code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
# +
import pandas as pd
file_path = "Resources/purchase_data.csv"
purchase_data = pd.read_csv(file_path)
purchase_data.head()
# -
# Player Count
purchase_data['SN'].count()
total_players = {'Total Players': [purchase_data['SN'].count()]}
player_count = pd.DataFrame(total_players)
player_count
# +
# Purchasing Analysis (Total)
purchasing = {
'Number of Unique Items': purchase_data['Item Name'].nunique(),
'Average Purchase Price': purchase_data['Price'].mean(),
'Total Number of Purchases': purchase_data['Purchase ID'].count(),
'Total Revenue': purchase_data['Price'].sum()
}
purchasing_analysis = pd.DataFrame(purchasing, index=[0])
purchasing_analysis
# -
purchasing_analysis['Average Purchase Price'] = purchasing_analysis['Average Purchase Price'].map('${:.2f}'.format)
purchasing_analysis['Total Revenue'] = purchasing_analysis['Total Revenue'].map('${:,}'.format)
purchasing_analysis
# Gender Demographics
|
HeroesOfPymoli/.ipynb_checkpoints/main-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Term Project - Milestone #3
#
# <NAME>
#
# MAT 328 - Techniques in Data Science
#
# Spring 2022
#
# Professor Owen
#
# ----
# -
# # 0. Environment configuration
# ## 0.1 Import libraries
# +
# Data handling
from sklearn.model_selection import train_test_split
import pandas as pd
import numpy as np
import os
# Models
from sklearn.neighbors import KNeighborsClassifier
# Model evaluation
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
# Feature selection
from sklearn.feature_selection import RFE
# Visualization
from sklearn.tree import plot_tree
import seaborn as sns
import matplotlib.pyplot as plt
# Environment setup
# %matplotlib inline
pd.set_option('display.max_columns', None)
# -
# ## 0.2 Load data
# Get path to the repository's data folder
data_path = "/".join(os.getcwd().split("/")[0:-1]) + "/data/"
chd = pd.read_csv(f"{data_path}processed/model_data.csv")
chd.head()
# ## 0.3 Utility functions
def rfe_selection(model, data):
# Assign data-related variables
features = ["age", "sex", "is_smoking", "cigsPerDay", "BPMeds", "prevalentStroke", "prevalentHyp", "diabetes", "totChol", "sysBP", "diaBP", "BMI", "heartRate", "glucose", "TenYearCHD"]
df_vals = data.values
# Identify the column names of the features
inputs = df_vals[:,0:len(features)-1]
response_variable = df_vals[:,len(features)-1]
# Feature extraction
#dt_classifier = DecisionTreeClassifier(max_depth = 5)
rfe = RFE(model, step = 3)
fit = rfe.fit(inputs, response_variable)
# Create list with the target feature names
selected_features = list(np.array(features[:-1])[np.array(fit.support_)])
# Return a list containing the column names of the most relevant features
return selected_features
# # 1. K-Nearest Neighbor clustering
#
# ----
# ## 1.1 Split model data
# +
# Split data before evaluation for consistency in results
knn_x = chd.drop(columns = ["TenYearCHD"]) #Use columns identified above
knn_y = chd[["TenYearCHD"]]
knn_x_train, knn_x_test, knn_y_train, knn_y_test = train_test_split(knn_x, knn_y, test_size=0.2, random_state = 42)
# -
# ## 1.2 Determine ideal value of `k`
# +
# Create empty lists
k_list = []
model_error = []
# Loop through each depth
for k in range(2,21):
# Append the depth to the list
k_list.append(k)
# Create and fit model
knn_classifier = KNeighborsClassifier(n_neighbors = k)
knn_classifier = knn_classifier.fit(knn_x_train, np.ravel(knn_y_train))
# Test model and log error
knn_y_pred = knn_classifier.predict(knn_x_test)
model_error.append(accuracy_score(knn_y_pred, np.ravel(knn_y_test)))
# -
plt.plot(k_list, model_error)
plt.xlabel("k")
plt.ylabel("Accuracy")
# ## 1.3 Train classifier
knn = KNeighborsClassifier(n_neighbors = k_list[model_error.index(max(model_error))])
knn = knn.fit(knn_x_train, np.ravel(knn_y_train))
# ## 1.4 Evaluate model performance
# Predict test data with classifier
knn_y_pred_test = knn.predict(knn_x_test)
# +
# Calculate the accuracy of the model
knn_classifier_accuracy = accuracy_score(knn_y_pred_test, knn_y_test)
# Output accuracy
print(f"KNN Classifier Accuracy (d={k_list[model_error.index(max(model_error))]}): {knn_classifier_accuracy}")
# -
cm = confusion_matrix(knn_y_test, knn_y_pred_test)
cmd = ConfusionMatrixDisplay(cm)
cmd.plot()
cmd.ax_.set(xlabel='Predicted', ylabel='Actual')
print(classification_report(knn_y_test, knn_y_pred_test))
# # 2. New graph
# Get path to the repository's data folder
fig_path = "/".join(os.getcwd().split("/")[0:-1]) + "/figures/"
# +
# Create figure and correlation matrix
corr = chd.corr()
fig, ax = plt.subplots(figsize=(10, 8))
# Create mask to remove all duplicated correlations
mask = np.triu(np.ones_like(corr, dtype=np.bool_))
mask = mask[1:, :-1]
# Create hte heatmap
corr = corr.iloc[1:,:-1].copy()
heatmap = sns.heatmap(corr, mask=mask, annot=True, fmt=".2f", cmap='RdBu', vmin=-1, vmax=1, cbar_kws={"shrink": .8})
# Save figure
plt.savefig(f'{fig_path}correlation_plot.jpeg', dpi=300)
# Plot the graph
plt.yticks(rotation=0)
plt.show()
|
notebooks/Milestone3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" id="ZvFIZlL63mXh" outputId="512160d7-1f7e-4ca7-800c-5255fcae5813"
import torch
import torch.nn as nn
import time
import argparse
import os
import datetime
import gc
from torch.distributions.categorical import Categorical
from torch.utils.data import DataLoader
import math
import numpy as np
import torch.nn.functional as F
import random
import torch.optim as optim
from torch.autograd import Variable
from torch.optim import lr_scheduler
import matplotlib.pyplot as plt
from tqdm import tqdm_notebook
from torch.utils.data import Dataset
from torch.autograd import Variable
import matplotlib
matplotlib.use('Agg')
# visualization
# %matplotlib inline
from IPython.display import set_matplotlib_formats, clear_output
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import warnings
warnings.filterwarnings("ignore", category=UserWarning)
device = torch.device("cpu"); gpu_id = -1 # select CPU
gpu_id = '0' # select a single GPU
#gpu_id = '2,3' # select multiple GPUs
os.environ["CUDA_VISIBLE_DEVICES"] = str(gpu_id)
if torch.cuda.is_available():
device = torch.device("cuda")
print('GPU name: {:s}, gpu_id: {:s}'.format(torch.cuda.get_device_name(0),gpu_id))
print(device)
# + id="9x_DxlIa3mXu"
import os
import numpy as np
import torch
from torch.utils.data import Dataset
from torch.autograd import Variable
import matplotlib
matplotlib.use('Agg')
import matplotlib.pyplot as plt
class VehicleRoutingDataset(Dataset):
def __init__(self, num_samples, input_size, max_load=20, max_demand=9):
super(VehicleRoutingDataset, self).__init__()
if max_load < max_demand:
raise ValueError(':param max_load: must be > max_demand')
self.num_samples = num_samples
self.max_load = max_load
self.max_demand = max_demand
# Depot location will be the first node in each
locations = torch.rand((num_samples, 2, input_size + 1))
self.static = locations
# All states will broadcast the drivers current load
# Note that we only use a load between [0, 1] to prevent large
# numbers entering the neural network
dynamic_shape = (num_samples, 1, input_size + 1)
loads = torch.full(dynamic_shape, 0)
# All states will have their own intrinsic demand in [1, max_demand),
# then scaled by the maximum load. E.g. if load=10 and max_demand=30,
# demands will be scaled to the range (0, 3)
HalfofTheDemands = torch.randint(1, max_demand + 1, (num_samples,1,int(input_size / 2)))
TheOtherHalfDemands = torch.randint(-1 * max_demand, 0, (num_samples,1,int(input_size / 2)))
# Cat both demands with each other
demands = torch.cat((HalfofTheDemands,TheOtherHalfDemands),dim = 2).squeeze(1)
# Shuffling the demands tensor over the col dim
demands = demands[:,torch.randperm(demands.size()[1])]
# Shuffling the demands "converting demands back into numpy array for shuffling"
# Adding zero demand for the depot
demands = torch.cat((torch.zeros((num_samples,1,1)),demands.unsqueeze(1)),dim = 2)
# Normlize demands with the maximum load
demands = demands / float(max_load)
self.dynamic = torch.cat((loads, demands), dim=1)
def __len__(self):
return self.num_samples
def __getitem__(self, idx):
# (static, dynamic, start_loc)
return (self.static[idx], self.dynamic[idx], self.static[idx, :, 0:1])
def reward_fn(static, tour_indices):
"""
Euclidean distance between all cities / nodes given by tour_indices
"""
# Convert the indices back into a tour
idx = tour_indices.unsqueeze(1).expand(-1, static.size(1), -1)
tour = torch.gather(static.data, 2, idx).permute(0, 2, 1)
# Ensure we're always returning to the depot - note the extra concat
# won't add any extra loss, as the euclidean distance between consecutive
# points is 0
start = static.data[:, :, 0].unsqueeze(1)
y = torch.cat((start, tour, start), dim=1)
# Euclidean distance between each consecutive point
tour_len = torch.sqrt(torch.sum(torch.pow(y[:, :-1] - y[:, 1:], 2), dim=2))
return tour_len.sum(1).detach()
def render_fn(static, tour_indices, save_path):
"""Plots the found solution."""
plt.close('all')
num_plots = 3 if int(np.sqrt(len(tour_indices))) >= 3 else 1
_, axes = plt.subplots(nrows=num_plots, ncols=num_plots,sharex='col', sharey='row')
if num_plots == 1:
axes = [[axes]]
axes = [a for ax in axes for a in ax]
for i, ax in enumerate(axes):
# Convert the indices back into a tour
idx = tour_indices[i]
if len(idx.size()) == 1:
idx = idx.unsqueeze(0)
idx = idx.expand(static.size(1), -1)
data = torch.gather(static[i].data, 1, idx).cpu().numpy()
start = static[i, :, 0].cpu().data.numpy()
x = np.hstack((start[0], data[0], start[0]))
y = np.hstack((start[1], data[1], start[1]))
# Assign each subtour a different colour & label in order traveled
idx = np.hstack((0, tour_indices[i].cpu().numpy().flatten(), 0))
where = np.where(idx == 0)[0]
for j in range(len(where) - 1):
low = where[j]
high = where[j + 1]
if low + 1 == high:
continue
ax.plot(x[low: high + 1], y[low: high + 1], zorder=1, label=j)
ax.legend(loc="upper right", fontsize=3, framealpha=0.5)
ax.scatter(x, y, s=4, c='r', zorder=2)
ax.scatter(x[0], y[0], s=20, c='k', marker='*', zorder=3)
ax.set_xlim(0, 1)
ax.set_ylim(0, 1)
plt.tight_layout()
plt.savefig(save_path, bbox_inches='tight', dpi=200)
# + id="C_qKmVDf3mXw"
import torch
import time
def update_state(demand,dynamic_capcity,selected,c = 0):#dynamic_capcity(num,1)
depot = selected.squeeze(-1).eq(0) # Is there a group to access the depot
current_demand = torch.gather(demand,1,selected)
dynamic_capcity = dynamic_capcity - current_demand
if depot.any():
dynamic_capcity[depot.nonzero().squeeze()] = c
return dynamic_capcity.detach()#(bach_size,1)
def update_mask(demand,capcity,selected,mask,i):
# If there is a route to select a depot, mask the depot, otherwise it will not mask the depot
go_depot = selected.squeeze(-1).eq(0)
mask1 = mask.scatter(1, selected.expand(mask.size(0), -1), 1)
if capcity.gt(1).any():
print("warning")
if (~go_depot).any():
mask1[(~go_depot).nonzero(),0] = 0
if i+1 > (demand.size(1) / 2):
is_done = (mask1[:, 1:].sum(1) >= (demand.size(1) - 1)).float()
combined = is_done.gt(0)
mask1[combined.nonzero(), 0] = 0
# Mask any city if its demand is greater than the current truck's cap
a = demand > capcity + 1e-3
# Mask any city if its demand and the remaining capcity are greater than the truck limit "1"
b = torch.neg(demand.masked_fill(demand.gt(0), 0.)) + capcity > 1
mask = a + mask1 + b
"""
print("mask",mask)
print("mask1",mask1)
print('demand',demand)
print('capcity',capcity)
print('*************************')
"""
return mask.detach(),mask1.detach()
# -
# # HPN for Random PDP
# + id="D6ststo33mXx"
class Transformer_encoder_net(nn.Module):
"""
Encoder network based on self-attention transformer
Inputs :
h of size (bsz, nb_nodes+1, dim_emb) batch of input cities
Outputs :
h of size (bsz, nb_nodes+1, dim_emb) batch of encoded cities
score of size (bsz, nb_nodes+1, nb_nodes+1) batch of attention scores
"""
def __init__(self, nb_layers, dim_emb, nb_heads, dim_ff, batchnorm):
super(Transformer_encoder_net, self).__init__()
assert dim_emb == nb_heads* (dim_emb//nb_heads) # check if dim_emb is divisible by nb_heads
self.MHA_layers = nn.ModuleList( [nn.MultiheadAttention(dim_emb, nb_heads) for _ in range(nb_layers)] )
self.linear1_layers = nn.ModuleList( [nn.Linear(dim_emb, dim_ff) for _ in range(nb_layers)] )
self.linear2_layers = nn.ModuleList( [nn.Linear(dim_ff, dim_emb) for _ in range(nb_layers)] )
if batchnorm:
self.norm1_layers = nn.ModuleList( [nn.BatchNorm1d(dim_emb) for _ in range(nb_layers)] )
self.norm2_layers = nn.ModuleList( [nn.BatchNorm1d(dim_emb) for _ in range(nb_layers)] )
else:
self.norm1_layers = nn.ModuleList( [nn.LayerNorm(dim_emb) for _ in range(nb_layers)] )
self.norm2_layers = nn.ModuleList( [nn.LayerNorm(dim_emb) for _ in range(nb_layers)] )
self.nb_layers = nb_layers
self.nb_heads = nb_heads
self.batchnorm = batchnorm
def forward(self, h):
# PyTorch nn.MultiheadAttention requires input size (seq_len, bsz, dim_emb)
h = h.transpose(0,1) # size(h)=(nb_nodes, bsz, dim_emb)
# L layers
for i in range(self.nb_layers):
h_rc = h # residual connection, size(h_rc)=(nb_nodes, bsz, dim_emb)
h, score = self.MHA_layers[i](h, h, h) # size(h)=(nb_nodes, bsz, dim_emb), size(score)=(bsz, nb_nodes, nb_nodes)
# add residual connection
h = h_rc + h # size(h)=(nb_nodes, bsz, dim_emb)
if self.batchnorm:
# Pytorch nn.BatchNorm1d requires input size (bsz, dim, seq_len)
h = h.permute(1,2,0).contiguous() # size(h)=(bsz, dim_emb, nb_nodes)
h = self.norm1_layers[i](h) # size(h)=(bsz, dim_emb, nb_nodes)
h = h.permute(2,0,1).contiguous() # size(h)=(nb_nodes, bsz, dim_emb)
else:
h = self.norm1_layers[i](h) # size(h)=(nb_nodes, bsz, dim_emb)
# feedforward
h_rc = h # residual connection
h = self.linear2_layers[i](torch.relu(self.linear1_layers[i](h)))
h = h_rc + h # size(h)=(nb_nodes, bsz, dim_emb)
if self.batchnorm:
h = h.permute(1,2,0).contiguous() # size(h)=(bsz, dim_emb, nb_nodes)
h = self.norm2_layers[i](h) # size(h)=(bsz, dim_emb, nb_nodes)
h = h.permute(2,0,1).contiguous() # size(h)=(nb_nodes, bsz, dim_emb)
else:
h = self.norm2_layers[i](h) # size(h)=(nb_nodes, bsz, dim_emb)
# Transpose h
h = h.transpose(0,1) # size(h)=(bsz, nb_nodes, dim_emb)
return h, score
class Attention(nn.Module):
def __init__(self, n_hidden):
super(Attention, self).__init__()
self.size = 0
self.batch_size = 0
self.dim = n_hidden
v = torch.FloatTensor(n_hidden)
self.v = nn.Parameter(v)
self.v.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
# parameters for pointer attention
self.Wref = nn.Linear(n_hidden, n_hidden)
self.Wq = nn.Linear(n_hidden, n_hidden)
def forward(self, q, ref): # query and reference
self.batch_size = q.size(0)
self.size = int(ref.size(0) / self.batch_size)
q = self.Wq(q) # (B, dim)
ref = self.Wref(ref)
ref = ref.view(self.batch_size, self.size, self.dim) # (B, size, dim)
q_ex = q.unsqueeze(1).repeat(1, self.size, 1) # (B, size, dim)
# v_view: (B, dim, 1)
v_view = self.v.unsqueeze(0).expand(self.batch_size, self.dim).unsqueeze(2)
# (B, size, dim) * (B, dim, 1)
u = torch.bmm(torch.tanh(q_ex + ref), v_view).squeeze(2)
return u, ref
class LSTM(nn.Module):
def __init__(self, n_hidden):
super(LSTM, self).__init__()
# parameters for input gate
self.Wxi = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whi = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wci = nn.Linear(n_hidden, n_hidden) # w(ct)
# parameters for forget gate
self.Wxf = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whf = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wcf = nn.Linear(n_hidden, n_hidden) # w(ct)
# parameters for cell gate
self.Wxc = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Whc = nn.Linear(n_hidden, n_hidden) # W(ht)
# parameters for forget gate
self.Wxo = nn.Linear(n_hidden, n_hidden) # W(xt)
self.Who = nn.Linear(n_hidden, n_hidden) # W(ht)
self.wco = nn.Linear(n_hidden, n_hidden) # w(ct)
def forward(self, x, h, c): # query and reference
# input gate
i = torch.sigmoid(self.Wxi(x) + self.Whi(h) + self.wci(c))
# forget gate
f = torch.sigmoid(self.Wxf(x) + self.Whf(h) + self.wcf(c))
# cell gate
c = f * c + i * torch.tanh(self.Wxc(x) + self.Whc(h))
# output gate
o = torch.sigmoid(self.Wxo(x) + self.Who(h) + self.wco(c))
h = o * torch.tanh(c)
return h, c
class HPN_PDP(nn.Module):
def __init__(self, n_feature, n_hidden):
super(HPN_PDP, self).__init__()
self.city_size = 0
self.batch_size = 0
self.dim = n_hidden
# pointer layer
self.pointer = Attention(n_hidden)
self.TransPointer = Attention(n_hidden)
# LSTM encoder
self.encoder = LSTM(n_hidden)
# trainable first hidden input
h0 = torch.FloatTensor(n_hidden)
c0 = torch.FloatTensor(n_hidden)
self.h0 = nn.Parameter(h0)
self.c0 = nn.Parameter(c0)
self.h0.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
self.c0.data.uniform_(-1/math.sqrt(n_hidden), 1/math.sqrt(n_hidden))
r1 = torch.ones(1)
r2 = torch.ones(1)
r3 = torch.ones(1)
self.r1 = nn.Parameter(r1)
self.r2 = nn.Parameter(r2)
self.r3 = nn.Parameter(r3)
# embedding for the feature tensor
self.embedding_all = nn.Linear(2 * n_feature , n_hidden)
self.fc = nn.Linear(n_hidden + 1, n_hidden, bias=False)
self.fc1 = nn.Linear(n_hidden, n_hidden, bias=False)
# transformer's encoder
self.Transembedding_all = Transformer_encoder_net(6, 128, 8, 512, batchnorm=True)
# weights for GNN
self.W1 = nn.Linear(n_hidden, n_hidden)
self.W2 = nn.Linear(n_hidden, n_hidden)
self.W3 = nn.Linear(n_hidden, n_hidden)
# aggregation function for GNN
self.agg_1 = nn.Linear(n_hidden, n_hidden)
self.agg_2 = nn.Linear(n_hidden, n_hidden)
self.agg_3 = nn.Linear(n_hidden, n_hidden)
def forward(self,static, dynamic,deterministic = False,decoder_input = None):
"""
Parameters
----------
static: Array of size (batch_size, feats, num_cities)
Defines the elements to consider as static. For the TSP, this could be
things like the (x, y) coordinates, which won't change
dynamic: Array of size (batch_size, feats, num_cities)
Defines the elements to consider as static. For the VRP, this can be
things like the (load, demand) of each city. If there are no dynamic
elements, this can be set to None
decoder_input: Array of size (batch_size, num_feats)
Defines the outputs for the decoder. Currently, we just use the
static elements (e.g. (x, y) coordinates), but this can technically
be other things as well
"""
tour_idx, tour_logp = [], []
self.batch_size, self.city_size,input_size= static.size() # (B,size,feat)
# Always use a mask - if no function is provided, we don't update it
mask1 = torch.zeros((self.batch_size, self.city_size)).to(device)
mask = torch.zeros((self.batch_size, self.city_size)).to(device)
dynamic_capcity = dynamic[:,0,0].view(self.batch_size,-1)#bat_size
demands = dynamic[:,:,1].view(self.batch_size, self.city_size)#(batch_size,seq_len)
# Handle hidden and cell state for LSTM
h0 = self.h0.unsqueeze(0).expand(self.batch_size, self.dim)
c0 = self.c0.unsqueeze(0).expand(self.batch_size, self.dim)
h0 = h0.unsqueeze(0).contiguous()
c0 = c0.unsqueeze(0).contiguous()
# let h0, c0 be the hidden variable of first turn
h = h0.squeeze(0)
c = c0.squeeze(0)
max_steps = 2 * self.city_size
# Special Embedding for depot
# Cat both feature for embedding
all_feature = torch.cat((static,demands.unsqueeze(2)),dim = 2)
all_feature = torch.cat((all_feature,torch.cdist(static,decoder_input[:,:2].unsqueeze(1),p=2)),dim = 2)
# init embedding for feature vector
context = self.embedding_all(all_feature) #(B,size,n_hidden)
# ==================================================
# graph neural network encoder & transformer encoder
# ==================================================
Trans_hidden,_ = self.Transembedding_all(context) # (B,size,n_hidden)
TransPooled = Trans_hidden.mean(dim=1)
#Trans_hidden = Trans_hidden.reshape(-1, self.dim) # (B*size,n_hidden)
context = context.reshape(-1, self.dim) # (B*size,n_hidden)
# GNN layers
context = self.r1 * self.W1(context) + (1-self.r1) * F.relu(self.agg_1(context/(self.city_size-1)))
context = self.r2 * self.W2(context) + (1-self.r2) * F.relu(self.agg_2(context/(self.city_size-1)))
context = self.r3 * self.W3(context) + (1-self.r3) * F.relu(self.agg_3(context/(self.city_size-1)))
contextPooled = context.reshape(self.batch_size,self.city_size,self.dim).mean(dim=1)
pool = TransPooled + contextPooled
index = torch.zeros(self.batch_size).to(device).long()
for t in range(max_steps):
if not mask1[:, 1:].eq(0).any():
break
if t == 0:
_input = Trans_hidden[:, 0, :] # depot
decoder_input = torch.cat((_input,dynamic_capcity),dim = 1)
decoder_input = self.fc(decoder_input)
pool = self.fc1(pool)
decoder_input = decoder_input + pool
if t == 0:
mask, mask1 = update_mask(demands, dynamic_capcity, index.unsqueeze(-1), mask1, t)
# LSTM encoder
h, c = self.encoder(decoder_input, h, c)
# pointer
u1, _ = self.pointer(h, context.reshape(-1, self.dim))
u2 ,_ = self.TransPointer(h,Trans_hidden.reshape(-1, self.dim))
u = u1 + u2
u = 10 * torch.tanh(u)
u = u.masked_fill(mask.bool(), float("-inf"))
probs = F.softmax(u, dim=1)
# When training, sample the next step according to its probability.
# During testing, we can take the greedy approach and choose highest
if deterministic:
prob, index = torch.max(probs,dim=1) # Greedy
logp = prob.log()
else:
# Sampling
m = torch.distributions.Categorical(probs)
index = m.sample()
logp = m.log_prob(index)
is_done = (mask1[:, 1:].sum(1) >= (Trans_hidden.size(1) - 1)).float()
logp = logp * (1. - is_done)
# After visiting a node update the dynamic representation
#dynamic = update_fn(dynamic.permute(0,2,1), ptr.data).permute(0,2,1)
# Since we compute the VRP in minibatches, some tours may have
# number of stops. We force the vehicles to remain at the depot
# in these cases, and logp := 0
#is_done = dynamic.permute(0,2,1)[:, 1].sum(1).eq(0).float()
#logp = logp * (1. - is_done)
dynamic_capcity = update_state(demands, dynamic_capcity, index.unsqueeze(-1),c = 0.5)
mask, mask1 = update_mask(demands, dynamic_capcity, index.unsqueeze(-1), mask1, t)
# And update the mask so we don't re-visit if we don't need to
tour_logp.append(logp.unsqueeze(1))
tour_idx.append(index.data.unsqueeze(1))
#mask = mask_fn(ptr_prev, dynamic.permute(0,2,1), ptr.data)
_input = torch.gather(Trans_hidden, 1,
index.unsqueeze(-1).unsqueeze(-1).expand(Trans_hidden.size(0), -1,
Trans_hidden.size(2))).squeeze(1)
tour_idx = torch.cat(tour_idx, dim=1) # (batch_size, seq_len)
tour_logp = torch.cat(tour_logp, dim=1) # (batch_size, seq_len)
return tour_idx, tour_logp
def validate(data_loader, Critic, reward_fn, render_fn=None, save_dir='.',num_plot=5):
"""Used to monitor progress on a validation set & optionally plot solution."""
if not os.path.exists(save_dir):
os.makedirs(save_dir)
rewards = []
for batch_idx, batch in enumerate(data_loader):
with torch.no_grad():
static, dynamic, x0 = batch
static = torch.movedim(static,1,2).to(device) # (B,size,feat)
dynamic = torch.movedim(dynamic,1,2).to(device) # (B,size,feat)
x0 = torch.movedim(x0,1,2).squeeze(1).to(device) if len(x0) > 0 else None # (B,size,feat)
with torch.no_grad():
tour_indices, _ = Critic(static, dynamic, decoder_input = x0,deterministic=True)
reward = reward_fn(static.permute(0,2,1), tour_indices).mean().item()
rewards.append(reward)
if render_fn is not None and batch_idx < num_plot:
name = 'batch%d_%2.4f.png'%(batch_idx, reward)
path = os.path.join(save_dir, name)
render_fn(static.permute(0,2,1), tour_indices, path)
return np.mean(rewards)
# + [markdown] id="NIuJiHBo3mX1"
# # Training
# + id="RaQiVFNW3mX5" outputId="63ffb291-201a-4944-a1a8-3796ec00ece9"
########################
# Training Hyperparameters
#######################
# Dynamic ones
size = 20 # Size of the CVRP Problem
max_load = 30 # Max load for the truck
MAX_DEMAND = 9 # Max Demand for each agent
# Fixed Parameter for all sizes
TOL = 1e-3 # Tolerance for Actor-critic
TINY = 1e-15
learn_rate = 1e-4 # learning rate
batch_size = 512 # batch_size
train_size = 512
compare_size = 512
valid_size = 10000 # validation size
valid_batch = 10000
B_valLoop = 20
steps = 2500 # training steps
n_epoch = 100 # epochs
print('=========================')
print('prepare to train')
print('=========================')
print('Hyperparameters:')
print('size', size)
print('learning rate', learn_rate)
print('batch size', batch_size)
print('validation size', valid_size)
print('steps', steps)
print('epoch', n_epoch)
print('=========================')
###################
# Instantiate a training network and a baseline network
###################
try:
del Actor # remove existing model
del Critic # remove existing model
except:
pass
valid_data = VehicleRoutingDataset(valid_size,size,max_load,MAX_DEMAND)
valid_loader = DataLoader(valid_data, valid_batch, False, num_workers=0)
Actor = HPN_PDP(n_feature=2, n_hidden=128)
Critic = HPN_PDP(n_feature=2, n_hidden=128)
optimizer = optim.Adam(Actor.parameters(), lr=learn_rate)
# Putting Critic model on the eval mode
Actor = Actor.to(device)
Critic = Critic.to(device)
Critic.eval()
# uncomment these lines if trained with multiple GPUs
print(torch.cuda.device_count())
if torch.cuda.device_count()>1:
Actor = nn.DataParallel(Actor)
Critic = nn.DataParallel(Critic)
# uncomment these lines if trained with multiple GPUs
########################
# Remember to first initialize the model and optimizer, then load the dictionary locally.
#######################
epoch_ckpt = 0
tot_time_ckpt = 0
val_mean = []
val_std = []
plot_performance_train = []
plot_performance_baseline = []
################################################################# Restart Training With Check Points ######################################################
#********************************************# Uncomment these lines to re-start training with saved checkpoint #********************************************#
#************************************************************************************************************************************************************#
"""
checkpoint_file = "../input/pdpsize50/checkpoint_21-11-22--01-27-17-n50-gpu0.pkl"
checkpoint = torch.load(checkpoint_file, map_location=device)
epoch_ckpt = checkpoint['epoch'] + 1
tot_time_ckpt = checkpoint['tot_time']
plot_performance_train = checkpoint['plot_performance_train']
plot_performance_baseline = checkpoint['plot_performance_baseline']
Critic.load_state_dict(checkpoint['model_baseline'])
Actor.load_state_dict(checkpoint['model_train'])
optimizer.load_state_dict(checkpoint['optimizer'])
print('Re-start training with saved checkpoint file={:s}\n Checkpoint at epoch= {:d} and time={:.3f}min\n'.format(checkpoint_file,epoch_ckpt-1,tot_time_ckpt/60))
del checkpoint
"""
#*************************************************************************************************************************************************************#
#*********************************************# Uncomment these lines to re-start training with saved checkpoint #********************************************#
###################
# Main training loop
###################
start_training_time = time.time()
time_stamp = datetime.datetime.now().strftime("%y-%m-%d--%H-%M-%S")
zero_to_bsz = torch.arange(batch_size, device=device) # [0,1,...,bsz-1]
R = 0
C = 0
for epoch in range(0,n_epoch):
# re-start training with saved checkpoint
epoch += epoch_ckpt
###################
# Train model for one epoch
###################
start = time.time()
Actor.train()
for i in range(1,steps+1):
train_data = VehicleRoutingDataset(train_size,size,max_load,MAX_DEMAND)
train_loader = DataLoader(train_data, batch_size, False, num_workers=0)
for batch_idx, batch in enumerate(train_loader):
static, dynamic, x0 = batch
static = torch.movedim(static,1,2).to(device) # (B,size,feat)
dynamic = torch.movedim(dynamic,1,2).to(device) # (B,size,feat)
x0 = torch.movedim(x0,1,2).squeeze(1).to(device) if len(x0) > 0 else None
tour_indices, logprobs = Actor(static, dynamic,decoder_input = x0,deterministic=False)
R = reward_fn(static.permute(0,2,1), tour_indices)
with torch.no_grad():
tour_indices, _ = Critic(static, dynamic, decoder_input = x0,deterministic=True)
C = reward_fn(static.permute(0,2,1), tour_indices)
###################
# Loss and backprop handling
###################
loss = torch.mean((R - C) * logprobs.sum(dim=1))
optimizer.zero_grad()
loss.backward()
optimizer.step()
if i % 50 == 0:
print("epoch:{}, batch:{}/{}, reward:{}".format(epoch, i, steps, R.mean().item()))
time_one_epoch = time.time() - start
time_tot = time.time() - start_training_time + tot_time_ckpt
###################
# Evaluate train model and baseline on 1k random TSP instances
###################
Actor.eval()
mean_tour_length_actor = 0
mean_tour_length_critic = 0
for step in range(0,B_valLoop):
# compute tour for model and baseline
comp_data = VehicleRoutingDataset(compare_size,size,max_load,MAX_DEMAND)
comp_loader = DataLoader(comp_data, compare_size, False, num_workers=0)
for batch_idx, batch in enumerate(comp_loader):
static, dynamic, x0 = batch
static = torch.movedim(static,1,2).to(device) # (B,size,feat)
dynamic = torch.movedim(dynamic,1,2).to(device) # (B,size,feat)
x0 = torch.movedim(x0,1,2).squeeze(1).to(device) if len(x0) > 0 else None # (B,size,feat)
with torch.no_grad():
tour_indicesActor, _ = Actor(static, dynamic, decoder_input = x0,deterministic = True)
tour_indicesCritic, _ = Critic(static, dynamic, decoder_input = x0,deterministic = True)
R = reward_fn(static.permute(0,2,1), tour_indicesActor)
C = reward_fn(static.permute(0,2,1), tour_indicesCritic)
mean_tour_length_actor += R.mean().item()
mean_tour_length_critic += C.mean().item()
mean_tour_length_actor = mean_tour_length_actor / B_valLoop
mean_tour_length_critic = mean_tour_length_critic / B_valLoop
# evaluate train model and baseline and update if train model is better
update_baseline = mean_tour_length_actor + TOL < mean_tour_length_critic
print('Avg Actor {} --- Avg Critic {}'.format(mean_tour_length_actor,mean_tour_length_critic))
if update_baseline:
Critic.load_state_dict(Actor.state_dict())
print('My actor is going on the right road Hallelujah :) Updated')
###################
# val train model and baseline on 1k random TSP instances
###################
# Saving checkpoint and valied images
checkpoint_dir = os.path.join("checkpoint")
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
with torch.no_grad():
tour_len = validate(valid_loader, Critic, reward_fn, render_fn,checkpoint_dir, num_plot=5)
print('validation tour length:', tour_len)
# For checkpoint
plot_performance_train.append([(epoch+1), mean_tour_length_actor])
plot_performance_baseline.append([(epoch+1), mean_tour_length_critic])
# Print and save in txt file
mystring_min = 'Epoch: {:d}, epoch time: {:.3f}min, tot time: {:.3f}day, L_actor: {:.3f}, L_critic: {:.3f}, update: {}'.format(
epoch, time_one_epoch/60, time_tot/86400, mean_tour_length_actor, mean_tour_length_critic, update_baseline)
print(mystring_min)
print('Save Checkpoints')
torch.save({
'epoch': epoch,
'time': time_one_epoch,
'tot_time': time_tot,
'loss': loss.item(),
'plot_performance_train': plot_performance_train,
'plot_performance_baseline': plot_performance_baseline,
'mean_tour_length_val': tour_len,
'model_baseline': Critic.state_dict(),
'model_train': Actor.state_dict(),
'optimizer': optimizer.state_dict(),
}, '{}.pkl'.format(checkpoint_dir + "/checkpoint_" + time_stamp + "-n{}".format(size) + "-gpu{}".format(gpu_id)))
# -
# # Simple Test
# + id="eCdLTr6E3mX6"
# Saving checkpoint and valied images
checkpoint_dir = os.path.join("checkpoint")
if not os.path.exists(checkpoint_dir):
os.makedirs(checkpoint_dir)
valid_data = VehicleRoutingDataset(1000,size,max_load,MAX_DEMAND)
valid_loader = DataLoader(valid_data, 1000, False, num_workers=0)
with torch.no_grad():
tour_len = validate(valid_loader, Critic, reward_fn, render_fn,checkpoint_dir, num_plot=5)
print('validation tour length:', tour_len)
|
Random demands/Size20/RandomDemandsSize20.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Kwi5qGAACfX8"
# #Create the environment
# + colab={"base_uri": "https://localhost:8080/"} id="DX5uA-CcCWtb" executionInfo={"status": "ok", "timestamp": 1632349517599, "user_tz": -120, "elapsed": 453, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="0b3e3219-9c09-4adc-964d-f78eda9b4aee"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="54aQjssdChxP" executionInfo={"status": "ok", "timestamp": 1632349517957, "user_tz": -120, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="715271ac-cbb9-42cb-dc18-cf93caaaf265"
# %cd /content/drive/My Drive/ESoWC
# + id="0Hi_PCdCCj1N" executionInfo={"status": "ok", "timestamp": 1632349519697, "user_tz": -120, "elapsed": 1744, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}}
import pandas as pd
import xarray as xr
import numpy as np
#Our class
from create_dataset.make_dataset import CustomDataset
# + [markdown] id="27WuamAPC2ta"
# #Create the NOX dataset
# + [markdown] id="2BK77F2F2qWw"
# I want to apply the following changes to the dataset:
# * cut_region function: the dataset contains a larger region than the one we are analyzing, so we proceed to eliminate the excess area
# * rescale function: we change the scale of the dataset from 0.75 degree to 0.25 degree
# * resample function: we create a time series with an hourly frequency
# + [markdown] id="QcpDI3ek5eyi"
# Extremes of the region that we are analyzing:
# + id="8Wa-QKfk5fNO" executionInfo={"status": "ok", "timestamp": 1632349519698, "user_tz": -120, "elapsed": 7, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}}
lat_s = 43.0
lat_n = 51.0
lon_e = 4.0
lon_w = 12.0
# + [markdown] id="QXZwp4Nd5psw"
# Here we load the dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 349} id="uo_XUWzy1ERz" executionInfo={"status": "ok", "timestamp": 1632349522291, "user_tz": -120, "elapsed": 2599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="9da7014e-a35e-47b3-e47c-ee48569b94a8"
NOX_instance = CustomDataset('Data/MAY_HOURLY_2019_NOX_reduced.nc')
NOX_instance.get_dataset()
# + [markdown] id="Pp_NBnwS9Qv-"
# Here we applay the functions
# + colab={"base_uri": "https://localhost:8080/", "height": 314} id="CwQTlm7G199j" executionInfo={"status": "ok", "timestamp": 1632349522723, "user_tz": -120, "elapsed": 438, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="1f5047f0-934a-44f7-ee1f-f6a91705c4d2"
NOX_instance.cut_region(lat_n, lat_s, lon_e, lon_w)
NOX_instance.rescale()
NOX_instance.resample("1H")
NOX_Dataset = NOX_instance.get_dataset()
NOX_Dataset
# + [markdown] id="UYsKqDzC2eEL"
# We delete the crs variable which is useless
# + colab={"base_uri": "https://localhost:8080/", "height": 293} id="rOt9QU7c8Sk4" executionInfo={"status": "ok", "timestamp": 1632349523145, "user_tz": -120, "elapsed": 426, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="2fd90da3-711d-4627-cc50-dad3462ef753"
NOX_Dataset = NOX_Dataset.drop_vars('crs')
NOX_Dataset
# + [markdown] id="K1HU6inqEP1R"
# #Load weather dataset
# + id="FqP8U3JwERYu" colab={"base_uri": "https://localhost:8080/", "height": 600} executionInfo={"status": "ok", "timestamp": 1632349527131, "user_tz": -120, "elapsed": 3988, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="ae1970c9-8feb-422f-861f-fd488c25e3b9"
weather_dataset = xr.open_dataset('Data/05_2019_weather.nc')
weather_dataset
# + [markdown] id="GJhjAiODEJve"
# #Togheter NO and weather
# + id="xYKTRzpA9LNc" colab={"base_uri": "https://localhost:8080/", "height": 272} executionInfo={"status": "ok", "timestamp": 1632349527132, "user_tz": -120, "elapsed": 20, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="51a1a70e-2418-4e3b-ba20-187b3719d11c"
dataset_togheter = NOX_Dataset.merge(weather_dataset)
dataset_togheter
# + [markdown] id="z7-nSxpZ1Sub"
# We make some checks
# + colab={"base_uri": "https://localhost:8080/", "height": 314} id="ZYuZV-KdyfVo" executionInfo={"status": "ok", "timestamp": 1632349527660, "user_tz": -120, "elapsed": 545, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="ba76bf0a-478e-46fc-d24b-63814b4784df"
dataset_togheter.isel(time=[10]).cvl.plot()
# + colab={"base_uri": "https://localhost:8080/", "height": 314} id="S88Z8pC_0UEN" executionInfo={"status": "ok", "timestamp": 1632349528089, "user_tz": -120, "elapsed": 434, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}} outputId="99005615-d76e-4288-d75d-dc8e5dfc18c8"
dataset_togheter.isel(time=[10]).EMISSIONS_2019.plot()
# + [markdown] id="ttjyLLPw0XJ4"
# #Save dataset
# + id="exEDdDraxr73" executionInfo={"status": "ok", "timestamp": 1632349532135, "user_tz": -120, "elapsed": 4051, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}}
dataset_togheter.to_netcdf('Data/05_2019_weather_and_NOX.nc', 'w', 'NETCDF4')
# + id="ra_b97t10Zj0" executionInfo={"status": "ok", "timestamp": 1632349532136, "user_tz": -120, "elapsed": 13, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "04111805272867115673"}}
|
notebooks/exploratory/3.0_05_2019_weather_and_NOX_create_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/LordofSea/larynx_old/blob/master/Untitled4.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="NCdS5BKf3OHE" outputId="dfa06a5c-0549-42d3-ce98-9a7171fd039d"
# !apt-get install sox jq parallel
# + colab={"base_uri": "https://localhost:8080/"} id="YbRky7Ri3gHA" outputId="5b794c5a-bbe5-42c3-c256-e88eeff80cb0"
# !git clone --recursive https://github.com/rhasspy/larynx_old.git
# + colab={"base_uri": "https://localhost:8080/"} id="nzDSLBWW3r5r" outputId="7fd19e61-70af-4a05-f416-ede9fe5efbb0"
# %cd /content/larynx_old/
# %cd TTS
!!pip install https://github.com/rhasspy/gruut/releases/download/v0.5.0/gruut-0.5.0.tar.gz
# !pip3 install -r requirements.txt
# !python3 setup.py develop
# %cd /content/larynx_old/
# !pip3 install -r requirements.txt
# + colab={"base_uri": "https://localhost:8080/"} id="hjmETlUh4IR1" outputId="1f68fb32-d7dd-4b5e-ec8c-268973d9b86c"
# !./bin/larynx
# !mkdir -p local
# + colab={"base_uri": "https://localhost:8080/"} id="wICDHDr1-lS4" outputId="8a36a470-da85-4b33-ef38-0e909a0081c0"
# !unzip /content/drive/MyDrive/Wotan.zip
# + id="eF7CoqkCsoqA"
# !mv /content/larynx_old/Wotan /content/larynx_old/local
# + colab={"base_uri": "https://localhost:8080/"} id="OBliyPrb_YrH" outputId="213a883b-8115-480c-a713-68172ddce64d"
# !ls local/Wotan/data/ | head
# + colab={"base_uri": "https://localhost:8080/"} id="wNgDSeyw_h7F" outputId="7be77ee9-7b56-4378-99d0-6b831f073405"
# !soxi local/Wotan/data/1.wav
# + id="XvgcUNDq_zr6"
# !mkdir -p local/Wotan/larynx
# + id="zOVm_M01__NN"
# !truncate -s 0 local/Wotan/larynx/metadata.csv
# + id="KiSlPyf-A_pI"
# !find local/Wotan/data -type f -name '*.txt' | \
# while read -r fname; do \
# id="$(basename "${fname}" .txt)"; \
# text="$(cat "${fname}")"; \
# printf '%s|%s\n' "${id}" "${text}" >> local/Wotan/larynx/metadata.csv; \
# done
# + colab={"base_uri": "https://localhost:8080/"} id="N8TJN7KrBiVK" outputId="7df00b08-6a92-4856-9d1e-906b116173ca"
# !find local/Wotan/data -type f -name '*.wav' -print0 | \
# parallel -0 sox {} -r 22050 -c 1 -e signed-integer -b 16 -t wav local/Wotan/larynx/{/}
# + id="nmxczMxdBqXV"
# !./bin/larynx init /content/drive/MyDrive/Wotan/ \
# --language de-de \
# --name Wotan-Wilke-Möhring \
# --model-type glowtts \
# --dataset local/Wotan/larynx \
# --debug
# + colab={"base_uri": "https://localhost:8080/"} id="f9q_CGGxDhnf" outputId="addefcef-9f60-4947-c96d-139ec819ad46"
# !python3 TTS/TTS/bin/train_glow_tts.py \
# --config_path /content/drive/MyDrive/Wotan/config.json
|
Train_Larynx_DE.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="rnpcJ1oUSL4A" colab={"base_uri": "https://localhost:8080/", "height": 748} outputId="8d1d37b5-8503-47cd-e3ef-ce9381f19c8d"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from datetime import datetime
# %matplotlib inline
df2=pd.read_csv("/content/KVK_Field_sensor1 modified - Soda.csv",parse_dates=['Date Time'])
df2.drop(['Wind direction','Snowfall','Snow depth','Short-wave irradiation'],axis='columns',inplace=True)
temp=df2[['Temperature']]
rel=df2[['Relative Humidity']]
pres=df2[['Pressure']]
wind=df2[['Wind speed']]
rain=df2[['Rainfall']]
df=pd.read_csv("/content/KVK_ORG1.xlsx - feeds.csv",parse_dates=['created_at'])
df['created_at'] = df.created_at.dt.strftime('%Y-%m-%d %H:%M:%S')
df1=df.drop(['Air Temperature (degree Centigrate)','entry_id','Relative Air Humidity (%)','Soil moisture (Resistance-Ohm)'],axis='columns')
merged1=pd.concat([df1,temp,rel,pres,wind,rain],axis='columns')
soil=df[['Soil moisture (Centi Bar)']]
merged2=pd.concat([df2,soil],axis='columns')
merged2
# + id="e-U03gK6i-oz" colab={"base_uri": "https://localhost:8080/", "height": 423} outputId="e0446202-629d-41a2-b20b-7eb6dbfc25ab"
merged1.rename(columns = {'created_at':'Date Time'}, inplace = True)
veg=pd.read_csv("/content/Vegetation indices modified ee-chart.csv",parse_dates=['Vegitation Index'])
veg.drop(0,axis='rows',inplace=True)
veg.rename(columns = {'Vegitation Index':'Date Time'}, inplace = True)
veg.drop(['NDWI1','CAI','EVI','GNDVI','SAVI','MSI','ARI','DWSI','CVI'], axis = 1,inplace=True)
veg.dropna(inplace=True)
merged1['Soil moisture (Centi Bar)'].fillna('0',inplace=True)
merged1['Temperature'].fillna('0',inplace=True)
merged1['Relative Humidity'].fillna('0',inplace=True)
merged1['Pressure'].fillna('0',inplace=True)
merged1['Wind speed'].fillna('0',inplace=True)
merged1['Rainfall'].fillna('0',inplace=True)
merged1.dropna(axis='rows',inplace=True)
merged2.dropna(axis='rows',inplace=True)
veg
# + id="VMV3aXWVifC1" colab={"base_uri": "https://localhost:8080/", "height": 852} outputId="f233a362-9d1c-4d68-ae0b-b108af204ab8"
merged2=merged2[['Date Time','Soil moisture (Centi Bar)','Temperature','Relative Humidity','Pressure','Wind speed','Rainfall']]
merged1['Date Time'] = pd.to_datetime(merged1['Date Time'])
merged2['Date Time'] = pd.to_datetime(merged2['Date Time'])
final=pd.merge_asof(merged1,merged2.sort_values(['Date Time','Temperature']),left_on='Date Time',right_on='Date Time')
final['TEMPERATURE']= final[['Temperature_y','Temperature_x']].median(axis='columns',skipna=True)
final.drop(['Temperature_y','Temperature_x'],axis='columns',inplace=True)
final['RELATIVE_HUMIDITY']= final[['Relative Humidity_x','Relative Humidity_y']].median(axis='columns',skipna=True)
final.drop(['Relative Humidity_x','Relative Humidity_y'],axis='columns',inplace=True)
final['SOIL_MOISTURE']= final[['Soil moisture (Centi Bar)_x','Soil moisture (Centi Bar)_y']].median(axis='columns',skipna=True)
final.drop(['Soil moisture (Centi Bar)_x','Soil moisture (Centi Bar)_y'],axis='columns',inplace=True)
final['PRESSURE']= final[['Pressure_x','Pressure_y']].median(axis='columns',skipna=True)
final.drop(['Pressure_x','Pressure_y'],axis='columns',inplace=True)
final['WIND_SPEED']= final[['Wind speed_x','Wind speed_y']].median(axis='columns',skipna=True)
final.drop(['Wind speed_x','Wind speed_y'],axis='columns',inplace=True)
final['RAINFALL']= final[['Rainfall_x','Rainfall_y']].median(axis='columns',skipna=True)
final.drop(['Rainfall_x','Rainfall_y'],axis='columns',inplace=True)
final_ds=pd.merge_asof(final,veg.sort_values(['Date Time']),left_on='Date Time',right_on='Date Time')
final_ds.dropna(axis='rows',inplace=True)
final_ds
# + id="lRYeTMM0QLv3"
final_ds.to_csv("/content/final_ds.csv",index=False)
|
preprocessingcode.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
import os
# ### Lecture 10:
#
# - Learn about "object oriented programming" (OOP)
#
# - Learn how to create a "class"
#
# - Learn more about namespaces
#
# - Learn more about copies
#
#
#
#
# ### Object oriented programming
#
# Until now we haven't mentioned object oriented programming (OOP), yet we have been using **objects**from the beginning. Knowing how to create and use **objects** in Python is very powerful. Examples of **objects** that we have already encountered are the various data containers we have been using and things like plots. **Objects** have **methods** that can be used to change an object and **attributes** that describe features of an object.
#
# Now we will learn how to make our own objects with our own special blend of **attributes** and **methods**. The trick is to make a **class** and define it to have the desired **attributes** and **methods**.
#
# ### Classes
#
# To create an object with methods, we use a **class** definition, which is a blueprint or recipe defining the **attributes** and **methods** associated with the **class**. When we call the class, we create an **instance** of the **class**, also known as an **object**.
#
# Here is an example of a **class** definition:
class Circle:
"""
This is an example of a class called Circle
"""
import numpy as np # get some math power
# define some attributes of the Circle class
pi=np.pi # pi is now an attribute of this class too.
# initialize the class with the attribute r (no parentheses when called)
def __init__(self,r):
self.r=r # define a variable, r
# define some methods (these have parentheses when called)
def area(self):
return (1./2.)*self.pi*self.r**2
def circumference(self):
return 2.*self.pi*self.r
# Now we can create an **instance** of the Circle **class** called C with a radius of $r$.
r=3.0 # assign 3 to a variable r
C=Circle(r) # create a class instance with radius of 3.0
# We can use any of the attributes or methods of this class like this:
print ("The value of pi is: ",C.pi) # no parentheses!
print ("The radius of this circle is: ",C.r)# no parentheses!
print ("The area of a circle with radius = ",r,'is: ',C.area()) # with parentheses!
print ("The circumference of that circle is: ",C.circumference()) # with parentheses!
# We can also save the Circle class in a module, just as we did in earlier Lectures for functions. Then we can import it into other notebooks of scripts as desired.
#
# %%writefile Shapes.py
class Circle:
"""
This is an example of a class called Circle
"""
import numpy as np # get some math power
# define some attributes of the Circle class
pi=np.pi # pi is now an attribute of this class too.
# initialize the class with the attribute r (no parentheses when called)
def __init__(self,r):
self.r=r # define a variable, r
# define some methods (these have parentheses when called)
def area(self):
return (1./2.)*self.pi*self.r**2
def circumference(self):
return 2.*self.pi*self.r
# Now we can use it! Here is an example how:
import Shapes as S
newCirc=S.Circle(6.0)
print (newCirc.pi)
# ### Attributes and methods
#
# You might be wondering about some things by now. For example, you should have noticed is that when we asked for **C.pi** there were no parentheses, but both **C.area( )** and **C.circumference( )** did have parentheses. Why?
#
# The answer is that __r__ and **pi** are **attributes**, and **area** and **circumference** are **methods**. Did you notice that the method definitions look a lot like functions, but are inside the class definition. A **method** really is a function, but it is special in that it belongs to a **class** and works on the **instance** of the **class**. They can only be called by using the name of the **instance**, followed by a dot, followed by the **method** (with parentheses).
#
# ### More about classes
#
# Classes are not the same as functions. Although our **Shape** module can be imported just the same as any other module, to use it, we first have to create a class **instance** (**C=Shapes.Circle(r)**).
#
#
# All _**methods** (parts that start with **def**), have an **argument** list. The first **argument** has to be a reference to the class instance itself, which is always **self**, followed by any variables you want to pass into the **method**.
#
# The "**\_\_init\_\_**" method initializes the **instance** attributes. In the Circle class, the **\_\_init\_\_** method defined the **attribute** **r**, which gets passed in when the class is first called.
# Asking for any **attribute**, retrieves the current value of that **attribute**.
#
# But. Attributes can be changed:
#
print (C.r)
C.r=7.
print (C.r)
# To summarize: The **methods** (**area** and **circumference**) are defined just like any function except note the use of **self** as the first argument. This is required in all class method definitions. In our case, no other variables are passed in because the only one used is $r$, so the argument list consists of only **self**. Calling these **methods** requires no further arguments (the parentheses are empty) and the class returns the current values.
C.area()
# You can make a subclass (child) of the parent class which has all the attributes and methods of the parent, but may have a few attributes and methods of its own. You do this by setting up another class definition within a class.
#
# So, the bottom line about classes is that they are in the same category of things as variables, lists, dictionaries, etc. That is, they are "data containers" but with benefits. They hold data, and they also hold the methods to process those data.
#
#
# If you are curious about classes, there's lots more to know about them that we don't have time to get into. You can find useful tutorials online: https://www.python-course.eu/python3_object_oriented_programming.php
#
# or
#
# http://www.sthurlow.com/python/lesson08/ [but be careful with this one as it is for Python 2.7, so the **print** statements won't work without parentheses, e.g., **print ('this way')**, not, **print 'not this way'**. ]
#
#
# ### Namespaces
#
# Another thing you might be wondering about is why did we import **NumPy** inside the class definition when it was imported into the notebook at the top? The answer is we didn't have to. The class definition works perfectly well without it in this case. But if we don't import **Numpy** within in the Shape module, the module won't work at all because it doesn't "know" about **NumPy**. So in the module, you have to import whatever you need to run the module.
#
#
# ### Copies
#
# Another issue we have been tiptoeing around is the concept of a copy of an object and what that means. In Python, this can be a bit confusing. When we define some simple variables, the behavior is pretty much what you might expect:
x=3 # define x
y=x # set y equal to x
print (y) # print out y
x=4 # change the value of X
print (y) # and y is still equal to its first definition.
# But if we define a list object (a _compound_ object with more than one variable), things get weird:
L1=['spam','ocelot',42] # define the list
L2=L1 # make a copy of the list
print (L2) # print the copy
L1[2]='not an ocelot' # change the original
print (L2) # and oops - the copy got changed too!
# This means that **L1** and **L2** refer to the SAME OBJECT. So how do I make a copy that is its own object (doesn't change)? For simple lists (that do not contain sublists), we already learned how to do this:
L3=L1[:]
print (L3)
L1[2]=42
print (L3)
# This approach breaks down if the object is more complicated. The copies will sometimes be subject to mutation. (Try this yourself!).
#
# To avoid this problem, there is a module called **copy** with a function called **deepcopy**, which will make an independent copy of the object in question:
# +
from copy import deepcopy
L1=['spam','ocelot',42] # define the list
L2=deepcopy(L1) # make a copy of the list
print ("L2: ",L2) # print the copy
L1[2]='not an ocelot' # change the original
print ("L1: ",L1)
print ("L2: ",L2) # and bingo, L2 didn't
# -
# clean up
os.remove('Shapes.py')
|
Lecture_10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numerical Methods in Scientific Computing
# # Assignment 5
# ## <NAME>
# ## (ME16B077)
# # Q1.
#
# Equioscillation theorem
# Let $f \in C[−1, 1]$ and $p(x)$ be a polynomial whose degree doesn’t exceed n. p minimizes $||f − p||_{\infty}$
# iff $f − p$ equioscillates at $n + 2$ points.
#
# p is the $minimax$ $approximation$ to f as the infinity norm tries to minimze the maximum error.
#
# This theorem can be proved by showing the assumption that p is not the minimax approximation cannot be true given $f − p$ equioscillates at $n + 2$ points.(reductio ad absurdum)
#
# Let $f − p$ equioscillates at $n + 2$ points and if $p$ is not the minimax approximation then there must be some polynomial $q \in P_n$ such that $p+q \in P_n$ is the minimax approximation. Since the curve $f-p-q$ must have smaller infinity norm, adding $q$ to $p$ reduces the size of the modulus of error function $f − p$ on all $n + 2$ equioscillation points.
#
# In particular, this must mean that $q$ alternates in sign on these $n + 2$ points, and thus must have at least $n + 1$ zeros. Since $q \in P_n$, this is impossible, contradicting our initial assumption that $p$ is not the minimax approximation. Hence p is the minimax approximation to f which is same as p minimizes $||f − p||_{\infty}$.
#
# To prove the uniqueness of p, let p is best approximation with equioscillation points $x_0, x_1, ..., x_{n+1}$ and $x_0< x_1< ...< x_{n+1}$. Suppose $||f − q||_{\infty} \leq ||f − p||_{\infty}$ for some $q \in P_n$. Then $p-q$ must be $\leq 0$ at $x_0, x_2, ...$ and $\geq 0$ at $x_1, x_3, ...$. Which implies p-q has atleast n+1 roots in total which is not possible for a poynomial $\in P_n$ unless p=q. Hence p is unique.
# # Q2.
#
# Let $E = |x| - x^2 - 1/8$. To find maximum modulus values of $E$ we can differentiate and equate to $0$ in two separate intervals [-1,0) and (0,1].
#
# \begin{equation}
# 1-2x=0 \Rightarrow x=1/2 \quad ; \quad -1-2x=0 \Rightarrow x=-1/2
# \end{equation}
#
# \begin{equation}
# E(x=-1/2) = 1/8 \quad ; \quad E(x=1/2) = 1/8
# \end{equation}
#
# Taking into consideration the boundary values and at x=0,
# \begin{equation}
# E(x=-1) = -1/8 \quad ; \quad E(x=0) = -1/8 \quad ; \quad E(x=1) = -1/8
# \end{equation}
# which are also the maximum modulus values.
#
# Therefore we can say that E equioscillates at $x=\pm1, \pm1/2, 0$. Hence, as $f-p$ equioscillates at $3+2$ points$(n=3)$, p is the unique polynomial of all polynomials whose degree doesn’t exceed 3 that minimizes $||f − p||_{\infty}$ and is the best approximation in infinity norm.
#
import numpy as np
import matplotlib.pyplot as plt
# +
# Legendre Nodes
x_L = [-0.8611363115940525752239464888928095050957253796297176376157219209065294714950488657041623398844793052105769209319781763249637438391157919764084938458618855762872931327441369944290122598469710261906458681564745219362114916066097678053187180580268539141223471780870198639372247416951073770551,
-0.3399810435848562648026657591032446872005758697709143525929539768210200304632370344778752804355548115489602395207464932135845003241712491992776363684338328221538611182352836311104158340621521124125023821932864240034767086752629560943410821534146791671405442668508151756169732898924953195536,
0.3399810435848562648026657591032446872005758697709143525929539768210200304632370344778752804355548115489602395207464932135845003241712491992776363684338328221538611182352836311104158340621521124125023821932864240034767086752629560943410821534146791671405442668508151756169732898924953195536,
0.8611363115940525752239464888928095050957253796297176376157219209065294714950488657041623398844793052105769209319781763249637438391157919764084938458618855762872931327441369944290122598469710261906458681564745219362114916066097678053187180580268539141223471780870198639372247416951073770551]
# Chebyshev Nodes
x_C = [np.cos((2*i-1)*np.pi/8) for i in range(1,5)]
nodes = {'Legendre':x_L, 'Chebyshev':x_C}
# Lagrange polynomials
def Lagrange(xnodes,x,i):
f = 1
nnodes = np.size(xnodes)
for j in range(0,nnodes):
if j==i:continue
f = f*(x-xnodes[j])/(xnodes[i]-xnodes[j])
return f
for node_type in nodes:
xnodes = np.array(nodes[node_type])
# fnodes = xnodes**2 + (1/8)
fnodes = np.abs(xnodes)
xplot = np.linspace(-1,1,50)
# f_plot = xplot**2 + (1/8)
f_plot = np.abs(xplot)
f_inter = 0
for i in range(0,4):
f_inter = f_inter + fnodes[i]*Lagrange(xnodes,xplot,i);
plt.rcParams["figure.figsize"] = [7,5]
plt.plot(xplot,f_inter,color='b', label='p_'+node_type[0]+'(x)')
plt.plot(xnodes,fnodes, marker='o',color='r', label=str(node_type)+"Nodes", linestyle=' ')
plt.legend()
plt.grid()
plt.xlabel('x')
plt.ylabel('y')
plt.title('p_'+node_type[0]+'(x)')
plt.show()
# -
xnodes = np.array(nodes['Legendre'])
print('Legendre Nodes and Function values')
print(list(map(lambda x, y:(x,y), xnodes, np.abs(xnodes))))
xnodes = np.array(nodes['Chebyshev'])
print('\nChebyshev Nodes and Function values')
print(list(map(lambda x, y:(x,y), xnodes, np.abs(xnodes))))
# - Norm 2 Error
#
# $p(x) = x^2 + 1/8$
# \begin{equation}
# || f(x)-p(x) ||_2 = \left(\int_{-1}^{1} (|x|-x^2-1/8)^2dx \right)^{1/2} = \left(\int_{-1}^{0} (-x-x^2-1/8)^2dx +\int_{0}^{1} (x-x^2-1/8)^2dx \right)^{1/2}
# \end{equation}
# \begin{equation}
# = \left((0+0+0+0+0+0)-(-1/5-1/3-1/64+1/2-1/12+1/8) + (1/5+1/3+1/64-1/2+1/12-1/8)-(0+0+0+0+0+0\right)^{1/2} = (7/480)^{1/2} = 0.12076147288491178
# \end{equation}
#
# $p_L(x) = 0.832558x^2 + 0.243748$
# \begin{equation}
# || f(x)-p_L(x) ||_2 = \left[ 0.832558^2x^5/5 + x^3/3 + 0.243748^2x + 0.832558*x^4/2 + 2*0.832558*0.243748*x^3/3 + 0.243748*x^2\right]_{-1}^0 + \left[ 0.832558^2x^5/5 + x^3/3 + 0.243748^2x - 0.832558*x^4/2 + 2*0.832558*0.243748*x^3/3 - 0.243748*x^2\right]_{0}^1
# \end{equation}
#
# \begin{equation}
# = ((-0.00663955) + 0.00663955)^{1/2} = 0.1152
# \end{equation}
#
# $p_C(x) = 0.765367x^2 + 0.270598$
# \begin{equation}
# || f(x)-p_C(x) ||_2 = \left[ 0.765367^2x^5/5 + x^3/3 + 0.243748^2x + 0.765367*x^4/2 + 2*0.765367*0.243748*x^3/3 + 0.243748*x^2\right]_{-1}^0 + \left[ 0.765367^2x^5/5 + x^3/3 + 0.243748^2x - 0.765367*x^4/2 + 2*0.765367*0.243748*x^3/3 - 0.243748*x^2\right]_{0}^1
# \end{equation}
#
# \begin{equation}
# = (-(-0.00784337) + 0.00784337)^{1/2} = 0.1252
# \end{equation}
#
# - Infinity Norm Error
# \begin{equation}
# || f(x)-p(x) ||_{\infty} = 0.125
# \end{equation}
#
# \begin{equation}
# || f(x)-p_L(x) ||_{\infty} = 0.2437
# \end{equation}
#
# \begin{equation}
# || f(x)-p_C(x) ||_{\infty} = 0.2705
# \end{equation}
# | Approximation | $||.||_2$ | $||.||_{\infty}$ |
# |---------------|-----------|------------------|
# | f(x)-p(x) | 0.1207 | 0.1250 |
# | f(x)-p_L(x) | 0.1152 | 0.2437 |
# | f(x)-p_C(x) | 0.1252 | 0.2705 |
#
# Within the polynomial approximation using Legendre nodes and Chebyshev nodes, the Legendre nodes give better polynomial approximation to the function in both 2-norm and infinity-norm.
#
# Based on the infinity-norm we can see that $p(x)$ has the minimum value which is in line with the equioscillation theorem. For the 2-norm the polynomial obtained using Legendre nodes is the best approximation.
#
# Accuracy based on 2-norm
# \begin{equation}
# p_L(x) [best] > p_C(x) > p(x) [worst]
# \end{equation}
#
# Accuracy based on infinity-norm
# \begin{equation}
# p(x) [best] > p_L(x) > p_C(x) [worst]
# \end{equation}
# # Q3.
#
# \begin{equation}
# \int_a^b w(x)f(x)dx - \sum_{i=1}^n w_if(x_i) = \frac{f^{(2n)}(\xi)}{(2n)!} ||p_n(x)||_w^2
# \end{equation}
#
# To prove this we need the error formula for the Hermite interpolation which is given by.
# \begin{equation}
# f(x)-p_{2n-1}(x) = \frac{f^{(2n)}(\xi)}{(2n)!} \prod_{j=1}^{n} (x-x_j)^2, \quad \xi \in (a,b)
# \end{equation}
#
# We know that Gaussian Quadrature $\sum_{i=1}^n w_if(x_i)$ yields an exact result for polynomials of degree 2n − 1 or less. Hence $\sum_{i=1}^n w_if(x_i)$ = $\int_a^b w(x)p_{2n-1}(x)dx$ where $p_{2n-1}(x)$ is a polynomial of degree atmost 2n-1,
# \begin{equation}
# \int_a^b w(x)f(x)dx - \sum_{i=1}^n w_if(x_i) = \int_a^b w(x)f(x)dx - \int_a^b w(x)p_{2n-1}(x)dx = \int_a^b w(x)(f(x)-p_{2n-1})dx
# \end{equation}
#
# Substituting the error formula for Hermite interpolation in the above equation,
# \begin{equation}
# \int_a^b w(x)(f(x)-p_{2n-1})dx = \int_a^b w(x)(\frac{f^{(2n)}(\xi)}{(2n)!} \prod_{j=1}^{n} (x-x_j)^2)dx = \frac{f^{(2n)}(\xi)}{(2n)!}\int_a^b w(x)(\prod_{j=1}^{n} (x-x_j))^2dx
# \end{equation}
#
# $\prod_{j=1}^{n} (x-x_j)$ has the leading coefficient as 1 and is the monic orthogonal polynomial corresponding to weight function $w(x)$. So, $\int_a^b w(x)(\prod_{j=1}^{n} (x-x_j))^2dx$ is $||p_n(x)||_w^2$.
# \begin{equation}
# \Rightarrow \int_a^b w(x)f(x)dx - \sum_{i=1}^n w_if(x_i) = \frac{f^{(2n)}(\xi)}{(2n)!} ||p_n(x)||_w^2
# \end{equation}
# # Q4.
#
# \begin{equation}
# f(x) = a_0 + \sum_{k=1}^n(a_kcos(2k\pi x)+b_ksin(2k\pi x))
# \end{equation}
#
# We can absorb $a_0$ from the equation into $f(x)$ as it is a constant term and after the integration in range $[-1,1]$ converts to $2a_0$. Re-writing the equation as
# \begin{equation}
# F(x) = f(x) - a_0 = \sum_{k=1}^n(a_kcos(2k\pi x)+b_ksin(2k\pi x))
# \end{equation}
#
# Exact integration is given by,
# \begin{equation}
# \int_{-1}^{1} F(x) = \left[\sum_{k=1}^n \frac{1}{2k\pi}(a_ksin(2k\pi x)-b_kcos(2k\pi x))\right]_{-1}^{1} = \left[\sum_{k=1}^n \frac{1}{2k\pi}(2a_ksin(2k\pi x))\right]_{0}^{1}
# \end{equation}
#
# \begin{equation}
# \left[\sum_{k=1}^n \frac{1}{2k\pi}(2a_ksin(2k\pi x))\right]_{0}^{1} = \sum_{k=1}^n \frac{1}{2k\pi}(2a_ksin(2k\pi 1)) = 0
# \end{equation}
#
# +
# Define Function
def func(N):
# N+1 points
h = 1/N
# X = np.array([h*i for i in range(N+1)])
X = np.linspace(-1,1,N+1)
a_k = np.linspace(-1,1,20)
b_k = np.linspace(-1,1,20)
F = 0
for i in range(20):
F = F + (a_k[i]*np.cos(2*(i+1)*np.pi*X) + b_k[i]*np.sin(2*(i+1)*np.pi*X))
return X, F
# Trapezoidal Rule
def trap_rule(N):
# N+1 points
h = 1/N
X, F = func(N)
I_trap = (h/2)*sum([F[i]+F[i+1] for i in range(0,N)])
return I_trap
def exact_integral():
return 0
I_trap_list = []
for k in range(1,81):
I_trap_list.append(trap_rule(k))
print('k \t Trapezoidal Rule')
print('-'*30)
for k in range(1,81):
print(k,'\t',I_trap_list[k-1])
# -
# The error alternates between close to 0 and significant values for k greater than n and only becomes close to zero for k greater than 2n.
# Plot of the function for reference.
x, y = func(10000)
plt.plot(x, y)
plt.show()
# # Q5.
# +
# Define the function
def func_q5(X):
return np.exp(-np.power(X,2))
# Calculate the Integral using Gauss Quadrature
N = [i for i in range(3, 52)]
I_Gauss = []
for n in N:
X, W = np.polynomial.legendre.leggauss(n)
F = func_q5(X)
I_Gauss.append(sum([W[i]*F[i] for i in range(n)]))
# Obtained from wolfram-alpha.
I_exact = 1.493648265624854050798934872263706010708999373625212658055
# Plot the results
plt.loglog(N, np.abs(np.subtract(I_Gauss,I_exact)),
marker='o',color='r', label="abs error", linestyle='dashed')
plt.grid()
plt.legend()
plt.xlabel('n')
plt.ylabel('Absolute error')
plt.title('Error using Gaussian Quadrature')
plt.show()
# -
# As we can after n=13 the integral using gaussian quadrature is matching with the exact integral upto machine precision. The oscillation towards the end are correspondingly the points closer to the lowest value representable in the machine or 0.
|
me16b077_5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Model features
# - augmentation (6 image generated)
NAME = 'augmentation'
# + _cell_guid="38d397d1-860d-4ca9-8b0b-9fe2113347f1" _kg_hide-output=false _uuid="20e67fe0e36a283ab6e899c9d45693e608dd5bdd"
import sys
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import cv2
import random
from tqdm import tqdm
from matplotlib import pyplot as plt
from sklearn.model_selection import train_test_split
import keras
from keras.applications.vgg16 import VGG16
from keras.models import Model
from keras.layers import Dense, Dropout, Flatten
# import third-party library
sys.path.append('./my_lib/')
from data_augmentation import DataAugmentation
# -
# import data
csv_train = pd.read_csv('../input/labels.csv')
csv_test = pd.read_csv('../input/sample_submission.csv')
## DEBUG: reduce data size
csv_train = csv_train.head(100)
csv_test = csv_test.head(100)
# + _cell_guid="fe883370-0270-4ad4-a8c0-05984af2900a" _uuid="6af41eb2be0b3d3f92a10a9f8bdf5fd7ae8bf94b"
# read training CSV
csv_train.head(10)
# + _cell_guid="639d3a2e-ef8c-4f27-b1a2-243cd5bc9ca9" _uuid="fc183bdf820d5f5daf0cd263868e1e3e774cb3a8"
# read test csv
csv_test.head(10)
# + _cell_guid="cc0fce21-3b53-42bf-8d0f-291eb020644f" _uuid="0c6118243a97075dd268239e31f7c851a9aac56b"
# Generate Labels
targets_series = pd.Series(csv_train['breed'])
# print(targets_series)
one_hot = pd.get_dummies(targets_series, sparse = True)
labels = np.asarray(one_hot)
n_check = random.randint(0, len(labels)-1)
print(csv_train['breed'][n_check], 'is encoded as', ''.join((str(i) for i in labels[n_check])))
# + _cell_guid="d2da7283-968a-42ec-95e0-2ecc42d9b027" _uuid="c5c96565ee76a1e9f22fcb9fd6f8305c73f05ad4"
im_size = 90
x_train = []
y_train = []
x_test = []
# + _cell_guid="07f84e2f-600a-47ee-9e7e-48ebac184cd5" _uuid="4478950c6500c326586489713b2c74c2ccf798e2"
for i, (f, breed) in enumerate(tqdm(csv_train.values)):
img = cv2.imread('../input/train/{}.jpg'.format(f))
x_train.append(cv2.resize(img, (im_size, im_size)))
y_train.append(labels[i])
# -
# Using the __stratify__ parameter on __treain_test_split__ the split should be equally distributed per classes.
x_train, x_valid, y_train, y_valid = train_test_split(x_train, y_train,
test_size=0.10, random_state=42,
#stratify=y_train
)
# Use external module to execute data augmentation.
# The module execute:
# - [ ] Inversion
# - [ ] Sobel derivative
# - [ ] Scharr derivative
# - [ ] Laplacian <!--**(error not used for now)**-->
# - [ ] Blur
# - [ ] Gaussian blur [disable]
# - [ ] Median blur
# - [ ] Bilateral blur
# - [x] Horizontal flips
# - [x] Rotation
# +
for i, images in enumerate(tqdm(DataAugmentation(x_train,
options={'horizontal_flips': True,
'rotation': True,
# 'rotation_config': [(10,1.2)],
}))):
for image in images:
if i == 4:
plt.imshow(image, cmap = 'gray', interpolation = 'bicubic')
plt.show()
x_train.append(image)
y_train.append(y_train[i])
print('dataset became:', len(x_train))
# + _cell_guid="7fd00196-2c2a-4264-b36c-10351dfc6781" _uuid="002beaed2ff53c46df2e47f3051ab19225c13f91"
# check train
n_check = random.randint(0, len(y_train)-1)
print('label:', ''.join((str(i) for i in y_train[n_check])))
plt.imshow(x_train[n_check], cmap = 'gray', interpolation = 'bicubic')
plt.show()
# + _cell_guid="bad58384-b970-4820-b5d1-168c31d6b863" _uuid="036e033fbb2f962bdcf03723d85fd0658a3635ce"
for f in tqdm(csv_test['id'].values):
img = cv2.imread('../input/test/{}.jpg'.format(f))
x_test.append(cv2.resize(img, (im_size, im_size)))
# -
# build np array and normalise them
x_train_raw = np.array(x_train, np.float32) / 255.
y_train_raw = np.array(y_train, np.uint8)
x_valid_raw = np.array(x_valid, np.float32) / 255.
y_valid_raw = np.array(y_valid, np.uint8)
x_test_raw = np.array(x_test, np.float32) / 255.
print("x_train shape:", x_train_raw.shape)
print("y_train shape:", y_train_raw.shape)
print("x_train shape:", x_valid_raw.shape)
print("y_train shape:", y_valid_raw.shape)
print("x_test shape:", x_test_raw.shape)
num_classes = y_train_raw.shape[1]
classes = csv_test.columns.values[1:]
# +
# Create the base pre-trained model
base_model = VGG16(weights="imagenet", include_top=False, input_shape=(im_size, im_size, 3))
# Add a new top layer
x = base_model.output
x = Flatten()(x)
predictions = Dense(num_classes, activation='softmax')(x)
# This is the model we will train
model = Model(inputs=base_model.input, outputs=predictions)
# First: train only the top layers (which were randomly initialized)
for layer in base_model.layers:
layer.trainable = False
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
callbacks_list = [
keras.callbacks.ModelCheckpoint('../output/weights.' + NAME + '.{epoch:02d}-{val_loss:.2f}.hdf5',
monitor='val_loss', verbose=0, save_best_only=True, save_weights_only=True,
mode='auto', period=1),
keras.callbacks.EarlyStopping(monitor='val_loss', patience=5, verbose=1)]
model.summary()
# -
history = model.fit(X_train, Y_train, epochs=40, batch_size=48,
validation_data=(X_valid, Y_valid),
callbacks=callbacks_list, verbose=1)
# list all data in history
print(history.history.keys())
# summarize history for accuracy
plt.plot(history.history['acc'])
plt.plot(history.history['val_acc'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
preds = model.predict(x_test_raw, verbose=1)
# save prediction to csv
frame = pd.DataFrame(preds, index=csv_test['id'].tolist(), columns=classes)
frame.to_csv("../output/predicted-{}.csv".format(NAME), index_label='id')
frame.head(10)
# + slideshow={"slide_type": "subslide"}
# check predict
n_check = random.randint(0, len(x_test_raw)-1)
plt.imshow(x_test_raw[n_check], cmap = 'gray_r', interpolation = 'bicubic')
plt.show()
pre = model.predict(np.array([x_test_raw[n_check]]))
arg_max = np.argmax(pre)
print(np.max(pre), arg_max, classes[arg_max])
|
notebook/augmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # How Do the Athletes at University of Nebraska-Lincoln Perform Academically
# ## Context
# The study was originally conducted by National Collegiate Athletic Association (NCAA) to track student-athletes' academic perfomance. The Academic Progress Rate (APR) was introduced to mearsure student-athletes' academic progress. The formula for the APR considers factors including student-athlete eligibility, retention and graduation. the NCAA recognizes the high APR scores indicate student-athletes academic sucess in the classroom, and the low APR scores raise the awareness of the poor performance at the institution. The institutions can be penalized by NCAA for consistently low APR scores.
#
# In this article, we will use the dataset provided by NCAA, and take a look at how athletes at University of Nebraska-Lincoln perform academically comparing to the overall academic scores.
library(dplyr)
library(ggplot2)
library(reshape2)
# ## Overview
scores <- read.csv("NCAA_data.csv")
head(scores)
# +
divisions <- scores %>%
group_by(NCAA_DIVISION) %>%
summarise(Athletes = sum(FOURYEAR_ATHLETES), Scores = mean(FOURYEAR_SCORE),
Eligibility = mean(FOURYEAR_ELIGIBILITY), Retention = mean(FOURYEAR_RETENTION))
divisions
# -
# ## Conference
# +
conferences <- scores %>%
group_by(NCAA_CONFERENCE) %>%
summarise(
Athletes = sum(FOURYEAR_ATHLETES), Scores = mean(FOURYEAR_SCORE),
Eligibility = mean(FOURYEAR_ELIGIBILITY), Retention = mean(FOURYEAR_RETENTION)
) %>% filter(Retention >= .94) %>%
arrange(desc(Scores, Retention))
conferences
# -
ggplot(conferences, aes(x = Scores, y = Retention, size= Athletes)) +
geom_point(alpha=.4, color="Blue") +
scale_size_continuous(range=c(1,12)) +
geom_text(data=subset(conferences,NCAA_CONFERENCE == "Big Ten Conference" | NCAA_CONFERENCE == "Atlantic Coast Conference"|
NCAA_CONFERENCE == "Big 12 Conference"), aes(label=NCAA_CONFERENCE), size=2.8, nudge_x=3, check_overlap=TRUE)
# University of Nebraska-Lincoln, who used to be a part of the Big 12 Conference, is now in the Big Ten Conference. Compare to other conferences, the Big Ten Conference has overall higher scores.
# ## College
# +
schools <- scores %>%
group_by(SCHOOL_NAME) %>%
summarise(
Athletes = sum(FOURYEAR_ATHLETES), Scores = mean(FOURYEAR_SCORE),
Eligibility = mean(FOURYEAR_ELIGIBILITY), Retention = mean(FOURYEAR_RETENTION)
) %>% filter(Eligibility >= .88 & Retention >= .88) %>%
arrange(desc(Scores, Retention))
head(schools)
count(schools)
# -
ggplot(schools, aes(x = Scores, y = Retention, size= Athletes)) +
geom_point(alpha=.3, color="Yellow") +
geom_point(data=subset(schools, SCHOOL_NAME == "University of Nebraska, Lincoln"), alpha=.8, color="Red") +
scale_size_continuous(range=c(1,12)) +
geom_text(data=subset(schools, SCHOOL_NAME == "University of Nebraska, Lincoln"),
aes(label=SCHOOL_NAME), size=3, nudge_x=3, check_overlap=TRUE)
# University of Nebraska-Lincoln has a adequately decent academic performance overall.
# +
sports <- scores %>%
group_by(SPORT_NAME) %>%
summarise(
Athletes = mean(FOURYEAR_ATHLETES), Scores = mean(FOURYEAR_SCORE),
Eligibility = mean(FOURYEAR_ELIGIBILITY), Retention = mean(FOURYEAR_RETENTION)
) %>% arrange(desc(Scores, Retention))
sports
# -
# By grouping sports, we realize that overall, female student-athletes perform better than male student-athletes.
# ## University of Nebraska-Lincoln
# +
unl <-scores %>% filter(SCHOOL_NAME == "University of Nebraska, Lincoln")
year <- unl[,c("SPORT_NAME", "X2014_SCORE", "X2013_SCORE", "X2012_SCORE", "X2011_SCORE",
"X2010_SCORE", "X2009_SCORE", "X2008_SCORE", "X2007_SCORE", "X2006_SCORE",
"X2005_SCORE","X2004_SCORE")]
names(year) <- c("Year", "2014", "2013", "2012", "2011", "2010", "2009", "2008", "2007", "2006", "2005", "2004")
temp <- setNames(data.frame(t(year[,-1])), year[,1])
unl_sports <- data.frame("Year"=rownames(temp),temp)
unl_sports
# -
ggplot(unl_sports, aes(x=Year, colour=Sports, group = 1)) +
geom_point(aes(y=Football, colour="Football")) +
geom_line(aes(y=Football, colour="Football")) +
geom_point(aes(y=Men.s.Basketball, colour="Men's Basketball")) +
geom_line(aes(y=Men.s.Basketball, colour="Men's Basketball")) +
geom_point(aes(y=Women.s.Volleyball, colour="Women's Volleyball")) +
geom_line(aes(y=Women.s.Volleyball, colour="Women's Volleyball")) +
geom_point(aes(y=Baseball, colour="Baseball")) +
geom_line(aes(y=Baseball, colour="Baseball")) +
geom_point(aes(y=Women.s.Soccer, colour="Women's Soccer")) +
geom_line(aes(y=Women.s.Soccer, colour="Women's Soccer")) +
geom_point(aes(y=Women.s.Basketball, colour="Women's Basketball")) +
geom_line(aes(y=Women.s.Basketball, colour="Women's Basketball")) +
labs(x="Year", y="Sports") + scale_y_continuous(limits = c(750,1000)) +
ggtitle("Most Popular Sports' APR Scores In the Past Decade")
# From the line chart above, we can see that among all the popular sports on campus, Men's basketball is the only sport that had APR scores below 900 in the past decade, and it also has higher flutuation compare to other popular sports. Overall, the academic perfomance for all the sports tends to become stablized between 950 and 1000 in recent years, which would be competitive among the entire NCAA.
# ## Conclusion
# As a public university with over 26,000 students on campus, including approximately 2,200 student-athletes, University of Nebraska-Lincoln has above average APR scores in the nation. Football, men's basketball, and women's volleyball are three of the most popular sports UNL campus. Their scores throughout the past decade have been flutuated in certain years.
|
NCAA_Academics_Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Roots: Bracketing Methods
# Bracketing methods determine successively smaller intervals (brackets) that contain a root. When the interval is small enough, then a root has been found. They generally use the intermediate value theorem, which asserts that if a continuous function has values of opposite signs at the end points of an interval, then the function has at least one root in the interval.
#
# Therefore, they require to start with an interval such that the function takes opposite signs at the end points of the interval. However, in the case of polynomials there are other methods for getting information on the number of roots in an interval.
#
# They lead to efficient algorithms for real-root isolation of polynomials, which ensure finding all real roots with a guaranteed accuracy.
# ## GRAPHICAL METHODS
# A simple method for obtaining an estimate of the root of the equation $f (x) = 0$ is to make
# a plot of the function and observe where it crosses the x axis.
# Given this function
#
# $$f(m) = \sqrt{\frac{gm}{c_d}}\tanh(\sqrt{\frac{gc_d}{m}}t) - v(t)$$
#
# We need to find the value of mass due some conditions
import numpy as np
import scipy as sc
import matplotlib.pyplot as plt
# +
# initial conditions
cd = 0.25
g = 9.81
v = 30
t = 5
x = np.linspace(20,50,100)
y = np.sqrt(g*x/cd)*np.tanh(np.sqrt(g*cd/x)*t) - v
# Plot
plt.plot(x,y)
plt.grid(color='k', linestyle='--', linewidth=1)
# -
# The function crosses the m axis between 25 and 30 kg. Visual inspection of the plot
# provides a rough estimate of the root of 28 kg. Assuming that the mass is 28kg, let's see the value of velocity
mass = 28
v_est = np.sqrt(g*mass/cd)*np.tanh(np.sqrt(g*cd/mass)*t)
v_est
# 29.8795 is not 30, right? But it's fine, for now.
#
# Graphical techniques are of limited practical value because they are not very precise.
# However, graphical methods can be utilized to obtain rough estimates of roots. These esti-
# mates can be employed as starting guesses for numerical methods
# ## BRACKETING METHODS AND INITIAL GUESSES
# If you had a roots problem in the days before computing, you’d often be told to use “trial and
# error” to come up with the root.
#
# But, for many other problems, it is preferable to have methods that come up with the
# correct answer automatically. Interestingly, as with trial and error, these approaches require
# an initial “guess” to get started
# ### Incremental Search
# Using the Bolzano theorm, if $f:[a,b]\to \Re$$ ,y = f(x)$ and continuous in the interval
# from $a$ to $b$ and $f(a)$ and $f(b)$ have opposite signs, that is $f(a).f(b) < 0$ then there is at least one real root betwen $[a,b]$
#
# Incremental search methods capitalize on this observation by locating an interval
# where the function changes sign
#
# A problem with an incremental search is the
# choice of the increment length. If the length is too small, the search can be very time
# consuming. On the other hand, if the length is too great, there is a possibility that closely
# spaced roots might be missed (Fig. 5.3). The problem is compounded by the possible exis-
# tence of multiple roots
# Identify brackets within the interval $[3,6]$ for the funciton $f(x) = sin(10x) + cos(3x)$
# +
def incremental_search(func, x_min, x_max, ns):
"""
incsearch: incremental search root locator
xb = incsearch(func,xmin,xmax,ns):
finds brackets of x that contain sign changes of a function on an interval
input:
func = name of function
xmin, xmax = endpoints of interval
ns = number of subintervals
output:
xb(k,1) is the lower bound of the kth sign change
xb(k,2) is the upper bound of the kth sign change
If no brackets found, xb = [].
if nargin < 3, error('at least 3 arguments required'), end
if nargin < 4, ns = 50; end %if ns blank set to 50
"""
# incremental search
x = np.linspace(x_min,x_max,ns)
f = func(x)
nb = 0
xb = np.zeros((x.size,2))
for i in range(0,len(x)-1):
if f[i]*f[i+1] < 0:
nb += 1
xb[nb][0] = x[i]
xb[nb][1] = x[i+1]
if not xb.any():
print("No brackets found")
print("Check interval or increase number of intervals")
else:
print("The number os brackets is: " + str(nb))
print("The solutions are: ")
return xb[1:nb+1]
# Test our function
incremental_search(lambda x: np.sin(10*x)+np.cos(3*x),3,6,50)
# -
# Plot and verify these roots
x = np.linspace(3,6,100)
y = np.sin(10*x)+np.cos(3*x)
plt.plot(x,y)
plt.grid
plt.axhline(y=0, color='r', linestyle='--')
plt.grid(color='k', linestyle='--', linewidth=1)
# The incremental search works fine, but could be missed some multiple roots.
# Just change the value of 'ns' in the code above, for example ns=100, and see what the solution.
def incremental_search_one(f, a, b, dx):
"""
Input:
f: The function to solve
a: The left boundary x-axis value
b: The right boundary x-axis value
dx: The incremental value in searching
Output:
roots: The x-axis value of the root,
interations: Number of iterations used
"""
fa = f(a)
c = a + dx
fc = f(c)
n = 1
while np.sign(fa) == np.sign(fc):
if a >= b:
return a - dx, n
a = c
fa = fc
c = a + dx
fc = f(c)
n += 1
if fa == 0:
return a, n
elif fc == 0:
return c, n
else:
return (a + c)/2., n
# +
y = lambda x: np.sin(10*x) + np.cos(3*x)
root, iterations = incremental_search_one(y, 3, 6, 1/50)
print("Root is: " + str(root))
print("Iterations: "+ str(iterations))
# -
# In this method we have missed some roots.
|
src/02_roots_optimization/01_bracketing_methods/bracketing_methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import graspy
import networkx as nx
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
from graspy.embed import select_dimension, AdjacencySpectralEmbed
from graspy.utils import remove_loops, symmetrize, binarize
from graspy.inference.latent_distribution_test import _median_sign_flips
from graspy.inference import LatentDistributionTest
from hyppo.ksample import KSample
from hyppo._utils import gaussian
import sys
sys.path.append("./")
from seedless_procrustes import *
# %matplotlib inline
print(graspy.__version__)
# -
def convert_classes(to_change):
simple_classes = ['PN', 'APL', 'MBIN', 'MBON', 'KC']
out = []
for string in to_change:
for simple_class in simple_classes:
if simple_class in string:
out.append(simple_class)
return out
# +
G = nx.read_graphml("./data/G.graphml")
G_l = G.subgraph([node for node, data in G.nodes(data=True) if data['Hemisphere'] == 'left'])
A_l = remove_loops(nx.to_numpy_array(G_l))
nodes_l = convert_classes([data['Class'] for node, data in G_l.nodes(data=True)])
G_r = G.subgraph([node for node, data in G.nodes(data=True) if data['Hemisphere'] in ['right', 'center']])
A_r = remove_loops(nx.to_numpy_array(G_r))
nodes_r = convert_classes([data['Class'] for node, data in G_r.nodes(data=True)])
hemispheres = ['Left'] * 163 + ['Right'] * 158
print(len(nodes_l), len(nodes_r))
# +
fig, ax = plt.subplots(1, 2, figsize=(20, 10))
graspy.plot.heatmap(A_l, inner_hier_labels=nodes_l, transform='binarize', hier_label_fontsize=15, sort_nodes=True, ax=ax[0], title="Left")
graspy.plot.heatmap(A_r, inner_hier_labels=nodes_r, transform='binarize', hier_label_fontsize=15, sort_nodes=True, ax=ax[1], title="Right")
# -
A_l_processed = binarize(symmetrize(A_l))
A_r_processed = binarize(symmetrize(A_r))
# +
fig, ax = plt.subplots(1, 2, figsize=(20, 10))
graspy.plot.heatmap(A_l_processed, inner_hier_labels=nodes_l, transform='binarize', hier_label_fontsize=15, sort_nodes=True, ax=ax[0], title="Left")
graspy.plot.heatmap(A_r_processed, inner_hier_labels=nodes_r, transform='binarize', hier_label_fontsize=15, sort_nodes=True, ax=ax[1], title="Right")
# +
num_dims1 = select_dimension(A_l_processed)[0][-1]
num_dims2 = select_dimension(A_r_processed)[0][-1]
n_components = max(num_dims1, num_dims2)
print(n_components)
# -
sns.set_context("talk")
# +
ase = AdjacencySpectralEmbed(n_components=n_components, diag_aug=True, check_lcc=False)
X1_hat = ase.fit_transform(A_l_processed)
X2_hat = ase.fit_transform(A_r_processed)
X1_hat_flipped, X2_hat_flipped = _median_sign_flips(X1_hat, X2_hat)
fig = graspy.plot.pairplot(np.vstack([X1_hat, X2_hat]), labels=hemispheres, title="Without Median Flip")
fig.savefig("flip_withoutgraspy.png", dpi=300, bbox_inches="tight")
fig = graspy.plot.pairplot(np.vstack([X1_hat_flipped, X2_hat_flipped]), labels=hemispheres, title="With Median Flip")
fig.savefig("flip_graspy.png", dpi=300, bbox_inches="tight")
# +
Q = SeedlessProcrustes().fit_predict(X1_hat, X2_hat)
X1_hat_seedless = X1_hat @ Q
fig = graspy.plot.pairplot(np.vstack([X1_hat_seedless, X2_hat]), labels=hemispheres, title="Seedless Flip")
fig.savefig("flip_seedless.png", dpi=300, bbox_inches="tight")
# +
tests = [KSample('Dcorr'), KSample('MGC'), KSample('HSic', compute_distance=gaussian)]
datas = [(X1_hat, X2_hat), (X1_hat_flipped, X2_hat_flipped), (X1_hat_seedless, X2_hat)]
pvals = []
for data in datas:
for test in tests:
res = test.test(*data)
pvals.append(res[1])
pvals = np.array(pvals).reshape(3, -1).T
df = pd.DataFrame(
pvals,
columns=['W/O Flip', 'Median Flip', 'Seedless Flip'],
index=['Dcorr', 'MGC', 'HSIC']
)
# -
df
# +
ase = AdjacencySpectralEmbed(n_components=n_components, algorithm='full', diag_aug=True, check_lcc=False)
X1_hat = ase.fit_transform(A_l_processed)
X2_hat = ase.fit_transform(A_r_processed)
X1_hat_flipped, X2_hat_flipped = _median_sign_flips(X1_hat, X2_hat)
fig = graspy.plot.pairplot(np.vstack([X1_hat, X2_hat]), labels=hemispheres, title="Without Median Flip")
fig.savefig("full_svd_flip_withoutgraspy.png", dpi=300, bbox_inches="tight")
fig = graspy.plot.pairplot(np.vstack([X1_hat_flipped, X2_hat_flipped]), labels=hemispheres, title="With Median Flip")
fig.savefig("full_svd_flip_graspy.png", dpi=300, bbox_inches="tight")
Q = SeedlessProcrustes().fit_predict(X1_hat, X2_hat)
X1_hat_seedless = X1_hat @ Q
fig = graspy.plot.pairplot(np.vstack([X1_hat_seedless, X2_hat]), labels=hemispheres, title="Seedless Flip")
fig.savefig("full_svd_flip_seedless.png", dpi=300, bbox_inches="tight")
# +
tests = [KSample('Dcorr'), KSample('MGC'), KSample('HSic', compute_distance=gaussian)]
datas = [(X1_hat, X2_hat), (X1_hat_flipped, X2_hat_flipped), (X1_hat_seedless, X2_hat)]
pvals = []
for data in datas:
for test in tests:
res = test.test(*data)
pvals.append(res[1])
pvals = np.array(pvals).reshape(3, -1).T
df = pd.DataFrame(
pvals,
columns=['W/O Flip', 'Median Flip', 'Seedless Flip'],
index=['Dcorr', 'MGC', 'HSIC']
)
df
|
ldt/drosophila.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#coding=utf-8
from sqlalchemy import create_engine
import pandas as pd
import cx_Oracle
import os
import matplotlib.pyplot as plt
import numpy as np
plt.rcParams['font.sans-serif'] = ['SimHei']
plt.rcParams['font.serif'] = ['SimHei']
os.environ['NLS_LANG'] = 'AMERICAN_AMERICA.ZHS16GBK'
trafficengine = create_engine('oracle+cx_oracle://gdata:gdata@10.50.0.212:1521/gdata')
# db = cx_Oracle.connect('gdata','gdata','10.50.0.212:1521/gdata')
# print(engine)
trafficsql = """
SELECT
to_char( to_date( xf_tc_countdata.XF_DATE_TIME, 'yyyy-mm-dd' ), 'WW' ) pdate,
round(nvl(
sum(
CASE
WHEN xf_tc_pass.xf_pass_no IN (
1,
2,
3,
4,
5,
6,
7,
8,
9,
10,
11,
12,
13,
14
) THEN
XF_INCOUNT
END
),
0
)/10000,2) AL
FROM
(
SELECT
xf_cameraid,
CASE
WHEN xf_cameraid IN ( 'GHGY5500033230', 'GHGY5500033196', 'GHGY5500033203', 'GHGY5500033197' )
AND XF_DATE_TIME > '2021-01-27' THEN
xf_cameraid || '02' ELSE xf_cameraid || '01'
END AS xf_vcameraid,
xf_startyear,
xf_startmonth,
xf_startday,
xf_starthour,
xf_startminute,
xf_endyear,
xf_endmonth,
xf_endday,
xf_endhour,
xf_endminute,
xf_incount,
xf_outcount,
xf_remarks,
xf_date_time,
xf_date_time_use
FROM
xf_tc_countdata
WHERE
XF_DATE_TIME BETWEEN '2021-01-01'
AND '2021-05-31'
) xf_tc_countdata,
xf_tc_pass
WHERE
xf_tc_countdata.xf_vcameraid = xf_tc_pass.xf_vcameraid
AND
xf_tc_countdata.XF_DATE_TIME BETWEEN '2021-01-01'
AND '2021-05-31'
GROUP BY
to_char( to_date( xf_tc_countdata.XF_DATE_TIME, 'yyyy-mm-dd' ), 'WW' ) order by 1
"""
saleengine = create_engine('oracle+cx_oracle://gl_mis:garland123321mis@10.10.10.20:1521/garland')
salesql = """
SELECT
to_char( to_date( gw_txdate, 'yyyy-mm-dd' ), 'WW' ) pdate,
round(sum( GW_SALESAMOUNT )/10000,2) money
FROM
gw_transsalestotal
WHERE
gw_txdate BETWEEN '2021-01-01'
AND '2021-05-31'
GROUP BY
to_char( to_date( gw_txdate, 'yyyy-mm-dd' ), 'WW' )
ORDER BY 1
"""
trafficdf = pd.read_sql_query(trafficsql, trafficengine)
saledf = pd.read_sql_query(salesql, saleengine)
datax = saledf['pdate']
# datax
datatraffic = trafficdf['al']
# datatraffic
datasale = saledf['money']
# datasale
fig = plt.figure()
# plt.rcParams['font.wqy-microhei']=['WenQuanYi Micro Hei Mono'] #用来正常显示中文标签
# plt.rcParams['axes.unicode_minus']=False #用来正常显示负号
ax1 = fig.add_subplot(111)
ax1 .plot(datax,datatraffic,'-ro',label='pflow')
# ax1.bar(datax,datatraffic,width=0.6,color='r', label='sale')
ax2 = ax1.twinx()
ax2.plot(datax,datasale,'-bo', label='sale')
# ax2.bar(datax,datasale,width=0.6,color='r', label='sale')
fig.legend(loc=1, bbox_to_anchor=(1,1), bbox_transform=ax1.transAxes)
ax1.set_xlabel("日期(周)")
ax1.set_ylabel("客流(人)")
ax2.set_ylabel("销售(元)")
# plt.savefigg()
# plt.plot(datax,datatraffic,"bo-",linewidth=2,markersize=8,label="pflow")
# plt.plot(datax,datasale,"ro-",linewidth=2,markersize=8,label="sale")
# plt.xlabel(r"week")
# plt.ylabel(r"nums")
# plt.legend(loc="upper left")
# import matplotlib
# matplotlib.matplotlib_fname()
# -
|
docker/work/datascience/trafficsSales.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Qiskit config
# +
from qiskit import IBMQ, QuantumCircuit, ClassicalRegister, QuantumRegister, execute, Aer, transpile
from qiskit.test.mock import FakeMontreal, FakeMumbai
from qiskit.providers.aer.backends import AerSimulator
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q-research') # 'ibm-q'
backend_0 = Aer.get_backend('qasm_simulator')
backend_1 = provider.get_backend('ibmq_qasm_simulator')
backend_2 = provider.get_backend('ibmq_santiago')
backend_3 = provider.get_backend('ibmq_manila')
backend_4 = provider.get_backend('ibmq_casablanca')
backend_5 = provider.get_backend('ibmq_jakarta')
backend_6 = FakeMontreal()
backend_7 = FakeMumbai()
shots = 8192
"""
Select the backends that will be compared.
"""
backends = [ backend_0, backend_7 ]
# -
# # Experiment procedures
# +
import numpy as np
import sys
sys.path.append('../../')
from qclib.state_preparation.schmidt import initialize
def measurement(circuit, c):
n = len(c)
circuit.measure(list(range(n)), c)
job = execute(circuit, backend, shots=shots) # , optimization_level=3)
counts = job.result().get_counts(circuit)
v = sum(counts.values())
counts2 = {}
for m in range(2**n):
pattern = '{:0{}b}'.format(m, n)
if pattern in counts:
counts2[pattern] = counts[pattern]
else:
counts2[pattern] = 0.0
return { key : value/v for (key, value) in counts2.items() }
def run_circuit(state, r=0):
circuit = initialize(state, r)
n = int(np.log2(len(input_state)))
c = ClassicalRegister(n)
circuit.barrier()
circuit.add_register(c)
prob = measurement(circuit, c)
return np.array([val for key, val in prob.items()])
def get_state_vector(circuit):
backend = AerSimulator()
tcircuit = transpile(circuit, backend)
tcircuit.save_statevector()
state_vector = backend.run(tcircuit).result().get_statevector()
return state_vector
def plot(r, mae):
# libraries
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# Data
df=pd.DataFrame({ 'x_values': r, 'mae': mae })
# multiple line plots
plt.plot( 'x_values', 'mae' , data=df, marker='o', markersize=8)
plt.xticks(r) # force integer on x axis
# show legend
plt.legend()
# config
#plt.legend(fontsize=12)
#plt.xticks(r, fontsize=14)
#plt.xlabel('s', fontsize=16)
#plt.gcf().set_dpi(200)
# show graph
plt.show()
def plot_result(n, result):
r_range = [r[0] for r in result[n]]
mae = [r[1] for r in result[n]]
l2n = [r[2] for r in result[n]]
#plot(r_range, mae)
plot(r_range, l2n)
# -
# # State Vector Experiment
# +
reps = 40
min_n = 3
max_n = 8
rnd = np.random.RandomState(42)
result2 = {}
for n in range(min_n, max_n+1):
print('\nn = {0}'.format(n))
result2[n] = []
for r in [2**m for m in range(n//2+1)][::-1]: # iso:{1<=rank<2**(n_qubits//2)};uni:{rank=2**(n_qubits//2)}
probs2 = []
ideals2 = []
for i in range(reps):
input_state = rnd.rand(2**n) + rnd.rand(2**n) * 1j
input_state = input_state/np.linalg.norm(input_state)
circuit = initialize(input_state, low_rank = r)
probs2.append(get_state_vector(circuit))
ideals2.append( input_state )
diff = np.mean( np.array(probs2) - np.array(ideals2), axis=0 )
mae = np.sum(np.abs( diff )) # / (2**n)
l2n = np.linalg.norm( diff ) # l2 norm (euclidian distance)
result2[n].append([r, mae, l2n])
print('\trank = {0}\tMAE = {1:.6f}\tMAE norm = {2:.6f}\tL2 norm = {3:.6f}'.format(r, mae, mae/(2**n), l2n))
# -
for n in range(min_n, max_n+1):
plot_result(n, result2)
# # Measurement Experiment
# +
reps = 10
min_n = 3
max_n = 8
rnd = np.random.RandomState(42)
result = {}
for j, backend in enumerate(backends):
backend_name = backend.name()
backend_config = backend.configuration()
backend_qubits = backend_config.n_qubits
print('\nExperiments using {0} backend, with {1} qubits available.'.format(backend_name, backend_qubits))
result[backend_name] = {}
for n in range(min_n, max_n+1):
print('\nn = {0}'.format(n))
result[backend_name][n] = []
for r in [2**m for m in range(n//2+1)][::-1]: # iso:{1<=rank<2**(n_qubits//2)};uni:{rank=2**(n_qubits//2)}
probs = []
probs2 = []
ideals = []
for i in range(reps):
input_state = ( ((1.0 - 0.001) * rnd.rand(2**n) + 0.001) +
((1.0 - 0.001) * rnd.rand(2**n) + 0.001) * 1j )
input_state = input_state/np.linalg.norm(input_state)
probs.append( run_circuit(input_state, r) )
ideals.append( np.power(np.abs(input_state), 2) )
circuit = initialize(input_state, low_rank = r)
probs2.append(get_state_vector_prob(circuit))
mae = np.sum(np.abs( np.mean( np.array(probs) - np.array(ideals), axis=0 ) )) / (2**n)
result[backend_name][n].append([r, mae])
print('\trank = {0}\tMAE = {1}\tMAE_r{0}/MAE_r{2} = {3}'.format(r,
mae,
result[backend_name][n][0][0],
mae/result[backend_name][n][0][1]))
# -
for j, backend in enumerate(backends):
backend_name = backend.name()
print('\n',backend_name,'\n')
for n in range(min_n, max_n+1):
plot_result(n, result[backend_name])
|
example/schmidt/Mean Absolute Error.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from urllib.parse import urlencode
from urllib.request import Request, urlopen
import json
from hashlib import md5
import time;
import math;
def URLRequest(url, params, headers={}):
data = urlencode(params).encode();
request = Request(url, data, headers)
res = urlopen(request).read().decode()
return json.loads(res);
def youdao_trans(text):
url = "http://fanyi.youdao.com/translate_o?smartresult=dict&smartresult=rule"
verify = "ebSeFb%=XZ%T[KZ)c(sy!"
salt = str(math.floor(time.time() * 1000))
client = "fanyideskweb"
sign = md5((client + text + salt + verify).encode('utf-8')).hexdigest()
params = {
'i': text,
'from': 'AUTO',
'to': 'AUTO',
'smartresult': 'dict',
'client': client,
'salt': salt,
'sign': sign,
'doctype': 'json',
'version': '2.1',
'keyfrom': 'fanyi.web',
'action': 'FY_BY_CLICKBUTTION',
'typoResult': 'false'
}
try:
json = URLRequest(url , params, {
'Referer': url,
'User-Agent':'Mozilla/4.0 (compatible; MSIE 9.0; Windows NT 6.1)',
'Cookie': 'YOUDAO_MOBILE_ACCESS_TYPE=1; OUTFOX_SEARCH_USER_ID=-123456789@0.0.0.0'
});
except Exception:
return "翻译失败,网络异常"
try:
return handleResult(json)
except Exception as e:
raise e
return "翻译失败,API异常"
def handleResult(res):
res = res['translateResult'];
res = list(map(lambda line:list(map(lambda block:block['tgt'],line)), res))
res = list(map(lambda line:' '.join(line), res))
res = '\n'.join(res);
return res;
def main(text):
info = youdao_trans(text)
sys.stdout.write(info)
import os
import sys
if __name__ == "__main__":
if os.environ['_'].endswith('ipython'):
text = "Email verification helps our support team verify ownership if you lose account access and allows you to receive all the notifications you ask for."
text = text + "\nEmail verification helps our support team verify ownership if you lose account access and allows you to receive all the notifications you ask for."
else:
text = ' '.join(sys.argv[1:])
main(text);
# -
|
yd.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img alt="QuantRocket logo" src="https://www.quantrocket.com/assets/img/notebook-header-logo.png">
# # Trade inspection
#
# This notebook shows how to inspect individual trades.
#
# First, we can load the transactions and identify a trade to look at. Here we load transactions for a particular month:
# +
from quantrocket.zipline import ZiplineBacktestResult
result = ZiplineBacktestResult.from_csv("sell_gap_backtest_results.csv")
transactions = result.transactions
# Set UTC to America/New_York time for convenience
transactions = transactions.tz_convert("America/New_York").tz_localize(None)
transactions.loc["2020-02"]
# -
# Let's pick a single stock, TTWO:
# +
SID = "FIBBG000BS1YV5"
DATE = "2020-02-07"
trade = transactions[transactions.symbol.str.contains(SID)].loc[DATE]
trade
# -
# We use the data object to load minutes prices for this sid and date, looking back 390 minutes from the session close to get the entire trading day:
# +
from zipline.research import get_data, sid
data = get_data(f"{DATE} 16:00:00")
minute_prices = data.history(sid(SID), "close", 390, "1m")
# Zipline timestamps are in UTC, convert to New York time for convenience
minute_prices.index = minute_prices.index.tz_convert("America/New_York").tz_localize(None)
# -
# Then we plot the minute prices and add trade markers for our buy and sell transactions:
# +
# Plot minute prices
ax = minute_prices.plot(title=f"Buy and sell transactions for {SID} on {DATE}")
# Add the trade markers
trade[trade.amount<0].price.plot(ax=ax, marker="v", color="red", label="Sell")
trade[trade.amount>0].price.plot(ax=ax, marker="^", color="green", label="Buy")
ax.legend()
# -
# ***
#
# ## *Next Up*
#
# Part 7: [Broker-Specific Steps for Live Trading](Part7-Broker-Specific-Steps.ipynb)
|
sell_gap/Part6-Trade-Inspection.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: wino
# language: python
# name: wino
# ---
# +
import pandas as pd
from pathlib import Path
from bs4 import BeautifulSoup as soup
from pprint import pprint as pp
from itertools import tee
import scrapy
from fire import Fire
from scrapy.crawler import CrawlerProcess
DATA = Path.cwd() / 'data'
PAGE = '.gr__wine_com_br'
FICHA = '.TechnicalDetails'
URL = 'https://www.wine.com.br/vinhos/casillero-reserva-limited-edition-cabernet-sauvignon/prod23130.html'
df = pd.read_csv(DATA / 'wines.csv')
TAG = '''<div class="TechnicalDetails">\n<div class="container visible-xs">\n<div class="row somelier">\n<div class="col-xs-12">
<h2 class="somelier__title">Comentário do Sommelier</h2>
<div class="somelier__description">\nEdição especial para o Halloween, esse Cabernet Sauvignon além de apresentar um rótulo ousado e divertido, também traz a elegância de um belo exemplar Concha Y Toro.O Casillero Del Diablo é um tinto frutado, equilibrado e saboroso! Uma edição limitada!\n</div>\n</div>\n
<div class="col-xs-12 content-space">\n</div>\n</div>\n</div>\n<div class="container">\n
<div class="row">\n<div class="col-xs-12 col-md-8 col-md-offset-4 col-sm-7 col-sm-offset-5">\n
<h2 class="TechnicalDetails-title">\nFicha Técnica\n </h2>\n</div>\n</div>\n<div class="TechnicalDetails-image">\n
<script src="https://ajax.cloudflare.com/cdn-cgi/scripts/7089c43e/cloudflare-static/rocket-loader.min.js" data-cf-settings="30065b171190d5cba9ad9c02-|49"></script><img title="Casillero Reserva Limited Edition Cabernet Sauvignon" alt="Casillero Reserva Limited Edition Cabernet Sauvignon" src="https://www.wine.com.br/cdn-cgi/image/f=auto,h=900,q=100/assets-images/produtos/23130-03.png" onerror="this.parentElement.parentElement.classList.add(\'js-wine-kit\')">\n</div>\n
<div class="row">\n<div class="col-xs-12 col-md-8 col-md-offset-4 col-sm-7 col-sm-offset-5">\n
<div class="row">\n<div class="col-xs-12 col-sm-6">\n<div class="row">\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--grape">\n
<div class="Left">\n<i class="Icon Icon--grapes"></i>\n</div>\n
<div class="Right">\n<dt>Tinto</dt>\n<dd>Cabernet Sauvignon (100.00%)</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--location">\n
<div class="Left">\n<i class="Icon Icon--location"></i>\n</div>\n
<div class="Right">\n<dt>Chile</dt>\n<dd>Valle Central </dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--winery">\n
<div class="Left">\n<i class="Icon Icon--winery"></i>\n</div>\n
<div class="Right">\n<dt>Vinícola</dt>\n<dd>Concha y Toro</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--alcoholic_strength">\n
<div class="Left">\n<i class="Icon Icon--wine-and-cup"></i>\n</div>\n
<div class="Right">\n<dt>Teor Alcoólico</dt>\n<dd>13.50% ABV</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--ageing">\n
<div class="Left">\n<i class="Icon Icon--ageing"></i>\n</div>\n
<div class="Right">\n<dt>Amadurecimento</dt>\n<dd>Em barricas de carvalho.</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--harvest">\n
<div class="Left">\n<i class="Icon Icon--harvest"></i>\n</div>\n
<div class="Right">\n<dt>Safra</dt>\n<dd>2017</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--classification">\n
<div class="Left">\n<i class="Icon Icon--classification"></i>\n</div>\n
<div class="Right">\n<dt>Classificação</dt>\n<dd>Seco</dd>\n</div>\n</div>\n</div>\n</div>\n
<div class="col-xs-12 col-sm-6">\n<div class="row">\n<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--appearance">\n
<div class="Left">\n<i class="Icon Icon--appearance"></i>\n</div>\n
<div class="Right">\n<dt>Visual</dt>\n <dd>Rubi intenso</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--aroma">\n
<div class="Left">\n<i class="Icon Icon--aroma"></i>\n</div>\n
<div class="Right">\n<dhttps://www.wine.com.br/vinhos/casillero-reserva-limited-edition-cabernet-sauvignon/prod23130.htmlt>Olfativo</dt>\n<dd>Cassis, cereja, ameixa e notas de tostado</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--taste">\n
<div class="Left">\n<i class="Icon Icon--taste"></i>\n</div>\n
<div class="Right">\n<dt>Gustativo</dt>\n<dd>Corpo médio, taninos sedosos e final longo e frutado.</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--temperature">\n
<div class="Left">\n<i class="Icon Icon--temperature"></i>\n</div>\n
<div class="Right">\n<dt>Temperatura de serviço</dt>\n<dd>15 °C</dd>\n</div>\n</div>\n
<div class="col-xs-8 col-sm-12 TechnicalDetails-description TechnicalDetails-description--temperature">\n
<div class="Left">\n<i class="Icon Icon--ageing_potential"></i>\n</div>\n
<div class="Right">\n<dt>Potencial de guarda</dt>\n<dd>4 anos</dd>\n</div>\n</div>\n</div>\n</div>\n</div>\n</div>\n</div>\n
<div class="row">\n<div class="col-xs-12 col-md-8 col-md-offset-4 col-sm-7 col-sm-offset-5">\n
<article class="TechnicalDetails-matching">\n<dt>Harmonização</dt>\n<dd>Carnes vermelhas, pratos condimentados e queijos envelhecidos como Gruyère e azuis.</dd>\n</article>\n</div>\n</div>\n</div>\n</div>
'''
# +
TIPOS = {'Branco', 'Espumante', 'Frisante', 'Licoroso', 'Rosé', 'Tinto'}
PAISES = {'África do Sul', 'Alemanha', 'Argentina', 'Austrália', 'Brasil', 'Chile'
'China', 'Espanha', 'Estados Unidos', 'França', 'Hungria', 'Itália', 'Líbano'
'Nova Zelândia', 'Portugal', 'Uruguai', 'Grécia', 'Marrocos'}
KEYS = {'Vinícola', 'Teor_Alcoólico', 'Amadurecimento', 'Safra','Classificação', 'Visual',
'Olfativo', 'Gustativo', 'Temperatura', 'Potencial_Guarda', 'Decantação''Harmonização'}
# -
#export
class FichaTecnica(scrapy.Spider):
name = "ficha_tecnica"
start_urls = df['link'].to_list()
def parse(self, response):
page = response.xpath('/html').get()
yield self.ficha_tecnica(page)
# Second parsing method
def ficha_tecnica(self, tag):
tag = soup(tag, 'lxml')
result = {}
key = []
val = []
v = tag.find(class_="PageHeader-title")
key.append("title")
val.append(v.string if v else '')
v = tag.find(class_="somelier__description")
key.append('somelier')
val.append(v.string.strip() if v else '')
precos = tag.find_all(class_='Price-raw')
if len(precos) >= 2:
precos = sorted(list(set([float(p.string) for p in precos])))
key.append('Preço_Sócio')
val.append(precos[0])
key.append('Preço_Normal')
val.append(precos[1])
keys = [t.string for t in tag.find_all('dt')]
vals = [t.string for t in tag.find_all('dd')]
for k,v in zip(keys, vals):
if k in TIPOS:
key.append('Tipo')
val.append(k)
elif k in PAISES:
key.append('Origem')
val.append(f'{k}-{v}')
else:
key.append(k)
val.append(v)
avaliação = tag.find("evaluation-tag")
# print(f"Avaliação: {avaliação.attrs}")
if avaliação:
key.append("Pontuação")
val.append(float(avaliação[':evaluation']))
rating = tag.find('a', class_='Rating-count', string=True)
if rating:
key.append("Avaliações")
rating = rating.string.replace("(", "")
rating = rating.replace(")", "")
val.append(rating)
return dict(zip(key, val))
#try:
# ficha.remove("Ficha Técnica")
#except:
# pass
#field, description = tee(ficha, 2)
#field, description = list(field)[::2], list(description)[1::2]
#return {f:d for f,d in zip(field, description)}
if __name__ == "__main__":
process = CrawlerProcess()
process.crawl(FichaTecnica)
process.start()
|
nbs/wine_extractor.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow-GPU
# language: python
# name: tensorflow
# ---
# +
# Importing Libraries
import tensorflow as tf
import os
import numpy as np
import sys
import random
import cv2
import glob
import matplotlib.pyplot as plt
import xlrd
import tensorflow.keras.backend as K
from tqdm import tqdm
from skimage.io import imread, imshow
from skimage.transform import resize
# -
seed = 42
np.random.seed = seed
# Input Image Dimensions
IMG_WIDTH = 256
IMG_HEIGHT = 256
IMG_CHANNELS = 3
# +
# Extracting the label data from excel sheet
path = ('dataset/CamVid/class_dict.xlsx')
labelNames = []
rgbColorCodes = []
wb = xlrd.open_workbook(path)
sheet = wb.sheet_by_index(0)
sheet.cell_value(0, 0)
for i in range(1, sheet.nrows):
labelNames.append(sheet.cell_value(i, 0))
# print(len(labelNames))
for i in range(1, sheet.nrows):
rgbColorCodes.append((int(sheet.cell_value(i, 1)), int(sheet.cell_value(i, 2)), int(sheet.cell_value(i, 3))))
# print(len(rgbColorCodes))
# Storing label names and corresponding RGB values in dictionaries
labelID = {i:j for i,j in enumerate(labelNames)}
rgbColorID = {i:j for i,j in enumerate(rgbColorCodes)}
print(labelID)
print(rgbColorID)
# +
def rgb_to_onehot(rgb_image, colormap):
'''Function to one hot encode RGB mask labels
Inputs:
rgb_image - image matrix (eg. 256 x 256 x 3 dimension numpy ndarray)
colormap - dictionary of color to label id
Output: One hot encoded image of dimensions (height x width x num_classes) where num_classes = len(colormap)
'''
num_classes = len(colormap)
# print(num_classes)
shape = rgb_image.shape[:2]+(num_classes,)
# print(type(shape))
encoded_image = np.zeros( shape, dtype=np.int8 )
for i, cls in enumerate(colormap):
encoded_image[:,:,i] = np.all(rgb_image.reshape( (-1,3) ) == colormap[i], axis=1).reshape(shape[:2])
return encoded_image
# +
# Processing training datasets
processing = False
X_train = []
Y_train = []
Y_trainRGB = []
count = 0
if processing:
print('Resizing training images')
for filename in glob.glob('dataset/CamVid/train/*.png'):
img = cv2.imread(filename)
img = cv2.resize(img, (IMG_WIDTH,IMG_HEIGHT), interpolation = cv2.INTER_AREA)
height, width, layers = img.shape
if count == 0:
print(height, width, layers)
print(type(img))
count += 1
size = (width,height)
X_train.append(img)
print(count)
print('Training Data resizing Finished')
print('Resizing training masks')
count = 0
for filename in glob.glob('dataset/CamVid/train_labels/*.png'):
img = imread(filename)
img = resize(img, (IMG_HEIGHT, IMG_HEIGHT), mode='constant', preserve_range=True)
Y_trainRGB.append(img)
img = rgb_to_onehot(img, rgbColorID)
count += 1
Y_train.append(img)
print(count)
print('Training Mask resizing Finished')
X_train = np.array(X_train, dtype='uint8')
print(type(X_train))
np.save('NPY files/trainingNPYfiles/MultiClassTrainingData', X_train)
Y_train = np.array(Y_train, dtype='uint8')
print(type(Y_train))
np.save('NPY files/trainingNPYfiles/MultiClassMasks', Y_train)
Y_trainRGB = np.array(Y_trainRGB, dtype='uint8')
print(type(Y_trainRGB))
np.save('NPY files/trainingNPYfiles/MultiClassMasksRGB', Y_trainRGB)
else:
print('Input image already processed. Modify processing parameter to False to process input images again.')
