, 818.97998],
[819, 823, 1198100, 816, 820.450012],
[811.700012, 815.25, 1098100, 809.780029, 813.669983],
[809.51001, 816.659973, 1398100, 804.539978, 809.559998]])
x_data = xy[:, 0:-1]
y_data = xy[:, [-1]]
X = tf.placeholder(tf.float32, shape=[None,4])
Y = tf.placeholder(tf.float32, shape=[None,1])
W = tf.Variable(tf.random_normal([4,1]))
b = tf.Variable(tf.random_normal([1]))
hypothesis = tf.matmul(X, W) + b
cost = tf.reduce_mean(tf.square(hypothesis - Y))
train = tf.train.GradientDescentOptimizer(learning_rate=1e-5).minimize(cost)
sess = tf.Session()
sess.run(tf.global_variables_initializer())
for step in range(2001):
cost_val, hy_val, _ = sess.run([cost, hypothesis, train], feed_dict={X:x_data, Y:y_data})
print(step, 'cost:',cost_val,'prediction:\n',hy_val)
# -
# <hr/>
# ## Normalized inputs - min-max scale
# <hr/>
#
# ```python
# xy = MinMaxScaler(xy)
#
# # datas changed to between 1 ~ 0
# ```
#
|
Lecture7 Training_Test dataset_Normalization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 02: Primitives
# [Download on GitHub](https://github.com/NumEconCopenhagen/lectures-2021)
#
# [<img src="https://mybinder.org/badge_logo.svg">](https://mybinder.org/v2/gh/NumEconCopenhagen/lectures-2021/master?urlpath=lab/tree/02/Primitives.ipynb)
# 1. [Your first notebook session](#Your-first-notebook-session)
# 2. [Fundamentals](#Fundamentals)
# 3. [Containers](#Containers)
# 4. [Conditionals and loops](#Conditionals-and-loops)
# 5. [Functions](#Functions)
# 6. [Floating point numbers](#Floating-point-numbers)
# 7. [Classes (user-defined types)](#Classes-(user-defined-types))
# 8. [Summary](#Summary)
# 9. [Extra: Iterators](#Extra:-Iterators)
# 10. [Extra: More on functions](#Extra:-More-on-functions)
#
# You will be given an in-depth introduction to the **fundamentals of Python** (objects, variables, operators, classes, methods, functions, conditionals, loops). You learn to discriminate between different **types** such as integers, floats, strings, lists, tuples and dictionaries, and determine whether they are **subscriptable** (slicable) and/or **mutable**. You will learn about **referencing** and **scope**. You will learn a tiny bit about **floating point arithmetics**.
# **Take-away:** This lecture is rather abstract compared to the rest of the course. The central take-away is **a language** to speak about programming in. An overview of the map, later we will study the terrain in detail. It is not about **memorizing**. Almost no code projects begin from scratch, you start by copying in similar code you have written for another project.
# Hopefully, this notebook can later be used as a **reference sheet**. When you are done with the DataCamp courses, read through this notebook, play around with the code, and ask questions if there is stuff you do not understand.
# **Links:**
#
# * **Tutorial:** A more detailed tutorial is provided [here](https://www.python-course.eu/python3_course.php).
# * **Markdown:** All text cells are written in *Markdown*. A guide is provided [here](https://www.markdownguide.org/basic-syntax/).
# <a id="Your-first-notebook-session"></a>
#
# # 1. Your first notebook session
# **Optimally:** You have this notebook open as well on your own computer.
# **Download guide:**
#
# 1. Follow the [installation guide](https://numeconcopenhagen.netlify.com/guides/python-setup/) in detail
# 2. Open VScode
# 3. Pres <kbd>Ctrl</kbd>+<kbd>Shift</kbd>+<kbd>P</kbd>
# 4. Write `git: clone` + <kbd>Enter</kbd>
# 5. Write `https://github.com/NumEconCopenhagen/lectures-2021` + <kbd>Enter</kbd>
# 6. You can always update to the newest version of the code with `git: sync` + <kbd>Enter</kbd>
# 7. Create a copy of the cloned folder, where you work with the code (otherwise you can not sync with updates)
# **PROBLEMS?** Ask your teaching asssistant ASAP.
# **Execution:**
#
# * **Movements**: Arrows and scrolling
# * **Run cell and advance:** <kbd>Shift</kbd>+<kbd>Enter</kbd>
# * **Run cell**: <kbd>Ctrl</kbd>+<kbd>Enter</kbd>
# * **Edit:** <kbd>Enter</kbd>
# * **Toggle sidebar:** <kbd>Ctrl</kbd>+<kbd>B</kbd>
# * **Change to markdown cell:** <kbd>M</kbd>
# * **Change to code cell:** <kbd>Y</kbd>
# <a id="Fundamentals"></a>
#
# # 2. Fundamentals
# All **variables** in Python is a **reference** to an **object** of some **type**.
# ## 2.1 Atomic types
# The most simple types are called **atomic**. Atomic indicates that they cannot be changed - only overwritten.
# **Integers (int):** -3, -2, -1, 0, 1, 2, 3, etc.
# +
x = 1
# variable x references an integer type object with a value of 1
print(type(x)) # prints the type of x
print(x) # prints the value of x
# -
# **Decimal numbers (float)**: 3.14, 2.72, 1.0, etc.
# +
x = 1.2
# variable x references an floating point (decimal number) type object
# with a value of 1.2
print(type(x))
print(x)
# -
# **Strings (str)**: 'abc', '123', 'this is a full sentence', etc.
# +
x = 'abc'
# variable x references a string type opbject
# with a value of 'abc'
print(type(x))
print(x)
# -
# **Note:** Alternatively, use double quotes instead of single quotes.
# +
x = "abc"
# variable x reference a string type opbject
# with a value of 'abc'
print(type(x))
print(x)
# -
# **Booleans (bool)**: True and False
# +
x = True
# variable x reference a boolean type opbject
# with a value of False
print(type(x))
print(x)
# -
# **Atomic types:**
#
# 1. Integers, *int*
# 2. Floating point numbers, *float*
# 3. Strings, *str*
# 4. Booleans, *bool*
# ## 2.2 Type conversion
# Objects of one type can (sometimes) be **converted** into another type.<br>For example, from float to string:
# +
x = 1.2
# variable x references an floating point (decimal number) type object
# with a value of 1.2
y = str(x)
# variable y now references a string type object
# with a value created based on x
print(y,type(y))
# -
# or from float to integer:
# +
x = 2.9
y = int(x)
# variable x now references an integer type object
# with a value created based on x (here rounded down)
print(y,type(y))
# -
# **Limitation:** You can, however, e.g. not convert a string to an integer.
try: # try to run this block
x = int('222a')
print('can be done')
print(x)
except: # if any error found run this block instead
print('canNOT be done')
# **Note**: The identation is required (typically 4 spaces).
# **Question**: Can you convert a boolean variable `x = False` to an integer?
#
# - **A:** No
# - **B:** Yes, and the result is 0
# - **C:** Yes, and the result is 1
# - **D:** Yes, and the result is -1
# - **E:** Don't know
# **Socrative room:** *NUMECON*
# ## 2.3 Operators
# Variables can be combined using **operators** (e.g. +, -, /, **).<br>For numbers we have:
x = 3
y = 2
print(x+y)
print(x-y)
print(x/y)
print(x*y)
# For strings we can use an overloaded '+' for concatenation:
x = 'abc'
y = 'def'
print(x+y)
# A string can also be multiplied by an integer:
x = 'abc'
y = 2
print(x*y)
# **Question**: What is the result of `x = 3**2`?
#
# - **A:** `x = 3`
# - **B:** `x = 6`
# - **C:** `x = 9`
# - **D:** `x = 12`
# - **E:** Don't know
#
# **Socrative room:** *NUMECON*
# **Note:** Standard division converts integers to floating point numbers.
x = 8
y = x/2 # standard division
z = x//3 # integer division
print(y,type(y))
print(z,type(z))
# ## 2.4 Augmentation
# Variables can be changed using **augmentation operators** (e.g. +=, -=, *=, /=)
x = 3
print(x)
x += 1 # same result as x = x+1
print(x)
x *= 2 # same result as x = x*2
print(x)
x /= 2 # same result as x = x/2
print(x)
# ## 2.5 Comparision
# Variables can be compared using **boolean operators** (e.g. ==, !=, <, <=, >, >=).
x = 3
y = 2
z = 10
print(x < y) # less than
print(x <= y) # less than or equal
print(x != y) # not equal
print(x == y) # equal
# The comparison returns a boolean variable:
z = x < y # z is now a boolean variable
print(z)
type(z)
# ## 2.6 Summary
# The new central concepts are:
#
# 1. Variable
# 2. Reference
# 3. Object
# 4. Type (int, float, str, bool)
# 5. Value
# 6. Operator (+, -, *, **, /, //, % etc.)
# 7. Augmentation (+=, -=, *=, /= etc.)
# 8. Comparison (==, !=, <, <= etc.)
# <a id="Containers"></a>
#
# # 3. Containers
# A more complicated type of object is a **container**. This is an object, which consists of serveral objects of e.g. an atomic type. They are also called **collection types**.
# ## 3.1 Lists
# A first example is a **list**. A list contains **variables** each **referencing** some **object**.
# +
x = [1,'abc']
# variable x references a list type object with elements
# referencing 1 and 'abc'
print(x,type(x))
# -
# The **length** of a list can be found with the **len** function.
print(f'the number of elements in x is {len(x)}')
# A list is **subscriptable** and starts, like everything in Python, from **index 0**. Beware!
print(x[0]) # 1st element
print(x[1]) # 2nd element
# A list is **mutable**, i.e. you can change its elements on the fly. Ie., you can change its **references** to objects.
x[0] = 'def'
x[1] = 2
print(x)
# and add more elements
x.append('new_element') # add new element to end of list
print(x)
# **Link:** [Why is 0 the first index?](http://python-history.blogspot.com/2013/10/why-python-uses-0-based-indexing.html)
# ### Slicing
# A list is **slicable**, i.e. you can extract a list from a list.
# +
x = [0,1,2,3,4,5]
print(x[0:3]) # x[0] included, x[3] not included
print(x[1:3])
print(x[:3])
print(x[1:])
print(x[:99]) # This is very particular to Python. Normally you'd get an error.
print(x[:-1]) # x[-1] is the last element
print(type(x[:-1])) # Slicing yields a list
print(type(x[-1])) # Unless only 1 element
# -
# **Explantion:** Slices are half-open intervals. I.e. ``x[i:i+n]`` means starting from element ``x[i]`` and create a list of (up to) ``n`` elements.
# splitting a list at x[3] and x[5] is:
print(x[0:3])
print(x[3:5])
print(x[5:])
# **Question**: Consider the following code:
x = [0,1,2,3,4,5]
# What is the result of `print(x[-4:-2])`?
#
# - **A:** [1,2,3]
# - **B:** [2,3,4]
# - **C:** [2,3]
# - **D:** [3,4]
# - **E:** Don't know
# ### Referencing
# **Important**: Multiple variables can refer to the **same** list.
x = [1,2,3]
y = x # y now references the same list as x
y[0] = 2 # change the first element in the list y
print(x) # x is also changed because it references the same list as y
# If you want to know if two variables contain the same reference, use the **is** operator.
print(y is x)
z = [1,2]
w = [1,2]
print(z is w) # z and w have the same numerical content, but do not reference the same object.
# **Conclusion:** The `=` sign copy the reference, not the content! What about the atomic types?
z = 10
w = z
print(z is w) # w is now the same reference as z
z += 5
print(z, w)
print(z is w) # z was overwritten in the augmentation statement.
# If one variable is deleted, the other one still references the list.
del x # delete the variable x
print(y)
# Instead, lists can by **copied** by using the copy-module:
# +
from copy import copy
x = [1,2,3]
y = copy(x) # y now a copy of x
y[0] = 2
print(y)
print(x) # x is not changed when y is changed
print(x is y) # as they are not the same reference
# -
# or by slicing:
x = [1,2,3]
y = x[:] # y now a copy of x
y[0] = 2
print(y)
print(x) # x is not changed when y is changed
# **Advanced**: A **deepcopy** is necessary, when the list contains mutable objects:
# +
from copy import deepcopy
a = [1,2,3]
x = [a,2,3] # x is a list of a list and two integers
y1 = copy(x) # y1 now a copy x
y2 = deepcopy(x) # y2 is a deep copy
a[0] = 10 # change1
x[-1] = 1 # change2
print(x) # Both changes happened
print(y1) # y1[0] reference the same list as x[0]. Only change1 happened
print(y2) # y2[0] is a copy of the original list referenced by x[0]
# -
# **Question**: Consider the following code:
x = [1,2,3]
y = [x,x]
z = x
z[0] = 3
z[2] = 1
# What is the result of `print(y[0])`?
#
# - **A:** 1
# - **B:** 3
# - **C:** [3,2,1]
# - **D:** [1,2,3]
# - **E:** Don't know
# ## 3.2 Tuples
# A **tuple** is an **immutable list**.<br>It is similar when extracting information:
x = (1,2,3) # note: parentheses instead of square backets
print(x,type(x))
print(x[2])
print(x[:2])
# But it **cannot be changed** (it is immutable):
try: # try to run this block
x[0] = 2
print('did succeed in setting x[0]=2')
except: # if any error found run this block instead
print('did NOT succeed in setting x[0]=2')
print(x)
# ## 3.3 Dictionaries
# A **dictionary** is a **key-based** (instead of index-based) container.
#
# * **Keys:** All immutable objects are valid keys.
# * **Values:** Fully unrestricted.
x = {} # create x as an empty dictionary
x['abc'] = '1' # key='abc', value = '1'
print(x['abc'])
x[('abc',1)] = 2 # key=('abc',1), value = 2
# Elements of a dictionary are **extracted** using their keyword:
key = 'abc'
value = x[key]
print(value)
key = ('abc',1)
value = x[key]
print(value)
# Dictionaries can also be **created with content**:
y = {'abc': '1', 'a': 1, 'b': 2, 'c': 3}
print(y['c'])
# **Content is deleted** using its key:
print(y)
del y['abc']
print(y)
# **Task:** Create a dictionary called `capitals` with the capital names of Denmark, Sweden and Norway as values and country names as keys.
# **Answer:**
# + jupyter={"source_hidden": true}
capitals = {}
capitals['denmark'] = 'copenhagen'
capitals['sweden'] = 'stockholm'
capitals['norway'] = 'oslo'
capital_of_sweden = capitals['sweden']
print(capital_of_sweden)
# -
# ## 3.4 Summary
# The new central concepts are:
#
# 1. Containers (lists, tuples, dictionaries)
# 2. Mutable/immutable
# 3. Slicing of lists and tuples
# 4. Referencing (copy and deepcopy)
# 5. Key-value pairs for dictionaries
# **Note:** All atomic types as immutable, and only strings are subscriptable.
x = 'abcdef'
print(x[:3])
print(x[3:5])
print(x[5:])
try:
x[0] = 'f'
except:
print('strings are immutable')
# **Advanced:** Other interesting containers are e.g. **namedtuple** and **OrderDict** (see [collections](https://docs.python.org/2/library/collections.html)), and [**sets**](https://docs.python.org/2/library/sets.html).
# <a id="Conditionals-and-loops"></a>
#
# # 4. Conditionals and loops
# ## 4.1 Conditionals
# You typically want your program to do one thing if some condition is met, and another thing if another condition is met.
#
# In Python this is done with **conditional statments**:
x = 3
if x < 2:
# happens if x is smaller than 2
print('first possibility')
elif x > 4: # elif = else if
# happens if x is not smaller than 2 and x is larger than 4
print('second possibility')
elif x < 0:
# happens if x is not smaller than 2, x is not larger than 4
# and x is smaller than 0
print('third posibility') # note: this can never happen
else:
# happens if x is not smaller than 2, x is not larger than 4
# and x is not smaller than 0
print('fourth possiblity')
# **Note:**
#
# 1. "elif" is short for "else if"
# 2. the **indentation** after if, elif and else is required (typically 4 spaces)
# An **equivalent formulation** of the above if-elif-else statement is:
# +
x = -1
cond_1 = x < 2 # a boolean (True or False)
cond_2 = x > 4 # a boolean (True or False)
cond_3 = x < 0 # a boolean (True or False)
if cond_1:
print('first possibility')
elif cond_2:
print('second possibility')
elif cond_3:
print('third posibility')
else:
print('fourth possiblity')
y = [1, 2]
if y:
print('y is not empty')
# -
# The above can also be written purely in terms of if-statements:
if cond_1:
print('first possibility')
if not cond_1 and cond_2:
print('second possibility')
if not (cond_1 or cond_2) and cond_3:
print('third posibility')
if not (cond_1 or cond_2 or cond_3):
print('fourth possiblity')
# ## 4.2 Simple loops
# You typically also want to **repeat a task multiple times**. But it is time-consuming and **error prone** to write:
x_list = [0,1,2,3,4]
y_list = [] # empty list
y_list.append(x_list[0]**2)
y_list.append(x_list[1]**2)
y_list.append(x_list[2]**2)
y_list.append(x_list[3]**2)
y_list.append(x_list[4]**2)
print(y_list)
# You should at **all costs** avoid repeating code. Therefore use a **for loop** instead:
y_list = [] # empty list
for x in x_list:
y_list.append(x**2)
print(y_list)
# Use a **while loop**:
y_list = [] # empty list
i = 0
while i <= 4:
y_list.append(x_list[i]**2)
i += 1
print(y_list)
# Use a **for loop** with **range** instead:
y_list = [] # empty list
for x in range(5):
print(x)
y_list.append(x**2)
print(y_list)
# Use a **list comprehension**:
y_list = [x**2 for x in x_list]
print(y_list)
# **Note:** List comprehension is the shortest (and fastest) code, but can become messy in more complicated situations.
# ## 4.3 More complex loops
# For loops can also be **enumerated**.
y_list = []
for i,x in enumerate(x_list):
print(i)
y_list.append(x**2)
print(y_list)
# Loops can be fine-tuned with **continue** and **break**.
# +
y_list = []
x_list = [*range(10)]
for i,x in enumerate(x_list):
if i == 1:
continue # go to next iteration
elif i == 4:
break # stop loop prematurely
y_list.append(x**2)
print(y_list)
# -
# **Task:** Create a list with the 10 first positive uneven numbers.
# +
# write your code here
# -
# **Answer:**
# + jupyter={"source_hidden": true}
my_list = []
for i in range(10):
my_list.append((i+1)*2-1)
print(my_list)
# -
# **Zip:** We can loop over **2 lists at the same time**:
# +
x = ['I', 'II', 'III']
y = ['a', 'b', 'c']
for i,j in zip(x,y):
print(i+j)
# -
# Iter(ation)tools enable us do complicated loops in a smart way. We can e.g. loop through **all combinations of elements in 2 lists**:
for i in x:
for j in y:
print(i+j)
import itertools as it
for i,j in it.product(x,y):
print(i,j)
# ## 4.4 Dictionaries
# We can loop throug keys, values or key-value pairs of a dictionary.
my_dict = {'a': '-', 'b': '--', 'c': '---'}
for key in my_dict.keys():
print(key)
for val in my_dict.values():
print(val)
for key,val in my_dict.items():
print(key,val)
# We can also **check whether a key exists**:
if 'a' in my_dict:
print('a is in my_dict with the value ' + my_dict['a'])
else:
print('a is not in my_dict')
if 'd' in my_dict:
print('d is in my_dict with the value ' + my_dict['d'])
else:
print('d is not in my_dict')
# **Note:** dictionaries can do this operation very quickly without looping through all elements. So use a dictionary when lookups are relevant.
# + [markdown] code_folding=[] heading_collapsed=true hidden=true
# ## 4.5 Summary
# -
# The new central concepts are:
#
# 1. Conditionals (if, elif, else)
# 2. Loops (for, while, range, enumerate, continue, break, zip)
# 3. List comprehensions
# 4. Itertools (product)
# <a id="Functions"></a>
#
# # 5. Functions
# The most simple function takes **one argument** and returns **one output**:
# +
def f(x):
return x**2
print(f(2))
# -
# **Note:** The identation after `def` is again required (typically 4 spaces).
# Alternatively, you can use a single-line **lambda formulation**:
g = lambda x: x**2
print(g(2))
# Introducing **multiple arguments** are straigtforward:
# +
def f(x,y):
return x**2 + y**2
print(f(2,2))
# -
# So are **multiple outputs**:
# +
def f(x,y):
z = x**2
q = y**2
return z,q
full_output = f(2,2) # returns a tuple
print(full_output)
# -
# The output tuple can be unpacked:
z,q = full_output # unpacking
print(z)
print(q)
# ## 5.1 No outputs...
# Functions without *any* output can be useful when arguments are mutable:
# +
def f(x): # assume x is a list
new_element = x[-1]+1
x.append(new_element)
x = [1,2,3] # original list
f(x) # update list (appending the element 4)
f(x) # update list (appending the element 5)
f(x)
print(x)
# -
# Note: this is called a side-effect, which is often best avoided.
# ## 5.2 Keyword arguments
# We can also have **keyword arguments** with default values (instead of **positionel** arguments):
# +
def f(x,y,a=2,b=2):
return x**a + y**b
print(f(2,4)) # 2**2 + 2**2
print(f(2,2,b=3)) # 2**3 + 2**2
print(f(2,2,a=3,b=3)) # 2**3 + 2**3
# -
# **Note:** Keyword arguments must come after positional arguments.
# **Advanced:** We can also use undefined keyword arguments:
def f(**kwargs):
# kwargs (= "keyword arguments") is a dictionary
for key,value in kwargs.items():
print(key,value)
f(a='abc',b='2',c=[1,2,3])
# and these keywords can come from *unpacking a dictionary*:
my_dict = {'a': 'abc', 'b': '2', 'c': [1,2,3]}
f(**my_dict)
# ## 5.3 A function is an object
# A function is an object and can be given to another functions as an argument.
# +
def f(x):
return x**2
def g(x,h):
temp = h(x) # call function h with argument x
return temp+1
print(g(2,f))
# -
# ## 5.4 Scope
# **Important:** Variables in functions can be either **local** or **global** in scope.
# +
a = 2 # a global variable
def f(x):
return x**a # a is global
def g(x,a=2):
# a's default value is fixed when the function is defined
return x**a
def h(x):
a = 2 # a is local
return x**a
print(f(2), g(2), h(2))
print('incrementing the global variable:')
a += 1
print(f(2), g(2), h(2)) # output is only changed for f
# -
# **Recommendation:** Never rely on global variables, they make it hard to understand what your code is doing.
# ## 5.5 Summary
# **Functions:**
#
# 1. are **objects**
# 2. can have multiple (or no) **arguments** and **outputs**
# 3. can have **positional** and **keyword** arguments
# 4. can use **local** or **global** variables (**scope**)
# **Task:** Create a function returning a person's full name from her first name and family name with middle name as an optional keyword argument with empty as a default.
# +
# write your code here
# -
# **Answer:**
# + jupyter={"source_hidden": true}
def full_name(first_name,family_name,middle_name=''):
name = first_name
if middle_name != '':
name += ' '
name += middle_name
name += ' '
name += family_name
return name
print(full_name('Jeppe','Druedahl','"Economist"'))
# -
# **Alternative answer** (more advanced, using a built-in list function):
# + jupyter={"outputs_hidden": true, "source_hidden": true}
def full_name(first_name,family_name,middle_name=''):
name = [first_name]
if middle_name != '':
name.append(middle_name)
name.append(family_name)
return ' '.join(name)
print(full_name('Jeppe','Druedahl','"Economist"'))
# -
# <a id="Floating-point-numbers"></a>
#
# # 6. Floating point numbers
# There are uncountable many real numbers. On a computer the real line is approximated with numbers on the form:
#
# $$\text{number} = \text{significand} \times \text{base}^{exponent}$$
#
# * **significand**: 1 bit, positive or negative
# * **base**: 52 bits
# * **exponent**: 11 bits
# All numbers is therefore *not* represented, but a *close* neighboring number is used.
x = 0.1
print(f'{x:.100f}') # printing x with 100 decimals
x = 17.2
print(f'{x:.100f}') # printing x with 100 decimals
# Simple sums might, consequently, not be exactly what you expect.
print(0.1 + 0.1 + 0.1 + 0.1 + 0.1 + 0.1 + 0.1 + 0.1 + 0.1 + 0.1)
# **Comparisions of floating point numbers** is therefore always problematic.<br>
# We know that
#
# $$\frac{a \cdot c}{b \cdot c} = \frac{a}{b}$$
#
# but:
a = 0.001
b = 11.11
c = 1000
test = (a*c) / (b*c) == a / b
print(test)
# **Underflow**: Multiplying many small numbers can result in an exact zero:
x = 1e-60
y = 1
for _ in range(6):
y *= x
print(y)
# **Overflow**: If intermediate results are too large to be represented, the final result may be wrong or not possible to calculate:
x = 1.0
y = 2.7
for i in range(200):
x *= (i+1)
y *= (i+1)
print(y/x) # should be 2.7
print(x,y)
# **Note:** `nan` is not-a-number. `inf` is infinite.
# **Note:** Order of additions matter, but not by that much:
sum1 = 10001234.0 + 0.12012 + 0.12312 + 1e-5
sum2 = 1e-5 + 0.12312 + 0.12012 + 10001234.0
print(sum1-sum2)
# ## 6.1 Summary
# The take-aways are:
#
# 1. Decimal numbers are **approximate** on a computer!
# 2. **Never compare floats with equality** (only use strict inequalities)
# 3. Underflow and overflow can create problem (not very important in practice)
#
# For further details see [here](https://docs.python.org/3/tutorial/floatingpoint.html).
#
# **Videos:**
#
# * [Why computers are bad at algebra - Infinite Series](https://www.youtube.com/watch?v=pQs_wx8eoQ8)
# * [Floating point numbers - Computerphile](https://www.youtube.com/watch?v=PZRI1IfStY0)
# <a id="Classes-(user-defined-types)"></a>
#
# # 7. Classes (user-defined types)
# **Advanced:** New types of objects can be defined using **classes**.
class human():
def __init__(self,name,height,weight): # called when created
# save the inputs as attributes
self.name = name # an attribute
self.height = height # an attribute
self.weight = weight # an attribute
def bmi(self): # a method
bmi = self.weight/(self.height/100)**2 # calculate bmi
return bmi # output bmi
# A class is used as follows:
# +
# a. create an instance of the human object called "jeppe"
jeppe = human('jeppe',182,80) # height=182, weight=80
print(type(jeppe))
# b. print an attribute
print(jeppe.name)
# c. print the result of calling a method
print(jeppe.bmi())
# -
# **Methods** are like functions, but can automatically use all the attributes of the class (saved in *self.*) without getting them as arguments.
# **Attributes** can be changed and extracted with **.-notation**
jeppe.height = 160
print(jeppe.height)
print(jeppe.bmi())
# Or with **setattr- and getatrr-notation**
setattr(jeppe,'height',182) # jeppe.height = 182
height = getattr(jeppe,'height') # height = jeppe.height
print(height)
print(jeppe.bmi())
# ## 7.1 Operator methods
# If the **appropriate methods** are defined, standard operators, e.g. +, and general functions such as print can be used.
#
# Define a new type of object called a **fraction**:
# + code_folding=[] hidden=true
class fraction:
def __init__(self,numerator,denominator): # called when created
self.num = numerator
self.denom = denominator
def __str__(self): # called when using print
return f'{self.num}/{self.denom}' # string = self.nom/self.denom
def __add__(self,other): # called when using +
new_num = self.num*other.denom + other.num*self.denom
new_denom = self.denom*other.denom
return fraction(new_num,new_denom)
# -
# **Note:** We use that
#
# $$\frac{a}{b}+\frac{c}{d}=\frac{a \cdot d+c \cdot b}{b \cdot d}$$
# We can now **add fractions**:
x = fraction(1,3)
print(x)
x = fraction(1,3) # 1/3 = 5/15
y = fraction(2,5) # 2/5 = 6/15
z = x+y # 5/15 + 6/15 = 11/15
print(z,type(z))
# Equivalent to:
z_alt = x.__add__(y)
print(z,type(z))
# But we **cannot multiply** fractions (yet):
try:
z = x*y
print(z)
except:
print('multiplication is not defined for the fraction type')
# **Extra task:** Implement multiplication for fractions.
# ## 7.2 Summary
# The take-aways are:
#
# 1. **A class is a user-defined type**
# 2. **Attributes** are like **variables** encapsulated in the class
# 3. **Methods** are like **functions** encapsulated in the class
# 4. Operators are fundamentally defined in terms of methods
# <a id="Summary"></a>
#
# # 8. Summary
# **This lecture:** We have talked about:
# 1. Types (int, str, float, bool, list, tuple, dict)
# 2. Operators (+, *, /, +=, *=, /=, ==, !=, <)
# 3. Referencing (=) vs. copying (copy, deepcopy)
# 4. Conditionals (if-elif-else) and loops (for, while, range, enumerate, zip, product)
# 5. Functions (positional and keyword arguments) and scope
# 6. Floating points
# 7. Classes (attributes, methods)
# **You work:** When you are done with the DataCamp courses read through this notebook, play around with the code and ask questions if there is stuff you don't understand.
# **Next lecture:** We will solve the consumer problem from microeconomics numerically.
# **Your to-do list:** You should be running JupyterLab on your own computer.
# <a id="Extra:-Iterators"></a>
#
# # 9. Extra: Iterators
# Consider the following loop, where my_list is said to be **iterable**.
my_list = [0,2,4,6,8]
for i in my_list:
print(i)
# Consider the same loop generated with an **iterator**.
for i in range(0,10,2):
print(i)
# This can also be written as:
x = iter(range(0,10,2))
print(x)
print(next(x))
print(next(x))
print(next(x))
# The main benefit here is that the, potentially long, my_list, is never created.
# We can also write **our own iterator class**:
class range_two_step:
def __init__(self, N):
self.i = 0
self.N = N
def __iter__(self):
return self
def __next__(self):
if self.i >= self.N:
raise StopIteration
temp = self.i
self.i = self.i + 2
return temp
# Can then be used as follows:
x = iter(range_two_step(10))
print(next(x))
print(next(x))
print(next(x))
# Or in a loop:
for i in range_two_step(10):
print(i)
# <a id="Extra:-More-on-functions"></a>
#
# # 10. Extra: More on functions
# We can have an **undefined number of input arguments**:
def f(*args):
out = 0
for x in args:
out += x**2
return out
print(f(2,2))
print(f(2,2,2,2))
# We can have **recursive functions** to calculate the Fibonacci sequence:
# $$
# \begin{aligned}
# F_0 &= 0 \\
# F_1 &= 1 \\
# F_n &= F_{n-1} + F_{n-2} \\
# \end{aligned}
# $$
# +
def fibonacci(n):
if n == 0:
return 0
elif n == 1:
return 1
else:
return fibonacci(n-1) + fibonacci(n-2)
y = fibonacci(7)
print(y)
|
web/02/Primitives.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
import copy
import logging
import sys
# from run_tests_201204 import *
import os
import sys
import importlib
from collections import defaultdict
sys.path.insert(0, '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc')
from tools_pattern import get_eucledean_dist
import compress_pickle
import my_plot
from my_plot import MyPlotData, my_box_plot
import seaborn as sns
script_n = 'plot_210603_grc_pcts_1024'
data_script = 'batch_210603_grc_pcts'
db_path = '/n/groups/htem/Segmentation/shared-nondev/cb2_segmentation/analysis_mf_grc/dimensionality_sim2/' \
f'{data_script}/'
scaled_noise = 0
core_noise = 0
n_mfs = 400
n_grcs = 2400
pattern_type = 'binary'
db = {}
model = 'global_random'
db[model] = compress_pickle.load(
db_path+f'{data_script}_{model}_{pattern_type}_{n_grcs}_{n_mfs}_0.3_1024_10.gz')
# avg_grc_dim_list = defaultdict(list)
# for ress in db['random']:
# ress_tries = ress
# for ress in ress_tries:
# # print(ress)
# for noise in ress:
# res = ress[noise]
# grc_dim = res['grc_dim']
# avg_grc_dim_list[noise].append(grc_dim)
# avg_grc_dim = {}
# for noise in avg_grc_dim_list:
# avg_grc_dim[noise] = sum(avg_grc_dim_list[noise])/len(avg_grc_dim_list[noise])
# +
name_map = {
'scaleup4': "Observed",
'global_random': "Global Random",
'random': "Global Random",
# 'naive_random_17': "Local Random",
'naive_random4': "Local Random",
}
palette = {
name_map['scaleup4']: sns.color_palette()[0],
name_map['global_random']: sns.color_palette()[1],
name_map['random']: sns.color_palette()[1],
name_map['naive_random4']: sns.color_palette()[2],
# name_map['naive_random_21']: sns.color_palette()[2],
}
mpd = MyPlotData()
ress_ref = db['global_random'][0][0]
resss_ref2 = db['global_random'][0]
for model_name in [
'global_random',
# 'naive_random_17',
# 'naive_random4',
# 'random',
# 'scaleup4',
]:
ress = db[model_name]
# print(ress)
ress_tries = ress[0] # get the first element in tuple
# ress = ress[0] # get the first try
for n_try, ress in enumerate(ress_tries):
# print(resss_ref2[0])
# print(resss_ref2.keys())
if n_try >= len(resss_ref2):
print(n_try)
continue
ress_ref2 = resss_ref2[n_try]
for noise in ress:
# print(noise)
res = ress[noise]
# res_ref = ress_ref[noise]
res_ref2 = ress_ref2[noise]
# hamming_distance_norm = res['hamming_distance']/res['num_grcs']
mpd.add_data_point(
model=name_map[model_name],
# noise=noise*100,
grc_pcts=noise*100*2400/1200,
grc_dim=res['grc_dim'],
grc_dim_norm=res['grc_dim']/res_ref2['grc_dim'],
# grc_dim_norm2=res['grc_dim']/avg_grc_dim[noise],
grc_by_mf_dim=res['grc_dim']/res['mf_dim'],
# grc_by_mf_dim_ref=res['grc_dim']/res_ref['mf_dim'],
num_grcs=res['num_grcs'],
num_mfs=res['num_mfs'],
voi=res['voi'],
grc_pop_corr=res['grc_pop_corr'],
grc_pop_corr_norm=res['grc_pop_corr']/res_ref2['grc_pop_corr'],
binary_similarity=res['binary_similarity'],
hamming_distance=res['hamming_distance'],
normalized_mse=res['normalized_mse'],
)
# importlib.reload(my_plot); my_plot.my_relplot(
# mpd,
# x='noise',
# y='grc_dim',
# hue='model',
# context='paper',
# palette=palette,
# linewidth=1,
# log_scale_y=True,
# width=10,
# # ylim=[0, None],
# y_axis_label='Dim. Expansion ($x$)',
# x_axis_label='MF Input Variation (%)',
# title='noise',
# save_filename=f'{script_n}_act_30.svg',
# show=True,
# )
# +
importlib.reload(my_plot); my_plot.my_relplot(
mpd,
x='grc_pcts',
y='grc_dim',
hue='model',
context='paper',
palette=palette,
linewidth=1,
# log_scale_y=True,
width=10,
ylim=[0, None],
y_axis_label='Dim. Expansion ($x$)',
x_axis_label='MF Input Variation (%)',
title='noise',
save_filename=f'{script_n}_act_30.svg',
show=True,
)
|
analysis/dimensionalty_sim/plot_210603_grc_pcts_1024.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/marisbotero/tensorflow_notes/blob/master/Housing_price.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="V45-ABwje8T9" colab_type="text"
# En este ejercicio intentará construir una red neuronal que prediga el precio de una casa de acuerdo con una fórmula simple.
#
# Entonces, imagine si el precio de la casa fuera tan fácil como una casa cuesta 50k + 50k por habitación, de modo que una casa de 1 habitación cuesta 100k, una casa de 2 habitaciones cuesta 150k, etc.
#
# ¿Cómo crearía una red neuronal que aprenda esta relación para predecir que una casa de 7 dormitorios costará cerca de 400k, etc.
#
# Sugerencia: Su red podría funcionar mejor si reduce el precio de la vivienda. No tiene que dar la respuesta 400 ... podría ser mejor crear algo que prediga el número 4, y luego su respuesta está en los 'cientos de miles', etc.
# + id="pN6tLjdVe9Iy" colab_type="code" colab={}
import tensorflow as tf
import numpy as np
from tensorflow import keras
# + id="Kq2DL_sifEGF" colab_type="code" colab={}
# GRADED FUNCTION: house_model
def house_model(y_new):
xs=[]
ys=[]
for i in range(1,10):
xs.append(i)
ys.append((1+float(i))*50)
xs=np.array(xs,dtype=float)
ys=np.array(ys, dtype=float)
model = keras.Sequential([keras.layers.Dense(units = 1, input_shape = [1])])
model.compile(optimizer='sgd', loss='mean_squared_error')
model.fit(xs, ys, epochs = 4500)
return (model.predict(y_new)[0]/100)
# + id="BGLnVrCxfMIW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f396c2b7-2413-4138-c8de-1270620c50c8"
prediction = house_model([7.0])
print(prediction)
# + id="BhUR4CuEfOvB" colab_type="code" colab={}
|
Housing_price.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from scipy.spatial import distance
# +
class Point:
def __init__(self, x, y):
self.x = x
self.y = y
@property
def as_np(self):
return np.array([self.y, self.x])
def parse_point(s):
x, y = s.split(',')
return Point(int(x), int(y))
# -
max_x = -np.inf
max_y = -np.inf
points = {}
with open("input.txt", "r") as fp:
for l in fp.readlines():
key = l.strip()
p = parse_point(key)
points[key] = p
if p.x > max_x:
max_x = p.x
if p.y > max_y:
max_y = p.y
p_list = list(points.keys())
masks = []
ogrid = np.ogrid[0:max_y+1, 0:max_x+1]
for k in points:
masks.append(distance.cityblock(points[k].as_np, ogrid))
masks = np.array(masks)
group_mask = np.argmin(masks, axis=0)
group_mask
dup_mask = np.ones(group_mask.shape)
z, y, x = masks.shape
for i in range(y):
for j in range(x):
arr = masks[:, i, j]
if len(arr[arr == arr.min()]) > 1:
dup_mask[i, j] = np.nan
dup_mask
region_mask = group_mask * dup_mask
region_mask
boundary = np.ones_like(region_mask)
boundary[1:-1, 1:-1] = np.nan
boundary_mask = boundary * region_mask
boundary_mask
max_area = 0
for i, p in enumerate(p_list):
if i in boundary_mask:
continue
else:
area = len(region_mask[region_mask == i])
if area > max_area:
max_area = area
max_p = p
print(max_area)
print(max_p)
s = masks.sum(axis=0)
len(s[s<10000])
|
day6/day 6 part 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# +
from math import *
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
# %matplotlib inline
from netCDF4 import Dataset
import numpy as np
import os
import pandas as pd
import pylab as pl
import scipy.io
import scipy as spy
import seaborn as sns
import sys
#lib_path = os.path.abspath('../../Building_canyon/BuildCanyon/PythonModulesMITgcm') # Add absolute path to my python scripts
lib_path = os.path.abspath('../BuildCanyon/PythonModulesMITgcm') # Add absolute path to my python scripts
sys.path.append(lib_path)
import ReadOutTools_MITgcm as rout
import MetricsPythonTools as mpt
# -
sns.set()
sns.set_style('darkgrid')
sns.set_context('poster')
# +
def get_metrics(exp, run, TrNum, key):
'''Get column from a tracer metrics pandas dataframe using the key name, run (01, 02, etc) and experiment
abreviated name (BAR, CNTDIFF, 3DDIFF, NOGMREDI). All input variables are strings. Returns the desired column from the dataframe'''
df = pd.read_csv(('results/metricsDataFrames/%srun%sTr%s.csv' %(exp,run,TrNum)))
col = df[key]
return col
def get_water(exp, run, key):
'''Get column from a tracer metrics pandas dataframe using the key name, run (01, 02, etc) and experiment
abreviated name (BAR, CNTDIFF, 3DDIFF, NOGMREDI). All input variables are strings. Returns the desired column from the dataframe'''
df = pd.read_csv(('results/metricsDataFrames/%srun%s.csv' %(exp,run)))
col = df[key]
return col
def get_areas(file, key):
'''Get column from a tracer metrics pandas dataframe using the key name, run (01, 02, etc) and experiment
abreviated name (BAR, CNTDIFF, 3DDIFF, NOGMREDI). All input variables are strings. Returns the desired column from the dataframe'''
df = pd.read_csv(file)
col = df[key]
return col
# +
#Exp
CGrid = '/Users/Karina/Research/PhD/Tracers/TemporaryData/BARKLEY/run01/gridGlob.nc' # Smallest volume grid, closed bdy, no canyon.
#CGrid = '/ocean/kramosmu/MITgcm/TracerExperiments/CNTDIFF/run03/gridGlob.nc' # Smallest volume grid, closed bdy, no canyon.
