code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Chapter 7 – Ensemble Learning and Random Forests**
# _This notebook contains all the sample code and solutions to the exercices in chapter 7._
# # Setup
# First, let's make sure this notebook works well in both python 2 and 3, import a few common modules, ensure MatplotLib plots figures inline and prepare a function to save the figures:
# +
# To support both python 2 and python 3
from __future__ import division, print_function, unicode_literals
# Common imports
import numpy as np
import os
# to make this notebook's output stable across runs
np.random.seed(42)
# To plot pretty figures
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
plt.rcParams['axes.labelsize'] = 14
plt.rcParams['xtick.labelsize'] = 12
plt.rcParams['ytick.labelsize'] = 12
# Where to save the figures
PROJECT_ROOT_DIR = "."
CHAPTER_ID = "ensembles"
def image_path(fig_id):
return os.path.join(PROJECT_ROOT_DIR, "images", CHAPTER_ID, fig_id)
def save_fig(fig_id, tight_layout=True):
print("Saving figure", fig_id)
if tight_layout:
plt.tight_layout()
plt.savefig(image_path(fig_id) + ".png", format='png', dpi=300)
# -
# # Voting classifiers
heads_proba = 0.51
coin_tosses = (np.random.rand(10000, 10) < heads_proba).astype(np.int32)
cumulative_heads_ratio = np.cumsum(coin_tosses, axis=0) / np.arange(1, 10001).reshape(-1, 1)
plt.figure(figsize=(8,3.5))
plt.plot(cumulative_heads_ratio)
plt.plot([0, 10000], [0.51, 0.51], "k--", linewidth=2, label="51%")
plt.plot([0, 10000], [0.5, 0.5], "k-", label="50%")
plt.xlabel("Number of coin tosses")
plt.ylabel("Heads ratio")
plt.legend(loc="lower right")
plt.axis([0, 10000, 0.42, 0.58])
save_fig("law_of_large_numbers_plot")
plt.show()
# +
from sklearn.model_selection import train_test_split
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=500, noise=0.30, random_state=42)
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import VotingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
log_clf = LogisticRegression(random_state=42)
rnd_clf = RandomForestClassifier(random_state=42)
svm_clf = SVC(random_state=42)
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('rf', rnd_clf), ('svc', svm_clf)],
voting='hard')
voting_clf.fit(X_train, y_train)
# +
from sklearn.metrics import accuracy_score
for clf in (log_clf, rnd_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, accuracy_score(y_test, y_pred))
# +
log_clf = LogisticRegression(random_state=42)
rnd_clf = RandomForestClassifier(random_state=42)
svm_clf = SVC(probability=True, random_state=42)
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('rf', rnd_clf), ('svc', svm_clf)],
voting='soft')
voting_clf.fit(X_train, y_train)
# +
from sklearn.metrics import accuracy_score
for clf in (log_clf, rnd_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, accuracy_score(y_test, y_pred))
# -
# # Bagging ensembles
# +
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
bag_clf = BaggingClassifier(
DecisionTreeClassifier(random_state=42), n_estimators=500,
max_samples=100, bootstrap=True, n_jobs=-1, random_state=42)
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
# -
from sklearn.metrics import accuracy_score
print(accuracy_score(y_test, y_pred))
tree_clf = DecisionTreeClassifier(random_state=42)
tree_clf.fit(X_train, y_train)
y_pred_tree = tree_clf.predict(X_test)
print(accuracy_score(y_test, y_pred_tree))
# +
from matplotlib.colors import ListedColormap
def plot_decision_boundary(clf, X, y, axes=[-1.5, 2.5, -1, 1.5], alpha=0.5, contour=True):
x1s = np.linspace(axes[0], axes[1], 100)
x2s = np.linspace(axes[2], axes[3], 100)
x1, x2 = np.meshgrid(x1s, x2s)
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = clf.predict(X_new).reshape(x1.shape)
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap, linewidth=10)
if contour:
custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])
plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo", alpha=alpha)
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs", alpha=alpha)
plt.axis(axes)
plt.xlabel(r"$x_1$", fontsize=18)
plt.ylabel(r"$x_2$", fontsize=18, rotation=0)
# -
plt.figure(figsize=(11,4))
plt.subplot(121)
plot_decision_boundary(tree_clf, X, y)
plt.title("Decision Tree", fontsize=14)
plt.subplot(122)
plot_decision_boundary(bag_clf, X, y)
plt.title("Decision Trees with Bagging", fontsize=14)
save_fig("decision_tree_without_and_with_bagging_plot")
plt.show()
# # Random Forests
bag_clf = BaggingClassifier(
DecisionTreeClassifier(splitter="random", max_leaf_nodes=16, random_state=42),
n_estimators=500, max_samples=1.0, bootstrap=True, n_jobs=-1, random_state=42)
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
# +
from sklearn.ensemble import RandomForestClassifier
rnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, n_jobs=-1, random_state=42)
rnd_clf.fit(X_train, y_train)
y_pred_rf = rnd_clf.predict(X_test)
# -
np.sum(y_pred == y_pred_rf) / len(y_pred) # almost identical predictions
from sklearn.datasets import load_iris
iris = load_iris()
rnd_clf = RandomForestClassifier(n_estimators=500, n_jobs=-1, random_state=42)
rnd_clf.fit(iris["data"], iris["target"])
for name, score in zip(iris["feature_names"], rnd_clf.feature_importances_):
print(name, score)
rnd_clf.feature_importances_
# +
plt.figure(figsize=(6, 4))
for i in range(15):
tree_clf = DecisionTreeClassifier(max_leaf_nodes=16, random_state=42 + i)
indices_with_replacement = np.random.randint(0, len(X_train), len(X_train))
tree_clf.fit(X[indices_with_replacement], y[indices_with_replacement])
plot_decision_boundary(tree_clf, X, y, axes=[-1.5, 2.5, -1, 1.5], alpha=0.02, contour=False)
plt.show()
# -
# ## Out-of-Bag evaluation
bag_clf = BaggingClassifier(
DecisionTreeClassifier(random_state=42), n_estimators=500,
bootstrap=True, n_jobs=-1, oob_score=True, random_state=40)
bag_clf.fit(X_train, y_train)
bag_clf.oob_score_
bag_clf.oob_decision_function_
from sklearn.metrics import accuracy_score
y_pred = bag_clf.predict(X_test)
accuracy_score(y_test, y_pred)
# ## Feature importance
from sklearn.datasets import fetch_mldata
mnist = fetch_mldata('MNIST original')
rnd_clf = RandomForestClassifier(random_state=42)
rnd_clf.fit(mnist["data"], mnist["target"])
def plot_digit(data):
image = data.reshape(28, 28)
plt.imshow(image, cmap = matplotlib.cm.hot,
interpolation="nearest")
plt.axis("off")
# +
plot_digit(rnd_clf.feature_importances_)
cbar = plt.colorbar(ticks=[rnd_clf.feature_importances_.min(), rnd_clf.feature_importances_.max()])
cbar.ax.set_yticklabels(['Not important', 'Very important'])
save_fig("mnist_feature_importance_plot")
plt.show()
# -
# # AdaBoost
# +
from sklearn.ensemble import AdaBoostClassifier
ada_clf = AdaBoostClassifier(
DecisionTreeClassifier(max_depth=1), n_estimators=200,
algorithm="SAMME.R", learning_rate=0.5, random_state=42)
ada_clf.fit(X_train, y_train)
# -
plot_decision_boundary(ada_clf, X, y)
# +
m = len(X_train)
plt.figure(figsize=(11, 4))
for subplot, learning_rate in ((121, 1), (122, 0.5)):
sample_weights = np.ones(m)
for i in range(5):
plt.subplot(subplot)
svm_clf = SVC(kernel="rbf", C=0.05, random_state=42)
svm_clf.fit(X_train, y_train, sample_weight=sample_weights)
y_pred = svm_clf.predict(X_train)
sample_weights[y_pred != y_train] *= (1 + learning_rate)
plot_decision_boundary(svm_clf, X, y, alpha=0.2)
plt.title("learning_rate = {}".format(learning_rate - 1), fontsize=16)
plt.subplot(121)
plt.text(-0.7, -0.65, "1", fontsize=14)
plt.text(-0.6, -0.10, "2", fontsize=14)
plt.text(-0.5, 0.10, "3", fontsize=14)
plt.text(-0.4, 0.55, "4", fontsize=14)
plt.text(-0.3, 0.90, "5", fontsize=14)
save_fig("boosting_plot")
plt.show()
# -
list(m for m in dir(ada_clf) if not m.startswith("_") and m.endswith("_"))
# # Gradient Boosting
np.random.seed(42)
X = np.random.rand(100, 1) - 0.5
y = 3*X[:, 0]**2 + 0.05 * np.random.randn(100)
# +
from sklearn.tree import DecisionTreeRegressor
tree_reg1 = DecisionTreeRegressor(max_depth=2, random_state=42)
tree_reg1.fit(X, y)
# -
y2 = y - tree_reg1.predict(X)
tree_reg2 = DecisionTreeRegressor(max_depth=2, random_state=42)
tree_reg2.fit(X, y2)
y3 = y2 - tree_reg2.predict(X)
tree_reg3 = DecisionTreeRegressor(max_depth=2, random_state=42)
tree_reg3.fit(X, y3)
X_new = np.array([[0.8]])
y_pred = sum(tree.predict(X_new) for tree in (tree_reg1, tree_reg2, tree_reg3))
y_pred
# +
def plot_predictions(regressors, X, y, axes, label=None, style="r-", data_style="b.", data_label=None):
x1 = np.linspace(axes[0], axes[1], 500)
y_pred = sum(regressor.predict(x1.reshape(-1, 1)) for regressor in regressors)
plt.plot(X[:, 0], y, data_style, label=data_label)
plt.plot(x1, y_pred, style, linewidth=2, label=label)
if label or data_label:
plt.legend(loc="upper center", fontsize=16)
plt.axis(axes)
plt.figure(figsize=(11,11))
plt.subplot(321)
plot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h_1(x_1)$", style="g-", data_label="Training set")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.title("Residuals and tree predictions", fontsize=16)
plt.subplot(322)
plot_predictions([tree_reg1], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1)$", data_label="Training set")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.title("Ensemble predictions", fontsize=16)
plt.subplot(323)
plot_predictions([tree_reg2], X, y2, axes=[-0.5, 0.5, -0.5, 0.5], label="$h_2(x_1)$", style="g-", data_style="k+", data_label="Residuals")
plt.ylabel("$y - h_1(x_1)$", fontsize=16)
plt.subplot(324)
plot_predictions([tree_reg1, tree_reg2], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1) + h_2(x_1)$")
plt.ylabel("$y$", fontsize=16, rotation=0)
plt.subplot(325)
plot_predictions([tree_reg3], X, y3, axes=[-0.5, 0.5, -0.5, 0.5], label="$h_3(x_1)$", style="g-", data_style="k+")
plt.ylabel("$y - h_1(x_1) - h_2(x_1)$", fontsize=16)
plt.xlabel("$x_1$", fontsize=16)
plt.subplot(326)
plot_predictions([tree_reg1, tree_reg2, tree_reg3], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="$h(x_1) = h_1(x_1) + h_2(x_1) + h_3(x_1)$")
plt.xlabel("$x_1$", fontsize=16)
plt.ylabel("$y$", fontsize=16, rotation=0)
save_fig("gradient_boosting_plot")
plt.show()
# +
from sklearn.ensemble import GradientBoostingRegressor
gbrt = GradientBoostingRegressor(max_depth=2, n_estimators=3, learning_rate=1.0, random_state=42)
gbrt.fit(X, y)
# -
gbrt_slow = GradientBoostingRegressor(max_depth=2, n_estimators=200, learning_rate=0.1, random_state=42)
gbrt_slow.fit(X, y)
# +
plt.figure(figsize=(11,4))
plt.subplot(121)
plot_predictions([gbrt], X, y, axes=[-0.5, 0.5, -0.1, 0.8], label="Ensemble predictions")
plt.title("learning_rate={}, n_estimators={}".format(gbrt.learning_rate, gbrt.n_estimators), fontsize=14)
plt.subplot(122)
plot_predictions([gbrt_slow], X, y, axes=[-0.5, 0.5, -0.1, 0.8])
plt.title("learning_rate={}, n_estimators={}".format(gbrt_slow.learning_rate, gbrt_slow.n_estimators), fontsize=14)
save_fig("gbrt_learning_rate_plot")
plt.show()
# -
# ## Gradient Boosting with Early stopping
# +
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_squared_error
X_train, X_val, y_train, y_val = train_test_split(X, y, random_state=49)
gbrt = GradientBoostingRegressor(max_depth=2, n_estimators=120, random_state=42)
gbrt.fit(X_train, y_train)
errors = [mean_squared_error(y_val, y_pred)
for y_pred in gbrt.staged_predict(X_val)]
bst_n_estimators = np.argmin(errors)
gbrt_best = GradientBoostingRegressor(max_depth=2,n_estimators=bst_n_estimators, random_state=42)
gbrt_best.fit(X_train, y_train)
# -
min_error = np.min(errors)
# +
plt.figure(figsize=(11, 4))
plt.subplot(121)
plt.plot(errors, "b.-")
plt.plot([bst_n_estimators, bst_n_estimators], [0, min_error], "k--")
plt.plot([0, 120], [min_error, min_error], "k--")
plt.plot(bst_n_estimators, min_error, "ko")
plt.text(bst_n_estimators, min_error*1.2, "Minimum", ha="center", fontsize=14)
plt.axis([0, 120, 0, 0.01])
plt.xlabel("Number of trees")
plt.title("Validation error", fontsize=14)
plt.subplot(122)
plot_predictions([gbrt_best], X, y, axes=[-0.5, 0.5, -0.1, 0.8])
plt.title("Best model (%d trees)" % bst_n_estimators, fontsize=14)
save_fig("early_stopping_gbrt_plot")
plt.show()
# +
gbrt = GradientBoostingRegressor(max_depth=2, warm_start=True, random_state=42)
min_val_error = float("inf")
error_going_up = 0
for n_estimators in range(1, 120):
gbrt.n_estimators = n_estimators
gbrt.fit(X_train, y_train)
y_pred = gbrt.predict(X_val)
val_error = mean_squared_error(y_val, y_pred)
if val_error < min_val_error:
min_val_error = val_error
error_going_up = 0
else:
error_going_up += 1
if error_going_up == 5:
break # early stopping
# -
print(gbrt.n_estimators)
# # Exercise solutions
# **Coming soon**
|
07_ensemble_learning_and_random_forests.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# +
using BenchmarkTools
# Create example array
S = (rand(4,4,1000) + 1im .* rand(4,4,1000)) ./ sqrt(2);
# Manual inversion
function f(A)
B = similar(A)
for i in 1:size(A,3)
B[:,:,i] = inv(@view A[:,:,i])
end
return B
end
@btime f($S)
# +
function g(A)
B = similar(A)
a = similar(A, size(A)[1:2])
u = one(a)
for i in 1:size(A,3)
a = @view A[:,:,i]
b = @view B[:,:,i]
b .= inv(a)
end
return B
end
@btime g($S)
# -
@btime mapslices(inv, $S; dims=(1,2))
@code_warntype mapslices(inv, S; dims=(1,2))
@code_warntype mapslices(identity, S; dims=(1,2))
@code_warntype Base.:(var"#mapslices#134")((1,2), mapslices, inv, S)
|
0017/type instability of mapslice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/livanshu/Data_Science_Portfolio/blob/main/TensorFlow/Intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="-5u3a4csUPyn"
# #TensorFlow 2.0 Introduction
# In this notebook you will be given an interactive introduction to TensorFlow 2.0. We will walk through the following topics within the TensorFlow module:
#
# - TensorFlow Install and Setup
# - Representing Tensors
# - Tensor Shape and Rank
# - Types of Tensors
#
#
# If you'd like to follow along without installing TensorFlow on your machine you can use **Google Collaboratory**. Collaboratory is a free Jupyter notebook environment that requires no setup and runs entirely in the cloud.
# + [markdown] id="F7ThfbiQl96l"
# ##Installing TensorFlow
# To install TensorFlow on your local machine you can use pip.
# ```console
# pip install tensorflow
# ```
# + [markdown] id="JYQWyAJ2mez6"
# If you have a CUDA enabled GPU you can install the GPU version of TensorFlow. You will also need to install some other software which can be found here: https://www.tensorflow.org/install/gpu
# ```console
# pip install tensorflow-gpu
# ```
# + [markdown] id="JJjNMaSClWhg"
# ## Importing TensorFlow
# The first step here is going to be to select the correct version of TensorFlow from within collabratory!
#
# + id="vGcE8x2Gkw9K"
# %tensorflow_version 2.x # this line is not required unless you are in a notebook
# + id="4N7XbNDVY8P3"
import tensorflow as tf # now import the tensorflow module
print(tf.version) # make sure the version is 2.x
# + [markdown] id="duDj86TfWFof"
# ##Tensors
# "A tensor is a generalization of vectors and matrices to potentially higher dimensions. Internally, TensorFlow represents tensors as n-dimensional arrays of base datatypes." (https://www.tensorflow.org/guide/tensor)
#
# It should't surprise you that tensors are a fundemental apsect of TensorFlow. They are the main objects that are passed around and manipluated throughout the program. Each tensor represents a partialy defined computation that will eventually produce a value. TensorFlow programs work by building a graph of Tensor objects that details how tensors are related. Running different parts of the graph allow results to be generated.
#
# Each tensor has a data type and a shape.
#
# **Data Types Include**: float32, int32, string and others.
#
# **Shape**: Represents the dimension of data.
#
# Just like vectors and matrices tensors can have operations applied to them like addition, subtraction, dot product, cross product etc.
#
# In the next sections we will discuss some different properties of tensors. This is to make you more familiar with how tensorflow represnts data and how you can manipulate this data.
#
# + [markdown] id="TAk6QhGUwQRt"
# ###Creating Tensors
# Below is an example of how to create some different tensors.
#
# You simply define the value of the tensor and the datatype and you are good to go! It's worth mentioning that usually we deal with tensors of numeric data, it is quite rare to see string tensors.
#
# For a full list of datatypes please refer to the following guide.
#
# https://www.tensorflow.org/api_docs/python/tf/dtypes/DType?version=stable
# + id="epGskXdjZHzu"
string = tf.Variable("this is a string", tf.string)
number = tf.Variable(324, tf.int16)
floating = tf.Variable(3.567, tf.float64)
# + [markdown] id="D0_H71HMaE-5"
# ###Rank/Degree of Tensors
# Another word for rank is degree, these terms simply mean the number of dimensions involved in the tensor. What we created above is a *tensor of rank 0*, also known as a scalar.
#
# Now we'll create some tensors of higher degrees/ranks.
# + id="hX_Cc5IfjQ6-"
rank1_tensor = tf.Variable(["Test"], tf.string)
rank2_tensor = tf.Variable([["test", "ok"], ["test", "yes"]], tf.string)
# + [markdown] id="55zuGMc7nHjC"
# **To determine the rank** of a tensor we can call the following method.
# + id="Zrj0rAWLnMNv" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b8b34a8c-fc6e-40c6-9fd6-294d1253631e"
tf.rank(rank2_tensor)
# + [markdown] id="hTv4Gz67pQbx"
# The rank of a tensor is direclty related to the deepest level of nested lists. You can see in the first example ```["Test"]``` is a rank 1 tensor as the deepest level of nesting is 1.
# Where in the second example ```[["test", "ok"], ["test", "yes"]]``` is a rank 2 tensor as the deepest level of nesting is 2.
# + [markdown] id="RaVrANK8q21q"
# ###Shape of Tensors
# Now that we've talked about the rank of tensors it's time to talk about the shape. The shape of a tensor is simply the number of elements that exist in each dimension. TensorFlow will try to determine the shape of a tensor but sometimes it may be unknown.
#
# To **get the shape** of a tensor we use the shape attribute.
#
# + id="L_NRXsFOraYa"
rank2_tensor.shape
# + [markdown] id="wVDmLJeFs086"
# ###Changing Shape
# The number of elements of a tensor is the product of the sizes of all its shapes. There are often many shapes that have the same number of elements, making it convient to be able to change the shape of a tensor.
#
# The example below shows how to change the shape of a tensor.
# + id="dZ8Rbs2xtNqj"
tensor1 = tf.ones([1,2,3]) # tf.ones() creates a shape [1,2,3] tensor full of ones
tensor2 = tf.reshape(tensor1, [2,3,1]) # reshape existing data to shape [2,3,1]
tensor3 = tf.reshape(tensor2, [3, -1]) # -1 tells the tensor to calculate the size of the dimension in that place
# this will reshape the tensor to [3,3]
# The numer of elements in the reshaped tensor MUST match the number in the original
# + [markdown] id="M631k7UDv1Wh"
# Now let's have a look at our different tensors.
# + id="IFNmUxaEv6s3"
print(tensor1)
print(tensor2)
print(tensor3)
# Notice the changes in shape
# + [markdown] id="q88pJucBolsp"
# ###Slicing Tensors
# You may be familiar with the term "slice" in python and its use on lists, tuples etc. Well the slice operator can be used on tensors to select specific axes or elements.
#
# When we slice or select elements from a tensor, we can use comma seperated values inside the set of square brackets. Each subsequent value refrences a different dimension of the tensor.
#
# Ex: ```tensor[dim1, dim2, dim3]```
#
# I've included a few examples that will hopefully help illustrate how we can manipulate tensors with the slice operator.
# + id="b0YrD-hRqD-W"
# Creating a 2D tensor
matrix = [[1,2,3,4,5],
[6,7,8,9,10],
[11,12,13,14,15],
[16,17,18,19,20]]
tensor = tf.Variable(matrix, dtype=tf.int32)
print(tf.rank(tensor))
print(tensor.shape)
# + id="Wd85uGI7qyfC"
# Now lets select some different rows and columns from our tensor
three = tensor[0,2] # selects the 3rd element from the 1st row
print(three) # -> 3
row1 = tensor[0] # selects the first row
print(row1)
column1 = tensor[:, 0] # selects the first column
print(column1)
row_2_and_4 = tensor[1::2] # selects second and fourth row
print(row2and4)
column_1_in_row_2_and_3 = tensor[1:3, 0]
print(column_1_in_row_2_and_3)
# + [markdown] id="UU4MMhB_rxvz"
# ###Types of Tensors
# Before we go to far, I will mention that there are diffent types of tensors. These are the most used and we will talk more in depth about each as they are used.
# - Variable
# - Constant
# - Placeholder
# - SparseTensor
#
# With the execption of ```Variable``` all these tensors are immuttable, meaning their value may not change during execution.
#
# For now, it is enough to understand that we use the Variable tensor when we want to potentially change the value of our tensor.
#
#
# + [markdown] id="F2OoXbe7aSVl"
# #Sources
# Most of the information is taken direclty from the TensorFlow website which can be found below.
#
# https://www.tensorflow.org/guide/tensor
|
TensorFlow/Intro.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''.venv'': venv)'
# language: python
# name: python3
# ---
# + [markdown] id="sZv_7gqAAA7t"
# ## Import Libraries
# + [markdown] id="RUqT3RIsEUOh"
# Since we will download a dataset from kaggle, we have to hand over our kaggle handle. You can find the handle in your kaggle account.
# + id="lm9TU3Nexubm"
#download kaggle api (kaggle.json) and import it here
from google.colab import files
files.upload()
# + id="SIa_ywoxfavs"
# !pip install -q tensorflow tensorflow-datasets
# + id="nCZ_Vy8YfZus"
# !pip install mlflow
# + id="gNdPBHTI_0zQ"
import os, sys
sys.path.append(os.path.dirname(os.path.realpath('/Users/paulosgidyelew/Desktop/cassava-classification-capstone/src')))
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import tensorflow_datasets as tfds
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import GlobalAveragePooling2D, Flatten, InputLayer, Dense, Dropout, BatchNormalization, Conv2D, Activation, MaxPooling2D
from tensorflow.keras.optimizers import RMSprop, Adam, SGD, Adagrad
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras.losses import SparseCategoricalCrossentropy
from keras.preprocessing.image import ImageDataGenerator
import tensorflow_hub as hub
import warnings
import mlflow
from sklearn.metrics import confusion_matrix, plot_confusion_matrix, classification_report, fbeta_score
from sklearn.utils import class_weight
import itertools, cv2
# user defined module
from src import confusion_matrix
# mlflow parameters:
EXPERIMENT_NAME = "Classava_capstone"
TRACKING_URI = "https://hudsju377cddpoevnjdkfnvpwovniewnipcdsnkvn.mlflow.neuefische.de"
warnings.filterwarnings('ignore')
RSEED = 42
tf.random.set_seed(RSEED)
# + [markdown] id="QYqFtwiTribr"
# ## Simple Convolutional Neural Network with balanced data... have to include that here.
#
# + [markdown] id="Tiom5-qIrib3"
# Now we want use the first model, but use balanced data. We chose a simple convolutional model in order to get a first glance at the results. We want to use this model as a low benchmark that we want to beat in more complex models that we will use afterwards. We were using the following tutorial as a guideline for the construction of the network: <a href= "https://www.youtube.com/watch?v=cAICT4Al5Ow&t=334s
# ">https://www.youtube.com/watch?v=cAICT4Al5Ow&t=334s</a>
#
# + [markdown] id="dCsdh_vzrib9"
# First we will set up MLflow to keep track of our experiments
# + id="YO6-BfL3rib-"
# setting the MLFlow connection and experiment
mlflow.set_tracking_uri(TRACKING_URI)
mlflow.set_experiment(EXPERIMENT_NAME)
mlflow.start_run(run_name='First, simple convolutional model')
run = mlflow.active_run()
# + [markdown] id="6AcfQa0fricA"
# Then we will create the architecture of the model. Here we are building three convolusional layers followed by one dense layers
# + id="Gq7dCw_aricC"
model = Sequential()
model.add(Conv2D(64, 3, 3, input_shape=(380, 380, 3)))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(64, 3, 3, activation='relu'))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Flatten())
model.add(Dense(64, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(5,'softmax'))
# + id="sI7oihkOricE"
model.compile(optimizer=Adam(learning_rate=0.001), loss='categorical_crossentropy',metrics=['accuracy'])
model.summary()
# + [markdown] id="l6vv3FC7ricH"
# The ImageDataGenerator is used to produce the train and validation sets.
# + id="h7UBWEXQricH"
image_data_generator = ImageDataGenerator(rescale=1./255,
rotation_range=90,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True,
vertical_flip=True,
validation_split=0.2)#,
train_set = image_data_generator.flow_from_directory('/content/train',
subset='training',
target_size=(380,380),
class_mode='categorical',
batch_size=32,
shuffle=True,
interpolation='nearest',
color_mode="rgb",
)
val_set = image_data_generator.flow_from_directory('/content/train',
subset='validation',
target_size=(380,380),
class_mode='categorical',
batch_size=32,
shuffle=False,
interpolation='nearest',
color_mode="rgb"
)
# + [markdown] id="yvSIjqkPricJ"
# We can look at the pictures and labels of one batch of the validation set:
# + id="9q8-n-h5ricJ"
#We can have a look at the images and labels in the batches
#The first [i] determines the batch number and the second [i] determines if we look at the images or its labels of this batch
val_set[1][1]
# + id="8IR3B0yMricK"
val_set[1][0]
# + [markdown] id="wA6T4fabricK"
# We can check out one instance of our set and its corresponding label:
# + id="EiwH3E9XricK"
plt.imshow(val_set[1][0][30])
print (val_set[1][1][30])
# + id="2euS6GY9ricK"
#the amount of batches in the train set are:
len(train_set)
# + [markdown] id="UF_LJUaJsL7N"
# We can include the class weights of the train and validation set, to balance out the training.
# + id="EE0Za6oUyFjs"
from collections import Counter
counter = Counter(train_set.classes)
max_val = float(max(counter.values())) #maximum value
class_weights = {class_id : max_val/num_images for class_id, num_images in counter.items()}
# + id="2ZF1fd5xvUVF"
#class_weights = class_weight.compute_class_weight(class_weight='balanced', classes=np.unique(train_set.classes), train_set.classes)
# + id="hMzTduzlricM"
model_checkpoint_filepath = 'checkpoints/simple_conv_model_balanced.ckpt'#h5
model_check_point = ModelCheckpoint(model_checkpoint_filepath,
verbose=1,
save_weights_only=True,
monitor='val_loss',
save_best_only=True,
mode='auto')
# the train_set contains both the pictures and the labels, so we do not have to define them separately
history = model.fit(train_set,
epochs=10,
verbose=1,
callbacks=[model_check_point],
validation_data=val_set,
steps_per_epoch=len(train_set),
validation_steps=len(val_set),
class_weight=class_weights
)
# + [markdown] id="oDbLBZ9-ricN"
# Let us plot the training-process.
# + id="IGro28HLricN"
plt.plot(history.history['accuracy'])
plt.plot(history.history['val_accuracy'])
plt.title('Model Accuracy')
plt.xlabel('epoch')
plt.ylabel('Accuracy')
plt.legend(['training','validation'], loc='lower right')
plt.show()
# + id="f8_scJwOricN"
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Loss Function')
plt.xlabel('epoch')
plt.ylabel('Loss')
plt.legend(['loss','val_loss'], loc='upper right')
plt.show()
# + [markdown] id="vN2H5-1UricN"
# ### Construction of the confusion matrix
# + id="5H8EJTgfricN"
#we can use model.predict to predict the validation set and argmax gives us the the highest number for each element
results = model.predict(val_set)
results = np.argmax(results, axis=1)
# + id="IOyW5NY6ricN"
#report = classification_report(list_of_true_labels,results)
report = classification_report(val_set.classes, results)
print (report)
# + id="-2ophpxwricO"
cm = confusion_matrix(val_set.classes, results)
confusion_matrix.plot_confusion_matrix(
cm, classes=['CBB', 'CBSD','CGM','CMD','Healthy'],
title='Pre-trained'
)
# + [markdown] id="CqvwwqhTricO"
# Calculation of the F2 score (description can be found in the simple model chapter)
# + id="dTkXuJodricO"
#Due to imbalance in our dataset we have to use 'macro' for averaging
F2_score = fbeta_score(val_set.classes,results, average='macro', beta=2)
print(F2_score)
# + [markdown] id="RtP4K56mricO"
# Now let us save the parameters of the model to MLflow:
# + id="K05Vly5-ricP"
#These are the parameters that will be transfered to MlFlow for logging our experiments
#Find meaningful parameters!
params = {
"number of epochs": 10,
"input_shape": val_set[0][0][0].shape,
"confusion matrix":cm
}
# + id="nRWu2wwRricP"
#logging params to mlflow
mlflow.log_params(params)
#setting tags
mlflow.set_tag("colab", "True")
#logging metrics
mlflow.log_metric("train-" + "accuracy", history.history['accuracy'][-1])
mlflow.log_metric("val-" + "accuracy", history.history['val_accuracy'][-1])
mlflow.log_metric("train-" + "loss", history.history['loss'][-1])
mlflow.log_metric("val-" + "loss", history.history['val_loss'][-1])
mlflow.log_metric("F2-score", F2_score)
# logging the model to mlflow will not work without a AWS Connection setup.. too complex for now
# but possible if running mlflow locally
# mlflow.log_artifact("../models")
# mlflow.sklearn.log_model(reg, "model")
mlflow.end_run()
# + [markdown] id="or8wawKb6WJz"
# Using the weighted classes for the fit did not deliver good results. Oversamplling of the data would be needed.
|
notebooks/CNN_Balanced_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import rgb2hex
# %matplotlib inline
# -
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score, confusion_matrix, classification_report, roc_auc_score
from keras.models import Model, load_model
from keras import initializers, regularizers, constraints
from keras.regularizers import l2
from keras.layers import Dense, Embedding, Input, concatenate, Flatten, Layer
from keras.layers import LSTM, Bidirectional, GlobalMaxPool1D, Dropout, Conv1D, MaxPool1D, BatchNormalization, TimeDistributed
from keras.preprocessing import text, sequence
from keras.callbacks import EarlyStopping, ModelCheckpoint
from keras.optimizers import SGD, Adam
from keras import backend as K
from IPython.display import SVG
from keras.utils.vis_utils import model_to_dot
from gensim.models import Word2Vec
import os, re, pickle
import spacy
from spacy.tokens.doc import Doc
# ### Import Data
train = pd.read_csv('data/labeledTrainData.tsv', sep='\t')
print(train.shape)
test = pd.read_csv('data/testData.tsv', sep='\t')
print(test.shape)
# ### Pre-process Data
MAX_FEATURES = 25000
MAX_LEN = 350
list_sentences_train = train['review'].fillna("UNKNOWN").values.tolist()
list_sentences_test = test['review'].fillna("UNKNOWN").values.tolist()
tokenizer = text.Tokenizer(num_words=MAX_FEATURES)
tokenizer.fit_on_texts(list_sentences_train)
list_tokenized_train = tokenizer.texts_to_sequences(list_sentences_train)
list_tokenized_test = tokenizer.texts_to_sequences(list_sentences_test)
X = sequence.pad_sequences(list_tokenized_train, maxlen=MAX_LEN)
X_test = sequence.pad_sequences(list_tokenized_test, maxlen=MAX_LEN)
y = train['sentiment'].values.reshape(-1,1)
y_softmax = np.array([np.array([0,1]) if x == 1 else np.array([1,0]) for x in y])
N_CLASSES = 2
X_train, X_val, y_train, y_val = train_test_split(X, y_softmax, test_size=0.1, random_state=42)
# ### Create Model - No External Knowledge
# +
EMBED_SIZE = 8
CNN_FILTER_SIZE = 8
CNN_KERNEL_SIZE = 3
def create_model():
input_sequence = Input(shape=(MAX_LEN, ))
x = Embedding(input_dim=MAX_FEATURES, output_dim=EMBED_SIZE)(input_sequence)
x = Dropout(0.5)(x)
x = Conv1D(filters=CNN_FILTER_SIZE, kernel_size=CNN_KERNEL_SIZE, padding='same', kernel_regularizer=l2(0.0001))(x)
#x = Bidirectional(LSTM(32,
# return_sequences=True,
# kernel_regularizer=l2(0.0001)))(x)
#x = GlobalMaxPool1D()(x)
#x = AttentionWithContext()(x)
x = TimeDistributed(Dense(1, activation="elu", kernel_regularizer=l2(0.0001)))(x)
x = Flatten()(x)
x = BatchNormalization()(x)
x = Dense(8, activation="elu", kernel_regularizer=l2(0.0001))(x)
prediction = Dense(N_CLASSES, activation="softmax")(x)
opt = Adam(lr=0.0001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
#opt = SGD(lr=0.001, momentum=0.0, decay=0.0, nesterov=False)
model = Model(inputs=input_sequence, outputs=prediction)
model.compile(loss='binary_crossentropy',
optimizer=opt,
metrics=['accuracy'])
return model
# +
model = create_model()
BATCH_SIZE = 512
EPOCHS = 50
# -
FILE_PATH = "models/keras_model_weights.hdf5"
checkpoint = ModelCheckpoint(FILE_PATH, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
early = EarlyStopping(monitor="val_loss", mode="min", patience=15)
callbacks_list = [checkpoint, early]
model.summary()
SVG(model_to_dot(model, show_shapes=True).create(prog='dot', format='svg'))
model.fit(X_train, y_train, batch_size=BATCH_SIZE, epochs=EPOCHS, validation_data=[X_val, y_val], callbacks=callbacks_list)
pd.DataFrame(model.history.history).plot(figsize=(12,8))
model = load_model(filepath=FILE_PATH, custom_objects={'AttentionWithContext': AttentionWithContext})
y_val_hat = model.predict(X_val)
print(accuracy_score(y_val, y_val_hat > 0.5))
print(roc_auc_score(y_val, y_val_hat))
print(confusion_matrix(y_val, y_val_hat > 0.5))
print(classification_report(y_val, y_val_hat > 0.5))
# Useful tutorials:
# * http://konukoii.com/blog/2018/02/19/twitter-sentiment-analysis-using-combined-lstm-cnn-models/
# #### Extract Activations
from IPython.display import display, HTML
def create_get_activation_function(model, output_layer_int):
inp = model.input
output = model.layers[output_layer_int].output
get_activations = K.function([inp]+ [K.learning_phase()], [output])
return get_activations
act_model = load_model(filepath=FILE_PATH, custom_objects={'AttentionWithContext': AttentionWithContext})
get_activations = create_get_activation_function(act_model, 3)
word_to_hash = tokenizer.word_index
hash_to_word = {v:k for k,v in word_to_hash.items()}
hash_to_word[0] = ''
cmap = plt.cm.get_cmap('RdYlGn')
# +
example = 4505
html_string = '<p>For training example: ' + str(example) + '</p>'
for node in range(CNN_FILTER_SIZE):
activations = get_activations([[X_train[example]], 0.])[0]
text = [hash_to_word[x] for x in X_train[example]]
scaled_activations = activations[0,:,node] - activations[0,:,node].min()
scaled_activations = scaled_activations / scaled_activations.max()
scaled_activations = pd.rolling_mean(scaled_activations, CNN_KERNEL_SIZE, min_periods=1)
new_string = ''
for i, t in enumerate(text):
new_string += '<span style="background-color: ' + str(rgb2hex(cmap(scaled_activations[i]))) + '">' + t + '</span>' + ' '
html_string += '<p>CNN Filter: ' + str(node) + '</p><p>' + new_string + '</p>'
h = HTML(html_string)
display(h)
# -
get_word_activations = create_get_activation_function(act_model, 5)
# +
example = 4505
html_string = '<p>For training example: ' + str(example) + '</p>'
activations = get_word_activations([[X_train[example]], 0.])[0]
text = [hash_to_word[x] for x in X_train[example]]
scaled_activations = activations[0,:] - activations[0,:].min()
scaled_activations = scaled_activations / scaled_activations.max()
new_string = ''
for i, t in enumerate(text):
new_string += '<span style="background-color: ' + str(rgb2hex(cmap(scaled_activations[i]))) + '">' + t + '</span>' + ' '
html_string += '<p>Time Distributed Dense Output: <p>' + new_string + '</p>'
h = HTML(html_string)
display(h)
# -
# #### Word Embeddings
from scipy.spatial.distance import pdist, squareform
emb_layer = model.layers[1]
emb_layer_weights = emb_layer.get_weights()[0]
emb_layer_weights.shape
x_sq = squareform(pdist(emb_layer_weights[0:10000,:], metric='cosine'))
df_x_sq = pd.DataFrame(x_sq)
df_x_edge = df_x_sq.where(np.triu(np.ones(df_x_sq.shape)).astype(np.bool)).stack().reset_index()
df_x_edge.columns = ['source','target','weight']
df_x_edge['weight'].hist(bins=50)
df_x_edge = df_x_edge[df_x_edge['weight'] < 0.1]
df_x_edge = df_x_edge[df_x_edge.source != df_x_edge.target]
df_x_edge.shape
df_x_edge['source_word'] = df_x_edge['source'].apply(lambda x: hash_to_word[x])
df_x_edge['target_word'] = df_x_edge['target'].apply(lambda x: hash_to_word[x])
df_x_edge.sort_values(by='weight')
df_x_edge.to_csv('../data/combine_activation_sim.csv', index=False)
df_node_text = pd.DataFrame(df['text'], columns=['text'])
df_node_text['Id'] = df_node_text.index
df_node_text = df_node_text[['Id', 'text']]
from IPython.core.display import display, HTML
from string import Template
import json, random
# +
random.seed(42)
n_nodes = 40
n_edges = 200
graph_data = { 'nodes': [], 'edges': [] }
for i in range(n_nodes):
graph_data['nodes'].append({
"id": "n" + str(i),
"label": "n" + str(i),
"x": random.uniform(0,1),
"y": random.uniform(0,1),
"size": random.uniform(0.2,1)
})
for j in range(n_edges):
x_center = random.uniform(0,1)
y_center = random.uniform(0,1)
x_dist = random.uniform(0.1,0.5)
y_dist = random.uniform(0.2,0.5)
neighborhood = []
for node in graph_data['nodes']:
if abs(node['x'] - x_center) < x_dist:
if abs(node['y'] - y_center) < y_dist:
neighborhood.append(int(node['id'].replace('n','')))
if len(neighborhood) >= 2:
ends = random.sample(neighborhood,2)
graph_data['edges'].append({
"id": "e" + str(j),
"source": "n" + str(ends[0]),
"target": "n" + str(ends[1])
})
# -
js_text_template = Template('''
var g = $graph_data ;
s = new sigma({graph: g, container: '$container', settings: { defaultNodeColor: '#ec5148'} });
s.graph.nodes().forEach(function(n) {
n.originalColor = n.color;
});
s.graph.edges().forEach(function(e) {
e.originalColor = e.color;
});
s.bind('clickNode', function(e) {
var nodeId = e.data.node.id,
toKeep = s.graph.neighbors(nodeId);
toKeep[nodeId] = e.data.node;
s.graph.nodes().forEach(function(n) {
if (toKeep[n.id])
n.color = n.originalColor;
else
n.color = '#eee';
});
s.graph.edges().forEach(function(e) {
if (toKeep[e.source] && toKeep[e.target])
e.color = e.originalColor;
else
e.color = '#eee';
});
s.refresh();
});
s.bind('clickStage', function(e) {
s.graph.nodes().forEach(function(n) {
n.color = n.originalColor;
});
s.graph.edges().forEach(function(e) {
e.color = e.originalColor;
});
s.refresh();
});
''')
js_text = js_text_template.substitute({'graph_data': json.dumps(graph_data),
'container': 'graph-div'})
'../ml-notebooks/js/sigma.min.js'
html_template = Template('''
<script src="../ml-notebooks/js/sigma.min.js"></script>
<div id="graph-div" style="height:800px"></div>
<script> $js_text </script>
''')
HTML(html_template.substitute({'js_text': js_text}))
# ### Create Model - Use Pretrained Embeddings, PoS Parsing
# #### Prep Data
MAX_FEATURES = 25000
MAX_LEN = 350
list_sentences_train = train['review'].fillna("UNKNOWN").values.tolist()
list_sentences_test = test['review'].fillna("UNKNOWN").values.tolist()
list_sentences_train_parsed = [transform_doc(x, MAX_LEN=1000) for x in list_sentences_train]
list_sentences_test_parsed = [transform_doc(x, MAX_LEN=1000) for x in list_sentences_test]
# +
with open('data/list_sentences_train_parsed.pkl', 'wb') as f:
pickle.dump(list_sentences_train_parsed, f)
with open('data/list_sentences_test_parsed.pkl', 'wb') as f:
pickle.dump(list_sentences_test_parsed, f)
# -
tokenizer = text.Tokenizer(num_words=MAX_FEATURES, filters='!"#$%&()*+,-/:;<=>?@[\\]^`{}~\t\n', lower=False)
tokenizer.fit_on_texts(list_sentences_train_parsed)
list_tokenized_train = tokenizer.texts_to_sequences(list_sentences_train_parsed)
list_tokenized_test = tokenizer.texts_to_sequences(list_sentences_test_parsed)
X_train = sequence.pad_sequences(list_tokenized_train, maxlen=MAX_LEN)
X_test = sequence.pad_sequences(list_tokenized_test, maxlen=MAX_LEN)
y = train['sentiment'].values.reshape(-1,1)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.1, random_state=42)
# #### Inspect
list_sentences_train_parsed[0]
list_tokenized_train[0]
index_word_dict = {v:k for k, v in tokenizer.word_index.items()}
word_index_dict = tokenizer.word_index
for w in list_tokenized_train[0]:
print(index_word_dict[w])
# #### Create Embedding Matrix
# (or load one if you have it)
w2v_model = Word2Vec.load('models/w2v_model_32_plaintext')
# +
EMBED_SIZE = w2v_model.vector_size
print('The size of the gensim word2vec vocab is: {}'.format(len(w2v_model.wv.vocab.items())))
unknown_word_count = 0
def choose_embedded_vector(word, unknown_word_count, verbose=False):
if word in w2v_model.wv.vocab:
return w2v_model.wv.word_vec(word), unknown_word_count
else:
if verbose:
print('Unknown word: {}'.format(word))
return np.random.rand(EMBED_SIZE), (unknown_word_count+1)
index_word_dict = {v:k for k, v in tokenizer.word_index.items()}
word_index_dict = tokenizer.word_index
num_words = tokenizer.num_words + 1
print('The size of the keras token vocab is: {}'.format(len(index_word_dict)))
print('The tokenizer vocab is limited to: {}'.format(tokenizer.num_words))
embedding_weights = np.zeros((num_words, EMBED_SIZE))
for word, index in word_index_dict.items():
if index < num_words:
embedding_weights[index,:], unknown_word_count = choose_embedded_vector(word, unknown_word_count)
print('Total amount of words not found in gensim word2vec model: {}'.format(unknown_word_count))
print('Embedding matrix shape: {}'.format(embedding_weights.shape))
# -
EMBED_SIZE
# #### Train Model
# +
CNN_FILTER_SIZE = 32
CNN_KERNEL_SIZE = 3
def create_model():
input_sequence = Input(shape=(MAX_LEN, ))
x = Embedding(input_dim=num_words, output_dim=EMBED_SIZE, input_length=MAX_LEN, mask_zero=False, weights=[embedding_weights], trainable=True)(input_sequence)
x = Dropout(0.5)(x)
x = Conv1D(filters=CNN_FILTER_SIZE, kernel_size=CNN_KERNEL_SIZE, padding='same', kernel_regularizer=l2(0.0001))(x)
#x = Bidirectional(LSTM(32,
# return_sequences=True,
# kernel_regularizer=l2(0.0001)))(x)
#x = GlobalMaxPool1D()(x)
x = AttentionWithContext()(x)
x = BatchNormalization()(x)
x = Dense(32, activation="elu", kernel_regularizer=l2(0.0001))(x)
prediction = Dense(N_CLASSES, activation="sigmoid")(x)
opt = Adam(lr=0.0001, beta_1=0.9, beta_2=0.999, epsilon=None, decay=0.0, amsgrad=False)
#opt = SGD(lr=0.001, momentum=0.0, decay=0.0, nesterov=False)
model = Model(inputs=input_sequence, outputs=prediction)
model.compile(loss='binary_crossentropy',
optimizer=opt,
metrics=['accuracy'])
return model
# +
model = create_model()
BATCH_SIZE = 512
EPOCHS = 50
# -
FILE_PATH = "models/keras_model_weights.hdf5"
checkpoint = ModelCheckpoint(FILE_PATH, monitor='val_loss', verbose=1, save_best_only=True, mode='min')
early = EarlyStopping(monitor="val_loss", mode="min", patience=15)
callbacks_list = [checkpoint, early]
model.summary()
model.fit(X_train, y_train, batch_size=BATCH_SIZE, epochs=EPOCHS, validation_data=[X_val, y_val], callbacks=callbacks_list)
pd.DataFrame(model.history.history).plot(figsize=(12,8))
model = load_model(filepath=FILE_PATH, custom_objects={'AttentionWithContext':AttentionWithContext})
y_hat = model.predict(X_val)
print(accuracy_score(y_val, y_hat > 0.5))
print(roc_auc_score(y_val, y_hat))
print(confusion_matrix(y_val, y_hat > 0.5))
print(classification_report(y_val, y_hat > 0.5))
# ### Helper Functions
# A lot of the spacy code was pulled from examples: https://github.com/explosion
# +
nlp = spacy.load('en_core_web_sm')
LABELS = {
'ENT': 'ENT',
'PERSON': 'ENT',
'NORP': 'ENT',
'FAC': 'ENT',
'ORG': 'ENT',
'GPE': 'ENT',
'LOC': 'ENT',
'LAW': 'ENT',
'PRODUCT': 'ENT',
'EVENT': 'ENT',
'WORK_OF_ART': 'ENT',
'LANGUAGE': 'ENT',
'DATE': 'DATE',
'TIME': 'TIME',
'PERCENT': 'PERCENT',
'MONEY': 'MONEY',
'QUANTITY': 'QUANTITY',
'ORDINAL': 'ORDINAL',
'CARDINAL': 'CARDINAL'
}
pre_format_re = re.compile(r'^[\`\*\~]')
post_format_re = re.compile(r'[\`\*\~]$')
url_re = re.compile(r'\[([^]]+)\]\(%%URL\)')
link_re = re.compile(r'\[([^]]+)\]\(https?://[^\)]+\)')
def strip_meta(text):
if type(text) == str:
text = link_re.sub(r'\1', text)
text = text.replace('>', '>').replace('<', '<')
text = pre_format_re.sub('', text)
text = post_format_re.sub('', text)
return text
else:
return ''
def represent_word(word):
if word.like_url:
return '%%URL|X'
text = re.sub(r'\s', '_', word.text)
tag = LABELS.get(word.ent_type_, word.pos_)
if not tag:
tag = '?'
return text + '|' + tag
def merge_clean_sentence(nlp, text, collapse_punctuation=True, collapse_phrases=True):
doc = nlp(text)
if collapse_punctuation:
spans = []
for word in doc[:-1]:
if word.is_punct:
continue
if not word.nbor(1).is_punct:
continue
start = word.i
end = word.i + 1
while end < len(doc) and doc[end].is_punct:
end += 1
span = doc[start : end]
spans.append(
(span.start_char, span.end_char,
{'tag': word.tag_, 'lemma': word.lemma_, 'ent_type': word.ent_type_})
)
for start, end, attrs in spans:
doc.merge(start, end, **attrs)
if collapse_phrases:
for np in list(doc.noun_chunks):
np.merge(tag=np.root.tag_, lemma=np.root.lemma_, ent_type=np.root.ent_type_)
return doc
def transform_doc(text, MAX_LEN):
d = merge_clean_sentence(nlp, text, collapse_punctuation=False, collapse_phrases=True)
strings = []
for sent in d.sents:
if sent.text.strip():
for w in sent:
if not w.is_space:
strings.append(represent_word(w))
if strings:
return ' '.join(strings[0:MAX_LEN])
else:
return ' '.join(['' for x in range(MAX_LEN)])
# -
# Attention adapted from: https://gist.github.com/cbaziotis/6428df359af27d58078ca5ed9792bd6d
# +
def dot_product(x, kernel):
"""
Wrapper for dot product operation, in order to be compatible with both
Theano and Tensorflow
Args:
x (): input
kernel (): weights
Returns:
"""
if K.backend() == 'tensorflow':
return K.squeeze(K.dot(x, K.expand_dims(kernel)), axis=-1)
else:
return K.dot(x, kernel)
class AttentionWithContext(Layer):
"""
Attention operation, with a context/query vector, for temporal data.
Supports Masking.
Follows the work of Yang et al. [https://www.cs.cmu.edu/~diyiy/docs/naacl16.pdf]
"Hierarchical Attention Networks for Document Classification"
by using a context vector to assist the attention
# Input shape
3D tensor with shape: `(samples, steps, features)`.
# Output shape
2D tensor with shape: `(samples, features)`.
How to use:
Just put it on top of an RNN Layer (GRU/LSTM/SimpleRNN) with return_sequences=True.
The dimensions are inferred based on the output shape of the RNN.
Note: The layer has been tested with Keras 2.0.6
Example:
model.add(LSTM(64, return_sequences=True))
model.add(AttentionWithContext())
# next add a Dense layer (for classification/regression) or whatever...
"""
def __init__(self,
W_regularizer=None, u_regularizer=None, b_regularizer=None,
W_constraint=None, u_constraint=None, b_constraint=None,
bias=True, **kwargs):
self.supports_masking = True
self.init = initializers.get('glorot_uniform')
self.W_regularizer = regularizers.get(W_regularizer)
self.u_regularizer = regularizers.get(u_regularizer)
self.b_regularizer = regularizers.get(b_regularizer)
self.W_constraint = constraints.get(W_constraint)
self.u_constraint = constraints.get(u_constraint)
self.b_constraint = constraints.get(b_constraint)
self.bias = bias
super(AttentionWithContext, self).__init__(**kwargs)
def build(self, input_shape):
assert len(input_shape) == 3
self.W = self.add_weight((input_shape[-1], input_shape[-1],),
initializer=self.init,
name='{}_W'.format(self.name),
regularizer=self.W_regularizer,
constraint=self.W_constraint)
if self.bias:
self.b = self.add_weight((input_shape[-1],),
initializer='zero',
name='{}_b'.format(self.name),
regularizer=self.b_regularizer,
constraint=self.b_constraint)
self.u = self.add_weight((input_shape[-1],),
initializer=self.init,
name='{}_u'.format(self.name),
regularizer=self.u_regularizer,
constraint=self.u_constraint)
super(AttentionWithContext, self).build(input_shape)
def compute_mask(self, input, input_mask=None):
# do not pass the mask to the next layers
return None
def call(self, x, mask=None):
uit = dot_product(x, self.W)
if self.bias:
uit += self.b
uit = K.tanh(uit)
ait = dot_product(uit, self.u)
a = K.exp(ait)
# apply mask after the exp. will be re-normalized next
if mask is not None:
# Cast the mask to floatX to avoid float64 upcasting in theano
a *= K.cast(mask, K.floatx())
# in some cases especially in the early stages of training the sum may be almost zero
# and this results in NaN's. A workaround is to add a very small positive number ε to the sum.
# a /= K.cast(K.sum(a, axis=1, keepdims=True), K.floatx())
a /= K.cast(K.sum(a, axis=1, keepdims=True) + K.epsilon(), K.floatx())
a = K.expand_dims(a)
weighted_input = x * a
return K.sum(weighted_input, axis=1)
def compute_output_shape(self, input_shape):
return input_shape[0], input_shape[-1]
# -
|
03_sequential_models.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/falconlee236/DeepLearningFrom_Scratch/blob/main/Chapter_2.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Mcqk8Lv3Eq8L"
# # **Chapter 2 Perceptron**
# + [markdown] id="6lEUjjw1E466"
# **2.3 Perceptron implementing**
# + [markdown] id="shfEFIhMGdPC"
# *2.3.1 from simple implementation*
# + id="v6mNJIGCGl56"
def AND(x1, x2):
w1, w2, theta = 0.5, 0.5, 0.7
tmp = x1 * w1 + x2 * w2
return 1 if tmp > theta else 0
# + id="vWYlP_e9Ik76"
AND(0, 0)
# + id="DlGWzaNgI55s"
AND(1, 0)
# + id="UmGiU2R_I8Nr"
AND(0, 1)
# + id="pPA3RlUZI9Cs"
AND(1, 1)
# + [markdown] id="Hj-66kRjI9xD"
# *2.3.2 introducing weight and bias* <br>
#
#
# ---
#
#
# from w1*x1 + w2*x2 > theta <br>
# to w1*x1 + w2*x2 + bias > 0 (bias = -theta)
# + id="yTM5oMJwKLJc"
import numpy as np
x = np.array([0, 1]) # input
w = np.array([0.5, 0.5]) # weight
b = -0.7 # bias
w * x
# + id="d2q1W6VoKc5e"
np.sum(w * x)
# + id="jdicyZhBKg6-"
np.sum(w * x) + b
# + [markdown] id="lgMzCNBmKpQ3"
# *2.3.3 implementing weight and bias*
# + id="znz9N8dvKz0r"
def AND(x1, x2):
x = np.array([x1, x2])
w = np.array([0.5, 0.5])
b = -0.7
return 1 if np.sum(x * w) + b > 0 else 0
# + id="SorkO_emLITH"
def NAND(x1, x2):
x = np.array([x1, x2])
w = np.array([-0.5, -0.5])
b = 0.7 # different only weight(w and b)
return 1 if np.sum(x * w) + b > 0 else 0
def OR(x1, x2):
x = np.array([x1, x2])
w = np.array([0.5, 0.5])
b = -0.2 # different only weight(w and b)
return 1 if np.sum(x * w) + b > 0 else 0
# + [markdown] id="N5kLkcVrMXq7"
# **2.5 IF Coming multi-layer perceptron**
# + [markdown] id="8MABLFTiPFDR"
# *2.5.2 implementing XOR gate*
# + id="2RQXl6-pPL7Z"
def XOR(x1, x2):
s1 = NAND(x1, x2)
s2 = OR(x1, x2)
return AND(s1, s2)
# + id="G1Z5PQW_PYhS"
XOR(0, 0)
# + id="WW2WT1lAPamG"
XOR(1, 0)
# + id="wQhr9XYnPeX-"
XOR(0, 1)
# + id="nQcr3s2UPjQr"
XOR(1, 1)
|
Book_1/ch02/Chapter_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import scda
import os
import shutil
import numpy as np
import matplotlib.pyplot as plt
# %pylab inline --no-import-all
matplotlib.rcParams['image.origin'] = 'lower'
matplotlib.rcParams['image.interpolation'] = 'nearest'
matplotlib.rcParams['image.cmap'] = 'gray'
import logging
scda.configure_log()
# -
# # Prepare two SCDA designs, with and without LS alignment tolerance
# ## Set the design parameters
pupil_params = {'N': 125, 'prim':'hex3', 'centobs':True, 'secobs':'X', 'thick':'025'}
fpm_params = {'rad': 4.}
ls_params_aligntol = {'obscure':0, 'id':25, 'od':80, 'aligntol':5, 'aligntolcon':2.9}
ls_params_noaligntol = {'obscure':0, 'id':25, 'od':80, 'aligntol':None}
image_params = {'c': 10., 'ida':3.5, 'oda':8., 'bw':0.10, 'Nlam':3}
work_dir = "/astro/opticslab1/SCDA/Scripts/AMPL/LDZ_test" # where to write the AMPL source code
if not os.path.exists(work_dir):
os.mkdir(work_dir)
input_dir = "/astro/opticslab1/SCDA/Apertures/InputMasks" # location of input TelAp, FPM, and LS arrays
TelAp_dir = os.path.join(input_dir, "TelAp")
FPM_dir = os.path.join(input_dir, "FPM")
LS_dir = os.path.join(input_dir, "LS")
design_params_aligntol = {'Pupil': pupil_params, 'FPM': fpm_params,
'LS': ls_params_aligntol, 'Image': image_params}
design_params_noaligntol = {'Pupil': pupil_params, 'FPM': fpm_params,
'LS': ls_params_noaligntol, 'Image': image_params}
fileorg = {'work dir': work_dir, 'TelAp dir': TelAp_dir,
'FPM dir': FPM_dir, 'LS dir': LS_dir}
bar_solver = {'method': 'bar'}
barhom_solver = {'method': 'barhom'}
# ## Initiate the coronagraph objects
hexap_coron_aligntol = scda.QuarterplaneAPLC(design=design_params_aligntol, fileorg=fileorg, solver=barhom_solver)
hexap_coron_noaligntol = scda.QuarterplaneAPLC(design=design_params_noaligntol, fileorg=fileorg, solver=bar_solver)
# ## Show the file organization for both coronagraphs
hexap_coron_aligntol.fileorg
hexap_coron_noaligntol.fileorg
# ### Check the status of input files needed to run the AMPL program
hexap_coron_aligntol.check_ampl_input_files()
print("All the input files for AMPL are in place? {0:}".format(hexap_coron_aligntol.ampl_infile_status))
hexap_coron_noaligntol.check_ampl_input_files()
print("All the input files for AMPL are in place? {0:}".format(hexap_coron_noaligntol.ampl_infile_status))
# ## Write the AMPL source file
hexap_coron_aligntol.write_ampl(override_infile_status=True, overwrite=True)
hexap_coron_noaligntol.write_ampl(override_infile_status=True, overwrite=True)
# ## Write serial bash execution script
bash_fname = os.path.join(hexap_coron_aligntol.fileorg['ampl src dir'], 'run_LDZ_test.sh')
bash_fobj = open(bash_fname, "w")
bash_fobj.write("#! /bin/bash -x\n")
bash_fobj.write("ampl {0:s}\n".format(hexap_coron_noaligntol.fileorg['ampl src fname']))
bash_fobj.write("ampl {0:s}\n".format(hexap_coron_aligntol.fileorg['ampl src fname']))
bash_fobj.close()
# ## Create a bundled source + input file subdirectory for both designs
bundled_dir = "/astro/opticslab1/SCDA/Scripts/AMPL/hex3_LStol25"
bundled_coron_list = scda.make_ampl_bundle([hexap_coron_noaligntol, hexap_coron_aligntol], bundled_dir)
os.listdir(bundled_dir)
# ## Evaluate basic coronagraph metrics
hexap_coron_noaligntol.get_metrics()
hexap_coron_aligntol.get_metrics()
# # Full-plane on-axis PSF evaluation
# +
TelAp_qp = np.loadtxt(hexap_coron_noaligntol.fileorg['TelAp fname'])
TelAp = np.concatenate((np.concatenate((TelAp_qp[::-1,::-1], TelAp_qp[:,::-1]),axis=0),
np.concatenate((TelAp_qp[::-1,:], TelAp_qp),axis=0)), axis=1)
FPM_qp = np.loadtxt(hexap_coron_noaligntol.fileorg['FPM fname'])
FPM = np.concatenate((np.concatenate((FPM_qp[::-1,::-1], FPM_qp[:,::-1]),axis=0),
np.concatenate((FPM_qp[::-1,:], FPM_qp),axis=0)), axis=1)
LS_qp = np.loadtxt(hexap_coron_noaligntol.fileorg['LS fname'])
LS = np.concatenate((np.concatenate((LS_qp[::-1,::-1], LS_qp[:,::-1]),axis=0),
np.concatenate((LS_qp[::-1,:], LS_qp),axis=0)), axis=1)
LDZ_qp = np.loadtxt(hexap_coron_aligntol.fileorg['LDZ fname'])
LDZ = np.concatenate((np.concatenate((LDZ_qp[::-1,::-1], LDZ_qp[:,::-1]),axis=0),
np.concatenate((LDZ_qp[::-1,:], LDZ_qp),axis=0)), axis=1)
An_col = np.loadtxt(hexap_coron_noaligntol.fileorg['sol fname'])[:,-1]
An_qp = An_col.reshape(TelAp_qp.shape)
An = np.concatenate((np.concatenate((An_qp[::-1,::-1], An_qp[:,::-1]),axis=0),
np.concatenate((An_qp[::-1,:], An_qp),axis=0)), axis=1)
#At_col = np.loadtxt(hexap_coron_aligntol.fileorg['sol fname'])[:,-1]
At_qp = At_col.reshape(TelAp_qp.shape)
#At = np.concatenate((np.concatenate((At_qp[::-1,::-1], At_qp[:,::-1]),axis=0),
# np.concatenate((At_qp[::-1,:], At_qp),axis=0)), axis=1)
plt.figure(figsize=(16,8))
plt.subplot(121)
plt.imshow(An*TelAp)
plt.title('Apodizer, no tol.')
plt.subplot(122)
plt.imshow(At*TelAp)
plt.title('Apodizer, with tol.')
# -
# ## Create translated Lyot stop, check against design tolerance
# +
test_shift = (0,1)
max_shift_tol = LS.shape[0]*float(hexap_coron_aligntol.design['LS']['aligntol'])/1000
max_shift_tol_int = int(np.floor(max_shift_tol))
print("The LDZ accomomdates a translation {0:.1f}% of D={1:d} pixels = {2:.2f} pixels, up to {3:d} whole pixels".format(
float(hexap_coron_aligntol.design['LS']['aligntol'])/10, LS.shape[0], max_shift_tol, max_shift_tol_int))
print("Testing an (x,y) translation of {0:} pixels. Beyond the design tolerance? {1:}".format(
test_shift, test_shift[0]**2 + test_shift[1]**2 > max_shift_tol))
LSe = np.roll(np.roll(LS, test_shift[0], axis=1), test_shift[1], axis=0)
LS_err_mask = np.ceil(np.abs(LSe - LS)).astype(bool)
print("LDZ encompasses the LS transmission error region? {0:}".format(
~np.any(np.logical_and(LS_err_mask, ~LDZ.astype(bool)))))
print("Total unconstrained \"leak\" area after translation = {0:d} pixels".format(
int(np.sum(np.logical_and(LS_err_mask, ~LDZ.astype(bool))))))
# -
plt.figure(figsize=(16,6))
plt.subplot(131)
plt.imshow(LSe - LS)
lims = plt.axis('off')
t=plt.title('Change in Lyot stop transmission profile')
plt.subplot(132)
plt.imshow(~LDZ.astype(bool))
lims = plt.axis('off')
t=plt.title('Inverse of LDZ mask')
plt.subplot(133)
plt.imshow(np.logical_and(LS_err_mask, ~LDZ.astype(bool)))
lims = plt.axis('off')
t=plt.title('Lyot leak region (black is good)')
# ## Define coordinates and dimensions
# +
D = 1.
N = hexap_coron_aligntol.design['Pupil']['N']
bw = hexap_coron_aligntol.design['Image']['bw']
Nlambda = hexap_coron_aligntol.design['Image']['Nlam']
M_fp1 = hexap_coron_aligntol.design['FPM']['M']
fpm_rad = hexap_coron_aligntol.design['FPM']['rad']
rho2 = hexap_coron_aligntol.design['Image']['oda'] + 1.
fp2res = 8.
M_fp2 = int(np.ceil(rho2*fp2res))
# pupil plane
dx = (D/2)/N
dy = dx
xs = np.matrix(np.linspace(-N+0.5,N-0.5,2*N)*dx)
ys = xs
# FPM
dmx = fpm_rad/M_fp1
dmy = dmx
mxs = np.matrix(np.linspace(-M_fp1+0.5,M_fp1-0.5,2*M_fp1)*dmx)
mys = mxs
# FP2
dxi = 1/fp2res
xis = np.matrix(np.linspace(-M_fp2+0.5,M_fp2-0.5,2*M_fp2)*dxi)
etas = xis
# wavelength ratios
wrs = np.linspace(1.-bw/2, 1.+bw/2, Nlambda)
# +
#wrs = [0.95, 1., 1.02]
# -
# ## Fourier propagation
# +
def get_onax_aplc_psf(TelAp, A, FPM, LS, xs, dx, mxs, dmx, xis, dxi, wrs):
intens_D_polychrom = []
for wr in wrs:
Psi_B = dx*dx/wr*np.dot(np.dot(np.exp(-1j*2*np.pi/wr*np.dot(mxs.T, xs)), TelAp*A ),
np.exp(-1j*2*np.pi/wr*np.dot(xs.T, mxs)))
Psi_B_stop = np.multiply(Psi_B, FPM)
Psi_C = A*TelAp - dmx*dmx/wr*np.dot(np.dot(np.exp(-1j*2*np.pi/wr*np.dot(xs.T, mxs)), Psi_B_stop),
np.exp(-1j*2*np.pi/wr*np.dot(mxs.T, xs)))
Psi_C_stop = np.multiply(Psi_C, LS)
Psi_D = dx*dx/wr*np.dot(np.dot(np.exp(-1j*2*np.pi/wr*np.dot(xis.T, xs)), Psi_C_stop),
np.exp(-1j*2*np.pi/wr*np.dot(xs.T, xis)))
Psi_D_0_peak = np.sum(A*TelAp*LS)*dx*dx/wr
# Psi_D_0 = dx*dx/wr*np.dot(np.dot(np.exp(-1j*2*np.pi/wr*np.dot(xis.T, xs)), A*TelAp*LS),
# np.exp(-1j*2*np.pi/wr*np.dot(xs.T, xis)))
# intens_D_0 = np.power(np.absolute(Psi_D_0), 2)
# intens_D_0_peak = Psi_D_0_peak**2
intens_D_polychrom.append(np.power(np.absolute(Psi_D)/Psi_D_0_peak, 2))
return intens_D_polychrom
# -
intens_n_polychrom = get_onax_aplc_psf(TelAp, An, FPM, LS, xs, dx, mxs, dmx, xis, dxi, wrs)
intens_ne_polychrom = get_onax_aplc_psf(TelAp, An, FPM, LSe, xs, dx, mxs, dmx, xis, dxi, wrs)
intens_t_polychrom = get_onax_aplc_psf(TelAp, At, FPM, LS, xs, dx, mxs, dmx, xis, dxi, wrs)
intens_te_polychrom = get_onax_aplc_psf(TelAp, At, FPM, LSe, xs, dx, mxs, dmx, xis, dxi, wrs)
# +
plt.figure(figsize=(16,12))
plt.subplot(221)
plt.imshow(np.log10(intens_n_polychrom[2]), vmin=-11, vmax=-7, cmap='CMRmap')
plt.colorbar()
plt.title('On-axis PSF, design without LS alignment tolerance, perfect alignment')
plt.subplot(222)
plt.imshow(np.log10(intens_ne_polychrom[1]), cmap='CMRmap')
plt.colorbar()
plt.title('On-axis PSF, design without LS alignment tolerance, translated LS')
plt.subplot(223)
plt.imshow(np.log10(intens_t_polychrom[1]), vmin=-11, vmax=-7, cmap='CMRmap')
plt.colorbar()
plt.title('On-axis PSF, design with LS alignment tolerance, perfect alignment')
plt.subplot(224)
plt.imshow(np.log10(intens_te_polychrom[1]), vmin=-11, vmax=-7, cmap='CMRmap')
plt.colorbar()
plt.title('On-axis PSF, design with LS alignment tolerance, translated LS')
# -
# ## Intensity curve
def get_radial_intens(intens_polychrom, xis, seps, wrs):
radial_intens_polychrom = np.zeros((len(wrs), len(seps)))
XXs = np.asarray(np.dot(np.matrix(np.ones(xis.shape)).T, xis))
YYs = np.asarray(np.dot(etas.T, np.matrix(np.ones(etas.shape))))
RRs = np.sqrt(XXs**2 + YYs**2)
for si, sep in enumerate(seps):
r_in = np.max([seps[0], sep-0.5])
r_out = np.min([seps[-1], sep+0.5])
meas_ann_mask = np.logical_and(np.greater_equal(RRs, r_in),
np.less_equal(RRs, r_out))
#meas_ann_ind = np.nonzero(meas_ann_mask)
meas_ann_ind = np.nonzero(np.logical_and(np.greater_equal(RRs, r_in).ravel(),
np.less_equal(RRs, r_out).ravel()))[0]
for wi, wr in enumerate(wrs):
radial_intens_polychrom[wi, si] = np.mean(np.ravel(intens_polychrom[wi])[meas_ann_ind])
return radial_intens_polychrom
# +
rho0 = hexap_coron_aligntol.design['Image']['ida']
rho1 = hexap_coron_aligntol.design['Image']['oda'] + 1
seps = np.arange(rho0, rho1, 0.25)
radial_intens_n_polychrom = get_radial_intens(intens_n_polychrom, xis, seps, wrs)
radial_intens_ne_polychrom = get_radial_intens(intens_ne_polychrom, xis, seps, wrs)
radial_intens_t_polychrom = get_radial_intens(intens_t_polychrom, xis, seps, wrs)
radial_intens_te_polychrom = get_radial_intens(intens_te_polychrom, xis, seps, wrs)
# +
plt.figure(figsize=(16,12))
plt.subplot(221)
plt.plot(seps, np.log10(radial_intens_n_polychrom[0]))
plt.plot(seps, np.log10(radial_intens_n_polychrom[1]))
plt.plot(seps, np.log10(radial_intens_n_polychrom[2]))
plt.legend(['blue','center','red'], loc='upper left')
plt.xlabel('angular sep. (lambda_0/D)')
plt.ylabel('log10(I/I0)')
plt.title('On-axis PSF, design without LS alignment tolerance, perfect alignment')
plt.subplot(222)
plt.plot(seps, np.log10(radial_intens_ne_polychrom[0]))
plt.plot(seps, np.log10(radial_intens_ne_polychrom[1]))
plt.plot(seps, np.log10(radial_intens_ne_polychrom[2]))
plt.legend(['blue','center','red'], loc='upper left')
plt.xlabel('angular sep. (lambda_0/D)')
plt.ylabel('log10(I/I0)')
plt.title('On-axis PSF, design without LS alignment tolerance, translated LS')
plt.subplot(223)
plt.plot(seps, np.log10(radial_intens_t_polychrom[0]))
plt.plot(seps, np.log10(radial_intens_t_polychrom[1]))
plt.plot(seps, np.log10(radial_intens_t_polychrom[2]))
plt.legend(['blue','center','red'], loc='upper left')
plt.xlabel('angular sep. (lambda_0/D)')
plt.ylabel('log10(I/I0)')
plt.title('On-axis PSF, design with LS alignment tolerance, perfect alignment')
plt.subplot(224)
plt.plot(seps, np.log10(radial_intens_te_polychrom[0]))
plt.plot(seps, np.log10(radial_intens_te_polychrom[1]))
plt.plot(seps, np.log10(radial_intens_te_polychrom[2]))
plt.legend(['blue','center','red'], loc='upper left')
plt.xlabel('angular sep. (lambda_0/D)')
plt.ylabel('log10(I/I0)')
plt.title('On-axis PSF, design with LS alignment tolerance, translated LS')
# -
|
old_notebooks/scda_LDZ_test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import sklearn as skl
from sklearn.cluster import DBSCAN
from sklearn.manifold import TSNE
from sklearn.metrics.cluster import homogeneity_score
from sklearn.metrics.cluster import completeness_score
from sklearn.metrics.cluster import v_measure_score
from sklearn.metrics.cluster import adjusted_rand_score
from sklearn.metrics import silhouette_score
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
import time
import warnings
import os
import seaborn as sns # for ploting
from sklearn.metrics import fbeta_score, precision_score, recall_score, confusion_matrix,f1_score
import itertools
import pickle
import matplotlib
from matplotlib import pyplot as plt,style
from multiprocessing import Pool
# +
style.use('ggplot')
np.random.seed(42)
def plot_confusion_matrix(cm, classes,
recall,precision,f2,f1,
normalize=False,
title='Confusion matrix',
cmap=plt.cm.Blues):
plt.figure()
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=0)
plt.yticks(tick_marks, classes)
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, cm[i, j],
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.text(0,2.3, f" Recall:{recall},\n Precision:{precision},\n F2 Score:{f2},\n F1 Score:{f1}", fontsize=12)
plt.show()
# +
warnings.simplefilter("ignore", category=DeprecationWarning)
root_feature = '/Users/abhijit/Desktop/GIT_Projects/intl-iot/model/features-testing1.1/us'
root_model='/Users/abhijit/Desktop/GIT_Projects/intl-iot/models_final/features-testing1.1/us'
root_output=root_model+'/output'
dir_tsne_plots = root_model + '/tsne-plots'
num_pools=12
# -
lfiles = []
lparas= []
ldnames = []
for csv_file in os.listdir(root_feature):
if csv_file.endswith('.csv'):
print(csv_file)
train_data_file = '%s/%s' % (root_feature, csv_file)
dname = csv_file[:-4]
lfiles.append(train_data_file)
ldnames.append(dname)
lparas.append((train_data_file, dname))
p = Pool(num_pools)
for i in lparas:
print(i)
lparas[0][0]
data = pd.read_csv(lparas[0][0])
data.head()
set(data['state'])
# +
anomaly_data = pd.read_csv('/Users/abhijit/Desktop/GIT_Projects/intl-iot/model/new-features-testing1.1-all/us/google-home-mini.csv')
anomaly_data.head()
# -
set(anomaly_data['state'])
anomaly_data = anomaly_data[(anomaly_data['state'] == 'local_voice') | (anomaly_data['state'] == 'android_lan_photo')]
anomaly_data
len(anomaly_data)
ss1 = StandardScaler()
ss2 = StandardScaler()
data_features = data.drop(['device','state'], axis=1).fillna(-1)
data_std = ss1.fit_transform(data_features)
data_std = pd.DataFrame(data_std, index=data_features.index,columns=data_features.columns)
data_std['state'] = data['state']
anomaly_features = anomaly_data.drop(['device','state'], axis=1).fillna(-1)
anomaly_std = ss2.fit_transform(anomaly_features)
anomaly_std = pd.DataFrame(anomaly_std, index=anomaly_features.index, columns=anomaly_features.columns)
anomaly_std['state'] = anomaly_data['state']
# +
train, normal_test, _, _ = train_test_split(data_std, data_std, test_size=.2, random_state=42)
normal_valid, normal_test, _, _ = train_test_split(normal_test, normal_test, test_size=.5, random_state=42)
anormal_valid, anormal_test, _, _ = train_test_split(anomaly_std, anomaly_std, test_size=.5, random_state=42)
# -
train = train.reset_index(drop=True)
valid = normal_valid.append(anormal_valid).sample(frac=1).reset_index(drop=True)
test = normal_test.append(anormal_test).sample(frac=1).reset_index(drop=True)
len(train)
# + language="markdown"
# ### Save testing data. (Uncomment if not available already)
# -
check = ss2.inverse_transform(train.drop('state', axis=1))
check = pd.DataFrame(check, columns=train.columns[train.columns != 'state'])
check['state'] = train['state']
check.to_csv('training_data_without_anomaly.csv')
check = ss2.inverse_transform(test.drop('state', axis=1))
check = pd.DataFrame(check, columns=test.columns[test.columns != 'state'])
check['state'] = test['state']
check.to_csv('testing_data_with_anomaly.csv')
train['state'] = train['state'].apply(lambda x: 1 if x == 'local_voice' else 0)
valid['state'] = valid['state'].apply(lambda x: 1 if x == 'local_voice' else 0)
test['state'] = test['state'].apply(lambda x: 1 if x == 'local_voice' else 0)
print('Train shape: ', train.shape)
print('Proportion os anomaly in training set: %.2f\n' % train['state'].mean())
print('Valid shape: ', valid.shape)
print('Proportion os anomaly in validation set: %.2f\n' % valid['state'].mean())
print('Test shape:, ', test.shape)
print('Proportion os anomaly in test set: %.2f\n' % test['state'].mean())
# + language="markdown"
# # Load original model
# -
original_model = pickle.load(open('/Users/abhijit/Desktop/GIT_Projects/intl-iot/model/tagged-models/us/yi-cameraknn.model','rb'))
labels = []
with open('/Users/abhijit/Desktop/GIT_Projects/intl-iot/model/tagged-models/us/yi-camera.label.txt') as ff:
for line in ff.readlines():
line = line.strip()
if line.startswith('#') or line == '':
continue
labels.append(line)
check = ss2.inverse_transform(test.drop('state', axis=1))
check = pd.DataFrame(check, columns=test.columns[test.columns != 'state'])
check['state'] = test['state']
ss_knn = original_model['standard_scaler']
pca = original_model['pca']
trained_model = original_model['trained_model']
unknown_data = ss_knn.transform(check.drop('state', axis=1))
unknown_data = pca.transform(unknown_data)
unknown_data = pd.DataFrame(unknown_data)
unknown_data = unknown_data.iloc[:, :4]
y_predict = trained_model.predict(unknown_data)
p_readable = []
for pindex in range(len(y_predict)):
label_predicted = labels[np.argmax(y_predict[pindex])]
p_readable.append(label_predicted)
y_hat_test = []
for i in p_readable:
if i:
y_hat_test.append(0)
else:
y_hat_test.append(1)
# +
recall = recall_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
precision = precision_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
f2 = fbeta_score(y_pred=y_hat_test, y_true=test['state'].values, beta=2,average='weighted')
f1 = f1_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
cnf_matrix = confusion_matrix(test['state'].values, y_hat_test)
plot_confusion_matrix(cnf_matrix, classes=['Normal','Anormal'],
recall=recall,precision=precision,f2 = f2,f1=f1,title='Confusion matrix')
# + language="markdown"
# # New model
# + language="markdown"
# ## Types of outliers
# In general, outliers can be classified into three categories, namely global outliers, contextual (or conditional) outliers, and collective outliers.
#
# Global outlier — Object significantly deviates from the rest of the data set
#
# Contextual outlier — Object deviates significantly based on a selected context. For example, 28⁰C is an outlier for a Moscow winter, but not an outlier in another context, 28⁰C is not an outlier for a Moscow summer.
#
# Collective outlier — A subset of data objects collectively deviate significantly from the whole data set, even if the individual data objects may not be outliers. For example, a large set of transactions of the same stock among a small party in a short period can be considered as an evidence of market manipulation.
# +
from scipy.stats import multivariate_normal
mu = train.drop('state', axis=1).mean(axis=0).values
sigma = train.drop('state', axis=1).cov().values
model = multivariate_normal(cov=sigma, mean=mu, allow_singular=True)
print(np.median(model.logpdf(valid[valid['state'] == 0].drop('state', axis=1).values)))
print(np.median(model.logpdf(valid[valid['state'] == 1].drop('state', axis=1).values)))
# +
tresholds = np.linspace(-100,-10, 300)
scores = []
for treshold in tresholds:
y_hat = (model.logpdf(valid.drop('state', axis=1).values) < treshold).astype(int)
scores.append([recall_score(y_pred=y_hat, y_true=valid['state'].values),
precision_score(y_pred=y_hat, y_true=valid['state'].values),
fbeta_score(y_pred=y_hat, y_true=valid['state'].values, beta=2)])
scores = np.array(scores)
print(scores[:, 2].max(), scores[:, 2].argmax())
# -
plt.plot(tresholds, scores[:, 0], label='$Recall$')
plt.plot(tresholds, scores[:, 1], label='$Precision$')
plt.plot(tresholds, scores[:, 2], label='$F_2$')
plt.ylabel('Score')
# plt.xticks(np.logspace(-10, -200, 3))
plt.xlabel('Threshold')
plt.legend(loc='best')
plt.show()
# +
final_tresh = tresholds[scores[:, 2].argmax()]
y_hat_test = (model.logpdf(test.drop('state', axis=1).values) < final_tresh).astype(int)
recall = recall_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
precision = precision_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
f2 = fbeta_score(y_pred=y_hat_test, y_true=test['state'].values, beta=2,average='weighted')
f1 = f1_score(y_pred=y_hat_test, y_true=test['state'].values,average='weighted')
cnf_matrix = confusion_matrix(test['state'].values, y_hat_test)
plot_confusion_matrix(cnf_matrix, classes=['Normal','Anormal'],
recall=recall,precision=precision,f2 = f2,f1=f1, title='Confusion matrix')
# -
combined_data = data_std.append(anomaly_std)
combined_data['state'] = combined_data['state'].apply(lambda x: 1 if x == 'local_voice' else 0)
columns_data = combined_data.columns
columns_data = columns_data.drop('state')
import seaborn as sns
import matplotlib.gridspec as gridspec
dataset = combined_data
# plt.figure(figsize=(12,28*5))
# gs = gridspec.GridSpec(28, 1)
for feature in columns_data:
ax = plt.subplot()
sns.distplot(dataset[feature][dataset.state == 1], bins=10, label='Anomaly')
sns.distplot(dataset[feature][dataset.state == 0], bins=10, label='Normal')
ax.set_xlabel('')
ax.set_title('histogram of feature: ' + str(feature))
plt.legend(loc='best')
plt.show()
# + language="markdown"
# ## Saving the new model.
# -
d = dict({'mvmodel' : model, 'treshold' : final_tresh})
f = open("multivariate_model.pkl", "wb")
pickle.dump(d, f)
f.close()
|
model/anomaly-detection-notebooks/anomaly_detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df=pd.read_excel("marketing_data.xlsx")
df
for x in df.index:
if df.loc[x,"Year_Birth"] >1954:
print(df.loc[x,"ID"])
df_sd=df["MntFruits"].std()
print(df_sd)
for y in df.index:
if df.loc[y,"Marital_Status"]=="Married":
print(df.loc[y,"ID"])
for z in df.index:
if df.loc[z,"Income"]>49000:
if df.loc[z,"Education"]=="Graduation":
print(df.loc[z,"ID"])
df.plot(x = 'MntFishProducts',y = 'NumCatalogPurchases')
df.mean
df.max
import pandas as pd
df=pd.read_excel("financial_analytics.xlsx")
df
income=df.loc[0,"Income"]
hra=df.loc[0,"HRA"]
insurance=df.loc[0,"Insurance"]
da=df.loc[0,"DA"]
if insurance<150000:
tax1=0
else:
tax1=(10/100)*(insurance-150000)
if income<=250000:
tax2=0
if income<=500000:
tax2=5/100*(income-250000)
if income<=750000:
tax2=12500+((10/100)*(income-500000))
if income<=1000000:
tax2=37500+((15/100)*(income-750000))
if income<=1250000:
tax2=75000+((20/100)*(income-1000000))
if income<=1500000:
tax2=125000+((25/100)*(income-1250000))
if income>1500000:
tax2=187500+((30-100)*(income-1500000))
if hra<(10/100)*x:
tax3=0
else:
tax3=(50/100)*(hra-(10/100)*income)
if da<10000:
tax4=0
else:
tax4=(10/100)*(da-10000)
finaltax=tax1+tax2+tax3+tax4
print("final_tax",finaltax)
print("case1",tax1)
print("case2",tax2)
print("case3",tax3)
print("case4",tax4)
|
exam (2).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: RAPIDS Stable
# language: python
# name: python3
# ---
# <a id="introduction"></a>
# # Ciencia de Datos Acelerada con GPU
# ## Introducción a cuDF para usuarios de pandas
# #### Presentador: <NAME> (BLAZINGSQL)
#
# Notebook adaptado a español desde [02_Introduction_to_cuDF_for_pandas.ipynb](https://github.com/BlazingDB/Welcome_to_BlazingSQL_Notebooks/blob/master/webinars/rapids_intro/02_Introduction_to_cuDF_for_pandas.ipynb) por <NAME>.
#
# -------
#
# En este Notebook presumiremos de los beneficios de usar cuDF. cuDF es una librería en Python de DataFrame GPU, para cargar, ejecutar joins, hacer agregaciones, filtrar y toda clase de manipulación de data tabular usando un DataFrame style API.
# # Primero, sumemos las librerías necesarias
# +
import os
import pandas as pd
import numpy as np
import cupy as cp
import cudf as cd
# -
# # Importemos data desde un archivo csv
# ### **movies_pdf** es nuestro DataFrame de Pandas
movies_pdf = pd.read_csv("s3://bsql/data/rapids_intro/movies.csv")
# ### **movies_cdf** es nuestro cuDF
movies_cdf = cd.read_csv("s3://bsql/data/rapids_intro/movies.csv", storage_options={'anon': True})
# # Ejemplos de estadísticas en dataset
print(movies_pdf.shape)
print(movies_pdf.ndim)
print(len(movies_pdf))
print(movies_cdf.shape)
print(movies_cdf.ndim)
print(len(movies_cdf))
# # Explorando la Data
movies_pdf.head()
movies_cdf.head()
movies_pdf.info()
movies_cdf.info()
movies_pdf.columns
movies_cdf.columns
# # Seleccionando subconjuntos del DataFrame
# ### Seleccionando campos que son data continua
movies_pdf.select_dtypes(include="number").head()
movies_cdf.select_dtypes(include="number").head()
# ### Seleccionando campos que son floats
movies_pdf.select_dtypes(include="float").head()
movies_cdf.select_dtypes(include="float").head()
# ### Seleccionando campos que son valores discretos
movies_pdf.select_dtypes(include="object").head()
movies_cdf.select_dtypes(include="object").head()
# # Análisis de Data
# ### Resumen estadístico para todos los campos de data continua
movies_pdf.select_dtypes(include="number").describe()
movies_cdf.select_dtypes(include="number").describe()
# ### Resumen estadístico para todos los campos con valores discretos
movies_pdf.select_dtypes(include="object").describe()
movies_cdf.select_dtypes(include="object").describe()
# ### Transpuesta de los resultados de la función describe() de cuDF
# (esto se puede hacer también en Pandas)
movies_cdf.select_dtypes(include="number").describe().T
# ### Cálculo de Covarianza de dos variables continuas
movies_pdf.movie_facebook_likes.cov(movies_pdf.actor_3_facebook_likes)
movies_cdf.movie_facebook_likes.cov(movies_cdf.actor_3_facebook_likes)
# ### Correlación Pearson de dos variables continuas
movies_pdf.movie_facebook_likes.corr(movies_pdf.actor_3_facebook_likes)
movies_cdf.movie_facebook_likes.corr(movies_cdf.actor_3_facebook_likes)
# ### Agrupaciones
# Analizando los montos brutos (gross amounts) generados por los dos actores principales
movies_pdf[['actor_1_name','actor_2_name','gross']].groupby(['actor_1_name','actor_2_name']).sum()
movies_cdf[['actor_1_name','actor_2_name','gross']].groupby(['actor_1_name','actor_2_name']).sum()
# # Preparación de la Data
# Algunos géneros (genres) son resultado de la combinación de otros varios géneros. Por ejemplo: `Action|Adventure|Comedy|Fantasy|Sci-Fi`
print('There are ' + str(len(movies_pdf.genres.unique())) + ' genre combinations in the genres field')
print('Examples:\n', movies_pdf.genres.unique()[:10])
# ## Dividiendo la columna Género (Genre) usando pandas
genres_pdf = movies_pdf.join(movies_pdf.genres.str.split('|', expand=True).add_prefix('genre_'))
genres_pdf
# ## Dividiendo la columna Género (Genre) usando cuDF
# cuDF no tiene la opción add_prefix() cuando dividimos una columna. Acá mostramos una opción para renombrar la columna en cuDF.
genre_fields = len(movies_cdf.genres.str.split('|', expand=True).columns)
print('There will be ' + str(genre_fields) + ' new columns that will be added into our dataframe\n')
genres_cdf = movies_cdf.join(movies_cdf.genres.str.split('|', expand=True))
genres_cdf.head()
# **Los nuevos nombres de columna son asignados a números, que no son strings**
col_numbers = genres_cdf.columns[-genre_fields:].to_list()
print(col_numbers)
# **Necesitas convertirlos a strings, asignarles un prefijo y convertir ambas listas a un diccionario para renombrar las nuevas columnas en cuDF**
# +
new_col_names = ['genre_' + str(x) for x in col_numbers]
print(new_col_names)
new_col_dict = dict(zip(col_numbers, new_col_names))
print(new_col_dict)
# -
genres_cdf = genres_cdf.rename(columns=new_col_dict)
genres_cdf.head()
# # One Hot Encoding
# ## OHE usando Pandas
# Aplicado a la columna genre_0
pd_ohe = pd.get_dummies(genres_pdf.genre_0, prefix='genre_0')
df = pd.concat([genres_pdf, pd_ohe], axis=1)
df.head()
# ## OHE usando cuDF
# Aplicado a la columna genre_0
cdf = cd.get_dummies(genres_cdf, prefix='genre_0', columns=['genre_0'])
cdf.head()
|
webinars/rapids_intro/02_Introduccion_a_cuDF_para_pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load data, train model
import sage
import numpy as np
from catboost import CatBoostClassifier
from sklearn.model_selection import train_test_split
# +
# Load data
df = sage.datasets.bank()
# Feature names and categorical columns (for CatBoost model)
feature_names = df.columns.tolist()[:-1]
categorical_cols = ['Job', 'Marital', 'Education', 'Default', 'Housing',
'Loan', 'Contact', 'Month', 'Prev Outcome']
categorical_inds = [feature_names.index(col) for col in categorical_cols]
# -
# Split data
train, test = train_test_split(
df.values, test_size=int(0.1 * len(df.values)), random_state=123)
train, val = train_test_split(
train, test_size=int(0.1 * len(df.values)), random_state=123)
Y_train = train[:, -1].copy().astype(int)
Y_val = val[:, -1].copy().astype(int)
Y_test = test[:, -1].copy().astype(int)
train = train[:, :-1].copy()
val = val[:, :-1].copy()
test = test[:, :-1].copy()
# +
model = CatBoostClassifier(iterations=100,
learning_rate=0.3,
depth=10)
model = model.fit(train, Y_train, categorical_inds, eval_set=(val, Y_val),
verbose=False)
# -
# # Shapley Effects cooperative game
from shapreg import removal, stochastic_games, shapley
from shapreg.utils import crossentropyloss
# +
# Make model callable
model_lam = lambda x: model.predict_proba(x)
# Model extension
marginal_extension = removal.MarginalExtension(train[:128], model_lam)
# -
# Set up game (output loss)
game = stochastic_games.DatasetOutputGame(marginal_extension, test, crossentropyloss)
# Run estimator
explanation = shapley.ShapleyRegression(game, thresh=0.02)
# Plot with 95% confidence intervals
explanation.plot(feature_names, title='Shapley Effects')
|
notebooks/bank.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="4f3CKqFUqL2-" slideshow={"slide_type": "slide"}
# # First Steps with TensorFlow
# + [markdown] colab_type="text" id="Bd2Zkk1LE2Zr"
# For the current exercises, see the following Colabs:
#
# - [Simple Linear Regression with Synthetic Data](https://colab.research.google.com/github/google/eng-edu/blob/main/ml/cc/exercises/linear_regression_with_synthetic_data.ipynb)
#
# - [Linear Regression with a Real Dataset](https://colab.research.google.com/github/google/eng-edu/blob/main/ml/cc/exercises/linear_regression_with_a_real_dataset.ipynb)
|
ml/cc/exercises/estimators/first_steps_with_tensor_flow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network Creation
# +
# %load_ext autoreload
# %autoreload
# #%autoreload
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
display(HTML("""<style>div.output_area{max-height:10000px;overflow:scroll;}</style>"""))
import networkx as nx
from networkTrips import organizeTrips
from timeUtils import clock, elapsed, getDateTime
from ioUtils import loadJoblib, saveFile, getFile
from fsUtils import mkDir, mkSubDir, setFile, setSubDir
from pandasUtils import getRowData, getColData, dropColumns
from geoUtils import convertMetersToLat, convertLatToMeters, convertMetersToLong, convertLongToMeters
from geocluster import geoClusters, geoCluster
from geoclusterUtils import genCenters, genCluster, genClusters, genTripsBetweenClusters
from networkClusterMaps import foliumMap
from networkOutput import printNetwork
from driverNetwork import driverNetwork
from networkFeatures import networkFeatures
import pandas as pd
pd.set_option("display.max_rows",1000)
pd.set_option('precision', 3)
import warnings
warnings.filterwarnings('ignore')
_, _ = clock("Last Run")
# -
savedir = "/Users/tgadf/Downloads/network"
mkDir(savedir)
# # Load/Generate Data
# +
#######################################################################################
# Generate Clusted Data
#######################################################################################
genData = False
if genData:
cls = 20
total = 500
genMax = 75
distMax = 500
raw = genClusters(cls, 250, latRange=[29.8, 30.2], lngRange=[49.8, 50.2], dist="gauss", maxrad=genMax)
gc = geoClusters(key="dummy", points=raw, distMax=distMax, debug=False)
gc.findClusters(seedMin=2, debug=False)
df = genTripsBetweenClusters(n=total, gc=gc, returnDF=True)
df["device"] = "dummy"
tmpdf = loadJoblib("/Users/tgadfort/Downloads/r4hIDs.p").sample(n=total, replace=True)
tojoin = tmpdf.sample(cls)
tojoin["cl"] = ["cl{0}".format(x) for x in range(cls)]
df['cl'] = df['cl0']
drops = [x for x in tojoin.columns if x.startswith("Geo1")]
tojoinCL0 = dropColumns(tojoin, columns=drops, inplace=False)
test = df.merge(tojoinCL0, on='cl')
test['cl'] = test['cl1']
drops = [x for x in tojoin.columns if x.startswith("Geo0")]
tojoinCL1 = dropColumns(tojoin, columns=drops, inplace=False)
test = test.merge(tojoinCL1, on='cl')
gpsdata = test
dropColumns(gpsdata, columns=["cl", "cl0", "cl1"])
gpsdata.replace('nan', 0, inplace=True)
else:
fname = "/Users/tgadf/Downloads/gpsTripsOakRidge (1).p"
print("Loading {0}".format(fname))
gpsdata = loadJoblib(fname)
_, _ = clock("Last Run")
# -
# ## Show Data (if needed)
# ## Subselect (if needed)
device = '352252060173789'
debug = True
gpsdata = gpsdata[gpsdata['device'] == device]
print("Keeping {0} rows".format(gpsdata.shape[0]))
_, _ = clock("Last Run")
# # Cluster and Sort Trips
i = 0
nd = gpsdata['device'].nunique()
for device, df in gpsdata.groupby('device'):
print('Key = {0}'.format(device),'\tRun = {0}/{1}'.format(i,nd),'\tTrips = {0}'.format(df.shape[0]))
i += 1
#######################################################################################
# Cluster Geo Data (Lat, Long)
#######################################################################################
points = df[["lat0", "long0"]]
points.columns = ["lat", "long"]
pnts = df[["lat1", "long1"]]
pnts.columns = ["lat", "long"]
points = points.append(pnts)
#######################################################################################
# Create Clusters
#######################################################################################True
gc = geoClusters(key="dummy", points=points, distMax=200, mergeFactor=2.5, debug=False)
gc.createCells(debug=False)
gc.createProtoClusters(seedMin=4, debug=False)
gc.createSeedlessClusters(seedMin=2, debug=False)
gc.mergeClusters(debug=False)
print("Found {0} geo clusters".format(gc.getNClusters()))
#######################################################################################
# Set Nearest Clusters
#######################################################################################
if debug:
start, cmt = clock("Finding Nearest Clusters for Start of Trips")
geoResults = df[['lat0', 'long0']].apply(gc.getNearestClusters, axis=1).values
df["geo0"] = [x[0] for x in geoResults]
if debug:
elapsed(start, cmt)
start, cmt = clock("Finding Nearest Clusters for End of Trips")
geoResults = df[['lat1', 'long1']].apply(gc.getNearestClusters, axis=1).values
df["geo1"] = [x[0] for x in geoResults]
if debug:
elapsed(start, cmt)
#######################################################################################
# Organize Trips for Network
#######################################################################################
trips = organizeTrips(df=df, gc=gc, debug=True, requireGood=False)
# ## Investigate Individual Cluster
clname = 'cl9'
clusterData = df[(df['geo0'] == clname) | (df['geo1'] == clname)]
# +
geocols = [x.replace("Geo0", "") for x in clusterData.columns if x.startswith("Geo0") and x.find("CENSUS") == -1 and x.find("ASDW") == -1]
from collections import Counter
cldata = Counter()
clprefix = None
for ir,row in clusterData.iterrows():
if row['geo0'] == clname:
clprefix = "Geo0"
elif row['geo1'] == clname:
clprefix = "Geo1"
for g in range(2):
for col in geocols:
colname = "{0}{1}".format(clprefix, col)
colval = row[colname]
try:
if colval > 0:
cldata[col] += 1
except:
pass
tripcldata = {}
for k,v in trips['vertexMetrics'][clname].items():
if not isinstance(v, dict):
continue
for k2,v2 in v.items():
if isinstance(v2, Counter):
if v2.get(1.0):
tripcldata[k2] = v2
### Show Results
print("Cluster {0} --> {1}".format(clname, cldata.most_common()))
print("Cluster {0} --> {1}".format(clname, tripcldata))
# -
# # Saved Data
# +
# Save trips/gc if needed
if False:
deviceDir = mkSubDir(savedir, device)
tripsfile = setFile(deviceDir, "trips.p")
gcfile = setFile(deviceDir, "gc.p")
loadTrips=False
if loadTrips:
trips = getFile(tripsfile)
gc = getFile(gcfile)
else:
print("Saving to {0}".format(deviceDir))
saveFile(ifile=gcfile, idata=gc)
saveFile(ifile=tripsfile, idata=trips)
_, _ = clock("Last Run")
# -
# Show data if needed
df.head().T
# # Driver Network
# +
# %load_ext autoreload
# %autoreload
from driverNetwork import driverNetwork
from edgeInfo import edgeInfo
from vertexInfo import vertexInfo
from networkCategories import categories
from networkAlgos import networkAlgos
start, cmt = clock("Running Driver Network")
dn = driverNetwork(trips)
dn.create(debug=True)
dn.computeNetworkAttrs(debug=True, level=1)
dn.fillVertexCensusData(debug=True)
dn.fillVertexGeospatialData(debug=True)
dn.fillVertexInternalData(debug=True)
dn.fillVertexNetworkData(debug=True)
dn.fillEdgeInternalData(debug=True)
dn.fillEdgeVertexData(debug=True)
dn.fillEdgeNetworkData(debug=True)
g = dn.getNetwork()
elapsed(start, cmt)
# -
if True:
# %load_ext autoreload
# %autoreload
from networkOutput import printNetwork
pn = printNetwork(dn)
pn.printVertices(minN=40)
pn.printEdges(minW=20)
dn.getVertexByName('cl9', 'attr')
# # Network Features
# +
# %load_ext autoreload
# %autoreload
from networkFeatures import networkFeatures
from networkAlgos import algos
nf = networkFeatures(dn)
## Vertex Counts
nf.fillVertexCensusCounts(debug=True)
nf.fillVertexInternalCounts(debug=True)
nf.fillVertexGeoSpatialCounts(debug=True)
nf.fillVertexProperties(debug=True)
## Edge Counts
nf.fillEdgeInternalCounts(debug=True)
nf.fillEdgeCensusCounts(debug=True)
nf.fillEdgeGeoSpatialCounts(debug=True)
nf.fillEdgeProperties(debug=True)
## Network Counts
nf.fillNetworkFeatures(debug=True)
## Home Counts
nf.fillHomeFeatures(debug=True)
## Indiv Vertex/Edge Values
#nf.fillIndividualVertexFeatures(debug=True)
#nf.fillIndividualEdgeFeatures(debug=True)
## Vertex/Edge Correlations
##nf.fillVertexFeatureCorrelations(debug=True)
#nf.fillEdgeFeatureCorrelations(debug=True)
_,_ = clock("Last Run")
# -
nf.getFeatures()
nf.getHomeFeatureDataFrame().T
# # GeoSpatial Maps
# +
from folium import PolyLine, CircleMarker, Circle, Marker, Icon, FeatureGroup, Map, LayerControl
import geohash
class foliumMap():
def __init__(self, df=None, pc=None, gc=None, dn=None, nf=None):
self.pc = None
if pc is not None:
self.setProtoClusters(pc)
self.gc = None
if gc is not None:
self.setGeoClusters(gc)
self.dn = None
if dn is not None:
self.setDriverNetwork(dn)
self.nf = None
if nf is not None:
self.setNetworkFeatures(nf)
self.df = None
if df is not None:
self.setTripsDataFrame(df)
self.m = None
self.init_zoom = 10
self.colors = ['red', 'blue', 'gray', 'darkred', 'lightred', 'orange', 'beige', 'green', 'darkgreen', 'lightgreen',
'darkblue', 'lightblue', 'purple', 'darkpurple', 'pink', 'cadetblue', 'lightgray']
########################################################################################
# Setters
########################################################################################
def setTripsDataFrame(self, df):
self.df = df
self.df = df[["lat0", "long0"]]
self.df.columns = ["lat", "long"]
pnts = df[["lat1", "long1"]]
pnts.columns = ["lat", "long"]
self.df = self.df.append(pnts)
def setProtoClusters(self, pc):
self.pc = pc
def setGeoClusters(self, gc):
self.gc = gc
def setDriverNetwork(self, dn):
self.dn = dn
def setNetworkFeatures(self, nf):
self.nf = nf
########################################################################################
# Getters
########################################################################################
def getMap(self):
return self.m
########################################################################################
########################################################################################
# Create Map
########################################################################################
########################################################################################
def createMapFromTripsDataFrame(self, zoom=None, debug=False):
if self.df is None:
print("There is no trips DataFrame object!")
return
try:
lat0 = self.df['lat'].mean()
long0 = self.df['long'].mean()
except:
raise ValueError("Could not get center of geo clusters and create map!")
if zoom is None:
zoom = self.init_zoom
self.m = Map(location=[lat0, long0], zoom_start=zoom)
def createMapFromProtoClusters(self, zoom=None, debug=False):
if self.pc is None:
print("There is no ProtoClusters object!")
return
try:
lats = Series(x["CoM"][0] for pcl,x in self.pc.items())
lngs = Series(x["CoM"][1] for pcl,x in self.pc.items())
lat0 = lats.mean()
long0 = lngs.mean()
except:
raise ValueError("Could not get center of geo clusters and create map!")
if zoom is None:
zoom = self.init_zoom
self.m = Map(location=[lat0, long0], zoom_start=zoom)
def createMapFromGeoClusters(self, zoom=None, debug=False):
if self.gc is None:
print("There is no GeoClusters object!")
return
try:
coms = self.gc.getClusterCoMs()
lat0 = coms[0].mean()
long0 = coms[1].mean()
except:
raise ValueError("Could not get center of geo clusters and create map!")
if zoom is None:
zoom = self.init_zoom
self.m = Map(location=[lat0, long0], zoom_start=zoom)
def createMapFromDriverNetwork(self, zoom=None, debug=False):
if self.dn is None:
print("There is no DriverNetwork object!")
return
if self.gc is None:
try:
self.dn.createGC()
self.gc = self.dn.getGC()
except:
raise ValueError("Could not get create geo clusters from driver network")
try:
coms = self.gc.getClusterCoMs()
lat0 = coms[0].mean()
long0 = coms[1].mean()
except:
raise ValueError("Could not get center of geo clusters and create map!")
if zoom is None:
zoom = self.init_zoom
self.m = Map(location=[lat0, long0], zoom_start=zoom)
def createMapFromNetworkFeatures(self, zoom=None, debug=False):
if self.nf is None:
print("There is no NetworkFeatures object!")
return
if zoom is None:
zoom = self.init_zoom
self.m = Map(location=[lat0, long0], zoom_start=zoom)
def createMap(self, zoom=None, debug=False):
if self.nf is not None:
self.createMapFromNetworkFeatures(zoom=zoom, debug=debug)
elif self.dn is not None:
self.createMapFromDriverNetwork(zoom=zoom, debug=debug)
elif self.gc is not None:
self.createMapFromGeoClusters(zoom=zoom, debug=debug)
elif self.pc is not None:
self.createMapFromProtoClusters(zoom=zoom, debug=debug)
elif self.df is not None:
self.createMapFromTripsDataFrame(zoom=zoom, debug=debug)
else:
raise ValueError("Cannot create map because there is no data object!")
########################################################################################
########################################################################################
# Points/Clusters
########################################################################################
########################################################################################
def addPointsFromTripsDataFrame(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.df is None:
print("DataFrame is None!")
return
cols = ['darkblue', 'lightblue', 'pink', 'lightgray']
rad = 5
weight = 1
for row in self.df.iterrows():
com = row[1].values
Circle(com, color=cols[1], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(self.m)
def addPointsFromGeoClusterCells(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cells = gc.getCells()
for geo,cnt in cells.iteritems():
com = geohash.decode_exactly(geo)[:2]
wgt = int(cnt)
popup = str(wgt)
Circle(com, color='black', radius=5, fill=True, fill_color='black', weight=wgt, opacity=0, popup=popup).add_to(self.m)
def addPointsFromProtoClusters(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cols = ['darkblue', 'lightblue', 'pink', 'lightgray']
from pandas import Series
for pcl,protoCluster in gc.getProtoClusters().items():
com = protoCluster["CoM"]
rad = max([protoCluster["Radius"], 5])
counts = protoCluster["Counts"]
weight = int(counts)
name = pcl
popup = "{0} : N = {1}".format(name, counts)
Circle(com, color=cols[0], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(self.m)
def addPointsFromSeedlessClusters(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cols = ['darkgreen']
from pandas import Series
for scl,seedlessCluster in gc.getSeedlessClusters().items():
com = seedlessCluster["CoM"]
rad = max([seedlessCluster["Radius"], 5])
counts = seedlessCluster["Counts"]
weight = int(counts)
name = scl
popup = "{0} : N = {1}".format(name, counts)
Circle(com, color=cols[0], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(self.m)
def addPointsFromMergedClusters(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cols = ['darkred']
from pandas import Series
for cl,cluster in gc.getMergedClusters().items():
com = cluster["CoM"]
rad = max([cluster["Radius"], 5])
counts = cluster["Counts"]
weight = int(counts)
name = cl
popup = "{0} : N = {1}".format(name, counts)
Circle(com, color=cols[0], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(self.m)
def addPointsFromGeoClusters(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cols = ['darkblue', 'lightblue', 'pink', 'lightgray']
from pandas import Series
feature_group_1 = FeatureGroup(name="Driver Top 90%")
feature_group_2 = FeatureGroup(name="Driver Top 75%")
feature_group_3 = FeatureGroup(name="Driver Top 50%")
feature_group_4 = FeatureGroup(name="Driver Low 50%")
weights = Series([cluster.getCounts() for cl,cluster in gc.getClusters().items()])
alpha = weights.quantile(0.9)
beta = weights.quantile(0.75)
gamma = weights.quantile(0.5)
for cl,cluster in gc.getClusters().items():
com = cluster.getCoM()
rad = max([cluster.getRadius(), 10])
counts = cluster.getCounts()
weight = float(counts)
quant = cluster.getQuantiles()
name = cl
cells = ",".join(cluster.getCells())
popup = "{0} : N = {1} : {2}".format(name, weight, com)
#popup = ""
if counts > alpha:
Marker(com, icon=Icon(color=cols[0], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_1)
Circle(com, color=cols[0], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(feature_group_1)
elif counts > beta:
Marker(com, icon=Icon(color=cols[1], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_2)
Circle(com, color=cols[1], radius=rad, fill=True, fill_color=cols[1], weight=weight, opacity=0).add_to(feature_group_2)
elif counts > gamma:
Marker(com, icon=Icon(color=cols[2], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_3)
Circle(com, color=cols[2], radius=rad, fill=True, fill_color=cols[2], weight=weight, opacity=0).add_to(feature_group_3)
else:
Marker(com, icon=Icon(color=cols[3], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_4)
Circle(com, color=cols[3], radius=rad, fill=True, fill_color=cols[3], weight=weight, opacity=0).add_to(feature_group_4)
feature_group_1.add_to(self.m)
feature_group_2.add_to(self.m)
feature_group_3.add_to(self.m)
feature_group_4.add_to(self.m)
LayerControl().add_to(self.m)
def addPointsFromDriverNetwork(self, debug=False):
if self.m is None:
print("Folium Map is None!")
return
if self.dn is None:
print("DrivingNetwork is None!")
return
if self.gc is None:
print("GeoClusters is None!")
return
cols = ['darkblue', 'lightblue', 'pink', 'lightgray']
from pandas import Series
feature_group_1 = FeatureGroup(name="Driver Home")
feature_group_2 = FeatureGroup(name="Daily Visits")
feature_group_3 = FeatureGroup(name="Weekly Visits")
feature_group_4 = FeatureGroup(name="Monthly Visits")
feature_group_5 = FeatureGroup(name="Infrequent Visits")
feature_group_5 = None
clusters = self.gc.getClusters()
for vertexName in self.dn.getVertices():
if vertexName == 'None':
continue
cluster = clusters[vertexName]
clname = cluster.getName()
com = cluster.getCoM()
rad = max([int(cluster.getRadius()), 10])
weight = 10
clusterFeatures = self.dn.getVertexByName(vertexName, "feat")
home = clusterFeatures["Internal"]["IsHome"]
place = clusterFeatures["Census"]["Place"]
active = clusterFeatures["Internal"]["FractionalVisits"]
visits = clusterFeatures["Internal"]["DailyVisits"]
n = int(clusterFeatures["Internal"]["N"])
pois = ", ".join([k for k,v in clusterFeatures["GeoSpatial"].items() if v > 0])
popup = "{0} ({1}) : N = {2} : fActive = {3} : POIs: {4} : {5}".format(vertexName, place, n, active, pois, com)
#print(vertexName,popup)
if home == 1:
Marker(com, icon=Icon(color='darkred', icon_color='white', icon="home", angle=0, prefix='fa'), popup=popup).add_to(feature_group_1)
Circle(com, color='darkred', radius=rad, fill=True, fill_color='darkred', weight=weight, opacity=0).add_to(feature_group_1)
elif active == "Daily":
Marker(com, icon=Icon(color=cols[0], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_2)
Circle(com, color=cols[0], radius=rad, fill=True, fill_color=cols[0], weight=weight, opacity=0).add_to(feature_group_2)
elif active == "Weekly":
Marker(com, icon=Icon(color=cols[1], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_3)
Circle(com, color=cols[1], radius=rad, fill=True, fill_color=cols[1], weight=weight, opacity=0).add_to(feature_group_3)
elif active == "Monthly":
Marker(com, icon=Icon(color=cols[2], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_4)
Circle(com, color=cols[2], radius=rad, fill=True, fill_color=cols[2], weight=weight, opacity=0).add_to(feature_group_4)
else:
if feature_group_5 is not None:
Marker(com, icon=Icon(color=cols[3], icon_color='white', icon="car", angle=0, prefix='fa'), popup=popup).add_to(feature_group_5)
Circle(com, color=cols[3], radius=rad, fill=True, fill_color=cols[3], weight=weight, opacity=0).add_to(feature_group_5)
feature_group_1.add_to(self.m)
feature_group_2.add_to(self.m)
feature_group_3.add_to(self.m)
feature_group_4.add_to(self.m)
if feature_group_5 is not None:
feature_group_5.add_to(self.m)
LayerControl().add_to(self.m)
def addPoints(self, debug=False):
if self.m is None:
raise ValueError("Cannot add points to an empty map")
if self.nf is not None:
self.addPointsFromNetworkFeatures(debug=debug)
elif self.dn is not None:
self.addPointsFromDriverNetwork(debug=debug)
elif self.pc is not None:
self.addPointsFromProtoClusters(debug=debug)
elif self.gc is not None:
self.addPointsFromGeoClusters(debug=debug)
elif self.df is not None:
self.addPointsFromTripsDataFrame(debug=debug)
else:
print("Cannot add points because there is map and object!")