# +
# Processing Validation datasets
processing = False
X_val = []
Y_val = []
Y_valRGB = []
count = 0
if processing:
print('Resizing validation images')
for filename in glob.glob('dataset/CamVid/val/*.png'):
img = cv2.imread(filename)
img = cv2.resize(img, (IMG_WIDTH,IMG_HEIGHT), interpolation = cv2.INTER_AREA)
height, width, layers = img.shape
if count == 0:
print(height, width, layers)
print(type(img))
count += 1
size = (width,height)
X_val.append(img)
print(count)
print('Validation Data resizing Finished')
print('Resizing Validation masks')
count = 0
for filename in glob.glob('dataset/CamVid/val_labels/*.png'):
img = imread(filename)
img = resize(img, (IMG_HEIGHT, IMG_HEIGHT), mode='constant', preserve_range=True)
Y_valRGB.append(img)
img = rgb_to_onehot(img, rgbColorID)
count += 1
Y_val.append(img)
print(count)
print('Validtion Mask resizing Finished')
X_val = np.array(X_val, dtype='uint8')
print(type(X_val))
np.save('NPY files/valNPYfiles/MultiClassValData', X_val)
Y_val = np.array(Y_val, dtype='uint8')
print(type(Y_val))
np.save('NPY files/valNPYfiles/MultiClassValMasks', Y_val)
Y_valRGB = np.array(Y_valRGB, dtype='uint8')
print(type(Y_valRGB))
np.save('NPY files/valNPYfiles/MultiClassValMasksRGB', Y_valRGB)
else:
print('Input image already processed. Modify processing parameter to False to process input images again.')
# +
X_train = np.load('NPY files/trainingNPYfiles/MultiClassTrainingData.npy')
Y_train = np.load('NPY files/trainingNPYfiles/MultiClassMasks.npy')
Y_trainRGB = np.load('NPY files/trainingNPYfiles/MultiClassMasksRGB.npy')
X_val = np.load('NPY files/valNPYfiles/MultiClassValData.npy')
Y_val = np.load('NPY files/valNPYfiles/MultiClassValMasks.npy')
Y_valRGB = np.load('NPY files/valNPYfiles/MultiClassValMasksRGB.npy')
# +
# Processing testing datasets
processTest = False
count = 0
X_test = []
Y_test = []
if processTest:
print('Resizing testing images')
for filename in glob.glob('dataset/CamVid/test/*.png'):
img = cv2.imread(filename)
img = cv2.resize(img, (IMG_WIDTH,IMG_HEIGHT), interpolation = cv2.INTER_AREA)
height, width, layers = img.shape
if count == 0:
print(height, width, layers)
print(type(img))
count += 1
size = (width,height)
X_test.append(img)
print(count)
print('Testing Data resizing Finished')
print('Resizing Testing masks')
count = 0
for filename in glob.glob('dataset/CamVid/test_labels/*.png'):
img = imread(filename)
img = resize(img, (IMG_HEIGHT, IMG_HEIGHT), mode='constant', preserve_range=True)
count += 1
Y_test.append(img)
print(count)
print('Testing Mask resizing Finished')
# This is done to avoid redundant processing
X_test = np.array(X_test, dtype='uint8')
print(type(X_test))
np.save('NPY files/testingNPYfiles/MultiClassTestingData', X_test)
Y_test = np.array(Y_test, dtype='uint8')
print(type(Y_test))
np.save('NPY files/testingNPYfiles/MultiClassTestingMasks', Y_test)
else:
print('Testing images already processed. Modify processTest parameter to False to process testing images again.')
# +
# Building the U-net model
inputs = tf.keras.layers.Input((IMG_WIDTH, IMG_HEIGHT, IMG_CHANNELS)) # Defining the input layer
s = tf.keras.layers.Lambda(lambda x: x/255)(inputs) # Converting input pixels to floating values
c1 = tf.keras.layers.Conv2D(32, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(s)
c1 = tf.keras.layers.Dropout(0.1)(c1) # To prevent neural net from over fitting
c1 = tf.keras.layers.Conv2D(32, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c1)
p1 = tf.keras.layers.MaxPooling2D((2,2))(c1)
c2 = tf.keras.layers.Conv2D(64, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(p1)
c2 = tf.keras.layers.Dropout(0.1)(c2) # To prevent neural net from over fitting
c2 = tf.keras.layers.Conv2D(64, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c2)
p2 = tf.keras.layers.MaxPooling2D((2,2))(c2)
c3 = tf.keras.layers.Conv2D(128, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(p2)
c3 = tf.keras.layers.Dropout(0.2)(c3) # To prevent neural net from over fitting
c3 = tf.keras.layers.Conv2D(128, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c3)
p3 = tf.keras.layers.MaxPooling2D((2,2))(c3)
c4 = tf.keras.layers.Conv2D(256, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(p3)
c4 = tf.keras.layers.Dropout(0.2)(c4) # To prevent neural net from over fitting
c4 = tf.keras.layers.Conv2D(256, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c4)
p4 = tf.keras.layers.MaxPooling2D((2,2))(c4)
c5 = tf.keras.layers.Conv2D(512, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(p4)
c5 = tf.keras.layers.Dropout(0.3)(c5) # To prevent neural net from over fitting
c5 = tf.keras.layers.Conv2D(512, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c5)
p5 = tf.keras.layers.MaxPooling2D((2,2))(c5)
c6 = tf.keras.layers.Conv2D(1024, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(p5)
c6 = tf.keras.layers.Dropout(0.3)(c6) # To prevent neural net from over fitting
c6 = tf.keras.layers.Conv2D(1024, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c6)
# Decoding the layers
u7 = tf.keras.layers.Conv2DTranspose(512, (2,2), strides=(2,2), padding='same')(c6)
u7 = tf.keras.layers.concatenate([u7, c5])
c7 = tf.keras.layers.Conv2D(512, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(u7)
c7 = tf.keras.layers.Dropout(0.2)(c7) # To prevent neural net from over fitting
c7 = tf.keras.layers.Conv2D(512, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c7)
u8 = tf.keras.layers.Conv2DTranspose(256, (2,2), strides=(2,2), padding='same')(c7)
u8 = tf.keras.layers.concatenate([u8, c4])
c8 = tf.keras.layers.Conv2D(256, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(u8)
c8 = tf.keras.layers.Dropout(0.2)(c8) # To prevent neural net from over fitting
c8 = tf.keras.layers.Conv2D(256, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c8)
u9 = tf.keras.layers.Conv2DTranspose(128, (2,2), strides=(2,2), padding='same')(c8)
u9 = tf.keras.layers.concatenate([u9, c3])
c9 = tf.keras.layers.Conv2D(128, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(u9)
c9 = tf.keras.layers.Dropout(0.1)(c9) # To prevent neural net from over fitting
c9 = tf.keras.layers.Conv2D(128, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c9)
u10 = tf.keras.layers.Conv2DTranspose(64, (2,2), strides=(2,2), padding='same')(c9)
u10 = tf.keras.layers.concatenate([u10, c2])
c10 = tf.keras.layers.Conv2D(64, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(u10)
c10 = tf.keras.layers.Dropout(0.1)(c10) # To prevent neural net from over fitting
c10 = tf.keras.layers.Conv2D(64, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c10)
u11 = tf.keras.layers.Conv2DTranspose(32, (2,2), strides=(2,2), padding='same')(c10)
u11 = tf.keras.layers.concatenate([u11, c1], axis=3)
c11 = tf.keras.layers.Conv2D(32, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(u11)
c11 = tf.keras.layers.Dropout(0.1)(c11) # To prevent neural net from over fitting
c11 = tf.keras.layers.Conv2D(32, (3,3), activation='relu', kernel_initializer='he_normal', padding='same')(c11)
outputs = tf.keras.layers.Conv2D(32, (1,1), activation='softmax', padding='same')(c11)
model = tf.keras.Model(inputs=[inputs], outputs=[outputs])
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['accuracy'])
model.summary()
# +
# Model Checkpoint
checkpointer = tf.keras.callbacks.ModelCheckpoint('model_for_nuclei.h5', verbose=1, save_best_only=True)
callbacks = [
tf.keras.callbacks.EarlyStopping(patience=2, monitor='val_loss'),
tf.keras.callbacks.TensorBoard(log_dir='logs')]
# -
results = model.fit(X_train, Y_train, validation_split=0.1, batch_size=5, epochs=100,callbacks=callbacks)
# +
X_test = np.load('NPY files/testingNPYfiles/MultiClassTestingData.npy')
Y_test = np.load('NPY files/testingNPYfiles/MultiClassTestingMasks.npy')
# -
preds_test = model.predict(X_test, verbose=1)
print(preds_test.shape)
preds_train = model.predict(X_train, verbose=1)
preds_val = model.predict(X_val, verbose=1)
# +
def onehot_to_rgb(onehot, colormap):
'''Function to decode encoded mask labels
Inputs:
onehot - one hot encoded image matrix (height x width x num_classes)
colormap - dictionary of color to label id
Output: Decoded RGB image (height x width x 3)
'''
single_layer = np.argmax(onehot, axis=-1)
output = np.zeros( onehot.shape[:2]+(3,) )
for k in colormap.keys():
output[single_layer==k] = colormap[k]
return np.uint8(output)
preds_testRGB = []
for i in range(len(X_test)):
preds_testRGB.append(onehot_to_rgb(preds_test[i], rgbColorID))
preds_valRGB = []
for i in range(len(X_val)):
preds_valRGB.append(onehot_to_rgb(preds_val[i], rgbColorID))
preds_trainRGB = []
for i in range(len(X_train)):
preds_trainRGB.append(onehot_to_rgb(preds_train[i], rgbColorID))
# +
preds_testRGB = np.array(preds_testRGB, dtype='uint8')
np.save('Output/MultiClassTestingDataOutput', preds_testRGB)
preds_valRGB = np.array(preds_valRGB, dtype='uint8')
np.save('Output/MultiClassValDataOutput', preds_valRGB)
preds_trainRGB = np.array(preds_trainRGB, dtype='uint8')
np.save('Output/MultiClassTrainingDataOutput', preds_trainRGB)
# +
# Comparing the results
ix = random.randint(0, len(preds_trainRGB))
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(1,3,1)
ax1.imshow(X_train[ix])
ax1.title.set_text('Actual Training Image')
ax2 = fig.add_subplot(1,3,2)
ax2.set_title('Expected Segmentation Output')
ax2.imshow(Y_trainRGB[ix])
ax3 = fig.add_subplot(1,3,3)
ax3.set_title('U-net Segmented Output')
ax3.imshow(preds_trainRGB[ix])
ix = random.randint(0, len(preds_valRGB))
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(1,3,1)
ax1.imshow(X_val[ix])
ax1.title.set_text('Actual Validation Image')
ax2 = fig.add_subplot(1,3,2)
ax2.set_title('Expected Segmentation Output')
ax2.imshow(Y_valRGB[ix])
ax3 = fig.add_subplot(1,3,3)
ax3.set_title('U-net Segmented Output')
ax3.imshow(preds_valRGB[ix])
ix = random.randint(0, len(preds_testRGB))
fig = plt.figure(figsize=(20,8))
ax1 = fig.add_subplot(1,3,1)
ax1.imshow(X_test[ix])
ax1.title.set_text('Actual Testing Image')
ax2 = fig.add_subplot(1,3,2)
ax2.set_title('Expected Segmentation Output')
ax2.imshow(Y_test[ix])
ax3 = fig.add_subplot(1,3,3)
ax3.set_title('U-net Segmented Output')
ax3.imshow(preds_testRGB[ix])
# +
N = len(results.history['loss'])
#Plot the model evaluation history
plt.style.use("ggplot")
fig = plt.figure(figsize=(20,8))
fig.add_subplot(1,2,1)
plt.title("Training Loss")
plt.plot(np.arange(0, N), results.history["loss"], label="train_loss")
plt.plot(np.arange(0, N), results.history["val_loss"], label="val_loss")
plt.ylim(0, 1)
fig.add_subplot(1,2,2)
plt.title("Training Accuracy")
plt.plot(np.arange(0, N), results.history["accuracy"], label="train_accuracy")
plt.plot(np.arange(0, N), results.history["val_accuracy"], label="val_accuracy")
plt.ylim(0, 1)
plt.xlabel("Epoch #")
plt.ylabel("Loss/Accuracy")
plt.legend(loc="lower left")
plt.show()
|
U-net/MulticlassUnet-Camvid.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="2RjXUbystvxg" colab_type="text"
# # Setup
# **For full setup read all 'NOTE:' comments and that should be enough**
# + [markdown] id="FQ4bfn-vihAW" colab_type="text"
# # NOTE: ctrl+shift+I and paste it to console
# function ConnectButton(){
# console.log("Connect pushed");
# document.querySelector("#top-toolbar > colab-connect-button").shadowRoot.querySelector("#connect").click()
# }
# setInterval(ConnectButton,60000);
# + [markdown] id="3iYof0Eog2bo" colab_type="text"
# NOTE: AFTER RESETING RUNTIME ENABLE GPU ACCELERATION!
# + id="AokMEAep3lbs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 125} outputId="b40bbc78-ccfe-449e-970c-cf04dea29285"
# Mount Drive
from google.colab import drive
drive.mount('/content/drive')
######### GLOBAL VIARIABLES SETUP ##########
##### NOTE: change directories to right ones #####
DIR_DETECTOR = "/content/drive/My Drive/Colab Content/GoldBagDetector/" # Will cd here
DIR_DATASET = "dataset-combined/"
PRETRAINED_MODEL = DIR_DATASET + "models/detection_model-ex-020--loss-0001.557.h5"
# PRETRAINED_MODEL = DIR_DATASET + "models/detection_model-ex-104--loss-2.93.h5"
global_id = 0 # To stop threads ath the end of program it's changed
print("Version should be 3.12.2 or higher. Otherwise upgrade protobuf")
# !pip show protobuf | grep Version
# %cd $DIR_DETECTOR
print("Dir check below. Good is no output:")
import os
if not os.path.exists(DIR_DETECTOR):
print("Path DIR_DETECTOR does not exist")
if not os.path.exists(DIR_DATASET):
print("Path DIR_DATASET does not exist")
if not os.path.exists(PRETRAINED_MODEL):
print("Path PRETRAINED_MODEL does not exist")
# + [markdown] id="1Ixfg69OrdVc" colab_type="text"
# # On Error: __init__() got an unexpected keyword argument 'read_only_collections'
#
# If validation does not work reinstall protobuf and restart runtime
# + id="uGcbJnb-qlKb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 302} outputId="3b298f81-f721-4661-9ee0-6b01f092d8e2"
# !pip uninstall --yes protobuf
# !pip install 'protobuf>=3.0.0a3'
# + [markdown] id="I4BhzLJ6eEgP" colab_type="text"
# Auth *pydrive* and install tensorflow
# + id="HlGQfyedtiQZ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="8fce232e-dde2-4598-d4f5-05fc200251f8"
# %cd $DIR_DETECTOR
# Init Drive autocleaner
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from google.colab import auth
from oauth2client.client import GoogleCredentials
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
my_drive = GoogleDrive(gauth)
# !pip3 install tensorflow-gpu==1.13.1
# !pip3 install imageai --upgrade
import tensorflow as tf
if tf.test.gpu_device_name() != "/device:GPU:0":
raise EnvironmentError("NO GPU ACCELERATION ENABLED")
else:
print("GPU GOOD")
# + [markdown] id="KQ4Llt-nt1pJ" colab_type="text"
# # Training
# + id="F_8qYPEJt3eg" colab_type="code" cellView="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="73041088-9bee-464f-9362-5b26976d12a5"
import tensorflow as tf
if tf.test.gpu_device_name() != "/device:GPU:0":
raise EnvironmentError("NO GPU ACCELERATION ENABLED")
# # %cd /
# %cd $DIR_DETECTOR
import os
import time
import glob
import random
import threading
from imageai.Detection.Custom import DetectionModelTrainer
######## VARIABLES ########
RUN_THREADS = True
JOB_DONE = False
THREAD_SLEEP = 300 # 300 = 5 min
global_id = 0
MODELS_TO_KEEP = 8 # NOTE: change depending on free space on drive
######## FUNCTIONS ########
def get_sorted_models_list():
model_list = glob.glob(DIR_DATASET + "models/*.h5")
# Sort by time modified oldest first
model_list = sorted(model_list, key=os.path.getmtime)
return model_list
def auto_clean_drive_trash():
global my_drive
try:
files_list = my_drive.ListFile({'q': "trashed = true"}).GetList()
print("Trash files to delete ({}): ".format(len(files_list)))
for file in files_list:
print("Trash \"{}\" deleted permanently.".format(file["title"]))
file.Delete()
except Exception:
pass
def manage_model_files(ID):
'''
Keep only best performing models
And autoclean trash
'''
global RUN_THREADS, THREAD_SLEEP, global_id
while True:
model_list = get_sorted_models_list()
# Shuffle worst performers
tmp_list = model_list[:-3]
random.shuffle(tmp_list)
model_list[:-3] = tmp_list
if len(model_list) > MODELS_TO_KEEP:
print("TOO MANY MODELS. LIST:")
print(model_list)
# Delete one bad model (shuffled)
print("Deleting: %s" % (model_list[0]))
os.remove(model_list[0])
# Autoclean Drive trash
auto_clean_drive_trash()
# Sleep
for i in range(THREAD_SLEEP):
if RUN_THREADS is False or ID != global_id:
return
time.sleep(1)
def log_time(ID):
global RUN_THREADS, THREAD_SLEEP, global_id
while True:
print (time.ctime())
filename = "my_log.txt"
if os.path.exists(filename):
append_write = 'a' # append if already exists
else:
append_write = 'w' # make a new file if not
f = open(filename, append_write)
f.write("{}\n{}\n".format(time.ctime(), get_sorted_models_list()))
f.close()
# Sleep
for i in range(THREAD_SLEEP):
if RUN_THREADS is False or ID != global_id:
return
time.sleep(1)
######## MAIN PROGRAM ########
# Run thread for file management
global_id = time.time()
t1 = threading.Thread(target=manage_model_files, kwargs=dict(ID=global_id))
t2 = threading.Thread(target=log_time, kwargs=dict(ID=global_id))
t1.start()
t2.start()
#Train model
##### NOTE: change train_from_pretrained_model to newest model every rerun #####
##### NOTE: change object_names_array= if data class have changed #####
trainer = DetectionModelTrainer()
trainer.setModelTypeAsYOLOv3()
trainer.setDataDirectory(data_directory=DIR_DATASET)
trainer.setTrainConfig(object_names_array=["goldbag"], batch_size=4,
num_experiments=150,
train_from_pretrained_model=PRETRAINED_MODEL)
trainer.trainModel()
print("Execution done. Going to sleep for 1 min")
RUN_THREADS = False
global_id = 0
time.sleep(60)
# + [markdown] id="XjbNsp6vmYla" colab_type="text"
# # Evaluation
# + id="2nONV3s2me-P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1596eddb-c9f1-44e9-9f92-66f67e45b938"
# %cd $DIR_DETECTOR
from imageai.Detection.Custom import DetectionModelTrainer
trainer = DetectionModelTrainer()
trainer.setModelTypeAsYOLOv3()
trainer.setDataDirectory(data_directory=DIR_DATASET)
trainer.evaluateModel(model_path=DIR_DATASET + "models", json_path=DIR_DATASET + "json/detection_config.json", iou_threshold=0.8, object_threshold=0.9, nms_threshold=0.5)
# + [markdown] id="usv22W4z09TI" colab_type="text"
# # Test Model
# + id="nAG5cJcI1DCx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="0fa97ca2-4458-4261-d734-25abd75577b6"
# %cd $DIR_DETECTOR
from imageai.Detection.Custom import CustomObjectDetection
import os
import cv2
from google.colab.patches import cv2_imshow
import glob
import time
import random
detector = CustomObjectDetection()
detector.setModelTypeAsYOLOv3()
detector.setJsonPath(DIR_DATASET + "json/detection_config.json")
models = glob.glob(DIR_DATASET + "models/*.h5")
models = sorted(models, key=os.path.getmtime, reverse=True) # Sort by time modified best first
print("All models: {}".format(models))
validation_imgs = glob.glob(DIR_DATASET + "validation/images/*.jpg")
random.shuffle(validation_imgs)
print("Validation images: {}".format(validation_imgs))
for model in models: # [0:2]:
print("#################################")
print("#################################")
print("##############MODEL##############")
print("Validating model: {}".format(model))
print("#################################")
print("#################################")
print("Showing max 25 random images per model")
detector.setModelPath(model)
detector.loadModel()
count_detections = 0
count_img = 0
for img in validation_imgs: # [:25]:
count_img += 1
frame = cv2.imread(img)
frame_out, detections = detector.detectObjectsFromImage(input_type="array", input_image=frame,
output_type="array", minimum_percentage_probability=10)
save_path = DIR_DATASET + "evaluation-images/{}/{}".format(model[-21:], img[-10:])
print("Save path: {}".format(save_path))
# if not os.path.exists(save_path):
# os.makedirs(save_path)
# cv2.imwrite(frame_out, save_path)
for eachObject in detections:
count_detections += 1
print(eachObject["name"] , " : ", eachObject["percentage_probability"], " : ", eachObject["box_points"] )
print("Detected: {}/{}".format(count_detections, count_img))
cv2_imshow(frame_out)
|
Python/GoldBagDetector/gold_bag_detector_gcolab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Proprietary content. ©Great Learning. All Rights Reserved. Unauthorized use or distribution prohibited
# + [markdown] colab_type="text" id="HyfIOUrSFwLb"
# ### Libraries along with their versions used at the time of making notebook-
# nltk 3.2.5
# + [markdown] colab_type="text" id="Bkhe6TWrxWDL"
# # Basic text reading
# + colab={} colab_type="code" id="jI8I-TDWxWDP"
import nltk # Import NLTK
from nltk.corpus import reuters # Import the Reuters corpus
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1008, "status": "ok", "timestamp": 1581770223019, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="mpqHmferxWDV" outputId="4d7d41fc-0633-43d0-c4fd-bbec5e993ff1"
nltk.download('reuters')
reuters.fileids() # List file-ids in the corpus
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 995, "status": "ok", "timestamp": 1581770247614, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="Etxq9RWbxWDZ" outputId="0ecadc59-a883-474d-8ca5-a163533f59c8"
reuters.categories() # List news categories in the corpus
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1071, "status": "ok", "timestamp": 1581770261619, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="THA37ZJYxWDd" outputId="7dca697a-2ac3-47a1-f867-951171234a52"
reuters.fileids(['wheat','rice']) # List file ids with either wheat or rice categories
# Some file ids may overlap as news covers multiple categories
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1842, "status": "ok", "timestamp": 1581770311010, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="hU8fJauJxWDh" outputId="1b26abc1-2b27-4475-d7aa-34d7045a0651"
# Let us see how many chars, words and sentences are in each file
nltk.download('punkt')
for fileid in reuters.fileids(['wheat','rice']):
num_chars = len(reuters.raw(fileid))
num_words = len(reuters.words(fileid))
num_sents = len(reuters.sents(fileid))
num_vocab = len(set(w.lower() for w in reuters.words(fileid)))
print(fileid, " : ",num_chars, num_words, num_sents, num_vocab)
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" executionInfo={"elapsed": 1226, "status": "ok", "timestamp": 1581770367032, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="gNImGLnwxWDl" outputId="54b692be-f67d-4fe2-9dfd-0eba991cc33a"
# Select one file for futher processing
fileid = 'test/15618'
reuters.raw(fileid) # See what is in the selected file
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1002, "status": "ok", "timestamp": 1581770396128, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="TlImJQZexWDo" outputId="b0b633c0-0584-48fb-956a-85343907547e"
reuters.words(fileid) # See individual words in the selected file
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" executionInfo={"elapsed": 991, "status": "ok", "timestamp": 1581770420555, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="CuTURxcdxWDr" outputId="cdf5b001-0edb-4ac2-bf37-2e6b000cb7dd"
# See sentences in the file. Notice the bracket within bracket for each sentence
reuters.sents(fileid)
# + colab={} colab_type="code" id="FwgOqSnq3Xmd"
# + [markdown] colab_type="text" id="kXqq84XwxWDu"
# # Pre-processing: lower case, tokenization, removing stop words, finding words
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1283, "status": "ok", "timestamp": 1581770476387, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="tdCnQOo-xWDv" outputId="cb52bbf8-f5be-41ad-bcfc-443ff374896e"
# See all the words in the file, lexicographically sorted
set(w.lower() for w in reuters.words(fileid))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1016, "status": "ok", "timestamp": 1581770526724, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="fujdCEOZxWDz" outputId="e2e5b670-89fe-4515-8de0-78e2379774f0"
#Remove stop words
nltk.download('stopwords')
from nltk.corpus import stopwords # Import stop words
wordList = [w for w in reuters.words(fileid) if w.lower() not in stopwords.words('english')]
wordList
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 988, "status": "ok", "timestamp": 1581770578748, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="u6AfvyOCxWD1" outputId="bb158a3c-8afa-49a0-fd8e-aab288eea6ff"
#Tokenize
from nltk import word_tokenize # Tokenize the file, which is similar to getting words
tokens = word_tokenize(reuters.raw(fileid))
wordList = reuters.words(fileid)
tokens
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1016, "status": "ok", "timestamp": 1581770609932, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="JXQs3vf5xWD4" outputId="6aa89460-bb4e-49d2-8023-2c573dca7f26"
# Check out the difference between tokens and words. Tokenization is more intelligence segmentation
wordList[12:20]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1001, "status": "ok", "timestamp": 1581770637540, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="Ub0mI8epxWD6" outputId="2f2bfe46-d485-419d-e6bf-69f5cf0d720b"
# Find position of a word
reuters.raw(fileid).find('MARKET')
# + [markdown] colab_type="text" id="QDTdUj-DxWD9"
# # Synonyms, PoS Tagging, Parsing: Chunking, Chinking, Syntax Trees
# + colab={"base_uri": "https://localhost:8080/", "height": 258} colab_type="code" executionInfo={"elapsed": 1010, "status": "ok", "timestamp": 1581770736550, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="wL3ybta5xWD9" outputId="f6faee8c-784e-45bb-a2ea-ccac447a4f0b"
# Check out some synonyms
nltk.download('wordnet')
from nltk.corpus import wordnet as wn # See the list of synonyms
wn.synsets('trade')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 1254, "status": "ok", "timestamp": 1581770814743, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="OLTWy4hbxWEC" outputId="a2f17fb8-5429-4a1e-cec5-103d4e2b7148"
wn.synset('trade.v.02').lemma_names() # Read one particular synonym
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1792, "status": "ok", "timestamp": 1581770874970, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="EIrd5zFwxWEF" outputId="4a6fbfef-0d75-4a4f-a42a-32ae7ead7308"
# Find text with similar context
text = nltk.Text(word.lower() for file_id in reuters.fileids(['wheat','rice']) for word in reuters.words(file_id))
text.similar('rice')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 987, "status": "ok", "timestamp": 1581770941957, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="MEn5GXINxWEH" outputId="894f318d-983f-442d-baef-6c3a63f328e2"
# See PoS of tokens (for some corpora, POS are already tagged in this corpus)
nltk.download('averaged_perceptron_tagger')
nltk.pos_tag(tokens)
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" executionInfo={"elapsed": 1045, "status": "ok", "timestamp": 1581771265025, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="s3U7kHeCxWEJ" outputId="ea35805f-9ad7-4de5-fa85-a712815f6e96"
# Parsing using regular expression with chunking (without chinking)
# We specify that noun phrase can have a determinant, adverb, gerund verb, or an adjective,
# but it must have a noun or a pronoun, e.g. "A fastly running beautiful deer..."
# The verb phrase should start with a verb, and then it can have anything.
pattern = """NP: {<DT>?<RB.?>?<VBG>?<JJ.?>*(<NN.?>|<PRP.?>)+}
VP: {<VB.?>+<.*>*}
"""
mySentence = 'A fastly running beautiful deer skidded off the road'
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
#myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize('The cat was going to eat bread but then he found a mouse')))
print(myParsedSentence)
# + colab={} colab_type="code" id="JwMuU4Ub1G53" outputId="3d138519-1271-4fd3-e0f0-0511e3014238"
# Displaying a parse (syntax) tree.
# Install Ghostscript: In Anacodna prompt, type 'conda install -c conda-forge ghostscript'
# This cell will not work in Google Colab
from IPython.display import display
display(myParsedSentence)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 978, "status": "ok", "timestamp": 1581771371704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="jzGGIbHwxWEO" outputId="bfdc677c-98ed-4fb7-cfe3-edc62fe26fa0"
# Let us try another sentence with chunking (without chinking)
mySentence = 'I left to do my homework'
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
print(myParsedSentence)
# + colab={} colab_type="code" id="fz1cGEGf1G6A" outputId="391d30ec-8d9c-43be-e59f-e06a9b48e555"
# This cell will not work in Google Colab
display(myParsedSentence)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" executionInfo={"elapsed": 880, "status": "ok", "timestamp": 1581769727331, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="WaOfnDUOxWEW" outputId="53a1e41f-8137-4e18-9c9d-89a35a0ee7a1"
# Parsing using regular expression with chunking and chinking (exclusion rule)
# Redefine pattern with chinking, to exclude "to do something"
pattern = """NP: {<DT>?<RB.?>?<VBG>?<JJ.?>*(<NN.?>|<PRP.?>)+}
VP: {<VB.?>+<.*>*}
}(<VBG>|(<TO><.*>*)){
"""
mySentence = 'I left to do my homework'
myParser = nltk.RegexpParser(pattern)
myParsedSentence = myParser.parse(nltk.pos_tag(nltk.word_tokenize(mySentence)))
print(myParsedSentence)
# + colab={} colab_type="code" id="JoEb8num1G6H" outputId="4c3479b5-616a-423a-ae16-e632ddf28009"
# This cell will not work in Google Colab
display(myParsedSentence)
# + [markdown] colab_type="text" id="2sZO3CZcxWEb"
# # Context free grammar (CFG)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1007, "status": "ok", "timestamp": 1581771616599, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="VgSVCpXzxWEc" outputId="f1e1a03f-a6a6-4b29-df6e-fb0072603207"
# Defining a grammar
from nltk import CFG
myGrammar = nltk.CFG.fromstring("""
S -> NP VP
VP -> VB NP
VP -> VB
VP -> VB PRP
NP -> DET NN
VB -> "chased"|"ate"
DET -> "another"|"the"
NN -> "cat"|"rat"|"snake"
PRP -> "it"
""")
# Generating sentences from the defined grammar
from nltk.parse.generate import generate
for sent in generate(myGrammar):
print(' '.join(sent))
# + [markdown] colab_type="text" id="60Byit-CxWEf"
# # Frequency of words, bi-grams, tri-grams
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1037, "status": "ok", "timestamp": 1581771669765, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="iIVgLK9FxWEg" outputId="a2d84b55-4e5a-4133-e1cd-7bda38321e7a"
# Frequency Distribution of words
fdist = nltk.FreqDist(wordList)
fdist
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1017, "status": "ok", "timestamp": 1581771722867, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="crfq_S2VxWEj" outputId="0f8fefbf-d7fa-4b04-ffe7-deff1ff2505a"
# Some n-gram examples in NLTK
from nltk.util import ngrams
bigrams = ngrams(tokens,2) # A bigram is specified by 2, trigram by 3, etc.
for b in bigrams:
print(b)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1263, "status": "ok", "timestamp": 1581771757405, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="2rp3udYExWEm" outputId="6587e469-e36a-4de4-b8b6-843f6089aeef"
trigrams = ngrams(tokens,3) # A bigram is specified by 2, trigram by 3, etc.
for t in trigrams:
print(t)
# + [markdown] colab_type="text" id="Oa7p8sucxWEp"
# # Stemming, lemmatization
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" executionInfo={"elapsed": 1165, "status": "ok", "timestamp": 1581771890013, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="k7ybqUuYxWEq" outputId="b8123abe-e452-4349-fc19-6e48741e2a68"
# Comparing stemming and lemmatization
# Reading corpus and removing stop words
nltk.download('brown')
from nltk.corpus import brown
fileid = 'ck23'
from nltk.corpus import stopwords # Remove stop words
wordList = [w for w in brown.words(fileid) if w.lower() not in stopwords.words('english')]
import string # Remove punctuation
wordList = [w for w in wordList if w not in string.punctuation]
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 1293, "status": "ok", "timestamp": 1581771954820, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="SJhwY8_ixWEs" outputId="e0b9ba9e-bb41-416e-e106-56ee2139bbf7"
# COMPARE TWO STEMMERS
from nltk.stem import PorterStemmer
from nltk.stem import LancasterStemmer
porter = PorterStemmer()
lancaster = LancasterStemmer()
StemmersCompared = [word+' : '+porter.stem(word)+' : '+lancaster.stem(word) for word in wordList]
StemmersCompared
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" executionInfo={"elapsed": 828, "status": "ok", "timestamp": 1581769771456, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mCX_PdUc-oYjLw3bTTsusGGiLA4h-CJUT94gBbkRw=s64", "userId": "14604457747665098998"}, "user_tz": -330} id="fEBSf6glxWEu" outputId="2964a58c-59c0-4685-a07c-1a50e8dda243"
# Lemmatization compared to Stemming
from nltk.stem import WordNetLemmatizer
wordNet = WordNetLemmatizer()
StemmersCompared = [word+' : '+porter.stem(word)+' : '+lancaster.stem(word)+' : '+wordNet.lemmatize(word) for word in wordList]
StemmersCompared
# + colab={} colab_type="code" id="XCtHZrDC00YQ"
|
.ipynb_checkpoints/NLP basics-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Arbitrage Pricing Theory
# By Evgenia "Jenny" Nitishinskaya and <NAME>
#
# Part of the Quantopian Lecture Series:
#
# * [www.quantopian.com/lectures](https://www.quantopian.com/lectures)
# * [github.com/quantopian/research_public](https://github.com/quantopian/research_public)
#
# Notebook released under the Creative Commons Attribution 4.0 License.
#
# ---
#
# Arbitrage pricing theory is a major asset pricing theory that relies on expressing the returns using a linear factor model:
#
# $$R_i = a_i + b_{i1} F_1 + b_{i2} F_2 + \ldots + b_{iK} F_K + \epsilon_i$$
#
# This theory states that if we have modelled our rate of return as above, then the expected returns obey
#
# $$ E(R_i) = R_F + b_{i1} \lambda_1 + b_{i2} \lambda_2 + \ldots + b_{iK} \lambda_K $$
#
# where $R_F$ is the risk-free rate, and $\lambda_j$ is the risk premium - the return in excess of the risk-free rate - for factor $j$. This premium arises because investors require higher returns to compensate them for incurring risk. This generalizes the capital asset pricing model (CAPM), which uses the return on the market as its only factor.
#
# We can compute $\lambda_j$ by constructing a portfolio that has a sensitivity of 1 to factor $j$ and 0 to all others (called a <i>pure factor portfolio</i> for factor $j$), and measure its return in excess of the risk-free rate. Alternatively, we could compute the factor sensitivities for $K$ well-diversified (no asset-specific risk, i.e. $\epsilon_p = 0$) portfolios, and then solve the resulting system of linear equations.
# ## Arbitrage
#
# There are generally many, many securities in our universe. If we use different ones to compute the $\lambda$s, will our results be consistent? If our results are inconsistent, there is an <i>arbitrage opportunity</i> (in expectation). Arbitrage is an operation that earns a profit without incurring risk and with no net investment of money, and an arbitrage opportunity is an opportunity to conduct such an operation. In this case, we mean that there is a risk-free operation with <i>expected</i> positive return that requires no net investment. It occurs when expectations of returns are inconsistent, i.e. risk is not priced consistently across securities.
#
# For instance, there is an arbitrage opportunity in the following case: say there is an asset with expected rate of return 0.2 for the next year and a $\beta$ of 1.2 with the market, while the market is expected to have a rate of return of 0.1, and the risk-free rate on 1-year bonds is 0.05. Then the APT model tells us that the expected rate of return on the asset should be
#
# $$ R_F + \beta \lambda = 0.05 + 1.2 (0.1 - 0.05) = 0.11$$
#
# This does not agree with the prediction that the asset will have a rate of return of 0.2. So, if we buy \$100 of our asset, short \$120 of the market, and buy \$20 of bonds, we will have invested no net money and are not exposed to any systematic risk (we are market-neutral), but we expect to earn $0.2 \cdot 100 - 0.1 \cdot 120 + 20 \cdot 0.05 = 9$ dollars at the end of the year.
#
# The APT assumes that these opportunities will be taken advantage of until prices shift and the arbitrage opportunities disappear. That is, it assumes that there are arbitrageurs who have sufficient amounts of patience and capital. This provides a justification for the use of empirical factor models in pricing securities: if the model were inconsistent, there would be an arbitrage opportunity, and so the prices would adjust.
|
lectures/drafts/Arbitrage Pricing Theory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# I simulate the photometric variable induced behavior as a random walk process.
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import MultipleLocator
from sklearn import linear_model
from sklearn.metrics import mean_squared_error, r2_score
from statsmodels.iolib.table import (SimpleTable, default_txt_fmt)
# %matplotlib inline
# # %config InlineBackend.figure_format = "svg"
# My progs
from my_progs.vlbi.ts_func import get_ts
np.random.seed(28)
# +
souname = "0552+398"
coordts = get_ts(souname, calc_oft=True)
plotdir = "../plots"
# +
fig, (ax0, ax1) = plt.subplots(nrows=2, sharex=True, sharey=True)
ax0.errorbar(coordts["jyear"], coordts["dra"], yerr=coordts["ra_err"],
fmt="b.", ecolor="grey", elinewidth=0.1, ms=1)
ax1.errorbar(coordts["jyear"], coordts["ddec"], yerr=coordts["dec_err"],
fmt="b.", ecolor="grey", elinewidth=0.1, ms=1)
ax0.xaxis.set_minor_locator(MultipleLocator(1))
ax0.set_ylabel("$\Delta\\alpha\,\cos\delta$ ($\mathrm{mas}$)")
ax1.set_ylabel("$\Delta\delta$ ($\mathrm{mas}$)")
ax1.set_xlabel("Year")
ax1.set_ylim([-5, 5])
ax1.set_xlim([1979, 2021])
# ax0.set_title("Coordinate offset for {}".format(souname))
plt.tight_layout()
plt.subplots_adjust(hspace=0)
plt.savefig("{:s}/{:s}.eps".format(plotdir, souname))
# -
# Simulate the random walk process.
from tool_func import random_walk
dra, ddec = random_walk(coordts["jyear"], t_scale=5, sigma_var=5)
# +
fig, (ax0, ax1) = plt.subplots(nrows=2, sharex=True, sharey=True)
ax0.errorbar(coordts["jyear"], dra,
fmt="b.", ecolor="grey", elinewidth=0.1, ms=1)
ax1.errorbar(coordts["jyear"], ddec,
fmt="b.", ecolor="grey", elinewidth=0.1, ms=1)
ax0.xaxis.set_minor_locator(MultipleLocator(1))
ax0.set_ylabel("$\Delta\\alpha\,\cos\delta$ ($\mathrm{mas}$)")
ax1.set_ylabel("$\Delta\delta$ ($\mathrm{mas}$)")
ax1.set_xlabel("Year")
ax1.set_ylim([-5, 5])
ax1.set_xlim([1979, 2021])
# ax0.set_title("Random walk simulation for {}".format(souname))
plt.tight_layout()
plt.subplots_adjust(hspace=0)
plt.savefig("{:s}/{:s}-RW.eps".format(plotdir, souname))
|
progs/.ipynb_checkpoints/random-walk-simulation-sample-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [ast542]
# language: python
# name: Python [ast542]
# ---
# %matplotlib inline
# %config InlineBackend.figure_format='retina'
import matplotlib.pyplot as plt
from matplotlib import colors, ticker, cm
import numpy as np
from scipy.interpolate import interp1d
from orbit_class import Orbit
import misc_utils as mu
import ifs_noise_model as snr_ifs
import imaging_noise_model as snr_im
import juneper_model as hpm
import coolTLUSTY_model as cpm
import simulate_data as simdata
import nice_plotting_forms as pf
epsEri = {}
epsEri.update({'d_obs':3.216}) # distance to the observer in parsecs
M = 3.73 - 5.0*np.log10(epsEri['d_obs']/10.0) # convert apparent (3.73) to absolute
epsEri.update({'Mstar':M}) # absolute stellar V-band magnitude
epsEri.update({'Nez':1.0}) # exozodi level
stellartype = 'k0v' # stellar type
stellar_mass = 0.781 # stellar mass in units of solar mass
# this is only used in orbit object, not really relevant for SNR calculations
stellar_spec = 'AuxiliaryData/'+stellartype+'.dat' # imports Hubble
ref_wl, ref_flambda = np.loadtxt(stellar_spec, unpack=True, usecols=(0,1))
fstar_lambda = interp1d(ref_wl,ref_flambda) # specific flux density W / m^2 / micron, for zero mag star
epsEri.update({'fstar_lambda':fstar_lambda}) # a function which returns
# specific flux density for any wavelength
epsEri.update({'rp':0.83294}) # planet radius in Jupiter radii
# epsEri.update({'met':3.0})
# epsEri.update({'Ag':cpm.ctlustyAg_more_wl(epsEri['met'])})
epsEri.update({'chromo':1.0,'meth':1.0})
epsEri.update({'Ag':hpm.juneper_Agfunc(epsEri['chromo'],epsEri['meth'])})
a = 3.4 # semimajor axis (in au)
ecc = 0.071 # eccentricity
inc = 90.0 # inclination (degrees)
ome = 180.0 # longitude of ascending node (degrees)
tp = 2530054 % (7.37*365) # epoch of perihelion passage (julian date) 2530054
argperi = 3.13 # argument of perihelion (degrees)
epsEri_orbit_pars = np.array([ecc,inc,ome,tp,a,argperi,
stellar_mass,epsEri['rp'],epsEri['d_obs']])
epsEri_orbit = Orbit(epsEri_orbit_pars)
ophase = 0.8 # orbital phase
pfunc = mu.avg_empirical # type of phase function to use
simdata.change_ophase(epsEri_orbit, epsEri, ophase, pfunc) # updates params['sep'], params['phi'], params['wa']
epsEri.update(mu.cbe_spc_pars)
# +
# look at prospects with the IFS
epsEri.update(mu.cbe_spc_pars)
wavelengths = np.arange(0.45,1.0,0.01)
plt.figure(figsize=(17.0, 9.0))
ax=plt.subplot(2,2,1)
epsEri.update({'met':3.0})
epsEri.update({'Ag':cpm.ctlustyAg_more_wl(epsEri['met'])})
pf.ophase_tsnr_plot(epsEri,epsEri_orbit,wavelengths,5.0,cbar=False)
ax.set_title('$\epsilon$ Eridani b 3.0 x Solar Metallicity',fontsize=17)
epsEri.update({'met':30.0})
epsEri.update({'Ag':cpm.ctlustyAg_more_wl(epsEri['met'])})
ax=plt.subplot(2,2,2)
pf.ophase_tsnr_plot(epsEri,epsEri_orbit,wavelengths,5.0,cbar=False)
ax.set_title('$\epsilon$ Eridani b 30.0 x Solar Metallicity',fontsize=17)
epsEri.update({'met':3.0})
epsEri.update({'Ag':cpm.ctlustyAg_more_wl(epsEri['met'])})
ax=plt.subplot(2,2,3)
pf.rp_tsnr_plot(epsEri,wavelengths,5.0,cbar=False)
plt.hlines(0.83294,0.4,1.0,color='grey')
plt.axis([0.45,1.0,0.2,2.0])
epsEri.update({'met':30.0})
epsEri.update({'Ag':cpm.ctlustyAg_more_wl(epsEri['met'])})
ax=plt.subplot(2,2,4)
pf.rp_tsnr_plot(epsEri,wavelengths,5.0,cbar=True)
plt.hlines(0.83294,0.4,1.0,color='grey')
plt.axis([0.45,1.0,0.2,2.0])
plt.tight_layout()
plt.show()
# +
# first lets look at the physical separation, phase angle, and projected separation as a function
# of time for 2019-2029
dates, seps, alphas, projs = np.loadtxt('phase_separation.csv',usecols=(0,2,5,8),unpack=True,skiprows=1,delimiter=',')
lower_bounds = np.loadtxt('phase_separation.csv',usecols=(0,1,4,7),unpack=True,skiprows=1,delimiter=',')
upper_bounds = np.loadtxt('phase_separation.csv',usecols=(0,3,6,9),unpack=True,skiprows=1,delimiter=',')
plt.figure(figsize=(20,10))
plt.subplot(1,3,1)
plt.fill_between(dates,lower_bounds[1],upper_bounds[1],alpha=0.1)
plt.plot(dates,seps,color='k')
plt.ylabel('physical separation, au',fontsize=17)
plt.subplot(1,3,2)
plt.fill_between(dates,lower_bounds[2],upper_bounds[2],alpha=0.1)
plt.plot(dates,alphas,color='k')
plt.xlabel('year',fontsize=17)
plt.ylabel('phase angle, degrees',fontsize=17)
plt.subplot(1,3,3)
plt.fill_between(dates,lower_bounds[3],upper_bounds[3],alpha=0.1)
plt.ylabel('projected separation, mas',fontsize=17)
plt.plot(dates,projs,color='k')
# +
# now lets incorporate our planet models to get a delta magnitude to go along with these...
plt.figure(figsize=(20,10))
lc_im1 = []
lc_imB = []
dates, seps, alphas, projs = np.loadtxt('phase_separation.csv',usecols=(0,2,5,8),unpack=True,skiprows=1,delimiter=',')
for alpha,date,sep,proj in zip(alphas,dates,seps,projs):
epsEri['sep'] = sep
epsEri['phi'] = mu.avg_empirical(alpha,degrees=True)
epsEri['wa'] = proj / 206264806.24709466 # want it to be in radians, not mas
epsEri.update(mu.cbe_hlc_pars)
lc_im1.append(simdata.imaging_delta_mag(0.575, 10.1, epsEri))
lc_imB.append(simdata.imaging_delta_mag(0.45, 22.0, epsEri))
lc_im1 = np.array(lc_im1)
lc_imB = np.array(lc_imB)
plt.plot(dates,lc_im1,color='black',linewidth=4,label='0.575 $\mu$m, 10% width (WFIRST-CGI)')
plt.plot(dates,lc_imB,color='blue',linewidth=4,label='0.450 $\mu$m, 22% width (~B band) ')
lc_im1 = []
lc_imB = []
dates, seps, alphas, projs = lower_bounds
for alpha,date,sep,proj in zip(alphas,dates,seps,projs):
epsEri['sep'] = sep
epsEri['phi'] = mu.avg_empirical(alpha,degrees=True)
epsEri['wa'] = proj / 206264806.24709466 # want it to be in radians, not mas
epsEri.update(mu.cbe_hlc_pars)
lc_im1.append(simdata.imaging_delta_mag(0.575, 10.1, epsEri))
lc_imB.append(simdata.imaging_delta_mag(0.45, 22.0, epsEri))
lc_im1_lb = np.array(lc_im1)
lc_imB_lb = np.array(lc_imB)
lc_im1 = []
lc_imB = []
dates, seps, alphas, projs = upper_bounds
for alpha,date,sep,proj in zip(alphas,dates,seps,projs):
epsEri['sep'] = sep
epsEri['phi'] = mu.avg_empirical(alpha,degrees=True)
epsEri['wa'] = proj / 206264806.24709466 # want it to be in radians, not mas
epsEri.update(mu.cbe_hlc_pars)
lc_im1.append(simdata.imaging_delta_mag(0.575, 10.1, epsEri))
lc_imB.append(simdata.imaging_delta_mag(0.45, 22.0, epsEri))
lc_im1_ub = np.array(lc_im1)
lc_imB_ub = np.array(lc_imB)
plt.fill_between(dates,lc_im1_lb,lc_im1_ub,facecolor='black',alpha=0.1)
plt.fill_between(dates,lc_imB_lb,lc_imB_ub,facecolor='blue',alpha=0.1)
plt.axis([2019,2029,25.0,20.0])
plt.hlines(-2.5*np.log10(10.0**-9.0),2019,2030,color='grey',linewidth=1)
plt.xlabel('year',fontsize=17)
plt.ylabel('M$_{planet}$ - M$_{star}$',fontsize=17)
plt.title('$\epsilon$ Eridani b assuming Jupiter\'s albedo and Phase Function',fontsize=17)
plt.legend(frameon=False,loc='lower left',fontsize=15)
# -
|
WFIRST_SIM/epsEri/epsEri.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GA4GH Variation Representation Schema
#
# This notebook demonstrates the use of the VR schema to represent variation in APOE. Objects created in this notebook are saved at the end and used by other notebooks to demonstrate other features of the VR specification.
#
#
# ## APOE Variation
#
# rs7412
# NC_000019.10:g.44908822
# NM_000041.3:c.526
# C T
# rs429358 C APOE-ε4 APOE-ε1
# NC_000019.10:g.44908684 T APOE-ε3 APOE-ε2
# NM_000041.3:c.388
#
# Note: The example currently uses only rs7412:T. Future versions of the schema will support haplotypes and genotypes, and these examples will be extended appropriately.
# ## Using the VR Reference Implemention
#
# See https://github.com/ga4gh/vr-python for information about installing the reference implementation.
from ga4gh.vrs import __version__, models
__version__
# ## Schema Overview
#
# <img src="images/schema-current.png" width="75%" alt="Current Schema"/>
# ## Sequences
#
# The VR Specfication expects the existence of a repository of biological sequences. At a minimum, these sequences must be indexed using whatever accessions are available. Implementations that wish to use the computed identifier mechanism should also have precomputed ga4gh sequence accessions. Either way, sequences must be referred to using [W3C Compact URIs (CURIEs)](https://w3.org/TR/curie/). In the examples below, we'll use "refseq:NC_000019.10" to refer to chromosome 19 from GRCh38.
# ## Locations
# A Location is an *abstract* object that refer to contiguous regions of biological sequences.
#
# In the initial release of VR, the only Location is a SequenceLocation, which represents a precise interval (`SimpleInterval`) on a sequence. GA4GH VR uses interbase coordinates exclusively; therefore the 1-based residue position 44908822 is referred to using the 0-based interbase interval <44908821, 44908822>.
#
# Future Location subclasses will provide for approximate coordinates, gene symbols, and cytogenetic bands.
# #### SequenceLocation
location = models.SequenceLocation(
sequence_id="refseq:NC_000019.10",
interval=models.SimpleInterval(start=44908821, end=44908822, type="SimpleInterval"),
type="SequenceLocation")
location.as_dict()
# ## Variation
#
# ### Text Variation
#
# The TextVariation class represents variation descriptions that cannot be parsed, or cannot be parsed yet. The primary use for this class is to allow unparsed variation to be represented within the VR framework and be associated with annotations.
variation = models.Text(definition="APO loss", type="Text")
variation.as_dict()
# ### Alleles
#
# An Allele is an asserion of a state of biological sequence at a Location. In the first version of the VR Schema, the only State subclass is SequenceState, which represents the replacement of sequence. Future versions of State will enable representations of copy number variation.
# ### "Simple" sequence replacements
# This case covers any "ref-alt" style variation, which includes SNVs, MNVs, del, ins, and delins.
allele = models.Allele(location=location,
state=models.SequenceState(sequence="A", type="SequenceState"),
type="Allele")
allele.as_dict()
# ----
#
# ## Saving the objects
#
# Objects created in this notebook will be saved as a json file and loaded by subsequent notebooks.
import json
filename = "objects.json"
data = {
"alleles": [allele.as_dict()],
"locations": [location.as_dict()]
}
json.dump(data, open(filename, "w"), indent=4)
|
notebooks/Schema.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.display import HTML
css_file = './custom.css'
HTML(open(css_file, "r").read())
# # Scaling and Outliers
# ## 1. Definition
#
# ***Scaling***, ***standardizing*** and ***normalizing*** are used somewhat interchangeably, even though they are ***not the same thing***.
#
# True, ***standardizing*** and ***normalizing*** can be seen as particular ways of ***scaling***. Let's sort the differences between all them before proceeding.
#
# ### 1.1 Scaling vs Standardizing vs Normalizing
#
# According to the implementations in Scikit-Learn library:
#
# 1. ***Scaling (MinMaxScaler)***:
#
# Transforms features by scaling each feature to a given range.
#
# This estimator scales and translates each feature individually such that it is in the given range on the training set, e.g. between zero and one.
#
#
# 2. ***Standardizing (StandardScaler)***:
#
# Standardize features by removing the mean and ***scaling to unit variance
#
# The standard score of a sample x is calculated as:
#
# z = (x - u) / s
#
# where u is the mean of the training samples or zero if with_mean=False, and s is the standard deviation of the training samples or one if with_std=False.
#
#
# 3. ***Normalizing (Normalizer)***:
#
# Normalize samples individually to unit norm.
#
# Each sample (i.e. each row of the data matrix) with at least one non zero component is rescaled independently of other samples so that its norm (l1 or l2) equals one.
#
# The main difference is: ***scaling*** and ***standardizing*** operate on ***features*** (columns of your dataset), while ***normalizing*** operates on ***individual samples*** (rows on your dataset).
#
# Even though it is very common for people to say *"you need to normalize on your features"*, they usually mean ***standardize your features***.
#
# Why ***standardize*** instead of simply ***scaling***? It turns out, ***scaling*** is fine when you have features with a ***limited range***, like ***age*** (0-120) or ***pixel values*** (0-255), but it is likely a bad choice whenever your features may have values ***very far apart***, like ***salaries*** (maybe not yours and mine, but think of some CEOs...).
#
# ### 1.2 Why Scaling?
#
# Most Machine Learning algorithms use ***gradient descent*** to learn the weights of the model. You'll see in the next lesson that ***scaling*** the features makes a ***BIG*** difference in performance.
#
# It is also important for other techniques, like ***Principal Component Analysis (PCA)*** for dimensionality reduction, and for identifying ***outliers***.
#
# 
# <center>Source: <a href="https://xkcd.com/1162/">XKCD</a></center>
#
# ### 1.3 Outliers
#
# What is an ***outlier***? It could be defined in several ways:
#
# - a point that is distant from the others (again, think of salaries, everyone is between USD 30k and 100k per year, and a CEO is making 50M!)
# - a point that is distinct from the others (think of black sheep in a flock of white sheep)
# - an error of measurement (think of someone listed as being 450 years old)
# - an anomaly / fraud (think of finding the purchase of a USD 10k Rolex on your credit card bill)
#
# The last case, anomalies / frauds, is a special case where ***your goal is to detect the outlier***.
#
# For now, let's focus on the other cases: in all of them, the ***presence of an outlier*** may ***hurt your model***, impacting its training and making its predictions less useful.
#
# So, how do you ***detect*** and ***remove or fence outliers***?
#
# #### 1.3.1 Tukey's Fences
#
# This is a very straightforward way of detecting ***possible*** outliers based on the ***InterQuartile Range (IQR)***.
#
# It defines a ***lower*** and an ***upper fence***, which are given by:
#
# $$
# IQR = Q_3 - Q_1
# \\
# lower fence = Q1 - k * IQR
# \\
# upper fence = Q3 + k * IQR
# $$
#
# Typical values for ***k*** are 1.5 (outlier) and 3.0 (far out).
#
# The plot below illustrates this:
#
# 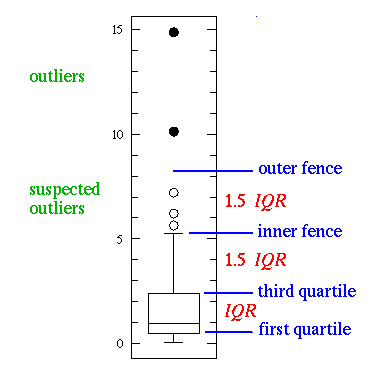
#
# <center>Source: <a href="http://www.physics.csbsju.edu/stats/box2.html">Box Plot: Display of Distribution</a></center>
#
# Although easy to compute, ***Tukey's Fences*** only consider the distribution of a ***single feature*** to assess values as outliers or not.
#
# What if we wanted to check if a value can be considered an outlier, ***given its many features***?
#
# #### 1.3.2 Mahalanobis Distance
#
# In a single dimension, we can easily compute how far (in standard deviations) a point is from the mean. This is what ***standardization*** does for a single feature (refer to the previous lesson for more details).
#
# Mahalanobis Distance is the generalization of the same idea in multiple dimensions.
#
# The ***Mahalanobis Distance*** is given by:
#
# $$
# \large\sqrt{(x_1 - x_2)\ S^{-1}\ (x_1 - x_2)}
# $$
#
# where ***S*** is the covariance matrix.
#
# If we ***standardize*** all features, the ***Mahalanobis Distance*** corresponds to the distance from the origin:
#
# $$
# \large\sqrt{x\ S^{-1}\ x}
# $$
#
# Knowing the distance is not enough, though. To determine if a given point is an ***outlier*** or not, we need to compare its computed distance to the ***cumulative chi-squared distribution*** using the ***number of features*** as ***degrees of freedom***. If it falls ***above a threshold***, like 99.9%, it is considered an ***outlier***.
#
# ```python
# from scipy.stats import chi2
#
# chi2.cdf(mahalanobis_distance, df=n_features)
# ```
# ## 2. Experiment
#
# Time to try it yourself!
#
# There are 200 data points (in blue).
#
# The controls below allow you:
#
# - change the ***scaling method***
# - obs.: MinMaxScaling is configured to scale features in [-5, 5] range
# - include ***ONE outlier*** (single red point)
# - plot ***Tukey's fences*** on horizontal and vertical axes (k = 1.5)
# - plot the ***Chi-Squared Probabilities*** contour plot for ***Mahalanobis Distance*** (under StandardScaling only!)
# - choose a ***threshold for Chi-Sq.Prob.*** between 99.1% and 99.9% for (under StandardScaling only!)
#
# Use the controls to play with different configurations and answer the questions below.
from intuitiveml.feature.Scaling import *
X = data()
mysc = plotScaling(X, outlier=(-9, 6))
vb = VBox(build_figure(mysc), layout={'align_items': 'center'})
vb
# #### Questions
#
# 1. Using ***no scaling***, turn ***Tukey's fences*** on:
# - how many ***outlier candidates*** you found on the horizontal axis (X1 feature)?
# - how many ***outlier candidates*** you found on the vertical axis (X2 feature)?
# - is there any point that is an ***outlier candidate*** on both features? If so, would you consider it an outlier or not? Why?
# - how do you like Tukey's method for outlier detection?
#
#
# 2. Include the ***outlier*** to the same configuration above:
# - is the ***red point*** an outlier according to ***Tukey's method***?
# - how do you compare the ***red point*** to any outliers you found in question 1?
#
#
# 3. Using ***MinMax Scaling***, make all boxes ***unchecked***:
# - take note of the general position of the blue points
# - include the ***outlier*** - what happens to the blue points? Why?
# - add ***Tukey's Fences*** - do you see any differences?
#
#
# 4. Using ***Standard Scaling***, make all boxes ***unchecked***:
# - take note of the general position of the blue points
# - include the ***outlier*** - what happens to the blue points? Why?
# - how is this different from what happened using ***MinMax Scaling***? Why?
# - add ***Tukey's Fences*** - do you see any differences?
#
#
# 5. Using ***Standard Scaling*** and check ONLY ***ChiSq.Prob. for L2 Norm***:
# - are there any points outside the 90% probability circle (dashed circle)?
# - include the ***outlier*** - is it outside the 90% probability circle?
# - change the ***probability threshold*** to different values and observe how the circle grows
# ## 3. Scikit-Learn
#
# [MinMaxScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
#
# [StandardScaler](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html)
#
# [Normalizer](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.Normalizer.html)
#
# [Comparison of the effect of different scalers on data with outliers](https://scikit-learn.org/stable/auto_examples/preprocessing/plot_all_scaling.html#sphx-glr-auto-examples-preprocessing-plot-all-scaling-py)
# ## 4. More Resources
#
# [About Feature Scaling and Normalization](https://sebastianraschka.com/Articles/2014_about_feature_scaling.html)
#
# [Outlier Detection with Isolation Forest](https://towardsdatascience.com/outlier-detection-with-isolation-forest-3d190448d45e)
# #### This material is copyright <NAME> and made available under the Creative Commons Attribution (CC-BY) license ([link](https://creativecommons.org/licenses/by/4.0/)).
#
# #### Code is also made available under the MIT License ([link](https://opensource.org/licenses/MIT)).
from IPython.display import HTML
HTML('''<script>
function code_toggle() {
if (code_shown){
$('div.input').hide('500');
$('#toggleButton').val('Show Code')
} else {
$('div.input').show('500');
$('#toggleButton').val('Hide Code')
}
code_shown = !code_shown
}
$( document ).ready(function(){
code_shown=false;
$('div.input').hide()
});
</script>
<form action="javascript:code_toggle()"><input type="submit" id="toggleButton" value="Show Code"></form>''')
|
5. Scaling and Outliers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
# -
# ## Read in Our Data
#
# The first step is to read out data into a `pandas` DataFrame. For an intro to using `pandas` I would highly suggest looking though [this 10 minute guide to `pandas`](http://pandas.pydata.org/pandas-docs/stable/10min.html).
df = pd.read_csv('npr_articles.csv')
# We can now checkout what our data consists of by using the `.head()` method on our DataFrame. By default, this will show the top 5 rows.
df.head()
# One of the first steps you should take is to get an overview of what kind of data we have but running the `.info()` method. Please [see the documentation](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) for more info (no pun intended).
df.info()
# We can see that the column `date_published` is being interpreted as an `object` and not a datetime. Let's change that by using the [`pandas.to_datetime()` function](http://pandas.pydata.org/pandas-docs/version/0.19.2/generated/pandas.to_datetime.html).
df['date_published'] = pd.to_datetime(df['date_published'])
df.info()
df['author'][:5]
# ## Number of Authors
#
# Let's say we wanted to add in another column that contains the number of authors that worked on a particular article. We could do this like so:
# +
# Let's create a mask for all rows that have a non-null value
mask = df['author'].notnull()
# When the data was saved to a csv, these lists were converted into strings, we can convert
# them back like so
from ast import literal_eval
if type(df['author'][0]) == type('abc'):
df.loc[mask, 'author'] = df.loc[mask, 'author'].map(literal_eval)
print(type(df['author'][0]))
# Initialize column with NaN's and then fill in the respective values
df['num_authors'] = np.nan
df.loc[mask, 'num_authors'] = df.loc[mask, 'author'].map(len)
# -
# We can now take a look at the summary statistics of any numeric columns by running the `.describe()` method.
df.describe()
df.head()
# ## Number of Unique Authors
#
# Let's say we wanted to get the number of unique authors that are represented in this dataframe. We could potentially use `df['author'].nunique()` but we are going to run into an error because each row contains a `list` which isn't hashable.
#
# Instead we could loop through each value and extend a set like so:
# Create a set to hold our authors
authors = set()
for lst in df.loc[mask, 'author']:
# For every row, update the authors set with those contained in that row
authors.update(lst)
# Print out the total authors seen
print(len(authors))
# If we also wanted the number of times a particular author was involved in writing an article we could leverage the power of `Counter`'s from the `collections` library. Refer to the [documentation](https://docs.python.org/2/library/collections.html) for more information.
from collections import Counter
authors = df.loc[mask, 'author'].map(Counter).sum()
authors
authors.most_common()
authors['<NAME>']
# Let's say we wanted to now subset down to the articles which Ari Shapiro worked on. There are a variety of way's we could do this but I will demo one possible avenue.
# +
# Because some rows have NaN's in them, we need to get clever with how we
# create our mask
mask = df['author'].map(lambda x: '<NAME>' in x if isinstance(x, list)
else False)
df.loc[mask, 'headline']
# +
# Here is another way we could acheive this
mask = df.loc[df['author'].notnull(), 'author'].map(lambda x: '<NAME>' in x)
df.loc[df['author'].notnull()].loc[mask, 'headline']
# -
# ## Most popular sections
#
# Let's find what the 5 most popular sections (as judged by the number of articles published within that article)
df['section'].value_counts(dropna=False)[:5]
# When we first were looking at our DataFrame, you may have noticed that there are quite a few rows missing author information. Maybe we have a hypothesis that there are certain sections that systemically weren't attaching author information. Let's dive deeper to try and prove/disprove this hypothesis...
# +
# Let's create a new column that indicates whether the author attribute was null or not
# This helps with the groupby below
df['author_null'] = df['author'].isnull()
# Get the mean amount of nulls for each section and sort descending
# NOTE: 1.0 indicates ALL Nulls
df.groupby('section')['author_null'].mean().sort_values(ascending=False)
# -
# As we can see, there are clearly sections that are consistently not attaching author information as well as many that are hit or miss with the author information.
# ## Article Count by Time
#
# Let's make a plot showing the frequency of articles published by day, week, and month.
# Create a pandas Series with 1's as the values and the date as the index
s = pd.Series([1], index=df['date_published'])
s[:10]
# Below we see how we could use the [resample function](http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.resample.html) to find the number of articles published per day.
#
# NOTE: Our DataFrame/Series must have a datetimeindex for this to work!
# Let's resample that Series and sum the values to find the number of articles by Day
s.resample('D').sum()
# There are, of course, many different offset alias' for passing to `resample`. For more options [see this page](http://pandas.pydata.org/pandas-docs/stable/timeseries.html#offset-aliases).
plt.plot(s.resample('D').sum())
plt.title('Article Count By Day')
plt.ylabel('Number of Articles')
plt.xlabel('Date')
locs, labels = plt.xticks()
plt.setp(labels, rotation=-45);
plt.plot(s.resample('W').sum())
plt.title('Article Count By Week')
plt.ylabel('Number of Articles')
plt.xlabel('date')
locs, labels = plt.xticks()
plt.setp(labels, rotation=-45);
# ## What hour is the most popular time for publishing articles?
#
# To answer this let's extract the hour when the article was published and create a histogram.
df['hour_published'] = df['date_published'].dt.hour
# We were able to run the above command because that particular column contains a datetime object. From there we can run `.dt` and then extract any aspect of that datetime (e.g. `.dt.hour`, `.dt.second`, `.dt.month`, `.dt.quarter`)
df['hour_published'].hist()
plt.ylabel('Number of Articles Published')
plt.xlabel('Hour Published (24Hr)');
# By default, the `.hist` method is going to plot 10 bins. Let's up that to 24 bins so we have a bin for each hour in the day...
# Let's force the plot to split into 24 bins, one for each hour
df['hour_published'].hist(bins=24)
plt.ylabel('Number of Articles Published')
plt.xlabel('Hour Published (24Hr)');
# Let's extract the relative frequency rather than the raw counts
df['hour_published'].hist(bins=24, normed=True, alpha=0.75)
plt.ylabel('Freq. of Articles Published')
plt.xlabel('Hour Published (24Hr)');
# We can also grab this information without plotting it using .value_counts
df['hour_published'].value_counts()
df['hour_published'].value_counts(normalize=True)
# Or we could leave them in the order of a day
df['hour_published'].value_counts().sort_index()
# ## Selecting Particular Dates
#
# Let's select articles which were published between 10 am and 2 pm on December 24th, 2016. There are a couple of ways we could do this, but let's start by making a mask.
mask = ((df['date_published'] >= '2016-12-24 10:00:00') &
(df['date_published'] <= '2016-12-24 14:00:00'))
df.loc[mask, :]
# Or we could reset or index and do it that way...
df2 = df.set_index('date_published')
df2.loc['2016-12-24 10:00:00': '2016-12-24 14:00:00', :]
# ## Length of Articles (# Words)
#
# Maybe we are interested in looking at the distribution of how long our articles are...
df['num_words'] = df['article_text'].map(lambda x: len(x.split()))
df['num_words'].describe()
# Let's create a histogram of the length of different articles...
df['num_words'].hist(bins=20, alpha=0.75)
plt.ylabel('Number of Articles Published')
plt.xlabel('Length of Article');
# Clearly there are some outliers in this data. Let's subset what we are plotting to cut out the top 2% of articles in terms of article length and see what the resulting histogram looks like...
#
# Refer to [the `numpy` percentile function](https://docs.scipy.org/doc/numpy-dev/reference/generated/numpy.percentile.html) for more information.
# +
cutoff = np.percentile(df['num_words'], 98)
df.loc[df['num_words'] <= cutoff, 'num_words'].hist(bins=20, alpha=0.75)
plt.ylabel('Number of Articles Published')
plt.xlabel('Length of Article');
# -
# ## Only rows that contain 'Obama' in the Headline
#
# We can also use standard string functions by using the `.str` functionality in `pandas`. Take a look at [this page](http://pandas.pydata.org/pandas-docs/stable/text.html) for more information.
df.loc[df['headline'].str.contains('Obama'), 'headline'].head()
# ## Looking at Average Hour Published by Section
#
# Maybe we have a hypothesis that different sections will vary in the time of day that they are publishing. We could try and get a sense for this like so:
# +
# Let's subset to just the 10 most popular sections
top_sections = df['section'].value_counts()[:10].index
df_sub = df.loc[df['section'].isin(top_sections), :]
# We are now grouping by the section and extracting the mean hour that articles were published
df_sub.groupby('section')['hour_published'].mean()
# -
|
live_coding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="pz2A3mM_Z_yf" colab_type="code" colab={}
from matplotlib import pyplot
# + id="iHQH97qAbBbh" colab_type="code" colab={}
def relu(x):
return max(0.0, x)
# + id="Vmi8yaaPbDy5" colab_type="code" colab={}
inputs = [x for x in range(-15, 15)]
outputs = [relu(x) for x in inputs]
# + id="kxgdvTO2bFnJ" colab_type="code" outputId="cfc942d3-cdb6-45ed-d212-34469e693e29" colab={"base_uri": "https://localhost:8080/", "height": 269}
pyplot.plot(inputs, outputs) #Plot the input against the output
pyplot.show()
|
Lesson 04/Exercise_19_Visualizing_ReLU (2).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to define compute magnetic flux with FEMM
#
# This tutorial shows the how to **compute magnetic flux and torque with FEMM**.
#
# The notebook related to this tutorial is available on [GitHub](https://github.com/Eomys/pyleecan/tree/master/Tutorials/tuto_Simulation_FEMM.ipynb).
#
#
# This tutorial is divided into four parts:
# - machine import
# - winding definition
# - magnetic simulation definition and running
# - plot of the magnetic flux for the first time step
#
# ### Loading Machine
#
# Before defining the simulation, one first has to define the machine. For this example we import the Toyota Prius 2004 defined in [this tutorial](https://www.pyleecan.org/tuto_Machine.html).
# +
# Add pyleecan to the Python path
import sys
sys.path.append('../..')
from pyleecan.Functions.load import load
# from pyleecan.Tests.Validation.Machine.SPMSM_003 import SPMSM_003 as IPMSM_A
# from pyleecan.Tests.Validation.Machine.IPMSM_A import IPMSM_A as IPMSM_A
# Import the machine from a script
IPMSM_A = load('../Data/Machine/IPMSM_A.json')
# %matplotlib notebook
im=IPMSM_A.plot()
# -
# ## Simulation definition
# ### Input currents
#
# To define the simulation, we use [Simu1](http://www.pyleecan.com/pyleecan.Classes.Simu1.html) and [InCurrent](http://www.pyleecan.com/pyleecan.Classes.InCurrent.html) to define the input such as stator currents, the angular and the time discretization.
# +
import numpy as np
from numpy import ones, pi, array, linspace
from pyleecan.Classes.Simu1 import Simu1
from pyleecan.Classes.InCurrent import InCurrent
from pyleecan.Classes.MagFEMM import MagFEMM
# Create the Simulation
mySimu = Simu1(name="EM_SIPMSM_AL_001", machine=IPMSM_A)
# Defining Simulation Input
mySimu.input = InCurrent()
# Electrical time vector without symmetry [s]
mySimu.input.time.value= np.linspace(start=0, stop=0, num=1, endpoint=False)
# Angular steps along the airgap circonference for flux density calculation
mySimu.input.angle.value = np.linspace(start = 0, stop = 2*np.pi, num=2048, endpoint=False) # 2048 steps
# Rotor speed as a function of time [rpm]
mySimu.input.Nr.value = ones(1) * 2504
# Stator currents as a function of time, each column correspond to one phase [A]
mySimu.input.Is.value = array(
[
[0, 12.2474, -12.2474],
]
)
# -
# To call FEMM, we need to define the magnetic part of the simulation with [MagFEMM](http://www.pyleecan.org/pyleecan.Classes.MagFEMM.html) class. As the simulation will only consider magnetic problem, we set the structural part as None to avoid computation.
#
from pyleecan.Classes.MagFEMM import MagFEMM
# Definition of the magnetic simulation (is_mmfr=False => no flux from the magnets)
mySimu.mag = MagFEMM(
is_stator_linear_BH=0, # 0 to use the B(H) curve,
# 1 to use linear B(H) curve according to mur_lin,
# 2 to enforce infinite permeability (mur_lin =100000)
is_rotor_linear_BH=0, # 0 to use the B(H) curve,
# 1 to use linear B(H) curve according to mur_lin,
# 2 to enforce infinite permeability (mur_lin =100000)
is_symmetry_a=True, # 0 Compute on the complete machine, 1 compute according to sym_a and is_antiper_a
sym_a = 4, # Number of symmetry for the angle vector
is_antiper_a=True, # To add an antiperiodicity to the angle vector
angle_stator=-np.pi / 6, # Angular position shift of the stator
)
mySimu.struct = None # We only use the magnetic part
# You can find all the parameters of _MagFEMM_ by looking at [Magnetics](http://www.pyleecan.org/pyleecan.Classes.Magnetics.html) and [MagFEMM](http://www.pyleecan.org/pyleecan.Classes.MagFEMM.html) classes.
#
# ### Run simulation
# To run the simulation, we first have to set the Output to store the results.
from pyleecan.Classes.Output import Output
myResults = Output(simu=mySimu)
mySimu.run()
# Once it is done, the results are stored in the magnetic part of the output (i.e. _out.mag_ ) and one can call different plots :
# %matplotlib notebook
myResults.plot_B_space()
|
Tutorials/tuto_Simulation_FEMM.ipynb
|
try:
import openmdao.api as om
import dymos as dm
except ImportError:
# !python -m pip install openmdao[notebooks]
# !python -m pip install dymos[docs]
import openmdao.api as om
import dymos as dm
# # Multibranch Trajectory
#
# This example demonstrates the use of a Trajectory to encapsulate a
# series of branching phases.
#
# ## Overview
#
# For this example, we build a system that contains two components: the
# first component represents a battery pack that contains multiple cells
# in parallel, and the second component represents a bank of DC electric
# motors (also in parallel) driving a gearbox to achieve a desired power
# output. The battery cells have a state of charge that decays as current
# is drawn from the battery. The open circuit voltage of the battery is a
# function of the state of charge. At any point in time, the coupling
# between the battery and the motor component is solved with a Newton
# solver in the containing group for a line current that satisfies the
# equations.
#
# Both the battery and the motor models allow the number of cells and the
# number of motors to be modified by setting the _n\_parallel_ option in
# their respective options dictionaries. For this model, we start with 3
# cells and 3 motors. We will simulate failure of a cell or battery by
# setting _n\_parallel_ to 2.
#
# Branching phases are a set of linked phases in a trajectory where the
# input ends of multiple phases are connected to the output of a single
# phase. This way you can simulate alternative trajectory paths in the
# same model. For this example, we will start with a single phase
# (_phase0_) that simulates the model for one hour. Three follow-on
# phases will be linked to the output of the first phase: _phase1_ will
# run as normal, _phase1\_bfail_ will fail one of the battery cells, and
# _phase1\_mfail_ will fail a motor. All three of these phases start
# where _phase0_ leaves off, so they share the same initial time and
# state of charge.
#
# ## Battery and Motor models
#
# The models are loosely based on the work done in Chin {cite}`chin2019battery`.
# +
"""
Simple dynamic model of a LI battery.
"""
import numpy as np
from scipy.interpolate import Akima1DInterpolator
import openmdao.api as om
# Data for open circuit voltage model.
train_SOC = np.array([0., 0.1, 0.25, 0.5, 0.75, 0.9, 1.0])
train_V_oc = np.array([3.5, 3.55, 3.65, 3.75, 3.9, 4.1, 4.2])
class Battery(om.ExplicitComponent):
"""
Model of a Lithium Ion battery.
"""
def initialize(self):
self.options.declare('num_nodes', default=1)
self.options.declare('n_series', default=1, desc='number of cells in series')
self.options.declare('n_parallel', default=3, desc='number of cells in parallel')
self.options.declare('Q_max', default=1.05,
desc='Max Energy Capacity of a battery cell in A*h')
self.options.declare('R_0', default=.025,
desc='Internal resistance of the battery (ohms)')
def setup(self):
num_nodes = self.options['num_nodes']
# Inputs
self.add_input('I_Li', val=np.ones(num_nodes), units='A',
desc='Current demanded per cell')
# State Variables
self.add_input('SOC', val=np.ones(num_nodes), units=None, desc='State of charge')
# Outputs
self.add_output('V_L',
val=np.ones(num_nodes),
units='V',
desc='Terminal voltage of the battery')
self.add_output('dXdt:SOC',
val=np.ones(num_nodes),
units='1/s',
desc='Time derivative of state of charge')
self.add_output('V_oc', val=np.ones(num_nodes), units='V',
desc='Open Circuit Voltage')
self.add_output('I_pack', val=0.1*np.ones(num_nodes), units='A',
desc='Total Pack Current')
self.add_output('V_pack', val=9.0*np.ones(num_nodes), units='V',
desc='Total Pack Voltage')
self.add_output('P_pack', val=1.0*np.ones(num_nodes), units='W',
desc='Total Pack Power')
# Derivatives
row_col = np.arange(num_nodes)
self.declare_partials(of='V_oc', wrt=['SOC'], rows=row_col, cols=row_col)
self.declare_partials(of='V_L', wrt=['SOC'], rows=row_col, cols=row_col)
self.declare_partials(of='V_L', wrt=['I_Li'], rows=row_col, cols=row_col)
self.declare_partials(of='dXdt:SOC', wrt=['I_Li'], rows=row_col, cols=row_col)
self.declare_partials(of='I_pack', wrt=['I_Li'], rows=row_col, cols=row_col)
self.declare_partials(of='V_pack', wrt=['SOC', 'I_Li'], rows=row_col, cols=row_col)
self.declare_partials(of='P_pack', wrt=['SOC', 'I_Li'], rows=row_col, cols=row_col)
self.voltage_model = Akima1DInterpolator(train_SOC, train_V_oc)
self.voltage_model_derivative = self.voltage_model.derivative()
def compute(self, inputs, outputs):
opt = self.options
I_Li = inputs['I_Li']
SOC = inputs['SOC']
V_oc = self.voltage_model(SOC, extrapolate=True)
outputs['V_oc'] = V_oc
outputs['V_L'] = V_oc - (I_Li * opt['R_0'])
outputs['dXdt:SOC'] = -I_Li / (3600.0 * opt['Q_max'])
outputs['I_pack'] = I_Li * opt['n_parallel']
outputs['V_pack'] = outputs['V_L'] * opt['n_series']
outputs['P_pack'] = outputs['I_pack'] * outputs['V_pack']
def compute_partials(self, inputs, partials):
opt = self.options
I_Li = inputs['I_Li']
SOC = inputs['SOC']
dV_dSOC = self.voltage_model_derivative(SOC, extrapolate=True)
partials['V_oc', 'SOC'] = dV_dSOC
partials['V_L', 'SOC'] = dV_dSOC
partials['V_L', 'I_Li'] = -opt['R_0']
partials['dXdt:SOC', 'I_Li'] = -1./(3600.0*opt['Q_max'])
n_parallel = opt['n_parallel']
n_series = opt['n_series']
V_oc = self.voltage_model(SOC, extrapolate=True)
V_L = V_oc - (I_Li * opt['R_0'])
partials['I_pack', 'I_Li'] = n_parallel
partials['V_pack', 'I_Li'] = -opt['R_0']
partials['V_pack', 'SOC'] = n_series * dV_dSOC
partials['P_pack', 'I_Li'] = n_parallel * n_series * (V_L - I_Li * opt['R_0'])
partials['P_pack', 'SOC'] = n_parallel * I_Li * n_series * dV_dSOC
# num_nodes = 1
# prob = om.Problem(model=Battery(num_nodes=num_nodes))
# model = prob.model
# prob.setup()
# prob.set_solver_print(level=2)
# prob.run_model()
# derivs = prob.check_partials(compact_print=True)
# +
"""
Simple model for a set of motors in parallel where efficiency is a function of current.
"""
import numpy as np
import openmdao.api as om
class Motors(om.ExplicitComponent):
"""
Model for motors in parallel.
"""
def initialize(self):
self.options.declare('num_nodes', default=1)
self.options.declare('n_parallel', default=3, desc='number of motors in parallel')
def setup(self):
num_nodes = self.options['num_nodes']
# Inputs
self.add_input('power_out_gearbox', val=3.6*np.ones(num_nodes), units='W',
desc='Power at gearbox output')
self.add_input('current_in_motor', val=np.ones(num_nodes), units='A',
desc='Total current demanded')
# Outputs
self.add_output('power_in_motor', val=np.ones(num_nodes), units='W',
desc='Power required at motor input')
# Derivatives
row_col = np.arange(num_nodes)
self.declare_partials(of='power_in_motor', wrt=['*'], rows=row_col, cols=row_col)
def compute(self, inputs, outputs):
current = inputs['current_in_motor']
power_out = inputs['power_out_gearbox']
n_parallel = self.options['n_parallel']
# Simple linear curve fit for efficiency.
eff = 0.9 - 0.3 * current / n_parallel
outputs['power_in_motor'] = power_out / eff
def compute_partials(self, inputs, partials):
current = inputs['current_in_motor']
power_out = inputs['power_out_gearbox']
n_parallel = self.options['n_parallel']
eff = 0.9 - 0.3 * current / n_parallel
partials['power_in_motor', 'power_out_gearbox'] = 1.0 / eff
partials['power_in_motor', 'current_in_motor'] = 0.3 * power_out / (n_parallel * eff**2)
# num_nodes = 1
# prob = om.Problem(model=Motors(num_nodes=num_nodes))
# model = prob.model
# prob.setup()
# prob.run_model()
# derivs = prob.check_partials(compact_print=True)
# +
"""
ODE for example that shows how to use multiple phases in Dymos to model failure of a battery cell
in a simple electrical system.
"""
import numpy as np
import openmdao.api as om
class BatteryODE(om.Group):
def initialize(self):
self.options.declare('num_nodes', default=1)
self.options.declare('num_battery', default=3)
self.options.declare('num_motor', default=3)
def setup(self):
num_nodes = self.options['num_nodes']
num_battery = self.options['num_battery']
num_motor = self.options['num_motor']
self.add_subsystem(name='pwr_balance',
subsys=om.BalanceComp(name='I_Li', val=1.0*np.ones(num_nodes),
rhs_name='pwr_out_batt',
lhs_name='P_pack',
units='A', eq_units='W', lower=0.0, upper=50.))
self.add_subsystem('battery', Battery(num_nodes=num_nodes, n_parallel=num_battery),
promotes_inputs=['SOC'],
promotes_outputs=['dXdt:SOC'])
self.add_subsystem('motors', Motors(num_nodes=num_nodes, n_parallel=num_motor))
self.connect('battery.P_pack', 'pwr_balance.P_pack')
self.connect('motors.power_in_motor', 'pwr_balance.pwr_out_batt')
self.connect('pwr_balance.I_Li', 'battery.I_Li')
self.connect('battery.I_pack', 'motors.current_in_motor')
self.nonlinear_solver = om.NewtonSolver(solve_subsystems=False, maxiter=20)
self.linear_solver = om.DirectSolver()
# -
# ## Building and running the problem
# +
import matplotlib.pyplot as plt
import openmdao.api as om
import dymos as dm
from dymos.examples.battery_multibranch.battery_multibranch_ode import BatteryODE
from dymos.utils.lgl import lgl
prob = om.Problem()
opt = prob.driver = om.ScipyOptimizeDriver()
opt.declare_coloring()
opt.options['optimizer'] = 'SLSQP'
num_seg = 5
seg_ends, _ = lgl(num_seg + 1)
traj = prob.model.add_subsystem('traj', dm.Trajectory())
# First phase: normal operation.
transcription = dm.Radau(num_segments=num_seg, order=5, segment_ends=seg_ends, compressed=False)
phase0 = dm.Phase(ode_class=BatteryODE, transcription=transcription)
traj_p0 = traj.add_phase('phase0', phase0)
traj_p0.set_time_options(fix_initial=True, fix_duration=True)
traj_p0.add_state('state_of_charge', fix_initial=True, fix_final=False,
targets=['SOC'], rate_source='dXdt:SOC')
# Second phase: normal operation.
phase1 = dm.Phase(ode_class=BatteryODE, transcription=transcription)
traj_p1 = traj.add_phase('phase1', phase1)
traj_p1.set_time_options(fix_initial=False, fix_duration=True)
traj_p1.add_state('state_of_charge', fix_initial=False, fix_final=False,
targets=['SOC'], rate_source='dXdt:SOC')
traj_p1.add_objective('time', loc='final')
# Second phase, but with battery failure.
phase1_bfail = dm.Phase(ode_class=BatteryODE, ode_init_kwargs={'num_battery': 2},
transcription=transcription)
traj_p1_bfail = traj.add_phase('phase1_bfail', phase1_bfail)
traj_p1_bfail.set_time_options(fix_initial=False, fix_duration=True)
traj_p1_bfail.add_state('state_of_charge', fix_initial=False, fix_final=False,
targets=['SOC'], rate_source='dXdt:SOC')
# Second phase, but with motor failure.
phase1_mfail = dm.Phase(ode_class=BatteryODE, ode_init_kwargs={'num_motor': 2},
transcription=transcription)
traj_p1_mfail = traj.add_phase('phase1_mfail', phase1_mfail)
traj_p1_mfail.set_time_options(fix_initial=False, fix_duration=True)
traj_p1_mfail.add_state('state_of_charge', fix_initial=False, fix_final=False,
targets=['SOC'], rate_source='dXdt:SOC')
traj.link_phases(phases=['phase0', 'phase1'], vars=['state_of_charge', 'time'])
traj.link_phases(phases=['phase0', 'phase1_bfail'], vars=['state_of_charge', 'time'])
traj.link_phases(phases=['phase0', 'phase1_mfail'], vars=['state_of_charge', 'time'])
prob.model.options['assembled_jac_type'] = 'csc'
prob.model.linear_solver = om.DirectSolver(assemble_jac=True)
prob.setup()
prob['traj.phase0.t_initial'] = 0
prob['traj.phase0.t_duration'] = 1.0*3600
prob['traj.phase1.t_initial'] = 1.0*3600
prob['traj.phase1.t_duration'] = 1.0*3600
prob['traj.phase1_bfail.t_initial'] = 1.0*3600
prob['traj.phase1_bfail.t_duration'] = 1.0*3600
prob['traj.phase1_mfail.t_initial'] = 1.0*3600
prob['traj.phase1_mfail.t_duration'] = 1.0*3600
prob.set_solver_print(level=0)
dm.run_problem(prob)
soc0 = prob['traj.phase0.states:state_of_charge']
soc1 = prob['traj.phase1.states:state_of_charge']
soc1b = prob['traj.phase1_bfail.states:state_of_charge']
soc1m = prob['traj.phase1_mfail.states:state_of_charge']
# Plot Results
t0 = prob['traj.phases.phase0.time.time']/3600
t1 = prob['traj.phases.phase1.time.time']/3600
t1b = prob['traj.phases.phase1_bfail.time.time']/3600
t1m = prob['traj.phases.phase1_mfail.time.time']/3600
plt.subplot(2, 1, 1)
plt.plot(t0, soc0, 'b')
plt.plot(t1, soc1, 'b')
plt.plot(t1b, soc1b, 'r')
plt.plot(t1m, soc1m, 'c')
plt.xlabel('Time (hour)')
plt.ylabel('State of Charge (percent)')
I_Li0 = prob['traj.phases.phase0.rhs_all.pwr_balance.I_Li']
I_Li1 = prob['traj.phases.phase1.rhs_all.pwr_balance.I_Li']
I_Li1b = prob['traj.phases.phase1_bfail.rhs_all.pwr_balance.I_Li']
I_Li1m = prob['traj.phases.phase1_mfail.rhs_all.pwr_balance.I_Li']
plt.subplot(2, 1, 2)
plt.plot(t0, I_Li0, 'b')
plt.plot(t1, I_Li1, 'b')
plt.plot(t1b, I_Li1b, 'r')
plt.plot(t1m, I_Li1m, 'c')
plt.xlabel('Time (hour)')
plt.ylabel('Line Current (A)')
plt.legend(['Phase 1', 'Phase 2', 'Phase 2 Battery Fail', 'Phase 2 Motor Fail'], loc=2)
plt.show()
# +
from openmdao.utils.assert_utils import assert_near_equal
# Final value for State of Chrage in each segment should be a good test.
print('State of Charge after 1 hour')
assert_near_equal(soc0[-1], 0.63464982, 1e-6)
print('State of Charge after 2 hours')
assert_near_equal(soc1[-1], 0.23794217, 1e-6)
print('State of Charge after 2 hours, battery fails at 1 hour')
assert_near_equal(soc1b[-1], 0.0281523, 1e-6)
print('State of Charge after 2 hours, motor fails at 1 hour')
assert_near_equal(soc1m[-1], 0.18625395, 1e-6)
# -
# ## References
#
# ```{bibliography}
# :filter: docname in docnames
# ```
|
docs/examples/multibranch_trajectory/multibranch_trajectory.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import os
import random
import copy
import time
from pathlib import Path
import hashlib as hl
import pickle
import somhos.resources.dataset as rd
import somhos.resources.queries as rq
import somhos.methods.useful as mu
import kleis.resources.dataset as kl
kleis = kl.load_corpus()
kleis.training(features_method="simple-posseq", filter_min_count=10)
#data_path = "resources/aminer/v1"
data_path = "../../src/somhos/resources/aminer/v9beta"
# Load document ids in the Same order than the matrices
test_preselected = set(rd.get_sample_ids(data_path, related_docs=True))
test_preselected = test_preselected | set(rd.get_sample_ids(data_path, related_docs=False))
test_dataset = copy.deepcopy(test_preselected)
train_dataset = set()
fixed_seed = 0
random.seed(fixed_seed)
threshold = 0.1
# Read artminer
datafiles = sorted(rd.get_filenames(data_path))
print(datafiles)
# +
filepath = ""
if datafiles:
filepath = datafiles[1]
print(filepath)
dociter = None
if Path(filepath).exists():
dociter = rd.get_aminer_txt(filepath, merge_text_title=True)
# +
start_time, elapsed_time = time.time(), 0
kps_count = 0
kps_hashes_counts = {}
kps_hashes_keyphrases = {}
kps_hashes_idocs = {}
# idocs_wl_content = {} # idocs with large content
kps_file_segment = 58
tmp_data_path = data_path + "/tmp-pickles"
if not Path(tmp_data_path).exists():
os.mkdir(tmp_data_path)
kps_tmp_counts = tmp_data_path + "/kps-tmp-counts-%d.pkl"
kps_tmp_keyphrases = tmp_data_path + "/kps-tmp-keyphrases-%d.pkl"
kps_tmp_idocs = tmp_data_path + "/kps-tmp-idocs-%d.pkl"
for i, (idoc, title, content) in enumerate(dociter):
# Check length of content
# idocs_wcontent[idoc] = True if content.split() > 50 else False
# Sampling test dataset
if random.random() <= threshold:
test_dataset.add(idoc)
else:
# Train dataset
train_dataset.add(idoc)
#if i > 10000:
# break
if i <= 700000:
continue
if i % 100000 == 0:
print("Progress: %d" % i, file=sys.stderr)
prev_elapsed_time = elapsed_time
elapsed_time = time.time() - start_time
print("Total time: %f" % (elapsed_time/60/60), file=sys.stderr)
print("Elapsed time: %f" % ((elapsed_time - prev_elapsed_time)/60/60), file=sys.stderr)
print("Hashes: %d\n" % len(kps_hashes_keyphrases), file=sys.stderr)
# Saving keywords
mu.save_pickle(kps_hashes_counts, kps_tmp_counts % kps_file_segment)
mu.save_pickle(kps_hashes_keyphrases, kps_tmp_keyphrases % kps_file_segment)
mu.save_pickle(kps_hashes_idocs, kps_tmp_idocs % kps_file_segment)
# increase no.
kps_file_segment += 1
# reset vars
del kps_hashes_counts
del kps_hashes_keyphrases
del kps_hashes_idocs
kps_hashes_counts = {}
kps_hashes_keyphrases = {}
kps_hashes_idocs = {}
# Avoid preselected documents
#if idoc in test_preselected:
# print("Pre-selected", idoc)
# continue
try:
text = title.strip(". ") + ". " + content
keyphrases = kleis.label_text(text, post_processing=False)
for kpid, (kplabel, (kpstart, kpend)), kptext in keyphrases:
kps_count += 1
kplower = mu.lower_utf8(kptext)
kps_hash_16 = mu.hash_16bytes(kplower)
# count
kps_hashes_counts.setdefault(kps_hash_16, 0)
kps_hashes_counts[kps_hash_16] += 1
# normalized keyphrase
kps_hashes_keyphrases[kps_hash_16] = kplower
# id docs
kps_hashes_idocs.setdefault(kps_hash_16, set())
kps_hashes_idocs[kps_hash_16].add(idoc)
except ValueError:
pass
# print("\nSkipped: %s\n" % idoc, file=sys.stderr)
# save last hashes
mu.save_pickle(kps_hashes_counts, kps_tmp_counts % kps_file_segment)
mu.save_pickle(kps_hashes_keyphrases, kps_tmp_keyphrases % kps_file_segment)
mu.save_pickle(kps_hashes_idocs, kps_tmp_idocs % kps_file_segment)
# reset vars
del kps_hashes_counts
del kps_hashes_keyphrases
del kps_hashes_idocs
# +
test_dataset_path = data_path + "/test-dataset-acm02.pkl"
if not Path(test_dataset_path).exists():
with open(test_dataset_path, "wb") as fout:
pickle.dump(test_dataset, fout, pickle.HIGHEST_PROTOCOL)
train_dataset_path = data_path + "/train-dataset-acm02.pkl"
if not Path(train_dataset_path).exists():
with open(train_dataset_path, "wb") as fout:
pickle.dump(train_dataset, fout, pickle.HIGHEST_PROTOCOL)
|
notebooks/keyphrases/beta-02-all-keyphrases-hashes-generate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import math as math
import os
exec(open("../header.py").read())
# -
raw_train_data_0 = pd.read_csv(processed_root("03-bag-of-words/threshold-40/bow_train_data.csv"))
raw_val_data_0 = pd.read_csv(processed_root("03-bag-of-words/threshold-40/bow_val_data.csv"))
raw_test_data_0 = pd.read_csv(processed_root("03-bag-of-words/threshold-40/bow_test_data.csv"))
raw_train_data = pd.read_csv(processed_root("04-bow-added-features/feat_train_data-40.csv"))
raw_val_data = pd.read_csv(processed_root("04-bow-added-features/feat_val_data-40.csv"))
raw_test_data = pd.read_csv(processed_root("04-bow-added-features/feat_test_data-40.csv"))
raw_train_data_0.columns
raw_train_data.columns
raw_train_data['poem_id']
print(len(raw_train_data_0.columns.values), len(raw_train_data.columns.values))
raw_train_data['num_syllables']
raw_train_data['num_rhymes']
raw_train_data_0
raw_train_data
# ### Import
list(set(raw_train_data['poetry_author'].values))
X_train = raw_train_data\
.drop(['poetry_text', 'poetry_author', 'num_syllables', 'num_rhymes', 'ave_word_len', 'ave_line_len', 'Unnamed: 0'], axis = 1)
X_val = raw_val_data\
.drop(['poetry_text', 'poetry_author', 'num_syllables', 'num_rhymes', 'ave_word_len', 'ave_line_len', 'Unnamed: 0'], axis = 1)
y_train = raw_train_data['poetry_author']
y_val = raw_val_data['poetry_author']
X_train = raw_train_data\
.drop(['poetry_text', 'poetry_author', 'Unnamed: 0'], axis = 1)
X_val = raw_val_data\
.drop(['poetry_text', 'poetry_author', 'Unnamed: 0'], axis = 1)
y_train = raw_train_data['poetry_author']
y_val = raw_val_data['poetry_author']
# ### Model
from sklearn.naive_bayes import MultinomialNB, GaussianNB, CategoricalNB
nb = MultinomialNB()
# nb = GaussianNB()
# nb = CategoricalNB()
# ### Train
y_pred = nb.fit(X_train, y_train)
# ### Evaluate
# +
# Training
y_pred_train = nb.predict(X_train)
train_accuracy = np.mean(y_pred_train == y_train) * 100
print("Training Accuracy: %.2f%%"%(train_accuracy))
# Validation
y_pred_val = nb.predict(X_val)
val_accuracy = np.mean(y_pred_val == y_val) * 100
print("Validation Accuracy: %.2f%%"%(val_accuracy))
# -
X_train.columns
nb.coef_.shape
np.mean(nb.coef_[:, 1897:], axis=0)
coeffs = np.round_(nb.coef_, 3)
max_coeffs = list(map(max, abs(coeffs)))
print(max_coeffs)
for i, elem in enumerate(max_coeffs):
print(len(np.where(abs(coeffs[i]) == elem)[0]))
for coefs in nb.coef_:
plt.hist(coefs, alpha=0.5)
plt.show()
# ### Weighted Accuracy
np.mean(raw_val_data[y_pred_val==y_val].groupby("poetry_author").count()["poetry_text"]/\
raw_val_data.groupby("poetry_author").count()["poetry_text"])
# ### Looking at misclassified poems
correct_poems = np.random.choice(raw_val_data[y_pred_val==y_val]["poetry_text"],3,replace=False)
for poem in correct_poems:
print("poem \n", poem)
incorrect_poems = np.random.choice(raw_val_data[y_pred_val!=y_val]["poetry_text"],3,replace=False)
for poem in incorrect_poems:
print("poem \n", poem)
raw_train_data[y_pred_train==y_train].groupby("poetry_author").count()["poetry_text"]
raw_train_data.groupby("poetry_author").count()["poetry_text"]
raw_train_data[y_pred_train==y_train].groupby("poetry_author").count()["poetry_text"]/raw_train_data.groupby("poetry_author").count()["poetry_text"]
# It classifies <NAME> and <NAME> and <NAME> perfectly. The worst is <NAME>.
correct_pred_val = raw_val_data[y_pred_val==y_val]
correct_pred_train = raw_train_data[y_pred_train==y_train]
incorrect_pred_val = raw_val_data[y_pred_val!=y_val]
incorrect_pred_train = raw_train_data[y_pred_train!=y_train]
correct_pred_val.groupby("poetry_author").count()["poetry_text"]/\
raw_val_data.groupby("poetry_author").count()["poetry_text"]
list(set(correct_pred_val["poetry_author"]))
print(np.mean(list(map(len,correct_pred_val["poetry_text"]))))
print(np.std(list(map(len,correct_pred_val["poetry_text"]))))
plt.hist(list(map(len,correct_pred_val["poetry_text"])), alpha=0.5, color='blue')
plt.hist(list(map(len,incorrect_pred_val["poetry_text"])), alpha=0.5, color= "red")
print(np.mean(list(map(len,incorrect_pred["poetry_text"]))))
print(np.std(list(map(len,incorrect_pred["poetry_text"]))))
print(np.mean(list(map(len,correct_pred_train["poetry_text"]))))
print(np.std(list(map(len,correct_pred_train["poetry_text"]))))
print(np.mean(list(map(len,incorrect_pred_train["poetry_text"]))))
print(np.std(list(map(len,incorrect_pred_train["poetry_text"]))))
# In general it has a harder time with shorter poems
raw_train_data.groupby("poetry_author").count()["poetry_text"]
import seaborn as sn
from sklearn.metrics import confusion_matrix
author_labels = list(y_val.value_counts().index)
# +
fig, ax = plt.subplots()
sn.heatmap(confusion_matrix(y_true = y_val, y_pred = y_pred_val, labels = author_labels),
annot = True,
xticklabels = author_labels,
yticklabels = author_labels,
cbar = False,
ax = ax)
ax.set_title("Confusion Matrix for Naive Bayes")
ax.set_xlabel("Predicted")
ax.set_ylabel("True")
# -
|
code/03-notebooks/Naive Bayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Logistic Regression for M&Ms
#
# Let's learn about logistic regression with candy! See the [accompanying episode]() for more information.
#
# First, we're going to import some Python packages.
# +
# display plots interactively
# %matplotlib notebook
# import matplotlib for plotting
import matplotlib.pyplot as plt
# optionally: import seaborn for making plots prettier
# you can comment this out if you don't have seaborn installed
import seaborn as sns
sns.set_style('whitegrid')
# numerical Python!
import numpy as np
# table reading fun
import pandas as pd
# -
# Let's first load out sweets data, which you should find in the `data` directory:
datafile = "../data/sweets_data_200611.tsv"
data = pd.read_csv(datafile, sep="\t")
data.head()
# First let's separate out our _labels_, i.e. the type of candy we're looking at:
labels = data["Type of Candy"]
# And we'll also separate out our _features_, i.e. the measurements we've taken:
data.columns
features = data[['has_m', 'has_s', 'length', 'width', 'height','colour']]
# Right now, we're only interested in peanut and plain M&Ms, so we're going to make a new
# array that has only those entries in it:
features_small = features[(labels == "peanut m&m") | (labels == "plain m&m")]
labels_small = np.array(labels[(labels == "peanut m&m") | (labels == "plain m&m")])
features_small.head()
# As you can see in the table above, our labels are currently in text form, that is, when I recorded the data, I wrote down in words what I was measuring, whether it was a plain M&M or a skittle or another type of candy.
# For logistic regression, we're going to turn our text labels for plain and meanut M&Ms into a numerical label, i.e. 0 and 1, like so:
#
# plain m&m = 0
# peanut m&m = 1
# +
labels_small[labels_small == "plain m&m"] = 0.0
labels_small[labels_small == "peanut m&m"] = 1.0
labels_small = np.array(labels_small, dtype=int)
# -
labels_small
# Great! Now we can plot a graph of the length, and the corresponding label:
# +
fig, ax = plt.subplots(1, 1, figsize=(8,4))
ax.scatter(features_small["length"], labels_small, s=20, c="black", marker="x")
ax.set_xlabel("height [mm]")
ax.set_ylabel("Class [0 = plain M&m, 1 = peanut M&M]")
# -
# ### The Step Function
#
# The function that may seem most appropriate is the step function:
def step_function(x, b):
"""
Function which will take the value 0 for
x <= b, and 1 for x > b
"""
y = np.zeros_like(x)
y[x > b] = 1.0
return y
# +
min_x = -1
max_x = 4
x = np.linspace(min_x, max_x, 1000)
a = 1.5
y = step_function(x, a)
fig, ax = plt.subplots(1, 1, figsize=(8,4))
ax.scatter(features_small["length"], labels_small, s=20, c="black", marker="x")
ax.plot(x, y, lw=2, color="red")
ax.set_xlabel("feature x")
ax.set_ylabel("y(x)")
# -
# We can see that peanut M&Ms are on average longer than plain M&Ms. This is perhaps not surprising, since they need to fit a peanut inside of them, and plain M&Ms are generally smaller and rounder.
#
# ### The Logistic Function
#
# Before we can try to draw a function through our data points on this graph, let's first define the logistic function:
def logistic(x, a, b):
"""
Logistic function.
Parameters
----------
x : float or iterable
A single value or list of features.
a, b: floats
Parameters for the linear model.
"""
z = a * x + b
y = 1.0 / (1. + np.exp(-z))
return y
# Before we see what this looks like let's plot just the straight line part of this:
def straight_line(x, a, b):
return a * x + b
# +
a = 1.0
b = -1.0
# make a list of equally spaced points between
# -5 and 5
min_x = 1.0
max_x = 3.0
x = np.linspace(min_x, max_x, 1000)
y = straight_line(x, a, b)
fig, ax = plt.subplots(1, 1, figsize=(8,4))
ax.scatter(features_small["length"], labels_small, s=20, c="black", marker="x")
ax.plot(x, y, lw=2, color="red")
ax.set_xlabel("height [mm]")
ax.set_ylabel("Class [0 = plain M&m, 1 = peanut M&M]")
# -
# Let's see what this looks like for `a=1` and `b=0`:
# +
a = 1.0
b = 0.0
# make a list of equally spaced points between
# -5 and 5
min_x = -10
max_x = 10
x = np.linspace(min_x, max_x, 1000)
y = logistic(x, a, b)
# +
fig, ax = plt.subplots(1, 1, figsize=(8,4))
ax.plot(x, y, lw=2, color="red")
ax.set_xlabel("feature x")
ax.set_ylabel("probability p(y | x)")
# -
# **Exercise**: Experiment with different values for `a` and `b`, and see how the shape of this function changes. What effect does changing `a` and `b` have on the shape of the function?
#
# Hint: If you change `a` and `b` and end up with a funny looking plot try changing the minimum and maximum values for x in `linspace`, `min_x` and `max_x` to expand the range of values being plotted.
#
#
# Let's now go back to our data, and plot both our M&Ms and the logistic function in a single plot:
# +
# make the overall plot
fig, ax = plt.subplots(1, 1, figsize=(6,4))
# add the data points for peanut and plain M&Ms
ax.scatter(features_small["length"], labels_small, s=20, c="black", marker="x")
ax.set_xlabel("height [mm]")
ax.set_ylabel("Class [0 = plain M&m, 1 = peanut M&M]")
# now let's make a logistic function in the range where we have data:
a = 60
b = -90
# make a list of equally spaced points between
# -5 and 5
min_x = np.min(features_small['length'])
max_x = np.max(features_small["length"])
x = np.linspace(min_x, max_x, 1000)
y = logistic(x, a, b)
ax.plot(x, y, lw=2, color="red")
# -
# Well, that doesn't look super great yet!
#
# **Exercise**: Based on your explorations above, can you find values for `a` and `b` that make the function go through the data points?
#
# +
# Solution
# a = 50
# b = -76.0
# -
# ### Classification with Two Features
#
# Let's now take a look at the case where we have more than one feature. So far, we've done all of this by hand, but that's not very practical when you add more features, because each feature adds a parameter, and a _dimension_ to your problem. This makes visualizing the data and model as we've done above more difficult, and it also makes it much harder to find good values for the parameters.
#
# We're going to use a standard library, `scikit-learn`, to help us find good values for `a` and `b`.
from sklearn.linear_model import LogisticRegression
# First, let's pull out the length and width of our M&Ms:
X = np.array(features_small[["length", "width"]])
# Let's see where they fall on a plot:
# +
fig, ax = plt.subplots(1, 1, figsize=(7,7))
ax.scatter(X[labels_small == 0,0], X[labels_small == 0,1], color="blue", s=20, marker="o", label="plain M&Ms")
ax.scatter(X[labels_small == 1,0], X[labels_small == 1,1], color="red", s=20, marker="x", label="peanut M&Ms")
ax.set_xlabel("length")
ax.set_ylabel("width")
ax.legend()
# -
# You can see that the plain M&Ms all cluster in the left side of the plot, and the peanut M&Ms on the right side.
#
# **Exercise**: Hold up a pen or a ruler to the screen and see if you can position it such that it makes a line separating the plain and the peanut M&Ms. Can you do it? If you use one of the other features (e.g. height), does that change your ability to separate the two classes?
#
# ### Fitting a Model
#
# Let's now fit our model to the data. We're going to use the [`LogisticRegression`](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LogisticRegression.html) model in `scikit-learn`:
# +
# set up the model
# it has lots of parameters, but let's not worry about those now
lr = LogisticRegression()
# you can have the computer estimate the parameters by using the `fit`
# function, and the features and labels for our data:
lr.fit(X, labels_small)
# -
# The parameters for our model, i.e. the slopes $a_1$ and $a_2$, and the intercept $b$ can be accessed by calling the `coef_` and `intercept_` keywords:
lr.intercept_
lr.coef_
# Let's calculate and draw the decision boundary. This is the line in the 2D space of our features where $p(y | X) = 0.5$:
# +
x = np.linspace(1.25, 1.75, 1000)
dec_boundary = -(lr.coef_[0,0] * x + lr.intercept_ ) / lr.coef_[0,1]
# +
fig, ax = plt.subplots(1, 1, figsize=(7,7))
ax.scatter(X[labels_small == 0,0], X[labels_small == 0,1], color="blue", s=20, marker="o", label="plain M&Ms")
ax.scatter(X[labels_small == 1,0], X[labels_small == 1,1], color="red", s=20, marker="x", label="peanut M&Ms")
ax.plot(x, dec_boundary, lw=2, color="black")
ax.set_xlabel("length")
ax.set_ylabel("width")
ax.legend()
# -
# That looks pretty good! It definitely separates out our two classes. Hooray! We've done machine learning!
|
code/ML_LogisticRegression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_mxnet_p36
# language: python
# name: conda_mxnet_p36
# ---
# # Image classification training with image format demo
#
# 1. [Introduction](#Introduction)
# 2. [Prerequisites and Preprocessing](#Prequisites-and-Preprocessing)
# 1. [Permissions and environment variables](#Permissions-and-environment-variables)
# 2. [Prepare the data](#Prepare-the-data)
# 3. [Fine-tuning The Image Classification Model](#Fine-tuning-the-Image-classification-model)
# 1. [Training parameters](#Training-parameters)
# 2. [Start the training](#Start-the-training)
# 4. [Inference](#Inference)
# ## Introduction
#
# Welcome to our end-to-end example of the image classification algorithm training with image format. In this demo, we will use the Amazon sagemaker image classification algorithm in transfer learning mode to fine-tune a pre-trained model (trained on imagenet data) to learn to classify a new dataset. In particular, the pre-trained model will be fine-tuned using [caltech-256 dataset](http://www.vision.caltech.edu/Image_Datasets/Caltech256/).
#
# To get started, we need to set up the environment with a few prerequisite steps, for permissions, configurations, and so on.
# ## Prequisites and Preprocessing
#
# ### Permissions and environment variables
#
# Here we set up the linkage and authentication to AWS services. There are three parts to this:
#
# * The roles used to give learning and hosting access to your data. This will automatically be obtained from the role used to start the notebook
# * The S3 bucket that you want to use for training and model data
# * The Amazon sagemaker image classification docker image which need not be changed
# +
# %%time
import sagemaker
from sagemaker import get_execution_role
role = get_execution_role()
print(role)
sess = sagemaker.Session()
bucket = sess.default_bucket()
prefix = "ic-lstformat"
# +
from sagemaker.amazon.amazon_estimator import get_image_uri
training_image = get_image_uri(sess.boto_region_name, "image-classification", repo_version="latest")
print(training_image)
# -
#
# ### Prepare the data
# The caltech 256 dataset consist of images from 257 categories (the last one being a clutter category) and has 30k images with a minimum of 80 images and a maximum of about 800 images per category.
#
# The image classification algorithm can take two types of input formats. The first is a [RecordIO format](https://mxnet.incubator.apache.org/tutorials/basic/record_io.html) (content type: application/x-recordio) and the other is a [lst format](https://mxnet.incubator.apache.org/how_to/recordio.html?highlight=im2rec) (content type: application/x-image). Files for both these formats are available at http://data.dmlc.ml/mxnet/data/caltech-256/. In this example, we will use the lst format for training and use the training/validation split [specified here](http://data.dmlc.ml/mxnet/data/caltech-256/).
# +
import os
import urllib.request
def download(url):
filename = url.split("/")[-1]
if not os.path.exists(filename):
urllib.request.urlretrieve(url, filename)
# Caltech-256 image files
download("http://www.vision.caltech.edu/Image_Datasets/Caltech256/256_ObjectCategories.tar")
# !tar -xf 256_ObjectCategories.tar
# Tool for creating lst file
download("https://raw.githubusercontent.com/apache/incubator-mxnet/master/tools/im2rec.py")
# + language="bash"
#
# mkdir -p caltech_256_train_60
# for i in 256_ObjectCategories/*; do
# c=`basename $i`
# mkdir -p caltech_256_train_60/$c
# for j in `ls $i/*.jpg | shuf | head -n 60`; do
# mv $j caltech_256_train_60/$c/
# done
# done
#
# python im2rec.py --list --recursive caltech-256-60-train caltech_256_train_60/
# python im2rec.py --list --recursive caltech-256-60-val 256_ObjectCategories/
# -
# A .lst file is a tab-separated file with three columns that contains a list of image files. The first column specifies the image index, the second column specifies the class label index for the image, and the third column specifies the relative path of the image file. The image index in the first column should be unique across all of the images. Here we make an image list file using the [im2rec](https://github.com/apache/incubator-mxnet/blob/master/tools/im2rec.py) tool from MXNet. You can also create the .lst file in your own way. An example of .lst file is shown as follows.