CGridOut = Dataset(CGrid)
# General input
nx = 360
ny = 360
nz = 90
nt = 19 # t dimension size
numTr = 21 # number of tracers in total
rc = CGridOut.variables['RC']
xc = rout.getField(CGrid, 'XC') # x coords tracer cells
yc = rout.getField(CGrid, 'YC') # y coords tracer cells
drF = CGridOut.variables['drF'] # vertical distance between faces
drC = CGridOut.variables['drC'] # vertical distance between centers
# -
# +
labels = ['Kv=1E-5, Ki=10',
'Kv=1E-4, Ki=10',
'Kv=1E-3, Ki=10', #run02
'Kv=1E-5, Ki=1',
'Kv=1E-4, Ki=1',
'Kv=1E-3, Ki=1', # run03
'Kv=1E-5, Ki=0.1',
'Kv=1E-4, Ki=0.1',
'Kv=1E-3, Ki=0.1', #run04
#'Kv_noc=1E-5, Ki=1',
#'Kv_noc=1E-4, Ki=1',
#'Kv_noc=1E-3, Ki=1',#run07
'Kv=3.8E-5, Ki=10',
'Kv=2.8E-5, Ki=10',
'Kv=1.3E-5, Ki=10', #run09
'Kv=3.8E-5, Ki=1',
'Kv=2.8E-5, Ki=1',
'Kv=1.3E-5, Ki=1',#run10
'Kv=KTv=KSv=1E-4, Ki=1', # run11
'Kv=KTv=KSv=1E-3, Ki=1,Kt=Ks',#run12
'Kv=1E-7(out), 1E-3(in), Ki=1',# 3Drun04
'Kv=1E-7(out), 1E-4(in), Ki=1', # 3Drun05
'Kv=1E-5(out), 1E-3(in), Ki=1',# 3Drun06
'Kv=1E-5(out), 1E-4(in), Ki=1']# 3Drun07
wlabels = ['run02 - CNT','run03 - CNT','run04 - CNT','run09 - CNT',
'run10 - CNT','run11 - CNT','run12 - CNT','run04 - 3D','run05 - 3D','run06 - 3D','run07 - 3D']
times = np.arange(0,nt,1)
# +
# LOAD AREAS
CS1A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS1area' )
CS2A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS2area' )
CS3A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS3area' )
CS3sbA = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS3sbarea' )
CS4A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS4area' )
CS5A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'CS5area' )
AS1A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'AS1area' )
AS2A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'AS2area' )
LID1A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'LID1area' )
LID2A = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'LID2area' )
VolHole = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'VolHole' )
VoleShwoHole = get_areas('results/metricsDataFrames/Canyon_AreasVolumes.csv', 'VolShNoHole' )
# +
tracers_3D = ['04','05','06','07'] #run number because there's only 1 tr per run
tracers_CNT03 = ['1','2','3'] # tracer number , constant runs
tracers_CNT09 = ['1','2','3'] # tracer number , average diffusivity runs
tracers_CNT07 = ['1','2','3'] # tracer number , no canyon case
tracers_CNT02 = ['1','2','3'] # tracer number , Kiso=0.1
tracers_CNT04 = ['1','2','3'] # tracer number , Kiso=10
tracers_CNT10 = ['1','2','3'] # tracer number , Kiso=1
tracers_CNT11 = ['2'] # tracer number , Kiso=1, Ks=Kt=10^4
tracers_CNT12 = ['3'] # tracer number , Kiso=1, Ks=Kt=10^3
# LOAD TRACER ON SHELF DATA
TrOnSh = np.zeros((nt,numTr))
HWC = np.zeros((nt,numTr))
kk = 0
fields = ['TronShelfwHole', 'HCWonShelfwHole','TronHole','HCWonHole']
for ii in tracers_CNT02:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '02',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '02',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '02',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '02', ii,fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
for ii in tracers_CNT03:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '03',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '03', ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '03', ii,fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '03',ii, fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
for ii in tracers_CNT04:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '04',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '04',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '04',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '04',ii, fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
#for ii in tracers_CNT07:
# TrSh = get_metrics('CNTDIFF_hole_', '07', ii, fields[0] )
# TrHole= get_metrics('CNTDIFF_hole_', '07', ii, fields[2] )
# HWCSh = get_metrics('CNTDIFF_hole_', '07', ii, fields[1] )
# HWCHole = get_metrics('CNTDIFF_hole_', '07', ii, fields[3] )
# TrOnSh[:,kk] = TrHole+ TrOnShwHole
# HWC[:,kk] = HWCHole+ HWCsh
# kk=kk+1
for ii in tracers_CNT09:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '09',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_','09',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '09',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '09',ii, fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
for ii in tracers_CNT10:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '10',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '10',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '10',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '10',ii, fields[3] )
HWC[:,kk] = HWChole + HWCsh
kk=kk+1
for ii in tracers_CNT11:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '11',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '11',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '11',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '11',ii, fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
for ii in tracers_CNT12:
TrOnShwHole = get_metrics('CNTDIFF_hole_', '12',ii, fields[0] )
TrOnHole = get_metrics('CNTDIFF_hole_', '12',ii, fields[2] )
TrOnSh[:,kk] = TrOnHole+ TrOnShwHole
HWCsh = get_metrics('CNTDIFF_hole_', '12',ii, fields[1] )
HWChole = get_metrics('CNTDIFF_hole_', '12',ii, fields[3] )
HWC[:,kk] = HWChole+ HWCsh
kk=kk+1
for ii in tracers_3D:
TrOnShwHole = get_metrics('3DDIFF_hole_', ii, '1', fields[0] )
TrOnHole = get_metrics('3DDIFF_hole_', ii, '1', fields[2] )
TrOnSh[:,kk] = TrOnHole + TrOnShwHole
HWCsh = get_metrics('3DDIFF_hole_', ii, '1', fields[1] )
HWChole = get_metrics('3DDIFF_hole_', ii, '1', fields[3] )
HWC[:,kk] = HWChole + HWCsh
kk=kk+1
# +
tracers_3D = ['04','05','06','07'] #run number because there's only 1 tr per run
tracers_CNT03 = ['1','2','3'] # tracer number , constant runs
tracers_CNT09 = ['1','2','3'] # tracer number , average diffusivity runs
#tracers_CNT07 = ['1','2','3'] # tracer number , no canyon case
tracers_CNT02 = ['1','2','3'] # tracer number , Kiso=0.1
tracers_CNT04 = ['1','2','3'] # tracer number , Kiso=10
tracers_CNT10 = ['1','2','3'] # tracer number , Kiso=1
tracers_CNT11 = ['2'] # tracer number , Kiso=1, Ks=Kt=10^4
tracers_CNT12 = ['3'] # tracer number , Kiso=1, Ks=Kt=10^3
# LOAD TRANSPORTS
CS1 = np.zeros((nt-1,numTr))
CS2 = np.zeros((nt-1,numTr))
CS3 = np.zeros((nt-1,numTr))
CS4 = np.zeros((nt-1,numTr))
CS5 = np.zeros((nt-1,numTr))
CS3sb = np.zeros((nt-1,numTr))
AS1 = np.zeros((nt-1,numTr))
AS2 = np.zeros((nt-1,numTr))
LID1 = np.zeros((nt-1,numTr))
LID2 = np.zeros((nt-1,numTr))
CS1a = np.zeros((nt-1,numTr))
CS2a = np.zeros((nt-1,numTr))
CS3a = np.zeros((nt-1,numTr))
CS4a = np.zeros((nt-1,numTr))
CS5a = np.zeros((nt-1,numTr))
CS3sba = np.zeros((nt-1,numTr))
AS1a = np.zeros((nt-1,numTr))
AS2a = np.zeros((nt-1,numTr))
LID1a = np.zeros((nt-1,numTr))
LID2a = np.zeros((nt-1,numTr))
CS1d = np.zeros((nt-1,numTr))
CS2d = np.zeros((nt-1,numTr))
CS3d = np.zeros((nt-1,numTr))
CS4d = np.zeros((nt-1,numTr))
CS5d = np.zeros((nt-1,numTr))
CS3sbd = np.zeros((nt-1,numTr))
AS1d = np.zeros((nt-1,numTr))
AS2d = np.zeros((nt-1,numTr))
LID1d = np.zeros((nt-1,numTr))
LID2d = np.zeros((nt-1,numTr))
kk = 0
fields = ['CS1','CS2','CS3','CS3sb','CS4','CS5','AS1' ,'AS2','LID1' ,'LID2']
fieldsDiff = ['CS1','CS2','CS3','CS3sb','CS4','CS5','AS1' ,'AS2','LID1' ,'LID2','LID1i' ,'LID2i']
for ii in tracers_CNT02:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '02', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '02', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_CNT03:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '03', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '03', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_CNT04:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '04', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[5] )
AS1d[:,kk] =get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '04', ii, fieldsDiff[11] ))
kk=kk+1
#for ii in tracers_CNT07:
# CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[0] )
# CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[1] )
# CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[2] )
# CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[3] )
# CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[4] )
# CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[5] )
# AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[6] )
# AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[7] )
# LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[8] )
# LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '07', ii, fields[9] )
# CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07',ii, fieldsDiff[0] )
# CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[1] )
# CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[2] )
# CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[3] )
# CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[4] )
# CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[5] )
# AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[6] )
# AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[7] )
# LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[8] )
# +get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[10] ))
# LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[9] )
# +get_metrics('CNTDIFF_CS_DIFFFLUX_', '07', ii, fieldsDiff[11] ))
# kk=kk+1
for ii in tracers_CNT09:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '09', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '09', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_CNT10:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '10', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '10', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_CNT11:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '11', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '11', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_CNT12:
CS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[0] )
CS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[1] )
CS3a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[2] )
CS3sba[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[3] )
CS4a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[4] )
CS5a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[5] )
AS1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[6] )
AS2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[7] )
LID1a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[8] )
LID2a[:,kk] = get_metrics('CNTDIFF_CS_ADVFLUX_', '12', ii, fields[9] )
CS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12',ii, fieldsDiff[0] )
CS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[1] )
CS3d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[3] )
CS4d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[4] )
CS5d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[5] )
AS1d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[6] )
AS2d[:,kk] = get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[8] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[9] )
+get_metrics('CNTDIFF_CS_DIFFFLUX_', '12', ii, fieldsDiff[11] ))
kk=kk+1
for ii in tracers_3D:
CS1a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[0] )
CS2a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[1] )
CS3a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[2] )
CS3sba[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[3] )
CS4a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[4] )
CS5a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[5] )
AS1a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[6] )
AS2a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[7] )
LID1a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[8] )
LID2a[:,kk] = get_metrics('3DDIFF_CS_ADVFLUX_', ii, '1', fields[9] )
CS1d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[0] )
CS2d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[1] )
CS3d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[2] )
CS3sbd[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[3] )
CS4d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[4] )
CS5d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[5] )
AS1d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[6] )
AS2d[:,kk] = get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[7] )
LID1d[:,kk] = (get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[8] )
+get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[10] ))
LID2d[:,kk] = (get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[9] )
+get_metrics('3DDIFF_CS_DIFFFLUX_', ii, '1', fieldsDiff[11] ))
kk=kk+1
CS1 = CS1a +CS1d
CS2 = CS2a +CS2d
CS3 = CS3a +CS3d
CS4 = CS4a +CS4d
CS5 = CS5a+ CS5d
CS3sb = CS3sba +CS3sbd
AS1 = AS1a +AS1d
AS2 = AS2a +AS2d
LID1 = LID1a+ LID1d
LID2 = LID2a +LID2d
# +
# LOAD WATER TRANSPORT
numWat = 11
water_3D = ['04','05','06','07'] #run number
water_CNT = ['02','03','04','09','10','11','12'] # run number , constant runs
wCS1 = np.zeros((nt-1,numWat))
wCS2 = np.zeros((nt-1,numWat))
wCS3 = np.zeros((nt-1,numWat))
wCS4 = np.zeros((nt-1,numWat))
wCS5 = np.zeros((nt-1,numWat))
wCS3sb = np.zeros((nt-1,numWat))
wAS1 = np.zeros((nt-1,numWat))
wAS2 = np.zeros((nt-1,numWat))
wLID1 = np.zeros((nt-1,numWat))
wLID2 = np.zeros((nt-1,numWat))
kk = 0
fields = ['CS1','CS2','CS3','CS3sb','CS4','CS5','AS1' ,'AS2','LID1' ,'LID2']
for ii in water_CNT:
wCS1[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[0] )
wCS2[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[1] )
wCS3[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[2] )
wCS3sb[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[3] )
wCS4[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[4] )
wCS5[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[5] )
wAS1[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[6] )
wAS2[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[7] )
wLID1[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[8] )
wLID2[:,kk] = get_water('CNTDIFF_WaterCSTRANS_', ii, fields[9] )
kk=kk+1
for ii in water_3D:
wCS1[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[0] )
wCS2[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[1] )
wCS3[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[2] )
wCS3sb[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[3] )
wCS4[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[4] )
wCS5[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[5] )
wAS1[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[6] )
wAS2[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[7] )
wLID1[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[8] )
wLID2[:,kk] = get_water('3DDIFF_WaterCSTRANS_', ii, fields[9] )
kk=kk+1
# +
### NON-DIMENSIONAL PARAMETERS
kdout = np.array([1.E-5,1.E-4,1.E-3, #run02
1.E-5,1.E-4,1.E-3, #run03
1.E-5,1.E-4,1.E-3, #run04
(3.81)*1.E-5,(2.8)*1.E-5,(1.3)*1.E-5, #run09
(3.81)*1.E-5,(2.8)*1.E-5,(1.3)*1.E-5, #run10
1.E-4, #run11
1.E-3, #run12
1.E-7, #3D run04
1.E-7, #3D run05
1.E-5, #3D run06
1.E-5]) #3D run07
kdTSout = np.array([1.E-5,1.E-5,1.E-5, #run02
1.E-5,1.E-5,1.E-5, #run03
1.E-5,1.E-5,1.E-5, #run04
(1)*1.E-5,(1)*1.E-5,(1)*1.E-5, #run09
(1)*1.E-5,(1)*1.E-5,(1)*1.E-5, #run10
1.E-4, #run11
1.E-3, #run12
1.E-7, #3D run04
1.E-7, #3D run05
1.E-5, #3D run06
1.E-5]) #3D run07
kdcan = np.array([1.E-5,1.E-4,1.E-3, #run02
1.E-5,1.E-4,1.E-3, #run03
1.E-5,1.E-4,1.E-3, #run04
(3.81)*1.E-5,(2.8)*1.E-5,(1.3)*1.E-5, #run09
(3.81)*1.E-5,(2.8)*1.E-5,(1.3)*1.E-5, #run10
1.E-4, #run11
1.E-3, #run12
1.E-3, #3D run04
1.E-4, #3D run05
1.E-3, #3D run06
1.E-4]) #3D run07
ki = np.array([10.0,10.0,10.0, #run02
1.0,1.0,1.0, #run03
0.1,0.1,0.1, #run04
10.0,10.0,10.0, #run09
1.0,1.0,1.0, #run10
1.0, #run11
1.0, #run12
1.0, #3D run04
1.0, #3D run05
1.0, #3D run06
1.0]) #3D run07
Z = np.array([75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,75.5,53.5,59.5,71.5,63.5,75.5])
#Z = 85.0
df = pd.read_csv('results/metricsDataFrames/OmegaUstarCNTand3D_tavg_zavg.csv')
U = df['Ustar']
Om = df['Omega']
Uvec = np.array([U[0],U[0],U[0],U[1],U[1],U[1],U[2],U[2],U[2],U[3],U[3],U[3],U[4],U[4],U[4],U[5],U[6],U[7],U[8],U[9],U[10]])
Omvec = np.array([Om[0],Om[0],Om[0],Om[1],Om[1],Om[1],Om[2],Om[2],Om[2],Om[3],Om[3],Om[3],Om[4],Om[4],Om[4],Om[5],Om[6],Om[7],Om[8],Om[9],Om[10]])
L = 6400.0 # meters
Peh = (L*Uvec)/ki
Pev_can = (Z*Omvec)/kdcan
Pev_TSout = (Z*Omvec)/kdTSout
Pev_out = (Z*Omvec)/kdout
K_out = ((Z**2)*ki)/(L*L*kdout)
K_can = ((Z**2)*ki)/(L*L*kdcan)
K_TScan = ((Z**2)*ki)/(L*L*kdTScan)
# +
vertical = LID1+LID2
total = CS1+CS2+CS3sb+CS4+CS5
verticala = LID1a+LID2a
totala = CS1a+CS2a+CS3sba+CS4a+CS5a
verticald = LID1d+LID2d
totald = CS1d+CS2d+CS3sbd+CS4d+CS5d
watVert = LID1A[0]*1000.0*wLID1 + LID2A[0]*1000.0*wLID2
watTot = (1000.0*CS1A[0]*wCS1+1000.0*CS2A[0]*wCS2+1000.0*CS3sbA[0]*wCS3sb+1000.0*CS4A[0]*wCS4+1000.0*CS5A[0]*wCS5)
## FIGURES ##
sns.set_palette( 'Set1',9)
marker = ['o','o','o','*','*','*','d','d','d','^','^','^','>','>','>','h','.','s','s','s','s']
wmarker = ['o','*','d','^','>','h','.','s','s','s','s']
indexList = [0,1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20]
windexList = [0,1,2,3,4,5,6,7,8,9,10]
transEqIndex = [0,3,6,9,12,15,16,17,18,19,20]
# +
## Pe_v
sns.set_context("talk", font_scale=0.9, rc={"lines.linewidth": 2.5})
fig42=plt.figure(figsize=(17,12))
jj=0
for ii in indexList:
ax1 = plt.subplot(3,3,1)
plt.plot( Pev_out[ii] ,np.mean(TrOnSh[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean tr mass (Mol)')
plt.xlabel('$Pe_{vOut}$')
plt.title(' Tracer in shelf box - advective phase ')
ax1.set_xscale("log", nonposy='clip')
ax2 = plt.subplot(3,3,2)
plt.plot(Pev_out[ii] , np.nanmean(HWC[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean volume ($m^3$)')
plt.xlabel('$Pe_{vOut}$')
plt.title('HCW on cshelf box ')
ax2.set_xscale("log", nonposy='clip')
ax4 = plt.subplot(3,3,4)
plt.plot(Pev_TSout[ii] , np.mean(total[10:,ii]+vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{vTSOut}$')
plt.title('Total transport through shelf box ')
ax4.set_xscale("log", nonposy='clip')
ax7 = plt.subplot(3,3,7)
plt.plot(Pev_TSout[ii] , np.mean(vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{vTSOut}$')
plt.title('Vertical ')
ax7.set_xscale("log", nonposy='clip')
ax8 = plt.subplot(3,3,8)
plt.plot(Pev_TSout[ii] , np.mean(verticala[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{vTSOut}$')
plt.title('Advective, vertical')
ax8.set_xscale("log", nonposy='clip')
ax9 = plt.subplot(3,3,9)
plt.plot(Pev_out[ii] ,np.mean(verticald[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{vOut}$')
plt.title('Diffusive, vertical ')
ax9.set_xscale("log", nonposy='clip')
jj = jj+1
for mm, ii in zip(transEqIndex,windexList):
ax5 = plt.subplot(3,3,5)
plt.plot(Pev_TSout[mm] , np.nanmean(watVert[10:,ii]),wmarker[ii], markersize = 13,alpha = 0.8,label = wlabels[ii])
plt.ylabel('Transport ($m^3/s$)')
plt.xlabel('$Pe_{vOut}$')
plt.title('Vertical transport water ')
ax5.set_xscale("log", nonposy='clip')
plt.tight_layout()
ax2.legend(loc ='upper right', bbox_to_anchor=(2.1,1))
#ax2.legend(loc =0)
plt.show()
#fig42.savefig('results/figures/PevAllMetricsAllShelfAdvPhase.eps', format='eps', dpi=1000, bbox_inches='tight')
# +
## Pe_h Time-dependent phase
sns.set_context("talk", font_scale=0.9, rc={"lines.linewidth": 2.5})
fig43=plt.figure(figsize=(15,12))
jj=0
for ii in indexList:
ax1 = plt.subplot(3,3,1)
plt.plot( Peh[ii] ,np.mean(TrOnSh[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean tr mass (Mol)')
plt.xlabel('$Pe_{h}$')
plt.title(' Tracer in shelf box - Advective phase ')
ax1.set_xscale("log", nonposy='clip')
ax2 = plt.subplot(3,3,2)
plt.plot(Peh[ii] , np.nanmean(HWC[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean volume ($m^3$)')
plt.xlabel('$Pe_{h}$')
plt.title('HCW on shelf box ')
ax2.set_xscale("log", nonposy='clip')
ax4 = plt.subplot(3,3,4)
plt.plot(Peh[ii] , np.mean(total[10:,ii]+vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{h}$')
plt.title('Total transport through shelf box ')
ax4.set_xscale("log", nonposy='clip')
ax7 = plt.subplot(3,3,7)
plt.plot(Peh[ii] , np.mean(vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{h}$')
plt.title('Vertical ')
ax7.set_xscale("log", nonposy='clip')
ax8 = plt.subplot(3,3,8)
plt.plot(Peh[ii] , np.mean(verticala[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{h}$')
plt.title('Advective, vertical')
ax8.set_xscale("log", nonposy='clip')
ax9 = plt.subplot(3,3,9)
plt.plot(Peh[ii] ,np.mean(verticald[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$Pe_{h}$')
plt.title('Diffusive, vertical ')
ax9.set_xscale("log", nonposy='clip')
jj = jj+1
for mm, ii in zip(transEqIndex,windexList):
ax5 = plt.subplot(3,3,5)
plt.plot(Peh[mm] , np.nanmean(watVert[10:,ii]),wmarker[ii], markersize = 13,alpha = 0.8,label = wlabels[ii])
plt.ylabel('Transport ($m^3/s$)')
plt.xlabel('$Pe_{h}$')
plt.title('Vertical transport water ')
ax5.set_xscale("log", nonposy='clip')
plt.tight_layout()
ax2.legend(loc ='upper right', bbox_to_anchor=(2.1,1))
#ax2.legend(loc =0)
plt.show()
#fig43.savefig('results/figures/PehAllMetricsAllShlefAdvPhase.eps', format='eps', dpi=1000, bbox_inches='tight')
# +
#-----------------------------------------------------------------------------------------------------------------------------
## Kappa Time-dependent phase
sns.set_context("talk", font_scale=0.9, rc={"lines.linewidth": 2.5})
fig44=plt.figure(figsize=(15,12))
jj=0
for ii in indexList:
ax1 = plt.subplot(3,3,1)
plt.plot( K_can[ii] ,np.mean(TrOnSh[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean tr mass (Mol)')
plt.xlabel('$\kappa_{can}$')
plt.title(' Tracer in shlef box - Advective phase ')
ax1.set_xscale("log", nonposy='clip')
ax2 = plt.subplot(3,3,2)
plt.plot(K_out[ii] , np.nanmean(HWC[10:,ii]),marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Mean volume ($m^3$)')
plt.xlabel('$\kappa_{out}$')
plt.title('HCW on shelf box ')
ax2.set_xscale("log", nonposy='clip')
ax4 = plt.subplot(3,3,4)
plt.plot(K_out[ii] , np.mean(total[10:,ii]+vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$\kappa_{out}$')
plt.title('Total transport through shelf box ')
ax4.set_xscale("log", nonposy='clip')
ax7 = plt.subplot(3,3,7)
plt.plot(K_can[ii] , np.mean(vertical[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$\kappa_{can}$')
plt.title('Vertical ')
ax7.set_xscale("log", nonposy='clip')
ax8 = plt.subplot(3,3,8)
plt.plot(K_can[ii] , np.mean(verticala[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$\kappa_{can}$')
plt.title('Advective, vertical')
ax8.set_xscale("log", nonposy='clip')
ax9 = plt.subplot(3,3,9)
plt.plot(K_can[ii] ,np.mean(verticald[10:,ii])*1000.0,marker[jj], markersize = 13,alpha = 0.8,label = labels[ii])
plt.ylabel('Tracer transport ($Mol/s$)')
plt.xlabel('$\kappa_{can}$')
plt.title('Diffusive, vertical ')
ax9.set_xscale("log", nonposy='clip')
jj = jj+1
for mm, ii in zip(transEqIndex,windexList):
ax5 = plt.subplot(3,3,5)
plt.plot(K_can[mm] , np.nanmean(watVert[10:,ii]),wmarker[ii], markersize = 13,alpha = 0.8,label = wlabels[ii])
plt.ylabel('Transport ($m^3/s$)')
plt.xlabel('$\kappa_{can}$')
plt.title('Vertical transport water ')
ax5.set_xscale("log", nonposy='clip')
plt.tight_layout()
ax2.legend(loc ='upper right', bbox_to_anchor=(2.1,1))
#ax2.legend(loc =0)
plt.show()
#fig44.savefig('results/figures/KappaAllMetricsAllShelfAdvPhase.eps', format='eps', dpi=1000, bbox_inches='tight')
#-----------------------------------------------------------------------------------------------------------------------------
# -
|
PlotNonDimVsMetricsShelfBox.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Creating Folds & Preprocessing
#
# We will be using Kaggle Kernels to test XGBoost and CatBoost models using GPU and using these local notebooks to test LightGBM models with CPU. We want our cross-validation scheme to be consistent across both so we define it here, save it using `feather` and upload it to Kaggle.
#
# The original data is very large and takes a long time to load. We use some techniques to save memory and also store the data more efficiently.
# Global variables for testing changes to this notebook quickly
FOLD_SEED = 0
MIN_FOLDS = 3
MAX_FOLDS = 6
# +
import os
import warnings
import numpy as np
import pandas as pd
import pyarrow
import time
# cross validation
from sklearn.model_selection import StratifiedKFold
# display options
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
#pd.set_option('float_format', '{:f}'.format)
warnings.filterwarnings('ignore')
# -
# # Loading Times
#
# We benchmark the `read_csv` method with our raw data:
# +
# %%time
train = pd.read_csv('../data/train.csv')
# +
# %%time
test = pd.read_csv('../data/test.csv')
# -
# # Data Size
#
# We also check how large our raw data is in memory:
print("Train:",
round(train.memory_usage().sum() / 1024 ** 2, 2), "Mb")
print("Test:",
round(test.memory_usage().sum() / 1024 ** 2, 2), "Mb")
# # Reduce Memory Usage
#
# We use a helper function to cast the numerical variables (all variables in this months dataset are numerical) to their lowest possible subtype. This idea was adapted from this [Kaggle notebook](https://www.kaggle.com/bextuychiev/how-to-work-w-million-row-datasets-like-a-pro).
# Creates a copy of the original data
def reduce_memory_usage(data, verbose=True):
df = data.copy()
start_mem = df.memory_usage().sum() / 1024 ** 2
for col, dtype in df.dtypes.iteritems():
if dtype.name.startswith('int'):
df[col] = pd.to_numeric(data[col], downcast ='integer')
elif dtype.name.startswith('float'):
df[col] = pd.to_numeric(data[col], downcast ='float')
if np.max(df[col] - data[col]) > 1:
df[col] = data[col]
end_mem = df.memory_usage().sum() / 1024 ** 2
if verbose:
print(
"Mem. usage decreased to {:.2f} Mb ({:.1f}% reduction)".format(
end_mem, 100 * (start_mem - end_mem) / start_mem
)
)
return df
# +
# %%time
train_new = reduce_memory_usage(train)
temp = train - train_new
print("Maximal Difference:", np.max(temp.max()))
# +
# %%time
test_new = reduce_memory_usage(test)
temp = test - test_new
print("Maximal Difference:", np.max(temp.max()))
# -
# # Stratified K-Fold
#
# We use `StratifiedKFold` to define our cross-validation scheme.
# +
# Create Folds
for NUM_FOLDS in range(MIN_FOLDS,MAX_FOLDS+1):
train_new[f"{NUM_FOLDS}fold"] = -1
kf = StratifiedKFold(NUM_FOLDS, shuffle = True, random_state = FOLD_SEED)
for fold, (train_idx, valid_idx) in enumerate(kf.split(train_new, train_new['claim'])):
train_new.loc[valid_idx,f"{NUM_FOLDS}fold"] = fold
# check output
train_new.head()
# -
# # Saving Output
#
# We want to save our output in a format that is easy to load and retains our memory usage savings. To this end we use the feather format in pandas.
# +
path_train = '../data/train.feather'
path_test = '../data/test.feather'
# save data
train_new.to_feather(path_train)
test_new.to_feather(path_test)
# -
# # Sanity Checks
#
# We check the following:
#
# 1. Speed at which we can load `.feather` files
# 2. Our `.feather` data is equivalent to the original data
# ## Loading Times
# +
# %%time
# reload data (for testing purposes)
train_df = pd.read_feather(path_train)
test_df = pd.read_feather(path_test)
# -
# ## Data Equivalence
# +
# Check train data types are preserved
types_1 = [train_new[x].dtype for x in train.columns]
types_2 = [train_df[x].dtype for x in train.columns]
assert types_1 == types_2
# Check test data types are preserved
types_1 = [test_new[x].dtype for x in test_new.columns]
types_2 = [test_df[x].dtype for x in test_df.columns]
assert types_1 == types_2
# Find the difference between the data
temp = train_new - train_df
print("Largest Difference (Train):", temp.max().max())
# Find the difference between the data
temp = test_new - test_df
print("Largest Difference (Test):", temp.max().max())
|
tps-2021-09/notebooks/Notebook 1 - Preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Aula 3 Video 1 - Entendendo os dados e o problema
import pandas as pd
from sklearn.model_selection import train_test_split
from collections import Counter
# Download Dataset: [avaliacoes_usuario.csv](https://s3.amazonaws.com/caelum-online-public/machine-learning-aprendizado-supervisionado/avaliacoes_usuario.csv)
movies = pd.read_csv('datasets/avaliacoes_usuario.csv')
movies.head(5)
Counter(movies['Gostou'])
print('Gostou {}'.format((543/(135+543))))
print('Não gostou {}'.format((135/(135+543))))
caract = movies[movies.columns[1:16]]
gostos = movies[movies.columns[16:]]
#treino, teste, treino_labels, teste_labels = train_test_split(caract, gostos, test_size=0.1)
treino, teste, treino_labels, teste_labels = train_test_split(caract, gostos)
type(treino)
Counter(treino_labels['Gostou'])
Counter(teste_labels['Gostou'])
print('Gostou % {} do treino_label (deve manter a proporção +- 80%)'.format(404/(104+404)))
print('Gostou % {} do teste_label (deve manter a proporção +- 80%)'.format(139/(31+139)))
# ## Aula 3 Video 2 - A regressão logística
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score
import numpy as np
from sklearn.naive_bayes import MultinomialNB,GaussianNB
type(treino)
treino[0:1]
treino = np.array(treino).reshape(len(treino), 15)
teste = np.array(teste).reshape(len(teste), 15)
type(treino)
treino[0]
type(treino_labels)
treino_labels.head(5)
treino_labels = treino_labels.values.ravel()
teste_labels = teste_labels.values.ravel()
type(treino_labels)
treino_labels[0:5]
treino_labels.shape
# #### Usando regressão logistica
modelo = LogisticRegression()
modelo.fit(treino, treino_labels)
previsoes = modelo.predict(teste)
previsoes
teste_labels.shape
previsoes.shape
acuracia = accuracy_score(teste_labels, previsoes)
acuracia
zootopia = [0,0,0,0,0,0,0,1,1,1,1,0,1,110,27.74456356]
modelo.predict([zootopia])
# #### Usando Multinomial Naive Bayes
modelo_NB = MultinomialNB()
modelo_NB.fit(treino,treino_labels)
previsoes_NB = modelo_NB.predict(teste)
acuracia = accuracy_score(teste_labels, previsoes_NB)
acuracia
zootopia = [0,0,0,0,0,0,0,1,1,1,1,0,1,110,27.74456356]
modelo_NB.predict([zootopia])
# #### EXTRA: Usando Gaussian Naive Bayes
modelo_GNB = GaussianNB()
modelo_GNB.fit(treino,treino_labels)
previsoes_GNB = modelo_GNB.predict(teste)
acuracia = accuracy_score(teste_labels, previsoes_GNB)
acuracia
zootopia = [0,0,0,0,0,0,0,1,1,1,1,0,1,110,27.74456356]
modelo_GNB.predict([zootopia])
|
Alura/MLAprendizadoSupervisionado/626-jupyter-notebooks/626-Aula3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # ODaly's Driverless Cars
# %run rallyman_internals.ipynb
show(grid([
plot_occurences(dc_champion.movements(), label="champion", color="red", range=MOVEMENT_RANGE),
plot_occurences(dc_veteran.movements(), label="veteran", color="coral", range=MOVEMENT_RANGE),
plot_occurences(dc_midfielder.movements(), label="midfielder", color="orange", range=MOVEMENT_RANGE),
plot_occurences(dc_silver.movements(), label="silver", color="yellow", range=MOVEMENT_RANGE),
plot_occurences(dc_rookie.movements(), label="rookie", color="green", range=MOVEMENT_RANGE),
plot_occurences(dc_amateur.movements(), label="amateur", color="blue", range=MOVEMENT_RANGE)
]))
|
driverless_cars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/John-G-Thomas/DS-Unit-2-Kaggle-Challenge/blob/master/module2-random-forests/LS_DS_222.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="UEHyIgGaaSEJ"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 2*
#
# ---
# + [markdown] colab_type="text" id="mRfPLX4WgLVJ"
# # Random Forests
# + [markdown] colab_type="text" id="jRRNhkxcgLVK"
# - use scikit-learn for **random forests**
# - do **ordinal encoding** with high-cardinality categoricals
# - understand how categorical encodings affect trees differently compared to linear models
# - understand how tree ensembles reduce overfitting compared to a single decision tree with unlimited depth
# + [markdown] colab_type="text" id="-3TH11e1gLVL"
# Today's lesson has two take-away messages:
#
# #### Try Tree Ensembles when you do machine learning with labeled, tabular data
# - "Tree Ensembles" means Random Forest or Gradient Boosting models.
# - [Tree Ensembles often have the best predictive accuracy](https://arxiv.org/abs/1708.05070) with labeled, tabular data.
# - Why? Because trees can fit non-linear, non-[monotonic](https://en.wikipedia.org/wiki/Monotonic_function) relationships, and [interactions](https://christophm.github.io/interpretable-ml-book/interaction.html) between features.
# - A single decision tree, grown to unlimited depth, will [overfit](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/). We solve this problem by ensembling trees, with bagging (Random Forest) or boosting (Gradient Boosting).
# - Random Forest's advantage: may be less sensitive to hyperparameters. Gradient Boosting's advantage: may get better predictive accuracy.
#
# #### One-hot encoding isn’t the only way, and may not be the best way, of categorical encoding for tree ensembles.
# - For example, tree ensembles can work with arbitrary "ordinal" encoding! (Randomly assigning an integer to each category.) Compared to one-hot encoding, the dimensionality will be lower, and the predictive accuracy may be just as good or even better.
#
# + [markdown] colab_type="text" id="r5PbOFEuFfGF"
# ### Setup
#
# Run the code cell below. You can work locally (follow the [local setup instructions](https://lambdaschool.github.io/ds/unit2/local/)) or on Colab.
#
# Libraries
#
# - **category_encoders**
# - **graphviz**
# - ipywidgets
# - matplotlib
# - numpy
# - pandas
# - seaborn
# - scikit-learn
# + colab_type="code" id="FStAplyRFoEu" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + [markdown] colab_type="text" id="ZL-yK8B7gLVW"
# # Use scikit-learn for random forests
# + [markdown] id="E1xsazz7BtC3" colab_type="text"
# ## Overview
#
# Let's fit a Random Forest!
#
# 
#
# [<NAME>, MachineLearningFlashcards.com](https://twitter.com/chrisalbon/status/1181261589887909889)
# + [markdown] colab_type="text" id="gHFxMCPSgLVM"
# ### Solution example
#
# First, read & wrangle the data.
#
# > Define a function to wrangle train, validate, and test sets in the same way. Clean outliers and engineer features. (For example, [what other columns have zeros and shouldn't?](https://github.com/Quartz/bad-data-guide#zeros-replace-missing-values) What other columns are duplicates, or nearly duplicates? Can you extract the year from date_recorded? Can you engineer new features, such as the number of years from waterpump construction to waterpump inspection?)
# + colab_type="code" id="YTLm-rDagLVM" colab={}
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# Split train into train & val
train, val = train_test_split(train, train_size=0.80, test_size=0.20,
stratify=train['status_group'], random_state=42)
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# + colab_type="code" id="m2HppBvZgLVP" colab={}
# The status_group column is the target
target = 'status_group'
# Get a dataframe with all train columns except the target
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
# Get a series with the cardinality of the nonnumeric features
cardinality = train_features.select_dtypes(exclude='number').nunique()
# Get a list of all categorical features with cardinality <= 50
categorical_features = cardinality[cardinality <= 50].index.tolist()
# Combine the lists
features = numeric_features + categorical_features
# + colab_type="code" id="aXmK2brXgLVR" colab={}
# Arrange data into X features matrix and y target vector
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
# + [markdown] id="WQwrzPlxBtDB" colab_type="text"
# ## Follow Along
#
# [Scikit-Learn User Guide: Random Forests](https://scikit-learn.org/stable/modules/ensemble.html#random-forests)
# + colab_type="code" id="57yyygsdgLVW" colab={}
# TODO
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.pipeline import Pipeline
from sklearn.feature_selection import SelectKBest
from category_encoders import OneHotEncoder, OrdinalEncoder
from sklearn.impute import SimpleImputer
# + id="G9-kan7pN5Xf" colab_type="code" colab={}
model = Pipeline([
('ohe', OneHotEncoder()),
('impute', SimpleImputer()),
('classifier', DecisionTreeClassifier())
])
# + id="gKt4jhq-OOF-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="36f9be97-17ef-47c2-9f8e-1839e0884603"
# %%time
model.fit(X_train, y_train)
print('training accuracy:', model.score(X_train, y_train))
print('validation accuracy:', model.score(X_val, y_val))
# + id="k6WcWJdfPgwI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="e6682460-001a-450c-e6f2-b95f83cae5f8"
# %%time
model = Pipeline([
('ohe', OneHotEncoder()),
('impute', SimpleImputer()),
('select', SelectKBest(k=20)),
('classifier', DecisionTreeClassifier())
])
model.fit(X_train, y_train)
print('training accuracy:', model.score(X_train, y_train))
print('validation accuracy:', model.score(X_val, y_val))
# + id="gwptatvXPDUF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="722ca8d1-7db6-48cb-e2bb-239b0826050e"
model.named_steps['classifier'].get_depth()
# + id="ulDnFHlpQ_jo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1f235443-b9a8-4e8d-8124-7c43eee8e0c9"
model.named_steps['classifier'].get_depth()
# + id="NfyXD36mR6G6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="fc1e1f6b-efa3-4a83-8b9a-6ea5f22164f8"
ohe = model.named_steps['ohe']
X_trans = ohe.transform(X_train)
print(X_train.shape)
print(X_trans.shape)
# + id="bABZnP1EUDcR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="f251bea0-beee-49a6-a1cd-9fc081a2b0c8"
# %%time
model = Pipeline([
('ohe', OrdinalEncoder()),
('impute', SimpleImputer()),
('classifier', DecisionTreeClassifier())
])
model.fit(X_train, y_train)
print('training accuracy:', model.score(X_train, y_train))
print('validation accuracy:', model.score(X_val, y_val))
# + id="XEm8JLTCWeBL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="0832dcd7-f835-4446-abd8-0d7b0c946a73"
# %%time
model = Pipeline([
('ohe', OneHotEncoder()),
('impute', SimpleImputer()),
('classifier', RandomForestClassifier(n_jobs=-1))
])
model.fit(X_train, y_train)
print('training accuracy:', model.score(X_train, y_train))
print('validation accuracy:', model.score(X_val, y_val))
# + [markdown] colab_type="text" id="yfyk_aa5gLVY"
# # Do ordinal encoding with high-cardinality categoricals
# + [markdown] id="UxtCY11gBtDG" colab_type="text"
# ## Overview
#
# https://contrib.scikit-learn.org/category_encoders/ordinal.html
# + [markdown] id="Hp2VoefdBtDG" colab_type="text"
# ## Follow Along
# + colab_type="code" id="b8d_WJtcgLVZ" colab={}
# TODO
# + [markdown] colab_type="text" id="xs2UPoVdgLVp"
# # Understand how categorical encodings affect trees differently compared to linear models
# + [markdown] id="98S2I4etBtDK" colab_type="text"
# ## Follow Along
# + [markdown] colab_type="text" id="z8V-A92mgLVp"
# ### Categorical exploration, 1 feature at a time
#
# Change `feature`, then re-run these cells!
# + colab_type="code" id="G35RAzVdgLVq" colab={}
feature = 'extraction_type_class'
# + colab_type="code" id="OuxHWiH8gLVr" colab={"base_uri": "https://localhost:8080/", "height": 151} outputId="bec48b9f-6248-47fc-8d4c-03d9c0a01f68"
X_train[feature].value_counts()
# + id="HctGcH9QdAbQ" colab_type="code" colab={}
import matplotlib.pyplot as plt
# + colab_type="code" id="pVxoC4NngLVt" colab={"base_uri": "https://localhost:8080/", "height": 434} outputId="32c4087b-f419-49ef-bedd-e3b5d04b5afa"
import seaborn as sns
plt.figure(figsize=(16,9))
sns.barplot(
x=train[feature],
y=train['status_group']=='functional',
color='grey'
);
# + colab_type="code" id="w99mek14gLVv" colab={"base_uri": "https://localhost:8080/", "height": 370} outputId="04f80999-ed31-4ab5-c916-9c17f5732cf5"
X_train[feature].head(20)
# + id="MkfDPY6BfKZK" colab_type="code" colab={}
import numpy as np
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.tree import DecisionTreeClassifier
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
import matplotlib.pyplot as plt
import category_encoders as ce
from sklearn.ensemble import RandomForestClassifier
from sklearn.impute import SimpleImputer
from sklearn.pipeline import make_pipeline
from sklearn.decomposition import PCA
# + [markdown] colab_type="text" id="ezzK2IdbgLVx"
# ### [One Hot Encoding](https://contrib.scikit-learn.org/category_encoders/onehot.html)
#
# > Onehot (or dummy) coding for categorical features, produces one feature per category, each binary.
#
# Warning: May run slow, or run out of memory, with high cardinality categoricals!
# + colab_type="code" id="HDQZtV6GgLVy" colab={"base_uri": "https://localhost:8080/", "height": 693} outputId="b2da7a75-f2dc-43d2-d6d0-d0ff532f364f"
encoder = ce.OneHotEncoder(use_cat_names=True)
encoded = encoder.fit_transform(X_train[[feature]])
print(f'{len(encoded.columns)} columns')
encoded.head(20)
# + [markdown] colab_type="text" id="1Ql9Qmw3sNJ7"
# #### One-Hot Encoding, Logistic Regression, Validation Accuracy
# + colab_type="code" id="mT4A-oDGpOss" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fc6e56cf-8d5f-4b11-b0bf-ef855de89ff0"
from sklearn.linear_model import LogisticRegressionCV
from sklearn.preprocessing import StandardScaler
lr = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
StandardScaler(),
LogisticRegressionCV(multi_class='auto', solver='lbfgs', cv=5, n_jobs=-1)
)
lr.fit(X_train[[feature]], y_train)
score = lr.score(X_val[[feature]], y_val)
print('Logistic Regression, Validation Accuracy', score)
# + [markdown] colab_type="text" id="EbH6wivpsRuV"
# #### One-Hot Encoding, Decision Tree, Validation Accuracy
# + colab_type="code" id="b6KUluFOqIdK" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="55df7128-3077-4520-fcb3-ea87e694ae73"
from sklearn.tree import DecisionTreeClassifier
dt = make_pipeline(
ce.OneHotEncoder(use_cat_names=True),
SimpleImputer(),
DecisionTreeClassifier(random_state=42)
)
dt.fit(X_train[[feature]], y_train)
score = dt.score(X_val[[feature]], y_val)
print('Decision Tree, Validation Accuracy', score)
# + [markdown] colab_type="text" id="8yg11_gTsUu6"
# #### One-Hot Encoding, Logistic Regression, Model Interpretation
# + colab_type="code" id="IxHwXGRornNI" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="62aa1c3c-3531-448b-b5b4-671ff8a68141"
model = lr.named_steps['logisticregressioncv']
encoder = lr.named_steps['onehotencoder']
encoded_columns = encoder.transform(X_val[[feature]]).columns
coefficients = pd.Series(model.coef_[0], encoded_columns)
coefficients.sort_values().plot.barh(color='grey');
# + [markdown] colab_type="text" id="0REZ8HdpsccR"
# #### One-Hot Encoding, Decision Tree, Model Interpretation
# + colab_type="code" id="gV-grmYKpDp9" colab={"base_uri": "https://localhost:8080/", "height": 954} outputId="6e67b9a3-3fc6-4235-be32-32b7ad55e821"
# Plot tree
# https://scikit-learn.org/stable/modules/generated/sklearn.tree.export_graphviz.html
import graphviz
from sklearn.tree import export_graphviz
model = dt.named_steps['decisiontreeclassifier']
encoder = dt.named_steps['onehotencoder']
encoded_columns = encoder.transform(X_val[[feature]]).columns
dot_data = export_graphviz(model,
out_file=None,
max_depth=7,
feature_names=encoded_columns,
class_names=model.classes_,
impurity=False,
filled=True,
proportion=True,
rounded=True)
display(graphviz.Source(dot_data))
# + [markdown] colab_type="text" id="QUd6gzcZgLVz"
# ### [Ordinal Encoding](https://contrib.scikit-learn.org/category_encoders/ordinal.html)
#
# > Ordinal encoding uses a single column of integers to represent the classes. An optional mapping dict can be passed in; in this case, we use the knowledge that there is some true order to the classes themselves. Otherwise, the classes are assumed to have no true order and integers are selected at random.
# + colab_type="code" id="CnBz2RbwgLVz" colab={"base_uri": "https://localhost:8080/", "height": 656} outputId="aea74b5b-b2bc-49fc-c9a7-96b1c38959be"
encoder = ce.OrdinalEncoder()
encoded = encoder.fit_transform(X_train[[feature]])
print(f'1 column, {encoded[feature].nunique()} unique values')
encoded.head(20)
# + [markdown] colab_type="text" id="Nd-ZWprasqUM"
# #### Ordinal Encoding, Logistic Regression, Validation Accuracy
# + colab_type="code" id="GJ1YpwjvrhfL" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6145a00f-85b9-4e04-afd1-1cf26bd39fe3"
lr = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
StandardScaler(),
LogisticRegressionCV(multi_class='auto', solver='lbfgs', cv=5, n_jobs=-1)
)
lr.fit(X_train[[feature]], y_train)
score = lr.score(X_val[[feature]], y_val)
print('Logistic Regression, Validation Accuracy', score)
# + [markdown] colab_type="text" id="9lO_R3SksuHs"
# #### Ordinal Encoding, Decision Tree, Validation Accuracy
# + colab_type="code" id="aOELD_roriVI" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e7bea5bd-04d5-415e-fa79-2dc1b7bad353"
dt = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
DecisionTreeClassifier(random_state=42)
)
dt.fit(X_train[[feature]], y_train)
score = dt.score(X_val[[feature]], y_val)
print('Decision Tree, Validation Accuracy', score)
# + [markdown] colab_type="text" id="7V2zHjiwswTg"
# #### Ordinal Encoding, Logistic Regression, Model Interpretation
# + colab_type="code" id="S9UPYPois8QR" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="612980d3-47fc-495c-f9c3-9a3d22be8e17"
model = lr.named_steps['logisticregressioncv']
encoder = lr.named_steps['ordinalencoder']
encoded_columns = encoder.transform(X_val[[feature]]).columns
coefficients = pd.Series(model.coef_[0], encoded_columns)
coefficients.sort_values().plot.barh(color='grey');
# + [markdown] colab_type="text" id="MvmmvE8fsymh"
# #### Ordinal Encoding, Decision Tree, Model Interpretation
# + colab_type="code" id="jCvUu4Oms88b" colab={"base_uri": "https://localhost:8080/", "height": 816} outputId="1f353fd6-5e5e-4dc7-9bc3-2be97f743d76"
model = dt.named_steps['decisiontreeclassifier']
encoder = dt.named_steps['ordinalencoder']
encoded_columns = encoder.transform(X_val[[feature]]).columns
dot_data = export_graphviz(model,
out_file=None,
max_depth=5,
feature_names=encoded_columns,
class_names=model.classes_,
impurity=False,
filled=True,
proportion=True,
rounded=True)
display(graphviz.Source(dot_data))
# + [markdown] colab_type="text" id="P4EJi2GvgLVa"
# # Understand how tree ensembles reduce overfitting compared to a single decision tree with unlimited depth
# + [markdown] id="uzMWiIRfBtD4" colab_type="text"
# ## Overview
# + [markdown] colab_type="text" id="0nNABF3HgLVg"
# ### What's "random" about random forests?
# 1. Each tree trains on a random bootstrap sample of the data. (In scikit-learn, for `RandomForestRegressor` and `RandomForestClassifier`, the `bootstrap` parameter's default is `True`.) This type of ensembling is called Bagging. (Bootstrap AGGregatING.)