# -
# ## Show Clustering GeoSpatial Map
fm = foliumMap(pc=gc.getProtoClusters(), gc=gc)
fm.createMap(zoom=11)
fm.addPointsFromGeoClusterCells()
fm.addPointsFromGeoClusters()
m = fm.getMap()
m.save("/Users/tgadf/Downloads/oak-ridge-map.html")
#m
# ## Show Driver Network GeoSpatial Map
fm = foliumMap(dn=dn, gc=gc)
fm.createMap(zoom=11)
fm.addPointsFromDriverNetwork()
m = fm.getMap()
#m.save("oak-ridge-map-dn2.html")
m
# +
fm = foliumMap(pc=gc.getProtoClusters(), gc=gc)
fm.createMap()
fm.addPointsFromGeoClusterCells()
fm.addPointsFromProtoClusters()
m = fm.getMap()
m
fm = foliumMap(df=df)
fm.createMap()
fm.addPoints()
m = fm.getMap()
m.save("oak-ridge-map-df2.html")
fm = foliumMap(gc=gc, df=df)
fm.createMap()
fm.addPoints()
fm.addPointsFromTripsDataFrame()
m = fm.getMap()
m.save("oak-ridge-map-gc2.html")
# +
# %load_ext autoreload
# %autoreload
from networkOutput import printNetwork
pn = printNetwork(dn)
pn.printVertices(minN=40)
pn.printEdges(minW=20)
pn.printFrequencies()
# -
dn.getVertexByName('cl9')
vertexData = dn.getVertexByName('cl0', 'feat')
vertexData
|
createNetwork.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import scipy.optimize as opt
#2.6mm wide
#1.9mm verticle
#Taken at 640x480
mmperpix = .004
fps = 1000
n = 3 #Number of epx files
#epx files renamed test0.txt through testn.txt
dt = 1/fps
def filterindex(maj):
z = 0
for entry in maj:
if entry<2*maj[0] and entry>.5*maj[0] and entry<400:
"""When the particle tracker loses the particle it creates a very large ellipse where the minor axis is
about the width of the screen. Also if tracking a small particle that overlaps with a dust spec, sometimes
the tracker will shrink the size of the ellipse and fit it to the dust. This if statement along with the
list slicing below discard any entries in the position arrays after there has been an error in tracking"""
z = z + 1
return z
alldata = np.array([np.delete(np.loadtxt('fast' + str(j) +'.txt', unpack=True),0,0) for j in range(n)])
alldata = alldata*mmperpix
alldata = np.array([entry for box in alldata for entry in box])
N = int(len(alldata)/5)
allx = np.array([alldata[5*i] for i in range(N)])
ally = np.array([alldata[1+5*i] for i in range(N)])
allmaj = np.array([alldata[2+5*i] for i in range(N)])
allmin = np.array([alldata[3+5*i] for i in range(N)])
x = np.array([allx[i][0:filterindex(allmaj[i])] for i in range(len(allx))])
x = np.array([entry for entry in x if len(entry)>=10])
y = np.array([ally[i][0:filterindex(allmaj[i])] for i in range(len(ally))])
y = np.array([entry for entry in y if len(entry)>=10])
xavg = np.array([np.mean(entry) for entry in x])
alldy = np.array([np.gradient(entry) for entry in y])
dyavg = -np.array([np.mean(entry) for entry in alldy])
vyavg = dyavg/dt
particles = len(x)
particles
def u(x,dp):
b = 1.25*.001
mu = .045
return -(x/mu)*dp*(b-x/2)
theta_best, theta_cov = opt.curve_fit(u, xavg[vyavg>0]*.001, vyavg[vyavg>0]*.001)
dp = theta_best[0]
dp
print('dp/dx = {0:.3f} +/- {1:.3f}'.format(dp, np.sqrt(theta_cov[0,0])))
plt.title('Average Y Velocity vs Average X Position');
plt.xlabel('X Position (mm)');
plt.ylabel('Y Velocity (mm/s)');
plt.scatter(xavg[vyavg>.2], vyavg[vyavg>.2]);
plt.plot(np.linspace(0,2.5,1000),u(np.linspace(0,2.5*.001,1000),dp)*1000)
plt.tick_params(axis='x',top='off',direction='out')
plt.tick_params(axis='y',right='off',direction='out')
plt.text(.4, 5, 'dP/dx = {0:.3f} +/- {1:.3f} Pa/m'.format(dp, np.sqrt(theta_cov[0,0])));
plt.text(.4, 2.5, 'Number of Particles = {} '.format(particles));
plt.plot(np.zeros(5),np.linspace(0,35,5),color = 'black',linewidth = '6')
plt.plot(np.array([2.5]*5),np.linspace(0,35,5),color = 'black',linewidth = '6')
plt.xlim(0,2.5)
plt.ylim(0,35)
plt.savefig('Fastrun.png')
|
FilmChannel/FastRun.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + pycharm={"name": "#%%\n", "is_executing": false}
import numpy as np
from scipy.stats import norm
from scipy.integrate import quad
from scipy.optimize import root
# + pycharm={"is_executing": false}
"""
%% Summary of CJK_Func.m %%
The function generates the value of CJK representation.
Note that, this function is not used directly, it is used to solve for Bt
--------------------------------------------------------------------------
Input:
Bt - the value of boundary at t
Bs - the value of boundary at s
k - strike price
r - risk-free interest rate
q - continuously compounded dividend rate
vol - annualized volatility
T - maturity
t - current time
--------------------------------------------------------------------------
Output:
y - value of CJK reprentation
--------------------------------------------------------------------------
Author:
<NAME>
<EMAIL>
MSc Financial Engineering, ICMA Centre, Henley Business School,
University of Reading, UK
24 July 2015
--------------------------------------------------------------------------
"""
def d1(x,y,z,b,vol):
return (np.log(x/y)+(b+0.5*vol**2)*z)/(vol*np.sqrt(z))
def d2(x,y,z,b,vol):
return d1(x,y,z,b,vol)-vol*np.sqrt(z)
def CJK_Func( Bt,Bs,k,r,q,vol,T,t ):
T1 = T-t
b=r-q
term = np.zeros(5)
term[0] = Bt
term[1] = -k
term[2] = k*np.exp(-r*T1)*norm.cdf(-d2(Bt,k,T1,b,vol))
term[3] = -Bt*np.exp(-q*T1)*norm.cdf(-d1(Bt,k,T1,b,vol))
integralFun = lambda s: r*k*np.exp(-r*(s))*norm.cdf(-d2(Bt,Bs,(s),b,vol)) - q*Bt*np.exp(-q*(s))*norm.cdf(-d1(Bt,Bs,(s),b,vol))
term[4] = quad(integralFun,t,T)[0]
y = np.sum(term)
return y
# + pycharm={"name": "#%%\n", "is_executing": false}
"""
%% Summary of Boundary.m %%
The function generates the early exercise boundary and spot of time by
CJK representation
--------------------------------------------------------------------------
Input:
k - strike price
r - risk-free interest rate
q - continuously compounded dividend rate
vol - annualized volatility
T - time to maturity
steps - a number of time steps in the calculation
--------------------------------------------------------------------------
Output:
B - the values of early exercise boundary
time - the point of time that each B-value is calculated
--------------------------------------------------------------------------
Author:
<NAME>
<EMAIL>
MSc Financial Engineering, ICMA Centre, Henley Business School,
University of Reading, UK
24 July 2015
--------------------------------------------------------------------------
"""
def Boundary( k,r,q,vol,T,steps ):
dt=T/steps
t=T
B = np.zeros(steps+1)
time = np.linspace(0,T,np.floor(dt).astype(np.int))
for i in range(steps,1,-1):
if i==steps:
if q == 0:
B[i]=np.min(k,k*r/q)
else:
B[i]=k
else:
t=t-dt
res=root(lambda Bt: CJK_Func(Bt,B[i+1],k,r,q,vol,T,t) ,k)
B[i] = res.x
return B
# + pycharm={"name": "#%%\n", "is_executing": false}
s0 = 100
k = 100
r = 0.07
q = 0.03
vol = 0.25
T=1
#paths = 100000
steps = 10000
dt=T/steps
B = Boundary( k,r,q,vol,T,steps)
# + pycharm={"name": "#%%\n", "is_executing": false}
from matplotlib import pyplot as plt
plt.plot(np.linspace(0,1,10001),B)
# + pycharm={"name": "#%%\n", "is_executing": false}
from scipy.stats.distributions import norm, lognorm, rv_frozen
class GeometricBrownianMotion:
'''Geometric Brownian Motion.(with optional drift).'''
def __init__(self, mu: float=0.0, sigma: float=1.0):
self.mu = mu
self.sigma = sigma
def simulate(self, t: np.array, n: int, rnd: np.random.RandomState) \
-> np.array:
assert t.ndim == 1, 'One dimensional time vector required'
assert t.size > 0, 'At least one time point is required'
dt = np.concatenate((t[0:1], np.diff(t)))
assert (dt >= 0).all(), 'Increasing time vector required'
# transposed simulation for automatic broadcasting
dW = (rnd.normal(size=(t.size, n)).T * np.sqrt(dt)).T
W = np.cumsum(dW, axis=0)
return np.exp(self.sigma * W.T + (self.mu - self.sigma**2 / 2) * t).T
def distribution(self, t: float) -> rv_frozen:
mu_t = (self.mu - self.sigma**2/2) * t
sigma_t = self.sigma * np.sqrt(t)
return lognorm(scale=np.exp(mu_t), s=sigma_t)
# + pycharm={"name": "#%%\n", "is_executing": false}
from scipy.optimize import newton
class LS:
def __init__(self, X, t, r, strike):
self.X = X
self.t = t
self.r = r
self.strike = strike
def _ls_american_option_quadratic_iter(self, X, t, r, strike):
# given no prior exercise we just receive the payoff of a European option
cashflow = np.maximum(strike - X[-1, :], 0.0)
# iterating backwards in time
for i in reversed(range(1, X.shape[1] - 1)):
# discount factor between t[i] and t[i+1]
df = np.exp(-r * (t[i+1]-t[i]))
# discount cashflows from next period
cashflow = cashflow * df
x = X[:, i]
# exercise value for time t[i]
exercise = np.maximum(strike - x, 0.0)
# boolean index of all in-the-money paths
itm = exercise > 0
# fit polynomial of degree 2
fitted = Polynomial.fit(x[itm], cashflow[itm], 2)
# approximate continuation value
continuation = fitted(x)
# boolean index where exercise is beneficial
ex_idx = itm & (exercise > continuation)
# update cashflows with early exercises
cashflow[ex_idx] = exercise[ex_idx]
func = cashflow - strike
res = newton(func,strike)
yield res,cashflow, x, fitted, continuation, exercise, ex_idx
def simulate(self):
for res,cashflow, *_ in self._ls_american_option_quadratic_iter(self.X, self.t, self.r, self.strike):
pass
return res,cashflow.mean(axis=0) * np.exp(-self.r * (self.t[1] - self.t[0]))
# + pycharm={"name": "#%%\n"}
|
code/notebook/test_dev.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Variational Autoencoders
#
# ## Introduction
#
# The variational autoencoder (VAE) is arguably the simplest setup that realizes deep probabilistic modeling. Note that we're being careful in our choice of language here. The VAE isn't a model as such—rather the VAE is a particular setup for doing variational inference for a certain class of models. The class of models is quite broad: basically
# any (unsupervised) density estimator with latent random variables. The basic structure of such a model is simple, almost deceptively so (see Fig. 1).
# + raw_mimetype="text/html" active=""
# <center><figure><img src="_static/img/vae_model.png" style="width: 200px;"><figcaption> <font size="+1"><b>Figure 1</b>: the class of deep models we're interested in.</font></figcaption></figure></center><br>
# -
#
# Here we've depicted the structure of the kind of model we're interested in as a graphical model. We have $N$ observed datapoints $\{ \bf x_i \}$. Each datapoint is generated by a (local) latent random variable $\bf z_i$. There is also a parameter $\theta$, which is global in the sense that all the datapoints depend on it (which is why it's drawn outside the rectangle). Note that since $\theta$ is a parameter, it's not something we're being Bayesian about. Finally, what's of particular importance here is that we allow for each $\bf x_i$ to depend on $\bf z_i$ in a complex, non-linear way. In practice this dependency will be parameterized by a (deep) neural network with parameters $\theta$. It's this non-linearity that makes inference for this class of models particularly challenging.
#
# Of course this non-linear structure is also one reason why this class of models offers a very flexible approach to modeling complex data. Indeed it's worth emphasizing that each of the components of the model can be 'reconfigured' in a variety of different ways. For example:
#
# - the neural network in $p_\theta({\bf x} | {\bf z})$ can be varied in all the usual ways (number of layers, type of non-linearities, number of hidden units, etc.)
# - we can choose observation likelihoods that suit the dataset at hand: gaussian, bernoulli, categorical, etc.
# - we can choose the number of dimensions in the latent space
#
# The graphical model representation is a useful way to think about the structure of the model, but it can also be fruitful to look at an explicit factorization of the joint probability density:
#
# $$ p({\bf x}, {\bf z}) = \prod_{i=1}^N p_\theta({\bf x}_i | {\bf z}_i) p({\bf z}_i) $$
#
# The fact that $p({\bf x}, {\bf z})$ breaks up into a product of terms like this makes it clear what we mean when we call $\bf z_i$ a local random variable. For any particular $i$, only the single datapoint $\bf x_i$ depends on $\bf z_i$. As such the $\{\bf z_i\}$ describe local structure, i.e. structure that is private to each data point. This factorized structure also means that we can do subsampling during the course of learning. As such this sort of model is amenable to the large data setting. (For more discussion on this and related topics see [SVI Part II](svi_part_ii.ipynb).)
#
# That's all there is to the model. Since the observations depend on the latent random variables in a complicated, non-linear way, we expect the posterior over the latents to have a complex structure. Consequently in order to do inference in this model we need to specify a flexibly family of guides (i.e. variational distributions). Since we want to be able to scale to large datasets, our guide is going to make use of amortization to keep the number of variational parameters under control (see [SVI Part II](svi_part_ii.ipynb) for a somewhat more general discussion of amortization).
#
# Recall that the job of the guide is to 'guess' good values for the latent random variables—good in the sense that they're true to the model prior _and_ true to the data. If we weren't making use of amortization, we would introduce variational parameters
# $\{ \lambda_i \}$ for _each_ datapoint $\bf x_i$. These variational parameters would represent our belief about 'good' values of $\bf z_i$; for example, they could encode the mean and variance of a gaussian distribution in ${\bf z}_i$ space. Amortization means that, rather than introducing variational parameters $\{ \lambda_i \}$, we instead learn a _function_ that maps each $\bf x_i$ to an appropriate $\lambda_i$. Since we need this function to be flexible, we parameterize it as a neural network. We thus end up with a parameterized family of distributions over the latent $\bf z$ space that can be instantiated for all $N$ datapoint ${\bf x}_i$ (see Fig. 2).
# + raw_mimetype="text/html" active=""
# <center><figure><img src="_static/img/vae_guide.png" style="width: 200px;"><figcaption> <font size="+1"><b>Figure 2</b>: a graphical representation of the guide. </font></figcaption></figure></center><br>
# -
# Note that the guide $q_{\phi}({\bf z} | {\bf x})$ is parameterized by a global parameter $\phi$ shared by all the datapoints. The goal of inference will be to find 'good' values for $\theta$ and $\phi$ so that two conditions are satisfied:
#
# - the log evidence $\log p_\theta({\bf x})$ is large. this means our model is a good fit to the data
# - the guide $q_{\phi}({\bf z} | {\bf x})$ provides a good approximation to the posterior
#
# (For an introduction to stochastic variational inference see [SVI Part I](svi_part_i.ipynb).)
#
# At this point we can zoom out and consider the high level structure of our setup. For concreteness, let's suppose the $\{ \bf x_i \}$ are images so that the model is a generative model of images. Once we've learned a good value of $\theta$ we can generate images from the model as follows:
#
# - sample $\bf z$ according to the prior $p({\bf z})$
# - sample $\bf x$ according to the likelihood $p_\theta({\bf x}|{\bf z})$
#
# Each image is being represented by a latent code $\bf z$ and that code gets mapped to images using the likelihood, which depends on the $\theta$ we've learned. This is why the likelihood is often called the decoder in this context: its job is to decode $\bf z$ into $\bf x$. Note that since this is a probabilistic model, there is uncertainty about the $\bf z$ that encodes a given datapoint $\bf x$.
#
# Once we've learned good values for $\theta$ and $\phi$ we can also go through the following exercise.
#
# - we start with a given image $\bf x$
# - using our guide we encode it as $\bf z$
# - using the model likelihood we decode $\bf z$ and get a reconstructed image ${\bf x}_\rm{reco}$
#
# If we've learned good values for $\theta$ and $\phi$, $\bf x$ and ${\bf x}_\rm{reco}$ should be similar. This should clarify how the word autoencoder ended up being used to describe this setup: the model is the decoder and the guide is the encoder. Together, they can be thought of as an autoencoder.
#
# ## VAE in Pyro
#
# Let's see how we implement a VAE in Pyro.
# The dataset we're going to model is MNIST, a collection of images of handwritten digits.
# Since this is a popular benchmark dataset, we can make use of PyTorch's convenient data loader functionalities to reduce the amount of boilerplate code we need to write:
# +
import os
import numpy as np
import torch
import torchvision.datasets as dset
import torch.nn as nn
import torchvision.transforms as transforms
import pyro
import pyro.distributions as dist
from pyro.infer import SVI, Trace_ELBO
from pyro.optim import Adam
# -
assert pyro.__version__.startswith('1.0.0')
pyro.enable_validation(True)
pyro.distributions.enable_validation(False)
pyro.set_rng_seed(0)
# Enable smoke test - run the notebook cells on CI.
smoke_test = 'CI' in os.environ
# for loading and batching MNIST dataset
def setup_data_loaders(batch_size=128, use_cuda=False):
root = './data'
download = True
trans = transforms.ToTensor()
train_set = dset.MNIST(root=root, train=True, transform=trans,
download=download)
test_set = dset.MNIST(root=root, train=False, transform=trans)
kwargs = {'num_workers': 1, 'pin_memory': use_cuda}
train_loader = torch.utils.data.DataLoader(dataset=train_set,
batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(dataset=test_set,
batch_size=batch_size, shuffle=False, **kwargs)
return train_loader, test_loader
# The main thing to draw attention to here is that we use `transforms.ToTensor()` to normalize the pixel intensities to the range $[0.0, 1.0]$.
#
# Next we define a PyTorch module that encapsulates our decoder network:
class Decoder(nn.Module):
def __init__(self, z_dim, hidden_dim):
super(Decoder, self).__init__()
# setup the two linear transformations used
self.fc1 = nn.Linear(z_dim, hidden_dim)
self.fc21 = nn.Linear(hidden_dim, 784)
# setup the non-linearities
self.softplus = nn.Softplus()
self.sigmoid = nn.Sigmoid()
def forward(self, z):
# define the forward computation on the latent z
# first compute the hidden units
hidden = self.softplus(self.fc1(z))
# return the parameter for the output Bernoulli
# each is of size batch_size x 784
loc_img = self.sigmoid(self.fc21(hidden))
return loc_img
# Given a latent code $z$, the forward call of `Decoder` returns the parameters for a Bernoulli distribution in image space. Since each image is of size
# $28\times28=784$, `loc_img` is of size `batch_size` x 784.
#
# Next we define a PyTorch module that encapsulates our encoder network:
class Encoder(nn.Module):
def __init__(self, z_dim, hidden_dim):
super(Encoder, self).__init__()
# setup the three linear transformations used
self.fc1 = nn.Linear(784, hidden_dim)
self.fc21 = nn.Linear(hidden_dim, z_dim)
self.fc22 = nn.Linear(hidden_dim, z_dim)
# setup the non-linearities
self.softplus = nn.Softplus()
def forward(self, x):
# define the forward computation on the image x
# first shape the mini-batch to have pixels in the rightmost dimension
x = x.reshape(-1, 784)
# then compute the hidden units
hidden = self.softplus(self.fc1(x))
# then return a mean vector and a (positive) square root covariance
# each of size batch_size x z_dim
z_loc = self.fc21(hidden)
z_scale = torch.exp(self.fc22(hidden))
return z_loc, z_scale
# Given an image $\bf x$ the forward call of `Encoder` returns a mean and covariance that together parameterize a (diagonal) Gaussian distribution in latent space.
#
# With our encoder and decoder networks in hand, we can now write down the stochastic functions that represent our model and guide. First the model:
# define the model p(x|z)p(z)
def model(self, x):
# register PyTorch module `decoder` with Pyro
pyro.module("decoder", self.decoder)
with pyro.plate("data", x.shape[0]):
# setup hyperparameters for prior p(z)
z_loc = x.new_zeros(torch.Size((x.shape[0], self.z_dim)))
z_scale = x.new_ones(torch.Size((x.shape[0], self.z_dim)))
# sample from prior (value will be sampled by guide when computing the ELBO)
z = pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# decode the latent code z
loc_img = self.decoder.forward(z)
# score against actual images
pyro.sample("obs", dist.Bernoulli(loc_img).to_event(1), obs=x.reshape(-1, 784))
# Note that `model()` is a callable that takes in a mini-batch of images `x` as input. This is a `torch.Tensor` of size `batch_size` x 784.
#
# The first thing we do inside of `model()` is register the (previously instantiated) decoder module with Pyro. Note that we give it an appropriate (and unique) name. This call to `pyro.module` lets Pyro know about all the parameters inside of the decoder network.
#
# Next we setup the hyperparameters for our prior, which is just a unit normal gaussian distribution. Note that:
# - we specifically designate independence amongst the data in our mini-batch (i.e. the leftmost dimension) via `pyro.plate`. Also, note the use of `.to_event(1)` when sampling from the latent `z` - this ensures that instead of treating our sample as being generated from a univariate normal with `batch_size = z_dim`, we treat them as being generated from a multivariate normal distribution with diagonal covariance. As such, the log probabilities along each dimension is summed out when we evaluate `.log_prob` for a "latent" sample. Refer to the [Tensor Shapes](tensor_shapes.ipynb) tutorial for more details.
# - since we're processing an entire mini-batch of images, we need the leftmost dimension of `z_loc` and `z_scale` to equal the mini-batch size
# - in case we're on GPU, we use `new_zeros` and `new_ones` to ensure that newly created tensors are on the same GPU device.
#
# Next we sample the latent `z` from the prior, making sure to give the random variable a unique Pyro name `'latent'`.
# Then we pass `z` through the decoder network, which returns `loc_img`. We then score the observed images in the mini-batch `x` against the Bernoulli likelihood parametrized by `loc_img`.
# Note that we flatten `x` so that all the pixels are in the rightmost dimension.
#
# That's all there is to it! Note how closely the flow of Pyro primitives in `model` follows the generative story of our model, e.g. as encapsulated by Figure 1. Now we move on to the guide:
# define the guide (i.e. variational distribution) q(z|x)
def guide(self, x):
# register PyTorch module `encoder` with Pyro
pyro.module("encoder", self.encoder)
with pyro.plate("data", x.shape[0]):
# use the encoder to get the parameters used to define q(z|x)
z_loc, z_scale = self.encoder.forward(x)
# sample the latent code z
pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# Just like in the model, we first register the PyTorch module we're using (namely `encoder`) with Pyro. We take the mini-batch of images `x` and pass it through the encoder. Then using the parameters output by the encoder network we use the normal distribution to sample a value of the latent for each image in the mini-batch. Crucially, we use the same name for the latent random variable as we did in the model: `'latent'`. Also, note the use of `pyro.plate` to designate independence of the mini-batch dimension, and `.to_event(1)` to enforce dependence on `z_dims`, exactly as we did in the model.
#
# Now that we've defined the full model and guide we can move on to inference. But before we do so let's see how we package the model and guide in a PyTorch module:
class VAE(nn.Module):
# by default our latent space is 50-dimensional
# and we use 400 hidden units
def __init__(self, z_dim=50, hidden_dim=400, use_cuda=False):
super(VAE, self).__init__()
# create the encoder and decoder networks
self.encoder = Encoder(z_dim, hidden_dim)
self.decoder = Decoder(z_dim, hidden_dim)
if use_cuda:
# calling cuda() here will put all the parameters of
# the encoder and decoder networks into gpu memory
self.cuda()
self.use_cuda = use_cuda
self.z_dim = z_dim
# define the model p(x|z)p(z)
def model(self, x):
# register PyTorch module `decoder` with Pyro
pyro.module("decoder", self.decoder)
with pyro.plate("data", x.shape[0]):
# setup hyperparameters for prior p(z)
z_loc = x.new_zeros(torch.Size((x.shape[0], self.z_dim)))
z_scale = x.new_ones(torch.Size((x.shape[0], self.z_dim)))
# sample from prior (value will be sampled by guide when computing the ELBO)
z = pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# decode the latent code z
loc_img = self.decoder.forward(z)
# score against actual images
pyro.sample("obs", dist.Bernoulli(loc_img).to_event(1), obs=x.reshape(-1, 784))
# define the guide (i.e. variational distribution) q(z|x)
def guide(self, x):
# register PyTorch module `encoder` with Pyro
pyro.module("encoder", self.encoder)
with pyro.plate("data", x.shape[0]):
# use the encoder to get the parameters used to define q(z|x)
z_loc, z_scale = self.encoder.forward(x)
# sample the latent code z
pyro.sample("latent", dist.Normal(z_loc, z_scale).to_event(1))
# define a helper function for reconstructing images
def reconstruct_img(self, x):
# encode image x
z_loc, z_scale = self.encoder(x)
# sample in latent space
z = dist.Normal(z_loc, z_scale).sample()
# decode the image (note we don't sample in image space)
loc_img = self.decoder(z)
return loc_img
# The point we'd like to make here is that the two `Module`s `encoder` and `decoder` are attributes of `VAE` (which itself inherits from `nn.Module`). This has the consequence they are both automatically registered as belonging to the `VAE` module. So, for example, when we call `parameters()` on an instance of `VAE`, PyTorch will know to return all the relevant parameters. It also means that if we're running on a GPU, the call to `cuda()` will move all the parameters of all the (sub)modules into GPU memory.
# ## Inference
#
# We're now ready for inference. Refer to the full code in the next section.
#
# First we instantiate an instance of the `VAE` module.
vae = VAE()
# Then we setup an instance of the Adam optimizer.
optimizer = Adam({"lr": 1.0e-3})
# Then we setup our inference algorithm, which is going to learn good parameters for the model and guide by maximizing the ELBO:
#
svi = SVI(vae.model, vae.guide, optimizer, loss=Trace_ELBO())
# That's all there is to it. Now we just have to define our training loop:
def train(svi, train_loader, use_cuda=False):
# initialize loss accumulator
epoch_loss = 0.
# do a training epoch over each mini-batch x returned
# by the data loader
for x, _ in train_loader:
# if on GPU put mini-batch into CUDA memory
if use_cuda:
x = x.cuda()
# do ELBO gradient and accumulate loss
epoch_loss += svi.step(x)
# return epoch loss
normalizer_train = len(train_loader.dataset)
total_epoch_loss_train = epoch_loss / normalizer_train
return total_epoch_loss_train
# Note that all the mini-batch logic is handled by the data loader. The meat of the training loop is `svi.step(x)`. There are two things we should draw attention to here:
#
# - any arguments to `step` are passed to the model and the guide. consequently `model` and `guide` need to have the same call signature
# - `step` returns a noisy estimate of the loss (i.e. minus the ELBO). this estimate is not normalized in any way, so e.g. it scales with the size of the mini-batch
#
# The logic for adding evaluation logic is analogous:
def evaluate(svi, test_loader, use_cuda=False):
# initialize loss accumulator
test_loss = 0.
# compute the loss over the entire test set
for x, _ in test_loader:
# if on GPU put mini-batch into CUDA memory
if use_cuda:
x = x.cuda()
# compute ELBO estimate and accumulate loss
test_loss += svi.evaluate_loss(x)
normalizer_test = len(test_loader.dataset)
total_epoch_loss_test = test_loss / normalizer_test
return total_epoch_loss_test
# Basically the only change we need to make is that we call evaluate_loss instead of step. This function will compute an estimate of the ELBO but won't take any gradient steps.
#
# The final piece of code we'd like to highlight is the helper method `reconstruct_img` in the VAE class: This is just the image reconstruction experiment we described in the introduction translated into code. We take an image and pass it through the encoder. Then we sample in latent space using the gaussian distribution provided by the encoder. Finally we decode the latent code into an image: we return the mean vector `loc_img` instead of sampling with it. Note that since the `sample()` statement is stochastic, we'll get different draws of z every time we run the reconstruct_img function. If we've learned a good model and guide—in particular if we've learned a good latent representation—this plurality of z samples will correspond to different styles of digit writing, and the reconstructed images should exhibit an interesting variety of different styles.
# ## Code and Sample results
#
# Training corresponds to maximizing the evidence lower bound (ELBO) over the training dataset. We train for 100 iterations and evaluate the ELBO for the test dataset, see Figure 3.
# +
# Run options
LEARNING_RATE = 1.0e-3
USE_CUDA = False
# Run only for a single iteration for testing
NUM_EPOCHS = 1 if smoke_test else 100
TEST_FREQUENCY = 5
# +
train_loader, test_loader = setup_data_loaders(batch_size=256, use_cuda=USE_CUDA)
# clear param store
pyro.clear_param_store()
# setup the VAE
vae = VAE(use_cuda=USE_CUDA)
# setup the optimizer
adam_args = {"lr": LEARNING_RATE}
optimizer = Adam(adam_args)
# setup the inference algorithm
svi = SVI(vae.model, vae.guide, optimizer, loss=Trace_ELBO())
train_elbo = []
test_elbo = []
# training loop
for epoch in range(NUM_EPOCHS):
total_epoch_loss_train = train(svi, train_loader, use_cuda=USE_CUDA)
train_elbo.append(-total_epoch_loss_train)
print("[epoch %03d] average training loss: %.4f" % (epoch, total_epoch_loss_train))
if epoch % TEST_FREQUENCY == 0:
# report test diagnostics
total_epoch_loss_test = evaluate(svi, test_loader, use_cuda=USE_CUDA)
test_elbo.append(-total_epoch_loss_test)
print("[epoch %03d] average test loss: %.4f" % (epoch, total_epoch_loss_test))
# + raw_mimetype="text/html" active=""
# <center>
# <figure>
# <img src="_static/img/vae_plots/test_elbo_vae.png" style="width: 550px;">
# <figcaption>
# <font size="+1"><b>Figure 3:</b> How the test ELBO evolves over the course of training. </font>
# </figcaption>
# </figure>
# </center>
# -
# Next we show a set of randomly sampled images from the model. These are generated by drawing random samples of `z` and generating an image for each one, see Figure 4.
# + raw_mimetype="text/html" active=""
# <center>
# <figure>
# <table>
# <tr>
# <td>
# <img src="_static/img/vae_plots/vae_embeddings_pt1.jpg" style="width: 350px;">
# </td>
# <td>
# <img src="_static/img/vae_plots/vae_embeddings_pt2.jpg" style="width: 350px;">
# </td>
# </tr>
# </table>
# <figcaption>
# <font size="+1"><b>Figure 4:</b> Samples from generative model.</font>
# </figcaption>
# </figure>
# </center>
# -
# We also study the 50-dimensional latent space of the entire test dataset by encoding all MNIST images and embedding their means into a 2-dimensional T-SNE space. We then color each embedded image by its class.
# The resulting Figure 5 shows separation by class with variance within each class-cluster.
# + raw_mimetype="text/html" active=""
# <center>
# <figure>
# <img src="_static/img/vae_plots/VAE_embedding.png" style="width: 550px;">
# <figcaption>
# <font size="+1"><b>Figure 5:</b> T-SNE Embedding of the latent z. The colors correspond to different classes of digits.</font>
# </figcaption>
# </figure>
# </center>
# -
# See the full code on [Github](https://github.com/uber/pyro/blob/dev/examples/vae/vae.py).
#
# ## References
#
# [1] `Auto-Encoding Variational Bayes`,<br/>
# <NAME>, <NAME>
#
# [2] `Stochastic Backpropagation and Approximate Inference in Deep Generative Models`,
# <br/>
# <NAME>, <NAME>, <NAME>
|
tutorial/source/vae.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # San Francisco Rental Prices Dashboard
# ____________________________________________
import panel as pn
pn.extension('plotly')
import plotly.express as px
import pandas as pd
import hvplot.pandas
import matplotlib.pyplot as plt
import os
from pathlib import Path
from dotenv import load_dotenv
# +
# Read the Mapbox API key
map_box_api = os.getenv("MAPBOX_API_KEY")
px.set_mapbox_access_token(map_box_api)
# +
def housing_units_per_year():
housing_units_yearly = census.groupby('year')['housing_units'].mean().hvplot.bar(xlabel="Year", ylabel="Housing Units",ylim=(370000,385000), title="Housing Units Per Year")
return housing_units_yearly
def average_gross_rent():
average_rent = census.groupby('year')['gross_rent'].mean().hvplot.line(xlabel='Year', ylabel='Gross Rent', title='Average Gross Rent in SF')
return average_rent
def average_sales_price():
average_sales = census.groupby('year')['sale_price_sqr_foot'].mean().hvplot.line(xlabel='Year', ylabel='Average Sales Price', title='Average Sale Price Per Year')
return average_sales
def average_price_by_neighborhood():
average_price = neighborhood_df.hvplot.line(x='year', y='sale_price_sqr_foot', groupby='neighborhood',title="Average Prices by Neighborhood.")
return average_price
def top_most_expensive_neighborhoods():
highest_priced_neighborhoods = top10_df_plot.head(10).plot(kind="bar",x="neighborhood",y="sale_price_sqr_foot",title="Top 10 Most Expensive Neighborhoods")
return highest_priced_neighborhoods
def get_parallel_coordinates():
parallel_coordinates = px.parallel_coordinates(top10_df_plot, color="sale_price_sqr_foot",title="Parallel Coordinates Plot")
return parallel_coordinates
def get_parallel_categories():
parallel_categories = px.parallel_categories(top10_df_plot,dimensions=["neighborhood","sale_price_sqr_foot","housing_units","gross_rent"],
color="sale_price_sqr_foot",title="Parallel Categories Plot.")
return parallel_categories
def get_neighborhood_map():
neighborhood_map = px.scatter_mapbox(joined_df,
lat="Lat",
lon="Lon",
size="sale_price_sqr_foot",
color="gross_rent",
title="Average Values Per Neighborhood In San Francisco")
return neighborhood_map
# -
# # Panel Dashboard
# +
#Basic data column
housing_data = pn.Column(
"##SF Housing Data",
housing_units_per_year(),
average_gross_rent(),
average_sales_price(),
)
#Neighborhood column
neigborhood_data = pn.Column(
"##SF Neighborhood Data",
average_price_by_neighborhood(),
top_most_expensive_neighborhoods(),
)
#Interactive column
interactive_data = pn.Column(
"##SF Interactive Data",
get_parallel_coordinates(),
get_parallel_categories()
)
#Map column
map_data = pn.Column(
"##SF Map",
get_neighborhood_map(),
)
# Create tabs
dashboard = pn.Tabs(
("Housing Data", housing_data), ("Neighborhood Data", neigborhood_data), ("Interactive Data", interactive_data), ("SF Map", map_data)
)
# -
dashboard.servable()
|
Dashboard.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This is the Jupyter Notebook for the MAGICODE project
# ### First install some modules that might not be installed
# Install a pip package in the current Jupyter kernel
# https://jakevdp.github.io/blog/2017/12/05/installing-python-packages-from-jupyter/
import sys
# !{sys.executable} -m pip install tensorflow
# install sklearn to use train_test_split function
# !{sys.executable} -m pip install sklearn
# install opencv to use cv2 module in get_preprocessed_img
# !{sys.executable} -m pip install opencv-python
# ### Then import some libraries and modules that are needed for the code to run
import os
import sklearn.model_selection as model_selection
import glob
import shutil
import datetime
import pydot
import numpy as np
from pathlib import Path
from os.path import join
# ### Define some values used later
SOURCE = 'dataset'
TRAINING_SET_NAME = 'training_set'
EVALUATION_SET_NAME = 'eval_set'
TRAINING_FEATURES = 'training_features'
EVAL_FEATURES = 'eval_features'
START_TOKEN = "<START>"
END_TOKEN = "<END>"
PLACEHOLDER = " "
SEPARATOR = '->'
CHECKPOINT_DIR = 'training'
CHECKPOINT_FILE_NAME = 'weights_checkpoint'
CHECKPOINT_FILE = join(CHECKPOINT_FILE_NAME + '.hdf5')
CHECKPOINT_ARCHIVE = join(CHECKPOINT_FILE_NAME + '.zip')
CHECKPOINT_PATH = join(CHECKPOINT_DIR, CHECKPOINT_FILE)
CHECKPOINT_ARCHIVE_PATH = join(CHECKPOINT_DIR, CHECKPOINT_ARCHIVE)
# ================================ HYPERPARAMETERS ================================
EPOCHS = 30 # Using early stopping, so training might run for less than X epochs
IMAGE_SIZE = 256
BATCH_SIZE = 128
CONTEXT_LENGTH = 50
PATIENCE = 5
LEARNING_RATE = 0.0001
LR_PATIENCE = 2
CLIPVALUE = 1.0
# ### Unzip the dataset
# +
# join the volumes together into a single zip file
# !zip -s 0 dataset.zip -O dataset_joined.zip
# unzip the newly assembled archive into the current folder
# !unzip dataset_joined.zip -d ./
if os.listdir(SOURCE):
print ('files unzipped')
# -
# ### Split dataset into training and evaluation sets
# +
# get all file paths
all_files = os.listdir(SOURCE)
# build a generic image path (e.g. 'all_data/dataset/*.png')
images_path = join(SOURCE, '*.png')
# get all images paths
img_files = glob.glob(images_path)
# remove files extension from files paths
img_files_without_extension = [Path(img_file).stem for img_file in img_files]
# splits randomly the files into two sets (train_set = 85% of dataset, eval_set = 15% of dataset)
train_set,eval_set = model_selection.train_test_split(img_files_without_extension, train_size=0.85)
# create the TRAINING_SET_NAME and EVALUATION_SET_NAME directories if they do not exist
if not os.path.exists(join(SOURCE, TRAINING_SET_NAME)):
os.makedirs(join(SOURCE, TRAINING_SET_NAME))
if not os.path.exists(join(SOURCE, EVALUATION_SET_NAME)):
os.makedirs(join(SOURCE, EVALUATION_SET_NAME))
# # copy the files (img and gui) from the all_data folder into the training_set folder
for file in train_set:
shutil.copyfile(join(SOURCE, file + '.png'), join(SOURCE, TRAINING_SET_NAME, file + '.png'))
shutil.copyfile(join(SOURCE, file + '.gui'), join(SOURCE, TRAINING_SET_NAME, file + '.gui'))
# # copy the files (img and gui) from the all_data folder into the eval_set folder
for file in eval_set:
shutil.copyfile(join(SOURCE, file + '.png'), join(SOURCE, EVALUATION_SET_NAME, file + '.png'))
shutil.copyfile(join(SOURCE, file + '.gui'), join(SOURCE, EVALUATION_SET_NAME, file + '.gui'))
print('Training dataset: {}'.format(join(SOURCE, TRAINING_SET_NAME)))
print('Evaluation dataset: {}'.format(join(SOURCE, EVALUATION_SET_NAME)))
# -
# ### Define some Classes and functions that will be used a few times
# +
class Utils:
@staticmethod
def sparsify(label_vector, output_size):
sparse_vector = []
for label in label_vector:
sparse_label = np.zeros(output_size)
sparse_label[label] = 1
sparse_vector.append(sparse_label)
return np.array(sparse_vector)
@staticmethod
def get_preprocessed_img(img_path, image_size):
import cv2
img = cv2.imread(img_path)
if not img is None:
img = cv2.resize(img, (image_size, image_size))
img = img.astype('float32')
img /= 255
return img
class Sampler:
def __init__(self, voc_path, input_shape, output_size, context_length):
self.voc = Vocabulary()
self.voc.retrieve(voc_path)
self.input_shape = input_shape
self.output_size = output_size
print('Vocabulary size: {}'.format(self.voc.size))
print('Input shape: {}'.format(self.input_shape))
print('Output size: {}'.format(self.output_size))
self.context_length = context_length
def predict_greedy(self, model, input_img, require_sparse_label=True, sequence_length=150, verbose=False):
current_context = [self.voc.vocabulary[PLACEHOLDER]] * (self.context_length - 1)
current_context.append(self.voc.vocabulary[START_TOKEN])
if require_sparse_label:
current_context = Utils.sparsify(current_context, self.output_size)
predictions = START_TOKEN
out_probas = []
for i in range(0, sequence_length):
if verbose:
print('predicting {}/{}...'.format(i, sequence_length))
probas = model.predict(input_img, np.array([current_context]))
prediction = np.argmax(probas)
out_probas.append(probas)
new_context = []
for j in range(1, self.context_length):
new_context.append(current_context[j])
if require_sparse_label:
sparse_label = np.zeros(self.output_size)
sparse_label[prediction] = 1
new_context.append(sparse_label)
else:
new_context.append(prediction)
current_context = new_context
predictions += self.voc.token_lookup[prediction]
if self.voc.token_lookup[prediction] == END_TOKEN:
break
return predictions, out_probas
class Vocabulary:
def __init__(self):
self.binary_vocabulary = {}
self.vocabulary = {}
self.token_lookup = {}
self.size = 0
self.append(START_TOKEN)
self.append(END_TOKEN)
self.append(PLACEHOLDER)
def append(self, token):
if token not in self.vocabulary:
self.vocabulary[token] = self.size
self.token_lookup[self.size] = token
self.size += 1
def create_binary_representation(self):
if sys.version_info >= (3,):
items = self.vocabulary.items()
else:
items = self.vocabulary.iteritems()
for key, value in items:
binary = np.zeros(self.size)
binary[value] = 1
self.binary_vocabulary[key] = binary
def get_serialized_binary_representation(self):
if len(self.binary_vocabulary) == 0:
self.create_binary_representation()
string = ''
if sys.version_info >= (3,):
items = self.binary_vocabulary.items()
else:
items = self.binary_vocabulary.iteritems()
for key, value in items:
array_as_string = np.array2string(value, separator=',', max_line_width=self.size * self.size)
string += '{}{}{}\n'.format(key, SEPARATOR, array_as_string[1:len(array_as_string) - 1])
return string
def save(self, path):
output_file_name = '{}/words.vocab'.format(path)
output_file = open(output_file_name, 'w')
output_file.write(self.get_serialized_binary_representation())
output_file.close()
def retrieve(self, path):
input_file = open('{}/words.vocab'.format(path), 'r')
buffer = ''
for line in input_file:
try:
separator_position = len(buffer) + line.index(SEPARATOR)
buffer += line
key = buffer[:separator_position]
value = buffer[separator_position + len(SEPARATOR):]
value = np.fromstring(value, sep=',')
self.binary_vocabulary[key] = value
self.vocabulary[key] = np.where(value == 1)[0][0]
self.token_lookup[np.where(value == 1)[0][0]] = key
buffer = ''
except ValueError:
buffer += line
input_file.close()
self.size = len(self.vocabulary)
class Generator:
@staticmethod
def data_generator(voc, gui_paths, img_paths, batch_size, generate_binary_sequences=False, verbose=False, loop_only_one=False):
assert len(gui_paths) == len(img_paths)
voc.create_binary_representation()
while 1:
batch_input_images = []
batch_partial_sequences = []
batch_next_words = []
sample_in_batch_counter = 0
for i in range(0, len(gui_paths)):
if img_paths[i].find('.png') != -1:
img = Utils.get_preprocessed_img(img_paths[i], IMAGE_SIZE)
else:
img = np.load(img_paths[i])['features']
gui = open(gui_paths[i], 'r')
token_sequence = [START_TOKEN]
for line in gui:
line = line.replace(',', ' ,').replace('\n', ' \n')
tokens = line.split(' ')
for token in tokens:
voc.append(token)
token_sequence.append(token)
token_sequence.append(END_TOKEN)
suffix = [PLACEHOLDER] * CONTEXT_LENGTH
a = np.concatenate([suffix, token_sequence])
for j in range(0, len(a) - CONTEXT_LENGTH):
context = a[j:j + CONTEXT_LENGTH]
label = a[j + CONTEXT_LENGTH]
batch_input_images.append(img)
batch_partial_sequences.append(context)
batch_next_words.append(label)
sample_in_batch_counter += 1
if sample_in_batch_counter == batch_size or (loop_only_one and i == len(gui_paths) - 1):
if verbose:
print('Generating sparse vectors...')
batch_next_words = Dataset.sparsify_labels(batch_next_words, voc)
if generate_binary_sequences:
batch_partial_sequences = Dataset.binarize(batch_partial_sequences, voc)
else:
batch_partial_sequences = Dataset.indexify(batch_partial_sequences, voc)
if verbose:
print('Convert arrays...')
batch_input_images = np.array(batch_input_images)
batch_partial_sequences = np.array(batch_partial_sequences)
batch_next_words = np.array(batch_next_words)
if verbose:
print('Yield batch')
yield ([batch_input_images, batch_partial_sequences], batch_next_words)
batch_input_images = []
batch_partial_sequences = []
batch_next_words = []
sample_in_batch_counter = 0
class Dataset:
def __init__(self):
self.input_shape = None
self.output_size = None
self.ids = []
self.input_images = []
self.partial_sequences = []
self.next_words = []
self.voc = Vocabulary()
self.size = 0
self.dataset_name = ''
@staticmethod
def load_paths_only(path):
print('Parsing data...')
gui_paths = []
img_paths = []
for f in os.listdir(path):
if f.find('.gui') != -1:
path_gui = join(path, f)
gui_paths.append(path_gui)
file_name = f[:f.find('.gui')]
if os.path.isfile(join(path, file_name + '.png')):
path_img = join(path, file_name + '.png')
img_paths.append(path_img)
elif os.path.isfile(join(path, file_name + '.npz')):
path_img = join(path, file_name + '.npz')
img_paths.append(path_img)
assert len(gui_paths) == len(img_paths)
return gui_paths, img_paths
def load(self, path, generate_binary_sequences=False):
print('Loading {} data...'.format(self.dataset_name))
for f in os.listdir(path):
if f.find('.gui') != -1:
gui = open(join(path, f), 'r')
file_name = Path(f).stem
if os.path.isfile(join(path, file_name + '.png')):
img = Utils.get_preprocessed_img(join(path, file_name + '.png'), IMAGE_SIZE)
self.append(file_name, gui, img)
elif os.path.isfile(join(path, file_name + '.npz')):
img = np.load(join(path, file_name + '.npz'))['features']
self.append(file_name, gui, img)
print('Generating sparse vectors...')
self.voc.create_binary_representation()
self.next_words = self.sparsify_labels(self.next_words, self.voc)
if generate_binary_sequences:
self.partial_sequences = self.binarize(self.partial_sequences, self.voc)
else:
self.partial_sequences = self.indexify(self.partial_sequences, self.voc)
self.size = len(self.ids)
assert self.size == len(self.input_images) == len(self.partial_sequences) == len(self.next_words)
assert self.voc.size == len(self.voc.vocabulary)
print('Dataset size: {}'.format(self.size))
print('Vocabulary size: {}'.format(self.voc.size))
self.input_shape = self.input_images[0].shape
self.output_size = self.voc.size
print('Input shape: {}'.format(self.input_shape))
print('Output size: {}'.format(self.output_size))
def convert_arrays(self):
print('Convert arrays...')
self.input_images = np.array(self.input_images)
self.partial_sequences = np.array(self.partial_sequences)
self.next_words = np.array(self.next_words)
def append(self, sample_id, gui, img, to_show=False):
if to_show:
pic = img * 255
pic = np.array(pic, dtype=np.uint8)
Utils.show(pic)
token_sequence = [START_TOKEN]
for line in gui:
line = line.replace(',', ' ,').replace('\n', ' \n')
tokens = line.split(' ')
for token in tokens:
self.voc.append(token)
token_sequence.append(token)
token_sequence.append(END_TOKEN)
suffix = [PLACEHOLDER] * CONTEXT_LENGTH
a = np.concatenate([suffix, token_sequence])
for j in range(0, len(a) - CONTEXT_LENGTH):
context = a[j:j + CONTEXT_LENGTH]
label = a[j + CONTEXT_LENGTH]
self.ids.append(sample_id)
self.input_images.append(img)
self.partial_sequences.append(context)
self.next_words.append(label)
@staticmethod
def indexify(partial_sequences, voc):
temp = []
for sequence in partial_sequences:
sparse_vectors_sequence = []
for token in sequence:
sparse_vectors_sequence.append(voc.vocabulary[token])
temp.append(np.array(sparse_vectors_sequence))
return temp
@staticmethod
def binarize(partial_sequences, voc):
temp = []
for sequence in partial_sequences:
sparse_vectors_sequence = []
for token in sequence:
sparse_vectors_sequence.append(voc.binary_vocabulary[token])
temp.append(np.array(sparse_vectors_sequence))
return temp
@staticmethod
def sparsify_labels(next_words, voc):
temp = []
for label in next_words:
temp.append(voc.binary_vocabulary[label])
return temp
def save_metadata(self, path):
np.save(join(path, 'meta_dataset'), np.array([self.input_shape, self.output_size, self.size], dtype=object), allow_pickle=True)
# -
# ### Transform training set into numpy arrays
# +
# define source and destination folders
source = join(SOURCE, TRAINING_SET_NAME)
destination = join(SOURCE, TRAINING_FEATURES)
# create the training_features directory if it does not exist
if not os.path.exists(destination):
os.makedirs(destination)
# transform images in training dataset (i.e. normalized pixel values and resized pictures) to numpy arrays (smaller files, useful if uploading the set to train a model in the cloud)
for f in os.listdir(source):
if f.find('.png') != -1:
img = Utils.get_preprocessed_img(join(source, f), IMAGE_SIZE)
file_name = f[:f.find('.png')]
np.savez_compressed(join(destination, file_name), features=img)
retrieve = np.load(join(destination, file_name + '.npz'))['features']
assert np.array_equal(img, retrieve)
shutil.copyfile(join(source, file_name + '.gui'), join(destination, file_name + '.gui'))
# -
# ### Transform evaluation set into numpy arrays
# +
# define source and destination folders
source = join(SOURCE, EVALUATION_SET_NAME)
destination = join(SOURCE, EVAL_FEATURES)
# create the eval_features directory if it does not exist
if not os.path.exists(destination):
os.makedirs(destination)
# transform images in eval dataset (i.e. normalized pixel values and resized pictures) to numpy arrays (smaller files, useful if uploading the set to train a model in the cloud)
for f in os.listdir(source):
if f.find('.png') != -1:
img = Utils.get_preprocessed_img(join(source, f), IMAGE_SIZE)
file_name = f[:f.find('.png')]
np.savez_compressed(join(destination, file_name), features=img)
retrieve = np.load(join(destination, file_name + '.npz'))['features']
assert np.array_equal(img, retrieve)
shutil.copyfile(join(source, file_name + '.gui'), join(destination, file_name + '.gui'))
# -
# make folder to store runtime files
if not os.path.exists('bin'):
os.mkdir('bin')
# ### Create folder to store checkpoint files
# make folder to store checkpoint files
if not os.path.exists(CHECKPOINT_DIR):
os.mkdir(CHECKPOINT_DIR)
# ### Declare class of callbacks - used to save model after each epoch
# +
from tensorflow.keras.callbacks import Callback
class TrainingCallback(Callback):
def on_train_begin(self, epoch, logs=None):
if os.path.isfile(CHECKPOINT_PATH):
print('Loading weights...')
model.load_weights(CHECKPOINT_PATH)
# -
# ### Declare magicode class
# +
from tensorflow.keras.layers import Input, Dense, Dropout, \
RepeatVector, LSTM, concatenate, \
Conv2D, MaxPooling2D, Flatten
from tensorflow.keras.models import Sequential, Model, model_from_json
from tensorflow.keras.optimizers import RMSprop
from tensorflow.keras import *
from tensorflow.keras.callbacks import EarlyStopping, TensorBoard
class magicode:
def __init__(self, input_shape, output_size, output_path):
self.model = None
self.name = 'magicode'
self.input_shape = input_shape
self.output_size = output_size
self.output_path = output_path
self.history = None
image_model = Sequential()
image_model.add(Conv2D(32, (3, 3), padding='valid', activation='relu', input_shape=input_shape))
image_model.add(Conv2D(32, (3, 3), padding='valid', activation='relu'))
image_model.add(MaxPooling2D(pool_size=(2, 2)))
image_model.add(Dropout(0.25))
image_model.add(Conv2D(64, (3, 3), padding='valid', activation='relu'))
image_model.add(Conv2D(64, (3, 3), padding='valid', activation='relu'))
image_model.add(MaxPooling2D(pool_size=(2, 2)))
image_model.add(Dropout(0.25))
image_model.add(Conv2D(128, (3, 3), padding='valid', activation='relu'))
image_model.add(Conv2D(128, (3, 3), padding='valid', activation='relu'))
image_model.add(MaxPooling2D(pool_size=(2, 2)))
image_model.add(Dropout(0.25))
image_model.add(Flatten())
image_model.add(Dense(1024, activation='relu'))
image_model.add(Dropout(0.3))
image_model.add(Dense(1024, activation='relu'))
image_model.add(Dropout(0.3))
image_model.add(RepeatVector(CONTEXT_LENGTH))
visual_input = Input(shape=input_shape)
encoded_image = image_model(visual_input)
language_model = Sequential()
language_model.add(LSTM(128, return_sequences=True, input_shape=(CONTEXT_LENGTH, output_size)))
language_model.add(LSTM(128, return_sequences=True))
textual_input = Input(shape=(CONTEXT_LENGTH, output_size))
encoded_text = language_model(textual_input)
decoder = concatenate([encoded_image, encoded_text])
decoder = LSTM(512, return_sequences=True)(decoder)
decoder = LSTM(512, return_sequences=False)(decoder)
decoder = Dense(output_size, activation='softmax')(decoder)
self.model = Model(inputs=[visual_input, textual_input], outputs=decoder)
optimizer = RMSprop(lr=LEARNING_RATE, clipvalue=CLIPVALUE)
self.model.compile(loss='categorical_crossentropy', optimizer=optimizer, metrics=['accuracy'])
def fit_generator(self, generator, eval_generator, steps_per_epoch, validation_steps):
# monitors the vali_loss value and when it stops decreasing the model will run for PATIENCE more times
# if the value doesn't decrease then restore the weights from the run with the smallest val_loss value
early_stopping = EarlyStopping(monitor='val_loss', mode = 'min', patience=PATIENCE, restore_best_weights=True, verbose = 1)
# Checkpoint callback to save the model's weights
checkpoint = tf.keras.callbacks.ModelCheckpoint(filepath=CHECKPOINT_PATH, save_weights_only=True, verbose=1)
self.history = self.model.fit(generator, steps_per_epoch=steps_per_epoch, epochs=EPOCHS, validation_data=eval_generator, validation_steps=validation_steps, verbose=1, callbacks=[early_stopping, checkpoint, TrainingCallback()])
self.save()
def predict(self, image, partial_caption):
return self.model.predict([image, partial_caption], verbose=0)[0]
def save(self):
model_json = self.model.to_json()
with open(join(self.output_path, self.name + '.json'), "w") as json_file:
json_file.write(model_json)
self.model.save_weights(join(self.output_path, self.name + '.h5'))
def load(self, name=""):
output_name = self.name if name == "" else name
with open(join(self.output_path, output_name + '.json'), "r") as json_file:
loaded_model_json = json_file.read()
self.model = model_from_json(loaded_model_json)
self.model.load_weights(join(self.output_path, output_name + '.h5'))
# -
# ### Train the model using a generator (to avoid having to fit all the data in memory)
# +
import tensorflow as tf
np.random.seed(1234)
training_features = join(SOURCE, TRAINING_FEATURES)
eval_features = join(SOURCE, EVAL_FEATURES)
output_path = join('bin')
dataset = Dataset()
dataset.dataset_name = 'training'
dataset.load(training_features, generate_binary_sequences=True)
dataset.save_metadata(output_path)
dataset.voc.save(output_path)
gui_paths, img_paths = Dataset.load_paths_only(training_features)
input_shape = dataset.input_shape
output_size = dataset.output_size
steps_per_epoch = dataset.size / BATCH_SIZE
voc = Vocabulary()
voc.retrieve(output_path)
generator = Generator.data_generator(voc, gui_paths, img_paths, batch_size=BATCH_SIZE, generate_binary_sequences=True)
eval_dataset = Dataset()
eval_dataset.dataset_name = 'evaluation'
eval_dataset.load(eval_features, generate_binary_sequences=True)
eval_gui_paths, eval_img_paths = Dataset.load_paths_only(eval_features)
validation_steps = eval_dataset.size / BATCH_SIZE
eval_generator = Generator.data_generator(voc, eval_gui_paths, eval_img_paths, batch_size=BATCH_SIZE, generate_binary_sequences=True)
model = magicode(input_shape, output_size, output_path)
# NOTE: upload any saved zip checkpoint to the training folder to start training from where it left off
model.fit_generator(generator, eval_generator, steps_per_epoch=steps_per_epoch, validation_steps=validation_steps)
# -
# ### Plot accuracy and loss from model history
# +
model.load(MODEL_NAME)
plt.figure(1)
# summarize history for accuracy
plt.subplot(211)
plt.plot(model.history.history['accuracy'])
plt.plot(model.history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
# summarize history for loss
plt.subplot(212)
plt.plot(model.history.history['loss'])
plt.plot(model.history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# -
# ### Create the directories for storing screenshots to "decode" and the resulting code
# create directory to store images to be "decoded"
if not os.path.exists('screenshots_to_convert'):
os.mkdir('screenshots_to_convert')
# create directory to store HTML and GUI code generated by "decoding" images in screenshots folder
if not os.path.exists('generated_code'):
os.mkdir('generated_code')
# ### Generate the code for provided screenshots - add files to the screenshots folder
# +
trained_weights_path = 'bin'
trained_model_name = 'magicode'
input_path = 'screenshots_to_convert'
output_path = 'generated_code'
meta_dataset = np.load(join(trained_weights_path, 'meta_dataset.npy'), allow_pickle=True)
input_shape = meta_dataset[0]
output_size = meta_dataset[1]
model = magicode(input_shape, output_size, trained_weights_path)
model.load(trained_model_name)
sampler = Sampler(trained_weights_path, input_shape, output_size, CONTEXT_LENGTH)
for f in os.listdir(input_path):
if f.find('.png') != -1:
evaluation_img = Utils.get_preprocessed_img(join(input_path,f), IMAGE_SIZE)
file_name = f[:f.find('.png')]
result, _ = sampler.predict_greedy(model, np.array([evaluation_img]))
print('Result greedy: {}'.format(result))
with open(join(output_path, file_name + '.gui'), 'w') as out_f:
out_f.write(result.replace(START_TOKEN, '').replace(END_TOKEN, ''))
# -
# ### Declare compiler
# +
import json
import string
import random
class Compiler:
def __init__(self, dsl_mapping_file_path):
with open(dsl_mapping_file_path) as data_file:
self.dsl_mapping = json.load(data_file)
self.opening_tag = self.dsl_mapping['opening-tag']
self.closing_tag = self.dsl_mapping['closing-tag']
self.content_holder = self.opening_tag + self.closing_tag
self.root = Node('body', None, self.content_holder)
def compile(self, input_file_path, output_file_path, rendering_function=None):
dsl_file = open(input_file_path)
current_parent = self.root
for token in dsl_file:
token = token.replace(' ', '').replace('\n', '')
if token.find(self.opening_tag) != -1:
token = token.replace(self.opening_tag, '')
element = Node(token, current_parent, self.content_holder)
current_parent.add_child(element)
current_parent = element
elif token.find(self.closing_tag) != -1:
current_parent = current_parent.parent
else:
tokens = token.split(',')
for t in tokens:
element = Node(t, current_parent, self.content_holder)
current_parent.add_child(element)
output_html = self.root.render(self.dsl_mapping, rendering_function=rendering_function)
with open(output_file_path, 'w') as output_file:
output_file.write(output_html)
class Node:
def __init__(self, key, parent_node, content_holder):
self.key = key
self.parent = parent_node
self.children = []
self.content_holder = content_holder
def add_child(self, child):
self.children.append(child)
def show(self):
print(self.key)
for child in self.children:
child.show()
def render(self, mapping, rendering_function=None):
content = ''
for child in self.children:
content += child.render(mapping, rendering_function)
value = mapping[self.key]
if rendering_function is not None:
value = rendering_function(self.key, value)
if len(self.children) != 0:
value = value.replace(self.content_holder, content)
return value
class CompilerUtils:
@staticmethod
def get_random_text(length_text=10, space_number=1, with_upper_case=True):
results = []
while len(results) < length_text:
char = random.choice(string.ascii_letters[:26])
results.append(char)
if with_upper_case:
results[0] = results[0].upper()
current_spaces = []
while len(current_spaces) < space_number:
space_pos = random.randint(2, length_text - 3)
if space_pos in current_spaces:
break
results[space_pos] = " "
if with_upper_case:
results[space_pos + 1] = results[space_pos - 1].upper()
current_spaces.append(space_pos)
return ''.join(results)
# -
# ### Compile the generated code
# +
FILL_WITH_RANDOM_TEXT = True
TEXT_PLACE_HOLDER = '[]'
dsl_path = join('compiler','assets','dsl-mapping.json')
compiler = Compiler(dsl_path)
def render_content_with_text(key, value):
text_inputs = ['input-text', 'input-password']
control_inputs = ['input-checkbox', 'input-radio']
if FILL_WITH_RANDOM_TEXT:
if key.find('btn') != -1:
value = value.replace(TEXT_PLACE_HOLDER, CompilerUtils.get_random_text())
elif key.find('title') != -1:
value = value.replace(TEXT_PLACE_HOLDER, CompilerUtils.get_random_text(length_text=5, space_number=0))
elif key.find('text') != -1:
value = value.replace(TEXT_PLACE_HOLDER,
CompilerUtils.get_random_text(length_text=56, space_number=7, with_upper_case=False))
elif any(text_input in key for text_input in text_inputs):
value = value.replace(TEXT_PLACE_HOLDER, CompilerUtils.get_random_text(length_text=30, space_number=0))
elif any(control_input in key for control_input in control_inputs):
value = value.replace(TEXT_PLACE_HOLDER, CompilerUtils.get_random_text(length_text=10, space_number=0))
return value
path = 'generated_code'
generated_code_files = os.listdir(path)
for file in generated_code_files:
file_uid = Path(file).stem
input_file_path = join(path, file_uid + '.gui')
output_file_path = join(path, file_uid + '.html')
compiler.compile(input_file_path, output_file_path, rendering_function=render_content_with_text)
|
Notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # NumPy: Numerical Arrays for Python
# **Learning Objectives:** Learn how to create, transform and visualize multidimensional data of a single type using Numpy.
#
# NumPy is the foundation for scientific computing and data science in Python. Its more data object is a multidimensional array with the following characteristics:
#
# * Any number of dimensions
# * All elements of an array have the same data type
# * Array elements are usually native data dtype
# * The memory for an array is a contiguous block that can be easily passed to other numerical libraries (BLAS, LAPACK, etc.).
# * Most of NumPy is implemented in C, so it is fast.
# ## Plotting
# While this notebook doesn't focus on plotting, Matplotlib will be used to make a few basic plots.
# %matplotlib inline
from matplotlib import pyplot as plt
import seaborn as sns
# The `vizarray` package will be used to visualize NumPy arrays:
import antipackage
from github.ellisonbg.misc import vizarray as va
# ## Multidimensional array type
# This is the canonical way you should import Numpy:
import numpy as np
data = [0,2,4,6]
a = np.array(data)
type(a)
a
# The `vz.vizarray` function can be used to visualize a 1d or 2d NumPy array using a colormap:
va.vizarray(a)
# The shape of the array:
a.shape
# The number of array dimensions:
a.ndim
# The number of array elements:
a.size
# The number of bytes the array takes up:
a.nbytes
# The `dtype` attribute describes the "data type" of the elements:
a.dtype
# ## Creating arrays
# Arrays can be created with nested lists or tuples:
data = [[0.0,2.0,4.0,6.0],[1.0,3.0,5.0,7.0]]
b = np.array(data)
b
va.vizarray(b)
b.shape, b.ndim, b.size, b.nbytes
# The `arange` function is similar to Python's builtin `range` function, but creates an array:
c = np.arange(0.0, 10.0, 1.0) # Step size of 1.0
c
# The `linspace` function is similar, but allows you to specify the number of points:
e = np.linspace(0.0, 5.0, 11) # 11 points
e
# There are also `empty`, `zeros` and `ones` functions:
np.empty((4,4))
np.zeros((3,3))
np.ones((3,3))
# See also:
#
# * `empty_like`, `ones_like`, `zeros_like`
# * `eye`, `identity`, `diag`
# ## dtype
# Arrays have a `dtype` attribute that encapsulates the "data type" of each element. It can be set:
#
# * Implicitely by the element type
# * By passing the `dtype` argument to an array creation function
#
# Here is an integer valued array:
a = np.array([0,1,2,3])
a, a.dtype
# All array creation functions accept an optional `dtype` argument:
b = np.zeros((2,2), dtype=np.complex64)
b
c = np.arange(0, 10, 2, dtype=np.float)
c
# You can use the `astype` method to create a copy of the array with a given `dtype`:
d = c.astype(dtype=np.int)
d
# IPython's tab completion is useful for exploring the various available `dtypes`:
# +
# np.float*?
# -
# The NumPy documentation on [dtypes](http://docs.scipy.org/doc/numpy/reference/arrays.dtypes.html) describes the many other ways of specifying dtypes.
# ## Array operations
# Basic mathematical operations are **elementwise** for:
#
# * Scalars and arrays
# * Arrays and arrays
#
# Fill an array with a value:
a = np.empty((3,3))
a.fill(0.1)
a
b = np.ones((3,3))
b
# Addition is elementwise:
a+b
# Division is elementwise:
b/a
# As are powers:
a**2
# Scalar multiplication is also elementwise:
np.pi*b
# ## Indexing and slicing
# Indexing and slicing provide an efficient way of getting the values in an array and modifying them.
a = np.random.rand(10,10)
# The `enable` function is part of `vizarray` and enables a nice display of arrays:
va.enable()
a
# List Python lists and tuples, NumPy arrays have zero-based indexing and use the `[]` syntax for getting and setting values:
a[0,0]
# An index of `-1` refers to the last element along that axis:
a[-1,-1] == a[9,9]
# Extract the 0th column using the `:` syntax, which denotes all elements along that axis.
a[:,0]
# The last row:
a[-1,:]
# You can also slice ranges:
a[0:2,0:2]
# Assignment also works with slices:
a[0:5,0:5] = 1.0
a
# Note how even though we assigned the value to the slice, the original array was changed. This clarifies that slices are **views** of the same data, not a copy.
va.disable()
# ### Boolean indexing
# Arrays can be indexed using other arrays that have boolean values.
ages = np.array([23,56,67,89,23,56,27,12,8,72])
genders = np.array(['m','m','f','f','m','f','m','m','m','f'])
# Boolean expressions involving arrays create new arrays with a `bool` dtype and the elementwise result of the expression:
ages > 30
genders == 'm'
# Boolean expressions provide an extremely fast and flexible way of querying arrays:
(ages > 10) & (ages < 50)
# You can use a boolean array to index into the original or another array. This selects the ages of all females in the `genders` array:
mask = (genders == 'f')
ages[mask]
ages[ages>30]
# ## Reshaping, transposing
va.enable()
a = np.random.rand(3,4)
a
# The `T` atrribute contains the transpose of the original array:
a.T
# The `reshape` method can be used to change the shape and even the number of dimensions:
a.reshape(2,6)
a.reshape(6,2)
# The `ravel` method strings the array out in one dimension:
a.ravel()
va.disable()
# ## Universal functions
# Universal function, or "ufuncs," are functions that take and return arrays or scalars. They have the following characteristics:
#
# * Vectorized C implementations, much faster than hand written loops in Python
# * Allow for concise Pythonic code
# * Here is a complete list of the [available NumPy ufuncs](http://docs.scipy.org/doc/numpy/reference/ufuncs.html#available-ufuncs) lists the available ufuncs.
va.set_block_size(5)
va.enable()
# Here is a linear sequence of values"
t = np.linspace(0.0, 4*np.pi, 100)
t
# Take the $sin$ of each element of the array:
np.sin(t)
# As the next two examples show, multiple ufuncs can be used to create complex mathematical expressions that can be computed efficiently:
np.exp(np.sqrt(t))
va.disable()
va.set_block_size(30)
plt.plot(t, np.exp(-0.1*t)*np.sin(t))
# In general, you should always try to use ufuncs rather than do computations using for loops. These types of array based computations are referred to as *vectorized*.
# ## Basic data processing
ages = np.array([23,56,67,89,23,56,27,12,8,72])
genders = np.array(['m','m','f','f','m','f','m','m','m','f'])
# Numpy has a basic set of methods and function for computing basic quantities about data.
ages.min(), ages.max()
# Compute the mean:
ages.mean()
# Compute the variance and standard deviation:
ages.var(), ages.std()
# The `bincount` function counts how many times each value occurs in the array:
np.bincount(ages)
# The `cumsum` and `cumprod` methods compute cumulative sums and products:
ages.cumsum()
ages.cumprod()
# Most of the functions and methods above take an `axis` argument that will apply the action along a particular axis:
a = np.random.randint(0,10,(3,4))
a
# With `axis=0` the action takes place along rows:
a.sum(axis=0)
# With `axis=1` the action takes place along columns:
a.sum(axis=1)
# The `unique` function is extremely useful in working with categorical data:
np.unique(genders)
np.unique(genders, return_counts=True)
# The where function allows you to apply conditional logic to arrays. Here is a rough sketch of how it works:
#
# ```python
# def where(condition, if_false, if_true):
# ```
np.where(ages>30, 0, 1)
# The `if_false` and `if_true` values can be arrays themselves:
np.where(ages<30, 0, ages)
# ## File IO
# NumPy has a a number of different function to reading and writing arrays to and from disk.
# ### Single array, binary format
a = np.random.rand(10)
a
# Save the array to a binary file named `array1.npy`:
np.save('array1', a)
# ls
# Using `%pycat` to look at the file shows that it is binary:
# %pycat array1.npy
# Load the array back into memory:
a_copy = np.load('array1.npy')
a_copy
# ### Single array, text format
b = np.random.randint(0,10,(5,3))
b
# The `savetxt` function saves arrays in a simple, textual format that is less effecient, but easier for other languges to read:
np.savetxt('array2.txt', b)
# ls
# Using `%pycat` to look at the contents shows that the files is indeed a plain text file:
# %pycat array2.txt
np.loadtxt('array2.txt')
# ### Multiple arrays, binary format
# The `savez` function provides an efficient way of saving multiple arrays to a single file:
np.savez('arrays.npz', a=a, b=b)
# The `load` function returns a dictionary like object that provides access to the individual arrays:
a_and_b = np.load('arrays.npz')
a_and_b['a']
a_and_b['b']
# ## Linear algebra
# NumPy has excellent linear algebra capabilities.
a = np.random.rand(5,5)
b = np.random.rand(5,5)
# Remember that array operations are elementwise. Thus, this is **not** matrix multiplication:
a*b
# To get matrix multiplication use `np.dot`:
np.dot(a, b)
# Or, NumPy as a `matrix` subclass for which matrix operations are the default:
m1 = np.matrix(a)
m2 = np.matrix(b)
m1*m2
# The `np.linalg` package has a wide range of fast linear algebra operations.
#
# Here is determinant:
np.linalg.det(a)
# Matrix inverse:
np.linalg.inv(a)
# Eigenvalues:
np.linalg.eigvals(a)
# NumPy can be built against fast BLAS/LAPACK implementation for these linear algebra operations.
c = np.random.rand(2000,2000)
# %timeit -n1 -r1 evs = np.linalg.eigvals(c)
# ## Random numbers
# NumPy has functions for creating arrays of random numbers from different distributions in `np.random`, as well as handling things like permutation, shuffling, and choosing.
#
# Here is the [numpy.random documentation](http://docs.scipy.org/doc/numpy/reference/routines.random.html).
plt.hist(np.random.random(250))
plt.title('Uniform Random Distribution $[0,1]$')
plt.xlabel('value')
plt.ylabel('count')
plt.hist(np.random.randn(250))
plt.title('Standard Normal Distribution')
plt.xlabel('value')
plt.ylabel('count')
# The `shuffle` function shuffles an array in place:
a = np.arange(0,10)
np.random.shuffle(a)
a
# The `permutation` function does the same thing but first makes a copy:
a = np.arange(0,10)
print(np.random.permutation(a))
print(a)
# The `choice` function provides a powerful way of creating synthetic data sets of discrete data:
np.random.choice(['m','f'], 20, p=[0.25,0.75])
# ## Resources
# * [NumPy Reference Documentation](http://docs.scipy.org/doc/numpy/reference/)
# * [Python Scientific Lecture Notes](http://scipy-lectures.github.io/index.html), Edited by <NAME>,
# <NAME> and <NAME>.
# * [Lectures on Scientific Computing with Python](https://github.com/jrjohansson/scientific-python-lectures), <NAME>.
# * [Introduction to Scientific Computing in Python](http://nbviewer.ipython.org/github/jakevdp/2014_fall_ASTR599/tree/master/), Jake Vanderplas.
|
days/day05/Numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1. Machine Learning
#
# ## Qué es Machine Learning?
#
# El término Machine Learning según la definición común es "el estudio de los algoritmos de computador que mejoran automáticamente a través de la experiencia".
#
#
# ## Aplicaciones recientes:
# ### Computación visual.
# - ImageNet
# - Vehículos conducidos autonomamente.
# - Redes Neuronales Convolucionales
# - Robots retroalimentados con CV.
#
# ### Reconocimiento de voz
# - Asistentes por voz.
# - Traductores automáticos en tiempo real.
#
# ### Traducción de idioma.
# - Traducción de idioma no supervisada.
#
# ### Videojuegos.
# - Aprendizaje por refuerzo para solucionar juegos.
# - Aprendizaje para mejorar experiencia del jugador.
#
# ### Ranking:
# - Sistemas de recomendación.
# ...
#
# ## Tipos de Aprendizaje
#
# Hay tres grandes categorías en las que se puede enmarcar el Machine Learning. Es importante poder identificar el tipo de algoritmo de Machine Learning que se va a utilizar dependiendo del caso de estudio ya que esto permite planear el ambiente más apropiado y entender qué hacer para que funcione.
#
# ## Aprendizaje Supervisado.
#
# El aprendizaje supervisado es el más común y popular de los paradigmas de Machine Learning. El aprendizaje supervisado logra, a partir de datos conocidos y respuestas conocidas a estos datos, aproximar una respuesta, de manera que ante la presencia de datos nuevos estima una nueva respuesta. (ie. Aprender $y=f(x)$ a partir de ${x_i,y_i}^{n}_{i=1}$)
#
# Aplicaciones:
# - Clasificación de Spam.
# - Reconocimiento facial.
#
# ## Aprendizaje No Supervisado.
#
# El aprendizaje no supervisado es opuesto al aprendizaje supervisado en varios sentidos. El objetivo del aprendizaje no supervisado es que, a partir de grandes cantidades de información no etiquetada, sea posible entender las propiedades de la información. A partir de aquí, es posible aprender a agrupar y organizar la información de manera que un humano (u otro algoritmo inteligente) pueda encontrar un sentido a partir de la información organizada. El aprendizaje no supervisado es interesante debido a que mucha información en el mundo real no está etiquetada. (ie. Aprender similaridades en ${x_i}^{n}_{i=1}$)
#
# Aplicaciones:
#
# - Sistemas generadores de recomendaciones.
#
# - Estimar hábitos de compra.
#
# Aprendizaje por Refuerzo.
#
# El aprendizaje por refuerzo es diferente a los aprendizajes supervisado y no supervisado. El objetivo del aprendizaje por refuerzo es determinar las políticas (o acciones) que debe tener un agente en orden de maximizar una función de recompensa. En Machine Learning, es muy común que el ambiente del agente sea modelado como un proceso de decisión de Markov. En general, se busca encontrar un equilibrio entre la exploración y la explotación.
#
# ## Aplicaciones:
#
# - Simulaciones industriales.
# - Manejo de recursos en ambientes complejos.
# - Videojuegos.
#
#
# ## Aspectos importantes de Machine Learning.
#
# - Validación y selección de Modelo.
# - Aprendizaje Estadístico.
# - Machine Learning Computacional.
# - Optimización.
#
#
#
#
# # 2. Aprendizaje Supervisado: Regresión Lineal
#
# A partir de un conjunto de variables de entrada, se busca predecir una característica objetivo de salida. Se denota una pareja $(x^{(i)},y^{(i)})$, la muestra de entrenamiento y el conjunto desde $i=1,...,n$ es denotado, conjunto de entrenamiento. El objetivo es a partir de nuestro conjunto deentrada $\mathcal{X}$ obtener una hipótesis $h:\mathcal{X} \rightarrow \mathcal{Y}$. Cuando $y$ únicamente puede tomar un pequeño número de valores discretos, se dice que este es un problema de clasificación.
#
#
# ## 2.1 Regresión Lineal
#
# Para realizar aprendizaje supervisado, se necesita decidir cómo representar funciones hipótesis $h$. Inicialmente, es posible decidir representarlas como funciones lineales de x:
#
# Denotaremos
#
# \begin{equation}
# h_{\theta}(x)=\theta_0 + \theta_1 x_1 +\theta_2 x_2
# \end{equation}
#
# Aquí, las $\theta_i$ son los parámetros, también llamados pesos (weights). También es posible escribir lo anterior de la siguiente manera.
#
# \begin{equation}
# h_{\theta}(x)=\sum_{i=0}^{d}\theta_i x_i=\theta^T x
# \end{equation}
#
# Ahora, es importante en un conjunto de entrenamiento, escoger una función de costo que permita hacer que $h(x)$ sea tan cercana a $y$ como sea posible para obtener unos parámetros $\theta$ apropiados. Definimos una función de costo:
#
# \begin{equation}
# J(\theta)=\frac{1}{2}\sum_{i=1}^{n}(h_{\theta}(x^{(i)})-y^{(i)})^2
# \end{equation}
#
# Y podemos recordar esta ecuación como la solución al modelo de mínimos cuadrados.
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
def linReg(X, y, theta=0, alpha=0.01, num_iters=100):
m = y.size
costs=[]
for i in range(num_iters):
y_hat = np.dot(X, theta)
costi =1.0/2.0 * sum((y_hat-y)**2)
costs.append(costi)
theta = theta - alpha * (1.0/m) * np.dot(X.T, y_hat-y)
return theta, np.array(costs)
#Generamos un conjunto de datos sintético de prueba.
x=np.linspace(-1,5,10)
y=7*x
y=y+(np.random.rand(10)*2-1)*7
#Realizamos el ajuste
t,c=linReg(x,y)
#Graficamos el ajuste
plt.plot(x,t*x,color='k',label='fit')
plt.scatter(x,y,color='blue',label='datos')
plt.legend()
plt.title('Regresión lineal, descenso de gradiente')
plt.show()
#Graficamos la función de costo en función de las iteraciones
plt.plot(c)
plt.xlabel('Iteración')
plt.ylabel(r'$J(\theta)$')
plt.title('Función de costo')
plt.show()
# -
# # 3. Mínimos cuadrados
# Es claro que a partir de la función de costo $J(\theta)$ que se definió previamente, el parámetro óptimo que se puede obtener utilizando mínimos cuadrados es $\hat{\theta}=(X^T X)^{-1}X^T \vec{y}$. Sin embargo, no siempre es factible llegar a esta solución debido a distintas ra
#
# ## 3.1 Interpretación probabilística
#
# Cuando se tiene un problema de regresión es común pensar en por qué la escogencia de la función de costo $J$ es apropiada. Asumamos que el target y las entradas están relacionadas a través de
#
# \begin{equation}
# y=\theta^T x+ \epsilon
# \end{equation}
#
# donde $\epsilon$ es un término de error que contiene toda la información que no se ha modelado. Se puede asumir que este término es IID y se modela gaussiano estándar. Por lo tanto, se puede decir que
#
# \begin{equation}
# p(y|x;\theta)=\frac{1}{\sqrt{2\pi \sigma^2}} \exp{\frac{-(y-\theta^T x)^2}{2\sigma^2}}
# \end{equation}
#
# A partir de esta probabilidad se desea observar esta función explicitamente como una función de $\theta$, a esta función se la llamará función de likelihood.
#
# \begin{equation}
# L(\theta)=L(\theta;X,y)=p(y|X;\theta)
# \end{equation}
#
# Donde
# \begin{equation}
# L(\theta)=\prod{\frac{1}{\sqrt{2\pi \sigma^2}} \exp{\frac{-(y-\theta^T x)^2}{2\sigma^2}}}
# \end{equation}
#
# Ahora, dado este modelo probabilístico que relaciona $y$ con $x$, se desea cuál es la manera razonable de escoger el parámetro $\theta$. Como se ha visto previamente, en lugar de maximizar esta función, es posible maximizar la función log-llikelihood $l(\theta)$
#
# Donde
# \begin{equation}
# l(\theta)=\log{\prod{\frac{1}{\sqrt{2\pi \sigma^2}} \exp{\frac{-(y-\theta^T x)^2}{2\sigma^2}}}}
# \end{equation}
#
# De manera que el máximo es
#
# Donde
# \begin{equation}
# \frac{1}{2}\sum_{i=1}^{n}{(y^{(i)}-\theta^T x^{(i)})^2}
# \end{equation}
#
# Por lo que la escogenca de nuestra función de costo es justificada.
#
#
#
#
# +
import matplotlib.pyplot as plt
import numpy as np
from sklearn import datasets, linear_model
from sklearn.metrics import mean_squared_error, r2_score
diabetes = datasets.load_diabetes()
diabetes_X = diabetes.data[:, np.newaxis, 2]
diabetes_X_train = diabetes_X[:-20]
diabetes_X_test = diabetes_X[-20:]
diabetes_y_train = diabetes.target[:-20]
diabetes_y_test = diabetes.target[-20:]
regr = linear_model.LinearRegression()
regr.fit(diabetes_X_train, diabetes_y_train)
diabetes_y_pred = regr.predict(diabetes_X_test)
print('Coefficients: \n', regr.coef_)
print("Mean squared error: %.2f"
% mean_squared_error(diabetes_y_test, diabetes_y_pred))
print('Variance score: %.2f' % r2_score(diabetes_y_test, diabetes_y_pred))
plt.scatter(diabetes_X_test, diabetes_y_test, color='blue')
plt.plot(diabetes_X_test, diabetes_y_pred, color='black', linewidth=3)
plt.title('Regresión lineal')
plt.xlabel('x')
plt.ylabel('y')
plt.show()
# -
|
9ML/.ipynb_checkpoints/1_IntroML-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Combining Datasets: Merge and Join
# One essential feature offered by Pandas is its high-performance, in-memory join and merge operations.
# If you have ever worked with databases, you should be familiar with this type of data interaction.
# The main interface for this is the ``pd.merge`` function, and we'll see few examples of how this can work in practice.
import pandas as pd
import numpy as np
# ## Relational Algebra
#
# The behavior implemented in ``pd.merge()`` is a subset of what is known as *relational algebra*, which is a formal set of rules for manipulating relational data, and forms the conceptual foundation of operations available in most databases.
# The strength of the relational algebra approach is that it proposes several primitive operations, which become the building blocks of more complicated operations on any dataset.
# With this lexicon of fundamental operations implemented efficiently in a database or other program, a wide range of fairly complicated composite operations can be performed.
#
# Pandas implements several of these fundamental building-blocks in the ``pd.merge()`` function and the related ``join()`` method of ``Series`` and ``Dataframe``s.
# As we will see, these let you efficiently link data from different sources.
# ## Categories of Joins
#
# The ``pd.merge()`` function implements a number of types of joins: the *one-to-one*, *many-to-one*, and *many-to-many* joins.
# All three types of joins are accessed via an identical call to the ``pd.merge()`` interface; the type of join performed depends on the form of the input data.
# Here we will show simple examples of the three types of merges, and discuss detailed options further below.
# ### One-to-one joins
#
# Perhaps the simplest type of merge expresion is the one-to-one join, which is in many ways very similar to the column-wise concatenation seen in NumPy.
# As a concrete example, consider the following two ``DataFrames`` which contain information on several employees in a company:
df1 = pd.DataFrame({'employee': ['Bob', 'Jake', 'Lisa', 'Sue'],
'group': ['Accounting', 'Engineering', 'Engineering', 'HR']})
df2 = pd.DataFrame({'employee': ['Lisa', 'Bob', 'Jake', 'Sue'],
'hire_date': [2004, 2008, 2012, 2014]})
df1
df2
# To combine this information into a single ``DataFrame``, we can use the ``pd.merge()`` function:
df3 = pd.merge(df1, df2)
df3
# The ``pd.merge()`` function recognizes that each ``DataFrame`` has an "employee" column, and automatically joins using this column as a key.
# The result of the merge is a new ``DataFrame`` that combines the information from the two inputs.
# Notice that the order of entries in each column is not necessarily maintained: in this case, the order of the "employee" column differs between ``df1`` and ``df2``, and the ``pd.merge()`` function correctly accounts for this.
# Additionally, keep in mind that the merge in general discards the index, except in the special case of merges by index (see the ``left_index`` and ``right_index`` keywords, discussed momentarily).
# ### Many-to-one joins
# Many-to-one joins are joins in which one of the two key columns contains duplicate entries.
# For the many-to-one case, the resulting ``DataFrame`` will preserve those duplicate entries as appropriate.
# Consider the following example of a many-to-one join:
df4 = pd.DataFrame({'group': ['Accounting', 'Engineering', 'HR'],
'supervisor': ['Carly', 'Guido', 'Steve']})
df3
df4
pd.merge(df3, df4)
# The resulting ``DataFrame`` has an aditional column with the "supervisor" information, where the information is repeated in one or more locations as required by the inputs.
# ### Many-to-many joins
# Many-to-many joins are a bit confusing conceptually, but are nevertheless well defined.
# If the key column in both the left and right array contains duplicates, then the result is a many-to-many merge.
# This will be perhaps most clear with a concrete example.
# Consider the following, where we have a ``DataFrame`` showing one or more skills associated with a particular group.
# By performing a many-to-many join, we can recover the skills associated with any individual person:
df5 = pd.DataFrame({'group': ['Accounting', 'Accounting',
'Engineering', 'Engineering', 'HR', 'HR'],
'skills': ['math', 'spreadsheets', 'coding', 'linux',
'spreadsheets', 'organization']})
df5
pd.merge(df1,df5)
# These three types of joins can be used with other Pandas tools to implement a wide array of functionality.
# But in practice, datasets are rarely as clean as the one we're working with here.
# In the following section we'll consider some of the options provided by ``pd.merge()`` that enable you to tune how the join operations work.
# ## Specification of the Merge Key
# We've already seen the default behavior of ``pd.merge()``: it looks for one or more matching column names between the two inputs, and uses this as the key.
# However, often the column names will not match so nicely, and ``pd.merge()`` provides a variety of options for handling this.
# ### The ``on`` keyword
#
# Most simply, you can explicitly specify the name of the key column using the ``on`` keyword, which takes a column name or a list of column names:
df1
df2
pd.merge(df1, df2, on='employee')
# This option works only if both the left and right ``DataFrame``s have the specified column name.
# ### The ``left_on`` and ``right_on`` keywords
#
# At times you may wish to merge two datasets with different column names; for example, we may have a dataset in which the employee name is labeled as "name" rather than "employee".
# In this case, we can use the ``left_on`` and ``right_on`` keywords to specify the two column names:
df3 = pd.DataFrame({'name': ['Bob', 'Jake', 'Lisa', 'Sue'],
'salary': [70000, 80000, 120000, 90000]})
df3
df1
pd.merge(df1, df3, left_on="employee", right_on="name")
# The result has a redundant column that we can drop if desired–for example, by using the ``drop()`` method of ``DataFrame``s:
pd.merge(df1, df3, left_on="employee", right_on="name").drop('name', axis=1)
# ### The ``left_index`` and ``right_index`` keywords
#
# Sometimes, rather than merging on a column, you would instead like to merge on an index.
# For example, your data might look like this:
df1a = df1.set_index('employee')
df2a = df2.set_index('employee')
df1a
df2a
# You can use the index as the key for merging by specifying the ``left_index`` and/or ``right_index`` flags in ``pd.merge()``:
pd.merge(df1a, df2a, left_index=True, right_index=True)
# For convenience, ``DataFrame``s implement the ``join()`` method, which performs a merge that defaults to joining on indices:
df1a.join(df2a)
# If you'd like to mix indices and columns, you can combine ``left_index`` with ``right_on`` or ``left_on`` with ``right_index`` to get the desired behavior:
pd.merge(df1a, df3, left_index=True, right_on='name')
# All of these options also work with multiple indices and/or multiple columns; the interface for this behavior is very intuitive.
# For more information on this, see the ["Merge, Join, and Concatenate" section](http://pandas.pydata.org/pandas-docs/stable/merging.html) of the Pandas documentation.
# ## Specifying Set Arithmetic for Joins
# In all the preceding examples we have glossed over one important consideration in performing a join: the type of set arithmetic used in the join.
# This comes up when a value appears in one key column but not the other. Consider this example:
df6 = pd.DataFrame({'name': ['Peter', 'Paul', 'Mary'],
'food': ['fish', 'beans', 'bread']},
columns=['name', 'food'])
df7 = pd.DataFrame({'name': ['Mary', 'Joseph'],
'drink': ['wine', 'beer']},
columns=['name', 'drink'])
df6
df7
pd.merge(df6,df7)
# Here we have merged two datasets that have only a single "name" entry in common: Mary.
# By default, the result contains the *intersection* of the two sets of inputs; this is what is known as an *inner join*.
# We can specify this explicitly using the ``how`` keyword, which defaults to ``"inner"``:
pd.merge(df6, df7, how='inner')
# Other options for the ``how`` keyword are ``'outer'``, ``'left'``, and ``'right'``.
# An *outer join* returns a join over the union of the input columns, and fills in all missing values with NAs:
pd.merge(df6, df7, how='outer')
# The *left join* and *right join* return joins over the left entries and right entries, respectively.
# For example:
pd.merge(df6, df7, how='left')
pd.merge(df6, df7, how='right')
# The output rows now correspond to the entries in the left input. Using
# ``how='right'`` works in a similar manner.
#
# All of these options can be applied straightforwardly to any of the preceding join types.
# ## Overlapping Column Names: The ``suffixes`` Keyword
# Finally, you may end up in a case where your two input ``DataFrame``s have conflicting column names.
# Consider this example:
df8 = pd.DataFrame({'name': ['Bob', 'Jake', 'Lisa', 'Sue'],
'rank': [1, 2, 3, 4]})
df9 = pd.DataFrame({'name': ['Bob', 'Jake', 'Lisa', 'Sue'],
'rank': [3, 1, 4, 2]})
df8
df9
pd.merge(df8, df9, on="name")
# Because the output would have two conflicting column names, the merge function automatically appends a suffix ``_x`` or ``_y`` to make the output columns unique.
# If these defaults are inappropriate, it is possible to specify a custom suffix using the ``suffixes`` keyword:
pd.merge(df8, df9, on="name", suffixes=["_L", "_R"])
# These suffixes work in any of the possible join patterns, and work also if there are multiple overlapping columns.
# ## Example: Indonesia Data
#
# Merge and join operations come up most often when combining data from different sources.
# Here we will consider an example of some data about Indonesia and their populations.
# Read `luas_wilayah.csv` and `jumlah_penduduk.csv`.
# Given this information, say we want to compute a relatively straightforward result: rank Indonesian provincies by their 2010 population density.
# We clearly have the data here to find this result, but we'll have to combine the datasets to find the result. Start with a many-to-one merge that will give us the full state name within the population ``DataFrame``. Use ``how='outer'`` to make sure no data is thrown away due to mismatched labels.
# Double-check whether there were any mismatches using `.isnull().any()`
luas = pd.read_csv('luas_wilayah.csv', delimiter=',')
jumlah = pd.read_csv('jumlah_penduduk.csv', delimiter=';', na_values=['-','-'])
|
day3/07. pandas - Merge-and-Join.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import tensorflow_probability as tfp
tfd = tfp.distributions
import numpy as np
import matplotlib.pyplot as plt
from tqdm import tqdm, trange
from rec.coding.samplers import ImportanceSampler
tf.config.experimental.set_visible_devices([], 'GPU')
# -
# # Importance Sampling Encoder and Decoder
# +
target = tfd.Normal(loc=[-1.], scale=[0.3])
proposal = tfd.Normal(loc=[1.], scale=[1.1])
kl = tf.reduce_sum(tfd.kl_divergence(target, proposal))
print(f"KL between target and proposal: {kl:.4f}")
# -
s = ImportanceSampler(alpha=1.)
# +
samps = []
indices = []
for i in trange(1000):
ind, samp = s.coded_sample(target=target,
coder=proposal,
seed=i)
samps.append(samp)
indices.append(ind)
samps = tf.concat(samps, axis=0)
# +
xs = tf.linspace(-4., 10., 300)
plt.plot(xs, target.prob(xs))
plt.plot(xs, proposal.prob(xs))
plt.hist(samps, bins=100, density=True)
plt.show()
tf.reduce_mean(tf.math.log(tf.cast(tf.stack(indices), tf.float32) + 1))
# +
dec_samps = []
for i in trange(1000):
dec_samp = s.decode_sample(coder=proposal,
sample_index=indices[i],
seed=i)
dec_samps.append(dec_samp)
dec_samps = tf.concat(dec_samps, axis=0)
# -
tf.reduce_sum(tf.abs(dec_samps - samps))
|
notebooks/coding/REC Coding.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="9_4mVKJel0VU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="85120fc6-3516-446e-a0d7-d3af0bd14221" executionInfo={"status": "ok", "timestamp": 1562488863882, "user_tz": -540, "elapsed": 2870, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}}
# !nvidia-smi
# + id="_k8u3RdK5Nn-" colab_type="code" colab={}
import torch
import torchvision
import torchvision.transforms as transforms
# + id="4uzh4w8zURRX" colab_type="code" colab={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from PIL import Image
# + id="cpDHCVJsXq7s" colab_type="code" colab={}
BATCH_SIZE = 96
# + id="ZLYvfeWDW_Vj" colab_type="code" colab={}
class RandomErasing(object):
def __init__(self, p=0.5, sl=0.02, sh=0.4, r1=0.3, r2=3):
self.p = p
self.sl = sl
self.sh = sh
self.r1 = r1
self.r2 = r2
def __call__(self, img):
if np.random.rand() > self.p:
return img
img = np.array(img)
while True:
img_h, img_w, img_c = img.shape
img_area = img_h * img_w
mask_area = np.random.uniform(self.sl, self.sh) * img_area
mask_aspect_ratio = np.random.uniform(self.r1, self.r2)
mask_w = int(np.sqrt(mask_area / mask_aspect_ratio))
mask_h = int(np.sqrt(mask_area * mask_aspect_ratio))
mask = np.random.rand(mask_h, mask_w, img_c) * 255
left = np.random.randint(0, img_w)
top = np.random.randint(0, img_h)
right = left + mask_w
bottom = top + mask_h
if right <= img_w and bottom <= img_h:
break
img[top:bottom, left:right, :] = mask
return Image.fromarray(img)
# + id="QANOPvH969Xs" colab_type="code" outputId="3ac0a23a-d5d5-4deb-e1d7-a202a61c2c7f" executionInfo={"status": "ok", "timestamp": 1562488871659, "user_tz": -540, "elapsed": 3230, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 102}
train_transform = transforms.Compose(
[transforms.RandomHorizontalFlip(),
RandomErasing(p=0.5, sh=0.3),
transforms.ToTensor()]
)
test_transform = transforms.Compose(
[transforms.RandomHorizontalFlip(),
transforms.ToTensor()]
)
trainval_dataset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=train_transform)
test_dataset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=test_transform)
train_size = int(len(trainval_dataset) * 0.8)
val_size = len(trainval_dataset)- train_size
train_dataset, val_dataset = torch.utils.data.random_split(trainval_dataset, [train_size, val_size])
print("train data num:", len(train_dataset))
print("val data num:", len(val_dataset))
print("test data num:", len(test_dataset))
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=BATCH_SIZE, shuffle=True, num_workers=2)
val_loader = torch.utils.data.DataLoader(val_dataset, batch_size=BATCH_SIZE, shuffle=False, num_workers=2)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=BATCH_SIZE, shuffle=False, num_workers=2)
classes = ('plane', 'car', 'bird', 'cat',
'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
# + id="Nk5qtY6aYEkq" colab_type="code" colab={}
def count_dataset(dataset, n_classes, normalize=False):
counter = [0] * n_classes
for _, label in dataset:
counter[label] += 1
if normalize:
sum_cnt = sum(counter)
normalized_counter = [c / sum_cnt for c in counter]
return normalized_counter
else:
return counter
def show_bar(val, labels, title=''):
left = [i for i in range(len(labels))]
plt.title(title)
plt.bar(left, val, tick_label=labels, align="center")
plt.show()
# + id="zQ_uV1VKYlhq" colab_type="code" colab={}
train_cnt = count_dataset(train_dataset, len(classes), normalize=False)
val_cnt = count_dataset(val_dataset, len(classes), normalize=False)
test_cnt = count_dataset(test_dataset, len(classes), normalize=False)
# + id="lricFt8pNeKA" colab_type="code" outputId="45cab389-c8d8-416d-af7c-9feeb0f11d49" executionInfo={"status": "ok", "timestamp": 1562488885890, "user_tz": -540, "elapsed": 15787, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 809}
show_bar(train_cnt, classes, title='train dataset')
show_bar(val_cnt, classes, title='val dataset')
show_bar(test_cnt, classes, title='test dataset')
# + id="Lt_mVMqO7DmU" colab_type="code" outputId="d4cbb175-04da-4025-a92f-0b4d075c6f6b" executionInfo={"status": "ok", "timestamp": 1562488886936, "user_tz": -540, "elapsed": 15538, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 787}
def show_img(img, figsize=(5,5)):
plt.figure(figsize=figsize)
if hasattr(img, 'numpy'):
img = img.numpy()
img = img.transpose((1, 2, 0)) # C H W -> H W C
img = np.squeeze(img)
plt.imshow(img)
plt.axis('off')
plt.show()
dataiter = iter(train_loader)
imgs, labels = dataiter.next()
imgs = imgs[:25]
labels = labels[:25]
grid = torchvision.utils.make_grid(imgs, nrow=5, padding=1)
show_img(grid, figsize=(12, 12))
class_list = [classes[labels[j]] for j in range(25)]
class_list = np.array(class_list)
print(class_list.reshape(5,5))
# + id="lTobLyqQQr7j" colab_type="code" colab={}
import torch.nn as nn
import torch.nn.functional as F
class Flatten(nn.Module):
def forward(self, x):
return x.view(x.size(0), -1)
class Network(nn.Module):
def __init__(self, num_classes=10):
super(Network, self).__init__()
self.features = nn.Sequential(
nn.Conv2d(3, 32, kernel_size=3, stride=1, padding=1),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
nn.Conv2d(32, 64, kernel_size=3, stride=1, padding=1),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
nn.Conv2d(64, 128, kernel_size=3, stride=1, padding=1),
nn.ReLU(inplace=True),
nn.Conv2d(128, 128, kernel_size=3, stride=1, padding=1),
nn.ReLU(inplace=True),
nn.MaxPool2d(kernel_size=2, stride=2),
)
self.flatten = Flatten()
self.classifier = nn.Sequential(
nn.Linear(128 * 4 * 4, 512),
nn.ReLU(inplace=True),
nn.Dropout(),
nn.Linear(512, num_classes),
)
def forward(self, x):
x = self.features(x)
x = self.flatten(x)
x = self.classifier(x)
return x
# + id="NWIeNd2cLDwz" colab_type="code" colab={}
def train(model, device, optimizer, criterion, train_loader, preprocess=None):
model.train()
running_loss = 0
correct = 0
for batch_idx, (data, labels) in enumerate(train_loader):
if preprocess:
data = preprocess(data)
data, labels = data.to(device), labels.to(device)
optimizer.zero_grad()
outputs = model(data)
loss = criterion(outputs, labels)
loss.backward()
optimizer.step()
pred = outputs.argmax(dim=1)
correct += (pred == labels).sum().item()
running_loss += loss.item()
train_acc = correct / len(train_loader.dataset)
train_loss = running_loss / len(train_loader)
return {'acc':train_acc, 'loss':train_loss}
# + id="MsKx5EC2LHJM" colab_type="code" colab={}
def validation(model, device, criterion, val_loader, preprocess=None):
model.eval()
runnning_loss = 0
correct = 0
with torch.no_grad():
for data, labels in val_loader:
if preprocess:
data = preprocess(data)
data, labels = data.to(device), labels.to(device)
output = model(data)
pred = output.argmax(dim=1)
correct += (pred == labels).sum().item()
runnning_loss += criterion(output, labels).item()
val_acc = correct / len(val_loader.dataset)
val_loss = runnning_loss / len(val_loader)
return {'acc':val_acc, 'loss':val_loss}
# + id="45IUwOPRLJBS" colab_type="code" colab={}
def run(epochs, model, device, optimizer, criterion, train_loader, val_loader, preprocess=None):
history = {'train_acc':[], 'train_loss':[], 'val_acc':[], 'val_loss':[]}
for epoch in range(1, epochs+1):
train_ret = train(model, device, optimizer, criterion, train_loader, preprocess=preprocess)
val_ret = validation(model, device, criterion, val_loader, preprocess=preprocess)
print("Epoch:{} train_acc:{:.4f}% train_loss:{:.4f} val_acc:{:.4f}% val_loss:{:.4f}".format(
epoch, train_ret['acc'], train_ret['loss'], val_ret['acc'], val_ret['loss']))
history['train_acc'].append(train_ret['acc'])
history['train_loss'].append(train_ret['loss'])
history['val_acc'].append(val_ret['acc'])
history['val_loss'].append(val_ret['loss'])
return history
# + id="cGrsOr8olNCQ" colab_type="code" outputId="3754a217-d977-46a6-b132-aa100e40e8d8" executionInfo={"status": "ok", "timestamp": 1562490268798, "user_tz": -540, "elapsed": 1377410, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 1000}
import torch.optim as optim
model = Network()
from torchsummary import summary
summary(model, input_size=(3, 32, 32), device="cpu")
criterion = nn.CrossEntropyLoss()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
epochs = 100
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("device:",device)
model = model.to(device)
history = run(epochs, model, device, optimizer, criterion, train_loader, val_loader)
# + id="LJ2ErwlIWDXY" colab_type="code" colab={}
save_model = False
if (save_model):
torch.save(model.state_dict(),"cifar10.pt")
# + id="yXuyk4FOMP9L" colab_type="code" outputId="bf1004bf-d5a4-4436-e995-78b70a2da280" executionInfo={"status": "ok", "timestamp": 1562490269275, "user_tz": -540, "elapsed": 1302220, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 549}
def plot_history(history):
epochs = len(history['train_acc'],)
xaxis = range(epochs)
plt.figure()
plt.plot(xaxis, history['train_acc'], 'r-', label='train_acc')
plt.plot(xaxis, history['val_acc'], 'b-', label='val_acc')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('acc')
plt.figure()
plt.plot(xaxis, history['train_loss'], 'r-', label='train_loss')
plt.plot(xaxis, history['val_loss'], 'b-', label='val_loss')
plt.legend()
plt.xlabel('epoch')
plt.ylabel('loss')
plot_history(history)
# + id="VMfBgjPZZsT6" colab_type="code" outputId="0acb32ad-ff93-424c-d969-5443259171db" executionInfo={"status": "ok", "timestamp": 1562490271967, "user_tz": -540, "elapsed": 1300392, "user": {"displayName": "\u4e2d\u5d8b\u62d3\u90ce", "photoUrl": "", "userId": "11259832953657969381"}} colab={"base_uri": "https://localhost:8080/", "height": 238}
def test(model, classes, device, criterion, test_loader):
outsize = len(classes)
class_correct = list(0. for i in range(outsize))
class_total = list(0. for i in range(outsize))
runnning_loss = 0
with torch.no_grad():
for data in test_loader:
imgs, labels = data
n_batch = imgs.size(0)
imgs, labels = imgs.to(device), labels.to(device)
outputs = model(imgs)
runnning_loss += criterion(outputs, labels).item()
pred = torch.argmax(outputs, 1)
c = (pred == labels).squeeze()
for i in range(n_batch):
label = labels[i]
class_correct[label] += c[i].item()
class_total[label] += 1
for i in range(outsize):
print('Accuracy of %5s : %2d %%' % (
classes[i], 100 * class_correct[i] / class_total[i]))
test_acc = int(sum(class_correct)/sum(class_total)*100)
test_loss = runnning_loss / len(test_loader)
print('acc : {:2d}%'.format(test_acc))
print('loss : {:.4f}'.format(test_loss) )
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print("device:",device)
model = model.to(device)
test(model, classes, device, criterion, test_loader)
|
PyTorch_RandomErasing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Week 04, Assignment 03: Methods II
# # Namespaces
# In this lesson we will continue practicing method coding and understanding namespaces
# ### Objective 1: Understanding namespaces
# Explain the difference between a local and global namespaces. What happens to the namespace when a function is defined?
#
# ### Objective 2: Namespaces in practice
# Run the following code and answer the questions below.
# +
# dice_simulator.py
from random import randint
SIDES = 6
def roll_dice(times):
result = []
for i in range(times):
result.append(randint(1,SIDES))
return result
roll_dice(10)
# -
# From all the variables `randint`, `SIDES`, `roll_dice`,`times`, and `result`. Which are local variables and which are global?