# !head -n 3 ./caltech-256-60-train.lst > example.lst
f = open("example.lst", "r")
lst_content = f.read()
print(lst_content)
# When you are bringing your own image files to train, please ensure that the .lst file follows the same format as described above. In order to train with the lst format interface, passing the lst file for both training and validation in the appropriate format is mandatory. Once we have the data available in the correct format for training, the next step is to upload the image and .lst file to S3 bucket.
# +
# Four channels: train, validation, train_lst, and validation_lst
s3train = "s3://{}/{}/train/".format(bucket, prefix)
s3validation = "s3://{}/{}/validation/".format(bucket, prefix)
s3train_lst = "s3://{}/{}/train_lst/".format(bucket, prefix)
s3validation_lst = "s3://{}/{}/validation_lst/".format(bucket, prefix)
# upload the image files to train and validation channels
# !aws s3 cp caltech_256_train_60 $s3train --recursive --quiet
# !aws s3 cp 256_ObjectCategories $s3validation --recursive --quiet
# upload the lst files to train_lst and validation_lst channels
# !aws s3 cp caltech-256-60-train.lst $s3train_lst --quiet
# !aws s3 cp caltech-256-60-val.lst $s3validation_lst --quiet
# -
# Now we have all the data stored in S3 bucket. The image and lst files will be converted to RecordIO file internelly by the image classification algorithm. But if you want do the conversion, the following cell shows how to do it using the [im2rec](https://github.com/apache/incubator-mxnet/blob/master/tools/im2rec.py) tool. Note that this is just an example of creating RecordIO files. We are **_not_** using them for training in this notebook. More details on creating RecordIO files can be found in this [tutorial](https://mxnet.incubator.apache.org/how_to/recordio.html?highlight=im2rec).
# + language="bash"
# python im2rec.py --resize 256 --quality 90 --num-thread 16 caltech-256-60-val 256_ObjectCategories/
# python im2rec.py --resize 256 --quality 90 --num-thread 16 caltech-256-60-train caltech_256_train_60/
# -
# After you created the RecordIO files, you can upload them to the train and validation channels for training. To train with RecordIO format, you can follow "[Image-classification-fulltraining.ipynb](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/imageclassification_caltech/Image-classification-fulltraining.ipynb)" and "[Image-classification-transfer-learning.ipynb](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/introduction_to_amazon_algorithms/imageclassification_caltech/Image-classification-transfer-learning.ipynb)". Again, we will **_not_** use the RecordIO file for the training. The following sections will only show you how to train a model with images and list files.
# Before training the model, we need to setup the training parameters. The next section will explain the parameters in detail.
# ## Fine-tuning the Image Classification Model
# Now that we are done with all the setup that is needed, we are ready to train our object detector. To begin, let us create a ``sageMaker.estimator.Estimator`` object. This estimator will launch the training job.
# ### Training parameters
# There are two kinds of parameters that need to be set for training. The first one are the parameters for the training job. These include:
#
# * **Training instance count**: This is the number of instances on which to run the training. When the number of instances is greater than one, then the image classification algorithm will run in distributed settings.
# * **Training instance type**: This indicates the type of machine on which to run the training. Typically, we use GPU instances for these training
# * **Output path**: This the s3 folder in which the training output is stored
s3_output_location = "s3://{}/{}/output".format(bucket, prefix)
ic = sagemaker.estimator.Estimator(
training_image,
role,
train_instance_count=1,
train_instance_type="ml.p2.xlarge",
train_volume_size=50,
train_max_run=360000,
input_mode="File",
output_path=s3_output_location,
sagemaker_session=sess,
)
# Apart from the above set of parameters, there are hyperparameters that are specific to the algorithm. These are:
#
# * **num_layers**: The number of layers (depth) for the network. We use 18 in this samples but other values such as 50, 152 can be used.
# * **use_pretrained_model**: Set to 1 to use pretrained model for transfer learning.
# * **image_shape**: The input image dimensions,'num_channels, height, width', for the network. It should be no larger than the actual image size. The number of channels should be same as the actual image.
# * **num_classes**: This is the number of output classes for the new dataset. Imagenet was trained with 1000 output classes but the number of output classes can be changed for fine-tuning. For caltech, we use 257 because it has 256 object categories + 1 clutter class.
# * **num_training_samples**: This is the total number of training samples. It is set to 15240 for caltech dataset with the current split.
# * **mini_batch_size**: The number of training samples used for each mini batch. In distributed training, the number of training samples used per batch will be N * mini_batch_size where N is the number of hosts on which training is run.
# * **epochs**: Number of training epochs.
# * **learning_rate**: Learning rate for training.
# * **top_k**: Report the top-k accuracy during training.
# * **resize**: Resize the image before using it for training. The images are resized so that the shortest side is of this parameter. If the parameter is not set, then the training data is used as such without resizing.
# * **precision_dtype**: Training datatype precision (default: float32). If set to 'float16', the training will be done in mixed_precision mode and will be faster than float32 mode
#
# + isConfigCell=true
ic.set_hyperparameters(
num_layers=18,
use_pretrained_model=1,
image_shape="3,224,224",
num_classes=257,
mini_batch_size=128,
epochs=2,
learning_rate=0.01,
top_k=2,
num_training_samples=15420,
resize=256,
precision_dtype="float32",
)
# -
# ## Input data specification
# Set the data type and channels used for training
# +
train_data = sagemaker.session.s3_input(
s3train,
distribution="FullyReplicated",
content_type="application/x-image",
s3_data_type="S3Prefix",
)
validation_data = sagemaker.session.s3_input(
s3validation,
distribution="FullyReplicated",
content_type="application/x-image",
s3_data_type="S3Prefix",
)
train_data_lst = sagemaker.session.s3_input(
s3train_lst,
distribution="FullyReplicated",
content_type="application/x-image",
s3_data_type="S3Prefix",
)
validation_data_lst = sagemaker.session.s3_input(
s3validation_lst,
distribution="FullyReplicated",
content_type="application/x-image",
s3_data_type="S3Prefix",
)
data_channels = {
"train": train_data,
"validation": validation_data,
"train_lst": train_data_lst,
"validation_lst": validation_data_lst,
}
# -
# ## Start the training
# Start training by calling the fit method in the estimator
ic.fit(inputs=data_channels, logs=True)
# # Inference
#
# ***
#
# A trained model does nothing on its own. We now want to use the model to perform inference. For this example, that means predicting the topic mixture representing a given document. You can deploy the created model by using the deploy method in the estimator
ic_classifier = ic.deploy(initial_instance_count=1, instance_type="ml.m4.xlarge")
# #### Download test image
# +
# !wget -O /tmp/test.jpg http://www.vision.caltech.edu/Image_Datasets/Caltech256/images/008.bathtub/008_0007.jpg
file_name = "/tmp/test.jpg"
# test image
from IPython.display import Image
Image(file_name)
# +
import json
import numpy as np
with open(file_name, "rb") as f:
payload = f.read()
payload = bytearray(payload)
ic_classifier.content_type = "application/x-image"
result = json.loads(ic_classifier.predict(payload))
# the result will output the probabilities for all classes
# find the class with maximum probability and print the class index
index = np.argmax(result)
object_categories = [
"ak47",
"american-flag",
"backpack",
"baseball-bat",
"baseball-glove",
"basketball-hoop",
"bat",
"bathtub",
"bear",
"beer-mug",
"billiards",
"binoculars",
"birdbath",
"blimp",
"bonsai-101",
"boom-box",
"bowling-ball",
"bowling-pin",
"boxing-glove",
"brain-101",
"breadmaker",
"buddha-101",
"bulldozer",
"butterfly",
"cactus",
"cake",
"calculator",
"camel",
"cannon",
"canoe",
"car-tire",
"cartman",
"cd",
"centipede",
"cereal-box",
"chandelier-101",
"chess-board",
"chimp",
"chopsticks",
"cockroach",
"coffee-mug",
"coffin",
"coin",
"comet",
"computer-keyboard",
"computer-monitor",
"computer-mouse",
"conch",
"cormorant",
"covered-wagon",
"cowboy-hat",
"crab-101",
"desk-globe",
"diamond-ring",
"dice",
"dog",
"dolphin-101",
"doorknob",
"drinking-straw",
"duck",
"dumb-bell",
"eiffel-tower",
"electric-guitar-101",
"elephant-101",
"elk",
"ewer-101",
"eyeglasses",
"fern",
"fighter-jet",
"fire-extinguisher",
"fire-hydrant",
"fire-truck",
"fireworks",
"flashlight",
"floppy-disk",
"football-helmet",
"french-horn",
"fried-egg",
"frisbee",
"frog",
"frying-pan",
"galaxy",
"gas-pump",
"giraffe",
"goat",
"golden-gate-bridge",
"goldfish",
"golf-ball",
"goose",
"gorilla",
"grand-piano-101",
"grapes",
"grasshopper",
"guitar-pick",
"hamburger",
"hammock",
"harmonica",
"harp",
"harpsichord",
"hawksbill-101",
"head-phones",
"helicopter-101",
"hibiscus",
"homer-simpson",
"horse",
"horseshoe-crab",
"hot-air-balloon",
"hot-dog",
"hot-tub",
"hourglass",
"house-fly",
"human-skeleton",
"hummingbird",
"ibis-101",
"ice-cream-cone",
"iguana",
"ipod",
"iris",
"jesus-christ",
"joy-stick",
"kangaroo-101",
"kayak",
"ketch-101",
"killer-whale",
"knife",
"ladder",
"laptop-101",
"lathe",
"leopards-101",
"license-plate",
"lightbulb",
"light-house",
"lightning",
"llama-101",
"mailbox",
"mandolin",
"mars",
"mattress",
"megaphone",
"menorah-101",
"microscope",
"microwave",
"minaret",
"minotaur",
"motorbikes-101",
"mountain-bike",
"mushroom",
"mussels",
"necktie",
"octopus",
"ostrich",
"owl",
"palm-pilot",
"palm-tree",
"paperclip",
"paper-shredder",
"pci-card",
"penguin",
"people",
"pez-dispenser",
"photocopier",
"picnic-table",
"playing-card",
"porcupine",
"pram",
"praying-mantis",
"pyramid",
"raccoon",
"radio-telescope",
"rainbow",
"refrigerator",
"revolver-101",
"rifle",
"rotary-phone",
"roulette-wheel",
"saddle",
"saturn",
"school-bus",
"scorpion-101",
"screwdriver",
"segway",
"self-propelled-lawn-mower",
"sextant",
"sheet-music",
"skateboard",
"skunk",
"skyscraper",
"smokestack",
"snail",
"snake",
"sneaker",
"snowmobile",
"soccer-ball",
"socks",
"soda-can",
"spaghetti",
"speed-boat",
"spider",
"spoon",
"stained-glass",
"starfish-101",
"steering-wheel",
"stirrups",
"sunflower-101",
"superman",
"sushi",
"swan",
"swiss-army-knife",
"sword",
"syringe",
"tambourine",
"teapot",
"teddy-bear",
"teepee",
"telephone-box",
"tennis-ball",
"tennis-court",
"tennis-racket",
"theodolite",
"toaster",
"tomato",
"tombstone",
"top-hat",
"touring-bike",
"tower-pisa",
"traffic-light",
"treadmill",
"triceratops",
"tricycle",
"trilobite-101",
"tripod",
"t-shirt",
"tuning-fork",
"tweezer",
"umbrella-101",
"unicorn",
"vcr",
"video-projector",
"washing-machine",
"watch-101",
"waterfall",
"watermelon",
"welding-mask",
"wheelbarrow",
"windmill",
"wine-bottle",
"xylophone",
"yarmulke",
"yo-yo",
"zebra",
"airplanes-101",
"car-side-101",
"faces-easy-101",
"greyhound",
"tennis-shoes",
"toad",
"clutter",
]
print("Result: label - " + object_categories[index] + ", probability - " + str(result[index]))
# -
# #### Clean up
#
# When we're done with the endpoint, we can just delete it and the backing instances will be released.
ic_classifier.delete_endpoint()
|
introduction_to_amazon_algorithms/imageclassification_caltech/Image-classification-lst-format-highlevel.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://www.datacamp.com/community/tutorials/autoencoder-keras-tutorial
# +
# https://github.com/davidflanagan/notMNIST-to-MNIST.git
# -
# # Denoising AutoEncoder
# ## Loading the Data
# The notMNIST dataset is an image recognition dataset of font glypyhs for the letters A through J. It is quite similar to the classic MNIST dataset, which contains images of handwritten digits 0 through 9: in this case, you'll find that the NotMNIST dataset comprises 28x28 grayscale images of 70,000 letters from A - J in total 10 categories, and 6,000 images per category.
#
# Tip: if you want to learn how to implement an Multi-Layer Perceptron (MLP) for classification tasks with the MNIST dataset, check out this tutorial.
#
# The NotMNIST dataset is not predefined in the Keras or the TensorFlow framework, so you'll have to download the data from this source. The data will be downloaded in ubyte.gzip format, but no worries about that just yet! You'll soon learn how to read bytestream formats and convert them into a NumPy array. So, let's get started!
#
# The network will be trained on a Nvidia Tesla K40, so if you train on a GPU and use Jupyter Notebook, you will need to add three more lines of code where you specify CUDA device order and CUDA visible devices using a module called os.
#
# In the code below, you basically set environment variables in the notebook using os.environ. It's good to do the following before initializing Keras to limit Keras backend TensorFlow to use first GPU. If the machine on which you train on has a GPU on 0, make sure to use 0 instead of 1. You can check that by running a simple command on your terminal: for example, nvidia-smi
# +
# import os
# os.environ["CUDA_DEVICE_ORDER"]="PCI_BUS_ID"
# os.environ["CUDA_VISIBLE_DEVICES"]="1" #model will be trained on GPU 1
# -
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices()) # list of DeviceAttributes
# +
import tensorflow as tf
tf.test.is_gpu_available() # True/False
# Or only check for gpu's with cuda support
tf.test.is_gpu_available(cuda_only=True)
# -
import keras
from matplotlib import pyplot as plt
import numpy as np
import gzip
# %matplotlib inline
from keras.layers import Input,Conv2D,MaxPooling2D,UpSampling2D
from keras.models import Model
from keras.optimizers import RMSprop
# Here, you define a function that opens the gzip file, reads the file using bytestream.read(). You pass the image dimension and the total number of images to this function. Then, using np.frombuffer(), you convert the string stored in variable buf into a NumPy array of type float32.
#
# Next, you reshape the array into a three-dimensional array or tensor where the first dimension is number of images, and the second and third dimension being the dimension of the image. Finally, you return the NumPy array data.
# # Loading Data Manually into RAM
import glob
import os
import cv2
# +
# """
# Preparing Data Manually
# """
# output_dataset_path = "F:/Datasets/DIV2K - Watermarks/train/class1"
# output_ground_path = "F:/Datasets/DIV2K - Watermarks/ground_truth/class1"
# # len(filenames_ground)
# filenames = glob.glob(output_dataset_path+"/*")
# filenames_ground = glob.glob(output_ground_path+"/*")
# assert (len(filenames) == len(filenames_ground)), "Shapes arent same"
# x_shape, y_shape, _ = cv2.imread(filenames[0]).shape
# print("shapes: ", len(filenames), ":", len(filenames_ground))
# import pandas as pd
# import os
# filenames_df = pd.Series(filenames)
# filenames_ground_df = pd.Series(filenames_ground)
# filenames_ground_df = filenames_ground_df.apply(lambda x: os.path.basename(x))
# filenames_df = filenames_df.apply(lambda x: os.path.basename(x))
# filenames_ground_df[~filenames_ground_df.isin(filenames_df.values)]
# filenames_df[~filenames_df.isin(filenames_ground_df.values)]
# filenames = filenames
# filenames_ground = filenames_ground
# # cv2.imread(filenames[0]).shape
# X = np.zeros((len(filenames), x_shape, y_shape, 3), dtype=np.float32)
# y = np.zeros((len(filenames_ground), x_shape, y_shape, 3), dtype=np.float32)
# import time
# from IPython.display import clear_output
# total_files = len(filenames)
# start_time = time.time()
# for i, (filename, filename_ground) in enumerate(zip(filenames, filenames_ground)):
# try:
# time_per_iteration = time.time()-start_time
# start_time = time.time()
# clear_output()
# print(f"{(np.round(i/total_files, 4))*100}% completed")
# print(f"{round(time_per_iteration*(total_files-i)/60)} minutes left")
# X[i, ...] = cv2.imread(filename)/255
# y[i, ...] = cv2.imread(filename_ground)/255
# except Exception as e:
# print("Exception")
# break
# -
# # Scaling
# +
# X = (X/np.max(X)).astype(np.float32)
# y = (y/np.max(y)).astype(np.float32)
# +
# np.save("X.npy", X)
# np.save("y.npy", y)
# +
# X = np.load("X.npy")
# y = np.save("y.npy")
# -
# # Train Test Split
# +
# from sklearn.model_selection import train_test_split
# X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.2, random_state=42)
# +
# np.save("X_train.npy", X_train)
# np.save("X_val.npy", X_val)
# np.save("y_train.npy", y_train)
# np.save("y_val.npy", y_val)
# -
base_dir = "F:\\Datasets\\temp\\WatermarkRemoval\\"
X_train = np.load(base_dir+"X_train.npy")
X_val = np.load(base_dir+"X_val.npy")
y_train = np.load(base_dir+"y_train.npy")
y_val = np.load(base_dir+"y_val.npy")
"""
RAM fully consumed. Using Scratch Disks!
will have to delete X and y after saving it in disk
"""
# ## Adding Noise to Images
# ### Note
# that for this task, you don't need training and testing labels. That's why you will pass the training images twice. Your training images will both act as the input as well as the ground truth similar to the labels you have in classification task.
#
# Now you are all set to define the network and feed the data into the network. So without any further ado, let's jump to the next step!
#
# ## The Convolutional Autoencoder
# The images are of size 28 x 28 x 1 or a 784-dimensional vector. You convert the image matrix to an array, rescale it between 0 and 1, reshape it so that it's of size 28 x 28 x 1, and feed this as an input to the network.
#
# Also, you will use a batch size of 128 using a higher batch size of 256 or 512 is also preferable it all depends on the system you train your model. It contributes heavily in determining the learning parameters and affects the prediction accuracy. You will train your network for 50 epochs.
# +
# batch_size = batch_size
# epochs = 50
# x = X_train[0].shape[0]
# y = X_train[0].shape[1]
# inChannel = X_train[0].shape[2]
# input_img = Input(shape = (x, y, inChannel))
# -
# As discussed before, the autoencoder is divided into two parts: there's an encoder and a decoder.
#
# ### Encoder
#
# The first layer will have 32 filters of size 3 x 3, followed by a downsampling (max-pooling) layer,
# The second layer will have 64 filters of size 3 x 3, followed by another downsampling layer,
# The final layer of encoder will have 128 filters of size 3 x 3.
#
#
# ### Decoder
# The first layer will have 128 filters of size 3 x 3 followed by a upsampling layer,/li>
# The second layer will have 64 filters of size 3 x 3 followed by another upsampling layer,
# The final layer of encoder will have 1 filter of size 3 x 3.
# The max-pooling layer will downsample the input by two times each time you use it, while the upsampling layer will upsample the input by two times each time it is used.
#
# ### Note:
# The number of filters, the filter size, the number of layers, number of epochs you train your model, are all hyperparameters and should be decided based on your own intuition, you are free to try new experiments by tweaking with these hyperparameters and measure the performance of your model. And that is how you will slowly learn the art of deep learning!
# # Loading Data
# +
# """
# Pretraining for model validation
# """
# # from keras.preprocessing.image import ImageDataGenerator
# batch_size = 64
# img_height = 200
# img_width = 200
# train_datagen = ImageDataGenerator(
# rescale=1./255,
# shear_range=0.2,
# zoom_range=0.2,
# horizontal_flip=True,
# validation_split=0.2
# ) # set validation split
# train_generator = train_datagen.flow_from_directory(
# "F:/Datasets/DIV2K - Watermarks/train",
# target_size=(img_height, img_width),
# batch_size=batch_size,
# class_mode="input",
# subset='training'
# ) # set as training data
# validation_generator = train_datagen.flow_from_directory(
# "F:/Datasets/DIV2K - Watermarks/train", # same directory as training data
# target_size=(img_height, img_width),
# batch_size=batch_size,
# class_mode="input",
# subset='validation'
# ) # set as validation data
# # train_datagen = ImageDataGenerator(
# # # rescale=1./255,
# # shear_range=0.2,
# # zoom_range=0.2,
# # horizontal_flip=True
# # )
# # train_generator = train_datagen.flow_from_directory(
# # 'F:/Datasets/DIV2K - Watermarks/train',
# # # target_size=(150, 150),
# # batch_size=32,
# # class_mode="input"
# # )
# +
# dir(tf.keras.optimizers)
# 'Adadelta','Adagrad', 'Adam', 'Adamax', 'Ftrl', 'Nadam', 'Optimizer', 'RMSprop', 'SGD'
# +
"""
Note: Training against Testing Data (one to one training)
If the architecture is fine, the loss should go to 0
"""
import time
batch_size = 32
epochs = 50
checkpoint_cb = keras.callbacks.ModelCheckpoint(f"models/{np.round(time.time()).astype(int)}_watermark_removal.h5", save_best_only=True)
def autoencoder(input_img):
#encoder
#input = 28 x 28 x 1 (wide and thin)
conv1 = Conv2D(8, (3, 3), activation='relu', padding='same')(input_img) #28 x 28 x 32
pool1 = MaxPooling2D(pool_size=(2, 2))(conv1) #14 x 14 x 32
conv2 = Conv2D(16, (3, 3), activation='relu', padding='same')(pool1) #14 x 14 x 64
pool2 = MaxPooling2D(pool_size=(2, 2))(conv2) #7 x 7 x 64
conv3 = Conv2D(32, (3, 3), activation='relu', padding='same')(pool2) #7 x 7 x 128 (small and thick)
#decoder
conv4 = Conv2D(32, (3, 3), activation='relu', padding='same')(conv3) #7 x 7 x 128
up1 = UpSampling2D((2,2))(conv4) # 14 x 14 x 128
conv5 = Conv2D(16, (3, 3), activation='relu', padding='same')(up1) # 14 x 14 x 64
up2 = UpSampling2D((2,2))(conv5) # 28 x 28 x 64
decoded = Conv2D(3, (3, 3), activation='sigmoid', padding='same')(up2) # 28 x 28 x 1
return decoded
autoencoder = Model(input_img, autoencoder(input_img))
# -
autoencoder.compile(loss='mean_squared_error', optimizer = keras.optimizers.Adam(learning_rate=0.01))
autoencoder.summary()
# +
# model = tf.keras.models.load_model("1585645914_watermark_removal.h5")
# from keras.models import load_model
# autoencoder = load_model('models/1585645914_watermark_removal.h5')
# -
epochs = 1000
autoencoder_train = autoencoder.fit(
X_train, y_train, batch_size=batch_size,
epochs=epochs,verbose=1,
validation_data=(X_val, y_val),
callbacks=[checkpoint_cb]
)
# autoencoder_train = autoencoder.fit(x_train_noisy, train_X, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(x_valid_noisy, valid_X))
dir(keras.optimizers)
# +
# import time
# checkpoint_cb = keras.callbacks.ModelCheckpoint(f"models/watermark_model_{np.round(time.time()).astype(int)}.h5", save_best_only=True)
# autoencoder.fit_generator(
# train_generator,
# steps_per_epoch = train_generator.samples // batch_size,
# validation_data = validation_generator,
# validation_steps = validation_generator.samples // batch_size,
# epochs = 50)
# +
# autoencoder_train = autoencoder.fit(
# X_train, y_train, batch_size=batch_size,
# epochs=epochs,verbose=1,
# validation_data=(X_val, y_val),
# callbacks=[checkpoint_cb]
# )
# # autoencoder_train = autoencoder.fit(x_train_noisy, train_X, batch_size=batch_size,epochs=epochs,verbose=1,validation_data=(x_valid_noisy, valid_X))
# -
fig_no = 567
pred = autoencoder.predict(np.expand_dims(X_val[fig_no], axis=0))
plt.subplots(figsize=(14,7))
plt.subplot(131)
plt.title("Watermarked Image")
plt.imshow(X_val[fig_no][:,:,::-1])
plt.subplot(132)
plt.title("Original Image")
plt.imshow(y_val[fig_no][:,:,::-1])
plt.subplot(133)
plt.title("Predicted Image")
plt.imshow(pred[0][:,:,::-1])
plt.show()
# # Training vs Validation Loss Plot
loss = autoencoder_train.history['loss']
val_loss = autoencoder_train.history['val_loss']
epochs = range(epochs)
plt.figure()
plt.plot(epochs, loss, 'bo', label='Training loss')
plt.plot(epochs, val_loss, 'b', label='Validation loss')
plt.title('Training and validation loss')
plt.legend()
plt.show()
# # Predicting on Test Data
pred = autoencoder.predict(np.expand_dims(X[0], axis=0))
plt.imshow(pred[0][:,:,::-1])
# +
plt.figure(figsize=(20, 4))
print("Test Images")
for i in range(10,20,1):
plt.subplot(2, 10, i+1)
plt.imshow(test_data[i, ..., 0], cmap='gray')
curr_lbl = test_labels[i]
plt.title("(Label: " + str(label_dict[curr_lbl]) + ")")
plt.show()
plt.figure(figsize=(20, 4))
print("Test Images with Noise")
for i in range(10,20,1):
plt.subplot(2, 10, i+1)
plt.imshow(x_test_noisy[i, ..., 0], cmap='gray')
plt.show()
plt.figure(figsize=(20, 4))
print("Reconstruction of Noisy Test Images")
for i in range(10,20,1):
plt.subplot(2, 10, i+1)
plt.imshow(pred[i, ..., 0], cmap='gray')
plt.show()
# -
|
WatermarkRemoval/AutoEncoder-watermark.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.0
# language: julia
# name: julia-0.6
# ---
Mod = include("../src/EventFiltering.jl")
using Mod.EventFiltering
# To compose more complex, repeating patterns of events, two event trains can be convolved:
# +
F = ExponentialFilter(0.5)
e1 = EventTrain([0.0, 5.0, 10.0], [1, 2, 0.5])
e2 = 2*EventTrain([0.1, 0.2, 0.5, 0.6])
e3 = e1 ∘ (F ∘ e2)
# -
# Let's plot the result:
using Plots
plot(e1, color=:red, linewidth=3)
plot!(e3, 0.0, 20.0, show_events=true, show_kernels=true)
# As we can see, the pattern defined by `e2`, convolved by `F`, repeats at every point defined by `e1`.
# Notice how each event is scaled according to `e1` and `e2`!
|
docs/complex_events.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Data Science lies at the intersection of :
# - Hacking Skills
# - Math and Statistics Knowledge
# - Substantive Expertise
#
# ### Python is well suited for learning Data Science :
# - It's Free.
# - It's relatively easy to code in and, in particular, to understand.
# - It has lots of useful data science related libraries.
# >Python uses *__indentation__* to delimit the execution of block of code
# +
#example
for i in [1,2]:
print(i)
for j in [1,2]:
print(j)
print(i+j)
print(i)
print("Loop completed")
# +
#indentation is ignored while working with inside parentheses and brackets
bracket_example = (1 + 2 + 3 + 4 + 5 + 6 + 7)
bracket_example
# +
#indentation also makes our code easier to read
list_of_lists = [[1, 2, 3], [4, 5, 6], [7, 8, 9]]
easier_to_read_list_of_lists = [[1, 2, 3],
[4, 5, 6],
[7, 8, 9]]
print(list_of_lists)
print(easier_to_read_list_of_lists)
# -
# > We can use a *__backslash ( \ )__* to indicate that a statement continues onto the next line.
a = 2 + \
3
print(a)
# ### Modules
# - Modules is collection of statements and definitions.
# - Modules are used to break larger programs into small manageable and organized files.
# - One can create modules on their own(module = example.py & module_name = example) or you can even install & import third party modules.
import matplotlib.pyplot as plt
# > Let's create one module on our own
# def add(a,b):
# """
# This module is used to add two numbers
# """
# result = a + b
# return result
import example
example.add(5,4)
# ### Functions
# - A function is a set or block of code which runs only when it is called.
# - *__def__* keyword is used to define a function.
# - A function takes inputs which are known as parameters and gives out outputs which is result.
# - Defining a function for writing any program is a good programming habit.
# - A function also contains docstrings which basically tells us about the use of the function.
def multiply_by_two(x):
"""
This function will multiply the input by 2
"""
result = x * 2
return result
multiply_by_two(25)
def double(x):
"""
Multiply x by 2
"""
return x*2
# > Python functions are amazing. We can assign them to variables and then pass variable into the functions.
def apply_to_one(f):
"""
Calls the function f with 1 as its argument.
"""
return f(1)
#refers to the previously defined function
my_double = double
x = apply_to_one(my_double)
print(x)
# > We can also create functions using Lambda
y = apply_to_one(lambda x:x+4)
print(y)
# > Function parameters can also be given default arguments which only need to be specified when you want a value other than the default.
def my_message(message="Hello World!"):
"""
This function will print the message when called.
"""
print(message)
my_message()
my_message("Hello World! Let's code in python")
def full_name(first="John",last="Doe"):
"""
This function is for displaying first name and last name
"""
print(first + " " + last)
full_name()
full_name("Sudhanshu","Mukherjee")
full_name(first="Dudh")
|
1 - Introduction, Modules and Functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (nlu)
# language: python
# name: nlu
# ---
# # Homework and bake-off: word-level entailment with neural networks
__author__ = "<NAME>"
__version__ = "CS224u, Stanford, Fall 2020"
# ## Contents
#
# 1. [Overview](#Overview)
# 1. [Set-up](#Set-up)
# 1. [Data](#Data)
# 1. [Baseline](#Baseline)
# 1. [Representing words: vector_func](#Representing-words:-vector_func)
# 1. [Combining words into inputs: vector_combo_func](#Combining-words-into-inputs:-vector_combo_func)
# 1. [Classifier model](#Classifier-model)
# 1. [Baseline results](#Baseline-results)
# 1. [Homework questions](#Homework-questions)
# 1. [Hypothesis-only baseline [2 points]](#Hypothesis-only-baseline-[2-points])
# 1. [Alternatives to concatenation [2 points]](#Alternatives-to-concatenation-[2-points])
# 1. [A deeper network [2 points]](#A-deeper-network-[2-points])
# 1. [Your original system [3 points]](#Your-original-system-[3-points])
# 1. [Bake-off [1 point]](#Bake-off-[1-point])
# ## Overview
# The general problem is word-level natural language inference. Training examples are pairs of words $(w_{L}, w_{R}), y$ with $y = 1$ if $w_{L}$ entails $w_{R}$, otherwise $0$.
#
# The homework questions below ask you to define baseline models for this and develop your own system for entry in the bake-off, which will take place on a held-out test-set distributed at the start of the bake-off. (Thus, all the data you have available for development is available for training your final system before the bake-off begins.)
# ## Set-up
# See [the first notebook in this unit](nli_01_task_and_data.ipynb) for set-up instructions.
from collections import defaultdict
import json
import numpy as np
import os
import pandas as pd
from torch_shallow_neural_classifier import TorchShallowNeuralClassifier
import nli
import utils
# +
DATA_HOME = 'data'
NLIDATA_HOME = os.path.join(DATA_HOME, 'nlidata')
wordentail_filename = os.path.join(
NLIDATA_HOME, 'nli_wordentail_bakeoff_data.json')
GLOVE_HOME = os.path.join(DATA_HOME, 'glove.6B')
# -
# ## Data
#
# I've processed the data into a train/dev split that is designed to put some pressure on our models to actually learn these semantic relations, as opposed to exploiting regularities in the sample.
#
# The defining feature of the dataset is that the `train` and `dev` __vocabularies__ are disjoint. That is, if a word `w` appears in a training pair, it does not occur in any text pair. It follows from this that there are also no word-pairs shared between train and dev, as you would expect. This should require your models to learn abstract relationships, as opposed to memorizing incidental properties of individual words in the dataset.
with open(wordentail_filename) as f:
wordentail_data = json.load(f)
# The keys are the splits plus a list giving the vocabulary for the entire dataset:
wordentail_data.keys()
wordentail_data['train'][: 5]
nli.get_vocab_overlap_size(wordentail_data)
# Because no words are shared between `train` and `dev`, no pairs are either:
nli.get_pair_overlap_size(wordentail_data)
# Here is the label distribution:
pd.DataFrame(wordentail_data['train']).head()
pd.DataFrame(wordentail_data['train'])[1].value_counts()
# This is a challenging label distribution – there are more than 5 times as more non-entailment cases as entailment cases.
# ## Baseline
# Even in deep learning, __feature representation is vital and requires care!__ For our task, feature representation has two parts: representing the individual words and combining those representations into a single network input.
# ### Representing words: vector_func
# Let's consider two baseline word representations methods:
#
# 1. Random vectors (as returned by `utils.randvec`).
# 1. 50-dimensional GloVe representations.
def randvec(w, n=50, lower=-1.0, upper=1.0):
"""Returns a random vector of length `n`. `w` is ignored."""
return utils.randvec(n=n, lower=lower, upper=upper)
# +
def load_glove50():
glove_src = os.path.join(GLOVE_HOME, 'glove.6B.50d.txt')
# Creates a dict mapping strings (words) to GloVe vectors:
GLOVE = utils.glove2dict(glove_src)
return GLOVE
GLOVE = load_glove50()
def glove_vec(w):
"""Return `w`'s GloVe representation if available, else return
a random vector."""
return GLOVE.get(w, randvec(w, n=50))
# -
# ### Combining words into inputs: vector_combo_func
# Here we decide how to combine the two word vectors into a single representation. In more detail, where `u` is a vector representation of the left word and `v` is a vector representation of the right word, we need a function `vector_combo_func` such that `vector_combo_func(u, v)` returns a new input vector `z` of dimension `m`. A simple example is concatenation:
def vec_concatenate(u, v):
"""Concatenate np.array instances `u` and `v` into a new np.array"""
return np.concatenate((u, v))
# `vector_combo_func` could instead be vector average, vector difference, etc. (even combinations of those) – there's lots of space for experimentation here; [homework question 2](#Alternatives-to-concatenation-[2-points]) below pushes you to do some exploration.
# ### Classifier model
#
# For a baseline model, I chose `TorchShallowNeuralClassifier`:
net = TorchShallowNeuralClassifier(early_stopping=True)
# ### Baseline results
#
# The following puts the above pieces together, using `vector_func=glove_vec`, since `vector_func=randvec` seems so hopelessly misguided for our problem!
baseline_experiment = nli.wordentail_experiment(
train_data=wordentail_data['train'],
assess_data=wordentail_data['dev'],
model=net,
vector_func=glove_vec,
vector_combo_func=vec_concatenate)
# ## Homework questions
#
# Please embed your homework responses in this notebook, and do not delete any cells from the notebook. (You are free to add as many cells as you like as part of your responses.)
# ### Hypothesis-only baseline [2 points]
#
# During our discussion of SNLI and MultiNLI, we noted that a number of research teams have shown that hypothesis-only baselines for NLI tasks can be remarkably robust. This question asks you to explore briefly how this baseline affects our task.
#
# For this problem, submit two functions:
#
# 1. A `vector_combo_func` function called `hypothesis_only` that simply throws away the premise, using the unmodified hypothesis (second) vector as its representation of the example.
#
# 1. A function called `run_hypothesis_only_evaluation` that does the following:
# 1. Loops over the two `vector_combo_func` values `vec_concatenate` and `hypothesis_only`, calling `nli.wordentail_experiment` to train on the 'train' portion and assess on the 'dev' portion, with `glove_vec` as the `vector_func`. So that the results are consistent, use an `sklearn.linear_model.LogisticRegression` with default parameters as the model.
# 1. Returns a `dict` mapping `function_name` strings to the 'macro-F1' score for that pair, as returned by the call to `nli.wordentail_experiment`. (Tip: you can get the `str` name of, e.g., `hypothesis_only` with `hypothesis_only.__name__`.)
#
# The functions `test_hypothesis_only` and `test_run_hypothesis_only_evaluation` will help ensure that your functions have the desired logic.
# +
from sklearn.linear_model import LogisticRegression
def hypothesis_only(u, v):
return v
def run_hypothesis_only_evaluation():
result = dict()
lr = LogisticRegression()
for combo in [hypothesis_only, vec_concatenate]:
experiment = nli.wordentail_experiment(
train_data=wordentail_data['train'],
assess_data=wordentail_data['dev'],
model=lr,
vector_func=glove_vec,
vector_combo_func=combo)
result[combo.__name__] = experiment['macro-F1']
return result
# -
def test_hypothesis_only(hypothesis_only):
v = hypothesis_only(1, 2)
assert v == 2
test_hypothesis_only(hypothesis_only)
def test_run_hypothesis_only_evaluation(run_hypothesis_only_evaluation):
results = run_hypothesis_only_evaluation()
assert all(x in results for x in ('hypothesis_only', 'vec_concatenate')), \
("The return value of `run_hypothesis_only_evaluation` does not "
"have the intended kind of keys.")
assert isinstance(results['vec_concatenate'], float), \
("The values of the `run_hypothesis_only_evaluation` result "
"should be floats.")
test_run_hypothesis_only_evaluation(run_hypothesis_only_evaluation)
# ### Alternatives to concatenation [2 points]
#
# We've so far just used vector concatenation to represent the premise and hypothesis words. This question asks you to explore two simple alternative:
#
# 1. Write a function `vec_diff` that, for a given pair of vector inputs `u` and `v`, returns the element-wise difference between `u` and `v`.
#
# 1. Write a function `vec_max` that, for a given pair of vector inputs `u` and `v`, returns the element-wise max values between `u` and `v`.
#
# You needn't include your uses of `nli.wordentail_experiment` with these functions, but we assume you'll be curious to see how they do!
# +
def vec_diff(u, v):
##### YOUR CODE HERE
return u - v
def vec_max(u, v):
##### YOUR CODE HERE
return np.maximum(u, v)
# -
def test_vec_diff(vec_diff):
u = np.array([10.2, 8.1])
v = np.array([1.2, -7.1])
result = vec_diff(u, v)
expected = np.array([9.0, 15.2])
assert np.array_equal(result, expected), \
"Expected {}; got {}".format(expected, result)
test_vec_diff(vec_diff)
def test_vec_max(vec_max):
u = np.array([1.2, 8.1])
v = np.array([10.2, -7.1])
result = vec_max(u, v)
expected = np.array([10.2, 8.1])
assert np.array_equal(result, expected), \
"Expected {}; got {}".format(expected, result)
test_vec_max(vec_max)
# ### A deeper network [2 points]
#
# It is very easy to subclass `TorchShallowNeuralClassifier` if all you want to do is change the network graph: all you have to do is write a new `build_graph`. If your graph has new arguments that the user might want to set, then you should also redefine `__init__` so that these values are accepted and set as attributes.
#
# For this question, please subclass `TorchShallowNeuralClassifier` so that it defines the following graph:
#
# $$\begin{align}
# h_{1} &= xW_{1} + b_{1} \\
# r_{1} &= \textbf{Bernoulli}(1 - \textbf{dropout_prob}, n) \\
# d_{1} &= r_1 * h_{1} \\
# h_{2} &= f(d_{1}) \\
# h_{3} &= h_{2}W_{2} + b_{2}
# \end{align}$$
#
# Here, $r_{1}$ and $d_{1}$ define a dropout layer: $r_{1}$ is a random binary vector of dimension $n$, where the probability of a value being $1$ is given by $1 - \textbf{dropout_prob}$. $r_{1}$ is multiplied element-wise by our first hidden representation, thereby zeroing out some of the values. The result is fed to the user's activation function $f$, and the result of that is fed through another linear layer to produce $h_{3}$. (Inside `TorchShallowNeuralClassifier`, $h_{3}$ is the basis for a softmax classifier; no activation function is applied to it because the softmax scaling is handled internally by the loss function.)
#
# For your implementation, please use `nn.Sequential`, `nn.Linear`, and `nn.Dropout` to define the required layers.
#
# For comparison, using this notation, `TorchShallowNeuralClassifier` defines the following graph:
#
# $$\begin{align}
# h_{1} &= xW_{1} + b_{1} \\
# h_{2} &= f(h_{1}) \\
# h_{3} &= h_{2}W_{2} + b_{2}
# \end{align}$$
#
# The following code starts this sub-class for you, so that you can concentrate on `build_graph`. Be sure to make use of `self.dropout_prob`.
#
# For this problem, submit just your completed `TorchDeepNeuralClassifier`. You needn't evaluate it, though we assume you will be keen to do that!
#
# You can use `test_TorchDeepNeuralClassifier` to ensure that your network has the intended structure.
# +
import torch.nn as nn
class TorchDeepNeuralClassifier(TorchShallowNeuralClassifier):
def __init__(self, dropout_prob=0.7, **kwargs):
self.dropout_prob = dropout_prob
super().__init__(**kwargs)
def build_graph(self):
"""Complete this method!
Returns
-------
an `nn.Module` instance, which can be a free-standing class you
write yourself, as in `torch_rnn_classifier`, or the outpiut of
`nn.Sequential`, as in `torch_shallow_neural_classifier`.
"""
##### YOUR CODE HERE
return nn.Sequential(
nn.Linear(self.input_dim, self.hidden_dim),
nn.Dropout(p=self.dropout_prob),
self.hidden_activation,
nn.Linear(self.hidden_dim, self.n_classes_))
# -
def test_TorchDeepNeuralClassifier(TorchDeepNeuralClassifier):
dropout_prob = 0.55
assert hasattr(TorchDeepNeuralClassifier(), "dropout_prob"), \
"TorchDeepNeuralClassifier must have an attribute `dropout_prob`."
try:
inst = TorchDeepNeuralClassifier(dropout_prob=dropout_prob)
except TypeError:
raise TypeError("TorchDeepNeuralClassifier must allow the user "
"to set `dropout_prob` on initialization")
inst.input_dim = 10
inst.n_classes_ = 5
graph = inst.build_graph()
assert len(graph) == 4, \
"The graph should have 4 layers; yours has {}".format(len(graph))
expected = {
0: 'Linear',
1: 'Dropout',
2: 'Tanh',
3: 'Linear'}
for i, label in expected.items():
name = graph[i].__class__.__name__
assert label in name, \
("The {} layer of the graph should be a {} layer; "
"yours is {}".format(i, label, name))
assert graph[1].p == dropout_prob, \
("The user's value for `dropout_prob` should be the value of "
"`p` for the Dropout layer.")
test_TorchDeepNeuralClassifier(TorchDeepNeuralClassifier)
# ### Your original system [3 points]
#
# This is a simple dataset, but its "word-disjoint" nature ensures that it's a challenging one, and there are lots of modeling strategies one might adopt.
#
# You are free to do whatever you like. We require only that your system differ in some way from those defined in the preceding questions. They don't have to be completely different, though. For example, you might want to stick with the model but represent examples differently, or the reverse.
#
# You are free to use different pretrained word vectors and the like.
#
# Please embed your code in this notebook so that we can rerun it.
#
# In the cell below, please provide a brief technical description of your original system, so that the teaching team can gain an understanding of what it does. This will help us to understand your code and analyze all the submissions to identify patterns and strategies. We also ask that you report the best score your system got during development, just to help us understand how systems performed overall.
# +
# PLEASE MAKE SURE TO INCLUDE THE FOLLOWING BETWEEN THE START AND STOP COMMENTS:
# 1) Textual description of your system.
# 2) The code for your original system.
# 3) The score achieved by your system in place of MY_NUMBER.
# With no other changes to that line.
# You should report your score as a decimal value <=1.0
# PLEASE MAKE SURE NOT TO DELETE OR EDIT THE START AND STOP COMMENTS
# NOTE: MODULES, CODE AND DATASETS REQUIRED FOR YOUR ORIGINAL SYSTEM
# SHOULD BE ADDED BELOW THE 'IS_GRADESCOPE_ENV' CHECK CONDITION. DOING
# SO ABOVE THE CHECK MAY CAUSE THE AUTOGRADER TO FAIL.
# START COMMENT: Enter your system description in this cell.
# My peak score was: MY_NUMBER
if 'IS_GRADESCOPE_ENV' not in os.environ:
from nltk.corpus import wordnet as wn
import torch.nn as nn
utils.fix_random_seeds()
class TorchDeepNCMultipleLayers(TorchShallowNeuralClassifier):
def __init__(self, nhidden=1, **kwargs):
"""
Generalisation of TorchShallowNeuralClassifier with multiple hidden layers. Each hidden layers keeps the
dimension of the hidden layer has defined in the super class.
:param nhidden: number of hidden layers
"""
super().__init__(**kwargs)
self.nhidden = nhidden
def build_graph(self):
"""
Returns
-------
an `nn.Module` instance, which can be a free-standing class you
write yourself, as in `torch_rnn_classifier`, or the output of
`nn.Sequential`, as in `torch_shallow_neural_classifier`.
"""
graph_list = [nn.Linear(self.input_dim, self.hidden_dim)]
graph_list.append(self.hidden_activation)
for i in range(self.nhidden - 1):
graph_list.append(nn.Linear(self.hidden_dim, self.hidden_dim))
graph_list.append(self.hidden_activation)
graph_list.append(nn.Linear(self.hidden_dim, self.n_classes_))
graph = nn.Sequential(*tuple(graph_list))
return graph
def wordnet_features(word1, word2, methodname):
"""
Returns 1 if the synsets extracted from the hierarchy methodname of the premise intersect the synsets of the
hypothesis. Otherwise returns -1
:param word1: word in premise
:param word2: word in hypothesis
:param methodname: wordnet hierarchy considered (hypernym or hyponym)
:return:
"""
try:
hyps = [h for ss in wn.synsets(word1) for h in getattr(ss, methodname)()]
except AttributeError:
hyps = list()
for ss in wn.synsets(word1):
try:
for h in getattr(ss.lemmas()[0], methodname)():
hyps.append(h)
except TypeError:
pass
syns = wn.synsets(word2)
output = 1 if set(hyps) & set(syns) else -1
return output
def word_entail_featurize_wordnet_1(data, vector_func, vector_combo_func, hierarchy, adjustment_param):
"""
Modified version of the featurize function used in nli.wordentail_experiment. Uses WordNet hierarchy.
:param data:
:param vector_func:
:param vector_combo_func:
:param hierarchy: hierarchy in wordnet considered, 'hypernyms' or 'hyponyms'
:param adjustment_param: Parameters by which we translate the vector output from vector_combo_func when
the hypothesis is included in the wordnet hierarchy of word1 (hypernyms or hyponyms)
:return:
"""
X = []
y = []
for (w1, w2), label in data:
is_related = list()
for h in hierarchy:
is_related.append(wordnet_features(w1, w2, h))
rep = vector_combo_func(vector_func(w1), vector_func(w2))
# build a vector of length vector_func(w1) with info from wordnet
vec_size = vector_func(w1).shape[0]
hierarchy_size = vec_size // len(hierarchy)
coeff_vec = np.array([x for item in is_related for x in [item] * hierarchy_size])
remaining_items = vec_size % len(hierarchy)
if remaining_items > 0:
# makes sure that coeff_vec has the right size
coeff_vec = np.concatenate((coeff_vec, np.zeros(remaining_items)))
adj_vec = coeff_vec * adjustment_param
rep = np.concatenate((rep, adj_vec))
X.append(rep)
y.append(label)
return X, y
def word_entail_featurize_wordnet(hierarchy, param):
"""
Wrapper of word_entail_featurize_wordnet_1, allowing to define parameters and adjustment_param
in the feature function.
"""
return lambda *args, **kwargs: word_entail_featurize_wordnet_1(*args, hierarchy=hierarchy,
adjustment_param=param, **kwargs)
def nli_model():
"""
Original system
"""
glove_dic = GLOVE
df = pd.DataFrame.from_dict(glove_dic, orient='index')
sum_rows = df.sum(axis=1)
# calculate param, average distance between the mean and the max/min sum of row values of the Glove embedded space.
# will be used in word_entail_featurize_wordnet to adjust vectors when they share an hypernym.
n = df.shape[1]
mean, max1, min1 = sum_rows.mean() / n, sum_rows.max() / n, sum_rows.min() / n
param = 0.25 * (abs(min1) + abs(max1)) + mean
featurize_func = word_entail_featurize_wordnet(['hypernyms', 'hyponyms', 'member_holonyms', 'antonyms', 'antonyms',
'derivationally_related_forms', 'pertainyms'], param)
model = TorchDeepNCMultipleLayers(hidden_dim=150, nhidden=3, batch_size=300, eta=0.001, hidden_activation=nn.Tanh())
experiment = nli.wordentail_experiment(
train_data=wordentail_data['train'],
assess_data=wordentail_data['dev'],
model=model,
vector_func=glove_vec,
vector_combo_func=vec_concatenate,
featurize_func=featurize_func)
return experiment
model_result = nli_model()
# STOP COMMENT: Please do not remove this comment.
# -
# ## Bake-off [1 point]
#
# The goal of the bake-off is to achieve the highest __macro-average F1__ score on a test set that we will make available at the start of the bake-off. The announcement will go out on the discussion forum. To enter, you'll be asked to run `nli.bake_off_evaluation` on the output of your chosen `nli.wordentail_experiment` run.
#
# The cells below this one constitute your bake-off entry.
#
# The rules described in the [Your original system](#Your-original-system-[3-points]) homework question are also in effect for the bake-off.
#
# Systems that enter will receive the additional homework point, and systems that achieve the top score will receive an additional 0.5 points. We will test the top-performing systems ourselves, and only systems for which we can reproduce the reported results will win the extra 0.5 points.
#
# Late entries will be accepted, but they cannot earn the extra 0.5 points. Similarly, you cannot win the bake-off unless your homework is submitted on time.
#
# The announcement will include the details on where to submit your entry.
# Enter your bake-off assessment code into this cell.
# Please do not remove this comment.
if 'IS_GRADESCOPE_ENV' not in os.environ:
pass
# Please enter your code in the scope of the above conditional.
##### YOUR CODE HERE
# On an otherwise blank line in this cell, please enter
# your macro-avg f1 value as reported by the code above.
# Please enter only a number between 0 and 1 inclusive.
# Please do not remove this comment.
if 'IS_GRADESCOPE_ENV' not in os.environ:
pass
# Please enter your score in the scope of the above conditional.
##### YOUR CODE HERE
|
hw_wordentail-training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2019 The TensorFlow Authors.
#
#
# + colab_type="code" id="tuOe1ymfHZPu" colab={} cellView="form"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # 추정기(Estimator)를 사용한 다중 워커(Multi-worker) 훈련
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/distribute/multi_worker_with_estimator"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />TensorFlow.org에서 보기</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/tutorials/distribute/multi_worker_with_estimator.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />구글 코랩(Colab)에서 실행하기</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/tutorials/distribute/multi_worker_with_estimator.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />깃허브(GitHub) 소스 보기</a>
# </td>
# </table>
# + [markdown] id="hYK9tHrcUfio" colab_type="text"
# Note: 이 문서는 텐서플로 커뮤니티에서 번역했습니다. 커뮤니티 번역 활동의 특성상 정확한 번역과 최신 내용을 반영하기 위해 노력함에도 불구하고 [공식 영문 문서](https://github.com/tensorflow/docs/blob/master/site/en/tutorials/distribute/multi_worker_with_estimator.ipynb)의 내용과 일치하지 않을 수 있습니다. 이 번역에 개선할 부분이 있다면 [tensorflow/docs](https://github.com/tensorflow/docs) 깃허브 저장소로 풀 리퀘스트를 보내주시기 바랍니다. 문서 번역이나 리뷰에 참여하려면 [<EMAIL>](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ko)로 메일을 보내주시기 바랍니다."
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## 개요
#
# Note: `tf.distribute` API와 함께 추정기를 사용할 수는 있지만, `tf.distribute`와 함께 케라스(Keras)를 사용하는 것을 추천합니다. [케라스를 사용한 다중 워커(Multi-worker) 훈련](../../guide/multi_worker_with_keras.ipynb)을 봐주세요. `tf.distribute.Strategy`를 추정기와 사용하는 것은 부분적으로만 지원하고 있습니다.
#
# 이 튜토리얼은 `tf.estimator`와 함께 분산 다중 워커 훈련을 하기 위하여 `tf.distribute.Strategy`를 어떻게 사용하는지 살펴봅니다. `tf.estimator`를 사용하여 코드를 작성하고 있고, 고성능의 장비 한 대로 다룰 수 있는 것보다 더 큰 작업을 하는 데에 관심이 있으시다면 이 튜토리얼이 알맞습니다.
#
# 시작하기 전에, [텐서플로로 분산 훈련하기](../../guide/distributed_training.ipynb)를 먼저 읽어주세요. [다중 GPU 훈련 튜토리얼](./keras.ipynb)도 관련이 있습니다. 이 튜토리얼과 같은 모델을 사용합니다.
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## 설정
#
# 먼저, 텐서플로를 설정하고 필요한 패키지들을 가져옵니다.
# + colab_type="code" id="IqR2PQG4ZaZ0" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
# + colab_type="code" id="bnYxvfLD-LW-" colab={}
try:
# # %tensorflow_version 기능은 코랩에서만 사용할 수 있습니다.
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow_datasets as tfds
import tensorflow as tf
tfds.disable_progress_bar()
import os, json
# + [markdown] colab_type="text" id="hPBuZUNSZmrQ"
# ## 입력 함수
#
# 이 튜토리얼은 [텐서플로 데이터셋(TensorFlow Datasets)](https://www.tensorflow.org/datasets)의 MNIST 데이터셋을 사용합니다. 코드 내용은 [다중 GPU 훈련 튜토리얼](./keras.ipynb)과 유사하지만 큰 차이점이 하나 있습니다. 바로 추정기를 써서 다중 워커 훈련을 할 때는 데이터셋을 워커 숫자대로 나누어 주어야 모델이 수렴합니다. 입력 데이터는 워커 인덱스로 샤딩(shard)합니다. 그러면 각 워커 프로세스가 데이터셋을 `1/워커 수` 만큼씩 겹치지 않게 나누어 갖습니다.
# + colab_type="code" id="dma_wUAxZqo2" colab={}
BUFFER_SIZE = 10000
BATCH_SIZE = 64
def input_fn(mode, input_context=None):
datasets, info = tfds.load(name='mnist',
with_info=True,
as_supervised=True)
mnist_dataset = (datasets['train'] if mode == tf.estimator.ModeKeys.TRAIN else
datasets['test'])
def scale(image, label):
image = tf.cast(image, tf.float32)
image /= 255
return image, label
if input_context:
mnist_dataset = mnist_dataset.shard(input_context.num_input_pipelines,
input_context.input_pipeline_id)
return mnist_dataset.map(scale).cache().shuffle(BUFFER_SIZE).batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="4BlcVXMhB59T"
# 훈련을 수렴시키기 위한 또 다른 방법으로 각 워커에서 데이터셋을 제각기 다른 시드 값으로 셔플하는 것도 있습니다.
# + [markdown] colab_type="text" id="8YFpxrcsZ2xG"
# ## 다중 워커 설정
#
# [다중 GPU 훈련 튜토리얼](./keras.ipynb)과 비교할 때 가장 큰 차이 중 하나는 다중 워커를 설정하는 부분입니다. `TF_CONFIG` 환경 변수는 클러스터를 이루는 각 워커에 클러스터 설정을 지정하는 표준 방법입니다.
#
# `TF_CONFIG`에는 `cluster`와 `task`라는 두 가지 구성요소가 있습니다. `cluster`는 전체 클러스터, 다시 말해 클러스터에 속한 워커와 파라미터 서버에 대한 정보를 제공합니다. `task`는 현재 작업에 대한 정보를 지정합니다. 이 예제에서는 작업의 `type`이 `worker`이고, 작업의 `index`는 `0`입니다.
#
# 예를 들기 위하여, 이 튜토리얼에서는 두 개의 워커를 localhost에 띄울 때의 `TF_CONFIG`를 보여드리겠습니다. 실제로는 각 워커를 다른 장비에서 띄울 텐데, 실제 IP 주소와 포트를 할당하고, 그에 맞게 TF_CONFIG를 지정해야 합니다. 예를 들어, 각 장비의 작업 `index`가 달라야 합니다.
#
# 주의: 아래 코드를 코랩에서 실행하지 마십시오. 텐서플로 런타임이 주어진 IP와 포트로 gRPC 서버를 띄우려고 할 텐데, 아마도 실패할 것입니다.
#
# ```
# os.environ['TF_CONFIG'] = json.dumps({
# 'cluster': {
# 'worker': ["localhost:12345", "localhost:23456"]
# },
# 'task': {'type': 'worker', 'index': 0}
# })
# ```
# + [markdown] colab_type="text" id="qDreJzTffAP5"
# ## 모델 정의하기
#
# 훈련을 위하여 레이어와 옵티마이저, 손실 함수를 정의하세요. 이 튜토리얼에서는 [다중 GPU 훈련 튜토리얼](./keras.ipynb)과 비슷하게 케라스 레이어로 모델을 정의합니다.
# + colab_type="code" id="WNvOn_OeiUYC" colab={}
LEARNING_RATE = 1e-4
def model_fn(features, labels, mode):
model = tf.keras.Sequential([
tf.keras.layers.Conv2D(32, 3, activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D(),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(10)
])
logits = model(features, training=False)
if mode == tf.estimator.ModeKeys.PREDICT:
predictions = {'logits': logits}
return tf.estimator.EstimatorSpec(labels=labels, predictions=predictions)
optimizer = tf.compat.v1.train.GradientDescentOptimizer(
learning_rate=LEARNING_RATE)
loss = tf.keras.losses.SparseCategoricalCrossentropy(
from_logits=True, reduction=tf.keras.losses.Reduction.NONE)(labels, logits)
loss = tf.reduce_sum(loss) * (1. / BATCH_SIZE)
if mode == tf.estimator.ModeKeys.EVAL:
return tf.estimator.EstimatorSpec(mode, loss=loss)
return tf.estimator.EstimatorSpec(
mode=mode,
loss=loss,
train_op=optimizer.minimize(
loss, tf.compat.v1.train.get_or_create_global_step()))
# + [markdown] colab_type="text" id="P94PrIW_kSCE"
# Note: 이 예제에서는 학습률이 고정되어있습니다. 하지만 실제로는 전역 배치 크기에 따라 학습률을 조정해야 할 수 있습니다.
# + [markdown] colab_type="text" id="UhNtHfuxCGVy"
# ## MultiWorkerMirroredStrategy
#
# 모델을 훈련하기 위하여 `tf.distribute.experimental.MultiWorkerMirroredStrategy`의 인스턴스를 사용하세요. `MultiWorkerMirroredStrategy`는 모든 워커의 각 장비에, 모델의 레이어에 있는 모든 변수의 복사본을 만듭니다. 이 전략은 `CollectiveOps`라는 수집을 위한 통신용 텐서플로 연산을 사용하여 그래디언트를 모으고, 변수들의 값을 동일하게 맞춥니다. [텐서플로로 분산 훈련하기](../../guide/distributed_training.ipynb)에 이 전략에 대한 더 자세한 내용이 있습니다.
# + colab_type="code" id="1uFSHCJXMrQ-" colab={}
strategy = tf.distribute.experimental.MultiWorkerMirroredStrategy()
# + [markdown] colab_type="text" id="H47DDcOgfzm7"
# ## 모델 훈련 및 평가하기
#
# 다음으로, 추정기의 `RunConfig`에 분산 전략을 지정하십시오. 그리고 `tf.estimator.train_and_evaluate`로 훈련 및 평가를 합니다. 이 튜토리얼에서는 `train_distribute`로만 전략을 지정하였기 때문에 훈련 과정만 분산 처리합니다. `eval_distribute`를 지정하여 평가도 분산 처리할 수 있습니다.
# + colab_type="code" id="BcsuBYrpgnlS" colab={}
config = tf.estimator.RunConfig(train_distribute=strategy)
classifier = tf.estimator.Estimator(
model_fn=model_fn, model_dir='/tmp/multiworker', config=config)
tf.estimator.train_and_evaluate(
classifier,
train_spec=tf.estimator.TrainSpec(input_fn=input_fn),
eval_spec=tf.estimator.EvalSpec(input_fn=input_fn)
)
# + [markdown] colab_type="text" id="XVk4ftYx6JAO"
# ## 훈련 성능 최적화하기
#
# 이제 모델과 `tf.distribute.Strategy`로 만든 다중 워커를 사용할 수 있는 추정기가 있습니다. 다중 워커 훈련 성능을 최적화하려면 다음과 같은 방법을 사용해 보십시오.
#
# * *배치 크기 늘리기:* 여기서 지정하는 배치 크기는 GPU당 크기입니다. 일반적으로, GPU 메모리 크기에 맞는 한 가장 크게 배치 크기를 잡는 것이 좋습니다.
# * *변수 형변환:* 가능하면 변수를 `tf.float` 타입으로 바꾸세요. 공식 ResNet 모델의 [예제](https://github.com/tensorflow/models/blob/8367cf6dabe11adf7628541706b660821f397dce/official/resnet/resnet_model.py#L466)에서 어떻게 변환하는지 볼 수 있습니다.
# * *집합 통신 구현을 사용하세요:* `MultiWorkerMirroredStrategy`는 여러 가지 [집합 통신 구현](https://github.com/tensorflow/tensorflow/blob/master/tensorflow/python/distribute/cross_device_ops.py)을 제공합니다.
# * `RING`은 장비 간 통신을 위하여 gRPC를 써서 링 네트워크 기반의 집합 통신을 구현한 것입니다.
# * `NCCL`은 [Nvidia의 NCCL](https://developer.nvidia.com/nccl)을 사용하여 수집 연산을 구현한 것입니다.
# * `AUTO`는 런타임이 알아서 고르도록 합니다.
#
# 어떤 집합 구현이 가장 좋은지는 GPU의 숫자와 종류, 클러스터 장비 간 네트워크 연결 등에 따라 다를 수 있습니다. 런타임 자동 선택을 오버라이드하려면, `MultiWorkerMirroredStrategy` 생성자의 `communication` 인자에 적절한 값을 주면 됩니다. 예를 들어 `communication=tf.distribute.experimental.CollectiveCommunication.NCCL`과 같이 주면 됩니다.
# + [markdown] colab_type="text" id="AW0Hb2xM6EGX"
# ## 다른 코드 예제
#
# 1. [처음부터 끝까지 살펴보는 예제](https://github.com/tensorflow/ecosystem/tree/master/distribution_strategy)에서는 tensorflow/ecosystem의 쿠버네티스(Kubernetes) 템플릿을 이용하여 다중 워커를 사용하여 훈련합니다. 이 예제에서는 케라스 모델을 만든 후, `tf.keras.estimator.model_to_estimator` API를 이용하여 추정기 모델로 변환합니다.
# 2. 다중 분산 전략으로 실행할 수 있는 공식 [ResNet50](https://github.com/tensorflow/models/blob/master/official/resnet/imagenet_main.py) 모델.
|
site/ko/tutorials/distribute/multi_worker_with_estimator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img align="left" src="https://ithaka-labs.s3.amazonaws.com/static-files/images/tdm/tdmdocs/CC_BY.png"><br />
#
# Created by [<NAME>](http://nkelber.com) and Ted Lawless for [JSTOR Labs](https://labs.jstor.org/) under [Creative Commons CC BY License](https://creativecommons.org/licenses/by/4.0/)<br />
# For questions/comments/improvements, email <EMAIL>.<br />
# ___
#
# # Latent Dirichlet Allocation (LDA) Topic Modeling
#
# **Description:**
# This [notebook](https://docs.constellate.org/key-terms/#jupyter-notebook) demonstrates how to do topic modeling. The following processes are described:
#
# * Using the `constellate` client to retrieve a dataset
# * Filtering based on a pre-processed ID list
# * Filtering based on a [stop words list](https://docs.constellate.org/key-terms/#stop-words)
# * Cleaning the tokens in the dataset
# * Creating a [gensim dictionary](https://docs.constellate.org/key-terms/#gensim-dictionary)
# * Creating a [gensim](https://docs.constellate.org/key-terms/#gensim) [bag of words](https://docs.constellate.org/key-terms/#bag-of-words) [corpus](https://docs.constellate.org/key-terms/#corpus)
# * Computing a topic list using [gensim](https://docs.constellate.org/key-terms/#gensim)
# * Visualizing the topic list with `pyldavis`
#
# **Use Case:** For Researchers (Mostly code without explanation, not ideal for learners)
#
# **Difficulty:** Intermediate
#
# **Completion time:** 30 minutes
#
# **Knowledge Required:**
# * Python Basics Series ([Start Python Basics I](./python-basics-1.ipynb))
#
# **Knowledge Recommended:**
# * [Exploring Metadata](./metadata.ipynb)
# * [Working with Dataset Files](./working-with-dataset-files.ipynb)
# * [Pandas I](./pandas-1.ipynb)
# * [Creating a Stopwords List](./creating-stopwords-list.ipynb)
# * A familiarity with [gensim](https://docs.constellate.org/key-terms/#gensim) is helpful but not required.
#
# **Data Format:** [JSON Lines (.jsonl)](https://docs.constellate.org/key-terms/#jsonl)
#
# **Libraries Used:**
# * [constellate](https://docs.constellate.org/key-terms/#tdm-client) client to collect, unzip, and read our dataset
# * [pandas](https://constellate.org/docs/key-terms/#pandas) to load a preprocessing list
# * `csv` to load a custom stopwords list
# * [gensim](https://docs.constellate.org/key-terms/#gensim) to accomplish the topic modeling
# * [NLTK](https://docs.constellate.org/key-terms/#nltk) to create a stopwords list (if no list is supplied)
# * `pyldavis` to visualize our topic model
#
# **Research Pipeline**
# 1. Build a dataset
# 2. Create a "Pre-Processing CSV" with [Exploring Metadata](./exploring-metadata.ipynb) (Optional)
# 3. Create a "Custom Stopwords List" with [Creating a Stopwords List](./creating-stopwords-list.ipynb) (Optional)
# 4. Complete the Topic Modeling analysis with this notebook
# ____
# ## What is Topic Modeling?
#
# **Topic modeling** is a **machine learning** technique that attempts to discover groupings of words (called topics) that commonly occur together in a body of texts. The body of texts could be anything from journal articles to newspaper articles to tweets.
#
# **Topic modeling** is an unsupervised, clustering technique for text. We give the machine a series of texts that it then attempts to cluster the texts into a given number of topics. There is also a *supervised*, clustering technique called **Topic Classification**, where we supply the machine with examples of pre-labeled topics and then see if the machine can identify them given the examples.
#
# **Topic modeling** is usually considered an exploratory technique; it helps us discover new patterns within a set of texts. **Topic Classification**, using labeled data, is intended to be a predictive technique; we want it to find more things like the examples we give it.
# ## Import your dataset
# We'll use the `constellate` client library to automatically retrieve the dataset in the JSON file format.
#
# Enter a [dataset ID](https://docs.constellate.org/key-terms/#dataset-ID) in the next code cell.
#
# If you don't have a dataset ID, you can:
# * Use the sample dataset ID already in the code cell
# * [Create a new dataset](https://constellate.org/builder)
# * [Use a dataset ID from other pre-built sample datasets](https://constellate.org/dataset/dashboard)
# +
# Creating a variable `dataset_id` to hold our dataset ID
# The default dataset is Independent Voices
# Independent Voices is an open access digital collection of alternative press newspapers, magazines and journals,
# drawn from the special collections of participating libraries. These periodicals were produced by feminists,
# dissident GIs, campus radicals, Native Americans, anti-war activists, Black Power advocates, Hispanics,
# LGBT activists, the extreme right-wing press and alternative literary magazines
# during the latter half of the 20th century.
dataset_id = "ac017df6-06b0-d415-7235-98fb729dac4e"
# -
# Next, import the `constellate` client, passing the `dataset_id` as an argument using the `get_dataset` method.
# +
# Importing your dataset with a dataset ID
import constellate
# Pull in the sampled dataset (1500 documents) that matches `dataset_id`
# in the form of a gzipped JSON lines file.
# The .get_dataset() method downloads the gzipped JSONL file
# to the /data folder and returns a string for the file name and location
dataset_file = constellate.get_dataset(dataset_id)
# To download the full dataset (up to a limit of 25,000 documents),
# request it first in the builder environment. See the Constellate Client
# documentation at: https://constellate.org/docs/constellate-client
# Then use the `constellate.download` method show below.
#dataset_file = constellate.download(dataset_id, 'jsonl')
# -
# ## Apply Pre-Processing Filters (if available)
# If you completed pre-processing with the "Exploring Metadata and Pre-processing" notebook, you can use your CSV file of dataset IDs to automatically filter the dataset. Your pre-processed CSV file must be in the root folder.
# +
# Import a pre-processed CSV file of filtered dataset IDs.
# If you do not have a pre-processed CSV file, the analysis
# will run on the full dataset and may take longer to complete.
import pandas as pd
import os
pre_processed_file_name = f'data/pre-processed_{dataset_id}.csv'
if os.path.exists(pre_processed_file_name):
df = pd.read_csv(pre_processed_file_name)
filtered_id_list = df["id"].tolist()
use_filtered_list = True
print('Pre-Processed CSV found. Successfully read in ' + str(len(df)) + ' documents.')
else:
use_filtered_list = False
print('No pre-processed CSV file found. Full dataset will be used.')
# -
# ## Load Stopwords List
#
# If you have created a stopword list in the stopwords notebook, we will import it here. (You can always modify the CSV file to add or subtract words then reload the list.) Otherwise, we'll load the NLTK [stopwords](https://docs.constellate.org/key-terms/#stop-words) list automatically.
# +
# Load a custom data/stop_words.csv if available
# Otherwise, load the nltk stopwords list in English
# The filename of the custom data/stop_words.csv file
stopwords_list_filename = 'data/stop_words.csv'
if os.path.exists(stopwords_list_filename):
import csv
with open(stopwords_list_filename, 'r') as f:
stop_words = list(csv.reader(f))[0]
print('Custom stopwords list loaded from CSV')
else:
# Load the NLTK stopwords list
from nltk.corpus import stopwords
stop_words = stopwords.words('english')
print('NLTK stopwords list loaded')
# -
list(stop_words)
# ## Define a Function to Process Tokens
# Next, we create a short function to clean up our tokens.
def process_token(token):
token = token.lower()
if token in stop_words:
return
if len(token) < 4:
return
if not(token.isalpha()):
return
return token
# +
# Limit to n documents. Set to None to use all documents.
limit = 5000
n = 0
documents = []
for document in constellate.dataset_reader(dataset_file):
processed_document = []
document_id = document["id"]
if use_filtered_list is True:
# Skip documents not in our filtered_id_list
if document_id not in filtered_id_list:
continue
unigrams = document.get("unigramCount", {})
for gram, count in unigrams.items():
clean_gram = process_token(gram)
if clean_gram is None:
continue
processed_document += [clean_gram] * count # Add the unigram as many times as it was counted
if len(processed_document) > 0:
documents.append(processed_document)
if n % 1000 == 0:
print(f'Unigrams collected for {n} documents...')
n += 1
if (limit is not None) and (n >= limit):
break
print(f'All unigrams collected for {n} documents.')
# -
# Build a gensim dictionary corpus and then train the model. More information about parameters can be found at the [Gensim LDA Model page](https://radimrehurek.com/gensim/models/ldamodel.html).
import gensim
dictionary = gensim.corpora.Dictionary(documents)
doc_count = len(documents)
num_topics = 10 # Change the number of topics
passes = 8 # The number of passes the model runs
# Remove terms that appear in less than 50 documents and terms that occur in more than 90% of documents.
dictionary.filter_extremes(no_below=50, no_above=0.9)
bow_corpus = [dictionary.doc2bow(doc) for doc in documents]
# Train the LDA model
model = gensim.models.LdaModel(
corpus=bow_corpus,
id2word=dictionary,
num_topics=num_topics,
passes=passes
)
# ## Display a List of Topics
# Print the most significant terms, as determined by the model, for each topic.
for topic_num in range(0, num_topics):
word_ids = model.get_topic_terms(topic_num)
words = []
for wid, weight in word_ids:
word = dictionary.id2token[wid]
words.append(word)
print("Topic {}".format(str(topic_num).ljust(5)), " ".join(words))
# ## Visualize the Topic Distances on a Flat Plane
#
# Visualize the model using [`pyLDAvis`](https://pyldavis.readthedocs.io/en/latest/). This visualization can take a while to generate depending on the size of your dataset.
import pyLDAvis.gensim
pyLDAvis.enable_notebook()
pyLDAvis.gensim.prepare(model, bow_corpus, dictionary)
# Export this visualization as an HTML file
# An internet connection is still required to view the HTML
p = pyLDAvis.gensim.prepare(model, bow_corpus, dictionary)
pyLDAvis.save_html(p, 'my_visualization.html')
|
topic-modeling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append("..")
from rosalind_tools.config import *
from typing import List
# Given: Six nonnegative integers, each of which does not exceed 20,000. The integers correspond to the number of couples in a population possessing each genotype pairing for a given factor. In order, the six given integers represent the number of couples having the following genotypes:
#
# 1. AA-AA
# 2. AA-Aa
# 3. AA-aa
# 4. Aa-Aa
# 5. Aa-aa
# 6. aa-aa
# Return: The expected number of offspring displaying the dominant phenotype in the next generation, under the assumption that every couple has exactly two offspring.
def expected_offspring(couples: List[int]) -> int:
# probablity of offspring displaying the dominant phenotype from genotypes of couples listed above
# [AA-AA, AA-Aa, AA-aa, Aa-Aa, Aa-aa, aa-aa]
prob = [1.00, 1.00, 1.00, 0.75, 0.5, 0.00]
expected = 0
for n, p in zip(couples, prob):
expected += n * p
print(expected * 2)
return
# Try sample dataset
data = '1 0 0 1 0 1'
couples = map(int, data.split(' '))
expected_offspring(couples)
# Try Rosalind dataset
with open(data_dir/"rosalind_iev.txt", 'r') as f:
couples = map(int, f.readline().rstrip().split(' '))
expected_offspring(couples)
|
notebooks/calculating_expected_offspring.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
import matplotlib.pyplot as plt
def display(image):
plt.imshow(image, cmap='gray')
plt.axis('off')
plt.show()
# +
image = cv2.imread("images/coins.jpg")
gray = cv2.cvtColor(image, cv2.COLOR_BGR2GRAY)
display(gray)
# +
# binary image
_, thresh = cv2.threshold(gray, 0, 255, cv2.THRESH_BINARY_INV+cv2.THRESH_OTSU)
display(thresh)
# +
# remove noise
kernel = np.ones((3,3), np.uint8)
opening = cv2.morphologyEx(thresh, cv2.MORPH_OPEN, kernel, iterations=2)
display(opening)
# +
sure_bg = cv2.dilate(opening,kernel, iterations=3)
display(sure_bg)
# +
dist_transform = cv2.distanceTransform(opening, cv2.DIST_L2, 5)
display(dist_transform)
# +
ret, sure_fg = cv2.threshold(dist_transform, 0.7*dist_transform.max(), 255, 0)
sure_fg
# -
sure_bg.dtype
sure_fg = sure_fg.astype('uint8')
sure_fg.dtype
display(sure_fg)
# +
unknown = cv2.subtract(sure_bg, sure_fg)
display(unknown)
# +
# Marker labelling
ret, markers = cv2.connectedComponents(sure_fg)
# Add one to all labels so that sure background is not 0, but 1
markers = markers+1
# Now, mark the region of unknown with zero
markers[unknown==255] = 0
# -
plt.imshow(markers, cmap='jet')
plt.show()
# apply the watershed algorithm
markers = cv2.watershed(image,markers)
plt.imshow(markers, cmap='jet')
plt.axis('off')
plt.show()
|
02-Advance_image_processing/watershed_algorithm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# Have you ever used "squeeze" function before? When should you use it? There are two use cases:
# 1. single column data frame to Series
# 2. single element Series or Index to scalar
# single column data frame to Series
a = pd.DataFrame({'col1':[1,2,3],'col2':[4,5,6],'col3':[7,8,9]})
a
a1 = a.loc[:,['col1']]
# a1
type(a1)
a1.shape
# this is a single column data frame ,which is unnecessary, let's squeeze it to Series
a1.squeeze()
# Second Scenario
# single element Series or Index to scalar
b = pd.Series([2])
b
b.squeeze()
|
pandas/examples/6_squeeze.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
ls1 = [1,5,6,4,1,2,3,5]
ls2 = [1,5,6,5,1,2,3,6]
ls = [1,1,5]
c = 0
r=0
for x in ls:
for y in ls1[r:]:
r+=1
if (x==y):
c+=1
break;
else:
pass
if(c==3):
print("it’s a Match")
else:
print("it’s Gone")
# +
c = 0
r=0
for x in ls:
for y in ls2[r:]:
r+=1
if (x==y):
c+=1
break;
else:
pass
if(c==3):
print("it’s a Match")
else:
print("it’s Gone")
# -
|
list comparing program.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
#hide
from nbdevtest import *
# # Nbdev Test
#
# > just we are testing nbdev
# This file will become your README and also the index of your documentation.
# ## Install
# `pip install nbdevtest`
# ## How to use
# Fill me in please! Don't forget code examples:
1+1
say_hello("JJ")
o = HelloSayer("Alexis")
o.say()
o = HelloSayer("A")
o.say()
C = CCac("Alexis")
C.sh()
|
index.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Homework: going neural (6 pts)
#
# We've checked out statistical approaches to language models in the last notebook. Now let's go find out what deep learning has to offer.
#
# <img src='https://raw.githubusercontent.com/yandexdataschool/nlp_course/master/resources/expanding_mind_lm_kn_3.png' width=300px>
#
# We're gonna use the same dataset as before, except this time we build a language model that's character-level, not word level. Before you go:
# * If you haven't done seminar already, use `seminar.ipynb` to download the data.
# * This homework uses TensorFlow v2.0: this is [how you install it](https://www.tensorflow.org/beta); and that's [how you use it](https://colab.research.google.com/drive/1YtfbZGgzKr7fpBTqkdEQtu4vUALoTv8A).
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# Working on character level means that we don't need to deal with large vocabulary or missing words. Heck, we can even keep uppercase words in text! The downside, however, is that all our sequences just got a lot longer.
#
# However, we still need special tokens:
# * Begin Of Sequence (__BOS__) - this token is at the start of each sequence. We use it so that we always have non-empty input to our neural network. $P(x_t) = P(x_1 | BOS)$
# * End Of Sequence (__EOS__) - you guess it... this token is at the end of each sequence. The catch is that it should __not__ occur anywhere else except at the very end. If our model produces this token, the sequence is over.
#
# +
BOS, EOS = ' ', '\n'
data = pd.read_json("./arxivData.json")
lines = data.apply(lambda row: (row['title'] + ' ; ' + row['summary'])[:512], axis=1) \
.apply(lambda line: BOS + line.replace(EOS, ' ') + EOS) \
.tolist()
# if you missed the seminar, download data here - https://yadi.sk/d/_nGyU2IajjR9-w
# -
# Our next step is __building char-level vocabulary__. Put simply, you need to assemble a list of all unique tokens in the dataset.
# +
# get all unique characters from lines (including capital letters and symbols)
tokens = <YOUR CODE>
tokens = sorted(tokens)
n_tokens = len(tokens)
print ('n_tokens = ',n_tokens)
assert 100 < n_tokens < 150
assert BOS in tokens, EOS in tokens
# -
# We can now assign each character with it's index in tokens list. This way we can encode a string into a TF-friendly integer vector.
# dictionary of character -> its identifier (index in tokens list)
token_to_id = <YOUR CODE>
# +
assert len(tokens) == len(token_to_id), "dictionaries must have same size"
for i in range(n_tokens):
assert token_to_id[tokens[i]] == i, "token identifier must be it's position in tokens list"
print("Seems alright!")
# -
# Our final step is to assemble several strings in a integet matrix `[batch_size, text_length]`.
#
# The only problem is that each sequence has a different length. We can work around that by padding short sequences with extra _EOS_ or cropping long sequences. Here's how it works:
def to_matrix(lines, max_len=None, pad=token_to_id[EOS], dtype='int32'):
"""Casts a list of lines into tf-digestable matrix"""
max_len = max_len or max(map(len, lines))
lines_ix = np.full([len(lines), max_len], pad, dtype=dtype)
for i in range(len(lines)):
line_ix = list(map(token_to_id.get, lines[i][:max_len]))
lines_ix[i, :len(line_ix)] = line_ix
return lines_ix
# +
#Example: cast 4 random names to matrices, pad with zeros
dummy_lines = [
' abc\n',
' abacaba\n',
' abc1234567890\n',
]
print(to_matrix(dummy_lines))
# -
# ### Neural Language Model (2 points including training)
#
# Just like for N-gram LMs, we want to estimate probability of text as a joint probability of tokens (symbols this time).
#
# $$P(X) = \prod_t P(x_t \mid x_0, \dots, x_{t-1}).$$
#
# Instead of counting all possible statistics, we want to train a neural network with parameters $\theta$ that estimates the conditional probabilities:
#
# $$ P(x_t \mid x_0, \dots, x_{t-1}) \approx p(x_t \mid x_0, \dots, x_{t-1}, \theta) $$
#
#
# But before we optimize, we need to define our neural network. Let's start with a fixed-window (aka convolutional) architecture:
#
# <img src='https://raw.githubusercontent.com/yandexdataschool/nlp_course/master/resources/fixed_window_lm.jpg' width=400px>
#
import tensorflow as tf
keras, L = tf.keras, tf.keras.layers
assert tf.__version__.startswith('2'), "Current tf version: {}; required: 2.0.*".format(tf.__version__)
class FixedWindowLanguageModel(L.Layer):
def __init__(self, n_tokens=n_tokens, emb_size=16, hid_size=64):
"""
A fixed window model that looks on at least 5 previous symbols.
Note: fixed window LM is effectively performing a convolution over a sequence of words.
This convolution only looks on current and previous words.
Such convolution can be represented as a sequence of 2 operations:
- pad input vectors by {strides * (filter_size - 1)} zero vectors on the "left", do not pad right
- perform regular convolution with {filter_size} and {strides}
- If you're absolutely lost, here's a hint: use ZeroPadding1D and Conv1D from keras.layers
You can stack several convolutions at once
"""
super().__init__() # initialize base class to track sub-layers, trainable variables, etc.
#YOUR CODE - create layers/variables and any metadata you want, e.g. self.emb = L.Embedding(...)
<...>
#END OF YOUR CODE
def __call__(self, input_ix):
"""
compute language model logits given input tokens
:param input_ix: batch of sequences with token indices, tf tensor: int32[batch_size, sequence_length]
:returns: pre-softmax linear outputs of language model [batch_size, sequence_length, n_tokens]
these outputs will be used as logits to compute P(x_t | x_0, ..., x_{t - 1})
"""
# YOUR CODE - apply layers, see docstring above
return <...>
def get_possible_next_tokens(self, prefix=BOS, temperature=1.0, max_len=100):
""" :returns: probabilities of next token, dict {token : prob} for all tokens """
prefix_ix = tf.convert_to_tensor(to_matrix([prefix]), tf.int32)
probs = tf.nn.softmax(self(prefix_ix)[0, -1]).numpy() # shape: [n_tokens]
return dict(zip(tokens, probs))
# +
model = FixedWindowLanguageModel()
# note: tensorflow and keras layers create variables only after they're first applied (called)
dummy_input_ix = tf.constant(to_matrix(dummy_lines))
dummy_logits = model(dummy_input_ix)
print('Weights:', tuple(w.name for w in model.trainable_variables))
# -
assert isinstance(dummy_logits, tf.Tensor)
assert dummy_logits.shape == (len(dummy_lines), max(map(len, dummy_lines)), n_tokens), "please check output shape"
assert np.all(np.isfinite(dummy_logits)), "inf/nan encountered"
assert not np.allclose(dummy_logits.numpy().sum(-1), 1), "please predict linear outputs, don't use softmax (maybe you've just got unlucky)"
# +
# test for lookahead
dummy_input_ix_2 = tf.constant(to_matrix([line[:3] + 'e' * (len(line) - 3) for line in dummy_lines]))
dummy_logits_2 = model(dummy_input_ix_2)
assert np.allclose(dummy_logits[:, :3] - dummy_logits_2[:, :3], 0), "your model's predictions depend on FUTURE tokens. " \
" Make sure you don't allow any layers to look ahead of current token." \
" You can also get this error if your model is not deterministic (e.g. dropout). Disable it for this test."
# -
# We can now tune our network's parameters to minimize categorical crossentropy over training dataset $D$:
#
# $$ L = {\frac1{|D|}} \sum_{X \in D} \sum_{x_i \in X} - \log p(x_t \mid x_1, \dots, x_{t-1}, \theta) $$
#
# As usual with with neural nets, this optimization is performed via stochastic gradient descent with backprop. One can also note that minimizing crossentropy is equivalent to minimizing model __perplexity__, KL-divergence or maximizng log-likelihood.
# +
def compute_lengths(input_ix, eos_ix=token_to_id[EOS]):
""" compute length of each line in input ix (incl. first EOS), int32 vector of shape [batch_size] """
count_eos = tf.cumsum(tf.cast(tf.equal(input_ix, eos_ix), tf.int32), axis=1, exclusive=True)
lengths = tf.reduce_sum(tf.cast(tf.equal(count_eos, 0), tf.int32), axis=1)
return lengths
print('matrix:\n', dummy_input_ix.numpy())
print('lengths:', compute_lengths(dummy_input_ix).numpy())
# -
def compute_loss(model, input_ix):
"""
:param model: language model that can compute next token logits given token indices
:param input ix: int32 matrix of tokens, shape: [batch_size, length]; padded with eos_ix
"""
input_ix = tf.convert_to_tensor(input_ix, dtype=tf.int32)
logits = model(input_ix[:, :-1])
reference_answers = input_ix[:, 1:]
# Your task: implement loss function as per formula above
# your loss should only be computed on actual tokens, excluding padding
# predicting actual tokens and first EOS do count. Subsequent EOS-es don't
# you will likely need to use compute_lengths and/or tf.sequence_mask to get it right.
<YOUR CODE>
return <YOUR CODE: return scalar loss>
loss_1 = compute_loss(model, to_matrix(dummy_lines, max_len=15))
loss_2 = compute_loss(model, to_matrix(dummy_lines, max_len=16))
assert (np.ndim(loss_1) == 0) and (0 < loss_1 < 100), "loss must be a positive scalar"
assert np.allclose(loss_1, loss_2), 'do not include AFTER first EOS into loss. '\
'Hint: use tf.sequence_mask. Beware +/-1 errors. And be careful when averaging!'
# ### Evaluation
#
# You will need two functions: one to compute test loss and another to generate samples. For your convenience, we implemented them both in your stead.
# +
def score_lines(model, dev_lines, batch_size):
""" computes average loss over the entire dataset """
dev_loss_num, dev_loss_len = 0., 0.
for i in range(0, len(dev_lines), batch_size):
batch_ix = to_matrix(dev_lines[i: i + batch_size])
dev_loss_num += compute_loss(model, batch_ix) * len(batch_ix)
dev_loss_len += len(batch_ix)
return dev_loss_num / dev_loss_len
def generate(model, prefix=BOS, temperature=1.0, max_len=100):
"""
Samples output sequence from probability distribution obtained by model
:param temperature: samples proportionally to model probabilities ^ temperature
if temperature == 0, always takes most likely token. Break ties arbitrarily.
"""
while True:
token_probs = model.get_possible_next_tokens(prefix)
tokens, probs = zip(*token_probs.items())
if temperature == 0:
next_token = tokens[np.argmax(probs)]
else:
probs = np.array([p ** (1. / temperature) for p in probs])
probs /= sum(probs)
next_token = np.random.choice(tokens, p=probs)
prefix += next_token
if next_token == EOS or len(prefix) > max_len: break
return prefix
# -
# ### Training loop
#
# Finally, let's train our model on minibatches of data
# +
from sklearn.model_selection import train_test_split
train_lines, dev_lines = train_test_split(lines, test_size=0.25, random_state=42)
batch_size = 256
score_dev_every = 250
train_history, dev_history = [], []
optimizer = keras.optimizers.Adam()
# score untrained model
dev_history.append((0, score_lines(model, dev_lines, batch_size)))
print("Sample before training:", generate(model, 'Bridging'))
# +
from IPython.display import clear_output
from random import sample
from tqdm import trange
for i in trange(len(train_history), 5000):
batch = to_matrix(sample(train_lines, batch_size))
with tf.GradientTape() as tape:
loss_i = compute_loss(model, batch)
grads = tape.gradient(loss_i, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_history.append((i, loss_i.numpy()))
if (i + 1) % 50 == 0:
clear_output(True)
plt.scatter(*zip(*train_history), alpha=0.1, label='train_loss')
if len(dev_history):
plt.plot(*zip(*dev_history), color='red', label='dev_loss')
plt.legend(); plt.grid(); plt.show()
print("Generated examples (tau=0.5):")
for _ in range(3):
print(generate(model, temperature=0.5))
if (i + 1) % score_dev_every == 0:
print("Scoring dev...")
dev_history.append((i, score_lines(model, dev_lines, batch_size)))
print('#%i Dev loss: %.3f' % dev_history[-1])
# +
assert np.mean(train_history[:10], axis=0)[1] > np.mean(train_history[-10:], axis=0)[1], "The model didn't converge."
print("Final dev loss:", dev_history[-1][-1])
for i in range(10):
print(generate(model, temperature=0.5))
# -
# ### RNN Language Models (3 points including training)
#
# Fixed-size architectures are reasonably good when capturing short-term dependencies, but their design prevents them from capturing any signal outside their window. We can mitigate this problem by using a __recurrent neural network__:
#
# $$ h_0 = \vec 0 ; \quad h_{t+1} = RNN(x_t, h_t) $$
#
# $$ p(x_t \mid x_0, \dots, x_{t-1}, \theta) = dense_{softmax}(h_{t-1}) $$
#
# Such model processes one token at a time, left to right, and maintains a hidden state vector between them. Theoretically, it can learn arbitrarily long temporal dependencies given large enough hidden size.
#
# <img src='https://raw.githubusercontent.com/yandexdataschool/nlp_course/master/resources/rnn_lm.jpg' width=480px>
class RNNLanguageModel(L.Layer):
def __init__(self, n_tokens=n_tokens, emb_size=16, hid_size=256):
"""
Build a recurrent language model.
You are free to choose anything you want, but the recommended architecture is
- token embeddings
- one or more LSTM/GRU layers with hid size
- linear layer to predict logits
"""
super().__init__() # initialize base class to track sub-layers, trainable variables, etc.
# YOUR CODE - create layers/variables/etc
<...>
#END OF YOUR CODE
def __call__(self, input_ix):
"""
compute language model logits given input tokens
:param input_ix: batch of sequences with token indices, tf tensor: int32[batch_size, sequence_length]
:returns: pre-softmax linear outputs of language model [batch_size, sequence_length, n_tokens]
these outputs will be used as logits to compute P(x_t | x_0, ..., x_{t - 1})
"""
#YOUR CODE
return <...>
def get_possible_next_tokens(self, prefix=BOS, temperature=1.0, max_len=100):
""" :returns: probabilities of next token, dict {token : prob} for all tokens """
prefix_ix = tf.convert_to_tensor(to_matrix([prefix]), tf.int32)
probs = tf.nn.softmax(self(prefix_ix)[0, -1]).numpy() # shape: [n_tokens]
return dict(zip(tokens, probs))
# +
model = RNNLanguageModel()
# note: tensorflow and keras layers create variables only after they're first applied (called)
dummy_input_ix = tf.constant(to_matrix(dummy_lines))
dummy_logits = model(dummy_input_ix)
assert isinstance(dummy_logits, tf.Tensor)
assert dummy_logits.shape == (len(dummy_lines), max(map(len, dummy_lines)), n_tokens), "please check output shape"
assert np.all(np.isfinite(dummy_logits)), "inf/nan encountered"
assert not np.allclose(dummy_logits.numpy().sum(-1), 1), "please predict linear outputs, don't use softmax (maybe you've just got unlucky)"
print('Weights:', tuple(w.name for w in model.trainable_variables))
# +
# test for lookahead
dummy_input_ix_2 = tf.constant(to_matrix([line[:3] + 'e' * (len(line) - 3) for line in dummy_lines]))
dummy_logits_2 = model(dummy_input_ix_2)
assert np.allclose(dummy_logits[:, :3] - dummy_logits_2[:, :3], 0), "your model's predictions depend on FUTURE tokens. " \
" Make sure you don't allow any layers to look ahead of current token." \
" You can also get this error if your model is not deterministic (e.g. dropout). Disable it for this test."
# -
# ### RNN training
#
# Our RNN language model should optimize the same loss function as fixed-window model. But there's a catch. Since RNN recurrently multiplies gradients through many time-steps, gradient values may explode, [ruining](https://raw.githubusercontent.com/yandexdataschool/nlp_course/master/resources/nan.jpg) your model.
# The common solution to that problem is to clip gradients either [individually](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/clip_by_value) or [globally](https://www.tensorflow.org/versions/r2.0/api_docs/python/tf/clip_by_global_norm).
#
# Your task here is to prepare tensorflow graph that would minimize the same loss function. If you encounter large loss fluctuations during training, please add gradient clipping using urls above.
#
# _Note: gradient clipping is not exclusive to RNNs. Convolutional networks with enough depth often suffer from the same issue._
# +
batch_size = 64 # <-- please tune batch size to fit your CPU/GPU configuration
score_dev_every = 250
train_history, dev_history = [], []
optimizer = keras.optimizers.Adam()
# score untrained model
dev_history.append((0, score_lines(model, dev_lines, batch_size)))
print("Sample before training:", generate(model, 'Bridging'))
# -
for i in trange(len(train_history), 5000):
batch = to_matrix(sample(train_lines, batch_size))
with tf.GradientTape() as tape:
loss_i = compute_loss(model, batch)
grads = tape.gradient(loss_i, model.trainable_variables)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
train_history.append((i, loss_i.numpy()))
if (i + 1) % 50 == 0:
clear_output(True)
plt.scatter(*zip(*train_history), alpha=0.1, label='train_loss')
if len(dev_history):
plt.plot(*zip(*dev_history), color='red', label='dev_loss')
plt.legend(); plt.grid(); plt.show()
print("Generated examples (tau=0.5):")
for _ in range(3):
print(generate(model, temperature=0.5))
if (i + 1) % score_dev_every == 0:
print("Scoring dev...")
dev_history.append((i, score_lines(model, dev_lines, batch_size)))
print('#%i Dev loss: %.3f' % dev_history[-1])
assert np.mean(train_history[:10], axis=0)[1] > np.mean(train_history[-10:], axis=0)[1], "The model didn't converge."
print("Final dev loss:", dev_history[-1][-1])
for i in range(10):
print(generate(model, temperature=0.5))
# ### Alternative sampling strategies (1 point)
#
# So far we've sampled tokens from the model in proportion with their probability.
# However, this approach can sometimes generate nonsense words due to the fact that softmax probabilities of these words are never exactly zero. This issue can be somewhat mitigated with sampling temperature, but low temperature harms sampling diversity. Can we remove the nonsense words without sacrificing diversity? __Yes, we can!__ But it takes a different sampling strategy.
#
# __Top-k sampling:__ on each step, sample the next token from __k most likely__ candidates from the language model.
#
# Suppose $k=3$ and the token probabilities are $p=[0.1, 0.35, 0.05, 0.2, 0.3]$. You first need to select $k$ most likely words and set the probability of the rest to zero: $\hat p=[0.0, 0.35, 0.0, 0.2, 0.3]$ and re-normalize:
# $p^*\approx[0.0, 0.412, 0.0, 0.235, 0.353]$.
#
# __Nucleus sampling:__ similar to top-k sampling, but this time we select $k$ dynamically. In nucleous sampling, we sample from top-__N%__ fraction of the probability mass.
#
# Using the same $p=[0.1, 0.35, 0.05, 0.2, 0.3]$ and nucleous N=0.9, the nucleous words consist of:
# 1. most likely token $w_2$, because $p(w_2) < N$
# 2. second most likely token $w_5$, $p(w_2) + p(w_5) = 0.65 < N$
# 3. third most likely token $w_4$ because $p(w_2) + p(w_5) + p(w_4) = 0.85 < N$
#
# And thats it, because the next most likely word would overflow: $p(w_2) + p(w_5) + p(w_4) + p(w_1) = 0.95 > N$.
#
# After you've selected the nucleous words, you need to re-normalize them as in top-k sampling and generate the next token.
#
# __Your task__ is to implement nucleus sampling variant and see if its any good.
def generate_nucleus(model, prefix=BOS, nucleus=0.9, max_len=100):
"""
Generate a sequence with nucleous sampling
:param prefix: a string containing space-separated previous tokens
:param nucleus: N from the formulae above, N \in [0, 1]
:param max_len: generate sequences with at most this many tokens, including prefix
:note: make sure that nucleous always contains at least one word, even if p(w*) > nucleus
"""
while True:
token_probs = model.get_possible_next_tokens(prefix)
tokens, probs = zip(*token_probs.items())
<YOUR CODE HERE>
prefix += <YOUR CODE>
if next_token == EOS or len(prefix) > max_len: break
return prefix
for i in range(10):
print(generate_nucleous(model, nucleous_size=PLAY_WITH_ME_SENPAI))
# ### Bonus quest I: Beam Search (2 pts incl. samples)
#
# At times, you don't really want the model to generate diverse outputs as much as you want a __single most likely hypothesis.__ A single best translation, most likely continuation of the search query given prefix, etc. Except, you can't get it.
#
# In order to find the exact most likely sequence containing 10 tokens, you would need to enumerate all $|V|^{10}$ possible hypotheses. In practice, 9 times out of 10 you will instead find an approximate most likely output using __beam search__.
#
# Here's how it works:
# 0. Initial `beam` = [prefix], max beam_size = k
# 1. for T steps:
# 2. ` ... ` generate all possible next tokens for all hypotheses in beam, formulate `len(beam) * len(vocab)` candidates
# 3. ` ... ` select beam_size best for all candidates as new `beam`
# 4. Select best hypothesis (-es?) from beam
from IPython.display import HTML
# Here's what it looks like:
# !wget -q https://raw.githubusercontent.com/yandexdataschool/nlp_course/2020/resources/beam_search.html
HTML("beam_search.html")
def generate_beamsearch(model, prefix=BOS, beam_size=4, length=5):
"""
Generate a sequence with nucleous sampling
:param prefix: a string containing space-separated previous tokens
:param nucleus: N from the formulae above, N \in [0, 1]
:param length: generate sequences with at most this many tokens, NOT INCLUDING PREFIX
:returns: beam_size most likely candidates
:note: make sure that nucleous always contains at least one word, even if p(w*) > nucleus
"""
<YOUR CODE HERE>
return <most likely sequence>
generate_beamsearch(model, prefix=' deep ', beam_size=4)
# +
# check it out: which beam size works best?
# find at least 5 prefixes where beam_size=1 and 8 generates different sequences
# -
# ### Bonus quest II: Ultimate Language Model (2+ pts)
#
# So you've learned the building blocks of neural language models, you can now build the ultimate monster:
# * Make it char-level, word level or maybe use sub-word units like [bpe](https://github.com/rsennrich/subword-nmt);
# * Combine convolutions, recurrent cells, pre-trained embeddings and all the black magic deep learning has to offer;
# * Use strides to get larger window size quickly. Here's a [scheme](https://storage.googleapis.com/deepmind-live-cms/documents/BlogPost-Fig2-Anim-160908-r01.gif) from google wavenet.
# * Train on large data. Like... really large. Try [1 Billion Words](http://www.statmt.org/lm-benchmark/1-billion-word-language-modeling-benchmark-r13output.tar.gz) benchmark;
# * Use training schedules to speed up training. Start with small length and increase over time; Take a look at [one cycle](https://medium.com/@nachiket.tanksale/finding-good-learning-rate-and-the-one-cycle-policy-7159fe1db5d6) for learning rate;
#
# _You are NOT required to submit this assignment. Please make sure you don't miss your deadline because of it :)_
|
week03_lm/homework_tf.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AXON by examples
# +
import axon
line = '\n'+32*'-'
# -
# ### Sequence of key:value pairs
vals = axon.loads("""
RU: "Moscow"
US: "Washington"
GB: "London"
""")
print(vals, line)
print(axon.dumps(vals))
# ### Sequence of ordered dicts
vals = axon.loads("""
[RU:"Moscow" FR:"Paris"]
[CN:"Beijin" JP:"Tokio"]
[US:"Washington" CA:"Ottava"]
""")
print(vals, line)
print(axon.dumps(vals), line)
print(axon.dumps(vals, pretty=1))
# ### Sequence of dicts
vals = axon.loads("""
{RU:"Moscow" FR:"Paris"}
{CN:"Beigin" JP:"Tokio"}
{US:"Washington" CA:"Ottava"}
""")
print(vals, line)
print(axon.dumps(vals, sorted=1), line)
print(axon.dumps(vals, sorted=1, pretty=1))
# ### Sequence of tuples
vals = axon.loads("""
("RU" "Moscow")
("US" "Washington")
("GB" "London")
""")
print(vals, line)
print(axon.dumps(vals))
# ### Sequence of lists
vals = axon.loads("""
["RU" "Moscow"]
["US" "Washington"]
["GB" "London"]
""")
print(vals, line)
print(axon.dumps(vals))
# ### List of atomic values
vals = axon.loads("""
[1 3.14 1000D "abc абв" 2015-12-01 12:00-03:00 2015-12-01T12:00+03:00 ? ∞]
""")
print(vals, line)
print(axon.dumps(vals))
# ### List of nodes
vals = axon.loads("""
country{id:"RU" capital:"Moscow"}
country{id:"USA" capital:"Washington"}
country{id:"GB" capital:"London"}
""")
print(vals, line)
print(axon.dumps(vals), line)
print(axon.dumps(vals, pretty=1), line)
print(axon.dumps(vals, pretty=1, braces=1))
# ### Sequence of objects
# +
class Country:
def __init__(self, id, capital):
self.id = id
self.capital = capital
def __repr__(self):
return "Country(id='{}', capital='{}')".format(self.id, self.capital)
@axon.reduce(Country)
def reduce_country(ob):
return axon.node("country", axon.odict([("id",ob.id), ("capital",ob.capital)]))
@axon.factory("country")
def factory_country(map, seq):
return Country(map["id"], map["capital"])
vals = axon.loads("""
country
id:"RU"
capital:"Moscow"
country
id:"USA"
capital:"Washington"
country
id:"GB"
capital:"London"
""", mode='strict')
print(vals, line)
print(axon.dumps(vals), line)
print(axon.dumps(vals, pretty=1, braces=1))
# -
|
examples/axon_by_examples2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from nltk.stem import PorterStemmer
from nltk.tokenize import word_tokenize
from nltk import pos_tag
from textblob import TextBlob
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import subprocess
import tensorflow as tf
stopwords=['i', 'me', 'my', 'myself', 'we', 'our', 'ours', 'ourselves', 'you', "you're", "you've", "you'll", "you'd", 'your', 'yours', 'yourself', 'yourselves', 'he', 'him', 'his', 'himself', 'she', "she's", 'her', 'hers', 'herself', 'it', "it's", 'its', 'itself', 'they', 'them', 'their', 'theirs', 'themselves', 'what', 'which', 'who', 'whom', 'this', 'that', "that'll", 'these', 'those', 'am', 'is', 'are', 'was', 'were', 'be', 'been', 'being', 'have', 'has', 'had', 'having', 'do', 'does', 'did', 'doing', 'a', 'an', 'the', 'and', 'but', 'if', 'or', 'because', 'as', 'until', 'while', 'of', 'at', 'by', 'for', 'with', 'about', 'against', 'between', 'into', 'through', 'during', 'before', 'after', 'above', 'below', 'to', 'from', 'up', 'down', 'in', 'out', 'on', 'off', 'over', 'under', 'again', 'further', 'then', 'once', 'here', 'there', 'when', 'where', 'why', 'how', 'all', 'any', 'both', 'each', 'few', 'more', 'most', 'other', 'some', 'such', 'no', 'nor', 'only', 'own', 'same', 'so', 'than', 'too', 'very', 's', 't', 'can', 'will', 'just', 'don', "don't", 'should', "should've", 'now', 'd', 'll', 'm', 'o', 're', 've', 'y', 'ain', 'aren', "aren't", 'couldn', "couldn't", 'didn', "didn't", 'doesn', "doesn't", 'hadn', "hadn't", 'hasn', "hasn't", 'haven', "haven't", 'isn', "isn't", 'ma', 'mightn', "mightn't", 'mustn', "mustn't", 'needn', "needn't", 'shan', "shan't", 'shouldn', "shouldn't", 'wasn', "wasn't", 'weren', "weren't", 'won', "won't", 'wouldn', "wouldn't"]
punkts='''"#$%&\'()*+,-./:;<=>@[\\]^_`{|}~'''
def CorFilt(i):
ps = PorterStemmer()
buff=word_tokenize(i.lower().replace("\n","").replace(" "," ").replace("n't"," not"))
buff2=""
for j in pos_tag(buff):
if j[-1]=='RB' and j[0]!="not":
pass
else:
buff2+=j[0]+" "
buff2=buff2.replace("not ","NOT")
buff=word_tokenize(buff2.strip())
ans=""
for j in buff:
if (j not in punkts) and (j not in stopwords):
if j=="!":
ans+=" XXEXCLMARK"
elif j=="?":
ans+=" XXQUESMARK"
else:
if j!="'s" and j!="``":
ans+=" "+ps.stem(j)
return ans.strip()
# +
import pickle
f=open("EmoVec","rb")
EmoVec=pickle.load(f)
f.close()
f=open("vectorizer","rb")
vectorizer=pickle.load(f)
f.close()
model=tf.keras.models.load_model("models/")
# -
def EmowavE(sent,vectorizer=vectorizer,EmoVec=EmoVec,trans=True):
transDict={'gu':'Gujarati',
'hi':'Hindi'}
# Translate from any language to english
if trans:
analysis = TextBlob(sent)
if analysis.detect_language()!='en':
try:
print(f"\nInput text was in {transDict[analysis.detect_language()]}")
except:
print(f"\nInput text was not in English")
print("\nTranslating...")
output=subprocess.check_output(['trans','-b',sent])
sent=output.decode('utf-8').strip()
print(f"\nTranslation in English: {sent}")
EmoBuff=vectorizer.transform([CorFilt(sent)])
EmoDict={0:'anger',
1:'disgust',
2:'fear',
3:'joy',
4:'sadness'}
weights = [float(cosine_similarity(EmoBuff.reshape(-1,1).T,EmoVec[i].reshape(-1,1).T)) for i in range(EmoVec.shape[0])]
if sum(weights)==0:
weights = [0 for i in range(5)]
else:
weights = [i/sum(weights) for i in weights]
return EmoDict[np.argmax(weights)], weights
def EmopreD(sent,model=model,vectorizer=vectorizer):
EmoDict={0:'anger',
1:'disgust',
2:'fear',
3:'joy',
4:'sadness'}
buff=vectorizer.transform([CorFilt(sent)]).toarray()
weights = model.predict(buff.reshape(1,1,buff.shape[1]))
return EmoDict[np.argmax(weights)], weights
sentence = "a perfectly parceled dead body"
print(f"\n\t>>> Emotion from VSM: {EmowavE(sentence)}")
print(f"\n\t>>> Emotion from LSTM: {EmopreD(sentence)}")
sentence = "i saw ghost"
print(f"\n\t>>> Emotion from VSM: {EmowavE(sentence)}")
print(f"\n\t>>> Emotion from LSTM: {EmopreD(sentence)}")
np.argmax(EmopreD(sentence)[1]) == np.argmax(EmowavE(sentence)[1])
def EnsemblE(sent):
EmoV, weightV = EmowavE(sentence)
EmoL, weightL = EmopreD(sentence)
if np.argmax(weightV)==np.argmax(weightL):
sureFLAG=True
else:
sureFLAG=False
if np.max(weightV)>=np.max(weightL):
method = "VSM"
Emo = EmoV
print(f"\n\t>>> Emotion from {method}: {EmoV}")
else:
method = "LSTM"
Emo = EmoL
print(f"\n\t>>> Emotion from {method}: {EmoL}")
if not sureFLAG:
print("EmowavE is not sure this time though!")
return sureFLAG, method, Emo
sentence = "i saw a friendly ghost"
EnsemblE(sentence)
|
Realtime Emotion Mining (ENSEMBLE).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import os
import torch
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import warnings
warnings.filterwarnings("ignore")
plt.ion()
# +
landmarks_frame = pd.read_csv('faces/face_landmarks.csv')
n = 65
img_name = landmarks_frame.iloc[n, 0]
landmarks = landmarks_frame.iloc[n, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
print('Image name: {}'.format(img_name))
print('Landmarks shape: {}'.format(landmarks.shape))
print('First 4 Landmarks: {}'.format(landmarks[:4]))
# +
def show_landmarks(image, landmarks):
plt.imshow(image)
plt.scatter(landmarks[:, 0], landmarks[:, 1], s=10, marker='.', c='r')
plt.pause(0.001)
plt.figure()
show_landmarks(io.imread(os.path.join('faces/', img_name)), landmarks)
plt.show()
# -
class FaceLandmarksDataset(Dataset):
def __init__(self, csv_file, root_dir, transform=None):
self.landmarks_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.landmarks_frame)
def __getitem__(self, idx):
img_name = os.path.join(self.root_dir, self.landmarks_frame.iloc[idx, 0])
image = io.imread(img_name)
landmarks = self.landmarks_frame.iloc[idx, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
sample = {'image': image, 'landmarks': landmarks}
if self.transform:
sample = self.transform(sample)
return sample
# +
face_dataset = FaceLandmarksDataset(csv_file='faces/face_landmarks.csv', root_dir='faces/')
fig = plt.figure()
for i in range(len(face_dataset)):
sample = face_dataset[i]
print(i, sample['image'].shape, sample['landmarks'].shape)
ax = plt.subplot(1, 4, i+1)
plt.tight_layout()
ax.set_title('sample #{}'.format(i))
ax.axis('off')
show_landmarks(**sample)
if i == 3:
plt.show()
break
# -
# # Transforms
# +
class Rescale(object):
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = transform.resize(image, (new_h, new_w))
landmarks = landmarks * [new_w / w, new_h / h]
return {'image': img, 'landmarks' : landmarks}
class RandomCrop(object):
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
landmarks = landmarks - [left, top]
return {'image': image, 'landmarks': landmarks}
class ToTensor(object):
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image), 'landmarks': torch.from_numpy(landmarks)}
# -
# # Compose transforms
# +
scale = Rescale(256)
crop = RandomCrop(128)
composed = transforms.Compose([Rescale(256), RandomCrop(224)])
fig = plt.figure()
sample = face_dataset[65]
for i, tsfrm in enumerate([scale, crop, composed]):
transformed_sample = tsfrm(sample)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tsfrm).__name__)
show_landmarks(**transformed_sample)
plt.show()
# -
# # Iterating through the dataset
transformed_dataset = FaceLandmarksDataset(csv_file='./faces/face_landmarks.csv', root_dir=('faces/'),
transform=transforms.Compose([Rescale(256),
RandomCrop(224),
ToTensor()
]))
for i in range(len(transformed_dataset)):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['landmarks'].size())
if i == 3:
break
# +
dataloader = DataLoader(transformed_dataset, batch_size=4, shuffle=True, num_workers=4)
def show_landmarks_batch(sample_batched):
images_batch, landmaks_batch = sample_batched['image'], sample_batched['landmarks']
batch_size = len(images_batch)
im_size = images_batch.size(2)
grid = utils.make_grid(images_batch)
plt.imshow(grid.numpy().transpose((1,2,0)))
for i in range(batch_size):
plt.scatter(landmaks_batch[i, :, 0].numpy() + i * im_size,
landmaks_batch[i, :, 1].numpy(),
s=10, marker='.', c='r')
plt.title('Batch from dataloader')
for i_batch, sample_batched in enumerate(dataloader):
print(i_batch, sample_batched['image'].size(), sample_batched['landmarks'].size())
if i_batch == 3:
plt.figure()
show_landmarks_batch(sample_batched)
plt.axis('off')
plt.ioff()
plt.show()
break
# -
# # Afterword: Torchvision
# +
import torch
from torchvision import transforms, datasets
data_transform = transforms.Compose([transforms.RandomResizedCrop(224),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
])
hymenoptera_dataset = datasets.ImageFolder(root='hymenoptera_data/train', transform=data_transform)
dataset_loader = torch.utils.data.DataLoader(hymenoptera_dataset, batch_size=4, shuffle=True, num_workers=4)
# -
|
tutorial/getting_started/data_loading_and_processing_tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SummerLife/EmbeddedSystem/blob/master/MachineLearning/gist/mobilenet_image_classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="-75DJsOSnkUM" colab_type="code" colab={}
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.layers import Dense, Activation
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.metrics import categorical_crossentropy
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.preprocessing import image
from tensorflow.keras.models import Model
from tensorflow.keras.applications import imagenet_utils
from sklearn.metrics import confusion_matrix
import itertools
import os
import shutil
import random
import matplotlib.pyplot as plt
# %matplotlib inline
# + id="u6X_vq8Xny9K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2085369a-1685-4692-f180-03ba20a5f9a3"
physical_devices = tf.config.experimental.list_physical_devices('GPU')
print("Num GPUs Available: ", len(physical_devices))
tf.config.experimental.set_memory_growth(physical_devices[0], True)
# + id="lL8hJKXUoAzE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="e5daa364-75d0-4ac0-8efc-3d20e2a52a88"
mobile = tf.keras.applications.mobilenet.MobileNet()
# + id="WQnBX_OioJl8" colab_type="code" colab={}
def prepare_image(file):
img_path = ''
img = image.load_img(img_path + file, target_size=(224, 224))
img_array = image.img_to_array(img)
img_array_expanded_dims = np.expand_dims(img_array, axis=0)
return tf.keras.applications.mobilenet.preprocess_input(img_array_expanded_dims)
# + id="2kT0Nwoto_Oq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="1dba1367-9fd9-465a-e259-410a02d9c6b5"
from IPython.display import Image
Image(filename='/content/drive/My Drive/train_data/test/dog/dog.10667.jpg', width=300,height=200)
# + id="aVf-sxHvpms0" colab_type="code" colab={}
preprocessed_image = prepare_image('/content/drive/My Drive/train_data/test/dog/dog.10667.jpg')
predictions = mobile.predict(preprocessed_image)
# + id="x14lvXMJpzUk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="e1fe685a-9b20-4f49-991d-0edbb783cbfa"
results = imagenet_utils.decode_predictions(predictions)
print(results)
# + id="Q9ic21deqDjA" colab_type="code" colab={}
|
MachineLearning/gist/mobilenet_image_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Calculate the mean value of the USD-EUR
import pandas as pd
import numpy as np
import json
import datetime
import pickle
import functions as fn
import matplotlib.pyplot as plt
# %matplotlib inline
import requests
# ## Select wanted exchange rate
exchange_currency = input()
desired_currency = input()
# ## Import data to calculate the exchange rate
#Retrieving my api keys information to access the Alpha Advantage API.
keys = fn.get_keys("/Users/jjherranzsarrion/.secret/alphaadvantage.json")
api_key = keys['api_key']
# Get the fx rate from Alpha Advantage
response_json = fn.get_fx_rates(api_key, exchange_currency, desired_currency)
# ## Graphing the exchange rate
response_df = fn.get_adjusted_rate(response_json)
response_df[-1:]['Adj Close Price'][0]
# Calculate 30 Day Moving Average, Std Deviation, Upper Band and Lower Band
response_bb_df = fn.get_bollinger_bands(response_df)
# Retrieve todays date
today = datetime.date.today()
today.replace(day=1)
datetime.timedelta(days=1)
print((today.replace(day=1) - datetime.timedelta(days=1)).replace(day=today.day).strftime("%Y-%m-%d"))
def get_graphical_view(response_df, exchange_currency, desired_currency, today):
"""Function that returns a graphic view of the exchange rate in question
and the corresponding bollinger bands."""