# 2. Each split considers a random subset of the features. (In scikit-learn, when the `max_features` parameter is not `None`.)
#
# For extra randomness, you can try ["extremely randomized trees"](https://scikit-learn.org/stable/modules/ensemble.html#extremely-randomized-trees)!
#
# >In extremely randomized trees (see [ExtraTreesClassifier](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.ExtraTreesClassifier.html) and [ExtraTreesRegressor](https://scikit-learn.org/stable/modules/generated/sklearn.ensemble.ExtraTreesRegressor.html) classes), randomness goes one step further in the way splits are computed. As in random forests, a random subset of candidate features is used, but instead of looking for the most discriminative thresholds, thresholds are drawn at random for each candidate feature and the best of these randomly-generated thresholds is picked as the splitting rule. This usually allows to reduce the variance of the model a bit more, at the expense of a slightly greater increase in bias
# + [markdown] id="qyRddy9qBtD5" colab_type="text"
# ## Follow Along
# + [markdown] colab_type="text" id="pUYP619CgLVb"
# ### Example: [predicting golf putts](https://statmodeling.stat.columbia.edu/2008/12/04/the_golf_puttin/)
# (1 feature, non-linear, regression)
# + colab_type="code" id="b4640ukxgLVc" colab={}
putts = pd.DataFrame(
columns=['distance', 'tries', 'successes'],
data = [[2, 1443, 1346],
[3, 694, 577],
[4, 455, 337],
[5, 353, 208],
[6, 272, 149],
[7, 256, 136],
[8, 240, 111],
[9, 217, 69],
[10, 200, 67],
[11, 237, 75],
[12, 202, 52],
[13, 192, 46],
[14, 174, 54],
[15, 167, 28],
[16, 201, 27],
[17, 195, 31],
[18, 191, 33],
[19, 147, 20],
[20, 152, 24]]
)
putts['rate of success'] = putts['successes'] / putts['tries']
putts_X = putts[['distance']]
putts_y = putts['rate of success']
# + colab_type="code" id="T0IpCcKggLVd" colab={"base_uri": "https://localhost:8080/", "height": 637, "referenced_widgets": ["c1e34079a2084ab3854a7988e7ed2ade", "e36e23878f8241f1b941c96f4a6fa7d0", "0e352bd2e0d84c959461e12295b30169", "5430d677c998434094dd7de04efbdeaa", "0a43a5f6fe8b4e05a15482d3c7b518c2", "8a0abac4fcd946eab2af110ab07abee6", "695185543cc84bef9e6b09c944351e55", "<KEY>", "f3e7161de13b481ab7be114a4325014c", "ffa1af1ab0d042bf8fc86be66487db64"]} outputId="0658bb38-7d97-4fd1-80dd-c0b9ec1245c0"
# %matplotlib inline
import matplotlib.pyplot as plt
from ipywidgets import interact
from sklearn.ensemble import RandomForestRegressor
from sklearn.tree import DecisionTreeRegressor
def putt_trees(max_depth=1, n_estimators=1):
models = [DecisionTreeRegressor(max_depth=max_depth),
RandomForestRegressor(max_depth=max_depth, n_estimators=n_estimators)]
for model in models:
name = model.__class__.__name__
model.fit(putts_X, putts_y)
ax = putts.plot('distance', 'rate of success', kind='scatter', title=name)
ax.step(putts_X, model.predict(putts_X), where='mid')
plt.show()
interact(putt_trees, max_depth=(1,6,1), n_estimators=(10,40,10));
# + [markdown] colab_type="text" id="rq4Z_wQ_gLVj"
# ### Go back to Tanzania Waterpumps ...
# + [markdown] colab_type="text" id="FoSE9iT6YXQz"
# #### Helper function to visualize predicted probabilities
#
#
# + colab_type="code" id="HzIAjGpJgLVj" colab={}
import itertools
import seaborn as sns
def pred_heatmap(model, X, features, class_index=-1, title='', num=100):
"""
Visualize predicted probabilities, for classifier fit on 2 numeric features
Parameters
----------
model : scikit-learn classifier, already fit
X : pandas dataframe, which was used to fit model
features : list of strings, column names of the 2 numeric features
class_index : integer, index of class label
title : string, title of plot
num : int, number of grid points for each feature
Returns
-------
y_pred_proba : numpy array, predicted probabilities for class_index
"""
feature1, feature2 = features
min1, max1 = X[feature1].min(), X[feature1].max()
min2, max2 = X[feature2].min(), X[feature2].max()
x1 = np.linspace(min1, max1, num)
x2 = np.linspace(max2, min2, num)
combos = list(itertools.product(x1, x2))
y_pred_proba = model.predict_proba(combos)[:, class_index]
pred_grid = y_pred_proba.reshape(num, num).T
table = pd.DataFrame(pred_grid, columns=x1, index=x2)
sns.heatmap(table, vmin=0, vmax=1)
plt.xticks([])
plt.yticks([])
plt.xlabel(feature1)
plt.ylabel(feature2)
plt.title(title)
plt.show()
return y_pred_proba
# + [markdown] colab_type="text" id="DiRfPqHjgLVl"
# ### Compare Decision Tree, Random Forest, Logistic Regression
# + colab_type="code" id="HKkMLXhMgLVl" colab={"base_uri": "https://localhost:8080/", "height": 864, "referenced_widgets": ["81f73c853ed643538e5c519d8f6b185b", "<KEY>", "0bd81b2db5f54c10b6042bc857e14cc1", "3434fa1cf3b443318f3e245b676629ce", "375cc66ab3944428b493c2e84b23b162", "<KEY>", "ac4237663e324fe6847de3ec27aebe8c", "e534a9b0253b40aaa16404097986cbc4", "<KEY>", "a35eeeeced4a42c9b0455f28907624cd"]} outputId="60f6e5bc-e650-44a1-c024-bf6e355bd1cc"
# Instructions
# 1. Choose two features
# 2. Run this code cell
# 3. Interact with the widget sliders
feature1 = 'longitude'
feature2 = 'quantity'
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
def get_X_y(df, feature1, feature2, target):
features = [feature1, feature2]
X = df[features]
y = df[target]
X = X.fillna(X.median())
X = OrdinalEncoder().fit_transform(X)
return X, y
def compare_models(max_depth=1, n_estimators=1):
models = [DecisionTreeClassifier(max_depth=max_depth),
RandomForestClassifier(max_depth=max_depth, n_estimators=n_estimators),
LogisticRegression(solver='lbfgs', multi_class='auto')]
for model in models:
name = model.__class__.__name__
model.fit(X, y)
pred_heatmap(model, X, [feature1, feature2], class_index=0, title=name)
X, y = get_X_y(train, feature1, feature2, target='status_group')
interact(compare_models, max_depth=(1,6,1), n_estimators=(10,40,10));
# + [markdown] colab_type="text" id="hOQqjLEDgLVn"
# ### Bagging
# + colab_type="code" id="Hm4aPgs2gLVn" colab={"base_uri": "https://localhost:8080/", "height": 864, "referenced_widgets": ["8008023b4f594171937124b79a042597", "11e3d86a493e47d8be0b8f74fb77f578", "c780736eda7c4e7980011a56fe543132", "82c8ac238a28404ca0ec052416e0fd0a", "<KEY>", "03a9fe0adb704add8e8baa6d8033db50", "1e40a1f65df24015a348d12fc0df8f1d", "<KEY>", "a1304aa6fd024221b84a2ac9419bf1e2", "835d85d01233478db266565741abb25b"]} outputId="08b5df0f-e85c-4faf-a354-4846852aa977"
# Do-it-yourself Bagging Ensemble of Decision Trees (like a Random Forest)
# Instructions
# 1. Choose two features
# 2. Run this code cell
# 3. Interact with the widget sliders
feature1 = 'longitude'
feature2 = 'latitude'
def waterpumps_bagging(max_depth=1, n_estimators=1):
predicteds = []
for i in range(n_estimators):
title = f'Tree {i+1}'
bootstrap_sample = train.sample(n=len(train), replace=True)
X, y = get_X_y(bootstrap_sample, feature1, feature2, target='status_group')
tree = DecisionTreeClassifier(max_depth=max_depth)
tree.fit(X, y)
predicted = pred_heatmap(tree, X, [feature1, feature2], class_index=0, title=title)
predicteds.append(predicted)
ensembled = np.vstack(predicteds).mean(axis=0)
title = f'Ensemble of {n_estimators} trees, with max_depth={max_depth}'
sns.heatmap(ensembled.reshape(100, 100).T, vmin=0, vmax=1)
plt.title(title)
plt.xlabel(feature1)
plt.ylabel(feature2)
plt.xticks([])
plt.yticks([])
plt.show()
interact(waterpumps_bagging, max_depth=(1,6,1), n_estimators=(2,5,1));
# + [markdown] colab_type="text" id="wYoSBi15akWP"
# # Review
#
# #### Try Tree Ensembles when you do machine learning with labeled, tabular data
# - "Tree Ensembles" means Random Forest or Gradient Boosting models.
# - [Tree Ensembles often have the best predictive accuracy](https://arxiv.org/abs/1708.05070) with labeled, tabular data.
# - Why? Because trees can fit non-linear, non-[monotonic](https://en.wikipedia.org/wiki/Monotonic_function) relationships, and [interactions](https://christophm.github.io/interpretable-ml-book/interaction.html) between features.
# - A single decision tree, grown to unlimited depth, will [overfit](http://www.r2d3.us/visual-intro-to-machine-learning-part-1/). We solve this problem by ensembling trees, with bagging (Random Forest) or boosting (Gradient Boosting).
# - Random Forest's advantage: may be less sensitive to hyperparameters. Gradient Boosting's advantage: may get better predictive accuracy.
#
# #### One-hot encoding isn’t the only way, and may not be the best way, of categorical encoding for tree ensembles.
# - For example, tree ensembles can work with arbitrary "ordinal" encoding! (Randomly assigning an integer to each category.) Compared to one-hot encoding, the dimensionality will be lower, and the predictive accuracy may be just as good or even better.
#
|
module2-random-forests/LS_DS_222.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from gs_quant.common import Currency, PayReceive
from gs_quant.instrument import IRSwap
from gs_quant.markets import PricingContext, OverlayMarket, MarketDataCoordinate
from gs_quant.session import GsSession
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
GsSession.use(client_id=None, client_secret=None, scopes=('run_analytics',))
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Retrieve market data
# + [markdown] pycharm={"name": "#%% md\n"}
# Let's create a swaption and retrieve the market data our instrument is sensitive to. To do so we can call `market()` on
# our instrument. This will give us the `OverlayMarket` object which contains the market data used to price our
# instrument.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
swap = IRSwap(PayReceive.Receive, '10y', Currency.EUR, fixed_rate=-0.025)
swap.resolve()
market = swap.market()
print(f'Base price: {swap.price()}')
# + [markdown] pycharm={"name": "#%% md\n"}
# Then, using the `market_data` attribute, we can access the market data coordinates and values directly
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
print(f'The value of the coordinate, {market.market_data[0].coordinate} is {market.market_data[0].value}')
# + [markdown] pycharm={"name": "#%% md\n"}
# ### Overwrite market data
# + [markdown] pycharm={"name": "#%% md\n"}
# We can also amend the market data of our instrument's `OverlayMarket` to pass-in our own market data value.
# To do so, we simply overwrite the `MarketDataCoordinate` of the instrument `OverlayMarket` to a given value.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
c_10y = MarketDataCoordinate.from_string('IR_EUR_SWAP_10Y.ATMRATE')
print(f'Current value of the EUR 10yr swap point is {market[c_10y]}')
market[c_10y] = -0.02
print(f'New value of the EUR 10yr swap point is {market[c_10y]}')
with PricingContext(market=market):
price_f = swap.price()
print(f'New price: {price_f.result()}')
# + [markdown] pycharm={"name": "#%% md\n"}
# ... or pass in an new `OverlayMarket` all together! Here we create a bespoke market with our own values for the 3m5y
# implied volatility and 10y swap rate. Note that the values that are not overwritten will be defaulted to their original
# value.
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
from gs_quant.instrument import IRSwaption
swaption = IRSwaption(PayReceive.Receive, '5y', Currency.EUR, expiration_date='3m')
swaption.resolve()
print(f'Base price: {swaption.price()}')
vol_3m5y = MarketDataCoordinate.from_string('IR VOL_EUR-EURIBOR-TELERATE_SWAPTION_5Y,3M')
market_data = {c_10y: 0.01, vol_3m5y: 40 / 1e4}
new_market = OverlayMarket(market_data)
with PricingContext(market=new_market):
price_f = swaption.price()
print(f'Price from new market data: {price_f.result()}')
|
gs_quant/documentation/02_pricing_and_risk/00_instruments_and_measures/examples/00_instrument_basics/0005_market-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
import json
import math
import os
import csv
from collections import Counter
from enum import Enum
username = 'login'
senha = 'psw'
class Operations(Enum):
Login = 'login'
Logout = 'logout'
DatasetFields = 'datasetfields'
Search = 'search'
DownloadOptions = 'downloadoptions'
Download = 'download'
IdLookUp = 'idlookup'
MetaData = 'metadata'
UpdateBulk = 'updatebulkdownloadscene'
SubmitBulk = 'submitbulkdownloadorder'
'a' + Operations.Login.value
# +
defaultUrl = 'https://earthexplorer.usgs.gov/inventory/json/v/1.4.0/'
legacyUrl = 'https://earthexplorer.usgs.gov/inventory/json/v/1.4.0/'
operations = {'login': 'login',
'logout' : 'logout',
'datasetfields' : 'datasetfields',
'search' : 'search',
'downloadoptions' : 'downloadoptions',
'download' : 'download',
'idlookup' : 'idlookup',
'metadata' : 'metadata',
'updateBulk' : 'updatebulkdownloadscene',
'submitBulk' : 'submitbulkdownloadorder'
}
productTypes = {'full' : 'STANDARD',
'preview' : 'FRB'
}
downloadIndexes = {'displayId' : 0,
'downloadUrl' : 1,
'entityId' : 2,
'startTime' : 3,
'endTime' : 4
}
# -
def login(usr,pwd, catalog = 'EE'):
payload = {"jsonRequest" : '{"username":"' + usr + '","password":"' + <PASSWORD> + '","catalogId":"' + catalog + '"}'}
response = requests.post(defaultUrl + operations['login'],data = payload)
if response.json()['errorCode'] is None:
return response.json()['data']
else:
print("Login inválido!")
return None
def logout(apiKey):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey+ '"}'}
return requests.post(defaultUrl + operations['logout'],data=payload).json()['data']
def calculateMBR(point,radius):
coef = radius * 0.0000089;
ll = (point[0] - coef,point[1] - coef / math.cos(point[0] * 0.018))
ur = (point[0] + coef,point[1] + coef / math.cos(point[0] * 0.018))
return ll, ur
def updateBulk(apiKey, downloadCodes, entityId, dataSetName = "SENTINEL_2A"):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + dataSetName + '","downloadCodes":'+ downloadCodes +',"entityId":' + entityId + '}'}
return requests.post(defaultUrl + operations['updateBulk'],data=payload).json()
updateBulk('db000f703052452ab43057bf0b764807','["' +productTypes['full'] + '","' + productTypes['preview'] + '"]','1100852')
updateBulk('db000f703052452ab43057bf0b764807','["' +productTypes['full'] + '","' + productTypes['preview'] + '"]','1464133')
def submitBulk(apiKey):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey+ '"}'}
return requests.post(defaultUrl + operations['submitBulk'],data=payload).json()
submitBulk("db000f703052452ab43057bf0b764807")
def metadata(apiKey, entityIds, dataSetName = "SENTINEL_2A"):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + dataSetName + '","entityIds":' + entityIds + '}'}
return requests.post(defaultUrl + operations['metadata'],data=payload).json()
a = metadata('d64e107c95c2457ea498cb20752f3d83','["1338121"]')
a
print(a['data'][0]['spatialFootprint'])
print(a['data'][0]['sceneBounds'])
for metaDataField in a['data'][0]['metadataFields']:
if metaDataField['fieldName'] == 'Cloud Cover':
print(metaDataField['value'])
break
def sceneSearch(apiKey,point,radius,startDate,endDate,
dataSetName = "SENTINEL_2A", months = None,
includeUnknownCloudCover = True,maxResults = 1000, sortOrder = 'ASC'):
ll, ur = calculateMBR(point,radius)
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + dataSetName + '","spatialFilter":{"filterType":"mbr","lowerLeft":{"latitude":"' + str(ll[0]) + '","longitude":"' + str(ll[1]) + '"},"upperRight":{"latitude":"' + str(ur[0]) + '","longitude":"' + str(ur[1]) + '"}},"temporalFilter":{"startDate":"' + startDate + '","endDate":"' + endDate + '"},'+( '' if months is None else '"months":' + str(months) + ',') + '"includeUnknownCloudCover":' + str(includeUnknownCloudCover).lower() + ',"maxResults":"' + str(maxResults) + '","sortOrder":"' + sortOrder + '"}'}
response = requests.post(defaultUrl + operations['search'], data = payload)
return response.json()
def getScenesDict():
if not os.path.isfile('scenes.json'):
return json.loads('{}')
else:
with open('scenes.json','r') as jsonFile:
return json.load(jsonFile)
def downloadFile(downloadInfo,apiKey,datasetName = "SENTINEL_2A",productType = "STANDARD"):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + datasetName + '","entityIds":"' + downloadInfo[downloadIndexes['entityId']] + '","products":"' + productType + '"}'}
with requests.post(defaultUrl + operations['download'], data=payload) as download:
downloadLink = (download.json())['data'][0]['url']
with requests.get(downloadLink, stream=True, allow_redirects=True) as request:
#lança exceção caso request de errado
request.raise_for_status()
filename = request.headers['content-disposition'].split("=")[1]
print("Baixando: " + filename)
with open(filename, 'wb') as file:
for chunk in request.iter_content(chunk_size=1024):
file.write(chunk)
return (True,downloadLink);
def downloadBulk(lat,lon,radius,startDate,endDate,usr,psw,batchSize,productType = "STANDARD", maxDownload = 1, driverPath = '/PATH_REPO/DesmateSeletivo/chromedriver',downloadPath ='/PATH_REPO/DesmateSeletivo'):
while (apiKey := login(usr,psw)) == None:
print("Erro ao logar tentando novamente")
scenes = sceneSearch(apiKey,(lat,lon),radius,startDate,endDate)
scenesJsonDict = getScenesDict()
downloadList = []
pointStr = f'({str(lat)},{str(lon)})'
for foundScene in scenes['data']['results']:
if foundScene['displayId'] in scenesJsonDict.keys():
print("A imagem " + foundScene['displayId'] + " já foi baixada!")
if pointStr not in scenesJsonDict[foundScene['displayId']]['points']:
print("Registrando o novo ponto: " + pointStr)
scenesJsonDict[foundScene['displayId']]['points'].append(pointStr)
else:
downloadList.append((foundScene['displayId'],foundScene['downloadUrl'],foundScene['entityId'],foundScene['startTime'],foundScene['endTime']))
print("Total de imagens para baixar:", len(downloadList))
for download in downloadList:
print(download[downloadIndexes['displayId']])
print("Adicionando para bulk order:")
updateBulk(apiKey,'["' +productTypes['full'] + '","' + productTypes['preview'] + '"]',download[downloadIndexes['entityId']])
print("Obtendo metadados:")
metaData = metadata(apiKey,'["' + download[downloadIndexes['entityId']] + '"]')
#print("metadata result: ",metaData)
cloudCover = None
for metaDataField in metaData['data'][0]['metadataFields']:
if metaDataField['fieldName'] == 'Cloud Cover':
cloudCover = metaDataField['value']
break
scenesJsonDict[download[downloadIndexes['displayId']]] = {"points" : [pointStr],
"startTime" : download[downloadIndexes['startTime']],
"endTime" : download[downloadIndexes['endTime']],
"cloudCover" : cloudCover,
"sceneBounds" : metaData['data'][0]['sceneBounds'],
"spatialFootprint" : metaData['data'][0]['spatialFootprint'],
"entityId" : download[downloadIndexes['entityId']],
"downloadUrl" : download[downloadIndexes['downloadUrl']],
}
batchSize += 1
with open('scenes.json','w+') as jsonFile:
json.dump(scenesJsonDict,jsonFile)
logout(apiKey)
return batchSize
downloadBulk(-10.463815 ,-58.537751, 500,'2016-01-01','2017-01-01',username,senha, productTypes['preview'])
downloadBulk(-10.463815 ,-58.537751, 500,'2016-01-01','2016-05-01',username,senha, productType=productTypes['preview'])
downloadBulk(-10.463815 ,-58.537751, 500,'2016-01-01','2016-01-07',username,senha, productTypes['preview'])
downloadBulk(-10.463815 ,-58.537751, 500,'2017-07-01','2017-09-30',username,senha, productTypes['preview'])
def getSceneSearchDisplayIds(apiKey,point,radius,startDate,endDate,
dataSetName = "SENTINEL_2A", months = None,
includeUnknownCloudCover = True,maxResults = 1000, sortOrder = 'ASC'):
ll, ur = calculateMBR(point,radius)
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + dataSetName + '","spatialFilter":{"filterType":"mbr","lowerLeft":{"latitude":"' + str(ll[0]) + '","longitude":"' + str(ll[1]) + '"},"upperRight":{"latitude":"' + str(ur[0]) + '","longitude":"' + str(ur[1]) + '"}},"temporalFilter":{"startDate":"' + startDate + '","endDate":"' + endDate + '"},'+( '' if months is None else '"months":' + str(months) + ',') + '"includeUnknownCloudCover":' + str(includeUnknownCloudCover).lower() + ',"maxResults":"' + str(maxResults) + '","sortOrder":"' + sortOrder + '"}'}
response = requests.post(defaultUrl + operations['search'], data = payload)
displayIdList = []
for scene in response.json()['data']['results']:
displayIdList.append(scene['displayId'])
return displayIdList
with open('Coordenadas desmatamento 2017.csv', mode='r') as original, open('CoordeanadasComNome2017.csv', mode='w') as new:
reader = csv.reader(original)
apiKey = login(username,senha)
writer = csv.writer(new)
header = next(reader)
header[1] = "Longitude"
header[2] = "Latitude"
print(header,type(header))
header.append('displayId')
header.append('displayIdList')
writer.writerow(header)
rowN = 0;
for row in reader:
listDisplayId = getSceneSearchDisplayIds(apiKey,(float(row[2]),float(row[1])),60,'2017-07-01','2017-09-30')
row.append(listDisplayId[-1])
row.append(listDisplayId)
writer.writerow(row)
rowN += 1
print(rowN)
if rowN > 1000:
apiKey = login(username,senha)
rowN = 0
with open('Coordenadas desmatamento 2017.csv', mode='r') as original, open('CoordeanadasComNome2017.csv', mode='a', newline='') as new:
reader = csv.reader(original)
apiKey = login(username,senha)
writer = csv.writer(new)
rowN = 0;
for i in range(10406):
next(reader)
for row in reader:
listDisplayId = getSceneSearchDisplayIds(apiKey,(float(row[2]),float(row[1])),60,'2017-07-01','2017-09-30')
row.append(listDisplayId[-1])
row.append(listDisplayId)
writer.writerow(row)
rowN += 1
print(rowN)
if rowN > 1000:
apiKey = login(username,senha)
rowN = 0
apiKey = 'be73bcb7be344d5face235f45e14d0c1'
lat = -9.504953
lon =
getSceneSearchDisplayIds(apiKey,(float(row[2]),float(row[1])),60,'2017-07-01','2017-09-30')
with open('Coordenadas desmatamento 2017.csv', mode='r') as original, open('CoordeanadasComNome2017.csv', mode='a', newline='') as new:
reader = csv.reader(original)
apiKey = login(username,senha)
writer = csv.writer(new)
rowN = 0;
for i in range(10406):
next(reader)
for row in reader:
print(row)
imgList = []
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for row in reader:
imgList.append(row[4])
c = Counter(imgList)
print(len(c))
print(c)
imgList = []
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for row in reader:
imgList.append(row[5])
c = Counter(imgList)
print(len(c))
print(c)
imgNameList = ['L1C_T21LTK_A010732_20170712T142431', 'L1C_T20LRQ_A010732_20170712T142431', 'L1C_T21LTK_A010832_20170719T141344', 'L1C_T21LTK_A011118_20170808T141047', 'L1C_T21LTK_A011304_20170821T142038', 'L1C_T20LRQ_A011304_20170821T142038', 'L1C_T21LTK_A011590_20170910T142035', 'L1C_T20LRQ_A011590_20170910T142035', 'L1C_T21LTK_A011690_20170917T141042', 'L1C_T20LRQ_A011876_20170930T142034', 'L1C_T21LTK_A011876_20170930T142034']
pointsByImage = dict()
for img in imgNameList:
pointsByImage[img] = []
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for row in reader:
rowList = eval(row[5])
for imgName in rowList:
if imgName in imgNameList:
pointsByImage[imgName].append((row[1],row[2]))
#print(len(pointsByImage))
#print(pointsByImage)
#pointsByImage
for img in pointsByImage:
print(len(img))
print(len(pointsByImage['L1C_T21LTK_A010732_20170712T142431']))
print(pointsByImage['L1C_T21LTK_A010732_20170712T142431'])
points = set()
for i in pointsByImage.values():
for p in i:
points.add(p)
len(points)
imgFullList = set()
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for row in reader:
rowList = eval(row[5])
for img in rowList:
imgFullList.add(img)
print(len(imgFullList))
line = 0
mean = 0
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for row in reader:
rowList = eval(row[5])
mean += len(rowList)
line +=1
print(mean/line)
def idLookup(apiKey,idList,inputField = 'displayId',datasetName = "SENTINEL_2A"):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + datasetName + '","idList":"' + idList + '","inputField":"' + inputField + '"}'}
return requests.post(defaultUrl + operations['idlookup'], data = payload).json()['data'][idList]
def getEntityId(displayId,login,usr,datasetName = ):
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + datasetName + '","entityIds":"' + downloadInfo[downloadIndexes['entityId']] + '","products":"' + productType + '"}'}
def downloadByName(nameList,usr,pwd,productType = "STANDARD",datasetName = "SENTINEL_2A"):
for displayId in nameList:
apiKey = login(usr,pwd)
entityId = idLookup(apiKey,displayId)
payload = {"jsonRequest" : '{"apiKey":"' + apiKey + '","datasetName":"' + datasetName + '","entityIds":"' + entityId + '","products":"' + productType + '"}'}
with requests.post(defaultUrl + operations['download'], data=payload) as download:
downloadLink = (download.json())['data'][0]['url']
with requests.get(downloadLink, stream=True, allow_redirects=True) as request:
#lança exceção caso request de errado
request.raise_for_status()
filename = request.headers['content-disposition'].split("=")[1]
print("Baixando: " + filename)
with open(filename, 'wb') as file:
for chunk in request.iter_content(chunk_size=1024):
file.write(chunk)
idLookup('bfe578d3be5140579115d00cbdf8d400','L1C_T21LXC_A001666_20170701T140052')
downloadByName(['L1C_T21LXC_A001666_20170701T140052'],username,senha)
orderSize = 20
actualSize = 0
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
next(reader)
next(reader)
for row in reader:
print("Linnha " + row[0])
if (actualSize := downloadBulk(float(row[2]),float(row[1]), 200,'2017-07-01','2017-09-30',username,senha,actualSize)) > orderSize:
break
apiKey = login(username,senha)
submitBulk(apiKey)
logout(apiKey)
orderSize = 10
actualSize = 0
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
next(reader)
next(reader)
for row in reader:
print("Linnha " + row[0])
if (actualSize := downloadBulk(float(row[2]),float(row[1]), 200,'2017-07-01','2017-09-30',username,senha,actualSize)) > orderSize:
break
apiKey = login(username,senha)
submitBulk(apiKey)
logout(apiKey)
orderSize = 15
actualSize = 0
lines = 200
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for i in range(8936):
next(reader)
for row in reader:
print("Linnha " + row[0])
actualSize = downloadBulk(float(row[2]),float(row[1]), 200,'2017-07-01','2017-09-30',username,senha,actualSize)
lines -= 1
if lines == 0:
break
if actualSize > 0:
apiKey = login(username,senha)
submitBulk(apiKey)
logout(apiKey)
apiKey = login(username,senha)
submitBulk(apiKey)
logout(apiKey)
actualSize
orderSize = 20
actualSize = 0
with open('CoordeanadasComNome2017-final.csv', mode='r') as file:
reader = csv.reader(file)
next(reader)
for i in range(736):
next(reader)
for row in reader:
print("Linnha " + row[0])
with open('/run/media/SEAGATE 1TB/ImagensSentinel2/md5sumNovo.txt') as o, \
open('/run/media/SEAGATE 1TB/ImagensSentinel2/md5sumNovoHd.txt') as h :
i = 0
for line in o:
i +=1
fName = line.split(' ')[1][-39:]
#print('XXXX' + fName)
for lineH in h:
fNameH = lineH.split(' ')[1][-39:]
#print(fName + '==' + fNameH)
if fName == fNameH:
#print(line.split(' ')[0] + '==' + lineH.split(' ')[0])
if line.split(' ')[0] != lineH.split(' ')[0]:
print(fName)
print(line + "!=" + lineH)
h.seek(0)
print(i)
import json
with open('scenes.json','r') as jsonFile:
j =json.load(jsonFile)
print(j["L1C_T21LXC_A001666_20170701T140052"])
import rasterio
#data link: https://earthexplorer.usgs.gov/download/external/options/SENTINEL_2A/3022286/INVSVC/
pathToImgFolder = "D:\RSDD\ImagensSentinel2\data\\"
pathData = pathToImgFolder + "L1C_T21LXC_A001666_20170701T140052\S2B_MSIL1C_20170701T140049_N0205_R067_T21LXC_20170701T140052.SAFE\GRANULE\L1C_T21LXC_A001666_20170701T140052\IMG_DATA\T21LXC_20170701T140049_B01.jp2"
boudingBoxCoordinates = (606859.0750363453, 8241169.269917269, 607219.0750363453, 8241529.269917269)
points = [(607014.0750363453,8241374.26991727),(607024.0750363444,8241374.26991727),\
(607034.0750363453,8241374.26991727),(607044.0750363453,8241374.26991727),\
(607054.0750363442,8241374.26991727)]
with rasterio.open(pathData) as img:
bandWindow = rasterio.windows.from_bounds(*boudingBoxCoordinates, img.transform)
winTransform = rasterio.windows.transform(bandWindow,img.transform)
bandData = img.read(1, window = bandWindow)
allData = img.read(1)
for point in points:
rBand,cBand = rasterio.transform.rowcol(winTransform,point[0],point[1])
rFull,cFull = img.index(point[0],point[1])
bandVal = bandData[rBand,cBand]
fullVal = allData[rFull,cFull]
print(f"{fullVal} {'=' if bandVal == fullVal else '!='} {bandVal}")
|
notebooks/DownloadEarthExplorer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:root] *
# language: python
# name: conda-root-py
# ---
# +
import sys
sys.path.append('..')
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from world_bank import countries, indices
from drake.utils import Entity
# %matplotlib inline
# +
cols = [str(i) for i in range(1991, 2019)]
cols = ["Country Name", "Indicator Name"] + cols
data = pd.read_csv(
"../data/WDI_csv/WDIData.csv",
usecols=cols
)
data = data[data["Country Name"].isin(countries)].reset_index(drop=True)
data = data[data["Indicator Name"].isin(indices)].reset_index(drop=True)
data
# +
grouped = data.groupby("Country Name")
entity_set = {}
for (name, group) in grouped:
entity = group.drop("Country Name", axis=1).set_index("Indicator Name").T
entity = entity[indices]
raw_mean = entity.mean().to_numpy()
raw_variance = entity.var().to_numpy()
scaled_mean = (raw_mean - np.mean(raw_mean)) / np.std(raw_mean)
scaled_variance = (raw_variance - np.mean(raw_variance)) / np.std(raw_variance)
corr = entity.corr().to_numpy()
entity_set[name] = Entity(name, corr, scaled_mean, scaled_variance)
# +
from pickle import load
clustered_countries = load(open("clustered_countries.pkl", "rb"))
clustered_countries
# -
for i, country_list in clustered_countries.items():
temp = []
for country in country_list:
temp.append(entity_set[country].correlation)
sns.heatmap(
np.array(temp).mean(axis=0),
cmap="viridis",
annot=True,
xticklabels=indices.values(),
yticklabels=indices.values()
)
plt.title(f"Cluster {i}")
print(f"Cluster {i}:", ", ".join(country_list))
plt.show()
|
notebooks/visualize_world_bank_results.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
import pandas as pd
import seaborn as sns
from scipy import optimize
from sklearn.metrics import mean_squared_error
class LMM():
def __init__(self):
pass
def fit(self, X, y, groups, method="bfgs"):
if method == "bfgs":
def f(x):
beta_shared, beta_fg = x
preds = np.squeeze(X) * beta_shared + np.matmul(groups, np.squeeze(X)) * beta_fg
# MSE
return np.mean((y - preds)**2)
# Initial value of x
x0 = np.random.normal(size=2)
# Try with BFGS
xopt = optimize.minimize(f,x0,method='bfgs',options={'disp':1})
self.coefs_shared = xopt.x[0]
self.coefs_fg = xopt.x[1]
elif method == "project":
# Regression on all samples
reg = LinearRegression().fit(X, y)
coefs_shared = reg.coef_
# Get residuals for foreground group
X_fg = X[groups == 1]
y_fg = y[groups == 1]
X_fg_preds = reg.predict(X_fg)
X_residuals = y_fg - X_fg_preds
# Regress residuals on the foreground
reg = LinearRegression().fit(X_fg, X_residuals)
coefs_fg = reg.coef_
self.coefs_shared = coefs_shared
self.coefs_fg = coefs_fg
else:
raise Exception("Method must be one of [bfgs, project]")
def predict(self, X, y, groups):
print(self.coefs_shared.shape)
print("FP: ", (np.squeeze(X) * self.coefs_shared).shape)
print("SP: ", np.matmul(groups, np.squeeze(X)).shape)
print("TP: ", self.coefs_fg.shape)
preds = np.squeeze(X) * self.coefs_shared + np.matmul(groups, np.squeeze(X)) * self.coefs_fg
return preds
# +
# simple example
n = 200
p = 12
coefs_shared_true = np.repeat([1], p)
coefs_shared_true = np.reshape(coefs_shared_true, (p, 1))
coefs_fg_true = np.repeat([4], p)
coefs_fg_true = np.reshape(coefs_fg_true, (p, 1))
X = np.random.normal(0, 1, size=(n, p))
groups = np.random.binomial(n=1, p=0.5, size=n)
# Shared effect
y = X @ coefs_shared_true
y = y.reshape((1, n))
y = y + np.random.normal(0, 1, n)
# Foreground-specific effect
y = y.reshape((n, 1))
y[groups == 1] = y[groups == 1] + X[groups == 1, :] @ coefs_fg_true
groups = np.reshape(groups, (1, n))
# -
# Fit LMM
lmm = LMM()
print("X shape: ", X.shape)
print("y shape: ", y.shape)
lmm.fit(X, y, groups=groups)
# +
# Test on a random test set
X_test = np.random.normal(0, 1, size=(n, p))
y_test = X_test @ coefs_shared_true + np.random.normal(0, 1, n)
groups_test = np.random.binomial(n=1, p=0.5, size=n)
y_test[groups_test == 1] = y_test[groups_test == 1] + X_test[groups_test == 1, :] @ coefs_fg_true
groups_test = np.reshape(groups_test, (1, n))
preds = lmm.predict(X_test, y_test, groups_test)
# -
preds.shape
|
simulated_fqi/notebooks/lmm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append("..")
import pandas as pd
import numpy as np
from numba import jit
import json
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import xmltodict
import numpy.polynomial as p
from multiprocessing import Pool
import time
from datetime import datetime , date, timedelta
from fottech_lib.market_data.dmds import DMDSServices
from fottech_lib import instrumentservice
from fottech_lib.market_data.repo import Repo
import project.market_data.repocurves as repoc
from project.market_data.repocurves import RepoCurves
# %matplotlib inline
# -
# ### Loading Indices
#Loading the indices
file_path = '../data/universe_indices.npy'
universe_indices = np.load(file_path)
# ### Computing Universe Repo
def get_repo_schedules(universe_indices_ric,business_date):
dictionary = {}
for ric in universe_indices_ric:
print('############################## Index {} ##############################'.format(ric))
try:
div_paths = 'RepoCurve/official/{}/PARIS/INTRADAY/equity/{}/sophis'.format(business_date,ric)
ds = DMDSServices('prod', 'APAC')
docs = ds.get_documents(div_paths)
d_s = docs['documents']['document'][0].__values__.get('content')
repo_schedule = xmltodict.parse(d_s)
date = repo_schedule['RepoCurve']['@businessDate']
df = pd.DataFrame(repo_schedule['RepoCurve']['repo'])
df['#text'] = df['#text'].astype(str)
df['@term'] = df['@term'].astype(str)
for i in range(df.shape[0]):
f_date = datetime.strptime(date, "%Y-%m-%d").date()
l_date = datetime.strptime(df['@term'][i], "%Y-%m-%d").date()
delta = l_date - f_date
if (delta.days >= 0):
df['@term'][i] = delta.days
else:
df = df.drop(i, axis = 0)
df = df.reset_index(drop=True)
df = df.get_values()
col1 = df[:,0].tolist()
col2 = df[:,1].tolist()
col = [col1 , col2, date]
dictionary[ric]=col
except:
dictionary[ric]=None
return dictionary
def save_dict(dictionary):
file_path = '../output/universe_repo_processed.json'
try:
with open(file_path, 'w') as fp:
json.dump(dictionary, fp)
print('file saved')
except:
print('For some reasons, the file couldnt be saved')
universe_indices_ric = []
B_to_R = instrumentservice.InstrumentService('prod','APAC')
for index in universe_indices:
index_ric = B_to_R.transcode(index, target='reuter', partial_match=False)
if(index_ric != None):
ric = index_ric[1:]
universe_indices_ric.append(ric)
dictionary = get_repo_schedules(universe_indices_ric,'latest')
save_dict(dictionary)
len(dictionary.keys())
# ### Now cleaning and preprocessing the universe repo curves
# +
path_to_data_Universe = '../output/universe_repo_processed.json'
path_to_cleaned_data_Universe = '../output/universe_repo_cleaned.json'
# +
print('################## Cleaning dividends for Universe index ##################')
new_dict = {}
with open(path_to_data_Universe) as json_file:
dictionary = json.load(json_file)
for key in list(dictionary.keys()):
if (dictionary[key]!=None):
if np.sum(np.isnan(dictionary[key][0]))==0 and np.sum(np.isnan(list(map(float,dictionary[key][1]))))==0 :
dictionary[key][1] = list(map(float,dictionary[key][1]))
new_dict[key] = dictionary[key]
xvals = [90, 180, 365, 730, 1095, 1460, 1825, 2190, 2555, 2920, 3285, 3650, 4015, 4380]
for key in new_dict.keys():
x = new_dict[key][0]
y = new_dict[key][1]
yinterp = np.interp(xvals, x, y)
#computing new interpolated values
new_dict[key][0] = xvals
new_dict[key][1] = yinterp.tolist()
with open(path_to_cleaned_data_Universe, 'w') as fp:
json.dump(new_dict, fp)
print('file saved')
# -
|
adam_api_repo_curve_anomaly_detection/notebooks/Preprocessing Universe Repo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import numpy as np
#import pandas as pd
import datetime
import json
from array import *
import os
import math
from random import randrange
import random
#from keras.models import Sequential
#from keras.models import model_from_json
#from keras.layers import Dense, Activation
#from keras import optimizers
from tensorflow.keras.models import Sequential
from tensorflow.keras.models import model_from_json
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras import optimizers
import tensorflow.keras as keras
import tensorflow.compat.v1 as tf
from tensorflow.compat.v1.keras import backend as K
tf.disable_v2_behavior()
# +
#Classes in GAME_SOCKET_DUMMY.py
class ObstacleInfo:
# initial energy for obstacles: Land (key = 0): -1, Forest(key = -1): 0 (random), Trap(key = -2): -10, Swamp (key = -3): -5
types = {0: -1, -1: 0, -2: -10, -3: -5}
def __init__(self):
self.type = 0
self.posx = 0
self.posy = 0
self.value = 0
class GoldInfo:
def __init__(self):
self.posx = 0
self.posy = 0
self.amount = 0
def loads(self, data):
golds = []
for gd in data:
g = GoldInfo()
g.posx = gd["posx"]
g.posy = gd["posy"]
g.amount = gd["amount"]
golds.append(g)
return golds
class PlayerInfo:
STATUS_PLAYING = 0
STATUS_ELIMINATED_WENT_OUT_MAP = 1
STATUS_ELIMINATED_OUT_OF_ENERGY = 2
STATUS_ELIMINATED_INVALID_ACTION = 3
STATUS_STOP_EMPTY_GOLD = 4
STATUS_STOP_END_STEP = 5
def __init__(self, id):
self.playerId = id
self.score = 0
self.energy = 0
self.posx = 0
self.posy = 0
self.lastAction = -1
self.status = PlayerInfo.STATUS_PLAYING
self.freeCount = 0
class GameInfo:
def __init__(self):
self.numberOfPlayers = 1
self.width = 0
self.height = 0
self.steps = 100
self.golds = []
self.obstacles = []
def loads(self, data):
m = GameInfo()
m.width = data["width"]
m.height = data["height"]
m.golds = GoldInfo().loads(data["golds"])
m.obstacles = data["obstacles"]
m.numberOfPlayers = data["numberOfPlayers"]
m.steps = data["steps"]
return m
class UserMatch:
def __init__(self):
self.playerId = 1
self.posx = 0
self.posy = 0
self.energy = 50
self.gameinfo = GameInfo()
def to_json(self):
return json.dumps(self, default=lambda o: o.__dict__, sort_keys=True, indent=4)
class StepState:
def __init__(self):
self.players = []
self.golds = []
self.changedObstacles = []
def to_json(self):
return json.dumps(self, default=lambda o: o.__dict__, sort_keys=True, indent=4)
# -
#Main class in GAME_SOCKET_DUMMY.py
class GameSocket:
bog_energy_chain = {-5: -20, -20: -40, -40: -100, -100: -100}
def __init__(self):
self.stepCount = 0
self.maxStep = 0
self.mapdir = "Maps" # where to load all pre-defined maps
self.mapid = ""
self.userMatch = UserMatch()
self.user = PlayerInfo(1)
self.stepState = StepState()
self.maps = {} # key: map file name, value: file content
self.map = [] # running map info: 0->Land, -1->Forest, -2->Trap, -3:Swamp, >0:Gold
self.energyOnMap = [] # self.energyOnMap[x][y]: <0, amount of energy which player will consume if it move into (x,y)
self.E = 50
self.resetFlag = True
self.craftUsers = [] # players that craft at current step - for calculating amount of gold
self.bots = []
self.craftMap = {} # cells that players craft at current step, key: x_y, value: number of players that craft at (x,y)
def init_bots(self):
self.bots = [Bot1(2), Bot2(3), Bot3(4)] # use bot1(id=2), bot2(id=3), bot3(id=4)
#for (bot) in self.bots: # at the beginning, all bots will have same position, energy as player
for bot in self.bots: # at the beginning, all bots will have same position, energy as player
bot.info.posx = self.user.posx
bot.info.posy = self.user.posy
bot.info.energy = self.user.energy
bot.info.lastAction = -1
bot.info.status = PlayerInfo.STATUS_PLAYING
bot.info.score = 0
self.stepState.players.append(bot.info)
self.userMatch.gameinfo.numberOfPlayers = len(self.stepState.players)
#print("numberOfPlayers: ", self.userMatch.gameinfo.numberOfPlayers)
def reset(self, requests): # load new game by given request: [map id (filename), posx, posy, initial energy]
# load new map
self.reset_map(requests[0])
self.userMatch.posx = int(requests[1])
self.userMatch.posy = int(requests[2])
self.userMatch.energy = int(requests[3])
self.userMatch.gameinfo.steps = int(requests[4])
self.maxStep = self.userMatch.gameinfo.steps
# init data for players
self.user.posx = self.userMatch.posx # in
self.user.posy = self.userMatch.posy
self.user.energy = self.userMatch.energy
self.user.status = PlayerInfo.STATUS_PLAYING
self.user.score = 0
self.stepState.players = [self.user]
self.E = self.userMatch.energy
self.resetFlag = True
self.init_bots()
self.stepCount = 0
def reset_map(self, id): # load map info
self.mapId = id
self.map = json.loads(self.maps[self.mapId])
self.userMatch = self.map_info(self.map)
self.stepState.golds = self.userMatch.gameinfo.golds
self.map = json.loads(self.maps[self.mapId])
self.energyOnMap = json.loads(self.maps[self.mapId])
for x in range(len(self.map)):
for y in range(len(self.map[x])):
if self.map[x][y] > 0: # gold
self.energyOnMap[x][y] = -4
else: # obstacles
self.energyOnMap[x][y] = ObstacleInfo.types[self.map[x][y]]
def connect(self): # simulate player's connect request
print("Connected to server.")