#
# What happens when we move `result` outside the method to line 4?
#
# Write the code `print(result)` one line 12 and re-run the cell. Explain what is happening
# ### Objective 3: locals() and globals()
# In the cell below call the function `locals()` and `globals()`. Explain what these functions return. *Hint: use help() to help your understanding*
#
# ### Objective 4: Methods in practice
# Copy and modify the code from Objective 2 to do the following:
# * Let `results` instead be a global variable will continuously add dice roll results to the list.
# * Create a new function called `clear_rolls` that removes the items from `results`
# * Create a new function called `average` which calculates the average between all current dice rolls in `results`
# ### Preperation for tomorrow...
# * Preread the section on lambda functions
# * Read up on the use of anonymous functions with iterators
# * Lookup the use of of the `map()` function
|
python/Assignment_One/Assignment 1.5.3_Methods_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/https-deeplearning-ai/tensorflow-1-public/blob/adding_C3/C3/W3/ungraded_labs/C3_W3_Lab_1_single_layer_LSTM.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="zX4Kg8DUTKWO"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="OSPyf0qgs_WX"
# **Note:** This notebook can run using TensorFlow 2.5.0
# + id="Dlj4bfz_s_WY"
# #!pip install tensorflow==2.5.0
# + [markdown] id="rFiCyWQ-NC5D"
# # Single Layer LSTM
# + id="Y20Lud2ZMBhW"
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow_datasets as tfds
import tensorflow as tf
print(tf.__version__)
# + id="AW-4Vo4TMUHb"
# Get the data
dataset, info = tfds.load('imdb_reviews/subwords8k', with_info=True, as_supervised=True)
train_dataset, test_dataset = dataset['train'], dataset['test']
# + id="DVfhKpHsPOxq"
tokenizer = info.features['text'].encoder
# + id="ffvRUI0_McDS"
BUFFER_SIZE = 10000
BATCH_SIZE = 64
train_dataset = train_dataset.shuffle(BUFFER_SIZE)
train_dataset = train_dataset.padded_batch(BATCH_SIZE, tf.compat.v1.data.get_output_shapes(train_dataset))
test_dataset = test_dataset.padded_batch(BATCH_SIZE, tf.compat.v1.data.get_output_shapes(test_dataset))
# + id="FxQooMEkMgur"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(tokenizer.vocab_size, 64),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)),
tf.keras.layers.Dense(64, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
# + id="QKI5dfPgMioL"
model.summary()
# + id="Uip7QOVzMoMq"
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="7mlgzaRDMtF6"
NUM_EPOCHS = 10
history = model.fit(train_dataset, epochs=NUM_EPOCHS, validation_data=test_dataset)
# + id="Mp1Z7P9pYRSK"
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.plot(history.history['val_'+string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.legend([string, 'val_'+string])
plt.show()
# + id="R_sX6ilIM515"
plot_graphs(history, 'accuracy')
# + id="RFEXtKtqNARB"
plot_graphs(history, 'loss')
# + id="hCq86yhZs_Wc"
|
C3/W3/ungraded_labs/C3_W3_Lab_1_single_layer_LSTM.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> Bivariate Analysis
#
# <font color=darkblue>
#
# Through bivariate analysis we try to analyze two variables simultaneously. As opposed to univariate analysis where we check the characteristics of a single variable, in bivariate analysis we try to determine if there is any relationship between two
# variables.
#
#
#
# There are essentially 3 major scenarios that we will come across when we perform bivariate analysis
# 1. Both variables of interest are qualitative
# 2. One variable is qualitative and the other is quantitative
# 3. Both variables are quantitative
#
#
#
# For the purpose of this exercise, we will explore few most popular techniques to perform bivariate analysis.
#
# The following plots are not limited to the headings they are under. They are the options we have if we face a certain scenario.
#
#
#
# ### Numerical vs. Numerical
# 1. Scatterplot
# 2. Line plot
# 3. Heatmap for correlation
# 4. Joint plot
# ### Categorical vs. Numerical
# 1. Bar chart
# 2. Violin plot
# 3. Categorical box plot
# 4. Swarm plot
#
# ## Two Categorical Variables
# 1. Bar chart
# 2. Grouped bar chart
# 3. Point plot
# <span style="font-family: Arial; font-weight:bold;font-size:1.9em;color:#0e92ea">Case Study
# <span style="font-family: Arial; font-weight:bold;font-size:0.7em;color:#b53f05"> (Suicide Rates Overview 1985 to 2016)
# + active=""
# Our aim is to explore the data of suicide rates.
#
# Lets start using the above techniques on a practical dataset. We will be using data about suicide rates from 1985 to 2026.
#
# source: https://www.kaggle.com/russellyates88/suicide-rates-overview-1985-to-2016
#
#
# Dataset Feature List
# country
# year
# sex
# age
# suicides_no
# population
# suicides/100k pop
# country-year
# HDI for year
# gdp_for_year
# gdp_per_capita
# generation
# -
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> 1.1 Loading the libraries
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> 1.2 Import the dataset
data=pd.read_csv('master.csv')
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.3 Check the data head()
data.head()
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.4 Check the data.describe()
data.describe()
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> 1.5 Check the columns
data.columns
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.6 Checking the shape of data
data.shape
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.7 Count the datatypes
data.dtypes.value_counts()
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.8 Check the data set information
data.info()
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.9 Checking the dataset missing values
def missing_check(df):
total = df.isnull().sum().sort_values(ascending=False) # total number of null values
percent = (df.isnull().sum()/df.isnull().count()).sort_values(ascending=False) # percentage of values that are null
missing_data = pd.concat([total, percent], axis=1, keys=['Total', 'Percent']) # putting the above two together
return missing_data # return the dataframe
missing_check(data)
data[['suicides_no','population','suicides/100k pop','gdp_per_capita ($)']].describe() #descriptive stats of continuous columns
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.10 Frequency table for Age
# One-Way Tables
my_tab = pd.crosstab(index=data["age"], # Make a crosstab
columns="count") # Name the count column
my_tab
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.11 Bar plot to check Number of Suicides by top Countries
# # This is an example of Numerical vs Categorical.
data.groupby(by=['country'])['suicides_no'].sum().reset_index().sort_values(['suicides_no']).tail(10).plot(x='country',
y='suicides_no',
kind='bar',
figsize=(15,5))
plt.show()
# <font color = darkblue>
#
# * Russia has the highest number of suicides followed by the US and Japan
# * Russia, the US and Japan have exceptionally high counts of suicides compared to the others in the lot
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.12 Bar plot to check Number of Suicides by bottom Countries
# +
data.groupby(by=['country'])['suicides_no'].sum().reset_index().sort_values(['suicides_no'],
ascending=True).head(10).plot(x='country',y='suicides_no',kind='bar', figsize=(15,5))
plt.show()
# -
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.13 Bar plot for Number of Suicides Vs Age
plt.figure(figsize=(10,5)) # setting the figure size
ax = sns.barplot(x='age', y='suicides_no', data=data, palette='muted') # barplot
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> 1.14 Bar plot Number of Suicides Vs Sex
plt.figure(figsize=(8,4))
ax = sns.barplot(x="sex", y="suicides_no", data=data)
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.15 Bar plot Number of Suicides Vs generation
# +
plt.figure(figsize=(15,5))
ax = sns.barplot(x='generation', y='suicides_no', data=data)
# -
# <font color = 'darkblue'>
#
# * Suicides are the highest among the Boomers and lowest among Generation Z
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.15 Scatter plot Number of Suicides Vs population
# # This is an examples of Numerical vs. Numerical
# +
figure = plt.figure(figsize=(15,5))
ax = sns.scatterplot(x=data['population'],y='suicides_no', data=data, size = "suicides_no") # scatter plot
# +
figure = plt.figure(figsize=(50,15))
ax = sns.regplot(x='population',y='suicides_no', data=data ) # regression plot - scatter plot with a regression line
# -
#Here we plotting a line plot.
sns.lineplot(x='population',y='suicides_no', data=data.head() )
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">Scatter plot Number of Suicides/100k Population Vs GDP Per Capita
# +
figure = plt.figure(figsize=(15,7))
sns.scatterplot(x='gdp_per_capita ($)', y='suicides/100k pop', data=data) # scatter plot
plt.show()
# -
# <font color = 'darkblue'>
#
# * Looks like higher suicide rates are a bit more prevalent in countries with higher GDP
# * However, it doesn't look like there any significant correlation between the two
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">Correlation among pairs of continuous variables
plt.figure(figsize=(10,5))
sns.heatmap(data.corr(), annot=True, linewidths=.5, fmt= '.1f', center = 1 ) # heatmap
plt.show()
# <font color = 'darkblue'>
#
# * Instead of doing a simple df.corr(), we can use the heatmap when there are large number of variables
# * The color helps with picking out the most correlated easily
# * In the above case, we don't have a lot of variables the plot is just for demonstration
# * The darker the color higher the correlation
# * None of the attributes seem have correlation of real significance
# * Some obvious correlations are that in a larger population, it is very likely that the number of suicides will be more
# * Human Development Index - gdp per capita is the only pair with the most correlation
#
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.17.1 Bar plot To check Number of suicides by sex and age (three variables used to generate a single plot)
# # This is an example of Numerical and 2 Categorical variables.
plt.figure(figsize=(15,5))
sns.barplot(data=data,x='sex',y='suicides_no',hue='age')
plt.show()
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">1.17.2 Bar plot To check Number of suicides by sex and Generation(three variables used to generate a single plot)
plt.figure(figsize=(15,5))
sns.barplot(data=data,x='sex',y='suicides_no',hue='generation')
plt.show()
# <font color = 'darkblue'>
#
# * Suicides in males apart from being higher, has a slight variation in distribution across generations compared to the suicides of females across generation
# * In case of males, Generation X has higher suicide numbers compared to G.I Generation but in case of females, it is the opposite
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> 1.18 Checking the No.of suicides: Country Vs Sex
# +
suic_sum_m = data['suicides_no'].groupby([data['country'],data['sex']]).sum() # number of suicides by country and sex
suic_sum_m = suic_sum_m.reset_index().sort_values(by='suicides_no',ascending=False) # sort in descending order
most_cont_m = suic_sum_m.head(10) # getting the top ten countries in terms of suicides
fig = plt.figure(figsize=(15,5))
plt.title('Count of suicides for 31 years.')
sns.barplot(y='country',x='suicides_no',hue='sex',data=most_cont_m,palette='Set2');
plt.ylabel('Count of suicides')
plt.tight_layout()
# -
# <font color = 'darkblue'>
#
# * Japan has a higher proportion of female suicides compared to the countries with overall suicide rates even more high
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea">Average number of suicides across each generation for a given gender along with the confidence intervals - Point Plot
# +
plt.figure(figsize=(15,5))
sns.pointplot(x="generation", y="suicides_no", hue = 'sex', data=data)
plt.show()
# -
# <font color = 'darkblue'>
#
# * The graph tells us the average suicides along with the confidence intervals
# * Suicides among females in general don't seem to be fluctuating a lot
# * Average suicides of Gen-Z are almost equally distributed across genders
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> Distribution of population across each generation - Violin plot
plt.figure(figsize=(10,5))
sns.violinplot(x=data.generation, y=data['population'])
plt.show()
# <font color = 'darkblue'>
#
# * The plot is similar to a box plot but here, we get a density function
# * Distribution of population across every generation is highly skewed
# * Potential of lot of outliers
# * Try it yourself - Go ahead and check if there really are a lot of outliers in populations across each generation
# Hint: use a boxplot
# # Checking trends with Temporal Data
# Temporal data is simply data that represents a state in time, such as the land-use patterns of Hong Kong in 1990,
# or total rainfall in Honolulu on July 1, 2009. Temporal data is collected to analyze weather patterns and other environmental variables, monitor traffic conditions, study demographic trends, and so on. This data comes from many sources ranging
# from manual data entry to data collected using observational sensors or generated from simulation models.
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> Checking pattern using Trend plot (1985-2015) suides Rate Vs Years
# +
data[['year','suicides_no']].groupby(['year']).sum().plot(figsize=(15,5))
plt.show()
# -
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> Checking pattern using Trend plot (1985-2015) Population Vs Years
# +
data[['year','population']].groupby(['year']).sum().plot(figsize=(15,5))
plt.show()
# -
# <span style="font-family: Arial; font-weight:bold;font-size:1.5em;color:#0e92ea"> Checking pattern using Trend plot (1985-2015) suicides/100k pop Vs Years
# +
data[['year','suicides/100k pop']].groupby(['year']).sum().plot(figsize=(15,5))
plt.show()
# -
# # Pandas Profiling with IPL data
# Generates profile reports from a pandas DataFrame.
# pandas_profiling extends the pandas DataFrame with df.profile_report() for quick data analysis.
#
# Link - https://pypi.org/project/pandas-profiling/
#Installation step
# #!pip install pandas-profiling
#or
import sys
# !{sys.executable} -m pip install pandas-profiling
#import pandas_profiling
import pandas_profiling
df = pd.read_csv('match_data.csv')
# It has many useful functionalities but the best one is to generate an EDA report as given below.
#Getting the pandas profiling report
pandas_profiling.ProfileReport(df)
#Getting an html file as output here
pandas_profiling.ProfileReport(df).to_file("output.html")
# You can see the report in your directory in html format. It will give you a structured summarized view of statistics of every columns with interactive options to explore more.
# # Steps Ahead
# Try it yourself -
#
# Create a Pandas Profile report for innings_data.csv in IPL Data.
# <span style="font-family: Arial; font-weight:bold;font-size:1.9em;color:#ez0028"> Happy Learning
|
01 Fundementals of AIML/Week 3 - Data Visualization and EDA/EDA-CaseStudy_and_PandasProfiling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Obsah dnesnej prednasky
# ## 1. Iterator a generator
# ## 2. Lenive vyhodnocovanie (Lazy evaluation)
# + [markdown] slideshow={"slide_type": "slide"}
# # 1. Iterator a Generator
#
# inspirovane http://www.python-course.eu/python3_generators.php
#
# Oba sa pouzivaju na postupne prechadzanie cez datovu strukturu alebo postupne vykonavanie algoritmu po krokoch.
# + [markdown] slideshow={"slide_type": "slide"}
# # Iterator
# * je objekt, ktory ma funkciu `__next__` a funkciu `__iter__`, ktora vracia `self`
# * je to vseobecnejsi pojem ako generator
# * da sa pouzivat napriklad na iterovanie cez kolekciu bez toho, aby sme vedeli aka je jej vnutorna struktura. Staci definovat funkciu `__next__`. Podobny koncept sa da najst vo vela jazykoch. Napriklad aj v Jave.
# + [markdown] slideshow={"slide_type": "slide"}
# # Iterator sa napriklad implicitne pouziva pri prechadzani kolekcii for cyklom
# -
cities = ["Paris", "Berlin", "Hamburg", "Frankfurt", "London", "Vienna", "Amsterdam", "Den Haag"]
for location in cities:
print("location: " + location)
# + slideshow={"slide_type": "slide"}
dir(cities.__iter__())
# + slideshow={"slide_type": "slide"}
type(cities.__iter__())
# -
print(type(cities.__iter__()))
print(type(cities.__iter__().__iter__()))
print(cities.__iter__().__next__())
# + [markdown] slideshow={"slide_type": "slide"}
# # Rovnako sa pouzivaju iteratory aj pri prechadzani inych kolekcii
# -
capitals = { "France":"Paris", "Netherlands":"Amsterdam", "Germany":"Berlin", "Switzerland":"Bern", "Austria":"Vienna"}
for country in capitals:
print("The capital city of " + country + " is " + capitals[country])
# + [markdown] slideshow={"slide_type": "slide"}
# # Generator
# * kazdy generator objekt je iterator, ale nie naopak
# * tento pojem sa pouziva na pomenovanie funkcie (generator funkcia) ako aj jej navratovej hodnoty (generator objekt)
# * generator objekt sa vytvara volanim funkcie (generator funkcie), ktora pouziva `yield`
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Generator pouziva vyraz `yield` na zastavenie vykonavania a na vratenie hodnoty
#
# * Vykonavanie sa spusta volanim funkcie `next()` (alebo metody `__next__()`)
# * Dalsie volanie zacina od posledneho `yield`
# * Medzi volaniami sa hodnoty lokalnych premennych uchovavaju.
#
# ## Pozor, toto nie je ten isty `yield` ako je v Ruby
# * V Ruby je `yield` volanie bloku asociovaneho s metodou
# * `yield` v Ruby vlastne odovzdáva kontrolu nejakému bloku kódu. Je to skôr podobné volanie lambda funkcie predanej parametrom, len je to inak zapísané
# * V Ruby je nieco podobne generatorom napriklad trieda `Enumerator`
#
# [http://stackoverflow.com/questions/2504494/are-there-something-like-python-generators-in-ruby](http://stackoverflow.com/questions/2504494/are-there-something-like-python-generators-in-ruby)
# + [markdown] slideshow={"slide_type": "slide"}
# # Jednoduchý príklad generátoru
# -
def city_generator():
yield "Konstanz"
yield "Zurich"
yield "Schaffhausen"
yield "Stuttgart"
gen = city_generator()
next(gen)
# + [markdown] slideshow={"slide_type": "slide"}
# # Vo vnutri generator funkcie mozem pouzivat cyklus
# -
cities = ["Konstanz", "Zurich", "Schaffhausen", "Stuttgart"]
def city_generator():
for city in cities:
yield city
gen = city_generator()
next(gen)
# Tento genereator vlastne len supluje iterator, ktory je nad polom, ale ten cyklus moze robit aj nieco viac a vtedy to uz moze byt zaujimavejsie (ukazem neskor)
# + [markdown] slideshow={"slide_type": "slide"}
# # Generator funkcia moze prijmat parametre
# -
def city_generator(local_cities):
for city in local_cities:
yield city
gen = city_generator(["Konstanz", "Zurich", "Schaffhausen", "Stuttgart"])
next(gen)
# + [markdown] slideshow={"slide_type": "slide"}
# # Trik ako napisat generator, ktory velmi casto funguje
#
# Uloha: Mame sekvenciu cisel a chceme vytvorit pohyblivy priemer dvoch po sebe nasledujucich cisel pre celu sekvenciu.
#
# napr:
#
# sekvencie = [1,2,3,4,5]
#
# pohyblivy priemer = [(0+1)/2, (1+2)/2, (2+3)/2, (3+4)/2, (4+5)/2] = [0.5, 1.5, 2.5, 3.5, 4.5]
# + [markdown] slideshow={"slide_type": "slide"}
# # Ako by ste to napisali imperativne ak chcete vysledok len zapisat do konzoly?
# + slideshow={"slide_type": "slide"}
sequence = [1,2,3,4,5]
previous = 0
for actual in sequence:
print((actual + previous) / 2)
previous = actual
# + [markdown] slideshow={"slide_type": "slide"}
# # Zabalim to do funkcie
# + slideshow={"slide_type": "-"}
sequence = [1,2,3,4,5]
def moving_average(sequence):
previous = 0
for actual in sequence:
print((actual + previous) * 0.5)
previous = actual
moving_average(sequence)
# + [markdown] slideshow={"slide_type": "slide"}
# # Vymenim print za `yield`
# -
sequence = [1,2,3,4,5]
def moving_average(sequence):
previous = 0
for actual in sequence:
yield (actual + previous) * 0.5
previous = actual
gen = moving_average(sequence)
next(gen)
print(list(moving_average(sequence)))
# + [markdown] slideshow={"slide_type": "slide"}
# # Hotovo
#
# Jednoduchy trik ako napisat generator
#
# 1. napiste kod, kde priebezne vysledky len zapsiujete funkciou print
# 2. zabalte kod do funkcie
# 3. nahradte print prikazom yield
# + [markdown] slideshow={"slide_type": "slide"}
# # Pomocou generatoru by sa dala napriklad spravit funkcia `map`
# -
def map(f, seq):
for x in seq:
print(f(x))
def map(f, seq):
for x in seq:
yield f(x)
# + [markdown] slideshow={"slide_type": "slide"}
# # Porovnajte si ako by vyzerala implementacia `map` v python2 a python3
# -
def map(f, seq): # V pythone 2 map vracia list, implementacia by m<NAME> napriklad takato
result = [] # mame premennu, ktoru postupne upravujeme a nafukujeme
for x in seq:
result.append(f(x))
return result
def map(f, seq): # V pythone 3 map je generator a zabera konstantne mnozstvo pamati
for x in seq:
yield f(x)
# + [markdown] slideshow={"slide_type": "slide"}
# # Niektore generatory sa daju nahradit funkciou `map`
# +
a, b = 1, 10
def squares(start, stop):
for i in range(start, stop):
yield i * i
generator = squares(a, b)
print(generator)
print(next(generator))
print(list(generator))
# + slideshow={"slide_type": "slide"}
generator = map(lambda i: i*i, range(a, b))
print(generator)
print(next(generator))
print(list(generator))
# + [markdown] slideshow={"slide_type": "slide"}
# # List comprehension tiez moze vytvarat generator
# -
generator = (i*i for i in range(a, b)) # rozdiel oproti kalsickemu LC je v zatvorkach [] => ()
print(generator)
print(next(generator))
print(list(generator))
# + [markdown] slideshow={"slide_type": "slide"}
# # Explicitny generator ma ale vacsiu vyjadrovaciu silu
#
# Nie je obmedzeny len na formu ktoru pouziva funkcia map:
# -
def generator(funkcia, iterator):
for i in iterator:
yield funkcia(i)
# + [markdown] slideshow={"slide_type": "slide"}
# # Na co je to cele dobre?
# + [markdown] slideshow={"slide_type": "slide"}
# Tu sa dostavame k druhej casti prednasky
# # 2. Lenive vyhodnocovanie - Lazy evaluation
# + [markdown] slideshow={"slide_type": "slide"}
# # Strategie vyhodnocovania
#
# ## Skratene vyhodocovanie (Short-circuit)
# ## Netrpezlive vyhodocovanie (Eager)
# ## Lenive vyhodnocovanie (Lazy)
# ## Vzdialene vyhodocovanie (Remote)
# ## Ciastocne vyhodocovanie (Partial)
#
# https://en.wikipedia.org/wiki/Evaluation_strategy
# + [markdown] slideshow={"slide_type": "slide"}
# # Skratene vyhodnocovanie
#
# Urcite si pamatate z Proceduralneho programovania
# +
def fun1():
print('prva')
return True
def fun2():
print('druha')
return True
if fun1() or fun2():
pass
# + [markdown] slideshow={"slide_type": "slide"}
# # Lenive vyhodocovanie
#
# Oddaluje vyhodnocovanie az do doby, ked je to treba
# -
pom = (x*x for x in range(5))
next(pom) #prvok z generatora sa vyberie az ked ho je treba a nie pri vytvoreni generatora
# + [markdown] slideshow={"slide_type": "slide"}
# # Nedockave vyhodocovanie
# Opak leniveho vyhodnotenia. Vyraz sa vyhodnoti hned ako je priradeny do premennej. Toto je typicky sposob vyhodnocaovania pri vacsine programovacich jazykoch.
# -
pom = [x*x for x in range(5)]
pom[4] # vyraz sa hned vyhodnocuje cely
# + [markdown] slideshow={"slide_type": "slide"}
# ## Vyhody nedockaveho vyhodnocovania
# * programator moze kontrolovat poradie vykonavania
# * nemusi sledovat a planovat poradie vyhodnocovania
#
# ## Nevyhody
# * neumoznuje vynechat vykonavanie kodu, ktory vobec nie je potrebny (spomente si na priklad so Sparkom z minuleho tyzdna)
# * neda sa vykonavat kod, ktory je v danej chvili dolezitejsi
# * programator musi organizovat kod tak, aby optimalizoval poradie vykonavania
#
# Moderne kompilatory ale uz niektore veci vedia optimalizovat za programatora
# + [markdown] slideshow={"slide_type": "slide"}
# # Vzdialene vyhodnocovanie
#
# * Vyhodnocovanie na vzdialenom pocitaci.
# * Hociaky vypoctovy model, ktory spusta kod na inom stroji.
# * Client/Server, Message passing, MapReduce, Remote procedure call (RPC)
# + [markdown] slideshow={"slide_type": "slide"}
# # Partial evaluation
#
# * Viacero optimalizacnych strategii na to aby sme vytvorili program, ktory bezi rychlejsie ako povodny program.
# * Napriklad predpocitavanie kodu na zaklade dat, ktore su zname uz v case kompilacie.
# * Mamoization (preklad Memoizacia?) - nevykonavanie (cistych) funkcii s rovnakymi vstupmi opakovane. V podstate ide o caschovanie vystupov volani funkcii
# * Partial application - fixovanie niektorych parametrov funkcie a vytvorenie novej s mensim poctom parametrov.
# + [markdown] slideshow={"slide_type": "slide"}
# # Lenive vyhodnocovanie moze zrychlit vyhodocovanie
# -
# %%time
print(2+2)
# + slideshow={"slide_type": "slide"}
# %%time
import time
def slow_square(x):
time.sleep(0.2)
return x**2
generator = map(slow_square, range(10))
print(generator)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Funkcia slow_square sa zatial nespustila ani raz. Preto je ten cas tak maly
# + slideshow={"slide_type": "slide"}
# %%time
# co sa stane ak budeme chciet transformovat generator na zoznam. Teda spustit ru pomalu funkciu na kazdom prvku?
print(list(generator))
# + slideshow={"slide_type": "slide"}
# %%time
# Aj ked chceme len cast pola, tak musime transformovat vsetky prvky
generator = map(slow_square, range(10))
pole = list(generator)
print(pole[:5])
# + slideshow={"slide_type": "slide"}
# Mozeme si ale skusit definovat funkciu, ktora nam vyberie len tu cast prvkov, ktore chceme
def head(iterator, n):
result = []
for _ in range(n):
result.append(next(iterator))
return result
# +
# %%time
print(head(map(slow_square, range(10)), 5))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Ta pomala operacia sa vykonala len tolko krat, kolko sme potrebovali a to co sme nepotrebovali sa nemuselo nikdy vykonat.
# + slideshow={"slide_type": "slide"}
# %%time
# tuto funkciu sme si ale nemuseli definovat sami. Nieco take uz existuje
from itertools import islice
generator = map(slow_square, range(10000))
print(list(islice(generator, 5)))
# + [markdown] slideshow={"slide_type": "slide"}
# # Funkciu `islice` si zapamatajte, este ju budeme vela krat pouzivat
# + [markdown] slideshow={"slide_type": "slide"}
# # Lenive vyhodnocovanie setri pamat
# +
from operator import add
from functools import reduce
print(reduce(add, [x*x for x in range(10000000)]))
print(reduce(add, (x*x for x in range(10000000)))) # rozdiel je len v zatvorkach
# + [markdown] slideshow={"slide_type": "slide"}
# ## skusim si vyrobit funkciu, ktora mi bude priebezne pocitat a vypisovat aktualnu spotrebu pamati premennych na halde pocas toho ako budeme spocitavat cisla
# + slideshow={"slide_type": "slide"}
from functools import reduce
import gc
import os
import psutil
process = psutil.Process(os.getpid())
def print_memory_usage():
print(process.memory_info().rss)
counter = [0] # Toto je hnusny hack a slubujem, ze nabuduce si povieme ako to spravit lepsie. Spytajte sa ma na to!
# Problem je v tom, ze potrebujem pocitadlo, ktore bude dostupne vo funkcii,
# ale zaroven ho potrebujem inicializovat mimo tejto funkcie.
# Teraz som zaspinil funkciu pouzitim mutable datovej struktury a globalneho priestoru mien. 2xFuj!!!
def measure_add(a, result, counter=counter):
if counter[0] % 2000000 == 0:
print_memory_usage()
counter[0] = counter[0] + 1
return a + result
# + slideshow={"slide_type": "slide"}
gc.collect()
counter[0] = 0
print_memory_usage()
print('vysledok', reduce(measure_add, [x*x for x in range(10000000)]))
# + slideshow={"slide_type": "slide"}
gc.collect()
counter[0] = 0
print_memory_usage()
print('vysledok', reduce(measure_add, (x*x for x in range(10000000))))
# + [markdown] slideshow={"slide_type": "slide"}
# # Ani ked su funkcie povnarane do seba a kolekcia sa predava ako parameter, nikdy nie je cela v pamati
# map, filter, reduce aj list comprehension vnutorne pracuju s kolekciami ako s iteratormi
# + slideshow={"slide_type": "-"}
gc.collect()
counter[0] = 0
print_memory_usage()
print('vysledok', reduce(measure_add, filter(lambda x: x%2 == 0, map(lambda x: x*x, range(10000000)))))
# a pokojne by som to mohol vnarat dalej
# + [markdown] slideshow={"slide_type": "slide"}
# # Ked vieme, ze generator sa vyhodnocuje lenivo, tak nam nic nebrani vlozit do neho nekonecny cyklus
# +
def fibonacci():
"""Fibonacci numbers generator"""
a, b = 1, 1
while True:
yield a
a, b = b, a + b
f = fibonacci()
# -
print(list(islice(f, 10)))
# + [markdown] slideshow={"slide_type": "slide"}
# # Voila, nekonecna datova struktura, ktora nezabera skoro ziadnu pamat dokedy ju nechcem materializovat celu.
# + slideshow={"slide_type": "-"}
# POZOR!!!
# Toto netreba pustat. Zozralo by to cely vykon procesoru a postupne aj celu pamat.
list(fibonacci())
# POZOR!!!
# + [markdown] slideshow={"slide_type": "slide"}
# # Vedeli by ste to pouzit na:
# * generator prvocisel?
# * citanie z velmi velkeho suboru, ktory vam nevojde do pamati?
# * citanie dat z nejakeho senzoru, ktory produkuje kludne nekonecne mnozstvo dat?
# + [markdown] slideshow={"slide_type": "slide"}
# # Dalo by sa to pouzit napriklad na cakanie na data
# Predstavte si, ze mate subor, do ktoreho nejaky proces zapisuje logy po riadkoch a vy ich spracovavate.
#
# Ako by ste spravili iterovanie cez riadky suboru tak, aby ste cakali na dalsie riadky ak dojdete na koniec suboru?
#
# inspirovane - http://stackoverflow.com/questions/6162002/whats-the-benefit-of-using-generator-in-this-case
# + slideshow={"slide_type": "slide"} language="bash"
# echo -n 'log line' > log.txt
# + slideshow={"slide_type": "slide"}
import time
# + slideshow={"slide_type": "-"}
# s generatorom napriklad takto
def read(file_name):
with open(file_name) as f:
while True:
line = f.readline()
if not line:
time.sleep(0.1)
continue
yield line
lines = read("log.txt")
print(next(lines))
# -
print(next(lines))
for line in lines:
print(line)
# + [markdown] slideshow={"slide_type": "slide"}
# # Toto by som vedel spravit aj bez generatora ale ...
# * nemal by som oddelenu logiku cakania a spracovavania riadku
# * zneuzivam necistu funkciu print
# * nevedel by som priamociaro znovupouzivat generator, vzdy by som to musel kodit odznova
# * jedine, ze by som pouzil funkciu ako parameter
# * stale tam ale zostava problem ako vratit viacero hodnot z jednej funkcie
# * nevedel by som pekne transparentne, lenivo iterovat
# -
while True:
line = logfile.readline()
if not line:
time.sleep(0.1)
continue
print line
# + [markdown] slideshow={"slide_type": "slide"}
# # Generator moze byt aj trochu zlozitejsi, napriklad rekurzivny
# + [markdown] slideshow={"slide_type": "slide"}
# # Predstavte si takuto stromovu strukturu
# + slideshow={"slide_type": "-"}
class Node(object): # toto je v podstate N-arny strom
def __init__(self, title, children=None):
self.title = title
self.children = children or []
tree = Node(
'A', [
Node('B', [
Node('C', [
Node('D')
]),
Node('E'),
]),
Node('F'),
Node('G'),
])
# + slideshow={"slide_type": "slide"}
def node_recurse_generator(node):
yield node
for n in node.children:
for rn in node_recurse_generator(n):
yield rn
[node.title for node in node_recurse_generator(tree)]
# -
# http://stackoverflow.com/posts/7634323/edit
# # Uloha na volny cas
#
# Vedeli by ste vytvorit datovu strukturu `list_r`, ktora by bola tvorena dvojicou prvy prvok zoznamu a jeho zvysok (first, rest)? Vedeli by ste vytvorit rekurzivne funkcie a generatory, ktore by spracovavali takyto zoznam (vratenie prvku na indexe, pridanie prvku, odstranenie prvku, prevratenie poradia, ..)?
#
# Ak ano, tak viete simulovat zakladnu datovu strukturu LISPu a mozete pracovat s Pythonom ako keby to bol LISP.
# + [markdown] slideshow={"slide_type": "slide"}
# # Ale castokrat sa to da aj bez pouzitia rekurzie
#
# http://stackoverflow.com/questions/26145678/implementing-a-depth-first-tree-iterator-in-python
# +
from collections import deque
def node_stack_generator(node):
stack = deque([node]) # tu si uchovavam stav prehladavania kedze nepouzivam call stack v rekurzii
while stack:
# Pop out the first element in the stack
node = stack.popleft()
yield node
# push children onto the front of the stack.
# Note that with a deque.extendleft, the first on in is the last
# one out, so we need to push them in reverse order.
stack.extendleft(reversed(node.children))
[node.title for node in node_stack_generator(tree)]
# + [markdown] slideshow={"slide_type": "slide"}
# # Uloha na volny cas
#
# Vedeli by ste tieto dva generatory upravit pre binarny strom?
# + [markdown] slideshow={"slide_type": "slide"}
# # Rekurzivny generator sa da napriklad pouzit na vyrabanie permutacii
# -
def permutations(items):
n = len(items)
if n==0:
yield []
else:
for i in range(len(items)):
for cc in permutations(items[:i]+items[i+1:]):
yield [items[i]]+cc
for p in permutations('red'):
print(''.join(p))
for p in permutations("game"):
print(''.join(p) + ", ", end="")
# + [markdown] slideshow={"slide_type": "slide"}
# # Spominate si na `from itertools import islice` ?
# +
def fibonacci():
"""Fibonacci numbers generator"""
a, b = 1, 1
while True:
yield a
a, b = b, a + b
print(list(islice(fibonacci(), 5)))
# + [markdown] slideshow={"slide_type": "slide"}
# # Pomocou generatoru si vieme vytvorit jej ekvivalent
# Generator generatorov alebo fukcia, ktora dostava ako parameter generator vracia iny generator
# -
def firstn(g, n): # generator objekt je parametrom generator funkcie
for i in range(n):
yield next(g)
list(firstn(fibonacci(), 10))
|
7/prednaska/Generator.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# +
# read base zoning input files
z_p = pd.read_csv(r'C:\Users\ywang\Box\Modeling and Surveys\Urban Modeling\Bay Area UrbanSim\PBA50\Current PBA50 Large General Input Data\2020_11_05_zoning_parcels_hybrid_pba50.csv',
usecols = ['PARCEL_ID', 'geom_id', 'zoning_id'])
z_lookup = pd.read_csv(r'C:\Users\ywang\Box\Modeling and Surveys\Urban Modeling\Bay Area UrbanSim\PBA50\Current PBA50 Large General Input Data\2020_11_05_zoning_lookup_hybrid_pba50.csv')
basezoning = z_p.merge(z_lookup, left_on = 'zoning_id', right_on = 'id', how='left')
print('basezoning has {} records'.format(basezoning.shape[0]))
# -
# rename columns
basezoning.rename(columns = {'juris': 'juris_basezoning',
'max_dua': 'max_dua_basezoning',
'max_far': 'max_far_basezoning',
'HS': 'HS_basezoning',
'HT': 'HT_basezoning',
'HM': 'HM_basezoning',
'OF': 'OF_basezoning',
'HO': 'HO_basezoning',
'SC': 'SC_basezoning',
'IL': 'IL_basezoning',
'IW': 'IW_basezoning',
'IH': 'IH_basezoning',
'RS': 'RS_basezoning',
'RB': 'RB_basezoning',
'MR': 'MR_basezoning',
'MT': 'MR_basezoning',
'ME': 'ME_basezoning'}, inplace=True)
display(basezoning.head())
# +
# read zoningmods input files
p_g = pd.read_csv(r'C:\Users\ywang\Box\Modeling and Surveys\Urban Modeling\Bay Area UrbanSim\PBA50\PBA50 Final Blueprint Large General Input Data\2020_11_10_parcels_geography.csv',
usecols = ['PARCEL_ID', 'fbpzoningmodcat', 'nodev'])
zmods = pd.read_csv(r'C:\Users\ywang\Documents\GitHub\bayarea_urbansim\data\zoning_mods_24.csv')
zoningmods = p_g.merge(zmods, on='fbpzoningmodcat', how='left')
print('zoningmods has {} records'.format(zoningmods.shape[0]))
# -
zoningmods.head()
# +
zoningmerge = basezoning.merge(zoningmods, on='PARCEL_ID', how='outer')
print('zoningmerge has {} records'.format(zoningmerge.shape[0]))
# create temporary fields to fill na in max_dua_basezoning and dua_up
zoningmerge['dua_up_temp'] = zoningmerge['dua_up'].fillna(0)
zoningmerge['max_dua_basezoning_temp'] = zoningmerge['max_dua_basezoning'].fillna(0)
display(zoningmerge[['PARCEL_ID', 'juris', 'max_dua_basezoning','dua_up']])
# -
# calculate final max_dua used in urbansim
zoningmerge['max_dua_urbansim'] = zoningmerge['max_dua_basezoning_temp']
zoningmerge['max_dua_source'] = 'basezoning'
zoningmerge.loc[zoningmerge.dua_up_temp > zoningmerge.max_dua_basezoning_temp,
'max_dua_source'] = 'zoningmods'
zoningmerge.loc[zoningmerge.dua_up_temp > zoningmerge.max_dua_basezoning_temp,
'max_dua_urbansim'] = zoningmerge['dua_up_temp']
# +
# label parcels with old building and parcels too small to develop
basemap_b10 = pd.read_csv(r'M:\Data\GIS layers\UrbanSim smelt\2020 03 12\b10.csv')
# conver PARCEL_ID to integer:
basemap_b10['parcel_id'] = basemap_b10['parcel_id'].apply(lambda x: int(round(x)))
print("Read {:,} rows from b10".format(basemap_b10.shape[0]))
display(basemap_b10.head())
print('b10 building data has {:,} unique PARCEL_ID'.format(len(basemap_b10.parcel_id.unique())))
basemap_p10 = pd.read_csv(r'M:\Data\GIS layers\UrbanSim smelt\2020 03 12\p10.csv',
usecols =['PARCEL_ID', 'ACRES', 'LAND_VALUE'])
# conver PARCEL_ID to integer:
basemap_p10['PARCEL_ID'] = basemap_p10['PARCEL_ID'].apply(lambda x: int(round(x)))
print("Read {:,} rows from p10".format(basemap_p10.shape[0]))
display(basemap_p10.head())
print('Number of unique PARCEL_ID: {}'.format(len(basemap_p10.PARCEL_ID.unique())))
# +
# join parcels to buildings which is used to determine current built-out condition
building_parcel = pd.merge(left=basemap_b10,
right=basemap_p10[['PARCEL_ID','LAND_VALUE','ACRES']],
left_on='parcel_id',
right_on='PARCEL_ID',
how='outer')
# aggregate buildings at the parcel level
building_groupby_parcel = building_parcel.groupby(['PARCEL_ID']).agg({
'ACRES' :'max',
'LAND_VALUE' :'max',
'improvement_value' :'sum',
'residential_units' :'sum',
'residential_sqft' :'sum',
'non_residential_sqft':'sum',
'building_sqft' :'sum',
'year_built' :'min',
'building_id' :'min'}) # all buildings must be vacant to call this building_vacant
# Identify parcels with old buildings which are protected (if multiple buildings on one parcel, take the oldest)
# and not build on before-1940 parcels
building_groupby_parcel['building_age'] = 'missing'
building_groupby_parcel.loc[building_groupby_parcel.year_built >= 2000, 'building_age' ] = 'after 2000'
building_groupby_parcel.loc[building_groupby_parcel.year_built < 2000, 'building_age' ] = '1980-2000'
building_groupby_parcel.loc[building_groupby_parcel.year_built < 1980, 'building_age' ] = '1940-1980'
building_groupby_parcel.loc[building_groupby_parcel.year_built < 1940, 'building_age' ] = 'before 1940'
building_groupby_parcel['has_old_building'] = False
building_groupby_parcel.loc[building_groupby_parcel.building_age == 'before 1940','has_old_building'] = True
print('Parcel statistics by the age of the oldest building: \n {}'.format(building_groupby_parcel.building_age.value_counts()))
# Identify single-family parcels smaller than 0.5 acre
building_groupby_parcel['small_HS_parcel'] = False
small_HS_idx = (building_groupby_parcel.residential_units == 1.0) & (building_groupby_parcel.ACRES < 0.5)
building_groupby_parcel.loc[small_HS_idx, 'small_HS_parcel'] = True
print("Small single-family parcel statistics: \n {}".format(building_groupby_parcel.small_HS_parcel.value_counts()))
# Identify parcels smaller than 2000 sqft
building_groupby_parcel['small_parcel'] = False
small_parcel_idx = (building_groupby_parcel.ACRES * 43560.0) < 2000
building_groupby_parcel.loc[small_parcel_idx, 'small_parcel'] = True
print("Small parcel (<2000 sqft) statistics: \n {}".format(building_groupby_parcel.small_parcel.value_counts()))
# +
# merge the needed fields with zoningmerge_dua
building_groupby_parcel.reset_index(inplace=True)
building_groupby_parcel['PARCEL_ID'] = building_groupby_parcel['PARCEL_ID'].apply(lambda x: int(round(x)))
parcel_type = building_groupby_parcel[['PARCEL_ID', 'has_old_building', 'small_HS_parcel','small_parcel']]
print('parcel_type has {} records, with {} unique PARCEL_IDs'.format(parcel_type.shape[0],
len(parcel_type.PARCEL_ID.unique())))
zoningmerge_dua = zoningmerge[['PARCEL_ID', 'juris', 'max_dua_urbansim', 'max_dua_source',
'max_dua_basezoning', 'dua_up', 'nodev']]
zoningmerge_dua = zoningmerge_dua.merge(parcel_type, on='PARCEL_ID', how='outer')
print('zoningmerge_dua has {} records, with {} unique PARCEL_IDs'.format(zoningmerge_dua.shape[0],
len(zoningmerge_dua.PARCEL_ID.unique())))
# +
# combine max_dua_source and 'nodev' information
zoningmerge_dua['max_dua_source_final'] = zoningmerge_dua['max_dua_source']
zoningmerge_dua.loc[zoningmerge_dua.has_old_building == True, 'max_dua_source_final'] = 'old building (pre 1940)'
zoningmerge_dua.loc[(zoningmerge_dua.small_HS_parcel == True) | (zoningmerge_dua.small_parcel == True),
'max_dua_source_final'] = 'small parcel'
zoningmerge_dua.loc[zoningmerge_dua.nodev == 1, 'max_dua_source_final'] = 'nodev'
# +
# export with key needed fields
display(zoningmerge_dua)
zoningmerge_dua.to_csv(r'M:\Data\Urban\BAUS\PBA50\Final_Blueprint\Zoning Modifications\final_zoning_urbansim\zoning_urbansim_max_dua.csv', index=False)
# +
# export San Francisco data
zoningmerge_dua_sanfrancisco = zoningmerge_dua.loc[zoningmerge_dua.juris == 'san_francisco']
print('export {} records for San Francisco'.format(zoningmerge_dua_sanfrancisco.shape[0]))
zoningmerge_dua_sanfrancisco.to_csv(r'M:\Data\Urban\BAUS\PBA50\Final_Blueprint\Zoning Modifications\final_zoning_urbansim\zoning_urbansim_max_dua_sanfrancisco.csv', index=False)
|
policies/plu/final_mix_zoning.ipynb
|
# https://eli5.readthedocs.io/en/latest/_modules/eli5/sklearn/explain_weights.html
# https://eli5.readthedocs.io/en/latest/tutorials/xgboost-titanic.html#explaining-predictions
#
# Also take a look at https://eli5.readthedocs.io/en/latest/blackbox/permutation_importance.html.
# +
import os
import random
import numpy as np
import pandas as pd
import xarray as xr
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('ticks')
from pathlib import Path
from tqdm.autonotebook import tqdm
from sklearn.model_selection import train_test_split # Split data into train and test set
from utils import evaluate_classifier, get_sklearn_df
# Allow more rows to be printed to investigate feature importance
pd.set_option('display.max_rows', 300)
# Automatically prints execution time for the individual cells
# %load_ext autotime
# Automatically reloads functions defined in external files
# %load_ext autoreload
# %autoreload 2
# Set xarray to use html as display_style
xr.set_options(display_style="html")
# Tell matplotlib to plot directly in the notebook
# %matplotlib inline
# The path to the project (so absoute file paths can be used throughout the notebook)
PROJ_PATH = Path.cwd().parent
# Mapping dict
mapping_dict_crop_types = {
'Kartofler, stivelses-': 'Potato',
'Kartofler, lægge- (egen opformering)': 'Potato',
'Kartofler, andre': 'Potato',
'Kartofler, spise-': 'Potato',
'Kartofler, lægge- (certificerede)': 'Potato',
'Vårbyg': 'Spring barley',
'Vinterbyg': 'Winter barley',
'Vårhvede': 'Spring wheat',
'Vinterhvede': 'Winter wheat',
'Vinterrug': 'Winter rye',
'Vårhavre': 'Spring oat',
'Silomajs': 'Maize',
'Vinterraps': 'Rapeseed',
'Permanent græs, normalt udbytte': 'Permanent grass',
'Pil': 'Willow',
'Skovdrift, alm.': 'Forest'
}
# Set seed for random generators
RANDOM_SEED = 42
# Seed the random generators
random.seed(RANDOM_SEED)
np.random.seed(RANDOM_SEED)
os.environ['PYTHONHASHSEED'] = str(RANDOM_SEED)
# -
netcdf_path = (PROJ_PATH / 'data' / 'processed' / 'FieldPolygons2019_stats').with_suffix('.nc')
ds = xr.open_dataset(netcdf_path, engine="h5netcdf")
ds # Remember to close the dataset before the netcdf file can be rewritten in cells above
ds.close()
# Convert the xarray dataset to pandas dataframe
df = ds.to_dataframe()
df = df.reset_index() # Removes MultiIndex
df = df.drop(columns=['cvr', 'gb', 'gbanmeldt', 'journalnr', 'marknr', 'pass_mode', 'relative_orbit'])
df = df.dropna()
# +
df_sklearn = get_sklearn_df(polygons_year=2019,
satellite_dates=slice('2018-01-01', '2019-12-31'),
fields='all',
satellite='S1B',
polarization='all',
crop_type='all',
netcdf_path=netcdf_path)
df_sklearn_remapped = df_sklearn.copy()
df_sklearn_remapped.insert(3, 'Crop type', '')
df_sklearn_remapped.insert(4, 'Label ID', 0)
mapping_dict = {}
class_names = []
i = 0
for key, value in mapping_dict_crop_types.items():
df_sklearn_remapped.loc[df_sklearn_remapped['afgroede'] == key, 'Crop type'] = value
if value not in class_names:
class_names.append(value)
mapping_dict[value] = i
i += 1
for key, value in mapping_dict.items():
df_sklearn_remapped.loc[df_sklearn_remapped['Crop type'] == key, 'Label ID'] = value
#print(f"Crop types: {class_names}")
# Get values as numpy array
array = df_sklearn_remapped.values
# Define the independent variables as features.
X = np.float32(array[:,5:]) # The features
# Define the target (dependent) variable as labels.
y = np.int8(array[:,4]) # The column 'afgkode'
# Create a train/test split using 30% test size.
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=RANDOM_SEED)
# Instantiate and evaluate classifier
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(solver='lbfgs', multi_class='auto', n_jobs=32, max_iter=1000)
clf_trained, _, accuracy_test, results_report = evaluate_classifier(
clf, X_train, X_test, y_train, y_test, class_names, feature_scale=True, plot_conf_matrix=False,
print_classification_report=True)
# +
# Get classfication report as pandas df
df_results = pd.DataFrame(results_report).transpose()
# Round the values to 2 decimals
df_results = df_results.round(2).astype({'support': 'int32'})
# Correct error when creating df (acc. is copied into 'precision')
df_results.loc[df_results.index == 'accuracy', 'precision'] = ''
# Correct error when creating df (acc. is copied into 'recall')
df_results.loc[df_results.index == 'accuracy', 'recall'] = ''
# Correct error when creating df (acc. is copied into 'recall')
df_results.loc[df_results.index == 'accuracy', 'recall'] = ''
# The number of samples ('support') was incorrectly parsed in to dataframe
df_results.loc[df_results.index == 'accuracy', 'support'] = df_results.loc[
df_results.index == 'macro avg', 'support'].values
# Print df in latex format (I normally add a /midrule above accuracy manually)
print(df_results.to_latex(index=True))
# -
df_results.loc[df_results.index == 'accuracy', 'precision'] = ''
df_results.loc[df_results.index == 'accuracy', 'recall'] = ''
df_results.loc[df_results.index == 'accuracy', 'support'] = df_results.loc[
df_results.index == 'macro avg', 'support'].values
df_results
try:
from eli5 import show_weights
except:
# !conda install -y eli5
# +
import eli5
from eli5.sklearn import PermutationImportance
feature_names = df_sklearn.columns[3:]
perm = PermutationImportance(clf, random_state=1).fit(X_test, y_test)
eli5.explain_weights(clf, feature_names=list(feature_names), target_names=class_names)
# Look at https://eli5.readthedocs.io/en/latest/autodocs/formatters.html#eli5.formatters.html.format_as_html
# IMPORTANT: LOOK HERE TO FIND IMPORTANCE FOR INDIVIDUAL CLASSES:
# https://stackoverflow.com/questions/59245580/eli5-explain-weights-does-not-returns-feature-importance-for-each-class-with-skl
# +
df_explanation = eli5.formatters.as_dataframe.explain_weights_df(perm, feature_names=list(feature_names))
df_explanation = df_explanation.sort_values(by=['feature'])
df_explanation['polarization'] = ''
features = df_explanation['feature'].unique()
for feature in features:
if feature[-5:] == 'VV-VH':
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VV-VH'
df_explanation = df_explanation.replace(feature, feature[:-6])
elif feature[-2:] == 'VV':
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VV'
df_explanation = df_explanation.replace(feature, feature[:-3])
else:
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VH'
df_explanation = df_explanation.replace(feature, feature[:-3])
# OLD CODE:
#df_explanation_vh = df_explanation.iloc[::3]
#df_explanation_vh['polarization'] = 'VH'
#df_explanation_vh['feature'] = df_explanation_vh['feature'].map(lambda x: str(x)[:-3])
#df_explanation_vv = df_explanation.iloc[1::3]
#df_explanation_vv['polarization'] = 'VV'
#df_explanation_vv['feature'] = df_explanation_vv['feature'].map(lambda x: str(x)[:-3])
#df_explanation_vvvh = df_explanation.iloc[2::3]
#df_explanation_vvvh['polarization'] = 'VV-VH'
#df_explanation_vvvh['feature'] = df_explanation_vvvh['feature'].map(lambda x: str(x)[:-6])
#df_explanation = pd.concat([df_explanation_vh, df_explanation_vv, df_explanation_vvvh])
# -
df_explanation.head(3)
plt.figure(figsize=(24, 8))
plt.xticks(rotation=90, horizontalalignment='center')
ax = sns.lineplot(x='feature', y='weight', hue='polarization', data=df_explanation, ci='sd')
# +
#df_explanation = eli5.formatters.as_dataframe.explain_prediction_df(perm, feature_names=list(feature_names))
# +
# Show the calculated stds in the df as confidence interval on the plot
# https://stackoverflow.com/questions/58399030/make-a-seaborn-lineplot-with-standard-deviation-confidence-interval-specified-f
#lower_bound = [M_new_vec[i] - Sigma_new_vec[i] for i in range(len(M_new_vec))]
#upper_bound = [M_new_vec[i] + Sigma_new_vec[i] for i in range(len(M_new_vec))]
#plt.fill_between(x_axis, lower_bound, upper_bound, alpha=.3)
# +
df_explanation = eli5.formatters.as_dataframe.explain_weights_df(clf, feature_names=list(feature_names), target_names=class_names)
df_explanation = df_explanation.sort_values(by=['feature', 'target'])
df_explanation['polarization'] = ''
features = df_explanation['feature'].unique()
features = features[:-1] # The last features are the bias values
df_bias_values = df_explanation[df_explanation['feature'] == '<BIAS>']
df_explanation = df_explanation[df_explanation['feature'] != '<BIAS>']
for feature in features:
if feature[-5:] == 'VV-VH':
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VV-VH'
df_explanation = df_explanation.replace(feature, feature[:-6])
elif feature[-2:] == 'VV':
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VV'
df_explanation = df_explanation.replace(feature, feature[:-3])
else:
df_explanation.loc[df_explanation['feature'] == feature, 'polarization'] = 'VH'
df_explanation = df_explanation.replace(feature, feature[:-3])
# -
df_bias_values
data = df_explanation[df_explanation['polarization'] == 'VH']
#data = data.loc[data['target'].isin(['Forest', 'Maize', 'Rapeseed'])]
plt.figure(figsize=(24, 8))
plt.xticks(rotation=90, horizontalalignment='center')
ax = sns.lineplot(x='feature', y='weight', hue='target', data=data, ci='sd')
data = df_explanation[df_explanation['polarization'] == 'VH']
data = data.loc[data['target'].isin(['Barley', 'Wheat', 'Forest', 'Potato'])]
plt.figure(figsize=(24, 8))
plt.xticks(rotation=90, horizontalalignment='center')
ax = sns.lineplot(x='feature', y='weight', hue='target', data=data, ci='sd')
|
notebooks/05_ExplainableClassification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PokeHackathonKedro
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
#Check preprocessing
preprocessed_available_pokemons = catalog.load("preprocessed_available_pokemons")
preprocessed_battle_results = catalog.load("preprocessed_battle_results")
all_pokemons = catalog.load("all_pokemons")
battle_results = catalog.load("battle_results")
# master = create_master_table(preprocessed_shuttles, preprocessed_companies, reviews)
# master.head()
# +
#Check for NaN values
print(battle_results.isnull().values.any())
print(battle_results.isnull().sum().sum())
print(all_pokemons.isnull().values.any())
print(all_pokemons.isnull().sum().sum())
# -
preprocessed_battle_results.head()
preprocessed_available_pokemons.head()
# all_pokemons.head()
pd.set_option("display.max_rows", 151, "display.max_columns", None)
print(all_pokemons)
pd.set_option("display.max_rows", 10, "display.max_columns", None)
# +
# print(preprocessed_available_pokemons)
# +
# Test master table for integrating type1,type2 in battle_results
def create_battle_results_with_types(
all_pokemons: pd.DataFrame, battle_results: pd.DataFrame) -> pd.DataFrame:
"""Integrates Type_1 and Type_2 from 'all_pokemons' into battle_results.
Gets projected onto Type_1_1 and Type_2_1 for Pokemon1 and Type_1_2 and Type_2_2 for Pokemon2.
'NaN' is replaced with '0'
Also integrates IDs for Pokemons => ID_1, ID_2
P1, P2 used for 'Pokemon 1' and 'Pokemon 2'
Args:
all_pokemons: Preprocessed data for All Pokemons.
battle_results: Preprocessed data for Battle Results.
Returns:
battle_results_with_types
"""
#Avoid Nidorana a second time (IDs 29 and 32) in all_pokemons
all_pokemons = all_pokemons[all_pokemons['ID'] != 32]
## Pokemon1
# Merge battle_results with all_pokemons based on the Name of P1
battle_results_with_types_P1 = battle_results.merge(all_pokemons, left_on="Name_1", right_on="Name")
# Rename Type_1 and Type_2 to Type_1_1 and Type_2_1
battle_results_with_types_P1.rename(columns={'Type_1': 'Type_1_1', 'Type_2': 'Type_2_1', 'ID':'ID_1'}, inplace=True)
#Drop 'Name' as it is already saved in 'Name_1'
battle_results_with_types_P1.drop(["Name"], axis=1, inplace=True)
## Adding Pokemon2
# Merge battle_results_with_types_P1 with all_pokemons based on the Name of P2
battle_results_with_types = battle_results_with_types_P1.merge(all_pokemons, left_on="Name_2", right_on="Name")
# Rename Type_1 and Type_2 to Type_1_1 and Type_2_1
battle_results_with_types.rename(columns={'Type_1': 'Type_1_2', 'Type_2': 'Type_2_2', 'ID':'ID_2'}, inplace=True)
#Drop 'Name' as it is already saved in 'Name_2'
battle_results_with_types.drop(["Name"], axis=1, inplace=True)
battle_results_with_types = battle_results_with_types.fillna('')
return battle_results_with_types
# -
battle_results_with_types = create_battle_results_with_types(all_pokemons, preprocessed_battle_results)
# +
print(len(preprocessed_battle_results))
print(len(battle_results_with_types))
print(len(battle_results_with_types)-len(preprocessed_battle_results))
print(len(preprocessed_battle_results['Name_1'].unique().tolist()))
# Here I found out that number of ID != number of names in all_pokemons because of Nidorana appearing twice (genderwise)
print(len(all_pokemons['ID'].unique().tolist()))
print(len(all_pokemons['Name'].unique().tolist()))
# +
#preprocessed_available_pokemons[preprocessed_available_pokemons['Name_1']=='Nidorana']
# +
pd.set_option("display.max_rows", 20, "display.min_rows", 10, "display.max_columns", None)
# battle_results_with_types['Name_1'].unique().tolist()
battle_results_with_types.loc[battle_results_with_types['Name_1'] == 'Nidorana']
# -
len(battle_results.loc[battle_results['Name_1'] == 'Nidorana'])
battle_results_with_types[['Name_1','ID_1','Type_1_1', 'Type_2_1','Name_2','ID_2','Type_1_2', 'Type_2_2']].head()
# +
# Test master table for feature hashing type1,type2 in battle_results
from sklearn.feature_extraction import FeatureHasher
def create_battle_results_with_hashed_types(
battle_results: pd.DataFrame) -> pd.DataFrame:
"""Feature hashes 'Type_1' and 'Type_2' and 'WeatherAndTime' from 'battle_results_with_types'.
Example for labeling: Type_1_1 => Type_1_1_a, Type_1_1_b, Type_1_1_c, Type_1_1_d, Type_1_1_e
number of features: 5 (labeled with a,b,c,d,e)
For transparency, original Types/WeatherandTime are not dropped in DataFrame
Args:
battle_results: Preprocessed data for Battle Results with types
Returns:
battle_results_with_hashed_types_weather
"""
##Feature Hasher for Type
h_type = FeatureHasher(n_features=5, input_type='string')
#Transform Types of Pokemon1: Type1_1, Type2_1 and of Pokemon2: Type1_2, Type2_2
dtype1_1 = h_type.fit_transform(battle_results["Type_1_1"])
dtype2_1 = h_type.fit_transform(battle_results["Type_2_1"])
dtype1_2 = h_type.fit_transform(battle_results["Type_1_2"])
dtype2_2 = h_type.fit_transform(battle_results["Type_2_2"])
#Transform into Pandas DataFrame and rename labels
dtype1_1 = pd.DataFrame(data=dtype1_1.toarray())
dtype1_1.rename(columns={0: 'Type_1_1_a', 1: 'Type_1_1_b', 2: 'Type_1_1_c', 3: 'Type_1_1_d', 4: 'Type_1_1_e'}, inplace=True)
dtype2_1 = pd.DataFrame(data=dtype2_1.toarray())
dtype2_1.rename(columns={0: 'Type_2_1_a', 1: 'Type_2_1_b', 2: 'Type_2_1_c', 3: 'Type_2_1_d', 4: 'Type_2_1_e'}, inplace=True)
dtype1_2 = pd.DataFrame(data=dtype1_2.toarray())
dtype1_2.rename(columns={0: 'Type_1_2_a', 1: 'Type_1_2_b', 2: 'Type_1_2_c', 3: 'Type_1_2_d', 4: 'Type_1_2_e'}, inplace=True)
dtype2_2 = pd.DataFrame(data=dtype2_2.toarray())
dtype2_2.rename(columns={0: 'Type_2_2_a', 1: 'Type_2_2_b', 2: 'Type_2_2_c', 3: 'Type_2_2_d', 4: 'Type_2_2_e'}, inplace=True)
##Feature Hasher for WeatherandTime
h_wt = FeatureHasher(n_features=5, input_type='string')
dwt = h_wt.fit_transform(battle_results["WeatherAndTime"])
dwt = pd.DataFrame(data=dwt.toarray())
dwt.rename(columns={0: 'WeatherAndTime_a', 1: 'WeatherAndTime_b', 2: 'WeatherAndTime_c', 3: 'WeatherAndTime_d', 4: 'WeatherAndTime_e'}, inplace=True)
#(No) Drop of Type_1_1, ... and WeatherandTime column.
#battle_results = battle_results.drop(columns = ["WeatherAndTime", "Type_1_1", "Type_2_1", "Type_1_2", "Type_2_2"])
#Concatenate battle_results with hashed features
battle_results_hashed = pd.concat([battle_results, dtype1_1, dtype1_2, dtype2_1, dtype2_2, dwt], axis=1)
return battle_results_hashed
# -
battle_results_hashed = create_battle_results_with_hashed_types(battle_results_with_types)
battle_results_hashed[['Name_1','Type_1_2','Type_1_1_a', 'Type_1_1_b', 'Type_1_1_c', 'Type_1_1_d', 'Type_1_1_e']]
battle_results_hashed.keys()
# +
# Create master table with Attack/Defense ratios
def create_battle_results_AD(
battle_results: pd.DataFrame) -> pd.DataFrame:
"""Creates additional features 'Attack1/Defense2', 'Attack2/Defense1'. Same for Special Attack/Special Defense
Args:
battle_results: Preprocessed data for Battle Results
Returns:
battle_results: Battle Results with A/D
"""
battle_results['A1/D2'] = battle_results['Attack_1']/battle_results['Defense_2']
battle_results['A2/D1'] = battle_results['Attack_2']/battle_results['Defense_1']
battle_results['Sp_A1/Sp_D2'] = battle_results['Sp_Atk_1']/battle_results['Sp_Def_2']
battle_results['Sp_A2/Sp_D1'] = battle_results['Sp_Atk_2']/battle_results['Sp_Def_1']
return battle_results
# -
battle_results_AD = create_battle_results_AD(battle_results_hashed)
battle_results_AD.head()
battle_results_AD.loc[battle_results_AD['Type_2_1'] == '0']
weakness_pokemons = catalog.load("weakness_pokemons")
# print(weakness_pokemons)
# weakness_pokemons.loc[weakness_pokemons['Types'] == 'Water']['Fire']
# battle_results_AD.keys()
weakness_pokemons
# +
# merge = pd.merge(battle_results_AD,weakness_pokemons,left_on=['Type_1_1','Type_1_2'],right_on=['Types', weakness_pokemons.columns])
# +
# merge.head()
# +
# pd.set_option("display.max_rows", 10, "display.max_columns", None)
#battle_results_weakness = battle_results_AD.loc[weakness_pokemons['Types'] == battle_results_AD['Type_1_1']].apply()#[battle_results_AD['Type_1_1']]
#m = weakness_pokemons['Types'] == battle_results_AD['Type_1_1']
#weakness_pokemons.loc[weakness_pokemons['Types'] == 'Water']['Fire']
#types = weakness_pokemons['Types']
#for i in range(len(types)):
#weakness_pokemons.loc[weakn
#np.where(weakness_pokemons['Types'] == battle_results_AD['Type_1_1']) #[battle_results_AD['Type_1_2']] for i
#print(next(battle_results_AD['Type_1_1'].iterrows())[1])
# print(types[0])
#(weakness_pokemons['Types'] == 'Water')
# battle_results_AD['Type_1_1'][1]
#weakness_pokemons.where(m)
# battle_results_AD['EFF1_2'] = battle_results_AD['Type_1_1'].map(weakness_pokemons)
# battle_results_AD['Type_1_1']
# weakness_pokemons
weakness_long = weakness_pokemons.melt(id_vars="Types")
weakness_long.columns = ["Type_P1", "Type_P2", "value"]
weakness_long
#Eff_11
battle_results_with_weakness = battle_results_AD.merge(weakness_long, left_on=["Type_1_1","Type_1_2"], right_on=["Type_P1","Type_P2"])
battle_results_with_weakness.rename(columns={'value': 'value_1_1ab'}, inplace=True)
battle_results_with_weakness.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness = battle_results_with_weakness.merge(weakness_long, left_on=["Type_1_2","Type_1_1"], right_on=["Type_P1","Type_P2"])
battle_results_with_weakness.rename(columns={'value': 'value_1_1ba'}, inplace=True)
battle_results_with_weakness.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
# battle_results_with_weakness = battle_results_with_weakness.fillna(1)
battle_results_with_weakness['Eff_11'] = battle_results_with_weakness['value_1_1ab']/battle_results_with_weakness['value_1_1ba']
# +
# battle_results_with_weakness.loc[battle_results_with_weakness['Eff_11'].isnull()
print(battle_results_with_weakness['Type_2_2'].isnull().values.any())
print(battle_results_with_weakness['Type_2_2'].isnull().sum())
print(battle_results_with_weakness.isnull().sum().sum())
# +
#Eff_12
battle_results_with_weakness_2 = battle_results_with_weakness.merge(weakness_long, left_on=["Type_1_1","Type_2_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_2.rename(columns={'value': 'value_1_2ab'}, inplace=True)
battle_results_with_weakness_2.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_2 = battle_results_with_weakness_2.merge(weakness_long, left_on=["Type_2_2","Type_1_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_2.rename(columns={'value': 'value_2_1ba'}, inplace=True)
battle_results_with_weakness_2.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_2 = battle_results_with_weakness_2.fillna(1)
battle_results_with_weakness_2['Eff_12'] = battle_results_with_weakness_2['value_1_2ab']/battle_results_with_weakness_2['value_2_1ba']
# np.where(battle_results_with_weakness_2['value_1_2ab'] == 'NaN', 1)
# +
#df['your column name'].isnull().values.any()
# battle_results_with_weakness_2.loc[battle_results_with_weakness_2['Type_2_2'].isnull()]
print(battle_results_with_weakness_2['Type_2_2'].isnull().values.any())
print(battle_results_with_weakness_2['Type_2_2'].isnull().sum())
print(battle_results_with_weakness_2['Type_1_2'].isnull().values.any())
print(battle_results_with_weakness_2['Name_1'].isnull().sum())
print(battle_results_with_weakness_2.isnull().sum().sum())
# -
battle_results_with_weakness_2.loc[battle_results_with_weakness['Type_2_2'] == '0']
# +
#Eff_21
battle_results_with_weakness_3 = battle_results_with_weakness_2.merge(weakness_long, left_on=["Type_2_1","Type_1_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_3.rename(columns={'value': 'value_2_1ab'}, inplace=True)
battle_results_with_weakness_3.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_3 = battle_results_with_weakness_3.merge(weakness_long, left_on=["Type_1_2","Type_2_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_3.rename(columns={'value': 'value_1_2ba'}, inplace=True)
battle_results_with_weakness_3.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_3 = battle_results_with_weakness_3.fillna(1)
battle_results_with_weakness_3['Eff_21'] = battle_results_with_weakness_3['value_2_1ab']/battle_results_with_weakness_3['value_1_2ba']
# -
battle_results_with_weakness_3.loc[battle_results_with_weakness_2['Type_2_1'] == '0']
# +
# Test whether there are Null values
# print(battle_results_with_weakness_3['Type_2_1'].isnull().values.any())
# print(battle_results_with_weakness_3['Type_2_1'].isnull().sum())
# print(battle_results_with_weakness_3['Type_1_2'].isnull().values.any())
# print(battle_results_with_weakness_3['Name_1'].isnull().sum())
print(battle_results_with_weakness_3.isnull().sum().sum())
# +
#Eff_22
battle_results_with_weakness_4 = battle_results_with_weakness_3.merge(weakness_long, left_on=["Type_2_1","Type_2_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_4.rename(columns={'value': 'value_2_2ab'}, inplace=True)
battle_results_with_weakness_4.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_4 = battle_results_with_weakness_4.merge(weakness_long, left_on=["Type_2_2","Type_2_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_4.rename(columns={'value': 'value_2_2ba'}, inplace=True)
battle_results_with_weakness_4.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_4 = battle_results_with_weakness_4.fillna(1)
battle_results_with_weakness_4['Eff_22'] = battle_results_with_weakness_4['value_2_2ab']/battle_results_with_weakness_4['value_2_2ba']
# -
battle_results_with_weakness_4.loc[(battle_results_with_weakness_4['Type_2_1'] == 'Water') & (battle_results_with_weakness_4['Type_1_2'] == 'Fire')]
print(battle_results_with_weakness_4.isnull().sum().sum())
# +
print(len(battle_results))
print(len(preprocessed_battle_results))
print(len(all_pokemons))
print('With Types: ', len(battle_results_with_types))
print(len(battle_results_AD))
print(len(battle_results_with_weakness))
print(len(battle_results_with_weakness_4))
#Super cool!
# +
battle_results_with_weakness_4.loc[battle_results_with_weakness_4['Name_1']=='Mew']
#Legendary seems to have no effect on Damage calculation (observed from Internet)
# +
# preprocessed_battle_results.loc[preprocessed_battle_results['Name_1']=='Mew']
# +
# battle_results_with_weakness_2.loc[battle_results_with_weakness_2['Type_2_1'] == '0']
# -
print(weakness_long)
print(18**2)
def create_battle_results_with_weakness(
battle_results: pd.DataFrame, weakness_pokemons: pd.DataFrame) -> pd.DataFrame:
"""Adds addtitional features for 'Effectiveness of Attacks' ('Eff_11', 'Eff_12', 'Eff_21', Eff_22').
These are based on weakness ratio between the Pokemon's types. The numbers refer only to the types
as the Pokemon can be omitted (ratio is 1/Eff) for the other => there is no new information.
Notation: 11 => Type_1_1 : Type_1_2 (this means Type1 of Pokemon1 divided by Type1 of Pokemon2)
When there is no Type 2 => Set Eff_12 (or Eff_21) to 1 as it should be neutral.
Final Effectiveness is Eff = Eff_11 * Eff_12 * Eff_21 * Eff_22
Args:
battle_results: Preprocessed data for Battle Results
weakness_pokemons: Preprocessed data for Weakness Pokemons.
Returns:
battle_results: Battle Results with Effectiveness
"""
#Unpivot pokemon_weakness to Table with Type1, Type2 and value as columns
weakness_long = weakness_pokemons.melt(id_vars="Types")
weakness_long.columns = ["Type_P1", "Type_P2", "value"]
#Eff_11
battle_results_with_weakness = battle_results.merge(weakness_long, left_on=["Type_1_1","Type_1_2"], right_on=["Type_P1","Type_P2"])
battle_results_with_weakness.rename(columns={'value': 'value_1_1ab'}, inplace=True)
battle_results_with_weakness.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness = battle_results_with_weakness.merge(weakness_long, left_on=["Type_1_2","Type_1_1"], right_on=["Type_P1","Type_P2"])
battle_results_with_weakness.rename(columns={'value': 'value_1_1ba'}, inplace=True)
battle_results_with_weakness.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness = battle_results_with_weakness.fillna(1)
battle_results_with_weakness['Eff_11'] = battle_results_with_weakness['value_1_1ab']/battle_results_with_weakness['value_1_1ba']
#Eff_12
battle_results_with_weakness_2 = battle_results_with_weakness.merge(weakness_long, left_on=["Type_1_1","Type_2_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_2.rename(columns={'value': 'value_1_2ab'}, inplace=True)
battle_results_with_weakness_2.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_2 = battle_results_with_weakness_2.merge(weakness_long, left_on=["Type_2_2","Type_1_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_2.rename(columns={'value': 'value_2_1ba'}, inplace=True)
battle_results_with_weakness_2.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_2 = battle_results_with_weakness_2.fillna(1)
battle_results_with_weakness_2['Eff_12'] = battle_results_with_weakness_2['value_1_2ab']/battle_results_with_weakness_2['value_2_1ba']
#Eff_21
battle_results_with_weakness_3 = battle_results_with_weakness_2.merge(weakness_long, left_on=["Type_2_1","Type_1_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_3.rename(columns={'value': 'value_2_1ab'}, inplace=True)
battle_results_with_weakness_3.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_3 = battle_results_with_weakness_3.merge(weakness_long, left_on=["Type_1_2","Type_2_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_3.rename(columns={'value': 'value_1_2ba'}, inplace=True)
battle_results_with_weakness_3.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_3 = battle_results_with_weakness_3.fillna(1)
battle_results_with_weakness_3['Eff_21'] = battle_results_with_weakness_3['value_2_1ab']/battle_results_with_weakness_3['value_1_2ba']
#Eff_22
battle_results_with_weakness_4 = battle_results_with_weakness_3.merge(weakness_long, left_on=["Type_2_1","Type_2_2"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_4.rename(columns={'value': 'value_2_2ab'}, inplace=True)
battle_results_with_weakness_4.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_4 = battle_results_with_weakness_4.merge(weakness_long, left_on=["Type_2_2","Type_2_1"], right_on=["Type_P1","Type_P2"], how='left')
battle_results_with_weakness_4.rename(columns={'value': 'value_2_2ba'}, inplace=True)
battle_results_with_weakness_4.drop(['Type_P1', 'Type_P2'], axis=1, inplace=True)
battle_results_with_weakness_4 = battle_results_with_weakness_4.fillna(1)
battle_results_with_weakness_4['Eff_22'] = battle_results_with_weakness_4['value_2_2ab']/battle_results_with_weakness_4['value_2_2ba']
#Final Effectiveness 'Eff'
battle_results_with_weakness_4['Eff'] = battle_results_with_weakness_4['Eff_11'] * battle_results_with_weakness_4['Eff_12'] * battle_results_with_weakness_4['Eff_21'] * battle_results_with_weakness_4['Eff_22']
return battle_results_with_weakness_4 #master_table
battle_results_with_weakness = create_battle_results_with_weakness(battle_results_AD, weakness_pokemons)
battle_results_with_weakness.head()
battle_results_with_weakness.loc[(battle_results_with_weakness['Type_2_1'] == 'Water') & (battle_results_with_weakness['Type_1_2'] == 'Fire')]
#Test lengths
print(len(battle_results))
print(len(preprocessed_battle_results))
print(len(all_pokemons))
print('With Types: ', len(battle_results_with_types))
print(len(battle_results_AD))
print(len(battle_results_with_weakness))
print(len(battle_results_with_weakness_4))
# +
#Garbage
# weakness_long = weakness_pokemons.melt(id_vars="Types")
# weakness_long.columns = ["Type_1", "Type_2", "value"]
# battle_results_AD.merge(weakness_long, left_on=["Type_1_1","Type_1_2"], right_on=["Type_1","Type_2"])
# battle_results['Eff1_1'] =
# battle_results.drop(['Type_1', 'Type_2'] axis=1, inplace=True)
# battle_results['Eff']
# battle_results.rename(columns={'value': 'Type_1_1')
# submissions = catalog.load("all_pokemons")
|
pokehackathon-kedro/notebooks/Preprocessing_to_master.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:conda]
# language: python
# name: conda-env-conda-py
# ---
# # RNN for Character Level Language Modeling
# ## Dataset pre-processing
#
# ### sample data
# +
import cPickle as pickle
import numpy as np
import argparse
# sys.path.append(os.path.join(os.path.dirname(__file__), '../../build/python'))
from singa import layer
from singa import loss
from singa import device
from singa import tensor
from singa import optimizer
from singa import initializer
from singa.proto import model_pb2
from tqdm import tnrange
# +
class Data(object):
def __init__(self, fpath, batch_size=32, seq_length=100, train_ratio=0.8):
'''Data object for loading a plain text file.
Args:
fpath, path to the text file.
train_ratio, split the text file into train and test sets, where
train_ratio of the characters are in the train set.
'''
self.raw_data = open(fpath, 'r').read() # read text file
chars = list(set(self.raw_data))
self.vocab_size = len(chars)
self.char_to_idx = {ch: i for i, ch in enumerate(chars)}
self.idx_to_char = {i: ch for i, ch in enumerate(chars)}
data = [self.char_to_idx[c] for c in self.raw_data]
# seq_length + 1 for the data + label
nsamples = len(data) / (1 + seq_length)
data = data[0:nsamples * (1 + seq_length)]
data = np.asarray(data, dtype=np.int32)
data = np.reshape(data, (-1, seq_length + 1))
# shuffle all sequences
np.random.shuffle(data)
self.train_dat = data[0:int(data.shape[0]*train_ratio)]
self.num_train_batch = self.train_dat.shape[0] / batch_size
self.val_dat = data[self.train_dat.shape[0]:]
self.num_test_batch = self.val_dat.shape[0] / batch_size
print 'train dat', self.train_dat.shape
print 'val dat', self.val_dat.shape
def numpy2tensors(npx, npy, dev):
'''batch, seq, dim -- > seq, batch, dim'''
tmpx = np.swapaxes(npx, 0, 1)
tmpy = np.swapaxes(npy, 0, 1)
inputs = []
labels = []
for t in range(tmpx.shape[0]):
x = tensor.from_numpy(tmpx[t])
y = tensor.from_numpy(tmpy[t])
x.to_device(dev)
y.to_device(dev)
inputs.append(x)
labels.append(y)
return inputs, labels
def convert(batch, batch_size, seq_length, vocab_size, dev):
'''convert a batch of data into a sequence of input tensors'''
y = batch[:, 1:]
x1 = batch[:, :seq_length]
x = np.zeros((batch_size, seq_length, vocab_size), dtype=np.float32)
for b in range(batch_size):
for t in range(seq_length):
c = x1[b, t]
x[b, t, c] = 1
return numpy2tensors(x, y, dev)
# -
# ## Create the network
# +
def get_lr(epoch):
return 0.001 / float(1 << (epoch / 50))
data = Data('static/linux_input.txt')
# SGD with L2 gradient normalization
opt = optimizer.RMSProp(constraint=optimizer.L2Constraint(5))
cuda = device.create_cuda_gpu()
rnn = layer.LSTM(name='lstm', hidden_size=32, num_stacks=1, dropout=0.5, input_sample_shape=(data.vocab_size,))
rnn.to_device(cuda)
rnn_w = rnn.param_values()[0]
rnn_w.uniform(-0.08, 0.08)
dense = layer.Dense('dense', data.vocab_size, input_sample_shape=(32,))
dense.to_device(cuda)
dense_w = dense.param_values()[0]
dense_b = dense.param_values()[1]
print 'dense w ', dense_w.shape
print 'dense b ', dense_b.shape
initializer.uniform(dense_w, dense_w.shape[0], 0)
print 'dense weight l1 = %f' % (dense_w.l1())
dense_b.set_value(0)
print 'dense b l1 = %f' % (dense_b.l1())
g_dense_w = tensor.Tensor(dense_w.shape, cuda)
g_dense_b = tensor.Tensor(dense_b.shape, cuda)
# -
# ## Conduct SGD
# +
lossfun = loss.SoftmaxCrossEntropy()
train_loss = 0
for epoch in range(3):
bar = tnrange(data.num_train_batch, desc='Epoch %d' % 0)
for b in bar:
batch = data.train_dat[b * batch_size: (b + 1) * batch_size]
inputs, labels = convert(batch, batch_size, seq_length, data.vocab_size, cuda)
inputs.append(tensor.Tensor())
inputs.append(tensor.Tensor())
outputs = rnn.forward(model_pb2.kTrain, inputs)[0:-2]
grads = []
batch_loss = 0
g_dense_w.set_value(0.0)
g_dense_b.set_value(0.0)
for output, label in zip(outputs, labels):
act = dense.forward(model_pb2.kTrain, output)
lvalue = lossfun.forward(model_pb2.kTrain, act, label)
batch_loss += lvalue.l1()
grad = lossfun.backward()
grad /= batch_size
grad, gwb = dense.backward(model_pb2.kTrain, grad)
grads.append(grad)
g_dense_w += gwb[0]
g_dense_b += gwb[1]
# print output.l1(), act.l1()
bar.set_postfix(train_loss=batch_loss / seq_length)
train_loss += batch_loss
grads.append(tensor.Tensor())
grads.append(tensor.Tensor())
g_rnn_w = rnn.backward(model_pb2.kTrain, grads)[1][0]
dense_w, dense_b = dense.param_values()
opt.apply_with_lr(epoch, get_lr(epoch), g_rnn_w, rnn_w, 'rnnw')
opt.apply_with_lr(epoch, get_lr(epoch), g_dense_w, dense_w, 'dense_w')
opt.apply_with_lr(epoch, get_lr(epoch), g_dense_b, dense_b, 'dense_b')
print '\nEpoch %d, train loss is %f' % (epoch, train_loss / data.num_train_batch / seq_length)
# -
# ## Checkpoint
with open('%s_%d.bin' % (model_path, epoch), 'wb') as fd:
print 'saving model to %s' % model_path
d = {}
for name, w in zip(['rnn_w', 'dense_w', 'dense_b'],[rnn_w, dense_w, dense_b]):
d[name] = tensor.to_numpy(w)
d['idx_to_char'] = data.idx_to_char
d['char_to_idx'] = data.char_to_idx
d['hidden_size'] = hidden_size
d['num_stacks'] = num_stacks
d['dropout'] = dropout
pickle.dump(d, fd)
|
doc/en/docs/notebook/rnn.ipynb
|
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="c294bc50-61c1-45c4-a00c-97fffa09c75b"
/ # Class 6
/ + [markdown] azdata_cell_guid="1517a44c-7b92-4003-be30-98027a475bba"
/ ### Few Quries
/ + [markdown] azdata_cell_guid="62516950-8d7f-4dd1-8c0c-6311f77f3256"
/ #### Select All
/ + azdata_cell_guid="8b096567-e09e-4ed6-8ca4-a0175ab02d40"
Select * from Customers;
/ + [markdown] azdata_cell_guid="1dc48f38-a4de-4cac-ae87-835cb1b213cd"
/ #### In
/ + azdata_cell_guid="f06401bd-53af-48a0-b661-98001fa57288"
Select * from Customers where City in ('Berlin', 'london')
/ + [markdown] azdata_cell_guid="fb5fa410-d445-45e6-b945-5fba0a32d295"
/ #### Between
/ + azdata_cell_guid="b79eabc0-2f67-406a-a56e-ce3a130bc9e6"
Select * from orders where OrderId between 10241 and 10254;
/ + [markdown] azdata_cell_guid="d6541ee5-4291-400c-af51-974efeea2694"
/ #### Like
/ + azdata_cell_guid="0c1584b8-4716-4e51-9475-bb7b865b9bd2"
Select * from Customers where City like 'B%';
-- x% :any number of characters after x
-- x_ :single character after x
-- [ABC]: A or B or C
-- ^:Negation
|
DB-CSC371/class6.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# $$\newcommand{\ket}[1]{\left|{#1}\right\rangle}$$
# $$\newcommand{\bra}[1]{\left\langle{#1}\right|}$$
#
# ## Tehtävä 1: Bellin tilat
#
# ### Johdanto
# Tehtävässä 1 tarkastellaan yksinkertaista kvanttipiiriä, jota käytetään Bellin parin luomiseen. Tavoitteena on myös tutustua syvemmin ket-notaation käyttöön kvanttitilan kuvauksessa.
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros4/bell_state00.png" width="200">
#
# Kuvan piirissä molemmat kubitit ovat alussa tilassa $\ket 0$. Tämän jälkeen 1. kubittiin operoidaan H-portilla, ja kubitit 1 ja 2 lomitetaan CNOT-portilla. 1. kubitti on tällöin ohjaava kubitti, ja 2. kobitti on kohde.
#
# Kuvataan ensin kahden kubitin kvanttipiirin tila $H$-portin jälkeen. 1. kubitti on tilojen superpositiossa ja 2. kubitti on edelleen tilassa $\ket 0$:
#
# $$
# \frac{1}{\sqrt 2} (\ket 0 + \ket 1) \otimes \ket 0
# $$
#
# Laskemalla tilojen tensoritulon saadaan kahden kubitin systeemin tila esitettyä muodossa
#
#
# $$
# \frac{1}{\sqrt 2} (\ket 0 \ket 0 + \ket 1 \ket 0) = \frac{1}{\sqrt 2}(\ket{00}+\ket{10}).
# $$
#
# Seuraavaksi esitetään, miten CNOT-portti vaikuttaa kahden kubitin systeemissä. Jos ohjaavan ensimmäisen kubitin tila on 1, niin kohdekubitin eli jälkimmäisen kubitin tila kääntyy:
#
# $$
# \frac{1}{\sqrt 2}(\ket{00}+\ket{10}) \xrightarrow{\text{CNOT}}\frac{1}{\sqrt 2}(\ket{00}+\ket{11})
# $$
#
# Tätä lopputilaa kutsutaan Bellin $\ket {\phi^+}$ tilaksi. Eli Bellin $\ket {\phi^+}$ tila määritellään seuraavasti:
#
# $$\ket {\phi^+}=\frac{1}{\sqrt 2}(\ket{00}+\ket{11}$$
#
# ### Tehtävät:
# A) Tutki saman H ja CNOT-porttien avulla toteutetun kvanttipiirin toiminta ket-notaation avulla alla olevan kuvan tilanteessa, jossa 1. kubitin alkutila on $\ket 1$ ja 2. kubitin alkutila on $\ket 0$.
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros4/bell_state10.png" width="200">
#
# Ohje: Muista 2. kierroksen tehtävien tulos, kuinka Hadamard-porttilla operointi vaikuttaa tilaan $\ket 1$, eli $H\ket 1 = \frac{1}{\sqrt 2} (\ket 0 - \ket 1)$.
#
#
# B) Laske piirin toiminta myös kubittien alkuarvoilla $\ket 0$ ja $\ket 1$ sekä $\ket 1$ ja $\ket 1$.
#
#
# C) Alla olevassa koodiesimerkissä luodaan tehtävän kvanttipiiri kubittien alkuarvoilla $\ket 0$ ja $\ket 0$ sekä tulostetaan piirin tila H ja CNOT -porttien jälkeen. Pythonissa piirin lopputilaa voidaan tutkia käyttämällä `'statevector_simulator'`-ajuria ja komentoa `result.get_statevector(circuit)`. Piiriin ei siis suoriteta mittausta, eli kahden kubitin systeemi ei romahda mihinkään tiettyyn lopputilaan ja voimme tarkastella superposition tarkkoja arvoja.
#
# Tutki `get_statevector(ciruit)`-metodin tulostetta eri alkuarvoilla 10, 01, ja 11. Kubitin tilan saa käännettyä NOT eli X-portilla, esimerkiksi `circuit.x(qr[0])`.
#
# +
import numpy as np
from qiskit import QuantumCircuit, QuantumRegister, ClassicalRegister
from qiskit import Aer, execute
from qiskit.tools.visualization import plot_state_city
from qiskit.providers.aer import StatevectorSimulator
# Määritellään kahden kubitin kvanttipiiri ilman mittausta
qr = QuantumRegister(2, 'qr')
circuit = QuantumCircuit(qr)
#circuit.x(qr[0])
#circuit.x(qr[1])
# H-portti operoi ensimmäiseen kubittiin
circuit.h(qr[0])
# CNOT-portti, jossa taulukon ensimmäinen kubitti qr[0] toimii ohjaavana
circuit.cx(qr[0], qr[1])
# Valitaan simulaattorityyppi, joka laskee lopullisen kvanttitilan numeerisesti
# eikä tee pelkästään yksittäisiä mittausottoja.
simulator = Aer.get_backend('statevector_simulator')
# Suoritetaan
result = execute(circuit, simulator).result()
statevector = result.get_statevector(circuit)
# Tulostetaan saatu tila ket-notaation kera
for i, c in enumerate(statevector):
print(c, f"|{i:02b}>")
# -
# Koodin tulosteessa merkintä `0.707...+0j` tarkoittaa, että kompleksiluvun reaaliosa on 0.707 ja `j` on imaginääriyksikkö. Huomaa, että $1 / \sqrt 2 \approx 0.707$.
# +
# Piirretään luotu piiri.
circuit.draw(output="mpl")
# -
# **LISÄTIETOJA TEHTÄVÄN 1 RATKAISUISTA:**
#
#
# Tehtävän A- ja B-kohdan ratkaisut ovat ns. Bellin tiloja, jotka määritellään seuraavasti:
#
#
# **A:** Kun alkuarvot ovat $\ket 1$ ja $\ket 0$, niin saadaan $\ket {\phi^-}=\frac{1}{\sqrt 2}(\ket{00}-\ket{11}$
#
# **B:**
#
# Kun alkuarvot ovat $\ket 0$ ja $\ket 1$, niin saadaan $\ket {\psi^+}=\frac{1}{\sqrt 2}(\ket{01}+\ket{10}$
#
# Kun alkuarvot ovat $\ket 1$ ja $\ket 1$, niin saadaan $\ket {\psi^-}=\frac{1}{\sqrt 2}(\ket{01}-\ket{10}$
#
#
# Vektorit $\ket {\phi^+}$ ja $\ket {\phi^-}$ sekä $\ket {\psi^+}$ ja $\ket {\psi^-}$ ovat ns. ortonormaaleja Bellin kantavektoreita, joita voidaan käyttää kvanttipiirin tilan kuvauksessa. Kun Bellin kantavektorit laskettiin kahden kubitin alkutiloista suoritetiin ns. kannanvaihto. H- ja CNOT-porttien avulla toteutettu kannanvaihto on unitaarinen, koska käytetyt portit ovat unitaarisia. Oikeastaan kaikki kannanvaihto-operaattorit ovat unitaarisia.
#
# +
# Kopioi tähän halutessasi yllä oleva koodiesimerkki ja tulosta arvoja eri input-arvoilla.
# Voit tehdä tehtävän myös suoraan yllä olevaan koodilaatikkoon, ja muokata siinä input-arvoja.
# -
# ## Tehtävä 2: Satunnaisluvun arpominen ja kvanttinoppa
#
# Tässä tehtävässä tutkitaan, miten kvanttitietokonetta voisi käyttää satunnaislukujen tuottamiseen.
#
#
# #### Muistutus binäärijärjestelmästä
#
# Esimerkiksi 10-järjestelmän luku 6 voidaan esittää binäärimuodossa $6=1\cdot 2^2+1\cdot2^1+0\cdot 2^0=110_2$.
#
#
#
# Seuraava kvanttipiiri käyttä 5 kubittia, joilla arvotaan 5 klassisesta bitistä koostuva satunnaisluku:
#
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros4/quantum_dice1.png" width="300">
#
#
#
# a) Selitä piirin toimintaperiaate. Miten kuvan kvanttipiirin avulla voidaan määrittää 5 bitistä koostuva satunnaisluku binäärimuodossa?
#
# b) Ohjelmoi kuvan kvanttipiiri. Jos osaat, kokeile myös simuloida piirin toimintaa.
#
# c) **Kvanttinoppa:**
#
# Aiemmalla esimerkillä saadaan vain noppia, joilla on jonkin kahden potenssin ($2^n$) verran eri silmälukuja. Seuraavassa yritämme luoda perinteisen 6-sivuisen nopan.
#
# Alla oleva piiri esittää kvanttinoppaa, joka arpoo yhden kuudesta luvusta: 0, 1, 3, 4, 5 tai 7.
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros4/quantum_dice2.jpeg" width="300">
#
# Tavoitteena on, että jokaisella loputuloksella olisi sama todennäköisyys. Ennen mittausta piirin luomaa kolmen kubitin tilaa voidaan esittää tilavektorilla
#
# $$
# \ket \psi =\frac1{\sqrt6}=(\ket {000}+\ket{001}+\ket{011}+\ket{100}+\ket{101}+\ket{111}).
# $$
#
# Kolmen kubitin systeemi on superpositiossa, ja kun systeemin tila mitataan, on jokaisella mahdollisella lopputilalla sama esiintymistodennäköisyys,
#
# $$P=\bra \psi \ket \psi =\frac 1 6$$.
#
# Kun tila mitataan, tallettuu tieto saadusta lopputuloksesta klassiseen rekisteriin. Esimerkiksi binääriluku $011_2= 0\cdot2^2+1\cdot2^1+1\cdot2^0=3_{10}$ on 10-järjestelmän luku 3.
#
# Piirissä on kaksi uutta porttia, ohjattu (controlled) Hadamard ja kubitin tilan kierto y-akselin suhteen Blochin pallolla. Kiertoportilla $R_y$ luodaan tila $\frac 1 { \sqrt{3}} (\ket 0 + 2\ket 1) $ ja ohjatulla Hadamard-portilla jaetaan lausekkeen jälkimmäinen termi kahteen osaan, jolloin saadaan $1{\sqrt3}(\ket {00}+\ket{01}+\ket{11})$. Kiertoportin parametri eli kääntökulma valitaan siten, että lopputulos painottuu oikein. (muista järjestys: ensimmäinen kubitti bittijonossa viimeisenä) Hadamard-muunnos tehdään siis vain niille tilan osille, joissa ensimmäinen kubitti on $\ket 1$.
#
#
#
# Seuraavassa piirin simuloinnista saadut tulokset:
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros4/dice_sim.jpeg" width="300">
#
#
#
#
# Alla on kvanttipiirin määrittelyssä käytetty koodi. Simuloi piiri. Suorita tämän jälkeen koodi oikella kvanttitietokoneella ja vertaile simuloinnin ja todellisten suoritusten eroja. Miten käytetyt portit vaikuttavat piirin toimintaan?
#
# _Taustatietoa:_ Yhden kubitin kääntöportit (tässä parametrisoitu $R_y$ ja Hadamard) toteutetaan suprajohtavien kubittien kvanttitietokoneessa eri mittaisilla mikroaaltopulsseilla. Pulssin pituus määrää kääntökulman. Voiko tämä kääntö olla tarkkaa?
#
# Monimutkaisia usean kubitin portteja, kuten ohjattua Hadamard-porttia ei ole yleensä suoraan toteutettu kvanttitietokoneessa, vaan sen toiminta pitää korvata useammalla alkeellisella portilla.
#
# +
from qiskit import *
import math
qreg = QuantumRegister(3)
creg = ClassicalRegister(3)
circuit = QuantumCircuit(qreg,creg)
circuit.draw(output='mpl')
# -
# Ry-portti ottaa parametrina haluamamme kulman
circuit.ry(2*acos(1/sqrt(3)), qreg[0])
circuit.ch(qreg[0], qreg[1])
circuit.h(qreg[2])
circuit.barrier()
circuit.measure(qreg[0], creg[0])
circuit.measure(qreg[1], creg[1])
circuit.measure(qreg[2],creg[2])
circuit.draw(output='mpl')
# +
# Tehtävä 2 jatkuu: simulointi
simulator = Aer.get_backend('qasm_simulator')
result = execute(circuit, backend=simulator).result()
# Vaihtoehtoisesti kokeile kommentoitua koodia,
# joka kertoo jokaisen nopanheiton tuloksen erikseen:
#execute(circuit, backend=simulator)
#result = execute(circuit, backend=simulator, shots=16, memory=True).result().get_memory()
#print(result)
from qiskit.tools.visualization import plot_histogram
plot_histogram(result.get_counts(circuit))
# -
# Yksi toisto, poista tarvittessa kommentit
simulator = Aer.get_backend('qasm_simulator')
result = execute(circuit, backend=simulator, shots=1).result()
counts = result.get_counts()
print(counts)
# +
IBMQ.load_account()
provider = IBMQ.get_provider('ibm-q')
# Vaihda tähän haluamasi IBM:n kvanttitietokoneen nimi
# (lista löytyy esim. IBM:n nettisivuilta)
qcomp = provider.get_backend('ibmq_essex')
job = execute(circuit, backend=qcomp)
from qiskit.tools.monitor import job_monitor
job_monitor(job) # Koodi jää odottamaan tähän, kun lähetys on jonossa ja käsiteltävänä
result = job.result()
plot_histogram(result.get_counts(circuit))
# -
# Tekstivastauskenttä. Akivoi kenttä klikkaamalla. Markdown-kentän voi suorittaa klikkaamalla yläpaneelista run.
#
#
#
# ## Tehtävä 3
#
# Tehtävässä 3 rakennetaan tutoriaalin avulla kvanttipiiri, joka ratkaisee binaarimuotoisen salasanan yhdellä yrityksellä. Salasana koostuu vain 0:sta ja 1:stä, salasanan pituus voi vaihdella. Annettuna on funktio eli musta laatikko, joka käyttäytyy sovitulla tavalla salasanan perusteella. Funktio käyttäytyy siten, että se vertailee salasanan jokaista bittiä ja syötteen jokaista bittiä. Funktio laskee AND-operaation salasanan $s$ ja syötteen $x$ jokaisen bitin välillä.
#
# $$
# \begin{array}{r|cccccc}
# s & 0 & 1 & 1 & 0 & 1 & 0 \\
# x & 0 & 0 & 1 & 1 & 1 & 0 \\
# \hline
# \mathit{AND} &0&0&1&0&1&0
# \end{array}
# $$
#
# Funktio palauttaa 0 jos tuloksena oli parillinen määrä ykkösbittejä, muuten funktio palauttaa 1. Toisin sanoen, funktio kertoo, kuinka monta kertaa salasanassa ja syötteessä on 1 bitti samassa kohdassa ja palauttaa tiedon onko näitä kohtia pariton vai parillinen määrä. Taulukon esimerkissä funktio palauttaisi arvon 0.
#
# Tässä termi "musta laatikko" ei tarkoita, ettemme tietäisi piirin toteutusta, päinvastoin: piirin jokainen portti on oltava tiedossa, jotta se voidaan suorittaa kvanttitietokonella. Tämä herättää kysymyksen, mitä hyötyä salasanan ratkaisemisesta on. Bernsteinin-Vaziranin algoritmi ei olekaan käytännössä kovin hyödyllinen, koska se rajaa mustan laatikon, eli funktion toiminnan todella tarkasti. Monimutkaisemmissa algoritmeissa funktio voidaan kuitenkin helposti määrittää tavalla, jolla emme pysty helposti ennustamaan sen käyttäytymistä, esimerkiksi matemaattisten laskutoimitusten avulla.
#
# ### Klassinen tausta:
# Jos salasana koostuu 6 bitistä, kuten esimerkkiksi 101001, niin klassisella tietkoneella tarvitaan salasanan selvittämiseen 6 kappaletta funktion kutsuja. Jos salasanan bittien määrä $n$ kasvaa, niin klassinen tietokone tarvitsee salasanan selvittämiseen $n$ kappaletta funktion kutsuja. Salasana saadaan selville testaamalla jokaista bittiä erikseen:
#
# $$
# f(100000)=1 \\
# f(010000)=0 \\
# f(001000)=1 \\
# f(000100)=0 \\
# f(000010)=0 \\
# f(000001)=1
# $$
#
# Kvanttitietokoneella salasanan selvittäminen onnistuu siten, että funktio ajaetaan vain kerran. Tätä kutsutaan Bernstein-Vazirani-algoritmiksi.
#
# Jos ehdimme, voimme palata Bernstein-Vaziranin algoritmiin ja pyrkiä näyttämään, miksi se toimii.
#
#
# ### Bernstein-Vazirani-algoritmi
#
# a) Rakenna salasanaa $101101$ vastaava mustan laatikon toteutus ja sitä ympäröivä ratkaisualgoritmi:
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros3/bv_piiri.png" width="360">
#
# Alla esimerkki mustan laatikon $U_f$ toteutuksesta, jota vastaava salasana on $1011$. Huomaa järjestys: salasanan ensimmäinen bitti vastaa syötekubiteista alimmaista, $x_3$. Salasanan $101101$ kanssa järjestyksellä ei ole kuitenkaan merkitystä.
#
# <img src="https://kvanttilaskenta.net/hub/static/content/kierros3/bv_box.png" width="220">
#
# Jos salasanassa on jossain kohdassa ykkösbitti, on mustan laatikon toteutuksessa CNOT-portti, joka kääntää tuloskubitin, jos syötteen vastaava bitti on yksi. Jos tuloskubittia käännetään parillinen määrä kertoja, ei piiri tee (tälle superposition osalle) mitään.
#
# Voit tarvittaessa hyödyntää myös ohjevideota:5
#
# <a href="https://youtu.be/sqJIpHYl7oo?t=241" target="_blank">Bernstein-Vazirani-algorimin tutoriaali</a>
#
# b) Simuloi kvanttipiirin toiminta, salasanan tulisi selvitä yhdellä algoritmin toistokerralla. Ohjeet tutoriaalissa.
#
# c) VAPAAEHTOINEN LISÄTEHTÄVÄ: Tee tutoriaalin loppuosassa oleva yleisempi algoritmi, jolla voit ratkaista $n$:stä bitistä koostuvan salasanan yhdellä yrityksellä. Tarkista piirin toiminta simuloimalla piiri.laskennallisesti
#
#
# OHJE A ja B kohdat:
# Jos sinulla on vaikeuksia kirjoittaa virheetöntä koodia Youtube-videosta, niin voit kopioida vaiheittain koodin tästä tietodostosta:
#
# <a href="https://kvanttilaskenta.net/hub/static/content/Bernstein_Vazirani_algorithm.ipynb" target="_blank">Bernstein_Vazirani_algorithm.ipynb </a>
#
# C-kohta:
# <a href="https://kvanttilaskenta.net/hub/static/content/Bernstein_Vazirani_algorithm_advanced.ipynb" target="_blank">Bernstein_Vazirani_algorithm_advanced.ipynb </a>
#
#
# Yritä kuitenkin koodata itse, näin opit paremmin.
#
# Vastuskenttä tehtävään 3. Voit myös tehdä oman jupyter notebook -tiedoston.
from qiskit import *
# %matplotlib inline
from qiskit.tools.visualization import plot_histogram
# jatka tästä
|
kv_laskuharjoitukset/kierros4/.ipynb_checkpoints/tehtavat_kierros4_v2-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Lab 12. Data Analysis In Python
# ## load data into pandas.dataframe
import pandas
# +
df = pandas.read_excel('s3://hackett-ia241-2021spring/house_price.xls')
df[:10]
# -
# ## 2.1 Unit Price
df['unit price'] = df['price']/df['area']
df[:10]
# ## 2.2 House Type
df['house_type'].value_counts()
# ## 2.3 Average Price/ more than two bathrooms
prc_more_2_bathrooms = df.loc[df['bathroom']>2]['price']
print('avgerage price of house more than two bathrooms is ${}'.format(prc_more_2_bathrooms.mean()))
# ## 2.4 mean/median unit price
print('mean unit price is ${}'.format(df['unit price'].mean()))
print('median unit price is ${}'.format(df['unit price'].median()))
# ## 2.5 avg price per house type
df.groupby('house_type').mean()['price']
# ## 2.6 predict price by house area
from scipy import stats
result = stats.linregress(df['area'],df['price'])
print('slope is {}'.format(result.slope))
print('intercept is {}'.format(result.intercept))
print('r square is {}'.format(result.rvalue*result.rvalue))
print('p value is {}'.format(result.pvalue))
# ## 2.7 predict price of house 2,000 sqft
print('price of a house with {} of sqft is ${}'.format(2000,2000*result.slope+result.intercept))
|
lab12.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
import tensorflow as tf
import numpy as np
import cv2
import matplotlib.pyplot as plt
from tensorflow.python.keras.engine import training
import hypar
import network_16 as net
# + id="NuxE9vJP90um"
# Model saved with Keras model.save()
MODEL_PATH = 'models/arcface_weights.h5'
cosine_threshold = 0.075 #emperically set threshold
def ResNet34():
img_input = tf.keras.layers.Input(shape=(112, 112, 3))
x = tf.keras.layers.ZeroPadding2D(padding=1, name='conv1_pad')(img_input)
x = tf.keras.layers.Conv2D(64, 3, strides=1, use_bias=False, kernel_initializer='glorot_normal', name='conv1_conv')(x)
x = tf.keras.layers.BatchNormalization(axis=3, epsilon=2e-5, momentum=0.9, name='conv1_bn')(x)
x = tf.keras.layers.PReLU(shared_axes=[1, 2], name='conv1_prelu')(x)
x = stack_fn(x)
model = training.Model(img_input, x, name='ResNet34')
return model
def block1(x, filters, kernel_size=3, stride=1, conv_shortcut=True, name=None):
bn_axis = 3
if conv_shortcut:
shortcut = tf.keras.layers.Conv2D(filters, 1, strides=stride, use_bias=False, kernel_initializer='glorot_normal', name=name + '_0_conv')(x)
shortcut = tf.keras.layers.BatchNormalization(axis=bn_axis, epsilon=2e-5, momentum=0.9, name=name + '_0_bn')(shortcut)
else:
shortcut = x
x = tf.keras.layers.BatchNormalization(axis=bn_axis, epsilon=2e-5, momentum=0.9, name=name + '_1_bn')(x)
x = tf.keras.layers.ZeroPadding2D(padding=1, name=name + '_1_pad')(x)
x = tf.keras.layers.Conv2D(filters, 3, strides=1, kernel_initializer='glorot_normal', use_bias=False, name=name + '_1_conv')(x)
x = tf.keras.layers.BatchNormalization(axis=bn_axis, epsilon=2e-5, momentum=0.9, name=name + '_2_bn')(x)
x = tf.keras.layers.PReLU(shared_axes=[1, 2], name=name + '_1_prelu')(x)
x = tf.keras.layers.ZeroPadding2D(padding=1, name=name + '_2_pad')(x)
x = tf.keras.layers.Conv2D(filters, kernel_size, strides=stride, kernel_initializer='glorot_normal', use_bias=False, name=name + '_2_conv')(x)
x = tf.keras.layers.BatchNormalization(axis=bn_axis, epsilon=2e-5, momentum=0.9, name=name + '_3_bn')(x)
x = tf.keras.layers.Add(name=name + '_add')([shortcut, x])
return x
def stack1(x, filters, blocks, stride1=2, name=None):
x = block1(x, filters, stride=stride1, name=name + '_block1')
for i in range(2, blocks + 1):
x = block1(x, filters, conv_shortcut=False, name=name + '_block' + str(i))
return x
def stack_fn(x):
x = stack1(x, 64, 3, name='conv2')
x = stack1(x, 128, 4, name='conv3')
x = stack1(x, 256, 6, name='conv4')
return stack1(x, 512, 3, name='conv5')
def loadModel():
base_model = ResNet34()
inputs = base_model.inputs[0]
arcface_model = base_model.outputs[0]
arcface_model = tf.keras.layers.BatchNormalization(momentum=0.9, epsilon=2e-5)(arcface_model)
arcface_model = tf.keras.layers.Dropout(0.4)(arcface_model)
arcface_model = tf.keras.layers.Flatten()(arcface_model)
arcface_model = tf.keras.layers.Dense(512, activation=None, use_bias=True, kernel_initializer="glorot_normal")(arcface_model)
embedding = tf.keras.layers.BatchNormalization(momentum=0.9, epsilon=2e-5, name="embedding", scale=True)(arcface_model)
model = tf.keras.models.Model(inputs, embedding, name=base_model.name)
model.load_weights(MODEL_PATH)
return model
# -
# Load your trained model
model = loadModel()
print("ArcFace expects ",model.layers[0].input_shape[1:]," inputs")
print("and it represents faces as ", model.layers[-1].output_shape[1:]," dimensional vectors")
# + id="Siy-Ydb8-POL"
def get_distance(emb1,emb2):
"""
emb1 & emb2: are both 512 dimensional vectors from the trained resnet model
get_distance: returns cosine_distance
Check Out "https://github.com/zestyoreo/Arcface/blob/main/get_distance()_test.ipynb" for clarity
"""
a = np.matmul(np.transpose(emb1), emb2)
b = np.sum(np.multiply(emb1, emb1))
c = np.sum(np.multiply(emb2, emb2))
cosine_distance = 1 - (a / (np.sqrt(b) * np.sqrt(c)))
return cosine_distance
def face_verify(img_path,img_path2,model):
face1x = cv2.imread(img_path, cv2.IMREAD_UNCHANGED)
if(face1x.shape[2]==4):
face1 = cv2.cvtColor(face1x, cv2.COLOR_BGRA2RGB)
else:
face1 = cv2.cvtColor(face1x, cv2.COLOR_BGR2RGB)
face2x = cv2.imread(img_path2, cv2.IMREAD_UNCHANGED)
if(face2x.shape[2]==4):
face2 = cv2.cvtColor(face2x, cv2.COLOR_BGRA2RGB)
else:
face2 = cv2.cvtColor(face2x, cv2.COLOR_BGR2RGB)
print('Original Dimensions : ',face1.shape)
plt.imshow(face1)
plt.show()
plt.imshow(face2)
plt.show()
# resize image
x1a = cv2.resize(face1, (112,112), interpolation = cv2.INTER_AREA)
print('Resized Dimensions : ',x1a.shape)
x1a = net.Resnet_preprocess(x1a)
img_pixels1 = np.expand_dims(x1a, axis = 0)
#img_pixels1 /= 255 #normalize input
print(img_pixels1.shape)
x2a = cv2.resize(face2, (112,112), interpolation = cv2.INTER_AREA)
x2a = net.Resnet_preprocess(x2a)
img_pixels2 = np.expand_dims(x2a, axis = 0)
#|img_pixels2 /= 255 #normalize input
# Be careful how your trained model deals with the input
# otherwise, it won't make correct prediction!
x1 = img_pixels1
x2 = img_pixels2
embedding1 = model.predict(x1)
embedding2 = model.predict(x2)
preds = "Different People"
cosine_distance = get_distance(embedding1.T,embedding2.T)
print(cosine_distance)
if cosine_distance<cosine_threshold:
preds = "Same People"
return preds
# + id="P_rIGH2O-aJ5"
# images are stored in the images folder. Change path here to do face verification with different images
img_path1 = 'images/a.jpeg'
img_path2 = 'images/b.jpeg'
# -
pred = face_verify(img_path1,img_path2,model)
print(pred)
|
GNR Submission/Codes/Arcface_Face_Verification.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Section 4: Hands-On Data Analysis Lab
#
# We will practice all that you’ve learned in a hands-on lab. This section features a set of analysis tasks that provide opportunities to apply the material from the previous sections. This lab comes in 2 formats – select the one that is most appropriate:
#
# - [Live Session](../notebooks/4-hands_on_data_analysis_lab.ipynb#Live-Session)
# - [Asynchronous Session](#Asynchronous-Session)
#
# ## Asynchronous Session
#
# The lab tasks for the asynchronous session can be found below. Note that these are different from those in the live session. Sample solutions can be found [here](solutions.ipynb).