# We only want to show the previous month, therefore subset the dataframe
one_month_ago = (today.replace(day=1) - datetime.timedelta(days=1)).replace(day=today.day).strftime("%Y-%m-%d")
print(one_month_ago)
date_15_days_ago = (today - datetime.timedelta(days=15)).strftime("%Y-%m-%d")
response_df = response_df.loc[(response_df.index >= one_month_ago) & (response_df.index <= today.strftime("%Y-%m-%d"))]
close = response_bb_df[-1:]['Adj Close Price']
# set style, empty figure and axes
fig = plt.figure(figsize=(10,5), facecolor='w')
ax = fig.add_subplot(111)
# Get index values for the X axis for exchange rate DataFrame
x_axis = response_df.index
# Plot shaded 21 Day Bollinger Band for exchange rate
#ax.fill_between(x_axis, response_df['Upper Band'], response_df['Lower Band'], color='white')
# Plot Adjust Closing Price and Moving Averages
ax.plot(x_axis, response_df['Adj Close Price'], color='blue', lw=2)
#ax.plot(x_axis, response_df['30 Day MA'], color='black', lw=2)
ax.plot(x_axis, response_df['Upper Band'], color='green', lw=2, )
ax.plot(x_axis, response_df['Lower Band'], color='red', lw=2)
ax.set_xticks([one_month_ago, date_15_days_ago, today.strftime("%Y-%m-%d")])
ax.yaxis.tick_right()
ax.set_facecolor('#ffffff')
ax.spines['left'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# Set Title & Show the Image
# Compare the value of the exchange rate currencies
compare = response_df.loc[response_df.index == today.strftime("%Y-%m-%d")]
if compare['Adj Close Price'].values > compare['Upper Band'].values:
print(f'The {exchange_currency} is strong, consider making your international transaction today.')
elif compare['Adj Close Price'].values > compare['Lower Band'].values:
print(f"The {exchange_currency} is currently trading according to its boundaries.")
else:
print(f"The {exchange_currency} is weak, consider making your international transaction another day.")
return response_df
# Return graphic showing the previous monthly history of the exchange rate in question and return recommendation
response_graphic = get_graphical_view(response_bb_df, exchange_currency, desired_currency, today)
# There is a problem with the dates on the x axis so check that out
response_bb_df[-1:]['Adj Close Price']
response_bb_df
response_df[-1:]['30 Day MA'][0]
|
currency_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], True)
# +
import numpy as np
import pickle
import operator
import cv2
from os import listdir
from sklearn.preprocessing import LabelBinarizer
from keras.models import Sequential
from keras.layers.normalization import BatchNormalization
from keras.layers.convolutional import Conv2D
from keras.layers.convolutional import MaxPooling2D
from keras.layers.core import Activation, Flatten, Dropout, Dense
from keras import backend as K
from keras.preprocessing.image import ImageDataGenerator
#from keras.optimizers import Adam
from keras.preprocessing import image
from keras.preprocessing.image import img_to_array
from sklearn.preprocessing import MultiLabelBinarizer
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import tensorflow as tf
from keras.models import Sequential
from keras.layers import Convolution2D, MaxPooling2D, Flatten, Dense, Dropout, GlobalAveragePooling2D
import tensorflow as tf
import keras
import os
import time
#import cv2
import numpy as np
#import matplotlib.pyplot as plt
from keras import optimizers
import keras
import tensorflow as tf
import keras.backend as K
from sklearn.metrics import confusion_matrix, classification_report
from keras.models import load_model
from keras.models import Sequential
from keras.regularizers import l2
from keras.applications.vgg16 import VGG16
from keras.preprocessing.image import ImageDataGenerator
from keras.callbacks import ModelCheckpoint, CSVLogger, EarlyStopping, ReduceLROnPlateau
from sklearn.metrics import accuracy_score
from sklearn.metrics import confusion_matrix, classification_report
from sklearn.metrics import roc_curve, auc, roc_auc_score
#import matplotlib.pyplot as plt
#from tqdm import tqdm
from keras.utils import np_utils
#from imgaug import augmenters as iaa
import itertools
# %matplotlib inline
#import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
#from keras.optimizers import RMSprop
from keras.layers import Conv2D, MaxPooling2D
from keras.layers import Activation, Dropout, Flatten, Dense
from keras.callbacks import CSVLogger
#from livelossplot import PlotLossesKeras
import os
#from classification_models.keras import Classifiers
# GPU test
from tensorflow.python.client import device_lib
np.random.seed(42)
from keras.models import load_model
# Print version
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from keras.layers import Input, Activation, merge, Dense, Flatten, Dropout, BatchNormalization, GlobalAveragePooling2D
from keras.models import Model
from keras.layers import Input, Activation, merge, Dense, Flatten, Dropout, BatchNormalization, GlobalAveragePooling2D
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.applications.vgg16 import VGG16
from keras.regularizers import l2
import os, sys
import scipy.misc
from glob import glob
import numpy as np
import random
import shutil
import keras
import time
import matplotlib.pyplot as plt
from keras.layers import Dense, Input, Conv2D, Flatten, MaxPool2D, Activation,Dropout, GlobalAveragePooling2D
from keras.models import Model
from keras.callbacks import ModelCheckpoint
from keras import backend as K
from keras.models import Sequential
from keras.applications.densenet import DenseNet121
from keras.models import load_model
#from keras.optimizers import Adam
from keras import optimizers
import pickle
import sys
import cv2
import tensorflow as tf
np.random.seed(1)
# +
DIRECTORY = r'D:\Leuk Dataset\new fol\train'
VAL_DIRECTORY = r'D:\Leuk Dataset\new fol\val'
CATEGORIES = ['all', 'hem']
# -
def crop_center(img, bounding):
start = tuple(map(lambda a, da: a//2-da//2, img.shape, bounding))
end = tuple(map(operator.add, start, bounding))
slices = tuple(map(slice, start, end))
return img[slices]
data = []
i = 0
plt.figure(figsize=(15, 15))
for category in CATEGORIES:
path = os.path.join(DIRECTORY, category)
for img in os.listdir(path):
img_path = os.path.join(path, img)
label = CATEGORIES.index(category)
arr = cv2.imread(img_path)
crop_arr = crop_center(arr, (210,210))
if 1 <= i+1 <= 140: # total 140 image
ax = plt.subplot(13, 11, i+1)
plt.imshow(crop_arr)
i += 1
data.append([crop_arr, label])
# +
random.shuffle(data)
x_train = []
y_train = []
for features, label in data:
x_train.append(features)
y_train.append(label)
x_train = np.array(x_train)
y_train = np.array(y_train)
print(x_train.shape)
np.save("x_train_leuknet",x_train)
np.save("y_train_leuknet",y_train)
# -
val_data = []
i = 0
plt.figure(figsize=(15, 15))
for category in CATEGORIES:
path = os.path.join(VAL_DIRECTORY, category)
for img in os.listdir(path):
img_path = os.path.join(path, img)
label = CATEGORIES.index(category)
arr = cv2.imread(img_path)
crop_arr = crop_center(arr, (210,210))
if 1 <= i+1 <= 70: # total image 70
ax = plt.subplot(10, 7, i+1)
plt.imshow(crop_arr)
i += 1
val_data.append([crop_arr, label])
# +
random.shuffle(val_data)
x_val = []
y_val = []
for features, label in val_data:
x_val.append(features)
y_val.append(label)
x_val = np.array(x_val)
y_val = np.array(y_val)
# +
from tensorflow.keras.applications.resnet50 import ResNet50
from tensorflow.keras.layers import Conv2D, BatchNormalization, Activation, MaxPool2D, Conv2DTranspose, Concatenate, Input
from tensorflow.keras.models import Model
from tensorflow.keras.applications import VGG16
import keras
from keras.models import Model, load_model
from keras.layers import Input
from keras.layers.core import Dropout, Lambda
from keras.layers.convolutional import Conv2D, Conv2DTranspose
from keras.layers.pooling import MaxPooling2D, AveragePooling2D
from keras.layers.merge import concatenate
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras import backend as K
import tensorflow as tf
from keras.optimizers import Adam, RMSprop, SGD
from keras.callbacks import ModelCheckpoint, CSVLogger, EarlyStopping, ReduceLROnPlateau
# load model without classifier layers
model = ResNet50(include_top=False, input_shape=(210, 210, 3))
# add new classifier layers
flat1 = Flatten()(model.layers[-1].output)
#class1 = Dense(1024, activation='relu')(flat1)
#class2 = Dense(1024, activation='relu')(class1)
output = Dense(1, activation='sigmoid')(flat1)
# define new model
model = Model(inputs=model.inputs, outputs=output)
# summarize
model.summary()
'''
def build_resnet50_unet(input_shape):
""" Input """
inputs = Input(input_shape)
s = Lambda(lambda x: x/255) (inputs)
""" Pre-trained VGG16 Model """
resnet50 = ResNet50(include_top=True, weights=None, input_tensor=s, input_shape=None,
pooling=None,
classes=1,
classifier_activation="sigmoid")
return resnet50
if __name__ == "__main__":
input_shape = (210, 210, 3)
model = build_resnet50_unet(input_shape)
model.summary()
'''
# +
#import tensorflow as tf
from keras.optimizers import Adam, RMSprop, SGD
adam_opt = Adam(lr=1e-5, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=1e-5)
#sgd_opt = SGD(learning_rate=1e-06, momentum=0.0, decay=0.0, nesterov=False)
#rmsp_opt = RMSprop(lr=1e-4, decay=0.9)
# eve_opt = Eve(lr=1e-4, decay=1E-4, beta_1=0.9, beta_2=0.999, beta_3=0.999, small_k=0.1, big_K=10, epsilon=1e-08)
model.compile(optimizer= adam_opt,
loss = 'binary_crossentropy',
metrics=['accuracy'])
# -
import tensorflow.keras as keras
callbacks = [
keras.callbacks.ModelCheckpoint('weighted_baseline_ResNet50.h5', monitor='val_acc', save_best_only=True, mode='max'),
keras.callbacks.ReduceLROnPlateau(monitor='val_acc', factor=0.1, verbose=1, patience=5, mode='max')]
print(x_train.shape)
print(x_val.shape)
'''
x_train = x_train.astype('float32')
x_val = x_val.astype('float32')
print(x_train.dtype)
print(x_val.dtype)
x_train = np.resize(x_train, (x_train.shape[0], 32, 32, 3))
x_val = np.resize(x_val, (x_val.shape[0], 32, 32, 3))
print(x_train.shape)
print(x_val.shape)
'''
# +
import math
BATCH_SIZE=32
TRAINING_SIZE = x_train.shape[0]
VALIDATION_SIZE = x_val.shape[0]
compute_steps_per_epoch = lambda x: int(math.ceil(1. * x / BATCH_SIZE))
train_steps_per_epoch = compute_steps_per_epoch(TRAINING_SIZE)
val_steps = compute_steps_per_epoch(VALIDATION_SIZE)
print(train_steps_per_epoch, val_steps)
# -
from keras.preprocessing.image import ImageDataGenerator
dataAugmentaion = ImageDataGenerator()
model.fit_generator(dataAugmentaion.flow(x_train, y_train, batch_size = 1),
validation_data = (x_val,y_val), steps_per_epoch = len(x_train) // 1,
epochs = 50, class_weight={0:0.73301705, 1:1.57288286})
|
others/Weighted_ResNet50_adam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PythonData
# language: python
# name: pythondata
# ---
# Add the dependencies
import pandas as pd
import os
# Files to load
school_data_to_load = os.path.join("Resources", "schools_complete.csv")
student_data_to_load = os.path.join("Resources", "students_complete.csv")
# Read the school data file and store it in a Pandas DataFrame
school_data_df = pd.read_csv(school_data_to_load)
student_data_df = pd.read_csv(student_data_to_load)
# Determine if there are any missing values in the school data.
school_data_df.count()
# Determine if there are an missing values in the student data.
student_data_df.count()
# Determine if there are any missing values in the school data.
school_data_df.isnull()
# Determine if there are any missing values in the student data.
student_data_df.isnull().sum()
# Determine if there are any missing values in the student data.
student_data_df.notnull().sum()
# Determine data types for the school DataFrame
school_data_df.dtypes
# Determine data types for the student DataFrame
student_data_df.dtypes
# Add each prefix and suffix to remove to a list.
prefixes_suffixes = ['Dr. ', 'Mr. ', 'Ms. ', 'Mrs. ', 'Miss ', ' MD', ' DDS', ' DVM', ' PhD']
# +
# Iterate through the words in the "prefixes_suffixes" list and replace them with an empty space, "".
for word in prefixes_suffixes:
student_data_df['student_name'] = student_data_df['student_name'].str.replace(word, '')
student_data_df.head(10)
# -
# Use cleaned student data from CSV
student_data_df = pd.read_csv("Resources/clean_students_complete.csv")
# Combine the data into a single dataset.
school_data_complete_df = pd.merge(student_data_df, school_data_df, on=['school_name', 'school_name'])
school_data_complete_df
# Get the total number of students.
student_count = school_data_complete_df['Student ID'].count()
student_count
# Calculate the total number of schools
school_count = len(school_data_complete_df["school_name"].unique())
print(school_count)
# Calculate the total budget
total_budget = school_data_df['budget'].sum()
total_budget
# Calculate the average reading score
average_reading_score = school_data_complete_df['reading_score'].mean()
average_reading_score
# Calculate the average math score
average_math_score = school_data_complete_df['math_score'].mean()
average_math_score
# Get all the students who are passing math in a new DataFrame
passing_math = school_data_complete_df[school_data_complete_df['math_score'] > 70]
passing_math.head()
# Get all the students that are passing reading in a new DataFrame.
passing_reading = school_data_complete_df[school_data_complete_df["reading_score"] > 70]
passing_math["student_name"].count()
# +
# Calculate the number of students passing math.
passing_math_count = passing_math["student_name"].count()
# Calculate the number of students passing reading.
passing_reading_count = passing_reading["student_name"].count()
# -
print(passing_math_count)
print(passing_reading_count)
# +
# Calculate the percent that passed math.
passing_math_percentage = passing_math_count / float(student_count) * 100
# Calculate the percent passing reading.
passing_reading_percentage = passing_reading_count / float(student_count) * 100
# -
print(passing_math_percentage)
print(passing_reading_percentage)
# Calculate the students who passed both math and reading.
passing_math_reading = school_data_complete_df[(school_data_complete_df['math_score'] >= 70) & (school_data_complete_df['reading_score'] >= 70)]
passing_math_reading.head()
# Calculate the number of students who passed both math and reading.
overall_passing_math_reading_count = passing_math_reading['student_name'].count()
overall_passing_math_reading_count
# Calculate the overall passing percentage
overall_passing_percentage = overall_passing_math_reading_count / student_count * 100
overall_passing_percentage
# Adding a list of values with keys to a create a new DataFrame
district_summary_df = pd.DataFrame(
[{'Total Schools': school_count,
'Total Students': student_count,
'Total Budget': total_budget,
'Average Math Score': average_math_score,
'Average Reading Score': average_reading_score,
'% Passing Math': passing_math_percentage,
'% Passing Reading': passing_reading_percentage,
'% Overall Passing': overall_passing_percentage}])
district_summary_df
# Format the Total Budget column to have a comma, a thousands separator, a decimal separator and a '$'
district_summary_df['Total Budget'] = district_summary_df['Total Budget'].map("${:,.2f}".format)
district_summary_df['Total Budget']
district_summary_df['Total Budget']
# Format the columns
district_summary_df['Average Math Score'] = district_summary_df['Average Math Score'].map("{:.1f}".format)
district_summary_df['Average Reading Score'] = district_summary_df['Average Reading Score'].map("{:.1f}".format)
district_summary_df['% Passing Math'] = district_summary_df['% Passing Math'].map("{:.1f}".format)
district_summary_df['% Passing Reading'] = district_summary_df['% Passing Reading'].map("{:.1f}".format)
district_summary_df['% Overall Passing'] = district_summary_df['% Overall Passing'].map("{:.1f}".format)
# Print the district summary DF
district_summary_df
# Reorder the columns in the order you want them to appear
new_column_order = ['Total Schools', 'Total Students', 'Total Budget', 'Average Math Score', 'Average Reading Score', '% Passing Math', '% Passing Reading', '% Overall Passing']
district_summary_df = district_summary_df[new_column_order]
district_summary_df
# Determine the school type.
per_school_types = school_data_df.set_index(['school_name'])['type']
per_school_types
# Add the per_school_types into a DataFrame for testing.
df = pd.DataFrame(per_school_types)
df
# Calculate the total student count.
per_school_counts = school_data_df.set_index(["school_name"])["size"]
per_school_counts
# Calculate the total student count.
per_school_counts = school_data_complete_df["school_name"].value_counts()
per_school_counts
# Calculate the total school budget.
per_school_budget = school_data_df.set_index(['school_name'])["budget"]
per_school_budget
# Calculate the per capita spending
per_school_capita = per_school_budget / per_school_counts
per_school_capita
# Calculate the math scores.
student_school_math = student_data_df.set_index(['school_name'])['math_score']
student_school_math
# +
# Calculate the average test scores
per_school_math = school_data_complete_df.groupby(['school_name']).mean()['math_score']
per_school_reading = school_data_complete_df.groupby(['school_name']).mean()['reading_score']
# -
per_school_math
per_school_reading
# +
# Calculate the passing scores by creating a filtered DataFrame
per_school_passing_math = school_data_complete_df[(school_data_complete_df['math_score'] >= 70)]
per_school_passing_reading = school_data_complete_df[(school_data_complete_df['reading_score'] >= 70)]
# -
per_school_passing_math
# +
# Calculate the number of students passing math and passing reading by school name
per_school_passing_math = per_school_passing_math.groupby(['school_name']).count()['student_name']
per_school_passing_reading = per_school_passing_reading.groupby(['school_name']).count()['student_name']
# -
per_school_passing_math
# +
# Calculate the percentage of passing math and reading score per school
per_school_passing_math = per_school_passing_math / per_school_counts * 100
per_school_passing_reading = per_school_passing_reading / per_school_counts * 100
# -
per_school_passing_math
# +
# Calculate the students who passed both math and reading.
per_passing_math_reading = school_data_complete_df[(school_data_complete_df['math_score'] >= 70) & (school_data_complete_df['reading_score'] >=70)]
per_passing_math_reading.head()
# -
# Calculate the number of students who passed both math and reading.
per_passing_math_reading = per_passing_math_reading.groupby(['school_name']).count()['student_name']
per_passing_math_reading
# Calculate the overall passing percentage.
per_overall_passing_percentage = per_passing_math_reading / per_school_counts * 100
per_overall_passing_percentage
# +
# Adding a list of values with keys to create a new DataFrame
per_school_summary_df = pd.DataFrame({
'School Type': per_school_types,
'Total Students': per_school_counts,
'Total School Budget': per_school_budget,
'Per Student Budget': per_school_capita,
'Average Math Score': per_school_math,
'Average Reading Score': per_school_reading,
'% Passing Math': per_school_passing_math,
'% Passing Reading': per_school_passing_reading,
'% Overall Passing': per_overall_passing_percentage
})
per_school_summary_df.head()
# +
# Format the Total School Budget and the Per Student Budget columns.
per_school_summary_df['Total School Budget'] = per_school_summary_df['Total School Budget'].map('${:,.2f}'.format)
per_school_summary_df['Per Student Budget'] = per_school_summary_df['Per Student Budget'].map('${:,.2f}'.format)
# Display the data frame
per_school_summary_df.head()
# +
# Sort and show the top five schools.
top_schools = per_school_summary_df.sort_values(['% Overall Passing'], ascending=False)
top_schools.head()
# +
# Sort and show bottom five schools.
bottom_schools = per_school_summary_df.sort_values(['% Overall Passing'], ascending=True)
bottom_schools.head()
# +
# Create grade level DataFrames.
ninth_graders = school_data_complete_df[(school_data_complete_df['grade'] == '9th')]
tenth_graders = school_data_complete_df[(school_data_complete_df['grade'] == '10th')]
eleventh_graders = school_data_complete_df[(school_data_complete_df['grade'] == '11th')]
twelfth_graders = school_data_complete_df[(school_data_complete_df['grade'] == '12th')]
# +
# Group each grade level DataFrame by the school name for the average math score.
ninth_grade_math_scores = ninth_graders.groupby(['school_name']).mean()['math_score']
tenth_grade_math_scores = tenth_graders.groupby(['school_name']).mean()['math_score']
eleventh_grade_math_scores = eleventh_graders.groupby(['school_name']).mean()['math_score']
twelfth_grade_math_scores = twelfth_graders.groupby(['school_name']).mean()['math_score']
# -
eleventh_grade_math_scores
# +
# Group each grade level DataFrame by the school name for the average math score.
ninth_grade_reading_scores = ninth_graders.groupby(['school_name']).mean()['reading_score']
tenth_grade_reading_scores = tenth_graders.groupby(['school_name']).mean()['reading_score']
eleventh_grade_reading_scores = eleventh_graders.groupby(['school_name']).mean()['reading_score']
twelfth_grade_reading_scores = twelfth_graders.groupby(['school_name']).mean()['reading_score']
# -
twelfth_grade_reading_scores
# +
# Combine each grade level Series for average math scores by school into a single DataFrame.
math_scores_by_grade = pd.DataFrame({
"9th": ninth_grade_math_scores,
"10th": tenth_grade_math_scores,
"11th": eleventh_grade_math_scores,
"12th": twelfth_grade_math_scores
})
math_scores_by_grade.head()
# +
# Combine each grade level Series for average reading scores by school into a single DataFrame.
reading_scores_by_grade = pd.DataFrame({
"9th": ninth_grade_reading_scores,
"10th": tenth_grade_reading_scores,
"11th": eleventh_grade_reading_scores,
"12th": twelfth_grade_reading_scores
})
reading_scores_by_grade.head()
# +
# Format each grade column.
math_scores_by_grade['9th'] = math_scores_by_grade['9th'].map('{:.1f}'.format)
math_scores_by_grade['10th'] = math_scores_by_grade['10th'].map('{:.1f}'.format)
math_scores_by_grade['11th'] = math_scores_by_grade['11th'].map('{:.1f}'.format)
math_scores_by_grade['12th'] = math_scores_by_grade['12th'].map('{:.1f}'.format)
# Make sure the columns are in the correct order.
math_scores_by_grade = math_scores_by_grade[
['9th', '10th', '11th', '12th']
]
# Remove the index name.
math_scores_by_grade.index.name = None
# Display the DataFrame
math_scores_by_grade
# +
# Format each grade column.
reading_scores_by_grade['9th'] = reading_scores_by_grade['9th'].map('{:.1f}'.format)
reading_scores_by_grade['10th'] = reading_scores_by_grade['10th'].map('{:.1f}'.format)
reading_scores_by_grade['11th'] = reading_scores_by_grade['11th'].map('{:.1f}'.format)
reading_scores_by_grade['12th'] = reading_scores_by_grade['12th'].map('{:.1f}'.format)
# Make sure the columns are in the correct order.
reading_scores_by_grade = reading_scores_by_grade[
['9th', '10th', '11th', '12th']
]
# Remove the index name.
reading_scores_by_grade.index.name = None
# Display the DataFrame
reading_scores_by_grade
# -
# Get the descriptive statistics for the per_school_capita
per_school_capita.describe()
# Cut the per_school_capita into spending ranges.
spending_bins = [0, 585, 615, 645, 675]
pd.cut(per_school_capita, spending_bins)
# Cut the per_school_capita into the spending ranges.
spending_bins = [0, 585, 630, 645, 675]
per_school_capita.groupby(pd.cut(per_school_capita, spending_bins)).count()
# Establish the spending bins and group names.
spending_bins = [0, 585, 630, 645, 675]
group_names = ["<$584", "$585-629", "$630-644", "$645-675"]
# +
# Categorize spending based on the bins.
per_school_summary_df['Spending Range (Per Student)'] = pd.cut(per_school_capita, spending_bins, labels=group_names)
per_school_summary_df
# +
# Calculate the averages for the desired columns.
spending_math_scores = per_school_summary_df.groupby(["Spending Range (Per Student)"]).mean()["Average Math Score"]
spending_reading_scores = per_school_summary_df.groupby(["Spending Range (Per Student)"]).mean()["Average Reading Score"]
spending_passing_math = per_school_summary_df.groupby(["Spending Range (Per Student)"]).mean()['% Passing Math']
spending_passing_reading = per_school_summary_df.groupby(["Spending Range (Per Student)"]).mean()['% Passing Reading']
overall_passing_spending = per_school_summary_df.groupby(["Spending Range (Per Student)"]).mean()['% Overall Passing']
# -
overall_passing_spending
# Assemble into DataFrame.
spending_summary_df = pd.DataFrame({
'Average Math Score': spending_math_scores,
'Average Reading Score': spending_reading_scores,
'% Passing Math': spending_passing_math,
'% Passing Reading': spending_passing_reading,
'% Overall Passing': overall_passing_spending
})
spending_summary_df
# +
# Formatting
spending_summary_df["Average Math Score"] = spending_summary_df["Average Math Score"].map("{:.1f}".format)
spending_summary_df["Average Reading Score"] = spending_summary_df["Average Reading Score"].map("{:.1f}".format)
spending_summary_df["% Passing Math"] = spending_summary_df["% Passing Math"].map("{:.0f}".format)
spending_summary_df["% Passing Reading"] = spending_summary_df["% Passing Reading"].map("{:.0f}".format)
spending_summary_df["% Overall Passing"] = spending_summary_df["% Overall Passing"].map("{:.0f}".format)
spending_summary_df
# -
# Establish the bins
size_bins = [0, 1000, 2000, 5000]
group_names = ["Small (<1000)", "Medium (1000-2000)", "Large (2000-5000)"]
# +
# Categorize spending based on the bins.
per_school_summary_df["School Size"] = pd.cut(per_school_summary_df["Total Students"], size_bins, labels=group_names)
per_school_summary_df.head()
# +
# Calculate averages for the desired columns.
size_math_scores = per_school_summary_df.groupby(["School Size"]).mean()["Average Math Score"]
size_reading_scores = per_school_summary_df.groupby(["School Size"]).mean()["Average Reading Score"]
size_passing_math = per_school_summary_df.groupby(["School Size"]).mean()["% Passing Math"]
size_passing_reading = per_school_summary_df.groupby(["School Size"]).mean()["% Passing Reading"]
size_overall_passing = per_school_summary_df.groupby(["School Size"]).mean()["% Overall Passing"]
# +
# Assemble into DataFrame.
size_summary_df = pd.DataFrame({
"Average Math Score" : size_math_scores,
"Average Reading Score": size_reading_scores,
"% Passing Math": size_passing_math,
"% Passing Reading": size_passing_reading,
"% Overall Passing": size_overall_passing})
size_summary_df
# +
# Formatting.
size_summary_df["Average Math Score"] = size_summary_df["Average Math Score"].map("{:.1f}".format)
size_summary_df["Average Reading Score"] = size_summary_df["Average Reading Score"].map("{:.1f}".format)
size_summary_df["% Passing Math"] = size_summary_df["% Passing Math"].map("{:.0f}".format)
size_summary_df["% Passing Reading"] = size_summary_df["% Passing Reading"].map("{:.0f}".format)
size_summary_df["% Overall Passing"] = size_summary_df["% Overall Passing"].map("{:.0f}".format)
size_summary_df
# +
# Calculate averages for the desired columns.
type_math_scores = per_school_summary_df.groupby(["School Type"]).mean()["Average Math Score"]
type_reading_scores = per_school_summary_df.groupby(["School Type"]).mean()["Average Reading Score"]
type_passing_math = per_school_summary_df.groupby(["School Type"]).mean()["% Passing Math"]
type_passing_reading = per_school_summary_df.groupby(["School Type"]).mean()["% Passing Reading"]
type_overall_passing = per_school_summary_df.groupby(["School Type"]).mean()["% Overall Passing"]
# +
# Assemble into DataFrame.
type_summary_df = pd.DataFrame({
"Average Math Score" : type_math_scores,
"Average Reading Score": type_reading_scores,
"% Passing Math": type_passing_math,
"% Passing Reading": type_passing_reading,
"% Overall Passing": type_overall_passing})
type_summary_df
# +
# Formatting
type_summary_df["Average Math Score"] = type_summary_df["Average Math Score"].map("{:.1f}".format)
type_summary_df["Average Reading Score"] = type_summary_df["Average Reading Score"].map("{:.1f}".format)
type_summary_df["% Passing Math"] = type_summary_df["% Passing Math"].map("{:.0f}".format)
type_summary_df["% Passing Reading"] = type_summary_df["% Passing Reading"].map("{:.0f}".format)
type_summary_df["% Overall Passing"] = type_summary_df["% Overall Passing"].map("{:.0f}".format)
type_summary_df
# -
|
PyCitySchools.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-3db1b7eb6aade580", "locked": true, "schema_version": 1, "solution": false}
# # BLU06 - Exercise Notebook
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-d9b262915f7cb1e5", "locked": true, "schema_version": 1, "solution": false}
import pandas as pd
import matplotlib.pyplot as plt
import warnings
idx = pd.IndexSlice
warnings.simplefilter(action='ignore', category=FutureWarning)
from random import seed
from sklearn.metrics import mean_absolute_error
import statsmodels.api as sm
from statsmodels.tsa.stattools import adfuller
from statsmodels.tsa.statespace.sarimax import SARIMAX
from sklearn.linear_model import LinearRegression
import numpy as np
from sklearn.metrics import mean_squared_error
import math
from sklearn.ensemble import GradientBoostingRegressor
import itertools
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa import stattools
import hashlib # for grading purposes
from sklearn.model_selection import ParameterGrid
from pandas.plotting import lag_plot
from statsmodels.tsa.arima_model import ARIMA
plt.rcParams['figure.figsize'] = (12, 4)
# %matplotlib inline
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-efc88f208fd6703c", "locked": true, "schema_version": 1, "solution": false}
# # Functions
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-bb6777a76facfe35", "locked": true, "schema_version": 3, "solution": false, "task": false}
# These functions will be necessary for the exercises. The only one you'll use is _predict_n_periods_. The others are used by the latter.
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-4b3975b136921032", "locked": true, "schema_version": 1, "solution": false}
def build_target(series_, number_of_periods_ahead):
"""
takes a series, turned it into a dataframe, and adds a new column called target
This column is the input series, lagged number_of_periods_ahead into the future
"""
# make a copy
series_ = series_.copy()
series_.name = 'customers'
# make a dataframe from the series
df_ = pd.DataFrame(series_)
# the target column will be the input series, lagged into the future
df_['target'] = series_.shift(-number_of_periods_ahead)
return df_
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-922878325abdd03c", "locked": true, "schema_version": 1, "solution": false}
def separate_last_day(df_):
"""
takes a dataset which has the target and features built
and separates it into the last day
"""
# take the last period
last_period = df_.iloc[-1]
# the last period is now a series, so it's name will be the timestamp
training_data = df_.loc[df_.index < last_period.name]
return last_period, training_data
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-84d68d31545e3063", "locked": true, "schema_version": 1, "solution": false}
def build_some_features(df_, num_periods_lagged=1, num_periods_diffed=0, weekday=False, month=False, rolling=[], holidays=False):
"""
Builds some features by calculating differences between periods
"""
# make a copy
df_ = df_.copy()
# for a few values, get the lags
for i in range(1, num_periods_lagged+1):
# make a new feature, with the lags in the observed values column
df_['lagged_%s' % str(i)] = df_['customers'].shift(i)
# for a few values, get the diffs
for i in range(1, num_periods_diffed+1):
# make a new feature, with the lags in the observed values column
df_['diff_%s' % str(i)] = df_['customers'].diff(i)
for stat in rolling:
df_['rolling_%s'%str(stat)] = df_['customers'].rolling('7D').aggregate(stat)
if weekday == True:
df_['sin_weekday'] = np.sin(2*np.pi*df_.index.weekday/7)
df_['cos_weekday'] = np.sin(2*np.pi*df_.index.weekday/7)
if month == True:
df_['sin_month'] = np.sin(2*np.pi*df_.index.month/12)
df_['cos_month'] = np.sin(2*np.pi*df_.index.month/12)
if holidays == True:
holidays = df_[((df_.index.month==12) & (df_.index.day==25))
|((df_.index.month==1) & (df_.index.day==1))].customers
df_['holidays'] = holidays + 1
df_['holidays'] = df_['holidays'].fillna(0)
return df_
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-f4d816f5cc808636", "locked": true, "schema_version": 1, "solution": false}
def separate_train_and_test_set(last_period_, training_data_, target='target'):
"""
separates training and test set (clue was in the name, really... )
Ok, we were lazy and left the target hardcoded as 'target'. Shame on us.
"""
# anything that isn't a target is a feature
features = [feature for feature in training_data_.columns if feature != target]
# adding a sneaky little dropna to avoid the missing data problem above
X_train = training_data_.dropna()[features]
y_train = training_data_.dropna()[target]
X_last_period = last_period_[features]
return X_train, y_train, X_last_period
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-bc08afac09669562", "locked": true, "schema_version": 1, "solution": false}
def prepare_for_prediction(series_, number_of_periods_ahead, num_periods_lagged, num_periods_diffed, weekday, month, rolling, holidays):
"""
Wrapper to go from the original series to X_train, y_train, X_last_period
"""
# build the target
data_with_target = build_target(series_,
number_of_periods_ahead)
# build the features
data_with_target_and_features = build_some_features(data_with_target,
num_periods_lagged=num_periods_lagged,
num_periods_diffed=num_periods_diffed,
weekday=weekday,
month=month,
rolling=rolling,
holidays=holidays)
# separate train and test data
last_period, training_data = separate_last_day(data_with_target_and_features)
# separate X_train, y_train, and X_test
X_train, y_train, X_last_period = separate_train_and_test_set(last_period,
training_data,
target='target')
# return ALL OF THE THINGS! (well, actually just the ones we need)
return X_train, y_train, X_last_period
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-3a743d99c1cb73f5", "locked": true, "schema_version": 1, "solution": false}
def predict_period_n(series_, model, number_of_periods_ahead, num_periods_lagged, num_periods_diffed, weekday, month, rolling, holidays):
X_train, y_train, X_last_period = prepare_for_prediction(series_,
number_of_periods_ahead,
num_periods_lagged,
num_periods_diffed,
weekday,
month,
rolling,
holidays)
model.fit(X_train, y_train)
return model.predict(X_last_period.values.reshape(1, -1))
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-2c4345e87909ae7f", "locked": true, "schema_version": 1, "solution": false}
def predict_n_periods(series_, n_periods, model, num_periods_lagged, num_periods_diffed=0, weekday=False, month=False,rolling=[], holidays=False):
predictions = []
for period_ahead in range(1, n_periods+1):
pred = predict_period_n(series_=series_,
model=model,
number_of_periods_ahead=period_ahead,
num_periods_lagged=num_periods_lagged,
num_periods_diffed=num_periods_diffed,
weekday=weekday,
month=month,
rolling=rolling,
holidays=holidays)
predictions.append(pred[0])
return predictions
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-c9a3e566cc624451", "locked": true, "schema_version": 1, "solution": false}
# # Let's predict store customers!
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-c89f8456e9e70392", "locked": true, "schema_version": 1, "solution": false}
store = pd.read_csv('data/stores_exercise.csv')
store['date'] = pd.to_datetime(store['date'])
store = store.set_index('date')
store = store.sort_index()
store = store[:-180]
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-5db0587a41ba75ab", "locked": true, "schema_version": 1, "solution": false}
# ##### Plot the series to get an idea of what's going on
# + deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-846a66e59e645d8e", "locked": true, "schema_version": 1, "solution": false}
store.plot(figsize=(16, 4));
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-50c774b21ff04b4b", "locked": true, "schema_version": 1, "solution": false}
# ### Q1: Are there any missing days in the time series? If so, inspect them and decide how to fill them.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-4673648012938702", "locked": false, "schema_version": 1, "solution": true}
# hint1: if the missing dates are holidays you can fill them with 0, since that's
# indication that the store was closed.
# hint2: the missing_value_mask should be a boolean Pandas Series with True or False according to if the value
# missing or not.
# hint3: the missing_value_dates should be a DatetimeIndex with the dates with missing values.
# store_resampled =
# missing_value_mask =
# missing_value_dates =
# store_cleaned =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-37f19bfc31b2d1e0", "locked": true, "points": 2, "schema_version": 1, "solution": false}
expected_hash = 'e9f26ecc6d60870d336e555719e6024c19a07e62c9e7174c36c42b847d50936a'
assert hashlib.sha256(str(missing_value_dates).encode()).hexdigest() == expected_hash
expected_hash = '5ddd3f2573d5c72b9718a3626e6478ec5f7f68f572e36dd3752cb962bf8602a5'
assert hashlib.sha256(str(store_cleaned).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-ef1184e061dcce01", "locked": true, "schema_version": 1, "solution": false}
# ### Q2: Formulate it as time series one-step-ahead prediction
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-52ebd77841c0b4ce", "locked": true, "schema_version": 1, "solution": false}
# ### Q2.1 Create the target, the lags and drop the missing values.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-b2bfb6431053bbb1", "locked": false, "schema_version": 1, "solution": true}
# Note: You should add as features the top 5 lags. To do that, look a the ACF of the time series and chose
# Remember from the previous BLU to look at the ACF you only need to run plot_acf(store)
# the top5 most correlated lags.
#store_features =
#store_features['lag_a'] =
#store_features['lag_b'] =
#store_features['lag_c'] =
#store_features['lag_d'] =
#store_features['lag_e'] =
#store_features['target'] =
#store_features =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-ec83e113b13f6716", "locked": true, "points": 2, "schema_version": 1, "solution": false}
expected_hash = '87d487ce996d6bdbf2f7f919c5c40afc5ab52775f9ddb4798572c3d32e585699'
assert hashlib.sha256(str(store_features.shape).encode()).hexdigest() == expected_hash
expected_hash = '33f8d758034dda4da49f9060da8a1870a17e39738a0a6c891ed0d76e5dbdff8c'
assert hashlib.sha256(str(store_features.iloc[0]).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-7ab428009d677b3c", "locked": true, "schema_version": 1, "solution": false}
# ### Q2.2 Separate the training and test set. The test set consists of the last 60 values, while the training set consists of the rest.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-0aa5b9495555cd7d", "locked": false, "schema_version": 1, "solution": true}
# note: this is a very straightforward question. But you may think: "isn't this one-step-ahead forecasting?
# Why does the test have 60 values" Well, basically this just means we are doing 60 one-step-ahead forecasts.
# This way we obtain a better estimate of how our one-step-ahead model would perform in real life.
# store_train =
# store_test =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-46d22f610161d58e", "locked": true, "points": 1, "schema_version": 1, "solution": false}
expected_hash = '0d597dc2afbcf77932523efe4fa118591cbc3f691191c2471ae95e465c918dd3'
assert hashlib.sha256(str(store_train.index[-1]).encode()).hexdigest() == expected_hash
expected_hash = '23a57d8694a6eb61e0232c384c6939676cf8f8d515298f7f388b9071c22af223'
assert hashlib.sha256(str(store_test.index[0]).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-328a6be750d6b545", "locked": true, "schema_version": 1, "solution": false}
# ### Q2.3 Fit a linear regression to the training set
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-c10406518994a44d", "locked": false, "schema_version": 1, "solution": true}
# X_store_train =
# y_store_train =
# model =
# model.fit()
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-ba2d8a4a2f417e25", "locked": true, "points": 2, "schema_version": 1, "solution": false}
expected_hash = 'cc02033f738b18730fa0d433c70f9f93d2db69418fd9a7bb1f3d1cd8683313ba'
assert hashlib.sha256(str(X_store_train.shape).encode()).hexdigest() == expected_hash
expected_hash = '0fabfb9c5d2a657b34aa1ce1474093e522fc2841b3665fdaaabb67e40eddcf44'
assert hashlib.sha256(str(y_store_train.shape).encode()).hexdigest() == expected_hash
expected_hash = '7624371ed342168f5e33937f92ba58381bb3d29a9a904b0e9e48940649c7e0cf'
assert hashlib.sha256(str(np.round(model.coef_,1)).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-f91888ec4ca37707", "locked": true, "schema_version": 1, "solution": false}
# ### Q2.4 Predict the test set and calculate the MAE
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-c702242ee9013f2d", "locked": false, "schema_version": 1, "solution": true}
# X_store_test =
# y_store_test =
# y_predict =
# test_mae =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-fa53ac00c701b6ad", "locked": true, "points": 2, "schema_version": 1, "solution": false}
expected_hash = 'eca663a9d471ccdde2af2ff1aafbb6bb843c6ef5e21bebc2321143a8166971a7'
assert hashlib.sha256(str(X_store_test.shape).encode()).hexdigest() == expected_hash
expected_hash = '9053d01727f35cfdbbbd7284dba5e54ee557e5fc33084045dbc6fec2cb8730f7'
assert hashlib.sha256(str(y_store_test.shape).encode()).hexdigest() == expected_hash
expected_hash = '8b940be7fb78aaa6b6567dd7a3987996947460df1c668e698eb92ca77e425349'
assert hashlib.sha256(str(np.int(test_mae)).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-1b64863f837f65e9", "locked": true, "schema_version": 1, "solution": false}
# ### Q3 Let's go into multi-step prediction!
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-b8175f4db9400784", "locked": true, "schema_version": 1, "solution": false}
# ### Q3.1 Separate the data into train and test. Use the _predict_n_periods_ function to predict 60 steps ahead using linear regression. Then, calculate the MAE for the test set.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-b4e7b5a3887dcbaa", "locked": false, "schema_version": 1, "solution": true}
# hint: use the cleaned dataset
# Use 7 lags
# store_multistep_train =
# store_multistep_test =
# predictions =
# test_mae =
# YOUR CODE HERE
raise NotImplementedError()
# -
test_mae
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-d64221e5c054b60c", "locked": true, "points": 3, "schema_version": 1, "solution": false}
expected_hash = '299fe0f9c075c40a21029166ce1bd2046db47fcb639a2a13240b5d974961cb0b'
assert hashlib.sha256(str(store_multistep_train.shape).encode()).hexdigest() == expected_hash
expected_hash = '01f6bb906f864c80f90afb9e7d9071b6f6e7222bf67f2de6bce28f55981fbab4'
assert hashlib.sha256(str(store_multistep_test.shape).encode()).hexdigest() == expected_hash
expected_hash = 'f1ee529ef49111208f1c1646c53c8c311c9f093fd7891c1b46d77e98210b018d'
assert hashlib.sha256(str(np.int(predictions[-1])).encode()).hexdigest() == expected_hash
expected_hash = '29db0c6782dbd5000559ef4d9e953e300e2b479eed26d887ef3f92b921c06a67'
assert hashlib.sha256(str(np.int(test_mae)).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-6cecdca4023a9b09", "locked": true, "schema_version": 1, "solution": false}
# ### Q3.2 Separate into train, val and test. Test corresponds to the last 60 values and Val corresponds to the 60 steps before test.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-b4676deaf08fb5b8", "locked": false, "schema_version": 1, "solution": true}
# hint: use the cleaned dataset
# store_multistep_train =
# store_multistep_val =
# store_multistep_test =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-144aa010340df42d", "locked": true, "points": 2, "schema_version": 1, "solution": false}
expected_hash = '98eb165d180aa2cd9255f5a5151c101fac78fba9ce6c24421daa8b64ca2a0288'
assert hashlib.sha256(str(store_multistep_train.shape).encode()).hexdigest() == expected_hash
expected_hash = '01f6bb906f864c80f90afb9e7d9071b6f6e7222bf67f2de6bce28f55981fbab4'
assert hashlib.sha256(str(store_multistep_val.shape).encode()).hexdigest() == expected_hash
expected_hash = '01f6bb906f864c80f90afb9e7d9071b6f6e7222bf67f2de6bce28f55981fbab4'
assert hashlib.sha256(str(store_multistep_test.shape).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-a252157fed7db039", "locked": true, "schema_version": 1, "solution": false}
# ### Q3.3 Are the holidays, weekday and the month of the year useful features to the model?
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-16a7b564ac712979", "locked": false, "schema_version": 1, "solution": true}
# %%time
# Create a parameter grid using the gradient boosting regressor as a model.
# Use 5 lags, zero diffs and no rollings
# For the gradient boosting regressor use n_estimators=20 and random_state=10
# Use a for cycle to find the group of params that minimizes the MAE on the validation set.
# hint: to have no rollings in the predict_n_periods you should send an empty lists of lists: [[]]
#param_grid =
# grid =
# for params in grid:
# predictions =
# best_params =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-27e82c63c7934897", "locked": true, "points": 3, "schema_version": 1, "solution": false}
expected_hash = '3cbc87c7681f34db4617feaa2c8801931bc5e42d8d0f560e756dd4cd92885f18'
assert hashlib.sha256(str(best_params['weekday']).encode()).hexdigest() == expected_hash
expected_hash = '3cbc87c7681f34db4617feaa2c8801931bc5e42d8d0f560e756dd4cd92885f18'
assert hashlib.sha256(str(best_params['month']).encode()).hexdigest() == expected_hash
expected_hash = '3cbc87c7681f34db4617feaa2c8801931bc5e42d8d0f560e756dd4cd92885f18'
assert hashlib.sha256(str(best_params['holidays']).encode()).hexdigest() == expected_hash
# + [markdown] deletable=false editable=false nbgrader={"grade": false, "grade_id": "cell-cd3355c62f859eeb", "locked": true, "schema_version": 1, "solution": false}
# ### Q3.4 Train a model with the best combination and predict the test set. Calculate the corresponding MAE.
# + deletable=false nbgrader={"grade": false, "grade_id": "cell-a8fdbd5c2eaf09dc", "locked": false, "schema_version": 1, "solution": true}
# For the gradient boosting regressor use n_estimators=20 and random_state=10
# We expect you to train the final model with train and val together.
# store_multistep_train_val =
# predictions =
# test_mae =
# YOUR CODE HERE
raise NotImplementedError()
# + deletable=false editable=false nbgrader={"grade": true, "grade_id": "cell-cc1f893477248c54", "locked": true, "points": 3, "schema_version": 1, "solution": false}
expected_hash = '434c9b5ae514646bbd91b50032ca579efec8f22bf0b4aac12e65997c418e0dd6'
assert hashlib.sha256(str(np.int(test_mae)).encode()).hexdigest() == expected_hash
|
S03 - Time Series/BLU06 - Machine Learning for Time Series/Exercise notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Edward's Personal Notebook
# dataset is the "Police Incident Blotter From 2015 - 2022". Or in other words, the amount of police incidents in Pittsburgh from 2015-2022
# importing required libraries and settings
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import geopandas
pd.set_option('display.max_colwidth', None)
# Reading in data
df = pd.read_csv("crimeall.csv", sep = ',')
df.head(5) # print dataframe sample
# +
#I removed unneeded columns like: incident tract, council district, and etc.
# The main idea is lower number of police incidents correlates to safety(my submetric), so I want to see each neighborhood's police incident count.
df_sub = df.INCIDENTNEIGHBORHOOD.value_counts() # stores count of each unique value
df_reverse = df_sub.iloc[::-1] # reversing data frame to show from lowest value....a better way might exist
df_reverse.head(15)
# +
# <NAME> appears 3 different times so there is something wrong with the dataset.
# Below removed series are sourced from the Official Designations from the City of Pittsburgh
df = df[df['INCIDENTNEIGHBORHOOD'] != 'Outside County'] # This is not a Pittsburgh Neighborhood
df = df[df['INCIDENTNEIGHBORHOOD'] != 'Outside State'] # This is not a Pittsburgh Neighborhood
df = df[df['INCIDENTNEIGHBORHOOD'] != 'Mt. <NAME>'] # This is not a Pittsburgh Neighborhood ***Mt. Oliver is a
# Pittsburgh neighborhood, but Mt. <NAME> is not.***
# I merged Mount Oliver and Mt. Oliver Neighborhood, I checked the street addresses of them and they are in the same neighborhood. This error
# in the dataset might be due to Mt. <NAME> being included but not being a neighborhood a part of Pittsburgh. So perhaps some officers added
# the "neighborhood" at the end to help clarify.
df = df.replace(to_replace = 'Mt. Oliver Neighborhood', value ='Mount Oliver')
# df # testing purposes
# -
# Showing neighbors again after removing the non-neighborhoods areas from data set
df_sub = df.INCIDENTNEIGHBORHOOD.value_counts() # stores count of each unique value
df_reverse = df_sub.iloc[::-1] # reversing data frame to show from lowest value....a better way might exist
df_reverse # showing the df
# +
plt.rcParams["figure.figsize"] = [20,9] # setting graph size or else it's too small
df_reverse10 = df_reverse.iloc[:10] # show only first 10 neighborhoods
df_reverse10.plot.bar() # graphing our dataframe
plt.title("Neighborhood Police Incidents From 2015 - 2022", fontsize = 14) # graph title
plt.xticks(fontsize = 12,rotation=45) # rorate x-axis for readability
plt.xlabel("Neighborhoods", fontsize = 12) # set x-axis label
plt.yticks(fontsize = 12) # rotate x-axis for readability
plt.ylabel("Police Incidents", fontsize = 12, rotation = 0, labelpad = 50) # y-axis label, padding because it enters the graph body
plt.show() # show graph more cleanly
# herrs island is part of troy hill neighborhood
# -
# ## Conclusion - Edward
# This data set had over 277000 rows of data, each representing 1 police incident in a Pittsburgh neighborhood over the span of 7 years. I organized the data so that each neighborhood had a score(the number of police incidents). The lower the score, the safer the neighborhood is. In result, the Troy Hill-Herrs Island Neighborhood had the least amount of incidents, and therefore was the "best" neighborhood for my submetric.
#
# Something I'd like to mention is: the data set didn't have population density data. I think with a larger population, it is only natural for police incidents to go up. So I think this factor would also affect the information gained from this data set and even potentially become a shortcoming for this interpretation.
#
# The data driven determination of the "best" neighbohood is different from my personal favorite. East Liberty is my personal favorite, simply because I live there.........although......definitely wouldn't recommend living here. Kinda dangerous at night. O.O
# +
# Merge map with csv
neighborhoods = geopandas.read_file("Neighborhoods/Neighborhoods_.shp") # read shp
df_sub['East Liberty'] = 9999999999
df_merge = df_sub.rename_axis('INCIDENTNEIGHBORHOOD').to_frame('count')
steps_map = neighborhoods.merge(df_merge, how='left', left_on='hood', right_on='INCIDENTNEIGHBORHOOD')
steps_map.plot(column='count', # set the data to be used for coloring
cmap='OrRd', # choose a color palette
edgecolor="white", # outline the districts in white
legend=True, # show the legend
legend_kwds={'label': "Lit-ness" }, # label the legend
figsize=(15, 10), # set the size
missing_kwds={"color": "white"} # set disctricts with no data to gray
)
plt.title("WHO DA BEST NEIGHBORHOOD...EAST LIBERTYYYYY", fontsize = 20) # graph title
plt.xticks(fontsize = 12,rotation=0) # rorate x-axis for readability
plt.show
# -
# _____________________________________________________________________________________________________________________________________________________________________________________________________
# Wentao's Personal Notebook
#read the data
df1 = pd.read_csv("ISP.csv")
#delete the invalid data
df2 = df1[df1.MaxCIRDown >= 16]
df3 = df1[df1.MaxCIRUp >= 16]
#sort
value_count = df3['Neighborhood'].value_counts()
df4 = value_count.rename_axis('Neighborhood').to_frame('count')
df4.head(20)
# +
plt.rcParams["figure.figsize"] = [20,9] # setting graph size or else it's too small
df5 = df4.iloc[:10] # show only first 10 neighborhoods
df5.plot.bar() # graphing our dataframe
plt.title("Best Internet Service of Neighborhood", fontsize = 14) # graph title
plt.xticks(fontsize = 12,rotation=45) # rorate x-axis for readability
plt.xlabel("Neighborhoods", fontsize = 12) # set x-axis label
plt.yticks(fontsize = 12) # rotate x-axis for readability
plt.ylabel("Number of ISPs", fontsize = 12, rotation = 0, labelpad = 50) # y-axis label, padding because it enters the graph body
plt.show() # show graph more cleanly
# -
neighborhoods = geopandas.read_file("Neighborhoods/Neighborhoods_.shp") # read in the shapefile
# do the merge
ISP_map = neighborhoods.merge(df4, how='left', left_on='hood', right_on='Neighborhood')
ISP_map.plot(column='count', # set the data to be used for coloring
cmap='OrRd', # choose a color palette
edgecolor="white", # outline the districts in white
legend=True, # show the legend
legend_kwds={'label': "Number of ISPs"}, # label the legend
figsize=(15, 10), # set the size
missing_kwds={"color": "lightgrey"} # set disctricts with no data to gray
)
plt.title("Best Internet Service - Downtown", fontsize = 20) # graph title
# ##### Wentao's Conclusion
#
# According to my analysis, Central Business District is the best Neighbourhood - Downtown. My data analysis also has some shortcomings. We consider the living experience of the residents, but my results show that the most convenient area for the Internet is dowtown, which has a large number of companies. It's not a residential area. In addition, the effect will be better if the population and area of each neighborhood are analyzed.
# _____________________________________________________________________________________________________________________________________________________________________________________________________
# ## Final Conclusion
# +
df_reverse10 = df_reverse.iloc[:19]
plt.rcParams["figure.figsize"] = [20,9] # setting graph size or else it's too small
df_reverse10.plot.bar() # graphing our dataframe
plt.title("Neighborhood Police Incidents From 2015 - 2022", fontsize = 14) # graph title
plt.xticks(fontsize = 12,rotation=45) # rorate x-axis for readability
plt.xlabel("Neighborhoods", fontsize = 12) # set x-axis label
plt.yticks(fontsize = 12) # rotate x-axis for readability
plt.ylabel("Police Incidents", fontsize = 12, rotation = 0, labelpad = 50) # y-axis label, padding because it enters the graph body
plt.show()
# +
plt.rcParams["figure.figsize"] = [20,9] # setting graph size or else it's too small
df6 = df4.iloc[:18] # show only first 10 neighborhoods
df6.plot.bar() # graphing our dataframe
plt.title("Best Internet Service of Neighborhood", fontsize = 14) # graph title
plt.xticks(fontsize = 12,rotation=45) # rorate x-axis for readability
plt.xlabel("Neighborhoods", fontsize = 12) # set x-axis label
plt.yticks(fontsize = 12) # rotate x-axis for readability
plt.ylabel("Number of ISPs", fontsize = 12, rotation = 0, labelpad = 50) # y-axis label, padding because it enters the graph body
plt.show()
# -
# ###### Conclude
# Yuxuan
# By analyzing the data, the neighborhood areas of Pittsburgh were first screened. I ranked the infection and reinfections rates,the number of hospitalizations, and mortality rates for each area by adding up several items of data and then ranking them from lowest to highest. I concluded that Chateau had the lowest risk of infection or death, so it was the safest place I selected.
# ## Overall Conclusion
# In order to combine the two submetrics, we had to find a common neighborhood between the two data sets (police incidents and internet service). In the end, we ended with Central Northside. As shown from the two modified graphs above, Central Northside was ranked 19th out of all neighborhoods in regards to police incidents and was ranked 18th in terms of having ISP providers. Our overall metric is living quality, and Central Northside was the "best" in our data driven determination. There might be some biase-ness however with our choice however. The reason Central Northside might rank relatively high in both submetrics might be because of its small area. If it has a small enough area, it can have a large amount of internet providers and also have a low amount of police incidents.
#
|
Final Notebook/Overall Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Generating C Code to implement Method of Lines Timestepping for Explicit Runge Kutta Methods
#
# ## Authors: <NAME> & <NAME>
#
# ## This tutorial notebook generates three blocks of C Code in order to perform Method of Lines timestepping.
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). All Runge-Kutta (RK) Butcher tables were validated using truncated Taylor series in [a separate module](Tutorial-RK_Butcher_Table_Validation.ipynb). Finally, C-code implementation of RK4 was validated against a trusted version. C-code implementations of other RK methods seem to work as expected in the context of solving the scalar wave equation in Cartesian coordinates.
#
# ### NRPy+ Source Code for this module:
# * [MoLtimestepping/C_Code_Generation.py](../edit/MoLtimestepping/C_Code_Generation.py)
# * [MoLtimestepping/RK_Butcher_Table_Dictionary.py](../edit/MoLtimestepping/RK_Butcher_Table_Dictionary.py) ([**Tutorial**](Tutorial-RK_Butcher_Table_Dictionary.ipynb)) Stores the Butcher tables for the explicit Runge Kutta methods
#
# ## Introduction:
#
# When numerically solving a partial differential equation initial-value problem, subject to suitable boundary conditions, we implement Method of Lines to "integrate" the solution forward in time.
#
#
# ### The Method of Lines:
#
# Once we have the initial data for a PDE, we "evolve it forward in time", using the [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html). In short, the Method of Lines enables us to handle
# 1. the **spatial derivatives** of an initial value problem PDE using **standard finite difference approaches**, and
# 2. the **temporal derivatives** of an initial value problem PDE using **standard strategies for solving ordinary differential equations (ODEs), like Runge Kutta methods** so long as the initial value problem PDE can be written in the first-order-in-time form
# $$\partial_t \vec{f} = \mathbf{M}\ \vec{f},$$
# where $\mathbf{M}$ is an $N\times N$ matrix containing only *spatial* differential operators that act on the $N$-element column vector $\vec{f}$. $\mathbf{M}$ may not contain $t$ or time derivatives explicitly; only *spatial* partial derivatives are allowed to appear inside $\mathbf{M}$.
#
# You may find the next module [Tutorial-ScalarWave](Tutorial-ScalarWave.ipynb) extremely helpful as an example for implementing the Method of Lines for solving the Scalar Wave equation in Cartesian coordinates.
#
# ### Generating the C code:
#
# This module describes how core C functions are generated to implement Method of Lines timestepping for a specified RK method. There are three core functions:
#
# 1. Allocate memory for gridfunctions.
# 1. Step forward the solution one full timestep.
# 1. Free memory for gridfunctions.
#
# The first function is called first, then the second function is repeated within a loop to a fixed "final" time (such that the end state of each iteration is the initial state for the next iteration), and the third function is called at the end of the calculation.
#
# The generated codes are essential for a number of Start-to-Finish example tutorial notebooks that demonstrate how to numerically solve hyperbolic PDEs.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Initialize needed Python/NRPy+ modules
# 1. [Step 2](#diagonal): Checking if Butcher Table is Diagonal
# 1. [Step 3](#ccode): Generating the C Code
# 1. [Step 3.a](#generategfnames): `generate_gridfunction_names()`: Uniquely and descriptively assign names to sets of gridfunctions
# 1. [Step 3.b](#alloc): Memory allocation: `MoL_malloc_y_n_gfs()` and `MoL_malloc_non_y_n_gfs()`
# 1. [Step 3.c](#molstep): Take one Method of Lines time step: `MoL_step_forward_in_time()`
# 1. [Step 3.d](#free): Memory deallocation: `MoL_free_memory()`
# 1. [Step 3.e](#nrpybasicdefines): Define & register `MoL_gridfunctions_struct` in `NRPy_basic_defines.h`: `NRPy_basic_defines_MoL_timestepping_struct()`
# 1. [Step 3.f](#setupall): Add all MoL C codes to C function dictionary, and add MoL definitions to `NRPy_basic_defines.h`: `register_C_functions_and_NRPy_basic_defines()`
# 1. [Step 4](#code_validation): Code Validation against `MoLtimestepping.RK_Butcher_Table_Generating_C_Code` NRPy+ module
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize needed Python/NRPy+ modules [Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# Let's start by importing all the needed modules from Python/NRPy+:
import sympy as sp # Import SymPy, a computer algebra system written entirely in Python
import os, sys # Standard Python modules for multiplatform OS-level functions
from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
from outputC import add_to_Cfunction_dict, indent_Ccode, outC_NRPy_basic_defines_h_dict, superfast_uniq, outputC # NRPy+: Basic C code output functionality
# <a id='diagonal'></a>
#
# # Step 2: Checking if a Butcher table is Diagonal [Back to [top](#toc)\]
# $$\label{diagonal}$$
#
# A diagonal Butcher table takes the form
#
# $$\begin{array}{c|cccccc}
# 0 & \\
# a_1 & a_1 & \\
# a_2 & 0 & a_2 & \\
# a_3 & 0 & 0 & a_3 & \\
# \vdots & \vdots & \ddots & \ddots & \ddots \\
# a_s & 0 & 0 & 0 & \cdots & a_s \\ \hline
# & b_1 & b_2 & b_3 & \cdots & b_{s-1} & b_s
# \end{array}$$
#
# where $s$ is the number of required predictor-corrector steps for a given RK method (see [<NAME>. (2008)](https://onlinelibrary.wiley.com/doi/book/10.1002/9780470753767)). One known diagonal RK method is the classic RK4 represented in Butcher table form as:
#
# $$\begin{array}{c|cccc}
# 0 & \\
# 1/2 & 1/2 & \\
# 1/2 & 0 & 1/2 & \\
# 1 & 0 & 0 & 1 & \\ \hline
# & 1/6 & 1/3 & 1/3 & 1/6
# \end{array} $$
#
# Diagonal Butcher tables are nice when it comes to saving required memory space. Each new step for a diagonal RK method, when computing the new $k_i$, does not depend on the previous calculation, and so there are ways to save memory. Significantly so in large three-dimensional spatial grid spaces.
# +
# Check if Butcher Table is diagonal
def diagonal(key):
Butcher = Butcher_dict[key][0]
L = len(Butcher)-1 # Establish the number of rows to check for diagonal trait, all bust last row
row_idx = 0 # Initialize the Butcher table row index
for i in range(L): # Check all the desired rows
for j in range(1,row_idx): # Check each element before the diagonal element in a row
if Butcher[i][j] != sp.sympify(0): # If any non-diagonal coeffcient is non-zero,
# then the table is not diagonal
return False
row_idx += 1 # Update to check the next row
return True
# Loop over all Butcher tables to check whether each is diagonal or not
for key, value in Butcher_dict.items():
if diagonal(key) == True:
print("The MoL method "+str(key)+" is diagonal!")
else:
print("The MoL method "+str(key)+" is NOT diagonal!")
# -
# <a id='ccode'></a>
#
# # Step 3: Generating the C Code [Back to [top](#toc)\]
# $$\label{ccode}$$
#
# The following sections build up the C code for implementing the [Method of Lines timestepping algorithm](http://www.scholarpedia.org/article/Method_of_lines) for solving hyperbolic PDEs.
#
# **First an important note on efficiency:**
#
# Memory efficiency is incredibly important here, as $\vec{f}$ is usually the largest object in memory.
#
# If we only made use of the Butcher tables without concern for memory efficiency, `generate_gridfunction_names()` and `MoL_step_forward_in_time()` would be very simple functions.
#
# It turns out that several of the Runge-Kutta-like methods in MoL can be made more efficient; for example "RK4" can be performed using only 4 "timelevels" of $\vec{f}$ in memory (i.e., a total memory usage of `sizeof(f) * 4`). A naive implementation might use 5 or 6 copies. RK-like methods that have diagonal Butcher tables can be made far more efficient than the naive approach.
#
# **Exercise to student:** Improve the efficiency of other RK-like methods.
# <a id='generategfnames'></a>
#
# ## Step 3.a: `generate_gridfunction_names()`: Uniquely and descriptively assign names to sets of gridfunctions [Back to [top](#toc)\]
# $$\label{generategfnames}$$
#
# `generate_gridfunction_names()` names gridfunctions to be consistent with a given RK substep. For example we might call the set of gridfunctions stored at substep $k_1$ `k1_gfs`.
# Each MoL method has its own set of names for groups of gridfunctions,
# aiming to be sufficiently descriptive. So for example a set of
# gridfunctions that store "k_1" in an RK-like method could be called
# "k1_gfs".
def generate_gridfunction_names(MoL_method = "RK4"):
# Step 3.a: MoL gridfunctions fall into 3 overlapping categories:
# 1) y_n=y_i(t_n) gridfunctions y_n_gfs, which stores data for the vector of gridfunctions y_i at t_n,
# the start of each MoL timestep.
# 2) non-y_n gridfunctions, needed to compute the data at t_{n+1}. Often labeled with k_i in the name,
# these gridfunctions are *not* needed at the start of each timestep, so are available for temporary
# storage when gridfunctions needed for diagnostics are computed at the start of each timestep.
# These gridfunctions can also be freed during a regrid, to enable storage for the post-regrid
# destination y_n_gfs.
# 3) Diagnostic output gridfunctions diagnostic_output_gfs, which simply uses the memory from auxiliary
# gridfunctions at one auxiliary time to compute diagnostics at t_n.
# Here we specify which gridfunctions fall into each category, starting with the obvious: y_n_gridfunctions
y_n_gridfunctions = "y_n_gfs"
# Next the less-obvious, which depend on non-y_n_gfs
non_y_n_gridfunctions_list = []
# No matter the method we define gridfunctions "y_n_gfs" to store the initial data
if diagonal(MoL_method) and "RK3" in MoL_method:
non_y_n_gridfunctions_list.append("k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs")
non_y_n_gridfunctions_list.append("k2_or_y_nplus_a32_k2_gfs")
diagnostic_gridfunctions_point_to = "k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs"
else:
if not diagonal(MoL_method): # Allocate memory for non-diagonal Butcher tables
# Determine the number of k_i steps based on length of Butcher Table
num_k = len(Butcher_dict[MoL_method][0])-1
# For non-diagonal tables an intermediate gridfunction "next_y_input" is used for rhs evaluations
non_y_n_gridfunctions_list.append("next_y_input_gfs")
for i in range(num_k): # Need to allocate all k_i steps for a given method
non_y_n_gridfunctions_list.append("k" + str(i + 1) + "_gfs")
diagnostic_gridfunctions_point_to = "k1_gfs"
else: # Allocate memory for diagonal Butcher tables, which use a "y_nplus1_running_total gridfunction"
non_y_n_gridfunctions_list.append("y_nplus1_running_total_gfs")
if MoL_method != 'Euler': # Allocate memory for diagonal Butcher tables that aren't Euler
# Need k_odd for k_1,3,5... and k_even for k_2,4,6...
non_y_n_gridfunctions_list.append("k_odd_gfs")
non_y_n_gridfunctions_list.append("k_even_gfs")
diagnostic_gridfunctions_point_to = "y_nplus1_running_total_gfs"
non_y_n_gridfunctions_list.append("auxevol_gfs")
return y_n_gridfunctions, non_y_n_gridfunctions_list, diagnostic_gridfunctions_point_to
# <a id='alloc'></a>
#
# ## Step 3.b: Memory allocation: `MoL_malloc_y_n_gfs()` and `MoL_malloc_non_y_n_gfs()`: [Back to [top](#toc)\]
# $$\label{alloc}$$
#
# Generation of C functions `MoL_malloc_y_n_gfs()` and `MoL_malloc_non_y_n_gfs()` read the full list of needed lists of gridfunctions, provided by (Python) function `generate_gridfunction_names()`, to allocate space for all gridfunctions.
# add_to_Cfunction_dict_MoL_malloc() registers
# MoL_malloc_y_n_gfs() and
# MoL_malloc_non_y_n_gfs(), which allocate memory for
# the indicated sets of gridfunctions
def add_to_Cfunction_dict_MoL_malloc(MoL_method, which_gfs):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = "Method of Lines (MoL) for \"" + MoL_method + "\" method: Allocate memory for \""+which_gfs+"\" gridfunctions\n"
desc += " * y_n_gfs are used to store data for the vector of gridfunctions y_i at t_n, at the start of each MoL timestep\n"
desc += " * non_y_n_gfs are needed for intermediate (e.g., k_i) storage in chosen MoL method\n"
c_type = "void"
y_n_gridfunctions, non_y_n_gridfunctions_list, diagnostic_gridfunctions_point_to = \
generate_gridfunction_names(MoL_method = MoL_method)
gridfunctions_list = []
if which_gfs == "y_n_gfs":
gridfunctions_list = [y_n_gridfunctions]
elif which_gfs == "non_y_n_gfs":
gridfunctions_list = non_y_n_gridfunctions_list
else:
print("ERROR: which_gfs = \"" + which_gfs + "\" unrecognized.")
sys.exit(1)
name = "MoL_malloc_" + which_gfs
params = "const paramstruct *restrict params, MoL_gridfunctions_struct *restrict gridfuncs"
body = "const int Nxx_plus_2NGHOSTS_tot = Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2;\n"
for gridfunctions in gridfunctions_list:
num_gfs = "NUM_EVOL_GFS"
if gridfunctions == "auxevol_gfs":
num_gfs = "NUM_AUXEVOL_GFS"
body += "gridfuncs->" + gridfunctions + " = (REAL *restrict)malloc(sizeof(REAL) * " + num_gfs + " * Nxx_plus_2NGHOSTS_tot);\n"
body += "\ngridfuncs->diagnostic_output_gfs = gridfuncs->" + diagnostic_gridfunctions_point_to + ";\n"
add_to_Cfunction_dict(
includes=includes,
desc=desc,
c_type=c_type, name=name, params=params,
body=indent_Ccode(body, " "),
rel_path_to_Cparams=os.path.join("."))
# <a id='molstep'></a>
#
# ## Step 3.c: Take one Method of Lines time step: `MoL_step_forward_in_time()` [Back to [top](#toc)\]
# $$\label{molstep}$$
#
# An MoL step consists in general of a series of Runge-Kutta-like substeps, and the `MoL_step_forward_in_time()` C function pulls together all of these substeps.
#
# The basic C code for an MoL substep, set up by the Python function `single_RK_substep()` below, is as follows.
#
# 1. Evaluate the right-hand side of $\partial_t \vec{f}=$ `RHS`, to get the time derivative of the set of gridfunctions $\vec{f}$ at our current time.
# 1. Perform the Runge-Kutta update, which depends on $\partial_t \vec{f}$ on the current and sometimes previous times.
# 1. Call post-right-hand side functions as desired.
#
# The `single_RK_substep_input_symbolic()` function generates the C code for performing the above steps, applying substitutions for e.g., `RK_INPUT_GFS` and `RK_OUTPUT_GFS` as appropriate. `single_RK_substep_input_symbolic()` supports SIMD-capable code generation.
# single_RK_substep_input_symbolic() performs necessary replacements to
# define C code for a single RK substep
# (e.g., computing k_1 and then updating the outer boundaries)
def single_RK_substep_input_symbolic(commentblock, RHS_str, RHS_input_str, RHS_output_str, RK_lhss_list, RK_rhss_list,
post_RHS_list, post_RHS_output_list, enable_SIMD=False,
enable_griddata=False, gf_aliases="", post_post_RHS_string=""):
return_str = commentblock + "\n"
if not isinstance(RK_lhss_list, list):
RK_lhss_list = [RK_lhss_list]
if not isinstance(RK_rhss_list, list):
RK_rhss_list = [RK_rhss_list]
if not isinstance(post_RHS_list, list):
post_RHS_list = [post_RHS_list]
if not isinstance(post_RHS_output_list, list):
post_RHS_output_list = [post_RHS_output_list]
indent = ""
if enable_griddata:
return_str += "{\n" + indent_Ccode(gf_aliases, " ")
indent = " "
# Part 1: RHS evaluation:
return_str += indent_Ccode(str(RHS_str).replace("RK_INPUT_GFS", str(RHS_input_str).replace("gfsL", "gfs")).
replace("RK_OUTPUT_GFS", str(RHS_output_str).replace("gfsL", "gfs"))+"\n", indent=indent)
# Part 2: RK update
if enable_SIMD:
return_str += "#pragma omp parallel for\n"
return_str += indent + "for(int i=0;i<Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2*NUM_EVOL_GFS;i+=SIMD_width) {\n"
else:
return_str += indent + "LOOP_ALL_GFS_GPS(i) {\n"
type = "REAL"
if enable_SIMD:
type = "REAL_SIMD_ARRAY"
RK_lhss_str_list = []
for i, el in enumerate(RK_lhss_list):
if enable_SIMD:
RK_lhss_str_list.append(indent + "const REAL_SIMD_ARRAY __RHS_exp_" + str(i))
else:
RK_lhss_str_list.append(indent + str(el).replace("gfsL", "gfs[i]"))
read_list = []
for el in RK_rhss_list:
for read in list(sp.ordered(el.free_symbols)):
read_list.append(read)
read_list_uniq = superfast_uniq(read_list)
for el in read_list_uniq:
if str(el) != "dt":
if enable_SIMD:
return_str += indent + " const " + type + " " + str(el) + " = ReadSIMD(&" + str(el).replace("gfsL", "gfs[i]") + ");\n"
else:
return_str += indent + " const " + type + " " + str(el) + " = " + str(el).replace("gfsL", "gfs[i]") + ";\n"
if enable_SIMD:
return_str += indent + " const REAL_SIMD_ARRAY DT = ConstSIMD(dt);\n"
preindent = "1"
if enable_griddata:
preindent = "2"
kernel = outputC(RK_rhss_list, RK_lhss_str_list, filename="returnstring",
params="includebraces=False,preindent="+preindent+",outCverbose=False,enable_SIMD="+str(enable_SIMD))
if enable_SIMD:
return_str += kernel.replace("dt", "DT")
for i, el in enumerate(RK_lhss_list):
return_str += " WriteSIMD(&" + str(el).replace("gfsL", "gfs[i]") + ", __RHS_exp_" + str(i) + ");\n"
else:
return_str += kernel
return_str += indent + "}\n"
# Part 3: Call post-RHS functions
for post_RHS, post_RHS_output in zip(post_RHS_list, post_RHS_output_list):
return_str += indent_Ccode(post_RHS.replace("RK_OUTPUT_GFS", str(post_RHS_output).replace("gfsL", "gfs")))
if enable_griddata:
return_str += "}\n"
for post_RHS, post_RHS_output in zip(post_RHS_list, post_RHS_output_list):
return_str += indent_Ccode(post_post_RHS_string.replace("RK_OUTPUT_GFS", str(post_RHS_output).replace("gfsL", "gfs")), "")
return return_str
# In the `add_to_Cfunction_dict_MoL_step_forward_in_time()` Python function below we construct and register the core C function for MoL timestepping: `MoL_step_forward_in_time()`. `MoL_step_forward_in_time()` implements Butcher tables for Runge-Kutta-like methods, leveraging the `single_RK_substep()` helper function above as needed. Again, we aim for maximum memory efficiency so that e.g., RK4 needs to store only 4 levels of $\vec{f}$.
def add_to_Cfunction_dict_MoL_step_forward_in_time(MoL_method,
RHS_string = "", post_RHS_string = "", post_post_RHS_string="",
enable_rfm=False, enable_curviBCs=False, enable_SIMD=False,
enable_griddata=False):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
if enable_SIMD:
includes += [os.path.join("SIMD", "SIMD_intrinsics.h")]
desc = "Method of Lines (MoL) for \"" + MoL_method + "\" method: Step forward one full timestep.\n"
c_type = "void"
name = "MoL_step_forward_in_time"
if enable_griddata:
params = "griddata_struct *restrict griddata, const REAL dt"
else:
params = "const paramstruct *restrict params, "
if enable_rfm:
params += "const rfm_struct *restrict rfmstruct, "
else:
params += "REAL *restrict xx[3], "
if enable_curviBCs:
params += "const bc_struct *restrict bcstruct, "
params += "MoL_gridfunctions_struct *restrict gridfuncs, const REAL dt"
indent = "" # We don't bother with an indent here.
body = indent + "// C code implementation of -={ " + MoL_method + " }=- Method of Lines timestepping.\n\n"
y_n_gridfunctions, non_y_n_gridfunctions_list, _throwaway = generate_gridfunction_names(MoL_method)
if enable_griddata:
gf_prefix = "griddata->gridfuncs."
else:
gf_prefix = "gridfuncs->"
gf_aliases = """// Set gridfunction aliases from gridfuncs struct
REAL *restrict """ + y_n_gridfunctions + " = "+gf_prefix + y_n_gridfunctions + """; // y_n gridfunctions
// Temporary timelevel & AUXEVOL gridfunctions:\n"""
for gf in non_y_n_gridfunctions_list:
gf_aliases += "REAL *restrict " + gf + " = "+gf_prefix + gf + ";\n"
if enable_griddata:
gf_aliases += "paramstruct *restrict params = &griddata->params;\n"
gf_aliases += "const rfm_struct *restrict rfmstruct = &griddata->rfmstruct;\n"
gf_aliases += "const bc_struct *restrict bcstruct = &griddata->bcstruct;\n"
for i in ["0", "1", "2"]:
gf_aliases += "const int Nxx_plus_2NGHOSTS" + i + " = griddata->params.Nxx_plus_2NGHOSTS" + i + ";\n"
if not enable_griddata:
body += gf_aliases
# Implement Method of Lines (MoL) Timestepping
Butcher = Butcher_dict[MoL_method][0] # Get the desired Butcher table from the dictionary
num_steps = len(Butcher)-1 # Specify the number of required steps to update solution
# Diagonal RK3 only!!!
dt = sp.Symbol("dt", real=True)
if diagonal(MoL_method) and "RK3" in MoL_method:
# In a diagonal RK3 method, only 3 gridfunctions need be defined. Below implements this approach.
y_n_gfs = sp.Symbol("y_n_gfsL", real=True)
k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs = sp.Symbol("k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfsL", real=True)
k2_or_y_nplus_a32_k2_gfs = sp.Symbol("k2_or_y_nplus_a32_k2_gfsL", real=True)
# k_1
body += """
// In a diagonal RK3 method like this one, only 3 gridfunctions need be defined. Below implements this approach.
// Using y_n_gfs as input, k1 and apply boundary conditions\n"""
body += single_RK_substep_input_symbolic(
commentblock="""// -={ START k1 substep }=-
// RHS evaluation:
// 1. We will store k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs now as
// ... the update for the next rhs evaluation y_n + a21*k1*dt
// Post-RHS evaluation:
// 1. Apply post-RHS to y_n + a21*k1*dt""",
RHS_str=RHS_string,
RHS_input_str=y_n_gfs,
RHS_output_str=k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs,
RK_lhss_list=[k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs],
RK_rhss_list=[Butcher[1][1]*k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs*dt + y_n_gfs],
post_RHS_list=[post_RHS_string], post_RHS_output_list=[k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string) + "// -={ END k1 substep }=-\n\n"
# k_2
body += single_RK_substep_input_symbolic(
commentblock="""// -={ START k2 substep }=-
// RHS evaluation:
// 1. Reassign k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs to be the running total y_{n+1}; a32*k2*dt to the running total
// 2. Store k2_or_y_nplus_a32_k2_gfs now as y_n + a32*k2*dt
// Post-RHS evaluation:
// 1. Apply post-RHS to both y_n + a32*k2 (stored in k2_or_y_nplus_a32_k2_gfs)
// ... and the y_{n+1} running total, as they have not been applied yet to k2-related gridfunctions""",
RHS_str=RHS_string,
RHS_input_str=k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs,
RHS_output_str=k2_or_y_nplus_a32_k2_gfs,
RK_lhss_list=[k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs, k2_or_y_nplus_a32_k2_gfs],
RK_rhss_list=[Butcher[3][1]*(k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs - y_n_gfs)/Butcher[1][1] + y_n_gfs + Butcher[3][2]*k2_or_y_nplus_a32_k2_gfs*dt,
Butcher[2][2]*k2_or_y_nplus_a32_k2_gfs*dt + y_n_gfs],
post_RHS_list=[post_RHS_string, post_RHS_string],
post_RHS_output_list=[k2_or_y_nplus_a32_k2_gfs,
k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string) + "// -={ END k2 substep }=-\n\n"
# k_3
body += single_RK_substep_input_symbolic(
commentblock="""// -={ START k3 substep }=-
// RHS evaluation:
// 1. Add k3 to the running total and save to y_n
// Post-RHS evaluation:
// 1. Apply post-RHS to y_n""",
RHS_str=RHS_string,
RHS_input_str=k2_or_y_nplus_a32_k2_gfs, RHS_output_str=y_n_gfs,
RK_lhss_list=[y_n_gfs],
RK_rhss_list=[k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs + Butcher[3][3]*y_n_gfs*dt],
post_RHS_list=[post_RHS_string],
post_RHS_output_list=[y_n_gfs],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string) + "// -={ END k3 substep }=-\n\n"
else:
y_n = sp.Symbol("y_n_gfsL", real=True)
if not diagonal(MoL_method):
for s in range(num_steps):
next_y_input = sp.Symbol("next_y_input_gfsL", real=True)
# If we're on the first step (s=0), we use y_n gridfunction as input.
# Otherwise next_y_input is input. Output is just the reverse.
if s == 0: # If on first step:
RHS_input = y_n
else: # If on second step or later:
RHS_input = next_y_input
RHS_output = sp.Symbol("k" + str(s + 1) + "_gfs", real=True)
if s == num_steps - 1: # If on final step:
RK_lhs = y_n
else: # If on anything but the final step:
RK_lhs = next_y_input
RK_rhs = y_n
for m in range(s + 1):
k_mp1_gfs = sp.Symbol("k" + str(m + 1) + "_gfsL")
if Butcher[s + 1][m + 1] != 0:
if Butcher[s + 1][m + 1] != 1:
RK_rhs += dt * k_mp1_gfs*Butcher[s + 1][m + 1]
else:
RK_rhs += dt * k_mp1_gfs
post_RHS = post_RHS_string
if s == num_steps - 1: # If on final step:
post_RHS_output = y_n
else: # If on anything but the final step:
post_RHS_output = next_y_input
body += single_RK_substep_input_symbolic(
commentblock="// -={ START k" + str(s + 1) + " substep }=-",
RHS_str=RHS_string,
RHS_input_str=RHS_input, RHS_output_str=RHS_output,
RK_lhss_list=[RK_lhs], RK_rhss_list=[RK_rhs],
post_RHS_list=[post_RHS],
post_RHS_output_list=[post_RHS_output],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string) + "// -={ END k" + str(s + 1) + " substep }=-\n\n"
else:
y_n = sp.Symbol("y_n_gfsL", real=True)
y_nplus1_running_total = sp.Symbol("y_nplus1_running_total_gfsL", real=True)
if MoL_method == 'Euler': # Euler's method doesn't require any k_i, and gets its own unique algorithm
body += single_RK_substep_input_symbolic(
commentblock=indent + "// ***Euler timestepping only requires one RHS evaluation***",
RHS_str=RHS_string,
RHS_input_str=y_n, RHS_output_str=y_nplus1_running_total,
RK_lhss_list=[y_n], RK_rhss_list=[y_n + y_nplus1_running_total*dt],
post_RHS_list=[post_RHS_string],
post_RHS_output_list=[y_n],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string)
else:
for s in range(num_steps):
# If we're on the first step (s=0), we use y_n gridfunction as input.
# and k_odd as output.
if s == 0:
RHS_input = sp.Symbol("y_n_gfsL", real=True)
RHS_output = sp.Symbol("k_odd_gfsL", real=True)
# For the remaining steps the inputs and ouputs alternate between k_odd and k_even
elif s % 2 == 0:
RHS_input = sp.Symbol("k_even_gfsL", real=True)
RHS_output = sp.Symbol("k_odd_gfsL", real=True)
else:
RHS_input = sp.Symbol("k_odd_gfsL", real=True)
RHS_output = sp.Symbol("k_even_gfsL", real=True)
RK_lhs_list = []
RK_rhs_list = []
if s != num_steps-1: # For anything besides the final step
if s == 0: # The first RK step
RK_lhs_list.append(y_nplus1_running_total)
RK_rhs_list.append(RHS_output*dt*Butcher[num_steps][s+1])
RK_lhs_list.append(RHS_output)
RK_rhs_list.append(y_n + RHS_output*dt*Butcher[s+1][s+1])
else:
if Butcher[num_steps][s+1] != 0:
RK_lhs_list.append(y_nplus1_running_total)
if Butcher[num_steps][s+1] != 1:
RK_rhs_list.append(y_nplus1_running_total + RHS_output*dt*Butcher[num_steps][s+1])
else:
RK_rhs_list.append(y_nplus1_running_total + RHS_output*dt)
if Butcher[s+1][s+1] != 0:
RK_lhs_list.append(RHS_output)
if Butcher[s+1][s+1] != 1:
RK_rhs_list.append(y_n + RHS_output*dt*Butcher[s+1][s+1])
else:
RK_rhs_list.append(y_n + RHS_output*dt)
post_RHS_output = RHS_output
if s == num_steps-1: # If on the final step
if Butcher[num_steps][s+1] != 0:
RK_lhs_list.append(y_n)
if Butcher[num_steps][s+1] != 1:
RK_rhs_list.append(y_n + y_nplus1_running_total + RHS_output*dt*Butcher[num_steps][s+1])
else:
RK_rhs_list.append(y_n + y_nplus1_running_total + RHS_output*dt)
post_RHS_output = y_n
body += single_RK_substep_input_symbolic(
commentblock=indent + "// -={ START k" + str(s + 1) + " substep }=-",
RHS_str=RHS_string,
RHS_input_str=RHS_input, RHS_output_str=RHS_output,
RK_lhss_list=RK_lhs_list, RK_rhss_list=RK_rhs_list,
post_RHS_list=[post_RHS_string],
post_RHS_output_list=[post_RHS_output],
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata, gf_aliases=gf_aliases,
post_post_RHS_string=post_post_RHS_string) + "// -={ END k" + str(s + 1) + " substep }=-\n\n"
enableCparameters=True
if enable_griddata:
enableCparameters=False
add_to_Cfunction_dict(
includes=includes,
desc=desc,
c_type=c_type, name=name, params=params,
body=indent_Ccode(body, " "),
enableCparameters=enableCparameters, rel_path_to_Cparams=os.path.join("."))
# <a id='free'></a>
#
# ## Step 3.d: Memory deallocation: `MoL_free_memory()` [Back to [top](#toc)\]
# $$\label{free}$$
#
# We define the function `MoL_free_memory()` which generates the C code for freeing the memory that was being occupied by the grid functions lists that had been allocated.
# add_to_Cfunction_dict_MoL_free_memory() registers
# MoL_free_memory_y_n_gfs() and
# MoL_free_memory_non_y_n_gfs(), which free memory for
# the indicated sets of gridfunctions
def add_to_Cfunction_dict_MoL_free_memory(MoL_method, which_gfs):
includes = ["NRPy_basic_defines.h", "NRPy_function_prototypes.h"]
desc = "Method of Lines (MoL) for \"" + MoL_method + "\" method: Free memory for \"" + which_gfs + "\" gridfunctions\n"
desc += " - y_n_gfs are used to store data for the vector of gridfunctions y_i at t_n, at the start of each MoL timestep\n"
desc += " - non_y_n_gfs are needed for intermediate (e.g., k_i) storage in chosen MoL method\n"
c_type = "void"
y_n_gridfunctions, non_y_n_gridfunctions_list, diagnostic_gridfunctions_point_to = \
generate_gridfunction_names(MoL_method=MoL_method)
gridfunctions_list = []
if which_gfs == "y_n_gfs":
gridfunctions_list = [y_n_gridfunctions]
elif which_gfs == "non_y_n_gfs":
gridfunctions_list = non_y_n_gridfunctions_list
else:
print("ERROR: which_gfs = \"" + which_gfs + "\" unrecognized.")
sys.exit(1)
name = "MoL_free_memory_" + which_gfs
params = "const paramstruct *restrict params, MoL_gridfunctions_struct *restrict gridfuncs"
body = ""
for gridfunctions in gridfunctions_list:
body += " free(gridfuncs->" + gridfunctions + ");\n"
add_to_Cfunction_dict(
includes=includes,
desc=desc,
c_type=c_type, name=name, params=params,
body=indent_Ccode(body, " "),
rel_path_to_Cparams=os.path.join("."))
# <a id='nrpybasicdefines'></a>
#
# ## Step 3.e: Define & register `MoL_gridfunctions_struct` in `NRPy_basic_defines.h`: `NRPy_basic_defines_MoL_timestepping_struct()` [Back to [top](#toc)\]
# $$\label{nrpybasicdefines}$$
#
# `MoL_gridfunctions_struct` stores pointers to all the gridfunctions needed by MoL, and we define this struct within `NRPy_basic_defines.h`.
# Register MoL_gridfunctions_struct in NRPy_basic_defines
def NRPy_basic_defines_MoL_timestepping_struct(MoL_method="RK4"):
y_n_gridfunctions, non_y_n_gridfunctions_list, diagnostic_gridfunctions_point_to = \
generate_gridfunction_names(MoL_method=MoL_method)
# Step 3.b: Create MoL_timestepping struct:
indent = " "
Nbd = "typedef struct __MoL_gridfunctions_struct__ {\n"
Nbd += indent + "REAL *restrict " + y_n_gridfunctions + ";\n"
for gfs in non_y_n_gridfunctions_list:
Nbd += indent + "REAL *restrict " + gfs + ";\n"
Nbd += indent + "REAL *restrict diagnostic_output_gfs;\n"
Nbd += "} MoL_gridfunctions_struct;\n"
Nbd += """#define LOOP_ALL_GFS_GPS(ii) _Pragma("omp parallel for") \\
for(int (ii)=0;(ii)<Nxx_plus_2NGHOSTS0*Nxx_plus_2NGHOSTS1*Nxx_plus_2NGHOSTS2*NUM_EVOL_GFS;(ii)++)\n"""
outC_NRPy_basic_defines_h_dict["MoL"] = Nbd
# <a id='setupall'></a>
#
# ## Step 3.f: Add all MoL C codes to C function dictionary, and add MoL definitions to `NRPy_basic_defines.h`: `register_C_functions_and_NRPy_basic_defines()` \[Back to [top](#toc)\]
# $$\label{setupall}$$
# Finally declare the master registration function
def register_C_functions_and_NRPy_basic_defines(MoL_method = "RK4",
RHS_string = "rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
post_RHS_string = "apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);", post_post_RHS_string = "",
enable_rfm=False, enable_curviBCs=False, enable_SIMD=False, enable_griddata=False):
for which_gfs in ["y_n_gfs", "non_y_n_gfs"]:
add_to_Cfunction_dict_MoL_malloc(MoL_method, which_gfs)
add_to_Cfunction_dict_MoL_free_memory(MoL_method, which_gfs)
add_to_Cfunction_dict_MoL_step_forward_in_time(MoL_method, RHS_string, post_RHS_string, post_post_RHS_string,
enable_rfm=enable_rfm, enable_curviBCs=enable_curviBCs,
enable_SIMD=enable_SIMD, enable_griddata=enable_griddata)
NRPy_basic_defines_MoL_timestepping_struct(MoL_method=MoL_method)
# <a id='code_validation'></a>
#
# # Step 4: Code Validation against `MoLtimestepping.MoL_new_way` NRPy+ module [Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# As a code validation check, we verify agreement in the dictionary of Butcher tables between
#
# 1. this tutorial and
# 2. the NRPy+ [MoLtimestepping.MoL_new_way](../edit/MoLtimestepping/MoL_new_way.py) module.
#
# We generate the header files for each RK method and check for agreement with the NRPy+ module.
# +
import sys
import MoLtimestepping.MoL_new_way as MoLC
import difflib
import pprint
# Courtesy https://stackoverflow.com/questions/12956957/print-diff-of-python-dictionaries ,
# which itself is an adaptation of some Cpython core code
def compare_dicts(d1, d2):
return ('\n' + '\n'.join(difflib.ndiff(
pprint.pformat(d1).splitlines(),
pprint.pformat(d2).splitlines())))
print("\n\n ### BEGIN VALIDATION TESTS ###")
import filecmp
for key, value in Butcher_dict.items():
register_C_functions_and_NRPy_basic_defines(key,
"rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
"apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);")
from outputC import outC_function_dict, outC_function_master_list
notebook_dict = outC_function_dict.copy()
notebook_master_list = list(outC_function_master_list)
outC_function_dict.clear()
del outC_function_master_list[:]
from outputC import outC_function_dict
if outC_function_dict != {}:
print("Error in clearing outC_function_dict.")
sys.exit(1)
MoLC.register_C_functions_and_NRPy_basic_defines(key,
"rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
"apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);")
from outputC import outC_function_dict
python_module_dict = outC_function_dict
if notebook_dict != python_module_dict:
print("VALIDATION TEST FAILED.\n")
print(compare_dicts(notebook_dict, python_module_dict))
sys.exit(1)
print("VALIDATION TEST PASSED on all files from "+str(key)+" method")
print("### END VALIDATION TESTS ###")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 5: Output this notebook to $\LaTeX$-formatted PDF \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-RK_Butcher_Table_Generating_C_Code.pdf](Tutorial-RK_Butcher_Table_Generating_C_Code.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-Method_of_Lines-C_Code_Generation_new_way")
|
Tutorial-Method_of_Lines-C_Code_Generation_new_way.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <br><br><br><br><br><h1 style="font-size:4em;color:#2467C0">Welcome to Week 3</h1><br><br><br>
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">This document provides a running example of completing the Week 3 assignment : </p>
# <ul class="simple">
# <li style="line-height:31px;">A shorter version with fewer comments is available as script: sparkMLlibClustering.py</li>
# <li style="line-height:31px;">To run these commands in Cloudera VM: first run the setup script: setupWeek3.sh</li>
# <li style="line-height:31px;">You can then copy paste these commands in pySpark. </li>
# <li style="line-height:31px;">To open pySpark, refer to : <a class="reference external" href="https://www.coursera.org/learn/machinelearningwithbigdata/supplement/GTFQ0/slides-module-2-lesson-3">Week 2</a> and <a class="reference external" href="https://www.coursera.org/learn/machinelearningwithbigdata/supplement/RH1zz/download-lesson-2-slides-spark-mllib-clustering">Week 4</a> of the Machine Learning course</li>
# <li style="line-height:31px;">Note that your dataset may be different from what is used here, so your results may not match with those shown here</li>
# </ul></div>
import pandas as pd
from pyspark.mllib.clustering import KMeans, KMeansModel
from numpy import array
# <br><br>
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
# <h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Step 1: Attribute Selection</h1>
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Import Data</h1><br><br>
#
#
# <p style="line-height:31px;">First let us read the contents of the file ad-clicks.csv. The following commands read in the CSV file in a table format and removes any extra whitespaces. So, if the CSV contained ' userid ' it becomes 'userid'. <br><br>
#
#
# Note that you must change the path to ad-clicks.csv to the location on your machine, if you want to run this command on your machine.
# </p>
#
# </div>
#
# <br><br><br><br>
adclicksDF = pd.read_csv('./ad-clicks.csv')
adclicksDF = adclicksDF.rename(columns=lambda x: x.strip()) #remove whitespaces from headers
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Let us display the first 5 lines of adclicksDF:</p>
#
# </div>
#
# <br><br><br><br>
adclicksDF.head(n=5)
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Next, We are going to add an extra column to the ad-clicks table and make it equal to 1. We do so to record the fact that each ROW is 1 ad-click.
# You will see how this will become useful when we sum up this column to find how many ads
# did a user click.</p>
#
# </div>
#
# <br><br><br><br>
adclicksDF['adCount'] = 1
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Let us display the first 5 lines of adclicksDF and see if
# a new column has been added:</p>
#
# </div>
#
# <br><br><br><br>
adclicksDF.head(n=5)
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Next, let us read the contents of the file buy-clicks.csv. As before, the following commands read in the CSV file in a table format and removes any extra whitespaces. So, if the CSV contained ' userid ' it becomes 'userid'. <br><br>
#
#
# Note that you must change the path to buy-clicks.csv to the location on your machine, if you want to run this command on your machine.
# </p>
#
# </div>
#
# <br><br><br><br>
buyclicksDF = pd.read_csv('./buy-clicks.csv')
buyclicksDF = buyclicksDF.rename(columns=lambda x: x.strip()) #removes whitespaces from headers
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Let us display the first 5 lines of buyclicksDF:</p>
#
# </div>
#
# <br><br><br><br>
buyclicksDF.head(n=5)
# <br><br>
#
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Feature Selection</h1><br><br>
#
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">For this exercise, we can choose from buyclicksDF, the 'price' of each app that a user purchases as an attribute that captures user's purchasing behavior. The following command selects 'userid' and 'price' and drops all other columns that we do not want to use at this stage.</p>
#
#
# </div>
#
# <br><br><br><br>
userPurchases = buyclicksDF[['userId','price']] #select only userid and price
userPurchases.head(n=5)
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Similarly, from the adclicksDF, we will use the 'adCount' as an attribute that captures user's inclination to click on ads. The following command selects 'userid' and 'adCount' and drops all other columns that we do not want to use at this stage.</p>
#
#
# </div>
#
# <br><br><br><br>
useradClicks = adclicksDF[['userId','adCount']]
useradClicks.head(n=5) #as we saw before, this line displays first five lines
# <br><br>
# <h1 style="font-family: Arial; font-size:1.5em;color:#2462C0; font-style:bold">Step 2: Training Data Set Creation</h1>
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Create the first aggregate feature for clustering</h1><br><br>
#
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">From each of these single ad-clicks per row, we can now generate total ad clicks per user. Let's pick a user with userid = 3. To find out how many ads this user has clicked overall, we have to find each row that contains userid = 3, and report the total number of such rows.
#
# The following commands sum the total number of ads per user and rename the columns to be called 'userid' and 'totalAdClicks'. <b> Note that you may not need to aggregate (e.g. sum over many rows) if you choose a different feature and your data set already provides the necessary information. </b> In the end, we want to get one row per user, if we are performing clustering over users.
#
# </div>
#
# <br><br><br><br>
adsPerUser = useradClicks.groupby('userId').sum()
adsPerUser = adsPerUser.reset_index()
adsPerUser.columns = ['userId', 'totalAdClicks'] #rename the columns
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Let us display the first 5 lines of 'adsPerUser' to see if there
# is a column named 'totalAdClicks' containing total adclicks per user.</p>
#
# </div>
#
# <br><br><br><br>
adsPerUser.head(n=5)
# <br><br>
#
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Create the second aggregate feature for clustering</h1><br><br>
#
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Similar to what we did for adclicks, here we find out how much money in total did each user spend on buying in-app purchases. As an example, let's pick a user with userid = 9. To find out the total money spent by this user, we have to find each row that contains userid = 9, and report the sum of the column'price' of each product they purchased.
#
# The following commands sum the total money spent by each user and rename the columns to be called 'userid' and 'revenue'.
# <br><br>
#
# <p style="line-height:31px;"> <b> Note: </b> that you can also use other aggregates, such as sum of money spent on a specific ad category by a user or on a set of ad categories by each user, game clicks per hour by each user etc. You are free to use any mathematical operations on the fields provided in the CSV files when creating features. </p>
#
#
# </div>
#
# <br><br><br><br>
revenuePerUser = userPurchases.groupby('userId').sum()
revenuePerUser = revenuePerUser.reset_index()
revenuePerUser.columns = ['userId', 'revenue'] #rename the columns
revenuePerUser.head(n=5)
# <br><br>
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Merge the two tables</h1><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Lets see what we have so far. We have a table called revenuePerUser, where each row contains total money a user (with that 'userid') has spent. We also have another table called adsPerUser where each row contains total number of ads a user has clicked. We will use revenuePerUser and adsPerUser as features / attributes to capture our users' behavior.<br><br>
#
# Let us combine these two attributes (features) so that each row contains both attributes per user. Let's merge these two tables to get one single table we can use for K-Means clustering.
# </div>
#
# <br><br><br><br>
combinedDF = adsPerUser.merge(revenuePerUser, on='userId') #userId, adCount, price
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
#
# <p style="line-height:31px;">Let us display the first 5 lines of the merged table. <b> Note: Depending on what attributes you choose, you may not need to merge tables. You may get all your attributes from a single table. </b></p>
#
# </div>
#
# <br><br><br><br>
combinedDF.head(n=5) #display how the merged table looks
# <br><br>
#
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Create the final training dataset</h1><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Our training data set is almost ready. At this stage we can remove the 'userid' from each row, since 'userid' is a computer generated random number assigned to each user. It does not capture any behavioral aspect of a user. One way to drop the 'userid', is to select the other two columns. </p>
#
# </div>
#
# <br><br><br><br>
trainingDF = combinedDF[['totalAdClicks','revenue']]
trainingDF.head(n=5)
# <br><br>
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Display the dimensions of the training dataset</h1><br><br>
#
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Display the dimension of the training data set. To display the dimensions of the trainingDF, simply add .shape as a suffix and hit enter.</p>
#
# </div>
#
# <br><br><br><br>
#
trainingDF.shape
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">The following two commands convert the tables we created into a format that can be understood by the KMeans.train function. <br><br>
#
# line[0] refers to the first column. line[1] refers to the second column. If you have more than 2 columns in your training table, modify this command by adding line[2], line[3], line[4] ...</p>
#
# </div>
#
# <br><br><br><br>
#
sqlContext = SQLContext(sc)
pDF = sqlContext.createDataFrame(trainingDF)
parsedData = pDF.rdd.map(lambda line: array([line[0], line[1]])) #totalAdClicks, revenue
# <br>
# <h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Step 3: Train to Create Cluster Centers</h1>
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Train KMeans model</h1><br><br>
# <br><br><br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Here we are creating two clusters as denoted in the second argument.</p>
#
# </div>
#
# <br><br><br><br>
#
my_kmmodel = KMeans.train(parsedData, 2, maxIterations=10, runs=10, initializationMode="random")
# <br><br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Display the centers of two clusters formed</h1><br><br>
print(my_kmmodel.centers)
# <br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Step 4: Recommend Actions</h1>
# <br><h1 style="font-family: Arial; font-size:1.5em;color:#2462C0">Analyze the cluster centers
# </h1>
#
# <br><br>
#
# <div style="color:black;font-family: Arial; font-size:1.1em;line-height:65%">
#
# <p style="line-height:31px;">Each array denotes the center for a cluster:<br><br>
# One Cluster is centered at ... array([ 29.43211679, 24.21021898])<br>
# Other Cluster is centered at ... array([ 42.05442177, 113.02040816])</p>
#
# <br><br>
#
# <p style="line-height:31px;"> First number (field1) in each array refers to number of ad-clicks and the second number (field2) is the revenue per user.
#
# Compare the 1st number of each cluster to see how differently users in each cluster behave when it comes to clicking ads.
#
# Compare the 2nd number of each cluster to see how differently users in each cluster behave when it comes to buying stuff.
#
# </p><br><br>
#
# <p style="line-height:31px;">In one cluster, in general, players click on ads much more often (~1.4 times) and spend more money (~4.7 times) on in-app purchases. Assuming that Eglence Inc. gets paid for showing ads and for hosting in-app purchase items, we can use this information to increase game's revenue by increasing the prices for ads we show to the frequent-clickers, and charge higher fees for hosting the in-app purchase items shown to the higher revenue generating buyers.</p>
#
# <br><br>
# <p style="line-height:31px;"> <b> Note: </b> This analysis requires you to compare the cluster centers and find any ‘significant’ differences in the corresponding feature values of the centers. The answer to this question will depend on the features you have chosen. <br><br> Some features help distinguish the clusters remarkably while others may not tell you much. At this point, if you don’t find clear distinguishing patterns, perhaps re-running the clustering model with different numbers of clusters and revising the features you picked would be a good idea. </p>
#
# </div>
#
|
Week 3/Week 3 pySpark MLlib Clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Understanding the SVD
import numpy as np
# ### Useful reference
#
# - [A Singularly Valuable Decomposition](https://datajobs.com/data-science-repo/SVD-[Dan-Kalman].pdf)
# ## Sketch of lecture
# ### Singular value decomposition
#
# Our goal is to understand the following forms of the SVD.
#
# $$
# A = U \Sigma V^T
# $$
#
# $$
# A = \begin{bmatrix}
# U_1 & U_2
# \end{bmatrix}\begin{bmatrix}
# \Sigma_1 & 0 \\
# 0 & 0
# \end{bmatrix}\begin{bmatrix}
# V_1^T \\
# V_2^T
# \end{bmatrix}
# $$
#
# $$
# A = \sum_{i=1}^r \sigma u_i v_i^T
# $$
# ### (1) The matrix A
# #### What does a matrix do?
#
# A linear function is one that satisfies the property that
#
# $$
# f(a_1x_1 + a_2x_2 + \cdots + a_nx_n) = a_1 f(x_1) + a_2 f(x_2) + \ldots + a_n f(x_n)
# $$
#
# Let $f(x) = Ax$, where $A$ is a matrix and $x$ is a vector. You can check that the matrix $A$ fulfills the property of being a linear function. If $A$ is $m \times n$, then it is a linear map from $\mathbb{R}^n \mapsto \mathbb{R}^m$.
#
# Let's consider: what does a matrix *do* to a vector? Matrix multiplication has a *geometric* interpretation. When we multiply a vector, we either rotate, reflect, dilate or some combination of those three. So multiplying by a matrix *transforms* one vector into another vector. This is known as a *linear transformation*.
#
# Important Facts:
#
# * Any matrix defines a linear transformation
# * The matrix form of a linear transformation is NOT unique
# * We need only define a transformation by saying what it does to a *basis*
#
# Suppose we have a matrix $A$ that defines some transformation. We can take any invertible matrix $B$ and
#
# $$BAB^{-1}$$
#
# defines the same transformation. This operation is called a *change of basis*, because we are simply expressing the transformation with respect to a different basis.
# **Example**
#
# Let $f(x)$ be the linear transformation that takes $e_1=(1,0)$ to $f(e_1)=(2,3)$ and $e_2=(0,1)$ to $f(e_2) = (1,1)$. A matrix representation of $f$ would be given by:
#
# $$A = \left(\begin{matrix}2 & 1\\3&1\end{matrix}\right)$$
#
# This is the matrix we use if we consider the vectors of $\mathbb{R}^2$ to be linear combinations of the form
#
# $$c_1 e_1 + c_2 e_2$$
#
# Now, consider a second pair of (linearly independent) vectors in $\mathbb{R}^2$, say $v_1=(1,3)$ and $v_2=(4,1)$. We first find the transformation that takes $e_1$ to $v_1$ and $e_2$ to $v_2$. A matrix representation for this is:
#
# $$B = \left(\begin{matrix}1 & 4\\3&1\end{matrix}\right)$$
#
# Our original transformation $f$ can be expressed with respect to the basis $v_1, v_2$ via
#
# $$B^{-1}AB$$
# #### Fundamental subspaces of $A$
#
# - Span and basis
# - Inner and outer products of vectors
# - Rank of outer product is 1
# - $C(A)$, $N(A)$, $(C(A^T))$ and $N(A^T)$ mean
# - Dimensions of each space and its rank
# - How to find a basis for each subspace given a $m \times n$ matrix $A$
# - Sketch the diagram relating the four fundamental subspaces
# ### (2) Orthogonal matrices $U$ and $V^T$
# - Orthogonal (perpendicular) vectors
# - Orthonormal vectors
# - Orthogonal matrix
# - $Q^TQ = QQ^T = I$
# - Orthogonal matrices are rotations (and reflections)
# - Orthogonal matrices preserve norms (lengths)
# - 2D orthogonal matrix is a rotation matrix
# $$ V =
# \begin{bmatrix}
# \cos\theta & -\sin \theta \\
# \sin \theta & \cos \theta
# \end{bmatrix}
# $$
# - $V^T$ rotates the perpendicular frame spanned by $V$ into the standard frame spanned by $e_i$
# - $V$ rotates the standard frame into the frame spanned by $V$
# -
# $$\text{proj}_v x = \frac{\langle x, v \rangle}{\langle v, v \rangle} v
# $$
# - Matrix form
# $$
# P = \frac{vv^T}{v^Tv}
# $$
# - Gram-Schmidt for converting $A$ into an orthogonal matrix $Q$
# - QR decomposition
# ### (3) Diagonal matrix $S$
# - Recall that a matrix $A$ is a transform with respect to some basis
# - It is desirable to find the simplest similar matrix $B$ in some other basis
# - $A$ and $B$ represent the exact same linear transform, just in different coordinate systems
# - $Av = \lambda v$ defines the eigenvectors and eigenvalues of $A$
# - When a square matrix $A$ is real, symmetric and has all non-negative eigenvalues, it has an eigen-space decomposition (ESD)
# $$
# A = V \Lambda V^T
# $$
# where $V$ is orthogonal and $\Lambda$ is diagonal
# - The columns of $V$ are formed from the eigenvectors of $A$
# - The diagonals of $\Lambda$ are the eigenvalues of $A$ (arrange from large to small in absolute value)
# ## (4) SVD $U\Sigma V^T$
# - The SVD is a generalization of ESD for general $m \times n$ matrices $A$
# - If $A$ is $(m \times n)$, we cannot perform an ESD
# - $A^TA$ is diagonalizable (note this is the dot product of all pairs of column vectors in $A$)
# -
# $$
# A^TA = V \Lambda V^T
# $$
# - Let $\Lambda = \Sigma^2$
# - Let $U = AV\Sigma^{-1}$
# - The $A = U\Sigma V^T$
# - Show $U$ is orthogonal
# - Show $U$ is formed from eigenvectors of $AA^T$
# - Geometric interpretation of SVD
# - rotate orthogonal frame $V$ onto standard frame
# - scale by $\Sigma$
# - rotate standard frame into orthogonal frame $U$
# ### Covariance, PCA and SVD
# Remember the formula for covariance
#
# $$
# \text{Cov}(X, Y) = \frac{\sum_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})}{n-1}
# $$
#
# where $\text{Cov}(X, X)$ is the sample variance of $X$.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import scipy.linalg as la
np.set_printoptions(precision=3)
def cov(x, y):
"""Returns covariance of vectors x and y)."""
xbar = x.mean()
ybar = y.mean()
return np.sum((x - xbar)*(y - ybar))/(len(x) - 1)
X = np.random.random(10)
Y = np.random.random(10)
np.array([[cov(X, X), cov(X, Y)], [cov(Y, X), cov(Y,Y)]])
# Using `numpy` function
np.cov(X, Y)
Z = np.random.random(10)
np.cov([X, Y, Z])
# #### Eigendecomposition of the covariance matrix
mu = [0,0]
sigma = [[0.6,0.2],[0.2,0.2]]
n = 1000
x = np.random.multivariate_normal(mu, sigma, n).T
A = np.cov(x)
m = np.array([[1,2,3],[6,5,4]])
ms = m - m.mean(1).reshape(2,1)
np.dot(ms, ms.T)/2
e, v = la.eigh(A)
plt.scatter(x[0,:], x[1,:], alpha=0.2)
for e_, v_ in zip(e, v.T):
plt.plot([0, 3*e_*v_[0]], [0, 3*e_*v_[1]], 'r-', lw=2)
plt.axis([-3,3,-3,3])
plt.title('Eigenvectors of covariance matrix scaled by eigenvalue.');
# ### PCA
#
# Principal Components Analysis (PCA) basically means to find and rank all the eigenvalues and eigenvectors of a covariance matrix. This is useful because high-dimensional data (with $p$ features) may have nearly all their variation in a small number of dimensions $k$, i.e. in the subspace spanned by the eigenvectors of the covariance matrix that have the $k$ largest eigenvalues. If we project the original data into this subspace, we can have a dimension reduction (from $p$ to $k$) with hopefully little loss of information.
#
# Numerically, PCA is typically done using SVD on the data matrix rather than eigendecomposition on the covariance matrix. The next section explains why this works. Numerically, the condition number for working with the covariance matrix directly is the square of the condition number using SVD, so SVD minimizes errors.
# For zero-centered vectors,
#
# \begin{align}
# \text{Cov}(X, Y) &= \frac{\sum_{i=1}^n(X_i - \bar{X})(Y_i - \bar{Y})}{n-1} \\
# &= \frac{\sum_{i=1}^nX_iY_i}{n-1} \\
# &= \frac{XY^T}{n-1}
# \end{align}
#
# and so the covariance matrix for a data set X that has zero mean in each feature vector is just $XX^T/(n-1)$.
#
# In other words, we can also get the eigendecomposition of the covariance matrix from the positive semi-definite matrix $XX^T$.
# Note: Here $x$ is a matrix of **row** vectors
X = np.random.random((5,4))
X
Y = X - X.mean(1)[:, None]
np.around(Y.mean(1), 5)
Y
np.cov(X)
np.cov(Y)
e1, v1 = np.linalg.eig(np.dot(x, x.T)/(n-1))
# #### Principal components
#
# Principal components are simply the eigenvectors of the covariance matrix used as basis vectors. Each of the original data points is expressed as a linear combination of the principal components, giving rise to a new set of coordinates.
plt.scatter(x[0,:], x[1,:], alpha=0.2)
for e_, v_ in zip(e1, v1.T):
plt.plot([0, 3*e_*v_[0]], [0, 3*e_*v_[1]], 'r-', lw=2)
plt.axis([-3,3,-3,3]);
# #### Change of basis
#
# Suppose we have a vector $u$ in the standard basis $B$ , and a matrix $A$ that maps $u$ to $v$, also in $B$. We can use the eigenvalues of $A$ to form a new basis $B'$. As explained above, to bring a vector $u$ from $B$-space to a vector $u'$ in $B'$-space, we multiply it by $Q^{-1}$, the inverse of the matrix having the eigenvctors as column vectors. Now, in the eigenvector basis, the equivalent operation to $A$ is the diagonal matrix $\Lambda$ - this takes $u'$ to $v'$. Finally, we convert $v'$ back to a vector $v$ in the standard basis by multiplying with $Q$.
#
# 
# #### Rotate the standard frame
#
# Principal components are simply the eigenvectors of the covariance matrix used as basis vectors. Each of the original data points is expressed as a linear combination of the principal components, giving rise to a new set of coordinates.
#
#
ys = np.dot(v1.T, x)
# #### We get the principal components by a change of basis
plt.scatter(ys[0,:], ys[1,:], alpha=0.2)
for e_, v_ in zip(e1, np.eye(2)):
plt.plot([0, 3*e_*v_[0]], [0, 3*e_*v_[1]], 'r-', lw=2)
plt.axis([-3,3,-3,3]);
# For example, if we only use the first column of `ys`, we will have the projection of the data onto the first principal component, capturing the majority of the variance in the data with a single feature that is a linear combination of the original features.
# #### Transform back to original coordinates
#
# We may need to transform the (reduced) data set to the original feature coordinates for interpretation. This is simply another linear transform (matrix multiplication).
zs = np.dot(v1, ys)
plt.scatter(zs[0,:], zs[1,:], alpha=0.2)
for e_, v_ in zip(e1, v1.T):
plt.plot([0, 3*e_*v_[0]], [0, 3*e_*v_[1]], 'r-', lw=2)
plt.axis([-3,3,-3,3]);
u, s, v = np.linalg.svd(x)
u.dot(u.T)
# #### Dimension reduction via PCA
#
# We have the spectral decomposition of the covariance matrix
#
# $$
# A = Q^{-1}\Lambda Q
# $$
#
# Suppose $\Lambda$ is a rank $p$ matrix. To reduce the dimensionality to $k \le p$, we simply set all but the first $k$ values of the diagonal of $\Lambda$ to zero. This is equivalent to ignoring all except the first $k$ principal components.
#
# What does this achieve? Recall that $A$ is a covariance matrix, and the trace of the matrix is the overall variability, since it is the sum of the variances.
A
A.trace()
e, v = np.linalg.eigh(A)
D = np.diag(e)
D
D.trace()
D[0,0]/D.trace()
# Since the trace is invariant under change of basis, the total variability is also unchanged by PCA. By keeping only the first $k$ principal components, we can still "explain" $\sum_{i=1}^k e[i]/\sum{e}$ of the total variability. Sometimes, the degree of dimension reduction is specified as keeping enough principal components so that (say) $90\%$ of the total variability is explained.
# ### Using SVD for PCA
#
# SVD is a decomposition of the data matrix $X = U S V^T$ where $U$ and $V$ are orthogonal matrices and $S$ is a diagonal matrix.