for mapid in range(len(Maps)):
filename = "map" + str(mapid)
print("Found: " + filename)
self.maps[filename] = str(Maps[mapid])
def map_info(self, map): # get map info
# print(map)
userMatch = UserMatch()
userMatch.gameinfo.height = len(map)
userMatch.gameinfo.width = len(map[0])
i = 0
while i < len(map):
j = 0
while j < len(map[i]):
if map[i][j] > 0: # gold
g = GoldInfo()
g.posx = j
g.posy = i
g.amount = map[i][j]
userMatch.gameinfo.golds.append(g)
else: # obstacles
o = ObstacleInfo()
o.posx = j
o.posy = i
o.type = -map[i][j]
o.value = ObstacleInfo.types[map[i][j]]
userMatch.gameinfo.obstacles.append(o)
j += 1
i += 1
return userMatch
def receive(self): # send data to player (simulate player's receive request)
if self.resetFlag: # for the first time -> send game info
self.resetFlag = False
data = self.userMatch.to_json()
for (bot) in self.bots:
bot.new_game(data)
# print(data)
return data
else: # send step state
self.stepCount = self.stepCount + 1
if self.stepCount >= self.maxStep:
for player in self.stepState.players:
player.status = PlayerInfo.STATUS_STOP_END_STEP
data = self.stepState.to_json()
#for (bot) in self.bots: # update bots' state
for bot in self.bots: # update bots' state
bot.new_state(data)
# print(data)
return data
def send(self, message): # receive message from player (simulate send request from player)
if message.isnumeric(): # player send action
self.resetFlag = False
self.stepState.changedObstacles = []
action = int(message)
# print("Action = ", action)
self.user.lastAction = action
self.craftUsers = []
self.step_action(self.user, action)
for bot in self.bots:
if bot.info.status == PlayerInfo.STATUS_PLAYING:
action = bot.next_action()
bot.info.lastAction = action
# print("Bot Action: ", action)
self.step_action(bot.info, action)
self.action_5_craft()
for c in self.stepState.changedObstacles:
self.map[c["posy"]][c["posx"]] = -c["type"]
self.energyOnMap[c["posy"]][c["posx"]] = c["value"]
else: # reset game
requests = message.split(",")
print("Reset game: ", requests[:3], end='')
self.reset(requests)
def step_action(self, user, action):
switcher = {
0: self.action_0_left,
1: self.action_1_right,
2: self.action_2_up,
3: self.action_3_down,
4: self.action_4_free,
5: self.action_5_craft_pre
}
func = switcher.get(action, self.invalidAction)
func(user)
def action_5_craft_pre(self, user): # collect players who craft at current step
user.freeCount = 0
if self.map[user.posy][user.posx] <= 0: # craft at the non-gold cell
user.energy -= 10
if user.energy <= 0:
user.status = PlayerInfo.STATUS_ELIMINATED_OUT_OF_ENERGY
user.lastAction = 6 #eliminated
else:
user.energy -= 5
if user.energy > 0:
self.craftUsers.append(user)
key = str(user.posx) + "_" + str(user.posy)
if key in self.craftMap:
count = self.craftMap[key]
self.craftMap[key] = count + 1
else:
self.craftMap[key] = 1
else:
user.status = PlayerInfo.STATUS_ELIMINATED_OUT_OF_ENERGY
user.lastAction = 6 #eliminated
def action_0_left(self, user): # user go left
user.freeCount = 0
user.posx = user.posx - 1
if user.posx < 0:
user.status = PlayerInfo.STATUS_ELIMINATED_WENT_OUT_MAP
user.lastAction = 6 #eliminated
else:
self.go_to_pos(user)
def action_1_right(self, user): # user go right
user.freeCount = 0
user.posx = user.posx + 1
if user.posx >= self.userMatch.gameinfo.width:
user.status = PlayerInfo.STATUS_ELIMINATED_WENT_OUT_MAP
user.lastAction = 6 #eliminated
else:
self.go_to_pos(user)
def action_2_up(self, user): # user go up
user.freeCount = 0
user.posy = user.posy - 1
if user.posy < 0:
user.status = PlayerInfo.STATUS_ELIMINATED_WENT_OUT_MAP
user.lastAction = 6 #eliminated
else:
self.go_to_pos(user)
def action_3_down(self, user): # user go right
user.freeCount = 0
user.posy = user.posy + 1
if user.posy >= self.userMatch.gameinfo.height:
user.status = PlayerInfo.STATUS_ELIMINATED_WENT_OUT_MAP
user.lastAction = 6 #eliminated
else:
self.go_to_pos(user)
def action_4_free(self, user): # user free
user.freeCount += 1
if user.freeCount == 1:
user.energy += int(self.E / 4)
elif user.freeCount == 2:
user.energy += int(self.E / 3)
elif user.freeCount == 3:
user.energy += int(self.E / 2)
else:
user.energy = self.E
if user.energy > self.E:
user.energy = self.E
def action_5_craft(self):
craftCount = len(self.craftUsers)
# print ("craftCount",craftCount)
if (craftCount > 0):
for user in self.craftUsers:
x = user.posx
y = user.posy
key = str(user.posx) + "_" + str(user.posy)
c = self.craftMap[key]
m = min(math.ceil(self.map[y][x] / c), 50)
user.score += m
# print ("user", user.playerId, m)
for user in self.craftUsers:
x = user.posx
y = user.posy
key = str(user.<KEY> + str(user.posy)
if key in self.craftMap:
c = self.craftMap[key]
del self.craftMap[key]
m = min(math.ceil(self.map[y][x] / c), 50)
self.map[y][x] -= m * c
if self.map[y][x] < 0:
self.map[y][x] = 0
self.energyOnMap[y][x] = ObstacleInfo.types[0]
for g in self.stepState.golds:
if g.posx == x and g.posy == y:
g.amount = self.map[y][x]
if g.amount == 0:
self.stepState.golds.remove(g)
self.add_changed_obstacle(x, y, 0, ObstacleInfo.types[0])
if len(self.stepState.golds) == 0:
for player in self.stepState.players:
player.status = PlayerInfo.STATUS_STOP_EMPTY_GOLD
break;
self.craftMap = {}
def invalidAction(self, user):
user.status = PlayerInfo.STATUS_ELIMINATED_INVALID_ACTION
user.lastAction = 6 #eliminated
def go_to_pos(self, user): # player move to cell(x,y)
if self.map[user.posy][user.posx] == -1:
user.energy -= randrange(16) + 5
elif self.map[user.posy][user.posx] == 0:
user.energy += self.energyOnMap[user.posy][user.posx]
elif self.map[user.posy][user.posx] == -2:
user.energy += self.energyOnMap[user.posy][user.posx]
self.add_changed_obstacle(user.posx, user.posy, 0, ObstacleInfo.types[0])
elif self.map[user.posy][user.posx] == -3:
user.energy += self.energyOnMap[user.posy][user.posx]
self.add_changed_obstacle(user.posx, user.posy, 3,
self.bog_energy_chain[self.energyOnMap[user.posy][user.posx]])
else:
user.energy -= 4
if user.energy <= 0:
user.status = PlayerInfo.STATUS_ELIMINATED_OUT_OF_ENERGY
user.lastAction = 6 #eliminated
def add_changed_obstacle(self, x, y, t, v):
added = False
for o in self.stepState.changedObstacles:
if o["posx"] == x and o["posy"] == y:
added = True
break
if added == False:
o = {}
o["posx"] = x
o["posy"] = y
o["type"] = t
o["value"] = v
self.stepState.changedObstacles.append(o)
def close(self):
print("Close socket.")
#Bots :bot1
class Bot1:
ACTION_GO_LEFT = 0
ACTION_GO_RIGHT = 1
ACTION_GO_UP = 2
ACTION_GO_DOWN = 3
ACTION_FREE = 4
ACTION_CRAFT = 5
def __init__(self, id):
self.state = State()
self.info = PlayerInfo(id)
def next_action(self):
if self.state.mapInfo.gold_amount(self.info.posx, self.info.posy) > 0:
if self.info.energy >= 6:
return self.ACTION_CRAFT
else:
return self.ACTION_FREE
if self.info.energy < 5:
return self.ACTION_FREE
else:
action = self.ACTION_GO_UP
if self.info.posy % 2 == 0:
if self.info.posx < self.state.mapInfo.max_x:
action = self.ACTION_GO_RIGHT
else:
if self.info.posx > 0:
action = self.ACTION_GO_LEFT
else:
action = self.ACTION_GO_DOWN
return action
def new_game(self, data):
try:
self.state.init_state(data)
except Exception as e:
import traceback
traceback.print_exc()
def new_state(self, data):
# action = self.next_action();
# self.socket.send(action)
try:
self.state.update_state(data)
except Exception as e:
import traceback
traceback.print_exc()
#Bots :bot2
class Bot2:
ACTION_GO_LEFT = 0
ACTION_GO_RIGHT = 1
ACTION_GO_UP = 2
ACTION_GO_DOWN = 3
ACTION_FREE = 4
ACTION_CRAFT = 5
def __init__(self, id):
self.state = State()
self.info = PlayerInfo(id)
def next_action(self):
if self.state.mapInfo.gold_amount(self.info.posx, self.info.posy) > 0:
if self.info.energy >= 6:
return self.ACTION_CRAFT
else:
return self.ACTION_FREE
if self.info.energy < 5:
return self.ACTION_FREE
else:
action = np.random.randint(0, 4)
return action
def new_game(self, data):
try:
self.state.init_state(data)
except Exception as e:
import traceback
traceback.print_exc()
def new_state(self, data):
# action = self.next_action();
# self.socket.send(action)
try:
self.state.update_state(data)
except Exception as e:
import traceback
traceback.print_exc()
#Bots :bot3
class Bot3:
ACTION_GO_LEFT = 0
ACTION_GO_RIGHT = 1
ACTION_GO_UP = 2
ACTION_GO_DOWN = 3
ACTION_FREE = 4
ACTION_CRAFT = 5
def __init__(self, id):
self.state = State()
self.info = PlayerInfo(id)
def next_action(self):
if self.state.mapInfo.gold_amount(self.info.posx, self.info.posy) > 0:
if self.info.energy >= 6:
return self.ACTION_CRAFT
else:
return self.ACTION_FREE
if self.info.energy < 5:
return self.ACTION_FREE
else:
action = self.ACTION_GO_LEFT
if self.info.posx % 2 == 0:
if self.info.posy < self.state.mapInfo.max_y:
action = self.ACTION_GO_DOWN
else:
if self.info.posy > 0:
action = self.ACTION_GO_UP
else:
action = self.ACTION_GO_RIGHT
return action
def new_game(self, data):
try:
self.state.init_state(data)
except Exception as e:
import traceback
traceback.print_exc()
def new_state(self, data):
# action = self.next_action();
# self.socket.send(action)
try:
self.state.update_state(data)
except Exception as e:
import traceback
traceback.print_exc()
# +
#MinerState.py
def str_2_json(str):
return json.loads(str, encoding="utf-8")
class MapInfo:
def __init__(self):
self.max_x = 0 #Width of the map
self.max_y = 0 #Height of the map
self.golds = [] #List of the golds in the map
self.obstacles = []
self.numberOfPlayers = 0
self.maxStep = 0 #The maximum number of step is set for this map
def init_map(self, gameInfo):
#Initialize the map at the begining of each episode
self.max_x = gameInfo["width"] - 1
self.max_y = gameInfo["height"] - 1
self.golds = gameInfo["golds"]
self.obstacles = gameInfo["obstacles"]
self.maxStep = gameInfo["steps"]
self.numberOfPlayers = gameInfo["numberOfPlayers"]
def update(self, golds, changedObstacles):
#Update the map after every step
self.golds = golds
for cob in changedObstacles:
newOb = True
for ob in self.obstacles:
if cob["posx"] == ob["posx"] and cob["posy"] == ob["posy"]:
newOb = False
#print("cell(", cob["posx"], ",", cob["posy"], ") change type from: ", ob["type"], " -> ",
# cob["type"], " / value: ", ob["value"], " -> ", cob["value"])
ob["type"] = cob["type"]
ob["value"] = cob["value"]
break
if newOb:
self.obstacles.append(cob)
#print("new obstacle: ", cob["posx"], ",", cob["posy"], ", type = ", cob["type"], ", value = ",
# cob["value"])
def get_min_x(self):
return min([cell["posx"] for cell in self.golds])
def get_max_x(self):
return max([cell["posx"] for cell in self.golds])
def get_min_y(self):
return min([cell["posy"] for cell in self.golds])
def get_max_y(self):
return max([cell["posy"] for cell in self.golds])
def is_row_has_gold(self, y):
return y in [cell["posy"] for cell in self.golds]
def is_column_has_gold(self, x):
return x in [cell["posx"] for cell in self.golds]
def gold_amount(self, x, y): #Get the amount of golds at cell (x,y)
for cell in self.golds:
if x == cell["posx"] and y == cell["posy"]:
return cell["amount"]
return 0
def get_obstacle(self, x, y): # Get the kind of the obstacle at cell(x,y)
for cell in self.obstacles:
if x == cell["posx"] and y == cell["posy"]:
return cell["type"]
return -1 # No obstacle at the cell (x,y)
class State:
STATUS_PLAYING = 0
STATUS_ELIMINATED_WENT_OUT_MAP = 1
STATUS_ELIMINATED_OUT_OF_ENERGY = 2
STATUS_ELIMINATED_INVALID_ACTION = 3
STATUS_STOP_EMPTY_GOLD = 4
STATUS_STOP_END_STEP = 5
def __init__(self):
self.end = False
self.score = 0
self.lastAction = None
self.id = 0
self.x = 0
self.y = 0
self.energy = 0
self.mapInfo = MapInfo()
self.players = []
self.stepCount = 0
self.status = State.STATUS_PLAYING
def init_state(self, data): #parse data from server into object
game_info = str_2_json(data)
self.end = False
self.score = 0
self.lastAction = None
self.id = game_info["playerId"]
self.x = game_info["posx"]
self.y = game_info["posy"]
self.energy = game_info["energy"]
self.mapInfo.init_map(game_info["gameinfo"])
self.stepCount = 0
self.status = State.STATUS_PLAYING
self.players = [{"playerId": 2, "posx": self.x, "posy": self.y},
{"playerId": 3, "posx": self.x, "posy": self.y},
{"playerId": 4, "posx": self.x, "posy": self.y}]
def update_state(self, data):
new_state = str_2_json(data)
for player in new_state["players"]:
if player["playerId"] == self.id:
self.x = player["posx"]
self.y = player["posy"]
self.energy = player["energy"]
self.score = player["score"]
self.lastAction = player["lastAction"]
self.status = player["status"]
self.mapInfo.update(new_state["golds"], new_state["changedObstacles"])
self.players = new_state["players"]
for i in range(len(self.players), 4, 1):
self.players.append({"playerId": i, "posx": self.x, "posy": self.y})
self.stepCount = self.stepCount + 1
# -
#MinerEnv.py
TreeID = 1
TrapID = 2
SwampID = 3
class MinerEnv:
def __init__(self):
self.socket = GameSocket()
self.state = State()
self.score_pre = self.state.score#Storing the last score for designing the reward function
def start(self): #connect to server
self.socket.connect()
def end(self): #disconnect server
self.socket.close()
def send_map_info(self, request):#tell server which map to run
self.socket.send(request)
def reset(self): #start new game
try:
message = self.socket.receive() #receive game info from server
self.state.init_state(message) #init state
except Exception as e:
import traceback
traceback.print_exc()
def step(self, action): #step process
self.socket.send(action) #send action to server
try:
message = self.socket.receive() #receive new state from server
self.state.update_state(message) #update to local state
except Exception as e:
import traceback
traceback.print_exc()
# Functions are customized by client
def get_state(self):
# Building the map
#view = np.zeros([self.state.mapInfo.max_x + 1, self.state.mapInfo.max_y + 1], dtype=int)
view = np.zeros([self.state.mapInfo.max_y + 1, self.state.mapInfo.max_x + 1], dtype=int)
for x in range(self.state.mapInfo.max_x + 1):
for y in range(self.state.mapInfo.max_y + 1):
if self.state.mapInfo.get_obstacle(x, y) == TreeID: # Tree
view[y, x] = -TreeID
if self.state.mapInfo.get_obstacle(x, y) == TrapID: # Trap
view[y, x] = -TrapID
if self.state.mapInfo.get_obstacle(x, y) == SwampID: # Swamp
view[y, x] = -SwampID
if self.state.mapInfo.gold_amount(x, y) > 0:
view[y, x] = self.state.mapInfo.gold_amount(x, y)
DQNState = view.flatten().tolist() #Flattening the map matrix to a vector
# Add position and energy of agent to the DQNState
DQNState.append(self.state.x)
DQNState.append(self.state.y)
DQNState.append(self.state.energy)
#Add position of bots
for player in self.state.players:
if player["playerId"] != self.state.id:
DQNState.append(player["posx"])
DQNState.append(player["posy"])
#Convert the DQNState from list to array for training
DQNState = np.array(DQNState)
return DQNState
def get_reward(self):
# Calculate reward
reward = 0
score_action = self.state.score - self.score_pre
self.score_pre = self.state.score
if score_action > 0:
#If the DQN agent crafts golds, then it should obtain a positive reward (equal score_action)
#reward += score_action
reward += score_action*5
##If the DQN agent crashs into obstacels (Tree, Trap, Swamp), then it should be punished by a negative reward
#if self.state.mapInfo.get_obstacle(self.state.x, self.state.y) == TreeID: # Tree
# reward -= TreeID
#if self.state.mapInfo.get_obstacle(self.state.x, self.state.y) == TrapID: # Trap
# reward -= TrapID
if self.state.mapInfo.get_obstacle(self.state.x, self.state.y) == SwampID: # Swamp
reward -= SwampID
if self.state.lastAction == 4:
reward -= 40
# If out of the map, then the DQN agent should be punished by a larger nagative reward.
if self.state.status == State.STATUS_ELIMINATED_WENT_OUT_MAP:
#if self.state.stepCount < 50:
# reward += -5*(50 - self.state.stepCount)
reward += -50
#Run out of energy, then the DQN agent should be punished by a larger nagative reward.
if self.state.status == State.STATUS_ELIMINATED_OUT_OF_ENERGY:
if self.state.stepCount < 50:
reward += -(50 - self.state.stepCount)
if self.state.lastAction != 4:
# 4 is taking a rest
reward += -10
# control comes to here \implies our agent is not dead yet
if self.state.status == State.STATUS_PLAYING:
if self.state.energy >= 45 and self.state.lastAction == 4:
reward -= 30
# print ("reward",reward)
return reward
def check_terminate(self):
#Checking the status of the game
#it indicates the game ends or is playing
return self.state.status != State.STATUS_PLAYING
#Creating Maps
#This function is used to create 05 maps instead of loading them from Maps folder in the local
def CreateMaps():
map0 = [
[0, 0, -2, 100, 0, 0, -1, -1, -3, 0, 0, 0, -1, -1, 0, 0, -3, 0, -1, -1,0],
[-1,-1, -2, 0, 0, 0, -3, -1, 0, -2, 0, 0, 0, -1, 0, -1, 0, -2, -1, 0,0],
[0, 0, -1, 0, 0, 0, 0, -1, -1, -1, 0, 0, 100, 0, 0, 0, 0, 50, -2, 0,0],
[0, 0, 0, 0, -2, 0, 0, 0, 0, 0, 0, 0, -1, 50, -2, 0, 0, -1, -1, 0,0],
[-2, 0, 200, -2, -2, 300, 0, 0, -2, -2, 0, 0, -3, 0, -1, 0, 0, -3, -1, 0,0],
[0, -1, 0, 0, 0, 0, 0, -3, 0, 0, -1, -1, 0, 0, 0, 0, 0, 0, -2, 0,0],
[0, -1, -1, 0, 0, -1, -1, 0, 0, 700, -1, 0, 0, 0, -2, -1, -1, 0, 0, 0,100],
[0, 0, 0, 500, 0, 0, -1, 0, -2, -2, -1, -1, 0, 0, -2, 0, -3, 0, 0, -1,0],
[-1, -1, 0,-2 , 0, -1, -2, 0, 400, -2, -1, -1, 500, 0, -2, 0, -3, 100, 0, 0,0]
]
map1 = [
[0, 0, -2, 0, 0, 0, -1, -1, -3, 0, 0, 0, -1, -1, 0, 0, -3, 0, -1, -1,0],
[-1,-1, -2, 100, 0, 0, -3, -1, 0, -2, 100, 0, 0, -1, 0, -1, 0, -2, -1, 0,0],
[0, 0, -1, 0, 0, 0, 0, -1, -1, -1, 0, 0, 0, 0, 0, 0, 50, 0, -2, 0,0],
[0, 200, 0, 0, -2, 0, 0, 0, 0, 0, 0, 0, -1, 50, -2, 0, 0, -1, -1, 0,0],
[-2, 0, 0, -2, -2, 0, 0, 0, -2, -2, 0, 0, -3, 0, -1, 0, 0, -3, -1, 0,0],
[0, -1, 0, 0, 300, 0, 0, -3, 0, 0, -1, -1, 0, 0, 0, 0, 0, 0, -2, 0,0],
[500, -1, -1, 0, 0, -1, -1, 0, 700, 0, -1, 0, 0, 0, -2, -1, -1, 0, 0, 0,0],
[0, 0, 0, 0, 0, 0, -1, 0, -2, -2, -1, -1, 0, 0, -2, 0, -3, 100, 0, -1,0],
[-1, -1, 0,-2 , 0, -1, -2, 400, 0, -2, -1, -1, 0, 500, -2, 0, -3, 0, 0, 100,0]
]
map2 = [
[0, 0, -2, 0, 0, 0, -1, -1, -3, 0, 100, 0, -1, -1, 0, 0, -3, 0, -1, -1,0],
[-1,-1, -2, 0, 0, 0, -3, -1, 0, -2, 0, 0, 0, -1, 0, -1, 0, -2, -1, 0,0 ],
[0, 0, -1, 0, 0, 0, 100, -1, -1, -1, 0, 0, 50, 0, 0, 0, 50, 0, -2, 0,0],
[0, 200, 0, 0, -2, 0, 0, 0, 0, 0, 0, 0, -1, 0, -2, 0, 0, -1, -1, 0,0],
[-2, 0, 0, -2, -2, 0, 0, 0, -2, -2, 0, 0, -3, 0, -1, 0, 0, -3, -1, 0,0],
[0, -1, 0, 300, 0, 0, 0, -3, 0, 0, -1, -1, 0, 0, 0, 0, 0, 0, -2, 0,0],
[0, -1, -1, 0, 0, -1, -1, 700, 0, 0, -1, 0, 0, 0, -2, -1, -1, 0, 0, 0,0],
[0, 0, 0, 0, 0, 500, -1, 0, -2, -2, -1, -1, 0, 0, -2, 0, -3, 0, 700, -1,0],
[-1, -1, 0,-2 , 0, -1, -2, 400, 0, -2, -1, -1, 0, 500, -2, 0, -3, 0, 0, 100,0]
]
map3 = [
[0, 0, -2, 0, 0, 0, -1, -1, -3, 0, 0, 0, -1, -1, 0, 0, -3, 0, -1, -1,0],
[-1,-1, -2, 0, 0, 0, -3, -1, 0, -2, 0, 0, 100, -1, 0, -1, 0, -2, -1, 0,0],
[0, 0, -1, 0, 100, 0, 0, -1, -1, -1, 0, 0, 0, 0, 50, 0, 50, 0, -2, 0,0],
[0, 200, 0, 0, -2, 0, 0, 0, 0, 0, 0, 0, -1, 0, -2, 0, 0, -1, -1, 0,0],
[-2, 0, 0, -2, -2, 0, 0, 0, -2, -2, 0, 0, -3, 0, -1, 0, 0, -3, -1, 0,0],
[0, -1, 0, 0, 0, 0, 300, -3, 0, 700, -1, -1, 0, 0, 0, 0, 0, 0, -2, 0,0],
[0, -1, -1, 0, 0, -1, -1, 0, 0, 0, -1, 0, 0, 0, -2, -1, -1, 0, 0, 100,0],
[500, 0, 0, 0, 0, 0, -1, 0, -2, -2, -1, -1, 0, 0, -2, 0, -3, 0, 0, -1,0],
[-1, -1, 0,-2 , 0, -1, -2, 400, 0, -2, -1, -1, 0, 500, -2, 0, -3, 0, 0, 100,0]
]
map4 = [
[0, 0, -2, 0, 100, 0, -1, -1, -3, 0, 0, 0, -1, -1, 0, 0, -3, 0, -1, -1,0],
[-1,-1, -2, 0, 0, 0, -3, -1, 0, -2, 100, 0, 0, -1, 0, -1, 0, -2, -1, 0,0],
[0, 0, -1, 0, 0, 0, 0, -1, -1, -1, 0, 0, 0, 0, 50, 0, 0, 0, -2, 0,0],
[0, 200, 0, 0, -2, 0, 0, 0, 0, 0, 0, 0, -1, 0, -2, 0, 50, -1, -1, 0,0],
[-2, 0, 0, -2, -2, 0, 0, 0, -2, -2, 0, 0, -3, 0, -1, 0, 0, -3, -1, 0,0],
[0, -1, 0, 0, 300, 0, 0, -3, 0, 0, -1, -1, 0, 0, 0, 0, 0, 0, -2, 0,0],
[500, -1, -1, 0, 0, -1, -1, 0, 0, 700, -1, 0, 0, 0, -2, -1, -1, 0, 0, 100,0],
[0, 0, 0, 0, 0, 0, -1, 0, -2, -2, -1, -1, 0, 0, -2, 0, -3, 0, 0, -1,0],
[-1, -1, 0,-2 , 0, -1, -2, 400, 0, -2, -1, -1, 0, 500, -2, 0, -3, 0, 0, 100,0]
]
Maps = (map0,map1,map2,map3,map4)
return Maps
game_over_reason = (
"playing",
"went_out_map",
"out_of_energy",
"invalid_action",
"no_more_gold",
"no_more_step",
)
# ## Start gaming
MAP_MAX_X = 21
MAP_MAX_Y = 9
available_actions = {
"up": '2',
"down": '3',
"left": '0',
"right": '1',
"rest": '4',
"dig": '5',
}
# ### fortran C- reshape?
# + active=""
# help(s.reshape)
# + active=""
# a.reshape(shape, order='C')
# +
Maps = CreateMaps()
minerEnv = MinerEnv()
minerEnv.start()
mapID = np.random.randint(0, 5)
mapID = 1
posID_x = np.random.randint(MAP_MAX_X)
posID_y = np.random.randint(MAP_MAX_Y)
request = ("map" + str(mapID) + "," + str(posID_x) + "," + str(posID_y) + ",50,100")
minerEnv.send_map_info(request)
minerEnv.reset()
s = minerEnv.get_state()
print()
print(s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='C'))
print(s[-9:])
# +
Maps = CreateMaps()
minerEnv = MinerEnv()
minerEnv.start()
mapID = np.random.randint(0, 5)
mapID = 1
posID_x = np.random.randint(MAP_MAX_X)
posID_y = np.random.randint(MAP_MAX_Y)
request = ("map" + str(mapID) + "," + str(posID_x) + "," + str(posID_y) + ",50,100")
minerEnv.send_map_info(request)
minerEnv.reset()
s = minerEnv.get_state()
print()
print(s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='F'))
print(s[-9:])
# +
Maps = CreateMaps()
minerEnv = MinerEnv()
minerEnv.start()
mapID = np.random.randint(0, 5)
mapID = 1
posID_x = np.random.randint(MAP_MAX_X)
posID_y = np.random.randint(MAP_MAX_Y)
request = ("map" + str(mapID) + "," + str(posID_x) + "," + str(posID_y) + ",50,100")
minerEnv.send_map_info(request)
minerEnv.reset()
s = minerEnv.get_state()
print()
print(s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='C'))
print(s[-9:])
# +
Maps = CreateMaps()
minerEnv = MinerEnv()
minerEnv.start()
mapID = np.random.randint(0, 5)
mapID = 1
posID_x = np.random.randint(MAP_MAX_X)
posID_y = np.random.randint(MAP_MAX_Y)
request = ("map" + str(mapID) + "," + str(posID_x) + "," + str(posID_y) + ",50,100")
minerEnv.send_map_info(request)
minerEnv.reset()
s = minerEnv.get_state()
print()
print(s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X)))
print(s[-9:])
# -
vec = np.arange(189).reshape((21,9))
vec
vec.flatten()
vec.flatten().tolist()
vec.flatten(order='F')
help(vec.flatten)
vec.flatten().reshape((21,9))
minerEnv.step(available_actions["down"])
s = minerEnv.get_state()
carte = s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='F')
position, energy = s[-9:-7], s[-7]
terminate = minerEnv.check_terminate()
gold = minerEnv.state.score
print(f"carte = \n{carte}")
print(f"position = {position}")
print(f"energy = {energy}")
print(f"terminate = {bool(terminate)}")
print(f"gold = {gold}")
minerEnv.step(available_actions["dig"])
s = minerEnv.get_state()
carte = s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='F')
position, energy = s[-9:-7], s[-7]
terminate = minerEnv.check_terminate()
gold = minerEnv.state.score
print(f"carte = \n{carte}")
print(f"position = {position}")
print(f"energy = {energy}")
print(f"terminate = {bool(terminate)}")
print(f"gold = {gold}")
minerEnv.step(available_actions["dig"])
s = minerEnv.get_state()
carte = s[:-9].reshape((MAP_MAX_Y, MAP_MAX_X), order='F')
position, energy = s[-9:-7], s[-7]
terminate = minerEnv.check_terminate()
gold = minerEnv.state.score
print(f"carte = \n{carte}")
print(f"position = {position}")
print(f"energy = {energy}")
print(f"terminate = {bool(terminate)}")
print(f"gold = {gold}")
s[-9:]
|
round01/10_no-need-Fortran-reshape.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Create rank 1 & 2 arrays
import numpy as np
# #### - First Way (Creating the array from a list)
# +
list1 = [1, 2, 3, 4]
list2 = [5, 6, 7, 8]
array1 = np.array(list1)
array2 = np.array(list2)
print(array1, array2)
# +
big_list = [list1, list2] # [[1, 2, 3, 4], [5, 6, 7, 8]]
array = np.array(big_list)
print(array, "Array dimensions: {}".format(array.shape), sep="\n\n")
# -
# #### - Second Way (Creating the array directly)
# +
a = np.array([1, 2, 3]) #1 dimensional array or rank 1 array
a[0] = 5
print("Type of the array: {}".format(type(a)), "Array dimensions: {}".format(a.shape), sep="\n")
# -
b = np.array([[1,2,3], [4,5,6]]) # Create a rank 2 array
print(b.shape)
print(b[0,0], b[0,1], b[1,0], sep=", ")
# ### Slicing arrays
# +
arr = np.array([[1,2,3,4], [5,6,7,8]])
arr1 = arr[0][:2]
arr2 = arr[1][1::2]
print(arr1, arr2)
# -
# ##### Keep in mind that if you do change in slice, it affects the parent array
arr1[:] = 5
print(arr)
arr_new = np.array(([0,1,2],[3,4,5]))
print(arr_new[:2, 1:3])
|
Numpy/create-rank-1-2-arrays.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="z00iGjLO8jzo"
# %matplotlib inline
import matplotlib.pyplot as plt
import cv2
import numpy as np
import pandas as pd
from tensorflow.keras.models import Model, Sequential
from tensorflow.keras.layers import Conv2D, Conv2DTranspose, Reshape,LeakyReLU, Dropout
import tensorflow as tf
from tensorflow.keras.layers import AveragePooling2D,UpSampling2D
from tensorflow import keras
# + id="aXFQ33yH87Qm"
ab = np.load('/content/drive/My Drive/colorization data/ab1.npy')
gray = np.load('/content/drive/My Drive/colorization data/gray_scale.npy')
# + id="D5godryMnugl"
def batch_prep (gray_img,batch_size=100):
img=np.zeros((batch_size,224,224,3))
for i in range (0,3):
img[:batch_size,:,:,i]=gray_img[:batch_size]
return img
img_in=batch_prep(gray,batch_size=300)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="1d5ZcF2xqviy" outputId="6b560eae-5cb5-4699-bbdb-c29b18286d97"
plt.imshow(gray[29],cmap=plt.cm.gray)
# + colab={"base_uri": "https://localhost:8080/"} id="fju2Sc9-spzJ" outputId="b4140683-1884-42bf-ed56-3b1ce6f32982"
gray.shape
# + colab={"base_uri": "https://localhost:8080/"} id="sLirkcTgtHek" outputId="05689807-b998-4067-9fdf-cde7f4873d5e"
img_in.shape
# + id="W79efNzutRxV"
def get_rbg(gray_imgs,ab_imgs,n=10):
img1=np.zeros((n,224,224,3))
img1[:,:,:,0]=gray_imgs[0:n:]
img1[:,:,:,1:]=ab_imgs[0:n]
img1=img1.astype('uint8')
imgs=[]
for i in range(0,n):
imgs.append(cv2.cvtColor(img1[i],cv2.COLOR_LAB2RGB))
imgs=np.array(imgs)
return imgs
# + id="S0ZusTGEvqov"
img_out = get_rbg(gray_imgs = gray, ab_imgs = ab, n = 300)
# + id="T8SHx6Dyy89Q"
model = Sequential()
model.add(Conv2D(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(Conv2DTranspose(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(LeakyReLU(0.6))
model.add(Dropout(0.4))
model.add(Conv2D(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(Conv2DTranspose(strides=1,kernel_size=3,filters=3,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(LeakyReLU(0.6))
model.add(Dropout(0.4))
model.add(Conv2D(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(Conv2DTranspose(strides=1,kernel_size=3,filters=3,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(LeakyReLU(0.6))
model.add(Dropout(0.4))
model.add(Conv2D(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(Conv2DTranspose(strides=1,kernel_size=3,filters=3,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(AveragePooling2D(pool_size=(2,2)))
model.add(UpSampling2D((2,2)))
model.add(LeakyReLU(0.6))
model.add(Dropout(0.4))
model.add(Conv2D(strides=1,kernel_size=3,filters=12,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(Conv2DTranspose(strides=1,kernel_size=3,filters=3,use_bias=True,bias_initializer=tf.keras.initializers.RandomUniform(minval=0.05,maxval=0.05),padding="valid",activation=tf.nn.relu))
model.add(LeakyReLU(0.6))
model.add(Dropout(0.4))
# + id="DY6PEn1D55gy"
model.compile(optimizer=tf.keras.optimizers.Adam(clipvalue=0.5),loss='mape',metrics=tf.keras.metrics.Accuracy())
# + id="LXIGGrCk6q7H" colab={"base_uri": "https://localhost:8080/"} outputId="8f9ccce2-b2c2-4c0b-e7ef-2fc6c5f0855d"
model.fit(img_in,img_out,epochs=100,batch_size=16)
# + id="gOqnPPNTABfx"
prediction=model.predict(img_in)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="RufP5IWJAIlS" outputId="15dbe61f-dd63-40c5-c755-8c87ea8a2031"
plt.imshow(prediction[29])
# + id="Y5qZIjDXDXlq"
model.save('model.h5')
# + id="BqZ-WCz8EjIr"
model2=keras.models.load_model('modelfinal.h5')
# + id="Po3QSJKUFhSy"
prediction = model2.predict(img_in)
# + colab={"base_uri": "https://localhost:8080/", "height": 286} id="xtCjXso0Fr85" outputId="a19770a9-94ea-438b-81aa-328ec9017454"
plt.imshow(prediction[29].astype('uint8'))
|
image_colorization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="3HWdNocrJzgY"
# <a href="https://colab.research.google.com/github/magenta/midi-ddsp/blob/main/midi_ddsp/colab/MIDI_DDSP_Demo.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
#
# ##### Copyright 2022 The MIDI-DDSP Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
#
#
#
#
# + cellView="form" id="ZBy5NF8iJ3q7"
#@title
# Copyright 2022 The MIDI-DDSP Authors.
# #
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
# #
# http://www.apache.org/licenses/LICENSE-2.0
# #
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="aZUBBozs6haz"
# # MIDI-DDSP Demo
# Here is the demo where you can automatically synthesize MIDI with the proposed model and then edit the note expression controls of each note. This can be seen as a prototype of our system where users can interact with the model and create the desired music audio together.
#
# [MIDI-DDSP ICLR paper]()
#
# [Audio Examples]()
#
# <img src="https://midi-ddsp.github.io/pics/midi-ddsp-diagram-hori.png" alt="MIDI-DDSP" width="700">
#
# ### Instructions for running:
#
# * Make sure to use a GPU runtime, click: __Runtime >> Change Runtime Type >> GPU__
# * Press ▶️ on the left of each of the cells
# * View the code: Double-click any of the cells
# * Hide the code: Double click the right side of the cell
#
# + id="dO4EmHiR3H70" cellView="form"
#@title #Install Dependencies, Import Code and Setup Models
#@markdown Run this cell to install dependencies, import codes,
#@markdown setup utility functions and load MIDI-DDSP model weights.
#@markdown Running this cell could take a while.
# !pip install -q git+https://github.com/lukewys/qgrid.git
# !pip install -q git+https://github.com/magenta/midi-ddsp
# !midi_ddsp_download_model_weights
# !git clone -q https://github.com/magenta/midi-ddsp.git
# !wget -q https://keymusician01.s3.amazonaws.com/FluidR3_GM.zip
# !unzip -q FluidR3_GM.zip
# Ignore a bunch of deprecation warnings
import sys
sys.path.append('./midi-ddsp')
import warnings
warnings.filterwarnings("ignore")
import os
import librosa
import matplotlib.pyplot as plt
import numpy as np
import tensorflow.compat.v2 as tf
import pandas as pd
import qgrid
import music21
from IPython.display import Javascript
import IPython.display as ipd
from google.colab import files
from google.colab import output
output.enable_custom_widget_manager()
from midi_ddsp import load_pretrained_model
from midi_ddsp.utils.training_utils import set_seed, get_hp
from midi_ddsp.hparams_synthesis_generator import hparams as hp
from midi_ddsp.modules.get_synthesis_generator import get_synthesis_generator, get_fake_data_synthesis_generator
from midi_ddsp.modules.expression_generator import ExpressionGenerator, get_fake_data_expression_generator
from midi_ddsp.utils.audio_io import save_wav
from midi_ddsp.utils.midi_synthesis_utils import synthesize_mono_midi, synthesize_bach
from midi_ddsp.midi_ddsp_synthesize import synthesize_midi
from midi_ddsp.utils.inference_utils import conditioning_df_to_audio, get_process_group
from midi_ddsp.data_handling.instrument_name_utils import INST_NAME_TO_ID_DICT, INST_NAME_LIST
set_seed(1234)
sample_rate = 16000
synthesis_generator, expression_generator = load_pretrained_model()
def plot_spec(wav, sr, title='', play=True, vmin=-8, vmax=1, save_path=None):
D = np.log(np.abs(librosa.stft(wav, n_fft=512 + 256)))
librosa.display.specshow(D, sr=sr, vmin=vmin, vmax=vmax, cmap='magma')
plt.title(title)
wav = np.clip(wav, -1, 1)
if play:
ipd.display(ipd.Audio(wav, rate=sr))
if save_path:
plt.savefig(save_path)
plt.close()
EDIT_DF_NAME_ORDER = ['volume', 'vol_fluc', 'vol_peak_pos', 'vibrato', 'brightness', 'attack', 'pitch', 'note_length']
COND_DF_NAME_ORDER = ['volume', 'vol_fluc', 'vibrato', 'brightness', 'attack', 'vol_peak_pos', 'pitch', 'onset', 'offset', 'note_length']
def conditioning_df_to_edit_df(conditioning_df):
edit_df = conditioning_df.copy()
return edit_df[EDIT_DF_NAME_ORDER]
def edit_df_to_conditioning_df(edit_df):
conditioning_df = edit_df.copy()
note_length = conditioning_df['note_length'].values
offset = np.cumsum(note_length)
onset = np.concatenate([[0],offset[:-1]])
conditioning_df['onset']=onset
conditioning_df['offset']=offset
return conditioning_df[COND_DF_NAME_ORDER]
GAIN_ADJUST_DB_DICT = {
'string_set': {
'Soprano': 2,
'Alto': 2,
'Tenor': -1,
'Bass': -1,
},
'woodwind_set': {
'Soprano': 1.5,
'Alto': 1.2,
'Tenor': 0,
'Bass': 1.8,
},
'brasswind_set': {
'Soprano': 2,
'Alto': 2,
'Tenor': 5.6,
'Bass': 2.9,
},
}
def upload_midi():
midi_files = files.upload()
fnames = list(midi_files.keys())
return fnames
print('Done!')
# + [markdown] id="CVRDl2QBf71C"
# ## Monophonic MIDI Synthesis
# + cellView="form" id="qkINRV9Tk8BT"
#@markdown Let's first synthesize a MIDI using MIDI-DDSP! By running this cell without any change, MIDI-DDSP will synthesis the MIDI of "ode to joy" using violin. This will take about a minute.
#@markdown You can also upload your own MIDI file for MIDI-DDSP to synthesize! (by changing the `midi_file` to "Upload (.mid)")
#@markdown There are 13 instruments available. You can select instrument in the dropdown menu next to "instrument".
#@markdown Besides changing instruments and MIDI file, there are two variables you could change to adjust the MIDI synthesis:
#@markdown - `pitch_offset`: transpose the MIDI file for `pitch_offset` semitones. (>0 is pitch up, <0 is pitch down).
#@markdown Different instrument has different pitch range. Please consider adjusting the `pitch_offset` for different instrument depending on the MIDI file.
#@markdown - `speed_rate`: adjust play speed of the MIDI (=1: original speed, >1: faster, <1: slower).
#@markdown In this cell, we will only synthesizing a single monophonic track. If you upload multi-track MIDI, only the first track will be used. You can synthesize multi-track MIDI using MIDI-DDSP in the cell below.
#@markdown The generation speed would be 2.5x-5x realtime. That is, one need to wait 24-50 seconds for render a 10 second MIDI.
midi_file = 'Ode to Joy' #@param ['Ode to Joy','Upload (.mid)']
instrument = "violin" #@param ['violin', 'viola', 'cello', 'double bass', 'flute', 'oboe', 'clarinet', 'saxophone', 'bassoon', 'trumpet', 'horn', 'trombone', 'tuba']
pitch_offset = 0#@param {type:"integer"}
speed_rate = 1#@param {type:"number", min:0}
if midi_file == 'Ode to Joy':
midi_file = r'./midi-ddsp/midi_example/ode_to_joy.mid'
else:
midi_file = upload_midi()[0]
instrument_name = instrument
instrument_id = INST_NAME_TO_ID_DICT[instrument_name]
midi_audio, midi_control_params, midi_synth_params, conditioning_df = synthesize_mono_midi(synthesis_generator, expression_generator, midi_file, instrument_id, output_dir=None, pitch_offset=pitch_offset, speed_rate=speed_rate)
plt.figure(figsize=(15,5))
plot_spec(midi_audio[0].numpy(), sr=16000)
plt.show()
# + [markdown] id="pVZ0hinyiUp7"
# ## Adjusting Note Expression
# + [markdown] id="XfPPrdPu5sSy"
# In MIDI-DDSP, six note expression controls are designed and used to control the expressive performance, all in range of [0,1]. You can adjust them to edit the performance aspect of the MIDI synthesis:
# - Volume (`volume`): Controls overall volume of a note. (larger value -> larger volume)
# - Volume Fluctuation (`vol_fluc`): Controls the extent of the volume changing in a note (crescendo & decrescendo or not). (larger value -> more extensive dynamic changing)
# - Volume Peak Position (`vol_peak_pos`): Controls the volume changing in a note (crescendo & decrescendo, together with amplitude_std). (larger value -> later reach maximum volume)
# - Attack Noise (`attack_noise`): Controls the extent of note attack (strong or soft). (larger value -> larger attack)
# - Brightness (`brightness`): Controls the timbre of a note. (larger value -> brighter / more amplitude on higher harmonics)
# - Vibrato (`vibrato`): Controls the extend of the vibrato of a note. (larger value -> larger vibrato extend)
#
#
# + id="5q2DmNEO3H8J" cellView="form"
#@markdown Run this cell to get an editable table for adjusting note expression.
#@markdown The values shown are predicted by note expression control generator.
#@markdown Each row is a note. Double click the item in the table and enter the value, click anywhere else to save the change.