#
# ### About the Data
#
# We will be using 2019 flight statistics from the United States Department of Transportation’s Bureau of Transportation Statistics (available [here](https://www.transtats.bts.gov/DL_SelectFields.asp?gnoyr_VQ=FMF&QO_fu146_anzr=Nv4%20Pn44vr45) and in this repository as `data/T100_MARKET_ALL_CARRIER.zip`). It contains 321,409 rows and 41 columns. Note that you don't need to unzip the file to read it in with `pd.read_csv()`.
#
# #### Exercises
#
# ##### 1. Read in the data and convert the column names to lowercase to make them easier to work with.
# ##### 2. What columns are in the data?
# ##### 3. How many distinct carrier names are in the dataset?
# ##### 4. Calculate the totals of the `distance`, `freight`, `mail`, and `passengers` columns for flights from the United Kingdom to the United States.
# ##### 5. Find the top 5 carriers for distance traveled.
# ##### 6. Find the total cargo transported (mail + freight) and distance traveled for the 10 carriers that transported the most cargo.
# ##### 7. Which 10 carriers flew the most passengers out of the United States to another country?
# ##### 8. For each of the carriers found in *#7*, find the most popular destination country outside of the United States.
# ##### 9. For each of the carriers found in *#7*, find the total number of passengers flown on international flights to/from the destinations in *#8* or the United States. Note that this dataset only has data for flights with an origin and/or destination of the United States.
# ##### 10. Between which two cities were the most passengers flown? Make sure to account for both directions.
# ##### 11. Find the top 3 carriers for the pair of cities found in *#10* and calculate the percentage of passengers each accounted for.
# ##### 12. Find the percentage of international travel per country using total passengers on class F flights.
# ##### 13. Using a crosstab, find the percentage of total passengers on class F international flights between US cities and the countries found in *#12* that used the carriers found in *#11*.
# ##### 14. Create a pivot table showing the total passengers transported between cities in the United States and other countries by the carriers identified in *#7*. Select the top 10 US cities and top 10 international countries from the result.
# ##### 15: For the top 15 international countries, find the percentage of class F passengers traveling to/from the top 10 US cities for international travel (e.g. if only cities A, B, and C flew into Aruba, the sum of the Aruba row/column would be 1). Plot the result as a heatmap.
|
asynchronous_lab/asynchronous_lab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn import datasets
diabetes = datasets.load_diabetes()
diabetes
print(diabetes.DESCR)
print(diabetes.feature_names)
X = diabetes.data
Y = diabetes.target
X.shape, Y.shape
from sklearn.model_selection import train_test_split
X_train,X_test,Y_train,Y_test = train_test_split(X,Y,test_size = 0.2)
X_train.shape,Y_train.shape
X_test.shape ,Y_test.shape
# +
#Linear regression Model
# -
from sklearn import linear_model
from sklearn.metrics import mean_squared_error,r2_score
model = linear_model.LinearRegression()
model.fit(X_train,Y_train)
Y_pred = model.predict(X_test)
# +
print('Coefficients:', model.coef_)
print('Intercept:', model.intercept_)
print('Mean squared error (MSE): %.2f'
% mean_squared_error(Y_test, Y_pred))
print('Coefficient of determination (R^2): %.2f'
% r2_score(Y_test, Y_pred))
# -
r2_score(Y_test, Y_pred)
import seaborn as sns
Y_test
# +
import numpy as np
np.array(Y_test)
# -
Y_pred
sns.scatterplot(Y_test, Y_pred)
sns.scatterplot(Y_test, Y_pred, marker="+")
sns.scatterplot(Y_test, Y_pred, alpha=0.5)
# +
# +
# -
|
Assignment 2 linear regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yohanesnuwara/python-bootcamp-for-geoengineers/blob/master/petroweek_notebooks/petroweek2020_unit1.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="H2iewYI6RV9N" colab_type="text"
# # Unit 1. Very Brief Intro to Numpy, Matplotlib, Pandas, and Scipy
# + [markdown] id="E1B31foxRch5" colab_type="text"
# This is our Google Colab notebook. A notebook is where we will write codes, stream and import dataset, run them, and see the results. Your local computer doesn't do the work, but your internet does (because Google Colab is a Cloud IDE).
#
# First of all, we will import our GitHub repository, that later on we can stream and import the data from.
# + id="3CRoRg32f_jR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="76175ff1-c60b-42f1-b58d-7d8de0c56540"
# !git clone 'https://github.com/yohanesnuwara/python-bootcamp-for-geoengineers'
# + [markdown] id="-HxANwbEf1Lz" colab_type="text"
# This notebook gives a very 3x brief introduction to Numpy, Matplotlib, Pandas, and Scipy.
# + [markdown] id="NEJv3uURSZZd" colab_type="text"
# Now, we will import libraries.
# + id="RHe8RkP1F-g3" colab_type="code" colab={}
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import scipy
# + [markdown] id="fJUQJ4oOSm7w" colab_type="text"
# After this we will go through each library (how to use them).
# + [markdown] id="ykQN00dQSurf" colab_type="text"
# # Numpy
# + [markdown] id="5lljabciSyDe" colab_type="text"
# Numpy library is widely used for numerical computations.
#
# The objectives of this section are:
#
# * Handle arrays (1D, 2D, and 3D)
# * List comprehension
# * Data cleansing
# * Element search
# * Read (and write) file
# + [markdown] id="kDc2q8ueTeg2" colab_type="text"
# ## Handle arrays (1D, 2D, and 3D)
# + [markdown] id="0Fz3pnLoYNUY" colab_type="text"
# ### 1D array
# + [markdown] id="eMPXYr7mUGgZ" colab_type="text"
# Ways to create an array are:
# * `np.array`: array consists of several values
# * `np.arange`: array of series from a start value to an end value, with a specified increment
# * `np.linspace`: array of series between two values, with a specified number o elements
# + id="NHi3zM2AUs2w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="bbaaaa2d-4bf1-4440-fed2-5eb1f1e5b13b"
# create array with specified values
array1 = np.array([10, 11, 45, 75, 65, 78, 90, 85, 56, 77])
print(array1)
# create an array, say from 1 to 50, with a specified increment, say 5
array2 = np.arange(1, 50, 5)
print(array2)
# create a 1D numpy array consisting of numbers from 1 to 100, divided uniformly into 100 numbers
array3 = np.linspace(1, 100, 10)
print(array3)
# + [markdown] id="_o7BJPoHVlIY" colab_type="text"
# Print the length of each array
# + id="rPy7jqBEVogC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="752db5c9-1e2d-412b-e6fd-adaa44f1e2b7"
print(len(array1))
print(len(array2))
print(len(array3))
# + [markdown] id="zWkulP6TV9dU" colab_type="text"
# Now, instead of repetitive codes, we can use `for` loop
# + id="drdBu9nxWB0V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="06b142a6-45ec-485d-a2b6-b7aa7c94cc8d"
arrays = np.array([array1, array2, array3])
for i in range(len(arrays)):
print(len(arrays[i]))
# + [markdown] id="9Tt2Vyw1Fnp9" colab_type="text"
# Now, sort the elements in the `array1` in ascending order (smallest to highest number)
# + id="i4Ps0w43FyUC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6965518c-c552-4dc8-f9c5-8a7f08d34f4f"
print('Sorted from smallest to highest:', np.sort(array1))
# + [markdown] id="0ROaB2WIF8nj" colab_type="text"
# To sort with descending order, we can use alternative `[::-1]`
# + id="oY8t5wsyGN7m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8082bb6b-ace9-40b0-df29-34c6c19b7fbc"
print('Sorted from highest to smallest:', array1[::-1])
# + [markdown] id="5ONFKcH2Cz9f" colab_type="text"
# Numpy array is unique because indexing starts from 0, not 1. Try printing the first element of the array.
# + id="7nDEh-YJC-8m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9fe4503a-26a6-48f4-9d60-a8800586bec1"
print('First element:', array1[0])
print('Second element:', array1[1])
# + [markdown] id="7sCrtzecDJLC" colab_type="text"
# Print the very last element $N$ and its preceding element $N-1$
# + id="ro66cMczDIgO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2f0d2527-03a9-49d4-c973-faf7928796d6"
print('Last element N:', array1[-1])
print('Element N-1:', array1[-2])
# + [markdown] id="8kgBFKXcD8K6" colab_type="text"
# We can also print the first three elements of the array
# + id="m6bF6nVTEBWq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fc63ac55-48de-4a74-86ee-0b44dbfb5d25"
print('First five elements:', array1[:3])
# + [markdown] id="omp9B96ZEvyC" colab_type="text"
# Print the 5th until 8th element
# + id="X3qZ-nqAE4-r" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="29194f50-b9d9-48f3-a78b-0ed37330d3c5"
print('Fifth until eighth elements:', array1[5:8])
# + [markdown] id="EBHFgJlAELZl" colab_type="text"
# Print the last four elements of the array
# + id="8mXPhlBDEQNe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8ec4a1a3-dad1-465c-c321-59e90bba9871"
print('Last four elements:', array1[-4:])
# + [markdown] id="c3zo9aKpYQc0" colab_type="text"
# ### 2D array (Matrix)
# + [markdown] id="aBY6YhgjYSX1" colab_type="text"
# In math, we know 2D array as a matrix. We create matrix also using `np.array`
# + [markdown] id="t0hWxcLEYK_j" colab_type="text"
# 
#
# To create a matrix is simple. Imagine you have 3 arrays, each consisting of 3 elements.
#
# Array 1: $[10, 20, 30]$
#
# Array 2: $[50, 70, 90]$
#
# Array 3: $[12, 14, 16]$
#
# Then, you stack them. Now, you will have a $(3 \times 3)$ matrix.
#
# $\begin{bmatrix} 10 & 20 & 30 \\ 50 & 70 & 90 \\ 12 & 14 & 16 \end{bmatrix}$
#
# Likewise, using Numpy to stack these arrays are very simple.
# + [markdown] id="jbOT2woKbNHy" colab_type="text"
# You can build it step-by-step, from the 1st row to the 3rd row
# + id="yjDCoL7iYJqj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="a30af472-ddf3-410c-9061-de66c1a11aae"
# each row as 1D array
first = np.array([10, 20, 30])
second = np.array([50, 70, 90])
third = np.array([12, 14, 16])
# stack them together into 2D array
M = np.array([first,
second,
third])
M
# + [markdown] id="Qr34zOcgnn1e" colab_type="text"
# OR, you can build it directly!
#
# You already know how to make 1D numpy array by `np.array([...])`. There's only one squared bracket. So, for a matrix, which is a 2D numpy array, use `np.array([[...]])` with two squared brackets.
# + id="5AAJxHWYbLEx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="8d6e2e1c-041c-4591-e552-172cb43c2c86"
M = np.array([[10, 20, 30],
[50, 70, 90],
[12, 14, 16]])
M
# + [markdown] id="qHQOKI7Gbd1b" colab_type="text"
# Print the matrix shape
# + id="AecXYAy7bgn5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="0edb163c-fcb3-4335-f967-b09f934a42ef"
M.shape
# + [markdown] id="VJhcVa-caRPc" colab_type="text"
# Remember that Python indexing starts from 0. So, if you want to print the element $M_{1,1}$, pass `M[0,0]`
# + id="vSl0s1pearRX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="765de006-1c17-41f3-dfbe-34ceea7e37e9"
print('Element 1,1:', M[0,0])
# + [markdown] id="jYsSp21tawWZ" colab_type="text"
# Likewise, print element $M_{1,3}$
# + id="j0u5ErXxa8tN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e5e54291-b306-464b-ecb8-a5e799b23c01"
print('Element 1,3:', M[0,2])
# + [markdown] id="MYYXCEakfxCO" colab_type="text"
# Print all elements in row 2, or element $M_{2,n}$
# + id="B8k61Xqdf6hd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="2550af56-3566-4772-e805-fa71fe8819cd"
print('All elements in second row:', M[1,:])
# + [markdown] id="UhOv-RLjgE4-" colab_type="text"
# And print all elements in column 2, or element $M_{n,2}$
# + id="1FL9VsVvgKLm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6e4346b7-99c1-47a3-b48a-6b13c69c9cb4"
print('All elements in second column:', M[:,1])
# + [markdown] id="d_zTU-8LaCJD" colab_type="text"
# ## Data cleansing
# + [markdown] id="Av21pss7fqyf" colab_type="text"
# Create an array that consists of NaN values with `np.nan`
# + id="sZNWheokeoFQ" colab_type="code" colab={}
my_array = np.array([np.nan, 15, np.nan, 20, np.nan, 34, np.nan, np.nan, 67, 30, 10, np.nan, 34, np.nan, 50, 25, np.nan])
# + [markdown] id="k2tlh0URgVNS" colab_type="text"
# Check if there is `NaN` values in an array. Returns `True` if there's any. Unless, it returns `False` (Boolean argument).
# + id="rCfG2YAmgaol" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="66dca012-e627-435c-a945-29e76dbf7b1a"
np.isnan(my_array).any()
# + [markdown] id="RbkTJut-iiwi" colab_type="text"
# Removing `NaN` values from the array (just delete the `NaN` elements)
# + id="TIAQgVhYioGA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="045984cc-64a1-470d-d1b5-639d5b1d9b3c"
nan_remove = my_array[~np.isnan(my_array)]
nan_remove
# + [markdown] id="SxyMcF9Wg0cG" colab_type="text"
# Replacing (imputing) `NaN` values with any number. E.g. 0
# + id="0eBGus-7hDU4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="e1bcb0c3-3b6e-4f18-f291-ca2a04ce3d36"
# replace with zeros
# first make a new array to store the imputed data, name it new_array
new_array = np.array(my_array)
# replace with zeros
new_array[np.isnan(new_array)]=0
new_array
# + [markdown] id="bM23Aux0h-r3" colab_type="text"
# Imputing `NaN` values with the mean of the data
# + id="O2xyvIFQiFK4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="c6004879-2089-4a61-e054-73bfe236eb36"
# calculate mean of the non-NaN values
# we already have nan_remove array
mean = np.mean(nan_remove)
# # first make a new array to store the imputed data, name it new_array
new_array = np.array(my_array)
# replace with the mean values
new_array[np.isnan(new_array)]=mean
new_array
# + [markdown] id="Xz0LHEMtac22" colab_type="text"
# ### Element Search
# + [markdown] id="owZ8sF-QkRdq" colab_type="text"
# Create arbitrary array
# + id="UNCEuLTzjwhZ" colab_type="code" colab={}
my_array = np.array([10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 110, 120, 130, 140, 150])
# + [markdown] id="MTa03iIzka7F" colab_type="text"
# Search if there is any element with value $70$ of the created array. Returns `True` if there's any. Unless, it returns `False` (Boolean argument).
# + id="txRlxZuNkk_K" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e6205a31-8f80-4cbc-fe6d-6379a26003b0"
np.any(my_array == 70)
# + [markdown] id="gUvbC1KjlIah" colab_type="text"
# Check what index of the element with value $70$ is in the array
# + id="6xPlQALMlWDd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="72579abe-d485-4f18-c207-293dd9a41e0a"
np.where(my_array == 70)[0]
# + [markdown] id="e7kFNx_Yl5tM" colab_type="text"
# Check what index of the element with values **less than** $70$
# + id="dZ-iObkql3Wa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9ad48840-42bb-47de-b171-6af8ba2c1b65"
np.where(my_array < 70)[0]
# + [markdown] id="4u7x4gB3m1h-" colab_type="text"
# Data QC: Replace all values that are **less than** $70$ with certain value
# + id="NtJ20TCfmeLK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="fc9600ef-78af-41ed-a482-cb61c9893829"
# first make a new array to store the replaced data, name it new_array
new_array = np.array(my_array)
# replace with value 10
new_array[new_array < 70] = 10
new_array
# + [markdown] id="7mpjnUVxyuaD" colab_type="text"
# ## Read text file
# + [markdown] id="7Ylxg1glyzjT" colab_type="text"
# We use `np.loadtxt` to read a text file.
#
# An example here is given a `sincos.txt` file inside the GitHub repository. The file contains numerical result of sine and cosine function.
# + [markdown] id="mHsoNYa87FC8" colab_type="text"
# First, we specify the file path.
# + id="OYI2CRltywY4" colab_type="code" colab={}
# # copy the path in "Table of Contents" tab in Colab, and paste
filepath = '/content/python-bootcamp-for-geoengineers/data/sincos.txt'
# + [markdown] id="bll3_3u07RX8" colab_type="text"
# Open with `np.loadtxt`
# + id="bk2xlordDpvB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="97fec210-959f-4835-c709-6c9443d1c0c7"
data = np.loadtxt(filepath)
data
# + [markdown] id="t5i31W8W7V_w" colab_type="text"
# Check its shape
# + id="pLo-6EPJ7XnT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e3c28c57-d12f-4d59-a350-a550209f0ad1"
data.shape
# + [markdown] id="hSLvnYSB7Zym" colab_type="text"
# It has 1,000 rows and 3 columns.
# + [markdown] id="S1phkb9QhgHF" colab_type="text"
# # Matplotlib
# + [markdown] id="lyF6MiagEwr1" colab_type="text"
# ### Plot and its accessories
# + [markdown] id="o4hwjVnl7crs" colab_type="text"
# In this session, we will use the data that we have imported using `np.loadtxt` earlier.
#
# It has 3 columns. First column is $x$ values, second column is result of $\sin(x)$, and the third column is result of $\cos(x)$.
# + id="QlKgIWTDiqp6" colab_type="code" colab={}
x = data[:,0]
sinx = data[:,1]
cosx = data[:,2]
# + [markdown] id="I90AFqCQ72CJ" colab_type="text"
# Next we make a plot using Matplotlib Pyplot (or `plt`).
# + id="edS8dnTI708y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 265} outputId="a36e52f1-c79a-40c2-c56e-856201e9ef17"
# plot
plt.plot(x, sinx)
plt.plot(x, cosx)
plt.show()
# + [markdown] id="eR90X42F79hW" colab_type="text"
# In every plot, we need to give plot attributes (title, label, legend) and may change the color of curve. We will modify this now.
# + id="0-MgALfnju2g" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 364} outputId="c31d9858-3158-40cc-d7e6-47bbf273809b"
# resize the plot
plt.figure(figsize=(10,5))
# plot, specify color, linetype, linewidth, and give labels
plt.plot(x, sinx, '.', color='purple', label='y=sin(x)')
plt.plot(x, cosx, '-', color='green', linewidth=3, label='y=cos(x)')
# give title, with size and pad
plt.title('Sine and Cosine Function', size=20, pad=10)
# give labels, with size
plt.xlabel('x', size=15)
plt.ylabel('y', size=15)
# limit the axes
plt.xlim(0, 360)
# show the legends and specify its location in the plot
plt.legend(loc='upper center')
# show the grids
plt.grid()
plt.show()
# + [markdown] id="w25FYkPHk3g0" colab_type="text"
# ### Subplot
# + [markdown] id="vlxUJQyv9G1P" colab_type="text"
# What we'll do now is to create our own data using a sine function.
# + id="-VKjAKuy7YK0" colab_type="code" colab={}
x = np.linspace(-3, 3, 1000)
y1 = np.sin(np.pi * x)
y2 = np.sin(np.pi * x) + (0.3 * (np.sin(3 * np.pi * x)))
y3 = np.sin(np.pi * x) + (0.3 * (np.sin(3 * np.pi * x))) + (0.2 * (np.sin(5 * np.pi * x)))
y4 = np.sin(np.pi * x) + (0.3 * (np.sin(3 * np.pi * x))) + (0.2 * (np.sin(5 * np.pi * x))) + (0.1 * (np.sin(7 * np.pi * x)))
# + [markdown] id="_p7jRMlJ9T3F" colab_type="text"
# Next, plot all of the results, using `subplots` so you will have all plots side by side.
# + id="mx0QFDBD-a6n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 558} outputId="97f07a83-e7ef-4523-dac6-3c90fa4154b9"
plt.figure(figsize=(15,10))
plt.suptitle('Fourier Series', size=20)
plt.subplot(2,2,1)
plt.plot(x, y1, color='black')
plt.title(r'$y=\sin(x)$', size=15, pad=10)
plt.xlabel('x', size=10)
plt.ylabel('y', size=10)
plt.xlim(min(x), max(x))
plt.grid()
plt.subplot(2,2,2)
plt.plot(x, y2, color='red')
plt.title(r'$y=\sin(x)+0.3\sin(3x)$', size=15, pad=10)
plt.xlabel('x', size=10)
plt.ylabel('y', size=10)
plt.xlim(min(x), max(x))
plt.grid()
plt.subplot(2,2,3)
plt.plot(x, y3, color='blue')
plt.title(r'$y=\sin(x)+0.3\sin(3x)+0.2\sin(5x)$', size=15, pad=10)
plt.xlabel('x', size=10)
plt.ylabel('y', size=10)
plt.xlim(min(x), max(x))
plt.grid()
plt.subplot(2,2,4)
plt.plot(x, y4, color='green')
plt.title(r'$y=\sin(x)+0.3\sin(3x)+0.2\sin(5x)+0.1\sin(7x)$', size=15, pad=10)
plt.xlabel('x', size=10)
plt.ylabel('y', size=10)
plt.xlim(min(x), max(x))
plt.grid()
# set distance between subplots
plt.tight_layout(4)
plt.show()
# + [markdown] id="frq0R658YviO" colab_type="text"
# # Pandas
# + [markdown] id="OiEpv7wtgXjF" colab_type="text"
# ### Create a Dataframe
# + id="mB1pWDhVgbWY" colab_type="code" colab={}
company = np.array(['ConocoPhillips', 'Royal Dutch Shell', 'Equinor ASA', 'Sonatrach', 'Petronas'])
country = np.array(['USA', 'Netherlands', 'Norway', 'Algeria', 'Malaysia'])
date = np.array(['2002/08/02', '1907/04/23', '1972/06/14', '1963/12/31', '1974/08/07'])
# + id="5RB0uWRLmG_L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="417659d1-3bb5-4fef-feea-6dc4edfeb5c5"
company_df = pd.DataFrame({'Company': company, 'Country': country, 'Date Founded': date})
company_df
# + [markdown] id="3YzEVp89rhAq" colab_type="text"
# #### Basic elements of Dataframe
# + id="J_0nORpkrjgm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="11d32345-ef7a-411a-a46e-1d9ed996d69b"
company_df['Company']
# + id="RwfzKGlsrp6i" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="240c002f-cb77-4482-8ffe-c61b8401a7ef"
company_df.iloc[2]
# + [markdown] id="LIHmexQIsCgx" colab_type="text"
# #### Convert a column data to Array
# + id="rw3-dItOsG0G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="7f88be3a-eef0-467e-e1d5-cbf1f19e0a36"
company_df['Company'].values
# + [markdown] id="pwJN4bHrrPra" colab_type="text"
# #### Adding new column to the dataframe
# + id="6F0NZS3uniEk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="4a4bed11-07c5-4735-98b6-7c9d60357258"
employee = np.array([11400, 82000, 20000, 120000, 51000])
company_df['Employee'] = employee
company_df
# + [markdown] id="AuiJ-PfcsvrG" colab_type="text"
# #### Display summary statistics
# + id="N3h1_Ygqrzrk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 288} outputId="a153ea1f-c92b-44fd-b6e4-836e24478d4f"
company_df.describe()
# + [markdown] id="3XKMMGlbrWuM" colab_type="text"
# #### Convert to Datetime Format
# + id="i-hENuOHqR6T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="f2db94eb-451e-4b13-e772-46e3c2cb2e69"
company_df['Date Founded']
# + [markdown] id="HOuXhzU5qKJ-" colab_type="text"
# Format check web: https://strftime.org/
# + id="eOXirsDHoZC0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="0238c326-71b3-4097-e075-36ba3c45c2f6"
company_df['Date Founded'] = pd.to_datetime(company_df['Date Founded'], format='%Y/%m/%d')
company_df
# + id="LV6zNmyCqoY5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="759ad0fd-612a-49bb-85c1-425e16330a69"
company_df['Date Founded']
# + [markdown] id="QZZmKS2rvPrT" colab_type="text"
# #### Accessing Dataframe columns and rows
# + [markdown] id="dBE8H5USvmHm" colab_type="text"
# Display the "Country" column
# + id="aa6jFUygvbJX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="6416eaea-3ca0-459c-aa7e-940a08b91554"
company_df['Country']
# + [markdown] id="CJt6aglev5sP" colab_type="text"
# Or alternatively we can search by its column index. Let's display the fourth column
# + id="WHYTYvypwBDp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 121} outputId="ccb54bcf-96b1-48ac-d995-446e8b16c559"
company_df.iloc[:,3]
# + [markdown] id="dozcLGAAvtv6" colab_type="text"
# Display the third row
# + id="oJHBXwEuvxJt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="5b6173d1-799e-4268-ad7f-2a2ae9ea647d"
company_df.iloc[2,:]
# + [markdown] id="VE3ctgu6wO2y" colab_type="text"
# ### Slicing dataframe
# + [markdown] id="et6GshiHwRqv" colab_type="text"
# In any case, we may want to select only portion of the dataframe. For example, we want to get the **first two colums** only.
# + id="pyYFwO2Uwd-y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="cc82493d-ed5b-41fe-a4b7-46966569556d"
company_df.iloc[:, 0:2]
# + [markdown] id="PrYSMrJQwzbj" colab_type="text"
# Also, we may want to get the **first two columns**, omitting the rest.
# + id="sUAZuY-ww6IL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 108} outputId="ff98abe2-1253-4478-97ea-d78cf3132192"
company_df.iloc[0:2, :]
# + [markdown] id="Yqqj-dXulmqr" colab_type="text"
# ## Data Analysis of PetroWeek 2020 Registrants
# + [markdown] id="tuU5aozRzRya" colab_type="text"
# Let us analyze the registrant data of this PetroWeek 2020 Python training. Data is in CSV format. First, we specify the file path.
# + id="b4JibTj4zaIR" colab_type="code" colab={}
filepath = '/content/python-bootcamp-for-geoengineers/data/registrant_data_petroweek2020.csv'
# + [markdown] id="iipNwcyl0XMW" colab_type="text"
# Then open the data using Pandas `read_csv`
# + id="9-0Y9wTL0hE9" colab_type="code" colab={}
registrant = pd.read_csv('/content/python-bootcamp-for-geoengineers/data/registrant_data_petroweek2020.csv',
encoding = "ISO-8859-1")
# + [markdown] id="i7bpmiWE3ZjN" colab_type="text"
# Now you can take the overview of data by viewing its `head` and `tail`.
# + id="v-qgME5K3Y0b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 470} outputId="946e41d1-4fc8-4553-b3ef-18c8747ec839"
registrant.head(10)
# + id="zogg-8V14u7W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 435} outputId="9b674cb5-bd3b-4d80-cd6a-60315bb69fc6"
registrant.tail(10)
# + [markdown] id="l6z-1uLE2H6q" colab_type="text"
# Now as you can see, how MESSED UP our data is. Normally, we don't use the `encoding`. However, this raw data requires `encoding`. Also you can see 2 things:
#
# * The last 3 `Unnamed` columns has all `NaN` values
# * The last 3 rows has all `NaN` values (look its `tail`!)
# + [markdown] id="ehdLXkFL63hQ" colab_type="text"
# ### Data cleansing 1: Delete Unwanted Columns
# + [markdown] id="ERKCup9GxNGE" colab_type="text"
# We want to remove the **last 3 rows** because they contain NaN values. We can use the `.iloc` technique that we have learnt just before.
# + id="XKqUKj5P2dNe" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 349} outputId="91fb58f7-900f-483f-be72-34f2cf66ea47"
registrant_edit = registrant.iloc[:,0:-3]
registrant_edit.tail(10)
# + [markdown] id="7cxrOW6s69iX" colab_type="text"
# ### Data cleansing 2: Delete Unwanted Rows
# + [markdown] id="XaY3HUVtxdfl" colab_type="text"
# Next, we want to remove the **last 3 columns** because they contain NaN values. Also we use `.iloc`
# + id="g4LbU-HN7I0H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 349} outputId="588c4d7b-9688-441b-ee70-9ce7a6401341"
registrant_edit = registrant_edit.iloc[0:-3,:]
registrant_edit.tail(10)
# + [markdown] id="mPfgiBuuEl2B" colab_type="text"
# ### How many registrants in this course ?
# + id="OY2oVe9AEp4L" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 86} outputId="5c295cef-8ca5-494c-c1c6-b5248c6e4c10"
registrant_edit.count()
# + [markdown] id="nZU5KHWsCNfl" colab_type="text"
# ### Data analysis: Visualize pie diagram based on registrants' major
# + [markdown] id="gSbK37LZCSET" colab_type="text"
# Now that our data has been edited, we'd like to visualize the majors of the registrants in this course !
#
# First, do a slicing of the column that contains major, `Major / Batch`
# + id="ukmorbw5EE3R" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="0e245437-8c93-489d-fe55-c3a3fd3c91a0"
major = registrant_edit['Major / Batch']
major
# + [markdown] id="5E9dzmUdGatW" colab_type="text"
# Before continuing, we need to REMOVE all rows that contain `NaN` values, so we can sort well.
# + id="6P7w7_sNGevV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="13f1eb8a-1fd0-4faa-cd93-950392a245dd"
major = major.dropna()
major
# + [markdown] id="Smaszqk7UPpi" colab_type="text"
# Now, do the sorting. Here we'd like to count: **How many participants are from major "X" ???**
#
# We do this using `str.contains`, meaning we find in the "Major" colum, which row has string that contains sub-string. Confused?
#
# For example, major Petroleum Engineering. We could find it by searching a substring such as `Petro`.
#
# Next on, we pass `.count()` to count all rows that contains this sub-string.
# + id="blCHzlB9CXwf" colab_type="code" colab={}
# count all majors (in English)
petroleum_major1 = major[major.str.contains('Petro')].count()
geophysics_major1 = major[major.str.contains('Geoph')].count()
geology_major = major[major.str.contains('Geol')].count()
mech_major = major[major.str.contains('Mech')].count()
electrical_major = major[major.str.contains('Electr')].count()
chemical_major = major[major.str.contains('Chem')].count()
material_major = major[major.str.contains('Material')].count()
metallurgy_major = major[major.str.contains('Metal')].count()
astronomy_major = major[major.str.contains('Astro')].count()
economy_major = major[major.str.contains('Econo')].count()
marine_major = major[major.str.contains('Marine')].count()
# + [markdown] id="hQxs74JgVBtz" colab_type="text"
# Because some participants are from Indonesia, they inputted in the registration form in Indonesian language. So, we apply the sorting too.
# + id="lQOinACFLyKE" colab_type="code" colab={}
# count all majors (in Bahasa)
petroleum_major2 = major[major.str.contains('minyak')].count()
geophysics_major2 = major[major.str.contains('Geof')].count()
# + [markdown] id="DNqS-LgCVRf7" colab_type="text"
# Next, we sum up the major which in English and Bahasa, into one individual sum.
# + id="0d17D7fIMhxB" colab_type="code" colab={}
# summing majors in English and in Bahasa
petroleum_major = np.sum([petroleum_major1, petroleum_major2])
geophysics_major = np.sum([geophysics_major1, geophysics_major2])
# + [markdown] id="k6TrAUwJVZMs" colab_type="text"
# Finally, we create a pie diagram using Matplotlib that we learnt before. Use: `plt.pie`
# + id="X2D98K69OZoC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 683} outputId="2033daf4-3548-457b-b0e8-e6547b5d345f"
major_name = ['Petroleum', 'Geophysics', 'Geology', 'Mechanical', 'Electrical',
'Chemical', 'Material', 'Metallurgy', 'Astronomy', 'Economy', 'Marine']
major_count = [petroleum_major, geophysics_major, geology_major,
mech_major, electrical_major, chemical_major, material_major,
metallurgy_major, astronomy_major, economy_major, marine_major]
explode = [0, 0.1, 0.1, 0.2, 0.2 , 0.2, 0.2, 0.2, 0.2, 0.2, 0.2]
plt.figure(figsize=(20,12))
plt.pie(major_count, labels=major_name, explode=explode, autopct='%1.1f%%')
plt.show()
|
petroweek_notebooks/petroweek2020_unit1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [File I/O](https://docs.python.org/3/tutorial/inputoutput.html#reading-and-writing-files)
# Reading and writing files.
# ## Working with paths
# +
import os
current_file = os.path.realpath('file_io.ipynb')
print('current file: {}'.format(current_file))
# Note: in .py files you can get the path of current file by __file__
current_dir = os.path.dirname(current_file)
print('current directory: {}'.format(current_dir))
# Note: in .py files you can get the dir of current file by os.path.dirname(__file__)
data_dir = os.path.join(os.path.dirname(current_dir), '/')
print('data directory: {}'.format(data_dir))
# -
# ### Checking if path exists
print('exists: {}'.format(os.path.exists(data_dir)))
print('is file: {}'.format(os.path.isfile(data_dir)))
print('is directory: {}'.format(os.path.isdir(data_dir)))
# ## Reading files
# +
file_path = os.path.join(data_dir, 'simple_file.txt')
with open(file_path, 'r') as simple_file:
for line in simple_file:
print(line.strip())
# -
# The [`with`](https://docs.python.org/3/reference/compound_stmts.html#the-with-statement) statement is for obtaining a [context manager](https://docs.python.org/3/reference/datamodel.html#with-statement-context-managers) that will be used as an execution context for the commands inside the `with`. Context managers guarantee that certain operations are done when exiting the context.
#
# In this case, the context manager guarantees that `simple_file.close()` is implicitly called when exiting the context. This is a way to make developers life easier: you don't have to remember to explicitly close the file you openened nor be worried about an exception occuring while the file is open. Unclosed file maybe a source of a resource leak. Thus, prefer using `with open()` structure always with file I/O.
#
# To have an example, the same as above without the `with`.
# +
file_path = os.path.join(data_dir, 'simple_file.txt')
# THIS IS NOT THE PREFERRED WAY
simple_file = open(file_path, 'r')
for line in simple_file:
print(line.strip())
simple_file.close() # This has to be called explicitly
# -
# ## Writing files
# +
new_file_path = os.path.join(data_dir, 'new_file.txt')
with open(new_file_path, 'w') as my_file:
my_file.write('This is my first file that I wrote with Python.')
# -
# Now go and check that there is a new_file.txt in the data directory. After that you can delete the file by:
if os.path.exists(new_file_path): # make sure it's there
os.remove(new_file_path)
|
experimental/notebook-workspace/examples/jupyter/fileio.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ## Figures for 'A review of Bayesian perspectives on sample size derivation for confirmatory trials'
#
# The following notebook contains all code required to reproduce the figures in the manuscript.
# +
suppressPackageStartupMessages(library(tidyverse))
source("R/priors.R")
source("R/functions.R")
# -
# ## Comparison of required sample sizes for various prior choices (Figure 2)
#
# We start by defining a baisc parameters for the situation we look at, like maximal type one error rate etc.
# maximal sample size cut-off
nmax <- 1000
# one sided maximal type one error rate
alpha <- 0.025
# minimal power/expected power/probability of success is 1 - beta
beta <- 0.2
# upper boundary of the null hypothesis for the location parameter
# H0: theta <= theta_null
theta_null <- 0.0
# minimal clinically important difference (MCID)
theta_mcid <- 0.1
# Next, we define the range of prior (hyper)parameters to look at. We consider truncated normal priors (maximal entropy given mean/variance on compact interval).
prior_lo <- -0.3
prior_hi <- 0.7
# range of prior means for location parameter
prior_mean_range <- seq(prior_lo, prior_hi, by = .01)
# range of prior standard deviations for location parameter
prior_sd_range <- seq(.01, 1, by = .01)
# We can then evaluate the respective required sample sizes for the different approaches depending on the prior mean and standard deviation.
# See `R/priors.R` and `R/functions.R` for details.
# +
# adjust plot output size
options(repr.plot.width = 8, repr.plot.height = 3.15)
# create 2d grid of prior mean/sd combinations
expand_grid(
mu = prior_mean_range,
tau = prior_sd_range
) %>%
# compute the required sample sizes
mutate(
prior = map2(mu, tau, ~Normal(.x, .y, lo = prior_lo, hi = prior_hi)),
`quantile 0.5` = map_dbl(prior, ~get_n_quantile(theta_null, ., .5, pwr = 1 - beta, alpha = alpha, upper_n = nmax)),
`quantile 0.9` = map_dbl(prior, ~get_n_quantile(theta_null, ., .9, pwr = 1 - beta, alpha = alpha, upper_n = nmax)),
EP = map_dbl(prior, ~get_n_ep(theta_null, ., pwr = 1 - beta, alpha = alpha, upper_n = nmax)),
PoS = map_dbl(prior, ~get_n_pos(theta_null, ., pwr = 1 - beta, alpha = alpha, upper_n = nmax))
) %>%
pivot_longer(
c(contains('quantile'), EP, PoS),
values_to = 'required sample size',
names_to = 'criterion'
) %>%
filter(
`required sample size` <= nmax, # make sure maximal sample size is respected
!is.na(`required sample size`) # throw out instances where the maximal sample size boundary was hit
) %>%
# plot
ggplot() +
aes(mu, tau, fill = `required sample size`, z = `required sample size`) +
geom_raster() + # geom_raster leads to some pdf viewers interpolating, do not want that!
scale_fill_gradient(limits = c(0, 1000), low = '#FFFFFF', high = '#000000') +
guides(
fill = guide_colorbar("required sample size",
barwidth = grid::unit(15, "lines"),
barheight = grid::unit(.5, "lines")
)
) +
coord_cartesian(expand = FALSE) +
xlab('prior mean') +
ylab('prior standard deviation') +
facet_wrap(~criterion, nrow = 1) +
theme_bw() +
theme(
panel.grid = element_blank(),
panel.spacing = unit(1.25, 'lines'),
legend.position = 'top'
)
# -
# save plot as pdf
ggsave("figures/fig2-required-sample-size-comparison.pdf", width = 8, height = 3.15)
# ## Probability of success as the basis for sample size derivation (Figure 3)
# function to compute all parts of PoS'
PoS_all <- function(prior, n, c, null = 0, mrv = null) {
part1 <- integrate(
function(Delta) pdf(prior, Delta) * power(Delta, n, c), prior$lower, null
)$value
part2 <- integrate(
function(Delta) pdf(prior, Delta) * power(Delta, n, c), null, mrv
)$value
part3 <- integrate(
function(Delta) pdf(prior, Delta) * power(Delta, n, c), mrv, prior$upper
)$value
return(tibble(a = part1, b = part2, c = part3))
}
# We now plot the relative composition of PoS' for varying prior mean and standard deviation.
# +
# adjust plot output size
options(repr.plot.width = 7.5, repr.plot.height = 2.75)
expand_grid(
prior_mean = seq(-0.1, .2, .025),
prior_sd = seq(.025, .15, .025)
) %>%
mutate(
tmp = pmap(
list(prior_mean, prior_sd),
~PoS_all(
Normal(..1, ..2, -.3, .7),
150,
qnorm(1 - .05),
mrv = theta_mcid)
)
) %>%
unnest(tmp) %>%
mutate(
id = row_number(),
total = a + b + c,
A = a/total,
B = b/total,
C = c/total
) %>% {
ggplot(.) +
scatterpie::geom_scatterpie(
aes(x = prior_mean, y = prior_sd, group = id, r = .01),
data = ., cols = c("A", "B", "C"), color = NA
) +
geom_text(
aes(x = prior_mean, y = prior_sd, label = sprintf("%.2f", total)),
size = 2
) +
theme_bw() +
coord_equal() +
scale_x_continuous("prior mean", breaks = seq(-.2, .3, .05)) +
scale_y_continuous("prior standard deviation", breaks = seq(0.05, .25, .05)) +
scale_fill_manual("",
values = c(
A = scales::muted("red", l = 75, c = 60),
B = scales::muted("blue", l = 75, c = 60),
C = scales::muted("green", l = 75, c = 60)
)
) +
theme(
legend.position = "right",
panel.grid = element_blank()
)}
# -
ggsave("figures/fig3-pos-prime-composition.pdf", width = 7.5, height = 2.75)
# ## Distribution of random power under a constraint on expected power (not included)
#
# We now look at three particular situations with the following prior means and standard deviations:
# +
# prior configurations to look at
tbl_poi <- tibble(
mu = c(-.25, .3, .5),
tau = c(.4, .125, .05)
) %>%
mutate(
prior = map2(mu, tau, ~Normal(..1, ..2, prior_lo, prior_hi)),
n = map_dbl(prior, ~get_n_ep(theta_null, ., theta_mcid, 1 - beta, alpha)),
label = sprintf("mean=%.2f, sd=%.2f, n=%i", mu, tau, round(n))
)
tbl_poi %>%
select(label, n)
# +
# adjust plot output size
options(repr.plot.width = 7.5, repr.plot.height = 6)
plt_priors <- tbl_poi %>%
mutate(
tmp = map(prior, ~tibble(
theta = seq(prior_lo - 0.1, prior_hi + 0.1, .01),
`unconditional prior` = pdf(., theta) %>%
{ifelse(theta == theta[which.min(abs(prior_lo - theta))] | (theta == theta[which.min(abs(prior_hi - theta))]), NA_real_, .)},
`conditional prior` = pdf(condition(., lo = theta_mcid), theta) %>%
{ifelse(theta == theta[which.min(abs(theta_mcid - theta))] | (theta == theta[which.min(abs(prior_hi - theta))]), NA_real_, .)}
)
)
) %>%
unnest(tmp) %>%
pivot_longer(c(`conditional prior`, `unconditional prior`)) %>%
ggplot(aes(theta, value, linetype = label)) +
geom_line() +
facet_wrap(~name) +
scale_linetype_discrete("prior") +
scale_x_continuous(expression(theta)) +
scale_y_continuous("PDF") +
theme_bw() +
theme(
legend.position = "top",
panel.grid.minor = element_blank(),
strip.text = element_text(size = 6)
)
plt_power_curves <- full_join(
tbl_poi,
expand_grid(
theta = seq(prior_lo, prior_hi, by = 0.01),
n = tbl_poi$n
),
by = "n"
) %>%
mutate(
power = map2_dbl(theta, n, ~power(..1, ..2, qnorm(1 - alpha)))
) %>%
ggplot() +
aes(theta, power, linetype = label) +
geom_line() +
scale_color_discrete("") +
scale_x_continuous(expression(theta), breaks = c(0, 0.5)) +
scale_y_continuous("probability to reject", breaks = seq(0, 1, .2)) +
scale_linetype("") +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
legend.position = "top"
)
set.seed(42)
tbl_samples <- tbl_poi %>%
mutate(
unconditional = map(prior, ~get_sample(., 1e5)),
conditional = map(prior, ~get_sample(condition(., lo = theta_mcid), 1e5))
) %>%
unnest(c(conditional, unconditional)) %>%
pivot_longer(c(conditional, unconditional), names_to = "type", values_to = "rtheta") %>%
mutate(
power = map2_dbl(rtheta, n, ~power(..1, ..2, qnorm(1 - alpha)))
)
tbl_probs <- tbl_samples %>%
group_by(label, type) %>%
summarise(
tmp = mean(power >= 1 - beta),
power = 1 - beta,
.groups = "drop"
)
plt_power_distribution <- tbl_samples %>%
mutate(
type = ifelse(type == "conditional",
"random power",
"random probability to reject"
)
) %>%
ggplot(aes(power)) +
geom_histogram(aes(y = stat(ndensity)), bins = 25, fill = "darkgray") +
geom_vline(aes(xintercept = 0.8), color = "black") +
geom_text(
aes(
label = sprintf("%.2f", tmp),
),
y = .9,
x = 0.91,
size = 3,
color = "black",
data = tbl_probs %>%
mutate(
type = ifelse(type == "conditional",
"random power",
"random probability to reject"
)
)
) +
scale_y_continuous("", breaks = c(), limits = c(0, 1.1)) +
scale_x_continuous("probability to reject", breaks = seq(0, 1, .2)) +
coord_cartesian(expand = FALSE) +
facet_grid(type ~ label, scales = "free_y") +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
panel.spacing = unit(1.25, "lines"),
strip.text = element_text(size = 6)
)
legend <- cowplot::get_legend(plt_power_curves)
cowplot::plot_grid(
legend,
cowplot::plot_grid(
plt_priors + theme(legend.position = "none"),
plt_power_curves + theme(legend.position = "none"),
rel_widths = c(2, 1.2),
nrow = 1
),
plt_power_distribution,
rel_heights = c(.1, 1, 1.75),
ncol = 1
)
# -
ggsave("figures/fig4-power-distribution-ep-approach.pdf", width = 7.5, height = 6)
# ## Distribution of random power under quantile approach (not included)
#
# We now restrict the analysis to a single prior configuration and explore the sensitivity of the prior-quantile approach towards the choice of $\gamma$ and $\beta$.
prior <- Normal(0.3, 0.2, prior_lo, prior_hi)
# +
# adjust plot output size
options(repr.plot.width = 7.5, repr.plot.height = 6)
# plot conditional and unconditional priors
plt_priors <- tibble(
mu = .3,
tau = .2,
prior = list(Normal(.3, .2, lo = prior_lo, hi = prior_hi)),
label = sprintf("mean=%.2f, sd=%.2f", mu, tau)
) %>%
mutate(
tmp = map(prior, ~tibble(
theta = seq(prior_lo - 0.1, prior_hi + 0.1, .005),
`unconditional prior` = pdf(., theta) %>%
{ifelse(theta == theta[which.min(abs(prior_lo - theta))] | (theta == theta[which.min(abs(prior_hi - theta))]), NA_real_, .)},
`conditional prior` = pdf(condition(., lo = theta_mcid), theta) %>%
{ifelse(theta == theta[which.min(abs(theta_mcid - theta))] | (theta == theta[which.min(abs(prior_hi - theta))]), NA_real_, .)}
)
)
) %>%
unnest(tmp) %>%
pivot_longer(c(`conditional prior`, `unconditional prior`)) %>%
ggplot(aes(theta, value, linetype = label)) +
geom_line(aes(linetype = name)) +
scale_x_continuous(expression(theta)) +
scale_y_continuous("PDF") +
scale_linetype_discrete("") +
theme_bw() +
theme(
legend.position = "top",
panel.grid.minor = element_blank()
)
tbl_gamma <- expand_grid(
gamma = c(.5, .9),
target_power = c(0.7, 0.8)
) %>%
mutate(
n = map2_dbl(
gamma,
target_power,
~get_n_quantile(theta_null, prior, ..1, theta_mcid, ..2, alpha)),
label = sprintf("gamma = %.2f\n1 - beta = %.2f\nn = %i", gamma, target_power, round(n))
)
plt_power_curves <- full_join(
tbl_gamma,
expand_grid(
theta = seq(prior_lo, prior_hi, by = 0.01),
n = tbl_gamma$n
),
by = "n"
) %>%
mutate(
power = map2_dbl(theta, n, ~power(..1, ..2, qnorm(1 - alpha)))
) %>%
ggplot() +
aes(theta, power, linetype = label) +
geom_line() +
scale_linetype_discrete("", guide = guide_legend(nrow = 1)) +
scale_x_continuous(expression(theta), breaks =c(0, 0.5)) +
scale_y_continuous("probability to reject", breaks = seq(0, 1, .2)) +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
legend.position = "top",
legend.text = element_text(size = 5),
legend.key.size = unit(0.9,"line")
)
set.seed(42)
tbl_samples <- tbl_gamma %>%
mutate(
`random probability to reject` = map(gamma, ~get_sample(prior, 1e5)),
`random power` = map(gamma, ~get_sample(condition(prior, lo = theta_mcid), 1e5))
) %>%
unnest(c(`random probability to reject`, `random power`)) %>%
pivot_longer(
c(`random probability to reject`, `random power`),
names_to = "type",
values_to = "rtheta"
) %>%
mutate(
power = map2_dbl(rtheta, n, ~power(..1, ..2, qnorm(1 - alpha)))
)
tbl_probs <- tbl_samples %>%
group_by(label, type) %>%
summarise(
tmp = mean(power >= 1 - beta),
power = 1 - beta,
.groups = "drop"
)
plt_power_distribution <- tbl_samples %>%
ggplot(aes(power)) +
geom_histogram(aes(y = stat(ndensity)), bins = 25, fill = "darkgray") +
geom_vline(aes(xintercept = 0.8), color = "black") +
geom_text(
aes(
label = sprintf("%.2f", tmp),
),
y = .9,
x = 0.91,
size = 3,
color = "black",
data = tbl_probs
) +
scale_y_continuous("", breaks = c(), limits = c(0, 1.1)) +
scale_x_continuous("probability to reject", breaks = seq(0, 1, .2)) +
coord_cartesian(expand = FALSE) +
facet_grid(type ~ label, scales = "free_y") +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
panel.spacing = unit(1.25, "lines"),
strip.text = element_text(size = 6)
)
legend_priors <- cowplot::get_legend(plt_priors)
legend_pwr <- cowplot::get_legend(plt_power_curves)
cowplot::plot_grid(
cowplot::plot_grid(
plt_priors , plt_power_curves,
rel_widths = c(1, 1)
),
plt_power_distribution,
rel_heights = c(1, 1.5),
ncol = 1
)
# -
ggsave("figures/fig5-power-distribution-quantile-approach.pdf", width = 7.5, height = 6)
# ## A clinical trial example / utility matching (Figures 4 + 5)
# change the power function to log-rank test statistic assuming an event rate of 1/3
power <- function(Delta, n, c) 1 - pnorm(c, mean = sqrt(n) * sqrt(1/3/4) * Delta, sd = 1)
# +
prior <- Normal(0.2, 0.2, -log(1.5), -log(0.5))
# minimal clinically important difference (MCID)
theta_mcid <- -log(0.95)
tranformed_prior_pdf <- function(hr) {
pdf(prior, -log(hr))/hr
}
# -
1 - cdf(prior, theta_mcid) # a priori probability of relevant effect
# +
# adjust plot output size
options(repr.plot.width = 10, repr.plot.height = 4)
# plot the prior pdf
plt_prior <- tibble(
`hazard ratio` = seq(0.4, 1.6, .001),
`prior PDF` = tranformed_prior_pdf(`hazard ratio`) %>%
{ifelse((abs(`hazard ratio` - 0.5) < 0.01) | (abs(`hazard ratio` - 1.5) < 0.001), NA_real_, .)}
) %>%
ggplot() +
aes(`hazard ratio`, `prior PDF`) +
geom_vline(xintercept = 1) +
geom_line() +
scale_x_continuous("hazard ratio", breaks = seq(0.4, 1.6, by = 0.2)) +
theme_bw() +
theme(
legend.position = "top"
)
# compute required sample sizes
tbl_samplesizes <- tibble(
"MCID" = get_n(theta_null, theta_mcid, upper_n = 1e6),
"EP" = get_n_ep(theta_null, prior, mrv = theta_mcid, pwr = 1 - beta, alpha = alpha, upper_n = 1e5),
"quantile, 0.9" = get_n_quantile(theta_null, prior, .9, mrv = theta_mcid, pwr = 1 - beta, alpha = alpha, upper_n = 1e5),
"quantile, 0.5" = get_n_quantile(theta_null, prior, .5, mrv = theta_mcid, pwr = 1 - beta, alpha = alpha, upper_n = 1e5)
) %>%
pivot_longer(everything(), names_to = "type", values_to = "n")
# plot power curves
plt_powr_curves <- full_join(
tbl_samplesizes,
expand_grid(
`hazard ratio` = seq(0.6, 1.1, .001),
n = tbl_samplesizes$n
),
by = "n"
) %>%
mutate(
power = map2_dbl(`hazard ratio`, n, ~power(-log(..1), ..2, qnorm(1 - alpha))),
name = sprintf("%s (n = %i)", type, n)
) %>%
ggplot() +
aes(`hazard ratio`, power, color = name) +
geom_line() +
scale_color_discrete("") +
scale_x_continuous("hazard ratio", breaks = seq(0.4, 1.6, by = 0.2)) +
scale_y_continuous("probability to reject", limits = c(0, 1), breaks = seq(0, 1, .1), expand = c(0, 0)) +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
legend.position = "top"
)
n <- 1e5
rtheta <- numeric(n)
cprior <- condition(prior, lo = theta_mcid)
i <- 1
while (i < n) {
sample <- rnorm(1, mean = cprior$mu, sd = cprior$tau)
if (between(sample, cprior$lower, cprior$upper)) {
rtheta[i] <- sample
i <- i + 1
}
}
plt_power_cdf <- full_join(
tbl_samplesizes,
expand_grid(
rtheta = rtheta,
n = tbl_samplesizes$n
),
by = "n"
) %>%
mutate(
rpower = map2_dbl(rtheta, n, ~power(..1, ..2, qnorm(1 - alpha))),
name = sprintf("%s (n = %i)", type, n)
) %>%
select(name, rpower) %>%
group_by(name) %>%
nest() %>%
transmute(
ecdf = map(data, ~tibble(
power = seq(0, 1, .01),
CDF = ecdf(.$rpower)(power)
))
) %>%
unnest(ecdf) %>%
ggplot(aes(power, CDF, color = name)) +
geom_line() +
scale_color_discrete("") +
scale_x_continuous("random power", breaks = seq(0, 1, .1)) +
scale_y_continuous(breaks = seq(0, 1, .1)) +
coord_cartesian(expand = FALSE) +
theme_bw() +
theme(
panel.grid.minor = element_blank(),
legend.position = "top"
)
legend <- cowplot::get_legend(plt_powr_curves)
cowplot::plot_grid(
legend,
cowplot::plot_grid(
plt_prior,
plt_powr_curves + theme(legend.position = "none"),
plt_power_cdf + theme(legend.position = "none"),
nrow = 1,
align = "h",
axis = "bt"
),
rel_heights = c(1, 8),
ncol = 1
)
# -
ggsave("figures/fig6-clinical-trial-example.pdf", width = 10, height = 4)
# probability of success
tbl_samplesizes %>%
mutate(
PoS = map_dbl(n, ~PoS(prior, ., qnorm(1 - alpha), theta_mcid))
)
# Next we plot implied utilities (Figure 7)
# +
utility <- function(n, lambda) {
lambda * PoS(prior, n, qnorm(1 - alpha), theta_mcid) - n
}
get_implied_lambda <- function(expected_power) uniroot(
function(lambda) {
EP(
prior,
round(optimize(function(n) utility(n, lambda), c(10, 1e5), maximum = TRUE)$maximum),
c = qnorm(1 - alpha),
mrv = theta_mcid
) - expected_power
},
c(1, 1e6)
)
tbl_implied_lambda <- tibble(
power = seq(0.6, 0.95, by = 0.01)
) %>%
mutate(
n_ep = map_dbl(
power,
~get_n_ep(theta_null, prior, mrv = theta_mcid, pwr = ., alpha = alpha, upper_n = 1e5)
),
lambda_implied = map_dbl(power, ~get_implied_lambda(.)$root)
)
ggplot(tbl_implied_lambda) +
aes(power, lambda_implied) +
geom_point() +
geom_line() +
scale_x_continuous("expected power", breaks = seq(0.6, 0.95, by = 0.05)) +
scale_y_continuous("implied reward") +
theme_bw() +
theme(
panel.grid.minor = element_blank()
)
# -
ggsave("figures/fig7-matched-reward.pdf", width = 8, height = 2.5)
# What's the implied reward for 80% expected power?
get_implied_lambda(0.8)$root %>% round
round(get_implied_lambda(0.8)$root) * 30000 / 1e6
get_implied_lambda(0.9)$root %>% round
round(get_implied_lambda(0.9)$root) * 30000 / 1e7
# Get utility maximising sample size for $\lambda = 10000$
optimize(function(n) utility(n, 10000), c(10, 1e4), maximum = TRUE)$maximum %>%
round
# ... and the corresponging expected power
EP(prior, 1590, c = qnorm(1 - alpha), mrv = theta_mcid)
|
sample-size-calculation-under-uncertainty.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + nbpresent={"id": "d3ec1b0f-bf7a-4d20-b1c7-68193dcce997"}
# %pylab inline
pylab.rcParams['figure.figsize'] = (16.0, 8.0)
# + [markdown] nbpresent={"id": "a4c18296-bca2-4559-b952-6a8b98f2f79f"}
# # Summarizing the Monte Carlo output
# + [markdown] nbpresent={"id": "4528ecfb-a3bd-45b7-bad2-0e941ccc995d"}
# The result of a Monte Carlo simulation is a set of samples from the probability distribution associated with the measurand
# $$ \{\mathbf{y}^{(k)},\,k=1,\ldots,M\} $$
# + [markdown] nbpresent={"id": "8dd06483-0984-4ef0-b23f-021f910f3c2a"}
# The aim is to derive characteristic information about the measurand from this set:
#
# 1) best estimate
#
# 2) uncertainty associated with the best estimate
#
# 3) intervals/regions with a prescribed coverage probability
# + [markdown] nbpresent={"id": "e9ebfef2-1094-4261-bd04-d89768f43dee"}
# ## Univariate measurand
# + [markdown] nbpresent={"id": "b8ecff65-d340-4f51-ab1f-aad9de54d603"}
# 1) best estimate
# \begin{align}
# y =& mean \{ y^{(k)}, k=1,\ldots,M\} \\
# =& \frac{1}{M} \sum_{k=1}^M y^{(k)}
# \end{align}
# 2) uncertainty associated with the best estimate
# \begin{align}
# u_y =& std\{ y^{(k)}, k=1,\ldots,M\} \\
# =& \frac{1}{M-1} \sum_{k=1}^M (y-y^{(k)})^2
# \end{align}
# + [markdown] nbpresent={"id": "b621afbc-6f09-4e86-9890-29aa858c93dc"}
# ### Exercise 4.1
#
# Draw randomly from the normal distribution with mean $\mu=1.3$ and standard deviation $\sigma=0.4$ and calculate best estimate and its uncertainy using 100, 200, 300, ..., 100 000 draws. Compare estimate and uncertainty with mean and standard deviation of the original distribution for the different number of draws.
# + nbpresent={"id": "d3eafb75-33f4-4196-9215-c7c9bafb276e"}
from scipy.stats import norm
# + [markdown] nbpresent={"id": "ab636e15-c803-42a8-9145-c0193b9dfab2"}
# Intervals with a prescribed coverage probability can be calculated from the Monte Carlo outcome as follows
#
# 1) Sort the Monte Carlo outcome in ascending order
# ``` python
# sort(Y)
# ```
# 2) For propability $P$ calculate the corresponding fraction of Monte Carlo draws $q=int(PM)$
#
# 3) Set lower bound of interval as $r=int(0.5(M-q))$ for a probabilistically symmetrical interval
#
# 4) Calculate upper bound of interval as $r+q$
# + [markdown] nbpresent={"id": "af5f004e-360d-45be-b201-2f17fb61a8c2"}
# ### Exercise 4.2
#
# Draw randomly from the distribution from Exercise 4.1 and calculate the 95% probabilistally symetrical coverage interval from 1000, 10000 and 100000 draws. Compare the result to the exact 95% interval.
# + nbpresent={"id": "a93357c4-9c90-4341-a185-217c0ade693d"}
from scipy.stats import norm
P = 0.95 # sought probability of coverage interval
# + [markdown] nbpresent={"id": "5a05fd6c-76d2-47a4-9c24-58b7f6404d86"}
# ## Multivariate measurand
# + [markdown] nbpresent={"id": "b1c8148b-2bd3-4132-96a3-16a6d316eac6"}
# 1) best estimate
# \begin{align}
# \mathbf{y} =& mean \{ \mathbf{y}^{(k)}, k=1,\ldots,M\} \\
# =& \frac{1}{M} \sum_{k=1}^M \mathbf{y}^{(k)}
# \end{align}
# 2) uncertainty associated with the best estimate
# \begin{align}
# U_\mathbf{y} =& cov\{ \mathbf{y}^{(k)}, k=1,\ldots,M\} \\
# =& \frac{1}{M-1} \sum_{k=1}^M (\mathbf{y}-\mathbf{y}^{(k)})(\mathbf{y}-\mathbf{y}^{(k)})^T
# \end{align}
# + [markdown] nbpresent={"id": "5d6183ef-814d-4ea6-87ae-7dc95e9daf05"}
# ### Exercise 4.3
#
# Draw randomly from the normal distribution with mean
# $$\mathbf{\mu}=\left( \begin{array}{c}
# 0.4 \\ -1.5
# \end{array}\right)
# $$
# and covariance
# $$
# \Sigma=\left(\begin{array}{cc}
# 0.09 & -0.2 \\ -0.2 & 1.44
# \end{array}\right)
# $$
# and calculate best estimate and its uncertainy using 1000, 10000 and 100 000 draws. Compare estimate and uncertainty with mean and covariance of the original distribution for the different number of draws.
# + nbpresent={"id": "a98199d3-752c-42fb-9d7e-b31d69e8da5f"}
from scipy.stats import multivariate_normal
# + [markdown] nbpresent={"id": "df956796-6546-4dff-ad19-2251246d7c37"}
# Regions with a prescribed coverage probability can be calculated from the multivariate Monte Carlo outcome as follows
#
# 1) Calculate the Cholesky decomposition of the sample covariance matrix $U_{\mathbf{y}}=\mathbf{LL}^T$
#
# 2) Transform the Monte Carlo outcomes
# $$ \mathbf{y}_{(k)} = \mathbf{L}^{-1}(\mathbf{y}^{(k)}-\mathbf{y})$$
# and sort according to the distance measure
# $$ d^2_{(k)} = \mathbf{y}_{(k)}^T\mathbf{y}_{(k)} $$
#
# 3) calculate $k_P$ such that a fraction $P$ of all Monte Carlo outcomes satisfies $d_{(k)}<k_P$
#
# This defines the ellipsoidal region $(\mathbf{\eta}-\mathbf{y})^TU_{\mathbf{y}}^{-1}(\mathbf{\eta}-\mathbf{y})<k^2_P$
#
# For a bivariate normal distribution, the factor for a 95% coverage ellipsoidal region is given as the 95% quantile of the $\chi^2$ distribution with 2 degrees of freedom.
# + [markdown] nbpresent={"id": "d249b64c-c403-4a93-9621-d1bc390c5d97"}
# ### Exercise 4.4
#
# Calculate 100 000 random draws from the distribution from Exercise 4.3 and calculate the 95% coverage region. Compare to the true 95% coverage region.
# + nbpresent={"id": "f9bd0acc-9db7-4ef8-869b-6b5c3b8bcbe4"}
def calculate_ellipse(mu, Sigma, kP):
vals, vecs = linalg.eigh(Sigma)
order = vals.argsort()[::-1]
vals = vals[order]
vecs = vecs[:,order]
theta = degrees(np.arctan2(*vecs[:,0][::-1]))
width, height = kP * sqrt(vals)
return width, height, theta
# + nbpresent={"id": "58bc76fa-e48c-44d4-81b0-39c72da4fa5f"}
from scipy.stats import multivariate_normal, chi2
from matplotlib.patches import Ellipse
mu = array([0.4, -1.5])
Sigma = array([[0.09, -0.2],[-0.2, 1.44]])
|
.ipynb_checkpoints/05 Summarizing the Monte Carlo output-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pyspark import SparkContext
from pyspark.sql import SQLContext
sc = SparkContext('local', 'Spark SQL')
spark = SQLContext(sc)
# -
spark
df = spark.read.csv("UTF8_2018_Viagem.csv", header=True, sep=";")
df.printSchema()
df.limit(5).toPandas()
from pyspark.sql import functions as F
from pyspark.sql.types import FloatType
def to_value(v):
try:
return float(v.replace(",","."))
except:
return 0.0
udf_to_value = F.udf(to_value, FloatType())
df_typed = df.withColumn("ValorPassagens", udf_to_value(df["Valor passagens"])) \
.withColumn("ValorDiarias", udf_to_value(df["Valor diárias"])) \
.withColumn("ValorOutros", udf_to_value(df["Valor outros gastos"]))
df_typed.limit(5).toPandas()
df_typed.select("Nome órgão solicitante","ValorPassagens").limit(5).toPandas()
pd = df_typed.groupBy("Nome do órgão superior") \
.agg((F.sum("ValorPassagens") / F.lit(1_000_000)).alias("Total")) \
.orderBy("Total", ascending=False).toPandas()
pd
# %matplotlib inline
pd.plot(kind="barh", x="Nome do órgão superior", figsize=[10, 8], width=0.9)
df_typed.groupBy("Nome do órgão superior","Nome","CPF viajante") \
.agg(((F.sum("ValorPassagens") + F.sum("ValorDiarias") + F.sum("ValorOutros"))/ F.lit(1_000)) \
.alias("ValorTotal"), \
F.count("ValorPassagens").alias("Qtd")) \
.filter("Nome !='Informações protegidas por sigilo'") \
.orderBy("ValorTotal", ascending=False) \
.limit(10) \
.toPandas()
pd_gastador = df_typed.filter((df_typed["Nome"]=="<NAME>") & \
(df_typed["CPF viajante"]=="***.678.252-**")) \
.withColumn("DataPartida", F.to_date(df_typed["Período - Data de início"], format="dd/MM/yyyy")) \
.groupBy(F.month("DataPartida").alias("MesPartida")) \
.agg(((F.sum("ValorPassagens") + F.sum("ValorDiarias") + F.sum("ValorOutros"))/ F.lit(1_000)) \
.alias("ValorTotal"), \
F.count("ValorPassagens").alias("Qtd")) \
.orderBy("MesPartida") \
.toPandas()
pd_gastador
pd_gastador.plot(x="MesPartida", figsize=[16, 8], xticks=range(1, 13))
pd_gastador
|
Viagens Csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: foursight
# language: python
# name: foursight
# ---
# # Testing Foursight Checks
# * This notebook provides a list of parameters you can tweak while you are testing your foursight code locally.
# * You can also use this notebook to run available checks locally for longer then 15 min
# * Autoreload of modules are also enabled, so if you make code changes, you don't need to restart the kernel.
# * Please copy the notebook before editing, all notebooks except this one will be ignored by github.
# NEED TO RUN ONCE
# %load_ext autoreload
import logging, sys, json
logging.disable(sys.maxsize)
# +
# %autoreload 2
import app
# check name
check = 'wfr_cgap_checks/cgapS2_status'
action = 'wfr_cgap_checks/cgapS2_start'
# WHICH ENV YOU WANT TO WORK ON (data, staging, cgapwolf, ...)
env = 'cgap'
# DEV OR PROD BUCKET FOR STORING RESULTS - dev or prod
stage= 'prod'
app.set_stage(stage)
connection = app.init_connection(env)
res = app.run_check_or_action(connection, check, {'primary': True})
result = json.dumps(res, indent=4)
print(result)
uuid = res['kwargs']['uuid']
# +
# %autoreload 2
import app
# if there is and action, you can run it on the check you run above
action_params = {'check_name': check.split('/')[1],'called_by': uuid}
res = app.run_check_or_action(connection, action, action_params)
result = json.dumps(res, indent=4)
print(result)
|
LocalTest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="SB93Ge748VQs"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" id="0sK8X2O9bTlz"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="HEYuO5NFwDK9"
# # 将 tf.summary 用法迁移到 TF 2.0
#
# <table class="tfo-notebook-buttons" align="left">
# <td><a target="_blank" href="https://tensorflow.google.cn/tensorboard/migrate"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png">在 TensorFlow.org 上查看 </a></td>
# <td><a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tensorboard/migrate.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png">在 Google Colab 中运行 </a></td>
# <td><a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tensorboard/migrate.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png">在 GitHub 中查看源代码</a></td>
# </table>
# + [markdown] id="56V5oun18ZdZ"
# > 注:本文档面向已经熟悉 TensorFlow 1.x TensorBoard 并希望将大型 TensorFlow 代码库从 TensorFlow 1.x 迁移至 2.0 的用户。如果您是 TensorBoard 的新用户,另请参阅[入门](get_started.ipynb)文档。如果您使用 `tf.keras`,那么可能无需执行任何操作即可升级到 TensorFlow 2.0。
#
# + id="c50hsFk2MiWs"
import tensorflow as tf
# + [markdown] id="56XvRdPy-ewT"
# TensorFlow 2.0 包含对 `tf.summary` API(用于写入摘要数据以在 TensorBoard 中进行可视化)的重大变更。
# + [markdown] id="V_JOBTVzU5Cx"
# ## 变更
#
# 将 `tf.summary` API 视为两个子 API 非常实用:
#
# - 一组用于记录各个摘要(`summary.scalar()`、`summary.histogram()`、`summary.image()`、`summary.audio()` 和 `summary.text()`)的运算,从您的模型代码内嵌调用。
# - 写入逻辑,用于收集各个摘要并将其写入到特殊格式化的日志文件中(TensorBoard 随后会读取该文件以生成可视化效果)。
# + [markdown] id="9-rVv-EYU8_E"
# ### 在 TF 1.x 中
#
# 上述二者必须手动关联在一起,方法是通过 `Session.run()` 获取摘要运算输出,并调用 `FileWriter.add_summary(output, step)`。`v1.summary.merge_all()` 运算通过使用计算图集合汇总所有摘要运算输出使这个操作更轻松,但是这种方式对 Eager Execution 和控制流的效果仍不尽人意,因此特别不适用于 TF 2.0。
# + [markdown] id="rh8R2g5FWbsQ"
# ### 在 TF 2.X 中
#
# 上述二者紧密集成。现在,单独的 `tf.summary` 运算在执行时可立即写入其数据。在您的模型代码中使用 API 的方式与以往类似,但是现在对 Eager Execution 更加友好,同时也保留了与计算图模式的兼容性。两个子 API 的集成意味着 `summary.FileWriter` 现已成为 TensorFlow 执行上下文的一部分,可直接通过 `tf.summary` 运算访问,因此配置写入器将是主要的差异。
# + [markdown] id="em7GQju5VA0I"
# Eager Execution 的示例用法(TF 2.0 中默认):
# + id="GgFXOtSeVFqP"
writer = tf.summary.create_file_writer("/tmp/mylogs/eager")
with writer.as_default():
for step in range(100):
# other model code would go here
tf.summary.scalar("my_metric", 0.5, step=step)
writer.flush()
# + id="h5fk_NG7QKve"
# ls /tmp/mylogs/eager
# + [markdown] id="FvBBeFxZVLzW"
# tf.function 计算图执行的示例用法:
# + id="kovK0LEEVKjR"
writer = tf.summary.create_file_writer("/tmp/mylogs/tf_function")
@tf.function
def my_func(step):
with writer.as_default():
# other model code would go here
tf.summary.scalar("my_metric", 0.5, step=step)
for step in tf.range(100, dtype=tf.int64):
my_func(step)
writer.flush()
# + id="Qw5nHhRUSM7_"
# ls /tmp/mylogs/tf_function
# + [markdown] id="5SY6eYitUJH_"
# 旧 TF 1.x 计算图执行的示例用法:
#
# + id="OyQgeqZhVRNB"
g = tf.compat.v1.Graph()
with g.as_default():
step = tf.Variable(0, dtype=tf.int64)
step_update = step.assign_add(1)
writer = tf.summary.create_file_writer("/tmp/mylogs/session")
with writer.as_default():
tf.summary.scalar("my_metric", 0.5, step=step)
all_summary_ops = tf.compat.v1.summary.all_v2_summary_ops()
writer_flush = writer.flush()
with tf.compat.v1.Session(graph=g) as sess:
sess.run([writer.init(), step.initializer])
for i in range(100):
sess.run(all_summary_ops)
sess.run(step_update)
sess.run(writer_flush)
# + id="iqKOyawnNQSH"
# ls /tmp/mylogs/session
# + [markdown] id="xEJIh4btVVRb"
# ## 转换您的代码
#
# 将现有的 `tf.summary` 用法转换至 TF 2.0 API 无法实现可靠的自动化,因此需要通过 [`tf_upgrade_v2` 脚本](https://tensorflow.google.cn/guide/upgrade)将其全部重写为 `tf.compat.v1.summary`。要迁移到 TF 2.0,您需要以如下方式修改代码:
# + [markdown] id="Pq4Fy1bSUdrZ"
# 1. 必须存在通过 `.as_default()` 设置的默认写入器才能使用摘要运算
#
# - 这意味着在 Eager Execution 模式下执行运算或在计算图构造中使用运算
# - 如果没有默认写入器,摘要运算将变为静默空运算
# - 默认写入器(尚)不跨 `@tf.function` 执行边界传播(仅在跟踪函数时对其进行检测),所以最佳做法是在函数体中调用 `writer.as_default()`,并确保在使用 `@tf.function` 时,写入器对象始终存在
#
# 2. 必须通过 `step` 参数将“步骤”值传入每个运算
#
# - TensorBoard 需要步骤值以将数据呈现为时间序列
# - 由于 TF 1.x 中的全局步骤已被移除,因此需要执行显式传递,以确保每个运算都知道要读取的所需步骤变量
# - 为了减少样板,对注册默认步骤值的实验性支持通过 `tf.summary.experimental.set_step()` 提供,但这是临时功能,如有更改,恕不另行通知
#
# 3. 各个摘要运算的函数签名已更改
#
# - 现在,返回值为布尔值(指示是否实际写入了摘要)
# - 第二个参数名称(如果使用)已从 `tensor` 更改为 `data`
# - `collections` 参数已被移除;集合仅适用于 TF 1.x
# - `family` 参数已被移除;仅使用 `tf.name_scope()`
#
# 4. [仅针对旧计算图模式/会话执行用户]
#
# - 首先使用 `v1.Session.run(writer.init())` 初始化写入器
#
# - 使用 `v1.summary.all_v2_summary_ops()` 获取当前计算图的所有 TF 2.0 摘要运算,例如通过 `Session.run()` 执行它们
#
# - 使用 `v1.Session.run(writer.flush())` 刷新写入器,并以同样方式使用 `close()`
#
# 如果您的 TF 1.x 代码已改用 `tf.contrib.summary` API,因其与 TF 2.0 API 更加相似,`tf_upgrade_v2` 脚本将能够自动执行大多数迁移步骤(并针对无法完全迁移的任何用法发出警告或错误)。在大多数情况下,它只是将 API 调用重写为 `tf.compat.v2.summary`;如果只需要与 TF 2.0+ 兼容,那么您可以删除 `compat.v2` 并将其作为 `tf.summary` 引用。
# + [markdown] id="1GUZRWSkW3ZC"
# ## 其他提示
#
# 除上述重要内容以外,一些辅助方面也进行了更改:
#
# - 条件记录(例如“每 100 个步骤记录一次”)有所更新
#
# - 要控制运算和相关代码,请将其包装在常规 if 语句(可在 Eager 模式下运行,以及[通过 AutoGraph](https://tensorflow.google.cn/alpha/guide/autograph) 在 `@tf.function` 中使用)或 `tf.cond` 中
# - 要仅控制摘要,请使用新的 `tf.summary.record_if()` 上下文管理器,并将其传递给您选择的布尔条件
# - 以下内容替换了 TF 1.x 模式:
# ```
# if condition:
# writer.add_summary()
# ```
#
# + [markdown] id="9VMYrKn4Uh52"
# - 不直接编写 `tf.compat.v1.Graph` - 改为使用跟踪函数
#
# - TF 2.0 中的计算图执行使用 `@tf.function`,而非显式计算图
# - 在 TF 2.0 中,使用新的跟踪样式 API `tf.summary.trace_on()` 和 `tf.summary.trace_export()` 记录执行的函数计算图
#
# + [markdown] id="UGItA6U0UkDx"
# - 不再使用 `tf.summary.FileWriterCache` 按 logdir 缓存全局写入器
#
# - 用户应实现自己的写入器对象缓存/共享方案,或者使用独立的写入器(TensorBoard [正在](https://github.com/tensorflow/tensorboard/issues/1063)实现对后者的支持)
#
# + [markdown] id="d7BQJVcsUnMp"
# - 事件文件的二进制表示已更改
#
# - TensorBoard 1.x 已支持新格式;此项变更仅对从事件文件手动解析摘要数据的用户存在影响
# - 摘要数据现在以张量字节形式存储;您可以使用 `tf.make_ndarray(event.summary.value[0].tensor)` 将其转换为 Numpy
|
site/zh-cn/tensorboard/migrate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import requests
from bs4 import BeautifulSoup
# ### Naver Crawling Test
# - 웹툰명(w_name)
# - 해당 웹툰 링크
#
# - 소개글
# - 장르 : 쪼개져있는 장르를 모아야한다.
# - 작가
# - 완결여부
# - 평점
# - 하트수
# +
web_dic = {}
web_dic['webtoon_name'] = []
web_dic['webtoon_link'] = []
web_dic['webtoon_finished'] = []
response = requests.get("https://comic.naver.com/webtoon/genre.nhn?genre=comic")
dom = BeautifulSoup(response.content, "html.parser")
webtoons = dom.select(".list_area > ul.img_list > li")
for webtoon in webtoons:
web_dic['webtoon_name'].append(webtoon.select_one("dl > dt > a").get("title"))
web_dic['webtoon_link'].append("https://comic.naver.com" + webtoon.select_one("dl > dt > a").get("href"))
# print(len(webtoon.select("div > a > img")))
if len(webtoon.select("div > a > img")) > 1:
# print(webtoon.select("div > a > img")[1].get("alt"))
web_dic['webtoon_finished'].append(1 if webtoon.select("div > a > img")[1].get("alt") == "완결" else 0)
else:
web_dic['webtoon_finished'].append(0)
# -
def naver_webtoon(genre):
dic = {}
dic['unique_id'] = []
dic['webtoon_name'] = []
dic['author'] = []
dic['total_score'] = []
dic['webtoon_link'] = []
dic['image_link'] = []
dic['webtoon_finished'] = []
# first url for title
# get title with full name
url = "https://comic.naver.com/webtoon/genre.nhn?view=list&order=ViewCount&genre=" + genre
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
webtoons = dom.select(".list_area > ul.img_list > li")
for webtoon in webtoons:
web_dic['webtoon_name'].append(webtoon.select_one("dl > dt > a").get("title"))
dic['author'].append(webtoon.select_one("td:nth-of-type(3) > a").text)
# second url for the others
url = "https://comic.naver.com/webtoon/genre.nhn?genre=" + genre
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
keywords = dom.select(".list_area > .img_list > li")
# author, total score
for keyword in keywords:
dic['total_score'].append(keyword.select_one("dl > dd > .rating_type > strong").text)
# unique id
unique_id_list = dom.select("dl > dt > a")
for i in range(len(unique_id_list)):
idx = unique_id_list[i].get('href').index("=")
dic['unique_id'].append(unique_id_list[i].get('href')[idx+1:])
# main link
main_link_list = dom.select("dl > dt > a")
for i in range(len(main_link_list)):
dic['main_link'].append("https://comic.naver.com" + main_link_list[i].get('href'))
# image link
image_link_list = dom.select(".list_area > .img_list > li > .thumb > a > img:nth-of-type(1)")
for i in range(len(image_link_list)):
dic['image_link'].append(image_link_list[i].get('src'))
df = pd.DataFrame(dic)
df[genre] = genre
df['author'] = df['author'].map(lambda x : "".join(x.split()))
return df
web_dic['webtoon_link']
web_dic['webtoon_finished']
def naver_webtoon(genre):
dic = {}
dic['unique_id'] = []
dic['title'] = []
dic['author'] = []
dic['total_score'] = []
dic['main_link'] = []
dic['image_link'] = []
dic['update_date'] = []
# first url for title
# get title with full name
url = "https://comic.naver.com/webtoon/genre.nhn?view=list&order=ViewCount&genre=" + genre
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
keywords = dom.select("#content > div.list_area.table_list_area > table > tbody > tr")
for keyword in keywords:
dic['title'].append(keyword.select_one("td > a > strong").text)
dic['author'].append(keyword.select_one("td:nth-of-type(3) > a").text)
# second url for the others
url = "https://comic.naver.com/webtoon/genre.nhn?genre=" + genre
response = requests.get(url)
dom = BeautifulSoup(response.content, "html.parser")
keywords = dom.select(".list_area > .img_list > li")
# author, total score
for keyword in keywords:
dic['total_score'].append(keyword.select_one("dl > dd > .rating_type > strong").text)
dic['update_date'].append(keyword.select_one("dl > dd.date2").text)
# unique id
unique_id_list = dom.select("dl > dt > a")
for i in range(len(unique_id_list)):
idx = unique_id_list[i].get('href').index("=")
dic['unique_id'].append(unique_id_list[i].get('href')[idx+1:])
# main link
main_link_list = dom.select("dl > dt > a")
for i in range(len(main_link_list)):
dic['main_link'].append("https://comic.naver.com" + main_link_list[i].get('href'))
# image link
image_link_list = dom.select(".list_area > .img_list > li > .thumb > a > img:nth-of-type(1)")
for i in range(len(image_link_list)):
dic['image_link'].append(image_link_list[i].get('src'))
df = pd.DataFrame(dic)
df[genre] = genre
df['author'] = df['author'].map(lambda x : "".join(x.split()))
return df
|
crawling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="n6ecAvsmQp1I"
# ## To run this colab, press the "Runtime" button in the menu tab and then press the "Run all" button.
# + [markdown] colab_type="text" id="77gENRVX40S7"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="both" colab={} colab_type="code" id="d8jyt37T42Vf"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="hRTa3Ee15WsJ"
# # Recognize Flowers using Transfer Learning
# + [markdown] colab_type="text" id="dQHMcypT3vDT"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/community/en/flowers_tf_lite.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/examples/blob/master/community/en/flowers_tf_lite.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + colab={} colab_type="code" id="iBMcobPHdD8O"
import tensorflow as tf
assert tf.__version__.startswith('2')
import os
import numpy as np
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="NOG3l_MsBO1A"
tf.__version__
# + [markdown] colab_type="text" id="v77rlkCKW0IJ"
# ## Setup Input Pipeline
# + [markdown] colab_type="text" id="j4QOy2uA3P_p"
# Download the flowers dataset.
# + colab={} colab_type="code" id="xxL2mjVVGIrV"
_URL = "https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz"
zip_file = tf.keras.utils.get_file(origin=_URL,
fname="flower_photos.tgz",
extract=True)
base_dir = os.path.join(os.path.dirname(zip_file), 'flower_photos')
# + [markdown] colab_type="text" id="z4gTv7ig2vMh"
# Use `ImageDataGenerator` to rescale the images.
#
# Create the train generator and specify where the train dataset directory, image size, batch size.
#
# Create the validation generator with similar approach as the train generator with the flow_from_directory() method.
# + colab={} colab_type="code" id="aCLb_yV5JfF3"
IMAGE_SIZE = 224
BATCH_SIZE = 64
datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rescale=1./255,
validation_split=0.2)
train_generator = datagen.flow_from_directory(
base_dir,
target_size=(IMAGE_SIZE, IMAGE_SIZE),
batch_size=BATCH_SIZE,
subset='training')
val_generator = datagen.flow_from_directory(
base_dir,
target_size=(IMAGE_SIZE, IMAGE_SIZE),
batch_size=BATCH_SIZE,
subset='validation')
# + colab={} colab_type="code" id="tx1L7fxxWA_G"
for image_batch, label_batch in train_generator:
break
image_batch.shape, label_batch.shape
# + [markdown] colab_type="text" id="ZrFFcwUb3iK9"
# Save the labels in a file which will be downloaded later.
# + colab={} colab_type="code" id="-QFZIhWs4dsq"
print (train_generator.class_indices)
labels = '\n'.join(sorted(train_generator.class_indices.keys()))
with open('labels.txt', 'w') as f:
f.write(labels)
# + colab={} colab_type="code" id="duxD_UDSOmng"
# !cat labels.txt
# + [markdown] colab_type="text" id="OkH-kazQecHB"
# ## Create the base model from the pre-trained convnets
#
# Create the base model from the **MobileNet V2** model developed at Google, and pre-trained on the ImageNet dataset, a large dataset of 1.4M images and 1000 classes of web images.
#
# First, pick which intermediate layer of MobileNet V2 will be used for feature extraction. A common practice is to use the output of the very last layer before the flatten operation, the so-called "bottleneck layer". The reasoning here is that the following fully-connected layers will be too specialized to the task the network was trained on, and thus the features learned by these layers won't be very useful for a new task. The bottleneck features, however, retain much generality.
#
# Let's instantiate an MobileNet V2 model pre-loaded with weights trained on ImageNet. By specifying the `include_top=False` argument, we load a network that doesn't include the classification layers at the top, which is ideal for feature extraction.
# + colab={} colab_type="code" id="19IQ2gqneqmS"
IMG_SHAPE = (IMAGE_SIZE, IMAGE_SIZE, 3)
# Create the base model from the pre-trained model MobileNet V2
base_model = tf.keras.applications.MobileNetV2(input_shape=IMG_SHAPE,
include_top=False,
weights='imagenet')
# + [markdown] colab_type="text" id="rlx56nQtfe8Y"
# ## Feature extraction
# You will freeze the convolutional base created from the previous step and use that as a feature extractor, add a classifier on top of it and train the top-level classifier.
# + colab={} colab_type="code" id="Tts8BbAtRGvk"
base_model.trainable = False
# + [markdown] colab_type="text" id="wdMRM8YModbk"
# ### Add a classification head
# + colab={} colab_type="code" id="eApvroIyn1K0"
model = tf.keras.Sequential([
base_model,
tf.keras.layers.Conv2D(32, 3, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.GlobalAveragePooling2D(),
tf.keras.layers.Dense(5, activation='softmax')
])
# + [markdown] colab_type="text" id="g0ylJXE_kRLi"
# ### Compile the model
#
# You must compile the model before training it. Since there are multiple classes, use a categorical cross-entropy loss.
# + colab={} colab_type="code" id="RpR8HdyMhukJ"
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss='categorical_crossentropy',
metrics=['accuracy'])
# + colab={} colab_type="code" id="I8ARiyMFsgbH"
model.summary()
# + colab={} colab_type="code" id="krvBumovycVA"
print('Number of trainable variables = {}'.format(len(model.trainable_variables)))
# + [markdown] colab_type="text" id="RxvgOYTDSWTx"
# ### Train the model
#
# <!-- TODO(markdaoust): delete steps_per_epoch in TensorFlow r1.14/r2.0 -->
# + colab={} colab_type="code" id="JsaRFlZ9B6WK"
epochs = 10
history = model.fit(train_generator,
steps_per_epoch=len(train_generator),
epochs=epochs,
validation_data=val_generator,
validation_steps=len(val_generator))
# + [markdown] colab_type="text" id="Hd94CKImf8vi"
# ### Learning curves
#
# Let's take a look at the learning curves of the training and validation accuracy/loss when using the MobileNet V2 base model as a fixed feature extractor.
# + colab={} colab_type="code" id="53OTCh3jnbwV"
acc = history.history['accuracy']
val_acc = history.history['val_accuracy']
loss = history.history['loss']
val_loss = history.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,1.0])
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.show()
# + [markdown] colab_type="text" id="CqwV-CRdS6Nv"
# ## Fine tuning
# In our feature extraction experiment, you were only training a few layers on top of an MobileNet V2 base model. The weights of the pre-trained network were **not** updated during training.
#
# One way to increase performance even further is to train (or "fine-tune") the weights of the top layers of the pre-trained model alongside the training of the classifier you added. The training process will force the weights to be tuned from generic features maps to features associated specifically to our dataset.
# + [markdown] colab_type="text" id="CPXnzUK0QonF"
# ### Un-freeze the top layers of the model
#
# + [markdown] colab_type="text" id="rfxv_ifotQak"
# All you need to do is unfreeze the `base_model` and set the bottom layers be un-trainable. Then, recompile the model (necessary for these changes to take effect), and resume training.
# + colab={} colab_type="code" id="4nzcagVitLQm"
base_model.trainable = True
# + colab={} colab_type="code" id="-4HgVAacRs5v"
# Let's take a look to see how many layers are in the base model
print("Number of layers in the base model: ", len(base_model.layers))
# Fine tune from this layer onwards
fine_tune_at = 100
# Freeze all the layers before the `fine_tune_at` layer
for layer in base_model.layers[:fine_tune_at]:
layer.trainable = False
# + [markdown] colab_type="text" id="4Uk1dgsxT0IS"
# ### Compile the model
#
# Compile the model using a much lower training rate.
# + colab={} colab_type="code" id="NtUnaz0WUDva"
model.compile(loss='categorical_crossentropy',
optimizer = tf.keras.optimizers.Adam(1e-5),
metrics=['accuracy'])
# + colab={} colab_type="code" id="WwBWy7J2kZvA"
model.summary()
# + colab={} colab_type="code" id="bNXelbMQtonr"
print('Number of trainable variables = {}'.format(len(model.trainable_variables)))
# + [markdown] colab_type="text" id="4G5O4jd6TuAG"
# ### Continue Train the model
# + colab={} colab_type="code" id="ECQLkAsFTlun"
history_fine = model.fit_generator(train_generator,
epochs=5,
validation_data=val_generator)
# + [markdown] colab_type="text" id="kRDabW_u1wnv"
# ## Convert to TFLite
# + [markdown] colab_type="text" id="hNvMl6CM6lG4"
# Saved the model using `tf.saved_model.save` and then convert the saved model to a tf lite compatible format.
# + colab={} colab_type="code" id="_LZiKVInWNGy"
saved_model_dir = 'save/fine_tuning'
tf.saved_model.save(model, saved_model_dir)
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_dir)
tflite_model = converter.convert()
with open('model.tflite', 'wb') as f:
f.write(tflite_model)
# + [markdown] colab_type="text" id="GE4w-9S410Dk"
# Download the converted model and labels
# + colab={} colab_type="code" id="x47uW_lI1DoV"
from google.colab import files
files.download('model.tflite')
files.download('labels.txt')
# + [markdown] colab_type="text" id="TfXEmsxQf6eP"
# Let's take a look at the learning curves of the training and validation accuracy/loss, when fine tuning the last few layers of the MobileNet V2 base model and training the classifier on top of it. The validation loss is much higher than the training loss, so you may get some overfitting.
#
# You may also get some overfitting as the new training set is relatively small and similar to the original MobileNet V2 datasets.
#
# + colab={} colab_type="code" id="chW103JUItdk"
acc = history_fine.history['accuracy']
val_acc = history_fine.history['val_accuracy']
loss = history_fine.history['loss']
val_loss = history_fine.history['val_loss']
plt.figure(figsize=(8, 8))
plt.subplot(2, 1, 1)
plt.plot(acc, label='Training Accuracy')
plt.plot(val_acc, label='Validation Accuracy')
plt.legend(loc='lower right')
plt.ylabel('Accuracy')
plt.ylim([min(plt.ylim()),1])
plt.title('Training and Validation Accuracy')
plt.subplot(2, 1, 2)
plt.plot(loss, label='Training Loss')
plt.plot(val_loss, label='Validation Loss')
plt.legend(loc='upper right')
plt.ylabel('Cross Entropy')
plt.ylim([0,1.0])
plt.title('Training and Validation Loss')
plt.xlabel('epoch')
plt.show()
# + [markdown] colab_type="text" id="_TZTwG7nhm0C"
# ## Summary:
#
# * **Using a pre-trained model for feature extraction**: When working with a small dataset, it is common to take advantage of features learned by a model trained on a larger dataset in the same domain. This is done by instantiating the pre-trained model and adding a fully-connected classifier on top. The pre-trained model is "frozen" and only the weights of the classifier get updated during training.
# In this case, the convolutional base extracted all the features associated with each image and you just trained a classifier that determines the image class given that set of extracted features.
#
# * **Fine-tuning a pre-trained model**: To further improve performance, one might want to repurpose the top-level layers of the pre-trained models to the new dataset via fine-tuning.
# In this case, you tuned your weights such that your model learned high-level features specific to the dataset. This technique is usually recommended when the training dataset is large and very similar to the orginial dataset that the pre-trained model was trained on.
#
|
community/en/flowers_tf_lite.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
# create dummy matrix
n_plots = 4
rows = cols = 80
plot_length = int(rows/n_plots)
dummy_plot = np.ones(plot_length**2).reshape((plot_length, plot_length))
A1 = dummy_plot
A2 = dummy_plot * 2
B1 = dummy_plot * 3
B2 = dummy_plot * 4
large_matrix = np.block([[A1, A2], [B1, B2]])
large_dummy_matrix = np.ones_like(large_matrix)
n = plot_length
coef_matrix = np.array([[1, 2], [3, 4]])
result = np.multiply(large_dummy_matrix, np.kron(coef_matrix, np.ones((n,n))))
matrix_indizes = np.indices((n_plots, n_plots), dtype="uint8") + 1
row_indizes, column_indizes = matrix_indizes[0], matrix_indizes[1]
plot_definition_matrix = np.char.add(row_indizes.astype(np.str), column_indizes.astype(np.str)).astype(np.uint8)
dummy_playing_field_matrix = np.ones(shape=(rows, cols), dtype=np.uint8)
large_plot_definition_matrix = np.multiply(
dummy_playing_field_matrix,
np.kron(plot_definition_matrix, np.ones(shape=(plot_length, plot_length)))
)
# +
lulc_matrix = dummy_playing_field_matrix # 80 x 80
cooperation_matrix = dummy_playing_field_matrix # 80 x 80
plot_definition_matrix # 4 x 4
tourism_matrix = plot_definition_matrix # 4 x 4
# -
tourism_matrix
# +
def teamwork(cooperation_matrix):
"""
assumes four players. if the four corner player say yes then it's true.
Args:
cooperation_matrix: a numpy matrix where the corner elements are a boolean that is true if the associated player wants teamwork.
Returns:
a boolean - true if each corner player wants teamwork.
"""
teamwork = False
row, col = cooperation_matrix.shape
if (
cooperation_matrix[0][0]
== cooperation_matrix[0][col - 1]
== cooperation_matrix[col - 1][0]
== cooperation_matrix[col - 1][col - 1]
== True
):
teamwork = True
return teamwork
# -
teamwork(cooperation_matrix)
# +
def unemployment_rate(earning, earning_new, unemployment):
'''
Args:
earning: how much money the player earned last round
earning_new: how much money the player earned this round
unemployment: unemployment rate of the player had last round
Returns: unemployment rate of the player has this round
'''
# based on okuns law
delta_unemployment = ((earning-earning_new)/earning)/1.8
new_unemployment = max(0,unemployment + delta_unemployment)
return new_unemployment
# get the total yield for the current map
# not sure why this is necessary - the yield map function calculates the number of pixel for a matrix with values of simplified_lulc_mapping
# -
unempl = unemployment_rate(2500, 2400, 0.05)
unempl
3.2/1.8
# +
def yield_map(field):
'''
Args:
field: a matrix with values corresponding to simplified_lulc_mapping
Returns: the number of pixel for each value of simplified_lulc_mapping
'''
tot_cattle = np.count_nonzero(field == simplified_lulc_mapping['Cattle Farming'])
tot_sheep = np.count_nonzero(field == simplified_lulc_mapping['Sheep Farming'])
tot_n_forest = np.count_nonzero(field == simplified_lulc_mapping['Commercial Forest'])
tot_c_forest = np.count_nonzero(field == simplified_lulc_mapping['Cattle Farming'])
return tot_cattle, tot_sheep, tot_n_forest, tot_c_forest
simplified_lulc_mapping = {
"Sheep Farming": 1,
"Native Forest": 2,
"Commercial Forest": 3,
"Cattle Farming": 4,
}
# -
c,s,n,c = yield_map (lulc_matrix)
s
m, n = tourism_matrix.shape
# +
def tourism(tourism_matrix, gdp_tourism):
'''
Args:
tourism_matrix: part of lulc mapping in which tourism takes place (20x20)
Returns: number of tourists and a factor "tourism" that improves earnings.
'''
cattle, sheep, n_forest, c_forest = yield_map(tourism_matrix)
sum = cattle + sheep + n_forest + c_forest
m, n = tourism_matrix.shape
sum = max(sum, m*n) # that line was added for testing- it is needed because tourism_matrix isn't part of lulc_matrix
number_tourists = sheep * 2 + n_forest * 3 - c_forest * 5
# a minimum number of tourits always find their way
number_tourists = max(number_tourists, sum * 0.75)
# no more increased sale - tourists start thinking this is too expensive and full
tourism_factor = min(number_tourists/sum, 1 + gdp_tourism/100)
#beach bonus
if sum < m**2 - 10:
tourism_factor = max(0.95, tourism_factor*1.2)
return number_tourists, tourism_factor
# -
nt, tf = tourism(tourism_matrix, 40)
nt
tf
def crop_field(field):
"""
Devides an 2D numpy array of quadratric form into four parts.
Args:
field: a 2D array
Returns: 4 2d arrays of 1/4 the size
"""
m, n = field.shape
indices1 = list(range(0, int(n / 2)))
indices2 = list(range(int(n / 2), n))
fa1 = field[np.ix_(indices1, indices1)]
fo1 = field[np.ix_(indices2, indices1)]
fa2 = field[np.ix_(indices1, indices2)]
fo2 = field[np.ix_(indices2, indices2)]
return fa1, fa2, fo1, fo2
fa1, fa2, fo1, fo2 = crop_field(lulc_matrix)
fa1.shape
SUBSIDIES = 0.8
# +
# adapt the prices
def price_per_pixel(
current_round,
brexit,
tot_sheep_0,
tot_cattle,
tot_n_forest,
tot_c_forest,
income_farmland_cattle,
income_farmland_sheep,
income_forest_native,
income_forest_commercial,
):
"""
calculates current prices depending on demand based on an estimate on what's produced in the beginning.
Args:
current_round: timeline of the game
brexit: time at which brexit happens
tot_c_forest_0: total amount of pixel at the start
tot_n_forest_0:
tot_sheep_0
tot_cattle: total amount of pixel for each the current round
tot_n_forest:
tot_c_forest:
income_farmland_cattle,
income_farmland_sheep,
income_forest_native,
income_forest_commercial,
Returns: the price of the current round for cattle, sheep, native forest and commercial forest
"""
# doesn't take tourism_factor effects into account yet. and the equations are pretty random.
cattle_price_new = income_farmland_cattle + (tot_cattle/(income_farmland_cattle/income_farmland_sheep)/tot_sheep_0)*(income_farmland_sheep-income_farmland_cattle)
sheep_price_new = income_farmland_sheep # assume sheep can go everywhere, eat everything and no degradation and its profit only influences cattle by competition
c_forest_price_new = income_forest_commercial + tot_c_forest /(income_forest_commercial/income_forest_native)/(tot_n_forest+tot_c_forest)*(income_forest_native - income_forest_commercial)
n_forest_price_new = income_forest_native # assumes native forest can grow everywhere and its profit only influences the commercial forest through competition in the timber market
if brexit > current_round:
n_forest_price_new = n_forest_price_new/(1+SUBSIDIES)*2 # less import of wood.
c_forest_price_new = c_forest_price_new/SUBSIDIES
return cattle_price_new, sheep_price_new, n_forest_price_new, c_forest_price_new
# -
c, s, n, cf = price_per_pixel(2,3,40,10,10,10,100,30,50,200)
n
LANDUSE_CHANGE = 0.25
COSTS_LANDUSE_CHANGE = 10
# +
def money_farmer(
current_round,
tourism_factor,
teams,
brexit,
teamwork,
area_sheep,
area_cattle,
area_c_forest,
area_n_forest,
sheep_price,
cattle_price,
n_forest_price,
c_forest_price,
bank_account_farmer_1,
bank,
gdp_pc_scotland,
):
if current_round == 0:
earning = gdp_pc_scotland
bank_current = bank_account_farmer_1
else:
# costs of landscape change
try:
d_sheep = area_sheep[current_round] - area_sheep[current_round - 1]
except:
pass
d_cattle = area_cattle[current_round] - area_cattle[current_round - 1]
d_n_forest = (
area_n_forest[current_round] - area_n_forest[current_round - 1]
) # necessary to potentially allow two changes (i.e. a rise or native forests and cattle on cost of sheep )
m_change = 0
m_brexit = 0
m_teamwork = 0
if d_n_forest < 0:
m_change += (
min([d_cattle, d_n_forest], key=abs)
* LANDUSE_CHANGE
* cattle_price[current_round]
)
m_change += (
min([d_sheep, d_n_forest], key=abs) * LANDUSE_CHANGE * sheep_price[current_round]
)
if d_sheep < 0:
m_change += (
min([d_cattle, d_sheep], key=abs) * LANDUSE_CHANGE * cattle_price[current_round]
)
m_change += (
min([d_sheep, d_n_forest], key=abs)
* LANDUSE_CHANGE
* n_forest_price[current_round]
)
if d_cattle < 0:
m_change += (
min([d_cattle, d_sheep], key=abs) * LANDUSE_CHANGE * sheep_price[current_round]
)
m_change += (
min([d_cattle, d_n_forest], key=abs)
* LANDUSE_CHANGE
* n_forest_price[current_round]
)
# earning from the area
m_area = (area_sheep[current_round] * sheep_price[current_round]) + (
area_cattle[current_round] * cattle_price[current_round]
)
#divided by 10 to reduce that profit
if teamwork == True and teams <= current_round:
m_teamwork = (
area_c_forest[current_round] * c_forest_price[current_round]/10
+ area_n_forest[current_round] * n_forest_price[current_round]/10
)
if brexit <= current_round:
m_brexit = (SUBSIDIES - 1) * (area_sheep[current_round] * sheep_price[current_round])
m_tourism = tourism_factor * m_area - m_area
earning = (m_area + m_change + m_tourism + m_teamwork + m_brexit)*gdp_pc_scotland/(50*(area_sheep[current_round]+area_cattle[current_round]))
bank_current = bank - (d_cattle + d_n_forest + d_sheep)/2*COSTS_LANDUSE_CHANGE + earning - gdp_pc_scotland
return earning, bank_current, m_area, m_change, m_tourism, m_brexit, m_teamwork
# -
money_farmer(
current_round = 2,
tourism_factor = 1.2,
teams=1,
brexit=1,
teamwork=True,
area_sheep=[6,2,8],
area_cattle=[2,6,0],
area_c_forest=[5,7,9],
area_n_forest=[5,3,1],
sheep_price=[3,3,50],
cattle_price=[3,3,100],
n_forest_price=[3,3,20],
c_forest_price=[3,3,200],
bank_account_farmer_1=1000,
bank=1000,
gdp_pc_scotland= 30000,
)
# +
#if d_n_forest > 0:
# m_brexit += d_n_forest * (SUBSIDIES - 1)
# -
def money_forester(
current_round,
tourism_factor,
teams,
brexit,
teamwork,
area_sheep,
area_c_forest,
area_n_forest,
sheep_price,
n_forest_price,
c_forest_price,
bank_account_forestry_1,
bank,
gdp_pc_scotland
):
'''
Args:
current_round:
tourism_factor: tourism_factor (from tourism function
teams: in which round are teams allowed
brexit: in which round does brexit happen - doesn't include higher timber prices here.
teamwork: is teamwork true or false
area_sheep: number of pixel with the respective landuse
area_cattle:
area_c_forest:
area_n_forest:
sheep_price: list of prices
cattle_price:
n_forest_price:
c_forest_price:
bank_account_farmer_1: assumes same bank_account at the begining for both farmer
bank: how much money is in the bank-account (in round 0 bank_account_farmer_1
gdp_pc_scotland:
Returns: earnings and bank-account
'''
if current_round == 0:
earning = gdp_pc_scotland
bank_current = bank_account_forestry_1
else:
d_n_forest = (
area_n_forest[current_round] - area_n_forest[current_round - 1]
) # necessary to potentially allow two changes (i.e. a rise or native forests and cattle on cost of sheep )
m_change = 0
m_brexit = 0
m_teamwork = 0
# ich habe momentan gemacht, dass man nur etwas verkleinern darf!
if d_n_forest < 0:
m_change += d_n_forest * LANDUSE_CHANGE * c_forest_price[current_round]
if d_n_forest > 0:
m_change += d_n_forest * LANDUSE_CHANGE * n_forest_price[current_round]
# earning from the area
m_area = (area_n_forest[current_round] * n_forest_price[current_round]) + (
(area_c_forest[current_round] * c_forest_price[current_round])
)
if teamwork == True and teams <= current_round:
m_teamwork = area_sheep[current_round] * sheep_price[current_round]
if brexit <= current_round:
if d_n_forest > 0:
m_brexit = d_n_forest * (SUBSIDIES - 1)
m_brexit += (SUBSIDIES - 1) * (area_sheep[current_round] * sheep_price[current_round])
m_tourism_factor = tourism_factor * m_area
# maybe return later on the performance of each landuse/industrie --> append() so that its easy to plot?
# has to make more - maybe beacause he has less land? has still much more...
earning = (m_area + m_change + m_tourism_factor + m_teamwork + m_brexit)*gdp_pc_scotland/(100*(area_n_forest[current_round]+area_c_forest[current_round]))
bank_current = bank - abs(d_n_forest)*COSTS_LANDUSE_CHANGE + earning - gdp_pc_scotland
return earning, bank_current#, m_area, m_change, m_tourism_factor, m_teamwork, m_brexit
money_forester(
current_round = 2,
tourism_factor = 1.2,
teams=1,
brexit=1,
teamwork=True,
area_sheep=[6,2,8],
area_c_forest=[5,7,5],
area_n_forest=[5,3,5],
sheep_price=[3,3,50],
n_forest_price=[3,3,20],
c_forest_price=[3,3,200],
bank_account_forestry_1=1000,
bank=1000,
gdp_pc_scotland= 30000,
)
# +
# maps = {}
# maps["round_0"] = sumplified_maps
#new_simplified_map = ...
#map["round_2"]= new_simplified_map
#field.simplified_maps["roudn_2"]
# assumes four players only / if the four corner player say yes then it's true.
#field.map_simplified
# cm = ConceptualModel()
#cm.bank_accound_farmer1["roudn:2"]
#game_model.model_parameters.loc[5, value]
#game_model.bank_account_farmers1["round_1"]
# -
|
notebooks/tryout_methods.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Root
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="YiUdpEJYUt2Z"
# # [Practical Text Classification With Python and Keras - CNN](https://realpython.com/python-keras-text-classification/?fbclid=IwAR0uuIjJZMfOsT0BDBT_-MQnW3MjwZqpFZ3xkEh2TwZLjbgki5zfeiP91KM)
#
# Imagine you could know the mood of the people on the Internet. Maybe you are not interested in its entirety, but only if people are today happy on your favorite social media platform. After this tutorial, you’ll be equipped to do this. While doing this, you will get a grasp of current advancements of (deep) neural networks and how they can be applied to text.
#
# Reading the mood from text with machine learning is called [sentiment analysis](https://en.wikipedia.org/wiki/Sentiment_analysis) , and it is one of the prominent use cases in text classification. This falls into the very active research field of natural language processing (NLP). Other common use cases of text classification include detection of spam, auto tagging of customer queries, and categorization of text into defined topics. So how can you do this?
#
# ### Reference:
#
# https://realpython.com/python-keras-text-classification
#
# ### Dataset
# https://archive.ics.uci.edu/ml/machine-learning-databases/00331/
# + colab={} colab_type="code" id="IZ3aj-kKE8xg"
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
from IPython.display import display
# + [markdown] colab_type="text" id="xk8c4UojGoo5"
# ## Dataset
# This dataset was created for the Paper 'From Group to Individual Labels using Deep Features', Kotzias et. al,. KDD 2015
# Please cite the paper if you want to use it :)
#
# It contains sentences labelled with positive or negative sentiment, extracted from reviews of products, movies, and restaurants
#
#
# ### Format:
# sentence \t score \n
#
#
# ### Details:
#
# Score is either 1 (for positive) or 0 (for negative)
# The sentences come from three different websites/fields:
#
# ```
# imdb.com
# amazon.com
# yelp.com
# ```
#
# For each website, there exist 500 positive and 500 negative sentences. Those were selected randomly for larger datasets of reviews.
#
# We attempted to select sentences that have a clearly positive or negative connotaton, the goal was for no neutral sentences to be selected.
#
# For the full datasets look:
#
# - imdb: Maas et. al., 2011 'Learning word vectors for sentiment analysis'
# - amazon: McAuley et. al., 2013 'Hidden factors and hidden topics: Understanding rating dimensions with review text'
# - yelp: Yelp dataset challenge http://www.yelp.com/dataset_challenge
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 73, "resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "headers": [["content-type", "application/javascript"]], "ok": true, "status": 200, "status_text": ""}}} colab_type="code" id="cC83fYPXGbm-" outputId="59fabb8a-5da6-422d-c86b-4d7547c2ec32"
from google.colab import files
uploaded = files.upload()
# + colab={"base_uri": "https://localhost:8080/", "height": 90} colab_type="code" id="RXqs8Xx-HZuy" outputId="96a4a2c8-d3a8-4d4c-cc6a-3b94fd6b6830"
# ! tar -xvf sentiment_label_sentences.tar
# ! rm -rf sentiment_label_sentences.tar
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="LPHRuM5kHek-" outputId="96b0a033-b106-4dde-fc3a-8ea1c111aa43"
# ! ls sentiment_label_sentences
# + colab={"base_uri": "https://localhost:8080/", "height": 562} colab_type="code" id="Lnlfb78HHe02" outputId="7cb64308-ff56-497b-efe3-1c9a49b7f027"
filepath_dict = {
'yelp': 'sentiment_label_sentences/yelp_labelled.txt',
'amazon': 'sentiment_label_sentences/amazon_cells_labelled.txt',
'imdb': 'sentiment_label_sentences/imdb_labelled.txt'}
df_list = []
for source, filepath in filepath_dict.items():
df = pd.read_csv(filepath, names=['sentence', 'label'], sep='\t')
df['source'] = source # Add another column filled with the source name
df_list.append(df)
df = pd.concat(df_list)
display(df.info())
display(df.head())
display(print('Label value count:'))
display(df['label'].value_counts())
display(print('Source value count:'))
display(df['source'].value_counts())
# + [markdown] colab_type="text" id="aw1dWjRvHp6n"
# ## Train/Test Split
# + colab={"base_uri": "https://localhost:8080/", "height": 199} colab_type="code" id="2cUbFChnHipN" outputId="7491d4bf-f87c-43a0-d110-04719e0a755e"
from sklearn.model_selection import train_test_split
# Train data from all dataset
X = df['sentence'].values
y = df['label'].values
# Train/test split
X_sentence_train, X_sentence_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=1000)
display(X_sentence_train.shape)
display(y_train.shape)
display(X_sentence_test.shape)
display(y_test.shape)
print('')
display(X_sentence_train[0])
display(y_train[0])
print('')
display(X_sentence_train[5])
display(y_train[5])
# + [markdown] colab_type="text" id="IisJ2u0-IKDU"
# ## Feature Engineering - Word Embeddings
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="uqkgavnGH6jH" outputId="db01c02a-08dc-4d44-a6a4-66b548f3c874"
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
# Torkenize the sentences
tokenizer = Tokenizer(num_words=5000)
tokenizer.fit_on_texts(X_sentence_train)
### Since you might not have the testing data available during training,
### you can create the vocabulary using only the training data
# Tokenization
X_train = tokenizer.texts_to_sequences(X_sentence_train)
X_test = tokenizer.texts_to_sequences(X_sentence_test)
# vocab size
vocab_size = len(tokenizer.word_index) + 1
# Padding
maxlen = 100
X_train_pad = pad_sequences(X_train, padding='post', maxlen=maxlen)
X_test_pad = pad_sequences(X_test, padding='post', maxlen=maxlen)
display(X_train_pad.shape)
display(X_test_pad.shape)
# -
# ## Model - CNN
# - Working with sequential data, like text
# -> work with one dimensional convolutions
#
# - one dimensional convnet is invariant to translations
# -> Certain sequences can be recognized at a different position
# -> Discover certain patterns in the text
#
# <img src="images/njanakiev-1d-convolution.jpg" alt="drawing" width="600"/>
#
# - CNN not work well on this dataset
# - There are not enough training samples
# - The data does not generalize well
#
# - CNNs work best with large training sets where they are able to find generalizations where a simple model like logistic regression won’t be able.