#
# Recall that the transpose of an orthogonal matrix is also its inverse, so if we multiply on the right by $X^T$, we get the following simplification
#
# \begin{align}
# X &= U S V^T \\
# X X^T &= U S V^T (U S V^T)^T \\
# &= U S V^T V S U^T \\
# &= U S^2 U^T
# \end{align}
#
# Compare with the eigendecomposition of a matrix $A = W \Lambda W^{-1}$, we see that SVD gives us the eigendecomposition of the matrix $XX^T$, which as we have just seen, is basically a scaled version of the covariance for a data matrix with zero mean, with the eigenvectors given by $U$ and eigenvealuse by $S^2$ (scaled by $n-1$)..
u, s, v = np.linalg.svd(x)
e2 = s**2/(n-1)
v2 = u
plt.scatter(x[0,:], x[1,:], alpha=0.2)
for e_, v_ in zip(e2, v2):
plt.plot([0, 3*e_*v_[0]], [0, 3*e_*v_[1]], 'r-', lw=2)
plt.axis([-3,3,-3,3]);
v1 # from eigenvectors of covariance matrix
v2 # from SVD
e1 # from eigenvalues of covariance matrix
e2 # from SVD
|
notebooks/T04_SVD.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="qSFjR-V8p-k1"
# ## Importando bilioteca Bibliotecas
# + id="CvcevCfT5b3R"
# !pip install boto3
# + [markdown] id="jlcMUQnNqKfV"
# ## Acessando Nuvem AWS
# + id="jNRIJQrQ6CUs"
key= "###########"
access= "###########"
token= "#############"
# + [markdown] id="8sMjgo9qqYuW"
# ## Importando Biblioteca boto 3
# + id="y6D_O77I6RWj"
# a bilbioteca boto3 permite acessar os serviços da AWS
import boto3
# Acessando o Dynamodb via biblioteca boto3
dynamodb = boto3.resource('dynamodb', aws_access_key_id=key,
aws_secret_access_key=access,
aws_session_token=token
,region_name = "us-east-1", use_ssl=False)
# + id="KR6vAq2_6fD1"
# Criando uma tabela não relacional no Dynamodb.
table_name = 'funcionario' #Nome da tabela
#Parametros de criação
params = {
'TableName': table_name, #nome da tabela
#Esquema das chaves
'KeySchema': [
{'AttributeName': 'MATRICULA', 'KeyType': 'HASH'}, # hash será a chave primaria
{'AttributeName': 'VERSAO', 'KeyType': 'RANGE'} #range, não obrigatória, formaria uma chave composta
],
#Tipo de dados da chave
'AttributeDefinitions': [
{'AttributeName': 'MATRICULA', 'AttributeType': 'S'}, #S de string
{'AttributeName': 'VERSAO', 'AttributeType': 'N'} #N de number
],
#Configuração de escrita e leitura por segundo
'ProvisionedThroughput': {
'ReadCapacityUnits': 3,
'WriteCapacityUnits': 3
}
}
# + colab={"base_uri": "https://localhost:8080/"} id="tBZ99bBG8nCM" outputId="1eba23d2-fa1a-4019-f642-dcc3023f37f1"
dynamodb.create_table(**params)
# + colab={"base_uri": "https://localhost:8080/"} id="fi2lW2ML_7iU" outputId="bd445570-2928-4eb0-9124-5a1b3a4fa5b6"
dynamodb.Table(name='funcionario')
# + [markdown] id="99ednjeorQ7s"
# #### Inserindo valores manualmente.
# + colab={"base_uri": "https://localhost:8080/"} id="bk4YDD23_-7a" outputId="5aaa76d7-7518-427a-a1ab-c1295f302ec6"
table = dynamodb.Table("funcionario")
table.put_item(
Item={"MATRICULA":'1010',
'VERSAO':1,
'NOME': '<NAME>',
'CARGO': 'ANALISTA JUNIOR',
'TELEFONE': ["22381823","2291012"],
'DATA_ATUALIZAÇÃO': '202120101',
'TIPO SANGUINIO':'A',
'CNH': '24343455324'
})
# + colab={"base_uri": "https://localhost:8080/"} id="fuFkEaqKpGxn" outputId="913410f5-0e79-4a5c-b405-679aa3c58ac0"
table = dynamodb.Table("funcionario")
table.put_item(
Item={"MATRICULA":'1020',
'VERSAO':1,
'NOME': '<NAME>',
'CARGO': 'ENGENHEIRO',
'TELEFONE': ["953434343"],
'DATA_ATUALIZAÇÃO': '202120101',
'TIPO SANGUINIO':'',
'CNH': '1121323213'
})
# + colab={"base_uri": "https://localhost:8080/"} id="LyA6MOrVpZOT" outputId="0fa9728e-cfc3-46f0-a7a3-f1063273353a"
table = dynamodb.Table("funcionario")
table.put_item(
Item={"MATRICULA":'1030',
'VERSAO':1,
'NOME': '<NAME>',
'CARGO': 'TECNICO INFO',
'TELEFONE': ["953434344"],
'DATA_ATUALIZAÇÃO': '202120101',
'TIPO SANGUINIO':'O',
'CNH': '243434544653'
})
# + colab={"base_uri": "https://localhost:8080/"} id="C6VKdP6cpfaD" outputId="9203af6f-a06e-438b-830c-3e178a65f07d"
table = dynamodb.Table("funcionario")
table.put_item(
Item={"MATRICULA":'1040',
'VERSAO':1,
'NOME': '<NAME>',
'CARGO': 'ANALISTA SENIOR',
'TELEFONE': ["953434322"],
'DATA_ATUALIZAÇÃO': '202120101',
'TIPO SANGUINIO':'',
'CNH': ' '
})
# + colab={"base_uri": "https://localhost:8080/"} id="Tr7UHtDFpmI4" outputId="e23c855c-88dc-4377-b735-2292d3fd37ea"
table = dynamodb.Table("funcionario")
table.put_item(
Item={"MATRICULA":'1010',
'VERSAO':2,
'NOME': '<NAME>',
'CARGO': '<NAME>',
'TELEFONE': ["22381823","2291012"],
'DATA_ATUALIZAÇÃO': '202120101',
'TIPO SANGUINIO':'B',
'CNH': '343243232'
})
|
AWS_Dynamodb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deep Convolutional GANs
#
# In this notebook, you'll build a GAN using convolutional layers in the generator and discriminator. This is called a Deep Convolutional GAN, or DCGAN for short. The DCGAN architecture was first explored in 2016 and has seen impressive results in generating new images; you can read the [original paper, here](https://arxiv.org/pdf/1511.06434.pdf).
#
# You'll be training DCGAN on the [Street View House Numbers](http://ufldl.stanford.edu/housenumbers/) (SVHN) dataset. These are color images of house numbers collected from Google street view. SVHN images are in color and much more variable than MNIST.
#
# <img src='assets/svhn_dcgan.png' width=80% />
#
# So, our goal is to create a DCGAN that can generate new, realistic-looking images of house numbers. We'll go through the following steps to do this:
# * Load in and pre-process the house numbers dataset
# * Define discriminator and generator networks
# * Train these adversarial networks
# * Visualize the loss over time and some sample, generated images
#
# #### Deeper Convolutional Networks
#
# Since this dataset is more complex than our MNIST data, we'll need a deeper network to accurately identify patterns in these images and be able to generate new ones. Specifically, we'll use a series of convolutional or transpose convolutional layers in the discriminator and generator. It's also necessary to use batch normalization to get these convolutional networks to train.
#
# Besides these changes in network structure, training the discriminator and generator networks should be the same as before. That is, the discriminator will alternate training on real and fake (generated) images, and the generator will aim to trick the discriminator into thinking that its generated images are real!
# +
# import libraries
import matplotlib.pyplot as plt
import numpy as np
import pickle as pkl
# %matplotlib inline
# -
# ## Getting the data
#
# Here you can download the SVHN dataset. It's a dataset built-in to the PyTorch datasets library. We can load in training data, transform it into Tensor datatypes, then create dataloaders to batch our data into a desired size.
# +
import torch
from torchvision import datasets
from torchvision import transforms
# Tensor transform
transform = transforms.ToTensor()
# SVHN training datasets
svhn_train = datasets.SVHN(root='data/', split='train', download=True, transform=transform)
batch_size = 128 * 2
num_workers = 0
# build DataLoaders for SVHN dataset
train_loader = torch.utils.data.DataLoader(dataset=svhn_train,
batch_size=batch_size,
shuffle=True,
num_workers=num_workers)
# -
# ### Visualize the Data
#
# Here I'm showing a small sample of the images. Each of these is 32x32 with 3 color channels (RGB). These are the real, training images that we'll pass to the discriminator. Notice that each image has _one_ associated, numerical label.
# +
# obtain one batch of training images
dataiter = iter(train_loader)
images, labels = dataiter.next()
# plot the images in the batch, along with the corresponding labels
fig = plt.figure(figsize=(25, 4))
plot_size=20
for idx in np.arange(plot_size):
ax = fig.add_subplot(2, plot_size/2, idx+1, xticks=[], yticks=[])
ax.imshow(np.transpose(images[idx], (1, 2, 0)))
# print out the correct label for each image
# .item() gets the value contained in a Tensor
ax.set_title(str(labels[idx].item()))
# -
# ### Pre-processing: scaling from -1 to 1
#
# We need to do a bit of pre-processing; we know that the output of our `tanh` activated generator will contain pixel values in a range from -1 to 1, and so, we need to rescale our training images to a range of -1 to 1. (Right now, they are in a range from 0-1.)
# +
# current range
img = images[0]
print('Min: ', img.min())
print('Max: ', img.max())
# -
# helper scale function
def scale(x, feature_range=(-1, 1)):
''' Scale takes in an image x and returns that image, scaled
with a feature_range of pixel values from -1 to 1.
This function assumes that the input x is already scaled from 0-1.'''
# assume x is scaled to (0, 1)
# scale to feature_range and return scaled x
min = feature_range[0]
max = feature_range[1]
x = x * (max - min) + min
return x
# +
# scaled range
scaled_img = scale(img)
print('Scaled min: ', scaled_img.min())
print('Scaled max: ', scaled_img.max())
# -
# ---
# # Define the Model
#
# A GAN is comprised of two adversarial networks, a discriminator and a generator.
# ## Discriminator
#
# Here you'll build the discriminator. This is a convolutional classifier like you've built before, only without any maxpooling layers.
# * The inputs to the discriminator are 32x32x3 tensor images
# * You'll want a few convolutional, hidden layers
# * Then a fully connected layer for the output; as before, we want a sigmoid output, but we'll add that in the loss function, [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss), later
#
# <img src='assets/conv_discriminator.png' width=80%/>
#
# For the depths of the convolutional layers I suggest starting with 32 filters in the first layer, then double that depth as you add layers (to 64, 128, etc.). Note that in the DCGAN paper, they did all the downsampling using only strided convolutional layers with no maxpooling layers.
#
# You'll also want to use batch normalization with [nn.BatchNorm2d](https://pytorch.org/docs/stable/nn.html#batchnorm2d) on each layer **except** the first convolutional layer and final, linear output layer.
#
# #### Helper `conv` function
#
# In general, each layer should look something like convolution > batch norm > leaky ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a convolutional + an optional batch norm layer. We'll create these using PyTorch's [Sequential container](https://pytorch.org/docs/stable/nn.html#sequential), which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
#
# Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for strided convolutions.
# +
import torch.nn as nn
import torch.nn.functional as F
# helper conv function
def conv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a convolutional layer, with optional batch normalization.
"""
layers = []
conv_layer = nn.Conv2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append conv layer
layers.append(conv_layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
# using Sequential container
return nn.Sequential(*layers)
# -
class Discriminator(nn.Module):
def __init__(self, conv_dim=32):
super(Discriminator, self).__init__()
# complete init function
in_channels = 3
out_channels = conv_dim
self.conv1 = conv(3, out_channels, 4, batch_norm = False) # 32 x 32 x 3 -> 16 x 16 x 32
in_channels = out_channels
out_channels *= 2
self.conv2 = conv(in_channels, out_channels, 4) # 16 x 16 x 32 -> 8 x 8 x 64
in_channels = out_channels
out_channels *= 2
self.conv3 = conv(in_channels, out_channels, 4) # 8 x 8 x 64 -> 4 x 4 x 128
self.fc = nn.Linear(4 * 4 * out_channels, 1)
self.leaky_relu = nn.LeakyReLU(negative_slope=0.2)
self.final_out_channels = out_channels
def forward(self, x):
# complete forward function
x = self.conv1(x)
x = self.leaky_relu(x)
x = self.conv2(x)
x = self.leaky_relu(x)
x = self.conv3(x)
x = self.leaky_relu(x)
x = x.view(-1, 4 * 4 * self.final_out_channels)
x = self.fc(x)
return x
# ## Generator
#
# Next, you'll build the generator network. The input will be our noise vector `z`, as before. And, the output will be a $tanh$ output, but this time with size 32x32 which is the size of our SVHN images.
#
# <img src='assets/conv_generator.png' width=80% />
#
# What's new here is we'll use transpose convolutional layers to create our new images.
# * The first layer is a fully connected layer which is reshaped into a deep and narrow layer, something like 4x4x512.
# * Then, we use batch normalization and a leaky ReLU activation.
# * Next is a series of [transpose convolutional layers](https://pytorch.org/docs/stable/nn.html#convtranspose2d), where you typically halve the depth and double the width and height of the previous layer.
# * And, we'll apply batch normalization and ReLU to all but the last of these hidden layers. Where we will just apply a `tanh` activation.
#
# #### Helper `deconv` function
#
# For each of these layers, the general scheme is transpose convolution > batch norm > ReLU, and so we'll define a function to put these layers together. This function will create a sequential series of a transpose convolutional + an optional batch norm layer. We'll create these using PyTorch's Sequential container, which takes in a list of layers and creates layers according to the order that they are passed in to the Sequential constructor.
#
# Note: It is also suggested that you use a **kernel_size of 4** and a **stride of 2** for transpose convolutions.
# helper deconv function
def deconv(in_channels, out_channels, kernel_size, stride=2, padding=1, batch_norm=True):
"""Creates a transposed-convolutional layer, with optional batch normalization.
"""
layers = []
layer = nn.ConvTranspose2d(in_channels, out_channels,
kernel_size, stride, padding, bias=False)
# append conv layer
layers.append(layer)
if batch_norm:
# append batchnorm layer
layers.append(nn.BatchNorm2d(out_channels))
# using Sequential container
return nn.Sequential(*layers)
class Generator(nn.Module):
def __init__(self, z_size, conv_dim=32):
super(Generator, self).__init__()
# complete init function
self.initial_channels = conv_dim * 4
self.fc = nn.Linear(z_size, 4 * 4 * self.initial_channels)
in_channels = self.initial_channels
out_channels = int(in_channels / 2)
self.deconv1 = deconv(in_channels, out_channels, 4)
in_channels = out_channels
out_channels = int(in_channels / 2)
self.deconv2 = deconv(in_channels, out_channels, 4)
in_channels = out_channels
out_channels = 3
self.deconv3 = deconv(in_channels, out_channels, 4, batch_norm=False)
self.relu = nn.ReLU()
self.tanh = nn.Tanh()
def forward(self, x):
# complete forward function
x = self.fc(x)
x = x.view(-1, self.initial_channels, 4, 4)
x = self.deconv1(x)
x = self.relu(x)
x = self.deconv2(x)
x = self.relu(x)
x = self.deconv3(x)
x = self.tanh(x)
return x
# ## Build complete network
#
# Define your models' hyperparameters and instantiate the discriminator and generator from the classes defined above. Make sure you've passed in the correct input arguments.
# +
# define hyperparams
conv_dim = 128
z_size = 100
# define discriminator and generator
D = nn.DataParallel(Discriminator(conv_dim))
G = nn.DataParallel(Generator(z_size=z_size, conv_dim=conv_dim))
print(D)
print()
print(G)
# -
# ### Training on GPU
#
# Check if you can train on GPU. If you can, set this as a variable and move your models to GPU.
# > Later, we'll also move any inputs our models and loss functions see (real_images, z, and ground truth labels) to GPU as well.
# +
train_on_gpu = torch.cuda.is_available()
if train_on_gpu:
# move models to GPU
G.cuda()
D.cuda()
print('GPU available for training. Models moved to GPU')
else:
print('Training on CPU.')
# -
# ---
# ## Discriminator and Generator Losses
#
# Now we need to calculate the losses. And this will be exactly the same as before.
#
# ### Discriminator Losses
#
# > * For the discriminator, the total loss is the sum of the losses for real and fake images, `d_loss = d_real_loss + d_fake_loss`.
# * Remember that we want the discriminator to output 1 for real images and 0 for fake images, so we need to set up the losses to reflect that.
#
# The losses will by binary cross entropy loss with logits, which we can get with [BCEWithLogitsLoss](https://pytorch.org/docs/stable/nn.html#bcewithlogitsloss). This combines a `sigmoid` activation function **and** and binary cross entropy loss in one function.
#
# For the real images, we want `D(real_images) = 1`. That is, we want the discriminator to classify the the real images with a label = 1, indicating that these are real. The discriminator loss for the fake data is similar. We want `D(fake_images) = 0`, where the fake images are the _generator output_, `fake_images = G(z)`.
#
# ### Generator Loss
#
# The generator loss will look similar only with flipped labels. The generator's goal is to get `D(fake_images) = 1`. In this case, the labels are **flipped** to represent that the generator is trying to fool the discriminator into thinking that the images it generates (fakes) are real!
# +
def real_loss(D_out, smooth=False):
batch_size = D_out.size(0)
# label smoothing
if smooth:
# smooth, real labels = 0.9
labels = torch.ones(batch_size)*0.9
else:
labels = torch.ones(batch_size) # real labels = 1
# move labels to GPU if available
if train_on_gpu:
labels = labels.cuda()
# binary cross entropy with logits loss
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
def fake_loss(D_out):
batch_size = D_out.size(0)
labels = torch.zeros(batch_size) # fake labels = 0
if train_on_gpu:
labels = labels.cuda()
criterion = nn.BCEWithLogitsLoss()
# calculate loss
loss = criterion(D_out.squeeze(), labels)
return loss
# -
# ## Optimizers
#
# Not much new here, but notice how I am using a small learning rate and custom parameters for the Adam optimizers, This is based on some research into DCGAN model convergence.
#
# ### Hyperparameters
#
# GANs are very sensitive to hyperparameters. A lot of experimentation goes into finding the best hyperparameters such that the generator and discriminator don't overpower each other. Try out your own hyperparameters or read [the DCGAN paper](https://arxiv.org/pdf/1511.06434.pdf) to see what worked for them.
# +
import torch.optim as optim
# params
lr = 0.0002
beta1 = 0.5
beta2 = 0.999
# Create optimizers for the discriminator and generator
d_optimizer = optim.Adam(D.parameters(), lr, [beta1, beta2])
g_optimizer = optim.Adam(G.parameters(), lr, [beta1, beta2])
# -
# ---
# ## Training
#
# Training will involve alternating between training the discriminator and the generator. We'll use our functions `real_loss` and `fake_loss` to help us calculate the discriminator losses in all of the following cases.
#
# ### Discriminator training
# 1. Compute the discriminator loss on real, training images
# 2. Generate fake images
# 3. Compute the discriminator loss on fake, generated images
# 4. Add up real and fake loss
# 5. Perform backpropagation + an optimization step to update the discriminator's weights
#
# ### Generator training
# 1. Generate fake images
# 2. Compute the discriminator loss on fake images, using **flipped** labels!
# 3. Perform backpropagation + an optimization step to update the generator's weights
#
# #### Saving Samples
#
# As we train, we'll also print out some loss statistics and save some generated "fake" samples.
#
# **Evaluation mode**
#
# Notice that, when we call our generator to create the samples to display, we set our model to evaluation mode: `G.eval()`. That's so the batch normalization layers will use the population statistics rather than the batch statistics (as they do during training), *and* so dropout layers will operate in eval() mode; not turning off any nodes for generating samples.
# +
import pickle as pkl
# training hyperparams
num_epochs = 60
# keep track of loss and generated, "fake" samples
samples = []
losses = []
print_every = 300
# Get some fixed data for sampling. These are images that are held
# constant throughout training, and allow us to inspect the model's performance
sample_size=16
fixed_z = np.random.uniform(-1, 1, size=(sample_size, z_size))
fixed_z = torch.from_numpy(fixed_z).float()
# train the network
for epoch in range(num_epochs):
for batch_i, (real_images, _) in enumerate(train_loader):
batch_size = real_images.size(0)
# important rescaling step
real_images = scale(real_images)
# ============================================
# TRAIN THE DISCRIMINATOR
# ============================================
d_optimizer.zero_grad()
# 1. Train with real images
# Compute the discriminator losses on real images
if train_on_gpu:
real_images = real_images.cuda()
D_real = D(real_images)
d_real_loss = real_loss(D_real)
# 2. Train with fake images
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
# move x to GPU, if available
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
D_fake = D(fake_images)
d_fake_loss = fake_loss(D_fake)
# add up loss and perform backprop
d_loss = d_real_loss + d_fake_loss
d_loss.backward()
d_optimizer.step()
# =========================================
# TRAIN THE GENERATOR
# =========================================
g_optimizer.zero_grad()
# 1. Train with fake images and flipped labels
# Generate fake images
z = np.random.uniform(-1, 1, size=(batch_size, z_size))
z = torch.from_numpy(z).float()
if train_on_gpu:
z = z.cuda()
fake_images = G(z)
# Compute the discriminator losses on fake images
# using flipped labels!
D_fake = D(fake_images)
g_loss = real_loss(D_fake) # use real loss to flip labels
# perform backprop
g_loss.backward()
g_optimizer.step()
# Print some loss stats
if batch_i % print_every == 0:
# append discriminator loss and generator loss
losses.append((d_loss.item(), g_loss.item()))
# print discriminator and generator loss
print('Epoch [{:5d}/{:5d}] | d_loss: {:6.4f} | g_loss: {:6.4f}'.format(
epoch+1, num_epochs, d_loss.item(), g_loss.item()))
## AFTER EACH EPOCH##
# generate and save sample, fake images
G.eval() # for generating samples
if train_on_gpu:
fixed_z = fixed_z.cuda()
samples_z = G(fixed_z)
samples.append(samples_z)
G.train() # back to training mode
# Save training generator samples
with open('train_samples.pkl', 'wb') as f:
pkl.dump(samples, f)
# -
# ## Training loss
#
# Here we'll plot the training losses for the generator and discriminator, recorded after each epoch.
fig, ax = plt.subplots()
losses = np.array(losses)
plt.plot(losses.T[0], label='Discriminator', alpha=0.5)
plt.plot(losses.T[1], label='Generator', alpha=0.5)
plt.title("Training Losses")
plt.legend()
# ## Generator samples from training
#
# Here we can view samples of images from the generator. We'll look at the images we saved during training.
# helper function for viewing a list of passed in sample images
def view_samples(epoch, samples):
fig, axes = plt.subplots(figsize=(16,4), nrows=2, ncols=8, sharey=True, sharex=True)
for ax, img in zip(axes.flatten(), samples[epoch]):
img = img.detach().cpu().numpy()
img = np.transpose(img, (1, 2, 0))
img = ((img +1)*255 / (2)).astype(np.uint8) # rescale to pixel range (0-255)
ax.xaxis.set_visible(False)
ax.yaxis.set_visible(False)
im = ax.imshow(img.reshape((32,32,3)))
_ = view_samples(-1, samples)
|
dcgan-svhn/DCGAN_Exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object detection: Added brightness
#
# <div class="alert alert-info">
#
# **Note:** [Object detection requirements](../installation.rst#object-detection-example-and-notebooks-requirements).
#
# </div>
#
# <div class="alert alert-warning">
#
# **Warning:** Runtimes can be several hours even on clusters.
#
# </div>
# We compared the performance of models from FaceBook's Detectron project and YOLOv3 model from <NAME>, when different error sources were added. The models from FaceBook's Detectron project were FasterRCNN, MaskRCNN and RetinaNet.
# + pycharm={"is_executing": false}
import re
from abc import ABC, abstractmethod
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
from dpemu import runner
from dpemu.dataset_utils import load_coco_val_2017
from dpemu.filters.image import Brightness
from dpemu.ml_utils import run_ml_module_using_cli, load_yolov3
from dpemu.nodes import Array, Series
from dpemu.plotting_utils import print_results_by_model, visualize_scores
from dpemu.utils import get_project_root
# -
# We used 118 287 jpg images (COCO train2017) as the train set and 5000 images (COCO val2017) as the test set to calculate the mAP-50 scores.
# + pycharm={"name": "#%%\n"}
def get_data():
imgs, _, _, img_filenames = load_coco_val_2017()
return imgs, img_filenames
# -
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
def get_err_root_node():
err_node = Array()
err_root_node = Series(err_node)
err_node.addfilter(Brightness("tar", "rat", "range"))
return err_root_node
# -
# ## Examples from run_yolo_example.py
#
# ### rat: 0.0
#
# 
#
# ### rat: 0.47
#
# 
#
# ### rat: 0.93
#
# 
#
# ### rat: 1.4
#
# 
# + pycharm={"is_executing": false, "name": "#%%\n"}
def get_err_params_list():
rat_steps = np.linspace(0, 1.4, num=8)
return [{"tar": 1, "rat": rat, "range": 255} for rat in rat_steps]
# -
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
class Preprocessor:
def run(self, _, imgs, params):
img_filenames = params["img_filenames"]
for i, img_arr in enumerate(imgs):
img = Image.fromarray(img_arr)
path_to_img = f"{get_project_root()}/tmp/val2017/" + img_filenames[i]
img.save(path_to_img, "jpeg", quality=100)
return None, imgs, {}
# -
# Detectron's model zoo had pretrained weights for FasterRCNN, MaskRCNN and RetinaNet. YOLOv3's weights were trained by us, using the Kale cluster of University of Helsinki. The training took approximately five days when two NVIDIA Tesla V100 GPUs were used.
# + pycharm={"is_executing": false, "name": "#%%\n"}
class YOLOv3Model:
def run(self, _, imgs, params):
path_to_yolov3_weights, path_to_yolov3_cfg = load_yolov3()
cline = f"{get_project_root()}/libs/darknet/darknet detector map {get_project_root()}/data/coco.data \
{path_to_yolov3_cfg} {path_to_yolov3_weights}"
out = run_ml_module_using_cli(cline, show_stdout=False)
match = re.search(r"\(mAP@0.50\) = (\d+\.\d+)", out)
return {"mAP-50": round(float(match.group(1)), 3)}
class AbstractDetectronModel(ABC):
def run(self, _, imgs, params):
path_to_cfg = self.get_path_to_cfg()
url_to_weights = self.get_url_to_weights()
cline = f"""{get_project_root()}/libs/Detectron/tools/test_net.py \
--cfg {path_to_cfg} \
TEST.WEIGHTS {url_to_weights} \
NUM_GPUS 1 \
TEST.DATASETS '("coco_2017_val",)' \
MODEL.MASK_ON False \
OUTPUT_DIR {get_project_root()}/tmp \
DOWNLOAD_CACHE {get_project_root()}/tmp"""
out = run_ml_module_using_cli(cline, show_stdout=False)
match = re.search(r"IoU=0.50 \| area= all \| maxDets=100 ] = (\d+\.\d+)", out)
return {"mAP-50": round(float(match.group(1)), 3)}
@abstractmethod
def get_path_to_cfg(self):
pass
@abstractmethod
def get_url_to_weights(self):
pass
class FasterRCNNModel(AbstractDetectronModel):
def get_path_to_cfg(self):
return f"{get_project_root()}/libs/Detectron/configs/12_2017_baselines/e2e_faster_rcnn_X-101-64x4d-FPN_1x.yaml"
def get_url_to_weights(self):
return (
"https://dl.fbaipublicfiles.com/detectron/35858015/12_2017_baselines/"
"e2e_faster_rcnn_X-101-64x4d-FPN_1x.yaml.01_40_54.1xc565DE/output/train/"
"coco_2014_train%3Acoco_2014_valminusminival/generalized_rcnn/model_final.pkl"
)
class MaskRCNNModel(AbstractDetectronModel):
def get_path_to_cfg(self):
return f"{get_project_root()}/libs/Detectron/configs/12_2017_baselines/e2e_mask_rcnn_X-101-64x4d-FPN_1x.yaml"
def get_url_to_weights(self):
return (
"https://dl.fbaipublicfiles.com/detectron/36494496/12_2017_baselines/"
"e2e_mask_rcnn_X-101-64x4d-FPN_1x.yaml.07_50_11.fkwVtEvg/output/train/"
"coco_2014_train%3Acoco_2014_valminusminival/generalized_rcnn/model_final.pkl"
)
class RetinaNetModel(AbstractDetectronModel):
def get_path_to_cfg(self):
return f"{get_project_root()}/libs/Detectron/configs/12_2017_baselines/retinanet_X-101-64x4d-FPN_1x.yaml"
def get_url_to_weights(self):
return (
"https://dl.fbaipublicfiles.com/detectron/36768875/12_2017_baselines/"
"retinanet_X-101-64x4d-FPN_1x.yaml.08_34_37.FSXgMpzP/output/train/"
"coco_2014_train%3Acoco_2014_valminusminival/retinanet/model_final.pkl"
)
# -
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
def get_model_params_dict_list():
return [
{"model": FasterRCNNModel, "params_list": [{}]},
{"model": MaskRCNNModel, "params_list": [{}]},
{"model": RetinaNetModel, "params_list": [{}]},
{"model": YOLOv3Model, "params_list": [{}]},
]
# -
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
def visualize(df):
visualize_scores(
df,
score_names=["mAP-50"],
is_higher_score_better=[True],
err_param_name="rat",
title="Object detection with added brightness"
)
plt.show()
# -
#
# + pycharm={"is_executing": false, "name": "#%%\n"}
def main():
imgs, img_filenames = get_data()
df = runner.run(
train_data=None,
test_data=imgs,
preproc=Preprocessor,
preproc_params={"img_filenames": img_filenames},
err_root_node=get_err_root_node(),
err_params_list=get_err_params_list(),
model_params_dict_list=get_model_params_dict_list(),
n_processes=1
)
print_results_by_model(df, dropped_columns=["tar", "range"])
visualize(df)
# -
#
# + pycharm={"is_executing": true, "name": "#%%\n"}
main()
# -
# The notebook for this case study can be found [here](https://github.com/dpEmu/dpEmu/blob/master/docs/case_studies/Object_Detection_Added_Brightness.ipynb).
|
docs/case_studies/Object_Detection_Added_Brightness.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from numpy.linalg import inv
X = np.array([[1, 50], [1, 60], [1,70], [1,100]])
X
Y = np.array([[10], [30], [40], [50]])
Y
# +
w = inv(
(X.T).dot(X)
).dot(
X.T
).dot(Y)
w
# +
margin = 10
X_min = X[:,1].min() - margin
X_max = X[:,1].max() + margin
X_support = np.linspace(X_min, X_max, num=100)
Y_model = w[0][0] + w[1][0] * X_support
# +
plt.xlim(X_min, X_max)
plt.ylim(0, Y[:,0].max() + margin)
plt.scatter(X[:,1], Y[:,0], 40, 'g', 'o', alpha=0.8)
plt.plot(X_support, Y_model)
plt.show()
# -
|
regression_very_low_level.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# The line below sets the environment
# variable CUDA_VISIBLE_DEVICES
get_ipython().magic('env CUDA_VISIBLE_DEVICES = ')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import multiprocessing as mp # will come in handy due to the size of the data
import os.path
import random
import time
from collections import OrderedDict
import io
from datetime import datetime
import gc # garbage collector
import sklearn
import lightgbm as lgb
from sklearn.preprocessing import LabelEncoder
import math
import sys
from collections import defaultdict
import re
import logging
from sklearn.model_selection import KFold
from xgboost.sklearn import XGBClassifier
from sklearn import cross_validation, metrics #Additional scklearn functions
from sklearn.grid_search import GridSearchCV #Perforing grid search
# This is a bit of magic to make matplotlib figures appear inline in the notebook
# rather than in a new window.
get_ipython().magic('matplotlib inline')
plt.rcParams['figure.figsize'] = (10.0, 8.0) # set default size of plots
plt.rcParams['image.interpolation'] = 'nearest'
plt.rcParams['image.cmap'] = 'gray'
# Some more magic so that the notebook will reload external python modules;
# see http://stackoverflow.com/questions/1907993/autoreload-of-modules-in-ipython
get_ipython().magic('load_ext autoreload')
get_ipython().magic('autoreload 2')
# -
# ## Write a pandas dataframe to disk as gunzip compressed csv
# - df.to_csv('dfsavename.csv.gz', compression='gzip')
#
# ## Read from disk
# - df = pd.read_csv('dfsavename.csv.gz', compression='gzip')
#
# ## Magic useful
# - %%timeit for the whole cell
# - %timeit for the specific line
# - %%latex to render the cell as a block of latex
# - %prun and %%prun
DATASET_PATH = '/media/rs/0E06CD1706CD0127/Kapok/WSDM/'
HDF_FILENAME = DATASET_PATH + 'datas.h5'
SUBMISSION_FILENAME = DATASET_PATH + 'submission_{}.csv'
VALIDATION_INDICE = DATASET_PATH + 'validation_indice.csv'
def set_logging(logger_name, logger_file_name):
log = logging.getLogger(logger_name)
log.setLevel(logging.DEBUG)
# create formatter and add it to the handlers
print_formatter = logging.Formatter('%(message)s')
file_formatter = logging.Formatter('%(asctime)s - %(name)s_%(levelname)s: %(message)s')
# create file handler which logs even debug messages
fh = logging.FileHandler(logger_file_name, mode='w')
fh.setLevel(logging.DEBUG)
fh.setFormatter(file_formatter)
log.addHandler(fh)
# both output to console and file
consoleHandler = logging.StreamHandler()
consoleHandler.setFormatter(print_formatter)
log.addHandler(consoleHandler)
return log
log = set_logging('MUSIC', DATASET_PATH + 'music_gbm.log')
log.info('here is an info message.')
# +
# TRAIN_FILE = DATASET_PATH + 'train.csv'
# TEST_FILE = DATASET_PATH + 'test.csv'
# MEMBER_FILE = DATASET_PATH + 'members.csv'
# SONG_FILE = DATASET_PATH + 'fix_songs.csv'
# SONG_EXTRA_FILE = DATASET_PATH + 'song_extra_info.csv'
# train_data = pd.read_csv(TRAIN_FILE)
# test_data = pd.read_csv(TEST_FILE)
# member_data = pd.read_csv(MEMBER_FILE)
# song_data = pd.read_csv(SONG_FILE)
# song_extra_data = pd.read_csv(SONG_EXTRA_FILE)
# songs_all = pd.merge(left = song_data, right = song_extra_data, how = 'left', on='song_id')
# train_with_mem = pd.merge(left = train_data, right = member_data, how = 'left', on='msno')
# train_all = pd.merge(left = train_with_mem, right = songs_all, how = 'left', on='song_id')
# test_with_mem = pd.merge(left = test_data, right = member_data, how = 'left', on='msno')
# test_all = pd.merge(left = test_with_mem, right = songs_all, how = 'left', on='song_id')
# del train_with_mem, test_with_mem; gc.collect()
# def convert_unicode_to_str(df):
# df.columns = df.columns.astype(str)
# types = df.apply(lambda x: pd.api.types.infer_dtype(df.values))
# #print(types)#mixed-integer
# for col in types[types == 'mixed-integer'].index:
# df[col] = df[col].astype(str)
# for col in types[types == 'mixed'].index:
# df[col] = df[col].astype(str)
# return df
# store = pd.HDFStore(HDF_FILENAME)
# store['train_data'] = convert_unicode_to_str(train_all)
# store['test_data'] = convert_unicode_to_str(test_all)
# store['song_data'] = convert_unicode_to_str(songs_all)
# store['test_id'] = test_data.id
# store.close()
# -
# store_test = pd.HDFStore(HDF_FILENAME)
# train = store_test['train_data'][0:100]
# test = store_test['test_data'][0:100]
# test_id = store_test['test_id'][0:100]
# store_test.close()
store_test = pd.HDFStore(HDF_FILENAME)
train = store_test['train_data']
test = store_test['test_data']
test_id = store_test['test_id']
store_test.close()
def split_country(input_data):
def get_country(isrc):
if isinstance(isrc, str) and isrc != 'nan':
return isrc[0:2]
else:
return np.nan
countries = train['isrc'].apply(get_country)
country_list = list(countries.value_counts().index)
country_map = dict(zip(country_list, country_list))
country_map['QM'] = 'QZ'
country_map['US'] = 'QZ'
return countries.map(country_map)
train['country'] = split_country(train)
test['country'] = split_country(test)
# +
def isrc_to_year(isrc):
if isinstance(isrc, str) and isrc != 'nan':
if int(isrc[5:7]) > 17:
return 1900 + int(isrc[5:7])
else:
return 2000 + int(isrc[5:7])
else:
return np.nan
train['song_year'] = train['isrc'].apply(isrc_to_year)
test['song_year'] = test['isrc'].apply(isrc_to_year)
train.drop(['isrc'], axis = 1, inplace = True)
test.drop(['isrc'], axis = 1, inplace = True)
# -
def split_reg_date(input_data):
input_data['registration_year'] = input_data['registration_init_time'].apply(lambda x : int(str(x)[0:4]))
input_data['registration_year'] = pd.to_numeric(input_data['registration_year'], downcast='unsigned')
input_data['registration_month'] = input_data['registration_init_time'].apply(lambda x : int(str(x)[4:6]))
input_data['registration_month'] = pd.to_numeric(input_data['registration_month'], downcast='unsigned')
input_data['registration_day'] = input_data['registration_init_time'].apply(lambda x : int(str(x)[6:8]))
input_data['registration_day'] = pd.to_numeric(input_data['registration_day'], downcast='unsigned')
return input_data
def split_expir_date(input_data):
input_data['expiration_year'] = input_data['expiration_date'].apply(lambda x : int(str(x)[0:4]))
input_data['expiration_year'] = pd.to_numeric(input_data['expiration_year'], downcast='unsigned')
input_data['expiration_month'] = input_data['expiration_date'].apply(lambda x : int(str(x)[4:6]))
input_data['expiration_month'] = pd.to_numeric(input_data['expiration_month'], downcast='unsigned')
input_data['expiration_day'] = input_data['expiration_date'].apply(lambda x : int(str(x)[6:8]))
input_data['expiration_day'] = pd.to_numeric(input_data['expiration_day'], downcast='unsigned')
return input_data
def date_to_day(input_data):
# 转换注册时间
input_data['registration_init_time'] = pd.to_datetime(input_data['registration_init_time'],format="%Y%m%d")
input_data['expiration_date'] = pd.to_datetime(input_data['expiration_date'],format="%Y%m%d")
days = input_data.expiration_date - input_data.registration_init_time
days = [d.days for d in days]
input_data['days']=days
return input_data
# +
train = split_reg_date(train)
test = split_reg_date(test)
train = split_expir_date(train)
test = split_expir_date(test)
train = date_to_day(train)
test = date_to_day(test)
train.drop('registration_init_time',axis=1,inplace=True)
train.drop('expiration_date',axis=1,inplace=True)
test.drop('registration_init_time',axis=1,inplace=True)
test.drop('expiration_date',axis=1,inplace=True)
# -
train['song_length'] = pd.to_numeric(train['song_length'].replace('nan', '235415'), downcast='unsigned')
test['song_length'] = pd.to_numeric(test['song_length'].replace('nan', '235415'), downcast='unsigned')
for col in train.columns: print(col, ':', train[col].dtype)
for col in [col for col in test.columns if col != 'id' ]:
if train[col].dtype == object:
train[col] = train[col].astype('category')
test[col] = test[col].astype('category')
# +
# # encode registered_via, the less number of occurrences are merged into the top item which has the max number of occurrences
# registered_via_hist = pd.concat([train['registered_via'], test['registered_via']], axis = 0).value_counts()
# registered_via_map = dict(zip(registered_via_hist.index, [int(s) for s in registered_via_hist.index.values]))
# registered_via_map[registered_via_hist.index[-1]] = int(str(registered_via_hist.index.values[0]))
# train['registered_via'] = train['registered_via'].map(registered_via_map)
# test['registered_via'] = test['registered_via'].map(registered_via_map)
# +
# # encode language, fill nan with most occurrences item
# language_hist = pd.concat([train['language'], test['language']], axis = 0).value_counts()
# language_map = dict(zip(language_hist.index, [int(float(s)) for s in language_hist.index.values if s != 'nan']))
# language_map['nan'] = int(float(str(language_hist.index.values[0])))
# train['language'] = train['language'].map(language_map)
# test['language'] = test['language'].map(language_map)
# +
# # encode country, fill nan with most occurrences item
# country_hist = pd.concat([train['country'], test['country']], axis = 0).value_counts()
# merge_per = 0.25
# country_map = dict(zip(country_hist.index, list(range(len(country_hist)))))
# for key in list(country_hist[-int(len(country_hist)*merge_per):].index):
# country_map[key] = int(len(country_hist)*(1-merge_per)) + 1
# train['country'] = train['country'].map(country_map)
# test['country'] = test['country'].map(country_map)
# +
# msno : category ; uinque values: 30755
# song_id : category ; uinque values: 359966
# - source_system_tab : category ; uinque values: 10
# - source_screen_name : category ; uinque values: 21
# - source_type : category ; uinque values: 13
# - target : object ; uinque values: 2
# - city : category ; uinque values: 21
# - bd : category ; uinque values: 92
# - gender : category ; uinque values: 3
# - registered_via : category ; uinque values: 5
# song_length : uint32 ; uinque values: 60271
# genre_ids : category ; uinque values: 573
# artist_name : category ; uinque values: 40587
# composer : category ; uinque values: 76072
# lyricist : category ; uinque values: 33895
# - language : category ; uinque values: 11
# name : category ; uinque values: 234144
# - country : category ; uinque values: 107
# - song_year : float64 ; uinque values: 100
# - registration_year : uint16 ; uinque values: 14
# - registration_month : uint8 ; uinque values: 12
# - registration_date : uint8 ; uinque values: 31
# - expiration_year : uint16 ; uinque values: 18
# - expiration_month : uint8 ; uinque values: 12
# -
def one_hot_transform(input_train_data, input_test_data, columns_to_transform):
for col in columns_to_transform:
le = LabelEncoder()
train_values = list(input_train_data[col].unique())
test_values = list(input_test_data[col].unique())
le.fit(train_values + test_values)
input_train_data[col] = le.transform(input_train_data[col])
input_test_data[col] = le.transform(input_test_data[col])
return input_train_data, input_test_data
# +
#train, test = one_hot_transform(train, test, ['source_system_tab', 'source_screen_name', 'source_type', 'city', 'gender', 'name'])#, 'artist_name', 'composer', 'lyricist'])
# -
# TODO: wether song_id should be merged like this or not? 231475 reserved and 188364 merged
def encode_with_merge(input_train, input_test, columns, merge_value):
for index, col in enumerate(columns):
values_hist = pd.concat([input_train[col], input_train[col]], axis = 0).value_counts()
reserve_rows = values_hist[values_hist!=merge_value[index]]
merge_rows = values_hist[values_hist==merge_value[index]]
reserve_dict = dict(zip(list(reserve_rows.index), list(range(len(reserve_rows)))))
merge_dict = dict(zip(list(merge_rows.index), [len(reserve_rows)+1]*len(merge_rows.index)))
map_dict = {**reserve_dict, **merge_dict}
language_map['nan'] = int(float(str(language_hist.index.values[0])))
input_train[col] = input_train[col].map(map_dict)
input_test[col] = input_test[col].map(map_dict)
return input_train, input_test
# +
#train, test = encode_with_merge(train, test, ['msno', 'song_id', 'genre_ids'], [1, 1, 1])
# print(train.head())
# print(test.head())
# -
train_org, test_org = train, test
train = train_org.copy(deep=True)
test = test_org.copy(deep=True)
store_test = pd.HDFStore(VALIDATION_INDICE)
validation_list = store_test['keep_index']['index'].values
store_test.close()
train['target'] = pd.to_numeric(train['target'], downcast='signed')
#validation_use = train.iloc[validation_list].copy(deep=True).reset_index(drop=True)
validation_use = train.iloc[list(range(7277417, 7377417))].copy(deep=True).reset_index(drop=True)
#train_use = train.drop(validation_list)
train_use = train.drop(list(range(7277417, 7377417)))
# train['target'] = pd.to_numeric(train['target'], downcast='signed')
# validation_use = train[50:].copy(deep=True).reset_index(drop=True)
# train_use = train.drop(list(range(50,100)))
# +
# for col in train_use.columns: print(col, ':', train_use[col].dtype, '; uinque values:', len(train_use[col].value_counts()))
# -
def log_transform(train_data, validation_data, test_data):
train_data['song_length'] = np.log(pd.to_numeric(train_data['song_length'], downcast='float') + 1)
validation_data['song_length'] = np.log(pd.to_numeric(validation_data['song_length'], downcast='float') + 1)
test_data['song_length'] = np.log(pd.to_numeric(test_data['song_length'], downcast='float') + 1)
return train_data, validation_data, test_data
train_use, validation_use, test = log_transform(train_use, validation_use, test)
def cal_composer_hot_rate(train_data, val_data, test_data):
temp_data = pd.concat([train_data[['composer']], val_data[['composer']], test_data[['composer']]], axis=0, join="outer")
temp_data['composer'] = temp_data['composer'].apply(lambda x : x.replace(u'、','|'))
df_temp = temp_data['composer'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['composer_{}'.format(x) for x in df_temp.columns]
temp_data = pd.concat([df_temp['composer_0'], df_temp['composer_1'], df_temp['composer_2']], axis=0, join="outer")
temp_data.reset_index(drop=True)
composer_hot = np.log(temp_data.value_counts()+1)
#composer_hot = temp_data.value_counts()
composer_hot['nan'] = 0.
composer_hot['nan'] = composer_hot.mean()
#print(composer_hot)
def encoder_each(input_data, hot_hist):
input_data = input_data.copy()
input_data['composer'] = input_data['composer'].apply(lambda x : x.replace(u'、','|'))
df_temp = input_data['composer'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['composer_{}'.format(x) for x in df_temp.columns]
hot_hist = hot_hist.reset_index()
hot_hist.index.name='index'
hot_hist.columns = ['composer_0', 'composer_0_score']
df_temp = df_temp.merge(right = hot_hist, how = 'left', on='composer_0')
hot_hist.columns = ['composer_1', 'composer_1_score']
df_temp = df_temp.merge(right = hot_hist, how = 'left', on='composer_1')
hot_hist.columns = ['composer_2', 'composer_2_score']
df_temp = df_temp.merge(right = hot_hist, how = 'left', on='composer_2')
df_temp['composer_score'] = df_temp[['composer_0_score','composer_1_score','composer_2_score']].max(axis=1)
#df_temp['composer_score'] = df_temp['composer_0_score']
input_data['composer_score'] = df_temp['composer_score']
input_data.drop('composer', inplace=True, axis = 1)
#input_data = input_data.drop('composer', inplace=False, axis = 1)
input_data['composer'] = df_temp['composer_0']
return input_data
train_data = encoder_each(train_data, composer_hot)
val_data = encoder_each(val_data, composer_hot)
test_data = encoder_each(test_data, composer_hot)
return train_data, val_data, test_data
train_use, validation_use, test = cal_composer_hot_rate(train_use, validation_use, test)
def cal_lyricist_hot_rate(train_data, val_data, test_data):
temp_data = pd.concat([train_data[['lyricist']], val_data[['lyricist']], test_data[['lyricist']]], axis=0, join="outer")
temp_data['lyricist'] = temp_data['lyricist'].apply(lambda x : x.replace(u'、','|'))
df_temp = temp_data['lyricist'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['lyricist_{}'.format(x) for x in df_temp.columns]
#temp_data = df_temp['lyricist_0']
temp_data = pd.concat([df_temp['lyricist_0'], df_temp['lyricist_1'], df_temp['lyricist_2']], axis=0, join="outer")
temp_data.reset_index(drop=True)
lyricist_hot = np.log(temp_data.value_counts()+1)
#composer_hot = temp_data.value_counts()
lyricist_hot['nan'] = 0.
lyricist_hot['nan'] = lyricist_hot.mean()
#print(lyricist_hot)
def encoder_each(input_data, hot_hist):
input_data = input_data.copy()
input_data['lyricist'] = input_data['lyricist'].apply(lambda x : x.replace(u'、','|'))
df_temp = input_data['lyricist'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['lyricist_{}'.format(x) for x in df_temp.columns]
hot_hist = hot_hist.reset_index()
hot_hist.index.name='index'
hot_hist.columns = ['lyricist_0', 'lyricist_0_score']
df_temp = df_temp.merge(right = hot_hist, how = 'left', on='lyricist_0')
df_temp['lyricist_score'] = df_temp['lyricist_0_score']
input_data['lyricist_score'] = df_temp['lyricist_score']
input_data.drop('lyricist', inplace=True, axis = 1)
input_data['lyricist'] = df_temp['lyricist_0']
return input_data
train_data = encoder_each(train_data, lyricist_hot)
val_data = encoder_each(val_data, lyricist_hot)
test_data = encoder_each(test_data, lyricist_hot)
return train_data, val_data, test_data
train_use, validation_use, test = cal_lyricist_hot_rate(train_use, validation_use, test)
def cal_artist_hot_rate(train_data, val_data, test_data):
temp_data = pd.concat([train_data[['artist_name']], val_data[['artist_name']], test_data[['artist_name']]], axis=0, join="outer")
temp_data['artist_name'] = temp_data['artist_name'].apply(lambda x : x.replace(u'、','|'))
df_temp = temp_data['artist_name'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['artist_name_{}'.format(x) for x in df_temp.columns]
#temp_data = df_temp['artist_name_0']
temp_data = pd.concat([df_temp['artist_name_0'], df_temp['artist_name_1'], df_temp['artist_name_2']], axis=0, join="outer")
temp_data.reset_index(drop=True)
artist_hot = np.log(temp_data.value_counts()+1)
#composer_hot = temp_data.value_counts()
artist_hot['nan'] = 0.
artist_hot['nan'] = artist_hot.mean()
#print(artist_hot)
def encoder_each(input_data, hot_hist):
input_data = input_data.copy()
input_data['artist_name'] = input_data['artist_name'].apply(lambda x : x.replace(u'、','|'))
df_temp = input_data['artist_name'].str.split('\s{0,}[\|\\\\/]\s{0,}', 3, expand=True)
df_temp.columns = ['artist_name_{}'.format(x) for x in df_temp.columns]
hot_hist = hot_hist.reset_index()
hot_hist.index.name='index'
hot_hist.columns = ['artist_name_0', 'artist_name_0_score']
df_temp = df_temp.merge(right = hot_hist, how = 'left', on='artist_name_0')
df_temp['artist_name_score'] = df_temp['artist_name_0_score']
input_data['artist_name_score'] = df_temp['artist_name_score']
input_data.drop('artist_name', inplace=True, axis = 1)
input_data['artist_name'] = df_temp['artist_name_0']
return input_data
train_data = encoder_each(train_data, artist_hot)
val_data = encoder_each(val_data, artist_hot)
test_data = encoder_each(test_data, artist_hot)
return train_data, val_data, test_data
train_use, validation_use, test = cal_artist_hot_rate(train_use, validation_use, test)
print(train_use.head().columns)
# +
# temp_data = pd.concat([train_use[['composer']], validation_use[['composer']], test[['composer']]], axis=0, join="inner")
# temp_data['composer'].apply(lambda x : len(x.replace(u'、','|').split('|'))).value_counts().plot()
# +
# df = train_use.head(100000).copy(deep=True)
# print(df['lyricist'].value_counts())
# +
time_wnd = [2018, 0]
#time_wnd = [2018, 0]#, 2000, 2010, 2014, 2018]
def cal_song_listen_times(train_data, test_data, val_data):
all_data = pd.concat([train_data[['song_id', 'song_year', 'msno']], val_data[['song_id', 'song_year', 'msno']], test_data[['song_id', 'song_year', 'msno']]], axis=0, join="inner")
#all_data['song_id'] = pd.to_numeric(all_data['song_id'], downcast='unsigned')
#all_data['msno'] = pd.to_numeric(all_data['msno'], downcast='unsigned')
for index, _ in enumerate(time_wnd[:-1]):
begin_time, end_time = time_wnd[index] < time_wnd[index+1] and (time_wnd[index], time_wnd[index+1]) or (time_wnd[index+1], time_wnd[index])
# begin_time = time_wnd[index]
# end_time = time_wnd[index+1]
select_data = all_data[all_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
#select_data['target'] = pd.to_numeric(select_data['target'], downcast='signed')
grouped = select_data[['song_id', 'msno']].groupby(['song_id'])
count_song = grouped.agg(['count'])
num_people_per_song = grouped.agg({"msno": lambda x: np.log(x.nunique()+1)})
popularity = pd.concat([np.log(count_song+1), num_people_per_song], axis=1, join="inner")
popularity.columns = ['popular_{}'.format(index), 'num_people_{}'.format(index)]
popularity = popularity.reset_index(drop=False)
train_data = train_data.merge(popularity, on='song_id', how ='left')
test_data = test_data.merge(popularity, on='song_id', how ='left')
val_data = val_data.merge(popularity, on='song_id', how ='left')
return train_data, test_data, val_data
def cal_song_listen_times_seperate(train_data, test_data, val_data):
def cal_each_of_them(input_data):
all_data = input_data[['song_id', 'song_year', 'msno']]
#all_data['song_id'] = pd.to_numeric(all_data['song_id'], downcast='unsigned')
#all_data['msno'] = pd.to_numeric(all_data['msno'], downcast='unsigned')
for index, _ in enumerate(time_wnd[:-1]):
begin_time, end_time = time_wnd[index] < time_wnd[index+1] and (time_wnd[index], time_wnd[index+1]) or (time_wnd[index+1], time_wnd[index])
# begin_time = time_wnd[index]
# end_time = time_wnd[index+1]
select_data = all_data[all_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
grouped = select_data[['song_id', 'msno']].groupby(['song_id'])
count_song = grouped.agg(['count'])
num_people_per_song = grouped.agg({"msno": lambda x: np.log(x.nunique()+1)})
popularity = pd.concat([np.log(count_song+1), num_people_per_song], axis=1, join="inner")
popularity.columns = ['popular_{}'.format(index), 'num_people_{}'.format(index)]
popularity = popularity.reset_index(drop=False)
all_data = input_data.merge(popularity, on='song_id', how ='left')
return all_data
return cal_each_of_them(train_data), cal_each_of_them(test_data), cal_each_of_them(val_data)
# time_wnd = [2018, 0, 2000, 2010, 2014, 2018]
# def cal_song_listen_times(train_data, test_data):
# test_data['song_id'] = pd.to_numeric(test_data['song_id'], downcast='unsigned')
# for index, _ in enumerate(time_wnd[:-1]):
# begin_time, end_time = time_wnd[index] < time_wnd[index+1] and (time_wnd[index], time_wnd[index+1]) or (time_wnd[index+1], time_wnd[index])
# # begin_time = time_wnd[index]
# # end_time = time_wnd[index+1]
# select_data = train_data[train_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
# select_data['target'] = pd.to_numeric(select_data['target'], downcast='signed')
# grouped = select_data[['song_id', 'target']].groupby(['song_id'])
# count_song = grouped.agg(['count'])
# mean_repeat_song = grouped['target'].mean()
# popularity = pd.concat([np.log(count_song+1), mean_repeat_song, np.log(count_song.multiply(mean_repeat_song, axis=0)+1)], axis=1, join="inner")
# popularity.columns = ['popular_{}'.format(index), 'mean_repeat_{}'.format(index), 'replay_prob_{}'.format(index)]
# popularity = popularity.reset_index(drop=False)
# test_data = test_data.merge(popularity, on='song_id', how ='left')
# train_data = train_data.merge(popularity, on='song_id', how ='left')
# return train_data, test_data
# time_wnd = [2018, 0, 2000, 2010, 2014, 2018]
# def cal_song_listen_times(train_data):
# train_data['song_id'] = pd.to_numeric(train_data['song_id'], downcast='unsigned')
# for index, _ in enumerate(time_wnd[:-1]):
# begin_time, end_time = time_wnd[index] < time_wnd[index+1] and (time_wnd[index], time_wnd[index+1]) or (time_wnd[index+1], time_wnd[index])
# # begin_time = time_wnd[index]
# # end_time = time_wnd[index+1]
# select_data = train_data[train_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
# select_data['target'] = pd.to_numeric(select_data['target'], downcast='signed')
# grouped = select_data[['song_id', 'target']].groupby(['song_id'])
# count_song = grouped.agg(['count'])
# mean_repeat_song = grouped['target'].mean()
# popularity = pd.concat([np.log(count_song+1), mean_repeat_song, np.log(count_song.multiply(mean_repeat_song, axis=0)+1)], axis=1, join="inner")
# popularity.columns = ['popular_{}'.format(index), 'mean_repeat_{}'.format(index), 'replay_prob_{}'.format(index)]
# popularity = popularity.reset_index(drop=False)
# train_data = train_data.merge(popularity, on='song_id', how ='left')
# return test_data
# -
train_use, test, validation_use = cal_song_listen_times(train_use, test, validation_use)
# train = cal_song_listen_times(train)
# test = cal_song_listen_times(test)
# +
#for col in train_use.columns: print(col, ':', train_use[col].dtype, '; uinque values:', len(train_use[col].value_counts()))
# +
people_time_wnd = [2018, 0]
#people_time_wnd = [2018, 0]#, 2000, 2010, 2014, 2018]
def get_people_active(train_data, test_data, val_data):
all_data = pd.concat([train_data[['song_id', 'song_year', 'msno']], val_data[['song_id', 'song_year', 'msno']], test_data[['song_id', 'song_year', 'msno']]], axis=0, join="inner")
#all_data['song_id'] = pd.to_numeric(all_data['song_id'], downcast='unsigned')
#all_data['msno'] = pd.to_numeric(all_data['msno'], downcast='unsigned')
for index, _ in enumerate(people_time_wnd[:-1]):
begin_time, end_time = people_time_wnd[index] < people_time_wnd[index+1] and (people_time_wnd[index], people_time_wnd[index+1]) or (people_time_wnd[index+1], people_time_wnd[index])
# begin_time = time_wnd[index]
# end_time = time_wnd[index+1]
select_data = all_data[all_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
#select_data['target'] = pd.to_numeric(select_data['target'], downcast='signed')
grouped = select_data[['song_id', 'msno']].groupby(['msno'])
count_song = grouped.agg(['count'])
num_people_per_song = grouped.agg({"song_id": lambda x: np.log(x.nunique()+1)})
popularity = pd.concat([np.log(count_song+1), num_people_per_song], axis=1, join="inner")
popularity.columns = ['active_{}'.format(index), 'num_song_{}'.format(index)]
popularity = popularity.reset_index(drop=False)
train_data = train_data.merge(popularity, on='msno', how ='left')
test_data = test_data.merge(popularity, on='msno', how ='left')
val_data = val_data.merge(popularity, on='msno', how ='left')
return train_data, test_data, val_data
def get_people_active_seperate(train_data, test_data, val_data):
def cal_each_of_them(input_data):
all_data = input_data[['song_id', 'song_year', 'msno']]
#all_data['song_id'] = pd.to_numeric(all_data['song_id'], downcast='unsigned')
#all_data['msno'] = pd.to_numeric(all_data['msno'], downcast='unsigned')
for index, _ in enumerate(people_time_wnd[:-1]):
begin_time, end_time = people_time_wnd[index] < people_time_wnd[index+1] and (people_time_wnd[index], people_time_wnd[index+1]) or (people_time_wnd[index+1], people_time_wnd[index])
# begin_time = time_wnd[index]
# end_time = time_wnd[index+1]
select_data = all_data[all_data['song_year'].map(lambda x: x>=begin_time and x < end_time)]
grouped = select_data[['song_id', 'msno']].groupby(['msno'])
count_song = grouped.agg(['count'])
num_people_per_song = grouped.agg({"song_id": lambda x: np.log(x.nunique()+1)})
popularity = pd.concat([np.log(count_song+1), num_people_per_song], axis=1, join="inner")
popularity.columns = ['active_{}'.format(index), 'num_song_{}'.format(index)]
popularity = popularity.reset_index(drop=False)
all_data = input_data.merge(popularity, on='msno', how ='left')
return all_data
return cal_each_of_them(train_data), cal_each_of_them(test_data), cal_each_of_them(val_data)
# test = test.reset_index(drop=False)
# #test['msno'] = test['msno'].astype(int)
# train['target'] = pd.to_numeric(train['target'], downcast='signed')
# grouped = train[['msno', 'target']].groupby(['msno'])
# count_msno = grouped.agg(['count'])
# mean_repeat_msno = grouped['target'].mean()
# popularity = pd.concat([np.log(count_msno+1), mean_repeat_msno, np.log(count_msno.multiply(mean_repeat_msno, axis=0)+1)], axis=1, join="inner")
# popularity.columns = ['ms_popular', 'ms_mean_repeat', 'ms_replay_prob']
# popularity = popularity.reset_index(drop=False)
# test = test.merge(popularity, on='msno', how ='left')
# train = train.merge(popularity, on='msno', how ='left')
# -
train_use, test, validation_use = get_people_active(train_use, test, validation_use)
def measure_by_different_city_lang_country(train_data, test_data, val_data):
temp_msno_songid = pd.concat([train_data[['composer', 'lyricist', 'artist_name', 'city', 'country', 'language']], val_data[['composer', 'lyricist', 'artist_name', 'city', 'country', 'language']], test_data[['composer', 'lyricist', 'artist_name', 'city', 'country', 'language']]], axis=0, join="outer")
count_dict = dict()
for col in ['composer', 'lyricist', 'artist_name']:
temp_df = None
for target in ['city', 'country', 'language']:
grouped = temp_msno_songid.groupby([col])
df = grouped.agg({target: lambda x: x.nunique()})
if temp_df is not None:
temp_df = pd.concat([temp_df, df], axis=1, join="inner")
else:
temp_df = df
temp_df.columns = [col + '_by_{}'.format(index) for index in target]
temp_msno_songid['composer'] = df_temp['composer_0']
#count_song = grouped.agg(['count'])
num_people_per_song = grouped.agg({"msno": lambda x: x.nunique()})
print(num_people_per_song)
num_people_per_song = grouped.agg({"song_id": lambda x: x.nunique()})#np.log(x.nunique()+1)})
print(num_people_per_song)
return
temp_data = df_temp['composer_0']
print(temp_data)
for col in ['source_system_tab', 'source_screen_name', 'source_type', 'city', 'gender',\
'name', 'artist_name', 'composer', 'lyricist', 'msno', 'song_id', 'genre_ids',\
'country', 'language', 'registered_via']:
train_use[col] = train_use[col].astype('category')
validation_use[col] = validation_use[col].astype('category')
test[col] = test[col].astype('category')
for col in train_use.columns: print(col, ':', train_use[col].dtype, '; uinque values:', len(train_use[col].value_counts()))
for col in test.columns: print(col, ':', test[col].dtype, '; uinque values:', len(test[col].value_counts()))
for col in train_use.columns: print(col, ':', train_use[col].dtype, '; uinque values:', len(train_use[col].value_counts()))
print(len(test_id), len(test))
# +
predictions = np.zeros(shape=[len(test)])
train_data = lgb.Dataset(train_use.drop(['target'],axis=1),label=train_use['target'])
val_data = lgb.Dataset(validation_use.drop(['target'],axis=1),label=validation_use['target'])
params = {
'objective': 'binary',
'boosting': 'gbdt',
'learning_rate': 0.1 ,
'verbose': 0,
'num_leaves': 108,
'bagging_fraction': 0.95,
'bagging_freq': 1,
'bagging_seed': 1,
'feature_fraction': 0.9,
'feature_fraction_seed': 1,
'max_bin': 128,
'max_depth': 10,
'num_rounds': 200,
'metric' : 'auc',
}
bst = lgb.train(params, train_data, 100, valid_sets=[val_data])
predictions+=bst.predict(test.drop(['id'],axis=1))
print('cur fold finished.')
submission = pd.DataFrame({'id': test_id, 'target': predictions})
submission.to_csv(SUBMISSION_FILENAME.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S')),index=False)
# -
def param_tune_with_val(params, tune_param, param_list, data_list, val_data, less_prefered = False):
#data_list = {'train':{'x':train_d,'y':train_y}, 'validation':{'x':valid_d,'y':valid_y}}
best_metric = (less_prefered and sys.float_info.max or -sys.float_info.max)
best_param = param_list[0]
for par_value in param_list:
params[tune_param] = par_value
# , num_boost_round=params['num_boost_round'], early_stopping_rounds = params['early_stopping_rounds']
model = lgb.train(params, data_list['train']['x'], valid_sets=[data_list['validation']['x']], \
feature_name='auto', #categorical_feature=['source_system_tab', 'source_screen_name', 'source_type', 'city', 'gender',\
# 'bd', 'name', 'artist_name', 'composer', 'lyricist', 'msno', 'song_id', 'genre_ids',\
# 'country', 'language', 'registered_via'],、
)
val_predprob = model.predict(val_data)
auroc_score = metrics.roc_auc_score(data_list['validation']['y'], val_predprob)
if (not less_prefered and auroc_score > best_metric) or (less_prefered and auroc_score < best_metric):
best_metric = auroc_score
best_param = par_value
log.info('best param for {}: {}, metric: {}'.format(tune_param, best_param, best_metric))
return best_param
# +
#{'top_k': 20, 'feature_fraction': 0.8, 'bagging_freq': 1, 'min_data_in_bin': 3, 'min_sum_hessian_in_leaf': 0.001, 'bagging_fraction': 0.9, 'max_depth': 12, 'num_leaves': 100, 'learning_rate': 0.01, 'objective': 'binary', 'lambda_l2': 0.01, 'feature_fraction_seed': 1024, 'min_data_in_leaf': 15, 'max_bin': 100, 'verbose': 0, 'bagging_seed': 6666, 'max_cat_to_onehot': 4, 'metric': 'auc', 'lambda_l1': 1e-05, 'num_threads': 16, 'boosting': 'gbdt', 'min_split_gain': 0.3}
#{'bagging_seed': 6666, 'lambda_l1': 1e-05, 'lambda_l2': 0.01, 'metric': 'auc', 'bagging_freq': 1, 'min_sum_hessian_in_leaf': 0.001, 'feature_fraction': 0.8, 'feature_fraction_seed': 1024, 'num_leaves': 90, 'boosting': 'gbdt', 'verbose': 0, 'min_data_in_leaf': 15, 'top_k': 20, 'objective': 'binary', 'min_data_in_bin': 3, 'num_threads': 16, 'max_cat_to_onehot': 4, 'max_depth': 10, 'bagging_fraction': 0.9, 'learning_rate': 0.01, 'max_bin': 80, 'min_split_gain': 0.3}
# +
def search_for_best_params(train, validation, test):
X_train = lgb.Dataset(np.array(train.drop(['target'], axis=1)), label=train['target'].values)
X_valid = lgb.Dataset(np.array(validation.drop(['target'], axis=1)), label=validation['target'].values)
y_train = train['target'].values
y_valid = validation['target'].values
X_test = np.array(test.drop(['id'], axis=1))
data_list = {'train':{'x':X_train,'y':y_train}, 'validation':{'x':X_valid,'y':y_valid}}
######## for value rather than catogory ################
# params_to_eval = OrderedDict(
# (
# ('num_boost_round', range(120,150,10)),
# ('num_leaves', range(80,100,10)), # number of leaves in one tree
# ('max_depth', range(8,12,1)),
# ('min_data_in_leaf', 15),
# ('min_sum_hessian_in_leaf', [0.001]),# too high will lead to under-fitting
# ('min_split_gain',[0.3]),# the minimum loss reduction required to make a split
# ('bagging_fraction',[0.9]),# [i/10.0 for i in range(6,10)]
# ('feature_fraction',[0.8]),# typical: 0.5-1
# ('max_bin', range(70,90,10)),
# ('lambda_l2',[0.01]),
# ('lambda_l1',[1e-5]),
# ('learning_rate',[0.01]), # typical: 0.01-0.2
# )
# )
# initial_params = {
# 'objective': 'binary',
# 'boosting': 'gbdt',
# 'num_boost_round': 140,
# 'learning_rate': 0.01 ,
# 'verbose': 0,
# 'num_leaves': 90,
# 'num_threads':16,
# 'max_depth': 9,
# 'min_data_in_leaf': 15, #minimal number of data in one leaf. Can be used to deal with over-fitting
# 'min_sum_hessian_in_leaf': 1e-3, #minimal sum hessian in one leaf. Like min_data_in_leaf, it can be used to deal with over-fitting
# 'feature_fraction': 0.8, #colsample_bytree
# 'feature_fraction_seed': 1024,
# 'bagging_fraction': 0.9, #subsample
# 'bagging_freq': 1, #frequency for bagging, 0 means disable bagging. k means will perform bagging at every k iteration
# 'bagging_seed': 6666,
# 'early_stopping_rounds':10,
# 'lambda_l1': 1e-5, #L1 regularization
# 'lambda_l2': 0.01, #L2 regularization
# 'max_cat_to_onehot': 4, #when number of categories of one feature smaller than or equal to max_cat_to_onehot, one-vs-other split algorithm will be used
# 'top_k': 20, #set this to larger value for more accurate result, but it will slow down the training speed
# 'min_split_gain': 0.3, #the minimal gain to perform split
# 'max_bin': 70, #max number of bins that feature values will be bucketed in. Small number of bins may reduce training accuracy but may increase general power (deal with over-fitting)
# 'min_data_in_bin': 3, #min number of data inside one bin, use this to avoid one-data-one-bin (may over-fitting)
# 'metric' : 'auc',
# }
params_to_eval = OrderedDict(
(
('num_boost_round', range(100,400,50)),
('num_leaves', range(80,160,10)), # number of leaves in one tree
('max_depth', range(8,18,1)),
('min_data_in_leaf', range(10,18,2)),
('min_sum_hessian_in_leaf', [0.001]),# too high will lead to under-fitting
('min_split_gain',[0.3]),# the minimum loss reduction required to make a split
('bagging_fraction',[0.9]),# [i/10.0 for i in range(6,10)]
('feature_fraction',[0.8]),# typical: 0.5-1
('max_bin', range(80,200,10)),
('lambda_l2',[0.01]),
('lambda_l1',[1e-5]),
('learning_rate',[0.01]), # typical: 0.01-0.2
)
)
initial_params = {
'objective': 'binary',
'boosting': 'gbdt',
'num_boost_round': 200,
'learning_rate': 0.1 ,
'verbose': 0,
'num_leaves': 120,
'num_threads':16,
'max_depth': 14,
'min_data_in_leaf': 16, #minimal number of data in one leaf. Can be used to deal with over-fitting
'min_sum_hessian_in_leaf': 1e-3, #minimal sum hessian in one leaf. Like min_data_in_leaf, it can be used to deal with over-fitting
'feature_fraction': 0.8, #colsample_bytree
'feature_fraction_seed': 1024,
'bagging_fraction': 0.9, #subsample
'bagging_freq': 1, #frequency for bagging, 0 means disable bagging. k means will perform bagging at every k iteration
'bagging_seed': 6666,
'early_stopping_rounds':10,
'lambda_l1': 1e-5, #L1 regularization
'lambda_l2': 0.01, #L2 regularization
'max_cat_to_onehot': 4, #when number of categories of one feature smaller than or equal to max_cat_to_onehot, one-vs-other split algorithm will be used
'top_k': 20, #set this to larger value for more accurate result, but it will slow down the training speed
'min_split_gain': 0.3, #the minimal gain to perform split
'max_bin': 140, #max number of bins that feature values will be bucketed in. Small number of bins may reduce training accuracy but may increase general power (deal with over-fitting)
'min_data_in_bin': 3, #min number of data inside one bin, use this to avoid one-data-one-bin (may over-fitting)
'metric' : 'auc',
}
# only param nin this list are tuned, total list are ['n_estimators', 'reg_alpha', 'reg_lambda', 'subsample', 'colsample_bytree', 'min_child_weight', 'max_depth', 'learning_rate', 'gamma']
#tuned_param_name = ['num_boost_round', 'num_leaves', 'max_depth', 'max_bin']
tuned_param_name = ['num_boost_round', 'num_leaves', 'max_depth', 'min_data_in_leaf', 'min_sum_hessian_in_leaf',\
'min_split_gain', 'bagging_fraction', 'feature_fraction', 'max_bin', 'lambda_l2', 'lambda_l1', 'learning_rate']
for par_name, par_list in params_to_eval.items():
if par_name in tuned_param_name:
log.info('tunning {}...'.format(par_name))
if len(par_list) > 1:
initial_params[par_name] = param_tune_with_val(initial_params, par_name, par_list, data_list, np.array(validation.drop(['target'], axis=1)))
else:
initial_params[par_name] = par_list[0]
return initial_params
# -
start_time = time.time()
best_param = search_for_best_params(train_use, validation_use, test)
log.info(best_param)
time_elapsed = time.time() - start_time
log.info('time used: {:.3f}sec'.format(time_elapsed))
# +
params = {
'objective': 'binary',
'boosting': 'gbdt',
'num_boost_round': 140,
'learning_rate': 0.01 ,
'verbose': 0,
'num_leaves': 90,
'num_threads':16,
'max_depth': 9,
'min_data_in_leaf': 15, #minimal number of data in one leaf. Can be used to deal with over-fitting
'min_sum_hessian_in_leaf': 1e-3, #minimal sum hessian in one leaf. Like min_data_in_leaf, it can be used to deal with over-fitting
'feature_fraction': 0.8, #colsample_bytree
'feature_fraction_seed': 1024,
'bagging_fraction': 0.9, #subsample
'bagging_freq': 1, #frequency for bagging, 0 means disable bagging. k means will perform bagging at every k iteration
'bagging_seed': 6666,
'early_stopping_rounds':10,
'lambda_l1': 1e-5, #L1 regularization
'lambda_l2': 0.01, #L2 regularization
'max_cat_to_onehot': 4, #when number of categories of one feature smaller than or equal to max_cat_to_onehot, one-vs-other split algorithm will be used
'top_k': 20, #set this to larger value for more accurate result, but it will slow down the training speed
'min_split_gain': 0.3, #the minimal gain to perform split
'max_bin': 70, #max number of bins that feature values will be bucketed in. Small number of bins may reduce training accuracy but may increase general power (deal with over-fitting)
'min_data_in_bin': 3, #min number of data inside one bin, use this to avoid one-data-one-bin (may over-fitting)
'metric' : 'auc',
}
X_train = lgb.Dataset(np.array(train_use.drop(['target'], axis=1)), label=train_use['target'].values)
X_valid = lgb.Dataset(np.array(validation_use.drop(['target'], axis=1)), label=validation_use['target'].values)
X_test = np.array(test.drop(['id'], axis=1))
model = lgb.train(params, X_train, valid_sets=[X_valid])
pred = model.predict(X_test)
submission = pd.DataFrame({'id': test_id, 'target': pred})
submission.to_csv(SUBMISSION_FILENAME.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S')),index=False)
# +
X_train = np.array(train_use.drop(['target'], axis=1))
y_train = train_use['target'].values
X_valid = np.array(validation_use.drop(['target'], axis=1))
y_valid = validation_use['target'].values
X_test = np.array(test.drop(['id'], axis=1))
# d_train = xgb.DMatrix(X_train)
# d_valid = xgb.DMatrix(X_valid)
# d_test = xgb.DMatrix(X_test)
data_list = {'train':{'x':X_train,'y':y_train}, 'validation':{'x':X_valid,'y':y_valid}}
# Train model, evaluate and make predictions
params={
'n_estimators':500,
'objective': 'binary:logistic',
'learning_rate': 0.75,
'gamma':0.1,
'subsample':0.8,
'colsample_bytree':0.3,
'min_child_weight':3,
'max_depth':16,
'seed':1024,
}
param_tune_with_val(params, 'max_depth', [5,1,6], data_list, 'auc', 20)
# model = xgb.train(params, d_train, 100, watchlist, early_stopping_rounds=20, \
# maximize=True, verbose_eval=5)
# +
X_train = np.array(train_use.drop(['target'], axis=1))
y_train = train_use['target'].values
X_valid = np.array(validation_use.drop(['target'], axis=1))
y_valid = validation_use['target'].values
X_test = np.array(test.drop(['id'], axis=1))
d_train = xgb.DMatrix(X_train, label=y_train)
d_valid = xgb.DMatrix(X_valid, label=y_valid)
d_test = xgb.DMatrix(X_test)
watchlist = [(d_train, 'train'), (d_valid, 'valid')]
# Train model, evaluate and make predictions
params = {}
params['objective'] = 'binary:logistic'
params['eta'] = 0.75
params['max_depth'] = 16
params['silent'] = 1
params['eval_metric'] = 'auc'
model = xgb.train(params, d_train, 100, watchlist, early_stopping_rounds=20, \
maximize=True, verbose_eval=5)
#Predict training set:
train_predictions = model.predict(X_train)
train_predprob = model.predict_proba(X_train)[:,1]
val_predictions = model.predict(X_valid)
val_predprob = model.predict_proba(X_valid)[:,1]
#Print model report:
print("\nModel Report")
print("Train Accuracy : %.4g" % metrics.accuracy_score(y_train, train_predictions))
print("Train AUC Score (Train): %f" % metrics.roc_auc_score(y_train, train_predprob))
print("ValAccuracy : %.4g" % metrics.accuracy_score(y_valid, val_predictions))
print("Validation AUC Score (Train): %f" % metrics.roc_auc_score(y_valid, val_predprob))
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
p_test = model.predict(d_test)
# -
xgb1 = XGBClassifier(
learning_rate =0.1,
n_estimators=1000,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb1, train_use.drop(['target'],axis=1), train_use['target'], validation_use.drop(['target'],axis=1), validation_use['target'])
def modelfit(alg, dtrain, predictors, useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(dtrain[predictors].values, label=dtrain[target].values)
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds,
metrics='auc', early_stopping_rounds=early_stopping_rounds, show_progress=False)
alg.set_params(n_estimators=cvresult.shape[0])
#Fit the algorithm on the data
alg.fit(dtrain[predictors], dtrain['Disbursed'], eval_metric='auc')
#Predict training set:
dtrain_predictions = alg.predict(dtrain[predictors])
dtrain_predprob = alg.predict_proba(dtrain[predictors])[:,1]
#Print model report:
print("\nModel Report")
print("Accuracy : %.4g" % metrics.accuracy_score(dtrain['Disbursed'].values, dtrain_predictions))
print("AUC Score (Train): %f" % metrics.roc_auc_score(dtrain['Disbursed'], dtrain_predprob))
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
def modelfit(alg, train, label, validation, val_label, useTrainCV=True, cv_folds=5, early_stopping_rounds=50):
if useTrainCV:
xgb_param = alg.get_xgb_params()
xgtrain = xgb.DMatrix(train.values, label=label.values)
cvresult = xgb.cv(xgb_param, xgtrain, num_boost_round=alg.get_params()['n_estimators'], nfold=cv_folds, metrics='auc', early_stopping_rounds=early_stopping_rounds, show_progress=False)
alg.set_params(n_estimators=cvresult.shape[0])
#Fit the algorithm on the data
alg.fit(train, label, eval_metric='auc')
#Predict training set:
train_predictions = alg.predict(train)
train_predprob = alg.predict_proba(train)[:,1]
val_predictions = alg.predict(validation)
val_predprob = alg.predict_proba(validation)[:,1]
#Print model report:
print("\nModel Report")
print("Train Accuracy : %.4g" % metrics.accuracy_score(label.values, train_predictions))
print("Train AUC Score (Train): %f" % metrics.roc_auc_score(label, train_predprob))
print("ValAccuracy : %.4g" % metrics.accuracy_score(val_label.values, val_predictions))
print("Validation AUC Score (Train): %f" % metrics.roc_auc_score(val_label, val_predprob))
feat_imp = pd.Series(alg.booster().get_fscore()).sort_values(ascending=False)
feat_imp.plot(kind='bar', title='Feature Importances')
plt.ylabel('Feature Importance Score')
xgb1 = XGBClassifier(
learning_rate =0.1,
n_estimators=1000,
max_depth=5,
min_child_weight=1,
gamma=0,
subsample=0.8,
colsample_bytree=0.8,
objective= 'binary:logistic',
nthread=4,
scale_pos_weight=1,
seed=27)
modelfit(xgb1, train_use.drop(['target'],axis=1), train_use['target'], validation_use.drop(['target'],axis=1), validation_use['target'])
# +
import lightgbm as lgb
predictions = np.zeros(shape=[len(test)])
train_data = lgb.Dataset(train_use.drop(['target'],axis=1), label=train_use['target'])
val_data = lgb.Dataset(validation_use.drop(['target'],axis=1), label=validation_use['target'])
params = {
'objective': 'binary',
'boosting': 'gbdt',
'learning_rate': 0.1 ,
'verbose': 0,
'num_leaves': 108,
'bagging_fraction': 0.95,
'bagging_freq': 1,
'bagging_seed': 1,
'feature_fraction': 0.9,
'feature_fraction_seed': 1,
'max_bin': 128,
'max_depth': 10,
'num_rounds': 200,
'metric' : 'auc',
}
bst = lgb.train(params, train_data, 100, valid_sets=[val_data])
predictions=bst.predict(test.drop(['id'],axis=1))
print('finished.')
predictions = predictions/3
submission = pd.DataFrame({'id': test_id, 'target': predictions})
submission.to_csv(SUBMISSION_FILENAME.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S')),index=False)
# +
import lightgbm as lgb
kf = KFold(n_splits=3)
predictions = np.zeros(shape=[len(test)])
for train_indices,val_indices in kf.split(train) :
train_data = lgb.Dataset(train.drop(['target'],axis=1).loc[train_indices,:],label=train.loc[train_indices,'target'])
val_data = lgb.Dataset(train.drop(['target'],axis=1).loc[val_indices,:],label=train.loc[val_indices,'target'])
params = {
'objective': 'binary',
'boosting': 'gbdt',
'learning_rate': 0.1 ,
'verbose': 0,
'num_leaves': 108,
'bagging_fraction': 0.95,
'bagging_freq': 1,
'bagging_seed': 1,
'feature_fraction': 0.9,
'feature_fraction_seed': 1,
'max_bin': 128,
'max_depth': 10,
'num_rounds': 200,
'metric' : 'auc',
}
bst = lgb.train(params, train_data, 100, valid_sets=[val_data])
predictions+=bst.predict(test.drop(['id'],axis=1))
print('cur fold finished.')
del bst
predictions = predictions/3
submission = pd.DataFrame({'id': test_id, 'target': predictions})
submission.to_csv(SUBMISSION_FILENAME.format(datetime.now().strftime('%Y-%m-%d %H:%M:%S')),index=False)
# +
# Preprocess songs data
songs_genres = np.array(songs['genre_ids']\
.apply(lambda x: [int(v) for v in str(x).split('|')]))
genres_list = songs_genres.ravel().unique()
print('Number of genres: ' + str(len(genres_list)))
ohe_genres = np.zeros((len(songs_genres), len(genres_list)))
for s_i, s_genres in enumerate(songs_genres):
for genre in s_genres:
g_i = genres_list.find(genre)
ohe_genres[s_i, g_i] = 1
for g_i, g in enumerate(genres_list):
songs['genre_' + str(g)] = ohe_genres[:, g_i]
print(songs.head())
songs = songs.drop(['genre_ids'], axis=1)
song_cols = songs.columns
# Preprocess dataset
train = train.fillna(-1)
test = test.fillna(-1)
cols = list(train.columns)
cols.remove('target')
for col in tqdm(cols):
if train[col].dtype == 'object':
train[col] = train[col].apply(str)
test[col] = test[col].apply(str)
le = LabelEncoder()
train_vals = list(train[col].unique())
test_vals = list(test[col].unique())
le.fit(train_vals + test_vals)
train[col] = le.transform(train[col])
test[col] = le.transform(test[col])
print(col + ': ' + str(len(train_vals)) + ', ' + str(len(test_vals)))
# +
########################################
## import packages
########################################
import datetime
import numpy as np
import pandas as pd
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import roc_auc_score
from keras.models import Model
from keras.layers import Dense, Input, Embedding, Dropout, Activation, Reshape
from keras.layers.merge import concatenate, dot
from keras.layers.normalization import BatchNormalization
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.regularizers import l2
from keras.initializers import RandomUniform
from keras.optimizers import RMSprop, Adam, SGD
########################################
## load the data
########################################
train = pd.read_csv('./data/train.csv')
uid = train.msno
sid = train.song_id
target = train.target
test = pd.read_csv('./data/test.csv')
id_test = test.id
uid_test = test.msno
sid_test = test.song_id
########################################
## encoding
########################################
usr_encoder = LabelEncoder()
usr_encoder.fit(uid.append(uid_test))
uid = usr_encoder.transform(uid)
uid_test = usr_encoder.transform(uid_test)
sid_encoder = LabelEncoder()
sid_encoder.fit(sid.append(sid_test))
sid = sid_encoder.transform(sid)
sid_test = sid_encoder.transform(sid_test)
u_cnt = int(max(uid.max(), uid_test.max()) + 1)
s_cnt = int(max(sid.max(), sid_test.max()) + 1)
########################################
## train-validation split
########################################
perm = np.random.permutation(len(train))
trn_cnt = int(len(train) * 0.85)
uid_trn = uid[perm[:trn_cnt]]
uid_val = uid[perm[trn_cnt:]]
sid_trn = sid[perm[:trn_cnt]]
sid_val = sid[perm[trn_cnt:]]
target_trn = target[perm[:trn_cnt]]
target_val = target[perm[trn_cnt:]]
########################################
## define the model
########################################
def get_model():
user_embeddings = Embedding(u_cnt,
64,
embeddings_initializer=RandomUniform(minval=-0.1, maxval=0.1),
embeddings_regularizer=l2(1e-4),
input_length=1,
trainable=True)
song_embeddings = Embedding(s_cnt,
64,
embeddings_initializer=RandomUniform(minval=-0.1, maxval=0.1),
embeddings_regularizer=l2(1e-4),
input_length=1,
trainable=True)
uid_input = Input(shape=(1,), dtype='int32')
embedded_usr = user_embeddings(uid_input)
embedded_usr = Reshape((64,))(embedded_usr)
sid_input = Input(shape=(1,), dtype='int32')
embedded_song = song_embeddings(sid_input)
embedded_song = Reshape((64,))(embedded_song)
preds = dot([embedded_usr, embedded_song], axes=1)
preds = concatenate([embedded_usr, embedded_song, preds])
preds = Dense(128, activation='relu')(preds)
preds = Dropout(0.5)(preds)
preds = Dense(1, activation='sigmoid')(preds)
model = Model(inputs=[uid_input, sid_input], outputs=preds)
opt = RMSprop(lr=1e-3)
model.compile(loss='binary_crossentropy', optimizer=opt, metrics=['acc'])
return model
########################################
## train the model
########################################
model = get_model()
early_stopping =EarlyStopping(monitor='val_acc', patience=5)
model_path = 'bst_model.h5'
model_checkpoint = ModelCheckpoint(model_path, save_best_only=True, \
save_weights_only=True)
hist = model.fit([uid_trn, sid_trn], target_trn, validation_data=([uid_val, sid_val], \
target_val), epochs=100, batch_size=32768, shuffle=True, \
callbacks=[early_stopping, model_checkpoint])
model.load_weights(model_path)
preds_val = model.predict([uid_val, sid_val], batch_size=32768)
val_auc = roc_auc_score(target_val, preds_val)
########################################
## make the submission
########################################
preds_test = model.predict([uid_test, sid_test], batch_size=32768, verbose=1)
sub = pd.DataFrame({'id': id_test, 'target': preds_test.ravel()})
sub.to_csv('./sub_%.5f.csv'%(val_auc), index=False)
# +
# Linear algebra:
import numpy as np
import pandas as pd
# Graphics:
import matplotlib.pyplot as plt
import seaborn as sns
# Frameworks:
import lightgbm as lgb # LightGBM
# Utils:
import gc # garbage collector
# %matplotlib inline
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
IDIR = '../input/' # main path
members = pd.read_csv(IDIR + 'members.csv')
songs = pd.read_csv(IDIR + 'songs.csv')
song_extra_info = pd.read_csv(IDIR + 'song_extra_info.csv')
train = pd.read_csv(IDIR + 'train.csv')
test = pd.read_csv(IDIR + 'test.csv')
# Adding songs' info:
train_aug1 = pd.merge(left=train, right=songs, on='song_id', how='left')
test_aug1 = pd.merge(left=test, right=songs, on='song_id', how='left')
# Adding extra info about songs:
train_aug2 = pd.merge(left=train_aug1, right=song_extra_info, on='song_id', how='left')
test_aug2 = pd.merge(left=test_aug1, right=song_extra_info, on='song_id', how='left')
del train_aug1, test_aug1
# Addind users' info:
train_aug3 = pd.merge(left=train_aug2, right=members, on='msno', how='left')
test_aug3 = pd.merge(left=test_aug2, right=members, on='msno', how='left')
del train_aug2, test_aug2
# Merging train and test data:
train_aug3.drop(['song_id'], axis=1, inplace=True)
train_aug3['set'] = 0
test_aug3.drop(['song_id'], axis=1, inplace=True)
test_aug3['set'] = 1
test_aug3['target'] = -1
all_aug = pd.concat([train_aug3, test_aug3], axis=0)
del train_aug3, test_aug3
gc.collect();
# source_system_tab/source_screen_name/source_type/genre_ids/artist_name/composer/lyricist/name/isrc/gender 用'NA'填补并one-hot编码
# genre_ids encoding:
all_aug['genre_ids'] = all_aug.genre_ids.fillna('NA')
all_aug['genre_ids'] = all_aug.genre_ids.astype(np.str)
genre_ids_le = LabelEncoder()
genre_ids_le.fit(all_aug.genre_ids)
all_aug['genre_ids'] = genre_ids_le.transform(all_aug.genre_ids).astype(np.int16)
# language encoding:
all_aug['language'] = all_aug.language.fillna(-2)
all_aug['language'] = all_aug.language.astype(np.int8)
# city encoding:
all_aug['city'] = all_aug.city.astype(np.int8)
# bd encoding:
all_aug['bd'] = all_aug.bd.astype(np.int16)
# registered_via encoding:
all_aug['registered_via'] = all_aug.registered_via.astype(np.int8)
# registration_init_time encoding:
all_aug['registration_init_time'] = all_aug.registration_init_time.astype(np.int32)
# expiration_date encoding:
all_aug['expiration_date'] = all_aug.expiration_date.astype(np.int32)
# Info:
all_aug.info(max_cols=0)
all_aug.head(2)
all_aug['exp_reg_time'] = all_aug.expiration_date - all_aug.registration_init_time
gc.collect();
d_train = lgb.Dataset(all_aug[all_aug.set == 0].drop(['target', 'msno', 'id', 'set'], axis=1),
label=all_aug[all_aug.set == 0].pop('target'))
ids_train = all_aug[all_aug.set == 0].pop('msno')
lgb_params = {
'learning_rate': 1.0,
'max_depth': 15,
'num_leaves': 250,
'objective': 'binary',
'metric': {'auc'},
'feature_fraction': 0.8,
'bagging_fraction': 0.75,
'bagging_freq': 5,
'max_bin': 100}
cv_result_lgb = lgb.cv(lgb_params,
d_train,
num_boost_round=5000,
nfold=3,
stratified=True,
early_stopping_rounds=50,
verbose_eval=100,
show_stdv=True)
num_boost_rounds_lgb = len(cv_result_lgb['auc-mean'])
print('num_boost_rounds_lgb=' + str(num_boost_rounds_lgb))
# %%time
ROUNDS = num_boost_rounds_lgb
print('light GBM train :-)')
bst = lgb.train(lgb_params, d_train, ROUNDS)
# lgb.plot_importance(bst, figsize=(9,20))
# del d_train
gc.collect()
plt.figure(figsize=(10,5))
plt.subplot(1,2,1)
feature_imp = pd.Series(dict(zip(d_train.feature_name,
bst.feature_importance()))).sort_values(ascending=False)
sns.barplot(x=feature_imp.values, y=feature_imp.index.values, orient='h', color='g')
plt.subplot(1,2,2)
train_scores = np.array(cv_result_lgb['auc-mean'])
train_stds = np.array(cv_result_lgb['auc-stdv'])
plt.plot(train_scores, color='green')
plt.fill_between(range(len(cv_result_lgb['auc-mean'])),
train_scores - train_stds, train_scores + train_stds,
alpha=0.1, color='green')
plt.title('LightGMB CV-results')
plt.show()
# -
|
MusicRecommendation/GBM-First-0.678.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Numpy example
#
# Numpy hands on examples to be familiar with numpy behavior.
# ## Creating numpy.ndarray
#
# numpy.ndarray can be created by following methods
# * array
# * asarray
# * empty
# * zeros
# * ones
# ## Attributes
#
# numpy has following attributes (showing only major attributes)
# * dtype
# * shape
# * ndim
# * size
# +
import numpy as np # numpy is often imported as np for short terminology
def print_numpy_attributes(a):
"""dumps ``numpy.ndarray`` attribute information
:param a:
:type a: np.ndarray
:return:
"""
print('----- numpy attributes info -----')
print('', a) # data will be printed. this is same with a.data
print('type', type(a)) # should be <class 'numpy.ndarray'>
print('data', a.data) # actual array data
print('dtype', a.dtype) # type of data (int, float32 etc)
print('shape', a.shape) # dimensional information of data (2, 3) etc.
print('ndim', a.ndim) # total dimension of shape. 0 means scalar, 1 is vector, 2 is matrix...
print('size', a.size) # total size of data, which is product sum of shape
print('---------------------------------')
# -
# 1. creating scalar
a1 = np.array(3) # note that np.array([3]) will create vector with 1 element
print_numpy_attributes(a1)
# 2. creating vector
l2 = [1, 2, 3]
a2 = np.array(l2)
print(l2, type(l2)) # l2 is list
print_numpy_attributes(a2) # a2 is numpy.ndarray
# 3. creating matrix
l3 = [[1, 2, 3], [4, 5, 6]]
a3 = np.array(l3)
# print(l3, type(l3))
print_numpy_attributes(a3)
# 4. creating general multi-dimensional array (tensor)
a4 = np.array([
[[1, 2, 3], [4, 5, 6]],
[[11, 22, 33], [44, 55, 66]]
]
)
print_numpy_attributes(a4)
|
src/00_preparation/numpy_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import scipy as sp
import scipy.stats
import matplotlib.pyplot as plt
# # Lab 1
#
# When reporting probabilities in a scientific context, the standard is to convert this probabilty into a sigma value, which represents the probabilty of the data To convert sigma values back to a probabilty, some convoluted integration is required. Luckily, python has these capabilites built in. The *erfc* function is one of several scipy functions which returns a probabilty given an input sigma value.
for x in range(1, 6):
print(sp.special.erfc(x))
# Here we can see that *erfc* values roughly match up with the negative 1, 2, 3, and 4 sigma values on a z table (or the positive values subtracted from 1), giving us the probabilty that our target event happened outside of our measured sigma value. As previously mentioned, the accepted standard is to convert this probabilty into a sigma value, but a probabilty can also be converted back into a sigma, as is shown below.
sp.stats.norm.ppf(3e-7)
# Here we see the sigma value reported as a negative number. This is probably due to the fact that the built in function uses the left side of the normal distribution to find the associated sigma value as this would come up first when searching from negative to positive.
# ### Rayleigh Distribution
# A Rayliegh distribution is distinct in that it is not identical on each side of its peak. Applications of Rayleigh distributions are most common in places where long-lived or large events are less common than those of shorter length. Examples are wave height or product lifespan. Let's create a sample set of data with built-in Python functions.
# +
#def prob_graph(loc, scale, xleft, xright, size)
d = sp.stats.rayleigh.rvs(loc = 2.0, scale = 1, size = 100000)
xleft = 1.95
xright = 7
fig, ax = plt.subplots(1, 1)
ax.hist(d,50, density=True)
plt.tick_params(labelsize = 20)
plt.xlim(xleft, xright)
x = np.linspace(xleft,xright,1000)
ax.plot(x, sp.stats.rayleigh.pdf(x,loc = 2, scale = 1),linewidth = 7,alpha = 0.6)
plt.show()
# -
# Looks pretty good! The 100,000 sample size seems to have created a pretty accurate distrobution. However, towards the top end (~X = 6), we can't really tell what is going on. The height of the distribution is controlled by the 'scale' factor in the pdf, with a higher scale representing a wider and shorter distribution. Plotting our data on a semilog graph reveals some interesting secrets.
# +
#def prob_graph(loc, scale, xleft, xright, size)
d = sp.stats.rayleigh.rvs(loc = 2.0, scale = 1, size = 100000)
xleft = 1.95
xright = 7
fig, ax = plt.subplots(1, 1)
ax.set_yscale('log')
ax.hist(d,50, density=True)
plt.tick_params(labelsize = 20)
plt.xlim([xleft, xright])
x = np.linspace(xleft,xright,1000)
ax.plot(x,sp.stats.rayleigh.pdf(x,loc = 2, scale = 1),linewidth = 7,alpha = 0.6)
plt.show()
# -
# Even with the large sample size, there is a suprisingly large amount of innacuracy towards the tail of the distribution.
# The theoretical data above could represent the lifetime of a company's product in years. If a similar, slightly redesigned product has a lifespan of 4.5 years, what is the chance that this is not an improvement over our original product?
sp.stats.rayleigh.cdf(4.5)
sp.stats.norm.ppf(0.999959934702607)
# This comes out to be a sigma value of 3.94, which while very significant, would not be accepted by the scientific community in a physics-related context.
# ### Binomial Distribution
# +
fig, ax = plt.subplots(1,1)
n, p = 100, .45
x = np.arange(sp.stats.binom.ppf(0.01, n, p), sp.stats.binom.ppf(0.99, n, p))
plt.xlim(30, 60)
ax.plot(x, sp.stats.binom.pmf(x, n, p), 'o')
ax.vlines(x, 0, sp.stats.binom.pmf(x, n, p))
# +
fig, ax = plt.subplots(1,1)
n, p = 100, .45
x = np.arange(sp.stats.binom.ppf(0.01, n, p), sp.stats.binom.ppf(0.99, n, p))
plt.xlim(30, 60)
ax.plot(x, sp.stats.binom.pmf(x, n, p), 'o')
ax.set_yscale('log')
ax.vlines(x, 0, sp.stats.binom.pmf(x, n, p))
# -
# In a semilog plot, the distribution takes on the shape of a slightly skewed parabola, looking very similar to, but slightly different from a Gaussian curve.
# Using the distrubition above, let's assume we flip a coin that is slightly biased to one side. We'd assume most outcomes would land around 45 on one side to 55 on the other, which is reflected in the graph. One difference in comparison to our previous question, which dealt with a continuous probabilty, is that our probabilty only takes on integer values, which makes the binomial distribution good for counting events if we know the general probability that it should happen. Unlike individual data points, statistics about the binomial distribution don't have to necessarily be an integer value. If the average family has 1.9 kids, that clearly does not mean that any familiy has that exact value.
# So what happens if we get 60 heads on a second coin with unknown properties? Could it be the same type of coin?
sp.stats.binom.cdf(60, n, p)
sp.stats.norm.ppf(0.9990617681011207)
# Most likely, this coin is different, but there wouldn't be enough results here to publish it in a paper.
|
labs/lab1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Pyspark 2
# language: python
# name: pyspark2
# ---
# ## Overview of Windowing Functions
#
# Let us get an overview of Windowing Functions using Spark.
# * These are available as part of SQL in most of the traditional databases.
# * In some databases they are also known as Analytical Functions.
# Let us start spark context for this Notebook so that we can execute the code provided. You can sign up for our [10 node state of the art cluster/labs](https://labs.itversity.com/plans) to learn Spark SQL using our unique integrated LMS.
# +
from pyspark.sql import SparkSession
import getpass
username = getpass.getuser()
spark = SparkSession. \
builder. \
config('spark.ui.port', '0'). \
config("spark.sql.warehouse.dir", f"/user/{username}/warehouse"). \
enableHiveSupport(). \
appName(f'{username} | Python - Windowing Functions'). \
master('yarn'). \
getOrCreate()
# -
# If you are going to use CLIs, you can use Spark SQL using one of the 3 approaches.
#
# **Using Spark SQL**
#
# ```
# spark2-sql \
# --master yarn \
# --conf spark.ui.port=0 \
# --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
# ```
#
# **Using Scala**
#
# ```
# spark2-shell \
# --master yarn \
# --conf spark.ui.port=0 \
# --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
# ```
#
# **Using Pyspark**
#
# ```
# pyspark2 \
# --master yarn \
# --conf spark.ui.port=0 \
# --conf spark.sql.warehouse.dir=/user/${USER}/warehouse
# ```
# * First let us understand relevance of these functions using `employees` data set.
employeesPath = '/public/hr_db/employees'
employees = spark. \
read. \
format('csv'). \
option('sep', '\t'). \
schema('''employee_id INT,
first_name STRING,
last_name STRING,
email STRING,
phone_number STRING,
hire_date STRING,
job_id STRING,
salary FLOAT,
commission_pct STRING,
manager_id STRING,
department_id STRING
'''). \
load(employeesPath)
employees.show()
employees.printSchema()
employees.count()
from pyspark.sql.functions import col
employees. \
select('employee_id',
col('department_id').cast('int').alias('department_id'),
'salary'
). \
orderBy('department_id', 'salary'). \
show()
# * Let us say we want to compare individual salary with department wise salary expense.
# * Here is one of the approach which require self join.
# * Compute department wise expense usig `groupBy` and `agg`.
# * Join with **employees** again on department_id.
from pyspark.sql.functions import sum, col
department_expense = employees. \
groupBy('department_id'). \
agg(sum('salary').alias('expense'))
department_expense.show()
employees. \
select('employee_id', 'department_id', 'salary'). \
join(department_expense, employees.department_id == department_expense.department_id). \
orderBy(employees.department_id, col('salary')). \
show()
# **However, using this approach is not very efficient and also overly complicated. Windowing functions actually simplify the logic and also runs efficiently**
#
# Now let us get into the details related to Windowing functions.
# * Main package `pyspark.sql.window`
# * It has classes such as `Window` and `WindowSpec`
# * `Window` have APIs such as `partitionBy`, `orderBy` etc
# * These APIs (such as `partitionBy`) return `WindowSpec` object. We can pass `WindowSpec` object to over on functions such as `rank()`, `dense_rank()`, `sum()` etc
# * Syntax: `sum().over(spec)` where `spec = Window.partitionBy('ColumnName')`
from pyspark.sql import window
help(window)
# | Functions | API or Function |
# | ------------- |:-------------:|
# | Aggregate Functions | <ul><li>sum</li><li>avg</li><li>min</li><li>max</li></ul> |
# | Ranking Functions | <ul><li>rank</li><li>dense_rank</li></ul><ul><li>percent_rank</li><li>row_number</li> <li>ntile</li></ul> |
# | Analytic Functions | <ul><li>cume_dist</li><li>first</li><li>last</li><li>lead</li> <li>lag</li></ul> |
|
07_windowing_functions/02_overview_of_windowing_functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/SathyaSudha-96/Erdos/blob/main/4L_Cluster_Monet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="M3tLJIwyBBaT" outputId="ce542149-44e2-46a2-fad7-0a6293a716d7"
# !pip install -q condacolab
import condacolab
condacolab.install_anaconda()
# + colab={"base_uri": "https://localhost:8080/"} id="E6fQlgyMBalD" outputId="f469d009-e871-4204-c3f4-74a6cbdf4671"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="s4BJubFEBbaM" outputId="3c9fb9ce-8342-428b-ee5e-b9ca9a2dfb13"
# %pwd
# + colab={"base_uri": "https://localhost:8080/"} id="iwh8nSnQBdNY" outputId="7ea815a1-42e1-4c69-d1de-b72369d23c46"
# Execution time 4m 18s
# %cp -av /content/drive/MyDrive/'Colab Notebooks'/benchmarking-gnns /content
# + colab={"base_uri": "https://localhost:8080/"} id="Rt4YLBzgBg5V" outputId="7273d9c9-6a6e-4af9-dbf7-0f03310b835f"
# %cd benchmarking-gnns
# + colab={"base_uri": "https://localhost:8080/"} id="AzfO4LL9BjJO" outputId="2667030a-c3b6-43ea-f888-5f4bd402daa0"
# %cd /content/benchmarking-gnns/
# + colab={"base_uri": "https://localhost:8080/"} id="Y8QzCUXHBlw4" outputId="1b7e1298-b554-4a9d-d221-78db554fdc32"
# !conda env create -f /content/benchmarking-gnns/environment_gpu.yml
# + colab={"base_uri": "https://localhost:8080/"} id="bAbKB5d6Bo0U" outputId="32f5bb33-26f6-4df2-afd5-3865e9b3d935"
# !conda activate benchmark_gnn
# + colab={"base_uri": "https://localhost:8080/", "height": 728} id="6LAj-ZWEBtCn" outputId="0d01824b-904b-48b4-91d3-0c2ebe333fb7"
# !pip install dgl-cu101
# #!pip install tensorboardX
# !pip install pytorch
# #!pip install dgl
# !pip install tensorboardX
# + colab={"base_uri": "https://localhost:8080/"} id="PaaU1_QiBxoU" outputId="7842400d-caf9-4635-e308-f0b10f9d0014"
import dgl
import numpy as np
import os
import socket
import time
import random
import glob
import argparse, json
import pickle
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import DataLoader
from tensorboardX import SummaryWriter
from tqdm import tqdm
class DotDict(dict):
def __init__(self, **kwds):
self.update(kwds)
self.__dict__ = self
# + colab={"base_uri": "https://localhost:8080/"} id="SXEV1sqwByX8" outputId="783aa365-2312-4ed6-e2db-f619f804bc51"
# """
# AUTORELOAD IPYTHON EXTENSION FOR RELOADING IMPORTED MODULES
# """
def in_ipynb():
try:
cfg = get_ipython().config
return True
except NameError:
return False
notebook_mode = in_ipynb()
print(notebook_mode)
if notebook_mode == True:
# %load_ext autoreload
# %autoreload 2
# + id="zUa6_u4XB2Ev"
"""
IMPORTING CUSTOM MODULES/METHODS
"""
from nets.SBMs_node_classification.load_net import gnn_model # import GNNs
from data.data import LoadData # import dataset
# + id="BuPu68YFCTRp"
"""
GPU Setup
"""
def gpu_setup(use_gpu, gpu_id):
os.environ["CUDA_DEVICE_ORDER"] = "PCI_BUS_ID"
os.environ["CUDA_VISIBLE_DEVICES"] = str(gpu_id)
if torch.cuda.is_available() and use_gpu:
print('cuda available with GPU:',torch.cuda.get_device_name(0))
device = torch.device("cuda")
else:
print('cuda not available')
device = torch.device("cpu")
return device
# select GPU or CPU
use_gpu = True; gpu_id = 0; device = None # default GPU
#use_gpu = False; gpu_id = -1; device = None # CPU
# + colab={"base_uri": "https://localhost:8080/"} id="CI-4Yxu2CjvU" outputId="dd0856d9-a849-45bb-d7de-3e5d09de81d1"
# """
# USER CONTROLS
# """
if notebook_mode == True:
#MODEL_NAME = '3WLGNN'
#MODEL_NAME = 'RingGNN'
#MODEL_NAME = 'GatedGCN'
#MODEL_NAME = 'GCN'
#MODEL_NAME = 'GAT'
#MODEL_NAME = 'GraphSage'
#MODEL_NAME = 'MLP'
#MODEL_NAME = 'GIN'
MODEL_NAME = 'MoNet'
DATASET_NAME = 'SBM_CLUSTER'
#DATASET_NAME = 'SBM_PATTERN'
out_dir = 'out/SBMs_node_classification/'
root_log_dir = out_dir + 'logs/' + MODEL_NAME + "_" + DATASET_NAME + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
root_ckpt_dir = out_dir + 'checkpoints/' + MODEL_NAME + "_" + DATASET_NAME + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
dataset = LoadData(DATASET_NAME)
trainset, valset, testset = dataset.train, dataset.val, dataset.test
# + id="xi_oQLmDCXTr"
#MODEL_NAME = 'RingGNN'
#MODEL_NAME = 'GatedGCN'
#MODEL_NAME = 'GCN'
#MODEL_NAME = 'GAT'
#MODEL_NAME = 'GraphSage'
#MODEL_NAME = 'MLP'
#MODEL_NAME = 'DiffPool'
#MODEL_NAME = 'GIN'
MODEL_NAME = 'MoNet'
# + id="ErnyyAXPJK3q"
# """
# PARAMETERS
# """
if notebook_mode == True:
n_heads = -1
edge_feat = False
pseudo_dim_MoNet = -1
kernel = -1
gnn_per_block = -1
embedding_dim = -1
pool_ratio = -1
n_mlp_GIN = -1
gated = False
self_loop = False
#self_loop = True
max_time = 12
pos_enc = True
#pos_enc = False
pos_enc_dim = 10
if MODEL_NAME == 'GatedGCN':
seed=41; epochs=1000; batch_size=5; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=70; out_dim=hidden_dim; dropout=0.0; readout='mean'
if MODEL_NAME == 'GCN':
seed=41; epochs=1000; batch_size=5; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=146; out_dim=hidden_dim; dropout=0.0; readout='mean'
if MODEL_NAME == 'GAT':
seed=41; epochs=1000; batch_size=50; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; n_heads=8; hidden_dim=19; out_dim=n_heads*hidden_dim; dropout=0.0; readout='mean'
print('True hidden dim:',out_dim)
if MODEL_NAME == 'GraphSage':
seed=41; epochs=1000; batch_size=50; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=108; out_dim=hidden_dim; dropout=0.0; readout='mean'
if MODEL_NAME == 'MLP':
seed=41; epochs=1000; batch_size=50; init_lr=5e-4; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
gated=False; # MEAN
L=4; hidden_dim=150; out_dim=hidden_dim; dropout=0.0; readout='mean'
gated=True; # GATED
L=4; hidden_dim=135; out_dim=hidden_dim; dropout=0.0; readout='mean'
if MODEL_NAME == 'DiffPool':
seed=41; epochs=1000; batch_size=50; init_lr=5e-4; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=32; out_dim=hidden_dim; dropout=0.0; readout='mean'
n_heads=8; gnn_per_block=3; embedding_dim=32; batch_size=128; pool_ratio=0.15
if MODEL_NAME == 'GIN':
seed=41; epochs=1000; batch_size=50; init_lr=5e-4; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=110; out_dim=hidden_dim; dropout=0.0; readout='mean'
n_mlp_GIN = 2; learn_eps_GIN=True; neighbor_aggr_GIN='sum'
if MODEL_NAME == 'MoNet':
seed=41; epochs=120; batch_size=50; init_lr=5e-4; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=90; out_dim=hidden_dim; dropout=0.0; readout='mean'
pseudo_dim_MoNet=2; kernel=3;
if MODEL_NAME == 'RingGNN':
seed=41; epochs=1000; batch_size=1; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
#L=4; hidden_dim=145; out_dim=hidden_dim; dropout=0.0; readout='mean'
L=4; hidden_dim=25; out_dim=hidden_dim; dropout=0.0
if MODEL_NAME == '3WLGNN':
seed=41; epochs=1000; batch_size=1; init_lr=5e-5; lr_reduce_factor=0.5; lr_schedule_patience=25; min_lr = 1e-6; weight_decay=0
L=4; hidden_dim=82; out_dim=hidden_dim; dropout=0.0
# generic new_params
net_params = {}
net_params['device'] = device
net_params['in_dim'] = torch.unique(trainset[0][0].ndata['feat'],dim=0).size(0) # node_dim (feat is an integer)
net_params['hidden_dim'] = hidden_dim
net_params['out_dim'] = out_dim
num_classes = torch.unique(trainset[0][1],dim=0).size(0)
net_params['n_classes'] = num_classes
net_params['L'] = L # min L should be 2
net_params['readout'] = "mean"
net_params['layer_norm'] = True
net_params['batch_norm'] = True
net_params['in_feat_dropout'] = 0.0
net_params['dropout'] = 0.0
net_params['residual'] = True
net_params['edge_feat'] = edge_feat
net_params['self_loop'] = self_loop
# for MLPNet
net_params['gated'] = gated
# for GAT
net_params['n_heads'] = n_heads
# for graphsage
net_params['sage_aggregator'] = 'meanpool'
# specific for GIN
net_params['n_mlp_GIN'] = n_mlp_GIN
net_params['learn_eps_GIN'] = True
net_params['neighbor_aggr_GIN'] = 'sum'
# specific for MoNet
net_params['pseudo_dim_MoNet'] = pseudo_dim_MoNet
net_params['kernel'] = kernel
# specific for RingGNN
net_params['radius'] = 2
num_nodes = [dataset.train[i][0].number_of_nodes() for i in range(len(dataset.train))]
net_params['avg_node_num'] = int(np.ceil(np.mean(num_nodes)))
# specific for 3WLGNN
net_params['depth_of_mlp'] = 2
# specific for pos_enc_dim
net_params['pos_enc'] = pos_enc
net_params['pos_enc_dim'] = pos_enc_dim
# + colab={"base_uri": "https://localhost:8080/"} id="a9Fb7bhUC2YB" outputId="0220bb8d-8e12-4119-c4df-aa931258ebb0"
"""
VIEWING MODEL CONFIG AND PARAMS
"""
def view_model_param(MODEL_NAME, net_params):
model = gnn_model(MODEL_NAME, net_params)
total_param = 0
print("MODEL DETAILS:\n")
print(model)
for param in model.parameters():
# print(param.data.size())
total_param += np.prod(list(param.data.size()))
print('MODEL/Total parameters:', MODEL_NAME, total_param)
return total_param
if notebook_mode == True:
view_model_param(MODEL_NAME, net_params)
# + id="sdg5T2gMJK3t"
"""
TRAINING CODE
"""
def train_val_pipeline(MODEL_NAME, dataset, params, net_params, dirs):
start0 = time.time()
per_epoch_time = []
DATASET_NAME = dataset.name
if MODEL_NAME in ['GCN', 'GAT']:
if net_params['self_loop']:
print("[!] Adding graph self-loops for GCN/GAT models (central node trick).")
dataset._add_self_loops()
if MODEL_NAME in ['GatedGCN']:
if net_params['pos_enc']:
print("[!] Adding graph positional encoding.")
dataset._add_positional_encodings(net_params['pos_enc_dim'])
print('Time PE:',time.time()-start0)
trainset, valset, testset = dataset.train, dataset.val, dataset.test
root_log_dir, root_ckpt_dir, write_file_name, write_config_file = dirs
device = net_params['device']
# Write network and optimization hyper-parameters in folder config/
with open(write_config_file + '.txt', 'w') as f:
f.write("""Dataset: {},\nModel: {}\n\nparams={}\n\nnet_params={}\n\n\nTotal Parameters: {}\n\n"""\
.format(DATASET_NAME, MODEL_NAME, params, net_params, net_params['total_param']))
log_dir = os.path.join(root_log_dir, "RUN_" + str(0))
writer = SummaryWriter(log_dir=log_dir)
# setting seeds
random.seed(params['seed'])
np.random.seed(params['seed'])
torch.manual_seed(params['seed'])
if device.type == 'cuda':
torch.cuda.manual_seed(params['seed'])
print("Training Graphs: ", len(trainset))
print("Validation Graphs: ", len(valset))
print("Test Graphs: ", len(testset))
print("Number of Classes: ", net_params['n_classes'])
model = gnn_model(MODEL_NAME, net_params)
model = model.to(device)
optimizer = optim.Adam(model.parameters(), lr=params['init_lr'], weight_decay=params['weight_decay'])
scheduler = optim.lr_scheduler.ReduceLROnPlateau(optimizer, mode='min',
factor=params['lr_reduce_factor'],
patience=params['lr_schedule_patience'],
verbose=True)
epoch_train_losses, epoch_val_losses = [], []
epoch_train_accs, epoch_val_accs = [], []
if MODEL_NAME in ['RingGNN', '3WLGNN']:
# import train functions specific for WL-GNNs
from train.train_SBMs_node_classification import train_epoch_dense as train_epoch, evaluate_network_dense as evaluate_network
train_loader = DataLoader(trainset, shuffle=True, collate_fn=dataset.collate_dense_gnn)
val_loader = DataLoader(valset, shuffle=False, collate_fn=dataset.collate_dense_gnn)
test_loader = DataLoader(testset, shuffle=False, collate_fn=dataset.collate_dense_gnn)
else:
# import train functions for all other GCNs
from train.train_SBMs_node_classification import train_epoch_sparse as train_epoch, evaluate_network_sparse as evaluate_network # import train functions
train_loader = DataLoader(trainset, batch_size=params['batch_size'], shuffle=True, collate_fn=dataset.collate)
val_loader = DataLoader(valset, batch_size=params['batch_size'], shuffle=False, collate_fn=dataset.collate)
test_loader = DataLoader(testset, batch_size=params['batch_size'], shuffle=False, collate_fn=dataset.collate)
# At any point you can hit Ctrl + C to break out of training early.
try:
with tqdm(range(params['epochs'])) as t:
for epoch in t:
t.set_description('Epoch %d' % epoch)
start = time.time()
if MODEL_NAME in ['RingGNN', '3WLGNN']: # since different batch training function for dense GNNs
epoch_train_loss, epoch_train_acc, optimizer = train_epoch(model, optimizer, device, train_loader, epoch, params['batch_size'])
else: # for all other models common train function
epoch_train_loss, epoch_train_acc, optimizer = train_epoch(model, optimizer, device, train_loader, epoch)
epoch_val_loss, epoch_val_acc = evaluate_network(model, device, val_loader, epoch)
_, epoch_test_acc = evaluate_network(model, device, test_loader, epoch)
epoch_train_losses.append(epoch_train_loss)
epoch_val_losses.append(epoch_val_loss)
epoch_train_accs.append(epoch_train_acc)
epoch_val_accs.append(epoch_val_acc)
writer.add_scalar('train/_loss', epoch_train_loss, epoch)
writer.add_scalar('val/_loss', epoch_val_loss, epoch)
writer.add_scalar('train/_acc', epoch_train_acc, epoch)
writer.add_scalar('val/_acc', epoch_val_acc, epoch)
writer.add_scalar('test/_acc', epoch_test_acc, epoch)
writer.add_scalar('learning_rate', optimizer.param_groups[0]['lr'], epoch)
t.set_postfix(time=time.time()-start, lr=optimizer.param_groups[0]['lr'],
train_loss=epoch_train_loss, val_loss=epoch_val_loss,
train_acc=epoch_train_acc, val_acc=epoch_val_acc,
test_acc=epoch_test_acc)
per_epoch_time.append(time.time()-start)
# Saving checkpoint
ckpt_dir = os.path.join(root_ckpt_dir, "RUN_")
if not os.path.exists(ckpt_dir):
os.makedirs(ckpt_dir)
torch.save(model.state_dict(), '{}.pkl'.format(ckpt_dir + "/epoch_" + str(epoch)))
files = glob.glob(ckpt_dir + '/*.pkl')
for file in files:
epoch_nb = file.split('_')[-1]
epoch_nb = int(epoch_nb.split('.')[0])
if epoch_nb < epoch-1:
os.remove(file)
scheduler.step(epoch_val_loss)
if optimizer.param_groups[0]['lr'] < params['min_lr']:
print("\n!! LR SMALLER OR EQUAL TO MIN LR THRESHOLD.")
break
# Stop training after params['max_time'] hours
if time.time()-start0 > params['max_time']*3600:
print('-' * 89)
print("Max_time for training elapsed {:.2f} hours, so stopping".format(params['max_time']))
break
except KeyboardInterrupt:
print('-' * 89)
print('Exiting from training early because of KeyboardInterrupt')
_, test_acc = evaluate_network(model, device, test_loader, epoch)
_, train_acc = evaluate_network(model, device, train_loader, epoch)
print("Test Accuracy: {:.4f}".format(test_acc))
print("Train Accuracy: {:.4f}".format(train_acc))
print("Convergence Time (Epochs): {:.4f}".format(epoch))
print("TOTAL TIME TAKEN: {:.4f}s".format(time.time()-start0))
print("AVG TIME PER EPOCH: {:.4f}s".format(np.mean(per_epoch_time)))
writer.close()
"""
Write the results in out_dir/results folder
"""
with open(write_file_name + '.txt', 'w') as f:
f.write("""Dataset: {},\nModel: {}\n\nparams={}\n\nnet_params={}\n\n{}\n\nTotal Parameters: {}\n\n
FINAL RESULTS\nTEST ACCURACY: {:.4f}\nTRAIN ACCURACY: {:.4f}\n\n
Convergence Time (Epochs): {:.4f}\nTotal Time Taken: {:.4f} hrs\nAverage Time Per Epoch: {:.4f} s\n\n\n"""\
.format(DATASET_NAME, MODEL_NAME, params, net_params, model, net_params['total_param'],
test_acc, train_acc, epoch, (time.time()-start0)/3600, np.mean(per_epoch_time)))
# + colab={"base_uri": "https://localhost:8080/"} id="hOAYM1MUDDcJ" outputId="45948f76-710e-4a3b-d3fa-08d88be8ba58"
# !pip install nbconvert
import nbconvert
# + colab={"base_uri": "https://localhost:8080/"} id="JHW73vmSJK3y" outputId="2e88fae6-f375-41d7-a6bf-f63ab5b607a0"
def main(notebook_mode=False,config=None):
"""
USER CONTROLS
"""
# terminal mode
if notebook_mode==False:
parser = argparse.ArgumentParser()
parser.add_argument('--config', help="Please give a config.json file with training/model/data/param details")
parser.add_argument('--gpu_id', help="Please give a value for gpu id")
parser.add_argument('--model', help="Please give a value for model name")
parser.add_argument('--dataset', help="Please give a value for dataset name")
parser.add_argument('--out_dir', help="Please give a value for out_dir")
parser.add_argument('--seed', help="Please give a value for seed")
parser.add_argument('--epochs', help="Please give a value for epochs")
parser.add_argument('--batch_size', help="Please give a value for batch_size")
parser.add_argument('--init_lr', help="Please give a value for init_lr")
parser.add_argument('--lr_reduce_factor', help="Please give a value for lr_reduce_factor")
parser.add_argument('--lr_schedule_patience', help="Please give a value for lr_schedule_patience")
parser.add_argument('--min_lr', help="Please give a value for min_lr")
parser.add_argument('--weight_decay', help="Please give a value for weight_decay")
parser.add_argument('--print_epoch_interval', help="Please give a value for print_epoch_interval")
parser.add_argument('--L', help="Please give a value for L")
parser.add_argument('--hidden_dim', help="Please give a value for hidden_dim")
parser.add_argument('--out_dim', help="Please give a value for out_dim")
parser.add_argument('--residual', help="Please give a value for residual")
parser.add_argument('--edge_feat', help="Please give a value for edge_feat")
parser.add_argument('--readout', help="Please give a value for readout")
parser.add_argument('--kernel', help="Please give a value for kernel")
parser.add_argument('--n_heads', help="Please give a value for n_heads")
parser.add_argument('--gated', help="Please give a value for gated")
parser.add_argument('--in_feat_dropout', help="Please give a value for in_feat_dropout")
parser.add_argument('--dropout', help="Please give a value for dropout")
parser.add_argument('--layer_norm', help="Please give a value for layer_norm")
parser.add_argument('--batch_norm', help="Please give a value for batch_norm")
parser.add_argument('--sage_aggregator', help="Please give a value for sage_aggregator")
parser.add_argument('--data_mode', help="Please give a value for data_mode")
parser.add_argument('--num_pool', help="Please give a value for num_pool")
parser.add_argument('--gnn_per_block', help="Please give a value for gnn_per_block")
parser.add_argument('--embedding_dim', help="Please give a value for embedding_dim")
parser.add_argument('--pool_ratio', help="Please give a value for pool_ratio")
parser.add_argument('--linkpred', help="Please give a value for linkpred")
parser.add_argument('--cat', help="Please give a value for cat")
parser.add_argument('--self_loop', help="Please give a value for self_loop")
parser.add_argument('--max_time', help="Please give a value for max_time")
parser.add_argument('--pos_enc_dim', help="Please give a value for pos_enc_dim")
parser.add_argument('--pos_enc', help="Please give a value for pos_enc")
args = parser.parse_args()
with open(args.config) as f:
config = json.load(f)
# device
if args.gpu_id is not None:
config['gpu']['id'] = int(args.gpu_id)
config['gpu']['use'] = True
device = gpu_setup(config['gpu']['use'], config['gpu']['id'])
# model, dataset, out_dir
if args.model is not None:
MODEL_NAME = args.model
else:
MODEL_NAME = config['model']
if args.dataset is not None:
DATASET_NAME = args.dataset
else:
DATASET_NAME = config['dataset']
dataset = LoadData(DATASET_NAME)
if args.out_dir is not None:
out_dir = args.out_dir
else:
out_dir = config['out_dir']
# parameters
params = config['params']
if args.seed is not None:
params['seed'] = int(args.seed)
if args.epochs is not None:
params['epochs'] = int(args.epochs)
if args.batch_size is not None:
params['batch_size'] = int(args.batch_size)
if args.init_lr is not None:
params['init_lr'] = float(args.init_lr)
if args.lr_reduce_factor is not None:
params['lr_reduce_factor'] = float(args.lr_reduce_factor)
if args.lr_schedule_patience is not None:
params['lr_schedule_patience'] = int(args.lr_schedule_patience)
if args.min_lr is not None:
params['min_lr'] = float(args.min_lr)
if args.weight_decay is not None:
params['weight_decay'] = float(args.weight_decay)
if args.print_epoch_interval is not None:
params['print_epoch_interval'] = int(args.print_epoch_interval)
if args.max_time is not None:
params['max_time'] = float(args.max_time)
# network parameters
net_params = config['net_params']
net_params['device'] = device
net_params['gpu_id'] = config['gpu']['id']
net_params['batch_size'] = params['batch_size']
if args.L is not None:
net_params['L'] = int(args.L)
if args.hidden_dim is not None:
net_params['hidden_dim'] = int(args.hidden_dim)
if args.out_dim is not None:
net_params['out_dim'] = int(args.out_dim)
if args.residual is not None:
net_params['residual'] = True if args.residual=='True' else False
if args.edge_feat is not None:
net_params['edge_feat'] = True if args.edge_feat=='True' else False
if args.readout is not None:
net_params['readout'] = args.readout
if args.kernel is not None:
net_params['kernel'] = int(args.kernel)
if args.n_heads is not None:
net_params['n_heads'] = int(args.n_heads)
if args.gated is not None:
net_params['gated'] = True if args.gated=='True' else False
if args.in_feat_dropout is not None:
net_params['in_feat_dropout'] = float(args.in_feat_dropout)
if args.dropout is not None:
net_params['dropout'] = float(args.dropout)
if args.layer_norm is not None:
net_params['layer_norm'] = True if args.layer_norm=='True' else False
if args.batch_norm is not None:
net_params['batch_norm'] = True if args.batch_norm=='True' else False
if args.sage_aggregator is not None:
net_params['sage_aggregator'] = args.sage_aggregator
if args.data_mode is not None:
net_params['data_mode'] = args.data_mode
if args.num_pool is not None:
net_params['num_pool'] = int(args.num_pool)
if args.gnn_per_block is not None:
net_params['gnn_per_block'] = int(args.gnn_per_block)
if args.embedding_dim is not None:
net_params['embedding_dim'] = int(args.embedding_dim)
if args.pool_ratio is not None:
net_params['pool_ratio'] = float(args.pool_ratio)
if args.linkpred is not None:
net_params['linkpred'] = True if args.linkpred=='True' else False
if args.cat is not None:
net_params['cat'] = True if args.cat=='True' else False
if args.self_loop is not None:
net_params['self_loop'] = True if args.self_loop=='True' else False
if args.pos_enc is not None:
net_params['pos_enc'] = True if args.pos_enc=='True' else False
if args.pos_enc_dim is not None:
net_params['pos_enc_dim'] = int(args.pos_enc_dim)
# notebook mode
if notebook_mode:
# parameters
params = config['params']
# dataset
DATASET_NAME = config['dataset']
dataset = LoadData(DATASET_NAME)
# device
device = gpu_setup(config['gpu']['use'], config['gpu']['id'])
out_dir = config['out_dir']
# GNN model
MODEL_NAME = config['model']
# network parameters
net_params = config['net_params']
net_params['device'] = device
net_params['gpu_id'] = config['gpu']['id']
net_params['batch_size'] = params['batch_size']
# SBM
net_params['in_dim'] = torch.unique(dataset.train[0][0].ndata['feat'],dim=0).size(0) # node_dim (feat is an integer)
net_params['n_classes'] = torch.unique(dataset.train[0][1],dim=0).size(0)
if MODEL_NAME == 'RingGNN':
num_nodes = [dataset.train[i][0].number_of_nodes() for i in range(len(dataset.train))]
net_params['avg_node_num'] = int(np.ceil(np.mean(num_nodes)))
root_log_dir = out_dir + 'logs/' + MODEL_NAME + "_" + DATASET_NAME + "_GPU" + str(config['gpu']['id']) + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
root_ckpt_dir = out_dir + 'checkpoints/' + MODEL_NAME + "_" + DATASET_NAME + "_GPU" + str(config['gpu']['id']) + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
write_file_name = out_dir + 'results/result_' + MODEL_NAME + "_" + DATASET_NAME + "_GPU" + str(config['gpu']['id']) + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
write_config_file = out_dir + 'configs/config_' + MODEL_NAME + "_" + DATASET_NAME + "_GPU" + str(config['gpu']['id']) + "_" + time.strftime('%Hh%Mm%Ss_on_%b_%d_%Y')
dirs = root_log_dir, root_ckpt_dir, write_file_name, write_config_file
if not os.path.exists(out_dir + 'results'):
os.makedirs(out_dir + 'results')
if not os.path.exists(out_dir + 'configs'):
os.makedirs(out_dir + 'configs')
net_params['total_param'] = view_model_param(MODEL_NAME, net_params)
train_val_pipeline(MODEL_NAME, dataset, params, net_params, dirs)
if notebook_mode==True:
config = {}
# gpu config
gpu = {}
gpu['use'] = use_gpu
gpu['id'] = gpu_id
config['gpu'] = gpu
# GNN model, dataset, out_dir
config['model'] = MODEL_NAME
config['dataset'] = DATASET_NAME
config['out_dir'] = out_dir
# parameters
params = {}
params['seed'] = seed
params['epochs'] = epochs
params['batch_size'] = batch_size
params['init_lr'] = init_lr
params['lr_reduce_factor'] = lr_reduce_factor
params['lr_schedule_patience'] = lr_schedule_patience
params['min_lr'] = min_lr
params['weight_decay'] = weight_decay
params['print_epoch_interval'] = 5
params['max_time'] = max_time
config['params'] = params
# network parameters
config['net_params'] = net_params
# convert to .py format
from utils.cleaner_main import *
cleaner_main('main_SBMs_node_classification')
main(True,config)
else:
main()
|
4L_Cluster_Monet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="SxIf0YKuI2Rw"
# Global data variables
SANDBOX_NAME = ''# Sandbox Name
DATA_PATH = "/data/sandboxes/" + SANDBOX_NAME + "/data/data/"
# + [markdown] id="upw-KTIBI2SO"
#
# # SparkSession & SparkContext
# + [markdown] id="_MAn2VvNI2SS"
#
#
# En versiones de Spark anteriores a 2.0.0, __sparkContext__ era usado como el canal para acceder a todas las funcionalidades de spark. Se encarga de gestionar la conexión con el resource manager y los ejecutores.