#@markdown You can also edit the note pitch and length just as editing a MIDI sequence.
#@markdown To add a note or remove a note, click "Add Row" and "Remove Row" on the upper left corner.
#@markdown After edit, the table is changed automatically and you can run the synthesize cell below the next cell to synthesize the result.
#@markdown **Run this cell again will reset the table to the initial value.**
qgrid_widget = qgrid.show_grid(conditioning_df_to_edit_df(conditioning_df), show_toolbar=True)
qgrid_widget
# + cellView="form" id="kBAjQYSW3H8K"
#@markdown Run this cell to synthesize with edited note expression controls.
conditioning_df_changed = edit_df_to_conditioning_df(qgrid_widget.get_changed_df())
midi_audio, midi_control_params, midi_synth_params = conditioning_df_to_audio(synthesis_generator, conditioning_df_changed, tf.constant([instrument_id]), display_progressbar=True)
plt.figure(figsize=(15,5))
plot_spec(midi_audio[0].numpy(), sr=16000)
plt.show()
# + [markdown] id="YWs-peNlsilU"
# ## Bach Chorales Synthesis
# + cellView="form" id="6PD7OeeKEYkX"
#@markdown Besides synthesizing monophonic MIDI, MIDI-DDSP can also synthesizing multi-track MIDI.
#@markdown Running this cell to synthesis quartet of [4-part Bach Chorales](https://en.wikipedia.org/wiki/List_of_chorale_harmonisations_by_Johann_Sebastian_Bach).
#@markdown You can synthesize any Bach Chorales by typing in piece number to `piece_number` below. A full list of bach chorales available can be found [here](https://github.com/cuthbertLab/music21/tree/master/music21/corpus/bach).
#@markdown We provide three quartet settings, string, woodwind and brass wind. You can change the ensemble by selecting from the drop down menu of `ensemble`.
piece_number = 'bwv227.1' #@param {type:"string"}
ensemble = 'string_set' #@param ['string_set', 'woodwind_set', 'brasswind_set']
score = music21.corpus.parse(f'bach/{piece_number}')
score.write('midi', fp=f'./{piece_number}.mid')
midi_file = f'./{piece_number}.mid'
midi_audio_mix, midi_audio_all, midi_control_params, midi_synth_params, conditioning_df_all = synthesize_bach(
synthesis_generator,
expression_generator,
midi_file,
quartet_set=ensemble,
pitch_offset=0,
speed_rate=1,
output_dir='./',
gain_adjust_db_dict=GAIN_ADJUST_DB_DICT[ensemble])
part_name = list(GAIN_ADJUST_DB_DICT[ensemble].keys())
plt.figure(figsize=(15,5))
plot_spec(midi_audio_mix, sr=16000, title='Mix')
plt.show()
for i in range(len(midi_audio_all)):
plt.figure(figsize=(15,5))
plot_spec(midi_audio_all[i], sr=16000, title=part_name[i])
plt.show()
# + [markdown] id="g0fzbo7Nz-sF"
# ## Pitch Bend by Editing Synthesis Parameters
# + cellView="form" id="bEpDheCtEnl4"
#@markdown Run this cell to generate a pitch bend by editing synthesis parameters.
#@markdown You can double-click the cell to see how we do it in the code.
#@markdown This is just an example.
#@markdown We encourage you to come up with smarter ways to smooth the connection and
#@markdown crazy ways to play with the synthesis parameters :).
# First define functions to generate pitch bend.
# You can come up with your own way of edit pitch or
# other synthesis parameters.
def get_pitch_bend(start_value, end_value, length, power=4,offset_1=-1,bend_type='exp'):
if bend_type=='exp':
if start_value <= end_value:
value = start_value + (np.power(np.linspace(0.0, 1.0, num=length)+offset_1, power)-offset_1) * (end_value-start_value)
return value
elif bend_type=='linear':
return np.linspace(start_value, end_value, length)
# First run the MIDI-DDSP to get the synthesis parameters predicted
# You don't need to run this if you already have the synthesis parameter prediction.
midi_file = r'./midi-ddsp/midi_example/ode_to_joy.mid'
instrument_name = 'violin'
instrument_id = INST_NAME_TO_ID_DICT[instrument_name]
midi_audio, midi_control_params, midi_synth_params, conditioning_df = synthesize_mono_midi(synthesis_generator, expression_generator, midi_file, instrument_id, output_dir=None, pitch_offset=pitch_offset, speed_rate=speed_rate)
# Assume we want to add a pitch bend in the middle of two notes.
# The first note ends at frame 368 while the next note start at the frame 375
prev_note_off = 368
next_note_on = 375
f0_ori = midi_synth_params['f0_hz'][0,...,0]
amps_ori = midi_synth_params['amplitudes'].numpy()[0,...,0]
noise_ori = midi_synth_params['noise_magnitudes'].numpy()
hd_ori = midi_synth_params['harmonic_distribution'].numpy()
# Edit the f0 to add the pitch bend, starting from
# 20 frames before the previous note off to 50 frames after next note on.
edit_frame_start = prev_note_off-20
edit_frame_end = next_note_on+50
edit_frame_duration = edit_frame_end - edit_frame_start
f0_changed = tf.concat([f0_ori[:edit_frame_start],
get_pitch_bend(f0_ori[edit_frame_start],
f0_ori[edit_frame_end],
(edit_frame_end)-(edit_frame_start),
power=7,
bend_type='exp'),
f0_ori[edit_frame_end:]], axis=0)
f0_changed = f0_changed[tf.newaxis, ..., tf.newaxis]
# For other synthesis parameters, use that from the start of the next note
# to replace the connection of notes.
# We also need to avoid the onset of the next note,
# thus that is the reason we use "next_note_on+5" as the start.
amps_changed = amps_ori
amps_changed[edit_frame_start:edit_frame_end] = amps_ori[next_note_on+5:next_note_on+edit_frame_duration+5]
amps_changed = amps_changed[tf.newaxis, ..., tf.newaxis]
noise_changed = noise_ori
noise_changed[0,edit_frame_start:edit_frame_end,:] = noise_ori[0,next_note_on+5:next_note_on+edit_frame_duration+5,:]
hd_changed = hd_ori
hd_changed[0,edit_frame_start:edit_frame_end,:] = hd_changed[0,next_note_on+5:next_note_on+edit_frame_duration+5,:]
# Resynthesis the audio using DDSP
processor_group = get_process_group(midi_synth_params['amplitudes'].shape[1], use_angular_cumsum=True)
midi_audio_changed = processor_group({'amplitudes': amps_changed,
'harmonic_distribution': hd_changed,
'noise_magnitudes': noise_changed,
'f0_hz': f0_changed,},
verbose=False)
if synthesis_generator.reverb_module is not None:
midi_audio_changed = synthesis_generator.reverb_module(midi_audio_changed, reverb_number=instrument_id, training=False)
plt.figure(figsize=(15,5))
# Just play the first 4 seconds
plot_spec(midi_audio[0].numpy()[:4*16000], sr=16000, title='Original')
plt.show()
plt.figure(figsize=(15,5))
plot_spec(midi_audio_changed[0].numpy()[:4*16000], sr=16000, title='Add pitch bend')
plt.show()
# + [markdown] id="5xYX-txJ1B13"
# ## Multi-track MIDI Synthesis
# + cellView="form" id="IpddFjD97QoV"
#@markdown Run this cell to upload and synthesize any multi-track MIDI.
#@markdown For midi programs that are not supported by MIDI-DDSP, we will use [FluidSynth](https://www.fluidsynth.org/) to synthesize the track.
#@markdown For midi programs supported by MIDI-DDSP, it will only synthesize a monophonic performance.
#@markdown That is, for a polyphonic track, only a polyphonic note sequence will be synthesized.
pitch_offset = 0#@param {type:"integer"}
speed_rate = 1#@param {type:"number", min:0}
midi_file = upload_midi()[0]
output = synthesize_midi(synthesis_generator, expression_generator, midi_file,
pitch_offset=pitch_offset, speed_rate=speed_rate,
output_dir=r'./',
use_fluidsynth=True,
sf2_path='./FluidR3_GM.sf2',
display_progressbar=True)
plot_spec(output['midi_audio_mix'], sr=16000, title='Mix')
for i in range(len(output['stem_audio'])):
plt.figure(figsize=(15,5))
plot_spec(output['stem_audio'][i], sr=16000, title=f'Track {i}')
plt.show()
|
midi_ddsp/colab/MIDI_DDSP_Demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/liganega/gmlcc/blob/master/notebooks_ko/Ch04C-synthetic_features_and_outliers.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="copyright-notice"
# #### Copyright 2017 Google LLC.
# + colab_type="code" id="copyright-notice2" cellView="both" colab={}
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="4f3CKqFUqL2-" colab_type="text"
# # 합성 특성과 이상점
# + [markdown] id="jnKgkN5fHbGy" colab_type="text"
# **학습 목표:**
# * 다른 두 특성의 비율로 합성 특성을 만든다
# * 새 특성을 선형 회귀 모델의 입력으로 사용한다
# * 입력 데이터에서 이상점을 식별 및 삭제하여 모델의 효율성을 개선한다
# + [markdown] id="VOpLo5dcHbG0" colab_type="text"
# 텐서플로우 첫걸음 실습에서 사용한 모델을 다시 살펴보겠습니다.
#
# 우선 캘리포니아 주택 데이터를 *pandas* `DataFrame`으로 가져옵니다.
# + [markdown] id="S8gm6BpqRRuh" colab_type="text"
# ## 설정
# + id="9D8GgUovHbG0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 456} outputId="830488c9-b53c-4233-9508-e3a870646916"
from __future__ import print_function
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn.metrics as metrics
# %tensorflow_version 1.x
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://download.mlcc.google.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
california_housing_dataframe["median_house_value"] /= 1000.0
california_housing_dataframe
# + [markdown] id="I6kNgrwCO_ms" colab_type="text"
# 다음으로, 입력 함수를 설정하고 모델 학습용 함수를 정의합니다.
# + id="5RpTJER9XDub" colab_type="code" colab={}
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model of one feature.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating.
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified.
if shuffle:
ds = ds.shuffle(buffer_size=10000)
# Return the next batch of data.
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + id="VgQPftrpHbG3" colab_type="code" colab={}
def train_model(learning_rate, steps, batch_size, input_feature):
"""Trains a linear regression model.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
input_feature: A `string` specifying a column from `california_housing_dataframe`
to use as input feature.
Returns:
A Pandas `DataFrame` containing targets and the corresponding predictions done
after training the model.
"""
periods = 10
steps_per_period = steps / periods
my_feature = input_feature
my_feature_data = california_housing_dataframe[[my_feature]].astype('float32')
my_label = "median_house_value"
targets = california_housing_dataframe[my_label].astype('float32')
# Create input functions.
training_input_fn = lambda: my_input_fn(my_feature_data, targets, batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(my_feature_data, targets, num_epochs=1, shuffle=False)
# Create feature columns.
feature_columns = [tf.feature_column.numeric_column(my_feature)]
# Create a linear regressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
linear_regressor = tf.estimator.LinearRegressor(
feature_columns=feature_columns,
optimizer=my_optimizer
)
# Set up to plot the state of our model's line each period.
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
plt.title("Learned Line by Period")
plt.ylabel(my_label)
plt.xlabel(my_feature)
sample = california_housing_dataframe.sample(n=300)
plt.scatter(sample[my_feature], sample[my_label])
colors = [cm.coolwarm(x) for x in np.linspace(-1, 1, periods)]
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print("Training model...")
print("RMSE (on training data):")
root_mean_squared_errors = []
for period in range (0, periods):
# Train the model, starting from the prior state.
linear_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period,
)
# Take a break and compute predictions.
predictions = linear_regressor.predict(input_fn=predict_training_input_fn)
predictions = np.array([item['predictions'][0] for item in predictions])
# Compute loss.
root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(predictions, targets))
# Occasionally print the current loss.
print(" period %02d : %0.2f" % (period, root_mean_squared_error))
# Add the loss metrics from this period to our list.
root_mean_squared_errors.append(root_mean_squared_error)
# Finally, track the weights and biases over time.
# Apply some math to ensure that the data and line are plotted neatly.
y_extents = np.array([0, sample[my_label].max()])
weight = linear_regressor.get_variable_value('linear/linear_model/%s/weights' % input_feature)[0]
bias = linear_regressor.get_variable_value('linear/linear_model/bias_weights')
x_extents = (y_extents - bias) / weight
x_extents = np.maximum(np.minimum(x_extents,
sample[my_feature].max()),
sample[my_feature].min())
y_extents = weight * x_extents + bias
plt.plot(x_extents, y_extents, color=colors[period])
print("Model training finished.")
# Output a graph of loss metrics over periods.
plt.subplot(1, 2, 2)
plt.ylabel('RMSE')
plt.xlabel('Periods')
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(root_mean_squared_errors)
# Create a table with calibration data.
calibration_data = pd.DataFrame()
calibration_data["predictions"] = pd.Series(predictions)
calibration_data["targets"] = pd.Series(targets)
display.display(calibration_data.describe())
print("Final RMSE (on training data): %0.2f" % root_mean_squared_error)
return calibration_data
# + [markdown] id="FJ6xUNVRm-do" colab_type="text"
# ## 작업 1: 합성 특성 사용해 보기
#
# `total_rooms` 특성과 `population` 특성은 모두 특정 지역의 합계를 계수합니다.
#
# 그런데 지역마다 인구밀도가 다르다면 어떻게 될까요? `total_rooms`와 `population`의 비율로 합성 특성을 만들면 지역의 인구밀도와 주택 가격 중앙값의 관계를 살펴볼 수 있습니다.
#
# 아래 셀에서 `rooms_per_person`이라는 특성을 만들고 `train_model()`의 `input_feature`로 사용합니다.
#
# 학습률을 조정하여 이 단일 특성으로 성능을 어디까지 올릴 수 있을까요? 성능이 높다는 것은 회귀선이 데이터에 잘 부합하고 최종 RMSE가 낮다는 의미입니다.
# + [markdown] id="isONN2XK32Wo" colab_type="text"
# **참고**: 아래에 코드 셀을 몇 개 추가하여 다양한 학습률을 실험하면서 결과를 비교해 보면 도움이 됩니다. 새 코드 셀을 추가하려면 이 셀 가운데 바로 아래에 마우스를 가져가고 **CODE**를 클릭합니다.
# + id="5ihcVutnnu1D" colab_type="code" cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 130} outputId="29aacc2d-4bcd-4d13-9858-5b75d7215ace"
#
# YOUR CODE HERE
#
california_housing_dataframe["rooms_per_person"] =
calibration_data = train_model(
learning_rate=0.00005,
steps=500,
batch_size=5,
input_feature="rooms_per_person"
)
# + [markdown] id="i5Ul3zf5QYvW" colab_type="text"
# ### 해결 방법
#
# 해결 방법을 보려면 아래를 클릭하세요.
# + id="Leaz2oYMQcBf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 894} outputId="f59aeb92-4aed-4b84-c66f-08593cb5df13"
california_housing_dataframe["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] / california_housing_dataframe["population"])
calibration_data = train_model(
learning_rate=0.05,
steps=500,
batch_size=5,
input_feature="rooms_per_person")
# + [markdown] id="ZjQrZ8mcHFiU" colab_type="text"
# ## 작업 2: 이상점 식별
#
# 예측과 목표값을 비교한 산포도를 작성하면 모델의 성능을 시각화할 수 있습니다. 이상적인 상태는 완벽한 상관성을 갖는 대각선이 그려지는 것입니다.
#
# 작업 1에서 학습한 rooms-per-person 모델을 사용한 예측과 타겟에 대해 Pyplot의 `scatter()`로 산포도를 작성합니다.
#
# 특이한 점이 눈에 띄나요? `rooms_per_person`의 값 분포를 조사하여 소스 데이터를 추적해 보세요.
# + id="P0BDOec4HbG_" colab_type="code" colab={}
# YOUR CODE HERE
# + [markdown] id="jByCP8hDRZmM" colab_type="text"
# ### 해결 방법
#
# 해결 방법을 보려면 아래를 클릭하세요.
# + id="s0tiX2gdRe-S" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 391} outputId="753b57ad-8194-41c4-b107-b5898f9051cb"
plt.figure(figsize=(15, 6))
plt.subplot(1, 2, 1)
plt.scatter(calibration_data["predictions"], calibration_data["targets"])
# + [markdown] id="kMQD0Uq3RqTX" colab_type="text"
# 보정 데이터를 보면 대부분의 산포점이 직선을 이룹니다. 이 선은 수직에 가까운데, 여기에 대해서는 나중에 설명합니다. 지금은 선에서 벗어난 점에 대해 집중할 때입니다. 이러한 점은 비교적 적은 편입니다.
#
# `rooms_per_person`의 히스토그램을 그려보면 입력 데이터에서 몇 개의 이상점을 발견할 수 있습니다.
# + id="POTM8C_ER1Oc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="ea3752a0-8882-4ac0-8952-10cbeba5d418"
plt.subplot(1, 2, 2)
_ = california_housing_dataframe["rooms_per_person"].hist()
# + [markdown] id="9l0KYpBQu8ed" colab_type="text"
# ## 작업 3: 이상점 삭제
#
# `rooms_per_person`의 이상점 값을 적당한 최소값 또는 최대값으로 설정하여 모델의 적합성을 더 높일 수 있는지 살펴보세요.
#
# 다음은 Pandas `Series`에 함수를 적용하는 방법을 간단히 보여주는 예제입니다.
#
# clipped_feature = my_dataframe["my_feature_name"].apply(lambda x: max(x, 0))
#
# 위와 같은 `clipped_feature`는 `0` 미만의 값을 포함하지 않습니다.
# + id="rGxjRoYlHbHC" colab_type="code" colab={}
# YOUR CODE HERE
# + [markdown] id="WvgxW0bUSC-c" colab_type="text"
# ### 해결 방법
#
# 해결 방법을 보려면 아래를 클릭하세요.
# + [markdown] id="8YGNjXPaSMPV" colab_type="text"
# 작업 2에서 작성한 히스토그램을 보면 대부분의 값이 `5` 미만입니다. `rooms_per_person`을 5에서 잘라내고 히스토그램을 작성하여 결과를 다시 확인해 보세요.
# + id="9YyARz6gSR7Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="b4dc4a1d-4491-4249-9338-6aa6f40eef12"
california_housing_dataframe["rooms_per_person"] = (
california_housing_dataframe["rooms_per_person"]).apply(lambda x: min(x, 5))
_ = california_housing_dataframe["rooms_per_person"].hist()
# + [markdown] id="vO0e1p_aSgKA" colab_type="text"
# 삭제가 효과가 있었는지 확인하기 위해 학습을 다시 실행하고 보정 데이터를 한 번 더 출력해 보겠습니다.
# + id="ZgSP2HKfSoOH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 894} outputId="f3e0a9ca-a497-403e-9250-b4e7dbf09e10"
calibration_data = train_model(
learning_rate=0.05,
steps=500,
batch_size=5,
input_feature="rooms_per_person")
# + id="gySE-UgfSony" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="c022fa47-477d-4c78-e12f-b32208fae7f1"
_ = plt.scatter(calibration_data["predictions"], calibration_data["targets"])
|
notebooks_ko/Ch04C-synthetic_features_and_outliers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="vAM__pSGf5Pt"
# ## Exercises
# + [markdown] colab_type="text" id="C_nNF6aAf5P8"
# #### Exercise 01: Simple Transaction Data
# + colab={} colab_type="code" id="o9cGDEXCf5QB"
import matplotlib.pyplot as plt
import mlxtend.frequent_patterns
import mlxtend.preprocessing
import numpy
import pandas
# + colab={} colab_type="code" id="cGZWj6G9f5QM"
example = [
['milk', 'bread', 'apples', 'cereal', 'jelly', 'cookies', 'salad', 'tomatoes'],
['beer', 'milk', 'chips', 'salsa', 'grapes', 'wine', 'potatoes', 'eggs', 'carrots'],
['diapers', 'baby formula', 'milk', 'bread', 'chicken', 'asparagus', 'cookies'],
['milk', 'cookies', 'chicken', 'asparagus', 'broccoli', 'cereal', 'orange juice'],
['steak', 'asparagus', 'broccoli', 'chips', 'salsa', 'ketchup', 'potatoes', 'salad'],
['beer', 'salsa', 'asparagus', 'wine', 'cheese', 'crackers', 'strawberries', 'cookies'],
['chocolate cake', 'strawberries', 'wine', 'cheese', 'beer', 'milk', 'orange juice'],
['chicken', 'peas', 'broccoli', 'milk', 'bread', 'eggs', 'potatoes', 'ketchup', 'crackers'],
['eggs', 'bread', 'cheese', 'turkey', 'salad', 'tomatoes', 'wine', 'steak', 'carrots'],
['bread', 'milk', 'tomatoes', 'cereal', 'chicken', 'turkey', 'chips', 'salsa', 'diapers']
]
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="lNQ8WGZhf5Qe" outputId="1d267ab7-619d-4460-a3ca-c8467c878d2e"
print(example)
# + [markdown] colab_type="text" id="jkXS9DdHf5Qw"
# #### Exercise 02: Computing Metrics
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="b5kmwORMf5Q5" outputId="8f28655c-d796-4c5b-91d1-110110e7bb86"
# the number of transactions
N = len(example)
# the frequency of milk
f_x = sum(['milk' in i for i in example])
# the frequency of bread
f_y = sum(['bread' in i for i in example])
# the frequency of milk and bread
f_x_y = sum([
all(w in i for w in ['milk', 'bread'])
for i in example
])
# print out the metrics computed above
print(
"N = {}\n".format(N) +
"Freq(x) = {}\n".format(f_x) +
"Freq(y) = {}\n".format(f_y) +
"Freq(x, y) = {}".format(f_x_y)
)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="DqCmaTqhf5RF" outputId="469031d1-ed7e-45c6-d7a6-0209c44b3ce9"
# support (supp)
support = f_x_y / N
print("Support = {}".format(round(support, 4)))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1hUYlCc0f5RZ" outputId="60fc36aa-df6e-4f2f-8744-f2e9ed4039dd"
# confidence: x -> y
confidence = support / (f_x / N)
print("Confidence = {}".format(round(confidence, 4)))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="PPNr72sGf5R6" outputId="b2d6c5dd-afe7-4af1-bb17-de423e497267"
# lift: x -> y
lift = confidence / (f_y / N)
print("Lift = {}".format(round(lift, 4)))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="YyEi8lKBf5SB" outputId="25f54324-96f2-4ec1-db72-acf261f2cfc2"
# leverage: x -> y
leverage = support - ((f_x / N) * (f_y / N))
print("Leverage = {}".format(round(leverage, 4)))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="qE5ft71gf5SJ" outputId="dfdcc58c-7c7e-40fc-9886-aa6b6efebd55"
# conviction: x -> y
conviction = (1 - (f_y / N)) / (1 - confidence)
print("Conviction = {}".format(round(conviction, 4)))
# + [markdown] colab_type="text" id="83FIxuA5f5SS"
# #### Exercise 03: Data Loading
# + colab={} colab_type="code" id="X3EuKRoKf5SU"
online = pandas.read_excel(
io="./Online Retail.xlsx",
sheet_name="Online Retail",
header=0
)
# + colab={"base_uri": "https://localhost:8080/", "height": 359} colab_type="code" id="Np-8F3Vyf5Sc" outputId="8082c199-40ec-4d5a-84b7-f1454e47e8c9"
online.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="vRV-SDQqf5Sj" outputId="41a7ffdb-4357-49a8-b187-adece691a48d"
online.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="gcvjFxLIf5Sw" outputId="53f6c2ef-13bb-4900-b89d-c63e39475725"
print("Data dimension (row count, col count): {dim}".format(dim=online.shape))
print("Count of unique invoice numbers: {cnt}".format(cnt=online.InvoiceNo.nunique()))
print("Count of unique customer ids: {cnt}".format(cnt=online.CustomerID.nunique()))
# + [markdown] colab_type="text" id="CMtjTyrUf5S-"
# #### Exercise 04: Data Cleaning and Formatting
# + colab={} colab_type="code" id="VWOs4yoKf5TB"
# create new column called IsCPresent
online['IsCPresent'] = (
# looking for C in InvoiceNo column
online['InvoiceNo']
# convert column to string type for the apply function below
.astype(str)
# set element to 1 if C present otherwise 0, this will helpful for step 2
.apply(lambda x: 1 if x.find('C') != -1 else 0)
)
# + colab={} colab_type="code" id="sHbaTS8af5TK"
online1 = (
online
# filter out non-positive quantity values
.loc[online["Quantity"] > 0]
# remove InvoiceNos starting with C
.loc[online['IsCPresent'] != 1]
# column filtering
.loc[:, ["InvoiceNo", "Description"]]
# dropping all rows with at least one missing value
.dropna()
)
# + colab={"base_uri": "https://localhost:8080/", "height": 359} colab_type="code" id="583aPQJXf5TT" outputId="352756fd-9c02-4bea-975d-bf89b9b7f961"
online1.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="TMu5T1mHf5TZ" outputId="08dbf241-5107-4dcb-fa2b-31e677c93dd6"
print("Data dimension (row count, col count): {dim}".format(dim=online1.shape))
print("Count of unique invoice numbers: {cnt}".format(cnt=online1.InvoiceNo.nunique()))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="_7-2yDpXf5Th" outputId="49027f3e-b7b9-42e0-898c-57fea96ab3b8"
# extract unique invoice numbers as list
invoice_no_list = online1.InvoiceNo.tolist()
invoice_no_list = list(set(invoice_no_list))
print("Length of list of invoice numbers: {ln}".format(ln=len(invoice_no_list)))
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="voKBMeFuf5To" outputId="465b403a-a09c-4f56-d587-20a31d818014"
# take subset of invoice number for exercises
subset_invoice_no_list = invoice_no_list[0:5000]
print("Length of subset list of invoice numbers: {ln}".format(ln=len(subset_invoice_no_list)))
# + colab={} colab_type="code" id="JHrfPpAzf5Tw"
# filter data set down to based on
# subset of invoice number list
online1 = online1.loc[online1["InvoiceNo"].isin(subset_invoice_no_list)]
# + colab={"base_uri": "https://localhost:8080/", "height": 359} colab_type="code" id="OPbiPimqf5T3" outputId="c6183dfd-e3ba-42a5-a6a8-18012433ba23"
online1.head(10)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="UBaP-HUqf5T9" outputId="1bccd28d-51e9-4e00-ae14-c17bf24c9185"
print("Data dimension (row count, col count): {dim}".format(dim=online1.shape))
print("Count of unique invoice numbers: {cnt}".format(cnt=online1.InvoiceNo.nunique()))
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="gvXo--fgf5UD" outputId="a4dff520-be59-45a9-8721-9115ce7bf592"
invoice_item_list = []
for num in list(set(online1.InvoiceNo.tolist())):
# filter data set down to one invoice number
tmp_df = online1.loc[online1['InvoiceNo'] == num]
# extract item descriptions and convert to list
tmp_items = tmp_df.Description.tolist()
# append list invoice_item_list
invoice_item_list.append(tmp_items)
print(invoice_item_list[1:5])
# + [markdown] colab_type="text" id="vn63DJR4f5UI"
# #### Exercise 05: Data Encoding
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="YFELanOIf5UM" outputId="28c080ad-fc08-40ce-aac9-a7417cb1ef1f"
online_encoder = mlxtend.preprocessing.TransactionEncoder()
online_encoder_array = online_encoder.fit_transform(invoice_item_list)
print(online_encoder_array)
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="eHjXS-Y-f5UW" outputId="82428cc4-67e8-4e23-c4ec-88571a8e7d0b"
online_encoder_df = pandas.DataFrame(
online_encoder_array,
columns=online_encoder.columns_
)
# this is a very big table, so for more
# easy viewing only a subset is printed
online_encoder_df.loc[
4970:4979,
online_encoder_df.columns.tolist()[0:8]
]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="8m78q5tKf5Uc" outputId="b63b24ba-cfa2-4dfa-cf6f-4907ed73fbf6"
print("Data dimension (row count, col count): {dim}".format(dim=online_encoder_df.shape))
# + [markdown] colab_type="text" id="0a-pXy-Df5Uj"
# #### Exercise 06: Apriori Algorithm
# + colab={"base_uri": "https://localhost:8080/", "height": 49} colab_type="code" id="KykyiljXf5Um" outputId="d64c8b98-7b41-44ab-cff0-0d68ba0916cb"
# default minimum support = 0.5
# does not use colnames (item names)
mod = mlxtend.frequent_patterns.apriori(online_encoder_df)
mod
# + colab={"base_uri": "https://localhost:8080/", "height": 266} colab_type="code" id="q9HszqxRf5Uu" outputId="28122b0e-41f6-4031-d451-56e887c193fa"
mod_minsupport = mlxtend.frequent_patterns.apriori(
online_encoder_df,
min_support=0.01
)
mod_minsupport.loc[0:6]
# + colab={"base_uri": "https://localhost:8080/", "height": 266} colab_type="code" id="GrYAJwlZf5U2" outputId="d8743786-47f3-4f24-8d02-696fc5fe719d"
# add colnames for easier interpretability
mod_colnames_minsupport = mlxtend.frequent_patterns.apriori(
online_encoder_df,
min_support=0.01,
use_colnames=True
)
mod_colnames_minsupport.loc[0:6]
# + colab={"base_uri": "https://localhost:8080/", "height": 266} colab_type="code" id="6Im6JOSpf5U8" outputId="69b2a640-52fa-4f2c-90e5-5d5521753d24"
mod_colnames_minsupport['length'] = (
mod_colnames_minsupport['itemsets'].apply(lambda x: len(x))
)
mod_colnames_minsupport.loc[0:6]
# + colab={"base_uri": "https://localhost:8080/", "height": 80} colab_type="code" id="Z3EeXNl4f5VE" outputId="aac55195-710a-44ce-e967-3edba9f927c0"
mod_colnames_minsupport[
mod_colnames_minsupport['itemsets'] == frozenset(
{'10 COLOUR SPACEBOY PEN'}
)
]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="Ksj9KdwZf5VM" outputId="745ca973-43df-42fa-ac25-d4c23f5f355e"
## ORDER OF ITEMSETS DIFFERS
mod_colnames_minsupport[
(mod_colnames_minsupport['length'] == 2) &
(mod_colnames_minsupport['support'] >= 0.02) &
(mod_colnames_minsupport['support'] < 0.021)
]
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="OBPXtIZJf5VQ" outputId="ab0fa2e6-e70c-46eb-957b-a880fba769e0"
mod_colnames_minsupport.hist("support", grid=False, bins=30)
plt.xlabel("Support of item")
plt.ylabel("Number of items")
plt.title("Frequency distribution of Support")
plt.show()
# + [markdown] colab_type="text" id="hXPCVbvLf5VU"
# #### Exercise 07: Association Rules
# + colab={"base_uri": "https://localhost:8080/", "height": 368} colab_type="code" id="frXsrktaf5VX" outputId="8f31aa06-56a8-41fa-e107-b8a7d4f8eb8f"
## ROW ORDER SLIGHTLY DIFFERENT
rules = mlxtend.frequent_patterns.association_rules(
mod_colnames_minsupport,
metric="confidence",
min_threshold=0.6,
support_only=False
)
rules.loc[0:6]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="grY0vp4lf5Ve" outputId="98958b8f-7395-4fc5-e682-516d520bdcee"
print("Number of Associations: {}".format(rules.shape[0]))
# + colab={"base_uri": "https://localhost:8080/", "height": 402} colab_type="code" id="EZwCkWt3f5Vm" outputId="7827163a-baae-4748-d1c9-c0e2e697c827"
## ORDER OF DF DIFFERENT SLIGHTLY
rules2 = mlxtend.frequent_patterns.association_rules(
mod_colnames_minsupport,
metric="lift",
min_threshold=50,
support_only=False
)
rules2.loc[0:6]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="SdSvkG4jf5Vs" outputId="b9d6675f-8690-4ba5-952f-3e5a599e2900"
print("Number of Associations: {}".format(rules2.shape[0]))
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="LPkOE9dzf5Vy" outputId="3caf4d70-ab40-4244-dcef-5e10e4f25508"
rules.plot.scatter("support", "confidence", alpha=0.5, marker="*")
plt.xlabel("Support")
plt.ylabel("Confidence")
plt.title("Association Rules")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="tiqS3o2Of5V8" outputId="75b8a82a-4c5d-4b32-b4d4-5b9619019ca5"
rules.hist("confidence", grid=False, bins=30)
plt.xlabel("Confidence of item")
plt.ylabel("Number of items")
plt.title("Frequency distribution of Confidence")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="GfMf2JU8f5WF" outputId="116cd4cd-6f12-4641-8c66-7e1119b27894"
rules.hist("lift", grid=False, bins=30)
plt.xlabel("Lift of item")
plt.ylabel("Number of items")
plt.title("Frequency distribution of Lift")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="xumdeHcDf5WQ" outputId="333a5a9d-a39a-43ba-9c9f-f4efb881985f"
rules.hist("leverage", grid=False, bins=30)
plt.xlabel("Leverage of item")
plt.ylabel("Number of items")
plt.title("Frequency distribution of Leverage")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="-OMNoQhDf5WV" outputId="d86cf803-8f45-4c65-f090-3d5ab7b8d620"
plt.hist(rules[numpy.isfinite(rules['conviction'])].conviction.values, bins = 30)
plt.xlabel("Conviction of item")
plt.ylabel("Number of items")
plt.title("Frequency distribution of Conviction")
plt.show()
|
Exercise01-Exercise07/Exercise01-Exercise07.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "832f2701f2fb2f8d43d940f8664b5186", "grade": false, "grade_id": "introduction", "locked": true, "schema_version": 3, "solution": false, "task": false}
# # R for Neuroscience (r4n)
# Made with 💖 by <NAME>
#
# 
# *Source: https://giphy.com/gifs/back-to-school-SSirUu2TrV65ymCi4J*
#
# ## Welcome NURC 2021!
# I created this worksheet for the 2021 [Neuroscience Undergraduate Research Conference (NURC)](https://ubcneuroscienceclub.wixsite.com/uncweb/nurc-2021). This worksheet is meant for beginners with little to no R background. I will introduce you to the modern tools such as tidyverse to show you how to conduct a clean and modern data analysis.
#
# ## Why should you learn R?
# During my undergrad, my PhD mentor told me to learn R if I wanted to attend graduate school.
#
# 1. R is free (unlike Matlab, SPSS, or Excel)! As well, many textbooks and resources (like this one) is open source and widely available. As well, there is a huge online R community.
#
# 2. R is the future of statistical analysis (present as well?). During the summer and fall, I have interviewed with many potential supervisors. Most assumed that you know R already or that you will be willing to learn in grad school. As such, it is a very useful skill to pick up.
#
# 3. R is very versatile. I have used R in so many different ways from statistical tests, web development, and modeling.
#
# 4. Analyses conducted in R are reproducible, reusable, and shareable. I was an undergraduate research assistant and my PhD mentor was showing me the data analysis and plots he created in Excel. He told me to do the other data set but I totally forgot how to do it when I got around to it.
#
# ## About the instructors
# [<NAME>](http://andrewcli.me) <br>
# [<NAME>](https://tigerthepro.github.io/TigerWu/)
#
# ## Contents
# 1. Introduction to Jupyter Notebooks
# * Text cells
# * Markdown cheatsheet
# * Code cells
# * Equations
# * Comments
# * Check your answers
# 2. Welcome to the tidyverse
# 3. Data collection
# * `readr`
# 4. Data wrangling
# * `select()`
# * `filter()`
# * `rename()`
# * `mutate()`
# 5. Data visualization
# * Themes
# * Grammar of Graphics
# 6. Putting it all together
# * Data collection
# * Data wrangling
# * Data visualization
# 7. Future direction
#
# Use the cell below to load (install if you haven't already) the packages we will need:
# +
# uncomment and run this cell if you need to install tidyverse.
# install.packages('tidyverse')
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "70932dacf2c490040082057b7d2301ee", "grade": false, "grade_id": "libraries", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Run this cell before continuing.
source("tests_nurc_2021.R")
suppressPackageStartupMessages(library(tidyverse))
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "5581133fb64adf3be9cc6066c5d1496f", "grade": false, "grade_id": "IntroJupyter1", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## 1. Introduction to Jupyter Notebooks
# If you go on to statistic or computer science courses, often time they will use Jupyter Notebooks. As such it is worthwhile to familiarize yourselves to it. This section will show you what Jupyter notebooks are and what they can do!
#
# ### 1.1 Text cells
# In a notebook, each rectangle is called a cell. This one is a text cell because it contains text. You can edit text cells by double clicking it. After you are done, simply press `control + enter` (mac and pc)or click run. Text cells are written in markdown. Markdown is a very simple markup language to format and edit the text. **Note** Jupyter lab does not have spell check, so be careful if you want to submit a future assignment/project!
#
# #### 1.1.1 Markdown cheatsheet
#
# Double click this cell to take a look at some common markdown tools:
#
# **This is bold**
#
# *This is italics*
# ## This is a header
# Here is an ordered list:
# 1. Thing 1
# 2. Thing 2
#
# Here is an unordered list:
# * Order doesn't matter
# * Still doesn't matter
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c2d307a0efe039ed7b42ed53ca41433a", "grade": false, "grade_id": "IntroJupyter2", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 1.2 Code cells
# Code cells allow you to input R or Python code. Pressing `control + enter` or clicking run will make Jupyter run the whole cell. You can run the entire sheet from top to bottom if you click the `run all` tab in the `cell` tab.
#
# Try to print "Hello World!"
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "68451e04558b38ee67bcffd8f74e4767", "grade": false, "grade_id": "IntroJupyter2_code", "locked": true, "schema_version": 3, "solution": false, "task": false}
print("Hello World!")
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "5d047f2892000a0a0cd47104e84f93f4", "grade": false, "grade_id": "IntroJupyter3", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 1.3 Equations
# In markdown, you can enter equations as well. Double click this cell to see how you can create in line equations such as this $Y = \beta_0 + \beta_1 = X$ or in the center like this
#
# $$
# \bar{X} = \frac{\sum{X_1}}{N} = \frac{X_1 + X_2 + X_3 + ... + X_N}{N}
# $$
#
# As you can see, Jupyter notebooks have become very popular because of how much you can do with them.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "792a96647b1b1d9b23413d858c6f622a", "grade": false, "grade_id": "IntroJupyter4", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 1.4 Comments
# In all programming languages, you can comment out chunk codes or sentences. This is so that you and others can more easily understand your thoughts and workflow. As well, it is really good for debugging and trying things out. In R, you comment things with `#`, everything behind # will be ignored. Other languages will have different syntax. In Jupyter notebooks, you can comment out large chunks of code by selecting everything you want to run and use the hot keys `command or control + /`. In RStudio, you can do this by using the hot keys `control + shift + c`.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "a344fcd49d764d6b9078ba143c4da2a2", "grade": false, "grade_id": "IntroTests1", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 1.5 Check your answer
#
# I have written tests for your answers and the autograder. You can check your for the correctness of your code by simply running the cells directly underneath the cells with a question. However, you must load the solution first (just as you need to load packages) before you can use the test functions. In every notebook, you will be prompt to run the packages needed and my tests.
# +
# you will find this cell near the top of every notebook
# Run this cell before continuing.
source("tests_nurc_2021.R")
# example of a test from this notebook
# test_2.3()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "9c4141dd1e5c0d98cdabbc35bd4c7b4d", "grade": false, "grade_id": "intro_text_1", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 1.1
#
# Multiple choice:
#
# Welcome to your first question! When you are working through this worksheet make sure you *read* and *follow* the instructions.
#
# To answer this multiple choice question, assign your answer to `answer1.1` and make sure your response(s) are in upper case and in quotation marks ("A").
#
# answer1.1 <- c(FILL_THIS_IN, FILL_THIS_IN)
#
# A. Correct
#
# B. Correct
#
# C. Incorrect
#
# D. Incorrect
# + deletable=false nbgrader={"cell_type": "code", "checksum": "dac0ff2ec3281a46961b968f074f0c66", "grade": false, "grade_id": "intro_q1_q", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "adc3b00bad8db2239d199c7c8f788977", "grade": true, "grade_id": "intro_q1_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_1.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "4d286fedf8c4baf3a5ed86ac243b6140", "grade": false, "grade_id": "intro_text_2", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 1.2
#
# True or false:
#
# R is similar to SPSS.
#
# To answer this true or false question, assign your response to `answer1.2`. Make sure your submission is in all lower case and to surround your answer in quotation marks ("true"/"false").
#
# answer1.2 <- FILL_THIS_IN
# + deletable=false nbgrader={"cell_type": "code", "checksum": "46be296116516233c965f488fd3f0de7", "grade": false, "grade_id": "intro_q2_q", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "836a28155a2235887593eb6a59951652", "grade": true, "grade_id": "intro_q2_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_1.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "eef7910368806ee30cc82e481898a75e", "grade": false, "grade_id": "intro_q3_text", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 1.3
#
# True or False:
#
# This problem set is for marks.
#
# To answer this true or false question, assign your response to `answer1.3`. Make sure your submission is in all lower case and to surround your answer in quotation marks ("true"/"false").
#
# answer1.3 <- FILL_THIS_IN
# + deletable=false nbgrader={"cell_type": "code", "checksum": "d273206666aab14bdd4495ebde9ce3b9", "grade": false, "grade_id": "intro_q3_q", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "260be5b6e470e0695997ff9e05fec703", "grade": true, "grade_id": "intro_q3_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_1.3()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "50f4bbbd929143f00232e127a02fb0ae", "grade": false, "grade_id": "variable_intro", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 2. Welcome to the tidyverse
# Tidyverse is a *collection* of packages designed for data science (often called a "meta"--package). When you install and load tidyverse version 1.2.0, you are actually several core pacakges including: `dplyr`, `forcats`, `ggplot2`, `purrr`, `readr`, `stringr`, `tibble`, and `tidyr`. These packages all share a high-level design philosophy and low-level grammer and data structures. This makes it so that learning one package makes it easier to learn the others as well.
#
# #### Installation
# * In order to use an R package, you need to install the package. Note that you only need to install packages once.
#
# install.packages("tidyverse")
#
# * After you have installed the package, you need to load the packages you want to use in your current R session.
#
# library(tidyverse)
#
# #### Core tidyverse packages
#
# **Bold** denotes packages used today
#
# | Package name | Description | Cheetsheet |
# |---------------|------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|----------------------------------------------------------------------------------------------------------------|
# | **ggplot2** | ggplot2 is a system for declaratively creating graphics, based on The Grammar of Graphics.<br>You provide the data, tell ggplot2 how to map variables to aesthetics, what graphical <br>primitives to use, and it takes care of the details. | Click [here](https://rstudio.com/wp-content/uploads/2015/03/ggplot2-cheatsheet.pdf) for cheatsheet! |
# | **dplyr** | dplyr provides a grammar of data manipulation, providing a consistent set of verbs that <br>solve the most common data manipulation challenges. | Click [here](<br>https://github.com/rstudio/cheatsheets/blob/master/data-transformation.pdf) for cheatsheet! |
# | tidyr | tidyr provides a set of functions that help you get to tidy data. Tidy data is data with <br>a consistent form: in brief, every variable goes in a column, and every column is a variable | Click [here](<br>https://rstudio.com/wp-content/uploads/2015/02/data-wrangling-cheatsheet.pdf) for cheatsheet! |
# | **readr** | readr provides a fast and friendly way to read rectangular data (like csv, tsv, and fwf).<br>It is designed to flexibly parse many types of data found in the wild, while still cleanly <br>failing when data unexpectedly changes. | Click [here](<br>https://github.com/rstudio/cheatsheets/blob/master/data-import.pdf) for cheatsheet! |
# | purrr | purrr enhances R’s functional programming (FP) toolkit by providing a complete and consistent <br>set of tools for working with functions and vectors. Once you master the basic concepts, <br>purrr allows you to replace many for loops with code that is easier to write and more expressive. | Click [here](<br>https://github.com/rstudio/cheatsheets/blob/master/purrr.pdf) for cheatsheet! |
# | tibble | tibble is a modern re-imagining of the data frame, keeping what time has proven to be effective, <br>and throwing out what it has not. Tibbles are data.frames that are lazy and surly: they do less <br>and complain more forcing you to confront problems earlier, typically leading to cleaner, more <br>expressive code. | Click [here](<br>https://miro.medium.com/max/700/1*fEGdnyXLzgeftfCLwvBZ5A.jpeg) for cheatsheet! |
# | stringr | stringr provides a cohesive set of functions designed to make working with strings as easy as <br>possible. It is built on top of stringi, which uses the ICU C library to provide fast, correct <br>implementations of common string manipulations. | Click [here](<br>https://evoldyn.gitlab.io/evomics-2018/ref-sheets/R_strings.pdf) for cheatsheet! |
# | forcats | forcats provides a suite of useful tools that solve common problems with factors. R uses factors <br>to handle categorical variables, variables that have a fixed and known set of possible values. | Click [here](http://www.flutterbys.com.au/stats/downloads/slides/figure/factors.pdf) for cheatsheet! |
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "217a9ea27897363ce4324242659059f2", "grade": false, "grade_id": "variable_q1_text", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 2.1
# Use the `library` function to fire up tidyverse! {Points:1}
#
# library(FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "3307e8dc69184a21cb72ac87fa702b46", "grade": false, "grade_id": "variable_q1_question", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "69e181f1794036c4bc461e1e7f87db00", "grade": true, "grade_id": "variable_q1_answer", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_2.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "22e108dfa6aa31f577bd511d2b1a2c11", "grade": false, "grade_id": "variable_q2_text", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 2.2
# Tidyverse crossword!
#
# You can complete this crossword about `tidyverse` functions, specifically, `dplyr`, `ggplot2`, and `readr`. You might not know about some of these functions so googling is highly encouraged. You do not need parentheses but some answers have underscores.