# + colab={"base_uri": "https://localhost:8080/", "height": 326} colab_type="code" id="pN-uKswfIQPX" outputId="9a9ef6cc-2be0-4d80-c3fa-da29b928d866"
from keras.models import Sequential
from keras import layers
import keras.backend as K
embedding_dim = 100
K.clear_session()
model = Sequential()
model.add(layers.Embedding(
vocab_size,
embedding_dim,
input_length=maxlen))
# CNN
model.add(layers.Conv1D(128, 5, activation='relu'))
model.add(layers.GlobalMaxPooling1D())
model.add(layers.Dense(10, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
model.summary()
# + colab={} colab_type="code" id="PFK17EDYJNJT"
from keras.callbacks import EarlyStopping
early_stopping = EarlyStopping(
monitor='val_loss',
min_delta=0,
patience=2,
verbose=0, mode='auto')
history = model.fit(
X_train_pad, y_train,
epochs=50,
verbose=False,
validation_split=0.3,
callbacks=[early_stopping],
batch_size=10)
# + colab={"base_uri": "https://localhost:8080/", "height": 347} colab_type="code" id="AI6aMuyLJQ_Z" outputId="368e889e-b900-441f-e980-74ae40681f2f"
historydf = pd.DataFrame(history.history, index=history.epoch)
historydf.plot();
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="1wCrP8mTJTcq" outputId="97cab74d-b668-4c3c-fbe3-6d246bf2798c"
loss, accuracy = model.evaluate(
X_train_pad, y_train,
verbose=False)
print("Training Accuracy: {:.4f}".format(accuracy))
# + [markdown] colab_type="text" id="CIkkc6DqJXmL"
# ## Evaluate
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="-KWAX46yJVRr" outputId="b352de90-926c-4170-e196-3ea8bfae5548"
loss, accuracy = model.evaluate(
X_test_pad, y_test,
verbose=False)
print("Testing Accuracy: {:.4f}".format(accuracy))
# + [markdown] colab_type="text" id="A6q3aUFGLnbe"
# ## Grid Search
# + colab={} colab_type="code" id="dAzezjk3JaoR"
def create_model(num_filters, kernel_size, vocab_size, embedding_dim, maxlen):
K.clear_session()
model = Sequential()
model.add(layers.Embedding(vocab_size, embedding_dim, input_length=maxlen))
model.add(layers.Conv1D(num_filters, kernel_size, activation='relu'))
model.add(layers.GlobalMaxPooling1D())
model.add(layers.Dense(10, activation='relu'))
model.add(layers.Dense(1, activation='sigmoid'))
model.compile(optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
return model
# + colab={"base_uri": "https://localhost:8080/", "height": 617} colab_type="code" id="rsx2w_irKDOw" outputId="306f4cf6-6889-44c6-c09a-601ee1ebb697"
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import RandomizedSearchCV
# Main settings
epochs = 20
embedding_dim = 50
maxlen = 100
# Run grid search for each source (yelp, amazon, imdb)
for source, frame in df.groupby('source'):
print('Running grid search for data set :', source)
# X, y
X = df['sentence'].values
y = df['label'].values
# Train-test split
X_sentence_train, X_sentence_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=1000)
# Torkenize the sentences
tokenizer = Tokenizer(num_words=5000)
tokenizer.fit_on_texts(X_sentence_train)
# Tokenization
X_train = tokenizer.texts_to_sequences(X_sentence_train)
X_test = tokenizer.texts_to_sequences(X_sentence_test)
# vocab size
vocab_size = len(tokenizer.word_index) + 1
# Padding
maxlen = 100
X_train_pad = pad_sequences(X_train, padding='post', maxlen=maxlen)
X_test_pad = pad_sequences(X_test, padding='post', maxlen=maxlen)
# Model
model = KerasClassifier(
build_fn=create_model,
epochs=epochs, batch_size=10,
verbose=False)
# Parameter grid for grid search
param_grid = dict(
num_filters=[32, 64, 128],
kernel_size=[3, 5, 7],
vocab_size=[vocab_size],
embedding_dim=[embedding_dim],
maxlen=[maxlen])
# Grid search
grid = RandomizedSearchCV(
estimator=model,
param_distributions=param_grid,
cv=4, verbose=1, n_iter=5)
grid_result = grid.fit(X_train_pad, y_train)
# Evaluate testing set
test_accuracy = grid.score(X_test_pad, y_test)
# Evaluate and print results
print('''
Running {} data set
Best Accuracy : {:.4f}
Best params: {}
Test Accuracy : {:.4f}\n\n
'''.format(
source,
grid_result.best_score_,
grid_result.best_params_,
test_accuracy))
# + colab={"base_uri": "https://localhost:8080/", "height": 199} colab_type="code" id="GStCr38_VkYt" outputId="cd042abb-858c-4c14-8197-0774ad2a2bb3"
from keras.wrappers.scikit_learn import KerasClassifier
from sklearn.model_selection import RandomizedSearchCV
# Main settings
epochs = 20
embedding_dim = 50
maxlen = 100
# X, y
X = df['sentence'].values
y = df['label'].values
# Train-test split
X_sentence_train, X_sentence_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=1000)
# Torkenize the sentences
tokenizer = Tokenizer(num_words=5000)
tokenizer.fit_on_texts(X_sentence_train)
# Tokenization
X_train = tokenizer.texts_to_sequences(X_sentence_train)
X_test = tokenizer.texts_to_sequences(X_sentence_test)
# vocab size
vocab_size = len(tokenizer.word_index) + 1
# Padding
maxlen = 100
X_train_pad = pad_sequences(X_train, padding='post', maxlen=maxlen)
X_test_pad = pad_sequences(X_test, padding='post', maxlen=maxlen)
# Model
model = KerasClassifier(
build_fn=create_model,
epochs=epochs,
batch_size=10,
validation_split=0.3,
verbose=False)
# Parameter grid for grid search
param_grid = dict(
num_filters=[32, 64, 128],
kernel_size=[3, 5, 7],
vocab_size=[vocab_size],
embedding_dim=[embedding_dim],
maxlen=[maxlen])
# Grid search
grid = RandomizedSearchCV(
estimator=model,
param_distributions=param_grid,
cv=4, verbose=1, n_iter=5)
early_stopping = EarlyStopping(
monitor='val_loss',
min_delta=0,
patience=2,
verbose=0, mode='auto')
grid_result = grid.fit(
X_train_pad, y_train,
callbacks=[early_stopping])
# Evaluate testing set
test_accuracy = grid.score(X_test_pad, y_test)
# Evaluate and print results
print('''
Best Accuracy : {:.4f}
Best params: {}
Test Accuracy : {:.4f}\n\n
'''.format(
grid_result.best_score_,
grid_result.best_params_,
test_accuracy))
|
Problems/Text_Classification/2_Text_Classification_Hyperparameters.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.0
# language: julia
# name: julia-1.6
# ---
srand(46);
# ベクトルについて
# ベクトルをつくる
x = []
# なんでも入る
x = Int[]
x = Vector{Int}()
# Int[]とおなじ
x = [1,2,3]
# 全て同じベクトルをつくる
fill(42, 10)
fill("foo", 4)
# ベクトルに追加する
x = [1,2,3]
push!(x,4) # 右に追加
println(x)
# unshift!(x,0) # 左に追加 julia1.6では廃止
pushfirst!(x,0)
println(x)
# 別のベクトルを追加
append!(x,[5,6,7])
println(x)
# ベクトル の要素を削除する
x = [1,2,3,4,5]
pop!(x)
println(x)
# shift!(x) # julia 1.6では廃止
popfirst!(x)# 左端を削除
println(x)
deleteat!(x,2) # 指定した場所を削除
println(x)
# べクトル がソート済みかどうか
println(issorted([1,2,3,4]))
println(issorted([2,1,4,5]))
# ベクトル のそーと
x = [1,2,4,3,5,7]
sort!(x)
println(x)
sort!(x,rev=true)
println(x)
# ベクトルを逆順にする
x = [1,2,3,4,5]
reverse!(x)
println(x)
|
JuliaLearning/src/cookbook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="m0hTjBtvgQ7-" colab_type="text"
# # Light GBM
# + id="QGDV7PxhgVxm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f298eac0-4c05-4ca7-8255-ff1f6434c579"
from google.colab import drive
drive.mount('/content/drive')
# + id="Mk_rJZZNQct3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="5cdb5f2c-7850-4493-900e-a9691ee666f4"
# !git clone --recursive https://github.com/Microsoft/LightGBM ;
# + id="3nFuDXadYj8o" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="20cb8c69-e018-4257-a0a7-672844a40467"
# %cd /content/LightGBM
# + id="pwHfAPbDYrLC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 256} outputId="8ae4696f-4ca1-4999-c1c2-d04c041f0227"
# !mkdir build
# !cmake -DUSE_GPU=1 #avoid ..
# !make -j$(nproc)
# + id="iRfVto7QYzEg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 109} outputId="af343bf9-09e2-4e59-e60c-ab7c1cab2c72"
# !sudo apt-get -y install python-pip
# + id="XhnbSlZNY6Ag" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 935} outputId="08f0371a-53f9-4a6b-c990-d73b3192dc68"
# !sudo -H pip install setuptools pandas numpy scipy scikit-learn mlflow datetime matplotlib -U
# + id="IXImMmHSZU9P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="e6fb83fe-aa2b-43f9-bf6d-aa2fbaaa57be"
# %cd /content/LightGBM/python-package
# + id="CMQ_m38yZZTs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 605} outputId="8f10f81f-495e-433d-dbe3-f3813ab8661c"
# !sudo python setup.py install --precompile
# + [markdown] id="i6OfvlzjgQ8A" colab_type="text"
# Use tree-based algorithms since they are powerful and do not impose strict assumptions on features like linearity or independence. Light GBM is a fast algorithm with lower memory usage.
# + id="lphSz2l0gQ8E" colab_type="code" colab={}
# #!pip install mlflow
# + id="Xf3T4hGhgQ8R" colab_type="code" colab={}
#conda install numpy
# + [markdown] id="APM4Y5eDgQ8a" colab_type="text"
# Import libraries
# + id="hn9GkhDYgQ8c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="7dc87725-f2e7-4a02-e5fb-8d6060dc5a6b"
import pandas as pd
import numpy as np
from datetime import datetime
from datetime import timedelta
import lightgbm as lgb
from sklearn import preprocessing, metrics
from sklearn.model_selection import ParameterGrid
import matplotlib.pyplot as plt
import seaborn as sns
import mlflow
import mlflow.lightgbm
# + [markdown] id="c3n1XWpWgQ8j" colab_type="text"
# Read data
# + id="q5ZjUiq5gQ8m" colab_type="code" colab={}
data_folder = '/content/drive/My Drive/Kaggle_M5/01_preprocessed_data/'
X_train = pd.read_pickle(data_folder + 'X_train.pkl')
X_val = pd.read_pickle(data_folder + 'X_val.pkl')
X_test = pd.read_pickle(data_folder + 'X_test.pkl')
y_train = X_train['demand']
y_val = X_val['demand']
# + id="XurGzYpHgQ8t" colab_type="code" colab={}
data_folder = '/content/drive/My Drive/Kaggle_M5/00_data/'
submission = pd.read_csv(data_folder + 'sample_submission.csv')
# + [markdown] id="zqDf9jw9gQ82" colab_type="text"
# Set parameters of the light GBM and select features to fit.
# + id="2DI8NAIggQ83" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 109} outputId="60536498-61f4-4f2f-fa5b-1a065741139d"
print(X_train.shape)
print(X_val.shape)
print(X_test.shape)
print(y_train.shape)
print(y_val.shape)
# + id="j6qZ9yWfgQ8_" colab_type="code" colab={}
params = {
'boosting_type': 'gbdt',
'metric': 'rmse',
'objective': 'poisson',
'n_jobs': -1,
'seed': 0,
'learning_rate': 0.1,
'bagging_fraction': 0.75,
'bagging_freq': 10,
'colsample_bytree': 0.75,
'num_iterations':2000,
'device': 'gpu'}
not_features = ['d', 'id', 'item_id', 'demand', 'date', 'start_date']
# + id="hxVO3ABkWTNX" colab_type="code" colab={}
params = {'bagging_fraction': 0.5,
'bagging_freq': 10,
'boosting_type': 'gbdt',
'colsample_bytree': 0.75,
'device': 'gpu',
'early_stopping_round': 500,
'learning_rate': 0.05,
'metric': 'rmse',
'n_jobs': -1,
'num_iterations': 5000,
'objective': 'tweedie',
'seed': 0}
# + id="uBUIAYVZgQ9D" colab_type="code" colab={}
param_grid = {'boosting_type': ['gbdt'],
'metric': ['rmse'],
'objective': ['poisson', 'tweedie'],
'n_jobs': [-1],
'seed': [0],
'learning_rate': [0.05, 0.075, 0.1],
'bagging_fraction': [0.5, 0.75, 1],
'bagging_freq': [10],
'colsample_bytree': [0.75],
'num_iterations': [1000, 5000],
'early_stopping_round': [500],
'device': ['gpu']}
# + id="uuE4wF1ggQ9K" colab_type="code" colab={}
features = X_test.columns[~X_test.columns.isin(not_features)]
# + id="nCXtaSTkgQ9T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 226} outputId="55c8042d-a880-4fa5-8962-ef3df4a5a2ae"
X_train.head()
# + id="kJyt5_WlgQ9c" colab_type="code" colab={}
X_train = X_train[X_train['date'] >= "2015-01-01"]
y_train = X_train['demand']
# + id="Knr52YANgQ9h" colab_type="code" colab={}
train_set = lgb.Dataset(X_train[features], y_train) #, categorical_feature = categorical_features)
val_set = lgb.Dataset(X_val[features], y_val)#, categorical_feature = categorical_features)
# + [markdown] id="rS9QupycgQ9t" colab_type="text"
# Train the model
# + id="unhYXPwd81Nz" colab_type="code" colab={}
#print(X_train.dtypes)
# + id="_NFr11KlgQ9u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 495} outputId="d6af30d9-af86-48ab-a218-252b9f681519"
model = lgb.train(params, train_set,
valid_sets = [train_set, val_set], verbose_eval = 100)
# + id="dBCAEeKvgQ9z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="3051f9ae-5bcf-4288-d143-00309e03fa41"
i = 0
data_folder = '/content/drive/My Drive/Kaggle_M5/04_submissions/'
for g in ParameterGrid(param_grid):
print(i)
print(g)
mlflow.lightgbm.autolog()
model = lgb.train(g, train_set,
valid_sets = [train_set, val_set], verbose_eval = 100)
y_test = model.predict(X_test[features])
X_test['demand'] = y_test
predictions = X_test[['id', 'days_from_start', 'demand']]
predictions = pd.pivot(predictions, index = 'id', columns = 'days_from_start', values = 'demand').reset_index()
predictions.columns = ['id'] + ['F' + str(i + 1) for i in range(28)]
evaluation_rows = [row for row in submission['id'] if 'evaluation' in row]
evaluation = submission[submission['id'].isin(evaluation_rows)]
validation = submission[['id']].merge(predictions, on = 'id')
final = pd.concat([validation, evaluation])
final.head()
final.to_csv(data_folder + 'lightGBM.csv', index = False)
final.to_csv(data_folder + 'lightGBM_{}.csv'.format(i), index = False)
i = i + 1
print("------------------------------------")
# + id="UzmCdHzKgQ96" colab_type="code" colab={} outputId="ff836e5b-fcad-494c-e262-a04f396f788c"
print(len(ParameterGrid(param_grid)))
# + [markdown] id="ZcYY014DgQ9-" colab_type="text"
# Calculate the RMSE on the validation set
# + id="-4TzBLpTgQ9_" colab_type="code" colab={}
y_test = model.predict(X_test[features])
# + id="Yvyp9HKxgQ-Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4b078341-0415-4617-d88c-a7e23a1e11fc"
val_pred = model.predict(X_val[features])
val_score = np.sqrt(metrics.mean_squared_error(val_pred, y_val))
X_val['demand'] = y_val
X_val['demand_pred'] = val_pred
X_val['abs_difference'] = abs(X_val['demand'] - X_val['demand_pred'])
print(f'Our val rmse score is {val_score}')
y_test = model.predict(X_test[features])
X_test['demand'] = y_test
# + id="C_OXP8B1gQ-U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="bd472778-63d9-4852-ba48-cf0e7892b4f3"
print(f'Our val mae score is {metrics.mean_absolute_error(val_pred, y_val)}')
# + [markdown] id="g8n_oM2DgQ-Y" colab_type="text"
# We look at the mean absolute error by forecastablity in order to indestand what category needs more improvement for prediction.
# + id="mDbKpqsHgQ-a" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="9ab96952-7ba9-40ee-c7e1-c82affab8ba4"
X_val.groupby(['demand_type'])['demand_type', 'abs_difference'].agg(['mean']).reset_index()
# + id="e6zQdIgggQ-h" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 313} outputId="ae495d25-fd3a-4aca-9be2-0863faaddb0b"
demans_vs_error = sns.scatterplot(x=X_val['demand'], y=X_val['abs_difference'])
demans_vs_error.set_title('Distribution of error over demand')
demans_vs_error.set_xlabel('absolute error')
# + [markdown] id="WUt3ggb1gQ-k" colab_type="text"
# Surprisingly smooth time series has a large mean absolute error. Erratic time series has a large mean absolute error; typically time series of this type is difficult to predict.
# + [markdown] id="7odHhbd-gQ-n" colab_type="text"
# ## Feature importance
# + id="Rgd_Aga_gQ-n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 731} outputId="84609a19-f05f-422f-ed76-6e27b73bcaf8"
def plotImp(model, X , num = 20):
feature_imp = pd.DataFrame({'Value':model.feature_importance(),'Feature':X.columns})
plt.figure(figsize=(40, 20))
sns.set(font_scale = 5)
sns.barplot(x="Value", y="Feature", data=feature_imp.sort_values(by="Value",
ascending=False)[0:num])
plt.title('LightGBM Features (avg over folds)')
plt.tight_layout()
plt.savefig('lgbm_importances-01.png')
plt.show()
plotImp(model, X_train[features], 30)
# + [markdown] id="hmXGtIragQ-t" colab_type="text"
# 'item_id' and 'id' are most important features. 'days_from_start' reflect the trend of data.
# + [markdown] id="0C3tI6e8gQ-u" colab_type="text"
# ## Transform prediction
# + [markdown] id="Lu30-XekgQ-v" colab_type="text"
# Transform predictions to the right format
# + id="WpJ6AAipgQ-w" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 226} outputId="4bfb216f-2b22-4a72-9a1e-ae34f17cb23e"
predictions = X_test[['id', 'days_from_start', 'demand']]
predictions = pd.pivot(predictions, index = 'id', columns = 'days_from_start', values = 'demand').reset_index()
predictions.columns = ['id'] + ['F' + str(i + 1) for i in range(28)]
evaluation_rows = [row for row in submission['id'] if 'evaluation' in row]
evaluation = submission[submission['id'].isin(evaluation_rows)]
validation = submission[['id']].merge(predictions, on = 'id')
final = pd.concat([validation, evaluation])
final.head()
# + [markdown] id="L5Qe2hsBgQ-1" colab_type="text"
# Save the submission, the features and the score to files.
# + id="-MWsOVyhgQ-3" colab_type="code" colab={}
data_folder = '/content/drive/My Drive/Kaggle_M5/04_submissions/'
final.to_csv(data_folder + 'lightGBM.csv', index = False)
# + id="e6QltxGWgQ-8" colab_type="code" colab={}
_features = X_test.columns[~X_test.columns.isin(not_features)]
features = features.to_list()
features.append(str(val_score))
features.append(str(params['objective']))
# + id="rLSNtwLHgQ_B" colab_type="code" colab={}
with open("../04_submissions/lgb_features_score.txt", "a") as outfile:
outfile.write("\n".join(features))
# + [markdown] id="WjC7Gr7egQ_F" colab_type="text"
# ## Score
# + [markdown] id="ISJsNaWtgQ_H" colab_type="text"
# The score of this submission is 0.58584, which is better than naive prediction.
# + id="KY966IMegQ_I" colab_type="code" colab={}
# + id="Z-biR6DegQ_M" colab_type="code" colab={}
|
03_lightGBM_GPU.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time Series Data Analysis of Selected Low-Cost Funds
# I have downloaded data of 10 different funds from finance.yahoo.com and manually merged their daily closing prices by date starting 01-01-2014. Each of these funds represent different classes of assets which vary in risk. For investors, higher risk can yield more attractive returns; however, if an investor is risk averse they would look to other asset classes to diversify their portfolio and the risk it carries.
#
# The ideal goal of this data analysis would be to understand how much weigh to give to each of these funds on a diversified portfolio in order to achieve a specified risk/return rate.
# #### Importing libraries I will need
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# #### Loading data and understanding what the data looks like
mfunds = pd.read_csv("MFData.csv")
mfunds.head()
# #### Need to clean the data to make it more workable
# +
# creating a new dataframe with 'date' column as index
data = mfunds.set_index('Date')
data.head()
# -
data.plot()
# #### Need to make my visualization more interpretable
# +
plt.style.use('ggplot')
# %pylab inline
pylab.rcParams['figure.figsize'] = (15, 9)
data.plot()
# -
# We can actually learn form the above plot that TLT is slightly counter-cyclical to the general market trends represented by the rest (We will see below that general trend is reflected by VOO when compared to the Dow Jones Index since VOO contains most of the securities that make up the index. In fact, the bottom couple of lines are also a bit difficult to explore so I will omit the higher priced funds to be able to look at them more closely.
columns = ['VGSIX', 'VFITX', 'VGTSX', 'VEIEX', 'VFITX', 'VIPSX', 'IBMH']
df1 = pd.DataFrame(data, columns=columns)
df1.plot()
# It seems that VGSIX does not always convey market trends. We can ignore the flatter lines for IBMH, VPSIX, and VFITX as they contains treasury bonds and assets that are designed to be stable and combat losses to inflation. I will study the correlation of each compared to the market index below. First I will have to retrieve the data for the Dow Jones Market Index and merge it to my current dataset.
# #### Merging the new data for Dow Jones Index to mycurrent dataset
mindex = pd.read_csv("DJI.csv") #get the data
mindex = pd.DataFrame(mindex, columns=['Date','Close']) # selecting the columns I want
mindex.columns= ['Date','DJI'] #Rename the column from Close to DJI
mindex.head()
#Merging mindex with data using pd.concat()
data = data.reset_index(drop=True) #requied to join without encountering a bug that wipes the data
newdata = pd.concat([data,mindex], axis=1)
#newdata = data.join(mindex)
newdata.set_index("Date", inplace=True, drop=True) #setting Date back as the index again
newdata.head()
# #### Correlation study with Dow Jones Market Index
newdata.corr()
# From the above correlation table, we recognize reduced correlation for VGTSX, VEIEX, and VGSIX. We have negative correlation for VFITX, VIPSX, TLT, and IBMH. This confirms our visual analysis from our first plots.
# Will re-raw the correlation table with a color map as reference in case we need to look for insights in the future. Source: https://stackoverflow.com/questions/29432629/plot-correlation-matrix-using-pandas
corr = newdata.corr()
corr.style.background_gradient(cmap='RdBu_r', axis=None).set_precision(3) # 'coolwarm', RdBu_r', & 'BrBG' good color maps
# ### Assessment of returns
# I need to build new tables from the original dataset (excluding the Dow Jones Index) to assess the returns for each investable fund. The return can be calculated by return(t,0) = price(t)/price(0). source: https://ntguardian.wordpress.com/2018/07/17/stock-data-analysis-python-v2/
# df.apply(arg) will apply the function arg to each column in df, and return a DataFrame with the result
# Recall that lambda x is an anonymous function accepting parameter x; in this case, x will be a pandas Series object
mfunds_return = data.apply(lambda x: x / x[0])
mfunds_return.head() - 1
mfunds_return.plot(grid = True).axhline(y = 1, color = "black", lw = 2)
cov = mfunds_return.cov()
cov.style.background_gradient(cmap='RdBu_r', axis=None).set_precision(2) # 'coolwarm', RdBu_r', & 'BrBG' good color maps
|
41fred/MFData.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### LightGBM + ADSTuner: do the ensemble
# * two models, one for causal and the other one for registerd
# * added feature engineering
# * added year, removed temp
# * removing day I got the best results. (The range of days in the train set don't match with test set)
# +
import pandas as pd
import numpy as np
import lightgbm as lgb
# to use ADSTuner
from ads.hpo.search_cv import ADSTuner
from ads.hpo.stopping_criterion import *
from ads.hpo.distributions import *
# to encode categoricals
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import make_scorer
import seaborn as sns
import matplotlib.pyplot as plt
# set seaborn look&feel
sns.set()
import logging
# +
#
# supporting functions
#
def add_features(df):
# feature engineering
df_new = df.copy()
df_new['datetime'] = pd.to_datetime(df_new['datetime'])
# this way I add 2 engineered features
df_new['hour'] = df_new['datetime'].dt.hour
df_new['year'] = df_new['datetime'].dt.year
return df_new
#
# define a custom scorer for ADSTuner
# aligned with the scorer used in Kaggle leaderboard
#
def rmsle(y_pred, y_actual):
diff = np.log(y_pred + 1) - np.log(y_actual + 1)
mean_error = np.square(diff).mean()
return np.sqrt(mean_error)
#
# functions for categorical encoding
#
# first train label encoder
TO_CODE = ['season', 'weather', 'year']
def train_encoders(df):
le_list = []
for col in TO_CODE:
print(f'train for coding: {col} ')
le = LabelEncoder()
le.fit(df[col])
le_list.append(le)
print()
return le_list
# then use it
def apply_encoders(df, le_list):
for i, col in enumerate(TO_CODE):
print(f'Coding: {col} ')
le = le_list[i]
df[col] = le.transform(df[col])
# special treatment for windspeed
# windpeed actually is integer badly rounded !!
print('Coding: windspeed')
df['windspeed'] = np.round(df['windspeed'].values).astype(int)
return df
def show_tuner_results(tuner):
print("ADSTuner session results:")
print(f"ADSTuner has completed {tuner.trials.shape[0]} trials")
print()
print(f"The best trial is the #: {tuner.best_index}")
print(f"Parameters for the best trial are: {tuner.best_params}")
print(f"The metric used to optimize is: {tuner.scoring_name}")
print(f"The best score is: {round(tuner.best_score, 4)}")
# +
# globals and load train dataset
FIGSIZE = (9, 6)
# number of folds for K-fold cv in ADSTuner
FOLDS = 5
# in secs
TIME_BUDGET = 1800
FILE_TRAIN = "train.csv"
FILE_TEST = "test.csv"
# train dataset
data_orig = pd.read_csv(FILE_TRAIN)
# +
#
# add features
#
data_extended = add_features(data_orig)
# have a look
data_extended.tail()
# +
# give a better look at cols with low cardinality
# to decide which one we want to treat as categoricals
THR = 100
# to get cols in alfabetical order
cols = sorted(data_extended.columns)
# changed using list comprehension, to shorten code
cols2 = [col for col in cols if data_extended[col].nunique() < THR]
list_count2 = [
data_extended[col].nunique() for col in cols if data_extended[col].nunique() < THR
]
# plot
plt.figure(figsize=FIGSIZE)
plt.title("Low cardinality features")
ax = sns.barplot(x=cols2, y=list_count2)
# to plot values on bar
ax.bar_label(ax.containers[0])
plt.xticks(rotation=90)
plt.ylabel("# of distinct values")
plt.grid(True)
# +
# ok, we will treat as categorical: holiday, hour, season, weather, windspeed, workingday, year
# +
all_columns = data_extended.columns
# cols to be ignored
# atemp and temp are strongly correlated (0.98) we're taking only one
del_columns = ['datetime', 'temp']
# drop ignored columns
data_used = data_extended.drop(del_columns, axis=1)
# let's code categorical
# windspeed need a special treatment
le_list = train_encoders(data_used)
# coding
data_used = apply_encoders(data_used, le_list)
# +
cat_cols = ['season', 'holiday','workingday', 'weather', 'hour', 'year']
num_cols = ['atemp', 'humidity', 'windspeed']
target_columns = ['casual', 'registered', 'count']
features = sorted(cat_cols + num_cols)
# define indexes for cat_cols
# # cat boost want indexes
cat_columns_idxs = [i for i, col in enumerate(features) if col in cat_cols]
print('All columns:', len(all_columns))
print('Ignored columns:', len(del_columns))
print('Categorical columns:', len(cat_cols))
print('Numerical columns:', len(num_cols))
print(f'All targets: {len(target_columns)}')
print('All the features', len(features))
# +
# loading the parameters for the two models
import pickle
with open("model1.pkl", "rb") as mode1_file:
params1 = pickle.load(mode1_file)
print(params1)
with open("model2.pkl", "rb") as mode1_file:
params2 = pickle.load(mode1_file)
print(params2)
# +
# %%time
# train model1
TARGET = "registered"
x_train = data_used[features]
y_train = data_used[TARGET]
# train the model with chosen parameters
model1 = lgb.LGBMRegressor(**params1)
model1.fit(x_train, y_train, categorical_feature=cat_columns_idxs)
# +
# %%time
# train model2
TARGET = "casual"
x_train = data_used[features]
y_train = data_used[TARGET]
# train the model with chosen parameters
model2 = lgb.LGBMRegressor(**params2)
model2.fit(x_train, y_train, categorical_feature=cat_columns_idxs)
# +
# now we must combine predictions from model1 (registered) and model2 (causal))
test_orig = pd.read_csv(FILE_TEST)
# add engineered features
# feature engineering
test_orig = add_features(test_orig)
# coding
test_orig = apply_encoders(test_orig, le_list)
# data on which do scoring
x_test = test_orig[features]
# +
# scoring
score_test1 = model1.predict(x_test)
score_test2 = model1.predict(x_test)
# -
score_test = score_test1 + score_test2
# +
df_sub = pd.read_csv("sampleSubmission.csv")
# remove decimals
df_sub["count"] = np.round(score_test, 0)
# remove eventual negative
condition = df_sub["count"] < 0
df_sub.loc[condition, "count"] = 0
# +
FILE_SUB_PREFIX = "sub-test11"
FILE_SUB = FILE_SUB_PREFIX + ".csv"
df_sub.to_csv(FILE_SUB, index=False)
# -
# !kaggle competitions submit -c "bike-sharing-demand" -f $FILE_SUB -m "adstuner, two models"
|
.ipynb_checkpoints/lightgbm-adstuner-casual-registered-ensemble-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # A notebook to Plot results
# ### Seaborn
# The following is based on the Seaborn tutorial: https://seaborn.pydata.org/tutorial/axis_grids.html
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="ticks")
# #### Read in your results
# Select your data file:
df = pd.read_csv('../data/final/IAEE_residential_results.csv')
df.tail()
# #### To plot all economies:
g = sns.FacetGrid(df, col="Economy", hue="Fuel" ,col_wrap=2, height=5, ylim=(0, 8000))
g.map(plt.plot,"Year","Demand", alpha=0.7)
g.add_legend()
# #### To plot one economy in more detail:
economy = '01_AUS'
to_plot = df[df['Economy']==economy]
h = sns.FacetGrid(to_plot, col="Fuel", col_wrap=3, height=3.5, ylim=(0, 8000))
h.map(plt.plot, "Year", "Demand")
# ### Bokeh
from bokeh.plotting import figure, show, output_notebook
from bokeh.layouts import gridplot
output_notebook()
# Make pivot table so columns are individual lines
df_pivot = df.pivot_table(index=['Economy','Year'],columns='Fuel',values='Demand')
df_pivot.head(2)
# #### Line plot
#
# This will loop through economies.
#
# Clicking on legend item will "mute" it.
# +
from bokeh.palettes import Spectral10
figures = []
economies = ['01_AUS','18_CT']
for economy in economies:
q = figure(width=500, height=300,title=economy)
_df = df_pivot.loc[(economy,),]
numlines=len(_df.columns)
for name, color in zip(_df,Spectral10):
q.line(_df.index.values, _df[name], line_width=2, color=color,
alpha=0.7, legend_label=name, muted_color=color, muted_alpha=0.2,)
q.legend.location = "top_left"
q.legend.click_policy="mute"
figures.append(q)
show(gridplot(figures, ncols=2, plot_width=400, plot_height=300))
# -
# #### Stacked area plot
# +
from bokeh.palettes import brewer
figures = []
economies = ['01_AUS','18_CT']
for economy in economies:
_df = df_pivot.loc[(economy,),]
numlines=len(_df.columns)
names = _df.columns.values.tolist()
_df = _df.reset_index()
v = figure(width=500, height=300,title=economy)
v.varea_stack(names, x='Year',
color=brewer['Spectral'][numlines],
alpha=0.7,
muted_color=brewer['Spectral'][numlines],
muted_alpha=0.2,
legend_label=names,
source=_df)
v.legend.location = "top_left"
v.legend.click_policy="mute"
figures.append(v)
show(gridplot(figures, ncols=2, plot_width=400, plot_height=300))
# -
|
notebooks/Plot Results.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .scala
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Apache Toree - Scala
// language: scala
// name: apache_toree_scala
// ---
// # Kmeans over a set of GeoTiffs
//
// This notebook loads a set of GeoTiffs into a **RDD** of Tiles, with each Tile being a band in the GeoTiff. Each GeoTiff file contains [**MODIS MCD12Q2 v005**](https://lpdaac.usgs.gov/dataset_discovery/modis/modis_products_table/mcd12q2) for one year over the entire USA. The data ranges from 2001 until 2014.
//
// Kmeans takes years as dimensions. Hence, the matrix has cells as rows and the years as columns. To cluster on all years, the matrix needs to be transposed. The notebook has two flavors of matrix transpose, locally by the Spark-driver or distributed using the Spark-workers. Once transposed the matrix is converted to a **RDD** of dense vectors to be used by **Kmeans** algorithm from **Spark-MLlib**. The end result is a grid where each cell has a cluster ID which is then saved into a SingleBand GeoTiff. By saving the result into a GeoTiff, the reader can plot it using a Python notebook as the one defined in the [python examples](../examples/python).
//
// <span style="color:red">In this notebook the reader only needs to modify the variables in **Mode of Operation Setup**</span>.
// ## Dependencies
// +
import sys.process._
import java.io.{ByteArrayInputStream, ByteArrayOutputStream, ObjectInputStream, ObjectOutputStream}
import geotrellis.proj4.CRS
import geotrellis.raster.{CellType, ArrayTile, DoubleArrayTile, Tile, UByteCellType}
import geotrellis.raster.io.geotiff._
import geotrellis.raster.io.geotiff.writer.GeoTiffWriter
import geotrellis.raster.io.geotiff.{GeoTiff, SinglebandGeoTiff}
import geotrellis.spark.io.hadoop._
import org.apache.hadoop.io._
import geotrellis.vector.{Extent, ProjectedExtent}
import org.apache.spark.broadcast.Broadcast
import org.apache.spark.mllib.clustering.{KMeans, KMeansModel}
import org.apache.spark.mllib.linalg.distributed.{CoordinateMatrix, MatrixEntry, RowMatrix}
import org.apache.spark.mllib.linalg.{Vector, Vectors}
import org.apache.spark.rdd.RDD
import org.apache.spark.{SparkConf, SparkContext}
import org.apache.hadoop.io.{IOUtils, SequenceFile}
import org.apache.hadoop.io.SequenceFile.Writer
//Spire is a numeric library for Scala which is intended to be generic, fast, and precise.
import spire.syntax.cfor._
// -
// ## Mode of operation
//
// Here the user can define the mode of operation.
// * **rdd_offline_mode**: If false it means the notebook will create all data from scratch and store protected_extent and num_cols_rows into HDFS. Otherwise, these data structures are read from HDFS.
// * **matrix_offline_mode**: If false it means the notebook will create a mtrix, transposed it and save it to HDFS. Otherwise, these data structures are read from HDFS.
// * **kmeans_offline_mode**: If false it means the notebook will train kmeans and run kemans and store kmeans model into HDFS. Otherwise, these data structures are read from HDFS.
//
// It is also possible to define which directory of GeoTiffs is to be used and on which **band** to run Kmeans. The options are
// * **all** which are a multi-band (**8 bands**) GeoTiffs
// * Or choose single band ones:
// 0. Onset_Greenness_Increase
// 1. Onset_Greenness_Maximum
// 2. Onset_Greenness_Decrease
// 3. Onset_Greenness_Minimum
// 4. NBAR_EVI_Onset_Greenness_Minimum
// 5. NBAR_EVI_Onset_Greenness_Maximum
// 6. NBAR_EVI_Area
// 7. Dynamics_QC
//
// For kmeans the user can define the **number of iterations** and **number of clusters** as an inclusive range. Such range is defined using **minClusters**, **maxClusters**, and **stepClusters**. These variables will set a loop starting at **minClusters** and stopping at **maxClusters** (inclusive), iterating **stepClusters** at the time. <span style="color:red">Note that when using a range **kemans offline mode** is not possible and it will be reset to **online mode**</span>.
// ### Mode of Operation setup
// <a id='mode_of_operation_setup'></a>
// +
//Operation mode
var rdd_offline_mode = true
var matrix_offline_mode = true
var kmeans_offline_mode = true
//GeoTiffs to be read from "hdfs:///user/hadoop/modis/"
var dir_path = "hdfs:///user/hadoop/avhrr/"
var offline_dir_path = "hdfs:///user/pheno/avhrr/"
/*
Choose all and then the band or the dir which has the band extracted.
0: Onset_Greenness_Increase
1: Onset_Greenness_Maximum
2: Onset_Greenness_Decrease
3: Onset_Greenness_Minimum
4: NBAR_EVI_Onset_Greenness_Minimum
5: NBAR_EVI_Onset_Greenness_Maximum
6: NBAR_EVI_Area
7: Dynamics_QC
for example:
var geoTiff_dir = "Onset_Greenness_Increase"
var band_num = 0
*/
var geoTiff_dir = "SOST"
var band_num = 0
//Years between (inclusive) 1989 - 2014
var satellite_first_year = 1989
var satellite_last_year = 2014
//Mask
val toBeMasked = true
val mask_path = "hdfs:///user/hadoop/usa_mask.tif"
//Kmeans number of iterations and clusters
var numIterations = 75
var minClusters = 100
var maxClusters = 100
var stepClusters = 10
var save_kmeans_model = false
// -
//
// <span style="color:red">DON'T MODIFY ANY PIECE OF CODE FROM HERE ON!!!</span>.
//
// ### Mode of operation validation
// +
//Validation, do not modify these lines.
var single_band = false
if (geoTiff_dir == "all") {
single_band = false
} else {
single_band = true
if (band_num > 0) {
println("Since it is single band, we will use band 0!!!")
band_num = 0
}
}
if (minClusters > maxClusters) {
maxClusters = minClusters
stepClusters = 1
}
if (stepClusters < 1) {
stepClusters = 1
}
//Paths to store data structures for Offline runs
var mask_str = ""
if (toBeMasked)
mask_str = "_mask"
var grid0_path = offline_dir_path + geoTiff_dir + "/grid0" + "_"+ band_num + mask_str
var grid0_index_path = offline_dir_path + geoTiff_dir + "/grid0_index" + "_"+ band_num + mask_str
var grids_noNaN_path = offline_dir_path + geoTiff_dir + "/grids_noNaN" + "_"+ band_num + mask_str
var metadata_path = offline_dir_path + geoTiff_dir + "/metadata" + "_"+ band_num + mask_str
var grids_matrix_path = offline_dir_path + geoTiff_dir + "/grids_matrix" + "_"+ band_num + mask_str
//Check offline modes
var conf = sc.hadoopConfiguration
var fs = org.apache.hadoop.fs.FileSystem.get(conf)
val rdd_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(grid0_path))
val matrix_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(grids_matrix_path))
if (rdd_offline_mode != rdd_offline_exists) {
println("\"Load GeoTiffs\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + rdd_offline_exists.toString())
rdd_offline_mode = rdd_offline_exists
}
if (matrix_offline_mode != matrix_offline_exists) {
println("\"Matrix\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + matrix_offline_exists.toString())
matrix_offline_mode = matrix_offline_exists
}
if (!fs.exists(new org.apache.hadoop.fs.Path(mask_path))) {
println("The mask path: " + mask_path + " is invalid!!!")
}
//Years
val satellite_years = 1989 to 2014
if (!satellite_years.contains(satellite_first_year) || !(satellite_years.contains(satellite_last_year))) {
println("Invalid range of years for " + geoTiff_dir + ". I should be between " + satellite_first_year + " and " + satellite_last_year)
System.exit(0)
}
var satellite_years_range = (satellite_years.indexOf(satellite_first_year), satellite_years.indexOf(satellite_last_year))
var num_kmeans :Int = 1
if (minClusters != maxClusters) {
num_kmeans = ((maxClusters - minClusters) / stepClusters) + 1
}
println(num_kmeans)
var kmeans_model_paths :Array[String] = Array.fill[String](num_kmeans)("")
var wssse_path :String = offline_dir_path + geoTiff_dir + "/" + numIterations + "_wssse"
var geotiff_hdfs_paths :Array[String] = Array.fill[String](num_kmeans)("")
var geotiff_tmp_paths :Array[String] = Array.fill[String](num_kmeans)("")
if (num_kmeans > 1) {
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
kmeans_model_paths(numClusters_id) = offline_dir_path + geoTiff_dir + "/kmeans_model_" + band_num + "_" + numClusters + "_" + numIterations
//Check if the file exists
val kmeans_exist = fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))
if (kmeans_exist && !kmeans_offline_mode) {
println("The kmeans model path " + kmeans_model_paths(numClusters_id) + " exists, please remove it.")
} else if (!kmeans_exist && kmeans_offline_mode) {
kmeans_offline_mode = false
}
geotiff_hdfs_paths(numClusters_id) = offline_dir_path + geoTiff_dir + "/clusters_" + band_num + "_" + numClusters + "_" + numIterations + ".tif"
geotiff_tmp_paths(numClusters_id) = "/tmp/clusters_" + band_num + "_" + geoTiff_dir + "_" + numClusters + "_" + numIterations + ".tif"
if (fs.exists(new org.apache.hadoop.fs.Path(geotiff_hdfs_paths(numClusters_id)))) {
println("There is already a GeoTiff with the path: " + geotiff_hdfs_paths(numClusters_id) + ". Please make either a copy or move it to another location, otherwise, it will be over-written.")
}
numClusters_id += 1
}
kmeans_offline_mode = false
} else {
kmeans_model_paths(0) = offline_dir_path + geoTiff_dir + "/kmeans_model_" + band_num + "_" + minClusters + "_" + numIterations
val kmeans_offline_exists = fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(0)))
if (kmeans_offline_mode != kmeans_offline_exists) {
println("\"Kmeans\" offline mode is not set properly, i.e., either it was set to false and the required file does not exist or vice-versa. We will reset it to " + kmeans_offline_exists.toString())
kmeans_offline_mode = kmeans_offline_exists
}
geotiff_hdfs_paths(0) = offline_dir_path + geoTiff_dir + "/clusters_" + band_num + "_" + minClusters + "_" + numIterations + ".tif"
geotiff_tmp_paths(0) = "/tmp/clusters_" + band_num + "_" + geoTiff_dir + "_" + minClusters + "_" + numIterations + ".tif"
if (fs.exists(new org.apache.hadoop.fs.Path(geotiff_hdfs_paths(0)))) {
println("There is already a GeoTiff with the path: " + geotiff_hdfs_paths(0) + ". Please make either a copy or move it to another location, otherwise, it will be over-written.")
}
}
// -
// ## Functions to (de)serialize any structure into Array[Byte]
// +
def serialize(value: Any): Array[Byte] = {
val out_stream: ByteArrayOutputStream = new ByteArrayOutputStream()
val obj_out_stream = new ObjectOutputStream(out_stream)
obj_out_stream.writeObject(value)
obj_out_stream.close
out_stream.toByteArray
}
def deserialize(bytes: Array[Byte]): Any = {
val obj_in_stream = new ObjectInputStream(new ByteArrayInputStream(bytes))
val value = obj_in_stream.readObject
obj_in_stream.close
value
}
// -
// ## Load GeoTiffs
//
// Using GeoTrellis all GeoTiffs of a directory will be loaded into a RDD. Using the RDD, we extract a grid from the first file to lated store the Kmeans cluster_IDS, we build an Index for populate such grid and we filter out here all NaN values.
// +
val t0 = System.nanoTime()
//Global variables
var projected_extent = new ProjectedExtent(new Extent(0,0,0,0), CRS.fromName("EPSG:3857"))
var grid0: RDD[(Long, Double)] = sc.emptyRDD
var grid0_index: RDD[Long] = sc.emptyRDD
var grids_noNaN_RDD: RDD[Array[Double]] = sc.emptyRDD
var num_cols_rows :(Int, Int) = (0, 0)
var cellT :CellType = UByteCellType
var grids_RDD :RDD[Array[Double]] = sc.emptyRDD
var mask_tile0 :Tile = new SinglebandGeoTiff(geotrellis.raster.ArrayTile.empty(cellT, num_cols_rows._1, num_cols_rows._2), projected_extent.extent, projected_extent.crs, Tags.empty, GeoTiffOptions.DEFAULT).tile
var grid_cells_size :Long = 0
//Load Mask
if (toBeMasked) {
val mask_tiles_RDD = sc.hadoopGeoTiffRDD(mask_path).values
val mask_tiles_withIndex = mask_tiles_RDD.zipWithIndex().map{case (e,v) => (v,e)}
mask_tile0 = (mask_tiles_withIndex.filter(m => m._1==0).values.collect())(0)
}
//Local variables
val pattern: String = "tif"
val filepath: String = dir_path + geoTiff_dir
if (rdd_offline_mode) {
grids_noNaN_RDD = sc.objectFile(grids_noNaN_path)
grid0 = sc.objectFile(grid0_path)
grid0_index = sc.objectFile(grid0_index_path)
val metadata = sc.sequenceFile(metadata_path, classOf[IntWritable], classOf[BytesWritable]).map(_._2.copyBytes()).collect()
projected_extent = deserialize(metadata(0)).asInstanceOf[ProjectedExtent]
num_cols_rows = (deserialize(metadata(1)).asInstanceOf[Int], deserialize(metadata(2)).asInstanceOf[Int])
cellT = deserialize(metadata(3)).asInstanceOf[CellType]
} else {
if (single_band) {
//Lets load a Singleband GeoTiffs and return RDD just with the tiles.
var tiles_RDD :RDD[Tile] = sc.hadoopGeoTiffRDD(filepath, pattern).values
//Retrive the numbre of cols and rows of the Tile's grid
val tiles_withIndex = tiles_RDD.zipWithIndex().map{case (e,v) => (v,e)}
val tile0 = (tiles_withIndex.filter(m => m._1==0).values.collect())(0)
num_cols_rows = (tile0.cols,tile0.rows)
cellT = tile0.cellType
if (toBeMasked) {
val mask_tile_broad :Broadcast[Tile] = sc.broadcast(mask_tile0)
grids_RDD = tiles_RDD.map(m => m.localInverseMask(mask_tile_broad.value, 1, -1000).toArrayDouble())
} else {
grids_RDD = tiles_RDD.map(m => m.toArrayDouble())
}
} else {
//Lets load Multiband GeoTiffs and return RDD just with the tiles.
val tiles_RDD = sc.hadoopMultibandGeoTiffRDD(filepath, pattern).values
//Retrive the numbre of cols and rows of the Tile's grid
val tiles_withIndex = tiles_RDD.zipWithIndex().map{case (e,v) => (v,e)}
val tile0 = (tiles_withIndex.filter(m => m._1==0).values.collect())(0)
num_cols_rows = (tile0.cols,tile0.rows)
cellT = tile0.cellType
//Lets read the average of the Spring-Index which is stored in the 4th band
val band_numB :Broadcast[Int] = sc.broadcast(band_num)
if (toBeMasked) {
val mask_tile_broad :Broadcast[Tile] = sc.broadcast(mask_tile0)
grids_RDD = tiles_RDD.map(m => m.band(band_numB.value).localInverseMask(mask_tile_broad.value, 1, -1000).toArrayDouble())
} else {
grids_RDD = tiles_RDD.map(m => m.band(band_numB.value).toArrayDouble())
}
}
//Retrieve the ProjectExtent which contains metadata such as CRS and bounding box
val projected_extents_withIndex = sc.hadoopGeoTiffRDD(filepath, pattern).keys.zipWithIndex().map{case (e,v) => (v,e)}
projected_extent = (projected_extents_withIndex.filter(m => m._1 == 0).values.collect())(0)
//Get Index for each Cell
val grids_withIndex = grids_RDD.zipWithIndex().map { case (e, v) => (v, e) }
if (toBeMasked) {
grid0_index = grids_withIndex.filter(m => m._1 == 0).values.flatMap(m => m).zipWithIndex.filter(m => m._1 != -1000.0).map { case (v, i) => (i) }
} else {
grid0_index = grids_withIndex.filter(m => m._1 == 0).values.flatMap(m => m).zipWithIndex.map { case (v, i) => (i) }
}
//Get the Tile's grid
grid0 = grids_withIndex.filter(m => m._1 == 0).values.flatMap( m => m).zipWithIndex.map{case (v,i) => (i,v)}
//Lets filter out NaN
if (toBeMasked) {
grids_noNaN_RDD = grids_RDD.map(m => m.filter(m => m != -1000.0))
} else {
grids_noNaN_RDD = grids_RDD
}
//Store data in HDFS
grid0.saveAsObjectFile(grid0_path)
grid0_index.saveAsObjectFile(grid0_index_path)
grids_noNaN_RDD.saveAsObjectFile(grids_noNaN_path)
val grids_noNaN_RDD_withIndex = grids_noNaN_RDD.zipWithIndex().map { case (e, v) => (v, e) }
grids_noNaN_RDD = grids_noNaN_RDD_withIndex.filterByRange(satellite_years_range._1, satellite_years_range._2).values
val writer: SequenceFile.Writer = SequenceFile.createWriter(conf,
Writer.file(metadata_path),
Writer.keyClass(classOf[IntWritable]),
Writer.valueClass(classOf[BytesWritable])
)
writer.append(new IntWritable(1), new BytesWritable(serialize(projected_extent)))
writer.append(new IntWritable(2), new BytesWritable(serialize(num_cols_rows._1)))
writer.append(new IntWritable(3), new BytesWritable(serialize(num_cols_rows._2)))
writer.append(new IntWritable(4), new BytesWritable(serialize(cellT)))
writer.hflush()
writer.close()
}
grid_cells_size = grid0_index.count().toInt
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// ## Matrix
//
// We need to do a Matrix transpose to have clusters per cell and not per year. With a GeoTiff representing a single year, the loaded data looks liks this:
// ```
// bands_RDD.map(s => Vectors.dense(s)).cache()
//
// //The vectors are rows and therefore the matrix will look like this:
// [
// Vectors.dense(0.0, 1.0, 2.0),
// Vectors.dense(3.0, 4.0, 5.0),
// Vectors.dense(6.0, 7.0, 8.0),
// Vectors.dense(9.0, 0.0, 1.0)
// ]
// ```
//
// To achieve that we convert the **RDD[Vector]** into a distributed Matrix, a [**CoordinateMatrix**](https://spark.apache.org/docs/latest/mllib-data-types.html#coordinatematrix), which as a **transpose** method.
// +
val t0 = System.nanoTime()
//Global variables
var grids_matrix: RDD[Vector] = sc.emptyRDD
val grid_cells_sizeB = sc.broadcast(grid_cells_size)
if (matrix_offline_mode) {
grids_matrix = sc.objectFile(grids_matrix_path)
} else {
//Dense Vector
//val mat :RowMatrix = new RowMatrix(grids_noNaN_RDD.map(m => Vectors.dense(m)))
//Sparse Vector
val mat :RowMatrix = new RowMatrix(grids_noNaN_RDD.map(m => m.zipWithIndex).map(m => m.filter(!_._1.isNaN)).map(m => Vectors.sparse(grid_cells_sizeB.value.toInt, m.map(v => v._2), m.map(v => v._1)))) // Split the matrix into one number per line.
val byColumnAndRow = mat.rows.zipWithIndex.map {
case (row, rowIndex) => row.toArray.zipWithIndex.map {
case (number, columnIndex) => new MatrixEntry(rowIndex, columnIndex, number)
}
}.flatMap(x => x)
val matt: CoordinateMatrix = new CoordinateMatrix(byColumnAndRow)
val matt_T = matt.transpose()
//grids_matrix = matt_T.toRowMatrix().rows
grids_matrix = matt_T.toIndexedRowMatrix().rows.sortBy(_.index).map(_.vector)
grids_matrix.saveAsObjectFile(grids_matrix_path)
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// ## Kmeans
//
// We use Kmeans from Sparl-MLlib. The user should only modify the variables on Kmeans setup.
// ### Kmeans Training
// +
val t0 = System.nanoTime()
//Global variables
var kmeans_models :Array[KMeansModel] = new Array[KMeansModel](num_kmeans)
var wssse_data :List[(Int, Int, Double)] = List.empty
if (kmeans_offline_mode) {
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
if (!fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))) {
println("One of the files does not exist, we will abort!!!")
System.exit(0)
} else {
kmeans_models(numClusters_id) = KMeansModel.load(sc, kmeans_model_paths(numClusters_id))
}
numClusters_id += 1
}
val wssse_data_RDD :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)
wssse_data = wssse_data_RDD.collect().toList
} else {
var numClusters_id = 0
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
val wssse_data_RDD :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)
wssse_data = wssse_data_RDD.collect().toList
}
grids_matrix.cache()
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
println(numClusters)
kmeans_models(numClusters_id) = {
KMeans.train(grids_matrix, numClusters, numIterations)
}
// Evaluate clustering by computing Within Set Sum of Squared Errors
val WSSSE = kmeans_models(numClusters_id).computeCost(grids_matrix)
println("Within Set Sum of Squared Errors = " + WSSSE)
wssse_data = wssse_data :+ (numClusters, numIterations, WSSSE)
//Save kmeans model
if (save_kmeans_model) {
if (!fs.exists(new org.apache.hadoop.fs.Path(kmeans_model_paths(numClusters_id)))) {
kmeans_models(numClusters_id).save(sc, kmeans_model_paths(numClusters_id))
}
}
numClusters_id += 1
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
println("We will delete the wssse file")
try { fs.delete(new org.apache.hadoop.fs.Path(wssse_path), true) } catch { case _ : Throwable => { } }
}
println("Lets create it with the new data")
sc.parallelize(wssse_data, 1).saveAsObjectFile(wssse_path)
}
//Un-persist it to save memory
grids_matrix.unpersist()
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// ### Inspect WSSSE
// +
val t0 = System.nanoTime()
//from disk
if (fs.exists(new org.apache.hadoop.fs.Path(wssse_path))) {
var wssse_data_tmp :RDD[(Int, Int, Double)] = sc.objectFile(wssse_path)//.collect()//.toList
println(wssse_data_tmp.collect().toList)
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// ### Run Kmeans clustering
//
// Run Kmeans and obtain the clusters per each cell.
// +
val t0 = System.nanoTime()
//Cache it so kmeans is more efficient
grids_matrix.cache()
var kmeans_res: Array[RDD[Int]] = Array.fill(num_kmeans)(sc.emptyRDD)
var kmeans_centroids: Array[Array[Double]] = Array.fill(num_kmeans)(Array.emptyDoubleArray)
var numClusters_id = 0
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
kmeans_res(numClusters_id) = kmeans_models(numClusters_id).predict(grids_matrix)
kmeans_centroids(numClusters_id) = kmeans_models(numClusters_id).clusterCenters.map(m => m(0))
numClusters_id += 1
}
//Un-persist it to save memory
grids_matrix.unpersist()
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// #### Sanity test
//
// It can be skipped, it only shows the cluster ID for the first 50 cells
// +
val t0 = System.nanoTime()
val kmeans_res_out = kmeans_res(0).filter(_!= 0).filter(_!=1).take(150)
kmeans_res_out.foreach(print)
println(kmeans_res_out.size)
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// -
// ## Build GeoTiff with Kmeans cluster_IDs
//
// The Grid with the cluster IDs is stored in a SingleBand GeoTiff and uploaded to HDFS.
// ### Assign cluster ID to each grid cell and save the grid as SingleBand GeoTiff
//
// To assign the clusterID to each grid cell it is necessary to get the indices of gird cells they belong to. The process is not straight forward because the ArrayDouble used for the creation of each dense Vector does not contain the NaN values, therefore there is not a direct between the indices in the Tile's grid and the ones in **kmeans_res** (kmeans result).
//
// To join the two RDDS the knowledge was obtaing from a stackoverflow post on [how to perform basic joins of two rdd tables in spark using python](https://stackoverflow.com/questions/31257077/how-do-you-perform-basic-joins-of-two-rdd-tables-in-spark-using-python).
val t0 = System.nanoTime()
var numClusters_id = 0
val grid0_index_I = grid0_index.zipWithIndex().map{ case (v,i) => (i,v)}
cfor(minClusters)(_ <= maxClusters, _ + stepClusters) { numClusters =>
//Merge two RDDs, one containing the clusters_ID indices and the other one the indices of a Tile's grid cells
val cluster_cell_pos = ((kmeans_res(numClusters_id).zipWithIndex().map{ case (v,i) => (i,v)}).join(grid0_index_I).map{ case (k,(v,i)) => (v,i)})
//Associate a Cluster_IDs to respective Grid_cell
val grid_clusters = grid0.map{ case (i, v) => if (v == 0.0) (i,Double.NaN) else (i,v)}.leftOuterJoin(cluster_cell_pos.map{ case (c,i) => (i.toLong, c)})
//Convert all None to NaN
val grid_clusters_res = grid_clusters.sortByKey(true).map{case (k, (v, c)) => if (c == None) (k, Int.MaxValue) else (k, c.get)}
//Define a Tile
val cluster_cellsID :Array[Int] = grid_clusters_res.values.collect()
var cluster_cells :Array[Double] = Array.fill(cluster_cellsID.length)(Double.NaN)
cfor(0)(_ < cluster_cellsID.size, _ + 1) { cellID =>
if (cluster_cellsID(cellID) != Int.MaxValue) {
cluster_cells(cellID) = kmeans_centroids(numClusters_id)(cluster_cellsID(cellID))
}
}
/*
//Convert all None to NaN
val grid_clusters_res = grid_clusters.sortByKey(true).map{case (k, (v, c)) => if (c == None) (k, Double.NaN) else (k, c.get.toDouble)}
//Define a Tile
val cluster_cells :Array[Double] = grid_clusters_res.values.collect()
*/
val cluster_cellsD = DoubleArrayTile(cluster_cells, num_cols_rows._1, num_cols_rows._2)
val geoTif = new SinglebandGeoTiff(cluster_cellsD, projected_extent.extent, projected_extent.crs, Tags.empty, GeoTiffOptions(compression.DeflateCompression))
//Save to /tmp/
GeoTiffWriter.write(geoTif, geotiff_tmp_paths(numClusters_id))
//Upload to HDFS
var cmd = "hadoop dfs -copyFromLocal -f " + geotiff_tmp_paths(numClusters_id) + " " + geotiff_hdfs_paths(numClusters_id)
println(cmd)
Process(cmd)!
//Remove from /tmp/
cmd = "rm -fr " + geotiff_tmp_paths(numClusters_id)
println(cmd)
Process(cmd)!
numClusters_id += 1
}
val t1 = System.nanoTime()
println("Elapsed time: " + (t1 - t0) + "ns")
// # [Visualize results](plot_kmeans_clusters.ipynb) --------------- [Plot WSSE](kmeans_wsse.ipynb)
|
applications/notebooks/stable/kmeans_satellite.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/fangyihaoo/beginners-pytorch-deep-learning/blob/master/pytorch_book_ian.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="g5wJzTg2F9mP" outputId="c078194a-0f77-4088-fe3c-1a96c0ac5f88" colab={"base_uri": "https://localhost:8080/"}
# !git clone https://github.com/fangyihaoo/beginners-pytorch-deep-learning.git
# !pip3 install -r /content/beginners-pytorch-deep-learning/requirements.txt
# + id="cPVVYXHxTNuf" outputId="c249fee0-3158-4464-c0d4-4d97d054c56e" colab={"base_uri": "https://localhost:8080/"}
import time
time.sleep(5)
print(time.ctime())
|
pytorch_book_ian.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="peQDo0VaZbGn"
# # Clase 10: Introducción a ML
#
# - ¿Qué es Machine Learning?
# - ¿Dónde se puede utilizar?
# - Divisiones de Machine Learning
# - Primer algoritmo: regresión lineal
# + [markdown] id="j9wYCr3UZ4_4"
# ## ¿Qué es Machine Learning?
# + [markdown] id="IS9E-HItPMwr"
# >"In particular, we define machine learning as a set of methods that can automatically detect patterns in data, and then use the uncovered patterns to predict future data, or to perform other kinds of decision making under uncertainty." (Murphy, 2012)
# + [markdown] id="P5GbRerOZ9L0"
# ## Tipos de Machine Learning
#
# El machine learning suele dividirse en dos grupos: supervisado y no supervisado.
#
# ### Aprendizaje supervisado
#
# Desde esta aproximación, se tiene
#
# $$D = \{(\textbf{x}_i,y_i)\}_{i=1}^N$$
#
# Donde $D$ es el conjunto de entrenamiento, $\textbf{x}$ son los features o variables (inputs) y $y$ es el output. N es el tamaño del conjunto de entrenamiento. El aprendizaje supervisado se emplea cuando se conocen los valores de $y$. Cuando este output tiene valores continuos, se denomina problema de regresión. Cuando es de tipo categórico, se denomina problema de clasificación.
#
# |Regresión|Clasificación|
# |----|----|
# |Salario|Trabaja o no trabaja|
# |Valor de la vivienda|Tipo de vivienda|
# |Demanda de transporte|Tipo de transporte|
#
#
#
# ### Aprendizaje no supervisado
#
# Por el lado del aprendizaje no supervisado se tiene
# $$D = \{\textbf{x}_i\}^N_{i=1}$$
#
# cuyo objetivo es encontrar o descubrir patrones en los datos.
#
#
# Al final, sin importar el algoritmo, el objetivo es descifrar comportamientos en los datos para que el computador aprenda de ellos y pueda entregar predicciones óptimas para la toma de decisiones.
# + [markdown] id="UxEHt_r9SpvR"
# ##Aspectos importantes iniciales
#
# - Overfitting vs. underfitting
#
# 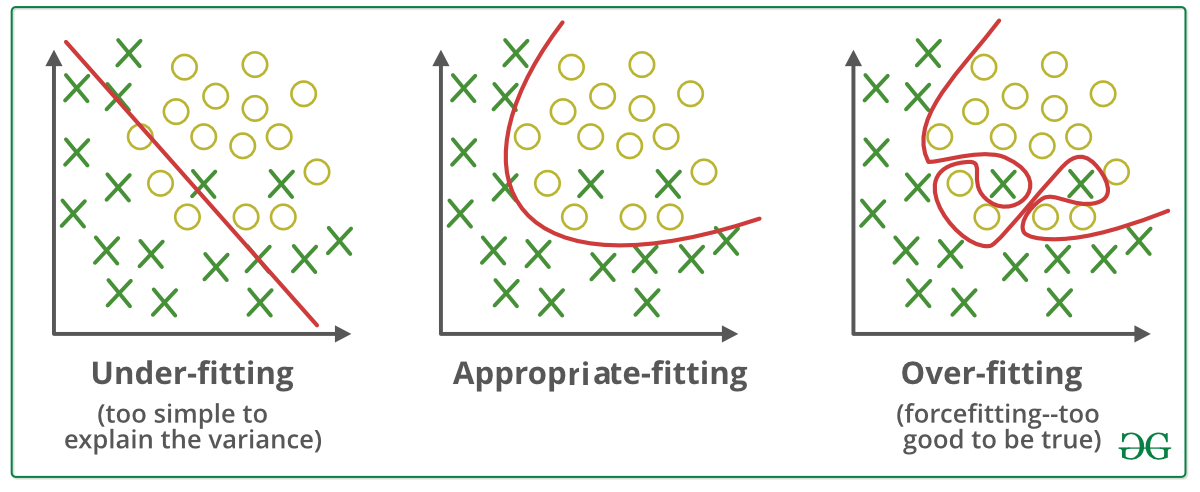
#
# - Varianza vs. sesgo
#
# 
#
#
#
#
# + [markdown] id="XcElvzoPZ_nU"
# ## Primer algoritmo: regresión lineal
# + id="M00N9iFlPmq0" colab={"base_uri": "https://localhost:8080/", "height": 609} outputId="ce78aa20-0e40-4b7e-ccb9-2f19753c174e"
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
a = np.linspace(0,100, 50)
e = np.random.normal(0,100,50)
f = 5 + 10*a + e
plt.figure(figsize=(20,10))
plt.scatter(a,f)
# + [markdown] id="TGNx1V3oSkcy"
#
#
# #### Función de pérdida
#
# $$J(\beta_0, \beta_1) =\sum_i^n (\hat{y}_i-y_i)^2$$
#
# $$J(\beta_0, \beta_1) =\sum_{i=1}^n [(\beta_0+\beta_1x_i)-y_i]^2$$
#
# #### Optimización con dos variables
#
# $$\min_{\beta_0,\beta_1} J(\beta_0, \beta_1)$$
#
# Encontramos el vector gradiente de la función de pérdida
#
# $$\nabla J(\beta_0, \beta_1) =
# \begin{bmatrix}
# \frac{\partial J}{\partial \beta_0} \\
# \frac{\partial J}{\partial \beta_1}
# \end{bmatrix}=\textbf{0}$$
#
# $$\frac{\partial J}{\partial \beta_0}=2\sum_{i=1}^n [(\beta_0+\beta_1x_i)-y_i]$$
#
# $$\frac{\partial J}{\partial \beta_1}=2\sum_{i=1}^n [(\beta_0+\beta_1x_i)-y_i]\cdot x_i$$
#
#
# $$\nabla J(\beta_0, \beta_1) =
# \begin{bmatrix}
# 2\sum_{i=1}^n [(\beta_0+\beta_1x_i)-y_i] \\
# 2\sum_{i=1}^n [(\beta_0+\beta_1x_i)-y_i]\cdot x_i
# \end{bmatrix}=\textbf{0}$$
#
# Al resolver el sistema de ecuaciones 2x2, encontramos el valor de los betas $\hat{\beta}_0,\hat{\beta}_1$
#
#
# #### Optimización con varias variables
#
# $$\min_{\textbf{b}} J(\textbf{b})$$
#
# $$\nabla J(\textbf{b})=\begin{bmatrix}
# \frac{\partial J}{\partial \beta_0} \\
# \frac{\partial J}{\partial \beta_1}\\
# \vdots \\
# \frac{\partial J}{\partial \beta_k}
# \end{bmatrix}=\begin{bmatrix}
# 2\sum_{i=1}^n [h(\textbf{b})-y_i]\cdot x_0^{(i)} \\
# 2\sum_{i=1}^n [h(\textbf{b})-y_i]\cdot x_1^{(i)}\\
# \vdots \\
# 2\sum_{i=1}^n [h(\textbf{b})-y_i]\cdot x_k^{(i)}
# \end{bmatrix}=\textbf{0}$$
#
# #### Optimización con ecuaciones normales
#
# $$\textbf{y} = \textbf{X}\textbf{b}$$
#
# $$\begin{bmatrix}y_i\\\vdots\\ y_n\end{bmatrix}_{n\cdot1}=\begin{bmatrix}
# 1 & x_{1i} & x_{2i} & \ldots & x_{ki} \\
# \vdots & \vdots & \vdots & \ddots & \vdots \\
# 1 & x_{1n} & x_{2n} & \ldots & x_{kn}
# \end{bmatrix}_{n\cdot k+1}\begin{bmatrix}\beta_0\\ \vdots \\ \beta_k\end{bmatrix}_{k+1\cdot 1}$$
#
# $$\min_{\textbf{b}} \textbf{e}'\textbf{e}$$
#
# $$\min_{\textbf{b}} (\textbf{X}\textbf{b}-\textbf{y})'(\textbf{X}\textbf{b}-\textbf{y})$$
#
# $$\textbf{b}=(\textbf{X}'\textbf{X})^{-1}\textbf{X}'\textbf{y}$$
# + [markdown] id="-fNMypwIYasn"
# ¡Esto es computacionalmente costoso!
#
# ### Aparece el descenso del gradiente
#
# Si quiero optimizar una función que depende de una variable
#
# $$\min_{x} f(x)$$
#
# derivo e igualo a 0
#
# $$f'(x)=0$$
#
# Si quiero optimizar una función que depende de varias variables, encuentro el vector gradiente
#
# $$\min_{\textbf{x}} f(\textbf{x})$$
#
# y lo igualo a un vector nulo
#
# $$\nabla f (\textbf{x})=\textbf{0}$$
#
# Ambos procesos me permitirán encontrar los puntos críticos que minimizan la función. Puedo llegar al mismo resultado a través de un método numérico: el descenso del gradiente. El vector gradiente me entrega la dirección en la cual la función cambia más rápido. Puedo aprovechar esto para poco a poco ir acercándome al punto donde ya no hay cambio, el extremo.
#
# Suponga la función $f(x) = x^2$ que por definición es convexa. Si se plantea el problema
#
# $$\min_{x} x^2$$
#
# derivo la función e igualo a 0. El punto crítico es 0.
#
# $$x^*=0$$
#
# + id="iEpM4_9mFupR"
import numpy as np
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="sxgFsnS1SkPq" outputId="e96d6937-8491-4b62-beae-afbd4e0bab90"
x = np.linspace(-5,5,100)
y = x**2
plt.plot(x,y)
# + [markdown] id="z9VDOTGBF0hK"
# El punto donde la función encuentra el mínimo es cuando
#
# $$x=0$$
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="1GFMDPgTF5MA" outputId="6915deaf-e026-475a-ee34-a232d639d3fa"
x = np.linspace(-5,5,100)
y = x**2
plt.plot(x,y)
plt.axhline(0,alpha=0.2,color='k', ls='--')
plt.axvline(0,alpha=0.2,color='k', ls='--')
# + [markdown] id="dglPGTPJGJVO"
# Con el descenso del gradiente, puedo llegar a la misma respuesta pero en lugar de ser un proceso analítico es un método numérico. El descenso del gradiente consta de tres partes:
#
# - Punto de partida
# - Tasa de aprendizaje o learning rate
# - El gradiente de la función que quiere optimizarse
#
# A través de un proceso iterativo se busca un punto de convergencia con la siguiente operación:
#
#
# $$x_{min}:= x_{min} - r\cdot\nabla f(x)$$
#
# Donde $r$ es la tasa de aprendizaje y puede optimizarse o seleccionarse un número arbitrario, usualmente de $0.01$ o inferior.
#
# Para el ejercicio anteriormente planteado, se debe iterar sobre la siguiente operación
#
# $$x_{min} := x_{min} - r\cdot 2x$$
# + colab={"base_uri": "https://localhost:8080/"} id="NawQB3EHGIjr" outputId="13d618f7-0183-4aa2-aaba-c54e69b89f2e"
grad = lambda x: 2*x #creamos el gradiente para la función
x = -1 #generamos un número aleatorio
r = 0.1 #seleccionamos una tasa de aprendizaje
X = []
for _ in range(1000):
X.append(x)
x = x - r * grad(x)
print(x)
# + [markdown] id="mzGzqmcFHur2"
# Podemos ver el comportamiento gráficamente
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="3V2w-qN5HoAt" outputId="0f8536b7-1151-4246-8bc1-b5f40eae6e79"
vec = np.linspace(-2,2,100)
y = vec**2
y_ = [i**2 for i in X]
plt.plot(vec,y)
plt.plot(X, y_, 'o-')
plt.scatter(x,x**2)
plt.axhline(0,alpha=0.2,color='k', ls='--')
plt.axvline(0,alpha=0.2,color='k', ls='--')
# + [markdown] id="vSvHxdQiIlAW"
# Si la tasa de aprendizaje es muy grande o muy pequeña, no convergerá fácilmente.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="DGWyigifItq7" outputId="7021f25d-4fc7-42bb-9ce5-af8a5591257f"
grad = lambda x: 2*x #creamos el gradiente para la función
x = -2#generamos un número aleatorio
r = 0.8 #seleccionamos una tasa de aprendizaje
X = []
for _ in range(1000):
X.append(x)
x = x - r * grad(x)
vec = np.linspace(-2,2,100)
y = vec**2
y_ = [i**2 for i in X]
plt.plot(vec,y)
plt.plot(X, y_, 'o-')
plt.scatter(x,x**2)
plt.axhline(0,alpha=0.2,color='k', ls='--')
plt.axvline(0,alpha=0.2,color='k', ls='--')
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="c7kbJGCwI1P7" outputId="00b010de-358b-41eb-e1d0-21a4ce45e699"
grad = lambda x: 2*x #creamos el gradiente para la función
x = 4#generamos un número aleatorio
r = 0.01 #seleccionamos una tasa de aprendizaje
X = []
for _ in range(1000):
X.append(x)
x = x - r * grad(x)
vec = np.linspace(-2,2,100)
y = vec**2
y_ = [i**2 for i in X]
plt.plot(vec,y)
plt.plot(X, y_, 'o-')
plt.scatter(x,x**2)
plt.axhline(0,alpha=0.2,color='k', ls='--')
plt.axvline(0,alpha=0.2,color='k', ls='--')
# + [markdown] id="jJHUvYuHZZmT"
# #### Descenso del gradiente con varias variables
#
# Dado que el ser humano no puede ver más allá de las tres dimensiones, analizaremos el descenso del gradiente con dos variables independientes. Supongamos la función:
#
# $$x^2 + y^2$$
#
# Esta función es convexa, dado que su matriz hessiana es definida positiva, por tanto el punto extremo que encontraremos será un mínimo.
#
# $$\min_{x,y} x^2 + y^2$$
#
# Encontramos el vector gradiente e igualamos a un vector nulo
#
# $$\nabla f(x,y)=\begin{bmatrix}
# 2x \\
# 2y
# \end{bmatrix}=\textbf{0}$$
#
# Al solucionar el sistema de ecuaciones encontramos que el punto mínimo se da cuando se tiene que
# $$(x,y)=(0,0)$$
#
# Gráficamente
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="X5STAwOnSkLW" outputId="d5196716-18a9-4357-e80d-c00d880465ed"
from mpl_toolkits import mplot3d
# %matplotlib inline
fig = plt.figure()
ax = plt.axes(projection='3d')
x = np.arange(-3,3,0.1)
y = np.arange(-3,3,0.1)
X, Y=np.meshgrid(x, y)
z=X**2+Y**2
ax.plot_surface(X,Y,z)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="FT2OdHe8LzwH" outputId="e4271ecb-0b0a-4f5d-ee0b-29499d132968"
plt.contourf(X,Y,z)
# + colab={"base_uri": "https://localhost:8080/"} id="dFAFhxyJMQGN" outputId="e2e5cbf2-8d50-4ab3-f939-1aae23208e07"
def grad(x,y):
return np.array([2*x,2*y])
p = np.random.randint(-3,3,2)
X_ = []
Y_ = []
lr = 0.8
for _ in range(1000):
X_.append(p[0])
Y_.append(p[1])
p = p - lr * grad(p[0],p[1])
print(p)
# + colab={"base_uri": "https://localhost:8080/", "height": 391} id="kDJGVqNZMsgl" outputId="0fb0f503-4575-4318-b285-c8e9a2d802cb"
plt.figure(figsize=(10,6))
plt.contourf(X,Y,z)
plt.scatter(X_,Y_, color='r')
# + [markdown] id="xZaM4bNhJ87F"
# El descenso del gradiente tiene un problema, es útil cuando la función tiene un solo punto mínimo. Ante varios, puede no converger hacia el global.
# + [markdown] id="bX6K1VvMZgMF"
# # ¡Vamos a crear un objeto!
# + id="AJTChUEWSkDr"
class Regression:
def __init__(self,X,y):
self.X = np.c_[np.ones(len(X)),X]
self.b = None
self.y = y
self.error = []
def fit(self, lr=0.01, iter: int=10000, method: str='gradiente'):
n, k = self.X.shape
if method =='gradiente':
self.b = np.random.randn(k,1)
for _ in range(iter):
Yp = np.dot(self.X,self.b).T
pred = Yp - self.y
self.b = self.b - (1/n)*(lr)*(self.X.T.dot(pred.T))
self.error.append((1/2*n)*np.sum(np.square(pred)))
elif method == 'normales':
self.b = np.linalg.inv((self.X.T @ self.X)) @ (self.X.T @ self.y)
else:
print('Método no válido')
def predict(self, X_test):
X_test = np.c_[np.ones(len(X_test)),X_test]
return np.dot(X_test,self.b)
# + id="pwQ1tgVkf2yZ"
X = np.arange(0,10,0.1)
e = np.random.normal(0,5,100)
y = 35 + 4*X + e
# + id="LDBcrlsMXrUF"
re = Regression(X,y)
# + id="AoALdw3rvqT2"
re.fit()
# + colab={"base_uri": "https://localhost:8080/"} id="-QlZUSIXv0aS" outputId="17404cbc-fd20-46e8-9956-27da9bff92e4"
re.b
# + id="kWW9uyCkv3O_"
re.fit(method='normales')
# + colab={"base_uri": "https://localhost:8080/"} id="XiBmaxdJv6_k" outputId="805f199f-c22c-40b1-900d-a3bda0fab413"
re.b
# + colab={"base_uri": "https://localhost:8080/", "height": 395} id="12gouejhwhT6" outputId="487ef902-9be2-4795-fd95-c57469861283"
plt.figure(figsize=(10,6))
plt.scatter(X,y)
plt.plot(X,re.b[0]+re.b[1]*X, color='r')
plt.scatter(np.array([5.4]), re.predict(np.array([5.4])), color='k')
# + colab={"base_uri": "https://localhost:8080/"} id="J62zQ9Vjw5bv" outputId="66a8206b-eb8b-4c50-e00e-81ec748f00a3"
re.b[0]
# + [markdown] id="YXqxsVU4ajZV"
# # scikit-learn
# + id="XZk75n_PajJI"
from sklearn.linear_model import LinearRegression, Ridge, Lasso
from sklearn.metrics import mean_squared_error, r2_score, mean_absolute_error
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_boston
# + id="inivgZy8Sj3h"
reg = LinearRegression()
# + id="HN1lekNtWI5Y"
boston_data = load_boston()
# + id="F0HlIiAaWIhD"
X_train, X_test, y_train, y_test = train_test_split(boston_data['data'], boston_data['target'], test_size=0.2, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="8AnIimiiWIJR" outputId="9b4ac78a-acc7-4953-f697-3ccd71b83792"
X_train.shape, X_test.shape
# + colab={"base_uri": "https://localhost:8080/"} id="tf7nIpi1Wewa" outputId="6d180171-995d-4478-f877-b82df1128a0c"
reg.fit(X_train, y_train)
# + id="rvCQHLIaWeW7"
y_pred = reg.predict(X_test)
# + [markdown] id="2H2souanaCIZ"
# ### Evaluar el modelo
#
# - Error absoluto medio
#
# $$\text{MAE} = \frac{1}{n}\sum_{i=1}^n |y_i-\hat{y}_i|$$
#
# - Error cuadrático medio
#
# $$\text{MSE} = \frac{1}{n}\sum_{i=1}^n (y_i-\hat{y}_i)^2$$
#
# - Raíz cuadrada del error cuadrático medio
#
# $$\text{RMSE} = \sqrt{\frac{1}{n}\sum_{i=1}^n (y_i-\hat{y}_i)^2}$$
#
# El primero no es diferenciable, por tanto, hace difícil la aplicación del descenso del gradiente. MSE/RMSE penalizan los errores grandes.
#
# - Coeficiente de determinación
#
# $$R^2=1-\frac{\sum_{i=1}^n (y_i-\hat{y}_i)^2}{\sum_{i=1}^n (y_i-\bar{y}_i)^2}$$
#
# Es un número entre 0 y 1 que indica el porcentaje de varianza de las variables de respuesta explicado por el modelo.
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="e4JKXoZgGm1K" outputId="c3b956b4-d9b3-49cc-b6e4-f65a930754cc"
sns.histplot((y_test - y_pred), bins = 40)
# + colab={"base_uri": "https://localhost:8080/"} id="pbxw1_D6avG5" outputId="bac56977-51fa-4b7c-9906-9b52317b9c5f"
mean_squared_error(y_test, y_pred), mean_absolute_error(y_test, y_pred), np.sqrt(mean_squared_error(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="6GF_abZSGJ_V" outputId="5ed1050d-bd09-4761-c383-c6861025b00c"
r2_score(y_pred, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="ACi59jRYGdhl" outputId="7f6339bf-3ea8-4bad-fb40-40bfa0794036"
reg.score(X_test,y_test)
# + [markdown] id="FKWr1ySFT429"
# ### No hay que olvidar lo aprendido
#
# El objetivo del Machine Learning es enseñarle a un computador a comprender los datos para que pueda predecir a partir de la inyección de nueva información. Es decir, la capacidad de predecir es lo importante. Pero después de tres años no se puede olvidar lo aprendido:
#
# - Supuestos de la regresión lineal
#
# - Linealidad de los parámetros
# - Rango completo
# - Exogeneneidad de las variables independientes
# - Homocedasticidad
# - Muestreo aleatorio
# - Distribución normal de los errores
#
# - Causalidad
#
# - Significancia estadística de los parámetros y del modelo
# - Porque unas variables guarden una relación no quiere decir que haya causalidad entre ellas
#
# - Estructuras de datos
#
# - Corte transversal
# - Panel de datos
# - Series de tiempo
# + [markdown] id="Wov-2eqkoByi"
# ## Regresión polinomial
# + id="U8mbtC8roBeQ"
from sklearn.preprocessing import PolynomialFeatures
from sklearn.model_selection import train_test_split
X = np.arange(0,60).reshape(60,1)
e = np.random.normal(0,25,60).reshape(60,1)
y = 35 + 4*X + e
# + id="ZSSBGnMbKib_"
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.2, random_state=42)
# + id="z8KzUQSV7zTv"
poly3 = PolynomialFeatures(3,include_bias=False)
poly10 = PolynomialFeatures(10, include_bias=False)
X_3_train = poly3.fit_transform(X_train)
X_3_test = poly3.fit_transform(X_test)
X_10_train = poly10.fit_transform(X_train)
X_10_test = poly10.fit_transform(X_test)
# + id="9ThWfuNHLXfN" colab={"base_uri": "https://localhost:8080/"} outputId="40c93d88-6fec-44c3-eb41-1a66cba9890d"
X_3_train[:6]
# + id="Z0YW9VWf9SGr"
reg = LinearRegression()
reg.fit(X_train,y_train)
model = reg.predict(np.sort(X_train, axis=0))
# + id="aGOr4Hny9p1h" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="345a35f0-4a63-4556-8849-68b87ec932d5"
plt.scatter(X_train,y_train)
plt.plot(np.sort(X_train, axis=0),model, color='red')
# + id="w5OMkl5pHxS1"
reg3 = LinearRegression()
reg3.fit(X_3_train,y_train)
model3 = reg3.predict(np.sort(X_3_train, axis=0))
# + id="5zTpRcMF_jKZ" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="6ddcf8be-4c0f-438d-b5b1-950839f5e93d"
plt.scatter(X_train,y_train)
plt.plot(np.sort(X_train,axis=0),model3, color='red')
# + id="Cy3jXiFPBTOc"
reg10 = LinearRegression()
reg10.fit(X_10_train,y_train)
model10 = reg10.predict(np.sort(X_10_train, axis=0))
# + id="X9s2AKvgBj1r" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="cde18dca-ab96-4c14-d04f-1ce9b8a42f25"
plt.scatter(X_train,y_train)
plt.plot(np.sort(X_train,axis=0),model10, color='red')
# + id="q3_HH2Lm9IPF" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="59a6035b-af89-472b-8646-4028bc1024f5"
fig, ax = plt.subplots(1,3, sharey=True, figsize=(20,4))
ax[0].plot(np.sort(X_train,axis=0),model, color='red')
ax[0].scatter(X_train,y_train)
ax[0].set_title('Lineal')
ax[1].plot(np.sort(X_train,axis=0),model3, color='red')
ax[1].scatter(X_train,y_train)
ax[1].set_title('Pol3')
ax[2].plot(np.sort(X_train,axis=0),model10, color='red')
ax[2].scatter(X_train,y_train)
ax[2].set_title('Pol10')
# + id="lirHg_DnPM0A"
y_pred = reg.predict(np.sort(X_test, axis=0))
y_pred_3 = reg3.predict(np.sort(X_3_test, axis=0))
y_pred_10 = reg10.predict(np.sort(X_10_test, axis=0))
# + id="X9xgPUI6Pgle" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="6de0f16e-3080-4097-a5b1-a4fe9d07b3b1"
fig, ax = plt.subplots(1,3, sharey=True, figsize=(20,4))
ax[0].plot(np.sort(X_test,axis=0),y_pred, color='red')
ax[0].scatter(X_test,y_test)
ax[0].set_title('Lineal')
ax[1].plot(np.sort(X_test,axis=0),y_pred_3, color='red')
ax[1].scatter(X_test,y_test)
ax[1].set_title('Pol3')
ax[2].plot(np.sort(X_test,axis=0),y_pred_10, color='red')
ax[2].scatter(X_test,y_test)
ax[2].set_title('Pol10')
# + id="mKCnADSiPgZ4" colab={"base_uri": "https://localhost:8080/", "height": 298} outputId="309634e2-b483-4628-d428-fcdbab3e2462"
fig, ax = plt.subplots(2,3, sharex=True, sharey=True, figsize=(20,4))
ax[0, 0].plot(np.sort(X_train,axis=0),model, color='red')
ax[0, 0].scatter(X_train,y_train)
ax[0, 0].set_title('Lineal')
ax[0, 1].plot(np.sort(X_train,axis=0),model3, color='red')
ax[0, 1].scatter(X_train,y_train)
ax[0, 1].set_title('Pol3')
ax[0, 2].plot(np.sort(X_train,axis=0),model10, color='red')
ax[0, 2].scatter(X_train,y_train)
ax[0, 2].set_title('Pol10')
ax[1, 0].plot(np.sort(X_test,axis=0),y_pred, color='red')
ax[1, 0].scatter(X_test,y_test)
ax[1, 1].plot(np.sort(X_test,axis=0),y_pred_3, color='red')
ax[1, 1].scatter(X_test,y_test)
ax[1, 2].plot(np.sort(X_test,axis=0),y_pred_10, color='red')
ax[1, 2].scatter(X_test,y_test)
# + id="RY1zOH69T1n6" colab={"base_uri": "https://localhost:8080/"} outputId="aaca81ae-267f-4d1b-d23c-ae1e71fc0efe"
print('Evaluación en el entrenamiento')
print(f'El error cuadrático medio del primer modelo es: {mean_squared_error(y_train,reg.predict(X_train))}')
print(f'El error cuadrático medio del segundo modelo es: {mean_squared_error(y_train,reg3.predict(X_3_train))}')
print(f'El error cuadrático medio del tercer modelo es: {mean_squared_error(y_train,reg10.predict(X_10_train))}')
# + id="PbN5aJdoVZKV" colab={"base_uri": "https://localhost:8080/"} outputId="adcc7280-180b-48d2-a9f5-00a734e4a4fe"
print('Evaluación en el testeo')
print(f'El error cuadrático medio del primer modelo es: {mean_squared_error(y_test,reg.predict(X_test))}')
print(f'El error cuadrático medio del segundo modelo es: {mean_squared_error(y_test,reg3.predict(X_3_test))}')
print(f'El error cuadrático medio del tercer modelo es: {mean_squared_error(y_test,reg10.predict(X_10_test))}')
# + id="tuolxNxTT4C6" colab={"base_uri": "https://localhost:8080/"} outputId="7806ad51-86ba-4e83-cb0c-f07dd0a8cca9"
print('Evaluación en el entrenamiento')
print(f'El error cuadrático medio del primer modelo es: {mean_absolute_error(y_train,reg.predict(X_train))}')
print(f'El error cuadrático medio del segundo modelo es: {mean_absolute_error(y_train,reg3.predict(X_3_train))}')
print(f'El error cuadrático medio del tercer modelo es: {mean_absolute_error(y_train,reg10.predict(X_10_train))}')
# + id="-VporoXCVvqK" colab={"base_uri": "https://localhost:8080/"} outputId="14754548-b93d-486f-ee00-5fd27b4ec30d"
print('Evaluación en el testeo')
print(f'El error cuadrático medio del primer modelo es: {mean_absolute_error(y_test,reg.predict(X_test))}')
print(f'El error cuadrático medio del segundo modelo es: {mean_absolute_error(y_test,reg3.predict(X_3_test))}')
print(f'El error cuadrático medio del tercer modelo es: {mean_absolute_error(y_test,reg10.predict(X_10_test))}')
# + id="f-VaT2TWVZ2B" colab={"base_uri": "https://localhost:8080/"} outputId="f26f2604-c306-4e0f-81a7-ba977a311f07"
print('Evaluación en el entrenamiento')
print(f'El error cuadrático medio del primer modelo es: {r2_score(y_train,reg.predict(X_train))}')
print(f'El error cuadrático medio del segundo modelo es: {r2_score(y_train,reg3.predict(X_3_train))}')
print(f'El error cuadrático medio del tercer modelo es: {r2_score(y_train,reg10.predict(X_10_train))}')
# + id="hMOo2nc5V8OG" colab={"base_uri": "https://localhost:8080/"} outputId="a84e1e63-bec1-418e-c529-9d279b21b9a0"
print('Evaluacion en el testeo')
print(f'El error cuadrático medio del primer modelo es: {r2_score(y_test,reg.predict(X_test))}')
print(f'El error cuadrático medio del segundo modelo es: {r2_score(y_test,reg3.predict(X_3_test))}')
print(f'El error cuadrático medio del tercer modelo es: {r2_score(y_test,reg10.predict(X_10_test))}')
# + [markdown] id="Eze6hJ8N0XVb"
# ## Regresión Ridge (L2)
#
# Ante la presencia de overfitting en una regresión lineal es posible emplear distintos métodos para penalizar los coeficientes de regresión, entre ellos, la regresión Ridge. El proceso de optimización que se realiza es el siguiente
#
# $$\min_{\textbf{b}}||\textbf{X}\textbf{b}-\textbf{y}||^2_2+\lambda||\textbf{b}||^2_2$$
#
# O escrito en forma matricial
#
# $$\min_{\textbf{b}}(\textbf{y}-\textbf{X}\textbf{b})'(\textbf{y}-\textbf{X}\textbf{b})+\lambda\textbf{b}'\textbf{b}$$
#
# $$\textbf{b}_{ridge}=(\textbf{X}'\textbf{X}-\lambda\textbf{I})^{-1}\textbf{X}'\textbf{y}$$
#
# Con variables correlacionadas en un modelo, los estimadores pueden ser pobremente estimados. Al establecer una restricción de tamaño en los coeficientes se soluciona el problema. Usualmente se normalizan los inputs antes de resolver el problema de optimización.
#
# En la regresión de Ridge, el valor de $\lambda$ es crítico. El valor óptimo puede encontrarse a través de validación cruzada, grid search, entre otros métodos.
#
# NOTA: el intercepto no es regularizado
# + id="WZylSg2fvavn"
from sklearn.linear_model import Ridge
from sklearn.model_selection import train_test_split
# + id="4id-wLpEJ4lD" colab={"base_uri": "https://localhost:8080/", "height": 653} outputId="7e657b4b-0e5c-43f0-b711-d511b0823e64"
plt.figure(figsize=(12,8))
for i in np.arange(0,100000,10000):
ridge = Ridge(alpha=i)
ridge.fit(X_train,y_train)
model = ridge.predict(X_train)
print(ridge.coef_, ridge.intercept_)
plt.plot(X_train, model, label=i)
plt.scatter(X_train, y_train)
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="FZ7sDI_vwoBc" outputId="8ab50de5-cdc6-43d6-c06f-b8841333b2da"
ridge10 = Ridge(alpha=10000000)
ridge10.fit(X_10_train,y_train)
model_r10 = ridge10.predict(X_10_train)
yridge_10 = ridge10.predict(X_10_test)
# + colab={"base_uri": "https://localhost:8080/"} id="p0Gwcq_vw9WG" outputId="2dd3514f-0e39-4f3f-c990-9ea85318a739"
mean_squared_error(y_train,model_r10), mean_squared_error(y_test,yridge_10)
# + [markdown] id="SL1HyzeW0ara"
# ## Regresión Lasso (L1)
#
# Sucede que con la regresión Ridge, el tamaño de los coeficientes es penalizado a través del $\lambda$. No obstante, aunque estos coeficientes pueden asintóticamente tender a 0, nunca van a llegar a serlo. Este problema es solucionado por Lasso, donde el problema de overfitting presente en la regresión lineal es solucionado eliminando aquellas variables que no son relevantes.
#
# $$ \min_{\textbf{b}}||\textbf{X}\textbf{b}-\textbf{y}||^2_2+\lambda||\textbf{b}||_1$$
#
#
# Al igual que en la regresión Ridge, el valor de $\lambda$ es crítico.
#
# + id="y1cRHIWin_y5"
from sklearn.linear_model import Lasso
# + id="smoGrx9LbEk5" colab={"base_uri": "https://localhost:8080/", "height": 775} outputId="caedcf41-d955-4c2b-87ad-79ea4b90c5ad"
plt.figure(figsize=(12,8))
for i in np.arange(0,10000,1000):
lasso = Lasso(alpha=i)
lasso.fit(X_train,y_train)
model = lasso.predict(X_train)
print(lasso.coef_, lasso.intercept_)
plt.plot(X_train, model, label=i)
plt.scatter(X_train, y_train)
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="J0ggWBJZ9hDj" outputId="a735a4a6-de3d-41cb-9056-b2cc444f7d1d"
lasso10 = Lasso(alpha=5)
lasso10.fit(X_10_train,y_train)
model_l10 = lasso10.predict(X_10_train)
ylasso_10 = lasso10.predict(X_10_test)
# + colab={"base_uri": "https://localhost:8080/"} id="x9UolX7U9htw" outputId="7b642deb-e466-4bb0-f4b6-848f38fe86a7"
mean_squared_error(y_train,model_l10), mean_squared_error(y_test,ylasso_10)
# + colab={"base_uri": "https://localhost:8080/"} id="AlvKyi8T9-Bt" outputId="a8e77db7-bbe4-4c9d-91fe-859625675940"
lasso10.coef_
# + [markdown] id="CYLZrAm50cnB"
# ## Elastic Net (L1 y L2)
#
# Es un modelo de regresión lineal que combina la regularización de los coeficientes de Lasso y de Ridge. Es útil cuando se tienen varias variables que están correlacionadas entre sí. Lasso elimina una de estas variables de manera aleatoria mienras que Elastic-Net las elimina todas.
#
# La función objetivo a minimizar es
#
# $$\min_{\textbf{b}}\frac{1}{2n_{muestras}}||\textbf{X}\textbf{b}-\textbf{y}||^2_2+\lambda\rho||\textbf{b}||_1+\frac{\lambda(1-\rho)}{2}||\textbf{b}||^2_2$$
#
# A través de validación cruzada se pueden encontrar los valores óptimos para $\lambda$ y $\rho$. Este último se emplea para controlar la combinación convexa de las dos formas de regularización.
# + id="lvalvCpcoAVz"
from sklearn.linear_model import ElasticNet
# + id="_xWILyGcdDkd" colab={"base_uri": "https://localhost:8080/", "height": 775} outputId="035f4603-10ba-46e2-913f-9e6ee4358e62"
plt.figure(figsize=(12,8))
for i in np.arange(0,1000,100):
en = ElasticNet(alpha=i,l1_ratio=0.2)
en.fit(X_train,y_train)
model = en.predict(X_train)
print(en.coef_, en.intercept_)
plt.plot(X_train, model, label=i)
plt.scatter(X_train, y_train)
plt.legend()
plt.show()
# + [markdown] id="28HostUJ1RxN"
# ## Un ejercicio
# + id="9Z5bAgxCdYKh"
from sklearn.datasets import load_boston
# + id="JhoVspWt1ROS"
X = load_boston()['data']
y = load_boston()['target']
# + id="s9CP3eVB1NgC"
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.25, random_state=42)
# + id="qPPXN4Hv0Wvs" colab={"base_uri": "https://localhost:8080/"} outputId="84d49934-d86b-408f-f0a3-25dababe393a"
load_boston()['feature_names']
# + [markdown] id="KplXXUQrfT7S"
# ### Regresión lineal
# + id="hkUTKl1BeBXE" colab={"base_uri": "https://localhost:8080/"} outputId="fa123a0b-3d40-44e5-de47-4a893305c6c9"
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler().fit(X_train)
X_scaled = scaler.transform(X_train)
reg = LinearRegression()
reg.fit(X_scaled, y_train)
y_pred_d = reg.predict(X_scaled)
y_pred = reg.predict(scaler.transform(X_test))
print(f'Evaluación en entrenamiento: {mean_squared_error(y_pred_d, y_train)}')
print(f'Evaluación en testeo: {mean_squared_error(y_pred, y_test)}')
# + [markdown] id="CV0mqG4dfWIA"
# ### Ridge
# + id="H5E3cSm619rb" colab={"base_uri": "https://localhost:8080/"} outputId="512f06c2-a706-48cd-8a79-ec69c5458672"
ridge = Ridge()
ridge.fit(X_scaled, y_train)
y_pred = ridge.predict(scaler.transform(X_test))
print(f'Evaluación en entrenamiento: {ridge.score(X_scaled, y_train)}')
print(f'Evaluación en testeo: {ridge.score(scaler.transform(X_test), y_test)}')
# + id="JRMyCOIC2X9t"
X_test.shape
# + id="2dmuI3sGfgrB" colab={"base_uri": "https://localhost:8080/"} outputId="470ac0f7-a7bc-4c93-e9e6-27bf3b9f8ab8"
ridge_coef = []
for i in np.arange(0,100,10):
ridge = Ridge(alpha=i)
ridge.fit(X_scaled, y_train)
y_pred = ridge.predict(scaler.transform(X_test))
ridge_coef.append(ridge.coef_)
print(f'Evaluación en entrenamiento: {ridge.score(X_scaled, y_train)}')
print(f'Evaluación en testeo: {ridge.score(scaler.transform(X_test), y_test)}')
# + id="nGDOPP58hMZy" colab={"base_uri": "https://localhost:8080/", "height": 483} outputId="c99cc0b8-f1b3-4f67-950b-388aca96d7bd"
from matplotlib.lines import Line2D as ld
mark = [i for i in ld.markers.keys()]
plt.figure(figsize=(12,8))
plt.plot(reg.coef_, 'ro', markersize=10)
plt.axhline(0, alpha=0.4, color='k', ls='--')
for i, j in zip(ridge_coef, mark[:10]):
plt.plot(i, j, markersize=10)
# + [markdown] id="ZaqI0OjWfXph"
# ### Lasso
# + id="BRPikm-re61h" colab={"base_uri": "https://localhost:8080/"} outputId="6348b369-3455-4b3b-a6ff-0edf10dfc745"
lasso = Lasso()
lasso.fit(X_scaled, y_train)
y_pred = lasso.predict(scaler.transform(X_test))
print(f'Evaluación en entrenamiento: {lasso.score(X_scaled, y_train)}')
print(f'Evaluación en testeo: {lasso.score(scaler.transform(X_test), y_test)}')
# + id="xuQHcyRHf7DO" colab={"base_uri": "https://localhost:8080/"} outputId="01d550ea-2f9b-4b51-ac7f-6b85b48f23ad"
lasso_coef = []
for i in np.arange(0,1,0.1):
lasso = Lasso(alpha=i)
lasso.fit(X_scaled, y_train)
y_pred = lasso.predict(scaler.transform(X_test))
lasso_coef.append(lasso.coef_)
# + id="Z6xHYw3SgQ3S" colab={"base_uri": "https://localhost:8080/", "height": 483} outputId="34ea6b91-b9fe-439a-ce0b-a863df908782"
plt.figure(figsize=(12,8))
plt.plot(reg.coef_, 'ro', markersize=10)
for i, j in zip(lasso_coef, mark[:10]):
plt.plot(i, j, markersize=10)
# + [markdown] id="-bquX67NfZ3_"
# ### ElasticNet
# + id="pxvI8xOqfDX-"
en = ElasticNet()
en.fit(X_train, y_train)
y_pred = en.predict(X_test)
print(f'Evaluación en entrenamiento: {en.score(X_train, y_train)}')
print(f'Evaluación en testeo: {en.score(X_test, y_test)}')
|
clase_10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TensorFlow 튜토리얼 #17
# # Estimator API
#
# 원저자 [<NAME>](http://www.hvass-labs.org/)
# / [GitHub](https://github.com/Hvass-Labs/TensorFlow-Tutorials) / [Videos on YouTube](https://www.youtube.com/playlist?list=PL9Hr9sNUjfsmEu1ZniY0XpHSzl5uihcXZ) / 번역 곽병권
# ## 개요
#
# 높은 수준의 API는 매우 복잡한 작업을 수행하는 데 필요한 추상화를 제공하기 때문에 모든 소프트웨어 개발에서 매우 중요합니다. 이렇게하면 소스 코드를 쉽게 작성하고 이해할 수 있으며 오류의 위험을 줄일 수 있습니다.
#
# 튜토리얼 #03에서 우리는 TensorFlow에서 신경망을 생성하기 위해 다양한 빌더 API를 사용하는 방법을 보았습니다. 그러나 모델을 교육하고 새로운 데이터에 사용하는 데 필요한 추가 코드가 많이 있었습니다. Estimator는 이것이 실제로 얼마나 간단한 지에 대해 논쟁이 가능할지라도 대부분의 것을 구현하는 또 다른 고수준 API입니다.
#
# Estimator API 사용은 여러 단계로 구성됩니다:
#
# 1. Estimator에 데이터를 입력하는 기능을 정의합니다.
# 2. 미리 만들어져 있거나 깡통에 든(Canned Estimator)라고도하는 기존의 Estimator(예: Deep Neural Network)를 사용합니다. 또는 자신의 Estimator를 작성합니다.이 경우 옵티마이저, 성능 메트릭 등을 정의해야합니다.
# 3. 1 단계에서 정의한 학습 세트를 사용하여 Estimator를 학습합니다.
# 4. 1 단계에서 정의한 테스트 세트에서 Estimator의 성능을 평가합니다.
# 5. 학습 된 Estimator를 사용하여 다른 데이터에 대한 예측을 합니다.
# ## Imports
# %matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
# 이 문서는 Python 3.6.1 (Anaconda) 및 아래의 TensorFlow 버전을 사용하여 개발되었습니다.
tf.__version__
# ## Load Data
# MNIST 데이터 세트는 약 12MB이며 주어진 경로에 위치하지 않으면 자동으로 다운로드 됩니다.
from tensorflow.examples.tutorials.mnist import input_data
data = input_data.read_data_sets('data/MNIST/', one_hot=True)
# MNIST 데이터 세트는 현재 로드 되었으며 70.000개의 이미지 및 관련 라벨 (즉, 이미지의 분류)로 구성됩니다. 데이터 집합은 3개의 상호 배타적인 하위 집합으로 나뉩니다. 이 튜토리얼에서는 훈련 및 테스트 세트 만 사용합니다.
print("Size of:")
print("- Training-set:\t\t{}".format(len(data.train.labels)))
print("- Test-set:\t\t{}".format(len(data.test.labels)))
print("- Validation-set:\t{}".format(len(data.validation.labels)))
# 클래스 레이블은 One-Hot로 인코딩 됩니다. 즉, 각 레이블은 하나의 요소를 제외하고 모두 0인 요소가 포함 된 10개의 벡터입니다. 이 요소의 색인은 클래스 번호, 즉 연관된 이미지에 표시된 숫자입니다. 테스트 집합에 대한 클래스 수를 정수로 필요로 하므로 지금 계산합니다.
data.train.cls = np.argmax(data.train.labels, axis=1)
data.test.cls = np.argmax(data.test.labels, axis=1)
# 이것은 원 핫 인코드 된 레이블의 예입니다.
data.train.labels[0:10]
# 이것들은 대응하는 클래스 번호들입니다.
data.train.cls[0:10]
# ## Data Dimensions
# 데이터 차원은 아래 소스 코드의 여러 위치에서 사용됩니다. 그것들은 한 번 정의되어 있으므로 아래의 소스 코드에서 숫자 대신 이러한 변수를 사용할 수 있습니다.
# +
# We know that MNIST images are 28 pixels in each dimension.
img_size = 28
# Images are stored in one-dimensional arrays of this length.
img_size_flat = img_size * img_size
# Tuple with height and width of images used to reshape arrays.
img_shape = (img_size, img_size)
# Number of colour channels for the images: 1 channel for gray-scale.
num_channels = 1
# Number of classes, one class for each of 10 digits.
num_classes = 10
# -
# ### 이미지를 그리는 도움 함수
# 3x3그리드에 9개의 이미지를 플롯하고 각 이미지 아래에 참 및 예측 클래스를 쓰는 데 사용되는 함수입니다.
def plot_images(images, cls_true, cls_pred=None):
assert len(images) == len(cls_true) == 9
# Create figure with 3x3 sub-plots.
fig, axes = plt.subplots(3, 3)
fig.subplots_adjust(hspace=0.3, wspace=0.3)
for i, ax in enumerate(axes.flat):
# Plot image.
ax.imshow(images[i].reshape(img_shape), cmap='binary')
# Show true and predicted classes.
if cls_pred is None:
xlabel = "True: {0}".format(cls_true[i])
else:
xlabel = "True: {0}, Pred: {1}".format(cls_true[i], cls_pred[i])
# Show the classes as the label on the x-axis.
ax.set_xlabel(xlabel)
# Remove ticks from the plot.
ax.set_xticks([])
ax.set_yticks([])
# Ensure the plot is shown correctly with multiple plots
# in a single Notebook cell.
plt.show()
# ### 일부의 이미지를 그려서 데이터가 정확한지 확인해 봅니다.
# +
# Get the first images from the test-set.
images = data.test.images[0:9]
# Get the true classes for those images.
cls_true = data.test.cls[0:9]
# Plot the images and labels using our helper-function above.
plot_images(images=images, cls_true=cls_true)
# -
# ## Estimator를 위한 입력 함수
# Estimator에 원시 데이터를 직접 제공하는 대신 데이터를 반환하는 함수를 제공해야 합니다. 이를 통해 데이터 소스에 더 많은 유연성과 데이터가 임의로 셔플 링되고 반복되는 방식을 제공할 수 있습니다.
#
# class-numbers가 정수라고 가정하는 `DNNClassifier`를 사용하여 Estimator를 생성 할 것이므로 one-hot으로 인코딩 된 배열 인 `data.train.labels` 대신 `data.train.cls`를 사용합니다.