#
# A partir de la versión 2.0.0, __sparkSession__ funciona como punto de entrada único para interactuar con spark, contiene toda la funcionalidad básica del _sparkContext_ y añade la API para interacturar con Dataframes.
#
# En el caso de Intelligence ambos objetos son creados automáticamente al iniciar un _kernel_ de _pyspark_.
# + id="9HwCK5UzI2SX" outputId="f1ba2bd1-dc59-4109-a4a4-0041902cc7f1"
spark
# + id="XOHY2_LpI2Sc" outputId="8cd8ba0e-62e9-416e-cacc-8a0f4b2d6198"
spark.version
# + id="8d1-4NHYI2Sg" outputId="5ffdee3d-8cec-487e-e8b2-3c4b6ee693dd"
sc
# + [markdown] id="Xw4hnTbWI2Sm"
#
#
# _sparkSession_ contiene al _sparkContext_
# + id="KY9FykAeI2So" outputId="131e2b4c-a6c9-4fcf-af1d-22d15217b6d5"
spark.sparkContext
# + [markdown] id="gFFFtFyFI2Ss"
#
#
# _sparkContext_ contiene la configuración de spark
# + id="dydqC1JMI2Sw" outputId="7aff4560-ffc0-4345-9c93-5a1dc6e86f09"
sc.getConf().getAll()
# + [markdown] id="BZEXS_wsI2S1"
#
# # DataFrames
# Es una colección inmutable y distribuida de datos organizados en columnas de forma tabular que permite una abstracción de los datos a alto nivel.
#
#
# __Características__
#
# - Conjunto de filas (Rows) con un esquema de datos (Schema)
# - Colección distribuida de datos organizados en filas y columnas nombradas.
# - Conceptualmente equivalente a una tabla en una BD relacional, o a un data frame de R o Python (pandas), pero con optimizaciones avanzadas para soportar aplicaciones Big Data y Data Science.
# - Pueden construirse a partir de distintas fuentes: ficheros de datos estructurados, tablas de Hive, BD externas, o RDDs
# - DataFrame API disponible en Scala, Java, Python, y R
# + [markdown] id="q-0JO6xcI2TA"
#
#
# En SparkSQL cada __transformación__ sobre un DataFrame se añade a lo que se conoce como *query plan*. Cuando se aplica una __acción__ sobre el DataFrame el *Catalyst Optimizer* analiza el *query plan* e intenta optimizarlo, luego selecciona el plan físico (transformación a operaciones RDD de bajo nivel) más eficiente para la ejecución del plan, y lo ejecuta.
#
# Se puede consultar el *query plan* usando la función `explain()` sobre el DataFrame.
# + [markdown] id="8x6nLHXSI2TD"
#
#
# ## Creación de DataFrames
# Normalmente un DataFrame se crea leyendo datos desde una fuente externa (S3, HDFS, etc). Por ejemplo para leer el fichero 'Building_Permits.csv' almacenado en el Sandbox Data:
# + id="lXgb_5hMI2TE"
file_name = "Building_Permits.csv"
buildings_df = spark.read.csv(DATA_PATH + file_name, sep=',', header=True, inferSchema=True)
# + id="gB0RDD5KI2TG" outputId="9bc46387-3b99-4ac2-c158-1bbb90fa19c9"
buildings_df.show(3)
# + [markdown] id="_fpY9DuBI2TH"
#
#
# En el caso de leer ficheros parquet no es necesario indicar una cabecera ni separador. Esta iformación ya está contenida en los propios ficheros parquet. Por ejemplo:
# + id="XOp8k7X5I2TK"
file_path = "parquet/online_retail"
online_df = spark.read.parquet(DATA_PATH + file_path)
# + id="oxUaLhObI2TM" outputId="77c3ee4b-5a50-409e-e5e3-f5787b5f31ee"
online_df.show(5)
# + [markdown] id="e5VXAKbDI2TP"
#
#
# También se puede crear un DataFrame de Spark a partir de un DataFrame de Pandas.
# + id="gdDOMitsI2TR" outputId="e625f753-4f91-489c-e733-91b89b0697a7"
import pandas as pd
df = pd.DataFrame({'city': ['Madrid', 'Birmingham', 'Barcelona', 'Mexico City'],
'population': [3.2, 0.2, 1.6, 8.8]})
df
# + id="7s-xp_lpI2TT" outputId="7d8e3c18-07e6-4453-d5be-27986a3e06f9"
df_s = spark.createDataFrame(df)
df_s
# + id="_K1AG1meI2Tu" outputId="e09e585a-85b6-449e-cfdf-c20e6568b9df"
df_s.show()
# + [markdown] id="_S-2PFS_I2Tw"
#
# ## Información básica de DataFrames
# + [markdown] id="2meMt1WzI2Tx"
#
#
# ### Previsualización
#
# `show` es un método que muestra por pantalla _n_ filas del DataFrame.
# + id="ZswmFAarI2Ty" outputId="836fe0e3-4275-4600-aa26-cf43da918337"
online_df.show(6)
# + [markdown] id="_5sCIWinI2T1"
#
#
# ### Dimensions
#
# En Spark, no existe un método *shape*, por lo que hay que contar por separados las filas y las columnas.
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="xQKP_BW2I2T2" executionInfo={"status": "error", "timestamp": 1614107663337, "user_tz": 180, "elapsed": 2231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="a4bea7e6-724c-4925-c2fb-bc46abeb1750"
buildings_df.count()
# + [markdown] id="Tct_bLRTI2T4"
#
#
# `columns` es un atributo que contiene los nombres de las columnas del DataFrame.
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="vO7vV4kbI2T5" executionInfo={"status": "error", "timestamp": 1614107685267, "user_tz": 180, "elapsed": 1770, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="a596f84e-453a-4dd2-9af0-816d36be14f4"
buildings_df.columns
# + id="wXW3neRDI2T6" executionInfo={"status": "aborted", "timestamp": 1614107663673, "user_tz": 180, "elapsed": 2517, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}}
len(buildings_df.columns)
# + [markdown] id="kZBKt84SI2T8"
#
#
# ### Schema
#
# El schema de un dataframe nos muestra como se interpretaran los datos. Esto no significa que los datos estén así. _schema_ es un atributo del objeto, no un método. _printSchema()_ es un método que muestra una versión más entendidible del _schema_.
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="GE_1dYYwI2T9" executionInfo={"status": "error", "timestamp": 1614108174622, "user_tz": 180, "elapsed": 583, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="35a8fe35-d988-4877-f4dd-0d13befe7924"
buildings_df.schema
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="8hBmHpXzI2T_" executionInfo={"status": "error", "timestamp": 1614108174971, "user_tz": 180, "elapsed": 906, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="523bd6de-36fb-4130-95f6-0d82a3a8e7dd"
buildings_df.printSchema()
# + [markdown] id="K4BxoaLPI2UA"
#
#
# ### dtypes
#
# El atributo `dtypes` contiene los nombres de las columnas del dataframe junto con su tipo. Esto permite seleccionar nombres de columnas basados en el tipo, normalmente las variables categóricas (string y boolean) tienen tratamientos distintos a las numéricas (enteras y decimales).
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="5iZ-qIz3I2UC" executionInfo={"status": "error", "timestamp": 1614108361339, "user_tz": 180, "elapsed": 733, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="9de2e623-3f00-4679-c8c0-87c5f1e96a4a"
online_df.dtypes
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="Ns6B8ckrI2UI" executionInfo={"status": "error", "timestamp": 1614108361780, "user_tz": 180, "elapsed": 1146, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="81ca374e-c8c2-4854-852f-3b0f90cbe27b"
categorial_columns = [c for c,t in online_df.dtypes if t in ['string', 'bool']]
categorial_columns
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="-_vS_f2JI2UK" executionInfo={"status": "error", "timestamp": 1614108361793, "user_tz": 180, "elapsed": 1140, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="bf9b5a98-ac2c-4135-d2cd-2ab565aa4707"
numerical_columns = [c for c,t in online_df.dtypes if t in ['int', 'double']]
numerical_columns
# + [markdown] id="sAc8PsHnI2UM"
#
# ## Acciones
#
# Las acciones principales de Spark que se van a tratar son las siguientes:
#
# - show()
# - count()
# - first()
# - take()
# - collect()
# - toPandas()
# - write()
# + [markdown] id="issqpP7EI2UN"
#
# ### show
# Muestra por pantalla _n_ filas del DataFrame (20 por defecto). Es una llamada a un `print`, no permite almacenar el resultado en una variable. El parámetro `truncate` limita el número de caracteres de cada campo.
# + id="spHh_QOrI2UO" outputId="bb063a0a-5c85-486d-db2e-ac80fd42a601"
online_df.show(5)
# + id="xU-2-Q3_I2UP" outputId="a921424e-e395-462c-8c85-fb9c978813d9"
online_df.show(5, truncate=False)
# + [markdown] id="10XM1anqI2UT"
#
#
# ### count
#
# Cuenta el número de filas del DataFrame
# + id="3lQOGP1DI2UV" outputId="b0495f31-6b8c-4f2b-a15b-5cde068d92f5"
online_df.count()
# + id="GIk-U5rTI2UV" outputId="46dad128-f19b-43c6-846a-6fe503c139f2"
buildings_df.count()
# + [markdown] id="4eauJ42rI2UX"
#
#
# ### first
#
# Devuelve una única fila del DataFrame como un objeto de tipo `Row`. A los elementos del objeto `Row` se puede acceder tanto por nombre como por posición.
# + colab={"base_uri": "https://localhost:8080/", "height": 191} id="IZAyMPOsI2Uc" executionInfo={"status": "error", "timestamp": 1614108771055, "user_tz": 180, "elapsed": 749, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="2f608619-0b0c-46b3-a9c8-cf6bcd81689c"
single_row = online_df.first()
single_row
# + colab={"base_uri": "https://localhost:8080/", "height": 172} id="474xdqmhI2Ue" executionInfo={"status": "error", "timestamp": 1614108771068, "user_tz": 180, "elapsed": 740, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}} outputId="5903c50e-4b00-4e18-b1e5-157d39c9bd14"
type(single_row)
# + id="pLx3eMKNI2Ug" executionInfo={"status": "aborted", "timestamp": 1614108771060, "user_tz": 180, "elapsed": 709, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}}
single_row[1]
# + id="KoK1k5VPI2Uh" executionInfo={"status": "aborted", "timestamp": 1614108771065, "user_tz": 180, "elapsed": 704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gg1TanzZFZvSS_bpI7mh9ac8jVLykMdhlIBCEsjCQ=s64", "userId": "15188645614769644944"}}
single_row['StockCode']
# + [markdown] id="zLPiRGcII2Uj"
#
#
# ### take
#
# Devuelve _n_ filas del DataFrame como una lista de objetos `Row`.
# + id="_zEzABgxI2Ul" outputId="c96d420a-4ef5-46bf-8b87-b071aeed9712"
row_list = online_df.take(4)
row_list
# + id="mK_p10eDI2Um" outputId="ddbec769-7915-452a-fff1-324a2b488f52"
row_list[1]
# + id="F9gUEem3I2Up" outputId="b3e88a86-7242-4c34-f850-58adc78d074b"
row_list[1][2]
# + id="F9TPzdwiI2Uq" outputId="64b1afac-a13c-4648-b8f6-adad275cc24f"
row_list[1]['Description']
# + [markdown] id="7R0YQ2ELI2Ur"
#
#
# ### collect
#
# Vuelca en un solo nodo todos los datos y los almacena en formato lista de Rows. Funciona como un `take` sin límite. Es una función a evitar salvo en ocasiones muy específicas donde sea necesario tener todos los datos en local y no haya otra forma de gestionarlos. Utilizar únicamente con datos filtrados y/o agregados.
# + id="ucWKTwBpI2Us"
all_rows = online_df.collect()
# + id="Os53_djiI2Uu" outputId="8dda5887-e841-4bda-fc75-8863709375de"
len(all_rows)
# + id="_PgdWaliI2Uv" outputId="2e6f3785-8469-40c3-cf3d-ee5f64382acb"
all_rows[:4]
# + [markdown] id="W7k7_41wI2Ux"
#
#
# ### toPandas
#
# Vuelca en un nodo todos los datos como un DataFrame de Pandas. Se deben seguir las mismas restricciones que con `collect`.
# + id="HHNpLHHlI2Uy"
online_pandas = online_df.toPandas()
# + id="FrhATGXwI2Uz" outputId="d0ad6709-79f4-4753-8570-2ff3b8021c7e"
type(online_pandas)
# + id="2WxUpeLVI2U1" outputId="71ba2443-d2bc-437b-8169-2d0b7cd723ad"
online_pandas.head(3)
# + [markdown] id="hmjTLH7OI2U1"
#
#
# ### write
#
# Guarda la información de la tabla en fichero. Se puede escribir tanto en texto plano (CSV) como en formato parquet.
# + id="4HWW5_FXI2U2"
online_df.write.csv(DATA_PATH + 'online_retail_csv')
# + id="D8tFCJ-FI2U2"
online_df.write.parquet(DATA_PATH + 'online_retail_parquet')
# + [markdown] id="iUnAzKefI2U4"
#
#
# Podemos comprobar que usando la misma ruta que hemos utilizado para escribir, podemos leer el DataFrame tanto de un CSV como de un Parquet
# + id="KlozuI-nI2U4" outputId="a2475c6c-87fe-442f-f128-7e3f896b49f7"
spark.read.csv(DATA_PATH + "online_retail_csv").show(5)
# + id="lpd3fCP8I2U5" outputId="badf5370-e42a-4868-9960-07d9f2deddc3"
spark.read.parquet(DATA_PATH + "online_retail_parquet").show(5)
# + [markdown] id="jxQVPl_wI2U8"
#
#
# La partición de tablas es un forma habitual de optimización. En una tabla particionada, los datos se almacenan en diferentes carpetas, donde los valores de la columna utilizada para particionar forman parte de la ruta de cada carpeta. Al leer un fichero particionado, Spark es capaz de descubrir e inferir la información de partición automáticamente para recuperar los datos correctamente.
# + id="pFOM3YpFI2U9"
online_df.write.partitionBy("Country").parquet(DATA_PATH + "online_retail_parquet_partition")
# + [markdown] id="OGJ4FfZLI2U_"
#
#
# Podemos leer los datos del parquet que acabamos de escribir:
#
# Leyendo todo el parquet completo (sin especificar la columna con la que hemos particionado):
# + id="C79g5A28I2VA" outputId="e21f5f20-48c1-438a-aeb8-6ee7c4049945"
spark.read.parquet(DATA_PATH + "online_retail_parquet_partition").show(5)
# + [markdown] id="P931CQBaI2VD"
#
#
# Leyendo solo la partición correspondiente a un valor de la columna utilizada para particionar:
# + id="56EwjRGmI2VF" outputId="53d87905-a20d-4ab6-ac46-15c27bdef4a0"
spark.read.parquet(DATA_PATH + "online_retail_parquet_partition/Country=United Kingdom").show(5)
|
2021Q1_DSF/5.- Spark/notebooks/spark_sql/01_spark_sql_basics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
movies_df = pd.read_csv("Movie.csv")
movies_df
movies_df.sort_values(by=['userId'])
#number of unique users in the dataset
len(movies_df.userId.unique())
len(movies_df.movie.unique())
user_movies_df = movies_df.pivot(index='userId',
columns='movie',
values='rating').reset_index(drop=True)
user_movies_df
user_movies_df.index = movies_df.userId.unique()
user_movies_df
#Impute those NaNs with 0 values
user_movies_df.fillna(0, inplace=True)
user_movies_df
#Calculating Cosine Similarity between Users
from sklearn.metrics import pairwise_distances
from scipy.spatial.distance import cosine, correlation
user_sim = 1 - pairwise_distances(user_movies_df.values,metric='cosine')
user_sim
#Store the results in a dataframe
user_sim_df = pd.DataFrame(user_sim)
user_sim_df
#Set the index and column names to user ids
user_sim_df.index = movies_df.userId.unique()
user_sim_df.columns = movies_df.userId.unique()
user_sim_df.iloc[0:5, 0:5]
np.fill_diagonal(user_sim, 0)
user_sim_df.iloc[0:5, 0:5]
#Most Similar Users
user_sim_df.idxmax(axis=1)[0:5]
movies_df[(movies_df['userId']==6) | (movies_df['userId']==168)]
user_1=movies_df[movies_df['userId']==6]
user_2=movies_df[movies_df['userId']==168]
user_2.movie
user_1.movie
pd.merge(user_1,user_2,on='movie',how='outer')
|
Recommendation Engine.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Project Euler: Problem 2
# + [markdown] nbgrader={}
# Each new term in the Fibonacci sequence is generated by adding the previous two terms. By starting with 0 and 1, the first 12 terms will be:
#
# 0, 1, 1, 2, 3, 5, 8, 13, 21, 34, 55, 89, ...
#
# By considering the terms in the Fibonacci sequence whose values do not exceed four million, find the sum of the even-valued terms.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
n=0 #this will be the fibonacci numbers
L=[0,1] #list of the numbers
s=0 #the sum (eventually)
while n<4000000: #limiting n
n=L[-1]+(L[-2]) #to define which terms i want
L.append(n) #to put n in the list
if n%2 == 0: #to get the even numbers
s += n #sum them
print (s) #print the sum
# + deletable=false nbgrader={"checksum": "e8afe8a5735f0fff949b706895f8583d", "grade": true, "grade_id": "projecteuler2", "points": 10}
# This cell will be used for grading, leave it at the end of the notebook.
|
assignments/assignment02/ProjectEuler2.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Basic Gates Kata
#
# **Basic Gates** quantum kata is a series of exercises designed
# to get you familiar with the basic quantum gates in Q#.
# It covers the following topics:
# * basic single-qubit and multi-qubit gates,
# * adjoint and controlled gates,
# * using gates to modify the state of a qubit.
#
# Each task is wrapped in one operation preceded by the description of the task.
# Your goal is to fill in the blank (marked with `// ...` comments)
# with some Q# code that solves the task. To verify your answer, run the cell using Ctrl/⌘+Enter.
#
# Most tasks in this kata can be done using exactly one gate.
# None of the tasks require measurement, and the tests are written so as to fail if qubit state is measured.
#
# The tasks are given in approximate order of increasing difficulty; harder ones are marked with asterisks.
# To begin, first prepare this notebook for execution (if you skip this step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells):
%package Microsoft.Quantum.Katas::0.10.2002.2610
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
#
# ## Part I. Single-Qubit Gates
#
#
# ### Theory
#
# * A list of most common gates can be found in [this Wikipedia article](https://en.wikipedia.org/wiki/Quantum_logic_gate).
# * [Quirk](http://algassert.com/quirk) is a convenient tool for visualizing the effect of gates on qubit states.
#
# ### Q# materials
#
# * Basic gates provided in Q# belong to the `Microsoft.Quantum.Intrinsic` namespace and are listed [here](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic).
#
# > Note that all operations in this section have `is Adj+Ctl` in their signature.
# This means that they should be implemented in a way that allows Q#
# to compute their adjoint and controlled variants automatically.
# Since each task is solved using only intrinsic gates, you should not need to put any special effort in this.
# ### Task 1.1. State flip: $|0\rangle$ to $|1\rangle$ and vice versa
#
# **Input:** A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
#
# **Goal:** Change the state of the qubit to $\alpha |1\rangle + \beta |0\rangle$.
#
# **Example:**
#
# If the qubit is in state $|0\rangle$, change its state to $|1\rangle$.
#
# If the qubit is in state $|1\rangle$, change its state to $|0\rangle$.
#
# > Note that this operation is self-adjoint: applying it for a second time
# > returns the qubit to the original state.
# +
%kata T101_StateFlip_Test
operation StateFlip (q : Qubit) : Unit is Adj+Ctl {
// The Pauli X gate will change the |0⟩ state to the |1⟩ state and vice versa.
// Type X(q);
// Then run the cell using Ctrl/⌘+Enter.
// ...
}
# -
# ### Task 1.2. Basis change: $|0\rangle$ to $|+\rangle$ and $|1\rangle$ to $|-\rangle$ (and vice versa)
#
# **Input**: A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
#
# **Goal**: Change the state of the qubit as follows:
# * If the qubit is in state $|0\rangle$, change its state to $|+\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle + |1\rangle\big)$.
# * If the qubit is in state $|1\rangle$, change its state to $|-\rangle = \frac{1}{\sqrt{2}} \big(|0\rangle - |1\rangle\big)$.
# * If the qubit is in superposition, change its state according to the effect on basis vectors.
#
# > Note:
# > $|+\rangle$ and $|-\rangle$ form a different basis for single-qubit states, called X basis.
# > $|0\rangle$ and $|1\rangle$ are called Z basis.
#
# +
%kata T102_BasisChange_Test
operation BasisChange (q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.3. Sign flip: $|+\rangle$ to $|-\rangle$ and vice versa.
#
# **Input**: A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
#
# **Goal** : Change the qubit state to $\alpha |0\rangle - \beta |1\rangle$ (flip the sign of $|1\rangle$ component of the superposition).
#
# +
%kata T103_SignFlip_Test
operation SignFlip (q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.4. Amplitude change: $|0\rangle$ to $\cos{α} |0\rangle + \sin{α} |1\rangle$.
#
# **Inputs:**
#
# 1. Angle α, in radians, represented as Double.
# 2. A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
#
# **Goal:** Change the state of the qubit as follows:
# - If the qubit is in state $|0\rangle$, change its state to $\cos{α} |0\rangle + \sin{α} |1\rangle$.
# - If the qubit is in state $|1\rangle$, change its state to $-\sin{α} |0\rangle + \cos{α} |1\rangle$.
# - If the qubit is in superposition, change its state according to the effect on basis vectors.
#
# > This is the first operation in this kata that is not self-adjoint, i.e., applying it for a second time
# > does not return the qubit to the original state.
# +
%kata T104_AmplitudeChange_Test
operation AmplitudeChange (alpha : Double, q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.5. Phase flip
#
# **Input:** A qubit in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$.
#
# **Goal:** Change the qubit state to $\alpha |0\rangle + \color{red}i\beta |1\rangle$ (add a relative phase $i$ to $|1\rangle$ component of the superposition).
#
# +
%kata T105_PhaseFlip_Test
operation PhaseFlip (q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.6. Phase change
#
# **Inputs:**
#
# 1. Angle α, in radians, represented as Double.
# 2. A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
#
# **Goal:** Change the state of the qubit as follows:
# - If the qubit is in state $|0\rangle$, don't change its state.
# - If the qubit is in state $|1\rangle$, change its state to $e^{i\alpha} |1\rangle$.
# - If the qubit is in superposition, change its state according to the effect on basis vectors: $\beta |0\rangle + \color{red}{e^{i\alpha}} \gamma |1\rangle$.
#
# +
%kata T106_PhaseChange_Test
operation PhaseChange (alpha : Double, q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.7. Global phase change
# **Input:** A qubit in state $|\psi\rangle = \beta |0\rangle + \gamma |1\rangle$.
#
# **Goal**: Change the state of the qubit to $- \beta |0\rangle - \gamma |1\rangle$.
#
# > Note: this change on its own is not observable - there is no experiment you can do on a standalone qubit to figure out whether it acquired the global phase or not.
# > However, you can use a controlled version of this operation to observe the global phase it introduces.
# > This is used in later katas as part of more complicated tasks.
# +
%kata T107_GlobalPhaseChange_Test
operation GlobalPhaseChange (q : Qubit) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.8. Bell state change - 1
#
# **Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
#
# **Goal:** Change the two-qubit state to $|\Phi^{-}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle - |11\rangle\big)$.
#
# +
%kata T108_BellStateChange1_Test
operation BellStateChange1 (qs : Qubit[]) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.9. Bell state change - 2
#
# **Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
#
# **Goal:** Change the two-qubit state to $|\Psi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|01\rangle + |10\rangle\big)$.
# +
%kata T109_BellStateChange2_Test
operation BellStateChange2 (qs : Qubit[]) : Unit is Adj+Ctl {
// ...
}
# -
# ### Task 1.10. Bell state change - 3
#
# **Input:** Two entangled qubits in Bell state $|\Phi^{+}\rangle = \frac{1}{\sqrt{2}} \big(|00\rangle + |11\rangle\big)$.
#
# **Goal:** Change the two-qubit state to $|\Psi^{-}\rangle = \frac{1}{\sqrt{2}} \big(|01\rangle - |10\rangle\big)$.
#
# +
%kata T110_BellStateChange3_Test
operation BellStateChange3 (qs : Qubit[]) : Unit is Adj+Ctl {
// ...
}
# -
# ## Part II. Multi-Qubit Gates
#
# ### Q# materials
#
# * Using controlled and adjoint versions of gates is covered in the Q# documentation on [operation types](https://docs.microsoft.com/quantum/language/type-model#operation-and-function-types).
# ### Task 2.1. Two-qubit gate - 1
#
# **Input:** Two unentangled qubits (stored in an array of length 2).
# The first qubit will be in state $|\psi\rangle = \alpha |0\rangle + \beta |1\rangle$, the second - in state $|0\rangle$
# (this can be written as two-qubit state $\big(\alpha |0\rangle + \beta |1\rangle \big) \otimes |0\rangle = \alpha |00\rangle + \beta |10\rangle$.
#
#
# **Goal:** Change the two-qubit state to $\alpha |00\rangle + \beta |11\rangle$.
#
# > Note that unless the starting state of the first qubit was $|0\rangle$ or $|1\rangle$,
# > the resulting two-qubit state can not be represented as a tensor product
# > of the states of individual qubits any longer; thus the qubits become entangled.
# +
%kata T201_TwoQubitGate1_Test
operation TwoQubitGate1 (qs : Qubit[]) : Unit is Adj {
// ...
}
# -
# ### Task 2.2. Two-qubit gate - 2
#
# **Input:** Two unentangled qubits (stored in an array of length 2) in state $|+\rangle \otimes |+\rangle = \frac{1}{2} \big( |00\rangle + |01\rangle + |10\rangle \color{blue}+ |11\rangle \big)$.
#
#
# **Goal:** Change the two-qubit state to $\frac{1}{2} \big( |00\rangle + |01\rangle + |10\rangle \color{red}- |11\rangle \big)$.
#
# > Note that while the starting state can be represented as a tensor product of single-qubit states,
# > the resulting two-qubit state can not be represented in such a way.
# +
%kata T202_TwoQubitGate2_Test
operation TwoQubitGate2 (qs : Qubit[]) : Unit is Adj {
// ...
}
# -
# ### Task 2.3. Two-qubit gate - 3
#
# **Input:** Two unentangled qubits (stored in an array of length 2) in an arbitrary two-qubit state $\alpha |00\rangle + \color{blue}\beta |01\rangle + \color{blue}\gamma |10\rangle + \delta |11\rangle$.
#
#
# **Goal:** Change the two-qubit state to $\alpha |00\rangle + \color{red}\gamma |01\rangle + \color{red}\beta |10\rangle + \delta |11\rangle$.
#
# > This task can be solved using one intrinsic gate; as an exercise, try to express the solution using several (possibly controlled) Pauli gates.
# +
%kata T203_TwoQubitGate3_Test
operation TwoQubitGate3 (qs : Qubit[]) : Unit is Adj {
// ...
}
# -
# ### Task 2.4. Toffoli gate
#
# **Input:** Three qubits (stored in an array of length 3) in an arbitrary three-qubit state
# $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \zeta|101\rangle + \color{blue}\eta|110\rangle + \color{blue}\theta|111\rangle$.
#
# **Goal:** Flip the state of the third qubit if the state of the first two is $|11\rangle$, i.e., change the three-qubit state to $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \zeta|101\rangle + \color{red}\theta|110\rangle + \color{red}\eta|111\rangle$.
# +
%kata T204_ToffoliGate_Test
operation ToffoliGate (qs : Qubit[]) : Unit is Adj {
// ...
}
# -
# ### Task 2.5. Fredkin gate
#
# **Input:** Three qubits (stored in an array of length 3) in an arbitrary three-qubit state
# $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \color{blue}\zeta|101\rangle + \color{blue}\eta|110\rangle + \theta|111\rangle$.
#
# **Goal:** Swap the states of second and third qubit if and only if the state of the first qubit is $|1\rangle$, i.e., change the three-qubit state to $\alpha |000\rangle + \beta |001\rangle + \gamma |010\rangle + \delta |011\rangle + \epsilon |100\rangle + \color{red}\eta|101\rangle + \color{red}\zeta|110\rangle + \theta|111\rangle$.
# +
%kata T205_FredkinGate_Test
operation FredkinGate (qs : Qubit[]) : Unit is Adj {
// ...
}
|
BasicGates/BasicGates.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gensim, logging
logging.basicConfig(format='%(asctime)s : %(levelname)s : %(message)s', level=logging.INFO)
gmodel = gensim.models.KeyedVectors.load_word2vec_format('GoogleNews-vectors-negative300.bin', binary=True)
gmodel['cat']
gmodel['dog']
gmodel['spatula']
gmodel.similarity('cat', 'dog')
gmodel.similarity('cat', 'spatula')
from gensim.models.doc2vec import TaggedDocument
from gensim.models import Doc2Vec
def extract_words(sent):
sent = sent.lower()
sent = re.sub(r'<[^>]+>', ' ', sent) # strip html tags
sent = re.sub(r'(\w)\'(\w)', '\1\2', sent) # remove apostrophes
sent = re.sub(r'\W', ' ', sent) # remove punctuation
sent = re.sub(r'\s+', ' ', sent) # remove repeated spaces
sent = sent.strip()
return sent.split()
# +
# unsupervised training data
import re
import os
unsup_sentences = []
# source: http://ai.stanford.edu/~amaas/data/sentiment/, data from IMDB
for dirname in ["train/pos", "train/neg", "train/unsup", "test/pos", "test/neg"]:
for fname in sorted(os.listdir("aclImdb/" + dirname)):
if fname[-4:] == '.txt':
with open("aclImdb/" + dirname + "/" + fname, encoding='UTF-8') as f:
sent = f.read()
words = extract_words(sent)
unsup_sentences.append(TaggedDocument(words, [dirname + "/" + fname]))
# source: http://www.cs.cornell.edu/people/pabo/movie-review-data/
for dirname in ["review_polarity/txt_sentoken/pos", "review_polarity/txt_sentoken/neg"]:
for fname in sorted(os.listdir(dirname)):
if fname[-4:] == '.txt':
with open(dirname + "/" + fname, encoding='UTF-8') as f:
for i, sent in enumerate(f):
words = extract_words(sent)
unsup_sentences.append(TaggedDocument(words, ["%s/%s-%d" % (dirname, fname, i)]))
# source: https://nlp.stanford.edu/sentiment/, data from Rotten Tomatoes
with open("stanfordSentimentTreebank/original_rt_snippets.txt", encoding='UTF-8') as f:
for i, line in enumerate(f):
words = extract_words(sent)
unsup_sentences.append(TaggedDocument(words, ["rt-%d" % i]))
# -
len(unsup_sentences)
unsup_sentences[0:10]
import random
class PermuteSentences(object):
def __init__(self, sents):
self.sents = sents
def __iter__(self):
shuffled = list(self.sents)
random.shuffle(shuffled)
for sent in shuffled:
yield sent
permuter = PermuteSentences(unsup_sentences)
model = Doc2Vec(permuter, dm=0, hs=1, size=50)
# done with training, free up some memory
model.delete_temporary_training_data(keep_inference=True)
model.save('reviews.d2v')
# in other program, we could write: model = Doc2Vec.load('reviews.d2v')
model.infer_vector(extract_words("This place is not worth your time, let alone Vegas."))
from sklearn.metrics.pairwise import cosine_similarity
cosine_similarity(
[model.infer_vector(extract_words("This place is not worth your time, let alone Vegas."))],
[model.infer_vector(extract_words("Service sucks."))])
cosine_similarity(
[model.infer_vector(extract_words("Highly recommended."))],
[model.infer_vector(extract_words("Service sucks."))])
# +
sentences = []
sentvecs = []
sentiments = []
for fname in ["yelp", "amazon_cells", "imdb"]:
with open("sentiment labelled sentences/%s_labelled.txt" % fname, encoding='UTF-8') as f:
for i, line in enumerate(f):
line_split = line.strip().split('\t')
sentences.append(line_split[0])
words = extract_words(line_split[0])
sentvecs.append(model.infer_vector(words, steps=10)) # create a vector for this document
sentiments.append(int(line_split[1]))
# shuffle sentences, sentvecs, sentiments together
combined = list(zip(sentences, sentvecs, sentiments))
random.shuffle(combined)
sentences, sentvecs, sentiments = zip(*combined)
# +
from sklearn.neighbors import KNeighborsClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
import numpy as np
clf = KNeighborsClassifier(n_neighbors=9)
clfrf = RandomForestClassifier()
# -
scores = cross_val_score(clf, sentvecs, sentiments, cv=5)
np.mean(scores), np.std(scores)
scores = cross_val_score(clfrf, sentvecs, sentiments, cv=5)
np.mean(scores), np.std(scores)
# bag-of-words comparison
from sklearn.pipeline import make_pipeline
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
pipeline = make_pipeline(CountVectorizer(), TfidfTransformer(), RandomForestClassifier())
scores = cross_val_score(pipeline, sentences, sentiments, cv=5)
np.mean(scores), np.std(scores)
|
Chapter03/SentimentAnalysisnew.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Test Gistify Markdown
# A few code blocks to test that gistify option works as expected
# this code block should not gistify as <5 lines
import datetime as dt
import matplotlib.pyplot as plt
import numpy as np
# +
# this code block should gistify as it is >5 lines
x = np.linspace(0, 1, 101)
y = x**2
fig, ax = plt.subplots(figsize=(20,10))
plt.plot(x, y)
ax.set_title('Graph showing y=x**2')
ax.set_xlabel('x')
ax.set_ylabel('y');
|
tests/notebooks/Test Gistify Markdown.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jhpytorch
# language: python
# name: pytorch
# ---
# # S-GAN sample test
# +
import argparse
from loss import sganloss
import os
import numpy as np
from dataloader import *
import math
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import torchvision.transforms as transforms
from torchvision.utils import save_image
from torch.utils.data import DataLoader
from torchvision import datasets
from torch.autograd import Variable
import torch.nn as nn
import torch.nn.functional as F
import torch
import torchvision.models as models
# -
cuda = True if torch.cuda.is_available() else False
cuda
# +
def weights_init_normal(m): #?
classname = m.__class__.__name__
if classname.find("Conv") != -1:
torch.nn.init.normal_(m.weight.data, 0.0, 0.02)
elif classname.find("BatchNorm") != -1:
torch.nn.init.normal_(m.weight.data, 1.0, 0.02)
torch.nn.init.constant_(m.bias.data, 0.0)
class IdentityPadding(nn.Module):
def __init__(self, in_channels, out_channels, stride):
super(IdentityPadding, self).__init__()
self.pooling = nn.MaxPool2d(1, stride=stride)
self.add_channels = out_channels - in_channels
def forward(self, x):
out = F.pad(x, (0, 0, 0, 0, 0, self.add_channels))
out = self.pooling(out)
return out
class ResidualBlock(nn.Module):
def __init__(self, in_channels, out_channels, stride=1, down_sample=False):
super(ResidualBlock, self).__init__()
self.conv1 = nn.Conv2d(in_channels, out_channels, kernel_size=3,
stride=stride, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(out_channels)
self.relu = nn.ReLU(inplace=True)
self.conv2 = nn.Conv2d(out_channels, out_channels, kernel_size=3,
stride=1, padding=1, bias=False)
self.bn2 = nn.BatchNorm2d(out_channels)
self.stride = stride
if down_sample:
self.down_sample = IdentityPadding(in_channels, out_channels, stride)
else:
self.down_sample = None
def forward(self, x):
shortcut = x
out = self.conv1(x)
out = self.bn1(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn2(out)
if self.down_sample is not None:
shortcut = self.down_sample(x)
out += shortcut
out = self.relu(out)
return out
class Generator(nn.Module):
def __init__(self):
super(Generator, self).__init__()
self.FaceOcclusion_1=nn.Sequential(
nn.Conv2d(3, 64, kernel_size=7, stride=1, padding=3),
nn.InstanceNorm2d(64),
nn.ReLU(),
# -----
nn.Conv2d(64, 128, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(128),
nn.ReLU(),
nn.Conv2d(128, 256, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(256),
nn.ReLU(),
# -----
ResidualBlock(256, 256),
ResidualBlock(256, 256),
ResidualBlock(256, 256),
# -----
nn.ConvTranspose2d(256, 128, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(128),
nn.ReLU(),
nn.ConvTranspose2d(128, 64, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(64),
nn.ReLU()
# -----
)
self.FaceOcclusion_2=nn.Sequential(
nn.Conv2d(64, 1, kernel_size=7, stride=1, padding=3),
nn.Sigmoid()
)
self.FaceCompletion=nn.Sequential(
nn.Conv2d(64, 128, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(512),
nn.ReLU(),
nn.Conv2d(128, 256, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(512),
nn.ReLU(),
nn.Conv2d(256, 512, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(512),
nn.ReLU(),
# -----
nn.ConvTranspose2d(512, 256, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(256),
nn.ReLU(),
nn.ConvTranspose2d(256, 128, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(128),
nn.ReLU(),
nn.ConvTranspose2d(128, 64, kernel_size=4, stride=2, padding=1),
nn.InstanceNorm2d(64),
nn.ReLU(),
# -----
nn.Conv2d(64, 3, kernel_size=7, stride=1, padding=3),
nn.Tanh()
)
def forward(self, x):
# occlusion aware module
out_predicted=self.FaceOcclusion_1(x)
# out_InvertedM = torch.ones(1, 1, 128, 128).cuda() - x
out_predictedM=self.FaceOcclusion_2(out_predicted)
out_InvertedM=torch.ones(1, 1, 128, 128) - out_predictedM
out_oa=torch.matmul(out_predicted, out_predictedM)
# face completion module
out_synth=self.FaceCompletion(out_oa)
out_fc=torch.matmul(out_InvertedM, out_synth)
out_filter=torch.matmul(x, out_predictedM)
out_final=out_filter + out_fc
return out_predictedM, out_InvertedM, out_synth, out_final
# +
class weight():
def __init__(self):
self.lam1 = 0.1
self.lam2 = 0.2
self.lam3 = 0.2
self.lam4 = 0.2
self.lam5 = 0.1
self.lam6 = 0.2
self.alpha = 0.5
self.beta = 0.5
w=weight()
# +
class Discriminator(nn.Module):
def __init__(self):
super(Discriminator, self).__init__()
self.discriminator_block = nn.Sequential(
nn.Conv2d(3, 64, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU(),
nn.Conv2d(64, 128, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU(),
nn.Conv2d(128, 256, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU(),
nn.Conv2d(256, 512, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU(),
nn.Conv2d(512, 1024, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU(),
nn.Conv2d(1024, 2048, kernel_size=4, stride=2, padding=1),
nn.LeakyReLU()
)
self.adv_layer = nn.Sequential(nn.Conv2d(2048, 1, kernel_size=3, stride=1, padding=1),
nn.Sigmoid()
)
self.attr_layer = nn.Sequential(nn.Conv2d(2048, 10, kernel_size=2, stride=1, padding=0),
nn.Softmax()) # attribute classification대신 얼굴 인식 수행
def forward(self, x):
out = self.discriminator_block(x)
# out = out.view(out.shape[0], -1)
validity = self.adv_layer(out)
label = self.attr_layer(out)
return validity, label
adversarial_loss = nn.BCELoss()
attribute_loss = nn.MSELoss() # discriminator에 사용되는 attribute loss
# Initialize generator and discriminator
generator = Generator()
discriminator = Discriminator()
if cuda:
generator.cuda()
discriminator.cuda()
adversarial_loss.cuda()
attribute_loss.cuda()
# Initialize weights
generator.apply(weights_init_normal)
discriminator.apply(weights_init_normal)
# -
# ## Dataloader에서 sample batch로 test
# +
def show(img,y,color=False): #미리보기
npimg=img.numpy()
y=y.numpy()
npimg_tr=np.transpose(npimg,(1,2,0))
y_tr=np.transpose(y,(1,2,0))
plt.subplot(2,2,1)
plt.imshow(npimg_tr)
plt.subplot(2,2,2)
plt.imshow(y_tr)
OAGan_dataset = OAGandataset( paired = True, folder_numbering = False )
train_dataloader = DataLoader(OAGan_dataset,
shuffle=True,
num_workers=0,
batch_size=3) #3 batch
dataiter = iter(train_dataloader)
example_batch = next(dataiter)
# -
show(example_batch[0][0],example_batch[1][0])
print(example_batch[2][0])
show(example_batch[0][1],example_batch[1][1])
print(example_batch[2][1])
show(example_batch[0][2],example_batch[1][2])
print(example_batch[2][2])
# +
optimizer_G = torch.optim.Adam(generator.parameters(), lr=0.0001, betas=(0.5, 0.999))
optimizer_D = torch.optim.Adam(discriminator.parameters(), lr=0.0001, betas=(0.5, 0.999))
FloatTensor = torch.cuda.FloatTensor if cuda else torch.FloatTensor
LongTensor = torch.cuda.LongTensor if cuda else torch.LongTensor
# +
imgs,imgs_gt,labels=example_batch
batch_size = imgs.shape[0]
# Adversarial ground truths
valid = Variable(FloatTensor(batch_size, 1, 2, 2).fill_(1.0), requires_grad=False)
fake = Variable(FloatTensor(batch_size, 1, 2, 2).fill_(0.0), requires_grad=False)
fake_attr_gt = Variable(LongTensor(batch_size).fill_(10), requires_grad=False)
# Configure input
real_imgs = Variable(imgs.type(FloatTensor))
labels = Variable(labels.type(LongTensor))
# -----------------
# Train Generator
# -----------------
optimizer_G.zero_grad()
# Sample noise and labels as generator input
# z = Variable(FloatTensor(np.random.normal(0, 1, (batch_size, opt.latent_dim))))
# Generate a batch of images
# gen_imgs = generator(z)
print("real_imgs: ", real_imgs.shape) #x_occ
# +
def show1(img,y,color=False): #미리보기
#npimg=img.detach().numpy()
#npimg=img.numpy()
#y=y.numpy()
npimg_tr=np.transpose(img,(1,2,0))
y_tr=np.transpose(y,(1,2,0))
plt.subplot(2,2,1)
plt.imshow(npimg_tr)
plt.subplot(2,2,2)
plt.imshow(y_tr)
#out_synth[0]
# +
out_predictionM, out_InvertedM, out_synth, out_final = generator(real_imgs)
loss = sganloss([out_final,
out_predictionM,
out_InvertedM,
imgs_gt,
out_synth])
#print("gen_imgs: ", gen_imgs.shape)
print("predictM: ",out_predictionM.shape)
print("out_InvertedM: ", out_InvertedM.shape)
print("out_synth: ", out_synth.shape)
print("out_final: ", out_final.shape)
print ("imgs_gt:", imgs_gt.shape)
# -
show1(np.array(out_predictionM[0].detach()),np.array(out_InvertedM[0].detach()))
# # loss test
# +
import numpy as np
# from sgan_main import *
import torch
# from gan_model import *
import torch.nn as nn
import torchvision.models as models
# import numpy.linalg
from torch import linalg as LA
class sganloss():
# TODO: vgg_features를 loss함수마다 계산해야 해서 메모리를 엄청 잡아먹음. __init__에서 사진마다 feature 저장해주고, conv layer마다 받아오는게 아니라 pooling layer마다 feature 받아오게 수정
def __init__(self, imgs):
self.final = imgs[0]
self.M = imgs[1]
self.inverse_M = imgs[2]
self.gt = imgs[3]
self.synth = imgs[4]
# self.img = torch.randn([1, 3, 128, 128])
self.vgg16 = models.vgg16(pretrained=True).cuda()
# self.vgg16_features = [self.vgg16.features[:5],
# self.vgg16.features[:10],
# self.vgg16.features[:17],
# self.vgg16.features[:24],
# self.vgg16.features[:31]]
#self.vgg16_features = self.vgg16.features[:31]
self.final_features = self.vgg16.features[:31](self.final)
self.gt_features = self.vgg16.features[:31](self.gt)
self.sy_features = self.vgg16.features[:31](self.sy)
# self.M_features = [model(self.M) for model in self.vgg16_features]
# self.iM_features = [model(self.inverse_M) for model in range(self.vgg16_features)]
# self.final_features = [model(self.final) for model in self.vgg16_features]
# self.final_features = [model(self.final) for model in self.vgg16_features]
# self.gt_features = [model(self.gt) for model in self.vgg16_features]
# self.sy_features = [model(self.synth) for model in self.vgg16_features]
def pixel_loss(self, alpa, beta):
one = torch.matmul(self.inverse_M, (self.final - self.gt))
one = torch.norm(one, 1)
one = alpa * one.detach()
two = torch.matmul(self.M, (self.final - self.gt))
two = torch.norm(two, 1)
two = beta * two.detach()
return one + two
def smooth_loss(self):
final = self.final
M = self.M
sm = 0
for batch in range(len(final)):
for channel in range(len(final[0])):
for i in range(len(final[0][0])):
a = final[batch, channel, i, 1:]
b = final[batch, channel, i, :-1]
sm += torch.norm(a - b, 1)
c = final[batch, channel, 1:, i]
d = final[batch, channel, :-1, i]
sm += torch.norm(c - d, 1)
for i in range(len(M[0][0])):
a = final[batch, channel, i, 1:]
b = final[batch, channel, i, :-1]
sm += torch.norm(a - b, 1)
c = final[batch, channel, 1:, i]
d = final[batch, channel, :-1, i]
sm += torch.norm(c - d, 1)
return sm
def perceptual_loss(self):
pc = 0
for i in range(len(self.vgg16_features)):
a = self.sy_features[i]
b = self.gt_features[i]
c = self.final_features[i]
pc += torch.norm(a - b, 1)
pc += torch.norm(c - b, 1)
return pc
# print(vgg16_features[0](img).size(2))
def style_loss(self):
sy = 0
for i in range(len(self.vgg16_features)):
kn = 1 / (self.sy_features[i].size(1) * self.sy_features[i].size(2) * self.sy_features[i].size(3))
s = self.sy_features[i]
sT = torch.transpose(s.detach(), 2, 3)
g = self.gt_features[i]
gT = torch.transpose(g.detach(), 2, 3)
f = self.final_features[i]
fT = torch.transpose(f.detach(), 2, 3)
a = torch.matmul(sT, s) - torch.matmul(gT, g)
a = torch.norm(a, 1)
b = torch.matmul(fT, f) - torch.matmul(gT, g)
b = torch.norm(b, 1)
sy += kn * (a + b)
return sy
def l2_norm(self):
x_normalized = 0
x = self.M.detach()
for i in range(len(x)):
# print(x[i].shape)
x_normalized += LA.norm(x[i])
# x_normalized+=np.linalg.norm(x[i], axis=1, ord=2)
# print("dd")
# x_normalized =sum(x_normalized[0])
# print(x_normalized)
# norm = x.norm(p=2, dim=1, keepdim=True)
# x_normalized = x.div(norm.expand_as(x))
return x_normalized
# -
vgg16 = models.vgg16(pretrained=True)
vgg16_features = vgg16.features[:31]
final_features = vgg16_features(out_final)
# [model(out_final) for model in vgg16_features]
# gt_features = [model(imgs_gt) for model in vgg16_features]
# sy_features = [model(out_synth) for model in vgg16_features]
final_features
sum(sum(sum(sum(gt_features[4]))))
a=perceptual_loss()
a
b=pixel_loss(out_InvertedM,out_predictionM,out_final,imgs_gt,0.5,0.5)
b
def perceptual_loss():
global sy_features
global gt_features
global final_features
pc = 0
for i in range(len(vgg16_features)):
a = sy_features[i]
b = gt_features[i]
c = final_features[i]
pc += torch.norm(a-b, 1)
pc += torch.norm(c-b,1)
return pc
out_final-imgs_gt
def pixel_loss(inverse_M,M,final,gt,alpa, beta):
one = torch.matmul(inverse_M, (final - gt))
one = torch.norm(one, 1)
one = np.dot(alpa, one.detach())
two = torch.matmul(M, (final - gt))
two = torch.norm(two, 1)
two = np.dot(beta, two.detach())
return one + two
out_predictionM[0]
show1(np.array(out_synth[0].detach()),np.array(out_final[0].detach()))
#print(example_batch[2][2])
# +
#validity, _ = discriminator(out_final)
#print('validity shape: ', validity.shape)
#print('valid shape: ', valid.shape)
g_loss = 0
g_loss += w.lam1*loss.perceptual_loss(out_synth, out_final, imgs_gt)
g_loss += w.lam2*loss.style_loss(out_synth, out_final, imgs_gt)
g_loss += w.lam3*loss.pixel_loss(out_final, imgs_gt, out_InvertedM, out_predictionM, w.alpha, w.beta)
g_loss += w.lam4*loss.smooth_loss(out_final, imgs_gt, out_predictionM)
g_loss += w.lam5*loss.l2_norm(out_predictionM)
g_loss += w.lam6*loss.adversarial_loss(validity,valid)
#g_loss += w.lam1*loss.pixel_loss(out_final, imgs_gt, out_InvertedM, out_predictionM, w.alpha, w.beta)
#g_loss += w.lam2*loss.smooth_loss(out_final, imgs_gt, out_predictionM)
#g_loss += w.lam3*loss.perceptual_loss(out_synth, out_final, imgs_gt)
#g_loss += w.lam4*loss.style_loss(out_synth, out_final, imgs_gt)
#g_loss += w.lam5*loss.l2_norm(out_predictionM)
#g_loss += w.lam6*adversarial_loss(validity, valid)
#g_loss.backward()
#optimizer_G.step()
# -
g_loss
# +
imgs,imgs_gt,labels=example_batch
batch_size = imgs.shape[0]
# Adversarial ground truths
valid = Variable(FloatTensor(batch_size, 1, 2, 2).fill_(1.0), requires_grad=False)
fake = Variable(FloatTensor(batch_size, 1, 2, 2).fill_(0.0), requires_grad=False)
fake_attr_gt = Variable(LongTensor(batch_size).fill_(10), requires_grad=False)
# Configure input
real_imgs = Variable(imgs.type(FloatTensor))
labels = Variable(labels.type(LongTensor))
# -----------------
# Train Generator
# -----------------
optimizer_G.zero_grad()
# Sample noise and labels as generator input
# z = Variable(FloatTensor(np.random.normal(0, 1, (batch_size, opt.latent_dim))))
# Generate a batch of images
# gen_imgs = generator(z)
print("real_imgs: ", real_imgs.shape)
out_predictionM, out_InvertedM, out_synth, out_final = generator(real_imgs)
print("gen_imgs: ", gen_imgs.shape)
# Loss measures generator's ability to fool the discriminator
validity, _ = discriminator(gen_imgs)
print('validity shape: ', validity.shape)
print('valid shape: ', valid.shape)
g_loss += loss.perceptual_loss(out_synth, out_final, )
g_loss += adversarial_loss(validity, valid)
g_loss.backward()
optimizer_G.step()
# ---------------------
# Train Discriminator
# ---------------------
optimizer_D.zero_grad()
# d_alpha, d_beta는 discriminator에 사용되는 2가지 loss함수에 대한 가중치값으로 우리가 결정해야 하는듯
d_alpha = 0.5
d_beta = 0.5
# Loss for real images
real_pred, real_attr = discriminator(real_imgs)
# d_real_loss = (adversarial_loss(real_pred, valid) + auxiliary_loss(real_aux, labels)) / 2
d_real_loss = d_alpha * adversarial_loss(real_pred, valid) + d_beta * attribute_loss(real_attr, labels)
# Loss for fake images
fake_pred, fake_attr = discriminator(gen_imgs.detach())
# d_fake_loss = (adversarial_loss(fake_pred, fake) + auxiliary_loss(fake_aux, fake_aux_gt)) / 2
d_fake_loss = d_alpha * adversarial_loss(fake_pred, fake) + d_beta * attribute_loss(fake_attr, fake_attr_gt)
# Total discriminator loss
d_loss = (d_real_loss + d_fake_loss) / 2
# Calculate discriminator accuracy
pred = np.concatenate([real_attr.data.cpu().numpy(), fake_attr.data.cpu().numpy()], axis=0)
gt = np.concatenate([labels.data.cpu().numpy(), fake_attr_gt.data.cpu().numpy()], axis=0)
d_acc = np.mean(np.argmax(pred, axis=1) == gt)
# print('d_loss type: ', type(d_loss))
d_loss = d_loss.type(torch.FloatTensor)
d_loss.backward()
optimizer_D.step()
print(
"[Epoch %d/%d] [Batch %d/%d] [D loss: %f, acc: %d%%] [G loss: %f]"
% (epoch, 100, i, len(dataloader), d_loss.item(), 100 * d_acc, g_loss.item())
)
batches_done = epoch * len(dataloader) + i
if batches_done % 400 == 0:
save_image(gen_imgs.data[:25], "images/%d.png" % batches_done, nrow=5, normalize=True)
# net = SiameseNetwork().cuda()
# criterion = ContrastiveLoss()
# optimizer = optim.Adam(net.parameters(),lr = 0.0005 )
# counter = []
# loss_history = []
# iteration_number= 0
|
implementations/sgan/.ipynb_checkpoints/SGAN-test-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import nltk
import string
from nltk.corpus import stopwords
from nltk.tokenize import RegexpTokenizer
from nltk.stem import WordNetLemmatizer
from nltk.stem.porter import PorterStemmer
# -
url = 'http://destroyblackmagic.com/quran/data/id.indonesian.json'
df = pd.read_json(url)
df = pd.DataFrame(df)
df
data = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/dataset/quran-json-indonesia/surah/1.json')
data.head()
data = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/dataset/quran-json-english/en_translation_1.json')
data6= pd.read_csv('https://raw.githubusercontent.com/langsari/quran-dataset/master/notebook/eda/Suraiya-Project/English.csv')
data
url = 'http://destroyblackmagic.com/quran/data/id.indonesian.json'
df = pd.read_json(url)
df
df
conda install -c anaconda beautifulsoup4
import requests
from bs4 import BeautifulSoup
url = "https://quran.com/1"
web_data = requests.get(url)
print(web_data.text)
soup = BeautifulSoup(web_data.text,'html.parser')
# +
find_word = soup.find_all("h2",{"class":"text-left text-translation times-new"})
for i in find_word:
print(i)
# -
import http.client
df1 = pd.DataFrame(data)
df1
data
data.describe()
data.columns
data.drop(['name','index','count'],axis=1,inplace=True)
data
verse_df['verse']
data.shape
verses = verse_df['verse'].str.split("_", n=1, expand=True)
data['verse']
verses = data['verse'].str.split("_", n=1, expand=True)
verses.head()
verses.head()
url = "https://quran.com/1"
web_data = requests.get(url)
data2 = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/dataset/quran-json-english/en_translation_1.json')
data2
data2.columns
data.drop(['name','index','count'],axis=1,inplace=True)
data
def remove_punctuation(text):
no_punct = "".join([c for c in text if c not in string.punctuation])
return no_punct
data2['verse']
data2['verse']=data2['verse'].apply(lambda x: remove_punctuation)
data2['verse'].head()
data3 = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/dataset/quran-json-english/en_translation_1.json')
data3
data3.drop(['name','index','count'],axis=1,inplace=True)
data
data3['verse']=data3['verse'].apply(lambda x: remove_punctuation)
data3['verse'].head()
data4 = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/dataset/quran-json-english/en_translation_1.json')
data4.drop(['name','index','count'],axis=1,inplace=True)
data
data4['verse']=data4['verse'].apply(lambda x:lower()))
tokenizer=RegexpTokenizer(r'\wt')
data4['verse'] = data4['verse'].apply(lambda x: tokenizer.tokenize(x.lower()))
data4['verse'].head()
data5 = pd.read_json('https://raw.githubusercontent.com/langsari/quran-dataset/master/notebook/eda/Suraiya-Project/English.csv')
data6 = pd.read_json('https://raw.githubusercontent.com/risan/quran-json/master/json/translations/en.pretty.json')
data6.head()
data6.describe()
data6.columns
data6.drop(['surah_number','verse_number'],axis=1,inplace=True)
data6
data6.info()
data6.describe()
data7 = pd.read_json('https://raw.githubusercontent.com/risan/quran-json/master/json/translations/en.pretty.json')
data7.head()
data7.columns
data7.drop(['surah_number','verse_number'],axis=1,inplace=True)
data7
data7.head(100)
tokenizer=RegexpTokenizer(r'\wt')
data7.rename(columns={'content':'translation'},inplace=True)
data7
data7
data7['translation'].head()
data6.head()
data6
from nltk.tokenize import word_tokenize
text = "In the Name of Allah the Most Compassionate, Most Merciful."
print(word_tokenize(text))
print(word_tokenize(text))
import nltk
nltk.download('punkt')
nltk.download('punkt')
print(word_tokenize(text))
s1 = "In the Name of Allah the Most Compassionate, Most Merciful."
word_tokenize(s1)
fdist1 = FreqDist(s1)
FreqDist= s1
|
notebook/eda/Suraiya-Project/old-scrap-nltk/ enlish-translation-cleandata+nltk .ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # BottomUpParceLiNGAM
# ## Import and settings
# In this example, we need to import `numpy`, `pandas`, and `graphviz` in addition to `lingam`.
# +
import numpy as np
import pandas as pd
import graphviz
import lingam
from lingam.utils import print_causal_directions, print_dagc, make_dot
import warnings
warnings.filterwarnings('ignore')
print([np.__version__, pd.__version__, graphviz.__version__, lingam.__version__])
np.set_printoptions(precision=3, suppress=True)
# -
# ## Test data
# First, we generate a causal structure with 7 variables. Then we create a dataset with 6 variables from x0 to x5, with x6 being the latent variable for x2 and x3.
# +
np.random.seed(1000)
x6 = np.random.uniform(size=1000)
x3 = 2.0*x6 + np.random.uniform(size=1000)
x0 = 0.5*x3 + np.random.uniform(size=1000)
x2 = 2.0*x6 + np.random.uniform(size=1000)
x1 = 0.5*x0 + 0.5*x2 + np.random.uniform(size=1000)
x5 = 0.5*x0 + np.random.uniform(size=1000)
x4 = 0.5*x0 - 0.5*x2 + np.random.uniform(size=1000)
# The latent variable x6 is not included.
X = pd.DataFrame(np.array([x0, x1, x2, x3, x4, x5]).T, columns=['x0', 'x1', 'x2', 'x3', 'x4', 'x5'])
X.head()
# +
m = np.array([[0.0, 0.0, 0.0, 0.5, 0.0, 0.0, 0.0],
[0.5, 0.0, 0.5, 0.0, 0.0, 0.0, 0.0],
[0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 2.0],
[0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 2.0],
[0.5, 0.0,-0.5, 0.0, 0.0, 0.0, 0.0],
[0.5, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0],
[0.0, 0.0, 0.0, 0.0, 0.0, 0.0, 0.0]])
dot = make_dot(m)
# Save pdf
dot.render('dag')
# Save png
dot.format = 'png'
dot.render('dag')
dot
# -
# ## Causal Discovery
# To run causal discovery, we create a `BottomUpParceLiNGAM` object and call the `fit` method.
model = lingam.BottomUpParceLiNGAM()
model.fit(X)
# Using the `causal_order_` properties, we can see the causal ordering as a result of the causal discovery. x2 and x3, which have latent confounders as parents, are stored in a list without causal ordering.
model.causal_order_
# Also, using the `adjacency_matrix_` properties, we can see the adjacency matrix as a result of the causal discovery. The coefficients between variables with latent confounders are np.nan.
model.adjacency_matrix_
# We can draw a causal graph by utility funciton.
make_dot(model.adjacency_matrix_)
# ## Independence between error variables
# To check if the LiNGAM assumption is broken, we can get p-values of independence between error variables. The value in the i-th row and j-th column of the obtained matrix shows the p-value of the independence of the error variables $e_i$ and $e_j$.
p_values = model.get_error_independence_p_values(X)
print(p_values)
# ## Bootstrapping
# We call `bootstrap()` method instead of `fit()`. Here, the second argument specifies the number of bootstrap sampling.
# +
import warnings
warnings.filterwarnings('ignore', category=UserWarning)
model = lingam.BottomUpParceLiNGAM()
result = model.bootstrap(X, n_sampling=100)
# -
# ## Causal Directions
# Since `BootstrapResult` object is returned, we can get the ranking of the causal directions extracted by `get_causal_direction_counts()` method. In the following sample code, `n_directions` option is limited to the causal directions of the top 8 rankings, and `min_causal_effect` option is limited to causal directions with a coefficient of 0.01 or more.
cdc = result.get_causal_direction_counts(n_directions=8, min_causal_effect=0.01, split_by_causal_effect_sign=True)
# We can check the result by utility function.
print_causal_directions(cdc, 100)
# ## Directed Acyclic Graphs
# Also, using the `get_directed_acyclic_graph_counts()` method, we can get the ranking of the DAGs extracted. In the following sample code, `n_dags` option is limited to the dags of the top 3 rankings, and `min_causal_effect` option is limited to causal directions with a coefficient of 0.01 or more.
dagc = result.get_directed_acyclic_graph_counts(n_dags=3, min_causal_effect=0.01, split_by_causal_effect_sign=True)
# We can check the result by utility function.
print_dagc(dagc, 100)
# ## Probability
# Using the `get_probabilities()` method, we can get the probability of bootstrapping.
prob = result.get_probabilities(min_causal_effect=0.01)
print(prob)
# ## Total Causal Effects
# Using the `get_total_causal_effects()` method, we can get the list of total causal effect. The total causal effects we can get are dictionary type variable.
# We can display the list nicely by assigning it to pandas.DataFrame. Also, we have replaced the variable index with a label below.
# +
causal_effects = result.get_total_causal_effects(min_causal_effect=0.01)
# Assign to pandas.DataFrame for pretty display
df = pd.DataFrame(causal_effects)
labels = [f'x{i}' for i in range(X.shape[1])]
df['from'] = df['from'].apply(lambda x : labels[x])
df['to'] = df['to'].apply(lambda x : labels[x])
df
# -
# We can easily perform sorting operations with pandas.DataFrame.
df.sort_values('effect', ascending=False).head()
df.sort_values('probability', ascending=True).head()
# And with pandas.DataFrame, we can easily filter by keywords. The following code extracts the causal direction towards x1.
df[df['to']=='x1'].head()
# Because it holds the raw data of the total causal effect (the original data for calculating the median), it is possible to draw a histogram of the values of the causal effect, as shown below.
# +
import matplotlib.pyplot as plt
import seaborn as sns
sns.set()
# %matplotlib inline
from_index = 0 # index of x0
to_index = 5 # index of x5
plt.hist(result.total_effects_[:, to_index, from_index])
# -
# ## Bootstrap Probability of Path
# Using the `get_paths()` method, we can explore all paths from any variable to any variable and calculate the bootstrap probability for each path. The path will be output as an array of variable indices. For example, the array `[3, 0, 1]` shows the path from variable X3 through variable X0 to variable X1.
# +
from_index = 3 # index of x3
to_index = 1 # index of x0
pd.DataFrame(result.get_paths(from_index, to_index))
# -
|
examples/BottomUpParceLiNGAM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
from functools import reduce
from sklearn.dummy import DummyRegressor
from sklearn import linear_model
from sklearn.model_selection import cross_val_score
from sklearn.metrics import mean_absolute_error
from xgboost import XGBRegressor
from sklearn.metrics import r2_score
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
from scipy import stats #libraries for plotting
from scipy import signal
import warnings
warnings.filterwarnings(action='ignore')
# -
# ### 2.1 Import data from CSV into pandas dataframe
pdata = pd.read_csv("./Programming_Part1_TTI_Challenge.csv", sep='\t')
stats = pd.DataFrame(pdata.iloc[:,2:].describe())## gather stastics for last two columns
gm = lambda n: reduce(lambda x,y: x*y, n) ** (1.0 / len(n)) ## Function to calculate geometric mean
stats.loc[8] = [gm(pdata.X) , gm(pdata.Y)]
pdata.head()
# ### 2.2 Print all data stastics in one table
stats.rename(index={8: 'geo.mean'}) ## Last row for geometric mean
# +
from scipy import stats
# %matplotlib inline
plt.figure(figsize=(8,8)) #set size
sns.distplot((pdata.X),color="red", label="X" ,kde=False, fit=stats.gamma) #try to approximate a distribution over data with fit()
sns.distplot((pdata.Y),color="green",label="Y", kde=False, fit=stats.gamma)
plt.ylabel('Y', fontsize=12)
plt.xlabel("X", fontsize=12)
plt.show()
# +
plt.figure(figsize=(14,10)) #set size
g = sns.jointplot(x="X", y="Y", data=pdata, kind="kde"); ## use KDE method to get more idea about the data
g.plot_joint(plt.scatter, c="w", s=30, linewidth=.5, marker="*") ## plot a line in data with all the data points
g.ax_joint.collections[0].set_alpha(0)
plt.ylabel('Y', fontsize=12)
plt.xlabel("X", fontsize=12)
plt.show()
# -
# #### From the plots we can see that the two features are highly correlated and form a clear function curve.
# #### 2.3 To write out data to a table, it can be done in many ways but for this exercise I am sqlite db since it is a lightweight and minimal to setup. The connection part can be replace with any database details/jdbc connections, such as Oracle or MySQL.
#
#
import sqlite3
conn = sqlite3.connect("new.db")
pdata.to_sql("p1", conn, if_exists="replace")
# #### The code also demonstrate that we can convert the data into a RDBMS table and fetch it back.
cur = conn.cursor()
cur.execute("select * from p1 limit 5;")
results = cur.fetchall()
print(results)
conn.commit()
conn.close()
# #### We can convert the Class feature to a categorical one.
pdata.nunique() #find number of unique categories
pdata.Class.value_counts()
pdata.Class = pdata["Class"].astype("category")
X_cat = pd.get_dummies(pdata , columns=["Class"])
X_cat.drop('Y', 1,inplace=True)
from sklearn.model_selection import train_test_split
X_cat.set_index("PartNo", inplace=True)
X_train, X_test, y_train, y_test = train_test_split(X_cat, pdata["Y"], test_size=0.33, random_state=2111)
# Keep 2/3 of the data for training and 1/3 for testing model performance
# ### 2.4 Predictive Model
# ### Establish a baseline with a dummy regressor, so we know that we are doing better than predicting mean.
model = DummyRegressor(strategy='mean')
model.fit(X_train, y_train)
Y_pred=model.predict(X_test)
print(mean_absolute_error(y_test,Y_pred))
X_train.head()
# +
model = linear_model.LinearRegression()
model.fit(X_train.loc[:,"X"].reshape([-1,1]), y_train)
Y_pred=model.predict(X_test.loc[:,"X"].reshape([-1,1]))
print("MAE", mean_absolute_error(y_test,Y_pred)) #MAE difference between observation
print("R2_score", r2_score(y_test,Y_pred, multioutput='variance_weighted')) #R2 models explained variance
# +
from sklearn.preprocessing import PolynomialFeatures
from sklearn import linear_model
for i in range(2,4):
poly = PolynomialFeatures(degree=i)
X_ = poly.fit_transform(X_train.loc[:,"X"].reshape([-1,1])) # 1-D data , needs to be reshaped to trick sklearn
predict_ = poly.fit_transform( X_test.loc[:,"X"].reshape([-1,1]) )
clf = linear_model.LinearRegression()
clf.fit(X_, y_train)
y_pred = clf.predict(predict_)
print("\n score with polynomial degree ", i)
print("MAE ", mean_absolute_error(y_test,y_pred))
print("R2_score", r2_score(y_test,y_pred, multioutput='variance_weighted'))
# -
# ### From plots we could see that the scatterplot has a very clear function curve, this can be leveraged by using a gradient based learner.
# +
from sklearn.metrics import mean_squared_error
from sklearn.ensemble import GradientBoostingRegressor
est = GradientBoostingRegressor(n_estimators=160, learning_rate=0.09,
max_depth=1, random_state=0, loss='ls').fit(X_train.loc[:,"X"].reshape([-1,1]), y_train)
print("MAE", mean_absolute_error(y_test, est.predict(X_test.loc[:,"X"].reshape([-1,1]))))
print("R2_score",r2_score(y_test,est.predict(X_test.loc[:,"X"].reshape([-1,1])), multioutput='variance_weighted'))
# -
# #### 2.5 Assess the accuracy of your predictive model
#
# Both gradientboosting and polynomial regression models have resulted in much better performance with near perfect R2 score and low mean absolute error.
# The reason that simple linear regression did not work so well here is, the simple linear model could not generalize/model the curve function so well. Where as in case of polynomial and Boosting, since the data curve represented a polunomial/differentiable function, at which polynomial/gradient models perform well.
#
# The low error means that the predictions by this model would only be off by a 0.05 of the actual values. High R2 square suggests that 99.8% of the data can be explained by the model
#
#
# Also, X is the most important feature for predicting the target value. While part number has a very little contribution.
# # 3 SQL Questions
# #### Write the SQL statements to:
# 1. Display Full_MFG_Name in Table B without the MFG Code ( Example: ‘Amphenol’)
# 2. Calculate Total Revenue from Table B
# 3. Display the top 10 Products from Table B which made highest profit
# 4. Display total cost, total Price and Margins grouped by Parent_MFG in table A
# 5. Display the highest selling product and the second highest selling product
# 6. Display the Total Cost and Total Revenue based on Type from Table C and order it in a descending order
# 7. Find which Quarter sold highest number of products
# 8. Find which quarter made the highest sale in ‘AUTOMOTIVE’ category In the last year
# 9. Find the Products in table C that haven’t sold
# +
s = "../MFG.txt"
conn = sqlite3.connect("new.db")
def get_dtype(s): #get data type with regex (is float?)
import re
if re.match("^\d+?\.\d+?$", s) is None:
if s.isdigit():
return "int"
else:
return "string"
return "float"
catFeatures = []
numFeatures = []
def process_dtypes(df): #read data, and assign data types accordingly
for col,x in df.iloc[1,:].iteritems():
t = get_dtype(str(x).strip())
if(t == "int" or t == "float" ):
numFeatures.append(col)
if t == "int":
df[col] = df[col].astype(int)
else:
df[col] = df[col].astype(float)
else:
catFeatures.append(col)
df[col] = df[col].astype(str)
return df
def file_to_table(fn, cols, conn=conn): #helper function to read data from a file and return a dataframe from it.
table = []
tname = fn.split(".")[2].strip("/")# table name from filename
f = open(fn, 'r')
for i in f:
table.append(i.strip()) #build a nested list of table rows
table = np.reshape(table, [-1,cols]) # reshape to build table
columns = table[0,:]
data = table[1:,:]
temp = pd.DataFrame(data, columns=columns) #convert into a dataframe
process_dtypes(temp)
temp.to_sql(tname, conn, if_exists="replace") # commit to a database
conn.commit()
f.close()
return temp
# +
MFG = file_to_table('../MFG.txt',4)
conn = sqlite3.connect("new.db")
products = file_to_table('../product_table.txt',5)
conn = sqlite3.connect("new.db")
sales = file_to_table('../sales_table.txt',7)
conn = sqlite3.connect("new.db")
# -
# #### 1. Display Full_MFG_Name in Table B without the MFG Code ( Example: ‘Amphenol’)
# ``` mysql
# select RIGHT(a.Full_MFG_Name, LENGTH(a.Full_MFG_Name) -6) as Name , b.* from MFG a, sales_table b where a.MFG_Code = b.MFG_code;
#
# Query #1 (Executed in 4ms)
# name mfg_code product quantity unit_price vaunitprice1 unit_cost dates
# KEMET KEM N 100 18.43 0.03 13.02 11/4/2017
# KEMET KEM D 100 18.43 0.03 13.02 21/4/2016
# KEMET KEM J 16 3.31 0.0001 2.45 26/8/2017
# KEMET KEM K 1700 0.545 0.072 0.44 6/9/2017
# KEMET KEM E 150 0.8 0.1666 0.21 4/12/2017
# Amphenol APH I 5 15.49 0.35 13.8618 15/5/2017
# Amphenol APH H 5 16.33 0.35 15.2708 19/8/2017
# .
# .
# .
# .
# TE Connectivity/Raychem Tubing RAY K 5000 0.03 0.0016 0.0172 5/2/2016
#
# ```
# #### 2. Calculate Total Revenue from Table B
# #### Total revenue
# ``` mysql
# select sum(quantity * unit_price) as revenue from sales_table;
#
# revenue
# 22595.3201681077
#
# ```
#
# #### Total Profit
# ``` mysql
# select sum((quantity * unit_price) - (quantity * unit_cost)) as profit from sales_table order by profit desc LIMIT 10;
# profit
# 4281.87903094292
# ```
# #### 3. Display the top 10 Products from Table B which made highest profit
#
# ``` mysql
#
# select product, sum((quantity * unit_price) - (quantity * unit_cost)) as profit from sales_table
# group by product
# order by profit desc LIMIT 10;
#
#
# product profit
# L 1068.800573349
# N 540.999984741211
# D 540.999984741211
# C 470.559993743896
# K 402.499992772937
# I 339.161009788513
# O 250
# E 248.499969393015
# H 165.295968055725
# F 91.1389923095703
#
# ```
#
# #### 4. Display total cost, total Price and Margins grouped by Parent_MFG in table A
#
#
#
# ``` mysql
# select a.Parent_MFG , sum(b.Unit_Cost) as Unit_Cost, sum(b.Unit_Price) as Unit_price, (sum( (b.quantity * b.unit_price) - (b.quantity * b.unit_cost) )/sum(b.quantity * b.unit_price) )*100 as profit_margin_pct
# from MFG a, sales_table b
# where a.MFG_Code = b.MFG_code
# group by a.Parent_MFG
# order by a.Parent_MFG;
#
#
# Query #4 (Executed in 3ms)
# parent_mfg | unit_cost | unit_price | profit_margin_pct|
# ACG | Amphenol Connector Group 970.132 1031.99 9.20290857973324
# AVC | AVX Corporation 59.7 75.85 14.0426265605136
# KCC | KOA Corporation 0.75 0.87 13.793100967702
# KCO | Kemet Corporation 29.14 41.515 28.4770947263481
# TEG | TE Connectivity Group 112.814 224.77 48.3635106304724
# ```
# #### 5. Display the highest selling product and the second highest selling product
#
# ```mysql
#
# select product,sum(quantity*unit_Cost) as sale from sales_table
# group by product
# order by sale desc limit 2;
#
# product sale
# L 7915.79961776733
# K 1833.99999886751
#
#
# ```
# #### 6. Display the Total Cost and Total Revenue based on Type from Table C and order it in a descending order
#
# Without the nested query, the values returned would consider duplicate values (x4 times) for product K
#
# ```mysql
# select
# abc.type as Type,
# sum(abc.total_revenue) as total_revenue,
# sum(abc.total_cost) as total_cost
# from
# (
# select
# distinct pt.product as product,
# pt.type as type,
# (st.quantity * st.unit_price) as total_revenue,
# (st.quantity * st.unit_cost) as total_cost
# from product_table pt
# inner join sales_table st
# on(st.product = pt.product)
# ) as abc
# group by abc.type
# order by abc.type desc;
#
# type total_revenue total_cost
# EREL 16534.8101039827 14065.0290335119
# COMM 4141.51003456116 2879.95005607605
# ```
# #### 7 Find which Quarter sold highest number of products
#
# ```mysql
#
# select EXTRACT(YEAR from to_date(dates,'DD/MM/YYYY')) as YEAR, EXTRACT(QUARTER from to_date(dates,'DD/MM/YYYY')) as quart, sum(quantity) as Number_of_products
# from sales_table
# group by year,quart
# order by Number_of_products desc LIMIT 1;
#
# year quart number_of_products
# 2017 1 5015
#
# ```
# #### 8. Find which quarter made the highest sale in ‘AUTOMOTIVE’ category In the last year
# ```mysql
# select stt.years, stt.Quarter_num, pt.category, sum(st.quantity * st.Unit_cost) as sale
# from sales_table st , product_table pt,
# (select EXTRACT(YEAR from to_date(dates,'DD/MM/YYYY')) as years, EXTRACT(QUARTER from to_date(dates,'DD MM YYYY')) as Quarter_num from sales_table) as stt
# where st.product = pt.product
# and pt.category ='AUTOMOTIVE'
# and stt.years = EXTRACT(year from CURRENT_DATE) -1
# group by stt.years, stt.Quarter_num ,pt.category
# order by sale desc limit 1 ;
#
# years quarter_num category sale
# 2017 3 AUTOMOTIVE 10210.0002765656
#
# ```
#
#
# #### 9. Find the Products in table C that haven’t sold anything ever
# ```mysql
# select * from product_table where product not in (select product from sales_table);
#
# product class commodity category type
# P 564 C/P AUTOMOTIVE COMM
# ```
# ## Part 4
# ### 4.1 In Python (or Pandas) write a code to import the transaction table
import pandas as pd
p3 = pd.read_excel(open("./ModelingDataSet.xlsx",'rb'))
# or using sheet index starting 0
p3.set_index("Transaction_ID", inplace=True)
p3.columns = [i.strip().lower() for i in p3.columns]
p3.head()
# #### A quick exploratory analysis of data.
# +
# %matplotlib inline
sns.set(rc={'figure.figsize':(10,8)})
corr = p3.corr()
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
# -
corr[abs(corr) > .3]
# #### From the correlation plot we can see that the extended_cost has very high correlation to revenue, which makes sense, higher extended cost will result in higher revenue. It also has some correlation to margin, when extended costs go up, marging goes down.
#
# #### A Quick look at distributions of all the features
g = sns.pairplot(p3, kind="reg")
sns.set(rc={'figure.figsize':(8,8)})
# #### From the plots we can see that the features do not follow any partucular distributions. Also many features contain extreme outlier values that coud impact our analysis.
#
# #### Some ireatment might be required to make this data more modelable.
# #### Performing log transformation could be an option
g = sns.pairplot(np.log(p3), kind="reg")
sns.set(rc={'figure.figsize':(8,8)})
# #### Logged version of the data looks much better, we can look at the histograms can see differentiable bins now!
# #### We can also process the outliers in this data, the extreme outliers can effect the binning algorithms. Since, the algorithms will have to compensate for the variance and mean introduced by the extreme values.
#
# #### We will test with both processes and unprocessed data.
#
# #### Below code will remove top and bottom 1 percentile values from data and take log transformation of data and return new dataframe
#
#
# +
import itertools
outliers_lst = []
log_data = p3.copy()
# For each feature find the data points with extreme high or low values
for feature in log_data.columns:
pdwn = np.percentile(log_data.loc[:, feature], 1) #values in bottom 1 percentile
pup = np.percentile(log_data.loc[:, feature], 99) #values in upper 1 percentile
#Using the interquartile range to calculate an outlier step (1.5 times the interquartile range)
step = 1.5 * (pup - pdwn)
# The tilde sign ~ means not
# So here, we're finding any points outside of Q1 - step and Q3 + step
outliers_rows = log_data.loc[~((log_data[feature] >= pdwn - step) & (log_data[feature] <= pup + step)), :]
print("Outliers for the feature '{}':".format(feature), len(outliers_rows[feature]))
outliers_lst.append(list(outliers_rows.index))
outliers = list(itertools.chain.from_iterable(outliers_lst))
uniq_outliers = list(set(outliers))
# List of duplicate outliers
dup_outliers = list(set([x for x in outliers if outliers.count(x) > 1]))
print( 'Total Number of outliers:\n', len(uniq_outliers))
# Remove duplicate outliers
# Only 5 specified
processed_data = log_data.drop(log_data.index[dup_outliers]).reset_index(drop = True)
processed_data = np.log(processed_data)
# Original Data
print( 'Original shape of data:\n', p3.shape)
# Processed Data
print( 'New shape of data:\n', processed_data.shape)
# -
# ### 4.2.1 In Python (or Pandas), write a code that will cluster the extended costs into bins
# ### Premise for algorithms
#
# #### To segment data in optimal bins we can consider multiple approaches. The best thing about the data here, since we are performing 1D segmentation we can leverage the fact that data can be fully sorted and processes. (This makes k-means not so optimal approach for this problem)
#
# #### After some research, I found some best fits for this problem ,
#
# ### The Jenks optimization method
#
# #### https://en.wikipedia.org/wiki/Jenks_natural_breaks_optimization
# Also called the Jenks natural breaks classification method, is a data clustering method designed to determine the best arrangement of values into different classes. This is done by seeking to minimize each class’s average deviation from the class mean, while maximizing each class’s deviation from the means of the other groups.
#
# #### The optimization technique for this algorithm is exactly what we are looking for,
# #### Algorithm
# Calculate the sum of squared deviations between classes (SDBC).
# Calculate the sum of squared deviations from the array mean (SDAM).
# Subtract the SDBC from the SDAM (SDAM-SDBC). This equals the sum of the squared deviations from the class means (SDCM).
# After inspecting each of the SDBC, a decision is made to move one unit from the class with the largest SDBC toward the class with the lowest SDBC.
#
# Finally, the goodness of variance fit (GVF) is calculated. GVF is defined as (SDAM - SDCM) / SDAM. GVF ranges from 0 (worst fit) to 1 (perfect fit).
#
#
#
#
#
# ### CK-Means
#
# #### reference paper https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5148156/
#
# Fast optimal univariate clustering and segementation by dynamic programming. Three types of problem including univariate k-means, k-median, and k-segments are solved with guaranteed optimality and reproducibility.
# The core algorithm minimizes the sum of within-cluster distances using respective metrics. Its advantage over heuristic clustering algorithms in efficiency and accuracy is increasingly pronounced as the number of clusters k increases. An auxiliary function generates histograms that are adaptive to patterns in data.
#
#
#
# ### KDE methods
#
# We can look at distribution densities, and use the maxim and minimas to create bins, or atleast to get estimates.
# ### KDE Method
# +
from numpy import array, linspace
from sklearn.neighbors.kde import KernelDensity
from matplotlib.pyplot import plot
a = p3.extended_cost.values.reshape(-1, 1)
kde = KernelDensity(kernel='gaussian', bandwidth=np.mean(p3.extended_cost)).fit(a)
s = linspace(min(p3.extended_cost),max(p3.extended_cost))
e = kde.score_samples(s.reshape(-1,1))
plot(s, e)
# -
from scipy.signal import argrelextrema
mi, ma = argrelextrema(e, np.less)[0], argrelextrema(e, np.greater)[0]
print("Minima:", s[mi]) # get minimas in densities for creating breaks
print( "Maxima:", s[ma]) #
print("nmber of bins", len(s[mi])+2)
splits = a[a < mi[0]], a[(a >= mi[0]) * (a <= mi[1])], a[a >= mi[1]]
plot(s[:mi[0]+1], e[:mi[0]+1], 'r',
s[mi[0]:mi[1]+1], e[mi[0]:mi[1]+1], 'g',
s[mi[1]:], e[mi[1]:], 'b',
s[ma], e[ma], 'go',
s[mi], e[mi], 'ro')
# ### Jenks natural breaks
# +
from jenks import jenks
import numpy as np
def goodness_of_variance_fit(array, classes):
classes = jenks(array, classes)
classified = np.array([classify(i, classes) for i in array])
#print(classified)
array = p3["margin%"].values
maxz = max(classified)
# nested list of zone indices
zone_indices = [[idx for idx, val in enumerate(classified) if zone + 1 == val] for zone in range(maxz)]
# Calculate the sum of squared deviations from the array mean (SDAM).
sdam = np.sum((array - array.mean()) ** 2)
# sorted polygon stats
array_sort = [np.array([array[index] for index in zone]) for zone in zone_indices]
# Calculate the sum of squared deviations between classes (SDBC).
sdcm = sum([np.sum((classified - classified.mean()) ** 2) for classified in array_sort])
# goodness of variance fit
gvf = (sdam - sdcm) / sdam
return gvf, classes
def classify(value, breaks):
for i in range(1, len(breaks)):
if value < breaks[i]:
return i
return len(breaks) - 1
# +
gvf = 0
gvfs = []
nclasses = 2
for nclasses in range(3,15):
gvf, cls = goodness_of_variance_fit(p3["extended_cost"].values, nclasses)
gvfs.append(gvf)
plt.plot(range(3,15), gvfs )
plt.xlabel="number_of_bins"
plt.xlabel="ratio_of_variance_to_mean_difference"
# -
# #### For jenks metrc was variance across clusters/variance within clusters. 0 being worst fit, 1 being best fit
# #### We can see that after 9-10 bins the metric does not improve much.
#
# ### Metric
#
# I extracted variance within cluster and average difference across clusters from data(SSW, SSB, SST) and generated stastics for both processed and unprocessed data in vars_stats function
# #.
#
# SSW - sum squared within cluster
#
# SSB - sum squared between cluster means
#
# SST - Total variance
#
# For calculations I have used a ratio on variance within to variance across. This fits the problem discription well.
#
# #### The function stat_n_bins takes in data frame, uses ckmeans, jenks and kmeans and returns statistics sorted by the metric. Optionally can also perform plotting for data.
#
# ex.
#
# stat_n_bins(range(3,12),["extended_cost", "margin%"], processed_data, lg=True)
#
# arg 1 - bin range to traverse
#
# arg 2 - primary and secondary feature (primary to create segments, secondary to find stats in those bins )
#
# arg 3 - dataframe to use
#
# arg 4 - was a log transformed dataframe passed
#
# #### All the stats were calculated with data in its original shape (exp ws taken while calculating stats)
#
# +
from sklearn.cluster import KMeans
from jenks import jenks #https://en.wikipedia.org/wiki/Jenks_natural_breaks_optimization
import ckmeans #https://journal.r-project.org/archive/2011-2/RJournal_2011-2_Wang+Song.pdf
import os
from contextlib import redirect_stdout
def vars_stats(df, method_col, feature,lg):
if lg==True:
df = np.exp(df)
k = len(df[feature].value_counts())
N = len(df.values)
n = df.groupby(method_col).size()
SSqX = sum([value**2 for value in df[feature]])
SqX = sum(df[feature])**2
SStotal = SSqX - (SqX/N)
SSbetween = sum(df.groupby(by=method_col).sum()[feature]**2 /n) - (SqX/N)
SSwithin = SStotal - SSbetween
return SSwithin, SSbetween
def stat_n_bins(n_bins, features, df,v=False, p=False, lg=False ):
stats = []
for i in n_bins:
n_bin = i
array = df[features[0]] ##feature
# generate labels
jclasses = jenks(array, n_bin)
kmeans = KMeans(n_clusters=n_bin, n_jobs=-1, precompute_distances=True)
reshaped = array.values.reshape([-1,1])
kmeans.fit(reshaped)
y_kmeans = kmeans.predict(reshaped)
k_classes = kmeans.labels_
with open(os.devnull, 'w') as devnull: #supress output
with redirect_stdout(devnull):
ck_classes = ckmeans.ckmeans(array,k=n_bin)[0]
# add assignment column to dataframe
df["jclasses"] = np.array([classify(i, jclasses) for i in array])
df["k_classes"] = k_classes
df["ck_classes"] = ck_classes
#gather statistics
jwvar, jbvar = vars_stats(df, "jclasses", features[1],lg) #stats are based on Margin
kwvar, kbvar = vars_stats(df, "k_classes", features[1], lg)
ckwvar, ckbvar = vars_stats(df, "ck_classes", features[1],lg)
# append stats
stats.append([n_bin,kwvar,kbvar,"kmeans"])
stats.append([n_bin,jwvar,jbvar, "jenk"])
stats.append([n_bin,ckwvar,ckbvar,"ckmeans"])
## Plotting Flag, plot segments formeda for each iteration depending upon p flag
if p==True:
sns.lmplot('margin%', 'extended_cost',
data=df.sample(10000),
fit_reg=False,
hue="ck_classes",
scatter_kws={"marker": "D", "s": 50})
plt.title("extended_cost vs margin")
plt.ylabel("extended_cost")
#plt.xlabel("margin")
plt.show()
if v == True: #verbose flag
print("\n Stats when number of bins is" , n_bin)
print("\nUsing Jenks")
print("Mean difference in margin across bins",jbvar )
print("Total variance within bins ", jwvar)
print("\nUsing K-Means")
print("Mean difference in margin across bins", kbvar )
print("Total variance within bins ", kwvar )
print("\nUsing 1D Optimal CKMeans")
print("Mean difference in margin across bins", ckbvar )
print("Total variance within bins ", ckwbvar )
else:
print("."*n_bin) #progress bar
# generate df of stastics
dstats = pd.DataFrame(stats)
dstats.columns = ["n_bin","var_within", "var_between", "method"]
dstats["ssb/sst"] = dstats["var_between"]/(dstats["var_between"]+dstats["var_within"])
dstats.sort_values(by="ssb/sst",ascending=False, inplace=True)
return dstats
# -
eng_data_stats = stat_n_bins(range(3,12),["extended_cost", "margin%"], processed_data, lg=True)
orig_stats = stat_n_bins(range(3,12),["extended_cost", "margin%"], p3)
# ### 4.2.2 Show your solution when the number of bins (nb_b) from 3 to 10 and measure the accuracy
# #### Some plotting on statistics, to understand the results better.
# #### First two plots show
# 1 variance between clusters # look for highest
#
# 2 variance within clusters, look for lowest
g = sns.factorplot(x="n_bin", y="var_between",
hue="method",
data=orig_stats, kind="bar",
size=6, aspect=.8);
g = sns.factorplot(x="n_bin", y="var_within",
hue="method",
data=orig_stats, kind="bar",
size=6, aspect=.8);
# #### We can see that around 7 number of bins the ration is optimal.
# Next plot shows the number of bins vs accuracy with the number of bins used.
# We can see that when original unprocessed data is used, a lower metric was achieved. but also with higher number of bins. Reason for this lower metric could be the impact of outliers
# Extra bins were needed to accomodate outliers
#
#
# For processed data a higher accuracy was achieved (0.6) with best values around 7 bins.
#
sns.lmplot(x="n_bin",y="ssb/sst", fit_reg=False, hue="method", data = orig_stats, size=7) #original
sns.lmplot(x="n_bin",y="ssb/sst", fit_reg=False, hue="method", data = eng_data_stats,size=7) #processed data
# ### Visualizing the optimal clusters
stat_n_bins(range(9,10),["extended_cost", "margin%"], p3,p=True)
stat_n_bins(range(7,8),["extended_cost", "margin%"], processed_data,p=True, lg=True)
# #### We can see the clear clusters in transformed data(thus the reason for higher accuracy) while in unprocessed data it is a little messy since most values are in bottom.
# #### Final bins
# +
## Finally what are the actual bins?
print("Original Data")
print(ckmeans.ckmeans(p3.extended_cost,k=7)[0])
print(jenks(p3.extended_cost,7))
print("processed_data Data")
print(ckmeans.ckmeans(processed_data.extended_cost,k=7)[0])
print(jenks(processed_data.extended_cost,7))
# -
# ### 4.2.3 What column in the Transaction table could help you to differentiate the margins even better?
#
# #### Since we are aiming to segment margins and not predict a feature's value, classifier/regressor models are not very useful here.
#
# #### Since the functions created earlier used extended_cost for segmentation, and used margin to calculate metric. To find out other useful feature we can can replace extended cost iteratively. Since, it was already written as a function, it will be simpler to do so.
#
ls = ['unit_cost', 'quantity', 'revenue']
for i in ls:
df = stat_n_bins(range(3,12),[i, "margin%"], processed_data, lg=True)
print(df.head(2))
ls = ['unit_cost', 'quantity', 'revenue']
for i in ls:
df = stat_n_bins(range(3,12),[i, "margin%"], p3, lg=True)
print(df.head(2))
#
# #### After using other columns as a segmentation feature to functions above and generating statistics again.
# #### We can see that, none of the other features are able to return better statistics than previously used feature, "extended_cost". Revenue is a distant second, with accuracy close at .40 (last set of values) #for both original and processed data.
#
# #### To leverage any information contained in revenue feature, We could derive a ratio (or some other form of interaction feature) from a combination of revenue and extended cost to better segment the margins.
|
EDA_Segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow2_p36)
# language: python
# name: conda_tensorflow2_p36
# ---
# # Building your own TensorFlow container from Amazon SageMaker Studio
#
# **STUDIO KERNEL NOTE:** If you are prompted for Kernel, choose 'Python 3 (TensorFlow CPU Optimized)
#
# With Amazon SageMaker, you can package your own algorithms that can then be trained and deployed in the SageMaker environment. This notebook guides you through an example using TensorFlow that shows you how to build a Docker container for SageMaker and use it for training and inference.
#
# This notebook contains a modified version of the existing [Tensorflow Bring Your Own](https://github.com/awslabs/amazon-sagemaker-examples/tree/master/advanced_functionality/tensorflow_bring_your_own) notebook created to run on Amazon SageMaker Notebook Instances. Because the underlying architecture between Amazon SageMaker Notebook Instances and Amazon SageMaker Studio Notebooks is different, this notebook is created specifically to illustrate a bring-your-own scenario within Amazon SageMaker Studio using the [SageMaker Studio Image Build CLI](https://github.com/aws-samples/sagemaker-studio-image-build-cli/blob/master/README.md)
#
# By packaging an algorithm in a container, you can bring almost any code to the Amazon SageMaker environment, regardless of programming language, environment, framework, or dependencies.
#
# 1. [Building your own TensorFlow container](#Building-your-own-tensorflow-container)
# 1. [When should I build my own algorithm container?](#When-should-I-build-my-own-algorithm-container?)
# 1. [Permissions](#Permissions)
# 1. [The example](#The-example)
# 1. [The workflow](#The-workflow)
# 1. [Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker](#Part-1:-Packaging-and-Uploading-your-Algorithm-for-use-with-Amazon-SageMaker)
# 1. [An overview of Docker](#An-overview-of-Docker)
# 1. [How Amazon SageMaker runs your Docker container](#How-Amazon-SageMaker-runs-your-Docker-container)
# 1. [Running your container during training](#Running-your-container-during-training)
# 1. [The input](#The-input)
# 1. [The output](#The-output)
# 1. [Running your container during hosting](#Running-your-container-during-hosting)
# 1. [The parts of the sample container](#The-parts-of-the-sample-container)
# 1. [The Dockerfile](#The-Dockerfile)
# 1. [Building and registering the container using the sagemaker-docker CLI](#Building-and-registering-the-container)
# 1. [Testing your algorithm on your local machine](#Testing-your-algorithm-on-your-local-machine)
# 1. [Part 2: Training and Hosting your Algorithm in Amazon SageMaker](#Part-2:-Training-and-Hosting-your-Algorithm-in-Amazon-SageMaker)
# 1. [Set up the environment](#Set-up-the-environment)
# 1. [Create the session](#Create-the-session)
# 1. [Upload the data for training](#Upload-the-data-for-training)
# 1. [Training On SageMaker](#Training-on-SageMaker)
# 1. [Optional cleanup](#Optional-cleanup)
# 1. [Reference](#Reference)
#
# _or_ I'm impatient, just [let me see the code](#The-Dockerfile)!
#
# ## When should I build my own algorithm container?
#
# You may not need to create a container to bring your own code to Amazon SageMaker. When you are using a framework such as Apache MXNet or TensorFlow that has direct support in SageMaker, you can simply supply the Python code that implements your algorithm using the SDK entry points for that framework. This set of supported frameworks is regularly added to, so you should check the current list to determine whether your algorithm is written in one of these common machine learning environments.
#
# Even if there is direct SDK support for your environment or framework, you may find it more effective to build your own container. If the code that implements your algorithm is quite complex or you need special additions to the framework, building your own container may be the right choice.
#
# Some of the reasons to build an already supported framework container are:
# 1. A specific version isn't supported.
# 2. Configure and install your dependencies and environment.
# 3. Use a different training/hosting solution than provided.
#
# This walkthrough shows that it is quite straightforward to build your own container. So you can still use SageMaker even if your use case is not covered by the deep learning containers that we've built for you.
#
# ## Permissions
#
# Running this notebook requires permissions in addition to the normal `SageMakerFullAccess` execution role permissions. This is because it:
#
# 1. Creates a new repository and pushes built images to [Amazon Elastic Container Registry](https://aws.amazon.com/ecr/)
# 2. Utilizes [AWS Code Build](https://aws.amazon.com/codebuild/) to build new docker images
#
#
# ## The example
#
# In this example we show how to package a custom TensorFlow container with Amazon SageMaker studio with a Python example which works with the CIFAR-10 dataset and uses TensorFlow Serving for inference. However, different inference solutions other than TensorFlow Serving can be used by modifying the docker container.
#
# In this example, we use a single image to support training and hosting. This simplifies the procedure because we only need to manage one image for both tasks. Sometimes you may want separate images for training and hosting because they have different requirements. In this case, separate the parts discussed below into separate Dockerfiles and build two images. Choosing whether to use a single image or two images is a matter of what is most convenient for you to develop and manage.
#
# If you're only using Amazon SageMaker for training or hosting, but not both, only the functionality used needs to be built into your container.
#
# [CIFAR-10]: http://www.cs.toronto.edu/~kriz/cifar.html
#
# ## The workflow
#
# This notebook is divided into two parts: _building_ the container and _using_ the container.
# # Part 1: Packaging and Uploading your Algorithm for use with Amazon SageMaker
#
# ### An overview of Docker
#
# If you're familiar with Docker already, you can skip ahead to the next section.
#
# For many data scientists, Docker containers are a new technology. But they are not difficult and can significantly simply the deployment of your software packages.
#
# Docker provides a simple way to package arbitrary code into an _image_ that is totally self-contained. Once you have an image, you can use Docker to run a _container_ based on that image. Running a container is just like running a program on the machine except that the container creates a fully self-contained environment for the program to run. Containers are isolated from each other and from the host environment, so the way your program is set up is the way it runs, no matter where you run it.
#
# Docker is more powerful than environment managers like conda or virtualenv because (a) it is completely language independent and (b) it comprises your whole operating environment, including startup commands, and environment variables.
#
# A Docker container is like a virtual machine, but it is much lighter weight. For example, a program running in a container can start in less than a second and many containers can run simultaneously on the same physical or virtual machine instance.
#
# Docker uses a simple file called a `Dockerfile` to specify how the image is assembled. An example is provided below. You can build your Docker images based on Docker images built by yourself or by others, which can simplify things quite a bit.
#
# Docker has become very popular in programming and devops communities due to its flexibility and its well-defined specification of how code can be run in its containers. It is the underpinning of many services built in the past few years, such as [Amazon ECS].
#
# Amazon SageMaker uses Docker to allow users to train and deploy arbitrary algorithms.
#
# In Amazon SageMaker, Docker containers are invoked in a one way for training and another, slightly different, way for hosting. The following sections outline how to build containers for the SageMaker environment.
#
# Some helpful links:
#
# * [Docker home page](http://www.docker.com)
# * [Getting started with Docker](https://docs.docker.com/get-started/)
# * [Dockerfile reference](https://docs.docker.com/engine/reference/builder/)
# * [`docker run` reference](https://docs.docker.com/engine/reference/run/)
#
# [Amazon ECS]: https://aws.amazon.com/ecs/
#
# ### How Amazon SageMaker runs your Docker container
#
# Because you can run the same image in training or hosting, Amazon SageMaker runs your container with the argument `train` or `serve`. How your container processes this argument depends on the container.
#
# * In this example, we don't define an `ENTRYPOINT` in the Dockerfile so Docker runs the command [`train` at training time](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html) and [`serve` at serving time](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-inference-code.html). In this example, we define these as executable Python scripts, but they could be any program that we want to start in that environment.
# * If you specify a program as an `ENTRYPOINT` in the Dockerfile, that program will be run at startup and its first argument will be `train` or `serve`. The program can then look at that argument and decide what to do.
# * If you are building separate containers for training and hosting (or building only for one or the other), you can define a program as an `ENTRYPOINT` in the Dockerfile and ignore (or verify) the first argument passed in.
#
# #### Running your container during training
#
# When Amazon SageMaker runs training, your `train` script is run, as in a regular Python program. A number of files are laid out for your use, under the `/opt/ml` directory:
#
# /opt/ml
# |-- input
# | |-- config
# | | |-- hyperparameters.json
# | | `-- resourceConfig.json
# | `-- data
# | `-- <channel_name>
# | `-- <input data>
# |-- model
# | `-- <model files>
# `-- output
# `-- failure
#
# ##### The input
#
# * `/opt/ml/input/config` contains information to control how your program runs. `hyperparameters.json` is a JSON-formatted dictionary of hyperparameter names to values. These values are always strings, so you may need to convert them. `resourceConfig.json` is a JSON-formatted file that describes the network layout used for distributed training.
# * `/opt/ml/input/data/<channel_name>/` (for File mode) contains the input data for that channel. The channels are created based on the call to CreateTrainingJob but it's generally important that channels match algorithm expectations. The files for each channel are copied from S3 to this directory, preserving the tree structure indicated by the S3 key structure.
# * `/opt/ml/input/data/<channel_name>_<epoch_number>` (for Pipe mode) is the pipe for a given epoch. Epochs start at zero and go up by one each time you read them. There is no limit to the number of epochs that you can run, but you must close each pipe before reading the next epoch.
#
# ##### The output
#
# * `/opt/ml/model/` is the directory where you write the model that your algorithm generates. Your model can be in any format that you want. It can be a single file or a whole directory tree. SageMaker packages any files in this directory into a compressed tar archive file. This file is made available at the S3 location returned in the `DescribeTrainingJob` result.
# * `/opt/ml/output` is a directory where the algorithm can write a file `failure` that describes why the job failed. The contents of this file are returned in the `FailureReason` field of the `DescribeTrainingJob` result. For jobs that succeed, there is no reason to write this file as it is ignored.