#
# To answer this question, assign the across words to lower case "a" followed by the number. Assign the down words to a lowercase "d" followed by the number. Make sure you answer all of the across questions in ascending order first before moving on to down for the autograder. Follow the guide below {Points: 1}
#
# a3 <- FILL_THIS_IN
# a5 <- FILL_THIS_IN
# ...
# ...
# d12 <- FILL_THIS_IN
#
# You can solve for this puzzle [here](https://crosswordlabs.com/embed/2020-12-08-749) as well!
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "5456cf69d666501af53419a364a7b52f", "grade": false, "grade_id": "crossword", "locked": true, "schema_version": 3, "solution": false, "task": false}
# <img src="img/crossword.png" width="800" height="800">
# + deletable=false nbgrader={"cell_type": "code", "checksum": "778c66a382327c7b0612c08782d18c0d", "grade": false, "grade_id": "variable_q2_question", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "9624157d2b930e765c8c1c9bf701acbf", "grade": true, "grade_id": "variable_q2_answer", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_2.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "353e6398ebc1ed9b6282b52c018365f4", "grade": false, "grade_id": "data_collection", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 3. Data collection
# 
# *Source: https://giphy.com/gifs/brain-anatomy-Vn9JVHDAzYw1O*
#
# #### Background
# For this workshop, we will be working through a MRI and Alzheimer's data set made publicly available by Kaggle. You can find the original data set [here](https://www.kaggle.com/jboysen/mri-and-alzheimers). The original data was made available by the [Open Access Series of Imaging Studies (OASIS)](http://www.oasis-brains.org/) project. This was aimed at making MRI data sets freely available to the scientific community. OASIS was made available by the [Neuroinformatics Research Group (NRG)](http://nrg.wustl.edu/) at Washington University and the [Howard Hughes Medical Institude (HHMI)](http://www.hhmi.org/) at Harvard University.
#
# #### Alzheimer's Disease
# Alzheimer's disease is an irreversible, progressive brain disorder that slowly destroys memory, cognitive functions, and eventually, the ability to carry out the simplest tasks. In late onset, symptoms typically appear in their mid-60s while early onset symptoms typically appear between a person's 30s and mid 60s. In 1906, Dr. <NAME> noticed significant changes in the brain tissue of a women who died of an unusual mental illness. She experienced memory loss, language problems, and unpredictable behavior. An autopsy showed that her brain had many abnormal clumps (amyloid plaques) and tangled bundles of fibers (neurofibrillary). The damage initially takes place in the hippocampus but spreads to other brain regions.
#
# Experts predict that 5.5 million Americans age 65 and older may suffer from Alzheimer's. As well many under 65 suffer from it as well.
#
# #### Cross sectional data
# Kaggle offers two data sets but we will be working with the cross sectional data set `oasis_cross-sectional.csv`.
#
# *Cross-sectional MRI Data in Young, Middle Aged, Nondemented and Demented Older Adults:* This set consists of a cross-sectional collection of 416 subjects aged 18 to 96. For each subject, 3 or 4 individual T1-weighted MRI scans obtained in single scan sessions are included. The subjects are all right-handed and include both men and women. 100 of the included subjects over the age of 60 have been clinically diagnosed with very mild to moderate Alzheimer's disease (AD). Additionally, a reliability data set is included containing 20 nondemented subjects imaged on a subsequent visit within 90 days of their initial session.
#
# | Variable | Description | Type |
# |----------|----------------------------------------------------------------------------------------------------------------------------------------------------------------------------------------|---------|
# | ID | Unique Id of the patient | String |
# | M/F | Gender | Boolean |
# | Hand | handedness | Boolean |
# | Age | Age in years | Integer |
# | Educ | Education level in years | Integer |
# | SES | Socioeconomic status as assessed by the Hollingshead Index of Social Position and <br>classified into categories from 1 (highest status) to 5 (lowest status) | Integer |
# | MNSE | Mini Mental State Examination is a test of cognitive function and classified via<br>(range is from 0 = worst to 30 = best) | Integer |
# | CDR | Clinical Dementia Rating (0 = no dementia, 0.5 = very mild AD, 1 = mild AD, <br> 2 = moderate AD, 3 = Severe AD) | Float |
# | eTIV | Estimated Total Intracranial Volume | Integer |
# | nWBV | Normalized Whole Brain Volume expressed as a percent of all voxels in the atlas-masked image <br>that are labeled as gray or white matter by the automated tissue segmentation process | Float |
# | ASF | Atlas scaling factor (unitless). Computed scaling factor that transforms native-space brain <br>and skull to the atlas target (i.e., the determinant of the transform matrix) | Float |
# | Delay | Delay time (contrast) | Integer |
#
# #### Reading in the data set
# Using the `read_csv` function from the `readr` package (that is in the tidyverse package), we can load this data set into R.
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c1b06167b89693bbd00d1d345db9c058", "grade": false, "grade_id": "mc_31", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 3.1
#
# Multiple choice:
#
# What is the `oasis_cross-sectional.csv` data set about?
#
# To answer this multiple choice question, assign your answer to `answer3.1` and make sure your response(s) are in upper case and in quotation marks ("A").
#
# answer3.1 <- FILL_THIS_IN
#
# A. Study conducted looking at predictors for drug addiction
#
# B. A data set that uses electroencephalography (EEG) to show brain activity in people with Alzheimer's disease
#
# C. A longitudinal MRI data set that looks at nondemented and demented oldr adults
#
# D. A cross-sectional MRI data set with young, middle-aged, nondemented, and demented older adults
#
# E. A data set that uses positron emission tomography (PET)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "76aec86891362a9c7e295301f0d82838", "grade": false, "grade_id": "mc31_answer", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "4150b9ae17062272bc15c5979ccb9a66", "grade": true, "grade_id": "mc31_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_3.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "fa6d28b2f457cb739c65f792732025ba", "grade": false, "grade_id": "mc32", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 3.2
# True or False:
#
# The Clinical Dementia Rating scale is classified from 1 (no Dementia) to 30 (severe Dementia).
#
# To answer this true or false question, assign your response to `answer3.2`. Make sure your submission is in all lower case and to surround your answer in quotation marks ("true"/"false").
#
# answer3.2 <- FILL_THIS_IN
# + deletable=false nbgrader={"cell_type": "code", "checksum": "cb64d1ef6c9a210889c678140df2e08a", "grade": false, "grade_id": "mc32_answer", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "5eb71163af85f202127389dd7ef8690b", "grade": true, "grade_id": "mc32_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_3.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c31ea013e847ab791e68c8fd916f043a", "grade": false, "grade_id": "read_in", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 3. 3
# If we just load the data into R, it will be printed on the screen but you cannot do anything to it. If we want to work with it, we need to give it a name so that we can call upon it and manipulate it moving forward. Create a new variable called `mri_data` and assign the `oasis_cross-sectional.csv` data set to it.
#
# FILL_THIS_IN <- read_csv("data/FILL_THIS_IN")
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9ff14f637099c26b7c85b8f316d37520", "grade": false, "grade_id": "read_in_answer", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(mri_data) # prints the first 6 lines
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "553745c69d0ff7bacaf4f13c85bbce98", "grade": true, "grade_id": "readin_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_3.3()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "2b51ab4c94da3ce87a171e8f8f96d9c6", "grade": false, "grade_id": "mc34_question", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 3.4
#
# Now that we have read in the data, let's take a look at it. In question 3.3, we used the `head()` function to look at the first 6 entries. Now, use the `tail()` function to take a look at the last 6 entries and find out the last patient's (`OAS1_0395_MR2`) age.
#
# To answer this question, assign your answer to `answer3.4`.
#
# answer3.4 <- FILL_THIS_IN
# + deletable=false nbgrader={"cell_type": "code", "checksum": "200a18bf35af6ba8c74925b22dd3b5ac", "grade": false, "grade_id": "34_answer", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "b6ecd43a621c5a4e356a039bbc05e04a", "grade": true, "grade_id": "mc34_test", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_3.4()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "61d7cf568cd01b8831725b7acdfd8dc3", "grade": false, "grade_id": "cell-4ad3707e95256dc6", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 4. Data Wrangling
#
# Before we create any graphs or models, we need to tweak the data to make it easier to work with. Some common manipulations include renaming variables or creating new variables. To do this, we will use the `dplyr` package. We will look at three of the most common functions in this package:
#
# 1. select()
# 2. filter()
# 3. mutate()
#
# <img src="img/dplyr.jpg" width="500" height="300">
#
# *Source: <NAME>*
#
# ### 4.1 `select()`
# The `select()` function allows you to select and work with the variable you want and find relevant.
#
# <img src="img/select.png" width="500" height="300">
#
# *Source: https://datacarpentry.org/r-intro-geospatial/06-dplyr/*
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "d3df73421f8fc9e7ea8b01a2f1b471ba", "grade": false, "grade_id": "cell-cd741374a2ff558f", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.1
# Say we are only interested in the age of the participants. Use the select function on `mri_data` so that only have the age of the participants.
#
# To answer this question, call the new data frame `answer4.1`
#
# answer4.1 <- select(mri_data, FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2e09e6beea7ac7bdae034550edc855ab", "grade": false, "grade_id": "cell-696091d66e2dbaf9", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.1)
# + nbgrader={"grade": false, "grade_id": "cell-cc8c7a7042cfa644", "locked": false, "schema_version": 3, "solution": false, "task": false}
test_4.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "7d74af7dce09e4cc08fde1880d5bc09c", "grade": false, "grade_id": "cell-614525b495eaa1c5", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.2
# Now, we are only interested in the demographics information. Select ID, M/F, Hand, Age, Educ, and SES.
#
# We can use this code:
#
# select(mri_data, ID, "M/F", Hand, Age, Educ, SES)
#
# But there is a much faster way to select multiple columns using ":". Use the select function and : to create a new variable called `answer4.2` that contains the demographic data.
#
# answer4.2 <- (FILL_THIS_IN, FILL_THIS_IN:FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "e66723197e55fe5c7c887fd1be02e29a", "grade": false, "grade_id": "cell-5e627fd68fdf67f9", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.2)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "b937a0e89981390c18793f71f41fa018", "grade": true, "grade_id": "cell-c10f35a40e9a818a", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "dd6baad4c39e75762ae35f7277a68336", "grade": false, "grade_id": "cell-ac7bef9f2bf2ab30", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.4
#
# Now, select *every* column except Delay. We will not be needing the variable moving forward.
#
# We can manually type out all of the columns we want
#
# select(mri_data, ID, "M/F", Hand, Age, Educ, SES ... ASF)
#
# Or, we can simply tell R which columns we *don't* want via the minus sign. create a new variable called `answer4.4` with every column except Delay.
#
# select(FILL_THIS_IN, -FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "6d9a771bda0dd01a5298f7f837e06cf8", "grade": false, "grade_id": "cell-d78858832fe238e1", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.3)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "da8777f07820b23f4ade2633d95dcc30", "grade": true, "grade_id": "cell-49e7ff2c14b6abaf", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.3()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "c94f4896caa82b5f739f5ffe00c764b8", "grade": false, "grade_id": "cell-64eaa814434123b0", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.4
# If you want to look at the gender distribution, you will encounter an error because of how it is named. We need to use quotations in order to use it. Let's rename this variable to something that is easier to work with. Rename M/F to `gender` and create a new object called `answer4.4`. Remember to use our new data frame `answer4.3`.
#
# Hint there is a `rename()` function!
#
# answer4.4 <- FILL_THIS_IN(answer4.3, gender = FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "26bdc4d5496a3368102a243ed8290aef", "grade": false, "grade_id": "cell-c81f7cc8e1f3dd30", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.4)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "61639f2bf10aaeb21e36a4d35939ae29", "grade": true, "grade_id": "cell-dcc7e59acdc8aea9", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.4()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "2c6974a16c118bd4a3a3e62eeba188a9", "grade": false, "grade_id": "cell-6dfd5bfbce392b2b", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 4.2 `filter()`
#
# We can use the `filter()` function to keep observations that meet our criteria.
#
# <img src="img/filter.jpg" width="600" height="300">
#
# *Source: <NAME>*
#
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "6c9c0b67c59c6d8073dfb90bebde74c6", "grade": false, "grade_id": "cell-61c1d0dc288a991d", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.5
#
# As you know, Alzheimer's mainly affect the older population. As such, we want to look filter our data so that we only have participants that are over 60 years old. Using `answer4.4` create a new object called `answer4.5` so that we only have older adults.
#
# answer4.5 <- filter(FILL_THIS_IN, FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "3801ce5898a1923fe16be94c5ad4df0d", "grade": false, "grade_id": "cell-749c5d0ab9006590", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.5)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3e0724531fec7a0aeac5290309b92248", "grade": true, "grade_id": "cell-5f518a84f1f01605", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.5()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "51dc3d487cf9e4b57ed35920c3c4f268", "grade": false, "grade_id": "cell-f3814a685c3dc383", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.6
#
# Now, to practice, from `answer4.5`, take only the data from females. Create a new object variable called `answer4.6`.
#
# answer4.6 <- FILL_THIS_IN(FILL_THIS_IN, FILL_THIS_IN == FILL_THIS_IN)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "909a632641f8ef6a4f83841705ebbff3", "grade": false, "grade_id": "cell-1ca7cacd67164887", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.6)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "3c1a52ecf7021dfa98335338e4babb1a", "grade": true, "grade_id": "cell-774967518f0d8ed0", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.6()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "66d88b002b256618834e59ed6c5b28e4", "grade": false, "grade_id": "cell-e4903a8edbae795b", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 4.3 `mutate()`
#
# We can use the `mutate()` function to create new variables while preserving existing ones. Mutate will create a new column at the end of your data set.
#
# <img src="img/mutate.jpg" width="400" height="400">
#
# *Source: <NAME>*
#
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "24c14fe21344e49055197881d5155b00", "grade": false, "grade_id": "cell-e4ae7713713b13ec", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 4.7
#
# We can calculate the "risk score" for developing Alzheimer's by adding up their socioeconomic status (SES), Mini Mental State Examination score (MMSE), and Clinical Dementia Rating (CDR). Create a new column in the `answer4.6` data frame called `risk_score`. Call this new data frame `answer4.7`. This new column should be the sum of the three predictive scores.
#
# answer4.7 <- FILL_THIS_IN(answer4.6,
# FILL_THIS_IN = FILL_THIS_IN + FILL_THIS_IN + FILL_THIS_IN)
#
# **NOTE:** Risk Score was completely made up for teaching purposes.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "b0556a680dbc7085fc74ce3682aecf69", "grade": false, "grade_id": "cell-0baad99234afbb91", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(answer4.7)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "4d78cedda8f083b8b6c9afa2cf71202b", "grade": true, "grade_id": "cell-2081062d4f6d98f2", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_4.7()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "accc83934023195ef2ab8451e2570d76", "grade": false, "grade_id": "cell-87f70c39efcd7eb2", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 5. Data Visualization
# In the final section of this workshop, we will cover how to make plots via `ggplot2`!
# <img src="img/ggplot2.jpg" width="400" height="400">
#
# *Source: <NAME>*
#
# ### 5.1 Grammar of graphics
# ggplot2 follows the grammar of graphics principle by <NAME>. Before this, there were functions for each graph - a line graph, bar graph, etc. Wilkinson did not like this because this became complex and unmanageable really fast (imagine creating a new function for every plot). As such, he set out to look for what all graphs have in common - the grammar of graphics. He proposed that these 8 constituents of graphics can become the bedrock for *every* plot.
#
# <img src="img/ggplot-2.png" width="400" height="400">
#
# *Source: https://www.science-craft.com/2014/07/08/introducing-the-grammar-of-graphics-plotting-concept/*
#
#
# ### 5.2 Themes
# I like to teach beginners how to use themes because the default theme R gives you is really ugly (in my opinion). You can easily change the theme by adding the theme name to your code. I created the graph below using the plot you will create in question 5.2. It is the same graph, I just applied all the different base themes available in R.
#
# plot + theme_bw() #example
#
# <img src="img/ggthemes.png" width="500" height="500">
#
# *Source: [<NAME>](http://andrewcli.me)*
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "66be18a0e2dac5d62d766ab10091f627", "grade": false, "grade_id": "cell-a94da89160424a78", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 5.1
#
# Let's bring it all together! Using our original data set `mri_data`, create a new data frame called `mri_tidy` that has the following:
#
# 1. Only contains participants who is 60 years old or greater
# 2. Create a new `risk_score` column which is the sum of SES, MMSE, and CDR
# 3. Rename the M/F column name to `gender`
#
# mri_tidy <- mri_data %>%
# FILL_THIS_IN(Age FILL_THIS_IN) %>%
# FILL_THIS_IN(FILL_THIS_IN = SES + MMSE + CDR) %>%
# FILL_THIS_IN(FILL_THIS_IN = "M/F")
#
# **NOTE:** The `%>%` is called the pipe operator. This comes in handy when you want to perform multiple functions like question 5.1. The pipe operator will forward the result of an expression into the next expression.
#
# filter(mri_data, SES == 3)
#
# Is the same as
#
# mri_data %>% filter(SES == 3)
# + deletable=false nbgrader={"cell_type": "code", "checksum": "1007238d7cc23d87b32bb794a702bdff", "grade": false, "grade_id": "cell-6de34feef99a37dc", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(mri_data)
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "b57893ddbb604ec804a1e58ae86d4536", "grade": true, "grade_id": "cell-3ea99e5e0bec60bc", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_5.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "093ca275e098792fefe5449fabcda8f6", "grade": false, "grade_id": "cell-23cf192dc2bb6ae4", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 5.2
#
# Let's take a look at relationship between the **Clinical Dementia Score** (CDR) and the **Mini Mental State Examination** (MMSE). As well, we want to see the difference between **genders**. Using the following parameters, create a boxplot to look at this relationship:
#
# * CDR should be on the x-axis and MMSE should be the y-axis
# * Color by gender
# * Set the transparency (alpha) to 0.8
# * Rename the x-axis, y-axis, and legend title so that it is descriptive and human readable
# * x-axis: `Clinical Dementia Score`
# * y-axis: `Mini Mental State Examination`
# * legend title: `Gender` (simply capitalize it)
# * Create a title `MRI and Alzheimer's`
# * Change the theme (optional)
#
# Store the result in a variable called `answer5.2`
#
# answer5.2 <- mri_tidy %>%
# ggplot(aes(x = as.factor(FILL_THIS_IN), y = FILL_THIS_IN, fill = FILL_THIS_IN)) +
# geom_boxplot(FILL_THIS_IN) +
# xlab(FILL_THIS_IN) +
# ylab(FILL_THIS_IN) +
# labs(fill = "Gender") +
# ggtitle(FILL_THIS_IN) +
# theme_classic() # you can play around and try different themes
#
# Or you can use this:
#
# answer5.2 <- mri_tidy %>%
# ggplot(aes(x = as.factor(FILL_THIS_IN), y = FILL_THIS_IN, fill = FILL_THIS_IN)) +
# geom_boxplot(FILL_THIS_IN) +
# labs(x = FILL_THIS_IN, y = FILL_THIS_IN, title = FILL_THIS_IN, fill = FILL_THIS_IN) +
# theme_classic() # you can play around and try different themes
# + deletable=false nbgrader={"cell_type": "code", "checksum": "dbc11a7c6cc8366dc472c576b7055e3e", "grade": false, "grade_id": "cell-7cd1d960a53ef016", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
answer5.2
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "c192446359a6a50bc0c8a6b7af1ccc31", "grade": true, "grade_id": "cell-e02b83869d434916", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_5.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "a0efae338e7a99d974e1152bf0e70835", "grade": false, "grade_id": "cell-394cb9fcb5729485", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### 6. Putting it all together
#
# In this last section, we will use everything we have learned thus far! We will be working on creating this graph below. __Note__ that this last section might be tricky but the goal of this section is to show you how customizable ggplot is. If there is something you haven't learned yet, I will provide the code.
#
# <img src="img/tbi_deaths.png" width="600" height="600">
#
# *Source: [<NAME>](http://andrewcli.me)*
#
# ### 6.1 Data collection - Traumatic Brain Injury (TBI)
#
# We will be taking the data from tidytuesday - a weekly social data project in R where users explore a new data set each week. This data set will look into [traumatic brain injury](https://github.com/rfordatascience/tidytuesday/blob/master/data/2020/2020-03-24/readme.md) and how common it is. The original data comes from the [CDC](https://www.cdc.gov/traumaticbraininjury/pdf/TBI-Surveillance-Report-FINAL_508.pdf) and [Veterans Brain Injury Center](https://dvbic.dcoe.mil/dod-worldwide-numbers-tbi).
#
# > Brain Injury Awareness Month, observed each March, was established 3 decades ago to educate the public about the incidence of brain injury and the needs of persons with brain injuries and their families (1). Caused by a bump, blow, or jolt to the head, or penetrating head injury, a traumatic brain injury (TBI) can lead to short- or long-term changes affecting thinking, sensation, language, or emotion.
#
# <img src="img/tbi_summary.png" width="800" height="200">
#
# *Source: [CDC](https://www.cdc.gov/mmwr/volumes/68/wr/mm6810a1.htm)*
#
# #### 6.1.1 Variables
#
# | Variable | Description | Type |
# |------------------|----------------------------------|--------|
# | age_group | Age group | string |
# | type | Type of measure | string |
# | injury_mechanism | Injury mechanism | string |
# | number_est | Estimated observed cases in 2014 | Integer|
# | rate_est | Rate/100,000 in 2014 | float |
#
# #### 6.1.2 The question we want to answer
#
# What are the leading causes of traumatic brain injury related deaths by age group?
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "05c626a8b9f84d69de6cd7ae17ba07d8", "grade": false, "grade_id": "cell-9bb81866008185ff", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 6.1
#
# Use the `read_csv()` function to load the data into this session. Name the variable `tbi_age`.
#
# FILL_THIS_IN <- FILL_THIS_IN('https://raw.githubusercontent.com/rfordatascience/tidytuesday/master/data/2020/2020-03-24/tbi_age.csv')
# + deletable=false nbgrader={"cell_type": "code", "checksum": "e8254b399d096647513f587f46dcb669", "grade": false, "grade_id": "cell-46c7f548488a001e", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(tbi_age) # prints the first 6 lines
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "17314e56293f26d6f163c8f116cda29c", "grade": true, "grade_id": "cell-35f12bc8c3292552", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_6.1()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "d70ecd0afd4685f4792acffeb34c3908", "grade": false, "grade_id": "cell-e28bb837e424d445", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 6.2
#
# Now that we have loaded the data, we need to clean it up a bit. Using `tbi_age` conduct the following manipulations:
#
# 1. Using the `filter()` function we will:
# * Notice that in the `age_group` column, **0-17** seems pretty useless. Let's get rid of it. Use the != operator.
# * We do not need the **Total** in `age_group`, let's get rid of it as well.
# * **Other or no mechanism specified** in the `injury_mechanism` column isn't that informative. Let's get rid of it.
# * We are only interested in **Deaths**. Use the == operator to filter that from `type`.
#
#
#
# 2. Using the `mutate()` function we will:
# * Create a new column called `pct` whereby we will divide `number_est` by the sum (`sum()`) of itself.
# * Turn all columns that are strings to factors. We will use the `factor()` function. Keep the original names of the columns.
# * Reorder `age_group` into ascending order. (See what happens to our plot if we skip this step!)
#
#
#
# 3. We need to get rid of missing values. We can do this via `na.omit()`.
#
#
#
# 4. Assign this new tidy data set to an object called `tbi_age_tidy`.
#
# Follow the following skeleton code provided to answer this question:
#
# FILL_THIS_IN <- FILL_THIS_IN %>%
# FILL_THIS_IN(FILL_THIS_IN != "0-17" & age_group != "FILL_THIS_IN"
# & FILL_THIS_IN != "Other or no mechanism specified",
# & FILL_THIS_IN == "Deaths") %>%
# na.omit() %>%
# FILL_THIS_IN(pct = number_est/sum(number_est)) %>%
# mutate(pct = pct * 10) %>%
# FILL_THIS_IN(age_group = factor(FILL_THIS_IN),
# injury_mechanism = factor(FILL_THIS_IN),
# type = factor(FILL_THIS_IN)) %>%
# mutate(age_group = fct_relevel(age_group, c("0-4", "5-14", "15-24",
# "25-34", "35-44", "45-54",
# "55-64", "65-74", "75+")))
# + deletable=false nbgrader={"cell_type": "code", "checksum": "09f0d03b98b102d382a9f4cfc853f435", "grade": false, "grade_id": "cell-e4ec9f65ca444f1f", "locked": false, "schema_version": 3, "solution": true, "task": false}
# your code here
fail() # No Answer - remove if you provide an answer
head(tbi_age_tidy) # prints the first 6 lines
# + deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "d3664dd34605e14629e56de7482a8d33", "grade": true, "grade_id": "cell-04d8a1292a8004d0", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
test_6.2()
# + [markdown] deletable=false editable=false nbgrader={"cell_type": "markdown", "checksum": "f2fd9b0caed9d37226d25f7802b9c46e", "grade": false, "grade_id": "cell-f85937b6f241986e", "locked": true, "schema_version": 3, "solution": false, "task": false}
# #### Question 6.3
#
# Now, using our new data, we will go on to answer our question, __What are the leading causes of traumatic brain injury related deaths by age group?__ As you can see, from our FMRI question, you can create a graph in R very easily. In this question, we will be trying to make this graph as aesthetically pleasing as possible. Here, I will show you some cool and easy ways to make your plot look really nice.
#
# 1. What data will we use?
# * We will use `tidy_age_group` for this graph.
#
#
# 2. What are our aesthetic mappings?
# * We will be mapping __age_group__ on the x-axis, __pct__ on the y-axis, and we will differentiate __injury_mechanism__ by color.
#
#
# 3. What geometric object will we use?
# * We will need a scatter plot and a line plot.
#
# *Nice! We have a plot now. Everything we do beyond step three can be optional but encouraged.*
#
#
# 4. We can create this graph without scales, but if we want to make it nicer, we can change the scales.
# * We can change the y-axis to percent via `scales` package.
# * We can change the default color R gives us.
#
#
# 5. Let's change the x-axis and y-axis to be something more human readable.
# * In this case, I would get rid of the x-axis and the y-axis names.
# * We can add a descriptive title as well.
# * Play around and add a subtitle or caption!
# * We can get rid of the legend title. The contents are self-explanatory and it seems redundant to add a title. Let's get rid of it.
#
#
# 6. I am not a big fan of the default theme that R gives us. Play around with the built in themes in R!
#
# *The graph we have now is publication ready. However, we can make it look even nicer.*
#
# 7. Finally, we can add the finishing touches to our graph.
# * I am not a fan of the legend placement. Let's change the position of the graph. Try changing it to be on top or on the bottom.
# * Let's make our title font size a bit bigger and make it bold.
# * Accordingly, let's make the axis text a bit easier to read. Make it **bold**.
#
#
# Use the following skeleton code to create this graph:
#
# *__Note__ that this question will not be graded. Play around with the colors, naming, themes, and more! Make this graph unique and aesthetically pleasing to look at!*
# -
# Step 1: Pipe the data set you will be using
tbi_age_tidy %>%
# Step 2: Add your aesthetic mappings
ggplot(aes(x = age_group, y = pct, color = injury_mechanism)) +
# Step 3: Choose what geometric object you will use
geom_point() +
geom_line(aes(group = injury_mechanism)) +
# Step 4: Change the scales
scale_y_continuous(labels = scales::percent) +
scale_color_manual(values = c("#1b9e77", "#d95f02", "#7570b3", #change to whatever color you want!
"#e7298a", "#66a61e", "#e6ab02")) +
# Step 5: Add human readable and descriptive titles
labs(x = "",
y = "",
color = "",
title = "Leading causes of TBI related deaths",
subtitle = "By age in 2014",
caption = "") +
# Step 6: Change the default theme to something more aesthetically pleasing
theme_minimal() +
# Step 7: Add the final touches!
theme(
legend.position = "bottom",
plot.title = element_text(face = "bold"),
axis.text = element_text(face = "bold")
)
# ### 7. Future directions
#
# If you enjoyed this workshop, please let me know by filling out our (very short) survey [here](https://ubc.ca1.qualtrics.com/jfe/form/SV_9BN4hBVzr9jRkZD)! I would love to create and host more workshops in the future. We covered a lot of topics today but we didn't really go over the basics or go deep into a specific topic. My goal was to show you the cool side of R to get you interested. If you want more workshops that go into more detail, shoot me an email!
|
content/problem_sets/nurc_2021-folder/nurc_2021.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/dlsun/pods/blob/master/Chapter_01_The_Data_Ecosystem/Chapter_1.4_Columns_and_Variables.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="jDl1ZPzz08kD"
# ## 1.3 Columns and Variables
#
# Recall that the columns of a tabular data set represent variables. They are the measurements that we make on each observation.
#
# As an example, let's consider the variables in the OKCupid data set. This data set does not have a natural index, so we use the default index (0, 1, 2, ...).
# + colab={} colab_type="code" id="-5F143Dj7OKh"
import pandas as pd
data_dir = "https://dlsun.github.io/pods/data/"
df_okcupid = pd.read_csv(data_dir + "okcupid.csv")
df_okcupid.head()
# + [markdown] colab_type="text" id="_CScd5-542Ci"
# ### 1.3.1 Types of Variables
#
# There is a fundamental difference between variables like `age` and `height`, which can be measured on a numeric scale, and variables like `religion` and `orientation`, which cannot be.
#
# Variables that can be measured on a numeric scale are called **quantitative variables**. Just because a variable happens to contain numbers does not necessarily make it "quantitative". For example, in the Framingham data set, the `SEX` column was coded as 1 for men and 2 for women. However, these numbers are not on any meaningful numerical scale; a woman is not "twice" a man.
#
# Variables that are not quantitative but take on a limited set of values are called **categorical variables**. For example, the variable `orientation` takes on one of three possible values (gay, straight, or bisexual), so it is a categorical variable. So is the variable `religion`, which takes on a larger, but still limited, set of values. We call each possible value of a categorical variable a "level". Levels are usually non-numeric.
#
# Some variables do not fit neatly into either classification. For example, the variable `essay1` contains users' answers to the prompt "What I’m doing with my life". This variable is obviously not quantitative, but it is not categorical either because every user has a unique answer. In other words, this variable does not take on a limited set of values. We will group such variables into an "other" category.
#
# Every variable can be classified into one of these three **types**:
# - quantitative,
# - categorical, or
# - other.
#
# The type of the variable often dictates how we analyze that variable, as we will see in the next two chapters.
# + [markdown] colab_type="text" id="xZlpl3SVELs4"
# ### 1.3.2 Selecting Variables
#
# Suppose we want to select the `age` column from the `DataFrame` above. There are three ways to do this.
# + [markdown] colab_type="text" id="vleFOfFYE82o"
# 1\. Use `.loc`, specifying both the rows and columns. (The colon `:` is Python shorthand for "all".)
# + colab={} colab_type="code" id="tYeLGCqtELgK"
df_okcupid.loc[:, "age"]
# + [markdown] colab_type="text" id="Zw1KEIjeFEbT"
# 2\. Access the column as you would a key in a `dict`.
# + colab={} colab_type="code" id="y7UY-QO_FEIT"
df_okcupid["age"]
# + [markdown] colab_type="text" id="kixEblNxFMjs"
# 3\. Access the column as an attribute of the `DataFrame`.
# + colab={} colab_type="code" id="Iivsx62yFMYm"
df_okcupid.age
# + [markdown] colab_type="text" id="MqwcwoI_FZ9p"
# Method 3 (attribute access) is the most concise. However, it does not work if the variable name contains spaces or special characters, begins with a number, or matches an existing attribute of `DataFrame`. For example, if `df_okcupid` had a column called `head`, `df_okcupid.head` would not return the column because `df_okcupid.head` is already reserved for something else.
# + [markdown] colab_type="text" id="TSTNcccjFx_Q"
# Notice that a `Series` is used here to store a single variable (across multiple observations). In the previous section, we saw that a `Series` can also be used to store a single observation (across multiple columns). To summarize, the `Series` data structure is used to store either a single row or a single column in a tabular data set. In other words, while a `DataFrame` is two-dimensional (containing both rows and columns), a `Series` is one-dimensional.
# + [markdown] colab_type="text" id="Oh5YgoA7GrgG"
# To select multiple columns, you would pass in a _list_ of variable names, instead of a single variable name. For example, to select both `age` and `religion`, either of the two methods below would work (and produce the same result):
# + colab={} colab_type="code" id="0WcfjkqGGvfw"
# METHOD 1
df_okcupid.loc[:, ["age", "religion"]].head()
# METHOD 2
df_okcupid[["age", "religion"]].head()
# + [markdown] colab_type="text" id="jHFdMuKxHQGh"
# ### 1.3.3 Type Inference and Casting
# + [markdown] colab_type="text" id="rQzO52mU49V5"
#
# `pandas` tries to infer the type of each variable automatically. If every value in a column (except for missing values) is a number, then `pandas` will treat that variable as quantitative. Otherwise, the variable is treated as categorical.
#
# To determine the type that Pandas inferred, simply select that variable using the methods above and look for its `dtype`. A `dtype` of `float64` or `int64` indicates that the variable is quantitative. For example, the `age` variable has a `dtype` of `int64`, so it is quantitative.
# + colab={} colab_type="code" id="ryMiIIc41cYZ"
df_okcupid.age
# + [markdown] colab_type="text" id="Hp9eJzrcHnj8"
# On the other hand, the `religion` variable has a `dtype` of `object`, so `pandas` will treat it as categorical.
# + colab={} colab_type="code" id="G3pTtkAYHeLC"
df_okcupid.religion
# + [markdown] colab_type="text" id="iL7_9NjKJWaC"
# Sometimes it is necessary to convert quantitative variables to categorical variables and vice versa. This can be achieved using the `.astype()` method of a `Series`. For example, to convert `age` to a categorical variable, we simply cast its values to strings.
# + colab={} colab_type="code" id="RYkKhlT8IFlz"
df_okcupid.age.astype(str)
# + [markdown] colab_type="text" id="7PjCiv8KMaep"
# To save this as a column in the `DataFrame`, we assign it to a column called `age_cat`. (Note that this column does not exist yet! It will be created at the time of assignment.)
# + colab={} colab_type="code" id="DYO8Nf7eMaMC"
df_okcupid["age_cat"] = df_okcupid.age.astype(str)
# Check that age_cat is a column in this DataFrame
df_okcupid.head()
# + [markdown] colab_type="text" id="46Toey3KJmkw"
# ### 1.3.4 Exercises
# + [markdown] colab_type="text" id="VPWWaiMkJ8je"
# Exercises 1-2 deal with the Titanic data set `https://dlsun.github.io/pods/data/titanic.csv`
# + [markdown] colab_type="text" id="0pHYmVsPJx5V"
# 1\. Read in the Titanic data set. Identify each variable in the Titanic data set as either quantitative, categorical, or other. Cast all variables to the right type and assign them back to the `DataFrame`.
# + colab={} colab_type="code" id="Ms1lIXsmJY7g"
# YOUR CODE HERE
# + [markdown] colab_type="text" id="yXE6l53nO14r"
# 2\. Create a `DataFrame` (not a `Series`) consisting of just the `class` column.
# + colab={} colab_type="code" id="iqTL7YA2Lz-j"
# YOUR CODE HERE
|
TesterBook/01_Data_Ecosystem/Chapter_1.4_Columns_and_Variables.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Run help
# !python src.py -h
# + pycharm={"name": "#%%\n"}
# Run `create` help
# !python src.py create -h
# + pycharm={"name": "#%%\n"}
# Run `remove` help
# !python src.py remove -h
# + pycharm={"name": "#%%\n"}
# Run `list` help
# !python src.py list -h
# + pycharm={"name": "#%%\n"}
# Run `create` command
# !python src.py create folder1
# + pycharm={"name": "#%%\n"}
# Run `remove` command
# !python src.py remove folder1 folder2
# + pycharm={"name": "#%%\n"}
# Run `list` command
# !python src.py list
|
docs/examples/2-sub-commands/usage.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# # READING THE DATASET
df=pd.read_csv("heart.csv")
# # DATA SUMMARY
df.head()
# +
df.tail()
#age age in years
#sex (1 = male; 0 = female)
# #cp chest pain type
#trestbps resting blood pressure (in mm Hg on admission to the hospital)
#chol serum cholestoral in mg/dl
#fbs (fasting blood sugar > 120 mg/dl) (1 = true; 0 = false)
#restecg resting electrocardiographic results
#thalach maximum heart rate achieved
#exang exercise induced angina (1 = yes; 0 = no)
#oldpeak ST depression induced by exercise relative to rest
#slope the slope of the peak exercise ST segment
#ca number of major vessels (0-3) colored by flourosopy
#thal 3 = normal; 6 = fixed defect; 7 = reversable defect
#target 1 or 0
# -
df.describe()
#checking for null values
df.isnull().sum()
plt.figure(figsize=(20,20))
sns.heatmap(df.corr(),vmax=.3,center=0,square=True,linewidths=.5,cbar_kws={"shrink":.5},annot=True)
plt.tight_layout()
plt.show()
sns.pairplot(df,hue="target")
plt.show()
# # ---------------------------------------------------------------------------------------------------------------
# # K-NN
from sklearn.neighbors import KNeighborsClassifier
# +
from sklearn.model_selection import train_test_split
X=df.iloc[:,:-1]
y=df.iloc[:,-1]
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=50)
print(len(X_train),len(X_test))
print(len(y_train),len(y_test))
# +
def neigh_test(n):
neigh=KNeighborsClassifier(n_neighbors=n)
neigh.fit(X_train,y_train)
return neigh.score(X_test,y_test)
n=[]
acc=[]
for i in range(2,15):
n.append(i)
acc.append(neigh_test(i))
fig=plt.figure(figsize=(10,10))
ax=fig.add_subplot(111)
line,=ax.plot(n,acc)
ymax=max(acc)
xpos=acc.index(ymax)
xmax=n[xpos]
print(ymax,xmax)
ax.annotate('maximum accuracy',xy=(xmax,ymax),xytext=(xmax,ymax),arrowprops=dict(facecolor='black',shrink=0.05),)
plt.plot(n,acc)
plt.title("K-NN")
plt.xlabel("neighbours")
plt.ylabel("accuracy")
plt.show()
# -
# # ---------------------------------------------------------------------------------------------------------------
# # LOGISTIC REGRESSION WITH PCA
from sklearn.linear_model import LogisticRegression
from sklearn.decomposition import PCA
# +
def pca_and_regression(n):
pca=PCA(n_components=n)
Xpca=X
#print(len(Xpca))
pca.fit(Xpca)
#print(pca)
Xpca=pca.fit_transform(Xpca)
#print(Xpca)
Xpca_train,Xpca_test,ypca_train,ypca_test=train_test_split(Xpca,y,test_size=0.3,random_state=50)
#print(Xpca_train)
clf=LogisticRegression(random_state=0,solver='lbfgs')
clf.fit(Xpca_train,ypca_train)
return clf.score(Xpca_test,ypca_test)
n=[i for i in range(1,14)]
acc=[]
for ele in n:
acc.append(pca_and_regression(ele))
fig=plt.figure(figsize=(10,10))
ax=fig.add_subplot(111)
line,=ax.plot(n,acc)
ymax=max(acc)
xpos=acc.index(ymax)
xmax=n[xpos]
print(ymax,xmax)
ax.annotate('maximum accuracy',xy=(xmax,ymax),xytext=(xmax,ymax),arrowprops=dict(facecolor='black',shrink=0.05),)
plt.plot(n,acc)
plt.title("Logistic regression")
plt.xlabel("features")
plt.ylabel("accuracy")
plt.show()
# -
# # ---------------------------------------------------------------------------------------------------------------
# # DECISION TREE
from sklearn import tree
X=df.iloc[:,:-1]
y=df.iloc[:,-1]
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=50)
print(len(X_train),len(X_test))
print(len(y_train),len(y_test))
clf=tree.DecisionTreeClassifier(criterion='entropy')
clf.fit(X_train,y_train)
print("accuracy with decision trees ",clf.score(X_test,y_test))
import graphviz
dot_data = tree.export_graphviz(clf, out_file=None)
graph = graphviz.Source(dot_data)
graph.render("heart")
# +
# dot_data = tree.export_graphviz(clf, out_file=None,
# feature_names=["age","sex","cp","trestbps","chol","fbs","restecg","thalach","exang","oldpeak","slope","ca","thal"],
# class_names=["0","1"],
# filled=True, rounded=True,
# special_characters=True)
# graph = graphviz.Source(dot_data)
# graph
# -
# # ---------------------------------------------------------------------------------------------------------------
# # SVM
# +
from sklearn.model_selection import train_test_split
from sklearn.svm import SVC
import pandas as pd
df=pd.read_csv("heart.csv")
X=df.iloc[:,:-1]
y=df.iloc[:,-1]
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=50)
# +
from sklearn.svm import SVC
clf = SVC(kernel = 'linear',random_state = 0)
clf.fit(X_train, y_train)
y_pred_svm = clf.predict(X_test)
print(y_pred_svm)
print(list(y_test))
print("accuracy ",clf.score(X_test,y_test))
# -
# # ---------------------------------------------------------------------------------------------------------------
#
# # AdaBoost
import pandas as pd
df=pd.read_csv("heart.csv")
X=df.iloc[:,:-1]
y=df.iloc[:,-1]
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=10)
print(len(X_train),len(X_test))
print(len(y_train),len(y_test))
from sklearn.model_selection import cross_val_score
from sklearn.ensemble import AdaBoostClassifier
from sklearn.model_selection import train_test_split
from sklearn import tree
from sklearn import metrics
from sklearn import model_selection
import matplotlib.pyplot as plt
# +
kfold=model_selection.KFold(n_splits=10,random_state=7)
clf=AdaBoostClassifier(tree.DecisionTreeClassifier(criterion='entropy'),n_estimators=10,random_state=2)
clf.fit(X_train,y_train)
result=model_selection.cross_val_score(clf,X_train,y_train,cv=kfold)
print(result)
print(result.mean())
# -
predicted=clf.predict(X_test)
print(metrics.accuracy_score(y_test,predicted))
|
V3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:python35]