#
# 이 함수는 또한 데이터 읽기를 더 세밀하게 제어하기 위해 `batch_size`, `queue_capacity` 및 `num_threads`를 위한 매개 변수를 가지고 있습니다. 여기서는 메모리의 numpy 배열로부터 직접 데이터를 전달하므로 필요하지 않습니다.
train_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": np.array(data.train.images)},
y=np.array(data.train.cls),
num_epochs=None,
shuffle=True)
# 이것은 실제로는 함수를 반환합니다.
train_input_fn
# 이 함수를 호출하면 입력 및 출력 데이터를 반환하기 위한 TensorFlow 연산이 포함 된 튜플이 반환됩니다.
train_input_fn()
# 마찬가지로 우리는 테스트 세트에 대한 데이터를 읽는 함수를 생성해야 합니다. 이 이미지를 한 번만 처리할 것 이므로 `num_epochs=1`이고 `shuffle=False`로 합니다.
test_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": np.array(data.test.images)},
y=np.array(data.test.cls),
num_epochs=1,
shuffle=False)
# 새로운 데이터의 클래스를 예측하기 위해 입력 함수가 필요합니다. 예를 들어, 테스트 세트의 이미지 몇 개를 사용합니다.
some_images = data.test.images[0:9]
predict_input_fn = tf.estimator.inputs.numpy_input_fn(
x={"x": some_images},
num_epochs=1,
shuffle=False)
# 클래스 번호는 실제로 예측에 필요하지 않으므로 실제로 입력 함수에서 사용되지 않습니다. 그러나 이미지를 아래에 더 자세히 표시하려면 실제 클래스 번호가 필요합니다.
some_images_cls = data.test.cls[0:9]
# ## 내장 Estimator (Pre-Made / Canned Estimator)
#
# 사전 제작 된 Estimator를 사용할 때 데이터의 입력 기능을 지정해야합니다. 여기서는 주어진 형태의 숫자 배열 인 데이터 세트의 이미지를 입력하려고 합니다.
feature_x = tf.feature_column.numeric_column("x", shape=img_shape)
# 목록에 여러 입력 feature들을 조합하여 사용할 수 있습니다.
feature_columns = [feature_x]
# 이 예에서는 각각 512, 256 및 128 단위의 3층 DNN을 사용하려고합니다.
num_hidden_units = [512, 256, 128]
# `DNNClassifier`는 신경망을 만들어 줍니다. 활성화 함수 및 다양한 다른 매개 변수도 지정할 수 있습니다 (문서 참조). 여기에서는 클래스의 수와 체크포인트가 저장 될 디렉토리를 지정합니다.
model = tf.estimator.DNNClassifier(feature_columns=feature_columns,
hidden_units=num_hidden_units,
activation_fn=tf.nn.relu,
n_classes=num_classes,
model_dir="./checkpoints_tutorial17-1/")
# ### 학습
#
# 이제 주어진 반복 횟수만큼 모델을 트레이닝 할 수 있습니다. 그러면 나중에 학습을 계속할 수 있도록 체크포인트를 자동으로 로드하고 저장합니다.
#
# `INFO:tensorflow:` 텍스트가 모든 행에 인쇄되어 실제 진행 상황을 빨리 읽는 것을 어렵게 만들기 때문에 한 줄에 인쇄해야합니다.
model.train(input_fn=train_input_fn, steps=2000)
# ### 평가
#
# 모델이 훈련되면 테스트 세트에서 성능을 평가할 수 있습니다.
result = model.evaluate(input_fn=test_input_fn)
result
print("Classification accuracy: {0:.2%}".format(result["accuracy"]))
# ### 예측
#
# 학습 된 모델은 또한 새로운 데이터에 대한 예측을 하는데 사용될 수 있습니다.
#
# TensorFlow 그래프가 다시 생성되고 새 데이터에 대한 예측을 할 때마다 체크 포인트가 다시 로드됩니다. 모델이 매우 큰 경우에는 상당한 오버 헤드가 추가 될 수 있습니다.
#
# Estimator는 항상 최신 체크포인트를 사용하는데, 이렇게 설계된 이유는 아마도 여러 컴퓨터에서 쉽게 사용할 수 있도록 하려는 것으로 보입니다.
predictions = model.predict(input_fn=predict_input_fn)
cls = [p['classes'] for p in predictions]
cls_pred = np.array(cls, dtype='int').squeeze()
cls_pred
plot_images(images=some_images,
cls_true=some_images_cls,
cls_pred=cls_pred)
# # Estimator 만들기
# 내장 Estimator 중 하나를 사용할 수 없으면 임의의 TensorFlow 모델을 직접 만들 수 있습니다. 이렇게 하려면 먼저 다음을 정의하는 함수를 만들어야합니다.
#
# 1. TensorFlow 모델 예) 컨볼루션 신경망
# 2. 모델의 출력
# 3. 최적화 과정에서 모델을 향상시키는 데 사용 된 손실 함수
# 4. 최적화 방법
# 5. 성능 메트릭
#
# Estimator 학습, 평가 또는 예측의 세 가지 모드로 실행할 수 있습니다. 코드는 대부분 같지만 예측 모드에서는 손실 함수 및 최적화 프로그램을 설정할 필요가 없습니다.
#
# 이는 Estimator API 설계의 또 다른 측면으로 예전에 구조체를 사용하던 ANSI C 프로그래밍을 수행 한 방식과 유사합니다. 아마도 이것을 여러 함수로 나누고 Estimator 클래스를 하위 클래스로 만들면 더 우아했을 것입니다.
def model_fn(features, labels, mode, params):
# Args:
#
# features: This is the x-arg from the input_fn.
# labels: This is the y-arg from the input_fn,
# see e.g. train_input_fn for these two.
# mode: Either TRAIN, EVAL, or PREDICT
# params: User-defined hyper-parameters, e.g. learning-rate.
# Reference to the tensor named "x" in the input-function.
x = features["x"]
# The convolutional layers expect 4-rank tensors
# but x is a 2-rank tensor, so reshape it.
net = tf.reshape(x, [-1, img_size, img_size, num_channels])
# First convolutional layer.
net = tf.layers.conv2d(inputs=net, name='layer_conv1',
filters=16, kernel_size=5,
padding='same', activation=tf.nn.relu)
net = tf.layers.max_pooling2d(inputs=net, pool_size=2, strides=2)
# Second convolutional layer.
net = tf.layers.conv2d(inputs=net, name='layer_conv2',
filters=36, kernel_size=5,
padding='same', activation=tf.nn.relu)
net = tf.layers.max_pooling2d(inputs=net, pool_size=2, strides=2)
# Flatten to a 2-rank tensor.
net = tf.contrib.layers.flatten(net)
# Eventually this should be replaced with:
# net = tf.layers.flatten(net)
# First fully-connected / dense layer.
# This uses the ReLU activation function.
net = tf.layers.dense(inputs=net, name='layer_fc1',
units=128, activation=tf.nn.relu)
# Second fully-connected / dense layer.
# This is the last layer so it does not use an activation function.
net = tf.layers.dense(inputs=net, name='layer_fc2',
units=10)
# Logits output of the neural network.
logits = net
# Softmax output of the neural network.
y_pred = tf.nn.softmax(logits=logits)
# Classification output of the neural network.
y_pred_cls = tf.argmax(y_pred, axis=1)
if mode == tf.estimator.ModeKeys.PREDICT:
# If the estimator is supposed to be in prediction-mode
# then use the predicted class-number that is output by
# the neural network. Optimization etc. is not needed.
spec = tf.estimator.EstimatorSpec(mode=mode,
predictions=y_pred_cls)
else:
# Otherwise the estimator is supposed to be in either
# training or evaluation-mode. Note that the loss-function
# is also required in Evaluation mode.
# Define the loss-function to be optimized, by first
# calculating the cross-entropy between the output of
# the neural network and the true labels for the input data.
# This gives the cross-entropy for each image in the batch.
cross_entropy = tf.nn.sparse_softmax_cross_entropy_with_logits(labels=labels,
logits=logits)
# Reduce the cross-entropy batch-tensor to a single number
# which can be used in optimization of the neural network.
loss = tf.reduce_mean(cross_entropy)
# Define the optimizer for improving the neural network.
optimizer = tf.train.AdamOptimizer(learning_rate=params["learning_rate"])
# Get the TensorFlow op for doing a single optimization step.
train_op = optimizer.minimize(
loss=loss, global_step=tf.train.get_global_step())
# Define the evaluation metrics,
# in this case the classification accuracy.
metrics = \
{
"accuracy": tf.metrics.accuracy(labels, y_pred_cls)
}
# Wrap all of this in an EstimatorSpec.
spec = tf.estimator.EstimatorSpec(
mode=mode,
loss=loss,
train_op=train_op,
eval_metric_ops=metrics)
return spec
# ### Estimator 인스턴스 만들기
#
# 예를 들어 하이퍼 매개 변수를 지정할 수 있습니다. 예를 들면, 최적화 알고리즘의 학습 속도와 같이 말입니다.
params = {"learning_rate": 1e-4}
# 그런 다음 새 Estimator 인스턴스를 만들 수 있습니다.
#
# `model_fn()`이 호출 될 때 데이터 함수로부터 자동으로 유추되기 때문에 우리는 여기에 feature-columns를 제공하지 않습니다.
#
# 위의 예제에서 `DNNClassifier`를 사용할 때 TensorFlow 문서에서 왜 feature-columns을 지정해야 하는지 명확하지 않습니다. 여기서는 필요하지 않습니다.
model = tf.estimator.Estimator(model_fn=model_fn,
params=params,
model_dir="./checkpoints_tutorial17-2/")
# ### 학습
#
# 새로운 Estimator가 만들어졌으므로, 우리는 그것을 학습시킬 수 있습니다.
model.train(input_fn=train_input_fn, steps=2000)
# ### 평가
#
# 모델이 훈련되면 테스트 세트에서 성능을 평가할 수 있습니다.
result = model.evaluate(input_fn=test_input_fn)
result
print("Classification accuracy: {0:.2%}".format(result["accuracy"]))
# ### 예측
#
# 모델을 사용하여 새 데이터에 대한 예측을 수행 할 수도 있습니다.
predictions = model.predict(input_fn=predict_input_fn)
cls_pred = np.array(list(predictions))
cls_pred
plot_images(images=some_images,
cls_true=some_images_cls,
cls_pred=cls_pred)
# ## 결론
#
# 이 튜토리얼에서는 TensorFlow에서 Estimator API를 사용하는 방법을 보여줍니다. 그것은 모델을 훈련하고 사용하는 것을 더 쉽게 만들어야 한다고 생각되지만 몇 가지 디자인 문제가 있어 보입니다.
#
# * Estimator API는 복잡하고 일관성이 없으며 혼란 스럽습니다.
# * 오류 - 메시지는 매우 길며 이해하기가 쉽지 않습니다.
# * TensorFlow 그래프가 다시 생성되고 새 데이터에 대한 예측을 수행하기 위해 학습 된 모델을 사용할 때마다 체크 포인트가 다시 로드됩니다. 일부 모델은 매우 커서 오버 헤드가 매우 커질 수 있습니다. 체크 포인트가 디스크에서 변경된 경우에만 모델을 다시 로드하는 것이 더 좋은 방법 일 수 있습니다.
# * 학습 된 모델에 액세스하는 방법이 불분명합니다 (예: 신경망의 가중치를 그리고 싶을때와 같이.)
#
# Estimator API는 훨씬 간단하고 사용하기 쉬울 수 있었습니다. 소규모 프로젝트의 경우에는 너무 복잡하고 혼란 스럽기 때문에 그만한 가치를 느끼기 어렵습니다. 그러나 매우 큰 데이터 집합이 있고 많은 컴퓨터에서 학습하는 경우 Estimator API가 유용 할 수 있습니다.
# ## 연습 문제
#
# TensorFlow 활용 기술을 향상 시키는데 도움이 될 수있는 몇 가지 제안 사항입니다. TensorFlow를 제대로 사용하는 방법을 배우려면 실습 경험을 쌓는 것이 중요합니다.
#
# 변경하기 전에 이 노트북을 백업 해 두세요.
#
# * 각 모델에 대해 10000회의 학습 반복을 실행합니다.
# * 최적화 전, 1000, 2000 및 10000 반복 후 테스트 세트의 분류 정확도를 인쇄합니다.
# * Estimator 내부의 신경망 구조를 변경하십시오. 체크포인트 파일을 삭제해야합니까? 왜 그럴까요?
# * 입력 함수의 batch 크기를 변경하십시오.
# * 많은 이전 튜토리얼에서 우리는 잘못 분류 된 이미지의 예를 도식화 했습니다. 여기에서도 똑같이 해보세요.
# * Estimator 정수 클래스 번호 대신 One-Hot 인코드된 레이블을 사용하도록 변경하십시오.
# * numpy 배열을 사용하는 대신 이미지 파일을 로드하도록 입력 함수를 변경하십시오.
# * 신경망의 가중치와 개별 레이어의 출력을 플로팅하는 방법을 찾을 수 있습니까?
# * Estimator API에 대해 좋아하는 것과 좋아하지 않는 5 가지를 나열하십시오. 개선을위한 제안 사항이 있습니까? 어쩌면 개발자에게 제안해야할까요?
# * 프로그램이 어떻게 작동하는지 동료에게 설명하십시오.
# ## 라이센스 (MIT)
#
# 저작권 (c) 2016-2017 [<NAME>] (http://www.hvass-labs.org/)
#
# 사용, 복사, 수정, 병합 할 수있는 권한을 포함하되 이에 국한되지 않고 소프트웨어를 취급하기 위해 이 소프트웨어 및 관련 문서 파일 (이하 "소프트웨어")의 사본을 얻는 모든 사람에게 사용 권한이 무료로 부여됩니다 다음 조건에 따라 소프트웨어의 사본을 게시, 배포, 재 라이센스 및 또는 판매 할 수 있고 소프트웨어를 제공받는 사람에게 허용 할 수 있습니다.
#
# 위의 저작권 고지 및 이 허가 고지는 소프트웨어의 모든 사본 또는 상당 부분에 포함되어야합니다.
#
# 소프트웨어는 상품성, 특정 목적에의 적합성 및 비 침해에 대한 보증을 포함하여 (단, 이에 한하지 않음) 명시적이거나 묵시적인 어떤 종류의 보증도 없이 "있는 그대로" 제공됩니다. 제작자 또는 저작권 보유자는 소프트웨어 또는 사용과 관련하여 발생했거나 발생했거나 발생했거나 또는 기타 행위로 인해 발생한 청구, 기타 책임에 대해 어떠한 경우에도 책임을 지지 않습니다.
|
17_Estimator_API.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # EEP/IAS 118 - Introductory Applied Econometrics
# ## Problem Set 1, Spring 2021, Villas-Boas
# #### <span style="text-decoration: underline">Due 9:30am on February 4, 2021</span>
#
#
# Submit materials (all handwritten/typed answers, Excel workbooks, and R reports) as one combined pdf on [Gradescope](https://www.gradescope.com/courses/226571). All students currently on the EEP118 bCourses have been added using the bCourses email. If you do not have access to the Gradescope course, please reach out to the GSI's.
#
# For help combining pdf's, see the recent announcement on bCourses.
# ## Exercise 1 (Excel)
# **Relationship between Gasoline Consumption Data and Price of Gasoline in 18 OECD countries.**
#
# We will use data from Baltagi (2005) on gasoline consumption for 18 OECD countries. The original data span 19 years. In this first problem set we will only use the year 1960.
#
# This exercise is to be completed using Excel. Looking at the first graph in the paper, there appears to be an association between Gasoline consumption per car and the price of gasoline. We will establish a simple linear relationship on a subset of 9 countries at a time.
#
# *Note: in economics, log always refers to the natural log, ln().*
#
# <center><b> Table 1: Log of Gasoline Consumption Per Car and Log of Price of Gasoline, Sample 1</b></center>
#
# |CountryName | log of Gasoline Consumption per Car | log of Gasoline Price |
# |-----------|---------------|---------------|
# |sample 1 | log of Y | log of X |
# |AUSTRIA| 4.173244195|-0.334547613|
# |BELGIUM|4.16401597|-0.165709611|
# |CANADA|4.855238441|-0.972106499|
# |DENMARK|4.50198595|-0.195702601|
# |FRANCE|3.907704233|-0.019598332|
# |GERMANY|3.916953172|-0.185910784|
# |GREECE|5.037405535|-0.083547398|
# |IRELAND|4.270420603|-0.076481181|
# |ITALY|4.050728238|0.165077076
# (a) Use Excel to create a scatter plot of these observations. Don't forget to (1) label the axes and their units, and (2) title your graph. **You should use the tables provided here for these calculations, not the actual observations from the .csv data file.**
# (b) This question has **two parts**.
#
# First: Estimate the linear relationship between the log of Gasoline
# consumption per car (log(Y)) and the log of gasoline prices (log(X)) by OLS, showing all intermediate
# calculations as we saw in the lecture 3 slides (use Excel to create the table and show all the steps).
#
# Second: interpret the value of the estimated parameters $\beta_0$ and $\beta_1$.
#
# $$ \widehat{log (Y_i)} = \hat{\beta_0} + \hat{\beta_1} log(X_i) \ \ \ \ \ \ \text{i = \{first 9 countries\}}$$
# ➡️ Type your answer to _Exercise 1 (b) Second Part_ here (replacing this text)
# (c) In your table, compute the fitted value and the residual for each observation, and verify that the residuals (approximately) sum to 0.
# (d) According to the estimated relation, what is the predicted $\hat{Y}$ (**level**, not log) for a country with a log price of -2? (Pay attention to units)
# ➡️ Type your answer to _Exercise 1 (d)_ here (replacing this text)
# (e) How much of the variation in per capita log gasoline consumption in these 9 countries is explained by the log of price of gasoline in the countries?
# ➡️ Type your answer to _Exercise 1 (e)_ here (replacing this text)
# <center><b> Table 2: Log of Gasoline Consumption Per Car and Log of Price of Gasoline, Sample 2</b></center>
#
# |CountryName | log of Gasoline Consumption per Car | log of Gasoline Price |
# |-----------|---------------|---------------|
# |sample 2 | log of Y | log of X |
# |JAPAN|5.995286556|-0.14532271|
# |NETHERLANDS|4.646268005|-0.201484804|
# |NORWAY|4.43504067|-0.139689574|
# |SPAIN|4.749409172|1.125310702|
# |SWEDEN|4.063010036|-2.52041588|
# |SWITZERLAND|4.397621493|-0.82321833|
# |TURKEY|6.129552849|-0.253408214|
# |U.K.|4.100244284|-0.391085814|
# |U.S.A.|4.823964512|-1.121114893|
#
#
# (f) Repeat exercise (b) for one additional set of 9 countries below. **You should use Table 2 provided
# above for these calculations, not the actual observations from the .csv data file.**
# ➡️ Type your answer to _Exercise 1 (e) Second Part_ here (replacing this text)
# (g) Do your estimates of $\hat{\beta_0}$ and $\hat{\beta_1}$ change between Tables 1, and 2? Why?
#
# ➡️ Type your answer to _Exercise 1 (g)_ here (replacing this text)
# (h) Save a copy of your Excel workbook as a pdf (OLS tables and scatter plot) to combine with the later work.
# ## Exercise 2 (Functional Forms)
# (a) Suppose you estimate alternative specifications as given below for the year of 1972 using all countries:
#
# $$ \text{A linear relationship:} ~~~~\hat{Y_i} = 121 + 2.23 X_i$$
# $$ \text{A linear-log relationship:} ~~~~\hat{Y_i} = 4.5 + 0.06 \log(X)_i$$
# $$ \text{A log-log relationship:} ~~~~\widehat{\log(Y)}_i = 4 + 0.09 \log(X)_i$$
#
# Note that it is convention to always use the natural log.
#
# i. Interpret the parameter on gasoline price X (or log of gasoline price log(X)) in each of these equations.
#
# ii. What is the predicted per car gasoline consumption in dollars for a country with a gasoline price of 2 in each of these equations?
#
# ➡️ Type your answer to _Exercise 2 (a) i_ here (replacing this text)
# ➡️ Type your answer to _Exercise 2 (a) ii_ here (replacing this text)
# ## Exercise 3. Importing Data into R and Basic R First Commands
#
# For the purposes of this class, we will be primarily Berkeley's _Datahub_ to conduct our analysis remotely using these notebooks.
#
# If instead you already have an installation of R/RStudio on your personal computer and prefer to work offline, you can download the data for this assignment from bCourses (Make sure to install/update all packages mentioned in the problem sets in order to prevent issues regarding deprecated or outdated packages). The data files can be accessed directly through $Datahub$ and do not require you to install anything on your computer. This exercise is designed to get you familiar with accessing the service, loading data, and obtaining summary statistics. To start off, we're going to use Jupyter notebooks to help familiarize you with some R commands.
#
# *Note: [Coding Bootcamp Part 1](https://bcourses.berkeley.edu/courses/1502259/external_tools/70734) covers all necessary R methods.
# (a) To access the Jupyter notebook for this problem set on Datahub, click the following link:
#
# *Skip! You are already here - nice work.*
#
# (b) Load the data file *dataPset1_1960.csv* into R (since this is a ".csv" file, you should use the `read.csv()` command).
# +
# insert code here
# -
# (c) Provide basic summary statistics on the log of Gas Consumption per car (*LGASPCAR*) in the dataframe. Use the `summary()` command. This command is part of base R, so you do not need to load any packages before using it. What is the median value of log gasoline consumption per car?
# +
# insert code here
# -
# (d) Next, generate custom summary statistics on the Log of Gasoline Price Variable(*LRPMG*) using the `summarise()` command provided by ***dplyr***. You will need to call the ***tidyverse*** package with the `library()` command to use it (***tidyverse*** is a collection of packages designed for data science. It includes ***dplyr*** and several other packages we'll use this term).
# +
# insert code here
# -
# (e) Create a scatter plot of the *LGASPCAR* and *LRPMG* data. Use
#
# `figureAsked <- plot(my_data$LRPMG, my_data$LGASPCAR,
# main = "Scatter of Log Y on Log X",
# xlab = "Log(X)=Log of Gas Price",
# ylab = "Log Y = Log of Gas Consumption per Car")`
#
# *Note:* Make sure to run the code cell to print the scatterplot in the notebook.
# +
# insert code here
# -
# (f) Save a pdf to your computer (note: this can be done by going to **File > Download As > PDF Via HTML (.pdf )** in the menu) and combine it with your excel workbook from Exercise 1 for submission on Gradescope.
|
Spring2021-J/Problem-Sets/ProblemSet1/ProblemSet1_2021.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:TestEnv]
# language: python
# name: conda-env-TestEnv-py
# ---
# + [markdown] colab_type="text" id="eNSkSfKvSXEE"
# ## Cosine Similarity Calculations
# Cosine similarity is a measure of similarity between two non-zero vectors of an inner product space that measures the cosine of the angle between them. Similarity measures have a multiude of uses in machine learning projects; they come in handy when matching strings, measuring distance, and extracting features. This similarity measurement is particularly concerned with orientation, rather than magnitude.
# In this case study, you'll use the cosine similarity to compare both a numeric data within a plane and a text dataset for string matching.
# + [markdown] colab_type="text" id="USGZdZYoVu7A"
# Load the Python modules, including cosine_similarity, from sklearn.metrics.pairwise
# + colab={} colab_type="code" id="_T-FTjWuSXEF"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# %matplotlib inline
plt.style.use('ggplot')
from scipy import spatial
from sklearn.metrics.pairwise import cosine_similarity
# + [markdown] colab_type="text" id="woBv3-cmV1R6"
# **<font color='teal'> Load the distance dataset into a dataframe. </font>**
# + colab={} colab_type="code" id="irpsaCuGSXEK"
df= pd.read_csv('distance_dataset.csv')
# -
df.head()
# + [markdown] colab_type="text" id="WWd4xbk1SXEO"
# ### Cosine Similarity with clusters and numeric matrices
#
# All points in our dataset can be thought of as feature vectors. We illustrate it here as we display the __Cosine Similarity__ between each feature vector in the YZ plane and the [5, 5] vector we chose as reference. The sklearn.metrics.pairwise module provides an efficient way to compute the __cosine_similarity__ for large arrays from which we can compute the similarity.
# + [markdown] colab_type="text" id="NeRvhNjLWaau"
# **<font color='teal'> First, create a 2D and a 3D matrix from the dataframe. The 2D matrix should contain the 'Y' and 'Z' columns and the 3D matrix should contain the 'X','Y', and 'Z' columns.</font>**
# + colab={} colab_type="code" id="aWkrVP8dWqZ-"
matYZ = df[['Y','Z']].to_numpy()
mat = df[['X','Y','Z']].to_numpy()
# + [markdown] colab_type="text" id="HlRZpDfzXJR9"
# Calculate the cosine similarity for those matrices with reference planes of 5,5 and 5,5,5. Then subtract those measures from 1 in new features.
# + colab={} colab_type="code" id="QmKPcdIjSXEP"
simCosine3D = 1. - cosine_similarity(mat, [[5,5,5]], 'cosine')
simCosine = 1. - cosine_similarity(matYZ, [[5,5]], 'cosine')
# + [markdown] colab_type="text" id="EieVXBAcX0OU"
# Using the 2D matrix and the reference plane of (5,5) we can use a scatter plot to view the way the similarity is calculated using the Cosine angle.
# + colab={} colab_type="code" id="Oga4-0A6SXEY"
figCosine = plt.figure(figsize=[10,8])
plt.scatter(df.Y, df.Z, c=simCosine[:,0], s=20)
plt.plot([0,5],[0,5], '--', color='dimgray')
plt.plot([0,3],[0,7.2], '--', color='dimgray')
plt.text(0.7,2.6,r'$\theta$ = 22.4 deg.', rotation=47, size=14)
plt.ylim([0,10])
plt.xlim([0,10])
plt.xlabel('Y', size=14)
plt.ylabel('Z', size=14)
plt.title('Cosine Similarity')
cb = plt.colorbar()
cb.set_label('Similarity with (5,5)', size=14)
#figCosine.savefig('similarity-cosine.png')
# + [markdown] colab_type="text" id="h5EERbkYXioh"
# Now, plot the 3D matrix with the similarity and the reference plane, (5,5,5).
# + colab={} colab_type="code" id="-f3ZhgoQSXEV"
from mpl_toolkits.mplot3d import Axes3D
figCosine3D = plt.figure(figsize=(10, 8))
ax = figCosine3D.add_subplot(111, projection='3d')
p = ax.scatter(mat[:,0], mat[:,1], mat[:,2], c=simCosine3D[:,0])
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
cb = figCosine3D.colorbar(p)
cb.set_label('Similarity with (5,5,5)', size=14)
figCosine3D.tight_layout()
#figCosine3D.savefig('cosine-3D.png', dpi=300, transparent=True)
# + [markdown] colab_type="text" id="_ysZ1wFMSXEt"
# ----
# + [markdown] colab_type="text" id="jGJ2vr60SXEu"
# ### Cosine Similarity with text data
# This is a quick example of how you can use Cosine Similarity to compare different text values or names for record matching or other natural language proecessing needs.
# First, we use count vectorizer to create a vector for each unique word in our Document 0 and Document 1.
# + colab={} colab_type="code" id="7oSm7linSXEv"
from sklearn.feature_extraction.text import CountVectorizer
count_vect = CountVectorizer()
Document1 = "Starbucks Coffee"
Document2 = "Essence of Coffee"
corpus = [Document1,Document2]
X_train_counts = count_vect.fit_transform(corpus)
pd.DataFrame(X_train_counts.toarray(),columns=count_vect.get_feature_names(),index=['Document 0','Document 1'])
# + [markdown] colab_type="text" id="W99LBHobSXE1"
# Now, we use a common frequency tool called TF-IDF to convert the vectors to unique measures.
# + colab={} colab_type="code" id="mA52-mj9SXE1"
from sklearn.feature_extraction.text import TfidfVectorizer
vectorizer = TfidfVectorizer()
trsfm=vectorizer.fit_transform(corpus)
pd.DataFrame(trsfm.toarray(),columns=vectorizer.get_feature_names(),index=['Document 0','Document 1'])
# + [markdown] colab_type="text" id="8T8ZOVnLSXE4"
# Here, we finally apply the __Cosine Similarity__ measure to calculate how similar Document 0 is compared to any other document in the corpus. Therefore, the first value of 1 is showing that the Document 0 is 100% similar to Document 0 and 0.26055576 is the similarity measure between Document 0 and Document 1.
# + colab={} colab_type="code" id="ZLbYTQfnSXE4"
cosine_similarity(trsfm[0:1], trsfm)
# + [markdown] colab_type="text" id="uSDf5EgJatlw"
# Replace the current values for `Document 0` and `Document 1` with your own sentence or paragraph and apply the same steps as we did in the above example.
# + [markdown] colab_type="text" id="S9FESS0IbF9I"
# **<font color='teal'> Combine the documents into a corpus.</font>**
# + colab={} colab_type="code" id="7aVkjj-xczS-"
Doc0 = 'The list of ship launches in 1901 includes a chronological list of ships launched in 1901. In cases where no official launching ceremony was held, the date built or completed may be used instead.'
Doc1 = 'This is a list of the governors of colonies, protectorates, or other dependencies in 1901. Where applicable, native rulers are also listed.'
corpus = [Doc0,Doc1]
# + [markdown] colab_type="text" id="Bng2fLV_bR-V"
# **<font color='teal'> Apply the count vectorizer to the corpus to transform it into vectors.</font>**
# + colab={} colab_type="code" id="FtW2_TcJczuh"
count_vect = CountVectorizer()
X_train_counts = count_vect.fit_transform(corpus)
# + [markdown] colab_type="text" id="55i9xWTYbc-r"
# **<font color='teal'> Convert the vector counts to a dataframe with Pandas.</font>**
# + colab={} colab_type="code" id="OaXsheskc0Ql"
pd.DataFrame(X_train_counts.toarray(),columns=count_vect.get_feature_names(),index=['Doc 0','Doc 1'])
# + [markdown] colab_type="text" id="dONg4uupcJAK"
# **<font color='teal'> Apply TF-IDF to convert the vectors to unique frequency measures.</font>**
# + colab={} colab_type="code" id="o2z2p_kMc1mF"
vectorizer = TfidfVectorizer()
trsfm=vectorizer.fit_transform(corpus)
pd.DataFrame(trsfm.toarray(),columns=vectorizer.get_feature_names(),index=['Doc 0','Doc 1'])
# + [markdown] colab_type="text" id="wd1JHVSxcl1D"
# **<font color='teal'> Use the cosine similarity function to get measures of similarity for the sentences or paragraphs in your original document.</font>**
# + colab={} colab_type="code" id="sJdk5sX5bcPO"
cosine_similarity(trsfm[0:1], trsfm)
# -
|
Cosine_Similarity_Case_Study/Cosine_Similarity_Case_Study.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Step #0: Import libraries
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# Step #1: Import the dataset
bank_df = pd.read_csv('Bank_Customer_retirement.csv')
bank_df
bank_df.keys()
bank_df.head(10)
bank_df.tail(10)
# Step #2: Visualize the data
sns.pairplot(bank_df, hue = 'Retire', vars = ['Age','401K Savings'])
sns.countplot(bank_df['Retire'], label = 'Retirement')
# Step #3: Model Training
bank_df = bank_df.drop(['Customer ID'], axis =1)
#Let's drop the target columns
X = bank_df.drop(['Retire'],axis = 1)
Y = bank_df['Retire']
from sklearn.model_selection import train_test_split
X_train, X_test, Y_train, Y_test = train_test_split(X, Y, test_size = 0.2, random_state = 5)
from sklearn.svm import SVC
from sklearn.metrics import classification_report, confusion_matrix
svc_model = SVC()
svc_model.fit(X_train, Y_train)
# Step #4: Evaluating the Model
Y_predict = svc_model.predict(X_test)
cm = confusion_matrix(Y_test, Y_predict)
sns.heatmap(cm, annot= True)
print(classification_report(Y_test, Y_predict))
# Step #5: Improve the Model
#Feature scaling
min_train = X_train.min()
range_train = (X_train - min_train).max()
X_train_scaled = (X_train - min_train) / range_train
X_train_scaled
sns.scatterplot(x =X_train['Age'], y= X_train['401K Savings'], hue = Y_train)
sns.scatterplot(x =X_train_scaled['Age'], y= X_train_scaled['401K Savings'], hue = Y_train)
min_test = X_test.min()
range_test = (X_test - min_test).max()
X_test_scaled = (X_test - min_test) / range_test
X_test_scaled
from sklearn.svm import SVC
from sklearn.metrics import classification_report, confusion_matrix
svc_model = SVC()
svc_model.fit(X_train_scaled, Y_train)
Y_predict = svc_model.predict(X_test_scaled)
cm = confusion_matrix(Y_test, Y_predict)
sns.heatmap(cm, annot = True)
print(classification_report(Y_test, Y_predict))
# Step #5: Improving the Model
param_grid = {'C': [0.1, 1, 10, 100], 'gamma' : [1, 0.1, 0.01, 0.001], 'kernel' : ['rbf']}
from sklearn.model_selection import GridSearchCV
grid = GridSearchCV(SVC(), param_grid, refit = True, verbose = 4)
grid.fit(X_train_scaled, Y_train)
grid.best_param_
grid_predict = grid.predict(X_test_scaled)
cm = confusion_matrix(Y_test, grid_predict)
sns.heatmap(cm, annot= True)
print(classification_report(Y_test, grid_predict))
|
SVM 2- Bank Customer Retirement Predictions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Y4flqGKuVzRE" outputId="07d64f1b-cb95-4487-be63-0a7a85f6afca"
# ! pip install livelossplot
# + id="WAiS6UoSqZ3A"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, OneHotEncoder, MinMaxScaler
from livelossplot import PlotLossesKeras
# + id="4l0vapZzP5XS"
import itertools
from sklearn.metrics import confusion_matrix, classification_report, plot_roc_curve, precision_recall_curve
def plot_confusion_matrix(model, X_test, y_test, normalize=False, cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
y_predict = model.predict(X_test)
cm = confusion_matrix(y_test, y_predict)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
title = 'Normalized confusion matrix'
else:
title = 'Confusion matrix, without normalization'
classes = np.arange(len(model.classes_))
plt.figure()
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45)
plt.yticks(tick_marks, classes)
np.set_printoptions(precision=2)
fmt = '.2f' if normalize else 'd'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
def print_roc(clf, X_test, y_test):
y_pred = clf.predict(X_test)
print(classification_report(y_test, y_pred))
plot_roc_curve(clf, X_test, y_test)
plt.plot([(0,0),(1,1)], '--y')
plt.title('ROC curve')
plt.xlabel('False positive rate')
plt.ylabel('True positive rate')
plt.show()
# + [markdown] id="JI0FlesMNnx6"
# # Data cleaning
# + id="W12B3p4lrCex"
df = pd.read_csv('KaggleV2-May-2016.csv')
df = df.rename(columns={'Hipertension': 'Hypertension', 'Handcap': 'Handicap', 'SMS_received': 'SMSReceived', 'No-show': 'NoShow'})
# + [markdown] id="uJXPdrkXOHDr"
# ## Encoding
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="FGI427WspogP" outputId="d1be8554-5571-45f4-f36c-ac221cddf9f1"
df.head()
# + id="q2UQyV2k5Vzz"
df_t = pd.DataFrame()
le_1 = LabelEncoder()
df_t['Gender'] = pd.Series(le_1.fit_transform(df['Gender']), index=df.index)
df_t['Age'] = df['Age']
df_t['Age'] = pd.Series(le_1.fit_transform(df['Age']), index=df.index)
df['ScheduledDay'] = pd.to_datetime(df['ScheduledDay']).dt.date.astype('datetime64[ns]')
df_t['ScheduledDay'] = df['ScheduledDay'].dt.weekday
df['AppointmentDay'] = pd.to_datetime(df['AppointmentDay']).dt.date.astype('datetime64[ns]')
df_t['AppointmentDay'] = df['AppointmentDay'].dt.weekday
df_t['wait_time'] = (df['AppointmentDay'] - df['ScheduledDay']).dt.days
df_t['Scholarship'] = df['Scholarship']
df_t['Hypertension'] = df['Hypertension']
df_t['Diabetes'] = df['Diabetes']
df_t['Alcoholism'] = df['Alcoholism']
df_t['Handicap'] = df['Handicap']
enc_1 = OneHotEncoder(handle_unknown='ignore')
enc_1.fit(df['Neighbourhood'].to_numpy().reshape(-1,1))
enc_arr = enc_1.transform(df['Neighbourhood'].to_numpy().reshape(-1,1)).toarray()
Neighbourhoods = enc_1.get_feature_names()
df_Neighbourhoods = pd.DataFrame(data=enc_arr, columns=enc_1.get_feature_names(),
index=df.index)
df_t = pd.concat([df_t, df_Neighbourhoods], axis=1)
le_2 = LabelEncoder()
df_t['NoShow'] = le_2.fit_transform(df['NoShow'])
# + colab={"base_uri": "https://localhost:8080/", "height": 287} id="IKccX5Ue46pO" outputId="2eee07f3-3e71-4c44-b272-6435fe49a43d"
for feature in df_t.columns:
df_t[feature] = df_t[feature].astype('int')
df_t.head()
# + id="Dme7PwPnOFs2"
# + [markdown] id="mOdsrRJ_Nu7r"
# # Model Building
# + [markdown] id="TY92zlBXSQTn"
# ## DNN
# + id="5tMIisadSGII"
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Dropout
from tensorflow.keras.callbacks import EarlyStopping
# + colab={"base_uri": "https://localhost:8080/"} id="NTSCNjmCEfwS" outputId="1d6f79e1-2eb1-4fc6-84bc-2444e93ba4d9"
df_t.shape
# + [markdown] id="bDAOuI2CSa_6"
# 91 Input features --
# 1 prediction
# + [markdown] id="JhDuhYCZXRNK"
# ### Feature pre-prossesing
# + id="1qOqjE2oYXeI"
X = df_t.drop(['NoShow'], axis=1)
y = df_t['NoShow']
# + colab={"base_uri": "https://localhost:8080/"} id="1cTBsB0j5GD_" outputId="df5e697b-b52f-4b75-e4fa-12f161e724ae"
scaler = MinMaxScaler()
X = scaler.fit_transform(X)
print(X.min(axis=0))
print(X.max(axis=0))
# + id="qu3UCiacHh28"
import pickle
encoders = [le_1, enc_1, scaler]
with open('encoders.pickle', 'wb') as handle:
pickle.dump(encoders, handle, protocol=4)
# + id="kwghfU2FNHiN"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
# + id="Z1NjMNnFSFit"
layers = []
layers.append(Dense(128, input_dim=91, activation='relu', name='fc_1')) # 1st fully connected layer
layers.append(Dense(64, activation='sigmoid', name='fc_2')) # 2nd connected layer
layers.append(Dropout(0.5, name='drop_out'))
layers.append(Dense(1, activation='sigmoid', name='fc_3')) # 3rd fully connected layer
model = Sequential(layers, name='show/no_show')
# + id="MoKyVHeMQ-D9"
model.compile(
optimizer='adam',
loss='mse',
metrics=['accuracy'])#, 'AUC'])
# + colab={"base_uri": "https://localhost:8080/"} id="zWEH2fqcTCiv" outputId="1a8908a1-59a1-46d7-80d0-45ca1a9706ff"
print(f'input shape: {model.input_shape[1]}')
print(f'out shape: {model.output_shape[1]}\n\n')
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 687} id="LwlxnYqDTG8L" outputId="c2e36690-e26c-4b21-894b-9caed96e21ab"
es = EarlyStopping(monitor='val_loss', min_delta=0, patience=5, verbose=0,
mode='auto', baseline=None, restore_best_weights=True)
history = model.fit(x=X_train, y=y_train, validation_split=0.2,
epochs=100, batch_size=50, verbose=-1,
callbacks=[PlotLossesKeras(), es])
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="y2BBU6S5ZZmG" outputId="045c0028-4ef1-4235-e2d8-ebdefc1aef41"
y_pred = model.predict(X_test)
plt.hist(y_pred);
# + colab={"base_uri": "https://localhost:8080/"} id="nqGQM0mdXuI4" outputId="4c789846-24a1-43f9-bcff-04ed886b7fa6"
y_pred.mean(), y_test.mean()
# + id="WeAofQSWcFVk"
precision, recall, threshold = precision_recall_curve(y_test, y_pred)
# + colab={"base_uri": "https://localhost:8080/", "height": 283} id="VLuzu2sWGlqd" outputId="17551fab-b856-4b50-97d4-623d3e6caf58"
plt.plot(threshold, precision[:-1], "r", label="precision")
plt.plot(threshold, recall[:-1], "b", label="recall")
plt.xlabel("threshold")
plt.legend()
plt.ylim([0, 1])
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="NGvHMTuCIpX-" outputId="3b8e9b95-8d16-48b2-9412-4944ced3573e"
model.save('model')
# + colab={"base_uri": "https://localhost:8080/"} id="2GNGxQIeJqYE" outputId="4f66d1b8-fca5-4166-f020-e03d5762b200"
# ! zip -r model.zip model
|
Data_Science/Doc_oclock_model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The transport-length hillslope diffuser
# # The basics:
#
# This component uses an approach similar to Davy and Lague (2009)'s equation for fluvial erosion and transport, and applies it to hillslope diffusion.
# Formulation and implementation were inspired by Carretier et al (2016), see this paper and references therein for justification.
#
# ## Theory
#
# The elevation z of a point of the landscape (grid node) changes according to:
#
# \begin{equation}
# \frac{\partial z}{\partial t} = -\epsilon + D + U \tag{1}\label{eq:1}
# \end{equation}
#
#
# and we define:
# \begin{equation}
# D = \frac{q_s}{L} \tag{2}\label{eq:2}
# \end{equation}
#
#
#
# where $\epsilon$ is the local erosion rate [*L/T*], *D* the local deposition rate [*L/T*], *U* the uplift (or subsidence) rate [*L/T*], $q_s$ the incoming sediment flux per unit width [*L$^2$/T*] and *L* is the **transport length**.
#
# We specify the erosion rate $\epsilon$ and the transport length *L*:
#
# \begin{equation}
# \epsilon = \kappa S \tag{3}\label{eq:3}
# \end{equation}
#
#
#
# \begin{equation}
# L = \frac{dx}{1-({S}/{S_c})^2} \tag{4}\label{eq:4}
# \end{equation}
#
# where $\kappa$ [*L/T*] is an erodibility coefficient, $S$ is the local slope [*L/L*] and $S_c$ is the critical slope [*L/L*].
#
# Thus, the elevation variation results from the difference between local rates of detachment and deposition.
#
# The detachment rate is proportional to the local gradient. However, the deposition rate (*$q_s$/L*) depends on the local slope and the critical slope:
# - when $S \ll S_c$, most of the sediment entering a node is deposited there, this is the pure diffusion case. In this case, the sediment flux $q_s$ does not include sediment eroded from above and is thus "local".
# - when $S \approx S_c$, *L* becomes infinity and there is no redeposition on the node, the sediments are transferred further downstream. This behaviour corresponds to mass wasting, grains can travel a long distance before being deposited. In that case, the flux $q_s$ is "non-local" as it incorporates sediments that have both been detached locally and transited from upslope.
# - for an intermediate $S$, there is a prgogressive transition between pure creep and "balistic" transport of the material. This is consistent with experiments (Roering et al., 2001; Gabet and Mendoza, 2012).
#
# ## Contrast with the non-linear diffusion model
#
# Previous models typically use a "non-linear" diffusion model proposed by different authors (e.g. Andrews and Hanks, 1985; Hanks, 1999; Roering et al., 1999) and supported by $^{10}$Be-derived erosion rates (e.g. Binnie et al., 2007) or experiments (Roering et al., 2001). It is usually presented in the followin form:
#
# $ $
#
# \begin{equation}
# \frac{\partial z}{\partial t} = \frac{\partial q_s}{\partial x} \tag{5}\label{eq:5}
# \end{equation}
#
# $ $
# \begin{equation}
# q_s = \frac{\kappa' S}{1-({S}/{S_c})^2} \tag{6}\label{eq:6}
# \end{equation}
#
# where $\kappa'$ [*L$^2$/T*] is a diffusion coefficient.
#
# This description is thus based on the definition of a flux of transported sediment parallel to the slope:
# - when the slope is small, this flux refers to diffusion processes such as aoil creep, rain splash or diffuse runoff
# - when the slope gets closer to critical slope, the flux increases dramatically, simulating on average the cumulative effect of mass wasting events.
#
#
# Despite these conceptual differences, Eq ($\ref{eq:3}$) and ($\ref{eq:4}$) predict similar topographic evolution to the 'non-linear' diffusion equations for $\kappa' = \kappa dx$, as shown in the following example.
#
# # Example 1:
# First, we import what we'll need:
# +
# %matplotlib inline
import numpy as np
from matplotlib.pyplot import figure, show, plot, xlabel, ylabel, title
import pymt.models
# -
# Set the initial and run conditions:
# +
total_t = 2000000. # total run time (yr)
dt = 1000. # time step (yr)
nt = int(total_t // dt) # number of time steps
uplift_rate = 0.0001 # uplift rate (m/yr)
kappa = 0.001 # erodibility (m/yr)
Sc = 0.6 # critical slope
# -
# Instantiate the components:
# The hillslope diffusion component must be used together with a flow router/director that provides the steepest downstream slope for each node, with a D4 method (creates the field *topographic__steepest_slope* at nodes).
fdir = pymt.models.FlowDirectorSteepest()
tl_diff = pymt.models.TransportLengthHillslopeDiffuser()
config_file, config_dir = fdir.setup(
grid_row_spacing=10.,
grid_column_spacing=10.,
grid_rows=100,
grid_columns=100,
clock_start=0.0,
clock_stop=total_t,
clock_step=dt,
)
fdir.initialize(config_file, config_dir)
config_file, config_dir = tl_diff.setup(
grid_row_spacing=10.,
grid_column_spacing=10.,
grid_rows=100,
grid_columns=100,
clock_start=0.0,
clock_stop=total_t,
clock_step=dt,
erodibility=kappa,
slope_crit=Sc,
)
tl_diff.initialize(config_file, config_dir)
# Set the boundary conditions. The **FlowDirector** component uses a variable called *boundary_condition_flag* to set its boundary conditions. A value of 1, means the boundary is open and sediment is free to leave the grid. A value of 4 means the nodes are closed and so there is no flux through them. The **TransportLengthHillslopeDiffuser** uses these boundary conditions as input so we'll set them both here.
status = fdir.get_value("boundary_condition_flag").reshape((100, 100))
status[:, (0, -1)] = 1 # E and W boundaries are open
status[(0, -1), :] = 4 # N and S boundaries are closed
fdir.set_value("boundary_condition_flag", status)
tl_diff.set_value("boundary_condition_flag", status)
# Start with an initial surface that's just random noise.
z = np.random.rand(100 * 100)
fdir.set_value("topographic__elevation", z)
# Get the input values for **TransportLengthHillslopeDiffuser** from the flow director.
tl_diff.set_value("topographic__elevation", z)
tl_diff.set_value("flow__receiver_node", fdir.get_value("flow__receiver_node"))
tl_diff.set_value("topographic__steepest_slope", fdir.get_value("topographic__steepest_slope"))
# Run the components for 2 Myr and trace an East-West cross-section of the topography every 100 kyr:
for t in range(nt - 1):
fdir.update()
tl_diff.set_value("topographic__elevation", z)
tl_diff.set_value(
"flow__receiver_node", fdir.get_value("flow__receiver_node")
)
tl_diff.set_value(
"topographic__steepest_slope",
fdir.get_value("topographic__steepest_slope"),
)
tl_diff.update()
z = tl_diff.get_value("topographic__elevation").reshape((100, 100))
z[1:-1, 1:-1] += uplift_rate * dt # add the uplift
fdir.set_value("topographic__elevation", z)
# add some output to let us see we aren't hanging:
if t % 100 == 0:
print(t * dt)
# plot east-west cross-section of topography:
x_plot = range(0, 1000, 10)
z_plot = z[1, :]
figure('cross-section')
plot(x_plot, z_plot)
# And plot final topography:
tl_diff.quick_plot("topographic__elevation")
# # Example 2
#
# In this example, we show that when the slope is steep ($S \geq S_c$), the transport-length hillsope diffusion simulates mass wasting, with long transport distances.
#
# First, we create a grid: the western half of the grid is flat at 0m of elevation, the eastern half is a 45-degree slope.
fdir = pymt.models.FlowDirectorSteepest()
tl_diff = pymt.models.TransportLengthHillslopeDiffuser()
# +
total_t = 1000000. # total run time (yr)
dt = 1000. # time step (yr)
nt = int(total_t // dt) # number of time steps
kappa = 0.001 # erodibility (m / yr)
Sc = 0.6 # critical slope
# -
grid_params = {
"grid_row_spacing": 10.,
"grid_column_spacing": 10.,
"grid_rows": 100,
"grid_columns": 100,
}
clock_params = {
"clock_start": 0.0,
"clock_stop": total_t,
"clock_step": dt,
}
config_file, config_dir = fdir.setup(**grid_params, **clock_params)
fdir.initialize(config_file, config_dir)
config_file, config_dir = tl_diff.setup(
**grid_params,
**clock_params,
erodibility=kappa,
slope_crit=Sc,
)
tl_diff.initialize(config_file, config_dir)
# As before, set the boundary conditions for both components.
status = fdir.get_value("boundary_condition_flag").reshape((100, 100))
status[:, (0, -1)] = 1 # E and W boundaries are open
status[(0, -1), :] = 4 # N and S boundaries are closed
fdir.set_value("boundary_condition_flag", status)
tl_diff.set_value("boundary_condition_flag", status)
# In this example, we'll use a different initial surface: a dipping plane.
grid = fdir.var_grid("topographic__elevation")
n_vals = fdir.grid_size(grid)
x, y = fdir.grid[0].node_x, fdir.grid[0].node_y
z = np.zeros(n_vals)
z[x > 500] = x[x < 490] / 10.0
# To make sure we've set things up correctly.
fdir.set_value("topographic__elevation", z)
fdir.quick_plot("topographic__elevation")
# Now time step through the model, plotting things along the way.
for t in range(1000):
fdir.update()
tl_diff.set_value("topographic__elevation", fdir.get_value("topographic__elevation"))
tl_diff.set_value("flow__receiver_node", fdir.get_value("flow__receiver_node"))
tl_diff.set_value("topographic__steepest_slope", fdir.get_value("topographic__steepest_slope"))
tl_diff.update()
fdir.set_value("topographic__elevation", tl_diff.get_value("topographic__elevation"))
# add some output to let us see we aren't hanging:
if t % 100 == 0:
print(t * dt)
z = tl_diff.get_value("topographic__elevation").reshape((100, 100))
# plot east-west cross-section of topography:
x_plot = range(0, 1000, 10)
z_plot = z[1, :]
figure('cross-section')
plot(x_plot, z_plot)
fdir.quick_plot("topographic__elevation")
# The material is diffused from the top and along the slope and it accumulates at the bottom, where the topography flattens.
# # Example 3
#
# As a comparison, the following code uses linear diffusion on the same slope. Instead of using the **TransportLengthHillslopeDiffuser** component, we'll swap in the **LinearDiffuser** component. Everything else will be pretty much the same.
fdir = pymt.models.FlowDirectorSteepest()
diff = pymt.models.LinearDiffuser()
# Setup and initialize the models.
config_file, config_dir = fdir.setup(**grid_params, **clock_params)
fdir.initialize(config_file, config_dir)
config_file, config_dir = diff.setup(
**grid_params,
**clock_params,
linear_diffusivity=0.1,
)
diff.initialize(config_file, config_dir)
# Set boundary conditions.
status = fdir.get_value("boundary_condition_flag").reshape((100, 100))
status[:, (0, -1)] = 1 # E and W boundaries are open
status[(0, -1), :] = 4 # N and S boundaries are closed
fdir.set_value("boundary_condition_flag", status)
diff.set_value("boundary_condition_flag", status)
# Set the initial topography.
grid = fdir.var_grid("topographic__elevation")
n_vals = fdir.grid_node_count(grid)
x, y = fdir.grid[0].node_x, fdir.grid[0].node_y
z = np.zeros(n_vals)
z[x > 500] = x[x < 490] / 10.0
fdir.set_value("topographic__elevation", z)
fdir.quick_plot("topographic__elevation")
# Run the model!
for t in range(1000):
fdir.update()
diff.set_value("topographic__elevation", fdir.get_value("topographic__elevation"))
diff.update()
fdir.set_value("topographic__elevation", diff.get_value("topographic__elevation"))
# add some output to let us see we aren't hanging:
if t % 100 == 0:
print(t * dt)
z = diff.get_value("topographic__elevation").reshape((100, 100))
# plot east-west cross-section of topography:
x_plot = range(0, 1000, 10)
z_plot = z[1, :]
figure('cross-section')
plot(x_plot, z_plot)
fdir.quick_plot("topographic__elevation")
|
nb/transport_length_hillslope_diffuser.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pphongsopa/MQP2019/blob/master/Pavee/qlearning_sample.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="TJSUsM1DsNFa" colab_type="code" colab={}
import numpy as np
import pylab as plt
# + id="vBHD-VcDDbC-" colab_type="code" colab={}
# map cell to cell, add circular cell to goal point
points_list = [(0,1), (1,5), (5,6), (5,4), (1,2), (2,3), (2,7)]
# + id="qIjGsDpyDdPW" colab_type="code" outputId="e5b93e58-af8b-46e2-c6e7-d49fa1641fd3" colab={"base_uri": "https://localhost:8080/", "height": 248}
goal = 7
import networkx as nx
G=nx.Graph()
G.add_edges_from(points_list)
pos = nx.spring_layout(G)
nx.draw_networkx_nodes(G,pos)
nx.draw_networkx_edges(G,pos)
nx.draw_networkx_labels(G,pos)
plt.show()
# + id="G9_JNGs2DdXv" colab_type="code" colab={}
# 8 points in the graph
MATRIX_SIZE = 8
# create matrix x*y
R = np.matrix(np.ones(shape=(MATRIX_SIZE, MATRIX_SIZE)))
R *= -1
# + id="kV5Tg4tODdad" colab_type="code" outputId="eb33f1cb-5524-4fc0-ac6f-2e9aecf199ca" colab={"base_uri": "https://localhost:8080/", "height": 134}
# assign zeros to paths and 100 to goal-reaching point
for point in points_list:
print(point)
if point[1] == goal:
R[point] = 100
else:
R[point] = 0
if point[0] == goal:
R[point[::-1]] = 100
else:
# reverse of point
R[point[::-1]]= 0
# add goal point round trip
R[goal,goal]= 100
# + id="N9f4fSVSDdcu" colab_type="code" colab={}
Q = np.matrix(np.zeros([MATRIX_SIZE,MATRIX_SIZE]))
# + id="qZm0JRztDwqL" colab_type="code" outputId="c332bda5-d581-40c8-8189-c165520b354f" colab={"base_uri": "https://localhost:8080/", "height": 34}
# learning parameter
gamma = 0.8
initial_state = 1
def available_actions(state):
current_state_row = R[state,]
av_act = np.where(current_state_row >= 0)[1]
return av_act
available_act = available_actions(initial_state)
def sample_next_action(available_actions_range):
next_action = int(np.random.choice(available_act,1))
return next_action
action = sample_next_action(available_act)
def update(current_state, action, gamma):
max_index = np.where(Q[action,] == np.max(Q[action,]))[1]
if max_index.shape[0] > 1:
max_index = int(np.random.choice(max_index, size = 1))
else:
max_index = int(max_index)
max_value = Q[action, max_index]
Q[current_state, action] = R[current_state, action] + gamma * max_value
#print('max_value', R[current_state, action] + gamma * max_value)
if (np.max(Q) > 0):
return(np.sum(Q/np.max(Q)*100))
else:
return (0)
update(initial_state, action, gamma)
# + id="XLlO4ffiHLq2" colab_type="code" outputId="4c3dec18-56b0-41e2-8a4e-a537b3531b6b" colab={"base_uri": "https://localhost:8080/", "height": 302}
# Training
scores = []
for i in range(700):
current_state = np.random.randint(0, int(Q.shape[0]))
available_act = available_actions(current_state)
action = sample_next_action(available_act)
score = update(current_state,action,gamma)
scores.append(score)
#print ('Score:', str(score))
print("Trained Q matrix:")
print(Q/np.max(Q)*100)
# + id="_SQEHX11HN8I" colab_type="code" outputId="547f3ba8-2c60-4f2a-a15f-f623d8f6742e" colab={"base_uri": "https://localhost:8080/", "height": 298}
# Testing
current_state = 0
steps = [current_state]
while current_state != 7:
next_step_index = np.where(Q[current_state,] == np.max(Q[current_state,]))[1]
if next_step_index.shape[0] > 1:
next_step_index = int(np.random.choice(next_step_index, size = 1))
else:
next_step_index = int(next_step_index)
steps.append(next_step_index)
current_state = next_step_index
print("Most efficient path:")
print(steps)
plt.plot(scores)
plt.show()
|
Pavee/qlearning_sample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assumptions
#
# - Population counts are taken at week 40 of each season
# - The population counts by age do not change significantly between the time when the population count is taken and when the breakpoints for age happen for sampling.
# - 0 year olds represent people > 6 mo old
#
# +
import pandas as pd
from matplotlib import pyplot as plt
import seaborn as sns
import numpy as np
from datetime import datetime, date
# %matplotlib inline
start_week = 40
pan_start = 15
# These are the breakpoints for which age was defined for a given season. I.e., if the breakpoint is 2007-01-01
# that means that all individuals need to be > 6 months old on or beofre 2007-01-01.
inferred_beakpoints = {'2008': '2008-01-01',
'2009': '2008-12-01',
'2009Pan': '2009-05-01',
'2010': '2009-12-01',
'2011': '2011-01-01',
'2012': '2011-09-01',
'2013': '2012-09-01',
'2014': '2013-09-01',
'2015': '2014-09-01',
'2016': '2015-09-01',
'2017': '2016-09-01',
'2018': '2017-09-01'}
eligible_demo = pd.read_csv('../raw_data/demography_by_age.csv')
all_seasons = list(range(2008, 2019)) + [2009.5]
index = pd.MultiIndex.from_product([range(1890, 2019), all_seasons], names=['Birth_year', 'Season'])
converted_df = pd.DataFrame(0,
index = index,
columns = ['Population'])
index2 = pd.MultiIndex.from_product([set(eligible_demo.Age), all_seasons], names=['Age', 'Season'])
extended_df = pd.DataFrame(index = index2)
for index, row in eligible_demo.iterrows():
if row.Year > 2007:
age = row.Age
pop = row.MESA_pop
week_of_eligibility = start_week
day_of_eligibility = inferred_beakpoints[str(int(row.Year))]
start = datetime.strptime(day_of_eligibility, '%Y-%m-%d')
year = start.year
ordinal_start_day = (start.date() - date(year, 1, 1)).days + 1
year_length_days = (date(year, 12, 31) - date(year, 1, 1)).days + 1
birth_year_1 = int(year - age)
birth_year_2 = int(year - age - 1)
if age != 0:
frac_1 = ordinal_start_day / year_length_days
else: # if the age is 0 then the eligible cohort only consists of children > 6 mo old
frac_1 = (ordinal_start_day - 183)/183
if frac_1 < 0:
frac_1 = 0
season = row.Year
converted_df.loc[(birth_year_1, season), 'Population'] += frac_1 * pop
converted_df.loc[(birth_year_1, season), 'a1_pop'] = frac_1 * pop
converted_df.loc[(birth_year_1, season), 'a1'] = age
converted_df.loc[(birth_year_2, season), 'Population'] += pop - (frac_1 * pop)
converted_df.loc[(birth_year_2, season), 'a2_pop'] = pop - (frac_1 * pop)
converted_df.loc[(birth_year_2, season), 'a2'] = age
extended_df.loc[(age, season), 'Population'] = pop
extended_df.loc[(age, season), 'y1'] = birth_year_1
extended_df.loc[(age, season), 'y2'] = birth_year_2
extended_df.loc[(age, season), 'f1'] = frac_1
extended_df.loc[(age, season), 'f2'] = (1-frac_1)
# Add in pandemic, we assume that the age distribution does not change
# between 2008-2009 and the pandemic
for index, row in eligible_demo.iterrows():
if row.Year == 2009:
age = row.Age
pop = row.MESA_pop
week_of_eligibility = start_week
day_of_eligibility = inferred_beakpoints['2009Pan']
start = datetime.strptime(day_of_eligibility, '%Y-%m-%d')
year = start.year
ordinal_start_day = (start.date() - date(year, 1, 1)).days + 1
year_length_days = (date(year, 12, 31) - date(year, 1, 1)).days + 1
birth_year_1 = int(year - age)
birth_year_2 = int(year - age - 1)
if age != 0:
frac_1 = ordinal_start_day / year_length_days
else: # if the age is 0 then the eligible cohort only consists of children > 6 mo old
frac_1 = (ordinal_start_day - 183)/183
if frac_1 < 0:
frac_1 = 0
converted_df.loc[(birth_year_1, 2009.5), 'Population'] += frac_1 * pop
converted_df.loc[(birth_year_1, 2009.5), 'a1_pop'] = frac_1 * pop
converted_df.loc[(birth_year_1, 2009.5), 'a1'] = age
converted_df.loc[(birth_year_2, 2009.5), 'Population'] += pop - (frac_1 * pop)
converted_df.loc[(birth_year_2, 2009.5), 'a2_pop'] = pop - (frac_1 * pop)
converted_df.loc[(birth_year_2, 2009.5), 'a2'] = age
extended_df.loc[(age, 2009.5), 'Population'] = pop
extended_df.loc[(age, 2009.5), 'y1'] = birth_year_1
extended_df.loc[(age, 2009.5), 'y2'] = birth_year_2
extended_df.loc[(age, 2009.5), 'f1'] = frac_1
extended_df.loc[(age, 2009.5), 'f2'] = (1-frac_1)
converted_df = converted_df[(converted_df.Population > 0) &
(converted_df.index.get_level_values('Birth_year') >= 1918)]
converted_df['f1'] = converted_df.a1_pop / converted_df.Population
converted_df['f2'] = 1.0 - converted_df.f1
# -
converted_df = converted_df.sort_values(['Birth_year', 'Season'])
converted_df.to_csv('../data/demography_by_birth_year.csv')
extended_df.to_csv('../data/demography_by_age_extended.csv')
|
data_processing/convert_demography_to_birth_year.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
from skmultilearn.problem_transform import ClassifierChain
from sklearn.svm import SVC
from sklearn.model_selection import KFold
###Laod data
df = pd.read_csv('../data/Finalplfam_id_Multilabel_Ecoli_data.csv', dtype=str, index_col=0)
Labeldf = pd.read_csv('../data/'+'AMR_LAbel_EColi.csv',index_col=0)
Ycolumns=Labeldf.columns.values
# +
Y=df[Ycolumns]
Y=Y.drop(columns=['genome_id', 'genome_name','taxon_id'])
X=df.drop(columns=Ycolumns)
# -
Labellist=Ycolumns.tolist()
Labellist.remove('genome_id')
Labellist.remove('genome_name')
Labellist.remove('taxon_id')
kfold = KFold(n_splits=5)
scorelist=[]
Y=Y.replace(to_replace=['Not defined','Susceptible-dose dependent'], value=[np.nan,0])
X=X.fillna(0)
X = X.apply(pd.to_numeric)
def hamming_score(y_pred,y_true):
i=0
scorelist=[]
for amrtrue in y_true:
match=0.0
total=0.0
#print(amrtrue, y_pred[i])
j=0
for trueentry in amrtrue:
#print(trueentry)
if (trueentry == y_pred[i][j]):
#print ('match found')
match=match+1
total=total+1
elif (not (np.isnan(trueentry))):
total = total + 1
j=j+1
i=i+1
#print (match, total, (match/total))
scorelist.append(match/total)
return np.mean(scorelist)
import numpy as np
def Modified_F1_Score(y_pred,y_true):
#i=0
scorelist=[]
for i,amrtrue in enumerate(y_true):
TP,FP,TN,FN=0.00000001,0.00000001,0.00000001,0.00000001
#j=0
for j,trueentry in enumerate(amrtrue):
if ((np.isnan(trueentry))):
continue
elif ((trueentry == 1) and (y_pred[i][j]==1)):
TP=TP+1
elif ((trueentry == 1) and (y_pred[i][j]==0)):
FN=FN+1
elif ((trueentry == 0) and (y_pred[i][j]==0)):
TN=TN+1
elif ((trueentry == 0) and (y_pred[i][j]==1)):
FP=FP+1
#j=j+1
#i=i+1
precision=TP/(TP+FP)
recall=TP/(TP+FN)
fscore=(2*precision*recall)/(precision+recall)
scorelist.append(fscore)
return np.mean(scorelist)
# +
import matplotlib.pyplot as plt
import seaborn as sb
sb.set_context("talk")
def plot_coefficients(feature_names, coef, name, top_features=20):
top_positive_coefficients = np.argsort(coef)[-top_features:]
#top_negative_coefficients = np.argsort(coef)[:top_features]
top_coefficients =top_positive_coefficients# np.hstack([top_positive_coefficients])
# create plot
plt.figure(figsize=(10, 5))
plt.title("Feature Importances Multi AMR for "+str(name), y=1.08)
colors = ['crimson' if c < 0 else 'cornflowerblue' for c in coef[top_coefficients]]
plt.bar(np.arange(1 * top_features), coef[top_coefficients], color=colors)
feature_names = np.array(feature_names)
plt.xticks(np.arange(0, 1 * top_features), feature_names[top_coefficients], rotation=60, ha='right')
plt.show()
np.asarray(feature_names)[top_positive_coefficients]
# -
from xgboost import XGBClassifier
from sklearn.ensemble import AdaBoostClassifier
####Base models
xgbmodel = XGBClassifier()
adaboostmodel = AdaBoostClassifier(n_estimators=100, random_state=0)
modellist=[xgbmodel,adaboostmodel]
namelist=['xgb','AdaBoost']
import time
times=[]
methods=[]
from skmultilearn.problem_transform import BinaryRelevance
def BinaryRelevanceModel(X,Y,kfold, basemodel, name, times, methods):
classifier = BinaryRelevance(basemodel)
hammingscorelist=[]
fscorelist=[]
for train_index, test_index in kfold.split(X, Y):
start = time.process_time()
# split data into train/test sets
x_train_tfidf = X.iloc[train_index]
y_train_tfidf = Y.iloc[train_index]
x_test_tfidf = X.iloc[test_index]
y_test_tfidf = Y.iloc[test_index]
y_train_tfidf=y_train_tfidf.fillna(0)
y_train_tfidf = y_train_tfidf.apply(pd.to_numeric)
y_train_tfidf=y_train_tfidf.astype(int)
y_test_tfidf = y_test_tfidf.apply(pd.to_numeric)
#y_test_tfidf=y_test_tfidf.astype(int)
classifier.fit(x_train_tfidf.values, y_train_tfidf.values)
# calculating test accuracy
prediction = classifier.predict(x_test_tfidf.values)
hammingscore=hamming_score(prediction.toarray(),y_test_tfidf.values)
fscore=Modified_F1_Score(prediction.toarray(),y_test_tfidf.values)
print(fscore, hammingscore)
hammingscorelist.append(hammingscore)
fscorelist.append(fscore)
times = np.append(times, (time.process_time() - start))
methods = np.append(methods, 'BR-'+name)
print ('Base model for BR is {}'.format(name))
print('Test Hamming accuracy for Binary relvance is {}'.format(np.mean(hammingscorelist)))
print('Test F-score accuracy for Binary relvance is {}'.format(np.mean(fscorelist)))
print('Hamming std'+str(np.std(hammingscorelist)))
print('F-Score std'+str(np.std(fscorelist)))
return np.mean(hammingscorelist),np.mean(fscorelist),np.std(hammingscorelist),np.std(fscorelist), times, methods
i=0
for basemodel in modellist:
hamming, fscore, hmstd, fstd, times, methods=BinaryRelevanceModel(X,Y,kfold,basemodel,namelist[i], times, methods)
i=i+1
print("Before removing low variance: ", X.shape)
from sklearn.feature_selection import VarianceThreshold
selector = VarianceThreshold(threshold=0.01)
selector.fit_transform(X)
X = X[X.columns[selector.get_support()]].copy()
print("After removing low variance: ", X.shape)
modellist=[xgbmodel]
namelist=['xgb']
from RectifiedClassifierChain import RectifiedClassiferChain
import shap
def RCCModelwithMR(X,Y,kfold, basemodel, name, times, methods,shapTotal, type=0):
sim_all_df = pd.DataFrame()
hammingscorelist=[]
fscorelist=[]
for train_index, test_index in kfold.split(X, Y):
# split data into train/test sets
start = time.process_time()
x_train_tfidf = X.iloc[train_index]
y_train_tfidf = Y.iloc[train_index]
x_test_tfidf = X.iloc[test_index]
y_test_tfidf = Y.iloc[test_index]
x_train_tfidf=x_train_tfidf.reset_index(drop=True)
y_train_tfidf=y_train_tfidf.reset_index(drop=True)
y_train_tfidf = y_train_tfidf.apply(pd.to_numeric)
#y_train_tfidf=y_train_tfidf.astype(int)
y_test_tfidf = y_test_tfidf.apply(pd.to_numeric)
classifier=RectifiedClassiferChain(basemodel,optimized=True,optimizedmethod='MissingRatio')
classifier.trainRCC(x_train_tfidf, y_train_tfidf)
# calculating test accuracy
x_test_tfidf=x_test_tfidf.reset_index(drop=True)
y_test_tfidf=y_test_tfidf.reset_index(drop=True)
prediction = classifier.predictRCC(x_test_tfidf)
hammingscore, fscore=classifier.Evaluate(y_test_tfidf,prediction)
#hammingscore1=hamming_score(np.array(prediction),y_test_tfidf.values)
#fscore1=Modified_F1_Score(np.array(prediction),y_test_tfidf.values)
print(hammingscore,fscore)
#print(hammingscore1,fscore1)
hammingscorelist.append(hammingscore)
fscorelist.append(fscore)
times = np.append(times, (time.process_time() - start))
methods = np.append(methods, 'RCC_MR-'+name)
label_order=classifier.getOptimizedLabelOrder()
print (label_order)
if(type !=2):
featuredf=classifier.getFeature(NoOfFeature=100,type=type,full=True)
#featuredf.to_csv('Test.csv')
sim_all_df = pd.concat([sim_all_df, featuredf], ignore_index=True)
if(type==3):
shapValue=classifier.getShapFeatures()
shapTotal.append(shapValue)
print ('Base model for RCC is {}'.format(name))
print('Test Hamming accuracy for RCC is {}'.format(np.mean(hammingscorelist)))
print('Test F-score accuracy for RCC is {}'.format(np.mean(fscorelist)))
print('Hamming std'+str(np.std(hammingscorelist)))
print('F-Score std'+str(np.std(fscorelist)))
if(type !=2):
sim_all_df_T = sim_all_df.transpose().copy()
sim_all_df_T["feature_weight_sum"] = sim_all_df_T.apply(lambda x: abs(x).sum(), axis=1)
sim_all_df_T_top = sim_all_df_T.sort_values("feature_weight_sum", ascending=False)[:30]
#sim_all_df_T_top.to_csv('Test.csv')
else:
sim_all_df_T_top=sim_all_df.copy()
return np.mean(hammingscorelist),np.mean(fscorelist),sim_all_df_T_top, times, methods, shapTotal
# +
i=0
for basemodel in modellist:
if(namelist[i] =='AdaBoost'):
type=1
elif(namelist[i] =='xgb'):
type=3
elif(namelist[i] =='Gaussian'):
type=2
else:
type=0
hamming, fscore, featuredf, times, methods, TotShape=RCCModelwithMR(X,Y,kfold,basemodel,namelist[i], times, methods, TotShape, type)
if(type!=2):
featuredf.to_csv('sim_all_df_T_RCC_MR_Sal'+str(namelist[i])+'.csv')
#print(featuredf)
plot_coefficients(list(featuredf.index),featuredf["feature_weight_sum"],namelist[i],30)
if (type==3):
TotShapedf = pd.DataFrame (np.vstack(TotShape), columns = [X.columns])
shap.summary_plot(np.vstack(TotShape), features=X, feature_names=X.columns, plot_type="bar")
TotShapedf.to_csv('Shape_Features_RCC_MR_all_Sal_'+str(namelist[i])+'.csv')
i=i+1
# -
from StackedClassifierChain import StackedClassifierChain
def SCCModelwithSSC(X,Y,kfold, basemodel, name, times, methods, type=0):
sim_all_df = pd.DataFrame()
hammingscorelist=[]
fscorelist=[]
for train_index, test_index in kfold.split(X, Y):
# split data into train/test sets
start = time.process_time()
x_train_tfidf = X.iloc[train_index]
y_train_tfidf = Y.iloc[train_index]
x_test_tfidf = X.iloc[test_index]
y_test_tfidf = Y.iloc[test_index]
x_train_tfidf=x_train_tfidf.reset_index(drop=True)
y_train_tfidf=y_train_tfidf.reset_index(drop=True)
y_train_tfidf=y_train_tfidf.fillna(0)
y_train_tfidf = y_train_tfidf.apply(pd.to_numeric)
y_train_tfidf=y_train_tfidf.astype(int)
y_test_tfidf = y_test_tfidf.apply(pd.to_numeric)
classifier=StackedClassifierChain(basemodel)
classifier.trainSCC(x_train_tfidf, y_train_tfidf)
# calculating test accuracy
x_test_tfidf=x_test_tfidf.reset_index(drop=True)
y_test_tfidf=y_test_tfidf.reset_index(drop=True)
prediction = classifier.predictSCC(x_test_tfidf)
hammingscore, fscore=classifier.Evaluate(y_test_tfidf,prediction)
#hammingscore1=hamming_score(np.array(prediction),y_test_tfidf.values)
#fscore1=Modified_F1_Score(np.array(prediction),y_test_tfidf.values)
print(hammingscore,fscore)
#print(hammingscore1,fscore1)
hammingscorelist.append(hammingscore)
fscorelist.append(fscore)
times = np.append(times, (time.process_time() - start))
methods = np.append(methods, 'SCC-'+name)
if(type !=2):
featuredf=classifier.getFeature(NoOfFeature=100,type=type,full=True)
#featuredf.to_csv('Test.csv')
sim_all_df = pd.concat([sim_all_df, featuredf], ignore_index=True)
print ('Base model for SCC is {}'.format(name))
print('Test Hamming accuracy for SCC is {}'.format(np.mean(hammingscorelist)))
print('Test F-score accuracy for SCC is {}'.format(np.mean(fscorelist)))
print('Hamming std'+str(np.std(hammingscorelist)))
print('F-Score std'+str(np.std(fscorelist)))
if(type !=2):
sim_all_df_T = sim_all_df.transpose().copy()
sim_all_df_T["feature_weight_sum"] = sim_all_df_T.apply(lambda x: abs(x).sum(), axis=1)
sim_all_df_T_top = sim_all_df_T.sort_values("feature_weight_sum", ascending=False)[:30]
sim_all_df_T_top.to_csv('SCCTest.csv')
else:
sim_all_df_T_top=sim_all_df.copy()
return np.mean(hammingscorelist),np.mean(fscorelist),sim_all_df_T_top, times, methods
i=0
for basemodel in modellist:
if(namelist[i] =='randomforest' or namelist[i] =='xgb' or namelist[i]=='AdaBoost'):
type=1
elif(namelist[i] =='Gaussian'):
type=2
else:
type=0
hamming, fscore, featuredf, times, methods=SCCModelwithSSC(X,Y,kfold,basemodel,namelist[i], times, methods,type)
if(type!=2):
featuredf.to_csv('sim_all_df_T_SCC_SSC_Sal'+str(namelist[i])+'.csv')
#print(featuredf)
plot_coefficients(list(featuredf.index),featuredf["feature_weight_sum"],namelist[i],30)
i=i+1
'''
XGB -A tree depth of 16 , 10-mers were used to build models from assembled genomes to ensure that all models would fit in memory
learning rate was set to 0.0625
column and row subsampling was set to 1.0
number of rounds of boosting was limited to 1000
'''
'''
AdaBoost
K=31
'''
'''
##https://journals.asm.org/doi/full/10.1128/JCM.01260-18
https://github.com/PATRIC3/mic_prediction
###https://github.com/TahaAslani/AAk-mer
https://www.mdpi.com/2079-7737/9/11/365/htm
##https://journals.plos.org/ploscompbiol/article?id=10.1371/journal.pcbi.1008319
'''
import shap
#explainer=shap.TreeExplainer(rf_classifier)
rf_shap_values = shap.KernelExplainer(rf_classifier.predict,X_test)
shap.initjs()
prediction=rf_classifier.predict(X_test)
shap.force_plot(rf_shap_values.expected_value,shap_values,X_test)
KErnalExplnanier = shap.TreeExplainer(classifer)
svm_shap_values = KErnalExplnanier.shap_values(X_test)
from skmultilearn.problem_transform import ClassifierChain
# +
# #!pip install shap
# -
|
source/Compare_With_SoA_Work.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import time
import numpy as np
import matplotlib.pyplot as plt
from tensorflow.examples.tutorials.mnist import input_data
import tensorflow as tf
def sample_noise(m, n):
return np.random.uniform(-1, 1, size = [m, n])
class Model:
def __init__(self, learning_rate):
self.X = tf.placeholder(tf.float32, shape = [None, 784])
self.discriminator_W = tf.Variable(tf.random_normal([784, 128], stddev = 0.1))
self.discriminator_b = tf.Variable(tf.zeros([128]))
self.discriminator_W_out = tf.Variable(tf.random_normal([128, 1], stddev = 0.1))
self.discriminator_b_out = tf.Variable(tf.zeros([1]))
backpropagate_discriminator = [self.discriminator_W, self.discriminator_b, self.discriminator_W_out, self.discriminator_b_out]
self.Z = tf.placeholder(tf.float32, shape = [None, 100])
self.generator_W = tf.Variable(tf.random_normal([100, 128], stddev = 0.1))
self.generator_b = tf.Variable(tf.zeros([128]))
self.generator_W_out = tf.Variable(tf.random_normal([128, 784], stddev = 0.1))
self.generator_b_out = tf.Variable(tf.zeros([784]))
backpropagate_generator = [self.generator_W, self.generator_W_out, self.generator_b, self.generator_b_out]
def discriminator(z):
discriminator_hidden1 = tf.nn.relu(tf.matmul(z, self.discriminator_W) + self.discriminator_b)
discriminator_out = tf.matmul(discriminator_hidden1, self.discriminator_W_out) + self.discriminator_b_out
return discriminator_out
def generator(z):
generator_hidden1 = tf.nn.relu(tf.matmul(z, self.generator_W) + self.generator_b)
generator_out = tf.matmul(generator_hidden1, self.generator_W_out) + self.generator_b_out
return tf.nn.sigmoid(generator_out)
self.generator_sample = generator(self.Z)
discriminator_real = discriminator(self.X)
discriminator_fake = discriminator(self.generator_sample)
discriminator_loss_real = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits = discriminator_real, labels = tf.ones_like(discriminator_real)))
discriminator_loss_fake = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits = discriminator_fake, labels = tf.zeros_like(discriminator_fake)))
self.discriminator_total_loss = discriminator_loss_real + discriminator_loss_fake
self.generator_loss = tf.reduce_mean(tf.nn.sigmoid_cross_entropy_with_logits(logits = discriminator_fake, labels = tf.ones_like(discriminator_fake)))
self.optimizer_discriminator = tf.train.AdamOptimizer(learning_rate).minimize(self.discriminator_total_loss, var_list = backpropagate_discriminator)
self.optimizer_generator = tf.train.AdamOptimizer(learning_rate).minimize(self.generator_loss, var_list = backpropagate_generator)
# +
sample_size_output = 16
sample_size_train = 128
learning_rate = 0.001
epoch = 30000
mnist = input_data.read_data_sets('/home/huseinzol05/Documents/MNIST/MNIST_data')
# +
EPOCH = []; DISCRIMINATOR_LOSS = []; GENERATOR_LOSS = []
sess = tf.InteractiveSession()
model = Model(learning_rate)
sess.run(tf.global_variables_initializer())
for i in xrange(epoch):
EPOCH.append(i)
last_time = time.time()
input_images, _ = mnist.train.next_batch(sample_size_train)
_, discriminator_loss = sess.run([model.optimizer_discriminator, model.discriminator_total_loss], feed_dict = {model.X : input_images, model.Z : sample_noise(sample_size_train, 100)})
_, generator_loss = sess.run([model.optimizer_generator, model.generator_loss], feed_dict = {model.Z : sample_noise(sample_size_train, 100)})
DISCRIMINATOR_LOSS.append(discriminator_loss); GENERATOR_LOSS.append(generator_loss)
if (i + 1) % 5000 == 0:
print "epoch: " + str(i + 1) + ", discriminator loss: " + str(discriminator_loss) + ", generator loss: " + str(generator_loss) + ", s / 5k epoch: " + str(time.time() - last_time)
fig = plt.figure(figsize = (4, 4))
samples = sess.run(model.generator_sample, feed_dict = {model.Z: sample_noise(sample_size_output, 100)})
for z in xrange(sample_size_output):
plt.subplot(sample_size_output / 4, 4, z + 1)
plt.imshow(samples[z].reshape(28, 28), cmap = 'Greys_r')
plt.show()
# -
import seaborn as sns
sns.set()
fig = plt.figure(figsize = (5, 5))
plt.plot(EPOCH, DISCRIMINATOR_LOSS, label = 'discriminator loss')
plt.plot(EPOCH, GENERATOR_LOSS, label = 'generator loss')
plt.xlabel('epoch'); plt.ylabel('loss')
plt.legend()
plt.show()
|
GAN/GAN/basicgan.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
from importer_snippets import load_experiment
from scipy.signal import find_peaks, savgol_filter
import matplotlib.pyplot as plt
#plt.style.use('JGW')
import os
os.getcwd()
# + [markdown] pycharm={"name": "#%% md\n"}
# # Cyclic Voltammograms
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
CV = load_experiment()
# + jupyter={"outputs_hidden": false} pycharm={"is_executing": true, "name": "#%%\n"}
# CV
fig = plt.figure()
ax = fig.add_subplot()
# ax.set_title('Cyclic Voltammogram')
ax.tick_params(axis='both', which='both', direction='in', right=True, top=True)
# ax.set_prop_cycle(cc2)
ax.scatter(CV.data['Ecell'], CV.data['iw'], label = '$v=$ '+str(CV.params.loc[CV.params['parameter']=='scan_rate', 'value'].item()) + ' V/s')
ax.set_xlabel('$E_{cell}$ / V')
ax.set_ylabel('$i_{w}$ / $A$')
ax.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
plt.style.use('JGW')
fig.legend()
# -
# ### Find CV Peaks
# + pycharm={"is_executing": true}
# Smooth data using the Savitski Golay method
smooth_iw = savgol_filter(CV.data['iw'], 25, 2)
# Check the result
# plt.plot(CV.data['Ecell'], smooth_iw, zorder=1, color='r')
# plt.plot(CV.data['Ecell'], CV.data['iw'], label = '$v=$ '+str(CV.params.loc[CV.params['parameter']=='scan_rate', 'value'].item()) + ' V/s', zorder=0)
# plt.xlim(-.75, -.74)
# plt.ylim(.00003, .00004)
# Maxima
find_ipa = find_peaks(smooth_iw, height=0, width=3600) # Must give minimum height to get peak_heights dict.
ipa, Epa = CV.data['iw'][find_ipa[0]].to_numpy(), CV.data['Ecell'][find_ipa[0]].to_numpy()
# Invert signal to find minima - Signals are negative but indicies are correct!
find_ipc = find_peaks(-smooth_iw, height=0, width=3600) # Remember to un-invert anytime if calling ipc values from find_ipc, but not indicies.
ipc, Epc = CV.data['iw'][find_ipc[0]].to_numpy(), CV.data['Ecell'][find_ipc[0]].to_numpy()
peaks = {'ipa': ipa[0], 'Epa': Epa[0], 'ipc': ipc[0], 'Epc': Epc[0]}
# df = pd.DataFrame(data=peaks, index=[CV.params.loc[CV.params['parameter']=='scan_rate', 'value'].item()]) # Gives a little table, but rounds values...
print(peaks)
# -
# #### Peak picking and figure saving
# + tags=[]
def save_CV_fig(fig, analyte, electrode, solvent, notebook):
ans = input('Save the figure?')
if ans in ['y', 'yes', 'Yes', 'yeye', 'yeah', 'why, certainly', True]:
fig.savefig("C:\\Users\\jgage\\Documents\GitHub\\CHEM274\\data\\figs\\CV_"+
analyte + electrode + solvent +
str(CV.params.loc[CV.params['parameter']=='scan_rate', 'value'].item()) + 'Vs-1_'+
notebook+'.jpg')
print('Saved')
else:
print('Not saved')
# + pycharm={"is_executing": true} tags=[]
# Draw same plot as above, but with peaks
fig = plt.figure()
ax = fig.add_subplot()
ax.tick_params(axis='both', which='both', direction='in', right=True, top=True)
ax.plot(CV.data['Ecell'], CV.data['iw'], label = '$v=$ '+str(CV.params.loc[CV.params['parameter']=='scan_rate', 'value'].item()) + ' V/s', zorder=0)
ax.plot(CV.data['Ecell'], smooth_iw, color='C2', label='Savitsky-Golay', lw=.5, zorder=.5)
ax.scatter([*Epa, *Epc], [*ipa, *ipc], color='C1', label='Peaks', marker='|', s=200, zorder=1)
fig.tight_layout()
# ax.set_title('Cyclic Voltammogram')
ax.set_xlabel('$E_{cell}$ / V')
ax.set_ylabel('$i_{w}$ / $A$')
ax.ticklabel_format(style='sci', axis='y', scilimits=(0,0))
fig.legend(loc=(0.75, 0.78))
analyte = 'FcStar_'
electrode = 'GC_'
solvent = 'MeAc'
notebook = '_P-09-03'
plt.show()
save_CV_fig(fig, analyte, electrode, solvent, notebook)
# -
# #####
|
.ipynb_checkpoints/CV Week 4 Offsetting and iR correction-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:dl]
# language: python
# name: conda-env-dl-py
# ---
# <div class="alert alert-block alert-info">
# <font size="5"><b><center> Section 5</font></center>
# <br>
# <font size="5"><b><center>Recurrent Neural Network in PyTorch with an Introduction to Natural Language Processing</font></center>
# </div>
# # Simple Text Processing
# ## Typically Data Preprocessing Steps before Modeling Training for NLP Applications
#
# * Read the data from disk
# * Tokenize the text
# * Create a mapping from word to a unique integer
# * Convert the text into lists of integers
# * Load the data in whatever format your deep learning framework requires
# * Pad the text so that all the sequences are the same length, so you can process them in batch
thor_review = "the action scenes were top notch in this movie. Thor has never been this epic in the MCU. He does some pretty epic sh*t in this movie and he is definitely not under-powered anymore. Thor in unleashed in this, I love that."
word_embedding = "word embeddings are a representation of the *semantics* of a word, efficiently encoding semantic information that might be relevant to the task at hand"
print(list(word_embedding))
# ## Converting text into words
print(word_embedding.split())
# ## N-grams and NLTK
from nltk import ngrams
print(list(ngrams(word_embedding.split(),2)))
# - `n-grams` is used in many supervised machine learning models, such as Naive Bayes, to improve feature space.
#
# - `n-grams` are also used for spelling correction and text-summarization tasks.
#
# - Limitation of `n-grams` representation: it loses the sequential nature of text
# **Do one-hot encoding using canned functions**
# A major limitation of the one-hot representation is that the data is too sparse, because the size of the vector quickly grows as the number of unique words in the vocabulary increases. As such, this representation is not typically used in deep learning.
# ## Word Embedding
# Word embedding is a very popular way of representing text data in problems that are solved by deep learning algorithms
#
# Word embedding provides a dense representation of a word filled with floating numbers.
#
# It drastically reduces the dimension of the dictionary
#
#
# + [markdown] heading_collapsed=true
# ### `Torchtext` and Training word embedding by building a sentiment classifier
# + [markdown] hidden=true
# Torchtext takes a declarative approach to loading its data:
#
# * you tell torchtext how you want the data to look like, and torchtext handles it for you
#
# * Declaring a Field: The Field specifies how you want a certain field to be processed
#
# The `Field` class is a fundamental component of torchtext and is what makes preprocessing very easy
#
#
# -
# ### Load Car Review into `torchtext.data`
# +
# pip install torchtext
from torchtext import data, datasets
from torchtext.vocab import GloVe,FastText,CharNGram
import pandas as pd
import numpy as np
import re
import string
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from nltk.tokenize import word_tokenize
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.autograd import Variable
import torch
# -
# * Define two `Field` objects for the actual texts and the labels.
#
# * For the actual texts, `torchtext` will lowercase, tokenize, and trim the text to a maximum length of 20. We could change the maximum length of the text to other numbers.
#Load Subaru data
review_df = pd.read_csv("data/Section5/car_review.csv")
review_df.info
review_df.head()
review_df.shape
#Word count describe stats for review text
review_df.review_text.apply(lambda x: len(x)).describe()
# +
#Split DF csv to training and testing data
def prepare_csv(df_file, text_clm, label_clm, test_ratio, seed):
"""
Split review_df to training and testing sets as csv
Arguments:
df_file, the full dataset csv about to be split
text_clm, the column name of the text in csv
label_clm, the column name of the label in csv
val_ration, validation set ratio to the overall data
seed, randmoization seed
Return:
save train and test csv files under data/Section5/cache folder
"""
#Read file
full_df = pd.read_csv(df_file)
#shuffle index
idx = np.arange(full_df.shape[0])
np.random.seed(seed)
np.random.shuffle(idx)
#test set size
test_size = int(len(idx) * test_ratio)
#Save train and test csv under cache folder
full_df.iloc[idx[test_size:], :].to_csv("data/Section5/cache/training.csv", index = False)
full_df.iloc[idx[:test_size], :].to_csv("data/Section5/cache/testing.csv", index = False)
#Print size of trainig and testing data
print("training set size: " + str(len(idx[test_size:])))
print("testing set size: " + str(len(idx[:test_size])))
# -
#Define tokenizer function for the Text data field
def tokenizer(comment):
"""
Tokenize reviews without punctuation and non-text signs
Arguments:
String text
Returns:
Tokenized clean words
"""
comment = re.sub(r"[\*\"“”\n\\…\+\-\/\=\(\)‘•:\[\]\|’\!;]", " ", str(comment))
comment = re.sub(r"[ ]+", " ", comment)
comment = re.sub(r"\!+", "!", comment)
comment = re.sub(r"\,+", ",", comment)
comment = re.sub(r"\?+", "?", comment)
return [x for x in word_tokenize(comment) if x not in string.punctuation]
# ## Build Train and Text `torch.data.TabularDataset`
# +
#Split Train and Test
prepare_csv("data/Section5/car_review.csv", "review_text", "label", 0.3, 42)
#construct field
TEXT = data.Field(sequential=True, fix_length=300, tokenize=tokenizer, pad_first=True,
#tensor_type=torch.cuda.LongTensor,
lower=True, batch_first = True)
LABEL = data.Field(sequential = False)
#Build training and testing datasets
train = data.TabularDataset(path='data/Section5/cache/training.csv', format='csv', skip_header=True,
fields=[('review_text', TEXT),
('label', LABEL),
])
test = data.TabularDataset(path='data/Section5/cache/testing.csv', format='csv', skip_header=True,
fields=[('review_text', TEXT),
('label', LABEL),
])
# -
# - **Read in car_review.csv and split it ito train and test datasets**
#
# - The following command abstracts away all the complexity involved in downloading, tokenizing, and splitting the database into train and test datasets
#
# - Torch comes with datasets to play with; e.g. IMDB datasets. `torch.datasets` come with many datasets, such as `IMDB` for sentiment analysis, `TREC` for question classification, `WikiText-2` for language modeling, etc.
#
# - For tokenization, `NLTK` can also be specified
#Number of training and testing reviews
len(train), len(test)
train.fields['review_text']
# `train.fields` contains a dictionary where `TEXT` is the key and the `value` LABEL
type(train.fields)
print('train.fields', train.fields)
train.fields.keys()
len(train.fields)
train.fields.values()
train.fields.items()
type(train)
len(train[0].review_text),
vars(train[120])
# ## Buidling a vocabulary
# The `torchtext` facilitate the creation of a dictionary from a corpus of text. Once a dataset is loaded, we can call `build_vocab` and pass the necessary arguments that will handle building the vocabulary for the data.
#
# For instance, in the code below, we can initialize vectors with `pretrained embeddings of dimensions 300`
#Build Vocabulary with Glove
TEXT.build_vocab(train, vectors=GloVe(name='6B', dim=300),max_size=10000,min_freq=5)
LABEL.build_vocab(train,)
# Once the vocabulary is built, we can obtain different values such as
# * frequency
# * word index
# * vector representation for each word
print(TEXT.vocab.vectors)
TEXT.vocab.vectors.shape
TEXT.vocab.vectors[3].shape
TEXT.vocab.vectors[234]
print(TEXT.vocab.stoi)
# ### Batching
# `Torchtext` provides `BucketIterator`, which helps in batching all the text and replacing the words with the index number of the words.
#
# * The `BucketIterator` instance comes with a lot of useful parameters like `batch_size`, `device` (GPU or CPU), and `shuffle`.
train_iter, test_iter = data.BucketIterator.splits((train, test), batch_size=128, device=-1,shuffle=True)
# **Create a batch**
# +
batch = next(iter(train_iter))
print(batch.review_text.shape)
print(batch.label.shape)
# batch_size x fix_len
batch.review_text
batch.label
# -
# ## Create a network with Word Embedding
# # Use LSTM for Sentiment Classification
# 1. Preparing the data (previous steps)
# 2. Creating the batches
# 3. Creating the network
# 4. Training the model
# +
#Split Train and Test
prepare_csv("data/Section5/car_review.csv", "review_text", "label", 0.3, 42)
#construct field
TEXT = data.Field(sequential=True, fix_length=200, tokenize=tokenizer,# pad_first=True,
lower=True, batch_first = False)
LABEL = data.Field(sequential = False,)
#Build training and testing datasets
train = data.TabularDataset(path='data/Section5/cache/training.csv', format='csv', skip_header=True,
fields=[('review_text', TEXT),
('label', LABEL),
])
test = data.TabularDataset(path='data/Section5/cache/testing.csv', format='csv', skip_header=True,
fields=[('review_text', TEXT),
('label', LABEL),
])
#Build dictionary
TEXT.build_vocab(train, vectors=GloVe(name='6B', dim=300),max_size=10000, min_freq=5)
LABEL.build_vocab(train,)
# -
#Break train, test data into batches(size 32)
train_iter, test_iter = data.BucketIterator.splits((train, test), batch_size=32, device=-1, sort_key = lambda x: len(x.review_text), sort_within_batch = False)
train_iter.repeat = False
test_iter.repeat = False
class IMDBRnn(nn.Module):
def __init__(self,vocab,hidden_size,n_cat,bs=1,nl=2):
super().__init__()
self.hidden_size = hidden_size
self.bs = bs
self.nl = nl
self.e = nn.Embedding(n_vocab,hidden_size)
self.rnn = nn.LSTM(hidden_size,hidden_size,nl)
self.fc2 = nn.Linear(hidden_size,n_cat)
self.softmax = nn.LogSoftmax(dim=-1)
def forward(self,inp):
bs = inp.size()[1]
if bs != self.bs:
self.bs = bs
e_out = self.e(inp)
h0 = c0 = Variable(e_out.data.new(*(self.nl,self.bs,self.hidden_size)).zero_())
rnn_o,_ = self.rnn(e_out,(h0,c0))
rnn_o = rnn_o[-1]
fc = F.dropout(self.fc2(rnn_o),p=0.50)
return self.softmax(fc)
n_vocab = len(TEXT.vocab)
n_hidden = 64
# + code_folding=[]
model = IMDBRnn(n_vocab,n_hidden,3,bs=32)
#model = model.cuda()
optimizer = optim.Adam(model.parameters(),lr=1e-3)
def fit(epoch,model,data_loader,phase='training',volatile=False):
if phase == 'training':
model.train()
if phase == 'validation':
model.eval()
volatile=True
running_loss = 0.0
running_correct = 0
for batch_idx , batch in enumerate(data_loader):
text,target = batch.review_text , batch.label
# if is_cuda:
# text,target = text.cuda(),target.cuda()
if phase == 'training':
optimizer.zero_grad()
output = model(text)
loss = F.nll_loss(output, target)
running_loss += F.nll_loss(output,target,size_average=False).data
preds = output.data.max(dim=1,keepdim=True)[1]
running_correct += preds.eq(target.data.view_as(preds)).cpu().sum()
if phase == 'training':
loss.backward()
optimizer.step()
loss = running_loss/len(data_loader.dataset)
accuracy = 100. * running_correct/len(data_loader.dataset)
print("loss: ", loss, "accuracy: ", accuracy)
#print(f'{phase} loss is {loss:{5}.{2}} and {phase} accuracy is {running_correct}/{len(data_loader.dataset)}{accuracy:{10}.{4}}')
return loss,accuracy
# +
import time
start = time.time()
train_losses , train_accuracy = [],[]
val_losses , val_accuracy = [],[]
for epoch in range(1,25):
print("epoch:" + str(epoch) )
epoch_loss, epoch_accuracy = fit(epoch,model,train_iter,phase='training')
val_epoch_loss, val_epoch_accuracy = fit(epoch,model,test_iter,phase='validation')
train_losses.append(epoch_loss)
train_accuracy.append(epoch_accuracy)
val_losses.append(val_epoch_loss)
val_accuracy.append(val_epoch_accuracy)
end = time.time()
print((end-start)/60)
print("Execution Time: ", round(((end-start)/60),1), "minutes")
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(range(1,len(train_losses)+1),train_losses,'bo',label = 'training loss')
plt.plot(range(1,len(val_losses)+1),val_losses,'r',label = 'validation loss')
plt.legend()
# -
plt.plot(range(1,len(train_accuracy)+1),train_accuracy,'bo',label = 'train accuracy')
plt.plot(range(1,len(val_accuracy)+1),val_accuracy,'r',label = 'val accuracy')
plt.legend()
|
Notebook/Section 5 - RNN and NLP - Car Example v3b.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pytorch
# language: python
# name: pytorch
# ---
# +
# imports
import torch
# +
# GPU CHECK
# cuda version
print("cuda version: " , torch.version.cuda)
# check fot the GPU
print("GPU available: ",torch.cuda.is_available())
# check device name
print("Device Name : ", torch.cuda.get_device_name)
# how many GPU's do we have
print("The number of GPU's we have are : ", torch.cuda.device_count() )
# switiching to GPU
if torch.cuda.is_available():
device = torch.device("cuda:0")
print("Running on the GPU")
else:
device = torch.device("cpu")
print("Running on the CPU")
|
Code/project code from discovery cluster/.ipynb_checkpoints/GPU -checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Find Errors
# *Learned Skills: basic python skills*
# Note: to use the value of a variable as part of a string, you can write `%X` in the string as a placeholder for the variable, and append `% variable` to the string to specify which variable should be used. `%X` specifies the type of the variable, e.g. use `%d` for integers and `%g` for floats. E.g.:
#
# `s = "integer %d, float %g" % (1, 3.14)`
#
# sets `s` to `"integer 1, float 3.14"`
# Find 3 programming errors in the following code:
# ```
# import random
#
# def compute(n):
# i = 0; s = 0
# while i <= n:
# s += random.random()
# i += 1
# return s/n
#
# n = raw_input("Enter a number: ")
#
# print('the average of %d random numbers is %g' % (n, compute(n)))
# ```
# # Number game
# *Learned Skills: Random numbers; Loops*
# Implement the following guessing game: Someone writes down a secret number between 00–99. The other players then take turns guessing numbers, with a catch: if someone says the secret number, that person loses. If the guessed number is not the secret one, it splits the range.
# The chooser then states the part which contains the chosen number. If the new region
# only has one number, the chooser loses. Here's an example:
#
# * Chooser writes down (secretly) his number (let’s say, 30).
# * Chooser: “State a number between 00 and 99.”
# * Player: “42”.
# * Chooser: “State a number between 00 and 42.”
# * Player: “26”.
# * Chooser: “State a number between 26 and 42.”
#
# $\vdots$
#
# * Chooser: “State a number between 29 and 32.”
# * Player: “31”.
# * Chooser loses.
#
# Implement this game in Python, where the computer is the chooser.
#
# **Useful:** $\mathtt{random.randint()}$ and $\mathtt{input()}$.
# # Rock-paper-scissors
# *Learned Skills: Random choices*
# - Implement the game of rock-paper-scissors.
# - Extra: make it rock-paper-scissors-lizard-spock (if you do not know it, Wikipedia should help).
#
# **Useful:** $\mathtt{random.choice()}$
# # Dice Simulation
# *Learned Skills: Random numbers; Monte-Carlo Simulation*
# Estimate the chance of an event in a dice game. What is the probability of getting
# at least one 6 when throwing two dice? While this question can be analyzed theoretically, here we want to simulate it.
# 1\. Create a script that $n$ times draws two uniform random integers
# between 1 and 6 and counts how many times $p$ a 6 shows up. Write out the
# estimated probability $p/n$ together with the exact result 11/36. Run the script a
# few times with different n values and determine from the experiments how large
# n must be to get at least three decimals (0.306) of the probability correct. Use
# the random module to draw random uniformly distributed integers in a specified
# interval.
# 2\. Generalize the script to an arbitrary number of dice, $N$.
# 3\. Determine if you win or lose a hazard game. Somebody suggests the following
# game. You pay 1 unit of money and are allowed to throw four dice. If the sum of
# the eyes on the dice is less than 9, you win 10 units of money, otherwise you lose
# your investment. Should you play this game?
# # Calculating a Histogram
# *Learned Skills: Creating a function; List handling; Random numbers*
# 1\. Write a function $\mathtt{histgram(data, numbins)}$ which calculates the histogram of a
# given data set, where ```numbins``` gives the number of intervals in which the data
# range is divided.
#
# The function should return a tuple of two lists of equal lengths. The first list
# contains the midpoints of the intervals and the second list contains the counts of
# data points in the interval.
# 2\. Give a pseudo-graphical representation of the distribution, by drawing a number of
# stars corresponding to the number of data elements in a given interval. Example:
#
# 0.0
# 0.5
# 1.0
# 1.5
# 2.0
# 2.5
# 3.0
# 3.5
# 4.0
#
# +++
#
# +++++
#
# ++++++++
#
# ++++++++++++++++++
#
# +++++++++++++
#
# ++++++++++
#
# ++++++++
#
# ++++++
#
# ++
# 3\. Test the function by drawing samples from different probability distributions from the package $\mathtt{random}$.
# # Word counting
# *Learned Skills: File input/output*
# 1\. Create a script that opens a text file for reading and report the number of lines,
# words and characters. Assume that words are separated by whitespace.
# 2\. Open a text file for writing and save the count in it.
# 3\. Extra: Choose an arbitrary lengthy text (in raw text format) and list the ten most frequently used words. You can use a Dictionary for counting.
# **Useful:** $\mathtt{string.split()}$ $\mathtt{open()}$, $\mathtt{file.read()}$ and $\mathtt{file.write()}$
|
module0/1_python_basics/python_basics_2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Example 7 - Hydrodinamic Bearings
# =====
# In this example, we use the rotor seen in Example 5.9.6 from 'Dynamics of Rotating Machinery' by <NAME>, <NAME>, <NAME> & <NAME>, published by Cambridge University Press, 2010.
#
# Same rotor of Example 3, but the bearings are replaced with hydrodynamic bearings. In order to instantiate them, rather than giving the stiffness and damping data, we will calculate them using their hydrodinamic data, as provided by Example 5.5.1 from the book: The oil-film bearings have a diameter of 100 mm, are 30 mm long, and each supports a static load of 525 N, which represents half of the weight of the rotor. The radial clearance in the bearings is 0.1 mm and the oil film has a viscosity of 0.1 Pa s.
#
from bokeh.io import output_notebook, show
import ross as rs
import numpy as np
output_notebook()
# +
# Classic Instantiation of the rotor
shaft_elements = []
bearing_seal_elements = []
disk_elements = []
Steel = rs.materials. steel
for i in range(6):
shaft_elements.append(rs.ShaftElement(L=0.25, material=Steel, n=i, i_d=0, o_d=0.05))
disk_elements.append(rs.DiskElement.from_geometry(n=2,
material=Steel,
width=0.07,
i_d=0.05,
o_d=0.28
)
)
disk_elements.append(rs.DiskElement.from_geometry(n=4,
material=Steel,
width=0.07,
i_d=0.05,
o_d=0.35
)
)
bearing_seal_elements.append(rs.BearingElement.from_fluid_flow(n=0, nz=30, ntheta=20, nradius=11,
length=0.03, omega=157.1, p_in=0,
p_out=0, radius_rotor=0.0499,
radius_stator=0.05, visc=0.1,
rho=860., load=525))
bearing_seal_elements.append(rs.BearingElement.from_fluid_flow(n=6, nz=30, ntheta=20, nradius=11,
length=0.03, omega=157.1, p_in=0,
p_out=0, radius_rotor=0.0499,
radius_stator=0.05, visc=0.1,
rho=860., load=525))
rotor596c = rs.Rotor(shaft_elements=shaft_elements,
bearing_seal_elements=bearing_seal_elements,
disk_elements=disk_elements, n_eigen=12)
show(rotor596c.plot_rotor())
# +
# From_section class method instantiation
bearing_seal_elements = []
disk_elements = []
shaft_length_data = 3*[0.5]
i_d = 3*[0]
o_d = 3*[0.05]
disk_elements.append(rs.DiskElement.from_geometry(n=1,
material=Steel,
width=0.07,
i_d=0.05,
o_d=0.28
)
)
disk_elements.append(rs.DiskElement.from_geometry(n=2,
material=Steel,
width=0.07,
i_d=0.05,
o_d=0.35
)
)
bearing_seal_elements.append(rs.BearingElement(n=0, kxx=1e6, kyy=1e6, cxx=3e3, cyy=3e3))
bearing_seal_elements.append(rs.BearingElement(n=3, kxx=1e6, kyy=1e6, cxx=3e3, cyy=3e3))
rotor596fs = rs.Rotor.from_section(brg_seal_data=bearing_seal_elements,
disk_data=disk_elements, leng_data=shaft_length_data,
i_ds_data=i_d,o_ds_data=o_d
)
show(rotor596fs.plot_rotor())
# +
# Obtaining results for w = 200 rpm
rotor596fs.w = 200*np.pi/30
print('Normal Instantiation =', rotor596c.wn/(2*np.pi), '[Hz]')
print('\n')
print('From Section Instantiation =', rotor596fs.wn/(2*np.pi), '[Hz]')
# +
# Obtaining results for w=4000RPM
rotor596c.w = 4000*np.pi/30
print('Normal Instantiation =', rotor596c.wn/(2*np.pi))
# -
show(rotor596c.run_campbell(np.linspace(0, 4000*np.pi/30, 50)).plot())
|
docs/examples/example_05_09_06.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Principal Component Analysis
# #### Introduction
# To extract prinicipal components Hebbian learning rule was modified by <NAME> and developed Oja's rule which is used to extract first prinicipal component (i.e. eigen vector corresponding to largest eigen value). Generalized Hebbian Algorithm (GHA) can be used to all prinicipal components. The Generalized Hebbian Algorithm (GHA), is also known in the literature as Sanger's rule, is a linear feedforward neural network model for unsupervised learning. GHA combines Oja's rule with the Gram-Schmidt process to produce a learning rule. Please refer to following links: https://en.wikipedia.org/wiki/Oja%27s_rule, https://en.wikipedia.org/wiki/Generalized_Hebbian_Algorithm. Many more links on this topic can be found in google.
# Import Required Libraries
import numpy
from matplotlib import pyplot as plt
from copy import deepcopy
# #### GHA
# Two kind of datasets are here one of the shape "X" and another which is just a diagonal line. Here we develop a single layer feedforward neural network with linear activation function and no bias. For this example since our dataset is two dimensional and we want to extract all the PCs we will have two outputs in the network.
#
# Note: result for the former dataset can reach a local solution ($[0.707, 0.707]$ and $[-0.707, 0.707]$) instead of global sol ($[1, 0]$ and $[0, 1]$).
# +
# PCA using GHA Extracting All PCs
# Parameters
# X = numpy.array([[1, 1], [1, -1], [-1, 1], [-1, -1], [2, 2], [2, -2], [-2, 2], [-2, -2]])
X = numpy.array([[-1, -1], [0, 0], [1, 1], [-2, -2], [2, 2], [-1.1, -0.8], [-2.1, -1.8], [1.1, 0.8], [2.1, 1.8]])
wig = numpy.random.normal(0, 0.5, (2, 2))
wig_Norm = wig/numpy.linalg.norm(wig, axis=1).reshape(2, 1)
eta = 0.2
epoch = 1
max_epoch = 200000
# Required Functions
def update_weights(lr, x, W, iterations):
y = numpy.dot(W, x)
LT = numpy.tril(numpy.matmul(y[:, numpy.newaxis], y[numpy.newaxis, :]))
W = W + lr/iterations * ((y[:, numpy.newaxis] * x) - (numpy.matmul(LT, W)))
return W
# Main
W_new = deepcopy(wig_Norm)
while epoch <= max_epoch:
for i in range(0, 8):
W_new = update_weights(eta, X[i], W_new, epoch)
# print('Epoch: ', epoch, ' LR: ', eta)
epoch += 1
print('Optimal Weights Reached!!!')
# -
# #### Singular Value Decompostion
# Analytically solution can be found for PCA using SVD.
# SVD
Mean = numpy.mean(X, axis=0)
X_Norm = X - Mean
row, col = X.shape
Sample_Cov_Matrix = numpy.matmul(X_Norm.T, X_Norm)/(row - 1)
Eigen_Values, Eigen_Vectors = numpy.linalg.eig(Sample_Cov_Matrix)
# #### Comparison
# Solution using SVD and GHA can be compared if they are close or not. GHA gives all the principal components and we can take dot product between them to see if they are orthogonal or not. To see if SVD and GHA solution are close we can take cross product between PC's found using GHA and SVD. If they are close then cross product should yield answer as zero.