#
# #### Running your container during hosting
#
# Hosting has a very different model than training because hosting is reponding to inference requests that come in via HTTP. In this example, we use [TensorFlow Serving](https://www.tensorflow.org/serving/), however the hosting solution can be customized. One example is the [Python serving stack within the scikit learn example](https://github.com/awslabs/amazon-sagemaker-examples/blob/master/advanced_functionality/scikit_bring_your_own/scikit_bring_your_own.ipynb).
#
# Amazon SageMaker uses two URLs in the container:
#
# * `/ping` receives `GET` requests from the infrastructure. Your program returns 200 if the container is up and accepting requests.
# * `/invocations` is the endpoint that receives client inference `POST` requests. The format of the request and the response is up to the algorithm. If the client supplied `ContentType` and `Accept` headers, these are passed in as well.
#
# The container has the model files in the same place that they were written to during training:
#
# /opt/ml
# `-- model
# `-- <model files>
#
#
# ### The parts of the sample container
#
# This directory has all the components you need to package the sample algorithm for Amazon SageMager:
#
# .
# |-- Dockerfile
# `-- cifar10
# |-- cifar10.py
# |-- resnet_model.py
# |-- nginx.conf
# |-- serve
# `-- train
#
# Let's discuss each of these in turn:
#
# * __`Dockerfile`__ describes how to build your Docker container image. More details are provided below.
# * __`cifar10`__ is the directory which contains the files that are installed in the container.
#
# In this simple application, we install only five files in the container. You may only need that many, but if you have many supporting routines, you may wish to install more. These five files show the standard structure of our Python containers, although you are free to choose a different toolset and therefore could have a different layout. If you're writing in a different programming language, you will have a different layout depending on the frameworks and tools you choose.
#
# The files that we put in the container are:
#
# * __`cifar10.py`__ is the program that implements our training algorithm.
# * __`resnet_model.py`__ is the program that contains our Resnet model.
# * __`nginx.conf`__ is the configuration file for the nginx front-end. Generally, you should be able to take this file as-is.
# * __`serve`__ is the program started when the container is started for hosting. It simply launches nginx and loads your exported model with TensorFlow Serving.
# * __`train`__ is the program that is invoked when the container is run for training. Our implementation of this script invokes cifar10.py with our our hyperparameter values retrieved from /opt/ml/input/config/hyperparameters.json. The goal for doing this is to avoid having to modify our training algorithm program.
#
# In summary, the two files you probably want to change for your application are `train` and `serve`.
# ### The Dockerfile
#
# The Dockerfile describes the image that we want to build. You can think of it as describing the complete operating system installation of the system that you want to run. A Docker container running is quite a bit lighter than a full operating system, however, because it takes advantage of Linux on the host machine for the basic operations.
#
# For the Python science stack, we start from an official TensorFlow docker image and run the normal tools to install TensorFlow Serving. Then we add the code that implements our specific algorithm to the container and set up the right environment for it to run under.
#
# Let's look at the [Dockerfile](./container/Dockerfile) for this example.
# ## Building and registering the container using the SageMaker Studio Image Build CLI
#
# There are two ways to build and push docker images to ECR from within an Amazon SageMaker Studio Notebook.
#
# 1. **Setup Your Own Integrations** Build the necessary integrations and workflow into your Studio environment that allow you to use a build service such as AWS Code Build to build your docker images as well as setup your ECR repository and pushes image to that respository.
#
# 2. **Utilize the SageMaker Studio Image Build CLI convenience package** This is the preferred approach as it removes the heavy lift of setting up your own workflows and docker build capabilities. The CLI provides an abstraction of those underlying integrations and workflows allowing you to easily build and push docker images using simple CLI commands.
# ## Using the SageMaker Studio Image Build CLI Convenience Package
#
# There are just a few steps to get started using the new convenience package.
#
# ### Step 1: Install the CLI
import sys
# !{sys.executable} -m pip install sagemaker_studio_image_build
# ### Step 2: Ensure IAM Role has access to necessary services
#
# The SageMaker Studio Image Build CLI uses Amazon Elastic Container Registry and AWS CodeBuild so we need to ensure that the role we provide as input to our CLI commands has the necessary policies and permissions attached.
#
# Two scenarios are supported including:
#
# 1. **Add IAM Permissions to SageMaker Execution Role**
#
# This scenario includes updating the Execution Role attached to this notebook instance with the required permissions. In this scenario, you need to get the current execution role and ensure the trust policy and additional permissions are associated with the role.
#
# 2. **Create/Utilize a secondary role with appropriate permissions attached**
#
# This scenario includes using a secondary role setup with the permissions below and identified in the --role argument when invoking the CLI (Example: *sm-docker build . --role build-cli-role*)
#
# For this example, we are going to **Add IAM Permissions to the current SageMaker Execution Role**.
#
# Let's first grab the current execution role...
# +
import sagemaker
import boto3
try:
role = sagemaker.get_execution_role()
except:
role = get_execution_role()
print("Using IAM role arn: {}".format(role))
# -
# Now we need to add the permissions below for the role identified above.
#
# **Update Trust Policy for CodeBuild**
# * Open [IAM](https://console.aws.amazon.com/iam/home#/roles) and search for the role listed above.
# * Select the Role and click on the **Trust relationships** tab.
# * Update the trust relationship using the JSON to establish a trust relationship with CodeBuild
# + active=""
# {
# "Version": "2012-10-17",
# "Statement": [
# {
# "Effect": "Allow",
# "Principal": {
# "Service": "sagemaker.amazonaws.com"
# },
# "Action": "sts:AssumeRole"
# },
# {
# "Effect": "Allow",
# "Principal": {
# "Service": [
# "codebuild.amazonaws.com"
# ]
# },
# "Action": "sts:AssumeRole"
# }
# ]
# }
# -
# * Once you've added the trust relationship above, click **Update Trust Policy**
#
# We also need to add some additional permissions to the execution role to be able to build the image with CodeBuild and push the image to ECR. You can update the existing execution policy attached to the role or create a new policy and attach it to the existing execution role. Whichever option you choose, ensure the policy has the correct permissions set for intended S3 bucket access. The sample policy in the [CLI README](https://github.com/aws-samples/sagemaker-studio-image-build-cli/tree/b0b8d337dba4f1ecc88f33f81e815fb44c4c9915) assumes access to the default session bucket so this may need to be modified for your use casee. For this example, we are going to create a new policy and attach it to the existing role.
#
# **Create policy allowing access to supporting services**
#
# * Open [Policies](https://console.aws.amazon.com/iam/home#/policies) in IAM
# * Click **Create policy**
# * Select the JSON tab and copy/paste the policy below
# + active=""
# {
# "Version": "2012-10-17",
# "Statement": [
# {
# "Effect": "Allow",
# "Action": [
# "codebuild:DeleteProject",
# "codebuild:CreateProject",
# "codebuild:BatchGetBuilds",
# "codebuild:StartBuild"
# ],
# "Resource": "arn:aws:codebuild:*:*:project/sagemaker-studio*"
# },
# {
# "Effect": "Allow",
# "Action": "logs:CreateLogStream",
# "Resource": "arn:aws:logs:*:*:log-group:/aws/codebuild/sagemaker-studio*"
# },
# {
# "Effect": "Allow",
# "Action": [
# "logs:GetLogEvents",
# "logs:PutLogEvents"
# ],
# "Resource": "arn:aws:logs:*:*:log-group:/aws/codebuild/sagemaker-studio*:log-stream:*"
# },
# {
# "Effect": "Allow",
# "Action": "logs:CreateLogGroup",
# "Resource": "*"
# },
# {
# "Effect": "Allow",
# "Action": [
# "ecr:CreateRepository",
# "ecr:BatchGetImage",
# "ecr:CompleteLayerUpload",
# "ecr:DescribeImages",
# "ecr:DescribeRepositories",
# "ecr:UploadLayerPart",
# "ecr:ListImages",
# "ecr:InitiateLayerUpload",
# "ecr:BatchCheckLayerAvailability",
# "ecr:PutImage"
# ],
# "Resource": "arn:aws:ecr:*:*:repository/sagemaker-studio*"
# },
# {
# "Effect": "Allow",
# "Action": "ecr:GetAuthorizationToken",
# "Resource": "*"
# },
# {
# "Effect": "Allow",
# "Action": [
# "s3:GetObject",
# "s3:DeleteObject",
# "s3:PutObject"
# ],
# "Resource": "arn:aws:s3:::sagemaker-*/*"
# },
# {
# "Effect": "Allow",
# "Action": [
# "s3:CreateBucket"
# ],
# "Resource": "arn:aws:s3:::sagemaker*"
# },
# {
# "Effect": "Allow",
# "Action": [
# "iam:GetRole",
# "iam:ListRoles"
# ],
# "Resource": "*"
# },
# {
# "Effect": "Allow",
# "Action": "iam:PassRole",
# "Resource": "arn:aws:iam::*:role/*",
# "Condition": {
# "StringLikeIfExists": {
# "iam:PassedToService": "codebuild.amazonaws.com"
# }
# }
# }
# ]
# }
# -
# * Click **Review policy**
# * Give the policy a name such as `Studio-Image-Build-Policy`
# * Click **Create policy**
#
# We now need to attach our policy to the Execution Role attached to this notebook environment.
#
# * Go back to [Roles](https://console.aws.amazon.com/iam/home#/roles) in IAM
# * Select the SageMaker Execution Role from abovee
# * On the **Permissions** tab, click **Attach policies**
# * Search for the Policy we created above `Studio-Image-Build-Policy`
# * Select the policy and click **Attach policy**
# ### Step 3: Building and registering the container using the SageMaker Studio Image Build CLI
#
# We will now create our training container image, using the SageMaker Studio Image Build CLI
#
# To do this we need to navigate to the directory containing our Dockerfile and simply execute the build command:
#
# sm-docker build .
#
# The build command can optionally take additional arguments depending on your needs:
#
# sm-docker build . --file /path/to/Dockerfile --build-arg foo=bar
#
# **TIP** If you receive a permissions error below, please ensure you have completed **both** permission setup items above: (1) Update Trust Policy (2) Create new policy & Attach it to the existing SageMaker Execution Role
# !sm-docker build .
# **NOTE** The Image URI output above will be used as the input training image for our training job
#
# ---
# ## Download the CIFAR-10 dataset
# Our training algorithm is expecting our training data to be in the file format of [TFRecords](https://www.tensorflow.org/guide/datasets), which is a simple record-oriented binary format that many TensorFlow applications use for training data.
#
# Below is a Python script adapted from the [official TensorFlow CIFAR-10 example](https://github.com/tensorflow/models/blob/451906e4e82f19712455066c1b27e2a6ba71b1dd/research/slim/datasets/download_and_convert_cifar10.py), which downloads the CIFAR-10 dataset and converts them into TFRecords.
#
# The adapted script has a dependency on ipywidgets so we will first need to install that dependencies in our notebook prior to executing the script.
import sys
# !{sys.executable} -m pip install ipywidgets
# ! python utils/generate_cifar10_tfrecords.py --data-dir=/tmp/cifar-10-data
# There should be three tfrecords. (eval, train, validation)
# ! ls /tmp/cifar-10-data
# # Part 2: Training and Hosting your Algorithm in Amazon SageMaker
# Once you have your container packaged, you can use it to train and serve models. Let's do that with the algorithm we made above.
#
# ## Set up the environment
# Here we specify the bucket to use
# S3 prefix
prefix = 'DEMO-tensorflow-cifar10'
# ## Create the session
#
# The session remembers our connection parameters to SageMaker. We use it to perform all of our SageMaker operations.
# +
import sagemaker as sage
sess = sage.Session()
# -
# ## Upload the data for training
#
# We will use the tools provided by the SageMaker Python SDK to upload the data to a default bucket.
# +
WORK_DIRECTORY = '/tmp/cifar-10-data'
data_location = sess.upload_data(WORK_DIRECTORY, key_prefix=prefix)
# -
# ## Train & Deploy on SageMaker
# Next, we will perform our training using SageMaker Training Instances. Because are bringing our own training image, we need to specify our ECR image URL. This is the Image URI that was output from our SageMaker Studio Image Build CLI that we executed above. Make sure you update the ECR Image value with that output value as indicated below.
#
# Finally, our local training dataset has to be in Amazon S3 and the S3 URL to our dataset is passed into the `fit()` call. After our model is trained, we will then use the `deploy()` call to deploy our model to a persistent endpoint using SageMaker Hosting.
#
# Let's first fetch our ECR image url that corresponds to the image we just built and pushed.
# +
import boto3
sm = boto3.client('sagemaker')
ecr = boto3.client('ecr')
account = sess.boto_session.client('sts').get_caller_identity()['Account']
region = boto3.session.Session().region_name
domain_id = 'sagemaker-studio-{}'.format(sm.list_apps()['Apps'][0]['DomainId'])
image_tag = ecr.list_images(repositoryName=domain_id, filter={'tagStatus':'TAGGED'})['imageIds'][0]['imageTag']
ecr_image = '{}.dkr.ecr.{}.amazonaws.com/{}:{}'.format(account, region, domain_id, image_tag)
print(ecr_image)
# -
# ### Train our model using SageMaker Training Instances
# +
from sagemaker.estimator import Estimator
hyperparameters = {'train-steps': 100}
instance_type = 'ml.m4.xlarge'
estimator = Estimator(role=role,
train_instance_count=1,
train_instance_type=instance_type,
image_name=ecr_image,
hyperparameters=hyperparameters)
estimator.fit(data_location)
# -
# ### Host our model using SageMaker Hosting
predictor = estimator.deploy(1, instance_type)
# ### Test Endpoint - Making predictions using Python SDK
#
# To make predictions, we use an image that is converted using Imageio into a json format to send as an inference request. We need to install Imageio to deserialize the image that is used to make predictions.
#
# The JSON reponse will be the probabilities of the image belonging to one of the 10 classes along with the most likely class the picture belongs to. The classes can be referenced from the [CIFAR-10 website](https://www.cs.toronto.edu/~kriz/cifar.html).
#
# **NOTE**: Since we didn't train the model for that long, we aren't expecting very accurate results. To improve results, consider experimennting with additional training optimizations.
#
# **Import Imageio**
import sys
# !{sys.executable} -m pip install imageio
# **View our sample image**
# +
import os
from IPython.display import Image, display
images = []
for entry in os.scandir('data'):
if entry.is_file() and entry.name.endswith("png"):
images.append('data/' + entry.name)
for image in images:
display(Image(image))
# -
# **Format Image for prediction**
# +
import imageio as imageio
import numpy
from sagemaker.predictor import json_serializer, json_deserializer
image = imageio.imread("data/cat.png")
print(image.shape)
data = {'instances': numpy.asarray(image).astype(float).tolist()}
# -
# **Send image to endpoint for prediction**
# + jupyter={"source_hidden": true}
# The request and response format is JSON for TensorFlow Serving.
# For more information: https://www.tensorflow.org/serving/api_rest#predict_api
predictor.accept = 'application/json'
predictor.content_type = 'application/json'
predictor.serializer = json_serializer
predictor.deserializer = json_deserializer
# For more information on the predictor class.
# https://github.com/aws/sagemaker-python-sdk/blob/master/src/sagemaker/predictor.py
predictor.predict(data)
# -
# As we mentioned above, we don't expect our model to perform well as we did not train it for very long. You can increase you experiements through additional training cycles to continue to improve your model.
# ## Optional cleanup
# When you're done with the endpoint, you should clean it up.
#
# All of the training jobs, models and endpoints we created can be viewed through the SageMaker console of your AWS account.
predictor.delete_endpoint()
# # Reference
# - [SageMaker Studio Image Build CLI](https://github.com/aws-samples/sagemaker-studio-image-build-cli/README.md)
# - [How Amazon SageMaker interacts with your Docker container for training](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-training-algo.html)
# - [How Amazon SageMaker interacts with your Docker container for inference](https://docs.aws.amazon.com/sagemaker/latest/dg/your-algorithms-inference-code.html)
# - [CIFAR-10 Dataset](https://www.cs.toronto.edu/~kriz/cifar.html)
# - [SageMaker Python SDK](https://github.com/aws/sagemaker-python-sdk)
# - [Dockerfile](https://docs.docker.com/engine/reference/builder/)
|
aws_sagemaker_studio/sagemaker_studio_image_build/tensorflow_bring_your_own/tensorflow_bring_your_own.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Monitoring change through time using satellite imagery filmstrip plots <img align="right" src="../Supplementary_data/DE_Africa_Logo_Stacked_RGB_small.jpg">
#
# * **Products used:**
# [ls8_usgs_sr_scene](https://explorer.digitalearth.africa/ls8_usgs_sr_scene),
# [ls7_usgs_sr_scene](https://explorer.digitalearth.africa/ls7_usgs_sr_scene),
# [ls5_usgs_sr_scene](https://explorer.digitalearth.africa/ls5_usgs_sr_scene)
#
# ## Background
#
# Understanding how natural and human landscapes have changed through time can provide vital information about the health of local ecosystems and development of the built environment.
# For example, data on changes in the distribution of vegetation in the landscape can be used to monitor the impact of deforestation, or track the recovery of forests after habitat restoration or extreme natural events (e.g. bushfires).
# Tracking changes within urban areas can be used to monitor the growth of infrastructure such as ports and transport networks, while data on coastal changes can be vital for predicting and managing the impacts of coastal erosion or the loss of coastal wetlands (e.g. mangroves).
#
# Although these examples of change can be tracked using direct on-the-ground monitoring (e.g. vegetation surveys), it can be extremely challenging and expensive to obtain a comprehensive understanding of these processes at a broader landscape scale.
# For many applications, it can also be extremely useful to obtain a record of the history of a location undergoing change.
# This typically requires historical monitoring data which is unlikely to be available for all but the most intensively monitored locations.
#
# ### Digital Earth Africa use case
#
# More than 30 years of satellite imagery from the [NASA/USGS Landsat program](https://www.usgs.gov/land-resources/nli/landsat) is freely available for Africa, making this a powerful resource for monitoring natural and human-driven change across the African continent.
# Because these satellites image every location over Africa regularly (approximately once every 16 days), they provide an unparalleled archive of how many of Africa's landscapes have changed through time.
#
# Analysing change from individual satellite images can be made difficult by the presence of clouds, cloud shadow, sunglint over water, and dynamic processes like changing tides along the coastline.
# By combining individual noisy images into cleaner, cloud-free "summary" images that cover a longer time period (e.g. one or multiple years), we can obtain a clear, consistent view of the African environment that can be compared to reveal changes in the landscape over time.
#
# ## Description
# In this example, Digital Earth Africa Landsat data is extracted for a given time range and location, and combined using the geometric median ("geomedian") statistic to reveal the median or 'typical' appearance of the landscape for a series of time periods (for more information about geomedians, see the [Geomedian composites notebook](../Frequently_used_code/Geomedian_composites.ipynb)).
#
# For coastal applications, the analysis can be customised to select only satellite images obtained during a specific tidal range (e.g. low, average or high tide).
#
# The results for each time period are combined into a 'filmstrip' plot which visualises how the landscape has changed in appearance across time, with a 'change heatmap' panel highlighting potential areas of greatest change:
#
# 
#
# ***
# ## Getting started
#
# To run this analysis, run all the cells in the notebook, starting with the "Load packages" cell.
# ### Load packages
# Import Python packages used for the analysis.
# +
# %matplotlib inline
import sys
import datacube
from datacube.helpers import write_geotiff
sys.path.append('../Scripts')
from notebookapp_changefilmstrips import run_filmstrip_app
# -
# ### Analysis parameters
#
# The following cell sets important required parameters for the analysis:
#
# * `output_name`: A name that will be used to name the output filmstrip plot file
# * `time_range`: The date range to analyse (e.g. `time_range = ('1984-01-01', '2019-12-31)`)
# * `time_step`: This parameter allows us to choose the length of the time periods we want to compare (e.g. `time_step = {'years': 5}` will generate one filmstrip plot for every five years of data in the dataset; `time_step = {'months': 18}` will generate one plot for each 18 month period etc.
# Time periods are counted from the first value given in `time_range`.
#
# Optional parameters:
#
# * `tide_range`: This parameter allows you to generate filmstrip plots based on specific ocean tide conditions.
# This can be valuable for analysing change consistently along the coast.
# For example, `tide_range = (0.0, 0.2)` will select only satellite images acquired at the lowest 20% of tides; `tide_range = (0.8, 1.0)` will select images from the highest 20% of tides.
# The default is `tide_range = (0.0, 1.0)` which will select all images regardless of tide.
# * `resolution`: The spatial resolution to load data.
# The default is `resolution = (-30, 30)`, which will load data at 30 m pixel resolution.
# Increasing this (e.g. to `resolution = (-100, 100)`) can be useful for loading large spatial extents.
# * `max_cloud`: This parameter allows you to exclude satellite images with excessive cloud.
# The default is `0.5`, which will keep all images with less than 50% cloud.
# * `ls7_slc_off`: Whether to include data from after the Landsat 7 SLC failure (i.e. SLC-off).
# Defaults to `False`, which removes all Landsat 7 observations after May 31 2003.
# Setting this to `True` will result in extra data, but can also introduce horizontal striping in the output filmstrip plots.
# * `size_limit`: An optional integer (in square kilometres) specifying the size limit for the data query. Queries larger than this size will receive a warning that he data query is too large (and may therefore result in memory errors).
#
#
# **If running the notebook for the first time**, keep the default settings below. This will demonstrate how the analysis works and provide meaningful results.
#
# +
# Required parameters
output_name = 'example'
time_range = ('1984-01-01', '2019-12-31')
time_step = {'years': 10}
# Optional parameters
tide_range = (0.0, 1.0)
resolution = (-30, 30)
max_cloud = 0.5
ls7_slc_off = False
size_limit = 100
# -
# ## Select location and generate filmstrips
# Run the following cell to start the analysis.
# This will plot an interactive map that is used to select the area to load satellite data for.
#
# Select the `Draw a rectangle` or `Draw a polygon` tool on the left of the map, and draw a shape around the area you are interested in.
#
# If running this notebook for the first time, **draw a rectangle around West African Cement Company plant** to see an example of change driven by urban development.
# You should draw a rectangle similar to the image shown below:
#
# 
#
# If running this notebook for a new area, zoom and pan around the map until you find an area of interest, then draw a rectangle or polygon as described above.
# You will need to check the availability of Landsat data in your chosen area using the [Digital Earth Africa Explorer](https://explorer.digitalearth.africa/ls8_usgs_sr_scene).
#
# When you are ready, press the green `done` button on the top right of the map.
# This will start loading the data, and then generate a filmstrips plot.
#
# > Depending on the size of the area you select, this step can take **several minutes to complete**.
# To keep load times reasonable, select an area **smaller than 10,000 hectares** in size.
# When using the rectangle drawing tool, the area in hectares will be displayed to help guide you.
# The 10,000 hectare limit can be overuled by supplying the `size_limit` parameter in the `Analysis Parameters` section above.
#
#
# > Once the analysis reaches the `Generating geomedian composites` step, you can check the status of the data load by clicking the **Dashboard** link under **Client** below.
#
output_data = run_filmstrip_app(output_name,
time_range,
time_step,
tide_range,
resolution,
max_cloud,
ls7_slc_off)
# ## Using filmstrip plots to identify change
#
# The filmstrip plot above contains several colour imagery panels that summarise the median or 'typical' appearance of the landscape for the time periods defined using `time_range` and `time_step`.
# If you ran the analysis over the West African Cemement Company plant, inspect each of the imagery plots.
# You should be able to see the expansion of the plant over time.
#
# ### Change heatmap
#
# To make it easier to identify areas that have changed between each filmstrip panel, the final panel provides a "change heatmap".
# This highlights pixels whose values vary greatly between the panels in the filmstrip plot.
# Bright colours indicate pixels that have changed; dark colours indicate pixels that have remained relatively similar across time.
#
# Compare the "change heatmap" panel against the colour imagery panels.
# You should be able to see the expansion clearly in bright pixels and the central plant buildings in the darkest pixels.
#
# > **Technical info:** The "change heatmap" is calculated by first taking a log transform of the imagery data to emphasize dark pixels, then calculating standard deviation across all of the filmstrip panels to reveal pixels that changed over time.
# ## Downloading filmstrip plot
# The high resolution version of the filmstrip plot generated above will be saved to the same location you are running this notebook from (e.g. typically `Real_world_examples`).
# In JupyterLab, use the file browser to locate the image file with a name in the following format:
#
# `filmstrip_{output_name}_{date_string}_{time_step}.png`
#
# You can download the image to your PC by right clicking on the image file and selecting `Download`.
# ## Export GeoTIFF data
# It can be useful to export each of the filmstrip panels generated above as GeoTIFF raster files so that they can be loaded into a Geographic Information System (GIS) software for further analysis.
# Because the filmstrip panels were generated using the "geomedian" statistic that preserves relationships between spectral bands, the resulting data can be validly analysed in the same way as we would analyse an individual satellite image.
#
# To export the GeoTIFFs, run the following cell then right click on the files in the JupyterLab file browser and select `Download`.
for i, ds in output_data.groupby('timestep'):
print(f'Exporting {i} data')
write_geotiff(dataset=ds, filename=f'geotiff_{output_name}_{i}.tif')
# ## Next steps
# When you are done, return to the [Analysis parameters](#Analysis-parameters) section, modify some values and rerun the analysis.
# For example, you could try:
#
# * Modify `time_range` to look at a specific time period of interest.
# * Setting a shorter `time_step` (e.g. `time_step = {'years': 5}`) for a more detailed look at how the landscape has changed over shorter time periods.
# * Move to a different area of interest.
#
# ***
#
# ## Additional information
#
# **License:** The code in this notebook is licensed under the [Apache License, Version 2.0](https://www.apache.org/licenses/LICENSE-2.0).
# Digital Earth Africa data is licensed under the [Creative Commons by Attribution 4.0](https://creativecommons.org/licenses/by/4.0/) license.
#
# **Contact:** If you need assistance, please post a question on the [Open Data Cube Slack channel](http://slack.opendatacube.org/) or on the [GIS Stack Exchange](https://gis.stackexchange.com/questions/ask?tags=open-data-cube) using the `open-data-cube` tag (you can view previously asked questions [here](https://gis.stackexchange.com/questions/tagged/open-data-cube)).
# If you would like to report an issue with this notebook, you can file one on [Github](https://github.com/digitalearthafrica/deafrica-sandbox-notebooks).
#
# **Last modified:** April 2020
#
# **Compatible datacube version:**
print(datacube.__version__)
# ## Tags
# Browse all available tags on the DE Africa User Guide's [Tags Index](https://) (placeholder as this does not exist yet)
# + raw_mimetype="text/restructuredtext" active=""
# **Tags**: :index:`landsat 5`, :index:`landsat 7`, :index:`landsat 8`, :index:`deafrica_datahandling`, :index:`deafrica_coastaltools`, :index:`deafrica_plotting`, :index:`load_ard`, :index:`mostcommon_crs`, :index:`tidal_tag`, :index:`rgb`, :index:`image compositing`, :index:`geomedian`, :index:`filmstrip plot`, :index:`change monitoring`, :index:`real world`, :index:`widgets`, :index:`interactive`
|
Real_world_examples/Change_filmstrips.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Examples with Different Protocols Showing Metrics
#
# ## Prerequisites
#
# * A kubernetes cluster with kubectl configured
# * curl
# * grpcurl
# * pygmentize
#
#
# ## Setup Seldon Core
#
# Install Seldon Core as described in [docs](https://docs.seldon.io/projects/seldon-core/en/latest/workflow/install.html)
#
# Then port-forward to that ingress on localhost:8003 in a separate terminal either with:
#
# * Ambassador:
#
# ```bash
# kubectl port-forward $(kubectl get pods -n seldon -l app.kubernetes.io/name=ambassador -o jsonpath='{.items[0].metadata.name}') -n seldon 8003:8080
# ```
# * Istio:
#
# ```bash
# kubectl port-forward $(kubectl get pods -l istio=ingressgateway -n istio-system -o jsonpath='{.items[0].metadata.name}') -n istio-system 8003:80
# ```
#
# !kubectl create namespace seldon
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
# ## Install Seldon Analytics
# !helm install seldon-core-analytics ../../../helm-charts/seldon-core-analytics \
# --set grafana_prom_admin_password=password \
# --set persistence.enabled=false \
# --namespace seldon-system \
# --wait
# Port forward to the Grafana dashboard
#
# ```bash
# kubectl port-forward $(kubectl get pods -n seldon-system -l app.kubernetes.io/name=grafana -o jsonpath='{.items[0].metadata.name}') 3000:3000 -n seldon-system
# ```
# %env RESOURCES=../../../notebooks/resources
# ## Seldon Protocol REST Model
#
# **Make sure your active namespace is seldon**
# !pygmentize ${RESOURCES}/model_seldon_rest.yaml
# !kubectl apply -f ${RESOURCES}/model_seldon_rest.yaml -n seldon
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=rest-seldon \
# -o jsonpath='{.items[0].metadata.name}')
# !for i in `seq 1 60`; do \
# sleep 1 && curl -d '{"data": {"ndarray":[[1.0, 2.0, 5.0]]}}' \
# -X POST http://localhost:8003/seldon/seldon/rest-seldon/api/v1.0/predictions \
# -H "Content-Type: application/json"; \
# done
# 
import time
for i in range(3):
# metric=!curl -s http://localhost:8003/seldon/seldon/rest-seldon/prometheus | grep seldon_api_executor_server_requests_seconds_count
if metric and len(metric)>0:
print(metric[0])
assert(not metric[0] == "")
break
else:
print("Failed to get metrics for rest-seldon")
time.sleep(2)
# !kubectl delete -f ${RESOURCES}/model_seldon_rest.yaml -n seldon
# ## Seldon Protocol GRPC Model
# !pygmentize ${RESOURCES}/model_seldon_grpc.yaml
# !kubectl apply -f ${RESOURCES}/model_seldon_grpc.yaml -n seldon
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=grpc-seldon \
# -o jsonpath='{.items[0].metadata.name}')
# !cd ../../../executor/proto && for i in `seq 1 60`; do \
# sleep 1 && grpcurl -d '{"data":{"ndarray":[[1.0,2.0]]}}' \
# -rpc-header seldon:grpc-seldon -rpc-header namespace:seldon \
# -plaintext \
# -proto ./prediction.proto 0.0.0.0:8003 seldon.protos.Seldon/Predict; \
# done
# 
for i in range(3):
# metric=!curl -s http://localhost:8003/seldon/seldon/grpc-seldon/prometheus | grep seldon_api_executor_server_requests_seconds_count
if metric and len(metric)>0:
print(metric[0])
assert(not metric[0] == "")
break
else:
print("Failed to get metrics for grpc-seldon")
time.sleep(2)
# !kubectl delete -f ${RESOURCES}/model_seldon_grpc.yaml -n seldon
# ## Tensorflow Protocol REST Model
# !pygmentize ${RESOURCES}/model_tfserving_rest.yaml
# !kubectl apply -f ${RESOURCES}/model_tfserving_rest.yaml -n seldon
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=rest-tfserving \
# -o jsonpath='{.items[0].metadata.name}')
# !for i in `seq 1 60`; do \
# sleep 1 && curl -d '{"instances": [1.0, 2.0, 5.0]}' \
# -X POST http://localhost:8003/seldon/seldon/rest-tfserving/v1/models/halfplustwo/:predict \
# -H "Content-Type: application/json"; \
# done
for i in range(3):
# metric=!curl -s http://localhost:8003/seldon/seldon/rest-tfserving/prometheus | grep seldon_api_executor_server_requests_seconds_count
if metric and len(metric)>0:
print(metric[0])
assert(not metric[0] == "")
break
else:
print("Failed to get metrics for rest-tfserving")
time.sleep(2)
# 
# !kubectl delete -f ${RESOURCES}/model_tfserving_rest.yaml -n seldon
# ## Tensorflow Protocol GRPC Model
# !pygmentize ${RESOURCES}/model_tfserving_grpc.yaml
# !kubectl apply -f ${RESOURCES}/model_tfserving_grpc.yaml -n seldon
# !kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id=grpc-tfserving \
# -o jsonpath='{.items[0].metadata.name}')
# !cd ../../../executor/proto && for i in `seq 1 60`; do \
# sleep 1 && grpcurl \
# -d '{"model_spec":{"name":"halfplustwo"},"inputs":{"x":{"dtype": 1, "tensor_shape": {"dim":[{"size": 3}]}, "floatVal" : [1.0, 2.0, 3.0]}}}' \
# -rpc-header seldon:grpc-tfserving -rpc-header namespace:seldon \
# -plaintext -proto ./prediction_service.proto \
# 0.0.0.0:8003 tensorflow.serving.PredictionService/Predict; \
# done
# 
for i in range(3):
# metric=!curl -s http://localhost:8003/seldon/seldon/grpc-tfserving/prometheus | grep seldon_api_executor_server_requests_seconds_count
if metric and len(metric)>0:
print(metric[0])
assert(not metric[0] == "")
break
else:
print("Failed to get metrics for grpc-tfserving")
time.sleep(2)
# !kubectl delete -f ${RESOURCES}/model_tfserving_grpc.yaml -n seldon
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon-system
# !helm delete seldon-core-analytics -n seldon-system
# !kubectl config set-context $(kubectl config current-context) --namespace=seldon
|
examples/models/metrics/metrics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using ssh-copy-id to add your key
# ## Overview:
# - **Teaching:** 15 min
# - **Exercises:** 0 min
#
# **Questions**
# - How do I add my own keys to unmanaged services?
# - Where does ssh store my public key
#
# **Objectives**
# - Know how to use `ssh-copy-id` to add your key to a remote service
# - Know that keys are added to `~/.ssh/authorized_keys` on the remote system
# ## Copy the public part of the key to the remote host
#
# When you create the key pair two files will be generated, a private key e.g. `id_ed25519` (or `id_rsa`) and the public key `id_ed25519.pub` (or `id_rsa.pub`). Your private key should never be copied to different machines, however, in order to use your key pair you do need to copy the public key to the remote machine.
#
# Using you normal login password, add the public part of your key pair to the authorized\_keys file on the remote host to which you wish to connect. We can use the utility `ssh-copy-id` to do this:
#
# ```bash
# ssh-copy-id -i ~/.ssh/id_ed25519.pub [userID]@<hpc-service>
# ```
#
# Now you can test that your key pair is working correctly by attempting to connect to the remote host and run a command. You should be asked for your key pair **passphase** (which you entered when you created the key pair) rather than your remote machine **password**.
#
# ```
# ssh [userID]@<hpc-service> 'date'
# Enter passphrase for key '/Home/user/.ssh/id_rsa': [Passphrase]
# Wed May 8 10:36:48 BST 2020
# ```
#
# We have run `date` on the remote server to confirm that we have been able to use the key pair, and **passphrase** to log in.
# ### Information: What is `ssh-copy-id` doing?
#
# `ssh-copy-id` is appending the contents of the public part of the key to the remote file `~/.ssh/authorized_keys`.
#
# You could also copy and paste your public key into the remote `~/.ssh/authorized_keys` but using the provided tool makes this easier.
#
# *If you do this make sure that you don't replace existing keys that you want to keep.*
# ## Exercise: Add your **public** key to the remote service
#
# ### Linux:
#
# Use `ssh-copy-id` to the remote service and verify that it works.
#
# If you have used a non-standard name or location you will have to explicitly use the key with:
#
# `ssh -i /path/to/id_key_name [userID]@<hpc-service>`
#
# ### Windows
#
# Add your key to the new connection you created under -> connection ->ssh -> auth `Private key file for authentication`
# ## Key Points:
#
# - use `ssh-copy-id` to add your key to a remote service
# - keys are added to `~/.ssh/authorized_keys` on the remote system
|
nbplain/06_sshcopy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img align="right" src="images/tf-small.png" width="128"/>
# <img align="right" src="images/etcbc.png"/>
# <img align="right" src="images/dans-small.png"/>
#
# You might want to consider the [start](search.ipynb) of this tutorial.
#
# Short introductions to other TF datasets:
#
# * [Dead Sea Scrolls](https://nbviewer.jupyter.org/github/annotation/tutorials/blob/master/lorentz2020/dss.ipynb),
# * [Old Babylonian Letters](https://nbviewer.jupyter.org/github/annotation/tutorials/blob/master/lorentz2020/oldbabylonian.ipynb),
# or the
# * [Q'uran](https://nbviewer.jupyter.org/github/annotation/tutorials/blob/master/lorentz2020/quran.ipynb)
#
# %load_ext autoreload
# %autoreload 2
from tf.app import use
A = use('bhsa', hoist=globals())
# A = use("bhsa:clone", checkout="clone", hoist=globals())
# # Quantifiers
# Quantifiers add considerable power to search templates.
#
# Quantifiers consist of full-fledged search templates themselves, and give rise to
# auxiliary searches being performed.
#
# The use of quantifiers may prevent the need to resort to hand-coding in many cases.
# That said, they can also be exceedingly tricky, so that it is advisable to check the results
# by hand-coding anyway, until you are perfectly comfortable with them.
# # Examples
# ## Lexemes
#
# It is easy to find the lexemes that occur in a specific book only.
# Because the `lex` node of such a lexeme is contained in the node of that specific book.
#
# Lets get the lexemes specific to Ezra and then those specific to Nehemiah.
# +
query = """
book book@en=Ezra
lex
"""
ezLexemes = A.search(query)
ezSet = {r[1] for r in ezLexemes}
query = """
book book@en=Nehemiah
lex
"""
nhLexemes = A.search(query)
nhSet = {r[1] for r in nhLexemes}
print(f"Total {len(ezSet | nhSet)} lexemes")
# -
# What if we want to have the lexemes that occur only in Ezra and Nehemia?
#
# If such a lexeme occurs in both books, it will not be contained by either book.
# So we have missed them by the two queries above.
#
# We have to find a different way. Something like: search for lexemes of which all words occur either in Ezra or in Nehemia.
#
# With the template constructions you have seen so far, this is impossible to say.
#
# This is where [*quantifiers*](https://annotation.github.io/text-fabric/tf/about/searchusage.html#quantifiers) come in.
# ## /without/
#
# First we are going to query for these lexemes by means of a `no:` quantifier.
query = """
lex
/without/
book book@en#Ezra|Nehemiah
w:word
w ]] ..
/-/
"""
query1results = A.search(query, shallow=True)
# ## /where/
#
# Now the `/without/` quantifier is a bit of a roundabout way to say what you really mean.
# We can also employ the more positive `/where/` quantifier.
query = """
lex
/where/
w:word
/have/
b:book book@en=Ezra|Nehemiah
w ]] b
/-/
"""
query2results = A.search(query, shallow=True)
# Check by hand coding:
A.silentOff()
A.indent(reset=True)
universe = F.otype.s("lex")
wordsEzNh = set(
L.d(T.bookNode("Ezra", lang="en"), otype="word")
+ L.d(T.bookNode("Nehemiah", lang="en"), otype="word")
)
handResults = set()
for lex in universe:
occs = set(L.d(lex, otype="word"))
if occs <= wordsEzNh:
handResults.add(lex)
A.info(len(handResults))
# Looks good, but we are thorough:
print(query1results == handResults)
print(query2results == handResults)
# ## Verb phrases
#
# Let's look for clauses with where all `Pred` phrases contain only verbs and look for `Subj`
# phrases in those clauses.
query = """
clause
/where/
phrase function=Pred
/have/
/without/
word sp#verb
/-/
/-/
phrase function=Subj
"""
queryResults = A.search(query)
A.show(queryResults, end=5, condenseType="sentence")
# Note that the pieces of template that belong to a quantifier, do not correspond to nodes in the result tuples!
# Check by hand:
A.indent(reset=True)
handResults = []
for clause in F.otype.s("clause"):
phrases = L.d(clause, otype="phrase")
preds = [p for p in phrases if F.function.v(p) == "Pred"]
good = True
for pred in preds:
if any(F.sp.v(w) != "verb" for w in L.d(pred, otype="word")):
good = False
if good:
subjs = [p for p in phrases if F.function.v(p) == "Subj"]
for subj in subjs:
handResults.append((clause, subj))
A.info(len(handResults))
queryResults == handResults
# ### Inspection
#
# We can see which templates are being composed in the course of interpreting the quantifier.
# We use the good old `S.study()`:
query = """
clause
/where/
phrase function=Pred
/have/
/without/
word sp#verb
/-/
/-/
phrase function=Subj
"""
S.study(query)
# Observe the stepwise unraveling of the quantifiers, and the auxiliary templates that are distilled
# from your original template.
#
# If you ever get syntax errors, run `S.study()` to find clues.
# ## Subject at start or at end
#
# We want the clauses that consist of at least two adjacent phrases, has a Subj phrase, which is either at the beginning or at the end.
# +
query = """
c:clause
/with/
=: phrase function=Subj
/or/
:= phrase function=Subj
/-/
phrase
<: phrase
"""
queryResults = sorted(A.search(query, shallow=True))
# -
# Check by hand:
A.indent(reset=True)
handResults = []
for clause in F.otype.s("clause"):
clauseWords = L.d(clause, otype="word")
phrases = set(L.d(clause, otype="phrase"))
if any(
L.n(p, otype="phrase") and (L.n(p, otype="phrase")[0] in phrases)
for p in phrases
):
# handResults.append(clause)
# continue
subjPhrases = [p for p in phrases if F.function.v(p) == "Subj"]
if any(L.d(p, otype="word")[0] == clauseWords[0] for p in subjPhrases) or any(
L.d(p, otype="word")[-1] == clauseWords[-1] for p in subjPhrases
):
handResults.append(clause)
A.info(len(handResults))
# A nice case where the search template performs better than this particular piece of hand-coding.
queryResults == handResults
# Let's also study this query:
S.study(query)
# ## Verb-containing phrases
#
# Suppose we want to collect all phrases with the condition that if they
# contain a verb, their `function` is `Pred`.
#
# This is a bit theoretical, but it shows two powerful constructs to increase readability
# of quantifiers.
# ### Unreadable
#
# First we express it without special constructs.
query = """
p:phrase
/where/
w:word pdp=verb
/have/
q:phrase function=Pred
q = p
/-/
"""
results = A.search(query, shallow=True)
# We check the query by means of hand-coding:
#
# 1. is every result a phrase: either without verbs, or with function Pred?
# 2. is every phrase without verbs or with function Pred contained in the results?
# +
allPhrases = set(F.otype.s("phrase"))
ok1 = all(
F.function.v(p) == "Pred" or all(F.pdp.v(w) != "verb" for w in L.d(p, otype="word"))
for p in results
)
ok2 = all(
p in results
for p in allPhrases
if (
F.function.v(p) == "Pred"
or all(F.pdp.v(w) != "verb" for w in L.d(p, otype="word"))
)
)
print(f"Check 1: {ok1}")
print(f"Check 2: {ok2}")
# -
# Ok, we are sure that the query does what we think it does.
# ### Readable
#
# Now let's make it more readable.
query = """
phrase
/where/
w:word pdp=verb
/have/
.. function=Pred
/-/
"""
# +
results2 = A.search(query, shallow=True)
print(f"Same results as before? {results == results2}")
# -
# Try to see how search is providing the name `parent` to the phrase atom and how it resolves the name `..`:
S.study(query)
# # All steps
#
# * **[start](start.ipynb)** your first step in mastering the bible computationally
# * **[display](display.ipynb)** become an expert in creating pretty displays of your text structures
# * **[search](search.ipynb)** turbo charge your hand-coding with search templates
#
# ---
#
# [advanced](searchAdvanced.ipynb)
# [sets](searchSets.ipynb)
# [relations](searchRelations.ipynb)
# quantifiers
#
# You have come far.
#
# Time to have a look at prior work.
#
# [fromMQL](searchFromMQL.ipynb)
# [rough](searchRough.ipynb)
# [gaps](searchGaps.ipynb)
#
# ---
#
# * **[exportExcel](exportExcel.ipynb)** make tailor-made spreadsheets out of your results
# * **[share](share.ipynb)** draw in other people's data and let them use yours
# * **[export](export.ipynb)** export your dataset as an Emdros database
# * **[annotate](annotate.ipynb)** annotate plain text by means of other tools and import the annotations as TF features
# * **[volumes](volumes.ipynb)** work with selected books only
# * **[trees](trees.ipynb)** work with the BHSA data as syntax trees
#
# CC-BY <NAME>
|
bhsa/searchQuantifiers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %config IPCompleter.greedy=True
2+2
print ("hola")
# # Carga de datos a traves de la funcion read_csv
import pandas as pd
import os
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets"
filename = "titanic/titanic3.csv"
fullpath = mainpath +"/"+ filename
df= pd.read_csv(fullpath)
df.head()
import os
# # otra forma de hacerlo con os (operating system) haciendo join
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets"
filename = "titanic/titanic3.csv "
fullpath = os.path.join(mainpath,filename)
df=pd.read_csv(fullpath)
df.head()
df2 = pd.read_csv(mainpath + "/" + "customer-churn-model/customer churn model.txt")
df2.columns.values
df2.head()
# # Para ver la cabecera de nuestro dataset
df2.columns.values
df = pd.read_csv("/Users/34686/Documents/GitHub/python-ml-course/datasets/titanic/titanic3.csv",
sep = ',', dtype=None, header=0, skip_blank_lines=True, skiprows=None)
df.head()
import pandas as pd
df3_cols = pd.read_csv ('/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Columns.csv')
df3_cols
df3_cols_list= df3_cols["Column_Names"].tolist()
df3_cols_list
df4_cols = pd.read_csv ("/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Model.txt")
df4_cols
df4_cols = pd.read_csv ("/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Model.txt")
df4_cols
df4_cols = pd.read_csv ("/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Model.txt")
df4_cols.columns.values
df3_cols = pd.read_csv ("/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Columns.csv")
df3_cols
df3_cols_list= df3_cols["Column_Names"].tolist()
df3_cols_list
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets/"
import pandas as pd
# # Cambiar nombres de las columnas
df3_cols = pd.read_csv(mainpath + "/" + "customer-churn-model/Customer Churn Columns.csv")
df3_cols_list= df3_cols["Column_Names"].tolist()
df4_cols = pd.read_csv ("/Users/34686/Documents/GitHub/python-ml-course/datasets/customer-churn-model/Customer Churn Model.txt",
header = None, names = df3_cols_list)
df4_cols.columns.values
df4_cols
# # Carga de datos a partir de la función open
# ## abrir la fila de las columnas para ver de cuantas columnas esta conformado
df5 = open (mainpath + "/" + "customer-churn-model/Customer Churn Model.txt", "r")
cols = df5.readline().strip().split(",")
cols
# # para ver el número de columnas
cols = df5.readline().strip().split(",")
n_cols = len(cols)
n_cols
counter = 0
main_dictionary = {}
for col in cols:
main_dict[col] = {}
main_dict
# # en el script anterior contábamos columnas y ahora filas
data3 = open(mainpath + "/" + "customer-churn-model/Customer Churn Model.txt",'r')
cols = data3.readline().strip().split(",")
n_cols = len(cols)
counter = 0
main_dict = {}
for col in cols:
main_dict[col] = []
# +
for line in data3:
values = line.strip().split(",")
for i in range(len(cols)):
main_dict[cols[i]].append(values[i])
counter += 1
print("El data set tiene %d filas y %d columnas"%(counter, n_cols))
# -
df3 = pd.DataFrame(main_dict)
df3.head()
# # Lectura y escritura de ficheros
infile = mainpath + "/" + "customer-churn-model/Customer Churn Model.txt"
outfile = mainpath + "/" + "customer-churn-model/tab2 Customer Churn Model.txt"
with open(infile, "r") as infile1:
with open(outfile, "w") as outfile1:
for line in infile1:
fields = line.strip().split(",")
outfile1.write("\t".join(fields)) # separamos los campos con un tab (\t)
outfile1.write("\n") # "("\n") es new line o INTRO (salto de línea)
df9 = pd.read_csv (outfile, sep = "\t")
df9
#
# # CARGAR DATOS DESDE UNA URL
medals_url = "http://winterolympicsmedals.com/medals.csv"
import pandas as pd
medals_data = pd.read_csv (medals_url) # Después leer el csv
medals_data.head()
medals_data.to_excel (mainpath + "/titanic/medals_data.xls") # envío el archivo a la ruta del pc en el formato xls
import urllib3
import http
medals_data.to_excel (mainpath + "/athletes/medals_juan.xls") # envío el archivo a la ruta del pc en el formato xls
medals_data.to_csv (mainpath + "/athletes/medals_juan.csv") # envío el archivo a la ruta del pc en el formato csv
medals_data.to_json (mainpath + "/athletes/medals_juan.json") # envío el archivo a la ruta del pc en el formato json
# # IMPORTAR LIBRERIAS CSV Y URLLIB3 DE PYTHON
import csv # TRAER INFORMACION DESDE FICHEROS CSV
import urllib3#TRAER O NAVEGAR POR INFORMACION DESDE UNA URL
import os
import http
import urllib3
# # Ejercicio de descarga MANUAL de datos video 30 con urllib3
def downloadFromURL(url, filename, sep = ",", delim = "\n", encoding="utf-8",
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets"):
#primero importamos la librería y hacemos la conexión con la web de los datos
import urllib3
http = urllib3.PoolManager()
r = http.request('GET', medals_url)# en este caso es medals_url porque lo definí en el anterior ejemplo
print("El estado de la respuesta es %d" %(r.status))
response = r.data ## CORREGIDO: eliminado un doble decode que daba error
#El objeto reponse contiene un string binario, así que lo convertimos a un string descodificándolo en UTF-8
str_data = response.decode(encoding)
#Dividimos el string en un array de filas, separándolo por intros
lines = str_data.split(delim)
#La primera línea contiene la cabecera, así que la extraemos
col_names = lines[0].split(sep)
n_cols = len(col_names)
#Generamos un diccionario vacío donde irá la información procesada desde la URL externa
counter = 0
main_dict = {}
for col in col_names:
main_dict[col] = []
#Procesamos fila a fila la información para ir rellenando el diccionario con los datos como hicimos antes
for line in lines:
#Nos saltamos la primera línea que es la que contiene la cabecera y ya tenemos procesada
if(counter > 0):
#Dividimos cada string por las comas como elemento separador
values = line.strip().split(sep)
#Añadimos cada valor a su respectiva columna del diccionario
for i in range(len(col_names)):
main_dict[col_names[i]].append(values[i])
counter += 1
print("El data set tiene %d filas y %d columnas"%(counter, n_cols))
#Convertimos el diccionario procesado a Data Frame y comprobamos que los datos son correctos
df10 = pd.DataFrame(main_dict)
print(df10.head())
#Elegimos donde guardarlo (en la carpeta athletes es donde tiene más sentido por el contexto del análisis)
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets"
filename = "/athletes/medals1_juan.xls"
fullpath = os.path.join(mainpath, filename)
#Lo guardamos en CSV, en JSON o en Excel según queramos
df10.to_csv (mainpath + "/athletes/medals2_juan.csv")
df10.to_json (mainpath + "/athletes/medals2_juan.json")
df10.to_excel (mainpath + "/athletes/medals2_juan.xls")
print("Los ficheros se han guardado correctamente en: "+ fullpath)
return df10
# # Tambien podemos realizar esta forma
medals_df = downloadFromURL(medals_url, "athletes/medals2_juan.xls")
medals_df.head()
# # Importar xls y xlsx
#
import pandas as pd
mainpath = "/Users/34686/Documents/GitHub/python-ml-course/datasets"
filename = "titanic/titanic3.xls"
fullpath = mainpath +"/"+ filename
titanic2 = pd.read_excel (fullpath, "titanic3") # titanic3 es la pestaña dentgro de la hoja excel de la ruta fullpath
titanic2.head()
# # Crear un CSV - excel - json
titanic3 = pd.read_excel(mainpath + "/" + filename, "titanic3")
titanic3.to_csv(mainpath + "/titanic/titanic_custom.csv")
titanic3.to_excel(mainpath + "/titanic/titanic_custom.xls")
titanic3.to_json(mainpath + "/titanic/titanic_custom.json"j
|
notebooks/01_Juan_mi primer notebook_data cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/nitish-01/vision/blob/main/LabelSmoothing.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g0GlIuz18vuL" colab={"base_uri": "https://localhost:8080/"} outputId="6f554bba-3799-40a6-8859-87c76ed92ab1"
from google.colab import drive
# Accessing My Google Drive
drive.mount('/content/drive', force_remount=True)
# + [markdown] id="ytAIwMpHDoFx"
# ## Unzip dataset on runtime session
# + id="pGfZJQqFDuEF" colab={"base_uri": "https://localhost:8080/"} outputId="346c4841-dffb-419c-affd-5c9603644a44"
# !unzip '/content/drive/My Drive/CSIE7512_DL_Project/STL10.zip'
# + [markdown] id="N1MCxs3h84lB"
# ## Define Model
# + id="YCCnwKmMM4iP"
import torch
import torch.nn as nn
import math
#from math import round
import torch.utils.model_zoo as model_zoo
def conv3x3(in_planes, out_planes, stride=1):
"3x3 convolution with padding"
return nn.Conv2d(in_planes, out_planes, kernel_size=3, stride=stride,
padding=1, bias=False)
class BasicBlock(nn.Module):
outchannel_ratio = 1
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(BasicBlock, self).__init__()
self.bn1 = nn.BatchNorm2d(inplanes)
self.conv1 = conv3x3(inplanes, planes, stride)
self.bn2 = nn.BatchNorm2d(planes)
self.conv2 = conv3x3(planes, planes)
self.bn3 = nn.BatchNorm2d(planes)
self.relu = nn.ReLU(inplace=True)
self.downsample = downsample
self.stride = stride
def forward(self, x):
out = self.bn1(x)
out = self.conv1(out)
out = self.bn2(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn3(out)
if self.downsample is not None:
shortcut = self.downsample(x)
featuremap_size = shortcut.size()[2:4]
else:
shortcut = x
featuremap_size = out.size()[2:4]
batch_size = out.size()[0]
residual_channel = out.size()[1]
shortcut_channel = shortcut.size()[1]
if residual_channel != shortcut_channel:
padding = torch.autograd.Variable(torch.cuda.FloatTensor(batch_size, residual_channel - shortcut_channel, featuremap_size[0], featuremap_size[1]).fill_(0))
out += torch.cat((shortcut, padding), 1)
else:
out += shortcut
return out
class Bottleneck(nn.Module):
outchannel_ratio = 4
def __init__(self, inplanes, planes, stride=1, downsample=None):
super(Bottleneck, self).__init__()
self.bn1 = nn.BatchNorm2d(inplanes)
self.conv1 = nn.Conv2d(inplanes, planes, kernel_size=1, bias=False)
self.bn2 = nn.BatchNorm2d(planes)
self.conv2 = nn.Conv2d(planes, (planes*1), kernel_size=3, stride=stride,
padding=1, bias=False)
self.bn3 = nn.BatchNorm2d((planes*1))
self.conv3 = nn.Conv2d((planes*1), planes * Bottleneck.outchannel_ratio, kernel_size=1, bias=False)
self.bn4 = nn.BatchNorm2d(planes * Bottleneck.outchannel_ratio)
self.relu = nn.ReLU(inplace=True)
self.downsample = downsample
self.stride = stride
def forward(self, x):
out = self.bn1(x)
out = self.conv1(out)
out = self.bn2(out)
out = self.relu(out)
out = self.conv2(out)
out = self.bn3(out)
out = self.relu(out)
out = self.conv3(out)
out = self.bn4(out)
if self.downsample is not None:
shortcut = self.downsample(x)
featuremap_size = shortcut.size()[2:4]
else:
shortcut = x
featuremap_size = out.size()[2:4]
batch_size = out.size()[0]
residual_channel = out.size()[1]
shortcut_channel = shortcut.size()[1]
if residual_channel != shortcut_channel:
padding = torch.autograd.Variable(torch.cuda.FloatTensor(batch_size, residual_channel - shortcut_channel, featuremap_size[0], featuremap_size[1]).fill_(0))
out += torch.cat((shortcut, padding), 1)
else:
out += shortcut
return out
class PyramidNet(nn.Module):
def __init__(self, dataset, depth, alpha, num_classes, bottleneck=False):
super(PyramidNet, self).__init__()
self.dataset = dataset
if self.dataset.startswith('cifar'):
self.inplanes = 16
if bottleneck == True:
n = int((depth - 2) / 9)
block = Bottleneck
else:
n = int((depth - 2) / 6)
block = BasicBlock
self.addrate = alpha / (3*n*1.0)
self.input_featuremap_dim = self.inplanes
self.conv1 = nn.Conv2d(3, self.input_featuremap_dim, kernel_size=3, stride=1, padding=1, bias=False)
self.bn1 = nn.BatchNorm2d(self.input_featuremap_dim)
self.featuremap_dim = self.input_featuremap_dim
self.layer1 = self.pyramidal_make_layer(block, n)
self.layer2 = self.pyramidal_make_layer(block, n, stride=2)
self.layer3 = self.pyramidal_make_layer(block, n, stride=2)
self.final_featuremap_dim = self.input_featuremap_dim
self.bn_final= nn.BatchNorm2d(self.final_featuremap_dim)
self.relu_final = nn.ReLU(inplace=True)
self.avgpool = nn.AvgPool2d(24)
self.fc = nn.Linear(self.final_featuremap_dim, num_classes)
elif dataset == 'imagenet':
blocks ={18: BasicBlock, 34: BasicBlock, 50: Bottleneck, 101: Bottleneck, 152: Bottleneck, 200: Bottleneck}
layers ={18: [2, 2, 2, 2], 34: [3, 4, 6, 3], 50: [3, 4, 6, 3], 101: [3, 4, 23, 3], 152: [3, 8, 36, 3], 200: [3, 24, 36, 3]}
if layers.get(depth) is None:
if bottleneck == True:
blocks[depth] = Bottleneck
temp_cfg = int((depth-2)/12)
else:
blocks[depth] = BasicBlock
temp_cfg = int((depth-2)/8)
layers[depth]= [temp_cfg, temp_cfg, temp_cfg, temp_cfg]
print('=> the layer configuration for each stage is set to', layers[depth])
self.inplanes = 64
self.addrate = alpha / (sum(layers[depth])*1.0)
self.input_featuremap_dim = self.inplanes
self.conv1 = nn.Conv2d(3, self.input_featuremap_dim, kernel_size=7, stride=2, padding=3, bias=False)
self.bn1 = nn.BatchNorm2d(self.input_featuremap_dim)
self.relu = nn.ReLU(inplace=True)
self.maxpool = nn.MaxPool2d(kernel_size=3, stride=2, padding=1)
self.featuremap_dim = self.input_featuremap_dim
self.layer1 = self.pyramidal_make_layer(blocks[depth], layers[depth][0])
self.layer2 = self.pyramidal_make_layer(blocks[depth], layers[depth][1], stride=2)
self.layer3 = self.pyramidal_make_layer(blocks[depth], layers[depth][2], stride=2)
self.layer4 = self.pyramidal_make_layer(blocks[depth], layers[depth][3], stride=2)
self.final_featuremap_dim = self.input_featuremap_dim
self.bn_final= nn.BatchNorm2d(self.final_featuremap_dim)
self.relu_final = nn.ReLU(inplace=True)
self.avgpool = nn.AvgPool2d(7)
self.fc = nn.Linear(self.final_featuremap_dim, num_classes)
for m in self.modules():
if isinstance(m, nn.Conv2d):
n = m.kernel_size[0] * m.kernel_size[1] * m.out_channels
m.weight.data.normal_(0, math.sqrt(2. / n))
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
def pyramidal_make_layer(self, block, block_depth, stride=1):
downsample = None
if stride != 1: # or self.inplanes != int(round(featuremap_dim_1st)) * block.outchannel_ratio:
downsample = nn.AvgPool2d((2,2), stride = (2, 2), ceil_mode=True)
layers = []
self.featuremap_dim = self.featuremap_dim + self.addrate
layers.append(block(self.input_featuremap_dim, int(round(self.featuremap_dim)), stride, downsample))
for i in range(1, block_depth):
temp_featuremap_dim = self.featuremap_dim + self.addrate
layers.append(block(int(round(self.featuremap_dim)) * block.outchannel_ratio, int(round(temp_featuremap_dim)), 1))
self.featuremap_dim = temp_featuremap_dim
self.input_featuremap_dim = int(round(self.featuremap_dim)) * block.outchannel_ratio
return nn.Sequential(*layers)
def forward(self, x):
if self.dataset == 'cifar10' or self.dataset == 'cifar100':
x = self.conv1(x)
x = self.bn1(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.bn_final(x)
x = self.relu_final(x)
x = self.avgpool(x)
x = x.view(x.size(0), -1)
x = self.fc(x)
elif self.dataset == 'imagenet':
x = self.conv1(x)
x = self.bn1(x)
x = self.relu(x)
x = self.maxpool(x)
x = self.layer1(x)
x = self.layer2(x)
x = self.layer3(x)
x = self.layer4(x)
x = self.bn_final(x)
x = self.relu_final(x)
x = self.avgpool(x)
x = x.view(x.size(0), -1)
x = self.fc(x)
return x
def Model():
r"""Return your custom model
"""
return PyramidNet('cifar10', 60, 48, 10)
# + [markdown] id="1CDgUpDcmq6-"
# ## utils
# + id="aw4uo1CL9M7j"
class AverageMeter(object):
r"""Computes and stores the average and current value
"""
def __init__(self, name, fmt=':f'):
self.name = name
self.fmt = fmt
self.reset()
def reset(self):
self.val = 0
self.avg = 0
self.sum = 0
self.count = 0
def update(self, val, n=1):
self.val = val
self.sum += val * n
self.count += n
self.avg = self.sum / self.count
def __str__(self):
fmtstr = '{name} {val' + self.fmt + '} ({avg' + self.fmt + '})'
return fmtstr.format(**self.__dict__)
class ProgressMeter(object):
def __init__(self, num_batches, *meters, prefix=""):
self.batch_fmtstr = self._get_batch_fmtstr(num_batches)
self.meters = meters
self.prefix = prefix
def print(self, batch):
entries = [self.prefix + self.batch_fmtstr.format(batch)]
entries += [str(meter) for meter in self.meters]
print('\t'.join(entries))
def _get_batch_fmtstr(self, num_batches):
num_digits = len(str(num_batches // 1))
fmt = '{:' + str(num_digits) + 'd}'
return '[' + fmt + '/' + fmt.format(num_batches) + ']'
def accuracy(output, target, topk=(1,)):
r"""Computes the accuracy over the $k$ top predictions for the specified values of k
"""
with torch.no_grad():
maxk = max(topk)
batch_size = target.size(0)
# _, pred = output.topk(maxk, 1, True, True)
# pred = pred.t()
# correct = pred.eq(target.view(1, -1).expand_as(pred))
# faster topk (ref: https://github.com/pytorch/pytorch/issues/22812)
_, idx = output.sort(descending=True)
pred = idx[:,:maxk]
pred = pred.t()
correct = pred.eq(target.view(1, -1).expand_as(pred))
res = []
for k in topk:
correct_k = correct[:k].reshape(-1).float().sum(0, keepdim=True)
res.append(correct_k.mul_(100.0 / batch_size))
return res
# + [markdown] id="_SxkjBIxolsm"
# ## Hyperparameters
# + id="Pv-hIWaJomEf"
SAVEPATH = '/content/drive/My Drive/CSIE7512_DL_Project/'
WEIGHTDECAY = 5e-4
MOMENTUM = 0.9
BATCHSIZE = 64
LR = 0.1
EPOCHS = 200
PRINTFREQ = 10
# + [markdown] id="9y7t4HTSm-9g"
# ## Train Model
# + id="yiGordRmyP-E" colab={"base_uri": "https://localhost:8080/"} outputId="7b76dfdc-684f-41f6-8252-c056c30dcbb4"
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
class LabelSmoothLoss(nn.Module):
def __init__(self, smoothing=0.0):
super(LabelSmoothLoss, self).__init__()
self.smoothing = smoothing
def forward(self, input, target):
log_prob = F.log_softmax(input, dim=-1)
weight = input.new_ones(input.size()) * \
self.smoothing / (input.size(-1) - 1.)
weight.scatter_(-1, target.unsqueeze(-1), (1. - self.smoothing))
loss = (-weight * log_prob).sum(dim=-1).mean()
return loss
def main():
model = Model()
##### optimizer / learning rate scheduler / criterion #####
optimizer = torch.optim.SGD(model.parameters(), lr=LR,
momentum=MOMENTUM, weight_decay=WEIGHTDECAY,
nesterov=True)
scheduler = torch.optim.lr_scheduler.MultiStepLR(optimizer, [100, 150],
gamma=0.1)
#criterion = torch.nn.CrossEntropyLoss()
criterion = LabelSmoothLoss(0.1)
###########################################################
model = model.cuda()
criterion = criterion.cuda()
# Check number of parameters your model
pytorch_total_params = sum(p.numel() for p in model.parameters())
print(f"Number of parameters: {pytorch_total_params}")
if int(pytorch_total_params) > 2000000:
print('Your model has the number of parameters more than 2 millions..')
exit()
normalize = transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225])
train_transform = transforms.Compose([
transforms.Resize((100,100)),
transforms.RandomRotation(15,),
transforms.RandomCrop(96),
transforms.RandomHorizontalFlip(),
transforms.ToTensor(),
normalize
])
train_dataset = torchvision.datasets.ImageFolder(
'./train', transform=train_transform)
train_loader = DataLoader(train_dataset,
batch_size=BATCHSIZE, shuffle=True,
num_workers=4, pin_memory=True)
last_top1_acc = 0
for epoch in range(EPOCHS):
print("\n----- epoch: {}, lr: {} -----".format(
epoch, optimizer.param_groups[0]["lr"]))
# train for one epoch
start_time = time.time()
last_top1_acc = train(train_loader, epoch, model, optimizer, criterion)
elapsed_time = time.time() - start_time
print('==> {:.2f} seconds to train this epoch\n'.format(
elapsed_time))
# learning rate scheduling
scheduler.step()
# Save model each epoch
if epoch > 150:
torch.save(model.state_dict(), SAVEPATH+'model_weight_'+ str(epoch).zfill(3) +'.pth')
print(f"Last Top-1 Accuracy: {last_top1_acc}")
print(f"Number of parameters: {pytorch_total_params}")
def train(train_loader, epoch, model, optimizer, criterion):
batch_time = AverageMeter('Time', ':6.3f')
data_time = AverageMeter('Data', ':6.3f')
losses = AverageMeter('Loss', ':.4e')
top1 = AverageMeter('Acc@1', ':6.2f')
top5 = AverageMeter('Acc@5', ':6.2f')
progress = ProgressMeter(len(train_loader), batch_time, data_time, losses,
top1, top5, prefix="Epoch: [{}]".format(epoch))
# switch to train mode
model.train()
end = time.time()
for i, (input, target) in enumerate(train_loader):
# measure data loading time
data_time.update(time.time() - end)
input = input.cuda()
target = target.cuda()
# compute output
output = model(input)
loss = criterion(output, target)
# measure accuracy and record loss, accuracy
acc1, acc5 = accuracy(output, target, topk=(1, 5))
losses.update(loss.item(), input.size(0))
top1.update(acc1[0].item(), input.size(0))
top5.update(acc5[0].item(), input.size(0))
# compute gradient and do SGD step
optimizer.zero_grad()
loss.backward()
optimizer.step()
# measure elapsed time
batch_time.update(time.time() - end)
end = time.time()
if i % PRINTFREQ == 0:
progress.print(i)
print('=> Acc@1 {top1.avg:.3f} Acc@5 {top5.avg:.3f}'
.format(top1=top1, top5=top5))
return top1.avg
if __name__ == "__main__":
main()
# + [markdown] id="EMzXV6lTykgJ"
# ## Make an evalutation csv file
#
# This code makes an evaluation csv file for kaggle submission.
#
# **Never change below code!!!**
# + id="MaUGSOCrAqx9" colab={"base_uri": "https://localhost:8080/"} outputId="f78382b0-09fd-4f31-89e0-bbcf0a7e4386"
import torch
import pandas as pd
import argparse
import time
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
def eval():
test_transform = transforms.Compose([
transforms.ToTensor()
])
test_dataset = torchvision.datasets.ImageFolder('./test', transform=test_transform)
test_loader = DataLoader(test_dataset, batch_size=BATCHSIZE, num_workers=4, shuffle=False)
model = Model()
model = model.cuda()
model.load_state_dict(torch.load(SAVEPATH+'model_weight_199.pth'))
print('Make an evaluation csv file for kaggle submission...')
Category = []
for input, _ in test_loader:
input = input.cuda()
output = model(input)
output = torch.argmax(output, dim=1)
Category = Category + output.tolist()
Id = list(range(0, 8000))
samples = {
'Id': Id,
'Category': Category
}
df = pd.DataFrame(samples, columns=['Id', 'Category'])
df.to_csv(SAVEPATH+'submission.csv', index=False)
print('Done!!')
if __name__ == "__main__":
eval()
|
LabelSmoothing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + hide_input=true init_cell=true
from IPython.core.display import display, HTML, Markdown, display_html, clear_output, Javascript
from ipywidgets import Button, HBox, VBox, widgets
from string import Template
import ipysheet
import pandas as pd
import json, random
import yaml
import math
import time
import re
import os
from os import path
from datetime import datetime
#verificatori
from exercise_3.verifier.lds import lds
from exercise_3.verifier.zSeq import zSequenza
from exercise_3.verifier.ldsWithNumber import ldsWithNumber
from exercise_3.verifier.ldscutrange import ldsSubwithoutElementInRange
from exercise_3.verifier.minNsubCre import minimoNumSubCRE
from exercise_3.verifier.final_results import save_results
# + hide_input=false init_cell=true
button_run = widgets.Button(description="Start exercise")
output_run = widgets.Output()
display(button_run, output_run)
def on_run_clicked1(b):
with output_run:
display(Javascript('''
function sleep2(milliseconds) {
var start = new Date().getTime();
for (var i = 0; i < 1e7; i++) {
if ((new Date().getTime() - start) > milliseconds){
break;
}
}
}
var k = 3
//IPython.notebook.execute_cells([0])
try{
while(k < 50){
IPython.notebook.execute_cells([k])
k = k+1
sleep2(50)
}
}catch{}
'''))
button_run.on_click(on_run_clicked1)
# + hide_input=true init_cell=true language="javascript"
# var kernel = IPython.notebook.kernel;
# var thename = window.document.getElementById("notebook_name").innerHTML;
# var command = "theNotebook = " + "'"+thename+"'";
# kernel.execute(command);
# + hide_input=true
str_list = theNotebook.split("_")
exercise = str_list[1]
# %run -i "exercise_3/script/load_config.py"
txt = open(yaml_text[0], 'r',encoding='utf-8')
content = txt.read()
text_str = "# "+ str_list[0].upper() + " " + str_list[1] + "\n" + content
Markdown(text_str)
# + hide_input=true
file = open(yaml_table_path, 'r',encoding='utf-8')
data = []
for line in file.readlines():
data.append(line.split(' '))
df = pd.DataFrame(data=data)
df.columns = [i for i in range(1, len(data[0])+1)]
display_html(df.to_html(index=False), raw=True)
global prog0
global command0
prog0=0
command0=0
# + hide_input=false
print("Qui puoi compilare la tabella di supporto")
if command0 == 0:
sheet1=ipysheet.sheet(rows=5,columns=len(df.columns),column_headers=False, row_headers=False, column_width=30)
values=df.values.tolist()
for i in range(0,len(df.columns)):
ipysheet.cell(0,i,"->",read_only=True)
ipysheet.cell(1,i,"0")
ipysheet.cell(2,i,values[0][i],read_only=True)
ipysheet.cell(3,i,"0")
ipysheet.cell(4,i,"<-",read_only=True)
if command0 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/tablesupp.txt", "r")
lines = f.readlines()
num=[]
for line in lines:
num.append([int(s) for s in re.findall(r'\b\d+\b', line)])
sheet1=ipysheet.sheet(rows=5,columns=len(df.columns),column_headers=False, row_headers=False, column_width=30)
values=df.values.tolist()
for i in range(0,len(df.columns)):
ipysheet.cell(0,i,"->",read_only=True)
ipysheet.cell(1,i,str(num[0][i]))
ipysheet.cell(2,i,values[0][i],read_only=True)
ipysheet.cell(3,i,str(num[1][i]))
ipysheet.cell(4,i,"<-",read_only=True)
HBox([sheet1], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet1
# + hide_input=true
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
center_box = VBox([items[1]])
right_box = VBox([items[2]])
def reset0(_):
global command0
command0=0
display(Javascript('IPython.notebook.execute_cells([5])'))
def save0(_):
DIR = "./exercise_3/data/log/checkpoint"
timestamp = datetime.timestamp(datetime.now())
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
f = open("./exercise_3/data/log/checkpoint/tablesupp_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet1)[1])))+"\n")
f.write(str(list(map(int,ipysheet.to_array(sheet1)[3]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/tablesupp.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet1)[1])))+"\n")
f.write(str(list(map(int,ipysheet.to_array(sheet1)[3]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([6])'))
def recall0(_):
global command0
if path.exists("./exercise_3/data/log/lastcheckpoint/tablesupp.txt"):
command0=1
else:
command0=0
display(Javascript('IPython.notebook.execute_cells([5])'))
items[0].on_click(reset0)
items[1].on_click(save0)
items[2].on_click(recall0)
HBox([left_box,center_box,right_box])
# + hide_input=true pycharm={"name": "#%%\n"}
global command1
command1 = 0
txt = open(yaml_text[1], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true
print("Lunghezza della sottosequenza decrescente più lunga:")
global maxsubseq
if command1 == 0:
maxsubseq=widgets.IntText(
value=1,
disabled=False
)
display(maxsubseq)
#9
# + hide_input=false
global maxsubseq
if command1 == 0:
lunghezza=int(maxsubseq.value)
sheet2=ipysheet.sheet(rows=1,columns=lunghezza,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza):
ipysheet.cell(0,i,"0")
if command1 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/query1.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza=len(num)
sheet2=ipysheet.sheet(rows=1,columns=lunghezza,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq
maxsubseq=widgets.IntText(
value=lunghezza,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([8])'))
if command1 == 2:
f = open("./exercise_3/data/log/lastans/ansquery1.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza=len(num)
sheet2=ipysheet.sheet(rows=1,columns=lunghezza,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq
maxsubseq=widgets.IntText(
value=lunghezza,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([8])'))
HBox([sheet2], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet2
#34, 42, 44, 49, 52, 63, 69, 79, 81
# + hide_input=false
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset1(_):
global command1
command1=0
display(Javascript('IPython.notebook.execute_cells([9])'))
def save1(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/checkpoint/query1_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet2)[0]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query1.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet2)[0]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([10])'))
def recall1(_):
global command1
if path.exists("./exercise_3/data/log/lastcheckpoint/query1.txt"):
command1=1
else:
command1=0
display(Javascript('IPython.notebook.execute_cells([9])'))
def verify1(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/ans/ansquery1_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet2)[0]))))
f.close()
f = open("./exercise_3/data/log/lastans/ansquery1.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet2)[0]))))
f.close()
res=lds(values[0], list(ipysheet.to_array(sheet2)[0]), lunghezza)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([10])'))
def recallverify1(_):
global command1
if path.exists("./exercise_3/data/log/lastans/ansquery1.txt"):
command1=2
else:
command1=0
display(Javascript('IPython.notebook.execute_cells([9])'))
items[0].on_click(reset1)
items[1].on_click(save1)
items[2].on_click(recall1)
items[3].on_click(verify1)
items[4].on_click(recallverify1)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=false pycharm={"name": "#%%\n"}
global command2
command2 = 0
txt = open(yaml_text[2], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true
print("Lunghezza della sottosequenza decrescente più lunga:")
global zseqlen
if command2==0:
zseqlen=widgets.IntText(
value=1,
disabled=False
)
display(zseqlen)
#14
# + hide_input=true
global zseqlen
if command2 == 0:
lunghezza2=int(zseqlen.value)
sheet3=ipysheet.sheet(rows=1,columns=lunghezza2,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza2):
ipysheet.cell(0,i,"0")
if command2 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/query2.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza2=len(num)
sheet3=ipysheet.sheet(rows=1,columns=lunghezza2,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza2):
ipysheet.cell(0,i,str(num[i]))
#global zseqlen
zseqlen=widgets.IntText(
value=lunghezza2,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([12])'))
if command2 == 2:
f = open("./exercise_3/data/log/lastans/ansquery2.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza2=len(num)
sheet3=ipysheet.sheet(rows=1,columns=lunghezza2,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza2):
ipysheet.cell(0,i,str(num[i]))
#global zseqlen
zseqlen=widgets.IntText(
value=lunghezza2,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([12])'))
HBox([sheet3], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet3
#34, 42, 44, 49, 52, 63, 69, 79, 81, 43, 46, 61, 64, 73
# + hide_input=true
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset2(_):
global command2
command2=0
display(Javascript('IPython.notebook.execute_cells([13])'))
def save2(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/checkpoint/query2_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet3)[0]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query2.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet3)[0]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([14])'))
def recall2(_):
global command2
if path.exists("./exercise_3/data/log/lastcheckpoint/query2.txt"):
command2=1
else:
command2=0
display(Javascript('IPython.notebook.execute_cells([13])'))
def verify2(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/ans/ansquery2_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet3)[0]))))
f.close()
f = open("./exercise_3/data/log/lastans/ansquery2.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet3)[0]))))
f.close()
res=zSequenza(values[0], list(ipysheet.to_array(sheet3)[0]), lunghezza2)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([14])'))
def recallverify2(_):
global command2
if path.exists("./exercise_3/data/log/lastans/ansquery2.txt"):
command2=2
else:
command2=0
display(Javascript('IPython.notebook.execute_cells([13])'))
items[0].on_click(reset2)
items[1].on_click(save2)
items[2].on_click(recall2)
items[3].on_click(verify2)
items[4].on_click(recallverify2)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=true pycharm={"name": "#%%\n"}
global command3
command3 = 0
txt = open(yaml_text[3], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true
print("Lunghezza della sottosequenza decrescente più lunga:")
global maxsubseq2
if command3 == 0:
maxsubseq2=widgets.IntText(
value=1,
disabled=False
)
display(maxsubseq2)
#7
# + hide_input=true
global maxsubseq2
if command3 == 0:
lunghezza3=int(maxsubseq2.value)
sheet4=ipysheet.sheet(rows=1,columns=lunghezza3,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza3):
ipysheet.cell(0,i,"0")
if command3==1:
f = open("./exercise_3/data/log/lastcheckpoint/query3.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza3=len(num)
sheet4=ipysheet.sheet(rows=1,columns=lunghezza3,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza3):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq2
maxsubseq2=widgets.IntText(
value=lunghezza3,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([16])'))
if command3==2:
f = open("./exercise_3/data/log/lastans/ansquery3.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza3=len(num)
sheet4=ipysheet.sheet(rows=1,columns=lunghezza3,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza3):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq2
maxsubseq2=widgets.IntText(
value=lunghezza3,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([16])'))
HBox([sheet4], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet4
#34, 40, 45, 54, 61, 64, 73
# + hide_input=true
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset3(_):
global command3
command3=0
display(Javascript('IPython.notebook.execute_cells([17])'))
def save3(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/checkpoint/query3_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet4)[0]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query3.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet4)[0]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([18])'))
def recall3(_):
global command3
if path.exists("./exercise_3/data/log/lastcheckpoint/query3.txt"):
command3=1
else:
command3=0
display(Javascript('IPython.notebook.execute_cells([17])'))
def verify3(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/ans/ansquery3_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet4)[0]))))
f.close()
f = open("./exercise_3/data/log/lastans/ansquery3.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet4)[0]))))
f.close()
res=ldsWithNumber(values[0], list(ipysheet.to_array(sheet4)[0]), lunghezza3, 59)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([18])'))
def recallverify3(_):
global command3
if path.exists("./exercise_3/data/log/lastans/ansquery3.txt"):
command3=2
else:
command3=0
display(Javascript('IPython.notebook.execute_cells([17])'))
items[0].on_click(reset3)
items[1].on_click(save3)
items[2].on_click(recall3)
items[3].on_click(verify3)
items[4].on_click(recallverify3)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=true pycharm={"name": "#%%\n"}
global command4
command4 = 0
txt = open(yaml_text[4], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true
print("Lunghezza della sottosequenza decrescente più lunga evitando i primi 4 elementi:")
global maxsubseq3
if command4 == 0:
maxsubseq3=widgets.IntText(
value=1,
disabled=False
)
display(maxsubseq3)
#6
# + hide_input=false
global maxsubseq3
if command4 == 0:
lunghezza4=int(maxsubseq3.value)
sheet5=ipysheet.sheet(rows=1,columns=lunghezza4,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza4):
ipysheet.cell(0,i,"0")
if command4 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/query4.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza4=len(num)
sheet5=ipysheet.sheet(rows=1,columns=lunghezza4,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza4):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq3
maxsubseq3=widgets.IntText(
value=lunghezza4,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([20])'))
if command4 == 2:
f = open("./exercise_3/data/log/lastans/ansquery4.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza4=len(num)
sheet5=ipysheet.sheet(rows=1,columns=lunghezza4,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza4):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq3
maxsubseq3=widgets.IntText(
value=lunghezza4,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([20])'))
HBox([sheet5], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet5
#41, 52, 63, 69, 79, 81
# + hide_input=true pycharm={"name": "#%%\n"}
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset4(_):
global command4
command4=0
display(Javascript('IPython.notebook.execute_cells([21])'))
def save4(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/checkpoint/query4_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet5)[0]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query4.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet5)[0]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([22])'))
def recall4(_):
global command4
if path.exists("./exercise_3/data/log/lastcheckpoint/query4.txt"):
command4=1
else:
command4=0
display(Javascript('IPython.notebook.execute_cells([21])'))
def verify4(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/ans/ansquery4_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet5)[0]))))
f.close()
f = open("./exercise_3/data/log/lastans/ansquery4.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet5)[0]))))
f.close()
res=ldsSubwithoutElementInRange(values[0], list(ipysheet.to_array(sheet5)[0]), lunghezza4, 0, 3)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([22])'))
def recallverify4(_):
global command4
if path.exists("./exercise_3/data/log/lastans/ansquery4.txt"):
command4=2
else:
command4=0
display(Javascript('IPython.notebook.execute_cells([21])'))
items[0].on_click(reset4)
items[1].on_click(save4)
items[2].on_click(recall4)
items[3].on_click(verify4)
items[4].on_click(recallverify4)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=true pycharm={"name": "#%%\n"}
global command5
command5 = 0
txt = open(yaml_text[5], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true
print("Lunghezza della sottosequenza decrescente più lunga evitando gli elementi dal 14-esimo al 17-esimo:")
global maxsubseq4
if command5 == 0:
maxsubseq4=widgets.IntText(
value=1,
disabled=False
)
display(maxsubseq4)
#9
# + hide_input=true
global maxsubseq4
if command5 == 0:
lunghezza5=int(maxsubseq4.value)
sheet6=ipysheet.sheet(rows=1,columns=lunghezza5,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza5):
ipysheet.cell(0,i,"0")
if command5 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/query5.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza5=len(num)
sheet6=ipysheet.sheet(rows=1,columns=lunghezza5,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza5):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq4
maxsubseq4=widgets.IntText(
value=lunghezza5,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([24])'))
if command5 == 2:
f = open("./exercise_3/data/log/lastans/ansquery5.txt", "r")
line = f.readline()
num=[int(s) for s in re.findall(r'\b\d+\b', line)]
lunghezza5=len(num)
sheet6=ipysheet.sheet(rows=1,columns=lunghezza5,column_headers=False, row_headers=False, column_width=30)
for i in range(0,lunghezza5):
ipysheet.cell(0,i,str(num[i]))
#global maxsubseq4
maxsubseq4=widgets.IntText(
value=lunghezza5,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([24])'))
HBox([sheet6], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet6
#34, 42, 44, 49, 52, 60, 61, 64, 73
# + hide_input=false pycharm={"name": "#%%\n"}
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset5(_):
global command5
command5=0
display(Javascript('IPython.notebook.execute_cells([25])'))
def save5(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/checkpoint/query5_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet6)[0]))))
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query5.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet6)[0]))))
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([26])'))
def recall5(_):
global command5
if path.exists("./exercise_3/data/log/lastcheckpoint/query5.txt"):
command5=1
else:
command5=0
display(Javascript('IPython.notebook.execute_cells([25])'))
def verify5(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
f = open("./exercise_3/data/log/ans/ansquery5_"+str(prog)+"_"+str(timestamp)+".txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet6)[0]))))
f.close()
f = open("./exercise_3/data/log/lastans/ansquery5.txt", "w")
f.write(str(list(map(int,ipysheet.to_array(sheet6)[0]))))
f.close()
res=ldsSubwithoutElementInRange(values[0], list(ipysheet.to_array(sheet6)[0]), lunghezza5, 13, 16)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([26])'))
def recallverify5(_):
global command5
if path.exists("./exercise_3/data/log/lastans/ansquery5.txt"):
command5=2
else:
command5=0
display(Javascript('IPython.notebook.execute_cells([25])'))
items[0].on_click(reset5)
items[1].on_click(save5)
items[2].on_click(recall5)
items[3].on_click(verify5)
items[4].on_click(recallverify5)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=true pycharm={"name": "#%%\n"}
global command6
command6 = 0
txt = open(yaml_text[6], 'r',encoding='utf-8')
content = txt.read()
Markdown(content)
# + hide_input=true pycharm={"name": "#%%\n"}
print("Inserisci il numero di sottosequenze:")
global nsubseq
if command6 == 0:
nsubseq=widgets.IntText(
value=1,
disabled=False
)
display(nsubseq)
#9
# + hide_input=true pycharm={"name": "#%%\n"}
print("Inserisci la dimensione della sottosequenza più lunga:")
global maxsubseq5
if command6 == 0:
maxsubseq5=widgets.IntText(
value=1,
disabled=False
)
display(maxsubseq5)
#4 sol errata
#5 sol corretta
# + hide_input=true pycharm={"name": "#%%\n"}
global nsubseq
global maxsubseq5
if command6 == 0:
r=int(nsubseq.value)
c=int(maxsubseq5.value)
sheet7=ipysheet.sheet(rows=r,columns=c,column_headers=False, row_headers=False, column_width=30)
for i in range(0,r):
for j in range(0,c):
ipysheet.cell(i,j,"0")
if command6 == 1:
f = open("./exercise_3/data/log/lastcheckpoint/query6.txt", "r")
lines = f.readlines()
num=[]
for line in lines:
num.append([int(s) for s in re.findall(r'\b\d+\b', line)])
r=len(num)
c=len(num[0])
sheet7=ipysheet.sheet(rows=r,columns=c,column_headers=False, row_headers=False, column_width=30)
for i in range(0,r):
for j in range(0,c):
ipysheet.cell(i,j,str(num[i][j]))
#global maxsubseq5
maxsubseq5=widgets.IntText(
value=c,
disabled=False
)
#global nsubseq
nsubseq=widgets.IntText(
value=r,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([28,29])'))
if command6 == 2:
f = open("./exercise_3/data/log/lastans/ansquery6.txt", "r")
lines = f.readlines()
num=[]
for line in lines:
num.append([int(s) for s in re.findall(r'\b\d+\b', line)])
r=len(num)
c=len(num[0])
sheet7=ipysheet.sheet(rows=r,columns=c,column_headers=False, row_headers=False, column_width=30)
for i in range(0,r):
for j in range(0,c):
ipysheet.cell(i,j,str(num[i][j]))
#global maxsubseq5
maxsubseq5=widgets.IntText(
value=c,
disabled=False
)
#global nsubseq
nsubseq=widgets.IntText(
value=r,
disabled=False
)
display(Javascript('IPython.notebook.execute_cells([28,29])'))
HBox([sheet7], layout={'max_width' : '1000px', 'overflow_y' : 'auto'})
#sheet7
#34 ; 42,41,40,38; 44,43; 49,45; 52,46; 63,60,54,48; 69,66,61; 86,79,64,47; 81,80,73
# + hide_input=false pycharm={"name": "#%%\n"}
words = ['Start from scratch', 'Make checkpoint', 'Recall last checkpoint', 'Get feedback', 'Recall last certificate']
items = [Button(description=w) for w in words]
left_box = VBox([items[0]])
cl_box = VBox([items[1]])
cc_box = VBox([items[2]])
cr_box = VBox([items[3]])
right_box = VBox([items[4]])
def reset6(_):
global command6
command6=0
display(Javascript('IPython.notebook.execute_cells([30])'))
def save6(_):
DIR = "./exercise_3/data/log/checkpoint"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
aux=ipysheet.to_array(sheet7)
f = open("./exercise_3/data/log/checkpoint/query6_"+str(prog)+"_"+str(timestamp)+".txt", "w")
for elem in aux:
f.write(str(list(map(int,elem)))+"\n")
f.close()
f = open("./exercise_3/data/log/lastcheckpoint/query6.txt", "w")
for elem in aux:
f.write(str(list(map(int,elem)))+"\n")
f.close()
print("Tabella salvata")
time.sleep(3)
display(Javascript('IPython.notebook.execute_cells([31])'))
def recall6(_):
global command6
if path.exists("./exercise_3/data/log/lastcheckpoint/query6.txt"):
command6=1
else:
command6=0
display(Javascript('IPython.notebook.execute_cells([30])'))
def verify6(_):
DIR = "./exercise_3/data/log/ans"
prog=len([name for name in os.listdir(DIR) if os.path.isfile(os.path.join(DIR, name))])+1
timestamp = datetime.timestamp(datetime.now())
aux=ipysheet.to_array(sheet7)
f = open("./exercise_3/data/log/ans/ansquery6_"+str(prog)+"_"+str(timestamp)+".txt", "w")
for elem in aux:
f.write(str(list(map(int,elem)))+"\n")
f.close()
f = open("./exercise_3/data/log/lastans/ansquery6.txt", "w")
for elem in aux:
f.write(str(list(map(int,elem)))+"\n")
f.close()
res=minimoNumSubCRE(values[0], list(ipysheet.to_array(sheet7)), r)
display(Markdown(res))
time.sleep(10)
display(Javascript('IPython.notebook.execute_cells([31])'))
def recallverify6(_):
global command6
if path.exists("./exercise_3/data/log/lastans/ansquery6.txt"):
command6=2
else:
command6=0
display(Javascript('IPython.notebook.execute_cells([30])'))
items[0].on_click(reset6)
items[1].on_click(save6)
items[2].on_click(recall6)
items[3].on_click(verify6)
items[4].on_click(recallverify6)
HBox([left_box,cl_box,cc_box,cr_box,right_box])
# + hide_input=false
button6 = widgets.Button(description='Invia tutti i risultati')
out = widgets.Output()
def on_button_clicked6(_):
with out:
save_results(res_path)
clear_output()
print("Risultati inviati")
button6.on_click(on_button_clicked6)
widgets.VBox([button6,out])
|
Applet/exercise_3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + raw_mimetype="text/restructuredtext" active=""
# Get Started
# ===========
#
# In this guide, you walk through examples where you initialize Woodwork on a DataFrame and on a Series. Along the way, you learn how to update and remove logical types and semantic tags. You also learn how to use typing information to select subsets of data.
#
# Types and Tags
# --------------
# Woodwork relies heavily on the concepts of physical types, logical types and semantic tags. These concepts are covered in detail in :doc:`guides/understanding_types_and_tags`, but we provide brief definitions here for reference:
#
# * Physical Type: defines how the data is stored on disk or in memory.
# * Logical Type: defines how the data should be parsed or interpreted.
# * Semantic Tag(s): provides additional data about the meaning of the data or how it should be used.
#
# Start learning how to use Woodwork by reading in a dataframe that contains retail sales data.
# +
import pandas as pd
df = pd.read_csv("https://api.featurelabs.com/datasets/online-retail-logs-2018-08-28.csv")
df.head(5)
# -
# As you can see, this is a dataframe containing several different data types, including dates, categorical values, numeric values, and natural language descriptions. Next, initialize Woodwork on this DataFrame.
# ## Initializing Woodwork on a DataFrame
# Importing Woodwork creates a special namespace on your DataFrames, `DataFrame.ww`, that can be used to set or update the typing information for the DataFrame. As long as Woodwork has been imported, initializing Woodwork on a DataFrame is as simple as calling `.ww.init()` on the DataFrame of interest. An optional name parameter can be specified to label the data.
# +
import woodwork as ww
df.ww.init(name="retail", make_index=True, index="order_product_id")
df.ww
# -
# Using just this simple call, Woodwork was able to infer the logical types present in the data by analyzing the DataFrame dtypes as well as the information contained in the columns. In addition, Woodwork also added semantic tags to some of the columns based on the logical types that were inferred. Because the original data did not contain an index column, Woodwork's `make_index` parameter was used to create a new index column in the DataFrame.
#
# All Woodwork methods and properties can be accessed through the `ww` namespace on the DataFrame. DataFrame methods called from the Woodwork namespace will be passed to the DataFrame, and whenever possible, Woodwork will be initialized on the returned object, assuming it is a Series or a DataFrame.
#
# As an example, use the `head` method to create a new DataFrame containing the first 5 rows of the original data, with Woodwork typing information retained.
head_df = df.ww.head(5)
head_df.ww
head_df
# + raw_mimetype="text/restructuredtext" active=""
# .. note::
# Once Woodwork is initialized on a DataFrame, it is recommended to go through the ``ww`` namespace when performing DataFrame operations to avoid invalidating Woodwork's typing information.
# -
# ## Updating Logical Types
# If the initial inference was not to our liking, the logical type can be changed to a more appropriate value. Let's change some of the columns to a different logical type to illustrate this process. In this case, set the logical type for the `order_product_id` and `country` columns to be `Categorical` and set `customer_name` to have a logical type of `PersonFullName`.
df.ww.set_types(logical_types={
'customer_name': 'PersonFullName',
'country': 'Categorical',
'order_id': 'Categorical'
})
df.ww.types
# Inspect the information in the `types` output. There, you can see that the Logical type for the three columns has been updated with the logical types you specified.
# ## Selecting Columns
#
# Now that you've prepared logical types, you can select a subset of the columns based on their logical types. Select only the columns that have a logical type of `Integer` or `Double`.
numeric_df = df.ww.select(['Integer', 'Double'])
numeric_df.ww
# This selection process has returned a new Woodwork DataFrame containing only the columns that match the logical types you specified. After you have selected the columns you want, you can use the DataFrame containing just those columns as you normally would for any additional analysis.
numeric_df
# ## Adding Semantic Tags
#
# Next, let’s add semantic tags to some of the columns. Add the tag of `product_details` to the `description` column, and tag the `total` column with `currency`.
df.ww.set_types(semantic_tags={'description':'product_details', 'total': 'currency'})
df.ww
# Select columns based on a semantic tag. Only select the columns tagged with `category`.
category_df = df.ww.select('category')
category_df.ww
# Select columns using multiple semantic tags or a mixture of semantic tags and logical types.
category_numeric_df = df.ww.select(['numeric', 'category'])
category_numeric_df.ww
mixed_df = df.ww.select(['Boolean', 'product_details'])
mixed_df.ww
# To select an individual column, specify the column name. Woodwork will be initialized on the returned Series and you can use the Series for additional analysis as needed.
total = df.ww['total']
total.ww
total
# Select multiple columns by supplying a list of column names.
multiple_cols_df = df.ww[['product_id', 'total', 'unit_price']]
multiple_cols_df.ww
# ## Removing Semantic Tags
# Remove specific semantic tags from a column if they are no longer needed. In this example, remove the `product_details` tag from the `description` column.
df.ww.remove_semantic_tags({'description':'product_details'})
df.ww
# Notice how the ``product_details`` tag has been removed from the ``description`` column. If you want to remove all user-added semantic tags from all columns, you can do that, too.
df.ww.reset_semantic_tags()
df.ww
# ## Set Index and Time Index
# At any point, you can designate certain columns as the Woodwork `index` or `time_index` with the methods [set_index](generated/woodwork.table_accessor.WoodworkTableAccessor.set_index.rst) and [set_time_index](generated/woodwork.table_schema.TableSchema.set_time_index.rst). These methods can be used to assign these columns for the first time or to change the column being used as the index or time index.
#
# Index and time index columns contain `index` and `time_index` semantic tags, respectively.
df.ww.set_index('order_product_id')
df.ww.index
df.ww.set_time_index('order_date')
df.ww.time_index
df.ww
# + raw_mimetype="text/restructuredtext" active=""
# Using Woodwork with a Series
# ----------------------------
#
# Woodwork also can be used to store typing information on a Series. There are two approaches for initializing Woodwork on a Series, depending on whether or not the Series dtype is the same as the physical type associated with the LogicalType. For more information on logical types and physical types, refer to :doc:`guides/understanding_types_and_tags`.
#
# If your Series dtype matches the physical type associated with the specified or inferred LogicalType, Woodwork can be initialized through the `ww` namespace, just as with DataFrames.
# -
series = pd.Series([1, 2, 3], dtype='int64')
series.ww.init(logical_type='Integer')
series.ww
# In the example above, we specified the `Integer` LogicalType for the Series. Because `Integer` has a physical type of `int64` and this matches the dtype used to create the Series, no Series dtype conversion was needed and the initialization succeeds.
#
# In cases where the LogicalType requires the Series dtype to change, a helper function `ww.init_series` must be used. This function will return a new Series object with Woodwork initialized and the dtype of the series changed to match the physical type of the LogicalType.
#
# To demonstrate this case, first create a Series, with a `string` dtype. Then, initialize a Woodwork Series with a `Categorical` logical type using the `init_series` function. Because `Categorical` uses a physical type of `category`, the dtype of the Series must be changed, and that is why we must use the `init_series` function here.
#
# The series that is returned will have Woodwork initialized with the LogicalType set to `Categorical` as expected, with the expected dtype of `category`.
string_series = pd.Series(['a', 'b', 'a'], dtype='string')
ww_series = ww.init_series(string_series, logical_type='Categorical')
ww_series.ww
# As with DataFrames, Woodwork provides several methods that can be used to update or change the typing information associated with the series. As an example, add a new semantic tag to the series.
series.ww.add_semantic_tags('new_tag')
series.ww
# As you can see from the output above, the specified tag has been added to the semantic tags for the series.
#
# You can also access Series properties methods through the Woodwork namespace. When possible, Woodwork typing information will be retained on the value returned. As an example, you can access the Series `shape` property through Woodwork.
series.ww.shape
# You can also call Series methods such as `sample`. In this case, Woodwork typing information is retained on the Series returned by the `sample` method.
sample_series = series.ww.sample(2)
sample_series.ww
sample_series
# ## List Logical Types
# Retrieve all the Logical Types present in Woodwork. These can be useful for understanding the Logical Types, as well as how they are interpreted.
# +
from woodwork.type_sys.utils import list_logical_types
list_logical_types()
|
docs/source/start.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# *Creating a Jupyter Book with The Turing Way*
#
#
# # Module 3: Creating a Jupyter Book with The Turing Way
#
# **Learning Objective:**
# - Explain what files exist in the repository that we will use for the hands on session in this module (if you haven't already, please download the data required for this tutorial described in [module 1](./1-welcome.ipynb))
# - Explore the important/minimal components for creating a Jupyter Book
# - Build the first minimal version of the Jupyter Book locally using example files from _The Turing Way_
# <!---Set up the repository, add a chapter, create a table of content, build the book--->
#
# 📹 [VIDEO](https://www.youtube.com/watch?v=Dr_4H0Xr2Ko&list=PLBxcQEfGu3Dmdo6oKg6o9V7Q_e7WSX-vu&index=4)
# ---
# ## Introduction to Jupyter Book
#
# Welcome! In this Jupyter Notebook we will introduce the basic commands to generate your first Jupyter Book.
#
# In the previous module, we briefly looked into the awesome and very detailed [documentation](https://jupyterbook.org/intro.html) of Jupyter Book, and its [GitHub repository](https://github.com/executablebooks/jupyter-book).
#
# Jupyter Book has a [command-line interface](https://jupyterbook.org/reference/cli.html), so in this tutorial we will show you how to build your book using iPython's special syntax that lets you execute shell commands from a Jupyter Notebook. In this example we will do so by prefixing `!` in each cell.
#
# *__TIP__: If you are unfamiliar with executing shell commands from Jupyter Notebooks, read this [tutorial by <NAME>](https://jakevdp.github.io/PythonDataScienceHandbook/01.05-ipython-and-shell-commands.html).*
# ## Creating the content of your book based on _The Turing Way_
#
# In order to build our Jupyter Book, we first need to create a folder where all the content of your book exists.
# This is also where Jupyter Book will create the `html` files in order to host the book online.
#
# As demonstrated for *The Turing Way*, we will store all the content of our book in a folder named `book` located in the main repository (it doesn't need to be named this way for Jupyter Book to work). Let's create it:
# !mkdir ../book/
# The folder should be empty as you have just created it!
#
# __NOTE__: If this folder already exists, you are probably re-running this notebook without having cleared the `book` folder. Make sure you remove it using this command:
#
# ```
# $ rm -R ../book/
# ```
# ### Example files for this tutorial
#
# We will now add some contents such as a book chapter and other important components to our `book` folder.
#
# In this tutorial, we will be using some of the chapters from _[The Turing Way](https://the-turing-way.netlify.app/welcome.html)_.
# We have provided some example files, which are placed under the folder `content` in the main repository.
#
# Let's inspect whats inside it:
# !ls ../content
# Inside `content` there are 2 Markdown files: `welcome.md` and `reproducible-research.md`.
# These are the [welcome page](https://the-turing-way.netlify.app/welcome) and the introduction to the [Guide for Reproducible Research](https://the-turing-way.netlify.app/reproducible-research/reproducible-research.html) in _The Turing Way_ respectively.
#
# The `references.bib` is a bibtex file, which contains all the references cited in the book (to read more about citing using sphinx and Jupyter Book go [here](https://jupyterbook.org/content/citations.html#)).
#
# The `figures` folder contains all the images and figures used in these files.
#
# There are also two folders, `overview` and `open`, that contain the content of the `Overview` and `Open Research` chapters respectively (you can see them in the online [_The Turing Way_ book](https://the-turing-way.netlify.app/reproducible-research/overview.html)).
#
# The folder `content` also contains some Jupyter Notebooks (titled `demo` and `demo_2`), but we will cover the use of these in a later module.
# ### Let's start!
#
# For our first Jupyter Book we don't want to include all these files.
# Let's import to our `book` folder the files we will use one by one.
#
# We first need an index page for our book, which corresponds to the file `welcome.md` in _The Turing Way_.
# Let's copy it into our `book` folder:
# !cp ../content/welcome.md ../book/
# We will now need a chapter. Let's use the second Markdown file `reproducible-research.md` as our first chapter.
# !cp ../content/reproducible-research.md ../book/reproducible-research.md
# These files link to some citations and images placed in the `references.bib` file and `figures` folder, which we will also copy to our `book` folder:
# !cp -R ../content/figures/ ../book/figures
# !cp ../content/references.bib ../book/references.bib
# Let's make sure our `book` folder now contains all the files we want:
# !ls ../book/
# You should see printed a `figures`, `references.bib`, `welcome.md` and `reproducible-research.md` file.
# **We are almost there!**
#
# But we can not yet build our Jupyter Book.
# We have to specify the structure of our book - how will your book look like.
# Let's do this in the next section.
# ## Creating the structure of your Jupyter Book (`_toc.yml`)
#
# A book always has a structure, that is, a __Table Of Contents (TOC)__ that defines the hierarchy and order of the content-files.
#
# Similarly, to create a Jupyter Book we need to have a file that specifies the TOC. Jupyter Book defines the TOC using a [yaml](https://en.wikipedia.org/wiki/YAML) file that needs to be named `_toc.yml`. This can be created manually, or automatically by running:
#
# ```shell
# $ jupyter-book toc {path}
# ```
#
# as long as the intended structure is also ordered alphanumerically.
# But in _The Turing Way_ we are always creating new content and restructuring our book. In these cases, is not practical to have our filenames ordered alphanumerically. We will thus show you how to manually create our `_toc.yml` using the python library [ruamel.yaml](https://yaml.readthedocs.io/en/latest/):
# +
from ruamel.yaml import YAML
yaml = YAML()
# Define the contents of our _toc.yml
toc_document = """
- file: welcome
- file: reproducible-research
"""
# Save _toc.yml in the book directory
toc_file = open('../book/_toc.yml', 'w')
yaml.dump(yaml.load(toc_document), toc_file)
# -
# Important things to note of our example are:
#
# - The `file:` specifies to the location of our markdown files relative to the main directory.
# - The first `file` defined in our `_toc.yml` will be the index (opening page) of our book.
#
# __Note:__ You don't need Python to create your `_toc.yml`, you can also do it with any text editor.
# ## Building your book
# Let's check that our `book` directory now contains the `_toc.yml` file:
# !ls ../book/
# It does! That means we can now create our first Jupyter Book. This can be achieved by executing the command
#
# ```shell
# $ jupyter-book build {path-to-book}
# ```
#
# Read more about this command and its options [here](https://jupyterbook.org/reference/cli.html#jupyter-book-build).
#
# Let's try it:
# !jupyter-book build ../book/
# ***Congratulations! You have built your first Jupyter Book!***
#
# We now have a `_build` folder in our book path that contains the `html` files of our book.
# !ls ../book/
# This means we can locally deploy our book!
#
# To get a preview, paste the path specified at the very bottom of the `jupyter-book build` output into a new tab on your browser.
# The link should look like this: `file:///Users/PATH/jupytercon-2020/jupytercon_tutorial/book/_build/html/index.html` (please update the `Users/PATH`).
# ## Adding sections and chapters to your Jupyter Book
# We will now add the `Overview` chapter to our Jupyter Book.
#
# To do so, let's first copy the `overview` folder from the `content` to the `book` folder:
# !cp -R ../content/overview ../book/overview
# Let's also inspect the files inside the overview folder.
# !ls ../book/overview/
# There are 4 Markdown files. One is `overview.md`, which is the landing page for the `Overview` chapter.
#
# We also have files corresponding to the 3 subsections of the `Overview` chapter: `overview-definitions.md`, `overview-benefit.md` and `overview-resources.md` (you can inspect them in *The Turing Way* [here](https://the-turing-way.netlify.app/reproducible-research/overview.html)).
# What happens if we re-build our book?
# !jupyter-book build ../book/
# Inspect your deployed Jupyter Book. It did not change at all! Why? Jupyter Book gives us a warning: the files that we have moved over have not been included in our `_toc.yml`.
#
# We will need to update the `_toc.yml` to include `Overview` as a chapter in our book, and define the rest of the files as sub-chapters of it by specifying them as `sections`, like so:
# +
from ruamel.yaml import YAML
yaml = YAML()
# Define the contents of our _toc.yml
toc_document = """
- file: welcome
- file: reproducible-research
- file: overview/overview
title: Overview
sections:
- file: overview/overview-definitions.md
- file: overview/overview-benefit.md
- file: overview/overview-resources.md
"""
# Save _toc.yml in the book directory
toc_file = open('../book/_toc.yml', 'w')
yaml.dump(yaml.load(toc_document), toc_file)
# -
# Note that we have used `title:` under the `file:` description of the overview chapter.
# This is to provide a title to this chapter that will appear in the index of the navigation bar (in the left hand side of the book).
#
# If `title` is not specified (as in our welcome file), the first title in our markdown file will be used.
#
# Let's build our Jupyter Book again to incorporate the new changes:
# !jupyter-book build ../book/
# #### EXERCISE
#
# So far, we only have one chapter in our book (`Overview`), let's add the `Open Research` chapter by taking the following steps:
# 1. Copy the `open` folder from `content` to your `book folder`
# !cp -r ../content/open ../book
# +
# 2. Inspect the files inside the `book` folder
# +
# 3. Update the _toc.yml file to include the chapter `open.md` and name it `Open Research`
# 4. Add subchapters (other .md) to the `open` chapter, in your preferred order
# +
# Solution: uncomment the command below and run it to see the yml file for steps 3 and 4
# #%load ../answers/module_3/exercise-module-3.txt
# +
# 5. Re-build the updated Jupyter Book
# -
# ### Defining nested sections
#
# We now have multiple chapters and subchapters in our Jupyter Book!
#
# What if the two chapters `Overview` and `Open Research` were displayed under `Guide for Reproducible Research` in our book - just how they appear in _[The Turing Way](https://the-turing-way.netlify.app/reproducible-research/reproducible-research.html)_?
#
# Turns out we can define this in our `_toc.yml` by nesting them as `sections` of `Guide for Reproducible Research`, like so:
# +
yaml = YAML()
# Define the contents of our _toc.yml
# Optional: add titles to all our chapters
toc_document = """
- file: welcome
- file: reproducible-research
sections:
- file: overview/overview
title: Overview
sections:
- file: overview/overview-definitions
title: Definitions
- file: overview/overview-benefit
title: Benefits
- file: overview/overview-resources
title: Resources
- file: open/open
title: Open Research
sections:
- file: open/open-data
title: Open Data
- file: open/open-source
title: Open Source
- file: open/open-hardware
title: Open Hardware
- file: open/open-access
title: Open Access
- file: open/open-notebooks
title: Open Notebooks
- file: open/open-scholarship
title: Open Scholarship
"""
# Save _toc.yml in the book directory
toc_file = open('../book/_toc.yml', 'w')
yaml.dump(yaml.load(toc_document), toc_file)
# -
# !jupyter-book build ../book/
# #### Further reading
#
# What if instead we would have wanted the section `Guide for Reproducible Research` to be a part of our book that is not associated with its own file, but has files underneath it?
#
# To complete this exercise:
#
# 1. Read more about Parts/Chapters/Sections and how to specify them in the [official documentation](https://jupyterbook.org/customize/toc.html).
# 2. Define the `Guide for Reproducible Research` as a part of your book, and `Overview` and `Open Research` as sections of it.
#
# __HINT__: You will no longer need the `reproducible-research.md` file.
#
# __Answer:__ Run the command below to check that your `_toc.yml` looks the same
# !cat ../answers/module_3/ex2.yml
# ## Numbering your Jupyter Book sections
#
# We can also decide to number the sections in our Jupyter Book. This can be achieved by using the key `numbered` and set it to true.
#
# Let's see an example:
# +
yaml = YAML()
# Define the contents of our _toc.yml
toc_document = """
- file: welcome
numbered: true
- file: reproducible-research
title: Reproducibility Guide
sections:
- file: overview/overview
title: Overview
sections:
- file: overview/overview-definitions
title: Definitions
- file: overview/overview-benefit
title: Benefits
- file: overview/overview-resources
title: Resources
- file: open/open
title: Open Research
sections:
- file: open/open-data
title: Open Data
- file: open/open-source
title: Open Source
- file: open/open-hardware
title: Open Hardware
- file: open/open-access
title: Open Access
- file: open/open-notebooks
title: Open Notebooks
- file: open/open-scholarship
title: Open Scholarship
"""
# Save _toc.yml in the book directory
toc_file = open('../book/_toc.yml', 'w')
yaml.dump(yaml.load(toc_document), toc_file)
# -
# !jupyter-book build ../book/
# #### Further reading
# We might want to create separate numberings for each section of the book. Read how to do so [here](https://jupyterbook.org/customize/toc.html#number-your-book-s-chapters-and-sections) and number the sections under the `Overview` and `Open Research` chapters separately.
#
# __Answer:__ Run the command below to check that your `_toc.yml` looks the same
# !cat ../answers/module_3/ex3.yml
# 🗝 Takeaway
# ---
#
# - Jupyter Book uses markdown files to create chapters of an online book. In this tutorial, these files are placed in a directory called `book`.
# - All the images used in the file should also be stored in `book` (we saved them under the folder `figures` in this tutorial).
# - References used in the book should be defined in a `references.bib` file, which should also be stored in `book`.
# - A `_toc.yml` should be generated inside the folder `book` to structure the Jupyter Book (`toc` is the abbreviation for table of content).
# - With these files in place, you can create a minimal book using the command `$ jupyter-book build {path-to-book}`.
#
# #### RESOURCES
#
# - Read more about customization of the table of contents in the [official documentation](https://jupyterbook.org/customize/toc.html).