# language: python
# name: conda-env-python35-py
# ---
# # This code based on these codes.
# https://www.kaggle.com/francksylla/titanic/titanic-machine-learning-from-disaster/code
#
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import Image, display
# %matplotlib inline
train_input = pd.read_csv("../input/train.csv", dtype={"Age": np.float64})
test_input = pd.read_csv("../input/test.csv", dtype={"Age": np.float64})
df = pd.concat([train_input, test_input], ignore_index=True)
df.head()
# -
print(df.hist())
# +
categorical_columns = ['Sex', 'Embarked']
numerical_columns = ['Pclass', 'Age', 'SibSp', 'Parch', 'Fare']
text_columns = ['Name', 'Ticket']
def category_to_numeric(df, column_name):
for category in df[column_name].unique():
category_column = column_name + '_' + str(category)
if category_column in df.columns:
df = df.drop(category_column, axis=1)
if category_column not in numerical_columns:
numerical_columns.append(category_column)
df= pd.concat([df,pd.get_dummies(df[column_name], prefix=column_name)],axis=1)
return df
# -
print(df.hist())
# +
# Sex
sns.set(style="whitegrid")
g = sns.factorplot(x="Sex", y="Survived", data=df, size=4, palette="muted")
g.despine(left=True)
g.set_ylabels("survival probability")
# +
def get_sex_adult(row):
age, sex = row
if age < 18:
return 'child'
elif sex == 'female':
return 'female_adult'
else:
return 'male_adult'
df['SexAdult'] = df[['Age', 'Sex']].apply(get_sex_adult, axis=1)
g = sns.factorplot(x="SexAdult", y="Survived", data=df, size=4, palette="muted")
if 'SexAdult' not in categorical_columns:
categorical_columns.append('SexAdult')
# +
# Embarked
df['Embarked'] = df['Embarked'].fillna('unknown')
if 'Embarked' not in categorical_columns:
categorical_columns.append('Embarked')
df["Embarked_Category"] = pd.Categorical.from_array(df.Embarked).codes
if 'Embarked_Category' not in categorical_columns:
categorical_columns.append('Embarked_Category')
g = sns.factorplot(x="Embarked_Category", y="Survived", data=df, size=4, palette="muted")
g.despine(left=True)
g.set_ylabels("survival probability")
# +
df_ticket = pd.DataFrame(df['Ticket'].value_counts())
df_ticket.rename(columns={'Ticket':'TicketMembers'}, inplace=True)
df_ticket['Ticket_perishing_women'] = df.Ticket[(df.SexAdult == 'female_adult')
& (df.Survived == 0.0)
& ((df.Parch > 0) | (df.SibSp > 0))].value_counts()
df_ticket['Ticket_perishing_women'] = df_ticket['Ticket_perishing_women'].fillna(0)
df_ticket['TicketGroup_include_perishing_women'] = df_ticket['Ticket_perishing_women'] > 0
df_ticket['TicketGroup_include_perishing_women'] = df_ticket['TicketGroup_include_perishing_women'].astype(int)
df_ticket['Ticket_surviving_men'] = df.Ticket[(df.SexAdult == 'male_adult')
& (df.Survived == 1.0)
& ((df.Parch > 0) | (df.SibSp > 0))].value_counts()
df_ticket['Ticket_surviving_men'] = df_ticket['Ticket_surviving_men'].fillna(0)
df_ticket['TicketGroup_include_surviving_men'] = df_ticket['Ticket_surviving_men'] > 0
df_ticket['TicketGroup_include_surviving_men'] = df_ticket['TicketGroup_include_surviving_men'].astype(int)
df_ticket["TicketId"] = pd.Categorical.from_array(df_ticket.index).codes
df_ticket.loc[df_ticket[df_ticket['TicketMembers'] < 3].index, "TicketId"] = -1
df_ticket["TicketMembers_Simple"] = pd.cut(df_ticket['TicketMembers'], bins=[0,1,4,20], labels=[0,1,2])
if 'TicketGroup_include_perishing_women' not in df.columns:
df = pd.merge(df, df_ticket, left_on="Ticket", right_index=True, how='left', sort=False)
if 'Ticket_perishing_women' not in numerical_columns:
numerical_columns.append('Ticket_perishing_women')
if 'TicketGroup_include_perishing_women' not in numerical_columns:
numerical_columns.append('TicketGroup_include_perishing_women')
if 'Ticket_surviving_men' not in numerical_columns:
numerical_columns.append('Ticket_surviving_men')
if 'TicketGroup_include_surviving_men' not in numerical_columns:
numerical_columns.append('TicketGroup_include_surviving_men')
if 'TicketId' not in numerical_columns:
numerical_columns.append('TicketId')
if 'TicketMembers' not in numerical_columns:
numerical_columns.append('TicketMembers')
g = sns.factorplot(x="TicketGroup_include_perishing_women", y="Survived", data=df, size=4, palette="muted")
g = sns.factorplot(x="Ticket_surviving_men", y="Survived", data=df, size=4, palette="muted")
# +
# surname
df['surname'] = df['Name'].apply(lambda x: x.split(',')[0].lower())
df_surname = pd.DataFrame(df['surname'].value_counts())
df_surname.rename(columns={'surname':'SurnameMembers'}, inplace=True)
df_surname['Surname_perishing_women'] = df.surname[(df.SexAdult == 'female_adult')
& (df.Survived == 0.0)
& ((df.Parch > 0) | (df.SibSp > 0))].value_counts()
df_surname['Surname_perishing_women'] = df_surname['Surname_perishing_women'].fillna(0)
df_surname['SurnameGroup_include_perishing_women'] = df_surname['Surname_perishing_women'] > 0
df_surname['SurnameGroup_include_perishing_women'] = df_surname['SurnameGroup_include_perishing_women'].astype(int)
df_surname['Surname_surviving_men'] = df.surname[(df.SexAdult == 'male_adult')
& (df.Survived == 1.0)
& ((df.Parch > 0) | (df.SibSp > 0))].value_counts()
df_surname['Surname_surviving_men'] = df_surname['Surname_surviving_men'].fillna(0)
df_surname['SurnameGroup_include_surviving_men'] = df_surname['Surname_surviving_men'] > 0
df_surname['SurnameGroup_include_surviving_men'] = df_surname['SurnameGroup_include_surviving_men'].astype(int)
df_surname["SurnameId"] = pd.Categorical.from_array(df_surname.index).codes
df_surname.loc[df_surname[df_surname['SurnameMembers'] < 3].index, "SurnameId"] = -1
df_surname["SurnameMembers_Simple"] = pd.cut(df_surname['SurnameMembers'], bins=[0,1,4,20], labels=[0,1,2])
if 'SurnameGroup_include_perishing_women' not in df.columns:
df = pd.merge(df, df_surname, left_on="surname", right_index=True, how='left', sort=False)
if 'Surname_perishing_women' not in numerical_columns:
numerical_columns.append('Surname_perishing_women')
if 'SurnameGroup_include_perishing_women' not in numerical_columns:
numerical_columns.append('SurnameGroup_include_perishing_women')
if 'Surname_surviving_men' not in numerical_columns:
numerical_columns.append('Surname_surviving_men')
if 'SurnameGroup_include_surviving_men' not in numerical_columns:
numerical_columns.append('SurnameGroup_include_surviving_men')
if 'SurnameId' not in numerical_columns:
numerical_columns.append('SurnameId')
if 'SurnameMembers' not in numerical_columns:
numerical_columns.append('SurnameMembers')
g = sns.factorplot(x="SurnameGroup_include_perishing_women", y="Survived", data=df, size=4, palette="muted")
g = sns.factorplot(x="SurnameGroup_include_surviving_men", y="Survived", data=df, size=4, palette="muted")
# -
# title
import re
df['Name_title'] = df['Name'].apply(lambda x: re.search(' ([A-Za-z]+)\.', x).group(1))
df.loc[df[df['Name_title'] == 'Ms'].index, 'Name_title'] = 'Miss'
print(df['Name_title'].unique())
if 'Name_title' not in categorical_columns:
categorical_columns.append('Name_title')
g = sns.factorplot(y="Name_title", x="Survived", data=df, size=4, palette="muted")
# +
title_mapping = {
"Mr": 1,
"Miss": 2,
"Ms": 2,
"Mlle": 2,
"Mrs": 3,
"Mme": 3,
"Master": 4,
"Dr": 5,
"Rev": 6,
"Major": 7,
"Capt": 7,
"Col": 7,
"Don": 9,
"Dona": 9,
"Sir": 9,
"Lady": 10,
"Countess": 10,
"Jonkheer": 10,
}
df["Name_titleCategory"] = df.loc[:,'Name_title'].map(title_mapping)
if 'Name_titleCategory' not in categorical_columns:
categorical_columns.append('Name_titleCategory')
g = sns.factorplot(x="Name_titleCategory", y="Survived", data=df, size=4, palette="muted")
# -
# FamilySize
df['FamilySize'] = df['SibSp'] + df['Parch']
if 'FamilySize' not in numerical_columns:
numerical_columns.append('FamilySize')
g = sns.factorplot(x="SibSp", y="Survived", data=df, size=4, palette="muted")
g = sns.factorplot(x="Parch", y="Survived", data=df, size=4, palette="muted")
g = sns.factorplot(x="FamilySize", y="Survived", data=df, size=4, palette="muted")
# Name Length?
df['NameLength'] = df["Name"].apply(lambda x: len(x))
if 'NameLength' not in numerical_columns:
numerical_columns.append('NameLength')
g = sns.factorplot(y="NameLength", x="Survived", data=df, size=4, palette="muted")
g.despine(left=True)
g.set_ylabels("survival probability")
# Pclass
g = sns.factorplot(x="Pclass", y="Survived", data=df, size=4, palette="muted")
# +
# cabin
# https://www.kaggle.com/c/titanic/prospector#1326
def get_cabin_location(cabin):
if cabin == ' ':
return 'no_cabin'
# The cabin info consists of a letter (corresponding to a deck)
# and a cabin number, which is odd for cabins on the starboard side and even for the port.
cabin_search_result = re.search('\d+', cabin)
if cabin_search_result:
type_code = np.int64(cabin_search_result.group(0))
if type_code % 2 == 0:
return 'port'
else:
return 'starboard'
return 'unknown'
def get_cabin_deck(cabin):
if cabin == ' ':
return 'no_cabin'
# The cabin info consists of a letter (corresponding to a deck)
# and a cabin number, which is odd for cabins on the starboard side and even for the port.
cabin_search_result = re.search('[A-z]+', cabin)
if cabin_search_result:
return cabin_search_result.group(0)
return 'unknown'
def get_cabin_count(cabin):
if cabin == ' ':
return 0
cabin_search_result = re.findall('([A-z]\d+)', cabin)
if cabin_search_result:
return len(cabin_search_result)
return 0
df['CabinLocation'] = df['Cabin'].fillna(' ').apply(get_cabin_location)
df['CabinDeck'] = df['Cabin'].fillna(' ').apply(get_cabin_deck)
df['CabinCount'] = df['Cabin'].fillna(' ').apply(get_cabin_count)
if 'CabinLocation' not in categorical_columns:
categorical_columns.append('CabinLocation')
if 'CabinDeck' not in categorical_columns:
categorical_columns.append('CabinDeck')
if 'CabinCount' not in numerical_columns:
numerical_columns.append('CabinCount')
g = sns.factorplot(x="Survived", y="CabinLocation", data=df, size=4, palette="muted")
g = sns.factorplot(x="Survived", y="CabinDeck", data=df, size=4, palette="muted")
g = sns.factorplot(x="CabinCount", y="Survived", data=df, size=4, palette="muted")
# -
df['CabinCategory'] = pd.Categorical.from_array(df.Cabin.fillna('0').apply(lambda x:x[0])).codes
g = sns.factorplot(y="Survived", x="CabinCategory", data=df, size=4, palette="muted")
if 'CabinCategory' not in categorical_columns:
categorical_columns.append('CabinCategory')
# Fare
# df['Fare'] = df['Fare'].fillna(df['Fare'].mean())
df["Fare"] = df["Fare"].fillna(8.05)
print(df['Fare'].describe())
print(df['Fare'].hist())
g = sns.factorplot(x="Survived", y="Fare", data=df, size=4, palette="muted")
df['TicketMembers'] = df['TicketMembers'].fillna(0)
print(df.head()[['Pclass','Fare', 'TicketMembers']])
df['Fare_per_ticket_member'] = df['Fare'] / (df['TicketMembers'])
print(df['Fare_per_ticket_member'].hist())
g = sns.factorplot(x="Survived", y="Fare_per_ticket_member", data=df, size=4, palette="muted")
# +
from math import log
class_fare = pd.DataFrame(columns=['count','mean','std','min','25%','50%','75%','max'])
class_fare.loc[1,:] = df[df['Pclass'] == 1]['Fare'].describe()
class_fare.loc[2,:] = df[df['Pclass'] == 2]['Fare'].describe()
class_fare.loc[3,:] = df[df['Pclass'] == 3]['Fare'].describe()
very_small_val = 0.01
df['Fare_standard_score_with_Pclass'] = df.apply(lambda row: (log(row['Fare'] + very_small_val) - log(class_fare.loc[row['Pclass'], 'mean'] + very_small_val)) / log(class_fare.loc[row['Pclass'], 'std'] + very_small_val), axis=1)
if 'Fare_standard_score_with_Pclass' not in numerical_columns:
numerical_columns.append('Fare_standard_score_with_Pclass')
# -
df[(df['Fare_standard_score_with_Pclass'] >= -0.5) & (df['Fare_standard_score_with_Pclass'] <= 0.5)]['Fare_standard_score_with_Pclass'].hist()
g = sns.factorplot(x="Survived", y="Fare_standard_score_with_Pclass", data=df, size=4, palette="muted")
# +
from math import log
class_fare = pd.DataFrame(columns=['count','mean','std','min','25%','50%','75%','max'])
class_fare.loc[1,:] = df[df['Pclass'] == 1]['Fare_per_ticket_member'].describe()
class_fare.loc[2,:] = df[df['Pclass'] == 2]['Fare_per_ticket_member'].describe()
class_fare.loc[3,:] = df[df['Pclass'] == 3]['Fare_per_ticket_member'].describe()
very_small_val = 0.01
df['Fare_per_ticket_member_standard_score_with_Pclass'] = df.apply(lambda row: (log(row['Fare_per_ticket_member'] + very_small_val) - log(class_fare.loc[row['Pclass'], 'mean'] + very_small_val)) / log(class_fare.loc[row['Pclass'], 'std'] + very_small_val), axis=1)
if 'Fare_per_ticket_member_standard_score_with_Pclass' not in numerical_columns:
numerical_columns.append('Fare_per_ticket_member_standard_score_with_Pclass')
# -
df[(df['Fare_per_ticket_member_standard_score_with_Pclass'] >= -0.5) & (df['Fare_per_ticket_member_standard_score_with_Pclass'] <= 0.5)]['Fare_per_ticket_member_standard_score_with_Pclass'].hist()
g = sns.factorplot(x="Survived", y="Fare_per_ticket_member_standard_score_with_Pclass", data=df, size=4, palette="muted")
# +
# https://www.kaggle.com/c/titanic/forums/t/11127/do-ticket-numbers-mean-anything
#print(df["Ticket"])
#print(df["Ticket"].value_counts())
def get_ticket_prefix(cabin):
# The cabin info consists of a letter (corresponding to a deck)
# and a cabin number, which is odd for cabins on the starboard side and even for the port.
cabin_search_result = re.search('[^\d]+', cabin)
if cabin_search_result:
return cabin_search_result.group(0).replace('/', '').replace('.', '').strip()
return 'unknown'
df['TicketPrefix'] = df['Ticket'].apply(get_ticket_prefix)
g = sns.factorplot(y="TicketPrefix", x="Survived", data=df, size=8, palette="muted")
if 'TicketPrefix' not in categorical_columns:
categorical_columns.append('TicketPrefix')
# -
for col in categorical_columns:
df = category_to_numeric(df, col)
# +
# age prediction
from sklearn.ensemble import ExtraTreesRegressor
age_prediction_features = ['Fare', 'Fare_standard_score_with_Pclass',
#'Fare_per_ticket_member', 'Fare_per_ticket_member_standard_score_with_Pclass',
'Parch', 'Pclass', 'SibSp', 'Sex_female', 'Sex_male', 'FamilySize',
'NameLength', 'TicketMembers', 'TicketId',
'Embarked_S', 'Embarked_C', 'Embarked_Q', 'Embarked_unknown',
'Name_title_Mr', 'Name_title_Mrs', 'Name_title_Miss', 'Name_title_Master',
'Name_title_Don', 'Name_title_Rev', 'Name_title_Dr', 'Name_title_Mme',
'Name_title_Major', 'Name_title_Lady', 'Name_title_Sir', 'Name_title_Mlle', 'Name_title_Col',
'Name_title_Capt', 'Name_title_Countess', 'Name_title_Jonkheer',
'CabinLocation_no_cabin', 'CabinLocation_starboard', 'CabinLocation_port', 'CabinDeck_no_cabin',
'CabinDeck_C', 'CabinDeck_E', 'CabinDeck_G', 'CabinDeck_D', 'CabinDeck_A', 'CabinDeck_B', 'CabinDeck_F', 'CabinDeck_T'
]
age_prediction_tree_regressor = ExtraTreesRegressor(n_estimators=200)
age_X_train = df[age_prediction_features][df['Age'].notnull()]
age_Y_train = df['Age'][df['Age'].notnull()]
age_prediction_tree_regressor.fit(age_X_train, np.ravel(age_Y_train))
# predict only isnull values
df['Age_pred'] = df['Age']
df.loc[df[df['Age'].isnull()].index, 'Age_pred'] = age_prediction_tree_regressor.predict(df[age_prediction_features][df['Age'].isnull()])
if 'Age_pred' not in numerical_columns:
numerical_columns.append('Age_pred')
# add ageGroup
df["AgeGroup"] = pd.cut(df['Age'], bins=[-2000,0,11,15,18,30,49,59,200], labels=[-1, 11,15,18,30,49,59,200])
df["AgeGroup_pred"] = pd.cut(df['Age_pred'], bins=[-2000,11,15,18,30,49,59,200], labels=[11,15,18,30,49,59,200])
if 'AgeGroup' not in numerical_columns:
numerical_columns.append('AgeGroup')
if 'AgeGroup_pred' not in numerical_columns:
numerical_columns.append('AgeGroup_pred')
g = sns.factorplot(y="Survived", x="AgeGroup", data=df, size=4, palette="muted")
g = sns.factorplot(y="Survived", x="AgeGroup_pred", data=df, size=4, palette="muted")
# -
# Frugal_First_Class_Single_Man
# midle age first class single man with large discounted and unknown prefixed ticket and without cabin.
print("died", df[(df['Survived'] == 0) & (df['Sex'] == 'male')
& (df['Pclass'] == 1)
& (df['Age_pred'] <= 45)
& (df['Fare'] > 0)
& (df['Fare_standard_score_with_Pclass'] < -0.25)
& (df['TicketPrefix_unknown'] == 1)
& (df['TicketMembers_Simple'] == 0)
& (df['CabinCount'] == 0)
])
print("survived", df[(df['Survived'] == 1) & (df['Sex'] == 'male')
& (df['Pclass'] == 1)
& (df['Age_pred'] <= 45)
& (df['Fare'] > 0)
& (df['Fare_standard_score_with_Pclass'] < -0.25)
& (df['TicketPrefix_unknown'] == 1)
& (df['TicketMembers_Simple'] == 0)
& (df['CabinCount'] == 0)
])
# +
df['Frugal_First_Class_Single_Man'] = 0
df.loc[df[(df['Sex'] == 'male')
& (df['CabinCount'] > 0)
& (df['Embarked'] == 'C')
& (df['SurnameMembers'] == 1)
& (df['TicketPrefix_unknown'] == 1.0)
& (df['Fare_standard_score_with_Pclass'] < -0.23)
& (df['Pclass'] == 1)]['Frugal_First_Class_Single_Man'].index, 'Frugal_First_Class_Single_Man'] = 1
display(df[(df['Frugal_First_Class_Single_Man'] == 1)])
if 'Frugal_First_Class_Single_Man' not in numerical_columns:
numerical_columns.append('Frugal_First_Class_Single_Man')
# +
display(df[(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age_pred'] > 30) &
(df['Pclass'] == 3) &
(df['Name_title_Miss'] == 1.0)
])
# poor old miss
df['Poor_Old_Miss_Third_Class'] = 0
df.loc[df[(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age'] > 30) &
(df['Pclass'] == 3) &
(df['Name_title_Miss'] == 1.0)].index, 'Poor_Old_Miss_Third_Class'] = 1
if 'Poor_Old_Miss_Third_Class' not in numerical_columns:
numerical_columns.append('Poor_Old_Miss_Third_Class')
# +
display(df[(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age_pred'] >= 38) &
(df['Pclass'] == 2) &
(df['Name_title_Miss'] == 1.0) &
(df['TicketPrefix_unknown'] == 1.0) &
(df['SurnameMembers_Simple'] == 0)
])
# poor old miss
df['Poor_Old_Miss_Second_Class'] = 0
df.loc[df[
(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age_pred'] >= 38) &
(df['Pclass'] == 2) &
(df['Name_title_Miss'] == 1.0) &
(df['TicketPrefix_unknown'] == 1.0) &
(df['SurnameMembers_Simple'] == 0)
].index, 'Poor_Old_Miss_Second_Class'] = 1
if 'Poor_Old_Miss_Second_Class' not in numerical_columns:
numerical_columns.append('Poor_Old_Miss_Second_Class')
# +
display(df[
(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age_pred'] >= 35) &
(df['Pclass'] == 1) &
(df['Name_title_Miss'] == 1.0) &
(df['SurnameMembers_Simple'] == 0)
])
# poor old miss
df['Poor_Old_Miss_First_Class'] = 0
df.loc[df[
(df['Sex'] == 'female') &
(df['Fare_standard_score_with_Pclass'] <= -0.18) &
(df['Age_pred'] >= 35) &
(df['Pclass'] == 1) &
(df['Name_title_Miss'] == 1.0) &
(df['SurnameMembers_Simple'] == 0)
].index, 'Poor_Old_Miss_First_Class'] = 1
if 'Poor_Old_Miss_First_Class' not in numerical_columns:
numerical_columns.append('Poor_Old_Miss_First_Class')
# +
df[(df['Sex'] == 'female') & (df['Fare'] <= 10) & (df['Age'] > 28) & (df['Name_title_Miss'] == 1.0)]
# poor old miss
df['Poor_Old_Miss'] = 0
df.loc[df[(df['Sex'] == 'female')
& (df['Fare'] <= 10)
& (df['Age_pred'] > 28)
& (df['Name_title_Miss'] == 1.0)].index, 'Poor_Old_Miss'] = 1
if 'Poor_Old_Miss' not in numerical_columns:
numerical_columns.append('Poor_Old_Miss')
# +
df[(df['Sex'] == 'female') & (df['Fare'] <= 10) & (df['Age'] > 26) & (df['Embarked'] == 'S') & (df['Name_title_Miss'] == 1.0)]
# poor Shouthampton old miss
df['Poor_Shouthampton_Old_Miss'] = 0
df.loc[df[(df['Sex'] == 'female')
& (df['Fare'] <= 10)
& (df['Age_pred'] > 26)
& (df['Embarked'] == 'S')
& (df['Name_title_Miss'] == 1.0)].index, 'Poor_Shouthampton_Old_Miss'] = 1
if 'Poor_Shouthampton_Old_Miss' not in numerical_columns:
numerical_columns.append('Poor_Shouthampton_Old_Miss')
# +
# feature selection
from sklearn.feature_selection import SelectKBest, f_classif
df_copied = df.copy()
df_copied['Name_titleCategory'] = df_copied['Name_titleCategory'].fillna(' ')
df_copied['Cabin'] = df_copied['Cabin'].fillna(' ')
df_copied['Age'] = df_copied['Age'].fillna(-300)
df_copied['AgeGroup'] = df_copied['AgeGroup'].fillna(-1.0)
train = df_copied[0:891].copy()
target = train["Survived"].values
selector = SelectKBest(f_classif, k=len(numerical_columns))
selector.fit(train[numerical_columns], target)
scores = -np.log10(selector.pvalues_)
indices = np.argsort(scores)[::-1]
print("Features importance :")
for f in range(len(scores)):
print("%0.2f %s" % (scores[indices[f]],numerical_columns[indices[f]]))
# +
# Random Forest
from sklearn.ensemble import RandomForestClassifier
from sklearn import cross_validation
random_forest = RandomForestClassifier(n_estimators=3000, min_samples_split=4, class_weight={0:0.745, 1:0.255})
kfold = cross_validation.KFold(train.shape[0], n_folds=3, random_state=42)
scores = cross_validation.cross_val_score(random_forest, train[numerical_columns], target, cv=kfold)
print("Accuracy: %0.3f (+/- %0.2f) [%s]" % (scores.mean() * 100, scores.std() * 100, 'Random Forest Cross Validation'))
random_forest.fit(train[numerical_columns], target)
score = random_forest.score(train[numerical_columns], target)
print("Accuracy: %0.3f [%s]" % (score * 100, 'Random Forest full test'))
importances = random_forest.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(len(numerical_columns)):
print("%d. feature %d (%f) %s" % (f + 1, indices[f] + 1, importances[indices[f]] * 100, numerical_columns[indices[f]]))
# +
import matplotlib.pyplot as plt
from collections import OrderedDict
from sklearn.datasets import make_classification
from sklearn.ensemble import RandomForestClassifier, ExtraTreesClassifier
ensemble_clfs = [
("RandomForestClassifier",
RandomForestClassifier(
n_estimators=3000,
n_jobs=8,
class_weight={0:0.745, 1:0.255},
min_samples_split=4,
random_state=42)),
]
kfold = cross_validation.KFold(train.shape[0], n_folds=3, random_state=42)
# +
# error_rate = OrderedDict((label, []) for label, _ in ensemble_clfs)
# for label, clf in ensemble_clfs:
# print("Classifier : %s" % label)
# for i in range(2, 10):
# clf.set_params(min_samples_split=i)
# clf.fit(train[numerical_columns], target)
# scores = cross_validation.cross_val_score(clf, train[numerical_columns], target, cv=kfold)
# error_rate[label].append((i, scores.mean() * 100))
# print("%d estimator" % i)
# for label, clf_err in error_rate.items():
# xs, ys = zip(*clf_err)
# plt.plot(xs, ys, label=label)
# plt.xlim(2, 10)
# plt.xlabel("min_samples_split")
# plt.ylabel("score")
# plt.legend(loc="upper right")
# plt.show()
# +
# error_rate = OrderedDict((label, []) for label, _ in ensemble_clfs)
# for label, clf in ensemble_clfs:
# print("Classifier : %s" % label)
# for i in range(1, 100, 10):
# clf.set_params(max_depth=i)
# clf.fit(train[numerical_columns], target)
# scores = cross_validation.cross_val_score(clf, train[numerical_columns], target, cv=kfold)
# # error_rate[label].append((i, scores.mean() * 100))
# print("max_depth : %d" % i)
# for label, clf_err in error_rate.items():
# xs, ys = zip(*clf_err)
# plt.plot(xs, ys, label=label)
# plt.xlim(0, 100)
# plt.xlabel("max_depth")
# plt.ylabel("score")
# plt.legend(loc="upper right")
# plt.show()
# +
features = [
'Sex_female','Sex_male',
'Age_pred',
'SexAdult_male_adult','SexAdult_female_adult', 'SexAdult_child',
'Name_titleCategory',
# 'Name_titleCategory_1',
# 'Name_titleCategory_2',
# 'Name_titleCategory_3',
# 'Name_titleCategory_4',
# 'Name_titleCategory_5',
# 'Name_titleCategory_6',
# 'Name_titleCategory_7',
# 'Name_titleCategory_9',
# 'Name_titleCategory_10',
# 'Name_title_Mr', 'Name_title_Mrs', 'Name_title_Miss', 'Name_title_Master',
# 'Name_title_Don', 'Name_title_Rev', 'Name_title_Dr', 'Name_title_Mme',
# 'Name_title_Major', 'Name_title_Lady', 'Name_title_Sir', 'Name_title_Mlle', 'Name_title_Col',
# 'Name_title_Capt', 'Name_title_Countess', 'Name_title_Jonkheer',
'Pclass',
'TicketId',
'NameLength',
'CabinLocation_no_cabin', 'CabinLocation_starboard', 'CabinLocation_port',
'CabinCategory',
# 'CabinCategory_0',
# 'CabinCategory_1',
# 'CabinCategory_2',
# 'CabinCategory_3',
# 'CabinCategory_4',
# 'CabinCategory_5',
# 'CabinCategory_6',
# 'CabinCategory_7',
# 'CabinCategory_8',
# 'CabinDeck_C', 'CabinDeck_E', 'CabinDeck_G', 'CabinDeck_D', 'CabinDeck_A', 'CabinDeck_B', 'CabinDeck_F', 'CabinDeck_T','CabinDeck_no_cabin',
'SibSp','Parch',
'Fare',
# 'Fare_per_ticket_member',
# 'Fare_standard_score_with_Pclass',
# 'Fare_per_ticket_member_standard_score_with_Pclass',
'Embarked_Category',
# 'Embarked_S','Embarked_Q','Embarked_C','Embarked_unknown',
'SurnameMembers_Simple','SurnameGroup_include_perishing_women','SurnameGroup_include_surviving_men',
'TicketMembers_Simple', 'TicketGroup_include_perishing_women','TicketGroup_include_surviving_men',
'FamilySize',
# 'Frugal_First_Class_Single_Man',
# 'Poor_Old_Miss',
# 'Poor_Shouthampton_Old_Miss',
# 'Poor_Old_Miss_Third_Class',
# 'Poor_Old_Miss_Second_Class',
# 'Poor_Old_Miss_First_Class',
# 'TicketPrefix_SOPP', 'TicketPrefix_WC',
# 'TicketPrefix_unknown',
# 'TicketPrefix_SCA','TicketPrefix_SP','TicketPrefix_SOP','TicketPrefix_Fa','TicketPrefix_SCOW','TicketPrefix_AS',
# 'TicketPrefix_FC','TicketPrefix_SOTONO','TicketPrefix_CASOTON','TicketPrefix_SWPP','TicketPrefix_SC','TicketPrefix_SCAH Basle',
# 'CabinCount',
]
# +
# analyze failed.
X_train, X_test, y_train, y_test = cross_validation.train_test_split(train, target, test_size=0.2, random_state=42)
random_forest = RandomForestClassifier(n_estimators=3000, min_samples_split=4, class_weight={0:0.745, 1:0.255})
kfold = cross_validation.KFold(X_train.shape[0], n_folds=3, random_state=42)
scores = cross_validation.cross_val_score(random_forest, X_train[features], y_train, cv=kfold)
print("Accuracy: %0.3f (+/- %0.2f) [%s]" % (scores.mean() * 100, scores.std() * 100, 'Random Forest Cross Validation'))
random_forest.fit(X_train[features], y_train)
score = random_forest.score(X_test[features], y_test)
print("Accuracy: %0.3f [%s]" % (score * 100, 'Random Forest full test'))
pred_test = random_forest.predict(X_test[features])
importances = random_forest.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(len(features)):
print("%d. feature %d (%f) %s" % (f + 1, indices[f] + 1, importances[indices[f]] * 100, features[indices[f]]))
# -
pd.set_option("display.max_columns",101)
X_test_reseted = X_test.reset_index()
X_test_reseted['Survived_'] = y_test
X_test_reseted['Prediction'] = pred_test
X_test_reseted['pred_result'] = pred_test == y_test
display(X_test_reseted[(X_test_reseted['Survived'] == 1.0) & (X_test_reseted['pred_result'] == False)])
display(X_test_reseted[(X_test_reseted['Survived'] == 0.0) & (X_test_reseted['pred_result'] == False)])
# +
# select specidic features
random_forest = RandomForestClassifier(n_estimators=3000, min_samples_split=4, class_weight={0:0.745, 1:0.255})
kfold = cross_validation.KFold(train.shape[0], n_folds=3, random_state=42)
scores = cross_validation.cross_val_score(random_forest, train[features], target, cv=kfold)
print("Accuracy: %0.3f (+/- %0.2f) [%s]" % (scores.mean() * 100, scores.std() * 100, 'Random Forest Cross Validation'))
random_forest.fit(train[features], target)
score = random_forest.score(train[features], target)
print("Accuracy: %0.3f [%s]" % (score * 100, 'Random Forest full test'))
importances = random_forest.feature_importances_
indices = np.argsort(importances)[::-1]
for f in range(len(features)):
print("%d. feature %d (%f) %s" % (f + 1, indices[f] + 1, importances[indices[f]] * 100, features[indices[f]]))
# -
random_forest = RandomForestClassifier(n_estimators=3000, min_samples_split=4, class_weight={0:0.745, 1:0.255})
test = df_copied[891:].copy()
random_forest.fit(train[features], target)
predictions = random_forest.predict(test[features])
PassengerId = np.array(test["PassengerId"]).astype(int)
submit_df = pd.DataFrame(predictions, PassengerId, columns = ['Survived']).astype(int)
submit_df.to_csv('titanic.csv', index_label=['PassengerId'])
wnot_subimt_df = pd.read_csv("wnot_submit.csv")
wnot_subimt_df = wnot_subimt_df.reset_index().drop('index', axis=1)
wnot_subimt_df = wnot_subimt_df.set_index('PassengerId')
diff = submit_df.copy()
diff['Survived_wnot'] = wnot_subimt_df['Survived']
diff['pred_result'] = diff['Survived_wnot'] == diff['Survived']
display(df_copied.loc[diff[(diff['pred_result'] == False)].index - 1, :])
print(diff[(diff['pred_result'] == False)])
|
titanic/notebook/titanic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
# I've noticed that the biomass reaction in the model doesn't match what is present in Beata's thesis. Tables 4.1, 4.2, 4.3, 4.4, 4.5 and 4.6 cover results of experimental determination of the biomass components. The biomass reaction should encompass all of these metabolites. I have noticed that quite some metabolites are missing, or that the coefficients do not exactly match the presented data. So here I will try to make the reaction match the data presented. I will go per category, and after adding each missing metabolite check biomass formation is still possible. If not, I will try to find out the root cause and tackle that problem.
#
# All the way at the end, when the model is mass and stoiciometrically balanced we can fit the GAM to the data presented in the thesis; for now I will leave it as it is.
import cameo
import pandas as pd
import cobra.io
import escher
from escher import Builder
from cobra import Metabolite, Reaction
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
model_e_coli = cameo.load_model ('iML1515')
model_b_sub = cameo.load_model ('iYO844')
#available at: https://github.com/SysBioChalmers/yeast-GEM/blob/master/ModelFiles/xml/yeastGEM.xml
model_yeast = cobra.io.read_sbml_model('../../Databases/yeastGEM.xml')
model.optimize().objective_value
# ## Ions (table 4.1)
# Table 4.1 shows the determination of the following ions: K, Mg, Fe3+, Ca, phosphate and diphosphate. Note that the units of the metabolites added should be mmol/gdcw.
#
# Our reaction currently has the ions: fe3, fe2 and cl. I don't know why or where they were added, but I will remove them. Here I will try to stringently match the data presented.
#
# Note: phosphate is also formed from the hydrolysis of ATP, so this number will need to change when the GAM is estimated.
#Potassium
model.reactions.biomass.add_metabolites({
model.metabolites.k_c: -0.7082
})
#we also supply potassium in the minimal medium, so add the exchange
model.add_boundary(model.metabolites.k_e, type= 'exchange',reaction_id = 'EX_k_e',lb = -1000, ub = 1000)
#magnesium needs to be added to the model first
model.add_metabolites(Metabolite(id='mg2_c'))
model.add_metabolites(Metabolite(id='mg2_e'))
model.metabolites.mg2_c.name = 'Magnesium'
model.metabolites.mg2_c.compartment= 'c'
model.metabolites.mg2_c.formula = 'Mg'
model.metabolites.mg2_c.annotation = model_e_coli.metabolites.mg2_c.annotation
model.metabolites.mg2_c.charge = 2
model.metabolites.mg2_e.name = 'Magnesium'
model.metabolites.mg2_e.compartment= 'e'
model.metabolites.mg2_e.formula = 'Mg'
model.metabolites.mg2_e.annotation = model_e_coli.metabolites.mg2_c.annotation
model.metabolites.mg2_e.charge = 2
#the mg channel has been found in geobacillus, so will add passive transport.
model.add_reaction(Reaction(id='MG2t'))
model.reactions.MG2t.name = 'Magnesium transport via channel'
model.reactions.MG2t.annotation['sbo'] = 'SBO:0000185'
model.reactions.MG2t.bounds = (-1000,1000)
model.reactions.MG2t.add_metabolites({model.metabolites.mg2_c: -1, model.metabolites.mg2_e: 1})
#add exchange
model.add_boundary(model.metabolites.mg2_e, type= 'exchange',reaction_id = 'EX_mg2_e',lb = -1000, ub = 1000)
#add mg to biomass
model.reactions.biomass.add_metabolites({
model.metabolites.mg2_c: -0.098
})
#Fe3+ is already in the model, just need to change unit
model.reactions.biomass.add_metabolites({
model.metabolites.fe3_c:-0.0041
})
#calcium needs to be added to the model first
model.add_metabolites(Metabolite(id='ca2_c'))
model.add_metabolites(Metabolite(id='ca2_e'))
model.metabolites.ca2_c.name = 'Calcium'
model.metabolites.ca2_c.compartment= 'c'
model.metabolites.ca2_c.formula = 'Ca'
model.metabolites.ca2_c.annotation = model_e_coli.metabolites.ca2_c.annotation
model.metabolites.ca2_c.charge = 2
model.metabolites.ca2_e.name = 'Calcium'
model.metabolites.ca2_e.compartment= 'e'
model.metabolites.ca2_e.formula = 'Ca'
model.metabolites.ca2_e.annotation = model_e_coli.metabolites.ca2_e.annotation
model.metabolites.ca2_e.charge = 2
#there is a calcium antiporter annotated in the genome
model.add_reaction(Reaction(id='CA2t'))
model.reactions.CA2t.name = 'Transport of calcium via antiport'
model.reactions.CA2t.annotation['sbo'] = 'SBO:0000185'
model.reactions.CA2t.bounds = (-1000,1000)
model.reactions.CA2t.add_metabolites({model.metabolites.ca2_c: -1, model.metabolites.ca2_e:1, model.metabolites.h_e: -1, model.metabolites.h_c:1})
#add exchange
model.add_boundary(model.metabolites.ca2_e, type= 'exchange',reaction_id = 'EX_ca2_e',lb = -1000, ub = 1000)
#add ca to biomass
model.reactions.biomass.add_metabolites({
model.metabolites.ca2_c: -0.00315
})
# __phosphate__ is orginally in the biomass reaction already: 104.9856 (coupled to the amount of ATP required for growth.
# As a substrate that is consumed, phosphate only needs 0,0173 mmol/gCDW. This will play only a small difference on the total reaction. I will not change this yet, but this should be modified when we fit the GAM.
#diphosphate
model.reactions.biomass.add_metabolites({model.metabolites.ppi_c:-0.0012})
#remove fe2 and cl from the biomass reaction
model.reactions.biomass.add_metabolites({model.metabolites.fe2_c:0.029903042, model.metabolites.cl_c:0.029903042})
#save & commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
model.optimize().objective_value
# ## Amino acids (table 4.2)
# I've already ensured all amino acids are present in the model. However, the coefficients associated to a majority of them do not fit what is included in table 4.2, so I will modify those where necessary to match the table.
model.reactions.biomass.add_metabolites({
model.metabolites.gly_c:-0.02407042,
model.metabolites.ala__L_c:-0.0303495,
model.metabolites.val__L_c:-0.02220057,
model.metabolites.leu__L_c:-0.02051081,
model.metabolites.ile__L_c:-0.0161025,
model.metabolites.ser__L_c:-0.01068746,
model.metabolites.thr__L_c:-0.0158668,
model.metabolites.phe__L_c:-0.00856894,
model.metabolites.tyr__L_c:0.060075203,
model.metabolites.trp__L_c:-0.00605801,
model.metabolites.cys__L_c:-0.00636758,
model.metabolites.met__L_c:-0.00301383,
model.metabolites.lys__L_c:-0.018717862,
model.metabolites.arg__L_c:-0.01105,
model.metabolites.his__L_c:-0.00459031,
model.metabolites.asp__L_c:-0.01310304,
model.metabolites.glu__L_c:-0.250734713,
model.metabolites.pro__L_c: -0.00983182,
model.metabolites.gln__L_c:0.322355,
model.metabolites.asn__L_c:0.18114312
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
model.optimize().objective_value
# ## DNA & RNA (table 4.3)
# Here table 4.3 estimates values around 0.2-0.3 per nucleotide, or even 8 mmol/gcdw for UMP and CMP. This is a factor 10 higher than the information about the DNA in figure 4.6. This makes it hard to decide which value to use, though for DNA I would expect values near 0.02 mmol/gCDW, and for RNA around 0.15 mm/gDCW.
#
# As the data is hard to really interpret and the outcome of it seems strange. E.g. From the DNA distribution in fig 4.6 we would have 75.4% GC content in the strain, but in the genome we know this should be around 43% (allowing some experimental error).
#
# Niko mentioned that we could alternatively estimate the DNA and RNA composition by knowing the GC-content of the genome and the total amounts of DNA and RNA in our bacteria. So instead I will estimate the values from this.
#
# GC-content: approx 43.4 mol%
# total DNA (1%) i.e. 1g DNA / 100 gDCW
# total RNA (16% of biomass) i.e. 16 g / 100 gDCQ
#
# From this data, together with the MW of each (d)NMP we can estimate the mmol/gDCW of each nucleotide present. see '../databases/Biomass_rct.xlsx' for the calculations and desired final values.
#
# I will correct the values in the model here.
model=cobra.io.read_sbml_model('../model/p-thermo.xml')
model.reactions.biomass.add_metabolites({
model.metabolites.damp_c:0.021231612,
model.metabolites.dcmp_c: 0.016684519,
model.metabolites.dtmp_c: 0.021231612,
model.metabolites.dgmp_c: 0.016684519,
model.metabolites.amp_c:-0.020399789,
model.metabolites.gmp_c: 0.043519524,
model.metabolites.cmp_c:-0.022435461,
model.metabolites.ump_c:-0.046781783,
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# ## Carbohydrates
# Beata's thesis identifies that 10% of DCW is composed of carbohydrates. One would also expect these to be present in the biomass. In the thesis, the data shows the carbohydrate component as split into basic sugars which is convenient from a modelling perspective. It may influence the energy requirement but this will be fit in the GAM anyway.
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
model.reactions.biomass.add_metabolites({
model.metabolites.arab__L_c: -0.045625,
model.metabolites.gal_c:-0.212236,
model.metabolites.glc__D_c:-0.010725,
model.metabolites.xyl__D_c: -0.204370,
model.metabolites.man_c:-0.006411,
model.metabolites.fru_c:-0.117367
})
# Adding the carbohydrates kills the biomass accumulation: so somehow we cannot produce one (or more) of the carbohydrates.
#
# The problematic carbohydrates are: arabinose, galactose, xylose and mannose. Here I will go through their biosynthesis one by one to ensure they can be made and fix this issue.
#
# __Arabinose__
# in the model, the arabinose can be made from ribulose. However ribulose cannot be made, so I will fix this. This is because the RBK_L1 reaction is irreversible as it consumes ATP. The options we have hear are either to make this reaction reversible and risk it results in the formation of ATP. Or we add a hydrolysis reaction that hydrolysis the Ru5p__L to rbl, which is used for biomass.
#
# I think this second option is better. We prevent extensive arabinose production for ATP production purposes in this way. Also the different in ATP requirement for the arabinose needed in growth will be captured in the fitted GAM anyway. The hydrolysis reaction will also be irreversible. By both being irreversible it will prevent a cycle as ATP consumption is coupled to one reaction.
model.add_reaction(Reaction(id='RU5PHY'))
model.reactions.RU5PHY.name = 'Hydrolysis of L-Ribulose-5-phosphate'
model.reactions.RU5PHY.notes['NOTES'] = 'Added reaction to allow biomass formation. ATP is captured in GAM'
model.reactions.RU5PHY.annotation['sbo'] = 'SBO:0000176'
model.reactions.RU5PHY.add_metabolites({
model.metabolites.ru5p__L_c: -1,
model.metabolites.rbl__L_c: 1,
model.metabolites.h2o_c: -1,
model.metabolites.pi_c:1,
model.metabolites.h_c: 1
})
# __Galactose__ Here the problem is the conversion from gal1p to galatose, captured in the GALKr reaction. In E. coli this reaction is reversible, even though it is associated with atp. Based on thermodynamic prediction, this reaction can be reversible still. of course SLP is possible, we just need to be sure that there is not a surplus of energy created by this reaction. As galactose is not exported out of the cell, I doubt this would not happen and so we will make the reaction reversible again.
model.reactions.GALKr.bounds=(-1000,1000)
model.reactions.MELIBHY.id = 'GALS3'
# __Xylose__ Here again the problem is the formation of xylulose from xylulose-5-phosphate. here based on thermodynamics I would not expect a reversible reaction. So instead, we again will add a hydrolysis of xu5p__D_c reaction to allow biomass consumption. Again the possible difference in ATP needed to make biomass will be reflected in the biomass GAM and so is not an issue here.
model.add_reaction(Reaction(id='XU5PHY'))
model.reactions.XU5PHY.name = 'Hydrolysis of L-Xylulose-5-phosphate'
model.reactions.XU5PHY.notes['NOTES'] = 'Added reaction to allow biomass formation. ATP is captured in GAM'
model.reactions.XU5PHY.annotation['sbo'] = 'SBO:0000176'
model.reactions.XU5PHY.add_metabolites({
model.metabolites.xu5p__D_c: -1,
model.metabolites.xylu__D_c: 1,
model.metabolites.h2o_c: -1,
model.metabolites.pi_c:1,
model.metabolites.h_c: 1
})
# __Mannose__ Here we are again missing a reaction that can convert the mannose-6-phosphate back into mannose. The reaction going to man6p is still mass imbalanced and will be fixed later. Based on the fix for that, it is unreasonable to expect a reverse reaction here, and so I will add a hydrolysis reaction again. (Note: same is done in the E. coli model here.)
#
model.add_reaction(Reaction(id='M6PHY'))
model.reactions.M6PHY.name = 'Hydrolysis of Mannose-6-phosphate'
model.reactions.M6PHY.notes['NOTES'] = 'Added reaction to allow biomass formation. ATP is captured in GAM'
model.reactions.M6PHY.annotation['sbo'] = 'SBO:0000176'
model.reactions.M6PHY.add_metabolites({
model.metabolites.man6p_c: -1,
model.metabolites.man_c: 1,
model.metabolites.h2o_c: -1,
model.metabolites.pi_c:1,
model.metabolites.h_c: 1
})
model.reactions.M6PHY.check_mass_balance()
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# # Lipids
# To include the lipids in the biomass equation, we need to do a significant amount of additional work. We need to first decide which approach we will use to capture lipid metabolism and then update the model to account for the production of all components we need for that approach. Therefore this is a much larger issue in itself, and so will be tackled in another notebook.