# Test
print('Initial Guess:')
print(wig)
print(wig_Norm)
print('Final Sol:')
print(W_new)
print(W_new/numpy.linalg.norm(W_new, axis=1).reshape(2, 1))
print('Check if GHA 2 PCs are orthogonal: ')
print(numpy.dot(W_new[0], W_new[1]))
print('SVD and GHA Sol:')
print(Eigen_Vectors)
print(W_new.T)
print('Check if SVD and GHA sol are close:')
print(numpy.cross(W_new.T[:, 0], Eigen_Vectors[:, 0]))
print(numpy.cross(W_new.T[:, 1], Eigen_Vectors[:, 1]))
# #### Plot
# Plot the dataset and PCs found using GHA and SVD.
# Plot Results
plt.plot(X[:, 0], X[:, 1], '.')
plt.plot([0, W_new[0, 0]], [0, W_new[0, 1]], 'r')
plt.plot([0, W_new[1, 0]], [0, W_new[1, 1]], 'g')
plt.plot([0, Eigen_Vectors[0, 0]], [0, Eigen_Vectors[1, 0]], 'k')
plt.plot([0, Eigen_Vectors[0, 1]], [0, Eigen_Vectors[1, 1]], 'k')
plt.grid()
plt.show()
|
PCA_HebbianLearning_SimpleExample.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # How to visualize your schema with biothings_schema package
# import python package
from biothings_schema import Schema
# load schema
schema_url = 'https://raw.githubusercontent.com/data2health/schemas/biothings/biothings/biothings_curie_kevin.jsonld'
se = Schema(schema_url)
# ## Visualize the full schema as a tree
## Visualize the full tree
# !pip install graphviz
se.full_schema_graph()
# ## Visualize only the parents of a specific class
# visualize the parents of "GenomicEntity" class as a tree
se.sub_schema_graph(source='GenomicEntity', include_children=False, include_parents=True)
# ## Visualize only the children of a specific class
# visualize the children of "GenomicEntity" class as a tree
se.sub_schema_graph(source='GenomicEntity', include_children=True, include_parents=False, size="9,6")
# ## Visualize all classes which are parents/children of a specific class
# visualize the children/parents of "GenomicEntity" class
se.sub_schema_graph(source='GenomicEntity', size="9,6")
se.sub_schema_graph(source="bts:Gene", size="9,6")
se.sub_schema_graph(source="http://schema.biothings.io/GenomicEntity", size="9,6")
|
jupyter notebooks/Visualizing Schema.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from alphamind.api import *
from PyFin.api import *
from PyFin.Math.Accumulators.StatefulAccumulators import MovingAverage
from PyFin.Math.Accumulators.StatefulAccumulators import MovingSharp
from PyFin.Math.Accumulators.StatefulAccumulators import MovingMaxDrawdown
plt.style.use('ggplot')
# +
"""
Back test parameter settings
"""
start_date = '2012-01-01'
end_date = '2018-01-02'
benchmark_code = 300
universe_name = ['zz500', 'hs300']
universe = Universe(universe_name, universe_name)
frequency = '5b'
batch = 16
method = 'risk_neutral'
use_rank = 100
industry_lower = 1.
industry_upper = 1.
neutralize_risk = ['SIZE', 'LEVERAGE'] + industry_styles
constraint_risk = ['SIZE', 'LEVERAGE'] + industry_styles
size_risk_lower = 0
size_risk_upper = 0
turn_over_target_base = 0.25
benchmark_total_lower = 0.8
benchmark_total_upper = 1.
horizon = map_freq(frequency)
hedging_ratio = 0.
executor = NaiveExecutor()
# +
"""
Model phase: we need 1 constant linear model and one alpha model
"""
const_features_keys = ['roe_q_raw',
'ep_q_raw',
'DivP_raw',
'cfinc1_q_raw',
'EBIT_raw',
'EARNYILD_raw',
'EPIBS_raw']
const_weights = [0.2,
0.2,
0.1,
0.15,
0.10,
0.10,
0.10]
const_weights = pd.Series(const_weights,
index=const_features_keys)
const_model = ConstLinearModel(features=const_features_keys,
weights=const_weights)
linear_model_features = {
'ep_q_raw': LAST('ep_q'),
'roe_q_raw': LAST('roe_q'),
'DivP_raw': LAST('DivP'),
'cfinc1_q_raw': LAST('cfinc1_q'),
'EBIT_raw': LAST('EBIT'),
'EARNYILD_raw': LAST('EARNYILD'),
'EPIBS_raw': LAST('EPIBS')
}
total_features = {
'IVR_raw': LAST('IVR'),
'DivP_raw': LAST('DivP'),
'cfinc1_q_raw': LAST('cfinc1_q'),
'CFinc1_raw': LAST('CFinc1'),
'roe_q_raw': LAST('roe_q'),
'ep_q_raw': LAST('ep_q'),
'EBIT_raw': LAST('EBIT'),
'EARNYILD_raw': LAST('EARNYILD'),
'EPIBS_raw': LAST('EPIBS')
}
total_features_keys = list(total_features.keys())
# +
"""
Data phase
"""
engine = SqlEngine()
linear_model_factor_data = fetch_data_package(engine,
alpha_factors=linear_model_features,
start_date=start_date,
end_date=end_date,
frequency=frequency,
universe=universe,
benchmark=benchmark_code,
batch=batch,
neutralized_risk=neutralize_risk,
pre_process=[winsorize_normal, standardize],
post_process=[winsorize_normal, standardize],
warm_start=batch)
linear_model_features_keys = linear_model_factor_data['x_names']
train_x = linear_model_factor_data['train']['x']
train_y = linear_model_factor_data['train']['y']
train_risk = linear_model_factor_data['train']['risk']
ref_dates = sorted(train_x.keys())
predict_x = linear_model_factor_data['predict']['x']
predict_y = linear_model_factor_data['predict']['y']
predict_risk = linear_model_factor_data['predict']['risk']
predict_code = linear_model_factor_data['predict']['code']
settlement = linear_model_factor_data['settlement']
# -
def cross_product(x, y):
n, m = x.shape
res = []
for j in range(m):
res.append(x[:, [j]] * y)
return np.concatenate(res, axis=1)
# +
"""
Training phase
"""
models_series = pd.Series()
for ref_date in ref_dates:
x = train_x[ref_date]
y = train_y[ref_date]
risk = train_risk[ref_date][:, 1:]
new_x = cross_product(x, risk)
#model = LinearRegression(features=linear_model_features_keys, fit_intercept=True) # n_jobs=8, min_samples_split=20)
model = LassoRegression(alpha=0.01, features=linear_model_features_keys, fit_intercept=True) # n_jobs=8, min_samples_split=20)
model.fit(new_x, y)
models_series.loc[ref_date] = model
alpha_logger.info('trade_date: {0} training finished'.format(ref_date))
frequency = '5b'
ref_dates = makeSchedule(start_date, end_date, frequency, 'china.sse')
const_model_factor_data = engine.fetch_data_range(universe,
total_features,
dates=ref_dates,
benchmark=benchmark_code)['factor']
horizon = map_freq(frequency)
# +
"""
Predicting and re-balance phase
"""
weight_gaps = [0.01, 0.02, 0.03, 0.04]
model1_weight = 1.0
model2_weight = 0.5
factor_groups = const_model_factor_data.groupby('trade_date')
for weight_gap in weight_gaps:
print("start {0} weight gap simulation ...".format(weight_gap))
rets = []
turn_overs = []
leverages = []
previous_pos = pd.DataFrame()
index_dates = []
for i, value in enumerate(factor_groups):
date = value[0]
data = value[1]
ref_date = date.strftime('%Y-%m-%d')
total_data = data.fillna(data[total_features_keys].median())
alpha_logger.info('{0}: {1}'.format(date, len(total_data)))
risk_exp = total_data[neutralize_risk].values.astype(float)
industry = total_data.industry_code.values
benchmark_w = total_data.weight.values
constraint_exp = total_data[constraint_risk].values
risk_exp_expand = np.concatenate((constraint_exp, np.ones((len(risk_exp), 1))), axis=1).astype(float)
risk_names = constraint_risk + ['total']
risk_target = risk_exp_expand.T @ benchmark_w
lbound = np.maximum(0., benchmark_w - weight_gap) # np.zeros(len(total_data))
ubound = weight_gap + benchmark_w
is_in_benchmark = (benchmark_w > 0.).astype(float)
risk_exp_expand = np.concatenate((risk_exp_expand, is_in_benchmark.reshape((-1, 1))), axis=1).astype(float)
risk_names.append('benchmark_total')
constraint = Constraints(risk_exp_expand, risk_names)
for i, name in enumerate(risk_names):
if name == 'total':
constraint.set_constraints(name,
lower_bound=risk_target[i],
upper_bound=risk_target[i])
elif name == 'SIZE':
base_target = abs(risk_target[i])
constraint.set_constraints(name,
lower_bound=risk_target[i] + base_target * size_risk_lower,
upper_bound=risk_target[i] + base_target * size_risk_upper)
elif name == 'benchmark_total':
base_target = benchmark_w.sum()
constraint.set_constraints(name,
lower_bound=benchmark_total_lower * base_target,
upper_bound=benchmark_total_upper * base_target)
else:
constraint.set_constraints(name,
lower_bound=risk_target[i] * industry_lower,
upper_bound=risk_target[i] * industry_upper)
factor_values = factor_processing(total_data[const_features_keys].values,
pre_process=[winsorize_normal, standardize],
risk_factors=risk_exp,
post_process=[winsorize_normal, standardize])
# const linear model
er1 = const_model.predict(factor_values)
# linear regression model
try:
model = models_series.ix[date]
except KeyError:
alpha_logger.info("model is not available. Omit this date")
continue
index_dates.append(date)
# x = predict_x[date]
x = predict_x[date]
risk = predict_risk[date][:, 1:]
this_predict_code = predict_code[date]
new_x = cross_product(x, risk)
er2 = model.predict(new_x)
# combine model
er1_table = pd.DataFrame({'er1': er1, 'code': total_data.code.values})
er2_table = pd.DataFrame({'er2': er2, 'code': this_predict_code})
er_table = pd.merge(er1_table, er2_table, on=['code'], how='left').fillna(0)
er1 = er_table.er1.values
er2 = er_table.er2.values
if er2.std() > 0:
print('using backbone model and alpha model')
er = model1_weight * er1 / er1.std() + model2_weight * er2 / er2.std()
else:
print('using only backbone modedl')
er = model1_weight * er1
codes = total_data['code'].values
if previous_pos.empty:
current_position = None
turn_over_target = None
else:
previous_pos.set_index('code', inplace=True)
remained_pos = previous_pos.loc[codes]
remained_pos.fillna(0., inplace=True)
turn_over_target = turn_over_target_base
current_position = remained_pos.weight.values
try:
target_pos, _ = er_portfolio_analysis(er,
industry,
None,
constraint,
False,
benchmark_w,
method=method,
use_rank=use_rank,
turn_over_target=turn_over_target,
current_position=current_position,
lbound=lbound,
ubound=ubound)
except ValueError:
alpha_logger.info('{0} full re-balance'.format(date))
target_pos, _ = er_portfolio_analysis(er,
industry,
None,
constraint,
False,
benchmark_w,
method=method,
use_rank=use_rank,
lbound=lbound,
ubound=ubound)
target_pos['code'] = total_data['code'].values
turn_over, executed_pos = executor.execute(target_pos=target_pos)
executed_codes = executed_pos.code.tolist()
dx_returns = engine.fetch_dx_return(date, executed_codes, horizon=horizon, offset=1)
result = pd.merge(executed_pos, total_data[['code', 'weight']], on=['code'], how='inner')
result = pd.merge(result, dx_returns, on=['code'])
leverage = result.weight_x.abs().sum()
ret = result.weight_x.values @ (np.exp(result.dx.values) - 1.)
rets.append(np.log(1. + ret))
executor.set_current(executed_pos)
turn_overs.append(turn_over)
leverages.append(leverage)
previous_pos = executed_pos
alpha_logger.info('{0} is finished'.format(date))
ret_df = pd.DataFrame({'returns': rets, 'turn_over': turn_overs, 'leverage': leverages}, index=index_dates)
# index return
index_return = engine.fetch_dx_return_index_range(benchmark_code, start_date, end_date, horizon=horizon,
offset=1).set_index('trade_date')
ret_df['index'] = index_return['dx']
ret_df.loc[advanceDateByCalendar('china.sse', ref_dates[-1], frequency)] = 0.
ret_df = ret_df.shift(1)
ret_df.iloc[0] = 0.
ret_df['tc_cost'] = ret_df.turn_over * 0.002
ret_df['returns'] = ret_df['leverage'] * (ret_df['returns'] - ret_df['index'])
ret_df[['returns', 'tc_cost']].cumsum().plot(figsize=(12, 6),
title='Fixed frequency rebalanced: {0}'.format(frequency),
secondary_y='tc_cost')
ret_df['ret_after_tc'] = ret_df['returns'] - ret_df['tc_cost']
sharp_calc = MovingSharp(49)
drawdown_calc = MovingMaxDrawdown(49)
max_drawdown_calc = MovingMaxDrawdown(len(ret_df))
res_df = pd.DataFrame(columns=['daily_return', 'cum_ret', 'sharp', 'drawdown', 'max_drawn', 'leverage'])
total_returns = 0.
for i, ret in enumerate(ret_df['ret_after_tc']):
date = ret_df.index[i]
total_returns += ret
sharp_calc.push({'ret': ret, 'riskFree': 0.})
drawdown_calc.push({'ret': ret})
max_drawdown_calc.push({'ret': ret})
res_df.loc[date, 'daily_return'] = ret
res_df.loc[date, 'cum_ret'] = total_returns
res_df.loc[date, 'drawdown'] = drawdown_calc.result()[0]
res_df.loc[date, 'max_drawn'] = max_drawdown_calc.result()[0]
res_df.loc[date, 'leverage'] = ret_df.loc[date, 'leverage']
if i < 10:
res_df.loc[date, 'sharp'] = 0.
else:
res_df.loc[date, 'sharp'] = sharp_calc.result() * np.sqrt(49)
res_df.to_csv('hs300_{0}.csv'.format(int(weight_gap * 100)))
# -
# ## combine
# -------------
# +
from pandas import ExcelWriter
writer = ExcelWriter('d:/多因子500+300合并.xlsx')
df1 = pd.read_csv('d:/hs300.csv', index_col=0)
df2 = pd.read_csv('d:/zz500.csv', index_col=0)
# -
weights_secnario = {
'3-1': (3., 1.),
'2-1': (2., 1.),
'1-1': (1., 1.),
}
for key, weights in weights_secnario.items():
weight300, weight500 = weights
ret_series = (weight300* df1['ret_after_tc'] + weight500 * df2['ret_after_tc']) / (weight300 + weight500)
sharp_calc = MovingSharp(49)
drawdown_calc = MovingMaxDrawdown(49)
max_drawdown_calc = MovingMaxDrawdown(len(ret_df))
res_df = pd.DataFrame(columns=['daily_return', 'cum_ret', 'sharp', 'drawdown', 'max_drawn'])
total_returns = 0.
for i, ret in enumerate(ret_series):
date = ret_series.index[i]
sharp_calc.push({'ret': ret, 'riskFree': 0.})
drawdown_calc.push({'ret': ret})
max_drawdown_calc.push({'ret': ret})
total_returns += ret
res_df.loc[date, 'daily_return'] = ret
res_df.loc[date, 'cum_ret'] = total_returns
res_df.loc[date, 'drawdown'] = drawdown_calc.result()[0]
res_df.loc[date, 'max_drawn'] = max_drawdown_calc.result()[0]
if i < 10:
res_df.loc[date, 'sharp'] = 0.
else:
res_df.loc[date, 'sharp'] = sharp_calc.result() * np.sqrt(49)
res_df.to_excel(writer, key)
writer.save()
|
notebooks/candidate_prod_model_20171204.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Thinkful Data Science Fundamentals: Capstone Data Analysis Report
# -------------------------------------------------------------------------------------------------------------------------
# ## _United States Oil Pipeline Accidents, Jan 2010 - Dec 2016_
# ###### Source of data (gathered August 2018): [https://www.kaggle.com/usdot/pipeline-accidents](https://www.kaggle.com/usdot/pipeline-accidents)
# ###### Date: August 10, 2018
# -------------------------------------------------------------------------------------------------------------------------
# # _Import modules and enable the display of plots in this notebook_
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from scipy.stats import ttest_ind
# %matplotlib inline
# -
# -------------------------------------------------------------------------------------------------------------------------
# # _Load the dataset into a DataFrame_
file = 'database.csv'
accidents = pd.read_csv(file)
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: What is some _key information_ that can be learned quickly about the content of the dataset?
# How many (rows, columns) are in the dataset?
accidents.shape
# What are the column names and how many of them contain data?
accidents.info()
# -------------------------------------------------------------------------------------------------------------------------
# # _Prepare the dataset for analysis_
# +
# Make a copy of the initial DataFrame
acc_copy = accidents.copy()
# Add a 'Count' column to the copy
acc_copy['Count'] = 1
# Add a 'Month' column to the copy
acc_copy['Month'] = acc_copy['Accident Date/Time'].apply(lambda dt: int(dt[:dt.find('/')]))
# Add an 'Hour' column to the copy
def get_hour(dt):
hour = int(dt[dt.find(' ')+1:dt.find(':')])
if dt.strip().endswith('AM') and hour == 12:
hour = 0
elif dt.strip().endswith('PM') and hour != 12:
hour += 12
return hour
acc_copy['Hour'] = acc_copy['Accident Date/Time'].apply(get_hour)
# The dataset includes accidents from Jan 2010 through Jan 2017.
# Since data exists for full years 2010-2016 but only exists for
# a small portion of the year 2017 (two records for Jan 2017),
# I excluded those two records. This should prevent any incorrect
# conclusions from being made about 2017 relative to other years.
acc_2010_2016 = acc_copy[acc_copy['Accident Year'] < 2017]
# -
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: Which accidents caused the _most fatalities and/or injuries_?
injuries_fatalities = acc_2010_2016[(acc_2010_2016['All Injuries'] > 0) |
(acc_2010_2016['All Fatalities'] > 0)].fillna(0)
injuries_fatalities[['Operator ID', 'Operator Name', 'Pipeline/Facility Name',
'Pipeline Type', 'Liquid Type', 'Liquid Ignition',
'Liquid Explosion', 'Pipeline Shutdown',
'All Injuries', 'All Fatalities', 'All Costs']].sort_values(by=['All Fatalities', 'All Injuries'], ascending=[False, False])
# ### [Colonial Pipeline Co - Line 01: Accident history leading up to and including the October 2016 disaster](https://medium.com/planet-stories/the-anatomy-of-a-pipeline-accident-the-colonial-pipeline-spill-d30bb2a5941d)
# +
colonial_line_01 = acc_2010_2016[acc_2010_2016['Pipeline/Facility Name'] == 'LINE 01']
# Accident History/Details for Colonial Pipeline Co. - Line 01
colonial_line_01[['Accident Date/Time', 'Operator Name',
'Pipeline/Facility Name', 'Accident Latitude',
'Accident Longitude', 'Cause Category',
'Cause Subcategory', 'All Injuries',
'All Fatalities', 'All Costs']]
# -
# ###### A common motto in industry is "Safety First" and the accident above is [very tragic](https://www.cbs42.com/news/local/2nd-colonial-pipeline-explosion-victim-has-died-osha-says/868185307). There (fortunately) are not enough safety-related incidents (injuries/fatalities) in this dataset to warrant a detailed statistical exploration in those categories. However, we should look for warning signs in historical data that may indicate heightened risk parameters, which might help prevent future loss of life, injuries, and environmental damage.
#
# ###### Numerical and statistical analysis will focus on some of the other columns in the dataset.
# -------------------------------------------------------------------------------------------------------------------------
# # _Limitations of the analyzed data_
# ### The dataset used in this analysis only includes information regarding "pipeline accidents" for the United States, from 2010-2016. This means that several noteworthy oil-related accidents, such as the Exxon Valdez "tanker" spill in 1989 and the Deepwater Horizon "offshore platform" accident from 2010, are not included in the dataset.
#
# ### Only seven years of U.S. pipeline accident data are included in this analysis. If the dataset was larger and included data for more than one decade, it might be necessary to adjust the values in the cost columns for inflation. No inflation adjustment of pipeline accident costs will be performed in the following numerical & statistical analyses.
# -------------------------------------------------------------------------------------------------------------------------
# # _Exploring the data for context & meaning_
# ### QUESTION: Which pipeline operators incurred the _greatest_ accident-related costs from 2010-2016, and _how many_ accidents did they experience over that timeframe?
# +
operator_acc_df = acc_2010_2016[['Operator ID', 'Operator Name', 'All Costs', 'Count']]
operator_acc_obj = operator_acc_df.groupby(['Operator ID', 'Operator Name'])
operator_acc = operator_acc_obj.sum().sort_values(by='All Costs', ascending=False)
# Top 15 Pipeline Operators with the Highest Incurred Costs for Pipeline Accidents
# from 2010-2016
# ('All Costs' includes totals from all associated Operator Pipeline/Facilites)
operator_acc.head(15)
# -
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: Did any Operator Pipelines/Facilities experience _more than one (1)_ accident from 2010-2016?
op_pipe_fac_acc_df = acc_2010_2016[['Operator ID', 'Operator Name',
'Pipeline/Facility Name', 'Pipeline Type',
'Liquid Type', 'Cause Category',
'Cause Subcategory', 'All Costs', 'Count']]
op_pipe_fac_acc_obj = op_pipe_fac_acc_df.groupby(['Operator ID', 'Operator Name',
'Pipeline/Facility Name', 'Pipeline Type'])
op_pipe_fac_acc = op_pipe_fac_acc_obj.sum()
op_pipe_fac_acc_mult = op_pipe_fac_acc[op_pipe_fac_acc['Count'] > 1]
# ###### Pipelines/Facilities with more than one (1) accident sorted by 'Count'
# Top 15 Pipeline/Facilities with the Most Accidents from 2010-2016
# The 'Count' column may reveal something about the age and/or
# quality control practices of the associated pipeline/facility.
op_pipe_fac_acc_mult.sort_values(by='Count', ascending=False).head(15)
# ###### Pipelines/Facilities with more than one (1) accident sorted by 'All Costs'
# Top 15 Pipeline/Facilities with the Most COSTLY Accidents from 2010-2016
# The 'Count' column may reveal something about the age and/or
# quality control practices of the associated operator pipeline/facility.
op_pipe_fac_acc_mult.sort_values(by='All Costs', ascending=False).head(15)
# ###### The above findings will be explored in more detail later in this analysis.
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: What is the _total number_ of pipeline accidents: by year, by month, & by hour?
# +
# Total accidents by year
# Have accident totals increased/decreased over the years?
acc_by_year = acc_2010_2016[['Accident Year', 'Count']].groupby('Accident Year').sum()
# Total accidents by month (over multiple years)
# Might season be a contributing factor to pipeline accidents?
acc_by_month = acc_2010_2016[['Month', 'Count']].groupby('Month').sum()
# Total accidents by the hour (over multiple years)
# Might time-of-day be a contributing factor to pipeline accidents?
acc_by_hour = acc_2010_2016[['Hour', 'Count']].groupby('Hour').sum()
# Plot the variables above
fig, axes = plt.subplots(1, 3, figsize=(18,4))
opacity = 0.5
# By Year
axes[0].bar(acc_by_year.index, acc_by_year['Count'], alpha=opacity, color='red')
axes[0].set_xlabel('Year')
axes[0].set_ylabel('Number of Accidents')
axes[0].set_title('Total Number of Pipeline Accidents\n(By Year)')
# By Month
axes[1].bar(acc_by_month.index, acc_by_month['Count'], alpha=opacity, color='orange')
axes[1].set_xlabel('Month')
axes[1].set_ylabel('Number of Accidents')
axes[1].set_title('Total Number of Pipeline Accidents\n(By Month, From Jan 2010 - Dec 2016)')
axes[1].set_xticks(acc_by_month.index)
axes[1].set_xticklabels(('Jan','Feb','Mar','Apr','May','Jun','Jul','Aug','Sept','Oct','Nov','Dec'))
# By Hour
axes[2].bar(acc_by_hour.index, acc_by_hour['Count'], alpha=opacity, color='blue')
axes[2].set_xlabel('Hour')
axes[2].set_ylabel('Number of Accidents')
axes[2].set_title('Total Number of Pipeline Accidents\n(By Hour - Starting From Midnight)')
axes[2].set_xticks(acc_by_hour.index)
axes[2].set_xticklabels(range(24))
plt.show()
# -
# ###### It is unknown as to whether the bell-shaped hourly chart (on the right) above may indicate:
# - ###### if pipeline accidents are more likely to happen during the daylight hours when more people are working, AND/OR
# - ###### if that is when people are more apt to notice that an accident has occurred and report it, assuming control systems have not logged the accidents automatically, AND/OR
# - ###### something else
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: What is the distribution of _total cost_ statistics for U.S. pipeline accidents, 2010-2016?
# Total cost statistics for all U.S. pipeline accidents, 2010-2016
all_pipeline_acc_2010_2016 = acc_2010_2016['All Costs']
all_pipeline_acc_2010_2016.describe()
# ###### The statistics above show that data (for 2,793 pipeline accidents) in the 'All Costs' column ranges from USD 0 to USD ~840,526,100. The average (mean) cost per accident (between 2010-2016) was USD 834,599 but has a standard deviation of USD ~16,584,220! Since 75% of the data points are less than USD 117,395 and 50% are less than USD 23,052 (the median), the average is weighted higher due to smaller numbers of much larger data points. This indicates that the total cost data for all pipeline accidents in the dataset is not normally distributed. This can be seen below where only pipeline accidents with total costs less than USD 1million are included in the histogram.
# +
plt.hist(acc_2010_2016[acc_2010_2016['All Costs'] < 1000000]['All Costs'], bins=20)
plt.xlabel('Cost of Pipeline Accident (USD)')
plt.ylabel('Number of Pipeline Accidents')
plt.title('Pipeline Accidents With Total Costs:\n USD 0 to 1,000,000 (ex)')
plt.show()
# -
# ###### However, there are many accidents in the dataset with total costs greater than USD 1million. Therefore, a better way to view the frequency distribution of this data may be to break it up into different cost tiers.
# ###### Create DataFrames for different cost tiers
# +
# Accidents where total costs were less than $10,000
cost_0_10k = acc_2010_2016[acc_2010_2016['All Costs'] < 10000]
# Accidents where total costs ranged from $10,000 to $100,000
cost_10k_100k = acc_2010_2016[(acc_2010_2016['All Costs'] >= 10000) &
(acc_2010_2016['All Costs'] < 100000)]
# Accidents where total costs ranged from $100,000 to $1,000,000
cost_100k_1m = acc_2010_2016[(acc_2010_2016['All Costs'] >= 100000) &
(acc_2010_2016['All Costs'] < 1000000)]
# Accidents where total costs ranged from $1,000,000 to $10,000,000
cost_1m_10m = acc_2010_2016[(acc_2010_2016['All Costs'] >= 1000000) &
(acc_2010_2016['All Costs'] < 10000000)]
# Accidents where total costs ranged from $10,000,000 to $100,000,000
cost_10m_100m = acc_2010_2016[(acc_2010_2016['All Costs'] >= 10000000) &
(acc_2010_2016['All Costs'] < 100000000)]
# Accidents where total costs ranged from $100,000,000 to $1,000,000,000
cost_100m_1b = acc_2010_2016[(acc_2010_2016['All Costs'] >= 100000000) &
(acc_2010_2016['All Costs'] < 1000000000)]
# -
# ###### Display frequency distributions for pipeline accidents where total costs fell within each of the above cost ranges
# +
# Plot the variables above
fig, axes = plt.subplots(2, 3, figsize=(18,12))
# [$0, $10,000)
axes[0,0].hist(cost_0_10k['All Costs'], bins=20)
axes[0,0].set_xlabel('Cost of Pipeline Accident (Thousand USD)')
axes[0,0].set_ylabel('Number of Pipeline Accidents')
axes[0,0].set_title('Pipeline Accidents With Total Costs:\n USD 0 to 10,000 (ex)')
# [$10,000, $100,000)
axes[0,1].hist(cost_10k_100k['All Costs'], bins=20)
axes[0,1].set_xlabel('Cost of Pipeline Accident (Ten Thousand USD)')
axes[0,1].set_ylabel('Number of Pipeline Accidents')
axes[0,1].set_title('Pipeline Accidents With Total Costs:\n USD 10,000 to 100,000 (ex)')
# [$100,000, 1,000,000)
axes[0,2].hist(cost_100k_1m['All Costs'], bins=20)
axes[0,2].set_xlabel('Cost of Pipeline Accident (Hundred Thousand USD)')
axes[0,2].set_ylabel('Number of Pipeline Accidents')
axes[0,2].set_title('Pipeline Accidents With Total Costs:\n USD 100k to 1m (ex)')
# [$1,000,000, $10,000,000)
axes[1,0].hist(cost_1m_10m['All Costs'], bins=20)
axes[1,0].set_xlabel('Cost of Pipeline Accident (Ten Million USD)')
axes[1,0].set_ylabel('Number of Pipeline Accidents')
axes[1,0].set_title('Pipeline Accidents With Total Costs:\n USD 1m to 10m (ex)')
# [$10,000,000, $100,000,000)
axes[1,1].hist(cost_10m_100m['All Costs'], bins=20)
axes[1,1].set_xlabel('Cost of Pipeline Accident (Hundred Million USD)')
axes[1,1].set_ylabel('Number of Pipeline Accidents')
axes[1,1].set_title('Pipeline Accidents With Total Costs:\n USD 10m to 100m (ex)')
# [$100,000,000, $1,000,000,000)
axes[1,2].hist(cost_100m_1b['All Costs'], bins=20)
axes[1,2].set_xlabel('Cost of Pipeline Accident (Hundred Million USD)')
axes[1,2].set_ylabel('Number of Pipeline Accidents')
axes[1,2].set_title('Pipeline Accidents With Total Costs:\n USD 100m to 1b (ex)')
plt.tight_layout()
plt.show()
# -
# ###### The frequency distribution of pipeline accident costs appears similarly shaped at each cost tier/range, with the exception of the plot on the bottom-right (as only three accidents totaling between USD 100m - 1b were recorded).
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: What are the _primary causes_ of pipeline accidents?
acc_2010_2016['Cause Category'].unique()
# +
causes = acc_2010_2016['Cause Category'].value_counts()
# Make a horizontal bar chart
fig, axes = plt.subplots(figsize=(14,6))
axes.barh(causes.index, causes.values)
axes.set_yticks(np.arange(len(causes.index)))
axes.set_yticklabels(causes.index)
axes.set_xlabel('Number of Accidents by Cause Category')
axes.set_title('Primary Causes of US Pipeline Accidents, 2010-2016\n')
plt.show()
# -
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: What are the most financially costly _types_ of pipeline accidents, grouped by: Pipeline Type, Liquid Type, & Cause Category?
# +
onshore_costs = acc_2010_2016[['Pipeline Type','Liquid Type',
'Cause Category','Count','All Costs']]
onshore_costs_obj = onshore_costs.groupby(['Pipeline Type',
'Liquid Type','Cause Category'])
grouped_costs = onshore_costs_obj.sum().sort_values(by='All Costs',
ascending=False)
grouped_costs['Avg Cost/Accident'] = grouped_costs['All Costs'] / grouped_costs['Count']
# Top 15 Most Financially Costly Types of Pipeline Accidents for 2010-2016
grouped_costs.head(15)
# -
# ###### When grouped by 'Pipeline Type,' 'Liquid Type,' and 'Cause Category,' accidents that are attributed to some variant of UNDERGROUND piping can be seen to comprise the Top 10 slots for most financially costly types of pipeline accidents for the timeframe 2010-2016.
#
# ###### 'MATERIAL/WELD/EQUIP FAILURE' and 'CORROSION' appear as the two (2) most costly causes of pipeline accidents.
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: How have the numbers of these two particular _causes_ of pipeline accidents changed over time?
# +
cause_by_year = acc_2010_2016[['Accident Year', 'Cause Category',
'Cause Subcategory', 'Count']]
# All Accidents Caused By Material / Weld / Equipment Failure
mwe_failure = cause_by_year[cause_by_year['Cause Category'] == 'MATERIAL/WELD/EQUIP FAILURE']
mwe_failure_by_year = mwe_failure.groupby('Accident Year').sum()
# All Accidents Caused By Corrosion
corr = cause_by_year[cause_by_year['Cause Category'] == 'CORROSION']
corr_by_year = corr.groupby('Accident Year').sum()
# Accidents Caused By External Corrosion
ext_corr = cause_by_year[(cause_by_year['Cause Category'] == 'CORROSION') &
(cause_by_year['Cause Subcategory'] == 'EXTERNAL')]
ext_corr_by_year = ext_corr.groupby('Accident Year').sum()
# Accidents Caused By Internal Corrosion
int_corr = cause_by_year[(cause_by_year['Cause Category'] == 'CORROSION') &
(cause_by_year['Cause Subcategory'] == 'INTERNAL')]
int_corr_by_year = int_corr.groupby('Accident Year').sum()
# Plot the variables above
fig, axes = plt.subplots(1, 2, figsize=(18,6))
opacity = 0.5
# Material / Weld / Equipment Failures By Year
axes[0].bar(mwe_failure_by_year.index, mwe_failure_by_year['Count'],
alpha=opacity, color='red')
axes[0].set_xlabel('Year')
axes[0].set_ylabel('Number of Accidents')
axes[0].set_title('Number of Pipeline Accidents Caused By:\n'
'Material, Weld, or Equipment Failure (By Year)')
# Corrosion By Year
axes[1].plot(corr_by_year.index, corr_by_year['Count'],
alpha=opacity, color='red', label='Total')
axes[1].plot(ext_corr_by_year.index, ext_corr_by_year['Count'],
alpha=opacity, color='brown', label='External Corrosion')
axes[1].plot(int_corr_by_year.index, int_corr_by_year['Count'],
alpha=opacity, color='orange', label='Internal Corrosion')
axes[1].set_xlabel('Year')
axes[1].set_ylabel('Number of Accidents')
axes[1].set_title('Number of Pipeline Accidents Caused By:\n'
'Corrosion (By Year)')
axes[1].legend()
plt.show()
# -
# ###### Although more annual data would be helpful, the 'Material, Weld, or Equipment Failure' plot on the left appears to trend upwards over time and is also similar in shape to the 'Total Number of Pipeline Accidents (By Year)' plot displayed previously. The 'Total' line-graph in the 'Corrosion' plot on the right also appears to have an upward trend, characterized by higher lows and higher highs for the given time period. It is likely attributed to some combination of aging infrastructure and/or better data collection over time.
# ###### Regarding corrosion, the National Association of Corrosion Engineers (NACE) states on its [website](http://impact.nace.org/economic-impact.aspx#):
#
# > ASSESSMENT OF THE GLOBAL COST OF CORROSION
#
# > The global cost of corrosion is estimated to be USD 2.5 trillion, which is equivalent to 3.4% of the global GDP (2013). By using available corrosion control practices, it is estimated that savings of between 15 and 35% of the cost of corrosion could be realized; i.e., between USD 375 and USD 875 billion annually on a global basis. These costs typically do not include individual safety or environmental consequences. Through near misses, incidents, forced shutdowns (outages), accidents, etc., several industries have come to realize that lack of corrosion management can be very costly and that, through proper corrosion management, significant cost savings can be achieved over the lifetime of an asset. To achieve the full extent of these savings, corrosion management and its integration into an organization’s management system must be accomplished by implementing a Corrosion Management System (CMS).
#
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: Did any operator pipelines/facilities experience _more than one (1)_ accident _due to the same causes_?
# +
pipe_fac_acc_causes_obj = op_pipe_fac_acc_df.groupby(['Operator ID', 'Operator Name',
'Pipeline/Facility Name',
'Pipeline Type', 'Liquid Type',
'Cause Category', 'Cause Subcategory'])
pipe_fac_acc_causes = pipe_fac_acc_causes_obj.sum()
pipe_fac_acc_same_causes = pipe_fac_acc_causes[pipe_fac_acc_causes['Count'] > 1]
# Make a scatter plot for Operator Pipelines/Facilities that have experienced
# the same types of accidents more than once and the total costs incurred
fig, axes = plt.subplots(1, 1, figsize=(10, 6))
opacity = 0.5
axes.scatter(pipe_fac_acc_same_causes['Count'], pipe_fac_acc_same_causes['All Costs'],
alpha=opacity, color='red')
axes.set_xlabel('Number of Same Types of Accidents')
axes.set_ylabel('Pipeline/Facility Total Costs Incurred (Ten Million USD)')
axes.set_title('Individual Operator Pipelines/Facilities Experiencing\nthe Same Types of Accidents More Than Once, From 2010-2016')
plt.show()
# -
# ###### See the DataFrame below for details regarding the five (5) most costly data points in the scatter plot above.
pipe_fac_acc_same_causes.sort_values(by='All Costs', ascending=False).head(5)
# -------------------------------------------------------------------------------------------------------------------------
# ### QUESTION: How _confidently_ can it be stated that, in general, 'UNDERGROUND' pipeline accidents are more costly than 'ABOVEGROUND' pipeline accidents?
# +
pop_underground_acc_costs = acc_2010_2016[acc_2010_2016['Pipeline Type'] == 'UNDERGROUND']['All Costs']
pop_aboveground_acc_costs = acc_2010_2016[acc_2010_2016['Pipeline Type'] == 'ABOVEGROUND']['All Costs']
t_stats = []
p_vals = []
for i in range(10000):
# I decided to use sample sizes equal to 1/2 of the number of accidents for each Pipeline Type
# Total underground accidents = 1474; total aboveground accidents = 984
sample_underground_costs = np.random.choice(pop_underground_acc_costs, 492, replace=False)
sample_aboveground_costs = np.random.choice(pop_aboveground_acc_costs, 737, replace=False)
ttest = ttest_ind(sample_underground_costs, sample_aboveground_costs, equal_var=False)
t_stats.append(ttest.statistic)
p_vals.append(ttest.pvalue)
# -
# After 10,000 iterations of sampling and averaging the p_values
avg_ttest_statistic = sum(t_stats) / len(t_stats)
avg_p_val = sum(p_vals) / len(p_vals)
print("Average t-statistic: ", avg_ttest_statistic)
print("Average p-value: ", avg_p_val)
# ###### Based on repeatedly sampling the dataset for the costs of accidents that happened 'UNDERGROUND' vs the costs of accidents that happened 'ABOVEGROUND,' comparing their sample statistics, and averaging the resulting t-statistic and p-value calculations, an average t-statistic of ~2.56 and an average p-value of ~0.05 are the results. The t-statistic indicates that the sample means are separated by ~2.56 standard errors. The p-value is close to 0.05, meaning that we can reject the hypothesis of both means being equal with 95% confidence.
# -------------------------------------------------------------------------------------------------------------------------
# ### _FURTHER RESEARCH - EVALUATING RISK FOR INDIVIDUAL PIPELINES/FACILITIES & OPERATORS_
# #### Based on all of the above findings, it would be interesting to explore the data further to determine:
# - ##### Do certain types of accidents occur more/less often depending on season?
# - ##### How concentrated (geographically and over time) are recurring types of accidents for the same pipelines/facilities?
#
# - ##### Which operators have _more than one_ pipeline/facility that have experienced the same types of accidents _more than once_?
# - #### This may reveal something about:
# - #### The age of operator assets, and/or
# - #### The rigor of QA/QC practices put in place during construction and/or maintenance
#
# #### It's possible that such research could also be applied to determine the likelihood (and locations at highest risk) of future accidents for those pipelines/facilities and others.
|
thinkful/data_science/my_progress/intro_data_science_fundamentals/unit_4_capstone/Analysis_of_Recent_Pipeline_Accidents_-_Final_Draft.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
#
#
# # Naive Bayes Classification
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In this assignement you will load a customer dataset, fit the data, and use Naive Bayes Classification to predict a data point.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Naive Bayes Classification** In statistics, Naive Bayes classifiers are a family of simple "probabilistic classifiers" based on applying Bayes' theorem with strong (naive) independence assumptions between the features. They are among the simplest Bayesian network models.But they could be coupled with Kernel density estimation and achieve higher accuracy levels.
#
# Naïve Bayes classifiers are highly scalable, requiring a number of parameters linear in the number of variables (features/predictors) in a learning problem. Maximum-likelihood training can be done by evaluating a closed-form expression,which takes linear time, rather than by expensive iterative approximation as used for many other types of classifiers.
#
# In the statistics and computer science literature, naive Bayes models are known under a variety of names, including simple Bayes and independence Bayes.All these names reference the use of Bayes' theorem in the classifier's decision rule, but naïve Bayes is not (necessarily) a Bayesian method.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Here's an explanation about Naive Bayes Classifier from wikipedia
#
# <img src="bayesClassifier.png">
#
# -
# <h1>Table of contents</h1>
#
# <div class="alert alert-block alert-info" style="margin-top: 20px">
# <ol>
# <li><a href="#about_dataset">About the dataset</a></li>
# <li><a href="#visualization_analysis">Data Visualization and Analysis</a></li>
# <li><a href="#classification">Classification</a></li>
# </ol>
# </div>
# <br>
# <hr>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets load required libraries
#
# + button=false new_sheet=false run_control={"read_only": false}
import itertools
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.ticker import NullFormatter
import pandas as pd
import numpy as np
import matplotlib.ticker as ticker
from sklearn import preprocessing
# %matplotlib inline
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <div id="about_dataset">
# <h2>About the dataset</h2>
# </div>
#
# -
# Imagine a telecommunications provider has segmented its customer base by service usage patterns, categorizing the customers into four groups. If demographic data can be used to predict group membership, the company can customize offers for individual prospective customers. It is a classification problem. That is, given the dataset, with predefined labels, we need to build a model to be used to predict class of a new or unknown case.
#
# The example focuses on using demographic data, such as region, age, and marital, to predict usage patterns.
#
# The target field, called **custcat**, has four possible values that correspond to the four customer groups, as follows:
# 1- Basic Service
# 2- E-Service
# 3- Plus Service
# 4- Total Service
#
# Our objective is to build a classifier, to predict the class of unknown cases. We will use a specific type of classification called K nearest neighbour.
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Data From CSV File
#
# + button=false new_sheet=false run_control={"read_only": false}
df = pd.read_csv('teleCust1000t.csv')
df.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <div id="visualization_analysis">
# <h2>Data Visualization and Analysis</h2>
# </div>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Let’s see how many of each class is in our data set
#
# + button=false new_sheet=false run_control={"read_only": false}
df['custcat'].value_counts()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### 281 Plus Service, 266 Basic-service, 236 Total Service, and 217 E-Service customers
#
# -
# You can easily explore your data using visualization techniques:
#
df.hist(column='income', bins=50)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Feature set
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Lets define feature sets, X:
#
# -
df.columns
# To use scikit-learn library, we have to convert the Pandas data frame to a Numpy array:
#
# + button=false new_sheet=false run_control={"read_only": false}
X = df[['region', 'tenure','age', 'marital', 'address', 'income', 'ed', 'employ','retire', 'gender', 'reside']] .values #.astype(float)
X[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# What are our labels?
#
# + button=false new_sheet=false run_control={"read_only": false}
y = df['custcat'].values
y[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Normalize Data
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Data Standardization give data zero mean and unit variance, it is good practice, especially for algorithms such as KNN which is based on distance of cases:
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# X = preprocessing.StandardScaler().fit(X).transform(X.astype(float))
# X[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Train Test Split
#
# Out of Sample Accuracy is the percentage of correct predictions that the model makes on data that that the model has NOT been trained on. Doing a train and test on the same dataset will most likely have low out-of-sample accuracy, due to the likelihood of being over-fit.
#
# It is important that our models have a high, out-of-sample accuracy, because the purpose of any model, of course, is to make correct predictions on unknown data. So how can we improve out-of-sample accuracy? One way is to use an evaluation approach called Train/Test Split.
# Train/Test Split involves splitting the dataset into training and testing sets respectively, which are mutually exclusive. After which, you train with the training set and test with the testing set.
#
# This will provide a more accurate evaluation on out-of-sample accuracy because the testing dataset is not part of the dataset that have been used to train the data. It is more realistic for real world problems.
#
# + button=false new_sheet=false run_control={"read_only": false}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.3, random_state=4)
print ('Train set:', X_train.shape, y_train.shape)
print ('Test set:', X_test.shape, y_test.shape)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <div id="classification">
# <h2>Classification</h2>
# </div>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <h3>Naive Bayes Classification</h3>
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Import library
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Classifier implementing the k-nearest neighbors vote.
#
# + button=false new_sheet=false run_control={"read_only": false}
from sklearn.naive_bayes import GaussianNB
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Training
#
# Lets start the algorithm with Gaussian Distribution now:
#
# + button=false new_sheet=false run_control={"read_only": false}
clf = GaussianNB()
clf.fit(X_train, y_train)
GaussianNB()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Predicting
#
# we can use the model to predict the test set:
#
# + button=false new_sheet=false run_control={"read_only": false}
yhat=(clf.predict(X_test))
yhat[0:5]
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Accuracy evaluation
#
# In multilabel classification, **accuracy classification score** is a function that computes subset accuracy. This function is equal to the jaccard_similarity_score function. Essentially, it calculates how closely the actual labels and predicted labels are matched in the test set.
#
# +
from sklearn import metrics
print("Test set Accuracy: ", metrics.accuracy_score(y_test, yhat))
# -
# ## Practice
#
# Please build your the Naive Bayes Classification Model with "numeric attributes tenure, age, income and employ" features here, with your own codes!
# (Calculate Normal Probabilitis for each feature and apply Bayes Theorem to decide tle class labels!)
#
# Compare your result with the given solution on upper part!
#
# +
# write your code here
class NaiveBayesClassifier:
def __init__(self):
self.class_attributes = ['custcat']
self.numeric_attributes = ['tenure', 'age', 'income', 'employ']
def train(self,X_train,Y_train):
X_train = np.delete(X_train,[0,3,4,6,8,9,10],axis = 1)
self.uniques, self.counts = np.unique(Y_train,return_counts=True)
self.classProbablity = self.counts/np.sum(self.counts)
self.matrixMean = np.zeros((len(self.numeric_attributes),len(self.uniques)))
self.matrixStd = np.zeros((len(self.numeric_attributes),len(self.uniques)))
for attr in range(len(self.numeric_attributes)):
for cls in range(len(self.uniques)):
mean = np.mean(X_train[np.where(Y_train==self.uniques[cls])][:,attr])
stdDev = np.std(X_train[np.where(Y_train==self.uniques[cls])][:,attr])
#matrislere kaydets
self.matrixMean[attr,cls] = mean
self.matrixStd[attr,cls] = stdDev
def Gaussian(self,value,means,std):
return np.exp((1/(2*(std**2))*(-((value-means)**2)) )) / (np.sqrt(2*np.pi)*std)
def predict(self,X_test):
X_test = np.delete(X_test,[0,3,4,6,8,9,10],axis = 1)
y_pred = np.zeros(len(X_test))
for sample in range(len(X_test)):
probs = np.ones(4)
for cls in range(len(self.uniques)):
for attr in range(len(X_test[sample,:])):
probs[cls]*=self.Gaussian(X_test[sample,attr],self.matrixMean[attr,cls],self.matrixStd[attr,cls])
probs[cls]*= self.classProbablity[cls]
y_pred[sample] = int(np.argmax(probs)+1)
print(y_pred)
return y_pred
# -
model = NaiveBayesClassifier()
model.train(X_train, y_train)
y_pred = model.predict(X_test)
np.where((y_pred == y_test),1,0).sum() / 200
|
naiveBayesClassifier/naiveBayes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Name: <NAME>
# # E-mail: <EMAIL>
# # Linear Regression
# # Project: Bike Sharing Demand
# # Analyzed from Kaggle Competition data set
# ## Agenda
#
# 1. Introducing the bikeshare dataset
# - Reading in the data
# - Visualizing the data
# 2. Linear regression basics
# - Form of linear regression
# - Building a linear regression model
# - Using the model for prediction
# - Does the scale of the features matter?
# 3. Working with multiple features
# - Visualizing the data (part 2)
# - Adding more features to the model
# 4. Choosing between models
# - Feature selection
# - Evaluation metrics for regression problems
# - Comparing models with train/test split and RMSE
# - Comparing testing RMSE with null RMSE
# 5. Creating features
# - Handling categorical features
# - Feature engineering
#
# <h1>Bike Sharing Demand</h1>
#
# <h2>Forecast use of a city bikeshare system</h2>
# <p/>
# You are provided hourly rental data spanning two years. For this competition, the training set is comprised of the first 19 days of each month, while the test set is the 20th to the end of the month. You must <b>predict the <u>total count of bikes rented</u> during each hour</b> covered by the test set, using only information available prior to the rental period.
# <p/>
# <b>Data Fields</b>
# <br/>
# datetime - hourly date + timestamp<br/>
# season - 1 = spring, 2 = summer, 3 = fall, 4 = winter <br/>
# holiday - whether the day is considered a holiday<br/>
# workingday - whether the day is neither a weekend nor holiday<br/>
# weather - 1: Clear, Few clouds, Partly cloudy, Partly cloudy
# 2: Mist + Cloudy, Mist + Broken clouds, Mist + Few clouds, Mist
# 3: Light Snow, Light Rain + Thunderstorm + Scattered clouds, Light Rain + Scattered clouds
# 4: Heavy Rain + Ice Pallets + Thunderstorm + Mist, Snow + Fog <br/>
# temp - temperature in Celsius<br/>
# atemp - "feels like" temperature in Celsius<br/>
# humidity - relative humidity<br/>
# windspeed - wind speed<br/>
# casual - number of non-registered user rentals initiated<br/>
# registered - number of registered user rentals initiated<br/>
# count - number of total rentals<br/>
# ## Reading in the data
#
# We'll be working with a dataset from Capital Bikeshare that was used in a Kaggle competition ([data dictionary](https://www.kaggle.com/c/bike-sharing-demand/data)).
# read the data and set the datetime as the index
import pandas as pd
url = 'https://raw.githubusercontent.com/codebuild81/Bike_Sharing_Demand/master/Data/bikeshare.csv'
bikes = pd.read_csv(url, index_col='datetime', parse_dates=True)
#len(bikes)
bikes.head()
# **Questions:**
#
# - What does each observation represent?
# - What is the response variable (as defined by Kaggle)?
# - How many features are there?
# "count" is a method, so it's best to name that column something else
bikes.rename(columns={'count':'total'}, inplace=True)
# Placeholder for any queries
#bikes.dtypes
bikes.describe()
# ## Visualizing the data
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (8, 6)
plt.rcParams['font.size'] = 14
# Pandas scatter plot
bikes.plot(kind='scatter', x='temp', y='total', alpha=0.2)
# Seaborn scatter plot with regression line
sns.lmplot(x='temp', y='total', data=bikes, aspect=1.5, scatter_kws={'alpha':0.2})
# ## Form of linear regression
#
# $y = \beta_0 + \beta_1x_1 + \beta_2x_2 + ... + \beta_nx_n$
#
# - $y$ is the response
# - $\beta_0$ is the intercept
# - $\beta_1$ is the coefficient for $x_1$ (the first feature)
# - $\beta_n$ is the coefficient for $x_n$ (the nth feature)
#
# The $\beta$ values are called the **model coefficients**:
#
# - These values are estimated (or "learned") during the model fitting process using the **least squares criterion**.
# - Specifically, we are going to find the line (mathematically) which minimizes the **sum of squared residuals** (or "sum of squared errors").
# - And once we've learned these coefficients, we can use the model to predict the response.
#
# <img src="https://raw.githubusercontent.com/upxacademy/ML_with_Python/master/images/estimating_coefficients.png?token=<PASSWORD>%3D%3D"></img>
#
# In the diagram above:
#
# - The black dots are the **observed values** of x and y.
# - The blue line is our **least squares line**.
# - The red lines are the **residuals**, which are the vertical distances between the observed values and the least squares line.
# ## Building a linear regression model
#
# Refer : http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.LinearRegression.html
# create X and y
feature_cols = ['temp']
X = bikes[feature_cols]
y = bikes.total
# import, instantiate, fit
from sklearn.linear_model import LinearRegression
linreg = LinearRegression()
linreg.fit(X, y)
# print the coefficients
print (linreg.intercept_)
print (linreg.coef_)
# Interpreting the **intercept** ($\beta_0$):
#
# - It is the value of $y$ when $x$=0.
# - Thus, it is the estimated number of rentals when the temperature is 0 degrees Celsius.
# - **Note:** It does not always make sense to interpret the intercept. (Why?)
#
# Interpreting the **"temp" coefficient** ($\beta_1$):
#
# - It is the change in $y$ divided by change in $x$, or the "slope".
# - Thus, a temperature increase of 1 degree Celsius is **associated with** a rental increase of 9.17 bikes.
# - This is not a statement of causation.
# - $\beta_1$ would be **negative** if an increase in temperature was associated with a **decrease** in rentals.
# ## Using the model for prediction
#
# How many bike rentals would we predict if the temperature was 25 degrees Celsius?
# manually calculate the prediction
linreg.intercept_ + linreg.coef_*25
# use the predict method
linreg.predict(25)
# ## Does the scale of the features matter?
#
# Let's say that temperature was measured in Fahrenheit, rather than Celsius. How would that affect the model?
# create a new column for Fahrenheit temperature
bikes['temp_F'] = bikes.temp * 1.8 + 32
bikes.head()
# Seaborn scatter plot with regression line
sns.lmplot(x='temp_F', y='total', data=bikes, aspect=1.5, scatter_kws={'alpha':0.2})
# +
# create X and y
feature_cols = ['temp_F']
X = bikes[feature_cols]
y = bikes.total
# instantiate and fit
linreg = LinearRegression()
linreg.fit(X, y)
# print the coefficients
print (linreg.intercept_)
print (linreg.coef_)
# -
# convert 25 degrees Celsius to Fahrenheit
25 * 1.8 + 32
# predict rentals for 77 degrees Fahrenheit
linreg.predict(77)
# **Conclusion:** The scale of the features is **irrelevant** for linear regression models. When changing the scale, we simply change our **interpretation** of the coefficients.
# remove the temp_F column
bikes.drop('temp_F', axis=1, inplace=True)
# ## Visualizing the data (part 2)
# explore more features
feature_cols = ['temp', 'season', 'weather', 'humidity']
# multiple scatter plots in Seaborn
sns.pairplot(bikes, x_vars=feature_cols, y_vars='total', kind='reg')
# multiple scatter plots in Pandas
fig, axs = plt.subplots(1, len(feature_cols), sharey=True)
for index, feature in enumerate(feature_cols):
bikes.plot(kind='scatter', x=feature, y='total', ax=axs[index], figsize=(16, 3))
# Are you seeing anything that you did not expect?
# box plot of rentals, grouped by season
bikes.boxplot(column='total', by='season')
# Notably:
#
# - A line can't capture a non-linear relationship.
# - There are more rentals in winter than in spring (?)
# line plot of rentals
bikes.total.plot()
# What does this tell us?
#
# There are more rentals in the winter than the spring, but only because the system is experiencing **overall growth** and the winter months happen to come after the spring months.
# ## Adding more features to the model
# create a list of features
feature_cols = ['temp', 'season', 'weather', 'humidity']
# +
# create X and y
X = bikes[feature_cols]
y = bikes.total
# instantiate and fit
linreg = LinearRegression()
linreg.fit(X, y)
# print the coefficients
print (linreg.intercept_)
print (linreg.coef_)
# -
# pair the feature names with the coefficients
list(zip(feature_cols, linreg.coef_))
# Interpreting the coefficients:
#
# - Holding all other features fixed, a 1 unit increase in **temperature** is associated with a **rental increase of 7.86 bikes**.
# - Holding all other features fixed, a 1 unit increase in **season** is associated with a **rental increase of 22.5 bikes**.
# - Holding all other features fixed, a 1 unit increase in **weather** is associated with a **rental increase of 6.67 bikes**.
# - Holding all other features fixed, a 1 unit increase in **humidity** is associated with a **rental decrease of 3.12 bikes**.
#
# Does anything look incorrect?
# ## Feature selection
#
# How do we choose which features to include in the model? We're going to use **train/test split** (and eventually **cross-validation**).
#
# Why not use of **p-values** or **R-squared** for feature selection?
#
# - Linear models rely upon **a lot of assumptions** (such as the features being independent), and if those assumptions are violated, p-values and R-squared are less reliable. Train/test split relies on fewer assumptions.
# - Features that are unrelated to the response can still have **significant p-values**.
# - Adding features to your model that are unrelated to the response will always **increase the R-squared value**, and adjusted R-squared does not sufficiently account for this.
# - p-values and R-squared are **proxies** for our goal of generalization, whereas train/test split and cross-validation attempt to **directly estimate** how well the model will generalize to out-of-sample data.
#
# More generally:
#
# - There are different methodologies that can be used for solving any given data science problem, and this course follows a **machine learning methodology**.
# - This course focuses on **general purpose approaches** that can be applied to any model, rather than model-specific approaches.
# example true and predicted response values
true = [10, 7, 5, 5]
pred = [8, 6, 5, 10]
# calculate these metrics by hand!
from sklearn import metrics
import numpy as np
print ('MAE:', metrics.mean_absolute_error(true, pred))
print ('MSE:', metrics.mean_squared_error(true, pred))
print ('RMSE:', np.sqrt(metrics.mean_squared_error(true, pred)))
# +
# same true values as above
true = [10, 7, 5, 5]
# new set of predicted values
pred = [10, 7, 5, 13]
# MAE is the same as before
print ('MAE:', metrics.mean_absolute_error(true, pred))
# MSE and RMSE are larger than before
print ('MSE:', metrics.mean_squared_error(true, pred))
print ('RMSE:', np.sqrt(metrics.mean_squared_error(true, pred)))
# -
# ## Comparing models with train/test split and RMSE
# +
from sklearn.model_selection import train_test_split
# define a function that accepts a list of features and returns testing RMSE
def train_test_rmse(feature_cols):
X = bikes[feature_cols]
y = bikes.total
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=123)
linreg = LinearRegression()
linreg.fit(X_train, y_train)
y_pred = linreg.predict(X_test)
return np.sqrt(metrics.mean_squared_error(y_test, y_pred))
# -
# compare different sets of features
print (train_test_rmse(['temp', 'season', 'weather', 'humidity']))
print (train_test_rmse(['temp', 'season', 'weather']))
print (train_test_rmse(['temp', 'season', 'humidity']))
# using these as features is not allowed!
print (train_test_rmse(['casual', 'registered']))
# ## Comparing testing RMSE with null RMSE
#
# Null RMSE is the RMSE that could be achieved by **always predicting the mean response value**. It is a benchmark against which you may want to measure your regression model.
# +
# split X and y into training and testing sets
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=123)
# create a NumPy array with the same shape as y_test
y_null = np.zeros_like(y_test, dtype=float)
# fill the array with the mean value of y_test
y_null.fill(y_test.mean())
y_null
# -
# compute null RMSE
np.sqrt(metrics.mean_squared_error(y_test, y_null))
# ## Handling categorical features
#
# scikit-learn expects all features to be numeric. So how do we include a categorical feature in our model?
# For season, we can't simply leave the encoding as 1 = spring, 2 = summer, 3 = fall, and 4 = winter, because that would imply an **ordered relationship**. Instead, we create **multiple dummy variables:**
# +
# create dummy variables
season_dummies = pd.get_dummies(bikes.season, prefix='season')
# print 5 random rows
season_dummies.sample(n=5, random_state=1)
# -
# However, we actually only need **three dummy variables (not four)**, and thus we'll drop the first dummy variable.
#
# Why? Because three dummies captures all of the "information" about the season feature, and implicitly defines spring (season 1) as the **baseline level:**
# +
# drop the first column
season_dummies.drop(season_dummies.columns[0], axis=1, inplace=True)
# print 5 random rows
season_dummies.sample(n=5, random_state=1)
# -
# In general, if you have a categorical feature with **k possible values**, you create **k-1 dummy variables**.
#
# If that's confusing, think about why we only need one dummy variable for holiday, not two dummy variables (holiday_yes and holiday_no).
# +
# concatenate the original DataFrame and the dummy DataFrame (axis=0 means rows, axis=1 means columns)
bikes = pd.concat([bikes, season_dummies], axis=1)
# print 5 random rows
bikes.sample(n=5, random_state=1)
# -
# include dummy variables for season in the model
feature_cols = ['temp', 'season_2', 'season_3', 'season_4', 'humidity']
X = bikes[feature_cols]
y = bikes.total
linreg = LinearRegression()
linreg.fit(X, y)
list(zip(feature_cols, linreg.coef_))
# How do we interpret the season coefficients? They are **measured against the baseline (spring)**:
#
# - Holding all other features fixed, **summer** is associated with a **rental decrease of 3.39 bikes** compared to the spring.
# - Holding all other features fixed, **fall** is associated with a **rental decrease of 41.7 bikes** compared to the spring.
# - Holding all other features fixed, **winter** is associated with a **rental increase of 64.4 bikes** compared to the spring.
#
# Would it matter if we changed which season was defined as the baseline?
#
# - No, it would simply change our **interpretation** of the coefficients.
#
# **Important:** Dummy encoding is relevant for all machine learning models, not just linear regression models.
# compare original season variable with dummy variables
print (train_test_rmse(['temp', 'season', 'humidity']))
print (train_test_rmse(['temp', 'season_2', 'season_3', 'season_4', 'humidity']))
# ## Feature engineering
#
# See if you can create the following features:
#
# - **hour:** as a single numeric feature (0 through 23)
# - **hour:** as a categorical feature (use 23 dummy variables)
# - **daytime:** as a single categorical feature (daytime=1 from 7am to 8pm, and daytime=0 otherwise)
#
# Then, try using each of the three features (on its own) with `train_test_rmse` to see which one performs the best!
# hour as a numeric feature
bikes['hour'] = bikes.index.hour
bikes.head(2)
# hour as a categorical feature
hour_dummies = pd.get_dummies(bikes.hour, prefix='hour')
hour_dummies.drop(hour_dummies.columns[0], axis=1, inplace=True)
bikes = pd.concat([bikes, hour_dummies], axis=1)
# daytime as a categorical feature
bikes['daytime'] = ((bikes.hour > 6) & (bikes.hour < 21)).astype(int)
print (train_test_rmse(['hour']))
print (train_test_rmse(bikes.columns[bikes.columns.str.startswith('hour_')]))
print (train_test_rmse(['daytime']))
#
|
code/ML_BikeSharingDemand_LR_Kaggle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (sklearn_env)
# language: python
# name: sklearn_env
# ---
print("hello")
# +
# work with strings
# (lists loops and conditionals)
# read external text as a file
# working with longer text documents
# cleaning, parsing, preparing a text document
# read in multiple text files
# build a pandas dataframe
# -
positive_review = """This was an excellent film. Good acting,
well written script, entertaining plot. "Not bad at all"."""
negative_review = """Poorly done film. Bad acting, boring plot, stilted dialogue, not good at all."""
print(positive_review)
print(negative_review)
len(positive_review)
len(negative_review)
positive_review[5]
positive_review[5:10]
positive_review[-1]
positive_review[10:-10]
# +
# this will print every character. (what if I wanted to print every *word*)
#for c in positive_review:
# print(c)
# -
pos_words = positive_review.split()
neg_words = negative_review.split()
print(pos_words)
print(neg_words)
for w in pos_words:
print(w)
# +
# think for 5 min - what would useful about the two sentences. what modifications might you want to make?
# resume at 10:52
# +
# punctuation
# capitalization
# stop words
# short words
# -
a_word = "HeLlo"
print(a_word)
print(a_word.lower())
a_word = a_word.lower()
a_word
# +
# write a loop to print the lower case version of all words in the positive review
# take 5 min 10:03
# -
pos_lower_case = []
for w in pos_words:
pos_lower_case.append(w.lower())
pos_lower_case
neg_lower_case = []
for w in neg_words:
neg_lower_case.append(w.lower())
print(pos_lower_case)
print(neg_lower_case)
import re
# +
# remove everything from a string that isn't alphanumeric
# -
a_word = "<>hello!%"
a_word
re.sub(r'\W+', '', a_word)
# +
# we can now write a loop that removes all punctuation from our string
# +
# make an empty list called "no_punct"
# write a loop to populate it with the words with all non-alphanum chars removed
# resume aat 10:16
# -
pos_no_punct = []
for w in pos_lower_case:
pos_no_punct.append(re.sub(r'\W+', '', w))
pos_no_punct
stop_words = ['this', 'the', 'a', 'an', 'in', 'at', 'by']
'this' not in stop_words
no_stop_words = []
for w in pos_no_punct:
if w not in stop_words:
no_stop_words.append(w)
no_stop_words
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
# !pip install nltk
stemmer = PorterStemmer()
lemmatizer = WordNetLemmatizer()
import nltk
nltk.download('wordnet')
print(stemmer.stem('narrators'))
print(lemmatizer.lemmatize('narrators'))
print(stemmer.stem('narrations'))
print(lemmatizer.lemmatize('narrations'))
stemmed_words = []
for w in no_stop_words:
stemmed_words.append(stemmer.stem(w))
stemmed_words
lemmatized_words = []
for w in no_stop_words:
lemmatized_words.append(lemmatizer.lemmatize((w)))
lemmatized_words
' '.join(stemmed_words)
' '.join(lemmatized_words)
# +
# we will wrap up at 11:30 today
# +
# take a 10 minute break
# resume at 10:55
# when we resume, we'll go over how to read lots of files into a pandas dataframe
# see the Read-All-Files notebook
# -
|
Working-With-Text.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os, shutil
# ## Data preprocessing
# +
# List all filenames in the master dataset and count how many samples there are
originalDir = './retinopathy-dataset-master'
class1 = 'nosymptoms'
originalNoSymptomsDir = os.path.join(originalDir, class1)
noSymptomsFnames = os.listdir(originalNoSymptomsDir)
class2 = 'symptoms'
originalSymptomsDir = os.path.join(originalDir, class2)
symptomsFnames = os.listdir(originalSymptomsDir)
# +
baseDir = './dataset'
#Training
subDir = 'train'
trainDir = os.path.join(baseDir, subDir)
trainNoSymptomsDir = os.path.join(baseDir, subDir, class1)
trainSymptomsDir = os.path.join(baseDir, subDir, class2)
#Validation
subDir = 'validation'
valDir = os.path.join(baseDir, subDir)
valNoSymptomsDir = os.path.join(baseDir, subDir, class1)
valSymptomsDir = os.path.join(baseDir, subDir, class2)
#Test
subDir = 'test'
testDir = os.path.join(baseDir, subDir)
testNoSymptomsDir = os.path.join(baseDir, subDir, class1)
testSymptomsDir = os.path.join(baseDir, subDir, class2)
# -
#Create folders
try:
os.mkdir(baseDir)
os.mkdir(trainDir)
os.mkdir(trainNoSymptomsDir)
os.mkdir(trainSymptomsDir)
os.mkdir(valDir)
os.mkdir(valNoSymptomsDir)
os.mkdir(valSymptomsDir)
os.mkdir(testDir)
os.mkdir(testNoSymptomsDir)
os.mkdir(testSymptomsDir)
except:
print('Error in creating folders. This step might be done already')
# +
from sklearn.model_selection import train_test_split
from numpy import random
trainSymptomsFnames, testSymptomsFnames = train_test_split(symptomsFnames, test_size=0.10)
trainSymptomsFnames, valSymptomsFnames = train_test_split(trainSymptomsFnames, test_size=0.20)
len(trainSymptomsFnames), len(valSymptomsFnames), len(testSymptomsFnames)
# +
trainNoSymptomsFnames, testNoSymptomsFnames = train_test_split(noSymptomsFnames, test_size=0.10)
trainNoSymptomsFnames, valNoSymptomsFnames = train_test_split(trainNoSymptomsFnames, test_size=0.20)
len(trainNoSymptomsFnames), len(valNoSymptomsFnames), len(testNoSymptomsFnames)
# +
import time
start = time.time()
#Training
#Disease
for fname in trainSymptomsFnames:
src = os.path.join(originalSymptomsDir, fname)
dst = os.path.join(trainSymptomsDir, fname)
shutil.copyfile(src, dst)
#Healthy
for fname in trainNoSymptomsFnames:
src = os.path.join(originalNoSymptomsDir, fname)
dst = os.path.join(trainNoSymptomsDir, fname)
shutil.copyfile(src, dst)
#Validation
#Disease
for fname in valSymptomsFnames:
src = os.path.join(originalSymptomsDir, fname)
dst = os.path.join(valSymptomsDir, fname)
shutil.copyfile(src, dst)
#Healthy
for fname in valNoSymptomsFnames:
src = os.path.join(originalNoSymptomsDir, fname)
dst = os.path.join(valNoSymptomsDir, fname)
shutil.copyfile(src, dst)
#Test
#Disease
for fname in testSymptomsFnames:
src = os.path.join(originalSymptomsDir, fname)
dst = os.path.join(testSymptomsDir, fname)
shutil.copyfile(src, dst)
#Healthy
for fname in testNoSymptomsFnames:
src = os.path.join(originalNoSymptomsDir, fname)
dst = os.path.join(testNoSymptomsDir, fname)
shutil.copyfile(src, dst)
end = time.time()
print('Time {:.2f}'.format(end-start))
# -
|
Case 2 preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import os
import torch
import pandas as pd
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
plt.ion() # interactive mode
# -
# ## Download the dataset
# +
import os
import zipfile
from six.moves import urllib
import zipfile
DATASET_URL = "https://download.pytorch.org/tutorial/faces.zip"
DATASET_PATH = os.path.join(os.getcwd(), "data", "")
DATASET_FILE = os.path.join(DATASET_PATH, "faces.zip")
def fetch_data(url=DATASET_URL, path=DATASET_PATH, zip_path=DATASET_FILE) -> bool:
if not os.path.isdir(path):
os.makedirs(path)
urllib.request.urlretrieve(url, zip_path)
if (os.path.isfile(zip_path)):
print(f'Download path: {zip_path}')
return True
else:
return False
def extract_zip(zip_file=DATASET_FILE, unzip_folder=DATASET_PATH):
with zipfile.ZipFile(zip_file, 'r') as zip_ref:
zip_ref.extractall(unzip_folder)
if fetch_data():
extract_zip()
# +
landmarks_frame = pd.read_csv('data/faces/face_landmarks.csv')
print(landmarks_frame.head(3))
n = 65
img_name = landmarks_frame.iloc[n, 0]
landmarks = landmarks_frame.iloc[n, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
print('Image name: {}'.format(img_name))
print('Landmarks shape: {}'.format(landmarks.shape))
print('First 4 Landmarks: {}'.format(landmarks[:4]))
# +
def show_landmarks(image, landmarks):
"""Show image with landmarks"""
plt.imshow(image)
plt.scatter(landmarks[:, 0], landmarks[:, 1], s=10, marker='.', c='r')
plt.pause(0.001) # pause a bit so that plots are updated
plt.figure()
show_landmarks(io.imread(os.path.join('data/faces/', img_name)),
landmarks)
plt.show()
# -
# ## Dataset class
class FaceLandmarksDataset(Dataset):
"""Face Landmarks dataset."""
def __init__(self, csv_file, root_dir, transform=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.landmarks_frame = pd.read_csv(csv_file)
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.landmarks_frame)
def __getitem__(self, idx):
img_name = os.path.join(self.root_dir,
self.landmarks_frame.iloc[idx, 0])
image = io.imread(img_name)
landmarks = self.landmarks_frame.iloc[idx, 1:].as_matrix()
landmarks = landmarks.astype('float').reshape(-1, 2)
sample = {'image': image, 'landmarks': landmarks}
if self.transform:
sample = self.transform(sample)
return sample
# +
face_dataset = FaceLandmarksDataset(csv_file='data/faces/face_landmarks.csv',
root_dir='data/faces/')
fig = plt.figure()
for i in range(len(face_dataset)):
sample = face_dataset[i]
print(i, sample['image'].shape, sample['landmarks'].shape)
ax = plt.subplot(1, 4, i + 1)
plt.tight_layout()
ax.set_title('Sample #{}'.format(i))
ax.axis('off')
show_landmarks(**sample)
if i == 3:
plt.show()
break
# -
# ## Transforms
# +
class Rescale(object):
"""Rescale the image in a sample to a given size.
Args:
output_size (tuple or int): Desired output size. If tuple, output is
matched to output_size. If int, smaller of image edges is matched
to output_size keeping aspect ratio the same.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
if isinstance(self.output_size, int):
if h > w:
new_h, new_w = self.output_size * h / w, self.output_size
else:
new_h, new_w = self.output_size, self.output_size * w / h
else:
new_h, new_w = self.output_size
new_h, new_w = int(new_h), int(new_w)
img = transform.resize(image, (new_h, new_w))
# h and w are swapped for landmarks because for images,
# x and y axes are axis 1 and 0 respectively
landmarks = landmarks * [new_w / w, new_h / h]
return {'image': img, 'landmarks': landmarks}
class RandomCrop(object):
"""Crop randomly the image in a sample.
Args:
output_size (tuple or int): Desired output size. If int, square crop
is made.
"""
def __init__(self, output_size):
assert isinstance(output_size, (int, tuple))
if isinstance(output_size, int):
self.output_size = (output_size, output_size)
else:
assert len(output_size) == 2
self.output_size = output_size
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
h, w = image.shape[:2]
new_h, new_w = self.output_size
top = np.random.randint(0, h - new_h)
left = np.random.randint(0, w - new_w)
image = image[top: top + new_h,
left: left + new_w]
landmarks = landmarks - [left, top]
return {'image': image, 'landmarks': landmarks}
class ToTensor(object):
"""Convert ndarrays in sample to Tensors."""
def __call__(self, sample):
image, landmarks = sample['image'], sample['landmarks']
# swap color axis because
# numpy image: H x W x C
# torch image: C X H X W
image = image.transpose((2, 0, 1))
return {'image': torch.from_numpy(image),
'landmarks': torch.from_numpy(landmarks)}
# -
# ## Compose transforms
# +
scale = Rescale(256)
crop = RandomCrop(128)
composed = transforms.Compose([Rescale(256),
RandomCrop(224)])
# Apply each of the above transforms on sample.
fig = plt.figure()
sample = face_dataset[65]
for i, tsfrm in enumerate([scale, crop, composed]):
transformed_sample = tsfrm(sample)
ax = plt.subplot(1, 3, i + 1)
plt.tight_layout()
ax.set_title(type(tsfrm).__name__)
show_landmarks(**transformed_sample)
plt.show()
# +
transformed_dataset = FaceLandmarksDataset(csv_file='data/faces/face_landmarks.csv',
root_dir='data/faces/',
transform=transforms.Compose([
Rescale(256),
RandomCrop(224),
ToTensor()
]))
for i in range(len(transformed_dataset)):
sample = transformed_dataset[i]
print(i, sample['image'].size(), sample['landmarks'].size())
if i == 3:
break
# +
dataloader = DataLoader(transformed_dataset, batch_size=4,
shuffle=True, num_workers=4)
# Helper function to show a batch
def show_landmarks_batch(sample_batched):
"""Show image with landmarks for a batch of samples."""
images_batch, landmarks_batch = \
sample_batched['image'], sample_batched['landmarks']
batch_size = len(images_batch)
im_size = images_batch.size(2)
print(f"batch size: {batch_size}")
print(f"images_batch shape: {images_batch.shape}")
print(f"image size: {im_size}")
grid = utils.make_grid(images_batch)
plt.imshow(grid.numpy().transpose((1, 2, 0)))
for i in range(batch_size):
plt.scatter(landmarks_batch[i, :, 0].numpy() + i * im_size,
landmarks_batch[i, :, 1].numpy(),
s=10, marker='.', c='r')
plt.title('Batch from dataloader')
for i_batch, sample_batched in enumerate(dataloader):
print(i_batch, sample_batched['image'].size(),
sample_batched['landmarks'].size())
# observe 4th batch and stop.
if i_batch == 3:
plt.figure()
show_landmarks_batch(sample_batched)
plt.axis('off')
plt.ioff()
plt.show()
break
# -
|
notebook/pytorch/data_loading.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Quantum Computing 101: Hello Quantum World!
# ### [WIQCA Seattle Meetup](https://www.wiqca.dev/events/quantum101-helloquantumworld.html) #
#
# <NAME> | @crazy4pi314 | April 1, 2020
#
# ---
#
# Talk slides/Jupyter Notebook can be found at [bit.ly/wiqca-quantum101](https://bit.ly/wiqca-quantum101)
#
#
# + [markdown] slideshow={"slide_type": "skip"}
# ### Abstract
# Quantum computing is an exciting and growing field. To realize quantum applications, we need a new generation of programmers ready to leverage quantum technologies. As one of the most actively used languages in scientific computing, Python lets us reduce barriers to engaging new learners in quantum computing.
#
# In this talk, Sarah will introduce quantum computing hands-on, and will show how Python can be used to work new quantum programming languages like Q#, an open-source high-level language for quantum computing from Microsoft. Using Python and Q# together, Sarah will demo core quantum computing, and will share resources on how you can get started learning and developing for a quantum computer, and how you can join the growing quantum community!
#
# ---
#
# #### Installation instructions for running this notebook on your machine can be found [here](https://docs.microsoft.com/en-ca/quantum/install-guide/python?view=qsharp-preview).
# + [markdown] slideshow={"slide_type": "skip"}
# ## `about_me.md`
#
#
# <figure style="text-align: center;">
# <img src="media/about_me.png" width="70%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## 💪Goals💪
# + [markdown] slideshow={"slide_type": "-"}
# I want to show **you**:
#
# - the skills you already have are a great start for quantum development
# - that you can learn the rest you need as you go
#
# in other words...
#
# _You can jump into writing code for a quantum computer today!_
# + [markdown] slideshow={"slide_type": "slide"}
# # Quantum Computing
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quantum computers *are not* :
#
# 🦇 spooky
#
# 🙃 weird
#
# 💞 in two places at once
#
# 💻 going to replace your regular computer
#
# 🙀 cats (dead or alive)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quantum programs are classical programs
#
# - Quantum programs are just classical programs that emit instructions for quantum hardware.
#
# ```c#
# operation SayHello(name: String) : Unit {
# Message($"Hello World! Nice to meet you, {name}!");
# }
# ```
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quantum computers *are* :
#
# 🚄🖥 hardware accelerators (think GPUs)
#
# <br>
#
# <figure style="text-align: center;">
# <img src="media/what-is-qc.png" width="70%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
# + [markdown] slideshow={"slide_type": "slide"}
# ## How can Python help?
#
# There are **tons** of packages that can help you learn quantum computing, as well as write code for quantum computers.
#
# The one we will look at today is:
#
# - [`qsharp`](https://docs.microsoft.com/en-us/quantum/?view=qsharp-preview) - Python interoperabilty with Q#, a domain-specific programming language for quantum computers
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Python + Q# = 💖
#
# - Q# is a domain-specific programming language, included in the [Quantum Development Kit](https://docs.microsoft.com/en-us/quantum/install-guide/?view=qsharp-preview) (QDK) which is a set of development tools used for expressing quantum algorithms.
# - ❤ Open source ❤
# - Allows you to write code the same way you think about it (high level of abstraction)
# <figure style="text-align: center;">
# <img src="media/qsharp_software_stack.png" width="70%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
# + [markdown] slideshow={"slide_type": "slide"}
# #### For this talk we will be using a simulator target machine:
#
# <figure style="text-align: center;">
# <img src="media/qsharp_software_stack_highlight.png" width="70%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What skills can help you program a quantum computer?
#
# <figure style="text-align: center;">
# <img src="media/danger.png" width="50%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## What skills can help you program a quantum computer?
#
#
# - Version control
# - Open source community tools (Pull requests, filing issues, etc)
# - Reading documentation
# - A bit of linear algebra, similar to data science or ML
# - A bit of math like complex numbers and trigonometry
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # ⌚Demo Time⌚
# + [markdown] slideshow={"slide_type": "-"}
# Let's get started with Python by loading the package for Q# interoperability called `qsharp`.
# + slideshow={"slide_type": "-"}
import qsharp
qsharp.component_versions()
# -
qsharp.reload()
# + [markdown] slideshow={"slide_type": "slide"}
# # Task: Quantum random numbers
#
# We want to use a truly* random source to generate a list of random bits like this:
# + slideshow={"slide_type": "-"}
randomness = [0,1,0,0,1,1,0,1,0,1]
# -
# _Bonus points_ : share this randomness without sending the classical bits (send quantum information instead)
#
# <tiny>*still simulated here so still pseudo-random</tiny>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Generating _quantum_ random numbers with Q\# ##
#
# ```c#
# // demo.qs
# namespace Wiqca.Demo {
# operation Qrng() : Result {
# using (qubit = Qubit()) { // Preparing the qubit
# H(qubit); // Do operation H
# return MResetZ(qubit); // Measure and reset qubit
# }
# }
# }
# ```
# How can we dive in to what is going on here?
# + [markdown] slideshow={"slide_type": "slide"}
# ### Let's load the Q# code from Python!
# + slideshow={"slide_type": "-"}
from Wiqca.Demo import Qrng
# + [markdown] slideshow={"slide_type": "slide"}
# ## Understanding `Qrng`
#
# We can use built-in documentation strings, just like we can with Python functions.
# + slideshow={"slide_type": "-"}
# ?Qrng
# + [markdown] slideshow={"slide_type": "fragment"}
# That tells us what we can **do** with `Qrng`:
# + slideshow={"slide_type": "fragment"}
[Qrng.simulate() for _ in range(10)]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Hold up: What is a qubit?
# + [markdown] slideshow={"slide_type": "-"}
# - Answer: a single unit of information in a quantum computer
# - _quantum + bit = qubit_
# + [markdown] slideshow={"slide_type": "fragment"}
# - We can predict what a single qubit will do by using a column vector of 2 complex numbers* like this:
#
# <!--$\left|{x}\right\rangle = \left[\begin{matrix} 1 + 0\times i \\0 + 0\times i \end{matrix}\right]$
#
#
# -->
# + slideshow={"slide_type": "-"}
import numpy as np
qubit = np.array([[1],[0]],dtype=complex)
print(qubit)
# + [markdown] slideshow={"slide_type": "slide"}
# ## What can we _do_ with a qubit?
# + [markdown] slideshow={"slide_type": "-"}
# Similar to classical bits on your computer, you can do three types of things with qubits:
#
# - Prepare a qubit
# - Do operations with a qubit
# - Measure a qubit : returns a 0 or 1
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# ```c#
# // demo.qs
# namespace Wiqca.Demo {
# operation Qrng() : Result {
# using (qubit = Qubit()) { // Preparing
# H(qubit); // Operation
# return MResetZ(qubit); // Measure and reset
# }
# }
# }
# ```
# + [markdown] slideshow={"slide_type": "slide"}
# ### How can we "get" a qubit?
# + slideshow={"slide_type": "-"}
prepare_qubit = qsharp.compile("""
open Microsoft.Quantum.Diagnostics;
operation PrepareQubit() : Unit {
using (qubit = Qubit()) { // We want 1 qubit to use for our task
DumpMachine(); // Print out what the simulator is keeping a record of
}
}
""")
# + slideshow={"slide_type": "-"}
prepare_qubit.simulate()
# + [markdown] slideshow={"slide_type": "skip"}
# You can read the above output like the vector we wrote above, where the first column is the index, the second is the real part of the vector at that position, and the second is the complex part of that vector entry.
# + [markdown] slideshow={"slide_type": "slide"}
# What does `DumpMachine` tell us?
#
# ```
# |0⟩ 1 + 0𝑖
# |1⟩ 0 + 0𝑖
# ```
#
# This is the same state we saw earlier!
# + slideshow={"slide_type": "-"}
print(qubit)
# + [markdown] slideshow={"slide_type": "slide"}
# ### Learning quantum operations by inspection
# + [markdown] slideshow={"slide_type": "-"}
# We can use `DumpMachine` again to understand see what the `H` operation does to our qubit.
# + slideshow={"slide_type": "-"}
from Wiqca.Demo import QrngWithDiagnostics
QrngWithDiagnostics.simulate()
# + [markdown] slideshow={"slide_type": "slide"}
# The operation `H` on our qubit puts our simulated qubit in **superposition**
#
# ```
# After using H(qubit) to create a superposition state:
# |0⟩ 0.7071067811865476 + 0𝑖
# |1⟩ 0.7071067811865476 + 0𝑖
# ```
#
# #### 🚨Note: `DumpMachine` is showing the information the simulator has!🚨
# + [markdown] slideshow={"slide_type": "slide"}
# # How about more qubits?!
#
#
#
# <figure style="text-align: center;">
# <img src="https://disneygenderevolution.files.wordpress.com/2014/12/ariel-the-little-mermaid-i-want-more-gif.gif" width="60%">
# <caption>
# <br>
# <strong></strong>
# </caption>
# </figure>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Operations with multiple qubits can create 💕entanglement💕
# + [markdown] slideshow={"slide_type": "-"}
# Using Q# with Python, we can also explore other quantum development tools, like **entanglement**.
# + slideshow={"slide_type": "-"}
from Wiqca.Demo import EntangleQubits
results = EntangleQubits.simulate(verbose=True)
# + [markdown] slideshow={"slide_type": "slide"}
# What does `DumpRegister` tell us this time?
# ```
# |0⟩ 0.7071067811865476 + 0𝑖
# |1⟩ 0 + 0𝑖
# |2⟩ 0 + 0𝑖
# |3⟩ 0.7071067811865476 + 0𝑖
# ```
# - ∣0❭➡ measuring both qubits give you (0,0)
# - ∣3❭➡ measuring both qubits give you (1,1)
# + [markdown] slideshow={"slide_type": "slide"}
# No matter how many times we run, both measurements are equal to each other!
# + slideshow={"slide_type": "-"}
[EntangleQubits.simulate(verbose=False) for _ in range(10)]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Next: share the randomness!
#
# - If you **entangle** two qubits and then share one, then you both measure you will have the same random number.
# - This can be useful for cryptographic protocols, like quantum key distribution!
#
# + slideshow={"slide_type": "-"}
[EntangleQubits.simulate(verbose=False) for _ in range(10)]
# + [markdown] slideshow={"slide_type": "slide"}
# ## Beam me up: Quantum teleportation
#
# - The most basic example of using entanglement in a quantum algoritihm is _Teleportation_.
# - This is similar to move operators and is used to move quantum data around.
#
# + slideshow={"slide_type": "-"}
from Wiqca.Demo import TeleportClassicalMessage
TeleportClassicalMessage.simulate(message=1)
# + [markdown] slideshow={"slide_type": "slide"}
# ```
# operation TeleportClassicalMessage (message : Bool) : Bool {
# // Ask for some qubits that we can use to teleport.
# using ((msg, target) = (Qubit(), Qubit())) {
#
# // Encode the message we want to send.
# if (message) {
# X(msg);
# }
#
# // Use the operation we defined above.
# Teleport(msg, target);
#
# // Check what message was sent.
# return MResetZ(target) == One;
# }
# }
# ```
# + [markdown] slideshow={"slide_type": "slide"}
#
# ```
# operation Teleport (msg : Qubit, target : Qubit) : Unit {
# using (register = Qubit()) {
# // Create some entanglement that we can use to send our message.
# H(register);
# CNOT(register, target);
#
# // Encode the message into the entangled pair.
# CNOT(msg, register);
# H(msg);
#
# // Measure the qubits to extract the classical data
# if (MResetZ(msg) == One) { Z(target); }
#
# if (IsResultOne(MResetZ(register))) { X(target); }
# }
# }
# ```
# + slideshow={"slide_type": "-"}
random_numbers = [Qrng.simulate() for _ in range(10)]
[(data, TeleportClassicalMessage.simulate(message=data)) for data in random_numbers]
# + [markdown] slideshow={"slide_type": "slide"}
# 
# + [markdown] slideshow={"slide_type": "slide"}
#
# ## Toy quantum algorithm: Deutsch–Jozsa
# + [markdown] slideshow={"slide_type": "-"}
# _If I had a function that had one bit input and output, how many different options would I have?_
# + [markdown] slideshow={"slide_type": "-"}
# <figure style="text-align: center;">
# <img src="media/twobit.png" width="50%">
# <caption>
# <br>
# <strong>Diagram of all possible one bit functions</strong>
# </caption>
# </figure>
# + [markdown] slideshow={"slide_type": "slide"}
# >#### Deutsch–Jozsa Algorithim ####
# >**Problem statement:**
# >
# >* **GIVEN:** A black box quantum operation which takes 1 input bit and produces either a 0 or a 1 as output. We are promised that the box is either _constant_ or _balanced_.
# >
# >* **GOAL:** to determine if the box output is _constant_ or _balanced_ by evaluating sample inputs.
#
# Deutsch–Jozsa can do this in **one** query to the black box!
# + [markdown] slideshow={"slide_type": "slide"}
# <figure style="text-align: center;">
# <img src="media/twobitDJ.png" width="40%">
# <caption>
# <br>
# <strong>Global property of the one bit functions: Constant or Balanced</strong>
# </caption>
# </figure>
# + [markdown] slideshow={"slide_type": "slide"}
# ## Let's load some Q\# code from the directory...
# + slideshow={"slide_type": "-"}
is_zero_oracle_balanced = qsharp.compile("""
open Wiqca.DeutschJozsa;
operation IsZeroOracleBalanced(): Bool {
return IsOracleBalanced(ZeroOracle);
}
""")
# + slideshow={"slide_type": "-"}
is_zero_oracle_balanced.simulate()
# + [markdown] slideshow={"slide_type": "slide"}
# <figure style="text-align: center;">
# <img src="media/twobitDJ.png" width="40%">
# <caption>
# <br>
# <strong>Global property of the one bit functions: Constant or Balanced</strong>
# </caption>
# </figure>
# + slideshow={"slide_type": "slide"}
is_not_oracle_balanced = qsharp.compile("""
open Wiqca.DeutschJozsa;
operation IsNotOracleBalanced(): Bool {
return IsOracleBalanced(NotOracle);
}
""")
# + slideshow={"slide_type": "-"}
is_not_oracle_balanced.simulate()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Putting it all together: One query, one answer!
# + slideshow={"slide_type": "-"}
from Wiqca.DeutschJozsa import RunDeutschJozsaAlgorithm
# + slideshow={"slide_type": "-"}
RunDeutschJozsaAlgorithm.simulate(verbose=False)
# + slideshow={"slide_type": "-"}
RunDeutschJozsaAlgorithm.simulate(verbose=True)
# + [markdown] slideshow={"slide_type": "slide"}
# # 📝 Review time 📝
#
# ## Tasks
# - Make random numbers ✔
# - Share classical data with quantum resources ✔
# - Use the Deutsch–Jozsa quantum algorithim to learn properties of a function with a single call ✔
#
# ## Concepts
# - Quantum computers use _superposition_ and _entanglement_ to enable their unique abilities ✔
# - Python + Q# tools and skills to learn quantum computing ✔
# + [markdown] slideshow={"slide_type": "slide"}
# ## What happens now?
#
# - Try Q# + Python for yourself!
# - Learn by teaching
# - Write blog posts
# - Make tutorials
# - Make the community better than you found it!
# - Contribute to docs
# - Fix bugs/File issues
# - Act intentionally to include everyone and expand the community 💖
# + [markdown] slideshow={"slide_type": "slide"}
# ## 👩💻Q# Quantum programming resources!👩💻
#
# - Community projects:
# - [qsharp.community](https://qsharp.community/)
# - [quantumcomputing.stackexchange.com](https://quantumcomputing.stackexchange.com/)
# - Q# Documentation: [docs.microsoft.com/quantum](docs.microsoft.com/quantum)
# - Books like [_Learn Quantum Computing with Python and Q#_](http://www.manning.com/?a_aid=learn-qc-kaiser) in early access
# - use **ctwmvp20** for 40% off at [bit.ly/qsharp-book](bit.ly/qsharp-book)
# - Seattle meetup group: **✨WIQCA✨**
# - Please join the Women in quantum computing and applications meetup page! [bit.ly/wiqca-email](http://bit.ly/wiqca-email)
#
#
# ### The Quantum 101 series will continue in May, watch the Meetup page or wiqca.dev for details!
#
#
#
# + [markdown] slideshow={"slide_type": "slide"}
# # Thank you!
#
# ## Tweet at @wiqca or hop on the slack if you want to chat in the meantime 🔮💖
# + [markdown] slideshow={"slide_type": "skip"}
# ---
#
# ## Helpful diagnostics :)
# + slideshow={"slide_type": "skip"}
for component, version in sorted(qsharp.component_versions().items(), key=lambda x: x[0]):
print(f"{component:20}{version}")
# + slideshow={"slide_type": "skip"}
import sys
print(sys.version)
|
demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PAMAP 2 Model 1: Artificial Neural Network
# #### Dataset Source: https://archive.ics.uci.edu/ml/datasets/PAMAP2+Physical+Activity+Monitoring
#
# This is the same model as our ANN, but tested on the dataset PAMAP2 to validate the architecture of our model. This does utilize the sliding window function.
#
# INPUT: pamap2_clean.csv. This file is over 1GB and can't be uploaded to our github. Please download this dataset from [Box link here?]()
#
# #### Cleaning code source: https://www.kaggle.com/avrahamcalev/time-series-models-pamap2-dataset
# ### Import libraries
# +
import pandas as pd
import numpy as np
import tensorflow as tf
import random
from numpy import mean
from numpy import std
import keras
from keras.models import Sequential
from keras.layers import Dense, Flatten, Dropout, LSTM, TimeDistributed, Conv1D, MaxPooling1D, Conv2D, MaxPooling2D, MaxPool2D
from keras.layers import Flatten
from keras.layers import Dropout
from keras.layers import LSTM
from keras.utils import to_categorical
from keras.utils import np_utils
from sklearn import metrics
from sklearn.metrics import classification_report
from sklearn.preprocessing import StandardScaler
# -
random.seed(321)
# ### Prepare data
df = pd.read_csv('../../../../10_code/40_usable_data_for_models/42_PAMAP2/pamap2_clean.csv')
df
# ### Take only relevant classes
# Classes 2, 3 and 4 represent the classes most representative for our data (walking, lying (DB), and standing).
# Take a look at the readme in the data source to pick other classes.
df = df[df['activity_id'].isin([2, 3, 4])]
# #### Drop these if your window size is 250 or below:
df1 = df.drop(df[(df['activity_id'] == 5) & (df['id'] == 104)].index)
df2 = df1.drop(df1[(df1['activity_id'] == 24) & (df['id'] == 106)].index)
no_act = ['time_stamp', 'id', 'heart_rate', 'hand_temperature',
'hand_3D_acceleration_16_x', 'hand_3D_acceleration_16_y',
'hand_3D_acceleration_16_z', 'hand_3D_acceleration_6_x',
'hand_3D_acceleration_6_y', 'hand_3D_acceleration_6_z',
'hand_3D_gyroscope_x', 'hand_3D_gyroscope_y', 'hand_3D_gyroscope_z',
'hand_3D_magnetometer_x', 'hand_3D_magnetometer_y',
'hand_3D_magnetometer_z', 'hand_4D_orientation_x',
'hand_4D_orientation_y', 'hand_4D_orientation_z',
'hand_4D_orientation_w', 'chest_temperature',
'chest_3D_acceleration_16_x', 'chest_3D_acceleration_16_y',
'chest_3D_acceleration_16_z', 'chest_3D_acceleration_6_x',
'chest_3D_acceleration_6_y', 'chest_3D_acceleration_6_z',
'chest_3D_gyroscope_x', 'chest_3D_gyroscope_y', 'chest_3D_gyroscope_z',
'chest_3D_magnetometer_x', 'chest_3D_magnetometer_y',
'chest_3D_magnetometer_z', 'chest_4D_orientation_x',
'chest_4D_orientation_y', 'chest_4D_orientation_z',
'chest_4D_orientation_w', 'ankle_temperature',
'ankle_3D_acceleration_16_x', 'ankle_3D_acceleration_16_y',
'ankle_3D_acceleration_16_z', 'ankle_3D_acceleration_6_x',
'ankle_3D_acceleration_6_y', 'ankle_3D_acceleration_6_z',
'ankle_3D_gyroscope_x', 'ankle_3D_gyroscope_y', 'ankle_3D_gyroscope_z',
'ankle_3D_magnetometer_x', 'ankle_3D_magnetometer_y',
'ankle_3D_magnetometer_z', 'ankle_4D_orientation_x',
'ankle_4D_orientation_y', 'ankle_4D_orientation_z',
'ankle_4D_orientation_w']
# ### Create sliding windows
# +
from window_slider import Slider
def make_windows(df, bucket_size, overlap_count):
window_list = []
final = pd.DataFrame()
activity_list = list(df['activity_id'].unique()) #list of the four activities
sub_id_list = list(df['id'].unique()) #list of the subject ids
df_list = []
for i in sub_id_list:
df_subject = df[df['id'] == i] #isolate a single subject id
for j in activity_list:
df_subject_activity_round = df_subject[df_subject['activity_id'] == j] #isolate by activity
final_df = pd.DataFrame()
if df_subject_activity_round.empty:
pass
else:
df_flat = df_subject_activity_round[no_act].T.values #array of arrays, each row is every single reading in an array for a sensor in that isolation
slider = Slider(bucket_size,overlap_count)
#print(i, j)
slider.fit(df_flat)
while True:
window_data = slider.slide()
if slider.reached_end_of_list(): break
window_list.append(list(window_data))
final_df = final.append(window_list)
final_df.columns = [no_act]
final_df.insert(53, "id_window", [i]*len(final_df), True)
final_df.insert(54, "activity_id", [j]*len(final_df), True)
df_list.append(final_df)
window_list = []
final = pd.DataFrame(columns = df_list[0].columns)
for l in df_list:
final = final.append(l)
final
final.columns = final.columns.map(''.join)
return final
# -
final = make_windows(df2, 500, 100)
# ### Number windows
final = final.assign(count=final.groupby(final.activity_id.ne(final.activity_id.shift()).cumsum()).cumcount().add(1))
final
final['count'] = final['count'].apply(lambda x: np.full(500, x))
# ### Create test/train split
# +
ID_list = df['id'].unique()
random.shuffle(ID_list)
train = pd.DataFrame()
test = pd.DataFrame()
#change size of train/test split
train = final[final['id_window'].isin(ID_list[:6])]
test = final[final['id_window'].isin(ID_list[6:])]
print(train.shape, test.shape)
# -
no_act.append('count')
id_list = train['id_window'].values
# +
X_train = train[no_act] #get only sensor values and subject ID
X_train = X_train.apply(pd.Series.explode).reset_index() #break windows into rows
X_test = test[no_act]
X_test = X_test.apply(pd.Series.explode).reset_index()
X_test = X_test.drop(['index', 'time_stamp'], axis = 1)
X_train = X_train.drop(['index', 'time_stamp'], axis = 1)
print(X_train.shape, X_test.shape)
train_shape = len(X_train)
# -
# ### Extract labels
y_train = train['activity_id'].values
y_test = test['activity_id'].values
X_train.shape
# ### Make dummy variables for subject ID
# +
X_train['train'] = 1
X_test['train'] = 0
combined = pd.concat([X_train, X_test])
combined = pd.concat([combined, pd.get_dummies(combined['id'])], axis =1)
combined = pd.concat([combined, pd.get_dummies(combined['count'])], axis =1)
# +
X_train = combined[combined['train'] == 1]
X_test = combined[combined['train'] == 0]
X_train.drop(["train", 'count'], axis = 1, inplace = True)
X_test.drop(["train", 'count'], axis = 1, inplace = True)
print(X_train.shape, X_test.shape, X_train.shape[0] + X_test.shape[0])
# -
X_train_df = combined[combined['train'] == 1].drop(["train", "id", 'count'], axis = 1, inplace = True)
X_train_df
# ### Scale/normalize features
# Scaling is used to change values without distorting differences in the range of values for each sensor. We do this because different sensor values are not in similar ranges of each other and if we did not scale the data, gradients may oscillate back and forth and take a long time before finding the local minimum. It may not be necessary for this data, but to be sure, we normalized the features.
#
# The standard score of a sample x is calculated as:
#
# $$z = \frac{x-u}{s}$$
#
# Where u is the mean of the data, and s is the standard deviation of the data of a single sample. The scaling is fit on the training set and applied to both the training and test set.
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
ss = StandardScaler()
X_train.iloc[:,1:53] = ss.fit_transform(X_train.iloc[:,1:53])
X_test.iloc[:,1:53] = ss.transform(X_test.iloc[:,1:53])
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# ### Encode y labels
le = LabelEncoder()
y_train = le.fit_transform(y_train)
y_test = le.transform(y_test)
y_train_dummy = np_utils.to_categorical(y_train)
y_test_dummy = np_utils.to_categorical(y_test)
activity_name_mapping = dict(zip(le.classes_, le.transform(le.classes_)))
print(activity_name_mapping)
# ### Reshaping windows as arrays
# Convert to transposed arrays
X_test = X_test.T.values
X_train = X_train.T.values
# +
X_test = X_test.astype('float64')
X_train = X_train.astype('float64')
# Reshape to -1, window_size, # features
X_train = X_train.reshape((-1, 500, X_train.shape[0]))
X_test = X_test.reshape((-1, 500, X_test.shape[0]))
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# -
# ### Neural Network
# #### Architecture:
# - 2 **Convolutional layers** that support 1D vectors as input. The first has a kernel (represents the amount of values as input to each convolution) of 3, and the second has 2
#
# - 1 **Pooling layer** that aggregates the convolutional layers by taking the maximum values of two consecutive convolutions.
#
# - 2 **Fully Connected layers**. The first has 100 nodes, and the second has 25 (the number of classes possible).
#
# - **Softmax** acitvation function - Used to generate probabilities for each class as an output in the final fully connected layer of the model
# +
from sklearn.model_selection import LeaveOneGroupOut
fold_no = 1
# Lists to store metrics
acc_per_fold = []
loss_per_fold = []
# Define the K-fold Cross Validator
groups = id_list
inputs = X_train
targets = y_train_dummy
logo = LeaveOneGroupOut()
logo.get_n_splits(inputs, targets, groups)
cv = logo.split(inputs, targets, groups)
for train, test in cv:
#Define the model architecture
model = Sequential()
model.add(Conv1D(filters=64, kernel_size=3, activation='relu'))
model.add(Conv1D(filters=64, kernel_size=2, activation='relu'))
model.add(MaxPooling1D(pool_size=2))
model.add(Flatten())
model.add(Dense(100, activation='relu'))
model.add(Dense(3, activation='softmax')) #3 outputs are possibl
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# Generate a print
print('------------------------------------------------------------------------')
print(f'Training for fold {fold_no} ...')
# Fit data to model
history = model.fit(inputs[train], targets[train],
batch_size=32,
epochs=10,
verbose=1)
# Generate generalization metrics
scores = model.evaluate(inputs[test], targets[test], verbose=0)
print(f'Score for fold {fold_no}: {model.metrics_names[0]} of {scores[0]}; {model.metrics_names[1]} of {scores[1]*100}%')
acc_per_fold.append(scores[1] * 100)
loss_per_fold.append(scores[0])
# Increase fold number
fold_no = fold_no + 1
# == Provide average scores ==
print('------------------------------------------------------------------------')
print('Score per fold')
for i in range(0, len(acc_per_fold)):
print('------------------------------------------------------------------------')
print(f'> Fold {i+1} - Loss: {loss_per_fold[i]} - Accuracy: {acc_per_fold[i]}%')
print('------------------------------------------------------------------------')
print('Average scores for all folds:')
print(f'> Accuracy: {np.mean(acc_per_fold)} (+- {np.std(acc_per_fold)})')
print(f'> Loss: {np.mean(loss_per_fold)}')
print('------------------------------------------------------------------------')
# -
# Model checkpoint is used to save weights of the best model. In the code below, we set the best model to be defined as the model with the least validation loss.
from keras.callbacks import ModelCheckpoint
import datetime
model_checkpoint = ModelCheckpoint('./models/HARnet.hdf5', monitor='val_loss', verbose=1, save_best_only=True)
log_dir = "logs/fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1)
# We decided to use ADAM as our optimizer as it is computationally efficient and updates the learning rate on a per-parameter basis, based on a moving estimate per-parameter gradient, and the per-parameter squared gradient.
model.summary()
model1.save("./model/TF_RNN.hdf5")
# ### Predict on test data and print accuracy/F1 scores
accuracy = model.evaluate(X_test, y_test_dummy, batch_size = 32, verbose = 1)
print(accuracy)
# Argmax is used to select the output class with the highest probability in the output as these are the prediction labels for our test data.
y_pred = np.argmax(model.predict(X_test), axis=-1)
# A **confusion matrix** is generated to observe where the model is classifying well and to see classes which the model is not classifying well. Check the label encodings to see which classes match to which encoding - they go in numerical order from up/left to down/right.
from sklearn.metrics import confusion_matrix, accuracy_score, f1_score
confusion_matrix(y_pred, y_test)
cm = confusion_matrix(y_pred, y_test)
cm = cm/cm.astype(np.float).sum(axis=1)
# +
import seaborn as sns
from matplotlib import pyplot as plt
ax = plt.subplot()
sns.heatmap(cm, annot = True, fmt = '.2f',cmap = 'Blues', xticklabels = ['Lying', 'Standing', 'Walking'], yticklabels = ['Lying', 'Standing', 'Walking'])
ax.set_xlabel("Predicted labels")
ax.set_ylabel('Actual labels')
plt.title('PAMAP2 Confusion Matrix')
plt.savefig('PAMAP2_ANN_conf_matrix.png')
# -
# The **accuracy** score represents the proportion of correct classifications over all classifications.
accuracy_score(y_pred, y_test)
# The **F1 score** is a composite metric of two other metrics:
#
# Specificity: proportion of correct 'positive predictions' over all 'positive' predictions.
#
# Sensitivity: number of correct 'negative' predictions over all 'negative' predictions.
#
# The F1 score gives insight as to whether all classes are predicted correctly at the same rate. A low F1 score and high accuracy can indicate that only a majority class is predicted.
f1_score(y_pred, y_test, average = 'weighted')
model.save_weights('./')
|
DigitalBiomarkers-HumanActivityRecognition/10_code/50_deep_learning/53_tensorflow_models/53_tensorFlow_PAMAP2/.ipynb_checkpoints/TF_CNN-PAMAP-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <center><img src="./images/logo_fmkn.png" width=300 style="display: inline-block;"></center>
#
# ## Машинное обучение
# ### Семинар 12. Введение в байесовские методы
#
# <br />
# <br />
# 2 декабря 2021
# ### Напоминание. <NAME>
#
# $$
# P(A | B) = \frac{P(B | A) P(A)}{\int P(B | A) P(A) dA} = \frac{P(B | A) P(A)}{P(B)}
# $$
#
# В машинном обучении это транслируется так:
# $$
# P(\theta | Y) = \frac{P(Y | \theta) P(\theta)}{\int P(Y | \theta) P(\theta)},
# $$
#
# где $P(Y | \theta)$ --- функция правдоподобия данных при заданных параметрах модели, $P(\theta)$ --- априорное распределение параметров, $P(Y) = \int P(Y | \theta) P(\theta)$ --- маргинальное правдоподобие данных.
#
# Общая схема применения байесовского подхода:
# 1. специфицировать модель (то есть задать функцию правдоподобия)
# 2. указать априорное распределение параметров. Cложная часть. Как правило, априорное распределение должно быть достаточно неинформативным, но при этом отражать какие-то _априорные_ знания о параметрах. Есть целый [документ](https://github.com/stan-dev/stan/wiki/Prior-Choice-Recommendations) с "рецептами" по выбору априорного распределения.
# 3. применить теорему Байеса :)
#
# На практике практически всегда вычисление апостериорного распределения оказывается невозможным.
# На помощь приходят либо различного рода аппроксимации, либо Markov Chain Monte Carlo.
# ### Сопряженные распределения. Пример
#
# Если апостериорное распределение $p(\theta | Y)$ для функции правдоподобия $p(y | \theta)$ из того же семейства распределений, что и априорное $p(\theta)$, такой prior называется сопряженным (conjugate) для правдоподобия $p(y | \theta)$. Такие распределения удобны для работы, потому что в этом случае апостериорное распределение считается аналитически.
#
# Примеры пар сопряженных распределений.
#
# | Likelihood | Prior |
# |------------|-------|
# | (Multivariate) Gaussian | (Multivariate) Gaussian |
# | Bernoulli | Beta |
# | Poisson | Gamma |
# | Gaussian | Gamma |
# Рассмотрим простейший пример с нечестной монеткой. Пусть монетка выпадает орлом с вероятностью $\theta$ (в случае $\theta = 0.5$ монетка честная). Результаты подкидывания монетки получаются из распределения $P(coin = head | \theta) = Bernoulli(\theta)$. Это будет нашей функцией правдоподобия.
#
# Как нам оценить параметр $\theta$ из данных?
# +
import numpy as np
import scipy.stats
import matplotlib.pyplot as plt
import seaborn as sns
import sklearn.model_selection
import sklearn.datasets
from scipy.special import expit
from sklearn.linear_model import LogisticRegression
SEED = 1234
np.random.seed(SEED) # for reproducibility
# -
# generate some data
theta = 0.7
coin_flips = scipy.stats.bernoulli.rvs(theta, size=10) # let's use small data
names = ['Head', 'Tails']
data = coin_flips.mean(), 1 - coin_flips.mean()
plt.bar(names, data)
# Простейший (фреквентистский) вариант --- получить максимум правдоподобия:
theta_mle = coin_flips.mean()
print('MLE for \\theta =', theta_mle)
# **Задание.** Что произошло? Докажите, что MLE-оценка для $\theta$ это выборочное среднее.
# Для правдоподобия Бернулли сопряженным априорным распределением будет бета-распределение $Beta(\alpha, \beta)$.
#
# Апостериорным распределением будет $Beta(\alpha + \sum x_i, \beta + n - \sum x_i)$.
#
# У нас нет оснований полагать что-то об $\alpha$ и $\beta$, поэтому выберем $\alpha=\beta=1$. Это распределение совпадает с равномерным распределением --- мы считаем, что $\theta$ может быть где угодно на отрезке $[0, 1]$.
a_prior = 1.
b_prior = 1.
beta_prior = scipy.stats.beta(a=a_prior, b=b_prior)
beta_posterior = scipy.stats.beta(a = a_prior + coin_flips.sum(),
b = b_prior + len(coin_flips) - coin_flips.sum())
# +
# let's plot
x_ = np.linspace(0, 1, 100)
plt.plot(x_, beta_prior.pdf(x_), label='Prior')
plt.plot(x_, beta_posterior.pdf(x_), label='Posterior')
plt.plot([theta_mle, theta_mle], [0, beta_posterior.pdf(theta_mle)], linestyle='--', alpha=0.9, label='MLE')
plt.grid()
plt.legend()
# -
# Давайте посмотрим, что будет в случае большего количества данных.
# +
coin_flips = scipy.stats.bernoulli.rvs(theta, size=1000) # let's use big data (ok, not so big)
beta_posterior = scipy.stats.beta(a = 1. + coin_flips.sum(), b = 1. + len(coin_flips) - coin_flips.sum())
theta_mle = coin_flips.mean()
x_ = np.linspace(0, 1, 1000)
plt.plot(x_, beta_prior.pdf(x_), label='Prior')
plt.plot(x_, beta_posterior.pdf(x_), label='Posterior')
plt.plot([theta_mle, theta_mle], [0, beta_posterior.pdf(theta_mle)], linestyle='--', alpha=0.9, label='MLE')
plt.grid()
plt.legend()
# -
# **Задание**. Попробуйте поменять параметры априорного распределение на другие. Что будет в этом случае c MLE-оценкой? Что произойдет с увеличением количества данных?