# 👉 [Next Module](./4-config-jupyterbook.ipynb)
# ---
|
notebooks/3-setup-jupyterbook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="052cb52c"
# [](https://colab.research.google.com/github/QData/super_sac/blob/master/docs/2demo/Demo.ipynb)
#
# [](https://github.com/QData/super_sac/blob/master/docs/2demo/Demo.ipynb)
# + [markdown] id="0Ie3gcwDiXFh"
# ## Demo to come
|
docs/2demo/Demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # Gmail - Automate response from keywords in mailbox
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/Gmail/Gmail_Automate_response_from_keywords_in_mailbox.ipynb" target="_parent">
# <img src="https://img.shields.io/badge/-Open%20in%20Naas-success?labelColor=000000&logo=data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iMTAyNHB4IiBoZWlnaHQ9IjEwMjRweCIgdmlld0JveD0iMCAwIDEwMjQgMTAyNCIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayIgdmVyc2lvbj0iMS4xIj4KIDwhLS0gR2VuZXJhdGVkIGJ5IFBpeGVsbWF0b3IgUHJvIDIuMC41IC0tPgogPGRlZnM+CiAgPHRleHQgaWQ9InN0cmluZyIgdHJhbnNmb3JtPSJtYXRyaXgoMS4wIDAuMCAwLjAgMS4wIDIyOC4wIDU0LjUpIiBmb250LWZhbWlseT0iQ29tZm9ydGFhLVJlZ3VsYXIsIENvbWZvcnRhYSIgZm9udC1zaXplPSI4MDAiIHRleHQtZGVjb3JhdGlvbj0ibm9uZSIgZmlsbD0iI2ZmZmZmZiIgeD0iMS4xOTk5OTk5OTk5OTk5ODg2IiB5PSI3MDUuMCI+bjwvdGV4dD4KIDwvZGVmcz4KIDx1c2UgaWQ9Im4iIHhsaW5rOmhyZWY9IiNzdHJpbmciLz4KPC9zdmc+Cg=="/>
# </a>
# + [markdown] papermill={} tags=[]
# <img width="20%" alt="Naas" src="https://cdn.vox-cdn.com/thumbor/K-q2WRPRyxxzzPLjxHGt26swMfM=/0x0:1320x880/1200x800/filters:focal(555x335:765x545)/cdn.vox-cdn.com/uploads/chorus_image/image/67587450/newgmaillogo.0.jpg"/>
# + [markdown] papermill={} tags=[]
# ## Account credentials
# + papermill={} tags=[]
username = "**********<EMAIL>"
to = "**********<EMAIL>"
password = "**********"
smtp_server = "imap.gmail.com"
box = "INBOX"
# + [markdown] papermill={} tags=[]
# ## Connect to email box
# + papermill={} tags=[]
import naas_drivers
emails = naas_drivers.email.connect(username,
password,
username,
smtp_server)
# + [markdown] papermill={} tags=[]
# ## Get email list
# + papermill={} tags=[]
dataframe = emails.get(criteria="ALL")
dataframe
# + [markdown] papermill={} tags=[]
# ## Automated reponse
# + papermill={} tags=[]
import naas
from re import search
for df in dataframe["text"]:
text = df.lower()
if search("sales report", text):
email_to = "<EMAIL>"
subject = "Sales Report"
content = "Hi \n,Here I am attaching the sales report as per your request\n.With Regards\n,NAAS Team"
files = ["Excel-Sales_Feb2020.csv"]
naas.notifications.send(email_to=email_to, subject=subject, html=content, files=files)
|
Gmail/Gmail_Automate_response_from_keywords_in_mailbox.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='top'></a>
# # แบบฝึกหัด Decision Tree
#
# ในแบบฝึกหัดนี้เราจะมาเขียนโปรแกรมทำ Decision Tree ใน Python กัน
# ในแบบฝึกหัด เราจะเรียนรู้เกี่ยวกับ
#
# 1. [การอ่านข้อมูลเข้าระบบ จากไฟล์ CSV](#section_1)
# 2. [การเตรียมข้อมูลเข้า](#section_2)
# 3. [การสร้าง Decision Tree โดยใช้เทคนิค CART](#section_3)
# 4. [การประเมินความแม่นยำของโมเดล Decision Tree ที่เราสร้าง](#section_4)
# ----
# <a id='section_1'></a>
# # 1. การอ่านข้อมูลเข้าระบบ จากไฟล์ CSV
#
# ในแบบฝึกหัดนี้จะใช้ข้อมูลจาก http://archive.ics.uci.edu/ml/datasets/banknote+authentication
#
# ซึ่งมีข้อมูล 5 คอลัมน์ดังนี้
# 1. variance of Wavelet Transformed image (continuous)
# 2. skewness of Wavelet Transformed image (continuous)
# 3. curtosis of Wavelet Transformed image (continuous)
# 4. entropy of image (continuous)
# 5. class (integer)
# +
# Test CART on Bank Note dataset
import random
import math
import pandas as pd
random.seed(1)
# -
# อ่านข้อมูลในไฟล์แต่ละบรรทัดเข้ามาเป็น `string`
data = pd.read_csv('http://archive.ics.uci.edu/ml/machine-learning-databases/00267/data_banknote_authentication.txt',
header=None)
print(f'ชุดข้อมูลมี : {len(data)} ตัวอย่าง')
# แปลงข้อมูลจาก `string` เป็น `list` ของ `float`
print(f'มีข้อมูล {len(set(data.iloc[:,-1]))} กลุ่ม คือ {set(data.iloc[:,-1])}')
# นับจำนวนตัวอย่างในแต่ละกลุ่ม
print(f'มีข้อมูลกลุ่ม 0 จำนวน {sum(data.iloc[:,-1]==0)} ตัวอย่าง')
print(f'มีข้อมูลกลุ่ม 1 จำนวน {sum(data.iloc[:,-1]==1)} ตัวอย่าง')
# ([กลับขึ้นข้างบน](#top))
#
# ----
# <a id='section_2'></a>
# # 2. การเตรียมข้อมูลเข้า
#
# แบ่งข้อมูลเป็น 2 กลุ่ม สำหรับ training และ test โดยแบ่งเป็น test data 20% และ training data 80%
def train_test_split(df, test_size = 0.2):
idx_train = random.choices([True,False], weights=[1-test_size, test_size], k=len(df))
idx_test = [not idx for idx in idx_train]
data_train = df.iloc[idx_train,:]
data_test = df.iloc[idx_test,:]
return data_train, data_test
test_size = 0.2
train_set, test_set = train_test_split(data, test_size)
# +
print(f'ข้อมูล training มี {train_set.shape[0]} ตัวอย่าง แต่ละตัวอย่างมี {train_set.shape[1]} คอลัมน์')
print(f'ข้อมูล training มี {len(set(train_set.iloc[:,-1]))} กลุ่ม คือ {set(train_set.iloc[:,-1])}')
print(f'มีข้อมูลกลุ่ม 0 จำนวน {sum(train_set.iloc[:,-1]==0)} ตัวอย่าง')
print(f'มีข้อมูลกลุ่ม 1 จำนวน {sum(train_set.iloc[:,-1]==1)} ตัวอย่าง')
print(f'ข้อมูล test มี {test_set.shape[0]} ตัวอย่าง แต่ละตัวอย่างมี {test_set.shape[1]} คอลัมน์')
print(f'ข้อมูล test มี {len(set(test_set.iloc[:,-1]))} กลุ่ม คือ {set(test_set.iloc[:,-1])}')
print(f'มีข้อมูลกลุ่ม 0 จำนวน {sum(test_set.iloc[:,-1]==0)} ตัวอย่าง')
print(f'มีข้อมูลกลุ่ม 1 จำนวน {sum(test_set.iloc[:,-1]==1)} ตัวอย่าง')
# -
# ([กลับขึ้นข้างบน](#top))
#
# ----
# <a id='section_3'></a>
# # 3. การสร้าง Decision Tree
#
# ## 3.1 Gini Index
# Calculate the Gini index for a split dataset
def gini_index(y_gr1, y_gr2):
# count all samples at split point
n_instances = float(len(y_gr1) + len(y_gr2))
classes = set(y_gr1 + y_gr2)
# sum weighted Gini index for each group
gini = 0.0
for group in [y_gr1, y_gr2]:
size = float(len(group))
# avoid divide by zero
if size == 0:
continue
score = 0.0
# score the group based on the score for each class
for class_val in classes:
p = group.count(class_val) / size
score += p * p
# weight the group score by its relative size
gini += (1.0 - score) * (size / n_instances)
return gini
print(gini_index([1, 0], [1, 0]))
# ผลลัพธ์ที่ควรจะได้
#
# `0.5`
print(gini_index([0, 0], [1, 1]))
# ผลลัพธ์ที่ควรจะได้
#
# `0.0`
# ## 3.2 Evaluate All Split
def test_split(index, thr, x_train, y_train):
left_x, left_y, right_x, right_y = list(), list(), list(), list()
for x, y in zip(x_train, y_train):
if x[index] <= thr:
left_x.append(x)
left_y.append(y)
else:
right_x.append(x)
right_y.append(y)
return left_x, left_y, right_x, right_y
def get_split(x_train, y_train):
b_index, b_value, b_score, b_groups = None, None, math.inf, None
for index in range(len(x_train[0])):
for x, y in zip(x_train, y_train):
groups = test_split(index, x[index], x_train, y_train)
left_x, left_y, right_x, right_y = groups
gini = gini_index(left_y, right_y)
if gini < b_score:
b_index, b_value, b_score, b_groups = index, x[index], gini, groups
return {'index':b_index, 'value':b_value, 'score':b_score, 'groups':b_groups}
x_sample = [[2.771244718,1.784783929],
[1.728571309,1.169761413],
[3.678319846,2.81281357],
[3.961043357,2.61995032],
[2.999208922,2.209014212],
[7.497545867,3.162953546],
[9.00220326,3.339047188],
[7.444542326,0.476683375],
[10.12493903,3.234550982],
[6.642287351,3.319983761]]
y_sample = [0, 0, 0, 0, 0, 1, 1, 1, 1, 1]
split = get_split(x_sample, y_sample)
print('Split: [X%d <= %.3f]' % ((split['index']+1), split['value']))
# ผลลัพธ์ที่ควรจะได้
#
# `Split: [X1 < 6.642]`
# ## 3.3 Build Tree
# ### Terminal Node
# Create a terminal node value
def to_terminal(outcomes):
return max(set(outcomes), key=outcomes.count)
to_terminal([0,1,1,1,1])
# ผลลัพธ์ที่ควรจะได้
#
# `1`
to_terminal([0,0,0,1,1])
# ผลลัพธ์ที่ควรจะได้
#
# `0`
# ### Recursive Splitting
# Create child splits for a node or make terminal
def split(node, max_depth, min_size, depth):
left_x, left_y, right_x, right_y = node['groups']
del(node['groups'])
# check for a no split
if not left_y or not right_y:
node['left'] = node['right'] = to_terminal(left_y + right_y)
return
# check for max depth
if depth >= max_depth:
node['left'], node['right'] = to_terminal(left_y), to_terminal(right_y)
return
# process left child
if len(left_y) <= min_size:
node['left'] = to_terminal(left_y)
else:
node['left'] = get_split(left_x, left_y)
split(node['left'], max_depth, min_size, depth+1)
# process right child
if len(right_y) <= min_size:
node['right'] = to_terminal(right_y)
else:
node['right'] = get_split(right_x, right_y)
split(node['right'], max_depth, min_size, depth+1)
# ### Put Everything Together
# Build a decision tree
def build_tree(x_train, y_train, max_depth, min_size):
root = get_split(x_train, y_train)
split(root, max_depth, min_size, 1)
return root
# Print a decision tree
def print_tree(node, depth=0):
if isinstance(node, dict):
print('%s[X%d <= %.3f]' % ((depth*' ', (node['index']), node['value'])))
print_tree(node['left'], depth+1)
print_tree(node['right'], depth+1)
else:
print('%s[%s]' % ((depth*' ', node)))
x_sample = [[2.771244718,1.784783929],
[1.728571309,1.169761413],
[3.678319846,2.81281357],
[3.961043357,2.61995032],
[2.999208922,2.209014212],
[7.497545867,3.162953546],
[9.00220326,3.339047188],
[7.444542326,0.476683375],
[10.12493903,3.234550982],
[6.642287351,3.319983761]]
y_sample = [0, 0, 0, 0, 0, 1, 1, 1, 1, 1]
tree = build_tree(x_sample, y_sample, 1, 1)
print_tree(tree)
# ผลลัพธ์ที่ควรจะได้
#
# ```
# [X1 < 6.642]
# [0]
# [1]
# ```
tree = build_tree(x_sample, y_sample, 2, 1)
print_tree(tree)
# ผลลัพธ์ที่ควรจะได้
#
# ```
# [X1 < 6.642]
# [X1 < 2.771]
# [0]
# [0]
# [X1 < 7.498]
# [1]
# [1]
# ```
tree = build_tree(x_sample, y_sample, 3, 1)
print_tree(tree)
# ผลลัพธ์ที่ควรจะได้
#
# ```
# [X1 < 6.642]
# [X1 < 2.771]
# [0]
# [X1 < 2.771]
# [0]
# [0]
# [X1 < 7.498]
# [X1 < 7.445]
# [1]
# [1]
# [X1 < 7.498]
# [1]
# [1]
# ```
# ### 3.4 Prediction
# Make a prediction with a decision tree
def predict(node, row):
if row[node['index']] < node['value']:
if isinstance(node['left'], dict):
return predict(node['left'], row)
else:
return node['left']
else:
if isinstance(node['right'], dict):
return predict(node['right'], row)
else:
return node['right']
# +
tree = build_tree(x_sample, y_sample, 1, 1)
for x, y in zip(x_sample, y_sample):
prediction = predict(tree, x)
print('Expected=%d, Got=%d' % (y, prediction))
# -
# ผลลัพธ์ที่ควรจะได้
#
# ```
# Expected=0, Got=0
# Expected=0, Got=0
# Expected=0, Got=0
# Expected=0, Got=0
# Expected=0, Got=0
# Expected=1, Got=1
# Expected=1, Got=1
# Expected=1, Got=1
# Expected=1, Got=1
# Expected=1, Got=1
# ```
# ([กลับขึ้นข้างบน](#top))
#
# ----
# <a id='section_4'></a>
# # 4. การประเมินความแม่นยำของโมเดล
y_train = train_set.iloc[:,-1].tolist()
x_train = train_set.iloc[:,:-1].values.tolist()
max_depth = 2
min_size = 10
tree = build_tree(x_train, y_train, max_depth, min_size)
y_test = test_set.iloc[:,-1].tolist()
x_test = test_set.iloc[:,:-1].values.tolist()
y_pred = []
for x in x_test:
y_pred.append(predict(tree, x))
assert(len(y_pred) == len(y_test))
# ## 4.1 Accuracy
def accuracy_metric(actual, predicted):
correct = 0
for i in range(len(actual)):
if actual[i] == predicted[i]:
correct += 1
return correct / float(len(actual)) * 100.0
accuracy_metric(y_test, y_pred)
# ## 4.2 Confusion Matrix
def confusion_matrix(actual, predicted):
y_g = set(actual)
y_p = set(predicted)
mat = [[0]*len(y_p) for i in range(len(y_g))]
for i, yg in enumerate(y_g):
for j, yp in enumerate(y_p):
for k in range(len(actual)):
if actual[k] == yg and predicted[k] == yp:
mat[i][j] += 1
return mat
def print_confusion_matrix(matrix):
print(f' Prediction')
print(f'Reference {0:5d} {1:5d}')
for i, row in enumerate(matrix):
print(f' {i:3d}', end=' ')
for num in row:
print(f'{num:5.0f}', end=' ')
print('')
mat = confusion_matrix(y_test, y_pred)
print_confusion_matrix(mat)
# ## 4.3 Precision and Recall
def precision_recall(actual, predicted):
tp = 0
for y_g, y_p in zip(actual, predicted):
tp += (y_g == 1) & (y_p == 1)
fp = 0
for y_g, y_p in zip(actual, predicted):
fp += (y_g == 0) & (y_p == 1)
fn = 0
for y_g, y_p in zip(actual, predicted):
fn += (y_g == 1) & (y_p == 0)
tn = 0
for y_g, y_p in zip(actual, predicted):
tn += (y_g == 0) & (y_p == 0)
precision = tp / float(tp + fp)
recall = tp / float(tp + fn)
return precision, recall
p, r = precision_recall(y_test, y_pred)
print(f'precision = {p:.3f}')
print(f'recall = {r:.3f}')
# ## 4.4 F1-score
def f1score(actual, predicted):
p, r = precision_recall(actual, predicted)
return (2*p*r) / (p+r)
f1 = f1score(y_test, y_pred)
print(f'F1-scre = {f1:.3f}')
# ([กลับขึ้นข้างบน](#top))
#
# ----
# # ใช้ Scikit-Learn
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier(max_depth=max_depth, min_samples_leaf=min_size)
clf = clf.fit(x_train, y_train)
import sklearn
import matplotlib.pyplot as plt
plt.figure(figsize=(10,6))
sklearn.tree.plot_tree(clf);
# เทียบกับของเดิม
print_tree(tree)
accuracy_metric(y_test, y_pred)
y_pred_sklearn = clf.predict(x_test)
accuracy_metric(y_test, y_pred_sklearn)
|
decision_tree_ex_sol.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="kJUj1SzWZUSD"
# <font color=gray>This Jupyter notebook was created by <NAME> for \the\world Girls' Machine Learning Day Camp. The license can be found at the bottom of the notebook.</font>
#
# # Section 1: Introduction to Jupyter and Output
#
# ## Introduction to Jupyter
#
# For this introduction to programming, we're using something called a "Jupyter Notebook" and we'll be programming in a language called "python." The Jupyter notebook allows us to include both text cells (called "markdown") and code cells. This text is all contained in a markdown cell, and the cell below this is a code cell (don't worry about the actual code for now, we'll learn about that later!). If you click the code cell and then type shift + enter, it will run the code in the cell. Try it out!
# + colab={} colab_type="code" id="BUqcz9pjZUSG"
coolness_factor = 10
if coolness_factor > 5:
print("The code ran, this is super awesome!")
else:
print("Sad face. We didn't make it.")
# + [markdown] colab_type="text" id="T0yDIhQDZUSO"
# Notice that the output from this code appears below the code cell. You can edit the text in both the text cells and the coding cells. For the text cells, double click, make your changes, and then type shift + enter to revert to the cleaner formatting. Feel free to do this to add your own notes, but please don't delete the instructions, they're meant to help you! You'll frequently edit the code cells to write your own code.
#
# For this introduction, we'll often act as though the code cells were completely independent blocks of code. However, in reality, the computer is still keeping track of any code that you ran previously in this notebook. If you would like to erase the computer's memory of everything you've run so far, you can click the button to "restart the kernel." Old output will still be displayed, even after you restart the kernel. If you'd like to erase all output, this is an option under "Cell."
#
# If you make a mistake while coding and the program doesn't finish quickly, you might want to manually stop it. There is a stop button near the top of the page, which will terminate the program. Try running the following code (which will run forever if you don't make it stop), and use the stop button to end it. Then restart the kernel, so it's (almost) as though it never happened. You can erase the output if you'd like.
# + colab={} colab_type="code" id="rjaW8hSuZUSQ"
while True:
print("this is the song that never ends")
# + [markdown] colab_type="text" id="IiFjshPGZUSV"
# Now, we're ready to start on the fun stuff: actually understanding and writing code.
#
# ## Output
#
# In programming, it can be very useful to print messages. Run the following code to print the message "hello world" to the screen.
# + colab={} colab_type="code" id="7s76JcAzZUSW"
print("hello world")
# + [markdown] colab_type="text" id="0yxDru0BZUSZ"
# Notice that the text inside of the quotation marks gets printed exactly as written - try this out by inserting your own message in the quotation marks!
# + colab={} colab_type="code" id="IRQQ-uLuZUSa"
print("WRITE YOUR MESSAGE HERE")
# + [markdown] colab_type="text" id="gQ5JqYw1ZUSd"
# If we have multiple print statements on different lines, the text output will be split between different lines.
# + colab={} colab_type="code" id="Z987EUugZUSe"
print("This is the 1st line.")
print("This is the 2nd line.")
print("y9q82!!!!")
# + [markdown] colab_type="text" id="JvSI7se-ZUSg"
# ## Exercises
# + [markdown] colab_type="text" id="8laZgNbaZUSg"
# Write code that produces the following output:
#
# `I like turtles` <br>
# `Yes I do!` <br>
# `I like turtles` <br>
# `How about you?` <br>
# + colab={} colab_type="code" id="bV2f7gmhZUSh"
# + [markdown] colab_type="text" id="hOth_Mt9ZUSj"
# For each line of code, try to run it to see what happens. Then find and correct the error.
# + colab={} colab_type="code" id="Z6ckyDvzZUSk"
print(What kind of pictures does a turtle take?)
# + colab={} colab_type="code" id="VtqKxFPaZUSm"
prnit("Shellfies!")
# + colab={} colab_type="code" id="2-vX-AdyZUSo"
print("What happens when you bring a turtle to a party?"):
# + colab={} colab_type="code" id="upK2fkooZUSq"
print["It becomes a shellebration!"]
# + [markdown] colab_type="text" id="zxCYCPxnZUSr"
# <font color=gray>Copyright (c) 2018 <NAME></font>
# <br><br>
# <font color=gray>Permission is hereby granted, free of charge, to any person obtaining a copy
# of this software and associated documentation files (the "Software"), to deal
# in the Software without restriction, including without limitation the rights
# to use, copy, modify, merge, publish, distribute, sublicense, and/or sell
# copies of the Software, and to permit persons to whom the Software is
# furnished to do so, subject to the following conditions:</font>
# <br><br>
# <font color=gray>The above copyright notice and this permission notice shall be included in all
# copies or substantial portions of the Software.</font>
# <br><br>
# <font color=gray>THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE
# AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM,
# OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER DEALINGS IN THE
# SOFTWARE.</font>
|
01-Monday/1 - Intro to Jupyter and Output.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit
# name: python38164bitf714261400ea47f6845c08cb7a4bef65
# ---
N = 2
# +
from random import shuffle
from itertools import product
class Board:
lambdas = {
# (x, y)
'U': lambda pos: (pos[0], pos[1] + 1),
'D': lambda pos: (pos[0], pos[1] - 1),
'L': lambda pos: (pos[0] + 1, pos[1]),
'R': lambda pos: (pos[0] - 1, pos[1]),
}
def __init__(self, matrix: list, wpos: tuple):
self.matrix = matrix
self.wpos = wpos
@staticmethod
def generate_board(N):
w_pos = None
flat_board = ['W'] + ['R'] * ((N**2 - 1) // 2) + ['B'] * (N**2 // 2)
shuffle(flat_board)
board = [flat_board[i: i + N] for i in range(0, len(flat_board), N)]
for y, x in product(range(N), repeat=2):
if board[y][x] == 'W':
w_pos = (x, y)
break
return Board(board, w_pos)
def compare_with(self, other):
part = 1 / (len(self.matrix) ** 2)
equality = 0
for y, x in product(range(len(self.matrix)), repeat=2):
if self.matrix[y][x] == other.matrix[y][x]:
equality += part
return equality
def move(self, movement):
x, y = self.wpos
tox, toy = self.wpos = self.lambdas[movement](self.wpos)
self.matrix[y][x] = self.matrix[toy][tox]
self.matrix[toy][tox] = 'W'
return self
def copy(self):
return Board(list(self.matrix), self.wpos)
def __repr__(self):
return '\n'.join(map(lambda b: ''.join(map(str, b)), self.matrix))
# -
class Node:
def __init__(self, board):
self.children = {}
self.board = board
self.data = None
self.parent = None
def fill_childs(self):
x, y = self.board.wpos
if x > 0:
self.children['R'] = Node(self.board.copy().move('R'))
if x < len(self.board.matrix) - 1:
self.children['L'] = Node(self.board.copy().move('L'))
if y > 0:
self.children['D'] = Node(self.board.copy().move('D'))
if y < len(self.board.matrix) - 1:
self.children['U'] = Node(self.board.copy().move('U'))
# +
initial = Board.generate_board(N)
target = Board.generate_board(N)
initial.compare_with(target)
print(initial)
print()
print(target)
# +
import numpy as np
from IPython.display import clear_output
from time import sleep
max_steps = 100
steps = 0
my_tree = Node(initial)
my_tree.fill_childs()
print(my_tree.children)
while True:
break
if steps > max_steps:
break
for child in my_tree.items():
pass
if equality(initial, target) == 1:
break
clear_output(wait=True)
print(np.matrix(initial))
sleep(0.5)
steps += 1
|
hackerrank/Solutions/euler_244.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
## https://github.com/sourcethemes/academic-admin
# -
# cd "/Users/erickaxxe/Documents/GitHub/famelo"
# !pip3 install -U git+https://github.com/wowchemy/hugo-academic-cli.git
# !academic import --bibtex cite.bib
|
BibTeX_Axxe_20200225.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 4: Introduction to Object-Oriented Programming
#
# We used Python so far for relatively simple scripting with the addition of an occasional function. Generally, we have been following what is called a **procedural programming** style: we define functions that combine several steps into a procedure that can be called and executed. Basically, the program execution follows a "top-down" approach: from defining some variables, we execute procedures (functions) that operate on these variables, collect the output, perform some other procedures, etc.
#
# This type of programming style is very common in scientific programming and there is nothing wrong with it: in fact, it is used successfully in highly complex scientific simulations. An advantage of this way of programming is that it usually follows a logical structure and is quite intuitive to follow.
#
# There is, however, another programming style that you have surely heard of: **object-oriented programming** (often abbreviated as OOP). Whereas in procedural programming we define variables and break down parts of codes in common functions/ procedures (often with the aim to ensure as much code re-use as possible), those two parts are (mostly) combined into single objects in OOP.
#
# If you think that this sounds strange then you are not alone - at first, OOP seems to be a lot of work without a clear benefit. However, once you get used to it, OOP can lead to very efficient code re-use and clear code structures.
#
# Most importantly, once you have gone through the hard work of defining classes, then using the objects in the program becomes the easier part.
#
# In this notebook, we'll have a look at some of the basic concepts of OOP in Python and some examples to give you a feel for how it works - and to see some of the advantages that you can get out of this programming style.
#
#
#
from IPython.core.display import HTML
css_file = 'nre2_style.css'
HTML(open(css_file, "r").read())
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# ## Defining a grid for numerical simulations
#
# We'll start straight away with the definition of a type of object that might actually be useful in the following parts of this course: the definition of a grid for numerical simulations.
#
# The simplest grid that can be used for numerical simulations is a completely regular grid with cell sizes that do not change along an axis direction:
# +
fig = plt.figure(figsize=(7,5))
# Set RC (Run & Configure, i.e. default) parameters for matplotlib
plt.rcParams['font.size'] = 16
ax1 = fig.add_subplot(111)
for i in range(10):
ax1.axvline(i, c='k')
ax1.axhline(i, c='k')
ax1.set_title("Example of a regular mesh")
plt.tight_layout()
# -
# Note that the cell width (let's call it $dx$) and hight ($dy$) can be different - but they are constant for all cells.
#
# Which kind of information do we need to represent this grid? As minimal requirements, we need:
#
# - the cell dimensions $dx$ and $dy$
# - the number of cells in each direction, $nx$, $ny$
# - some way to store the data of the cell values
#
# In this notebook, we will use both a procedural and an object-oriented approach to deal with the data in this grid to clarify the differences between (and advantages of) the approaches.
#
# First, we will define the grid properties that we need - and generate some random grid values to work with:
#
dx = 100
dy = 100
nx = 10
ny = 10
grid_data = np.random.randint(0,10,size= (nx, ny))
# We generated some random grid data with the command `np.random.randint`, in this case: random integer values between 0 and 10, stored in a 2-D array of size (nx, ny). Here just a quick view of the generated random grid:
plt.imshow(grid_data, origin = 'lower', interpolation = 'nearest', cmap='viridis')
# ## The procedural way: storing data in a dictionary
#
# We will first look at the procedural implementation of this grid data. Of course, we could simply work with the variables that we already defined above and use functions that directly work with these variables - but we would have to pass a lot of variables around. Also, when we define a second grid (as we will do in a second), we'd have to have a new name for all variables (e.g. `nx_grid_2`, `dx_grid_2`, etc.).
#
# One possibility to combine several variables for simpler procedural programming applications is to store the information in a Python dictionary:
grid_dict = {'dx' : dx,
'dy' : dy,
'nx' : nx,
'ny' : ny,
'grid_data' : grid_data}
# We can now have a look at all the data in the grid and, of course, access the single variables with the common dictionary access methods (e.g. `grid_dict['nx']` to get the nx values):
print(grid_dict)
print("Number of grid cells in x-direction: %d" % grid_dict['nx'])
# ## The OOP-way: creating a class for numerical grids
#
# Before we start looking at more complex object-oriented programming concepts, we will first simply replicate the storage of the grid information as above, but this time within a defined grid class.
#
# The absolutely simplest way to create a class is not so different to a dictionary:
#
#
class Grid():
pass
# The previous line is the blueprint for a class that, basically, does absolutely nothing (if you want Python to do nothing, you simply write `pass`)...
#
# However, we can now create an instance (object) of the class and then add the information that we want to store:
# create an instance of the grid
grid_obj = Grid()
# now: store information as grid attributes:
grid_obj.nx = nx
grid_obj.ny = ny
grid_obj.dx = dx
grid_obj.dy = dy
grid_obj.grid_data = grid_data
# We can now access the grid information. Here the first difference: if we simply use a `print` statement as above, we get information *about the object class* - and not about the data:
print(grid_obj)
# To get the data, we can use the '.'-operator to access the object attributes:
print("Number of grid cells in x-direction: %d" % grid_obj.nx)
# And we can actually get all variables back as a dictionary - and get a very similar printout to before:
print (grid_obj.__dict__)
# Ok, what we have done now is, basically, very similar to using a dictionary above: we simply have a "container" to store the relevant information of the grid. However, we will make this container a lot more sophisticated while we go along - and add features that are specific to object-oriented programming.
# ## Adding class methods to access data
#
# The first difference that we will now look at is typical (but not essential!) for object-oriented programming: instead of simply adding variables as before with the '.'-operator, we write class methods (functions) to store and access the data.
#
# I am sure that you'll wonder about what exactly this should be good for. Basically, there are two reasons:
#
# 1. We can "hide" the actual place where (and how) we store information from the user - mostly to make the access clearer and easier (this is the OOP concept of "encapsulation")
# 2. It is possible to directly react to the supplied data, e.g. to check for consistency, to react to different data types, etc.
#
# Let's start with the description of class methods to add the variables:
# The keyword "class" is used to start the class definition, followed by the name:
class Grid():
# we now define the class methods (i.e. the functions)
def set_grid(self, grid_data):
self.grid_data = grid_data
def set_nxny(self, nx, ny):
self.nx = nx
self.ny = ny
def set_dxdy(self, dx, dy):
self.dx = dx
self.dy = dy
# We now call these methods to assign data to the object:
# re-create the object to consider the new methods:
grid_obj = Grid()
# now, add data:
grid_obj.set_nxny(nx, ny)
grid_obj.set_dxdy(dx, dy)
grid_obj.set_grid(grid_data)
# Here is the first advantage of using this approach to store data: we can include simple checks of the data type. There are several ways to do this. What we will use below is already a quite advanced way of dealing with errors: "exception handling". If some condition is not met, we "raise" an error - an this causes the execution to abort:
#
# raise AttributeError("Some error description")
#
# This is an advanced topic and you don't have to understand the details now - we will see below what happens:
class Grid():
# we now define the class methods (i.e. the functions)
def set_grid(self,grid_data):
# before we store the data, let's check if it is actually a 2-D array.
# 1) check that the object is a numpy array:
if type(grid_data) != np.ndarray:
raise AttributeError("The provided grid data is of the wrong type!")
# 2) check that it is of dimension 2:
if grid_data.ndim != 2:
raise AttributeError("The numpy array is not of dimension 2!")
self.grid_data = grid_data
def set_nxny(self, nx, ny):
self.nx = nx
self.ny = ny
def set_dxdy(self, dx, dy):
self.dx = dx
self.dy = dy
# If we now pass the wrong type or the wrong dimension, then (a) the object will not be created, and (b) we get some information on the problem:
# wrong object
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
grid_obj.set_grid("a") # grid_data)
grid_obj.set_dxdy(dx, dy)
# wrong dimension
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
grid_obj.set_grid(np.array([1,2,3])) # grid_data)
grid_obj.set_dxdy(dx, dy)
# So what we get with these methods is now an automatic testing of the variables that are stored in our object - with the same object method that we use to add the data.
#
# Of course, we could replicate this behaviour with a procedural equivalent: we could write a function that adds the data to the dictionary and does these checks, e.g.:
def add_grid_to_dict(grid_dict, grid_data):
"""Add grid data to grid dictionary"""
# now: perform checks, exactly the same code as above:
# 1) check that the object is a numpy array:
if type(grid_data) != np.ndarray:
raise AttributeError("The provided grid data is of the wrong type!")
# 2) check that it is of dimension 2:
if grid_data.ndim != 2:
raise AttributeError("The numpy array is not of dimension 2!")
# if no errors were raised, add the data:
grid_dict['grid_data'] = grid_data
# wrong object
add_grid_to_dict(grid_dict, "a")
# wrong dimension
add_grid_to_dict(grid_dict, np.ndarray([1,2,3]))
# So: we get a similar possibility to perform checks: both with the procedural style of writing a function, as with the object-oriented style. The main difference at the moment is that we have to execute a separate function for the procedural style (`add_grid_to_dict`), whereas the object oriented method performs the step automatically.
#
# We will now look at a slightly different implementation to highlight the difference of the approaches.
#
# ## Encapsulation and private data in OOP
#
# Let's assume that we would like to force the tests for consistency (for data type and grid dimension) to be executed. In the case of the dictionary implementation, nobody forces you to use the `add_grid_to_dict` function that performs the tests. You can always just add the grid yourself to the dictionary (and this is actually the simpler approach - as you don't have to remember the name of the function, so: people *will* do it!).
#
# For the OOP implementation, we can - at least in principle - strongly suggest that the tests are executed when the data is added. This approach is based on encapsulation.
#
# The general idea in OOP encapsulation is that, as a convention, variables and data are added and extracted from an object with access methods (for example our `set_nxny()` function above, etc.) and not accessed "in place".
#
# For example: we could, theoretically, set the `nx` and `ny` values of our grid class directly:
#
#
grid_obj = Grid()
grid_obj.nx = nx
grid_obj.ny = ny
print(dir(grid_obj))
# Now, in Python there is a way to store data and mark them as "protected": an **underscore** (or double underscore) is **added as a prefix** to the class variable name, i.e. instead of:
#
#
# def set_nxny(self, nx, ny):
# self.nx = nx
# self.ny = ny
#
# we can write:
#
#
# def set_nxny(self, nx, ny):
# self._nx = nx
# self._ny = ny
#
# Using these variables is a basic Python **convention: do not change variables with a leading underscore** outside the class definition!
#
#
# Here an example implementation:
#
# +
# Adjusted grid class
class Grid():
# we now define the class methods (i.e. the functions)
def set_grid(self,grid_data):
# before we store the data, let's check if it is actually a 2-D array.
# 1) check that the object is a numpy array:
if type(grid_data) != numpy.ndarray:
raise AttributeError("The provided grid data is of the wrong type!")
# 2) check that it is of dimension 2:
if grid_data.ndim != 2:
raise AttributeError("The numpy array is not of dimension 2!")
self._grid_data = grid_data
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
# -
#
# Note that, *in principle*, you could still access and change the variable `grid_obj._nx' directly:
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
print(grid_obj.__dict__)
grid_obj._nx = 2
print(grid_obj.__dict__)
# **BUT**: the leading underscore of a class variable tells you that *you should not change it directly* - but rather use the appropriate functions (e.g. the `grid_obj.set_nxny()` function).
#
# If you still change it: *your responsibility* if things do not work out as intended (e.g. in our case: the test for grid consistency would be skipped).
#
# The reason why Python does not enforce complete privacy (as, for example, C++ and Java allow) has to do with the general culture of open-source programs in Python (as opposed to C++): basically, as you have access to the source code anyway, there is no real purpose for fully hiding information. If you are interested, you can find many discussions on this topic on the net, for example here:
#
# http://stackoverflow.com/questions/1641219/does-python-have-private-variables-in-classes
#
# Following these considerations, we could extend our grid class with methods to get the (`nx, ny`), (`dx, dy`) and grid values (in addition to setting them), and store the actual values in "private" variables:
class Grid():
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
def get_dxdy(self):
return (self._dx, self._dy)
# +
# and here a test:
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
grid_obj.set_dxdy(dx, dy)
grid_obj.set_grid(grid_data)
# now: access the information:
print(grid_obj.get_nxny())
print(grid_obj.get_dxdy())
print(grid_obj.get_grid())
# -
print(dir(grid_obj))
# A last note on encapsulation: all this "setting" and "getting" data seems extremely cumbersome - and it is, for small scripts or implementations that are only for a limited purpose. In this case, you don't need to use these concepts.
#
# However, once you create classes that you will use for longer - or that you even might pass on to other people - using these ideas becomes very powerful. Most importantly, it adds a great flexibility to the way your object is used: it allows you, for example to:
#
# - perform tests along the way (as we do for grid dimensions)
# - change the way the data is stored and accessed (for example in a different type of object - if you discover a better method)
# - add features to your code.
#
# And, here is the main point: you can do all of these things while making sure that **previously written code** (by you - or someone else) **will still work!** Ensuring this type of "backwards compatibility" is very difficult to achieve with standard procedural programming methods.
#
# </p>
#
#
# ## Adding more features to the grid class
#
# After the theoretical considerations of encapsulation, we are not getting back to actually doing something interesting with our grid class. First, we will add some more features that could be useful. First of all, we will add a function to calculate the grid extent.
#
# We have both the number of cells as well as the cell width (`dx, dy`) given in each direction, so we can simply calculate the grid extent. Actually, we would like to update the extent every time when either the number of cells `nx, ny` or the cell dimensions `dx, dy` are changed. We therefore include the following steps:
#
# 1. Create a method to calculate the extent (`self.update_extent()`)
# 2. Execute this function when either `self.set_nxny` or `self.set_dxdy` are called
# 3. Include a `get_extent()` function to access the information:
class Grid():
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# Here an example:
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
grid_obj.set_dxdy(dx, dy)
# now: get information
(extent_x, extent_y) = grid_obj.get_extent()
print("Extent in x-direction: %.1f, y-direction: %.1f" % (extent_x, extent_y))
# Of course, a similar behaviour could be replicated with the dictionary implementation in a purely procedural programming fashion - but an additional function would have to be called each time to calculate (or update) the extent. In the OOP implementation, this is done automatically!
#
# ## OOP-specific features (1): enabling expected behaviour
#
# So, after looking at implementations that could also be achieved with standard procedural methods (although with the requirement to call additional functions, etc.), we will now have a look at useful features that are only possible with an OOP implementation: concepts based on function overloading and defined object namespaces.
#
# <hr>
#
# Here is the **basic idea**: a good class definition should create an object that performs or reacts in an *intuitive* way.
#
# <hr>
#
# What does this mean? Basically, it means that the object should - if possible - have functions that work in a way that a user would expect - without having to read through the source code, etc. And these functions should perform actions that make sense in the context of the object and its variables.
#
# We will first look at standard functions to explain what is meant with this concept of "intuitive behaviour".
#
# We have seen this behaviour several times before. Here one example: if we create a numeric variables, for example a float, and we double it, then we actually obtain a variable with the doubled value. If we create a string and double it, then we obtain the string back twice:
numeric_val = 10.
print (2 * numeric_val)
string_val = "hello"
print (2 * string_val)
# As trivial as this may seem - it encapsulates exactly what was described above: the object (either a float or a string value) "knows" when you type
#
# 2 *
#
# in front of it - and acts accordingly. The same happens, for example, if you type the `print` statement. For `numeric_val` and `string_val` the behaviour is, again, quite trivial. But have a look what you get when you print a numpy array variable:
print(numeric_val)
print(string_val)
# now: let's look at a numpy array:
array_val = np.arange(1,10)
print(array_val)
# Again, maybe not too impressive - but look what happens if you increase the size of the array, for example:
array_val = np.arange(1,100000)
print(array_val)
# We actually observe two specific behaviours that are implemented here:
#
# 1. The numpy array object knows that you want to know the array values when you type `print` - so this is what you get back
# 2. The object checks the size of the array - and returns the appropriate type of information - instead of simply dumping all values out on the screen which would lead to a pretty unreadable output!
#
# The type of behaviour for an object with respect to the `print` function can be implemented in an own class definition, as well. First, let's look what we get if we simply type a `print` in front of an object that we created before:
print(grid_obj)
# We get back the information that this is an instance of the class grid - *not* very useful. Let's change that behaviour. First, we have to think what we would intuitively expect as information from the grid object. What I would consider useful:
#
# - Information what it actually is (a 2-D grid object)
# - range, extent, spacing of the grid
# - some information on the stored data.
#
# The way we tell the grid class that we would like to get this information when it is used in a `print` statement is that we create a special class function, the
# `__repr()__`-function (for representation). Here an example:
#
#
class Grid():
# function that provides information when used with print statement:
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# Let's test it:
grid_obj = Grid()
grid_obj.set_nxny(nx, ny)
grid_obj.set_dxdy(dx, dy)
grid_obj.set_grid(grid_data)
# now: access all the information with a simple "print" statement:
print(grid_obj)
# The type of information that you would like to show might be different in your case - but the essential point here is that you don't have to supply an extra function to the user that, for example, provides information about the data in a dictionary, but simply use the standard `print` statement that everyone knows and would intuitively use as a first try to get information on an object.
#
# Another advantage: the type of information that you might like to present might change over time. For example, we could add information about the units (would make sense, anyway) of the dimension and extent ([m], [km], etc.) and the data (Temperature, pressure, ...). You could easily add this information with an update of the class definition - and previously created code would still work!
#
# ## OOP-specific features (2): Object initialisation
#
# You might have realised that we always use the same steps to create an object: we first create an empty object, then add (nx,ny), (dx,dy) and then the data. This is actually quite annoying and repetitive.
#
# Class definitions provide a method to combine all of these steps directly in the object generation call. We have to define a special object method, called `__init__`, where we define how to consider this information automatically.
#
# Here is what this can look like:
class Grid():
# define the object initialisation function
def __init__(self, **kwds):
if 'nx' in kwds and 'ny' in kwds:
self.set_nxny(kwds['nx'], kwds['ny'])
if 'dx' in kwds and 'dy' in kwds:
self.set_dxdy(kwds['dx'], kwds['dy'])
if 'grid_data' in kwds:
self.set_grid(kwds['grid_data'])
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self, grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# Now, we can directly create an object with the grid information and the grid data:
grid_obj = Grid(nx = 10, ny = 10,
dx = 100., dy = 100.,
grid_data = np.random.randint(1,10,size=(nx,ny)))
print(grid_obj)
# There are many more ways to initialise objects - we will leave it here for now,
# ## OOP-sepcific features (3): Operator overloading of +,-,\*,/
#
# Getting information easily and intuitively is useful - but there are some much more useful possibilities: we can define the behaviour of an object when it is accessed with the standard arithmetic operators! This is now where object-oriented methods are becoming extremely powerful.
#
# The idea is now similar to before: we want to implement a behaviour of a grid object that is *intuitive* when we access it with a standard arithmetic operator.
#
#
#
class Grid():
# operator overloading
def __add__(self, rhs):
self._grid_data = self._grid_data + rhs
# now we return the updated object entirely:
return self
def __init__(self, **kwds):
if 'nx' in kwds and 'ny' in kwds:
self.set_nxny(kwds['nx'], kwds['ny'])
if 'dx' in kwds and 'dy' in kwds:
self.set_dxdy(kwds['dx'], kwds['dy'])
if 'grid_data' in kwds:
self.set_grid(kwds['grid_data'])
# function that provides information when used with print statement
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# +
grid_obj = Grid(nx = nx, ny = ny, dx = dx, dy = dy, grid_data = grid_data)
# Let's have a look at our grid data:
print(grid_obj)
# now: add a scalar value to the grid:
grid_obj_new = grid_obj + 10
print(grid_obj_new)
# -
# We can see from the min/max grid values that the scalar value was correctly added to the grid values!
#
# Here another option: we can check which type of object is passed after the standard "+"-operator and react accordingly. For example, we might want to add the values of two grids together. How we can do this:
#
# 1. Check if the passed object is of type "Grid"
# 2. Create a copy of the existing object (stored in the class "self" variable in the class definition)
# 3. Perform the addition operation and return the new grid:
#
# library copy is required to create a full copy of the object itself
import copy
class Grid():
# operator overloading
def __add__(self, rhs):
# (1) Check the type of the "rhs" variable
if isinstance(rhs, Grid):
# (2) create a copy of the
new_grid = copy.deepcopy(self)
# (3) perform addition of grid data
new_grid._grid_data = self._grid_data + rhs._grid_data
else:
new_grid = copy.deepcopy(self)
new_grid._grid_data = self._grid_data + rhs
# now we return the updated object entirely:
return new_grid
def __init__(self, **kwds):
if 'nx' in kwds and 'ny' in kwds:
self.set_nxny(kwds['nx'], kwds['ny'])
if 'dx' in kwds and 'dy' in kwds:
self.set_dxdy(kwds['dx'], kwds['dy'])
if 'grid_data' in kwds:
self.set_grid(kwds['grid_data'])
# function that provides information when used with print statement
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# Now, we can simply create a new grid with the added values:
# +
g1 = Grid(nx = nx, ny = ny, dx = dx, dy = dy,
grid_data = np.random.randint(1,10,size=(10,10)))
g2 = Grid(nx = nx, ny = ny, dx = dx, dy = dy,
grid_data = np.random.randint(1,10,size=(10,10)))
# and now: this is all we have to do to create a new grid with added values:
g_combined = g1 + g2
print("\n\tNew grid:\n")
print(g_combined)
print("\n\tOriginal grid:\n")
print(g1)
# -
# Now we get what we expected: we get back a new grid with the values of the two other grids added together.
#
# More ideas to make the "+" operator even more powerful:
#
# 1. We could first check if the grids are actually defined on the same range (same number of cells, dimensions, etc.);
# 2. We could define the addition with a simple numpy 2-D array: if this type of array is passed, then add it to the grid data;
#
# Here some more Python operators that can be overloaded in a similar way:
#
# Subtract values:
#
# __sub__(self, other)
#
# Multiply:
#
# __mul__(self, other)
#
# Divide:
#
# __div__(self, other)
#
# Power:
#
# __pow__(self, other)
#
# Technical detail: these previous operators are called "regular binary operators" they perform a binary operation for the object itself and another object *after* the operator. If you want to obtain an operator with the reversed order, i.e.: first the *other* object, then operator, then the own object (for example: `2 + g1`), you can define "reversed binary operator". For the addition, this is:
#
# __radd__(self, other)
#
# There are many more operators and standard actions that can be defined for your own class! Here a list: http://www.siafoo.net/article/57
#
# A note on the comparison to the procedural approach:
#
# Again - you could replicate a similar behaviour with a purely procedural approach and write a function, maybe called (`add_grids()`) that adds the grid values of two grid-dictionaries. However, you would have to:
#
# 1. Remember the name of the function to perform the operation;
# 2. Write one separate function (and remember its name!) for each type of operation
#
# Whereas, in the case of object overloading, you can intuitively "guess" that the addition operator performs a grid operation and that other operators (-, \*, / etc) work accordingly.
#
# ## Intuitive functions in object namespace
#
# Apart from overloading common operators, we can, of course, define our own functions to perform objet-specific operations that go beyond the simple "get" and "set" methods that we used so far.
#
# For example, for our grid we might want to define functions to create plot of the grid with imshow. You know quite well by now that you have to combine a couple of commands to create the image:
#
# 1. open a figure,
# 2. plot the data,
# 3. adjust extent,
# 4. and, of course: add labels and a title.
#
# We can combine all these steps into one object function and give it an intuitive name - for example `plot()`. The relevant information can be obtained from the grid variables:
#
#
import matplotlib.pyplot as plt
import copy
class Grid():
# include the plot function here:
def plot(self):
# (1) open figure, add axis
plt.figure(figsize=(8,8))
# x = fig.add_subplot(111)
# (3+4) create plot with automatically adjusted extent and some other
# useful settings
extent = [0., self._extent_x, 0., self._extent_y]
im = plt.imshow(self._grid_data, extent = extent,
interpolation = 'nearest', cmap = 'viridis')
cbar = plt.colorbar(im, orientation = 'horizontal', fraction=0.042, pad=0.14)
cbar.set_label("Grid data")
plt.xlabel("X")
plt.ylabel("Y")
plt.title("2D grid")
plt.show()
# operator overloading
def __add__(self, rhs):
# (1) Check the type of the "rhs" variable
if isinstance(rhs, Grid):
# (2) create a copy of the
new_grid = copy.deepcopy(self)
# (3) perform addition of grid data
new_grid._grid_data = self._grid_data + rhs._grid_data
else:
new_grid = copy.deepcopy(self)
new_grid._grid_data = self._grid_data + rhs
# now we return the updated object entirely:
return new_grid
def __init__(self, **kwds):
if 'nx' in kwds and 'ny' in kwds:
self.set_nxny(kwds['nx'], kwds['ny'])
if 'dx' in kwds and 'dy' in kwds:
self.set_dxdy(kwds['dx'], kwds['dy'])
if 'grid_data' in kwds:
self.set_grid(kwds['grid_data'])
# function that provides information when used with print statement
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self, grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# +
# define a grid with random values and create plot
nx = 10
ny = 10
dx = 100
dy = 100
g1 = Grid(nx = nx, ny = ny,
dx = dx, dy = dy,
grid_data = np.random.randint(1,10,size=(nx,ny)))
# create the plot
g1.plot()
# -
g3 = Grid()
print (g3.__dict__)
print(dir(Grid()))
# The definition of this plot function within the object is not only useful because it combines a lot of commands into one function (we could do the same, of course, for a standard procedural approach with one function!), but because we gave it a very simple, intuitive and easy to remember name: `plot`!
#
# Here a bit of more thought on this point: the function `plot` is defined as an object function - so it is available only to the object and strictly defined in its "namespace". This means that we can, without causing any problem, have another object that *also* has a function `plot()` - for example: think about a class definition for a line plot, maybe for a gamma-ray borehole log: an object of this class can have a `plot()` method as well - which creates an appropriate plot for this type of data!
# ## Exercise: extend the grid class
#
# Of course, it would be nice if we would get a proper name for the grid (in the title) and for the meaning of the grid data in the colorbar label. Here a little exercise for you:
#
# Extend the Grid class definition:
#
# 1. Add a function to provide a name for the grid (as title) and store it in a local variable (e.g. "Temperature distribution")
# 2. Add a function to provide a name for the data type and unit and store it in variables (e.g. type: Temperature, unit: C)
# 3. Extend the '__init__()' function to optionally set these variables during object generation (like dx, dy etc.)
# 4. Extend the `plot()` function to set plot title and colorbar label accordingly - if these names are defined!
#
# I included the template for you - now add the required features:
#
import matplotlib.pyplot as plt
import copy
class Grid():
#
# To Do (1): include appropriate checks for keywords and set variables, if provided:
#
# Add the keywords as:
# grid_title : the title of the grid (e.g. "Temperature distrubution")
# var_name : name of data type (e.g. "Temperature")
# var_unit : unit of data (e.g. "C")
def __init__(self, **kwargs):
if 'nx' in kwargs and 'ny' in kwargs:
self.set_nxny(kwargs['nx'], kwargs['ny'])
if 'dx' in kwargs and 'dy' in kwargs:
self.set_dxdy(kwargs['dx'], kwargs['dy'])
if 'grid_data' in kwargs:
self.set_grid(kwargs['grid_data'])
if 'grid_title' in kwargs:
self.set_grid_title(kwargs['grid_title'])
if 'var_name' in kwargs:
self.set_variable_name(kwargs['var_name'])
if 'var_unit' in kwargs:
self.set_variable_unit(kwargs['var_unit'])
#
# To Do (2): complete the following functions to set title, name and unit:
#
def set_grid_title(self, grid_title):
self._grid_title = grid_title
def set_variable_name(self, var_name):
self._var_name = var_name
def set_variable_unit(self, var_unit):
self._var_unit = var_unit
# include the plot function here:
def plot(self):
plt.figure(figsize=(8,8))
extent = [0., self._extent_x, 0., self._extent_y]
im = plt.imshow(self._grid_data, extent = extent,
interpolation = 'nearest', cmap = 'viridis')
cbar = plt.colorbar(im, orientation = 'horizontal', fraction=0.042, pad=0.14)
#
# To Do (3): check if variable name and unit are defined, and if so: adjust label
if hasattr(self, '_var_name') and hasattr(self, '_var_unit'):
cbar.set_label("%s [%s]" % (self._var_name, self._var_unit))
else:
cbar.set_label("Grid data")
#
plt.xlabel("X")
plt.ylabel("Y")
#
# To Do (4): check if grid title is defined, and if so: adjust title
if hasattr(self, '_grid_title'):
plt.title("%s" % (self._grid_title))
else:
plt.title("2D grid")
#
plt.show()
# operator overloading
def __add__(self, rhs):
# (1) Check the type of the "rhs" variable
if isinstance(rhs, Grid):
# (2) create a copy of the
new_grid = copy.deepcopy(self)
# (3) perform addition of grid data
new_grid._grid_data = self._grid_data + rhs._grid_data
else:
new_grid = copy.deepcopy(self)
new_grid._grid_data = self._grid_data + rhs
# now we return the updated object entirely:
return new_grid
# function that provides information when used with print statement
def __repr__(self):
# Idea: we construct a string with the information and return it:
# (1) basic information:
info_string = "Grid object with 2-D grid data\n"
# (2) info on grid cells, spacing, extent:
info_string += "Number of cells\t= (%d, %d)\n" % (self._nx, self._ny)
info_string += "Cell dimension\t= (%.1f, %.1f)\n" % (self._dx, self._dy)
info_string += "Grid extent\t= (%.1f, %.1f)\n" % (self._extent_x, self._extent_y)
# (3) add some information and statistics on grid values:
info_string += "Grid data is of type %s with:\n" % type(self._grid_data[0,0])
info_string += "Min value = %.1f\n" % (np.min(self._grid_data))
info_string += "Max value = %.1f\n" % (np.max(self._grid_data))
info_string += "Mean value = %.1f\n" % (np.mean(self._grid_data))
return info_string
def set_grid(self,grid_data):
new_nx, new_ny = grid_data.shape
if new_nx != self._nx:
raise AttributeError("nx is not the right dimension!")
if new_ny != self._ny:
raise AttributeError("nx is not the right dimension!")
self._grid_data = grid_data
def get_grid(self):
return self._grid_data
def update_extent(self):
# first: check if both number of cells and cell widths are already defined
if hasattr(self, '_dx') and hasattr(self, '_nx'):
self._extent_x = self._nx * self._dx
self._extent_y = self._ny * self._dy
def get_extent(self):
return (self._extent_x, self._extent_y)
def set_nxny(self, nx, ny):
self._nx = nx
self._ny = ny
self.update_extent()
def get_nxny(self):
return (self._nx, self._ny)
def set_dxdy(self, dx, dy):
self._dx = dx
self._dy = dy
self.update_extent()
def get_dxdy(self):
return (self._dx, self._dy)
# And if you implemented everything correctly, then the commands in the next cell should work well:
# define a grid with random values
g1 = Grid(nx = nx, ny = ny, dx = dx, dy = dy,
grid_title = "Temperature distribution",
var_name = "Temperature",
var_unit = "C",
grid_data = np.random.randint(1,10,size=(nx,ny)))
print(g1.__dict__)
# create the plot
g1.plot()
# ## An interesting extension
#
# Now think of the following: you have defined a grid, ideally suited for an investigation related to our reservoir enigneering problems. Think about the assignment 1: what would you have to do to extend the example here, so that you can add wells at certain locations, and simulated the effect of fluid extraction/ injection?
import numpy as np
import math
import matplotlib.pylab as plt
from mpl_toolkits.mplot3d import Axes3D
# +
# plt.plot?
# -
class Hydrogrid():
# define the object initialization function
def __init__(self, nx, ny, dx, dy, **kwargs):
"""This is an awesome method to perform analytic grounwater estimations
**Arguments**:
- nx
- ny
- dx
- dy
**Optional Keywords**:
- origin: ['lower']
By default the origin of the grid is at the center of the grid.
Entering 'lower' for the origin, puts the origin at the lower left corner of the grid.
"""
self._nx = nx
self._ny = ny
self._dx = dx
self._dy = dy
if 'origin' in kwargs:
if kwargs['origin'] == 'lower':
x = np.arange(0, self._nx, self._dx)
y = np.arange(0, self._ny, self._dy)
self._X, self._Y = np.meshgrid(x, y)
else:
raise AttributeError("origin can only have 'lower' as input!")
else:
x = np.arange((-0.5*self._nx + 0.5*self._dx), (0.5*self._nx + 0.5*self._dx))
y = np.arange((-0.5*self._ny + 0.5*self._dy), (0.5*self._ny + 0.5*self._dy))
self._X, self._Y = np.meshgrid(x, y)
if 'wx' in kwargs and 'wy' in kwargs:
self.set_wxwy(kwargs['wx'], kwargs['wy'])
def distance(self, wx, wy):
if np.size(wx) != np.size(wy):
raise AttributeError("wx and wy do not have the same size!")
self._wx = wx
self._wy = wy
self._r = np.zeros((2, Hg1._ny, Hg1._nx), dtype = np.float)
for i in range(np.size(wx)):
r_i = np.sqrt((self._X - wx[i])**2 + (self._Y - wy[i])**2)
self._r[i,:,:] = r_i
self._r[self._r == 0] = 0.5 * np.min([self._dx, self._dy])
def drawdown(self, t, T, S, Q):
if np.size(Q) != np.shape(Hg1._r)[0]:
raise AttributeError("Number of wells doesn't match number of pumping rates!")
self._t = t
self._T = T
self._S = S
self._Q = Q
u = (self._r**2*S) / (4*T*t)
W = -0.5772 - np.log(u) + u - (u**2)/(2*math.factorial(2)) + (u**3)/(3*math.factorial(3)) - (u**4)/(4*math.factorial(4))
self._s = np.zeros_like(self._r)
for i in range(np.size(Q)):
s_i = (Q[i]*W[i,:,:])/(4*np.pi*T)
self._s[i,:,:] = s_i
self._s_tot = np.sum(self._s, axis = 0)
def hydro_head(self, h0):
# if np.size(h0) != 1:
# raise AttributeError("h0 has to either be a scalar, or of the same shape as the grid")
self._h0 = h0 * np.ones(np.shape(self._X))
self._h = self._h0 - self._s_tot
def contour_plot(self, n):
self._fig_contf = plt.figure(figsize=(6,4))
ax1 = self._fig_contf.add_subplot(111)
cont1 = ax1.contourf(self._X, self._Y, self._h, n)
cbar = plt.colorbar(cont1, shrink = 1)
cbar.set_label("Drawdown [m]", fontsize = 12)
ax1.set_xlabel("x [m]", fontsize = 12)
ax1.set_ylabel("y [m]", fontsize = 12)
ax1.set_title("Drawdown due to pumping and injection", fontsize = 16)
plt.tight_layout()
def perspective_plot(self):
self._fig_persp = plt.figure(figsize=(18,12))
ax = self._fig_persp.add_subplot(111, projection='3d')
surf = ax.plot_surface(self._X, self._Y, self._h, cmap = 'viridis')
cbar = plt.colorbar(surf, format = '%.3f', shrink = 0.4)
cbar.set_label("Hydraulic head [m]", fontsize = 12)
ax.set_xlabel("x [m]", fontsize = 12)
ax.set_ylabel("y [m]", fontsize = 12)
ax.set_zlabel("Hydraulic head [m]", fontsize = 12, labelpad = 10)
ax.set_title("Hydraulic head field due to pumping and injection", fontsize = 24)
plt.tight_layout()
Hg1 = Hydrogrid(nx = 300, ny = 200, dx = 1, dy = 1)
# print(Hg1.__dict__)
# r = np.zeros((2, Hg1._ny, Hg1._nx), dtype = np.float)
print(np.shape(Hg1._X))
wx = [50, -25, 25]
wy = [50, 0, 0]
Hg1.distance(wx, wy)
print(type(Hg1._r))
print(np.shape(Hg1._r))
print(np.min(Hg1._r))
print(Hg1)
t = 10 * 3600 # time [s]
T = 10E-2 # transmissivity [m^2/s]
S = 10E-1 # storativity [-]
Q = [5E-3, 10E-3, -10E-3] # [pumping flow rate [m^3/s], injection flow rate [m^3/s]]
Hg1.drawdown(t, T, S, Q)
print(type(Hg1._s))
print(np.shape(Hg1._s))
print(np.shape(Hg1._s_tot))
h0 = 100
# h0 =h0 * np.ones(np.shape(Hg1._X))
# h0 - Hg1._s_tot
Hg1.hydro_head(h0)
# +
# print(Hg1.__dict__)
# -
Hg1.contour_plot(n = 30)
Hg1._fig_contf
Hg1.perspective_plot()
Hg1._fig_persp
# +
# Hydrogrid?
# -
|
hellopython/nre_notebooks/rwth_nre_oop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from probability_trees import PTree
from itertools import combinations
def simple_var(name, prob):
return [(prob, f"{name}=1"), (1-prob, f"{name}=0")]
def leaky_or(parents_prob, child, bvars, leak):
result = []
active_parents = [p for p in parents_prob.keys() if bvars[p]=="1"]
for l in range(len(active_parents)+1):
for c in combinations(active_parents, l):
gprob = 1
pvars = []
for parent in active_parents:
prob = parents_prob[parent]
if parent in c:
pvars.append(f"U_{parent}_{child}=1")
gprob *= prob
else:
pvars.append(f"U_{parent}_{child}=0")
gprob *= (1-prob)
if l:
result.append((gprob, ",".join(pvars+[f"{child}=1"])))
else:
result.append((gprob * leak,
",".join(pvars+[f"L_{child}=1,{child}=1"])))
result.append((gprob * (1-leak),
",".join(pvars+[f"L_{child}=0,{child}=0"])))
return result
def asia_model( bvar ):
if 'Bronchitis' not in bvar:
return simple_var("Bronchitis", 0.2)
if 'LungCancer' not in bvar:
return simple_var("LungCancer", 0.07)
if 'Dyspnea' not in bvar:
return leaky_or({"LungCancer": 0.3,
"Bronchitis": 0.1},
"Dyspnea", bvar, 0.3)
if 'Xray' not in bvar:
return leaky_or({"LungCancer": 0.3}, "Xray", bvar, 0.03)
return None
# +
asia = PTree.fromFunc(asia_model, 'Root=1')
xray_and_lung_cancer = asia.prop('Xray=1') & asia.prop('LungCancer=1')
display(asia.show(show_prob=True, cut=xray_and_lung_cancer, crit=asia.critical(xray_and_lung_cancer)))
asia.prob(xray_and_lung_cancer)
# -
def asia_model( bvar ):
p_bronchitis = 0.2
p_lung_cancer = 0.07
u_bronchitis_dyspnea = 0.1
u_lung_cancer_dyspnea = 0.3
u_lung_cancer_xray = 0.3
l_xray = 0.03
l_dyspnea = 0.3
if 'Bronchitis' not in bvar:
return bronchitis_is_exogenous( p_bronchitis, u_bronchitis_dyspnea )
if 'LungCancer' not in bvar:
return lung_cancer_is_exogenous( p_lung_cancer, u_lung_cancer_dyspnea, u_lung_cancer_xray )
if 'Dyspnea' not in bvar:
return dyspnea_if_lung_cancer_or_bronchitis( bvar['LungCancer'], bvar['U_lung_cancer_dyspnea'],
bvar['Bronchitis'], bvar['U_bronchitis_dyspnea'],
l_dyspnea
)
if 'Xray' not in bvar:
return xray_if_lung_cancer( bvar['LungCancer'], bvar['U_lung_cancer_xray'], l_xray )
return None
def bronchitis_is_exogenous( p, q):
return [( (1-p), 'U_bronchitis_dyspnea=0,Bronchitis=0'),
(p*(1-q) , 'U_bronchitis_dyspnea=0,Bronchitis=1'),
(p*q, 'U_bronchitis_dyspnea=1,Bronchitis=1')]
def lung_cancer_is_exogenous( p, q, r):
return [( (1-p) , 'LungCancer=0,U_lung_cancer_dyspnea=0,U_lung_cancer_xray=0'),
(p*(1-q)*(1-r) , 'LungCancer=1,U_lung_cancer_dyspnea=0,U_lung_cancer_xray=0'),
(p*(1-q)*r , 'LungCancer=1,U_lung_cancer_dyspnea=0,U_lung_cancer_xray=1'),
(p*q*(1-r) , 'LungCancer=1,U_lung_cancer_dyspnea=1,U_lung_cancer_xray=0'),
(p*q*r , 'LungCancer=1,U_lung_cancer_dyspnea=1,U_lung_cancer_xray=1')
]
def dyspnea_if_lung_cancer_or_bronchitis(lung_cancer, u_l, bronchitis, u_b, leak):
if (lung_cancer == '1' and u_l == '1') or (bronchitis == '1' and u_b == '1'):
return [(1, 'Dyspnea=1'),
(0, 'Dyspnea=0')]
else:
return [(leak, 'Dyspnea=1'),
(1-leak, 'Dyspnea=0')]
# +
def xray_if_lung_cancer(lung_cancer, u_l, leak):
if (lung_cancer == '1' and u_l == '1'):
return [(1, 'Xray=1'),
(0, 'Xray=0')]
else:
return [(leak, 'Xray=1'),
(1-leak, 'Xray=0')]
# -
def
|
notebooks/Richens-with-Ptrees.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="MhoQ0WE77laV"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" id="_ckMIh7O7s6D"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + cellView="form" id="vasWnqRgy1H4"
#@title MIT License
#
# Copyright (c) 2017 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a
# # copy of this software and associated documentation files (the "Software"),
# to deal in the Software without restriction, including without limitation
# the rights to use, copy, modify, merge, publish, distribute, sublicense,
# and/or sell copies of the Software, and to permit persons to whom the
# Software is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in
# all copies or substantial portions of the Software.
#
# THE SOFTWARE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR
# IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY,
# FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL
# THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER
# LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING
# FROM, OUT OF OR IN CONNECTION WITH THE SOFTWARE OR THE USE OR OTHER
# DEALINGS IN THE SOFTWARE.
# + [markdown] id="jYysdyb-CaWM"
# # 기본 분류: 의류 이미지 선형 분류
# + [markdown] id="S5Uhzt6vVIB2"
# <table class="tfo-notebook-buttons" align="left">
# <td> <a target="_blank" href="https://www.tensorflow.org/tutorials/keras/classification"><img src="https://www.tensorflow.org/images/tf_logo_32px.png">TensorFlow.org에서 보기</a> </td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ko/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png">Google Colab에서 실행</a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ko/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png">GitHub에서 소스 보기</a></td>
# <td><a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ko/tutorials/keras/classification.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png">노트북 다운로드</a></td>
# </table>
# + [markdown] id="FbVhjPpzn6BM"
# Note: 이 실습은 Tensorflow에 있는 자료를 바탕으로 간단하게 변형하여 ML 기초학습을 한 실습 정리 및 메모한 자료입니다.
#
# 이 튜토리얼에서는 운동화나 셔츠 같은 옷 이미지를 분류하는 신경망 모델을 훈련합니다. 상세 내용을 모두 이해하지 못해도 괜찮습니다. 여기서는 완전한 텐서플로(TensorFlow) 프로그램을 빠르게 살펴 보겠습니다. 자세한 내용은 앞으로 배우면서 더 설명합니다.
#
# 여기에서는 텐서플로 모델을 만들고 훈련할 수 있는 고수준 API인 [tf.keras](https://www.tensorflow.org/guide/keras)를 사용합니다.
# + id="dzLKpmZICaWN" colab={"base_uri": "https://localhost:8080/"} outputId="5c23731e-f251-401c-ed23-f8ef93dab101"
# tensorflow와 tf.keras를 임포트합니다
import tensorflow as tf
from tensorflow import keras
# 헬퍼(helper) 라이브러리를 임포트합니다
import numpy as np
import matplotlib.pyplot as plt
print(tf.__version__)
# + [markdown] id="yR0EdgrLCaWR"
# ## 패션 MNIST 데이터셋 임포트하기
# + [markdown] id="DLdCchMdCaWQ"
# 10개의 범주(category)와 70,000개의 흑백 이미지로 구성된 [패션 MNIST](https://github.com/zalandoresearch/fashion-mnist) 데이터셋을 사용하겠습니다. 이미지는 해상도(28x28 픽셀)가 낮고 다음처럼 개별 옷 품목을 나타냅니다:
#
# <table>
# <tr><td> <img src="https://tensorflow.org/images/fashion-mnist-sprite.png" alt="Fashion MNIST sprite" width="600"> </td></tr>
# <tr><td align="center"> <b>그림 1.</b> <a href="https://github.com/zalandoresearch/fashion-mnist">패션-MNIST 샘플</a> (Zalando, MIT License).<br>{nbsp} </td></tr>
# </table>
#
# 패션 MNIST는 컴퓨터 비전 분야의 "Hello, World" 프로그램격인 고전 [MNIST](http://yann.lecun.com/exdb/mnist/) 데이터셋을 대신해서 자주 사용됩니다. MNIST 데이터셋은 손글씨 숫자(0, 1, 2 등)의 이미지로 이루어져 있습니다. 여기서 사용하려는 옷 이미지와 동일한 포맷입니다.
#
# 패션 MNIST는 일반적인 MNIST 보다 조금 더 어려운 문제이고 다양한 예제를 만들기 위해 선택했습니다. 두 데이터셋은 비교적 작기 때문에 알고리즘의 작동 여부를 확인하기 위해 사용되곤 합니다. 코드를 테스트하고 디버깅하는 용도로 좋습니다.
#
# 여기에서 60,000개의 이미지를 사용하여 네트워크를 훈련하고 10,000개의 이미지를 사용하여 네트워크에서 이미지 분류를 학습한 정도를 평가합니다. TensorFlow에서 직접 Fashion MNIST에 액세스할 수 있습니다. TensorFlow에서 직접 [Fashion MNIST 데이터](https://www.tensorflow.org/api_docs/python/tf/keras/datasets/fashion_mnist/load_data)를 가져오고 로드합니다.
# + id="7MqDQO0KCaWS" colab={"base_uri": "https://localhost:8080/"} outputId="2dd156bc-ee7d-4a79-8480-2ec25e5eb0c2"
fashion_mnist = keras.datasets.fashion_mnist # 케라스에서 기본적으로 제공하는 데이터셋
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# + [markdown] id="t9FDsUlxCaWW"
# load_data() 함수를 호출하면 네 개의 넘파이(NumPy) 배열이 반환됩니다:
#
# - `train_images`와 `train_labels` 배열은 모델 학습에 사용되는 *훈련 세트*입니다.
# - `test_images`와 `test_labels` 배열은 모델 테스트에 사용되는 *테스트 세트*입니다.
#
# 이미지는 28x28 크기의 넘파이 배열이고 픽셀 값은 0과 255 사이입니다. *레이블*(label)은 0에서 9까지의 정수 배열입니다. 이 값은 이미지에 있는 옷의 *클래스*(class)를 나타냅니다:
#
# <table>
# <tr>
# <th>레이블</th>
# <th>클래스</th>
# </tr>
# <tr>
# <td>0</td>
# <td>T-shirt/top</td>
# </tr>
# <tr>
# <td>1</td>
# <td>Trouser</td>
# </tr>
# <tr>
# <td>2</td>
# <td>Pullover</td>
# </tr>
# <tr>
# <td>3</td>
# <td>Dress</td>
# </tr>
# <tr>
# <td>4</td>
# <td>Coat</td>
# </tr>
# <tr>
# <td>5</td>
# <td>Sandal</td>
# </tr>
# <tr>
# <td>6</td>
# <td>Shirt</td>
# </tr>
# <tr>
# <td>7</td>
# <td>Sneaker</td>
# </tr>
# <tr>
# <td>8</td>
# <td>Bag</td>
# </tr>
# <tr>
# <td>9</td>
# <td>Ankle boot</td>
# </tr>
# </table>
#
# 각 이미지는 하나의 레이블에 매핑되어 있습니다. 데이터셋에 *클래스 이름*이 들어있지 않기 때문에 나중에 이미지를 출력할 때 사용하기 위해 별도의 변수를 만들어 저장합니다:
# + id="IjnLH5S2CaWx"
# 데이터셋에 숫자로만 되어 있는 클래스를 문자로 정의
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + [markdown] id="Brm0b_KACaWX"
# ## 데이터 탐색
#
# 모델을 훈련하기 전에 데이터셋 구조를 살펴보죠. 다음 코드는 훈련 세트에 60,000개의 이미지가 있다는 것을 보여줍니다. 각 이미지는 28x28 픽셀로 표현됩니다:
# + id="zW5k_xz1CaWX" colab={"base_uri": "https://localhost:8080/"} outputId="1f0fdf64-251c-40bc-b5b8-20997b4bd2f1"
train_images.shape
# + colab={"base_uri": "https://localhost:8080/"} id="6D0Dj3lbIJ0p" outputId="a3cde4a7-e798-4742-c158-a49e77d28f74"
test_images.shape
# + [markdown] id="cIAcvQqMCaWf"
# 비슷하게 훈련 세트에는 60,000개의 레이블이 있습니다:
# + id="TRFYHB2mCaWb" colab={"base_uri": "https://localhost:8080/"} outputId="89d8b668-dbe4-4a05-e9dd-a3fee236561b"
len(train_labels), len(test_labels) # y값
# + [markdown] id="YSlYxFuRCaWk"
# 각 레이블은 0과 9사이의 정수입니다:
# + id="XKnCTHz4CaWg" colab={"base_uri": "https://localhost:8080/"} outputId="c03f53a0-2aa4-479e-a0da-889e0d481657"
train_labels
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="a3z6JiIHIibV" outputId="ecde55c0-14cb-4ee0-a786-a11cf4767182"
class_names[train_labels[0]]
# + [markdown] id="TMPI88iZpO2T"
# 테스트 세트에는 10,000개의 이미지가 있습니다. 이 이미지도 28x28 픽셀로 표현됩니다:
# + id="2KFnYlcwCaWl" colab={"base_uri": "https://localhost:8080/"} outputId="aadd6b87-4d0c-486f-bcc8-f4143580c5f9"
test_images.shape
# + [markdown] id="rd0A0Iu0CaWq"
# 테스트 세트는 10,000개의 이미지에 대한 레이블을 가지고 있습니다:
# + id="iJmPr5-ACaWn" colab={"base_uri": "https://localhost:8080/"} outputId="9df36e95-7e4d-44fa-ef05-9fcc2ec5ef91"
len(test_labels)
# + [markdown] id="ES6uQoLKCaWr"
# ## 데이터 전처리
#
# 네트워크를 훈련하기 전에 데이터를 전처리해야 합니다. 훈련 세트에 있는 첫 번째 이미지를 보면 픽셀 값의 범위가 0~255 사이라는 것을 알 수 있습니다:
# + id="m4VEw8Ud9Quh" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="2005af86-9bdc-44f0-d9ca-d010985a7d46"
plt.figure()
plt.imshow(train_images[0])
plt.colorbar()
plt.grid(False)
plt.show()
# + [markdown] id="Wz7l27Lz9S1P"
# 신경망 모델에 주입하기 전에 이 값의 범위를 0~1 사이로 조정하겠습니다. 이렇게 하려면 255로 나누어야 합니다. *훈련 세트*와 *테스트 세트*를 동일한 방식으로 전처리하는 것이 중요합니다:
# + id="bW5WzIPlCaWv"
# 정규화 (0~1 범위로)
train_images = train_images / 255.0
test_images = test_images / 255.0
# + [markdown] id="Ee638AlnCaWz"
# *훈련 세트*에서 처음 25개 이미지와 그 아래 클래스 이름을 출력해 보죠. 데이터 포맷이 올바른지 확인하고 네트워크 구성과 훈련할 준비를 마칩니다.
# + id="oZTImqg_CaW1" colab={"base_uri": "https://localhost:8080/", "height": 589} outputId="93aebc7c-ad6d-4c40-b08f-660d3e0e1935"
plt.figure(figsize=(10,10))
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[train_labels[i]])
plt.show()
# + [markdown] id="59veuiEZCaW4"
# ## 모델 구성
#
# 신경망 모델을 만들려면 모델의 층을 구성한 다음 모델을 컴파일합니다.
# + [markdown] id="Gxg1XGm0eOBy"
# ### 층 설정
#
# 신경망의 기본 빌딩 블록은 [*레이어*](https://www.tensorflow.org/api_docs/python/tf/keras/layers) 입니다. 레이어는 레이어에 공급된 데이터로부터 표현을 추출합니다. 이러한 표현은 당면한 문제에 의미가 있어야 합니다.
#
# 대부분 딥러닝은 간단한 층을 연결하여 구성됩니다. `tf.keras.layers.Dense`와 같은 층들의 가중치(parameter)는 훈련하는 동안 학습됩니다.
# + id="e5Cg7wLFKh5y"
from tensorflow.keras import layers
# + id="hN3ACzf_I_ZK"
inputs = keras.Input(shape=(28,28))
h = layers.Flatten()(inputs) # 일렬로 쭉 펴주는 작업 (행렬과 곱해질려면 28x28이 쫙펴져야함)
h = layers.Dense(128)(h)
outputs = layers.Dense(10)(h) # 10가지 클래스로 분류하는 모델이기에 각 10개의 클래스마다 점수를 부여
outputs = layers.Activation('softmax')(outputs) # softmax : 전체의 합을 1로 만들어줌 (이전 regression과의 차이)
model = keras.Model(inputs=inputs, outputs=outputs)
# + colab={"base_uri": "https://localhost:8080/"} id="wdcPb_rtKmSK" outputId="f746f679-5a99-46fa-a9ad-252135b7111a"
model.summary()
# + [markdown] id="gut8A_7rCaW6"
# 이 네트워크의 첫 번째 층인 `tf.keras.layers.Flatten`은 2차원 배열(28 x 28 픽셀)의 이미지 포맷을 28 * 28 = 784 픽셀의 1차원 배열로 변환합니다. 이 층은 이미지에 있는 픽셀의 행을 펼쳐서 일렬로 늘립니다. 이 층에는 학습되는 가중치가 없고 데이터를 변환하기만 합니다.
#
# 픽셀을 펼친 후에는 두 개의 `tf.keras.layers.Dense` 층이 연속되어 연결됩니다. 이 층을 밀집 연결(densely-connected) 또는 완전 연결(fully-connected) 층이라고 부릅니다. 첫 번째 `Dense` 층은 128개의 노드(또는 뉴런)를 가집니다. 두 번째 (마지막) 층은 10개의 노드의 *소프트맥스*(softmax) 층입니다. 이 층은 10개의 확률을 반환하고 반환된 값의 전체 합은 1입니다. 각 노드는 현재 이미지가 10개 클래스 중 하나에 속할 확률을 출력합니다.
#
# ### 모델 컴파일
#
# 모델을 훈련할 준비가 되기 전에 몇 가지 설정이 더 필요합니다. 다음은 모델의 [*컴파일*](https://www.tensorflow.org/api_docs/python/tf/keras/Model#compile) 단계에서 추가됩니다.
#
# - [*손실 함수*](https://www.tensorflow.org/api_docs/python/tf/keras/losses) - 훈련 중 모델이 얼마나 정확한지 측정합니다. 모델을 올바른 방향으로 "조정"하려면 이 함수를 최소화해야 합니다.
# - [*옵티마이저*](https://www.tensorflow.org/api_docs/python/tf/keras/optimizers) - 모델이 인식하는 데이터와 해당 손실 함수를 기반으로 모델이 업데이트되는 방식입니다.
# - [*메트릭*](https://www.tensorflow.org/api_docs/python/tf/keras/metrics) — 훈련 및 테스트 단계를 모니터링하는 데 사용됩니다. 다음 예에서는 올바르게 분류된 이미지의 비율인 *정확도*를 사용합니다.
# + id="Lhan11blCaW7"
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy', # 회귀에서는 'mse','mae'같은걸 사용하지만 / 분류에서는 카테고리칼 크로스엔트로피 사용(분류의 로스계산)
metrics=['accuracy'])
# + [markdown] id="qKF6uW-BCaW-"
# ## 모델 훈련
#
# 신경망 모델을 훈련하려면 다음 단계가 필요합니다.
#
# 1. 훈련 데이터를 모델에 주입합니다-이 예에서는 `train_images`와 `train_labels` 배열입니다.
# 2. 모델이 이미지와 레이블을 매핑하는 방법을 배웁니다.
# 3. 테스트 세트에 대한 모델의 예측을 만듭니다-이 예에서는 `test_images` 배열입니다. 이 예측이 `test_labels` 배열의 레이블과 맞는지 확인합니다.
# 4. 예측이 `test_labels` 배열의 레이블과 일치하는지 확인합니다.
#
# + [markdown] id="Z4P4zIV7E28Z"
# ### 모델 피드
#
# 훈련을 시작하려면 [`model.fit`](https://www.tensorflow.org/api_docs/python/tf/keras/Model#fit) 메서드를 호출합니다. 모델을 훈련 데이터에 "맞추기(fit)" 때문에 이렇게 불립니다.
# + id="xvwvpA64CaW_" colab={"base_uri": "https://localhost:8080/"} outputId="6ea79688-1242-401f-818b-e012faec244b"
model.fit(train_images, train_labels, epochs=5, validation_split=0.1)
# 트레이닝 로스랑 벨리데이션 로스랑 비슷한걸 보니 오버피팅은 아님
# + [markdown] id="W3ZVOhugCaXA"
# 모델이 훈련되면서 손실과 정확도 지표가 출력됩니다. 이 모델은 훈련 세트에서 약 0.88(88%) 정도의 정확도를 달성합니다.
# + [markdown] id="wCpr6DGyE28h"
# ### 정확도 평가
#
# 다음으로, 모델이 테스트 데이터세트에서 작동하는 방식을 비교합니다.
# + id="VflXLEeECaXC" colab={"base_uri": "https://localhost:8080/"} outputId="30873232-ac3d-4026-c02b-53f75bb23238"
test_loss, test_acc = model.evaluate(test_images, test_labels, verbose=2)
print('\n테스트 정확도:', test_acc)
# + [markdown] id="yWfgsmVXCaXG"
# 테스트 세트의 정확도가 훈련 세트의 정확도보다 조금 낮습니다. 훈련 세트의 정확도와 테스트 세트의 정확도 사이의 차이는 *과대적합*(overfitting) 때문입니다. 과대적합은 머신러닝 모델이 훈련 데이터보다 새로운 데이터에서 성능이 낮아지는 현상을 말합니다.
#
# - [과대적합 시연](https://www.tensorflow.org/tutorials/keras/overfit_and_underfit#demonstrate_overfitting)
# - [과대적합을 방지하기 위한 전략](https://www.tensorflow.org/tutorials/keras/overfit_and_underfit#strategies_to_prevent_overfitting)
# + [markdown] id="v-PyD1SYE28q"
# ### 예측하기
#
# 훈련된 모델을 사용하여 일부 이미지에 대한 예측을 수행할 수 있습니다. 모델의 선형 출력, [로짓](https://developers.google.com/machine-learning/glossary#logits). 소프트맥스 레이어를 연결하여 로짓을 해석하기 쉬운 확률로 변환합니다.
# + id="Gl91RPhdCaXI"
predictions = model.predict(test_images)
# + [markdown] id="x9Kk1voUCaXJ"
# 여기서는 테스트 세트에 있는 각 이미지의 레이블을 예측했습니다. 첫 번째 예측을 확인해 보죠:
# + id="3DmJEUinCaXK" colab={"base_uri": "https://localhost:8080/"} outputId="9ea7ad9b-0141-4efe-92e8-738c77753748"
predictions[0] # output이 10개여서 클래스 10개에 대한 각 점수, 이중 최대값이 이 0번째 그림의 클래스로 분류
# + [markdown] id="-hw1hgeSCaXN"
# 이 예측은 10개의 숫자 배열로 나타납니다. 이 값은 10개의 옷 품목에 상응하는 모델의 신뢰도(confidence)를 나타냅니다. 가장 높은 신뢰도를 가진 레이블을 찾아보죠:
# + id="qsqenuPnCaXO" colab={"base_uri": "https://localhost:8080/"} outputId="7cfcae9e-169d-4caa-9e17-84dc5e38e60d"
np.argmax(predictions[0]) # 최종 예측값 확인
# + [markdown] id="E51yS7iCCaXO"
# 모델은 이 이미지가 앵클 부츠(`class_name[9]`)라고 가장 확신하고 있습니다. 이 값이 맞는지 테스트 레이블을 확인해 보죠:
# + id="Sd7Pgsu6CaXP" colab={"base_uri": "https://localhost:8080/"} outputId="98a44c25-0d82-4f43-a3d7-b7097ebabb7d"
test_labels[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="0Db8nP9fMzHc" outputId="865d23c2-70db-42c6-cf1c-ae2213b520cb"
import matplotlib.pyplot as plt
plt.imshow(test_images[0], cmap=plt.cm.binary)
plt.xlabel(class_names[np.argmax(predictions[0])])
plt.show()
|
opensourceSW_DeveloperContest/ML_03_linear_classification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + id="06CfGvQe8rTp"
pip install PyPortfolioOpt
# + id="pTyGEAap8tEz"
pip install yfinance
# + id="MIZdovsh8u7X"
pip install pulp
# + id="3slTC5ml8xQB"
pip install quandl
# + colab={"base_uri": "https://localhost:8080/", "height": 238} executionInfo={"elapsed": 53, "status": "ok", "timestamp": 1633585901469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GggkDwz2kbsiN3be9s1OYgcBFC5-sc3hTfGRlNnyg=s64", "userId": "17101909825884887186"}, "user_tz": -330} id="W7_ne6HBSz2j" outputId="e65c531b-d411-46ae-eb21-739e44f55e44"
#equal risk contribution / risk parity
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
df= pd.read_csv("capstone_input_data.csv")
df.Date = pd.to_datetime(df.Date, dayfirst = True)
df.set_index("Date", inplace = True)
Portfolio_value = 10000000
returns = df.pct_change()
returns = returns[1:]
returns.shape
returns.tail()
# -
Portfolio_value = 10000000
weights = pd.read_csv("weightsHRP.csv")
weights= np.array(weights)
weights.shape
pnl = (weights * returns.values).sum(axis=1)
pnl.shape
historic_var95 = np.percentile(pnl, 5, interpolation = "lower")
historic_var99 = np.percentile(pnl, 1, interpolation = "lower")
historic_var99
Daily_VaR95 = Portfolio_value*historic_var95
Monthly_VaR95 = Daily_VaR95 * np.sqrt(22)
Monthly_VaR95
Daily_VaR99 = Portfolio_value*historic_var99
Monthly_VaR99 = Daily_VaR99 * np.sqrt(22)
Monthly_VaR99
Daily_VaR99 = Portfolio_value*historic_var99
Monthly_VaR99 = Daily_VaR99 * np.sqrt(22)
Monthly_VaR99
# ### CVaR
var_level = 95
var_95 = np.percentile(pnl, 100 - var_level)
cvar_95 = pnl[pnl <= var_95].mean()
cvar_95
CVaR_port =cvar_95*Portfolio_value
CVaR_port
var_level2 = 99
var_99 = np.percentile(pnl, 100 - var_level2)
cvar_99 = pnl[pnl <= var_99].mean()
CVaR_port99 =cvar_99*Portfolio_value
CVaR_port99
output = [['Portfolio Value', Portfolio_value], ['Daily_VaR_95', Daily_VaR95],['Monthly_VaR95', Monthly_VaR95],['Daily_VaR_99', Daily_VaR99],['Monthly_VaR99', Monthly_VaR99], ['Daily_CVAR_95', CVaR_port],['Daily_CVAR_99', CVaR_port99]]
output2 = pd.DataFrame(output, columns=['Details', " Amount in Mn"])
output2
# - VaR is conventionally reported as a positive number.
# - A negative VaR would imply the portfolio has a high probability of making a profit
# - For example a one-day 5% VaR of negative 1 million implies the portfolio has a 95 percent chance of making more than 1 million over the next day.
|
Hierarchical_Risk_Parity-with-VaR-CVaR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programmatic Replacements for NICMOS Units Conversion Form
#
# This notebook illustrates programmatic ways to perform unit conversions as provided by [NICMOS Units Conversion Form](http://www.stsci.edu/hst/nicmos/tools/conversion_form.html). The examples here are not exhaustive and will never be. They are meant to give users enough basic knowledge to then calculate what they really want.
# ## Imports
# - All the examples require `astropy.units`. See `Units` and `Quantities` in Astropy for more information.
# - Some examples require the package `snyphot`.
# - Some examples require the `math` library.
# +
from astropy import units as u
from synphot import SourceSpectrum
from synphot import units as syn_u
from synphot.models import BlackBodyNorm1D
import math
# -
# ## Example 1: Simple Jy to AB mag
#
# ```
# INPUT FROM THE FORM IS
# Input units = Jy
# Output units = AB magnitude
# Input flux = 1.00000E-13 Jy
# Temperature of the blackbody = 5500.00
# INPUT wavelength = 1.00000 micron
# OUTPUT wavelength = 1.00000 micron
# AB magnitude = 41.43
# ```
#
# In plain English, user wants to know what is 1e-13 Jy converted to AB magnitude at 1 micron. Blackbody is not needed for this conversion.
input_flux_ex1 = 1e-13 * u.Jy
input_wave_ex1 = 1 * u.micron
abmag_ex1 = input_flux_ex1.to(u.ABmag, u.spectral_density(input_wave_ex1))
print('{:.2f}'.format(abmag_ex1))
# ## Example 2: PHOTLAM to FNU for Blackbody
#
# ```
# INPUT FROM THE FORM IS
# Input units = photons/cm2/s/A
# Output units = erg/cm2/s/Hz
# Input flux = 1.00000 photons/cm2/s/A
# Temperature of the blackbody = 5500.00
# INPUT wavelength = 0.500000 micron
# OUTPUT wavelength = 0.600000 micron
# Flux=4.62E-23 erg/cm2/s/Hz
# ```
# This example is more complicated in that it requires assumption of a blackbody and has different input and output units.
#
# First, we create a source spectrum with a blackbody model with the given temperature of 5500 K.
bb_ex2 = SourceSpectrum(BlackBodyNorm1D, temperature=5500*u.K)
# Then, we calculate the normalization factor required for the blackbody to have the given input flux at the given input wavelength.
input_flux_ex2 = 1 * syn_u.PHOTLAM
input_wave_ex2 = 0.5 * u.micron
factor_ex2 = input_flux_ex2 / bb_ex2(input_wave_ex2)
# We apply this factor to our blackbody source.
input_ex2 = bb_ex2 * factor_ex2
# Finally, we calculate the desired flux in given output unit and wavelength.
output_wave_ex2 = 0.6 * u.micron
flux_ex2 = input_ex2(output_wave_ex2, flux_unit=syn_u.FNU)
print('{:.2e}'.format(flux_ex2))
# ## Example 3: W/m2/Hz to I-mag for Power-Law
#
#
# ```
# INPUT FROM THE FORM IS
# Input units = W/m2/Hz
# Output units = magnitude I
# Input flux = 1.00000E-23 W/m2/Hz
# Index of the power-law spectrum as a function of frequency = 0.250000
# INPUT wavelength = 1.00000 micron
# OUTPUT wavelength = 0.900000 micron
# I = 0.85
# ```
#
# First, we define the input flux and wavelength.
input_flux_ex3 = 1e-23 * (u.W / (u.m * u.m * u.Hz))
input_wave_ex3 = 1 * u.micron
# Then, we define a power-law function.
def powerlaw_ex3(nu):
"""F(nu)=nu**(spectral index)
nu is a Quantity.
"""
spectral_index = 0.25
return (nu.to(u.Hz, u.spectral()) ** spectral_index).value
# We use this power-law and a normalization factor based on input flux and wavelength to calculate output flux at output wavelength in input flux unit.
output_wave_ex3 = 0.9 * u.micron
flux_ex3 = powerlaw_ex3(output_wave_ex3) * input_flux_ex3 / powerlaw_ex3(input_wave_ex3)
# Finally, we convert the flux to *I* magnitude, as defined by the converter as:
#
# ```
# magnitude I: zero-flux=2250 Jy; central wavelength=0.90 microns;
# ```
imag_zpt = 2250 * u.Jy
i = -2.5 * math.log10(flux_ex3 / imag_zpt)
print('{:.2f}'.format(i))
|
notebooks/NICMOS/nicmos_unit_conversion/nicmos_unit_conversion.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# _The following excercises are adapted from <NAME>'s Introducing Python_
# 1- How many seconds are in an hour?
# 2- Assign the result from the previous task to a variable called seconds_per_hour
# 3- How many seconds are in a day? Use your seconds_per_hour variable
# 4- Calculate seconds per day again, but this time save the result in a variable called seconds_per_day
# 5- Divide seconds_per_day by seconds_per_hour. Use floating-point (/) division.
# 6- Divide seconds_per_day by seconds_per_hour using integer (//) division. Did this number agree with the floating-point value from the previous question, aside from the final .0?
|
03-warmup-blank_variables.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
from fastai.gen_doc.gen_notebooks import *
from pathlib import Path
# ### To update this notebook
# Run `tools/sgen_notebooks.py
# Or run below:
# You need to make sure to refresh right after
import glob
for f in Path().glob('*.ipynb'):
generate_missing_metadata(f)
# # Metadata generated below
update_nb_metadata('callbacks.csv_logger.ipynb',
summary='Callbacks that saves the tracked metrics during training',
title='callbacks.csv_logger')
update_nb_metadata('callbacks.tracker.ipynb',
summary='Callbacks that take decisions depending on the evolution of metrics during training',
title='callbacks.tracker')
# + hide_input=false
update_nb_metadata('torch_core.ipynb',
summary='Basic functions using pytorch',
title='torch_core')
# + hide_input=false
update_nb_metadata('gen_doc.convert2html.ipynb',
summary='Converting the documentation notebooks to HTML pages',
title='gen_doc.convert2html')
# + hide_input=false
update_nb_metadata('metrics.ipynb',
summary='Useful metrics for training',
title='metrics')
# + hide_input=false
update_nb_metadata('callbacks.fp16.ipynb',
summary='Training in mixed precision implementation',
title='callbacks.fp16')
# + hide_input=false
update_nb_metadata('callbacks.general_sched.ipynb',
summary='Implementation of a flexible training API',
title='callbacks.general_sched')
# + hide_input=false
update_nb_metadata('text.ipynb',
keywords='fastai',
summary='Application to NLP, including ULMFiT fine-tuning',
title='text')
# + hide_input=false
update_nb_metadata('callback.ipynb',
summary='Implementation of the callback system',
title='callback')
# + hide_input=false
update_nb_metadata('tabular.models.ipynb',
keywords='fastai',
summary='Model for training tabular/structured data',
title='tabular.models')
# + hide_input=false
update_nb_metadata('callbacks.mixup.ipynb',
summary='Implementation of mixup',
title='callbacks.mixup')
# + hide_input=false
update_nb_metadata('applications.ipynb',
summary='Types of problems you can apply the fastai library to',
title='applications')
# + hide_input=false
update_nb_metadata('vision.data.ipynb',
summary='Basic dataset for computer vision and helper function to get a DataBunch',
title='vision.data')
# + hide_input=false
update_nb_metadata('overview.ipynb',
summary='Overview of the core modules',
title='overview')
# + hide_input=false
update_nb_metadata('training.ipynb',
keywords='fastai',
summary='Overview of fastai training modules, including Learner, metrics, and callbacks',
title='training')
# + hide_input=false
update_nb_metadata('text.transform.ipynb',
summary='NLP data processing; tokenizes text and creates vocab indexes',
title='text.transform')
# + hide_input=false
# do not overwrite this notebook, or changes may get lost!
# update_nb_metadata('jekyll_metadata.ipynb')
# + hide_input=false
update_nb_metadata('collab.ipynb',
summary='Application to collaborative filtering',
title='collab')
# + hide_input=false
update_nb_metadata('text.learner.ipynb',
summary='Easy access of language models and ULMFiT',
title='text.learner')
# + hide_input=false
update_nb_metadata('gen_doc.nbdoc.ipynb',
summary='Helper function to build the documentation',
title='gen_doc.nbdoc')
# + hide_input=false
update_nb_metadata('vision.learner.ipynb',
summary='`Learner` support for computer vision',
title='vision.learner')
# + hide_input=false
update_nb_metadata('core.ipynb',
summary='Basic helper functions for the fastai library',
title='core')
# + hide_input=false
update_nb_metadata('fastai_typing.ipynb',
keywords='fastai',
summary='Type annotations names',
title='fastai_typing')
# + hide_input=false
update_nb_metadata('gen_doc.gen_notebooks.ipynb',
summary='Generation of documentation notebook skeletons from python module',
title='gen_doc.gen_notebooks')
# + hide_input=false
update_nb_metadata('basic_train.ipynb',
summary='Learner class and training loop',
title='basic_train')
# + hide_input=false
update_nb_metadata('gen_doc.ipynb',
keywords='fastai',
summary='Documentation modules overview',
title='gen_doc')
# + hide_input=false
update_nb_metadata('callbacks.rnn.ipynb',
summary='Implementation of a callback for RNN training',
title='callbacks.rnn')
# + hide_input=false
update_nb_metadata('callbacks.one_cycle.ipynb',
summary='Implementation of the 1cycle policy',
title='callbacks.one_cycle')
# + hide_input=false
update_nb_metadata('vision.ipynb',
summary='Application to Computer Vision',
title='vision')
# + hide_input=false
update_nb_metadata('vision.transform.ipynb',
summary='List of transforms for data augmentation in CV',
title='vision.transform')
# + hide_input=false
update_nb_metadata('callbacks.lr_finder.ipynb',
summary='Implementation of the LR Range test from <NAME>',
title='callbacks.lr_finder')
# + hide_input=false
update_nb_metadata('text.data.ipynb',
summary='Basic dataset for NLP tasks and helper functions to create a DataBunch',
title='text.data')
# + hide_input=false
update_nb_metadata('text.models.ipynb',
summary='Implementation of the AWD-LSTM and the RNN models',
title='text.models')
# + hide_input=false
update_nb_metadata('tabular.data.ipynb',
summary='Base class to deal with tabular data and get a DataBunch',
title='tabular.data')
# + hide_input=false
update_nb_metadata('callbacks.ipynb',
keywords='fastai',
summary='Callbacks implemented in the fastai library',
title='callbacks')
# + hide_input=false
update_nb_metadata('train.ipynb',
summary='Extensions to Learner that easily implement Callback',
title='train')
# + hide_input=false
update_nb_metadata('callbacks.hooks.ipynb',
summary='Implement callbacks using hooks',
title='callbacks.hooks')
# + hide_input=false
update_nb_metadata('vision.image.ipynb',
summary='Image class, variants and internal data augmentation pipeline',
title='vision.image')
# + hide_input=false
update_nb_metadata('vision.models.unet.ipynb',
summary='Dynamic Unet that can use any pretrained model as a backbone.',
title='vision.models.unet')
# + hide_input=false
update_nb_metadata('vision.models.ipynb',
keywords='fastai',
summary='Overview of the models used for CV in fastai',
title='vision.models')
# + hide_input=false
update_nb_metadata('tabular.transform.ipynb',
summary='Transforms to clean and preprocess tabular data',
title='tabular.transform')
# + hide_input=false
update_nb_metadata('index.ipynb',
keywords='fastai',
toc='false',
title='Welcome to fastai')
# + hide_input=false
update_nb_metadata('layers.ipynb',
summary='Provides essential functions to building and modifying `Model` architectures.',
title='layers')
# + hide_input=false
update_nb_metadata('tabular.ipynb',
keywords='fastai',
summary='Application to tabular/structured data',
title='tabular')
# + hide_input=false
update_nb_metadata('basic_data.ipynb',
summary='Basic classes to contain the data for model training.',
title='basic_data')
# + hide_input=false
update_nb_metadata('datasets.ipynb')
# + hide_input=false
update_nb_metadata('tmp.ipynb',
keywords='fastai')
# + hide_input=false
update_nb_metadata('callbacks.tracking.ipynb')
# + hide_input=false
update_nb_metadata('data_block.ipynb',
keywords='fastai',
summary='The data block API',
title='data_block')
# + hide_input=false
update_nb_metadata('callbacks.tracker.ipynb',
keywords='fastai',
summary='Callbacks that take decisions depending on the evolution of metrics during training',
title='callbacks.tracking')
# + hide_input=false
update_nb_metadata('widgets.ipynb')
# + hide_input=false
update_nb_metadata('text_tmp.ipynb')
# + hide_input=false
update_nb_metadata('tabular_tmp.ipynb')
|
docs_src/jekyll_metadata.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="z1y8_SXpdine"
# # Import statements
# + colab={"base_uri": "https://localhost:8080/"} id="Iogvt5eI-NOk" executionInfo={"status": "ok", "timestamp": 1618644843706, "user_tz": -330, "elapsed": 21774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="619a22c6-aad3-44be-adfc-e88e132ebe85"
from google.colab import drive
drive.mount('/content/drive')
# + id="utDoiASySB39" executionInfo={"status": "ok", "timestamp": 1618644864550, "user_tz": -330, "elapsed": 1292, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
from my_ml_lib import MetricTools, PlotTools
# + id="2GPNbC8UdlZX" executionInfo={"status": "ok", "timestamp": 1618644868299, "user_tz": -330, "elapsed": 4535, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
import os
import numpy as np
import matplotlib.pyplot as plt
import pickle
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.pyplot import figure
import json
import datetime
import copy
from PIL import Image as im
import joblib
from sklearn.model_selection import train_test_split
# import math as Math
import random
import torch.optim
# + id="XznHCx6ydmmS" executionInfo={"status": "ok", "timestamp": 1618644868960, "user_tz": -330, "elapsed": 5107, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
from torch.utils.data import DataLoader
from torch.utils.data import Dataset
import torchvision
# + id="2FRmNHO_dmrF" executionInfo={"status": "ok", "timestamp": 1618644869436, "user_tz": -330, "elapsed": 5501, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
import cv2
# + id="vqL1Kq0idnDC" executionInfo={"status": "ok", "timestamp": 1618644869436, "user_tz": -330, "elapsed": 5418, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# + [markdown] id="HVR5HtsIdncF"
# # Saving and Loading code
# + id="P9yFkBbUdoz7" executionInfo={"status": "ok", "timestamp": 1618644869437, "user_tz": -330, "elapsed": 4299, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# Saving and Loading models using joblib
def save(filename, obj):
with open(filename, 'wb') as handle:
joblib.dump(obj, handle, protocol=pickle.HIGHEST_PROTOCOL)
def load(filename):
with open(filename, 'rb') as handle:
return joblib.load(filename)
# + [markdown] id="WVxLGeMgdqkW"
# # Importing Dataset
# + id="B__Ue0eRdsaj" executionInfo={"status": "ok", "timestamp": 1618644871030, "user_tz": -330, "elapsed": 5300, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
p = "/content/drive/MyDrive/A3/"
data_path = p + "dataset/train.pkl"
x = load(data_path)
# + id="4j-AhjELo0At" executionInfo={"status": "ok", "timestamp": 1618644871031, "user_tz": -330, "elapsed": 5199, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# save_path = "/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/dataset/"
# # saving the images and labels array
# save(save_path + "data_image.pkl",data_image)
# save(save_path + "data_label.pkl",data_label)
# # dict values where labels key and image arrays as vlaues in form of list
# save(save_path + "my_dict.pkl",my_dict)
# + id="zKurY8cao0HV" executionInfo={"status": "ok", "timestamp": 1618644872657, "user_tz": -330, "elapsed": 6758, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
save_path = p + "dataset/"
# saving the images and labels array
data_image = load(save_path + "data_image.pkl")
data_label = load(save_path + "data_label.pkl")
# dict values where labels key and image arrays as vlaues in form of list
my_dict = load(save_path + "my_dict.pkl")
# + colab={"base_uri": "https://localhost:8080/"} id="K1Mjytyoo0OM" executionInfo={"status": "ok", "timestamp": 1618644874093, "user_tz": -330, "elapsed": 1419, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="ea17611e-39c6-4d61-b673-4527b4ebbe0d"
len(data_image) , len(data_label), my_dict.keys()
# + [markdown] id="reu4bmKXzZch"
# # Data Class and Data Loaders and Data transforms
# + colab={"base_uri": "https://localhost:8080/"} id="BUHDt_iEzbtB" executionInfo={"status": "ok", "timestamp": 1618644874093, "user_tz": -330, "elapsed": 1412, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="09cb080a-7f78-48bd-b2bc-03797ed9424f"
len(x['names']) ,x['names'][4999] , data_image[0].shape
# + [markdown] id="7UY8CjMb3L45"
# ## Splitting the data into train and val
# + id="QaTSdOG63Qk_" executionInfo={"status": "ok", "timestamp": 1618644874094, "user_tz": -330, "elapsed": 1410, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
X_train, X_test, y_train, y_test = train_test_split(data_image, data_label, test_size=0.10, random_state=42,stratify=data_label )
# + colab={"base_uri": "https://localhost:8080/"} id="WrRaxUfb30Nh" executionInfo={"status": "ok", "timestamp": 1618644874094, "user_tz": -330, "elapsed": 936, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="1d080805-682c-4957-df55-dc995656681f"
len(X_train) , len(y_train) , len(X_test) ,len(y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="-PYnG0eY4idN" executionInfo={"status": "ok", "timestamp": 1618644874518, "user_tz": -330, "elapsed": 1043, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="595611cf-c37d-47f2-8ce5-fd4bbae208a4"
pd.DataFrame(y_test).value_counts()
# + [markdown] id="8DBejOVP3Ts9"
# ## Data Class
# + id="pd72TAxBwLkw" executionInfo={"status": "ok", "timestamp": 1618644884381, "user_tz": -330, "elapsed": 1332, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
train_transforms = transforms.Compose([
transforms.RandomHorizontalFlip(),
])
val_transforms = transforms.Compose([
])
# + id="LiDsHK8MzcD4" executionInfo={"status": "ok", "timestamp": 1618644884382, "user_tz": -330, "elapsed": 940, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
class myDataClass(Dataset):
"""Custom dataset class"""
def __init__(self, images, labels , transform=None):
"""
Args:
images : Array of all the images
labels : Correspoing labels of all the images
"""
self.images = images
self.labels = labels
self.transform = transform
def __len__(self):
return len(self.images)
def __getitem__(self, idx):
# converts image value between 0 and 1 and returns a tensor C,H,W
img = torchvision.transforms.functional.to_tensor(self.images[idx])
target = self.labels[idx]
if self.transform:
img = self.transform(img)
return img,target
# + [markdown] id="P-M6IB6o3WQh"
# ## Data Loaders
# + id="TGBYO6J_zcGe" executionInfo={"status": "ok", "timestamp": 1618644903371, "user_tz": -330, "elapsed": 1345, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
batch = 64
train_dataset = myDataClass(X_train, y_train,train_transforms)
test_dataset = myDataClass(X_test, y_test)
train_dataloader = DataLoader(train_dataset, batch_size= batch, shuffle=True)
test_dataloader = DataLoader(test_dataset, batch_size= batch, shuffle=True)
# + id="XlZB_gOB2_KQ" executionInfo={"status": "ok", "timestamp": 1618644903373, "user_tz": -330, "elapsed": 886, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# next(iter(train_dataloader))[0].shape
# + colab={"base_uri": "https://localhost:8080/"} id="NxSyJNaI26T_" executionInfo={"status": "ok", "timestamp": 1618644904045, "user_tz": -330, "elapsed": 1055, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="4455ef4e-f716-434a-c935-8b1696d14d6c"
len(train_dataloader) , len(test_dataloader)
# + [markdown] id="bpz1XVRu_ql9"
# # Train and Test functions
# + id="0_3OoxMmNg1t" executionInfo={"status": "ok", "timestamp": 1618644904045, "user_tz": -330, "elapsed": 886, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
def load_best(all_models,model_test):
FILE = all_models[-1]
criterion = nn.CrossEntropyLoss()
optimizer = optim.Adam(model_test.parameters(), lr=0)
checkpoint = torch.load(FILE)
model_test.load_state_dict(checkpoint['model_state'])
optimizer.load_state_dict(checkpoint['optim_state'])
epoch = checkpoint['epoch']
model_test.eval()
return model_test
# + id="IawJoK7T_xqi" executionInfo={"status": "ok", "timestamp": 1618644904577, "user_tz": -330, "elapsed": 1319, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
def train(save_path,epochs,train_dataloader,model,test_dataloader,optimizer,criterion,basic_name):
model_no = 1
c = 1
all_models = []
valid_loss_min = np.Inf
train_losses = []
val_losses = []
for e in range(epochs):
train_loss = 0.0
valid_loss = 0.0
model.train()
for idx, (images,labels) in enumerate(train_dataloader):
images, labels = images.to(device) , labels.to(device)
optimizer.zero_grad()
log_ps= model(images)
loss = criterion(log_ps, labels)
loss.backward()
optimizer.step()
train_loss += ((1 / (idx + 1)) * (loss.data - train_loss))
else:
accuracy = 0
correct = 0
model.eval()
with torch.no_grad():
for idx, (images,labels) in enumerate(test_dataloader):
images, labels = images.to(device) , labels.to(device)
log_ps = model(images)
_, predicted = torch.max(log_ps.data, 1)
loss = criterion(log_ps, labels)
# correct += (predicted == labels).sum().item()
equals = predicted == labels.view(*predicted.shape)
accuracy += torch.mean(equals.type(torch.FloatTensor))
valid_loss += ((1 / (idx + 1)) * (loss.data - valid_loss))
print('Epoch: {} \tTraining Loss: {:.6f} \tValidation Loss: {:.6f}'.format(
e+1,
train_loss,
valid_loss
), "Test Accuracy: {:.3f}".format(accuracy/len(test_dataloader)))
train_losses.append(train_loss)
val_losses.append(valid_loss)
if valid_loss < valid_loss_min:
print('Saving model..' + str(model_no))
valid_loss_min = valid_loss
checkpoint = {
"epoch": e+1,
"model_state": model.state_dict(),
"optim_state": optimizer.state_dict(),
"train_losses": train_losses,
"test_losses": val_losses,
}
FILE = save_path + basic_name +"_epoch_" + str(e+1) + "_model_" + str(model_no)
all_models.append(FILE)
torch.save(checkpoint, FILE)
model_no = model_no + 1
save(save_path + basic_name + "_all_models.pkl", all_models)
return model, train_losses, val_losses, all_models
# + id="9S6qsEzj_x9n" executionInfo={"status": "ok", "timestamp": 1618644904577, "user_tz": -330, "elapsed": 1237, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
def plot(train_losses,val_losses,title='Training Validation Loss with CNN'):
plt.plot(train_losses, label='Training loss')
plt.plot(val_losses, label='Validation loss')
plt.xlabel('Iterations')
plt.ylabel('Loss')
plt.legend()
_ = plt.ylim()
plt.title(title)
# plt.savefig('plots/Training Validation Loss with CNN from scratch.png')
plt.show()
# + id="GkMgxEVD_yJN" executionInfo={"status": "ok", "timestamp": 1618644904578, "user_tz": -330, "elapsed": 1167, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
def test(loader, model, criterion, device, name):
test_loss = 0.
correct = 0.
total = 0.
y = None
y_hat = None
model.eval()
for batch_idx, (images, labels) in enumerate(loader):
# move to GPU or CPU
images, labels = images.to(device) , labels.to(device)
target = labels
# forward pass: compute predicted outputs by passing inputs to the model
output = model(images)
# calculate the loss
loss = criterion(output,labels)
# update average test loss
test_loss = test_loss + ((1 / (batch_idx + 1)) * (loss.data - test_loss))
# convert output probabilities to predicted class
pred = output.data.max(1, keepdim=True)[1]
if y is None:
y = target.cpu().numpy()
y_hat = pred.data.cpu().view_as(target).numpy()
else:
y = np.append(y, target.cpu().numpy())
y_hat = np.append(y_hat, pred.data.cpu().view_as(target).numpy())
correct += np.sum(pred.view_as(labels).cpu().numpy() == labels.cpu().numpy())
total = total + images.size(0)
# if batch_idx % 20 == 0:
# print("done till batch" , batch_idx+1)
print(name + ' Loss: {:.6f}\n'.format(test_loss))
print(name + ' Accuracy: %2d%% (%2d/%2d)' % (
100. * correct / total, correct, total))
return y, y_hat
# + id="efG8-oAkEv3z" executionInfo={"status": "ok", "timestamp": 1618644904578, "user_tz": -330, "elapsed": 1081, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + id="wpqqkahKHE4l" executionInfo={"status": "ok", "timestamp": 1618644905518, "user_tz": -330, "elapsed": 718, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# def train(save_path,epochs,train_dataloader,model,test_dataloader,optimizer,criterion,basic_name)
# def plot(train_losses,val_losses,title='Training Validation Loss with CNN')
# def test(loader, model, criterion, device)
# + [markdown] id="wec0leCLEwYo"
# # Relu [ X=2 Y=3 Z=1 ]
# + [markdown] id="uzqyfCNfqiXN"
# ## CNN-Block-123
# + [markdown] id="sVJG2N7gqmoh"
# ### model
# + id="bs9JxROAqmHf" executionInfo={"status": "ok", "timestamp": 1618644906811, "user_tz": -330, "elapsed": 1245, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
cfg3 = {
'B123': [16,16,'M','D',32,32,32,'M','D',64,'M','D'],
}
# + id="Iu5D4gUWqq2t" executionInfo={"status": "ok", "timestamp": 1618644907392, "user_tz": -330, "elapsed": 888, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
def make_layers3(cfg, batch_norm=True):
layers = []
in_channels = 3
for v in cfg:
if v == 'M':
layers += [nn.AvgPool2d(kernel_size=2, stride=2)]
elif v == 'M1':
layers += [nn.AvgPool2d(kernel_size=4, stride=3)]
elif v == 'D':
layers += [nn.Dropout(p=0.5)]
else:
conv2d = nn.Conv2d(in_channels, v, kernel_size=3)
if batch_norm:
layers += [conv2d, nn.BatchNorm2d(v), nn.ReLU(inplace=True)]
else:
layers += [conv2d, nn.ReLU(inplace=True)]
in_channels = v
return nn.Sequential(*layers)
# + id="ewWE-5oTrA1v" executionInfo={"status": "ok", "timestamp": 1618644908021, "user_tz": -330, "elapsed": 1028, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
class Model_B123(nn.Module):
'''
Model
'''
def __init__(self, features):
super(Model_B123, self).__init__()
self.features = features
self.classifier = nn.Sequential(
# nn.Linear(1600, 512),
# nn.ReLU(True),
# nn.Linear(512, 256),
# nn.ReLU(True),
# nn.Linear(256, 64),
# nn.ReLU(True),
nn.Linear(64, 10),
)
def forward(self, x):
x = self.features(x)
# print(x.shape)
x = x.view(x.size(0), -1)
x = self.classifier(x)
return x
# + id="gOOCCTbarA-A" executionInfo={"status": "ok", "timestamp": 1618644908021, "user_tz": -330, "elapsed": 946, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# m = Model_B123(make_layers3(cfg3['B123']))
# for i,l in train_dataloader:
# o = m(i)
# + colab={"base_uri": "https://localhost:8080/"} id="v6ZRV-LOrrTr" executionInfo={"status": "ok", "timestamp": 1618644918970, "user_tz": -330, "elapsed": 11809, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="31abea75-4986-4e16-de8b-7693e70f78b2"
model3 = Model_B123(make_layers3(cfg3['B123'])).to(device)
learning_rate = 0.001
criterion3 = nn.CrossEntropyLoss()
optimizer3 = optim.Adam(model3.parameters(), lr=learning_rate)
print(model3)
# + [markdown] id="J64Z8g_Jqr7V"
# ### train
# + id="JeJ9mIPcqrAP" executionInfo={"status": "ok", "timestamp": 1618644931696, "user_tz": -330, "elapsed": 1308, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
# # !rm '/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/models_saved_Q1/1_3/bw_blocks/Dropout(0.5)/cnn_block123/'*
# # !ls '/content/drive/MyDrive/SEM-2/05-DL /Assignments/A3/models_saved_Q1/1_3/bw_blocks/Dropout(0.5)/cnn_block123/'
# + id="ktg9rq0EqrH0" executionInfo={"status": "ok", "timestamp": 1618644931697, "user_tz": -330, "elapsed": 930, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
save_path3 = p + "models_saved_Q1/1_4/colab_notebooks /Batchnorm_and_pooling/models/"
# + colab={"base_uri": "https://localhost:8080/"} id="Oi-qyO4ir16f" executionInfo={"status": "ok", "timestamp": 1618645021290, "user_tz": -330, "elapsed": 90107, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="332ccb94-3293-4b85-e4c2-fa4a907aaf11"
m, train_losses, val_losses,m_all_models = train(save_path3,100,train_dataloader,model3,test_dataloader,optimizer3,criterion3,"cnn_b123_x2_y3_z1_with_BN_avg_pool_dp_without_WD_0.001_with_aug")
# + [markdown] id="iEF6Xr44qtf3"
# ### Tests and Plots
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="OJME3KDGqrQW" executionInfo={"status": "ok", "timestamp": 1618645023179, "user_tz": -330, "elapsed": 1883, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="3b16d78d-adf7-42f5-eb12-f0ce9fea1bc2"
plot(train_losses,val_losses,'Training Validation Loss with CNN-block1')
# + id="aGueEimwr6hw" executionInfo={"status": "ok", "timestamp": 1618645055791, "user_tz": -330, "elapsed": 1271, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
all_models3 = load(save_path3 + "cnn_b123_x2_y3_z1_with_BN_avg_pool_dp_without_WD_0.001_with_aug_all_models.pkl")
FILE = all_models3[-1]
m3 = Model_B123(make_layers3(cfg3['B123'])).to(device)
# + id="Lp0nDcyjr6pW" executionInfo={"status": "ok", "timestamp": 1618645055791, "user_tz": -330, "elapsed": 798, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
m3 = load_best(all_models3,m3)
# + colab={"base_uri": "https://localhost:8080/"} id="Hht6dGo_r6w4" executionInfo={"status": "ok", "timestamp": 1618645056921, "user_tz": -330, "elapsed": 1528, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="3571b7ca-99eb-4244-9f8a-26ab8f977864"
train_y, train_y_hat = test(train_dataloader, m3, criterion3, device, "TRAIN")
# + colab={"base_uri": "https://localhost:8080/", "height": 441} id="JOH7tQRUr64k" executionInfo={"status": "ok", "timestamp": 1618645057982, "user_tz": -330, "elapsed": 2085, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="eb32e173-cfc0-4a9c-ee52-a9f77e38d7b1"
cm = MetricTools.confusion_matrix(train_y, train_y_hat, nclasses=10)
PlotTools.confusion_matrix(cm, [i for i in range(10)], title='',
filename='Confusion Matrix with CNN', figsize=(6,6))
# + colab={"base_uri": "https://localhost:8080/"} id="cZcUExXEsGA5" executionInfo={"status": "ok", "timestamp": 1618645057983, "user_tz": -330, "elapsed": 2001, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="0b3931c2-0887-4ee1-cac3-6f3b1cc0c858"
test_y, test_y_hat = test(test_dataloader, m3, criterion3, device,"TEST")
# + colab={"base_uri": "https://localhost:8080/", "height": 441} id="iU7y_iHHsGLO" executionInfo={"status": "ok", "timestamp": 1618645058507, "user_tz": -330, "elapsed": 2425, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}} outputId="23671418-eb08-423b-8c18-85053730a2d0"
cm = MetricTools.confusion_matrix(test_y, test_y_hat, nclasses=10)
PlotTools.confusion_matrix(cm, [i for i in range(10)], title='',
filename='Confusion Matrix with CNN', figsize=(6,6))
# + id="Xy--Uk5_y1y1" executionInfo={"status": "ok", "timestamp": 1618645058508, "user_tz": -330, "elapsed": 2374, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Gj5-9IST_h6Gk-NrsbfHFXN6c1NPVKRjReYd9J54Q=s64", "userId": "09190534878071493768"}}
|
IIIT_Delhi_assignments/003_Convolutional_Neural_networks/A3_MT20055_Akanksha_MT20018_Vaibhav_MT20121_Shivam/Part(1)/1_4/data_augumentation/Deliverable_Part_1_4_Layers_(2,3,1)_with_BN_with_avg_pooling_dp_without_weight_Decay_LR(0_001)_augmentation.ipynb
|