# # Cofactors
# According to Beata's thesis, there should also be NAD, NADP and NADPH included in the model, as well as quinol. For these cofactors there are experimental values available and so this should be adapted in the biomass reaction.
#
# In the model currently, we also have ribflv, coA and fad present. It could be expected that these are required in the formation of biomass. However their stoichiometric coefficients are very high. After meeting with Ben, we discussed a way to get more realistic coefficients for these components: looking into the yeast, e. coli and b. sub models for the values they have and using those as quidelines. Of course, ideally one would have experimental data on this, however this is unavailable, so we can approach it like this. As these numbers are very small and play only a slight role in biomass formation, slight deviations will not show a huge effect and so it is oke to include it like this.
#
# Please se '../databases/Biomass_rct.xlsx' for a summary of the values used for all cofactors.
# For NADH, FAD, ribflav and CoA I will use the values of the e. coli model. There is still quite some variance that is observed bbetween the e. coli and yeast model. But the values are often of similar magnitude. As our organism is a bacteria, I will use the e. coli values.
#
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
model.reactions.biomass.add_metabolites({
model.metabolites.nad_c: 0.013703042,
model.metabolites.nadh_c: -0.000045,
model.metabolites.nadp_c: 0.025203042,
model.metabolites.nadph_c: 0.026903042,
model.metabolites.fad_c: 0.029680042,
model.metabolites.qh2_c: 0.029603042,
model.metabolites.ribflv_c: 0.029680042,
model.metabolites.coa_c: 0.029735042
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# # Other
# Aside from the above categories, there are also some other metabolites that should be added or looked into. In Beata's thesis there is data about 10-Formyltetrahydrofolate presence that is currently lacking in the biomass reaction. Here i will add this back in.
#
# Also for the accoa_c, succoa_c, chor_c, thmpp_c, gthrd_c and ptrc_c metabolites I will check the stoichiometry that is associated to them by comparing it to the other models that are present and adapt them accordingly as they seem way too high for our model currently.
#
# Again, here I will use the E. coli data as a guide, as those numbers seem more realistic than what has been added into our model.
model=cobra.io.read_sbml_model('../model/p-thermo.xml')
model.reactions.biomass.add_metabolites({
model.metabolites.get_by_id('10fthf_c'): -0.0004,
model.metabolites.accoa_c:0.029624042,
model.metabolites.chor_c:0.029680042,
model.metabolites.gthrd_c:0.029680042,
model.metabolites.ptrc_c: -0.003366958,
model.metabolites.succoa_c:0.03,
model.metabolites.thmpp_c:0.030126042
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# By adding 10fthf_c, the model no longer grows. So now I will check how to resolve this bloackage here. If you supply folate to the model, it grows again so the issue lies somewhere in the folate biosynthesis pathway.
#
# Folate can be made from chorismate or GTP. Supplying either of these doesn't rescue biomass production and so we need to go through both pathways completely to make sure they are correct.
# Aside from some reactions that are reverisible that shouldn't be, the part leading to 4-aminobenzoate seems to be correct. So it seems it must be further upstream of the 6hmhpt_c metabolite.
#
# Also the pathway from GTP Seems to be correct, so there must be some co-factor or side meetabolite that is the issue blocking this pathway.
model.reactions.DHFR.bounds = (-1000,0)
model.reactions.HMHPTMT.bounds = (0,1000)
# Problem is the conversion to form 6hmhpt_c, as supplying dhnpt_c doesnt rescue biomass formation. In the DHNPA2r reaction, the only other involved metabolite is glycoaldehyde. So possibly this cannot be converted sufficiently and so blocks the pathway. Adding the removal of glycoalehyde restores biomass so this is the issue we face currently.
# I will check the glycoaldehyde consumption here.
#seems to miss the GCALDD reaction
model.add_reaction(Reaction(id='GCALDD'))
model.reactions.GCALDD.name = 'Glycolaldehyde dehydrogenase'
model.reactions.GCALDD.annotation = model_e_coli.reactions.GCALDD.annotation
model.reactions.GCALDD.add_metabolites({
model.metabolites.gcald_c:-1,
model.metabolites.h2o_c: -1,
model.metabolites.nad_c:-1,
model.metabolites.glyclt_c:1,
model.metabolites.h_c: 2,
model.metabolites.nadh_c:1
})
#GLYCLTDx and GLYCNOR are duplicate reactions
model.remove_reactions(model.reactions.GLYCNOR)
# +
#problem is: glx can only be converted back into glyclt.
# -
#MALS in the glyoxylate cycle is in the wrong direction!
model.reactions.MALS.bounds=(-1000,0)
# The Bacillus genome encodes a Glyoxalate carboligase. A tBlastn search shows that there is a significant hit in our strain. For that reason, it can be expected that this enzyme is also present in our strain. Hence I will add the reaction here.
model.add_reaction(Reaction(id='GLXCBL'))
model.reactions.GLXCBL.name= 'Glyoxalate carboligase'
model.reactions.GLXCBL.annotation = model_b_sub.reactions.GLXCBL.annotation
model.reactions.GLXCBL.add_metabolites({
model.metabolites.glx_c: -2,
model.metabolites.h_c: -1,
model.metabolites.hop_c:1,
model.metabolites.co2_c:1
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# # Fix Succo and thmpp
# Ben spotted that our succoa and thmpp metabolites ended up with positive coefficients, which they should not have. So here I will quickly fix this.
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
model.reactions.biomass.add_metabolites({
model.metabolites.succoa_c: -1.95E-04,
model.metabolites.thmpp_c:-0.000446
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# # Carbohydrates
# So apparently I miss understood Ben; in the biomass reaction it is best to include the UDP-bound variant of the carbohydrates where that makes sense. This is because adding the carbohydrates in itself can lead to gluconeogenesis being triggered in the model to meetbiomass requirements.Also, for example for glycogen, it is known that this is made by assembling UDP-glucose, not free glucose. Then the energy accounting for polymerization is better and more correctly captured. (i.e. add the activated sugars)
#
# Note: with this, we will leave the stoichiometry the same. because the data as a weight percentage was determined based on just the glucose, not UDP-glucose. so I will not modify these values. As UDP will be released later this will correct for the additional mass that is brought into the reaction.
#
# In general, the synthesis of the activated sugars comes from a pyrophosphorylase, that takes the NTP and reacts with the sugar-1-phosphate to give the the NDP-sugar moeity that is used for the polymers. So according to this I will add these variants into the biomass reaction.
#
# So what to do with each carb:
# - glucose: becomes UDP-glucose, from g1p.
# - Galactose: from UDP-gal, from gal1p or udpg
# - Arabinose: from UDP-arabinose via ara1p and udp-xylose
# - xylose: from UDP-xylose via xylose-1-phosphate, or from UDP arabinose or UDPglucoronate (udpglcur_c)
# - mannose: polymer is mannan. make from mannose-1-phosphate to GDP-mannose and then consume GDP mannose.
#
# - fructose: polymer is inulin. the genes for the production of inulin seem to be missing. So for fructose I will just leave fructose in the reaction, as per bens recommendation.
#
model = cobra.io.read_sbml_model('../model/p-thermo.xml')
#for UDP-arbinose, add UDP-arabinose metabolite and synthesis
model.add_metabolites(Metabolite(id='udparab_c', name = 'UDP-L-arabinose', compartment = 'c', charge = 0, formula = 'C14H22N2O16P2'))
model.metabolites.udparab_c.annotation['kegg.compound'] = 'C00935'
model.metabolites.udparab_c.annotation['chebi'] = 'CHEBI:17983'
model.metabolites.udparab_c.annotation['metanetx.chemical'] = 'MNXM5976'
#add arabinose-1-phosphate
model.add_metabolites(Metabolite(id='ara1p_c', name = 'beta-L-Arabinose 1-phosphate', compartment = 'c', charge = 0, formula = 'C5H11O8P'))
model.metabolites.ara1p_c.annotation['kegg.compound'] = 'C03906'
model.metabolites.ara1p_c.annotation['chebi'] = 'CHEBI:15807'
#add reaction from ara1p to udparab
model.add_reaction(Reaction(id='ARAAT', name = 'UDP-sugar pyrophosphorylase '))
model.reactions.ARAAT.annotation['kegg.reaction'] = 'R08845'
model.reactions.ARAAT.annotation['ec-code'] = '2.7.7.64'
model.reactions.ARAAT.annotation['sbo'] = 'SBO:0000176'
model.groups.get_by_id('00520 - Amino sugar and nucleotide sugar metabolism').add_members(model.reactions.ARAAT)
model.reactions.ARAAT.add_metabolites({
model.metabolites.utp_c:-1,
model.metabolites.ara1p_c:-1,
model.metabolites.ppi_c:1,
model.metabolites.udparab_c:1
})
#add reaction from arabinose to ara1p
model.add_reaction(Reaction(id='ARAK', name = 'L-arabinokinase'))
model.reactions.ARAK.annotation['sbo'] = 'SBO:0000176'
model.reactions.ARAK.annotation['kegg.reaction'] = 'R01754'
model.reactions.ARAK.annotation['ec-code'] = '2.7.1.46'
model.groups.get_by_id('00520 - Amino sugar and nucleotide sugar metabolism').add_members(model.reactions.ARAK)
model.reactions.ARAK.add_metabolites({
model.metabolites.arab__L_c:-1,
model.metabolites.atp_c:-1,
model.metabolites.ara1p_c:1,
model.metabolites.adp_c:1,
model.metabolites.h_c:-1
})
#add udp-xylose met
model.add_metabolites(Metabolite(id='udpxyl_c', name='UDP-D-xylose', compartment ='c', formula ='C14H22N2O16P2', charge=0 ))
model.metabolites.udpxyl_c.annotation['kegg.compound'] = 'C00190'
model.metabolites.udpxyl_c.annotation['chebi'] = 'CHEBI:16082'
#add conversion from UDP-arabinose to UDP.xylose
model.add_reaction(Reaction(id='XYLAT', name = 'UDP-arabinose 4-epimerase'))
model.reactions.XYLAT.bounds = (-1000,1000)
model.reactions.XYLAT.annotation['sbo'] = 'SBO:0000176'
model.reactions.XYLAT.annotation['ec-code'] = '5.1.3.5'
model.reactions.XYLAT.annotation['kegg.reaction'] = 'R01473'
model.groups.get_by_id('00520 - Amino sugar and nucleotide sugar metabolism').add_members(model.reactions.XYLAT)
model.reactions.XYLAT.add_metabolites({
model.metabolites.udparab_c:-1,
model.metabolites.udpxyl_c:1
})
#add conversion of xylUDP from UDP-glucuronate.
model.add_reaction(Reaction(id='UDPGDC', name = 'UDP-glucuronate decarboxylase'))
model.reactions.UDPGDC.bounds = (0,1000)
model.reactions.UDPGDC.annotation['sbo'] = 'SBO:0000176'
model.reactions.UDPGDC.annotation['ec-code'] = '4.1.1.35'
model.reactions.UDPGDC.annotation['kegg.reaction'] = 'R01384'
model.groups.get_by_id('00520 - Amino sugar and nucleotide sugar metabolism').add_members(model.reactions.UDPGDC)
model.reactions.UDPGDC.add_metabolites({
model.metabolites.udpglcur_c:-1,
model.metabolites.udpxyl_c:1,
model.metabolites.co2_c:1,
model.metabolites.h_c:-3
})
model.reactions.UDPGDC.check_mass_balance()
# +
#conversion of udpglcur from udpg is already present.
#model.reactions.UDPGD
# +
#GDP-mannose from man1p: raction MAN1PT
#man1P is coupled to carbon metabolism.
# -
#remove all carbohydrates metabolites from biomass rct
model.reactions.biomass.add_metabolites({
model.metabolites.arab__L_c: 0.045625,
model.metabolites.gal_c:0.212236,
model.metabolites.glc__D_c:0.010725,
model.metabolites.xyl__D_c:0.204370,
model.metabolites.man_c:0.006411
})
#add udp/gdp-variant of carbohydrates and expell the udp/gdp too
model.reactions.biomass.add_metabolites({
model.metabolites.udparab_c: -0.045625,
model.metabolites.udpgal_c:-0.212236,
model.metabolites.udpg_c: -0.010725,
model.metabolites.udpxyl_c: -0.204370,
model.metabolites.gdpmann_c: -0.006411,
model.metabolites.udp_c:0.472956, #releasing the UDP, from the total of the above UDP related metabolites
model.metabolites.gdp_c: 0.006411 #releasing the GDP
})
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# Add SBO to metabolites where it was missing.
model.metabolites.udparab_c.annotation['sbo'] = 'SBO:0000247'
model.metabolites.ara1p_c.annotation['sbo'] = 'SBO:0000247'
model.metabolites.udpxyl_c.annotation['sbo'] = 'SBO:0000247'
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# # remove Glutathione
# In issue 64, we discussed why glutathione is in the model biomass pseudo-reaction. It seems to be unrealistic that it is included, instead bacillitiol should be added. Martyn will look into and include the bacillithiol steps. Our model is dependent on glutathione uptake as there is no biosynthesis pathway (as there shouldn't be). This reallly skews some other work we need to do. So in the mean time, while Martyn works on this bacillithiol addition, at least we can remove glutathione from the biomass reaction and from the medium so that we can continue and work further with some other things.
model.optimize().objective_value
#remove from biomass rct
model.reactions.biomass.add_metabolites({
model.metabolites.gthrd_c: 0.000223000000000001
})
#remove glutathione from the medium
model.reactions.EX_gthrd_e.bounds = (0,1000)
#check our model is no longer dependany on gthrd supply
model.optimize().objective_value
#save&commit
cobra.io.write_sbml_model(model,'../model/p-thermo.xml')
# Also Ben was curious what the current g/gCDW total of our biomass reaction was. So here i will write the script that can calculate that for us.
#check total g/gdcw biomass
biomass_g_g = []
for met in model.reactions.biomass.metabolites:
stoich = model.reactions.biomass.metabolites.get(met) #get stoichiometry in mmol/gcdw
if stoich == 1 :#ignore the biomass part, but need to include the ADP/Pi/UDP that is also made here
continue
else:
mw = met.formula_weight # get molecular weight of metabolite in g/mol
try:
weight_fraction = (mw/1000)*stoich #the g/gcdw of this metabolite
biomass_g_g.append(weight_fraction)
except TypeError: #for the met with R in it, just ignore
print(met.id)
#to find total g/gcdw of all mets, sum up this list
abs(sum(biomass_g_g))
|
notebooks/30. Update biomass reaction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 07_Convolutional_Neural_Networks
# In this notebook, we will see how to define simple convolutional neural networks.
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
import matplotlib.pyplot as plt
# %matplotlib inline
torch.manual_seed(777) # reproducibility
# -
# ### Convolutional Neural Network
# +
# Hyper parameters
num_epochs = 5
num_classes = 10
batch_size = 100
learning_rate = 0.001
# Device configuration
# device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
device = torch.device('cpu')
# Convolutional neural network (two convolutional layers)
class ConvNet(nn.Module):
def __init__(self, num_classes):
super(ConvNet, self).__init__()
# torch.nn.Conv2d(in_channels, out_channels, kernel_size, stride=1, padding=0, bias=True)
# torch.nn.MaxPool2d(kernel_size, stride=None, padding=0)
# torch.nn.Sequential(*args): a sequential container.
# Modules will be added to it in the order they are passed in the constructor.
# floor((28-5+2*2)/1)+1 = 28, 28 / 2 = 14
self.layer1 = nn.Sequential(
nn.Conv2d(1, 16, kernel_size=5, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
# floor((14-5+2*2)/1)+1 = 14, 14 / 2 = 7
self.layer2 = nn.Sequential(
nn.Conv2d(16, 32, kernel_size=5, stride=1, padding=2),
nn.ReLU(),
nn.MaxPool2d(kernel_size=2, stride=2))
self.fc = nn.Linear(7*7*32, num_classes)
def forward(self, x):
# x: (batch_size, 1, 28, 28) -> out: (batch_size, 16, 14, 14)
out = self.layer1(x)
# out: (batch_size, 16, 14, 14) -> out: (batch_size, 32, 7, 7)
out = self.layer2(out)
# out: (batch_size, 32, 7, 7) -> out: (batch_size, 32*7*7)
out = out.reshape(out.size(0), -1)
# out: (batch_size, 32*7*7) -> out: (batch_size, num_classes)
out = self.fc(out)
return out
model = ConvNet(num_classes).to(device)
print(model)
# -
# ### Loss function and Optimizer
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# ### DataLoader
# +
# MNIST dataset
train_dataset = torchvision.datasets.MNIST(root='./data',
train=True,
transform=transforms.ToTensor(),
download=True)
test_dataset = torchvision.datasets.MNIST(root='./data',
train=False,
transform=transforms.ToTensor())
# Data loader
train_loader = torch.utils.data.DataLoader(dataset=train_dataset,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_dataset,
batch_size=batch_size,
shuffle=False)
# -
# ### Train the network
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
running_loss = 0.0
for i, (images, labels) in enumerate(train_loader):
# Move tensors to the configured device
images = images.to(device)
labels = labels.to(device)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# zero the parameter gradients
optimizer.zero_grad()
# backward + optimize
loss.backward()
optimizer.step()
running_loss += loss.item()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, running_loss / 100))
running_loss = 0.0
# ### Test the network
# Test the model
# In test phase, we don't need to compute gradients (for memory efficiency)
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
# ### Save/Load the network parameters
# +
# Save the model checkpoint
torch.save(model.state_dict(), './data/cnn_model.ckpt')
# Load the model checkpoint if needed
# new_model = NeuralNet(input_size, hidden_size, num_classes).to(device)
# new_model.load_state_dict(torch.load('./data/cnn_model.ckpt'))
# -
# ## Practice: CIFAR10
#
# <img src="images/cifar10.png" width="400">
#
# The CIFAR-10 dataset has the following specification:
# - The images in CIFAR-10 are of size 3x32x32, i.e. 3-channel color images of 32x32 pixels in size.
# - CIFAR-10 has the ten classes: ‘airplane’, ‘automobile’, ‘bird’, ‘cat’, ‘deer’, ‘dog’, ‘frog’, ‘horse’, ‘ship’, ‘truck’.
#
# You have to define a convolutional neural network for performing image classifcation on the CIFAR-10 dataset as well as train and test the network.
# The architecture of the network should be:
# - first layer: conv2d(in_channels=3, out_channels=6, kernel_size=5) + ReLU
# - second layer: MaxPool2d(kernel_size=2, stride=2)
# - third layer: conv2d(in_channels=6, out_channels=16, kernel_size=5) + ReLU
# - fourth layer: Linear(in_features=?, out_features=120) + ReLU
# - fifth layer: Linear(in_features=120, out_features=84) + ReLU
# - sixth layer: Linear(84, num_classes)
# +
# Hyper parameters
num_epochs = 5
num_classes = 10
batch_size = 100
learning_rate = 0.001
# Device configuration
device = torch.device('cpu')
# transform images to tensors of normalized range [-1, 1]
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
train_dataset = torchvision.datasets.CIFAR10(root='./data', train=True,
download=True, transform=transform)
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size,
shuffle=True, num_workers=2)
test_dataset = torchvision.datasets.CIFAR10(root='./data', train=False,
download=True, transform=transform)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size,
shuffle=False, num_workers=2)
# Write the code to define the convolutional neural network for CIFAR-10
class Net(nn.Module):
def __init__(self, num_classes):
super(Net, self).__init__()
self.conv1 = nn.Conv2d(3, 6, 5)
self.pool = nn.MaxPool2d(2, 2)
self.conv2 = nn.Conv2d(6, 16, 5)
self.fc1 = nn.Linear(16 * 5 * 5, 120)
self.fc2 = nn.Linear(120, 84)
self.fc3 = nn.Linear(84, num_classes)
def forward(self, x):
x = self.pool(F.relu(self.conv1(x)))
x = self.pool(F.relu(self.conv2(x)))
x = x.view(-1, 16 * 5 * 5)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
model = Net(num_classes).to(device)
print(model)
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.Adam(model.parameters(), lr=learning_rate)
# Train the model
total_step = len(train_loader)
for epoch in range(num_epochs):
running_loss = 0.0
for i, (images, labels) in enumerate(train_loader):
# Move tensors to the configured device
images = images.to(device)
labels = labels.to(device)
# Forward pass
outputs = model(images)
loss = criterion(outputs, labels)
# Backward and optimize
# zero the parameter gradients
optimizer.zero_grad()
# backward + optimize
loss.backward()
optimizer.step()
running_loss += loss.item()
if (i+1) % 100 == 0:
print ('Epoch [{}/{}], Step [{}/{}], Loss: {:.4f}'
.format(epoch+1, num_epochs, i+1, total_step, running_loss / 100))
running_loss = 0.0
# Test the model
with torch.no_grad():
correct = 0
total = 0
for images, labels in test_loader:
images = images.to(device)
labels = labels.to(device)
outputs = model(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test images: {} %'.format(100 * correct / total))
|
07_Convolutional_Neural_Networks_answer.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.4-pre
# language: julia
# name: julia-0.4
# ---
# # Dynamic Modeling in Behavorial Ecology: Julia Edition
#
# In which we work through several examples from Mangel & Clark in Julia.
# patches are characterized the by following four parameters
type Patch
β::Float64 # MC's probability of predation
λ::Float64 # MC's probability of finding food
α::Int64 # MC's cost to visit
Y::Int64 # MC's value of food
end
# an Ecosystem is essentially an array of patches
typealias Ecosystem Array{Patch,1};
# +
# the patches from page 54
myplace = Ecosystem([
Patch(0., 0., 1, 0),
Patch(0.004, 0.4, 1, 3),
Patch(0.02, 0.6, 1, 5)
]);
x_c = 3::Int64; # critical energy threshold
C = 10::Int64; # max capacity
T = 20::Int64; # time horizon
# +
function xprime(x::Int64, α::Int64, y::Int64, x_critical::Int64, C::Int64)
return clamp(x - α + y, x_critical, C)
end;
function xprimeprime(x::Int64, α::Int64, x_critical::Int64, C::Int64)
return clamp(x - α, x_critical, C)
end;
# -
function findOptimalSelections(
aplace::Ecosystem,
C::Int64,
T::Int64,
x_c::Int64)
Fitness = zeros(Float64, C+1, T)
Selection = zeros(Int64, C+1, T)
for t = T:-1:1
for x = 0:C
if t == T
Fitness[x+1, t] = x > x_c ? 1.0 : 0.0
elseif x <= x_c
Fitness[x+1, t] = 0.0
else
patchoptions = [(1-P.β)*
(P.λ*Fitness[xprime(x, P.α, P.Y, x_c, C)+1,t+1] +
(1.0-P.λ)*Fitness[xprimeprime(x, P.α, x_c, C)+1,t+1])
for P in aplace]
(Fitness[x+1, t], Selection[x+1, t]) = findmax(patchoptions)
end
end
end
return Selection, Fitness
end;
(Selection, Fitness) = findOptimalSelections(myplace, C, T, x_c);
## compare to Mangel and Clark page 55
Fitness[5:11,19:-1:1]
|
patch_selection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (Data Science)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/datascience-1.0
# ---
# # Predicting Whether a Breast Cancer Sample is Benign or Malignant
#
# ## Learning Objectives:
#
#
# 1. Understand what SageMaker Script Mode is, and how it can be leveraged.
# 2. Read in data from S3 to SageMaker
# 3. User prebuilt SageMaker containers to build, train, and deploy customer sklearn model
# 4. Use batch transform to perform inferences and measure model performance.
#
#
# ## Introduction
# This is a breast cancer diagnoses dataset, where, for each sample, the sample is diagnosed as "Benign" or "Malignant". For each sample, a number of features are given as well. The source of the dataset is the [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Breast+Cancer+Wisconsin+(Diagnostic)).
#
# For this model, we will build, train and deploy a [Multi-layer Perceptron](https://scikit-learn.org/stable/modules/generated/sklearn.neural_network.MLPClassifier.html) using the sklearn library.
#
#
# ## Setup
# Ensure we have the right version of sagemaker
# %pip install sagemaker==2.48.0
# +
# Import required libraries and create necessary clients
import boto3
import matplotlib.pyplot as plt
import pandas
import sagemaker
from sagemaker.sklearn.estimator import SKLearn
from sagemaker.s3 import S3Downloader, S3Uploader
import sklearn
import sklearn.metrics as metrics
from sklearn import model_selection
import s3fs
role = sagemaker.get_execution_role()
sagemaker_session = sagemaker.Session()
BUCKET = sagemaker_session.default_bucket()
PREFIX = "breast_cancer"
# -
# ## Process the Data
# +
# Download the sample data
S3Downloader.download(
s3_uri="s3://sagemaker-sample-files/datasets/tabular/breast_cancer/wdbc.csv",
local_path="data",
sagemaker_session=sagemaker_session,
)
df_data = pandas.read_csv(
"data/wdbc.csv",
names=[
"id",
"diagnosis",
"radius_mean",
"texture_mean",
"perimeter_mean",
"area_mean",
"smoothness_mean",
"compactness_mean",
"concavity_mean",
"concave points_mean",
"symmetry_mean",
"fractal_dimension_mean",
"radius_se",
"texture_se",
"perimeter_se",
"area_se",
"smoothness_se",
"compactness_se",
"concavity_se",
"concave points_se",
"symmetry_se",
"fractal_dimension_se",
"radius_worst",
"texture_worst",
"perimeter_worst",
"area_worst",
"smoothness_worst",
"compactness_worst",
"concavity_worst",
"concave points_worst",
"symmetry_worst",
"fractal_dimension_worst",
],
)
df_data.head()
# -
# Get the feature names for analysis
features = list(set(df_data.columns) - set(["id", "diagnosis"]))
# One-hot encode the diagnosis column
df_data = pandas.get_dummies(df_data, columns=["diagnosis"])
# Get the data with encoded features. Malignant is now 1, Benign is 0
df_data = df_data.rename(columns={"diagnosis_M": "truth"})
df_data = df_data[features + ["truth"]]
# Preview the feature data frame
df_data.head()
# Split the data into training (70%) and test (30%) sets
train_df, test_df = model_selection.train_test_split(df_data, test_size=0.3)
# Move the truth column to the front of the training data set
train_df = train_df[["truth"] + features]
# Process the test data set
x_test = test_df[features]
y_test = test_df["truth"].tolist()
print(f"The test data has shape {x_test.shape}")
# +
# Copy the training data to s3 so that sagemaker can read it
train_df.to_csv("data/train_data.csv", index=False)
training_data_path = S3Uploader.upload(
local_path="data/train_data.csv",
desired_s3_uri=f"s3://{BUCKET}/{PREFIX}",
sagemaker_session=sagemaker_session,
)
# Do the same for the test data
x_test.to_csv("data/x_test.csv", index=False, header=False)
test_data_path = S3Uploader.upload(
local_path="data/x_test.csv",
desired_s3_uri=f"s3://{BUCKET}/{PREFIX}",
sagemaker_session=sagemaker_session,
)
# -
# ## Train the Model
# #### Define a training script
# +
# %%writefile train.py
import argparse
import joblib
import numpy as np
import os
import pandas as pd
from sklearn.neural_network import MLPClassifier
if __name__ == "__main__":
parser = argparse.ArgumentParser()
# Sagemaker specific arguments. Defaults are set in the environment variables.
parser.add_argument("--output-data-dir", type=str, default=os.environ.get("SM_OUTPUT_DATA_DIR"))
parser.add_argument("--model-dir", type=str, default=os.environ.get("SM_MODEL_DIR"))
parser.add_argument("--train", type=str, default=os.environ.get("SM_CHANNEL_TRAIN"))
args = parser.parse_args()
# Take the set of files and read them all into a single pandas dataframe
train_data=pd.read_csv(os.path.join(args.train, "train_data.csv"))
# Extract the labels from the first column
train_y = train_data["truth"]
train_X = train_data[train_data.columns[1:len(train_data)]]
# Use scikit-learn's MLP Classifier to train the model.
regr = MLPClassifier(random_state=1, max_iter=500).fit(train_X, train_y)
regr.get_params()
# Print the coefficients of the trained classifier, and save the coefficients
joblib.dump(regr, os.path.join(args.model_dir, "model.joblib"))
def model_fn(model_dir):
"""Deserialized and return fitted model
Note that this should have the same name as the serialized model in the main method
"""
regr = joblib.load(os.path.join(model_dir, "model.joblib"))
return regr
def predict_fn(input_data, model):
"""return the class and the probability of the class"""
prediction = model.predict(input_data)
pred_prob = model.predict_proba(input_data) # A numpy array
return np.array(pred_prob)
# -
# Create the estimator
sklearn = SKLearn(
entry_point="train.py",
instance_type="ml.c4.xlarge",
role=role,
py_version="py3",
framework_version="0.23-1",
sagemaker_session=sagemaker_session,
)
# Kick off the training job
sklearn.fit({"train": training_data_path})
# ## Make Batch Predictions
# Set up a batch transformer for predictions
transformer = sklearn.transformer(
instance_count=1, instance_type="ml.m4.xlarge", accept="text/csv"
)
# Start a transform job and wait for it to finish
batch_input_s3 = test_data_path
transformer.transform(batch_input_s3, content_type="text/csv", split_type="Line")
print("Waiting for transform job: " + transformer.latest_transform_job.job_name)
transformer.wait()
# Download the output data from S3 to local filesystem
batch_output = transformer.output_path
print(f"Batch transform results saved to {batch_output}")
S3Downloader.download(
s3_uri=batch_output,
local_path="data/output",
sagemaker_session=sagemaker_session,
)
# Preview the batch transform results
# !head data/output/*
# Load the predictions and measure performance
predictions = pandas.read_csv("data/output/x_test.csv.out", header=None)
predictions.reset_index(drop=True, inplace=True)
results = pandas.concat([predictions, pandas.Series(y_test)], axis=1)
results.columns = ["pred_0", "pred_1", "true"]
results["true"] = results["true"].astype(int)
# +
# Plot the AUC-ROC curve
fpr, tpr, threshold = metrics.roc_curve(results["true"], results["pred_1"])
roc_auc = metrics.auc(fpr, tpr)
plt.title("Receiver Operating Characteristic")
plt.plot(fpr, tpr, "b", label="AUC = %0.2f" % roc_auc)
plt.legend(loc="lower right")
plt.plot([0, 1], [0, 1], "r--")
plt.xlim([0, 1])
plt.ylim([0, 1])
plt.ylabel("True Positive Rate")
plt.xlabel("False Positive Rate")
plt.show()
# -
# ## Closing
# In this notebook we used SageMaker script mode to build, train, and deploy a sklearn model.
|
workshops/Bring_Your_Own_Sklearn_Classifier/byo_mlp_classifier/sklearn_bring_your_own_MLP_Classifier_Breast_Diagnostic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Parameter sweeps for replay speed analysis
# ## (Runs param sweep for Fig 3G)
# ## (No manuscript figures in this notebook)
# +
# %matplotlib inline
from copy import deepcopy as copy
import json
import sys
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from scipy import stats
from disp import set_font_size, set_n_x_ticks, set_n_y_ticks
from replay import smln, analysis, plot
cc = np.concatenate
LOAD_PRFX = 'PARAM_SWEEP_RESULTS'
SAVE_PRFX = 'PARAM_SWEEP_RESULTS_EXT'
# model
M_PARAMS = {
# Exc. PC membrane
'T_M_PC': 0.05, 'E_L_PC': -0.068, 'V_TH_PC': -0.036, 'V_R_PC': -0.068, 'T_R_PC': 0.008,
# Exc. PC architecture
'N_PC': 3000,
'W_PC_PC': .026, 'L_PC_PC': 0.053, 'W_MIN_PC_PC': 0.001,
# "Place-tuned" PC inputs
'R_MAX': 20, 'L_PL': .15,
'W_PC_PL': 0.02, 'S_PC_PL': 0,
# Inh. membrane
'T_M_INH': 0.005, 'E_L_INH': -0.06, 'V_TH_INH': -0.05, 'V_R_INH': -0.06, 'T_R_INH': 0.002,
# Inh. architecture
'N_INH': 300, # unused
'W_INH_PC': 0.003, 'W_PC_INH': 0.0002,
# gating inputs
'W_PC_G': 0.008216, 'S_PC_G': 0,
# synaptic reversal potentials & time constants
'E_E': 0, 'E_I': -0.08, 'T_E': 0.002, 'T_I': 0.002,
# Gating input firing rates for traj and replay epochs
'R_G': 125,
# potentiation
'SGM_MAX': 2, 'B_SGM': 1, 'R_SGM': 10
}
# simulation
S_PARAMS = {
"RNG_SEED": None,
"DT": 0.0005, "BOX_H": 2, "BOX_W": 2, "X_TRG": 1, "Y_TRG": -0.75,
'SPD': 0.2, 'TRJ': [(-1, .75), (0, .75), (0, -.75), (1, -.75)],
'TRG': [
{'T': 0.15, 'A': 0.02, 'R': 0.3, 'D': 0.002, 'X': 1, 'Y': -0.75},
],
"schedule": {"D_SMLN": .6},
}
# ANALYSIS
A_PARAMS = {
'MIN_START': 0.1, # s
'SMOOTH_FR': 4, # 2 ms
'MIN_GAP_DUR': .01, # s
'MIN_EVT_DUR': .03, # s
'EVT_DTCN_TH': 0.5, # Hz
'POS_T_CORR_TH': 0.85, # correlation
'SPEED_CALC_TRNC_FRAC': 0.1, # fraction
'MIN_DUR_SPD_CALC': 0.05, # s
'MAX_DUR_SPD_CALC': 0.4, # s
'MIN_ONE_WAY_EVTS_SPD_CALC': 5,
'SPD_CALC_TRNC_FRAC': 0.1, # fraction
'FR_NTRJ_MIN_BLOWUP': 1.5, # Hz
'FR_NTRJ_TRJ_BLOWUP_RATIO': .3, # frac
'SPD_CALC_MAX_ITER': 20,
'N_SPD_CALC_TRIALS': 10,
'SPD_CALC_START_OFFSET': 0.015, # s
}
# -
# ## Test replay speed calculation
# +
# run smln
m_params = copy(M_PARAMS)
s_params = copy(S_PARAMS)
s_params['RNG_SEED'] = 0
print('Running smln...')
rslt = smln.run(m_params, s_params)
print('Smln complete.')
# make plots
XYS_RASTER = [
(-.75, .75), (-.5, .75), (-.25, .75), (0, .75),
(0, .5), (0, .25), (0, 0), (0, -.25), (0, -.5),
(0, -.75), (.25, -.75), (.5, -.75), (.75, -.75)
]
EPOCHS = [(.156, .35)]
## potentiation profile
ax, c_ax = plot.ltp_ie_profile(rslt)
## add PC labels
for ctr, (x, y) in enumerate(XYS_RASTER):
ax.text(x, y, str(ctr), fontsize=14, horizontalalignment='center', verticalalignment='center')
## selected replay epoch spike sequences
for epoch in EPOCHS:
fig, axs = plot.spike_seq(rslt, epoch=epoch)
## raster for selected PCs
fig, axs = plot.raster_with_pc_inh(
rslt,
XYS_RASTER,
colors=np.zeros(len(XYS_RASTER)),
cmap='inferno',
nearest=1,
epoch=(0, .6),
trg_plt=[],
y_lim=(-1, 13),
y_ticks=[0, 3, 6, 9, 12],
smoothness=4)
## shade selected replay epochs
for epoch in EPOCHS:
axs[0].axvspan(*epoch, color='b', alpha=0.15)
axs[1].axvspan(*epoch, color='b', alpha=0.15)
axs[2].axvspan(*epoch, color='b', alpha=0.15)
## plot decoded trajectories during replay epochs
axs = plt.subplots(1, len(EPOCHS), figsize=(len(EPOCHS)*4, 3.5), tight_layout=True, squeeze=False)[1][0]
for epoch, ax in zip(EPOCHS, axs):
t, xy = analysis.decode_trj(rslt, epoch[0], epoch[1], .005, min_spks_wdw=10)
plot.decoded_trj(ax, rslt, t, xy);
# +
# test spd calc
starts, ends = analysis.get_evts(rslt, A_PARAMS)
start = starts[0] + 0.015
end = ends[0]
spd, debug = analysis.calc_spd(rslt, start, end, A_PARAMS, debug=True)
pfxs_spk = debug['pfxs_spk']
pfys_spk = debug['pfys_spk']
ds_trj_spk = debug['ds_trj_spk']
ts_spk = debug['ts_spk']
slp = debug['slp']
icpt = debug['icpt']
# plot xs and ys vs spike time
fig, axs = plt.subplots(4, 1, figsize=(12, 12), tight_layout=True)
axs[0].scatter(ts_spk, pfxs_spk)
axs[0].set_xlabel('Time (s)')
axs[0].set_ylabel('X (m)')
axs[0].set_title('Place fields (x)')
axs[1].scatter(ts_spk, pfys_spk)
axs[1].set_xlabel('Time (s)')
axs[1].set_ylabel('Y (m)')
axs[1].set_title('Place fields (y)')
axs[2].scatter(np.arange(len(debug['d_trj'])), debug['d_trj'])
axs[2].set_xlabel('Trj array index')
axs[2].set_ylabel('Dist along trj')
axs[2].set_title('Trajectory structure')
# scatter
axs[3].scatter(ts_spk, ds_trj_spk)
# line
x_line = np.array([ts_spk[0], ts_spk[-1]])
y_line = slp*x_line + icpt
axs[3].plot(x_line, y_line, c='r', lw=3)
axs[3].set_xlabel('Time (s)')
axs[3].set_ylabel('Dist along trj')
axs[3].set_title('Speed = {} m/s'.format(spd))
for ax in axs:
set_font_size(ax, 16)
# -
# # Param sweep
def spd_calc_sweep(sfx, a_params):
"""Calculate virtual speeds of replay events given existing parameter sweep file."""
# load existing param sweep results into dataframe
df, rslts, header_ = analysis.make_df('{}_{}'.format(LOAD_PRFX, sfx))
# write header of new save file
with open('{}_{}'.format(SAVE_PRFX, sfx), 'w') as f:
header = {
'SWEEP_PARAMS': header_['sweep_params'],
'M_PARAMS': header_['m_params'],
'S_PARAMS': header_['s_params'],
'A_PARAMS': a_params,
}
f.write(json.dumps(header) + '\n\n')
# get model and smln params
m_params = copy(header['M_PARAMS'])
s_params = copy(header['S_PARAMS'])
# set trigger and short smln dur for speed calc reruns
s_params['TRG'] = [{'T': 0.15, 'A': 0.01, 'R': 0.2, 'D': 0.002, 'X': 1, 'Y': -0.75}]
s_params['schedule']['D_SMLN'] = 0.6
offset = a_params['SPD_CALC_START_OFFSET']
# id all smln_ids w/o blowup
cond_0_blowup = df['FR_NTRJ'] >= a_params['FR_NTRJ_MIN_BLOWUP']
cond_1_blowup = (df['FR_NTRJ']/df['FR_TRJ']) >= a_params['FR_NTRJ_TRJ_BLOWUP_RATIO']
no_blowup = ~(cond_0_blowup | cond_1_blowup)
df_rerun = df[no_blowup]
# loop through valid smlns and rerun
for ctr, (smln_id, row) in enumerate(df_rerun.iterrows()):
print('Rerunning smln # {}/{}: SMLN_ID {}'.format(ctr, len(df_rerun), smln_id))
# set variable params for this smln
params_varied = {param: row[param] for param in header['SWEEP_PARAMS']['VARY']}
for param, param_val in params_varied.items():
m_params[param] = param_val
# loop over short trials for calculating speeds
spds = {}
durs = {}
for tr_ctr in range(a_params['SPD_CALC_MAX_ITER']):
sys.stdout.write('.')
# run smln with new rng_seed
rng_seed = np.random.randint(0, 1000)
s_params['RNG_SEED'] = rng_seed
rslt = smln.run(m_params, s_params)
# check for event in proper time window
starts, ends = analysis.get_evts(rslt, a_params)
for start_, end_ in zip(starts, ends):
if s_params['TRG'][0]['T'] - .01 <= start_ < s_params['TRG'][0]['T'] + .02:
start = start_
end = end_
break
else:
continue
# if evt is sufficiently long, calc speed
if a_params['MIN_DUR_SPD_CALC'] <= (end - start - offset) < a_params['MAX_DUR_SPD_CALC']:
durs[rng_seed] = end - start
spds[rng_seed] = analysis.calc_spd(rslt, start + offset, end, a_params)
else:
continue
if len(spds) >= a_params['N_SPD_CALC_TRIALS']:
break
print('')
# curate re-run results
save_dict = {
'SMLN_ID': smln_id,
'PARAMS': copy(params_varied),
'SPDS': copy(spds),
'DURS': copy(durs),
}
# append results to save file
with open('{}_{}'.format(SAVE_PRFX, sfx), 'a') as f:
f.write(json.dumps(save_dict) + '\n')
# ### SWEEP 0: W_PC_PC vs L_PC_PC
spd_calc_sweep('0_W_PC_PC_L_PC_PC.txt', A_PARAMS)
# ### SWEEP 1: W_PC_PC vs W_PC_INH
spd_calc_sweep('1_W_PC_PC_W_PC_INH.txt', A_PARAMS)
|
3B_param_regimes_sweep_ext.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Lecture 01 Python juypter notebook
# ### Use Spyder or Canopy Editor to create a new juypter (ipython) notebook
#
# Jupyter notebooks are useful for debugging and exploring. Use Jupyter notebooks to explore how Python works, expolore data, and to learn new functions.
#
# Jupyter notebooks and ipython are useful for figuring things out, but I want your HW to be submitted as a single *_py2.py* or *_py3.py* file. Where *.py* files are Python scripts.
# ### Be careful with pointers
x = [0,1,2]
y = x
x[0] = 3
print(x)
print(y)
import numpy as np
x = np.array([10])
y = x
z = x.copy()
#x = x + 10
x += 10
print(x,y,z)
x = x + 5
print(x,y)
x = x + 10
print(x,y,z)
# ### Be careful with Division!!!
a = 3
b = 2
print(a/b)
# +
from __future__ import division
'''
this line imports Python 3 division into Python 2
if you are using a script, these __future__
must be the first things in your Python script
'''
a = 3
b = 2
print(a/b)
print(a//b)
# -
a = 4
b = 2
c = a/b
print(c,type(c))
x[0.0]
print(0 == False)
print(1 == False)
print(0 == True)
print(1 == True)
# ### Floating point precission rounding error
0.1 + 0.2 == 0.3
np.isclose(0.1+0.2, 0.3)
# ### Sets
{1,2,3}
{1,5,5,5,2}
[1,2,3]
# ### Dictionary
param = {a:'hello'}
param[b] = 'hi'
print(param)
# ### Assigning new values at any time
a = None
b = None
a = 1
a
# ### The append function is soooooo useful
# ### We have lists in lists in lists
x = []
x.append(1)
x.append(2)
x.append(3)
x.append('string')
x.append([3,4,4,[1,2,2,3,3,0]])
print(x)
x[4]
x[4][3]
x[4][3][3]
x = 2
if x < 3:
print('x is 3')
print('end of the code block')
print('outside code block')
# ### This is how you can continue onto another line
x = 1 + 3 + \
2+4
print(x)
# ### If y is a tuple, you can assign y a new value, despite being a tuple
# The thing with tuples is that you can't change the
y = (1,2,3)
y = None
print(y)
|
lectures/lecture01/lecture01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Add event callbacks for on_hover and on_click
#
# Using the [set_map_event_handlers](https://docs.unfolded.ai/map-sdk/api/set-map-event-handlers) function it is possible to define event callbacks for the `on_hover` and `on_click` events. These events can return the data from the layer points or polygons if the user clicks or hovers on them.
from unfolded.map_sdk import UnfoldedMap, models
unfolded_map = UnfoldedMap(mapUUID='fb6aad80-eb4c-4f33-86eb-668772cc5fc4')
from sidecar import Sidecar
sc = Sidecar(title='Unfolded map', anchor='split-right')
with sc:
display(unfolded_map)
# We define the `on_hover` callback function:
import ipywidgets as widgets
output = widgets.Output()
@output.capture(clear_output=True)
def on_hover_output(info):
print('Hover event')
print(info)
output
# We define the `on_click` callback function:
import ipywidgets as widgets
output = widgets.Output()
@output.capture(clear_output=True)
def on_click_output(info):
print('Click event')
print(info)
output
# Here we register the defined callback functions. These functions will be called once you hover or click on the points or on the empty part of the map for the corresponding function.
unfolded_map.set_map_event_handlers({
'on_hover': on_hover_output,
'on_click': on_click_output
})
|
notebooks/Jupyter/Event Handling.ipynb
|