# Этот пример слишком простой. Во-первых, пространство параметров у нас одномерно. Во-вторых, нам "повезло" и для функции правдоподобия есть сопряженное распределение.
#
# В многомерном случае (как правило), нет такой роскоши.
# ### Логистическая регрессия
#
# Логистическая регрессия моделирует вероятность, что классифицируемый объект окажется в положительном классе:
#
# $$ p(y_i | \theta) = Bernoulli(p_i) $$
# $$ p_i = p(y_i = 1| \theta) = \sigma(\theta_1 x_{i1} + \theta_2 x_{i2} + \ldots + \theta_m x_{im} )$$
#
# Это наша функция правдоподобия.
#
# (Напоминание: $\sigma(t) = 1 / (1 + \exp(-t))$)
#
# Для набора данных $Y = [y_1, \ldots, y_N]^\top$ мы будем предполагать условную независимость:
#
# $$
# P(Y | \theta) = \prod_{i=1}^{N} p(y_i | \theta)
# $$
#
# Для такой функции правдоподобия нет сопряженного распределения :( Поэтому нам придется применять другие методы для нахождения апостериорного.
# +
# load some data
raw_data = sklearn.datasets.load_breast_cancer()
X = raw_data['data'][:, [0, 4]]
feature_names = raw_data['feature_names'][[0, 4]]
Y = raw_data['target']
target_names = raw_data['target_names']
# normalize the data
X_mean, X_std = X.mean(axis=0), X.std(axis=0)
X_std = np.where(np.isclose(X_std, 0), 1., X_std)
X_normed = (X - X_mean) / X_std
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X_normed, Y)
print('Predicting breast cancer based on %s' % ', '.join(feature_names))
# -
# Посмотрим на датасет:
def plot_classes(X, Y, feature_names, target_names, title):
colors = ['red', 'green']
for i, cancer_type in enumerate(target_names):
where = np.where(Y == i)
plt.scatter(X[where, 0], X[where, 1], c=colors[i], label=cancer_type)
plt.title(title)
plt.xlabel(feature_names[0])
plt.ylabel(feature_names[1])
plt.legend(loc='best')
plot_classes(X_train, Y_train, feature_names, target_names, title='Cancer')
# Давайте выберем априорное распределение для наших параметров $\theta$. Для простоты возьмем нормальное распределение с большой дисперсией:
theta_prior = scipy.stats.multivariate_normal(mean = np.zeros(2), cov=10*np.eye(2))
# Лог-правдоподобие данных:
def log_likelihood(theta):
dot_product = theta.dot(X_train.T)
sign = -np.sign(Y_train - 0.5) # -1 if Y==1, +1 if Y==0
log_likelihood = -np.log(1.0 + np.exp(sign * dot_product)).sum(axis=-1)
return log_likelihood
# Ненормированное апостериорное распределение:
# $$ p(\theta | Y) \propto p(Y | \theta) p(\theta) $$
#
# Нам удобнее считать логарифм:
# $$ \log p(\theta | Y) = \log p(Y | \theta) + \log p(\theta) + C $$
def log_unnormed_posterior(theta):
return log_likelihood(theta) + theta_prior.logpdf(theta)
# +
theta_1, theta_2 = np.meshgrid(np.linspace(-6., -3., 100), np.linspace(-2, -.5, 100)) # (100, 100)
theta = np.dstack((theta_1, theta_2)) # (100, 100, 2)
plt.contourf(theta_1, theta_2, np.exp(log_unnormed_posterior(theta)), levels=10)
plt.colorbar()
plt.title('Unnormed posterior')
plt.xlabel('$\\theta_1$')
plt.ylabel('$\\theta_2$')
# -
# ### MCMC
# #### Метрополис-Хастингс
#
# Напомним схему алгоритма Метрополиса-Хастингса.
# 0. Выберем произвольную точку $x_0$ и симметричное распределение $Q(x_{n+1} | x_n)$ (proposal rule)
#
# На каждой итерации:
# 1. Сгенерируем точку $x_{n+1}$ из распределения $Q(x_{n+1} | x_n)$
# 2. Посчитаем acceptance ratio
# $$ \alpha = \frac{p(x_{n+1})}{p(x_n)} $$
# 3. Сгенерируем случайное $u \sim U[0, 1]$.
# 4. Примем новый сэмпл, если $u \leq \alpha$
# +
x_0 = np.zeros(2) # why not?
x_current = x_0
n_accepts = 0
n_iter = 10000
metropolis_samples = []
for _ in range(n_iter):
# write your code here
# -
print('Efficiency: %.2f' % (n_accepts / n_iter))
# Как правило, из полученных сэмплов выкидывают первые, начальные сэмплы, полученные, когда марковская цепь еще не сошлась к стационарному распределению. Этот период называют burn in или warp up (прогрев). Кроме того, соседние сэмплы оказываются коррелированными, поэтому обычно их "разреживают", беря не все сэмплы, а только каждый N-ый.
#
# Для простоты мы зададим какие-то относительно разумные значения для прогрева и разреживания.
# +
burnin = 1000
sample_sparsifier = 5
theta_samples = np.array(metropolis_samples[burnin:][::sample_sparsifier])
# -
sns.jointplot(x=theta_samples[:,0], y=theta_samples[:,1], kind='kde')
# Довольно неплохо приблизили апостериорное распределение. Можно видеть, что апостериорное распределение параметров довольно широкое --- в случае "точечной" оценки эта информация теряется.
# ### Предсказательное распределение
#
# Окей, но как нам предсказывать класс для _новых_ пациентов?
#
# Посмотрим на апостериорное распределение для $y_*$:
# $$ p(y_* | Y) = \frac{p(y_*, Y)}{p(Y)} = \int \frac{p(y_*, Y, \theta)}{p(Y)} d\theta = \int \frac{p(y_* | Y, \theta) p(Y, \theta)}{p(Y)} d\theta = \int p(y_* | Y, \theta) p(\theta | Y) d\theta = \int p(y_* | \theta) p(\theta | Y) d\theta $$
#
# В последнем равенстве мы воспользовались условной независимостью исходов.
#
# Это можно интепретировать так: мы _усредняем_ предсказания $p(y_* | \theta)$ по _всем_ возможным моделям $\theta$, взвешивая их настолько, насколько они вероятны при имеющихся данных.
#
# Как обычно, этот интеграл не вычислим :) Мы можем воспользоваться методом Монте-Карло:
# 1. Сэмплируем $\theta_i$ из распределения $p(\theta | Y)$.
# 2. Сэмплируем $y_*$ из распределения $p(y_* | \theta_i)$.
#
# Таким образом, получим сэмплы из распределения $p(y_* | Y)$.
#
# Давайте посмотрим на тестовую выборку:
def predict(X):
prob = expit(theta_samples.dot(X.T)) # logistic function
rv = scipy.stats.bernoulli.rvs(prob) # samples from predictive distribution
means = np.mean(rv, axis=0)
variances = np.var(rv, axis=0)
return means, variances
means, variances = predict(X_test)
# +
plt.figure(figsize=(15, 5))
plt.subplot(131)
plot_classes(X_test, Y_test, feature_names, target_names, title='Truth')
plt.subplot(132)
plt.scatter(X_test[..., 0], X_test[..., 1], c=means, cmap='RdYlGn')
plt.title("Predictive mean")
plt.colorbar()
plt.subplot(133)
plt.scatter(X_test[..., 0], X_test[..., 1], c=variances, cmap='viridis')
plt.title("Predictive variance")
plt.colorbar()
# -
# Как видим, на границе классов среднее наших предсказаний близко к 0.5, а дисперсия выше, чем в глубине классов. Это демонстрирует неуверенность (неопределенность) модели --- действительно, классы перемешаны, и сделать уверенное предсказание на границе сложно в рамках нашей модели.
# ### Фреймворки
#
# Мы написали много бойлерплейт-кода, но для Python есть несколько фреймворков для байесовского вывода, упрощающих жизнь.
#
# * **PyMC3**
#
# Библиотека на основе `theano` (эффективно мертвом фреймворке). Тем не менее, хорошая документация, много примеров и туториалов. А также очень простая визуализация.
#
# * **Bambi**
#
# Надстройка над PyMC3 для упрощения реализации Generalized Linear Models.
#
# * **Pyro**
#
# Библиотека на основе `pytorch`.
#
# * **TensorFlow Probability**
#
# Модуль для `tensorflow`.
# Давайте попробуем вывести апостериорные распределения для параметров логистической регрессии по _всем_ признакам. Воспользуемся `pyro`.
import torch
import pyro
import pyro.distributions as dist
import pyro.infer.mcmc as mcmc
from pyro.infer import Predictive
# Подготовим данные
# +
X = raw_data['data']
feature_names = raw_data['feature_names']
Y = raw_data['target']
target_names = raw_data['target_names']
# normalize the data
X_mean, X_std = X.mean(axis=0), X.std(axis=0)
X_std = np.where(np.isclose(X_std, 0.), 1., X_std)
X_normed = (X - X_mean) / X_std
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X_normed, Y)
# convert to torch
X_train, X_test, Y_train, Y_test = torch.tensor(X_train).float(), torch.tensor(X_test).float(), \
torch.tensor(Y_train).float(), torch.tensor(Y_test).float()
n_features = X.shape[-1]
# -
# Заведем модель логистической регрессии. В качестве априорных распределений снова возьмем "широкие" гауссианы.
# +
burnin = 1000
n_samples = 9000
def logistic_regression(x, y):
intercept = pyro.sample("intercept", dist.Normal(torch.tensor([0.0]), torch.tensor([10.0])))
theta = pyro.sample("theta", dist.Normal(torch.zeros(n_features), 10.0*torch.ones(n_features)))
linear = intercept + torch.matmul(theta, x)
obs = pyro.sample("obs", dist.Bernoulli(logits = linear), obs=y)
return obs
# -
# Используем более "продвинутый" метод MCMC, который называется NUTS (No U-Turn Sampler).
#
# NUTS использует градиент, чтобы более эффективно исследовать апостериорное распределение. (То есть proposal rule более сложное, чем просто гауссиана).
#
# В детали NUTS вдаваться не будем, поскольку это выходит за рамки курса.
#
# **Warning:** следующая клетка может выполнятся долго.
# +
kernel = mcmc.NUTS(logistic_regression, jit_compile=False)
posterior = mcmc.MCMC(kernel,
num_samples=n_samples,
warmup_steps=burnin,
num_chains=1,
disable_progbar=False)
posterior.run(X_train.T, Y_train)
# -
# Получим сэмплы из апостериорного распределения
samples = posterior.get_samples()
samples['theta'] = samples['theta'][::sample_sparsifier]
samples['intercept'] = samples['intercept'][::sample_sparsifier]
# Давайте посмотрим на маргинальные распределения сэмплов.
theta_samples = samples['theta'].numpy()
# +
n_rows = 5
n_cols = 6
fig, axes = plt.subplots(n_rows, n_cols, figsize=(18, 15), sharey=True, sharex=True)
for i in range(theta_samples.shape[-1]):
ax = plt.subplot(n_rows, n_cols, i+1)
sns.kdeplot(ax=ax, x=theta_samples[:, i])
ax.set_title('$\\theta_{%d}$' % (i+1))
# -
sns.displot(x=theta_samples[:, 0], y=theta_samples[:, 1], kind='kde')
# **Вопрос**: что можно сказать о распределениях параметров? Какие параметры оценились уверенно, а в каких есть большая степень неопределенности?
# Попробуем что-нибудь предсказать.
predictive = Predictive(logistic_regression, samples)(X_test.T, None)
means = predictive['obs'].mean(axis=0)
variance = predictive['obs'].var(axis=0)
# +
plt.figure(figsize=(15, 5))
plt.subplot(131)
plot_classes(X_test[:, [0, 4]], Y_test, feature_names, target_names, title='Truth')
plt.subplot(132)
plt.scatter(X_test[:, 0], X_test[:, 4], c=means, cmap='RdYlGn')
plt.title("Predictive mean")
plt.colorbar()
plt.subplot(133)
plt.scatter(X_test[:, 0], X_test[:, 4], c=variances, cmap='viridis')
plt.title("Predictive variance")
plt.colorbar()
# -
# ### Материалы
#
# Интерактивная визуализация различных алгоритмов MCMC: https://chi-feng.github.io/mcmc-demo/
#
# Концептуальное введение в MCMC: https://arxiv.org/abs/1909.12313
# ### Это еще не всё
#
# Байесовский взгляд учитывает неопределенность, присущую задачам машинного обучения. Байесовские модели выдают не фиксированные ответы, а распределение ответов.
#
# MCMC -- один из методов получения апостериорного распределения. Его главный недостаток -- высокая вычислительная сложность. Помимо этого, есть и другие способы получать (аппроксимированно) апостериорные распределения (такие как вариационный вывод).
#
# Кроме того, как вы помните, есть "особое" правдоподобие Multivariate Gaussian, для которого сопряженным априорным будет Multivariate Gaussian. Это приводит к еще одной мощной байесовской модели гауссовских процессов. Но это уже отдельная история для следующего семестра :)
|
2021-fall-part-1/seminars/12_bayes/12_bayes_practice.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Plotting uncertainty
# In this example we will go over plotting uncertainties in various ways:
# + y errorbars
# + x errorbars
# + x and y errorbars (no covariance)
# + x and y error-ellipse (covariance)
#
# ## Packages being used
# + `matplotlib`: all the plotting
# + `astropy`: read in the data table
# + `numpy` and `scipy`: convert cov matrix to ellipse params
#
# ## Relevant documentation
# + `matplotlib`: http://matplotlib.org/2.0.2/api/pyplot_api.html#matplotlib.pyplot.errorbar
from astropy.table import Table
import scipy.linalg as sl
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.patches import Ellipse
import mpl_style
# %matplotlib notebook
plt.style.use(mpl_style.style1)
# Our data contains $(x, y)$ positions with 1-$\sigma$ uncertainties and covariance values:
t = Table.read('data.csv', format='ascii.csv')
print(t)
# **Note** the full covariance matrix for each data point is:
# $\left[ \begin{array}{ccc} \sigma_x^2 & \rho_{xy}\sigma_x \sigma_y \\ \rho_{xy}\sigma_x \sigma_y & \sigma_y^2 \end{array} \right]$
#
# ## y-uncertanties or x-uncertanties only
# The most common type of data you will work with will only have (significant) uncertainties in one direction. In this case it is very easy to plot using `errorbar`:
plt.figure(1)
plt.errorbar(
t['x'],
t['y'],
yerr=t['sy'],
ls='None',
mfc='k',
mec='k',
ms=5,
marker='s',
ecolor='k'
)
plt.xlabel('x')
plt.ylabel('y')
plt.ylim(0, 700)
plt.figure(2)
plt.errorbar(
t['x'],
t['y'],
xerr=t['sx'],
ls='None',
mfc='k',
mec='k',
ms=5,
marker='s',
ecolor='k'
)
plt.xlabel('x')
plt.ylabel('y')
plt.ylim(0, 700)
# ## Uncertainties in both x and y with no cov
# If your data has no cov you can still use `errorbar`:
plt.figure(3)
plt.errorbar(
t['x'],
t['y'],
yerr=t['sy'],
xerr=t['sx'],
ls='None',
mfc='k',
mec='k',
ms=5,
marker='s',
ecolor='k'
)
plt.xlabel('x')
plt.ylabel('y')
plt.ylim(0, 700)
# ## Uncertainties in both x and y with cov
# If your data does have cov you should plot a 1-$\sigma$ ellipse around each point. There is no built in function to do this, so we will have to write our own. We will start by writing a function to turn a cov matrix into the parameters for an ellipse and draw it on a figure.
# +
def cov_to_ellipse(cov, pos, **kwargs):
eigvec,eigval,V = sl.svd(cov,full_matrices=False)
# the angle the first eigenvector makes with the x-axis
theta = np.degrees(np.arctan2(eigvec[1, 0], eigvec[0, 0]))
# full width and height of ellipse, not radius
# the eigenvalues are the variance along the eigenvectors
width, height = 2 * np.sqrt(eigval)
return Ellipse(xy=pos, width=width, height=height, angle=theta, **kwargs)
def plot_ellipse(t, ax=None, **kwargs):
if ax is None:
ax = plt.gca()
for row in t:
cov = np.array(
[[row['sx']**2, row['pxy'] * row['sx'] * row['sy']],
[row['pxy'] * row['sx'] * row['sy'], row['sy']**2]]
)
ellip = cov_to_ellipse(cov, [row['x'], row['y']], **kwargs)
ax.add_artist(ellip)
plt.figure(4)
plt.plot(
t['x'],
t['y'],
's',
mfc='k',
mec='k',
ms=5
)
plot_ellipse(
t,
lw=1.5,
fc='none',
ec='C0'
)
plt.xlabel('x')
plt.ylabel('y')
plt.ylim(0, 700)
plt.draw()
# -
|
Uncertainty_plotting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="I08sFJYCxR0Z"
# 
# + [markdown] id="FwJ-P56kq6FU"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/tutorials/Certification_Trainings/Healthcare/4.6.Clinical_Deidentification_in_Italian.ipynb)
# + [markdown] id="Z7X1prqVxdB2"
# # Clinical Deidentification in Italian
#
# **Protected Health Information**:
#
# Individual’s past, present, or future physical or mental health or condition
# provision of health care to the individual
# past, present, or future payment for the health care
# Protected health information includes many common identifiers (e.g., name, address, birth date, Social Security Number) when they can be associated with the health information.
# + id="RWh6i1PtvE77"
import json
import os
from google.colab import files
if 'spark_jsl.json' not in os.listdir():
license_keys = files.upload()
os.rename(list(license_keys.keys())[0], 'spark_jsl.json')
with open('spark_jsl.json') as f:
license_keys = json.load(f)
# Defining license key-value pairs as local variables
locals().update(license_keys)
os.environ.update(license_keys)
# + id="MfOGbhC2wTyp"
# Installing pyspark and spark-nlp
# ! pip install --upgrade -q pyspark==3.1.2
# Installing Spark NLP Healthcare
# ! pip install --upgrade -q spark-nlp-jsl==$JSL_VERSION --extra-index-url https://pypi.johnsnowlabs.com/$SECRET
# + id="nou6cgDm35Vq" colab={"base_uri": "https://localhost:8080/", "height": 254} outputId="f1a31923-5dba-4a34-cb93-54cc0a6de6dd" executionInfo={"status": "ok", "timestamp": 1649533697923, "user_tz": -180, "elapsed": 24040, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
import sys
import os
import json
import pandas as pd
import string
import numpy as np
import sparknlp
import sparknlp_jsl
from pyspark.ml import Pipeline, PipelineModel
from pyspark.sql import functions as F
from pyspark.sql import SparkSession
from sparknlp.base import *
from sparknlp.annotator import *
from sparknlp.pretrained import ResourceDownloader
from sparknlp.util import *
from sparknlp_jsl.annotator import *
params = {"spark.driver.memory":"16G",
"spark.kryoserializer.buffer.max":"2000M",
"spark.driver.maxResultSize":"2000M"}
spark = sparknlp_jsl.start(SECRET, params=params)
print ("Spark NLP Version :", sparknlp.version())
print ("Spark NLP_JSL Version :", sparknlp_jsl.version())
spark
# + [markdown] id="VAyEoiHVuhbp"
# # 1. Italian NER Deidentification Models
# We have two different models you can use:
# * `ner_deid_generic`, detects 8 entities
# * `ner_deid_subentity`, detects 19 entities
# + [markdown] id="V1eksdJQoF4e"
# ### Creating pipeline
# + colab={"base_uri": "https://localhost:8080/"} id="snKDCdXwoNy4" outputId="40a458af-1b00-4ad3-ae03-0b4e51634040" executionInfo={"status": "ok", "timestamp": 1649533855707, "user_tz": -180, "elapsed": 95095, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
documentAssembler = DocumentAssembler()\
.setInputCol("text")\
.setOutputCol("document")
sentencerDL = SentenceDetectorDLModel.pretrained("sentence_detector_dl", "xx") \
.setInputCols(["document"])\
.setOutputCol("sentence")
tokenizer = Tokenizer()\
.setInputCols(["sentence"])\
.setOutputCol("token")
word_embeddings = WordEmbeddingsModel.pretrained("w2v_cc_300d", "it")\
.setInputCols(["document","token"])\
.setOutputCol("embeddings")
# + [markdown] id="3ZuJrSX6tT9m"
# ## 1.1. NER Deid Generic
#
# **`ner_deid_generic`** extracts:
# - Name
# - Profession
# - Age
# - Date
# - Contact (Telephone numbers, Email addresses)
# - Location (Address, City, Postal code, Hospital Name, Organization)
# - ID (Social Security numbers, Medical record numbers)
# - Sex
# + colab={"base_uri": "https://localhost:8080/"} id="C-qUIvqpsjV0" outputId="250df55b-f2e4-449c-9ceb-e3efff6bafc7" executionInfo={"status": "ok", "timestamp": 1649533869336, "user_tz": -180, "elapsed": 8709, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
ner_generic = MedicalNerModel.pretrained("ner_deid_generic", "it", "clinical/models")\
.setInputCols(["sentence","token","embeddings"])\
.setOutputCol("ner_deid_generic")
ner_converter_generic = NerConverter()\
.setInputCols(["sentence","token","ner_deid_generic"])\
.setOutputCol("ner_chunk_generic")
# + colab={"base_uri": "https://localhost:8080/"} id="ElhQ25TPwlYF" outputId="90f32379-6a40-4368-82d5-19c454975199" executionInfo={"status": "ok", "timestamp": 1649533869336, "user_tz": -180, "elapsed": 15, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
ner_generic.getClasses()
# + [markdown] id="3NjkLF70tYdZ"
# ## 1.2. NER Deid Subentity
#
# **`ner_deid_subentity`** extracts:
#
# - Patient
# - Doctor
# - Hospital
# - Date
# - Organization
# - City
# - Street
# - Username
# - Profession
# - Phone
# - Country
# - Age
# - Sex
# - Email
# - ZIP
# - Medical Record Number
# - Social Security Number
# - ID Number
# - URL
# + colab={"base_uri": "https://localhost:8080/"} id="C8n-h6D9tJXx" outputId="c5334c6b-68a6-419f-a76a-038d5a988b2c" executionInfo={"status": "ok", "timestamp": 1649533875318, "user_tz": -180, "elapsed": 5994, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
ner_subentity = MedicalNerModel.pretrained("ner_deid_subentity", "it", "clinical/models")\
.setInputCols(["sentence","token","embeddings"])\
.setOutputCol("ner_deid_subentity")
ner_converter_subentity = NerConverter()\
.setInputCols(["sentence", "token", "ner_deid_subentity"])\
.setOutputCol("ner_chunk_subentity")
# + colab={"base_uri": "https://localhost:8080/"} id="Oda9sjHDxRyi" outputId="d051fcee-2190-4e75-c80d-9f774cf3fbf1"
ner_subentity.getClasses()
# + [markdown] id="Eqg29dPuvl5f"
# ## 1.3. Pipeline
# + id="P0HZXLF6ueWi" executionInfo={"status": "ok", "timestamp": 1649533880854, "user_tz": -180, "elapsed": 3419, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentencerDL,
tokenizer,
word_embeddings,
ner_generic,
ner_converter_generic,
ner_subentity,
ner_converter_subentity,
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="fG4Vc36EwhFk" executionInfo={"status": "ok", "timestamp": 1649533882755, "user_tz": -180, "elapsed": 1904, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
text = "Ho visto <NAME> (49 anni), virologo, riferito all' Ospedale San Camillo per diabete mal controllato con sintomi risalenti a marzo 2015."
text_df = spark.createDataFrame([[text]]).toDF("text")
result = model.transform(text_df)
# + [markdown] id="MNUKLQMi0GjT"
# ### Results for `ner_generic`
# + colab={"base_uri": "https://localhost:8080/"} id="K2wDmdiFzwDb" outputId="b63718ba-63bd-424b-9868-6f3d3c11b189" executionInfo={"status": "ok", "timestamp": 1649533890170, "user_tz": -180, "elapsed": 7417, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
result.select(F.explode(F.arrays_zip('ner_chunk_generic.result', 'ner_chunk_generic.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + [markdown] id="y-CMPYTHz-L2"
# ### Results for `ner_subentity`
# + colab={"base_uri": "https://localhost:8080/"} id="vEtBcyIjzLA3" outputId="16e89cfb-0715-4bf0-d87c-310d9095eded" executionInfo={"status": "ok", "timestamp": 1649533891570, "user_tz": -180, "elapsed": 1404, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
result.select(F.explode(F.arrays_zip('ner_chunk_subentity.result', 'ner_chunk_subentity.metadata')).alias("cols")) \
.select(F.expr("cols['0']").alias("chunk"),
F.expr("cols['1']['entity']").alias("ner_label")).show(truncate=False)
# + [markdown] id="ZmClPVDfddNv"
# ## DeIdentification
# + [markdown] id="u-OTUyBK6yrt"
# ### Obfuscation mode
# + id="pbJisU_u7Kpl" executionInfo={"status": "ok", "timestamp": 1649533891571, "user_tz": -180, "elapsed": 4, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
# Downloading faker entity list.
# ! wget -q https://raw.githubusercontent.com/JohnSnowLabs/spark-nlp-workshop/master/tutorials/Certification_Trainings/Healthcare/data/obfuscate_it.txt
# + id="CBo2T-sZ64IJ" executionInfo={"status": "ok", "timestamp": 1649533891963, "user_tz": -180, "elapsed": 3, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
deid_masked_entity = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_entity")\
.setMode("mask")\
.setMaskingPolicy("entity_labels")
deid_masked_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_chars")\
.setMode("mask")\
.setMaskingPolicy("same_length_chars")
deid_masked_fixed_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_fixed_length_chars")\
.setMode("mask")\
.setMaskingPolicy("fixed_length_chars")\
.setFixedMaskLength(4)
deid_obfuscated = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"]) \
.setOutputCol("obfuscated") \
.setMode("obfuscate")\
.setObfuscateDate(True)\
.setObfuscateRefFile('obfuscate_it.txt')\
.setObfuscateRefSource("file")
# + id="h9pmXn0f75ST" executionInfo={"status": "ok", "timestamp": 1649533895263, "user_tz": -180, "elapsed": 865, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentencerDL,
tokenizer,
word_embeddings,
ner_subentity,
ner_converter_subentity,
deid_masked_entity,
deid_masked_char,
deid_masked_fixed_char,
deid_obfuscated
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="oVOL3bwr8J18" executionInfo={"status": "ok", "timestamp": 1649533896401, "user_tz": -180, "elapsed": 1, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
deid_lp = LightPipeline(model)
# + id="Maco1EiD8TK4" executionInfo={"status": "ok", "timestamp": 1649533896738, "user_tz": -180, "elapsed": 1, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
text = "Ho visto <NAME> (49 anni), virologo, riferito all' Ospedale San Camillo per diabete mal controllato con sintomi risalenti a marzo 2015."
# + colab={"base_uri": "https://localhost:8080/"} id="CXEmE1i78PX4" outputId="d22ab701-e6e0-4fcb-a34f-b4eeb7c1eb4e" executionInfo={"status": "ok", "timestamp": 1649533897882, "user_tz": -180, "elapsed": 359, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
result = deid_lp.annotate(text)
print("\n".join(result['masked_with_entity']))
print("\n")
print("\n".join(result['masked_with_chars']))
print("\n")
print("\n".join(result['masked_fixed_length_chars']))
print("\n")
print("\n".join(result['obfuscated']))
# + id="v8KakNhlMtnp" outputId="76cacbc3-7f21-4fd7-8a86-aaa9c07e5f89" colab={"base_uri": "https://localhost:8080/", "height": 211} executionInfo={"status": "ok", "timestamp": 1649533919172, "user_tz": -180, "elapsed": 355, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
pd.set_option("display.max_colwidth", 200)
df = pd.DataFrame(list(zip(result["masked_with_entity"],
result["masked_with_chars"],
result["masked_fixed_length_chars"],
result["obfuscated"])),
columns= ["Masked_with_entity", "Masked with Chars", "Masked with Fixed Chars", "Obfuscated"])
df
# + [markdown] id="DyA_2E7PMuLq"
# ### Faker mode
# + id="4WULhUB-MuLr" executionInfo={"status": "ok", "timestamp": 1649533922335, "user_tz": -180, "elapsed": 2, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
deid_masked_entity = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_entity")\
.setMode("mask")\
.setMaskingPolicy("entity_labels")
deid_masked_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_with_chars")\
.setMode("mask")\
.setMaskingPolicy("same_length_chars")
deid_masked_fixed_char = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"])\
.setOutputCol("masked_fixed_length_chars")\
.setMode("mask")\
.setMaskingPolicy("fixed_length_chars")\
.setFixedMaskLength(4)
deid_obfuscated = DeIdentification()\
.setInputCols(["sentence", "token", "ner_chunk_subentity"]) \
.setOutputCol("obfuscated") \
.setMode("obfuscate")\
.setLanguage('it')\
.setObfuscateDate(True)\
.setObfuscateRefSource('faker')
# + id="y0r2kTyGMuLr" executionInfo={"status": "ok", "timestamp": 1649533925189, "user_tz": -180, "elapsed": 1355, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
nlpPipeline = Pipeline(stages=[
documentAssembler,
sentencerDL,
tokenizer,
word_embeddings,
ner_subentity,
ner_converter_subentity,
deid_masked_entity,
deid_masked_char,
deid_masked_fixed_char,
deid_obfuscated
])
empty_data = spark.createDataFrame([[""]]).toDF("text")
model = nlpPipeline.fit(empty_data)
# + id="ayrfGKu8MuLr" executionInfo={"status": "ok", "timestamp": 1649533925681, "user_tz": -180, "elapsed": 495, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
deid_lp = LightPipeline(model)
# + id="OWJdfWjVMuLr" executionInfo={"status": "ok", "timestamp": 1649533925681, "user_tz": -180, "elapsed": 2, "user": {"displayName": "<NAME>", "userId": "08787989274818793476"}}
text = "Ho visto <NAME> (49 anni), virologo, riferito all' Ospedale San Camillo per diabete mal controllato con sintomi risalenti a marzo 2015."
# + colab={"base_uri": "https://localhost:8080/"} outputId="646a7c9b-a705-402c-b842-7ad95f2c9ff4" id="XdJmsPEsMuLr" executionInfo={"status": "ok", "timestamp": 1649533934070, "user_tz": -180, "elapsed": 7589, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
result = deid_lp.annotate(text)
print("\n".join(result['masked_with_entity']))
print("\n")
print("\n".join(result['masked_with_chars']))
print("\n")
print("\n".join(result['masked_fixed_length_chars']))
print("\n")
print("\n".join(result['obfuscated']))
# + id="_u6d1Q6aMvQr" outputId="bfd3d6a0-f3c7-4b93-faf1-abbf67a0d4c1" colab={"base_uri": "https://localhost:8080/", "height": 211} executionInfo={"status": "ok", "timestamp": 1649533936283, "user_tz": -180, "elapsed": 369, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
pd.set_option("display.max_colwidth", 200)
df = pd.DataFrame(list(zip(result["masked_with_entity"],
result["masked_with_chars"],
result["masked_fixed_length_chars"],
result["obfuscated"])),
columns= ["Masked_with_entity", "Masked with Chars", "Masked with Fixed Chars", "Obfuscated"])
df
# + [markdown] id="RRuq9NeemSaf"
# # 2. Pretrained Italian Deidentification Pipeline
#
# - We developed a clinical deidentification pretrained pipeline that can be used to deidentify PHI information from Italian medical texts. The PHI information will be masked and obfuscated in the resulting text.
# - The pipeline can mask and obfuscate:
# - Patient
# - Doctor
# - Hospital
# - Date
# - Organization
# - Sex
# - City
# - Street
# - Country
# - ZIP
# - Username
# - Profession
# - Phone
# - Email
# - Age
# - ID number
# - Medical record number
# - Account number
# - SSN
# - Plate Number
# - IP address
# - URL
# + colab={"base_uri": "https://localhost:8080/"} id="_vBWjdVTFGHD" outputId="54a89a96-d7d3-44e1-a4f3-542eb6fc5598" executionInfo={"status": "ok", "timestamp": 1649534027969, "user_tz": -180, "elapsed": 86347, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
from sparknlp.pretrained import PretrainedPipeline
deid_pipeline = PretrainedPipeline("clinical_deidentification", "it", "clinical/models")
# + colab={"base_uri": "https://localhost:8080/"} id="Esy1Yis8Hn1X" outputId="69a3925f-2033-4275-b7e8-2695c26b9546" executionInfo={"status": "ok", "timestamp": 1649534030980, "user_tz": -180, "elapsed": 3013, "user": {"displayName": "Monster C", "userId": "08787989274818793476"}}
text = """RAPPORTO DI RICOVERO
NOME: Lodovico Fibonacci
CODICE FISCALE: MVANSK92F09W408A
INDIRIZZO: <NAME> 7
CITTÀ : Napoli
CODICE POSTALE: 80139
DATA DI NASCITA: 03/03/1946
ETÀ: 70 anni
SESSO: M
EMAIL: <EMAIL>
DATA DI AMMISSIONE: 12/12/2016
DOTTORE: <NAME>
RAPPORTO CLINICO: 70 anni, pensionato, senza allergie farmacologiche note, che presenta la seguente storia: ex incidente sul lavoro con fratture vertebrali e costali; operato per la malattia di Dupuytren alla mano destra e un bypass ileo-femorale sinistro; diabete di tipo II, ipercolesterolemia e iperuricemia; alcolismo attivo, fuma 20 sigarette/giorno.
È stato indirizzato a noi perché ha presentato un'ematuria macroscopica post-evacuazione in un'occasione e una microematuria persistente in seguito, con un'evacuazione normale.
L'esame fisico ha mostrato buone condizioni generali, con addome e genitali normali; l'esame digitale rettale era coerente con un adenoma prostatico di grado I/IV.
L'analisi delle urine ha mostrato 4 globuli rossi/campo e 0-5 leucociti/campo; il resto del sedimento era normale.
L'emocromo è normale; la biochimica ha mostrato una glicemia di 169 mg/dl e trigliceridi 456 mg/dl; la funzione epatica e renale sono normali. PSA di 1,16 ng/ml.
INDIRIZZATO A: <NAME> - ASL Napoli 1 Centro, Dipartimento di Endocrinologia e Nutrizione - Stretto Scamarcio 320, 80138 Napoli
EMAIL: <EMAIL>
"""
result = deid_pipeline.annotate(text)
print("\n".join(result['masked_with_chars']))
print("\n")
print("\n".join(result['masked']))
print("\n")
print("\n".join(result['masked_fixed_length_chars']))
print("\n")
print("\n".join(result['obfuscated']))
# + [markdown] id="qRd-bpDenVsr"
# The results can also be inspected vertically by creating a Pandas dataframe as such:
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="YvTVq5JEmrc4" outputId="d5355c02-bcee-4f97-d9e1-35c784070455"
pd.set_option("display.max_colwidth", None)
df = pd.DataFrame(list(zip(result["sentence"],
result["masked"],
result["masked_with_chars"],
result["masked_fixed_length_chars"],
result["obfuscated"])),
columns= ["Sentence", "Masked", "Masked with Chars", "Masked with Fixed Chars", "Obfuscated"])
df
|
tutorials/Certification_Trainings/Healthcare/4.6.Clinical_Deidentification_in_Italian.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 12 - Advanced Pandas
# ## Advanced GroupBy Use
import pandas as pd
# Read from CSV file
df = pd.read_csv('dataset-A3-loans.csv')
df2 = df.copy()[['id', 'grade', 'funded_amount', 'interest_rate']]
df2 = df2[df2.grade.isin(['C', 'D', 'E', 'F'])]
_ = df2.reset_index(inplace=True, drop=True)
display(df2)
g = df2.groupby('grade')['funded_amount']
print(g)
g = df2.groupby('grade').funded_amount
g.mean()
df2['grade_mean'] = g.transform(lambda x : x.mean())
display(df2)
def normalise(x):
return (x - x.mean())/x.std()
display(df2.groupby('grade')['funded_amount'].transform(normalise))
display(df2.groupby('grade')['funded_amount'].apply(normalise))
# **References:**
#
# Python for Data Analysis, 2nd Edition, McKinney (2017)
|
12-2-Advanced-Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import json
import glob
# Start by importing the data and splitting it by lines
raw = []
with open("blaseball-log-season3.json", 'r') as f:
raw += f.read().split("\n")
if(raw[-1] == ""):
raw = raw[:-1]
json_lines = []
for line in raw:
json_lines.append(json.loads(line))
# How many lines did we get?
print(len(json_lines))
game_to_look_at = "ff1e878a-72d7-4162-bb10-3205fa4fa8d5"
games = {}
for line in json_lines:
for update in line["schedule"]:
if(update["_id"] == game_to_look_at):
#print((update["awayTeamBatterCount"], update["homeTeamBatterCount"]), update["lastUpdate"])
print(update["atBatBalls"], update["lastUpdate"])
|
Investigate single field.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# Nesse arquivo utilizamos uma abordagem mais complexa. Diante de cada caminho possuimos diversas variáveis
# (4 no total, cada variável representa um roteador no momento em que o usuario captura a informação).
#
# E fica a pergunta de:
#
# >Como podemos melhor utilizar esses dados?
#
# Aqui apresentamos uma estratégia de estrair informações de uma séries temporais através de [Wavelet](https://cran.r-project.org/web/packages/wavelets/wavelets.pdf).
# Simplementes tentamos extrair o máximo de informações que possam mostrar a similaridade ou disimilaridade de cada série. Para isso usamos a decomposição da série, extraindo mais detalhes da série.
#
# Aqui usamos o método **DWT**(Discrete Wavelet Transforms) com um filtro **haar** (são os filtro mais simples, básico degrais unitários, utilizados também em processamento de imagem como no algoritmo de viola jones para detecção de faces), mas como já havia dito, isso é apenas uma das diversas estratégias possíveis. Poderiamos usar **DFT**(acredito que **DFT** não é uma boa escolha desde que **DWT** trabalha melhor que **DFT**(Discrete Fourier Transform) em diversos artigos usando bases temporais (som, video, etc...) como é o nosso caso) ou outros filtros e até mesmo criar novos filtros.
# +
library(wavelets)
library(caret)
library(kernlab)
library(pROC)
print("Target")
groups <- read.csv(file="./MovementAAL/groups/MovementAAL_DatasetGroup.csv",head=TRUE,sep=",")
targetAll <- read.csv(file="./MovementAAL/dataset/MovementAAL_target.csv",head=TRUE,sep=",")
head(targetAll)
# +
#Group 1
allDataGroup1<-list()
allDataGroup1Target<-list()
groups1 = groups[groups$dataset_ID==1, ]
index<-1
for (id in groups1$X.sequence_ID){
caminho <-paste("./MovementAAL/dataset/MovementAAL_RSS_",id,".csv",sep="")
allDataGroup1[[index]]<-read.csv(file=caminho,head=TRUE,sep=",")
allDataGroup1Target[index]<-targetAll[[2]][id]
index<-index+1
}
wtData <- NULL
minStepsBack = 17
for (i in 1:length(allDataGroup1)){
aMatrix <- data.matrix(allDataGroup1[[i]], rownames.force = NA)
wt <- dwt(aMatrix[1:minStepsBack,], filter="haar", boundary="periodic")
wtData <- rbind(wtData, unlist(c(wt@W,wt@V[[wt@level]])))
}
wtData <- as.data.frame(wtData)
data = unlist(allDataGroup1Target)
target = factor(data,labels=c("No","Yes"))
frameDataFinal <- data.frame(cbind(target, wtData))
tail(frameDataFinal)
# -
# #### Média e Desvio padrão respectivamente.
# ##### Group 1, com Cross Validation tipo 10-fold
# +
inTraining <- createDataPartition(frameDataFinal$target, p = .7, list = TRUE,times=10)
allAccuracyGroup1 <- c()
for( i in 1:length(inTraining)){
training <- frameDataFinal[ inTraining[[i]],]
testing <- frameDataFinal[-inTraining[[i]],]
fitControl <- trainControl(method = "none", classProbs = TRUE)
svmLinearFit <- train(target ~ ., data = training,
method = "svmLinear",
trControl = fitControl,
family=binomial)
preds<- predict(svmLinearFit, newdata = testing)
matrix <- confusionMatrix(preds,frameDataFinal$target[-inTraining[[i]]])
allAccuracyGroup1 <- c(allAccuracyGroup1,matrix[3]$overall[[1]])
}
mean(allAccuracyGroup1)
sd(allAccuracyGroup1)
# +
#Group 2
allDataGroup2<-list()
allDataGroup2Target<-list()
groups2 = groups[groups$dataset_ID==2, ]
index<-1
for (id in groups2$X.sequence_ID){
caminho <-paste("./MovementAAL/dataset/MovementAAL_RSS_",id,".csv",sep="")
allDataGroup2[[index]]<-read.csv(file=caminho,head=TRUE,sep=",")
allDataGroup2Target[index]<-targetAll[[2]][id]
index<-index+1
}
wtData <- NULL
minStepsBack = 17
for (i in 1:length(allDataGroup2)){
aMatrix <- data.matrix(allDataGroup2[[i]], rownames.force = NA)
wt <- dwt(aMatrix[1:minStepsBack,], filter="haar", boundary="periodic")
wtData <- rbind(wtData, unlist(c(wt@W,wt@V[[wt@level]])))
}
wtData <- as.data.frame(wtData)
data = unlist(allDataGroup2Target)
target = factor(data,labels=c("No","Yes"))
frameDataFinal <- data.frame(cbind(target, wtData))
tail(frameDataFinal)
# -
# #### Média e Desvio padrão respectivamente.
# ##### Group 2, com Cross Validation tipo 10-fold
# +
inTraining <- createDataPartition(frameDataFinal$target, p = .7, list = TRUE,times=10)
allAccuracyGroup2 <- c()
for( i in 1:length(inTraining)){
training <- frameDataFinal[ inTraining[[i]],]
testing <- frameDataFinal[-inTraining[[i]],]
fitControl <- trainControl(method = "none", classProbs = TRUE)
svmLinearFit <- train(target ~ ., data = training,
method = "svmLinear",
trControl = fitControl,
family=binomial)
preds<- predict(svmLinearFit, newdata = testing)
matrix <- confusionMatrix(preds,frameDataFinal$target[-inTraining[[i]]])
allAccuracyGroup2 <- c(allAccuracyGroup2,matrix[3]$overall[[1]])
}
mean(allAccuracyGroup2)
sd(allAccuracyGroup2)
# +
#Group 3
allDataGroup3<-list()
allDataGroup3Target<-list()
groups3 = groups[groups$dataset_ID==3, ]
index<-1
for (id in groups3$X.sequence_ID){
caminho <-paste("./MovementAAL/dataset/MovementAAL_RSS_",id,".csv",sep="")
allDataGroup3[[index]]<-read.csv(file=caminho,head=TRUE,sep=",")
allDataGroup3Target[index]<-targetAll[[2]][id]
index<-index+1
}
wtData <- NULL
minStepsBack = 17
for (i in 1:length(allDataGroup3)){
aMatrix <- data.matrix(allDataGroup3[[i]], rownames.force = NA)
wt <- dwt(aMatrix[1:minStepsBack,], filter="haar", boundary="periodic")
wtData <- rbind(wtData, unlist(c(wt@W,wt@V[[wt@level]])))
}
wtData <- as.data.frame(wtData)
data = unlist(allDataGroup3Target)
target = factor(data,labels=c("No","Yes"))
frameDataFinal <- data.frame(cbind(target, wtData))
tail(frameDataFinal)
# -
# #### Média e Desvio padrão respectivamente.
# ##### Group 3, com Cross Validation tipo 10-fold
# +
inTraining <- createDataPartition(frameDataFinal$target, p = .7, list = TRUE,times=10)
allAccuracyGroup3 <- c()
for( i in 1:length(inTraining)){
training <- frameDataFinal[ inTraining[[i]],]
testing <- frameDataFinal[-inTraining[[i]],]
fitControl <- trainControl(method = "none", classProbs = TRUE)
svmLinearFit <- train(target ~ ., data = training,
method = "svmLinear",
trControl = fitControl,
family=binomial)
preds<- predict(svmLinearFit, newdata = testing)
matrix <- confusionMatrix(preds,frameDataFinal$target[-inTraining[[i]]])
allAccuracyGroup3 <- c(allAccuracyGroup3,matrix[3]$overall[[1]])
}
mean(allAccuracyGroup3)
sd(allAccuracyGroup3)
# +
#All Groups
allData<-list()
allDataTarget<-list()
targetAll <- read.csv(file="./MovementAAL/dataset/MovementAAL_target.csv",head=TRUE,sep=",")
index<-1
for (id in targetAll$X.sequence_ID){
caminho <-paste("./MovementAAL/dataset/MovementAAL_RSS_",id,".csv",sep="")
allData[[index]]<-read.csv(file=caminho,head=TRUE,sep=",")
allDataTarget[index]<-targetAll[[2]][id]
index<-index+1
}
wtData <- NULL
minStepsBack = 17
for (i in 1:length(allData)){
aMatrix <- data.matrix(allData[[i]], rownames.force = NA)
wt <- dwt(aMatrix[1:minStepsBack,], filter="haar", boundary="periodic")
wtData <- rbind(wtData, unlist(c(wt@W,wt@V[[wt@level]])))
}
wtData <- as.data.frame(wtData)
data = unlist(allDataTarget)
target = factor(data,labels=c("No","Yes"))
frameDataFinal <- data.frame(cbind(target, wtData))
tail(frameDataFinal)
# -
# #### Média e Desvio padrão respectivamente.
# ##### Todos os Groups em uma base apenas, com Cross Validation tipo 10-fold
# +
inTraining <- createDataPartition(frameDataFinal$target, p = .7, list = TRUE,times=10)
allAccuracy <- c()
for( i in 1:length(inTraining)){
training <- frameDataFinal[ inTraining[[i]],]
testing <- frameDataFinal[-inTraining[[i]],]
fitControl <- trainControl(method = "none", classProbs = TRUE)
svmLinearFit <- train(target ~ ., data = training,
method = "svmLinear",
trControl = fitControl,
family=binomial)
preds<- predict(svmLinearFit, newdata = testing)
matrix <- confusionMatrix(preds,frameDataFinal$target[-inTraining[[i]]])
allAccuracy <- c(allAccuracy,matrix[3]$overall[[1]])
}
mean(allAccuracy)
sd(allAccuracy)
# -
# #### Matrix de confusão
# #### Todos os Groups em uma base apenas
# +
#All groups datasets Confusion Matrix
inTraining <- createDataPartition(frameDataFinal$target, p = .7, list = TRUE,times=1)
training <- frameDataFinal[ inTraining[[1]],]
testing <- frameDataFinal[-inTraining[[1]],]
fitControl <- trainControl(method = "none", classProbs = TRUE)
svmLinearFit <- train(target ~ ., data = training,
method = "svmLinear",
trControl = fitControl,
family=binomial)
preds<- predict(svmLinearFit, newdata = testing)
matrix <- confusionMatrix(preds,frameDataFinal$target[-inTraining[[1]]])
matrix
# -
# #### Curva ROC e AUC
# #### Todos os Groups em uma base apenas
#ROC CURVE AND AUC
predsProb<- predict(svmLinearFit, newdata = testing,type="prob")
outcome<- predsProb[,2]
classes <- frameDataFinal$target[-inTraining[[1]]]
rocobj <- roc(classes, outcome,levels=c("No","Yes"))
plot(rocobj)
|
With Extractor of Caracteristics.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from PIL import Image
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
#pathBase = 'E://Chuan//Documents//GitHub//AusculPi//Data//VSD//'
#pathBase = 'I://Chuan//Documents//GitHub//AusculPi//Data//VSD//'
#pathBase = 'C://Users//Chuan//OneDrive//Dowrun//Text//Manuscripts//Sound//TAVR_post//'
# -
#pathBase = 'C://Users//triti//OneDrive//Dowrun//Text//Manuscripts//Data//HouCuiyu//AusculPi//'
pathBase = 'C://Users//Chuan//OneDrive//Dowrun//Text//Manuscripts//Data//JiangWenxia//AusculPi//'
filename = 'Numpy_Array_File_2020-07-02_13_57_25.npy'
line = pathBase + filename
arr = np.load(line)
arr
arr.shape
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[0], linewidth=1.0, color='black')
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[:,200], linewidth=1.0, color='black')
# +
start = 472
end = 1400
start_adj = int(start * 2583 / 3000)
end_adj = int(end * 2583 / 3000)
# -
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[start_adj:end_adj,170], linewidth=0.6, color='black')
fig = plt.figure()
s = fig.add_subplot(111)
s.plot(arr[start_adj:end_adj,100:500], linewidth=0.2, color='black')
fig.savefig('Phonocardiograph_JiangWenxia.png', dpi=500)
s.show()
|
SoundParser_JiangWenxia.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# The [iris dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) is a standard example used to illustrate machine-learning and visualization techniques. Here, we show how to use [Panel](http://panel.pyviz.org) to create a dashboard for visualizing the dataset. The Panel dashboard uses [hvPlot](http://hvplot.pyviz.org) to create plots and [Param](https://param.pyviz.org) objects to create options for selecting the `X` and `Y` axis for the plot. First, let's import the packages we are using:
# +
import hvplot.pandas
import param
import panel as pn
import pandas as pd
from bokeh.sampledata.iris import flowers
pn.extension(embed=True)
# -
# The `flowers` dataset we imported from Bokeh has five columns: `sepal_length`, `sepal_width`, `petal_length`, `petal width`, and `species`.
flowers.head(2)
# We will start by using the dataframe with these five features and then create a `Selector` object to develop menu options for different input features. Later we will define the core plotting function in a `plot` method and define the layout in the `panel` method of the `IrisDashboard` class.
#
# The `plot` method watches the `X_variable` and `Y_variable` using the `param.depends` [decorator](https://www.google.com/search?q=python+decorator), setting the `watch` option of this decorator to `True`. The `plot` method plots the features selected for `X_variable` and `Y_variable` and colors them using the `species` column.
# +
inputs = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width']
class IrisDashboard(param.Parameterized):
X_variable = param.Selector(inputs, default=inputs[0])
Y_variable = param.Selector(inputs, default=inputs[1])
@param.depends('X_variable', 'Y_variable', watch=True)
def plot(self):
return flowers.hvplot.scatter(x=self.X_variable, y=self.Y_variable, by='species')
def panel(self):
return pn.Row(self.param, self.plot)
dashboard = IrisDashboard(name='Iris_Dashboard')
# -
# And now you can explore how each of the input columns relate to each other, either here in the notebook or when served as a separate dashboard using `panel serve --show Iris_dataset.ipynb`:
dashboard.panel().servable()
|
examples/gallery/param/reactive_plots.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import cobra
import numpy as np
import matplotlib.pyplot as plt
from scipy.optimize import linprog
import scipy
import mip
from copy import deepcopy
def save_dict(data, name):
with open(name, 'w' ) as file:
json.dump( data, file )
# -
# # Community modeling
#
# In this notebook we will implement a method to create community models of two or more species specific metabolic models using cobrapy.
model_DP = cobra.io.read_sbml_model("models/consistent_DP_SNM.xml")
model_SA = cobra.io.read_sbml_model("models/consistent_iYS854_SNM.xml")
print("Growth: ", model_DP.slim_optimize())
print("Growth: ", model_SA.slim_optimize())
for rec in model_SA.reactions:
rec.lower_bound = max(rec.lower_bound, -1000)
rec.upper_bound = min(rec.upper_bound, 1000)
snm3 = pd.read_csv("SNM3.csv", sep =";")
snm3.head()
BIOMASS_DP = "Growth"
BIOMASS_SA = "BIOMASS_iYS_wild_type"
models = [model_DP.copy(), model_SA.copy()]
from community_models import *
# +
import json
compm_SA = json.loads(open("compm_SA.json").read())
compm_DP = json.loads(open("compm_DP.json").read())
model_DP.medium = compm_DP
model_SA.medium = compm_SA
# +
model1 = Model(model_DP, BIOMASS_DP)
model2 = Model(model_SA, BIOMASS_SA)
community_model1 = model1 + model2
# -
community_model2 = MIP_community_model(model1, model2)
# + tags=[]
community_model3_1_1 = create_bag_of_react_model([model_DP, model_SA],[BIOMASS_DP, BIOMASS_SA], [1,1])
community_model3_10_1 = create_bag_of_react_model([model_DP, model_SA],[BIOMASS_DP, BIOMASS_SA], [10,1])
# -
community_model1.set_weights([1,1])
print("MBR Weights 1:1: ", community_model1.slim_optimize())
single_growth = community_model1.optimize().x[community_model1.objective_c != 0]
print("DP growth: ", single_growth[0])
print("SA growth: ", single_growth[1])
community_model1.set_weights([10,1])
print("MBR Weights 10:1: ", community_model1.slim_optimize())
single_growth = community_model1.optimize().x[community_model1.objective_c != 0]
print("DP growth: ", single_growth[0])
print("SA growth: ", single_growth[1])
community_model2.weights = [1,1]
print("MBR Weights 1:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
community_model2.weights = [10,1]
print("MBR Weights 10:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
print("MBR Weights 1:1: ", community_model3_1_1.slim_optimize())
print("SA growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_DP).flux))
print("MBR Weights 10:1: ", community_model3_10_1.slim_optimize())
print("SA growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_DP).flux))
coopm = community_model2.compute_coopm()
coopm2 = optimize_coopm_community(community_model3_1_1, community_model3_1_1.slim_optimize(), [BIOMASS_DP, BIOMASS_SA], [1,1])
coopm
coopm2
community_model2.set_medium(coopm)
community_model2.weights = [1,1]
print("MBR Weights 1:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
community_model2.weights = [10,1]
print("MBR Weights 10:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
community_model3_1_1.medium = coopm
print("MBR Weights 1:1: ", community_model3_1_1.slim_optimize())
print("SA growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_DP).flux))
community_model3_10_1.medium = coopm
print("MBR Weights 10:1: ", community_model3_10_1.slim_optimize())
print("SA growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_DP).flux))
community_model2.set_medium(coopm2)
community_model2.weights = [1,1]
print("MBR Weights 1:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
community_model2.weights = [10,1]
print("MBR Weights 10:1: ", community_model2.optimize())
print("SA growth: ", community_model2.x2[community_model2.obj2].x)
print("DP growth: ", community_model2.x1[community_model2.obj1].x)
community_model3_1_1.medium = coopm2
print("MBR Weights 1:1: ", community_model3_1_1.slim_optimize())
print("SA growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_1_1.reactions.get_by_id(BIOMASS_DP).flux))
community_model3_10_1.medium = coopm2
print("MBR Weights 10:1: ", community_model3_10_1.slim_optimize())
print("SA growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_SA).flux))
print("DP growth: " + str(community_model3_10_1.reactions.get_by_id(BIOMASS_DP).flux))
# ## COOPM alphas model
#
# Here is a collection of COOPM medias for different alpha values
community_model2 = MIP_community_model(model1, model2)
alphas = [0.,0.01,0.1,0.2,0.5,0.8,0.9,0.99,1.]
coopms = []
for alpha in alphas:
coopms.append(community_model2.compute_alpha_coopm(alpha))
df = pd.DataFrame(coopms)
df.index = alphas
df.T.plot.bar(figsize=(20,10))
plt.yscale("log")
plt.ylabel("COOPM medium flux")
plt.xlabel("COOPM medium")
plt.savefig("COOPM_alpha_plot.pdf")
|
community_model_approaches_26.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import geohunter as gh
import pandas as pd
import geopandas as gpd
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
import matplotlib
matplotlib.rcParams['mathtext.fontset'] = 'stix'
matplotlib.rcParams['font.family'] = 'STIXGeneral'
warnings.filterwarnings("ignore")
# -
api = gh.osm.Eagle()
# # Study area
#
# [Natal, Brazil](https://en.wikipedia.org/wiki/Natal,_Rio_Grande_do_Norte)
city = api.get('(-8.02, -41.01, -3.0, -33.0)',
largest_geom=True,
name='Natal')
city.plot();
# # Grids
resolution = 1
p_grid = gh.util.make_gridpoints(city, resolution=resolution/2)
p_grid = p_grid.drop(columns=['lat', 'lon'])
p_grid.plot(markersize=1);
s_grid = gh.util.make_gridsquares(city, resolution)[['geometry']]
s_grid.plot(edgecolor='black')
# # Extracting Points-of-Interest data
#
# [OpenStreetMap's map features](https://wiki.openstreetmap.org/wiki/Map_Features)
poi = api.get(city,
amenity=['school', 'hospital', 'restaurant', 'police', 'place_of_worship'],
leisure='*',
highway=['primary', 'residential', 'bus_stop'],
tourism='*',
natural=['sand', 'wood', 'beach'],
shop='*'
)
poi.loc[poi['key'].isin(['leisure', 'tourism', 'shop']), 'item'] = '*'
poi.plot(column='key')
poi.head()
poi.groupby(['key','item']).size()
# # Extracting geographic features/covariates
# ## 1. Quadrat method
# +
x = []
for pair in poi[['key', 'item']].drop_duplicates().iterrows():
key, item = pair[1].values
x.append(gpd.sjoin(poi.loc[(poi['key']==key) & (poi['item']==item)], s_grid)\
.groupby('index_right').size().to_frame(f'{key}_{item}'))
X_quadrat = pd.concat(x, axis=1)\
.join(s_grid, how='outer')\
.fillna(0)
X_quadrat = gpd.GeoDataFrame(X_quadrat)
# -
feature = 'amenity_school'
X_quadrat.plot(column=feature)
# ## 2. KDE method
area_sgrid = gpd.GeoDataFrame(geometry=[s_grid.geometry.unary_union])
x = []
for pair in poi[['key', 'item']].drop_duplicates().iterrows():
key, item = pair[1].values
x.append(gh.util.kde_interpolation(poi.loc[(poi['key']==key) & (poi['item']==item)],
bw='silverman', area=area_sgrid, resolution=resolution/2)['density']\
.to_frame(f'{key}_{item}'))
X_kde = pd.concat(x, axis=1)
X_kde = p_grid.join(X_kde)
feature = 'amenity_school'
X_kde.plot(column=feature)
# Average values within the square grid cells:
X_kde = gpd.sjoin(X_kde, s_grid)\
.groupby('index_right').mean()\
.join(s_grid, how='outer')\
.fillna(0)
X_kde = gpd.GeoDataFrame(X_kde)
X_kde.plot(feature)
X_kde.head()
# # Qualitative assessment
# +
fig, ax = plt.subplots(ncols=3, nrows=3, figsize=(14,12))
for i in range(3):
for j in range(3):
s_grid.plot(edgecolor='black', color='white', ax=ax[i][j])
poi.loc[(poi['key']=='amenity') & (poi['item']=='hospital')]['geometry'].centroid.plot(ax=ax[0][0], color='red', markersize=5)
poi.loc[(poi['key']=='highway') & (poi['item']=='bus_stop')]['geometry'].centroid.plot(ax=ax[0][1], color='green', markersize=5)
poi.loc[(poi['key']=='highway') & (poi['item']=='residential')]['geometry'].centroid.plot(ax=ax[0][2], markersize=3)
X_quadrat.plot(column='amenity_hospital', ax=ax[1][0], legend=True, edgecolor='black', cmap='Reds')
X_quadrat.plot(column='highway_bus_stop', ax=ax[1][1], legend=True, edgecolor='black', cmap='Greens')
X_quadrat.plot(column='highway_residential', ax=ax[1][2], legend=True, edgecolor='black', cmap='Blues')
X_kde.plot(column='amenity_hospital', ax=ax[2][0], legend=True, edgecolor='black', cmap='Reds')
X_kde.plot(column='highway_bus_stop', ax=ax[2][1], legend=True, edgecolor='black', cmap='Greens')
X_kde.plot(column='highway_residential', ax=ax[2][2], legend=True, edgecolor='black', cmap='Blues')
ax[0][0].set_ylabel('PoI data', fontsize=18)
ax[1][0].set_ylabel('Quadrat', fontsize=18)
ax[2][0].set_ylabel('KDE', fontsize=18)
ax[2][0].set_xlabel('amenity_hospital', fontsize=18)
ax[2][1].set_xlabel('highway_bus_stop', fontsize=18)
ax[2][2].set_xlabel('highway_residential', fontsize=18)
sns.despine(trim=True)
fig.tight_layout()
fig.savefig('qualitative.pdf', dpi=200)
# -
# # Quantitive assessment
# Spatial heterogeneity (q) needs to use stratas on its calculation. We chose to get the city's neighborhoods as stratas.
nhoods = api.get(city, admin_level=10)[['geometry', 'name']]
nhoods.plot();
s_grid = s_grid.join(gpd.sjoin(s_grid, nhoods)['name'].to_frame('strata'))
s_grid = gpd.GeoDataFrame(s_grid).drop_duplicates(subset=['geometry'])
s_grid.plot(column='strata');
scores_quadrat, scores_kde = {}, {}
for feature in X_kde.drop(columns=['geometry']).columns:
scores_kde[feature] = {'q':gh.util.q_ongrid(X_kde[feature], s_grid, strata_col='strata'),
'I':gh.util.moran_i_ongrid(X_kde[feature],
s_grid['geometry'].centroid.apply(lambda x:x.coords[0]),
d_threshold=resolution/110.5)[0]}
scores_quadrat[feature] = {'q':gh.util.q_ongrid(X_quadrat[feature], s_grid, strata_col='strata'),
'I':gh.util.moran_i_ongrid(X_quadrat[feature],
s_grid['geometry'].centroid.apply(lambda x:x.coords[0]),
d_threshold=resolution/110.5)[0]}
# +
scores_kde = scores_kde = pd.DataFrame(scores_kde).T.reset_index()\
.melt(value_vars=['q','I'], id_vars=['index'])\
.assign(Method='KDE')
scores_quadrat = pd.DataFrame(scores_quadrat).T.reset_index()\
.melt(value_vars=['q','I'], id_vars=['index'])\
.assign(Method='Quadrat')
scores = pd.concat([scores_quadrat, scores_kde]).rename(columns={'index':'feature'})
# +
fig, ax = plt.subplots(ncols=2, figsize=(12, 5))
sns.barplot(y='feature', x='value', hue='Method', data=scores.loc[scores['variable']=='q'], ax=ax[0], palette='viridis')
sns.barplot(y='feature', x='value', hue='Method', data=scores.loc[scores['variable']=='I'], ax=ax[1], palette='viridis')
ax[0].yaxis.set_tick_params(labelsize=14)
ax[0].xaxis.set_tick_params(labelsize=12)
ax[0].set_ylabel('')
ax[0].set_xlabel('Spatial heterogeneity\nq', fontsize=18)
ax[0].get_legend().remove()
ax[1].legend(fontsize=18)
ax[1].xaxis.set_tick_params(labelsize=12)
ax[1].set_yticks([])
ax[1].set_ylabel('', fontsize=0)
ax[1].set_xlabel('Spatial autocorrelation\nI', fontsize=18)
fig.tight_layout()
fig.savefig('quantitative.pdf', dpi=200)
# -
# # Demonstration and applications
# +
from sklearn.cluster import KMeans
features = ['amenity_hospital', 'highway_bus_stop', 'highway_residential']
X = X_kde[features]
kmeans = KMeans(3).fit(X)
X['Clusters'] = kmeans.labels_
X['Clusters'] = X['Clusters'].apply(lambda x: [f'C{i}' for i in range(5)][x])
# -
fig = sns.pairplot(X, hue='Clusters', size=1.5, palette='Set1')
fig.savefig('cluster_histograms.png', dpi=500)
# +
norm=plt.Normalize(-2,2)
cmap = matplotlib.colors.LinearSegmentedColormap.from_list("", ["#E53631","#377EB8","#50AF4A"])
fig, ax = plt.subplots()
X['geometry'] = X_kde['geometry']
gpd.GeoDataFrame(X).plot(column='Clusters', cmap=cmap, edgecolor='black', alpha=0.9, legend=True, ax=ax)
sns.despine(trim=True)
fig.savefig('clusters_map.png', dpi=200)
|
examples/paper_experiment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Talktorial 3
#
# # Molecular filtering: unwanted substructures
#
# #### Developed in the CADD seminars 2017 and 2018, AG Volkamer, Charité/FU Berlin
#
# <NAME> and <NAME>
# ## Aim of this talktorial
#
# There are some substructures we prefer not to enclose into our screening library. In this talktorial, we learn about different types of such unwanted substructures and how to find and highlight them with RDKit.
#
# ## Learning goals
#
# ### Theory
#
# * What are unwanted substructures?
# * Pan Assay Interference Compounds (PAINS)
#
# ### Practical
#
# * Read a set of compounds from ChEMBL database (prepared in **talktorial 2**)
# * Filter out unwanted substructures as implemented in RDKit
# * Provide your own list of unwanted substructures and filter your data based on them
# * Search and highlight substructures
#
# ## References
#
# * Brenk et al.: "Lessons learnt from assembling screening libraries for drug discovery for neglected diseases" <i>Chem. Med. Chem.</i> (2008), <b>3</b>,435-444 (https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)
# * Brenk et al.: SMARTS definitions of unwanted groups (Table 1 of the Supporting
# information of <i>Chem. Med. Chem.</i> (2008), <b>3</b>,435-444 (https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)
# * Baell et al.: "New substructure filters for removal of Pan Assay Interference Compounds (PAINS) from screening libraries and for their exclusion in bioassays" <i>J. Med. Chem.</i> (2010), <b>53(7)</b>,2719-2740 (https://pubs.acs.org/doi/abs/10.1021/jm901137j)
# * Conversion of PAINS format from SLN (Baell et al. publication) to SMARTS by <NAME>: http://blog.rguha.net/?p=850; SMARTS that are used in the RDKit are the ones curated by <NAME> from <NAME>'s KNIME workflow: http://rdkit.blogspot.com/2015/08/curating-pains-filters.html
# * https://en.wikipedia.org/wiki/Pan-assay_interference_compounds
# * TDT -Tutorial2014 - (https://github.com/sriniker/TDT-tutorial-2014/blob/master/TDT_challenge_tutorial.ipynb)
# _____________________________________________________________________________________________________________________
#
#
# ## Theory
#
# ### Unwanted substructures
#
# Substructures can be unfavorable, e.g. because they are toxic or reactive, due to unfavorable pharmacokinetic properties, or because they likely interfere with certain assays.
# Nowadays, drug discovery often involves high throughput screening ([HTS wikipedia](https://en.wikipedia.org/wiki/High-throughput_screening)). Filtering unwanted substructures can support assembling more efficient screening libraries. Hence, reducing a library before screening can save time and resources.
#
# Brenk *et al.* ([<i>Chem. Med. Chem.</i> (2008), <b>3</b>,435-444](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)) have constructed a list of unfavorable substructures to filter their libraries used to screen for compounds to treat neglected diseases.
# Examples of such unwanted features are nitro groups (mutagenic), sulfates and phosphates (likely resulting in unfavorable pharmacokinetic properties), 2-halopyridines and thiols (reactive).
#
# This list of undesired substructures was published in the above mentioned paper and will be used in the practical part of this talktorial.
# ### Pan Assay Interference Compounds (PAINS)
#
# #### General information
# PAINS ([PAINS wikipedia](https://en.wikipedia.org/wiki/Pan-assay_interference_compounds)) are compounds that often occur as hits in HTS even though they actually are false positives. PAINS tend to react unspecifically with numerous targets rather than one specific target. Apparent binding in various assays and against a range of proteins is usually due to unspecific binding or interaction with assay components.
# [](https://commons.wikimedia.org/wiki/File:PAINS_Figure.tif)
# <div align="center">Figure 1: Specific and unspecific binding in the context of Pan Assay Interference Compounds. Figure is taken from [wikipedia](https://commons.wikimedia.org/wiki/File:PAINS_Figure.tif). </div>
# #### Filtering in the paper of Baell et al.
#
# Baell *et al.* ([<i>J. Med. Chem.</i> (2010), <b>53(7)</b>,2719-2740](https://pubs.acs.org/doi/abs/10.1021/jm901137j)) focused on substructures interfering in assay signaling. They described substructures which can help to identify such PAINS and provided a list which can be used for substructure filtering.
# ## Practical
#
# ### Load and visualize data
# First, we import the required libraries, load our filtered dataset from **talktorial T2**, and draw the first molecules.
import pandas
from rdkit import Chem
from rdkit.Chem.Draw import IPythonConsole
from rdkit.Chem import rdFMCS
from rdkit.Chem import AllChem
from rdkit.Chem import Descriptors
from rdkit.Chem import Draw
from rdkit import DataStructs
from rdkit.Chem import PandasTools
import matplotlib.pyplot as plt
filteredData = pandas.read_csv("../data/T2/EGFR_compounds_lipinski.csv", delimiter=";", index_col=0)
filteredData.drop(['HBD','HBA','MW','LogP'], inplace=True, axis=1) # Drop unnecessary information
print ('Dataframe shape: ', filteredData.shape) # Print dimension of the dataframe
filteredData.head(5)
PandasTools.AddMoleculeColumnToFrame(filteredData, smilesCol='smiles') # Add molecule column
# Draw first 20 molecules
Draw.MolsToGridImage(list(filteredData.ROMol[0:20]),
legends=list(filteredData.molecule_chembl_id[0:20]),
molsPerRow=4)
# ### Filtering for PAINS using RDKit
#
# The PAINS filter is already implemented in RDKit ([RDKit Documentation](http://rdkit.org/docs/source/rdkit.Chem.rdfiltercatalog.html)), let's learn how it can be used.
from rdkit.Chem.FilterCatalog import *
params = FilterCatalogParams()
# Build a catalog from all PAINS (A, B and C)
params.AddCatalog(FilterCatalogParams.FilterCatalogs.PAINS)
catalog = FilterCatalog(params)
# +
# Create empty dataframes for filtered data
rdkit_highLightFramePAINS = pandas.DataFrame(columns=('CompID', 'CompMol', 'unwantedID'))
rdkit_noPAINS = pandas.DataFrame(columns=('ChEMBL_ID', 'smiles','pIC50'))
rdkit_withPAINS = pandas.DataFrame(columns=('ChEMBL_ID', 'smiles', 'pIC50','unwantedID'))
# For index and row in the filtered df
for i,row in filteredData.iterrows():
curMol = Chem.MolFromSmiles(row.smiles) # Current molecule
match = False # Set match to false
rdkit_PAINSList = []
# Get the first match
entry = catalog.GetFirstMatch(curMol)
if entry!=None:
# Add name of current unwanted subsftructure to list
rdkit_PAINSList.append(entry.GetDescription().capitalize())
# Add relevant matching information to dataframe
rdkit_highLightFramePAINS.loc[len(rdkit_highLightFramePAINS)] = [row.molecule_chembl_id, curMol,
entry.GetDescription().capitalize()]
match = True
if not match:
# Add to frame of PAINS free compounds
rdkit_noPAINS.loc[len(rdkit_noPAINS)] = [row.molecule_chembl_id, row.smiles, row.pIC50]
else:
# Add to frame of compounds that contain PAINS
# Put the relevant information in the dataframe with the unwanted substructures
rdkit_withPAINS.loc[len(rdkit_withPAINS)] = [row.molecule_chembl_id, row.smiles, row.pIC50, entry.GetDescription().capitalize()]
print("Number of compounds with PAINS: %i"%(len(rdkit_withPAINS)))
print("Number of compounds without PAINS: %i (=remaining compounds)"%(len(rdkit_noPAINS)))
rdkit_highLightFramePAINS.head(10)
# -
# ### Filtering for unwanted/toxic substructures (Brenk list)
#
# Some lists of unwanted substructures, like PAINS, are already implemented in RDKit. However, it is also possible to use an external list and get the substructure matches.
# Here, we use the list provided in the supporting information by Brenk et al. ([<i>Chem. Med. Chem.</i> (2008), <b>3</b>,435-444](https://onlinelibrary.wiley.com/doi/full/10.1002/cmdc.200700139)).
#
# Note, we downloaded the data and stored it as csv file in the data folder (the format is name-space-SMARTS).
# First, we load the data.
# +
unwantedSubs = []
unwantedNames = []
for line in open('../data/T3/unwantedSubstructures.csv', 'r'):
if not line.startswith("#"): # Ignore header
splitted = line.strip().split(" ") # Split each line
m = Chem.MolFromSmarts(splitted[1]) # Generate a molecule from smarts
name = splitted[0].capitalize() # Store the name in name
unwantedNames.append(name) # Append the names of the unwanted substructes to a list
unwantedSubs.append(m) # Append the unwanted substructes to a list
print("Number of unwanted substructures in list =", len(unwantedSubs)) # Show the number of unwanted substructures
# -
# Let's have a look at a few substructures (not all SMARTS can be displayed, therefore we select a subset).
Chem.Draw.MolsToGridImage(list(unwantedSubs[2:5]), subImgSize=(200, 300), legends=unwantedNames[2:5])
# Search our filtered dataframe for matches with these unwanted substructures.
# +
# Creating dataFrames for filtered data
highLightFrameUNW = pandas.DataFrame(columns=('CompID', 'CompMol', 'unwantedID', 'unwSubstr'))
noUnwanted = pandas.DataFrame(columns=('ChEMBL_ID', 'smiles','pIC50'))
withUnwanted = pandas.DataFrame(columns=('ChEMBL_ID', 'smiles', 'pIC50','unwantedID'))
molsToDraw = []
# For each compound in data set
for i,row in filteredData.iterrows(): # For index and row in the filtered dataframe
curMol = Chem.MolFromSmiles(row.smiles) # Current molecule
match = False # Set match to false
unwantedList = []
molsToDraw.append(curMol)
# Search for all unwanted substructures
for idx, unwSub in enumerate(unwantedSubs):
# Check if it has the substructure
if curMol.HasSubstructMatch(unwSub): # If the current molecule has the unwanted substructure
match = True # Set match to True
unwantedList.append(unwantedNames[idx]) # Add name of unwanted substructure to list
# Put the relevant information in the dataframe
highLightFrameUNW.loc[len(highLightFrameUNW)] = [row.molecule_chembl_id, curMol, unwantedNames[idx], unwSub]
if not match: # If no match was found
noUnwanted.loc[len(noUnwanted)] = [row.molecule_chembl_id, row.smiles, row.pIC50]
# Add relevant information to dataframe of the wanted substructes
else: # If a match was found
withUnwanted.loc[len(withUnwanted)] = [row.molecule_chembl_id, row.smiles, row.pIC50, unwantedList] #put the relevant information in the dataframe with the unwanted substructures
print("Number of compounds with unwanted substructures: %i"%(len(withUnwanted)))
print("Number of compounds without unwanted substructures: %i (=remaining compounds)"%(len(noUnwanted)))
# -
highLightFrameUNW.head(8)
# The substructures can also be highlighted, directly within the molecule.
# +
first_highLightFrameUNW = highLightFrameUNW.head(8) # Subset of the first 8 entries in the list
# Draw molecules and highlight the unwanted substructure
Draw.MolsToGridImage(list(first_highLightFrameUNW["CompMol"]), subImgSize=(400,300),
molsPerRow=2, highlightAtomLists=
[m.GetSubstructMatch(first_highLightFrameUNW["unwSubstr"][i]) for i,m in enumerate(first_highLightFrameUNW["CompMol"])],
legends=list(first_highLightFrameUNW["CompID"]+": "+first_highLightFrameUNW["unwantedID"]))
# -
# We save the example as SVG file.
# +
# Save image to file
img = Draw.MolsToGridImage(list(first_highLightFrameUNW["CompMol"]), subImgSize=(400,300),
molsPerRow=3, highlightAtomLists=
[m.GetSubstructMatch(first_highLightFrameUNW["unwSubstr"][i]) for i,m in enumerate(first_highLightFrameUNW["CompMol"])],
legends=list(first_highLightFrameUNW["unwantedID"]), useSVG=True)
# Get SVG data
molsvg = img.data
# Replace non-transparent to transparent background
molsvg = molsvg.replace("opacity:1.0", "opacity:0.0");
molsvg = molsvg.replace("12px", "24px");
# Save altered SVG data to file
f = open("../data/T3/substructures.svg", "w")
f.write(molsvg)
f.close()
# -
# The lists of compounds with and without unwanted substructures are saved.
# +
# Write the compounds with unwanted substructures in a csv file
withUnwanted.to_csv("../data/T3/EGFR_compounds_lipinski_noPAINS.csv", sep=',')
# Write the compounds without unwanted substructures in a csv file
noUnwanted.to_csv("../data/T3/EGFR_compounds_lipinski_noPAINS_noBrenk.csv", sep=',')
# Show the head of the csv file with the compounds with the unwanted substructures i a csv file
noUnwanted.head()
# -
# The unwanted substructures found are further analyzed.
# +
# Count the most frequent compounds
unwCounts = {}
for ele in highLightFrameUNW.values:
unw = ele[2] # ID of the unwanted substructure from the dataframe highLightFrameUNW
if unwCounts.get(unw, "empty") == "empty": # If the ID of the unwanted structure is not yet in the dictionary
unwCounts[unw] = [1, ele[3]] # Put a 1 and the unwanted structure in the dictionary
else: # If the key (unwanted structure ID) already exists, increment the value of occurences
unwCounts[unw][0] += 1
frequentUNW = []
frequentUNWNames = []
frequentUNWList = []
# Structure of unwCounts: Dictionary with the ID as key, and the number of occurences and molecule as values
# E.g. ('acyclic-C=C-O', [7, <rdkit.Chem.rdchem.Mol object at 0x7fa58fc06710>])
# Sort the dictionary by frequencies of substructures
for key, value in sorted(unwCounts.items(), key=lambda kv: kv[1][0], reverse=True):
frequentUNW.append(value[1]) # Substructure
frequentUNWNames.append(key)
frequentUNWList.append((value[0], key))
frequentUNWList[:8]
# -
# Draw the eight frequent substructures.
# +
# Eight most frequent substructeres
Draw.MolsToGridImage(mols=list(frequentUNW[0:2]), subImgSize=(400,300),
legends=list(frequentUNWNames[0:2]),
molsPerRow=4)
# -
# ## Discussion
# In this talktorial, we learned two ways to perform a search for unwanted substructures:
# * Once with the class `FilterCatalog` which is already implemented in RDKit, and
# * Once with an external list and the `HasSubstructMatch()` function from RDKit.
#
# Actually, PAINS could also be implemented via substructure search. And also the list by Brenk et al. is already in RDKit. Further lists that are implemented can be found in the ([RDKit Documentation](http://rdkit.org/docs/source/rdkit.Chem.rdfiltercatalog.html)).
#
# So far, we have been using the function `HasSubstructMatch()` which only yields one match per compound. With [`GetSubstructMatches()`](http://www.rdkit.org/Python_Docs/rdkit.Chem.rdchem.Mol-class.html#GetSubstructMatches), we also have the option to find all substructures matching one compound.
# Similarly for PAINS: We have only looked at the first match `GetFirstMatch()` per molecule. If we want to filter out all PAINS this is enough. However, we could also use `GetMatches()` in order to see all critical substructures of a molecule. Since we did not take all matches per molecule into account, we cannot state that the last substructures we drew, are actually the most frequent ones. However, they definitely do occur very often.
#
# Detected substructures can be handled in two different ways.
# * Either, the substructure search is applied as a filter and the compounds are excluded from further testing to reduce the money and time.
# * Or, they can be used as warnings. Molecules containing unwanted substructures can be flagged. Expert's (e.g. chemists, toxicologists...) eyes might be able to judge from their experience. If the respective substructures are less critical, they might still include them for screening.
#
# We will not filter out the unwanted substructures as we do not want to lose too many compounds for machine learning. Also, the substructure filters could still be applied in a later stage before actual experimental screening. There's the possibility to set alert flags, so the information about unfavorable substructures (according to any list, e.g. PAINS or Brenk) could be kept and considered later.
# ## Quiz
# * Why should we consider removing "PAINS" from a screening library? What is the issue with these compounds?
# * Can you find situations when some unwanted substructures would not need to be removed?
# * How are the substructures we used in this tutorial encoded?
|
talktorials/3_substructure_filtering/T3_substructure_filtering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from IPython.core.display import HTML
HTML('<style> .container{ width:90%; } </style>')
# # Load Trained Model From Pickle
import numpy as np
import pandas as pd
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
import joblib
create_model=None
trained_model = joblib.load('models/LightGBM.pkl')
columns = joblib.load('models/columns.pkl')
# # Predict Category For Sample Data
data = pd.get_dummies(pd.DataFrame({'age':43,
'workclass':'Private',
'fnlwgt':100000,
'education':'Bachelors',
'education-num':13,
'marital-status':'Married-civ-spouse',
'occupation':'Sales',
'relationship':'Husband',
'race':'White',
'sex':'Male',
'capital-gain':0,
'capital-loss':0,
'hours-per-week':40,
'native-country':'Spain'}, index=[0]))
data = data.reindex(columns=columns, fill_value=0)
trained_model.predict(data)
# # Explain Weights and Prediction Using ELI5
# Split the pipeline between estimator (last step) and transformer (all steps bar last). Then get the transformed data and columns
# +
from sklearn.pipeline import Pipeline
estimator = trained_model
transformed_data = data
feature_names = columns
if type(trained_model) == Pipeline:
if len(trained_model.steps) == 1:
estimator = trained_model.steps[0][1]
else:
estimator = trained_model.steps[-1][1]
transformer = Pipeline(trained_model.steps[:-1])
transformed_data = transformer.transform(data)
if 'select' in transformer.named_steps:
feature_names = data.columns[transformer.named_steps.select.get_support()].tolist()
# -
import eli5
eli5.show_weights(estimator, feature_names=feature_names, top=None, show=eli5.formatters.fields.ALL)
eli5.show_prediction(estimator, transformed_data[0], feature_names=feature_names,
top=None, show=eli5.formatters.fields.ALL)
|
Census Income Prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Leer y almacenar datos usando *Pandas*
#
# Lo que nos interesa durante el curso es poder hacer análisis de datos, al menos que se ingrese de manera manual cada dato en una sesión de Python sera necesario poder tener acceso a distintas fuentes de información, es decir poder cargar datos. Todos los datos se guradan en forma binaria (ceros y unos), en los principios de la computación cada persona decidía como se guardaba la información, esto generó muchos tipos de formatos y muchos problemas para compartir la información. Por tal motivo los distintos tipos de formatos se estandarizaron generando así formatos restrictivo pero que todo podian ser compartidos entre las personas.
#
# La intención del curso no es que sepan como leer archivos en su forma binaria, si no hacer uso de las herramientas que se tienen disponibles para tal proposito. Nos enfocaremos en como usar las herramientas de *Pandas* para poder cargar y almacenar los datos de fuentes externa principalmenta a partir de archivos de texto que guerdan datos, formatos especializados para el almacenamiento de datos como son los archivos de excel (*.xsl*) o extraccion de fuentes de información como bases de datos.
#
# ## Archivos CSV
#
# Los archivos que comunmente se utilizan para almacenar datos son conocidos como *CSV* que tienen terminación `.csv`, este tipo de archivos guarda los datos de una tabla en forma de texto usando comas para la separación de los datos, de hay el nombre de este tipo de archivo *coma separated value (CSV)*.
#
# Podemos generar un archivo *csv* usando un editor de texto escribiendo el siguiente texto
#
# `AKBA,"Akebia Therapeutics Inc.", $2.69, -7.30, -73.05%
# CYCN,"Cyclerion Therapeutics Inc." , $6.44 , -1.25 , -16.36%
# EVGN,"Evogene Ltd.",$2.37, -0.44, -15.95%
# SPWH,"Sportsmans Warehouse Holdings Inc.",$14.50,-2.75,-15.94%
# CONN,"Conns Inc.", $11.54 ,-2.16,-15.77%
# `
#
# Y guardarlo con con el nombre `prueba.csv`.
#
# A continuación vamos a guardar el archivo usando únicamente usando código de *Python* lo ponemos aquí únicamente como referencia, por si se desea aprender como almacenar archivos de texto.
#
archivo_ob= open('prueba.csv', 'w')
texto = """AKBA,"Akebia Therapeutics Inc.", $2.69, -7.30, -73.05% \n
CYCN,"Cyclerion Therapeutics Inc." , $6.44 , -1.25 , -16.36% \n
EVGN,"Evogene Ltd.",$2.37, -0.44, -15.95% \n
SPWH,"Sportsmans Warehouse Holdings Inc.",$14.50,-2.75,-15.94%\n
CONN,"Conns Inc.", $11.54 ,-2.16,-15.77% """
archivo_ob.write(texto)
archivo_ob.close()
# La función `open` nos permite generar un objeto de tipo archivo, el cual se abre en modo de escritura (para abrirlo en modo de lectura se utiliza el parámetro `'r'`), una vez con el objeto se utiliza el método `write` para escribir la cadena de texto dentro del archivo. Despues se cierra el archivo para almacenarlo dentro de disco.
#
# Para leer un archivo *csv* dentro de *Pandas* se utiliza la función `read_csv` la cual toma puede tomar distintos parámetros, los cuales dependen de como se encuentren nuestros datos, por lo cual es recomendable hacer una inspección del archivo *csv* con la intención de conocer lo que contiene el archivo, en algunas ocaciones esto no es posible debido al tamaño del archivo.
#
# Como nosotros conocemos como se encuentra nuestro archivo
import pandas as pd
df_1 = pd.read_csv('prueba.csv', header=None)
df_1
#
df_1.columns= ['Simbolo', 'Nombre', 'Ultimo','Cambio', 'Porcentage cambio']
df_1
df_1.to_csv('prueba_1.csv')
df_1
|
Leer_guardar.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
# %matplotlib inline
df1 = pd.read_csv('xbtusdt_orderbook1.csv', index_col=0)
df2 = pd.read_csv('xbtusdt_orderbook2.csv', index_col=0)
df3 = pd.read_csv('xbtusdt_orderbook3.csv', index_col=0)
df1[['ask_p_0', 'bid_p_0']].plot()
df2[['ask_p_0', 'bid_p_0']].plot()
df3[['ask_p_0', 'bid_p_0']].plot()
# +
import numpy as np
def feature(df):
for i in range(10):
ind = str(i)
df['spreed_'+ind] = df['ask_p_'+ind] - df['bid_p_'+ind]
df['mid_p'+ind] = (df['ask_p_'+ind] + df['bid_p_'+ind]) / 2
for i in range(9):
ind = str(i+1)
df['ask_p_diff_'+ind+'_0'] = df['ask_p_'+ind] - df['ask_p_0']
df['bid_p_diff_'+ind+'_0'] = df['bid_p_'+ind] - df['bid_p_0']
ind1 = str(i)
df['ask_p_diff_'+ind+'_'+ind1] = df['ask_p_'+ind] - df['ask_p_'+ind1]
df['bid_p_diff_'+ind+'_'+ind1] = df['bid_p_'+ind] - df['bid_p_'+ind1]
l1, l2, l3, l4 = [], [], [], []
for i in range(10):
ind = str(i)
l1.append('ask_p_'+ind)
l2.append('bid_p_'+ind)
l3.append('ask_vol_'+ind)
l4.append('bid_vol_'+ind)
df['ask_p_mean'] = df[l1].mean(axis=1)
df['bid_mean'] = df[l2].mean(axis=1)
df['ask_vol_mean'] = df[l3].mean(axis=1)
df['bid_vol_mean'] = df[l4].mean(axis=1)
l = []
template = 'spreed_{0}'
for i in range(10):
l.append(template.format(i))
df['accum_spreed'] = df[l].sum(axis=1)
for i in range(10):
ind = str(i)
df['spreed_vol_'+ind] = df['ask_vol_'+ind] - df['bid_vol_'+ind]
l = []
template = 'spreed_vol_{0}'
for i in range(10):
l.append(template.format(i))
df['accum_spreed_vol'] = df[l].sum(axis=1)
df['y'] = np.where(df['mid_p0'] > df['mid_p0'].shift(1) , 1, 0)
df['y'] = np.where(df['mid_p0'] < df['mid_p0'].shift(1) , -1, df['y'])
return df
# -
df1 = feature(df1)
df2 = feature(df2)
df3 = feature(df3)
df1['y'].value_counts()
df2['y'].value_counts()
df3['y'].value_counts()
df = pd.concat([df1, df2, df3])
df.describe()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(df.drop(['y'], axis=1), df['y'], test_size=0.7, random_state=42)
# +
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.metrics import mean_squared_error
from sklearn.feature_selection import SelectKBest
from sklearn.feature_selection import SelectPercentile, f_classif
import xgboost as xgb
from xgboost.sklearn import XGBClassifier
from sklearn import metrics #Additional scklearn functions
import matplotlib.pyplot as plt
ind_params = {'n_estimators': 100, 'seed':0, 'colsample_bytree': 0.8,
'objective': 'multi:softmax', 'min_child_weight': 3, 'max_depth': 7,
'subsample': 0.9, 'learning_rate': 0.1}
# clf = XGBClassifier(learning_rate =0.01, n_estimators=2000, max_depth=6,
# min_child_weight=1, max_delta_step = 5, gamma=0, subsample=0.85, colsample_bytree=0.8,
# objective= 'binary:logistic', nthread=4, scale_pos_weight=1, seed=27, reg_alpha= 0.01)
clf = XGBClassifier(**ind_params)
clf.fit(X_train, y_train, eval_set=[(X_train, y_train), (X_test, y_test)], verbose=True)
# -
from sklearn.metrics import f1_score
y_pred = clf.predict(X_test)
def feature_importance(df, clf):
columns = [x for x in df.columns.tolist() if x != 'y']
t = zip(columns, clf.feature_importances_.tolist())
nonzero_feature_cnt = 0
fi = sorted(t, key=lambda tup: tup[1], reverse=True)
for i in fi:
print(i)
if i[1] > 0:
nonzero_feature_cnt += 1
print("non zero feature cnt: {0}, feature total cnt:{1}".format(nonzero_feature_cnt, len(clf.feature_importances_)))
return fi
feature_importance(df, clf)
f1_score(y_test, y_pred, average='macro')
report = metrics.classification_report(y_test, y_pred)
print(report)
# +
from matplotlib import pyplot
import scikitplot as skplt
skplt.plotters.plot_feature_importances(clf, feature_names=df.columns, figsize=(11.7, 8.27))
plt.show()
# -
X = df.drop(['y'], axis=1)
y = df['y']
predict = clf.predict(X)
report = metrics.classification_report(y, predict)
print(report)
df['predict'] = predict
# +
def profit_long(df, amount=1000):
long, short = None, None
xbt = None
profit = 0
for index, row in df.iterrows():
descision = row['predict']
best_bid = row['bid_p_0']
best_ask = row['ask_p_0']
if descision == 1:
if long is None:
long = best_ask
xbt = amount / long
print('long: price-{0}, vol-{1}'.format(long, row['ask_vol_0']))
elif descision == -1:
if long is not None:
tmp = xbt * best_bid
profit += (tmp - amount)
amount = tmp
print('sell long: price-{0}, vol-{1}, profit: {2}'.format(best_bid, row['bid_vol_0'], profit))
long = None
print("net return: {0}".format(profit))
def profit_short(df, amount=1000):
short = None
xbt = None
profit = 0
for index, row in df.iterrows():
descision = row['predict']
best_bid = row['bid_p_0']
best_ask = row['ask_p_0']
if descision == 1:
if short is not None:
tmp = sxbt * best_ask
profit += (amount - tmp)
amount = tmp
print('sell short: price-{0}, vol-{1}, profit: {2}'.format(best_ask, row['ask_vol_0'], profit))
short = None
elif descision == -1:
if short is None:
short = best_bid
sxbt = amount / short
print('short: price-{0}, vol-{1}'.format(short, row['bid_vol_0']))
print("net return: {0}".format(profit))
# -
X3 = df3.drop(['y'], axis=1)
y3 = df3['y']
predict3 = clf.predict(X3)
report3 = metrics.classification_report(y3, predict3)
print(report3)
profit_long(df, 500)
profit_short(df, 500)
44.820041660407924*30*6
import requests
import json
from sklearn.externals import joblib
clf1 = joblib.load('order_book_flow.joblib')
while True:
res = requests.get('http://localhost:4444/orderBookL2?symbol=XBTUSD')
ob = res.json()
asks = filter(lambda item: item['side'] == 'Sell', ob)
asks = sorted(asks, key=lambda a: a['price'])[:10]
bids = filter(lambda item: item['side'] == 'Buy', ob)
bids = sorted(bids, key=lambda b: b['price'], reverse=True)[:10]
row = {}
for index, val in enumerate(asks):
ind = str(index)
row['ask_p_' + ind] = val['price']
row['ask_vol_' + ind] = val['size']
for index, val in enumerate(bids):
ind = str(index)
row['bid_p_' + ind] = val['price']
row['bid_vol_' + ind] = val['size']
df = pd.DataFrame(row)
df = feature(df)
pred = clf.predict(df)
best_bid = row['bid_p_0']
best_ask = row['ask_p_0']
if descision == 1:
if long is None:
long = best_ask
xbt = amount / long
print('bid: {0}'.format(long))
elif descision == -1:
if long is not None:
tmp = xbt * best_bid
profit += (tmp - amount)
amount = tmp
print('ask: {0}, profit: {1}'.format(best_bid, profit))
long = None
from sklearn.externals import joblib
joblib.dump(clf, 'order_book_flow.joblib')
clf1 = joblib.load('order_book_flow.joblib')
clf1.predict(X.head(1))
|
.ipynb_checkpoints/order_flow_prediction-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Nov05/yelp-dataset-challenge/blob/master/notebooks/2019_12_05_json_to_csv.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="aX5GMld-16H0" colab_type="code" colab={}
# created by nov05 on 2019-12-05
# + id="vUvK72Tc9q-h" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="dEv5nJCu2PdJ" colab_type="code" outputId="0aad866a-1029-4b52-e650-f8dd0555cdc5" colab={"base_uri": "https://localhost:8080/", "height": 35}
# %%file json_to_csv.py
'''
Load Yelp JSON files and spit out CSV files
Does not try to reinvent the wheel and uses pandas json_normalize
Kinda hacky and requires a bit of RAM. But works, albeit naively.
Tested with Yelp JSON files in dataset challenge round 12:
https://www.yelp.com/dataset/challenge
Source Code:
https://gist.github.com/emredjan/6fffc4f696d2201d1e3697b783f9590b
'''
import json
from pathlib import Path
from time import clock
from typing import Dict, List
import click
import pandas as pd
from pandas.io.json import json_normalize
def read_json_as_array(json_file: Path) -> str:
'''
Read a given Yelp JSON file as string, adding opening / closing
brackets and commas to convert from separate JSON objects to
an array of JSON objects, so JSON aware libraries can properly read
Parameters
-----------
json_file: path-like
Returns
-------
json_data: str
String representation of JSON array
'''
json_data = ''
with open(json_file, 'r', encoding='utf-8') as in_file:
for i, line in enumerate(in_file):
if i == 0 and line:
json_data += '[' + line
elif line:
json_data += ',' + line
else:
pass
json_data += ']\n'
return json_data
def load_json(json_data: str) -> pd.DataFrame:
'''
Read and normalize a given JSON array into a pandas DataFrame
Parameters
-----------
json_data: str
String representation of JSON array
Returns
-------
df: pandas.DataFrame
DataFrame containing the normalized JSON data
'''
data = json.loads(json_data)
df = json_normalize(data)
return df
def write_csv(df: pd.DataFrame, out_file: Path) -> None:
'''
Write a given DataFrame to csv without index
Parameters
-----------
df: pandas.DataFrame
DataFrame containing the normalized JSON data
out_file: pathlib.Path
A proper path of CSV file name
'''
df.to_csv(out_file, index=False)
@click.command()
@click.argument('json-dir', type=click.Path(exists=True, dir_okay=True))
def main(json_dir):
'''
Read a given directory containing Yelp JSON data and convert those
files to CSV under 'csv_out' in the same directory
'''
t0 = clock()
json_dir = Path(json_dir)
csv_dir = json_dir / 'csv_out'
csv_dir.mkdir(exist_ok=True)
file_list: List[Path] = json_dir.glob('*.json')
with click.progressbar(file_list, label='Processing files..') as bar:
for file in bar:
csv_file = csv_dir / (file.stem + '.csv')
data = read_json_as_array(file)
df = load_json(data); del data
write_csv(df, csv_file); del [df]
t1 = clock()
mins = (t1 - t0) // 60
secs = int((t1 - t0) % 60)
timing = f'Conversion finished in {mins} minutes and {secs} seconds'
click.secho(timing, fg='green')
if __name__ == '__main__':
main() # pylint: disable=E1120
# + id="6OaTtFjn9L9w" colab_type="code" colab={}
# # !python json_to_csv.py /content/drive/My\ Drive/data/2019-12-06\ yelp/yelp_dataset
# + id="ihMr-KZKE3tH" colab_type="code" outputId="d6b793d8-4ec0-4675-90d1-2adba37d7219" colab={"base_uri": "https://localhost:8080/", "height": 123}
import glob
path = "/content/drive/My Drive/data/2019-12-06 yelp/yelp_dataset"
filenames = glob.glob(path+'/*.json')
for i in range(len(filenames)):
print(filenames[i])
# + id="wAg8z7L8glqT" colab_type="code" colab={}
import json
import pandas as pd
from pandas.io.json import json_normalize
from pathlib import Path
def read_json_as_array(json_file: Path) -> str:
json_data = ''
with open(json_file, 'r', encoding='utf-8') as in_file:
for i, line in enumerate(in_file):
if i == 0 and line:
json_data += '[' + line
elif line:
json_data += ',' + line
else:
pass
json_data += ']\n'
return json_data
def load_json(json_data: str) -> pd.DataFrame:
data = json.loads(json_data); del data
df = json_normalize(data)
return df
def write_csv(df: pd.DataFrame, out_file: Path) -> None:
df.to_csv(out_file, index=False)
def convert_json_to_csv(filename):
path = "/content/drive/My Drive/data/2019-12-06 yelp/yelp_dataset"
filename_json = path + "/" + filename
filename_csv = path + "/csv_out/" + filename[:-5] + ".csv"
data = read_json_as_array(filename_json)
df = load_json(data); del data
write_csv(df, filename_csv); del [df]
# + id="qIxrzC0-kj1q" colab_type="code" colab={}
"""
So there are some interesting numbers.
The size of review.json is about 5GB.
The size of converted CSV file is about 4.39GB.
And using pandas to convert it took about 23GB memory... orz
"""
# convert_json_to_csv("photo.json")
# convert_json_to_csv("review.json")
# convert_json_to_csv("tip.json")
# convert_json_to_csv("user.json")
|
notebooks/2019_12_05_json_to_csv.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: ipynb,.pct.py:percent
# text_representation:
# extension: .py
# format_name: percent
# format_version: '1.3'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %% [markdown]
# # Stochastic Variational Inference for scalability with SVGP
# %% [markdown]
# One of the main criticisms of Gaussian processes is their scalability to large datasets. In this notebook, we illustrate how to use the state-of-the-art Stochastic Variational Gaussian Process (SVGP) (*Hensman, et. al. 2013*) to overcome this problem.
# %%
# %matplotlib inline
import itertools
import numpy as np
import time
import gpflow
import tensorflow as tf
import matplotlib.pyplot as plt
from gpflow.ci_utils import ci_niter
plt.style.use("ggplot")
# for reproducibility of this notebook:
rng = np.random.RandomState(123)
tf.random.set_seed(42)
# %% [markdown]
# ## Generating data
# For this notebook example, we generate 10,000 noisy observations from a test function:
# \begin{equation}
# f(x) = \sin(3\pi x) + 0.3\cos(9\pi x) + \frac{\sin(7 \pi x)}{2}
# \end{equation}
# %%
def func(x):
return np.sin(x * 3 * 3.14) + 0.3 * np.cos(x * 9 * 3.14) + 0.5 * np.sin(x * 7 * 3.14)
N = 10000 # Number of training observations
X = rng.rand(N, 1) * 2 - 1 # X values
Y = func(X) + 0.2 * rng.randn(N, 1) # Noisy Y values
data = (X, Y)
# %% [markdown]
# We plot the data along with the noiseless generating function:
# %%
plt.plot(X, Y, "x", alpha=0.2)
Xt = np.linspace(-1.1, 1.1, 1000)[:, None]
Yt = func(Xt)
_ = plt.plot(Xt, Yt, c="k")
# %% [markdown]
# ## Building the model
# The main idea behind SVGP is to approximate the true GP posterior with a GP conditioned on a small set of "inducing" values. This smaller set can be thought of as summarizing the larger dataset. For this example, we will select a set of 50 inducing locations that are initialized from the training dataset:
# %%
M = 50 # Number of inducing locations
kernel = gpflow.kernels.SquaredExponential()
Z = X[:M, :].copy() # Initialize inducing locations to the first M inputs in the dataset
m = gpflow.models.SVGP(kernel, gpflow.likelihoods.Gaussian(), Z, num_data=N)
# %% [markdown]
# ## Likelihood computation: batch vs. minibatch
# First we showcase the model's performance using the whole dataset to compute the ELBO.
# %%
elbo = tf.function(m.elbo)
# %%
# TensorFlow re-traces & compiles a `tf.function`-wrapped method at *every* call if the arguments are numpy arrays instead of tf.Tensors. Hence:
tensor_data = tuple(map(tf.convert_to_tensor, data))
elbo(tensor_data) # run it once to trace & compile
# %%
# %%timeit
elbo(tensor_data)
# %% [markdown]
# We can speed up this calculation by using minibatches of the data. For this example, we use minibatches of size 100.
# %%
minibatch_size = 100
train_dataset = tf.data.Dataset.from_tensor_slices((X, Y)).repeat().shuffle(N)
train_iter = iter(train_dataset.batch(minibatch_size))
ground_truth = elbo(tensor_data).numpy()
# %%
# %%timeit
elbo(next(train_iter))
# %% [markdown]
# ### Stochastical estimation of ELBO
# The minibatch estimate should be an unbiased estimator of the `ground_truth`. Here we show a histogram of the value from different evaluations, together with its mean and the ground truth. The small difference between the mean of the minibatch estimations and the ground truth shows that the minibatch estimator is working as expected.
# %%
evals = [elbo(minibatch).numpy() for minibatch in itertools.islice(train_iter, 100)]
# %%
plt.hist(evals, label="Minibatch estimations")
plt.axvline(ground_truth, c="k", label="Ground truth")
plt.axvline(np.mean(evals), c="g", ls="--", label="Minibatch mean")
plt.legend()
plt.title("Histogram of ELBO evaluations using minibatches")
print("Discrepancy between ground truth and minibatch estimate:", ground_truth - np.mean(evals))
# %% [markdown]
# ### Minibatches speed up computation
# The reason for using minibatches is that it decreases the time needed to make an optimization step, because estimating the objective is computationally cheaper with fewer data points. Here we plot the change in time required with the size of the minibatch. We see that smaller minibatches result in a cheaper estimate of the objective.
# %%
# Evaluate objective for different minibatch sizes
minibatch_proportions = np.logspace(-2, 0, 10)
times = []
objs = []
for mbp in minibatch_proportions:
batchsize = int(N * mbp)
train_iter = iter(train_dataset.batch(batchsize))
start_time = time.time()
objs.append([elbo(minibatch) for minibatch in itertools.islice(train_iter, 20)])
times.append(time.time() - start_time)
# %%
f, (ax1, ax2) = plt.subplots(1, 2, figsize=(16, 6))
ax1.plot(minibatch_proportions, times, "x-")
ax1.set_xlabel("Minibatch proportion")
ax1.set_ylabel("Time taken")
ax2.plot(minibatch_proportions, np.array(objs), "kx")
ax2.set_xlabel("Minibatch proportion")
ax2.set_ylabel("ELBO estimates")
# %% [markdown]
# ### Running stochastic optimization
# %% [markdown]
# First we create a utility function that plots the model's predictions:
# %%
def plot(title=""):
plt.figure(figsize=(12, 4))
plt.title(title)
pX = np.linspace(-1, 1, 100)[:, None] # Test locations
pY, pYv = m.predict_y(pX) # Predict Y values at test locations
plt.plot(X, Y, "x", label="Training points", alpha=0.2)
(line,) = plt.plot(pX, pY, lw=1.5, label="Mean of predictive posterior")
col = line.get_color()
plt.fill_between(
pX[:, 0],
(pY - 2 * pYv ** 0.5)[:, 0],
(pY + 2 * pYv ** 0.5)[:, 0],
color=col,
alpha=0.6,
lw=1.5,
)
Z = m.inducing_variable.Z.numpy()
plt.plot(Z, np.zeros_like(Z), "k|", mew=2, label="Inducing locations")
plt.legend(loc="lower right")
plot(title="Predictions before training")
# %% [markdown]
# Now we can train our model. For optimizing the ELBO, we use the Adam Optimizer *(Kingma and Ba 2015)* which is designed for stochastic objective functions. We create a `run_adam` utility function to perform the optimization.
# %%
minibatch_size = 100
# We turn off training for inducing point locations
gpflow.set_trainable(m.inducing_variable, False)
def run_adam(model, iterations):
"""
Utility function running the Adam optimizer
:param model: GPflow model
:param interations: number of iterations
"""
# Create an Adam Optimizer action
logf = []
train_iter = iter(train_dataset.batch(minibatch_size))
training_loss = model.training_loss_closure(train_iter, compile=True)
optimizer = tf.optimizers.Adam()
@tf.function
def optimization_step():
optimizer.minimize(training_loss, model.trainable_variables)
for step in range(iterations):
optimization_step()
if step % 10 == 0:
elbo = -training_loss().numpy()
logf.append(elbo)
return logf
# %% [markdown]
# Now we run the optimization loop for 20,000 iterations.
# %%
maxiter = ci_niter(20000)
logf = run_adam(m, maxiter)
plt.plot(np.arange(maxiter)[::10], logf)
plt.xlabel("iteration")
_ = plt.ylabel("ELBO")
# %% [markdown]
# Finally, we plot the model's predictions.
# %%
plot("Predictions after training")
# %% [markdown]
# ## Further reading
#
# Several notebooks expand on this one:
#
# - [Advanced Sparse GP regression](../advanced/advanced_many_points.ipynb), which goes into deeper detail on sparse Gaussian process methods.
# - [Optimization](../advanced/optimisation.ipynb) discussing optimizing GP models.
# - [Natural gradients](../advanced/natural_gradients.ipynb) for optimizing SVGP models efficiently.
# %% [markdown]
# ## References:
# Hensman, James, <NAME>, and <NAME>. "Gaussian processes for big data." Uncertainty in Artificial Intelligence (2013).
#
# Kingma, <NAME>., and <NAME>. "Adam: A method for stochastic optimization." arXiv preprint arXiv:1412.6980 (2014).
|
doc/source/notebooks/advanced/gps_for_big_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Summarizing Data in a Stream
#
# __Requirement__: Count the number of occurrences of "HELLOWORLD" (case sensitive) in a text stream by applying stateful updateStateByKey() on a DStream
#
# __Learning Objective:__
#
# * Summarizing Data in entire Stream using updateStateByKey of DStream
# * See usage of a stateful updateStateByKey operation of DStream
# __Types of Stream Transformations__ =
# * Stateless - Transformations which are applied on single RDD like map(), filter(), reduceByKey()
# * Stateful - Transformations which accumulate across multiple RDDs across a longer time interval (i.e. entire stream or window)
# ### Prerequisites
#
# Run Netcat (a small utility found in most Unix-like systems) as a data server by using: `nc -lc 9999`
#
# For windows: [download netcat](https://joncraton.org/blog/46/netcat-for-windows) and run: `nc -l -p 9999`
# ### Create a StreamingContext and a DStream
#
# <font color="red">TODO: Configure checkpoint path</red>
# +
from pyspark.streaming import StreamingContext
ssc = StreamingContext(sc, 5)
# ssc.checkpoint("file:///tmp/spark")
ssc.checkpoint("D:\\x-temp\\x-temp-spark\\gs-spark-python\\notebooks\\22")
lines = ssc.socketTextStream("localhost", 9999)
# -
# ### Perform Stateful Operation (e.g. updateStateByKey) on DStream and Print
# +
# function to do sum of new value and last global sum
def countWords(newValues, lastSum):
if lastSum is None:
lastSum = 0
return sum(newValues, lastSum)
# Find count of "HELLOWORLD" word in the entire Stream (i.e. multiple RDDs)
counts = lines.flatMap(lambda line: line.split(" "))\
.filter(lambda word:"HELLOWORLD" in word)\
.map(lambda word: (word, 1))\
.updateStateByKey(countWords)
# -
# Print a counts of "HELLOWORLD" words in the entire Stream
counts.pprint()
# ### Start Listening for Streaming Data
# Run "netcat" on localhost:9999 and start entering few lines/words including few occurences of "HELLOWORLD" word
# +
ssc.start() # Start the computation
ssc.awaitTermination() # Wait for the computation to terminate
|
gs-spark/gs-spark-python/notebooks/22/Streaming-UpdateStateByKey.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 安装git
#下载
wget https://www.kernel.org/pub/software/scm/git/git-2.0.0.tar.gz
#然后解压
tar -zvxf git-2.0.0.tar.gz
#进入git解压目录,然后配置、安装:
autoconf
./configure --with-curl=/usr/local
make
sudo make install
#这里之后应该就会安装成功,可以通过下面代码查看版本
git --version
sudo service sshd restart > /dev/null 2>&1
|
notebooks/Untitled.ipynb
|